code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from plotnine import *
train = pd.read_csv('data/train.csv', encoding='euc-kr')
test = pd.read_csv('data/test.csv', encoding='euc-kr')
train.shape
test.shape
train.head()
train.info()
train.describe()
train.isnull().sum()
# Age와 Cabin의 경우 결측치가 많아 처리하여야 합니다.
train['Age_mean'] = train['Age']
test['Age_mean'] = test['Age']
train['Age_mean'].fillna(train['Age'].mean(), inplace=True)
test['Age_mean'].fillna(test['Age'].mean(), inplace=True)
train['Age_mean'].mean()
test['Age_mean'].mean()
# ## One-hot-Encoding
train['Sex'].value_counts()
train['Gender'] = (train['Sex'] == 'female')
train['Gender']
train['Embarked'].value_counts()
train['Embarked'].isnull().sum()
test['Embarked'].value_counts()
test['Embarked'].isnull().sum()
train['Embarked_S'] = train['Embarked'] == "S"
train['Embarked_C'] = train['Embarked'] == "C"
train['Embarked_Q'] = train['Embarked'] == "Q"
train['Embarked_S'].sum()
train['Embarked_C'].head()
train['Embarked_Q'].head()
# boolean 데이터도 사용가능
# int 로 수치형 데이터로 바꾸기 가능
train[['Embarked', 'Embarked_S', 'Embarked_C', 'Embarked_Q']].head()
test['Embarked_S'] = test['Embarked'] == "S"
test['Embarked_C'] = test['Embarked'] == "C"
test['Embarked_Q'] = test['Embarked'] == "Q"
# ## 가족수 구하기
train.columns
train['FamilySize'] = train['SibSp'] + train['Parch'] + 1
# 가족 수에 자신도 포함되게 하여 1명 이상이 되도록 만들어준다.
train['FamilySize'].value_counts
# 혼자 탄 사람이 많다.
train['Family'] = train['FamilySize']
train.loc[train['FamilySize'] == 1, 'Family'] = 'S'
train.loc[(train['FamilySize'] > 1) & (train['FamilySize'] < 5), 'Family'] = 'M'
train.loc[train['FamilySize'] > 4, 'Family'] = 'L'
train[['Family', 'FamilySize']].head()
train['Family_S'] = train['Family'] == "S"
train['Family_M'] = train['Family'] == "M"
train['Family_L'] = train['Family'] == "L"
train[['Family', 'Family_S', 'Family_M', 'Family_L', 'FamilySize']].head()
test['FamilySize'] = test['SibSp'] + test['Parch'] + 1
test['Family'] = test['FamilySize']
test.loc[test['FamilySize'] == 1, 'Family'] = 'S'
test.loc[(test['FamilySize'] >1) & (test['FamilySize'] < 4), 'Family'] = 'M'
test.loc[test['FamilySize'] > 4, 'Family'] = 'L'
test[['Family', 'FamilySize']].head()
test['Family_S'] = test['Family'] == "S"
test['Family_M'] = test['Family'] == "M"
test['Family_L'] = test['Family'] == "L"
test[['Family', 'Family_S', 'Family_M', 'Family_L', 'FamilySize']].head()
# ## feature selection
feature_names = ['Gender', 'Age_mean', 'Embarked_S', 'Embarked_C', 'Embarked_Q', 'Family_S', 'Family_M', 'Family_L']
x_train = train[feature_names]
x_train.head()
y_label = train['Survived']
print(y_label.shape)
y_label.head()
# !pip install graphviz
import os
os.environ["PATH"] += os.pathsep + 'C:/Program Files (x86)/Graphviz2.38/bin/'
# +
from sklearn.tree import export_graphviz
import graphviz
export_graphviz(model,
feature_names=feature_names,
class_names=["Perish", "Survived"],
out_file="decision-tree.dot")
with open("decision-tree.dot") as f:
dot_graph = f.read()
graphviz.Source(dot_graph)
# +
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier(max_depth=3, random_state=2018)
model
# -
model.fit(x_train, y_label)
# prediction = model.predict(x_test)
# +
test['Gender'] = (test['Sex'] == 'female')
x_test = test[feature_names]
prediction = model.predict(x_test)
prediction[:10]
# -
test['Survived'] = prediction
test.columns
submission = test[['PassengerId','Survived']]
submission.head()
# +
submission.to_csv("submission_ML.csv", index=False)
pd.read_csv("submission_ML.csv").head()
# -
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
k_fold = KFold(n_splits=9, shuffle=True, random_state=2018)
scoring = 'accuracy'
score = cross_val_score(model, x_train, y_label, cv=k_fold, n_jobs=-1, scoring=scoring)
print(score)
round(np.mean(score)*100, 2)
prediction = model.predict(x_test)
prediction
| Decision Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:naacl] *
# language: python
# name: conda-env-naacl-py
# ---
# +
# %load_ext autoreload
# %autoreload 2
import torch
import os
import time
import json
import numpy as np
from collections import defaultdict
from speaker import Speaker
from utils import read_vocab,write_vocab,build_vocab,Tokenizer,padding_idx,timeSince, read_img_features, read_graph_features, read_graph_features_parallel
import utils
from env import R2RBatch
from eval import Evaluation
from param import args
from agent import ActiveExplore_v1
import warnings
warnings.filterwarnings("ignore")
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
from tensorboardX import SummaryWriter
print('current directory',os.getcwd())
os.chdir('..')
print('current directory',os.getcwd())
args.name = 'active'
args.attn = 'soft'
args.train = 'listener'
args.featdropout = 0.3
args.angle_feat_size = 128
args.feedback = 'sample'
args.ml_weight = 0.2
args.sub_out = 'max'
args.dropout = 0.5
args.optim = 'adam'
args.lr = 1e-4
args.iters = 80000
args.maxAction = 35
args.batchSize = 64
args.self_train = True
args.aug = 'tasks/R2R/data/aug_paths.json'
# args.aug = 'tasks/R2R/data/aug_paths_unseenvalid.json'
args.speaker = 'snap/speaker/state_dict/best_val_unseen_bleu'
args.featdropout = 0.4
args.iters = 200000
if args.optim == 'rms':
print("Optimizer: Using RMSProp")
args.optimizer = torch.optim.RMSprop
elif args.optim == 'adam':
print("Optimizer: Using Adam")
args.optimizer = torch.optim.Adam
elif args.optim == 'sgd':
print("Optimizer: sgd")
args.optimizer = torch.optim.SGD
elif args.optim == 'adabound':
print("Optimizer: adabound")
args.optimizer = adabound.AdaBound
log_dir = 'snap/%s' % args.name
if not os.path.exists(log_dir):
os.makedirs(log_dir)
TRAIN_VOCAB = 'tasks/R2R/data/train_vocab.txt'
TRAINVAL_VOCAB = 'tasks/R2R/data/trainval_vocab.txt'
IMAGENET_FEATURES = 'img_features/ResNet-152-imagenet.tsv'
PLACE365_FEATURES = 'img_features/ResNet-152-places365.tsv'
if args.features == 'imagenet':
features = IMAGENET_FEATURES
if args.fast_train:
name, ext = os.path.splitext(features)
features = name + "-fast" + ext
feedback_method = args.feedback # teacher or sample
print(args)
def setup():
torch.manual_seed(1)
torch.cuda.manual_seed(1)
# Check for vocabs
if not os.path.exists(TRAIN_VOCAB):
write_vocab(build_vocab(splits=['train']), TRAIN_VOCAB)
if not os.path.exists(TRAINVAL_VOCAB):
write_vocab(build_vocab(splits=['train','val_seen','val_unseen']), TRAINVAL_VOCAB)
#
setup()
vocab = read_vocab(TRAIN_VOCAB)
tok = Tokenizer(vocab=vocab, encoding_length=args.maxInput)
feat_dict = read_img_features(features)
print('start extract keys...')
featurized_scans = set([key.split("_")[0] for key in list(feat_dict.keys())])
print('keys extracted...')
# Load the augmentation data
aug_path = args.aug
# Create the training environment
train_env = R2RBatch(feat_dict, batch_size=args.batchSize,
splits=['train'], tokenizer=tok)
aug_env = R2RBatch(feat_dict, batch_size=args.batchSize,
splits=[aug_path], tokenizer=tok, name='aug')
aug_unseen_env = R2RBatch(feat_dict, batch_size=args.batchSize,
splits=['tasks/R2R/data/aug_paths_unseenvalid.json'], tokenizer=tok, name='aug')
# Printing out the statistics of the dataset
stats = train_env.get_statistics()
print("The training data_size is : %d" % train_env.size())
print("The average instruction length of the dataset is %0.4f." % (stats['length']))
print("The average action length of the dataset is %0.4f." % (stats['path']))
# stats = aug_env.get_statistics()
# print("The augmentation data size is %d" % aug_env.size())
# print("The average instruction length of the dataset is %0.4f." % (stats['length']))
# print("The average action length of the dataset is %0.4f." % (stats['path']))
# Setup the validation data
val_envs = {split: (R2RBatch(feat_dict, batch_size=args.batchSize, splits=[split],
tokenizer=tok), Evaluation([split], featurized_scans, tok))
for split in ['train', 'val_seen', 'val_unseen']}
# +
import imp
import traceback
args.load = 'snap/agent/state_dict/best_val_unseen'
torch.autograd.set_detect_anomaly(True)
def test(train_env, tok, n_iters, log_every=100, val_envs={}, aug_env=None):
writer = SummaryWriter(logdir=log_dir)
listner = ActiveExplore_v1(train_env, "", tok, episode_len=args.maxAction)
speaker = None
if args.self_train:
speaker = Speaker(train_env, listner, tok)
if args.speaker is not None:
print("Load the speaker from %s." % args.speaker)
speaker.load(args.speaker)
start_iter = 0
if args.load is not None:
print("LOAD THE listener from %s" % args.load)
start_iter = listner.load(os.path.join(args.load))
start_iter = 0
ths = np.ones([args.maxAction])*3
ths[0] = 0.456
ths[1] = 0.485
ths[2] = 0.493
ths[3] = 0.584
ths[4] = 0.574
ths[5] = 0.418
ths[6] = 0.675
start = time.time()
best_val = {'val_seen': {"accu": 0., "state":"", 'update':False},
'val_unseen': {"accu": 0., "state":"", 'update':False}}
if args.fast_train:
log_every = 40
try:
loss_str = ""
for env_name, (env, evaluator) in val_envs.items():
listner.env = env
listner.logs = defaultdict(list)
# Get validation loss under the same conditions as training
iters = None if args.fast_train or env_name != 'train' else 20 # 20 * 64 = 1280
# iters = 5
# Get validation distance from goal under test evaluation conditions
listner.test(use_dropout=False, feedback='argmax', iters=iters, train_exp=True, ths=ths)
result = listner.get_results()
score_summary, score_details = evaluator.score(result)
with open('traj.json','w') as fp:
json.dump(result,fp,indent=4)
loss_str += ", %s \n" % env_name
for metric,val in score_summary.items():
loss_str += ', %s: %.3f' % (metric, val)
loss_str += '\n'
print(loss_str)
torch.cuda.empty_cache()
except Exception as e:
del listner
traceback.print_exc()
torch.cuda.empty_cache()
test(train_env, tok, args.iters,log_every=20, val_envs=val_envs, aug_env=[aug_env,aug_unseen_env])
# listener
# -
| evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Defining your own classes
# ### User Defined Types
# A **class** is a user-programmed Python type (since Python 2.2!)
# It can be defined like:
class Room(object):
pass
# Or:
class Room():
pass
# Or:
class Room:
pass
# What's the difference? Before Python 2.2 a class was distinct from all other Python types, which caused some odd behaviour. To fix this, classes were redefined as user programmed types by extending `object`, e.g., class `room(object)`.
#
# So most Python 2 code will use this syntax as very few people want to use old style python classes. Python 3 has formalised this by removing old-style classes, so they can be defined without extending `object`, or indeed without braces.
#
# Just as with other python types, you use the name of the type as a function to make a variable of that type:
zero = int()
type(zero)
myroom = Room()
type(myroom)
# In the jargon, we say that an **object** is an **instance** of a particular **class**.
#
# `__main__` is the name of the scope in which top-level code executes, where we've defined the class `Room`.
# Once we have an object with a type of our own devising, we can add properties at will:
myroom.name = "Living"
myroom.name
# The most common use of a class is to allow us to group data into an object in a way that is
# easier to read and understand than organising data into lists and dictionaries.
myroom.capacity = 3
myroom.occupants = ["Graham", "Eric"]
# ### Methods
# So far, our class doesn't do much!
# We define functions **inside** the definition of a class, in order to give them capabilities, just like the methods on built-in
# types.
class Room:
def overfull(self):
return len(self.occupants) > self.capacity
myroom = Room()
myroom.capacity = 3
myroom.occupants = ["Graham", "Eric"]
myroom.overfull()
myroom.occupants.append(['TerryG'])
myroom.occupants.append(['John'])
myroom.overfull()
# When we write methods, we always write the first function argument as `self`, to refer to the object instance itself,
# the argument that goes "before the dot".
# This is just a convention for this variable name, not a keyword. You could call it something else if you wanted.
# ### Constructors
# Normally, though, we don't want to add data to the class attributes on the fly like that.
# Instead, we define a **constructor** that converts input data into an object.
class Room:
def __init__(self, name, exits, capacity, occupants=[]):
self.name = name
self.occupants = occupants # Note the default argument, occupants start empty
self.exits = exits
self.capacity = capacity
def overfull(self):
return len(self.occupants) > self.capacity
living = Room("Living Room", {'north': 'garden'}, 3)
living.capacity
# Methods which begin and end with **two underscores** in their names fulfil special capabilities in Python, such as
# constructors.
# ### Object-oriented design
# In building a computer system to model a problem, therefore, we often want to make:
#
# * classes for each *kind of thing* in our system
# * methods for each *capability* of that kind
# * properties (defined in a constructor) for each *piece of information describing* that kind
#
# For example, the below program might describe our "Maze of Rooms" system:
# We define a "Maze" class which can hold rooms:
class Maze:
def __init__(self, name):
self.name = name
self.rooms = {}
def add_room(self, room):
room.maze = self # The Room needs to know which Maze it is a part of
self.rooms[room.name] = room
def occupants(self):
return [occupant for room in self.rooms.values()
for occupant in room.occupants.values()]
def wander(self):
"""Move all the people in a random direction"""
for occupant in self.occupants():
occupant.wander()
def describe(self):
for room in self.rooms.values():
room.describe()
def step(self):
self.describe()
print("")
self.wander()
print("")
def simulate(self, steps):
for _ in range(steps):
self.step()
# And a "Room" class with exits, and people:
class Room:
def __init__(self, name, exits, capacity, maze=None):
self.maze = maze
self.name = name
self.occupants = {} # Note the default argument, occupants start empty
self.exits = exits # Should be a dictionary from directions to room names
self.capacity = capacity
def has_space(self):
return len(self.occupants) < self.capacity
def available_exits(self):
return [exit for exit, target in self.exits.items()
if self.maze.rooms[target].has_space()]
def random_valid_exit(self):
import random
if not self.available_exits():
return None
return random.choice(self.available_exits())
def destination(self, exit):
return self.maze.rooms[self.exits[exit]]
def add_occupant(self, occupant):
occupant.room = self # The person needs to know which room it is in
self.occupants[occupant.name] = occupant
def delete_occupant(self, occupant):
del self.occupants[occupant.name]
def describe(self):
if self.occupants:
print(f"{self.name}: " + " ".join(self.occupants.keys()))
# We define a "Person" class for room occupants:
class Person:
def __init__(self, name, room=None):
self.name = name
def use(self, exit):
self.room.delete_occupant(self)
destination = self.room.destination(exit)
destination.add_occupant(self)
print("{some} goes {action} to the {where}".format(some=self.name,
action=exit,
where=destination.name))
def wander(self):
exit = self.room.random_valid_exit()
if exit:
self.use(exit)
# And we use these classes to define our people, rooms, and their relationships:
graham = Person('Graham')
eric = Person('Eric')
terryg = Person('TerryG')
john = Person('John')
living = Room('livingroom', {'outside': 'garden',
'upstairs': 'bedroom', 'north': 'kitchen'}, 2)
kitchen = Room('kitchen', {'south': 'livingroom'}, 1)
garden = Room('garden', {'inside': 'livingroom'}, 3)
bedroom = Room('bedroom', {'jump': 'garden', 'downstairs': 'livingroom'}, 1)
house = Maze('My House')
for room in [living, kitchen, garden, bedroom]:
house.add_room(room)
living.add_occupant(graham)
garden.add_occupant(eric)
garden.add_occupant(terryg)
bedroom.add_occupant(john)
# And we can run a "simulation" of our model:
house.simulate(3)
# ### Object oriented design
# There are many choices for how to design programs to do this. Another choice would be to separately define exits as a different class from rooms. This way,
# we can use arrays instead of dictionaries, but we have to first define all our rooms, then define all our exits.
class Maze:
def __init__(self, name):
self.name = name
self.rooms = []
self.occupants = []
def add_room(self, name, capacity):
result = Room(name, capacity)
self.rooms.append(result)
return result
def add_exit(self, name, source, target, reverse=None):
source.add_exit(name, target)
if reverse:
target.add_exit(reverse, source)
def add_occupant(self, name, room):
self.occupants.append(Person(name, room))
room.occupancy += 1
def wander(self):
"Move all the people in a random direction"
for occupant in self.occupants:
occupant.wander()
def describe(self):
for occupant in self.occupants:
occupant.describe()
def step(self):
house.describe()
print("")
house.wander()
print("")
def simulate(self, steps):
for _ in range(steps):
self.step()
class Room:
def __init__(self, name, capacity):
self.name = name
self.capacity = capacity
self.occupancy = 0
self.exits = []
def has_space(self):
return self.occupancy < self.capacity
def available_exits(self):
return [exit for exit in self.exits if exit.valid()]
def random_valid_exit(self):
import random
if not self.available_exits():
return None
return random.choice(self.available_exits())
def add_exit(self, name, target):
self.exits.append(Exit(name, target))
class Person:
def __init__(self, name, room=None):
self.name = name
self.room = room
def use(self, exit):
self.room.occupancy -= 1
destination = exit.target
destination.occupancy += 1
self.room = destination
print("{some} goes {action} to the {where}".format(some=self.name,
action=exit.name,
where=destination.name))
def wander(self):
exit = self.room.random_valid_exit()
if exit:
self.use(exit)
def describe(self):
print("{who} is in the {where}".format(who=self.name,
where=self.room.name))
class Exit:
def __init__(self, name, target):
self.name = name
self.target = target
def valid(self):
return self.target.has_space()
house = Maze('My New House')
living = house.add_room('livingroom', 2)
bed = house.add_room('bedroom', 1)
garden = house.add_room('garden', 3)
kitchen = house.add_room('kitchen', 1)
house.add_exit('north', living, kitchen, 'south')
house.add_exit('upstairs', living, bed, 'downstairs')
house.add_exit('outside', living, garden, 'inside')
house.add_exit('jump', bed, garden)
house.add_occupant('Graham', living)
house.add_occupant('Eric', garden)
house.add_occupant('TerryJ', bed)
house.add_occupant('John', garden)
house.simulate(3)
# This is a huge topic, about which many books have been written. The differences between these two designs are important, and will have long-term consequences for the project. That is the how we start to think about **software engineering**, as opposed to learning to program, and is an important part of this course.
# ### Exercise: Your own solution
# Compare the two solutions above. Discuss with a partner which you like better, and why. Then, starting from scratch, design your own. What choices did you make that are different from mine?
| ch00python/101Classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python Django
# language: python
# name: django
# ---
# +
# #!pip install cssselect
# -
import requests
import lxml.html
# HTML 소스 코드를 읽어 들입니다.
r = requests.get("http://wikibook.co.kr/python-for-web-scraping/")
html = r.text
# +
#r.text
# +
# HTML을 HtmlElement 객체로 변환합니다.
root = lxml.html.fromstring(html)
# XPath를 사용해서 요소를 추출합니다.
#titleElement = root.xpath('//*[@id="content"]/div[1]/div[2]/ul/li[1]')
# -
titleElement = root.xpath('//*[@id="content"]/div[1]/div[2]/h1')
type(titleElement)
# 리스트의 첫 번째 요소가 가진 텍스트를 출력합니다.
print(titleElement[0].text)
titleElement[0].tag
titleElement[0].attrib
import cssselect
# CSS 선택자를 사용해서 요소를 추출합니다.
linkAs = root.cssselect('#content > div:nth-child(1) > div.col-md-9 > h1')
len(linkAs)
## for 반복문으로 추출한 요소의 href 속성을 추출합니다.
for linkA in linkAs:
print(linkA.attrib["class"])
| Web_Crawling/python-crawler/chapter_4/html.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup
import pandas as pd
# Windows users
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)# Visit the Quotes to Scrape site
# Visit the mars nasa news site
url = 'https://mars.nasa.gov/news/'
browser.visit(url)
# Optional delay for loading the page
browser.is_element_present_by_css("ul.item_list li.slide", wait_time=1)
html = browser.html
news_soup = BeautifulSoup(html, 'html.parser')
slide_elem = news_soup.select_one('ul.item_list li.slide')
print(slide_elem)
slide_elem.find("div", class_='content_title')
# Use the parent element to find the first `a` tag and save it as `news_title`
news_title = slide_elem.find("div", class_='content_title').get_text()
news_title
# Use the parent element to find the paragraph text
news_p = slide_elem.find('div', class_="article_teaser_body").get_text()
news_p
# ### Featured Images
# Visit URL
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(url)
# Find and click the full image button
full_image_elem = browser.find_by_id('full_image')
full_image_elem.click()
# Find the more info button and click that
browser.is_element_present_by_text('more info', wait_time=1)
more_info_elem = browser.links.find_by_partial_text('more info')
more_info_elem.click()
# Parse the resulting html with soup
html = browser.html
img_soup = soup(html, 'html.parser')
# Find the relative image url
img_url_rel = img_soup.select_one('figure.lede a img').get("src")
img_url_rel
# Use the base URL to create an absolute URL
img_url = f'https://www.jpl.nasa.gov{img_url_rel}'
img_url
# tables into pandas
# pull only first table it finds
df = pd.read_html('http://space-facts.com/mars/')[0]
df.columns=['description', 'value']
df.set_index('description', inplace=True)
df
browser.quit()
# Import Splinter and BeautifulSoup
from splinter import Browser
from bs4 import BeautifulSoup
import pandas as pd
# Windows users
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)# Visit the Quotes to Scrape site
# Visit the mars nasa news site
url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url)
# Optional delay for loading the page
browser.is_element_present_by_css("div.collapsible.results", wait_time=1)
html = browser.html
news_soup = BeautifulSoup(html, 'html.parser')
news_soup
results = news_soup.find('div', class_='results')
results
all_hems = news_soup.find_all('div', class_='item')
all_hems
len(all_hems)
# +
info = []
for hem in all_hems:
d = {}
d["title"] = hem.h3.text
d["img_url"] = f"https://astrogeology.usgs.gov{hem.a['href']}"
d["thumb"] = f"https://astrogeology.usgs.gov{hem.img['src']}"
d["description"] = hem.p.text
info.append(d)
# d["thumb"] = hem.img['src']
# d["description"] = hem.p.text
# print(hem.h3.text)
# print(hem.a['href'])
# print(hem.img['src'])
# print(hem.img['alt'])
# print(hem.p.text)
info
# -
len(info)
for item in info:
print(item)
browser.quit()
import pandas as pd
df = pd.DataFrame(info)
df
df['img_url']
print(info[0]['img_url'])
print(info[1]['img_url'])
res = [sub['img_url'] for sub in info]
res
df.to_html(classes="table table-hemisphere")
# +
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
print(f'Opening URL:{url}')
browser.visit(url)
# Find and click the full image button
full_image_elem = browser.find_by_id('full_image')
full_image_elem.click()
# Find the more info button and click that
browser.is_element_present_by_text('more info', wait_time=1)
more_info_elem = browser.find_link_by_partial_text('more info')
more_info_elem.click()
# Parse the resulting html with soup
html = browser.html
img_soup = soup(html, 'html.parser')
try:
# find the relative image url
img_url_rel = img_soup.select_one('figure.lede a img').get("src")
except AttributeError:
print("Error")
# Use the base url to create an absolute url
img_url = f'https://www.jpl.nasa.gov{img_url_rel}'
img_url
# -
| Mission_to_Mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Carving Unit Tests
#
# So far, we have always generated _system input_, i.e. data that the program as a whole obtains via its input channels. If we are interested in testing only a small set of functions, having to go through the system can be very inefficient. This chapter introduces a technique known as _carving_, which, given a system test, automatically extracts a set of _unit tests_ that replicate the calls seen during the unit test. The key idea is to _record_ such calls such that we can _replay_ them later – as a whole or selectively.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Prerequisites**
#
# * Carving makes use of dynamic traces of function calls and variables, as introduced in the [chapter on configuration fuzzing](ConfigurationFuzzer.ipynb).
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## System Tests vs Unit Tests
#
# Remember the URL grammar introduced for [grammar fuzzing](Grammars.ipynb)? With such a grammar, we can happily test a Web browser again and again, checking how it reacts to arbitrary page requests.
#
# Let us define a very simple "web browser" that goes and downloads the content given by the URL.
# -
import urllib.parse
import urllib.request
import fuzzingbook_utils
def webbrowser(url):
"""Download the http/https resource given by the URL"""
response = urllib.request.urlopen(url)
if response.getcode() == 200:
contents = response.read()
return contents.decode("utf8")
# Let us apply this on [fuzzingboook.org](https://www.fuzzingbook.org/) and measure the time, using the [Timer class](Timer.ipynb):
from Timer import Timer
# +
with Timer() as webbrowser_timer:
fuzzingbook_contents = webbrowser(
"http://www.fuzzingbook.org/html/Fuzzer.html")
print("Downloaded %d bytes in %.2f seconds" %
(len(fuzzingbook_contents), webbrowser_timer.elapsed_time()))
# -
fuzzingbook_contents[:100]
# A full webbrowser, of course, would also render the HTML content. We can achieve this using these commands (but we don't, as we do not want to replicate the entire Web page here):
#
# ```python
# from IPython.core.display import HTML, display
# HTML(fuzzingbook_contents)
# ```
# Having to start a whole browser (or having it render a Web page) again and again means lots of overhead, though – in particular if we want to test only a subset of its functionality. In particular, after a change in the code, we would prefer to test only the subset of functions that is affected by the change, rather than running the well-tested functions again and again.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let us assume we change the function that takes care of parsing the given URL and decomposing it into the individual elements – the scheme ("http"), the network location (`"www.fuzzingbook.com"`), or the path (`"/html/Fuzzer.html"`). This function is named `urlparse()`:
# -
from urllib.parse import urlparse
urlparse('https://www.fuzzingbook.com/html/Carver.html')
# You see how the individual elements of the URL – the _scheme_ (`"http"`), the _network location_ (`"www.fuzzingbook.com"`), or the path (`"//html/Carver.html"`) are all properly identified. Other elements (like `params`, `query`, or `fragment`) are empty, because they were not part of our input.
# The interesting thing is that executing only `urlparse()` is orders of magnitude faster than running all of `webbrowser()`. Let us measure the factor:
# +
runs = 1000
with Timer() as urlparse_timer:
for i in range(runs):
urlparse('https://www.fuzzingbook.com/html/Carver.html')
avg_urlparse_time = urlparse_timer.elapsed_time() / 1000
avg_urlparse_time
# -
# Compare this to the time required by the webbrowser
webbrowser_timer.elapsed_time()
# The difference in time is huge:
webbrowser_timer.elapsed_time() / avg_urlparse_time
# Hence, in the time it takes to run `webbrowser()` once, we can have _tens of thousands_ of executions of `urlparse()` – and this does not even take into account the time it takes the browser to render the downloaded HTML, to run the included scripts, and whatever else happens when a Web page is loaded. Hence, strategies that allow us to test at the _unit_ level are very promising as they can save lots of overhead.
# ## Carving Unit Tests
#
# Testing methods and functions at the unit level requires a very good understanding of the individual units to be tested as well as their interplay with other units. Setting up an appropriate infrastructure and writing unit tests by hand thus is demanding, yet rewarding. There is, however, an interesting alternative to writing unit tests by hand. The technique of _carving_ automatically _converts system tests into unit tests_ by means of recording and replaying function calls:
#
# 1. During a system test (given or generated), we _record_ all calls into a function, including all arguments and other variables the function reads.
# 2. From these, we synthesize a self-contained _unit test_ that reconstructs the function call with all arguments.
# 3. This unit test can be executed (replayed) at any time with high efficiency.
#
# In the remainder of this chapter, let us explore these steps.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Recording Calls
#
# Our first challenge is to record function calls together with their arguments. (In the interest of simplicity, we restrict ourself to arguments, ignoring any global variables or other non-arguments that are read by the function.) To record calls and arguments, we use the mechanism [we introduced for coverage](Coverage.ipynb): By setting up a tracer function, we track all calls into individual functions, also saving their arguments. Just like `Coverage` objects, we want to use `Carver` objects to be able to be used in conjunction with the `with` statement, such that we can trace a particular code block:
#
# ```python
# with Carver() as carver:
# function_to_be_traced()
# c = carver.calls()
# ```
#
# The initial definition supports this construct:
# -
import sys
class Carver(object):
def __init__(self, log=False):
self._log = log
self.reset()
def reset(self):
self._calls = {}
# Start of `with` block
def __enter__(self):
self.original_trace_function = sys.gettrace()
sys.settrace(self.traceit)
return self
# End of `with` block
def __exit__(self, exc_type, exc_value, tb):
sys.settrace(self.original_trace_function)
# The actual work takes place in the `traceit()` method, which records all calls in the `_calls` attribute. First, we define two helper functions:
import inspect
def get_qualified_name(code):
"""Return the fully qualified name of the current function"""
name = code.co_name
module = inspect.getmodule(code)
if module is not None:
name = module.__name__ + "." + name
return name
def get_arguments(frame):
"""Return call arguments in the given frame"""
# When called, all arguments are local variables
arguments = [(var, frame.f_locals[var]) for var in frame.f_locals]
arguments.reverse() # Want same order as call
return arguments
class CallCarver(Carver):
def add_call(self, function_name, arguments):
"""Add given call to list of calls"""
if function_name not in self._calls:
self._calls[function_name] = []
self._calls[function_name].append(arguments)
# Tracking function: Record all calls and all args
def traceit(self, frame, event, arg):
if event != "call":
return None
code = frame.f_code
function_name = code.co_name
qualified_name = get_qualified_name(code)
arguments = get_arguments(frame)
self.add_call(function_name, arguments)
if qualified_name != function_name:
self.add_call(qualified_name, arguments)
if self._log:
print(simple_call_string(function_name, arguments))
return None
# Finally, we need some convenience functions to access the calls:
class CallCarver(CallCarver):
def calls(self):
"""Return a dictionary of all calls traced."""
return self._calls
def arguments(self, function_name):
"""Return a list of all arguments of the given function
as (VAR, VALUE) pairs.
Raises an exception if the function was not traced."""
return self._calls[function_name]
def called_functions(self, qualified=False):
"""Return all functions called."""
if qualified:
return [function_name for function_name in self._calls.keys()
if function_name.find('.') >= 0]
else:
return [function_name for function_name in self._calls.keys()
if function_name.find('.') < 0]
# ### Recording my_sqrt()
# Let's try out our new `Carver` class – first on a very simple function:
from Intro_Testing import my_sqrt
with CallCarver() as sqrt_carver:
my_sqrt(2)
my_sqrt(4)
# We can retrieve all calls seen...
sqrt_carver.calls()
sqrt_carver.called_functions()
# ... as well as the arguments of a particular function:
sqrt_carver.arguments("my_sqrt")
# We define a convenience function for nicer printing of these lists:
def simple_call_string(function_name, argument_list):
"""Return function_name(arg[0], arg[1], ...) as a string"""
return function_name + "(" + \
", ".join([var + "=" + repr(value)
for (var, value) in argument_list]) + ")"
for function_name in sqrt_carver.called_functions():
for argument_list in sqrt_carver.arguments(function_name):
print(simple_call_string(function_name, argument_list))
# This is a syntax we can directly use to invoke `my_sqrt()` again:
eval("my_sqrt(x=2)")
# ### Carving urlparse()
# What happens if we apply this to `webbrowser()`?
with CallCarver() as webbrowser_carver:
webbrowser("http://www.example.com")
# We see that retrieving a URL from the Web requires quite some functionality:
print(webbrowser_carver.called_functions(qualified=True))
# Among several other functions, we also have a call to `urlparse()`:
urlparse_argument_list = webbrowser_carver.arguments("urllib.parse.urlparse")
urlparse_argument_list
# Again, we can convert this into a well-formatted call:
urlparse_call = simple_call_string("urlparse", urlparse_argument_list[0])
urlparse_call
# Again, we can re-execute this call:
eval(urlparse_call)
# We now have successfully carved the call to `urlparse()` out of the `webbrowser()` execution.
# ## Replaying Calls
# Replaying calls in their entirety and in all generality is tricky, as there are several challenges to be addressed. These include:
#
# 1. We need to be able to _access_ individual functions. If we access a function by name, the name must be in scope. If the name is not visible (for instance, because it is a name internal to the module), we must make it visible.
#
# 2. Any _resources_ accessed outside of arguments must be recorded and reconstructed for replay as well. This can be difficult if variables refer to external resources such as files or network resources.
#
# 3. _Complex objects_ must be reconstructed as well.
#
# These constraints make carving hard or even impossible if the function to be tested interacts heavily with its environment. To illustrate these issues, consider the `email.parser.parse()` method that is invoked in `webbrowser()`:
email_parse_argument_list = webbrowser_carver.arguments("email.parser.parse")
# Calls to this method look like this:
email_parse_call = simple_call_string(
"email.parser.parse",
email_parse_argument_list[0])
email_parse_call
# We see that `email.parser.parse()` is part of a `email.parser.Parser` object and it gets a `StringIO` object. Both are non-primitive values. How could we possibly reconstruct them?
# ### Serializing Objects
#
# The answer to the problem of complex objects lies in creating a _persistent_ representation that can be _reconstructed_ at later points in time. This process is known as _serialization_; in Python, it is also known as _pickling_. The `pickle` module provides means to create a serialized representation of an object. Let us apply this on the `email.parser.Parser` object we just found:
import pickle
parser_object = email_parse_argument_list[0][0][1]
parser_object
pickled = pickle.dumps(parser_object)
pickled
# From this string representing the serialized `email.parser.Parser` object, we can recreate the Parser object at any time:
unpickled_parser_object = pickle.loads(pickled)
unpickled_parser_object
# The serialization mechanism allows us to produce a representation for all objects passed as parameters (assuming they can be pickled, that is). We can now extend the `simple_call_string()` function such that it automatically pickles objects. Additionally, we set it up such that if the first parameter is named `self` (i.e., it is a class method), we make it a method of the `self` object.
def call_value(value):
value_as_string = repr(value)
if value_as_string.find('<') >= 0:
# Complex object
value_as_string = "pickle.loads(" + repr(pickle.dumps(value)) + ")"
return value_as_string
def call_string(function_name, argument_list):
"""Return function_name(arg[0], arg[1], ...) as a string, pickling complex objects"""
if len(argument_list) > 0:
(first_var, first_value) = argument_list[0]
if first_var == "self":
# Make this a method call
method_name = function_name.split(".")[-1]
function_name = call_value(first_value) + "." + method_name
argument_list = argument_list[1:]
return function_name + "(" + \
", ".join([var + "=" + call_value(value)
for (var, value) in argument_list]) + ")"
# Let us apply the extended `call_string()` method to create a call for `email.parser.parse()`, including pickled objects:
call = call_string("email.parser.parse", email_parse_argument_list[0])
print(call)
# With this call involvimng the pickled object, we can now re-run the original call and obtain a valid result:
eval(call)
# ### All Calls
#
# So far, we have seen only one call of `webbrowser()`. How many of the calls within `webbrowser()` can we actually carve and replay? Let us try this out and compute the numbers.
import traceback
import enum
import socket
all_functions = set(webbrowser_carver.called_functions(qualified=True))
call_success = set()
run_success = set()
# +
exceptions_seen = set()
for function_name in webbrowser_carver.called_functions(qualified=True):
for argument_list in webbrowser_carver.arguments(function_name):
try:
call = call_string(function_name, argument_list)
call_success.add(function_name)
result = eval(call)
run_success.add(function_name)
except Exception as exc:
exceptions_seen.add(repr(exc))
# print("->", call, file=sys.stderr)
# traceback.print_exc()
# print("", file=sys.stderr)
continue
# -
print("%d/%d calls (%.2f%%) successfully created and %d/%d calls (%.2f%%) successfully ran" % (
len(call_success), len(all_functions), len(
call_success) * 100 / len(all_functions),
len(run_success), len(all_functions), len(run_success) * 100 / len(all_functions)))
# About half of the calls succeed. Let us take a look into some of the error messages we get:
for i in range(10):
print(list(exceptions_seen)[i])
# We see that:
#
# * **A large majority of calls could be converted into call strings.** If this is not the case, this is mostly due to having unserialized objects being passed.
# * **About half of the calls could be executed.** The error messages for the failing runs are varied; the most frequent being that some internal name is invoked that is not in scope.
# Our carving mechanism should be taken with a grain of salt: We still do not cover the situation where external variables and values (such as global variables) are being accessed, and the serialization mechanism cannot recreate external resources. Still, if the function of interest falls among those that _can_ be carved and replayed, we can very effectively re-run its calls with their original arguments.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# * _Carving_ allows for effective replay of function calls recorded during a system test
# * A function call can be _orders of magnitude faster_ than a system invocation.
# * _Serialization_ allows to create persistent representations of complex objects.
# * Functions that heavily interact with their environment and/or access external resources are difficult to carve.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Next Steps
#
# The following chapters make use of the concepts defined here:
#
# * In the chapter on [fuzzing APIs](APIFuzzer.ipynb), we discuss how to use carving to _fuzz functions with combinations of carved and newly generated values_. This effectively joins the strengths of carving and fuzzing.
# -
# ## Background
#
# Carving was invented by Elbaum et al. \cite{Elbaum2006} and originally implemented for Java. In this chapter, we follow several of their design choices (including recording and serializing method arguments only).
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Exercises
#
# ### Exercise 1: Carving for Regression Testing
#
# So far, during carving, we only have looked into reproducing _calls_, but not into actually checking the _results_ of these calls. This is important for _regression testing_ – i.e. checking whether a change to code does not impede existing functionality. We can build this by recording not only _calls_, but also _return values_ – and then later compare whether the same calls result in the same values. This may not work on all occasions; values that depend on time, randomness, or other external factors may be different. Still, for functionality that abstracts from these details, checking that nothing has changed is an important part of testing.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# Our aim is to design a class `ResultCarver` that extends `CallCarver` by recording both calls and return values.
#
# In a first step, create a `traceit()` method that also tracks return values by extending the `traceit()` method. The `traceit()` event type is `"return"` and the `arg` parameter is the returned value. Here is a prototype that only prints out the returned values:
# -
class ResultCarver(CallCarver):
def traceit(self, frame, event, arg):
if event == "return":
if self._log:
print("Result:", arg)
super().traceit(frame, event, arg)
# Need to return traceit function such that it is invoked for return
# events
return self.traceit
with ResultCarver(log=True) as result_carver:
my_sqrt(2)
# + [markdown] solution2="hidden" solution2_first=true
# #### Part 1: Store function results
#
# Extend the above code such that results are _stored_ in a way that associates them with the currently returning function (or method). To this end, you need to keep track of the _current stack of called functions_.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Here's a solution, building on the above:
# + slideshow={"slide_type": "skip"} solution2="hidden"
class ResultCarver(CallCarver):
def reset(self):
super().reset()
self._call_stack = []
self._results = {}
def add_result(self, function_name, arguments, result):
key = simple_call_string(function_name, arguments)
self._results[key] = result
def traceit(self, frame, event, arg):
if event == "call":
code = frame.f_code
function_name = code.co_name
qualified_name = get_qualified_name(code)
self._call_stack.append(
(function_name, qualified_name, get_arguments(frame)))
if event == "return":
result = arg
(function_name, qualified_name, arguments) = self._call_stack.pop()
self.add_result(function_name, arguments, result)
if function_name != qualified_name:
self.add_result(qualified_name, arguments, result)
if self._log:
print(
simple_call_string(
function_name,
arguments),
"=",
result)
# Keep on processing current calls
super().traceit(frame, event, arg)
# Need to return traceit function such that it is invoked for return
# events
return self.traceit
# + slideshow={"slide_type": "skip"} solution2="hidden"
with ResultCarver(log=True) as result_carver:
my_sqrt(2)
result_carver._results
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# #### Part 2: Access results
#
# Give it a method `result()` that returns the value recorded for that particular function name and result:
#
# ```python
# class ResultCarver(CallCarver):
# def result(self, function_name, argument):
# """Returns the result recorded for function_name(argument"""
# ```
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** This is mostly done in the code for part 1:
# + slideshow={"slide_type": "skip"} solution2="hidden"
class ResultCarver(ResultCarver):
def result(self, function_name, argument):
key = simple_call_string(function_name, arguments)
return self._results[key]
# + [markdown] solution2="hidden" solution2_first=true
# #### Part 3: Produce assertions
#
# For the functions called during `webbrowser()` execution, create a set of _assertions_ that check whether the result returned is still the same. Test this for `urllib.parse.urlparse()` and `urllib.parse.urlsplit()`.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Not too hard now:
# + slideshow={"slide_type": "skip"} solution2="hidden"
with ResultCarver() as webbrowser_result_carver:
webbrowser("http://www.example.com")
# + slideshow={"slide_type": "skip"} solution2="hidden"
for function_name in ["urllib.parse.urlparse", "urllib.parse.urlsplit"]:
for arguments in webbrowser_result_carver.arguments(function_name):
try:
call = call_string(function_name, arguments)
result = webbrowser_result_carver.result(function_name, arguments)
print("assert", call, "==", call_value(result))
except Exception:
continue
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We can run these assertions:
# + slideshow={"slide_type": "skip"} solution2="hidden"
from urllib.parse import SplitResult, ParseResult, urlparse, urlsplit
# + slideshow={"slide_type": "skip"} solution2="hidden"
assert urlparse(
url='http://www.example.com',
scheme='',
allow_fragments=True) == ParseResult(
scheme='http',
netloc='www.example.com',
path='',
params='',
query='',
fragment='')
assert urlsplit(
url='http://www.example.com',
scheme='',
allow_fragments=True) == SplitResult(
scheme='http',
netloc='www.example.com',
path='',
query='',
fragment='')
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We can now add these carved tests to a _regression test suite_ which would be run after every change to ensure that the functionality of `urlparse()` and `urlsplit()` is not changed.
| notebooks/Carver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Interactive Proteus parallel example
#
# First go to your notebook Home page, select the mpi cluster, and start some tasks. It should look like this: 
#
#
#
# ## Load IPython support for working with MPI tasks
import ipyparallel
import os
rc = ipyparallel.Client(profile="mpi")
view = rc[:]
view.apply(os.chdir, os.getcwd())
print view
# ## Load interactive Proteus module, physics, and numerics on the MPI tasks and solve problem
# +
# %%px --block
from proteus.iproteus import *
from proteus import default_n as n
from proteus import default_p as p
from proteus import default_s,default_so
from proteus.TransportCoefficients import *
class LAD(TC_base):
"""
The coefficients of the linear advection-diffusion equation
"""
def __init__(self,M,A,B):
self.nd=2
TC_base.__init__(self,
nc=1, #number of components
variableNames=['u'],
mass = {0:{0:'linear'}},
advection = {0:{0:'linear'}},
diffusion = {0:{0:{0:'constant'}}},
potential = {0:{0:'u'}},
reaction = {0:{0:'linear'}})
self.M=M;
self.A=A;
self.B=B;
def evaluate(self,t,c):
c[('m',0)][:] = self.M*c[('u',0)]
c[('dm',0,0)][:] = self.M
c[('f',0)][...,0] = self.B[0]*c[('u',0)]
c[('f',0)][...,1] = self.B[1]*c[('u',0)]
c[('df',0,0)][...,0] = self.B[0]
c[('df',0,0)][...,1] = self.B[1]
c[('a',0,0)][...,0,0] = self.A[0][0]
c[('a',0,0)][...,1,1] = self.A[1][1]
#physics
p.name = "ladr_2d"
p.nd = 2; #Two dimensions
p.L=(1.0,1.0,1.0);
p.T=1.0
p.coefficients=LAD(M=1.0,
A=[[0.001,0.0],
[0.0,0.001]],
B=[2.0,1.0])
def getDBC(x,flag):
if x[0] == 0.0 or x[1] == 0.0:
return lambda x,t: 1.0
elif x[0] == p.L[0] or x[1] == p.L[1]:
return lambda x,t: 0.0
else:
return None
p.dirichletConditions = {0:getDBC}
p.advectiveFluxBoundaryConditions = {}
p.diffusiveFluxBoundaryConditions = {0:{}}
p.periodicDirichletConditions = None
class IC:
def __init__(self):
pass
def uOfXT(self,x,t):
if x[0] <= 0.0 or x[1] <= 0.0:
return 1.0
else:
return 0.0
p.initialConditions = {0:IC()}
#numerics
n.timeIntegration = n.BackwardEuler_cfl
n.stepController = n.Min_dt_cfl_controller
n.runCFL=1.0
n.femSpaces = {0:n.C0_AffineLinearOnSimplexWithNodalBasis}
n.elementQuadrature = n.SimplexGaussQuadrature(p.nd,3)
n.elementBoundaryQuadrature = n.SimplexGaussQuadrature(p.nd-1,3)
n.subgridError = n.AdvectionDiffusionReaction_ASGS(p.coefficients,p.nd,lag=False)
n.shockCapturing = n.ResGradQuad_SC(p.coefficients,p.nd,
shockCapturingFactor=0.99,
lag=True)
n.numericalFluxType = n.Advection_DiagonalUpwind_Diffusion_SIPG_exterior
n.nnx=41; n.nny=41
n.tnList=[float(i)/40.0 for i in range(11)]
n.matrix = n.SparseMatrix
n.multilevelLinearSolver = n.KSP_petsc4py
n.linearSmoother = None#n.Jacobi
n.l_atol_res = 1.0e-8
n.parallelPartitioningType = n.MeshParallelPartitioningTypes.node
n.nLayersOfOverlapForParallel = 0
n.periodicDirichletConditions = None
from proteus import Comm
comm = Comm.get()
pList = [p]
nList = [n]
so = default_so
so.name = pList[0].name = "ladr_2d"
so.sList=[default_s]
so.tnList = n.tnList
nList[0].multilevelLinearSolver=default_n.KSP_petsc4py
ns = NumericalSolution.NS_base(so,pList,nList,so.sList,opts)
ns.calculateSolution('run1')
x = ns.modelList[0].levelModelList[-1].mesh.nodeArray[:,0]
y = ns.modelList[0].levelModelList[-1].mesh.nodeArray[:,1]
triangles = ns.modelList[0].levelModelList[-1].mesh.elementNodesArray
u = ns.modelList[0].levelModelList[-1].u[0].dof
n = len(x)
# -
# ## Combine subdomain solutions and plot
from parplot import parallel_plot
import numpy as np
r = ipyparallel.Reference
png = parallel_plot(view, 'tricontourf',
r('x'),
r('y'),
r('triangles'),
r('u'),
levels=np.linspace(0,1.01,6),
xlim=(0,1),
ylim=(0,1),
)
| notebooks/Tutorials/Tutorial_3/adr-complete-parallel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This is based on the tutorial from
# https://data-flair.training/blogs/advanced-python-project-detecting-fake-news/
import numpy as np
import pandas as pd
import itertools
import pickle
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
# +
df = pd.read_csv('news.csv')
df.shape
df.head()
# -
labels = df.label
labels.head()
x_train, x_test, y_train, y_test = train_test_split(df['text'], labels, test_size=0.2, random_state = 7)
# +
# initialize a tfidfvectorizer
tfidf_vectorizer=TfidfVectorizer(stop_words='english', max_df=0.7)
tfidf_train = tfidf_vectorizer.fit_transform(x_train)
tfidf_test = tfidf_vectorizer.transform(x_test)
# +
pac=PassiveAggressiveClassifier(max_iter=50)
pac.fit(tfidf_train, y_train)
y_pred=pac.predict(tfidf_test)
score=accuracy_score(y_test, y_pred)
print(f'Accuarcy: {round(score*100, 2)}%')
# -
confusion_matrix(y_test,y_pred, labels=['FAKE','REAL'])
model_file = 'model.pkl'
pickle.dump(pac, open())
| Detector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mds572]
# language: python
# name: conda-env-mds572-py
# ---
# 
# # Lab 2: Stochastic Gradient Descent & Pytorch
#
# **<NAME>, January 2021**
# 
# ## Table of Contents
# <hr>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Instructions" data-toc-modified-id="Instructions-2">Instructions</a></span></li><li><span><a href="#Imports" data-toc-modified-id="Imports-3">Imports</a></span></li><li><span><a href="#Exercise-1:-Stochastic-Gradient-Descent" data-toc-modified-id="Exercise-1:-Stochastic-Gradient-Descent-4">Exercise 1: Stochastic Gradient Descent</a></span></li><li><span><a href="#Exercise-2:-SGDClassifier-and-SGDRegresor" data-toc-modified-id="Exercise-2:-SGDClassifier-and-SGDRegresor-5">Exercise 2: <code>SGDClassifier</code> and <code>SGDRegresor</code></a></span></li><li><span><a href="#Exercise-3:-Neural-Networks-"By-Hand"" data-toc-modified-id="Exercise-3:-Neural-Networks-"By-Hand"-6">Exercise 3: Neural Networks "By Hand"</a></span></li><li><span><a href="#Exercise-4:-Predicting-Fashion" data-toc-modified-id="Exercise-4:-Predicting-Fashion-7">Exercise 4: Predicting Fashion</a></span></li><li><span><a href="#(Optional)-Exercise-5:-Implementing-Adam-Optimization-From-Scratch" data-toc-modified-id="(Optional)-Exercise-5:-Implementing-Adam-Optimization-From-Scratch-8">(Optional) Exercise 5: Implementing Adam Optimization From Scratch</a></span></li><li><span><a href="#(Optional)-Exercise-6:-Gif-or-Jiff" data-toc-modified-id="(Optional)-Exercise-6:-Gif-or-Jiff-9">(Optional) Exercise 6: Gif or Jiff</a></span></li><li><span><a href="#Submit-to-Canvas-and-GitHub" data-toc-modified-id="Submit-to-Canvas-and-GitHub-10">Submit to Canvas and GitHub</a></span></li></ul></div>
# ## Instructions
# <hr>
#
# rubric={mechanics:3}
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-4aecf0223bc592cc", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Link to your GitHub repository:**
#
# You will receive marks for correctly submitting this assignment. To submit this assignment you should:
#
# 1. Push your assignment to your GitHub repository!
# 2. Provide a link to your repository in the space provided above.
# 2. Upload a HTML render of your assignment to Canvas. The last cell of this notebook will help you do that.
# 3. Be sure to follow the [General Lab Instructions](https://ubc-mds.github.io/resources_pages/general_lab_instructions/). You can view a description of the different rubrics used for grading in MDS [here](https://github.com/UBC-MDS/public/tree/master/rubric).
#
# Here's a break down of the required and optional exercises in this lab:
#
# | | Number of Exercises | Points |
# |:-------:|:-------------------:|:------:|
# | Required| 13 | 35 |
# | Optional| 2 | 2 |
# -
# ## Imports
# <hr>
import numpy as np
import pandas as pd
import time
import torch
from torch import nn
from torchvision import datasets, transforms
from sklearn.datasets import load_boston
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LinearRegression, LogisticRegression, SGDClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from canvasutils.submit import submit, convert_notebook
import matplotlib.pyplot as plt
plt.style.use('ggplot')
plt.rcParams.update({'font.size': 16, 'figure.figsize': (5,5), 'axes.grid': False})
# ## Exercise 1: Stochastic Gradient Descent
# <hr>
# Below is a super-streamlined Python function that performs gradient descent, it's much like the code you saw in Lectures and in Lab 1. I just made a few small changes:
#
# - Removed the docstring and `print` statements for brevity.
# - The algorithm stops after `n_iters`.
#
# I also include functions to calculate the MSE and gradient of MSE for a linear model $\hat{y}=w^Tx$.
# +
def gradient_descent(f, f_grad, w0, X, y, n_iters=1000, α=0.001):
w = w0
for _ in range(n_iters):
w = w - α * f_grad(w, X, y)
return w
def mse(w, X, y):
return np.mean((X @ w - y) ** 2)
def mse_grad(w, X, y):
return X.T @ (X @ w) - X.T @ y
# -
# The code works fine, although it is quite slow as per usual for vanilla gradient descent. For example, here we do ordinary least squares linear regression on the boston house-prices dataset from sklearn:
X, y = load_boston(return_X_y=True)
X = MinMaxScaler().fit_transform(X)
start = time.time()
w0 = np.zeros(X.shape[1])
w_gd = gradient_descent(mse, mse_grad, w0, X, y, n_iters=10**5)
print(f"Fitting time = {time.time() - start:.4f}s")
print("Weights:")
w_gd
# Compared to sklearn's `LinearRegression()`:
start = time.time()
lr = LinearRegression(fit_intercept=False).fit(X, y)
print(f"Fitting time = {time.time() - start:.4f}s")
print("Weights:")
lr.coef_
# As we can see, the coefficients obtained from gradient descent are very similar to those obtained by sklearn's `LinearRegression()` (although sklearn is much faster). All is well so far.
# ### 1.1
# rubric={accuracy:5}
# In this exercise your task is to implement a function `stochastic_gradient_descent`, that performs SGD, _using_ the `gradient_descent` function provided above. You can have your function accept the same arguments as the `gradient_descent` function above, except:
#
# - Change `n_iters` to `n_epochs`.
# - Add an extra `batch_size` argument.
# - Your implementation of SGD should follow "[Approach 1](https://pages.github.ubc.ca/MDS-2020-21/DSCI_572_sup-learn-2_students/lectures/lecture3_stochastic-gradient-descent.html#sampling-with-or-without-replacement)" from Lecture 3: shuffle the dataset and then divide it into batches. If the numer of samples is not divisible by the `batch_size`, your last batch will have less than `batch_size` samples - you can either throw this last batch away or use it (I usually choose to use it and that's the default in PyTorch, the library we'll be using to build neural networks starting next lecture).
# - You can leave `α` constant for all iterations.
#
# >The pedagogical goal here is to help you see how SGD relates to regular "vanilla" gradient descent. In reality it would be fine to implement SGD "from scratch" without calling a GD function.
# +
def stochastic_gradient_descent(f, f_grad, w0, X, y, n_epochs=1, α=0.001, batch_size=1):
pass # Your solution goes here.
stochastic_gradient_descent(mse, mse_grad, w0, X, y) # Test your function with defaults (results probably won't be very good)
# -
# ### 1.2
# rubric={accuracy:3}
# Show that when the batch size is set to the whole training set (i.e., `batch_size=len(X)`), you get exactly the same estimated coefficients with SGD and GD. Use the same learning rate (`α=0.001`) and number of epochs (`10 ** 5`) for both algorithms.
# +
w_gd = gradient_descent(mse, mse_grad, w0, X, y, n_iters=10**5) # GD coefficients
print(w_gd)
w_sgd = None # Your solution goes here.
print(w_sgd)
# -
# ## Exercise 2: `SGDClassifier` and `SGDRegresor`
# <hr>
# In this exercise we'll explore training a classifier with SGD on the [Sentiment140 dataset](http://help.sentiment140.com/home), which contains tweets labeled with sentiment associated with a brand, product, or topic. Please start by doing the following:
#
# 1. Download the corpus from [here](http://cs.stanford.edu/people/alecmgo/trainingandtestdata.zip).
# 2. Unzip.
# 3. Copy the file `training.1600000.processed.noemoticon.csv` into the current directory.
# 4. Create a `.gitignore` file so that you don't accidentally commit the dataset (I've tried to do this for you but please double check it).
#
# Once you're done the above, steps, run the starter code below:
# Data loading and preprocessing
df = pd.read_csv(
"training.1600000.processed.noemoticon.csv",
encoding="ISO-8859-1",
names=["label", "id", "date", "no_query", "name", "text"],
)
df["label"] = df["label"].map({0: "neg", 4: "pos"}) # change 0's to "neg" and 4's to "pos"
df.head()
# Now we split the data:
X, y = df["text"], df["label"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=2021)
# And then we encode it using `CountVectorizer`, which may take a minute or so:
vec = CountVectorizer(stop_words='english')
X_train = vec.fit_transform(X_train)
X_test = vec.transform(X_test)
# Note that our training data is rather large compared to datasets we've explored in the past:
X_train.shape
# Luckily, `CountVectorizer()` returns us a sparse matrix:
type(X_train)
# Recall that a sparse matrix is a more efficient representation of a matrix that contains many 0's. What percentage of our array is non-zero?
print(f"{X_train.nnz / np.prod(X_train.shape) * 100:.5f}%")
# So few non-zero values - lucky we have a sparse matrix! Anyway, let's train a classifier (this may take a while!):
lr = LogisticRegression()
t = time.time()
lr.fit(X_train, y_train)
print(f"Training took {time.time() - t:.1f} seconds")
print(f"Train score: {lr.score(X_train, y_train):.2f}")
print(f"Test score: {lr.score(X_test, y_test):.2f}")
# ### 2.1
# rubric={accuracy:3}
# In sklearn, there is a classifier called `linear_model.SGDClassifier()` - [see the docs](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html) (there's also a `SGDRegressor` but we won't look at that here). As the name suggests, this model can train linear classifiers with SGD - in the true sense of the algorithm, 1 sample per iteration (i.e., batch size of 1).
#
# Train a logistic regression model on the same dataset above using `SGDClassifier`. Compare the training time of your `SGDClassifier` to `LogisticRegression()`. You'll need to specify the correct `loss` argument in `SGDClassifier()` to train a logistic regression model. [Read the docstring](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html) to find the appropriate `loss`.
#
# > The pedagogical goal here is to demonstrate how using just one sample per iteration in SGD can significantly speed up training, and accuracy-wise, is not a terrible idea!
# +
# Your solution goes here.
# -
# ### 2.2
# rubric={reasoning:2}
# Discuss the training and test accuracies of your `SGDClassifier()` and `LogisticRegression()` models. Is there any difference between the two models?
# +
# Your solution goes here.
# -
# ### 2.3
# rubric={reasoning:3}
# One possible explanation for `SGDClassifier`'s speed is that it's just doing fewer iterations (epochs) before converging. `SGDClassifier` and `LogisticRegression` have an `n_iter_` attribute which you can check after fitting (in these sklearn models, `n_iter_` is equivalent to "number of epochs").
# 1. Compare these values and discuss them in the context of the above hypothesis.
# 2. Then, using the `max_iter` parameter, do a "fair" experiment where `SGDClassifier` and `LogisticRegression` do the same number of passes through the dataset, and comment on the results.
#
# >To be completely "fair" we should also make sure that the tolerance and regularization strength in both models is the same (by default they are not) - but you don't need to worry about that here. Just focus on the number of iterations/epochs.
# Your solution goes here.
#
#
# ## Exercise 3: Neural Networks "By Hand"
# <hr>
# ### 3.1
# rubric={accuracy:4}
# The neural networks we've seen in [Lecture 4](https://pages.github.ubc.ca/MDS-2020-21/DSCI_572_sup-learn-2_students/lectures/lecture4_pytorch-neural-networks-pt1.html) are just recursive functions where each layer is made up of the previous layer's output, multiplied by some weights, with some biases added, and passed through an activation function (we call these networks "fully-connected feed-forward networks", but more on that next week). We don't usually include the input layer when counting up the layers in a network. So a 2 layer network (`L=2`) really means: 1 input layer, 1 hidden layer, 1 output layer. A 3 layer network (`L=3`) really means: 1 input layer, 2 hidden layers, 1 output layer.
#
# Mathematically, our networks have the form:
#
# $$ x^{(l+1)} = h\left( W^{(l)} x^{(l)} + b^{(l)}\right) $$
#
# where:
# - $W^{(l)}$ is a matrix of weights.
# - $b^{(l)}$ is a vector of biases.
# - $x^{(l)}$ is the output of layer $l$:
# - $x^{(0)}$ are the inputs
# - $x^{(1)}$ are the outputs of the first hidden layers, i.e., $x^{(1)} = h\left( W^{(0)} x^{(0)} + b^{(0)}\right)$
# - etc.
# - $x^{(L)} = \hat{y}$
# - Classification: $\hat{y} = h\left( W^{(L-1)} x^{(L-1)} + b^{(L-1)}\right)$
# - Regression: $\hat{y} = W^{(L-1)} x^{(L-1)} + b^{(L-1)}$ (no activation!)
#
# Suppose that we use a neural network with one hidden layer with a **ReLU activation** for a **regression** problem. After training, we obtain the following parameters:
#
# $$\begin{align}W^{(0)} &= \begin{bmatrix}-2 & 2 & -1\\-1 & -2 & 0\end{bmatrix}, &b^{(0)}&=\begin{bmatrix}2 \\ 0\end{bmatrix} \\ W^{(1)} &= \begin{bmatrix}3 & 1\end{bmatrix}, &b^{(1)}&=-10\end{align}$$
#
# For a training example with features $x = \begin{bmatrix}3 \\-2 \\ 2\end{bmatrix}$ what are the values in this network of $x^{(1)}$ and $\hat{y}$? Show your work using code cells or LaTeX.
# +
# Your answer goes here.
# -
# ### 3.2
# rubric={reasoning:4}
# Draw this neural network using a circle/arrow diagram (similar to [the ones I drew in Lecture 4](https://pages.github.ubc.ca/MDS-2020-21/DSCI_572_sup-learn-2_students/lectures/lecture4_pytorch-neural-networks-pt1.html#neural-network-basics)). Label the diagram with the weight/bias values given above. If you want to draw this diagram by hand, that is fine: you can take a photo of the drawing and put it in here. If you are doing so, make sure you upload the image to your repo!
# Your drawing goes here.
#
#
# ## Exercise 4: Predicting Fashion
# <hr>
# In this Exercise I'm going to get you to train a neural network using the Fashion-MNIST dataset. Fashion-MNIST is a set of 28 x 28 pixel greyscale images of clothes. Some of you may have worked with this dataset before - it's a classic. I promise that our datasets will get more interesting than this, but this dataset is ideal for your first PyTorch exercise. Below is a sample of some of the images in the dataset - we have 10 classes in the dataset: T-shirt/tops, Trousers, Pullovers, Dresss, Coats, Sandals, Shirts, Sneakers, Bags, and Ankle Boots.
#
#
# 
# The goal of this exercise is to develop a network that can correctly predict a given image of "fashion" into one of the 10 classes. This is a multi-class classification task, our model should spit out 10 probabilities for a given image - one probability for each class. Ideally the class our model predicts with maximum probability is the correct one!
#
# The below cell will download and load in the Fashion-MNIST data for you. We'll talk more about this process next week, but briefly:
# - Think of images as ndarrays of data, in the case of grayscale images like we have here, each pixel has a value between 0 and 1 indicating how "bright" that pixel is. So each image here is just a 28 x 28 ndarray with values ranging from 0 to 1! (when we get to colour images, it's exactly the same, except each pixel has 3 values, one for each of the colour channels Red, Blue, Green. If we had colour images here our array would be 28 x 28 x 3).
# - `transform`: applies some transformations to the images. Here we are converting the data to tensors. We'll work with these more next week so don't worry too much about them.
# - `torch.utils.data.DataLoader`: these are "data loaders". Think of them as generators. During training/testing, we can easily query them for a batch of data of size `BATCH_SIZE`. Cool!
# +
BATCH_SIZE = 64
# Define a transform to normalize the data, which usually helps with training
transform = transforms.Compose([transforms.ToTensor()])
# Download data
trainset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=True, transform=transform)
testset = datasets.FashionMNIST('~/.pytorch/F_MNIST_data/', download=True, train=False, transform=transform)
# Create data loaders (these are just generators that will give us `batch_size` samples as a time)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# Class labels
class_labels = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle Boot']
# -
# Let's plot a random image (run this cell as many times as you like to see different images):
image, label = next(iter(trainloader)) # Get a random batch of 64 images
i = np.random.randint(0, 64) # Choose one image at random
plt.imshow(image[i, 0], cmap='gray') # Plot
plt.title(class_labels[label[i]]);
# ### 4.1
# rubric={accuracy:3}
# Notice in the plot above that our image is 28 x 28 pixels. How do we feed this into a neural network? Well we can flatten it out into a vector of 784 elements (28 x 28 = 784) and create 784 input nodes! We'll do this later on - for now, all I want you to do is create a new class defining a classifier with the following architecture ([this section of Lecture 4](https://pages.github.ubc.ca/MDS-2020-21/DSCI_572_sup-learn-2_students/lectures/lecture4_pytorch-neural-networks-pt1.html#non-linear-regression-with-a-neural-network) might help here):
# - linear layer that goes from `input_size` -> 256 nodes
# - ReLU activation function
# - linear layer that goes from 256 nodes -> 128 nodes
# - ReLU activation function
# - linear layer that goes from 128 nodes -> 64 nodes
# - ReLU activation function
# - output layer that goes from 64 nodes -> `output_size` nodes
#
# I've given you some starter code to get you going.
#
# >When we create our model in a later exercise we will specify `input_size=784` and `output_size=10`. The `784` is the flattened 28 x 28 image and the output size of 10 is so that we have one node for each item of clothing (remember we have 10 classes), and each node will contain the probability of that item of clothing being the one in a particular input image.
class Classifier(nn.Module):
def __init__(self, input_size, output_size):
super().__init__()
self.main = None # Your code goes here
def forward(self, x):
out = self.main(x)
return out
# ### 4.2
# rubric={accuracy:1}
# If your model has `input_size = 784` and `output_size=10`, how many parameters does your model have? (Ideally you can work this out by hand but you can use `torchsummary` like I showed in Lecture 4 if you must...)
# +
# Your answer goes here.
# -
# ### 4.3
# rubric={accuracy:3}
# We haven't trained yet, but let's test out your network. The below function will help you plot your network's predictions for a particular image using `matplotlib`, run it:
def plot_prediction(image, label, predictions):
"""Plot network predictions with matplotlib."""
fig, (ax1, ax2) = plt.subplots(figsize=(8, 4), ncols=2) # Plot
ax1.imshow(image[0], cmap='gray')
ax1.axis('off')
ax1.set_title(class_labels[label])
ax2.barh(np.arange(10), predictions.data.numpy().squeeze())
ax2.set_title("Predictions")
ax2.set_yticks(np.arange(10))
ax2.set_yticklabels(class_labels)
ax2.set_xlim(0, 1)
plt.tight_layout();
model = Classifier(input_size=784, output_size=10)
# Test on training images (run as many times as you like!)
image, label = next(iter(trainloader)) # Get a random batch of 64 images
predictions = model(image[0].view(1, -1)) # Get first image, flatten to shape (1, 784) and predict it
predictions = nn.Softmax(dim=1)(predictions) # Coerce predictions to probabilities using Softmax()
plot_prediction(image[0], label[0], predictions)
# Okay, those predictions are probably pretty bad. We need to train! Below is a training function (the same one we saw in Lecture 4). The only difference is that when I'm creating `y_hat` (my model predictions), I'm reshaping my data to be of shape `(batch_size, 784)` using `X.view(X.shape[0], -1)` so we can feed it into our network (`X.shape[0]` is the batch size, and the -1 just says "flatten remaining dimensions into a single dimension", so this turns something of shape `(64, 28, 28)` to `(64, 784)` which in English reads as "64 examples of vectors with length 784", i.e. 64 flattened images).
def trainer(model, criterion, optimizer, dataloader, epochs=5, verbose=True):
"""Simple training wrapper for PyTorch network."""
for epoch in range(epochs):
losses = 0
for X, y in dataloader:
optimizer.zero_grad() # Clear gradients w.r.t. parameters
y_hat = model(X.view(X.shape[0], -1)) # Reshape data to (batch_size, 784) and forward pass to get output
loss = criterion(y_hat, y) # Calculate loss
loss.backward() # Getting gradients w.r.t. parameters
optimizer.step() # Update parameters
losses += loss.item() # Add loss for this batch to running total
if verbose: print(f"epoch: {epoch + 1}, loss: {losses / len(dataloader):.4f}")
# Define an appropriate `criterion` and `optimizer` to train your model with.
# - We are doing multi-class classification here, what loss function do we use for this case (hint: see [this part of Lecture 4](https://pages.github.ubc.ca/MDS-2020-21/DSCI_572_sup-learn-2_students/lectures/lecture4_pytorch-neural-networks-pt1.html#multiclass-classification-optional))?
# - Use any optimizer you like but I recommend Adam (by the way, optional Exercise 5 of this lab gets you to implement Adam from scratch 😉 )
#
# I already created the dataloader `trainloader` for you at the start of this exercise. Pass all these things to `trainer()` to train your model (it may take a few minutes):
# +
# Your answer goes here.
# Uncomment and run the below once you've defined a criterion and optimizer
# trainer(model, criterion, optimizer, trainloader, epochs=5, verbose=True)
# -
# ### 4.4
# rubric={accuracy:1}
# Test out your newly trained network on the training data:
# Test model on training images (run as many times as you like!)
image, label = next(iter(trainloader)) # Get a random batch of 64 images
predictions = model(image[0].view(1, -1)) # Get first image, flatten to shape (1, 784) and predict it
predictions = nn.Softmax(dim=1)(predictions) # Coerce predictions to probabilities using Softmax()
plot_prediction(image[0], label[0], predictions)
# And test it out on the test data:
# Test model on testing images (run as many times as you like!)
image, label = next(iter(testloader)) # Get a random batch of 64 images
predictions = model(image[0].view(1, -1)) # Get first image, flatten to shape (1, 784) and predict it
predictions = nn.Softmax(dim=1)(predictions) # Coerce predictions to probabilities using Softmax()
plot_prediction(image[0], label[0], predictions)
# Pretty sweet! That's all there is to it - you just created your first legitimate classifier! WELL DONE! I personally am super excited for you.
#
# Oh right, I need to give you an exercise to answer. In this exercise we used a `BATCH_SIZE = 64`. This is a pretty common size to use, in fact the most common sizes are: 32, 64, 128, 256, and 512. In terms of optimizing a network, list one difference between using a small batch size vs large batch size?
# Your answer goes here.
#
#
# ### 4.5
# No marks and nothing to do for this question, just a bit of fun. Our network was trained on clothing images, but that doesn't mean we can't use it to predict other images (but this is probably a bad idea)! Let's see what our model thinks of a 28 x 28 image of me:
image = torch.from_numpy(plt.imread("img/tomas_beuzen.png"))
predictions = model(image.view(1, -1)) # Flatten image to shape (1, 784) and predict it
predictions = nn.Softmax(dim=1)(predictions) # Coerce predictions to probabilities using Softmax()
label = predictions.argmax(dim=1) # Get class label from max probability
plot_prediction(image.numpy()[None, :, :], label, predictions)
# ## (Optional) Exercise 5: Implementing Adam Optimization From Scratch
# <hr>
#
# rubric={accuracy:1}
# Adam is an optimization algorithm that we'll be using a lot for the rest of the course. [Here's the original paper](https://arxiv.org/abs/1412.6980) that proposed it. It is essentially a fancier version of SGD. Without getting too technical, Adam really adds two additional features to SGD:
# 1. Momentum: which uses past gradients to help improve convergence speed, reduce noise in the path to the minimum, and avoid local minima.
# 2. Per-parameter learning rate: a learning rate is maintained and adapted for each parameter as iterations of optimization proceed.
#
# Pretty cool! I recommend [reading this article](https://ruder.io/optimizing-gradient-descent/index.html) or [watching this video](https://www.youtube.com/watch?v=JXQT_vxqwIs) to learn more, but Adam boils down to the following equations:
#
# Weight updating:
#
# $$\mathbf{w}_{t+1} = \mathbf{w}_{t} - \frac{\alpha}{\sqrt{\hat{v}_{t}} + \epsilon} \hat{m}_{t}$$
#
# The various components required for that equation:
#
# $$\begin{align}
# \hat{m}_{t} &= \frac{m_{t}}{1 - \beta_{1}^{t}}\\
# \hat{v}_{t} &= \frac{v_{t}}{1 - \beta_{2}^{t}}
# \end{align}$$
#
# $$\begin{align}
# m_{t} &= \beta_{1} m_{t-1} + (1 - \beta_{1}) g_{t}\\
# v_{t} &= \beta_{2} v_{t-1} + (1 - \beta_{2}) g_{t}^{2}
# \end{align}$$
#
# Where:
# - $t$ is the iteration of optimization, it increments up by one each time you update $\mathbf{w}$. Note that in the equation for $\hat{m}_{t}$ and $\hat{v}_{t}$, $\beta_{1}$ and $\beta_{2}$ are raised to the power of $t$.
# - $g_{t}$ is the gradient of the loss function w.r.t to the parameters $w$.
# - $m_{t}$ is known as the first moment (the mean) of the gradients. Initialize as 0.
# - $v_{t}$ is known as the second moment (the uncentered variance) of the gradients. Initialize as 0.
# - $\alpha$ is the learning rate. 0.1 is a good start.
# - $\epsilon$ is just a term to prevent division by zero. Default: $10^{-8}$.
# - $\beta_{1}$ is a hyperparameter that controls the influence of past gradients on subsequent updates. Default: $0.9$.
# - $\beta_{2}$ is a hyperparameter that controls the influence of past gradients on subsequent updates. Default: $0.999$.
#
# Here's a squiggly function for you to try and find the minimum parameter for. I've hard-coded the "optimum parameter" as $w_{opt}=4$ but I want you to find this value using Adam optimization and starting at $w \neq w_{opt}$. I've provided you the function (`f()`), the MSE loss w.r.t this function (`loss()`), and the gradient of the loss (`loss_grad()`). Run the cell below:
# +
def f(w, X):
"""Squiggly function"""
return w * np.cos(w * X)
def loss(w, X, y):
"""MSE loss."""
return np.mean((f(w, X) - y) ** 2)
def loss_grad(w, X, y):
"""Gradient of MSE."""
t = np.cos(w * X) - w * X * np.sin(w * X)
return np.mean((f(w, X) - y) * t)
w_opt = 4
X = np.arange(-3, 3, 0.1)
y = f(w_opt, X)
l = [loss(w, X, y) for w in np.arange(-10, 11, 0.1)]
plt.plot(np.arange(-10, 11, 0.1), l)
plt.xlabel("w")
plt.ylabel("MSE")
plt.grid(True);
# -
# Your task here is to implement Adam from scratch. Then use it to find $w_{opt}$ for the above function. I've provided some code below that you should run when you're ready. Note:
# - I've specified a default of 100 epochs. We have a *tiny* dataset here of 60 samples so this is nothing really. Feel free to add more epochs if you wish.
# - You can start with the default values for the various Adam terms I give in the equations above.
# - You *may* need to play around with the hyperparameter $\alpha$ to get to the minimum (I've given a default of 0.3 in the starter code below - I didn't need to change this value in my solution). You can leave $\beta_{1}$, $\beta_{2}$ as is - often we don't tune those ones and I didn't in my solution, but you can tune them if you want.
# - Adam is generally used with batches like SGD so my solution has the ability to accept a `batch_size` argument, but you don't have to code up this functionality and I didn't include that argument in the starter code below. So feel free to just use all the data each iteration for simplicity like vanilla GD would do. But if you're feeling adventurous, thrown in a `batch_size` argument 😉
#
# >The pedagogical goal here is to get you to implement Adam and play around with it to see how it can "jump over" local minima. If you get it working, it's pretty awesome!
# +
def Adam(X, y, w0, loss, loss_grad, n_epochs=100, alpha=0.3, beta1=0.9, beta2=0.999, eta=10e-8):
# Your code goes here.
w = None
return w
w0 = [9]
w = Adam(X, y, w0, loss, loss_grad)
print(w) # Should be close to 4
# -
# ## (Optional) Exercise 6: Gif or Jiff
# <hr>
#
# rubric={accuracy:1}
# Practically a free mark here for making it to the end. But I do have a question for you.
#
# .gif files are animations made of a series of images. For example, this is me when someone tells me they have 80% accuracy with their ML model:
#
# 
#
# My question to you - are these files pronounced "Gif" or "Jiff"?
#
# Your answer here:
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-f7d15fa1e22d412c", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Submit to Canvas and GitHub
# <hr>
# -
# When you are ready to submit your assignment do the following:
# 1. Run all cells in your notebook to make sure there are no errors by doing `Kernel -> Restart Kernel and Run All Cells...`
# 2. Save your notebook.
# 3. Convert your notebook to `.html` format using the `convert_notebook()` function below or by `File -> Export Notebook As... -> Export Notebook to HTML`
# 4. Run the code `submit()` below to go through an interactive submission process to Canvas.
# 5. Finally, push all your work to GitHub (including the rendered html file).
# + nbgrader={"grade": false, "grade_id": "cell-cd03c1d25ce1d32e", "locked": true, "schema_version": 3, "solution": false, "task": false}
# convert_notebook("lab2.ipynb", "html") # save your notebook, then uncomment and run when you want to convert to html
# +
# submit(course_code=59090) # uncomment and run when ready to submit to Canvas
# -
# 
| release/lab2/lab2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn import decomposition
from sklearn.preprocessing import MinMaxScaler
from sklearn.decomposition import PCA
import seaborn as sns
from scipy.io import loadmat
X = loadmat('PaviaU.mat')['paviaU']
y = loadmat('PaviaU_gt.mat')['paviaU_gt']
print("X:", (X.shape))
print("y:", (y.shape))
np.unique(y)
plt.figure(figsize=(10, 10))
plt.imshow(y, cmap='jet')
plt.colorbar()
plt.axis('off')
plt.title('Ground Truth')
plt.gca().legend(['0', '1', '2', '3', '4', '5', '6', '7', '8', '9'], loc='upper left')
#plt.savefig('ground_truth.png')
plt.show()
_,F,S = X.shape
print(F," ",S)
D1= X[:,:,0:36]
print(D1.shape)
D2= X[:,:,36:76]
print(D2.shape)
D3= X[:,:,76:]
print(D3.shape)
_,F1,S1 = D1.shape
_,F2,S2 = D2.shape
_,F3,S3 = D3.shape
print(F1," ",S1)
print(F2," ",S2)
print(F3," ",S3)
M = X.T.mean(axis=0)
M.shape
M=M.T;
M.shape
M1=D1.T.mean(axis=0)
M1.shape
M1=M1.T;
M1.shape
M2=D2.T.mean(axis=0)
M2.shape
M2=M2.T;
M2.shape
M3=D3.T.mean(axis=0)
M3.shape
M3=M3.T;
M3.shape
np.stack([M1 for _ in range(S1)], axis=2).shape
I1 = (D1 - np.stack([M1 for _ in range(S1)], axis=2));
I1 = I1.reshape(I1.shape[0]*I1.shape[1], I1.shape[2])
I1.shape
I2 = (D2 - np.stack([M2 for _ in range(S2)], axis=2));
I2 = I2.reshape(I2.shape[0]*I2.shape[1], I2.shape[2])
I2.shape
I3 = (D3 - np.stack([M3 for _ in range(S3)], axis=2));
I3 = I3.reshape(I3.shape[0]*I3.shape[1], I3.shape[2])
I3.shape
def applyPCA(X, numComponents, isReshape = True, drawPlot = False, layerNo = 1):
if isReshape == True:
newX = np.reshape(X, (-1, X.shape[2]))
pca = PCA(n_components=numComponents, whiten=True)
newX = pca.fit_transform(newX)
newX = np.reshape(newX, (X.shape[0],X.shape[1], numComponents))
else:
pca = PCA(n_components=numComponents, svd_solver='full')
newX = pca.fit_transform(X)
if drawPlot == True:
drawPCAPlot(pca.explained_variance_ratio_,numComponents, layerNo)
return newX, pca.explained_variance_ratio_
SX1,pca1 = applyPCA(I1,numComponents = 0.99,isReshape = False)
SX1.shape
SX2,pca2 = applyPCA(I2,numComponents = 0.99,isReshape = False)
SX2.shape
SX3,pca3 = applyPCA(I3,numComponents = 0.99,isReshape = False)
SX3.shape
SX= np.concatenate([SX1.T,SX2.T,SX3.T])
SX = SX.T
SX.shape
pca1
pca_var = []
pca_var = np.concatenate([pca1,pca2,pca3])
pca_var = sorted(pca_var, reverse = True)
MAX = max(np.cumsum(pca_var))
pca_var = np.round((pca_var)/MAX,3)
np.cumsum(pca_var)
print(pca_var)
plt.figure()
plt.plot(np.cumsum(pca_var))
plt.xlabel('Number of Components')
plt.ylabel('Cumulative Variance') #for each component
#plt.title('Pavia University Dataset Explained Variance')
plt.savefig('Pavia_University_explainedvariance_final1')
plt.show()
y = y.ravel()
print(SX.shape)
print(y.shape)
X = (SX[y>0,:])
Y = (y[y>0])
print(X.shape)
print(Y.shape)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, Y, random_state = 10, test_size = 0.25, stratify=Y )
print(X_train.shape)
print(X_test.shape)
label_tr,counts_tr=np.unique(y_train,return_counts=True)
label_te,counts_te=np.unique(y_test,return_counts=True)
print(pd.DataFrame(counts_tr,label_tr))
print(pd.DataFrame(counts_te,label_te))
# +
#Applying Scalar to train and test Dataset
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(X_train)
X_train= scaler.transform(X_train)
X_test=scaler.transform(X_test)
# +
#Appplying SVM
from sklearn.svm import SVC
from sklearn import metrics
# +
classifiers = []
Train_acc=[]
Valid_acc=[]
accuracy = -1
accuracy_train = -1
for C in np.arange(1,102,25):
for gamma in np.arange(0.001,0.1,0.005):
clf = SVC(C=C, gamma=gamma)
print("----------------------------------------------------------------")
clf.fit(X_train, y_train)
classifiers.append((C, gamma, clf))
y_pred = clf.predict(X_train)
acc_train =metrics.accuracy_score(y_train, y_pred)
Train_acc.append(acc_train)
y_pred = clf.predict(X_test)
acc=metrics.accuracy_score(y_test, y_pred)
Valid_acc.append(acc)
if (acc_train>accuracy_train):
accuracy_train=acc_train
best_c_train = C
best_g_train=gamma
print("C_train= ",C," Best C_train = ",best_c_train," gamma_train = ",gamma," best_gamma_train= ",best_g_train," Accuracy_train = ",acc_train," Best Accuracy_train = ",accuracy_train,"\n")
if (acc>accuracy):
accuracy=acc
best_c = C
best_g=gamma
print("C= ",C," Best C = ",best_c," gamma = ",gamma," best_gamma= ",best_g," Accuracy = ",acc," Best Accuracy = ",accuracy,"\n")
# -
clf = SVC(C=101,gamma=0.006,decision_function_shape='ovo')
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print("Accuracy :",metrics.accuracy_score(y_pred, y_test)*100)
from sklearn.metrics import cohen_kappa_score
print("Cohen Kappa Score :",cohen_kappa_score(y_pred, y_test)*100)
from sklearn.metrics import confusion_matrix,classification_report
mat = confusion_matrix(y_pred, y_test)
pd.DataFrame(mat)
print(confusion_matrix(y_test, y_pred ))
print(classification_report(y_test, y_pred ))
| spca+svm[pavia university].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Return the node with largest number:
# +
class BinaryTreeNode:
def __init__(self, data):
self.data = data
self.left = None
self.right = None
def treeInput():
rootData = int(input())
if rootData == -1:
return None
root = BinaryTreeNode(rootData)
root.left = treeInput()
root.right = treeInput()
return root
def printTreeDetailed(root):
if root == None:
return
print(root.data, end = ":")
if root.left is not None:
print("L", root.left.data, end = ",")
if root.right is not None:
print("R", root.right.data, end = " ")
print()
printTreeDetailed(root.left)
printTreeDetailed(root.right)
def findLargest(root):
if root == None:
return -1 # Ideally return -infinity
leftLargest = findLargest(root.left)
rightLargest = findLargest(root.right)
m = max(root.data, leftLargest, rightLargest) # Max functions works for 3 numbers as well
return m
root = treeInput()
print(findLargest(root))
| 13 Binary Trees - 1/13.07 Node with Largest Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem set 3: Sentiment
#
# ## Description
#
# The goal of this problem set is to compare the sentiments associated with male and female characters in a small corpus of 41 novels.
#
# This task has a bit in common with <NAME>'s work on the adjectives and body parts associated with male and female characters, though our task is significantly less complex (because we are not doing dependency parsing in order to associate specific words with individual character references).
#
# Here's the approach you should implement:
#
# 1. For each text in the corpus ...
# 1. Break the text into sentences and tokenize each sentence.
# 1. Assign a subject gender to each sentence by counting occurrences of gendered words ("he", "she", etc.). Refine these lists as you see fit.
# 1. Optional: devise ways to treat gender as a non-binary variable.
# 1. Assign sentiment scores to each sentence using the 10 sentiment types included in the NRC EmoLex lexicon ("anticipation", "disgust", "trust", etc.).
# 1. Calculate the mean sentiment score(s) for the sentences in the text that you identified as male-centric and female-centric.
# 1. Optional: If you chose to work with non-binary gender assignments, devise an appropriate way to group gender-similar sentences using your assignments.
# 1. Compare your grouped sentiment scores among the novels in the corpus.
# 1. Write roughly one paragraph of analysis and conclusions.
#
# Let's get to it.
#
# The next block of code contains some variables and import statements. **Make sure your variables point to the correct files on your system.** Get these files from the course GitHub site.
# +
from collections import defaultdict
from glob import glob
from nltk import word_tokenize, sent_tokenize
import numpy as np
import os
import string
# Files and locations
novel_files = glob(os.path.join('..', '..', 'data', 'texts', '*.txt'))
emolex_file = os.path.join('..', '..', 'data', 'lexicons', 'emolex.txt')
# Female and male wordlist from <NAME> via Bengfort et al.
# See bit.ly/2GJBGfV
male_words = set([
'guy','spokesman','chairman',"men's",'men','him',"he's",'his',
'boy','boyfriend','boyfriends','boys','brother','brothers','dad',
'dads','dude','father','fathers','fiance','gentleman','gentlemen',
'god','grandfather','grandpa','grandson','groom','he','himself',
'husband','husbands','king','male','man','mr','nephew','nephews',
'priest','prince','son','sons','uncle','uncles','waiter','widower',
'widowers'
])
female_words = set([
'heroine','spokeswoman','chairwoman',"women's",'actress','women',
"she's",'her','aunt','aunts','bride','daughter','daughters','female',
'fiancee','girl','girlfriend','girlfriends','girls','goddess',
'granddaughter','grandma','grandmother','herself','ladies','lady',
'lady','mom','moms','mother','mothers','mrs','ms','niece','nieces',
'priestess','princess','queens','she','sister','sisters','waitress',
'widow','widows','wife','wives','woman'
])
dev_text = '''\
He was the father of modern gardening.
She is the mother of computer science.
They are both excellent at their jobs.
Jane is generally happy with her work.
Jonathan had a bad day, but he's fine.
He and she are the parents of a girl.'''
# -
# ## Tokenize and preprocess text (10 points)
#
# **Write a function, `tokenize_text`, that takes as input a text string and returns a list sentences, each of which contains a list of tokens.** We'll use this function to tokenize each novel.
#
# Your tokenized text should be all lowercase. You may remove punctuation and stopwords, but be careful not to remove any of the gender-indicative pronouns that are included in the `male_words` and `female_words` lists above.
#
# **Run your `tokenize_text` function on the `dev_text` string and print the result.**
# +
def tokenize_text(text, stopwords=None):
'''
Takes a string.
Returns a list of tokenized sentences.
'''
tokenized = []
for sent in sent_tokenize(text):
tokens = word_tokenize(sent)
# if stopwords != None:
# tokens = [token for token in tokens if token not in stopwords]
tokenized.append(tokens)
return tokenized
# Display your results
display(tokenize_text(dev_text))
# -
# ## Assign gender scores (10 points)
#
# **Write a function, `gender_score`, that takes as input a list of word-level tokens and returns a gender score. Run this function over the tokenized sentences in `dev_text` and print the resulting scores.**
#
# The list of input tokens represents a single, tokenized sentence.
#
# The gender score may be either binary for female/male (according to the prepoderance of male or female terms in the sentence, `1 = female`, `0 = male`) or a floating-point number between 0 (male) and 1 (female) that represents the proportion of female and male words from the lists above. In either case, think about how you want to represent sentences that have no gender terms.
from collections import Counter
def gender_score(token_list, male_words, female_words, binary=True):
'''
Takes a list of tokens.
Returns a gender score between 0 and 1, or None.
'''
if binary is False:
gender_c = 0
all_c = 0
for i in token_list:
i = i.lower()
if i in male_words or i in female_words:
gender_c += 1
all_c += 1
if gender_c == 0:
return None
else:
return(gender_c/all_c)
else:
gender_c = Counter()
for i in token_list:
i = i.lower()
if i in male_words:
gender_c['male'] += 1
elif i in female_words:
gender_c['female'] +=1
if gender_c['male'] > gender_c['female']:
return 0
elif gender_c['male'] < gender_c['female']:
return 1
else:
return None
# Run your gender-scoring function on the tokenized dev_text
for sent in enumerate(tokenize_text(dev_text),1):
print("Sent:", sent[0], "\tGender:", gender_score(sent[1],male_words, female_words))
# ## Calculate sentence-level sentiment scores (20 points)
#
# **Write a function, `sentiment_score`, that takes as input a list of word-level tokens and an EmoLex lexicon and returns a dictionary of length-normalized EmoLex sentiment scores for every available emotion type. Run this function over the tokenized sentences in `dev_text` and print the resulting sentence-level scores.**
#
# The list of tokens used as input data represents a single, tokenized sentence.
# By "length-normalized," I mean that each sentence-level score should be divided by the number of tokens in the sentence.
#
# Your output dictionary should look like this (sample output on the fourth `dev_text` sentence):
#
# ```
# {
# 'anger': 0.0,
# 'anticipation': 0.125,
# 'disgust': 0.0,
# 'fear': 0.0,
# 'joy': 0.125,
# 'negative': 0.0,
# 'positive': 0.125,
# 'sadness': 0.0,
# 'surprise': 0.0,
# 'trust': 0.125
# }
# ```
# +
# A freebie helper function to read and parse the emolex file
def read_emolex(filepath=None):
'''
Takes a file path to the emolex lexicon file.
Returns a dictionary of emolex sentiment values.
'''
if filepath==None: # Try to find the emolex file
filepath = os.path.join('..','..','data','lexicons','emolex.txt')
if os.path.isfile(filepath):
pass
elif os.path.isfile('emolex.txt'):
filepath = 'emolex.txt'
else:
raise FileNotFoundError('No EmoLex file found')
emolex = defaultdict(dict) # Like Counter(), defaultdict eases dictionary creation
with open(filepath, 'r') as f:
# emolex file format is: word emotion value
for line in f:
word, emotion, value = line.strip().split()
emolex[word][emotion] = int(value)
return emolex
# Get EmoLex data. Make sure you set the right file path above.
emolex = read_emolex(emolex_file)
# +
# Sentiment scoring function
def sentiment_score(token_list, lexicon=None):
'''
Takes a tokenized sentence.
Returns a dictionary of length-normalized EmoLex sentiment scores.
'''
sent_score = {
'anger': 0.0,
'anticipation': 0.0,
'disgust': 0.0,
'fear': 0.0,
'joy': 0.0,
'negative': 0.0,
'positive': 0.0,
'sadness': 0.0,
'surprise': 0.0,
'trust': 0.0
}
count = 0
for i in token_list:
if i in emolex:
for x in sent_score:
sent_score[x] = sent_score[x] + emolex[i][x]
count += 1
for i in sent_score:
sent_score[i] = sent_score[i]/count
return(sent_score)
# Run scoring function on tokenized dev text and print results
for sent in enumerate(tokenize_text(dev_text),1):
print("Sent:", sent[0], "\tSentiment:", sentiment_score(sent[1]))
# -
# ## Compare emotions in female- and male-tagged sentences (20 points)
#
# **Write a function, `gendered_sentiment`, that uses `gender_score` and `sentiment_score` to calulate the mean sentiment scores for each EmoLex emotion type in female- and male-tagged sentences in a tokenized input text. Then print the output of this function when run on tokenized `dev_text`.**
#
# We've done this somewhat ineffeciently, since each of those functions iterates over all tokens (so we make two passes over the text). But this inefficiency made it easier to break up the task into its constituent parts.
#
# Your function should take as input a tokenized text and should return a dictionary with keys for each emotion type, subkeys `male` and `female`, and values representing the mean score for that emotion in sentences corresponding to the indicated gender. For example (invented, not real data):
#
# ```
# {
# 'anger': {
# 'male' : 0.02,
# 'female' : 0.03
# },
# ...
# }
# ```
#
# If you chose to operationalize gender as a nonbinary value, you'll need to decide how to treat values other than 0 and 1. You could split at 0.5 (how do you want to treat gender scores equal to 0.5?) or discard mid-level scores (between 0.3 and 0.7, say), or some other approach. And, no matter how you calculated your gender score, you'll need to handle genderless sentences appropriately (by ignoring them).
# +
# Sentiment comparison by gender
def gendered_sentiment(tokenized_text, lexicon=None, binary=True, female_level=0.7, male_level=None):
all_score = {
'anger': {'male': 0.0, 'female':0.0},
'anticipation': {'male': 0.0, 'female':0.0},
'disgust': {'male': 0.0, 'female':0.0},
'fear': {'male': 0.0, 'female':0.0},
'joy': {'male': 0.0, 'female':0.0},
'negative': {'male': 0.0, 'female':0.0},
'positive': {'male': 0.0, 'female':0.0},
'sadness': {'male': 0.0, 'female':0.0},
'surprise': {'male': 0.0, 'female':0.0},
'trust': {'male': 0.0, 'female':0.0}
}
sent_score = {
'anger': 0.0,
'anticipation': 0.0,
'disgust': 0.0,
'fear': 0.0,
'joy': 0.0,
'negative': 0.0,
'positive': 0.0,
'sadness': 0.0,
'surprise': 0.0,
'trust': 0.0
}
#iterate through tokenized text
for i in tokenized_text:
gender_c = Counter()
emo_c = Counter()
#interate through each element in tokenized text (so, each word)
for x in i:
#lowercase each word
x = x.lower()
#if a word is in male words, counter for male goes up 1
if x in male_words:
gender_c['male'] += 1
#if a word is in female words, counter for female goes up 1
if x in female_words:
gender_c['female'] += 1
#if a word is in emolex, then using sent_score as the basis for the emotional words used, we add a counter of the emotion to the emotion counter
#but by doing emo_c[y] I am making a counter per word, so say for the word anger-> emo_c['anger'] +1 and emo_c[negative] +1
if x in emolex:
for y in sent_score:
emo_c[y] += emolex[x][y]
#if gender counter is above 0 then interate through sent_score to add the emo counter numbers to all_score
if gender_c['male'] > 0:
for a in sent_score:
all_score[a]['male'] += emo_c[a]
#if gender counter is above 0 then interate through sent_score to add the emo counter numbers to all_score
elif gender_c['female'] > 0 :
for a in sent_score:
all_score[a]['female'] += emo_c[a]
#normalize the numbers from all_score- max equal .5
#method: after counting the scores from emotion per gender- I decided to multiply each number by normalize to make the maximum value .5
for a in sent_score:
if sum(all_score[a].values()) == 0:
all_score[a]['female'] = 0
all_score[a]['male'] = 0
else:
normalize = .5/sum(all_score[a].values())
all_score[a]['female'] = all_score[a]['female']*normalize
all_score[a]['male'] = all_score[a]['male']*normalize
return(all_score)
#I am not sure if this is what the question was asking to do, but this is how I interpreted the question.
# Run function and display results
scores = gendered_sentiment(tokenize_text(dev_text), lexicon=emolex)
display(scores)
# -
# ## Visualize your output (5 points)
#
# This one's a freebie, so long as the output of your `gendered_sentiment` function meets the problem spec.
#
# **Use the `visualize_emotion_scores` function below to produce a plot of your comparative gender scores in `dev_text` for all 10 emotion categories.**
# %matplotlib inline
def visualize_emotion_scores(scores):
import pandas as pd
import seaborn as sns
df = pd.DataFrame.from_dict(scores)
values = df.columns
df = df.reset_index().melt(id_vars='index', value_vars=values, var_name='emotion').rename(columns={'index':'gender'})
sns.set_context('poster')
g = sns.catplot(x='gender', y='value', col='emotion', data=df, kind='bar', col_wrap=4)
return g
g = visualize_emotion_scores(scores)
# ## Calculate scores in a novel (5 points)
#
# **Use the code you've written to calculate and plot the gendered emotion scores in *Madame Bovary*.**
#
# Your output should include both the mean scores printed to standard output (here in the notebook) and a visualization created using the `visualize_emotion_scores` function.
#
# Incidentally, working with large texts can be slow. But scoring a novel as we've done here shouldn't take more than a few seconds. If you're waiting minutes for your scoring to finish, something is wrong.
bovary_path = os.path.join('..','..','data','texts','F-Flaubert-Madame_Bovary-1857-M.txt')
with open(bovary_path, 'r') as f:
bovary_text = f.read()
bovary = tokenize_text(bovary_text)
bov_score = gendered_sentiment(bovary, lexicon=emolex)
display(bov_score)
visualize_emotion_scores(bov_score)
# ## Grand finale: Sentiment and gender in a small corpus (20 points)
#
# **Do the same thing you just did for one novel, but for all 41 novels in the `data/texts` directory (from GitHub). Calculate the mean sentiment score across the corpus for each emotion and each gender, as well as the standard deviation for each of those means. Display your results in a legible form.**
# +
# %%time
# Score all 41 novels in the corpus
# Takes about a minute on my aging laptop
corpus_scores = {'gender':[],'sentiment':[]} # Dictionary to hold results
for novel in novel_files: # Iterate over novels
with open(novel, 'r') as f:
novel_text = f.read() # Read a novel as a string
novel_label = os.path.split(novel)[1].rstrip('.txt') # Get convenience label for novel
tokenized = tokenize_text(novel_text)
for i in tokenized:
case = gender_score(i, male_words, female_words)
corpus_scores['gender'].append(case)
case_sent = sentiment_score(i)
corpus_scores['sentiment'].append(case_sent)
# -
# %%time
corpus_scores1 = {'gendered_sent':[]}
for novel in novel_files: # Iterate over novels
with open(novel, 'r') as f:
novel_text = f.read() # Read a novel as a string
novel_label = os.path.split(novel)[1].rstrip('.txt') # Get convenience label for novel
# for i in tokenized:
scores = gendered_sentiment(tokenize_text(novel_text), lexicon=emolex)
corpus_scores1['gendered_sent'].append(scores)
# %%time
for x in corpus_scores['gender']:
if x is None:
corpus_scores['gender'].remove(x)
# Aggregate scores for all novels by emotion type and gender
all_gender = {'male': 0, 'female': 0}
for i in corpus_scores['gender']:
if i == 1:
all_gender['female'] += 1
if i == 0:
all_gender['male'] += 1
print(all_gender) #comparing how many gendered sentences there are per gender
all_sentiment = {
'anger': 0.0,
'anticipation': 0.0,
'disgust': 0.0,
'fear': 0.0,
'joy': 0.0,
'negative': 0.0,
'positive': 0.0,
'sadness': 0.0,
'surprise': 0.0,
'trust': 0.0
}
n_sent = {
'anger': [],
'anticipation': [],
'disgust': [],
'fear': [],
'joy': [],
'negative': [],
'positive': [],
'sadness': [],
'surprise': [],
'trust': []
}
for i in range(len(corpus_scores['sentiment'])):
for x in all_sentiment:
all_sentiment[x] += corpus_scores['sentiment'][i][x]
n_sent[x].append(corpus_scores['sentiment'][i][x])
for i in all_sentiment:
all_sentiment[i]= all_sentiment[i]/len(corpus_scores['sentiment']) #calculating mean per all setiment scored item
print(all_sentiment)
# print(n_sent)
gs = {
'anger': {'male': [], 'female':[]},
'anticipation': {'male': [], 'female':[]},
'disgust': {'male': [], 'female':[]},
'fear': {'male': [], 'female':[]},
'joy': {'male': [], 'female':[]},
'negative': {'male': [], 'female':[]},
'positive': {'male': [], 'female':[]},
'sadness': {'male': [], 'female':[]},
'surprise': {'male': [], 'female':[]},
'trust': {'male': [], 'female':[]}
}
for i in range(len(corpus_scores1['gendered_sent'])):
for x in gs:
gs[x]['female'].append(corpus_scores1['gendered_sent'][i][x]['female'])
gs[x]['male'].append(corpus_scores1['gendered_sent'][i][x]['male'])
# Calculate corpus-wide means and standard deviations
# calculating the sentiment score means and std (same as above)
import statistics
from statistics import mean
from statistics import stdev
mean_and_std_sent = {
'anger': {'mean': 0.0, 'std':0.0},
'anticipation': {'mean': 0.0, 'std':0.0},
'disgust': {'mean': 0.0, 'std':0.0},
'fear': {'mean': 0.0, 'std':0.0},
'joy': {'mean': 0.0, 'std':0.0},
'negative': {'mean': 0.0, 'std':0.0},
'positive': {'mean': 0.0, 'std':0.0},
'sadness': {'mean': 0.0, 'std':0.0},
'surprise': {'mean': 0.0, 'std':0.0},
'trust': {'mean': 0.0, 'std':0.0}
}
for i in n_sent:
m = statistics.mean(n_sent[i])
std = statistics.stdev(n_sent[i])
mean_and_std_sent[i]['mean'] = m
mean_and_std_sent[i]['std'] = std
# print(i)
print(mean_and_std_sent) #same as all_sentiments
print(all_sentiment)
#mean and std of gendered sentiments
mean_and_std_gs = {
'anger': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'anticipation': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'disgust': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'fear': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'joy': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'negative': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'positive': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'sadness': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'surprise': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}},
'trust': {'male': {'mean': 0.0, 'std':0.0}, 'female':{'mean': 0.0, 'std':0.0}}
}
for i in gs:
m_f = statistics.mean(gs[i]['female'])
m_m = statistics.mean(gs[i]['male'])
std_f = statistics.stdev(gs[i]['female'])
std_m = statistics.stdev(gs[i]['male'])
mean_and_std_gs[i]['female']['mean'] = m_f
mean_and_std_gs[i]['male']['mean'] = m_m
mean_and_std_gs[i]['female']['std'] = std_f
mean_and_std_gs[i]['male']['std'] = std_m
print(mean_and_std_gs)
# ## Discuss your results (10 points)
#
# Write a short paragraph in which you summarize your findings and suggest ways in which the experiment might be improved.
# I found that these results in total were very interesting. I think the obvious points here are that thre seems to be a gendered bias in the language used in novels. Looking at the means and std of the sentiment scoring, it is hard to find a narrative, but I think the means and std of the gendered sentiment scoring shows another picture. It shows a picture of confirmed bias in writing. How I set up the gendered sentiment scoring was that I counted all the genered words and the emotional words within a sentence and weighed the emotions based on the amount of genered words (ex. in a sentence with more male words- male will be more weighed), so when looking at these means and stds, it can be seen that the language is heavily skewed in one direction. It seems that there is an overwhemingly large amount of male words, skewing the data- showing that all emotions are scored in favor of male words. However, maybe it is not much of a surprise, when looking at the how less frequently female words are used (nearly 20,000 times less). In conclusion, these novels have a tendancy to use more male words.
# ## Optional bonus (10 points)
#
# Extend your analysis to different author genders, nationalities, or time periods.
#
# The files in the corpus are informatively named:
#
# ```
# Nation-Author-Title-Year-Gender.txt
# ```
#
# Use the convenience function below to parse the corpus file paths into a dictionary of metadata, then segment the corpus-wide emotion scores into one or more dimensions by author gender, nation of origin, or era of publication. Do you notice any interesting trends in this segmented data?
#
# **NB.** We're obviously not imposing much statistical rigor here, and we're working with a very small corpus. Treat your results as a toy example!
# Convenience function to parse file paths into metadata
def parse_filename(path):
'''
Takes a formatted file path string.
Returns a dictionary of metadata about that file.
'''
name = os.path.split(path)[1]
metadata = {} # Dict to hold filename:[categories] mappings
parsed = name.rstrip('.txt').split('-') # strip extension and split on hyphens
metadata['nation'] = parsed[0]
metadata['author'] = parsed[1]
metadata['title'] = parsed[2]
metadata['year'] = parsed[3]
metadata['gender'] = parsed[4]
return metadata
# %%time
m_lst = []
f_lst = []
for novel in novel_files:
with open(novel, 'r') as f:
novel_text = f.read() # Read a novel as a string
novel_label = os.path.split(novel)[1].rstrip('.txt') # Get convenience label for novel
# print(parse_filename(novel))
if parse_filename(novel)['gender'] == 'M':
m_lst.append((novel))
if parse_filename(novel)['gender'] == 'F':
f_lst.append((novel))
# +
sent_f = {'sent':[]}
sent_m = {'sent':[]}
for i in range(len(m_lst)):
with open(m_lst[i], 'r') as f:
m_txt = f.read()
m_label = os.path.split(m_lst[i])[1].rstrip('.txt')
tokenized = tokenize_text(m_txt)
for i in tokenized:
case = sentiment_score(i)
sent_m['sent'].append(case)
# -
for i in range(len(f_lst)):
with open(f_lst[i], 'r') as f:
f_txt = f.read()
f_label = os.path.split(f_lst[i])[1].rstrip('.txt')
tokenized = tokenize_text(f_txt)
for i in tokenized:
case = sentiment_score(i)
sent_f['sent'].append(case)
emo_m = {
'anger': 0.0,
'anticipation': 0.0,
'disgust': 0.0,
'fear': 0.0,
'joy': 0.0,
'negative': 0.0,
'positive': 0.0,
'sadness': 0.0,
'surprise': 0.0,
'trust': 0.0
}
for i in range(len(sent_m['sent'])):
for x in emo_m:
emo_m[x] += sent_m['sent'][i][x]
for i in emo_m:
emo_m[i]= emo_m[i]/len(sent_m['sent'])
print(emo_m)
emo_f= {
'anger': 0.0,
'anticipation': 0.0,
'disgust': 0.0,
'fear': 0.0,
'joy': 0.0,
'negative': 0.0,
'positive': 0.0,
'sadness': 0.0,
'surprise': 0.0,
'trust': 0.0
}
for i in range(len(sent_f['sent'])):
for x in emo_f:
emo_f[x] += sent_f['sent'][i][x]
for i in emo_f:
emo_f[i]= emo_f[i]/len(sent_f['sent'])
print(emo_f)
for i in emo_f:
if emo_f[i] > emo_m[i]:
print('emo_f higher')
else:
print('emo_m higher')
# # Analysis:
# I am not sure why this may be the case(maybe because there is an uneven number of corpuses), but it seems that the novels with a female author have a higher senitment score, on average.
| problem_sets/ps_03/ps_03_sentiment_jh976.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class DFA:
def __init__(self, states, finals, transitions, start=0):
self.states = states
self.start = start
self.current = start
self.finals = set(finals)
self.transitions = { state: {} for state in range(states) }
for (origin, symbol), destination in transitions.items():
self.transitions[origin][symbol] = destination
def _move(self, symbol):
# :solution:
if symbol not in self.transitions[self.current]:
return False
self.current = self.transitions[self.current][symbol]
return True
# :final:
# Your code here
pass
# :end:
def _reset(self):
self.current = self.start
def recognize(self, string):
# :solution:
self._reset()
for c in string:
if not self._move(c):
return False
return self.current in self.finals
# :final:
# Your code here
pass
# :end:
automaton = DFA(states=3, finals=[2], transitions={
(0,'a'): 1,
(1,'a'): 2,
(2,'a'): 2,
(0,'b'): 0,
(1,'b'): 0,
(2,'b'): 2
})
assert automaton.recognize('baab')
assert not automaton.recognize('abba')
| notebooks/2-automata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
# # Import dependencies
import tensorflow as tf
import tensorflow_io as tfio
from tensorflow.keras.layers.experimental.preprocessing import PreprocessingLayer
from typing import List, Optional
# # Augmentor
class SpecAugment(PreprocessingLayer):
def __init__(self,
freq_mask_prob: float = 0.5,
freq_mask_param: float = 10,
time_mask_prob: float = 0.5,
time_mask_param: float = 10):
self.freq_mask_prob = freq_mask_prob
self.freq_mask_param = freq_mask_param
self.time_mask_prob = time_mask_prob
self.time_mask_param = time_mask_param
def call(self, features):
prob = tf.random.uniform([])
augmented = tfio.audio.freq_mask(features, param=self.freq_mask_param)
features = tf.cond(prob >= self.freq_mask_prob,
lambda: augmented,
lambda: features)
prob = tf.random.uniform([])
augmented = tfio.audio.time_mask(features, param=self.time_mask_param)
features = tf.cond(prob >= self.time_mask_prob,
lambda: augmented,
lambda: features)
return features
# # Convolution Subsampling
# +
class ConvSubsampling(tf.keras.layers.Layer):
def __init__(self,
filters: List[int],
kernel_size: List[int] = [3, 3],
num_blocks: int = 1,
num_layers_per_block: int = 2,
dropout_rate: float = 0.0,
name: str = "ConvSubsampling",
**kwargs):
super(ConvSubsampling, self).__init__(name=name, **kwargs)
self.conv_blocks = tf.keras.Sequential()
for i in range(num_blocks):
convs = tf.keras.Sequential()
for _ in range(num_layers_per_block):
conv = tf.keras.layers.Conv2D(filters=filters[i],
kernel_size=kernel_size[i],
padding='same')
dropout = tf.keras.layers.Dropout(rate=dropout_rate)
relu = tf.keras.layers.ReLU()
convs.add(conv)
convs.add(dropout)
convs.add(relu)
self.conv_blocks.add(convs)
def call(self, inputs, training=False, **kwargs):
outputs = self.conv_blocks(inputs, training=training)
return outputs
batch_size, seq_len1, seq_len2, dim = 3, 1, 15, 512
a = tf.random.uniform([batch_size, seq_len1, seq_len2, dim],
minval=-40,
maxval=40)
conv_sub = ConvSubsampling(filters=[512, 512])
b = conv_sub(a)
print(b)
# -
# # Feed Forward Module
# +
class FeedForwardModule(tf.keras.layers.Layer):
def __init__(self,
ffn_dim: int,
dropout_rate: float = 0.4,
expansion_factor: int = 4,
output_reduction_factor: int = 0.5,
name: str = "FeedForwardModule",
**kwargs):
super(FeedForwardModule, self).__init__(name=name, **kwargs)
self.output_reduction_factor = output_reduction_factor
self.ffn = tf.keras.Sequential([
tf.keras.layers.LayerNormalization(),
tf.keras.layers.Dense(ffn_dim * expansion_factor),
tf.keras.layers.Activation(tf.nn.silu), # Swish activation with beta=1
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.Dense(ffn_dim),
tf.keras.layers.Dropout(dropout_rate)
])
self.add = tf.keras.layers.Add()
def call(self, inputs, training=False, **kwargs):
outputs = self.ffn(inputs, training=training)
outputs = self.add([inputs, outputs * self.output_reduction_factor])
return outputs
x = tf.random.uniform([batch_size, seq_len2, dim],
minval=-40,
maxval=40)
ffn_module = FeedForwardModule(ffn_dim=512)
y = ffn_module(x)
print(y)
# -
# # Convolution Module
class GLU(tf.keras.layers.Layer):
def __init__(self,
name: str = "GLU",
**kwargs):
super().__init__(name=name, **kwargs)
def call(self, inputs, **kwargs):
mat1, mat2 = tf.split(inputs, 2, axis=-1)
mat2 = tf.nn.sigmoid(mat2)
return tf.math.multiply(mat1, mat2)
# +
class ConvolutionModule(tf.keras.layers.Layer):
def __init__(self,
filters: int,
expansion_factor: int = 2,
kernel_size: int = 3,
dropout_rate: float = 0.4,
name: str = "ConvolutionModule",
**kwargs):
super(ConvolutionModule, self).__init__(name=name, **kwargs)
self.conv_module = tf.keras.Sequential([
tf.keras.layers.LayerNormalization(),
tf.keras.layers.Conv1D(filters=filters * expansion_factor, # Pointwise Conv
kernel_size=1),
GLU(),
tf.keras.layers.Conv1D(filters=filters, # 1D Depthwise Conv
kernel_size=kernel_size,
padding='same',
groups=filters),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation(tf.nn.silu),
tf.keras.layers.Conv1D(filters=filters, # Pointwise Conv
kernel_size=1),
tf.keras.layers.Dropout(rate=dropout_rate)
])
self.add = tf.keras.layers.Add()
def call(self, inputs, training=False, **kwargs):
outputs = self.conv_module(inputs, training=training)
outputs = self.add([inputs, outputs])
return outputs
x = tf.random.uniform([batch_size, seq_len2, dim],
minval=-40,
maxval=40)
cv = ConvolutionModule(filters=512)
y = cv(x)
print(y)
# -
# # Multi Headed Self-Attention Module
# +
class PositionalEncoding(tf.keras.layers.Layer):
"""
Implements the sinusoidal positional encoding function
Based on https://nlp.seas.harvard.edu/2018/04/03/attention.html#positional-encoding
"""
def __init__(self,
d_model: int = 512,
name: str = "PositionalEncoding",
**kwargs):
self.d_model = d_model
super(PositionalEncoding, self).__init__(name=name, **kwargs)
def build(self, input_shape):
d_model = input_shape[-1]
assert d_model == self.d_model, f"d_model must be equal to the last dimension of the input, which is {self.d_model}"
@staticmethod
def encode(max_len, d_model):
pe = tf.zeros([max_len, d_model])
position = tf.expand_dims(tf.range(0, max_len), axis=1)
position = tf.cast(position, dtype=tf.float32)
div_term = tf.math.exp(tf.range(0, d_model, 2, dtype=tf.float32) * -(tf.math.log(10000.0) / float(d_model)))
# Have to set up this way cause Tensorflow not allow assigning to EagerTensor
pe = tf.Variable(pe)
pe[:, 0::2].assign(tf.math.sin(position * div_term))
pe[:, 1::2].assign(tf.math.cos(position * div_term))
pe = tf.convert_to_tensor(pe)
pe = tf.expand_dims(pe, axis=0)
return pe
def call(self, inputs, **kwargs):
print(tf.shape(inputs))
max_len, d_model = tf.shape(inputs)[-2], tf.shape(inputs)[-1]
pe = self.encode(max_len, d_model)
# outputs = tf.math.add(inputs, pe)
return pe
pos = PositionalEncoding()
b = pos(a)
print(b)
# -
class RelativeMHA(tf.keras.layers.Layer):
"""
Multi-head Attention with Relative Positional Embedding
Based on https://github.com/sooftware/conformer/blob/main/conformer/attention.py
"""
def __init__(self,
num_heads: int = 8,
d_model: int = 512,
dropout_rate: float = 0.4,
name: str = "RelativeMHA",
**kwargs):
super(RelativeMHA, self).__init__(name=name, **kwargs)
self.num_heads = num_heads
self.d_model = d_model
self.d_head = d_model // num_heads
self.dropout = tf.keras.layers.Dropout(rate=dropout_rate)
self.query_linear = tf.keras.layers.Dense(d_model)
self.key_linear = tf.keras.layers.Dense(d_model)
self.value_linear = tf.keras.layers.Dense(d_model)
self.pos_linear = tf.keras.layers.Dense(d_model)
self.out_linear = tf.keras.layers.Dense(d_model)
self.u_bias = tf.Variable(tf.keras.initializers.HeUniform()([self.num_heads, self.d_head]))
self.v_bias = tf.Variable(tf.keras.initializers.HeUniform()([self.num_heads, self.d_head]))
def build(self, input_shape):
d_model = input_shape[-1]
assert d_model == self.d_model, f"d_model must be equal to the last dimension of the input, which is {self.d_model}"
assert d_model % self.num_heads == 0, f"num_heads must be divisible by {d_model}"
def call(self,
query: tf.Tensor,
key: tf.Tensor,
value: tf.Tensor,
pos_embedding: tf.Tensor,
training=False,
attention_mask: Optional[tf.Tensor] = None) -> tf.Tensor:
batch_size, seq_len = tf.shape(query)[0], tf.shape(query)[2]
query = tf.reshape(self.query_linear(query, training=training), [batch_size, -1, self.num_heads, self.d_head])
key = tf.transpose(tf.reshape(self.key_linear(key, training=training), [batch_size, -1, self.num_heads, self.d_head]), perm=[0, 2, 1, 3])
value = tf.transpose(tf.reshape(self.value_linear(value, training=training), [batch_size, -1, self.num_heads, self.d_head]), perm=[0, 2, 1, 3])
pos_embedding = tf.reshape(self.pos_linear(pos_embedding, training=training), [batch_size, -1, self.num_heads, self.d_head])
content_score = tf.linalg.matmul(tf.transpose(query + self.u_bias, perm=[0, 2, 1, 3]), tf.transpose(key, perm=[0, 1, 3, 2]))
pos_score = tf.linalg.matmul(tf.transpose(query + self.v_bias, perm=[0, 2, 1, 3]), tf.transpose(pos_embedding, perm=[0, 2, 3, 1]))
pos_score = self._relative_shift(pos_score)
score = (content_score + pos_score) / tf.math.sqrt(float(self.d_model))
if attention_mask is not None:
attention_mask = tf.expand_dims(attention_mask, axis=1)
score = tf.where(attention_mask, tf.fill(tf.shape(score), -1e9), score)
attn = tf.nn.softmax(score, axis=-1)
attn = self.dropout(attn, training=training)
context = tf.transpose(tf.linalg.matmul(attn, value), perm=[0, 2, 1, 3])
context = self.out_linear(tf.reshape(context, [batch_size, -1, seq_len, self.d_model]), training=training)
return context
@staticmethod
def _relative_shift(pos_score: tf.Tensor) -> tf.Tensor:
batch_size, num_heads, seq_len1, seq_len2 = tf.shape(pos_score)
zeros = tf.zeros([batch_size, num_heads, seq_len1, 1])
padded_pos_score = tf.concat([zeros, pos_score], axis=-1)
padded_pos_score = tf.reshape(padded_pos_score, [batch_size, num_heads, seq_len2 + 1, seq_len1])
pos_score = tf.reshape(padded_pos_score[:, :, 1:], tf.shape(pos_score))
return pos_score
class MultiHeadedSelfAttention(tf.keras.layers.Layer):
def __init__(self,
num_heads: int = 8,
d_model: int = 512,
dropout_rate: float = 0.4,
name: str = "MultiHeadedSelfAttention",
**kwargs):
super(MultiHeadedSelfAttention, self).__init__(name=name, **kwargs)
self.layer_norm = tf.keras.layers.LayerNormalization()
self.positional_encoding = PositionalEncoding(d_model)
self.attention = RelativeMHA(num_heads=num_heads, d_model=d_model, dropout_rate=dropout_rate)
self.dropout = tf.keras.layers.Dropout(dropout_rate)
def call(self, inputs: tf.Tensor, training=False, mask: Optional[tf.Tensor] = None) -> tf.Tensor:
batch_size = tf.shape(inputs)[0]
pos_embedding = self.positional_encoding(inputs)
pos_embedding = tf.concat([pos_embedding for _ in range(batch_size)], axis=0)
x = self.layer_norm(inputs, training=training)
x = self.attention(x, x, x, pos_embedding, training=training, attention_mask=mask)
x = self.dropout(x, training=training)
return x
class MHSAModule(tf.keras.layers.Layer):
def __init__(self,
head_size: int,
num_heads: int = 8,
d_model: int = 512,
dropout_rate: float = 0.4,
name: str = "MHSAModule",
**kwargs):
super(MHSAModule, self).__init__(name=name, **kwargs)
self.layer_norm = tf.keras.layers.LayerNormalization()
self.positional_encoding = PositionalEncoding(d_model)
self.attention = tf.keras.layers.MultiHeadAttention(num_heads=num_heads,
key_dim=head_size)
self.dropout = tf.keras.layers.Dropout(dropout_rate)
self.add = tf.keras.layers.Add()
def call(self, inputs: tf.Tensor, training=False, mask: Optional[tf.Tensor] = None) -> tf.Tensor:
batch_size = tf.shape(inputs)[0]
pos_embedding = self.positional_encoding(inputs)
pos_embedding = tf.concat([pos_embedding for _ in range(batch_size)], axis=0)
pos_embedding = tf.cast(pos_embedding, dtype=inputs.dtype)
outputs = self.layer_norm(inputs, training=training)
outputs = self.add([outputs, pos_embedding])
outputs = self.attention(outputs, outputs, outputs, attention_mask=mask, training=training)
outputs = self.dropout(outputs, training=training)
outputs = self.add([inputs, outputs])
return outputs
# # Conformer Block
class ConformerBlock(tf.keras.layers.Layer):
def __init__(self,
num_blocks: int = 1,
encoder_dim: int = 512,
num_heads: int = 8,
dropout_rate: float = 0.4,
name: str = "ConformerBlock",
**kwargs):
super(ConformerBlock, self).__init__(name=name, **kwargs)
self.num_blocks = num_blocks
self.ff_module = FeedForwardModule(encoder_dim)
self.attention = MultiHeadedSelfAttention(num_heads=num_heads, d_model=encoder_dim, dropout_rate=dropout_rate)
# self.attention = MHSAModule(head_size=encoder_dim,
# num_heads=num_heads,
# d_model=encoder_dim,
# dropout_rate=dropout_rate)
self.conv = ConvolutionModule(encoder_dim)
self.layer_norm = tf.keras.layers.LayerNormalization()
def call(self, inputs: tf.Tensor, training=False, mask: Optional[tf.Tensor] = None) -> tf.Tensor:
for _ in range(self.num_blocks):
x = self.ff_module(inputs, training=training)
x = self.attention(x, training=training, mask=mask)
x = self.conv(x, training=training)
x = self.ff_module(x, training=training)
x = self.layer_norm(x, training=training)
return x
# # Conformer
class ConformerEncoder(tf.keras.Model):
def __init__(self,
num_conv_filters: List[int],
num_blocks: int = 1,
encoder_dim: int = 512,
num_heads: int = 8,
dropout_rate: float = 0.4,
num_classes:int = 10,
include_top: bool = True,
name: str = "ConformerEncoder",
**kwargs):
super(ConformerEncoder, self).__init__(name=name, **kwargs)
self.include_top = include_top
self.conv_subsampling = ConvSubsampling(filters=num_conv_filters, dropout_rate=dropout_rate)
self.linear = tf.keras.layers.Dense(encoder_dim)
self.out_linear = tf.keras.layers.Dense(num_classes)
self.relu = tf.keras.layers.Activation(tf.nn.relu)
self.log_softmax = tf.keras.layers.Activation(tf.nn.log_softmax)
self.dropout = tf.keras.layers.Dropout(dropout_rate)
self.conformer_block = ConformerBlock(num_blocks=num_blocks, encoder_dim=encoder_dim, num_heads=num_heads, dropout_rate=dropout_rate)
def call(self, inputs: tf.Tensor, training=False, mask: Optional[tf.Tensor] = None) -> tf.Tensor:
x = self.conv_subsampling(inputs, training=training)
x = self.linear(x, training=training)
x = self.relu(x, training=training)
x = self.dropout(x, training=training)
x = self.conformer_block(x, training=training, mask=mask)
if self.include_top:
x = self.out_linear(x, training=training)
x = self.log_softmax(x, training=training)
return x
# # Test
# +
batch_size, seq_len, dim = 3, 15, 512
inputs = tf.random.uniform((batch_size, seq_len, dim),
minval=-40,
maxval=40)
model = ConformerEncoder(num_conv_filters=[512, 512], num_blocks=1, encoder_dim=512, num_heads=8, dropout_rate=0.4, num_classes=10, include_top=True)
# -
inputs = tf.expand_dims(inputs, axis=1)
outputs = model(inputs)
print(tf.shape(outputs))
| conformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hamming loss
import pandas as pd
from io import StringIO
from sklearn.metrics import hamming_loss
def hammingLoss(y_true, y_pred):
"""
Computes the hamming loss. Used for multiclass and multilabel classification.
"""
from sklearn import preprocessing
from sklearn.metrics import hamming_loss
import numpy as np
from itertools import chain
def to_2d_array(df):
MULTI_CLASS_CONDITION=True
_dict = {}
for [index,value] in df.as_matrix():
if index in _dict.keys():
_dict[index].append(value)
MULTI_CLASS_CONDITION = False
else:
_dict[index]=[value]
return list(_dict.keys()), list(_dict.values()), MULTI_CLASS_CONDITION
(y_true_keys, y_true_mat, b_multiclass) = to_2d_array(y_true)
(y_pred_keys, y_pred_mat, b_multiclass) = to_2d_array(y_pred)
assert y_true_keys==y_pred_keys
if b_multiclass:
# print('this is a multiclass case')
y_true_label_encoded = np.array(y_true_mat).ravel()
y_pred_label_encoded = np.array(y_pred_mat).ravel()
else: # MULTI_LABEL_CONDITION
# print('this is a multilabel case')
y_true_classes=(set(list(chain.from_iterable(y_true_mat))))
y_pred_classes=(set(list(chain.from_iterable(y_pred_mat))))
all_classes = list(y_true_classes.union(y_pred_classes))
lb = preprocessing.MultiLabelBinarizer(classes=all_classes)
y_true_label_encoded = lb.fit_transform(y_true_mat)
y_pred_label_encoded = lb.transform(y_pred_mat)
return hamming_loss(y_true_label_encoded, y_pred_label_encoded)
# +
# Testcase 1: MultiLabel, typical
y_true = pd.read_csv(StringIO("""
d3mIndex,class_label
3,happy-pleased
3,relaxing-calm
7,amazed-suprised
7,happy-pleased
13,quiet-still
13,sad-lonely
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,class_label
3,happy-pleased
3,sad-lonely
7,amazed-suprised
7,happy-pleased
13,quiet-still
13,happy-pleased
"""))
hammingLoss(y_true, y_pred)
# +
# Testcase 2: MultiLabel, Zero loss
y_true = pd.read_csv(StringIO("""
d3mIndex,class_label
3,happy-pleased
3,relaxing-calm
7,amazed-suprised
7,happy-pleased
13,quiet-still
13,sad-lonely
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,class_label
3,happy-pleased
3,relaxing-calm
7,amazed-suprised
7,happy-pleased
13,quiet-still
13,sad-lonely
"""))
hammingLoss(y_true, y_pred)
# +
# Testcase 3: MultiLabel, Complete loss
y_true = pd.read_csv(StringIO("""
d3mIndex,class_label
3,happy-pleased
3,relaxing-calm
7,amazed-suprised
7,happy-pleased
13,quiet-still
13,sad-lonely
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,class_label
3,ecstatic
3,sad-lonely
3,quiet-still
3,amazed-suprised
7,ecstatic
7,sad-lonely
7,relaxing-calm
7,quiet-still
13,ecstatic
13,happy-pleased
13,relaxing-calm
13,amazed-suprised
"""))
hammingLoss(y_true, y_pred)
# +
# Testcase 4: Multiclass, Typical
y_true = pd.read_csv(StringIO("""
d3mIndex,species
1,versicolor
2,versicolor
16,virginica
17,setosa
22,versicolor
26,versicolor
30,versicolor
31,virginica
33,versicolor
37,virginica
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,species
1,setosa
2,versicolor
16,virginica
17,setosa
22,versicolor
26,virginica
30,versicolor
31,virginica
33,versicolor
37,virginica
"""))
hammingLoss(y_true, y_pred)
# +
# Testcase 5: Multiclass, Zero loss
y_true = pd.read_csv(StringIO("""
d3mIndex,species
1,versicolor
2,versicolor
16,virginica
17,setosa
22,versicolor
26,versicolor
30,versicolor
31,virginica
33,versicolor
37,virginica
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,species
1,versicolor
2,versicolor
16,virginica
17,setosa
22,versicolor
26,versicolor
30,versicolor
31,virginica
33,versicolor
37,virginica
"""))
hammingLoss(y_true, y_pred)
# +
# Testcase 6: Multiclass, Complete loss
y_true = pd.read_csv(StringIO("""
d3mIndex,species
1,versicolor
2,versicolor
16,versicolor
17,virginica
22,versicolor
26,versicolor
30,versicolor
31,virginica
33,versicolor
37,virginica
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,species
1,setosa
2,setosa
16,setosa
17,setosa
22,setosa
26,setosa
30,setosa
31,setosa
33,setosa
37,setosa
"""))
hammingLoss(y_true, y_pred)
# -
# # RMSE
def rootMeanSquaredError(y_true, y_pred):
"""
Computes the root mean squared error, for both univariate and multivariate case
"""
import numpy as np
from sklearn.metrics import mean_squared_error
from math import sqrt
rmse = None
# perform some checks
assert 'd3mIndex' in y_true.columns
assert 'd3mIndex' in y_pred.columns
assert y_true.shape == y_pred.shape
# preprocessing
y_true.set_index('d3mIndex', inplace=True)
y_pred.set_index('d3mIndex', inplace=True)
# determine the dimension
y_true_dim=y_true.shape[1]
# univariate case
if y_true_dim == 1:
y_true_array = y_true.as_matrix().ravel()
y_pred_array = y_pred.as_matrix().ravel()
mse = mean_squared_error(y_true, y_pred)
rmse = sqrt(mse)
# multivariate case
elif y_true_dim > 1:
y_true_array = y_true.as_matrix()
y_pred_array = y_pred.as_matrix()
mse = mean_squared_error(y_true_array, y_pred_array, multioutput='uniform_average')
rmse = sqrt(mse)
return rmse
# +
# test case 1
# y_true_uni=[3, -1., 2, 7]
# y_pred_uni=[2.1, 0.0, 2, 8]
# expected rmse = 0.8381527307120105
y_true = pd.read_csv(StringIO("""
d3mIndex,value
1,3
2,-1.0
16,2
17,7
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,value
1,2.1
2,0.0
16,2
17,8
"""))
rootMeanSquaredError(y_true, y_pred)
# +
# test case 2
# y_true_multi=[[0.5, 1],[-1, 1],[7, -6]]
# y_pred_multi=[[0, 2],[-1, 2],[8, -5]]
# expected rmse = 0.8416254115301732
y_true = pd.read_csv(StringIO("""
d3mIndex,value1, value2
1,0.5,1
2,-1,1
16,7,-6
"""))
y_pred = pd.read_csv(StringIO("""
d3mIndex,value1,value2
1,0,2
2,-1,2
16,8,-5
"""))
rootMeanSquaredError(y_true, y_pred)
# -
# # Object detection average precision
# +
def group_gt_boxes_by_image_name(gt_boxes):
gt_dict: typing.Dict = {}
for box in gt_boxes:
image_name = box[0]
bounding_polygon = box[1:]
bbox = convert_bouding_polygon_to_box_coords(bounding_polygon)
if image_name not in gt_dict.keys():
gt_dict[image_name] = []
gt_dict[image_name].append({'bbox': bbox})
return gt_dict
def convert_bouding_polygon_to_box_coords(bounding_polygon):
# box_coords = [x_min, y_min, x_max, y_max]
box_coords = [bounding_polygon[0], bounding_polygon[1],
bounding_polygon[4], bounding_polygon[5]]
return box_coords
def voc_ap(rec, prec):
import numpy
# First append sentinel values at the end.
mrec = numpy.concatenate(([0.], rec, [1.]))
mpre = numpy.concatenate(([0.], prec, [0.]))
# Compute the precision envelope.
for i in range(mpre.size - 1, 0, -1):
mpre[i - 1] = numpy.maximum(mpre[i - 1], mpre[i])
# To calculate area under PR curve, look for points
# where X axis (recall) changes value.
i = numpy.where(mrec[1:] != mrec[:-1])[0]
# And sum (\Delta recall) * prec.
ap = numpy.sum((mrec[i + 1] - mrec[i]) * mpre[i + 1])
return float(ap)
def object_detection_average_precision(y_true, y_pred):
"""
This function takes a list of ground truth bounding polygons (rectangles in this case)
and a list of detected bounding polygons (also rectangles) for a given class and
computes the average precision of the detections with respect to the ground truth polygons.
Parameters:
-----------
y_true: list
List of ground truth polygons. Each polygon is represented as a list of
vertices, starting in the upper-left corner going counter-clockwise.
Since in this case, the polygons are rectangles, they will have the
following format:
[image_name, x_min, y_min, x_min, y_max, x_max, y_max, x_max, y_min].
y_pred: list
List of bounding box polygons with their corresponding confidence scores. Each
polygon is represented as a list of vertices, starting in the upper-left corner
going counter-clockwise. Since in this case, the polygons are rectangles, they
will have the following format:
[image_name, x_min, y_min, x_min, y_max, x_max, y_max, x_max, y_min, confidence_score].
Returns:
--------
ap: float
Average precision between detected polygons (rectangles) and the ground truth polylgons (rectangles).
(it is also the area under the precision-recall curve).
Example 1:
>> predictions_list_1 = [['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 5, 10, 5, 20, 20, 20, 20, 10, 0.9],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6]]
>> ground_truth_list_1 = [['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]]
>> ap_1 = object_detection_average_precision(ground_truth_list_1, predictions_list_1)
>> print(ap_1)
0.667
Example 2:
>> predictions_list_2 = [['img_00285.png', 330, 463, 330, 505, 387, 505, 387, 463, 0.0739],
['img_00285.png', 420, 433, 420, 498, 451, 498, 451, 433, 0.0910],
['img_00285.png', 328, 465, 328, 540, 403, 540, 403, 465, 0.1008],
['img_00285.png', 480, 477, 480, 522, 508, 522, 508, 477, 0.1012],
['img_00285.png', 357, 460, 357, 537, 417, 537, 417, 460, 0.1058],
['img_00285.png', 356, 456, 356, 521, 391, 521, 391, 456, 0.0843],
['img_00225.png', 345, 460, 345, 547, 415, 547, 415, 460, 0.0539],
['img_00225.png', 381, 362, 381, 513, 455, 513, 455, 362, 0.0542],
['img_00225.png', 382, 366, 382, 422, 416, 422, 416, 366, 0.0559],
['img_00225.png', 730, 463, 730, 583, 763, 583, 763, 463, 0.0588]]
>> ground_truth_list_2 = [['img_00285.png', 480, 457, 480, 529, 515, 529, 515, 457],
['img_00285.png', 480, 457, 480, 529, 515, 529, 515, 457],
['img_00225.png', 522, 540, 522, 660, 576, 660, 576, 540],
['img_00225.png', 739, 460, 739, 545, 768, 545, 768, 460]]
>> ap_2 = object_detection_average_precision(ground_truth_list_2, predictions_list_2)
>> print(ap_2)
0.125
Example 3:
>> predictions_list_3 = [['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6],
['img_00002.png', 5, 8, 5, 16, 15, 16, 15, 8, 0.9],
['img_00002.png', 11, 12, 11, 18, 21, 18, 21, 12, 0.9]]
>> ground_truth_list_3 = [['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]]
>> ap_3 = object_detection_average_precision(ground_truth_list_3, predictions_list_3)
>> print(ap_3)
0.444
Example 4:
(Same as example 3 except the last two box predictions in img_00002.png are switched)
>> predictions_list_4 = [['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6],
['img_00002.png', 11, 12, 11, 18, 21, 18, 21, 12, 0.9],
['img_00002.png', 5, 8, 5, 16, 15, 16, 15, 8, 0.9]]
>> ground_truth_list_4 = [['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]]
>> ap_4 = object_detection_average_precision(ground_truth_list_4, predictions_list_4)
>> print(ap_4)
0.444
"""
"""
This function is different from others because ``y_true`` and ``y_pred`` are not vectors but arrays.
