code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Before we begin
#
# To talk to a MySQL database, you need to have a mysql "client". **I have already installed this for you!** But if you ever need to do it yourself, the commands are:
#
#
# sudo apt-get update
# sudo apt-get install mysql-client
#
#
# # Introduction to Databases and Structured Query Language (SQL)
#
# As Data Scientists, you will frequently want to store data in an organized, structured manner that allows you to do complex queries. Because you are good Data Scientists, [**you do not use Excel!!!**](https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-5-80)
#
# In this course, we will only discuss **Relational Databases**, because those are the most common in bioinformatics. (There are other kinds!!). So when I say "database" I mean "relational database".
#
# Databases are used to store information in a manner that, when used properly, is:
# a) highly structured
# b) constrained (i.e. detects errors)
# c) transactional (i.e. can undo a command if it discovers a problem)
# d) indexed (for speed of search)
# e) searchable
#
# The core concept of a database is a **Table**. **Tables contain one particular "kind" of information** (e.g. a Table could represent a Student, a University, a Book, or a portion of a Clinical Record. **Try not to mix info about different things in the same table**
#
# Tables contain **Rows** and **Columns** where, generally, every column represents a "feature" of that information (e.g. a Student table might have **["name", "gender", "studentID", "age"]** as its columns/features). Every row represents an "individual", and their values for each feature (e.g. a Row in a Student table might have **["<NAME>", "M", "163483", "35"]** as its values.
#
# A Database may have many Tables that represent various kinds of related information. For example, a library database might have a Books table, a Publishers table, and a Locations table. A Book has a Publisher, and a Location, so the tables need to be **connected to one another**. This is achieved using **keys**. Generally, **every row (individual)** in a table has a **unique identifier** (generally a number), and this is called its **key**. Because it is unique, it is possible to refer unambiguously to that individual record.
#
# I think the easiest way to learn about databases and SQL is to start building one! We will use the MySQL Docker Container that we created in the previous lesson. We are going to create a Germplasm database (seed stocks). It will contain information about the seed (its amount, its harvest date, its location), the germplasm (its species, the allele it carries), and about the genetics related to that allele (the gene_id, the gene name, the protein name, and a link to the GenBank record)
#
# (if that Docker Container isn't running, please **docker start course-mysql** now!)
#
# **Note: This Jupyter Notebook is running the Python kernel. This allows us to use some nice tools in Python (the sql extension and SqlMagic) that provide access to the mysql database server from inside of the Notebook. You don't need to know any Python to do this. Note also that you can do exactly the same commands in your Terminal window.**
#
# To connect to the MySQL Docker Container from your terminal window, type:
#
# mysql -h 127.0.0.1 -P 3306 --protocol=tcp -u root -p
#
# -h: host
# the address in every linux computer is always 127.0.0.1
# -P: port (when starting the container we communicated 3306 to 3306 host to container)
# --protocol: the protocol
# -u: username
# my user is root
# -p: request to ask for password (also root)
#
# (then enter your password '<PASSWORD>' to access the database)
#
# Ahora nos sale mysql> y eso es que estamos en nuestro Docker container
#
# para salir: exit;
# todos los comandos de mysql acaban en ; como en java
# <pre>
#
#
# </pre>
# # SQL
#
# Structured Query Language is a way to interact with a database server. It is used to create, delete, edit, fill, and query tables and their contents.
#
# First, we will learn the SQL commands that allow us to explore the database server, and create new databases and tables.. Later, we will use SQL to put information into those tables. Finally, we will use SQL to query those tables.
#
# ## Python SQL Extension
#
# The commands below are used to connect to the MySQL server in our Docker Container. You need to execute them ONCE. In every subsequent Juputer code window, you will have access to the database.
#
# all SQL commands are preceded by
#
# # %sql
#
# (**only in the Python extension! Not in your terminal window!**)
#
# all SQL commands end with a ";"
# %load_ext sql
# #%config SqlMagic.autocommit=False
# %sql mysql+pymysql://root:root@127.0.0.1:3306/mysql
# #%sql mysql+pymysql://anonymous@ensembldb.ensembl.org/homo_sapiens_core_92_38
# ## show databases
#
# **show databases** is the command to see what databases exist in the server. The ones you see now are the default databases that MySQL uses to organize itself. _**DO NOT TOUCH THESE DATABASES**_ **EVER EVER EVER EVER**
# +
# %sql show databases;
# -
# todo lo que hacemos en python (con %sql) lo podemos hacer desde la terminal en el Docker container (sin %sql) cuando estamos dentro de mysql>
# ## create database
#
# The command to create a database is **create database** (surprise! ;-) )
#
# We will create a database called "germplasm"
#
#
#
# %sql create database germplasm;
# %sql show databases
# ## use database_name
#
# the **use** command tells the server which database you want to interact with. Here we will use the database we just created
# +
# %sql use germplasm; # when we want to interact with a database we first have to do this
# -
# ## show tables
#
# The show tables command shows what tables the database contains (right now, none!)
# %sql show tables;
# # Planning your data structure
#
# This is the hard part. What does our data "look like" in a well-structured, relational format?
#
# Starting simply:
#
# <center>stock table</center>
#
# amount | date | location
# --- | --- | ---
# 5 | 10/5/2013 | Room 2234
# 9.8 | 12/1/2015 | Room 998
#
#
# -----------------------------
#
#
# <center>germplasm table</center>
#
# taxonid | allele
# --- | ---
# 4150 | def-1
# 3701 | ap3
#
# --------------------------------
#
# <center>gene table</center>
#
# gene | gene_name | embl
# --- | --- | ---
# DEF | Deficiens | https://www.ebi.ac.uk/ena/data/view/AB516402
# AP3 | Apetala3 | https://www.ebi.ac.uk/ena/data/view/AF056541
#
#
#
#
# now we need to connect each row of each table
# ## add indexes
#
# It is usually a good idea to have an index column on every table, so let's add that first:
#
#
# <center>stock table</center>
#
# id | amount | date | location
# --- | --- | --- | ---
# 1 | 5 | 10/5/2013 | Room 2234
# 2 | 9.8 | 12/1/2015 | Room 998
#
#
# -----------------------------
#
#
# <center>germplasm table</center>
#
# id | taxonid | allele
# --- | --- | ---
# 1 | 4150 | def-1
# 2 | 3701 | ap3
#
# --------------------------------
#
# <center>gene table</center>
#
# id | gene | gene_name | embl
# --- | --- | --- | ---
# 1 | DEF | Deficiens | https://www.ebi.ac.uk/ena/data/view/AB516402
# 2 | AP3 | Apetala3 | https://www.ebi.ac.uk/ena/data/view/AF056541
#
#
# ## find linkages
#
# * Every germplasm has a stock record. This is a 1:1 relationship.
# * Every germplasm represents a specific gene. This is a 1:1 relationship
#
# So every germplasm must point to the index of a stock, and also to the index of a gene
#
# Adding that into our tables we have:
#
#
# Our germplasm points to the stock --> the germplasm table has an additional column, the stock table doesnt change
#
# Our germplasm points to the gene --> the germplasm table has an additional column, the gene table doesnt change
#
# **the numbers dont need to be the same!!**
#
#
# -----------------------------
#
#
# <center>stock table</center>
#
# id | amount | date | location
# --- | --- | --- | ---
# 1 | 5 | 10/5/2013 | Room 2234
# 2 | 9.8 | 12/1/2015 | Room 998
#
#
# -----------------------------
#
#
# <center>germplasm table</center>
#
# id | taxonid | allele | stock_id | genetics_id
# --- | --- | --- | --- | ---
# 1 | 4150 | def-1 | 2 | 1
# 2 | 3701 | ap3 | 1 | 2
#
# --------------------------------
#
# <center>gene table</center>
#
# id | gene | gene_name | embl
# --- | --- | --- | ---
# 1 | DEF | Deficiens | https://www.ebi.ac.uk/ena/data/view/AB516402
# 2 | AP3 | Apetala3 | https://www.ebi.ac.uk/ena/data/view/AF056541
#
#
# ## data types in MySQL
#
# I will not discuss [all MySQL Datatypes](https://dev.mysql.com/doc/refman/5.7/en/data-types.html), but we will look at only the ones we need. We need:
#
# * Integers (type INTEGER) - integers
# * Floating point (type FLOAT) - integers/decimals
# * Date (type DATE [in yyyy-mm-dd format](https://dev.mysql.com/doc/refman/5.7/en/datetime.html) ) ISO 8601
# * Characters (small, variable-length --> type [VARCHAR(x)](https://dev.mysql.com/doc/refman/5.7/en/char.html) ) x is the maximum length of the varchar - text
#
# <pre>
#
#
# </pre>
# ## create table
#
# tables are created using the **create table** command (surprise!)
#
# The [syntax of create table](https://dev.mysql.com/doc/refman/5.7/en/create-table.html) can be quite complicated, but we are only going to do the most simple examples.
#
# create table table_name (column_name column_definition, column_name column_definition, ........)
#
# column definitions include the data-type, and other options like if it is allowed to be null(blank), or if it should be treated as an "index" column.
#
# Examples are easier to understand than words... so here are our table definitions:
#
#
# #%sql drop table stock
# %sql CREATE TABLE stock(id INTEGER NOT NULL AUTO_INCREMENT PRIMARY KEY, amount FLOAT NOT NULL, date DATE NOT NULL, location VARCHAR(20) NOT NULL);
# %sql DESCRIBE stock
# NOT NULL: it cannot be blank
#
# AUTO_INCREMENT: so it goes up 1 by 1, useful with the id, not with the other numbers
#
# PRIMARY KEY: the key (in germplasm table, stock id WILL point to this id)
#
# to recreate the table we first have to delete the table with drop table
# #%sql drop table germplasm
# %sql CREATE TABLE germplasm(id INTEGER NOT NULL AUTO_INCREMENT PRIMARY KEY, taxonid INTEGER NOT NULL, allele VARCHAR(10) NOT NULL, stock_id INTEGER NOT NULL, gene_id INTEGER NOT NULL);
# %sql DESCRIBE germplasm
# #%sql drop table gene
# %sql CREATE TABLE gene(id INTEGER NOT NULL AUTO_INCREMENT PRIMARY KEY, gene VARCHAR(10) NOT NULL, gene_name VARCHAR(30) NOT NULL, embl VARCHAR(70) NOT NULL);
# %sql DESCRIBE gene
# %sql show tables;
# ## loading data
#
# There are many ways to import data into MySQL. If you have data in another (identical) MySQL database, you can "dump" the data (download it entirely), and then import it directly. If you have tab or comma-delimited (tsv, csv) you can **sometimes** import it directly from these formats. You can also **enter data using SQL itself. This is usually the safest way, when you have to keep multiple tables synchronized** (as we do, since the germplasm table is "linked to" the other two tables)
#
# ## insert into
#
# The command to load data is:
#
# insert into table_name (field1, field2, field3) values (value1, value2, value3)
#
# Now... what data do we need to add, **in what order?** --> first the things that dont point to other things (the things that dont have the id of another table)
#
# **we start with the independents and go ""inwards"" once we have the information that the others depend on**
#
# The germplasm table needs the ID number from both the gene table and the stock table, so we cannot enter the germplasm information first. We must therefore enter the gene and stock data first.
#
# **WE DONT PUT DATA INTO THE ID BECAUSE WE SET IT AS AUTO_INCREMENT**
# NOTE - we DO NOT put data into the "id" column! This column is auto_increment, so it "magically" creates its own value
# %sql INSERT INTO gene (gene, gene_name, embl) VALUES ('DEF', "Deficiens", 'https://www.ebi.ac.uk/ena/data/view/AB516402');
# %sql INSERT INTO gene (gene, gene_name, embl) VALUES ('AP3', "Apetala3", 'https://www.ebi.ac.uk/ena/data/view/AF056541');
# To show the **last inserted id** we use:
# %sql SELECT last_insert_id(); # just to show you that this function exists!
# %sql INSERT INTO stock(amount, date, location) VALUES (5, '2013-05-10', 'Room 2234');
# %sql INSERT INTO stock(amount, date, location) VALUES (9.8, '2015-01-12', 'Room 998');
# #### Almost ready!
#
# We now need to know the index numbers from the stock and gene databases that correspond to the data for the germplasm table. For this, we need to learn another function: **select**
#
# ## Select statements
#
# **Select** is the command used to query the database. We will look in more detail later, but now all you need to know is that the most basic structure is:
#
# select * from table_name
#
#
# %sql SELECT * FROM stock; # notice that the id number was automatically generated
# %sql SELECT * FROM gene;
# <pre>
#
#
# </pre>
#
# Just a reminder, our germplasm data is:
#
# id | taxonid | allele | stock_id | gene_id
# --- | --- | --- | --- | --- |
# 1 | 4150 |def-1
# 2 | 3701 | ap3
#
#
# We need to connect the *germplasm* table **gene_id** to the appropriate **id** from the *gene* table. i.e.
#
# def-1 allele ---> DEF gene (id = 1)
# ap3 allele ---> AP3 gene (id = 2)
#
# We need to connect the *germplasm* table **stock_id** to the appropriate **id** from the *stock* table. i.e.
#
# def-1 allele ---> Room 998 (id = 2)
# ap3 allele ---> Room 2234 (id = 1)
#
# Now we are ready to do our ("manual") insert of data into the *germplasm* table:
#
# %sql INSERT INTO germplasm (taxonid, allele, stock_id, gene_id) VALUES (4150, 'def-1', 2, 1 );
# %sql INSERT INTO germplasm (taxonid, allele, stock_id, gene_id) VALUES (3701, 'ap3', 1, 2 );
# %sql SELECT * FROM germplasm;
# This is the manual way, it is not very elegant nor efficient. but it is ok. We'll see alternatives later.
#
# ## SQL UPDATE & SQL WHERE
#
# Imagine that we are going to plant some seed from our def-1 germplasm. We need to update the *stock* record to show that there is now less seed available. We do this using an [UPDATE statement](https://www.techonthenet.com/mysql/update.php). **UPDATE is used to change the values of a particular column or set of columns**. But we **don't want to change ALL** of the values in that column, we only want to change the values for the DEF stock. For that, we need a WHERE clause.
#
# WHERE allows you to set the conditions for an update. The general form is:
#
# UPDATE table_name SET column = value WHERE column = value;
#
# the second column may or may not be the same
#
# Positions of the database: table.column
#
# We will sow 1g of seed from DEF (stock.id = 2) (note that I am now starting to use the MySQL syntax for referring to *table*.**column** - the tablename followed by a "." followed by the column name).
#
# The simplest UPDATE statement is:
#
# %sql UPDATE stock SET amount = 8.8 WHERE id = 2;
# %sql SELECT * FROM stock;
#
# <pre>
#
# </pre>
# This simple solution is not very "friendly"... you are asking the database user to already know what the remaining amount is! It would be better if we simply reduced the amount by 1g.
#
# We needed to do SELECT * first, we want to avoid these steps and make the calculations automatically (removing 1 g).
#
# That is done using in-line equations, like this:
#
#
# +
# %sql UPDATE stock SET amount = amount-1 WHERE id = 2;
# %sql SELECT * FROM stock;
# This way we didn't have to know what the amount was, we only know that we took 1 g from it.
# -
# <pre>
#
#
# </pre>
# ## Using indexes and 'joining' tables
#
# Index columns were keys, and there is one per table which is the primary key.
#
# The UPDATE we did is still not very friendly! My stock table does not have any information about what gene or allele is in that stock, so we have to **know** that the stock record is stock.id=2. This is bad!
#
# It would be better if we could say **"plant 1 gram of the stock that represents gene record DEF"**, but that information exists in two different tables. How do we **join tables?**
#
# This is the main purpose of the "id" column. Note that, when we defined that column, we said that it is "auto_increment, not null, primary key", meaning that every record must have an id, and every id must be unique (**NOTE: an auto-increment id *should NEVER NEVER NEVER NEVER be manually modified/added*!!!** <span style="color:red;">You Have Been Warned!!!</span>). Being a 'primary key' means that this column was intended to be the "pointer" from other tables in the database (like our germplasm table, that points to the id of the stock, and the id of the gene, tables)
#
# When using UPDATE with multiple tables, we must name all of the tables, and then make the connection between them in the "where" clause, using *table*.**column** notation.
#
# The update clause below shows how this is done (a "\\" character means that the command continues on the next line):
#
# +
# %sql UPDATE stock, germplasm SET stock.amount = stock.amount-1 \
# WHERE \
# stock.id = germplasm.stock_id \
# AND \
# germplasm.allele = 'def-1';
# %sql SELECT * FROM stock;
# -
# <pre>
#
# </pre>
# # Challenges for you!
#
# 1. (hard) when we plant our seeds, we should update both the quantity, and the date. What does that UPDATE statement look like?
#
# change the date manually first and then do it automatically
#
# 2. (very hard!) when we plant our seed, instead of using the allele designation (def-1) I want to use the gene designation (DEF). This query spans **all three tables**. What does the UPDATE statement look like?
#
# <span style="visibility:hidden;">
# Challenge 1
# # %sql UPDATE stock,germplasm SET stock.amount=stock.amount-1, stock.date="2018-09-06" WHERE \
# # stock.id = germplasm.stock_id AND \
# # germplasm.allele='def-1';
# Challenge2
# # %sql UPDATE stock,germplasm,gene SET stock.amount=stock.amount-0.2, stock.date="2018-09-06" WHERE \
# # stock.id = germplasm.stock_id AND \
# # gene.id = germplasm.gene_id AND \
# # gene.gene='DEF';
# </span>
# +
# challenge 1
# changing date manually:
# %sql UPDATE stock, germplasm SET stock.amount = stock.amount - 1, stock.date = "2020-01-01" \
# WHERE stock.id = germplasm.stock_id AND germplasm.allele = "def-1"
# %sql SELECT * FROM stock
# +
# challenge 1
# changing date to today's date automatically: CURRENT_DATE (format yyyy-mm-dd), o CURDATE()
# %sql UPDATE stock, germplasm SET stock.amount = stock.amount - 1, stock.date = CURRENT_DATE \
# WHERE stock.id = germplasm.stock_id AND germplasm.allele = "def-1"
# %sql SELECT * FROM stock
# +
# challenge 2
# %sql UPDATE stock, germplasm, gene SET stock.amount = stock.amount - 1 \
# WHERE \
# stock.id = germplasm.stock_id \
# AND \
# germplasm.gene_id = gene.id \
# AND \
# gene.gene = "DEF"
# %sql SELECT * FROM stock
# -
# <pre>
#
#
# </pre>
#
# # SELECT queries
#
# Querying the data is the most common operation on a database. You have seen simple SELECT queries, but now we will look at more complex ones.
#
# The general structure is:
#
# SELECT table1.column1, ... FROM table1, ... WHERE condition1 [AND|OR] condition2....
#
# You probably understand this enough to show you the query that will show you all of the data:
# +
# %sql SELECT * FROM gene, stock, germplasm WHERE \
# germplasm.stock_id = stock.id AND \
# germplasm.gene_id = gene.id;
# you are linking the three tables together, so you display everything doing this (* is eveything)
# -
#
# # Dealing with missing records - JOIN clauses
#
# **Credit for the Venn diagrams used in this section goes to [Flatiron School](https://learn.co/) and are linked from their tutorial on [JOINs in SQL](https://learn.co/lessons/sql-complex-joins-readme) published under the [CC-BY-NC 4.0 license](https://creativecommons.org/licenses/by-nc-sa/4.0/)**
#
# Your first database will probably be complete, and perfect! You will be very proud of it! ;-)
#
# Over time, things will happen. Records will be deleted, and records will be added where there is incomplete information, for example, a germplasm stock record where the gene is unknown. You should think about these situations, because there are NO RULES telling you what you should do! You have to make a decision - a *policy* for your database - and you should follow that policy like a religion!
#
# For example:
#
# * If there is no known gene for a given germplasm, what does in the stock.allele column? What goes in the stock.gene_id column? What goes in the gene table? Discuss....
#
# Charlita:
#
# Si usamos NULL en un id, no podemos porque hemos usado NOT NULL al declarar. Lo podemos llamar "NULL".
#
# Hacemos un mismo NULL para todos??? o un NULL distinto para cada uno???
#
# ?????
#
# One of the really common queries that we do is SELECT DISTINCT gene name from the gene table. If we put the name "NULL" into the gene name column, WE WONT BE ABLE TO COUNT THE NuMBER OF GENES IN THE DATABASE because all of those are called the same. We there could have the policy of SELECT DISTINCT gene id because that is for sure different, even though it is not intuitive, but that is why we have a policy fot that.
#
# One good posibility is: adding something that is CLEARLY not a gene id but also DISTINCT: for example NULL+current_date or something like that, some identifier that is not a name but is not the same always.
#
# Another is having another column, a flag that is yes or no depending on whether it is unknown or not.
#
# **It is important to make these choices in the best way (there is no right or wrong really, but we have to think) and be ALWAYS CONSISTENT AND DO THE SAME THING SO IT IS CONSISTENT
#
# -----------------------------------
#
# * If a stock is fully planted - no more seeds - what do you do? Should you delete the record? If you do, then your germplasm table is linked through the stock_id to a stock that no longer exists. If you gather more seed in the future, is that the same stock? (answer: NO, it is not!!!).... so do you update the existing stock record to say there is now 10g of seed?
#
# Possibility: adding a new germplasm to refer to the new stock.
#
# **Never refill (in this case) an empty stock because it is not the same stock**
#
# 0 is already a flag for an empty stock, we don't need another flag.
#
# Do NOT delete the record so the database is not interrupted.
#
# The suggests the correct solution would be: create a new germplasm and stock so that both of those are correct, the germplasm would be identical but with another id.
#
# ------------------------------------
#
# * Remember, you are trapped! In your table definition you declared all columns to be "NOT NULL", meaning that if the row exists, there must be a value for each column in the row! What do you do if there isn't a value to put into that column?
# * zero?
# * What if you change the column definition to allow NULL?
# * What does NULL mean? What does zero mean?
# * How does software respond to NULL or zero values? (you don't know this yet, but we can talk about it)
#
# If a column is null, what does it mean.
# - never tested?
# - they can't test it?
#
# We have to define what null means, how 0 means, and think how the software responds to this.
#
# Some languages consider null and 0 equivalent to false. But they dont necessarily mean false in the context of the database, they can mean a different thing. So careful!
#
# ---------------------------
#
# **Policies: decisions that we have to make, and we follow them ALWAYS so we know afterwards years later how to interpret some weird things of the database**
#
# For our database, I am going to suggest this policy:
# 1. if we don't know the allele, we put "unknown" in the allele column
# 2. We put '0' into the gene_id column (auto_increment starts with 1 in the gene table, so a 0 will match nothing!) -> our database is only slightly inconsistent.
# 3. We DO NOT add a gene record at all.
#
# He doesn't say this is a good policy, but we'll follow it in this notebook.
#
# Let's add a record like this one to our database:
#
# %sql INSERT INTO stock(amount, date, location) VALUES (23, '2018-05-12', 'Room 289');
# %sql INSERT INTO germplasm (taxonid, allele, stock_id, gene_id) VALUES (4150, 'unknown', LAST_INSERT_ID(), 0 );
# note that I am using LAST_INSERT_ID to capture the auto_increment value from the stock table insert
# this ensures that the germplasm and stock tables are 'synchronized'
# it has to be IMMEDIATELY AFTER TRIGGERING AN AUTO-INCREMENT
# %sql SELECT * FROM germplasm;
# #%sql SELECT * FROM gene;
# <pre>
#
# </pre>
# That looks good! ...but we have just created a problem! **gene_id=0 doesn't exist in the gene table**, 0 is broken (not a legitimate value)
#
#
# so what happens with our beautiful SELECT query that we just created above?
#
#
# %sql SELECT * FROM gene, germplasm WHERE \
# germplasm.gene_id = gene.id;
# If the person writing the query doesn't know about our policy, it will seem to them that there is no more germplasm. There will be missing records this way.
#
# Therefore there are JOINs in SQL:
#
# ### OH CRAP!!!! We lost our data!!
#
# Our "unknown" germplasm has disappeared!! Or has it?
#
# The problem is that stock.gene_id = gene.id failed for the "unknown" record, and so it isn't reflected in the output from the query. THIS IS BAD, if (for example) you were trying to take an inventory of all germplasm stocks you had!
#
# How do we solve this? The answer is to use SQL's "JOIN" instruction.
#
# There are four kinds of JOIN: INNER, LEFT OUTER, RIGHT OUTER, and FULL OUTER.
#
# **DEFAULT: INNER -> if something doesn't exist in one of the sides, it doesn't show. It only shows the common ones.**
#
# The join we are doing with our current SELECT query is an INNER join. Using a Venn diagram, the query looks like this:
#
# <a href='https://learn.co/lessons/sql-complex-joins-readme'><img src='http://readme-pics.s3.amazonaws.com/Inner%20Join%20Venn%20Diagram.png' width=300px/></a>
#
#
# Effectively, the intersection where BOTH the 'left' (gene.id) and 'right' (germplasm.id) are true.
#
# You can duplicate this behavior using the INNER JOIN instruction. The syntax is a little bit different - look:
#
# +
# %sql SELECT * FROM gene INNER JOIN germplasm ON \
# germplasm.gene_id = gene.id;
# we replace the table WHERE condition with table1 INNER JOIN table2 ON condition
# -
# <pre>
#
#
# </pre>
# What we want is a query that allows one side to be "missing/absent/NULL", but the other side to exist.
#
# Perhaps we need a "LEFT JOIN"?
#
# ... gene LEFT JOIN germplasm ...
#
#
# Again, in this situation, "LEFT" means the table on the Left side of the SQL JOIN statement (gene)
#
# As a Venn diagram, Left joins look like this (ignore the :
#
# <a href='https://learn.co/lessons/sql-complex-joins-readme'><img src='http://readme-pics.s3.amazonaws.com/Left%20Outer%20Join%20Venn%20Diagram.png' width=300px/></a>
#
# What it means is that, **in addition to the perfect matches at the intersection**, the record on the left (the gene record) should be included in the result set, **even if it doesn't match** with a germplasm record (on the right). Is that the solution to our problem?
#
# %sql SELECT * FROM gene LEFT JOIN germplasm ON \
# germplasm.gene_id = gene.id;
# gene (left) LEFT JOIN germplasm (right) -> gene doesn't have extra things, the extra things are on the right. We could do it reversed or do RIGHT JOIN
# ## PFFFFFF!! No, that was not the solution
#
# Why?
#
# What about a RIGHT JOIN?
#
# ... gene RIGHT JOIN germplasm ...
#
# <a href='https://learn.co/lessons/sql-complex-joins-readme'><img src='http://readme-pics.s3.amazonaws.com/Right%20Outer%20Join%20Venn%20Diagram.png' width=300px/></a>
#
#
#
# Again, in this situation, "RIGHT" means the table on the Right side of the SQL JOIN statement (germplasm). So the germplasm record should be included in the result set, even if a gene record does not exist. ...that sounds much more likely to be correct!
#
# %sql SELECT * FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id;
# ### Voila!!
#
#
#
#
# ## Your turn
#
# 1) Create another record, where in this case, there is no **stock**, but there is a germplasm and a gene record.
# 2) Create the JOIN query between germplasm and stock that includes all germplasm records
#
# +
# %sql INSERT INTO gene(gene, gene_name, embl) VALUES ("CRO", "Crocodile", "crocodile_url");
# #%sql INSERT INTO germplasm (taxonid, allele, stock_id, gene_id) VALUES (4322, "cro-4", 0, LAST_INSERT_ID());
# you don't necessarily know the allele because you don't have the stock!!
# #%sql INSERT INTO germplasm (taxonid, allele, stock_id, gene_id) VALUES (5122, "unknown", 0, LAST_INSERT_ID());
# %sql SELECT * FROM germplasm
# -
# %sql SELECT * FROM germplasm LEFT JOIN stock ON \
# germplasm.stock_id = stock.id
# <pre>
#
#
# </pre>
# # Other SELECT "magic"
#
# You can do many other useful things with SELECTS, such as:
#
# ## COUNT()
#
# If you want to count the number of records returned from a query, use the **COUNT() AS your_name** function:
#
# %sql SELECT COUNT(*) AS "Number Of Matches" FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id;
# **Important in code:** the table header becomes Number Of Matches. We need to choose what the column header is going to be in order to extract the quantity afterwards.
# ## SUM(), AVG(), MAX()
#
# You can do mathematical functions on results also, for example, you can take the SUM of a column - how much seed do we have in total?
#
# (look carefully at this query! It's quite complicated!):
#
# %sql SELECT SUM(stock.amount) FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id \
# INNER JOIN stock ON germplasm.stock_id = stock.id;
# <pre>
#
# </pre>
# Or you could take the **average AVG()** of a column - what is the average quantity of seed we have?
#
# %sql SELECT AVG(stock.amount) FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id \
# INNER JOIN stock ON germplasm.stock_id = stock.id;
# <pre>
#
# </pre>
# Or you could take the **max MAX()** value of a column - what is the largest quantity of seed we have in our stocks?
# %sql SELECT MAX(stock.amount) FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id \
# INNER JOIN stock ON germplasm.stock_id = stock.id;
# ## ORDER BY
#
# You can put your results in a specific order:
#
# Final clause: ORDER BY
#
# %sql SELECT gene.gene_name, stock.amount FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id \
# INNER JOIN stock ON germplasm.stock_id = stock.id \
# ORDER BY stock.amount DESC; # change this to ASC
# %sql SELECT gene.gene_name, stock.amount FROM gene RIGHT JOIN germplasm ON \
# germplasm.gene_id = gene.id \
# INNER JOIN stock ON germplasm.stock_id = stock.id \
# ORDER BY stock.amount ASC; # change this to ASC
# <pre>
#
#
# </pre>
# ## Conclusion
#
# 1) Databases are a very powerful way to store structured information - far far better than Excel Spreadsheets! -> **Don't use Excel as a database**
#
# 2) It will take you years (literally, years!) to become an expert in MySQL! We have only explored the most common functions here.