"""
import numpy
ovthresh = 0.5
# y_true = typing.cast(Truth, unvectorize(y_true))
# y_pred = typing.cast(Predictions, unvectorize(y_pred))
# Load ground truth.
gt_dict = group_gt_boxes_by_image_name(y_true)
# Extract gt objects for this class.
recs = {}
npos = 0
imagenames = sorted(gt_dict.keys())
for imagename in imagenames:
Rlist = [obj for obj in gt_dict[imagename]]
bbox = numpy.array([x['bbox'] for x in Rlist])
det = [False] * len(Rlist)
npos = npos + len(Rlist)
recs[imagename] = {'bbox': bbox, 'det': det}
# Load detections.
det_length = len(y_pred[0])
# Check that all boxes are the same size.
for det in y_pred:
assert len(det) == det_length, 'Not all boxes have the same dimensions.'
image_ids = [x[0] for x in y_pred]
BP = numpy.array([[float(z) for z in x[1:-1]] for x in y_pred])
BB = numpy.array([convert_bouding_polygon_to_box_coords(x) for x in BP])
confidence = numpy.array([float(x[-1]) for x in y_pred])
boxes_w_confidences_list = numpy.hstack((BB, -1 * confidence[:, None]))
boxes_w_confidences = numpy.empty((boxes_w_confidences_list.shape[0],),
dtype=[('x_min', float), ('y_min', float),
('x_max', float), ('y_max', float),
('confidence', float)])
boxes_w_confidences[:] = [tuple(i) for i in boxes_w_confidences_list]
# Sort by confidence.
#sorted_ind = numpy.argsort(-confidence)
sorted_ind = numpy.argsort(
boxes_w_confidences, kind='mergesort',
order=('confidence', 'x_min', 'y_min'))
BB = BB[sorted_ind, :]
image_ids = [image_ids[x] for x in sorted_ind]
# Go down y_pred and mark TPs and FPs.
nd = len(image_ids)
tp = numpy.zeros(nd)
fp = numpy.zeros(nd)
for d in range(nd):
R = recs[image_ids[d]]
bb = BB[d, :].astype(float)
ovmax = -numpy.inf
BBGT = R['bbox'].astype(float)
if BBGT.size > 0:
# Compute overlaps.
# Intersection.
ixmin = numpy.maximum(BBGT[:, 0], bb[0])
iymin = numpy.maximum(BBGT[:, 1], bb[1])
ixmax = numpy.minimum(BBGT[:, 2], bb[2])
iymax = numpy.minimum(BBGT[:, 3], bb[3])
iw = numpy.maximum(ixmax - ixmin + 1., 0.)
ih = numpy.maximum(iymax - iymin + 1., 0.)
inters = iw * ih
# Union.
uni = ((bb[2] - bb[0] + 1.) * (bb[3] - bb[1] + 1.) +
(BBGT[:, 2] - BBGT[:, 0] + 1.) *
(BBGT[:, 3] - BBGT[:, 1] + 1.) - inters)
overlaps = inters / uni
ovmax = numpy.max(overlaps)
jmax = numpy.argmax(overlaps)
if ovmax > ovthresh:
if not R['det'][jmax]:
tp[d] = 1.
R['det'][jmax] = 1
else:
fp[d] = 1.
else:
fp[d] = 1.
# Compute precision recall.
fp = numpy.cumsum(fp)
tp = numpy.cumsum(tp)
rec = tp / float(npos)
# Avoid divide by zero in case the first detection matches a difficult ground truth.
prec = tp / numpy.maximum(tp + fp, numpy.finfo(numpy.float64).eps)
ap = voc_ap(rec, prec)
return ap
if __name__ == "__main__":
predictions_list_1 = [
['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 5, 10, 5, 20, 20, 20, 20, 10, 0.9],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6]
]
ground_truth_list_1 = [
['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]
]
ap_1 = object_detection_average_precision(
ground_truth_list_1, predictions_list_1)
print('TEST CASE 1 --- AP: ', ap_1)
predictions_list_2 = [
['img_00285.png', 330, 463, 330, 505, 387, 505, 387, 463, 0.0739],
['img_00285.png', 420, 433, 420, 498, 451, 498, 451, 433, 0.0910],
['img_00285.png', 328, 465, 328, 540, 403, 540, 403, 465, 0.1008],
['img_00285.png', 480, 477, 480, 522, 508, 522, 508, 477, 0.1012],
['img_00285.png', 357, 460, 357, 537, 417, 537, 417, 460, 0.1058],
['img_00285.png', 356, 456, 356, 521, 391, 521, 391, 456, 0.0843],
['img_00225.png', 345, 460, 345, 547, 415, 547, 415, 460, 0.0539],
['img_00225.png', 381, 362, 381, 513, 455, 513, 455, 362, 0.0542],
['img_00225.png', 382, 366, 382, 422, 416, 422, 416, 366, 0.0559],
['img_00225.png', 730, 463, 730, 583, 763, 583, 763, 463, 0.0588],
]
ground_truth_list_2 = [
['img_00285.png', 480, 457, 480, 529, 515, 529, 515, 457],
['img_00285.png', 480, 457, 480, 529, 515, 529, 515, 457],
['img_00225.png', 522, 540, 522, 660, 576, 660, 576, 540],
['img_00225.png', 739, 460, 739, 545, 768, 545, 768, 460],
]
ap_2 = object_detection_average_precision(
ground_truth_list_2, predictions_list_2)
print('TEST CASE 2 --- AP: ', ap_2)
predictions_list_3 = [
['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6],
['img_00002.png', 5, 8, 5, 16, 15, 16, 15, 8, 0.9],
['img_00002.png', 11, 12, 11, 18, 21, 18, 21, 12, 0.9]
]
ground_truth_list_3 = [
['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]
]
ap_3 = object_detection_average_precision(
ground_truth_list_3, predictions_list_3)
print('TEST CASE 3 --- AP: ', ap_3)
predictions_list_4 = [
['img_00001.png', 110, 110, 110, 210, 210, 210, 210, 110, 0.6],
['img_00002.png', 120, 130, 120, 200, 200, 200, 200, 130, 0.6],
['img_00002.png', 11, 12, 11, 18, 21, 18, 21, 12, 0.9],
['img_00002.png', 5, 8, 5, 16, 15, 16, 15, 8, 0.9]
]
ground_truth_list_4 = [
['img_00001.png', 100, 100, 100, 200, 200, 200, 200, 100],
['img_00002.png', 10, 10, 10, 20, 20, 20, 20, 10],
['img_00002.png', 70, 80, 70, 150, 140, 150, 140, 80]
]
ap_4 = object_detection_average_precision(
ground_truth_list_4, predictions_list_4)
print('TEST CASE 4 --- AP: ', ap_4)
# -
| datasets/data-supply/documentation/code/consolidated-new-metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Content :
# - Initiating A String
# - Concatenating Strings
# - Adjusting Cases
# - String Methods
# - Regex
# [recording](https://drive.google.com/drive/folders/15PfOCaTMCPRepwMHC8y0gyPIIPL3pQVG?usp=sharing)
# Why do I need to know how to work with strings?
# - Clean a dataset
# - Prepare your data for a machine learning algorithm
# - Parse specific data from a website
#
# ### Initiating a string
name = "sam"
name1 = 'sam'
wrong_name = 'Sam'
# - Length of a string -> `len`
len('name')
# - Converting another data type to a string -> `str`
str(2.9)
# ### Concatenating Strings
# We can concatenate (add) two strings together with the `+` operator.
greeting = "Hello"
best_programming_language = "Python"
print(greeting + " " + best_programming_language)
# 🤔What if we wanted to concatenate "I'm" + age + "years old"?
#Insert your code here
print("I'm" + " " + "40"+ " " + "Years Old")
# ### String Indexing
from IPython import display
display.Image("string_indexing.png")
sth = "hiiii???@@**&&*"
type(sth)
example = "MY_STRING"
example[0]
example[-9]
example[1:4]
example[::-1]
# ## Adjusting Cases
# - lower( ) and upper( )
cat = "mEoWW hoomAn"
cat.lower()
cat.upper()
# *why would I ever need this in data science?*
# - capitalize( )
cat.capitalize()
cat.title()
# ### split( )
examples = "I'm a sample string"
examples.split()
# `split()` splittes the string into a list of **substrings**
# We can control how we split a string with a couple of parameters that come with our method.
examples.split(sep=" ", maxsplit = 2)
# ### splitlines( )
# Usually, when we have a long text, defferent lines are separted by `\n`.
#
# for example :
#
# **"I'm the first line\n and I'm the second line"**
sentence = "I'm the first line\nand I'm the second line"
sentences = sentence.splitlines()
sentences[1]
type(sentences)
# ### joining( )
# As you see till now, when we use methods like `.splitlines()`, the result is a **list**. so we can't use string methods on them anymore.
#
# We can solve this problem by using `.join()`
# take `sentences` that we created just now.
type(sentences)
sentences_str.lower()
sentences_str = " ".join(sentences)
# Let's see the result:
sentences_str
# Let's check the type :
type(sentences_str)
# ### strip( )
# Usually, our text datasets are not clean. meaning they usually look like this:
# **" The goverment did something sketchy again\n"**
news = " The goverment did something sketchy again\n"
# We can use `.strip()` to remove both the **leading** and **trailing** characters.
news.strip()
# - rstrip( )
news.rstrip()
# - lstrip( )
news.lstrip()
# ### replace( )
# We have the following string:
text = "The Game Of Thrones series is my favorite series of all time.The Game Of Thrones series has the best music as well."
# well... after that horrible ending we no longer stan game of thrones. now dark is our favorite series.
# We can find every occurance of `game of thrones`, and replace them with `replace()`
text = text.lower()
text = text.replace('game of thrones', 'dark')
text
# What would happen if we didn't lower( ) before using `replace()`?
# **✨Task**
# - Use `lower()` on *review* and store the result in a variable called `review_lower`
# - Use `.splitlines()` on `review_lower` to divide the two different sentences. Store the result in a variable called `sentences`
# - Check the *type* of sentences
# - Turn sentences into a string again. (use `.join`)
# - Replace *hair dryer* with *laptop*
review = "I really liked using this hair dryer\nThe price of this hair dryer was nice as well."
#Insert your code here
review_lower = review.lower()
sentences = review_lower.splitlines()
sentences = " ".join(sentences)
sentences.replace("hair dryer", "laptop")
# ## String formatting
# When we calculate a certain variable, we want to show-case it to the use at some point.
custom_string ="string formatting"
print(f"{custom_string} is a powerful technique.")
# **✨Task**
# - print -> Square with a side {side} has the area of {area}
side = 21
area = side**2
#Insert your code here
print(f"Square with a side {side} has the area of {area}")
# ## REGEX (Regular Expressions)
# `A regular expression is a sequence of characters that specifies a search pattern.`
# First we need to import the module **re**
import re
# - Find all matches of a pattern :
text = "#data science has been a fast growing job! I love #data science! yayyyyy."
re.findall(r"#data science", text )
# - Split String at each match :
re.split(r"!", text)
# - replace a single match or many matches with a string :
re.sub(r"#data\sscience", "cryptography", text)
# ***
# First of all, we have something called `Metacharacters` which mean -> match something specific.
#
# For example :
# - **\d** -> Matches any digit, ie. 0-9
# - **\D** -> Matches any non-digit character
# - **\s** -> Matches any whitespace character
# - **\w** -> Matches any digit or normal character
# - **\W** -> Matches any Non word
# - Finding all matches of **user** which have a number followed them :
text = "The winners are : user9, userN, user8"
re.findall(r"user\d", text)
# **✨Task**
# - Find all matches of **user** followed by a non-digit character :
# +
#insert your code here
# -
# - Find all matches of **user** followed by any digit or normal character:
#insert your code here
re.findall(r"user\w", text)
# **✨Task**
# - Find the price ($7) in the following string using REGEX
# hint : you can use two metacharacters consecutively.
skirt = "This skirt is on sale and it's $7 or €9 code N6 now"
#insert your code here
re.findall(r"\W\d", skirt)
# **✨Task**
# The company that you are working for asked you to perform a sentiment analysis using a dataset with tweets. First of all, you need to do some cleaning and extract some information.
# While printing out some text, you realize that some tweets contain user mentions. Some of these mentions follow a very strange pattern. A few examples that you notice: @robot3!, @robot5& and @robot7#
# - Import the re module.
# - Write a regex that matches the user mentions that starts with @ and follows the pattern, e.g. @robot3!.
# - Find all the matches of the pattern in the `sentiment_analysis` variable.
sentiment_analysis = "@robot9! @robot4& I have a good feeling that the show isgoing to be amazing! @robot9$ @robot7%"
#insert your code here
pattern = r"@robot\d\W"
re.findall(pattern, sentiment_analysis)
# ## Additional Stuff :
# [Regex Essentials](https://towardsdatascience.com/the-essentials-of-regular-expressions-b52af8fe271a)
#
# [9 Most Essential REGEX Patterns](https://appcodelabs.com/stop-being-afraid-of-regular-expressions-9-essential-patterns-you-should-learn)
| week-3/strings/Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import skallel_tensor
from skallel_tensor import genotypes_count_alleles
from numba import cuda
skallel_tensor.__version__
# # Smaller dataset (400MB)
# simulate some genotype data
gt = np.random.choice(np.array([-1, 0, 1, 2, 3], dtype='i1'),
p=[.01, .7, .2, .08, .01],
size=(200_000, 1_000, 2))
gt.nbytes / 1e6
gt.dtype
gt
gt.shape
# ## Single CPU core
# warm-up jit
genotypes_count_alleles(gt, max_allele=3)
# %%timeit -n1 -r3
genotypes_count_alleles(gt, max_allele=3)
# ## Multiple CPU cores via Dask
import dask.array as da
gt_dask = da.from_array(gt, chunks=(10_000, None, None))
gt_dask
# %%timeit -n1 -r5
genotypes_count_alleles(gt_dask, max_allele=3).compute()
# ## Single GPU
# !nvidia-smi | head
# %%time
gt_cuda = cuda.to_device(gt)
gt_cuda
# warm up jit
genotypes_count_alleles(gt_cuda, max_allele=3).copy_to_host()
# %%timeit -n1 -r5
genotypes_count_alleles(gt_cuda, max_allele=3)
cuda.synchronize()
ac_cuda = genotypes_count_alleles(gt_cuda, max_allele=3)
ac_cuda
# %time ac_cuda.copy_to_host()
# ## GPU via Dask
gt_dask_cuda = gt_dask.map_blocks(cuda.to_device)
genotypes_count_alleles(gt_dask_cuda, max_allele=3).compute()
# %%timeit -n1 -r3
genotypes_count_alleles(gt_dask_cuda, max_allele=3).compute()
# # Larger dataset (4GB)
# +
# gt_random_big = da.random.choice(
# np.array([-1, 0, 1, 2, 3], dtype='i1'),
# p=[.01, .7, .2, .08, .01],
# size=(2_000_000, 1_000, 2),
# chunks=(50_000, None, None))
# gt_random_big
# +
# gt_random_big.to_zarr('example.zarr', component='gt')
# -
import zarr
gt_big_zarr = zarr.open('example.zarr')['gt']
gt_big_zarr.info
gt_big_dask = da.from_array(gt_big_zarr)
gt_big_dask
# ## Multiple CPUs via Dask
# %%time
genotypes_count_alleles(gt_big_dask, max_allele=3).compute()
# ## Single GPU via Dask
gt_big_dask_cuda = gt_big_dask.map_blocks(cuda.to_device)
# %%time
genotypes_count_alleles(gt_big_dask_cuda, max_allele=3).compute(num_workers=1)
# %%time
genotypes_count_alleles(gt_big_dask_cuda, max_allele=3).compute(num_workers=2)
# %%time
genotypes_count_alleles(gt_big_dask_cuda, max_allele=3).compute(num_workers=3)
# %%time
genotypes_count_alleles(gt_big_dask_cuda, max_allele=3).compute(num_workers=4)
# Interesting thing, even with just a single GPU, we can improve performance and get better saturation by having multiple Dask workers.
| examples/genotypes_count_alleles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "-"}
# nbi:hide_in
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import nbinteract as nbi
from IPython.display import display
import matplotlib.pyplot as plt
import numpy as np
# -
# # <span style="color:blue"> Mediciones y Errores <span>
#
# ## <span style="color:blue">Introducción <span>
#
# Uno de los primeros conceptos desarrollados por el hombre fue el de número, ya que tenía la necesidad de poder expresar numéricamente todo lo que se encontraba a su alrededor. El ser humano necesitaba comparar objetos como animales y alimentos o eventos como estaciones del año y temperatura, que le permitieran desarrollar la agricultura.
#
# Mas tarde usó la medición como un forma de entender el mundo que nos rodea y expandir las fronteras del conocimiento, cosa que seguimos haciendo hasta el día de hoy.
#
# Gracias a la medición, el hombre pudo construir refugios, navegar por lugares desconocidos lejos de las costas, construir los grandes monumentos que hasta el día de hoy se mantienen en pie como las pirámides y el Coliseo Romano. Ya en nuestra era nos permitió viajar al espació y a otro planetas, nos ayuda a entender el mundo en el que estamos, y hasta entender cosas que no vemos pero si podemos medir como las partículas subatómicas.
#
# La medición es una herramienta fundamental para nuestra vida diaria y profesional, por tal motivo es necesario entender los fundamentos de la misma.
#
#
# ## <span style="color:blue">¿Qué es medir?<span>
#
# En nuestra vida cotidiana estamos acostumbrados a medir todo el tiempo, aunque nunca nos tomamos el tiempo de reflexionar que es lo que estamos haciendo. Medimos nuestra altura, cuanto tardamos en ir al colegio, cuanto tiempo tardamos en realizar una tarea, la cantidad correcta de ingredientes para seguir una receta, etc.
#
# Si nos detenemos un momento podemos darnos cuenta que al medir estamos comparando una cosa con otra, estamos comparando una **magnitud** con otra.
#
# ### <span style="color:blue">¿Qué es Magnitud?<span>
#
# La magnitud son los atributos o características de un objeto, fenómeno o una sustancia susceptible de ser medido. Existen magnitudes básicas y derivadas, que constituyen ejemplos de magnitudes físicas: la masa, la longitud, el tiempo, la carga eléctrica, la densidad, la temperatura, la velocidad, la aceleración y la energía, etc.
#
# Las magnitudes se representan con símbolos y normalmente las denominamos **variables**
#
# ### <span style="color:blue">¿Y cantidad? <span>
# La cantidad, son las magnitudes de un cuerpo, fenómeno o una sustancia en particular que que permite comparar cualitativamente respecto de la que se tomó como unidad de la magnitud, por ejemplo la longitud de una mesa en particular **"mi escritorio"**, es una cantidad. Otro ejemplo sería la masa de **mi** libro de analógica es una **cantidad** de masa.
#
#
# Podemos definir a la **_medida_** como:
# <div class="alert alert-block alert-info">
# <b>Medir</b> una cantidad <b>X</b> (o variable x) es <b>compararla</b> con otra cantidad arbitraria <b>U</b> de la misma magnitud denominada <b> Unidad </b> (simbólicamente se indica como el cociente $\frac{A}{U}$). El resultado, es el número de veces que la cantidad contiene a la unidad, es un número real denominado <b> Medida</b> que no es otra cosa que el valor de la cantidad
#
# $$ \boxed {A = X * U} $$
#
# donde:
# $$$$
# * <b>A</b>: es la es el valor de la medida
# $$$$
# * <b>X</b>: es la cantidad de la medida
# $$$$
# * <b>U</b>: es la unidad con la que estamos comparando.
# </div>
#
# Podemos ver esta misma ecuación como $$\boxed{X = \frac{A}{U}}$$ donde queda mas claro que la cantidad se obtiene de comparar el valor de nuestra medida con un patrón que es la unidad.
#
# A lo largo de este apunte realizaremos muchos ejemplos para que las cosas que vista desde su definición parecen complejas, se entiendan.
#
#
# <div class="alert alert-block alert-success">
# <b> Ejemplo n° 1</b>
# Si tomamos una resistencia de valor 220 $\Omega$, $$R = \frac{220\Omega}{1\Omega}$$, esto quiere decir que la cantidad de la medida es <b>220</b>
# </div>
#
# Lo que hacemos durante el proceso de medición es comparar una cantidad de una magnitud de un objeto con un patrón preestablecido de esa magnitud, en este caso ohms.
#
# ## <span style="color:blue">Proceso de medición</span>
#
# Medir no representa en la mayoría de los casos una tarea sencilla. La **_Metrología_** es la ciencia que estudia las mediciones de las magnitudes garantizando su normalización mediante la trazabilidad. Se encargan de diseñar el proceso de medición mas indicado para cada caso, reduciendo las **incertidumbres** de las mediciones. También se encargan de definir los patrones que utilizamos para medir como el metro, Kelvin, Ampere, etc.
#
# Podemos definir tres sistemas que se encuentran en todo proceso de medición:
#
# * <span style="color:red"><b>Sistema objeto</b></span> Es lo que se desea medir. COnsiste en una cantidad de cierta magnitud física. Por ejemplo medir la longitud de una maesa.
#
# * <span style="color:red"><b>Sistema de medición</b></span> Es el dispositivo con el que se efectuará la medición. La elección de este dispositivo depende la magnitud a medir y de cuan precisa debe ser la medida. Por ejemplo para medir una longitud pequeña se puede usar una regla, un calibre o un micrómetro. Otro ejemplo puede ser que para medir tiempo podemos utilizar un reloj o un cronómetro.
#
# * <span style="color:red"><b>Operador</b></span> El operador es la persona que se encarga de llevar a cavo la medición en forma **correcta**, efectuando las lecturas en la escala del instrumento utilizado. Para lograr esto es necesario conocer como utilizar el instrumento y las unidades. Por ejemplo, para medir la longitud de una varilla, se debe ubicar el extremo izquierdo de la misma en coincidencia con el cero de la regla y ver en el otro extremo con que división de la regla coincide.
#
# <img src="Imagenes/sistema.png" width="600" height="360">
#
# ### <span style="color:blue">Protocolo de medición</span>
#
# Antes de medir nos tenemos que hacer estas preguntas
#
# * <span style="color:red"><b>¿Para qué medimos?</b></span> Debemos saber cual es el objetivo de la medición para saber que precisión(vamos a ver esta palabra mas adelante) necesitamos.
#
# * <span style="color:red"><b>¿Qué medimos?</b></span> Tenemos que seleccionar las <b>magnitudes</b> adecuadas de acuerdo a nuestro objetivo.
#
# * <span style="color:red"><b>¿Cómo se mide?</b></span> Debemos seleccionar el/los instrumentos necesarios.
#
# * <span style="color:red"><b>*¿Cómo vamos a medir?</b></span> Tenemos que establecer los pasos a seguir para determinar la magnitud correspondiente.
#
# * <span style="color:red"><b>¿Cuantas veces se mide?</b></span> Hay que medir varias veces para minimizar los <span style="color:red"><b>errores</b></span>
#
# ### <span style="color:blue"><b>¿Errores?</b></span>
#
# Tal vez seria mas correcto llamarlos incertezas, pero también reciben el nombre de errores. Como vimos al medir una magnitud física, tenemos diferentes forma de hacerlo y es muy probable que al medirla de las distintas formas, el resultado de cada medición sea distinta, incluso si medimos varias veces de la misma forma. ¿Esto quiere decir que los resultados son incorrectos?, ¿Alguno de ellos es correcto? ¿Qué valor tomo como resultado de la medición?
#
# La respuesta a la primera pregunta es no, solo significa que tenemos un cierto grado de incerteza en todas las medidas que realicemos, lo que quiere decir que **nunca vamos a conocer el valor verdadero de una medición**. La respuesta a la segunda pregunta es puede ser que si y puede ser que no, pero nunca lo vamos a saber. La tercer pregunta la vamos a responder un poco mas adelante.
#
# ## <span style="color:red"><b>Podemos concluir que al realizar una medición podemos establecer el valor verdadero de la misma entre ciertos límites pero nunca su valor verdadero. Esto se debe a que hay múltiples factores que afectan a la medición y estudiaremos en detalle en la siguiente sección.</b></span>
#
# ### Podemos definir al **_error_** como:
# <div class="alert alert-block alert-info">
# El grado de incerteza de una medición debido a las limitaciones inevitables de los instrumentos y del observador, a condiciones exteriores por lo general fluctuantes. Impidiendo que la medición sea perfecta y solo permitiéndonos caracterizar a la medida por un intervalo llamado <span style="color:brown"><b>intervalo de indeterminación o incertidumbre</b></span> dentro del cual se puede afirmar con un alto grado de certeza que se halla el valor verdadero
# </div>
#
# En la siguiente imagen se puede observar un intervalo de incerteza que se limita por los valores máximos y mínimos que puede tomar la medición.
# <img src="Imagenes/medida1.png" width="600" height="300">
#
# Podemos mejorar la calidad de la medición reduciendo el intervalo de indeterminación, utilizando un instrumento mas preciso, por ejemplo usando un calibre en vez de una regla.
#
# <img src="Imagenes/medida2.png" width="600" height="300">
#
# Dependiendo de lo que estemos midiendo no siempre es necesario hacerlo, por ejemplo al usar cuando un cocinero utiliza un termómetro para controlar la temperatura, una incertidumbre de 1°C es suficiente, no necesitan medir la temperatura con décimas o milésimas de grados, lo cual sumaría un costo mayor sin ningún tipo de beneficio.
#
# <div class="alert alert-block alert-warning">
# Matemáticamente lo podemos expresar como:
#
# $$ \boxed{x_{min} \leq x \leq x_{máx}}$$
#
# donde <b>x</b> es el valor verdadero de la medida.
# </div>
#
# Para terminar, toda medida debe expresarse de la siguiente forma:
#
# <div class="alert alert-block alert-warning">
# Matemáticamente lo podemos expresar como:
#
# $$ \boxed{X = X_o \pm \Delta X}$$
#
# donde:
# $$$$
# * <b>X</b> es el resultado de la medición.
# $$$$
# * <b>$X_o$</b> es el valor representativo de la medición (el valor que coincidieramos como verdadero).
# $$$$
# * <b>$\Delta X$</b> es el error absoluto, incerteza o indeterminación de la medición.
# $$$$
# </div>
#
# Profundizaremos en esto el la siguiente sección.
#
# ## <span style="color:red"><b>Toda medición que se realiza debe expresarse de esta forma, de lo contrario no esta bien representada y su resultado carece de sentido. </b></span>
#
# A continuación les dejo algunas preguntas para que se aseguren de haber comprendido el contenido.
#
# **Antes de comenzar no olvide presionar el boton <span style="color:red">Show Widgets</span> para poder responder las preguntas**
# +
# nbi:hide_in
nbi.multiple_choice(question="Magnitud es lo mismoque cantidad",
choices=['Verdadero', 'Falso'],
answers=1)
nbi.short_answer('Como se llama la ciencia que estudio la medición', answers='metrología')
nbi.multiple_choice(question="La medición da como resultado siempre el valor verdadero",
choices=['Verdadero', 'Falso', 'a veces'],
answers=1)
# -
# <form method="get" action="https://nicolasferragamo.github.io/AEA/index.html">
# <button type="submit">Inicio
# </form>
#
# <form method="get" action="https://nicolasferragamo.github.io/AEA/Mediciones/errores.html">
# <button type="submit">Siguiente
# </form>
# **Copyright**
#
# The notebook is provided as [Open Educational Resource]('https://en.wikipedia.org/wiki/Open_educational_resources'). Feel free to use the notebook for your own educational purposes. The text is licensed under [Creative Commons Attribution 4.0]('https://creativecommons.org/licenses/by/4.0/'), the code of the IPython examples under the [MIT license]('https://opensource.org/licenses/MIT'). Please attribute the work as follows: Aplicaciones de electrónica Analógica by <NAME>.
#
| Mediciones/introduccion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
ax = plt.gca()
ax.plot([1,2],[21,10])
ax.plot([2,0],[2,5])
ax.plot([2,3],[10,1])
ax.scatter([1,2,3],[2,4,5])
plt.ylimit(0,100)
| week_2/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### preqs
# `start bert-en`
import pandas as pd
dfjson = pd.read_json(f'/pi/stack/crawlers/langcrs/all_zh.json')
dfjson.head()
# +
# quotes['text'].to_list()
# -
dfjson_ja = pd.read_json(f'/pi/stack/crawlers/langcrs/all_ja.json')
dfjson_id = pd.read_json(f'/pi/stack/crawlers/langcrs/all_id.json')
set_ja=set(dfjson_ja['text'].to_list())
set_id=set(dfjson_id['text'].to_list())
print(len(set_ja), len(set_id), set_ja-set_id)
new_df = pd.merge(dfjson_ja, dfjson_id, how='left', left_on=['text','chapter'], right_on = ['text','chapter'])
new_df.head()
# +
import json
def search_in(text, lang):
with open(f'/pi/stack/crawlers/langcrs/all_{lang}.json') as json_file:
sents=json.load(json_file)
return [sent for sent in sents if sent['text']==text]
def search_in_list(text, langs):
rs={}
for lang in langs:
rs[lang]=search_in(text, lang)
return rs
search_in('both of us', 'ja')
# -
search_in_list('I write a letter.', ['ja', 'fa', 'id'])
# +
import pandas as pd
from bert_serving.client import BertClient
# Bert Client must be running locally
bc = BertClient()
# -
def train(quotes):
embeddings = bc.encode(quotes.text.to_list())
quotes['EMBEDDINGS'] = embeddings.tolist()
# Persist to pickle
quotes.to_pickle('data/embedded_corpus.pkl')
train(dfjson)
# +
def load_quotes_and_embeddings(file):
quotes = pd.read_pickle(file)
# change dtype in place for memory efficiency
quotes['EMBEDDINGS'] = quotes['EMBEDDINGS'].apply(
lambda arr: np.array(arr, dtype='float32')
)
quote_embeddings = np.stack(quotes.EMBEDDINGS.values)
# reduce memory footprint by dropping column
quotes.drop('EMBEDDINGS', axis='columns')
# normalize embeddings for cosine distance
embedding_sums = quote_embeddings.sum(axis=1)
normed_embeddings = quote_embeddings / embedding_sums[:, np.newaxis]
return quotes, normed_embeddings
quotes, embeddings = load_quotes_and_embeddings('data/embedded_corpus.pkl')
# +
def create_index(embeddings):
import faiss
"""
Create an index over the quote embeddings for fast similarity search.
"""
dim = embeddings.shape[1]
index = faiss.IndexFlatL2(dim)
index.add(embeddings)
return index
index = create_index(embeddings)
# -
def search(text):
text_embedding = bc.encode([text])
normalized_text_embedding = text_embedding / text_embedding.sum()
_, idx = index.search(normalized_text_embedding, 5)
relevant_quotes = quotes.iloc[idx.flatten()]['text'].values
relevant_chapters = quotes.iloc[idx.flatten()]['chapter'].values
return relevant_quotes, relevant_chapters
# +
# bc = BertClient()
text="I dreamed a dream."
relevant_quotes, relevant_chapters=search(text)
for q in range(5):
print('>'+relevant_quotes[q])
print(relevant_chapters[q])
# +
text="I write a letter."
relevant_quotes, relevant_chapters=search(text)
for q in range(5):
print('>'+relevant_quotes[q])
print(relevant_chapters[q])
# -
x,y=[1,2]
print(y)
| spacy-2.2/procs-faiss-corpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unit 5 - Financial Planning
# +
# Initial imports
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
# %matplotlib inline
# -
# Load .env enviroment variables
load_dotenv()
# ## Part 1 - Personal Finance Planner
# ### Collect Crypto Prices Using the `requests` Library
# Set current amount of crypto assets
# YOUR CODE HERE!
my_btc = 1.2
my_eth = 5.3
# Crypto API URLs
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=CAD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=CAD"
# +
# Fetch current BTC price
# YOUR CODE HERE!
btc_res = requests.get(btc_url).json()
my_btc_value = btc_res["data"]["1"]["quotes"]["USD"]["price"]
# Fetch current ETH price
# YOUR CODE HERE!
eth_res = requests.get(eth_url).json()
my_eth_value = eth_res["data"]["1027"]["quotes"]["USD"]["price"]
# Compute current value of my crpto
# YOUR CODE HERE!
print()
# Print current crypto wallet balance
print(f"The current value of your {my_btc} BTC is ${my_btc_value:0.2f}")
print(f"The current value of your {my_eth} ETH is ${my_eth_value:0.2f}")
# -
# ### Collect Investments Data Using Alpaca: `SPY` (stocks) and `AGG` (bonds)
# Set current amount of shares
my_agg = 200
my_spy = 50
# +
# Set Alpaca API key and secret
# YOUR CODE HERE!
# for some reason os.getenv was not returning string type for keys, had to hardcode for it to work
# alpaca_api_key = os.getenv("ALPACA_API_KEY")
# alpaca_secret_key = os.getenv("ALPACA_SECRET_KEY")
alpaca_api_key = "PK9JHYIUPWNUF0WX73IF"
alpaca_secret_key = "EuicuLgc9efvy081jfUI47msOkjuCItYyKaKyL0f"
# verification of type
# print(f"Alpaca Key type: {type(alpaca_api_key)}")
# print(f"Alpaca Secret Key type: {type(alpaca_secret_key)}")
# Create the Alpaca API object
# YOUR CODE HERE!
alpaca = tradeapi.REST(
alpaca_api_key,
alpaca_secret_key,
api_version="v2")
# +
# Format current date as ISO format
# YOUR CODE HERE!
today = pd.Timestamp("2021-06-15", tz="America/New_York").isoformat()
# Set the tickers
tickers = ["AGG", "SPY"]
# Set timeframe to '1D' for Alpaca API
timeframe = "1D"
# Get current closing prices for SPY and AGG
# (use a limit=1000 parameter to call the most recent 1000 days of data)
# YOUR CODE HERE!
df_portfolio = alpaca.get_barset(
tickers,
timeframe,
start = today,
end = today,
limit=1000
).df
# Preview DataFrame
# YOUR CODE HERE!
df_portfolio
# +
# Pick AGG and SPY close prices
# YOUR CODE HERE!
df_cp = pd.DataFrame()
df_cp['AGG'] = df_portfolio['AGG']["close"]
df_cp['SPY'] = df_portfolio['SPY']["close"]
# Drop the time component of the date
df_cp.index = df_cp.index.date
agg_close_price = df_cp['AGG']
spy_close_price = df_cp['SPY']
# Print AGG and SPY close prices
print(f"Current AGG closing price: ${agg_close_price}")
print(f"Current SPY closing price: ${spy_close_price}")
# +
# Compute the current value of shares
# YOUR CODE HERE!
# Print current value of shares
print(f"The current value of your {my_spy} SPY shares is ${my_spy_value:0.2f}")
print(f"The current value of your {my_agg} AGG shares is ${my_agg_value:0.2f}")
# -
# ### Savings Health Analysis
# +
# Set monthly household income
# YOUR CODE HERE!
# Consolidate financial assets data
# YOUR CODE HERE!
# Create savings DataFrame
# YOUR CODE HERE!
# Display savings DataFrame
display(df_savings)
# +
# Plot savings pie chart
# YOUR CODE HERE!
# +
# Set ideal emergency fund
emergency_fund = monthly_income * 3
# Calculate total amount of savings
# YOUR CODE HERE!
# Validate saving health
# YOUR CODE HERE!
# -
# ## Part 2 - Retirement Planning
#
# ### Monte Carlo Simulation
# Set start and end dates of five years back from today.
# Sample results may vary from the solution based on the time frame chosen
start_date = pd.Timestamp('2016-05-01', tz='America/New_York').isoformat()
end_date = pd.Timestamp('2021-05-01', tz='America/New_York').isoformat()
# +
# Get 5 years' worth of historical data for SPY and AGG
# (use a limit=1000 parameter to call the most recent 1000 days of data)
# YOUR CODE HERE!
# Display sample data
df_stock_data.head()
# +
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
# YOUR CODE HERE!
# +
# Printing the simulation input data
# YOUR CODE HERE!
# +
# Running a Monte Carlo simulation to forecast 30 years cumulative returns
# YOUR CODE HERE!
# +
# Plot simulation outcomes
# YOUR CODE HERE!
# +
# Plot probability distribution and confidence intervals
# YOUR CODE HERE!
# -
# ### Retirement Analysis
# +
# Fetch summary statistics from the Monte Carlo simulation results
# YOUR CODE HERE!
# Print summary statistics
# YOUR CODE HERE!
# -
# ### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `$20,000` initial investment.
# +
# Set initial investment
initial_investment = 20000
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000
# YOUR CODE HERE!
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
# -
# ### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `50%` increase in the initial investment.
# +
# Set initial investment
initial_investment = 20000 * 1.5
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000
# YOUR CODE HERE!
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
# -
# ## Optional Challenge - Early Retirement
#
#
# ### Five Years Retirement Option
# +
# Configuring a Monte Carlo simulation to forecast 5 years cumulative returns
# YOUR CODE HERE!
# +
# Running a Monte Carlo simulation to forecast 5 years cumulative returns
# YOUR CODE HERE!
# +
# Plot simulation outcomes
# YOUR CODE HERE!
# +
# Plot probability distribution and confidence intervals
# YOUR CODE HERE!
# +
# Fetch summary statistics from the Monte Carlo simulation results
# YOUR CODE HERE!
# Print summary statistics
# YOUR CODE HERE!
# +
# Set initial investment
# YOUR CODE HERE!
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000
# YOUR CODE HERE!
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 5 years will end within in the range of"
f" ${ci_lower_five} and ${ci_upper_five}")
# -
# ### Ten Years Retirement Option
# +
# Configuring a Monte Carlo simulation to forecast 10 years cumulative returns
# YOUR CODE HERE!
# +
# Running a Monte Carlo simulation to forecast 10 years cumulative returns
# YOUR CODE HERE!
# +
# Plot simulation outcomes
# YOUR CODE HERE!
# +
# Plot probability distribution and confidence intervals
# YOUR CODE HERE!
# +
# Fetch summary statistics from the Monte Carlo simulation results
# YOUR CODE HERE!
# Print summary statistics
# YOUR CODE HERE!
# +
# Set initial investment
# YOUR CODE HERE!
# Use the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000
# YOUR CODE HERE!
# Print results
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 10 years will end within in the range of"
f" ${ci_lower_ten} and ${ci_upper_ten}")
# -
| Starter_Code/financial-planner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ravi02512/Sentiment-classification-using-LSTM-Network/blob/master/sentiment%20Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="vgZwOBH0rcV2" colab_type="code" outputId="bb4d3ed2-09f0-4542-de7c-d137093c9c95" colab={"base_uri": "https://localhost:8080/", "height": 124}
from google.colab import drive
drive.mount('/content/drive')
# + id="l_E4BNMzrBx3" colab_type="code" outputId="fd9c7dfd-77c2-4b65-b362-3879f47d0d30" colab={"base_uri": "https://localhost:8080/", "height": 34}
import numpy as np
np.random.seed(0)
from keras.models import Model
from keras.layers import Dense, Input, Dropout, LSTM, Activation
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
from keras.initializers import glorot_uniform
np.random.seed(1)
import pandas as pd
import numpy as np
# + id="UC2lhKn0tsZL" colab_type="code" colab={}
#helper functions
import numpy as np
import pandas as pd
def read_glove_vecs(glove_file):
with open(glove_file, 'r') as f:
words = set()
word_to_vec_map = {}
for line in f:
line = line.strip().split()
curr_word = line[0]
words.add(curr_word)
word_to_vec_map[curr_word] = np.array(line[1:], dtype=np.float64)
i = 1
words_to_index = {}
index_to_words = {}
for w in sorted(words):
words_to_index[w] = i
index_to_words[i] = w
i = i + 1
return words_to_index, index_to_words, word_to_vec_map
def convert_to_one_hot(Y, C):
Y = np.eye(C)[Y.reshape(-1)]
return Y
# + id="_XVSlEQzrLVm" colab_type="code" colab={}
train = pd.read_csv('/content/drive/My Drive/sentiment/train_2kmZucJ.csv')
test= pd.read_csv('/content/drive/My Drive/sentiment/test_oJQbWVk.csv')
# + id="TT12wV5hF-Xr" colab_type="code" colab={}
word_to_index, index_to_word, word_to_vec_map = read_glove_vecs('/content/drive/My Drive/sentiment/glove.6B/glove.6B/glove.6B.300d.txt')
dictionary=word_to_vec_map.keys()
# + id="aFsbO2ZMEYzd" colab_type="code" colab={}
import re
clean_train=[]
for i in range(len(train)):
new=re.sub('[^A-Za-z ]+', ' ', train['tweet'][i])
clean_train.append(new)
# + id="SQKmuAJULr4m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2e585cde-1caa-4f60-a367-1256930115e7"
import nltk
nltk.download('stopwords')
stopwords = nltk.corpus.stopwords.words('english')
# + id="jr9Clj9VL1cT" colab_type="code" colab={}
def clean_sentence(sentence):
tokens=[]
word_list=sentence.split(' ')
for w in word_list:
if w in dictionary:
# if w not in stopwords:
if set(w) & set('aeiou')!=set():
tokens.append(w)
return ' '.join(tokens)
# + id="upxy6XdoLh6d" colab_type="code" colab={}
final_train=[]
for i in range(len(clean_train)):
final_train.append(clean_sentence(clean_train[i]))
# + id="hsJewaTQOzdE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a2ebd80e-617e-47f8-ac78-5f4ef97943ba"
final_train[1]
# + id="dvjjQoK8posN" colab_type="code" colab={}
tokens=[]
for i in range(len(clean_train)):
tokens.extend(clean_train[i].split(' '))
# + id="0ncGFWt_qKxz" colab_type="code" outputId="61e62682-d8be-45fa-9d6f-e8cc9e8ee658" colab={"base_uri": "https://localhost:8080/", "height": 206}
count_df=pd.DataFrame(tokens,columns=['words'])
count_df=count_df['words'].value_counts().reset_index().rename(columns={'index':'words', 'words':'counts'})
count_df['word length']=[len(w) for w in count_df['words']]
count_df.head()
# + id="8eaNa1VIigXL" colab_type="code" outputId="1aeb915b-e2bf-4c94-ece2-adedc5502181" colab={"base_uri": "https://localhost:8080/", "height": 132}
g=[]
for w in count_df['words']:
if w in dictionary:
g.append(w)
print('number of words belong to dictionary'len(g))
# + id="AyQ-sqrYsES_" colab_type="code" colab={}
length=[]
for i in range(len(final_train)):
length.append(len([word.lower() for word in final_train[i].split()]))
# + id="rCY5IBmDsEQ-" colab_type="code" outputId="93673a83-9cb0-41b1-cb58-82e654968d97" colab={"base_uri": "https://localhost:8080/", "height": 34}
max_len=np.max(length)
print(max_len)
# + id="EXQqwmumsEN1" colab_type="code" colab={}
def sentences_to_indices(X, word_to_index, max_len):
"""
Arguments:
X -- array of sentences (strings), of shape (m, 1)
word_to_index -- a dictionary containing the each word mapped to its index
max_len -- maximum number of words in a sentence. You can assume every sentence in X is no longer than this.
Returns:
X_indices -- array of indices corresponding to words in the sentences from X, of shape (m, max_len)
"""
m = X.shape[0]
X_indices = np.zeros((m,max_len))
for i in range(m):
sentence_words = [word.lower() for word in X[i].split()]
j = 0
for w in sentence_words:
if w.lower() in dictionary:
X_indices[i, j] = word_to_index[w.lower()]
#
j = j + 1
return X_indices
# + id="t3HVjNLKDY34" colab_type="code" outputId="c383ca83-50dd-4369-f8aa-61be7b49e93f" colab={"base_uri": "https://localhost:8080/", "height": 34}
# + id="Hs8UpgX-sELn" colab_type="code" colab={}
def pretrained_embedding_layer(word_to_vec_map, word_to_index):
"""
Arguments:
word_to_vec_map -- dictionary mapping words to their GloVe vector representation.
word_to_index -- dictionary mapping from words to their indices in the vocabulary (400,001 words)
Returns:
embedding_layer -- pretrained layer Keras instance
"""
vocab_len = len(word_to_index) + 1
emb_dim = word_to_vec_map["cucumber"].shape[0]
emb_matrix = np.zeros((vocab_len, emb_dim))
for word, index in word_to_index.items():
if index!= 1800:
emb_matrix[index, :] = word_to_vec_map[word]
embedding_layer = Embedding(vocab_len, emb_dim, trainable = False)
embedding_layer.build((None,))
embedding_layer.set_weights([emb_matrix])
return embedding_layer
# + id="hDcP1lvKsEHw" colab_type="code" colab={}
def my_model(input_shape, word_to_vec_map, word_to_index):
"""
Arguments:
input_shape -- shape of the input, usually (max_len,)
word_to_vec_map -- dictionary mapping every word in a vocabulary into its 50-dimensional vector representation
word_to_index -- dictionary mapping from words to their indices in the vocabulary (400,001 words)
Returns:
model -- a model instance in Keras
"""
sentence_indices = Input(input_shape, dtype = 'int32')
embedding_layer = pretrained_embedding_layer(word_to_vec_map, word_to_index)
embeddings = embedding_layer(sentence_indices)
X = LSTM(128, return_sequences = True)(embeddings)
X = Dropout(0.5)(X)
X = LSTM(128, return_sequences=False)(X)
X = Dropout(0.5)(X)
X = Dense(2)(X)
X = Activation('softmax')(X)
model = Model(inputs = sentence_indices, outputs = X)
return model
# + id="dpHEXa0QCvNX" colab_type="code" outputId="4caa77c4-9333-4027-a148-e8de8c65309c" colab={"base_uri": "https://localhost:8080/", "height": 34}
import gc
gc.collect()
# + id="oDfwEKRxsEFx" colab_type="code" outputId="79866b41-d9ab-4f1b-c1cf-e48e2b90a7cf" colab={"base_uri": "https://localhost:8080/", "height": 434}
model = my_model((max_len,), word_to_vec_map, word_to_index)
model.summary()
# + id="jJ3CuK71sEBs" colab_type="code" colab={}
from keras import optimizers
# sgd = optimizers.SGD(lr=0.1, decay=1e-4, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="9i-HaP1jwcbj" colab_type="code" colab={}
xtrain=np.asarray(final_train)
y_train=np.asarray(ytrain)
# + id="rWNA39B-s0lP" colab_type="code" colab={}
X_train_indices = sentences_to_indices(xtrain, word_to_index, max_len)
Y_train_oh = convert_to_one_hot(y_train, C = 2)
# + id="DJoWeCSTs3qL" colab_type="code" outputId="5853f115-941e-4377-a6b2-81f67c9d7cc7" colab={"base_uri": "https://localhost:8080/", "height": 399}
model.fit(X_train_indices, Y_train_oh,validation_split=0.2, epochs = 10, batch_size = 32, shuffle=True)
# + id="Jhiwk32J9LFF" colab_type="code" colab={}
test_clean=[]
for i in range(len(test)):
new=re.sub('[^A-Za-z ]+', ' ', test['tweet'][i])
test_clean.append(new)
final_test=[]
for i in range(len(test_clean)):
final_test.append(clean_sentence(test_clean[i]))
xtest=np.asarray(final_test)
X_test_indices = sentences_to_indices(xtest, word_to_index, max_len)
y_pred=model.predict(X_test_indices)
pred=[]
for i in range(len(y_pred)):
pred.append(y_pred[i].tolist().index(y_pred[i].max()))
# + id="eS7uPWT0s3fy" colab_type="code" colab={}
import os
sample_sub=pd.read_csv('/content/drive/My Drive/sentiment/sample_submission_LnhVWA4.csv')
sample_sub['label']=pred
os.chdir('/content/drive/My Drive/sentiment')
sample_sub.to_csv('sub17.csv', index=False)
# + id="9Aja_cZ7s3Zg" colab_type="code" colab={}
| sentiment Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py27]
# language: python
# name: conda-env-py27-py
# ---
# # The Skeleton Notebook
# This notebook/template serves as a starting point for most Supervised Machine Learning projects that involve common tasks such as data exploration, cleaning, transformation and preparation, and data modeling (using machine learning or deep learning techniques).
#
# I've tried to build the notebook to provide a set workflow in which to handle the above tasks. These are arranged in sections, encouraged to be expanded to into sub-sections to handle approproate tasks. I've also tried to include common sub-tasks, and the code required to do them (usually Pandas or scikit-learn).
#
# Sections included:
#
# - Housekeeping and Imports
# - Data Loading
# - Data Exploration
# - Data Cleaning
# - Feature Engineering
# - Data Transformation and Preparation
# - Model Exploration and Performance Analysis
# - Final Model Building
#
# It is suggested to use separate notebooks if any of the above tasks are performed in depth.
# ## Housekeeping and Imports
# For importing libraries necessary for the project, and for basic preprocessing functions (ex: typset conversion for NLP projects).
#
# We're going to import commonly used Data Science libraries, so make sure they're available for your Python set-up.
# +
# Import libraries necessary for projects
import numpy as np
import pandas as pd
from time import time
from IPython.display import display # Allows the use of display() for DataFrames
# Import visualisation libraries
import seaborn as sns
import matplotlib.pyplot as plt
# Pretty display for notebooks
# %matplotlib inline
# Make division futuristic for Python 2
from __future__ import division
# +
#Cell for Housekeeping code
# -
# ## Data Loading
# For loading data files into appropriate variables.
# +
#Loading the data file (ex: csv) using Pandas
# data = pd.read_csv('') #insert path to file
#Next steps?:
# Loading the test data?
# Loading the feaure vectors (X) and the prediction vector (Y) into different variables
# -
# ## Data Exploration
# Section for **exploratory analysis** on the available data.
#
# The exploration techniques vary for numerical, categorical, or time-series variables. Currently,
#
# Here we typically:
#
# - look at example records in the dataset
# - investigate the datatypes of variables in the dataset
# - calculate and investigate descriptive statistics (ex: central tendencies, variability etc.)
# - investigate distribution of feature vectors (ex: to check for skewness and outliers)
# - investigate distribution of prediction vector
# - check out the relationship (ex: correlation) between different features
# - check out the relationship between feature vectors and prediction vector
#
# Common steps to check the health of the data:
#
# - Check for missing data
# - Check the skewness of the data, outlier detection
# - etc...
# ### Look at Example Records
# +
# data.head(5) #Display out the first 5 records
# Additional:
# Look at last few records using data.tail()
# -
# ### Data-types, completeness Information
#
# Using the Pandas "info" function, in addition to the data-type information for the dataset, we can look at counts of available records/missing records too.
# +
# data.info()
# -
# ### Descriptive Statistics
# +
# data.describe()
# Additonal:
# We can also make a guess at the skewness of the data at this stage by looking at the difference between
# the means and medians of numerical features
# -
# ### Visualizaton: Distribution of features
# *Section has great potential for expansion.*
#
# Visualization techniques differ depending on the type of the feature vector (i.e. numerical: continuous or discrete, categorical: ordinal etc). Techniques will also depend on the type of data being dealt with, and the insight that we want to extract from it.
#
# Common visualization techniques include:
# - Bar Plots: Visualize the frequency distribution of categorical features.
# - Histograms: Visualize the frequency distribution of numerical features.
# - Box Plots: Visualize a numerical feature, while providing more information like the median, lower/upper quantiles etc..
# - Scatter Plots: Visualize the relationship (usually the correlation) between two features. Can include a goodness of fit line, to serve as a regression plot.
#
# Below are example code snippets to draw these using seaborn.
# +
#Example: drawing a seaborn barplot
#sns.barplot(x="",y="",hue="",data="")
#Can also use pandas/matplotlib for histograms (numerical features) or barplots ()
# +
# Example: drawing a seaborn regplot
# sns.regplot(data[feature1],data[feature2])
# +
#Example: drawing a pandas scatter_matrix
# pd.scatter_matrix(data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
# -
# ### Investigating correlations between features
# ### Visualizing prediction vector
# ### Investigating missing values
# ### Outlier Detection
# The presence of outliers can often skew results which take into consideration these data points.
#
# One approach to detect outliers is to use Tukey's Method for identfying them: An outlier step is calculated as 1.5 times the interquartile range (IQR). A data point with a feature that is beyond an outlier step outside of the IQR for that feature is considered abnormal.
#
# One such pipeline for detecting outliers is below:
# +
# def find_outliers(data):
# #Checking for outliers that occur for more than one feature
# outliers = []
# # For each feature find the data points with extreme high or low values
# for feature in [list of features to investigate]:
# # TODO: Calculate Q1 (25th percentile of the data) for the given feature
# Q1 = np.percentile(data[feature],25)
# # TODO: Calculate Q3 (75th percentile of the data) for the given feature
# Q3 = np.percentile(data[feature],75)
# # TODO: Use the interquartile range to calculate an outlier step (1.5 times the interquartile range)
# step = (Q3-Q1) * 1.5
# # Display the outliers
# out = data[~((data[feature] >= Q1 - step) & (data[feature] <= Q3 + step))]
# print "Number of outliers for the feature '{}': {}".format(feature, len(out))
# outliers = outliers + list(out.index.values)
# #Creating list of more outliers which are the same for multiple features.
# outliers = list(set([x for x in outliers if outliers.count(x) > 1]))
# return outliers
# print "Data points considered outliers for more than one feature: {}".format(find_outliers(data))
# -
# ## Data Cleaning
# ### Imputing missing values
# ### Cleaning outliers or error values
# +
# Remove the outliers, if any were specified
# good_data = data.drop(data.index[outliers]).reset_index(drop = True)
# print "The good dataset now has {} observations after removing outliers.".format(len(good_data))
# -
# ## Feature Engineering
# Section to extract more features from those currently available.
# +
# code
# -
# ## Data Transformation and Preparation
# ### Transforming Skewed Continous Features
# It is common practice to apply a logarthmic transformation to highly skewed continuous feature distributions. A typical flow for this is in a commented code block below.
# +
# skewered = [list of skewed continuous features]
# raw_features[skewed] = data[skewed].apply(lambda x: np.log(x+1))
# -
# ### Normalizing Numerical Features
# Another common practice is to perform some type of scaling on numerical features. Applying scaling doesn't change the shape of each feature's distribution; but ensures that each feature is treated equally when applying supervised learners. An example workflow of achieving normalisation using the MinMaxScaler module of sklearn is below:
# +
# from sklearn.preprocessing import MinMaxScaler
# scaler = MinMaxScaler()
# numerical = [list of skewed numerical features]
# raw_features[numerical] = scaler.fit_transform(data[numerical])
# +
# Checking examples after transformation
# raw_features.head()
# -
# ### One Hot Encoding Categorical Features
# +
# Using Pandas get_dummies function
# features = pd.get_dummies(raw_features)
# +
#Encoding categorical prediction vector to numerical ?
# -
# It is encouraged to create a pipeline function for data preprocessing, rather than separate script blocks.
# ### Shuffle and Split Data
# +
# from sklearn.cross_validation import train_test_split
# X_train, X_test, y_train, y_test = train_test_split(features, prediction_vector, test_size = 0.2, random_state = 0)
# Show the results of the split
# print "Training set has {} samples.".format(X_train.shape[0])
# print "Testing set has {} samples.".format(X_test.shape[0])
# -
# ## Model Exploration
# ### Naive Predictor Performance
# To set a baseline for the performance of the predictor.
#
# Common techniques:
# - For categorical prediction vector, choose the most common class
# - For numerical prediction vector, choose a measure of central tendency
#
# Then calculate the evalation metric (accuracy, f-score etc)
# +
#Code to implement the above
# -
# ### Choosing scoring metrics
# +
# from sklearn.metrics import accuracy_score, fbeta_score
# -
# ### Creating a Training and Prediction Pipeling
# +
#Importing models from sklearn, or tensorflow/keras components
# -
# Change below as seen fit
# +
# def train_predict(learner, sample_size, X_train, y_train, X_test, y_test):
# '''
# inputs:
# - learner: the learning algorithm to be trained and predicted on
# - sample_size: the size of samples (number) to be drawn from training set
# - X_train: features training set
# - y_train: income training set
# - X_test: features testing set
# - y_test: income testing set
# '''
# results = {}
# # TODO: Fit the learner to the training data using slicing with 'sample_size'
# start = time() # Get start time
# learner = learner.fit(X_train[:sample_size],y_train[:sample_size])
# end = time() # Get end time
# # TODO: Calculate the training time
# results['train_time'] = end - start
# # TODO: Get the predictions on the test set,
# # then get predictions on the first 300 training samples
# start = time() # Get start time
# predictions_test = learner.predict(X_test)
# predictions_train = learner.predict(X_train[:300])
# end = time() # Get end time
# # TODO: Calculate the total prediction time
# results['pred_time'] = end - start
# # TODO: Compute accuracy on the first 300 training samples
# results['acc_train'] = accuracy_score(y_train[:300],predictions_train)
# # TODO: Compute accuracy on test set
# results['acc_test'] = accuracy_score(y_test,predictions_test)
# # TODO: Compute F-score on the the first 300 training samples
# results['f_train'] = fbeta_score(y_train[:300],predictions_train,0.5)
# # TODO: Compute F-score on the test set
# results['f_test'] = fbeta_score(y_test,predictions_test,0.5)
# # Success
# print "{} trained on {} samples.".format(learner.__class__.__name__, sample_size)
# # Return the results
# return results
# -
# ### Model Evaluation
# +
# Change the list of classifiers and code below as seen fit. we probably also don't need to see the effects of
# different sample sizes
# +
# # TODO: Import the three supervised learning models from sklearn
# from sklearn.tree import DecisionTreeClassifier
# from sklearn.svm import SVC
# from sklearn.ensemble import AdaBoostClassifier
# # TODO: Initialize the three models, the random states are set to 101 so we know how to reproduce the model later
# clf_A = DecisionTreeClassifier(random_state=101)
# clf_B = SVC(random_state = 101)
# clf_C = AdaBoostClassifier(random_state = 101)
# # TODO: Calculate the number of samples for 1%, 10%, and 100% of the training data
# samples_1 = int(round(len(X_train) / 100))
# samples_10 = int(round(len(X_train) / 10))
# samples_100 = len(X_train)
# # Collect results on the learners in a dictionary
# results = {}
# for clf in [clf_A, clf_B, clf_C]:
# clf_name = clf.__class__.__name__
# results[clf_name] = {}
# for i, samples in enumerate([samples_1, samples_10, samples_100]):
# results[clf_name][i] = \
# train_predict(clf, samples, X_train, y_train, X_test, y_test)
# -
# Printing out the results
# +
# #Printing out the values
# for i in results.items():
# print i[0]
# display(pd.DataFrame(i[1]).rename(columns={0:'1%', 1:'10%', 2:'100%'}))
# -
# ## Final Model Building
# Using grid search (GridSearchCV) with different parameter/value combinations, we can tune our model for even better results.
#
# Example with Adaboost below
# +
# # TODO: Import 'GridSearchCV', 'make_scorer', and any other necessary libraries
# from sklearn.grid_search import GridSearchCV
# from sklearn.metrics import make_scorer
# # TODO: Initialize the classifier
# clf = AdaBoostClassifier(base_estimator=DecisionTreeClassifier())
# # TODO: Create the parameters list you wish to tune
# parameters = {'n_estimators':[50, 120],
# 'learning_rate':[0.1, 0.5, 1.],
# 'base_estimator__min_samples_split' : np.arange(2, 8, 2),
# 'base_estimator__max_depth' : np.arange(1, 4, 1)
# }
# # TODO: Make an fbeta_score scoring object
# scorer = make_scorer(fbeta_score,beta=0.5)
# # TODO: Perform grid search on the classifier using 'scorer' as the scoring method
# grid_obj = GridSearchCV(clf, parameters,scorer)
# # TODO: Fit the grid search object to the training data and find the optimal parameters
# grid_fit = grid_obj.fit(X_train,y_train)
# # Get the estimator
# best_clf = grid_fit.best_estimator_
# # Make predictions using the unoptimized and model
# predictions = (clf.fit(X_train, y_train)).predict(X_test)
# best_predictions = best_clf.predict(X_test)
# # Report the before-and-afterscores
# print "Unoptimized model\n------"
# print "Accuracy score on testing data: {:.4f}".format(accuracy_score(y_test, predictions))
# print "F-score on testing data: {:.4f}".format(fbeta_score(y_test, predictions, beta = 0.5))
# print "\nOptimized Model\n------"
# print "Final accuracy score on the testing data: {:.4f}".format(accuracy_score(y_test, best_predictions))
# print "Final F-score on the testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5))
# print best_clf
# -
# Next steps can include feature importance extraction, predictions on the test set.. etc
# ## Predictions on Test Set
| Skeleton Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Import libraries
import numpy as np
import math
import random
import matplotlib.pyplot as plt
from matplotlib import ticker
import itertools
from matplotlib.ticker import FixedLocator, FixedFormatter
from scipy import stats
from sklearn.preprocessing import normalize
from sklearn.preprocessing import minmax_scale
#Ensure reproducibility
random.seed(1)
np.random.seed(1)
class model(object):
'''
This class implements the CP model of human vision
as described in "Modeling learned categorical perception in human vision"
<NAME> (2012)
'''
def __init__(self, M_PC=7, M_EC=7, M_VV=7, W_c=0.4, mu_PC=0.6, mu_EC=0.4, mu_VV=0.2, rho=1, eta=0.01):
#Activation vectors
self.f_u_PC = np.zeros(M_PC) #Pre-cortical activity
self.f_u_EC = np.zeros(M_EC) #Early cortical activity
self.f_u_VV = np.zeros(M_VV) #Ventral visual activity
#Weights
self.W_PC = np.random.normal(0, 0.5, 9*M_PC).reshape((9, M_PC)) #Pre-cortical weights
self.W_EC = np.random.normal(0, 0.5, M_PC*M_EC).reshape((M_PC, M_EC)) #Early visual cortex weights
self.W_VV = np.random.normal(0, 0.5, M_EC*M_VV).reshape((M_EC, M_VV)) #Ventral visual weights
self.W_c = W_c #Category input weight
#Parameters
self.mu_PC = mu_PC #Pre-cortical Inhibtion rate
self.mu_EC = mu_EC #Early cortical Inhibtion rate
self.mu_VV = mu_VV #Ventral visual Inhibtion rate
self.eta = eta #Learning rate
self.rho = rho #Weight change threshold
def activation(self, x, module, cat_input=0):
'''
This function computes the activation of the module specified as an argument and given an input.
Cat_input is only 1 when a stimulus belonging to category B is presented AND the model is in category training
Returns the activation vector.
Module 0: precortical, module 1:early cortical, module 2:ventral visual
'''
if module == 0:
weights = self.W_PC
mu = self.mu_PC
elif module == 1:
weights = self.W_EC
mu = self.mu_EC
else:
weights = self.W_VV
mu = self.mu_VV
# Equations 4 and 6
f_u = np.dot(x, weights)
y_win_i = np.argmax(f_u)
if not np.isscalar(y_win_i):
y_win_i = np.random.choice(y_win_i)
for k in range(len(f_u)):
if f_u[k] > 1:
f_u[k] = 1
elif f_u[k] < 0:
f_u[k] = 0
y_win = np.amax(f_u)
#Provide category input to module 2
if module == 2:
f_u += self.W_c*cat_input
# Equation 5
for i in f_u:
if i < f_u[y_win_i]:
i -= mu*y_win
if module == 0:
self.f_u_PC = f_u
elif module == 1:
self.f_u_EC = f_u
else:
self.f_u_VV = f_u
return f_u
def update(self, x, module, cat_input=0, category_training=False):
'''
This function updates the weights of the module specified as an argument given an input x
Trigger category training for the 3rd module only.
Layer 0: PC; Layer 1: EC; Layer 2: VV
'''
# Equation 8
if category_training:
ar = np.append(x, cat_input)
mean_x = np.mean(ar)
else:
mean_x = np.mean(x)
# Equation 7
if module == 0:
w_ = np.zeros((self.W_PC.shape[0], self.W_PC.shape[1]))
for i in range(self.W_PC.shape[0]):
for j in range(self.W_PC.shape[1]):
if x[i] > self.rho*mean_x:
w_[i, j] = self.W_PC[i,j] + self.eta*x[i]*self.f_u_PC[j]
else:
w_[i, j] = self.W_PC[i,j]
elif module == 1:
w_ = np.zeros((self.W_EC.shape[0], self.W_EC.shape[1]))
for i in range(self.W_EC.shape[0]):
for j in range(self.W_EC.shape[1]):
if x[i] > self.rho*mean_x:
w_[i, j] = self.W_EC[i,j] + self.eta*x[i]*self.f_u_EC[j]
else:
w_[i, j] = self.W_EC[i,j]
else:
w_ = np.zeros((self.W_VV.shape[0], self.W_VV.shape[1]))
for i in range(self.W_VV.shape[0]):
for j in range(self.W_VV.shape[1]):
if x[i] > self.rho*mean_x:
w_[i, j] = self.W_VV[i,j] + self.eta*x[i]*self.f_u_VV[j]
else:
w_[i, j] = self.W_VV[i,j]
if category_training:
w_ = np.concatenate((w_, [np.repeat(self.W_c, self.W_VV.shape[1])]))
i = w_.shape[0] -1
x_cat = cat_input*self.W_c
for j in range(self.W_VV.shape[1]):
if x_cat > self.rho*mean_x:
w_[i, j] += self.eta*x_cat*self.f_u_VV[j]
# Equation 9
mean_w = np.mean(w_, axis=0)
if module == 0:
self.W_PC = w_/mean_w
elif module == 1:
self.W_EC = w_/mean_w
else:
if category_training:
self.W_VV = w_[:w_.shape[0]-1]/mean_w
else:
self.W_VV = w_/mean_w
def gabor_stimuli(S_o=45, o=45, lambda_p=106, lambda_o=30, f_comp=0):
#Categories
A = []
B = []
#Phases
P_a = [[0, 45, 90, 135, 180, 225, 270, 225], [0, 45, 90, 135, 180, 225, 270, 315],
[0, 45, 90, 135, 180, 225, 270, 315], [360, 45, 90, 135, 180, 225, 270, 315]]
P_b = [[0, 45, 90, 135, 180, 135, 90, 45], [0, 45, 90, 135, 180, 225, 180, 135],
[360, 405, 90, 135, 180, 225, 270, 315], [360, 405, 450, 135, 180, 225, 270, 315]]
P_A = [90, 135, 180, 225]
P_B = [0, 45, 270, 315]
#Parameters
Lambda_p = (-math.log(1/2))/math.pow(lambda_p/2, 2) # Phase bandwidth parameter (Equation 2)
Lambda_o = (-math.log(1/2))/math.pow(lambda_o/2, 2) # Orientation bandwidth parameter (Equation 3)
for i in range(4):
S_p_A = P_A[i]
S_p_B = P_B[i]
stim_A = []
stim_B = []
for (p_a, p_b) in zip(P_a[i], P_b[i]):
x_A = math.exp(-Lambda_p*math.pow(p_a-S_p_A, 2)-Lambda_o*math.pow(o-S_o, 2)) # Equation 1
x_B = math.exp(-Lambda_p*math.pow(p_b-S_p_B, 2)-Lambda_o*math.pow(o-S_o, 2)) # Equation 1
stim_A.append(x_A)
stim_B.append(x_B)
#f phase input for completeness with constant value of 0
stim_A.append(f_comp)
stim_B.append(f_comp)
A.append(stim_A)
B.append(stim_B)
return A, B
A, B = gabor_stimuli()
#Plot examples of stimuli at 0deg and 225deg
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12,4))
ax1.set_xlabel('3f phase')
ax2.set_xlabel('3f phase')
ax1.set_title('Gaussian pattern of activity with mean at 0 degrees')
ax2.set_title('Gaussian pattern of activity with mean at 225 degrees')
ax1.set_ylabel('Activity')
ax2.set_ylabel('Activity')
ax1.set_xlim((0, 1))
ax2.set_xlim((0, 1))
ax1.set_ylim((0, 1))
ax2.set_ylim((0, 1))
ax1.set_xticks(np.arange(0, 360, 45))
ax2.set_xticks(np.arange(0, 360, 45))
ax1.plot(np.arange(0, 360, 45), B[0][:8],'o-')
ax2.plot(np.arange(0, 360, 45), A[2][:8],'o-')
# +
def test_model(lVF, rVF, trials, stimuli, labels):
left_resp = np.zeros((7, 8, 3, trials))
right_resp = np.zeros((7, 8, 3, trials))
for t in range(trials):
for i, s in enumerate(stimuli):
y_l = lVF.activation(s, 0)
y_r = rVF.activation(s, 0)
left_resp[:,i, 0,t] = y_l
right_resp[:,i, 0,t] = y_r
y_l = lVF.activation(y_l, 1)
y_r = rVF.activation(y_r, 1)
left_resp[:,i, 1,t] = y_l
right_resp[:,i, 1,t] = y_r
y_l = lVF.activation(y_l, 2, cat_input=labels[i])
y_r = rVF.activation(y_r, 2, cat_input=labels[i])
left_resp[:,i, 2, t] = y_l
right_resp[:,i, 2, t] = y_r
return left_resp, right_resp
def pre_train_model(lVF, rVF, epochs, A, B):
#Generate pre-training sequence where 0 is left and 1 is right
training_sequence = np.repeat([0,1], epochs/2)
np.random.shuffle(training_sequence)
pre_training_set = np.concatenate((A,B))
zero_stim = np.zeros(9)
for vf in training_sequence:
np.random.shuffle(pre_training_set)
for stim in pre_training_set:
if vf == 0:
gets_stim = lVF
no_stim = rVF
else:
gets_stim = rVF
no_stim = lVF
#Precortical for both visual fields
y_stim0 = gets_stim.activation(stim, 0)
y_no0 = no_stim.activation(zero_stim, 0)
gets_stim.update(stim, 0)
no_stim.update(zero_stim, 0)
#Early cortical
y_stim1 = gets_stim.activation(y_stim0, 1)
y_no1 = no_stim.activation(y_no0, 1)
gets_stim.update(y_stim0, 1)
no_stim.update(y_no0, 1)
#Ventral visual
gets_stim.activation(y_stim1, 2)
no_stim.activation(y_no1, 2)
gets_stim.update(y_stim1, 2)
no_stim.update(y_no1, 2)
def category_train(lVF, rVF, epochs, A, B):
stim = np.concatenate((B[:2], A, B[2:]))
zero_stim = np.zeros(9)
a = [2,3,4,5]
b = [0,1,6,7]
labels = np.array([1, 1, 0, 0, 0, 0, 1, 1])
within_A = np.array(list(itertools.permutations(a, 2)))
within_B = np.array(list(itertools.permutations(b, 2)))
all_within = np.concatenate((within_A, within_B))
between = np.array(list(itertools.permutations(np.arange(0, 8, 1), 2)))
between = between[np.all(np.any((between-all_within[:, None]), axis=2), axis=0)]
stim_ind = np.concatenate((all_within, between))
#Double training sequence
random_sequence_stim = np.arange(0, 56, 1)
#Single training sequence
random_sequence_vf = np.repeat([0,1], 4)
np.random.shuffle(random_sequence_vf)
random_sequence_stim2 = np.arange(0, 8, 1)
for e in range(epochs):
np.random.shuffle(random_sequence_stim)
np.random.shuffle(random_sequence_stim2)
#Double training
for i in random_sequence_stim:
s = stim_ind[i]
s_left = stim[s[0]]
s_right = stim[s[1]]
l_left = labels[s[0]]
l_right = labels[s[1]]
#Precortical for both visual fields
y_l = lVF.activation(s_left, 0)
y_r = rVF.activation(s_right, 0)
lVF.update(s_left, 0)
rVF.update(s_right, 0)
#Early cortical
y_l = lVF.activation(y_l, 1)
y_r = rVF.activation(y_r, 1)
lVF.update(y_l, 1)
rVF.update(y_r, 1)
#Ventral visual
lVF.activation(y_l, 2, cat_input=l_left)
rVF.activation(y_r, 2, cat_input=l_right)
lVF.update(y_l, 2, cat_input=l_left, category_training=True)
rVF.update(y_r, 2, cat_input=l_right, category_training=True)
#Single training
for i in range(3):
for j in range(8):
if random_sequence_vf[j] == 0:
gets_stim = lVF
no_stim = rVF
else:
gets_stim = rVF
no_stim = lVF
s = random_sequence_stim2[j]
s_stim = stim[s]
l_stim = labels[s]
#Precortical for both visual fields
y_stim = gets_stim.activation(s_stim, 0)
y_zero = no_stim.activation(zero_stim, 0)
gets_stim.update(s_stim, 0)
no_stim.update(zero_stim, 0)
#Early cortical
y_stim = gets_stim.activation(y_stim, 1)
y_zero = no_stim.activation(y_zero, 1)
gets_stim.update(y_stim, 1)
no_stim.update(y_zero, 1)
#Ventral visual
gets_stim.activation(y_stim, 2, cat_input=l_stim)
no_stim.activation(y_zero, 2)
gets_stim.update(y_stim, 2, cat_input=l_stim, category_training=True)
no_stim.update(y_zero, 2)
| Categorical perception model of human vision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sqlalchemy import create_engine
import time
# -
tweet_df = pd.read_csv('expert_tweets_week_3.csv')
tweet_df.shape
# Merge Data & Save to SQLite Database
# +
###FUNCTION CREATE TABLES IN OUR SQLITE DATABASE - USE A FIXED DATABASE NAME OF fantasy_football_2018.db###
#function inputs are the dataframe, the table_name, and the
#if_exists_action can be 'fail', 'replace', 'append' (default is set to 'append' if user doesn't put in anything)
def add_to_SQLite(df, table_name, if_exists_action='append'):
default_database_path = 'sqlite:///fantasy_football_2018.db'
disk_engine = create_engine(default_database_path)
df.to_sql(table_name, disk_engine,
if_exists=if_exists_action, index=False,
chunksize=100) #chunksize limits how many variables get added at a time (SQLite needed max of 100 or would error out)
print(f"Table {table_name} was added to the database at {default_database_path}")
# -
###ADD WEEK 3 TWEETS TO SQL DB
add_to_SQLite(df=tweet_df,
table_name='week_3_tweets',
if_exists_action='fail')
week_3_ou_df = pd.read_csv('week_3_score_sent.csv')
###ADD WEEK 3 OVER UNDERS TO SQL DB
add_to_SQLite(df=week_3_ou_df,
table_name='week_3_over_unders',
if_exists_action='fail')
def_scores_df = pd.read_csv('binned_defenses.csv')
###ADD WEEK 3 OVER UNDERS TO SQL DB
add_to_SQLite(df=def_scores_df,
table_name='def_scores',
if_exists_action='replace')
| data/connor_to_sqlite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: D2GO
# language: python
# name: d2go
# ---
# +
import urllib
import json
import numpy as np
import itertools
import torch
from os import listdir
import os
from os.path import isfile, join
import time
import requests
import json
import pickle
import base64
from io import BytesIO
from PIL import Image
from tqdm import tqdm
import cv2
from matplotlib import pyplot as plt
def find_image_by_id(id: int):
for i in data['images']:
if i['id'] == id:
return i
f = open('fishial_collection_correct.json',)
data = json.load(f)
f.close()
bodyes_shapes_ids = []
for i in data['categories']:
if i['name'] == 'General body shape':
bodyes_shapes_ids.append(int(i['id']))
# -
folder_name = "fishial_collection"
os.makedirs(folder_name, exist_ok=True)
os.makedirs("{}/Train".format(folder_name), exist_ok=True)
os.makedirs("{}/Test".format(folder_name), exist_ok=True)
list_sd = []
for i in tqdm(range(len(data['images']))):
if 'train_data' not in data['images'][i]:
continue
list_sd.append(data['images'][i]['file_name'])
folder_type = 'Train' if data['images'][i]['train_data'] else 'Test'
path = os.path.join(os.path.join(folder_name, folder_type), data['images'][i]['file_name'])
start_time = time.time()
r = requests.get(data['images'][i]['coco_url'], allow_redirects=True) # to get content after redirection
with open(path, 'wb') as f:
f.write(r.content)
print("[FISHIAL] Request to fishial: {} path: {}".format(time.time() - start_time, path), end='\r')
def find_image_by_id(id: int):
for i in data['images']:
if i['id'] == id:
return i
count_withount_poly = 0
for i in range(len(data['annotations'])):
if len(data['annotations'][i]['segmentation'][0]) < 30:
poly_gon = []
for z in range(int(len(data['annotations'][i]['segmentation'][0])/2) - 1):
poly_gon.append([data['annotations'][i]['segmentation'][0][z*2],
data['annotations'][i]['segmentation'][0][z*2 +1]])
image_instance = find_image_by_id(data['annotations'][i]['image_id'])
file_name = image_instance['file_name']
img_path = image_instance['coco_url']
r = requests.get(img_path, allow_redirects=True) # to get content after redirection
with open(file_name, 'wb') as f:
f.write(r.content)
count_withount_poly += 1
image = cv2.cvtColor(cv2.imread(file_name), cv2.COLOR_BGR2RGB)
poly_gon_transf = np.array([poly_gon], np.int32)
cv2.polylines(image, [poly_gon_transf], True, (0,255,0), thickness=3)
cv2.imshow('Check', image)
cv2.waitKey(0)
cv2.destroyAllWindows()
# print(count_withount_poly)
#Import image
image = cv2.imread("input_path")
#Show the image with matplotlib
plt.imshow(image)
plt.show()
| helper/segmentation/examples/Preprocess_Fishial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MLSMOTE
#
# A python code that implemented the [MLSMOTE](https://www.sciencedirect.com/science/article/abs/pii/S0950705115002737) algorithm was available here: https://github.com/niteshsukhwani/MLSMOTE. However, the code had a bug and wasn't efficiently using the pandas. I fixed and modified the code, and here it is.
#
# **If you find this notebook useful, please don't forget to upvote.**
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np
import pandas as pd
import random
from sklearn.datasets import make_classification
from sklearn.neighbors import NearestNeighbors
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
def create_dataset(n_sample=1000):
'''
Create a unevenly distributed sample data set multilabel
classification using make_classification function
args
nsample: int, Number of sample to be created
return
X: pandas.DataFrame, feature vector dataframe with 10 features
y: pandas.DataFrame, target vector dataframe with 5 labels
'''
X, y = make_classification(n_classes=5, class_sep=2,
weights=[0.1,0.025, 0.205, 0.008, 0.9], n_informative=3, n_redundant=1, flip_y=0,
n_features=10, n_clusters_per_class=1, n_samples=1000, random_state=10)
y = pd.get_dummies(y, prefix='class')
return pd.DataFrame(X), y
def get_tail_label(df: pd.DataFrame, ql=[0.05, 1.]) -> list:
"""
Find the underrepresented targets.
Underrepresented targets are those which are observed less than the median occurance.
Targets beyond a quantile limit are filtered.
"""
irlbl = df.sum(axis=0)
irlbl = irlbl[(irlbl > irlbl.quantile(ql[0])) & ((irlbl < irlbl.quantile(ql[1])))] # Filtering
irlbl = irlbl.max() / irlbl
threshold_irlbl = irlbl.median()
tail_label = irlbl[irlbl > threshold_irlbl].index.tolist()
return tail_label
def get_minority_samples(X: pd.DataFrame, y: pd.DataFrame, ql=[0.05, 1.]):
"""
return
X_sub: pandas.DataFrame, the feature vector minority dataframe
y_sub: pandas.DataFrame, the target vector minority dataframe
"""
tail_labels = get_tail_label(y, ql=ql)
index = y[y[tail_labels].apply(lambda x: (x == 1).any(), axis=1)].index.tolist()
X_sub = X[X.index.isin(index)].reset_index(drop = True)
y_sub = y[y.index.isin(index)].reset_index(drop = True)
return X_sub, y_sub
def nearest_neighbour(X: pd.DataFrame, neigh) -> list:
"""
Give index of 10 nearest neighbor of all the instance
args
X: np.array, array whose nearest neighbor has to find
return
indices: list of list, index of 5 NN of each element in X
"""
nbs = NearestNeighbors(n_neighbors=neigh, metric='euclidean', algorithm='kd_tree').fit(X)
euclidean, indices = nbs.kneighbors(X)
return indices
def MLSMOTE(X, y, n_sample, neigh=5):
"""
Give the augmented data using MLSMOTE algorithm
args
X: pandas.DataFrame, input vector DataFrame
y: pandas.DataFrame, feature vector dataframe
n_sample: int, number of newly generated sample
return
new_X: pandas.DataFrame, augmented feature vector data
target: pandas.DataFrame, augmented target vector data
"""
indices2 = nearest_neighbour(X, neigh=5)
n = len(indices2)
new_X = np.zeros((n_sample, X.shape[1]))
target = np.zeros((n_sample, y.shape[1]))
for i in range(n_sample):
reference = random.randint(0, n-1)
neighbor = random.choice(indices2[reference, 1:])
all_point = indices2[reference]
nn_df = y[y.index.isin(all_point)]
ser = nn_df.sum(axis = 0, skipna = True)
target[i] = np.array([1 if val > 0 else 0 for val in ser])
ratio = random.random()
gap = X.loc[reference,:] - X.loc[neighbor,:]
new_X[i] = np.array(X.loc[reference,:] + ratio * gap)
new_X = pd.DataFrame(new_X, columns=X.columns)
target = pd.DataFrame(target, columns=y.columns)
return new_X, target
# -
X, y = create_dataset() # Creating a Dataframe
X_sub, y_sub = get_minority_samples(X, y) # Getting minority samples of that datframe
X_res, y_res = MLSMOTE(X_sub, y_sub, 100, 5) # Applying MLSMOTE to augment the dataframe
y_res.head()
X_res.head()
| kaggle_notebooks/upsampling-multilabel-data-with-mlsmote.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
# %matplotlib inline
pd.__version__
np.__version__
sns.__version__
# 1. class: lung, head & neck, esophasus, thyroid, stomach, duoden & sm.int,
# colon, rectum, anus, salivary glands, pancreas, gallblader,
# liver, kidney, bladder, testis, prostate, ovary, corpus uteri,
# cervix uteri, vagina, breast
# 2. age: <30, 30-59, >=60
# 3. sex: male, female
# 4. histologic-type: epidermoid, adeno, anaplastic
# 5. degree-of-diffe: well, fairly, poorly
# 6. bone: yes, no
# 7. bone-marrow: yes, no
# 8. lung: yes, no
# 9. pleura: yes, no
# 10. peritoneum: yes, no
# 11. liver: yes, no
# 12. brain: yes, no
# 13. skin: yes, no
# 14. neck: yes, no
# 15. supraclavicular: yes, no
# 16. axillar: yes, no
# 17. mediastinum: yes, no
# 18. abdominal: yes, no
col_names = ['class', 'age', 'sex', 'histologic-type','degree-of-diffe', 'bone', 'bone-narrow',
'lung', 'pleura', 'peritoneum', 'liver', 'brain', 'skin', 'neck', 'supraclavicular',
'axillar','mediastinum', 'abdominal']
dataset = pd.read_csv('data/primary-tumor.data',names = col_names, sep=',')
dataset.head(15)
dataset.replace('?', np.nan, inplace = True)
dataset.head(15)
dataset.info()
dataset.isnull().values.any()
dataset.isnull().sum()
| projects/project_01_AVC_risk/.ipynb_checkpoints/primary_tumor-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# Create a practice set of random latitude and longitude combinations.
x = [25.12903645, 25.92017388, 26.62509167, -59.98969384, 37.30571269]
y = [-67.59741259, 11.09532135, 74.84233102, -76.89176677, -61.13376282]
coordinates = zip(x, y)
# Iterate to display combinations
for coordinate in coordinates:
print(coordinate[0], coordinate[1])
# Use citipy module to determine city from latitude and longitude
from citipy import citipy
for coordinate in coordinates:
print(citipy.nearest_city(coordinate[0], coordinate[1]).city_name,
citipy.nearest_city(coordinate[0], coordinate[1]).country_code)
# +
# Import the requests library.
import requests
# Import the API key.
from config import open_weather_api_key
# -
# Starting URL for Weather Map API Call.
url = "http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID=" + open_weather_api_key
# Create endppoint url for city
city_url = url + "&q=" + "Boston"
# Make a 'Get' request for the city weather.
city_weather = requests.get(city_url)
city_weather
# Create a bad endpoint URL for a city.
city_url = url + "&q=" + "Bston"
city_weather = requests.get(city_url)
city_weather
# Turn get request into text.
city_url = url + "&q=" + "Boston"
city_weather = requests.get(city_url)
city_weather.text
# Get the JSON text of the 'Get' request.
city_weather.json()
# Create an endpoint URL for a city.
city_url = url + "&q=" + "Boston"
city_weather = requests.get(city_url)
if city_weather.status_code == 200:
print(f"City Weather found.")
else:
print(f"City weather not found.")
# Create an endpoint URL for a city.
city_url = url + "&q=" + "Bston"
city_weather = requests.get(city_url)
if city_weather.json():
print(f"City Weather found.")
else:
print(f"City weather not found.")
# Create an endpoint URL for a city.
city_url = url + "&q=" + "Boston"
city_weather = requests.get(city_url)
city_weather.json()
# Get JSON data
boston_data = city_weather.json()
boston_data["sys"]
boston_data["sys"]["country"]
boston_data["dt"]
lat = boston_data["coord"]["lat"]
lon = boston_data["coord"]["lon"]
max_temp = boston_data["main"]["temp_max"]
humidity = boston_data["main"]["humidity"]
clouds = boston_data["clouds"]["all"]
wind = boston_data["wind"]["speed"]
print(lat, lon, max_temp, humidity, clouds, wind)
# Import the datetime module from the datetime library.
from datetime import datetime
# Get the date from the JSON file.
date = boston_data["dt"]
# Convert the UTC date to a date format with year, month, day, hours, minutes, and seconds.
datetime.utcfromtimestamp(date)
# Import the time module.
import time
# Get today's date in seconds.
today = time.time()
today
import time
today = time.strftime("%x")
today
| API_practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
from pixell import utils, enmap, enplot, curvedsky
import healpy as hp
from multiprocessing import Pool
from cosmikyu import sehgal, stats, utils as cutils
from orphics import sehgal as osehgal
import matplotlib.pyplot as plt
import scipy
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
input_dir = "/home/dwhan89/scratch/data/sehgal_et_al_sims/cosmo_sim_maps/July2009/output/catalogues"
def input_path(file_name):
return os.path.join(input_dir, file_name)
# +
overwrite=False
radio_source_file = input_path("radio.cat")
radio_spectra_index_file = input_path("radio_spectra_indexv2.npz")
if not os.path.exists(radio_spectra_index_file) or overwrite:
print("computing")
## 1.4, 30, 90, 148, 219, 277, 350 GHz
radio_cat = np.loadtxt(radio_source_file, usecols=(3,4,5,6,7,8,9), dtype=np.float32)
loc = np.where(radio_cat[:,3]<=7)
radio_cat = radio_cat[loc[0],:]
radio_spectra_index = radio_cat/radio_cat[:,3][:,None]
radio_spectra_index = radio_spectra_index[:,[0,1,2,4,5,6]]
radio_spectra_index = np.log(radio_spectra_index)
freq_ratio = np.array([1.4, 30, 90, 219, 277, 350])/148
ln_freq_ratio = np.log(freq_ratio)
radio_spectra_index = radio_spectra_index/(ln_freq_ratio[:,None].T)
mean = np.mean(radio_spectra_index,axis=0)
cov = np.cov(radio_spectra_index.T)
radio_spectra_index = {"mean": mean[1:], "cov": cov[1:,1:]} ## drop 1.4
#radio_spectra_index = np.mean(radio_spectra_index, axis=1)
np.savez(radio_spectra_index_file, **radio_spectra_index)
else:
print("loading")
radio_spectra_index = np.load(radio_spectra_index_file)
# -
radio_spectra_index_file
np.empty(shape=(1,1), dtype=np.float32)
np.random.multivariate_normal([0,0],[[1,0],[0,1]], size=(2,2)).shape
# +
fig = plt.figure(figsize=(6,6))
ax = fig.gca()
#freqs = [1.4, 30, 90, 219, 277, 350]
freqs = [30, 90, 219, 277, 350]
std = np.sqrt(np.diag(radio_spectra_index['cov']))
plt.errorbar(freqs, radio_spectra_index['mean'], std, marker="o", color="r", markersize=10, ls="", label="per freq")
plt.ylabel("Radio Spectra Index", fontsize=15)
plt.xlabel("Freq (GHz)", fontsize=15)
upper = np.array([-0.81 + 0.11]*5)
lower = np.array([-0.81 - 0.11]*5)
ax.fill_between(freqs, lower, upper, color="blue", alpha=0.2, label="average spectral index")
plt.axhline(-0.81, color="b")
plt.legend(fontsize=15)
ax.tick_params(axis='both', which='major', labelsize=15)
ax.tick_params(axis='both', which='minor', labelsize=15)
plt.ylim(-1,-0.6)
plt.show()
cov = radio_spectra_index['cov'].copy()
corr = cov/np.outer(std,std)
fig, ax = plt.subplots(figsize=(6,6))
im = ax.imshow(corr, vmin=-1, vmax=1)
plt.colorbar(im)
# We want to show all ticks...
ax.set_xticks(np.arange(len(freqs)))
ax.set_yticks(np.arange(len(freqs)))
# ... and label them with the respective list entries
ax.set_xticklabels(freqs)
ax.set_yticklabels(freqs)
plt.title("Correlation Matrix", fontsize=15)
plt.xlabel("Freq (GHz)", fontsize=15)
plt.ylabel("Freq (GHz)", fontsize=15)
plt.show()
# +
overwrite=True
blast_source_file = input_path("IRBlastPop.dat")
cib_spectra_index_file = input_path("cib_spectra_indexv2.npz")
cib_spectra_index = None
input_files = [blast_source_file]
for i in range(1,11):
input_files.append(input_path(f"IRgal_S_{i}.dat"))
if not os.path.exists(cib_spectra_index_file) or overwrite:
for input_file in input_files:
print(f"processing {input_file}")
#30, 90, 148, 219, 277, 350
cib_cat = np.loadtxt(input_file, usecols=(4,5,6,7,8,9), dtype=np.float32)
print(cib_cat.shape)
cib_cat *= 0.75 ## apply scaling <= Double check this !!!
loc = np.where(cib_cat[:,2]<=7)
if len(loc[0]) == 0:
print(f"skipping {input_file}")
continue
cib_cat = cib_cat[loc[0],:]
index = cib_cat/cib_cat[:,2][:,None]
index = index[:,[0,1,3,4,5]]
index = np.log(index)
freq_ratio = np.array([30, 90, 219, 277, 350])/148
ln_freq_ratio = np.log(freq_ratio)
index = index/(ln_freq_ratio[:,None].T)
#index = np.mean(index, axis=1)
if cib_spectra_index is None:
cib_spectra_index = index
else:
cib_spectra_index = np.vstack((cib_spectra_index, index))
print(cib_spectra_index.shape)
mean = np.mean(cib_spectra_index,axis=0)
cov = np.cov(cib_spectra_index.T)
cib_spectra_index = {"mean": mean, "cov":cov}
np.savez(cib_spectra_index_file, **cib_spectra_index)
else:
cib_spectra_index = np.load(cib_spectra_index_file)
# +
fig = plt.figure(figsize=(6,6))
ax = fig.gca()
freqs = [30, 90, 219, 277, 350]
std = np.sqrt(np.diag(cib_spectra_index['cov']))
plt.errorbar(freqs, cib_spectra_index['mean'], std, marker="o", color="r", markersize=10, ls="", label="per freq")
plt.ylabel("CIB Spectra Index", fontsize=15)
plt.xlabel("Freq (GHz)", fontsize=15)
upper = np.array([3.02 + 0.17]*5)
lower = np.array([3.02 - 0.17]*5)
ax.fill_between(freqs, lower, upper, color="blue", alpha=0.2, label="average spectral index")
plt.axhline(3.02, color="b")
plt.legend(fontsize=15)
ax.tick_params(axis='both', which='major', labelsize=15)
ax.tick_params(axis='both', which='minor', labelsize=15)
plt.ylim(2,4)
plt.show()
cov = cib_spectra_index['cov'].copy()
corr = cov/np.outer(std,std)
fig, ax = plt.subplots(figsize=(6,6))
im = ax.imshow(corr, vmin=-1, vmax=1)
plt.colorbar(im)
# We want to show all ticks...
ax.set_xticks(np.arange(len(freqs)))
ax.set_yticks(np.arange(len(freqs)))
# ... and label them with the respective list entries
ax.set_xticklabels(freqs)
ax.set_yticklabels(freqs)
plt.title("Correlation Matrix", fontsize=15)
plt.xlabel("Freq (GHz)", fontsize=15)
plt.ylabel("Freq (GHz)", fontsize=15)
plt.show()
# +
nbins = 100
minval = np.min(cib_spectra_index)
maxval = np.max(cib_spectra_index)
mean = np.mean(cib_spectra_index)
std = np.std(cib_spectra_index)
FB = stats.FastBINNER(minval, maxval, nbins)
bin_center, index_binned = FB.bin(cib_spectra_index)
fig = plt.figure(figsize=(12, 12))
ax = plt.gca()
plt.plot(bin_center, index_binned, label=f"mean={mean:.2f}, std={std:.2f}")
plt.yscale("log")
plt.legend(fontsize=30)
plt.title("CIB Spectra Index (Histogram)", fontsize=30)
plt.ylabel("N", fontsize=30)
plt.xlabel(r"Spectral Index$", fontsize=30)
plt.yscale("linear")
#plt.ylim(1e-1,0.25)
plt.minorticks_on()
ax.tick_params(axis='x', which='minor', bottom=True)
plt.grid()
plt.show()
# -
cib_spectra_index
| notebooks/040821_spectral_indxes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
# +
from math import log
from statistics import mean, median, stdev
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# -
datasets = ["abalone", "adult", "cancer", "card", "covtype", "gene", "glass", "heart", "horse", "madelon", "optdigits", "page-blocks", "pendigits", "poker", "satimage", "segmentation", "shuttle", "soybean", "spect", "thyroid", "vehicle", "waveform"]
ms = [110, 150, 5, 5, 180, 5, 5, 5, 5, 5, 175, 100, 185, 185, 180, 25, 70, 5, 150, 145, 115, 60]
# Difference in training time for Stepwise between mingood and 25
mediandf = []
for file in ["stepwise", "mingood_stepwise"]:
df = pd.read_csv(f"../log/performance_{file}.txt")[["dataset", "time"]]
df = pd.pivot_table(df, index="dataset", aggfunc=median)
mediandf.append(df)
mediandf = pd.DataFrame({'mmingood': ms, 'time25': mediandf[0].time.to_list(), 'timemingood': mediandf[1].time.to_list()}, index=datasets)
mediandf["gain"] = mediandf.timemingood - mediandf.time25
mediandf["pct"] = (mediandf.gain / mediandf.time25 * 100)
mediandf["mgain"] = mediandf.mmingood - 25
mediandf.time25 = mediandf.time25.map(lambda t: "%s" % float("%.4g" % t))
mediandf.timemingood = mediandf.timemingood.map(lambda t: "%s" % float("%.4g" % t))
mediandf.gain = mediandf.gain.map(lambda t: "%s" % float("%.4g" % t))
mediandf.pct = mediandf.pct.map(lambda p: f'{"%s" % float("%.4g" % p)}%')
mediandf.to_csv("stepwise_time_diff.csv")
mediandf
| dataproc/misctest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import json
pd.set_option('display.max_rows', 500)
import requests
from bs4 import BeautifulSoup
# -
# # Data Understanding
#
# ## Data Sources
# * RKI, webscraping https://www.rki.de/DE/Content/InfAZ/N/Neuartiges_Coronavirus/Fallzahlen.html
# * <NAME> (GIT) https://github.com/CSSEGISandData/COVID-19.git
# * Rest API to retrieve covid data from NPGEO https://npgeo-corona-npgeo-de.hub.arcgis.com/
#
# ### <NAME>ins Source
data_path='../data/raw/COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv'
pd_raw=pd.read_csv(data_path)
pd_raw.head()
# ### Web Scraping from RKI
#
page = requests.get("https://www.rki.de/DE/Content/InfAZ/N/Neuartiges_Coronavirus/Fallzahlen.html")
# + tags=[]
soup = BeautifulSoup(page.content, 'html.parser')
html_table = soup.find('table')
all_rows = html_table.find_all('tr')
print(all_rows)
# + tags=[]
final_table_data = []
for pos,rows in enumerate(all_rows):
col_list=[each_col.get_text(strip=True) for each_col in rows.find_all('td')] #td for data element
final_table_data.append(col_list)
print(final_table_data)
# + tags=[]
pd.DataFrame(final_table_data).dropna().rename(columns={0:'state',
1:'cases',
2:'changes',
3:'cases_per_100k',
4:'fatality',
5:'comment'}).head()
# -
# ## REST API CALLS
## data request for Germany
data=requests.get('https://services7.arcgis.com/mOBPykOjAyBO2ZKk/arcgis/rest/services/Coronaf%C3%A4lle_in_den_Bundesl%C3%A4ndern/FeatureServer/0/query?where=1%3D1&outFields=*&outSR=4326&f=json')
json_object=json.loads(data.content)
json_object.keys()
full_list=[]
for pos,each_dict in enumerate (json_object['features'][:]):
full_list.append(each_dict['attributes'])
pd_full_list=pd.DataFrame(full_list)
pd_full_list.head()
pd_full_list.to_csv('../data/raw/NPGEO/GER_state_data.csv',sep=';')
pd_full_list
pd_full_list.shape[0]
# ## API access via REST service, e.g. USA data
#
# + tags=[]
# US for full list
headers = {
'Cache-Control': 'no-cache',
'Subscription-Key': '28ee4219700f48718be78b057beb7eb4',
}
response = requests.get('https://api.smartable.ai/coronavirus/stats/US', headers=headers)
if response.status_code != 200:
print("Something Wrong with Request!!")
else:
print("Request Success!")
# -
US_dict=json.loads(response.content) # imports string
with open('../data/raw/SMARTABLE/US_data.json', 'w') as outfile:
json.dump(US_dict, outfile,indent=2)
# + tags=["outputPrepend"]
print(json.dumps(US_dict,indent=2))
# + tags=[]
full_list_US_country=[]
for pos,each_dict in enumerate (US_dict['stats']['breakdowns'][:]):
flatten_dict=each_dict['location']
flatten_dict.update(dict(list(US_dict['stats']['breakdowns'][pos].items())[1: 7])
)
full_list_US_country.append(flatten_dict)
# -
pd.DataFrame(full_list_US_country).to_csv('../data/raw/SMARTABLE/full_list_US_country.csv',sep=';',index=False)
| notebooks/Data Understanding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BiGAN with MNIST (or Fashion MNIST)
#
# * `Adversarial Feature Learning`, [arXiv:1605.09782](https://arxiv.org/abs/1605.09782)
# * <NAME>, <NAME>, and <NAME>
#
# * This code is available to tensorflow version 2.0
# * Implemented by [`tf.keras.layers`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers) [`tf.losses`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/losses)
# * Use `transposed_conv2d` and `conv2d` for Generator and Discriminator, respectively.
# * I do not use `dense` layer for model architecture consistency. (So my architecture is different from original dcgan structure)
# * based on DCGAN model
# ## Import modules
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __future__ import unicode_literals
import os
import sys
import time
import glob
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import PIL
import imageio
from IPython import display
import tensorflow as tf
from tensorflow.keras import layers
sys.path.append(os.path.dirname(os.path.abspath('.')))
from utils.image_utils import *
from utils.ops import *
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# -
# ## Setting hyperparameters
# +
# Training Flags (hyperparameter configuration)
model_name = 'bigan'
train_dir = os.path.join('train', model_name, 'exp1')
dataset_name = 'mnist'
assert dataset_name in ['mnist', 'fashion_mnist']
max_epochs = 50
save_model_epochs = 10
print_steps = 100
save_images_epochs = 1
batch_size = 256
learning_rate_D = 1e-4
learning_rate_G = 1e-4
k = 1 # the number of step of learning D before learning G (Not used in this code)
num_examples_to_generate = 25
noise_dim = 100
# -
# ## Load the MNIST dataset
# +
# Load training and eval data from tf.keras
if dataset_name == 'mnist':
(train_images, train_labels), (test_images, test_labels) = \
tf.keras.datasets.mnist.load_data()
else:
(train_images, train_labels), (test_images, test_labels) = \
tf.keras.datasets.fashion_mnist.load_data()
train_images = train_images.reshape(-1, MNIST_SIZE, MNIST_SIZE, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
test_images = test_images.reshape(-1, MNIST_SIZE, MNIST_SIZE, 1).astype('float32')
test_images = (test_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
# -
# ## Set up dataset with `tf.data`
#
# ### create input pipeline with `tf.data.Dataset`
# +
#tf.random.set_seed(219)
# for train
N = len(train_images)
train_dataset = tf.data.Dataset.from_tensor_slices(train_images)
train_dataset = train_dataset.shuffle(buffer_size=N)
train_dataset = train_dataset.batch(batch_size=batch_size, drop_remainder=True)
print(train_dataset)
# for test
test_dataset = tf.data.Dataset.from_tensor_slices(test_images)
test_dataset = test_dataset.shuffle(buffer_size=N)
test_dataset = test_dataset.batch(batch_size=num_examples_to_generate, drop_remainder=True)
print(test_dataset)
# + [markdown] cell_style="center"
# ## Create the generator and discriminator models
# + cell_style="center"
class Generator(tf.keras.Model):
"""Build a generator that maps latent space to real space.
G(z): z -> x
"""
def __init__(self):
super(Generator, self).__init__()
self.conv1 = ConvTranspose(256, 3, padding='valid')
self.conv2 = ConvTranspose(128, 3, padding='valid')
self.conv3 = ConvTranspose(64, 4)
self.conv4 = ConvTranspose(1, 4, apply_batchnorm=False, activation='tanh')
def call(self, inputs, training=True):
"""Run the model."""
# inputs: [1, 1, 100]
conv1 = self.conv1(inputs, training=training) # conv1: [3, 3, 256]
conv2 = self.conv2(conv1, training=training) # conv2: [7, 7, 128]
conv3 = self.conv3(conv2, training=training) # conv3: [14, 14, 64]
generated_images = self.conv4(conv3, training=training) # generated_images: [28, 28, 1]
return generated_images
# -
class Encoder(tf.keras.Model):
"""Build a encoder that maps real space to latent space.
E(x): x -> z
"""
def __init__(self):
super(Encoder, self).__init__()
self.conv1 = Conv(64, 4, 2, apply_batchnorm=False, activation='leaky_relu')
self.conv2 = Conv(128, 4, 2, activation='leaky_relu')
self.conv3 = Conv(256, 3, 2, padding='valid', activation='leaky_relu')
self.conv4 = Conv(100, 3, 1, padding='valid', apply_batchnorm=False, activation='none')
def call(self, inputs, training=True):
"""Run the model."""
# inputs: [28, 28, 1]
conv1 = self.conv1(inputs, training=training) # conv1: [14, 14, 64]
conv2 = self.conv2(conv1, training=training) # conv2: [7, 7, 128]
conv3 = self.conv3(conv2, training=training) # conv3: [3, 3, 256]
z_vector = self.conv4(conv3, training=training) # z_vector: [1, 1, 100]
return z_vector
class Discriminator(tf.keras.Model):
"""Build a discriminator that discriminate tuple (x, z) whether real or fake.
D(x, z): (x, z) -> [0, 1]
"""
def __init__(self):
super(Discriminator, self).__init__()
self.fc = layers.Dense(MNIST_SIZE * MNIST_SIZE * 2, activation=tf.nn.leaky_relu)
self.conv1 = Conv(64, 4, 2, apply_batchnorm=False, activation='leaky_relu')
self.conv2 = Conv(128, 4, 2, activation='leaky_relu')
self.conv3 = Conv(256, 3, 2, padding='valid', activation='leaky_relu')
self.conv4 = Conv(1, 3, 1, padding='valid', apply_batchnorm=False, activation='none')
def call(self, image_inputs, latent_codes, training=True):
"""Run the model."""
# latent_code: [1, 1, 100]
latent_codes = tf.squeeze(latent_codes, axis=[1, 2])
fc = self.fc(latent_codes)
fc = tf.reshape(fc, [-1, MNIST_SIZE, MNIST_SIZE, 2])
# concatnate image_inputs + latent code
# image_inputs: [28, 28, 1] + fc: [28, 28, 2]
inputs = tf.concat([image_inputs, fc], axis=3) # inputs: [28, 28, 3]
conv1 = self.conv1(inputs) # conv1: [14, 14, 64]
conv2 = self.conv2(conv1) # conv2: [7, 7, 128]
conv3 = self.conv3(conv2) # conv3: [3, 3, 256]
conv4 = self.conv4(conv3) # conv4: [1, 1, 1]
discriminator_logits = tf.squeeze(conv4, axis=[1, 2]) # discriminator_logits: [1,]
return discriminator_logits
generator = Generator()
encoder = Encoder()
discriminator = Discriminator()
# ### Plot generated image via generator network
# +
noise = tf.random.normal([1, 1, 1, noise_dim])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
# -
# ### Test encoder network
z_hat = encoder(generated_image)
print(z_hat)
# ### Test discriminator network
#
# * **CAUTION**: the outputs of discriminator is **logits** (unnormalized probability) NOT probabilites
decision = discriminator(generated_image, noise)
print(decision)
# ## Define the loss functions and the optimizer
# use logits for consistency with previous code I made
# `tf.losses` and `tf.keras.losses` are the same API (alias)
bce = tf.losses.BinaryCrossentropy(from_logits=True)
def GANLoss(logits, is_real=True):
"""Computes standard GAN loss between `logits` and `labels`.
Args:
logits (`2-rank Tensor`): logits.
is_real (`bool`): True means `1` labeling, False means `0` labeling.
Returns:
loss (`0-rank Tensor`): the standard GAN loss value. (binary_cross_entropy)
"""
if is_real:
labels = tf.ones_like(logits)
else:
labels = tf.zeros_like(logits)
return bce(labels, logits)
def discriminator_loss(real_logits, fake_logits):
# losses of real with label "1"
real_loss = GANLoss(logits=real_logits, is_real=True)
# losses of fake with label "0"
fake_loss = GANLoss(logits=fake_logits, is_real=False)
return real_loss + fake_loss
def generator_loss(real_logits, fake_logits):
# losses of fake with label "0" that used to fool the discriminator
real_loss = GANLoss(logits=real_logits, is_real=False)
# losses of Generator with label "1" that used to fool the discriminator
fake_loss = GANLoss(logits=fake_logits, is_real=True)
return real_loss + fake_loss
#discriminator_optimizer = tf.keras.optimizers.Adam(learning_rate_D, beta_1=0.5)
discriminator_optimizer = tf.keras.optimizers.RMSprop(learning_rate_D)
generator_optimizer = tf.keras.optimizers.Adam(learning_rate_G, beta_1=0.5)
# ## Checkpoints (Object-based saving)
checkpoint_dir = train_dir
if not tf.io.gfile.exists(checkpoint_dir):
tf.io.gfile.makedirs(checkpoint_dir)
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# ## Training
# keeping the random image constant for generation (prediction) so
# it will be easier to see the improvement of the gan.
# To visualize progress in the animated GIF
for inputs in test_dataset.take(1):
constant_test_input = inputs
# ### Define training one step function
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
# generating noise from a uniform distribution
noise = tf.random.uniform([batch_size, 1, 1, noise_dim], minval=-1.0, maxval=1.0)
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
# generating images from generator() via random noise vector z
generated_images = generator(noise, training=True)
# inference latent codes from encoder via real images
inference_codes = encoder(images, training=True)
# discriminating tuple (real images, inference_codes) by discriminator()
real_logits = discriminator(images, inference_codes, training=True)
# discriminating tuple (generated_images, random_z) by discriminator()
fake_logits = discriminator(generated_images, noise, training=True)
gen_loss = generator_loss(real_logits, fake_logits)
disc_loss = discriminator_loss(real_logits, fake_logits)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables + encoder.trainable_variables)
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
generator_optimizer.apply_gradients(zip(gradients_of_generator,
generator.trainable_variables + encoder.trainable_variables))
return gen_loss, disc_loss
# ### Train full steps
# +
print('Start Training.')
num_batches_per_epoch = int(N / batch_size)
global_step = tf.Variable(0, trainable=False)
for epoch in range(max_epochs):
for step, images in enumerate(train_dataset):
start_time = time.time()
gen_loss, disc_loss = train_step(images)
global_step.assign_add(1)
if global_step.numpy() % print_steps == 0:
epochs = epoch + step / float(num_batches_per_epoch)
duration = time.time() - start_time
examples_per_sec = batch_size / float(duration)
display.clear_output(wait=True)
print("Epochs: {:.2f} global_step: {} loss_D: {:.3g} loss_G: {:.3g} ({:.2f} examples/sec; {:.3f} sec/batch)".format(
epochs, global_step.numpy(), disc_loss, gen_loss, examples_per_sec, duration))
for images in test_dataset.take(1):
sample_inference_codes = encoder(images, training=False)
reconstruction_images = generator(sample_inference_codes, training=False)
print_or_save_sample_images_two(images.numpy(),
reconstruction_images.numpy(),
num_examples_to_generate)
if (epoch + 1) % save_images_epochs == 0:
display.clear_output(wait=True)
print("This images are saved at {} epoch".format(epoch+1))
sample_inference_codes = encoder(constant_test_input, training=False)
reconstruction_images = generator(sample_inference_codes, training=False)
print_or_save_sample_images_two(constant_test_input.numpy(),
reconstruction_images.numpy(),
num_examples_to_generate,
is_save=True, epoch=epoch+1,
checkpoint_dir=checkpoint_dir)
# saving (checkpoint) the model every save_epochs
if (epoch + 1) % save_model_epochs == 0:
checkpoint.save(file_prefix=checkpoint_prefix)
print('Training Done.')
# -
# generating after the final epoch
display.clear_output(wait=True)
sample_inference_codes = encoder(constant_test_input, training=False)
reconstruction_images = generator(sample_inference_codes, training=False)
print_or_save_sample_images_two(constant_test_input.numpy(),
reconstruction_images.numpy(),
num_examples_to_generate,
is_save=True, epoch=epoch+1,
checkpoint_dir=checkpoint_dir)
# ## Restore the latest checkpoint
# restoring the latest checkpoint in checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# ## Display an image using the epoch number
display_image(max_epochs, checkpoint_dir=checkpoint_dir)
# ## Generate a GIF of all the saved images.
filename = model_name + '_' + dataset_name + '.gif'
generate_gif(filename, checkpoint_dir)
display.Image(filename=filename + '.png')
| gans/bigan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # fastgpu
#
# > API details
# +
# default_exp core
# -
#export
import os
os.environ['CUDA_DEVICE_ORDER']='PCI_BUS_ID'
from fastcore.all import *
from pynvml import *
from fastcore.nb_imports import *
from threading import Thread
import subprocess
from uuid import uuid4
#hide
from nbdev.showdoc import *
# ## Overview
# Here's what fastgpu does:
#
# 1. poll `to_run`
# 1. find first file
# 1. check there's an available worker id
# 1. move it to `running`
# 1. handle the script
# 1. create lock file
# 1. redirect stdout/err to `out`
# 1. run it
# 1. when done, move it to `complete` or `failed`
# 1. unlock
# For demonstrating how to use `fastgpu`, we first create a directory to store our scripts and outputs:
path = Path('data')
#export
def setup_dirs(path):
"Create and return the following subdirs of `path`: to_run running complete fail out"
path.mkdir(exist_ok=True)
dirs = L(path/o for o in 'to_run running complete fail out'.split())
for o in dirs: o.mkdir(exist_ok=True)
return dirs
# These are all the subdirectories that are created for us. Your scripts go in `to_run`.
path_run,path_running,path_complete,path_fail,path_out = setup_dirs(path)
# Let's create a scripts directory with a couple of "scripts" (actually symlinks for this demo) in it.
def _setup_test_env():
shutil.rmtree('data')
res = setup_dirs(path)
os.symlink(Path('test_scripts/script_succ.sh').absolute(), path_run/'script_succ.sh')
os.symlink(Path('test_scripts/script_fail.sh').absolute(), path_run/'script_fail.sh')
(path_run/'test_dir').mkdir(exist_ok=True)
_setup_test_env()
# ## Helper functions for scripts
# These functions are used to find and run scripts, and move scripts to the appropriate subdirectory at the appropriate time.
#export
def find_next_script(p):
"Get the first script from `p` (in sorted order)"
files = p.ls().sorted().filter(Self.is_file())
if files: return files[0]
test_eq(find_next_script(path_run).name, 'script_fail.sh')
assert not find_next_script(path_complete)
#export
def safe_rename(file, dest):
"Move `file` to `dest`, prefixing a random uuid if there's a name conflict"
to_name = dest/file.name
if to_name.exists():
u = uuid4()
to_name = dest/f'{name}-{u}'
warnings.warn(f'Using unique name {to_name}')
file.replace(to_name)
return to_name
# ## ResourcePoolBase -
#export
class ResourcePoolBase():
def __init__(self, path):
self.path = Path(path)
setup_dirs(self.path)
def _lockpath(self,ident): return self.path/f'{ident}.lock'
def _is_locked(self,ident): return self._lockpath(ident).exists()
def lock(self,ident, txt='locked'): self._lockpath(ident).write_text(str(txt))
def unlock(self,ident): return self._lockpath(ident).unlink() if self._is_locked(ident) else None
def is_available(self,ident): return not self._is_locked(ident)
def all_ids(self): raise NotImplementedError
def find_next(self, n=1):
selected = []
for i in range(n):
ident = first(o for o in self.all_ids() if ((not o in selected) and self.is_available(o)))
selected.append(ident)
if None in selected:
return None
else:
return selected
def lock_next(self, n=1):
idents = self.find_next(n)
if idents is None: return
for ident in idents:
self.lock(ident)
return idents
def _launch(self, script, idents, env):
with (self.path/'out'/f'{script.name}.stderr').open("w") as stderr:
with (self.path/'out'/f'{script.name}.stdout').open("w") as stdout:
process = subprocess.Popen(str(script), env=env, stdout=stdout, stderr=stderr)
for ident in idents:
self.lock(ident, process.pid)
return process.wait()
def _run(self, script, idents):
failed = False
env=copy(os.environ)
env['FASTGPU_ID'] = str(idents)
try: res = self._launch(script, idents, env=env)
except Exception as e: failed = str(e)
(self.path/'out'/f'{script.name}.exitcode').write_text(failed if failed else str(res))
dest = self.path/'fail' if failed or res else self.path/'complete'
finish_name = safe_rename(script, dest)
for ident in idents:
self.unlock(ident)
def run(self, *args, **kwargs):
thread = Thread(target=self._run, args=args, kwargs=kwargs)
thread.start()
def poll_scripts(self, poll_interval=0.1, exit_when_empty=True, num_workers=1):
while True:
sleep(poll_interval)
script = find_next_script(self.path/'to_run')
if script is None:
if exit_when_empty: break
else: continue
idents = self.lock_next(num_workers)
if idents is None: continue
run_name = safe_rename(script, self.path/'running')
self.run(run_name, idents)
#export
add_docs(ResourcePoolBase, "Base class for locked access to list of idents",
unlock="Remove lockfile for `ident`",
lock="Create lockfile for `ident`",
is_available="Is `ident` available",
all_ids="All idents (abstract method)",
find_next="Finds next available resource, or None",
lock_next="Locks an available resource and returns its ident, or None",
run="Run `script` using resource `ident`",
poll_scripts="Poll `to_run` for scripts and run in parallel on available resources")
# This abstract class locks and unlocks resources using lockfiles. Override `all_ids` to make the list of resources available. See `FixedWorkerPool` for a simple example and details on each method.
#export
class FixedWorkerPool(ResourcePoolBase):
"Vends locked access to fixed list of idents"
def __init__(self, worker_ids, path):
super().__init__(path)
self.worker_ids = worker_ids
def all_ids(self):
"All available idents"
return self.worker_ids
# The simplest possible `ResourcePoolBase` subclass - the resources are just a list of ids. For instance:
_setup_test_env()
wp = FixedWorkerPool(L.range(4), path)
show_doc(FixedWorkerPool.unlock)
# If there are no locks, this does nothing:
wp.unlock(0)
show_doc(FixedWorkerPool.find_next)
# Initially all resources are available (unlocked), so the first from the provided list will be returned:
test_eq(wp.find_next(), [0])
show_doc(FixedWorkerPool.lock)
# After locking the first resource, it is no longer returned next:
wp.lock(0)
test_eq(wp.find_next(), [1])
show_doc(FixedWorkerPool.lock_next)
# This is the normal way to access a resource - it simply combines `find_next` and `lock`:
wp.lock_next()
test_eq(wp.find_next(), [2])
show_doc(FixedWorkerPool.run)
_setup_test_env()
wp = FixedWorkerPool(L.range(4), path)
# +
_setup_test_env()
f = find_next_script(path_run)
wp._run(f, [0])
test_eq(find_next_script(path_run), path_run/'script_succ.sh')
test_eq((path_out/'script_fail.sh.exitcode').read_text(), '1')
assert (path_fail/'script_fail.sh').exists()
# -
show_doc(FixedWorkerPool.poll_scripts)
# +
_setup_test_env()
wp.poll_scripts()
assert not find_next_script(path_run), find_next_script(path_run)
test_eq((path_out/'script_fail.sh.exitcode').read_text(), '1')
test_eq((path_out/'script_succ.sh.exitcode').read_text(), '0')
assert not (path_run/'script_fail.sh').exists()
assert (path_fail/'script_fail.sh').exists()
assert (path_complete/'script_succ.sh').exists()
test_eq((path_out/'script_succ.sh.stdout').read_text(), '[0]\n')
# -
# ## GPU
#export
class ResourcePoolGPU(ResourcePoolBase):
"Vends locked access to NVIDIA GPUs"
def __init__(self, path):
super().__init__(path)
nvmlInit()
# NVML doesn't respect CUDA_VISIBLE_DEVICES, so we need to query manually
envvar = os.environ.get("CUDA_VISIBLE_DEVICES", '')
self.devs = (L(envvar.split(',')).map(int) if envvar!=''
else L.range(nvmlDeviceGetCount()))
self.ids = L.range(self.devs)
def _launch(self, script, ident, env):
env['CUDA_VISIBLE_DEVICES'] = str(self.devs[ident])[1:-1]
return super()._launch(script, ident, env)
# with (self.path/'out'/f'{script.name}.stderr').open("w") as stderr:
# with (self.path/'out'/f'{script.name}.stdout').open("w") as stdout:
# process = subprocess.Popen(str(script), env=env, stdout=stdout, stderr=stderr)
# self.lock(ident, process.pid)
# return process.wait()
# with (self.path/'out'/f'{script.name}.stderr').open("w") as stderr:
# with (self.path/'out'/f'{script.name}.stdout').open("w") as stdout:
# return subprocess.call(str(script), env=env, stdout=stdout, stderr=stderr)
def is_available(self,ident):
"If a GPU's used_memory is less than 1G and is running no procs then it will be regarded as available"
if not super().is_available(ident): return False
device = nvmlDeviceGetHandleByIndex(self.devs[ident])
if nvmlDeviceGetComputeRunningProcesses(device): return False
return nvmlDeviceGetMemoryInfo(device).used <= 1e9
def all_ids(self):
"All GPUs"
return self.ids
# +
# These only work if you have an NVIDIA GPU installed
# wp = ResourcePoolGPU('data')
# wp.find_next()
# -
# This is a resource pool that uses [pynvml](https://pypi.org/project/pynvml/) to find GPUs that aren't being used (based on whether they have memory allocated). It is implemented by overriding two methods from `ResourcePoolBase`. Usage is identical to `FixedWorkerPool`, except that you don't need to pass in `worker_ids`, since available GPUs are considered to be the resource pool.
show_doc(ResourcePoolGPU.is_available)
show_doc(ResourcePoolGPU.all_ids)
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| 00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import json
import os
import pandas as pd
from datetime import datetime
from dotenv import load_dotenv, find_dotenv
from pathlib import Path
from src.func import tweet_utils
from src.func import regex
from src.func import labmtgen
from src.sentiment.senti_utils import *
from labMTsimple.storyLab import *
import random
import numpy as np
from collections import Counter
import matplotlib.pyplot as plt
tweet_dir= Path("../data/processed/tweets/")
city = Path("../data/processed/tweets/CA_San_Diego_0666000.json")
# +
lang = 'english'
labMT,labMTvector,labMTwordList = emotionFileReader(stopval=1.0,lang=lang,returnVector=True)
# Find Most influential words using labmt shift method
for city in tweet_dir.glob("*.json"):
city_name = city.stem
if city_name not in old_city:
print("{}".format(city_name))
PVec, CVec,pStops = getVecs(city)
PVec, Pval = labmtSimpleSentiment(PVec,pStops)
CVec, Cval = labmtSimpleSentiment(CVec,pStops)
sortedMag,sortedWords,sortedType,sumTypes = shift(CVec, PVec,labMTvector, labMTwordList,sort=True)
city_words = []
for i in range(50):
word = sortedWords[i]
entry = {"word":word, "valence":labMTvector[labMTwordList.index(word)], "contribution":sortedMag[i]}
city_words.append(entry)
city_df = pd.DataFrame.from_records(city_words, index='word')
with pd.ExcelWriter('./results/city_words.xlsx', mode='a') as writer:
city_df.to_excel(writer, sheet_name=city.stem)
# -
xls = pd.ExcelFile('./results/city_words.xlsx')
sheets = pd.read_excel(xls, None)
words_investigate = {}
for city, df in sheets.items():
words_investigate[city] = list(df[df.remove ==1.0].word.values)
words_investigate
| notebooks/process/process_word_magnitude.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 在一棵无限的二叉树上,每个节点都有两个子节点,树中的节点 逐行 依次按 “之” 字形进行标记。
# 如下图所示,在奇数行(即,第一行、第三行、第五行……)中,按从左到右的顺序进行标记;
# 而偶数行(即,第二行、第四行、第六行……)中,按从右到左的顺序进行标记。
# 给你树上某一个节点的标号 label,请你返回从根节点到该标号为 label 节点的路径,该路径是由途经的节点标号所组成的。
#
# 示例 1:
# 输入:label = 14
# 输出:[1,3,4,14]
#
# 示例 2:
# 输入:label = 26
# 输出:[1,2,6,10,26]
#
# 提示:
# 1、1 <= label <= 10^6
# -
# <img src='https://assets.leetcode-cn.com/aliyun-lc-upload/uploads/2019/06/28/tree.png'>
| Tree/1022/1104. Path In Zigzag Labelled Binary Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing Libraries and Getting Help
# ===
# ------
# ## Most of the power of a programming language is in its libraries.
#
# * A *library* is a collection of specialized functions that can be used in analysis.
# * E.g. the "math" Library has functions like `log` that can compute logarithms
#
#
# * Many additional libraries are available that for a whole variety of applications.
#
#
# ## To use a library, you have to import it at the beginning of your script
#
# * Use `import` to load a library into a script.
# * * Function and values from the Library can be called with a "`.`"
# * Python uses "`.`" to mean "part of".
#
# +
import math
print('pi is', math.pi) # pi is a number available from the math library
print('cos(pi) is', math.cos(math.pi)) # cos() is a function from the math library
# -
# ## Import specific items from a library to shorten programs.
#
# * Use `from...import...` to load only specific items from a library.
# * Then refer to them directly without library name as prefix.
# +
from math import pi, cos
print('pi is', pi)
print('cos(pi) is', cos(pi))
# -
# From the `os` library, import the path class
from os import path
# ## Create an alias for a library when importing it to shorten programs.
#
# * Use `import...as...` to give a library a short *alias* while importing it.
# * Then refer to items in the library using that shortened name.
# +
import math as m
print('cos(pi) is', m.cos(m.pi))
# -
# * Commonly used for libraries that are frequently used or have long names.
# * E.g., `matplotlib` plotting library is often aliased as `mpl`.
# * But can make programs harder to understand,
# since readers must learn your program's aliases.
# ---
# ## EXERCISE:
# Given the following section of code...
# ~~~
# import math as m
# angle = ____.degrees(____.pi / 2)
# print(____)
# ~~~
# 1. Fill in the blanks so that the program below prints `90.0`.
# 2. Rewrite the program so that it uses `import` *without* `as`.
# 3. Which form do you find easier to read?
#
# ---
# +
import math as m
angle = m.degrees(m.pi / 2)
print (angle)
# convert from degrees to radians and pass to cos()
print(m.cos(m.radians(angle)))
# -
import math
angle = math.degrees(math.pi / 2)
print (angle)
# ## Use `help` to find out more about a library's contents.
#
# * Works just like help for a function.
help(math)
# ---
# ## EXERCISE:
# When a colleague of yours types `help(math)`,
# Python reports an error:
# ~~~
# NameError: name 'math' is not defined
# ~~~
# 1. What has your colleague forgotten to do?
import math
# ---
# ## EXERCISE:
# 1. What function from the `math` library can you use to calculate a square root?
#
# ---
# ## Add documentation to your own code
#
# You can document your own code to help make it useable and readable by other people.
#
# We've been use `#` to add comments to specific lines in our code, but it's also easy to document your functions to make them available to pythons `help`
#
# Immediately after defining the function add a documenation block with triple-quotes: either single (`'''`) or double (`"""`)
# +
# An adding function
def adding_fifty(num):
"""
This is function takes a given number and adds 50 to it.
"""
return num + 50
help(adding_fifty)
# -
# ---
# ## EXERCISE:
# You want to select a random value from your data:
# ~~~
# ids = [1, 2, 3, 4, 5, 6]
# ~~~
# 1. What standard library would you most expect to help?
# 2. Which function would you select from that library?
#
# ---
import random
help(random)
# ---
# ## EXERCISE:
# Jigsaw Puzzle (Parson’s Problem) Programming Example:
#
# Rearrange the following statements so that a random DNA base is printed. Not all statements may be needed. Feel free to use/add intermediate variables.
# ~~~
# dna = "ACTG"
# length = 10
# print(bases)
# bases += random.choice(dna)
# for count in range(length):
#
# bases = ""
# import random
# ~~~
import random
length = 10
dna = "ACTG"
bases = ""
for count in range(length):
bases += random.choice(dna)
print(bases)
help(random.choice)
# # -- COMMIT YOUR WORK TO GITHUB --
# ---
# ## Key Points
# * Most of the power of a programming language is in its libraries.
# * A program must import a library module in order to use it.
# * Use help to learn about the contents of a library module.
# * Import specific items from a library to shorten programs.
# * Create an alias for a library when importing it to shorten programs.
| python-lessons/03 - Importing Libraries and Getting Help.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
from sklearn.linear_model import Ridge
from sklearn.linear_model import RidgeCV
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
# -
# Load training data CSV using Pandas
filename = 'train.csv'
data = pd.read_csv(filename, header=0)
# Separate training data into Y and X
array = data.values
Y = array[:, 1]
X = array[:, 2:12]
# +
# Set up cross validation, grid search, model
lambda_values = np.array([0.1, 1, 10, 100, 1000])
# solvers = ['svd', 'cholesky', 'sparse_cg', 'lsqr', 'sag', 'saga']
# param_grid = dict(alpha=lambda_values, solver=solvers)
param_grid = dict(alpha=lambda_values)
k_folds = 10
seed = 3
scoring = 'neg_mean_squared_error'
model = Ridge(fit_intercept=False, max_iter=1e6, tol=1e-5, random_state=seed)
grid = GridSearchCV(estimator=model, param_grid=param_grid, scoring=scoring,
n_jobs=-1, cv=k_folds, verbose=1, return_train_score=True)
# -
# Run grid search on lambdas with cross validation using Ridge() model
grid.fit(X, Y)
# Print all results for inspection
print(grid.cv_results_)
# Extract mean scores for result w/ (convoluted) way to get RMSE out of sklearn
scores = np.sqrt(-grid.cv_results_['mean_test_score'])
result = pd.DataFrame(scores)
# +
# Alternatively, i.s.o. GridSearchCV, use cross_val_score to obtain same result
scores2 = []
for l in lambda_values:
model = Ridge(alpha=l, fit_intercept=False, random_state=seed)
scores2.append(np.sqrt(-np.mean(cross_val_score(model, X, Y, scoring=scoring,
cv=k_folds, n_jobs=-1))))
print(scores-scores2)
result2 = pd.DataFrame(scores2)
# -
# Alternatively, use KFold
def KFold_on_Ridge(n_splits=10, random_state=42, shuffle=False):
kf = KFold(n_splits=n_splits, random_state=random_state, shuffle=shuffle)
scores = []
for l in lambda_values:
model = Ridge(alpha=l, fit_intercept=False, random_state=seed)
error = []
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = Y[train_index], Y[test_index]
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
error.append(np.sqrt(mean_squared_error(y_test, y_pred)))
# print('Lambda: {}'.format(l), '10-fold mean RMSE: {:4f}'.format(np.mean(error)))
scores.append(np.mean(error))
return scores
scores3 = KFold_on_Ridge()
print(scores-scores3)
result3 = pd.DataFrame(scores3)
# And now run KFold several times w/ shuffle and take average
n_repeats = 500
multi_scores = np.ndarray(shape=(n_repeats, len(lambda_values)))
for i in range(n_repeats):
multi_scores[i] = KFold_on_Ridge(random_state=i, shuffle=True)
scores5 = np.mean(multi_scores, axis=0)
print(scores-scores5)
result5 = pd.DataFrame(scores5)
# Different route: I.s.o. all above, simply use CV-specialized model RidgeCV
model2 = RidgeCV(alphas=(0.1, 1, 10, 100, 1000), fit_intercept=False,
normalize=False, scoring=scoring, cv=None, store_cv_values=True)
model2.fit(X, Y)
scores4 = []
for j in range(5):
scores4.append(np.sqrt(mean_squared_error(Y, model2.cv_values_[:,j])))
print(scores-scores4)
result4 = pd.DataFrame(scores4)
# Final step for all options: Write chosen final result to output file
filename = 'result.csv'
final_result = result3
#final_result = result2
#final_result = result3
final_result.to_csv(filename, header=False, index=False)
| task1a_lm1d1z/task1a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Victoria, Australia
# **Source of original dataset:** https://discover.data.vic.gov.au/dataset/fatal-crashes-lives-lost-last-5-years-to-date
#
# **Location of accidents:** Latitude, Longitude
#
# **Date of accidents:** Date
#
# **Outcome of accidents:** Fatality
import pandas as pd
pd.set_option('max_columns', None)
pd.set_option('display.max_colwidth', -1)
import numpy as np
from plotly import graph_objects as go
import plotly.express as px
from itertools import chain
import matplotlib.pyplot as plt
import geopandas as gpd
# Setup input files
data_dir = "../data/victoria/"
geodata_file = data_dir + "Fatal_Crashes__Lives_Lost_Last_5_Years_to_Date.geojson"
# Read original data
data = gpd.read_file(geodata_file)
# Create Datetime column
data['Date'] = pd.to_datetime(data['ACCIDENT_DATE'])
# Setup bicycle filter
mask = (data["ROAD_USER_TYPE"] == 6)
# Setup latitude & longitude column
data['Longitude'] = data['geometry'].apply(lambda p: p.x)
data['Latitude'] = data['geometry'].apply(lambda p: p.y)
# Some key statistics
# Slice all bicycle accidents
data_bicycles = data.loc[mask].copy().reset_index()
data_bicycles.head()
# Save to file
print(data_bicycles.shape)
data_bicycles.to_csv('cycling_safety_victoria.csv')
print('Wrote file to: cycling_safety_victoria.csv')
| dataset_curation/victoria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Revised and fixed code from https://github.com/JayParks/transformer (MT) for LM
# Stripped the code from the JayParks repo for MT Transformer. Introduced a few updates and changes for speed, but it's still frustratingly slow. Possible improvement - speed it up.
#
# Another issue - hyperparameter search for language modelling (number of heads, number of self-attention layers, etc). Does not work well from the box. This might be of help https://arxiv.org/pdf/1804.00247.pdf.
#
# Also consider parallelizing.
# # TODO
# * Clean up
# * Add MoS
# # Random sequence length batching
#
# This version of Transformer LM usesrandom sequence length batching.
#
# **NB** Make sure the src code does not assuem the existence of PAD.
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]="1"
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from showprogress import showprogress
import torch
torch.cuda.device(0)
import torch.nn as nn
import torch.optim as optim
from torch.nn.utils.rnn import pack_padded_sequence as pack
from torch.nn.utils.rnn import pad_packed_sequence as pad
from torch.nn.utils import clip_grad_norm_ as clip
from torch.optim.lr_scheduler import StepLR
import const
from data import *
from transformermos import *
# -
d_model = 512
voc_size = 10000
batch_size = 12
seq_len = 74
n_experts = 10
a = torch.ones([batch_size, seq_len, d_model])
l1 = torch.nn.Linear(d_model, n_experts*d_model)
l2 = torch.nn.Linear(d_model, voc_size)
l3 = torch.nn.Linear(d_model, n_experts, bias=False)
output = a
latent = l1(output) # h [batch_size x seq_len x n_experts * d_model]
logit = l2(latent.view(-1, d_model)) # HW [batch_size * seq_len * n_experts x d_model]
print(logit.shape)
prior_logit = l3(output).contiguous().view(-1, n_experts)
prior = torch.nn.functional.softmax(prior_logit, dim=1) # pi
print(prior.shape)
prob = torch.nn.functional.softmax(logit.view(-1, voc_size), dim=1).view(-1, n_experts, voc_size) # exp(hw) / sum(exp(hw))
prob = (prob * prior.unsqueeze(2).expand_as(prob)).sum(1)
print(prob.shape)
prob.size(-1)
def logging(s, print_=True, log_=True):
if print_:
print(s)
# if log_:
# with open(os.path.join(args.save, 'log.txt'), 'a+') as f_log:
# f_log.write(s + '\n')
# +
# ptb_datapath_train = 'data/penn/train.txt'
# ptb_datapath_valid = 'data/penn/valid.txt'
# ptb_datapath_test = 'data/penn/test.txt'
# batch_size = 128
# ptb_train = DataSet(ptb_datapath_train, batch_size, display_freq=0, max_len=90, trunc_len=90)
# ptb_valid = DataSet(ptb_datapath_valid, batch_size, display_freq=0, max_len=90, trunc_len=90)
# ptb_test = DataSet(ptb_datapath_test, batch_size, display_freq=0, max_len=90, trunc_len=90)
# +
# ptb_train.build_dict()
# ptb_valid.change_dict(ptb_train.dictionary)
# ptb_test.change_dict(ptb_train.dictionary)
# +
############ Optional: get data by tokens ###############
corpus = Corpus('data/penn')
eval_batch_size = 10
test_batch_size = 1
batch_size = 12
train_data = batchify(corpus.train, batch_size, )
val_data = batchify(corpus.valid, eval_batch_size)
test_data = batchify(corpus.test, test_batch_size)
#### how to take a batch ####
# the data is already splitten into batch_size(now we need to decide about seq length)
# batch_num = 2
# batch = get_batch(train_data, batch_num, seq_len=35)
#### TODO (if needed) ###
# 1) repackage hiddens for learning by tokens
# 2) learn not every step (depends on 1st point)
# 3) add grad clipping
# -
val_data.device
voc_size = len(corpus.dictionary) #corpus.dictionary.total # ptb_train.num_vocb
n_tokens = voc_size
emb_dim = 512
d_k = 64
d_v = 64
n_layers = 2
n_heads = 4
d_ff = 2048
max_tgt_seq_len = 90
dropout = 0.1
weighted_model = False
share_proj_weight = True
lr = 1e-6
n_epochs = 1000
clip_grad = 5
warmup_steps = 2000
log_interval = 200
# +
model = LMTransformer(n_layers, d_k, d_v, emb_dim, d_ff,
n_heads, max_tgt_seq_len, voc_size,
dropout, weighted_model, share_proj_weight)
criterion = nn.CrossEntropyLoss(ignore_index=const.PAD)
if torch.cuda.is_available():
model = model.cuda()
criterion = criterion.cuda()
#opt = optim.Adam(model.trainable_params(), lr=lr)
# lr_lambda = lambda epoch: 0.99 ** epoch
#lrsched = StepLR(opt, step_size=10, gamma=0.5)
# -
bptt0 = 70
max_seq_len_delta = 40
# Evaluate model
# seq_len is strange parameter
def evaluate(data_source, model, ntokens, seq_len):
model.eval()
total_loss = 0
batch = 0
for i in range(0, data_source.size(0) - 1, seq_len):
data, targets = get_batch(data_source, i, seq_len=seq_len)
seq_len = data.shape[1]
lengths = torch.ones(data.shape[0], device=device, dtype=torch.long) * seq_len
log_prob, self_attn = model(data, lengths)
loss = criterion(log_prob, targets.view(-1))
total_loss += loss.item()
batch += 1
return total_loss / batch
# +
n_epochs = 500
# -
opt = optim.Adam(model.trainable_params(),betas=(0.9, 0.98), eps=1e-09, lr=lr, weight_decay=1e-5)
i=0
best_val_loss = []
# +
try:
for epoch in range(n_epochs):
epoch_start_time = time.time()
total_loss = 0
print('Start epoch %d, learning rate %f '%(epoch + 1, opt.state_dict()['param_groups'][0]['lr']))
start_time = time.time()
model.train()
batch, i = 0, 0
while i < train_data.size(0) - 2:
bptt = bptt0 if np.random.random() < 0.95 else bptt0 / 2.