# +
# Dropping the database, DO THIS BEFORE THE EXAM
# %sql drop database germplasm;
| Lesson 3 - Introduction to Databases and SQL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.6 64-bit (''alexl'': virtualenv)'
# name: python36664bitalexlvirtualenvb9f0b0a3af2a4e06a89ee778b9503914
# ---
# +
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
from sklearn.cluster import SpectralClustering, KMeans
from sklearn.metrics import pairwise_distances
from sklearn import metrics
import os
import networkx as nx
import numpy as np
import pandas as pd
from sklearn.linear_model import LogisticRegressionCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
# %matplotlib inline
# +
data_dir = os.getcwd()
cora_location = os.path.expanduser(os.path.join(data_dir, "cora/cora.cites"))
g_nx = nx.read_edgelist(path=cora_location)
cora_data_location = os.path.expanduser(os.path.join(data_dir, "cora/cora.content"))
node_attr = pd.read_csv(cora_data_location, sep='\t', header=None)
values = { str(row.tolist()[0]): row.tolist()[-1] for _, row in node_attr.iterrows()}
nx.set_node_attributes(g_nx, values, 'subject')
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
node_data = pd.read_table(os.path.join(data_dir, "cora/cora.content"), header=None, names=column_names)
# +
g_nx_ccs = (g_nx.subgraph(c).copy() for c in nx.connected_components(g_nx))
g_nx = max(g_nx_ccs, key=len)
node_ids = list(g_nx.nodes())
print("Largest subgraph statistics: {} nodes, {} edges".format(
g_nx.number_of_nodes(), g_nx.number_of_edges()))
node_targets = [ g_nx.nodes[node_id]['subject'] for node_id in node_ids]
print(f"There are {len(np.unique(node_targets))} unique labels on the nodes.")
print(f"There are {len(g_nx.nodes())} nodes in the network.")
# +
s = set(node_data["subject"])
#build a dictionary to convert string to numbers
convert_table = {e:idx for idx, e in enumerate(s)}
def word2idx(word):
return convert_table[word]
ground_truth = [word2idx(i) for i in node_targets]
# -
A = nx.to_numpy_array(g_nx)
D = np.diag(A.sum(axis=1))
print(D)
L = D-A
print(L)
# +
eigenvalues, eigenvectors = np.linalg.eig(L)
eigenvalues = np.real(eigenvalues)
eigenvectors = np.real(eigenvectors)
order = np.argsort(eigenvalues)
eigenvalues = eigenvalues[order]
# -
embedding_size = 32
v_0 = eigenvectors[:, order[0]]
v = eigenvectors[:, order[1:(embedding_size+1)]]
convert_table
# +
n = 7 #number of clusters
#Spectral Clustering method
model = SpectralClustering(n_clusters = n, n_init=100,assign_labels='discretize')
#model.fit(vecs[:,1])
labels = model.fit_predict(v[:,1:8])
#labels = [abs(i-1) for i in labels]
print(metrics.adjusted_rand_score(ground_truth, labels))
print(metrics.adjusted_mutual_info_score(ground_truth, labels))
print(metrics.accuracy_score(ground_truth, labels))
print(ground_truth)
print(labels)
# -
| term-paper-GNN/Part3_Cora_Spectral2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### 2.1: Bot Design Principles
# -
# This session focuses on the design aspects of bots with an emphasis on good bot design that has been established by conducting engagements with partners and customers. At the end of this session you will be able to:
#
# 1. Conduct effective design research.
# 2. Enhance and optimize conversational flow.
# 3. Map Bot capabilities to organizational objectives.
# ### Section 1: Effective Design Research
# In this section, you will explore the steps required to conduct effective design research, including:
#
# 1. Understanding the organizational requirements.
# 2. Understanding the types of conversation patterns.
# 3. Building a personality profile for your bot.
# ### Section 1.1: Understanding the organizational requirements.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# During the design phase, it is typical to perform a discovery workshop on the requirements for implementing a bot for the organization. The purpose of the workshop is to establish the problem/use case that the bot is going to solve, assess the desirability, feasibility and viability criteria, and ensure that the specific use cases/bot conversational flows and intents are appropriately captured.
#
# * Desirability = includes the human factors (is there a human need for this innovation? Or are we making this bot for the sake of making a bot?)
# * Feasibility = technically feasible (is the technology available to actually do this?)
# * Viability = financially viable (is this sustainable from a business perspective)
#
# Bots and Conversation as a Platform in general is a major technology shift and is considered a megatrend in the digital transformation space. Setting appropriate expectations, project goals and design guidelines are essential in ensuring that customers that embark on a conversation as a platform initiative drive the best value from their investment, and in-turn continue to build and grow their conversation platform initiative, driving future growth and continued end-user adoption.
#
# It is important to dedicate the right amount of focus on design and not get deep into Bot Architecture, Toolkits and other tech-led conversations. This is not just a technology, it facilitates interaction, a human-to-computer interaction. It’s an experience! which is why we have UX and Architect leading the design conversations with other business stakeholders, including:
#
# * Project Sponsor and other business leaders that can speak to the desired outcome from the engagement and conditions of satisfaction
# * Business Domain experts specific to the scenario and use cases being explored with Bot.
# * Subject Matter experts that can speak to the knowledge domain and are familiar with the conversations we expert bots to have with end-users
# * Technical leads that understand the overall Systems Architecture of knowledge sources and can cover any aspect of backend knowledge/integration possibilities
# * Core Project team members, including Project Managers and Technical Consultants to ensure they are aligned with overall design requirements and business expectations.
#
# The following are some general guidelines on running a workshop. These are harnessed through previous engagements conducted by the Microsoft Services teams.
#
# * Start the discussion broadly with Design Led Thinking making sure Desirability, Feasibility and Viability criteria are top of mind.
# * Use existing logs/fact based analysis for driving towards top intents and key uses cases.
# * Use the Pareto Principle to make the cases if applicable
# * In a complete green-field scenario, start by defining the personas and then use the lifecycle of the personas to identifying specific use-cases
# * Gauge the level of understanding around Bot terminologies in the audience and feel free to set the stage with key terms and taxonomies, this allows the audience to stay engaged during the workshop and not get lost in taxonomies.
# ### Section 1.2: Understanding the types of conversation patterns.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# While working on defining use cases, there is a varying level of complexity involved with certain types of conversational flows. For example, a one-turn dialog that is simply responding by pairing an answer to a question (or set of questions) is a low complexity use case. A multi-turn dialog that needs to extract multiple entities, has to remain context sensitive and is doing some type of task completion which also requires backend integration is a high complexity use case. Then there are other patterns that fall somewhere in between these two extremes. To manage priorities across use cases and overall acceptance criteria for the project, within the constraints of the project scope, it is important to bring in a complexity discussion while walking through the use cases.
#
# One way to manage these conversations is by defining a set of conversational patterns that are organized by the level of complexity where each use case, as it is discovered, is mapped to one of those conversational patterns. These can be organized as follow:
#
# * One-Turn FAQ
# These are general FAQ style answers that is based on one-turn responses.
#
# * Intelligent Notification
# This is a notification pattern because it is not in response to any query initiated by the user. Design guidance suggests caution when it comes to the bot providing proactive notification.
#
# * One-Turn Intelligent Response
# These are also one-turn responses which don’t typically have a follow-up or additional conversational flow. These are “intelligent” terms as they need to be provided within the context of a conversation or activity. For example, a customer may ask, what time does flight XX065 leave LAX today?
#
# * Contextual Guided Assistance
# In this scenario, context is associated with the page/url the user is on as opposed to who the user is and would like the bot to be able to answer questions within this context of the page/url.
#
# * Multi-Turn Process Guidance
# This is a unique scenario within the users domain - there are a number of process flows established that a bot can be enabled to do a user walk-through. While these are multi-turn dialog scenarios, they are not truly conversational as the user responses are restricted to Yes/No only with potentially an option to opt out or cancel the guided assistance.
#
# * Multi-Turn Conversational Task Completion
# This is a multi-turn conversational pattern typically leading to a task completion scenario. In this type of pattern, the user is not restricted to fixed responses and can potentially provide free-form natural language response. The bot will need to interpret this within the context of the process the user is in.
#
# ### Section 1.3: Building a personality profile for you bot.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# Would you trust someone not in your organization to speak on behalf of your organization?
#
# Your bot is going to be the spokesperson for your company. That’s why it is important to have a mix of business stakeholders ranging from UX leads, project owners and other stakeholders in the design phase of the bot. Most customers are looking at either knowledge orientated or task completion bots, and expect bots to be able to answer general knowledge questions, integrating that is not a big deal if you use knowledge search. What is often overlooked is the personality of the bot. Personality makes a difference, even if it's in subtle ways. Remember that it will be representing your brand, and it is important to consider the personality profile so that it represents your company at its’ best.
#
# Options to consider when defining a bot personality profile include:
#
# * Demographics
# * BIO
# * Goals of the bot
# * Pain Points of the bot
# * Personality
# * Character
# * Target users
# ### Section 1.4: Discussion. Bottlenecks to Effective Design Research?
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# The following are potential answers to the questions, but we encourage you to come up with more.
#
# What factors could affect effective design research?
#
# There are a range of answers here that could impact the effective design research including the following:
# Project lacks diversity and depth for a full design review.
# Time limitations leading to an incomplete or partial research.
# A lack of understanding of the corporate, transformation or project requirements
# Lack of support from senior stakeholders
#
#
# How would you overcome such factors?
#
# * Project lacks diversity and depth for a full design review.
# Consider seeking buy in from senior stakeholders to allow a variety of people with different backgrounds to take part in the review. PR and marketing professionals can be very helpful, as well as technologists and Project Managers. Consider what the actions are of the bot and seek an understanding of the role. For example, if you were creating a concierge service for a hotel, you should include people from that role.
#
# * Time limitations leading to an incomplete or partial research.
# It’s important to set the correct expectations for how long effective design research will take. It is not uncommon for a number of meetings to take place as research activities are delegated to members of the team to research and report back. However, having a deadline is also important for setting a line in the sand. Without this control, the research phase could go on and on
#
# * A lack of understanding of the corporate, transformation or project requirements
# It is important that there is a clear understanding of the corporate, transformation or project requirements. This acts as the north star for the project and should include measures for success. This should be the first aspect of the effective design research phase that should be completed and is of the highest priority. If there is still uncertainty with goals, revisit them and investigate.
#
# * Lack of support from senior stakeholders
# This is perhaps the most difficult to solve should the stakeholders in question not see any benefits to the solution and serious impacts the desirability criteria. If this is the case, it may be more pragmatic to shelve the project until there is more desire. However, before giving up, make sure that the senior stakeholders in question understand the commercial benefits of taking on the project. Be it to reduce operational costs, improve customer service (and therefore retain more new customers), or use the bot for upsell opportunities.
#
#
# * Bot personality profiling may be viewed as non serious. How would you address this mindset?
# As a technologist or Project Manager, you wouldn’t. You would lean on the expertise of the PR and marketing team to communicate the importance of the bot representing the company and how it would come across. This enables their expertise to come to the fore and cement the reason for their presence on the team.
#
# ### Section 2: Enhance and optimize conversational flow
# In this section, you will explore the steps required to conduct effective design research, including:
#
# 1. Microsoft Bot Framework Capabilities
# 2. Supporting technologies
# ### Section 2.1: Microsoft Bot Framework Capabilities.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# The Microsoft bot framework is the #1 bot framework in the market today. This framework provides 3 key capabilities for developers. This open source framework allows developers to build bots in the languages they know and love - .NET, Node.js, Python, or Java The framework provides connectors to over 15 different channels. This allows developers to build a bot once, and connect it to SMS, Email, Facebook, skype, slack, kik, web chat, etc. Finally the framework allows you to publish bots to be discovered via search, Cortana and other web services. It is important to establish the skillsets of those who work at the company. From this perspective it can be established if the bot solution can be developed in-house, or establish the level of support that is required for the project to succeed. Furthermore, understanding the differences between each channel that will be used for the bot.
# ### Section 2.2: Supporting technologies.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# There are a range of supporting technologies that can be integrated into the bot framework to enhance and optimize the conversational flow. Understanding the capabilities of each technology is important as it can help you map a technology to a conversation pattern. The image shows the mappings that can be used for conversational flow and technology.
#
# * [QnA Maker](https://www.qnamaker.ai/)
# * [FormFlow](https://docs.microsoft.com/en-us/azure/bot-service/dotnet/bot-builder-dotnet-formflow)
# * [LUIS](https://www.luis.ai/)
# * [Machine Learning](https://docs.microsoft.com/en-us/azure/machine-learning/)
# * [Cognitive Services](https://azure.microsoft.com/en-gb/services/cognitive-services/)
# ### Section 3: Map Bot capabilities to organizational objectives
# In this section, you will explore examples of mapping Bot capability to organizational objectives, including:
#
# * Microsoft Bot Logic Flow
# * Bot Evolution Roadmap
# ### Section 3.1: Microsoft Bot Logic Flow.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# The slide shows an example of how you can map the domain and use case intents at a high level. Remember that the intention here is not to focus on the technology, but more on the business domain, intents and potential conversational flow. In addition, you can annotate the diagram to outline any additional requirements such a personalization. The High Level Bot Logic Flow does not have to use this format, but it is important to ensure that the format is understood by the intended audience. Furthermore, this document is not fixed and the expectation should be set with the customer that the initial findings could change with further investigation.
# ### Section 3.2: Bot Evolution Roadmap.
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# The development of bots will likely be an evolution, not a revolution. It may not be possible to implement the solution in one release. It is therefore pragmatic to develop a roadmap of the functionality of the solution. The slide shows an example of a roadmap.
# ### Section 3.3: Discussion. The importance of Stakeholder Sign-off
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# The following are potential answers to the questions, but we encourage you to come up with more.
#
# How would you deal with individuals who required more detail than the High Level Bot Flow Logic would provide?
# Outline to those concerned that the purpose of the effective design research is to establish the high level objectives of the project, understand the personality of the bot, and get an understanding of the tasks that it is trying to solve so the team can start to get an understanding of what technologies may be used. The output is to create a high level bot logic flow and a roadmap with approximate timescales in the first instance. At this stage, a discovery exercise is being performed on what could be possible and more details will be flushed out in other portions of the project such as the LUIS schema design and the physical architectures.
#
# What artefacts in addition to those seen in this module would you require to ensure sign-off?
# A statement of work with signatures of senior stakeholders
| 02-bot_design/1_session.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width="100%"> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="35%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <NAME> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# <br>
# updated by <NAME> | July 05, 2020
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2>Inversion About the Mean</h2>
# We play a simple game to give some ideas about how Grover's search algorithm works.
#
# We have a list of N elements.
#
# Some of them are marked ones.
#
# At the beginning each has a value of 1.
#
# Each iteration of the game has two phases:
#
# <ol>
# <li><b>Query</b>: In this phase, each marked element is detected, and then its sign is flipped.</li>
# <li><b>Inversion</b>: In this phase, the value of each element is reflected over the mean of all values.</li>
# </ol>
# <h3>Task 1</h3>
#
# We play this game for $ N = 8 $.
#
# Suppose that only the 4th element is marked.
#
# We can visualize the values of elements in the list in the beginning as follows.
# +
from matplotlib.pyplot import bar
labels = []
L = []
for i in range(8):
labels = labels + [i+1]
L = L + [1]
# visualize the values of elements in the list
bar(labels,L)
# -
# Iterate the game for one step and visualize the values of elements in the list after each phase.
# 1st step - query phase:
# +
#
# 1st step - query
#
# visualize the values of elements in the list
bar(labels,L)
# -
# 1st step - inversion phase:
# +
#
# 1st step - inversion
#
# visualize the values of elements in the list
bar(labels,L)
# -
# Iterate the game for one more step and visualize the values of elements in the list after each phase.
# 2nd step - query phase:
# +
#
# 2nd step - query
#
# visualize the values of elements in the list
bar(labels,L)
# -
# 2nd step - inversion phase:
# +
#
# 2nd step - inversion
#
# visualize the values of elements in the list
bar(labels,L)
# -
# Iterate the game three more steps and visualize the values of elements in the list at the end.
# +
#
# your code is here
#
# visualize the values of elements in the list
bar(labels,L)
# -
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task1">click for our solution</a>
# <h3>Task 2</h3>
#
# We play this game for $ N = 16 $.
#
# Suppose that only the 11th element is marked.
#
# Play the game for 20 iterations, and print the value of the 11th element after each phase of every iteration.
#
# your code is here
#
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task2">click for our solution</a>
# <b> Observations: </b>
#
# The absolute value of the marked element is increasing and decreasing during the iterations.
#
# Its behavior is similar to rotations.
# <h3> Modified Game </h3>
#
# We modify the game by guaranteeing that the list represents a quantum state.
# <h3> Task 3</h3>
#
# What are the initial values for the modifed game if $ N=8 $?
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task3">click for our solution</a>
# <h3> Task 4</h3>
#
# Iterate the modified game for $ N = 8 $ where the second element is the only marked element.
#
# Print the list after each phase.
#
# Check whether the length of list is 1 after each iteration.
#
# your code is here
#
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task4">click for our solution</a>
# <h3> Task 5</h3>
#
# Repeat Task 4 for $ N = 16 $ where the marked elements are the first, third and tenth elements.
#
# your code is here
#
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task5">click for our solution</a>
# <h3> Task 6</h3>
#
# Repeat Task 4 for $ N = 16 $ where the first 12 elements are marked.
#
# your code is here
#
# <a href="B80_Inversion_About_the_Mean_Solutions.ipynb#task6">click for our solution</a>
| bronze/B80_Inversion_About_the_Mean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import turtle
def turtleReset():
turtle.reset()
turtle.mode("logo")
turtle.degrees()
turtle.up()
turtle.setposition(0,-200)
turtle.down()
turtle.tracer(5, 25)
# # Pseudocode for Triangle
# - Start: F
# - Rules:
# - F=G-F-G
# - G=F+G+F
# - Symbols:
# - F: Go forward
# - G: Go forward
# - +: Turn right n degrees
# - -: Turn left n degrees
# - Angle: 60
def buildTriangleString(nIterations):
string = 'F'
for i in range(nIterations):
string = string.replace('F', 'g-f-g').replace('G', 'f+g+f').upper()
return string
LENGTH = 5
DEBUG = False
def renderTriangle(nIterations):
turtleReset()
string = buildTriangleString(nIterations)
print(string)
for char in string:
if char is 'F' or char is 'G':
if DEBUG:
print('Advancing')
turtle.forward(LENGTH)
elif char is '+':
turtle.right(60)
if DEBUG:
print('Rotating right to {}'.format(turtle.heading()))
elif char is '-':
turtle.left(60)
if DEBUG:
print('Rotating left to {}'.format(turtle.heading()))
renderTriangle(7)
# # Tree pseudocode
# - Start: F
# - Rules:
# - F\[+F\]F\[-F\]\[F\]
# - Symbols:
# - [: Remember this location
# - ]: Teleport back to the most recent location
# - Angle: 20
def buildTreeString(nIterations):
string = 'F'
for i in range(nIterations):
string = string.replace('F', 'F[+F]F[-F][F]')
return string
LENGTH = 10
def renderTree(nIterations):
turtleReset()
string = buildTreeString(nIterations)
print(string)
myStack = []
for char in string:
if char is 'F':
turtle.forward(LENGTH)
elif char is '+':
turtle.right(20)
elif char is '-':
turtle.left(20)
elif char is '[':
myStack.append((turtle.pos(), turtle.heading()))
elif char is ']':
nPos, nHead = myStack.pop()
turtle.up()
turtle.setposition(nPos)
turtle.setheading(nHead)
turtle.down()
renderTree(5)
# # Random Examples
turtle.reset()
def sq():
turtle.fd(100)
turtle.rt(90)
turtle.fd(100)
turtle.rt(90)
turtle.fd(100)
turtle.rt(90)
turtle.fd(100)
turtle.rt(90)
for i in range(72):
sq()
turtle.rt(5)
turtle.fd(20)
def polyspi(angle,inc,side,times):
if times > 0:
turtle.fd(side)
turtle.rt(angle)
polyspi(angle,inc, (side + inc),(times - 1))
turtle.reset()
polyspi(90,5,50,50)
turtle.reset()
polyspi(95,1,50,100)
| Jupyter/Turtle-Graphics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from helpers.utilities import *
# %run helpers/notebook_setup.ipynb
# %R source('plots/colors.R');
# + tags=["parameters"]
clinical_path = 'data/clean/clinical/data_with_derived_variables.csv'
zz_log_path = 'data/clean/protein/zz_log_10.csv'
# -
# I am using double z-score transformed log10 abundance levels here. This has an advantage of reducing the effect of technical variation (i.e. more material taken from a patient) and in the second step - centering the protein levels around the mean (so that we can separate easily into two groups i.e. low and high),
clinical = read_csv(clinical_path, index_col=0)
protein_levels = read_csv(zz_log_path, index_col=0)
# ## Survival analysis
# Also see the survival analysis on clinical variables only: [Clinical_survival.ipynb](../Clinical_survival.ipynb).
# How feasible is survival analysis for protein data?
sum(~clinical['survival'].isnull())
# Feasible (15/22 patients ho have deceased have protein data), but any split (esp. other than in half) will have little power, though the censored data may be informative too.
# Important piece: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3071962/
pd.options.mode.chained_assignment = None
c = clinical[['censored_survival', 'Death', 'Sex', 'HIVResult', 'Meningitis', 'Tuberculosis']]
data = concat([protein_levels.T, c], axis=1)
# + language="R"
# library("survminer")
# library("survival")
# source('helpers/survival.R')
# -
data['Albumin_high_low'] = (data['Albumin'] > 0).map({True: 'High', False: 'Low'})
# + magic_args="-i data" language="R"
# fit <- survfit(Surv(data$censored_survival, data$Death) ~ Albumin_high_low, data=data)
# strata = strip_strata_prefix(fit)
#
# ggsurvplot(
# fit, data=data,
# legend.labs=strata,
# risk.table=T, ggtheme=theme_bw()
# )
| analyses/protein_vs_clinical/Survival.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_project_X
# language: python
# name: venv_project_x
# ---
# # Time Series Forecasting in Python
# https://www.analyticsvidhya.com/blog/2016/02/time-series-forecasting-codes-python/
# ## Loading and Handling Time Series in Pandas
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from statsmodels.tsa.stattools import adfuller
from statsmodels.tsa.stattools import acf, pacf
from statsmodels.tsa.arima_model import ARIMA
# Now, we will load the data set and look at some initial rows and data types of the columns:
data = pd.read_csv('data/AirPassengers.csv')
print (data.tail())
print ('\n Data Types:')
print (data.dtypes)
import datetime
# The data contains a particular month and number of passengers travelling in that month. In order to read the data as a time series, we have to pass special arguments to the read_csv command:
dateparse = lambda dates: pd.datetime.strptime(dates, '%Y-%m')
data = pd.read_csv('data/AirPassengers.csv', parse_dates=['Month'], index_col='Month', date_parser=dateparse)
print ('\n Parsed Data:')
print (data.head())
data.index
data['#Passengers']
ts = data["#Passengers"]
ts.head(10)
ts['1949-01-01']
ts[:"1949-05-01"]
# ## Check for stationarity
# plot the time series
plt.plot(ts)
plt.show()
# function to compute rolling stats and Dickey-Fuller test
def test_stationarity(timeseries):
# Determing rolling statistics
# rolmean = pd.rolling_mean(timeseries, window=12)
# rolstd = pd.rolling_std(timeseries, window=12)
rolmean = timeseries.rolling(12).mean()
rolstd = timeseries.rolling(12).std()
#Plot rolling statistics:
orig = plt.plot(timeseries, color='blue',label='Original')
mean = plt.plot(rolmean, color='red', label='Rolling Mean')
std = plt.plot(rolstd, color='black', label = 'Rolling Std')
plt.legend(loc='best')
plt.title('Rolling Mean & Standard Deviation')
plt.show(block=False)
#Perform Dickey-Fuller test:
print('Results of Dickey-Fuller Test:')
dftest = adfuller(timeseries, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
print(dfoutput)
# compute statistics on the time series
test_stationarity(ts)
# # Processing the series
# # Estimating the trend
# take the log
ts_log = np.log(ts)
plt.plot(ts_log); plt.show()
# let's take a difference
ts_diff = ts.diff()
plt.plot(ts_diff)
plt.show()
ts_diff['1949-01-01'] = 0.
ts_diff.head()
# compute dickey-fuller test on difference time series
test_stationarity(ts_diff)
# ## Moving average
moving_avg = ts_log.rolling(12).mean()
plt.plot(ts_log)
plt.plot(moving_avg, color='red')
plt.show()
ts_log_moving_avg_diff = ts_log - moving_avg
ts_log_moving_avg_diff.dropna(inplace=True)
test_stationarity(ts_log_moving_avg_diff)
# ## Exponentially weighted moving average
expwighted_avg = ts_log.ewm(halflife=12).mean()
expwighted_avg.head()
plt.plot(ts_log)
plt.plot(expwighted_avg, color='red')
plt.show()
ts_log_ewma_diff = ts_log - expwighted_avg
test_stationarity(ts_log_ewma_diff)
# # Eliminating Trend and Seasonality
# ## Differencing
# first order difference of the log series
ts_log_diff = ts_log - ts_log.shift()
plt.plot(ts_log_diff)
plt.show()
ts_log_diff.dropna(inplace=True)
test_stationarity(ts_log_diff)
# ## Decompose trend and seasonality
# +
from statsmodels.tsa.seasonal import seasonal_decompose
decomposition = seasonal_decompose(ts_log)
trend = decomposition.trend
seasonal = decomposition.seasonal
residual = decomposition.resid
plt.subplot(411)
plt.plot(ts_log, label='Original')
plt.legend(loc='best')
plt.subplot(412)
plt.plot(trend, label='Trend')
plt.legend(loc='best')
plt.subplot(413)
plt.plot(seasonal,label='Seasonality')
plt.legend(loc='best')
plt.subplot(414)
plt.plot(residual, label='Residuals')
plt.legend(loc='best')
plt.tight_layout()
plt.show()
# -
ts_log_decompose = residual
ts_log_decompose.dropna(inplace=True)
test_stationarity(ts_log_decompose)
# # Forecasting a Time Series
# ## ACF and PACF plots
lag_acf = acf(ts_log_diff, nlags=20)
lag_pacf = pacf(ts_log_diff, nlags=20, method='ols')
# +
#Plot ACF:
plt.subplot(121)
plt.plot(lag_acf)
plt.axhline(y=0,linestyle='--',color='gray')
plt.axhline(y=-1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.axhline(y=1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.title('Autocorrelation Function')
#Plot PACF:
plt.subplot(122)
plt.plot(lag_pacf)
plt.axhline(y=0,linestyle='--',color='gray')
plt.axhline(y=-1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.axhline(y=1.96/np.sqrt(len(ts_log_diff)),linestyle='--',color='gray')
plt.title('Partial Autocorrelation Function')
plt.tight_layout()
# -
# ## ARIMA (2, 1, 0)
model = ARIMA(ts_log, order=(2, 1, 0))
results_AR = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_AR.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_AR.fittedvalues-ts_log_diff)**2))
# ## ARIMA (0, 1, 2)
model = ARIMA(ts_log, order=(0, 1, 2))
results_MA = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_MA.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_MA.fittedvalues-ts_log_diff)**2))
# ## ARIMA (2, 1, 2)
model = ARIMA(ts_log, order=(2, 1, 2))
results_ARIMA = model.fit(disp=-1)
plt.plot(ts_log_diff)
plt.plot(results_ARIMA.fittedvalues, color='red')
plt.title('RSS: %.4f'% sum((results_ARIMA.fittedvalues-ts_log_diff)**2))
# ## Take the value back to the original scale
predictions_ARIMA_diff = pd.Series(results_ARIMA.fittedvalues, copy=True)
predictions_ARIMA_diff_cumsum = predictions_ARIMA_diff.cumsum()
predictions_ARIMA_diff_cumsum.head()
predictions_ARIMA_log = pd.Series(ts_log[0], index=ts_log.index)
predictions_ARIMA_log = predictions_ARIMA_log.add(predictions_ARIMA_diff_cumsum, fill_value=0)
predictions_ARIMA_log.head()
predictions_ARIMA = np.exp(predictions_ARIMA_log)
plt.plot(ts)
plt.plot(predictions_ARIMA)
plt.title('RMSE: %.4f'% np.sqrt(sum((predictions_ARIMA-ts)**2)/len(ts)))
results_ARIMA.predict('1961-01')
# ## Make one step ahead predictions
# compute predictions
model = ARIMA(ts_log, order=(2, 1, 2))
results = model.fit(disp=-1)
preds = results.predict(start='1959-01', dynamic=False)
ts_log['1958-12'][0]
# +
# Correct for first order difference
preds_ARIMA_diff = pd.Series(preds, copy=True)
preds_ARIMA_diff_cumsum = preds_ARIMA_diff.cumsum()
preds_ARIMA_log = pd.Series(ts_log[119], index=ts_log.index[120:])
preds_ARIMA_log = preds_ARIMA_log.add(preds_ARIMA_diff_cumsum, fill_value=0)
preds_ARIMA_log.head()
# +
# correct for logarithm
preds_ARIMA = np.exp(preds_ARIMA_log)
# -
plt.plot(ts)
plt.plot(preds_ARIMA)
plt.title('RMSE: %.4f'% np.sqrt(sum((predictions_ARIMA-ts)**2)/len(ts)))
# ## Make forecasts
print(type(results_ARIMA.fittedvalues))
print(results_ARIMA.fittedvalues)
predictions = pd.Series(results_ARIMA.fittedvalues, copy=True)
print(type(predictions))
print(predictions[0])
print("")
print(predictions.index)
forecasts = pd.Series.copy(predictions[0:5])
print(forecasts)
forecasts[0] = 111
print(predictions[0])
print("")
print(forecasts[0])
results_ARIMA.predict('1961')
| TimeSeriesForecastingARIMA.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
// # STIL Integration
//
// [STIL](http://www.star.bristol.ac.uk/~mbt/stil/), the Starlink Tables Infrastructure Library, is a Java API for working with astronomical data, including VOTable, FITS, SQL, ASCII, CSV, CDF, and GBIN formats. This notebook shows how to load STIL, and configure BeakerX to display STIL StarTables with the BeakerX interactive table widget.
// +
// %classpath add mvn commons-io commons-io 2.6
import org.apache.commons.io.FileUtils
stilUrl = "http://www.star.bristol.ac.uk/~mbt/stil/stil.jar"
stilFile = System.getProperty("java.io.tmpdir") + "/stilFiles/stil.jar"
FileUtils.copyURLToFile(new URL(stilUrl), new File(stilFile));
// %classpath add dynamic stilFile
// +
import uk.ac.starlink.table.StarTable
import uk.ac.starlink.table.Tables
import jupyter.Displayer
import jupyter.Displayers
Displayers.register(StarTable.class, new Displayer<StarTable>() {
def getColumnNames(t){
names = []
nCol = t.getColumnCount();
for ( int icol = 0; icol < nCol; icol++ ) {
names.add(t.getColumnInfo(icol).getName())
}
names
}
@Override
public Map<String, String> display(StarTable table) {
columnNames = getColumnNames(table)
columnInfos = Tables.getColumnInfos(table)
MAXCHAR = 64
new TableDisplay(
(int)table.getRowCount(),
(int)table.getColumnCount(),
columnNames,
new TableDisplay.Element() {
@Override
public String get(int columnIndex, int rowIndex) {
Object cell = table.getCell(rowIndex, columnIndex);
return columnInfos[columnIndex].formatValue(cell, MAXCHAR)
}
}
).display();
return OutputCell.DISPLAYER_HIDDEN;
}
});
// +
import org.apache.commons.io.FileUtils
messierUrl = "http://andromeda.star.bristol.ac.uk/data/messier.csv"
messierFile = System.getProperty("java.io.tmpdir") + "/stilFiles/messier.csv"
FileUtils.copyURLToFile(new URL(messierUrl), new File(messierFile));
"Done"
// +
import uk.ac.starlink.table.StarTable
import uk.ac.starlink.table.StarTableFactory
import uk.ac.starlink.table.Tables
starTable = new StarTableFactory().makeStarTable( messierFile, "csv" );
starTable = Tables.randomTable(starTable)
| doc/groovy/STIL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from switss.model import DTMC, ReachabilityForm
from switss.solver import MILP, LP
from switss.problem import MILPExact, Subsystem, QSHeur
# +
P = [[0.3, 0.7, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0],
[0.0, 0.1, 0.0, 0.7, 0.0, 0.1, 0.0, 0.0, 0.1, 0.0],
[0.0, 0.1, 0.0, 0.0, 0.0, 0.1, 0.0, 0.0, 0.8, 0.0],
[0.0, 0.2, 0.0, 0.4, 0.2, 0.0, 0.1, 0.1, 0.0, 0.0],
[0.0, 0.0, 0.0, 0.0, 0.7, 0.0, 0.0, 0.1, 0.2, 0.0],
[0.0, 0.0, 0.0, 0.1, 0.0, 0.8, 0.0, 0.0, 0.1, 0.0],
[0.0, 0.0, 0.7, 0.3, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0],
[0.0, 0.0, 0.0, 0.0, 0.3, 0.0, 0.0, 0.6, 0.0, 0.1],
[0.0, 0.0, 0.0, 0.0, 0.0, 0.9, 0.0, 0.0, 0.1, 0.0],
[0.0, 0.0, 0.1, 0.9, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]]
labels = { "target" : {8},
"init" : {0},
"group1" : {1,3,6},
"group2" : {7,9,2},
"group3" : {4,5}
}
mc = DTMC(P, labels)
rf, _, _ = ReachabilityForm.reduce(mc, "init", "target")
rf.system.digraph()
# -
# Try to find a subsystem that has probability at least 0.5 and sees as few groups as possible:
qs_min_heur = QSHeur(solver="cbc")
result = qs_min_heur.solve(rf, 0.5, "min", labels=["group1", "group2", "group3"])
result.subsystem.digraph()
qs_min_heur = QSHeur(solver="cbc")
result = qs_min_heur.solve(rf, 0.5, "max")
result.subsystem.digraph()
| examples/groups.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How many children, 16 and under, have been shot in the 11th District each year since 2012?
# We're going to do this two ways. First, by checking if the coordinates in each record are in the District 11 boundary and then by checking the explicit `District` column in each record.
# ## Load the shooting victims data from NewsroomDB
#
# NewsroomDB is the Tribune's proprietary database for tracking data that needs to be manually entered and validated rather than something that can be ingested from an official source. It's mostly used to track shooting victims and homicides. As far as I know, CPD doesn't provide granular data on shooting victims and the definition of homicide can be tricky (and vary from source to source).
#
# We'll grab shooting victims from the `shootings` collection.
# +
import os
import requests
def get_table_url(table_name, base_url=os.environ['NEWSROOMDB_URL']):
return '{}table/json/{}'.format(os.environ['NEWSROOMDB_URL'], table_name)
def get_table_data(table_name):
url = get_table_url(table_name)
try:
r = requests.get(url)
return r.json()
except:
print("Request failed. Probably because the response is huge. We should fix this.")
return get_table_data(table_name)
shooting_victims = get_table_data('shootings')
print("Loaded {} shooting victims".format(len(shooting_victims)))
# -
# ## Load police district boundaries
# +
import requests
from shapely.geometry import shape
# The City of Chicago's Socrata-based Data Portal provides a GeoJSON export of its spatial datasets.
# We'll use this so we don't have to save spatial data to the repo.
POLICE_DISTRICT_BOUNDARIES_GEOJSON_URL = "https://data.cityofchicago.org/api/geospatial/fthy-xz3r?method=export&format=GeoJSON"
r = requests.get(POLICE_DISTRICT_BOUNDARIES_GEOJSON_URL)
police_districts = r.json()
# Get the District 11 GeoJSON feature
district_11_feature = next(f for f in police_districts['features']
if f['properties']['dist_num'] == "11")
# Convert it to a Shapely shape so we can detect our
district_11_boundary = shape(district_11_feature['geometry'])
# -
# ## Annotate and filter shooting victims
# +
from datetime import datetime
import re
import pandas as pd
def parse_date(s):
try:
return datetime.strptime(s, '%Y-%m-%d').date()
except ValueError:
return None
def parse_coordinates(coordinate_str):
"""Convert a lat, lng string to a pair of lng, lat floats"""
try:
lat, lng = [float(c) for c in re.sub(r'[\(\) ]', '', coordinate_str).split(',')]
return lng, lat
except ValueError:
return None
def parse_age(age_str):
try:
return int(age_str)
except ValueError:
return None
def get_year(shooting_date):
try:
return shooting_date.year
except AttributeError:
return None
shooting_victims_df = pd.DataFrame(shooting_victims)
shooting_victims_df['Date'] = shooting_victims_df['Date'].apply(parse_date)
shooting_victims_df['Age'] = shooting_victims_df['Age'].apply(parse_age)
shooting_victims_df['coordinates'] = shooting_victims_df['Geocode Override'].apply(parse_coordinates)
shooting_victims_df['year'] = shooting_victims_df['Date'].apply(get_year)
# -
child_shooting_victims = shooting_victims_df[shooting_victims_df['Age'] < 18]
child_shooting_victims_16_and_under = child_shooting_victims[child_shooting_victims['Age'] <= 16]
child_shooting_victims_since_2012 = child_shooting_victims[child_shooting_victims['year'] >= 2012]
child_shooting_victims_16_and_under_since_2012 = child_shooting_victims_16_and_under[child_shooting_victims_16_and_under['year'] >= 2012]
# +
from shapely.geometry import Point
def is_in_11th_district(shooting_coordinates):
try:
shooting_point = Point(shooting_coordinates[0], shooting_coordinates[1])
return district_11_boundary.contains(shooting_point)
except TypeError:
return False
child_shooting_victims_since_2012_in_11th_dist = child_shooting_victims_since_2012[
child_shooting_victims_since_2012['coordinates'].apply(is_in_11th_district)
]
child_shooting_victims_16_and_under_since_2012_in_11th_dist = child_shooting_victims_16_and_under_since_2012[
child_shooting_victims_16_and_under_since_2012['coordinates'].apply(is_in_11th_district)
]
print("There have been {} victims, under 18 years of age, who have been shot in the 11th district since 2012".format(
len(child_shooting_victims_since_2012_in_11th_dist)))
print("There have been {} victims, age 16 or under, who have been shot in the 11th district since 2012".format(
len(child_shooting_victims_16_and_under_since_2012_in_11th_dist)))
# -
# Sanity check our filter
# It looks like one of our rows has a district of 10. Maybe this is because of bad
# data entry
for i, victim in child_shooting_victims_16_and_under_since_2012_in_11th_dist.iterrows():
print(victim['District'])
child_shooting_victims_since_2012_in_11th_dist_by_year = pd.DataFrame(
child_shooting_victims_since_2012_in_11th_dist.groupby('year').size(),
columns=['num_victims']
)
child_shooting_victims_since_2012_in_11th_dist_by_year
child_shooting_victims_16_and_under_since_2012_in_11th_dist_by_year = pd.DataFrame(
child_shooting_victims_16_and_under_since_2012_in_11th_dist.groupby('year').size(),
columns=['num_victims']
)
child_shooting_victims_16_and_under_since_2012_in_11th_dist_by_year
# ## Let's just use the "District" column
#
# After doing all the spatial stuff, I realized there is a "District" column I could have used to filter. Note that in the above data, there is one row with a "District" column of "10". Perhaps this is because it was mislabeled.
# +
def is_11th_district(district):
try:
return int(district) == 11
except ValueError:
return False
child_shooting_victims_since_2012_in_11th_dist = child_shooting_victims_since_2012[child_shooting_victims_since_2012['District'].apply(is_11th_district)]
print("There have been {} victims, under age 18, who have been shot in the 11th district since 2012".format(
len(child_shooting_victims_since_2012_in_11th_dist)))
child_shooting_victims_16_and_under_since_2012_in_11th_dist = child_shooting_victims_16_and_under_since_2012[child_shooting_victims_16_and_under_since_2012['District'].apply(is_11th_district)]
print("There have been {} victims, under age 18, who have been shot in the 11th district since 2012".format(
len(child_shooting_victims_since_2012_in_11th_dist)))
# -
child_shooting_victims_since_2012_in_11th_dist_by_year = pd.DataFrame(
child_shooting_victims_since_2012_in_11th_dist.groupby('year').size(),
columns=['num_victims']
)
child_shooting_victims_since_2012_in_11th_dist_by_year
# So, it looks like using this method there are 3 more victims in 2014. This could be due to bad or missing coordinates in some of the rows that cause them to not be categorized when we use the coordinates to detect district.
child_shooting_victims_16_and_under_since_2012_in_11th_dist_by_year = pd.DataFrame(
child_shooting_victims_16_and_under_since_2012_in_11th_dist.groupby('year').size(),
columns=['num_victims']
)
child_shooting_victims_16_and_under_since_2012_in_11th_dist_by_year
| Child shooting victims in the 11th district.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import pickle
import numpy as np
from sklearn.metrics import matthews_corrcoef, confusion_matrix
from torch.utils.data import (DataLoader, RandomSampler, SequentialSampler,
TensorDataset)
from torch.nn import CrossEntropyLoss, MSELoss
from tqdm import tqdm_notebook, trange
import os
from pytorch_pretrained_bert import BertForSequenceClassification, BertForTokenClassification
from pytorch_pretrained_bert.optimization import BertAdam, WarmupLinearSchedule
from transformers import BertModel, BertTokenizer
from model.MedClinical import Biobert_fc
from multiprocessing import Pool, cpu_count
from util.tools import *
from util import convert_examples_to_features
# OPTIONAL: if you want to have more information on what's happening, activate the logger as follows
import logging
logging.basicConfig(level=logging.INFO)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# device = "cpu"
# +
DATA_DIR = "data/"
# Bert pre-trained model selected in the list: bert-base-uncased,
# bert-large-uncased, bert-base-cased, bert-large-cased, bert-base-multilingual-uncased,
# bert-base-multilingual-cased, bert-base-chinese.
BERT_MODEL = 'bert-base-cased'
# The name of the task to train.I'm going to name this 'yelp'.
TASK_NAME = 'Relation Extraction'
# The output directory where the fine-tuned model and checkpoints will be written.
OUTPUT_DIR = f'outputs/{TASK_NAME}/'
# The directory where the evaluation reports will be written to.
REPORTS_DIR = f'reports/{TASK_NAME}_evaluation_report/'
# This is where BERT will look for pre-trained models to load parameters from.
CACHE_DIR = 'cache/'
# The maximum total input sequence length after WordPiece tokenization.
# Sequences longer than this will be truncated, and sequences shorter than this will be padded.
MAX_SEQ_LENGTH = 128
TRAIN_BATCH_SIZE = 24
EVAL_BATCH_SIZE = 8
LEARNING_RATE = 1e-5
NUM_TRAIN_EPOCHS = 10
RANDOM_SEED = 42
GRADIENT_ACCUMULATION_STEPS = 1
WARMUP_PROPORTION = 0.1
CONFIG_NAME = "config.json"
WEIGHTS_NAME = "pytorch_model.bin"
# -
if os.path.exists(REPORTS_DIR) and os.listdir(REPORTS_DIR):
REPORTS_DIR += f'/report_{len(os.listdir(REPORTS_DIR))}'
os.makedirs(REPORTS_DIR)
if not os.path.exists(REPORTS_DIR):
os.makedirs(REPORTS_DIR)
REPORTS_DIR += f'/report_{len(os.listdir(REPORTS_DIR))}'
os.makedirs(REPORTS_DIR)
# tokenizer = BertTokenizer.from_pretrained(OUTPUT_DIR + 'vocab.txt', do_lower_case=False)
tokenizer = BertTokenizer.from_pretrained('bert-base-cased', do_lower_case=False)
processor = MultiClassificationProcessor()
eval_examples = processor.get_dev_examples(DATA_DIR)
eval_examples_len = len(eval_examples)
label_list = processor.get_labels() # [0, 1] for binary classification
num_labels = len(label_list)
num_labels
eval_examples_for_processing = [(example, MAX_SEQ_LENGTH, tokenizer) for example in eval_examples]
process_count = cpu_count() - 1
with Pool(process_count) as p:
eval_features = list(tqdm_notebook(p.imap(convert_examples_to_features.convert_example_to_feature, eval_examples_for_processing), total=eval_examples_len))
all_input_ids = torch.tensor([f.input_ids for f in eval_features], dtype=torch.long)
all_input_mask = torch.tensor([f.input_mask for f in eval_features], dtype=torch.long)
all_segment_ids = torch.tensor([f.segment_ids for f in eval_features], dtype=torch.long)
all_label_ids = torch.tensor([int(f.label_id) for f in eval_features], dtype=torch.long)
eval_data = TensorDataset(all_input_ids, all_input_mask, all_segment_ids, all_label_ids)
# Run prediction for full data
eval_sampler = SequentialSampler(eval_data)
eval_dataloader = DataLoader(eval_data, sampler=eval_sampler, batch_size=EVAL_BATCH_SIZE)
# +
# model = BertForSequenceClassification.from_pretrained(CACHE_DIR + BERT_MODEL, cache_dir=CACHE_DIR, num_labels=len(label_list))
# model = BertForSequenceClassification.from_pretrained(BERT_MODEL, cache_dir=CACHE_DIR, num_labels=num_labels)
# model = BertForSequenceClassification.from_pretrained(BERT_MODEL, cache_dir=CACHE_DIR, num_labels=num_labels)
model = Biobert_fc()
# model = BertModel.from_pretrained((BERT_MODEL))
path = OUTPUT_DIR + 'pytorch_model.bin'
model.load_state_dict(torch.load(path))
model.eval()
# -
model.to(device)
# +
model.eval()
eval_loss = 0
nb_eval_steps = 0
preds = []
for input_ids, input_mask, segment_ids, label_ids in tqdm_notebook(eval_dataloader, desc="Evaluating"):
input_ids = input_ids.to(device)
input_mask = input_mask.to(device)
segment_ids = segment_ids.to(device)
label_ids = label_ids.to(device)
with torch.no_grad():
logits = model(input_ids, segment_ids, input_mask)
# create eval loss and other metric required by the task
loss_fct = CrossEntropyLoss()
tmp_eval_loss = loss_fct(logits.view(-1, num_labels), label_ids.view(-1))
eval_loss += tmp_eval_loss.mean().item()
nb_eval_steps += 1
if len(preds) == 0:
preds.append(logits.detach().cpu().numpy())
else:
preds[0] = np.append(
preds[0], logits.detach().cpu().numpy(), axis=0)
eval_loss = eval_loss / nb_eval_steps
preds = preds[0]
preds = np.argmax(preds, axis=1)
# -
len(preds), len(all_label_ids)
# +
def get_eval_report(task_name, labels, preds):
mcc = matthews_corrcoef(labels, preds)
cm = confusion_matrix(labels, preds)
return {
"task": task_name,
"mcc": mcc,
"cm": cm
}
def compute_metrics(task_name, labels, preds):
assert len(preds) == len(labels)
return get_eval_report(task_name, labels, preds)
# -
import json
CONFIG_FOLDER = 'config/'
id_label_file = 'id_2_label.json'
with open(CONFIG_FOLDER + id_label_file) as infile:
id2label = json.load(infile)
# +
preds_labels = [id2label[str(p)] for p in preds]
all_labels = [id2label[str(l)] for l in all_label_ids.numpy()]
mcc = matthews_corrcoef(all_labels, preds_labels)
print('Correlation Coefficient is ', mcc)
mismatches = []
all_rels = []
for row in range(len(all_labels)):
all_rels.append([all_labels[row], preds_labels[row]])
if preds_labels[row] != all_labels[row]:
mismatches.append([all_labels[row], preds_labels[row]])
# +
# %matplotlib inline
from sklearn.metrics import plot_confusion_matrix
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sn
df = pd.DataFrame(all_rels, columns = ['labels', 'predicted'])
# df.head(10)
plt.figure(figsize=(24,14))
plt.title(" all relationships")
confusion_matrix = pd.crosstab(df['labels'], df['predicted'], rownames=['Actual'], colnames=['Predicted'])
sn.heatmap(confusion_matrix, annot=True)
plt.show()
# -
df
from sklearn import metrics
metrics.f1_score(df["labels"], df["predicted"], average='micro')
df["matched"] = df["labels"] == df["predicted"]
# df["nomatch"] = df["labels"] != df["predicted"]
df.groupby(["labels", "matched"]).count()
| project_re/BERT_RE_TACRED_eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import keras
# from keras import backend as K
# K.tensorflow_backend._get_available_gpus()
from keras.models import Sequential
from keras.layers import Dense, LSTM, Dropout, Flatten, Activation
from sklearn.preprocessing import MinMaxScaler
import numpy as np
import pandas as pd
from datetime import datetime
#import warnings
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
from sklearn.metrics import mean_squared_error
from math import sqrt
import time
def parser(x):
return datetime.strptime(x, '%d-%b-%Y')
def get_data(path, date):
data = pd.read_csv(path , header=0, parse_dates=[date], index_col=date, squeeze=True, date_parser=parser)
return data
class LSTM_RNN:
def __init__(self, look_back, batch_size, ip_size, dropout_probability = 0.2, init ='he_uniform', loss='mse', optimizer='adam'):
self.batch_size = batch_size
self.look_back = look_back
self.rnn = Sequential()
self.rnn.add(LSTM(units=256,input_shape=(None, ip_size), init=init, return_sequences=True))
self.rnn.add(Dropout(dropout_probability))
self.rnn.add(Dense(1))
self.rnn.add(Activation("linear"))
# self.rnn.add(Flatten())
self.rnn.add(Dense(1, init=init))
self.rnn.compile(loss=loss, optimizer=optimizer)
print(self.rnn.summary())
def train(self, X, Y, nb_epoch=150):
self.rnn.fit(X, Y, nb_epoch=nb_epoch, batch_size=self.batch_size, verbose=2)
def evaluate(self, X, Y):
score = self.rnn.evaluate(X, Y, batch_size = self.batch_size, verbose=0)
print(score)
return score
def predict(self, X):
return self.rnn.predict(X)
def evaluate_models(look_back, batch_size, trainX, trainY, testX, testY, ret=False):
print('Training & evaluating LSTM-RNN for batch size = ' + str(batch_size) + '...')
(_, _, j) = trainX.shape
if j > look_back:
lstm_model = LSTM_RNN(look_back, batch_size, ip_size=j)
else:
lstm_model = LSTM_RNN(look_back, batch_size, ip_size=look_back)
lstm_model.train(trainX, trainY)
lstm_test_mse = lstm_model.evaluate(testX, testY)
print("With batch size = " + str(batch_size) + ", Score: " + str(lstm_test_mse))
print('Completed model evaluation for batch size = ' + str(batch_size) + '...')
if ret:
yhat = lstm_model.predict(testX)
return lstm_test_mse, yhat
del lstm_model
return lstm_test_mse
def print_results(lstm_mse_vals):
print('Completed model evaluation for all lookback values...')
lstm_mse_min = min(lstm_mse_vals)
lstm_mse_argmin = np.argmin(lstm_mse_vals) + 1
print('Best mse with an LSTM recurrent neural network was ' + str(lstm_mse_min) + ' with a batch size of ' + str(lstm_mse_argmin))
return lstm_mse_argmin
# +
# #Ran this on kaggle to get an optimum value of 43
# mse_vals = []
# for batch_size in range(1, 51):
# np.random.seed(10)
# mse_vals.append(evaluate_models(look_back, batch_size, x, y, act, act_y))
# min_bs = print_results(mse_vals)
# np.random.seed(10)
# mse_val, y_hat = evaluate_models(look_back, min_bs, x, y, act, act_y, True)
# +
np.random.seed(10)
title = 'NIFTY LSTM'
look_back = 5
x = get_data('../ml-project-data/NIFTY_train.csv', 0)
y = x.loc[: , "High":"Low"].mean(axis=1)
data_y = y
x_lag = pd.DataFrame()
for i in range(look_back,0,-1):
x_lag['t-'+str(i)] = y.shift(i)
x = x_lag
x = x.iloc[look_back:]
y = y.iloc[look_back:]
act_data = get_data('../ml-project-data/NIFTY_test.csv', 0)
act = act_data.loc[: , "High":"Low"].mean(axis=1)
idx = act_data.index
act_y = act
act = pd.concat([y[-5:], act])
act_lag = pd.DataFrame()
for i in range(look_back,0,-1):
act_lag['t-'+str(i)] = act.shift(i)
act = act_lag[look_back:]
scaler = MinMaxScaler()
x = scaler.fit_transform(x.values)
y = scaler.fit_transform(y.values.reshape(-1, 1))
act = scaler.fit_transform(act.values)
act_y = scaler.fit_transform(act_y.values.reshape(-1, 1))
i, j = x.shape
x = x.reshape(1, i , j)
y = y.reshape(1, len(y), 1)
i,j = act.shape
act = act.reshape(1, i , j)
act_y = act_y.reshape(1, len(act_y), 1)
min_bs = 43
np.random.seed(10)
mse_val, y_hat = evaluate_models(look_back, min_bs, x, y, act, act_y, True)
y_hat = y_hat.reshape(len(y_hat[0]))
act = act_y.reshape(len(act_y[0]))
preds = pd.DataFrame(y_hat,columns=['Prediction'],index=idx)
actuals = pd.DataFrame(act,columns=['Actual'],index=idx)
preds = scaler.inverse_transform(preds)
actuals = scaler.inverse_transform(actuals)
plt.figure(figsize=(14,6))
plt.plot(idx, preds, label="Prediction")
plt.plot(idx, actuals, label="Actual")
plt.ylabel("Average Price")
plt.xlabel('Date')
plt.title(title)
plt.legend(loc="best")
plt.show()
rms = sqrt(mean_squared_error(actuals, preds))
print('RMSE: ' + str(rms))
mape = np.mean(np.abs((actuals - preds) / actuals)) * 100
print('MAPE: ' + str(mape))
# -
def company_without_nifty(company):
np.random.seed(10)
if(company == 1):
title = 'TCS LSTM w/o NIFTY'
tcs_x = get_data('../ml-project-data/TCS_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/TCS_test.csv', 2)['Average Price']
if(company == 2):
title = 'INFY LSTM w/o NIFTY'
tcs_x = get_data('../ml-project-data/INFY_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/INFY_test.csv', 2)['Average Price']
if(company == 3):
title = 'TECHM LSTM w/o NIFTY'
tcs_x = get_data('../ml-project-data/TECHM_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/TECHM_test.csv', 2)['Average Price']
if(company == 4):
title = 'HCL LSTM w/o NIFTY'
tcs_x = get_data('../ml-project-data/HCL_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/HCL_test.csv', 2)['Average Price']
if(company == 5):
title = 'WIPRO LSTM w/o NIFTY'
tcs_x = get_data('../ml-project-data/WIPRO_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/WIPRO_test.csv', 2)['Average Price']
look_back = 5
tcs_y = tcs_x
tcs_x_lag = pd.DataFrame()
for i in range(look_back,0,-1):
tcs_x_lag['t-'+str(i)] = tcs_y.shift(i)
tcs_x = tcs_x_lag
tcs_x = tcs_x.iloc[look_back:]
tcs_y = tcs_y.iloc[look_back:]
tcs_idx = tcs_act.index
tcs_act_y = tcs_act
tcs_act = pd.concat([tcs_y[-5:], tcs_act])
tcs_act_lag = pd.DataFrame()
for i in range(look_back,0,-1):
tcs_act_lag['t-'+str(i)] = tcs_act.shift(i)
tcs_act = tcs_act_lag[look_back:]
scaler = MinMaxScaler()
tcs_x = scaler.fit_transform(tcs_x.values)
tcs_y = scaler.fit_transform(tcs_y.values.reshape(-1, 1))
tcs_act = scaler.fit_transform(tcs_act.values)
tcs_act_y = scaler.fit_transform(tcs_act_y.values.reshape(-1, 1))
i, j = tcs_x.shape
tcs_x = tcs_x.reshape(1, i , j)
tcs_y = tcs_y.reshape(1, len(tcs_y), 1)
i,j = tcs_act.shape
tcs_act = tcs_act.reshape(1, i , j)
tcs_act_y = tcs_act_y.reshape(1, len(tcs_act_y), 1)
np.random.seed(10)
min_bs = 1
tcs_mse_val, tcs_y_hat = evaluate_models(look_back, min_bs, tcs_x, tcs_y, tcs_act, tcs_act_y, True)
tcs_y_hat = tcs_y_hat.reshape(len(tcs_y_hat[0]))
tcs_act = tcs_act_y.reshape(len(tcs_act_y[0]))
tcs_preds = pd.DataFrame(tcs_y_hat,columns=['Prediction'],index=tcs_idx)
tcs_actuals = pd.DataFrame(tcs_act,columns=['Actual'],index=tcs_idx)
tcs_preds = scaler.inverse_transform(tcs_preds)
tcs_actuals = scaler.inverse_transform(tcs_actuals)
plt.figure(figsize=(14,6))
plt.plot(tcs_idx, tcs_preds, label="Prediction")
plt.plot(tcs_idx, tcs_actuals, label="Actual")
plt.ylabel("Average Price")
plt.xlabel('Date')
plt.title(title)
plt.legend(loc="best")
plt.show()
rms = sqrt(mean_squared_error(tcs_actuals, tcs_preds))
print('RMSE: ' + str(rms))
mape = np.mean(np.abs((tcs_actuals - tcs_preds) / tcs_actuals)) * 100
print('MAPE: ' + str(mape))
# +
# mse_vals = []
# for batch_size in range(1, 51):
# np.random.seed(10)
# mse_vals.append(evaluate_models(look_back, batch_size, tcs_x, tcs_y, tcs_act, tcs_act_y))
# min_bs = print_results(mse_vals)
# -
company_without_nifty(1) #TCS
company_without_nifty(2) #INFY
company_without_nifty(3) #TECHM
company_without_nifty(4) #HCL
company_without_nifty(5) #WIPRO
def company_with_nifty(company):
np.random.seed(10)
if(company == 1):
title = 'TCS LSTM w NIFTY'
tcs_x = get_data('../ml-project-data/TCS_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/TCS_test.csv', 2)['Average Price']
if(company == 2):
title = 'INFY LSTM w NIFTY'
tcs_x = get_data('../ml-project-data/INFY_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/INFY_test.csv', 2)['Average Price']
if(company == 3):
title = 'TECHM LSTM w NIFTY'
tcs_x = get_data('../ml-project-data/TECHM_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/TECHM_test.csv', 2)['Average Price']
if(company == 4):
title = 'HCL LSTM w NIFTY'
tcs_x = get_data('../ml-project-data/HCL_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/HCL_test.csv', 2)['Average Price']
if(company == 5):
title = 'WIPRO LSTM w NIFTY'
tcs_x = get_data('../ml-project-data/WIPRO_train.csv', 2)['Average Price']
tcs_act = get_data('../ml-project-data/WIPRO_test.csv', 2)['Average Price']
look_back = 5
tcs_y = tcs_x
tcs_x_lag = pd.DataFrame()
for i in range(look_back,0,-1):
tcs_x_lag['t-'+str(i)] = tcs_y.shift(i)
tcs_x = tcs_x_lag
tcs_x = tcs_x.iloc[look_back:]
tcs_y = tcs_y.iloc[look_back:]
tcs_idx = tcs_act.index
tcs_act_y = tcs_act
tcs_act = pd.concat([tcs_y[-5:], tcs_act])
tcs_act_lag = pd.DataFrame()
for i in range(look_back,0,-1):
tcs_act_lag['t-'+str(i)] = tcs_act.shift(i)
tcs_act = tcs_act_lag[look_back:]
tcs_x['nifty'] = data_y
tcs_act['nifty'] = preds
scaler = MinMaxScaler()
tcs_x = scaler.fit_transform(tcs_x.values)
tcs_y = scaler.fit_transform(tcs_y.values.reshape(-1, 1))
tcs_act = scaler.fit_transform(tcs_act.values)
tcs_act_y = scaler.fit_transform(tcs_act_y.values.reshape(-1, 1))
i, j = tcs_x.shape
tcs_x = tcs_x.reshape(1, i , j)
tcs_y = tcs_y.reshape(1, len(tcs_y), 1)
i,j = tcs_act.shape
tcs_act = tcs_act.reshape(1, i , j)
tcs_act_y = tcs_act_y.reshape(1, len(tcs_act_y), 1)
np.random.seed(10)
min_bs = 49
tcs_mse_val, tcs_y_hat = evaluate_models(look_back, min_bs, tcs_x, tcs_y, tcs_act, tcs_act_y, True)
tcs_y_hat = tcs_y_hat.reshape(len(tcs_y_hat[0]))
tcs_act = tcs_act_y.reshape(len(tcs_act_y[0]))
tcs_preds = pd.DataFrame(tcs_y_hat,columns=['Prediction'],index=tcs_idx)
tcs_actuals = pd.DataFrame(tcs_act,columns=['Actual'],index=tcs_idx)
tcs_preds = scaler.inverse_transform(tcs_preds)
tcs_actuals = scaler.inverse_transform(tcs_actuals)
plt.figure(figsize=(14,6))
plt.plot(tcs_idx, tcs_preds, label="Prediction")
plt.plot(tcs_idx, tcs_actuals, label="Actual")
plt.ylabel("Average Value")
plt.xlabel('Date')
plt.title(title)
plt.legend(loc="best")
plt.show()
rms = sqrt(mean_squared_error(tcs_actuals, tcs_preds))
print('RMSE: ' + str(rms))
mape = np.mean(np.abs((tcs_actuals - tcs_preds) / tcs_actuals)) * 100
print('MAPE: ' + str(mape))
# +
# mse_vals = []
# for batch_size in range(1, 51):
# np.random.seed(10)
# mse_vals.append(evaluate_models(look_back, batch_size, tcs_x, tcs_y, tcs_act, tcs_act_y))
# min_bs = print_results(mse_vals)
# -
company_with_nifty(1) #TCS
company_with_nifty(2) #INFY
company_with_nifty(3) #TECHM
company_with_nifty(4) #HCL
company_with_nifty(5) #WIPRO
| ipython notebooks/Mutivariate-LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="agEHopLVkhki" colab_type="text"
# #Exploring Corpora with Text Normalization
# + [markdown] id="YTyX6w9SkwqJ" colab_type="text"
# In this assignment, I will perform text normalization, using Regular Expression, then explore corpora.
#
# The corpora are in Arabic language, and I want to find count of most 10 frequest words without text normalization and with text normalization.