# Prevent excessively small or negative sequence lengths
seq_len = max(5, int(np.random.normal(bptt, 5))) # loc 70, scale 5
# There's a very small chance that it could select a very long sequence length resulting in OOM
seq_len = min(seq_len, bptt + max_seq_len_delta)
data, targets = get_batch(train_data, i, seq_len=seq_len)
seq_len = data.shape[1]
lengths = torch.ones(data.shape[0], device=device, dtype=torch.long) * seq_len
opt.zero_grad()
output, self_attn = model.forward(data, lengths)
# break
# break
loss = criterion(output, targets.view(-1))
loss.backward()
opt.step()
batch += 1
i += seq_len
new_lr = np.power(emb_dim, -0.5) * np.min([
np.power((batch), -0.5),
np.power(warmup_steps, -1.5) * (batch)])
for param_group in opt.param_groups:
param_group['lr'] = new_lr
if batch % log_interval == 0 and batch > 0:
cur_loss = loss.item()
elapsed = time.time() - start_time
logging('| epoch {:3d} | {}/{} batches | lr {:02.4f} | ms/batch {:5.2f} | '
'loss {:5.2f} | ppl {:8.2f}'.format(
epoch, batch, len(train_data) // bptt0, opt.param_groups[0]['lr'],
elapsed * 1000 / log_interval, cur_loss, math.exp(cur_loss)))
total_loss = 0
start_time = time.time()
val_loss = evaluate(val_data, model, voc_size, bptt0)
logging('-' * 89)
logging('| end of epoch {:3d} | time: {:5.2f}s | valid loss {:5.2f} | '
'valid ppl {:8.2f}'.format(epoch, (time.time() - epoch_start_time),
val_loss, math.exp(val_loss)))
logging('-' * 89)
best_val_loss.append(val_loss)
except KeyboardInterrupt:
logging('-' * 89)
logging('Exiting from training early')
# -
| TransformerLM_mos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('src/')
import numpy as np
import torch, torch.nn
from library_function import library_1D
from neural_net import LinNetwork
from DeepMod import DeepMod
import matplotlib.pyplot as plt
plt.style.use('seaborn-notebook')
import torch.nn as nn
from torch.autograd import grad
from scipy.io import loadmat
# %load_ext autoreload
# %autoreload 2
# -
# # Preparing data
# +
np.random.seed(34)
number_of_samples = 1000
data = np.load('data/burgers.npy', allow_pickle=True).item()
X = np.transpose((data['x'].flatten(), data['t'].flatten()))
y = np.real(np.transpose((data['u'].flatten(),data['u'].flatten())))
# -
idx = np.random.permutation(y.shape[0])
X_train = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y[idx, :][:number_of_samples], dtype=torch.float32)
rawdata = loadmat('data/kinetics_new.mat')
raw = np.real(rawdata['Expression1'])
raw= raw.reshape((1901,3))
t = raw[:-1,0].reshape(-1,1)
X1= raw[:-1,1]
X2 = raw[:-1,2]
X = np.float32(t.reshape(-1,1))
y= np.vstack((X1,X2))
y = np.transpose(y)
number_of_samples = 1000
idx = np.random.permutation(y.shape[0])
X_train = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y[idx, :][:number_of_samples], dtype=torch.float32)
y_train.shape
# # Building network
optim_config ={'lambda':1e-6,'max_iteration':50000}
lib_config={'poly_order':1, 'diff_order':2, 'total_terms':4}
network_config={'input_dim':1, 'hidden_dim':20, 'layers':5, 'output_dim':2}
sparse_weight_vector, sparsity_pattern, prediction, network = DeepMod(X_train, y_train,network_config, lib_config, optim_config)
prediction = network(torch.tensor(X, dtype=torch.float32))
prediction = prediction.detach().numpy()
x, y = np.meshgrid(X[:,0], X[:,1])
mask = torch.tensor((0,1,3))
mask
sparse_coefs = torch.tensor((0.1,0.2,0.4)).reshape(-1,1)
sparse_coefs
dummy = torch.ones((5,3,1))
dummy2 = torch.ones((5,1,4))
(dummy @ dummy2).shape
dummy.shape
dummy.reshape(-1,3,1).shape
dummy = dummy.reshape(2,2)
torch.where(coefs(mask),coefs,dummy)
x = np.linspace(0, 1, 100)
X, Y = np.meshgrid(x, x)
Z = np.sin(X)*np.sin(Y)
b = torch.ones((10, 2), dtype=torch.float32, requires_grad=True)
a = torch.tensor(np.ones((2,10)), dtype=torch.float32)
test=torch.tensor([[0.3073, 0.4409],
[0.0212, 0.6602]])
torch.where(test>torch.tensor(0.3),test, torch.zeros_like(test))
# + active=""
# test2=torch.reshape(test, (1,4))
# -
test2[0,:].reshape(-1,1)
mask=torch.nonzero(test2[0,:])
mask=torch.reshape(torch.nonzero(test2), (1,4))
mask
test2[mask[1]]
a.shape[1]
| .ipynb_checkpoints/PyTorch_V4_multi_traj-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda2]
# language: python
# name: conda-env-anaconda2-py
# ---
# # Dataset creation for "Crystal-structure identification using Bayesian deep learning"
#
# Author: <NAME> (<EMAIL>; <EMAIL>)
#
#
# ### Brief summary
# This notebook shows how to create the datasets starting from the tar files which contain the descriptor.
#
# The tar files for different displacements and vacancies are extracted, and a dataset is created using the function *prepare_dataset* from *ai4materials.dataprocessing.preprocessing*; a dataset consists of a numpy array containing the diffraction intensity in spherical harmonics (DISH) descriptor, a numpy array containing the correct labels (the label of the pristine structure), and a *json* files providing a human-readable summary of the content of the dataset.
#
# Moreover, a ASE database containing structures for each defective transformation (displacements and vacancies) is written to disk using the *write_ase_db* function from *ai4materials.utils.utils_data_retrieval*.
# + code_folding=[0]
# load libraries
from ai4materials.dataprocessing.preprocessing import load_dataset_from_file
from ai4materials.wrappers import load_descriptor
from ai4materials.utils.utils_config import set_configs
from ai4materials.utils.utils_config import setup_logger
from ai4materials.dataprocessing.preprocessing import load_dataset_from_file
from ai4materials.dataprocessing.preprocessing import prepare_dataset
from ai4materials.utils.utils_data_retrieval import write_ase_db
from ase.spacegroup import get_spacegroup as ase_get_spacegroup
from collections import Counter
import itertools
import matplotlib.pyplot as plt
import numpy as np
import os
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
# %matplotlib inline
main_folder = '/home/ziletti/Documents/calc_nomadml/rot_inv_3d/'
dataset_folder = os.path.abspath(os.path.normpath(os.path.join(main_folder, 'datasets')))
desc_folder = os.path.abspath(os.path.normpath(os.path.join(main_folder, 'desc_folder')))
configs = set_configs(main_folder=main_folder)
logger = setup_logger(configs, level='ERROR', display_configs=False)
configs['io']['dataset_folder'] = dataset_folder
configs['io']['desc_folder'] = desc_folder
# -
# ## 1. Pristine structures
# + code_folding=[0]
# hcp - spacegroup 194
filenames_pristine_hcp = ['hcp/pristine/A_hP2_194_c_target_nb_atoms128_rotid0_pristine.tar.gz',
'hcp/pristine/A_hP2_194_c_target_nb_atoms128_rotid1_pristine.tar.gz',
'hcp/pristine/A_hP2_194_c_target_nb_atoms128_rotid2_pristine.tar.gz',
'hcp/pristine/A_hP2_194_c_target_nb_atoms128_rotid3_pristine.tar.gz',
'hcp/pristine/A_hP2_194_c_target_nb_atoms128_rotid4_pristine.tar.gz']
# sc - spacegroup 221
filenames_pristine_sc = ['sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid0_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid1_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid2_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid3_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid4_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid5_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid6_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid7_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid8_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid9_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid10_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid11_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid12_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid13_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid14_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid15_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid16_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid17_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid18_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid19_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid20_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid21_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid22_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid23_pristine.tar.gz',
'sc/pristine/A_cP1_221_a_target_nb_atoms128_rotid24_pristine.tar.gz']
# fcc - spacegroup 225
filenames_pristine_fcc = ['fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid0_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid1_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid2_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid3_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid4_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid5_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid6_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid7_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid8_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid9_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid10_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid11_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid12_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid13_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid14_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid15_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid16_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid17_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid18_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid19_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid20_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid21_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid22_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid23_pristine.tar.gz',
'fcc/pristine/A_cF4_225_a_target_nb_atoms128_rotid24_pristine.tar.gz']
# diam - spacegroup 227
filenames_pristine_diam = ['diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid0_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid1_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid2_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid3_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid4_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid5_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid6_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid7_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid8_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid9_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid10_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid11_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid12_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid13_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid14_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid15_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid16_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid17_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid18_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid19_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid20_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid21_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid22_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid23_pristine.tar.gz',
'diam/pristine/A_cF8_227_a_target_nb_atoms128_rotid24_pristine.tar.gz']
# bcc - spacegroup 229
filenames_pristine_bcc = ['bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid0_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid1_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid2_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid3_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid4_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid5_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid6_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid7_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid8_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid9_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid10_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid11_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid12_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid13_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid14_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid15_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid16_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid17_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid18_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid19_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid20_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid21_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid22_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid23_pristine.tar.gz',
'bcc/pristine/A_cI2_229_a_target_nb_atoms128_rotid24_pristine.tar.gz']
desc_files_pristine_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_pristine_hcp]
desc_files_pristine_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_pristine_sc]
desc_files_pristine_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_pristine_fcc]
desc_files_pristine_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_pristine_diam]
desc_files_pristine_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_pristine_bcc]
y_true = []
target_list_hcp_pristine, structure_list_hcp_pristine = load_descriptor(desc_files=desc_files_pristine_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_pristine)
target_list_sc_pristine, structure_list_sc_pristine = load_descriptor(desc_files=desc_files_pristine_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_pristine)
target_list_fcc_pristine, structure_list_fcc_pristine = load_descriptor(desc_files=desc_files_pristine_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_pristine)
target_list_diam_pristine, structure_list_diam_pristine = load_descriptor(desc_files=desc_files_pristine_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_pristine)
target_list_bcc_pristine, structure_list_bcc_pristine = load_descriptor(desc_files=desc_files_pristine_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_pristine)
# + code_folding=[0]
# load structures
structure_list_pristine = structure_list_hcp_pristine + structure_list_sc_pristine + structure_list_fcc_pristine + structure_list_diam_pristine + structure_list_bcc_pristine
target_list_pristine = target_list_hcp_pristine + target_list_sc_pristine + target_list_fcc_pristine + target_list_diam_pristine + target_list_bcc_pristine
for idx, item in enumerate(target_list_pristine):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_pristine):
structure.info['target'] = y_true[idx]
#y_pred_disp04 = []
#for structure in structure_list_04:
# y_pred_disp04.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0, 2]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_pristine,
target_list=target_list_pristine,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_pristine-large',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures pristine. 25 rotations for cubic, 5 for hcp.")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_pristine, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_pristine-large', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ## 2. Structures with displacements
# ### 2.1 Displacements 0.1%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp01_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp01.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp01.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp01.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp01.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp01.tar.gz']
# sc - spacegroup 221
filenames_disp01_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp01.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp01.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp01.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp01.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp01.tar.gz']
# fcc - spacegroup 225
filenames_disp01_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp01.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp01.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp01.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp01.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp01.tar.gz']
# diam - spacegroup 227
filenames_disp01_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp01.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp01.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp01.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp01.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp01.tar.gz']
# bcc - spacegroup 229
filenames_disp01_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp01.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp01.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp01.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp01.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp01.tar.gz']
desc_files_disp01_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp01_hcp]
desc_files_disp01_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp01_sc]
desc_files_disp01_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp01_fcc]
desc_files_disp01_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp01_diam]
desc_files_disp01_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp01_bcc]
y_true = []
target_list_hcp_disp01, structure_list_hcp_disp01 = load_descriptor(desc_files=desc_files_disp01_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp01)
target_list_sc_disp01, structure_list_sc_disp01 = load_descriptor(desc_files=desc_files_disp01_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp01)
target_list_fcc_disp01, structure_list_fcc_disp01 = load_descriptor(desc_files=desc_files_disp01_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp01)
target_list_diam_disp01, structure_list_diam_disp01 = load_descriptor(desc_files=desc_files_disp01_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp01)
target_list_bcc_disp01, structure_list_bcc_disp01 = load_descriptor(desc_files=desc_files_disp01_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp01)
# + code_folding=[0]
# read structures
structure_list_disp01 = structure_list_hcp_disp01 + structure_list_sc_disp01 + structure_list_fcc_disp01 + structure_list_diam_disp01 + structure_list_bcc_disp01
target_list_disp01 = target_list_hcp_disp01 + target_list_sc_disp01 + target_list_fcc_disp01 + target_list_diam_disp01 + target_list_bcc_disp01
for idx, item in enumerate(target_list_disp01):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp01):
structure.info['target'] = y_true[idx]
#y_pred_disp04 = []
#for structure in structure_list_04:
# y_pred_disp04.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp01,
target_list=target_list_disp01,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-0.1%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 0.1% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp01, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-0.1%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.2 Displacements 0.2%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp02_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp02.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp02.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp02.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp02.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp02.tar.gz']
# sc - spacegroup 221
filenames_disp02_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp02.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp02.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp02.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp02.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp02.tar.gz']
# fcc - spacegroup 225
filenames_disp02_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp02.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp02.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp02.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp02.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp02.tar.gz']
# diam - spacegroup 227
filenames_disp02_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp02.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp02.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp02.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp02.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp02.tar.gz']
# bcc - spacegroup 229
filenames_disp02_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp02.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp02.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp02.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp02.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp02.tar.gz']
desc_files_disp02_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp02_hcp]
desc_files_disp02_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp02_sc]
desc_files_disp02_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp02_fcc]
desc_files_disp02_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp02_diam]
desc_files_disp02_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp02_bcc]
y_true = []
target_list_hcp_disp02, structure_list_hcp_disp02 = load_descriptor(desc_files=desc_files_disp02_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp02)
target_list_sc_disp02, structure_list_sc_disp02 = load_descriptor(desc_files=desc_files_disp02_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp02)
target_list_fcc_disp02, structure_list_fcc_disp02 = load_descriptor(desc_files=desc_files_disp02_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp02)
target_list_diam_disp02, structure_list_diam_disp02 = load_descriptor(desc_files=desc_files_disp02_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp02)
target_list_bcc_disp02, structure_list_bcc_disp02 = load_descriptor(desc_files=desc_files_disp02_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp02)
# + code_folding=[0]
# read structures
structure_list_disp02 = structure_list_hcp_disp02 + structure_list_sc_disp02 + structure_list_fcc_disp02 + structure_list_diam_disp02 + structure_list_bcc_disp02
target_list_disp02 = target_list_hcp_disp02 + target_list_sc_disp02 + target_list_fcc_disp02 + target_list_diam_disp02 + target_list_bcc_disp02
for idx, item in enumerate(target_list_disp02):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp02):
structure.info['target'] = y_true[idx]
#y_pred_disp04 = []
#for structure in structure_list_04:
# y_pred_disp04.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp02,
target_list=target_list_disp02,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-0.2%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 0.2% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp02, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-0.2%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.3 Displacements 0.6%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp06_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp06.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp06.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp06.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp06.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp06.tar.gz']
# sc - spacegroup 221
filenames_disp06_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp06.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp06.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp06.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp06.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp06.tar.gz']
# fcc - spacegroup 225
filenames_disp06_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp06.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp06.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp06.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp06.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp06.tar.gz']
# diam - spacegroup 227
filenames_disp06_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp06.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp06.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp06.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp06.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp06.tar.gz']
# bcc - spacegroup 229
filenames_disp06_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp06.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp06.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp06.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp06.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp06.tar.gz']
desc_files_disp06_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp06_hcp]
desc_files_disp06_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp06_sc]
desc_files_disp06_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp06_fcc]
desc_files_disp06_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp06_diam]
desc_files_disp06_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp06_bcc]
y_true = []
target_list_hcp_disp06, structure_list_hcp_disp06 = load_descriptor(desc_files=desc_files_disp06_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp06)
target_list_sc_disp06, structure_list_sc_disp06 = load_descriptor(desc_files=desc_files_disp06_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp06)
target_list_fcc_disp06, structure_list_fcc_disp06 = load_descriptor(desc_files=desc_files_disp06_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp06)
target_list_diam_disp06, structure_list_diam_disp06 = load_descriptor(desc_files=desc_files_disp06_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp06)
target_list_bcc_disp06, structure_list_bcc_disp06 = load_descriptor(desc_files=desc_files_disp06_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp06)
# + code_folding=[0]
# read structures
structure_list_disp06 = structure_list_hcp_disp06 + structure_list_sc_disp06 + structure_list_fcc_disp06 + structure_list_diam_disp06 + structure_list_bcc_disp06
target_list_disp06 = target_list_hcp_disp06 + target_list_sc_disp06 + target_list_fcc_disp06 + target_list_diam_disp06 + target_list_bcc_disp06
for idx, item in enumerate(target_list_disp06):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp06):
structure.info['target'] = y_true[idx]
#y_pred_disp04 = []
#for structure in structure_list_04:
# y_pred_disp04.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp06,
target_list=target_list_disp06,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-0.6%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 0.6% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp06, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-0.6%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.4 Displacements 1.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp1_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp1.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp1.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp1.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp1.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp1.tar.gz']
# sc - spacegroup 221
filenames_disp1_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp1.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp1.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp1.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp1.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp1.tar.gz']
# fcc - spacegroup 225
filenames_disp1_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp1.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp1.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp1.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp1.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp1.tar.gz']
# diam - spacegroup 227
filenames_disp1_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp1.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp1.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp1.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp1.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp1.tar.gz']
# bcc - spacegroup 229
filenames_disp1_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp1.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp1.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp1.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp1.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp1.tar.gz']
desc_files_disp1_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp1_hcp]
desc_files_disp1_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp1_sc]
desc_files_disp1_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp1_fcc]
desc_files_disp1_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp1_diam]
desc_files_disp1_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp1_bcc]
y_true = []
target_list_hcp_disp1, structure_list_hcp_disp1 = load_descriptor(desc_files=desc_files_disp1_hcp, configs=configs)
y_true = y_true + [194]*len(target_list_hcp_disp1)
target_list_sc_disp1, structure_list_sc_disp1 = load_descriptor(desc_files=desc_files_disp1_sc, configs=configs)
y_true = y_true + [221]*len(target_list_sc_disp1)
target_list_fcc_disp1, structure_list_fcc_disp1 = load_descriptor(desc_files=desc_files_disp1_fcc, configs=configs)
y_true = y_true + [225]*len(target_list_fcc_disp1)
target_list_diam_disp1, structure_list_diam_disp1 = load_descriptor(desc_files=desc_files_disp1_diam, configs=configs)
y_true = y_true + [227]*len(target_list_diam_disp1)
target_list_bcc_disp1, structure_list_bcc_disp1 = load_descriptor(desc_files=desc_files_disp1_bcc, configs=configs)
y_true = y_true + [229]*len(target_list_bcc_disp1)
# + code_folding=[0]
# read structures
structure_list_disp1 = structure_list_hcp_disp1 + structure_list_sc_disp1 + structure_list_fcc_disp1 + structure_list_diam_disp1 + structure_list_bcc_disp1
target_list_disp1 = target_list_hcp_disp1 + target_list_sc_disp1 + target_list_fcc_disp1 + target_list_diam_disp1 + target_list_bcc_disp1
for idx, item in enumerate(target_list_disp1):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp1):
structure.info['target'] = y_true[idx]
#y_pred_disp1 = []
#for structure in structure_list_1:
# y_pred_disp1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp1,
target_list=target_list_disp1,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-1%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 1% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp1, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-1%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.5 Displacements 2.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp2_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp2.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp2.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp2.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp2.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp2.tar.gz']
# sc - spacegroup 221
filenames_disp2_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp2.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp2.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp2.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp2.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp2.tar.gz']
# fcc - spacegroup 225
filenames_disp2_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp2.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp2.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp2.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp2.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp2.tar.gz']
# diam - spacegroup 227
filenames_disp2_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp2.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp2.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp2.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp2.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp2.tar.gz']
# bcc - spacegroup 229
filenames_disp2_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp2.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp2.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp2.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp2.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp2.tar.gz']
desc_files_disp2_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp2_hcp]
desc_files_disp2_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp2_sc]
desc_files_disp2_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp2_fcc]
desc_files_disp2_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp2_diam]
desc_files_disp2_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp2_bcc]
y_true = []
target_list_hcp_disp2, structure_list_hcp_disp2 = load_descriptor(desc_files=desc_files_disp2_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp2)
target_list_sc_disp2, structure_list_sc_disp2 = load_descriptor(desc_files=desc_files_disp2_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp2)
target_list_fcc_disp2, structure_list_fcc_disp2 = load_descriptor(desc_files=desc_files_disp2_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp2)
target_list_diam_disp2, structure_list_diam_disp2 = load_descriptor(desc_files=desc_files_disp2_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp2)
target_list_bcc_disp2, structure_list_bcc_disp2 = load_descriptor(desc_files=desc_files_disp2_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp2)
# + code_folding=[0]
# read structures
structure_list_disp2 = structure_list_hcp_disp2 + structure_list_sc_disp2 + structure_list_fcc_disp2 + structure_list_diam_disp2 + structure_list_bcc_disp2
target_list_disp2 = target_list_hcp_disp2 + target_list_sc_disp2 + target_list_fcc_disp2 + target_list_diam_disp2 + target_list_bcc_disp2
for idx, item in enumerate(target_list_disp2):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp2):
structure.info['target'] = y_true[idx]
#y_pred_disp2 = []
#for structure in structure_list_2:
# y_pred_disp2.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp2,
target_list=target_list_disp2,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-2%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 2% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp2, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-2%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.6 Displacements 4.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp4_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp4.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp4.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp4.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp4.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp4.tar.gz']
# sc - spacegroup 221
filenames_disp4_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp4.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp4.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp4.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp4.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp4.tar.gz']
# fcc - spacegroup 225
filenames_disp4_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp4.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp4.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp4.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp4.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp4.tar.gz']
# diam - spacegroup 227
filenames_disp4_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp4.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp4.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp4.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp4.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp4.tar.gz']
# bcc - spacegroup 229
filenames_disp4_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp4.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp4.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp4.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp4.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp4.tar.gz']
desc_files_disp4_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp4_hcp]
desc_files_disp4_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp4_sc]
desc_files_disp4_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp4_fcc]
desc_files_disp4_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp4_diam]
desc_files_disp4_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp4_bcc]
y_true = []
target_list_hcp_disp4, structure_list_hcp_disp4 = load_descriptor(desc_files=desc_files_disp4_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp4)
target_list_sc_disp4, structure_list_sc_disp4 = load_descriptor(desc_files=desc_files_disp4_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp4)
target_list_fcc_disp4, structure_list_fcc_disp4 = load_descriptor(desc_files=desc_files_disp4_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp4)
target_list_diam_disp4, structure_list_diam_disp4 = load_descriptor(desc_files=desc_files_disp4_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp4)
target_list_bcc_disp4, structure_list_bcc_disp4 = load_descriptor(desc_files=desc_files_disp4_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp4)
# + code_folding=[0]
# read structures
structure_list_disp4 = structure_list_hcp_disp4 + structure_list_sc_disp4 + structure_list_fcc_disp4 + structure_list_diam_disp4 + structure_list_bcc_disp4
target_list_disp4 = target_list_hcp_disp4 + target_list_sc_disp4 + target_list_fcc_disp4 + target_list_diam_disp4 + target_list_bcc_disp4
for idx, item in enumerate(target_list_disp4):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp4):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp4,
target_list=target_list_disp4,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-4%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 4% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp4, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-4%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.7 Displacements 5.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp5_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp5.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp5.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp5.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp5.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp5.tar.gz']
# sc - spacegroup 221
filenames_disp5_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp5.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp5.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp5.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp5.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp5.tar.gz']
# fcc - spacegroup 225
filenames_disp5_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp5.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp5.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp5.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp5.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp5.tar.gz']
# diam - spacegroup 227
filenames_disp5_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp5.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp5.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp5.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp5.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp5.tar.gz']
# bcc - spacegroup 229
filenames_disp5_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp5.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp5.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp5.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp5.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp5.tar.gz']
desc_files_disp5_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp5_hcp]
desc_files_disp5_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp5_sc]
desc_files_disp5_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp5_fcc]
desc_files_disp5_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp5_diam]
desc_files_disp5_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp5_bcc]
y_true = []
target_list_hcp_disp5, structure_list_hcp_disp5 = load_descriptor(desc_files=desc_files_disp5_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp5)
target_list_sc_disp5, structure_list_sc_disp5 = load_descriptor(desc_files=desc_files_disp5_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp5)
target_list_fcc_disp5, structure_list_fcc_disp5 = load_descriptor(desc_files=desc_files_disp5_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp5)
target_list_diam_disp5, structure_list_diam_disp5 = load_descriptor(desc_files=desc_files_disp5_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp5)
target_list_bcc_disp5, structure_list_bcc_disp5 = load_descriptor(desc_files=desc_files_disp5_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp5)
# + code_folding=[0]
# read structures
structure_list_disp5 = structure_list_hcp_disp5 + structure_list_sc_disp5 + structure_list_fcc_disp5 + structure_list_diam_disp5 + structure_list_bcc_disp5
target_list_disp5 = target_list_hcp_disp5 + target_list_sc_disp5 + target_list_fcc_disp5 + target_list_diam_disp5 + target_list_bcc_disp5
for idx, item in enumerate(target_list_disp5):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp5):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp5,
target_list=target_list_disp5,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-5%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 5% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp5, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-5%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.8 Displacements 8.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp8_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp8.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp8.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp8.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp8.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp8.tar.gz']
# sc - spacegroup 221
filenames_disp8_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp8.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp8.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp8.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp8.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp8.tar.gz']
# fcc - spacegroup 225
filenames_disp8_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp8.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp8.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp8.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp8.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp8.tar.gz']
# diam - spacegroup 227
filenames_disp8_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp8.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp8.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp8.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp8.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp8.tar.gz']
# bcc - spacegroup 229
filenames_disp8_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp8.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp8.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp8.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp8.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp8.tar.gz']
desc_files_disp8_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp8_hcp]
desc_files_disp8_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp8_sc]
desc_files_disp8_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp8_fcc]
desc_files_disp8_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp8_diam]
desc_files_disp8_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp8_bcc]
y_true = []
target_list_hcp_disp8, structure_list_hcp_disp8 = load_descriptor(desc_files=desc_files_disp8_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp8)
target_list_sc_disp8, structure_list_sc_disp8 = load_descriptor(desc_files=desc_files_disp8_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp8)
target_list_fcc_disp8, structure_list_fcc_disp8 = load_descriptor(desc_files=desc_files_disp8_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp8)
target_list_diam_disp8, structure_list_diam_disp8 = load_descriptor(desc_files=desc_files_disp8_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp8)
target_list_bcc_disp8, structure_list_bcc_disp8 = load_descriptor(desc_files=desc_files_disp8_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp8)
# + code_folding=[0]
# read structures
structure_list_disp8 = structure_list_hcp_disp8 + structure_list_sc_disp8 + structure_list_fcc_disp8 + structure_list_diam_disp8 + structure_list_bcc_disp8
target_list_disp8 = target_list_hcp_disp8 + target_list_sc_disp8 + target_list_fcc_disp8 + target_list_diam_disp8 + target_list_bcc_disp8
for idx, item in enumerate(target_list_disp8):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp8):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp8,
target_list=target_list_disp8,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-8%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 8% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp8, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-8%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.9 Displacements 10.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp10_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp10.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp10.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp10.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp10.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp10.tar.gz']
# sc - spacegroup 221
filenames_disp10_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp10.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp10.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp10.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp10.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp10.tar.gz']
# fcc - spacegroup 225
filenames_disp10_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp10.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp10.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp10.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp10.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp10.tar.gz']
# diam - spacegroup 227
filenames_disp10_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp10.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp10.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp10.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp10.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp10.tar.gz']
# bcc - spacegroup 229
filenames_disp10_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp10.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp10.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp10.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp10.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp10.tar.gz']
desc_files_disp10_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp10_hcp]
desc_files_disp10_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp10_sc]
desc_files_disp10_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp10_fcc]
desc_files_disp10_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp10_diam]
desc_files_disp10_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp10_bcc]
y_true = []
target_list_hcp_disp10, structure_list_hcp_disp10 = load_descriptor(desc_files=desc_files_disp10_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp10)
target_list_sc_disp10, structure_list_sc_disp10 = load_descriptor(desc_files=desc_files_disp10_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp10)
target_list_fcc_disp10, structure_list_fcc_disp10 = load_descriptor(desc_files=desc_files_disp10_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp10)
target_list_diam_disp10, structure_list_diam_disp10 = load_descriptor(desc_files=desc_files_disp10_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp10)
target_list_bcc_disp10, structure_list_bcc_disp10 = load_descriptor(desc_files=desc_files_disp10_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp10)
# + code_folding=[0]
# read structures
structure_list_disp10 = structure_list_hcp_disp10 + structure_list_sc_disp10 + structure_list_fcc_disp10 + structure_list_diam_disp10 + structure_list_bcc_disp10
target_list_disp10 = target_list_hcp_disp10 + target_list_sc_disp10 + target_list_fcc_disp10 + target_list_diam_disp10 + target_list_bcc_disp10
for idx, item in enumerate(target_list_disp10):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp10):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp10,
target_list=target_list_disp10,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-10%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 10% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp10, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-10%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.10 Displacements 12.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp12_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp12.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp12.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp12.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp12.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp12.tar.gz']
# sc - spacegroup 221
filenames_disp12_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp12.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp12.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp12.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp12.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp12.tar.gz']
# fcc - spacegroup 225
filenames_disp12_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp12.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp12.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp12.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp12.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp12.tar.gz']
# diam - spacegroup 227
filenames_disp12_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp12.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp12.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp12.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp12.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp12.tar.gz']
# bcc - spacegroup 229
filenames_disp12_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp12.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp12.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp12.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp12.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp12.tar.gz']
desc_files_disp12_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp12_hcp]
desc_files_disp12_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp12_sc]
desc_files_disp12_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp12_fcc]
desc_files_disp12_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp12_diam]
desc_files_disp12_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp12_bcc]
y_true = []
target_list_hcp_disp12, structure_list_hcp_disp12 = load_descriptor(desc_files=desc_files_disp12_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp12)
target_list_sc_disp12, structure_list_sc_disp12 = load_descriptor(desc_files=desc_files_disp12_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp12)
target_list_fcc_disp12, structure_list_fcc_disp12 = load_descriptor(desc_files=desc_files_disp12_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp12)
target_list_diam_disp12, structure_list_diam_disp12 = load_descriptor(desc_files=desc_files_disp12_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp12)
target_list_bcc_disp12, structure_list_bcc_disp12 = load_descriptor(desc_files=desc_files_disp12_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp12)
# + code_folding=[0]
# read structures
structure_list_disp12 = structure_list_hcp_disp12 + structure_list_sc_disp12 + structure_list_fcc_disp12 + structure_list_diam_disp12 + structure_list_bcc_disp12
target_list_disp12 = target_list_hcp_disp12 + target_list_sc_disp12 + target_list_fcc_disp12 + target_list_diam_disp12 + target_list_bcc_disp12
for idx, item in enumerate(target_list_disp12):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp12):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp12,
target_list=target_list_disp12,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-12%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 12% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp12, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-12%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.11 Displacements 20.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp20_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp20.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp20.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp20.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp20.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp20.tar.gz']
# sc - spacegroup 221
filenames_disp20_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp20.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp20.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp20.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp20.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp20.tar.gz']
# fcc - spacegroup 225
filenames_disp20_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp20.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp20.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp20.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp20.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp20.tar.gz']
# diam - spacegroup 227
filenames_disp20_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp20.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp20.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp20.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp20.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp20.tar.gz']
# bcc - spacegroup 229
filenames_disp20_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp20.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp20.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp20.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp20.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp20.tar.gz']
desc_files_disp20_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp20_hcp]
desc_files_disp20_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp20_sc]
desc_files_disp20_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp20_fcc]
desc_files_disp20_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp20_diam]
desc_files_disp20_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp20_bcc]
y_true = []
target_list_hcp_disp20, structure_list_hcp_disp20 = load_descriptor(desc_files=desc_files_disp20_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp20)
target_list_sc_disp20, structure_list_sc_disp20 = load_descriptor(desc_files=desc_files_disp20_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp20)
target_list_fcc_disp20, structure_list_fcc_disp20 = load_descriptor(desc_files=desc_files_disp20_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp20)
target_list_diam_disp20, structure_list_diam_disp20 = load_descriptor(desc_files=desc_files_disp20_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp20)
target_list_bcc_disp20, structure_list_bcc_disp20 = load_descriptor(desc_files=desc_files_disp20_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp20)
# + code_folding=[0]
# read structures
structure_list_disp20 = structure_list_hcp_disp20 + structure_list_sc_disp20 + structure_list_fcc_disp20 + structure_list_diam_disp20 + structure_list_bcc_disp20
target_list_disp20 = target_list_hcp_disp20 + target_list_sc_disp20 + target_list_fcc_disp20 + target_list_diam_disp20 + target_list_bcc_disp20
for idx, item in enumerate(target_list_disp20):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp20):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp20,
target_list=target_list_disp20,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-20%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 20% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp20, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-20%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.12 Displacements 30.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp30_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp30.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp30.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp30.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp30.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp30.tar.gz']
# sc - spacegroup 221
filenames_disp30_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp30.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp30.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp30.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp30.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp30.tar.gz']
# fcc - spacegroup 225
filenames_disp30_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp30.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp30.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp30.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp30.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp30.tar.gz']
# diam - spacegroup 227
filenames_disp30_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp30.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp30.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp30.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp30.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp30.tar.gz']
# bcc - spacegroup 229
filenames_disp30_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp30.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp30.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp30.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp30.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp30.tar.gz']
desc_files_disp30_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp30_hcp]
desc_files_disp30_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp30_sc]
desc_files_disp30_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp30_fcc]
desc_files_disp30_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp30_diam]
desc_files_disp30_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp30_bcc]
y_true = []
target_list_hcp_disp30, structure_list_hcp_disp30 = load_descriptor(desc_files=desc_files_disp30_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp30)
target_list_sc_disp30, structure_list_sc_disp30 = load_descriptor(desc_files=desc_files_disp30_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp30)
target_list_fcc_disp30, structure_list_fcc_disp30 = load_descriptor(desc_files=desc_files_disp30_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp30)
target_list_diam_disp30, structure_list_diam_disp30 = load_descriptor(desc_files=desc_files_disp30_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp30)
target_list_bcc_disp30, structure_list_bcc_disp30 = load_descriptor(desc_files=desc_files_disp30_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp30)
# + code_folding=[0]
# read structures
structure_list_disp30 = structure_list_hcp_disp30 + structure_list_sc_disp30 + structure_list_fcc_disp30 + structure_list_diam_disp30 + structure_list_bcc_disp30
target_list_disp30 = target_list_hcp_disp30 + target_list_sc_disp30 + target_list_fcc_disp30 + target_list_diam_disp30 + target_list_bcc_disp30
for idx, item in enumerate(target_list_disp30):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp30):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp30,
target_list=target_list_disp30,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-30%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 30% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp30, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-30%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 2.13 Displacements 50.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_disp50_hcp = ['hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid0_disp50.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid1_disp50.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid2_disp50.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid3_disp50.tar.gz',
'hcp/disp/A_hP2_194_c_target_nb_atoms128_rotid4_disp50.tar.gz']
# sc - spacegroup 221
filenames_disp50_sc = ['sc/disp/A_cP1_221_a_target_nb_atoms128_rotid0_disp50.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid1_disp50.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid2_disp50.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid3_disp50.tar.gz',
'sc/disp/A_cP1_221_a_target_nb_atoms128_rotid4_disp50.tar.gz']
# fcc - spacegroup 225
filenames_disp50_fcc = ['fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid0_disp50.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid1_disp50.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid2_disp50.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid3_disp50.tar.gz',
'fcc/disp/A_cF4_225_a_target_nb_atoms128_rotid4_disp50.tar.gz']
# diam - spacegroup 227
filenames_disp50_diam = ['diam/disp/A_cF8_227_a_target_nb_atoms128_rotid0_disp50.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid1_disp50.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid2_disp50.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid3_disp50.tar.gz',
'diam/disp/A_cF8_227_a_target_nb_atoms128_rotid4_disp50.tar.gz']
# bcc - spacegroup 229
filenames_disp50_bcc = ['bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid0_disp50.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid1_disp50.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid2_disp50.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid3_disp50.tar.gz',
'bcc/disp/A_cI2_229_a_target_nb_atoms128_rotid4_disp50.tar.gz']
desc_files_disp50_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp50_hcp]
desc_files_disp50_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp50_sc]
desc_files_disp50_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp50_fcc]
desc_files_disp50_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp50_diam]
desc_files_disp50_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_disp50_bcc]
y_true = []
target_list_hcp_disp50, structure_list_hcp_disp50 = load_descriptor(desc_files=desc_files_disp50_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_disp50)
target_list_sc_disp50, structure_list_sc_disp50 = load_descriptor(desc_files=desc_files_disp50_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_disp50)
target_list_fcc_disp50, structure_list_fcc_disp50 = load_descriptor(desc_files=desc_files_disp50_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_disp50)
target_list_diam_disp50, structure_list_diam_disp50 = load_descriptor(desc_files=desc_files_disp50_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_disp50)
target_list_bcc_disp50, structure_list_bcc_disp50 = load_descriptor(desc_files=desc_files_disp50_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_disp50)
# + code_folding=[0]
# read structures
structure_list_disp50 = structure_list_hcp_disp50 + structure_list_sc_disp50 + structure_list_fcc_disp50 + structure_list_diam_disp50 + structure_list_bcc_disp50
target_list_disp50 = target_list_hcp_disp50 + target_list_sc_disp50 + target_list_fcc_disp50 + target_list_diam_disp50 + target_list_bcc_disp50
for idx, item in enumerate(target_list_disp50):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_disp50):
structure.info['target'] = y_true[idx]
#y_pred_disp4 = []
#for structure in structure_list_4:
# y_pred_disp4.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_disp50,
target_list=target_list_disp50,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_displacement-50%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 50% displacement")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_disp50, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_displacement-50%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ## 3. Structures with vacancies
# ### 3.1 Vacancies 1.0%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_vac1_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac01.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac01.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac01.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac01.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac01.tar.gz']
# sc - spacegroup 221
filenames_vac1_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac01.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac01.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac01.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac01.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac01.tar.gz']
# fcc - spacegroup 225
filenames_vac1_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac01.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac01.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac01.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac01.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac01.tar.gz']
# diam - spacegroup 227
filenames_vac1_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac01.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac01.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac01.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac01.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac01.tar.gz']
# bcc - spacegroup 229
filenames_vac1_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac01.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac01.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac01.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac01.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac01.tar.gz']
desc_files_vac1_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac1_hcp]
desc_files_vac1_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac1_sc]
desc_files_vac1_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac1_fcc]
desc_files_vac1_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac1_diam]
desc_files_vac1_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac1_bcc]
y_true = []
target_list_hcp_vac1, structure_list_hcp_vac1 = load_descriptor(desc_files=desc_files_vac1_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac1)
target_list_sc_vac1, structure_list_sc_vac1 = load_descriptor(desc_files=desc_files_vac1_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac1)
target_list_fcc_vac1, structure_list_fcc_vac1 = load_descriptor(desc_files=desc_files_vac1_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac1)
target_list_diam_vac1, structure_list_diam_vac1 = load_descriptor(desc_files=desc_files_vac1_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac1)
target_list_bcc_vac1, structure_list_bcc_vac1 = load_descriptor(desc_files=desc_files_vac1_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac1)
# + code_folding=[0]
# read structure list
structure_list_vac1 = structure_list_hcp_vac1 + structure_list_sc_vac1 + structure_list_fcc_vac1 + structure_list_diam_vac1 + structure_list_bcc_vac1
target_list_vac1 = target_list_hcp_vac1 + target_list_sc_vac1 + target_list_fcc_vac1 + target_list_diam_vac1 + target_list_bcc_vac1
for idx, item in enumerate(target_list_vac1):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac1):
structure.info['target'] = y_true[idx]
#y_pred_vac1 = []
#for structure in structure_list_vac1:
# y_pred_vac1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac1,
target_list=target_list_vac1,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-1%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 1% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac1, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-1%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.2 Vacancies 2%
# + code_folding=[0]
#hcp - spacegroup 194
filenames_vac2_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac02.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac02.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac02.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac02.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac02.tar.gz']
# sc - spacegroup 221
filenames_vac2_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac02.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac02.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac02.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac02.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac02.tar.gz']
# fcc - spacegroup 225
filenames_vac2_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac02.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac02.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac02.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac02.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac02.tar.gz']
# diam - spacegroup 227
filenames_vac2_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac02.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac02.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac02.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac02.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac02.tar.gz']
# bcc - spacegroup 229
filenames_vac2_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac02.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac02.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac02.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac02.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac02.tar.gz']
desc_files_vac2_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac2_hcp]
desc_files_vac2_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac2_sc]
desc_files_vac2_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac2_fcc]
desc_files_vac2_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac2_diam]
desc_files_vac2_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac2_bcc]
y_true = []
target_list_hcp_vac2, structure_list_hcp_vac2 = load_descriptor(desc_files=desc_files_vac2_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac2)
target_list_sc_vac2, structure_list_sc_vac2 = load_descriptor(desc_files=desc_files_vac2_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac2)
target_list_fcc_vac2, structure_list_fcc_vac2 = load_descriptor(desc_files=desc_files_vac2_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac2)
target_list_diam_vac2, structure_list_diam_vac2 = load_descriptor(desc_files=desc_files_vac2_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac2)
target_list_bcc_vac2, structure_list_bcc_vac2 = load_descriptor(desc_files=desc_files_vac2_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac2)
# + code_folding=[0]
# load structure list
structure_list_vac2 = structure_list_hcp_vac2 + structure_list_sc_vac2 + structure_list_fcc_vac2 + structure_list_diam_vac2 + structure_list_bcc_vac2
target_list_vac2 = target_list_hcp_vac2 + target_list_sc_vac2 + target_list_fcc_vac2 + target_list_diam_vac2 + target_list_bcc_vac2
for idx, item in enumerate(target_list_vac2):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac2):
structure.info['target'] = y_true[idx]
#y_pred_vac1 = []
#for structure in structure_list_vac1:
# y_pred_vac1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac2,
target_list=target_list_vac2,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-2%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 2% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac2, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-2%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.3 Vacancies 5%
# + code_folding=[0]
#hcp - spacegroup 194
filenames_vac5_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac05.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac05.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac05.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac05.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac05.tar.gz']
# sc - spacegroup 221
filenames_vac5_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac05.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac05.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac05.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac05.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac05.tar.gz']
# fcc - spacegroup 225
filenames_vac5_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac05.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac05.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac05.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac05.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac05.tar.gz']
# diam - spacegroup 227
filenames_vac5_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac05.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac05.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac05.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac05.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac05.tar.gz']
# bcc - spacegroup 229
filenames_vac5_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac05.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac05.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac05.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac05.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac05.tar.gz']
desc_files_vac5_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac5_hcp]
desc_files_vac5_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac5_sc]
desc_files_vac5_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac5_fcc]
desc_files_vac5_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac5_diam]
desc_files_vac5_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac5_bcc]
y_true = []
target_list_hcp_vac5, structure_list_hcp_vac5 = load_descriptor(desc_files=desc_files_vac5_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac5)
target_list_sc_vac5, structure_list_sc_vac5 = load_descriptor(desc_files=desc_files_vac5_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac5)
target_list_fcc_vac5, structure_list_fcc_vac5 = load_descriptor(desc_files=desc_files_vac5_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac5)
target_list_diam_vac5, structure_list_diam_vac5 = load_descriptor(desc_files=desc_files_vac5_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac5)
target_list_bcc_vac5, structure_list_bcc_vac5 = load_descriptor(desc_files=desc_files_vac5_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac5)
# + code_folding=[0]
# load structure list
structure_list_vac5 = structure_list_hcp_vac5 + structure_list_sc_vac5 + structure_list_fcc_vac5 + structure_list_diam_vac5 + structure_list_bcc_vac5
target_list_vac5 = target_list_hcp_vac5 + target_list_sc_vac5 + target_list_fcc_vac5 + target_list_diam_vac5 + target_list_bcc_vac5
for idx, item in enumerate(target_list_vac5):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac5):
structure.info['target'] = y_true[idx]
#y_pred_vac1 = []
#for structure in structure_list_vac1:
# y_pred_vac1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac5,
target_list=target_list_vac5,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-5%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 5% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac5, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-5%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.4 Vacancies 10%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_vac10_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac10.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac10.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac10.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac10.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac10.tar.gz']
# sc - spacegroup 221
filenames_vac10_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac10.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac10.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac10.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac10.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac10.tar.gz']
# fcc - spacegroup 225
filenames_vac10_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac10.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac10.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac10.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac10.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac10.tar.gz']
# diam - spacegroup 227
filenames_vac10_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac10.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac10.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac10.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac10.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac10.tar.gz']
# bcc - spacegroup 229
filenames_vac10_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac10.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac10.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac10.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac10.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac10.tar.gz']
desc_files_vac10_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac10_hcp]
desc_files_vac10_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac10_sc]
desc_files_vac10_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac10_fcc]
desc_files_vac10_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac10_diam]
desc_files_vac10_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac10_bcc]
y_true = []
target_list_hcp_vac10, structure_list_hcp_vac10 = load_descriptor(desc_files=desc_files_vac10_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac10)
target_list_sc_vac10, structure_list_sc_vac10 = load_descriptor(desc_files=desc_files_vac10_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac10)
target_list_fcc_vac10, structure_list_fcc_vac10 = load_descriptor(desc_files=desc_files_vac10_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac10)
target_list_diam_vac10, structure_list_diam_vac10 = load_descriptor(desc_files=desc_files_vac10_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac10)
target_list_bcc_vac10, structure_list_bcc_vac10 = load_descriptor(desc_files=desc_files_vac10_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac10)
# + code_folding=[0]
# load structure list
structure_list_vac10 = structure_list_hcp_vac10 + structure_list_sc_vac10 + structure_list_fcc_vac10 + structure_list_diam_vac10 + structure_list_bcc_vac10
target_list_vac10 = target_list_hcp_vac10 + target_list_sc_vac10 + target_list_fcc_vac10 + target_list_diam_vac10 + target_list_bcc_vac10
for idx, item in enumerate(target_list_vac10):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac10):
structure.info['target'] = y_true[idx]
#y_pred_vac1 = []
#for structure in structure_list_vac1:
# y_pred_vac1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac10,
target_list=target_list_vac10,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-10%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 10% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac10, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-10%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.5 Vacancies 20%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_vac20_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac20.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac20.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac20.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac20.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac20.tar.gz']
# sc - spacegroup 221
filenames_vac20_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac20.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac20.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac20.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac20.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac20.tar.gz']
# fcc - spacegroup 225
filenames_vac20_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac20.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac20.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac20.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac20.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac20.tar.gz']
# diam - spacegroup 227
filenames_vac20_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac20.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac20.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac20.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac20.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac20.tar.gz']
# bcc - spacegroup 229
filenames_vac20_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac20.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac20.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac20.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac20.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac20.tar.gz']
desc_files_vac20_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac20_hcp]
desc_files_vac20_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac20_sc]
desc_files_vac20_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac20_fcc]
desc_files_vac20_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac20_diam]
desc_files_vac20_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac20_bcc]
y_true = []
target_list_hcp_vac20, structure_list_hcp_vac20 = load_descriptor(desc_files=desc_files_vac20_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac20)
target_list_sc_vac20, structure_list_sc_vac20 = load_descriptor(desc_files=desc_files_vac20_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac20)
target_list_fcc_vac20, structure_list_fcc_vac20 = load_descriptor(desc_files=desc_files_vac20_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac20)
target_list_diam_vac20, structure_list_diam_vac20 = load_descriptor(desc_files=desc_files_vac20_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac20)
target_list_bcc_vac20, structure_list_bcc_vac20 = load_descriptor(desc_files=desc_files_vac20_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac20)
# + code_folding=[0]
# load structure list
structure_list_vac20 = structure_list_hcp_vac20 + structure_list_sc_vac20 + structure_list_fcc_vac20 + structure_list_diam_vac20 + structure_list_bcc_vac20
target_list_vac20 = target_list_hcp_vac20 + target_list_sc_vac20 + target_list_fcc_vac20 + target_list_diam_vac20 + target_list_bcc_vac20
for idx, item in enumerate(target_list_vac20):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac20):
structure.info['target'] = y_true[idx]
#y_pred_vac1 = []
#for structure in structure_list_vac1:
# y_pred_vac1.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac20,
target_list=target_list_vac20,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-20%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 20% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac20, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-20%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.6 Vacancies 25%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_vac25_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac25.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac25.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac25.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac25.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac25.tar.gz']
# sc - spacegroup 221
filenames_vac25_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac25.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac25.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac25.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac25.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac25.tar.gz']
# fcc - spacegroup 225
filenames_vac25_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac25.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac25.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac25.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac25.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac25.tar.gz']
# diam - spacegroup 227
filenames_vac25_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac25.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac25.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac25.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac25.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac25.tar.gz']
# bcc - spacegroup 229
filenames_vac25_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac25.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac25.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac25.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac25.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac25.tar.gz']
desc_files_vac25_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac25_hcp]
desc_files_vac25_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac25_sc]
desc_files_vac25_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac25_fcc]
desc_files_vac25_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac25_diam]
desc_files_vac25_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac25_bcc]
y_true = []
target_list_hcp_vac25, structure_list_hcp_vac25 = load_descriptor(desc_files=desc_files_vac25_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac25)
target_list_sc_vac25, structure_list_sc_vac25 = load_descriptor(desc_files=desc_files_vac25_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac25)
target_list_fcc_vac25, structure_list_fcc_vac25 = load_descriptor(desc_files=desc_files_vac25_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac25)
target_list_diam_vac25, structure_list_diam_vac25 = load_descriptor(desc_files=desc_files_vac25_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac25)
target_list_bcc_vac25, structure_list_bcc_vac25 = load_descriptor(desc_files=desc_files_vac25_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac25)
# + code_folding=[0]
# read structure lists
structure_list_vac25 = structure_list_hcp_vac25 + structure_list_sc_vac25 + structure_list_fcc_vac25 + structure_list_diam_vac25 + structure_list_bcc_vac25
target_list_vac25 = target_list_hcp_vac25 + target_list_sc_vac25 + target_list_fcc_vac25 + target_list_diam_vac25 + target_list_bcc_vac25
for idx, item in enumerate(target_list_vac25):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac25):
structure.info['target'] = y_true[idx]
#y_pred_vac25 = []
#for structure in structure_list_vac25:
# y_pred_vac25.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac25,
target_list=target_list_vac25,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-25%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 25% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac25, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-25%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
# ### 3.7 Vacancies 50%
# + code_folding=[0]
# hcp - spacegroup 194
filenames_vac50_hcp = ['hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid0_vac50.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid1_vac50.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid2_vac50.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid3_vac50.tar.gz',
'hcp/vac/A_hP2_194_c_target_nb_atoms128_rotid4_vac50.tar.gz']
# sc - spacegroup 221
filenames_vac50_sc = ['sc/vac/A_cP1_221_a_target_nb_atoms128_rotid0_vac50.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid1_vac50.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid2_vac50.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid3_vac50.tar.gz',
'sc/vac/A_cP1_221_a_target_nb_atoms128_rotid4_vac50.tar.gz']
# fcc - spacegroup 225
filenames_vac50_fcc = ['fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid0_vac50.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid1_vac50.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid2_vac50.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid3_vac50.tar.gz',
'fcc/vac/A_cF4_225_a_target_nb_atoms128_rotid4_vac50.tar.gz']
# diam - spacegroup 227
filenames_vac50_diam = ['diam/vac/A_cF8_227_a_target_nb_atoms128_rotid0_vac50.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid1_vac50.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid2_vac50.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid3_vac50.tar.gz',
'diam/vac/A_cF8_227_a_target_nb_atoms128_rotid4_vac50.tar.gz']
# bcc - spacegroup 229
filenames_vac50_bcc = ['bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid0_vac50.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid1_vac50.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid2_vac50.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid3_vac50.tar.gz',
'bcc/vac/A_cI2_229_a_target_nb_atoms128_rotid4_vac50.tar.gz']
desc_files_vac50_hcp = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac50_hcp]
desc_files_vac50_sc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac50_sc]
desc_files_vac50_fcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac50_fcc]
desc_files_vac50_diam = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac50_diam]
desc_files_vac50_bcc = [os.path.join(configs['io']['desc_folder'], item) for item in filenames_vac50_bcc]
y_true = []
target_list_hcp_vac50, structure_list_hcp_vac50 = load_descriptor(desc_files=desc_files_vac50_hcp, configs=configs)
y_true = y_true + [194]*len(structure_list_hcp_vac50)
target_list_sc_vac50, structure_list_sc_vac50 = load_descriptor(desc_files=desc_files_vac50_sc, configs=configs)
y_true = y_true + [221]*len(structure_list_sc_vac50)
target_list_fcc_vac50, structure_list_fcc_vac50 = load_descriptor(desc_files=desc_files_vac50_fcc, configs=configs)
y_true = y_true + [225]*len(structure_list_fcc_vac50)
target_list_diam_vac50, structure_list_diam_vac50 = load_descriptor(desc_files=desc_files_vac50_diam, configs=configs)
y_true = y_true + [227]*len(structure_list_diam_vac50)
target_list_bcc_vac50, structure_list_bcc_vac50 = load_descriptor(desc_files=desc_files_vac50_bcc, configs=configs)
y_true = y_true + [229]*len(structure_list_bcc_vac50)
# + code_folding=[0]
# read structures
structure_list_vac50 = structure_list_hcp_vac50 + structure_list_sc_vac50 + structure_list_fcc_vac50 + structure_list_diam_vac50 + structure_list_bcc_vac50
target_list_vac50 = target_list_hcp_vac50 + target_list_sc_vac50 + target_list_fcc_vac50 + target_list_diam_vac50 + target_list_bcc_vac50
for idx, item in enumerate(target_list_vac50):
item['data'][0]['target'] = y_true[idx]
for idx, structure in enumerate(structure_list_vac50):
structure.info['target'] = y_true[idx]
#y_pred_vac50 = []
#for structure in structure_list_vac25:
# y_pred_vac50.append(ase_get_spacegroup(structure, symprec=1e-1).no)
# + code_folding=[0]
# make dataset
path_to_x, path_to_y, path_to_summary = prepare_dataset(
structure_list=structure_list_vac50,
target_list=target_list_vac50,
desc_metadata='diffraction_3d_sh_spectrum',
dataset_name='hcp-sc-fcc-diam-bcc_vacancies-50%',
target_name='target',
target_categorical=True,
input_dims=(50, 32),
configs=configs,
dataset_folder=dataset_folder,
main_folder=configs['io']['main_folder'],
desc_folder=configs['io']['desc_folder'],
tmp_folder=configs['io']['tmp_folder'],
notes="Hcp, sc, fcc, diam and sc structures with 50% vacancies")
x, y, dataset_info = load_dataset_from_file(path_to_x=path_to_x, path_to_y=path_to_y,
path_to_summary=path_to_summary)
# write ASE database (wirting a json database does not work
# probably because there is a bug in ASE that prevents to write a json database from Jupyter Notebooks
write_ase_db(structure_list_vac50, main_folder=main_folder, db_name='hcp-sc-fcc-diam-bcc_vacancies-50%', db_type='db',
overwrite=True, folder_name='db_ase')
Counter(y)
# -
| examples/dataset_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DJCordhose/ml-workshop/blob/master/notebooks/data-science/1-3-matplotlib-oo-api.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="RRKecEtUhKC2"
# # Matplotlib OO API
#
# for more complex plots you will want to use the object oriented API
#
# * https://matplotlib.org/api/index.html#the-object-oriented-api
# * https://matplotlib.org/gallery/lines_bars_and_markers/simple_plot.html
# * https://matplotlib.org/api/axes_api.html#matplotlib.axes.Axes
# + colab_type="code" id="ehZLYjeFhKC7" colab={}
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
# + colab_type="code" id="arjNFGNuhKDB" colab={}
t = np.arange(0.0, 2.0, 0.01)
s = 1 + np.sin(2 * np.pi * t)
# + colab_type="code" id="yalsufnVhKDG" outputId="7ff495e2-2441-4602-8550-7b3b78bcce3e" colab={"base_uri": "https://localhost:8080/", "height": 350}
# just one subplot, also called axes (pl. of axis)
fig, ax = plt.subplots(1, figsize=(10, 5))
ax.plot(t, s)
ax.set_xlabel('time (s)')
ax.set_ylabel('voltage (mV)')
ax.set_title('About as simple as it gets, folks')
ax.grid(True)
fig.savefig("subplot.png")
# + colab_type="code" id="4HxHr4BShKDL" outputId="bffec7bb-2494-4b0b-bdce-9a0faf9104db" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls -l subplot.png
# + [markdown] id="1eM6xVYIta6j" colab_type="text"
# ### subplots vs subplot
#
# * <em>subplot</em> is an API inspired by MATLAB
# * its API is totally weird if MATLAB is not your mind set
# * use <em>subplots</em> as shown before if you want an intuitive interface
#
# because their names are so close to each other, if things do not behave as expected make sure you are using the right API
#
# + [markdown] id="XT6dPJJyta6j" colab_type="text"
# # More complex figures
# + [markdown] id="nElWq91Dta6k" colab_type="text"
# ## Naming can be confusing
#
# <img src='https://github.com/djcordhose/ml-workshop/blob/master/notebooks/data-science/img/figure_axes_axis_labeled.png?raw=1'>
#
# https://raw.githubusercontent.com/matplotlib/AnatomyOfMatplotlib/master/images/figure_axes_axis_labeled.png
# + colab_type="code" id="uO0tDd2XhKDO" outputId="ceadf876-0644-472a-c645-8bc690d91e0c" colab={"base_uri": "https://localhost:8080/", "height": 350}
fig, ax = plt.subplots(2, sharex='all', figsize=(10, 5))
ax[0].plot(t, s)
ax[0].set_ylabel('voltage (mV)')
ax[0].set_title('Linear')
ax[0].grid(True)
ax[0].set_yscale('linear')
ax[1].plot(t, s)
ax[1].set_xlabel('time (s)')
ax[1].set_ylabel('voltage (mV)')
ax[1].set_title('Log')
ax[1].grid(True)
ax[1].set_yscale('log')
# + [markdown] id="j7iulrtbta6m" colab_type="text"
# ## Anatomy of a figure
#
# <img src='https://github.com/djcordhose/ml-workshop/blob/master/notebooks/data-science/img/anatomy.png?raw=1'>
#
# https://matplotlib.org/tutorials/introductory/usage.html
# + [markdown] id="Lf1RAh_1vupe" colab_type="text"
# # Exercise: create a plot with more than one row and more than one column
# * use plt.subplots? to find out how to do that
# * what do you get as a result?
# * How to access the individual subplots?
# + id="58_LHKBTta6n" colab_type="code" colab={}
# plt.subplots?
# + [markdown] id="syDSSOnnB03x" colab_type="text"
# ## STOP HERE
#
# .
# .
# .
#
# ---
#
# .
# .
# .
#
#
#
# ---
#
# .
# .
# .
#
#
#
# ---
#
# .
# .
# .
#
#
#
# ---
#
# .
# .
# .
#
#
#
# ---
#
# .
# .
# .
#
#
#
# ---
#
# .
# .
# .
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# + id="7c4HePJnB1Zw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="fbf34f42-06b5-453f-97e5-70afca64a536"
fig, ax = plt.subplots(nrows=2, ncols=2)
# + id="nRR5DTZgCDyE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="568a84ea-7e99-4e5d-8664-ad33849e1a76"
type(ax)
# + id="8UVRADhMCA5Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cb8b48a5-df99-4bff-bff9-87f6874daba8"
ax.shape
# + id="8gvq3cycCFZ0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="9dfb000b-6951-435f-da10-facac55b361b"
ax
# + id="Ld-4TKJRCMRy" colab_type="code" colab={}
((ax1, ax2), (ax3, ax4)) = ax
# + id="3bf1FWE8CRhG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5b3694e5-e088-4df9-b00f-d9da332cfff5"
ax4
| notebooks/data-science/1-3-matplotlib-oo-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/muditbit/Diabetic-logistic/blob/master/diabetes_prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Z-x8rq8hURQw"
#importing some neccesary libraries
import numpy as np
import pandas as pd
import seaborn as sns
import sklearn
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="MiwHLS9dUZJ0" outputId="e97f178d-85f7-4d3e-c400-d6f3261b06ea"
#loading the dataset
df = pd.read_csv('diabetes.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="g0t4cl-u4jrN" outputId="ff33fb92-509d-4696-9e68-8a02c0bf0b23"
#Summarizing or getting some insight of the dataset that to be worked on
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="gzW1AApJ4pdM" outputId="c5191e0f-5ac1-4e30-9a9e-16f1e0fdff54"
#Checking the dataset for null values
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="5q5HTAD_4wB_" outputId="ada8bb5d-9d58-4422-fc8e-2d44e8cd5f7e"
#visualing the the balance of data i.e. no of true and false values or 0-1 values etc
df.Outcome.value_counts().plot(kind = 'bar').set_title('Diabetes Outcomes Value Count')
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="64yyqaIo5LZ0" outputId="d0ed7b0b-5b9f-4028-cfd1-7ada0f889cc2"
df.describe().round(decimals=2)
# + colab={"base_uri": "https://localhost:8080/", "height": 622} id="hbi1RraX5_oR" outputId="bbb74952-ee4b-4db0-faa1-5d7f702f36df"
#creating HeatMap of visualization it generates correlation matrix
sns.set(style='white')
mask = np.zeros_like(df.corr(), dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
#to Control the fig size
fig, ax = plt.subplots(figsize=(8,8))
cmap = sns.diverging_palette(255, 10, as_cmap=True)
sns.heatmap(df.corr(), mask=mask, annot=True, square=True, cmap=cmap , vmin=-1, vmax=1, ax=ax)
bottom, top = ax.get_ylim()
ax.set_ylim(bottom+0.5, top-0.5)
# + colab={"base_uri": "https://localhost:8080/", "height": 451} id="PsVYjeym9y4L" outputId="9f908ffb-2110-4014-885f-9a0030ff841f"
# To analyse feature-outcome distribution in visualisation
features = ['Pregnancies', 'Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'DiabetesPedigreeFunction', 'Age']
ROWS, COLS = 2, 4
fig, ax = plt.subplots(ROWS, COLS, figsize=(18,8) )
row, col = 0, 0
for i, feature in enumerate(features):
if col == COLS - 1:
row += 1
col = i % COLS
# df[feature].hist(bins=35, color='green', alpha=0.5, ax=ax[row, col]).set_title(feature) #show all, comment off below 2 lines
df[df.Outcome==0][feature].hist(bins=35, color='blue', alpha=0.5, ax=ax[row, col]).set_title(feature)
df[df.Outcome==1][feature].hist(bins=35, color='orange', alpha=0.7, ax=ax[row, col])
plt.legend(['No Diabetes', 'Diabetes'])
fig.subplots_adjust(hspace=0.3)
# + [markdown] id="g7YXZgo-_ua0"
# there are zero values in Glucose, BloodPressure, SkinThickness, Insulin, BMI as outliers
# + [markdown] id="QjEnvBfsKUtB"
# We need to fix these outlliers with the median of the column.
# + id="_nmbNjOC_lEl"
#Glucose
df.Glucose.replace(0, np.nan, inplace=True)
df.Glucose.replace(np.nan, df['Glucose'].median(), inplace=True)
#BloodPressure
df.BloodPressure.replace(0, np.nan, inplace=True)
df.BloodPressure.replace(np.nan, df['BloodPressure'].median(), inplace=True)
#skinThickness
df.SkinThickness.replace(0, np.nan, inplace=True)
df.SkinThickness.replace(np.nan, df['SkinThickness'].median(), inplace=True)
#insulin
df.Insulin.replace(0, np.nan, inplace=True)
df.Insulin.replace(np.nan, df['Insulin'].median(), inplace=True)
#BMI
df.BMI.replace(0, np.nan, inplace=True)
df.BMI.replace(np.nan, df['BMI'].median(), inplace=True)
# + id="Sw4HnyEBat_O"
# + colab={"base_uri": "https://localhost:8080/", "height": 451} id="ueGC42hajZQ8" outputId="c9de0ee6-654c-4e60-fa8a-35eef88b3fe0"
# To analyse feature-outcome distribution in visualisation
features = ['Pregnancies', 'Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'DiabetesPedigreeFunction', 'Age']
ROWS, COLS = 2, 4
fig, ax = plt.subplots(ROWS, COLS, figsize=(18,8) )
row, col = 0, 0
for i, feature in enumerate(features):
if col == COLS - 1:
row += 1
col = i % COLS
# df[feature].hist(bins=35, color='green', alpha=0.5, ax=ax[row, col]).set_title(feature) #show all, comment off below 2 lines
df[df.Outcome==0][feature].hist(bins=35, color='blue', alpha=0.5, ax=ax[row, col]).set_title(feature)
df[df.Outcome==1][feature].hist(bins=35, color='orange', alpha=0.7, ax=ax[row, col])
plt.legend(['No Diabetes', 'Diabetes'])
fig.subplots_adjust(hspace=0.3)
# + id="00ZWJnjXLaRq"
#Preprocessing
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
norm = scaler.fit_transform(df[['Pregnancies', 'Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'DiabetesPedigreeFunction', 'Age']])
df_norm = pd.DataFrame({'Pregnancies': norm[ :, 0], 'Glucose' : norm[ :, 1], 'BloodPressure' : norm[ :, 2], 'SkinThickness' : norm[ :, 3],
'Insulin' : norm[ :, 4], 'BMI' : norm[ :, 5], 'DiabetesPedigreeFunction' : norm[ :, 5], 'Age' : norm[ :, 6]},
columns=['Pregnancies', 'Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'DiabetesPedigreeFunction', 'Age'])
df_norm['Outcome'] = df['Outcome']
# split
x = df.drop(['Outcome'], axis=1)
y = df['Outcome']
# + id="SnqlqvGQ79c2" colab={"base_uri": "https://localhost:8080/"} outputId="54fe0af5-bdc4-4fee-cb06-b70bb37b1455"
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import SMOTE
x_train,x_test,y_train,y_test = train_test_split(x,y,random_state=42,
test_size = 0.3)
os = SMOTE(random_state=42)
columns = x_train.columns
os_data_x,os_data_y = os.fit_resample(x_train, y_train)
# + id="ZC4G5NHl_V2C"
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix,classification_report, accuracy_score
#Random Forest Classifier
rf_params = {'criterion' : ['gini', 'entropy'],
'n_estimators': list(range(5, 26, 5)),
'max_depth': list(range(3, 20, 2))}
rf_model = GridSearchCV(RandomForestClassifier(), rf_params, cv=5)
rf_model.fit(os_data_x, os_data_y)
rf_predict = rf_model.predict(x_test)
rf_cm = confusion_matrix(y_test, rf_predict)
rf_score = rf_model.best_score_
# + id="vCtNpf4sTxHp"
from sklearn.tree import DecisionTreeClassifier
#Decision Tree
dt_params = {'criterion' : ['gini', 'entropy'],
'splitter': ['random', 'best'],
'max_depth': [3, 5, 7, 9, 11, 13]}
dt_model = GridSearchCV(DecisionTreeClassifier(), dt_params, cv=5)
dt_model.fit(os_data_x, os_data_y)
dt_predict = dt_model.predict(x_test)
dt_cm = confusion_matrix(y_test, dt_predict)
dt_score = dt_model.best_score_
# + colab={"base_uri": "https://localhost:8080/"} id="cJuFcdgCUtAB" outputId="ca917608-9864-425a-eb5c-9ed4492892f6"
# xgb
import xgboost as xgb
xgb_params = {'max_depth': [3, 5, 7, 9],
'n_estimators': [5, 10, 15, 20, 25, 50, 100],
'learning_rate': [0.01, 0.05, 0.1]}
xgb_model = GridSearchCV(xgb.XGBClassifier(eval_metric='logloss'), xgb_params, cv=5)
xgb_model.fit(os_data_x, os_data_y)
xgb_predict = xgb_model.predict(x_test.values)
xgb_cm = confusion_matrix(y_test, xgb_predict)
xgb_score = xgb_model.best_score_
xgb_score
# + colab={"base_uri": "https://localhost:8080/"} id="YkH2fq48VHRY" outputId="195d2e59-b8e9-4c0b-8b4b-2d41f67499ae"
# svc
from sklearn.svm import SVC
svc_params = {'C': [0.001, 0.01, 0.1, 1],
'kernel': [ 'linear' , 'poly' , 'rbf' , 'sigmoid' ]}
svc_model = GridSearchCV(SVC(), svc_params, cv=5)
svc_model.fit(os_data_x, os_data_y)
svc_predict = svc_model.predict(x_test)
svc_cm = confusion_matrix(y_test, svc_predict)
svc_score = svc_model.best_score_
print(svc_score)
# + colab={"base_uri": "https://localhost:8080/"} id="7PTQVVeZWDDy" outputId="aded1da5-f858-4b53-8880-7bc58a7b30f4"
# knn
from sklearn.neighbors import KNeighborsClassifier
knn_params = {'n_neighbors': list(range(3, 20, 2)),
'weights':['uniform', 'distance'],
'algorithm':['auto', 'ball_tree', 'kd_tree', 'brute'],
'metric':['euclidean', 'manhattan', 'chebyshev', 'minkowski']}
knn_model = GridSearchCV(KNeighborsClassifier(), knn_params, cv=5)
knn_model.fit(os_data_x, os_data_y)
knn_predict = knn_model.predict(x_test)
knn_cm = confusion_matrix(y_test, knn_predict)
knn_score = knn_model.best_score_
print(knn_score)
# + colab={"base_uri": "https://localhost:8080/"} id="zOekTbergoVC" outputId="221a6425-24bc-4ae1-d7bd-d962e69c26d6"
# !pip install catboost
# + id="chytjBRBgsSi"
# cb
from catboost import CatBoostClassifier
cb_params = {'learning_rate': [0.01, 0.05, 0.1],
'depth': [3, 5, 7, 9]}
cb_model = GridSearchCV(CatBoostClassifier(verbose=False), cb_params, cv=5)
cb_model.fit(os_data_x, os_data_y)
cb_predict = cb_model.predict(x_test)
cb_cm = confusion_matrix(y_test, cb_predict)
cb_score = cb_model.best_score_
# + id="sBc8zrT-WwV1"
# Deciding the winner
mdls = {'DecisionTree': dt_score*100,
'CatBoost':cb_score*100,
'XGBClassifier':xgb_score*100,
'SVM':svc_score*100,
'RandomForest':rf_score*100,
'KNN':knn_score*100}
# + colab={"base_uri": "https://localhost:8080/"} id="V_NRpNdfYGHl" outputId="d9f5aaa9-8c52-4dfe-cc40-79a398b80a31"
mdls
# + id="SoT3qe5vYHf-"
win = pd.DataFrame(list(mdls.items()),columns=['Model','Score'])
# + id="ie1uZtCaYmkM"
win.sort_values(by=['Score'],inplace=True,ascending=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="Epjz47J7YqD0" outputId="5bd15b61-9f8d-48f7-e1b4-3c71cfc22234"
win
# + colab={"base_uri": "https://localhost:8080/", "height": 322} id="IJuBI2c-h4nz" outputId="3bebd263-bcb6-4703-ffdf-a167aae51f03"
plt.figure(figsize= (10,5))
plt.bar(win.Model,win.Score, align = 'center', width =0.8,
color = 'green',edgecolor='yellow',linewidth=1)
plt.show()
# + id="zkdm_5Umh4iV"
# + id="QRWi67I_ZSfh" colab={"base_uri": "https://localhost:8080/"} outputId="709c20ba-c3c7-4385-8dfe-e6ec6598e5e5"
# !pip freeze
# + id="v0p27FjEabve"
| diabetes_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Computational Molecular Medicine Course Project
#
#
# <NAME>, <NAME>
# ### Frame the Problem
#
# - Background
#
# One of the most intense research areas in computational molecular medicine over the past fifteen years is the prediction of cellular phenotypes, e.g., properties of cancerous growths, based on gene expression profiles. <br>
#
# This project is about head and neck cancers, which arise in cells lining mucosal surfaces inside the head and neck (e.g., mouth, nose, and throat). This type of cancer is further categorized by the site of origination (oral cavity, pharynx, larynx, salivary glands, nasal cavity, and paranasal sinuses). These cancers account for about 4%-5% of all cancers in the US, are more common in men, and are diagnosed more often among in people over 50. About 65,000 individuals are diagnosed with head and neck cancers in the US yearly.<br>
#
# The most important causes of this type of cancer are alcohol, smoking, smokeless tobacco, and HPV infection. Treatment will depend on the exact tumor location, cancer stage, extension to local lymph nodes, the presence of metastasis, age, and general health. Treatment usually is a combination of surgery, radiation therapy, chemotherapy, and targeted therapy. <br>
#
#
# Cancer might present with or without metastasis to loco-regional lymph-nodes (in this case the neck). The presence of such metastases requires more aggressive treatments. **Unfortunately non-invasive methods to assess metastases in the neck lymph-nodes are not accurate and it is usually necessary to surgically remove the lymph-nodes to investigate the presence or absence of metastases** (neck lymph-nodes dissection).
#
# For oral cavity cancers with stage T3 and T4 the probability of lymph-node metastases is high, hence neck lymph-nodes dissection is required. In early stage cancers (stage T1 and T2), however, such probabilities of metastases are around 6% (T1) and 20% (T2). **Therefore, in this group of patients a method to accurately identify patients with and without lymph node metastases could greatly reduce over-treatment by sparing neck lymph-nodes dissection to patients who do not have metastases.**
#
# - Data
#
#
# We have assembled gene expression data from the public domain for a population of patients with early-stage oral cavity head and neck cancer, predominantly HPV negative, and with known lymph-node metastasis status ("NodalStatus"), which can be positive (Y = 1) or negative (Y = 0). <br>
#
# There are two datasets. One dataset was generated using RNAseq (TCGA project) and has a total of 109 samples (27 with positive nodes and 82 with negative nodes). It consists of expression data and phenotype information. The second dataset was generated with a different technology using microarrays and has a total of 112 samples (46 with positive nodes and 66 with negative nodes). It also consists of expression data and phenotype information.
#
# - Objective
#
# Build a classifier to predict "NodalStatus."
#
# - Evaluation
#
# Whereas obtaining high accuracy may be the clinical objective, it is not the primary measure of performance for this assignment. Your effort will be evaluated by various criteria, including **creativity, (mathematical) coherence, parsimony, and proper validation**. Finally, in order to allow us to compare your results and frame them in a more clinically realistic setting, **determine the specificity on your test dataset that can be achieved while maintaining 80% sensitivity, where the classifier is trained on your training dataset. Associate sensitivity with positive node status, and just show the ROC curve and report spec(t80).**
#
# ---
# Before we start to build a model, we should answer the question first. **What is the specific gap in the ideal world and the real one that requires machine learning to fill? **
#
#
# Traditionally non-invasive methods to assess metastases in the neck lymph-nodes are not accurate and it is usually necessary to surgically remove the lymph-nodes to investigate the presence or absence of metastases (neck lymph-nodes dissection). Using Machine Learning predictive algorithms to build a more precise model could help us reduce over-treatment by sparing neck lymph-nodes dissection to patients who do not have metastases.
#
#
# import some libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import random
import seaborn as sns
# common model helpers
from sklearn.preprocessing import StandardScaler
from sklearn import metrics
from sklearn.model_selection import train_test_split,cross_val_score
from sklearn.metrics import roc_curve, auc
import scipy
from scipy.stats import ranksums,mannwhitneyu
# Machine Learning algorithms
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.linear_model import LogisticRegression
# Hyper-parameter tuning
from sklearn.model_selection import GridSearchCV,KFold
# ### Get the Data
micro_exprs = pd.read_csv('arrayStudyExprs.csv')
micro_exprs = micro_exprs.T
micro_exprs = micro_exprs.rename(columns=micro_exprs.iloc[0])
micro_exprs = micro_exprs.reset_index()
micro_exprs = micro_exprs.drop([0])
micro_exprs = micro_exprs.rename(index=str, columns={"index": "sampleID"})
micro_pheno = pd.read_csv('arrayStudyPheno.csv')
micro_pheno = micro_pheno.drop('Unnamed: 0',axis=1)
rna_exprs = pd.read_csv('rnaSeqStudyExprs.csv')
rna_exprs = rna_exprs.T
rna_exprs = rna_exprs.rename(columns=rna_exprs.iloc[0])
rna_exprs = rna_exprs.reset_index()
rna_exprs = rna_exprs.drop([0])
rna_exprs = rna_exprs.rename(index=str, columns={"index": "sampleID"})
rna_pheno = pd.read_csv('rnaSeqStudyPheno.csv')
rna_pheno = rna_pheno.drop('Unnamed: 0',axis=1)
# Dataset generated using RNA-seq
rna_pheno.head()
print(rna_pheno.shape)
print('Variable', ' '*13, 'count of unique value',' '*15, 'content')
for column in rna_pheno.columns:
uniques = rna_pheno[column].unique()
print('{0:26s} {1:13d}\t'.format(column, len(uniques)), uniques[:4])
print('-'*120)
print("Total records in the RNAsequence phenotype information set:",len(rna_pheno))
print("Total vairables in the RNAsequence phenotype information set:",rna_pheno.shape[1])
print('Total missing values in the RNAsequence phenotype information set:\n',rna_pheno.isnull().sum())
rna_exprs.head()
print("Total records in the RNAsequence expression set:",len(rna_exprs))
print("Total vairables in the RNAsequence expression set:",rna_exprs.shape[1]-1)
print('Total missing values in the RNAsequence expression set:',rna_exprs.isnull().sum().sum())
# We can see that dataset generated by RNAsequence includes 109 patients(data instances), for gene expression data, it can be viewed as 109 observations with 9223 features; for phenotype information data, it can be considered as 109 observations with 16 variables. Some of the variables are with many missing values. Additionally, the last phenotype variable 'NodeStatus' which is the target variable of our task, has two unique values with positive and negtive.
# Dataset generated using Microarray
micro_pheno.head()
print(micro_pheno.shape)
print('Variable', ' '*13, 'count of unique value',' '*15, 'content')
for column in micro_pheno.columns:
uniques = micro_pheno[column].unique()
print('{0:26s} {1:13d}\t'.format(column, len(uniques)), uniques[:2])
print('-'*120)
print("Total records in the microarray phenotype information set:",len(micro_pheno))
print("Total variables in the microarray phenotype information set:",micro_pheno.shape[1])
print("Total missing values in the microarray phenotype information set:\n",micro_pheno.isnull().sum())
micro_exprs.head()
print("Total records in the microarray gene expression set:",len(micro_exprs))
print("Total vairables in the microarray gene expression set:",micro_exprs.shape[1]-1)
print("Total missing values in the microarray gene expression set:",micro_exprs.isnull().sum().sum())
# It is clear that dataset generated by microarray method includes 112 patients(data instances), for gene expression data, it can be viewed as 112 observations with 9223 features; for phenotype information data, it can be considered as 112 observations with 16 variables. Additionally, the last phenotype variable 'NodeStatus' which is the target variable of our task, has two unique values with positive and negtive. Except for the number of patients, the rest set up is the same as RNAsequence dataset.
# - It is clear that the number of features is much larger than the amount of data, but we could not combine two dataset together because they are generated using different technologies and may occur 'platform effects'.
#
#
# **Differentially Expressed Genes**
# A typical bioinformatics problem (e.g. classification) with high-dimensional omics data includes redundant and irrelevant features that can result, in the worst-case scenario, in false positive results. Then, feature selection becomes significantly important. The classification of gene expression data samples involves feature selection and classifier design.
# RNA-seq and microarray are two main technologies for profiling gene expression levels. As we observed before, the two datasets holds thousands of gene expression with hundreds of observations, which is the main characteristic but not friendly to applying machine learning algorithms. In the absence of feature selection, classification accuracy on the training data is typically good, but not replicated on the testing data. Except for considering accuracy of model, determining differentially expressed genes (DEGs) between biological samples is the key to understand how genotype gives rise to phenotype, here is 'NodeStatus' variable. A gene is declared differentially expressed if an observed difference or change in expression levels between two experimental conditions is statistically significant.
#
#
# A variety of methods have been published to detect differentially expressed genes. Some methods are based on non-statistical quantification of expression differences, but most methods are based on statistical tests to quantify the significance of differences in gene expression between samples.
# These statistical methods can furthermore be divided into two methodological categories: parametric tests and non-parametric tests. Here I will use non-parametric test Wilcoxon rank sum test to select a small subset(DEGs) out of the 9,223 of genes in microarray data and RNA-seq data for further study.
#
# ### Prepare the data
# **1. Adding some promising variables from phenotype dataset**
#
# After checking the missing values, I decide to combine some of phenotype variables with gene expression data to classify 'NodeStatus' variable. Further, since observations are limited to a small magnitude, we will not choose the variable with too many missing values which may add noise to the dataset. Consequently, we add 'gender', 'smoking', 'Tstage' and 'age' features, and convert their formats for running algorithms.
rna_pheno.loc[rna_pheno['gender']=='F','gender']=0
rna_pheno.loc[rna_pheno['gender']=='M','gender']=1
rna_pheno.loc[rna_pheno['smoking']=='YES','smoking']=0
rna_pheno.loc[rna_pheno['smoking']=='NO','smoking']=1
rna_pheno.loc[rna_pheno['Tstage']=='T1','Tstage']=0.06
rna_pheno.loc[rna_pheno['Tstage']=='T2','Tstage']=0.2
rna_pheno['smoking'].fillna(np.median(rna_pheno['smoking']), inplace=True)
rna_pheno['age'].fillna(round(np.mean(rna_pheno['age']),0), inplace=True)
rna_pheno.head()
micro_pheno.head()
micro_pheno.loc[micro_pheno['gender']=='F','gender']=0
micro_pheno.loc[micro_pheno['gender']=='M','gender']=1
micro_pheno.loc[micro_pheno['smoking']=='YES','smoking']=0
micro_pheno.loc[micro_pheno['smoking']=='NO','smoking']=1
micro_pheno.loc[micro_pheno['Tstage']=='T1','Tstage']=0.06
micro_pheno.loc[micro_pheno['Tstage']=='T2','Tstage']=0.2
micro_pheno.head()
# **2. Process target variable 'NodeStatus'**
#
# Replacing 'NEG' negtive with zero and 'POS' positive with value one for 'NodeStatus' and combining it with gene expression dataset.
rna_pheno.loc[rna_pheno['NodeStatus']=='NEG','NodeStatus']=0
rna_pheno.loc[rna_pheno['NodeStatus']=='POS','NodeStatus']=1
rna = pd.merge(rna_pheno[['sampleID','NodeStatus','age','gender','smoking','Tstage']],rna_exprs, how = 'right', on = 'sampleID')
rna.head()
micro_pheno.loc[micro_pheno['NodeStatus']=='NEG','NodeStatus']=0
micro_pheno.loc[micro_pheno['NodeStatus']=='POS','NodeStatus']=1
micro = pd.merge(micro_pheno[['sampleID','NodeStatus','age','gender','smoking','Tstage']],micro_exprs, how = 'right', on = 'sampleID')
micro.head()
# **2. Train vs Validation vs Test**
# We create a function specifically to divide the data into training and test sets because of the imbalance in the target variables. Training set will further be splitted into validation and train set.
def split_data(dataset):
random.seed(7)
pos = sum(dataset['NodeStatus']==1)
neg = sum(dataset['NodeStatus']==0)
index0 = list(dataset[dataset['NodeStatus']==0].index)
index1 = list(dataset[dataset['NodeStatus']==1].index)
random.shuffle(index0)
random.shuffle(index1)
test_index = index0[:round(neg*0.35)] + index1[:round(pos*0.35)]
random.shuffle(test_index)
train_index = index0[round(neg*0.35):] + index1[round(pos*0.35):]
random.shuffle(train_index)
Test = dataset.iloc[test_index]
Train = dataset.iloc[train_index]
X_test = Test.drop(['NodeStatus'],axis=1)
y_test = Test['NodeStatus']
X_train = Train.drop(['NodeStatus'],axis=1)
y_train = Train['NodeStatus']
return X_test,y_test,X_train,y_train,Train
rna_X_test, rna_y_test, rna_X_train, rna_y_train, rna_Train = split_data(rna)
micro_X_test, micro_y_test, micro_X_train, micro_y_train, micro_Train = split_data(micro)
# **3. Feature selection using Wilcoxon rank sum test**
# Wilcoxon rank sum test is a non-parametric method to detect differentially expressed genes and Python has already designed that test for us. After running Wilcoxon rank sum test, we have to order the gene expression by putting p-value from small to large, and select the gene with p-value as small as possible, which means the genes whose expression levels are statistically different between groups negative and groups positive. Obviously, it is more effective and helpful for our classification task. Additionally, we are planning to select various volume of subsets based on training set for RNA-seq and microarray dataset because we have no idea how many features are appropriate for our model.
rna_Train.head()
# +
def detect_genes(dataset):
stat = []
pval = []
for i in range(6,dataset.shape[1]):
data0 = dataset[dataset['NodeStatus']==0].iloc[:,[i]].values
data1 = dataset[dataset['NodeStatus']==1].iloc[:,[i]].values
stat.append([scipy.stats.mannwhitneyu(data0,data1)[0],i])
pval.append([scipy.stats.mannwhitneyu(data0,data1)[1],i])
return stat, pval
## When the number of observation in each sample is > 20 and you have 2 independent samples of ranks, scipy.stats.mannwhitneyu
## will perform better than scipy.stats.ranksums, but basically they are the same.
# -
stat_rna, pval_rna = detect_genes(rna_Train)
stat_micro, pval_micro = detect_genes(micro_Train)
# **4. According to test result, we could select any different number of features which directly affects the performance of the model. Consequently, It should be considered the number of features as a hyper-parameter and using cross-validation to determine the precise value.**
#
def gene_subset(dataset,pval,num):
pval_sort = sorted(pval)[:num]
index = []
for i in range(num):
index.append(pval_sort[i][1])
index.extend([1,2,3,4])
subset = dataset.iloc[:,index]
return subset
print('The first 20 most differentially expressed genes set for RNA-seq dataset includes:')
print(sorted(gene_subset(rna_X_train,pval_rna,20).columns))
print('The first 20 most differentially expressed genes set for Microarray dataset includes:')
print(sorted(gene_subset(micro_X_train,pval_micro,20).columns))
# From above detection results, we can find that for these two datasets, their differentially expressed genes at least the first 20 genes are totally different. Consequently, we could not directly integrate gene expression profiles across two different platforms. Since there's no requirement to combine them together, we will build models for these two different datasets, respectively.
# **5. Visualize some of the important features.**
print('Distributions of the first 16 differentially expressed genes based on RNA-seq training data.')
print('-'*90)
rna_subset_16 = gene_subset(rna_X_train,pval_rna,16)
rna_subset_16 = rna_subset_16.astype(float)
rna_subset_16.hist();
plt.tight_layout()
print('Distributions of the first 16 differentially expressed genes based on microarray training data.')
print('-'*90)
micro_subset_16 = gene_subset(micro_X_train,pval_micro,16)
micro_subset_16 = micro_subset_16.astype(float)
micro_subset_16.hist();
plt.tight_layout()
# **6. Feature scaling**
# From the distributions of some features, we could find that it seems like they all fall into the same magnitude which is good. However, some of algorithms may be sensitive to it for example Support Vector Machines. I tend to standardize features by removing the mean and scaling to unit variance.
#
#
def standprocess(train,test):
stdSc = StandardScaler()
train = stdSc.fit_transform(train)
test = stdSc.transform(test)
return train, test
rnas = rna_subset_16
micros = micro_subset_16
train_rna_16, train_micro_16 = standprocess(rnas,micros)
print('Distributions of the first 16 differentially expressed genes after scaling based on RNA-seq training data.')
print('-'*110)
train_rna_16 = pd.DataFrame(train_rna_16)
train_rna_16.columns = list(rna_subset_16.columns)
train_rna_16.hist();
plt.tight_layout()
print('Distributions of the first 16 differentially expressed genes after scaling based on microarray training data.')
print('-'*110)
train_micro_16 = pd.DataFrame(train_micro_16)
train_micro_16.columns = list(micro_subset_16.columns)
train_micro_16.hist();
plt.tight_layout()
# ### Short-List Promising Models
#Validation function
n_folds = 5
def rmsle_cv(model,x_train,y_train):
kf = KFold(n_folds, shuffle=True, random_state=777).get_n_splits(x_train)
auc_score= np.mean(cross_val_score(model, x_train, y_train, cv = kf,scoring='roc_auc'))
return(auc_score)
DT = DecisionTreeClassifier(random_state=777)
svm = SVC(probability = True)
rf = RandomForestClassifier(random_state=777,class_weight='balanced')
knn = KNeighborsClassifier()
ada = AdaBoostClassifier(random_state=777)
lr = LogisticRegression(random_state=777)
x_train_m = gene_subset(micro_X_train,pval_micro,250)
x_train_r = gene_subset(rna_X_train,pval_rna,250)
scorem = rmsle_cv(DT,x_train_m,micro_y_train)
print("\nDecision Tree Auc score on microarray dataset: {:.4f})".format(scorem.mean()))
scorer = rmsle_cv(DT,x_train_r,rna_y_train)
print("Decision Tree Auc score on RNA-seq dataset: {:.4f})".format(scorer.mean()))
scorem = rmsle_cv(svm,x_train_m,micro_y_train)
print("\nSupport Vector Machine Auc score on microarray dataset: {:.4f}".format(scorem.mean()))
scorer = rmsle_cv(svm,x_train_r,rna_y_train)
print("Support Vector Machine Auc score on RNA-seq dataset: {:.4f}".format(scorer.mean()))
scorem = rmsle_cv(rf,x_train_m,micro_y_train)
print("\nRandom Forest Auc score on microarray dataset: {:.4f}".format(scorem.mean()))
scorer = rmsle_cv(rf,x_train_r,rna_y_train)
print("Random Forest Auc score on RNA-seq dataset: {:.4f}".format(scorer.mean()))
scorem = rmsle_cv(knn,x_train_m,micro_y_train)
print("\nk-nearest neighbors Auc score on microarray dataset: {:.4f}".format(scorem.mean()))
scorer = rmsle_cv(knn,x_train_r,rna_y_train)
print("k-nearest neighbors Auc score on RNA-seq dataset: {:.4f}".format(scorer.mean()))
scorem = rmsle_cv(ada,x_train_m,micro_y_train)
print("\nAdaboost Auc score on microarray dataset: {:.4f}".format(scorem.mean()))
scorer = rmsle_cv(ada,x_train_r,rna_y_train)
print("Adaboost Auc score on RNA-seq dataset: {:.4f}".format(scorer.mean()))
scorem = rmsle_cv(lr,x_train_m, micro_y_train)
print("\nLogistic Regression Auc score on microarray dataset: {:.4f}".format(scorem.mean()))
scorer = rmsle_cv(lr,x_train_r, rna_y_train)
print("Logistic Regression Auc score on RNA-seq dataset: {:.4f}".format(scorer.mean()))
# We try many quick and dirty models from different categories using standard parameters. After measuring and comparing their performance, we plan to pick Random Forest, Logistic Regression and KNN alogrithms because they are fitting well now and seem like promising models after fine tuning the hyperparameters. Since the number of data instances is relatively small, we do not need to consider running time and space. I will use **GridSearchCV** function to tweak parameters.
# ### Fine-Tune the System
# In this part, we will use grid search cross-validation method to tweak the hyper-parameters for different models based on two different datasets and pick the best model with the best subset of genes to make prediction on the test set. Then plot the ROC curve and return the specificity on the test dataset that can be achieved while maintaining 80% sensitivity for each case.
def Gridsearchcv(x_train,y_train,model,param,cv_num):
# random search + cross validation
gridsearch = GridSearchCV(model,param,cv = cv_num,scoring='roc_auc',n_jobs = 7)
# train model in train set
gridsearch.fit(x_train,y_train)
# return search results for each case
result = gridsearch.cv_results_
for mean_score, params in zip(result["mean_test_score"], result["params"]):
print(np.mean(mean_score), params)
# get the best estimator
best_model = gridsearch.best_estimator_
print('Best model:\n')
print(best_model)
print('\n Optimal parameters:')
print(gridsearch.best_params_)
best_model.fit(x_train,y_train)
return best_model
def ROC_curve(x_train,y_train,x_test,y_test,model):
fprs = []
tprs = []
threshold = []
# use model fit training data
model.fit(x_train,y_train)
y_train_pred = model.predict_proba(x_train)
# compute tpr, fpr to plot ROC curve for training set
fpr_train, tpr_train, thresholds = roc_curve(y_train, y_train_pred[:,1])
roc_auc_train = auc(fpr_train, tpr_train)
# use model predict test data
y_pred = model.predict_proba(x_test)
y_preds = model.predict(x_test)
# compute tpr, fpr to plot ROC curve for test set
fpr_test, tpr_test, thresholds = roc_curve(y_test, y_pred[:,1])
fprs.append(fpr_test)
tprs.append(tpr_test)
threshold.append(thresholds)
roc_auc_test = auc(fpr_test, tpr_test)
# Plot ROC curve and compare them
plt.plot(fpr_train, tpr_train, linewidth=2, label='Train AUC = %0.2f'% roc_auc_train);
plt.plot(fpr_test, tpr_test, linewidth=2, label='Test AUC = %0.2f'% roc_auc_test)
plt.title("Receiving Operating Characteristic")
plt.legend(loc="lower right")
plt.plot([0, 1], [0, 1], 'r--')
plt.axis([0, 1, 0, 1])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
return tprs, fprs, threshold
# **1. Random Forest**
# - Tuning parameters based on microarray datset with the first 80 differentially expressed genes.
## Microarray dataset
train_m = gene_subset(micro_X_train,pval_micro,80)
test_m = gene_subset(micro_X_test,pval_micro,80)
train_M, test_M = standprocess(train_m, test_m)
rf = RandomForestClassifier(random_state=11,class_weight={0:0.46,1:0.54})
print('Grid search results based on microarray dataset:\n')
param = {'n_estimators':[200,300],
'max_depth':[10,20],
'min_samples_leaf':[5,7],
'min_samples_split':[2,5],
'class_weight':[{0:0.4,1:0.6}, {0:0.42,1:0.58},{0:0.45, 1:0.55}]}
best_rf = Gridsearchcv(train_m,micro_y_train,rf,param,3)
# - Evaluation on the test set
# plot roc curve
x_train = train_m; y_train = micro_y_train; x_test = test_m; y_test = micro_y_test; model = best_rf
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# Visualize feature importance
feature_imp = pd.Series(best_rf.feature_importances_,
index=train_m.columns).sort_values(ascending=False)
a=sns.barplot(x=feature_imp[:20], y=feature_imp.index[:20])
# Add labels to your graph
plt.xlabel('Feature Importance Score')
a.set_xscale('log')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.show()
print('Feature rank:')
genes = []
for score, feature in sorted(zip(best_rf.feature_importances_, train_m.columns), reverse=True)[:20]:
if score >0.01:
genes.append(feature)
print('{0:26} {1:3}'.format(feature,score))
# - Tuning parameters based on RNA-seq datset with the first 450 differentially expressed genes.
## RNA-seq dataset #450
train_r = gene_subset(rna_X_train,pval_rna,450)
test_r = gene_subset(rna_X_test,pval_rna,450)
train_R, test_R = standprocess(train_r, test_r)
rf = RandomForestClassifier(random_state=7,class_weight={0:0.46,1:0.54})
print('Grid search results based on microarray dataset:\n')
param = {'n_estimators':[200,300],
'max_depth':[10,20],
'min_samples_leaf':[5,7],
'min_samples_split':[2,5],
'class_weight':[{0:0.4,1:0.6}, {0:0.42,1:0.58},{0:0.45, 1:0.55}]}
best_rf = Gridsearchcv(train_r,rna_y_train,rf,param,3)
# - Evaluation on the test set
# plot roc curve
x_train = train_r; y_train = rna_y_train; x_test = test_r; y_test = rna_y_test; model = best_rf
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# Visualize feature importance
feature_imp = pd.Series(best_rf.feature_importances_,
index=train_r.columns).sort_values(ascending=False)
a=sns.barplot(x=feature_imp[:20], y=feature_imp.index[:20])
# Add labels to your graph
plt.xlabel('Feature Importance Score')
a.set_xscale('log')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.show()
print('Feature rank:')
genes = []
for score, feature in sorted(zip(best_rf.feature_importances_, train_r.columns), reverse=True)[:20]:
if score >0.01:
genes.append(feature)
print('{0:26} {1:3}'.format(feature,score))
#
# **2. Logistic Regression**
#
# - Based on RNA-seq datset with the first 510 differentially expressed genes.
## RNA-seq dataset
train_r = gene_subset(rna_X_train,pval_rna,510)
test_r = gene_subset(rna_X_test,pval_rna,510)
## standardize the dataset
train_R, test_R = standprocess(train_r,test_r)
lr = LogisticRegression(random_state=111,class_weight='balanced')
print('Grid search results based on microarray dataset:\n')
param = {'C':[1,10,20],'penalty':['l1','l2']}
best_lr = Gridsearchcv(train_R,rna_y_train,lr,param,5)
x_train = train_R; y_train = rna_y_train; x_test = test_R; y_test = rna_y_test; model = best_lr
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# - Based on microarray datset with the first 200 differentially expressed genes.
## Microarray dataset
train_m = gene_subset(micro_X_train,pval_micro,200)
test_m = gene_subset(micro_X_test,pval_micro,200)
## standardize the dataset
train_M, test_M = standprocess(train_m,test_m)
lr = LogisticRegression(random_state=77,class_weight='balanced')
print('Grid search results based on microarray dataset:\n')
param = {'C':[1,10,15,20],'penalty':['l1','l2']}
best_lr = Gridsearchcv(train_M,micro_y_train,lr,param,5)
x_train = train_M; y_train = micro_y_train; x_test = test_M; y_test = micro_y_test; model = best_lr
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# **3. k-nearest neighbors**
#
# - Based on RNA-seq datset with the first 40 differentially expressed genes.
## RNA-seq dataset
train_r = gene_subset(rna_X_train,pval_rna,40)
test_r = gene_subset(rna_X_test,pval_rna,40)
## standardize the dataset
train_R, test_R = standprocess(train_r,test_r)
knn = KNeighborsClassifier()
print('Grid search results based on microarray dataset:\n')
param = {'n_neighbors':[6,7,8,9,10],'weights':['uniform','distance'],
'p':[1,2,3]}
random.seed(7)
best_knn = Gridsearchcv(train_R,rna_y_train,knn,param,5)
x_train = train_R; y_train = rna_y_train; x_test = test_R; y_test = rna_y_test; model = best_knn
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# - Based on microarray datset with the first 40 differentially expressed genes.
#
## Microarray dataset
train_m = gene_subset(micro_X_train,pval_micro,40)
test_m = gene_subset(micro_X_test,pval_micro,40)
## standardize the dataset
train_M, test_M = standprocess(train_m,test_m)
knn = KNeighborsClassifier()
random.seed(77)
print('Grid search results based on microarray dataset:\n')
param = {'n_neighbors':[6,7,8,9],'weights':['uniform','distance'],
'p':[1,2,3,4]}
best_knn = Gridsearchcv(train_M,micro_y_train,knn,param,5)
x_train = train_M; y_train = micro_y_train; x_test = test_M; y_test = micro_y_test; model = best_knn
tpr, fpr, threshold = ROC_curve(x_train,y_train,x_test,y_test,model)
print('Sensitivity:',tpr[0][tpr[0]>=0.8][0])
print('Specificity:',1-fpr[0][np.where(tpr[0] >= 0.8)[0][0]])
# ### Present the Solution
# In our project, random forest, k-Nearest Neighbors, and logistic regression are utilized to establish a classifier.
# In order to obtain comparable results, we try all three methods on RNA sequence dataset and microarrays dataset. First, we process the target variable "NodeStatus", which equals to 1 corresponding to "positive" and 0 corresponding to "negative". Second, we pick some relevant features 'age', 'smoking', and 'gender' from the phenotype dataset, convert and combine them with gene expressions dataset. Then we split both datasets randomly into the training set and test set. Training set will be further split into validation set which helps us pick parameters and train set which is utilized to train models. We will not touch test set until the last moment comes when all models are set up and tuned well. Additionally, detecting differentially expressed genes is very important for building our classifier. There are various methods to identify DEGs and here we use a non-parametric one Wilcoxon rank sum test because the amount of our dataset is relatively too small. We create functions to process, split datasets and standardize features which are prerequisite for some of machine learning algorithms. After preparing the data, we try several categories methods, compare their performance and pick some of the promising ones and use GridSearch cross-validation method to extensively tune hyper-parameters. Evaluate the results on RNA sequence dataset and microarray dataset with a different number of differentially expressed genes by plotting ROC curve and reporting Sensitivity and Specificity, respectively. For each ROC curve, we find the point which sens($t_{80}$) ≥ 0.80 and report the corresponding specificity spec($t_{80}$).
#
# - Comparison of results
#
#
#
#
# | Dataset | |Random Forest | Logistic Regression | K-Nearest Neighbors |
# | :------ | :--------------------------------: |
# | RNA-seq | [Sensitivity](https://en.wikipedia.org/wiki/Sensitivity_and_specificity) | 0.8889(450DEGs)|0.8889(510DEGs)|0.8889(40DEGs)|
# | | [Specificity](https://en.wikipedia.org/wiki/Sensitivity_and_specificity) | 0.4828(450DEGs)|0.4138(510DEGs)|0.3793(40DEGs)|
# |Microarray | [Sensitivity](https://en.wikipedia.org/wiki/Sensitivity_and_specificity) | 0.8125(80DEGs) |0.8125(200DEGs)|0.8125(40DEGs)|
# | | [Specificity](https://en.wikipedia.org/wiki/Sensitivity_and_specificity) | 0.3478(80DEGs) |0.5217(200DEGs)|0.3043(40DEGs)|
# > DEGs is short for differentially expressed genes.
#
#
# - Describe what worked, what did not and what can try in the future.
# 1. Standardizing differentially expressed genes and adjusting the number of DEGs in the training and test set help us improve the AUC score.
# 2. Since the number of features is much larger than the amount of dataset, for different classification algorithms, the performance will be significantly influenced by selected features. The optimal number of genes to be selected is variant and can be considered as a hyper-parameter.
# 3. Directly combining RNA-seq with microarray dataset together to fit the model does not improve classification performance.
# 4. Domain knowledge is a requirement for merging two datasets collected by different technology, but if it works, it will greatly improve the accuracy of our classification.
# 5. Other detecting methods of DEGs may work, except for some statistical methods, it is a good idea to use feature importance of tree models, combining them with genes selected by Wilcoxon rank sum test to train a model.
#
#
# - Reference<br>
# [Microarray Data Analysis](https://discover.nci.nih.gov/microarrayAnalysis/Statistical.Tests.jsp)<br>
# [Integrating gene expression profiles across different platforms](https://www.rna-seqblog.com/integrating-gene-expression-profiles-across-different-platforms/)<br>
# [An Improved Method on Wilcoxon Rank Sum Test for Gene Selection from Microarray Experiments](https://www.tandfonline.com/doi/abs/10.1080/03610918.2012.667479)
| Detection of lymph node metastases in head and neck cancer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting
### Sempre começar com essas duas linhas para fazer plots
import numpy as np
import matplotlib.pyplot as plt
# If the `x` array is missing, that is, if there is only a single array, as in our example above, the plot function uses `0, 1,..., N-1` for the `x` array, where *N* is the size of the `y` array. Thus, the plot function provides a quick graphical way of examining a data set.
#
plt.plot([1, 2, 3, 2, 3, 4, 3, 4, 5])
plt.show()
x = np.linspace(0, 4. * np.pi, 33)
y = np.sin(x)
plt.plot(x, y)
plt.show()
x = np.linspace(0, 4. * np.pi, 129) # aumentou número de pontos
y = np.sin(x)
plt.plot(x, y)
plt.show()
# !cat wavePulseData.txt
# +
# read data from file
xdata, ydata = np.loadtxt('wavePulseData.txt', unpack=True)
# create x and y arrays for theory
x = np.linspace(-10., 10., 200) # muitos pontos para uma curva smooth
y = np.sin(x) * np.exp(-(x / 5.0)**2)
# create plot
plt.figure(1, figsize=(6, 4)) #'figsize = (width, height)' inches
plt.plot(x, y, 'b-', label='theory')
plt.plot(xdata, ydata, 'ro', label="data") # r = red, o = circle
plt.xlabel('x') #título eixo x
plt.ylabel('transverse displacement') #título eixo y
plt.legend(
loc='upper right') #localização. Legenda com base nas kwargs 'label'
'''axhline() draws a horizontal line across the width of the plot at y=0.
The optional keyword argument color is a string that specifies the color of the
line. The default color is black. The optional keyword argument zorder is an
integer that specifies which plotting elements are in front of or behind others.
By default, new plotting elements appear on top of previously plotted elements
and have a value of zorder=0. By specifying zorder=-1, the horizontal line is
plotted behind all existing plot elements that have not be assigned an explicit
zorder less than -1.
axvline() draws a vertical line from the top to the bottom of the plot at x=0.
See axhline() for explanation of the arguments.'''
plt.axhline(color='gray', zorder=-1)
plt.axvline(color='gray', zorder=-1)
# save plot to file
plt.savefig('WavyPulse.pdf')
# display plot on screen
plt.show()
# -
# ## Format specifiers
# 
#
plt.plot(x, y, 'ro') # plots red circles
plt.show()
plt.plot(x, y, 'ks-') # plot black squares connected by black lines
plt.show()
plt.plot(x, y, 'g^') # plots green triangles that point up
plt.show()
plt.plot(x, y, 'k-') # plots a black line between the points
plt.show()
plt.plot(x, y, 'ms') # plots magenta squares
plt.show()
plt.plot(x,
y,
color='green',
linestyle='dashed',
marker='d',
markerfacecolor='yellow',
markersize=12,
markeredgecolor='blue')
plt.show()
# ## Error bars
# !cat expDecayData.txt
# +
import numpy as np
import matplotlib.pyplot as plt
# read data from file
xdata, ydata, yerror = np.loadtxt('expDecayData.txt', unpack=True)
# create theoretical fitting curve
x = np.linspace(0, 45, 128)
y = 1.1 + 3.0 * x * np.exp(-(x / 10.0)**2)
# create plot
plt.figure(1, figsize=(6, 4))
plt.plot(x, y, 'b-', label="theory")
plt.errorbar(xdata,
ydata,
fmt='ro',
label="data",
xerr=0.75,
yerr=yerror,
ecolor='black')
plt.xlabel('x')
plt.ylabel('transverse displacement')
plt.legend(loc='upper right')
# save plot to file
plt.savefig('ExpDecay.pdf')
# display plot on screen
plt.show()
# -
# ## Setting plotting limits
# +
import numpy as np
import matplotlib.pyplot as plt
theta = np.arange(0.01, 10., 0.04)
ytan = np.tan(theta)
ytanM = np.ma.masked_where(np.abs(ytan) > 20., ytan) # evita linhas verticais
# ligando "infinitos"
plt.figure()
plt.plot(theta, ytanM)
plt.ylim(-8, 8) # restricts range of y axis
plt.axhline(color="gray", zorder=-1) # cria um eixo x horizontal
plt.show()
# -
# ## Subplots
#
# - Figura de cima:
# $$
# \tan\theta \text{ vs } \theta \\
# \sqrt{(8 / \theta)^2-1} \text{ vs } \theta \\
# $$
# As curvas se cruzam onde $\tan\theta = \sqrt{(8 / \theta)^2-1}$.
#
#
# - Figura de baixo:
# $$
# \cot\theta \text{ vs } \theta \\
# -\sqrt{(8 / \theta)^2-1} \text{ vs } \theta \\
# $$
# As curvas se cruzam onde $\cot\theta = -\sqrt{(8 / \theta)^2-1}$.
# +
import numpy as np
import matplotlib.pyplot as plt
theta = np.arange(0.01, 8., 0.04)
y = np.sqrt((8. / theta)**2 - 1.)
ytan = np.tan(theta)
ytan = np.ma.masked_where(np.abs(ytan) > 20., ytan)
ycot = 1. / np.tan(theta)
ycot = np.ma.masked_where(np.abs(ycot) > 20., ycot)
plt.figure(1)
plt.subplot(2, 1, 1) #(rows, columns, position)
plt.plot(theta, y)
plt.plot(theta, ytan)
plt.ylim(-8, 8)
plt.axhline(color="gray", zorder=-1)
plt.axvline(x=np.pi / 2., color="gray", linestyle='--', zorder=-1)
plt.axvline(x=3. * np.pi / 2., color="gray", linestyle='--', zorder=-1)
plt.axvline(x=5. * np.pi / 2., color="gray", linestyle='--', zorder=-1)
plt.xlabel("theta")
plt.ylabel("tan(theta)")
plt.subplot(2, 1, 2)
plt.plot(theta, -y)
plt.plot(theta, ycot)
plt.ylim(-8, 8)
plt.axhline(color="gray", zorder=-1)
plt.axvline(x=np.pi, color="gray", linestyle='--', zorder=-1)
plt.axvline(x=2. * np.pi, color="gray", linestyle='--', zorder=-1)
plt.xlabel("theta")
plt.ylabel("cot(theta)")
plt.show()
# -
# ## Semi-log plots
# +
import numpy as np
import matplotlib.pyplot as plt
# read data from file
time, counts, unc = np.loadtxt('SemilogDemo.txt', unpack=True)
# create theoretical fitting curve
tau = 20.2 # Phosphorus-32 half life = 14 days; tau = t_half/ln(2)
N0 = 8200. # Initial count rate (per second)
t = np.linspace(0, 180, 128)
N = N0 * np.exp(-t / tau)
# creat plot
plt.figure(1, figsize=(10, 4))
plt.subplot(1, 2, 1)
plt.plot(t, N, 'b-', label="theory")
plt.plot(time, counts, 'ro', label="data")
plt.xlabel('time (days)')
plt.ylabel('counts per second')
plt.legend(loc='upper right')
plt.subplot(1, 2, 2)
plt.semilogy(t, N, 'b-', label="theory")
plt.semilogy(time, counts, 'ro', label="data")
plt.xlabel('time (days)')
plt.ylabel('counts per second')
plt.legend(loc='upper right')
plt.tight_layout(
) # adjusts sizes of the plots to make room for the axes labels
#display plot on screen
plt.show()
# -
# ## More advanced graphical output
# +
# Demonstrates the following:
# plotting logarithmic axes
# user-defined functions
# "where" function, NumPy array conditional
import numpy as np
import matplotlib.pyplot as plt
# Define the sinc function, with output for x=0 defined
# as a special case to avoid division by zero
def s(x):
a = np.where(x == 0., 1., np.sin(x) / x)
return a
# create arrays for plotting
x = np.arange(0., 10., 0.1)
y = np.exp(x)
t = np.linspace(-10., 10., 100)
z = s(t)
# create a figure window
fig = plt.figure(1, figsize=(9, 8))
# subplot: linear plot of exponential
ax1 = fig.add_subplot(2, 2, 1)
ax1.plot(x, y) # ax --> axes
ax1.set_xlabel('time (ms)')
ax1.set_ylabel('distance (mm)')
ax1.set_title('exponential')
# subplot: semi-log plot of exponential
ax2 = fig.add_subplot(2, 2, 2)
ax2.plot(x, y)
ax2.set_yscale('log')
ax2.set_xlabel('time (ms)')
ax2.set_ylabel('distance (mm)')
ax2.set_title('exponential')
# subplot: wide subplot of sinc function
ax3 = fig.add_subplot(2, 1, 2)
ax3.plot(t, z, 'r')
ax3.axhline(color='gray')
ax3.axvline(color='gray')
ax3.set_xlabel('angle (deg)')
ax3.set_ylabel('electric field')
ax3.set_title('sinc function')
# Adjusts while space around plots to avoid collisions between subplots
fig.tight_layout()
plt.savefig("MultPlotDemo.pdf")
plt.show()
# +
import numpy as np
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(9, 8))
ax1 = fig.add_subplot(2, 2, 1)
plt.show()
# +
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(223)
ax4 = fig.add_subplot(224)
for ax in [ax1, ax2, ax3, ax4]:
ax.plot([3, 5, 8], [6, 3, 1])
plt.show()
# -
| chap5/chapter_05_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "a13ec6289580145af918a00feb38de76", "grade": false, "grade_id": "cell-e0ef790a27391356", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## **Write your answers directly in the cells on this notebook.**
#
# Use as name of the variables the ones provided as placeholders (often `sol=0`. Delete the `raise NotImplementedError()` line.
#
# ### Show the entire process, from the initial fractions to the final result.
# The probability results must be a decimal value between 0.0 and 1.0. Round your number to the third decimal (e.g 0.574).**
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "e74739a6b924b6e23e1af548c4b59006", "grade": false, "grade_id": "cell-ebda7b65cd913cca", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Consider the following boxes (two boxes (red/blue) with two types of balls (green/orange)):
# 
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "de6a8f795ebc041caae1fb763a767dd7", "grade": false, "grade_id": "cell-abf5d87eb6226183", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Exercise 1: which is the probability of selecting the red box?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "93c7f4b0c233348b2be8ef996352808d", "grade": false, "grade_id": "cell-7969bda5a10dc7ac", "locked": false, "schema_version": 3, "solution": true, "task": false}
sol = 0
# YOUR CODE HERE
Total_no_of_boxes = 2
red_boxes = 1
blue_boxes = 1
Total_no_of_balls = 12
no_of_balls_in_red_boxes = 8
gbr = 2 #green balls in red box
obr = 6 #orange balls in red box
gbb = 3 #green balls in blue box
obb = 1 #orange balls in blue box
tob = 7 #Total no of orange balls in both the boxes
tgb = 5 #Total no of green balls in both the boxes
no_of_balls_in_blue_boxes = 4
sol = red_boxes/Total_no_of_boxes
#raise NotImplementedError()
# + deletable=false nbgrader={"cell_type": "code", "checksum": "8359b7bc05770fd1e1e55d723927ed1b", "grade": true, "grade_id": "cell-2b9f005060121b41", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert sol == 0.5
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "28ffea54b9f99049680e9c788b467a1e", "grade": false, "grade_id": "cell-6deca18d77abc906", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Exercise 2: if you want to run out of balls both boxes at the same time, how often should you select the blue box?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "6a83bfbcf112deeec82137cc84ef1807", "grade": false, "grade_id": "cell-10a736364bdf6ef5", "locked": false, "schema_version": 3, "solution": true, "task": false}
#sol = 0
sol = no_of_balls_in_blue_boxes/Total_no_of_balls
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f1c39a9b2b7ebbdd562454f3051d6920", "grade": true, "grade_id": "cell-d06dc523a3f300ce", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.3333333333333333) < 0.01
# -
# #### Exercise 3: which is the probability of picking up a green ball if you are using the red box?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "701ca27bb44fb1547bb23c5b57d487d4", "grade": false, "grade_id": "cell-335a720d5716ad4b", "locked": false, "schema_version": 3, "solution": true, "task": false}
#sol = 0
sol = gbr/no_of_balls_in_red_boxes
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false nbgrader={"cell_type": "code", "checksum": "aa20451ae4ee5dcde17e766c1df9d9c3", "grade": true, "grade_id": "cell-3cc76e304b24df43", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert sol == 0.25
# -
# #### Exercise 4: which is the probability of picking up an orange ball (any of the boxes)?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "6b031c5732125e59cb486706d217d3e1", "grade": false, "grade_id": "cell-6be260afb8041372", "locked": false, "schema_version": 3, "solution": true, "task": false}
sol = tob/Total_no_of_balls
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false nbgrader={"cell_type": "code", "checksum": "7c0c56c33bacdefb861d8833b26e1e17", "grade": true, "grade_id": "cell-57ee289d4e5df3f0", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.5833333333333334) < 0.01
# -
# #### Exercise 5: given that you picked up a green ball, which is the probability of having selected the red box?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "147c928e559035adb661616e98e13b51", "grade": false, "grade_id": "cell-9c4af4d3ec207bb2", "locked": false, "schema_version": 3, "solution": true, "task": false}
sol = gbr/tgb
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false nbgrader={"cell_type": "code", "checksum": "8e456f643cc92ff22b4178a60b2bf9fc", "grade": true, "grade_id": "cell-10164afff838da6f", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.125) < 0.3
# -
# #### Optional Exercise 6: I have chosen a ball from a box and I picked up a green one. Which is the probability of having used the blue box?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "d37eef12ddf7c25e42efac84fe979775", "grade": false, "grade_id": "cell-ac2e94b60b1ed6fc", "locked": false, "schema_version": 3, "solution": true, "task": false}
# No need to use the Bayes Theorem as it has not been explained in class
sol = gbb/tgb
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d0effe3c87dedc925f42fa749f891a52", "grade": true, "grade_id": "cell-1ca0be55ea9d081c", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.375) < 0.9
# -
# Now consider a standard deck of cards (the one used in Poker):
# 
#
# #### Exercise 7: which is the probability of drawing an Ace?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "c0e38430d25b053c64a6e34cd0ee698a", "grade": false, "grade_id": "cell-f7a947cdf92e17c3", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Create function that returns probability percent rounded to three decimal places
def event_probability(event_outcomes, sample_space):
probability = event_outcomes/sample_space
return probability
sol = event_probability(4,52)
print(round(sol,3))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3a9f2c7239011d2a9aa6b41fee62a2f9", "grade": true, "grade_id": "cell-adb42109bd9b7996", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Using the previously defined function: Determine the probability of drawing an Ace
assert abs(sol-0.077 ) < 0.01
# -
# #### Exercise 8: which is the probability of drawing a card that is a Heart?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "8a33dcb8fbbea5116bd11304098083da", "grade": false, "grade_id": "cell-e261cc3812382a18", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Using the previously defined function: Determine the probability of drawing a heart
sol = event_probability(13,52)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "bbb0458877d5a323e0441efb5d733514", "grade": true, "grade_id": "cell-2bcb2371e498a429", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.25 ) < 0.01
# -
# #### Exercise 9: which is the probability of drawing a face card (such as Jacks, Queens, or Kings)?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "ecbb9fcd5df828494a1215f36a5dc6d2", "grade": false, "grade_id": "cell-fc4fcd733d3e1ca7", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Using the previously defined function: Determine the probability of drawing a face card
sol = event_probability(12,52)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "48c66896a88667f97093e50b05e4e811", "grade": true, "grade_id": "cell-f17c3aa2f33022be", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.231 ) < 0.01
# -
# #### Exercise 10: which is the probability of drawing a face card which is also a Heart?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9669ca6fd185e41f0bd534eef82cd7bf", "grade": false, "grade_id": "cell-c50a860ddb2f18f9", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Using the previously defined function: Determine the probability of drawing the jack, queen or king of hearts
sol = event_probability(3,52)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "eedcc36846411cf784ec8076f41c11ff", "grade": true, "grade_id": "cell-dcd84807c13f2154", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.058 ) < 0.01
# -
# #### Exercise 11: which is the probability of drawing an Ace on the second draw, if the first card drawn was a King?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "234cb4223c601ce08bb7f52d89c70081", "grade": false, "grade_id": "cell-c953e962deacd22d", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Using the previously defined function: Determine the probability of drawing an Ace after drawing a King on the first draw
cards = 52
cards_drawn = 1
Total_cards = cards - cards_drawn
aces = 4
sol = event_probability(aces,Total_cards)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "12ca141c71f705c7b1e5667a892f879c", "grade": true, "grade_id": "cell-c0fd1832c0fa3b01", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.078 ) < 0.01
# -
# #### Exercise 12: which is the probability of drawing an Ace on the second draw, if the first card drawn was an Ace?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "fa3f03da3d2969f08347a8219e30c37a", "grade": false, "grade_id": "cell-899b031f866ba3b2", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Using the previously defined function: Determine the probability of drawing an Ace after drawing an Ace on the first draw
cards = 52
cards_drawn = 1
ace_cards_drawn = 1
Total_cards = cards - cards_drawn
Total_aces = 4
aces = Total_aces-ace_cards_drawn
sol = event_probability(aces,Total_cards)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "e5f9d7129b1a72ed7ee2cd3672e8b99d", "grade": true, "grade_id": "cell-9109c59db8e329c8", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.059) < 0.01
# -
# Now consider the following situation:
#
# You are playing Poker in the [Texas Holdem variant](https://en.wikipedia.org/wiki/Texas_hold_%27em). In case you are not familiar, it is a variant of poker in which each player has two cards and there is a set of community cards to play from.
# 
# #### Exercise 13: what is the probability that the next card drawn will be a Diamond card?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "ec2c2ce83b2175f2a1a9986996fe4398", "grade": false, "grade_id": "cell-17b8aec4013e6b9c", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Sample Space
cards = 52
player_cards = 2
turn_community_cards = 4
available_cards = cards - (player_cards+turn_community_cards)
diamonds = 13
diamonds_drawn = 4
available_diamonds = diamonds - diamonds_drawn
sol = event_probability(available_diamonds,available_cards)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "756e81e0fdc5a32be37a25d0d9c5c250", "grade": true, "grade_id": "cell-ebe4abfecd5e8690", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.196) < 0.01
# -
# Now consider the following situation:
# 
# #### Exercise 14: which is the probability that with the next card drawn there will be five cards in sequential order?
#
# (Notice that any Eight ( 8, 9, 10, Jack, Queen) or any King (9, 10, Jack, Queen, King) will complete the straight)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "fd4058cda5116880eef40aa7789067b1", "grade": false, "grade_id": "cell-f05c41e0e869e186", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Sample Space
cards = 52
player_cards = 2
turn_community_cards = 4
available_cards = cards - (player_cards+turn_community_cards)
available_cards_of_eights_and_kings = 8
sol = event_probability(available_cards_of_eights_and_kings,available_cards)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "fff1dae11dacb0907d324307f22ee39d", "grade": true, "grade_id": "cell-e46db60e9b2dfa4f", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.174) < 0.01
# -
# ## Now, we keep playing Texas Holdem
# 
# ### But looking only at the cards we could have in our hand
# ### (2 possible drawing events, ignore community cards)
# #### Exercise 15: which is the probability of drawing a heart OR drawing a club?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f68ad264c169753b051d3e63ea8d8be0", "grade": false, "grade_id": "cell-5b6c82ce08d8c47f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Calculate the probability of drawing a heart or a club
cards = 52
player_cards = 2
#turn_community_cards = 4
#available_cards = cards - (player_cards)#+turn_community_cards)
print(cards)
hearts = 13
clubs = 13
#sol1 = event_probability(hearts,available_cards)
#sol2 = event_probability(clubs,available_cards)
sol1 = event_probability(hearts+clubs,cards)
sol2 = event_probability(clubs+hearts,cards-1)
sol = 1-(sol1*sol2)
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
sol
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3cc558ba56a2dfe83f6ae2cc1a506fb2", "grade": true, "grade_id": "cell-dedacb0828533e9c", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.755) < 0.01
# -
# #### Exercise 16: which is the probability of drawing an ace, a king or a queen?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "c6932d09078beb0ab84b632362b58892", "grade": false, "grade_id": "cell-0b50174237f37b03", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Calculate the probability of drawing an ace, king, or a queen
cards = 52
player_cards = 2
#turn_community_cards = 4
#available_cards = cards - (player_cards+turn_community_cards)
#print(available_cards)
aces = 4
king = 4
queen = 4
sol1 = event_probability(aces+king+queen,cards)
sol2 = event_probability((king+aces+queen)-1,cards-1)
#sol3 = event_probability((queen+aces+king)-2,cards-2)
sol = sol1*sol2
print(round(sol,3))
# YOUR CODE HERE
#raise NotImplementedError()
sol
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3628cc71fe5948228e7eba7948f5434d", "grade": true, "grade_id": "cell-1c3bec1e6a4f0726", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.41171499999999994) < 0.01
# -
# #### Exercise 17: which is the probability of drawing a heart or an ace?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "5edf1bc7e82a89d77a3ede4255b85997", "grade": false, "grade_id": "cell-3ff5cdeedc739059", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Calculate the probability of drawing a heart or an ace
# YOUR CODE HERE
#raise NotImplementedError()
cards = 52
hearts = 13
aces = 4
ace_of_hearts = 1
sol = event_probability(hearts, cards) + event_probability(aces, cards) - event_probability(ace_of_hearts, cards)
print(round(sol,3))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "67e592333b8ab3fcc55bb59f8651caa3", "grade": true, "grade_id": "cell-a74509b22cdaf4bd", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.525288) < 0.01
# -
# #### Exercise 18: which is the probability of drawing a red card or drawing a face card?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b87522461ae9c12b6c6e7190fcd7674d", "grade": false, "grade_id": "cell-9c43e1f91aac785f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Calculate the probability of drawing a red card or a face card
# YOUR CODE HERE
#raise NotImplementedError()
red_cards = 26
face_cards = 12
red_face_cards = 6
sol = event_probability(red_cards, cards) + event_probability(face_cards, cards) - event_probability(red_face_cards, cards)
print(round(sol, 1))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "82f6de4209429091bcb59c905a5e7db9", "grade": true, "grade_id": "cell-47f891a62515e8d4", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.856395) < 0.01
# -
# #### Exercise 19: which is the probability of drawing an Ace from a deck of cards, replacing it, reshuffling the deck, and drawing another Ace?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "a21f3711e715072fcf4245676d2ef938", "grade": false, "grade_id": "cell-18d5038a32ffeac9", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Sample Space
cards = 52
# Outcomes
aces = 4
ace_probability1 = aces / cards
ace_probability2 = aces / cards
# YOUR CODE HERE
#raise NotImplementedError()
sol = ace_probability1*ace_probability2
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "8eb9ce68fd76ca02e4ffedd850a7e603", "grade": true, "grade_id": "cell-536e3de73c88e083", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.005929) < 0.001
# -
# #### Exercise 20: which is the probability of being dealt two Aces (drawing one Ace after the other starting with a full deck)?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "36f83abcab2e34194ee7339a7687fb5d", "grade": false, "grade_id": "cell-9ec7e543991e2e7c", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Sample Space first draw
cards = 52
# Outcomes first draw
aces = 4
# YOUR CODE HERE
raise NotImplementedError()
sol
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "4dacaae00d1222957a4d7ec6a6c1748e", "grade": true, "grade_id": "cell-0651acc2d4c97ad3", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.004542999999999999) < 0.001
# -
# ### Permutations and combinations
# #### Exercise 21: How many different 5-letter arrangements are there of the letters in the word morse?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "4043edd3b7280831dffe83277d53579d", "grade": false, "grade_id": "cell-5d16f7fa95eb0955", "locked": false, "schema_version": 3, "solution": true, "task": false}
import math
sol = 0
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "2b7ac1eb0e11f7e443cfc6290dcccedd", "grade": true, "grade_id": "cell-290944fdf9fb212b", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert sol == 120
# -
# #### Exercise 22: How many different seven-letter arrangements of the letters in the word HEXAGON can be made if each letter is used only once?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "08fb1d68735a4cc34ff629be615de787", "grade": false, "grade_id": "cell-2b2af5a344e47a8b", "locked": false, "schema_version": 3, "solution": true, "task": false}
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "a1c691147ff054172038dc9d28650cd0", "grade": true, "grade_id": "cell-9eed40b79a9bc42b", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert sol == 5040
# -
# #### Exercise 23: Two cards are drawn at random from a standard deck of cards, without replacement. Find the probability of drawing an 8 and a queen in that order.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "dc1c1dcb93108518e5548db1e0615b0a", "grade": false, "grade_id": "cell-0772318d22f07226", "locked": false, "schema_version": 3, "solution": true, "task": false}
# YOUR CODE HERE
raise NotImplementedError()
sol
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c5c5a1f2cea0046812e93dea9452f7a2", "grade": true, "grade_id": "cell-a85db4f2cf65b50d", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
assert abs(sol-0.006006) < 0.001
| Chapter 1/12. Probability/mt1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:fastscape]
# language: python
# name: conda-env-fastscape-py
# ---
# # Modeling an orogenic system using FastScape: 6. Response to tectonic variation
# 
# +
import numpy as np
import matplotlib.pyplot as plt
#plt.style.use('dark_background')
import xsimlab as xs
import xarray as xr
# %load_ext xsimlab.ipython
from fastscape.models import basic_model
# -
# Finally we are going to see how the source area (the mountain) reacts to cyclic perturbation in uplift rate.
#
# This does not make much sense as tectonics is unlikely to vary in a periodic manner. However, we will apply the *standard* method of computing the response of a system to periodic perturbation to characterize it.
#
# You should go through this notebook step-by-step and try to understand what is being done. It is very similar to the previous notebook so this should not be too difficult. You should also try to run this notebook after changing the period of the climate cycles to see how it affects the system response (in terms of flux amplitude and lag).
spl_model = basic_model.drop_processes('diffusion')
# +
m = 0.4
n = 1
neq = 101
Kf = 1e-5
U = 1e-3
nstep = 201
teq = 1e7
period = 1e8
tfinal = teq + 5*period
tim1 = np.linspace(0,teq,101)
tim2 = np.linspace(teq + period/10, tfinal, 100)
tim = np.concatenate((tim1,tim2))
tecto = 1 + 0.5*np.sin(2*np.pi*(tim-teq)/period)
U_tim = np.where(tim>teq, U*tecto, U)
U_xr = xr.DataArray(data=U_tim, dims='time')
fig, ax = plt.subplots(nrows = 1, ncols = 1, sharex=False, sharey=True, figsize=(12,7))
ax.plot(tim, U_tim)
# +
# # %create_setup spl_model --default --verbose
import xsimlab as xs
ds_in = xs.create_setup(
model=spl_model,
clocks={'time': tim},
input_vars={
# nb. of grid nodes in (y, x)
'grid__shape': [101,101],
# total grid length in (y, x)
'grid__length': [1e5,1e5],
# node status at borders
'boundary__status': ['fixed_value', 'core', 'looped', 'looped'],
# uplift rate
'uplift__rate': U_xr,
# random seed
'init_topography__seed': None,
# bedrock channel incision coefficient
'spl__k_coef': Kf,
# drainage area exponent
'spl__area_exp': m,
# slope exponent
'spl__slope_exp': n,
},
output_vars={'topography__elevation': 'time'}
)
# -
import zarr
zgroup = zarr.group("output.zarr", overwrite=True)
with xs.monitoring.ProgressBar():
ds_out = ds_in.xsimlab.run(model=spl_model, store=zgroup)
# +
from ipyfastscape import TopoViz3d
app = TopoViz3d(ds_out, canvas_height=300, time_dim="time")
app.components['background_color'].set_color('lightgray')
app.components['vertical_exaggeration'].set_factor(5)
app.components['timestepper'].go_to_time(ds_out.time[-1])
app.show()
# -
# The flux of material, $\phi$, coming out of the orogen can be calculated according to:
#
# $$\phi=\frac{U-<\frac{\partial h}{\partial t}>}{U}$$
# +
fig, ax = plt.subplots(nrows=1, ncols=1, sharex=False, sharey=True, figsize=(12, 7))
((ds_out.uplift__rate - ds_out.topography__elevation.differentiate("time").mean(("x", "y"))) / ds_out.uplift__rate
).plot(ax=ax, label="Flux")
(ds_out.uplift__rate / U).plot(ax=ax, label="Tecto")
(ds_out.topography__elevation.mean(("x", "y")) / ds_out.topography__elevation.mean()).plot(ax=ax, label="Topo")
ax.legend()
ax.set_xlim((teq,tfinal))
# -
| notebooks/FastScape_CRC_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # Module 2. GluonTS Training on Amazon SageMaker
# ---
#
# 본 모듈에서는 Amazon SageMaker API를 호출하여 모델 훈련을 수행합니다. 노트북 실행에는 약 10분 가량 소요되며, 핸즈온 실습 시에는 25분을 권장드립니다.
#
# Amazon SageMaker는 완전관리형 머신 러닝 서비스로 인프라 관리에 대해 걱정할 필요가 없으며, 딥러닝 프레임워크의 훈련/배포 컨테이너 이미지를 가져 와서
# 여러분의 스크립트 코드를 쉽게 통합할 수 있습니다.
# <br>
#
# ## 1. Training script
# ---
#
# 아래 코드 셀은 `src` 디렉토리에 SageMaker 훈련 스크립트인 `train.py`를 저장합니다.
# 아래 스크립트가 이전 모듈의 코드와 대부분 일치하다는 점을 알 수 있습니다. 다시 말해, SageMaker 훈련 스크립트 파일은 기존 온프레미스에서 사용했던 Python 스크립트 파일과 크게 다르지 않으며, SageMaker 훈련 컨테이너에서 수행하기 위한 추가적인 환경 변수들만 설정하시면 됩니다.