#
# The text normalization includes
#
# 1. علامات الترقيم
# 2. الحركات
# 3. التنوين
#
#
# + [markdown] id="rKMdtQ9ksqDV" colab_type="text"
# ### Functions
# + id="fjH743cZstvb" colab_type="code" colab={}
import re
from google.colab import drive
def load_corpus(file_name):
text = open(file_name, encoding='utf-8').read()
return text
def words_count(text):
text = text.split()
word_counts = {}
for word in text:
word_counts[word] = word_counts.get(word, 0) + 1
return list(word_counts.items())
def regex(term):
return '["\',-\\/:،؛ًٌٍَُِّ]?' + term + '["\',-\\/:،؛ًٌٍَُِّ]?'
# + [markdown] id="NjtQ8kNyswaY" colab_type="text"
# ### Files
# + id="phX4Anies0EB" colab_type="code" colab={}
jsc_file = 'https://github.com/motazsaad/NLP-ICTS6361/blob/master/corpus/aljazeera.net_20190419_titles.txt'
cnn_file = 'https://github.com/motazsaad/NLP-ICTS6361/blob/master/corpus/arabic.cnn.com_20190419_titles.txt'
euro_file = 'https://github.com/motazsaad/NLP-ICTS6361/blob/master/corpus/arabic.euronews.com_20190409_titles.txt'
rt_file = 'https://github.com/motazsaad/NLP-ICTS6361/blob/master/corpus/arabic.rt.com_20190419_titles.txt'
bbc_file = 'https://github.com/motazsaad/NLP-ICTS6361/blob/master/corpus/bbc.com_20190409_titles.txt'
jsc = load_corpus(jsc_file)
cnn = load_corpus(cnn_file)
euro = load_corpus(euro_file)
rt = load_corpus(rt_file)
bbc = load_corpus(bbc_file)
# + [markdown] id="KOjdYZ8as2CA" colab_type="text"
# ### Count
# + id="sZXOBFvns3_Y" colab_type="code" colab={}
# count
jsc_count = words_count(jsc)
cnn_count = words_count(cnn)
euro_count = words_count(euro)
rt_count = words_count(rt)
bbc_count = words_count(bbc)
sorted_jsc_count = sorted([(v,k) for k,v in jsc_count], reverse=True)
sorted_cnn_count = sorted([(v,k) for k,v in cnn_count], reverse=True)
sorted_euro_count = sorted([(v,k) for k,v in euro_count], reverse=True)
sorted_rt_count = sorted([(v,k) for k,v in rt_count], reverse=True)
sorted_bbc_count = sorted([(v,k) for k,v in bbc_count], reverse=True)
# + [markdown] id="LngYCluJt5ng" colab_type="text"
# ### Explore
# + id="zf2cWZDGt6-g" colab_type="code" colab={}
norm_jsc_count = {}
norm_cnn_count = {}
norm_euro_count = {}
norm_rt_count = {}
norm_bbc_count = {}
for t in sorted_jsc_count[:10]:
word = t[1]
norm_jsc_count[word] = len(re.findall(regex(word), jsc))
for t in sorted_cnn_count[:10]:
word = t[1]
norm_cnn_count[word] = len(re.findall(regex(word), cnn))
for t in sorted_euro_count[:10]:
word = t[1]
norm_euro_count[word] = len(re.findall(regex(word), euro))
for t in sorted_rt_count[:10]:
word = t[1]
norm_rt_count[word] = len(re.findall(regex(word), rt))
for t in sorted_bbc_count[:10]:
word = t[1]
norm_bbc_count[word] = len(re.findall(regex(word), bbc))
# + [markdown] id="ZqPynhlNt9BI" colab_type="text"
# ### Results
# + [markdown] id="yslyIRDUuD6I" colab_type="text"
# #### JSC
# + [markdown] id="9nJPwQ15uJ64" colab_type="text"
# without text normalization
# + id="JwhcuaJQuNKx" colab_type="code" colab={}
print (sorted_jsc_count[:10])
# + [markdown] id="YmsigeEq50Ao" colab_type="text"
# [(21042, 'في'), (10199, 'من'), (6902, 'على'), (4616, 'مصر'), (3732, 'عن'), (3335, 'مقتل'), (2711, 'قتلى'), (2597, 'غزة'), (2359, 'مع'), (2349, 'إلى')]
# + [markdown] id="WvkgwJxAuPwI" colab_type="text"
# with text normalization
# + [markdown] id="NO29wszG549l" colab_type="text"
# [('في', 28108), ('من', 19135), ('على', 7163), ('مصر', 9002), ('عن', 5510), ('مقتل', 3463), ('قتلى', 3113), ('غزة', 3546), ('مع', 7303), ('إلى', 2349)]
# + id="wihE7c7ZuR5S" colab_type="code" colab={}
print (list(norm_jsc_count.items())[:10])
# + [markdown] id="F-f6oYE_uTly" colab_type="text"
# #### Euro
# + [markdown] id="4I-cQZhPucsI" colab_type="text"
# without text normalization
# + id="brl7Cn56udwJ" colab_type="code" colab={}
print (sorted_euro_count[:10])
# + [markdown] id="ab7F32-49YBi" colab_type="text"
# [(21101, 'في'), (8009, 'من'), (7730, 'على'), (3033, 'مع'), (2753, 'إلى'), (2701, 'شاهد:'), (2663, 'بعد'), (2550, 'عن'), (1725, 'ترامب'), (1321, 'السعودية')]
# + [markdown] id="YAG6oYBkugGJ" colab_type="text"
# with text normalization
# + id="QXDKh9LHugcI" colab_type="code" colab={}
print (list(norm_euro_count.items())[:10])
# + [markdown] id="obHoYFNT9a_d" colab_type="text"
# [('في', 26389), ('من', 13929), ('على', 8098), ('مع', 5970), ('إلى', 2769), ('شاهد:', 2718), ('بعد', 2845), ('عن', 3709), ('ترامب', 2149), ('السعودية', 1518)]
# + [markdown] id="05JV5nOsuoBr" colab_type="text"
# #### CNN
# + [markdown] id="CohUilXQuqew" colab_type="text"
# without text normalization
# + id="_dWnZZH-usU5" colab_type="code" colab={}
print (sorted_cnn_count[:10])
# + [markdown] id="6qWiLiGd9dMF" colab_type="text"
# [(8905, 'في'), (6401, 'من'), (5167, 'على'), (2318, 'عن'), (2276, 'بعد'), (1937, 'إلى'), (1476, 'داعش'), (1470, 'لـCNN:'), (1433, 'مع'), (1172, 'هل')]
# + [markdown] id="DTbhckaNuts8" colab_type="text"
# with text normalization
# + id="ozlujhUEuwOR" colab_type="code" colab={}
print (list(norm_cnn_count.items())[:10])
# + [markdown] id="AfqqXpcS9euP" colab_type="text"
# [('في', 13652), ('من', 12166), ('على', 5321), ('عن', 3507), ('بعد', 2536), ('إلى', 1954), ('داعش', 3144), ('لـCNN:', 1472), ('مع', 4165), ('هل', 1942)]
# + [markdown] id="_LXddXl2ux6m" colab_type="text"
# #### RT
# + [markdown] id="kZjFLJJRu1z4" colab_type="text"
# without text normalization
# + id="2czGJEugu2N4" colab_type="code" colab={}
print (sorted_rt_count[:10])
# + [markdown] id="K9KbzEKQ9gYe" colab_type="text"
# [(119313, 'في'), (49560, 'من'), (48777, 'على'), (20839, 'عن'), (20554, 'إلى'), (16939, 'روسيا'), (16372, 'مع'), (9323, 'بعد'), (9322, 'الروسية'), (8982, 'سوريا')]
# + [markdown] id="V2oUExt0u3uB" colab_type="text"
# with text normalization
# + id="j-T2gUThu5ug" colab_type="code" colab={}
print (list(norm_rt_count.items())[:10])
# + [markdown] id="NGHzf3Sl9hmI" colab_type="text"
# [('في', 170496), ('من', 89792), ('على', 49696), ('عن', 27359), ('إلى', 20620), ('روسيا', 19939), ('مع', 34430), ('بعد', 10746), ('الروسية', 11311), ('سوريا', 10872)]
# + [markdown] id="Jy1rGuC9u7Kw" colab_type="text"
# #### BBC
# + [markdown] id="PlCoG9xtu8mA" colab_type="text"
# without text normalization
# + id="jSHVTeGIu-NS" colab_type="code" colab={}
print (sorted_bbc_count[:10])
# + [markdown] id="3ZdoP3gE9jUp" colab_type="text"
# [(43957, 'في'), (14610, 'على'), (14051, 'من'), (5865, 'عن'), (4184, 'مقتل'), (4111, 'إلى'), (3648, 'مع'), (3423, 'بعد'), (2635, 'بين'), (2236, 'سوريا')]
# + [markdown] id="8Dt5HHKAu_ot" colab_type="text"
# with text normalization
# + id="gigpFSo8vA0x" colab_type="code" colab={}
print (list(norm_bbc_count.items())[:10])
# + [markdown] id="ZxSiFkU79khN" colab_type="text"
# [('في', 54271), ('على', 14877), ('من', 26774), ('عن', 8263), ('مقتل', 4586), ('إلى', 4127), ('مع', 10640), ('بعد', 3831), ('بين', 3919), ('سوريا', 3386)]
| 1 - Exploring Corpora/code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import re
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('../Data/datos_previos_clean.csv',index_col=0)
data.head()
data.columns
data['Provincia'].unique()
data['Zona'].unique()
print(data['Provincia'].value_counts(sort=True)[0:8].reset_index()['index'].values)
print()
#print(data['Provincia'].value_counts(sort=True)[0:8])
print(data['Provincia'].value_counts(sort=True))
a = pd.DataFrame(data['Provincia'].value_counts(sort=True)[0:8]).reset_index()
datos_por_prov = a.copy()
datos_por_prov['Prov'] = datos_por_prov['index']
datos_por_prov['Datos'] = datos_por_prov['Provincia']
datos_por_prov.drop(columns=['index','Provincia'],inplace=True)
datos_por_prov.rename(columns={'Prov':'Provincia'},inplace=True)
datos_por_prov.sort_values(by='Datos',ascending=True,inplace=True)
datos_por_prov = datos_por_prov.reset_index().drop(columns=['index'])
datos_por_prov
plt.figure(figsize=(10,8))
plt.barh(y=datos_por_prov['Provincia'],width=datos_por_prov['Datos'])
for i,v in enumerate(datos_por_prov['Provincia']):
plt.text(datos_por_prov['Datos'][i],i,datos_por_prov['Datos'][i],fontsize=13)
plt.xlim([0,13300])
plt.title('Cantidad de Datos por Provincias',fontsize=16)
plt.savefig('../Images/Cant. de Datos por Provincias.png')
datos_por_prov.plot(kind='barh',y='Datos',x='Provincia',figsize=(10,8))
data['Zona'].value_counts(sort=True)
data['Zona'].value_counts(sort=True).reset_index()['index'].values
capital_df=data.loc[data['Provincia']=='Capital Federal'].copy().reset_index().drop(columns='index')
gbanorte_df=data.loc[data['Provincia']=='Bs.As. G.B.A. Zona Norte'].copy().reset_index().drop(columns='index')
costa_df=data.loc[data['Provincia']=='Buenos Aires Costa Atlántica'].copy().reset_index().drop(columns='index')
gbasur_df=data.loc[data['Provincia']=='Bs.As. G.B.A. Zona Sur'].copy().reset_index().drop(columns='index')
gbaoeste_df=data.loc[data['Provincia']=='Bs.As. G.B.A. Zona Oeste'].copy().reset_index().drop(columns='index')
cordoba_df=data.loc[data['Provincia']=='Córdoba'].copy().reset_index().drop(columns='index')
santafe_df=data.loc[data['Provincia']=='Santa Fe'].copy().reset_index().drop(columns='index')
bsasint_df=data.loc[data['Provincia']=='Buenos Aires Interior'].copy().reset_index().drop(columns='index')
capital_df.to_csv('../Data/data_capital.csv')
#gbanorte_df.to_csv('../Data/data_gbanorte.csv')
#gbaoeste_df.to_csv('../Data/data_gbaoeste.csv')
#gbasur_df.to_csv('../Data/data_gbasur.csv')
#costa_df.to_csv('../Data/data_costa.csv')
#cordoba_df.to_csv('../Data/data_cordoba.csv')
#santafe_df.to_csv('../Data/data_santafe.csv')
#bsasint_df.to_csv('../Data/data_bsasint.csv')
# +
gba_df = data.loc[data['Zona']=='GBA'].copy().reset_index().drop(columns='index')
bsas_df = data.loc[data['Zona']=='BsAs'].copy().reset_index().drop(columns='index')
resto_df = data.loc[data['Zona']=='Resto País'].copy().reset_index().drop(columns='index')
gba_df.to_csv('../Data/data_todo_gba.csv')
#bsas_df.to_csv('../Data/data_resto_bsas.csv')
#resto_df.to_csv('../Data/data_resto_pais.csv')
# -
combo_provs_df = data.loc[(data['Provincia']=='Buenos Aires Costa Atlántica')|(data['Provincia']=='Córdoba')|(data['Provincia']=='Santa Fe')|(data['Provincia']=='Buenos Aires Interior')].copy().reset_index().drop(columns='index')
combo_provs_df.to_csv('../Data/combo_provincias.csv')
| TP2-Predictor-Regresion-Lineal-Properati/Notebooks/Division datos por ciudades.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp ecoenv
# -
# ## Google Colab preparations
# +
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
IN_MAIN = __name__ == '__main__'
# +
#Infrastructure for copying notebooks
if IN_COLAB and IN_MAIN:
home_dir = '/content/drive/MyDrive/Colab Notebooks/Ecosystems/v3'
if IN_COLAB and IN_MAIN:
from google.colab import drive
drive.mount('/content/drive')
import sys
sys.path.append(home_dir)
# %cd $home_dir
# !pip -q install import-ipynb
# -
#export
import gym
import numpy as np
import matplotlib.pyplot as plt
from gym import spaces
# import import_ipynb
from ecotwins.utility import distance, motion_diagram
from ecotwins.animal_classes import Ecosystem, Terrain, Animal, SimpleSheep
# +
# Can we (re)move this cell?
ORIGIN = np.array([0.,0.])
SIDE = 20
N_OBJECTS = 10
# Agent settings
RADIUS = SIDE/2
DELTA = 0.01 #1e-2
REWARD_RADIUS = SIDE/10
# TRACE_LENGTH = 1000
# Training settings
EPISODE_LENGTH = 2000
# EPISODE_LENGTH = np.int(np.round(SIDE/DELTA))
# +
# Can we (re)move this cell?
def generate_objects(side=None, n_objects=N_OBJECTS):
"""Generates a random map of objects
param side: size of each side of the grid
param n_objects: (initial) number of objects in the ecosystem
"""
objects=(np.random.rand(n_objects,2)-0.5)*side # Takes values in (-ecoenv.SIDE/2,+ecoenv.SIDE/2)
return objects
# -
# # The EcoEnv class
# +
#export
class EcoEnv(gym.Env):
"""An ecosystem environment for OpenAI gym"""
metadata = {'render.modes': ['human']} #Ta ej bort
#A functon that takes an ecosystem and returns a gym
def __init__(self, ecosystem):
super(EcoEnv, self).__init__()
self.ecosystem = ecosystem
self.agent = ecosystem.agent
self.perception = ecosystem.agent.perception
self.position = self.agent.position.copy()
self.p_happiness = self.agent.happiness(self.ecosystem.terrain)
self.current_step = 0
self.action_space = self.agent.action_space
self.observation_space = self.agent.observation_space
# debug
self.total_reward = 0
# An attempt to reward stayin alive
self.age_reward = (self.agent.happiness(self.ecosystem.terrain) /
self.agent.hyperparameters['max_age']
)
def _next_observation(self):
return self.agent.observation(self.ecosystem.terrain)
# Helper function to step.
def _take_action(self, action):
# Maybe this should be handled by the ecosystem ie the call should be
# self.ecosystem.update(action, agent)
# print(type(self))
self.agent.update(action, self.ecosystem.terrain)
# terrain update as well
self.position = self.agent.position.copy()
return self.position
# Reward functions
def _reward(self):
happiness = self.agent.happiness(self.ecosystem.terrain)
r = happiness - self.p_happiness
self.p_happiness = happiness
# TODO: Clean handling of resource punishment
return r + self.age_reward - 3 * self.agent.out_of_resources()
def _is_done(self):
"""Done if maximum number of step exceed or if
we are outside the region. """ # or if we die?
return self.current_step > EPISODE_LENGTH or self._is_outside()
def _is_outside(self): return np.abs(self.position).max() > self.side / 2
def _is_close(self):
return distance(self.position, self.objects).min() < self.reward_radius
# Execute one time step within the environment
def step(self, action):
self._take_action(action)
self.current_step += 1
reward = self._reward()
self.total_reward += reward
obs = self._next_observation()
done = self.ecosystem.is_done()
return obs, reward, done, {}
# Reset the state of the environment to an initial state
def reset(self):
print(f"Reset@{self.current_step}, accumulated reward: {self.total_reward:.2f}", end="")
print(", Interoception levels: ", end="")
print(*[f'{k}:{v:.2f}' for k,v in self.agent.interoception.items()], sep=', ', end="")
print(f' happiness: {self.agent.happiness(self.ecosystem.terrain):.2f}')
self.total_reward = 0
self.ecosystem.reset()
self.position = self.agent.position.copy()
self.current_step = 0
self.p_happiness = self.agent.happiness(self.ecosystem.terrain)
# self.consumed = []
return self._next_observation()
# Render the environment to the screen
def render(self, trace, mode='human', close=False):
# @TODO This should be handled properly
side = self.ecosystem.terrain.space[0,1] - self.ecosystem.terrain.space[0,0]
# @TODO Should support any number of object types not just one.
objects = next(iter(self.ecosystem.terrain.objects.values()))
motion_diagram(objects, trace, side)
# -
if IN_MAIN:
# t = Terrain(objects={'dandelion':(np.random.random((10,2)) - 0.5) * 20})
t = Terrain(objects={'dandelion': 10})
hyperparameters = {'max_age': 2000, 'delta': 0.1, 'close': 1, 'gamma': 0.9}
agent = SimpleSheep(distances={'dandelion':10})
eco = Ecosystem(t, agent)
env = EcoEnv(eco)
from nbdev.export import notebook2script; notebook2script()
| ecoenv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Import NumPy and SciPy (not needed when using --pylab)
# %pylab inline
import scipy as sp
# Load data from file
zz = np.loadtxt('wiggleZ_DR1_z.dat',dtype='float'); # Load WiggleZ redshifts
# Check bounds
np.min() # Check bounds
np.max()
# **Construct histogram from data**
# There are several histrogram commands: hist() will be fine here, but note the syntax below. Also note that the bin *edges* are returned, so that there will nbins+1 of these.
nbins = 50; # Is this a good choice?
n, bins, patches = hist() # With hist, one needs to (spuriously) request the patch objects as well
x = bins[0:nbins] + (bins[2]-bins[1])/2; # Convert bin edges to centres, chopping the last
# Interpolate histogram output -> p(z); n.b. that you can also use numerical quadrature to get $P(z)$ directly.
# Import the function you need
from scipy.interpolate import interp1d
# +
# Build an interpolation function for p(z) that accepts an arbitrary redshift z
# -
z = linspace(0,2,100); plot(z,p(z)) # Test your interpolation function out
# Use numerical integration to get $P(z) = \int_0^\infty p(z') dz'$
# Import the function you need
from scipy import integrate
Pz = lambda : ... # Use integrate inside a lambda function to define P(z)?
total = Pz(5) # Get normalisation constant by evaluating P(z->\infty)
total # Check that this worked
| Day_02/00_Scipy/scipy_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Meat consumption worldwide
# ## Analysis questions
# Analyze from wikipedia, life expectancy and meat consumption across the world.
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
from sklearn import metrics
# # Data
# ## Life expectancy by country (2019):
lifedata= pd.read_html('https://en.wikipedia.org/wiki/List_of_countries_by_life_expectancy')
lifedata= lifedata[0]
lifedata.head()
lifedata= lifedata.drop(index= [0]) #first row dropped
birth= lifedata[['Country', 'Life expectancy at birth']] #life expectancy at birth
birth.head()
# We only need country and life expectancy at birth of all genders:
birth_all= birth.loc[:, [('Country', 'Country'), ('Life expectancy at birth', 'All')]]
birth_all
birth_all= pd.concat([birth_all['Country'], birth_all['Life expectancy at birth']], axis=1 )
birth_all
birth_all= birth_all.set_index('Country')
birth_all
# ## Meat consumption by country (2017):
meatconsumptiondata= pd.read_html('https://en.wikipedia.org/wiki/List_of_countries_by_meat_consumption')
meatconsumptiondata= meatconsumptiondata[0]
meatconsumptiondata
meatdata2017= meatconsumptiondata.loc[:, ['Country', 'kg/person (2017) [11]']]
meatdata2017
meatdata2017= meatdata2017.rename(columns= {'kg/person (2017) [11]': 'kg/person'})
meatdata2017= meatdata2017.set_index('Country')
meatdata2017
# Now we merge data based on the country:
concatenated_data= pd.merge(birth_all.reset_index(), meatdata2017.reset_index() ).set_index('Country')
concatenated_data.head()
concatenated_data= concatenated_data.rename(columns={'All': 'Life expectancy (Age)', 'kg/person': 'Meat consumption (kg/person)'}) #renaming columns
concatenated_data.head()
# Now we drop rows that have any null values:
concatenated_data= concatenated_data.dropna()
concatenated_data.head()
# there is some stuff in meat consumption column
concatenated_data.iloc[:, 1]= concatenated_data.iloc[:, 1].replace('32[15]', '32').astype('float')
concatenated_data.head() # now it is ok
# # Data analysis
# ## Top 20 countries in terms of life expectancy
birth_all.head()
birth_all.info()
# +
N=10
top= birth_all.sample(frac=N/190) #shuffling countries and then get N samples
top
# -
# Let's plot scatter plot and compare relative sizes
# +
randomgene= np.random.RandomState(300)
topvalues= (np.concatenate(top.values))**2
topindex= top.index
x= randomgene.randn(N)
y =randomgene.randn(N)
colors= np.random.randn(1)**2
plt.figure(figsize=(10,10))
for i, j, index, values in zip(x, y, topindex, topvalues ):
plt.scatter(i, j, s=values , alpha=0.5 )
plt.annotate(index, (i, j), c='k')
# -
# ## Exploring relation between meat comsumption and life expectancy
# Constructing models:
#
# 2nd, 10th order polynomial fits
# +
#fitting observations
polynomialfit= np.polyfit(concatenated_data['Meat consumption (kg/person)'], concatenated_data['Life expectancy (Age)'], 2) #10th polynomial fit
polynomialfit2= np.polyfit(concatenated_data['Meat consumption (kg/person)'], concatenated_data['Life expectancy (Age)'], 10) #2nd order polynomial fit
#models
model= np.poly1d(polynomialfit) #2nd order model
model10 = np.poly1d(polynomialfit2) #10th order model
#fits
xfit=np.linspace(concatenated_data.iloc[:, 1].min(), concatenated_data.iloc[:, 1].max(), 10000)
yfit= model(xfit) #2n order
yfit10= model10(xfit) #10th order
# +
concatenated_data.plot.scatter(x= 'Meat consumption (kg/person)',y= 'Life expectancy (Age)', figsize=(10,10)) #scatter plot of original data points
plt.plot(xfit, yfit,c= 'r', linestyle='-', linewidth=3, label='2n order order polynomial fit') # 2nd order fit
plt.plot(xfit, yfit10, c='g', linestyle=':', linewidth= 8, label='10th order polynomial fit') #10th order fit
plt.legend()
plt.grid()
# -
# Calculating coefficient of determination ($R^2$)
# +
r2_2nd= metrics.r2_score(concatenated_data['Life expectancy (Age)'], model(concatenated_data['Meat consumption (kg/person)']) )
r2_10th= metrics.r2_score(concatenated_data['Life expectancy (Age)'], model10(concatenated_data['Meat consumption (kg/person)']) )
print(f'(2n order, 10th order) : ({r2_2nd}, {r2_10th}) ' )
# -
# Constructing linear fit
# +
from scipy import stats
slope, intercept, r, p, std_err= stats.linregress(concatenated_data.iloc[:, 1], concatenated_data.iloc[:, 0])
def linearmodel(x, slope, intercept):
return slope*x+intercept
# -
# Plot linear fit
# +
concatenated_data.plot.scatter(x= 'Meat consumption (kg/person)',y= 'Life expectancy (Age)', figsize=(10,10), label='Datapoints') #scatter plot of original data points
plt.plot(xfit, linearmodel(xfit, slope, intercept), label='Linear fit', c='r')
plt.legend()
plt.grid()
# -
# # Testing statistical significance
stats.spearmanr(concatenated_data)
stats.pearsonr(x= concatenated_data['Meat consumption (kg/person)'], y= concatenated_data['Life expectancy (Age)'] )
# It seems like when eating more meat, it is better, since it **increases your life expectancy.** However, there is possiblity that wealthiness have influenced this result. Country, which eats less meat, tends to be poorer, hence they are more undernourished. Wealthiness (hence undernourishment) may have affected the result.
# # Another research: How about comparing countries with same wealthiness
# Let's compare life expectancy against meat consumption among countries with same relative purchasing power index. In this way, we can lessen the effect of wealthiness affecting the result.
# # Data
# **Data about life expectancy and meat consumption:**
concatenated_data
# **Data of purchasing power index:**
df_ppp= pd.read_html('https://www.numbeo.com/quality-of-life/rankings_by_country.jsp?title=2021&displayColumn=1')
df_ppp= df_ppp[1]
df_ppp.head()
# Cleaning data of ppp:
# remove Rank row
df_ppp= df_ppp.drop(columns=['Rank'])
df_ppp
# Merging purchasing power index data with meat consumption and life expectancy data:
pppmeatlifdat= pd.merge(concatenated_data.reset_index(), df_ppp).set_index('Country')
pppmeatlifdat.head()
# # 2. Data analysis
# Let's check if there is relationship between purchasing power index and meat consumption:
# +
pppmeatlifdat.plot.scatter(x= 'Purchasing Power Index', y= 'Meat consumption (kg/person)', figsize=(10,10))
plt.grid()
# -
# Seems like there is a linear relationship when ppp is small, but the effect of ppp is lost when ppp is over 40.
# Let's check if we are right. Applying Pearson correlation test:
# +
sorted_pppmeatlifdat= pppmeatlifdat.sort_values(by= ['Purchasing Power Index']) # sorting values
sorted_pppmeatlifdat
# + tags=[]
countriessmall= sorted_pppmeatlifdat[sorted_pppmeatlifdat.iloc[:, -1]<45] #extracting countries with ppp smalle than 45
countriesbig= sorted_pppmeatlifdat[sorted_pppmeatlifdat.iloc[:, -1]>45] #extracting countries with ppp bigger than 45
# -
stats.pearsonr(x= countriessmall['Purchasing Power Index'], y=countriessmall['Meat consumption (kg/person)'] )
stats.pearsonr(x= countriesbig['Purchasing Power Index'], y=countriesbig['Meat consumption (kg/person)'] )
# After calculating Pearson's tests, we notice that there is a weak but very significant **positive correlation** between PPP and meat consumption for PPP values that is smaller than 45. But the correlation for bigger values of PPP had no significance.
# # Re-exploring the relationship between meat consumption and life expectancy between countries that are equal in wealthiness.
# Since wealthiest countries had no correlation between PPP and meat consumption, we can conclude that wealthiness does not contribute to the life expectancy.
#
# Let's explore the relationship in wealthiest countries.
countriesbig.head()
countriesbig.plot.scatter(x= 'Meat consumption (kg/person)', y='Life expectancy (Age)' )
stats.pearsonr(x= countriesbig['Meat consumption (kg/person)'], y=countriesbig['Life expectancy (Age)'])
# # Conclusion
# There is no relationship between meat consumption and life expectancy.
| worldwidemeat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# ## The CUTE Classification Scheme
#
# Each _CUTE_ problem is assigned a string identifire in it's SIF encoding, which has the form:
#
# `**XXXr-XX-n-m**`
#
# The **X** characters do not need to be present in the origonal FORTRAN tools that queries SIF CLASS.DB file, see
# https://www.cuter.rl.ac.uk/Problems/classification.shtml for more information.
#
# ## CUTEst.jl
#
# It appears the CUTEst.jl package contains problems classified under the scope of Test, which is encoded SIF classification in the first **X** to the right of the first hyphen. Such test set problems are listed here https://www.cuter.rl.ac.uk/Problems/mastsif.html. Furthermore, every problem in the test set has a first charachter of '2' left of first hypen; suggesting that we test our algorithms on problems that have an analytical computation for the Hessian. (Note, CUTEst.jl belongs to the JuliaSmoothOptimizers orginization)
#
# In CUTEst.jl (v0.12) they have a tool called `CUTEst.select(...)` that scans the set of Test Problems and queiries a subset corresponding to the given arguments.
# For more information `ctrl+F` _Selection tool_ here http://juliasmoothoptimizers.github.io/CUTEst.jl/v0.12/tutorial/#Selection-tool
#
#
#
# ## FORTRAN Tool
# There does exist a tool in the SIFDecode artifact directory that is created when adding CUTEst.jl, called `slct.f`. The command line tool `slct.f` should work when your enviroment variables are exported into your shells path, as explained in https://github.com/ralna/CUTEst. When mine are not exported to my _~/.zshrc_, a segmentation fault occurs. You can find the `slct.f` tool in the path relative to your Julia installation directory, i.e. .julia/artifacts/{long shasum hash}/libexec/SIFDecode-2.0.3/src/select
#
# +
using CUTEst, NLPModels
# selecting unconstrained problems:
problems = CUTEst.select(contype="unc")
length(problems)
# -
# ## JuliaSmoothOptimizers (JSO)
#
# The orginization behind CUTEst.jl, NLPModels.jl, ADNLPModels.jl (an abstract framework for AD in NLP models developed with ForwardDiff.jl in mind) is JuliaSmoothOptimizers.
#
# #### Code disscusion
# The **newton_cg** function below is a JSO complient solver that constructs a trust-region sub-problem using Krlov.jl. Krylov.jl performs a conjugate gradient method to solve the subproblem.
#
# **Refrence:**
# https://juliasmoothoptimizers.github.io/pages/tutorials/creating-a-jso-compliant-solver/
# +
using Krylov, LinearAlgebra
function newton_cg(nlp :: AbstractNLPModel)
x = nlp.meta.x0
fx = obj(nlp, x)
gx = grad(nlp, x)
ngx = norm(gx)
while norm(gx) > 1e-6
Hx = hess_op(nlp, x)
d, _ = cg(Hx, -gx)
slope = dot(gx, d)
if slope >= 0 # Not a descent direction
d = -gx
slope = -dot(d,d)
end
t = 1.0
xt = x + t * d
ft = obj(nlp, xt)
while ft > fx + 0.5 * t * slope
t *= 0.5
xt = x + t * d
ft = obj(nlp, xt)
end
x = xt
fx = ft
gx = grad(nlp, x)
ngx = norm(gx)
end
return x, fx, ngx
end
# test it on 2D-Rosenbrock function
nlp = CUTEstModel("ROSENBR")
print(newton_cg(nlp))
finalize(nlp) # you must always finalize the model
# -
# ## LinearOperators.jl
#
# This package is the cornerstone of efficient design of nonlinear optimization algorithms through the JuliaSmoothOptimizers package.
# We perform an exploration of the package below, which is compatible with Julia 1.3 and up.
#
# `LinearOperator():` defines a linear transformation
# - v -> Av.
# - v -> A'v
# - v -> A*v
#
# There are many advantages of using LinearOperators instead of working with matrices
#
# **Reference:**
# https://juliasmoothoptimizers.github.io/LinearOperators.jl/stable/
# +
using LinearOperators
prod(v) = [v[1] + v[2]; 2v[1] + 3v[3]]
tprod(v) = [v[1] + 2v[2]; v[1] + 3v[2]]
A = LinearOperator(Float64, 2, 2, false, false, prod, tprod, tprod)
# -
A = rand(500, 500)
B = rand(500, 500)
@time A*B;
opA = LinearOperator(A)
opB = LinearOperator(B)
@time opA*opB;
# +
v = rand(500)
@time (A * B) * v
@time A * (B*v)
@time (opA * opB) * v
@time opA * (opB * v);
# -
# Note a linear operator is nearly a wrapper of a matrix, but there are some differences (e.g. slicing)
A * ones(500) == opA * ones(500)
| CUTEstExploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Rappi
# https://www.rappi.com.ar/restaurantes/rapanui
#
# Gráficar el histograma de precios de rapanui
# +
import requests
def rapanui_products_from_rappi():
headers = {
'authority': 'services.rappi.com.ar',
'accept': 'application/json, text/plain, */*',
'dnt': '1',
'authorization': 'Bearer <KEY>',
'accept-language': 'es-AR',
'user-agent': 'Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/86.0.4240.183 Safari/537.36',
'sentry-trace': '3a8cf9c44c784450a1a5deb254324537-94c2d6b8cb1e5b36-1',
'content-type': 'application/json',
'origin': 'https://www.rappi.com.ar',
'sec-fetch-site': 'same-site',
'sec-fetch-mode': 'cors',
'sec-fetch-dest': 'empty',
'referer': 'https://www.rappi.com.ar/',
}
data = '{"store_type":"restaurant","lat":-34.5984904,"lng":-58.427746}'
response = requests.post('https://services.rappi.com.ar/api/ms/web-proxy/restaurants-bus/store/rapanui', headers=headers, data=data)
return response.json()
def rapanui_prices():
prices = []
rapanui_data = rapanui_products_from_rappi()
for corridor in rapanui_data["corridors"]:
for product in corridor["products"]:
prices.append(product["price"])
return prices
# -
import pandas as pd
rapanui_df = pd.DataFrame({"Precios de Rapanui (ARS)":rapanui_prices()})
rapanui_df.hist()
# +
# YAPA
# Version mejorada que no duplica productos en caso de estar repetidos
def rapanui_prices():
products = {}
rapanui_data = rapanui_products_from_rappi()
for corridor in rapanui_data["corridors"]:
for product in corridor["products"]:
products[product["name"]] = product["price"]
prices = list(products.values())
return prices
import pandas as pd
rapanui_df = pd.DataFrame({"Precios de Rapanui (ARS)":rapanui_prices()})
rapanui_df.hist()
| Scraping/2_HTTP_Avanzado/ejercicio/rappi-rapanui_solucion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
df = pd.read_csv('data/Minjust2018.csv',
header=None,
usecols=[4, 27, 28, 33])
df.fillna('', inplace=True)
df.rename(columns={4: 'name',
27: 'director',
28: 'activity',
33: 'founders'},
inplace = True)
# -
df
def count_letters(words):
return pd.Series([len(word) for word in words])
df.loc[count_letters(df['director']) == count_letters(df['name'])]
df.loc[count_letters(df['director']) <= 4]
| df_loc_select_with_callable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jinseongdu/ERC223-token-standard/blob/master/The_tensorflow_version_magic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nPDc8yQtVxk4"
# #TensorFlow versions in Colab
# + [markdown] id="N2y2uqx9GfA5"
#
# ##Background
# Colab has two versions of TensorFlow pre-installed: a 2.x version and a 1.x version. Colab uses TensorFlow 2.x by default, though you can switch to 1.x by the method shown below.
#
# + [markdown] id="aR_btJrKGdw7"
# ##Specifying the TensorFlow version
#
# Running `import tensorflow` will import the default version (currently 2.x). You can use 1.x by running a cell with the `tensorflow_version` magic **before** you run `import tensorflow`.
# + colab={"base_uri": "https://localhost:8080/"} id="NeWVBhf1VxlH" outputId="6251a6d7-9ce4-4058-a56f-64e847dc3a66"
# %tensorflow_version 1.x
# + [markdown] id="8dSlimhOVxlQ"
# Once you have specified a version via this magic, you can run `import tensorflow` as normal and verify which version was imported as follows:
# + colab={"base_uri": "https://localhost:8080/"} id="-XbfkU7BeziQ" outputId="3de79323-bec5-4b43-eaa0-e963050e91cc"
import tensorflow
print(tensorflow.__version__)
# + id="DR4L_vyOFtPd"
# + [markdown] id="nv75qFFRFtse"
# okwwo
#
# + [markdown] id="uBIKyjpEVxlU"
# If you want to switch TensorFlow versions after import, you **will need to restart your runtime** with 'Runtime' -> 'Restart runtime...' and then specify the version before you import it again.
# + [markdown] id="8UvRkm1JGUrk"
# ## Avoid Using ``pip install`` with GPUs and TPUs
#
# We recommend against using ``pip install`` to specify a particular TensorFlow version for both GPU and TPU backends. Colab builds TensorFlow from source to ensure compatibility with our fleet of accelerators. Versions of TensorFlow fetched from PyPI by ``pip`` may suffer from performance problems or may not work at all.
| The_tensorflow_version_magic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Normal model for <NAME>'s experiment (BDA3 p.66)
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# with open('../data/light.txt', 'r') as f:
# data = f.readlines()
# y = np.asarray(y, dtype=int)
y = np.loadtxt('../data/light.txt')
plt.hist(y, bins=30)
plt.title('Histogram of Newcomb\'s measurements');
# +
# sufficient statistics
n = len(y)
y_mean = np.mean(y)
y_var = np.var(y, ddof=1) # ddof=1 -> sample estimate
# grid for computing density of mu
mu_grid = np.linspace(np.min(y[y>0]), np.max(y), 100)
# compute the exact marginal posterior density for mu
# multiplication by 1./sqrt(y_var/n) is due to the transformation of variable
pm_mu = stats.t.pdf((mu_grid - y_mean) / np.sqrt(y_var/n), n-1) / np.sqrt(y_var/n)
mu025, mu975 = y_mean + stats.t.ppf(0.025, n-1), y_mean + stats.t.ppf(0.975, n-1)
# plot the posterior of mu
plt.plot(mu_grid, pm_mu)
plt.axvline(mu025, color='red')
plt.axvline(mu975, color='red')
axes = plt.gca()
plt.text(
mu025,
axes.get_ylim()[1]+0.03,
'2.5%',
horizontalalignment='right'
)
plt.text(
mu975,
axes.get_ylim()[1]+0.03,
'97.5%',
horizontalalignment='left'
)
plt.xlabel(r'$\mu$')
plt.title(r'Maginal posterior distribution for $\mu$');
# +
# calculate posterior interval by simulation
n_sample = 1000
# draw sigma squares
sigma2_sample = (n-1)* y_var / stats.chi2.rvs(df=n-1, size=n_sample)
mu_sample = stats.norm.rvs(y_mean, sigma2_sample/n, size=(1, n_sample))
# posterior median and 95% posterior interval
mu_sample_median = np.median(mu_sample)
mu_sample_025, mu_sample_975 = np.percentile(mu_sample, [2.5, 97.5])
print('mu sample median: {0:.2f}\n95% posterior interval:[{1:.2f}, {2:.2f}]'.format(mu_sample_median, mu_sample_025, mu_sample_975))
| content/zh/projects/bda3/Estimate_the_speed_of_light.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # Lists of Other Types
//
// Watch the full [C# 101 video](https://www.youtube.com/watch?v=oIQdb93xewE&list=PLdo4fOcmZ0oVxKLQCHpiUWun7vlJJvUiN&index=14) for this module.
//
// You've been practicing lists of strings, but you can make a list of anything! Here's a number example.
//
// ## Fibonacci
//
// Fibonacci is a cool number sequence. It adds up the last two numbers up to make the next number. You start with 1 and 1
// 1 + 1 = 2 (1, 1, 2)
// 1 + 2 = 3 (1, 1, 2, 3)
// 2 + 3 = 5 (1, 1, 2, 3, 5)
// 3 + 5 = 8 (1, 1, 2, 3, 5, 8)
// and so on. There are lots of stuff in nature that follow this number sequence, and has lots of cool stuff if you want to look it up!
//
// > Start with the base numbers: Here's a list with just 1, 1 in it. Run it and see what happens.
// + dotnet_interactive={"language": "csharp"}
var fibonacciNumbers = new List<int> {1, 1};
foreach (var item in fibonacciNumbers)
Console.WriteLine(item);
// -
// Now, you don't want just 1,1 in it! You want more of the sequence. In this code, you're using the last two numbers of the list, adding them together to make the next number, then adding it to the list.
//
// > Run the code to try it out.
// + dotnet_interactive={"language": "csharp"}
var fibonacciNumbers = new List<int> {1, 1}; // Starting the list off with the basics
var previous = fibonacciNumbers[fibonacciNumbers.Count - 1]; // Take the last number in the list
var previous2 = fibonacciNumbers[fibonacciNumbers.Count - 2]; // Take the second to last number in the list
fibonacciNumbers.Add(previous + previous2); // Add the previous numbers together, and attach the sum to the end of the list
foreach (var item in fibonacciNumbers) // Print out the list
Console.WriteLine(item);
// -
// ## Count -1
//
// Why do you need to do `fibonacciNumbers.Count -1` to get the last number of the list? Well, `Count` tells you how many items are in a list. However, the index of an item starts at zero. So, if you only had one item in your list, the count would be one, but the index of the item would be 0. The index and count of the last item is always one off.
// # Challenge: Fibonacci to 20th number
//
// We've given you a base of code that deals with Fibonacci. Can you make a list that has the first 20 fibonacci numbers?
//
// > Make and print a list that has the first 20 fibonacci numbers.
// + dotnet_interactive={"language": "csharp"}
Console.WriteLine("Challenge");
// -
// ## Tips and tricks
//
// - The final number should be 6765.
// - Could you make a `for` loop? A `foreach` loop? A `while` loop? Which kind of loop do you prefer and which would be more useful?
// - Are you getting close, but are you one number off? That's a really common issue! Remember that `>` and `>=` are similar, but they end up being one off from the other. Try playing around with that?
// - Remember that you're starting with two items in the list already.
// - Stuck? Watch the [C# 101 video](https://www.youtube.com/watch?v=oIQdb93xewE&list=PLdo4fOcmZ0oVxKLQCHpiUWun7vlJJvUiN&index=14) for this module. Try out pausing once you get an idea and trying it out first before watching the rest.
// # Continue learning
//
// There are plenty more resources out there to learn!
//
// > [⏩ Next Module - Objects and Classes](http://tinyurl.com/csharp101-notebook13-ipynb)
// >
// > [⏪ Last Module - Search, Sort, and Index Lists](http://tinyurl.com/csharp101-notebook11-ipynb)
// >
// > [Watch the video](https://www.youtube.com/watch?v=oIQdb93xewE&list=PLdo4fOcmZ0oVxKLQCHpiUWun7vlJJvUiN&index=14)
// >
// > [Documentation: Arrays, Lists, and Collections](https://docs.microsoft.com/dotnet/csharp/tour-of-csharp/tutorials/arrays-and-collections?WT.mc_id=Educationalcsharp-c9-scottha)
// >
// > [Start at the beginning: What is C#?](https://www.youtube.com/watch?v=BM4CHBmAPh4&list=PLdo4fOcmZ0oVxKLQCHpiUWun7vlJJvUiN&index=1)
// # Other resources
//
// Here's some more places to explore:
// > [Other 101 Videos](https://dotnet.microsoft.com/learn/videos?WT.mc_id=csharpnotebook-35129-website)
// >
// > [Microsoft Learn](https://docs.microsoft.com/learn/dotnet/?WT.mc_id=csharpnotebook-35129-website)
// >
// > [C# Documentation](https://docs.microsoft.com/dotnet/csharp/?WT.mc_id=csharpnotebook-35129-website)
// >
// > [Download Visual Studio](https://visualstudio.microsoft.com/downloads/)
// >
| csharp-101/12-Lists of Other Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import numpy as np
import pandas as pd
from datetime import datetime
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn-whitegrid')
from sklearn.svm import SVC
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
# -
# # Load the data
df = pd.read_csv('00 df.csv')
train = df[df['flag']=='train']
test = df[df['flag']=='test']
# +
cat_feats = ['age_bin','capital_gl_bin','education_bin','hours_per_week_bin','msr_bin','occupation_bin','race_sex_bin']
y_train = train['y']
x_train = train[['age_bin','capital_gl_bin','education_bin','hours_per_week_bin','msr_bin','occupation_bin','race_sex_bin']]
x_train = pd.get_dummies(x_train,columns=cat_feats,drop_first=True)
y_test = test['y']
x_test = test[['age_bin','capital_gl_bin','education_bin','hours_per_week_bin','msr_bin','occupation_bin','race_sex_bin']]
x_test = pd.get_dummies(x_test,columns=cat_feats,drop_first=True)
# -
# # Support Vector Machine
svm = SVC(kernel="rbf", C=0.025,random_state=101)
svm.fit(x_train, y_train)
y_pred=svm.predict(x_test)
# +
test_calc = pd.concat([pd.DataFrame(y_test).reset_index(drop=True),pd.DataFrame(y_pred).reset_index(drop=True)],axis=1)
test_calc.rename(columns={0: 'predicted'}, inplace=True)
test_calc['predicted'] = test_calc['predicted'].apply(lambda x: 1 if x > 0.5 else 0)
df_table = confusion_matrix(test_calc['y'],test_calc['predicted'])
print (df_table)
print('accuracy:', (df_table[0,0] + df_table[1,1]) / (df_table[0,0] + df_table[0,1] + df_table[1,0] + df_table[1,1]))
print ('precision:', df_table[1,1] / (df_table[1,1] + df_table[0,1]))
print('recall:', df_table[1,1] / (df_table[1,1] + df_table[1,0]))
p = df_table[1,1] / (df_table[1,1] + df_table[0,1])
r = df_table[1,1] / (df_table[1,1] + df_table[1,0])
print('f1 score: ', (2*p*r)/(p+r))
# -
svm = SVC(kernel="linear", C=0.025,random_state=101)
svm.fit(x_train, y_train)
y_pred=svm.predict(x_test)
# +
test_calc = pd.concat([pd.DataFrame(y_test).reset_index(drop=True),pd.DataFrame(y_pred).reset_index(drop=True)],axis=1)
test_calc.rename(columns={0: 'predicted'}, inplace=True)
test_calc['predicted'] = test_calc['predicted'].apply(lambda x: 1 if x > 0.5 else 0)
df_table = confusion_matrix(test_calc['y'],test_calc['predicted'])
print (df_table)
print('accuracy:', (df_table[0,0] + df_table[1,1]) / (df_table[0,0] + df_table[0,1] + df_table[1,0] + df_table[1,1]))
print ('precision:', df_table[1,1] / (df_table[1,1] + df_table[0,1]))
print('recall:', df_table[1,1] / (df_table[1,1] + df_table[1,0]))
p = df_table[1,1] / (df_table[1,1] + df_table[0,1])
r = df_table[1,1] / (df_table[1,1] + df_table[1,0])
print('f1 score: ', (2*p*r)/(p+r))
# -
svm = SVC(kernel="poly", C=0.025,random_state=101)
svm.fit(x_train, y_train)
y_pred=svm.predict(x_test)
# +
test_calc = pd.concat([pd.DataFrame(y_test).reset_index(drop=True),pd.DataFrame(y_pred).reset_index(drop=True)],axis=1)
test_calc.rename(columns={0: 'predicted'}, inplace=True)
test_calc['predicted'] = test_calc['predicted'].apply(lambda x: 1 if x > 0.5 else 0)
df_table = confusion_matrix(test_calc['y'],test_calc['predicted'])
print (df_table)
print('accuracy:', (df_table[0,0] + df_table[1,1]) / (df_table[0,0] + df_table[0,1] + df_table[1,0] + df_table[1,1]))
print ('precision:', df_table[1,1] / (df_table[1,1] + df_table[0,1]))
print('recall:', df_table[1,1] / (df_table[1,1] + df_table[1,0]))
p = df_table[1,1] / (df_table[1,1] + df_table[0,1])
r = df_table[1,1] / (df_table[1,1] + df_table[1,0])
print('f1 score: ', (2*p*r)/(p+r))
# -
| 09SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib.ticker as mticker
import matplotlib
from mpl_finance import candlestick_ohlc
from datetime import datetime
import seaborn as sns
sns.set()
df = pd.read_csv('../dataset/GOOG2020.csv')
df.head()
date = [datetime.strptime(d, '%Y-%m-%d') for d in df['Date']]
candlesticks = list(zip(mdates.date2num(date),df['Open'],
df['High'],df['Low'],df['Close'],df['Volume']))
# +
fig = plt.figure(figsize = (15, 15))
ax = fig.add_subplot(1,1,1)
ax.set_ylabel('Quote ($)', size=20)
dates = [x[0] for x in candlesticks]
dates = np.asarray(dates)
volume = [x[5] for x in candlesticks]
volume = np.asarray(volume)
candlestick_ohlc(ax, candlesticks, width=1,
colorup='g', colordown='r')
pad = 0.25
yl = ax.get_ylim()
ax.set_ylim(yl[0]-(yl[1]-yl[0])*pad,yl[1])
ax2 = ax.twinx()
ax2.set_position(matplotlib.transforms.Bbox([[0.125,0],[0.9,0.32]]))
pos = df['Open'] - df['Close']<0
neg = df['Open'] - df['Close']>0
ax2.bar(dates[pos],volume[pos],color='green',width=1,align='center')
ax2.bar(dates[neg],volume[neg],color='red',width=1,align='center')
ax2.set_xlim(min(dates),max(dates))
yticks = ax2.get_yticks()
ax2.set_yticks(yticks[::3])
ax2.yaxis.set_label_position("right")
ax2.set_ylabel('Volume', size=20)
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax.xaxis.set_major_locator(mticker.MaxNLocator(10))
plt.show()
# +
def removal(signal, repeat):
copy_signal = np.copy(signal)
for j in range(repeat):
for i in range(3, len(signal)):
copy_signal[i - 1] = (copy_signal[i - 2] + copy_signal[i]) / 2
return copy_signal
def get(original_signal, removed_signal):
buffer = []
for i in range(len(removed_signal)):
buffer.append(original_signal[i] - removed_signal[i])
return np.array(buffer)
signal = np.copy(df.Open.values)
removed_signal = removal(signal, 30)
noise_open = get(signal, removed_signal)
signal = np.copy(df.High.values)
removed_signal = removal(signal, 30)
noise_high = get(signal, removed_signal)
signal = np.copy(df.Low.values)
removed_signal = removal(signal, 30)
noise_low = get(signal, removed_signal)
signal = np.copy(df.Close.values)
removed_signal = removal(signal, 30)
noise_close = get(signal, removed_signal)
# +
noise_candlesticks = list(zip(mdates.date2num(date),noise_open,
noise_high,noise_low,noise_close))
fig = plt.figure(figsize = (15, 5))
ax = fig.add_subplot(1,1,1)
ax.set_ylabel('Quote ($)', size=20)
candlestick_ohlc(ax, noise_candlesticks, width=1,
colorup='g', colordown='r')
ax.plot(dates, [np.percentile(noise_close, 95)] * len(noise_candlesticks), color = (1.0, 0.792156862745098, 0.8, 0.7),
linewidth=10.0, label = 'overbought line')
ax.plot(dates, [np.percentile(noise_close, 10)] * len(noise_candlesticks),
color = (0.6627450980392157, 1.0, 0.6392156862745098, 0.7),
linewidth=10.0, label = 'oversold line')
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax.xaxis.set_major_locator(mticker.MaxNLocator(10))
plt.legend()
plt.show()
# +
fig = plt.figure(figsize = (15, 12))
ax1 = plt.subplot2grid((3, 1), (0, 0), rowspan=2)
ax1.set_ylabel('Quote ($)', size=20)
dates = [x[0] for x in candlesticks]
dates = np.asarray(dates)
volume = [x[5] for x in candlesticks]
volume = np.asarray(volume)
candlestick_ohlc(ax1, candlesticks, width=1,
colorup='g', colordown='r')
pad = 0.25
yl = ax1.get_ylim()
ax1.set_ylim(yl[0]-(yl[1]-yl[0])*pad,yl[1])
ax2 = ax1.twinx()
pos = df['Open'] - df['Close']<0
neg = df['Open'] - df['Close']>0
ax2.bar(dates[pos],volume[pos],color='green',width=1,align='center')
ax2.bar(dates[neg],volume[neg],color='red',width=1,align='center')
ax2.set_xlim(min(dates),max(dates))
yticks = ax2.get_yticks()
ax2.set_yticks(yticks[::3])
ax2.yaxis.set_label_position("right")
ax2.set_ylabel('Volume', size=20)
ax1.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax1.xaxis.set_major_locator(mticker.MaxNLocator(10))
ax2 = plt.subplot2grid((3, 1), (2, 0))
ax2.set_ylabel('Quote ($)', size=20)
candlestick_ohlc(ax2, noise_candlesticks, width=1,
colorup='g', colordown='r')
ax2.plot(dates, [np.percentile(noise_close, 95)] * len(noise_candlesticks), color = (1.0, 0.792156862745098, 0.8, 1.0),
linewidth=5.0, label = 'overbought line')
ax2.plot(dates, [np.percentile(noise_close, 10)] * len(noise_candlesticks),
color = (0.6627450980392157, 1.0, 0.6392156862745098, 1.0),
linewidth=5.0, label = 'oversold line')
ax2.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax2.xaxis.set_major_locator(mticker.MaxNLocator(10))
plt.legend()
plt.show()
# -
| misc/overbought-oversold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Create a list for each level of a binary tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Is this a binary search tree?
# * Yes
# * Should each level be a list of nodes?
# * Yes
# * Can we assume we already have a Node class with an insert method?
# * Yes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * 5, 3, 8, 2, 4, 1, 7, 6, 9, 10, 11 -> [[5], [3, 8], [2, 4, 7, 9], [1, 6, 10], [11]]
#
# Note: Each number in the result is actually a node containing the number
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/graphs_trees/tree_level_lists/tree_level_lists_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
# %run ../bst/bst.py
# %load ../bst/bst.py
class BstLevelLists(Bst):
def create_level_lists(self):
levelLists = [[self.root]]
stax = levelLists[-1][:]
while stax:
stax = [c for n in stax for c in [n.left, n.right] if c]
levelLists.append(stax)
return levelLists
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# %run ../utils/results.py
# +
# # %load test_tree_level_lists.py
import unittest
class TestTreeLevelLists(unittest.TestCase):
def test_tree_level_lists(self):
bst = BstLevelLists(Node(5))
bst.insert(3)
bst.insert(8)
bst.insert(2)
bst.insert(4)
bst.insert(1)
bst.insert(7)
bst.insert(6)
bst.insert(9)
bst.insert(10)
bst.insert(11)
levels = bst.create_level_lists()
results_list = []
for level in levels:
results = Results()
for node in level:
results.add_result(node)
results_list.append(results)
self.assertEqual(str(results_list[0]), '[5]')
self.assertEqual(str(results_list[1]), '[3, 8]')
self.assertEqual(str(results_list[2]), '[2, 4, 7, 9]')
self.assertEqual(str(results_list[3]), '[1, 6, 10]')
self.assertEqual(str(results_list[4]), '[11]')
print('Success: test_tree_level_lists')
def main():
test = TestTreeLevelLists()
test.test_tree_level_lists()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/graphs_trees/tree_level_lists/tree_level_lists_solution.ipynb) for a discussion on algorithms and code solutions.
| graphs_trees/tree_level_lists/tree_level_lists_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prep
# +
import sqlite3
import matplotlib
import pandas
import scipy.stats
# %matplotlib inline
# -
# ### Data
db = sqlite3.connect('immigration_analysis.sqlite.db')
source = pandas.read_sql(
'''
SELECT
occupation,
majorSocCode + 0.0 AS majorSocCode,
may2020UnemployedCnt + 0.0 AS may2020UnemployedCnt,
may2020UnemployedRate + 0.0 AS may2020UnemployedRate,
may2019UnemployedCnt + 0.0 AS may2019UnemployedCnt,
may2019UnemployedRate + 0.0 AS may2019UnemployedRate,
totalPositionsCnt + 0.0 AS totalPositionsCnt
FROM
unemployment_with_immigration
''',
db
)
source['occupationShort'] = source['occupation'].apply(lambda x: x.replace(' occupations', ''))
# ### Utility Functions
FONT_FAMILY = 'Lato'
def style_graph(ax, title, x_label, y_label, x_range=None, y_range=None):
"""Style a matplotlib graph.
Args:
ax: The matplob axes to manipulate.
title: The string title to have appear at the top of the graphic.
x_label: The label for the hoizontal axis.
y_label: The label for the vertical axis.
x_range: Two element tuple or list with the minimum and maximum values for the horizontal axis.
y_range: Two element tuple or list with the minimum and maximum values for the vertical axis.