#
# 환경 변수 설정의 code snippet은 아래과 같습니다.
#
# ```python
# # SageMaker Container environment
# parser.add_argument('--model_dir', type=str, default=os.environ['SM_MODEL_DIR'])
# parser.add_argument('--data_dir', type=str, default=os.environ['SM_CHANNEL_TRAINING'])
# parser.add_argument('--num_gpus', type=int, default=os.environ['SM_NUM_GPUS'])
# parser.add_argument('--output_dir', type=str, default=os.environ.get('SM_OUTPUT_DATA_DIR'))
# ```
# +
# %%writefile ./src/train.py
import os
import pandas as pd
import gluonts
import numpy as np
import argparse
import json
import pathlib
from mxnet import gpu, cpu
from mxnet.context import num_gpus
import matplotlib.pyplot as plt
from gluonts.dataset.util import to_pandas
from gluonts.mx.distribution import DistributionOutput, StudentTOutput, NegativeBinomialOutput, GaussianOutput
from gluonts.model.simple_feedforward import SimpleFeedForwardEstimator
from gluonts.model.deepar import DeepAREstimator
from gluonts.mx.trainer import Trainer
from gluonts.evaluation import Evaluator
from gluonts.evaluation.backtest import make_evaluation_predictions, backtest_metrics
from gluonts.model.predictor import Predictor
from gluonts.dataset.field_names import FieldName
from gluonts.dataset.common import ListDataset
def train(args):
# Parse arguments
epochs = args.epochs
pred_length = args.pred_length
batch_size = args.batch_size
lr = args.lr
model_dir = args.model_dir
data_dir = args.data_dir
num_gpus = args.num_gpus
output_dir = args.output_dir
device = "gpu" if num_gpus > 0 else "cpu"
FREQ = 'H'
target_col = 'traffic_volume'
related_cols = ['holiday', 'temp', 'rain_1h', 'snow_1h', 'clouds_all', 'weather_main', 'weather_description']
# Get training data
target_train_df = pd.read_csv(os.path.join(data_dir, 'target_train.csv'), index_col=0)
related_train_df = pd.read_csv(os.path.join(data_dir, 'related_train.csv'), index_col=0)
num_steps, num_series = target_train_df.shape
target = target_train_df.values
start_train_dt = '2017-01-01 00:00:00'
custom_ds_metadata = {'num_series': num_series,
'num_steps': num_steps,
'prediction_length': pred_length,
'freq': FREQ,
'start': start_train_dt
}
# Prepare GlounTS Dataset
related_list = [related_train_df[c].values for c in related_cols]
train_lst = []
target_vec = target[:-pred_length].squeeze()
related_vecs = [related[:-pred_length].squeeze() for related in related_list]
dic = {FieldName.TARGET: target_vec,
FieldName.START: start_train_dt,
FieldName.FEAT_DYNAMIC_REAL: related_vecs
}
train_lst.append(dic)
test_lst = []
target_vec = target.squeeze()
related_vecs = [related.squeeze() for related in related_list]
dic = {FieldName.TARGET: target_vec,
FieldName.START: start_train_dt,
FieldName.FEAT_DYNAMIC_REAL: related_vecs
}
test_lst.append(dic)
train_ds = ListDataset(train_lst, freq=FREQ)
test_ds = ListDataset(test_lst, freq=FREQ)
# Define Estimator
trainer = Trainer(
ctx=device,
epochs=epochs,
learning_rate=lr,
batch_size=batch_size
)
mlp_estimator = SimpleFeedForwardEstimator(
num_hidden_dimensions=[50],
prediction_length=pred_length,
context_length=2*pred_length,
freq=FREQ,
trainer=trainer
)
# Train the model
mlp_predictor = mlp_estimator.train(train_ds)
# Evaluate trained model on test data
forecast_it, ts_it = make_evaluation_predictions(test_ds, mlp_predictor, num_samples=100)
forecasts = list(forecast_it)
tss = list(ts_it)
evaluator = Evaluator(quantiles=[0.1, 0.5, 0.9])
agg_metrics, item_metrics = evaluator(iter(tss), iter(forecasts), num_series=len(test_ds))
metrics = ['RMSE', 'MAPE', 'wQuantileLoss[0.1]', 'wQuantileLoss[0.5]', 'wQuantileLoss[0.9]', 'mean_wQuantileLoss']
metrics_dic = dict((key,value) for key, value in agg_metrics.items() if key in metrics)
print(json.dumps(metrics_dic, indent=2))
# Save the model
mlp_predictor.serialize(pathlib.Path(model_dir))
return mlp_predictor
def parse_args():
parser = argparse.ArgumentParser()
# Hyperparameter Setting
parser.add_argument('--epochs', type=int, default=10)
parser.add_argument('--pred_length', type=int, default=24*7)
parser.add_argument('--batch_size', type=float, default=32)
parser.add_argument('--lr', type=float, default=0.001)
# SageMaker Container Environment
parser.add_argument('--model_dir', type=str, default=os.environ['SM_MODEL_DIR'])
parser.add_argument('--data_dir', type=str, default=os.environ['SM_CHANNEL_TRAINING'])
parser.add_argument('--num_gpus', type=int, default=os.environ['SM_NUM_GPUS'])
parser.add_argument('--output_dir', type=str, default=os.environ.get('SM_OUTPUT_DATA_DIR'))
args = parser.parse_args()
return args
if __name__ == '__main__':
args = parse_args()
train(args)
# -
# <br>
#
# ## 2. Training
# ---
#
# 스크립트가 준비되었다면 SageMaker 훈련을 수행하는 법은 매우 간단합니다. SageMaker Python SDK 활용 시, Estimator 인스턴스를 생성하고 해당 인스턴스의 `fit()` 메서드를 호출하는 것이 전부입니다. 좀 더 자세히 기술해 보면 아래와 같습니다.
#
# #### 1) Estimator 인스턴스 생성
# 훈련 컨테이너에 필요한 설정들을 지정합니다. 본 핸즈온에서는 훈련 스크립트 파일이 포함된 경로인 소스 경로와(source_dir)와 훈련 스크립트 Python 파일만 엔트리포인트(entry_point)로 지정해 주면 됩니다.
#
# #### 2) `fit()` 메서드 호출
# `estimator.fit(YOUR_TRAINING_DATA_URI)` 메서드를 호출하면, 훈련에 필요한 인스턴스를 시작하고 컨테이너 환경을 시작합니다. 필수 인자값은 훈련 데이터가 존해자는 S3 경로(`s3://`)이며, 로컬 모드로 훈련 시에는 S3 경로와 로컬 경로(`file://`)를 모두 지정할 수 있습니다.
#
# 인자값 중 wait은 디폴트 값으로 `wait=True`이며, 모든 훈련 작업이 완료될 때까지 코드 셀이 freezing됩니다. 만약 다른 코드 셀을 실행하거나, 다른 훈련 job을 시작하고 싶다면 `wait=False`로 설정하여 Asynchronous 모드로 변경하면 됩니다.
#
# **SageMaker 훈련이 끝나면 컨테이너 환경과 훈련 인스턴스는 자동으로 삭제됩니다.** 이 때, SageMaker는 자동으로 `SM_MODEL_DIR` 경로에 저장된 최종 모델 아티팩트를 `model.tar.gz`로 압축하여 훈련 컨테이너 환경에서 S3 bucket으로 저장합니다. 당연히, S3 bucket에 저장된 모델 아티팩트를 다운로드받아 로컬 상에서 곧바로 테스트할 수 있습니다.
# +
import os
import boto3
import sagemaker
from sagemaker.mxnet import MXNet
boto_session = boto3.Session()
sagemaker_session = sagemaker.Session(boto_session=boto_session)
role = sagemaker.get_execution_role()
bucket = sagemaker.Session().default_bucket()
# -
# ### Upload data to Amazon S3
#
# Amazon SageMaker로 모델 훈련을 실행하기 위해, 데이터를 S3에 업로드합니다. 참고로, 로컬 모드에서 테스트 시에는 S3에 업로드할 필요 없이 로컬 상에서도 훈련이 가능합니다.
# +
prefix = 'timeseries-hol/traffic-volume/train'
s3_bucket = boto3.Session().resource('s3').Bucket(bucket)
s3_bucket.Object(os.path.join(prefix, 'target_train.csv')).upload_file('data/target_train.csv')
s3_bucket.Object(os.path.join(prefix, 'related_train.csv')).upload_file('data/related_train.csv')
# -
# ### Local Mode Training
#
# 로컬 모드는 여러분이 작성한 훈련 및 배포 스크립트를 SageMaker에서 관리하는 클러스터에서 실행하기 전에 로컬 상(예: SageMaker 노트북 인스턴스, 개인 랩탑, 온프레미스)에서 여러분의 스크립트가 잘 동작하는지 빠르게 디버깅할 수 있는 방법입니다. 로컬 모드 훈련을 위해서는 docker-compose 또는 nvidia-docker-compose (GPU 인스턴스인 경우)가 필요하지만, SageMaker 노트북 인스턴스는 이미 docker가 설치되어 있습니다.
estimator = MXNet(entry_point='train.py',
source_dir='src',
role=role,
instance_type='local',
instance_count=1,
framework_version='1.6.0',
py_version='py3',
hyperparameters = {'epochs': 2,
'lr': 0.001
}
)
s3_input = sagemaker.inputs.TrainingInput(s3_data='s3://{}/{}'.format(bucket, prefix))
estimator.fit(s3_input)
# 현재 실행 중인 도커 컨테이너가 없는 것을 확인할 수 있습니다.
# !docker ps
# ### SageMaker Hosted Training
#
# 훈련 코드가 로컬에서 잘 작동하므로, 이제 SageMaker에서 관리하는 훈련 인스턴스를 사용하여 훈련을 수행하겠습니다. 로컬 모드 훈련과 달리 호스팅 훈련은
# 노트북 인스턴스 대신에 SageMaker에서 관리하는 별도의 클러스터에서 수행합니다. 본 핸즈온의 데이터셋 사이즈가 작기 때문에 체감이 되지 않겠지만, 대규모 데이터 및 복잡한 모델에 대한 분산 훈련은 SageMaker 호스팅 훈련 방법을 사용하는 것을 권장합니다.
estimator = MXNet(entry_point='train.py',
source_dir='src',
role=role,
train_instance_type='ml.c5.xlarge',
train_instance_count=1,
framework_version='1.6.0',
py_version='py3',
hyperparameters = {'epochs': 20,
'lr': 0.001,
}
)
s3_input = sagemaker.inputs.TrainingInput(s3_data='s3://{}/{}'.format(bucket, prefix))
estimator.fit(s3_input)
# <br>
#
# ## 3. Getting Model Artifacts
# ---
#
# 훈련이 완료된 모델 아티팩트를 로컬(노트북 인스턴스 or 온프레미스)로 복사합니다. 훈련 완료 시 `SM_MODEL_DIR`에 있는 파일들이
# `model.tar.gz`로 자동으로 압축되며, 압축을 해제하여 로컬 상에서도 추론을 수행할 수 있습니다.
local_model_dir = './model'
# !rm -rf $local_model_dir
# +
import json , os
s3_model_dir = estimator.model_data.replace('model.tar.gz', '')
print(s3_model_dir)
# !aws s3 ls {s3_model_dir}
if not os.path.exists(local_model_dir):
os.makedirs(local_model_dir)
# !aws s3 cp {s3_model_dir}model.tar.gz {local_model_dir}/model.tar.gz
# !tar -xzf {local_model_dir}/model.tar.gz -C {local_model_dir}
# -
# 다음 모듈에서 활용할 변수들을 저장합니다.
# %store s3_model_dir
# %store prefix
| traffic-volume/2.training_on_sagemaker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
import tensorflow as tf
def lost(y,A,gamma,w):
return np.linalg.norm(y-A*w)**2+1/2*gamma*np.linalg.norm(w)**2
y=np.ones(2)
A=np.array([[1.0,3.0],[1.0,3.0]])
#w=torch.ones(2)
gamma=0.5
#lost(y,A,gamma,w)
def grad(w):
u=lost(y,A,gamma,w)
u.backward()
return w.grad
#grad(w)
w=torch.ones(2,requires_grad=True)
grad(w)
def f(x):
return (x-5)**2
def grad(x):
y=f(x)
y.backward()
return x.grad
x=torch.ones(1,requires_grad=True)
q=grad(x)
grad(x)-x
x=torch.ones(1,requires_grad=True)
q=x=torch.ones(1,requires_grad=True)
abs(x)>0
w0=torch.ones(1,requires_grad=True)
q=grad(w0)
for i in range(10):
q=w0-grad(w0)
w0=torch.tensor(q,requires_grad=True)
print(q)
w0#-grad(w0)
print(grad(w0))
w1=torch.ones(2,requires_grad=True)
grad(w1)
w0=torch.ones(2)#,requires_grad=True)
w0
np.linalg.norm(w0)
w0=torch.ones(2)#,requires_grad=True)
#w0=torch.ones(2,requires_grad=True)
w0@w0
a=torch.tensor([[1.0,3.0],[1.0,3.0]])
tf.norm(w0,1)+torch.ones(2)
xs = tf.Variable(np.array([[1, 2],[3, 4]]), dtype = tf.float32)
l1 = tf.norm(xs, axis = None)
xs = tf.Variable(np.array([[1, 2],[3, 4]]), dtype = tf.float32)
xs
import pandas as pd
a=np.array([[1,2,3,4,5],[1,2,3,4,5]])
A={'A':'louis','B':'eno','C':'kk'}
B={'Name':'jdh','nom':'kp','nm':'i'}
data=pd.DataFrame(A,B)
data
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="7XDBpzBjlV9X"
# ## Retrieving data from the web
# + colab={} colab_type="code" id="FxcJAU4plV9Y"
# You tell Python that you want to use a library with the import statement.
import requests
# + colab={} colab_type="code" id="X5ezX4D6lV9c"
# Get the HU Wikipedia page
req = requests.get("https://en.wikipedia.org/wiki/Harvard_University")
# + [markdown] colab_type="text" id="Qp5f0iI4lV9f"
# Another very nifty Python function is `dir`. You can use it to list all the properties of an object.
#
# + colab={} colab_type="code" id="fHLQCt4LlV9f"
dir(req)
# + [markdown] colab_type="text" id="3tKlDZhnlV9i"
# Right now `req` holds a reference to a *Request* object; but we are interested in the text associated with the web page, not the object itself.
#
# So the next step is to assign the value of the `text` property of this `Request` object to a variable.
# + colab={} colab_type="code" id="_ALSXUOmlV9j"
page = req.text
page
# + [markdown] colab_type="text" id="EwaIhp4-lV9m"
# # 1) Import BeautifulSoup
# + colab={} colab_type="code" id="UVf4tQVwlV9m"
from bs4 import BeautifulSoup
# + [markdown] colab_type="text" id="YW5H9wQolV9p"
# BeautifulSoup can deal with HTML or XML data, so the next line parser the contents of the `page` variable using its HTML parser, and assigns the result of that to the `soup` variable.
# + [markdown] colab_type="text" id="W4qMWAjjlV9q"
# # 2) Create a Soup variable to store the parsed contents of the page
# + colab={} colab_type="code" id="ETgsZQ54lV9q"
soup=BeautifulSoup(page,'html.parser')
# + [markdown] colab_type="text" id="A44wD93llV9t"
# Let's check the string representation of the `soup` object.
# + colab={} colab_type="code" id="U1Yc1O0wlV9t"
soup_s=str(soup).split()
Is=soup_s
print(Is[50])
# + [markdown] colab_type="text" id="yeiPGaJLlV9w"
# Doesn't look much different from the `page` object representation. Let's make sure the two are different types.
# + colab={} colab_type="code" id="XpB2HMKQlV9w"
type(page)
# + colab={} colab_type="code" id="DXQkdjPYlV9z"
type(soup)
# + [markdown] colab_type="text" id="guVuqnfVlV92"
# Looks like they are indeed different.
# + [markdown] colab_type="text" id="qGosO-26lV92"
# # 3) Display the title of the webpage
# + colab={} colab_type="code" id="zlq9qwTxlV93"
soup.find_all('title')
# + [markdown] colab_type="text" id="IWCsGEtNlV95"
# # 4) Display all p tags from the webpage
# + [markdown] colab_type="text" id="_SrRxjBMlV96"
# #### You may use find_all method!
# + colab={} colab_type="code" id="S-IDFqAblV96"
# Be careful with elements that show up multiple times.
soup.find_all('p')
# + [markdown] colab_type="text" id="O527OjrFlV99"
# # 5) How may p tags are present?
# + colab={} colab_type="code" id="VENpIgJylV99"
len(soup.find_all('p'))
# + [markdown] colab_type="text" id="0e3gFB18lV9_"
# ---
# + colab={} colab_type="code" id="YUbXHoK6lV-A"
soup.table["class"]
# + [markdown] colab_type="text" id="KzoUTUo6lV-C"
# ---### List Comprehensions
# + [markdown] colab_type="text" id="acLP3SvSlV-D"
# # 6) Create a nested list containing classes of all the table tags
# + colab={} colab_type="code" id="RcdwNrL7lV-F"
[ table['class'] for table in soup.find_all('table')]
# + [markdown] colab_type="text" id="fOLdxC_FlV-J"
# As I mentioned, we will be using the Demographics table for this lab. The next cell contains the HTML elements of said table. We will render it in different parts of the notebook to make it easier to follow along the parsing steps.
# + [markdown] colab_type="text" id="oHZQxxdqlV-K"
# # 7) Check the classes and find the Demographics Table
# #### Use find method to find the table using the correct class , convert it into string format and store it in table_html also stored the original form in html_soup
# + colab={} colab_type="code" id="xAFCqdBdlV-K"
table_html = str(soup.find_all('table', class_ = 'wikitable'))
# + colab={} colab_type="code" id="C4DDJmZblV-O"
table_html
# + colab={} colab_type="code" id="1DzOJCEllV-Q"
from IPython.core.display import HTML
HTML(table_html)
# + [markdown] colab_type="text" id="1e3ACBh9lV-T"
# # 8) Extract the rows from the Demographics table and store it in rows variable
# + colab={} colab_type="code" id="e1W00hrLlV-U"
table = soup.find_all('table', class_ = 'wikitable')
print(table[2])
# + colab={} colab_type="code" id="xjBPQa9flV-X"
# Lambda expressions return the value of the expression inside it.
# In this case, it will return a string with new line characters replaced by spaces.
rem_nl = lambda s: s.replace("\n", " ")
# + [markdown] colab_type="text" id="851BgCf6lV-Z"
# # 8) Extract the columns from the Demographics table and store it in columns variable
# + colab={} colab_type="code" id="KWwqgo15lV-a"
headers = table[2].find_all('th')[0:4]
headers = headers[1:]
headers
columns = []
for header in headers:
columns.append(header.text.strip())
print(columns)
# + [markdown] colab_type="text" id="RdYgTEiNlV-b"
# Now let's do the same for the rows. Notice that since we have already parsed the header row, we will continue from the second row. The `[1:]` is a slice notation and in this case it means we want all values starting from the second position.
# + [markdown] colab_type="text" id="RZ4Skq2QlV-c"
# # 9) Extract the indexes from the rows variable
# ### Store it in a variable named indexes
# + colab={} colab_type="code" id="-KH6yroVlV-c"
column = table[2].find_all('th')[4:]
column = column[0:]
indexes = []
for i in column:
indexes.append(i.text.strip())
print(indexes)
# + colab={} colab_type="code" id="AcZcaD1PlV-f"
indexes
# + colab={} colab_type="code" id="GJxoc0kElV-h"
# Here's the original HTML table.
HTML(table_html)
# + [markdown] colab_type="text" id="Ea2avEaklV-k"
# # 10) Convert the percentages to integers
# ### Store it in a variable named values
# + colab={} colab_type="code" id="o4_XUfDIlV-k"
# Your Code Here
data =table[2].findAll('td')
data1 = []
values = []
for i in data:
data1.append(i.getText(strip =True))
for i in data1:
if i[-1] == '%':
values.append(int(i[:-1]))
else:
values.append(None)
values
# + [markdown] colab_type="text" id="0in8D_gdlV-p"
# The problem with the list above is that the values lost their grouping.
#
# The `zip` function is used to combine two sequences element wise. So `zip([1,2,3], [4,5,6])` would return `[(1, 4), (2, 5), (3, 6)]`.
#
# This is the first time we see a container bounded by parenthesis. This is a tuple, which you can think of as an immutable list (meaning you can't add, remove, or change elements from it). Otherwise they work just like lists and can be indexed, sliced, etc.
# + colab={} colab_type="code" id="9U6avqTylV-q"
c1=[values[i] for i in range(0,len(values),3)]
c2=[values[i] for i in range(1,len(values),3)]
c3=[values[i] for i in range(2,len(values),3)]
stacked_values = list(zip(c1,c2,c3))
stacked_values
# + colab={} colab_type="code" id="5DkiF6GelV-s"
# Here's the original HTML table.
HTML(table_html)
# + colab={} colab_type="code" id="Hf3F1YKRlV-v"
import pandas as pd
# + [markdown] colab_type="text" id="Z7_L6B71lV-x"
# # 11) Create the DataFrame
# ### Use stacked_values, columns and indexes to create the Demographics DataFrame
# #### Name the DataFrame df
# + colab={} colab_type="code" id="fYG5DvVBlV-x"
# Your Code Here
df=pd.DataFrame(stacked_values,index=indexes,columns=columns)
df.head()
#df['Undergrad'].corr(df['Grad/prof'])
df['Undergrad'].corr(df['US census'])
#df_clean.mean(axis=0)
# + colab={} colab_type="code" id="bWEzcjnilV-z"
# Here's the original HTML table.
HTML(table_html)
# + [markdown] colab_type="text" id="F8iqHSV1lV-2"
# ---
#
# ### DataFrame cleanup
#
# Our DataFrame looks nice; but does it have the right data types?
# + [markdown] colab_type="text" id="bnZ8vzR5lV-2"
# # 12) Display the datatypes of all the columns
# + colab={} colab_type="code" id="6PjFH677lV-3"
# Your Code Here
df.dtypes
# + [markdown] colab_type="text" id="qxBtq4QXlV-7"
# # 13) Drop the row containing NaN value.
# ### After droping the row store it in df_clean_row
# + colab={} colab_type="code" id="agY6jks7lV-9"
# Your Code Here
df.dropna(axis = 0)
# + [markdown] colab_type="text" id="ka5mJhBmlV-_"
# # 13) Drop the column containing NaN value.
# ### After droping the row store it in df_clean_column
# + colab={} colab_type="code" id="VoRXibnVlV-_"
# Your Code Here
df.dropna(axis = 1)
# + [markdown] colab_type="text" id="nrOc2WkxlV_B"
# We will take a less radical approach and replace the missing value with a zero. In this case this solution makes sense, since 0% value meaningful in this context. We will also transform all the values to integers at the same time.
# + [markdown] colab_type="text" id="dleQ3exZlV_B"
# # 13) Fill the NaN value with 0
# ### After filling the NaN value with 0 store it in df_clean
# + colab={} colab_type="code" id="iE-C7eA1lV_B"
# Your Code Here
df_clean = df.fillna(0)
# + colab={} colab_type="code" id="Bg9w39lplV_D"
df_clean.dtypes
# + [markdown] colab_type="text" id="psKN1wZ6lV_F"
# Now our table looks good!
#
# + colab={} colab_type="code" id="jUep-9NllV_G"
import numpy as np
# + [markdown] colab_type="text" id="lxkkHR_3lV_I"
# The `values` method of the DataFrame will return a two-dimensional `array` with the DataFrame values. The `array` is a NumPy structure that we will be using a lot during this class.
# + colab={} colab_type="code" id="DG0k2_UVlV_I"
df_clean.values
# + [markdown] colab_type="text" id="Fi-YMfJ5lV_K"
# Let's see if this is indeed a NumPy type...
# + colab={} colab_type="code" id="Y99gwFjKlV_K"
type(df_clean.values)
# + [markdown] colab_type="text" id="3t1WCFI_lV_M"
# NumPy also offers many functions that can operate directly on the DataFrame.
# + [markdown] colab_type="text" id="gfRSm1AolV_N"
# # 14) Find the mean for the column 'Undergrad' from the cleaned dataset
# + colab={} colab_type="code" id="4Ae8SICFlV_N"
# Your Code Here
print(df['Undergrad'].mean())
print(df['Grad/prof'].min())
print(df['Grad/prof'].max())
X = df_clean/(df_clean.sum()).max()
df_clean/df_clean.sum()
# + [markdown] colab_type="text" id="U1aE66TMlV_O"
# # 15) Find the standard deviation for all the columns of the cleaned dataset
# + colab={} colab_type="code" id="6GOMSOKnlV_R"
# Your Code Here
df_clean.std()
| Web_scrapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#rdkit関連のimport
from rdkit import Chem
from rdkit.Chem import PandasTools
#randamforest
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
#その他のimport
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import glob
import re
#データの規格化
def rescaling(features):
norm_features = []
max_value = max(features)
min_value = min(features)
for feature in features:
norm_feature = (feature - min_value)/(max_value - min_value)
norm_features.append(norm_feature)
return norm_features
#SDFファイルの読み込み
mol_list = Chem.SDMolSupplier("../../ForMolPredict/SDF_files/SOL/SOL_AllMOL.sdf",removeHs=False)
mol_num = len(mol_list)
print("there are {} molecules".format(mol_num))
# -
#データの抽出
SOL_class_list = [mol.GetProp('SOL_class') for mol in mol_list]
mol_props = ['Volume', 'Energy', 'HOMO', 'LUMO', 'HLgap', 'Mcharge_ave', 'Mcharge_var', 'Lcharge_ave', 'Lcharge_var', 'dipole', 'Atom_num', 'Mass', 'Density']
rdkit_fp = []
calc_list = []
for mol in mol_list:
fp = [x for x in Chem.RDKFingerprint(mol)]
rdkit_fp.append(fp)
mol_data = [mol.GetDoubleProp(prop) for prop in mol_props]
calc_list.append(mol_data)
FP_df = pd.DataFrame(rdkit_fp)
Calc_df = pd.DataFrame(calc_list, columns=mol_props)
FC_df = pd.concat([Calc_df, FP_df], axis=1 )
#Rescaling
resc_list = []
for prop_name, prop_data in Calc_df.iteritems():
resc_list.append(rescaling(prop_data))
rescCalc_df = pd.DataFrame(resc_list, index=mol_props)
rescCalc_df = rescCalc_df.T
rescFC_df = pd.concat([rescCalc_df, FP_df], axis=1 )
print('number of Fingerprint:',len(FP_df.columns))
print('number of Calclation Data:', len(Calc_df.columns))
print('number of All Data:', len(FC_df.columns))
print('number of Rescaled Calclation Data:', len(rescCalc_df.columns))
print('number of Rescaled All Data:', len(rescFC_df.columns))
#activityのカウント
print('number of (A) low: ',SOL_class_list.count('(A) low'))
print('number of (B) medium: ',SOL_class_list.count('(B) medium'))
print('number of (C) high: ',SOL_class_list.count('(C) high'))
#Only Rdkit Finger Print
X_train, X_test, y_train, y_test = train_test_split(FP_df, SOL_class_list, random_state=0)#test_size=0.25(default)
forest = RandomForestClassifier(max_depth=100, n_estimators=500, random_state=0)
forest.fit(X_train, y_train)
y_pred = forest.predict(X_test) #予測値算出
print('=======Only Rdkit Finger Print========')
print('train accuracy: {:.3f}'.format(forest.score(X_train, y_train)))
print('accuracy: {:.3f}'.format(accuracy_score(y_test, y_pred)))
#Only Calculation Results
X2_train, X2_test, y2_train, y2_test = train_test_split(Calc_df, SOL_class_list, random_state=0)
forest2 = RandomForestClassifier(max_depth=100, n_estimators=500, random_state=0)
forest2.fit(X2_train, y2_train)
y2_pred = forest2.predict(X2_test) #予測値算出
print('=======Only Calculation Results========')
print('train accuracy: {:.3f}'.format(forest2.score(X2_train, y2_train)))
print('accuracy: {:.3f}'.format(accuracy_score(y2_test, y2_pred)))
#Calculation Results + Rdkit Finger Print
X3_train, X3_test, y3_train, y3_test = train_test_split(FC_df, SOL_class_list, random_state=0)
forest3 = RandomForestClassifier(max_depth=100, n_estimators=500, random_state=0)
forest3.fit(X3_train, y3_train)
y3_pred = forest3.predict(X3_test) #予測値算出
print('=======Calculation Results + Rdkit Finger Print========')
print('train accuracy: {:.3f}'.format(forest3.score(X3_train, y3_train)))
print('test accuracy: {:.3f}'.format(accuracy_score(y3_test, y3_pred)))
print('/////////////Rescaled Results//////////////')
#Only Calculation Results
X4_train, X4_test, y4_train, y4_test = train_test_split(rescCalc_df, SOL_class_list, random_state=0)
forest4 = RandomForestClassifier(max_depth=100, n_estimators=500, random_state=0)
forest4.fit(X4_train, y4_train)
y4_pred = forest4.predict(X4_test) #予測値算出
print('=======Only Calculation Results========')
print('train accuracy: {:.3f}'.format(forest4.score(X4_train, y4_train)))
print('test accuracy: {:.3f}'.format(accuracy_score(y4_test, y4_pred)))
#Calculation Results + Rdkit Finger Print
X5_train, X5_test, y5_train, y5_test = train_test_split(rescFC_df, SOL_class_list, random_state=0)
forest5 = RandomForestClassifier(max_depth=100, n_estimators=500, random_state=0)
forest5.fit(X5_train, y5_train)
y5_pred = forest3.predict(X5_test) #予測値算出
print('=======Calculation Results + Rdkit Finger Print========')
print('train accuracy: {:.3f}'.format(forest5.score(X5_train, y5_train)))
print('test accuracy: {:.3f}'.format(accuracy_score(y5_test, y5_pred)))
#activityごとの評価
print('=======Only Rdkit Finger Print========')
print(classification_report(y_test,y_pred))
print('=======Only Calculation Results========')
print(classification_report(y2_test,y2_pred))
print('=======Calculation Results + Rdkit Finger Print========')
print(classification_report(y3_test,y3_pred))
print('/////////////Rescaled Results//////////////')
print('=======Only Calculation Results========')
print(classification_report(y4_test,y4_pred))
print('=======Calculation Results + Rdkit Finger Print========')
print(classification_report(y5_test,y5_pred))
#confusion matrix
print('=======Only Rdkit Finger Print========')
matrix_fp = confusion_matrix(y_test, y_pred)
sns.heatmap(matrix_fp, square=True, annot=True, cbar=False, fmt='d', cmap='RdPu')
plt.xlabel('solubility')
plt.ylabel('true value')
#confusion matrix
print('=======Only Calculation Results========')
matrix_calc = confusion_matrix(y2_test, y2_pred)
sns.heatmap(matrix_calc, square=True, annot=True, cbar=False, fmt='d', cmap='RdPu')
plt.xlabel('solubility')
plt.ylabel('true value')
#confusion matrix
print('=======Calculation Results + Rdkit Finger Print========')
matrix_FC = confusion_matrix(y3_test, y3_pred)
sns.heatmap(matrix_FC, square=True, annot=True, cbar=False, fmt='d', cmap='RdPu')
plt.xlabel('solubility')
plt.ylabel('true value')
#confusion matrix
print('=======Only Rescaled Calculation Results========')
matrix_calc = confusion_matrix(y4_test, y4_pred)
sns.heatmap(matrix_calc, square=True, annot=True, cbar=False, fmt='d', cmap='RdPu')
plt.xlabel('solubility')
plt.ylabel('true value')
#confusion matrix
print('=======Rescaled Calculation Results + Rdkit Finger Print========')
matrix_FC = confusion_matrix(y5_test, y5_pred)
sns.heatmap(matrix_FC, square=True, annot=True, cbar=False, fmt='d', cmap='RdPu')
plt.xlabel('solubility')
plt.ylabel('true value')
# Feature Importance
fti = forest.feature_importances_
fti2 = forest2.feature_importances_
fti3 = forest3.feature_importances_
fti4 = forest4.feature_importances_
fti5 = forest5.feature_importances_
fti_df = pd.DataFrame(fti, columns = ['importance'])
fti_Calc_index = mol_props
fti_FC_index = mol_props+list(fti_df.index)
fti2_df = pd.DataFrame(fti2, columns = ['importance'], index = fti_Calc_index)
fti3_df = pd.DataFrame(fti3, columns = ['importance'], index = fti_FC_index)
fti4_df = pd.DataFrame(fti4, columns = ['importance'], index = fti_Calc_index)
fti5_df = pd.DataFrame(fti5, columns = ['importance'], index = fti_FC_index)
#sort
fti_df = fti_df.sort_values("importance", ascending=False)
fti2_df = fti2_df.sort_values("importance", ascending=False)
fti3_df = fti3_df.sort_values("importance", ascending=False)
fti4_df = fti4_df.sort_values("importance", ascending=False)
fti5_df = fti5_df.sort_values("importance", ascending=False)
print('=======Only Rdkit Finger Print========')
plt.figure()
fti_df[:20].plot.bar() #Top20
print('=======Only Calculation Results========')
plt.figure()
fti2_df[:20].plot.bar() #Top20
print('=======Calculation Results + Rdkit Finger Print========')
plt.figure()
fti3_df[:20].plot.bar() #Top20
print('=======Only Rescaled Calculation Results========')
plt.figure()
fti4_df[:20].plot.bar() #Top20
print('=======Rescaled Calculation Results + Rdkit Finger Print========')
plt.figure()
fti5_df[:20].plot.bar() #Top20
| RandomForest/Sol_SDF_3RandomForest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Propagación hacia atrás
#
# En esta sección vamos a crear nuestra primera red neuronal capaz de aprender.
# ## Cargamos los datos
# Para hacer un ejemplo relativamente sencillo, vamos a resolver un problema de clasificación binaria usando el dataset del Cancer de Mama, que ya hemos visto anteriormente. Recordemos que es un dataset donde las variables independientes son medidas hechas de una imágen de un posible tumor y la variable objetivo es un 0 (cancer maligno) o un 1 (cancer benigno)
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (6, 6)
# +
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
X, y = data.data, data.target
# -
# Usamos solo las 4 primeras variables para que el input coincida con la capa de entrada de la red
X = data.data[:,:4]
X.shape
# Para entrenar una red neuronal es muy importante que los datos estén normalizados, si no lo son la red le va a dar más importancia a aquellas variables cuyo rango sea más grande.
from sklearn.preprocessing import StandardScaler
x_estandardizador = StandardScaler()
X_std = x_estandardizador.fit_transform(X)
x0 = X_std[0]
y0 = y[0]
print(x0, y0)
# ### Creación de la red neuronal
# E implementaremos el algoritmo de propagación hacia atrás (backpropagation) que es lo que permitirá que la red aprenda.
# Es decir, una capa de entrada con 4 neuronas (*también llamadas **unidades**^), una capa oculta con 5 neuronas y una capa de salida que convertirá los outputs (o **activaciones** de la capa oculta en clase positiva o negativa
# En primer lugar definimos las funciones de activación distintas:
# - ** Función identidad**, que se usa en la capa de entrada y no hace nada (o sea, $f(x)=x$)
# - **Función Sigmoide**, que aplica la función sigmoide $f(x)=\frac{1}{1+e^{-x}}$ que convierte los números al rango `[0,1]` y que se usa para problemas de clasificación binaria
# +
def fn_identidad(x, derivada=False):
if derivada:
return np.ones(x.shape)
return x
def fn_sigmoide(x, derivada=False):
if derivada:
return x*(1-x)
return 1/(1+np.exp(-x))
# -
# también tenemos que definir una manera de computar el error de una predicción (error, coste y perdida se usan indistintamente).
# Para un problema de clasificación binaria, una buena métrica es la pérdida logarítmica ([logloss](http://wiki.fast.ai/index.php/Log_Loss))
def error_logloss(y_pred, y):
p = np.clip(y_pred, 1e-15, 1 - 1e-15)
if y == 1:
return -np.log(p)
else:
return -np.log(1 - p)
# En primer lugar, definimos la capa básica, que tiene un número de unidades un bias, y una función de activación
class Layer:
def __init__(self, n_unidades, fn_activacion, bias=True):
self.n_unidades = n_unidades
self.fn_activacion = fn_activacion
self.dim_output = n_unidades
# añadimos un peso más para la unidad de bias
self.bias = bias
self.dimensiones = "no generada"
self.w = None
def __repr__(self):
return """
Capa {}. dimensiones = {}.
pesos: {}
""".format(
self.nombre, self.dimensiones, self.w)
def generar_pesos(self, dim_output_anterior):
if self.bias:
self.dimensiones = (self.n_unidades, dim_output_anterior+1)
else:
self.dimensiones = (self.n_unidades, dim_output_anterior)
self.w = np.random.random(self.dimensiones)
def add_bias(self, x):
if not self.bias:
return x
x_con_bias_1d = np.append(1, x)
# append convierte en array 1dimensional necesitamos 2d
return x_con_bias_1d.reshape(
x_con_bias_1d.shape[0], 1
)
def activar(self, x):
x_con_bias_2d = self.add_bias(x)
return self.fn_activacion( self.w @ x_con_bias_2d )
def calcular_delta(self, producto_capa, output_capa):
return producto_capa * self.fn_activacion(output_capa, derivada=True)
# Tenemos 3 tipos de Capas de neuronas.
#
# - **Capa de Entrada**, no hace nada, simplemente conecta el input con el resto de la red
# - **Capa Oculta**, también llamada capa densa, realiza el algoritmo perceptrón con una función de activación no lineal
# - **Capa de Salida**, esta capa traduce el output de la capa antepenúltima a la variable objetivo deseada
# +
class InputLayer(Layer):
nombre = "entrada"
def generar_pesos(self):
pass
def activar(self, x):
return x
class HiddenLayer(Layer):
nombre = "oculta"
class OutputLayer(Layer):
nombre = "salida"
# -
# Ahora creamos la red neuronal, que es simplemente una lista de capas y con capacidad de hacer propagación hacia delante y hacia atrás.
class RedNeuronal:
def __init__(self, ratio_aprendizaje, fn_error):
self.layers = []
self.ratio_aprendizaje = ratio_aprendizaje
self.fn_error = fn_error
def add_layer(self, layer):
if layer.nombre == "entrada":
layer.generar_pesos()
else:
layer.generar_pesos(self.layers[-1].dim_output)
self.layers.append(layer)
def __repr__(self):
info_red = ""
for layer in self.layers:
info_red += "\nCapa: {} Nº unidades: {}".format(
layer.nombre, layer.n_unidades)
return info_red
def forward(self, x):
for layer in self.layers:
layer.input = layer.add_bias(x).T
x = layer.activar(x)
layer.output = x
return x
def calcular_error_prediccion(self, y_pred, y):
return self.fn_error(y_pred, y)
def backward(self, y_pred, y):
# El error de prediccion final
delta_capa = self.calcular_error_prediccion(y_pred, y)
for layer in reversed(self.layers):
if layer.nombre == "entrada":
continue
if layer.nombre == "salida":
producto_capa = delta_capa @ layer.w
else:
#quitamos el error del bias de la capa anterior
producto_capa = delta_capa[:,1:] @ layer.w
delta_capa = layer.calcular_delta(producto_capa, layer.output)
layer.delta = delta_capa
def actualizar_pesos(self):
"""
Actualiza pesos mediante el descenso de gradiente"""
for layer in self.layers[1:]:
layer.w = layer.w - self.ratio_aprendizaje \
*layer.delta * layer.input
def aprendizaje(self, x, y):
"""
Función principal para entrenar la red
"""
y_pred = self.forward(x)
self.backward(y_pred, y)
self.actualizar_pesos()
error_prediccion = self.calcular_error_prediccion(y_pred, y)
return error_prediccion
def predict_proba(self, x):
return self.forward(x)
def predict(self, x):
probabilidad = self.predict_proba(x)
if probabilidad>=0.5:
return 1
else:
return 0
# #### Creación de la red neuronal
# En primer lugar tenemos que definir los tamaños de cada capa, y si van a incluir sesgo (bias) o no.
# +
n_input = 4
n_oculta = 5
n_output = 1
RATIO_APRENDIZAJE = 0.0001
N_ITERACIONES=1000
# +
red_sigmoide = RedNeuronal(ratio_aprendizaje=RATIO_APRENDIZAJE, fn_error=error_logloss)
red_sigmoide.add_layer(InputLayer(n_input, bias=False, fn_activacion=fn_identidad))
red_sigmoide.add_layer(HiddenLayer(n_oculta, fn_activacion=fn_sigmoide))
red_sigmoide.add_layer(OutputLayer(n_output, fn_activacion=fn_sigmoide))
# -
# Inicialmente la red tiene unos pesos aleatorios
red_sigmoide.layers
# Si ahora hacemos una iteración del proceso de aprendizaje:
red_sigmoide.aprendizaje(x0, y0)
# Vemos que los pesos de las capas se han actualizado
red_sigmoide.layers
# Esto es el equivalente a hacer los siguientes pasos:
prediccion = red_sigmoide.forward(x0)
prediccion
red_sigmoide.backward(prediccion, y0)
red_sigmoide.actualizar_pesos()
red_sigmoide.layers
# Ya tenemos una red neuronal que aprende para optimizar una observación usando el método del descenso de gradiente. Ahora solo tenemos que implementar el método de descenso estocástico de gradiente (SGD) para iterar en todo el dataset de entrenamiento e ir modificando los pesos para minimizar los errores de entrenamiento
# +
def iteracion_sgd(red, X, y):
# barajamos los datos de entrenamiento
indice_aleatorio = np.random.permutation(X.shape[0])
error = []
# iteramos todo el dataset
for i in range(indice_aleatorio.shape[0]):
x0 = X[indice_aleatorio[i]]
y0 = y[indice_aleatorio[i]]
err = red.aprendizaje(x0, y0)
error.append(err)
return np.nanmean(np.array(error))
def entrenar_sgd(red, n_epocas, X, y):
epocas = []
for epoca in range(n_epocas):
error_epoca = iteracion_sgd(red, X, y)
epocas.append([epoca, error_epoca])
return np.array(epocas)
# -
# Ahora por ejemplo corremos el algoritmo durante varias iteraciones.
resultados_sigmoide = entrenar_sgd(red_sigmoide, N_ITERACIONES, X_std, y)
# Si ahora visualizamos la evolución del error medio
plt.scatter(x=resultados_sigmoide[:,0], y=resultados_sigmoide[:,1])
plt.title("Error para red con funcion sigmoide en capa oculta")
plt.xlabel("Número de Iteraciones")
plt.ylabel("Error medio");
# Vemos que a cada iteración (época) de aprendizaje el error medio total se va reduciendo
# Lo bueno de las redes neuronales es que tienen una flexibilidad que otros modelos no tienen.
#
# Por ejemplo podemos cambiar la función de activación de la capa oculta.
#
# En la práctica la función sigmoide no se usa para capas ocultas, se suele usar más la **Unidad Linear rectificada (ReLU)**.
# +
def fn_relu(x, derivada=False):
if derivada:
return 1. * (x>0.)
return np.maximum(x, 0.)
def fn_leakyrelu(x, derivada=False):
if derivada:
if x.any()>0:
return 1.
else:
return 0.01
return np.maximum(x, 0.01*x)
# -
# o incluso modificar la red y añadir otra capa con el doble de unidades
red_relu = RedNeuronal(ratio_aprendizaje=RATIO_APRENDIZAJE, fn_error=error_logloss)
red_relu.add_layer(InputLayer(n_input, bias=False, fn_activacion=fn_identidad))
red_relu.add_layer(HiddenLayer(n_oculta, fn_activacion=fn_relu))
red_relu.add_layer(HiddenLayer(n_oculta, fn_activacion=fn_relu))
red_relu.add_layer(OutputLayer(n_output, fn_activacion=fn_sigmoide))
resultados_relu = entrenar_sgd(red_relu, N_ITERACIONES, X_std, y)
plt.scatter(x=resultados_relu[:,0], y=resultados_relu[:,1])
plt.title("Error para red con funcion ReLU en capa oculta");
| Deep_Learning/3_Propagacion_Atrás.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="H1In7nWTdKQg" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2fee8b07-3fea-4674-a04d-abbe409c2d8b"
import numpy
import numpy as np
import numpy.random as rn
import matplotlib.pyplot as plt # to plot
import matplotlib as mpl
from scipy import optimize # to compare
'''
DESCRIPTION
Calculates Annual Energy Production (AEP) of a Wind Farm
============================================================
This is vectorzied version of Farm_Evalautor.py.
Farm_Evalautor_Vec.py is a python file that calculates AEP (GWh)
of a certain arrangement of wind turbines in a farm, under
given annual wind conditions.
The code in this script for wake-effect modeling is based on
standard Jensen (PARK) model.
PACKAGE LIST
You may need to install the package Shapely in your
python distribution. These are not pre-installed.
=============================================================
Packages Used:
Numpy
Pandas
Shapely
math (built-in)
OPTIMIZATION USAGE
This vectorized version is faster than unvectorized version
Farm_Evalautor.py. Due to speed benefits, we advise you to use
the function getAEP in this script while developing the optimizer.
=============================================================
One way to use getAEP function as AEP evaluator while optimizing is:
- STEP 1. Import the relevant function from Farm_Evalautor_Vec.
from Farm_Evalautor_Vec import getTurbLoc, loadPowerCurve,
binWindResourceData, preProcessing, getAEP
- STEP 2. Set Turbine Radius to 50.0. First arg of getAEP
- STEP 3. Load Turbine Locations. Using function getTurbLoc
- STEP 4. Load Power Curve. Using function loadPowerCurve
- STEP 5. Load wind instance probabilities.
Using function binWindResourceData
- STEP 6. Perform Preprocessing by calling function preProcessing.
We do preprocessing to avoid same repeating calculations.
Do them once.
- STEP 7. Finally, call function getAEP
This makes it easy to control the reloading of data and hence achieve
better performance. '''
# Module List
import numpy as np
import pandas as pd
from math import radians as DegToRad # Degrees to radians Conversion
from shapely.geometry import Point # Imported for constraint checking
from shapely.geometry.polygon import Polygon
#from geneticalgorithm import geneticalgorithm as ga
import warnings
warnings.filterwarnings("ignore")
def getTurbLoc(turb_loc_file_name):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Returns x,y turbine coordinates
:Called from
main function
:param
turb_loc_file_name - Turbine Loc csv file location
:return
2D array
"""
df = pd.read_csv(turb_loc_file_name, sep=',', dtype = np.float32)
turb_coords = df.to_numpy(dtype = np.float32)
return(turb_coords)
def loadPowerCurve(power_curve_file_name):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Returns a 2D numpy array with information about
turbine thrust coeffecient and power curve of the
turbine for given wind speed
:called_from
main function
:param
power_curve_file_name - power curve csv file location
:return
Returns a 2D numpy array with cols Wind Speed (m/s),
Thrust Coeffecient (non dimensional), Power (MW)
"""
powerCurve = pd.read_csv(power_curve_file_name, sep=',', dtype = np.float32)
powerCurve = powerCurve.to_numpy(dtype = np.float32)
return(powerCurve)
def binWindResourceData(wind_data_file_name):
r"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Loads the wind data. Returns a 2D array with shape (36,15).
Each cell in array is a wind direction and speed 'instance'.
Values in a cell correspond to probability of instance
occurence.
:Called from
main function
:param
wind_data_file_name - Wind Resource csv file
:return
1-D flattened array of the 2-D array shown below. Values
inside cells, rough probabilities of wind instance occurence.
Along: Row-direction (drct), Column-Speed (s). Array flattened
for vectorization purpose.
|0<=s<2|2<=s<4| ... |26<=s<28|28<=s<30|
|_____________|______|______|______|________|________|
| drct = 360 | -- | -- | -- | -- | -- |
| drct = 10 | -- | -- | -- | -- | -- |
| drct = 20 | -- | -- | -- | -- | -- |
| .... | -- | -- | -- | -- | -- |
| drct = 340 | -- | -- | -- | -- | -- |
| drct = 350 | -- | -- | -- | -- | -- |
"""
# Load wind data. Then, extracts the 'drct', 'sped' columns
df = pd.read_csv(wind_data_file_name)
wind_resource = df[['drct', 'sped']].to_numpy(dtype = np.float32)
# direction 'slices' in degrees
slices_drct = np.roll(np.arange(10, 361, 10, dtype=np.float32), 1)
## slices_drct = [360, 10.0, 20.0.......340, 350]
n_slices_drct = slices_drct.shape[0]
# speed 'slices'
slices_sped = [0.0, 2.0, 4.0, 6.0, 8.0, 10.0, 12.0, 14.0, 16.0,
18.0, 20.0, 22.0, 24.0, 26.0, 28.0, 30.0]
n_slices_sped = len(slices_sped)-1
# placeholder for binned wind
binned_wind = np.zeros((n_slices_drct, n_slices_sped),
dtype = np.float32)
# 'trap' data points inside the bins.
for i in range(n_slices_drct):
for j in range(n_slices_sped):
# because we already have drct in the multiples of 10
foo = wind_resource[(wind_resource[:,0] == slices_drct[i])]
foo = foo[(foo[:,1] >= slices_sped[j])
& (foo[:,1] < slices_sped[j+1])]
binned_wind[i,j] = foo.shape[0]
wind_inst_freq = binned_wind/np.sum(binned_wind)
wind_inst_freq = wind_inst_freq.ravel()
return(wind_inst_freq)
def searchSorted(lookup, sample_array):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Returns lookup indices for closest values w.r.t sample_array elements
:called_from
preProcessing, getAEP
:param
lookup - The lookup array
sample_array - Array, whose elements need to be matched
against lookup elements.
:return
lookup indices for closest values w.r.t sample_array elements
"""
lookup_middles = lookup[1:] - np.diff(lookup.astype('f'))/2
idx1 = np.searchsorted(lookup_middles, sample_array)
indices = np.arange(lookup.shape[0])[idx1]
return indices
def preProcessing(power_curve):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Doing preprocessing to avoid the same repeating calculations.
Record the required data for calculations. Do that once.
Data are set up (shaped) to assist vectorization. Used later in
function totalAEP.
:called_from
main function
:param
power_curve - 2D numpy array with cols Wind Speed (m/s),
Thrust Coeffecient (non dimensional), Power (MW)
:return
n_wind_instances - number of wind instances (int)
cos_dir - For coordinate transformation
2D Array. Shape (n_wind_instances,1)
sin_dir - For coordinate transformation
2D Array. Shape (n_wind_instances,1)
wind_sped_stacked - column staked all speed instances n_turb times.
C_t - 3D array with shape (n_wind_instances, n_turbs, n_turbs)
Value changing only along axis=0. C_t, thrust coeff.
values for all speed instances.
"""
# number of turbines
n_turbs = 50
# direction 'slices' in degrees
slices_drct = np.roll(np.arange(10, 361, 10, dtype=np.float32), 1)
## slices_drct = [360, 10.0, 20.0.......340, 350]
n_slices_drct = slices_drct.shape[0]
# speed 'slices'
slices_sped = [0.0, 2.0, 4.0, 6.0, 8.0, 10.0, 12.0, 14.0, 16.0,
18.0, 20.0, 22.0, 24.0, 26.0, 28.0, 30.0]
n_slices_sped = len(slices_sped)-1
# number of wind instances
n_wind_instances = (n_slices_drct)*(n_slices_sped)
# Create wind instances. There are two columns in the wind instance array
# First Column - Wind Speed. Second Column - Wind Direction
# Shape of wind_instances (n_wind_instances,2).
# Values [1.,360.],[3.,360.],[5.,360.]...[25.,350.],[27.,350.],29.,350.]
wind_instances = np.zeros((n_wind_instances,2), dtype=np.float32)
counter = 0
for i in range(n_slices_drct):
for j in range(n_slices_sped):
wind_drct = slices_drct[i]
wind_sped = (slices_sped[j] + slices_sped[j+1])/2
wind_instances[counter,0] = wind_sped
wind_instances[counter,1] = wind_drct
counter += 1
# So that the wind flow direction aligns with the +ve x-axis.
# Convert inflow wind direction from degrees to radians
wind_drcts = np.radians(wind_instances[:,1] - 90)
# For coordinate transformation
cos_dir = np.cos(wind_drcts).reshape(n_wind_instances,1)
sin_dir = np.sin(wind_drcts).reshape(n_wind_instances,1)
# create copies of n_wind_instances wind speeds from wind_instances
wind_sped_stacked = np.column_stack([wind_instances[:,0]]*n_turbs)
# Pre-prepare matrix with stored thrust coeffecient C_t values for
# n_wind_instances shape (n_wind_instances, n_turbs, n_turbs).
# Value changing only along axis=0. C_t, thrust coeff. values for all
# speed instances.
# we use power_curve data as look up to estimate the thrust coeff.
# of the turbine for the corresponding closest matching wind speed
indices = searchSorted(power_curve[:,0], wind_instances[:,0])
C_t = power_curve[indices,1]
# stacking and reshaping to assist vectorization
C_t = np.column_stack([C_t]*(n_turbs*n_turbs))
C_t = C_t.reshape(n_wind_instances, n_turbs, n_turbs)
return(n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t)
def getAEP(turb_rad, turb_coords, power_curve, wind_inst_freq,
n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Calculates AEP of the wind farm. Vectorised version.
:called from
main
:param
turb_diam - Radius of the turbine (m)
turb_coords - 2D array turbine euclidean x,y coordinates
power_curve - For estimating power.
wind_inst_freq - 1-D flattened with rough probabilities of
wind instance occurence.
n_wind_instances - number of wind instances (int)
cos_dir - For coordinate transformation
2D Array. Shape (n_wind_instances,1)
sin_dir - For coordinate transformation
2D Array. Shape (n_wind_instances,1)
wind_sped_stacked - column staked all speed instances n_turb times.
C_t - 3D array with shape (n_wind_instances, n_turbs, n_turbs)
Value changing only along axis=0. C_t, thrust coeff.
values for all speed instances.
:return
wind farm AEP in Gigawatt Hours, GWh (float)
"""
# number of turbines
n_turbs = turb_coords.shape[0]
assert n_turbs == 50, "Error! Number of turbines is not 50."
# Prepare the rotated coordinates wrt the wind direction i.e downwind(x) & crosswind(y)
# coordinates wrt to the wind direction for each direction in wind_instances array
rotate_coords = np.zeros((n_wind_instances, n_turbs, 2), dtype=np.float32)
# Coordinate Transformation. Rotate coordinates to downwind, crosswind coordinates
rotate_coords[:,:,0] = np.matmul(cos_dir, np.transpose(turb_coords[:,0].reshape(n_turbs,1))) - \
np.matmul(sin_dir, np.transpose(turb_coords[:,1].reshape(n_turbs,1)))
rotate_coords[:,:,1] = np.matmul(sin_dir, np.transpose(turb_coords[:,0].reshape(n_turbs,1))) +\
np.matmul(cos_dir, np.transpose(turb_coords[:,1].reshape(n_turbs,1)))
# x_dist - x dist between turbine pairs wrt downwind/crosswind coordinates)
# for each wind instance
x_dist = np.zeros((n_wind_instances,n_turbs,n_turbs), dtype=np.float32)
for i in range(n_wind_instances):
tmp = rotate_coords[i,:,0].repeat(n_turbs).reshape(n_turbs, n_turbs)
x_dist[i] = tmp - tmp.transpose()
# y_dist - y dist between turbine pairs wrt downwind/crosswind coordinates)
# for each wind instance
y_dist = np.zeros((n_wind_instances,n_turbs,n_turbs), dtype=np.float32)
for i in range(n_wind_instances):
tmp = rotate_coords[i,:,1].repeat(n_turbs).reshape(n_turbs, n_turbs)
y_dist[i] = tmp - tmp.transpose()
y_dist = np.abs(y_dist)
# Now use element wise operations to calculate speed deficit.
# kw, wake decay constant presetted to 0.05
# use the jensen's model formula.
# no wake effect of turbine on itself. either j not an upstream or wake
# not happening on i because its outside of the wake region of j
# For some values of x_dist here RuntimeWarning: divide by zero may occur
# That occurs for negative x_dist. Those we anyway mark as zeros.
sped_deficit = (1-np.sqrt(1-C_t))*((turb_rad/(turb_rad + 0.05*x_dist))**2)
sped_deficit[((x_dist <= 0) | ((x_dist > 0) & (y_dist > (turb_rad + 0.05*x_dist))))] = 0.0
# Calculate Total speed deficit from all upstream turbs, using sqrt of sum of sqrs
sped_deficit_eff = np.sqrt(np.sum(np.square(sped_deficit), axis = 2))
# Element wise multiply the above with (1- sped_deficit_eff) to get
# effective windspeed due to the happening wake
wind_sped_eff = wind_sped_stacked*(1.0-sped_deficit_eff)
# Estimate power from power_curve look up for wind_sped_eff
indices = searchSorted(power_curve[:,0], wind_sped_eff.ravel())
power = power_curve[indices,2]
power = power.reshape(n_wind_instances,n_turbs)
# Farm power for single wind instance
power = np.sum(power, axis=1)
# multiply the respective values with the wind instance probabilities
# year_hours = 8760.0
AEP = 8760.0*np.sum(power*wind_inst_freq)
# Convert MWh to GWh
AEP = AEP/1e3
return(AEP)
def checkConstraints(turb_coords, turb_diam):
"""
-**-THIS FUNCTION SHOULD NOT BE MODIFIED-**-
Checks if the turbine configuration satisfies the two
constraints:(i) perimeter constraint,(ii) proximity constraint
Prints which constraints are violated if any. Note that this
function does not quantifies the amount by which the constraints
are violated if any.
:called from
main
:param
turb_coords - 2d np array containing turbine x,y coordinates
turb_diam - Diameter of the turbine (m)
:return
None. Prints messages.
"""
bound_clrnc = 50
prox_constr_viol = False
peri_constr_viol = False
# create a shapely polygon object of the wind farm
farm_peri = [(0, 0), (0, 4000), (4000, 4000), (4000, 0)]
farm_poly = Polygon(farm_peri)
# checks if for every turbine perimeter constraint is satisfied.
# breaks out if False anywhere
for turb in turb_coords:
turb = Point(turb)
inside_farm = farm_poly.contains(turb)
correct_clrnc = farm_poly.boundary.distance(turb) >= bound_clrnc
if (inside_farm == False or correct_clrnc == False):
peri_constr_viol = True
break
# checks if for every turbines proximity constraint is satisfied.
# breaks out if False anywhere
for i,turb1 in enumerate(turb_coords):
for turb2 in np.delete(turb_coords, i, axis=0):
if np.linalg.norm(turb1 - turb2) < 4*turb_diam:
prox_constr_viol = True
break
# print messages
viol_overall = False
if peri_constr_viol == True and prox_constr_viol == True:
# print('Somewhere both perimeter constraint and proximity constraint are violated\n')
viol_overall = True
elif peri_constr_viol == True and prox_constr_viol == False:
# print('Somewhere perimeter constraint is violated\n')
viol_overall = True
elif peri_constr_viol == False and prox_constr_viol == True:
# print('Somewhere proximity constraint is violated\n')
viol_overall = True
# else:
# print('Both perimeter and proximity constraints are satisfied !!\n')
return viol_overall
def f(X):
coord = []
counter = 0
for i in range(0,Num_turbines):
coo = []
coo.append(X[counter])
counter = counter + 1
coo.append(X[counter])
counter = counter + 1
coord.append(coo)
coord = np.array(coord)
if(checkConstraints(coord, turb_diam)):
obj_value = 0.001*getAEP(turb_rad, coord, power_curve, wind_inst_freq, n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t)
else:
obj_value = getAEP(turb_rad, coord, power_curve, wind_inst_freq, n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t)
# print(type(X), len(X))
return obj_value
# Turbine Specifications.
# -**-SHOULD NOT BE MODIFIED-**-
turb_specs = {
'Name': '<NAME>',
'Vendor': 'Anon Vendor',
'Type': 'Anon Type',
'Dia (m)': 100,
'Rotor Area (m2)': 7853,
'Hub Height (m)': 100,
'Cut-in Wind Speed (m/s)': 3.5,
'Cut-out Wind Speed (m/s)': 25,
'Rated Wind Speed (m/s)': 15,
'Rated Power (MW)': 3
}
turb_diam = turb_specs['Dia (m)']
turb_rad = turb_diam/2
# Turbine x,y coordinates
turb_coords = getTurbLoc('turbine_loc_test.csv')
# Load the power curve
power_curve = loadPowerCurve('power_curve.csv')
# Pass wind data csv file location to function binWindResourceData.
# Retrieve probabilities of wind instance occurence.
wind_inst_freq = binWindResourceData('wind_data_2009.csv')
# Doing preprocessing to avoid the same repeating calculations. Record
# the required data for calculations. Do that once. Data are set up (shaped)
# to assist vectorization. Used later in function totalAEP.
n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t = preProcessing(power_curve)
# check if there is any constraint is violated before we do anything. Comment
# out the function call to checkConstraints below if you desire. Note that
# this is just a check and the function does not quantifies the amount by
# which the constraints are violated if any.
# checkConstraints(turb_coords, turb_diam)
bound_min = 50
bound_max = 3950
elements = 100
Num_turbines = 50
interval = (bound_min, bound_max)
def annealing(random_start,
cost_function,
random_neighbour,
acceptance,
temperature,
maxsteps=10000,
debug=True):
""" Optimize the black-box function 'cost_function' with the simulated annealing algorithm."""
state = random_start()
cost = cost_function(state)
states, costs = [state], [cost]
best_state = state
best_cost = cost
for step in range(maxsteps):
fraction = step / float(maxsteps)
T = temperature(fraction)
new_state = random_neighbour(state, fraction)
new_cost = cost_function(new_state)
print("Step #",step,", T = ",T,", cost = ",cost,", new_cost = ",new_cost)
if acceptance_probability(cost, new_cost, T) > rn.random():
state, cost = new_state, new_cost
states.append(state)
costs.append(cost)
if best_cost >= new_cost:
best_state = new_state
best_cost = new_cost
return best_state, best_cost
def f(x):
""" Function to minimize."""
coord = []
counter = 0
for i in range(0,50):
coo = []
coo.append(x[counter])
counter = counter + 1
coo.append(x[counter])
counter = counter + 1
coord.append(coo)
coord = np.array(coord)
if(checkConstraints(coord, turb_diam)):
obj_value = -0.001*getAEP(turb_rad, coord, power_curve, wind_inst_freq, n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t)
else:
obj_value = -getAEP(turb_rad, coord, power_curve, wind_inst_freq, n_wind_instances, cos_dir, sin_dir, wind_sped_stacked, C_t)
return obj_value
def clip(x):
""" Force x to be in the interval."""
a, b = interval
return max(min(x, b), a)
def random_start():
""" Random point in the interval."""
coord = []
for i in range(0,50):
coord.append(turb_coords[i][0])
coord.append(turb_coords[i][1])
coord = np.array(coord)
return coord
def cost_function(x):
""" Cost of x = f(x)."""
return f(x)
def random_neighbour(x, fraction=1):
"""Move a little bit x, from the left or the right."""
amplitude = (max(interval) - min(interval)) * fraction / 10
vector = []
for i in x:
delta = (-amplitude/2.) + amplitude * rn.random_sample()
vector.append(clip(i + delta))
return vector
def acceptance_probability(cost, new_cost, temperature):
if new_cost < cost:
# print(" - Acceptance probabilty = 1 as new_cost = {} < cost = {}...".format(new_cost, cost))
return 1
else:
p = np.exp(- (new_cost - cost) / temperature)
# print(" - Acceptance probabilty = {:.3g}...".format(p))
return p
def temperature(fraction):
""" Example of temperature dicreasing as the process goes on."""
return max(0.001, min(1, 1 - fraction))
best_state, best_cost = annealing(random_start, cost_function, random_neighbour, acceptance_probability, temperature, maxsteps=10000, debug=True);
print(best_state)
print(best_cost)
# + id="BxoUIPwGOAQI" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9c6e799d-2f15-4fbc-d914-b5bf3ed9bbd4"
best_cost
# + id="y_zRHD7tei1F"
best=np.array(best_state)
# + id="RCkpUW63eRph"
sa=pd.DataFrame(np.ndarray.reshape(best,(50,2)))
sa.to_csv('sa_2017.csv')
# + id="f8EkHa2YPZun"
| Simulated_Anneling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="4UkbwMULuiC2" colab_type="text"
# # Physics analysis on a GPU!
#
# <NAME> (NICPB, Estonia)
#
# <EMAIL>
# + [markdown] id="n7DsM_p3uqqV" colab_type="text"
# In this notebook, we'll show how to connect `uproot`, `awkward-array`, `cupy` and `numba` to do numerically intensive physics data processing such as histogramming directly on a GPU!
#
#
# This is an entry-level tutorial, we don't focus on highly technical GPU optimizations at this time. It's mostly based on work that happened prior to the latest exciting developments (`awkward1`, `NanoEvents`, ...) in the PyHEP world, some of which we have already seen today.
# + id="9iHI8kJMnEAw" colab_type="code" colab={}
#switch to a GPU runtime, let's check which GPU we got
# + id="qgkMDr4vWA0p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="e102f70c-a409-42ff-9c93-28036ce8bdbb"
# !nvidia-smi
# + [markdown] id="Dum1pkkEv4MG" colab_type="text"
# Let's download an opendata ntuple for local use.
# + id="Qg3oO40faBuW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="94065907-6282-4f1a-ceaf-8c4b36d35d9d"
# !wget -nc http://jpata.web.cern.ch/jpata/opendata_files/DYJetsToLL-merged/1.root
# + [markdown] id="NOwbVFuZA0_K" colab_type="text"
# While the file is downloading, here's an overview of what we'll be doing.
#
# 1. Load a file with uproot, get the awkward array content
# 2. Compile a custom CUDA kernel with numba
# 3. Process an array with with threads in a kernel
# 3. Create a kernel to process jagged array events
# 4. Compute the invariant mass of jets in an event on the GPU and fill a weighted histogram
# 5. ADL exercise 7: mask jets with respect to leptons with dR, compute jet sum pt and fill into a histogram
# + [markdown] id="dA6kfll31DbL" colab_type="text"
# In CUDA, we execute functions (kernels) in many simultaneous threads. Each thread is indexed by a block and thread index and gets assigned a chunk of work based on this index. Threads in the same block run in lock step on the same GPU core (SM), each of which consists of many CUDA cores.
# + [markdown] id="WttCC9zv1INQ" colab_type="text"
# 
# + id="Nw5Rr_V9VPtM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 700} outputId="caf88a37-12a5-46ae-c77f-4c6d37a6885d"
# !pip install uproot uproot-methods awkward1 xxhash lz4 wurlitzer mplhep
# + id="RFaJOnMJk5Bm" colab_type="code" colab={}
# capture stdout directly, as it is not piped to jupyter otherwise
# https://github.com/jupyterhub/jupyterhub/issues/2035
from wurlitzer import pipes
def run_and_print(function, *args):
with pipes() as (out, err):
function(*args)
stdout = out.read()
print(stdout)
# + [markdown] id="ErQy2nKYMrv7" colab_type="text"
# # 1. Load ROOT files with uproot
# + id="3STBj3BbVRWb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="fdfdf5db-ff46-41db-c7cf-865d349fd722"
import uproot
fi = uproot.open("1.root")
evs = fi["Events"]
evs.keys()
# + id="hf_006L3a1ch" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="08e68441-b8f1-4494-9049-f32c40a95abc"
jet_pt = evs.array("Jet_pt")
jet_eta = evs.array("Jet_eta")
jet_phi = evs.array("Jet_phi")
jet_mass = evs.array("Jet_mass")
jet_pt
# + [markdown] id="Ii5R9JctMnvl" colab_type="text"
# # 2. Create an example kernel
# + id="hdVRE5kbfBgM" colab_type="code" colab={}
import numpy as np
import numba
import cupy
from numba import cuda
import math
import mplhep
import matplotlib.pyplot as plt
@cuda.jit
def example_kernel(array_in, array_out):
#just print out the thread and block indices of this kernel invocation
print("blockIdx", cuda.blockIdx.x, "blockDim", cuda.blockDim.x, "threadIdx", cuda.threadIdx.x)
#We run a kernel by calling it on a thread pool made up of nblocks and nthreads
def run(nblocks, nthreads, *args):
example_kernel[nblocks, nthreads](*args)
#the kernel itself runs asynchronously, i.e. returns immediately
#if we want to wait for all open kernels to conclude, we need to call the following:
cuda.synchronize()
#we need to copy the input to the GPU, and create a placeholder array for the output
array_in = cupy.array(jet_pt.content[:10])
array_out = cupy.zeros_like(array_in)
# + id="XtG6wnLez-M_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="c8d6d3fe-c281-461d-9089-bb7e480915b2"
array_in
# + id="Gb7PYoQGfJpl" colab_type="code" colab={}
#Let's call this function with a single thread
run(1, 1, array_in, array_out)
# + id="TuA9uiTVj_xi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="cd0b3f90-32cf-44f8-bb43-6c7bc1111a03"
#If we want to be able to access the stdout for debugging, we need to use the following wrapper function.
run_and_print(run, 1, 1, array_in, array_out)
# + id="fWudeSOulSc7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="228e03b2-3ce0-4809-c844-dd15516d2935"
#Now lets run 5 threads in one block.
run_and_print(run, 1, 5, array_in, array_out)
# + id="dPGZoB_plzgv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="258d8f11-6dca-4707-830e-9687d1282418"
#Now let's run it on two blocks, 5 threads each.
run_and_print(run, 2, 5, array_in, array_out)
# + [markdown] id="Vw1blKJnQbnO" colab_type="text"
# # 3. Process an array with a CUDA kernel
# + [markdown] id="rkMHqerZjDMO" colab_type="text"
# We can use the block and thread indices to process individual array elements in different threads.
# + id="GsS1uU2joJCe" colab_type="code" colab={}
@cuda.jit
def example_kernel_idx(array_in, array_out):
idx = cuda.blockIdx.x * cuda.blockDim.x + cuda.threadIdx.x
#just a simple example transformation within the kernel
array_out[idx] = 2.0 * array_in[idx]
print("blockIdx", cuda.blockIdx.x, "blockDim", cuda.blockDim.x, "threadIdx", cuda.threadIdx.x, "idx", idx)
def run(nblocks, nthreads):
example_kernel_idx[nblocks, nthreads](array_in, array_out)
cuda.synchronize()
# + id="__DES6Lc0qRE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4efd1111-ab19-4f59-b402-548d5d377940"
array_in
# + id="DIyx7M_0oXnb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="fba251fe-3396-47fd-d242-42cf9b164e3f"
array_out[:] = 0.0
run_and_print(run, 2, 5)
array_out
# + [markdown] id="-LDwG7cyj3Lu" colab_type="text"
# This approach requires you to choose the block size appropriately for the data array. Numba has helper functions to simplify thread and block indexing. In the following, we create a 1D grid and use threads to process slices of the input array.
#
# + id="y2sjQj-Go1RL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="0ce1a057-7a08-4923-e553-1ec115de5560"
@cuda.jit
def example_kernel_grid(array_in, array_out):
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
#each thread processes a part of the array
for i in range(xi, len(array_in), xstride):
print("arrayIdx", i, "threadIdx", cuda.threadIdx.x, "blockIdx", cuda.blockIdx.x)
def run(nblocks, nthreads):
example_kernel_grid[nblocks, nthreads](array_in, array_out)
cuda.synchronize()
#Now we can process the whole array, regardless of size, with a fixed number of threads
run_and_print(run, 2, 2)
# + [markdown] id="3OIvilGmrpDW" colab_type="text"
# # 4. Creating a jagged kernel
# + id="kEFagwWgruD2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="e6fdc483-f18c-4c9c-f695-19422b281419"
#Suppose we have the following jagged array with jet pT values in events (in GeV).
jet_pt
# + id="S5P4EnjWrvPn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="97d8c033-e2ae-4196-bc2d-876ee4c56c3b"
#Thanks to awkward-array, we can compute the per-event sum pt in a familiar way.
jet_pt.sum()
# + [markdown] id="vcIIxADTkwrc" colab_type="text"
# CuPy as a numpy replacement that runs on a CUDA GPU. Simple, universal array functions (e.g. `sin`, `cos`, `sqrt`) can be applied directly using CuPy.
# + id="J7sanM5QlBYM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="be478493-e259-4da1-9d1b-e990eab7b7bd"
import cupy
jet_pt_cuda = cupy.array(jet_pt.content)
jet_phi_cuda = cupy.array(jet_phi.content)
jet_px_cuda = jet_pt_cuda * cupy.cos(jet_phi_cuda)
jet_px_cuda
# + [markdown] id="kpC_JEJ-z1nd" colab_type="text"
# We can also compare the speed of CuPy and numpy (single-thread).
# + id="6HbVXPkJBrkv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ff2eff2f-9426-40d6-d190-757993749a26"
# %timeit np.cos(jet_phi.content)*jet_pt.content
# + id="Z-cxM6juCKb_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e1bbdec0-eb75-470a-bf96-727aca517c84"
# %timeit cupy.cos(jet_phi_cuda)*jet_pt_cuda
# + [markdown] id="mI12iwQuld7r" colab_type="text"
# What about computing event-wise quantities with jagged arrays? Normally, we would do this with awkward, using `jet_pt.sum()` to get the per-event jet $p_t$ sum. In order to run this on a GPU, we can use the awkward array structure to write a specialized kernel for computing this jagged sum. Note that the awkward1 release has support for GPU kernels, therefore this approach with numba should be considered as purely illustrational / a stop-gap.
#
# We can interpret the 1-deep jagged array using the float32 content array and the int64 offset array. Note that this is specific to the old version of awkward and will be different in awkward1.
# + id="IC6Ar3fbge14" colab_type="code" colab={}
@cuda.jit
def sum_in_offsets(offsets: numba.int64[:], content: numba.float32[:], out: numba.float32[:]):
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
#each thread gets a chunk of the events
#each event is only processed by one thread, once
for iev in range(xi, offsets.shape[0] - 1, xstride):
#process the jets in the event within the thread
#Note that this naive way will cause threads within the same block to
#possibly process a varying number of jets, causing thread divergence and resulting
#in inefficiency. Nevertheless, we can give it a try.
start = offsets[iev]
end = offsets[iev + 1]
for ielem in range(start, end):
out[iev] += content[ielem]
# + id="yNNOqinvhv43" colab_type="code" colab={}
array_in = cupy.array(jet_pt.content)
array_offsets = cupy.array(jet_pt.offsets)
array_out = cupy.zeros(len(jet_pt.offsets) - 1, dtype=cupy.float32)
sum_in_offsets[256, 1024](array_offsets, array_in, array_out); cuda.synchronize();
# + id="3_mPDbo9q1zw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6a0b4956-10ca-4abc-a4bd-f00b9bcb68ce"
array_out
# + id="lw2ycVmCq0xX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="912f29dc-0900-4752-ce96-8d7a3a067106"
jet_pt.sum()
# + [markdown] id="eNF4T2CPsR2Y" colab_type="text"
# Small differences from floating point arithmetics can be expected, but we can see that the GPU code returns the same result for this 1-deep jagged array as awkward-array. For more complicated nesting structures, awkward1 should be preferred.
#
# We can also observe that the CUDA code is relatively fast compared to a single CPU thread:
#
# + id="aQdbWtf8gfTu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="89d6cfae-f132-4b3d-ce14-0171fa085079"
# %timeit jet_pt.sum()
# + id="d8fZlKGerRED" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="845f78cc-8eb6-4815-daea-c81bf122b12a"
#Note, this does not include array allocation time
# %timeit sum_in_offsets[256, 1024](array_offsets, array_in, array_out); cuda.synchronize();
# + [markdown] id="Dqq3RrOyQw5I" colab_type="text"
# # 5. Computing the invariant mass
# + [markdown] id="HGIa9OzZsaGU" colab_type="text"
# Now let's compute the invariant mass of jets in the event.
# + id="eSRkfTdQejzj" colab_type="code" colab={}
#device functions can be used to compute quantities within kernels
@cuda.jit(device=True)
def spherical_to_cartesian_devfunc(pt: numba.float32, eta: numba.float32, phi: numba.float32, mass: numba.float32):
px = pt * math.cos(phi)
py = pt * math.sin(phi)
pz = pt * math.sinh(eta)
e = math.sqrt(px**2 + py**2 + pz**2 + mass**2)
return px, py, pz, e
@cuda.jit
def spherical_to_cartesian_kernel(
offsets: numba.int64[:],
pt: numba.float32[:],
eta: numba.float32[:],
phi: numba.float32[:],
mass: numba.float32[:],
out_inv_mass: numba.float32[:]):
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
for iev in range(xi, offsets.shape[0] - 1, xstride):
sum_px = 0.0
sum_py = 0.0
sum_pz = 0.0
sum_e = 0.0
start = offsets[iev]
end = offsets[iev + 1]
for ielem in range(start, end):
px, py, pz, e = spherical_to_cartesian_devfunc(pt[ielem], eta[ielem], phi[ielem], mass[ielem])
sum_px += px
sum_py += py
sum_pz += pz
sum_e += e
inv_mass = math.sqrt(-(sum_px**2 + sum_py**2 + sum_pz**2 - sum_e**2))
out_inv_mass[iev] = inv_mass
# + id="D-vSDIluCqb7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="992b7480-a1ed-4fbf-dba9-26acc922d8e0"
jet_offsets_cuda = cupy.array(jet_pt.offsets)
jet_pt_cuda = cupy.array(jet_pt.content, dtype=cupy.float32)
jet_eta_cuda = cupy.array(jet_eta.content, dtype=cupy.float32)
jet_phi_cuda = cupy.array(jet_phi.content, dtype=cupy.float32)
jet_mass_cuda = cupy.array(jet_mass.content, dtype=cupy.float32)
inv_mass = cupy.zeros(len(jet_offsets_cuda)-1, dtype=cupy.float32)
spherical_to_cartesian_kernel[256, 1024](
jet_offsets_cuda, jet_pt_cuda, jet_eta_cuda, jet_phi_cuda, jet_mass_cuda,
inv_mass
)
cuda.synchronize()
inv_mass
# + [markdown] id="y-O3Ntyb_GOj" colab_type="text"
# We can create a simple histogram from this data using cupy:
# + id="ZPZSeOVK_JQv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 293} outputId="a37515d1-b15a-41c5-9bfc-506e533b2705"
contents, bin_edges = cupy.histogram(inv_mass, cupy.linspace(0,10000,100, dtype=cupy.float32))
mplhep.histplot(cupy.asnumpy(contents), cupy.asnumpy(bin_edges))
# + [markdown] id="MqJHrKXU98uc" colab_type="text"
# Weighted histograms are not supported out of the box in `cupy`, so we create our own method. This also serves as a good example on how to use atomic operations.
# + id="oQffIZBiWPHu" colab_type="code" colab={}
@cuda.jit
def fill_histogram(
data: numba.float32[:],
weights: numba.float32[:],
bins: numba.float32[:],
out_w: numba.float32[:, :],
out_w2: numba.float32[:, :]):
assert(len(data) == len(weights))
assert(len(bins)-1 == out_w.shape[1])
assert(len(bins)-1 == out_w2.shape[1])
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
bi = cuda.blockIdx.x
bd = cuda.blockDim.x
ti = cuda.threadIdx.x
nbins = out_w.shape[1]
#loop over the data array
for i in range(xi, len(data), xstride):
#numba.cuda does not support searchsorted, so we need to define our own method
#bin_idx = np.searchsorted(bins, data[i])
bin_idx = searchsorted_devfunc_right(bins, data[i]) - 1
#put overflow in the last visible bin, as is typica for ROOT histograms
if bin_idx >= nbins:
bin_idx = nbins - 1
#here we define a 2D index based on the block and bin
bin_idx_histo = (bi, bin_idx)
if bin_idx >= 0 and bin_idx < nbins:
wi = weights[i]
#In order to prevent a race condition, we need to add the weights to the bin
#contents atomically. As we use the block index as an additional temporary indexing dimension
#for the histogram, threads can block each other only within the block.
cuda.atomic.add(out_w, bin_idx_histo, wi)
cuda.atomic.add(out_w2, bin_idx_histo, wi ** 2)
# Copied from numba source
@cuda.jit(device=True)
def searchsorted_inner_right(a, v):
n = len(a)
lo = np.int32(0)
hi = np.int32(n)
while hi > lo:
mid = (lo + hi) >> 1
if a[mid] <= v:
# mid is too low => go up
lo = mid + 1
else:
# mid is too high, or is a NaN => go down
hi = mid
return lo
@cuda.jit(device=True)
def searchsorted_devfunc_right(bins, val):
ret = searchsorted_inner_right(bins, val)
if val < bins[0]:
ret = 0
if val >= bins[len(bins) - 1]:
ret = len(bins) - 1
return ret
# + id="_o74vIicWd3Q" colab_type="code" colab={}
arr = cupy.random.randn(1000000, dtype=cupy.float32)
weights = cupy.ones_like(arr)
bins = cupy.linspace(-5,5, 100, dtype=cupy.float32)
nblocks = 128
nthreads = 1024
out_w = cupy.zeros((nblocks, len(bins) - 1), dtype=cupy.float32)
out_w2 = cupy.zeros((nblocks, len(bins) - 1), dtype=cupy.float32)
fill_histogram[nblocks, nthreads](arr, weights, bins, out_w, out_w2)
cuda.synchronize()
histo_w = cupy.asnumpy(out_w.sum(axis=0))
histo_w2 = cupy.asnumpy(out_w2.sum(axis=0))
# + id="m18QNMQ-vEzI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="f0f48b90-4c0f-4a45-bf99-45f03025b1c4"
#Comparing the cupy and numpy histogram functions
mplhep.histplot(cupy.asnumpy(cupy.histogram(arr, bins=bins)[0]), cupy.asnumpy(bins), ls="-", color="blue")
mplhep.histplot(np.histogram(cupy.asnumpy(arr), bins=cupy.asnumpy(bins))[0], cupy.asnumpy(bins), ls="--", color="red")
# + id="z-q_-qVLvJgB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="5dd10c31-806e-4ee5-bb1d-778b67192289"
#Comparing our custom weighted histogram with numpy/cupy
mplhep.histplot(histo_w, cupy.asnumpy(bins), yerr=np.sqrt(histo_w2), lw=0.5)
mplhep.histplot(np.histogram(cupy.asnumpy(arr), bins=cupy.asnumpy(bins))[0], cupy.asnumpy(bins), ls="-", lw=0.5, color="red")
# + id="bUsw1mh9BKDP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f360c28b-d52b-471c-d9b8-199321430722"
def test1():
fill_histogram[nblocks, nthreads](arr, weights, bins, out_w, out_w2)
cuda.synchronize()
# %timeit test1()
# + id="XbgNTM2jBX8W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8cc819e9-ac67-41d8-935e-da2c8d3b36a4"
arr_np = cupy.asnumpy(arr)
bins_np = cupy.asnumpy(bins)
# %timeit np.histogram(arr_np, bins_np)
# + [markdown] id="JKGm8Y1b0mh6" colab_type="text"
# # 6. Putting it together: Analysis Description Languages exercise 7.
# + [markdown] id="vHEPZ5QkQ-z7" colab_type="text"
# Plot the sum of pT of jets with pT > 30 GeV that are not within 0.4 in ΔR of any lepton with pT > 10 GeV.
# + id="DkwvVpnq0ohw" colab_type="code" colab={}
nblocks = 64
nthreads = 256
#This is a naive implementation and might result in thread divergence.
#Nevertheless, let's start here.
@cuda.jit(device=True)
def deltaphi_devfunc(phi1, phi2):
dphi = phi1 - phi2
out_dphi = 0
if dphi > math.pi:
dphi = dphi - 2 * math.pi
out_dphi = dphi
elif (dphi + math.pi) < 0:
dphi = dphi + 2 * math.pi
out_dphi = dphi
else:
out_dphi = dphi
return out_dphi
#This kernel masks objects in the first collection based on proximity to the second
#collection.
@cuda.jit
def mask_deltar_first_cudakernel(
etas1, phis1, mask1, offsets1, etas2, phis2, mask2, offsets2, dr2, mask_out
):
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
for iev in range(xi, len(offsets1) - 1, xstride):
a1 = np.uint64(offsets1[iev])
b1 = np.uint64(offsets1[iev + 1])
a2 = np.uint64(offsets2[iev])
b2 = np.uint64(offsets2[iev + 1])
#loop over the first collection
for idx1 in range(a1, b1):
if not mask1[idx1]:
continue
eta1 = np.float32(etas1[idx1])
phi1 = np.float32(phis1[idx1])
#loop over the second collection
for idx2 in range(a2, b2):
if not mask2[idx2]:
continue
eta2 = np.float32(etas2[idx2])
phi2 = np.float32(phis2[idx2])
deta = abs(eta1 - eta2)
dphi = deltaphi_devfunc(phi1, phi2)
passdr = (deta**2 + dphi**2) < dr2
mask_out[idx1] = passdr
#This is the same as we had above, but includes a mask
@cuda.jit
def sum_in_offsets(offsets, content, mask_rows, mask_content, out):
xi = cuda.grid(1)
xstride = cuda.gridsize(1)
for iev in range(xi, offsets.shape[0] - 1, xstride):
if not mask_rows[iev]:
continue
start = offsets[iev]
end = offsets[iev + 1]
for ielem in range(start, end):
if mask_content[ielem]:
out[iev] += content[ielem]
# + id="SgdznRy4RYB9" colab_type="code" colab={}
jet_offsets = cupy.array(evs.array("Jet_pt").offsets)
jet_pt = cupy.array(evs.array("Jet_pt").content)
jet_eta = cupy.array(evs.array("Jet_eta").content)
jet_phi = cupy.array(evs.array("Jet_phi").content)
jet_mass = cupy.array(evs.array("Jet_mass").content)
mu_offsets = cupy.array(evs.array("Muon_pt").offsets)
mu_pt = cupy.array(evs.array("Muon_pt").content)
mu_eta = cupy.array(evs.array("Muon_eta").content)
mu_phi = cupy.array(evs.array("Muon_phi").content)
#Choose the passing muons and jets
mask_mu = mu_pt > 10.0
mask_jet = jet_pt > 30.0
mask_jet_dr = cupy.zeros(len(mask_jet), dtype=cupy.bool)
# + id="KCSfd31xRagc" colab_type="code" colab={}
#Find which jets are closer than dR<0.4 to the muons
mask_deltar_first_cudakernel[nblocks, nthreads](
jet_eta, jet_phi, mask_jet, jet_offsets,
mu_eta, mu_phi, mask_mu, mu_offsets, 0.4**2, mask_jet_dr)
cuda.synchronize()
#Choose the final jets
= mask_jet & ~mask_jet_dr
# + id="avXENbCe4--R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8ed75ec4-0e21-49e1-be7c-fa336fe303c6"
selected_jets
# + id="FWj4qaDmRc2s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="06f4bccf-5deb-4a2a-83a9-5129463d0a13"
#Now compute the sum pt
nev = len(jet_offsets) - 1
sum_pt = cupy.zeros(nev, dtype=cupy.float32)
sum_in_offsets[nblocks, nthreads](
jet_offsets, jet_pt,
cupy.ones(nev, dtype=cupy.bool),
selected_jets,
sum_pt)
cuda.synchronize()
sum_pt
# + id="8ac7NcQzRfB-" colab_type="code" colab={}
#now fill the histogram
weights = cupy.ones(nev, dtype=cupy.float32)
bins = cupy.linspace(50, 500, 100, dtype=cupy.float32)
out_w = cupy.zeros((nblocks, len(bins) - 1), dtype=cupy.float32)
out_w2 = cupy.zeros((nblocks, len(bins) - 1), dtype=cupy.float32)
mask_events = sum_pt > 90.0
# + id="oOz_40CuRhg1" colab_type="code" colab={}
fill_histogram[nblocks, nthreads](sum_pt[mask_events], weights[mask_events], bins, out_w, out_w2)
cuda.synchronize()
hist_content = cupy.asnumpy(out_w.sum(axis=0))
hist_content2 = cupy.asnumpy(out_w2.sum(axis=0))
bins = cupy.asnumpy(bins)
# + id="QpE3Q3CO2XdD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 268} outputId="2b9502ef-6dc6-4c72-ad30-505834ed405b"
mplhep.histplot(hist_content, bins, yerr=np.sqrt(hist_content2))
plt.yscale("log")
# + id="SrgPUSlp5QcR" colab_type="code" colab={}
| notebooks/pyhep_gpu.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # List and Map
// In Java, the most used data structures are List (an indexed list) and Map (a dictionary)
//
// ## List
// To create a simple list
//
var numbers = List.of(1, 2, 3);
System.out.println(numbers);
// a list is an indexed data structure that stores object in the order of insertions
// get() access to an element given an index
//
var firstElement = numbers.get(0);
var lastElement = numbers.get(numbers.size() - 1);
// contains return true if a value is contained in the list
//
System.out.println(numbers.contains(4));
// indexOf returns the first index of the element in the list
//
System.out.println(numbers.indexOf(2));
// a list also defines the method equals()/hashCode() and toString(), so
// you can print a list or test if two list are equals
//
var friends = List.of("cesar", "rosalie", "david");
System.out.println(friends);
System.out.println(friends.hashCode());
System.out.println(friends.equals(numbers));
// ### Unmodifiable/modifiable list
// in Java, depending on how you create a data structure it can be changed
// after creation or not. Implementation that allow mutation after creation
// are called modifiable
//
// by example, the list above (created with the static method of()) is not modifiable
//
var countries = List.of("UK", "US", "France");
countries.set(0, "Poland"); // throws an UnsupportedOperationException
// To create a modifiable list, we use an ArrayList, created using the operator 'new'
// and because because there is no element in the list, the compiler has no way to know
// the type of the elements so we have to provide it in between angle brackets ('<' and '>')
//
var modifiableCountries = new ArrayList<String>();
System.out.println(modifiableCountries);
// To add elements in a list, we have the method add()
//
modifiableCountries.add("UK");
modifiableCountries.add("US");
modifiableCountries.add("France");
modifiableCountries.add("Poland");
System.out.println(modifiableCountries);
// to remove an element, we have the method remove()
//
modifiableCountries.remove("UK");
System.out.println(modifiableCountries);
// ### Iterating
// an unmodifiable list or a modifiable list have the same set of methods.
// to loop over the elements of a list, we have a special syntax using the keyword 'for'
//
var countries = List.of("UK", "US", "France");
for(var country: countries) {
System.out.println(country);
}
// you can also loop over the elements using a method forEach
// if you don't understand this one, don't panic, we will see it later
//
countries.forEach(country -> System.out.println(country));
// ### Conversions
// To can create an unmodifiable list from a modifiable one with List.copyOf()
//
var unmodifiableList = List.copyOf(modifiableCountries);
System.out.println(unmodifiableList);
// To create a modifiable list from an unmodifiable one using `new ArrayList(List)`
// In that case you don't have to specify the type of the elements
// the compiler already knows the type of list hence the <> (diamond)
//
var modifiableList = new ArrayList<>(unmodifiableList);
System.out.println(modifiableList);
// ### Useful patterns
// To remove some elements depending on a predicate (if something is true)
//
var elements = new ArrayList<>(List.of("table", "chair", "stool"));
elements.removeIf(element -> element.charAt(0) == 'c');
System.out.println(elements);
// ## Map
// A Map associate a value to a key
// To create a simple Map
//
var petCost = Map.of("cat", 200, "dog", 350, "lion", 5000);
System.out.println(petCost);
// to get the value from a key
//
var costOfADog = petCost.get("dog");
System.out.println(costOfADog);
// __Warning__! __warning__! asking for a key which is not in the map will return null
//
var costOfAGirafe = petCost.get("girafe");
System.out.println(costOfAGirafe);
// to avoid null, use the method `getOrDefault()` that let you specify a default value
//
var costOfAGirafe = petCost.getOrDefault("girafe", 0);
System.out.println(costOfAGirafe);
// And like a list, a map defines the method `equals()`/`hashCode()` and `toString()`
//
var lighter = Map.of("blue", "lightblue", "gray", "white");
var darker = Map.of("blue", "darkblue", "gray", "black");
System.out.println(lighter);
System.out.println(darker.hashCode());
System.out.println(lighter.equals(darker));
// ### Unmodifiable/modifiable
// The Map create by Map.of() are non modifiable
// To create a modifiable map, we use new HashMap to create the map
// and map.put to put key/value in it
//
var modifiableMap = new HashMap<String, String>();
modifiableMap.put("blue", "lightblue");
modifiableMap.put("gray", "white");
System.out.println(modifiableMap);
// Removing the key, remove the couple key/value
//
modifiableMap.remove("blue");
System.out.println(modifiableMap);
// ### Iterating on `keySet()` or `entrySet()`
// you can not use a for loop directly on a Map
// but you can do it on the set of keys
//
var petCost = Map.of("cat", 200, "dog", 350, "lion", 5000);
for(var pet: petCost.keySet()) {
System.out.println(pet);
}
// or on the set of couples key/value (a `Map.Entry`)
//
for(var entry: petCost.entrySet()) {
var pet = entry.getKey();
var cost = entry.getValue();
System.out.println(pet + " " + cost);
}
// You can also loop over the entries using the method `forEach()`
//
petCost.forEach((pet, cost) -> {
System.out.println(pet + " " + cost);
});
// ### Conversions
// To create a unmodifiableMap from a modifiable map, use `Map.copyOf()`
//
var modifiableMap = new HashMap<String, String>();
modifiableMap.put("<NAME>", "pirate");
modifiableMap.put("<NAME>", "pirate");
var unmodifiableMap = Map.copyOf(modifiableMap);
System.out.println(unmodifiableMap);
// To create a modifiableMap from an unmodifiable map, use `new HashMap<>()`
//
var unmodifiableMap = Map.of("<NAME>", "pirate", "<NAME>", "pirate");
var modifiableMap = new HashMap<>(unmodifiableMap);
System.out.println(modifiableMap);
// ### Useful patterns
// To make the Map acts as a cache, use `computeIfAbsent()`
//
record Person(String name, int age) { }
var persons = List.of(new Person("Bob", 23), new Person("Anna", 32), new Person("Bob", 12));
var group = new HashMap<String, List<Person>>();
persons.forEach(person -> group.computeIfAbsent(person.name(), name -> new ArrayList<>())
.add(person));
System.out.println(group);
// to count the number of occurrence, use `merge()` that takes a key, a value and the function
// to call if there is already an existing value to combine them
//
var letters = List.of("a", "b", "e", "b");
var occurrenceMap = new HashMap<String, Integer>();
letters.forEach(letter -> occurrenceMap.merge(letter, 1, Integer::sum));
System.out.println(occurrenceMap);
// ### More on transformations
// To do more transformations of lists and maps, use the Stream API.
//
| jupyter/chapter09-list_and_map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Семинар 6. Контейнеры и итераторы.
# <br />
# ##### begin && end
# `begin()`, `end()` - методы у контейнеров, возвращающие итераторы:
# * `begin()` на первый элемент в контейнере
# * `end()` на следующий за последним элементом
# Как, зная `begin()` и `end()`, проверить, что контейнер пуст?
# ```c++
# std::vector<int> v = {10, 20, 30, 40};
# ```
# 
# ```c++
# auto it = v.begin();
# std::cout << *it; // 10
#
# ++it;
# std::cout << *it; // 20
#
# ++it;
# std::cout << *it; // 30
#
# ++it;
# std::cout << *it; // 40
#
# ++it; // it == v.end()
# std::cout << *it; // UB - you should never dereference end()!
# ```
# <br />
# ##### Итераторы и ассоциативные контейнеры
# Обратить внимание на сложности итерирования по ассоциативным контейнерам
# 
# <br />
# ##### range for
# https://en.cppreference.com/w/cpp/language/range-for
# ```c++
# std::vector<int> v = {10, 20, 30, 40};
# for (int x : v)
# std::cout << x; // во что разворачивается range-for? (упрощённо)
# ```
# во что разворачивается range-for? (упрощённо)
# ```c++
# for (auto x : v) { ... }
# ```
# ```c++
# {
# auto&& __range = v;
# auto __begin = __range.begin();
# auto __end = __range.end();
# for (; __begin != __end; ++__begin) {
# auto x = *__begin;
# ...
# }
# }
# ```
# Смысл в том, что как только для пользовательского контейнера определены итераторы и методы `begin()`, `end()`, `cbegin()`, `cend()`, для него "из коробки" начинает работать range-for.
# <br />
# ##### Инвалидация итераторов
# Итераторы могут быть инвалидированы, если контейнер меняется.
#
# Рассмотрим пример `std::vector` в предположении, что итератор - указатель на элемент в `std::vector`.
#
# (нарисовать происходящее на доске)
# ```c++
# std::vector<int> v = {10, 20, 30, 40};
#
# auto v_end = v.end();
# auto it = v.begin();
#
# v.push_back(50);
# // at this point:
# // |it| - invalidated
# // |v_end| - invalidated
#
# std::cout << *it; // oooops, ub
# if (v.begin() == v_end) // ooooops, ub
# ...;
# ```
# https://en.cppreference.com/w/cpp/container/vector/push_back
# ```c++
# std::set<int> s = {20, 30, 40, 50};
#
# auto it = s.begin();
# std::cout << *it; // 20
#
# s.insert(10);
#
# std::cout << *it; // ok, 20
# ```
# Почему так? Потому что документация:
#
# https://en.cppreference.com/w/cpp/container/set/insert
# *У каждого контейнера у каждого метода прописан контракт на валидность итераторов (когда и какие итераторы инвалидируются).*
#
# *Читайте документацию внимательно!*
# <br />
# ##### Правильное удаление элементов из map/set/vector/... по условию
# Как неправильно удалять элементы из `std::set`:
# ```c++
# std::set<int> s = {1, 2, 3, 4, 5};
#
# auto it = s.begin();
# for(; it != s.end(); ++it)
# if((*it) % 2 == 1)
# s.erase(it);
# ```
# В каком месте баг?
# Правильное удаление:
# ```c++
# std::set<int> s = {1, 2, 3, 4, 5};
# for(auto it = s.begin(); it != s.end();)
# {
# if((*it) % 2 == 1)
# it = s.erase(it);
# else
# ++it;
# }
# ```
# <br />
# ##### Операции над итераторами доступа
# ```c++
# std::vector<int> v = {10, 20, 30, 40, 50};
#
# auto it = v.begin();
#
# // некоторые итераторы позволяют брать следующий элемент:
# auto jt_1 = it + 1;
# auto jt_2 = std::next(it);
#
# // некоторые итераторы позволяют брать предыдущий элемент:
# auto jt_3 = it - 1; // ?
# auto jt_4 = std::prev(it); // ?
#
# // некоторые итераторы позволяют прыгать на n элементов вперёд:
# auto jt_5 = it + 4;
# auto jt_6 = std::advance(it, 4);
#
# // некоторые итераторы позволяют прыгать на n элементов назад:
# auto jt_7 = it - 4; // ?
# auto jt_8 = std::advance(it, -4); // ?
#
# // некоторые итераторы позволяют считать расстояние между ними:
# std::cout << std::distance(it, jt_5); // 4
# ```
# Стандартные операции над итераторами доступа (access iterators):
# * `std::next`
# * `std::prev`
# * `std::advance`
# * `std::distance`
# <br />
# ##### Типы итераторов
# Как вы помните, у access iterators у `std::forward_list` нельзя делать `--it`, они только для хождения вперёд и только по одному шагу. А у `std::vector` можно вперёд-назад и на любое число шагов за раз. По этому принципу классифицируются итераторы доступа:
# * [Forward Iterator](https://en.cppreference.com/w/cpp/named_req/ForwardIterator)
# * [Bidirectional Iterator](https://en.cppreference.com/w/cpp/named_req/BidirectionalIterator)
# * [Random Access Iterator](https://en.cppreference.com/w/cpp/named_req/RandomAccessIterator)
# Классификация имеет важное значение для алгоритмов. Например, алгоритм сортировки работает только с Random Access Iterator:
# ```c++
# std::vector<int> v = {20, 30, 10};
# std::sort(v.begin(), v.end()); // ok
#
# std::list<int> l = {20, 30, 10};
# std::sort(l.begin(), l.end()); // compile-time error
# ```
# И это отражено в требованиях к алгоритму:
#
# https://en.cppreference.com/w/cpp/algorithm/sort
# Поэтому для `std::list` реализовали свой `sort`:
#
# https://en.cppreference.com/w/cpp/container/list/sort
# ```c++
# std::list<int> l = {20, 30, 10};
# l.sort();
# ```
# <br />
# Прочие типы итераторов:
# * [Input Iterator](https://en.cppreference.com/w/cpp/named_req/InputIterator)
# * [Output Iterator](https://en.cppreference.com/w/cpp/named_req/OutputIterator)
# * [Reverse Iterator](https://en.cppreference.com/w/cpp/iterator/reverse_iterator)
# Мощь и безобразие итераторов в двух примерах:
# ```c++
# std::istringstream str("0.1 0.2 0.3 0.4");
# const double sum = std::accumulate(std::istream_iterator<double>(str),
# std::istream_iterator<double>(),
# 0.);
#
#
# std::vector<int> v = {1, 2, 3, 4, 5};
# std::copy(v.begin(),
# v.end(),
# std::ostream_iterator<int>(std::cout, " "));
# ```
# <br />
# ##### reverse_iterator
# 
# ```c++
# std::vector<int> v = {10, 20, 30, 40};
#
# // обход в прямом направлении в стиле до С++11
# std::copy(v.begin(),
# v.end(),
# std::ostream_iterator<int>(std::cout, " ")); // 10 20 30 40
#
# // обход в обратном направлении:
# std::copy(v.rbegin(),
# v.rend(),
# std::ostream_iterator<int>(std::cout, " ")); // 40 30 20 10
#
#
# // сортировка по возрастанию:
# std::sort(v.begin(), v.end());
#
# // сортировка по убыванию:
# std::sort(v.rbegin(), v.rend());
# ```
# Конвертация iterator <-> reverse_iterator:
#
# Обратите внимание на перескакивание итератора на предыдущий элемент при конвертации.
# ```c++
# std::vector<int> v = {10, 20, 30, 40};
#
# auto it = v.begin() + 2; // 30
#
# auto rit = std::make_reverse_iterator(it); // 20 !!!!
#
# auto it2 = rit.base(); // 30 !!!!
# ```
# <br />
# Задача: найти последнее число 5 в последовательности, предшествующее первому 10
# Вариант решения:
# ```c++
# template<typename It>
# It function(It begin, It end)
# {
# auto it = std::find(begin, end, 10);
#
# if (it == end)
# return end; // no 10
#
# auto rit = std::find(std::make_reverse_iterator(it),
# std::make_reverse_iterator(begin),
# 5);
#
# if (rit == std::make_reverse_iterator(begin))
# return end; // no 5 before 10
#
# return std::next(rit.base());
# }
#
# std::list<int> l = {1, 2, 3, 5, 5, 10};
# auto it = function(l.begin(), l.end());
# auto rit = function(l.rbegin(), l.rend());
# ```
| 2019/sem1/seminar6_containers_iterators/seminar6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Mask-RCNN Model Inference in Amazon SageMaker
#
# This notebook is a step-by-step tutorial on [Mask R-CNN](https://arxiv.org/abs/1703.06870) model inference using [Amazon SageMaker model deployment hosting service](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-hosting.html).
#
# To get started, we initialize an Amazon execution role and initialize a `boto3` session to find our AWS region name.
# +
import boto3
import sagemaker
from sagemaker import get_execution_role
role = get_execution_role() # provide a pre-existing role ARN as an alternative to creating a new role
print(f'SageMaker Execution Role:{role}')
session = boto3.session.Session()
aws_region = session.region_name
print(f'AWS region:{aws_region}')
# -
# ## Build and Push Amazon SageMaker Serving Container Images
#
# For this step, the [IAM Role](https://docs.aws.amazon.com/IAM/latest/UserGuide/id_roles.html) attached to this notebook instance needs full access to [Amazon ECR service](https://aws.amazon.com/ecr/). If you created this notebook instance using the ```./stack-sm.sh``` script in this repository, the IAM Role attached to this notebook instance is already setup with full access to Amazon ECR service.
#
# Below, we have a choice of two different models for doing inference:
#
# 1. [TensorPack Faster-RCNN/Mask-RCNN](https://github.com/tensorpack/tensorpack/tree/master/examples/FasterRCNN)
#
# 2. [AWS Samples Mask R-CNN](https://github.com/aws-samples/mask-rcnn-tensorflow)
#
# It is recommended that you build and push both Amazon SageMaker <b>serving</b> container images below and use one of the two container images for serving the model from an Amazon SageMaker Endpoint.
#
# ### Build and Push TensorPack Faster-RCNN/Mask-RCNN Serving Container Image
#
# Use ```./container-serving/build_tools/build_and_push.sh``` script to build and push the [TensorPack Faster-RCNN/Mask-RCNN](https://github.com/tensorpack/tensorpack/tree/master/examples/FasterRCNN) <b>serving</b> container image to Amazon ECR.
# !pygmentize ./container-serving/build_tools/build_and_push.sh
# Using your *AWS region* as argument, run the cell below.
# %%time
# ! ./container-serving/build_tools/build_and_push.sh {aws_region}
# Set ```tensorpack_image``` below to Amazon ECR URI of the <b>serving</b> image you pushed above.
tensorpack_image = #<amazon-ecr-uri>
# ### Build and Push AWS Samples Mask R-CNN Serving Container Image
# Use ```./container-serving-optimized/build_tools/build_and_push.sh``` script to build and push the [AWS Samples Mask R-CNN](https://github.com/aws-samples/mask-rcnn-tensorflow) <b>serving</b> container image to Amazon ECR.
# !pygmentize ./container-serving-optimized/build_tools/build_and_push.sh
# Using your *AWS region* as argument, run the cell below.
# %%time
# ! ./container-serving-optimized/build_tools/build_and_push.sh {aws_region}
# Set ```aws_samples_image``` below to Amazon ECR URI of the <b>serving</b> image you pushed above.
aws_samples_image = #<amazon-ecr-uri>
# ## Select Serving Container Image
# Above, we built and pushed [TensorPack Faster-RCNN/Mask-RCNN](https://github.com/tensorpack/tensorpack/tree/master/examples/FasterRCNN) and [AWS Samples Mask R-CNN](https://github.com/aws-samples/mask-rcnn-tensorflow) serving container images to Amazon ECR. Now we are ready to deploy our trained model to an Amazon SageMaker Endpoint using one of the two container images.
#
# Next, we set ```serving_image``` to either the `tensorpack_image` or the `aws_samples_image` variable you defined above, making sure that the serving container image we set below matches our trained model.
serving_image = # set to tensorpack_image or aws_samples_image variable (no string quotes)
print(f'serving image: {serving_image}')
# ## Create Amazon SageMaker Session
# Next, we create a SageMaker session.
sagemaker_session = sagemaker.session.Session(boto_session=session)
# ## Define Amazon SageMaker Model
# Next, we define an Amazon SageMaker Model that defines the deployed model we will serve from an Amazon SageMaker Endpoint.
model_name= 'mask-rcnn-model-1'# Name of the model
# This model assumes you are using ResNet-50 pre-trained model weights for the ResNet backbone. If this is not true, please adjust `PRETRAINED_MODEL` value below. Please ensure that the `s3_model_url` of your trained model used below is consistent with the container `serving_image` you set above.
# +
s3_model_url = # Trained Model Amazon S3 URI in the format s3://<your path>/model.tar.gz
serving_container_def = {
'Image': serving_image,
'ModelDataUrl': s3_model_url,
'Mode': 'SingleModel',
'Environment': { 'SM_MODEL_DIR' : '/opt/ml/model',
'PRETRAINED_MODEL': '/ImageNet-R50-AlignPadding.npz'}
}
create_model_response = sagemaker_session.create_model(name=model_name,
role=role,
container_defs=serving_container_def)
print(create_model_response)
# -
# Next, we set the name of the Amaozn SageMaker hosted service endpoint configuration.
endpoint_config_name=f'{model_name}-endpoint-config'
print(endpoint_config_name)
# Next, we create the Amazon SageMaker hosted service endpoint configuration that uses one instance of `ml.p3.2xlarge` to serve the model.
epc = sagemaker_session.create_endpoint_config(
name=endpoint_config_name,
model_name=model_name,
initial_instance_count=1,
instance_type='ml.p3.2xlarge')
print(epc)
# Next we specify the Amazon SageMaker endpoint name for the endpoint used to serve the model.
endpoint_name=f'{model_name}-endpoint'
print(endpoint_name)
# Next, we create the Amazon SageMaker endpoint using the endpoint configuration we created above. This should take about 10 minutes.
ep=sagemaker_session.create_endpoint(endpoint_name=endpoint_name, config_name=epc, wait=True)
print(ep)
# Now that the Amazon SageMaker endpoint is in service, we will use the endpoint to do inference for test images.
#
# Next, we download [COCO 2017 Test images](http://cocodataset.org/#download).
# !wget -O ~/test2017.zip http://images.cocodataset.org/zips/test2017.zip
# We extract the downloaded COCO 2017 Test images to the home directory.
# !unzip -q -d ~/ ~/test2017.zip
# !rm ~/test2017.zip
# Below, we will use the downloaded COCO 2017 Test images to test our deployed Mask R-CNN model. However, in order to visualize the detection results, we need to define some helper functions.
# ## Visualization Helper Functions
# Next, we define a helper function to convert COCO Run Length Encoding (RLE) to a binary image mask.
#
# The RLE encoding is a dictionary with two keys `counts` and `size`. The `counts` value is a list of counts of run lengths of alternating 0s and 1s for an image binary mask for a specific instance segmentation, with the image is scanned row-wise. The `counts` list starts with a count of 0s. If the binary mask value at `(0,0)` pixel is 1, then the `counts` list starts with a `0`. The `size` value is a list containing image height and width.
# +
import numpy as np
def rle_to_binary_mask(rle, img_shape):
value = 0
mask_array = []
for count in rle:
mask_array.extend([int(value)]*count)
value = (value + 1) % 2
assert len(mask_array) == img_shape[0]*img_shape[1]
b_mask = np.array(mask_array, dtype=np.uint8).reshape(img_shape)
return b_mask
# -
# Next, we define a helper function for generating random colors for visualizing detection results.
# +
import colorsys
import random
def random_colors(N, bright=False):
brightness = 1.0 if bright else 0.7
hsv = [(i / N, 1, brightness) for i in range(N)]
colors = list(map(lambda c: colorsys.hsv_to_rgb(*c), hsv))
random.shuffle(colors)
return colors
# -
# Next, we define a helper function to apply an image binary mask for an instance segmentation to the image. Each image binary mask is of the size of the image.
def apply_mask(image, mask, color, alpha=0.5):
a_mask = np.stack([mask]*3, axis=2).astype(np.int8)
for c in range(3):
image[:, :, c] = np.where(mask == 1, image[:, :, c] *(1 - alpha) + alpha * color[c]*255,image[:, :, c])
return image
# Next, we define a helper function to show the applied detection results.
# +
import matplotlib.pyplot as plt
from matplotlib import patches
def show_detection_results(img=None,
annotations=None):
"""
img: image numpy array
annotations: annotations array for image where each annotation is in COCO format
"""
num_annotations = len(annotations)
colors = random_colors(num_annotations)
fig,ax = plt.subplots(figsize=(img.shape[1]//50, img.shape[0]//50))
for i, a in enumerate(annotations):
segm = a['segmentation']
img_shape = tuple(segm['size'])
rle = segm['counts']
binary_image_mask = rle_to_binary_mask(rle, img_shape)
bbox = a['bbox']
category_id = a['category_id']
category_name = a['category_name']
# select color from random colors
color = colors[i]
# Show bounding box
bbox_x, bbox_y, bbox_w, bbox_h = bbox
box_patch = patches.Rectangle((bbox_x, bbox_y), bbox_w, bbox_h,
linewidth=1,
alpha=0.7, linestyle="dashed",
edgecolor=color, facecolor='none')
ax.add_patch(box_patch)
label = f'{category_name}:{category_id}'
ax.text(bbox_x, bbox_y + 8, label,
color='w', size=11, backgroundcolor="none")
# Show mask
img = apply_mask(img, binary_image_mask.astype(np.bool), color)
ax.imshow(img.astype(int))
plt.show()
# -
# ## Visualize Detection Results
# Next, we select a random image from COCO 2017 Test image dataset. After you are done visualizing the detection results for this image, you can come back to the cell below and select your next random image to test.
# +
import os
import random
test2017_dir=os.path.join(os.environ['HOME'], "test2017")
img_id=random.choice(os.listdir(test2017_dir))
img_local_path = os.path.join(test2017_dir,img_id)
print(img_local_path)
# -
# Next, we read the image and convert it from BGR color to RGB color format.
# +
import cv2
img=cv2.imread(img_local_path, cv2.IMREAD_COLOR)
print(img.shape)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# -
# Next, we show the image that we randomly selected.
fig,ax = plt.subplots(figsize=(img.shape[1]//50, img.shape[0]//50))
ax.imshow(img.astype(int))
plt.show()
# Next, we invoke the Amazon SageMaker Endpoint to detect objects in the test image that we randomly selected.
#
# This REST API endpoint only accepts HTTP POST requests with `ContentType` set to `application/json`. The content of the POST request must conform to following JSON schema:
#
# `{
# "img_id": "YourImageId",
# "img_data": "Base64 encoded image file content, encoded as utf-8 string"
# }`
#
# The response of the POST request conforms to following JSON schema:
#
# `{
# "annotations": [
# {
# "bbox": [X, Y, width, height],
# "category_id": "class id",
# "category_name": "class name",
# "segmentation": { "counts": [ run-length-encoding, ], "size": [height, width]}
# },
# ]
# }`
# +
import boto3
import base64
import json
client = boto3.client('sagemaker-runtime')
with open(img_local_path, "rb") as image_file:
img_data = base64.b64encode(image_file.read())
data = {"img_id": img_id}
data["img_data"] = img_data.decode('utf-8')
body=json.dumps(data).encode('utf-8')
response = client.invoke_endpoint(EndpointName=endpoint_name,
ContentType="application/json",
Accept="application/json",
Body=body)
body=response['Body'].read()
msg=body.decode('utf-8')
data=json.loads(msg)
assert data is not None
# -
# The response from the endpoint includes annotations for the detected objects in COCO annotations format.
# Next, we aplly all the detection results to the image.
annotations = data['annotations']
show_detection_results(img, annotations)
# ## Delete SageMaker Endpoint, Endpoint Config and Model
# If you are done testing, delete the deployed Amazon SageMaker endpoint, endpoint config, and the model below. The trained model in S3. bucket is not deleted. If you are not done testing, go back to the section <b>Visualize Detection Results</b> and select another test image.
sagemaker_session.delete_endpoint(endpoint_name=endpoint_name)
sagemaker_session.delete_endpoint_config(endpoint_config_name=endpoint_config_name)
sagemaker_session.delete_model(model_name=model_name)
| mask_rcnn/mask-rcnn-inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] origin_pos=0 id="n5SYQLj6neFt"
# # 数据预处理
# :label:`sec_pandas`
#
# 为了能用深度学习来解决现实世界的问题,我们经常从预处理原始数据开始,
# 而不是从那些准备好的张量格式数据开始。
# 在Python中常用的数据分析工具中,我们通常使用`pandas`软件包。
# 像庞大的Python生态系统中的许多其他扩展包一样,`pandas`可以与张量兼容。
# 本节我们将简要介绍使用`pandas`预处理原始数据,并将原始数据转换为张量格式的步骤。
# 我们将在后面的章节中介绍更多的数据预处理技术。
#
# ## 读取数据集
#
# 举一个例子,我们首先(**创建一个人工数据集,并存储在CSV(逗号分隔值)文件**)
# `../data/house_tiny.csv`中。
# 以其他格式存储的数据也可以通过类似的方式进行处理。
# 下面我们将数据集按行写入CSV文件中。
#
# + id="I1FjQFo4ytQY"
from google.colab import drive
drive.mount('/content/drive')
# + origin_pos=1 tab=["pytorch"] id="osTfG1oeneFx"
import os
os.makedirs(os.path.join('..', 'data'), exist_ok=True)
data_file = os.path.join('..', 'data', 'house_tiny.csv')
with open(data_file, 'w') as f:
f.write('NumRooms,Alley,Price\n') # 列名
f.write('NA,Pave,127500\n') # 每行表示一个数据样本
f.write('2,NA,106000\n')
f.write('4,NA,178100\n')
f.write('NA,NA,140000\n')
# + [markdown] origin_pos=2 id="xtKJAucKneFy"
# 要[**从创建的CSV文件中加载原始数据集**],我们导入`pandas`包并调用`read_csv`函数。该数据集有四行三列。其中每行描述了房间数量(“NumRooms”)、巷子类型(“Alley”)和房屋价格(“Price”)。
#
# + origin_pos=3 tab=["pytorch"] id="vLNw-ne8neFy" outputId="c9181cac-230c-48f1-d3c0-96ce0bafb21d" colab={"base_uri": "https://localhost:8080/", "height": 175}
# 如果没有安装pandas,只需取消对以下行的注释来安装pandas
# # !pip install pandas
import pandas as pd
data = pd.read_csv(data_file)
data
# + [markdown] origin_pos=4 id="KaKfBqQ4neFz"
# ## 处理缺失值
#
# 注意,“NaN”项代表缺失值。
# [**为了处理缺失的数据,典型的方法包括*插值法*和*删除法*,**]
# 其中插值法用一个替代值弥补缺失值,而删除法则直接忽略缺失值。
# 在(**这里,我们将考虑插值法**)。
#
# 通过位置索引`iloc`,我们将`data`分成`inputs`和`outputs`,
# 其中前者为`data`的前两列,而后者为`data`的最后一列。
# 对于`inputs`中缺少的数值,我们用同一列的均值替换“NaN”项。
#
# + origin_pos=5 tab=["pytorch"] id="Y-f3or5nneF0" outputId="4229edda-2939-4abb-8d2c-e0737a5d6915" colab={"base_uri": "https://localhost:8080/"}
inputs, outputs = data.iloc[:, 0:2], data.iloc[:, 2]
inputs = inputs.fillna(inputs.mean())
print(inputs)
# + [markdown] origin_pos=6 id="GU56xP4PneF0"
# [**对于`inputs`中的类别值或离散值,我们将“NaN”视为一个类别。**]
# 由于“巷子类型”(“Alley”)列只接受两种类型的类别值“Pave”和“NaN”,
# `pandas`可以自动将此列转换为两列“Alley_Pave”和“Alley_nan”。
# 巷子类型为“Pave”的行会将“Alley_Pave”的值设置为1,“Alley_nan”的值设置为0。
# 缺少巷子类型的行会将“Alley_Pave”和“Alley_nan”分别设置为0和1。
#
# + origin_pos=7 tab=["pytorch"] id="kCbwvPAoneF1" outputId="296d1857-4668-4c20-e79c-2181e83f9ea4" colab={"base_uri": "https://localhost:8080/"}
inputs = pd.get_dummies(inputs, dummy_na=True)
print(inputs)
# + [markdown] origin_pos=8 id="Q3ICudEKneF2"
# ## 转换为张量格式
#
# [**现在`inputs`和`outputs`中的所有条目都是数值类型,它们可以转换为张量格式。**]
# 当数据采用张量格式后,可以通过在 :numref:`sec_ndarray`中引入的那些张量函数来进一步操作。
#
# + origin_pos=10 tab=["pytorch"] id="pMANNZwCneF2" outputId="18b6b196-ac54-413f-e475-29e0a9a24ef1" colab={"base_uri": "https://localhost:8080/"}
import torch
X, y = torch.tensor(inputs.values), torch.tensor(outputs.values)
X, y
# + [markdown] origin_pos=12 id="V0YH-LsNneF3"
# ## 小结
#
# * `pandas`软件包是Python中常用的数据分析工具中,`pandas`可以与张量兼容。
# * 用`pandas`处理缺失的数据时,我们可根据情况选择用插值法和删除法。
#
# ## 练习
#
# 创建包含更多行和列的原始数据集。
#
# 1. 删除缺失值最多的列。
# 2. 将预处理后的数据集转换为张量格式。
#
# + [markdown] origin_pos=14 tab=["pytorch"] id="ggvHxjFqneF3"
# [Discussions](https://discuss.d2l.ai/t/1750)
#
| chapter_preliminaries/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Fitting Models Exercise 1
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt
# + [markdown] nbgrader={}
# ## Fitting a quadratic curve
# + [markdown] nbgrader={}
# For this problem we are going to work with the following model:
#
# $$ y_{model}(x) = a x^2 + b x + c $$
#
# The true values of the model parameters are as follows:
# + nbgrader={}
a_true = 0.5
b_true = 2.0
c_true = -4.0
# + [markdown] nbgrader={}
# First, generate a dataset using this model using these parameters and the following characteristics:
#
# * For your $x$ data use 30 uniformly spaced points between $[-5,5]$.
# * Add a noise term to the $y$ value at each point that is drawn from a normal distribution with zero mean and standard deviation 2.0. Make sure you add a different random number to each point (see the `size` argument of `np.random.normal`).
#
# After you generate the data, make a plot of the raw data (use points).
# +
# np.random.normal?
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
xdata=np.linspace(-5,5,30)
dy=2
sigma=np.random.normal(0,dy,30)
ydata=a_true*xdata**2+b_true*xdata+c_true+sigma
# + deletable=false nbgrader={"checksum": "3acfeb5975cc4a690bc60e56103ce367", "grade": true, "grade_id": "fittingmodelsex01a", "points": 5}
assert True # leave this cell for grading the raw data generation and plot
# + [markdown] nbgrader={}
# Now fit the model to the dataset to recover estimates for the model's parameters:
#
# * Print out the estimates and uncertainties of each parameter.
# * Plot the raw data and best fit of the model.
# -
def model(x,a,b,c):
y=a*x**2+b*x+c
return y
def deviation(theta,x,y,dy):
a=theta[0]
b=theta[1]
c=theta[2]
return (y-a*x**2-b*x-c)/dy
xdata,ydata,sigma
# +
# opt.leastsq?
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
model_best,error_best=opt.curve_fit(model,xdata,ydata,dy)
# -
best_fit=opt.leastsq(deviation,np.array((1,2,-5)), args=(xdata, ydata, dy), full_output=True)
theta_best=best_fit[0]
theta_cov=best_fit[1]
print('a=',theta_best[0],'+/-',np.sqrt(theta_cov[0,0]))
print('b=',theta_best[1],'+/-',np.sqrt(theta_cov[1,1]))
print('c=',theta_best[2],'+/-',np.sqrt(theta_cov[2,2]))
plt.errorbar(xdata,ydata,dy,fmt='k.')
xfit=np.linspace(-5,5,100)
yfit=theta_best[0]*xfit**2+theta_best[1]*xfit+theta_best[2]
plt.plot(xfit,yfit)
plt.ylabel('y')
plt.xlabel('x')
plt.title('Quadratic Fit')
# + deletable=false nbgrader={"checksum": "5c7b35cc43322f076fb2acf1cddfc759", "grade": true, "grade_id": "fittingmodelsex01b", "points": 5}
assert True # leave this cell for grading the fit; should include a plot and printout of the parameters+errors
| assignments/assignment12/FittingModelsEx01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Some References
# * [The iris dataset and an intro to sklearn explained on the Kaggle blog](http://blog.kaggle.com/2015/04/22/scikit-learn-video-3-machine-learning-first-steps-with-the-iris-dataset/)
# * [sklearn: Conference Notebooks and Presentation from Open Data Science Conf 2015](https://github.com/amueller/odscon-sf-2015) by <NAME>
# * [real-world example set of notebooks for learning ML from Open Data Science Conf 2015](https://github.com/cmmalone/malone_OpenDataSciCon) by <NAME>
# * [PyCon 2015 Workshop, Scikit-learn tutorial](https://www.youtube.com/watch?v=L7R4HUQ-eQ0) by <NAME> (Univ of Washington, eScience Dept)
# * [Data Science for the Rest of Us](https://channel9.msdn.com/blogs/Cloud-and-Enterprise-Premium/Data-Science-for-Rest-of-Us) great introductory webinar (no math) by <NAME> (Microsoft)
# * [A Few Useful Things to Know about Machine Learning](http://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf) with useful ML "folk wisdom" by <NAME> (Univ of Washington, CS Dept)
# * [Machine Learning 101](http://www.astroml.org/sklearn_tutorial/general_concepts.html) associated with `sklearn` docs
#
# ### Some Datasets
# * [Machine learning datasets](http://mldata.org/)
# * [Make your own with sklearn](http://scikit-learn.org/stable/datasets/index.html#sample-generators)
# * [Kaggle datasets](https://www.kaggle.com/datasets)
#
# ### Contact Info
#
# <NAME><br>
# email: <EMAIL>
| Resources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Maximum Likelihood
# :label:`sec_maximum_likelihood`
#
# One of the most commonly encountered way of thinking in machine learning is the maximum likelihood point of view. This is the concept that when working with a probabilistic model with unknown parameters, the parameters which make the data have the highest probability are the most likely ones.
#
# ## The Maximum Likelihood Principle
#
# This has a Bayesian interpretation which can be helpful to think about. Suppose that we have a model with parameters $\boldsymbol{\theta}$ and a collection of data examples $X$. For concreteness, we can imagine that $\boldsymbol{\theta}$ is a single value representing the probability that a coin comes up heads when flipped, and $X$ is a sequence of independent coin flips. We will look at this example in depth later.
#
# If we want to find the most likely value for the parameters of our model, that means we want to find
#
# $$\mathop{\mathrm{argmax}} P(\boldsymbol{\theta}\mid X).$$
# :eqlabel:`eq_max_like`
#
# By Bayes' rule, this is the same thing as
#
# $$
# \mathop{\mathrm{argmax}} \frac{P(X \mid \boldsymbol{\theta})P(\boldsymbol{\theta})}{P(X)}.
# $$
#
# The expression $P(X)$, a parameter agnostic probability of generating the data, does not depend on $\boldsymbol{\theta}$ at all, and so can be dropped without changing the best choice of $\boldsymbol{\theta}$. Similarly, we may now posit that we have no prior assumption on which set of parameters are better than any others, so we may declare that $P(\boldsymbol{\theta})$ does not depend on theta either! This, for instance, makes sense in our coin flipping example where the probability it comes up heads could be any value in $[0,1]$ without any prior belief it is fair or not (often referred to as an *uninformative prior*). Thus we see that our application of Bayes' rule shows that our best choice of $\boldsymbol{\theta}$ is the maximum likelihood estimate for $\boldsymbol{\theta}$:
#
# $$
# \hat{\boldsymbol{\theta}} = \mathop{\mathrm{argmax}} _ {\boldsymbol{\theta}} P(X \mid \boldsymbol{\theta}).
# $$
#
# As a matter of common terminology, the probability of the data given the parameters ($P(X \mid \boldsymbol{\theta})$) is referred to as the *likelihood*.
#
# ### A Concrete Example
#
# Let us see how this works in a concrete example. Suppose that we have a single parameter $\theta$ representing the probability that a coin flip is heads. Then the probability of getting a tails is $1-\theta$, and so if our observed data $X$ is a sequence with $n_H$ heads and $n_T$ tails, we can use the fact that independent probabilities multiply to see that
#
# $$
# P(X \mid \theta) = \theta^{n_H}(1-\theta)^{n_T}.
# $$
#
# If we flip $13$ coins and get the sequence "HHHTHTTHHHHHT", which has $n_H = 9$ and $n_T = 4$, we see that this is
#
# $$
# P(X \mid \theta) = \theta^9(1-\theta)^4.
# $$
#
# One nice thing about this example will be that we know the answer going in. Indeed, if we said verbally, "I flipped 13 coins, and 9 came up heads, what is our best guess for the probability that the coin comes us heads?, " everyone would correctly guess $9/13$. What this maximum likelihood method will give us is a way to get that number from first principals in a way that will generalize to vastly more complex situations.
#
# For our example, the plot of $P(X \mid \theta)$ is as follows:
#
# + origin_pos=2 tab=["pytorch"]
# %matplotlib inline
from d2l import torch as d2l
import torch
theta = torch.arange(0, 1, 0.001)
p = theta**9 * (1 - theta)**4.
d2l.plot(theta, p, 'theta', 'likelihood')
# + [markdown] origin_pos=4
# This has its maximum value somewhere near our expected $9/13 \approx 0.7\ldots$. To see if it is exactly there, we can turn to calculus. Notice that at the maximum, the function is flat. Thus, we could find the maximum likelihood estimate :eqref:`eq_max_like` by finding the values of $\theta$ where the derivative is zero, and finding the one that gives the highest probability. We compute:
#
# $$
# \begin{aligned}
# 0 & = \frac{d}{d\theta} P(X \mid \theta) \\
# & = \frac{d}{d\theta} \theta^9(1-\theta)^4 \\
# & = 9\theta^8(1-\theta)^4 - 4\theta^9(1-\theta)^3 \\
# & = \theta^8(1-\theta)^3(9-13\theta).
# \end{aligned}
# $$
#
# This has three solutions: $0$, $1$ and $9/13$. The first two are clearly minima, not maxima as they assign probability $0$ to our sequence. The final value does *not* assign zero probability to our sequence, and thus must be the maximum likelihood estimate $\hat \theta = 9/13$.
#
# ## Numerical Optimization and the Negative Log-Likelihood
#
# The previous example is nice, but what if we have billions of parameters and data examples.
#
# First notice that, if we make the assumption that all the data examples are independent, we can no longer practically consider the likelihood itself as it is a product of many probabilities. Indeed, each probability is in $[0,1]$, say typically of value about $1/2$, and the product of $(1/2)^{1000000000}$ is far below machine precision. We cannot work with that directly.
#
# However, recall that the logarithm turns products to sums, in which case
#
# $$
# \log((1/2)^{1000000000}) = 1000000000\cdot\log(1/2) \approx -301029995.6\ldots
# $$
#
# This number fits perfectly within even a single precision $32$-bit float. Thus, we should consider the *log-likelihood*, which is
#
# $$
# \log(P(X \mid \boldsymbol{\theta})).
# $$
#
# Since the function $x \mapsto \log(x)$ is increasing, maximizing the likelihood is the same thing as maximizing the log-likelihood. Indeed in :numref:`sec_naive_bayes` we will see this reasoning applied when working with the specific example of the naive Bayes classifier.
#
# We often work with loss functions, where we wish to minimize the loss. We may turn maximum likelihood into the minimization of a loss by taking $-\log(P(X \mid \boldsymbol{\theta}))$, which is the *negative log-likelihood*.
#
# To illustrate this, consider the coin flipping problem from before, and pretend that we do not know the closed form solution. We may compute that
#
# $$
# -\log(P(X \mid \boldsymbol{\theta})) = -\log(\theta^{n_H}(1-\theta)^{n_T}) = -(n_H\log(\theta) + n_T\log(1-\theta)).
# $$
#
# This can be written into code, and freely optimized even for billions of coin flips.
#
# + origin_pos=6 tab=["pytorch"]
# Set up our data
n_H = 8675309
n_T = 25624
# Initialize our paramteres
theta = torch.tensor(0.5, requires_grad=True)
# Perform gradient descent
lr = 0.00000000001
for iter in range(10):
loss = -(n_H * torch.log(theta) + n_T * torch.log(1 - theta))
loss.backward()
with torch.no_grad():
theta -= lr * theta.grad
theta.grad.zero_()
# Check output
theta, n_H / (n_H + n_T)
# + [markdown] origin_pos=8
# Numerical convenience is only one reason people like to use negative log-likelihoods. Indeed, there are a several reasons that it can be preferable.
#
#
#
# The second reason we consider the log-likelihood is the simplified application of calculus rules. As discussed above, due to independence assumptions, most probabilities we encounter in machine learning are products of individual probabilities.
#
# $$
# P(X\mid\boldsymbol{\theta}) = p(x_1\mid\boldsymbol{\theta})\cdot p(x_2\mid\boldsymbol{\theta})\cdots p(x_n\mid\boldsymbol{\theta}).
# $$
#
# This means that if we directly apply the product rule to compute a derivative we get
#
# $$
# \begin{aligned}
# \frac{\partial}{\partial \boldsymbol{\theta}} P(X\mid\boldsymbol{\theta}) & = \left(\frac{\partial}{\partial \boldsymbol{\theta}}P(x_1\mid\boldsymbol{\theta})\right)\cdot P(x_2\mid\boldsymbol{\theta})\cdots P(x_n\mid\boldsymbol{\theta}) \\
# & \quad + P(x_1\mid\boldsymbol{\theta})\cdot \left(\frac{\partial}{\partial \boldsymbol{\theta}}P(x_2\mid\boldsymbol{\theta})\right)\cdots P(x_n\mid\boldsymbol{\theta}) \\
# & \quad \quad \quad \quad \quad \quad \quad \quad \quad \quad \vdots \\
# & \quad + P(x_1\mid\boldsymbol{\theta})\cdot P(x_2\mid\boldsymbol{\theta}) \cdots \left(\frac{\partial}{\partial \boldsymbol{\theta}}P(x_n\mid\boldsymbol{\theta})\right).
# \end{aligned}
# $$
#
# This requires $n(n-1)$ multiplications, along with $(n-1)$ additions, so it is total of quadratic time in the inputs! Sufficient cleverness in grouping terms will reduce this to linear time, but it requires some thought. For the negative log-likelihood we have instead
#
# $$
# -\log\left(P(X\mid\boldsymbol{\theta})\right) = -\log(P(x_1\mid\boldsymbol{\theta})) - \log(P(x_2\mid\boldsymbol{\theta})) \cdots - \log(P(x_n\mid\boldsymbol{\theta})),
# $$
#
# which then gives
#
# $$
# - \frac{\partial}{\partial \boldsymbol{\theta}} \log\left(P(X\mid\boldsymbol{\theta})\right) = \frac{1}{P(x_1\mid\boldsymbol{\theta})}\left(\frac{\partial}{\partial \boldsymbol{\theta}}P(x_1\mid\boldsymbol{\theta})\right) + \cdots + \frac{1}{P(x_n\mid\boldsymbol{\theta})}\left(\frac{\partial}{\partial \boldsymbol{\theta}}P(x_n\mid\boldsymbol{\theta})\right).
# $$
#
# This requires only $n$ divides and $n-1$ sums, and thus is linear time in the inputs.
#
# The third and final reason to consider the negative log-likelihood is the relationship to information theory, which we will discuss in detail in :numref:`sec_information_theory`. This is a rigorous mathematical theory which gives a way to measure the degree of information or randomness in a random variable. The key object of study in that field is the entropy which is
#
# $$
# H(p) = -\sum_{i} p_i \log_2(p_i),
# $$
#
# which measures the randomness of a source. Notice that this is nothing more than the average $-\log$ probability, and thus if we take our negative log-likelihood and divide by the number of data examples, we get a relative of entropy known as cross-entropy. This theoretical interpretation alone would be sufficiently compelling to motivate reporting the average negative log-likelihood over the dataset as a way of measuring model performance.
#
# ## Maximum Likelihood for Continuous Variables
#
# Everything that we have done so far assumes we are working with discrete random variables, but what if we want to work with continuous ones?
#
# The short summary is that nothing at all changes, except we replace all the instances of the probability with the probability density. Recalling that we write densities with lower case $p$, this means that for example we now say
#
# $$
# -\log\left(p(X\mid\boldsymbol{\theta})\right) = -\log(p(x_1\mid\boldsymbol{\theta})) - \log(p(x_2\mid\boldsymbol{\theta})) \cdots - \log(p(x_n\mid\boldsymbol{\theta})) = -\sum_i \log(p(x_i \mid \theta)).
# $$
#
# The question becomes, "Why is this OK?" After all, the reason we introduced densities was because probabilities of getting specific outcomes themselves was zero, and thus is not the probability of generating our data for any set of parameters zero?
#
# Indeed, this is the case, and understanding why we can shift to densities is an exercise in tracing what happens to the epsilons.
#
# Let us first re-define our goal. Suppose that for continuous random variables we no longer want to compute the probability of getting exactly the right value, but instead matching to within some range $\epsilon$. For simplicity, we assume our data is repeated observations $x_1, \ldots, x_N$ of identically distributed random variables $X_1, \ldots, X_N$. As we have seen previously, this can be written as
#
# $$
# \begin{aligned}
# &P(X_1 \in [x_1, x_1+\epsilon], X_2 \in [x_2, x_2+\epsilon], \ldots, X_N \in [x_N, x_N+\epsilon]\mid\boldsymbol{\theta}) \\
# \approx &\epsilon^Np(x_1\mid\boldsymbol{\theta})\cdot p(x_2\mid\boldsymbol{\theta}) \cdots p(x_n\mid\boldsymbol{\theta}).