"""
ax.spines['top'].set_color('#ffffff')
ax.spines['right'].set_color('#ffffff')
ax.spines['bottom'].set_color('#ffffff')
ax.spines['left'].set_color('#ffffff')
ax.set_xlabel(x_label, fontname=FONT_FAMILY, fontweight='medium', fontsize=13)
ax.set_ylabel(y_label, fontname=FONT_FAMILY, fontweight='medium', fontsize=13)
ax.xaxis.label.set_color('#555555')
ax.yaxis.label.set_color('#555555')
ax.tick_params(axis='x', colors='#555555')
ax.tick_params(axis='y', colors='#555555')
if x_range:
ax.set_xlim(x_range)
if y_range:
ax.set_ylim(y_range)
if title:
ax.set_title(title, fontname=FONT_FAMILY, fontweight='medium', fontsize=16, color="#505050")
ax.title.set_position([.5, 1.05])
for tick in ax.get_xticklabels():
tick.set_fontname(FONT_FAMILY)
tick.set_fontweight('medium')
for tick in ax.get_yticklabels():
tick.set_fontweight('medium')
# <br>
#
# # EDA / Hypothesis Testing
# ### Hypothesis 1: The proportion of visa holders is small part of the working force
source['may2020EmployedRate'] = (100 - source['may2020UnemployedRate'])
source['may2020EmployedCnt'] = source['may2020UnemployedCnt'] / source['may2020UnemployedRate'] * source['may2020EmployedRate']
source['percentVisa'] = source['totalPositionsCnt'] / source['may2020EmployedCnt'] * 100
# +
ax = source.sort_values('percentVisa').plot.barh(
x='occupationShort',
y='percentVisa',
figsize=(9, 9),
legend=None,
colors=['#8da0cb'] * 22
)
style_graph(ax, 'Percent of Employed Workforce is on Visa', 'Percent', 'Occupation')
for p in ax.patches:
label_val = '%.1f%%' % p.get_width()
end_x = p.get_x() + p.get_width() + 0.02
ax.annotate(label_val, (end_x, p.get_y() + 0.05), color='#8da0cb')
# -
percent_under_1_percent = source[source['percentVisa'] < 1].shape[0] / source.shape[0] * 100
print('Percent occupations under 1%% visa: %.2f%%' % percent_under_1_percent)
# ### Hypothesis 2: Occupations with higher unemployment have < 1% H1B
source['changeUnemployedRate'] = source['may2020UnemployedRate'] - source['may2019UnemployedRate']
# +
ax = source.sort_values('may2020UnemployedRate').plot.barh(
x='occupationShort',
y=['changeUnemployedRate', 'percentVisa'],
figsize=(9, 9),
colors=['#8da0cb', '#fc8d62'] * 22
)
style_graph(
ax,
'Occupations with More Unemployment Have Fewer Visa Workers',
'Percent',
'Occupation'
)
# -
high_visa = source[source['changeUnemployedRate'] >= 5]
low_visa = source[source['changeUnemployedRate'] < 5]
p_value = scipy.stats.mannwhitneyu(high_visa['changeUnemployedRate'], low_visa['changeUnemployedRate'])[1]
if p_value < 0.05:
print('Hypothesis accepted (%.2f).' % p_value)
print(
'High unemployment had %.2f%% while low unemployment had %.2f%%.' % (
high_visa['percentVisa'].mean(),
low_visa['percentVisa'].mean()
)
)
# ### Hypothesis 3: If all visa jobs went to unemployed, unemployment would not improve substantially
source['hypotheticalUnemploymentCnt'] = source.apply(
lambda row: max([row['may2020UnemployedCnt'] - row['totalPositionsCnt'], 0]),
axis=1
)
source['hypotheticalUnemployment'] = source.apply(
lambda row: row['may2020UnemployedRate'] * (row['hypotheticalUnemploymentCnt'] / row['may2020UnemployedCnt']),
axis=1
)
source['hypotheticalChangeInUnemployment'] = source['may2020UnemployedRate'] - source['hypotheticalUnemployment']
# +
ax = source.sort_values('may2020UnemployedRate').plot.barh(
x='occupationShort',
y=['hypotheticalUnemployment', 'may2020UnemployedRate'],
figsize=(9,9),
colors=['#8da0cb', '#fc8d62'] * 22
)
style_graph(
ax,
'Unemployment Rate Does Not Reduce Substantially in Most Occupations',
'Percent',
'Occupation'
)
# -
avg_change_in_unemployment = source['hypotheticalChangeInUnemployment'].mean()
print('Avg change in unemployment: %.2f%%' % avg_change_in_unemployment)
source[source['hypotheticalChangeInUnemployment'] > 1].shape[0] / source.shape[0]
new_unemployment_rate = source['hypotheticalUnemploymentCnt'].sum() / source['may2020UnemployedCnt'].sum() * 13.3
print('New unemployment rate: %.2f%%' % new_unemployment_rate)
# # Overall tabs
source['totalPositionsCnt'].sum() / source['may2020EmployedCnt'].sum()
visa_class_counts = pandas.read_sql(
'''
SELECT
visaClass,
sum(totalWorkerPositionsCnt) AS cnt
FROM
immigration_data
WHERE
visaActiveDuringMay2020 = "1"
GROUP BY
visaClass
''',
db
)
visa_class_counts.to_csv('visa_counts.csv')
visa_class_counts
pandas.read_sql(
'''
SELECT
sum(totalWorkerPositionsCnt) AS cnt
FROM
immigration_data
WHERE
visaActiveDuringMay2020 = "1"
''',
db
)
pandas.read_sql(
'''
SELECT
src.visaClassSimplified AS visaClassSimplified,
sum(src.cnt) AS cnt
FROM
(
SELECT
(
CASE
WHEN visaClass = "H-1B" THEN "H-1B or Similar"
WHEN visaClass = "H-1B1 Chile" THEN "H-1B or Similar"
WHEN visaClass = "H-1B1 Singapore" THEN "H-1B or Similar"
WHEN visaClass = "E-3 Australian" THEN "H-1B or Similar"
ELSE visaClass
END
) AS visaClassSimplified,
totalWorkerPositionsCnt AS cnt
FROM
immigration_data
WHERE
visaActiveDuringMay2020 = "1"
) src
GROUP BY
src.visaClassSimplified
''',
db
)
# # Efficiency
# +
ax = source.sort_values('may2020UnemployedRate').plot.scatter(
x='changeUnemployedRate',
y='percentVisa',
figsize=(7,5),
color='#8da0cb',
alpha=0.8,
s=20
)
style_graph(
ax,
'Increase in Unemployment Rate vs Percent of Workers on Visa',
'Increase in Unemployment Rate (05/2020 - 05/2019)',
'Percent Workers on Visa'
)
# -
total_visa_positions = source['totalPositionsCnt'].sum()
total_new_jobs = (source['may2020UnemployedCnt'] - source['hypotheticalUnemploymentCnt']).sum()
print('Percent of jobs lost: %.2f%%' % ((1 - total_new_jobs / total_visa_positions) * 100))
# # Output
source.to_csv('unemployment_and_counts_extended.csv')
source.head(5)
| Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit ('.venv')
# metadata:
# interpreter:
# hash: aa53c8c6e6798222a2084c11cc25017700a8d3ad495b587e3a634f357767115f
# name: python3
# ---
# # Usage: quickest tour
#
# [](https://colab.research.google.com/github/lisphilar/covid19-sir/blob/master/example/usage_quickest.ipynb)
#
# Thank you for using CovsirPhy!!
# This is the quickest tour to get an overview of CovsirPhy.
#
# - Download datasets
# - Parameter estimation with phase-dependent SIR-derived models
# - Simulate the number of cases
# ## Preparation
# Prepare the packages.
# + tags=[]
# # !pip install covsirphy --upgrade
from pprint import pprint
import covsirphy as cs
cs.__version__
# -
# ## Dataset preparation
# Download the datasets to "../input" directory and load them.
# If "../input" directory has the datasets, `DataLoader` will load the local files. If the datasets were updated in remote servers, `DataLoader` will update the local files automatically.
# Please refer to [Usage: datasets](https://lisphilar.github.io/covid19-sir/usage_dataset.html) for the details.
# + tags=[]
# Standard users and developers
data_loader = cs.DataLoader("../input")
# The number of cases and population values
jhu_data = data_loader.jhu()
# -
# We can select the following countries.
# + tags=[]
pprint(jhu_data.countries(), compact=True)
# -
# ## Start scenario analysis
# As an example, we will analysis the number of cases in Italy using `Scenario` class. To initialize this class, we need to specify the country name.
snl = cs.Scenario(country="Italy", province=None)
snl.register(jhu_data)
# ## Check records
# Let's see the records at first. `Scenario.records()` method return the records as a pandas dataframe and show a line plot. Some kind of complement will be done for analysis, if necessary.
df = snl.records()
df.tail()
# ## S-R trend analysis
# S-R trend analysis finds the change points of SIR-derived ODE parameters. This is a significant step of analysis because we assume that ODE parameter values will be changed phase by phase (not daily basis, not constant through the outbreak).
# Details will be explained in [Usage: phases](https://lisphilar.github.io/covid19-sir/usage_phases.html).
_ = snl.trend()
# Summarize the phases.
#
# - Type: "Past" or "Future"
# - Start: start date of the phases
# - End: end date of the phases
# - Population: total population in the phases
snl.summary()
# ## Hyperparameter estimation of ODE models
# Here, we will estimate the parameter values of SIR-derived models. As an example, we use SIR-F model. Details of models will be explained in [Usage: SIR-derived models](https://lisphilar.github.io/covid19-sir/usage_theoretical.html).
# + tags=[]
# Default value of timeout is 180 sec
snl.estimate(cs.SIRF, timeout=30)
# -
# ## History of reproduction number
# Let's see the history of parameter values. Reproduction number is here.
_ = snl.history(target="Rt")
# ## History of parameters
# History of each parameter. Values will be divided by the values in 0th phase.
_ = snl.history_rate()
# ## Simulate the number of cases
# How many cases will be in 30 days if the parameter values will not be changed from today?
# Add a phase with 30 days from the date of the last record
snl.add(days=30)
_ = snl.simulate()
# Next, please see [Usage: scenario analysis](https://lisphilar.github.io/covid19-sir/usage_quick.html) to find details of datasets and how to perform scenario analysis.
# Thank you!
| example/usage_quickest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Information Extraction Applications
# - Tagging news an other contents
# - Chatbots
# - Applications in social media
# - Extracting data from forms and receipts
# # Information Extraction Tasks
# - Keyphrase extraction (KPE)
# - Named entity recognition
# - Named entity disambiguation and linking
# - Relation extraction
# # The General Pipeline for IE
# 
| 05_Information Extraction/01_Information Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as mp
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# https://www.kaggle.com/khotijahs1/predict-who-will-move-to-a-new-job
data=pd.read_csv('aug_train.csv')
data.head()
# 'target'= dependent variable
# getting job or not
data['target'].value_counts().plot(kind='bar',title='job or not')
data.columns
# data imputation (missing values)
data.isnull().sum().plot(kind='barh',figsize=(13,5),title='missing values')
data['gender'].value_counts()
data.plot(y=['experience','education_level'], x="gender", kind='bar')
| HR Analysis/HR anlysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Caesar Cipher
#
# **By <NAME>, Byte Sized Code**
# ## What is Caesar Cipher?
#
# The Caesar Cipher is one of the simplest [ciphers](https://en.wikipedia.org/wiki/Cipher) in cryptography.
# It is used to encrypt or decrypt messages/texts between two parties.
# It is a kind of **Monoalphabetic Substitution cipher.**
#
# A [Substitution cipher](https://en.wikipedia.org/wiki/Substitution_cipher) is a simple cipher where some characters of the plaintext are replaced by specific characters to produce the ciphertext.
#
# To encrypt text using classic Caesar cipher, we just shift the alphaber by 3 and substitute the values. For example, A is replaced by D, B -> E, C -> F, and so on, till Y -> B and Z -> C.
# Its really simple. Let's have a look at a simple example.
# ## Working Example
#
# Lets take a simple example.
# Our enemy is stationed in a palace in a valley. Unsuspecting of any attach they are enjoying in their palace. Our army is at one of the hill peaks near the valley and our allies are at the other end on the opposite hill.
# We want to encrypt the phrase **ATTACK AT ONCE** and send it to our allies on the other side, so that we both can start the attack at the same time and crush our enemy from both sides.
# We disregard spaces as we don't have any method to encrypt spaces using the Caesar Cipher, thus, the plaintext becomes **ATTACKATONCE**
#
# To encrypt using the traditional Caesar Cipher, we shift alphabets to the right by 3 places. Thus, we need to make the following changes:
# - A -> D
# - T -> W
# - C -> F
# - K -> N
# - O -> R
# - N -> Q
# - E -> H
# Making the necessary changes:
# plaintext - **ATTACKATONCE**
# encrypted - **DWWDFNDWRQFH**
#
# Once the message is delivered on the other side, our allies (who know how to encrypt and decrypt) can then just reverse shift the alphabet by 3 places to get the message that we sent.
# The message they receive: **DWWDFNDWRQFH**
# They make the following changes:
# - D -> A
# - W -> T
# - F -> C
# - N -> K
# - R -> O
# - Q -> N
# - H -> E
# Making the changes:
# encrypted message - **DWWDFNDWRQFH**
# decrypted message - **ATTACKATONCE**
#
# The allies then add the necessary spaces, look at the message and launch their attack! Together, we defeat the enemies and have a nice day hanging out at their palace.
#
# Wikipedia has another [working example](https://en.wikipedia.org/wiki/Caesar_cipher#Example) if you want to check it out.
# ## What are we going to do?
#
# We will code two simple functions that will help us encrypt and decrypt text using the Caesar Cipher. As the project is aimed at beginners, we will be using conventions and code that is easy to understand but not high performance.
# Once you are familiar and comfortable with the code, I would suggest you to try to improve on it.
# ## Actual Code
# ### Data structure for conversion
#
# To make things simpler, we will use dictionaries to use for the encryption and decryption process.
# We will have one dictionary for encryption and another for decryption.
# Encryption dictionary
encrypt_key = {"A": "D", "B": "E", "C": "F", "D": "G", "E": "H", "F": "I", "G": "J",
"H": "K", "I": "L", "J": "M", "K": "N", "L": "O", "M": "P", "N": "Q",
"O": "R", "P": "S", "Q": "T", "R": "U", "S": "V", "T": "W", "U": "X",
"V": "Y", "W": "Z", "X": "A", "Y": "B", "Z": "C"}
print(encrypt_key)
# Decryption dictionary
# Just reverse the encryption dictionary
decrypt_key = {value: key for key, value in encrypt_key.items()}
print(decrypt_key)
# ### Encryption
#
# We will write one function to help with the encryption process.
# The function will take in the plaintext (string) as input and return the encrypted text (string) as output.
# Encryption function
def encrypt(plaintext):
# Initialize an empty string
encrypted = ""
# We use .upper() to convert all characters to uppercase as our dictionary supports uppercase only
for character in plaintext.upper():
# Add the encrypted character to the initialized string
encrypted += encrypt_key[character]
# Return the encrypted string
return encrypted
# We can use the function to encrypt the initial example phrase, "ATTACKATONCE" and check if it matches the answer that we got above.
encrypt("ATTACKATONCE")
# Yes it does! Hooray! We've done half of the job, the other part is quite as simple too.
# We just need a decryption function.
# ### Decryption
#
# The decryption function is the inverse of the encryption function.
# It takes in an encrypted string and converts it to the corresponding plaintext (string).
# The function will be quite similar to the encryption function.
# Decryption function
def decrypt(encrypted):
# Initialize the plaintext string
plaintext = ""
# Same logic for using the upper function
for character in encrypted.upper():
plaintext += decrypt_key[character]
# Return the final plaintext
return plaintext
# Now that we have a function for decryption, lets test it out using the encrypted text we got above:
decrypt('DWWDFNDWRQFH')
# Yay! We got our initial value for "ATTACKATONCE" from the encrypted text!
# So, that's all for the script.
# You can surely compile it into a single pytho file.
# I'd suggest improving on the code a bit once you fully understand it (look below for other interesting thing to try out).
# If this was your first project after learning Python, then congratulations on completing it!
# Watchout for more projects to come and keep coding meanwhile!
# ### What to do next?
#
# First of all, compile it into a single script and try to make it a commandline tool.
# Where you can ask the user for inputs and then return the corresponding encrypted and decrypted texts.
#
# Here are a few ideas of improving on the project (and making some similar projects):
# - Rework the current data structure for conversions (dictionary) and implement a modulo based conversion
# - Expand the project to include any number of character shifts from 1 to 26 (we just worked with a shift of 3)
# - If Cryptography interests you (like me), try coding some other simple ciphers using python. Here are two to get you started:
# - [ROT 13](https://en.wikipedia.org/wiki/ROT13)
# - [Vigenere Cipher](https://en.wikipedia.org/wiki/Vigen%C3%A8re_cipher)
| Projects/Caesar Cipher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false" id="cOPFNyxk7dPV"
# # 2022-03 DoD Training
#
# Outline
#
# 1. The Grants data model - quick walkthrough
# 2. Useful queries from a Funder GRID ID
# 3. Querying using arbitrary lists of IDs
#
# + [markdown] Collapsed="false" id="hMaQlB7DG8Vw"
# ## Prerequisites
#
# This notebook assumes you have installed the [Dimcli](https://pypi.org/project/dimcli/) library and are familiar with the [API LAB](https://api-lab.dimensions.ai/) *Getting Started* tutorials.
#
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 34959, "status": "ok", "timestamp": 1646231321904, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="MEsDfbBt7dPX" outputId="519ca9ea-e95e-46d3-d3b3-e1184732b2e4"
# !pip install dimcli --quiet
import dimcli
from dimcli.utils import *
import json
import sys
import pandas as pd
import plotly.express as px
from IPython.display import Image
print("==\nLogging in..")
ENDPOINT = "https://app.dimensions.ai"
if 'google.colab' in sys.modules:
import getpass
KEY = getpass.getpass(prompt='API Key: ')
dimcli.login(key=KEY, endpoint=ENDPOINT)
else:
KEY = ""
dimcli.login()
dsl = dimcli.Dsl()
# + [markdown] Collapsed="false" id="HCpneDnCB4-U" tags=[]
# ## PART 1. The grants data model - quick walkthrough
#
# + [markdown] Collapsed="false" id="-2viO5qLB4-U"
# Fields reference: https://docs.dimensions.ai/dsl/datasource-grants.html
# + [markdown] Collapsed="false" id="9g2OGneGB4-U"
# ### 1.1 Dimensions grant `ID` VS 'original' `grant_number`
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 232} executionInfo={"elapsed": 404, "status": "ok", "timestamp": 1646231322303, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="ujGhYY6wB4-U" outputId="c864f455-48a1-4896-c494-8bb9916e92e7"
# %%dsldf
search grants
for "detector AND chemicals"
return grants[id+grant_number+title] limit 5
# + [markdown] Collapsed="false" id="QCCwln2FB4-V"
# Once you have the IDs, you can search using them - especially when using related models e.g. publications, patents etc.. (more on this in the next section).
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 232} executionInfo={"elapsed": 431, "status": "ok", "timestamp": 1646231322730, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="CjI8w2QWB4-V" outputId="12da65ec-ed49-4e6e-b4fd-b34f237215f2"
# %%dsldf
search grants
where id in ["grant.9971026", "grant.9967366"]
or grant_number in ["ST/W000830/1", "2151709", "201547"]
return grants[id+grant_number+title]
# + [markdown] Collapsed="false" id="PlP_rgRLB4-W"
# ### 1.2 Field types: atomic data type (`funding_org_name`) VS lists (`funder_countries`) VS entities (eg `funders`)
# + colab={"base_uri": "https://localhost:8080/", "height": 397} executionInfo={"elapsed": 307, "status": "ok", "timestamp": 1646231323033, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="CoX8OyDVB4-W" outputId="c9e9f3fb-25cb-498a-80a5-f323f3bc5929"
# %%dsldf
search grants
for "detector AND chemicals"
return grants[id+funding_org_name+funder_countries+funders] limit 5
# + [markdown] Collapsed="false" id="2-MDr9udB4-W"
# Use `unnest(funders)` or `unnest(funder_countries)` to unpack the contents of complex fields (docs:https://docs.dimensions.ai/dsl/language.html#unnesting-multi-value-entity-fields)
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 232} executionInfo={"elapsed": 241, "status": "ok", "timestamp": 1646231323269, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="3S4sUP_EB4-W" outputId="66deb1a3-a1b8-4edd-c2cb-6c4afd30ba6b"
# %%dsldf
search grants
for "detector AND chemicals"
return grants[id+funding_org_name+unnest(funder_countries)] limit 5
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 412} executionInfo={"elapsed": 350, "status": "ok", "timestamp": 1646231323617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="jYCmUmsFB4-W" outputId="d18163a0-0084-4176-9438-168c4f48001e"
# %%dsldf
search grants
for "detector AND chemicals"
return grants[id+unnest(funders)] limit 5
# + [markdown] Collapsed="false" id="0gb5AJDoB4-X" tags=[]
# ### 1.3 Funder GRID identifiers (`funders.id`)
#
# + [markdown] id="hcxUJZqcB4-X"
# Let's find the GRID identifier we are interested in, using the [organizations API](https://docs.dimensions.ai/dsl/datasource-organizations.html)).
# + colab={"base_uri": "https://localhost:8080/", "height": 184} executionInfo={"elapsed": 484, "status": "ok", "timestamp": 1646231324099, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="yhq4nfts7dPY" outputId="05f0109d-50da-4192-aaea-ce80c4cf9bfb"
# %%dsldf
search organizations for "Department of Defense"
where country_name="United States" and types in ["Government"]
return organizations limit 100
# + [markdown] id="qYZpyW_sB4-X"
# Now let's find **grants** data using `grid.420391.d`
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 1238} executionInfo={"elapsed": 327, "status": "ok", "timestamp": 1646231324420, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="ffDzAENoB4-X" outputId="9f3d18c3-b627-4cbc-c49e-01fc475b167f"
# %%dsldf
search grants for "detector AND chemicals"
where funders.id = "grid.420391.d"
return grants limit 5
# + [markdown] id="tAjvi_urB4-X"
# ## PART 2. Useful queries using a Funder GRID identifier
# + colab={"base_uri": "https://localhost:8080/", "height": 664} executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1646231324420, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="PQ13xVN2B4-X" outputId="b25059c6-4769-44e2-c1b7-f49e5d59d4a9"
Image(url= "https://docs.dimensions.ai/dsl/_images/data-model-grants.png", width=1000)
# + [markdown] Collapsed="false" id="M-RgrBweB4-Y"
# ### 2.1 How many grants?
#
# The total number of results should match [what you see in Dimensions](https://app.dimensions.ai/discover/publication?search_mode=content&and_facet_funder=grid.420391.d).
#
# + colab={"base_uri": "https://localhost:8080/", "height": 555} executionInfo={"elapsed": 259, "status": "ok", "timestamp": 1646231324672, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="-aTqHF6nB4-Y" outputId="363f5c75-3efd-4286-9a68-d21e2045f222"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
return grants[id+title]
# + [markdown] Collapsed="false" id="oVDCAbaNB4-Y"
# ### 2.2 Downloading all grants records
#
# We can use a little Python and the `query_iterative` method.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 742} executionInfo={"elapsed": 6176, "status": "ok", "timestamp": 1646231330844, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="ekm4azh5B4-Y" outputId="ae4c4e26-27a4-4661-cde0-7ed7ad684aea"
# %%dslloopdf
search grants
where funders.id = "grid.420391.d"
return grants[id+title]
# + [markdown] id="QaEoKMoDB4-Y"
# Save the data so that we can reuse it later.
# + executionInfo={"elapsed": 4, "status": "ok", "timestamp": 1646231330844, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="sN2m_iliB4-Y"
all_grants = dsl_last_results
# + [markdown] id="VDZir8fiB4-Y"
# ### 2.3 Now that we have the query, we can experiment with other facets..
#
# #### Top organizations
#
# Change the final query bit to return `research_orgs`
# + colab={"base_uri": "https://localhost:8080/", "height": 1250} executionInfo={"elapsed": 713, "status": "ok", "timestamp": 1646231331554, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="OktoQsOPB4-Y" outputId="dd31505a-8afc-4b3e-8e0e-6d2c3fbc5ee0"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
and active_year >= 2010
return research_orgs
aggregate funding
limit 1000
# + colab={"base_uri": "https://localhost:8080/", "height": 917} executionInfo={"elapsed": 2117, "status": "ok", "timestamp": 1646231333665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="fLoQ3bAqB4-Z" outputId="2cced5a9-7504-4935-87b8-a466d2e12993"
px.scatter(dsl_last_results[:100], y="funding",
x="name",
color="state_name",
size="count",
height=900,
title="Funded organizations overview")
# + [markdown] id="U7Z_hDCZB4-Z"
# #### Top Countries
#
# Change the last query bit to return `research_org_countries`
# + colab={"base_uri": "https://localhost:8080/", "height": 254} executionInfo={"elapsed": 339, "status": "ok", "timestamp": 1646231334003, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="TVO4_SNwB4-Z" outputId="91740cde-1daa-464a-99f3-475b9a2b3a5c"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
and active_year >= 2010
return research_org_countries limit 1000
# + [markdown] id="lMkCbBtVB4-Z"
# #### Who is getting funded outside the US?
#
# Use the filter `research_org_countries.name != "United States"`
# + colab={"base_uri": "https://localhost:8080/", "height": 189} executionInfo={"elapsed": 511, "status": "ok", "timestamp": 1646231334510, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="Cf4HVI4jB4-Z" outputId="e5fa913e-ca35-473a-85fd-e0b53d3b4088"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
and active_year >= 2010
and research_org_countries.name != "United States"
return research_orgs limit 1000
# + [markdown] id="OTpzO3gVB4-Z"
# #### How do I see the specific grants information?
#
# Eg if I want to follow up on those non-US grants
# + colab={"base_uri": "https://localhost:8080/", "height": 1005} executionInfo={"elapsed": 315, "status": "ok", "timestamp": 1646231334821, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="ZC93gO2IB4-Z" outputId="aef06927-3d84-4b00-ed9b-47bec949b3e4"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
and active_year >= 2010
and research_org_countries.name != "United States"
return grants[id+dimensions_url+active_year+funding_org_name+funding_usd+grant_number+title+unnest(research_org_countries)+unnest(research_org_names)+research_orgs] limit 1000
# + [markdown] id="bsNk7awpB4-Z"
# #### Top Researchers
#
# + colab={"base_uri": "https://localhost:8080/", "height": 953} executionInfo={"elapsed": 634, "status": "ok", "timestamp": 1646231335450, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="pZEYZsvzB4-Z" outputId="949a190c-c575-46e2-9340-75c2c0c40d06"
# %%dsldf
search grants
where funders.id = "grid.420391.d"
and active_year >= 2010
return researchers[basics+current_research_org] limit 1000
# + [markdown] Collapsed="false" id="qLknsnV8B4-Z" tags=[]
# ### 2.4 Getting Publications from a funder ID
#
# + [markdown] Collapsed="false" id="8YQHc5pdB4-Z" tags=[]
# We can use the field `funders`, which is a direct link from Publications to Funder Organizations (a 'shortcut'), using the funders IDs.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 722} executionInfo={"elapsed": 308, "status": "ok", "timestamp": 1646231335753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="vUz4YnsaB4-Z" outputId="8a60e440-f15b-48d5-ee8c-71cb9f6ac9ba"
# %%dsldf
search publications
where funders.id = "grid.420391.d"
return publications[id+title+year+unnest(supporting_grant_ids)]
# + Collapsed="false" executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1646231335753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="sdiLRl7TB4-Z"
pubs = dsl_last_results
# + [markdown] Collapsed="false" id="9fYLmonlB4-Z"
# Now we can retrieve the associated grants.
#
#
#
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 1244} executionInfo={"elapsed": 226, "status": "ok", "timestamp": 1646231335976, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="srKfHKtyB4-a" outputId="e2fdeb51-b6de-4c5f-9147-e5c47399b8d2"
temp = pubs.dropna(subset=['supporting_grant_ids'])
grantids = temp['supporting_grant_ids'].to_list()
query = f"""
search grants
where id in {json.dumps(grantids)}
return grants[id+title+funders]
"""
linkedgrants = dsl.query(query).as_dataframe()
linkedgrants
# + [markdown] id="zK9wP3O-B4-a"
# ### 2.5 Getting Patents from a funder ID
# + colab={"base_uri": "https://localhost:8080/", "height": 672} executionInfo={"elapsed": 500, "status": "ok", "timestamp": 1646231336475, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="P4xQwNMZB4-a" outputId="ec475837-68ed-433c-8a9a-5de61faa640f"
# %%dsldf
search patents
where funders.id = "grid.420391.d"
return patents[id+title+year+unnest(associated_grant_ids)]
# + [markdown] Collapsed="false" id="2XskjV8WB4-a" tags=[]
# ## 3. Querying using arbitrary lists of IDs
#
# See also these tutorials
#
# * [Enriching Grants part 2: Adding Publications Information from Dimensions](https://api-lab.dimensions.ai/cookbooks/3-grants/2-grants-enrichment-adding-publications-information.html#)
# * [Working with longs lists of IDs](https://api-lab.dimensions.ai/cookbooks/1-getting-started/6-Working-with-lists.html#5.-How-Long-can-lists-get?)
#
# ### 3.1 From grants to other entities: the links available
# + Collapsed="false" colab={"base_uri": "https://localhost:8080/", "height": 649} executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1646231336477, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="qJMxo3SAB4-a" outputId="1b25da95-6c12-4880-cce0-f664b0352c91"
Image(url= "https://docs.dimensions.ai/dsl/_images/data-model-overview-1.png", width=1000)
# + [markdown] Collapsed="false" id="t7NxFPZeB4-a"
# ### 3.2 Example: Patents funders analysis
#
# Example - we want to get an overview of who's funded patents on a specific topic.
#
# **Two steps**
#
# * first, we create a patents dataset using any criteria of choice
# * second, we find associated grants for those patents/grants IDs
# * since we can have a large number of IDs, we need to pay attention to the API query 'size'
# + colab={"base_uri": "https://localhost:8080/", "height": 1105} executionInfo={"elapsed": 6342, "status": "ok", "timestamp": 1646231342808, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="SJ36boIFB4-a" outputId="bd619fd8-c778-4bcd-f860-f6909c9f5e3f"
# %%dslloopdf
search patents
for "detection AND explosives"
where year >= 2000
and associated_grant_ids is not empty
return patents[id+title+year+unnest(associated_grant_ids)]
# + executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1646231342809, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="FJv7jhUpB4-a"
patents = dsl_last_results
# + colab={"base_uri": "https://localhost:8080/", "height": 2382, "referenced_widgets": ["9a3db0e012ef406787e4b621a95ad6d2", "f73f97e8d2314099b1088a2dabe841cf", "c316616ec9b243bc97b137dc9e2bb206", "96d23661e633465a9718cb54569ff596", "d2722811c8bd4fe68a7c58a1c31a253e", "adf4e5f44fef443ba67e39f2f567c39a", "e6f5ee4e9b294e85b877f1ed952ed659", "6c0668b8a4ae48af833c1fd324cd4452", "d95450c7783c41e6830ce16d778116a0", "e9c70703143f464ca63addde46e5ca3d", "7ce3d52688de4113a1fdd73996c4c4f5"]} executionInfo={"elapsed": 47192, "status": "ok", "timestamp": 1646231389990, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="u8JNJGVLB4-a" outputId="40e995c0-bc03-4452-9e9c-99d38674fe80"
# we get grants for those patents, by segmenting the associated_grant_ids list into groups of 300 IDs
# this is because each DSL query can take max ~300 IDs at a time
from tqdm.notebook import tqdm as progressbar
associated_grant_ids = dsl_last_results['associated_grant_ids'].to_list()
#
# the main API query
#
q = """search grants
where id in {}
return grants[id+dimensions_url+researchers+title+active_year+funding_usd+funding_org_name+unnest(funder_countries)+unnest(research_org_countries)+unnest(research_org_names)]"""
#
# let's loop through all IDs in chunks and query Dimensions
#
results = []
for chunk in progressbar(list(chunks_of(list(associated_grant_ids), 300))):
data = dsl.query_iterative(q.format(json.dumps(chunk)), verbose=True)
results += data.grants
time.sleep(1)
#
# put the data into a dataframe, remove duplicates and save
#
grants = pd.DataFrame().from_dict(results)
print("Grants: ", len(grants))
grants.drop_duplicates(subset='id', inplace=True)
print("Unique Grants: ", len(grants))
#
# preview
#
print("Example:")
grants.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 817} executionInfo={"elapsed": 361, "status": "ok", "timestamp": 1646231390347, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="ollQG9wqB4-a" outputId="6cc5c5f1-a6e5-4dc5-964f-b0b31b0732e3"
# fix empty values
grants.fillna(0, inplace=True)
gsubset = grants[grants["funder_countries.name"] == 'United States']
px.scatter(gsubset,
x="funding_org_name",
y="funding_usd",
marginal_x="histogram",
color="funder_countries.name",
height=800,
title="US funding for selected patents dataset")
# + [markdown] id="jI3SvTo3B4-b"
# ### 3.3 Example: For a given list of DoD awards, how to find out if the awardees have awards from other funders?
#
# **Approach: Three steps**
#
# * first get all grants from DoD
# * second get all awards for those researchers (not just the DoD ones)
# * third, keep only the non-DoD awards
# + colab={"base_uri": "https://localhost:8080/", "height": 335} executionInfo={"elapsed": 7991, "status": "ok", "timestamp": 1646231463331, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="QGGaY-u47dPZ" outputId="a5d40442-ec13-4c83-f368-3311526c62be"
grants = dsl.query_iterative(f"""
search grants
where funders.id = "grid.420391.d"
return grants[id+title+grant_number+researchers]
""").as_dataframe()
grants.head(5)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 300, "status": "ok", "timestamp": 1646231463627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="iYR3cfJCB4-b" outputId="c514fbaf-10e8-4b77-e8b5-aa5acbcab181"
# ignore grants with no researchers info
grants.dropna(subset=['researchers'], inplace=True)
# get the IDS only
researchers = set()
for index, row in grants.iterrows():
res = [el['id'] for el in row['researchers']]
researchers.update(res)
print("Unique researchers: ", len(researchers))
# + colab={"base_uri": "https://localhost:8080/", "height": 2004, "referenced_widgets": ["34b25edbd0224bc1aa661546a63da31f", "c1dcb547065843dd8deed7f2c6be39d3", "5fc06415d43640bfacd4710cb947a39c", "85b87ce4cfb44ffea735821fd98eb0be", "ce96b05994a34f6aa5d94063d47395a1", "80236186b29740928bd89d083a7e40fe", "d7a34d372ca849d7b225d92e946a07c8", "5d54ba63419f4bae8911080a9ebe7791", "59490cde272c40fd9ca5fdda8323f6ee", "71c0e6c1153443fdb81073ea2eb56bcd", "f27eba9577cf49b28bfb642464fec989"]} executionInfo={"elapsed": 61609, "status": "ok", "timestamp": 1646231525232, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="6pBQ1yniB4-b" outputId="c101ec8c-44dd-4657-95f9-9fc7ef25e4b3"
# we get grants for all researchers, by segmenting the researchers list into groups of 300 IDs
# this is because each DSL query can take max ~300 researchers at a time
from tqdm.notebook import tqdm as progressbar
researcher_ids = researchers
#
# TRIAL RUN: Uncomment this line to use less researchers and speed things up
#
# researcher_ids= researcher_ids[:200]
#
# the main API query
#
q = """search grants
where researchers in {}
return grants[id+dimensions_url+researchers+title+active_year+funding_usd+funding_org_name+unnest(funder_countries)+unnest(research_org_countries)+unnest(research_org_names)]"""
#
# let's loop through all researcher IDs in chunks and query Dimensions
#
results = []
for chunk in progressbar(list(chunks_of(list(researcher_ids), 200))):
data = dsl.query_iterative(q.format(json.dumps(chunk)), verbose=True)
results += data.grants
time.sleep(1)
#
# put the data into a dataframe, remove duplicates and save
#
grants = pd.DataFrame().from_dict(results)
print("Grants: ", len(grants))
grants.drop_duplicates(subset='id', inplace=True)
print("Unique Grants: ", len(grants))
#
# preview
#
print("Example:")
grants.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} executionInfo={"elapsed": 225, "status": "ok", "timestamp": 1646231525453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="tac6EfICB4-b" outputId="9c3a7507-c2a4-4904-97e9-5356e7c3389b"
# fix empty values
grants.fillna(0, inplace=True)
# add aggregated counts
grants['country_count'] = grants.groupby('funder_countries.name')['id'].transform('count')
grants['country_funding'] = grants.groupby('funder_countries.name')['funding_usd'].transform('sum')
gsubset = grants[['funder_countries.name', 'country_count']]
px.choropleth(gsubset.drop_duplicates(),
locations="funder_countries.name",
locationmode="country names",
color="country_count",
hover_name="funder_countries.name",
color_continuous_scale=px.colors.sequential.Plasma,
title="Researchers' funding: number of grants by funder countries")
# + colab={"base_uri": "https://localhost:8080/", "height": 817} executionInfo={"elapsed": 210, "status": "ok", "timestamp": 1646231525659, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="DKMnF1eNB4-b" outputId="e19813d0-3c40-4107-cd98-9f2a93ed527e"
gsubset = grants[grants["funder_countries.name"] != 'United States']
px.scatter(gsubset,
y="funding_org_name",
x="research_org_names",
marginal_x="histogram",
color="funder_countries.name",
height=800,
title="Researchers' funding: funders from outside the US")
# + [markdown] id="nDDg0Fz1B4-b"
# #### To find out who are the researchers
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1646231525659, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYfmLTPbeMuYDDrETLbTVXTXnfVr9f7eBtkmR73A=s64", "userId": "10309320684375994511"}, "user_tz": 0} id="sepUb2DrB4-b" outputId="b9f458cb-516d-4ac8-cd77-51125d737b4d"
# eg for Japan
COUNTRY = "Japan"
focus_set = grants[grants["funder_countries.name"] == COUNTRY]['researchers'].to_list()
for res in focus_set[0]:
if res['id'] in researcher_ids:
print(res['first_name'], res['last_name'], "\n", dimensions_url(res['id'], 'researchers'))
# + [markdown] Collapsed="false" id="C3mPj2JP7dPZ"
# ## More tutorials
#
# * Doing this at scale: see the [Working with lists tutorial](https://api-lab.dimensions.ai/cookbooks/1-getting-started/6-Working-with-lists.html) that shows how to deal with list of researchers / grants of any size, using pagination and 'chunking' methods
# * Makings the analysis more specific by using the [investigators details data structure](https://docs.dimensions.ai/dsl/datasource-grants.html#grants-investigators-long-desc) in the API grants model
# * Going from grants to [Related publications](https://api-lab.dimensions.ai/cookbooks/3-grants/2-grants-enrichment-adding-publications-information.html)
# * Going from grants to [Related Patents and Clinical Trials](https://api-lab.dimensions.ai/cookbooks/3-grants/3-grants-enrichment-adding-patents-cltrials-information.html)
#
| archive/2022-02-BRO-Training/2022-02-BRO-Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GiulioCMSanto/HDSIdent/blob/master/notebooks/MIMO%20Systems/Segmentation/numerical_conditioning_mimo_laguerre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8CTy2vo0rzZR"
# # Numerical Conditioning: MIMO Laguerre Approach
#
# [HDSIdent: Historical Data Segmentation for System Identification](https://github.com/GiulioCMSanto/HDSIdent)
#
# This notebook explores how to obtain intervals suitable for system identification through a numerical conditioning Laguerre Filter approach, considering multivariable systems.
#
# **How to reference this work?**
#
# [SANTO, <NAME>. Data Mining Techniques Applied to Historical Data of Industrial Processes
# as a Tool to Find Time Intervals Suitable for System Identification. Masters dissertation
# – Polytechnic School of the University of São Paulo, São Paulo, Brasil, 2020.
# DOI: 10.13140/RG.2.2.13295.46240](https://www.researchgate.net/publication/347511108_Data_Mining_Techniques_Applied_to_Historical_Data_of_Industrial_Processes_as_a_Tool_to_Find_Time_Intervals_Suitable_for_System_Identification?channel=doi&linkId=5fdf5293a6fdccdcb8e856c4&showFulltext=true)
#
#
#
# + [markdown] id="IUqMw5e0IiHL"
# **About the Method**
#
# The multivariable approach here presented is inspired in the following work:
#
# ```
# PATEL, A. Data Mining of Process Data in Mutlivariable Systems.
# Degree project in electrical engineering — Royal Institute of Technology,
# Stockholm, Sweden, 2016.
# ```
#
# This approach uses concepts originally proposed in:
#
# ```
# <NAME>. et al. Data mining of historic data for process identification.
# In: Proceedings of the 2011 AIChE Annual Meeting, p. 1027–1033, 2011.
#
# <NAME>. et al. An algorithm for finding process identification
# intervals from normal operating data. Processes, v. 3, p. 357–383, 2015.
#
# <NAME>.; <NAME>. Selecting transients automatically for the
# identification of models for an oil well. IFAC-PapersOnLine, v. 48, n. 6, p.
# 154–158, 2015
# ```
#
# An interesting related work is the following:
#
# ```
# <NAME>.; <NAME>. A Search Method for Selecting Informative Data in Predominantly
# Stationary Historical Records for Multivariable System Identification.
# In: Proceedings of the 21st International Conference on System Theory,
# Control and Computing (ICSTCC). Sinaia, Romenia: IEEE, 2017a. p. 100–105.
# ```
# + [markdown] id="VQ0ojKXSIiYn"
# **About the [Dataset](https://github.com/GiulioCMSanto/HDSIdent/tree/master/data/distillation_column)**
#
# The dataset here adopted was produced through simulation in the dissertation (SANTO, <NAME>., 2020). The transfer functions adopted in the simulation were directly extracted from (WOOD; BERRY, 1973) and the operating conditions adopted were extracted from (JULIANI, 2017). The simulation idea was based in (PATEL, 2016), with similar signals being produced.
#
# **References**:
#
# ```
# SANTO, <NAME>. Data Mining Techniques Applied to Historical Data of Industrial Processes
# as a Tool to Find Time Intervals Suitable for System Identification. Masters dissertation
# – Polytechnic School of the University of São Paulo, São Paulo, Brasil, 2020.
# DOI: 10.13140/RG.2.2.13295.46240.
#
# <NAME>. Plantwide control: a review and proposal of an augmented
# hierarchical plantwide control design technique. Thesis — Polytechnic School of
# the University of São Paulo, São Paulo, Brasil, 2017.
#
# PATEL, A. Data Mining of Process Data in Mutlivariable Systems. 606–610 p.
# Degree project in electrical engineering — Royal Institute of Technology,
# Stockholm, Sweden, 2016.
#
# <NAME>.; <NAME>. Terminal composition control of a binary distillation
# column. Chemical Engineering Science, v. 28, n. 9, p. 1707–1717, 1973.
# ```
# + id="FN4OPByxO5l2" colab={"base_uri": "https://localhost:8080/"} outputId="848665e3-a639-46f6-f680-f5ad5455c4ff"
# !git clone https://github.com/GiulioCMSanto/HDSIdent.git
# + id="14Cd790WO_V8" colab={"base_uri": "https://localhost:8080/"} outputId="9108d48c-5a7d-40af-9b6a-27ab411382ca"
# Change into the directory for install
# %cd HDSIdent/
# + id="Hi0f1L7gPHxY" colab={"base_uri": "https://localhost:8080/"} outputId="dc9f6e81-5b03-4602-b348-c77f2b14f87c"
# !python setup.py install
# + id="__KZwYwlPDGq"
import pandas as pd
import numpy as np
from scipy.stats import chi2
import matplotlib.pyplot as plt
import seaborn as sns
from time import time
import plotly
import plotly.graph_objects as go
from plotly.offline import init_notebook_mode
plotly.io.renderers.default = 'colab'
# %matplotlib inline
sns.set_style('darkgrid')
# + id="UAKS4RjHPI5E"
from HDSIdent.data_treatment.data_preprocessing import Preprocessing
from HDSIdent.initial_intervals.exponentially_weighted import ExponentiallyWeighted
from HDSIdent.initial_intervals.bandpass_filter import BandpassFilter
from HDSIdent.initial_intervals.sliding_window import SlidingWindow
from HDSIdent.segmentation_methods.mimo_segmentation import MIMOSegmentation
from HDSIdent.model_structures.ar_structure import ARStructure
from HDSIdent.model_structures.arx_structure import ARXStructure
from HDSIdent.model_structures.laguerre_filter import LaguerreStructure
# + [markdown] id="fguMLxwtPSFf"
# ## **1. Read Data**
# + id="Z_-eww7_PLXi"
u1_url = "https://raw.githubusercontent.com/GiulioCMSanto/HDSIdent/master/data/distillation_column/mimo_simu_u1.csv"
u2_url = "https://raw.githubusercontent.com/GiulioCMSanto/HDSIdent/master/data/distillation_column/mimo_simu_u2.csv"
y1_url = "https://raw.githubusercontent.com/GiulioCMSanto/HDSIdent/master/data/distillation_column/mimo_simu_y1.csv"
y2_url = "https://raw.githubusercontent.com/GiulioCMSanto/HDSIdent/master/data/distillation_column/mimo_simu_y2.csv"
# + id="nq7wJ0HjPwLH"
u1 = pd.read_csv(u1_url, error_bad_lines=False, header=None)
u2 = pd.read_csv(u2_url, error_bad_lines=False, header=None)
y1 = pd.read_csv(y1_url, error_bad_lines=False, header=None)
y2 = pd.read_csv(y2_url, error_bad_lines=False, header=None)
# + [markdown] id="SuyuK4c7P7MO"
# ## **2. Data Preprocessing**
# + id="pUG77dj1P4DH"
pp = Preprocessing(
scaler='MinMaxScaler',
feature_range=(-0.5,0.5),
k=100);
# + id="08dMbC_sP-b2"
X_clean, Y_clean = pp.fit_transform(X=np.concatenate([u1,u2],axis=1),
y=np.concatenate([y1,y2],axis=1))
# + [markdown] id="wKSBFMNAQH-M"
# ## **3. Define Potential Intervals - Exponentially Weighted Moving Average (EWMA) Filter**
# + id="T47j_nFzP_Yi"
df = pd.DataFrame()
df['U1'] = np.squeeze(X_clean[:,0])
df['U2'] = np.squeeze(X_clean[:,1])
df['Y1'] = np.squeeze(Y_clean[:,0])
df['Y2'] = np.squeeze(Y_clean[:,1])
# + id="0w7i5Es0QNXK"
EW = ExponentiallyWeighted(
forgetting_fact_v = np.array([0.006,0.006,0.006,0.006]),
forgetting_fact_u = np.array([0.006,0.006,0.006,0.006]),
H_v = [0.005,0.005,0.005,0.005],
num_previous_indexes=50,
verbose=0,
n_jobs=-1);
EW.fit(X=df[['U1','U2']], y=df[['Y1','Y2']]);
# + id="RfH_gMzDQTTx" colab={"base_uri": "https://localhost:8080/", "height": 729} outputId="06e90499-88f0-4379-97b4-1a365c16c729"
plt.figure(figsize=(14,10));
plt.subplot(4,1,1);
plt.plot(X_clean[:,0], color='darkred');
plt.title("Reflux Flow Rate", fontsize=20);
plt.ylabel("Flow rate (lb/s)", fontsize=20);
plt.xticks(fontsize=20);
plt.yticks(fontsize=20);
for key, interval in EW.unified_intervals.items():
plt.axvline(np.min(interval))
plt.axvline(np.max(interval))
plt.subplot(4,1,2);
plt.plot(X_clean[:,1], color='darkgreen');
plt.title("Steam Flow Rate", fontsize=20);
plt.ylabel("Flow rate (lb/s)", fontsize=20);
plt.xticks(fontsize=20);
plt.yticks(fontsize=20);
for key, interval in EW.unified_intervals.items():
plt.axvline(np.min(interval))
plt.axvline(np.max(interval))
plt.subplot(4,1,3);
plt.plot(Y_clean[:,0], color='darkmagenta');
plt.title("Overhead Composition", fontsize=20);
plt.ylabel("Composition (%)", fontsize=20);
plt.xticks(fontsize=20);
plt.yticks(fontsize=20);
for key, interval in EW.unified_intervals.items():
plt.axvline(np.min(interval))
plt.axvline(np.max(interval))
plt.subplot(4,1,4);
plt.plot(Y_clean[:,1], color='purple');
plt.title("Bottom Composition", fontsize=20);
plt.ylabel("Composition (%)", fontsize=20);
plt.xlabel("Time (Minutes)", fontsize=20);
plt.xticks(fontsize=20);
plt.yticks(fontsize=20);
for key, interval in EW.unified_intervals.items():
plt.axvline(np.min(interval))
plt.axvline(np.max(interval))
plt.tight_layout();
# + [markdown] id="0XD9rtYyQnZT"
# ## **4. Apply Laguere Filter Method**
# + id="6R4YynkmQbKD"
LG = LaguerreStructure(
Nb=10,
p=0.92,
delay=10,
cc_alpha=0.05,
initial_intervals=EW.unified_intervals,
efr_type='type_2',
sv_thr=0.5,
n_jobs = -1,
verbose = 0
)
# + id="hE32sVUzQuNF"
start = time()
LG.fit(X=df[['U1','U2']],
y=df[['Y1','Y2']]);
end = time()
# + id="EUpnX4YfQz--" colab={"base_uri": "https://localhost:8080/"} outputId="6b0fdd5e-9cca-4acf-c58b-d11dd32f24de"
print("Execution Time: {}".format(end-start))
# + id="5RCSI2IAQ15r" colab={"base_uri": "https://localhost:8080/", "height": 189} outputId="0989a714-acee-4155-90f3-d28bfaa433cc"
pd.DataFrame(LG.miso_ranks).T
# + id="7NbMo9XcQ3Gw" colab={"base_uri": "https://localhost:8080/", "height": 189} outputId="9bc7f778-da63-4e58-e902-6b95d8836bdd"
pd.DataFrame(LG.cond_num_dict).T
# + id="1NR3X118RPeH" colab={"base_uri": "https://localhost:8080/", "height": 189} outputId="3bc451f4-7084-49b3-8030-3b31a34da0de"
pd.DataFrame(LG.chi_squared_dict).T
# + id="T-RDXoMwRRPo" colab={"base_uri": "https://localhost:8080/", "height": 189} outputId="d522aea4-ae09-432e-a539-8272e1c564ae"
pd.DataFrame(LG.miso_correlations).T
# + [markdown] id="5DOX2PxXRb4P"
# ### **4.1 Case 1: at least one input-output pair must satisfy the required criteria**
# + id="N81Ir28JRSXW"
MS_1 = MIMOSegmentation(
model_structure=[LG],
segmentation_method=['method1'],
parameters_dict={'Laguerre':{'chi2_p_value_thr':0.01,
'cond_thr':15000,
'min_input_coupling':1,
'min_output_coupling':1}
},
segmentation_type='stationary',
n_jobs=-1,
verbose=1);
# + id="p5mjkNycRgkO" colab={"base_uri": "https://localhost:8080/"} outputId="3a6c68fc-2fba-4199-9272-4cc6b3b5abc1"
MS_1.fit(X=df[['U1','U2']],
y=df[['Y1','Y2']]);
# + id="rGzStLLERiBO" colab={"base_uri": "https://localhost:8080/"} outputId="d37d3507-aef0-4c6c-dca3-f8512a4ead6b"
print("Approved Intervals: {}".format(
MS_1.sucessed_intervals['method1']['Laguerre'].keys()))
# + [markdown] id="p67NoWONRwD9"
# ### **4.2 Case 2: all inputs and all outputs must satisfy the required criteria**
# + id="n2LdptfeRlnH"
MS_2 = MIMOSegmentation(
model_structure=[LG],
segmentation_method=['method1'],
parameters_dict={'Laguerre':{'chi2_p_value_thr':0.01,
'cond_thr':15000,
'min_input_coupling':2,
'min_output_coupling':2}
},
segmentation_type='stationary',
n_jobs=-1,
verbose=1);
# + id="0Nc6eFEOSAec" colab={"base_uri": "https://localhost:8080/"} outputId="bf326e91-b8de-4928-b28c-2c1ca74be60b"
MS_2.fit(X=df[['U1','U2']],
y=df[['Y1','Y2']]);
# + id="fHiDveTXSBv1" colab={"base_uri": "https://localhost:8080/"} outputId="7343a16a-d51c-42e1-bda0-7b50ccb02da3"
print("Approved Intervals: {}".format(
MS_2.sucessed_intervals['method1']['Laguerre'].keys()))
| notebooks/MIMO Systems/Segmentation/numerical_conditioning_mimo_laguerre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cxbxmxcx/Evolutionary-Deep-Learning/blob/main/EDL_5_DE_HPO_PCA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7eo9h5KA5UPe"
# # Setup
# + colab={"base_uri": "https://localhost:8080/"} id="jpHn_-RfV0_n" outputId="d994134e-4469-4916-a5e6-7b8a265e3110"
#@title Install DEAP
# !pip install deap --quiet
# + id="FslekLiVp_Il"
#@title Defining Imports
#numpy
import numpy as np
#DEAP
from deap import algorithms
from deap import base
from deap import benchmarks
from deap import creator
from deap import tools
#PyTorch
import torch
import torch.nn as nn
from torch.autograd import Variable
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import TensorDataset, DataLoader
#SkLearn
from sklearn.decomposition import PCA
#plotting
from matplotlib import pyplot as plt
from matplotlib import cm
from IPython.display import clear_output
#utils
import random
import math
import array
import time
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="MQCdK2oWJlid" outputId="238daf7f-25f4-4557-acd6-d2871d80c094"
#@title Setup Target Function and Data
def function(x):
return (2*x + 3*x**2 + 4*x**3 + 5*x**4 + 6*x**5 + 10)
data_min = -5
data_max = 5
data_step = .5
Xi = np.reshape(np.arange(data_min, data_max, data_step), (-1, 1))
yi = function(Xi)
inputs = Xi.shape[1]
yi = yi.reshape(-1, 1)
plt.plot(Xi, yi, 'o', color='black')
# + id="eY30bijNqNBL"
#@title Define the Model
class Net(nn.Module):
def __init__(self, inputs, middle):
super().__init__()
self.fc1 = nn.Linear(inputs,middle)
self.fc2 = nn.Linear(middle,middle)
self.out = nn.Linear(middle,1)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.out(x)
return x
# + id="nsBRAusqy7Uz"
#@title Define HyperparametersEC Class
class HyperparametersEC(object):
def __init__(self, **kwargs):
self.__dict__.update(kwargs)
self.hparms = [d for d in self.__dict__]
def __str__(self):
out = ""
for d in self.hparms:
ds = self.__dict__[d]
out += f"{d} = {ds} "
return out
def values(self):
vals = []
for d in self.hparms:
vals.append(self.__dict__[d])
return vals
def size(self):
return len(self.hparms)
def next(self, individual):
dict = {}
#initialize generators
for i, d in enumerate(self.hparms):
next(self.__dict__[d])
for i, d in enumerate(self.hparms):
dict[d] = self.__dict__[d].send(individual[i])
return HyperparametersEC(**dict)
def clamp(num, min_value, max_value):
return max(min(num, max_value), min_value)
def linespace(min,max):
rnge = max - min
while True:
i = yield
i = (clamp(i, -1.0, 1.0) + 1.0) / 2.0
yield i * rnge + min
def linespace_int(min,max):
rnge = max - min
while True:
i = yield
i = (clamp(i, -1.0, 1.0) + 1.0) / 2.0
yield int(i * rnge) + min
def static(val):
while True:
yield val
# + [markdown] id="2Y9-31iHVPqm"
# # Create the HyperparamtersEC Object
# + colab={"base_uri": "https://localhost:8080/"} id="ciFUE2XDzhMk" outputId="e978e725-1d9c-46d8-b64a-2efb632d2a69"
#@title Instantiate the HPO
hp = HyperparametersEC(
middle_layer = linespace_int(8, 64),
learning_rate = linespace(3.5e-02,3.5e-01),
batch_size = linespace_int(4,20),
epochs = linespace_int(50,400)
)
ind = [-.5, -.3, -.1, .8]
print(hp.next(ind))
# + colab={"base_uri": "https://localhost:8080/"} id="ubSMH-0uhuCO" outputId="2bf1cc33-da7a-4662-e4f4-00a0d9bdd1b3"
#@title Check for Cuda/GPU
cuda = True if torch.cuda.is_available() else False
print("Using CUDA" if cuda else "Not using CUDA")
Tensor = torch.cuda.FloatTensor if cuda else torch.Tensor
# + colab={"base_uri": "https://localhost:8080/", "height": 276} id="PXPpiBDyTRXh" outputId="4b818847-4749-4fac-baf6-523a523b543e"
#@title Setup Principle Component Analysis
#create example individuals
pop = np.array([[-.5, .75, -.1, .8], [-.5, -.3, -.5, .8]])
pca = PCA(n_components=2)
reduced = pca.fit_transform(pop)
t = reduced.transpose()
plt.scatter(t[0], t[1])
plt.show()
# + [markdown] id="SiomzsQfWoL5"
# [link text](https://)# Setup DEAP for DE Search
# + id="yTrNof0Cub4F"
#@title DE Bounding Hyperparameters
NDIM = hp.size()
CR = 0.25
F_ = 1
MU = 50
NGEN = 10
# + id="j6sqsbkKWmw_"
#@title Setup Fitness Criteria
creator.create("FitnessMin", base.Fitness, weights=(-1.0,))
creator.create("Individual", array.array, typecode='d', fitness=creator.FitnessMin)
# + id="J161LLLguzUf"
#@title Add Genetic Operators to Toolbox
toolbox = base.Toolbox()
toolbox.register("attr_float", random.uniform, -1, 1)
toolbox.register("individual", tools.initRepeat, creator.Individual, toolbox.attr_float, NDIM)
toolbox.register("population", tools.initRepeat, list, toolbox.individual)
toolbox.register("select", tools.selRandom, k=3)
# + [markdown] id="zCbrGhsr6IhK"
# # Create a Training Function
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="je4Jf9Vp88Rq" outputId="388e7037-1af8-407d-a9f3-ec46527ec9a9"
#@title Wrapper Function for DL
loss_fn = nn.MSELoss()
if cuda:
loss_fn.cuda()
def train_function(hp):
X = np.reshape(
np.arange(
data_min,
data_max,
data_step)
, (-1, 1))
y = function(X)
inputs = X.shape[1]
tensor_x = torch.Tensor(X) # transform to torch tensor
tensor_y = torch.Tensor(y)
dataset = TensorDataset(tensor_x,tensor_y) # create your datset
dataloader = DataLoader(dataset, batch_size= hp.batch_size, shuffle=True) # create your dataloader
model = Net(inputs, hp.middle_layer)
optimizer = optim.Adam(model.parameters(), lr=hp.learning_rate)
if cuda:
model.cuda()
history=[]
start = time.time()
for i in range(hp.epochs):
for X, y in iter(dataloader):
# wrap the data in variables
x_batch = Variable(torch.Tensor(X).type(Tensor))
y_batch = Variable(torch.Tensor(y).type(Tensor))
# forward pass
y_pred = model(x_batch)
# compute and print loss
loss = loss_fn(y_pred, y_batch)
ll = loss.data
history.append(ll)
# reset gradients
optimizer.zero_grad()
# backwards pass
loss.backward()
# step the optimizer - update the weights
optimizer.step()
end = time.time() - start
return end, history, model, hp
hp_in = hp.next(ind)
span, history, model, hp_out = train_function(hp_in)
plt.plot(history)
print(min(history).item())
# + [markdown] id="NRQ0U-6UaUlJ"
# # DE Evaluate Function in Toolbox
# + id="SfgZvw9haOQX"
#@title Create Evaluation Function and Register
run_history = []
def evaluate(individual):
hp_in = hp.next(individual)
span, history, model, hp_out = train_function(hp_in)
y_ = model(torch.Tensor(Xi).type(Tensor))
fitness = loss_fn(y_, torch.Tensor(yi).type(Tensor)).data.item()
run_history.append([fitness,*hp_out.values()])
return fitness,
# fitness eval
toolbox.register("evaluate", evaluate)
# + [markdown] id="8-M8o_OJVjg5"
# # Perform the HPO
# + id="xxp8Ir2kvQ6N"
random.seed(64)
pop = toolbox.population(n=MU);
hof = tools.HallOfFame(1)
stats = tools.Statistics(lambda ind: ind.fitness.values)
stats.register("avg", np.mean)
stats.register("std", np.std)
stats.register("min", np.min)
stats.register("max", np.max)
logbook = tools.Logbook()
logbook.header = "gen", "evals", "std", "min", "avg", "max"
# Evaluate the individuals
fitnesses = toolbox.map(toolbox.evaluate, pop)
for ind, fit in zip(pop, fitnesses):
ind.fitness.values = fit
record = stats.compile(pop)
logbook.record(gen=0, evals=len(pop), **record)
print(logbook.stream)
start = time.time()
for g in range(1, NGEN):
for k, agent in enumerate(pop):
a,b,c = toolbox.select(pop)
y = toolbox.clone(agent)
index = random.randrange(NDIM)
for i, value in enumerate(agent):
if i == index or random.random() < CR:
y[i] = a[i] + F_*(b[i]-c[i])
y.fitness.values = toolbox.evaluate(y)
if y.fitness > agent.fitness:
pop[k] = y
hof.update(pop)
record = stats.compile(pop)
#logbook.record(gen=g, evals=len(pop), **record)
best = hof[0]
span, history, model, hp_out = train_function(hp.next(best))
y_ = model(torch.Tensor(Xi).type(Tensor))
fitness = loss_fn(y_, torch.Tensor(yi).type(Tensor)).data.item()
run_history.append([fitness,*hp_out.values()])
best_hp = hp_out
clear_output()
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(18,6))