# \end{aligned}
# $$
#
# Thus, if we take negative logarithms of this we obtain
#
# $$
# \begin{aligned}
# &-\log(P(X_1 \in [x_1, x_1+\epsilon], X_2 \in [x_2, x_2+\epsilon], \ldots, X_N \in [x_N, x_N+\epsilon]\mid\boldsymbol{\theta})) \\
# \approx & -N\log(\epsilon) - \sum_{i} \log(p(x_i\mid\boldsymbol{\theta})).
# \end{aligned}
# $$
#
# If we examine this expression, the only place that the $\epsilon$ occurs is in the additive constant $-N\log(\epsilon)$. This does not depend on the parameters $\boldsymbol{\theta}$ at all, so the optimal choice of $\boldsymbol{\theta}$ does not depend on our choice of $\epsilon$! If we demand four digits or four-hundred, the best choice of $\boldsymbol{\theta}$ remains the same, thus we may freely drop the epsilon to see that what we want to optimize is
#
# $$
# - \sum_{i} \log(p(x_i\mid\boldsymbol{\theta})).
# $$
#
# Thus, we see that the maximum likelihood point of view can operate with continuous random variables as easily as with discrete ones by replacing the probabilities with probability densities.
#
# ## Summary
# * The maximum likelihood principle tells us that the best fit model for a given dataset is the one that generates the data with the highest probability.
# * Often people work with the negative log-likelihood instead for a variety of reasons: numerical stability, conversion of products to sums (and the resulting simplification of gradient computations), and theoretical ties to information theory.
# * While simplest to motivate in the discrete setting, it may be freely generalized to the continuous setting as well by maximizing the probability density assigned to the datapoints.
#
# ## Exercises
# 1. Suppose that you know that a random variable has density $\frac{1}{\alpha}e^{-\alpha x}$ for some value $\alpha$. You obtain a single observation from the random variable which is the number $3$. What is the maximum likelihood estimate for $\alpha$?
# 2. Suppose that you have a dataset of samples $\{x_i\}_{i=1}^N$ drawn from a Gaussian with unknown mean, but variance $1$. What is the maximum likelihood estimate for the mean?
#
# + [markdown] origin_pos=10 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1096)
#
| d2l-en/pytorch/chapter_appendix-mathematics-for-deep-learning/maximum-likelihood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Lindronics/honours_project/blob/master/notebooks/classification/Image_classification_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_J49473r3x_G" colab_type="text"
# # Image classification test 1
#
# The purpose of this test is to:
# * familiarize myself with Pytorch
# * test classification performance of visible light vs visible light plus FIR
#
# It seems as if the FIR and visible light images aren't always taken at the same time, which might be the reason why the advanced classifier performs worse.
#
# + [markdown] id="RLxw-eJHDk0S" colab_type="text"
# ## Load dataset
# + id="9OqSTYG1DCWn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="f7105130-aeb6-4906-c326-a4248593fb7a"
# !wget http://adas.cvc.uab.es/webfiles/datasets/CVC-14-Visible-Fir-Day-Night/CVC-14.rar
# + id="FX_y0ZwvDcMg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d2036b86-6ea4-4496-b6b3-4ffc873da17f"
# !unrar x CVC-14.rar > /dev/null
# !echo "finished"
# + [markdown] id="ERjCYTn44sbK" colab_type="text"
# ## Data loader
#
# Loads visible light + FIR data.
# + id="f0_003dyDe2r" colab_type="code" colab={}
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [15, 10]
# + id="iRKs9HqmDpbU" colab_type="code" colab={}
from os import listdir
from os import path
import cv2
class CVC14Dataset(torch.utils.data.Dataset):
def __init__(self, root_dir, train_test):
self.root_dir = root_dir
self.train_test = train_test
self.visible_dir = path.join(root_dir, "Visible", train_test)
self.thermal_dir = path.join(root_dir, "FIR", train_test)
# Intersection of visible and thermal data
intersection = list(
set(listdir(path.join(self.visible_dir, "CropsPos"))) & \
set(listdir(path.join(self.thermal_dir, "CropsPos"))) \
)
self.intersection = sorted(filter(lambda x: x.endswith(".tif"), intersection))
def __len__(self):
return len(self.intersection) * 2
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
if idx >= len(self.intersection):
def random_cutout(vis, thr):
w = 64
h = 128
x = np.random.randint(0, vis.shape[0]-w)
y = np.random.randint(0, vis.shape[1]-h)
return vis[x:x+w, y:y+h, :], thr[x:x+w, y:y+h, :]
fname = sorted(filter(lambda x: x.endswith(".tif"), listdir(path.join(self.visible_dir, "FramesPos"))))[idx-len(self.intersection)]
print(idx, fname)
fpath = path.join(self.visible_dir, "FramesPos", fname)
vis = cv2.imread(fpath) / 128 - 1
fpath = path.join(self.thermal_dir, "FramesPos", fname)
thr = cv2.imread(fpath) / 128 - 1
vis, thr = random_cutout(vis, thr)
vis = cv2.resize(vis, (128, 128))
vis = np.einsum("hwc->chw", vis)
thr = cv2.resize(thr, (128, 128))
thr = np.mean(thr, axis=2)[None, ...]
return {
"visible": vis,
"thermal": thr,
"label": 0
}
else:
fname = sorted(filter(lambda x: x.endswith(".tif"), self.intersection))[idx]
print(idx, fname)
# Visible
fpath = path.join(self.visible_dir, "CropsPos", fname)
vis = cv2.imread(fpath) / 128 - 1
vis = cv2.resize(vis, (128, 128))
vis = np.einsum("hwc->chw", vis)
# Thermal
fpath = path.join(self.thermal_dir, "CropsPos", fname)
thr = cv2.imread(fpath) / 128 - 1
thr = cv2.resize(thr, (128, 128))
thr = np.mean(thr, axis=2)[None, ...] * -1
# thr = thr[None, ..., 0]
return {
"visible": vis,
"thermal": thr,
"label": 1
}
dataset = CVC14Dataset("Day", "Train")
# + id="IfRDNw-DHwgG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f063609f-cefa-46d5-dbdf-41e30482c827"
dataset[303]["visible"].min()
# + id="mZqiUatRImOy" colab_type="code" colab={}
loader = torch.utils.data.DataLoader(dataset, batch_size=4, shuffle=True)
classes = (0, 1)
# + id="uZ199fG8JwXL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 577} outputId="42aa5ae5-9be7-48ca-fb94-828a7f1cbaec"
def imshow(img):
img = img / 2 + 0.5
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
def show_batch(batch):
visible, thermal, labels = batch["visible"], batch["thermal"], batch["label"]
imshow(torchvision.utils.make_grid(visible))
imshow(torchvision.utils.make_grid(thermal))
dataiter = iter(loader)
show_batch(dataiter.next())
# + [markdown] id="M1MiSExhOfUB" colab_type="text"
# ## The model
# + id="ys5vBFS-KjMk" colab_type="code" colab={}
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16*29*29, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 2)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16*29*29)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
net = net.float()
# net(dataiter.next()["visible"].float())
# + id="mdI2ipX3W9Wa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f86964e6-eaaa-459c-a173-2da71c301596"
len(loader)
# + id="mLMb1uanOtzj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="5b7b8eac-e472-4952-e59b-faec28b72f6b"
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
for epoch in range(5):
print(f"Starting epoch {epoch}")
running_loss = 0.0
for i, data in enumerate(loader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data["visible"].float(), data["label"]
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 100 == 99: # print every 100 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 100))
running_loss = 0.0
print('Finished Training')
# + id="a3IIeSlISNvg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="51a1952e-cda3-4dbc-e807-4b7cad526137"
from sklearn.metrics import classification_report
test_dataset = CVC14Dataset("Night", "Train")
print(len(test_dataset))
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=4, shuffle=True)
all_outputs = []
all_labels = []
for i, data in enumerate(test_loader, 1000):
inputs, labels = data["visible"].float(), data["label"]
outputs = net(inputs)
_, predicted = torch.max(outputs, 1)
all_outputs += predicted.tolist()
all_labels += labels.tolist()
# + id="4MOulaaidBod" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="cb7685e9-3219-40f5-bb05-2aff61e06415"
print(classification_report(all_labels, all_outputs, target_names=["nothing", "human"]))
# + [markdown] id="vnVZut-G4xDI" colab_type="text"
# ## Advanced model (including FIR)
# + id="HXEx3mGLfk-y" colab_type="code" colab={}
class CombinedNet(nn.Module):
def __init__(self):
super(CombinedNet, self).__init__()
self.conv1 = nn.Conv2d(4, 8, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(8, 16, 5)
self.fc1 = nn.Linear(16 * 29 * 29, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 2)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16*29*29)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
comb_net = CombinedNet()
comb_net = comb_net.float()
# + id="ot3JaApbjCcP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="d5b147f0-488e-4bbb-de90-946536c3f7e0"
dataiter = iter(loader)
data = dataiter.next()
visible, thermal, labels = data["visible"].float(), data["thermal"].float(), data["label"]
inputs = torch.cat([visible, thermal], 1)
print(inputs.shape)
comb_net(inputs)
# + id="xuZtVw_QioSG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="66e65575-42a6-421f-fb14-ffaf45910d81"
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(comb_net.parameters(), lr=0.001, momentum=0.9)
for epoch in range(5):
print(f"Starting epoch {epoch}")
running_loss = 0.0
for i, data in enumerate(loader, 0):
# get the inputs; data is a list of [inputs, labels]
visible, thermal, labels = data["visible"].float(), data["thermal"].float(), data["label"]
inputs = torch.cat([visible, thermal], 1)
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = comb_net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 100 == 99: # print every 100 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 100))
running_loss = 0.0
print('Finished Training')
# + id="50GQEAkQj-Zb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c06e9d5b-2a25-459c-d70d-88396531afe1"
test_dataset = CVC14Dataset("Night", "Train")
print(len(test_dataset))
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=4, shuffle=True)
all_outputs = []
all_labels = []
for i, data in enumerate(test_loader, 1000):
visible, thermal, labels = data["visible"].float(), data["thermal"].float(), data["label"]
inputs = torch.cat([visible, thermal], 1)
outputs = comb_net(inputs)
_, predicted = torch.max(outputs, 1)
all_outputs += predicted.tolist()
all_labels += labels.tolist()
# + id="2lKlCFZ9oYwk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="3aba630f-e375-4dd9-d407-4e59de4598e5"
print(classification_report(all_labels, all_outputs, target_names=["ham", "human"]))
# + id="dznNWWDrok6X" colab_type="code" colab={}
| notebooks/classification/Image_classification_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/garima-mahato/END2/blob/main/END2_Session3_PytorchAssignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hcvnZC9gxS7l"
# # Identity Adder
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Z6bXKR5cBaA5" outputId="53e8f04e-972f-423d-eb43-90c3e710e343"
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
# + colab={"base_uri": "https://localhost:8080/"} id="u9GguNhPByDT" outputId="cb575290-9002-4544-c652-451ecd42fde3"
# %cd 'gdrive/MyDrive/END2'
# + [markdown] id="55ML3A-3vekT"
# # Code
# ---
# + id="b4r-b51WYQh4"
import os
import numpy as np
import torch
from torch.utils.data import Dataset, DataLoader
from torchvision import datasets, transforms
from torchvision.transforms import Compose, ToTensor
from torch import nn
import torch.nn.functional as F
import torch.optim as optim
from torchsummary import summary
from torch.utils.data.sampler import SubsetRandomSampler
import matplotlib.pyplot as plt
from random import randrange
# + id="Evu8eAVWrrJU" colab={"base_uri": "https://localhost:8080/"} outputId="39222588-8b3d-4c12-91ed-6f4937ebb266"
from torch_lr_finder import *
# + id="3yBPaxtabF7G"
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
# + [markdown] id="UkFKae0WwPn4"
# ## Utility
# ---
# + id="RMbBUOTIDQxP"
# visualize accuracy and loss graph
def visualize_graph(train_losses, train_acc, test_losses, test_acc):
fig, axs = plt.subplots(2,2,figsize=(15,10))
axs[0, 0].plot(train_losses)
axs[0, 0].set_title("Training Loss")
axs[1, 0].plot(train_acc)
axs[1, 0].set_title("Training Accuracy")
axs[0, 1].plot(test_losses)
axs[0, 1].set_title("Test Loss")
axs[1, 1].plot(test_acc)
axs[1, 1].set_title("Test Accuracy")
# + id="Ir-zaWMEMUVV"
def visualize_save_train_vs_test_graph(EPOCHS, dict_list, title, xlabel, ylabel, PATH, name="fig"):
plt.figure(figsize=(20,10))
#epochs = range(1,EPOCHS+1)
for label, item in dict_list.items():
x = np.linspace(1, EPOCHS+1, len(item))
plt.plot(x, item, label=label)
plt.title(title)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
plt.legend()
plt.savefig(PATH+"/"+name+".png")
# + id="f7t06Q3uQBEB"
def display_model_eval(model, test_loader):
# get some random training images
dataiter = iter(test_loader)
images, numbers, target_nums, target_sums = dataiter.next()
with torch.no_grad():
images, numbers = images.to(device), numbers.to(device)
nums, sums = model(images, numbers)
pred_num = (F.log_softmax(nums, dim=1)).argmax(dim=1, keepdim=True) # get the index of the max log-probability
pred_sum = (F.log_softmax(sums, dim=1)).argmax(dim=1, keepdim=True)
figure = plt.figure(figsize=(12, 12))
figure.suptitle('Model Prediction Results', fontsize=16)
cols, rows = 3, 3
for i in range(1, cols * rows + 1):
image = images[i-1].cpu()
number = numbers[i-1].cpu().item()
num = pred_num[i-1].cpu().item()
sum = pred_sum[i-1].cpu().item()
target_num = target_nums[i-1].cpu().item()
target_sum = target_sums[i-1].cpu().item()
figure.add_subplot(rows, cols, i)
plt.title(f'Target: {target_num} + {number} = {target_sum} \n Predicted: {num} + {number} = {sum}')
plt.axis("off")
plt.imshow(image.squeeze(), cmap="gray")
plt.show()
# + id="lWSdt2dDR-or"
#view dataset
def view_dataset(loader, title):
# get some random images
dataiter = iter(loader)
images, numbers, nums, sums = dataiter.next()
figure = plt.figure(figsize=(10, 10))
figure.suptitle(title, fontsize=16)
cols, rows = 3, 3
for i in range(1, cols * rows + 1):
image = images[i].cpu().numpy()
number = numbers[i].cpu().numpy()
num= nums[i].cpu().item()
sum = sums[i].cpu().item()
figure.add_subplot(rows, cols, i)
plt.title(f'{num} + {number} = {sum}')
plt.axis("off")
plt.imshow(image.squeeze(), cmap="gray")
plt.show()
# + [markdown] id="9d5ZVPPQvoGV"
# ## Dataset Creation
# ---
# + id="ThtB7yJc5iL4"
class IdentityAdderDataset(Dataset):
def __init__(self, train=True, transform=None):
self.transform = transform
self.mnist_data = datasets.MNIST(root="data", train=train, download=True, transform=None)
self.max_rand = 10
self.max_sum = 19
def __len__(self):
return len(self.mnist_data)
def __getitem__(self, idx):
img_input, number = self.mnist_data[idx]
rand = randrange(self.max_rand)
sum = number + rand
if self.transform:
img_input = self.transform(img_input)
return img_input, rand, number, sum
# + id="DafrDoYdNvEL"
train_transforms = transforms.Compose([
#transforms.ToPILImage(),
transforms.RandomAffine(degrees=20, translate=(0.1,0.1), scale=(0.9, 1.1)),
transforms.ColorJitter(brightness=0.2, contrast=0.2),
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
test_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
train_dataset = IdentityAdderDataset(train=True, transform=train_transforms)
test_dataset = IdentityAdderDataset(train=True, transform=test_transforms)
eval_dataset = IdentityAdderDataset(train=False, transform=test_transforms)
# + id="FG6Ge-9cTBMF"
# set seed
random_seed = 1
torch.manual_seed(random_seed)
batch_size = 128
kwargs = {'num_workers': 2, 'pin_memory': True} if use_cuda else {}
ratio = 0.2
num_train = len(train_dataset)
indices = list(range(num_train))
split = int(np.floor(ratio * num_train))
np.random.seed(random_seed)
np.random.shuffle(indices)
train_idx, test_idx = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_idx)
test_sampler = SubsetRandomSampler(test_idx)
# creating train data loader with transformations
# train_loader = DataLoader(train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
train_loader = DataLoader(train_dataset, batch_size=batch_size, sampler=train_sampler, **kwargs)
# creating test data loader with transformations
#test_loader = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, **kwargs)
test_loader = DataLoader(test_dataset, batch_size=batch_size, sampler=test_sampler, **kwargs)
eval_loader = DataLoader(eval_dataset, batch_size=batch_size, shuffle=True, **kwargs)
# + colab={"base_uri": "https://localhost:8080/"} id="6E2uxs1-Xov6" outputId="6deff0b4-cc57-4e57-dd4d-50691785a3d3"
print(f'Training Dataset size: {len(train_idx)}')
print(f'Validation Dataset size: {len(test_idx)}')
print(f'Test.Evaluation Dataset size: {len(eval_dataset)}')
# + [markdown] id="xrPrNX7UvwxF"
# ### View dataset
# ---
#
# 1) Training dataset
# + id="aXPI2tqQAIEO" colab={"base_uri": "https://localhost:8080/", "height": 646} outputId="3ed4be24-61d4-460a-ef98-a7258d9532b4"
view_dataset(train_loader,'Train Data')
# + [markdown] id="Uccx5iHGv526"
# 2) Test dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 646} id="QBg1ElBSt59V" outputId="a0623f26-ac80-4e63-ce97-762b25e1a5c3"
view_dataset(test_loader, 'Test Data')
# + [markdown] id="7D5ZrI5uxJHG"
# 3) Evaluation Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 646} id="GJmqT1A3Seaa" outputId="b9885353-dae5-4358-eb7b-f249372e876c"
view_dataset(eval_loader, 'Evaluation Data')
# + [markdown] id="9eXyvUafQLC2"
# ## Model
# ---
#
# + id="0_ZzeNH5P4Cl"
class IdentityAdderModel(nn.Module):
def __init__(self, num_channels=1, img_classes=10, sum_classes=19, dropout=0.1):
super(IdentityAdderModel, self).__init__()
self.num_channels = num_channels
self.img_classes = img_classes
self.sum_classes = sum_classes
self.dropout = dropout
# block 1
self.conv1 = nn.Sequential(
nn.Conv2d(self.num_channels, 8, 3, padding=1,bias=False), #Input : 28*28*1 - Output : 28*28*8 - RF : 3*3
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(self.dropout)
)
self.conv2 = nn.Sequential(
nn.Conv2d(8, 8, 3, padding=1,bias=False), #Input : 28*28*8 - Output : 28*28*8 - RF : 5*5
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(self.dropout)
)
self.pool1 = nn.MaxPool2d(2, 2) #Input : 28*28*8 - Output : 14*14*8 - RF : 6*6
# block 2
self.conv3 = nn.Sequential(
nn.Conv2d(8, 16, 3, padding=1,bias=False), #Input : 14*14*8 - Output : 14*14*16 - RF : 10*10
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(self.dropout)
)
self.conv4 = nn.Sequential(
nn.Conv2d(16, 16, 3, padding=1,bias=False), #Input : 14*14*16 - Output : 14*14*16 - RF : 14*14
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(self.dropout)
)
self.pool2 = nn.MaxPool2d(2, 2) #Input : 14*14*16 - Output : 7*7*16 - RF : 16*16
# block 3
self.conv5 = nn.Sequential(
nn.Conv2d(16, 16, 3, padding=1, bias=False), #Input : 7*7*16 - Output : 7*7*16 - RF : 24*24
nn.ReLU(),
nn.BatchNorm2d(16)
)
self.conv6 = nn.Sequential(
nn.Conv2d(16, 16, 3,bias=False), #Input : 7*7*16 - Output : 5*5*32 - RF : 32*32
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(self.dropout)
)
self.gap = nn.AvgPool2d(kernel_size=5)
self.flatten = nn.Flatten()
self.fc1 = nn.Sequential(
nn.Linear(16, 20, bias=False),
nn.BatchNorm1d(20),
nn.ReLU()
)
self.fc2 = nn.Sequential(
nn.Linear(30, 64, bias=False),
nn.BatchNorm1d(64),
nn.ReLU()
)
self.out_num = nn.Sequential(
nn.Linear(64, self.img_classes, bias=False)
)
self.out_sum = nn.Sequential(
nn.Linear(64, self.sum_classes, bias=False)
)
def forward(self, img, num):
x = self.conv1(img)
x = self.conv2(x)
x = self.pool1(x)
x = self.conv3(x)
x = self.conv4(x)
x = self.pool2(x)
x = self.conv5(x)
x = self.conv6(x)
x = self.gap(x)
x = self.flatten(x)
x = self.fc1(x)
inp2 = torch.squeeze(F.one_hot(num.long(), num_classes=self.img_classes), dim=1)
inp = torch.cat([x, inp2], 1)
x = self.fc2(inp)
y1 = self.out_num(x)
y2 = self.out_sum(x)
return y1, y2
# + [markdown] id="AmEfCn78w7B3"
# **Model Summary**
# + id="e1_tiWckxjk9" colab={"base_uri": "https://localhost:8080/"} outputId="45c133a2-415d-4d77-c904-17f15ce1153b"
model = IdentityAdderModel().to(device)
summary(model, input_size=[[1, 28, 28],[1]], batch_size=1)
# + [markdown] id="TzjRDAhDxpi2"
# ## Training, Testing and Evaluation Code
# ---
# + id="c8cNXZBEWyNV"
from tqdm import tqdm
train_losses = []
test_losses = []
train_acc = []
test_acc = []
def train(model, device, train_loader, criterion, optimizer, epoch, scheduler=None):
model.train()
pbar = tqdm(train_loader)
correct = 0
processed = 0
for batch_idx, (inp1, inp2, num, sum) in enumerate(pbar):
# get samples
inp1, inp2, num, sum = inp1.to(device), inp2.to(device), num.to(device), sum.to(device)
# Init
optimizer.zero_grad()
# In PyTorch, we need to set the gradients to zero before starting to do backpropragation because PyTorch accumulates the gradients on subsequent backward passes.
# Because of this, when you start your training loop, ideally you should zero out the gradients so that you do the parameter update correctly.
# Predict
out1, out2 = model(inp1, inp2)
# Calculate loss
loss = criterion(out1, num) + criterion(out2, sum)
train_losses.append(loss)
# Backpropagation
loss.backward()
optimizer.step()
if scheduler is not None:
scheduler.step()
# Update pbar-tqdm
pred1 = (F.log_softmax(out1, dim=1)).argmax(dim=1, keepdim=True) #out1.argmax(dim=1, keepdim=True) # get the index of the max log-probability
pred2 = (F.log_softmax(out2, dim=1)).argmax(dim=1, keepdim=True) #out2.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += torch.logical_and(pred1.eq(num.view_as(pred1)), pred2.eq(sum.view_as(pred2))).sum().item()
processed += len(inp1)
pbar.set_description(desc= f'Loss={loss.item()} Batch_id={batch_idx} Accuracy={100*correct/processed:0.2f}')
train_acc.append(100*correct/processed)
def test(model, device, test_loader, criterion, scheduler=None):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for inp1, inp2, num, sum in test_loader:
# get samples
inp1, inp2, num, sum = inp1.to(device), inp2.to(device), num.to(device), sum.to(device)
out1, out2 = model(inp1, inp2)
# Calculate loss
loss = criterion(out1, num) + criterion(out2, sum)
test_loss += torch.sum(loss).item() # sum up batch loss
pred1 = (F.log_softmax(out1, dim=1)).argmax(dim=1, keepdim=True) #out1.argmax(dim=1, keepdim=True) # get the index of the max log-probability
pred2 = (F.log_softmax(out2, dim=1)).argmax(dim=1, keepdim=True) #out2.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += torch.logical_and(pred1.eq(num.view_as(pred1)), pred2.eq(sum.view_as(pred2))).sum().item()
test_loss /= len(test_loader.sampler)
if scheduler is not None:
scheduler.step(test_loss)
test_losses.append(test_loss)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
test_loss, correct, len(test_loader.sampler),
100. * correct / len(test_loader.sampler)))
test_acc.append(100. * correct / len(test_loader.sampler))
# Evaluate model
def evaluate(model, device, test_loader, criterion):
model.eval()
test_loss = 0
correct = 0
loss_num = 0
loss_sum = 0
correct_num = 0
correct_sum = 0
with torch.no_grad():
for image, rand_num, num, sum in test_loader:
# get samples
image, rand_num, num, sum = image.to(device), rand_num.to(device), num.to(device), sum.to(device)
out_num, out_sum = model(image, rand_num)
# Calculate loss
loss_num += torch.sum(criterion(out_num, num)).item()
loss_sum += torch.sum(criterion(out_sum, sum)).item()
loss = criterion(out_num, num) + criterion(out_sum, sum)
test_loss += torch.sum(loss).item() # sum up batch loss
pred_num = (F.log_softmax(out_num, dim=1)).argmax(dim=1, keepdim=True) #out1.argmax(dim=1, keepdim=True) # get the index of the max log-probability
pred_sum = (F.log_softmax(out_sum, dim=1)).argmax(dim=1, keepdim=True) #out2.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct_num += pred_num.eq(num.view_as(pred_num)).sum().item()
correct_sum += pred_sum.eq(sum.view_as(pred_sum)).sum().item()
correct += torch.logical_and(pred_num.eq(num.view_as(pred_num)), pred_sum.eq(sum.view_as(pred_sum))).sum().item()
test_loss /= len(test_loader.dataset)
test_losses.append(test_loss)
avg_loss_num = loss_num / len(test_loader.dataset)
avg_loss_sum = loss_sum / len(test_loader.dataset)
avg_loss = test_loss
acc_num = 100. * correct_num / len(test_loader.dataset)
acc_sum = 100. * correct_sum / len(test_loader.dataset)
acc = 100. * correct / len(test_loader.dataset)
print('+--------------------------+')
print(' Model Evaluation Result')
print('+--------------------------+')
print(f'Average Loss for MNIST Number Prediction: {avg_loss_num}')
print(f'Average Loss for Sum Prediction: {avg_loss_sum}')
print(f'Average Loss for both Prediction: {avg_loss_num}')
print(f'Accuracy for Sum Prediction: {acc_sum}')
print(f'Accuracy for MNIST Number Prediction: {acc_num}')
print(f'Accuracy for both Prediction: {acc}')
print('+--------------------------+')
# + [markdown] id="87Xg2WYnxKby"
# ## Experimentation
# ---
# + [markdown] id="23A8wa-xyreA"
# #### With LR=0.01
# ---
#
# **Model Building and Training**
# + colab={"base_uri": "https://localhost:8080/"} id="2X5fXKXBneh9" outputId="1b5fb16f-c160-4928-a2b8-fa904d9d87d4"
# Instantiate the model
model = IdentityAdderModel().to(device)
criterion = nn.CrossEntropyLoss()
# Define the optimizer
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
epochs = 15
for epoch in range(1, epochs+1):
print(f'\nEPOCH {epoch}/{epochs}')
train(model, device, train_loader, criterion, optimizer, epoch)
test(model, device, test_loader, criterion)
# + [markdown] id="Sdg_jPhowmX4"
# Save model
# + colab={"base_uri": "https://localhost:8080/"} id="YmHGgIlB4KBs" outputId="8f5b1aa9-8a78-45d1-941b-1bb5ea5c31f0"
print("\n Saving trained model...")
torch.save(model.state_dict(), "./model/final_model.pth")
# + [markdown] id="A173fgRhwZb7"
# #### View Accuracy and Loss while model training and testing
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 641} id="9Wppq7iNzKvT" outputId="b85380f1-5a13-4f5f-c536-c499b3e91ada"
print("\n Visualizing:")
visualize_graph(train_losses, train_acc, test_losses, test_acc)
# + [markdown] id="0Gfjo9owwPEo"
# #### Train versus Test Accuracy Plot
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="iyWLNdC-zW6n" outputId="2295cda0-53a2-466d-d30a-3b12114ee630"
dict_list = {'Training Accuracy': train_acc, 'Test Accuracy': test_acc}
title = "Training vs Test Accuracy"
xlabel = "Epochs"
ylabel = "Accuracy(in Percentage)"
name = "train_vs_test_acc_comparison_graph"
visualize_save_train_vs_test_graph(epochs, dict_list, title, xlabel, ylabel, "./visualization", name=name)
# + [markdown] id="4SAKGEA_wDIi"
# #### Evalution Metrics on Evaluation Dataset
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="gJGY52wszmwA" outputId="ad4f76ac-a1b2-4688-d45d-dceb4db03703"
evaluate(model, device, eval_loader, criterion)
# + [markdown] id="NwoDAqTwv7M9"
# #### View Model Predictions for Evaluation dataset
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 770} id="_sjQU_4czrQn" outputId="1b0f4114-f966-4036-a020-87a725a28dd5"
display_model_eval(model, eval_loader)
# + [markdown] id="D0YEThoRvz2f"
# ### Save as ONNX model
# + colab={"base_uri": "https://localhost:8080/"} id="00MwHKPM45xs" outputId="b39b04ee-7005-455f-a3e6-079ec8e95aed"
import torch.onnx
import netron
dataiter = iter(test_loader)
images, numbers, _, _ = dataiter.next()
with torch.no_grad():
images, numbers = images.to(device), numbers.to(device)
images, numbers = images.cpu(), numbers.cpu()
image = images[0].cpu().unsqueeze(0)
number = numbers[0].cpu().unsqueeze(0).unsqueeze(0)
m = IdentityAdderModel()
o = m(images, numbers)
onnx_path = "onnx_identity_adder_model.onnx"
torch.onnx.export(m, (image, number), onnx_path)
netron.start(onnx_path)
| Session3-PyTorch/END2_Session3_PytorchAssignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Множества и словари
#
# *<NAME>, НИУ ВШЭ*
[1, 1.2, 'hey', [1, 24, 5]]
t = (1, 1.2, 'hey', [1, 24, 5])
t[-1]
t[-1][0] = 99
t
t[-1] = 99
s = set()
set([1, 1, 1, 2, 2, 3, 3])
s1 = set([1, 1, 1, 2, 2, 3, 3])
s1
# # Множества (set)
#
# Мы уже знаем списки и кортежи - упорядоченные структуры, которые могут хранить в себе объекты любых типов, к которым мы можем обратиться по индексу. Теперь поговорим о стуктурах неупорядоченных - множествах и словарях.
#
# Множества хранят некоторое количество объектов, но, в отличие от списка, один объект может храниться в множестве не более одного раза. Кроме того, порядок элементов множества произволен, им нельзя управлять.
#
# Тип называется set, это же является конструктором типа, т.е. в функцию set можно передать произвольную последовательность, и из этой последовательности будет построено множество:
x = set()
type(x)
set([1, 1, 1, 2, 2, 3])
x = (4, 5, 6)
print(set(x)) # передаем список
set(range(10))
print(set()) # передаем tuple
print(set(range(10)))
print(set()) # пустое множество
#
# Другой способ создать множество - это перечислить его элементы в фигурных скобках (список - в квадратных, кортеж в круглых, а множество - в фигурных)
# +
primes = {2, 2, 3, 5, 7}
animals = {"cat", "dog", "horse", 'cat'}
print(primes)
print(animals)
# -
primes
x = [1,2,3,2,2,2,2,2,2,2,'dog',1]
print(set(x))
# Кстати, обратите внимание, что множество может состоять только из уникальных объектов. Выше множество animals включает в себя только одну кошку несмотря на то, что в конструктор мы передали 'cat' два раза. Преобразовать в список в множество - самый простой способ узнать количество уникальных объектов.
#
# Со множествами работает почти всё, что работает с последовательностями (но не работают индексы, потому что элементы не хранятся упорядоченно).
print(len(primes))
primes = {1,11,22,34,5}
11 in primes
animals = {"cat", "dog", "horse", 'cat'}
"cow" in animals
animals
len(animals)
# длина
print(11 in primes) # проверка на наличие элемента in хорошо и быстро работает для множеств
print("cow" in animals)
# Все возможные операции с множествами: https://docs.python.org/3/library/stdtypes.html#set-types-set-frozenset
#
# Отдельно мы посмотрим на так называемые операции над множествами. Если вы знаете круги Эйлера, то помните как различают объекты множеств - пересечение, объекты, которые принадлежат множеству а, но не принадлежат b и так далее. Давайте посмотрим, как эти операции реализовани в питоне.
a = {1, 2, 3, 4}
b = {3, 4, 5, 6}
c = {2, 3}
print(c <= a)
a1 = {1, 2}
a2 = {1, 2}
a1 < a2
a1.issubset(a2)
a1 <= a2
a1 < a2
print(c >= a)
c <= b
a = {1, 2, 3, 4}
b = {3, 4, 5, 6}
c = {2, 3}
print(a | b)
a.intersection(b)
a.union(b)
a ^ b
b.difference(a)
b - a
# +
a = {1, 2, 3, 4}
b = {3, 4, 5, 6}
c = {2, 3}
# проверка на подмножество (с подномжество a)
print(c <= b) # не подмножество, т.к. в b нет 2
print(a >= c)
print(a | b) # объединение a.union(b) aka a+b
print(a & b) # пересечение a.intersection(b)
print(a - b) # разность множеств (все что в a, кроме b) a.difference(b)
print(a ^ b) # симметрическая разность множеств (объединение без пересечения)
c = a.copy() # копирование множества, или set(a)
print(c)
# -
a = [1, 2, 3]
b = a
b[-1] = 999
b
a
a = [1, 2, 3]
b = a.copy()
a = [1, 2, 3]
b = a[:]
b[-1] = 999
b
a
a = {1, 2, 3, 4}
c = a.copy()
c
print(c.issubset(a)) # c <= a
print(c.isdisjoint(a)) # a и c не пересекаются?
print(a.issuperset(c)) # a включает в себя с как подмножество
s = {1, 2, 3}
s
s.add(10)
s
s.add(10)
s
s.discard(500)
s
s.remove(500)
x = s.pop()
print(s)
print(x)
s
s.clear()
s
# Предыдущие операции не меняли множества, создавали новые. А как менять множество:
#
s = {1, 2, 3}
s.add(10) # добавить
print(s) # обратите внимание, что порядок элементов непредсказуем
s.remove(1) # удаление элемента
s.discard(1) # аналогично, но не будет ошибки, если вдруг такого элемента нет в множестве
print(s)
x = s.pop() # удаляет и возвращает один произвольный элемент множества (можем сохранить его в переменную)
print(s)
print(x)
s.clear() # очистить
print(s)
x = 1
x += 1 # x = x + 1
# Как мы сокращали арифметические операции раньше (например, +=), так же можно сокращать операции над множествами.
s |= {1.5, 20} # s = s | {10, 20} # объединение множества s с {10,20}
print(s)
# s ^=, s &= и т.п.
# # Словари (dict)
# Обычный массив (в питоне это список) можно понимать как функцию, которая сопоставляет начальному отрезку натурального ряда какие-то значения.
# Давайте посмотрим на списки непривычным способом. Списки - это функции (отображения), которые отображают начальный ряд натуральных чисел в объекты (проще говоря - преводят число 0,1,2,3... во что-то):
l = [10, 20, 30, 'a']
print(l[0])
print(l[1])
print(l[2])
print(l[3])
# В словарях отображать можно не только начала натурального ряда, а произвольные объекты. Представьте себе настоящий словарь или телефонную книжку. Имени человека соответствует номер телефона.
#
# Классическое использование словарей в анализе данных: хранить частоту слова в тексте.
#
# кот $\rightarrow$ 10
#
# и $\rightarrow$ 100
#
# Тейлора $\rightarrow$ 2
# Словарь состоит из набора ключей и соответствующих им значений. Значения могут быть любыми объектами (также как и в списке, хранить можно произвольные объекты). А ключи могут быть почти любыми объектами, но только неизменяемыми. В частности числами, строками, кортежами. Список или множество не могут быть ключом.
#
# Одному ключу соответствует ровно одно значение. Но одно и то же значение, в принципе, можно сопоставить разным ключам.
d = dict()
d = {}
type({})
s = set()
type(s)
{1: None}
{'Ivanov': [1, 2, 3, 4], 'Petrov': [6, 7]}
1, 1.5, 'hey', (1, 2)
d = {1: 1, 1.5:2, 'hey': 1, (1, 2): 5}
{[1, 2]: 'a'}
d = {1: 1, 1.5:2, 'hey': 1, (1, 2): 5, 'hey': 100}
d
s = 'aaaabbbccc'
s[0]
d[0]
d
d[1]
d[1.5]
d['hey']
d[(1, 2)]
a = dict()
type(a)
a['chapter1'] = 'ghfdlksgrjkasgdjkagrdjksargs'
a
a[1] = 'hrjegrejk'
a
x = (3,4,5)
a[x] = 'hrjekwslerw'
a
x = (1,2,3,'str')
a[x] = 'fuidaslt'
a
a
a = dict()
a[(2,3)] = [2,3] # кортеж может быть ключом, потому что он неизменямый
a
b = dict()
b[[2,3]] = [2,3] # а список уже нет, получим ошибку
print(b)
len(a.keys())
a.keys()
a.values()
a.values()[0]
list(a.values())[0]
students = {'Petrov': [1, 2]}
students['Petrov'].append(10)
students
# ### Создание словаря
# В фигурных скобках (как множество), через двоеточие ключ:значение
d = dict()
d
# +
# d[ключ] = значение
# -
# {ключ: значение, ключ2: значение2}
d1 = {"кот": 10, "и": 100, "Тейлора": 2}
print(d1)
d1["кот"]
# Через функцию dict(). Обратите внимание, что тогда ключ-значение задаются не через двоеточие, а через знак присваивания. А строковые ключи пишем без кавычек - по сути мы создаем переменные с такими названиями и присваиваим им значения (а потом функция dict() уже превратит их в строки).
d2 = dict(кот=10, и=100, Тейлора=2)
print(d2) # получили тот же результат, что выше
# И третий способ - передаем функции dict() список списков или кортежей с парами ключ-значение.
d3 = dict([("и", 100), ("кот", 10), ("Тейлора", 2)]) # перечисление (например, список) tuple
print(d3)
# Помните, когда мы говорили про списки, мы обсуждали проблему того, что важно создавать именно копию объекта, чтобы сохранять исходный список. Копию словаря можно сделать так
d4 = dict(d3) # фактически, копируем dict который строчкой выше
print(d4)
d1 == d2 == d3 == d4 # Содержание всех словарей одинаковое
z = [1, 2, 3]
d15 = {1: z.copy()}
d15[1].append(99)
d15
z
# Пустой словарь можно создать двумя способами.
d2 = {} # это пустой словарь (но не пустое множество)
d4 = dict()
print(d2, d4)
type({})
type(d2)
type({1, 2, 3})
x = {}
type(x)
# ### Операции со словарями
# Как мы уже говорили, словари неупорядоченные структуры и обратиться по индексу к объекту уже больше не удастся.
d1
d1[1] # выдаст ошибку во всех случах кроме того, если в вашем словаре вдруг есть ключ 1
# Но можно обращаться к значению по ключу.
d3 = dict([("кот", 10), ("и", 100), ("Тейлора", 2)])
print(d1['кот'])
d1[1]
# Можно создать новую пару ключ-значение. Для этого просто указываем в квадратных скобках название нового ключа.
d1[1] = 'test'
print(d1[1]) # теперь работает!
d1
# Внимание: если элемент с указанным ключом уже существует, новый с таким же ключом не добавится! Ключ – это уникальный идентификатор элемента. Если мы добавим в словарь новый элемент с уже существующим ключом, мы просто изменим старый – словари являются изменяемыми объектами.
d1[1.5] = 'test2'
d1
d1.keys()
d1.values()
d1.items()
list(d1.keys())[0]
d1[list(d1.keys())[0]]
d3 = dict([("кот", 10), ("и", 100), ("Тейлора", 2)])
d1["кот"] = 11 # так же как в списке по индексу - можно присвоить новое значение по ключу
d1
d1["кот"] += 1 # или даже изменить его за счет арифметической операции
d1
# А вот одинаковые значения в словаре могут быть.
d1['собака'] = 13
print(d1)
# Кроме обращения по ключу, можно достать значение с помощью метода .get(). Отличие работы метода в том, что если ключа еще нет в словаре, он не генерирует ошибку, а возвращает объект типа None ("ничего"). Это очень полезно в решении некоторых задач.
d1
d1[2.5]
d1.get(2.5)
d1.get(2.5, 'такого нет!')
d1.get('кот', 'такого нет!')
print(d1['кот'])
print(d1.get("ктоо")) # вернут None
# Удобство метода .get() заключается в том, что мы сами можем установить, какое значение будет возвращено, в случае, если пары с выбранным ключом нет в словаре. Так, вместо None мы можем вернуть строку Not found, и ломаться ничего не будет:
print(d1.get("ктоо", 'Not found')) # передаем вторым аргументом, что возвращать
print(d1.get("ктоо", False)) # передаем вторым аргументом, что возвращать
# Также со словарями работают уже знакомые нам операции - проверка количества элементов, проверка на наличие объектов.
d1
print(d1)
print("кот" in d1) # проверка на наличие ключа
print("ктоо" not in d1) # проверка на отстуствие ключа
# Удалить отдельный ключ или же очистить весь словарь можно специальными операциями.
del d1["кот"]
d1
d1.clear()
d1
del d1["кот"] # удалить ключ со своим значением
print(d1)
d1.clear() # удалить все
print(d1)
d1 = dict([("кот", 10), ("и", 10), ("Тейлора", 2)])
# У словарей есть три метода, с помощью которых мы можем сгенерировать список только ключей, только значений и список пар ключ-значения (на самом деле там несколько другая структура, но ведет себя она очень похоже на список).
print(d1.values()) # только значения
print(d1.keys()) # только ключи
print(d1.items()) # только ключ-значение
d1
for x in d1:
print(x)
for x in d1.keys():
print(x)
for x in d1.values():
print(x)
for x in d1.items():
print(x)
for key, value in d1.items():
print(f'{key} is {value}')
# Ну, и раз уж питоновские словари так похожи на обычные, давайте представим, что у нас есть словарь, где все слова многозначные. Ключом будет слово, а значением ‒ целый список.
my_dict = {'swear' : {'swear' : ['клясться', 'ругаться'],
'dream' : ['СПАТЬ', 'МЕЧТАТЬ']},
'dream' : ['спать', 'мечтать']}
my_dict['swear']
my_dict['swear']['dream']
my_dict['swear']['dream'][0]
my_dict['swear']['dream'][0] + '!!!!!'
my_dict.keys()
my_dict['swear']
my_dict['swear']['swear']
my_dict['swear']['dream']
my_dict['swear']['dream'][1]
my_dict['dream']
my_dict['dream']['dream']
my_dict.items()
type(my_dict.items())
len(my_dict.items())
# По ключу мы получим значение в виде списка:
my_dict['swear']['swear'][0]
# Так как значением является список, можем отдельно обращаться к его элементам:
my_dict['swear'][0] # первый элемент
# Можем пойти дальше и создать словарь, где значениями являются словари! Например, представим, что в некотором сообществе проходят выборы, и каждый участник может проголосовать за любое число кандидатов. Данные сохраняются в виде словаря, где ключами являются имена пользователей, а значениями – пары *кандидат-голос*.
votes = {'user1': {'cand1': '+', 'cand2': '-'},
'user2' : {'cand1': 0, 'cand3' : '+'}} # '+' - за, '-' - против, 0 - воздержался
votes
# По аналогии с вложенными списками по ключам мы сможем обратиться к значению в словаре, который сам является значением в `votes` (да, эту фразу нужно осмыслить):
votes['user1']['cand1'] # берем значение, соответствующее ключу user1, в нем – ключу cand1
dict1 = {'a': 1, 'b': 2}
dict2 = {'c': 3, 'd': 4}
dict3 = dict1.copy()
dict3.update(dict2)
print(dict3)
dict1 = {'a': 1, 'b': 2}
dict2 = {'a': 3, 'd': 4}
dict3 = {**dict1, **dict2}
print(dict3)
| class_5/class_5_set_dict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aaptecode/DS-Unit-3-Sprint-1-Software-Engineering/blob/master/PyPi_Check.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="kJzgBAxohXH0" colab_type="code" outputId="ce5db77c-330a-40fc-e135-67ee7a73407b" colab={"base_uri": "https://localhost:8080/", "height": 190}
# !pip install -U -i https://test.pypi.org/simple/ lambdata-aaptecode
# + id="JBFktEigh61L" colab_type="code" outputId="76fdbf0b-d82d-4d1c-b2e5-7854694c2746" colab={"base_uri": "https://localhost:8080/", "height": 190}
import lambdata_aaptecode.mltools as mltools
dir(mltools)
# + id="gsmKOKCZAqCy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="35b03d27-c8cf-44f9-95c3-cdb9558bca8f"
help(mltools.conf_matrix_2d)
# + id="-JA6jBvTAulN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="bbbd8b08-655f-4895-a052-949a29a12c5d"
help(mltools.train_validation_test_split)
| PyPi_Check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Jaira21/OOP-58002/blob/main/FUNDAMENTALS_OF_PYTHON.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rQp9WsN429L_"
# Python Identation
# + colab={"base_uri": "https://localhost:8080/"} id="K4VzowGeIkSB" outputId="7c0ed508-52ef-4930-8d14-1427b2634fe7"
if 5>2:
print("Yes")
# + [markdown] id="GuhRiZtGInJY"
# Python Comments
# + colab={"base_uri": "https://localhost:8080/"} id="3iJNT0w3Io-Q" outputId="2ad1af3e-9473-4faa-e8e7-679777034397"
#This is a comment
print("Hello, World")
# + [markdown] id="t7QoLUowJILw"
# Python Variable
# + colab={"base_uri": "https://localhost:8080/"} id="XSrHF846JKJY" outputId="58f3dd44-3890-4429-f91b-fc2ec6675802"
x = "Sally"
a = 0
a,b,c=0,1,2
print(x)
print(a)
print(b)
print(c)
print(a,b,c)
# + [markdown] id="p8KlBVOVJMaY"
# Casting
# + colab={"base_uri": "https://localhost:8080/"} id="vewfdPNDJOhw" outputId="e327e703-86db-4a8c-e881-86f699a9c356"
d = 4
d = int(4)
print(d)
# + [markdown] id="os2lA4dJJXFo"
# Type () Function
# + colab={"base_uri": "https://localhost:8080/"} id="za6mFOHhJZ6o" outputId="9edf98b4-255f-4eca-b1f7-e0d44bd388b6"
d = 4
d = int(4)
print(type(d))
# + [markdown] id="w1iXiT8-JjLQ"
# Double quotes and single quotes
# + colab={"base_uri": "https://localhost:8080/"} id="iSaIgWk8JlUI" outputId="be739dda-db33-464b-d5f5-86bb4a6397d4"
#y = "Ana"
#print(y)
y = 'Ana'
Y = "Robert"
print(y)
print(Y)
# + [markdown] id="DLGKmXXQJzdJ"
# Multiple Variables with one value
# + colab={"base_uri": "https://localhost:8080/"} id="057XcGZBJ1Iw" outputId="db557382-72eb-47bd-f70c-f1dfdc864786"
#x = "Tony"
k=l=m="four"
print(k)
print(l)
print(m)
print(k,l,m)
# + [markdown] id="DkmgkwdEJ3iP"
# Output Variable
# + colab={"base_uri": "https://localhost:8080/"} id="HBWqtNTmJ5Zf" outputId="f3328448-2e90-4164-afd2-5a3a79f6632d"
print("Python programming is enjoying")
h = "enjoying"
p = "Python programming is"
print("Python programming is" + h)
print(p+ "" + "" +h)
# + [markdown] id="2UCbFzKyJ8un"
# Arithmetic Operations
# + colab={"base_uri": "https://localhost:8080/"} id="QCtOoNDcJ_Df" outputId="18d530d0-2346-4cf3-bd13-cb97de7d632b"
print(c+d) # c = 2, d = 4
print(d-c)
print(d*c)
print(int(d/c))
print(d%c)
print(3//2) # 1.50
print(d**c)
# + [markdown] id="nqHLy5KmKBp_"
# Assignment Operators
# + colab={"base_uri": "https://localhost:8080/"} id="YlfodzMXKDe_" outputId="c65e1386-424f-4ae2-e51a-4057a6453ff0"
q=10
q+=5 #same as q=q+5
print(q) #sane as q = q+5, q 10+5=15
# + [markdown] id="wftSqVMWKGtP"
# Boolean Logic
# + colab={"base_uri": "https://localhost:8080/"} id="gbVAykBQKHzA" outputId="fd5df271-8347-4928-fcf1-5de01eabd3b4"
s = 10
print(s^2)
print(s|2)
# + [markdown] id="NI1p4APMKJ_o"
# Comparison Operators
# + colab={"base_uri": "https://localhost:8080/"} id="iN8_nCv1KMDU" outputId="4d3be6f1-fcbf-4ec7-9998-20e88ff725fb"
print(s>q)
print(s==s)
print(q==q)
# + [markdown] id="09ux4wGBKN63"
# Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="jYdGx6A4KP0A" outputId="e504110f-b35a-48b6-9dc3-6ca455e528df"
s>q and s==s #False
s>q or s==s #True
print(s>q and s==s)
print(s>q or s==s)
# + [markdown] id="l8PI0uUQKSGw"
# Identity Operators
# + colab={"base_uri": "https://localhost:8080/"} id="2-_1cVAhKVHY" outputId="f34e9317-3e7d-46cb-e219-659d7624927b"
s is q
print(s is not q)
# + [markdown] id="9shfngmwKXVX"
# Bitwise Operators Shift right
# + id="vyeXz_9eKZL3"
u = 2 #0001
print(u<<1)
g = 10
print(10<<1)
| FUNDAMENTALS_OF_PYTHON.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: 928b61f5e44213fcfe6dada5ff80a864126784261d6cf6ac4f3c3dedebb569eb
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
def isNaN(num):
return num != num
csv = pd.read_csv('hungary_deaths.csv', sep=',')
csv
last_year = 2014
for i in range(0, len(csv)):
if (not pd.isnull(csv["Év"][i])):
last_year = last_year + 1
csv["Év"][i] = last_year
csv
deaths_per_week = csv[["Összesen.2"]]
deaths_per_week
# +
deaths_per_year = pd.DataFrame(columns=range(2015, 2020+1), index=pd.RangeIndex(1, 54, name='week'), dtype='Int64')
for index, row in csv[["Év", "A hét sorszáma", "Összesen.2"]].iterrows():
year = int(row["Év"])
week = int(row["A hét sorszáma"])
deaths_per_year.loc[week, year] = int(row["Összesen.2"].replace(",",""))
# -
deaths_per_year
# +
def data_for_year(y):
year = deaths_per_year[y].dropna().to_numpy()
if y == 2020:
num_weeks = len(year)
day_of_the_year = num_weeks*7 - 3 # ex. week 46 -> november 15 -> day 319
theta = np.linspace(0, (day_of_the_year/365)*2*np.pi, num_weeks)
else:
# append first week of next year for correct radial plotting
year = np.append(year, deaths_per_year.loc[1, y+1])
theta = np.linspace(0, 2*np.pi, len(year))
return (theta, year)
def plot_year(ax, y, **kwargs):
ax.plot(*data_for_year(y), label=f"{y}", **kwargs)
def setup_polar_plot(figsize=(8, 6), constrained_layout=True):
fig = plt.figure(figsize=figsize, constrained_layout=constrained_layout)
ax = fig.add_subplot(111, projection='polar')
ax.set_theta_zero_location('N')
ax.set_theta_direction(-1)
months = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']
ax.set_xticks(np.arange(0, 2*np.pi, np.pi/6))
ax.set_xticklabels(months)
ax.set_rlabel_position(180)
ax.set_yticklabels(['', '1000', '', '2000', ''])
return fig, ax
fig, ax = setup_polar_plot()
# plot_year(ax, 2000, linewidth=0.5)
# plot_year(ax, 2001, linewidth=0.5)
# plot_year(ax, 2002, linewidth=0.5)
# plot_year(ax, 2003, linewidth=0.5)
# plot_year(ax, 2004, linewidth=0.5)
# plot_year(ax, 2005, linewidth=0.5)
# plot_year(ax, 2006, linewidth=0.5)
# plot_year(ax, 2007, linewidth=0.5)
# plot_year(ax, 2008, linewidth=0.5)
# plot_year(ax, 2009, linewidth=0.5)
# plot_year(ax, 2010, linewidth=0.5)
plot_year(ax, 2015, color='tab:gray')
plot_year(ax, 2016, color='tab:blue', linestyle='dashdot')
plot_year(ax, 2017, color='tab:blue')
plot_year(ax, 2018, color='tab:green')
plot_year(ax, 2019, color='tab:orange')
plot_year(ax, 2020, color='tab:red', linewidth=3)
ax.set_rmax(5000)
fig.legend(loc='center right')
fig.suptitle("Deaths in Hungary per Week", fontsize=14, y=1.04)
ax.set_title("Source: KSH", fontsize=10, y=1.08)
plt.savefig('deathshun.png', dpi=300, bbox_inches='tight', facecolor='white')
# +
years = deaths_per_year.iloc[:, :-1] # excluding 2020
mean = years.mean(axis=1)
mean[53] = mean[1]
median = years.median(axis=1)
median[53] = median[1]
min = years.min(axis=1)
min[53] = min[1]
max = years.max(axis=1)
max[53] = max[1]
q25 = deaths_per_year.quantile(0.25, axis=1).astype(float)
q25[53] = q25[1]
q75 = deaths_per_year.quantile(0.75, axis=1).astype(float)
q75[53] = q75[1]
fig, ax = setup_polar_plot()
# ax.plot(np.linspace(0, 2*np.pi, len(mean)), mean, label="5y mean")
# ax.plot(np.linspace(0, 2*np.pi, len(data_for_year(2019))), data_for_year(2019), label="2019")
ax.fill_between(np.linspace(0, 2*np.pi, len(min)), min, max, alpha=0.2, label="min/max")
ax.fill_between(np.linspace(0, 2*np.pi, len(q25)), q25, q75, alpha=0.3, label="50%", color='tab:blue')
plot_year(ax, 2020, color='tab:red', linewidth=3)
ax.plot(np.linspace(0, 2*np.pi, len(median)), median, label="Median", linestyle='dashed')
ax.set_rmax(5000)
fig.legend(loc='lower right')
fig.suptitle(f"Deaths in Hungary per week", fontsize=14, y=1.04)
ax.set_title("Source: KSH", fontsize=10, y=1.08)
plt.savefig('median.png', dpi=300, bbox_inches='tight', facecolor='white')
# + tags=[]
fig, ax = setup_polar_plot(figsize=(6, 6.2), constrained_layout=False)
# nudge ax position
pos = ax.get_position()
pos.y0 -= 0.05
pos.y1 -= 0.05
pos.x0 -= 0.012
pos.x1 -= 0.012
ax.set_position(pos)
fig.suptitle("Deaths in Hungary per week", fontsize=14)
ax.set_title("Source: KSH", fontsize=10, y=1.1)
old, = ax.plot([], [], color='tab:blue', linewidth=0.5, linestyle='dotted', label="Vorjahre")
prev, = ax.plot([], [], color='tab:blue', label="2019")
current, = ax.plot([], [], color='tab:red', linewidth=3, label="2020")
center = ax.text(0, 25, "2015", horizontalalignment='center', fontsize=18)
ax.set_rmax(4000)
start_year = 2015
def year_and_week_for_index(i):
y = start_year
while True:
len_year = len(deaths_per_year[y].dropna()) + 1
if len_year > i:
return (y, i+1)
else:
y += 1
i -= (len_year-1)
def data_for_index(i):
y, w = year_and_week_for_index(i)
print(y, w)
theta, year = data_for_year(y)
return theta[:w], year[:w]
def init():
old.set_data([], [])
prev.set_data([], [])
current.set_data([], [])
center.set_text("")
return old, prev, current, center
def animate(i):
y = year_and_week_for_index(i)[0]
if y > start_year:
old_theta = np.array([])
old_data = np.array([])
for year in range(start_year, y-1):
theta, data = data_for_year(year)
old_theta = np.append(old_theta, theta)
old_data = np.append(old_data, data)
old.set_data(old_theta, old_data)
prev.set_data(*data_for_year(y-1))
current.set_data(*data_for_index(i))
center.set_text(f"{y}")
return old, prev, current, center
num_frames = len(deaths_per_week)
anim = mpl.animation.FuncAnimation(fig, animate, init_func=init, frames=num_frames, interval=50, blit=True)
anim.save('deaths.mp4', writer='ffmpeg', dpi=300, extra_args=['-vf', 'tpad=stop_mode=clone:stop_duration=5'])
fig.legend(loc='lower right')
plt.savefig('deaths.png', dpi=300, bbox_inches='tight', facecolor='white')
# anim.save('anim.gif', writer='imagemagick', dpi=300, savefig_kwargs={'facecolor': 'white'})
# plt.savefig('anim.png', dpi=300, bbox_inches='tight', facecolor='white')
# $ ffmpeg -i deaths.mp4 -vf "fps=15,scale=900:-1:flags=lanczos,split[s0][s1];[s0]palettegen[p];[s1][p]paletteuse" -loop 0 deaths.gif
# -
| sterblichkeit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelos de Categorización
# Modelos de categorización siguiendo la referencia de <NAME>., & <NAME>. (2010). Cognitive modeling. Sage.
# ## Modelos de prototipos
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 12, 120, endpoint=True)
y1 = np.exp(-(np.abs(x-3)/1)**2)
y2 = np.exp(-(np.abs(x-7)/1)**2)
plt.plot(x, y1, 'r', x, y2, 'b')
plt.title('Activación para el modelo de prototipos')
plt.xlabel('Valor del Estímulo')
plt.ylabel('Activación de los nodos')
plt.show()
# ## Modelos Ejemplares
# +
# %matplotlib notebook
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter, MultipleLocator
import numpy as np
# -
fig = plt.figure()
ax = fig.gca(projection='3d')
fig = plt.figure()
ax = fig.gca(projection='3d')
X = np.linspace(0, 12, 120)
Y = np.linspace(0, 12, 120)
X, Y = np.meshgrid(X, Y)
Z = (np.exp(-(np.abs(X-3)/3)**2))*(np.exp(-(np.abs(Y-7)/3)**2))
ax.contour3D(X, Y, Z, 120, cmap='plasma')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z');
ax = plt.axes(projection='3d')
ax.plot_surface(X, Y, Z, rstride=1, cstride=1,
cmap='viridis', edgecolor='none')
ax.set_title('Activación producida por el modelo Ejemplar')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('Activación del nodo');
ax.set_zlim(0, 1)
ax.zaxis.set_major_locator(MultipleLocator(0.1))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
| Categorizacion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # NRPy+'s Finite Difference Interface
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# ### NRPy+ Source Code for this module: [finite_difference.py](../edit/finite_difference.py)
#
# <a id='toc'></a>
#
# # Table of Contents \[Back to [top](#toc)\]
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Preliminaries](#fdd): Introduction to Finite Difference Derivatives
# 1. [Step 1](#fdmodule): The finite_difference NRPy+ module
# 1. [Step 1.a](#fdcoeffs_func): The `compute_fdcoeffs_fdstencl()` function
# 1. [Step 1.a.i](#exercise): Exercise: Using `compute_fdcoeffs_fdstencl()`
# 1. [Step 1.b](#fdoutputc): The `FD_outputC()` function
# 1. [Step 2](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF
# <a id='fdd'></a>
#
# # Preliminaries: Introduction to Finite Difference Derivatives \[Back to [top](#toc)\]
# $$\label{fdd}$$
#
# Suppose we have a *uniform* numerical grid in one dimension; say, the Cartesian $x$ direction. Since the grid is uniform, the spacing between successive grid points is $\Delta x$, and the position of the $i$th point is given by
#
# $$x_i = x_0 + i \Delta x.$$
#
# Then, given a function $u(x)$ on this uniform grid, we will adopt the notation
#
# $$u(x_i) = u_i.$$
#
# We wish to approximate derivatives of $u_i$ at some nearby point (in this tutorial, we will consider derivatives at one of the sampled points $x_i$) using [finite difference](https://en.wikipedia.org/wiki/Finite_difference). (FD) techniques.
#
# FD techniques are usually constructed as follows:
# * First, find the unique $N$th-degree polynomial that passes through $N+1$ sampled points of our function $u$ in the neighborhood of where we wish to find the derivative.
# * Then, provided $u$ is smooth and properly-sampled, the $n$th derivative of the polynomial (where $n\le N-1$; *Exercise: Justify this inequality*) is approximately equal to the $n$th derivative of $u$. We call this the **$n$th-order finite difference derivative of $u$**.
# * So long as the function $u$ is smooth and properly sampled, the relative error between the exact and the finite difference derivative $u^{(n)}$ will generally decrease as the polynomial degree or sampling density increases.
#
# The $n$th finite difference derivative of $u(x)$ at $x=x_i$ can then be written in the form
# $$u^{(n)}(x_i)_{\text{FD}} = \sum_{j=0}^{N} u_j a_j,$$
# where the $a_j$'s are known as *finite difference coefficients*. So long as the $N$th-degree polynomial that passes through the $N+1$ points is unique, the corresponding set of $a_j$'s are unique as well.
#
# There are multiple ways to compute the finite difference coefficients $a_j$, including solving for the $N$th-degree polynomial that passes through the function at the sampled points. However, the most popular and most straightforward way involves Taylor series expansions about sampled points near the point where we wish to evaluate the derivative.
#
# **Recommended: Learn more about the algorithm NRPy+ adopts to automatically compute finite difference derivatives: ([How NRPy+ Computes Finite Difference Coefficients](Tutorial-How_NRPy_Computes_Finite_Difference_Coeffs.ipynb))**
#
# <a id='fdmodule'></a>
#
# # Step 1: The finite_difference NRPy+ module \[Back to [top](#toc)\]
# $$\label{fdmodule}$$
#
# The finite_difference NRPy+ module contains one parameter:
#
# * **FD_CENTDERIVS_ORDER**: An integer indicating the requested finite difference accuracy order (not the order of the derivative) , where FD_CENTDERIVS_ORDER = [the size of the finite difference stencil in each direction, plus one].
#
# The finite_difference NRPy+ module contains two core functions: `compute_fdcoeffs_fdstencl()` and `FD_outputC()`. The first is a low-level function normally called only by `FD_outputC()`, which computes and outputs finite difference coefficients and the numerical grid indices (stencil) corresponding to each coefficient:
# <a id='fdcoeffs_func'></a>
#
# ## Step 1.a: The `compute_fdcoeffs_fdstencl()` function \[Back to [top](#toc)\]
# $$\label{fdcoeffs_func}$$
#
# **compute_fdcoeffs_fdstencl(derivstring,FDORDER=-1)**:
# * Output nonzero finite difference coefficients and corresponding numerical stencil as lists, using as inputs:
# * **derivstring**: indicates the precise type and direction derivative desired:
# * **Centered derivatives**, where the center of the finite difference stencil corresponds to the point where the derivative is desired:
# * For a first-order derivative, set derivstring to "D"+"dirn", where "dirn" is an integer denoting direction. For a second-order derivative, set derivstring to "DD"+"dirn1"+"dirn2", where "dirn1" and "dirn2" are integers denoting the direction of each derivative. Currently only $1 \le N \le 2$ supported (extension to higher-order derivatives is straightforward). Examples in 3D Cartesian coordinates (x,y,z):
# * the derivative operator $\partial_x^2$ corresponds to derivstring = "DD00"
# * the derivative operator $\partial_x \partial_y$ corresponds to derivstring = "DD01"
# * the derivative operator $\partial_z$ corresponds to derivstring = "D2"
# * **Up- or downwinded derivatives**, where the center of the finite difference stencil is *one gridpoint* up or down from where the derivative is requested.
# * Set derivstring to "upD"+"dirn" or "dnD"+"dirn", where "dirn" is an integer denoting direction. Example in 3D Cartesian coordinates (x,y,z):
# * the upwinded derivative operator $\partial_x$ corresponds to derivstring = "dupD0"
# * **Kreiss-Oliger dissipation derivatives**, where the center of the finite difference stencil corresponds to the point where the dissipation will be applied.
# * Set derivstring to "dKOD"+"dirn", where "dirn" is an integer denoting direction. Example in 3D Cartesian coordinates (x,y,z):
# * the Kreiss-Oliger derivative operator $\partial_z^\text{KO}$ corresponds to derivstring = "dKOD2"
# * **FDORDER**: an *optional* parameter that, if set to a positive even integer, overrides FD_CENTDERIVS_ORDER
#
# Within NRPy+, `compute_fdcoeffs_fdstencl()` is only called from `FD_outputC()`. Regardless, this function provides a nice interface for evaluating finite difference coefficients, as shown below:
# +
# Import the finite difference module
import finite_difference as fin # NRPy+: Finite difference C code generation module
fdcoeffs, fdstencl = fin.compute_fdcoeffs_fdstencl("dDD00")
print(fdcoeffs)
print(fdstencl)
# -
# Interpreting the output, notice first that $\texttt{fdstencl}$ is a list of coordinate indices, where up to 4 dimension indices are supported (higher dimensions are possible and can be straightforwardly added, though be warned about [The Curse of Dimensionality](https://en.wikipedia.org/wiki/Curse_of_dimensionality)).
#
# Thus NRPy+ found that for some function $u$, the fourth-order accurate finite difference operator at point $x_{i0}$ is given by
#
# $$[\partial_{x}^{2} u]^\text{FD4}_{i0} = \frac{1}{\Delta x^{2}} \left[ -\frac{1}{12} \left(u_{i0-2,i1,i2,i3} + u_{i0+2,i1,i2,i3}\right) - \frac{5}{2}u_{i0,i1,i2,i3} + \frac{4}{3}\left(u_{i0-1,i1,i2,i3} + u_{i0+1,i1,i2,i3}\right)\right]$$
#
# Notice also that multiplying by the appropriate power of $\frac{1}{\Delta x}$ term is up to the user of this function.
#
# In addition, if the gridfunction $u$ exists on a grid that is less than four (spatial) dimensions, it is up to the user to truncate the additional index information.
# <a id='exercise'></a>
#
# ### Step 1.a.i: Exercise: Using `compute_fdcoeffs_fdstencl()` \[Back to [top](#toc)\]
# $$\label{exercise}$$
#
# Using `compute_fdcoeffs_fdstencl()` write the necessary loops to output the finite difference coefficient tables in the Wikipedia article on [finite difference coefficients](https://en.wikipedia.org/wiki/Finite_difference_coefficients), for first and second centered derivatives (i.e., up to $\partial_i^2$) up to eighth-order accuracy. [Solution, courtesy <NAME>](Tutorial-Finite_Difference_Derivatives-FDtable_soln.ipynb).
# <a id='fdoutputc'></a>
#
# ## Step 1.b: The `FD_outputC()` function \[Back to [top](#toc)\]
# $$\label{fdoutputc}$$
#
# **FD_outputC(filename,sympyexpr_list)**: C code generator for finite-difference expressions.
#
# C codes that evaluate expressions with finite difference derivatives on numerical grids generally consist of three components, all existing within a loop over "interior" gridpoints; at a given gridpoint, the code must
# 1. Read gridfunctions from memory at all points needed to evaluate the finite difference derivatives or the gridfunctions themselves.
# 2. Perform arithmetic, including computation of finite difference stencils.
# 3. Write the output from the arithmetic to other gridfunctions.
#
# To minimize cache misses and maximize potential compiler optimizations, it is generally recommended to segregate the above three steps. FD_outputC() first analyzes the input expressions, searching for derivatives of gridfunctions. The search is very easy, as NRPy+ requires a very specific syntax for derivatives:
# * gf_dD0 denotes the first derivative of gridfunction "gf" in direction zero.
# * gf_dupD0 denotes the upwinded first derivative of gridfunction "gf" in direction zero.
# * gf_ddnD0 denotes the downwinded first derivative of gridfunction "gf" in direction zero.
# * gf_dKOD2 denotes the Kreiss-Oliger dissipation operator of gridfunction "gf" in direction two.
# Each time `FD_outputC()` finds a derivative (including references to the gridfunction directly \["zeroth"-order derivatives\]) in this way, it calls `compute_fdcoeffs_fdstencl()` to record the specific locations in memory from which the underlying gridfunction must be read to evaluate the appropriate finite difference derivative.
#
# `FD_outputC()` then orders this list of points for all gridfunctions and points in memory, optimizing memory reads based on how the gridfunctions are stored in memory (set via parameter MemAllocStyle in the NRPy+ grid module). It then completes step 1.
#
# For step 2, `FD_outputC()` exports all of the finite difference expressions, as well as the original expressions input into the function, to outputC() to generate the optimized C code. Step 3 follows trivally from just being careful with the bookkeeping in the above steps.
#
# `FD_outputC()` takes two arguments:
# * **filename**: Set to "stdout" to print to screen. Otherwise specify a filename.
# * **sympyexpr_list**: A single named tuple or list of named tuples of type "lhrh", where the lhrh type refers to the simple structure:
# * **lhrh(left-hand side of equation, right-hand side of the equation)**
#
# Time for an example: let's compute
# $$
# \texttt{output} = \text{phi_dDD00} = \partial_x^2 \phi(x,t),
# $$
# where $\phi$ is a function of space and time, though we only store its spatial values at a given time (*a la* the [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html), described & implemented in next the [Scalar Wave Equation module](Tutorial-Start_to_Finish-ScalarWave.ipynb)).
#
# As detailed above, the suffix $\text{_dDD00}$ tells NRPy+ to construct the second finite difference derivative of gridfunction $\texttt{phi}$ with respect to coordinate $xx0$ (in this case $xx0$ is simply the Cartesian coordinate $x$). Here is the NRPy+ implementation:
# +
import sympy as sp # SymPy, Python's core symbolic algebra package on which NRPy+ depends
from outputC import lhrh # NRPy+: Core C code output module
import NRPy_param_funcs as par # NRPy+: parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import finite_difference as fin # NRPy+: Finite difference C code generation module
# Set the spatial dimension to 1
par.set_paramsvals_value("grid::DIM = 1")
# Register the input gridfunction "phi" and the gridfunction to which data are output, "output":
phi, output = gri.register_gridfunctions("AUX",["phi","output"])
# Declare phi_dDD as a rank-2 indexed expression: phi_dDD[i][j] = \partial_i \partial_j phi
phi_dDD = ixp.declarerank2("phi_dDD","nosym")
# Set output to \partial_0^2 phi
output = phi_dDD[0][0]
# Output to the screen the core C code for evaluating the finite difference derivative
fin.FD_outputC("stdout",lhrh(lhs=gri.gfaccess("out_gf","output"),rhs=output))
# -
# Some important points about the above code:
# * The gridfunction PHIGF samples some function $\phi(x)$ at discrete uniform points in $x$, labeled $x_i$ at all points $i\in [0,N]$, so that
# $$\phi(x_i) = \phi_{i}=\text{in_gfs[IDX2(PHIGF, i)]}.$$
# * For a *uniformly* sampled function with constant grid spacing (sample rate) $\Delta x$, $x_i$ is defined as $x_i = x_0 + i \Delta x$.
# * The variable $\texttt{invdx0}$ must be defined by the user in terms of the uniform gridspacing $\Delta x$ as $\texttt{invdx0} = \frac{1}{\Delta x}$.
# * *Aside*: Why do we choose to multiply by $1/\Delta x$ instead of dividing the expression by $\Delta x$, which would seem much more straightforward?
# * *Answer*: as discussed in the [first part of the tutorial](Tutorial-Coutput__Parameter_Interface.ipynb), division of floating-point numbers on modern CPUs is far more expensive than multiplication, usually by a factor of ~3 or more.
# <a id='latex_pdf_output'></a>
#
# # Step 2: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-Finite_Difference_Derivatives.pdf](Tutorial-Finite_Difference_Derivatives.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-Finite_Difference_Derivatives")
| Tutorial-Finite_Difference_Derivatives.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-wmdecomp
# language: python
# name: venv-wmdecomp
# ---
# +
from collections import defaultdict
from wmdecompose.documents import Document
from wmdecompose.gale_shapeley import Matcher
from wmdecompose.models import LC_RWMD, WMD, WMDPairs
from wmdecompose.utils import *
from gensim.models import KeyedVectors
from nltk.corpus import stopwords
from nltk.tokenize import ToktokTokenizer
from random import shuffle
from scipy.spatial.distance import is_valid_dm, cdist
from sklearn.decomposition import PCA
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.manifold import TSNE
from sklearn.metrics import silhouette_score
from sklearn import cluster
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import random
import re
import seaborn as sns
import umap
# %load_ext autoreload
# %autoreload 2
random.seed(42)
# -
# ## 1. Prepare IMDB data
# ### 1.1 Load, filter, and sample data
# Load Yelp data. Dataset can be downloaded [here](https://www.yelp.com/dataset/download). After entering your contact information, select "Download JSON."
# +
# %%time
PATH = "../data/yelp_dataset/"
yelp_data = []
r_dtypes = {"review_id":str,
"user_id":str,
"business_id":str,
"stars": np.int32,
"date":str,
"text":str,
"useful": np.int32,
"funny": np.int32,
"cool": np.int32}
drop = ['review_id', 'user_id', 'useful', 'funny', 'cool']
query = "date >= '2017-12-01' and (stars==1 or stars ==5)"
with open(f"{PATH}yelp_academic_dataset_review.json", "r") as f:
reader = pd.read_json(f, orient="records", lines=True, dtype=r_dtypes, chunksize=1000)
for chunk in reader:
reduced_chunk = chunk.drop(columns=drop).query(query)
yelp_data.append(reduced_chunk)
yelp_data = pd.concat(yelp_data, ignore_index=True)
# -
yelp_data.shape
# Merge reviews and metadata on businesses.
yelp_business = pd.read_json(f"{PATH}yelp_academic_dataset_business.json", orient="records", lines=True)
yelp_business.shape
# Select businesses in Portland and Atlanta only.
yelp_business = yelp_business[yelp_business.city.isin(["Portland", "Atlanta"])]
yelp_business.shape
# Merge review and business data.
yelp_merged = yelp_data.merge(yelp_business, on='business_id')
yelp_merged.shape
yelp_merged = yelp_merged.rename(columns={"stars_x":"stars"})
# Split string category column into list.
categories = [c.split(",") for c in yelp_merged.categories if c is not None]
c_count = Counter(c.strip() for c_list in categories for c in set(c_list))
c_count.most_common()[:30]
yelp_categorized = yelp_merged.assign(categories=yelp_merged['categories'].str.split(',')).explode('categories')
yelp_categorized.categories = yelp_categorized.categories.str.strip()
yelp_categorized.shape
# Filter data on category for Restaurants and 'Health & Medical'.
yelp_filtered = yelp_categorized[yelp_categorized.categories.isin(['Restaurants','Health & Medical'])]
yelp_filtered.shape
# Sample data, so that we get 1000 of each category and of 1 or 5 stars.
s_size = 1000
rs = 42
sample = yelp_filtered.groupby(["stars", "categories"]).sample(n=s_size, random_state=rs).reset_index()
sample.shape
stopword_list=stopwords.words('english')
# ### 1.2 Phrase data, remove special formatting and stopwords
# Initialize tokenizer.
tokenizer = ToktokTokenizer()
# Remove stopwords before denoising, lemmatizing and removing special characters.
# +
# %%time
sample['review_clean']= [remove_stopwords(r, stopword_list, tokenizer) for r in sample['text']]
# -
# Denoise, remove special characters, lemmatize.
# +
# %%time
sample['review_clean']=sample['review_clean'].apply(denoise_text)
sample['review_clean']=sample['review_clean'].apply(remove_special_characters)
sample['review_clean']=sample['review_clean'].apply(simple_lemmatizer)
# -
# Remove stopwords again, after other preprocessing.
# +
# %%time
sample['review_clean']= [remove_stopwords(r, stopword_list, tokenizer) for r in sample['review_clean']]
# -
# Find phrases.
# Load phrases found in data during finetuning. This will require that you have a folder called 'embeddings' in the root of the 'paper' folder with files that can be found here. Alternatively, you can finetune and phrase vectors yourself, using the 'FinetuneYelp.py' script.
PHRASING = True
MIN = 500
THRESHOLD = 200
# +
# %%time
if PHRASING:
sample['review_clean']= get_phrases([tokenizer.tokenize(i) for i in sample['review_clean']],
min_count = MIN,
threshold = THRESHOLD,
save=False,
load=True,
PATH="../embeddings/")
# -
# Data _before_ preprocessing and phrasing.
sample['text'][6]
# Data _after_ preprocessing and phrasing.
#sample['review_clean'][0]
" ".join(sample['review_clean'][6])
sample["sentiment"] = ['positive' if s == 5 else 'negative' for s in sample['stars']]
sample[["sentiment", "stars"]]
sample.head()
sample_sorted = sample.sort_values(["sentiment", "categories"]).reset_index()
sample_sorted
# ### 1.3 Separate pos and neg reviews
pos = sample_sorted[sample_sorted.sentiment == "positive"].reset_index(drop=True)
neg = sample_sorted[sample_sorted.sentiment == "negative"].reset_index(drop=True)
pos = pos.review_clean.tolist()
neg = neg.review_clean.tolist()
# ## 2. WMD
# ### 2.1 Tokenize data
# Separate data into positive and negative reviews and tokenize each review.
pos_tok = list(map(lambda x: tokenize(x, tokenizer), pos))
neg_tok = list(map(lambda x: tokenize(x, tokenizer), neg))
pos_sample = [" ".join(doc) for doc in pos_tok]
neg_sample = [" ".join(doc) for doc in neg_tok]
print(len(pos_sample))
print(len(neg_sample))
# ### 2.2 Load pretrained Google News W2V model
# Load word vectors. You can use vectors finetuned for the paper (found here), pretrained vectors from Google (found here), or finetune vectors yourself using the 'FineTuneYelp.py' script.
finetuned = True
if not finetuned:
print("Loading GoogleNews Vectors")
# %time model = KeyedVectors.load_word2vec_format('../embeddings/GoogleNews-vectors-negative300.bin.gz', binary=True)
else:
print("Loading GoogleNews Vectors finetuned using Yelp review data.")
# %time model = KeyedVectors.load_word2vec_format('../embeddings/yelp_w2v.txt', binary=False)
model.distance("taco", "burrito")
model.distance("quesadilla","tostada")
model.distance("sushi", "sashimi")
# ### 2.3 Load corpus and remove OOV words
# Form a corpus for running the TfIdf vectorizer with l1 normalization.
# +
# %%time
corpus = pos_sample + neg_sample
vectorizer = TfidfVectorizer(use_idf=True, tokenizer=tfidf_tokenize, norm='l1')
vectorizer.fit(corpus)
# -
# Find lingering out-of-vocabulary words.
# %time oov = [word for word in vectorizer.get_feature_names() if word not in model.key_to_index.keys()]
len(oov)
print(oov[:50])
# Remove oov words.
# %time pos_sample = list(map(lambda x: remove_oov(x, tokenizer, oov), pos_sample[:2000]))
# %time neg_sample = list(map(lambda x: remove_oov(x, tokenizer, oov), neg_sample[:2000]))
pos_sample[5]
# Now transform the corpus into bag-of-words.
# %time
pos_nbow = vectorizer.transform(pos_sample)
neg_nbow = vectorizer.transform(negT_sample)
# Tokenize samples.
pos_tok = list(map(lambda x: tokenize(x, tokenizer), pos_sample))
neg_tok =list(map(lambda x: tokenize(x, tokenizer), neg_sample))
print(pos_tok[5][:20])
# %time oov_ = [word for word in vectorizer.get_feature_names() if word not in model.key_to_index.keys()]
len(oov_)
# ### 2.4 Get features and embeddings
features = vectorizer.get_feature_names()
word2idx = {word: idx for idx, word in enumerate(vectorizer.get_feature_names())}
idx2word = {idx: word for idx, word in enumerate(vectorizer.get_feature_names())}
# Get the embedding matrix "E" for all features.
E = model[features]
# ### 2.5 Cluster
# In order to make the results of the WMD model more interpretable, we add the option to inspect the output not only by individual words, but also by *word clusters*. We do this by clustering the input words with two different algorithms
# and assigning each word to a cluster.
# #### 2.5.1 Kmeans
# First, we select the number of clusters we want to search to determine the optimal size of K.
# +
# %%time
K = range(10,210, 10)
# -
# Now, let's search through the different Ks to see what the optimal K is using the [silhouette score](https://towardsdatascience.com/silhouette-coefficient-validating-clustering-techniques-e976bb81d10c). The silhouette score can take on values between -1 and 1, with values closer to 1 indicating nicely separated clusters. We are also getting the sum of squared differences in order to plot the so called [elbow plot](https://blog.cambridgespark.com/how-to-determine-the-optimal-number-of-clusters-for-k-means-clustering-14f27070048f).
# +
# %%time
wvec_ssd, wvec_silhouette = kmeans_search(E, K)
# -
# Plotting the elbow plot. A good heuritic for K is if there are any clear and "sharp" corners in the plot.
plot_kmeans(K,wvec_ssd,"elbow")
# Plotting the silhouette score, with values closer to 1 indicating nicely separated clusters.
plot_kmeans(K,wvec_silhouette,"silhouette")
# #### 2.5.2 T-SNE + Kmeans
# As we saw above, using Kmeans on the raw embeddings did not produce very satisfactory results. To get clusters that are better separated, we will try to first reduce the vectors into two dimensions using the t-SNE algorithm, and then running Kmeans on these reduced embeddings.
method='barnes_hut'
n_components = 2
verbose = 1
E_tsne = TSNE(n_components=n_components, method=method, verbose=verbose).fit_transform(E)
# A scatterplot of the reduced vectors indicates that there are indeed quite clear (albeit very many) clusters in the data.
plt.scatter(E_tsne[:, 0], E_tsne[:, 1], s=1);
# Run the same diagnostics as above, using the silhouette score and the elbow plot.
# +
# %%time
tsne_ssd, tsne_silhouette = kmeans_search(E_tsne, K)
# -
# Elbow plot is much clearer than above, with a break at 20 or 25, or alternatively closer to 100.
plot_kmeans(K,tsne_ssd,"elbow")
# The silhouette score indicates that results are better than above, but the differences in values are quite nominal.
plot_kmeans(K,tsne_silhouette,"silhouette")
# #### 2.5.3 PCA + Kmeans
# While t-SNE gave us promising diagnostics, another approach would be to use dimensionality reduction with PCA. However, as can be seen inspecting the diagnostics below, PCA reduced vectors are actually much less clearly separated than the t-SNE reduced vectors.
n_components = 0.9
verbose = 1
pca_fit = PCA(n_components = n_components).fit(E)
print(len(pca_fit.explained_variance_ratio_))
print(pca_fit.explained_variance_ratio_)
print(np.sum(pca_fit.explained_variance_ratio_))
E_pca = pca_fit.transform(E)
plt.scatter(E_pca[:, 0], E_pca[:, 1], s=1);
# +
# %%time
pca_ssd, pca_silhouette = kmeans_search(E_pca, K)
# -
plot_kmeans(K,pca_ssd,"elbow")
plot_kmeans(K,pca_silhouette,"silhouette")
# #### 2.5.4 UMAP + Kmeans
# Finally, we also try the increasingly popular UMAP dimensionality reduction technique. The results are comparible to the t-SNE reduced vectors, but because t-SNE is better known, we stick with that algorithm. However, UMAP could be a good alternative to explore in more detail in the future.
# +
# %%time
metric = 'cosine'
dm = cdist(E, E, metric)
np.fill_diagonal(dm, 0)
# -
is_valid_dm(dm)
mean, std = np.mean(dm), np.std(dm)
print(mean, std)
min_dist=mean - 2*std
n_neighbors = int(0.001*len(E))
n_components=2
print(f"Min distance: {min_dist}")
print(f"N. neighbors: {n_neighbors}")
print(f"N. compontents: {n_components}")
# +
# %%time
E_umap = umap.UMAP(
n_neighbors=n_neighbors,
min_dist=min_dist,
n_components=n_components,
random_state=42,
verbose=verbose
).fit_transform(E)
# -
plt.scatter(E_umap[:, 0], E_umap[:, 1], s=1);
# +
# %%time
umap_ssd, umap_silhouette = kmeans_search(E_umap, K)
# -
plot_kmeans(K,umap_ssd,"elbow")
plot_kmeans(K,umap_silhouette,"silhouette")
# #### 2.5.5 Choose clustering model
# Becaus silhouette scores were so even across K and because the elbow plots were ultimately quite ambigious, we use 100 clusters as a simple heuristic.
k_base = 100
k_pca = k_base
k_tsne = k_base
k_umap = k_base
# +
# %%time
km_base = cluster.KMeans(n_clusters=k_base,max_iter=300).fit(E)
labels = km_base.labels_
centroids = km_base.cluster_centers_
km_pca = cluster.KMeans(n_clusters=k_pca,max_iter=300).fit(E_pca)
labels_pca = km_pca.labels_
km_tsne = cluster.KMeans(n_clusters=k_tsne,max_iter=300).fit(E_tsne)
labels_tsne = km_tsne.labels_
km_umap = cluster.KMeans(n_clusters=k_umap,max_iter=300).fit(E_umap)
labels_umap=km_umap.labels_
# -
# Create an index that maps each word to a cluster.
word2cluster = {features[idx]: cl for idx, cl in enumerate(labels)}
print(take(10, word2cluster.items()))
# Now, conversely, create an index that maps each cluster to a word.
cluster2words = defaultdict(list)
for key, value in word2cluster.items():
cluster2words[value].append(key)
print(len(cluster2words[0]))
print(cluster2words[0][:100])
print(len(cluster2words[1]))
print(cluster2words[1][:100])
# ### 2.6 Initialize documents
# Transform all reviews into "documents", each with a set of weights per word in the corpus ("nbow"), the sum of these weights ("weights_sum"), the indeces of the words in the documents ("idxs") and the word vectors corresponding to each word ("vecs").
pos_nbow[0]
# +
# %%time
pos_docs, neg_docs = [], []
for idx, doc in enumerate(pos_tok):
pos_docs.append(Document(doc, pos_nbow[idx], word2idx, E))
for idx, doc in enumerate(neg_tok):
neg_docs.append(Document(doc, neg_nbow[idx], word2idx, E))
# -
pos_docs[0].nbow
pos_docs[0].weights_sum
pos_docs[0].idxs[:10]
pos_docs[0].vecs[:1][0][:10]
# ### 2.7 Linear-Complexity Relaxed WMD (LC-RWMD)
# Run the [Linear-Complexity Relaxed WMD](https://arxiv.org/abs/1711.07227) to get the distances between all positive and all negative reviews.
metric = "cosine"
lc_rwmd = LC_RWMD(pos_docs, neg_docs,pos_nbow,neg_nbow,E)
# %time lc_rwmd.get_D(metric=metric)
# ### 2.8 Gale-Shapeley Pairing
# Use the [Gale-Shapeley matching algorithm](https://en.wikipedia.org/wiki/Gale%E2%80%93Shapley_algorithm) to find the optimal pairs between positive and negative reviews. This iterates over all the reviews and finds the set of matches that pairs each review with its optimal match given that all positive reviews have to be matched with a negative review and vice versa. The output is a dictionary of key-value pairs, where each pair represents an optimal match.
#
# Alternatively, you can run full pairs or random pairs.
# +
# Options: 'gale_shapeley','random','full'
pairing = 'gale_shapeley'
# +
# %%time
if pairing == 'gale_shapeley':
print("Running Gale-Shapeley pairing.")
matcher = Matcher(lc_rwmd.D)
engaged = matcher.matchmaker()
print(f"Pairing is stable: {matcher.check()}")
pairs = [(k, v) for k, v in engaged.items()]
if pairing == 'random':
print("Running random pairing.")
pos_idx = list(range(0,len(pos_docs)))
neg_idx = list(range(0,len(neg_docs)))
shuffle(pos_idx)
shuffle(neg_idx)
pairs = list(zip(pos_idx, neg_idx))
if pairing == 'full':
print("Running full pairing.")
pos_idx = list(range(0,len(pos_docs)))
neg_idx = list(range(0,len(neg_docs)))
pairs = [(i,j) for i in pos_idx for j in neg_idx]
# -
# Let's look at the output of our pairing (by default Gale-Shapeley, see above):
print(pairs[:20])
example_pairs = [(' '.join(word for word in pos_docs[p[0]].words),
' '.join(word for word in neg_docs[p[1]].words))
for p in pairs]
example_pairs[1][0]
example_pairs[1][1]
len(pairs)
# ### 2.9 Pairwise WMD with Baseline Kmeans
# Calculate the pairwise distances between the documents selected by the Galey-Shapeley algorithm _without_ returning the flow between individual words.
# +
# %%time
wmd_pairs = WMDPairs(pos_docs,neg_docs,pairs,E,idx2word, metric=metric)
wmd_pairs.get_distances(decompose=False,thread=False,relax=True)
# -
# The return value is a matrix of distances between the document pairs.
np.max(wmd_pairs.distances)
wmd_dists = np.concatenate(wmd_pairs.distances)
wmd_dists[wmd_dists != 0].mean()
# Calculate the pairwise distances between the documents selected by the Galey-Shapeley algorithm, this time also returning the flow between individual words.
# +
# %%time
wmd_pairs_flow = WMDPairs(pos_docs,neg_docs,pairs,E,idx2word, metric=metric)
wmd_pairs_flow.get_distances(decompose = True,
sum_clusters = True,
w2c = word2cluster,
c2w = cluster2words,
thread = False,
relax = True)
# -
# Now we have three return values.
#
# The first one is again a matrix of distances between the document pairs.
np.max(wmd_pairs_flow.distances)
wmd_flow_dists = np.concatenate(wmd_pairs_flow.distances)
wmd_flow_dists[wmd_flow_dists != 0].mean()
# The second return value is a list of tuples with all the words that contributed the most to the distance from the positive documents to the negative ones. These are _not_ sorted from high to low or vice versa.
take(10, wmd_pairs_flow.wd_source.items())
# The third return value is a list of tuples with all the words that contributed the most to the distance from the negative documents to the positive ones. Again, these are _not_ sorted from high to low or vice versa.
take(10, wmd_pairs_flow.wd_sink.items())
take(10, wmd_pairs_flow.cd_source.items())
take(10, wmd_pairs_flow.cd_sink.items())
{k: v for k, v in sorted(wmd_pairs_flow.cd_source.items(), key=lambda item: item[1], reverse=True)[:10]}
# Calculate the WMD with difference (see paper).
wmd_pairs_flow.get_differences()
# ### 3.1 Intepreting pairwise WMD flows
# Now, let's sort the distances of the words that created the most distance from the positive to the negative reviews.
top_words_source = {k: v for k, v in sorted(wmd_pairs_flow.wd_source_diff.items(), key=lambda item: item[1], reverse=True)[:30]}
top_words_source_df = pd.DataFrame.from_dict(top_words_source, orient='index', columns = ["distance"])
top_words_source_df['cost'] = top_words_source_df.distance.round(2)
top_words_source_df
# Next, let's see what added most distance when moving from the negative to the positive reviews.
# +
top_words_sink = {k: v for k, v in sorted(wmd_pairs_flow.wd_sink_diff.items(), key=lambda item: item[1], reverse=True)[:30]}
top_words_sink_df = pd.DataFrame.from_dict(top_words_sink, orient='index', columns = ["distance"])
top_words_sink_df['distance'] = top_words_sink_df.distance.round(2)
top_words_sink_df
# -
# Finally, let's look at the distances between the two sets by clustering similar words, in order to get a better sense of what kind of "topics" that separate them. Each cluster has a weight that matches the sum of the words belonging to that cluster. We choose *n* top clusters to inspect. To make the clusters interpretable, we also represent each of them by *m* keywords, selected based on the cost they individually add between the two sets.
n_clusters = 10
n_words = 10
c1 = output_clusters(wd=wmd_pairs_flow.wd_source_diff.items(),
cd=wmd_pairs_flow.cd_source.items(),
c2w=cluster2words,
n_clusters=n_clusters,
n_words=n_words)
c2 = output_clusters(wd=wmd_pairs_flow.wd_sink_diff.items(),
cd=wmd_pairs_flow.cd_sink.items(),
c2w=cluster2words,
n_clusters=n_clusters,
n_words=n_words)
# Positive to negative clusters.
c1
# Negative to positive clusters.
c2
sample.shape
# ### 3.2. Explore differences by category
# Because our sample is stratified by city and by category, we also offer this small example of how to explore the differences by city and category using facetted boxplots.
source_dists = pd.DataFrame(wmd_pairs_flow.source_feat)
source_dists.index = [p[0] for p in pairs]
source_dists = source_dists.sort_index()
source_dists = source_dists[c1.columns]
source_dists['categories'] = sample[:2000].categories
Counter(source_dists.categories)
source_dists_long = pd.melt(source_dists, id_vars=['categories']).rename(columns={"variable":"cluster"})
source_dists_long.head()
source_dists_long = source_dists_long[source_dists_long.value != 0]
g = sns.catplot(x="categories",
y="value",
col="cluster",
data=source_dists_long,
kind="box",
height=5,
aspect=.7,
col_wrap=5,
margin_titles=True);
g.map_dataframe(sns.stripplot,
x="categories",
y="value",
palette=["#404040"],
alpha=0.2, dodge=True)
g.set_axis_labels("Category", "Distance")
for ax in g.axes.flatten():
ax.tick_params(labelbottom=True)
sink_dists = pd.DataFrame(wmd_pairs_flow.sink_feat)
sink_dists.index = [p[1] for p in pairs]
sink_dists = sink_dists.sort_index()
sink_dists = sink_dists[c2.columns]
sink_dists['categories'] = sample[2000:4000].categories.tolist()
sink_dists_long = pd.melt(sink_dists, id_vars=['categories']).rename(columns={"variable":"cluster"})
sink_dists_long.head()
sink_dists_long = sink_dists_long[sink_dists_long.value != 0]
g = sns.catplot(x="categories",
y="value",
col="cluster",
data=sink_dists_long,
kind="box",
height=5,
aspect=.7,
col_wrap=5,
margin_titles=True);
g.map_dataframe(sns.stripplot,
x="categories",
y="value",
palette=["#404040"],
alpha=0.2, dodge=True)
g.set_axis_labels("Category", "Distance")
for ax in g.axes.flatten():
ax.tick_params(labelbottom=True)
# ## 4 Better clusters with t-SNE
# Repeat the steps in section 3, but with clusters based on t-SNE reduced embeddings.
word2cluster_t = {features[idx]: cl for idx, cl in enumerate(labels_tsne)}
print(take(10, word2cluster_t.items()))
# Now, conversely, create an index that maps each cluster to a word.
cluster2words_t = defaultdict(list)
for key, value in word2cluster_t.items():
cluster2words_t[value].append(key)
# +
# %%time
wmd_pairs_flow_t = WMDPairs(pos_docs,neg_docs,pairs,E,idx2word)
wmd_pairs_flow_t.get_distances(decompose = True,
sum_clusters = True,
w2c = word2cluster_t,
c2w = cluster2words_t,
thread = False,
relax = True)
# -
wmd_pairs_flow_t.get_differences()
c1_t = output_clusters(wd=wmd_pairs_flow_t.wd_source_diff.items(),
cd=wmd_pairs_flow_t.cd_source.items(),
c2w=cluster2words_t,
n_clusters=n_clusters,
n_words=n_words)
c2_t = output_clusters(wd=wmd_pairs_flow_t.wd_sink_diff.items(),
cd=wmd_pairs_flow_t.cd_sink.items(),
c2w=cluster2words_t,
n_clusters=n_clusters,
n_words=n_words)
c1_t
c2_t
| paper/notebooks/Yelp_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import cv2 as cv
import json
"""缩小图像,方便看效果
resize会损失像素,造成边缘像素模糊,不要再用于计算的原图上使用
"""
def resizeImg(src):
height, width = src.shape[:2]
size = (int(width * 0.3), int(height * 0.3))
img = cv.resize(src, size, interpolation=cv.INTER_AREA)
return img
"""找出ROI,用于分割原图
原图有四块区域,一个是地块区域,一个是颜色示例区域,一个距离标尺区域,一个南北方向区域
理论上倒排后的最大轮廓的是地块区域
"""
def findROIContours(src):
copy = src.copy()
gray = cv.cvtColor(copy, cv.COLOR_BGR2GRAY)
# cv.imshow("gray", gray)
# 低于thresh都变为黑色,maxval是给binary用的
# 白底 254, 255 黑底 0, 255
threshold = cv.threshold(gray, 0, 255, cv.THRESH_BINARY)[1]
# cv.imshow("threshold", threshold)
contours, hierarchy = cv.findContours(threshold, cv.RETR_EXTERNAL, cv.CHAIN_APPROX_SIMPLE)
sortedCnts = sorted(contours, key = cv.contourArea, reverse=True)
# cv.drawContours(copy, [maxCnt], -1, (255, 0, 0), 2)
# cv.imshow("roi contours", copy)
return sortedCnts
"""按照mask,截取roi
"""
def getROIByContour(src, cnt):
copy = src.copy()
# black background
mask = np.zeros(copy.shape[:2], np.uint8)
mask = cv.fillConvexPoly(mask, cnt, (255,255,255))
# cv.imshow("mask", resizeImg(mask))
# print(mask.shape)
# print(copy.dtype)
roi = cv.bitwise_and(copy, copy, mask=mask)
# cv.imshow("roi", roi)
# white background for none roi and fill the roi's backgroud
mask = cv.bitwise_not(mask)
whiteBg = np.full(copy.shape[:2], 255, dtype=np.uint8)
whiteBg = cv.bitwise_and(whiteBg, whiteBg, mask=mask)
whiteBg = cv.merge((whiteBg,whiteBg,whiteBg))
# cv.imshow("whiteBg", resizeImg(whiteBg))
roiWithAllWhite = cv.bitwise_or(roi, whiteBg)
return roiWithAllWhite
"""找出所有的地块轮廓
"""
def findAllBlockContours(src):
copy = src.copy()
contours = findExternalContours(copy)
return contours
"""找出颜色示例里的颜色BGR
根据限定长,高,比例来过滤出实例颜色区域
"""
def findBGRColors(cnts):
W_RANGE = [170,180]
H_RANGE = [75, 85]
RATIO_RANGE = [0.40, 0.50]
colors = []
# TODO 如果可以知道颜色示例的个数可以提前统计退出循环
for cnt in cnts:
x,y,w,h = cv.boundingRect(cnt)
ratio = round(h/w, 2)
if ratio > RATIO_RANGE[0] and ratio < RATIO_RANGE[1] \
and w > W_RANGE[0] and w < W_RANGE[1] \
and h > H_RANGE[0] and h < H_RANGE[1]:
# print(ratio,x,y,w,h)
# 因为,原图色块矩形的周边和mask出来的颜色区都有模糊渐变的线
# 无法使用cv.mean(colorRegion, meanMask)来计算实际颜色
# 所以,取矩形的中心点的颜色最为准确
cx,cy = round(x+w/2), round(y+h/2)
bgr = img_white_bg[cy, cx]
# print(bgr)
colors.append(bgr)
return colors
def drawnForTest(img, contours, rect=False):
img = img.copy()
for i in contours:
if rect:
x, y, w, h = cv.boundingRect(i)
cv.rectangle(img, (x, y), (x + w, y + h), (255, 0, 0), 2)
cv.putText(img, 'Area' + str(cv.contourArea(i)), (x+5,y+15), cv.FONT_HERSHEY_PLAIN, 1,(255,0,0), 1, cv.LINE_AA)
else:
cv.drawContours(img, [i], -1, (0, 255, 0), 2)
cv.imshow("detect", resizeImg(img))
cv.waitKey(0)
"""Find Original Contours
Find Original Contours from source image, we only need external contour.
Args:
src: source image
Returns:
Original contours
"""
def findExternalContours(src):
# 必须是白色背景的图片,如果是黑色背景,将黑色改成白色
# src[np.where((src == [0,0,0]).all(axis = 2))] = [255,255,255]
# preprocess, remove noise, a lot noise on the road
gray = cv.cvtColor(src, cv.COLOR_BGR2GRAY)
# 测试这里道理有没有必要高斯
# blur = cv.GaussianBlur(gray, (3,3), 0)
thresVal = 254
maxVal = 255
ret,thresh1 = cv.threshold(gray, thresVal, maxVal, cv.THRESH_BINARY)
kernel = np.ones((7,7),np.uint8)
morph = cv.morphologyEx(thresh1, cv.MORPH_CLOSE, kernel)
# ??threshold怎么计算的?
edges = cv.Canny(morph,100,200)
# edges找出来,但是是锯齿状,会在找轮廓时形成很多点,这里加一道拉普拉斯锐化一下
edges = cv.Laplacian(edges, -1, (3,3))
contours, hierarchy = cv.findContours(edges, cv.RETR_EXTERNAL, cv.CHAIN_APPROX_SIMPLE)
# contours, hierarchy = cv.findContours(edges, cv.RETR_EXTERNAL, cv.CHAIN_APPROX_SIMPLE)
# cv.imshow('gray', resizeImg(gray))
# cv.imshow('thresh1', resizeImg(thresh1))
# cv.imshow('edges', resizeImg(edges))
# cv.imshow('opening', opening)
# cv.imshow('opening', resizeImg(opening))
return contours, hierarchy
"""根据找出的轮廓和层级关系计算地块和色块的父子关系
"""
def getBlockColorTree(copy, blockCnts, hierarchy):
# print(hierarchy)
# hierarchy [Next, Previous, First_Child, Parent]
currentRootIndex = -1
rootRegions = {}
for i,cnt in enumerate(blockCnts):
x,y,w,h = cv.boundingRect(cnt)
cntArea = cv.contourArea(cnt)
if cntArea > 1000:
continue
cv.putText(copy, str(i), (x+5,y+15), cv.FONT_HERSHEY_PLAIN, 1,(255,0,0), 1, cv.LINE_AA)
print(i, hierarchy[0][i])
if hierarchy[0][i][3] == -1:
# root region
currentRootIndex = i
if currentRootIndex == len(blockCnts):
break
rootRegion = {'index': i, 'contour': cv.contourArea(blockCnts[currentRootIndex]), 'childRegion': []}
rootRegions[currentRootIndex] = rootRegion
elif hierarchy[0][i][3] == currentRootIndex:
rootRegions[currentRootIndex]['childRegion'].append({'index': i, 'contour': cv.contourArea(cnt)})
cv.imshow("blockCnts with debug info", resizeImg(copy))
print(rootRegions)
data2 = json.dumps(rootRegions, sort_keys=True, indent=4, separators=(',', ': '))
print(data2)
"""使用颜色来分块,并返回所有地块和色块父子关系
debug 只演示前三个地块的识别过程,可以通过debugFrom:debugLen来调整debug开始位置和长度
"""
def findColorRegionsForAllBlocks(img_white_bg, blockCnts, debug=False, debugFrom=0, debugLen=3):
filteredBlockCnts = [cnt for cnt in blockCnts if cv.contourArea(cnt) > 100]
if debug:
filteredBlockCnts = filteredBlockCnts[debugFrom:debugLen]
for blockCnt in filteredBlockCnts:
findColorRegionsForBlock(img_white_bg, blockCnt, debug)
"""根据threshold重新计算BGR的值
"""
def bgrWithThreshold(bgr, threshold):
newBgr = []
for x in bgr.tolist():
if x + threshold < 0:
newBgr.append(0)
elif x + threshold > 255:
newBgr.append(255)
else:
newBgr.append(x + threshold )
return newBgr
"""使用颜色来找出单个地块内的色块
"""
def findColorRegionsForBlock(img_white_bg, blockCnt, debug=False):
blockWithColorsDict = {'area': cv.contourArea(blockCnt), 'points':[] , 'children': []}
blockRegion = getROIByContour(img_white_bg, blockCnt)
if debug:
cv.imshow("blockRegions", np.hstack([resizeImg(img_white_bg), resizeImg(blockRegion)]))
colorCnts = []
for bgr in bgrColors:
# 图片里的颜色可能和示例颜色不相等,适当增加点阈值来防色差
threshold = 5
lower = np.array(bgrWithThreshold(bgr, -threshold), dtype="uint8")
upper = np.array(bgrWithThreshold(bgr, threshold), dtype="uint8")
# 根据阈值找到对应颜色区域,黑底白块
mask = cv.inRange(blockRegion, lower, upper)
# cv.imshow("mask", resizeImg(mask))
# 过滤出于颜色匹配的色块
nonZeroCount = cv.countNonZero(mask)
# print('none zero count', nonZeroCount)
if nonZeroCount == 0:
continue
contours, hierarchy = cv.findContours(mask.copy(), cv.RETR_EXTERNAL, cv.CHAIN_APPROX_SIMPLE)
# print('external', len(contours))
# print(hierarchy)
colorCnts.extend(contours)
# contours, hierarchy = cv.findContours(mask.copy(), cv.RETR_TREE, cv.CHAIN_APPROX_SIMPLE)
# print('tree', len(contours))
# print(hierarchy)
if debug:
# 黑白变白黑
mask_inv = 255 - mask
# cv.imshow("mask_inv", resizeImg(mask_inv))
# 展示图片
output = cv.bitwise_and(blockRegion, blockRegion, mask=mask_inv)
# cv.drawContours(output, contours, -1, (0, 0, 255), 3)
cv.imshow("images", np.hstack([resizeImg(blockRegion), resizeImg(output)]))
cv.waitKey(0)
# 色块在同一个地块内也可能是多个的,色块内又内嵌了色块
# TODO 嵌套的
# colorCnts 循环递归,不到叶子节点,所有的色块统一当作地块继续处理,这样就可以解决嵌套问题
# 所以要先构建一个三层数据模型
# 第一层 是一个列表 存放所有的地块
# 第二层 是以色块为节点,色块内无嵌套,则为叶子节点,有嵌套,色块升级为地块,继续深入查找色块,直到没有内嵌,成为叶子节点
# 第三层 是以色块为叶子节点
colorDicts = []
for colorCnt in contours:
colorDict = {'area': cv.contourArea(colorCnt), 'points':[], 'color': bgr.tolist()}
colorDicts.append(colorDict)
blockWithColorsDict['children'].extend(colorDicts)
# print(blockWithColorsDict)
jsonData = json.dumps(blockWithColorsDict, sort_keys=True, indent=4, separators=(',', ': '))
print(jsonData)
return colorCnts
# 用于找ROI
img = cv.imread('data_hierarchy2.png')
# 用于真实计算,
# 1. 色调区间误差,原图4识别最高,图3识别一般,需要增加threshold到5,
# 2. 间隙与边框误差,色块面积总和与地块面积相差3000左右,应该是线的面积没计算进去
# 3.
img_white_bg = cv.imread('data_hierarchy4.png')
# 将图片按照轮廓排序,最大的是总地块
# 按照原图中的比例和实际距离来分割图片,参考findBGRColors的计算方式
sortedCnts = findROIContours(img)
# print(len(sortedCnts[2:]))
# drawnForTest(img_white_bg, sortedCnts[3], rect=True)
# print(sortedCnts[3])
print(cv.boundingRect(sortedCnts[3]))
print(img_white_bg.shape)
# 2401 * 3151
# 670
px_km_scale = 670/1000
area_px = (2401*3151)
area_km2 = (2401*px_km_scale*3151*px_km_scale)
print(area_px/area_km2)
print(1/(px_km_scale*px_km_scale))
# 获取总地块
rootRegion = getROIByContour(img_white_bg, sortedCnts[0])
# cv.imshow("rootRegion", resizeImg(rootRegion))
# 找出色块的颜色
bgrColors = findBGRColors(sortedCnts[1:])
# print(bgrColors)
# 找出地块
copy = rootRegion.copy()
blockCnts, hierarchy = findAllBlockContours(copy)
# print(len(blockCnts))
# drawnForTest(img_white_bg, blockCnts, rect=False)
# 通过颜色来检测地块内色块
# findColorRegionsForAllBlocks(img_white_bg, blockCnts, debug=True, debugLen=1)
# findColorRegionsForAllBlocks(img_white_bg, blockCnts)
# 根据轮廓找地块的方法首先hierarchy的转换父子关系,还有很多小的干扰项待解决
# getBlockColorTree(copy, blockCnts, hierarchy)
cv.waitKey(0)
cv.destroyAllWindows()
# +
a = [1]
b = [2,3]
a.extend(b)
a
c=1
-c
| opencv_tutorial/contour-hierarchy/contour-hierachy-bgr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Meiyi-Wu/blooket-hack/blob/main/Numpy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="uLHV3CxqzVgn" outputId="e0d62d32-dd21-4124-90ac-1594f831524e"
import numpy as np
np_city = np.array(["NYC","LA","Miami","Houston"])
np_city_state = np.array([[["NYC","LA","Miami","Houston"],["NY","CA","FL","TX"],[100,200,50,5]]])
print(np_city_state)
np_city_state.ndim
# + colab={"base_uri": "https://localhost:8080/"} id="NC5oSe7X_GJR" outputId="494b908c-be4c-491a-eb01-8c1028cba6cb"
trial_1=[10,15,17,26]
trial_2=[2]
np_trial_1=np.array(trial_1)
np_trial_2=np.array(trial_2)
np_total =np_trial_1/np_trial_2
np_total
all = np.array([[10,15,17,26],[12,11,21,24]])
all
#first_trial = all[0]
#second_trial = all[1]
#first_element = all[0,1]
#first_element
# + colab={"base_uri": "https://localhost:8080/"} id="1IeRDroA_GNY" outputId="ced9c145-7c36-41c2-8455-0308fc6b286b"
trial_1=[10,15,17,26]
trial_2=[2]
np_trial_1=np.array(trial_1)
np_trial_2=np.array(trial_2)
np_total =np_trial_1/np_trial_2
np_total
all = np.array([[10,15,17,26],[12,11,21,24]])
all[1,0]
all[1,3]
first_trial = all[0]
second_trial = all[1]
print(second_trial[1])
all[1,1]
# + colab={"base_uri": "https://localhost:8080/"} id="8kfIpImP_Gmb" outputId="1f66ada0-d229-42cc-c143-a5b9e9cbc564"
test_scores = np.array([[83,71,57,63],[54,68,81,45]])
pass_scores = test_scores >60
print(pass_score)
# + colab={"base_uri": "https://localhost:8080/"} id="KaqjkEtC_G0J" outputId="cb48f7e4-343b-47fb-a6d7-547a5164afae"
test_scores= np.array([[83,71,57,63], [54,68,81,45]])
pass_score = test_scores > 60
array_pass = test_scores[pass_score]
print(sorted(array_pass))
# + [markdown] id="_gVaZkQM_HMG"
#
# + colab={"base_uri": "https://localhost:8080/"} id="VyqKW3D9_Irz" outputId="1883dcec-3206-4823-e5f8-9eed3ef22c13"
NYC_Borough = np.array(["manhattan", "Bronx", "Brooklyn", "Queens", "Staten Island"])
NYC_Borough
Boroughs_In_NYC = NYC_Borough
Boroughs_In_NYC in NYC_Borough
# + colab={"base_uri": "https://localhost:8080/"} id="xi2l5VYR_I_X" outputId="084740b0-bbe7-4a08-c41c-5abb4be60cc0"
Boroughs_In_NYC
view_Boroughs_In_NYC = Boroughs_In_NYC.view()
view_Boroughs_In_NYC[4] = 'Central Park'
print(view_Boroughs_In_NYC)
print(Boroughs_In_NYC)
# + colab={"base_uri": "https://localhost:8080/"} id="kVy5_56e_J2B" outputId="5cc24779-911c-4c4d-c9dd-bbc6159a23dc"
copy_of_Boroughs_In_NYC = NYC_Borough.copy()
copy_of_Boroughs_In_NYC
NYC_Borough
copy_of_Boroughs_In_NYC[4] = 'Staten Island'
copy_of_Boroughs_In_NYC
NYC_Borough
# + colab={"base_uri": "https://localhost:8080/"} id="Mty8Ialv_DyI" outputId="6fa840ac-0f75-4082-a774-4deeb7e4ab4e"
np_sqrt = np.sqrt([2,4,9,16])
np_sqrt
from numpy import pi
np.sin(pi/2)
np.cos(pi)
np.floor([1.5,2.3,6.7])
np.exp([0,1,5])
# + colab={"base_uri": "https://localhost:8080/"} id="cJ1IlNvB_EZt" outputId="890945fb-a99e-4261-d8ae-226726d06322"
test_scores = np.array([[83,71,57,63],[54,68,81,45]])
print(test_scores)
print(test_scores.transpose())
# + colab={"base_uri": "https://localhost:8080/"} id="YMsxeup8_FHe" outputId="d6ab20f9-8134-471a-d01f-d6b3f01682f1"
10^-1
| Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# # Phoneme Data Analysis
#
# This notebook will analyze and preprocess the phoneme data. The dataset was used in the European ESPRIT 5516 project (ROARS), in which the objective is to develove and inmplementing a real time analytical system of speech recognition for the French and Spanish languages.
#
# As said by the words of [Phoneme Data Description](https://raw.githubusercontent.com/jbrownlee/Datasets/master/phoneme.names):
#
# The aim of the present database is to distinguish between nasal and
# oral vowels. There are thus two different classes:
#
# Class 0 : Nasals
# Class 1 : Orals
#
# This database contains vowels coming from 1809 isolated syllables (for
# example: pa, ta, pan,...). Five different attributes were chosen to
# characterize each vowel: they are the amplitudes of the five first
# harmonics AHi, normalised by the total energy Ene (integrated on all
# the frequencies): AHi/Ene. Each harmonic is signed: positive when it
# corresponds to a local maximum of the spectrum and negative otherwise.
# ## Dependencies
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import seaborn as sns
# -
phoneme = pd.read_csv("phoneme.csv")
phoneme.head(10)
# The dataset has no feature names, neither the class column, so we will give their names based upon the [Phoneme Data Description](https://raw.githubusercontent.com/jbrownlee/Datasets/master/phoneme.names)
# Get the columns names as array
new_header = phoneme.columns.values
new_header
# +
# Create a DataFrame from the hedader
ph = pd.DataFrame(new_header.astype(float)[None,:],columns=list(new_header))
# Transform the type of classes as integer
ph['0'] = ph['0'].astype('int64')
# Append both the dataframes
ph = ph.append(phoneme)
# Rename the columns
cols = {new_header[i]:'H{}'.format(i+1) for i in range(new_header.size-1)}
cols['0']='target'
ph=ph.rename(columns=cols)
ph.head(10)
# -
# ## Dataset Information and Description
#
# Let's analyze the information and the data description.
ph.info()
# So we can see that there are no missing data, which is a good thing, and all the features are floats, so for while we do not need to change any data type as much as the **target** feature, which is already of integer type.
#
# Now, let's see the description.
ph.describe()
# The means and the stds shows that the data hasn't so much variation, which is not necessary to standarlize the data.
#
# Now I will analyze the counting of labeled data, to see if there is some tendency.
count = ph['target'].value_counts().rename_axis('Unique').to_frame('Counts')
count['Counts']
plt.bar(np.arange(1),count['Counts'][0],color='b',alpha=0.5, label='Nasal')
plt.bar(np.arange(1)+1,count['Counts'][1],color='r',alpha=0.5, label='Oral')
plt.xlabel('Classes',fontsize=20, labelpad=30)
plt.ylabel('Frequency',fontsize=20, labelpad=30)
plt.xticks(np.arange(2),['0','1'],fontsize=20)
plt.yticks(fontsize=20)
plt.legend()
plt.show()
print("Nasal Vowels: {}%".format(round(3818*100/count['Counts'].sum(),2)))
print("Oral Vowels: {}%".format(round(1586*100/count['Counts'].sum(),2)))
print("Ratio Nasal/Oral: {}%".format(round((3818/1586)*100,2)))
# So we can see that there are 70% of the data labeled as Nasal, and it's 240% higher than the oral vowels. It's clear that the Nasal label is biased.
# Now, let's save our dataset.
ph.to_csv("phoneme.data")
| tests/11 - NSC Data Tests/All folders/Data-tests/comparison/datas/Phoneme_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hcad_pred
# language: python
# name: hcad_pred
# ---
# + [markdown] papermill={"duration": 0.044661, "end_time": "2020-09-13T12:44:15.367203", "exception": false, "start_time": "2020-09-13T12:44:15.322542", "status": "completed"} tags=[]
# # Find the comparables: real_acc.txt
#
# The file `real_acc.txt` contains important property information like number total appraised value (the target on this exercise), neighborhood, school district, economic group, land value, and more. Let's load this file and grab a subset with the important columns to continue our study.
# + papermill={"duration": 0.049398, "end_time": "2020-09-13T12:44:15.447405", "exception": false, "start_time": "2020-09-13T12:44:15.398007", "status": "completed"} tags=[]
# %load_ext autoreload
# %autoreload 2
# + papermill={"duration": 0.478058, "end_time": "2020-09-13T12:44:15.973517", "exception": false, "start_time": "2020-09-13T12:44:15.495459", "status": "completed"} tags=[]
from pathlib import Path
import pickle
import numpy as np
import pandas as pd
from src.definitions import ROOT_DIR
from src.data.utils import Table, save_pickle
# + papermill={"duration": 0.053467, "end_time": "2020-09-13T12:44:16.058458", "exception": false, "start_time": "2020-09-13T12:44:16.004991", "status": "completed"} tags=[]
real_acct_fn = ROOT_DIR / 'data/external/2016/Real_acct_owner/real_acct.txt'
assert real_acct_fn.exists()
# + papermill={"duration": 0.047817, "end_time": "2020-09-13T12:44:16.165004", "exception": false, "start_time": "2020-09-13T12:44:16.117187", "status": "completed"} tags=[]
real_acct = Table(real_acct_fn, '2016')
# + papermill={"duration": 0.05578, "end_time": "2020-09-13T12:44:16.250909", "exception": false, "start_time": "2020-09-13T12:44:16.195129", "status": "completed"} tags=[]
real_acct.get_header()
# + [markdown] papermill={"duration": 0.030536, "end_time": "2020-09-13T12:44:16.336836", "exception": false, "start_time": "2020-09-13T12:44:16.306300", "status": "completed"} tags=[]
# # Load accounts and columns of interest
# Let's remove the account numbers that don't meet free-standing single-family home criteria that we found while processing the `building_res.txt` file.
#
# Also, the columns above show a lot of value information along property groups that might come in handy when predicting the appraised value. Now let's get a slice of some of the important columns.
# + papermill={"duration": 9.845532, "end_time": "2020-09-13T12:44:26.213431", "exception": false, "start_time": "2020-09-13T12:44:16.367899", "status": "completed"} tags=[]
skiprows = real_acct.get_skiprows()
# + papermill={"duration": 0.046552, "end_time": "2020-09-13T12:44:26.310241", "exception": false, "start_time": "2020-09-13T12:44:26.263689", "status": "completed"} tags=[]
cols = [
'acct',
'site_addr_3', # Zip
'school_dist',
'Neighborhood_Code',
'Market_Area_1_Dscr',
'Market_Area_2_Dscr',
'center_code',
'bld_ar',
'land_ar',
'acreage',
'land_val',
'tot_appr_val', # Target
'prior_land_val',
'prior_tot_appr_val',
'new_own_dt', # New owner date
]
# + papermill={"duration": 8.120532, "end_time": "2020-09-13T12:44:34.490774", "exception": false, "start_time": "2020-09-13T12:44:26.370242", "status": "completed"} tags=[]
real_acct_df = real_acct.get_df(skiprows=skiprows, usecols=cols)
# + papermill={"duration": 0.068312, "end_time": "2020-09-13T12:44:34.604541", "exception": false, "start_time": "2020-09-13T12:44:34.536229", "status": "completed"} tags=[]
real_acct_df.head()
# + [markdown] papermill={"duration": 0.032287, "end_time": "2020-09-13T12:44:34.694488", "exception": false, "start_time": "2020-09-13T12:44:34.662201", "status": "completed"} tags=[]
# Double check if the there is only one account number per row
# + papermill={"duration": 0.161511, "end_time": "2020-09-13T12:44:34.890245", "exception": false, "start_time": "2020-09-13T12:44:34.728734", "status": "completed"} tags=[]
assert real_acct_df['acct'].is_unique
# + [markdown] papermill={"duration": 0.045413, "end_time": "2020-09-13T12:44:34.970733", "exception": false, "start_time": "2020-09-13T12:44:34.925320", "status": "completed"} tags=[]
# # Describe and clean the columns
#
# Now we must describe each column by answering:
#
# * Meaning
# * Descriptive statistics or value counts
# * Data type
#
# There is no explicit document provided by HCAD explaining all the variables, but most are easy to guess for using their name.
# + [markdown] papermill={"duration": 0.031929, "end_time": "2020-09-13T12:44:35.036116", "exception": false, "start_time": "2020-09-13T12:44:35.004187", "status": "completed"} tags=[]
# ## Fix column names
# We would like the column names to be all lower case, with no spaces nor non-alphanumeric characters.
# + papermill={"duration": 0.054339, "end_time": "2020-09-13T12:44:35.122711", "exception": false, "start_time": "2020-09-13T12:44:35.068372", "status": "completed"} tags=[]
from src.data.utils import fix_column_names
# + papermill={"duration": 0.056537, "end_time": "2020-09-13T12:44:35.210342", "exception": false, "start_time": "2020-09-13T12:44:35.153805", "status": "completed"} tags=[]
real_acct_df.columns
# + papermill={"duration": 0.050233, "end_time": "2020-09-13T12:44:35.292064", "exception": false, "start_time": "2020-09-13T12:44:35.241831", "status": "completed"} tags=[]
real_acct_df = fix_column_names(real_acct_df)
# + papermill={"duration": 0.056703, "end_time": "2020-09-13T12:44:35.382049", "exception": false, "start_time": "2020-09-13T12:44:35.325346", "status": "completed"} tags=[]
real_acct_df.columns
# + [markdown] papermill={"duration": 0.031789, "end_time": "2020-09-13T12:44:35.445168", "exception": false, "start_time": "2020-09-13T12:44:35.413379", "status": "completed"} tags=[]
# ### Find duplicated rows
# + papermill={"duration": 1.618926, "end_time": "2020-09-13T12:44:37.096863", "exception": false, "start_time": "2020-09-13T12:44:35.477937", "status": "completed"} tags=[]
cond0 = real_acct_df.duplicated()
real_acct_df.loc[cond0, :]
# + [markdown] papermill={"duration": 0.034682, "end_time": "2020-09-13T12:44:37.167685", "exception": false, "start_time": "2020-09-13T12:44:37.133003", "status": "completed"} tags=[]
# ## Zip code: site_addr_3
# These are the property zip code. They should be non-zero integers. Most should start with 77.
# + papermill={"duration": 0.055076, "end_time": "2020-09-13T12:44:37.256889", "exception": false, "start_time": "2020-09-13T12:44:37.201813", "status": "completed"} tags=[]
# Let's change the column name
real_acct_df.rename({'site_addr_3': 'zip_code'}, axis=1, inplace=True)
# + papermill={"duration": 0.293237, "end_time": "2020-09-13T12:44:37.582175", "exception": false, "start_time": "2020-09-13T12:44:37.288938", "status": "completed"} tags=[]
# there were some empty zip code: ' '
real_acct_df['zip_code'] = real_acct_df['zip_code'].apply(lambda code: code if code.startswith('77') else np.nan )
# + papermill={"duration": 0.540348, "end_time": "2020-09-13T12:44:38.160290", "exception": false, "start_time": "2020-09-13T12:44:37.619942", "status": "completed"} tags=[]
real_acct_df['zip_code'] = pd.to_numeric(real_acct_df['zip_code'], downcast='unsigned')
# + papermill={"duration": 0.059985, "end_time": "2020-09-13T12:44:38.253380", "exception": false, "start_time": "2020-09-13T12:44:38.193395", "status": "completed"} tags=[]
real_acct_df['zip_code'].value_counts()
# + papermill={"duration": 0.088312, "end_time": "2020-09-13T12:44:38.374867", "exception": false, "start_time": "2020-09-13T12:44:38.286555", "status": "completed"} tags=[]
real_acct_df['zip_code'].describe()
# + papermill={"duration": 0.13045, "end_time": "2020-09-13T12:44:38.540407", "exception": false, "start_time": "2020-09-13T12:44:38.409957", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['zip_code'].isnull())}")
# + [markdown] papermill={"duration": 0.033488, "end_time": "2020-09-13T12:44:38.606904", "exception": false, "start_time": "2020-09-13T12:44:38.573416", "status": "completed"} tags=[]
# ## School district: school_dist
# The school district values come coded as two digit integers. There are a few numbers missing from the code sequence, i.e. 10, 11, 13, 14, and 22. This is consistent with the ISD codes in the `code_jur_list` table. Let's decode the school district values using this table, but first let's represent the missing data with NaNs.
# + papermill={"duration": 0.055, "end_time": "2020-09-13T12:44:38.696408", "exception": false, "start_time": "2020-09-13T12:44:38.641408", "status": "completed"} tags=[]
from src.data.utils import decode_isd
# + papermill={"duration": 0.059664, "end_time": "2020-09-13T12:44:38.789439", "exception": false, "start_time": "2020-09-13T12:44:38.729775", "status": "completed"} tags=[]
real_acct_df['school_dist'].head()
# + papermill={"duration": 1.690019, "end_time": "2020-09-13T12:44:40.543613", "exception": false, "start_time": "2020-09-13T12:44:38.853594", "status": "completed"} tags=[]
real_acct_df['school_dist'] = decode_isd(real_acct_df['school_dist'])
# + papermill={"duration": 0.148512, "end_time": "2020-09-13T12:44:40.741003", "exception": false, "start_time": "2020-09-13T12:44:40.592491", "status": "completed"} tags=[]
real_acct_df['school_dist'].value_counts(normalize=True)
# + papermill={"duration": 0.206561, "end_time": "2020-09-13T12:44:40.982137", "exception": false, "start_time": "2020-09-13T12:44:40.775576", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['school_dist'].isnull())}")
# + [markdown] papermill={"duration": 0.03551, "end_time": "2020-09-13T12:44:41.052232", "exception": false, "start_time": "2020-09-13T12:44:41.016722", "status": "completed"} tags=[]
# ## neighborhood_code
#
# From [Definition help](https://pdata.hcad.org/Desc/Definition_help.pdf) file:
# > Residential valuation neighborhoods are groups of comparable properties whose
# boundaries were developed based on location and similarity of property data
# characteristics. Each neighborhood in a school district has a unique identifier known as a
# residential valuation number. These neighborhood boundaries are maintained via an ongoing office and field review. As neighborhoods change, neighborhood lines are redrawn
# to reflect the changes, and maintain the homogeneity of the neighborhood.
# + papermill={"duration": 0.050828, "end_time": "2020-09-13T12:44:41.139070", "exception": false, "start_time": "2020-09-13T12:44:41.088242", "status": "completed"} tags=[]
from src.data.utils import decode_nhood
# + papermill={"duration": 65.413156, "end_time": "2020-09-13T12:45:46.587627", "exception": false, "start_time": "2020-09-13T12:44:41.174471", "status": "completed"} tags=[]
real_acct_df['neighborhood_code'] = decode_nhood(real_acct_df['neighborhood_code'])
# + papermill={"duration": 0.172113, "end_time": "2020-09-13T12:45:46.799483", "exception": false, "start_time": "2020-09-13T12:45:46.627370", "status": "completed"} tags=[]
real_acct_df['neighborhood_code'].value_counts(normalize=True)
# + papermill={"duration": 0.208026, "end_time": "2020-09-13T12:45:47.043300", "exception": false, "start_time": "2020-09-13T12:45:46.835274", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['neighborhood_code'].isnull())}")
# + [markdown] papermill={"duration": 0.046268, "end_time": "2020-09-13T12:45:47.142657", "exception": false, "start_time": "2020-09-13T12:45:47.096389", "status": "completed"} tags=[]
# ## Market area description: market_area_1_dscr
# + papermill={"duration": 0.126174, "end_time": "2020-09-13T12:45:47.305823", "exception": false, "start_time": "2020-09-13T12:45:47.179649", "status": "completed"} tags=[]
real_acct_df['market_area_2_dscr'].value_counts(normalize=True)
# + papermill={"duration": 0.691673, "end_time": "2020-09-13T12:45:48.033733", "exception": false, "start_time": "2020-09-13T12:45:47.342060", "status": "completed"} tags=[]
cond0 = real_acct_df['market_area_2_dscr'].str.strip() == ''
real_acct_df.loc[cond0, :]
# + papermill={"duration": 0.167605, "end_time": "2020-09-13T12:45:48.241933", "exception": false, "start_time": "2020-09-13T12:45:48.074328", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['market_area_2_dscr'].isnull())}")
# + [markdown] papermill={"duration": 0.038843, "end_time": "2020-09-13T12:45:48.317854", "exception": false, "start_time": "2020-09-13T12:45:48.279011", "status": "completed"} tags=[]
# ## center_code
# From [Definition help](https://pdata.hcad.org/Desc/Definition_help.pdf) file:
# > The center code indicates the individual (HCAD employee or contractor) that is
# responsible for working and valuing the property account.
# + papermill={"duration": 0.150791, "end_time": "2020-09-13T12:45:48.507195", "exception": false, "start_time": "2020-09-13T12:45:48.356404", "status": "completed"} tags=[]
real_acct_df['center_code'].value_counts()
# + papermill={"duration": 0.240193, "end_time": "2020-09-13T12:45:48.805699", "exception": false, "start_time": "2020-09-13T12:45:48.565506", "status": "completed"} tags=[]
real_acct_df['center_code'] = real_acct_df['center_code'].apply(lambda x: np.nan if x.isspace() else x)
# + papermill={"duration": 0.174209, "end_time": "2020-09-13T12:45:49.017737", "exception": false, "start_time": "2020-09-13T12:45:48.843528", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['center_code'].isnull())}")
# + [markdown] papermill={"duration": 0.03893, "end_time": "2020-09-13T12:45:49.094634", "exception": false, "start_time": "2020-09-13T12:45:49.055704", "status": "completed"} tags=[]
# ## Building area: 'bld_ar'
# This seems to be the total area occupied by buildings in the parcel. It possibly includes non-livable buildings like barns, sheds, and other outbuildings.
# + papermill={"duration": 0.059701, "end_time": "2020-09-13T12:45:49.194765", "exception": false, "start_time": "2020-09-13T12:45:49.135064", "status": "completed"} tags=[]
from src.data.utils import fix_area_column
# + papermill={"duration": 0.572448, "end_time": "2020-09-13T12:45:49.805217", "exception": false, "start_time": "2020-09-13T12:45:49.232769", "status": "completed"} tags=[]
real_acct_df = fix_area_column(real_acct_df, 'bld_ar')
# + [markdown] papermill={"duration": 0.040751, "end_time": "2020-09-13T12:45:49.890849", "exception": false, "start_time": "2020-09-13T12:45:49.850098", "status": "completed"} tags=[]
# ## Land area: land_ar
# Land area in square feet.
# + papermill={"duration": 0.581363, "end_time": "2020-09-13T12:45:50.512369", "exception": false, "start_time": "2020-09-13T12:45:49.931006", "status": "completed"} tags=[]
real_acct_df = fix_area_column(real_acct_df, 'land_ar')
# + [markdown] papermill={"duration": 0.053683, "end_time": "2020-09-13T12:45:50.611618", "exception": false, "start_time": "2020-09-13T12:45:50.557935", "status": "completed"} tags=[]
# ## Land appraised value: land_val
# Appraised value of the land. It is represented in US dollars, should integers unless there are NaNs.
# + papermill={"duration": 0.075326, "end_time": "2020-09-13T12:45:50.757982", "exception": false, "start_time": "2020-09-13T12:45:50.682656", "status": "completed"} tags=[]
real_acct_df['land_val'].head()
# + papermill={"duration": 0.171761, "end_time": "2020-09-13T12:45:50.970415", "exception": false, "start_time": "2020-09-13T12:45:50.798654", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['land_val'].isnull())}")
# + papermill={"duration": 0.110843, "end_time": "2020-09-13T12:45:51.121394", "exception": false, "start_time": "2020-09-13T12:45:51.010551", "status": "completed"} tags=[]
real_acct_df['land_val'].describe().apply(lambda x: format(x, 'f'))
# + [markdown] papermill={"duration": 0.042066, "end_time": "2020-09-13T12:45:51.213840", "exception": false, "start_time": "2020-09-13T12:45:51.171774", "status": "completed"} tags=[]
# ## Total appraised value: tot_appr_val
# This is the target to be predicted. It is represented in US dollars, should integers unless there are NaNs.
# + papermill={"duration": 0.063069, "end_time": "2020-09-13T12:45:51.318916", "exception": false, "start_time": "2020-09-13T12:45:51.255847", "status": "completed"} tags=[]
from src.data.utils import fix_appraised_values
# + papermill={"duration": 0.214078, "end_time": "2020-09-13T12:45:51.572873", "exception": false, "start_time": "2020-09-13T12:45:51.358795", "status": "completed"} tags=[]
real_acct_df = fix_appraised_values(real_acct_df, 'tot_appr_val')
# + [markdown] papermill={"duration": 0.043013, "end_time": "2020-09-13T12:45:51.658697", "exception": false, "start_time": "2020-09-13T12:45:51.615684", "status": "completed"} tags=[]
# ## Prior land value: prior_land_val
# Last year's appraised land value. It is represented in US dollars, should integers unless there are NaNs.
# + papermill={"duration": 0.177925, "end_time": "2020-09-13T12:45:51.879632", "exception": false, "start_time": "2020-09-13T12:45:51.701707", "status": "completed"} tags=[]
real_acct_df = fix_appraised_values(real_acct_df, 'prior_land_val')
# + [markdown] papermill={"duration": 0.043062, "end_time": "2020-09-13T12:45:51.999154", "exception": false, "start_time": "2020-09-13T12:45:51.956092", "status": "completed"} tags=[]
# ## Prior total appraised value: prior_tot_appr_val
# Last year's total appraised land value. It is represented in US dollars, should integers unless there are NaNs.
# + papermill={"duration": 0.18162, "end_time": "2020-09-13T12:45:52.222373", "exception": false, "start_time": "2020-09-13T12:45:52.040753", "status": "completed"} tags=[]
real_acct_df = fix_appraised_values(real_acct_df, 'prior_tot_appr_val')
# + [markdown] papermill={"duration": 0.042673, "end_time": "2020-09-13T12:45:52.308034", "exception": false, "start_time": "2020-09-13T12:45:52.265361", "status": "completed"} tags=[]
# ## New owner date: new_own_dt
# When was the last recorded ownership changed on the property. Should be datetime type.
# + papermill={"duration": 0.261477, "end_time": "2020-09-13T12:45:52.611532", "exception": false, "start_time": "2020-09-13T12:45:52.350055", "status": "completed"} tags=[]
real_acct_df['new_own_dt'] = pd.to_datetime(real_acct_df['new_own_dt'])
# + papermill={"duration": 0.137212, "end_time": "2020-09-13T12:45:52.791843", "exception": false, "start_time": "2020-09-13T12:45:52.654631", "status": "completed"} tags=[]
print(f"The number of missing values is: {sum(real_acct_df['new_own_dt'].isnull())}")
# + papermill={"duration": 0.063468, "end_time": "2020-09-13T12:45:52.897043", "exception": false, "start_time": "2020-09-13T12:45:52.833575", "status": "completed"} tags=[]
real_acct_df['new_own_dt']
# + papermill={"duration": 0.098105, "end_time": "2020-09-13T12:45:53.044625", "exception": false, "start_time": "2020-09-13T12:45:52.946520", "status": "completed"} tags=[]
real_acct_df['new_own_dt'].describe()
# + [markdown] papermill={"duration": 0.043733, "end_time": "2020-09-13T12:45:53.132974", "exception": false, "start_time": "2020-09-13T12:45:53.089241", "status": "completed"} tags=[]
# # Export real_acct
# + papermill={"duration": 0.433891, "end_time": "2020-09-13T12:45:53.610211", "exception": false, "start_time": "2020-09-13T12:45:53.176320", "status": "completed"} tags=[]
save_fn = ROOT_DIR / 'data/raw/2016/real_acct_comps.pickle'
save_pickle(real_acct_df, save_fn)
| notebooks/01_Exploratory/output/1.2-rp-hcad-data-view-real-acct_20200913.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from cloudmesh.compute.vm.Provider import Provider
provider = Provider(name="chameleon")
flavors = provider.flavors()
flavors[0]['name']
provider.Print(flavors)
| notebooks/cloudmesh-flavor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading the Open COVID-19 Dataset
# This very short notebook showcases how to load the [Open COVID-19 datset](https://github.com/open-covid-19/data), including some examples for commonly performed operations.
#
# First, loading the data is very simple with `pandas`. We can use the CSV master table to download the entire Open COVID-19 dataset in a single step:
# +
import pandas as pd
# Load CSV data directly from the URL with pandas
data = pd.read_csv("https://open-covid-19.github.io/data/v2/master.csv")
# Print a small snippet of the dataset
print(f"The dataset currently contains {len(data)} records, here is a sample:"
data.sample(5)
# -
# ### Looking at country-level data
# Some records contain country-level data, in other words, data that is aggregated at the country level. Other records contain region-level data, which are subdivisions of a country; for example, Chinese provinces or USA states. A few regions also report at an even smaller subdivision, i.e. county/municipality level.
#
# To filter only country-level data from the dataset, look for records that have a `aggregation_level == 0` or, alternatively, null value for the `subregion1_code` (or `subregion1_name`) field:
# +
# Look for rows with country level data
# Same as `data[data.subregion2_code.isna()]`
countries = data[data.aggregation_level == 0]
# We no longer need the subregion-level columns
countries = countries.drop(columns=['subregion1_code', 'subregion1_name', 'subregion2_code', 'subregion2_name'])
countries.tail()
# -
# ### Looking at state/province data
# Conversely, to filter state/province data for a specific country, we need to look for records where the aggregation level is `1` (or where the region columns have non-null values). The following snippet extracts data related to Spain's subregions from the dataset:
# +
# Filter records that have the right country code AND a non-null region code
# Same as `data[(data.country_code == 'ES') & ~(data.subregion`_code.isna())]`
spain_regions = data[(data.country_code == 'ES') & (data.aggregation_level == 1)]
# We no longer need the municipality-level columns
spain_regions = spain_regions.drop(columns=['subregion2_code', 'subregion2_name'])
spain_regions.tail()
# -
# ### Using the `key` column
# The `key` column is present in all datasets and is unique for each combination of country, province/state and municipality/county. This way, we can retrieve a specific country or region using a single filter for the data. The `key` column is built using `country_code` for country-level data, `${country_code}_${subregion1_code}` for province/state level data, and `${country_code}_${subregion1_code}_${subregion2_code}` for municipality/county data:
# +
# Filter records for Spain at the country-level
spain_country = data[data.key == 'ES']
# We no longer need the subregion-level columns
spain_country = spain_country.drop(columns=['subregion1_code', 'subregion1_name', 'subregion2_code', 'subregion2_name'])
spain_country.tail()
# +
# Filter records for Madrid, one of the subregions of Spain
madrid = data[data.key == 'ES_MD']
madrid.tail()
# -
# ### Dataset Subsets
# The master table can be large and cumbersome depending on your application. If you only need a subset of the data, you can consult each table individually. For a list of all the available tables, see the [README](../README.md) of the repo. For example, here's how you would get only epidemiology data for Madrid:
# +
# Load the epidemiology table
# Note that all the helper columns such as country code, country name, aggregation level, etc. are present in the
# `index` table; we only have the key here
epi = pd.read_csv('https://open-covid-19.github.io/data/v2/epidemiology.csv')
# Filter records for Madrid, one of the subregions of Spain
madrid = epi[epi.key == 'ES_MD']
madrid.tail()
# -
# ### Data consistency
# Often, region-level data and country-level data will come from different sources. This will lead to numbers not adding up exactly, or even date misalignment (the data for the region may be reported sooner or later than the whole country). However, country- and region- level data will *always* be self-consistent
mob = pd.read_csv('https://open-covid-19.github.io/data/v2/mobility.csv')
mob = data[['date', 'key', 'country_code', 'subregion1_name', 'aggregation_level', 'mobility_driving']].dropna()
mob = mob[(mob.country_code == 'US') & (mob.aggregation_level == 1)].set_index(['key', 'date'])
import seaborn
seaborn.set()
mob.loc['US_OH', 'mobility_driving'].plot()
| examples/data_loading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chiaramarzi/ML-models-validation/blob/main/models_validation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="kTe8Jj3qhlw5"
# # Artificial intelligence (AI) for health - potentials
#
# + [markdown] id="BGoOZvGALKgw"
# * **Data mining**: finding pattern in big data
# * **Biomarker discovery**: determining potential (compound) biomarkers
# * The **predicitive nature** of machine learning strategies is highly in line with the aim of clinical diagnosis and prognosis **in the single patient**
# + [markdown] id="UUGpx9Z7iqL0"
# # Models validation
#
#
# + [markdown] id="4JwXyfF-LRej"
# In machine learning, model validation is referred to as the process where a trained model is evaluated with a testing data set. The testing data set is a separate portion of the same data set from which the training set is derived.
# Model validation is carried out after model training.
#
# Estimation of **unbiased generalization performance** of the model
# + [markdown] id="iZ_Jo9T8SBbc"
# # Outline
# + [markdown] id="z9T0es-dSFsQ"
# * Holdout validation
# * K-fold cross-validation (CV)
# * Leave-One-Out CV (LOOCV)
# * Hyperparameters tuning
# * Training, validation and test set: the holdout validation
# * Training, validation and test set: the nested CV
# * Sampling bias
# * Repetition of holdout validation
# * Repetition of CV
# * Unbalanced datasets
# + [markdown] id="kfjOSjyakq9H"
# # Age prediction based on neuroimaging features
# + [markdown] id="OTl_nz8OLv_C"
#
#
# * Data: T1-weighted images of 86 healthy subjects with age ranging from 19 to 85 years (41 males and 45 females, age 44.2 ± 17.1 years, mean ± standard deviation). Data are freely accessible at [here](https://fcon_1000.projects.nitrc.org/) and described in (Mazziotta et al., 2001)
# * Features:
# * Cortical thickness (mCT)
# * Gyrification index (Pial_mean_GI)
# * Fractal dimension (FD)
# * Task:
# * Regression:
# * Estimator: Support Vector Machines (SVR)
# * Performance: Mean Absolute Error (MAE)
# * Classification ("young" vs. "old"):
# * Estimator: Support Vector Machines (SVC)
# * Performance: Accuracy
#
# The same data and features have been previously investigated in (Marzi et al., 2020).
#
#
# **References**
#
# <NAME>., <NAME>., <NAME>. et al. Toward a more reliable characterization of fractal properties of the cerebral cortex of healthy subjects during the lifespan. Sci Rep 10, 16957 (2020). https://doi.org/10.1038/s41598-020-73961-w
#
# <NAME>. et al. A probabilistic atlas and reference system for the human brain: International Consortium for Brain Mapping (ICBM). Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 1293–1322. https://doi.org/10.1098/rstb.2001.0915 (2001).
# + [markdown] id="Im3BWk_5OVtS"
# # Cloning repository, libraries and data loading
# + id="-MZY9faAOjaj"
# My repo cloning
# ! git clone https://github.com/chiaramarzi/ML-models-validation
# %cd /content/ML-models-validation
# ! git pull
# + id="4vmE6GnKUpQ7"
# Libraries loading
from IPython.display import Image
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split, KFold, GridSearchCV, cross_validate
from sklearn.svm import SVR, SVC
from sklearn.metrics import mean_absolute_error, accuracy_score
# %run utils.ipynb import *
# Regression data
reg_data = pd.read_csv('data_regression.csv')
# Balanced classification data
class_data = pd.read_csv('data_classification_balanced.csv')
# Unbalanced classification data
unbal_class_data = pd.read_csv('data_classification_unbalanced.csv')
# + id="tSE0z7esU8Ky"
reg_data
# + id="ndM8efnpVGAP"
class_data
# + id="2YHx6EyaVHwK"
unbal_class_data
# + [markdown] id="E6G4jorBVPOI"
# # Holdout validation
# + [markdown] id="wZRcU-8ZtQ0h"
# The principle is simple, you simply split your data randomly into roughly 70% used for training the model and 30% for testing the model.
#
# 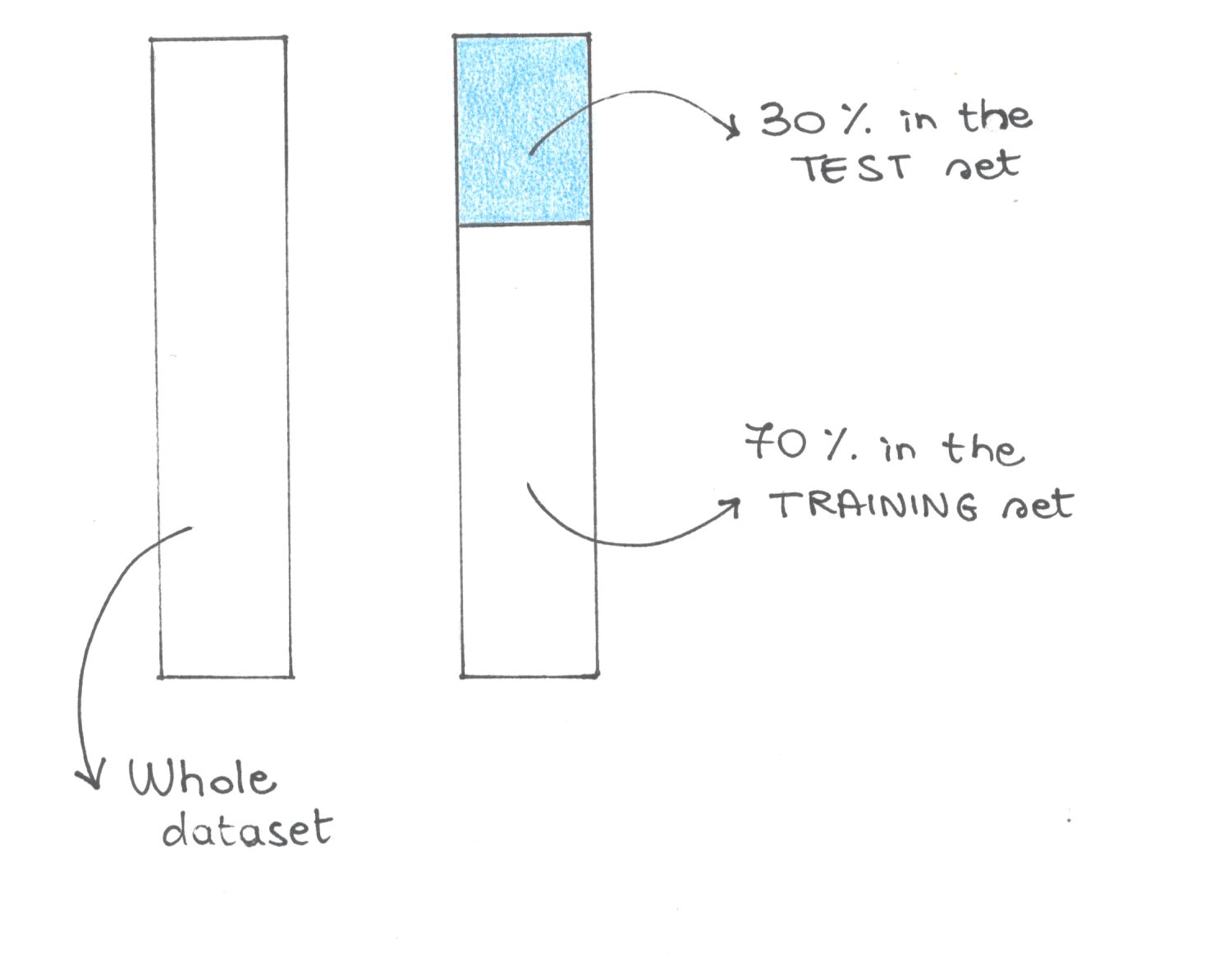
#
#
#
#
#
#
#
#
#
# + id="4z4UNX79uxKr"
#Image('figures/IMG_4103.png')
# + id="5ftXYNS7hM81"
#SEED = 42 #563: good, 0: perfect, 42: worse
### REGRESSION ###
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
regression_holdout(X, y, seed = 42, test_size = 0.25)
# + id="8J2_qJfzk-xl"
### CLASSIFICATION ###
print('***Classification task')
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" (<= 30 years) subjects in the sample:", np.sum(y==0))
print("\"old\" (>= 56 years) subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
classification_holdout(X, y, seed = 42, stratify = None, test_size = 0.25)
# + [markdown] id="vBrm5c1DlgTV"
# # K-fold cross-validation (CV)
# + [markdown] id="5DJkmJ3alik5"
# It splits the data into k folds, then trains the data on k-1 folds and test on the one fold that was left out. It does this for all combinations and averages the result on each instance.
#
# 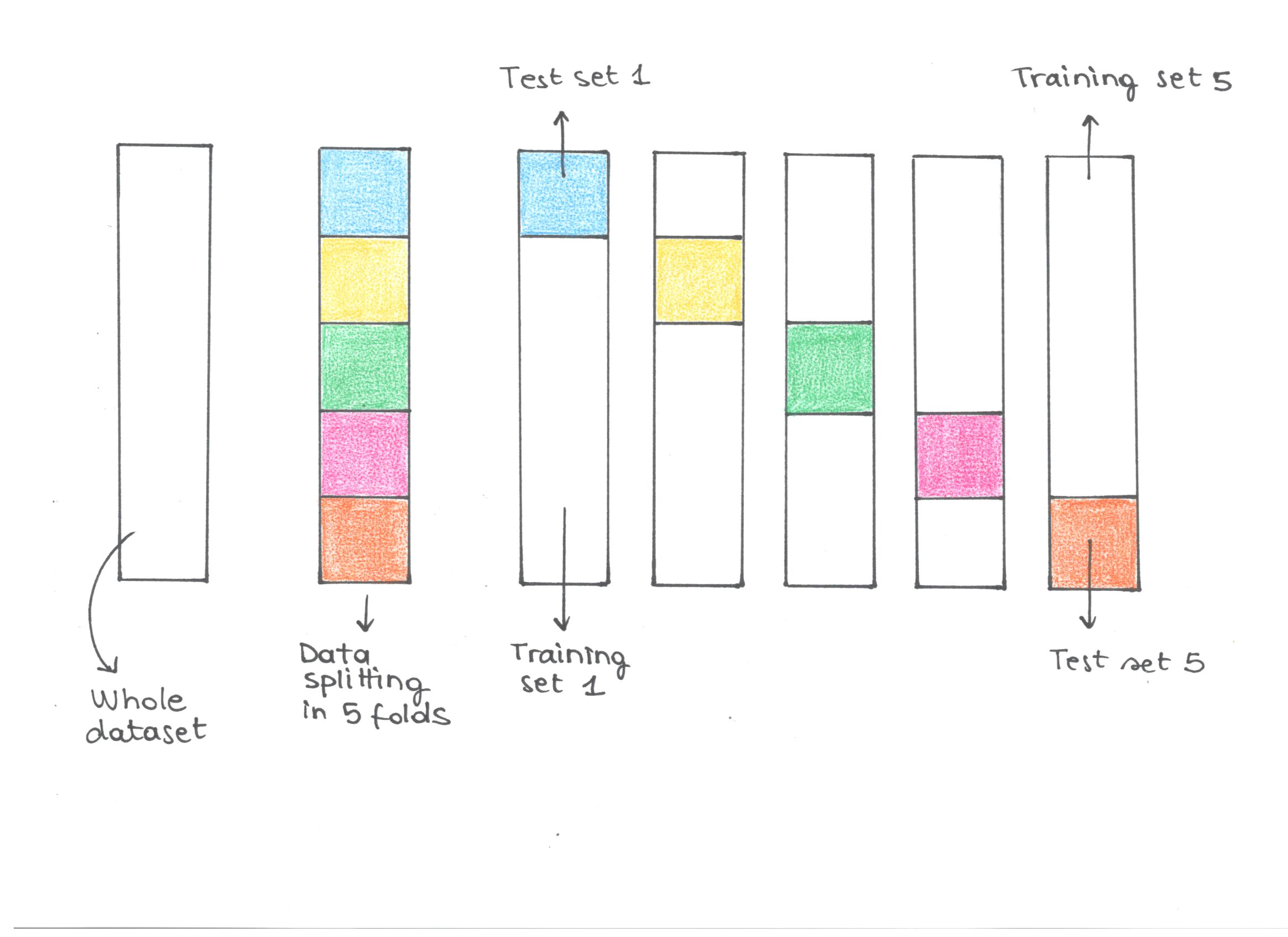
#
# The advantage is that all observations are used for both training and validation, and each observation is used once for validation.
#
# We typically choose either k=5 or k=10 as they find a nice balance between computational complexity and validation accuracy.
#
# The scores of each fold from CV techniques are more insightful than one may think. They are mostly used to simply extract the average performance. However, one might also look at the variance or standard deviation of the resulting folds as it will give information about the stability of the model across different data inputs.
#
# + id="HHpBE5XxmCpJ"
### REGRESSION ###
n_folds = 5 # for LOOCV insert n_fold = 86
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
MAE_train, MAE_test = regression_CV(X, y, seed = 42, n_folds = n_folds)
print_to_std(MAE_train, MAE_test, "MAE")
# + id="zQ2aP-OsmhDK"
### CLASSIFICATION ###
n_folds = 5 # for LOOCV insert n_fold = 50
print('***Classification task')
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
ACC_train, ACC_test = classification_CV(X, y, seed = 42, n_folds = n_folds)
print_to_std(ACC_train, ACC_test, "ACC")
# + [markdown] id="7XT91xJqm3in"
# # Leave-one-out CV (LOOCV)
# + [markdown] id="Sgu3uRNBm-Jf"
# A variant of k-fold CV is Leave-one-out Cross-Validation (LOOCV).
#
# LOOCV uses each sample in the data as a separate test set while all remaining samples form the training set. This variant is identical to k-fold CV when k = n (number of observations).
#
# LOOCV is computationally very costly as the model needs to be trained n times. Only do this if the dataset is small or if you can handle that many computations.
#
#
# + id="tuoxXbBGnPi0"
### REGRESSION ###
n_folds = 86
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
MAE_train, MAE_test = regression_CV(X, y, seed = 42, n_folds = n_folds)
print_to_std(MAE_train, MAE_test, "MAE")
# + id="pBzy85wWncCP"
### CLASSIFICATION ###
n_folds = 50
print('***Classification task')
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
ACC_train, ACC_test = classification_CV(X, y, seed = 42, n_folds = n_folds)
print_to_std(ACC_train, ACC_test, "ACC")
# + [markdown] id="d-0dPq40nlXl"
# # Hyperparameters tuning
# + [markdown] id="6xN-9qQFnvl_"
# Support vector machines (SVMs) are a set of supervised learning methods used for classification, regression and outliers detection.
#
# The advantages of support vector machines are:
#
# * Effective in high dimensional spaces.
# * Still effective in cases where number of dimensions is greater than the number of samples.
# * Uses a subset of training points in the decision function (called support vectors), so it is also memory efficient.
# * Versatile: different Kernel functions can be specified for the decision function. Common kernels are provided, but it is also possible to specify custom kernels.
#
# Both SVR() and SVC() classes have, among others, a **regularization parameter**. The parameter C, common to all SVM kernels, trades off misclassification of training examples against simplicity of the decision surface. The strength of the regularization is inversely proportional to C. Must be strictly positive. A low C makes the decision surface smooth, while a high C aims at classifying all training examples correctly.
# + [markdown] id="CtagDWcOn6ur"
# # Training, validation and test set: the holdout validation
# + [markdown] id="7Y1riVe2oLTV"
# When optimizing the hyperparameters of your model, you might overfityour model if you were to optimize using the train/test split.
# Why? Because the model searches for the hyperparameters that fit the specific train/test you made.
#
# To solve this issue, you can create an additional holdout set. This is often 10% of the data which you have not used in any of your processing/validation steps.
#
# 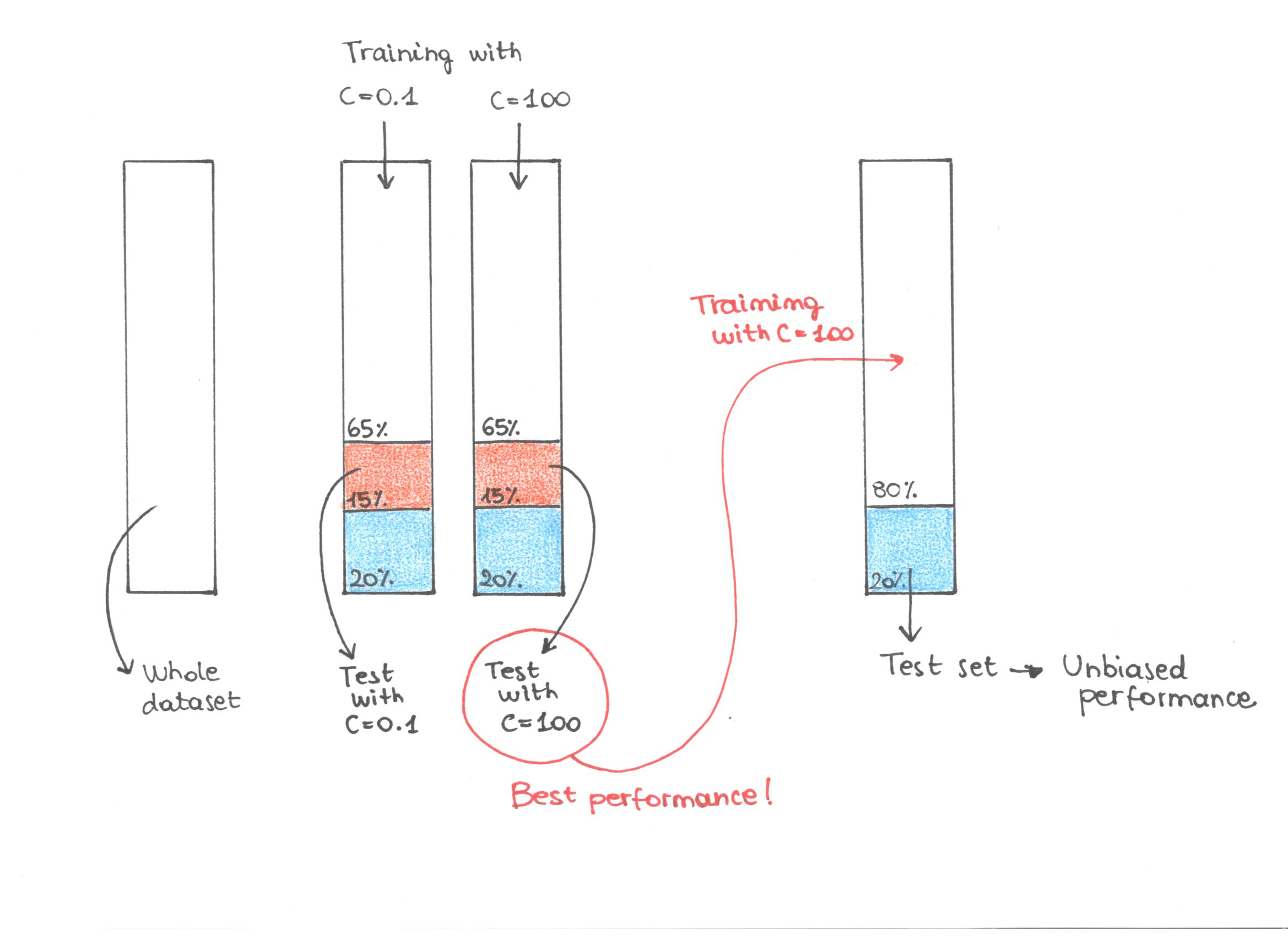
#
# + id="pLqXCvQhoVFP"
SEED = 42 #563: good, 0: perfect
C = [0.1, 1, 10, 100]
### REGRESSION ###
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
regression_holdout_val_set(X, y, SEED, test_set_size = 0.25, val_set_size = 0.15, C=C)
# + id="YHnjLjXLqI1-"
### CLASSIFICATION ###
print('***Classification task')
C = [0.1, 1, 10, 100]
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
classification_holdout_val_set(X, y, SEED, test_set_size = 0.25, val_set_size = 0.15, C=C)
# + [markdown] id="CzqJx2cMqalD"
# # Training, validation and test set: the nested CV
# + [markdown] id="eaADFev8rolQ"
# When you are optimizing the hyperparameters of your model and you use the same k-Fold CV strategy to tune the model and evaluate performance you run the risk of overfitting. You do not want to estimate the accuracy of your model on the same split that you found the best hyperparameters for.
#
#
# Instead, we use a Nested Cross-Validation strategy allowing to separate the hyperparameter tuning step from the error estimation step. To do this, we nest two k-fold cross-validation loops:
#
#
# * The inner loop for hyperparameter tuning and
# * the outer loop for estimating accuracy
#
# 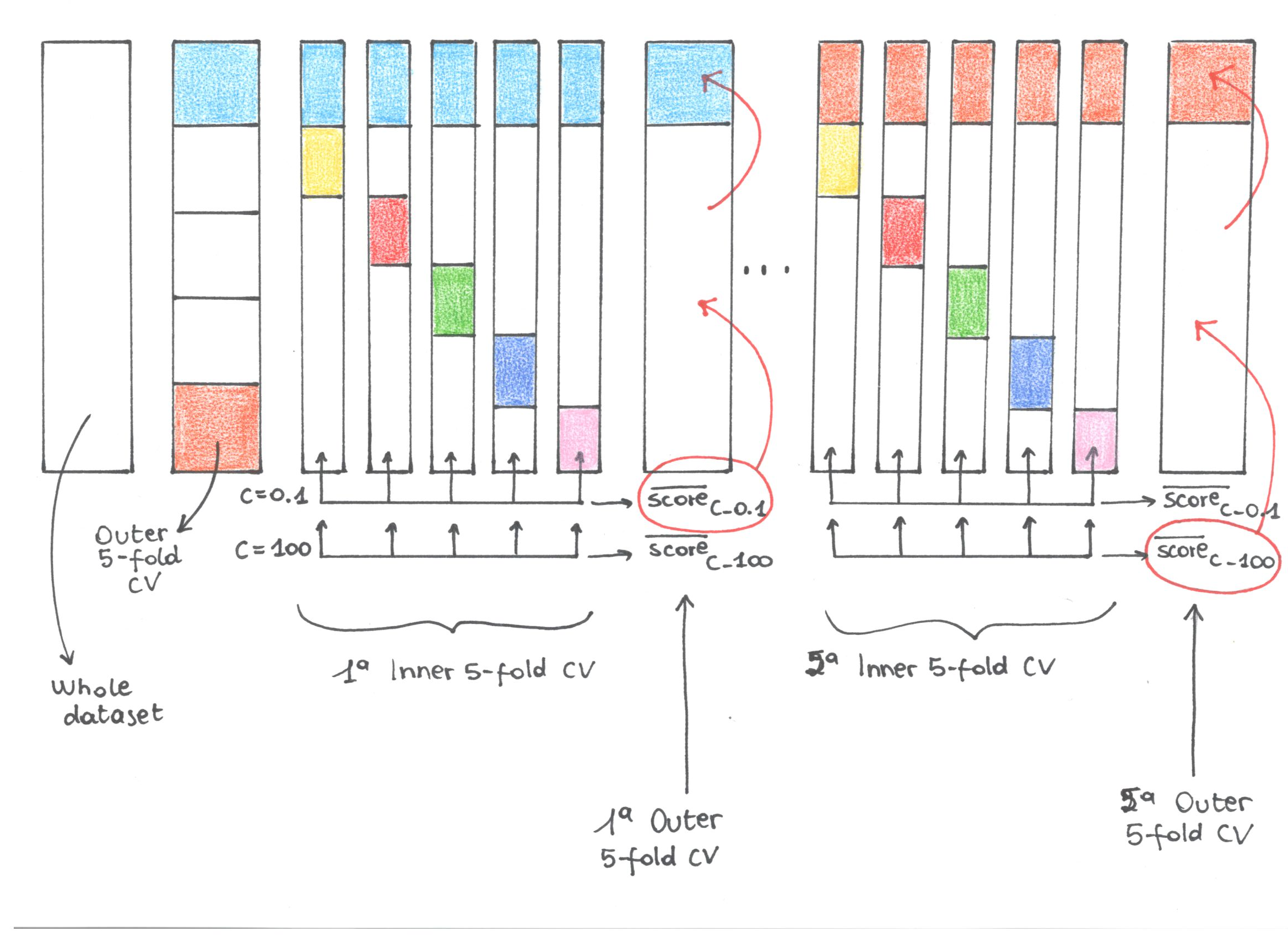
#
# + id="y-U58KpmsGlN"
### REGRESSION ###
print('***Regression task')
SEED = 42
outer_n_folds = 5
inner_n_folds = 5
C = [0.1, 1, 10]
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
MAE_tr_val, MAE_test = regression_nestedCV(X, y, SEED, outer_n_folds, inner_n_folds, C)
print_to_std(MAE_tr_val, MAE_test, "MAE")
# + id="zb2xDS8rHz5Y"
# NestedCV implemented in scikit-learn
outer_cv = KFold(n_splits=outer_n_folds, shuffle=True, random_state=SEED)
inner_cv = KFold(n_splits=inner_n_folds, shuffle=True, random_state=SEED)
clf = SVR(kernel='rbf', degree=3, gamma='scale', coef0=0.0, tol=0.001, C=0.1, epsilon=0.1, shrinking=True, cache_size=200, verbose=0, max_iter=- 1)
p_grid = [{'C': C}]
X = np.asarray(X)
y = np.asarray(y)
clf_gs = GridSearchCV(clf, param_grid=p_grid, cv=inner_cv, refit='neg_mean_absolute_error', scoring='neg_mean_absolute_error', n_jobs=1, verbose = 4)
nested_score = cross_validate(clf_gs, X=X, y=y, cv=outer_cv, return_train_score=True, return_estimator=True, scoring = 'neg_mean_absolute_error', n_jobs=1)
#print(np.abs(nested_score['train_score']))
#print(np.abs(nested_score['test_score']))
print("Average MAE train:", np.abs(np.mean(nested_score['train_score'])))
print("Average MAE test:", np.abs(np.mean(nested_score['test_score'])))
# + id="lA-GopPOsmo4"
### CLASSIFICATION ###
print('***Classification task')
SEED = 42
outer_n_folds = 5
inner_n_folds = 5
C = [0.1, 1, 10]
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
ACC_train, ACC_test = classification_nestedCV(X, y, SEED, outer_n_folds, inner_n_folds, C)
print_to_std(ACC_train, ACC_test, "ACC")
# + [markdown] id="iEm4D3AVs1hX"
# # Sampling bias
# + [markdown] id="ugdUBZ_Ys-A4"
# What if one subset of our data only have people of a certain age or income levels? This is typically referred to as a sampling bias.
#
# **Sampling bias** is systematic error due to a non-random sample of a population, causing some members of the population to be less likely to be included than others, resulting in a biased sample.
#
# 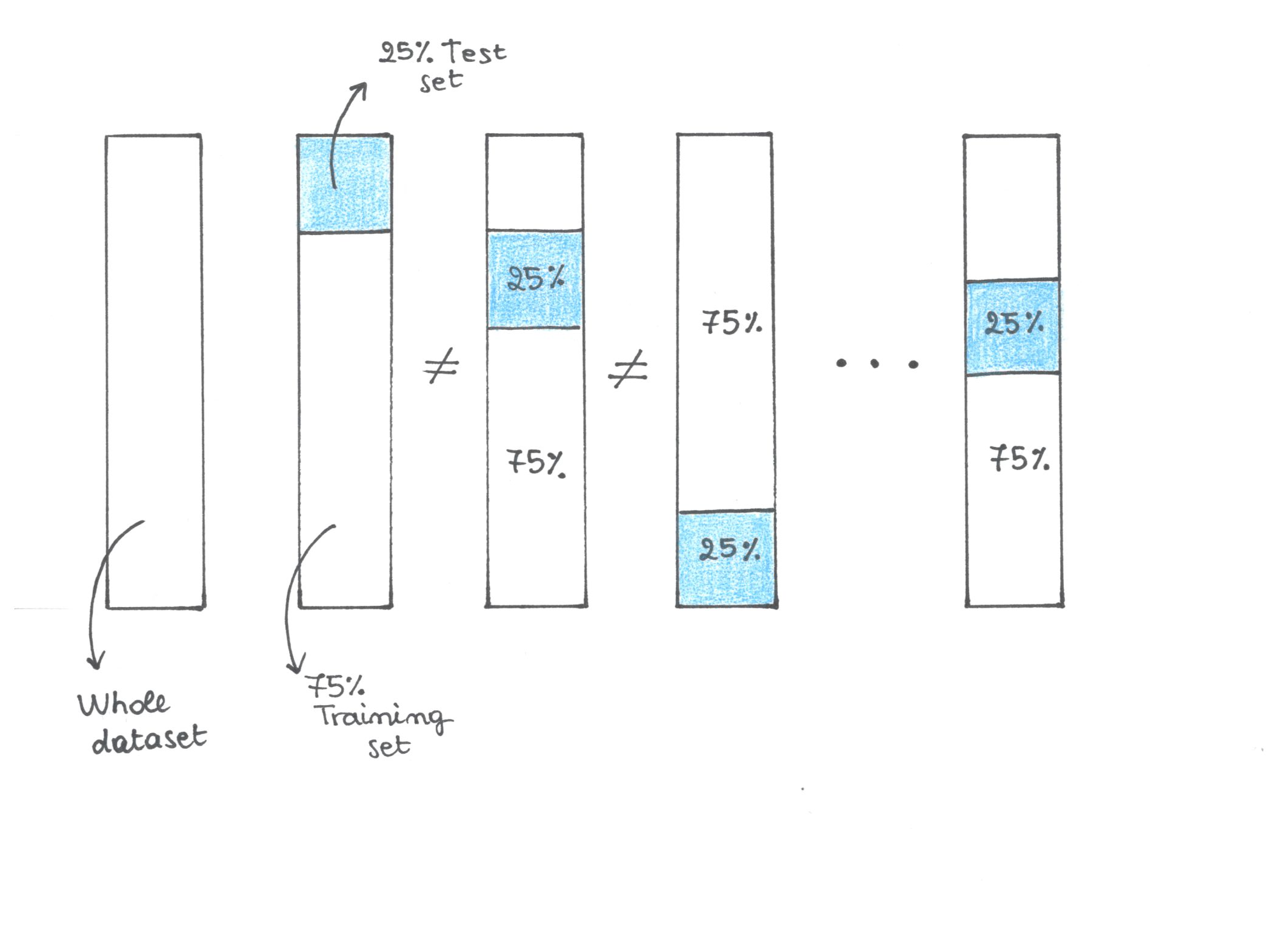
# + [markdown] id="1T1cTdbQtITQ"
# # Repetition of holdout validation
# + [markdown] id="sh2_22VkuYRq"
# One way to overcome the sampling bias is the **n-times repetition** of the validation method, changing the seed of the pseudo-random numbers generator, that determines the data splitting.
#
# The performances on the different test sets will be averaged.
#
# 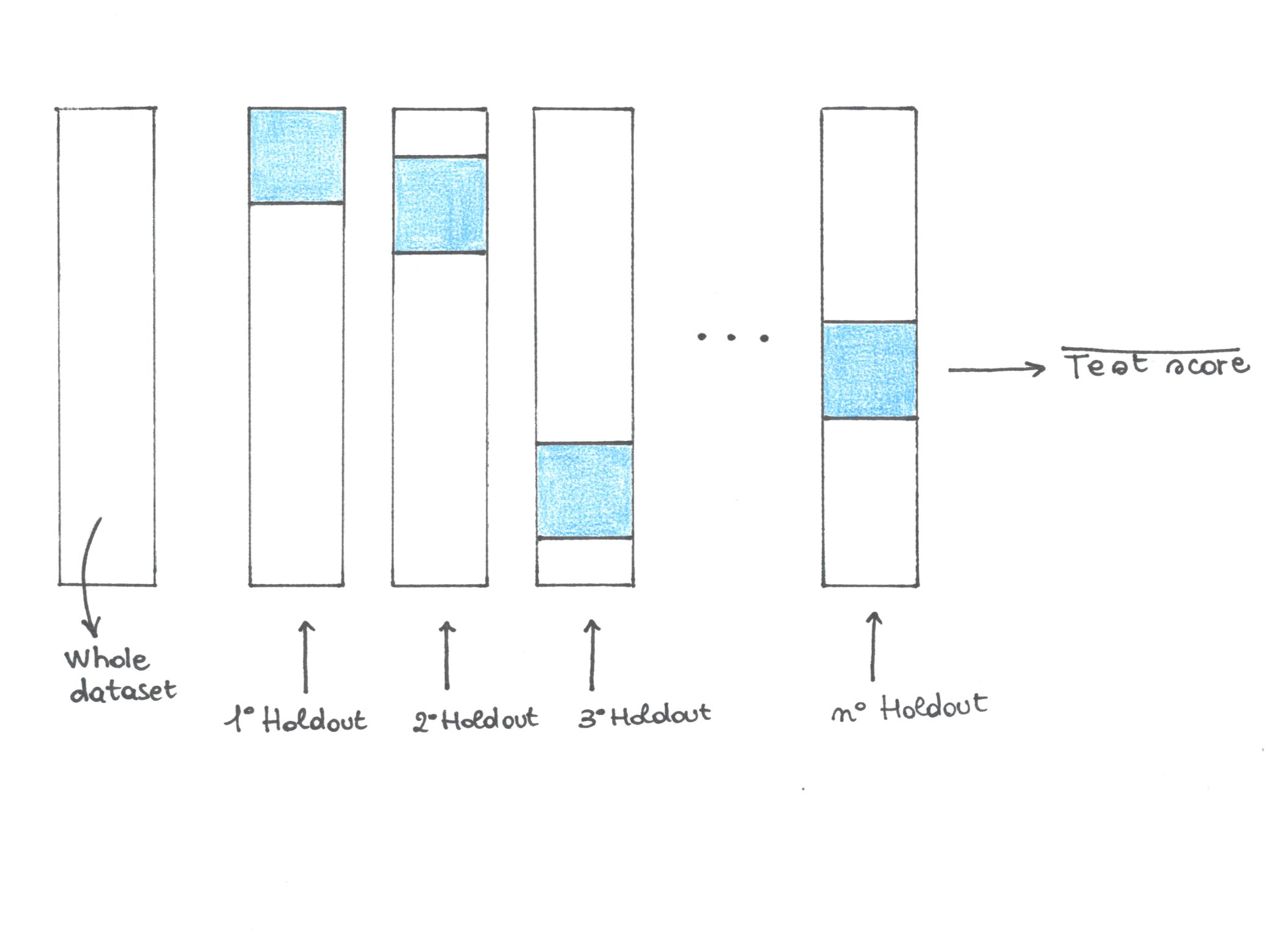
# + id="ATIUK0JItLdc"
print('#### SAMPLING BIAS: HOLDOUT REPETITION')
### REGRESSION ###
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
for SEED in range(0,10):
print("Seed:", SEED)
regression_holdout(X, y, seed = SEED, test_size = 0.25)
# + id="re_UEY8xte1a"
### CLASSIFICATION ###
print('***Classification task')
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
for SEED in range(0,10):
print("Seed:", SEED)
classification_holdout(X, y, SEED, test_size = 0.25, stratify = None)
# + [markdown] id="uDKhgOEptvRx"
# # Repetition of CV
# + id="cd9vC_WFtw_6"
print('#### SAMPLING BIAS: K-FOLD CV REPETITION')
### REGRESSION ###
n_folds = 5
print('***Regression task')
X = reg_data.iloc[:,2:5]
y = reg_data['Age']
print('The whole dataset contains ' + str(np.shape(reg_data)[0]) + ' subjects')
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
for SEED in range(1,10):
print("Seed:", SEED)
MAE_train, MAE_test = regression_CV(X, y, SEED, n_folds)
print_to_std(MAE_train, MAE_test, "MAE")
# + id="byN32Bd6uCZo"
### CLASSIFICATION ###
n_folds = 5
print('***Classification task')
X = class_data.iloc[:,2:5]
y = class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(class_data)[0]) + ' subjects')
print("\"young\" subjects in the sample:", np.sum(y==0))
print("\"old\" subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
for SEED in range(1,10):
print("Seed:", SEED)
ACC_train, ACC_test = classification_CV(X, y, SEED, n_folds)
print_to_std(ACC_train, ACC_test, "ACC")
# + [markdown] id="JQOYF10-uSR8"
# # Unbalanced datasets
# + [markdown] id="KlVF4wxeuX94"
# In some cases, there may be a large imbalance in the response variables.
#
# For example, in case of classification, there might be several times more negative samples than positive samples. For such problems, a slight variation in the K-Fold cross validation technique is made, such that each fold contains approximately the same percentage of samples of each target class as the complete set, or in case of prediction problems, the mean response value is approximately equal in all the folds.
#
# This variation is also known as **Stratified** K-Fold CV
# + id="gAITJP9Wuj2Z"
SEED = 95
#### UNBALANCED DATASETS ###
print('#### UNBALANCED DATASETS')
### CLASSIFICATION ###
print('***Classification task')
X = unbal_class_data.iloc[:,2:5]
y = unbal_class_data['Age_class']
print('The whole dataset contains ' + str(np.shape(unbal_class_data)[0]) + ' subjects')
print("\"young\" (<= 30 years) subjects in the sample:", np.sum(y==0))
print("\"old\" (> 30 years) subjects in the sample:",np.sum(y==1))
print()
print('The age prediction will be performed using ' + str(np.shape(X)[1]) + ' MRI-derived features')
print()
'''
for SEED in range(0,100):
print("SEED:", SEED)
classification_holdout(X, y, SEED, stratify = None)
classification_holdout(X, y, SEED, stratify = y)
'''
print("Unbalanced dataset - unstratified holdout")
classification_holdout(X, y, SEED, test_size = 0.25, stratify = None)
print("Unbalanced dataset - stratified holdout")
classification_holdout(X, y, SEED, test_size = 0.25, stratify = y)
# SEED = 95, 91, 88
| models_validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://github.com/jchan-gi/TORI/blob/master/docs/assets/Tori.png?raw=true" width="720px">
#
#
# # Tori (Trend- & oversell-based rebalancing instructions) in BCT MPF ProChoice MPF
# # 銀聯強積金積金之選應用TORI進行資產調配
#
#
# ## Disclaimer
# 1. The author (jchan-gi) expressly stated that nothing my repositories and webpages constitutes any advices or recommendation on investing or allocating assets on any stock or financial products.
# 2. No content in my study have been prepared with respect to specific personal need and investment objects of individuals. All materials reveal to the public are applicable to the author (jchan-gi) only.
# 3. All results from this repository are simulated from historical data only. It may portray inaccurate information and hence the repository must not applied in production and must used as academic exchange only. The author is not responsible in any loss induced from using this repository.
# 4. All investors shall obtain kind advices from professional financial consultants before making any decisions. Professional financial consultants should recommend products or decisions that suit your needs and objectives.
# 5. The author is not licensed nor professional in the field hence this studies are not professional advice and may not be suitable for anyone except myself.
# 6. You shall not invest on any financial products using the information on this code repository, website or any information made public by jchan-gi alone. You agree you have fully understand and willing to undertake the risks of this script in any circumstances involves in using this script.
# 7. English version shall be used if any discrepencies exists between English and Chinese version.
#
# ## 免責聲明
# 1. 本文作者現此聲明,本人的程式及網頁並非對閤下提供或作出對任何金融產品的投資或資產調配的建議。
# 2. 本文內容沒有針對閣下或其他人的實際個人需要而編撰。所有公開的內容只適用於本文作者。
# 3. 本文內容根據歷史數據模擬回報率等一切資訊。該等資訊可能非常不準確,本文及程式庫任何內容不應於實際環境使用,及只供學術討論。如因使用本程式庫任何內容而招致損失,作者一概不會負責。
# 4. 所有投資者應在作出任何決定前,先向專業理財顧問查詢及要求提供意見。只有理財顧問的意見才能符合閣下的實際理財需要及風險胃納。
# 5. 本文作者非專業人士或持牌從業者,因此,本文內容並非專業意見。而且,本文內容極有可能並不合適於除作者以外的任何人。
# 6. 在作出任何投資決定前,你不應單靠此程序作出決定。當你作出任何與本程序算法有關的投資決定時,你已完全知悉並願意接受本程序或算法所帶來的風險。
# 7. 本聲明如有歧義,將以英文版本作準。
#
#
# ## 最後更新日期 Last update: 2020/08/13.
# ### Changelog
# ### 更新記錄
#
# 2020-08-13 v3.0.2b
# Repair calcuation method of IRR.
# 修正內部回報率IRR的計算方式。
#
# 2020-08-11 v3.0.1bII
# Update project dependencies (Python part).
# 更新所需Python程式庫。
#
# 2020-08-11 v3.0.1bI
# Adjust the substitute candidates and their weights for BCEPCHK.
# 更改中港股票基金的替代基金及其比重。
#
# 2020-08-03 v3.0.1b
# Release results accounted up to 2020/07/30.
# 公佈直至2020/07/30的結果。
#
# 2020-07-29 v3.0.1a
# NAV storage media are now using SQLite3 database for capability in automated loading.
# 改用SQLite3數據庫儲存NAV以加強支援自動載入功能。
#
# 2020-07-25 v3.0.0a
# Release results accounted up to 2020/06/30.
# 公佈直至2020/06/30的結果。
#
# 2020-07-24 v3.0.0rc1
# Converted to Jupyter Notebook + R format for better performance.
# 轉換成Jupyter Notebook + R方式計算。
#
# 2020-05-03 v2.1.3b
# Release results accounted up to 2020/04/28.
# 公佈直至2020/04/28的結果。
#
# 2020-04-02 v2.1.3a
# Release results accounted up to 2020/03/31.
# 公佈直至2020/03/31的結果。
#
# 2020-03-27 v2.1.1a
# 1. Fix return matrix column misalignment in Tori @ BCT.
# 2. Fix faulty code on inconsistent total asset price across time. Description: Subsetting would not work to produce consistent total asset value when the data is splited by train-test partition. This is due to the hedge flag detect the future signal if the subsetting is being applied to the weight but not the return. This is a characteristics of Return.portfolio but not a glitch of the algorithm.
# 1. 修正回報欄位錯置。
# 2. 修正於不同時間下出現不一致的總資產。
#
# 2020-03-25 v2.1.1
# 1. Adding Golden Butterfly Portfolio as a benchmark portfolio.
# 1. 增加Golden Butterfly Portfolio作為基準指標。
#
# 2020-03-24 v2.1
# 1. Further diversify the hedge mode with Sixty-Fifth Plus Fund.
# 2. HSI tracking fund is no longer a substitute candidate of HKCE fund. Currently only Fidelity SaveEasy 2040 and GCE tracking fund are candidates of substituting HKCE fund.
# 3. Performance indicator calculation on first installment revised extensively, includes:
# * Adding SPY (SPDR S&P 500 ETF), QQQ (Invesco QQQ Trust), and HSI (Hang Seng Index) as a benchmark.
# * Captial Asset Pricing Model CAPM and Information Ration results are available. (The author disapprove the use of CAPM, the results of CAPM are for reference only)
# * Charting with benchmark are accomplished through tidyquant and ggplot now.
# 1. 使用65歲後基金進一步分散避險狀態下的風險。
# 2. 取代中港股票基金的替代品不再包括恒指基金。富達2040及大中華基金仍為中港股票基金的替代品。
# 3. 計算第一期投資的回報已作出重大修改,包括:
# 加入SPY,QQQ及恒生指數作基準
# 以三項基準計算資本資產定價模型CAPM及資訊比率Information Ratio的比較。
# (作者不認同CAPM是有效的評估方法。數值僅供參考。)
# 圖表改以tidyquant + ggplot展示,並加入與基準的比較。
#
# 2020-03-18 v2
# 1. Separate the return used in calculating AKANE, TORI (log) and return (simple).
# 1. 修正計算回報時使用的方式
#
# 2020-01-17 v1.3
# 1. Minor enhancement on background information.
# 1. 更改背景資料
#
# 2020-01-11 v1.2
# 1. Monthly installment: now only sell units in excess
# 1. 月供投資中,現只出售/買入所需的單位,而非全數賣出買入
#
# 2020-01-07 v1.1
# 1. Use top 2 best performing funds instead of 3
# 1. 改為只使用頭2名最好表現的基金
#
# 2020-01-07 v1
# 1. TORI is officially published and backtested with BCT MPF
# 1. 正式推出以銀聯強積金作回測的TORI介紹
#
#
# ## WARNING: Please understand nature of Tori in this section before reading further:
# This script relies on BIAS indicator and RSI indicator for investment rebalancing.
# Please understand BIAS indicator and RSI indicator before reading further.
# The script perform walk-forward validation only.
# A model will be created for exactly next re-balance only based on most updated data for that moment.
# There is no other validation or assessment on the data.
# Please be reminded that backtesting and/or walk-forward validation is no way to be accurate in predicting future performance.
# Using this script may exposed to various risk, including but not limited to overfitting, market, timing, etc.
#
# ## 警告:在應用Tori之前,請先於此節了解TORI的特性
# Tori根據乖離值及相對強弱指數作投資組合再配置。
# 閱讀下文前,請先了解乖離值及相對強弱指數。
# 此模式只使用了前移式回測(walk-forward validation)。
# 每次資產調配都會基於擁有當時而言的最新資料。
# 然而,請切記過往表現不代表將來表現。
# 使用此代碼將會受到包括但不限於以下的風險所影響:過擬合風險、市場風險、時差風險等。
#
#
# ## Background Introduction
# Mandatory Provident Fund (MPF) is one of the legal pensions in Hongkong.
# All Employers and some employees are required to pay at least 5% of monthly salary to MPF services provider.
# MPF providers are required to provide approved funds for employees to invest,
# while the team could earn management fees.
#
# However, the average annualized return of MPF is 3.1% only in from 2000-2016 (Mandatory Provident Fund Schemes Authority, 2016).
# Most Hongkong employees feels they are forced to give their salary to MPF fund manager.
# Indeed, the reasons of low return may due to inactive management by employees.
# In this example, we will explore n-m MA BIAS to rise the annualized return in Tori.
#
# ## 背景簡介
# 在香港,強積性公積金(強積金, Mandatory Provident Fund, MPF)是其中一種法定退休金。
# 僱主及部份僱員須扣除5%薪金供強積金營運商。
# 強積金營運商將每月收取管理費,並提供認可基金供僱員作投資儲蓄。
# 但是,強積金的平均回報僅3.1%,因而被批評回報有三份之一被基金經理蠶食。
# 誠言,每位僱員如以主動方式管理強積金,有助提升回報而減少收費的影響。
# 這文記錄了作者以時差平均線乖離率(n-m MA BIAS)提升強積金回報的探索(即Tori)。
#
#
# ## Tori used NMMA BIAS & Relative Strength Index (RSI)
# The author used n-m MA BIAS indicator (NMMA BIAS) to extract trend and RSI for overbuy/oversell status in Tori.
# This script combine two indicators and consider recent stock returns for the three outcome:
# 1. Tactical Asset Allocation (TAA) using Tori
# 2. Strategic Asset Allocation (SAA) using Tori and designated funds
# 3. Hedging mode (cash mode)
#
# ## Tori 應用了時差平均線乖離率及相對強
# 作者嘗試以時差平均線乖離率(n-m MA BIAS, NMMA BIAS)及相對強弱指數(RSI)分配投資物。
# 時差平均線乖離率的預測考慮了投資物的趨勢(Trend)。
# 相對強弱指數則考慮投資物的超買超賣狀態。
# 本程序結合兩項指標後,再考慮考慮最近三至六個月回報後,程式將自動決定以TORI佈置戰術性資產部署(tactical asset allocation, TAA) 、或是以TORI結果配合債券及現金的策略性資產部署(Strategic asset allocation)、或進入避險模式。
#
#
# ### When to manage my portfolio using Tori?
# The author collect historical pricing in the last day of every month to calculate NMMA BIAS and RSI in Tori.
# The same day would be the day of successful rebalance.
# In this example, MPF would have T+2 time delay for obtaining historical price, hence it is expected to have timing risks in Tori in MPF.
#
# ### 何時使用Tori管理及調配投資組合?
# 作者以每月最後一日取得投資物的歷史價格計算每月回報、NMMA BIAS及相對強弱指數。
# 然後於同日重新配置投資物。
# 在本示例中,強積金大部分均有T+2的時延,因此實際操作將有時延誤差。
#
#
# ### Tori in action: Walk-forwark validation results
# ### Tori 前移式回測結果
# Bottom of page 本頁底部
# + tags=[]
# Import section
# Internal packages
import decimal
import math
import datetime
import os
# Computation packages
import numpy as np
import numpy_financial as npf
import pandas as pd
from pyarrow import feather as f
import sqlite3 as sql3
from statsmodels.distributions.empirical_distribution import ECDF
import talib
# RPy2 packages
import rpy2.robjects as ro
from rpy2.robjects import pandas2ri, numpy2ri
numpy2ri.activate()
pandas2ri.activate()
from rpy2.robjects.packages import importr
from rpy2.robjects.pandas2ri import py2rpy as p2r
from rpy2.robjects.pandas2ri import rpy2py as r2p
# Import R packages
quantmod = importr('quantmod')
ttr = importr('TTR', robject_translations = {".env": "__env_ttr__"})
xts = importr('xts', robject_translations = {".subset.xts": "_subset_xts2",
"to.period": "to_period2"})
# +
# Constant defintion
# Paths
WORKSPACE_DIR = './'
DB_PATH = os.path.join(WORKSPACE_DIR, 'raw_data/mpf_bct.sqlite')
# trading days
TRADING_DAYS = 21
EVALUATE_TO = '2020/07/30'
# n-m MA BIAS parameters
MA_PERIOD_LONG = 6
MA_PERIOD_SHORT = 2
NM_LAG = 1
# RSI-weighted parameters
RSI_OVERBUY = 0.85
RSI_OVERBUY_VALUE = 0.2
RSI_OVERBUY_FALSE_VALUE = 1.2
RSI_PERIOD = 18
# nmMA calculation parameters
MIN_NMMA = 0.00002
DOWN_N_QUANTILE = 0.35
DOWN_O_QUANTILE = 0.45
EOHEDGE_N_QUANTILE = 0.45
EOHEDGE_O_QUANTILE = 0.35
GREEDY_MARKET_O_QUANTILE = 0.85
GREEDY_MARKET_N_QUANTILE = 0
# Fund Parameters
TOP_FUNDS = 2
EQUITY = [x for x in range(2,9)]
MIXED_ASSET_HIGH = [13, 14, 15, 18]
MIXED_ASSET_LOW = [0, 1, 16, 17]
MONEY_MARKET = [12]
FIXED_INCOME = [9, 10, 11]
HIGH_RISKS = []
HIGH_RISKS.extend(EQUITY)
HIGH_RISKS.extend(MIXED_ASSET_HIGH)
LOWER_RISKS = []
LOWER_RISKS.extend(MIXED_ASSET_LOW)
LOWER_RISKS.extend(FIXED_INCOME)
LOWER_RISKS.extend(MONEY_MARKET)
# Hedge mixing weight
HEDGE_COMPONENTS = [12, 9, 1]
HEDGE_MIXED_RETURN = -0.075
HEDGE_FULL_WEIGHT = {HEDGE_COMPONENTS[0]: [100, 50, 30], HEDGE_COMPONENTS[1]: [None, 50, 30], HEDGE_COMPONENTS[2]: [None, None, 40]}
HEDGE_MIX_WEIGHT = {HEDGE_COMPONENTS[0]: [30, 10, 8], HEDGE_COMPONENTS[1]: [None, 20, 10], HEDGE_COMPONENTS[2]: [None, None, 12]}
# +
# %%capture
# Function declaration
def ecdf(x):
last_value = x.tail(1)
temp_ecdf = ECDF(x)
if temp_ecdf(last_value) >= (RSI_OVERBUY - 1/math.sqrt(x.size)):
return RSI_OVERBUY_VALUE
else:
return RSI_OVERBUY_FALSE_VALUE-math.pow(temp_ecdf(last_value),3)
def check_down(stock_return, i, up, new_quantile=DOWN_N_QUANTILE, old_quantile=DOWN_O_QUANTILE):
if i > 7 and stock_return.iloc[i] < stock_return.iloc[0:(i+1)].quantile(new_quantile) and \
stock_return.iloc[i-3] < stock_return.iloc[0:(i+1)].quantile(old_quantile):
up = False
return up
def check_end_of_hedge(stock_return, i, up, hedge, new_quantile=EOHEDGE_N_QUANTILE, old_quantile=EOHEDGE_O_QUANTILE):
if i > 7 and hedge and stock_return.iloc[i] > stock_return.iloc[0:(i+1)].quantile(new_quantile) and \
stock_return.iloc[i-3] > stock_return.iloc[0:(i+1)].quantile(old_quantile):
up = True
hedge = False
return (up, hedge)
def check_greedy_hedge(stock_return, i, hedge, new_quantile = GREEDY_MARKET_N_QUANTILE, old_quantile=GREEDY_MARKET_O_QUANTILE):
if i > 7 and stock_return.iloc[i] < new_quantile and \
stock_return.iloc[i-1] > stock_return.iloc[0:(i+1)].quantile(old_quantile):
hedge = True
return hedge
ro.r('''
r_hedge_down <- function(portf_return, month, draw_down, up, hedge){
library("PerformanceAnalytics")
month.date <- as.Date(month, "%Y-%m-%d")
p.return.xts <- as.xts(portf_return, order.by = month.date)
is.nan.xts <- function(x) {
do.call(cbind, lapply(x, is.nan))
}
p.return.xts[is.nan(p.return.xts)] <- NA
mpf.portf.drawdown <- Drawdowns(p.return.xts, geometric=TRUE)
if (tail(na.omit(mpf.portf.drawdown), 1) < -0.065 && up == FALSE) {
hedge <- TRUE
}
r <- hedge
}
r_portf <- function(return, weight, i, month){
library("PerformanceAnalytics")
month.date <- as.Date(month, "%Y-%m-%d")
row <- i+1
is.nan.xts <- function(x) {
do.call(cbind, lapply(x, is.nan))
}
return.xts <- as.xts(return, order.by = month.date)
return.xts[is.nan(return.xts)] <- NA
weight.xts <- as.xts(weight, order.by = month.date)
weight.xts[is.nan(weight.xts)] <- NA
if (row == 1) {
r <- sum(na.fill(return.xts[row,],0) * weight.xts[row,])
} else {
r <- Return.portfolio(
na.fill(return.xts[1:row,], 0),
weight = weight.xts[1:row,],
geometric = TRUE,
rebalance_on = "months"
)
}
}
r_annual <- function(portf_return, month){
library("PerformanceAnalytics")
month.date <- as.Date(month, "%Y-%m-%d")
portf.rebal.fm <- as.xts(portf_return, order.by = month.date)
temp <- Return.annualized(portf.rebal.fm, geometric = TRUE)
}
''')
# +
# Read SQLite3 Connection
conn = sql3.connect(DB_PATH)
dcur = conn.cursor()
dcur.execute('SELECT * FROM bctMpfNav')
mpf_bct_db = dcur.fetchall()
conn.close()
# Table reshape
mpf_bct = pd.DataFrame(mpf_bct_db, columns=['id', 'Date', 'Fund', 'Nav']).drop('id', axis=1)
mpf_bct = mpf_bct.drop('Date', axis=1).set_index(pd.to_datetime(mpf_bct['Date'], format='%Y/%m/%d'))
mpf_bct = mpf_bct.pivot(columns='Fund', values='Nav').reset_index()
mpf_bct = mpf_bct.drop('Date', axis=1).set_index(pd.to_datetime(mpf_bct['Date'], format='%Y/%m/%d'))
mpf_bct = mpf_bct.truncate(after=pd.to_datetime(EVALUATE_TO, format='%Y/%m/%d'))
daily_str = np.datetime_as_string(mpf_bct.index.values, unit='D')
# +
sma_long = lambda x: talib.SMA(x, TRADING_DAYS*MA_PERIOD_LONG)
sma_short = lambda x: talib.SMA(x, TRADING_DAYS*MA_PERIOD_SHORT)
rsi = lambda x: talib.RSI(x, TRADING_DAYS*RSI_PERIOD)
mpf_bct_sma_l = mpf_bct.apply(sma_long)
mpf_bct_sma_s = mpf_bct.apply(sma_short)
mpf_bct_rsi = mpf_bct.apply(rsi)
# +
log = lambda x: np.log(x) - np.log(x.shift(1))
mpf_bct_monthly = mpf_bct.groupby(pd.Grouper(freq='BM')).last()
mpf_bct_monthly_simple = mpf_bct_monthly.pct_change()
mpf_bct_monthly_simple.iloc[0,:] = (mpf_bct_monthly.iloc[0,:] - mpf_bct.iloc[0,:]) / mpf_bct.iloc[0,:]
mpf_bct_monthly_log = mpf_bct_monthly.apply(log)
mpf_bct_monthly_log.iloc[0,:] = np.log(mpf_bct_monthly.iloc[0,:]) - np.log(mpf_bct.iloc[0,:])
monthly_str = np.datetime_as_string(mpf_bct_monthly_simple.index.values, unit='D')
# +
mpf_bct_rsi_monthly = mpf_bct_rsi.fillna(0).groupby(pd.Grouper(freq='BM')).mean()
mpf_bct_rsi_ecdf = mpf_bct_rsi_monthly.copy().fillna(0)
mpf_bct_rsi_ecdf = mpf_bct_rsi_ecdf.where(mpf_bct_rsi_ecdf != np.nan, 0)
mpf_bct_rsi_monthly_ecdf = mpf_bct_rsi_monthly.expanding(1).apply(lambda x: ecdf(x))
#mpf_bct_rsi_monthly_ecdf.MPFC = 1
mpf_bct_rsi_monthly_ecdf = mpf_bct_rsi_monthly_ecdf.apply(lambda x: x / float(x.sum()), axis=1).pow(0.25)
# + tags=[]
mpf_bct_bias_diff = (mpf_bct_sma_s - mpf_bct_sma_l)/(mpf_bct_sma_l)
mpf_bct_bias_diff_dt = (mpf_bct_bias_diff - mpf_bct_bias_diff.shift(TRADING_DAYS*1)).fillna(0).groupby(pd.Grouper(freq='BM')).mean()
mpf_bct_bias_diff_dt = mpf_bct_bias_diff_dt * mpf_bct_rsi_monthly_ecdf
mpf_bct_stock_return = mpf_bct_monthly_log.iloc[:,HIGH_RISKS].mean(axis=1)
mpf_bct_bias_diff_dt.fillna(0)
#mpf_bct_bias_diff_dt.iloc[:,] = np.where(mpf_bct_bias_diff_dt.iloc[:,] < 0, 0, mpf_bct_bias_diff_dt.iloc[:,])
mpf_bct_bias_diff_dt.iloc[:, HIGH_RISKS] = np.where(mpf_bct_bias_diff_dt.iloc[:,HIGH_RISKS] < 1e-6, 0, mpf_bct_bias_diff_dt.iloc[:,HIGH_RISKS])
mpf_bct_bias_diff_dt.iloc[:, LOWER_RISKS] = np.where(mpf_bct_bias_diff_dt.iloc[:,LOWER_RISKS] < 0, 0, mpf_bct_bias_diff_dt.iloc[:,LOWER_RISKS])
mpf_bct_bias_diff_dt = mpf_bct_bias_diff_dt.mask(mpf_bct_bias_diff_dt.rank(axis=1, method="min", ascending=False) > TOP_FUNDS, 0)
num_of_months = mpf_bct_bias_diff_dt.shape[0]
mpf_bct_bias_p = pd.DataFrame().reindex_like(mpf_bct_bias_diff_dt).fillna(0).astype(int)
mpf_portf_weight = pd.DataFrame().reindex_like(mpf_bct_monthly_simple).fillna(0).astype(np.float64)
mpf_bct_portf_return = np.zeros(num_of_months)
r_portf = ro.globalenv['r_portf']
r_hedge_down = ro.globalenv['r_hedge_down']
up = True
hedge = False
for i in range(num_of_months):
up = check_down(mpf_bct_stock_return, i, up)
up, hedge = check_end_of_hedge(mpf_bct_stock_return, i, up, hedge)
hedge = check_greedy_hedge(mpf_bct_stock_return, i, hedge)
mpf_bct_sum = mpf_bct_bias_diff_dt.iloc[i,:].sum()
#print(mpf_bct_sum)
if i <= 11 or mpf_bct_sum == mpf_bct_bias_diff_dt.iloc[i,MONEY_MARKET[0]] or mpf_bct_sum < 1e-6 or hedge:
if i >= 197:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[2]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[2]][2]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[1]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[1]][2]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[0]][2]
elif i >= 23:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[1]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[1]][1]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[0]][1]
else:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] = HEDGE_FULL_WEIGHT[HEDGE_COMPONENTS[0]][0]
elif mpf_bct_stock_return.iloc[(i-3):(i+1)].min() < HEDGE_MIXED_RETURN:
mpf_bct_bias_p.iloc[i,:] = np.around(mpf_bct_bias_diff_dt.iloc[i,:]/mpf_bct_sum*70).astype(np.int)
#print(mpf_bct_bias_p.iloc[i,:].sum())
if i >= 197:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[2]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[2]][2]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[1]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[1]][2]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[0]][2]
elif i >= 23:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[1]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[1]][1]
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[0]][1]
else:
mpf_bct_bias_p.iloc[i, HEDGE_COMPONENTS[0]] += HEDGE_MIX_WEIGHT[HEDGE_COMPONENTS[0]][0]
else:
mpf_bct_bias_p.iloc[i,:] = np.around(mpf_bct_bias_diff_dt.iloc[i,:]/mpf_bct_sum*100).astype(np.int)
# Change Global Equity to World Equity
if mpf_bct_bias_p.iloc[i, 5] != 0 and mpf_bct_monthly.iloc[i,:].name > np.datetime64('2012-08-31'):
mpf_bct_bias_p.iloc[i, 6] += mpf_bct_bias_p.iloc[i, 5]
mpf_bct_bias_p.iloc[i, 5] = 0
# Change HKCE to portfolio of Fidelity SE 2040
if mpf_bct_bias_p.iloc[i, 3] != 0 and mpf_bct_monthly.iloc[i,:].name > np.datetime64('2012-08-31'):
mpf_bct_bias_p.iloc[i, 18] += (mpf_bct_bias_p.iloc[i, 3] / 4)
mpf_bct_bias_p.iloc[i, 6] += (mpf_bct_bias_p.iloc[i, 3] / 4)
mpf_bct_bias_p.iloc[i, 7] += (mpf_bct_bias_p.iloc[i, 3] / 4)
mpf_bct_bias_p.iloc[i, 2] += (mpf_bct_bias_p.iloc[i, 3] / 4)
mpf_bct_bias_p.iloc[i, 3] = 0
elif mpf_bct_bias_p.iloc[i, 3] != 0 and mpf_bct_monthly.iloc[i,:].name > np.datetime64('2008-11-30'):
mpf_bct_bias_p.iloc[i, 18] += (mpf_bct_bias_p.iloc[i, 3] / 2)
mpf_bct_bias_p.iloc[i, 2] += (mpf_bct_bias_p.iloc[i, 3] / 2)
mpf_bct_bias_p.iloc[i, 3] = 0
elif mpf_bct_bias_p.iloc[i, 3] != 0:
mpf_bct_bias_p.iloc[i, 2] += (mpf_bct_bias_p.iloc[i, 3])
mpf_bct_bias_p.iloc[i, 3] = 0
mpf_portf_weight.iloc[i,:] = np.around(mpf_bct_bias_p.iloc[i, :] / 100, 2)
r_monthly_simple = p2r(mpf_bct_monthly_simple)
r_portf_weight = p2r(mpf_portf_weight)
mpf_bct_portf_return[i] = r_portf(r_monthly_simple, r_portf_weight, i, monthly_str)[-1]
hedge = bool(r_hedge_down(mpf_bct_portf_return, monthly_str, -0.065, up, hedge))
#print(hedge)
# + tags=[]
f.write_feather(mpf_bct_monthly_simple, "./feather/sr_mpf_bct", compression='uncompressed')
f.write_feather(mpf_portf_weight, "./feather/w_mpf_bct", compression='uncompressed')
f.write_feather(pd.DataFrame({"monthly": monthly_str}), "./feather/m_mpf_bct", compression='uncompressed')
f.write_feather(mpf_bct_monthly, "./feather/p_mpf_bct", compression='uncompressed')
# %load_ext rpy2.ipython
# + tags=[]
# %%capture
# %%R -o output1,output2,output3,fig1,fig2 -w 8 -h 4.25 --units in -r 96
library(zoo)
library(xts)
library(PerformanceAnalytics)
library(quantmod)
library(arrow)
monthly <- as.Date(as.matrix(read_feather("./feather/m_mpf_bct")))
MPF.BCT.simple.returns <- as.data.frame(read_feather("./feather/sr_mpf_bct"))
MPF.portf.weight <- as.data.frame(read_feather("./feather/w_mpf_bct"))
MPF.BCT.monthly.price <- as.data.frame(read_feather("./feather/p_mpf_bct"))
MPF.BCT.simple.returns <- as.xts(MPF.BCT.simple.returns, order.by=monthly)
MPF.portf.weight <- as.xts(MPF.portf.weight, order.by=monthly)
MPF.BCT.monthly.price <- as.xts(MPF.BCT.monthly.price, order.by=monthly)
library(tidyverse)
library(tidyquant)
library(reshape)
library(ggplot2)
portf.rebal.fm <- Return.portfolio(
MPF.BCT.simple.returns,
weight = MPF.portf.weight,
geometric = TRUE,
rebalance_on = "months"
)
etfs <- c('SPY', 'QQQ', '^HSI')
golden.butterfly <- c('VTI', 'IWM', 'SHY', 'TLT', 'GLD')
weights <- c(
1,0,0,
0,1,0,
0,0,1
)
weights.table <- tibble(etfs) %>%
tq_repeat_df(n = 3) %>%
bind_cols(tibble(weights)) %>%
group_by(portfolio)
etf.dat <- c(etfs) %>%
map_df(function(i) i %>%
tq_get("stock.prices",
from = "2001-01-01" %>%
as.Date,
to = "2020-08-01" %>%
as.Date
)
) %>% group_by(symbol) %>%
tq_transmute(select = adjusted,
mutate_fun = periodReturn,
period = "monthly",
col_rename = "Rb")
GoldBut.dat <- c(golden.butterfly) %>%
map_df(function(i) i %>%
tq_get("stock.prices",
from = "2004-11-01" %>%
as.Date,
to = "2020-08-01" %>%
as.Date
)
) %>% group_by(symbol) %>%
tq_transmute(select = adjusted,
mutate_fun = periodReturn,
period = "monthly",
col_rename = "Rb")
etf.by.column <- spread(etf.dat, symbol, Rb)
SPY.etf <- na.omit(select(etf.by.column, 'date', 'SPY')) %>% mutate(symb = 'SPY')
QQQ.etf <- na.omit(select(etf.by.column, 'date', 'QQQ')) %>% mutate(symb = 'QQQ')
HSI.etf <- na.omit(select(etf.by.column, 'date', '^HSI')) %>% mutate(symb = 'HSI')
SPY.return.monthly <- SPY.etf %>%
tq_portfolio(assets_col = symb,
returns_col = SPY,
col_rename = "Rb.SPY")
QQQ.return.monthly <- QQQ.etf %>%
tq_portfolio(assets_col = symb,
returns_col = QQQ,
col_rename = "Rb.QQQ")
HSI.return.monthly <- HSI.etf %>%
tq_portfolio(assets_col = symb,
returns_col = `^HSI`,
col_rename = "Rb.HSI")
GoldBut.return.monthly <- GoldBut.dat %>%
tq_portfolio(assets_col = symbol,
returns_col = Rb,
col_rename = "Rb.GoldBut",
rebalance_on = "years")
portf.rebal.fm.dollar <- Return.portfolio(
MPF.BCT.simple.returns,
weight = MPF.portf.weight,
geometric = TRUE,
rebalance_on = "months"
)
portf.rebal.fm.dollar.cpy <- Return.portfolio(
MPF.BCT.simple.returns,
weight = MPF.portf.weight,
geometric = TRUE,
rebalance_on = "months",
wealth.index = TRUE
)
portf.rebal.fm.dollar.gb <- Return.portfolio(
tail(MPF.BCT.simple.returns, -46),
weight = tail(MPF.portf.weight, -46),
geometric = TRUE,
rebalance_on = "months"
)
portf.rebal.fm.dollar.gb.cpy <- Return.portfolio(
tail(MPF.BCT.simple.returns,-46),
weight = tail(MPF.portf.weight,-46),
geometric = TRUE,
rebalance_on = "months",
wealth.index = TRUE
)
xts2df <- function(x) {
data.frame(date=index(x), coredata(x))
}
colnames(portf.rebal.fm.dollar) <- "Ra"
portf.rebal.fm.dollar.tb <- tbl_df(melt(xts2df(portf.rebal.fm.dollar), id="date"))
portf.rebal.fm.dollar.tb <- portf.rebal.fm.dollar.tb[,-2]
colnames(portf.rebal.fm.dollar.tb) <- c("date","Ra")
portf.rebal.fm.dollar.gb.tb <- tbl_df(melt(xts2df(portf.rebal.fm.dollar.gb), id="date"))
portf.rebal.fm.dollar.gb.tb <- portf.rebal.fm.dollar.gb.tb[,-2]
colnames(portf.rebal.fm.dollar.gb.tb) <- c("date","Ra")
portf.rebal.fm.dollar.tb[,1] <- SPY.etf[,1]
RaSPY.multiple.portfolio <- left_join(portf.rebal.fm.dollar.tb,
SPY.return.monthly, by = "date")
portf.rebal.fm.dollar.tb[,1] <- QQQ.etf[,1]
RaQQQ.multiple.portfolio <- left_join(portf.rebal.fm.dollar.tb,
QQQ.return.monthly, by = "date")
portf.rebal.fm.dollar.gb.tb[,1] <- head(GoldBut.dat$date, length(GoldBut.dat$date) / 5)
RaGoldBut.multiple.portfolio <- left_join(portf.rebal.fm.dollar.gb.tb,
GoldBut.return.monthly, by="date")
portf.rebal.fm.dollar.tb[,1] <- HSI.etf[,1]
RaHSI.multiple.portfolio <- left_join(portf.rebal.fm.dollar.tb,
HSI.return.monthly, by = "date")
#print("Annualised return of SPY:")
TORI.SPY <- RaSPY.multiple.portfolio %>%
tq_performance(Rb.SPY, NULL, performance_fun = table.AnnualizedReturns)
SPY.annualized <- TORI.SPY$AnnualizedReturn
SPY.Sharpe <- TORI.SPY$`AnnualizedSharpe(Rf=0%)`
SPY.StdDev <- TORI.SPY$AnnualizedStdDev
#print(TORI.SPY)
#print("Annualised return of QQQ:")
TORI.QQQ <- RaQQQ.multiple.portfolio %>%
tq_performance(Rb.QQQ, NULL, performance_fun = table.AnnualizedReturns)
QQQ.annualized <- TORI.QQQ$AnnualizedReturn
QQQ.Sharpe <- TORI.QQQ$`AnnualizedSharpe(Rf=0%)`
QQQ.StdDev <- TORI.QQQ$AnnualizedStdDev
#print(TORI.QQQ)
#print("Annualised return of Golden Butterfly:")
TORI.GoldBut <- RaGoldBut.multiple.portfolio %>%
tq_performance(Rb.GoldBut, NULL, performance_fun = table.AnnualizedReturns)
GB.annualized <- TORI.GoldBut$AnnualizedReturn
GB.Sharpe <- TORI.GoldBut$`AnnualizedSharpe(Rf=0%)`
GB.StdDev <- TORI.GoldBut$AnnualizedStdDev
#print(TORI.GoldBut)
#print("Annualised return of HSI:")
TORI.HSI <- RaHSI.multiple.portfolio %>%
tq_performance(Rb.HSI, NULL, performance_fun = table.AnnualizedReturns)
HSI.annualized <- TORI.HSI$AnnualizedReturn
HSI.Sharpe <- TORI.HSI$`AnnualizedSharpe(Rf=0%)`
HSI.StdDev <- TORI.HSI$AnnualizedStdDev
#print(TORI.HSI)
#print("CAPM Information against SPY:")
TORI.SPY <- RaSPY.multiple.portfolio %>%
tq_performance(Ra, Rb.SPY, performance_fun = table.CAPM)
SPY.alpha <- TORI.SPY$AnnualizedAlpha
SPY.beta <- TORI.SPY$Beta
SPY.corr <- TORI.SPY$Correlation
SPY.info <- TORI.SPY$InformationRatio
#print(TORI.SPY)
#print("CAPM Information against QQQ:")
TORI.QQQ <- RaQQQ.multiple.portfolio %>%
tq_performance(Ra, Rb.QQQ, performance_fun = table.CAPM)
QQQ.alpha <- TORI.QQQ$AnnualizedAlpha
QQQ.beta <- TORI.QQQ$Beta
QQQ.corr <- TORI.QQQ$Correlation
QQQ.info <- TORI.QQQ$InformationRatio
#print(TORI.QQQ)
#print("CAPM Information against GoldBut:")
TORI.GoldBut <- RaGoldBut.multiple.portfolio %>%
tq_performance(Ra, Rb.GoldBut, performance_fun = table.CAPM)
GB.alpha <- TORI.GoldBut$AnnualizedAlpha
GB.beta <- TORI.GoldBut$Beta
GB.corr <- TORI.GoldBut$Correlation
GB.info <- TORI.GoldBut$InformationRatio
#print(TORI.GoldBut)
#print("CAPM Information against HSI:")
TORI.HSI <- RaHSI.multiple.portfolio %>%
tq_performance(Ra, Rb.HSI, performance_fun = table.CAPM)
HSI.alpha <- TORI.HSI$AnnualizedAlpha
HSI.beta <- TORI.HSI$Beta
HSI.corr <- TORI.HSI$Correlation
HSI.info <- TORI.HSI$InformationRatio
#print(TORI.HSI)
#print("Information Ratio against SPY:")
#TORI.SPY <- RaSPY.multiple.portfolio %>%
# tq_performance(Ra, Rb.SPY, performance_fun = table.InformationRatio)
#print(TORI.SPY)
#print("Information Ratio against QQQ:")
#TORI.QQQ <- RaQQQ.multiple.portfolio %>%
# tq_performance(Ra, Rb.QQQ, performance_fun = table.InformationRatio)
#print(TORI.QQQ)
#print("Information Ratio against GoldBut:")
#TORI.GoldBut <- RaGoldBut.multiple.portfolio %>%
# tq_performance(Ra, Rb.GoldBut, performance_fun = table.InformationRatio)
#print(TORI.GoldBut)
#print("Information Ratio against HSI:")
#TORI.HSI <- RaHSI.multiple.portfolio %>%
# tq_performance(Ra, Rb.HSI, performance_fun = table.InformationRatio)
#print(TORI.HSI)
SPY.dollar.monthly <- SPY.etf %>%
tq_portfolio(assets_col = symb,
returns_col = SPY,
col_rename = "Rb.SPY",
wealth.index = TRUE) %>%
mutate(Rb.SPY = Rb.SPY * 2301)
QQQ.dollar.monthly <- QQQ.etf %>%
tq_portfolio(assets_col = symb,
returns_col = QQQ,
col_rename = "Rb.QQQ",
wealth.index = TRUE) %>%
mutate(Rb.QQQ = Rb.QQQ * 2301)
GoldBut.dollar.monthly <- GoldBut.dat %>%
tq_portfolio(assets_col = symbol,
returns_col = Rb,
col_rename = "Rb.GoldBut",
rebalance_on = "years",
wealth.index = TRUE) %>%
mutate(Rb.GoldBut = Rb.GoldBut * 2301)
HSI.dollar.monthly <- HSI.etf %>%
tq_portfolio(assets_col = symb,
returns_col = `^HSI`,
col_rename = "Rb.HSI",
wealth.index = TRUE) %>%
mutate(Rb.HSI = Rb.HSI * 2301)
portf.rebal.fm.dollar.tb.cpy <- tbl_df(melt(xts2df(portf.rebal.fm.dollar.cpy),
id="date", variable_name = "Ra"))
portf.rebal.fm.dollar.tb.cpy <- portf.rebal.fm.dollar.tb.cpy[,-2]
colnames(portf.rebal.fm.dollar.tb.cpy) <- c("date","Ra")
portf.rebal.fm.dollar.tb.cpy[,1] <- HSI.etf[,1]
portf.dollar.monthly <- portf.rebal.fm.dollar.tb.cpy %>%
mutate(Ra = Ra * 2301)
portf.rebal.fm.dollar.gb.tb.cpy <- tbl_df(melt(xts2df(portf.rebal.fm.dollar.gb.cpy),
id="date", variable_name = "Ra"))
portf.rebal.fm.dollar.gb.tb.cpy <- portf.rebal.fm.dollar.gb.tb.cpy[,-2]
colnames(portf.rebal.fm.dollar.gb.tb.cpy) <- c("date","Ra")
portf.rebal.fm.dollar.gb.tb.cpy[,1] <- head(GoldBut.dat$date, length(GoldBut.dat$date) / 5)
portf.dollar.monthly.gb <- portf.rebal.fm.dollar.gb.tb.cpy %>%
mutate(Ra = Ra * 2301)
fig1 <- ggplot()+
scale_color_manual(labels = c("SPY", "QQQ", "HSI", "Tori @ BCT"),
values = c(1,2,3,"pink3")) +
geom_line(aes(x = date, y = Rb.SPY, color = as.factor(1)), SPY.dollar.monthly, size=0.8)+
geom_line(aes(x = date, y = Rb.QQQ, color = as.factor(2)), QQQ.dollar.monthly, size=0.8)+
geom_line(aes(x = date, y = Rb.HSI, color = as.factor(3)), HSI.dollar.monthly, size=0.8)+
geom_line(aes(x = date, y = Ra, color = "pink3"), portf.dollar.monthly, size=1.8)+
labs(title = "Tori @ BCT versus SPY, QQQ, and Hang Seng Index",
subtitle = "How Tori @ BCT outperform common index fund",
x = "", y = "Portfolio Value",
color = "Portfolio or Index ETFs") +
theme_tq() +
scale_y_continuous(labels = scales::dollar)
fig2 <- ggplot()+
scale_color_manual(labels = c("Golden Butterfly", "Tori @ BCT"), values = c(4,"pink3")) +
geom_line(aes(x = date, y = Rb.GoldBut, color = as.factor(3)), GoldBut.dollar.monthly, size=0.8)+
geom_line(aes(x = date, y = Ra, color = "pink3"), portf.dollar.monthly.gb, size=1.8)+
labs(title = "Tori @ BCT versus Golden Butterfly Portfolio",
subtitle = "How Tori @ BCT outperform golden butterfly portfolio",
x = "", y = "Portfolio Value",
color = "Portfolio") +
theme_tq() +
scale_y_continuous(labels = scales::dollar)
#cat("Cumulative return: ", Return.cumulative(portf.rebal.fm, geometric=TRUE), "\n")
portf.rebal.fm.sharpe <- Return.annualized(portf.rebal.fm, geometric=TRUE) /
(StdDev.annualized(portf.rebal.fm))
rownames(portf.rebal.fm.sharpe) <- "Sharpe Ratio"
#cat("Annualised return: ", Return.annualized(portf.rebal.fm, geometric=TRUE), "\n")
mean.annual.return <-
mean(do.call(rbind, lapply(split(portf.rebal.fm, "years"), function(x)
colMeans(x))) * 12)
#cat("Mean annual return (for ref. only): ", mean.annual.return, "\n")
portf.rebal.fm.sharpe.mean <- mean.annual.return / (StdDev.annualized(portf.rebal.fm))
rownames(portf.rebal.fm.sharpe.mean) <- "Sharpe Ratio (Mean annual return)"
SharpeRatio(portf.rebal.fm, annualize = TRUE, method="modified")
#cat("Annualised Standard Deviation: ", StdDev.annualized(portf.rebal.fm), "\n")
#cat("Sortino Ratio: ", SortinoRatio(portf.rebal.fm) * sqrt(12), "\n")
#cat("Coditional Value of Risk (CVaR, Expected Shortfall ES",
# ES(portf.rebal.fm, method="modified"), "\n")
MPF.monthly.asset <- as.matrix(MPF.BCT.simple.returns)
MPF.monthly.asset[, ] <- 0
MPF.time <- 0:length(MPF.BCT.simple.returns[, 1]) / 12
MPF.pay <- -2301 + 0 * MPF.time
for (row in 1:length(MPF.BCT.simple.returns[, 1])) {
this.price <- as.matrix(MPF.BCT.monthly.price[row])
if (row == 1) {
last.value <- 2301
this.value <- last.value * this.price[13]
MPF.monthly.asset[row,] <-
na.fill(((this.value + 2301) / this.price * as.matrix(MPF.portf.weight[row, ])),0)
last.price <- this.price
} else {
last.value <-
as.numeric(sum(na.fill(last.price * MPF.monthly.asset[row - 1, ], 0)))
this.value <-
as.numeric(sum(na.fill(this.price * MPF.monthly.asset[row - 1, ], 0)))
MPF.monthly.asset[row,] <-
na.fill(((this.value + 2301) / this.price * as.matrix(MPF.portf.weight[row, ])),0)
last.price <- this.price
}
}
total.asset.value <- sum(MPF.monthly.asset[row, ] * this.price)
total.contribution <- 2301 * length(MPF.BCT.monthly.price[, 1])
library(knitr)
df <-
data.frame(
c(
"年率化回報 Annualized Return",
"累積回報 Cumulative Return",
"年率化標準誤差 Annualized SD",
"夏普比率 Sharpe Ratio",
"索丁諾比率 Sortino Ratio",
"資訊比率 Information Ratio",
"Annualised Jensen's Alpha",
"Beta",
"關聯系數 Correlation Coeff.",
"預期損失 Expected Shortfall:"
),
c(
sprintf("%.2f%%", 100*(Return.annualized(portf.rebal.fm, geometric = TRUE))),
sprintf("%.2f%%", 100*(Return.cumulative(portf.rebal.fm, geometric = TRUE))),
sprintf("%.4f%%", 100*(StdDev.annualized(portf.rebal.fm))),
sprintf("%.4f", SharpeRatio(portf.rebal.fm, annualize = TRUE, method = "modified", FUN="StdDev")),
sprintf("%.4f", (SortinoRatio(portf.rebal.fm) * sqrt(12))),
"-->",
"-->",
"-->",
"-->",
round(ES(portf.rebal.fm, method = "historical"),4)
),
c(
sprintf("%.2f%%", 100*(SPY.annualized)),
"/",
sprintf("%.2f%%", 100*(SPY.StdDev)),
sprintf("%.4f", SPY.Sharpe),
"/",
sprintf("%.4f", SPY.info),
sprintf("%.4f", SPY.alpha),
sprintf("%.4f", SPY.beta),
sprintf("%.4f", SPY.corr),
"/"
),
c(
sprintf("%.2f%%", 100*(HSI.annualized)),
"/",
sprintf("%.2f%%", 100*(HSI.StdDev)),
sprintf("%.4f", HSI.Sharpe),
"/",
sprintf("%.4f", HSI.info),
sprintf("%.4f", HSI.alpha),
sprintf("%.4f", HSI.beta),
sprintf("%.4f", HSI.corr),
"/"
),
c(
sprintf("%.2f%%", 100*(GB.annualized)),
"/",
sprintf("%.2f%%", 100*(GB.StdDev)),
sprintf("%.4f", GB.Sharpe),
"/",
sprintf("%.4f", GB.info),
sprintf("%.4f", GB.alpha),
sprintf("%.4f", GB.beta),
sprintf("%.4f", GB.corr),
"/"
)
)
output1 <- kable(df, "pandoc",
col.names = c("指標 Indicator", "Tori @ BCT", "SPY", "HSI", "Golden Butterfly"))
output2 <- c(sprintf("%.2f", total.asset.value),
sprintf("%.2f", total.contribution))
output3 <- tail(head(MPF.portf.weight, n=-1), n = 6)
# -
# ### Tori @ BCT Estimated result (based on real price published)
# ### Tori @ BCT 估計結果 (以公佈的價值計算)
#
# + tags=[]
print("First installment 第一次供款")
print("====================================================")
print(*output1, sep="\n")
print()
print()
print("Continuous installment 連續供款")
print("====================================================")
print("Monthly installment amount 每月供款額: 2301")
mpf_payment = np.append(np.full(len(mpf_portf_weight), -2301), float(output2[0])).astype(np.float32)
irr = ((1+npf.irr(mpf_payment))**12)-1
monthly_installment = ["Latest asset value 最新結餘", "Total contribution 總供款", "Internal Rate of Return (IRR) 內部回報率"]
print(f"{monthly_installment[0]}: {output2[0]} HKD")
print(f"{monthly_installment[1]}: {output2[1]} HKD")
print(f"{monthly_installment[2]}: {irr*100:.2f}%")
print()
print()
print("Last six rebalance weight 最後六次再配置的比重")
print("====================================================")
pd.DataFrame(output3, columns=mpf_bct_monthly_simple.columns.values, index=mpf_bct_monthly_simple.iloc[-7:-1].index.values)
# + magic_args="-i fig1,fig2 -w 8 -h 4.25 --units in -r 96" language="R"
# print(fig1)
# print(fig2)
| Tori_MPF_BCT_v3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this example, we will use tensorflow.keras package to create a keras image classification application using model MobileNetV2, and transfer the application to Cluster Serving step by step.
# ### Original Keras application
# We will first show an original Keras application, which download the data and preprocess it, then create the MobileNetV2 model to predict.
import tensorflow as tf
import os
import PIL
tf.__version__
# +
# Obtain data from url:"https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip"
zip_file = tf.keras.utils.get_file(origin="https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip",
fname="cats_and_dogs_filtered.zip", extract=True)
# Find the directory of validation set
base_dir, _ = os.path.splitext(zip_file)
test_dir = os.path.join(base_dir, 'validation')
# Set images size to 160x160x3
image_size = 160
# Rescale all images by 1./255 and apply image augmentation
test_datagen = tf.keras.preprocessing.image.ImageDataGenerator(rescale=1./255)
# Flow images using generator to the test_generator
test_generator = test_datagen.flow_from_directory(
test_dir,
target_size=(image_size, image_size),
batch_size=1,
class_mode='binary')
# -
# Create the base model from the pre-trained model MobileNet V2
IMG_SHAPE=(160,160,3)
model = tf.keras.applications.MobileNetV2(input_shape=IMG_SHAPE,
include_top=False,
weights='imagenet')
# In keras, input could be ndarray, or generator. We could just use `model.predict(test_generator)`. But to simplify, here we just input the first record to model.
prediction=model.predict(test_generator.next()[0])
print(prediction)
# Great! Now the Keras application is completed.
# ### Export TensorFlow SavedModel
# Next, we transfer the application to Cluster Serving. The first step is to save the model to SavedModel format.
# Save trained model to ./transfer_learning_mobilenetv2
model.save('/tmp/transfer_learning_mobilenetv2')
# ! ls /tmp/transfer_learning_mobilenetv2
# ### Deploy Cluster Serving
# After model prepared, we start to deploy it on Cluster Serving.
#
# First install Cluster Serving
# ! pip install analytics-zoo-serving
# we go to a new directory and initialize the environment
# ! mkdir cluster-serving
os.chdir('cluster-serving')
# ! cluster-serving-init
# ! tail wget-log.2
# +
# if you encounter slow download issue like above, you can just use following command to download
# # ! wget https://repo1.maven.org/maven2/com/intel/analytics/zoo/analytics-zoo-bigdl_0.12.1-spark_2.4.3/0.9.0/analytics-zoo-bigdl_0.12.1-spark_2.4.3-0.9.0-serving.jar
# if you are using wget to download, or get "analytics-zoo-xxx-serving.jar" after "ls", please call mv *serving.jar zoo.jar after downloaded.
# -
# After initialization finished, check the directory
# ! ls
# We config the model path in `config.yaml` to following (the detail of config is at [Cluster Serving Configuration](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#2-configuration))
# +
## Analytics-zoo Cluster Serving
model:
# model path must be provided
path: /tmp/transfer_learning_mobilenetv2
# -
# ! head config.yaml
# ### Start Cluster Serving
#
# Cluster Serving requires Flink and Redis installed, and corresponded environment variables set, check [Cluster Serving Installation Guide](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#1-installation) for detail.
#
# Flink cluster should start before Cluster Serving starts, if Flink cluster is not started, call following to start a local Flink cluster.
# ! $FLINK_HOME/bin/start-cluster.sh
# After configuration, start Cluster Serving by `cluster-serving-start` (the detail is at [Cluster Serving Programming Guide](https://github.com/intel-analytics/analytics-zoo/blob/master/docs/docs/ClusterServingGuide/ProgrammingGuide.md#3-launching-service))
# ! cluster-serving-start
# ### Prediction using Cluster Serving
# Next we start Cluster Serving code at python client.
from zoo.serving.client import InputQueue, OutputQueue
input_queue = InputQueue()
# In Cluster Serving, only NdArray is supported as input. Thus, we first transform the generator to ndarray (If you do not know how to transform your input to NdArray, you may get help at [data transform guide](https://github.com/intel-analytics/analytics-zoo/tree/master/docs/docs/ClusterServingGuide/OtherFrameworkUsers#data))
arr = test_generator.next()[0]
arr
# Use async api to put and get, you have pass a name arg and use the name to get
input_queue.enqueue('my-input', t=arr)
output_queue = OutputQueue()
prediction = output_queue.query('my-input')
# Use sync api to predict, this will block until the result is get or timeout
prediction = input_queue.predict(arr)
prediction
# If everything works well, the result `prediction` would be the exactly the same NdArray object with the output of original Keras model.
# don't forget to delete the model you save for this tutorial
# ! rm -rf /tmp/transfer_learning_mobilenetv2
# This is the end of this tutorial. If you have any question, you could raise an issue at [Analytics Zoo Github](https://github.com/intel-analytics/analytics-zoo/issues).
| docs/docs/ClusterServingGuide/OtherFrameworkUsers/keras-to-cluster-serving-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:geog0111-geog0111]
# language: python
# name: conda-env-geog0111-geog0111-py
# ---
# # Energy Balance Model Practical
# This practical uses a simple Energy Balance Model that is in the book by McGuffie & Henderson-Sellers. We will use it to investigate ice-albedo feedbacks and irreversibility.
#
# The [model documentation](https://moodle.ucl.ac.uk/mod/resource/view.php?id=292610) and the [book](https://moodle.ucl.ac.uk/mod/book/view.php?id=292625) can be found on the Moodle page. But I will run through the basics here first.
#
# **_You will probably want to save your own version of this file in your home directory, so that you can edit it. Select "Save as" under the "File" menu and remove the preceeding directory names_**
#
# ## Model Description
# This model is one of the simplest around, and has been constructed in a top-down fashion (rather than the bottom-up principles used for a general circulation model). It was originally proposed by [Budyko (1969)](https://moodle.ucl.ac.uk/mod/resource/view.php?id=292627).
#
# It divides the globe into 9 different boxes (each with an average temperature) and considers the impact of three different fluxes on each box. The boxes span from the equator to the pole, with evenly spaced boxes in the middle.
# 
# ## Model Equation
# The three fluxes considered by in this model are
# 1. The incoming solar radiation ($S_o$) of which a fraction, $\alpha$, is reflected back away from the box (yellow).
# 2. An outgoing longwave radiation ($L$), which depends on the temperautre of the box (red).
# 3. A latitudinal heat transport ($F$), which depends how warm the box is with respect to the rest of the globe (blue arrows, defined as positive when the transport is ouf to the box).
#
# The model is an equilibrium model (so it has no time variation in it). Therefore all the fluxes are in balance and we can write following equation for the $i$ th box.
#
# $S_i(1-\alpha_i)=L_i+F_i$
# ## Parameterisations
# #### Albedo
# Albedo, $\alpha$, is high (0.62) if ice-covered, low if not (0.3). We shall assume a grid-box is ice-covered is if is colder than a critical temperature, $T_{crit}$.
#
# $\alpha_i =
# \begin{cases}
# 0.3 & \text{if } T_i > T_{crit} \\
# 0.62 & \text{if } T_i \leq T_{crit}
# \end{cases}$
#
# #### Outgoing longwave radiation
# Top of atmosphere radiation is related to $\sigma$$T^4$ and the properties of atmosphere. As this model will only vary over a small range of $T$ (in Kelvin), we shall use a linear approximation:
#
# $L_i=A+B T_i$
#
# where $A$ and $B$ are constants derived from observations.
#
# #### Heat transports
# Heat transport is related to temperature gradients (here modelled as a form of conduction for simplicity). Having already computed the global mean temperature ($T_g$), the amount of heat transport *out* of the grid box is simply related to the a grid boxes temperature difference from the global mean.
#
# $F=C(T_i-T_g)$
#
# where C is a constant of proportionality, which has been derived from observational analysis.
# ## Numerical Approach
# We shall describe how the model is programmed below, as we start coding it up. However the basic way we will solve this model is **iteratively**. What this means is that we will
# * set the model up
# * feed in an initial guess
# * run that guess through the equations, and see what temperatures it gives
# * put those new temperatures through the equations
# * repeat that process many times - moving ever slowly closer to the correct solution
# * once the answer is steady, then it has **converged** on a stable answer
# # Program
# Start by loading the required packages into the kernal. We need *numpy* to do mathematics, *matplotlib* so that we can visualise the output, and then *pandas*: the [Python Data Analysis Library](https://pandas.pydata.org/) to build a data frame to store the model output with some metadata.
#Import packages
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Define some tunable parameters (we shall be altering these later in the exercise)
#Input quantities
FracSC=1 #solar constant as fraction of present day
A=204 #thermal A
B=2.17 #thermal B
C=3.81 #transport C
t_crit=-10 #critical temp
# Specify some important constants, and set up some essential factors related to the grid boxes. Each grid box is defined by its mean latitude, which helps specify the proportion of incoming solar radiation it receives, `SunWt`, and how much it contributes to the global average temperature, `cos_lat`.
#constant para. and names
SC=1370 #solar constant of present day in W/m^2
a=0.3 #albedo without ice
a_ice=0.62 #ice albedo
zones=['80-90','70-80','60-70','50-60','40-50','30-40','20-30','10-20','0-10'] #znoal bands
zones_mean=np.array([85,75,65,55,45,35,25,15,6]) #mean lat. of each zonal band
SunWt=np.array([0.5,0.531,0.624,0.77,0.892,1.021,1.12,1.189,1.219])
cos_lat=np.cos(zones_mean*3.1415/180)
R_in=SC/4*FracSC*SunWt #compute the incoming solar radiation at each latitude
# We need to specify our initial temperatures. These then allow us to determine the albedo at each latitude
#initial condition
init_t=[-15,-15,-5,5,10,15,18,22,24] #initial temp
init_a=np.zeros(len(init_t)) #create an array to store initial albedo
for i in range(len(zones)):
if init_t[i]<t_crit:
init_a[i]=a_ice
else:
init_a[i]=a
# Run these initial temperatures through the equation set to see what it does...
#Step 1
Tcos=init_t*cos_lat
mean_T=np.sum(Tcos)/np.sum(cos_lat)
Temp=(R_in*(1-init_a)+C*mean_T-A)/(B+C)
albedo=np.zeros(len(zones)) #create an array to store initial albedo
for i in range(len(zones)):
if Temp[i]<t_crit:
albedo[i]=a_ice
else:
albedo[i]=a
# Now that we've done it once, let's iterate it again many times (25 here) to allow it to converge.
#following steps
step_no=25 #stop number
for i in range(step_no):
Tcos=Temp*cos_lat
mean_T=np.sum(Tcos)/np.sum(cos_lat)
Temp=(R_in*(1-albedo)+C*mean_T-A)/(B+C)
albedo=np.zeros(len(zones)) #create an array to store initial albedo
for i in range(len(zones)):
if Temp[i]<t_crit:
albedo[i]=a_ice
else:
albedo[i]=a
print("Running the model results in a global mean temperature of % 5.2f oC." %mean_T)
# So far everything is stored as separate vectors. It would be much more useful to store things in a structure that can also have some metadata (information about the data). We are going to use a DataFrame, which is provided by `pandas`.
pd.DataFrame({'zones':zones,'Albedo':albedo,'Temp':Temp})
# Finally let us plot the output. Which we are going to do using the routines in `matplotlib`.
# +
#plotting
fig, ax1 = plt.subplots()
ax1.set_xlabel('lat. zones')
ax1.set_ylabel('temperature',color='r')
ax1.plot(zones, Temp,'r' )
ax1.tick_params(axis='y')
ax2 = ax1.twinx() # create a second axes that shares the same x-axis
ax2.set_ylabel('albedo',color='b')
ax2.plot(zones, albedo,'b')
ax2.tick_params(axis='y')
fig.tight_layout()
plt.show()
# -
# # Exercise and investigations with the model
#
# Having build a program of this model, we now need to use it learn something about the climate system. I want you to answer five different questions:
# 1. What fractional decrease of the solar constant is required to glaciate the world (i.e. make a snowball Earth)?
# 2. What would be the impact of increasing $C$, the heat transport parameter? Write down a hypothesis first.
# 3. The critical temperatures over land and sea are different ($0 ^oC$ and $-13 ^oC$), yet this model just represents a single hemisphere. Do you expect the Arctic or Antarctic to be colder? Test this by altering $T_{crit}$
# 4. Early estimates for $B$ ranged from 1.45 to 1.6, but how does the higher value of $B$ used here influence the climate? What does it correspond to physically?
# 5. If you set the initial temperatures to those of a snowball Earth, what fraction of solar constant is required to remove the ice from the Equator. What does this mean for the climate?
#
#
# All of these questions can be investigated by altering the "tunable parameters" and the initial temperatures. Below we have gathered these together in a single cell, followed by a single cell for you to run and plot the output (which you should not edit).
#Edit these values
FracSC=1.0 #solar constant as fraction of present day
A=204 #thermal A
B=2.17 #thermal B
C=3.81 #transport C
t_crit=-10. #critical temp
init_t=[-15.,-15.,-5.,5.,10.,15.,18.,22.,24.] #initial temp
# +
SC=1370 #solar constant of present day in W/m^2
a=0.3 #albedo without ice
a_ice=0.62 #ice albedo
zones=['80-90','70-80','60-70','50-60','40-50','30-40','20-30','10-20','0-10'] #znoal bands
zones_mean=np.array([85,75,65,55,45,35,25,15,6]) #mean lat. of each zonal band
SunWt=np.array([0.5,0.531,0.624,0.77,0.892,1.021,1.12,1.189,1.219])
cos_lat=np.cos(zones_mean*3.1415/180)
R_in=SC/4*FracSC*SunWt #compute the incoming solar radiation at each latitude
init_a=np.zeros(len(init_t)) #create an array to store initial albedo
for i in range(len(zones)):
if init_t[i]<t_crit:
init_a[i]=a_ice
else:
init_a[i]=a
#Step 1
Tcos=init_t*cos_lat
mean_T=np.sum(Tcos)/np.sum(cos_lat)
Temp=(R_in*(1-init_a)+C*mean_T-A)/(B+C)
albedo=np.zeros(len(zones)) #create an array to store initial albedo
for i in range(len(zones)):
if Temp[i]<t_crit:
albedo[i]=a_ice
else:
albedo[i]=a
#following steps
step_no=25 #stop number
for i in range(step_no):
Tcos=Temp*cos_lat
mean_T=np.sum(Tcos)/np.sum(cos_lat)
Temp=(R_in*(1-albedo)+C*mean_T-A)/(B+C)
albedo=np.zeros(len(zones)) #create an array to store initial albedo
for i in range(len(zones)):
if Temp[i]<t_crit:
albedo[i]=a_ice
else:
albedo[i]=a
print("Running the model results in a global mean temperature of % 5.2f oC." %mean_T)
pd.DataFrame({'zones':zones,'Albedo':albedo,'Temp':Temp})
#plotting
fig, ax1 = plt.subplots()
ax1.set_xlabel('lat. zones')
ax1.set_ylabel('temperature',color='r')
ax1.plot(zones, Temp,'r' )
ax1.tick_params(axis='y')
ax2 = ax1.twinx() # create a second axes that shares the same x-axis
ax2.set_ylabel('albedo',color='b')
ax2.plot(zones, albedo,'b')
ax2.tick_params(axis='y')
fig.tight_layout()
plt.show()
| practicals/Energy_Balance/EBM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SQL Lab
#
# In this lab we will learn how to use execute SQL from the ipython notebook and practice some queries on the [Northwind sample database](https://northwinddatabase.codeplex.com/) that we used in Lesson 3.1.
#
# You can access the data with this command:
#
# psql -h dsi.c20gkj5cvu3l.us-east-1.rds.amazonaws.com -p 5432 -U dsi_student northwind
# password: <PASSWORD>
#
# First of all let's install the ipython-sql extension. You can find instructions [here](https://github.com/catherinedevlin/ipython-sql).
# +
# # !pip install ipython-sql
# -
# Let's see if it works:
# %load_ext sql
# + magic_args="postgresql://dsi_student:gastudents@dsi.c20gkj5cvu3l.us-east-1.rds.amazonaws.com/northwind" language="sql"
#
# select * from orders limit 5;
# -
# Nice!!! We can now go ahead with the lab!
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# ## 1: Inspect the database
#
# If we were connected via console, it would be easy to list all tables using `\dt`. We can however access table information performing a query on the `information_schema.tables` table.
#
# ### 1.a: List Tables
#
# 1. write a `SELECT` statement that lists all the tables in the public schema of the `northwind` database, sorted alphabetically
# + language="sql"
# SELECT table_schema,table_name
# FROM information_schema.tables
# WHERE table_schema = 'public'
# ORDER BY table_name;
# -
# ### 1.b: Print Schemas
#
# The table `INFORMATION_SCHEMA.COLUMNS` contains schema information on each.
#
# Query it to display schemas of all the public tables. In particular we are interested in the column names and data types. Make sure you only include public schemas to avoid cluttering your results with a bunch of postgres related stuff.
# + language="sql"
# select table_name, column_name, data_type
# from INFORMATION_SCHEMA.COLUMNS
# where table_catalog = 'northwind'
# and table_schema = 'public'
# -
# ### 1.c: Table peek
#
# Another way of quickly looking at table information is to query the first few rows. Do this for a couple of tables, for example: `orders`, `products`, `usstates`. Display only the first 3 rows.
#
# orders
# %%sql
select * from orders limit 3
# +
# products
# +
# usstates
# -
# As you can see, some tables (like `usstates` or `region`) contain information that is probably less prone to change than other tables (like `orders` or `order_details`). This database is well organized to avoid unnecessary duplication. Let's start digging deeper in the data.
# ## 2: Products
#
# What products is this company selling? The `products` and `categories` tables contain information to answer this question.
#
# Use a combination of SQL queries and Pandas merge to answer the following questions:
#
# - What categories of products is the company selling?
# - How many products per category does the catalog contain?
# - Let's focus only on products that have not been discontinued => how many products per category?
# - What are the most expensive 5 products (not discontinued)?
# - How many units of each of these 5 products are there in stock?
# - Draw a pie chart of the categories, with slices that have the size of the number of products in that category (use non discontinued products)
# ### 2.a: What categories of products is the company selling?
#
# Remember that PostgreSQL is case sensitive.
# ### 2.b: How many products per category does the catalog contain?
#
# Keep in mind that you can cast a %sql result to a pandas dataframe using the `.DataFrame()` method.
# ### 2.c: How many not discontinued products per category?
# ### 2.d: What are the most expensive 5 products (not discontinued)?
# ### 2.e: How many units of each of these 5 products are there in stock?
# ### 2.f: Pie Chart
#
# Use pandas to make a pie chart plot.
# ## 3: Orders
#
# Now that we have a better understanding of products, let's start digging into orders.
#
# - How many orders in total?
# - How many orders per year
# - How many orders per quarter
# - Which country is receiving the most orders
# - Which country is receiving the least
# - What's the average shipping time (ShippedDate - OrderDate)
# - What customer is submitting the highest number of orders?
# - What customer is generating the highest revenue (need to pd.merge with order_details)
# - What fraction of the revenue is generated by the top 5 customers?
# ### 3.a: How many orders in total?
# ### 3.b: How many orders per year?
# ### 3.c: How many orders per quarter?
#
# Make a line plot for these.
# ### 3.d: Which country is receiving the most orders?
# ### 3.e: Which country is receiving the least?
# ### 3.f: What's the average shipping time (ShippedDate - OrderDate)?
# ### 3.g: What customer is submitting the highest number of orders?
# ### 3.h: What customer is generating the highest revenue (need to pd.merge with order_details)?
# ### 3.i: What fraction of the revenue is generated by the top 5 customers?
#
# Compare that with the fraction represented by 5 customers over the total number of customers.
# Wow!! 5.5% of the customers generate a third of the revenue!!
# ## Bonus: Other tables
#
# Investigate the content of other tables. In particular lookt at the `suppliers`, `shippers` and `employees` tables.
| general_assembly/17_databases/SQL-lab-starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Examples of hyperparameter optimization
#
# In this notebook, we provide examples for how to optimize hyperparameters of a decoder and then use the decoder with those hyperparameters. We demonstrate how to use 2 different hyperparameter optimization packages, ["BayesianOptimization"](https://github.com/fmfn/BayesianOptimization) and ["hyperopt"](http://hyperopt.github.io/hyperopt/). Both give similar performance. In the arXiv manuscript, I used "BayesianOptimization" (simply because I discovered it first).
# - The first few sections (1-3) just import packages, load the files, and preprocess them
# - Section 4 shows examples of [BayesianOptimization](https://github.com/fmfn/BayesianOptimization) for 3 decoders: Wiener Cascade, XGBoost, and Feedforward Neural Net
# - Section 5 shows examples of [hyperopt](http://hyperopt.github.io/hyperopt/) for 3 decoders: Wiener Cascade, XGBoost, and Feedforward Neural Net
# - Section 6 shows examples of making test-set predictions using the decoders with the fit hyperparameters
#
# Note that the example using the Wiener Cascade is quick, but the examples with XGBoost and the Feedforward Neural Net are slower (depending on your computer, potentially 10's of minutes).
# ## 1. Import Packages
#
# Below, we import both standard packages, and functions from the accompanying .py files
# +
#Import standard packages
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import io
from scipy import stats
import pickle
import time
# If you would prefer to load the '.h5' example file rather than the '.pickle' example file. You need the deepdish package
# import deepdish as dd
#Import function to get the covariate matrix that includes spike history from previous bins
from preprocessing_funcs import get_spikes_with_history
#Import metrics
from metrics import get_R2
from metrics import get_rho
#Import decoder functions
from decoders import WienerCascadeDecoder
from decoders import WienerFilterDecoder
from decoders import DenseNNDecoder
from decoders import SimpleRNNDecoder
from decoders import GRUDecoder
from decoders import LSTMDecoder
from decoders import XGBoostDecoder
from decoders import SVRDecoder
#Import hyperparameter optimization packages
#If either are not installed, give a warning
try:
from bayes_opt import BayesianOptimization
except ImportError:
print("\nWARNING: BayesianOptimization package is not installed. You will be unable to use section 4.")
pass
try:
from hyperopt import fmin, hp, Trials, tpe, STATUS_OK
except ImportError:
print("\nWARNING: hyperopt package is not installed. You will be unable to use section 5.")
pass
# +
#Turn off deprecation warnings
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
# ## 2. Load Data
# The data for this example can be downloaded at this [link](https://www.dropbox.com/sh/n4924ipcfjqc0t6/AACPWjxDKPEzQiXKUUFriFkJa?dl=0&preview=example_data_s1.pickle). It was recorded by <NAME> from Lee Miller's lab at Northwestern.
#
#
# The data that we load is in the format described below. We have another example notebook, "Example_format_data", that may be helpful towards putting the data in this format.
#
# Neural data should be a matrix of size "number of time bins" x "number of neurons", where each entry is the firing rate of a given neuron in a given time bin
#
# The output you are decoding should be a matrix of size "number of time bins" x "number of features you are decoding"
#
#
# +
# folder='' #ENTER THE FOLDER THAT YOUR DATA IS IN
folder='/home/jglaser/Data/DecData/'
# folder='/Users/jig289/Dropbox/Public/Decoding_Data/'
with open(folder+'example_data_s1.pickle','rb') as f:
# neural_data,vels_binned=pickle.load(f,encoding='latin1') #If using python 3
neural_data,vels_binned=pickle.load(f) #If using python 2
# #If you would prefer to load the '.h5' example file rather than the '.pickle' example file.
# data=dd.io.load(folder+'example_data_s1.h5')
# neural_data=data['neural_data']
# vels_binned=data['vels_binned']
# -
# ## 3. Preprocess Data
# ### 3A. User Inputs
# The user can define what time period to use spikes from (with respect to the output).
# I am using fewer bins in this example than in the manuscript and other examples, to make it run faster
bins_before=3 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=3 #How many bins of neural data after the output are used for decoding
# ### 3B. Format Covariates
# #### Format Input Covariates
# +
# Format for recurrent neural networks (SimpleRNN, GRU, LSTM)
# Function to get the covariate matrix that includes spike history from previous bins
X=get_spikes_with_history(neural_data,bins_before,bins_after,bins_current)
# Format for Wiener Filter, Wiener Cascade, XGBoost, and Dense Neural Network
#Put in "flat" format, so each "neuron / time" is a single feature
X_flat=X.reshape(X.shape[0],(X.shape[1]*X.shape[2]))
# -
# #### Format Output Covariates
#Set decoding output
y=vels_binned
# ### 3C. Split into training / testing / validation sets
# Note that hyperparameters should be determined using a separate validation set.
# Then, the goodness of fit should be be tested on a testing set (separate from the training and validation sets).
# #### User Options
#Set what part of data should be part of the training/testing/validation sets
#I made the ranges smaller for this example so that the hyperparameter optimization runs faster
training_range=[0.6, 0.7]
testing_range=[0.7, 0.8]
valid_range=[0.8,0.9]
# #### Split Data
# +
num_examples=X.shape[0]
#Note that each range has a buffer of"bins_before" bins at the beginning, and "bins_after" bins at the end
#This makes it so that the different sets don't include overlapping neural data
training_set=np.arange(np.int(np.round(training_range[0]*num_examples))+bins_before,np.int(np.round(training_range[1]*num_examples))-bins_after)
testing_set=np.arange(np.int(np.round(testing_range[0]*num_examples))+bins_before,np.int(np.round(testing_range[1]*num_examples))-bins_after)
valid_set=np.arange(np.int(np.round(valid_range[0]*num_examples))+bins_before,np.int(np.round(valid_range[1]*num_examples))-bins_after)
#Get training data
X_train=X[training_set,:,:]
X_flat_train=X_flat[training_set,:]
y_train=y[training_set,:]
#Get testing data
X_test=X[testing_set,:,:]
X_flat_test=X_flat[testing_set,:]
y_test=y[testing_set,:]
#Get validation data
X_valid=X[valid_set,:,:]
X_flat_valid=X_flat[valid_set,:]
y_valid=y[valid_set,:]
# -
# ### 3D. Process Covariates
# We normalize (z_score) the inputs and zero-center the outputs.
# Parameters for z-scoring (mean/std.) should be determined on the training set only, and then these z-scoring parameters are also used on the testing and validation sets.
# +
#Z-score "X" inputs.
X_train_mean=np.nanmean(X_train,axis=0)
X_train_std=np.nanstd(X_train,axis=0)
X_train=(X_train-X_train_mean)/X_train_std
X_test=(X_test-X_train_mean)/X_train_std
X_valid=(X_valid-X_train_mean)/X_train_std
#Z-score "X_flat" inputs.
X_flat_train_mean=np.nanmean(X_flat_train,axis=0)
X_flat_train_std=np.nanstd(X_flat_train,axis=0)
X_flat_train=(X_flat_train-X_flat_train_mean)/X_flat_train_std
X_flat_test=(X_flat_test-X_flat_train_mean)/X_flat_train_std
X_flat_valid=(X_flat_valid-X_flat_train_mean)/X_flat_train_std
#Zero-center outputs
y_train_mean=np.mean(y_train,axis=0)
y_train=y_train-y_train_mean
y_test=y_test-y_train_mean
y_valid=y_valid-y_train_mean
# -
# ## 4. Optimize Hyperparameters of decoders using "BayesianOptimization"
# - The general idea is that we will try to find the decoder hyperparameters that produce the highest R2 values on the validation set.
#
# - We will provide examples for a few decoders (Wiener Cascade, XGBoost, Feedforward Neural Net)
#
# A potential downside of BayesianOptimization is that it optimizes over a continuous space. So if a hyperparameter has integer values, the optimizer may unnecessarily test many nearby values (e.g. 2.05, 2.1, and 2.2) which are all treated the same (as 2), when it could just test the one integer value (2).
# ### 4A. Wiener Cascade (Linear Nonlinear Model)
# - The hyperparameter we are trying to optimize is "degree" (the degree of the polynomial).
# - Note that a sophisticated hyperparameter optimization technique is not needed for this decoder with a single hyperparameter - you could easily do a grid search. However, we show the example since it's the simplest and runs quickly.
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameter (degree)
def wc_evaluate(degree):
model_wc=WienerCascadeDecoder(degree) #Define model
model_wc.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_wc=model_wc.predict(X_flat_valid) #Validation set predictions
return np.mean(get_R2(y_valid,y_valid_predicted_wc)) #R2 value of validation set (mean over x and y position/velocity)
# #### Set range of hyperparameters, and run optimization
#
#Define Bayesian optimization, and set limits of hyperparameters
#Here, we set the limit of "degree" to be [1, 6.99], so we test degrees 1,2,3,4,5,6
wcBO = BayesianOptimization(wc_evaluate, {'degree': (1, 6.99)}, verbose=0)
#Set number of initial runs (init_points) and subsequent tests (n_iter), and do the optimization
#kappa is a parameter that sets exploration vs exploitation in the algorithm
#We set kappa=10 (greater than the default) so there is more exploration when there are more hyperparameters
wcBO.maximize(init_points=5, n_iter=5, kappa=10)
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in "wcBO.res": (each hyperparameter tested and the resulting R2 value)
#Print out the best parameters and associated R2 value (called "max_val")
wcBO.res['max']
#Assign the best hyperparameter to a variable
best_params=wcBO.res['max']['max_params']
degree=best_params['degree']
# ### 4B. XGBoost
# The hyperparameters we are trying to optimize are:
# - "max_depth" (maximum depth of the trees)
# - "num_round" (number of trees for fitting)
# - "eta" (learning rate)
#
# Note that this example can be somewhat slow (depending on your computer, potentially 10's of minutes).
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameters
def xgb_evaluate(max_depth,num_round,eta):
#The parameters need to be in the correct format for the decoder, so we do that below
max_depth=int(max_depth)
num_round=int(num_round)
eta=float(eta)
#Define model
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta)
model_xgb.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_xgb=model_xgb.predict(X_flat_valid) #Get validation set predictions
return np.mean(get_R2(y_valid,y_valid_predicted_xgb)) #Return mean validation set R2
# #### Set range of hyperparameters, and run optimization
# If you want to keep track of progress, set verbose=1 in the cell below
#Do bayesian optimization, and set limits of hyperparameters
xgbBO = BayesianOptimization(xgb_evaluate, {'max_depth': (2, 6.99), 'num_round': (100,600.99), 'eta': (0.01, 0.8)},verbose=0) #Define Bayesian optimization, and set limits of hyperparameters
#Set number of initial runs and subsequent tests, and do the optimization. Also, we set kappa=10 (greater than the default) so there is more exploration when there are more hyperparameters
xgbBO.maximize(init_points=10, n_iter=10, kappa=10)
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in "xgbBO.res": (each hyperparameter tested and the resulting R2 value)
#Print out the best parameters and associated R2 value (called "max_val")
xgbBO.res['max']
#Assign the best hyperparameters to variables, and put them in the correct format
best_params=xgbBO.res['max']['max_params'] #Get the hyperparameters that give rise to the best fit
num_round=np.int(best_params['num_round']) #We want the integer value associated with the best "num_round" parameter (which is what the xgb_evaluate function does above)
max_depth=np.int(best_params['max_depth']) #We want the integer value associated with the best "max_depth" parameter (which is what the xgb_evaluate function does above)
eta=best_params['eta']
# ### 4C. Feedforward (Dense) Neural Net
# The hyperparameters we are trying to optimize are:
# - "num_units" (the number of hidden units in each layer)
# - "frac_dropout" (the proportion of units that are dropped out"
# - "n_epochs" (the number of epochs used for fitting)
#
# Note that this example can be somewhat slow (depending on your computer, potentially 10's of minutes).
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameters
def dnn_evaluate(num_units,frac_dropout,n_epochs):
#The parameters need to be in the correct format for the decoder, so we do that below
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
#Declare and fit decoder
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs)
model_dnn.fit(X_flat_train,y_train)
#Make predictions and get R2 values on validation set
y_valid_predicted_dnn=model_dnn.predict(X_flat_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_dnn))
# #### Set range of hyperparameters, and run optimization
# If you want to keep track of progress, set verbose=1 in the cell below
# +
#Do bayesian optimization, and set limits of hyperparameters
dnnBO = BayesianOptimization(dnn_evaluate, {'num_units': (50, 700.99), 'frac_dropout': (0,.5), 'n_epochs': (2,15.99)},verbose=0)
#Set number of initial runs (init_points) and subsequent tests (n_iter), and do the optimization
#kappa is a parameter that sets exploration vs exploitation in the algorithm - 10 seems to work pretty welldnnBO = BayesianOptimization(dnn_evaluate, {'num_units': (50, 500), 'frac_dropout': (0.,.5), 'n_epochs': (2,15)})
dnnBO.maximize(init_points=10, n_iter=10, kappa=10)
# -
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in "dnnBO.res": (each hyperparameter tested and the resulting R2 value)
#Print out the best parameters and associated R2 value
dnnBO.res['max']
#Assign the best hyperparameters to variables, and put them in the correct format
best_params=dnnBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# ## 5. Optimize Hyperparameters of decoders using "Hyperopt
#
# - The general idea is that we will try to find the decoder hyperparameters that produce the highest R2 values on the validation set.
#
# - We will provide examples for a few decoders (Wiener Cascade, XGBoost, Feedforward Neural Net)
# ### 5A. Wiener Cascade
# - The hyperparameter we are trying to optimize is "degree" (the degree of the polynomial).
# - Note that a sophisticated hyperparameter optimization technique is not needed for this decoder with a single hyperparameter - you could easily do a grid search. However, we show the example since it's the simplest.
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameter (degree)
# - hyperopt minimizes the parameter, so we will return -R2 (in order to maximize R2)
def wc_evaluate2(degree):
model_wc=WienerCascadeDecoder(degree) #Define model
model_wc.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_wc=model_wc.predict(X_flat_valid) #Validation set predictions
return -np.mean(get_R2(y_valid,y_valid_predicted_wc)) #-R2 value of validation set (mean over x and y position/velocity)
# #### Set range of hyperparameters, and run optimization
# +
#The range of values I'll look at for the parameter
#"hp.quniform" will allow us to look at integer (rather than continuously spaced) values.
#Below we consider values of "degree" starting at 1, going until 6, and spaced at values of 1 (i.e., 1,2,3,4,5,6)
space = hp.quniform('degree', 1, 6, 1)
#object that holds iteration results
trials = Trials()
# -
#Do optimization
#Set the number of evaluations below (10 in this example)
hyperoptBest = fmin(wc_evaluate2, space, algo=tpe.suggest, max_evals=10, trials=trials)
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in the "trials" object. "trials.results" will give the R2 value for each hyperparameters tested, and "trials.vals" will give you the values of the hyperparameters.
print("R2_validation",-trials.best_trial['result']['loss'])
print(hyperoptBest)
degree=hyperoptBest['degree']
# ### 5B. XGBoost
# The hyperparameters we are trying to optimize are:
# - "max_depth" (maximum depth of the trees)
# - "num_round" (number of trees for fitting)
# - "eta" (learning rate)
#
# Note that this example can be somewhat slow (depending on your computer, potentially 10's of minutes).
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameter (degree)
# - hyperopt minimizes the parameter, so we will return -R2 (in order to maximize R2)
def xgb_evaluate2(params):
#Put parameters in correct formats
num_round=int(params['num_round'])
eta=float(params['eta'])
max_depth=int(params['max_depth'])
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta) #Define model
model_xgb.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_xgb=model_xgb.predict(X_flat_valid) #Get validation set predictions
return -np.mean(get_R2(y_valid,y_valid_predicted_xgb)) #Return mean validation set R2
# #### Set range of hyperparameters, and run optimization
# +
#The range of values I'll look at for the parameter
#"hp.quniform" will allow us to look at integer (rather than continuously spaced) values.
#So for "num_round", we are looking at values between 100 and 600 by 50 (100,150,200,...600)
#"hp.uniform" looks at continuously spaced values
space = {
'eta': hp.uniform('eta', 0.01, 0.8),
'num_round': hp.quniform('num_round', 100,600,50),
'max_depth': hp.quniform('max_depth', 2,6,1),
}
#object that holds iteration results
trials = Trials()
# -
#Do optimization
#Set the number of evaluations below (20 in this example)
hyperoptBest = fmin(xgb_evaluate2, space, algo=tpe.suggest, max_evals=20, trials=trials)
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in the "trials" object. "trials.results" will give the R2 value for each hyperparameters tested, and "trials.vals" will give you the values of the hyperparameters.
print("R2_validation",-trials.best_trial['result']['loss'])
# +
print(hyperoptBest)
best_params=hyperoptBest #Just renamed so it was in the same format as I used with BayesOptimization
num_round=np.int(best_params['num_round']) #We want the integer value associated with the best "num_round" parameter (which is what the xgb_evaluate function does above)
max_depth=np.int(best_params['max_depth']) #We want the integer value associated with the best "max_depth" parameter (which is what the xgb_evaluate function does above)
eta=best_params['eta']
# -
# ### 5C. Feedforward (Dense) Neural Net
# The hyperparameters we are trying to optimize are:
# - "num_units" (the number of hidden units in each layer)
# - "frac_dropout" (the proportion of units that are dropped out"
# - "n_epochs" (the number of epochs used for fitting)
#
# Note that this example can be somewhat slow (depending on your computer, potentially 10's of minutes).
# #### Define a function that returns the metric we are trying to optimize (R2 value of the validation set) as a function of the hyperparameter (degree)
# - hyperopt minimizes the parameter, so we will return -R2 (in order to maximize R2)
def dnn_evaluate2(params):
#Put parameters in proper format
num_units=int(params['num_units'])
frac_dropout=float(params['frac_dropout'])
n_epochs=int(params['n_epochs'])
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs) #Define model
model_dnn.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_dnn=model_dnn.predict(X_flat_valid) #Get validation set predictions
return -np.mean(get_R2(y_valid,y_valid_predicted_dnn)) #Return -R2 value of validation set
# #### Set range of hyperparameters, and run optimization
# +
#The range of values I'll look at for the parameter
#"hp.quniform" will allow us to look at integer (rather than continuously spaced) values.
#So for "num_units", we are looking at values between 50 and 700 by 10 (50,60,70,...700)
#"hp.uniform" looks at continuously spaced values
space = {
'frac_dropout': hp.uniform('frac_dropout', 0., 0.5),
'num_units': hp.quniform('num_units', 50,700,10),
'n_epochs': hp.quniform('n_epochs', 2,15,1),
}
#object that holds iteration results
trials = Trials()
# -
#Do optimization
#Set the number of evaluations below (20 in this example)
hyperoptBest = fmin(dnn_evaluate2, space, algo=tpe.suggest, max_evals=20, trials=trials)
# #### Get best hyperparameters
# Note that you can also find out more information about each tested hyperparameter in the "trials" object. "trials.results" will give the R2 value for each hyperparameters tested, and "trials.vals" will give you the values of the hyperparameters.
print("R2_validation",-trials.best_trial['result']['loss'])
# +
print(hyperoptBest)
best_params=hyperoptBest #Just renamed so it was in the same format as I used with BayesOptimization
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# -
# ## 6. Use the optimal hyperparameters to fit the decoder on the test set
# This can be run after running either section 4 or section 5 (both don't need to be run).
#
# ### 6A. Wiener Cascade
#"degree" was determined during hyperparameter optimization
model_wc=WienerCascadeDecoder(degree) #Declare model w/ fit hyperparameter
model_wc.fit(X_flat_train,y_train) #Fit model on training data
y_test_predicted_wc=model_wc.predict(X_flat_test) #Get test set predictions
#Print R2 values on test set
R2s_wc=get_R2(y_test,y_test_predicted_wc)
print('R2s_wc:', R2s_wc)
# ### 6B. XGBoost
# Run model w/ above hyperparameters
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta) #Declare model w/ fit hyperparameters
model_xgb.fit(X_flat_train,y_train) #Fit model
y_test_predicted_xgb=model_xgb.predict(X_flat_test) #Get test set predictions
#Print R2 values on test set
R2s_xgb=get_R2(y_test,y_test_predicted_xgb)
print('R2s:', R2s_xgb)
# ### 6C. Feedforward Neural Net
# Run model w/ above hyperparameters
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs) #Declare model w/ fit hyperparameters
model_dnn.fit(X_flat_train,y_train) #Fit model
y_test_predicted_dnn=model_dnn.predict(X_flat_test) #Get test set predictions
#Print R2 values on test set
R2s_dnn=get_R2(y_test,y_test_predicted_dnn)
print('R2s:', R2s_dnn)
| Example_hyperparam_opt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from statsmodels.graphics.tsaplots import plot_acf
from statsmodels.tsa.stattools import acf, pacf
import seaborn as sns
import matplotlib.pyplot as plt
# +
data = pd.read_csv("/Users/alexanderkell/Documents/PhD/Projects/18-battery-optimisation/data/processed/lagged_2012-2013-solar-electricity-data.csv")
data.datetime = pd.to_datetime(data.datetime)
data.head(20)
# -
solar = data[(data['Consumption Category'] == 'solar_generation') & (data['Customer'] == 1)]
solar.head()
solar_acf_coef = acf(solar.consumption, nlags=300)
# plot_acf(solar_acf_coef, lags=200)
solar_acf_coef
for category, grouped_data in data.groupby("Consumption Category"):
print(category)
data_category = data[(data['Consumption Category'] == category) & (data['Customer'] == 1)]
dat_acf_coef = acf(data_category.consumption, nlags=300)
plot_acf(dat_acf_coef, lags=300)
plot_acf(dat_acf_coef, lags=60)
plt.show()
plt.close()
| notebooks/features/2.0-ajmk-feature-selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="oJN5A4eKyT2b"
# # Setup
# + [markdown] id="q9uCUBN6yT2c"
# Import a few common modules
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 172, "status": "ok", "timestamp": 1636633385878, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="R1uvP5vDyT2d" outputId="819b2639-a78e-43bb-e459-e66e6bd3b4f6"
#CODE for points = 1
# import sklearn, numpy, os
import sklearn
import seaborn as sns
import numpy as np
import os
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
# TensorFlow ≥2.0 is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
# %load_ext tensorboard
# to make this notebook's output stable across runs
np.random.seed(42)
# + [markdown] id="jzMYiB0EyT2f"
# # Vanishing/Exploding Gradients Problem
# + executionInfo={"elapsed": 14, "status": "ok", "timestamp": 1636633386037, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="XFRcZCHRyT2g"
def logit(z):
return 1 / (1 + np.exp(-z))
# + executionInfo={"elapsed": 14, "status": "ok", "timestamp": 1636633386038, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="Q14TCXqxyT2g"
z = np.linspace(-5, 5, 200)
# + [markdown] id="jf45uObOyT2i"
# ## Xavier and He Initialization
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 15, "status": "ok", "timestamp": 1636633386039, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="iQqSsCCxyT2j" outputId="7be28639-bca8-4df5-d6f0-b19dc14fcc6b"
#CODE for points = 1
# set activation to relu and kernel initializer to he_normal
keras.layers.Dense(10, activation='relu', kernel_initializer='he_normal')
# + [markdown] id="VKeun9hyyT2k"
# ### Leaky ReLU
# + executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1636633386040, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="c1HuzTv0yT2k"
#CODE for points = 1
# remember α is the hyperparameter that defines how much the function “leaks”
# set the value of alpha, use the value typically set
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
# + [markdown] id="WUD0kkZdyT2l"
# Let's train a neural network on Fashion MNIST using the Leaky ReLU:
# + executionInfo={"elapsed": 1166, "status": "ok", "timestamp": 1636633387195, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="-ijUxHfhyT2m"
#CODE for points = 1
#load MNIST dataset from keras
(X_train_full, y_train_full), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train_full = X_train_full / 255.0
X_test = X_test / 255.0
X_valid, X_train = X_train_full[:5000], X_train_full[5000:]
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
# + executionInfo={"elapsed": 191, "status": "ok", "timestamp": 1636633387383, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="GDnaASswyT2m"
#CODE for points = 1
# initialize kernel_initializer to "he_normal" and activation function to softmax
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(100, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(10, activation="softmax")
])
# + executionInfo={"elapsed": 154, "status": "ok", "timestamp": 1636633395542, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="2qfd7pXdyT2m"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 61967, "status": "ok", "timestamp": 1636632425512, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="nEyLFBtmyT2m" outputId="7eae651a-1d09-41d7-db28-54e7b9a0d545"
history = model.fit(X_train, y_train, epochs=5,
validation_data=(X_valid, y_valid))
# + [markdown] id="wdphkBRoyT2s"
# Now look at what happens if we try to use the ReLU activation function instead:
# + executionInfo={"elapsed": 129, "status": "ok", "timestamp": 1636632432199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="szqns6OGyT2s"
np.random.seed(42)
tf.random.set_seed(42)
# + executionInfo={"elapsed": 1246, "status": "ok", "timestamp": 1636632436528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="Zu5g45C5yT2s"
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="relu", kernel_initializer="he_normal"))
for layer in range(99):
model.add(keras.layers.Dense(100, activation="relu", kernel_initializer="he_normal"))
model.add(keras.layers.Dense(10, activation="softmax"))
# + executionInfo={"elapsed": 157, "status": "ok", "timestamp": 1636632438947, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="b8ziJs2PyT2s"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 206299, "status": "ok", "timestamp": 1636632647953, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="1ATK0oSHyT2s" outputId="2cf7bb01-ede5-4818-a46a-99e3ac30642e"
history = model.fit(X_train, y_train, epochs=5,
validation_data=(X_valid, y_valid))
# + [markdown] id="Zsa-9WY5yT2t"
# Not great at all, we suffered from the vanishing/exploding gradients problem.
# + [markdown] id="NODlyFfnyT2t"
# # Batch Normalization
# + executionInfo={"elapsed": 249, "status": "ok", "timestamp": 1636632657711, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="IANuTmvzyT2t"
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(100, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(10, activation="softmax")
])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1636632657883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="jpkrSpmOyT2t" outputId="dd9a45b2-65cb-4abb-dfd2-e5bdbf9c375c"
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1636632657884, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="SeHquM9ByT2t" outputId="f96e1452-1804-449d-95c3-80e9bc6a9b33"
bn1 = model.layers[1]
[(var.name, var.trainable) for var in bn1.variables]
# + executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1636632657884, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="nzC0HW2byT2u"
#bn1.updates #deprecated
# + executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1636632657884, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="wsaT9GAeyT2u"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 82427, "status": "ok", "timestamp": 1636632753208, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="71G_x6M7yT2u" outputId="89ad1b7c-0ac4-4f84-9db4-15a095efe805"
history = model.fit(X_train, y_train, epochs=5,
validation_data=(X_valid, y_valid))
# + [markdown] id="MVtHTRXPyT2u"
# Sometimes applying BN before the activation function works better (there's a debate on this topic). Moreover, the layer before a `BatchNormalization` layer does not need to have bias terms, since the `BatchNormalization` layer some as well, it would be a waste of parameters, so you can set `use_bias=False` when creating those layers:
# + executionInfo={"elapsed": 14, "status": "ok", "timestamp": 1636632753209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="AOHLeX3gyT2v"
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(100, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(10, activation="softmax")
])
# + executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1636632753209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="xMcbKZ0OyT2v"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 51936, "status": "ok", "timestamp": 1636632880991, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="gGco3TNkyT2v" outputId="521a34d9-13ce-475f-eb80-baed0c30f1d7"
history = model.fit(X_train, y_train, epochs=5,
validation_data=(X_valid, y_valid))
# + [markdown] id="gZcjpY7TyT2z"
# # Faster Optimizers
# + [markdown] id="QzvjID6IyT20"
# ## Momentum optimization
# + executionInfo={"elapsed": 21, "status": "ok", "timestamp": 1636632880992, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="W_8slx2ByT20"
#CODE for points = 1
# initialize lr and momentum to typical values
optimizer = keras.optimizers.SGD(learning_rate=0.01, momentum=0.9)
# + [markdown] id="QTF0A3-oyT20"
# ## Nesterov Accelerated Gradient
# + executionInfo={"elapsed": 21, "status": "ok", "timestamp": 1636632880993, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="piqxQeEwyT21"
#CODE for points = 1
# initialize lr and momentum to typical values. Set nesterov so that it is used
optimizer = keras.optimizers.SGD(learning_rate=0.1, momentum=0.9, nesterov=True)
# + [markdown] id="G7VG-ueuyT2-"
# # Avoiding Overfitting Through Regularization
# + [markdown] id="to1x3hcIyT2-"
# ## $\ell_1$ and $\ell_2$ regularization
# + executionInfo={"elapsed": 210, "status": "ok", "timestamp": 1636632992431, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="saUIENUUyT2-"
#CODE for points = 0.5
# Use syntax for assigning l2 regularization with a factor 0.01 as given here - https://keras.io/api/layers/regularizers/
layer = keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=tf.keras.regularizers.l2(0.01))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 28924, "status": "ok", "timestamp": 1636633021500, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="EBqxkuQCyT2-" outputId="0fb21ca3-ecca-4774-a910-464a88435391"
#CODE for points = 1
# Use syntax for assigning l2 regularization with a factor 0.01 as given here - https://keras.io/api/layers/regularizers/
# nadam optimizer
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=tf.keras.regularizers.l2(0.01)),
keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=tf.keras.regularizers.l2(0.01)),
keras.layers.Dense(10, activation="softmax",
kernel_regularizer=tf.keras.regularizers.l2(0.01))
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train, y_train, epochs=n_epochs,
validation_data=(X_valid, y_valid))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 29454, "status": "ok", "timestamp": 1636633050937, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14784762166639519889"}, "user_tz": 300} id="IyzDPdgbyT2-" outputId="778609a2-40b4-465e-a72f-d99896d7a948"
#CODE for points = .5
# Use syntax for assigning l2 regularization with a factor 0.01 as given here - https://keras.io/api/layers/regularizers/
from functools import partial
RegularizedDense = partial(keras.layers.Dense,
activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=tf.keras.regularizers.l2(0.01))
# + id="oW0OIFKABTNZ"
#CODE for points = 1
# activation function initialized as softmax
# nadam optimizer
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
RegularizedDense(300),
RegularizedDense(100),
RegularizedDense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train, y_train, epochs=n_epochs,
validation_data=(X_valid, y_valid))
# -
| Assignments/Class Exercise 11_11 Omkar Khanvilkar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="dab5021d92c7210270cec270470e4ae179a1de01" _cell_guid="5fdcd462-5466-40f5-9992-dc31a9a174f2"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
from collections import Counter
# %matplotlib inline
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import warnings
warnings.filterwarnings('ignore')
from subprocess import check_output
print(check_output(["ls", "../input"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# -
# # Read datas
median_house_hold_in_come = pd.read_csv('../input/MedianHouseholdIncome2015.csv', encoding="windows-1252")
percentage_people_below_poverty_level = pd.read_csv('../input/PercentagePeopleBelowPovertyLevel.csv', encoding="windows-1252")
percent_over_25_completed_highSchool = pd.read_csv('../input/PercentOver25CompletedHighSchool.csv', encoding="windows-1252")
share_race_city = pd.read_csv('../input/ShareRaceByCity.csv', encoding="windows-1252")
kill = pd.read_csv('../input/PoliceKillingsUS.csv', encoding="windows-1252")
# # Poverty rate of each state
percentage_people_below_poverty_level.head()
percentage_people_below_poverty_level.shape
percentage_people_below_poverty_level['Geographic Area'].unique()
percentage_people_below_poverty_level['Geographic Area']
# *Need to plot a graph between Poverty Rate and Geographic Area*
percentage_people_below_poverty_level.poverty_rate.replace(['-'],0.0,inplace = True)
percentage_people_below_poverty_level
# +
percentage_people_below_poverty_level.poverty_rate.replace(['-'],0.0,inplace = True)
percentage_people_below_poverty_level.poverty_rate = percentage_people_below_poverty_level.poverty_rate.astype(float)
area_list = list(percentage_people_below_poverty_level['Geographic Area'].unique())
area_poverty_ratio = []
for i in area_list:
x = percentage_people_below_poverty_level[percentage_people_below_poverty_level['Geographic Area']==i]
area_poverty_rate = sum(x.poverty_rate)/len(x)
area_poverty_ratio.append(area_poverty_rate)
data = pd.DataFrame({'area_list': area_list,'area_poverty_ratio':area_poverty_ratio})
new_index = (data['area_poverty_ratio'].sort_values(ascending=False)).index.values
sorted_data = data.reindex(new_index)
sorted_data
# -
# visualization
plt.figure(figsize=(15,10))
sns.barplot(x=sorted_data['area_list'], y=sorted_data['area_poverty_ratio'])
plt.xticks(rotation= 45)
plt.xlabel('States')
plt.ylabel('Poverty Rate')
plt.title('Poverty Rate Given States')
# # Most common 15 Name or Surname of killed people
kill.head()
kill.name.value_counts()
separate = kill.name[kill.name != 'TK TK'].str.split()
separate
a,b = zip(*separate)
a
a,b = zip(*separate)
name_list = a+b
print("Name List:",name_list, "\n")
name_count = Counter(name_list)
print("Name Count:",name_count)
most_common_names = name_count.most_common(10)
print("Most Common Names:", most_common_names)
x,y = zip(*most_common_names)
x,y = list(x),list(y)
print("X List", x)
print("Y List", y)
plt.figure(figsize=(15,10))
ax= sns.barplot(x=x, y=y,palette = sns.cubehelix_palette(len(x)))
plt.xlabel('Name or Surname of killed people')
plt.ylabel('Frequency')
plt.title('Most common 10 Name or Surname of killed people')
percent_over_25_completed_highSchool.head()
percent_over_25_completed_highSchool.info()
# High school graduation rate of the population that is older than 25 in states
percent_over_25_completed_highSchool.percent_completed_hs.replace(['-'],0.0,inplace = True)
percent_over_25_completed_highSchool.percent_completed_hs = percent_over_25_completed_highSchool.percent_completed_hs.astype(float)
area_list = list(percent_over_25_completed_highSchool['Geographic Area'].unique())
area_highschool = []
for i in area_list:
x = percent_over_25_completed_highSchool[percent_over_25_completed_highSchool['Geographic Area']==i]
area_highschool_rate = sum(x.percent_completed_hs)/len(x)
area_highschool.append(area_highschool_rate)
# sorting
data = pd.DataFrame({'area_list': area_list,'area_highschool_ratio':area_highschool})
new_index = (data['area_highschool_ratio'].sort_values(ascending=True)).index.values
sorted_data2 = data.reindex(new_index)
# visualization
plt.figure(figsize=(15,10))
sns.barplot(x=sorted_data2['area_list'], y=sorted_data2['area_highschool_ratio'])
plt.xticks(rotation= 90)
plt.xlabel('States')
plt.ylabel('High School Graduate Rate')
plt.title("Percentage of Given State's Population Above 25 that Has Graduated High School")
share_race_city.head()
share_race_city.info()
# Percentage of state's population according to races that are black,white,native american, asian and hispanic
share_race_city.replace(['-'],0.0,inplace = True)
share_race_city.replace(['(X)'],0.0,inplace = True)
share_race_city.loc[:,['share_white','share_black','share_native_american','share_asian','share_hispanic']] = share_race_city.loc[:,['share_white','share_black','share_native_american','share_asian','share_hispanic']].astype(float)
area_list = list(share_race_city['Geographic area'].unique())
share_white = []
share_black = []
share_native_american = []
share_asian = []
share_hispanic = []
for i in area_list:
x = share_race_city[share_race_city['Geographic area']==i]
share_white.append(sum(x.share_white)/len(x))
share_black.append(sum(x.share_black) / len(x))
share_native_american.append(sum(x.share_native_american) / len(x))
share_asian.append(sum(x.share_asian) / len(x))
share_hispanic.append(sum(x.share_hispanic) / len(x))
# +
# visualization
f,ax = plt.subplots(figsize = (9,15))
sns.barplot(x=share_white,y=area_list,color='red',alpha = 0.5,label='White' )
sns.barplot(x=share_black,y=area_list,color='black',alpha = 0.7,label='African American')
sns.barplot(x=share_native_american,y=area_list,color='cyan',alpha = 0.6,label='Native American')
sns.barplot(x=share_asian,y=area_list,color='yellow',alpha = 0.6,label='Asian')
sns.barplot(x=share_hispanic,y=area_list,color='blue',alpha = 0.6,label='Hispanic')
ax.legend(loc='lower right',frameon = True) # legend<NAME>
ax.set(xlabel='Percentage of Races', ylabel='States',title = "Percentage of State's Population According to Races ")
# -
# # Point Plot
# +
# high school graduation rate vs Poverty rate of each state
sorted_data['area_poverty_ratio'] = sorted_data['area_poverty_ratio']/max( sorted_data['area_poverty_ratio'])
sorted_data2['area_highschool_ratio'] = sorted_data2['area_highschool_ratio']/max( sorted_data2['area_highschool_ratio'])
data = pd.concat([sorted_data,sorted_data2['area_highschool_ratio']],axis=1)
data.sort_values('area_poverty_ratio',inplace=True)
# visualize
f,ax1 = plt.subplots(figsize =(20,10))
sns.pointplot(x='area_list',y='area_poverty_ratio',data=data,color='lime',alpha=0.8)
sns.pointplot(x='area_list',y='area_highschool_ratio',data=data,color='red',alpha=0.8)
plt.text(40,0.6,'high school graduate ratio',color='red',fontsize = 17,style = 'italic')
plt.text(40,0.55,'poverty ratio',color='lime',fontsize = 18,style = 'italic')
plt.xlabel('States',fontsize = 15,color='blue')
plt.ylabel('Values',fontsize = 15,color='blue')
plt.title('High School Graduate VS Poverty Rate',fontsize = 20,color='blue')
plt.grid()
# -
# # Joint Plot
#
# Visualization of high school graduation rate vs Poverty rate of each state with different style of seaborn code
# joint kernel density
# pearsonr= if it is 1, there is positive correlation and if it is, -1 there is negative correlation.
# If it is zero, there is no correlation between variables
# Show the joint distribution using kernel density estimation
g = sns.jointplot(data.area_poverty_ratio, data.area_highschool_ratio, kind="kde", size=7)
plt.savefig('graph.png')
plt.show()
data .head()
# you can change parameters of joint plot
# kind : { “scatter” | “reg” | “resid” | “kde” | “hex” }
# Different usage of parameters but same plot with previous one
g = sns.jointplot("area_poverty_ratio", "area_highschool_ratio", data=data,size=5, ratio=3, color="r")
# Pie Chart¶
kill.race.head(15)
kill.race.value_counts()
# +
# Race rates according in kill data
kill.race.dropna(inplace = True)
labels = kill.race.value_counts().index
colors = ['grey','blue','red','yellow','green','brown']
explode = [0,0,0,0,0,0]
sizes = kill.race.value_counts().values
# visual
plt.figure(figsize = (7,7))
plt.pie(sizes, explode=explode, labels=labels, colors=colors, autopct='%1.1f%%')
plt.title('Killed People According to Races',color = 'blue',fontsize = 15)
# -
# Lm Plot
# Visualization of high school graduation rate vs Poverty rate of each state with different style of seaborn code
# lmplot
# Show the results of a linear regression within each dataset
sns.lmplot(x="area_poverty_ratio", y="area_highschool_ratio", data=data)
plt.show()
# # KDE Plot
# Visualization of high school graduation rate vs Poverty rate of each state with different style of seaborn code
# cubehelix plot
sns.kdeplot(data.area_poverty_ratio, data.area_highschool_ratio, shade=True, cut=3)
plt.show()
# # Violin Plot
data.tail()
# Show each distribution with both violins and points
# Use cubehelix to get a custom sequential palette
pal = sns.cubehelix_palette(2, rot=-.5, dark=.3)
sns.violinplot(data=data, palette=pal, inner="points")
plt.show()
# # Heatmap
#correlation map
# Visualization of high school graduation rate vs Poverty rate of each state with different style of seaborn code
f,ax = plt.subplots(figsize=(5, 5))
sns.heatmap(data.corr(), annot=True, linewidths=0.5,linecolor="red", fmt= '.1f',ax=ax)
plt.show()
# # Swarm Plot
#
# swarm plot
# manner of death(olum sekli) : ates edilerek, ates edilerek ve sok tabancasiyla
# gender cinsiyet
# age: yas
sns.swarmplot(x="gender", y="age",hue="manner_of_death", data=kill)
plt.show()
# # Pair Plot
# pair plot
sns.pairplot(data)
plt.show()
# # Count Plot
#
kill.gender.value_counts()
# kill properties
# Manner of death
sns.countplot(kill.gender)
#sns.countplot(kill.manner_of_death)
plt.title("gender",color = 'blue',fontsize=15)
# kill weapon
armed = kill.armed.value_counts()
#print(armed)
plt.figure(figsize=(10,7))
sns.barplot(x=armed[:7].index,y=armed[:7].values)
plt.ylabel('Number of Weapon')
plt.xlabel('Weapon Types')
plt.title('Kill weapon',color = 'blue',fontsize=15)
# age of killed people
above25 =['above25' if i >= 25 else 'below25' for i in kill.age]
df = pd.DataFrame({'age':above25})
sns.countplot(x=df.age)
plt.ylabel('Number of Killed People')
plt.title('Age of killed people',color = 'blue',fontsize=15)
# Race of killed people
sns.countplot(data=kill, x='race')
plt.title('Race of killed people',color = 'blue',fontsize=15)
# Most dangerous cities
city = kill.city.value_counts()
plt.figure(figsize=(10,7))
sns.barplot(x=city[:12].index,y=city[:12].values)
plt.xticks(rotation=45)
plt.title('Most dangerous cities',color = 'blue',fontsize=15)
# most dangerous states
state = kill.state.value_counts()
plt.figure(figsize=(10,7))
sns.barplot(x=state[:20].index,y=state[:20].values)
plt.title('Most dangerous state',color = 'blue',fontsize=15)
# Having mental ilness or not for killed people
sns.countplot(kill.signs_of_mental_illness)
plt.xlabel('Mental illness')
plt.ylabel('Number of Mental illness')
plt.title('Having mental illness or not',color = 'blue', fontsize = 15)
# Threat types
sns.countplot(kill.threat_level)
plt.xlabel('Threat Types')
plt.title('Threat types',color = 'blue', fontsize = 15)
# Kill numbers from states in kill data
sta = kill.state.value_counts().index[:10]
sns.barplot(x=sta,y = kill.state.value_counts().values[:10])
plt.title('Kill Numbers from States',color = 'blue',fontsize=15)
# +
# Having body cameras or not for police
sns.countplot(kill.body_camera)
plt.xlabel('Having Body Cameras')
plt.title('Having body cameras or not on Police',color = 'blue',fontsize = 15)
| Seaborn Basicis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import random
import time
import numpy as np
from indices import Indices
from indices_group import Indices_Group
from map import Map
import itertools
from functions2 import quadruplet_metric, pair_metric3, triplet_metric2, pair_metric4, rm_duplicate_ciphertexts3
from functions2 import pair_metric, cycle, cycle_list, rm_duplicate_arrays, cycle_list_gen, pair_metric2, swap, swap_list, triplet_metric, swap_list_gen
metric_results = np.zeros(len(self.ciphertexts))
for i, text in enumerate(self.ciphertexts):
metric = metric_function(text, self.natural_text)
metric_results[i] = metric
ranking = np.argsort(metric_results)
ranked_texts = [self.ciphertexts[x] for x in ranking]
filtered_ranked_texts = rm_duplicate_ciphertexts(ranked_texts, number_retained)
self.ciphertexts = filtered_ranked_texts
metric_results = np.zeros(len(trial_texts))
for i, text in enumerate(trial_texts):
metric = pair_metric(text, natural_text)
metric_results[i] = metric
ranking = np.argsort(metric_results)
ranked_texts = [trial_texts[x] for x in ranking]
filtered_ranked_texts = rm_duplicate_ciphertexts3(ranked_texts, 10)
len(metric_results)
# +
with open('message4.txt','r') as source:
message_text = source.read()
message = Indices()
message.text_in(message_text)
original = Indices()
original.text_in(message_text)
print message.text_out()
message.randomise()
print message.text_out()
with open('sample.txt','r') as source:
sample_text = source.read()
natural_text = Indices()
natural_text.text_in(sample_text)
natural_text.group_frequencies(1)
natural_text.pair_frequencies()
message.group_frequencies(1)
freq_attempt = Map()
freq_attempt.freq_key(message, natural_text)
identity = np.arange(27)
decryption_attempt = grand_permute(identity, message.rates[0])
decryption_attempt = [substitute(map,freq_attempt.map) for map in decryption_attempt]
trial_texts = [message.map(map) for map in decryption_attempt]
group = Indices_Group(trial_texts,natural_text)
natural_text.pair_frequencies()
natural_text.triplet_frequencies()
zero_map = Map(np.arange(27))
cycle_maps = zero_map.proliferate([cycle_list_gen(i) for i in range(2,5)]) + [np.arange(27)]
swap_maps = zero_map.proliferate([swap_list_gen(3)])
d_swap_maps = zero_map.proliferate([swap_list_gen(15)])
# -
len(decryption_attempt)
#initial discrimination
for i in range(1):
group.rank(pair_metric,10)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
backup_group = group
a1 = group
a2 = group
a3 = group
a1
for i in range(1):
a1.proliferate(d_swap_maps,1)
a1.rank(quadruplet_metric,1)
print a1.ciphertexts[0].text_out()
print a1.ciphertexts[0].map_record
print pair_metric3(a1.ciphertexts[0],natural_text)
for i in range(5):
a2.proliferate(d_swap_maps,1)
a2.rank(triplet_metric,1)
print a2.ciphertexts[0].text_out()
print a2.ciphertexts[0].map_record
print pair_metric3(a2.ciphertexts[0],natural_text)
print len(group.ciphertexts)
print pair_metric3(original,natural_text)
print pair_metric3(group.ciphertexts[0],natural_text)
print pair_metric3(group.ciphertexts[1],natural_text)
print freq_attempt.map
repeat_indices(message.rates[0])
for i in range(25):
group.proliferate(d_swap_maps,1)
group.rank(triplet_metric,1)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
print pair_metric3(group.ciphertexts[0],natural_text)
my_map = np.arange(27)
my_map[15] = 3
my_map[3] = 15
trial1 = my_trial.map(my_map)
print trial1.text_out()
print pair_metric3(trial1,natural_text)
print pair_metric3(my_trial,natural_text)
a = (swap_maps == my_map)
for i in range(10):
group.proliferate(d_swap_maps,20)
group.rank(triplet_metric,20)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
print pair_metric3(group.ciphertexts[0],natural_text)
for i in range(1):
group.proliferate(swap_maps,10)
group.rank(quadruplet_metric,10)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
print pair_metric3(group.ciphertexts[0],natural_text)
#generate Indices object for Indices_Group
with open('message.txt','r') as source:
original_text = source.read()
original = Indices()
original.text_in(original_text)
with open('sample.txt','r') as source:
sample_text = source.read()
natural_text = Indices()
natural_text.text_in(sample_text)
natural_text.group_frequencies(1)
freq_attempt = Map()
decryption_attempts = []
for i in range(100):
with open('message.txt','r') as source:
message_text = source.read()
message = Indices()
message.text_in(message_text)
message.randomise()
freq_attempt.freq_key(message, natural_text)
decryption_attempt = message.map(freq_attempt.map)
decryption_attempts.append(decryption_attempt.map_record)
decryption_attempts = rm_duplicate_arrays(decryption_attempts)
decryption_attempts = [original.map(x) for x in decryption_attempts]
group = Indices_Group(decryption_attempts,natural_text)
#GENEREATE MAPS
with open('sample.txt','r') as source:
sample_text = source.read()
natural_text = Indices()
natural_text.text_in(sample_text)
natural_text.group_frequencies(1)
freq_attempt = Map()
decryption_attempts = []
for i in range(100):
with open('message.txt','r') as source:
message_text = source.read()
message = Indices()
message.text_in(message_text)
message.randomise()
freq_attempt.freq_key(message, natural_text)
decryption_attempt = message.map(freq_attempt.map)
decryption_attempts.append(decryption_attempt.map_record)
decryption_attempts = rm_duplicate_arrays(decryption_attempts)
range(-2,5)
natural_text.pair_frequencies()
for i in range(20):
group.proliferate(swap_maps,20)
group.rank(pair_metric,20)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
natural_text.triplet_frequencies()
for i in range(1):
group.proliferate(d_swap_maps,10)
group.rank(triplet_metric,10)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
natural_text.group_frequencies(4)
for i in range(1):
group.proliferate(d_swap_maps,10)
group.rank(triplet_metric,10)
print group.ciphertexts[0].text_out()
print group.ciphertexts[0].map_record
ciphertexts = [sample2 for i in range(1)]
ciphertexts[0].randomise()
natural_sample.pair_frequencies()
group = Indices_Group(ciphertexts, natural_sample)
# %%timeit
group.ranking(pair_metric)
import numpy as np
values = np.array([1,2,3,1,2,4,5,6,3,2,1])
searchval = 3
ii = np.where(values == searchval)[0]
ii
rates2 = original.rates[0]
print rates2
rates[1] = 1.5
repeated_elements
repeat_indices(rates)
# +
def repeat_indices(rates):
added = []
indices = []
repeated_elements = []
for x in rates:
if (x not in added and x !=0):
added.append(x)
indices.append(np.where(rates==x)[0])
for x in indices:
if len(x) != 1:
repeated_elements.append(x)
return repeated_elements
def perm_maps(indices):
perms = itertools.permutations(indices)
subs = [np.array([indices,x]) for x in perms]
maps = [sub_to_map(sub) for sub in subs]
return maps
def sub_to_map(sub):
map = np.arange(27)
for i in range(sub[0].size):
map[sub[0][i]] = sub[1][i]
return map
def substitute(map, s_map):
new_map = np.copy(map)
for i, x in enumerate(new_map):
new_map[i] = s_map[x]
return new_map
def permute(old_maps, indices):
shuffle_maps = perm_maps(indices)
permuted_maps = []
for o_map in old_maps:
for s_map in shuffle_maps:
permuted_maps.append(substitute(o_map, s_map))
return permuted_maps
def grand_permute(map,rates):
maps = [map]
indices = repeat_indices(rates)
for index_set in indices:
maps = permute(maps, index_set)
return maps
# -
rates = np.array([1,1,2,3,2,2])
map = np.arange(6)
new_maps = grand_permute(map,rates)
new_maps
perms = itertools.permutations(indices)
subs = [np.array([indices,x]) for x in perms]
maps = [sub_to_map(sub) for sub in subs]
sub = np.array([[0,1],[0,1]])
sub_to_map(sub)
def sub_to_map(sub):
map = np.arange(27)
for i in range(sub[0].size):
map[sub[0][i]] = sub[1][i]
return map
sub_to_map(sub)
subs
def sub_to_map(sub):
#print sub
map = np.arange(27)
for i in range(sub[0].size):
print map[sub[0][i]], sub[1][i]
# map[subs[0][i]] = subs[1][i]
return map
map = np.arange(27)
for i in range(subs[0].size):
map[i] = -1
print map[subs[0][i]], subs[1][i]
#map[subs[0][i]] = subs[1][i]
a = np.arange(1000000)
random.shuffle(a)
np.argsort(a)
import random
a = np.arange(5)
random.shuffle(a)
a
| index_based/test_pad5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="63vUbrY8ooij"
# <div style="color:Navy">
#
# <div style="text-align:center">
#
# ***
# # <u>TP4:</u>
# # Filtrage et Convolutions
#
# <p style="text-align: center; color:gray"><i>@Author:</i> <NAME></p>
#
# ***
#
# </div>
#
# <u>**Plan:**</u>
#
# 1. [**Filtrage & Convolutions : notions**](#1):
# 1. Filtrage spatial: convolutions
# 2. Filtrage fréquentiel: transformée de Fourier
#
#
# 2. [**Lissage & Filtres passe-bas**](#2): Moyen, Gaussien, Median
#
#
# 3. [**Accentuation & Filtres passe-haut**](#3): Prewitt, Sobel, Laplace, Canny
#
# </div>
# + colab={"base_uri": "https://localhost:8080/", "height": 139} colab_type="code" executionInfo={"elapsed": 33681, "status": "ok", "timestamp": 1580589097214, "user": {"displayName": "Rivi\u00e8<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="Q9sJbZ6kooim" outputId="9487d9e0-5f97-49a7-e268-ecd8d5fef0c4"
'''''''''''''''''''''''''''''''''
#################################
# Code global pour tout le TP #
#################################
'''''''''''''''''''''''''''''''''
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
print("Running on Colaboratory")
from google.colab import drive, files
drive.mount('/content/gdrive', force_remount=True)
root_path = 'gdrive/My Drive/3. Doctorat/Enseignements/[Intro] Image Processing/TP4/' # A modifier à votre chemin d'accès
img_path = root_path + "img/"
else:
print("Not running on Colaboratory")
root_path = "/"
img_path = "img/"
# + [markdown] colab_type="text" id="WjVMwT41ooiy"
# # <span style="color: Green;text-decoration: underline" id="1">I. Filtrage & Convolution : notions</span>
# ***
# + [markdown] colab_type="text" id="S2G6EQodooi2"
# Le principe du **filtrage** est de modifier la valeur d'un signal (ici les pixels d'une image) sur la base des valeurs des ses voisins. C'est donc une **opération locale**, qui va se baser soit sur les voisins du pixel dans le temps (pour les séries/signaux temporels), ou dans l'espace (pour les données structurées, comme les images).
#
# Cette opération peut avoir différents objectifs: changer l'aspect du signal (i.e. améliorer ou débruiter une image) ou en extraire des caractéristiques intéressantes (coins, bordures, ...).
#
# N'entrent dans cette catégorie que les transformations se basant sur les valeurs des pixels de l'image. D'autres transformations comme les changements d'échelles, projections, translations, rotations, ... ne sont pas considérées comme des opérations de filtrage.
#
#
# > Une opération de filtrage est au **domaine fréquentiel** ce que la convolution est au **domaine spatial** (e.g. images).
# + [markdown] colab_type="text" id="6skxXikKooi6"
# ## <span style="color: DodgerBlue;text-decoration: underline">I.1 Filtrage spatial : convolutions</span>
# + [markdown] colab_type="text" id="UGiOvcHbooi-"
# ### I.1.a Principes généraux:
# + [markdown] colab_type="text" id="tq9yMKf4oojA"
# La **convolution** est une opération mathématique de deux fonctions $f$ et $g$ qui produit une troisième fonction représentant comment la forme de la première est modifiée par la seconde.
#
# Mathématiquement, le **produit de convolution** entre une fonction $f$ et $g$ est dénotée :
#
# $$
# \Large{f * g = y}
# $$
#
# Appliquée à un signal continu 1D, elle peut être formalisée comme :
#
# $$
# \Large (f*g)(t)\,=\, f(t) * g(t) \,=\, \int _{-\infty }^{\infty }f(t-\tau )g(\tau )\,d\tau
# $$
#
# Pour un filtre de taille finie $n$, appliqué à un signal discret 1D, elle peut être représentée comme:
#
# $$
# \Large (f*g)(t)\,=\, f(t) * g(t) \,=\, \sum _{\tau \ = \ 1}^{n}f(t-\tau )g(\tau )
# $$
#
#
# <br>
#
# <u>Illustration</u>: cas d'une convolution 1D de la fonction $f(t)$ par le noyau $g(t)$
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/6/6a/Convolution_of_box_signal_with_itself2.gif" width="150%">
#
# ***
#
# Dans le cas des images (signaux 2D ou 3D), la **convolution** transforme une image en modifiant la valeur de chaque pixel par une **somme pondérée des valeurs des pixels avoisinants.** Il s'agit d'une **transformation linéaire locale** vu que la nouvelle valeur d'un pixel peut être exprimée comme une somme pondérée (des valeurs de ses voisins sur l'image non-modifiée).
#
# Elle peut être formalisée comme:
#
# $$
# \Large I'(x, y) = I(x,y) * K(x,y) = \sum_{k_1\ =\ 1}^{n} \sum_{k_2\ =\ 1}^{n} I(x - k_1, y - k_2)K(k_1, k_2)
# $$
#
# Avec :
# * $K$ le *kernel*, une matrice carrée impaire de dimension $n*n$
# * $I'$ l'image convoluée résultant du produit de convolution de $I$ et de $K$
# + [markdown] colab_type="text" id="VSpzI6ySoojC"
# Ce sont les **coefficients** de cette pondération effectuée entre les deux fonctions (dans notre cas, l'image et la fonction de convolution) qui influencent l'effet de la convolution sur l'image d'origine (lissage, extraction de bordures, ...). Ces coefficients sont représentée par une matrice (carrée, impaire) que l'on appelle communément le **noyau** (*kernel*) de la convolution.
#
# Certains *kernels* possèdent une dénomination spécifique, généralement découlant d'un opérateur connu (dont ils sont une approximation discrète - e.g. filtre Gaussien ou Laplacien), ou de leur inventeur (e.g. filtre de Sobel).
#
# <img src="https://benchpartner.com/wp-content/uploads/2019/06/sobel-filter-edge-detctor.png">
#
# <u>Remarque</u>: les convolutions peuvent être vues comme une extension des opérations morphologiques :
# * Dans les OM, le noyau était binaire (ou éventuellement trinaire), et la nouvelle valeur du pixel devient 0 ou 1 en fonction de la correspondance (*hit, fit, miss*) entre l'image et le noyau.
# * Dans les convolutions, le noyau prends des valeurs entières $[0, 255]$, et la nouvelle du pixel sera également entre $[0, 255]$, calculée par un produit de convolution entre le noyau et l'image.
#
#
# ***
#
# L'objectif d'une convolution est de **transformer l'image afin de faire ressortir / extraire des informations utiles**. Etant une opération locale, la convolution suppose que l'information utile à la transformation d'un pixel donné est entièrement (ou du moins, majoritairement) contenue que dans les $3*3 = 9$ ou $5*5 = 25$ (etc, selon le choix de taille du noyau) voisins immédiats (pour une image 2D). La dimension du noyau, donc la distance de "visibilité" de la convolution à un moment donné, est également appelé son champ récepteur (*receptive field*).
#
# Cela suppose également que la valeur (et l'information contenue) d'un pixel est liée à celle de ses voisins immédiats, ce qui est le cas quand cette image contient des objets (et donc du signal), et pas uniquement du bruit (aléa). Par exemple, il est naturel de supposer que si un pixel $i$ appartient à un objet, ses voisins immédiats ont plus de chance d'y appartenir également (comparé à un pixel plus éloigné). Leurs valeurs de luminance ou de chrominance ont donc de grandes chances d'être corrélées. On peut émmettre l'hypothèse que cette dépendance des pixels avoisinants est présente dans toutes les directions (e.g. *8-connectivity*) si l'on se trouve à l'intérieur de l'objet, ou seulement dans certaines directions (e.g. *horizontal / vertical 2-connectivity*) si l'on se trouve sur une bordure, ...
#
# <img src="https://images.deepai.org/converted-papers/1906.03366/images/Connectedness.jpg" width=300>
#
# ***
# + [markdown] colab_type="text" id="qXHWABlfoojE" inputHidden=false outputHidden=false
# Une convolution est une opération en **fenêtre glissante** (*sliding window*): l'on applique le noyau de manière successive à chaque pixel de l'image, du coin haut-gauche `(0,0)` au coin bas-droit `(img.shape[0], img.shape[1])`.
#
# <img src="https://mlnotebook.github.io/img/CNN/convSobel.gif">
#
# <u>Légende</u>: La matrice centrale représente le noyau de convolution, et la matrice bleue represent l'image de sortie, également appelée *Feature Map* (surtout dans le domaine du *Deep Learning*).
#
# <br>
#
# <u>Exemple de convolution avec le filtre de Sobel horizontal</u>:
#
# 
#
# <u>Remarque</u>: Pour une illustration interactive, visiter cette page: http://setosa.io/ev/image-kernels/
#
# ***
# + [markdown] colab_type="text" id="aRkc5UikoojG"
# ### I.1.b *Strides & Padding* :
# + [markdown] colab_type="text" id="lEuWm6dioojJ"
# #### Strides:
#
# Pas de progression du kernel dans chaque direction (x et y). Par défaut, il avance de 1 en 1 pour parcourir l'ensemble des pixels.
#
# <img src="https://miro.medium.com/max/790/1*L4T6IXRalWoseBncjRr4wQ@2x.gif" alt="Exemple de convolution avec Stride = 1" title="Exemple de convolution avec Stride = 1" width="40%">
#
# <img src="https://miro.medium.com/max/721/1*4wZt9G7W7CchZO-5rVxl5g@2x.gif" alt="Exemple de convolution avec Stride = 2" title="Exemple de convolution avec Stride = 2" width="40%">
#
# Augmenter le pas permet d'avoir moins de superposition / redondance entre les champs récepteurs, et va diminuer la taille de l'image résultante.
#
# ***
# #### Padding:
#
# Ajout de marges (lignes et colonnes de pixels vides - à 0 - autour de l'image) afin d'éviter les effets de bords (qui nous feront perdre un certain nombre de lignes et de colonnes sur chaque bord de l'image). Le *padding* permet donc d'obtenir un résultat de la même dimension que l'image d'entrée.
#
# <img src="https://miro.medium.com/max/1063/1*W2D564Gkad9lj3_6t9I2PA@2x.gif" alt="Exemple de convolution avec Padding = Same" title="Exemple de convolution avec Padding = Same" width="40%">
#
# <u>Remarque</u>: le paramètre de padding d'une convolution prends généralemens l'une des 2 valeurs suivantes:
# * `Valid`: pas de padding appliqué $\rightarrow$ donc perte d'informations aux bordures, et la taille de l'image résultante sera inférieure à celle d'origine.
# * `Same`: ajout de marges dont la taille est calculée automatiquement de sorte à ce que les dimensions des images d'entrée et de sortie soient identiques.
# + [markdown] colab_type="text" id="5Jkkltv0oojL"
# ### I.1.c Types de convolutions:
# + [markdown] colab_type="text" id="iSwuq8h9oojP"
# Au delà du type de kernel employé, il existe différents types de convolutions qui dépendent de comment le noyau est appliqué à l'image :
# * Convolutions 2D (`conv2D`), mono ou multi-canaux.
# * Convolutions 3D (`conv3D`)
# * Transposed convolutions (aka Deconvolution)
# * Atrous convolutions (aka Dilated convolutions)
# * Depthwise separable convolutions
# * Flattened convolutions
# * Grouped convolutions
# * ...
#
# #### Convolutions 2D:
#
# Jusque-là, les exemples que nous avons vus étaient des convolutions 2D mono-canal : elles s'appliquaient à une image 2D avec un seul canal (niveaux de gris). Une convolution peut également être multi-canaux, par exemple si elle s'applique à chaque canal d'une image couleur. Un kernel multi-canaux peut-être calculé de deux manières différentes :
# 1. Convolution séparée (de taille `n*n*1`) sur chaque canal, et fusion des 3 images resultantes à postériori.
# 2. Convolution combinée des 3 cannaux (filtre 3D de taille `n*n*3`) donnant une image en sortie.
#
# <u>Illustration du cas 1</u>:
#
# 
#
# 
#
# <u>Illustration du cas 2</u>:
#
# 
#
# Utiliser un kernel 3D ne donne pas le même résultat de combiner le résultat de 3 convolutions 2D appliquées séparément aux 3 canaux: une convolution 3D va capter les interactions entre les valeurs / pixels des 3 canaux (en appliquant des sommes pondérées entre les mêmes pixels des différents canaux), alors que combiner les conv2D des 3 canaux suppose qu'ils sont indépendants.
#
#
# <u>Remarque</u>: plusieurs kernels différents peuvent être appliquée à une même image d'entrée pour en extraire différentes caractéristiques (*features*) :
#
# <img src="https://miro.medium.com/max/1124/1*45GSvnTvpHV0oiRr78dBiw@2x.png" width="40%">
# <img src="https://miro.medium.com/max/1022/1*hbp1VRfeWnaREPrRLnxtqQ@2x.png" width="40%">
#
# De même, l'ensemble des *feature maps* obtenues à la suite de l'application de ce groupe de filtres (*filter bank*) peuvent être passées dans une autre série de convolution, qui seront donc appliquées au résultats de la première série de convolution, pour en extraire des *features* de plus haut niveau en combinant celles obtenues précédemment. C'est un des principes fondateurs des réseaux de neurones à convolutions (*CNN*).
#
# <img src="https://miro.medium.com/max/1740/1*uUYc126RU4mnTWwckEbctw@2x.png" width="50%">
#
# ***
# + [markdown] colab_type="text" id="5au-RU8hoojR"
# #### Convolutions 3D:
#
# Les convolutions 3D sont les convolutions enployant un filtre 3D qui va parcourir l'image d'entrée dans les 3 dimensions (à ne pas confondre avec le cas 2. des conv2D ou le filtre était 3D mais ne se déplaçait que selon les axes x et y).
#
# *Similar as 2D convolutions which encode spatial relationships of objects in a 2D domain, 3D convolutions can describe the spatial relationships of objects in the 3D space. Such 3D relationship is important for some applications, such as in 3D segmentations / reconstructions of biomedical imagining, e.g. CT and MRI where objects such as blood vessels meander around in the 3D space.*
#
# <img src="https://miro.medium.com/max/1610/1*wUVVgZnzBwYKgQyTBK_5sg.png" width="40%">
# + [markdown] colab_type="text" id="Ms1kCskwoojT"
# ### I.1.d Fonctions utiles :
# + [markdown] colab_type="text" id="yIrA3CeBoojV"
# Pour les convolutions, nous allons utiliser :
# * `convolve` et `convolve2D` de Scipy
# * `filter2D` d'OpenCV
#
# ```python
# cv2.filter2D(src, ddepth, kernel)
#
# scipy.ndimage.convolve(src, kernel, mode)
# scipy.signal.convolve2D(src, kernel, mode)
# ```
# **Paramètres**
# * `src` : image de source
# * `kernel` : masque (ou noyau) de convolution à coefficients réels
# * `ddepth` : profondeur de l'image destination. Si valeur à -1: l'image destination aura la même profondeur que celle d'entrée.
# * `mode`: spécifier le comportement du filtre aux bordures (padding, marges, ...)
# + [markdown] colab_type="text" id="y_4r32jRoojY"
# ## <span style="color: DodgerBlue;text-decoration: underline">I.2 Filtrage fréquentiel</span>
# + [markdown] colab_type="text" id="45gY3-HQooji"
# ### I.2.a Domaine fréquentiel :
# + [markdown] colab_type="text" id="AQCZNyMIoojk"
# Un signal (1D, comme un son) peut être représenté de deux manières:
# * Dans le domaine temporel, ou est représentée l'évolution du signal dans le temps.
# * Dans le domaine fréquentiel, ou sont représentées les différentes **composantes fréquentielles** qui forment ce signal quand combinées ensemble.
#
# <img src="https://qph.fs.quoracdn.net/main-qimg-0cb386984a92c405f0aaeb2594761884">
#
# <img src="https://thepracticaldev.s3.amazonaws.com/i/v1p6fhprekoheceqafw1.png">
#
# Chacune de ces composantes fréquentielles est caractérisée par une amplitude, une fréquence, et une phase (son décalage dans le temps par rapport a un point d'origine commun). Décomposer un signal en une somme de composantes de différentes fréquences est permis par la **Transformée de Fourier** (TF) (cf. section suivante)
#
# #### Fréquence et images:
#
# La fréquence, dans une image, représente les **variations de l’intensité des pixels** dans l'espace (2D) :
# * Les **basses fréquences**, qui correspondent à des changements d’intensité lents, correspondent aux régions homogènes et floues de l'image.
# * Les **hautes fréquences**, qui correspondent à des changements d’intensité rapides, correspondant souvent à des petits objets, ou aux contours / frontières d'objets.
#
# Représenter une image dans le **domaine fréquentiel**, c'est représenter quelles composantes fréquentielles sont présentes dans l'image, et à quelle proportion (amplitude) elles contribuent à l'image globale.
# + [markdown] colab_type="text" id="NsigEasYoojm"
# ### I.2.b Transformée de Fourier :
# + [markdown] colab_type="text" id="2uB1-DeNoojq"
# La **Transformée de Fourier** (ou TF) décompose un signal (continu ou discret) en une série (somme pondérée) de signaux périodiques.
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/2/2b/Fourier_series_and_transform.gif" width="70%">
#
# La Transformée de Fourier peut être formalisée comme :
#
# $$
# \Large s_{N}(x) = a_n \cos_n(x) + b_n \sin_n(x)
# $$
#
# Avec :
#
# $$
# \Large a_{n} = {\frac {2}{P}}\int _{P}s(x)\cdot \cos \left(2\pi x{\tfrac {n}{P}}\right)\,dx
# $$
#
# $$
# \Large b_{n} = {\frac {2}{P}}\int _{P}s(x)\cdot \sin \left(2\pi x{\tfrac {n}{P}}\right)\,dx
# $$
#
# Qui peuvent être combinées en :
#
# $$
# \Large s_{N}(x)={\frac {a_{0}}{2}}+\sum _{n=1}^{N}\left(a_{n}\cos \left({\tfrac {2\pi nx}{P}}\right)+b_{n}\sin \left({\tfrac {2\pi nx}{P}}\right)\right)
# $$
#
# Chacune des composantes de la décomposition de Fourier est un signal périodique caractérisé par une amplitude, une fréquence et une phase. La combinaison de ces composantes permet de retracer le signal d'origine: la variation dans le temps (ou dans l'espace) de la série recrée le signal (ou image) d'origine.
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/1/1a/Fourier_series_square_wave_circles_animation.gif" width="70%">
#
# La transformée de Fourier est une **opération inversible**: l'on peut récupérer le signal (ou l'image) d'origine à partir de sa représentation dans le domaine fréquentiel, sans perte d'informations.
#
# Cette transformée permet donc de **passer du domaine temporel ou spatial au domaine fréquentiel**, et vice-versa.
# + [markdown] colab_type="text" id="i7HZuKu5oojr"
# ### I.2.c TF Discrète et images :
# + [markdown] colab_type="text" id="CtYFh6Smoojv"
# Une image peut être considérée comme un signal discret (et cyclique, si l'on considère qu'elle boucle sur elle-même) qui varie en deux dimensions spatiales: les axes $x$ et $y$ de l'image. Cela nous permet d'appliquer une TF discrète (*DFT*) à l'image selon les dimensions x et y pour obtenir la représentation fréquentielle de celle-ci.
#
# *The DFT is the sampled Fourier Transform and therefore does not contain all frequencies forming an image, but only a set of samples which is large enough to fully describe the spatial domain image. The number of frequencies corresponds to the number of pixels in the spatial domain image, i.e. the image in the spatial and Fourier domain are of the same size.*
#
# Pour une image carrée $N*N$, la DFT est donnée par (en notation exponentielle complexe) :
#
# $$
# \Large F(u,v) = \sum _{x=0}^{N-1}\sum _{y=0}^{N-1} F(x,y)\cdot e^{-i2\pi \left(\frac{u \cdot x}{N} + \frac{v \cdot y}{N}\right)}
# $$
#
#
# L'équation peut être interprétée comme: la valeur de chaque point de l'espace de Fourier ($F(u,v)$) est obtenue en multipliant la valeur du pixel dans l'image spatiale $F(x,y)$ par la *base function* correspondante $ e^{-i2\pi \left(\frac{u \cdot x}{N} + \frac{v \cdot y}{N}\right)} $, sommé pour tout x et y.
#
#
# ****
# Chaque valeur retournée par la TF est un nombre complexe qui peut être divisé en deux composantes:
#
# $$
# \Large F(u,v) = F_R(u,v) + j*F_I(u,v)
# $$
#
# Avec:
# * La composante réelle, $F_R(u,v) = |F(u,v)|$, également appelée le **spectre de magnitude** (*magnitude spectrum*)
# * La composante imaginaire, $F_I(u,v) = \arctan{\frac{F_I(u,v)}{F_R(u,v)}}$, également appelé **spectre de phase**.
#
# ****
# Une image représentée dans l'espace de Fourier est une représentation visuelle de son spectre de magnitude, ou l'intensité de chaque pixel est donnée par l'importance de la fréquence qu'il représente :
#
# <img src="https://akshaysin.github.io/images/beforeNafter.JPG">
#
# Cet espace représente l'ensemble des composantes fréquentielles présentes dans l'image d'origine:
# * Leur **fréquence** (basse ou haute) est donnée par leur éloignement au centre de l'image: les hautes fréquences sont celles qui sont éloignées du centre.
# * Leur **amplitude**, alias l'importance de leur contribution à l'image d'origine (donc si elles sont très présentes ou non dedans) est représentée par l'intensité lumineuse du pixel qui correspond à cette composante fréquentielle dans l'espace de Fourier.
# * Leur **orientation**, représentée par leur position sur le graph du domaine fréquentiel, qui correspond à l'orientation spatiale de cette composante fréquentielle dans l'image d'origine.
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/f/fa/2D_Fourier_Transform_and_Base_Images.png">
#
# *Ici avec $k_x$ = $u$ et $k_y$ = $v$*
# + [markdown] colab_type="text" id="CRDTaaBroojx"
# ### I.2.d Filtrage :
# + [markdown] colab_type="text" id="kN9TozZUoojy"
# Filtrer un signal, c'est **modifier l'importance de certaines composantes fréquentielle** de ce signal (à partir de sa décomposition de Fourier). Leur importance (amplitude) sera modulée par les valeurs du filtre qui sera appliqué (passe-bas, passe-haut, ou la combinaison des deux: passe-bande).
#
# Le processus filtrage consiste en une succession d'étapes :
# 1. On transforme le signal / image vers le domaine fréquentiel.
# 2. L'on applique un filtre à l'image transformée, ce qui correspond **appliquer un masque (en niveaux de gris)** afin d'éliminer (ou diminuer l'importance de) certaines bandes de fréquence.
# 3. L'on applique la TF inverse à la décomposition de Fourier filtrée pour retourner dans le domaine temporel / spatial et ainsi récupérer le signal ou l'image filtrée.
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/8/8b/Fourier_filtering.png">
#
# <u>Remarque</u>: L'avantage d'effectuer un filtrage dans le domaine fréquentiel plutot qu'une convolution dans le domaine temporel ou spatial est le gain en temps de calcul (pour les images assez grandes, i.e. généralement de plus de 500 * 500 pixels). La plupart des algorithmes de convolution modernes choisissent d'appliquer le filtrage en fréquentielle automatiquement si l'image dépasse une certaine résolution.
# + [markdown] colab_type="text" id="hW3vvBLcooj0"
# ### I.2.e Fonctions utiles :
# + [markdown] colab_type="text" id="joWLCUTuooj1"
# Pour le filtrage en domaine fréquentiel, nous allons nous servir des méthodes suivantes:
# * Numpy: `np.fft.fft2()` et son inverse, `np.fft.ifft2()`
# * OpenCV: `cv2.dft()` et son inverse, `cv2.idft()`
# * Scipy: `scipy.signal.fftconvolve()`
#
# <u>Remarque</u>: *FFT* signifie *Fast Fourier Transform*
#
# <u>Remarque 2</u>: les méthodes d'`OpenCV` sont mieux optimisées que celle de `numpy`pour les TF:
#
# 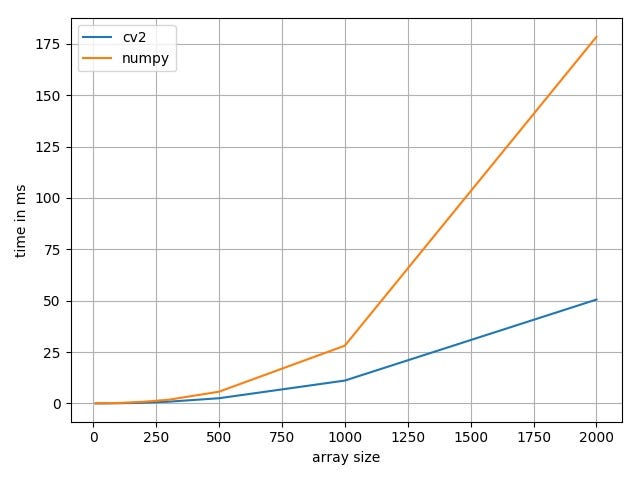
# + [markdown] colab_type="text" id="EA4WLRZnooj2"
# # <span style="color: Green;text-decoration: underline" id="2">II. Lissage & Filtres passe-bas</span>
# + [markdown] colab_type="text" id="m_ynzzHpooj3"
# Un filtre de **lissage** (*smoothing*) est un filtre **passe-bas** (*low-pass*), car il laisse passer les basses-fréquences (évolutions "lentes"), et élimine les hautes-fréquences (évolutions "rapides / soudaines" : les détails). Il va donc rendre l'image plus **floue** en éliminant les détails fins.
#
# Il existe différents types de filtres passe-bas, qui vont flouter l'image de manière différente:
# * Filtre moyen
# * Filtre Gaussien
# * Filtre Médian
# * Filtre Bilatéral
#
# Dans le **domaine fréquentiel**, un filtre passe-bas est tout simplement un masque (en niveaux de gris) où les valeurs sont plus élevées près du centre, afin de laisser passer les basses fréquences (qui sont proches du centre) préférentiellement.
#
# Exemple de filtre passe-bas, appelé la **fenêtre de Hamming**:
# 
# + [markdown] colab_type="text" id="x_sy5PUVooj5"
# **Remarque:** les filtres les plus connus (Gaussien, Median, ...) sont implémentés par des fonctions spécifiques dans la plupart des libraries de CV.
#
# **Avec OpenCV:**
# ```python
# cv2.blur()
# cv2.GaussianBlur()
# cv2.medianBlur()
# cv2.bilateralFilter()
# ```
#
# **Avec Scipy:**
# ```Python
# filters.gaussian_filter(img, 5)
# ```
#
# **Avec Pillow:**
# ```Python
# img.filter(ImageFilter.GaussianBlur(5))
# ```
# + colab={} colab_type="code" id="apJYScivooj7"
### Imports et fonctions utiles à cette partie
import os, cv2
import numpy as np
import pandas as pd
from PIL import Image, ImageOps
from scipy.signal import convolve, fftconvolve, convolve2d
# Interactivité
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import plotly.express as px # Librairie de visualisation avancée (similaire à Seaborn)
import plotly.graph_objects as go
from matplotlib.pylab import *
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits import mplot3d
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina' # Améliorer la netteté des figures
plt.rcParams["figure.figsize"] = 12.8, 9.6
import warnings # "Do not disturb" mode
warnings.filterwarnings('ignore')
# Ajout de bruit Gaussien
def gaussian_noise(img, mean = 0.0, std = 10.0):
noisy_gauss = img + np.random.normal(mean, std, img.shape)
return np.array(np.clip(noisy_gauss, 0, 255), dtype='uint8')
def affichage_1x2(src, dst):
fig = plt.figure(figsize=(12, 6))
plt.subplot(121), plt.imshow(src, cmap="gray", origin="upper"), plt.title("Image d'origine")
plt.subplot(122), plt.imshow(dst, cmap="gray", origin="upper"), plt.title("Image filtrée")
plt.show()
def affichage_2x2(img1, img2, img3, img4):
fig = plt.figure(figsize=(12, 12))
plt.subplot(221), plt.imshow(img1, cmap="gray", origin="upper"), plt.title("Original")
plt.subplot(222), plt.imshow(img2, cmap="gray", origin="upper"), plt.title("Filtrée (FFT)")
plt.subplot(223), plt.imshow(img3, cmap="hot", extent=(-img3.shape[0]//2, img3.shape[0]//2, -img3.shape[1]//2, img3.shape[1]//2)), plt.title("[Fourier] Original")
plt.subplot(224), plt.imshow(img4, cmap="hot", extent=(-img4.shape[0]//2, img4.shape[0]//2, -img4.shape[1]//2, img4.shape[1]//2)), plt.title("[Fourier] Filtré")
plt.show()
def kernel_heatmap(kernel, cm="gray"):
fig = px.imshow(kernel, color_continuous_scale=cm)
fig.show()
def kernel_3D(kernel):
z_data = pd.DataFrame(kernel)
fig = go.Figure(data=[go.Surface(z=z_data.values)])
fig.update_layout(title='Kernel', autosize=True,
width=500, height=500,
margin=dict(l=65, r=50, b=65, t=90))
fig.show()
def kernel_3D_continuous(data, color_map="viridis"):
x,y,z = data
X1 = np.reshape(x, -1)
Y1 = np.reshape(y, -1)
Z1 = np.reshape(z, -1)
# Normalize the colors based on Z value
norm = plt.Normalize(z.min(), z.max())
colors = cm.jet(norm(z))
ax = plt.axes(projection='3d')
ax.get_proj = lambda: np.dot(mplot3d.Axes3D.get_proj(ax), np.diag([1, 1, 1, 1]))
ax.plot_trisurf(X1, Y1, Z1, cmap=color_map)
# + [markdown] colab_type="text" id="8b28bJYzookA" inputHidden=false outputHidden=false
# ## <span style="color: DodgerBlue;text-decoration: underline">II.1 Le filtre moyen</span>
# + [markdown] colab_type="text" id="JIl9h2zXookC" inputHidden=false outputHidden=false
# ``` python
# K = 1/9 * np.array([[1, 1, 1],[1, 1, 1],[1, 1, 1],])
# ```
#
# <img src="https://opencv-python-tutroals.readthedocs.io/en/latest/_images/math/42f61cdcb41615a23af32b0fd95e674090afdc8d.png">
#
# Appliquer cette matrice revient en fait à remplacer la valeur de chaque pixel par la moyenne du pixel en cours et de ses 8 voisins immédiats (matrice 3x3).
#
# **Remarque**: Rappelons que les valeurs des composantes des pixels sont des nombres entiers compris entre 0 et $255$. Si les valeurs post-convolution ne sont plus des entiers, il faudra les arrondir : min(x, 255) et max(x, 0), où x est la nouvelle valeur.
# + [markdown] colab_type="text" id="_DVinpOnookD"
# ### II.1.a Dans le domaine spatial:
# + [markdown] colab_type="text" id="OwA9D7NQcU9m"
# Convolution "manuelle": on définit nous-même le noyau de convolution puis on l'applique à l'image.
# + colab={"base_uri": "https://localhost:8080/", "height": 464, "referenced_widgets": ["c2291b6350e049a7977f81e73f452f87", "4613ba379a3346b2baf334de2de1019f", "<KEY>", "0c6eed08da6b41c0a70fea7b6d5fec2d", "174ba2f81bed488bb3be9a5a61bc5278", "283d228f653043ddafef33dd4356c549", "66a6aa950de04e64b3e478bbf89e10c2", "25e59bebe92048a0b812f9a8495ab4b0", "abedb46ec07a49408cc93db1dda0c24f", "cbce81a642534fdba2cff8af7e6cc0b4", "01d8a11a66044902a5d0a32b6e445343", "7354962fc4e04321954e12859ba8c3ab"]} colab_type="code" executionInfo={"elapsed": 1288, "status": "ok", "timestamp": 1580589452733, "user": {"displayName": "Rivi\u00<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="XhME_-BjookE" outputId="fdd8e224-3843-43d5-d3dc-5cdc8df7df51"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact_manual
def mean_filter(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], kernel_size=kernel_size_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
#Application du bruit puis filtrage
noisy = gaussian_noise(img)
# Création du noyau de convolution (kernel)
kernel = 1/kernel_size * np.ones((kernel_size,kernel_size), dtype="uint8")
# Débruitage
denoised_average = convolve2d(noisy, kernel, mode="same")
# On affiche
affichage_1x2(noisy, denoised_average)
# + [markdown] colab_type="text" id="IJOHi99fookI"
# ## <span style="color: DodgerBlue;text-decoration: underline">II.2 Le filtre médian</span>
# + [markdown] colab_type="text" id="7u0k_tNgookJ"
# Le **filtre médian** permet de réduire le bruit tout en conservant les contours de l'image.
# L'idée principale du filtre médian est de remplacer chaque pixel par la valeur médiane de son voisinage immédiat.
#
# Le filtre médian est efficace pour nettuyer du bruit multiplicatif, comme le bruit Sel&Poivre.
#
# <img src="https://www.mathworks.com/matlabcentral/mlc-downloads/downloads/submissions/46563/versions/2/screenshot.jpg">
# -
# #### Avec `cv2.medianBlur` :
# + colab={"base_uri": "https://localhost:8080/", "height": 432, "referenced_widgets": ["e90dfd1838a74399add0184db5aaff50", "2282ea182a0b4986b9f7062a6a0cf5d8", "2c78a27e022f4b4dbeeb29389ad4ee68", "b0a463360b2c42ec98e9cd5552bbc772", "786ebc6be1a84f959ea2d8f5f0649600", "<KEY>", "077a79c23bbf4310af3f27e5ffbb1ef1", "<KEY>", "116ba43b9ce946d582d85db19536d86d"]} colab_type="code" executionInfo={"elapsed": 3350, "status": "ok", "timestamp": 1580478684114, "user": {"displayName": "Rivi\u00e8<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="JwTXi7fQookK" outputId="50aa2170-4267-4b01-e30c-8d2c6f342199"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact_manual
def median_filter(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], kernel_size=kernel_size_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
noisy = gaussian_noise(img)
denoised_median = cv2.medianBlur(noisy, ksize=kernel_size)
# On affiche
affichage_1x2(noisy, denoised_median)
# + [markdown] colab_type="text" id="QtQRB3h3ookO"
# ## <span style="color: DodgerBlue;text-decoration: underline">II.3 Filtre Gaussien</span>
# + [markdown] colab_type="text" id="X3OEIad-ookR"
# Le **filtre Gaussien** permet de faire un moyennage pondéré des pixels l'image: chaque pixel est remplacé par une somme pondérée de ses voisins, et les poids de cette pondération suivent l'évolution d'une Gaussienne 2D.
#
# <u>Illustration</u>: Gaussienne 2D de variances $\sigma_x$ et $\sigma_y$, donnée par l'équation:
#
# $$
# G(x,y) = \frac{1}{2\pi\sigma^2} e^{-\frac{x^2+y^2}{2\sigma^2}}
# $$
# +
from scipy.stats import multivariate_normal
def Gaussian(mu=[0.0, 0.0], sigma=[.5, .5]):
n = sigma[0] + sigma[1] *3
x, y = np.mgrid[-n:n:30j, -n:n:30j]
xy = np.column_stack([x.flat, y.flat])
covariance = np.diag(np.array(sigma)**2)
z = multivariate_normal.pdf(xy, mean=np.array(mu), cov=covariance)
z = z.reshape(x.shape)
return x,y,z
@interact
def gaussiant_filter(sigmaX=(0.5,5,0.5), sigmaY=(0.5,5,0.5)):
kernel_3D_continuous(Gaussian([0.0, 0.0], [sigmaX, sigmaY]))
# + colab={"base_uri": "https://localhost:8080/", "height": 464, "referenced_widgets": ["76949b4910b24361abb3534b42031db7", "fe5d2d2bbd964afca9d15a9d256e638f", "<KEY>", "990b18e934d7489fb3d34d601e3fbca9", "<KEY>", "55b068e3036548f385920952fed6c05b", "e54998040b2c4e98bee8181756ebe9cc", "191a0d754bf44002b91ac7efdd615352", "<KEY>", "<KEY>", "02bfd6c1e9d44e0f9891c0a1096c5272", "83f4f4280c004fbcb081de8f9081c216"]} colab_type="code" executionInfo={"elapsed": 3438, "status": "ok", "timestamp": 1580478776160, "user": {"displayName": "Rivi\u00e8<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="8hDx9LTzookT" outputId="8e72640b-816d-42ca-9f0f-f484fbf52d1d"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact
def gaussiant_filter(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], kernel_size=kernel_size_slider, noise=(0,30,1), sigma=(0.5,5,0.5)):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# Application d'un bruit Gaussien puis filtrage Gaussien
noisy = gaussian_noise(img, 0, noise)
denoised_gaussian = cv2.GaussianBlur(noisy, ksize=(kernel_size,kernel_size), sigmaX=sigma, sigmaY=sigma, borderType=cv2.BORDER_DEFAULT)
# On affiche
affichage_1x2(noisy, denoised_gaussian)
# + [markdown] colab_type="text" id="FxwjTGmJookX"
# ## <span style="color: DodgerBlue;text-decoration: underline">II.4 Filtrage fréquentiel : filtre passe-bas</span>
# + [markdown] colab_type="text" id="2y7y8UXwookY"
# ##### Avec Numpy:
# + colab={"base_uri": "https://localhost:8080/", "height": 811, "referenced_widgets": ["216913aa7d7e434bb4872c7453479a7d", "<KEY>", "<KEY>", "<KEY>", "66e63540e7b8404eb70eaea351c35284", "<KEY>", "e33d97bc0cce476daeb5b0e738ddf933", "9d445774dfbd47f9a454de63dc0ada3f", "<KEY>", "a369dc78ffbe4ab09b3647dbeb5e892f", "774c53c2c00a4e2190c5f1ad9fc149c1", "cb0ed6657fdc4fcf8ef6146d3f15c11a"]} colab_type="code" executionInfo={"elapsed": 939, "status": "ok", "timestamp": 1580589488364, "user": {"displayName": "Rivi\u00e8<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="ObUhIpsRookZ" outputId="217b9a8d-b89a-470c-8c57-29c4d45830b2"
@interact_manual
def mean_filter1_fft(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], mask_size=(0,100,10)):
img = np.array(Image.open(img_path + image).convert("L"))
## Spatial --> Fréquentiel
# On applique la transformée de Fourier discrète
dft = np.fft.fft2(img)
# On shift le résultat pour le centrer
dft_shift = np.fft.fftshift(dft)
# Calcul du spectre de magnitude (représentation de Fourier)
magnitude_img = 20 * np.log(np.abs(dft_shift))
# Filtrage: création d'un mask binaire noir carré, sauf en son centre
rows, cols = img.shape
crow, ccol = rows//2 , cols//2
mask = np.zeros((rows,cols), np.uint8)
mask[crow-mask_size:crow+mask_size, ccol-mask_size:ccol+mask_size] = 1
## On applique le masque
img_filtered = dft_shift * mask
# Et calcule sa représentation de Fourier
magnitude_img_filtered = 20 * np.log(np.abs(img_filtered))
### Fréquentiel --> Spatial
# Shift inverse
img_filtered = np.fft.ifftshift(img_filtered)
# Inversion de la TF
img_filtered = np.fft.ifft2(img_filtered)
img_filtered = np.abs(img_filtered)
affichage_2x2(img, img_filtered, magnitude_img, magnitude_img_filtered)
# + [markdown] colab_type="text" id="sfzijli5ooke"
# ##### Avec OpenCV:
# + colab={"referenced_widgets": ["8cc84ab8a46f4b27bf0fad837254effd"]} colab_type="code" id="-zCICIZqookh" outputId="c4747513-b8ab-4f6e-f41d-efc7ee82b17e"
@interact_manual
def mean_filter1_fft(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], mask_size=(0,100,10)):
img = np.array(Image.open(img_path + image).convert("L"))
# Spatial --> Fréquentiel
dft = cv2.dft(np.float32(img), flags=cv2.DFT_COMPLEX_OUTPUT)
dft_shift = np.fft.fftshift(dft)
# Calcul du spectre de magnitude (représentation de Fourier)
magnitude_img = 20 * np.log(cv2.magnitude(dft_shift[:,:,0], dft_shift[:,:,1]))
# Filtrage: création d'un mask binaire noir carré, sauf en son centre
rows, cols = img.shape
crow, ccol = rows//2 , cols//2
mask = np.zeros((rows,cols,2), np.uint8)
mask[crow-mask_size:crow+mask_size, ccol-mask_size:ccol+mask_size] = 1
# On applique le masque
img_filtered = dft_shift * mask
# Et calcule sa représentation de Fourier
magnitude_img_filtered = 20 * np.log(cv2.magnitude(img_filtered[:,:,0], img_filtered[:,:,1]))
# Fréquentiel --> Spatial
img_filtered = np.fft.ifftshift(img_filtered)
img_filtered = cv2.idft(img_filtered)
img_filtered = cv2.magnitude(img_filtered[:,:,0], img_filtered[:,:,1])
affichage_2x2(img, img_filtered, magnitude_img, magnitude_img_filtered)
# + [markdown] colab_type="text" id="pu5eG4lJookl"
# #### Avec `fftconvolve`de `Scipy`, qui s'occupe de la TF et TF inverse après filtrage :
# + [markdown] colab_type="text" id="Zn40intLookn"
# Cette méthode permet d'appliquer directement un kernel (spatial) à l'image, en passant par l'espace de Fourier.
# + colab={"base_uri": "https://localhost:8080/", "height": 464, "referenced_widgets": ["1e6c8ec076fc413c95c2b8857cf5206b", "<KEY>", "<KEY>", "323e660a89c141ca87c8b37d5557dd1d", "<KEY>", "6fff9e64f34742f4b6e8dfacda5e8be6", "<KEY>", "61c7644ee6004f0ea0843ddaf8c02a6c", "70ce5b2c38974a488fb92e3a542eec41", "34539a0210004cfdaa90ad68b63a22fe", "b91317996d774d8a92054a9096c2daaf", "20c598b6a1f6412cbb908561d0aa1101"]} colab_type="code" executionInfo={"elapsed": 1142, "status": "ok", "timestamp": 1580479091758, "user": {"displayName": "Rivi\u00e8<NAME>\u00e8le", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBNfjDm1Y0wcm0EDe7v_Sdi-WtWb5EtJA-TV9SO=s64", "userId": "04667194983314352464"}, "user_tz": -60} id="AbKjF57aooko" outputId="e389ee49-adac-4c92-a938-67aa4d69171a"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact_manual
def mean_filter3_fft(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], kernel_size=kernel_size_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
kernel = 1/kernel_size * np.ones((kernel_size,kernel_size), dtype="uint8")
noisy = gaussian_noise(img, 0, 15)
fft_filtered = fftconvolve(noisy, kernel, mode='same')
affichage_1x2(noisy, fft_filtered)
# + [markdown] colab_type="text" id="5svVFQT7ookr"
# ### <span style="color:crimson">**[<u>Exercice</u>]** A vous de jouer:</span>
# ***
# <div style="color:DarkSlateBlue">
#
# 1. **Créez une méthode interactive permettant de générer une image carrée (en niveaux de gris) dont les valeurs seront affectées selon l'équation $f(x,y) = \sin(2\pi (\lambda_x*x + \lambda_y*y))$ avec des sliders pour modifier les valeurs de $\lambda_x$ et $\lambda_y$ (entre 0 et 1, avec un pas de 0.1).**
#
# > <u>Astuce</u>:
# ```python
# n = 300
# xs = np.linspace(-2*np.pi, 2*np.pi, n)
# ys = np.linspace(-2*np.pi, 2*np.pi, n)
# x, y = np.meshgrid(xs, ys)
# z = # TODO: calcul de l'onde sin 2D
# ```
#
#
# 2. **Créer une méthode permettant de visualiser l'image résultante dans le domaine spatial et dans le domaine fréquentiel (représentation logarithmique absolue shiftée).**
#
# > <u>Astuce</u>: vous pouvez vous inspirer de la méthode `kernel_heatmap` pour l'affichage dans le domaine spatial.
#
# > Faire varier $\lambda_x$ et $\lambda_y$ et observer l'effet sur le spectre de magnitude de la TF.
#
# > Ajoutez une option pour binariser l'image générée.
#
#
# 3. **Créer une méthode interactive permettant:**
# * De sélectionner une image
# * De lui appliquer du bruit Gaussien ou Sel&Poivre
# * De la débruiter / lisser en choisissant un filtre (moyen, médian, Gaussien) ainsi que la taille du dit filtre.
# * Affiche le résultat sous la forme d'une grille 2x3:
#
# | | | |
# |:--------:|:-------------:|:------:|
# | image | image_bruitée | image_débruitée |
# | fft_image | fft_image_bruitée | fft_image_débruitée |
#
# > Observer les différences de qualité de débruitage des différents filtres selon le type de bruit.
#
#
# 4. **Ajouter la possibilité de générer des masques binaires circulaires (dont le rayon est réglable de manière interactive), qui auront un centre blanc et l'extérieur noir.**
#
# > Appliquer le masque à la représentation fréquentielle de votre image et observer le résultat sur l'image recomposée.
#
#
# 5. **Modifiez votre code de sorte à ce que les filtres moyens et Gaussiens soient appliqués "manuellement": créez le kernel et appliquez-le via la méthode `fftconvolve` de `Scipy`.**
#
#
# 6. **Modifiez votre code de sorte à pouvoir appliquer le filtre à une image couleur, en implémentant 3 méthodes:**
# 1. L'applique à chacun des canaux de l'image et combine le résultat
# 2. L'applique au canal de luminance d'un espace colorimétrique approprié
# 3. Applique un filtre 3D (`n*n*3`)
#
# > Afficher les 3 résultats côte-à-côte et comparer.
#
#
# 7. **Proposez une solution pour extraire les contours / hautes-fréquences d'une image en utilisant uniquement un filtre Gaussien.**
#
#
# 8. **Etendez le principe de la question 7. en l'appliquant à la différence de deux Gaussiennes, i.e. une même image lissée selon deux kernels Gaussiens (avec la première image moins floutée que la seconde).**
#
# > Vous venez d'appliquer une Différence de Gaussiennes (*DoG*), qui est une technique d'extraction de contours.
#
#
# 8. **[<u>Bonus</u>] Proposez une méthode basique pour quantifier la différence de qualité entre l'image et sa version débruitée, et se servir de cette méthode pour comparer l'efficacité des différents filtres selon le type de bruit appliqué.**
#
# </div>
# + colab={} colab_type="code" id="TbG86DcIookt"
# > Emplacement exercice <
# + [markdown] colab_type="text" id="7EMgeXj_ooky"
# ## <span style="color: Green;text-decoration: underline" id="3">III. Accentuation (*sharpening*) et Filtres passe-haut</span>
# + [markdown] colab_type="text" id="vUeHpXg8ookz" inputHidden=false outputHidden=false
# Un filtre de **sharpening** est un **filtre passe-haut**, car il laisse passer les hautes-fréquences et élimine les basses-fréquences. Ces filtres réagiront au maximum là ou les gradients (de luminance) sont maximums (et concordants avec le filtre). Ils vont donc rendre l'image plus nette, en faire ressortir les contrastes, les gradients, les bordures.
#
# Il existe différents types de filtres passe-haut, qui vont faire ressortir différents types de détails de l'image:
# * Filtre de Prewitt
# * Filtre de Sobel
# * Filtre Laplacien (LoG)
# * Filtre de Canny
#
# Dans le **domaine fréquentiel**, un filtre passe-haut est tout simplement un masque (en niveaux de gris) où les valeurs sont plus élevées loin du centre, afin de laisser passer les hautes fréquences (qui sont éloignées du centre) préférentiellement.
# -
# ***
# Les hautes fréquences d'une image correspondent aux changements rapides de luminance. Ces changements rapides peuvent être détectés en analysant le **gradient de luminance de l'image**: le gradient de l'image, noté $\nabla I$ ("nabla de I"), caractérise la vitesse de variation d'une quantité (ici la luminance) par rapport aux variations d'une ou plusieurs autres quantités.
#
#
# Pour une image, $\nabla I(x,y)$ est calculé selon les variations conjointes de $x$ et $y$, et peut s'exprimer par les dérivés partielles de la luminance selon $x$ et $y$:
#
# $$
# \nabla I(x,y) = \left( \frac{\partial I(x,y)}{\partial x},\frac{\partial I(x,y)}{\partial y} \right)
# $$
#
# La dérivée partielle par rapport à x (ou y) permet d'étudier les variations d'intensités de l'image dans la direction de l'axe des abscisses (ou des ordonnées).
#
# Le gradient étant un vecteur, il possède une norme (ou amplitude) $||\nabla I||$ et une direction donnée par l'angle $\theta$ :
#
# \begin{align*}
# ||\nabla I(x,y)||_2 &= \sqrt{\left( \frac{\partial I (x,y) }{ \partial x}\right)^2 + \left( \frac{\partial I (x,y) }{ \partial y}\right)^2} \\
# \Theta &= \arctan \left( \frac{ \frac{\partial I(x,y)}{\partial y } }{ \frac{\partial I(x,y)}{\partial x}} \right)
# \end{align*}
#
# Le gradient pointe dans la direction $\theta$ vers laquelle l'intensité varie le plus dans le voisinage du pixel $(x,y)$. L'amplitude $||\nabla I(x,y)||$ indique à quel point cette variation est importante : plus elle est élevée, plus le changement est brusque. Mathématiquement, détecter les hautes fréquences (comme les bords) revient donc à trouver les maxima locaux de $||\nabla I||$.
#
# ***
# Dans une image numérique, la fonction d'intensité n'est pas continue, mais est définie en un nombre fini de pixels. Le calcul des dérivées partielles ne peut donc être qu'approximé par la méthode des différences finies: le gradient est approximé par les variations d'un pixel au suivant (selon l'axe des $x$ ou $y$):
#
# $$
# \frac{\partial I (x,y)}{\partial x} ≈ I(x+1, y) − I(x,y)
# \\
# \frac{\partial I (x,y)}{\partial y} ≈ I(x, y+1) − I(x,y)
# $$
#
# Cette approximation peut être reformulée en calcul matriciel par:
#
# $$
# \frac{\partial I (x,y) }{\partial x} \approx [0\ -1\ 1] * I(x,y)
# \\
# \frac{\partial I (x,y) }{\partial y} \approx [0\ -1\ 1]^T * I(x,y)
# $$
#
# Avec $[−1\ 0\ 1]$ et $[−1\ 0\ 1]^T$ les noyaux de la convolution appliqués à l'image $I$ pour approximer ses gradients de luminance selon $x$ et $y$.
#
# <u>Remarque</u>: Ces noyaux ne s'utilisent jamais en pratique puisqu'ils ne sont pas centrés: l'on préfère $[−1\ 0\ 1]$ pour approximer les variations selon $x$, et $[−1\ 0\ 1]^T$ pour les variations selon $y$. Ces nouveaux masques exploitent les informations provenant de part et d'autre du pixel, et non d'un seul côté.
#
# <u>Remarque</u>: Etendus en deux dimensions, ces noyaux vont devenir les noyaux du filtre de Prewitt, l'un des opérateur les plus basiques pour la détection de contours (cf. section III.1).
# + colab={} colab_type="code" id="zny7Hhn1ook0"
### Imports et fonctions utiles à cette partie
import os, cv2
import numpy as np
from PIL import Image, ImageOps
from scipy.signal import convolve as scipy_convolve, convolve2d, fftconvolve
from scipy.ndimage import sobel
# Interactivité
from IPython.display import display, Markdown
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import plotly.express as px # Librairie de visualisation avancée (similaire à Seaborn)
import plotly.graph_objects as go
from matplotlib.pylab import *
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits import mplot3d
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina' # Améliorer la netteté des figures
plt.rcParams["figure.figsize"] = 12.8, 9.6
import warnings # "Do not disturb" mode
warnings.filterwarnings('ignore')
def get_power_spectrum(img):
return np.log(np.abs(np.fft.fftshift(np.fft.fft2(np.array(img, dtype="uint8")))))
def affichage_1x2(src, dst):
fig = plt.figure(figsize=(12, 6))
plt.subplot(121), plt.imshow(src, cmap="gray", origin="upper"), plt.title("Image d'origine")
plt.subplot(122), plt.imshow(dst, cmap="gray", origin="upper"), plt.title("Image filtrée")
plt.show()
def affichage_2x2_Fourier(img, filtered, titles=["Original", "Filtered", "[Fourier] Original", "[Fourier] Filtered"]):
fig = plt.figure(figsize=(12, 12))
magnitude_img = get_power_spectrum(img)
magnitude_filtered = get_power_spectrum(filtered)
plt.subplot(221), plt.imshow(img, cmap="gray", origin="upper"), plt.title(titles[0])
plt.subplot(222), plt.imshow(filtered, cmap="gray", origin="upper"), plt.title(titles[1])
plt.subplot(223), plt.imshow(magnitude_img, cmap="hot", extent=(-magnitude_img.shape[0]//2, magnitude_img.shape[0]//2, -magnitude_img.shape[1]//2, magnitude_img.shape[1]//2)), plt.title(titles[2])
plt.subplot(224), plt.imshow(magnitude_filtered, cmap="hot", extent=(-magnitude_filtered.shape[0]//2, magnitude_filtered.shape[0]//2, -magnitude_filtered.shape[1]//2, magnitude_filtered.shape[1]//2)), plt.title(titles[3])
plt.show()
def affichage_2x3_Fourier(img, filtered1, filtered2, titles=["Original", "Filtered", "Filtered2", "[Fourier] Original", "[Fourier] Filtered1", "[Fourier] Filtered2"]):
fig = plt.figure(figsize=(18, 12))
magnitude_img = get_power_spectrum(img)
magnitude_filtered1 = get_power_spectrum(filtered1)
magnitude_filtered2 = get_power_spectrum(filtered2)
plt.subplot(231), plt.imshow(img, cmap="gray", origin="upper"), plt.title(titles[0])
plt.subplot(232), plt.imshow(filtered1, cmap="gray", origin="upper"), plt.title(titles[1])
plt.subplot(233), plt.imshow(filtered2, cmap="gray", origin="upper"), plt.title(titles[2])
plt.subplot(234), plt.imshow(magnitude_img, cmap="hot", extent=(-magnitude_img.shape[0]//2, magnitude_img.shape[0]//2, -magnitude_img.shape[1]//2, magnitude_img.shape[1]//2)), plt.title(titles[3])
plt.subplot(235), plt.imshow(magnitude_filtered1, cmap="hot", extent=(-magnitude_filtered1.shape[0]//2, magnitude_filtered1.shape[0]//2, -magnitude_filtered1.shape[1]//2, magnitude_filtered1.shape[1]//2)), plt.title(titles[4])
plt.subplot(236), plt.imshow(magnitude_filtered2, cmap="hot", extent=(-magnitude_filtered2.shape[0]//2, magnitude_filtered2.shape[0]//2, -magnitude_filtered2.shape[1]//2, magnitude_filtered2.shape[1]//2)), plt.title(titles[5])
plt.show()
def kernel_heatmap(kernel, cm="gray"):
fig = px.imshow(kernel, color_continuous_scale=cm)
fig.show()
def kernel_3D(kernel):
z_data = pd.DataFrame(kernel)
fig = go.Figure(data=[go.Surface(z=z_data.values)])
fig.update_layout(title='Kernel', autosize=True,
width=500, height=500,
margin=dict(l=65, r=50, b=65, t=90))
fig.show()
def kernel_3D_continuous(data, color_map="viridis"):
x,y,z = data
X1 = np.reshape(x, -1)
Y1 = np.reshape(y, -1)
Z1 = np.reshape(z, -1)
# Normalize the colors based on Z value
norm = plt.Normalize(z.min(), z.max())
colors = cm.jet(norm(z))
ax = plt.axes(projection='3d')
ax.get_proj = lambda: np.dot(mplot3d.Axes3D.get_proj(ax), np.diag([1, 1, 1, 1]))
ax.plot_trisurf(X1, Y1, Z1, cmap=color_map)
# + [markdown] colab_type="text" id="YS2y5o_HoolK"
# ## <span style="color: DodgerBlue;text-decoration: underline">III.1 Filtre de Prewitt</span>
# + [markdown] colab_type="text" id="Y49TAzSfoolL"
# Le **filtre de Prewitt** est un **opérateur différentiel du premier ordre** (car basé sur le calcul de la dérivée première de la luminance) permettant de **détecter des bords orientés** (verticaux ou horizontaux). Il est basé sur deux noyaux, l'un pour l'axe horizontal ($G_x$) et l'autre pour l'axe vertical ($G_y$). Il permet de calculer une **approximation du gradient** (horizontal ou vertical) de la luminance de l'image.
#
# <img src="https://homepages.inf.ed.ac.uk/rbf/HIPR2/figs/prwgrad.gif">
# -
@interact
def prewitt_manual(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))]):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# Noyau de convolution
prewitt_x_kernel = np.array([[1, 1, 1],[0, 0, 0],[-1, -1, -1]])
prewitt_y_kernel = np.array([[-1, 0, 1],[-1, 0, 1],[-1, 0, 1]])
# Application du filtre avec la fonction fftconvolve de Scipy.signal
prewitt_x = fftconvolve(img, prewitt_x_kernel, mode="same")
prewitt_y = fftconvolve(img, prewitt_y_kernel, mode="same")
# On combine les deux sous-filtres
prewitt = np.sqrt(prewitt_x**2 + prewitt_y**2)
titles=["Orignal", "Prewitt", "[Fourier] Original", "[Fourier] Prewitt"]
affichage_2x2_Fourier(img, prewitt, titles)
# + [markdown] colab_type="text" id="N3mIiUfsoolE" inputHidden=false outputHidden=false
# ## <span style="color: DodgerBlue;text-decoration: underline">III.2 Filtres de Sobel & Scharr</span>
# + [markdown] colab_type="text" id="HuRaFbacoolF"
# Le **filtre de Sobel** est un opérateur différentiel discret du premier ordre, permettant de détecter des bords orientées, horizontaux ($G_x$) ou verticaux ($G_y$) (ou une combinaison des deux). Il permet de calculer une **approximation du gradient** (horizontal ou vertical) de la luminance de l'image.
#
# 
#
# L'opérateur de Sobel à plus tard été optimisé par `Scharr`, aboutissant à **l'opérateur de Scharr** dont deux discrétisations connues sont :
#
# 
#
#
# 
# -
# #### Avec `cv2.sobel` :
# + colab={"referenced_widgets": ["0f4b8eccd1c147c6929a3b69f5c5eac6"]} colab_type="code" id="A1fzLg_koolG" outputId="96eb17dc-bd3c-4c74-826b-142d5fc99c4e"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact
def sobel(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], kernel_size=kernel_size_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# Sobel sur l'axe des X
sobel_horizontal = cv2.Sobel(img, cv2.CV_64F, dx=1, dy=0, ksize=kernel_size)
sobel_horizontal = np.uint8(np.absolute(sobel_horizontal))
# Sobel sur l'axe des Y
sobel_vertical = cv2.Sobel(img, cv2.CV_64F, dx=0, dy=1, ksize=kernel_size)
sobel_vertical = np.uint8(np.absolute(sobel_vertical))
titles=["Orignal", "Sobel X", "Sobel Y", "[Fourier] Original", "[Fourier] Sobel X", "[Fourier] Sobel Y"]
affichage_2x3_Fourier(img, sobel_horizontal, sobel_vertical, titles)
display(Markdown("Le filtre de Sobel Horizontal détecte les variations (gradients) d'intensité selon l'axe des X, ce qui correspond aux lignes verticales !"))
# -
# #### Application de l'opérateur de `Sobel` manuellement:
@interact
def sobel_manual(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))]):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# Noyau de convolution
sobel_x_kernel = np.array([[1, 0, -1],[2, 0, -2],[1, 0, -1]])
sobel_y_kernel = np.array([[1, 2, 1],[0, 0, 0],[-1, -2, -1]])
# Application du filtre avec la fonction convolve de Scipy.signal (en mode fft)
sobel_x = scipy_convolve(img, sobel_x_kernel, mode="same", method="fft")
sobel_y = scipy_convolve(img, sobel_y_kernel, mode="same", method="fft")
# On combine les deux sous-filtres
sobel_x = np.uint8(np.absolute(sobel_x))
sobel_y = np.uint8(np.absolute(sobel_y))
sobel = 0.5 * sobel_x + 0.5 * sobel_y
titles=["Sobel X", "Sobel Y", "Sobel", "[Fourier] Sobel X", "[Fourier] Sobel Y", "[Fourier] Sobel"]
affichage_2x3_Fourier(sobel_x, sobel_y, sobel, titles)
# + [markdown] colab_type="text" id="dhRoL0W4oolO"
# ## <span style="color: DodgerBlue;text-decoration: underline">III.3 Filtre Laplacien</span>
# + [markdown] colab_type="text" id="aTeDA4iWoolP"
# Le **filtre Laplacien** (ou opérateur de Laplace) est un **opérateur différentiel du second ordre** fréquemment utilisé pour **réhausser les contours des images**. Cet opérateur correspond à la dérivée seconde d'une fonction 2D, généralement une Gaussienne (on parle alors de Laplacien de Gaussienne - **_LoG_**).
#
# Les filtres Laplaciens (comme les autres filtres dérivatifs) sont utilisés pour faire ressortir les zones de changements rapides, i.e. les hautes fréquences de l'image.
#
# <u>Illustration</u>: visualisation en 3D d'une LoG
# +
# Tracé d'une Laplacienne de Gaussienne (LoG) continue
def LoG(n, sigma=8):
half_n = n // 2
x, y = np.meshgrid(range(n), range(n))
x = x - half_n
y = y - half_n
temp = (x ** 2 + y ** 2) / (2 * sigma ** 2)
return x, y, -1 / (np.pi * sigma ** 4) * (1 - temp) * np.exp(-temp)
kernel_3D_continuous(LoG(50), color_map="viridis_r")
# -
# Cette fonction peut être discrétisée de plusieurs manières, aboutissant à différentes variantes du LoG discret (pour une même taille de noyau) :
#
# <img src="http://www.rroij.com/articles-images/IJAREEIE-2107-e004.gif">
#
# ```python
# h1 = np.array([[0.0, 1.0, 0.0],[1.0, -4.0, 1.0],[0.0, 1.0, 0.0],])
# h2 = np.array([[1.0, 1.0, 1.0],[1.0, -8.0, 1.0],[1.0, 1.0, 1.0],])
# h3 = np.array([[-1.0, 2.0, -1.0],[2.0, -4.0, 2.0],[-1.0, 2.0, -1.0],])
# ```
#
# Comme pour beaucoup d'autres filtres, il en existe également des discrétisation de plus grandes taille (5x5, 7x7, ...).
#
# *Remarque:* Ce filtre est tres sensible au bruit, du coup il est souvent précédé par un lissage Gaussien.
# #### Avec `cv2.Laplacian`:
# + colab={"referenced_widgets": ["c1501e1d9a8140d5a45691751ff07822"]} colab_type="code" id="hpD0-6XDoolQ" outputId="5f185f7f-6338-4e1d-93d3-714f43652542"
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
@interact_manual
def LoG(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], blurr_kernel_size=(3,9,2), kernel_size=kernel_size_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# On applique le filtre LoG
laplacian = cv2.Laplacian(img, cv2.CV_64F, ksize=kernel_size)
laplacian = np.uint8(np.absolute(laplacian))
# On applique le filtre LoG après lissage Gaussien
blurred = cv2.GaussianBlur(img, (blurr_kernel_size, blurr_kernel_size), 0)
laplacian_blurred = cv2.Laplacian(blurred, cv2.CV_64F, ksize=kernel_size)
laplacian_blurred = np.uint8(np.absolute(laplacian_blurred))
titles=["Orignal", "Laplacian", "Smoothed Laplacian", "[Fourier] Original", "[Fourier] Laplacian", "[Fourier] Smoothed Laplacian"]
affichage_2x3_Fourier(img, laplacian, laplacian_blurred, titles)
# + [markdown] colab_type="text" id="XziqWeSBoolV"
# ## <span style="color: DodgerBlue;text-decoration: underline">III.4 Filtre de Canny</span>
# + [markdown] colab_type="text" id="Z0mFIpiXoolW"
# Le **filtre de Canny** (*Canny Edge Detector*) est un filtre d'extraction de contours multi-étapes permettant d'extraire un grand nombre de contours de tailles et orientations différentes.
#
# La méthode d'OpenCV qui permet de réaliser un filtre de Canny est :
# ```python
# cv2.Canny(image, threshold_lower, threshold_upper)
# ```
#
# Elle se basent sur deux seuils (`threshold_lower` et `threshold_upper`) qui affectent la sensibilité de l'opétation d'extraction de contours.
# + colab={"referenced_widgets": ["4716da00ec9a49bb842405eb6c7e038e"]} colab_type="code" id="CWOMIkoboolY" outputId="7d8c8703-6502-4e4d-9902-f4b18d9f9706"
# Méthode permettant de définir les seuils de la méthode cv2.Canny automatiquement
def auto_canny(image, sigma=0.33):
# compute the median of the single channel pixel intensities
v = np.median(image)
# apply automatic Canny edge detection using the computed median
lower = int(max(0, (1.0 - sigma) * v))
upper = int(min(255, (1.0 + sigma) * v))
edged = cv2.Canny(image, lower, upper)
return edged, lower, upper
#######
kernel_size_slider = widgets.IntSlider(min=3, max=7, step=2, value=3)
thresh_low_slider = widgets.IntSlider(min=25, max=250, step=25, value=100)
thresh_up_slider = widgets.IntSlider(min=25, max=250, step=25, value=200)
@interact_manual
def canny(image=[f for f in os.listdir(img_path) if os.path.isfile(os.path.join(img_path, f))], thresh_low=thresh_low_slider, thresh_up=thresh_up_slider):
img = np.array(Image.open(img_path + image).convert("L")).astype("uint8")
# On applique le filtre de Canny après filtrage (smoothing) Bilateral
img_bilateral = cv2.bilateralFilter(img, 7, 50, 50)
canny_bilateral_manuel = cv2.Canny(img_bilateral, thresh_low, thresh_up)
canny_bilateral_auto,_,_ = auto_canny(img_bilateral)
titles=["Orignal", "Canny", "Canny auto", "[Fourier] Original", "[Fourier] Canny", "[Fourier] Canny auto"]
affichage_2x3_Fourier(img, canny_bilateral_manuel, canny_bilateral_auto, titles)
# + [markdown] colab_type="text" id="iL7f4Rjloolc"
# ### <span style="color:crimson">**[<u>Exercice</u>]** A vous de jouer:</span>
# ***
# <div style="color:DarkSlateBlue">
#
# 1. **A partir de votre code de l'exercice de la partie II :**
# * Ajouter le filtre Bilateral à vos options de débruitage / lissage.
# * Ajouter la possibilité de choisir entre deux types de masques binaires pour le filtrage fréquentiel:
# * _On_ : centre blanc, extérieur noir
# * _Off_ : centre noir, extérieur blanc
#
# > Appliquer le nouveau type de masque (*Off*) à la représentation fréquentielle de votre image et observer le résultat sur l'image recomposée.
#
# 2. **Modifiez votre code de sorte à pouvoir faire varier la valeur de deux rayons $R1$ et $R2$ de manière interactive :**
# * Avec $R1 < R2$
# * $R1$ utilisé pour créer un masque Off
# * $R2$ utilisé pour créer un masque On
# * Combinez les deux masques avec l'opérateur `bitwise_and`
#
# > Appliquez le masque résultant aux images, faites varier l'écart entre $R1$ et $R2$, et observez.
#
# > <u>Remarque</u>: Le filtre que vous venez d'appliquer est un **filtre passe-bande**, qui ne laisse passer qu'une certaine bande de fréquences, bande ici définie par la différence entre $R1$ et $R2$.
#
#
# 3. **Implémentez la technique _d'Unsharp Masking_ qui consiste à ajouter à une image un masque généré en soustrayant une version floutée de l'image à elle même (pour en faire ressortir les contours). Cette technique permet de mettre les contours de l'image en surbrillance.**
#
# Ce principe suit l'équation: $sharpened = original + \lambda*(original − blurred)$.
#
# Avec $\lambda$ le coefficient d'amplification du masque.
#
# > Faites varier $\lambda$ ($<1$ ou $>1$) et observer.
#
#
# 4. **Afficher côte-à-côte (dans une grille 2x2) et comparer les contours extraits par DoG (*cf. exercice II*), LoG, Sobel (en combinant $S_x$ et $S_y$ selon la formule: $||S|| = \sqrt{S_x^2 + S_y^2}$), et Canny (auto).**
#
# </div>
# + colab={} colab_type="code" id="DXIpePJZoold"
# > Emplacement exercice <
# + [markdown] colab_type="text" id="D7NLN3g5oolh"
# <div style="color:Navy">
#
# ***
# # Fin du TP4
# ***
#
# </div>
| TP4/IP - TP4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _nb_constraint_handling:
# -
# # Constraint Handling
# Constraint handling is often neglected in frameworks but is indeed an essential aspect of optimization.
# Depending on the characteristics of the constraints, one method might be more suitable and lead to better results than another.
# In principle, two ways exist how constraints can be dealt with: a) use an algorithm that can handle constraints internally; b) redefine the problem to have no constraints but still guarantee the optimum to be a feasible solution.
# Both strategies will be elaborated in the following.
# By default, the returned optimum is **always** required to be feasible. It might be required to obtain the **least infeasible** solution if no feasible could have been found in some situations. To enable this behavior, you can pass `return_least_infeasible=True` to the `minimize` function.
# +
from pymoo.algorithms.soo.nonconvex.ga import GA
from pymoo.factory import get_problem
from pymoo.optimize import minimize
res = minimize(get_problem("g05"),
GA(),
return_least_infeasible=True,
seed=1,
verbose=False)
print("CV:", res.CV[0])
print("F:", res.F[0])
# -
# ## Inequality Constraints
# In pymoo, inequality constraints are always defined as "$\leq 0$" constraints. Thus, constraint violation is defined as follows: A solution is considered as **feasible** of all constraint violations are **less than** zero. A solution is considered as **infeasible** if at least one constraint violation is **larger than** zero.
# $$
# - x^4 \geq - 2
# $$
# Has to be converted to a "$\leq 0$" constraints to satisfy the implementation of constraints in pymoo:
# $$
# x^4 \leq 2\\
# x^4 - 2 \leq 0
# $$
# Let us assume the whole optimization problem is then given by:
# \begin{align}
# \begin{split}
# \min \;\; & f(x) = 1 + x^3 \\
# \text{s.t.} \;\; & x^4 - 2\leq 0\\
# & -2 \leq x \leq 2
# \end{split}
# \end{align}
# The figure below shows the corresponding function and the optimum to be found. The constraint function is shown with a dashed line greater than zero (violated) and with a solid line where less or equal to zero (satisfied). The optimum is shown by a red cross, which is minimum in the feasible region.
# + nbsphinx="hide_input"
import numpy as np
import matplotlib.pyplot as plt
plt.rc('font', family='serif')
X = np.linspace(-2, 2, 10000)
F = 1 + X ** 3
G = X**4 - 2
plt.figure(figsize=(7, 5))
plt.plot(X, F, linewidth=2.0, color="blue", label="$f(x) = 1 + x^3$")
I = G<=0
plt.plot(X[I], G[I], linewidth=4.0, color="green")
plt.plot(X, G, linewidth=2.0, color="green", label="$g(x) = x^4 - 2$", linestyle='dotted')
plt.scatter([-1.18920654], [-0.6817904], color="red", s=100, marker="X", label="Optimum")
plt.xlim(-2, 2)
plt.xlabel("$x$")
plt.ylabel("$f$")
plt.legend(loc='upper center', bbox_to_anchor=(0.5, 1.12),
ncol=4, fancybox=True, shadow=False)
plt.tight_layout()
plt.show()
# -
# The problem can be implemented straightforwardly:
# +
import numpy as np
from pymoo.util.misc import stack
from pymoo.core.problem import Problem
class MyProblem(Problem):
def __init__(self):
super().__init__(n_var=1,
n_obj=1,
n_constr=1,
xl=-2,
xu=2)
def _evaluate(self, X, out, *args, **kwargs):
out["F"] = 1 + X ** 3
out["G"] = X**4 - 2
# -
# And then being solved as usually by initiating the problem, algorithm and running the minimize method:
# +
from pymoo.algorithms.soo.nonconvex.pso import PSO
from pymoo.optimize import minimize
problem = MyProblem()
algorithm = PSO()
res = minimize(problem,
algorithm,
seed=1,
verbose=False)
print("Best solution found: \nX = %s\nF = %s\nCV = %s" % (res.X, res.F, res.CV))
# -
# For a bi-objective example with constraints, we like to refer to our [Getting Started Guide](../getting_started/index.ipynb) where a similar example is shown.
# ## Equality to Inequality Constraints
# We got a couple of questions about how equality constraints should be handled in a genetic algorithm. In general, functions without any smoothness are challenging to handle for genetic algorithms. An equality constraint is an extreme case, where the constraint violation is 0 at precisely one point and otherwise 1.
#
# Let us consider the following constraint $g(x)$ where $x$ represents a variable:
# $g(x): x = 5$
# An equality constraint can be expressed by an inequality constraint:
# $g(x): |x - 5| \leq 0$
# or
# $g(x): (x-5)^2 \leq 0$
# However, all of the constraints above are very strict and make **most search space infeasible**. Without providing more information to the algorithm, those constraints are challenging to satisfy.
# Thus, the constraint can be smoothed by adding an epsilon to it and, therefore, having two inequality constraints:
# $g'(x): 5 - \epsilon \leq x \leq 5 + \epsilon$
# Also, it can be simply expressed in one inequality constraint by:
# $g'(x): (x-5)^2 - \hat{\epsilon} \leq 0$
# Depending on the $\epsilon$, the solutions will be more or less close to the desired value. However, the genetic algorithm does not know anything about the problem itself, making it challenging to handle and focus the search in the infeasible space.
# ## Parameter-less Approach / Feasibility First
# Parameter-less constraint handling is the default constraint handling approach in **pymoo**. It strictly prefers feasible over infeasible solutions.
# $$
# F(x)=
# \begin{cases}
# f(x), \quad \text{if feasible} \\
# f_{\max} + \sum_{j=1}^{|J|} \, \langle \, g_j (x) \, \rangle, \quad \rm{otherwise}
# \end{cases}
# $$
# where $\langle \cdot \rangle$ is defined by
# $$
# \langle a \rangle =
# \begin{cases}
# a, \quad a \geq 0 \\
# 0, \quad \rm{otherwise}
# \end{cases}
# $$
# This approach works well if an algorithm is based on comparisons. For instance, a genetic algorithm has to determine what individuals survive. Thus, this mapping can be utilized to create a rank for each individual.
# If the $f_\max$ of the function is known beforehand, it can also be used to redefine the problem beforehand.
# Redefining the problem is very similar to the approach discussed next.
# ## Penalty-based Approach / Lagrange Multipliers
# The penalty-based constraint handling approach internally converts equality constraints to inequality constraints. Then, the violation of inequality and equality constraints penalize the objective value. This makes all non-constrained algorithms applicable; however, it also might create a more difficult fitness landscape to solve.
#
# **NOTE**: Genetic algorithms are not made to handle equality constraints out of the box, but often need to be customized to satisfy the equality constraints directly. The approach below only works for a very simple and a very limited amount of equality constraints.
# ### Single-objective
# \begin{align}
# \begin{split}
# \min \;\; & f(x) = x_1^2 + x_2^2 + x_3^2 \\
# \text{s.t.} \;\; & x_1 \cdot x_2 \geq 1\\
# & x_1 \cdot x_2 \leq 5\\
# & x_2 + x_3 = 1\\
# & 0 \leq x \leq 5
# \end{split}
# \end{align}
# By converting the constraints to $\leq 0$ constraints the implementation of the problem looks as follows:
# +
def func_obj(x):
return x[0] ** 2 + x[1] ** 2 + x[2] ** 2
constr_ieq = [
lambda x: 1 - x[0] * x[1],
lambda x: x[0] * x[1] - 5
]
constr_eq = [
lambda x: 1 - x[1] - x[2]
]
# -
# For this example, we use the definition of a problem using the factory method `create_problem_from_funcs`:
# +
from pymoo.problems.functional import FunctionalProblem
from pymoo.problems.constr_as_penalty import ConstraintsAsPenalty
problem = FunctionalProblem(3,
func_obj,
xl=0,
xu=5,
constr_ieq=constr_ieq,
constr_eq=constr_eq
)
problem = ConstraintsAsPenalty(problem, penalty=1e6)
# -
# Then, an algorithm can be used to solve the problem as usual:
# +
from pymoo.algorithms.soo.nonconvex.pso import PSO
from pymoo.optimize import minimize
algorithm = PSO(pop_size=50)
res = minimize(problem, algorithm, seed=1)
opt = res.opt[0]
X, F, CV = opt.get("X", "__F__", "__CV__")
print("Best solution found: \nX = %s\nF = %s\nCV = %s\n" % (X, F, CV))
# -
# ### Multi-objective
# The almost equivalent could be applied for multi-objective optimization problems. The example below shows the [getting started guide](../getting_started/index.ipynb) example with a penalty being used to deal with the constraints:
# #### Functions
# +
objs = [
lambda x: x[0]**2 + x[1]**2,
lambda x: (x[0]-1)**2 + x[1]**2
]
constr_ieq = [
lambda x: 2*(x[0]-0.1) * (x[0]-0.9) / 0.18,
lambda x: - 20*(x[0]-0.4) * (x[0]-0.6) / 4.8
]
constr_eq = []
# -
# #### Problem
# +
from pymoo.problems.functional import FunctionalProblem
from pymoo.problems.constr_as_penalty import ConstraintsAsPenalty
problem = FunctionalProblem(2,
objs,
xl=-2,
xu=2,
constr_ieq=constr_ieq,
constr_eq=constr_eq
)
problem = ConstraintsAsPenalty(problem, penalty=1e6)
# -
# #### Optimize
# +
from pymoo.algorithms.moo.nsga2 import NSGA2
from pymoo.optimize import minimize
algorithm = NSGA2(pop_size=40)
res = minimize(problem,
algorithm,
seed=1)
# -
# #### Visualize
# +
from pymoo.visualization.scatter import Scatter
plot = Scatter(title = "Objective Space")
plot.add(res.F)
plot.show()
# -
# ## Repair Operator
# A simple approach is to handle constraints through a repair function. This is only possible if the equation of the constraint is known. The repair makes sure every solution that is evaluated is, in fact, feasible. Let us consider the following example where
# the equality constraints need to consider more than one variable:
# \begin{align}
# \begin{split}
# \min \;\; & f_1(x) = (x_1^2 + x_2^2) \\
# \max \;\; & f_2(x) = -(x_1-1)^2 - x_2^2 \\[1mm]
# \text{s.t.} \;\; & g_1(x_1, x_3) : x_1 + x_3 = 2\\[2mm]
# & -2 \leq x_1 \leq 2 \\
# & -2 \leq x_2 \leq 2 \\
# & -2 \leq x_3 \leq 2
# \end{split}
# \end{align}
# We implement the problem using by squaring the term and using an $\epsilon$ as we have explained above. The source code for the problem looks as follows:
# +
import numpy as np
from pymoo.core.problem import Problem
class MyProblem(Problem):
def __init__(self):
super().__init__(n_var=3,
n_obj=2,
n_constr=1,
xl=np.array([-2, -2, -2]),
xu=np.array([2, 2, 2]))
def _evaluate(self, x, out, *args, **kwargs):
f1 = x[:, 0] ** 2 + x[:, 1] ** 2
f2 = (x[:, 0] - 1) ** 2 + x[:, 1] ** 2
g1 = (x[:, 0] + x[:, 2] - 2) ** 2 - 1e-5
out["F"] = np.column_stack([f1, f2])
out["G"] = g1
# -
# As you might have noticed, the problem has similar characteristics to the problem in our getting started.
# Before a solution is evaluated, a repair function is called. To make sure a solution is feasible, an approach would be to either set $x_3 = 2 - x_1$ or $x_1 = 2 - x_3$. Additionally, we need to consider that this repair might produce a variable to be out of bounds.
# +
from pymoo.core.repair import Repair
class MyRepair(Repair):
def _do(self, problem, pop, **kwargs):
for k in range(len(pop)):
x = pop[k].X
if np.random.random() < 0.5:
x[2] = 2 - x[0]
if x[2] > 2:
val = x[2] - 2
x[0] += val
x[2] -= val
else:
x[0] = 2 - x[2]
if x[0] > 2:
val = x[0] - 2
x[2] += val
x[0] -= val
return pop
# -
# Now the algorithm object needs to be initialized with the repair operator and then can be run to solve the problem:
# +
from pymoo.algorithms.moo.nsga2 import NSGA2
from pymoo.optimize import minimize
from pymoo.visualization.scatter import Scatter
algorithm = NSGA2(pop_size=100, repair=MyRepair(), eliminate_duplicates=True)
res = minimize(MyProblem(),
algorithm,
('n_gen', 20),
seed=1,
verbose=True)
plot = Scatter()
plot.add(res.F, color="red")
plot.show()
# -
# In our case it is easy to verify if the constraint is violated or not:
print(res.X[:, 0] + res.X[:, 2])
# If you would like to compare the solution without a repair you will see how searching only in the feasible space helps:
# +
algorithm = NSGA2(pop_size=100, eliminate_duplicates=True)
res = minimize(MyProblem(),
algorithm,
('n_gen', 20),
seed=1,
verbose=True)
plot = Scatter()
plot.add(res.F, color="red")
plot.show()
# -
print(res.X[:, 0] + res.X[:, 2])
# Here, in fact, the $\epsilon$ term is necessary to find any feasible solution at all.
| source/misc/constraints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="G5HpRApza9UR"
#Importando as bibliotecas
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use("seaborn")
# -
df = pd.read_excel("/home/wesley/GitHub/Pandas_dio/datasets/AdventureWorks.xlsx")
# + colab={"base_uri": "https://localhost:8080/", "height": 391} colab_type="code" id="DPOEg0MikIXG" outputId="633ff3c2-2339-4b86-8110-b65b6d29e2d2"
#Visualizando as 5 primeiras linhas
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="UCJpu--kK9wo" outputId="8d55d823-0742-4a54-fcb2-16df652007da"
#Quantidade de linhas e colunas
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="P9S1i8o1lUu-" outputId="6197dad1-cbfe-444a-a212-1bfa12541ebd"
#Verificando os tipos de dados
df.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="duheNX1GlhWw" outputId="0b047796-3ab7-4dd4-a370-ecce8e256679"
#Qual a Receita total?
df["Valor Venda"].sum()
# + colab={} colab_type="code" id="IHop-35BlyDO"
#Qual o custo Total?
df["custo"] = df["Custo Unitário"].mul(df["Quantidade"]) #Criando a coluna de custo
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="3fy4QmNLmMWd" outputId="b08296d5-566d-4123-a8d7-b2c4d959e935"
df.head(1)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Uj7LTfyumqcn" outputId="ed0c7864-fc3c-40e2-e2e1-04182d758866"
#Qual o custo Total?
round(df["custo"].sum(), 2)
# + colab={} colab_type="code" id="dcL7yq6dm6-R"
#Agora que temos a receita e custo e o total, podemos achar o Lucro total
#Vamos criar uma coluna de Lucro que será Receita - Custo
df["lucro"] = df["Valor Venda"] - df["custo"]
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="AESBzwFuqgy4" outputId="a832d8aa-bbee-41c6-e823-5844dd61890c"
df.head(1)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="odfh78ayqpN4" outputId="8e29504f-0eb5-4bc7-8312-ae1df3d7de5f"
#Total Lucro
round(df["lucro"].sum(),2)
# + colab={} colab_type="code" id="dOlaVDsFqv-t"
#Criando uma coluna com total de dias para enviar o produto
df["Tempo_envio"] = df["Data Envio"] - df["Data Venda"]
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="xzf6mIH5r3vy" outputId="e5444795-08c5-434f-b88a-0043c6fb2c66"
df.head(1)
# + [markdown] colab_type="text" id="tYKqnysZthDh"
# **Agora, queremos saber a média do tempo de envio para cada Marca, e para isso precisamos transformar a coluna Tempo_envio em númerica**
# + colab={} colab_type="code" id="eUAJwu45uVV-"
#Extraindo apenas os dias
df["Tempo_envio"] = (df["Data Envio"] - df["Data Venda"]).dt.days
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="MngNW5dZxjh_" outputId="ce20f74d-8730-463e-ed5e-b0a40104bcd7"
df.head(1)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="k9le4YEvxlow" outputId="4fa5b1a9-e7c4-43ba-f074-7e539c3faf8e"
#Verificando o tipo da coluna Tempo_envio
df["Tempo_envio"].dtype
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="VtCqhtr60byy" outputId="8f08f2ff-50b9-40c3-b103-153a19a4e335"
#Média do tempo de envio por Marca
df.groupby("Marca")["Tempo_envio"].mean().plot.bar()
plt.xlabel=''
# + [markdown] colab_type="text" id="I1sg7kwKjuU1"
# **Missing Values**
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="a26UV-kTjmog" outputId="2db6fd94-9426-4085-c484-81ac68e32e6c"
#Verificando se temos dados faltantes
df.isnull().sum()
# + [markdown] colab_type="text" id="Mh40m00N0lQE"
# **E, se a gente quiser saber o Lucro por Ano e Por Marca?**
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="7CPhZjrJ00a1" outputId="08a87137-f56d-4a34-891d-679beef065b8"
#Vamos Agrupar por ano e marca
df.groupby([df["Data Venda"].dt.year, "Marca"])["lucro"].sum()
# + colab={} colab_type="code" id="kZ3lxKGabXeq"
pd.options.display.float_format = '{:20,.2f}'.format
# + colab={"base_uri": "https://localhost:8080/", "height": 235} colab_type="code" id="knQfX6NC3GMc" outputId="8030fd21-9a09-4637-fe4f-97361465cbf4"
#Resetando o index
lucro_ano = df.groupby([df["Data Venda"].dt.year, "Marca"])["lucro"].sum().reset_index()
lucro_ano
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="0xu9qx1x4WM6" outputId="28166c13-06a7-4532-908a-2cd10156422b"
#Qual o total de produtos vendidos?
df.groupby("Produto")["Quantidade"].sum().sort_values(ascending=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="Ov8qN2bI56NI" outputId="f7f81f4d-dd72-4500-dd5d-3459964fbaff"
#Gráfico Total de produtos vendidos
df.groupby("Produto")["Quantidade"].sum().plot.barh(title="Total Produtos Vendidos", color = "green")
plt.xlabel("Total")
plt.ylabel("Produto");
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="4-FPJ5dP5saX" outputId="3f945085-2546-4643-86b3-bbb2f0f9d235"
#Gráfico
df.groupby(df["Data Venda"].dt.year)["lucro"].sum().plot.bar()
# + colab={} colab_type="code" id="qEjCs7y77966"
#Selecionando apenas as vendas de 2009
df_2009 = df[df["Data Venda"].dt.year == 2009]
# + colab={"base_uri": "https://localhost:8080/", "height": 496} colab_type="code" id="GiL4JRnU_LSf" outputId="f55f66df-5bd7-408b-9113-c7bf694fd2fb"
df_2009.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="xaH-Ym6h_SG9" outputId="1ea0cbdf-ecea-4722-e6c4-9e0557b931c1"
df_2009.groupby(df_2009["Data Venda"].dt.month)["lucro"].sum().plot(title="Lucro x Mês")
#plt.xlabel("Mês")
#plt.ylabel("Lucro");
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="8HDLr3pp_hqf" outputId="e476ce16-0d5f-4bf6-c5c5-7cec1e4ec576"
df_2009.groupby("Marca")["lucro"].sum().plot.bar(title="Lucro x Marca", color = "red")
plt.ylabel("Lucro")
plt.xticks(rotation='horizontal');
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="xguSC8ya_mr7" outputId="bec6125e-6b51-4aac-b9df-d6826c26afdc"
df_2009.groupby("Classe")["lucro"].sum().plot.bar(title="Lucro x Classe", color = "gray")
plt.ylabel("Lucro", color = "red")
plt.xticks(rotation='horizontal');
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="IbO8CjekDdbk" outputId="0f917524-9158-48af-b8d2-a91dfa5d348b"
df["Tempo_envio"].describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 347} colab_type="code" id="yVBuChl7D-LK" outputId="4a661f48-67f7-414e-9993-941548179c6d"
#Gráfico de Boxplot
plt.boxplot(df["Tempo_envio"]);
# + colab={"base_uri": "https://localhost:8080/", "height": 347} colab_type="code" id="AAso8LU5GiFN" outputId="69919c67-916c-490c-a647-1462e363612f"
#Histograma
plt.hist(df["Tempo_envio"]);
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="hkxhLlATHMN3" outputId="c5e19e4f-d830-416f-ce78-7367926adb04"
#Tempo mínimo de envio
df["Tempo_envio"].min()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qg1q3fAKIDtM" outputId="9f4667ac-1557-4591-9fdc-5436c14099c5"
#Tempo máximo de envio
df['Tempo_envio'].max()
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="BiOyhekfIgLb" outputId="732adad6-aa24-4067-a926-7aacf59992e0"
#Identificando o Outlier
df[df["Tempo_envio"] == 20]
# + colab={} colab_type="code" id="xL5IKMeeLI6v"
df.to_csv("df_vendas_novo.csv", index=False)
# + colab={} colab_type="code" id="NLtTuecu62_h"
# -
| Projeto_Exploratoria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Série 8 - Exercices - Questions
#
# Le but de cette série est de s'exercer à l'utilisation des techniques de calcul d'intervalle de confiance sur la moyenne et de les appliquer sur un cas pratique. Il s'agit d'analyser des données collectées au Groënland et en Antarctique pour essayer de déterminer si l'atmosphère se mélange de manière homogène à l'échelle globale de la terre ou pas.
#
# L'idée est que la neige et la glace contiennent des poussières ayant une taille de quelques microns. Ces poussières proviennent de différents processus (volcanismes, tempêtes de poussières, micro-météorites, pollution, etc.). Comme elles sont très petites et légères elle ne retombe pas naturellement. Elle servent toutefois de nucléus pour la formation de la neige.
#
# Elles retournent donc au sol avec la neige et peuvent être incorporées dans de la glace dans les régions polaires. Si les courants atmosphériques mélangent de manière homogène l'atmosphère, on devrait trouver approximativement les mêmes quantités de poussières dans les échantillons en Antarctiques et Groënland.
#
# Pour étudier cette hypothèse, vous allez donc comparer les jeux de données collectés en Antarctique et au Groënland, et en particulier vous allez tenter de déterminer si les moyennes des concentrations en microparticules dans les neiges de ces deux régions sont semblables ou pas.
#
# # Chargement des données
#
# Le fichier `microparticles.txt` contient les données. La première colonne contient un code qui décrit le lieu de collecte de l'échantillon, 0 signifie "Antarctique", 1 signifie "Groënland". La deuxième colonne contient la teneur en microparticules en ppb pour chaque mesure.
#
# **Question 1** - Chargez les données. Calculer les moyennes pour les teneurs. Sont-elles identiques ? Qu'en pensez-vous, commentez ?
# # Visualisation des données
#
# **Question 2** Pour mieux comprendre les données, faites un histogramme et un graphe de type boite à moustache pour les mesures en Antarctique et au Groënland et commentez vos observations.
# # Erreur standard sur la moyenne
#
# Pour y voir plus clair, nous devons estimer des intervalles de confiance sur la moyenne.
#
# **Question 3** - Calculer l'erreur standard sur la moyenne pour les deux jeux de données. Quels sont les intervalles de confiance ?
#
# **Question 4** -Faire un graphe avec les moyennes et barres d'erreur pour montrer visuellement les intervales de confiance des deux jeux de données. Commentez le résultat. Quelle conclusion en tirez-vous ?
# # Bootstrap
#
# **Question 5** - La conclusion ci-dessus est claire. Mais on vous demande de vérifier si la même conclusion est obtenue en utilisant le bootstrap pour faire un calcul alternatif d'intervalle de confiance. Faites le calcul et comparez ? Commentaires ?
#
#
| series/serie08_questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Cloblak/aipi540_deeplearning/blob/main/1D_CNN_Attempts/CDT1D_CNN_181154FEB.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Xj0pR3efRVrc" colab={"base_uri": "https://localhost:8080/"} outputId="5c2b33d2-633d-4f81-9cd4-55ac43b8468f"
# !pip install alpaca_trade_api
# + [markdown] id="hdKRKIogGAu6"
# Features To Consider
# - Targets are only predicting sell within market hours, i.e. at 1530, target is prediciting price for 1100 the next day. Data from pre and post market is taken into consideration, and a sell or buy will be indicated if the price will flucuate after close.
# + id="J1fWNRnTQZX-"
# Import Dependencies
import numpy as np
import pandas as pd
import torch
from torch.utils.data import DataLoader, TensorDataset
from torch.autograd import Variable
from torch.nn import Linear, ReLU, CrossEntropyLoss, Sequential, Conv2d, MaxPool2d, Module, Softmax, BatchNorm2d, Dropout
from torch.optim import Adam, SGD
from torch.utils.data import DataLoader, TensorDataset
from torch.utils.tensorboard import SummaryWriter
from torchsummary import summary
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from tqdm.notebook import tqdm
import alpaca_trade_api as tradeapi
from datetime import datetime, timedelta, tzinfo, timezone, time
import os.path
import ast
import threading
import math
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, MinMaxScaler
import warnings
# + colab={"base_uri": "https://localhost:8080/"} id="DrI_WR501Iis" outputId="1d71636f-388e-4da7-cbe8-35f70ad9497a"
random_seed = 182
torch.manual_seed(random_seed)
# + id="IXnO8ykgRIuv"
PAPER_API_KEY = "<KEY>"
PAPER_SECRET_KEY = "<KEY>"
PAPER_BASE_URL = 'https://paper-api.alpaca.markets'
# + id="_3XShkLcRQMs"
api = tradeapi.REST(PAPER_API_KEY, PAPER_SECRET_KEY, PAPER_BASE_URL, api_version='v2')
# + id="tINNlljbRaDs" colab={"base_uri": "https://localhost:8080/"} outputId="c7b660c5-1d42-480a-9bca-bf284c3d2d94"
def prepost_train_test_validate_offset_data(api, ticker, interval, train_days=180, test_days=60, validate_days=30, offset_days = 0):
ticker_data_dict = None
ticker_data_dict = {}
monthly_data_dict = None
monthly_data_dict = {}
interval_loop_data = None
interval_loop_data = pd.DataFrame()
stock_data = None
days_to_collect = train_days + test_days + validate_days + offset_days
TZ = 'US/Eastern'
start = pd.to_datetime((datetime.now() - timedelta(days=days_to_collect)).strftime("%Y-%m-%d %H:%M"), utc=True)
end = pd.to_datetime(datetime.now().strftime("%Y-%m-%d %H:%M"), utc=True)
stock_data = api.get_bars(ticker, interval, start = start.isoformat(), end=end.isoformat(), adjustment="raw").df
interval_loop_data = interval_loop_data.append(stock_data)
df_start_ref = interval_loop_data.index[0]
start_str_ref = pd.to_datetime(start, utc=True)
while start_str_ref.value < ( pd.to_datetime(df_start_ref, utc=True) - pd.Timedelta(days=2.5)).value:
end_new = pd.to_datetime(interval_loop_data.index[0].strftime("%Y-%m-%d %H:%M"), utc=True).isoformat()
stock_data_new = None
stock_data_new = api.get_bars(ticker, interval, start=start, end=end_new, adjustment="raw").df
#stock_data_new = stock_data_new.reset_index()
interval_loop_data = interval_loop_data.append(stock_data_new).sort_values(by=['index'], ascending=True)
df_start_ref = interval_loop_data.index[0]
stock_yr_min_df = interval_loop_data.copy()
stock_yr_min_df["Open"] = stock_yr_min_df['open']
stock_yr_min_df["High"]= stock_yr_min_df["high"]
stock_yr_min_df["Low"] = stock_yr_min_df["low"]
stock_yr_min_df["Close"] = stock_yr_min_df["close"]
stock_yr_min_df["Volume"] = stock_yr_min_df["volume"]
stock_yr_min_df["VolumeWeightedAvgPrice"] = stock_yr_min_df["vwap"]
stock_yr_min_df["Time"] = stock_yr_min_df.index.tz_convert(TZ)
stock_yr_min_df.index = stock_yr_min_df.index.tz_convert(TZ)
final_df = stock_yr_min_df.filter(["Time", "Open", "High", "Low", "Close", "Volume", "VolumeWeightedAvgPrice"], axis = 1)
first_day = final_df.index[0]
traintest_day = final_df.index[-1] - pd.Timedelta(days= test_days+validate_days+offset_days)
valtest_day = final_df.index[-1] - pd.Timedelta(days= test_days+offset_days)
last_day = final_df.index[-1] - pd.Timedelta(days= offset_days)
training_df = final_df.loc[first_day:traintest_day] #(data_split - pd.Timedelta(days=1))]
validate_df = final_df.loc[traintest_day:valtest_day]
testing_df = final_df.loc[valtest_day:last_day]
full_train = final_df.loc[first_day:last_day]
offset_df = final_df.loc[last_day:]
return training_df, validate_df, testing_df, full_train, offset_df, final_df, traintest_day, valtest_day
from datetime import date
train_start = date(2017, 2, 18)
train_end = date(2020, 3, 29)
train_delta = train_end - train_start
print(f'Number of days of Training Data {train_delta.days}')
val_day_num = 400
print(f'Number of days of Validation Data {val_day_num}')
test_start = train_end + timedelta(val_day_num)
test_end = date.today()
test_delta = (test_end - test_start)
print(f'Number of days of Holdout Test Data {test_delta.days}')
ticker = "WEAT" # Ticker Symbol to Test
interval = "5Min" # Interval of bars
train_day_int = train_delta.days # Size of training set (Jan 2010 - Oct 2017)
val_day_int = val_day_num # Size of validation set
test_day_int = test_delta.days # Size of test set
offset_day_int = 60 # Number of days to off set the training data
train_raw, val_raw, test_raw, full_raw, offset_raw, complete_raw, traintest_day, testval_day = prepost_train_test_validate_offset_data(api, ticker,
interval,
train_days=train_day_int,
test_days=test_day_int,
validate_days=val_day_int,
offset_days = offset_day_int)
def timeFilterAndBackfill(df):
"""
Prep df to be filled out for each trading day:
Time Frame: 0930-1930
Backfilling NaNs
Adjusting Volume to Zero if no Trading data is present
- Assumption is that there were no trades duing that time
We will build over lapping arrays by 30 min to give ourselfs more
oppurtunities to predict during a given trading day
"""
df = df.between_time('07:29','17:29') # intial sorting of data
TZ = 'US/Eastern' # define the correct timezone
start_dateTime = pd.Timestamp(year = df.index[0].year,
month = df.index[0].month,
day = df.index[0].day,
hour = 7, minute = 25, tz = TZ)
end_dateTime = pd.Timestamp(year = df.index[-1].year,
month = df.index[-1].month,
day = df.index[-1].day,
hour = 17, minute = 35, tz = TZ)
# build blank index that has ever 5 min interval represented
dateTime_index = pd.date_range(start_dateTime,
end_dateTime,
freq='5min').tolist()
dateTime_index_df = pd.DataFrame()
dateTime_index_df["Time"] = dateTime_index
filtered_df = pd.merge_asof(dateTime_index_df, df,
on='Time').set_index("Time").between_time('09:29','17:29')
# create the close array by back filling NA, to represent no change in close
closeset_list = []
prev_c = None
for c in filtered_df["Close"]:
if prev_c == None:
if math.isnan(c):
prev_c = 0
closeset_list.append(0)
else:
prev_c = c
closeset_list.append(c)
elif prev_c != None:
if c == prev_c:
closeset_list.append(c)
elif math.isnan(c):
closeset_list.append(prev_c)
else:
closeset_list.append(c)
prev_c = c
filtered_df["Close"] = closeset_list
# create the volume
volumeset_list = []
prev_v = None
for v in filtered_df["Volume"]:
if prev_v == None:
if math.isnan(v):
prev_v = 0
volumeset_list.append(0)
else:
prev_v = v
volumeset_list.append(v)
elif prev_v != None:
if v == prev_v:
volumeset_list.append(0)
prev_v = v
elif math.isnan(v):
volumeset_list.append(0)
prev_v = 0
else:
volumeset_list.append(v)
prev_v = v
filtered_df["Volume"] = volumeset_list
adjvolumeset_list = []
prev_v = None
for v in filtered_df["VolumeWeightedAvgPrice"]:
if prev_v == None:
if math.isnan(v):
prev_v = 0
adjvolumeset_list.append(0)
else:
prev_v = v
adjvolumeset_list.append(v)
elif prev_v != None:
if v == prev_v:
adjvolumeset_list.append(0)
prev_v = v
elif math.isnan(v):
adjvolumeset_list.append(0)
prev_v = 0
else:
adjvolumeset_list.append(v)
prev_v = v
filtered_df["VolumeWeightedAvgPrice"] = adjvolumeset_list
preped_df = filtered_df.backfill()
return preped_df
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="J6Cw_PVrh2lJ" outputId="03786954-4f27-4fd5-e19b-eb70d1cb85ff"
train_raw[0:300]
# + id="fWqLBPQPbjYZ" outputId="36c46af7-d97b-4a1a-e83d-99ddaf74da21" colab={"base_uri": "https://localhost:8080/", "height": 432}
def buildTargets_VolOnly(full_df = full_raw, train_observations = train_raw.shape[0],
val_observations = val_raw.shape[0],
test_observations = test_raw.shape[0],
alph = .55, volity_int = 10):
"""
This function will take a complete set of train, val, and test data and return the targets.
Volitility will be calculated over the 252 5min incriments
The Target shift is looking at 2 hours shift from current time
"""
returns = np.log(full_df['Close']/(full_df['Close'].shift()))
returns.fillna(0, inplace=True)
volatility = returns.rolling(window=(volity_int)).std()*np.sqrt(volity_int)
return volatility
#return train_targets, val_targets, test_targets, full_targets
volatility = buildTargets_VolOnly()
fig = plt.figure(figsize=(15, 7))
ax1 = fig.add_subplot(1, 1, 1)
volatility.plot(ax=ax1, color = "red")
ax1.set_xlabel('Date')
ax1.set_ylabel('Volatility', color = "red")
ax1.set_title(f'Annualized volatility for {ticker}')
ax2 = ax1.twinx()
full_raw.Close.plot(ax=ax2, color = "blue")
ax2.set_ylabel('Close', color = "blue")
ax2.axvline(x=full_raw.index[train_raw.shape[0]])
ax2.axvline(x=full_raw.index[val_raw.shape[0]+train_raw.shape[0]])
plt.show()
# + id="0_yU14VEM37m"
train = timeFilterAndBackfill(train_raw)
val = timeFilterAndBackfill(val_raw)
test = timeFilterAndBackfill(test_raw)
train = train[train.index.dayofweek <= 4].copy()
val = val[val.index.dayofweek <= 4].copy()
test = test[test.index.dayofweek <= 4].copy()
train["Open"] = np.where((train["Volume"] == 0), train["Close"], train["Open"])
train["High"] = np.where((train["Volume"] == 0), train["Close"], train["High"])
train["Low"] = np.where((train["Volume"] == 0), train["Close"], train["Low"])
val["Open"] = np.where((val["Volume"] == 0), val["Close"], val["Open"])
val["High"] = np.where((val["Volume"] == 0), val["Close"], val["High"])
val["Low"] = np.where((val["Volume"] == 0), val["Close"], val["Low"])
test["Open"] = np.where((test["Volume"] == 0), test["Close"], test["Open"])
test["High"] = np.where((test["Volume"] == 0), test["Close"], test["High"])
test["Low"] = np.where((test["Volume"] == 0), test["Close"], test["Low"])
def strided_axis0(a, L, overlap=1):
if L==overlap:
raise Exception("Overlap arg must be smaller than length of windows")
S = L - overlap
nd0 = ((len(a)-L)//S)+1
if nd0*S-S!=len(a)-L:
warnings.warn("Not all elements were covered")
m,n = a.shape
s0,s1 = a.strides
return np.lib.stride_tricks.as_strided(a, shape=(nd0,L,n), strides=(S*s0,s0,s1))
# OLDER CODE WITHOUT OVERLAP OF LABELING
# def blockshaped(arr, nrows, ncols):
# """
# Return an array of shape (n, nrows, ncols) where
# n * nrows * ncols = arr.size
# If arr is a 2D array, the returned array should look like n subblocks with
# each subblock preserving the "physical" layout of arr.
# """
# h, w = arr.shape
# assert h % nrows == 0, f"{h} rows is not evenly divisible by {nrows}"
# assert w % ncols == 0, f"{w} cols is not evenly divisible by {ncols}"
# return np.flip(np.rot90((arr.reshape(h//nrows, nrows, -1, ncols)
# .swapaxes(1,2)
# .reshape(-1, nrows, ncols)), axes = (1, 2)), axis = 1)
def blockshaped(arr, nrows, ncols, overlapping_5min_intervals = 12):
"""
Return an array of shape (n, nrows, ncols) where
n * nrows * ncols = arr.size
If arr is a 2D array, the returned array should look like n subblocks with
each subblock preserving the "physical" layout of arr.
"""
h, w = arr.shape
assert h % nrows == 0, f"{h} rows is not evenly divisible by {nrows}"
assert w % ncols == 0, f"{w} cols is not evenly divisible by {ncols}"
return np.flip(np.rot90((strided_axis0(arr, 24, overlap=overlapping_5min_intervals).reshape(-1, nrows, ncols)), axes = (1, 2)), axis = 1)
train_tonp = train[["Open", "High", "Low", "Close", "Volume"]]
val_tonp = val[["Open", "High", "Low", "Close", "Volume"]]
test_tonp = test[["Open", "High", "Low", "Close", "Volume"]]
train_array = train_tonp.to_numpy()
val_array = val_tonp.to_numpy()
test_array = test_tonp.to_numpy()
X_train_pre_final = blockshaped(train_array, 24, 5, overlapping_5min_intervals = 12)
X_val_pre_final = blockshaped(val_array, 24, 5, overlapping_5min_intervals = 12)
X_test_pre_final = blockshaped(test_array, 24, 5, overlapping_5min_intervals = 12)
# X_train_pre_final = blockshaped(train_array, 24, 5)
# X_val_pre_final = blockshaped(val_array, 24, 5)
# X_test_pre_final = blockshaped(test_array, 24, 5)
# + colab={"base_uri": "https://localhost:8080/"} id="V8KcE_XCidzo" outputId="0a03568d-48af-4229-e4aa-f366052eab28"
np.set_printoptions(edgeitems=10,linewidth=580)
X_train_pre_final[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 833} id="29LbU_ukUK8j" outputId="5719ecad-f34c-4312-f77a-088614606ad0"
train_tonp[0:24]
# + colab={"base_uri": "https://localhost:8080/"} id="Y6w8fb_TU-Tx" outputId="bd31c150-62dc-4a94-a432-00a86cab4ea1"
X_train_pre_final[1]
# + id="Pe89LdnsLltO"
# create target from OHLC and Volume Data
###### THIS IS FOR 3 CLASS FROM PAPER ########
# def buildTargets(obs_array,
# alph = .55,
# volity_int = 10):
# """
# This function will take a complete set of train, val, and test
# data and return the targets. Volitility will be calculated over
# the 24 5min incriments. The Target shift is looking at 2 hours
# shift from current time
# shift_2hour = The amount of time the data interval take to equal 2 hours
# (i.e. 5 min data interval is equal to 24)
# alph = The alpha value for calculating the shift in price
# volity_int = the number of incriments used to calculate volitility
# """
# target_close_list =[]
# for arr in obs_array:
# target_close_list.append(arr[3][-1])
# target_close_df = pd.DataFrame()
# target_close_df["Close"] = target_close_list
# target_close_df["Volitility"] = target_close_df["Close"].rolling(volity_int).std()
# # print(len(volatility), len(target_close_df["Close"]))
# targets = [0] * len(target_close_df.Close)
# targets = np.where(target_close_df.Close.shift() >= (target_close_df.Close * (1 + alph * target_close_df["Volitility"])),
# 2, targets)
# targets = np.where(target_close_df.Close.shift() <= (target_close_df.Close * (1 - alph * target_close_df["Volitility"])),
# 1, targets)
# return targets
#####DISREGUARD THE VOLITLITY######
def buildTargets(obs_array,
alph = .55,
volity_int = 10):
"""
This function will take a complete set of train, val, and test
data and return the targets. Volitility will be calculated over
the 24 5min incriments. The Target shift is looking at 2 hours
shift from current time
shift_2hour = The amount of time the data interval take to equal 2 hours
(i.e. 5 min data interval is equal to 24)
alph = The alpha value for calculating the shift in price
volity_int = the number of incriments used to calculate volitility
"""
target_close_list =[]
for arr in obs_array:
target_close_list.append(arr[3][-1])
target_close_df = pd.DataFrame()
target_close_df["Close"] = target_close_list
target_close_df["Volitility"] = target_close_df["Close"].rolling(volity_int).std()
targets = [0] * len(target_close_df.Close)
targets = np.where(target_close_df.Close.shift() >= (target_close_df.Close * (1 + alph)),
2, targets)
targets = np.where(target_close_df.Close.shift() <= (target_close_df.Close * (1 - alph)),
1, targets)
return targets
#####Binary Class######
# def buildTargets(obs_array,
# alph = .55,
# volity_int = 10):
# """
# This function will take a complete set of train, val, and test
# data and return the targets. Volitility will be calculated over
# the 24 5min incriments. The Target shift is looking at 2 hours
# shift from current time
# shift_2hour = The amount of time the data interval take to equal 2 hours
# (i.e. 5 min data interval is equal to 24)
# alph = The alpha value for calculating the shift in price
# volity_int = the number of incriments used to calculate volitility
# """
# target_close_list =[]
# for arr in obs_array:
# target_close_list.append(arr[3][-1])
# target_close_df = pd.DataFrame()
# target_close_df["Close"] = target_close_list
# target_close_df["Volitility"] = target_close_df["Close"].rolling(volity_int).std()
# # print(len(volatility), len(target_close_df["Close"]))
# targets = [0] * len(target_close_df.Close)
# targets = np.where(target_close_df.Close.shift() >= (target_close_df.Close * (1 + alph)),
# 1, targets)
# return targets
# + id="4aYPOa7INyAl"
volity_val = 10
alph = .0015
y_train_pre_final = buildTargets(X_train_pre_final, alph=alph, volity_int = volity_val)
y_val_pre_final = buildTargets(X_val_pre_final, alph=alph, volity_int = volity_val)
y_test_pre_final = buildTargets(X_test_pre_final, alph=alph, volity_int = volity_val)
# + id="vWIY2rwEYCfM"
def get_class_distribution(obj):
count_dict = {
"up": 0,
"flat": 0,
"down": 0,
}
for i in obj:
if i == 2:
count_dict['up'] += 1
elif i == 1:
count_dict['down'] += 1
elif i == 0:
count_dict['flat'] += 1
else:
print("Check classes.")
return count_dict
# + id="-BsVCfr8YCiX" outputId="28df8d72-8669-4d05-d612-c41d10a7e84b" colab={"base_uri": "https://localhost:8080/", "height": 475}
bfig, axes = plt.subplots(nrows=1, ncols=3, figsize=(25,7))
# Train
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_train_pre_final)]).melt(), x = "variable", y="value", hue="variable", ax=axes[0]).set_title('Class Distribution in Train Set')
# Validation
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_val_pre_final)]).melt(), x = "variable", y="value", hue="variable", ax=axes[1]).set_title('Class Distribution in Val Set')
# Test
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_test_pre_final)]).melt(), x = "variable", y="value", hue="variable", ax=axes[2]).set_title('Class Distribution in Test Set')
# + id="1XdpMcVCo2_b"
def createFinalData_RemoveLateAfternoonData(arr, labels):
assert arr.shape[0] == len(labels), "X data do not match length of y labels"
step_count = 0
filtered_y_labels = []
for i in range(arr.shape[0]):
if i == 0:
final_arr = arr[i]
filtered_y_labels.append(labels[i])
#print(f'Appending index {i}, step_count: {step_count}')
step_count += 1
elif i == 1:
final_arr = np.stack((final_arr, arr[i]))
filtered_y_labels.append(labels[i])
step_count += 1
elif step_count == 0:
final_arr = np.vstack((final_arr, arr[i][None]))
filtered_y_labels.append(labels[i])
#print(f'Appending index {i}, step_count: {step_count}')
step_count += 1
elif (step_count) % 5 == 0:
#print(f'skipping {i} array, step_count: {step_count}')
step_count += 1
elif (step_count) % 6 == 0:
#print(f'skipping {i} array, step_count: {step_count}')
step_count += 1
elif (step_count) % 7 == 0:
#print(f'skipping {i} array, step_count: {step_count}')
step_count = 0
else:
final_arr = np.vstack((final_arr, arr[i][None]))
filtered_y_labels.append(labels[i])
#print(f'Appending index {i}, step_count: {step_count}')
step_count += 1
return final_arr, filtered_y_labels
X_train, y_train = createFinalData_RemoveLateAfternoonData(X_train_pre_final, y_train_pre_final)
X_val, y_val = createFinalData_RemoveLateAfternoonData(X_val_pre_final, y_val_pre_final)
X_test, y_test = createFinalData_RemoveLateAfternoonData(X_test_pre_final, y_test_pre_final)
y_train = np.array(y_train)
y_val = np.array(y_val)
y_test = np.array(y_test)
# + id="ZyvritE4qPNR" outputId="8a90d454-14b2-47d9-cc7b-5581ac9d58fa" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Check it arrays are made correctly
train[12:48]
# + id="Bi-1VYrmn0Eb" outputId="c42bcc7d-b645-4cbc-8a3b-a2ebbdb5e2a4" colab={"base_uri": "https://localhost:8080/"}
np.set_printoptions(threshold=200)
y_train_pre_final[50:75]
# + id="xPkrkhqV4Ef-"
######
# Code fro scaling at a later date
######
# from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
scalers = {}
for i in range(X_train.shape[1]):
scalers[i] = StandardScaler()
X_train[:, i, :] = scalers[i].fit_transform(X_train[:, i, :])
for i in range(X_val.shape[1]):
X_val[:, i, :] = scalers[i].transform(X_val[:, i, :])
for i in range(X_test.shape[1]):
X_test[:, i, :] = scalers[i].transform(X_test[:, i, :])
# + id="kNH38ORXLGfn"
def get_class_distribution(obj):
count_dict = {
"up": 0,
"flat": 0,
"down": 0,
}
for i in obj:
if i == 2:
count_dict['up'] += 1
elif i == 1:
count_dict['down'] += 1
elif i == 0:
count_dict['flat'] += 1
else:
print("Check classes.")
return count_dict
# + outputId="1888427c-9ee1-4731-dcf5-0ced8f985700" colab={"base_uri": "https://localhost:8080/", "height": 475} id="qqTz9-J7LGft"
bfig, axes = plt.subplots(nrows=1, ncols=3, figsize=(25,7))
# Train
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_train)]).melt(), x = "variable", y="value", hue="variable", ax=axes[0]).set_title('Class Distribution in Train Set')
# Validation
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_val)]).melt(), x = "variable", y="value", hue="variable", ax=axes[1]).set_title('Class Distribution in Val Set')
# Test
sns.barplot(data = pd.DataFrame.from_dict([get_class_distribution(y_test)]).melt(), x = "variable", y="value", hue="variable", ax=axes[2]).set_title('Class Distribution in Test Set')
# + [markdown] id="aXbU1Pz7oKam"
# # PyTorch Model Conv2D
# + id="tdeD0Rsc4rqJ"
###### ONLY EXECUTE FOR PYTORCH 2D CNN #####
X_train = X_train.reshape(X_train.shape[0],
1,
X_train.shape[1],
X_train.shape[2]
)
X_val = X_val.reshape(X_val.shape[0],
1,
X_val.shape[1],
X_val.shape[2],
)
X_test = X_test.reshape(X_test.shape[0],
1,
X_test.shape[1],
X_test.shape[2],
)
# + colab={"base_uri": "https://localhost:8080/"} id="QLv4oER44svp" outputId="4cfbc7db-b185-4e3b-a109-0731974a6c81"
print(f'X Train Length {X_train.shape}, y Train Label Length {y_train.shape}')
print(f'X Val Length {X_val.shape}, y Val Label Length {y_val.shape}')
print(f'X Test Length {X_test.shape}, y Test Label Length {y_test.shape}')
# + id="_ncbTnJ4oPhc"
trainset = TensorDataset(torch.from_numpy(X_train).float(),
torch.from_numpy(y_train).long())
testset = TensorDataset(torch.from_numpy(X_test).float(),
torch.from_numpy(y_test).long())
# + colab={"base_uri": "https://localhost:8080/"} id="TQl5pGL7opIj" outputId="749c4e44-1924-48c4-8da0-78d76b4b1887"
train_loader = torch.utils.data.DataLoader(trainset, shuffle=False)
i1, l1 = next(iter(train_loader))
print(i1.shape)
# val_data = []
# for i in range(len(X_val)):
# val_data.append([X_val[i].astype('float'), y_val[i]])
val_loader = torch.utils.data.DataLoader(testset, shuffle=False)
i1, l1 = next(iter(val_loader))
print(i1.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 139} id="WOiXPp7popOT" outputId="da04bfaa-6e23-4cd9-d3f1-3510eb277697"
# Get next batch of training images
windows, labels = iter(train_loader).next()
print(windows.shape)
windows = windows.numpy()
batch_size = 5
# plot the windows in the batch, along with the corresponding labels
fig = plt.figure(figsize=(15, 5))
for idx in range(batch_size):
print(labels)
# + id="1hHxQyq1opQ2"
class StockShiftClassification(nn.Module):
def __init__(self):
super(StockShiftClassification, self).__init__()
self.conv1 = nn.Conv2d(1, 32, kernel_size = (1,3), stride=1, padding = 1)
self.pool1 = nn.MaxPool2d(4,1)
self.conv2 = nn.Conv2d(32, 64, kernel_size = (1,3), stride=1, padding = 1)
self.pool2 = nn.MaxPool2d(3,1)
self.conv3 = nn.Conv2d(64, 128, kernel_size = (1,3), stride=1, padding = 1)
self.pool3 = nn.MaxPool2d(2,1)
self.fc1 = nn.Linear(11520,1000) #calculate this
self.fc2 = nn.Linear(1000,500)
self.fc3 = nn.Linear(500,3)
self.drop = nn.Dropout(p=0.7)
def forward(self, x):
x = F.relu(self.conv1(x))
x = self.pool1(x)
x = self.drop(x)
x = F.relu(self.conv2(x))
x = self.pool2(x)
x = self.drop(x)
x = F.relu(self.conv3(x))
x = self.pool3(x)
x = self.drop(x)
x = x.view(x.size(0), -1)
# Linear layer
x = self.fc1(x)
x = self.fc2(x)
x = self.fc3(x)
output = x #F.log_softmax(x, dim=1)
return output
# + colab={"base_uri": "https://localhost:8080/"} id="_QVWUvZ2opU6" outputId="f0a90a25-df96-4efc-b8ed-c545ff1ac551"
# Instantiate the model
net = StockShiftClassification().float()
# Display a summary of the layers of the model and output shape after each layer
summary(net,(windows.shape[1:]),batch_size=batch_size,device="cpu")
# + id="Mg6bEz7orEsC"
train_x = torch.from_numpy(X_train).float()
train_y = torch.from_numpy(y_train).long()
val_x = torch.from_numpy(X_val).float()
val_y = torch.from_numpy(y_val).long()
# + id="7-A3kueTqlai"
# defining the model
model = net
# defining the optimizer
optimizer = Adam(model.parameters(), lr=0.001, weight_decay=0.00001)
# defining the loss function
criterion = CrossEntropyLoss()
# checking if GPU is available
if torch.cuda.is_available():
model = model.cuda()
criterion = criterion.cuda()
# + id="cnrwBuUoqljR"
from torch.autograd import Variable
def train(epoch, train_x, train_y, val_x, val_y):
model.train()
tr_loss = 0
# getting the training set
x_train, y_train = Variable(train_x), Variable(train_y)
# getting the validation set
x_val, y_val = Variable(val_x), Variable(val_y)
# converting the data into GPU format
if torch.cuda.is_available():
x_train = x_train.cuda()
y_train = y_train.cuda()
x_val = x_val.cuda()
y_val = y_val.cuda()
# clearing the Gradients of the model parameters
optimizer.zero_grad()
# prediction for training and validation set
output_train = model(x_train)
output_val = model(x_val)
# computing the training and validation loss
loss_train = criterion(output_train, y_train)
loss_val = criterion(output_val, y_val)
train_losses.append(loss_train)
val_losses.append(loss_val)
# computing the updated weights of all the model parameters
loss_train.backward()
optimizer.step()
tr_loss = loss_train.item()
if epoch%2 == 0:
# printing the validation loss
print('Epoch : ',epoch+1, '\t', 'loss :', loss_val)
# + id="y23Cfuj6sGPv"
import torch
torch.cuda.empty_cache()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="BF4n8j85qloZ" outputId="5c349331-9d61-4f40-c68c-828e3b84bf95"
# defining the number of epochs
n_epochs = 550
# empty list to store training losses
train_losses = []
# empty list to store validation losses
val_losses = []
# training the model
for epoch in range(n_epochs):
train(epoch, train_x, train_y, val_x, val_y)
# + [markdown] id="qtbdKWSK7l7o"
# # KERAS 1D Model
# + id="lNIh0H7JVnF5"
from tensorflow.keras.utils import to_categorical
y_train = to_categorical(y_train, 3)
y_val = to_categorical(y_val, 3)
y_test = to_categorical(y_test, 3)
# + colab={"base_uri": "https://localhost:8080/"} outputId="cad8a3cf-7c19-41d2-bba5-56463dab1e0d" id="xUtWCihJoI1e"
print(f'X Train Length {X_train.shape}, y Train Label Length {y_train.shape}')
print(f'X Val Length {X_val.shape}, y Val Label Length {y_val.shape}')
print(f'X Test Length {X_test.shape}, y Test Label Length {y_test.shape}')
# + id="MUE5rS-l7z9B"
import tensorflow as tf
import keras.backend as K
from keras.models import Sequential
from keras.layers import Dense, BatchNormalization
from keras.layers import Flatten
from keras.layers import Dropout
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from tensorflow.keras.optimizers import Adam, Nadam
# + colab={"base_uri": "https://localhost:8080/"} id="zYwZW6eJ2HBu" outputId="2bc0ccce-6884-4f9f-be93-86c75631dc72"
# Simple Model
model = Sequential()
model.add(Conv1D(filters=32, kernel_size=3, padding='same', activation='relu', input_shape=(X_train.shape[1],X_train.shape[2])))
model.add(MaxPooling1D(pool_size=3, strides=1, padding='same'))
model.add(Dropout(0.7))
model.add(Conv1D(filters=64, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=2, strides=3, padding='same'))
model.add(Dropout(0.7))
model.add(Conv1D(filters=128, kernel_size=1, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=1, strides=4, padding='same'))
model.add(Dropout(0.7))
model.add(Flatten())
model.add(Dense(1000,activation="relu"))
model.add(Dense(500,activation="relu"))
model.add(Dense(3,activation="softmax"))
model.summary()
# + id="SOmAeEzs2HGb"
adam = Adam(learning_rate=0.001, decay=0.0001)
nadam = Nadam(learning_rate=0.002, beta_1=0.9, beta_2=0.999, epsilon=1e-08, schedule_decay=0.004)
def custom_loss_function(y_true, y_pred):
squared_difference = tf.square(y_true - y_pred)
return tf.reduce_mean(squared_difference, axis=-1)
model.compile(loss='categorical_crossentropy', optimizer=adam, metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="ux9JsTrE2HJz" outputId="f5f69606-1105-44bd-de38-d792e7710063"
batch_size = 12
hist = model.fit(X_train, y_train, batch_size=batch_size, epochs=350, validation_data=(X_val, y_val)) #, class_weight={0:1, 1:1.5})
# + id="CMsqd9n02HMv"
plt.plot(hist.history['loss'], label='train loss')
plt.plot(hist.history['val_loss'], label='val loss')
plt.legend()
plt.show()
plt.savefig('LossVal_loss')
# + id="M_S_jaA_2ubl"
plt.plot(hist.history['accuracy'], label='train acc')
plt.plot(hist.history['val_accuracy'], label='val acc')
plt.legend()
plt.show()
plt.savefig('AccVal_acc')
# + id="Gct8IuVa25GW"
y_pred = model.predict(X_test)
# + id="rk2qkycw25Gc"
# Calculate the accuracy
test_preds = np.argmax(y_pred, axis=1)
y_true = np.argmax(y_test, axis=1)
test_acc = np.sum(test_preds == y_true)/y_true.shape[0]
# Recall for each class
recall_vals = []
for i in range(3):
class_idx = np.argwhere(y_true==i)
total = len(class_idx)
correct = np.sum(test_preds[class_idx]==i)
recall = correct / total
recall_vals.append(recall)
classes = [0,1,2]
# Calculate the test set accuracy and recall for each class
print('Test set accuracy is {:.3f}'.format(test_acc))
for i in range(3):
print('For class {}, recall is {:.3f}'.format(classes[i],recall_vals[i]))
print("Accuracy is {:.3f}".format(test_acc))
# print("Weighted F score is {:.3f}".format(calculate_weighted_f_score(y_true, y_pred)))
# + id="coQSJfRo2uoe"
from sklearn.metrics import confusion_matrix
def plot_confusion_matrix(cm,
target_names,
title='Confusion matrix',
cmap=None,
normalize=True):
"""
given a sklearn confusion matrix (cm), make a nice plot
Arguments
---------
cm: confusion matrix from sklearn.metrics.confusion_matrix
target_names: given classification classes such as [0, 1, 2]
the class names, for example: ['high', 'medium', 'low']
title: the text to display at the top of the matrix
cmap: the gradient of the values displayed from matplotlib.pyplot.cm
see http://matplotlib.org/examples/color/colormaps_reference.html
plt.get_cmap('jet') or plt.cm.Blues
normalize: If False, plot the raw numbers
If True, plot the proportions
Usage
-----
plot_confusion_matrix(cm = cm, # confusion matrix created by
# sklearn.metrics.confusion_matrix
normalize = True, # show proportions
target_names = y_labels_vals, # list of names of the classes
title = best_estimator_name) # title of graph
Citiation
---------
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
"""
import matplotlib.pyplot as plt
import numpy as np
import itertools
accuracy = np.trace(cm) / np.sum(cm).astype('float')
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass))
plt.show()
nb_classes = 2
# Confusion matrix
conf_mat=confusion_matrix(y_true, np.argmax(y_pred, axis=-1))
plot_confusion_matrix(conf_mat, [0,1,2])
from sklearn.metrics import precision_score
precision_score(y_true, np.argmax(y_pred, axis=-1), average='weighted')
from sklearn.metrics import classification_report
print(classification_report(y_true, np.argmax(y_pred, axis=-1), target_names=["flat","down", "up"], digits=4))
# + id="F7n6lPZp2uvK"
def calculate_weighted_f_score(y_true, y_pred):
test_preds = np.argmax(y_pred, axis=-1)
Ntu = sum((test_preds == 2) & (y_true == 2))
Ntd = sum((test_preds == 1) & (y_true == 1))
Ntf = sum((test_preds == 0) & (y_true == 0))
Ewutd = sum((test_preds == 2) & (y_true == 1))
Ewdtu = sum((test_preds == 1) & (y_true == 2))
Ewutf = sum((test_preds == 2) & (y_true == 0))
Ewdtf = sum((test_preds == 1) & (y_true == 0))
Ewftu = sum((test_preds == 0) & (y_true == 2))
Ewftd = sum((test_preds == 0) & (y_true == 1))
beta_1 = 0.5
beta_2 = 0.125
beta_3 = 0.125
Ntp = Ntu + Ntd + beta_3**2 * Ntf
E1 = Ewutd + Ewdtu
E2 = Ewutf + Ewdtf
E3 = Ewftu + Ewftd
F = (1 + beta_1**2 + beta_2**2) * Ntp / ((1+beta_1**2+beta_2**2) * Ntp + E1 + beta_1**2 * E2 + beta_2**2 * E3)
return F
print(f'Weight CDT F Score: {calculate_weighted_f_score(y_true, y_pred)}')
# + id="7v0QHbxx2uyQ"
# + id="Z9EfhZEA70xZ"
def calculate_weighted_f_score(y_true, y_pred):
test_preds = np.argmax(y_pred, axis=1)
Ntu = sum((test_preds == 1) & (y_true == 1))
Ntd = sum((test_preds == 0) & (y_true == 0))
Ntf = sum((test_preds == 2) & (y_true == 2))
Ewutd = sum((test_preds == 1) & (y_true == 0))
Ewdtu = sum((test_preds == 0) & (y_true == 1))
Ewutf = sum((test_preds == 1) & (y_true == 2))
Ewdtf = sum((test_preds == 0) & (y_true == 2))
Ewftu = sum((test_preds == 2) & (y_true == 1))
Ewftd = sum((test_preds == 2) & (y_true == 0))
beta_1 = 0.5
beta_2 = 0.125
beta_3 = 0.125
Ntp = Ntu + Ntd + beta_3**2 * Ntf
E1 = Ewutd + Ewdtu
E2 = Ewutf + Ewdtf
E3 = Ewftu + Ewftd
F = (1 + beta_1**2 + beta_2**2) * Ntp / ((1+beta_1**2+beta_2**2) * Ntp + E1 + beta_1**2 * E2 + beta_2**2 * E3)
return F
# + colab={"base_uri": "https://localhost:8080/"} id="9ytUuvub8xcJ" outputId="d7408444-1252-4153-95d4-1d7e14e813f6"
print(X_train.shape[1],X_train.shape[2])
# + colab={"base_uri": "https://localhost:8080/"} id="q50kVGG076-F" outputId="99b837fa-103d-4eee-dc58-98a42a0fdee3"
model = Sequential()
model.add(Conv1D(filters=32, kernel_size=3, padding='same', activation='relu', input_shape=(X_train.shape[1],X_train.shape[2])))
model.add(MaxPooling1D(pool_size=3, strides=4, padding='same'))
model.add(Dropout(0.7))
model.add(Conv1D(filters=64, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=3, strides=3, padding='same'))
model.add(Dropout(0.7))
model.add(Conv1D(filters=128, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPooling1D(pool_size=3, strides=2, padding='same'))
model.add(Dropout(0.7))
model.add(Flatten())
model.add(Dense(1000,activation="relu"))
model.add(Dense(500,activation="relu"))
model.add(Dense(2,activation="softmax"))
model.summary()
# + id="vKDmE-h59Lv9"
adam = Adam(learning_rate=0.001, decay=0.0001)
nadam = Nadam(learning_rate=0.002, beta_1=0.9, beta_2=0.999, epsilon=1e-08, schedule_decay=0.004)
def custom_loss_function(y_true, y_pred):
squared_difference = tf.square(y_true - y_pred)
return tf.reduce_mean(squared_difference, axis=-1)
def calculate_weighted_f_score(y_true, y_pred):
test_preds = np.argmax(y_pred, axis=-1)
print(test_preds)
Ntu = sum((test_preds == 1) & (y_true == 1))
Ntd = sum((test_preds == 0) & (y_true == 0))
Ntf = sum((test_preds == 2) & (y_true == 2))
Ewutd = sum((test_preds == 1) & (y_true == 0))
Ewdtu = sum((test_preds == 0) & (y_true == 1))
Ewutf = sum((test_preds == 1) & (y_true == 2))
Ewdtf = sum((test_preds == 0) & (y_true == 2))
Ewftu = sum((test_preds == 2) & (y_true == 1))
Ewftd = sum((test_preds == 2) & (y_true == 0))
beta_1 = 0.5
beta_2 = 0.125
beta_3 = 0.125
Ntp = Ntu + Ntd + beta_3**2 * Ntf
E1 = Ewutd + Ewdtu
E2 = Ewutf + Ewdtf
E3 = Ewftu + Ewftd
F = (1 + beta_1**2 + beta_2**2) * Ntp / ((1+beta_1**2+beta_2**2) * Ntp + E1 + beta_1**2 * E2 + beta_2**2 * E3)
return F
prec = tf.keras.metrics.Precision()
model.compile(loss='binary_crossentropy', optimizer=adam, metrics=[prec])
# + colab={"base_uri": "https://localhost:8080/"} id="qK4V5UIa9ORe" outputId="8b6e9317-44ba-438e-aab9-e9ed3113f482"
batch_size = 12
hist = model.fit(X_train, y_train, batch_size=batch_size, epochs=50, validation_data=(X_val, y_val)) #, class_weight={0:2, 1:3, 2:1})
# + colab={"base_uri": "https://localhost:8080/"} id="Nj9-LfyGhwZz" outputId="b3e8eb0b-0d48-4d9b-91af-3f1dc7360708"
hist.history
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="PP75W71D9ij8" outputId="93ecd498-1356-42bf-bd10-6db42c5d3c63"
plt.plot(hist.history['loss'], label='train loss')
plt.plot(hist.history['val_loss'], label='val loss')
plt.legend()
plt.show()
plt.savefig('LossVal_loss')
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="esQzOZTb9p48" outputId="8ac28837-c403-4d2a-ed6b-58803bd453b0"
plt.plot(hist.history['precision_7'], label='train acc')
plt.plot(hist.history['val_precision_7'], label='val acc')
plt.legend()
plt.show()
plt.savefig('AccVal_acc')
# + id="MNh51nKK9u15"
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="IuLzuden9xuZ" outputId="440cf5f8-14b0-4d88-ecc1-12a09fae9cb1"
# Calculate the accuracy
test_preds = np.argmax(y_pred, axis=1)
y_true = np.argmax(y_test, axis=1)
test_acc = np.sum(test_preds == y_true)/y_true.shape[0]
# Recall for each class
recall_vals = []
for i in range(3):
class_idx = np.argwhere(y_true==i)
total = len(class_idx)
correct = np.sum(test_preds[class_idx]==i)
recall = correct / total
recall_vals.append(recall)
classes = [0,1]
# Calculate the test set accuracy and recall for each class
print('Test set accuracy is {:.3f}'.format(test_acc))
for i in range(2):
print('For class {}, recall is {:.3f}'.format(classes[i],recall_vals[i]))
print("Weighted F score is {:.3f}".format(test_acc))
# print("Weighted F score is {:.3f}".format(calculate_weighted_f_score(y_true, y_pred)))
# + [markdown] id="KHDKw2Fj3Q9Z"
# # Completely New Model Build
# + id="uBUl4s7dF9ni"
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import Dropout
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
# + id="hFN0cl_cSjwq"
model = Sequential()
model.add(Conv2D(32, (1, 3), activation='relu', padding='same', input_shape=(X_train.shape[1], X_train.shape[2], X_train.shape[3])))
model.add(MaxPooling2D((1, 4), strides=4))
model.add(Dropout(0.7))
model.add(Conv2D(64, (1, 3), activation='relu', padding='same',))
model.add(MaxPooling2D((1, 3), strides=3))
model.add(Dropout(0.7))
model.add(Conv2D(128, (1, 3), activation='relu', padding='same',))
model.add(MaxPooling2D((1, 2), strides=2))
model.add(Dropout(0.7))
model.add(Flatten())
model.add(Dense(1000,activation="relu"))
model.add(Dense(500,activation="relu"))
model.add(Dense(3,activation="softmax"))
# + colab={"base_uri": "https://localhost:8080/"} id="-KKO5LSUTB1v" outputId="f16f4296-8e3b-4f7b-b141-9a6623e31a22"
model.summary()
# + id="NRIZjV2-THkm"
model.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="2EDJJqlJUMHb" outputId="a214b6fc-c9d2-435d-9a02-b8a3db3829cf"
batch_size = 12
hist = model.fit(X_train, y_train, batch_size=batch_size, epochs=50, validation_data=(X_val, y_val))
##### LABLES NEED TO BY 1x3 ARRAY
# + id="5S0IDmg7URVU"
| 1D_CNN_Attempts/CDT1D_CNN_181154FEB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (BoW)
# language: python
# name: pycharm-81ce5fe2
# ---
# + pycharm={"name": "#%%\n"}
from pathlib import Path
# %pip install -U tensorflow-addons
# %pip show numpy
# +
import pandas as pd
import re
import nltk
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sn
import tensorflow_addons as tfa
LABELS = ['positive', 'neutral', 'negative']
SAVE_MODEL = True
DUMP_DIRECTORY = 'created_models'
TRAIN_RANDOM_STATE = 42
DEV_RANDOM_STATE = 42
DROP=0.2
LAYER_1_DENSITY = 1024
LAYER_2_DENSITY = 512
LAYER_3_DENSITY = 256
ACTIVATION = 'sigmoid'
EMBEDDING_DIMENSIONS = 100
LAST_DIMENSIONS = 3 #based on labels
LAST_ACTIVATION = 'softmax'
MAX_LENGTH = 1000
NUM_EPOCHS = 50
BATCH_SIZE = 600
COMPILE_LOSS = 'categorical_crossentropy'
COMPILE_METRICS = ['accuracy']
GLOVE_DIMENSIONS = 100
# + pycharm={"name": "#%%\n"}
import os
from datetime import datetime
import joblib
def dump_file(o, object_name, name):
if not os.path.exists(DUMP_DIRECTORY):
os.mkdir(DUMP_DIRECTORY)
time = datetime.now().strftime("%d%b%Y%H%M%S")
created_model_path = DUMP_DIRECTORY + '/' + name + '_' + time + '.dump'
joblib.dump(o, created_model_path)
# + pycharm={"name": "#%%\n"}
from nltk.corpus import stopwords
from sklearn.utils import shuffle
HEADER = ['id1', 'id2', 'sentiment', 'tweet_text']
HEADER_TO_DELETE = ['id1', 'id2']
def remove_stopwords(input_text):
stopwords_list = stopwords.words('english')
whitelist = ["n't", "not", "no"]
words = input_text.split()
clean_words = [word for word in words if (word not in stopwords_list or word in whitelist) and len(word) > 1]
return " ".join(clean_words)
def remove_mentions(input_text):
URL_RE = 'https?:\/\/(www\.)?[-a-zA-Z0-9@:%._\+~#=]{1,256}\.[a-zA-Z0-9()]{1,6}([-a-zA-Z0-9()@:%_+.~#?&/=]*)'
input_text = re.sub(URL_RE, '', input_text)
input_text = re.sub(r'@\w+', '', input_text)
return re.sub(r'#\w+', '', input_text)
def clean(dataset, col='tweet_text', not_equals_text='Not Available'):
return dataset[dataset[col] != not_equals_text]
def merge_neutrals(dataset):
neutral_sentiments = ['objective', 'objective-OR-neutral']
dataset['sentiment'] = dataset['sentiment'].apply(lambda x: 'neutral' if x in neutral_sentiments else x)
return dataset
def evaluate(predict, labels):
print('Classification report:')
print(classification_report(labels, predict))
print('Accuracy:')
print(accuracy_score(labels, predict))
print('Confusion matrix:')
df_cm = pd.DataFrame(confusion_matrix(labels, predict),
index=[i for i in ['positive', 'neutral', 'negative']],
columns=[i for i in ['positive', 'neutral', 'negative']])
plt.figure(figsize=(10,7))
hm = sn.heatmap(df_cm, annot=True, fmt='g', cmap="Blues")
hm.set(ylabel='True label', xlabel='Predicted label')
plt.show()
train_dataset = pd.read_csv('db/train.tsv', sep='\t', header=None, names=HEADER)
train_dataset = shuffle(train_dataset, random_state=TRAIN_RANDOM_STATE)
train_dataset = clean(train_dataset)
train_dataset = merge_neutrals(train_dataset)
train_dataset.drop(HEADER_TO_DELETE, axis=1, inplace=True)
train_dataset['tweet_text'] = train_dataset['tweet_text'].apply(remove_stopwords).apply(remove_mentions)
train_dataset.info()
dev_dataset = pd.read_csv('db/dev-full.tsv', sep='\t', header=None, names=HEADER)
dev_dataset = shuffle(dev_dataset, random_state=DEV_RANDOM_STATE)
dev_dataset = clean(dev_dataset)
dev_dataset = merge_neutrals(dev_dataset)
dev_dataset.drop(HEADER_TO_DELETE, axis=1, inplace=True)
dev_dataset['tweet_text'] = dev_dataset['tweet_text'].apply(remove_stopwords).apply(remove_mentions)
dev_dataset.info()
# + pycharm={"name": "#%%\n"}
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
def evaluate(predict, labels, history):
print('Classification report:')
print(classification_report(labels, predict))
print('Accuracy:')
print(accuracy_score(labels, predict))
print('Confusion matrix:')
df_cm = pd.DataFrame(confusion_matrix(labels, predict),
index=[i for i in ['positive', 'neutral', 'negative']],
columns=[i for i in ['positive', 'neutral', 'negative']])
plt.figure(figsize=(10,7))
hm = sn.heatmap(df_cm, annot=True, fmt='g', cmap="Blues")
hm.set(ylabel='True label', xlabel='Predicted label')
plt.show()
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train acc', 'val acc'], loc='upper left')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'val loss'], loc='upper left')
plt.show()
x_train = train_dataset['tweet_text'].values
y_train = train_dataset['sentiment'].values
x_dev = dev_dataset['tweet_text'].values
y_dev = dev_dataset['sentiment'].values
train_dataset.head(5)
# + pycharm={"name": "#%%\n"}
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
# tk = Tokenizer(num_words=NUM_WORDS, filters='!"#$%&()*+,-./:;<=>?@[\]^_`{"}~\t\n',
# lower=True, split=" ")
tk = Tokenizer()
all_tweets = np.append(x_train, x_dev)
tk.fit_on_texts(all_tweets)
vocab_size = len(tk.word_index) + 1
x_train_seq = tk.texts_to_sequences(x_train)
x_dev_seq = tk.texts_to_sequences(x_dev)
x_train_seq_trunc = pad_sequences(x_train_seq, maxlen=MAX_LENGTH, padding='post')
x_dev_seq_trunc = pad_sequences(x_dev_seq, maxlen=MAX_LENGTH, padding='post')
# + pycharm={"name": "#%%\n"}
from sklearn.preprocessing import LabelEncoder
from keras.utils.np_utils import to_categorical
le = LabelEncoder()
y_train_le = le.fit_transform(y_train)
y_dev_le = le.fit_transform(y_dev)
y_train_categorical = to_categorical(y_train_le)
y_dev_categorical = to_categorical(y_dev_le)
# + pycharm={"name": "#%%\n"}
from keras import models
from keras import layers
def create_model(embedding_layer):
sequence_input = models.Input(shape=(MAX_LENGTH,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
x = layers.Conv1D(128, 5, activation='relu')(embedded_sequences)
x = layers.MaxPooling1D(5)(x)
x = layers.Conv1D(128, 5, activation='relu')(x)
x = layers.MaxPooling1D(5)(x)
x = layers.Conv1D(128, 5, activation='relu')(x)
x = layers.MaxPooling1D(35)(x)
x = layers.Flatten()(x)
x = layers.Dense(128, activation='relu')(x)
preds = layers.Dense(LAST_DIMENSIONS, activation='softmax')(x)
return sequence_input, preds
embedding_layer = layers.Embedding(vocab_size, GLOVE_DIMENSIONS, input_length=max_length)
sequence_input, preds = create_model(embedding_layer)
emb_model = models.Model(sequence_input, preds)
emb_model.summary()
# + pycharm={"name": "#%%\n"}
tqdm_callback = tfa.callbacks.TQDMProgressBar()
from tensorflow.python.keras.callbacks import EarlyStopping
def deep_model(model, x_train, y_train, x_dev, y_dev):
model.compile(loss=COMPILE_LOSS, metrics=COMPILE_METRICS, optimizer='adam')
history = model.fit(x_train, y_train, epochs=NUM_EPOCHS, batch_size=BATCH_SIZE,
validation_data=(x_dev, y_dev), verbose=0, callbacks=[EarlyStopping(patience=2), tqdm_callback])
# TODO: shuffle -> random state, with seeds
# if SAVE_MODEL:
# dump_file(model, 'neural net', 'deep_model')
result = model.evaluate(x_dev, y_dev)
predict = model.predict_classes(x_dev)
return history, result, predict
# + pycharm={"name": "#%%\n"}
emb_history, emb_result, emb_predict = deep_model(emb_model, x_train_seq_trunc, y_train_categorical,
x_dev_seq_trunc, y_dev_categorical)
# + pycharm={"name": "#%%\n"}
print(emb_result)
evaluate(emb_predict, y_dev_le, emb_history)
# + pycharm={"name": "#%%\n"}
glove_file = 'glove.twitter.27B.' + str(GLOVE_DIMENSIONS) + 'd.txt'
emb_dict = {}
glove = open(Path('./db') / glove_file, encoding="utf8")
for line in glove:
values = line.split()
word = values[0]
vector = np.asarray(values[1:], dtype='float32')
emb_dict[word] = vector
glove.close()
# + pycharm={"name": "#%%\n"}
emb_matrix = np.zeros((vocab_size, GLOVE_DIMENSIONS))
for w, i in tk.word_index.items():
if i < vocab_size:
vect = emb_dict.get(w)
if vect is not None:
emb_matrix[i] = vect
else:
break
# + pycharm={"name": "#%%\n"}
embedding_layer = layers.Embedding(vocab_size, GLOVE_DIMENSIONS, input_length=max_length,
weights=[emb_matrix], trainable=False)
sequence_input, preds = create_model(embedding_layer)
glove_model = models.Model(sequence_input, preds)
glove_model.summary()
# + pycharm={"name": "#%%\n"}
glove_model.layers[0].set_weights([emb_matrix])
glove_model.layers[0].trainable = False
glove_history, glove_result, glove_predict = deep_model(glove_model, x_train_seq_trunc, y_train_categorical,
x_dev_seq_trunc, y_dev_categorical)
# + pycharm={"name": "#%%\n"}
print(glove_result)
evaluate(glove_predict, y_dev_le, glove_history)
| word_embedding_try_to_save.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: My CogPhenoPark Kernel
# language: python
# name: cogphenoparkernel
# ---
# +
import os
import argparse
import math
from decimal import Decimal
from os.path import join
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats
import pandas as pd
from tqdm import tqdm
from tractseg.data import dataset_specific_utils
from tractseg.libs.AFQ_MultiCompCorrection import AFQ_MultiCompCorrection
from tractseg.libs.AFQ_MultiCompCorrection import get_significant_areas
from tractseg.libs import metric_utils
from tractseg.libs import plot_utils
from tractseg.libs import tracking
import glob as glob
import nibabel as nib
from scipy.stats import t as t_dist
# -
def parse_subjects_file(file_path):
with open(file_path) as f:
l = f.readline().strip()
if l.startswith("# tractometry_path="):
base_path = l.split("=")[1]
else:
raise ValueError("Invalid first line in subjects file. Must start with '# tractometry_path='")
bundles = None
plot_3D_path = None
# parse bundle names
for i in range(2):
l = f.readline().strip()
if l.startswith("# bundles="):
bundles_string = l.split("=")[1]
bundles = bundles_string.split(" ")
valid_bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
for bundle in bundles:
if bundle not in valid_bundles:
raise ValueError("Invalid bundle name: {}".format(bundle))
print("Using {} manually specified bundles.".format(len(bundles)))
elif l.startswith("# plot_3D="):
plot_3D_path = l.split("=")[1]
#if bundles is None:
# bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
bundles = dataset_specific_utils.get_bundle_names("All")[1:]
df = pd.read_csv(file_path, sep=" ", comment="#")
df["subject_id"] = df["subject_id"].astype(str)
# Check that each column (except for first one) is correctly parsed as a number
for col in df.columns[1:]:
if not np.issubdtype(df[col].dtype, np.number):
raise IOError("Column {} contains non-numeric values".format(col))
#if df.columns[1] == "group":
# if df["group"].max() > 1:
# raise IOError("Column 'group' may only contain 0 and 1.")
return base_path, df, bundles, plot_3D_path
def correct_for_confounds(values, meta_data, bundles, selected_bun_indices, NR_POINTS, analysis_type, confound_names):
values_cor = np.zeros([len(bundles), NR_POINTS, len(meta_data)])
for b_idx in selected_bun_indices:
for jdx in range(NR_POINTS):
target = np.array([values[s][b_idx][jdx] for s in meta_data["subject_id"]])
if analysis_type == "group":
target_cor = metric_utils.unconfound(target, meta_data[["group"] + confound_names].values,
group_data=True)
else:
target_cor = metric_utils.unconfound(target, meta_data[confound_names].values,
group_data=False)
meta_data["target"] = metric_utils.unconfound(meta_data["target"].values,
meta_data[confound_names].values,
group_data=False)
values_cor[b_idx, jdx, :] = target_cor
# Restore original data structure
values_cor = values_cor.transpose(2, 0, 1)
# todo: nicer way: use numpy array right from beginning instead of dict
values_cor_dict = {}
for idx, subject in enumerate(list(meta_data["subject_id"])):
values_cor_dict[subject] = values_cor[idx]
return values_cor_dict
def get_corrected_alpha(values_allp, meta_data, analysis_type, subjects_A, subjects_B, alpha, bundles, nperm, b_idx):
if analysis_type == "group":
y = np.array((0,) * len(subjects_A) + (1,) * len(subjects_B))
else:
y = meta_data["target"].values
alphaFWE, statFWE, clusterFWE, stats = AFQ_MultiCompCorrection(np.array(values_allp), y,
alpha, nperm=nperm)
#print("Processing {}...".format(bundles[b_idx]))
#print(" cluster size: {}".format(clusterFWE))
#print(" alphaFWE: {}".format(format_number(alphaFWE)))
#return alphaFWE, clusterFWE,stats
return alphaFWE, clusterFWE
def format_number(num):
if abs(num) > 0.00001:
return round(num, 4)
else:
return '%.2e' % Decimal(num)
# +
#FWE_method = "alphaFWE"
FWE_method = "clusterFWE"
show_detailed_p = False
hide_legend = False
show_color_bar = True # colorbar on 3D plot
nperm = 5000
alpha=0.05
correct_mult_tract_comp = False
base_path, meta_data, selected_bundles, plot_3D_path = parse_subjects_file("/mnt/d//LINUX/CogPhenoPark/dataTractSeg/Tractometry_FA.txt")
analysis_type = "group"
plot_3D_type="pval"
tracking_format="tck"
tracking_dir="FOD_iFOD2_trackings"
output_path="/mnt/d//LINUX/CogPhenoPark"
# +
all_bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
values = {}
for subject in meta_data["subject_id"]:
raw = np.loadtxt(base_path.replace("SUBJECT_ID", subject), delimiter=";", skiprows=1).transpose()
values[subject] = raw
NR_POINTS = values[meta_data["subject_id"][0]].shape[1]
#selected_bun_indices = [bundles.index(b) for b in selected_bundles]
selected_bun_indices = [all_bundles.index(b) for b in selected_bundles]
print(selected_bun_indices)
# -
# # Two T_Test
correct_mult_tract_comp=True
for withCofound in [True]:
for para in (['''RD''','''FA''','''MD''','''AD''','''density''']):
#for para in ['''RD''']:
print("***************** "+ para+" *********************")
for group_vs in (['''G1VsG3''','''G3VsG4''','''G2VsG3''','''G1VsG2''','''G1VsG4''','''G2VsG4''']):
#for group_vs in ['''G1VsG3''']:
print("***************** "+ group_vs +" *********************")
#for ind in [True,False]:
for ind in [False]:
for show_detailed_p in [True,False]:
###############
FWE_method = "alphaFWE"
show_detailed_p = False
hide_legend = False
show_color_bar = True # colorbar on 3D plot
nperm = 5000
alpha=0.05
correct_mult_tract_comp = False
base_path, meta_data, selected_bundles, plot_3D_path = parse_subjects_file("/mnt/d//LINUX/CogPhenoPark/dataTractSeg/Tractometry_template_"+group_vs+".txt")
analysis_type = "group"
plot_3D_type="none"
###########
all_bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
values = {}
for subject in meta_data["subject_id"]:
raw = np.loadtxt(base_path.replace("SUBJECT_ID", subject).replace("PARA",para), delimiter=";", skiprows=1).transpose()
values[subject] = raw
NR_POINTS = values[meta_data["subject_id"][0]].shape[1]
selected_bun_indices = [all_bundles.index(b) for b in selected_bundles]
confound_names = list(meta_data.columns[2:])
cols = 3
rows = math.ceil(len(selected_bundles) / cols)
a4_dims = (cols*3, rows*5)
f, axes = plt.subplots(rows, cols, figsize=a4_dims)
axes = axes.flatten()
sns.set(font_scale=1.2)
sns.set_style("whitegrid")
subjects_A = list(meta_data[meta_data["group"] == 0]["subject_id"])
subjects_B = list(meta_data[meta_data["group"] == 1]["subject_id"])
# Correct for confounds
if withCofound :
values = correct_for_confounds(values, meta_data, all_bundles, selected_bun_indices, NR_POINTS, analysis_type,confound_names)
# Significance testing with multiple correction of bundles
if correct_mult_tract_comp:
values_allp = [] # [subjects, NR_POINTS * nr_bundles]
for s in meta_data["subject_id"]:
print(s)
values_subject = []
for i, b_idx in enumerate(selected_bun_indices):
# print(b_idx)
# print(np.mean(values[s][b_idx]))
values_subject += list(values[s][b_idx]) # concatenate all bundles
values_allp.append(values_subject)
alphaFWE, clusterFWE = get_corrected_alpha(values_allp, meta_data, analysis_type, subjects_A, subjects_B, alpha,all_bundles, nperm, b_idx)
for i, b_idx in enumerate(selected_bun_indices):
############
vals_thresA=np.zeros([])
subjects_AA=subjects_A[:]
for subject in subjects_A:
if np.all(values[subject][b_idx]>0) :
vals_thresA=np.append(vals_thresA,np.mean(values[subject][b_idx]))
vals_thresA=vals_thresA[1:]
vals_thresA = vals_thresA[~ np.isnan(vals_thresA)]
val_thresA=np.mean(vals_thresA)-2*np.std(vals_thresA)
if val_thresA < 0 : val_thresA = 0
#print("valeur seuil G0= "+str(val_thresA))
vals_thresB=np.zeros([])
subjects_BB=subjects_B[:]
for j, subject in enumerate(subjects_B):
if np.all(values[subject][b_idx]>0) :
vals_thresB=np.append(vals_thresB,np.mean(values[subject][b_idx]))
vals_thresB=vals_thresB[1:]
vals_thresB = vals_thresB[~ np.isnan(vals_thresB)]
val_thresB=np.mean(vals_thresB)-2*np.std(vals_thresB)
if val_thresB < 0 : val_thresB = 0
#print("valeur seuil G1= "+str(val_thresB))
# Bring data into right format for seaborn
data = {"position": [],
"fa": [],
"group": [],
"subject": []}
subjects_AA=subjects_A[:]
for j, subject in enumerate(subjects_A):
if ((np.mean(values[subject][b_idx]) > val_thresA) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append(group_vs[0:2])
else :
#print(group_vs[0:2] + " : "+subject+" "+str(np.mean(values[subject][b_idx])))
subjects_AA.remove(subject)
subjects_BB=subjects_B[:]
for j, subject in enumerate(subjects_B):
if ((np.mean(values[subject][b_idx]) > val_thresB) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append(group_vs[-2:])
else :
#print(group_vs[-2:]+ " : "+subject+" "+str(np.mean(values[subject][b_idx])))
subjects_BB.remove(subject)
# Plot
if ind :
ax = sns.lineplot(x="position", y="fa", data=data,markers=True,ax=axes[i], hue="group",units="subject",estimator=None, lw=1) # each subject as single line
else :
ax = sns.lineplot(x="position", y="fa", data=data,markers=True,ax=axes[i], hue="group")
ax.set(xlabel='position along tract', ylabel=para)
ax.set_title(all_bundles[b_idx])
if analysis_type == "correlation" or hide_legend:
ax.legend_.remove()
elif analysis_type == "group" and i > 0:
ax.legend_.remove() # only show legend on first subplot
alpha=0.05
nperm=1000
# Significance testing without multiple correction of bundles
if not correct_mult_tract_comp:
values_allp = [values[s][b_idx] for s in subjects_A + subjects_B] # [subjects, NR_POINTS]
#alphaFWE, clusterFWE = get_corrected_alpha(values_allp, meta_data, analysis_type, subjects_A, subjects_B,alpha, bundles, nperm, b_idx)
alphaFWE, clusterFWE = get_corrected_alpha(values_allp, meta_data, analysis_type, subjects_A, subjects_B,alpha,all_bundles, nperm, b_idx)
# Calc p-values
pvalues = np.zeros(NR_POINTS)
stats = np.zeros(NR_POINTS) # for ttest: t-value, for pearson: correlation
for jdx in range(NR_POINTS):
if analysis_type == "group":
values_controls = [values[s][b_idx][jdx] for s in subjects_AA]
values_patients = [values[s][b_idx][jdx] for s in subjects_BB]
stats[jdx], pvalues[jdx] = scipy.stats.ttest_ind(values_controls, values_patients)
else:
values_controls = [values[s][b_idx][jdx] for s in subjects_A]
stats[jdx], pvalues[jdx] = scipy.stats.pearsonr(values_controls, meta_data["target"].values)
# Plot significant areas
if show_detailed_p:
ax2 = axes[i].twinx()
ax2.bar(range(len(pvalues)), -np.log10(pvalues), color="gray", edgecolor="none", alpha=0.5)
ax2.plot([0, NR_POINTS-1], (-np.log10(alphaFWE),)*2, color="red", linestyle=":")
ax2.set(xlabel='position', ylabel='-log10(p)')
else:
sig_areas = get_significant_areas(pvalues, 1, alphaFWE)
sig_areas = sig_areas * np.quantile(np.array(data["fa"]), 0.98)
sig_areas[sig_areas == 0] = np.quantile(np.array(data["fa"]), 0.02)
axes[i].plot(range(len(sig_areas)), sig_areas, color="red", linestyle=":")
sig_areas2 = get_significant_areas(pvalues, clusterFWE, alpha)
sig_areas2 = sig_areas2 * np.quantile(np.array(data["fa"]), 0.98)
sig_areas2[sig_areas2 == 0] = np.quantile(np.array(data["fa"]), 0.02)
axes[i].plot(range(len(sig_areas2)), sig_areas2, color="green", linestyle="-")
if np.any(pvalues<alphaFWE):
#print(pvalues)
print(all_bundles[b_idx])
print(para)
print(len(subjects_A)+len(subjects_B))
print(len(subjects_A))
print(len(subjects_B))
print(len(subjects_AA)+len(subjects_BB))
print(len(subjects_AA))
print(len(subjects_BB))
# Plot text
axes[i].annotate("alphaFWE: {}".format(format_number(alphaFWE)),
(0, 0), (0, -45), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
axes[i].annotate("min p-value: {}".format(format_number(pvalues.min())),
(0, 0), (0, -65), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
axes[i].annotate("clusterFWE: {}".format(clusterFWE),
(0, 0), (0, -55), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
STR="n : "+str(len(subjects_AA)+len(subjects_BB))+"/"+str(len(subjects_AA))+"/"+str(len(subjects_BB))
axes[i].annotate(STR,
(0, 0), (0, -75), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
stats_label = "t-value: " if analysis_type == "group" else "corr.coeff.: "
#axes[i].annotate(stats_label + " {}".format(format_number(stats[pvalues.argmin()])),
# (0, 0), (0, -55), xycoords='axes fraction', textcoords='offset points', va='top',
# fontsize=10)
if plot_3D_type != "none":
print(plot_3D_type)
if plot_3D_type == "metric":
metric = np.array([values[s][b_idx] for s in subjects_A + subjects_B]).mean(axis=0)
else:
#metric = pvalues # use this code if you want to plot the pvalues instead of the FA
metric = sig_areas
#bundle = bundles[b_idx]
bundle = all_bundles[b_idx]
output_path_3D = output_path.split(".")[0] +"/"+bundle+"_"+para+"_"+group_vs+"_2std_c._3D.png"
if tracking_dir == "auto":
tracking_dir = tracking.get_tracking_folder_name("fixed_prob", False)
if tracking_format == "tck":
tracking_path = join(plot_3D_path, tracking_dir, bundle + ".tck")
else:
tracking_path = join(plot_3D_path, tracking_dir, bundle + ".trk")
ending_path = join(plot_3D_path, "endings_segmentations", bundle + "_b.nii.gz")
mask_path = join(plot_3D_path, "nodif_brain_mask.nii.gz")
if not os.path.isfile(tracking_path):
raise ValueError("Could not find: " + tracking_path)
if not os.path.isfile(ending_path):
raise ValueError("Could not find: " + ending_path)
if not os.path.isfile(mask_path):
raise ValueError("Could not find: " + mask_path)
print(tracking_path)
print(ending_path)
print(mask_path)
print(bundle)
print(metric)
print(output_path_3D)
print(tracking_format)
print(show_color_bar)
plot_utils.plot_bundles_with_metric(tracking_path, ending_path, mask_path, bundle, metric,
output_path_3D, tracking_format, show_color_bar)
plt.tight_layout()
plt.savefig("/mnt/d//LINUX/CogPhenoPark/"+para+"_"+group_vs+"_ind_"+str(ind)+'_p_'+str(show_detailed_p)+"_cofound_"+str(withCofound)+"_2std_cc_mtc.png", dpi=200)
plt.close ('all')
# +
plot_3D_type="pval"
tracking_format="tck"
tracking_dir="FOD_iFOD2_trackings"
output_path="/mnt/d//LINUX/CogPhenoPark"
print(plot_3D_path)
if plot_3D_type != "none":
if plot_3D_type == "metric":
metric = np.array([values[s][b_idx] for s in subjects_A + subjects_B]).mean(axis=0)
else:
# metric = pvalues # use this code if you want to plot the pvalues instead of the FA
metric = sig_areas
#bundle = bundles[b_idx]
bundle = all_bundles[b_idx]
output_path_3D = output_path.split(".")[0] + "_" + bundle + "_3D.png"
if tracking_dir == "auto":
tracking_dir = tracking.get_tracking_folder_name("fixed_prob", False)
if tracking_format == "tck":
tracking_path = join(plot_3D_path, tracking_dir, bundle + ".tck")
else:
tracking_path = join(plot_3D_path, tracking_dir, bundle + ".trk")
ending_path = join(plot_3D_path, "endings_segmentations", bundle + "_b.nii.gz")
mask_path = join(plot_3D_path, "nodif_brain_mask.nii.gz")
if not os.path.isfile(tracking_path):
raise ValueError("Could not find: " + tracking_path)
if not os.path.isfile(ending_path):
raise ValueError("Could not find: " + ending_path)
if not os.path.isfile(mask_path):
raise ValueError("Could not find: " + mask_path)
plot_utils.plot_bundles_with_metric(tracking_path, ending_path, mask_path, bundle, metric,
output_path_3D, tracking_format, show_color_bar)
# -
output_path_3D
# # ANOVA
for para in (['''FA''','''RD''','''MD''','''AD''','''density''']):
for group_vs in ([''' ''']):
###############
FWE_method = "alphaFWE"
show_detailed_p = False
hide_legend = False
show_color_bar = True # colorbar on 3D plot
nperm = 5000
alpha=0.05
correct_mult_tract_comp = False
base_path, meta_data, selected_bundles, plot_3D_path = parse_subjects_file("/mnt/d//LINUX/CogPhenoPark/dataTractSeg/Tractometry_template.txt")
analysis_type = "group"
plot_3D_type="none"
###########
all_bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
values = {}
for subject in meta_data["subject_id"]:
raw = np.loadtxt(base_path.replace("SUBJECT_ID", subject).replace("PARA",para), delimiter=";", skiprows=1).transpose()
values[subject] = raw
NR_POINTS = values[meta_data["subject_id"][0]].shape[1]
#selected_bun_indices = [bundles.index(b) for b in selected_bundles]
selected_bun_indices = [all_bundles.index(b) for b in selected_bundles]
############
confound_names = list(meta_data.columns[2:])
cols = 3
rows = math.ceil(len(selected_bundles) / cols)
a4_dims = (cols*3, rows*5)
f, axes = plt.subplots(rows, cols, figsize=a4_dims)
axes = axes.flatten()
sns.set(font_scale=1.2)
sns.set_style("whitegrid")
subjects_A = list(meta_data[meta_data["group"] == 0]["subject_id"])
subjects_B = list(meta_data[meta_data["group"] == 1]["subject_id"])
subjects_C = list(meta_data[meta_data["group"] == 2]["subject_id"])
subjects_D = list(meta_data[meta_data["group"] == 3]["subject_id"])
# Correct for confounds
values = correct_for_confounds(values, meta_data, all_bundles, selected_bun_indices, NR_POINTS, analysis_type,confound_names)
for i, b_idx in enumerate(tqdm(selected_bun_indices)):
# print(all_bundles[b_idx])
############
vals_thresA=np.zeros([])
subjects_AA=subjects_A[:]
for subject in subjects_A:
if np.all(values[subject][b_idx]>0) :
vals_thresA=np.append(vals_thresA,np.mean(values[subject][b_idx]))
vals_thresA=vals_thresA[1:]
vals_thresA = vals_thresA[~ np.isnan(vals_thresA)]
val_thresA=np.mean(vals_thresA)-2*np.std(vals_thresA)
if val_thresA < 0 : val_thresA = 0
#print("valeur seuil G0= "+str(val_thresA))
vals_thresB=np.zeros([])
subjects_BB=subjects_B[:]
for j, subject in enumerate(subjects_B):
if np.all(values[subject][b_idx]>0) :
vals_thresB=np.append(vals_thresB,np.mean(values[subject][b_idx]))
vals_thresB=vals_thresB[1:]
vals_thresB = vals_thresB[~ np.isnan(vals_thresB)]
val_thresB=np.mean(vals_thresB)-2*np.std(vals_thresB)
if val_thresB < 0 : val_thresB = 0
#print("valeur seuil G1= "+str(val_thresB))
vals_thresC=np.zeros([])
subjects_CC=subjects_C[:]
for j, subject in enumerate(subjects_C):
if np.all(values[subject][b_idx]>0) :
vals_thresC=np.append(vals_thresC,np.mean(values[subject][b_idx]))
vals_thresC=vals_thresC[1:]
vals_thresC = vals_thresC[~ np.isnan(vals_thresC)]
val_thresC=np.mean(vals_thresC)-2*np.std(vals_thresC)
if val_thresC < 0 : val_thresC = 0
#print("valeur seuil G2= "+str(val_thresC))
vals_thresD=np.zeros([])
subjects_DD=subjects_D[:]
for j, subject in enumerate(subjects_D):
if np.all(values[subject][b_idx]>0) :
vals_thresD=np.append(vals_thresD,np.mean(values[subject][b_idx]))
vals_thresD=vals_thresD[1:]
vals_thresD = vals_thresD[~ np.isnan(vals_thresD)]
val_thresD=np.mean(vals_thresD)-2*np.std(vals_thresD)
if val_thresD < 0 : val_thresD = 0
#print("valeur seuil G3= "+str(val_thresD))
# Bring data into right format for seaborn
data = {"position": [],
"fa": [],
"group": [],
"subject": []}
for j, subject in enumerate(subjects_A):
if ((np.mean(values[subject][b_idx]) > val_thresA) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G1")
else :
subjects_AA.remove(subject)
for j, subject in enumerate(subjects_B):
if ((np.mean(values[subject][b_idx]) > val_thresB) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G2")
else :
subjects_BB.remove(subject)
for j, subject in enumerate(subjects_C):
if ((np.mean(values[subject][b_idx]) > val_thresC) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G3")
else :
subjects_CC.remove(subject)
for j, subject in enumerate(subjects_D):
if ((np.mean(values[subject][b_idx]) > val_thresD) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G4")
else :
subjects_DD.remove(subject)
# Plot
ax = sns.lineplot(x="position", y="fa", data=data, ax=axes[i], hue="group")
# units="subject", estimator=None, lw=1) # each subject as single line
print(all_bundles[b_idx])
ax.set(xlabel='position along tract', ylabel='metric')
ax.set_title(all_bundles[b_idx])
if analysis_type == "correlation" or hide_legend:
ax.legend_.remove()
elif analysis_type == "group" and i > 0:
ax.legend_.remove() # only show legend on first subplot
alpha=0.05
nperm=1000
# Significance testing without multiple correction of bundles
if not correct_mult_tract_comp:
values_allp = [values[s][b_idx] for s in subjects_AA + subjects_BB + subjects_CC + subjects_DD ] # [subjects, NR_POINTS]
#alphaFWE, clusterFWE = get_corrected_alpha(values_allp, meta_data, analysis_type, subjects_AA, subjects_BB,alpha,all_bundles, nperm, b_idx)
clusterFWE=10
alphaFWE=0.05
# Calc p-values
pvalues = np.zeros(NR_POINTS)
stats = np.zeros(NR_POINTS) # for ttest: t-value, for pearson: correlation
for jdx in range(NR_POINTS):
if analysis_type == "group":
values_AA = [values[s][b_idx][jdx] for s in subjects_AA]
values_BB = [values[s][b_idx][jdx] for s in subjects_BB]
values_CC = [values[s][b_idx][jdx] for s in subjects_CC]
values_DD = [values[s][b_idx][jdx] for s in subjects_DD]
#stats[jdx], pvalues[jdx] = scipy.stats.kruskal(values_A, values_B,values_C, values_D)
stats[jdx], pvalues[jdx]=scipy.stats.f_oneway(values_AA, values_BB, values_CC, values_DD)
else:
values_controls = [values[s][b_idx][jdx] for s in subjects_A]
stats[jdx], pvalues[jdx] = scipy.stats.pearsonr(values_controls, meta_data["target"].values)
if np.any(pvalues<alphaFWE):
print(all_bundles[b_idx])
print(para)
print(len(subjects_A)+len(subjects_B))
print(len(subjects_A))
print(len(subjects_B))
print(len(subjects_AA)+len(subjects_BB))
print(len(subjects_AA))
print(len(subjects_BB))
# Plot significant areas
if show_detailed_p:
ax2 = axes[i].twinx()
ax2.bar(range(len(pvalues)), -np.log10(pvalues), color="gray", edgecolor="none", alpha=0.5)
ax2.plot([0, NR_POINTS-1], (-np.log10(alphaFWE),)*2, color="red", linestyle=":")
ax2.set(xlabel='position', ylabel='-log10(p)')
else:
sig_areas = get_significant_areas(pvalues, 1, alphaFWE)
sig_areas = sig_areas * np.quantile(np.array(data["fa"]), 0.98)
sig_areas[sig_areas == 0] = np.quantile(np.array(data["fa"]), 0.02)
axes[i].plot(range(len(sig_areas)), sig_areas, color="red", linestyle=":")
sig_areas2 = get_significant_areas(pvalues, clusterFWE, alpha)
sig_areas2 = sig_areas2 * np.quantile(np.array(data["fa"]), 0.98)
sig_areas2[sig_areas2 == 0] = np.quantile(np.array(data["fa"]), 0.02)
axes[i].plot(range(len(sig_areas2)), sig_areas2, color="green", linestyle="-")
# Plot text
if FWE_method == "alphaFWE":
#axes[i].annotate("alphaFWE: {}".format(format_number(alphaFWE)),
# (0, 0), (0, -35), xycoords='axes fraction', textcoords='offset points', va='top',
# fontsize=10)
axes[i].annotate("min p-value: {}".format(format_number(pvalues.min())),
(0, 0), (0, -45), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
else:
axes[i].annotate("clusterFWE: {}".format(clusterFWE),
(0, 0), (0, -35), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
STR=str(len(subjects_AA)+len(subjects_BB)+len(subjects_CC)+len(subjects_DD))+"/"+str(len(subjects_AA))+"/"+str(len(subjects_BB))+"/"+str(len(subjects_CC))+"/"+str(len(subjects_DD))
axes[i].annotate(STR,
(0, 0), (0, -55), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
stats_label = "t-value: " if analysis_type == "group" else "corr.coeff.: "
#axes[i].annotate(stats_label + " {}".format(format_number(stats[pvalues.argmin()])),
# (0, 0), (0, -55), xycoords='axes fraction', textcoords='offset points', va='top',
# fontsize=10)
plt.tight_layout()
plt.savefig("/mnt/d//LINUX/CogPhenoPark/"+para+"_groups_2std_ANOVA_cc.png", dpi=200)
sig_areas2 = get_significant_areas(pvalues, clusterFWE, alpha)
sig_areas2
# # Boxplot
values = {}
len(values)
for para in (['''MD''','''FA''','''RD''','''AD''','''density''']):
for group_vs in ([''' ''']):
###############
FWE_method = "alphaFWE"
show_detailed_p = False
hide_legend = False
show_color_bar = True # colorbar on 3D plot
nperm = 5000
alpha=0.05
correct_mult_tract_comp = False
base_path, meta_data, selected_bundles, plot_3D_path = parse_subjects_file("/mnt/d//LINUX/CogPhenoPark/dataTractSeg/Tractometry_template.txt")
analysis_type = "group"
plot_3D_type="none"
###########
#all_bundles = dataset_specific_utils.get_bundle_names("All_tractometry")[1:]
all_bundles = dataset_specific_utils.get_bundle_names("All")[1:]
values = {}
for subject in meta_data["subject_id"]:
raw = np.loadtxt(base_path.replace("SUBJECT_ID", subject).replace("PARA",para), delimiter=";", skiprows=1).transpose()
values[subject] = raw
NR_POINTS = values[meta_data["subject_id"][0]].shape[1]
#selected_bun_indices = [bundles.index(b) for b in selected_bundles]
selected_bun_indices = [all_bundles.index(b) for b in selected_bundles]
confound_names = list(meta_data.columns[2:])
cols = 5
rows = math.ceil(len(selected_bundles) / cols)
a4_dims = (cols*3, rows*7)
f, axes = plt.subplots(rows, cols, figsize=(a4_dims))
axes = axes.flatten()
sns.set(font_scale=1.2)
sns.set_style("whitegrid")
subjects_A = list(meta_data[meta_data["group"] == 0]["subject_id"])
subjects_B = list(meta_data[meta_data["group"] == 1]["subject_id"])
subjects_C = list(meta_data[meta_data["group"] == 2]["subject_id"])
subjects_D = list(meta_data[meta_data["group"] == 3]["subject_id"])
# Correct for confounds
values = correct_for_confounds(values, meta_data, all_bundles, selected_bun_indices, NR_POINTS, analysis_type,confound_names)
print(len(subjects_A))
print(len(subjects_B))
print(len(subjects_C))
print(len(subjects_D))
cpt=0
for i,b_idx in enumerate(selected_bun_indices):
# print(all_bundles[b_idx])
############
vals_thresA=np.zeros([])
subjects_AA=subjects_A[:]
for subject in subjects_A:
if np.all(values[subject][b_idx]>0) :
vals_thresA=np.append(vals_thresA,np.mean(values[subject][b_idx]))
vals_thresA=vals_thresA[1:]
vals_thresA = vals_thresA[~ np.isnan(vals_thresA)]
val_thresA=np.mean(vals_thresA)-2*np.std(vals_thresA)
if val_thresA < 0 : val_thresA = 0
#print("valeur seuil G0= "+str(val_thresA))
vals_thresB=np.zeros([])
subjects_BB=subjects_B[:]
for j, subject in enumerate(subjects_B):
if np.all(values[subject][b_idx]>0) :
vals_thresB=np.append(vals_thresB,np.mean(values[subject][b_idx]))
vals_thresB=vals_thresB[1:]
vals_thresB = vals_thresB[~ np.isnan(vals_thresB)]
val_thresB=np.mean(vals_thresB)-2*np.std(vals_thresB)
if val_thresB < 0 : val_thresB = 0
#print("valeur seuil G1= "+str(val_thresB))
vals_thresC=np.zeros([])
subjects_CC=subjects_C[:]
for j, subject in enumerate(subjects_C):
if np.all(values[subject][b_idx]>0) :
vals_thresC=np.append(vals_thresC,np.mean(values[subject][b_idx]))
vals_thresC=vals_thresC[1:]
vals_thresC = vals_thresC[~ np.isnan(vals_thresC)]
val_thresC=np.mean(vals_thresC)-2*np.std(vals_thresC)
if val_thresC < 0 : val_thresC = 0
#print("valeur seuil G2= "+str(val_thresC))
vals_thresD=np.zeros([])
subjects_DD=subjects_D[:]
for j, subject in enumerate(subjects_D):
if np.all(values[subject][b_idx]>0) :
vals_thresD=np.append(vals_thresD,np.mean(values[subject][b_idx]))
vals_thresD=vals_thresD[1:]
vals_thresD = vals_thresD[~ np.isnan(vals_thresD)]
val_thresD=np.mean(vals_thresD)-2*np.std(vals_thresD)
if val_thresD < 0 : val_thresD = 0
#print("valeur seuil G3= "+str(val_thresD))
# Bring data into right format for seaborn
data = {"position": [],
"fa": [],
"group": [],
"subject": []}
for j, subject in enumerate(subjects_A):
if ((np.mean(values[subject][b_idx]) > val_thresA) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G1")
else :
subjects_AA.remove(subject)
for j, subject in enumerate(subjects_B):
if ((np.mean(values[subject][b_idx]) > val_thresB) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G2")
else :
subjects_BB.remove(subject)
for j, subject in enumerate(subjects_C):
if ((np.mean(values[subject][b_idx]) > val_thresC) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G3")
else :
subjects_CC.remove(subject)
for j, subject in enumerate(subjects_D):
if ((np.mean(values[subject][b_idx]) > val_thresD) & (np.all(values[subject][b_idx]>0))) :
for position in range(NR_POINTS):
data["position"].append(position)
data["subject"].append(subject)
data["fa"].append(values[subject][b_idx][position])
data["group"].append("G4")
else :
subjects_DD.remove(subject)
values_AA=vals_thresA[vals_thresA>val_thresA]
values_BB=vals_thresB[vals_thresB>val_thresB]
values_CC=vals_thresC[vals_thresC>val_thresC]
values_DD=vals_thresD[vals_thresD>val_thresD]
stat_val,p_val=scipy.stats.f_oneway(vals_thresA[vals_thresA>val_thresA], vals_thresB[vals_thresB>val_thresB], vals_thresC[vals_thresC>val_thresC], vals_thresD[vals_thresD>val_thresD])
# Plot
if p_val < 0.05 :
ax = sns.violinplot(x="group", y="fa", data=data,ax=axes[cpt],inner="point")
cpt=cpt+1
ax.set_title(all_bundles[b_idx])
axes[cpt].annotate("p-value: {}".format(format_number(p_val)),
(0, 0), (0, -25), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
STR=str(len(subjects_AA)+len(subjects_BB)+len(subjects_CC)+len(subjects_DD))+"/"+str(len(subjects_AA))+"/"+str(len(subjects_BB))+"/"+str(len(subjects_CC))+"/"+str(len(subjects_DD))
axes[cpt].annotate(STR,
(0, 0), (0, -35), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
print(p_val)
print(all_bundles[b_idx])
print(para)
print(len(subjects_AA)+len(subjects_BB)+len(subjects_CC)+len(subjects_DD))
print(len(subjects_AA))
print(len(subjects_BB))
print(len(subjects_CC))
print(len(subjects_DD))
STR="two_samples_ttest : "
stat, pvalue = scipy.stats.ttest_ind(values_AA, values_BB)
if (pvalue<0.05):
STR=STR+" G1 Vs G2 : "+ str(format_number(pvalue))
stat, pvalue = scipy.stats.ttest_ind(values_AA, values_CC)
if (pvalue<0.05):
STR=STR+" G1 Vs G3 : "+ str(format_number(pvalue))
stat, pvalue = scipy.stats.ttest_ind(values_AA, values_DD)
if (pvalue<0.05):
STR=STR+" G1 Vs G4 : "+ str(format_number(pvalue))
stat, pvalue = scipy.stats.ttest_ind(values_BB, values_CC)
if (pvalue<0.05):
STR=STR+" G2 Vs G3 : "+ str(format_number(pvalue))
stat, pvalue = scipy.stats.ttest_ind(values_BB, values_DD)
if (pvalue<0.05):
STR=STR+" G2 Vs G4 : "+ str(format_number(pvalue))
stat, pvalue = scipy.stats.ttest_ind(values_CC, values_DD)
if (pvalue<0.05):
STR=STR+" G3 Vs G4 : "+ str(format_number(pvalue))
axes[cpt].annotate(STR,
(0, 0), (0, -45), xycoords='axes fraction', textcoords='offset points', va='top',
fontsize=10)
print(STR)
plt.tight_layout()
plt.show()
plt.savefig("/mnt/d//LINUX/CogPhenoPark/"+para+"_violinplot_2std_72.png", dpi=200)
plt.close()
print(len(subjects_A))
print(len(vals_thresA))
print(len(subjects_B))
print(len(vals_thresB))
print(len(subjects_C))
print(len(vals_thresC))
print(len(subjects_D))
print(len(vals_thresD))
meta_data
vals_thresD[vals_thresD>val_thresD]
print(subjects_B)
print(vals_thresB)
print(type(subjects_B))
print(type(vals_thresB))
print(len(subjects_B))
print(len(vals_thresB))
for j, subject in enumerate(subjects_B):
if np.all(values[subject][b_idx]>0) :
print(subject)
print(np.mean(values[subject][b_idx]))
#vals_thresD=np.append(vals_thresD,)
plot_tractometry_with_pvalue(values, meta_data, all_bundles,selected_bundles,"/NAS/dumbo/protocoles/CogPhenoPark/",
0.05, FWE_method, analysis_type, correct_mult_tract_comp,
show_detailed_p, nperm=5000, hide_legend=False,
plot_3D_path=plot_3D_path, plot_3D_type="pval",
tracking_format="tck", tracking_dir="auto",
show_color_bar=show_color_bar)
def t_stat(y, X, c):
""" betas, t statistic and significance test given data, design matrix, contrast
This is OLS estimation; we assume the errors to have independent
and identical normal distributions around zero for each $i$ in
$\e_i$ (i.i.d).
"""
# Make sure y, X, c are all arrays
y = np.asarray(y)
X = np.asarray(X)
c = np.atleast_2d(c).T # As column vector
# Calculate the parameters - b hat
beta = npl.pinv(X).dot(y)
# The fitted values - y hat
fitted = X.dot(beta)
# Residual error
errors = y - fitted
# Residual sum of squares
RSS = (errors**2).sum(axis=0)
# Degrees of freedom is the number of observations n minus the number
# of independent regressors we have used. If all the regressor
# columns in X are independent then the (matrix rank of X) == p
# (where p the number of columns in X). If there is one column that
# can be expressed as a linear sum of the other columns then
# (matrix rank of X) will be p - 1 - and so on.
df = X.shape[0] - npl.matrix_rank(X)
# Mean residual sum of squares
MRSS = RSS / df
# calculate bottom half of t statistic
SE = np.sqrt(MRSS * c.T.dot(npl.pinv(X.T.dot(X)).dot(c)))
t = c.T.dot(beta) / SE
# Get p value for t value using cumulative density dunction
# (CDF) of t distribution
ltp = t_dist.cdf(t, df) # lower tail p
p = 1 - ltp # upper tail p
return beta, t, df, p
# +
from openpyxl import load_workbook
import glob as glob
from shutil import copyfile
import pandas as pd
file_names=glob.glob('/mnt/d/LINUX/CogPhenoPark/dataTractSeg/CogPhenoPark_csvTractSeg_72/*_L2_tractometry_72.csv')
for file_nameL2 in file_names :
file_nameL3 = file_nameL2.replace("_L2_tractometry_72.csv","_L3_tractometry_72.csv")
print(file_nameL2)
df_L2 = pd.read_csv(file_nameL2,header=0,index_col=False,sep = ';')
df_L3 = pd.read_csv(file_nameL3,header=0,index_col=False,sep = ';')
file_nameRD = file_nameL2.replace("_L2_tractometry_72.csv","_RD_tractometry_72.csv")
RD=(df_L2.values+df_L3.values)/2
df = pd.DataFrame(RD)
print(df_L2.columns)
df.to_csv(file_nameRD,sep = ';',header=list(df_L2.columns),index=False)
#name1=file_name.replace("/mnt/d/LINUX/CogPhenoPark/dataTractSeg/ind_stats/","").replace(".csv","")
#name2=file_nameR.replace("/mnt/d/LINUX/CogPhenoPark/dataTractSeg/ind_stats/","").replace(".csv","")
#wb1 = load_workbook(filename = file_name)
#wb2 = load_workbook(filename = file_nameR)
#sheet_ranges1 = wb1[name1]
#sheet_ranges2 = wb1[name2]
#print(sheet_ranges1["A2"].value)
#print(sheet_ranges2["A2"].value)
#copyfile(file_name,file_nameR)
# +
from openpyxl import load_workbook
import glob as glob
from shutil import copyfile
import pandas as pd
file_names=glob.glob('/mnt/d/LINUX/CogPhenoPark/dataTractSeg/ind_stats/*.csv')
for file_nameL2 in file_names :
df_L2 = pd.read_csv(file_nameL2,index_col = 0,sep = ';')
if (np.any(df_L2.values<0)) :
print(file_nameL2)
# -
import pandas as pd
df_L2 = pd.read_csv("/mnt/d/LINUX/CogPhenoPark/dataTractSeg/nbTrack_t.csv",index_col = 0,sep = ';')
boxplot = df_L2.boxplot(column=["AF_left"])
plt.show()
df_L2.describe()
fig, axes = plt.subplots()
sns.violinplot('AF_left',df_L2)
plt.show()
# +
file_names=glob.glob('/mnt/d/LINUX/CogPhenoPark/dataTractSeg/CogPhenoPark_tracto_MNI/w_AF_left.*.nii.gz')
cpt=0
restot=np.zeros((122,182, 218, 182))
for i,file_name in enumerate(file_names) :
img = nib.load(file_name)
restot[i,:,:,:] = img.get_fdata()
test=np.mean(restot,axis=0)
# +
img = nib.Nifti1Image(test, np.eye(4))
img.get_data_dtype() == np.dtype(np.int16)
nib.save(img, '/mnt/d/LINUX/CogPhenoPark/dataTractSeg/w_AF_left_mean.nii.gz')
# -
def mutual_information(hgram):
""" Mutual information for joint histogram
"""
# Convert bins counts to probability values
pxy = hgram / float(np.sum(hgram))
px = np.sum(pxy, axis=1) # marginal for x over y
py = np.sum(pxy, axis=0) # marginal for y over x
px_py = px[:, None] * py[None, :] # Broadcast to multiply marginals
# Now we can do the calculation using the pxy, px_py 2D arrays
nzs = pxy > 0 # Only non-zero pxy values contribute to the sum
return np.sum(pxy[nzs] * np.log(pxy[nzs] / px_py[nzs]))
time()
# +
from scipy.spatial import distance
from sklearn.metrics.cluster import normalized_mutual_info_score
import pandas as pd
df_full = pd.read_csv("/mnt/d/LINUX/CogPhenoPark//dataTractSeg/nbTrack_t.csv",index_col = 0,sep = ';')
df_tractometry=df_L2 = pd.read_csv("/mnt/d/LINUX/CogPhenoPark/dataTractSeg/ind_stats/AD__sub_100269SD100714.csv",sep = ';')
resSimi=np.zeros((129,129,72))
for z,trk in enumerate(df_full.columns[0:-1]):
print(trk)
file_names=glob.glob('/mnt/d/LINUX/CogPhenoPark/dataTractSeg/CogPhenoPark_tracto_MNI/w_'+trk+'.*.nii.gz')
file_names2=glob.glob('/mnt/d/LINUX/CogPhenoPark/dataTractSeg/CogPhenoPark_tracto_MNI/w_'+trk+'.*.nii.gz')
for i,file_name in enumerate(file_names) :
img = nib.load(file_name)
data= img.get_fdata()
print(i)
for j,file_name2 in enumerate(file_names2) :
if ( resSimi[j,i,z] == 0 ) :
img2 = nib.load(file_name2)
data2= img2.get_fdata()
#hist_2d, x_edges, y_edges = np.histogram2d(data.ravel(),data2.ravel(),bins=200)
tmp_val=normalized_mutual_info_score(data.ravel(),data2.ravel())#mutual_information(hist_2d)
#print(file_name)
print(j)
#print(file_name2)
#print(tmp_val)
resSimi[i,j,z]=tmp_val
else :
resSimi[i,j,z] == resSimi[j,i,z]
| CogPhenoPark-TractSeg-plot-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UyiZb68MQl4y"
# # Amostras Sintéticas
# -
import warnings
warnings.simplefilter('ignore')
import IPython.display as ipd
import librosa
# %matplotlib inline
import matplotlib.pyplot as plt
import librosa.display
import numpy as np
print('Onda Quadrada')
ipd.Audio('../_static/audio/quadrada.wav')
x, sr = librosa.load('../_static/audio/quadrada.wav', duration=0.02)
plt.figure(figsize=(14, 10))
librosa.display.waveplot(x, sr=sr, alpha=0.8)
# +
S = librosa.feature.melspectrogram(y=x, sr=sr, n_mels=128,
fmax=8000)
plt.figure(figsize=(14, 5))
S_dB = librosa.power_to_db(S, ref=np.max)
librosa.display.specshow(S_dB, sr=sr, x_axis='time', y_axis='hz')
plt.colorbar(format='%+2.0f dB')
plt.title('Mel-frequency spectrogram')
plt.tight_layout()
plt.show()
# -
print('Onda Senoide')
ipd.Audio('../_static/audio/senoide.wav')
x, sr = librosa.load('../_static/audio/senoide.wav', duration=0.02)
plt.figure(figsize=(14, 10))
librosa.display.waveplot(x, sr=sr, alpha=0.8)
S = librosa.feature.melspectrogram(y=x, sr=sr, n_mels=128,
fmax=8000)
plt.figure(figsize=(14, 5))
S_dB = librosa.power_to_db(S, ref=np.max)
librosa.display.specshow(S_dB, sr=sr, x_axis='time', y_axis='hz')
plt.colorbar(format='%+2.0f dB')
plt.title('Mel-frequency spectrogram')
plt.tight_layout()
plt.show()
print('Onda Triangular')
ipd.Audio('../_static/audio/triangular.wav')
x, sr = librosa.load('../_static/audio/triangular.wav', duration=0.02)
plt.figure(figsize=(14, 10))
librosa.display.waveplot(x, sr=sr, alpha=0.8)
S = librosa.feature.melspectrogram(y=x, sr=sr, n_mels=128,
fmax=8000)
plt.figure(figsize=(14, 5))
S_dB = librosa.power_to_db(S, ref=np.max)
librosa.display.specshow(S_dB, sr=sr, x_axis='time', y_axis='hz')
plt.colorbar(format='%+2.0f dB')
plt.title('Mel-frequency spectrogram')
plt.tight_layout()
plt.show()
| results/artificial_samples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Create TensorFlow model </h1>
#
# This notebook illustrates:
# <ol>
# <li> Creating a model using the high-level Estimator API
# </ol>
# change these to try this notebook out
BUCKET = 'qwiklabs-gcp-fac41c9e0b13d442'
PROJECT = 'qwiklabs-gcp-fac41c9e0b13d442'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# + language="bash"
# if ! gsutil ls | grep -q gs://${BUCKET}/; then
# gsutil mb -l ${REGION} gs://${BUCKET}
# fi
# -
# <h2> Create TensorFlow model using TensorFlow's Estimator API </h2>
# <p>
# First, write an input_fn to read the data.
# <p>
#
# ## Lab Task 1
# Verify that the headers match your CSV output
import shutil
import numpy as np
import tensorflow as tf
# +
# Determine CSV, label, and key columns
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,plurality,gestation_weeks,key'.split(',')
LABEL_COLUMN = 'weight_pounds'
KEY_COLUMN = 'key'
# Set default values for each CSV column
DEFAULTS = [[0.0], ['null'], [0.0], ['null'], [0.0], ['nokey']]
TRAIN_STEPS = 1000
# -
# ## Lab Task 2
#
# Fill out the details of the input function below
# Create an input function reading a file using the Dataset API
# Then provide the results to the Estimator API
def read_dataset(filename_pattern, mode, batch_size = 512):
def _input_fn():
def decode_csv(line_of_text):
# TODO #1: Use tf.decode_csv to parse the provided line
columns = tf.decode_csv(line_of_text, record_defaults=DEFAULTS)
# TODO #2: Make a Python dict. The keys are the column names, the values are from the parsed data
features = dict(zip(CSV_COLUMNS, columns))
# TODO #3: Return a tuple of features, label where features is a Python dict and label a float
label = features.pop(LABEL_COLUMN)
return features, label
# TODO #4: Use tf.gfile.Glob to create list of files that match pattern
file_list = tf.gfile.Glob(filename_pattern)
# Create dataset from file list
dataset = (tf.data.TextLineDataset(file_list).map(decode_csv)) # Transform each elem by applying decode_csv fn
# TODO #5: In training mode, shuffle the dataset and repeat indefinitely
# (Look at the API for tf.data.dataset shuffle)
# The mode input variable will be tf.estimator.ModeKeys.TRAIN if in training mode
# Tell the dataset to provide data in batches of batch_size
if mode == tf.estimator.ModeKeys.TRAIN:
epochs = None
dataset = dataset.shuffle(buffer_size = 10 * batch_size)
else:
epochs = 1
dataset = dataset.repeat(epochs).batch(batch_size)
# This will now return batches of features, label
return dataset
return _input_fn
# ## Lab Task 3
#
# Use the TensorFlow feature column API to define appropriate feature columns for your raw features that come from the CSV.
#
# <b> Bonus: </b> Separate your columns into wide columns (categorical, discrete, etc.) and deep columns (numeric, embedding, etc.)
# Define feature columns
def get_wide_deep():
is_male,mother_age,plurality,gestation_weeks = \
[\
tf.feature_column.categorical_column_with_vocabulary_list('is_male',
['True', 'False', 'Unknown']),
tf.feature_column.numeric_column('mother_age'),
tf.feature_column.categorical_column_with_vocabulary_list('plurality',
['Single(1)', 'Twins(2)', 'Triplets(3)',
'Quadruplets(4)', 'Quintuplets(5)','Multiple(2+)']),
tf.feature_column.numeric_column('gestation_weeks')
]
# Discretize
age_buckets = tf.feature_column.bucketized_column(mother_age,
boundaries=np.arange(15,45,1).tolist())
gestation_buckets = tf.feature_column.bucketized_column(gestation_weeks,
boundaries=np.arange(17,47,1).tolist())
# Sparse columns are wide, have a linear relationship with the output
wide = [is_male,
plurality,
age_buckets,
gestation_buckets]
# Feature cross all the wide columns and embed into a lower dimension
crossed = tf.feature_column.crossed_column(wide, hash_bucket_size=20000)
embed = tf.feature_column.embedding_column(crossed, 3)
# Continuous columns are deep, have a complex relationship with the output
deep = [mother_age,
gestation_weeks,
embed]
return wide, deep
# ## Lab Task 4
#
# To predict with the TensorFlow model, we also need a serving input function (we'll use this in a later lab). We will want all the inputs from our user.
#
# Verify and change the column names and types here as appropriate. These should match your CSV_COLUMNS
# Create serving input function to be able to serve predictions later using provided inputs
def serving_input_fn():
feature_placeholders = {
'is_male': tf.placeholder(tf.string, [None]),
'mother_age': tf.placeholder(tf.float32, [None]),
'plurality': tf.placeholder(tf.string, [None]),
'gestation_weeks': tf.placeholder(tf.float32, [None])
}
features = {
key: tf.expand_dims(tensor, -1)
for key, tensor in feature_placeholders.items()
}
return tf.estimator.export.ServingInputReceiver(features, feature_placeholders)
# ## Lab Task 5
#
# Complete the TODOs in this code:
# %ls
# Create estimator to train and evaluate
def train_and_evaluate(output_dir):
wide, deep = get_wide_deep()
EVAL_INTERVAL = 300
run_config = tf.estimator.RunConfig(save_checkpoints_secs = EVAL_INTERVAL,
keep_checkpoint_max = 3)
# TODO #1: Create your estimator
estimator = tf.estimator.DNNLinearCombinedRegressor(
model_dir = output_dir,
linear_feature_columns = wide,
dnn_feature_columns = deep,
dnn_hidden_units = [64, 32],
config = run_config)
train_spec = tf.estimator.TrainSpec(
# TODO #2: Call read_dataset passing in the training CSV file and the appropriate mode
input_fn = read_dataset('train.csv', mode=tf.estimator.ModeKeys.TRAIN),
max_steps = TRAIN_STEPS)
exporter = tf.estimator.LatestExporter('exporter', serving_input_fn)
eval_spec = tf.estimator.EvalSpec(
# TODO #3: Call read_dataset passing in the evaluation CSV file and the appropriate mode
input_fn = read_dataset('eval.csv', mode=tf.estimator.ModeKeys.EVAL),
steps = None,
start_delay_secs = 60, # start evaluating after N seconds
throttle_secs = EVAL_INTERVAL, # evaluate every N seconds
exporters = exporter)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# Finally, train!
# Run the model
shutil.rmtree('babyweight_trained', ignore_errors = True) # start fresh each time
train_and_evaluate('babyweight_trained')
# When I ran it, the final lines of the output (above) were:
# <pre>
# INFO:tensorflow:Saving dict for global step 1000: average_loss = 1.2693067, global_step = 1000, loss = 635.9226
# INFO:tensorflow:Restoring parameters from babyweight_trained/model.ckpt-1000
# INFO:tensorflow:Assets added to graph.
# INFO:tensorflow:No assets to write.
# INFO:tensorflow:SavedModel written to: babyweight_trained/export/exporter/temp-1517899936/saved_model.pb
# </pre>
# The exporter directory contains the final model and the final RMSE (the average_loss) is 1.2693067
# <h2> Monitor and experiment with training </h2>
from google.datalab.ml import TensorBoard
TensorBoard().start('./babyweight_trained')
for pid in TensorBoard.list()['pid']:
TensorBoard().stop(pid)
print('Stopped TensorBoard with pid {}'.format(pid))
# Copyright 2017-2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| week2/3_tensorflow_wd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "14d82f0b-99cd-4117-9a57-3beb5ca12f56", "showTitle": false, "title": ""}
# Instalacija neophodnih modula:
# * __gdown__ - preuzimanje baze podataka sa _Google Drive_ platforme
# * __pymongo__ - konektovanje na udaljenu _MongoDB_ bazu podataka
# * __pymongo-schema__ - ekstrahovanje šeme _MongoDB_ baze podataka
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "96b019fd-75d2-4d46-8b8e-169a02e6f74d", "showTitle": false, "title": ""}
# !pip install gdown
# !pip install 'pymongo[srv]'
# !pip install --upgrade https://github.com/pajachiet/pymongo-schema/archive/master.zip
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1cb0f645-41b8-45a4-ac41-85fced67f9b6", "showTitle": false, "title": ""}
# Preuzimanje baze podataka sa _Google Drive_ platforme.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c18a645e-b3b6-4ccb-ada1-1656ed7bf6a7", "showTitle": false, "title": ""}
# !gdown --id 1A3ILSwvIBbNTEnl5Av2EU450SdBbaNS7
# !unzip database.sqlite.zip
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "65fee49b-8a5a-4d73-b057-649ad23e4520", "showTitle": false, "title": ""}
# Importovanje neophodnih modula.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "42d21d7e-8fe6-4543-9954-4f976ec234ca", "showTitle": false, "title": ""}
import math
import json
import sqlite3
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from time import sleep
from functools import reduce
from pymongo import MongoClient
from itertools import combinations
from pyspark.sql.types import *
from pyspark.sql.functions import *
from pymongo_schema.extract import extract_pymongo_client_schema
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0759261f-edf9-4f07-9e67-0eab5ad15ee6", "showTitle": false, "title": ""}
# Konektovanje na udaljenu _MongoDB_ bazu podataka.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d069fece-e87a-4e23-b5f0-96c9c3b76912", "showTitle": false, "title": ""}
# Definisanje kredencijala za MongoDB bazu podataka
MONGO_URL = 'mongodb+srv://databricks:databricks@vp-vezbe.leebs.mongodb.net/vp-projekat?retryWrites=true&w=majority'
MONGO_DB = 'vp-projekat'
# Konektovanje na udaljenu MongoDB bazu podataka
client = MongoClient(MONGO_URL)
db = client[MONGO_DB]
# Ispisivanje šeme MongoDB baze podataka
schema = extract_pymongo_client_schema(client, database_names=[MONGO_DB])
print(json.dumps(schema, indent=4, sort_keys=True))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "dceb7ca2-c346-446a-be8a-8c9911b6662f", "showTitle": false, "title": ""}
# Konvertovanje _SQLite_ tabela u _MongoDB_ kolekcije uz izbacivanje suvišnih kolona.
#
# * Tabela: __League__
# * Kolona __id__ - jedinstveni identifikator lige
# * Kolona __name__ - naziv lige
#
# * Tabela: __Match__
# * Kolona __id__ - jedinstveni identifikator utakmice
# * Kolona __league_id__ - identifikator lige u kojoj je utakmica odigrana
# * Kolona __season__ - sezona u kojoj je utakmica odigrana
# * Kolona __date__ - datum odigravanje utakmice
# * Kolona __home_team_goal__ - ukupno golova datih od strane domaćeg tima
# * Kolona __away_team_goal__ - ukupno golova datih od strane gostujućeg tima
# * Kolona __B365H__ - kvota u slučaju da domaći tim pobedi
# * Kolona __B365D__ - kvota u slučaju da rezultat bude nerešen
# * Kolona __B365A__ - kvota u slučaju da gostujući tim pobedi
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c472376c-49b4-43b4-baf6-18dccfefd75d", "showTitle": false, "title": ""}
# Konektovanje na lokalnu SQLite bazu podataka
cnx = sqlite3.connect('database.sqlite')
# Definisanje SQLite tabela i kolona koje je potrebno konvertovati u MongoDB kolekcije
metadata = [
{'table':'League', 'columns':['id', 'name']},
{'table':'Match', 'columns':['id', 'league_id', 'season', 'date', 'home_team_goal', 'away_team_goal', 'B365H', 'B365D', 'B365A']},
{'table':'Team', 'columns':['id', 'team_long_name', 'team_short_name']}
]
# Iteriranje kroz sve tabele/kolekcije
for meta in metadata:
# Proveravanje postojanja i brisanje MongoDB kolekcija sa istim nazivom
if db[meta['table']].count_documents({}) > 0:
db[meta['table']].delete_many({})
# Formiranje stringa za izolovanje odabranih kolona SQLite tabele
table, select = meta['table'], ', '.join(meta['columns'])
# Izolovanje odabranih kolona SQLite tabele i kovertovanje u MongoDB kolekciju
df = pd.read_sql_query(f'SELECT {select} FROM {table}', cnx)
df.reset_index(inplace=True)
records = df.to_dict('records')
db[table].insert_many(records)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b66c2c41-0309-46ee-91e3-35c2ed6d65a5", "showTitle": false, "title": ""}
# Konvertovanje _MongoDB_ kolekcija u _Pandas_ okvire podataka. Tabela __Match__ se sortira po datumu odigravanja utakmice. Izbacuju se utakmice koje nemaju definisane kvote.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "46ca1791-a888-4f0b-b41a-f1f066afb113", "showTitle": false, "title": ""}
# Učitavanje MongoDB kolekcija u Pandas okvire podataka
leagues = pd.DataFrame.from_dict(db['League'].find())
matches = pd.DataFrame.from_dict(db['Match'].find().sort('date'))
# Brisanje utakmica bez definisanih kvota
matches.dropna(inplace=True)
# Ispisivanje šema Pandas okvira podataka
print('--------- Leagues ---------')
print(leagues.infer_objects().dtypes)
print('--------- Matches ---------')
print(matches.infer_objects().dtypes)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ec17c7db-c492-4c42-ab71-56cb6e9eee8c", "showTitle": false, "title": ""}
# Kreira se novi _Pandas_ okvir podataka koji za ključ ima ligu i sezonu, a za vrednost broj utakmica te lige odigranih u toj sezoni.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "85a06e07-452b-4e4d-81b6-ba01ca33d09d", "showTitle": false, "title": ""}
# Prebrojavanje ukupnog broja odigranih utakmica po ligi po sezoni
league_season = pd.DataFrame(matches.groupby(['league_id', 'season'], as_index=False)['_id'].count())
league_season.rename(columns={'_id':'matches_total'}, inplace=True)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fe52a5a8-e7d4-464b-b34d-543b9550a1c6", "showTitle": false, "title": ""}
# Definiše se funkcija __predicted__ za evidentiranje da li se rezultat prosleđene utakmice može predvideti na osnovu kvota. Definiše se funkcija __entropy__ za evidentiranje mere entropije za određenu sezonu određene lige na osnovu broja tačno predviđenih rezultata utakmica.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9a8dc2e8-4dd1-477d-a0a9-22caf9e22f32", "showTitle": false, "title": ""}
def _predicted(x):
# Čuvanje kvota i rezultata utakmice
gh, ga = x['home_team_goal'], x['away_team_goal']
oh, od, oa = x['B365H'], x['B365D'], x['B365A']
rh, rd, ra = gh > ga, gh == ga, gh < ga
omin = np.min([oh, od, oa])
bh, bd, ba = oh == omin, od == omin, oa == omin
# Ispitivanje da li se rezultat utakmice može predvideti na osnovu kvota
return 1 if rh and bh or rd and bd or ra and ba else 0
def _entropy(x):
# Računanje mere entropije kao količnik tačno predviđenih rezultata kroz ukupan broj odigranih utakmica
return x['matches_predicted'] / x['matches_total']
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9b7b20ae-e669-4a46-85ee-7d2d7c3bbd43", "showTitle": false, "title": ""}
# Funkcija __predicted__ se poziva nad svakom utakmicom. Dobijeni rezultati se agregiraju kako bi se dobila mera entropije pozivom funkcije __entropy__ nad svakom sezonom svake lige.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d558e4af-fe37-4149-88d2-92ebbd8bca84", "showTitle": false, "title": ""}
# Prebrojavanje tačno predviđenih rezultata svih utakmica
matches['predicted'] = matches.apply(lambda x: _predicted(x), axis=1)
# Agregiranje tačno predviđenih rezultata po ligi po sezoni
matches_predicted = pd.DataFrame(matches.groupby(['league_id', 'season'])['predicted'].sum())
matches_predicted.rename(columns={'predicted':'matches_predicted'}, inplace=True)
# Brisanje privremene kolone kreirane prilikom prebrojavanja
matches = matches.drop(columns=['predicted'])
# Dodeljivanje broja predviđenih utakmica i mere entropije svakoj ligi svake sezone
league_season = pd.merge(league_season, matches_predicted, on=['league_id', 'season'])
league_season['entropy'] = league_season.apply(lambda x: _entropy(x), axis=1)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3f9eef84-e904-4c2f-a436-2e2aa99b1037", "showTitle": false, "title": ""}
# Definiše se funkcija __simulate__ koja za sve utakmice određene ligu određene sezone simulira klađenje po prosleđenom sistemu sa početnim ulaganjem. Definiše se pomoćna funkcija __comb__ koja računa broj kombinacija sa parametrima __n__ i __k__. Definiše se pomoćna funkcija __bet__ koja evidentira rezultat klađenja, tj. u slučaju da je pogođen rezultat utakmice, vraća vrednost pobedničke kvote koja kasnije služi za računanje dobitka tiketa u kojem se nalazi prosleđena utakmica, inače, vraća nulu što odgovara gubitku uloženog novca.
#
# * Moguće vrednosti promenljive __type__:
# * __h__ - Domaći tim pobeđuje
# * __d__ - Rezultat nerešen
# * __a__ - Gostujući tim pobeđuje
# * __min__ - Klađenje na najmanju kvotu (ishod sa najvećom verovatnoćom)
# * __max__ - Klađenje na najveću kvotu (ishod sa najmanjom verovatnoćom)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "193174c8-c6e4-4aa3-be13-f0c43b0a9bc1", "showTitle": false, "title": ""}
def _simulate(league_id, season, output=False, system=(6, 4), bet_size=1000, type_='min'):
def _comb(n, k):
# Računanje broja kombinacija po standardnoj formuli
nf, kf, nkf = math.factorial(n), math.factorial(k), math.factorial(n-k)
return nf / (kf * nkf);
def _bet(x, t):
# Čuvanje kvota i rezultata utakmice
gh, ga = x['home_team_goal'], x['away_team_goal']
oh, od, oa = x['B365H'], x['B365D'], x['B365A']
odds = {'h':oh, 'd':od, 'a':oa}
rh, rd, ra = gh > ga, gh == ga, gh < ga
# Proveravanje tipa klađenja
if t in 'hda':
# Ispitivanje da li je pogođen rezultat utakmice
return odds[t] if rh and t == 'h' or rd and t == 'd' or ra and t == 'a' else 0
elif t == 'min' or t == 'max':
minmax = np.min(odds.values()) if t == 'min' else np.max(odds.values())
# Ispitivanje da li je pogođen rezultat utakmice
if rh and oh == minmax:
return oh
elif rd and od == minmax:
return od
elif ra and oa == minmax:
return oa
else:
return 0
# Čuvanje prosleđene lige i sezone
season = matches.loc[(matches['league_id'] == league_id) & (matches['season'] == season)]
# Definisanje početnih parametara
balance = 0
balances = []
cash_per_ticket = bet_size / _comb(*system)
# Iteriranje kroz sve utakmice prosleđene lige i sezone
for s in range(0, season.shape[0], system[0]):
# Umanjenje bilansa stanja sa definisani ulog
balance -= bet_size
# Proveravnje rezultata klađenja
result = season[s:s+system[0]].apply(lambda x: _bet(x, type_), axis=1).to_numpy()
# Čuvanje dobitnih tiketa
correct = result[result > 0]
# Proveravanje da li je klađenje po sistemu uspešno
if correct.shape[0] >= system[1]:
# Generisanje svih mogućih tiketa po prosleđenom sistemu
tickets = [x for x in combinations(correct, system[1])]
# Iteriranje kroz sve tikete
for t in tickets:
# Računanje novčanog dobitka
prod = reduce((lambda x, y: x * y), t)
balance += prod * cash_per_ticket
# Čuvanje trenutnog bilansa stanja
balances.append(balance)
# Proveravanje da li je potrebno kreirati JSON datoteke za Apache Spark Structured Streaming
if output:
# Simuliranje računanja radi testiranja Apache Spark Structured Streaming
sleep(1)
# Kreiranje JSON datoteke sa informacija o tiketu i bilansu stanja
path = STREAM_PATH + str(s) + '.json'
dbutils.fs.put(path, "{'ticket':" + str(s) + ", 'balance':" + str(balance) + "}")
# Računanje najmanjeg i najvećeđ bilansa stanja
balances = np.array(balances)
min_balance, max_balance = np.min(balances), np.max(balances)
# Vraćanje završnog, najmanjeg i najvećeg bilansa stanja
return pd.Series([balance, min_balance, max_balance])
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9f7d4442-8dee-4009-bcb9-b8ba20a25bbc", "showTitle": false, "title": ""}
# Definiše se funkcija __simulate_all__ koja sa svaku ligu svake sezone poziva funkciju __simulate__ radi testiranja hipoteze o predidivosti liga. Definiše se pomoćna funkcija __scatter__ za iscrtavanje grafika zavisnosti novčanog dobitka/gubitka od prediktivnosti liga.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fa13d999-e1a1-4fd7-8597-5f45d0fc8c50", "showTitle": false, "title": ""}
def _simulate_all():
def _scatter(data, index, total, cols=4):
# Pozicioniranje grafika
plt.subplot(total // cols, cols, index + 1)
# Iscrtavanje tačaka
plt.scatter(data[:,1], data[:,2])
# Pridruživanje naziva lige svakoj tački na grafiku
for i, ann in enumerate(data[:,0]):
plt.annotate(ann, (data[i,1], data[i,2]))
# Postavljanje dimenzija grafika
plt.figure(figsize=(30, 15))
# Čuvanje jedinstvenih sezona
seasons = league_season['season'].unique()
# Iteriranje kroz sve sezone
for i, season in enumerate(seasons):
# Odabiranje svih liga trenutne sezone
selected_season = league_season.query(f'season == "{season}"')
selected_season.index.name = '_id'
# Simuliranje klađenja po sistemu za svaku sezonu svake lige
predictive_leagues = selected_season.apply(lambda x: _simulate(x['league_id'], season), axis=1)
# Čuvanje završnog, najmanjeg i najvećeg bilansa stanja
predictive_leagues.rename(columns={0:'balance', 1:'min_balance', 2:'max_balance'}, inplace=True)
# Spajanje okvira podataka radi povezivanja konkretne lige sa isplativošću klađenja po sistemu
selected_season = pd.merge(selected_season, predictive_leagues, on='_id')
selected_season = pd.merge(leagues, selected_season, left_on='id', right_on='league_id')
# Iscrtavanje pojedinačnih grafika
data = selected_season[['name', 'entropy', 'balance']].to_numpy()
_scatter(data, i, seasons.shape[0])
# Prikazivanje svih pojedinačnih grafika
plt.show()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c8033a69-c532-4ba2-b744-788a933d45b1", "showTitle": false, "title": ""}
# __Hipoteza 1:__ Predvidivost lige utiče na visinu novčanog dobitka/gubitka.
#
# __Rezultat:__ U opštem slučaju, klađenje po sistemu se ne isplati nezavisno od prediktivnosti lige.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e14e44df-c378-48fd-b3a2-6d544c1f379b", "showTitle": false, "title": ""}
_simulate_all()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fc8d2ce1-dd9a-4fb1-b684-01ec71aea858", "showTitle": false, "title": ""}
# Pokreće se _Apache Spark Structured Streaming_ za uživo praćenje simulacije klađenja po sistemu. Definiše se šema _JSON_ dokumenta koji se kreira u funkciji __simulate__ za svaki tiket odigran po sistemu. Iscrtava se grafik koji pokazuje bilans stanja kroz vreme koje se označava rednim brojem utakmice odabrane sezone u odabranoj ligi.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "763e841c-f550-46c4-b438-68ee2b3c64ca", "showTitle": false, "title": ""}
# Definisanje direktorijuma na kojoj će se čuvati JSON datoteke simulacije
STREAM_PATH = '/test/'
# Brisanje JSON datoteka sačuvanih u prethodnoj simulaciji
dbutils.fs.rm(STREAM_PATH, True)
# Kreiranje direktorijuma na kojoj će se čuvati JSON datoteke trenutne simulacije
dbutils.fs.mkdirs(STREAM_PATH)
# Prikazivanje grafika koji pokazuje promenu bilansa stanja kroz vreme
display(
spark
.readStream
# Pridruživanje šeme JSON datoteka
.schema(StructType([StructField('ticket', LongType(), False), StructField('balance', DecimalType(), False)]))
# Učitavanje JSON datoteka
.json(STREAM_PATH)
)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "065ac315-4cb6-44d2-9cc2-42a2040b3940", "showTitle": false, "title": ""}
# __Hipoteza 2:__ Klađenje po sistemu se isplati.
#
# __Rezultat:__ U opštem slučaju, klađenje po sistemu se ne isplati, ali se može doći do izuzetaka u zavisnosti od parametara sistema i tipa klađenja.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "aeb8d525-34e6-4269-8264-e1c61413bd72", "showTitle": false, "title": ""}
# Definisanje indeksa željene kombinacije lige i sezone
# Klađenje po sistemu se isplati za STREAM_INDEX=6
STREAM_INDEX = 7
# Definisanje željenog sistema klađenja
SIMULATE_SYSTEM = (6, 4)
# Definisanje željenog tipa klađenja
SIMULATE_TYPE = 'h'
# Odabiranje kombinacije lige i sezone
selected = league_season.iloc[STREAM_INDEX]
# Simuliranje klađenja po sistemu za odabranu ligu i sezonu
balances = _simulate(selected['league_id'], selected['season'], output=True, system=SIMULATE_SYSTEM, type_=SIMULATE_TYPE).to_numpy()
# Ispisivanje završnog, najmanjeg i najvećeg bilansa stanja
print('Current balance:', balances[0])
print('Minimum balance:', balances[1])
print('Maximum balance:', balances[2])
| project/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="JwAn1Jr86udO"
# # Introduction to Logistic Regression
#
#
#
# ## Learning Objectives
#
# 1. Create Seaborn plots for Exploratory Data Analysis
# 2. Train a Logistic Regression Model using Scikit-Learn
#
#
# ## Introduction
#
# This lab is an introduction to logistic regression using Python and Scikit-Learn. This lab serves as a foundation for more complex algorithms and machine learning models that you will encounter in the course. In this lab, we will use a synthetic advertising data set, indicating whether or not a particular internet user clicked on an Advertisement on a company website. We will try to create a model that will predict whether or not they will click on an ad based off the features of that user.
#
#
# Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/intro_logistic_regression.ipynb).
#
# -
# ### Import Libraries
# + colab={} colab_type="code" id="8z-hq2Co6udQ"
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + [markdown] colab_type="text" id="XuU2AljQ6udT"
# ### Load the Dataset
#
# We will use a synthetic [advertising](https://www.kaggle.com/fayomi/advertising) dataset. This data set contains the following features:
#
# * 'Daily Time Spent on Site': consumer time on site in minutes
# * 'Age': customer age in years
# * 'Area Income': Avg. Income of geographical area of consumer
# * 'Daily Internet Usage': Avg. minutes a day consumer is on the internet
# * 'Ad Topic Line': Headline of the advertisement
# * 'City': City of consumer
# * 'Male': Whether or not consumer was male
# * 'Country': Country of consumer
# * 'Timestamp': Time at which consumer clicked on Ad or closed window
# * 'Clicked on Ad': 0 or 1 indicated clicking on Ad
# + colab={} colab_type="code" id="MXAy-rgp6udU"
# TODO 1: Read in the advertising.csv file and set it to a data frame called ad_data.
# TODO: Your code goes here
# + [markdown] colab_type="text" id="lqtY61dE6udW"
# **Check the head of ad_data**
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="wS9c7okn6udX" outputId="3be715b4-850a-4ece-f565-9d9ebcc8ad52"
ad_data.head()
# + [markdown] colab_type="text" id="WWCSKrwq6udZ"
# **Use info and describe() on ad_data**
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="0LTHowWE6uda" outputId="b22829e2-f0fa-49db-89c9-2e8c0ae23d7e"
ad_data.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="7exNAo7r6udc" outputId="b6777af0-bfff-4ac9-89ff-8f324b133fe6"
ad_data.describe()
# -
# Let's check for any null values.
ad_data.isnull().sum()
# + [markdown] colab_type="text" id="tNDtwGLU6ude"
# ## Exploratory Data Analysis (EDA)
#
# Let's use seaborn to explore the data! Try recreating the plots shown below!
# -
# TODO 1: **Create a histogram of the Age**
# + colab={"base_uri": "https://localhost:8080/", "height": 296} colab_type="code" id="_cZQWiIa6udf" outputId="5064a339-ccdb-46c6-ea47-d5705185eb49"
# TODO: Your code goes here
# + [markdown] colab_type="text" id="gc8IkCJV6udh"
# TODO 1: **Create a jointplot showing Area Income versus Age.**
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="uoI729Bk6udh" outputId="3907e402-448c-4568-cb7b-8a387ce61cf2"
# TODO: Your code goes here
# + [markdown] colab_type="text" id="y_eq6CJs6udj"
# TODO 2: **Create a jointplot showing the kde distributions of Daily Time spent on site vs. Age.**
# + colab={"base_uri": "https://localhost:8080/", "height": 441} colab_type="code" id="0w695Wda6udk" outputId="956e69b1-a108-48d3-d0a0-20424369c65a"
# TODO: Your code goes here
# + [markdown] colab_type="text" id="tDM8yyQR6udm"
# TODO 1: **Create a jointplot of 'Daily Time Spent on Site' vs. 'Daily Internet Usage'**
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="TygXpR2Y6udm" outputId="e16c0cba-f120-4191-cb2e-5581168d0972"
# TODO: Your code goes here
# + [markdown] colab_type="text" id="AHeR3th76udr"
# # Logistic Regression
#
# Logistic regression is a supervised machine learning process. It is similar to linear regression, but rather than predict a continuous value, we try to estimate probabilities by using a logistic function. Note that even though it has regression in the name, it is for classification.
# While linear regression is acceptable for estimating values, logistic regression is best for predicting the class of an observation
#
# Now it's time to do a train test split, and train our model! You'll have the freedom here to choose columns that you want to train on!
# + colab={} colab_type="code" id="3Z2BXtOt6uds"
from sklearn.model_selection import train_test_split
# -
# Next, let's define the features and label. Briefly, feature is input; label is output. This applies to both classification and regression problems.
# + colab={} colab_type="code" id="Kjlnug3W6udt"
X = ad_data[['Daily Time Spent on Site', 'Age', 'Area Income','Daily Internet Usage', 'Male']]
y = ad_data['Clicked on Ad']
# -
# TODO 2: **Split the data into training set and testing set using train_test_split**
# + colab={} colab_type="code" id="WtK2DMqd6udv"
# TODO: Your code goes here
# + [markdown] colab_type="text" id="Hh-cXKCb6udy"
# **Train and fit a logistic regression model on the training set.**
# + colab={} colab_type="code" id="0PnZOrge6udy"
from sklearn.linear_model import LogisticRegression
# + colab={"base_uri": "https://localhost:8080/", "height": 156} colab_type="code" id="XhqwY1ZG6ud0" outputId="6688f1e4-4ade-4b8c-d0e2-90a9fb662cbe"
logmodel = LogisticRegression()
logmodel.fit(X_train,y_train)
# + [markdown] colab_type="text" id="QV3-c2qF6ud2"
# ## Predictions and Evaluations
# **Now predict values for the testing data.**
# + colab={} colab_type="code" id="UA72s_Vt6ud2"
predictions = logmodel.predict(X_test)
# + [markdown] colab_type="text" id="vYpC2Yc46ud4"
# **Create a classification report for the model.**
# + colab={} colab_type="code" id="dFD-TA186ud5"
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="NrQ6CswF6ud8" outputId="6725c756-4394-4b14-821e-205c6775e8fb"
print(classification_report(y_test,predictions))
# + [markdown] colab_type="text" id="1-ruS5jy6ud9"
# Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| courses/machine_learning/deepdive2/launching_into_ml/labs/intro_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sazio/GAMELEON/blob/master/Analysis/Curfew_Processing_Beta0.1_1k.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="S7csjosew-lk"
import numpy as np
import pandas as pd # data handling
from tqdm.notebook import tqdm # measuring for loops runtime
import matplotlib.pyplot as plt # plot
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# + id="52Pgfx-Iw_U1"
#Import data from GDrive - if you run this notebook in your local machine mute this cell and change path below
from google.colab import drive
drive.mount('/content/drive', force_remount= True)
# + id="GnKR6ECZw-lt" colab={"referenced_widgets": ["e627377f31a8427f954b134694320b40"]} outputId="2a4cf886-5fdc-4113-8ffb-ae445d557696"
path = "drive/MyDrive/MAS/Data/Output_GAMA/Data/People/Restrictions_Beta0.1_1k/" # path to folder & files
num_people = 1020
curfew_time = [18,19,20] # hour when the curfew starts
curfew_delay = [5, 10] #days after the start of the simulation for curfew
people_files = []
for i in tqdm(range(0,num_people)):
people_files.append(path + "people" + str(i) + ".txt")
# + id="vNpEwrUww-lv"
def preprocess_people(people_file):
DF_people = pd.read_csv(people_file, sep = ",", names = ["cycle","beta","curfew_time", "curfew_delay", "working_place","living_place","is_infected","is_immune","is_dead"])
DF_people = DF_people[DF_people["cycle"] != "cycle"]
DF_people = DF_people.reset_index(drop = True)
DF_people["cycle"] = DF_people["cycle"].astype(np.int16) # dtype conversion
DF_people["beta"] = DF_people["beta"].astype(np.float32)
DF_people["curfew_time"] = DF_people["curfew_time"].astype(np.int16)
DF_people["curfew_delay"] = DF_people["curfew_delay"].astype(np.int16)
DF_people = DF_people.replace({"is_infected":{"true":1, "false":0}})
DF_people = DF_people.replace({"is_immune":{"true":1, "false":0}})
DF_people = DF_people.replace({"is_dead":{"true":1, "false":0}})
DF_people["is_susceptible"] = 1 - (DF_people.is_infected + DF_people.is_immune + DF_people.is_dead)
mean_ = DF_people.groupby(['curfew_time', 'curfew_delay', 'cycle'])['is_susceptible', 'is_immune', 'is_infected', 'is_dead'].mean()
var_ = DF_people.groupby(['curfew_time', 'curfew_delay', 'cycle'])['is_susceptible', 'is_immune', 'is_infected', 'is_dead'].var()
return mean_, var_
# + id="OjQoK4odw-lv" colab={"referenced_widgets": ["544254d72675450b8a2964d02cfed94c"]} outputId="17179e9d-d319-49ce-f28a-dd900d88bb4a"
mean_0, var_0 = preprocess_people(people_files[0])
mean = mean_0
var = var_0
for file in tqdm(people_files[1:], total = len(people_files)-1):
pivot_mean, pivot_var = preprocess_people(file)
mean += pivot_mean
var += pivot_var
# + id="jnBLm_mTw-lw"
n_batch = 30
aggr_people = mean.reset_index()
aggr_people["days"] = aggr_people["cycle"]/24
aggr_people_var = var.reset_index()
aggr_people_var["days"] = aggr_people_var["cycle"]/24
aggr_people_var["is_susceptible"] = aggr_people_var["is_susceptible"] /n_batch
aggr_people_var["is_immune"] = aggr_people_var["is_immune"]/n_batch
aggr_people_var["is_infected"] = aggr_people_var["is_infected"]/n_batch
aggr_people_var["is_dead"] = aggr_people_var["is_dead"]/n_batch
aggr_people["std_infected"] = np.sqrt(aggr_people_var["is_infected"])
aggr_people["lower_bound"] = aggr_people["is_infected"] - 3*aggr_people["std_infected"] # 3 sigma
aggr_people["upper_bound"] = aggr_people["is_infected"] + 3*aggr_people["std_infected"]
# + id="d7jV8ZKRw-lw"
curfew_0 = aggr_people[(aggr_people.curfew_time == curfew_time[0]) & (aggr_people.curfew_delay == curfew_delay[0]) ] # corresponding to beta = 0.075......
curfew_1 = aggr_people[(aggr_people.curfew_time == curfew_time[1]) & (aggr_people.curfew_delay == curfew_delay[0]) ]
curfew_2 = aggr_people[(aggr_people.curfew_time == curfew_time[2]) & (aggr_people.curfew_delay == curfew_delay[0]) ]
curfew_3 = aggr_people[(aggr_people.curfew_time == curfew_time[0]) & (aggr_people.curfew_delay == curfew_delay[1]) ] # corresponding to beta = 0.075......
curfew_4 = aggr_people[(aggr_people.curfew_time == curfew_time[1]) & (aggr_people.curfew_delay == curfew_delay[1]) ]
curfew_5 = aggr_people[(aggr_people.curfew_time == curfew_time[2]) & (aggr_people.curfew_delay == curfew_delay[1]) ]
# + id="nOyXcofww-lx" outputId="c0023368-9140-4685-f06f-d85bdc8ea068"
# Beta = .05 curfew_time = 18, curfew_delay = 5
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_0,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_0.days, curfew_0.lower_bound, curfew_0.upper_bound, alpha=.3)
plt.show()
# + id="IyQ59Ok0w-ly" outputId="4e3951ff-cd03-418b-9437-7976ea906e45"
# Beta = .05 curfew_time = 19, curfew_delay = 5
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_1,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_1.days, curfew_1.lower_bound, curfew_1.upper_bound, alpha=.3)
plt.show()
# + id="jfIS9YWAw-ly" outputId="1ee02494-15b1-422c-d5b1-c0cba17c7e3c"
# Beta = .05 curfew_time = 20, curfew_delay = 5
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_2,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_2.days, curfew_2.lower_bound, curfew_2.upper_bound, alpha=.3)
plt.show()
# + id="yalmmD2yw-lz" outputId="5b81c472-5b83-4ce8-b5ee-e2a279984a7e"
# Beta = .05 curfew_time = 18, curfew_delay = 10
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_3,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_3.days, curfew_3.lower_bound, curfew_3.upper_bound, alpha=.3)
plt.show()
# + id="VX2RvV9mw-l0" outputId="549aae01-f6aa-4e43-bc53-ec237458e9d9"
# Beta = .05 curfew_time = 19, curfew_delay = 10
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_4,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_4.days, curfew_4.lower_bound, curfew_4.upper_bound, alpha=.3)
plt.show()
# + id="y3GRoZZQw-l0" outputId="1224e49d-4013-49c0-e243-422997c7632f"
# Beta = .05 curfew_time = 20, curfew_delay = 10
sns.set_theme()
plt.figure(figsize = (15,8))
sns.lineplot(
data= curfew_5,
x="days", y="is_infected" ,
markers=True, dashes=False
)
plt.fill_between(curfew_5.days, curfew_5.lower_bound, curfew_5.upper_bound, alpha=.3)
plt.show()
# + id="mMkp-gt7w-l1" outputId="6dff69c3-55c2-4805-c611-2150058fdee7"
# Beta = .05 curfew_time = 18, curfew_delay = 5
sns.set_theme()
fig, axes = plt.subplots(2,3,figsize = (16,9))
sns.lineplot(ax = axes[0,0],
data= curfew_0,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[0,0].fill_between( curfew_0.days, curfew_0.lower_bound, curfew_0.upper_bound, alpha=.3)
axes[0,0].legend(["CT = 6p.m. CD = 5"])
axes[0,0].set_xlabel("Days")
axes[0,0].set_ylabel("Infected Citizens")
# Beta = .05 curfew_time = 19, curfew_delay = 5
sns.lineplot(ax = axes[0,1],
data= curfew_1,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[0,1].fill_between(curfew_1.days, curfew_1.lower_bound, curfew_1.upper_bound, alpha=.3)
axes[0,1].legend(["CT = 7p.m., CD = 5"])
axes[0,1].set_xlabel("Days")
axes[0,1].set_ylabel("Infected Citizens")
# Beta = .05 curfew_time = 20, curfew_delay = 5
sns.lineplot(ax = axes[0,2],
data= curfew_2,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[0,2].fill_between( curfew_2.days, curfew_2.lower_bound, curfew_2.upper_bound, alpha=.3)
axes[0,2].legend(["CT = 8p.m. CD = 5"])
axes[0,2].set_xlabel("Days")
axes[0,2].set_ylabel("Infected Citizens")
####------------------------------------------------------------------------------------------------------------####
# Beta = .05 curfew_time = 18, curfew_delay = 10
sns.lineplot(ax = axes[1,0],
data= curfew_3,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[1,0].fill_between(curfew_3.days, curfew_3.lower_bound, curfew_3.upper_bound, alpha=.3)
axes[1,0].legend(["CT = 6p.m. CD = 10"])
axes[1,0].set_xlabel("Days")
axes[1,0].set_ylabel("Infected Citizens")
# Beta = .05 curfew_time = 19, curfew_delay = 10
sns.lineplot(ax = axes[1,1],
data= curfew_4,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[1,1].fill_between(curfew_4.days, curfew_4.lower_bound, curfew_4.upper_bound, alpha=.3)
axes[1,1].legend(["CT = 7p.m. CD = 10"])
axes[1,1].set_xlabel("Days")
axes[1,1].set_ylabel("Infected Citizens")
# Beta = .05 curfew_time = 20, curfew_delay = 10
sns.lineplot(ax = axes[1,2],
data= curfew_5,
x="days", y="is_infected" ,
markers=True, dashes=False
)
axes[1,2].fill_between(curfew_5.days, curfew_5.lower_bound, curfew_5.upper_bound, alpha=.3)
axes[1,2].legend(["CT = 8p.m. CD = 10"])
axes[1,2].set_xlabel("Days")
axes[1,2].set_ylabel("Infected Citizens")
#plt.xlabel("Days")
#plt.ylabel("Infected Citizens")
fig.suptitle("MAS Epidemics - Different Curfew_Time & Curfew_Delay")
plt.savefig("Curfew_Beta0.1_1k.png", dpi = 600)
plt.show()
# + [markdown] id="5_i6dBJtfzZs"
# ## Maximum Percentage of Infected
# + id="3dPLHUM3gZLu"
c0_max_val = curfew_0.is_infected.max()
c1_max_val = curfew_1.is_infected.max()
c2_max_val = curfew_2.is_infected.max()
c3_max_val = curfew_3.is_infected.max()
c4_max_val = curfew_4.is_infected.max()
c5_max_val = curfew_5.is_infected.max()
# + id="ATWPRXX-f4Wq"
curfew_0[curfew_0.is_infected == c0_max_val]
# + id="0mLT7CMRf5a2"
curfew_1[curfew_1.is_infected == c1_max_val]
# + id="zQShO4CLf6cb"
curfew_2[curfew_2.is_infected == c2_max_val]
# + id="IPz_dOWsf7mR"
curfew_3[curfew_3.is_infected == c3_max_val]
# + id="Ukj01s10f8s9"
curfew_4[curfew_4.is_infected == c4_max_val]
# + id="NquEa3Vxf9n4"
curfew_5[curfew_5.is_infected == c5_max_val]
# + [markdown] id="XeQitj_9gprE"
#
| Analysis/Curfew_Processing_Beta0.1_1k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
## impoting the libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import sweetviz as sv
from pandas_profiling import ProfileReport
# %matplotlib inline
from sklearn.model_selection import train_test_split
##importing tge datasets
data = pd.read_csv('Dataset/Train.csv')
test = pd.read_csv('Dataset/Test.csv')
#taking a glance at data
data.head()
# Let's make a copy of our data so original file remains safe.
#making copy
df = data.copy()
#Getting information about datatypes
df.info()
# As we can see we have two categorical, 1 floattype and rest int type features.And we can see X_12 column has some missing values.
#lets see some description of our numerical features
df.describe()
#checking the total null values
df.isnull().sum()
# Here X_12 has 182 missing values.
#checking if dataset has any duplicated values
df.duplicated().sum()
#locating missing values in our data
df.loc[df['X_12'].isnull()]
# As we have enough amount of data so we can get rid of missing values directly by droping the same rows.
#droping the missing value rows
df.dropna()
##Bringing the numerical features together
features = ['X_1','X_2','X_3','X_4','X_5','X_6','X_7','X_8','X_9','X_10','X_11','X_12','X_13','X_14','X_15']
#Our target is to predict weather it is Multiple_Offense(1) or not(0)
target = df['MULTIPLE_OFFENSE']
# ### ---- Explore the data (EDA) ----
# Let's plot our features using sweetviz and pandas profiling libraries.These libraries are awesome they create analysis with just one line of code into html files.So lets see what we got here:
#creating report on df using sweetviz
my_report = sv.analyze([df, "Train"],target_feat= 'MULTIPLE_OFFENSE')
#storing the report into html file
my_report.show_html('Report.html')
### To Create the Simple report quickly
profile = ProfileReport(df, title='Pandas Profiling Report', explorative=True)
## stoting the report into html file
profile.to_file("output.html")
# The two above files gives us a lot of information about each feature like mean, standard deviation,values and bar chart shows the percentage of each numerical value with respect to that column.
# Lets plot heat map for see correlation between features.
#correlation plot
# Set the width and height of the figure
plt.figure(figsize=(20,14))
corr = df.corr()
sns.heatmap(corr, annot=False)
#annot=True - This ensures that the values for each cell appear on the chart.
#(Leaving this out removes the numbers from each of the cells!)
# As we can see here X_10 and X_10shows negative correlation with our target and many of them shows very little positive correlation.
# Lets see if our data has any skewness. If yes then we will normalize by using numpy's logarithmic function.
#transform the numeric features using log(x + 1)
from scipy.stats import skew
skewed = df[features].apply(lambda x: skew(x.dropna().astype(float)))
skewed = skewed[skewed > 0.75]
skewed = skewed.index
df[skewed] = np.log1p(df[skewed])
# Lets scale our data by using standardscaler function.
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df[features])
scaled = scaler.transform(df[features])
for i, col in enumerate(features):
df[col] = scaled[:,i]
# -
# Created a pipeline for filling missing values and standardizing the data.
# 1) Imputer function fill the missing values by mean or meadian, we are using meadian.
#
# 2) Standard Scaler apply scaling to the numerical features.
# +
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
num_pipeline = Pipeline([
('imputer', SimpleImputer(strategy="median")),
('std_scaler', StandardScaler()),
])
df_num_tr = num_pipeline.fit_transform(df[features])
# -
# convert the target into an numnpy array as neural network expects it to be.
target = np.array(target.values)
# split train data into train and validation
x_train, x_test, y_train, y_test = train_test_split(
df_num_tr, target,
test_size=0.2,
random_state=42
)
# ### ---- Establish a baseline ----
#
# Lets create our first classifier using randomforest classifier and checking the classification report using our validation dataset using confusion_matrix.
# ## DEVELOP MODELS
#
# We will cycle through creating features, tuning models, and training/validing models until We've reached our efficacy goal.
#
# #### our metric will be Recall and our goal is:
# - Recall ~= 1
#
# ## ---- Create models ----
# +
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(random_state=42)
forest_clf.fit(x_train, y_train)
frst_prd = forest_clf.predict(x_test)
# -
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report (frst_prd, y_test))
# Wow, we have got very good accuracy and f1_score at our first attempt.
# Lets create another classifier using Support Vector Classifier and let the randomizedsearchcv deals with the hyperparameter tunning out of the range we are giving.
# +
# traing the data by svm classifier
from sklearn.svm import SVC
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(SVC(), param_distributions, n_iter=10, verbose=2, cv=3, random_state=42)
rnd_search_cv.fit(x_train, y_train)
# -
rnd_search_cv.best_estimator_
#lets predict our validation using best estimator
y_pred = rnd_search_cv.best_estimator_.predict(x_test)
## check how our predictions worked
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report (y_pred, y_test))
# oohh!This score is lesser than what we got with randomforestclassifier.
#
# Lets try Deep Neural Network as neural networks are good in finding patterns from complex data.
# FIRST we flatten the data and then used 2 hidden layers each with 3 neurons and relu activation function.
# Then the final output layer with 2 neuron(as we have binary classes our output) with softmax activation function.
import tensorflow as tf
import keras
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(8, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.relu),
tf.keras.layers.Dense(1, activation=tf.nn.softmax)])
# Now compile the model with 'adam' optimizer and categorical loss function.
model.compile(optimizer = tf.train.AdamOptimizer(),
loss = 'categorical_crossentropy',
metrics=['accuracy'])
# +
# Now train the model using spited dataset and get the loss and accuracy as score.
batch_size = 80
history = model.fit(x_train, y_train, batch_size=batch_size, epochs=10, validation_data = (x_test,y_test), verbose=2)
# +
# Ploting the loss and accuracy curves for training and validation
fig, ax = plt.subplots(2,1)
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['acc'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_acc'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
# -
# DNN has not been its best today so lets try some other machine learning algorithams.
#glancing at test dataset
test.head()
# ching if test set has any missing values
test.isnull().sum()
# +
#test.dropna()
# -
#check the shape of test set
test.shape
##lets standardize the test features
tst_num = num_pipeline.fit_transform(test[features])
##predicting using DNN
pred = model.predict(tst_num)
pred
sub = pd.DataFrame()
# +
#sub['INCIDENT_ID'] = test['INCIDENT_ID']
# -
sub['MULTIPLE_OFFENSE'] = pred
# +
#sub.head()
# -
# We have dropped some of the rows in our test dataset so lets load it again and to deal with full dataset.
ts = pd.read_csv('Dataset/Test.csv')
ts_nm = num_pipeline.fit_transform(ts[features])
# Making prediction using rnd_svc
ts_pred = rnd_search_cv.best_estimator_.predict(ts_nm)
sbm = pd.DataFrame()
sbm['INCIDENT_ID'] = ts['INCIDENT_ID']
sbm['MULTIPLE_OFFENSE'] = ts_pred
# +
#sbm.head()
# + active=""
# del sbm['MULTIPLE_OFFENCE']
# -
sbm.to_csv('submission.csv', index = False)
chk = pd.read_csv('submission.csv')
chk.head()
This submission gave us score of 79.2.
##Lets predict the test data using random forest
prediction = forest_clf.predict(ts_nm)
#create a empty dataframe
frst = pd.DataFrame()
frst['INCIDENT_ID'] = ts['INCIDENT_ID']
frst['MULTIPLE_OFFENSE']= prediction
frst.head()
#convert dataframe to csv for submission
frst.to_csv('submission_2.csv', index = False)
# This submssion got us a suffosticated score of 92 which is quite good upgrade.
#
#
#
# Now it needs to improve.For that we will be using xgboost and ensemble method Voteclassifier.
import xgboost as xgb
from sklearn.model_selection import cross_val_score
model1=xgb.XGBClassifier(colsample_bylevel= 1, learning_rate= 0.1,max_depth=10, n_estimators= 1000)
result=cross_val_score(estimator=model1,X=x_train,y=y_train,cv=10)
print(result)
print(result.mean())
model1.fit(x_train, y_train)
xg_pred = model1.predict(x_test)
print(classification_report (xg_pred, y_test))
# ## ---- Feature Importance ----
indices=np.argsort(model1.feature_importances_)
plt.figure(figsize=(10,10))
g = sns.barplot(y=df[features].columns[indices][:40],x = model1.feature_importances_[indices][:40] , orient='h')
# This result is mirroring our correlation matrix.Showing that X_10,X_12,X_15 and X_11 are some of the most important features here.And seems like X_1 and X_5 are almost uneesential features.
from sklearn.ensemble import AdaBoostClassifier, GradientBoostingClassifier, ExtraTreesClassifier, VotingClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV, cross_val_score, StratifiedKFold, learning_curve
# Cross validate model with Kfold stratified cross val
kfold = StratifiedKFold(n_splits=10)
# +
## using gradientboostingclassifier
GBC = GradientBoostingClassifier()
gb_param_grid = {'loss' : ["deviance"],
'n_estimators' : [400,500],
'learning_rate': [0.1, 0.2],
'max_depth': [4, 8],
'min_samples_leaf': [100,150],
'max_features': [0.3, 0.1]
}
gsGBC = GridSearchCV(GBC,param_grid = gb_param_grid, cv=kfold, scoring="accuracy", n_jobs= 4, verbose = 1)
gsGBC.fit(x_train,y_train)
model2 = gsGBC.best_estimator_
# Best score
print(gsGBC.best_score_)
print(gsGBC.best_params_)
# -
model2 = gsGBC.best_estimator_
model2.fit(x_train, y_train)
y_prd = model2.predict(x_test)
print(classification_report (y_prd, y_test))
# Excellent! almost perfect score we have got here.
rslt = model2.predict(ts_nm)
sbmsn = pd.DataFrame()
sbmsn['INCIDENT_ID'] = ts['INCIDENT_ID']
sbmsn['MULTIPLE_OFFENSE'] = rslt
sbmsn.head()
sbmsn.to_csv('submission_3.csv', index = False)
# Now we have got our highest score of 98.46.Now we will proceed further to increase score upto perfect score(100).
# For that we will use an ensemble method called voting classifier.
# ## ---- Ensembling models ----
# +
from sklearn.ensemble import VotingClassifier
votingC = VotingClassifier(estimators=[('gbc',model2),('rfc',forest_clf),('xgb',model1)], voting='soft', n_jobs=4)
votingC.fit(x_train,y_train)
y_pred2=votingC.predict(x_test)
# -
print(classification_report (y_prd, y_test))
votes = votingC.predict(ts_nm)
submission=pd.DataFrame()
submission['INCIDENT_ID'] = ts['INCIDENT_ID']
submission['MULTIPLE_OFFENSE'] = votes
submission.to_csv('submission_4.csv', index = False)
# ### ---- Select best model ----
xg_votes = model1.predict(ts_nm)
submit=pd.DataFrame()
submit['INCIDENT_ID'] = ts['INCIDENT_ID']
submit['MULTIPLE_OFFENSE'] = xg_votes
submit.to_csv('submission_5.csv', index = False)
# Using XGboost gave us the highest score of 98.79457.And by the barplot of feature importances it is clear that X-1 and X_5 are of no importance with respect to the offence.While X_10 and X_12 are of very musch importance and X_15 and X_11 and others are also quite important.
# So we can perform some feature selection further to increase our score.
#
# ### ---- Saving best model ----
#
# First save our model for future predictions.
## Create a Pickle file using serialization
import pickle
pickle_out = open("classifier.pkl","wb")
pickle.dump(model1, pickle_out)
pickle_out.close()
# ### ---- Feature Selection ----
new_feat = ['X_10','X_11','X_12','X_15']
df_nm_new = num_pipeline.fit_transform(df[new_feat])
model1.fit(df_nm_new, target)
df_prd = model1.predict(df_nm_new)
print(classification_report (df_prd, target))
new_nm = num_pipeline.fit_transform(ts[new_feat])
new_prd = model1.predict(new_nm)
new_subm = pd.DataFrame()
new_subm['INCIDENT_ID'] = ts['INCIDENT_ID']
new_subm['MULTIPLE_OFFENSE'] = new_prd
new_subm.to_csv('submission_6.csv', index = False)
# This submission got us score of 99.00. WHich is our highest score so far.
mod_feat = ['X_10','X_11','X_12','X_15','X_2','X_4','X_6', 'X_8', 'X_9','X_13']
df_nm_mod = num_pipeline.fit_transform(df[mod_feat])
model1.fit(df_nm_mod, target)
mod_nm = num_pipeline.fit_transform(ts[mod_feat])
mod_prd = model1.predict(mod_nm)
mod_sub = pd.DataFrame()
mod_sub['INCIDENT_ID'] = ts['INCIDENT_ID']
mod_sub['MULTIPLE_OFFENSE'] = mod_prd
mod_sub.to_csv('submission_7.csv', index = False)
# Okay! once again we got the same score 99.00.
# So our final submission file is 'submission_7.csv' with the score of 99.00.
#
# ### ---- Furthure Scope ----
#
# Here we got the final score of 99.00.It is not the perfect score but we can improve it by doing some feature engineering.Rather than doing random feature engineering it would be good to have some domain knowledge.That we can discuss with our domain expert like what is the significance of the features,other scanerios that causes hacking,any kind of security issues and weather our system was working propely or not where hacking has happened,any other loopholes in our security system etc.
# This discussion will surely help us creating more features and do proper feature engineering and finally improve our prediction.
#
# ### ---- THANK YOU ----
# +
from zipfile import ZipFile
#creating a zipfile object
zipobj = ZipFile('submission.zip','w')
## Adding multiple files to the zip
zipobj.write('output.html')
zipobj.write('Report.html')
zipobj.write('Untitled.ipynb')
zipobj.write('classifier.pkl')
##closing the zip_file
zipobj.close()
# -
| Untitled.ipynb |