fig.suptitle(f"Best Fitness {best.fitness} \n{best_hp}")
fig.text(0,0,f"Generation {g+1}/{NGEN} Current Fitness {fitness} \n{hp_out}")
ax1.plot(history)
ax1.set_xlabel("iteration")
ax1.set_ylabel("loss")
ax2.plot(Xi, yi, 'o', color='black')
ax2.plot(Xi,y_.detach().cpu().numpy(), 'r')
ax2.set_xlabel("X")
ax2.set_ylabel("Y")
rh = np.array(run_history)
M = rh[:,1:NDIM+1]
reduced = pca.fit_transform(M)
t = reduced.transpose()
hexbins = ax3.hexbin(t[0], t[1], C=rh[:, 0],
bins=50, gridsize=50, cmap=cm.get_cmap('gray'))
ax3.set_xlabel("PCA X")
ax3.set_ylabel("PCA Y")
plt.show()
time.sleep(1)
| EDL_5_DE_HPO_PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!-- <div>
# <img src="attachment:image.png" align="right" width="150">
# </div> -->
#
# # <font color='#5D6D7E '> <center >Forecasting - AVAX</center>
#
# ### Master Degree Program in Data Science and Advanced Analytics
#
# ### <font color='#5D6D7E '> Business Cases with Data Science Project:
# > #### Group AA
#
# ### <font color='#5D6D7E '> Done by:
# > #### - <NAME>, m20210545
# > #### - <NAME>, m20210547
# > #### - <NAME>, m20201076
# > #### - <NAME>, m20210587
# ---
# <div>
#
# # Table of Content<a class="anchor"><a id='toc'></a>
#
# ### <font color='#5D6D7E '> Import and Data Integration
# - [<font color='#000000'>Import the needed Libraries</font>](#third-bullet)<br>
#
# ### <font color='#5D6D7E '> Data Exploration and Understanding
# - [<font color='#000000'>Initial Analysis (EDA - Exploratory Data Analysis)</font>](#fifth-bullet)<br>
# - [<font color='#000000'>Variables Distribution</font>](#seventh-bullet)<br>
#
# ### <font color='#5D6D7E '> Data Preparation
# - [<font color='#000000'>Data Transformation</font>](#eighth-bullet)<br>
#
# ### <font color='#5D6D7E '> Modelling
# - [<font color='#000000'>Building LSTM Model</font>](#twentysecond-bullet)<br>
# - [<font color='#000000'>Get Best Parameters for LSTM</font>](#twentythird-bullet)<br>
# - [<font color='#000000'>Run the LSTM Model and Get Predictions</font>](#twentyfourth-bullet)<br>
# - [<font color='#000000'>Recursive Predictions</font>](#twentysixth-bullet)<br>
#
#
# </div>
# ---
# # Import and Data Integration
#
#
# ## <font color='#5D6D7E '>Import the needed Libraries</font> <a class="anchor" id="third-bullet"></a>
# [Back to TOC](#toc)
# +
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# # Data Exploration and Understanding
#
# ## <font color='#5D6D7E'>Initial Analysis (EDA - Exploratory Data Analysis) </font> <a class="anchor" id="fifth-bullet"></a>
# [Back to TOC](#toc)
#
df = pd.read_csv('../data/data_aux/df_AVAX.csv')
df
# ### Data Types
# Get to know the number of instances and Features, the DataTypes and if there are missing values in each Feature
df.info()
# ### Missing Values
# Count the number of missing values for each Feature
df.isna().sum().to_frame().rename(columns={0: 'Count Missing Values'})
# ### Descriptive Statistics
# Descriptive Statistics Table
df.describe().T
# +
# settings to display all columns
pd.set_option("display.max_columns", None)
# display the dataframe head
df.sample(n=10)
# -
#CHECK ROWS THAT HAVE ANY MISSING VALUE IN ONE OF THE COLUMNS
is_NaN = df.isnull()
row_has_NaN = is_NaN.any(axis=1)
rows_with_NaN = df[row_has_NaN]
rows_with_NaN
#FILTER OUT ROWS THAT ARE MISSING INFORMATION
df = df[~row_has_NaN]
df.reset_index(inplace=True, drop=True)
df
# # Data Preparation
#
#
# ## <font color='#5D6D7E'>Data Transformation</font> <a class="anchor" id="eighth-bullet"></a>
# [Back to TOC](#toc)
# __`Duplicates`__
# Checking if exist duplicated observations
print(f'\033[1m' + "Number of duplicates: " + '\033[0m', df.duplicated().sum())
# __`Convert Date to correct format`__
df['Date'] = pd.to_datetime(df['Date'], format='%Y-%m-%d')
df
# __`Get percentual difference between open and close values and low and high values`__
df['pctDiff_CloseOpen'] = abs((df[df.columns[2]]-df[df.columns[5]])/df[df.columns[2]])*100
df['pctDiff_HighLow'] = abs((df[df.columns[3]]-df[df.columns[4]])/df[df.columns[4]])*100
df.head()
def plot_coinValue(df):
#Get coin name
coin_name = df.columns[2].split('-')[0]
#Get date and coin value
x = df['Date']
y = df[df.columns[2]] # ADA-USD_CLOSE
#Get the volume of trades
v = df[df.columns[-3]]/1e9
#Get percentual diferences
y2 = df[df.columns[-1]] # pctDiff_HighLow
y1= df[df.columns[-2]] # pctDiff_CloseOpen
fig, axs = plt.subplots(3, 1, figsize=(12,14))
axs[0].plot(x, y)
axs[2].plot(x, v)
# plotting the line 1 points
axs[1].plot(x, y1, label = "Close/Open")
# plotting the line 2 points
axs[1].plot(x, y2, label = "High/Low")
axs[1].legend()
axs[0].title.set_text('Time Evolution of '+ coin_name)
axs[0].set(xlabel="", ylabel="Close Value in USD$")
axs[2].title.set_text('Volume of trades of '+ coin_name)
axs[2].set(xlabel="", ylabel="Total number of trades in billions")
axs[1].title.set_text('Daily Market percentual differences of '+ coin_name)
axs[1].set(xlabel="", ylabel="Percentage (%)")
plt.savefig('../analysis/'+coin_name +'_stats'+'.png')
return coin_name
coin_name = plot_coinValue(df)
#FILTER DATASET
df = df.loc[df['Date']>= '2021-09-01']
df
# # Modelling
#
#
# ## <font color='#5D6D7E'>Building LSTM Model</font> <a class="anchor" id="twentysecond-bullet"></a>
# [Back to TOC](#toc)
# ## Strategy
#
# Create a DF (windowed_df) where the middle columns will correspond to the close values of X days before the target date and the final column will correspond to the close value of the target date. Use these values for prediction and play with the value of X
def get_windowed_df(X, df):
start_Date = df['Date'] + pd.Timedelta(days=X)
perm = np.zeros((1,X+1))
#Get labels for DataFrame
j=1
labels=[]
while j <= X:
label = 'closeValue_' + str(j) + 'daysBefore'
labels.append(label)
j+=1
labels.append('closeValue')
for i in range(X,df.shape[0]):
temp = np.zeros((1,X+1))
#Date for i-th day
#temp[0,0] = df.iloc[i]['Date']
#Close values for k days before
for k in range(X):
temp[0,k] = df.iloc[i-k-1,2]
#Close value for i-th date
temp[0,-1] = df.iloc[i,2]
#Add values to the permanent frame
perm = np.vstack((perm,temp))
#Get the array in dataframe form
windowed_df = pd.DataFrame(perm[1:,:], columns = labels)
return windowed_df
#Get the dataframe and append the dates
windowed_df = get_windowed_df(15, df)
windowed_df['Date'] = df.iloc[15:]['Date'].reset_index(drop=True)
windowed_df
# +
#Get the X,y and dates into a numpy array to apply on a model
def windowed_df_to_date_X_y(windowed_dataframe):
df_as_np = windowed_dataframe.to_numpy()
dates = df_as_np[:, -1]
middle_matrix = df_as_np[:, 0:-2]
X = middle_matrix.reshape((len(dates), middle_matrix.shape[1], 1))
Y = df_as_np[:, -2]
return dates, X.astype(np.float32), Y.astype(np.float32)
dates, X, y = windowed_df_to_date_X_y(windowed_df)
dates.shape, X.shape, y.shape
# +
#Partition for train, validation and test
q_80 = int(len(dates) * .8)
q_90 = int(len(dates) * .9)
dates_train, X_train, y_train = dates[:q_80], X[:q_80], y[:q_80]
dates_val, X_val, y_val = dates[q_80:q_90], X[q_80:q_90], y[q_80:q_90]
dates_test, X_test, y_test = dates[q_90:], X[q_90:], y[q_90:]
fig,axs = plt.subplots(1, 1, figsize=(12,5))
#Plot the partitions
axs.plot(dates_train, y_train)
axs.plot(dates_val, y_val)
axs.plot(dates_test, y_test)
axs.legend(['Train', 'Validation', 'Test'])
fig.savefig('../analysis/'+coin_name +'_partition'+'.png')
# -
# ## <font color='#5D6D7E'>Get Best Parameters for LSTM</font> <a class="anchor" id="twentythird-bullet"></a>
# [Back to TOC](#toc)
# +
# #!pip install tensorflow
# +
#import os
#os.environ['PYTHONHASHSEED']= '0'
#import numpy as np
#np.random.seed(1)
#import random as rn
#rn.seed(1)
#import tensorflow as tf
#tf.random.set_seed(1)
#
#from tensorflow.keras.models import Sequential
#from tensorflow.keras.optimizers import Adam
#from tensorflow.keras import layers
#from sklearn.metrics import mean_squared_error
#
## Function to create LSTM model and compute the MSE value for the given parameters
#def check_model(X_train, y_train, X_val, y_val, X_test, y_test, learning_rate,epoch,batch):
#
# # create model
# model = Sequential([layers.Input((15, 1)),
# layers.LSTM(64),
# layers.Dense(32, activation='relu'),
# layers.Dense(32, activation='relu'),
# layers.Dense(1)])
# # Compile model
# model.compile(loss='mse', optimizer=Adam(learning_rate=learning_rate), metrics=['mean_absolute_error'])
#
# model.fit(X_train, y_train, validation_data=(X_val, y_val), epochs=epoch, shuffle=False, batch_size=batch, verbose=2)
#
# test_predictions = model.predict(X_test).flatten()
#
# LSTM_mse = mean_squared_error(y_test, test_predictions)
#
# return LSTM_mse
#
##Function that iterates the different parameters and gets the ones corresponding to the lowest MSE score.
#def search_parameters(batch_size, epochs, learn_rate, X_train, y_train, X_val, y_val, X_test, y_test):
#
# best_score = float('inf')
#
# for b in batch_size:
# for e in epochs:
# for l in learn_rate:
# print('Batch Size: ' + str(b))
# print('Number of Epochs: ' + str(e))
# print('Value of Learning Rate: ' + str(l))
# try:
# mse = check_model(X_train, y_train, X_val, y_val, X_test, y_test,l,e,b)
# print('MSE=%.3f' % (mse))
# if mse < best_score:
# best_score = mse
# top_params = [b, e, l]
# except:
# continue
#
# print('Best MSE=%.3f' % (best_score))
# print('Optimal Batch Size: ' + str(top_params[0]))
# print('Optimal Number of Epochs: ' + str(top_params[1]))
# print('Optimal Value of Learning Rate: ' + str(top_params[2]))
#
#
## define parameters
#batch_size = [10, 100, 1000]
#epochs = [50, 100]
#learn_rate = np.linspace(0.001,0.1, num=10)
#
#warnings.filterwarnings("ignore")
#search_parameters(batch_size, epochs, learn_rate, X_train, y_train, X_val, y_val, X_test, y_test)
# -
# ## <font color='#5D6D7E'>Run the LSTM Model and Get Predictions</font> <a class="anchor" id="twentyfourth-bullet"></a>
# [Back to TOC](#toc)
# +
#BEST SOLUTION OF THE MODEL
# MSE=48.801
# Batch Size: 10
# Number of Epochs: 100
# Value of Learning Rate: 0.012
model = Sequential([layers.Input((15, 1)),
layers.LSTM(64),
layers.Dense(32, activation='relu'),
layers.Dense(32, activation='relu'),
layers.Dense(1)])
model.compile(loss='mse',
optimizer=Adam(learning_rate=0.012),
metrics=['mean_absolute_error'])
model.fit(X_train, y_train, validation_data=(X_val, y_val), epochs=100, shuffle=False, batch_size=10, verbose=2)
# +
#PREDICT THE VALUES USING THE MODEL
train_predictions = model.predict(X_train).flatten()
val_predictions = model.predict(X_val).flatten()
test_predictions = model.predict(X_test).flatten()
fig,axs = plt.subplots(3, 1, figsize=(14,14))
axs[0].plot(dates_train, train_predictions)
axs[0].plot(dates_train, y_train)
axs[0].legend(['Training Predictions', 'Training Observations'])
axs[1].plot(dates_val, val_predictions)
axs[1].plot(dates_val, y_val)
axs[1].legend(['Validation Predictions', 'Validation Observations'])
axs[2].plot(dates_test, test_predictions)
axs[2].plot(dates_test, y_test)
axs[2].legend(['Testing Predictions', 'Testing Observations'])
plt.savefig('../analysis/LTSM_recursive/'+coin_name +'_modelPredictions'+'.png')
# -
# ## <font color='#5D6D7E'>Recursive Predictions</font> <a class="anchor" id="twentysixth-bullet"></a>
# [Back to TOC](#toc)
# +
from copy import deepcopy
#Get prediction for future dates recursively based on the previous existing information. Then update the window of days upon
#which the predictions are made
recursive_predictions = []
recursive_dates = np.concatenate([dates_test])
last_window = deepcopy(X_train[-1])
for target_date in recursive_dates:
next_prediction = model.predict(np.array([last_window])).flatten()
recursive_predictions.append(next_prediction)
last_window = np.insert(last_window,0,next_prediction)[:-1]
# +
fig,axs = plt.subplots(2, 1, figsize=(14,10))
axs[0].plot(dates_train, train_predictions)
axs[0].plot(dates_train, y_train)
axs[0].plot(dates_val, val_predictions)
axs[0].plot(dates_val, y_val)
axs[0].plot(dates_test, test_predictions)
axs[0].plot(dates_test, y_test)
axs[0].plot(recursive_dates, recursive_predictions)
axs[0].legend(['Training Predictions',
'Training Observations',
'Validation Predictions',
'Validation Observations',
'Testing Predictions',
'Testing Observations',
'Recursive Predictions'])
axs[1].plot(dates_test, y_test)
axs[1].plot(recursive_dates, recursive_predictions)
axs[1].legend(['Testing Observations',
'Recursive Predictions'])
plt.savefig('../analysis/LTSM_recursive/'+ coin_name +'_recursivePredictions'+'.png')
# -
| BC4_crypto_forecasting/scripts/AVAX_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qiskit Visualizations
from qiskit import *
from qiskit.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
# ## Plot histogram <a name='histogram'></a>
#
# To visualize the data from a quantum circuit run on a real device or `qasm_simulator` we have made a simple function
#
# `plot_histogram(data)`
#
# As an example we make a 2-qubit Bell state
# +
# quantum circuit to make a Bell state
bell = QuantumCircuit(2, 2)
bell.h(0)
bell.cx(0, 1)
meas = QuantumCircuit(2, 2)
meas.measure([0,1], [0,1])
# execute the quantum circuit
backend = BasicAer.get_backend('qasm_simulator') # the device to run on
circ = bell + meas
result = execute(circ, backend, shots=1000).result()
counts = result.get_counts(circ)
print(counts)
# -
plot_histogram(counts)
# ### Options when plotting a histogram
#
# The `plot_histogram()` has a few options to adjust the output graph. The first option is the `legend` kwarg. This is used to provide a label for the executions. It takes a list of strings use to label each execution's results. This is mostly useful when plotting multiple execution results in the same histogram. The `sort` kwarg is used to adjust the order the bars in the histogram are rendered. It can be set to either ascending order with `asc` or descending order with `dsc`. The `number_to_keep` kwarg takes an integer for the number of terms to show, the rest are grouped together in a single bar called rest. You can adjust the color of the bars with the `color` kwarg which either takes a string or a list of strings for the colors to use for the bars for each execution. You can adjust whether labels are printed above the bars or not with the `bar_labels` kwarg. The last option available is the `figsize` kwarg which takes a tuple of the size in inches to make the output figure.
# Execute 2-qubit Bell state again
second_result = execute(circ, backend, shots=1000).result()
second_counts = second_result.get_counts(circ)
# Plot results with legend
legend = ['First execution', 'Second execution']
plot_histogram([counts, second_counts], legend=legend)
plot_histogram([counts, second_counts], legend=legend, sort='desc', figsize=(15,12),
color=['orange', 'black'], bar_labels=False)
# ### Using the output from plot_histogram()
#
# When using the plot_histogram() function it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object from `plot_histogram()` to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png`. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
# ## Plot State <a name='state'></a>
# In many situations you want to see the state of a quantum computer. This could be for debugging. Here we assume you have this state (either from simulation or state tomography) and the goal is to visualize the quantum state. This requires exponential resources, so we advise to only view the state of small quantum systems. There are several functions for generating different types of visualization of a quantum state
#
# ```
# plot_state_city(quantum_state)
# plot_state_qsphere(quantum_state)
# plot_state_paulivec(quantum_state)
# plot_state_hinton(quantum_state)
# plot_bloch_multivector(quantum_state)
# ```
#
# A quantum state is either a state matrix $\rho$ (Hermitian matrix) or statevector $|\psi\rangle$ (complex vector). The state matrix is related to the statevector by
#
# $$\rho = |\psi\rangle\langle \psi|,$$
#
# and is more general as it can represent mixed states (positive sum of statevectors)
#
# $$\rho = \sum_k p_k |\psi_k\rangle\langle \psi_k |.$$
#
# The visualizations generated by the functions are:
#
# - `'plot_state_city'`: The standard view for quantum states where the real and imaginary (imag) parts of the state matrix are plotted like a city.
#
# - `'plot_state_qsphere'`: The Qiskit unique view of a quantum state where the amplitude and phase of the state vector are plotted in a spherical ball. The amplitude is the thickness of the arrow and the phase is the color. For mixed states it will show different `'qsphere'` for each component.
#
# - `'plot_state_paulivec'`: The representation of the state matrix using Pauli operators as the basis $\rho=\sum_{q=0}^{d^2-1}p_jP_j/d$.
#
# - `'plot_state_hinton'`: Same as `'city'` but where the size of the element represents the value of the matrix element.
#
# - `'plot_bloch_multivector'`: The projection of the quantum state onto the single qubit space and plotting on a bloch sphere.
from qiskit.visualization import plot_state_city, plot_bloch_multivector
from qiskit.visualization import plot_state_paulivec, plot_state_hinton
from qiskit.visualization import plot_state_qsphere
# execute the quantum circuit
backend = BasicAer.get_backend('statevector_simulator') # the device to run on
result = execute(bell, backend).result()
psi = result.get_statevector(bell)
plot_state_city(psi)
plot_state_hinton(psi)
# + tags=["nbsphinx-thumbnail"]
plot_state_qsphere(psi)
# -
plot_state_paulivec(psi)
plot_bloch_multivector(psi)
# Here we see that there is no information about the quantum state in the single qubit space as all vectors are zero.
# ### Options when using state plotting functions
#
# The various functions for plotting quantum states provide a number of options to adjust how the plots are rendered. Which options are available depends on the function being used.
# **plot_state_city()** options
#
# - **title** (str): a string that represents the plot title
# - **figsize** (tuple): figure size in inches (width, height).
# - **color** (list): a list of len=2 giving colors for real and imaginary components of matrix elements.
plot_state_city(psi, title="My City", color=['black', 'orange'])
# **plot_state_hinton()** options
#
# - **title** (str): a string that represents the plot title
# - **figsize** (tuple): figure size in inches (width, height).
plot_state_hinton(psi, title="My Hinton")
# **plot_state_paulivec()** options
#
# - **title** (str): a string that represents the plot title
# - **figsize** (tuple): figure size in inches (width, height).
# - **color** (list or str): color of the expectation value bars.
plot_state_paulivec(psi, title="My Paulivec", color=['purple', 'orange', 'green'])
# **plot_state_qsphere()** options
#
# - **figsize** (tuple): figure size in inches (width, height).
# **plot_bloch_multivector()** options
#
# - **title** (str): a string that represents the plot title
# - **figsize** (tuple): figure size in inches (width, height).
plot_bloch_multivector(psi, title="My Bloch Spheres")
# ### Using the output from state plotting functions
#
# When using any of the state plotting functions it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png` in the current working directory. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
# ## Plot Bloch Vector <a name='bloch'></a>
#
# A standard way of plotting a quantum system is using the Bloch vector. This only works for a single qubit and takes as input the Bloch vector.
#
# The Bloch vector is defined as $[x = \mathrm{Tr}[X \rho], y = \mathrm{Tr}[Y \rho], z = \mathrm{Tr}[Z \rho]]$, where $X$, $Y$, and $Z$ are the Pauli operators for a single qubit and $\rho$ is the state matrix.
#
from qiskit.visualization import plot_bloch_vector
plot_bloch_vector([0,1,0])
# ### Options for plot_bloch_vector()
#
# - **title** (str): a string that represents the plot title
# - **figsize** (tuple): Figure size in inches (width, height).
plot_bloch_vector([0,1,0], title='My Bloch Sphere')
# ### Adjusting the output from plot_bloch_vector()
#
# When using the `plot_bloch_vector` function it returns a `matplotlib.Figure` for the rendered visualization. Jupyter notebooks understand this return type and render it for us in this tutorial, but when running outside of Jupyter you do not have this feature automatically. However, the `matplotlib.Figure` class natively has methods to both display and save the visualization. You can call `.show()` on the returned object to open the image in a new window (assuming your configured matplotlib backend is interactive). Or alternatively you can call `.savefig('out.png')` to save the figure to `out.png` in the current working directory. The `savefig()` method takes a path so you can adjust the location and filename where you're saving the output.
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| tutorials/circuits/2_plotting_data_in_qiskit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Face Recognition
#
# Welcome! In this assignment, you're going to build a face recognition system. Many of the ideas presented here are from [FaceNet](https://arxiv.org/pdf/1503.03832.pdf). In the lecture, you also encountered [DeepFace](https://research.fb.com/wp-content/uploads/2016/11/deepface-closing-the-gap-to-human-level-performance-in-face-verification.pdf).
#
# Face recognition problems commonly fall into one of two categories:
#
# **Face Verification** "Is this the claimed person?" For example, at some airports, you can pass through customs by letting a system scan your passport and then verifying that you (the person carrying the passport) are the correct person. A mobile phone that unlocks using your face is also using face verification. This is a 1:1 matching problem.
#
# **Face Recognition** "Who is this person?" For example, the video lecture showed a [face recognition video](https://www.youtube.com/watch?v=wr4rx0Spihs) of Baidu employees entering the office without needing to otherwise identify themselves. This is a 1:K matching problem.
#
# FaceNet learns a neural network that encodes a face image into a vector of 128 numbers. By comparing two such vectors, you can then determine if two pictures are of the same person.
#
# By the end of this assignment, you'll be able to:
#
# * Differentiate between face recognition and face verification
# * Implement one-shot learning to solve a face recognition problem
# * Apply the triplet loss function to learn a network's parameters in the context of face recognition
# * Explain how to pose face recognition as a binary classification problem
# * Map face images into 128-dimensional encodings using a pretrained model
# * Perform face verification and face recognition with these encodings
#
# **Channels-last notation**
#
# For this assignment, you'll be using a pre-trained model which represents ConvNet activations using a "channels last" convention, as used during the lecture and in previous programming assignments.
#
# In other words, a batch of images will be of shape $(m, n_H, n_W, n_C)$.
# ## Table of Contents
#
# - [1 - Packages](#1)
# - [2 - Naive Face Verification](#2)
# - [3 - Encoding Face Images into a 128-Dimensional Vector](#3)
# - [3.1 - Using a ConvNet to Compute Encodings](#3-1)
# - [3.2 - The Triplet Loss](#3-2)
# - [Exercise 1 - triplet_loss](#ex-1)
# - [4 - Loading the Pre-trained Model](#4)
# - [5 - Applying the Model](#5)
# - [5.1 - Face Verification](#5-1)
# - [Exercise 2 - verify](#ex-2)
# - [5.2 - Face Recognition](#5-2)
# - [Exercise 3 - who_is_it](#ex-3)
# - [6 - References](#6)
# <a name='1'></a>
# ## 1 - Packages
#
# Go ahead and run the cell below to import the packages you'll need.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, ZeroPadding2D, Activation, Input, concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import MaxPooling2D, AveragePooling2D
from tensorflow.keras.layers import Concatenate
from tensorflow.keras.layers import Lambda, Flatten, Dense
from tensorflow.keras.initializers import glorot_uniform
from tensorflow.keras.layers import Layer
from tensorflow.keras import backend as K
K.set_image_data_format('channels_last')
import os
import numpy as np
from numpy import genfromtxt
import pandas as pd
import tensorflow as tf
import PIL
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# <a name='2'></a>
# ## 2 - Naive Face Verification
#
# In Face Verification, you're given two images and you have to determine if they are of the same person. The simplest way to do this is to compare the two images pixel-by-pixel. If the distance between the raw images is below a chosen threshold, it may be the same person!
#
# <img src="images/pixel_comparison.png" style="width:380px;height:150px;">
# <caption><center> <u> <font color='purple'> <b>Figure 1</b> </u></center></caption>
#
# Of course, this algorithm performs poorly, since the pixel values change dramatically due to variations in lighting, orientation of the person's face, minor changes in head position, and so on.
#
# You'll see that rather than using the raw image, you can learn an encoding, $f(img)$.
#
# By using an encoding for each image, an element-wise comparison produces a more accurate judgement as to whether two pictures are of the same person.
# <a name='3'></a>
# ## 3 - Encoding Face Images into a 128-Dimensional Vector
#
# <a name='3-1'></a>
# ### 3.1 - Using a ConvNet to Compute Encodings
#
# The FaceNet model takes a lot of data and a long time to train. So following the common practice in applied deep learning, you'll load weights that someone else has already trained. The network architecture follows the Inception model from [Szegedy *et al*..](https://arxiv.org/abs/1409.4842) An Inception network implementation has been provided for you, and you can find it in the file `inception_blocks_v2.py` to get a closer look at how it is implemented.
#
# *Hot tip:* Go to "File->Open..." at the top of this notebook. This opens the file directory that contains the `.py` file).
#
# The key things to be aware of are:
#
# - This network uses 160x160 dimensional RGB images as its input. Specifically, a face image (or batch of $m$ face images) as a tensor of shape $(m, n_H, n_W, n_C) = (m, 160, 160, 3)$
# - The input images are originally of shape 96x96, thus, you need to scale them to 160x160. This is done in the `img_to_encoding()` function.
# - The output is a matrix of shape $(m, 128)$ that encodes each input face image into a 128-dimensional vector
#
# Run the cell below to create the model for face images!
# +
from tensorflow.keras.models import model_from_json
json_file = open('keras-facenet-h5/model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
model = model_from_json(loaded_model_json)
model.load_weights('keras-facenet-h5/model.h5')
# -
# Now summarize the input and output shapes:
print(model.inputs)
print(model.outputs)
# By using a 128-neuron fully connected layer as its last layer, the model ensures that the output is an encoding vector of size 128. You then use the encodings to compare two face images as follows:
#
# <img src="images/distance_kiank.png\" style="width:680px;height:250px;">
# <caption><center> <u> <font color='purple'> <b>Figure 2:</b> <br> </u> <font color='purple'>By computing the distance between two encodings and thresholding, you can determine if the two pictures represent the same person</center></caption>
#
# So, an encoding is a good one if:
#
# - The encodings of two images of the same person are quite similar to each other.
# - The encodings of two images of different persons are very different.
#
# The triplet loss function formalizes this, and tries to "push" the encodings of two images of the same person (Anchor and Positive) closer together, while "pulling" the encodings of two images of different persons (Anchor, Negative) further apart.
#
# <img src="images/triplet_comparison.png" style="width:280px;height:150px;"><br>
# <caption><center> <u> <font color='purple'> <b>Figure 3: </b> <br> </u> <font color='purple'> In the next section, you'll call the pictures from left to right: Anchor (A), Positive (P), Negative (N)</center></caption>
# <a name='3-2'></a>
# ### 3.2 - The Triplet Loss
#
# **Important Note**: Since you're using a pretrained model, you won't actually need to implement the triplet loss function in this assignment. *However*, the triplet loss is the main ingredient of the face recognition algorithm, and you'll need to know how to use it for training your own FaceNet model, as well as other types of image similarity problems. Therefore, you'll implement it below, for fun and edification. :)
#
# For an image $x$, its encoding is denoted as $f(x)$, where $f$ is the function computed by the neural network.
#
# <img src="images/f_x.png" style="width:380px;height:150px;">
#
# Training will use triplets of images $(A, P, N)$:
#
# - A is an "Anchor" image--a picture of a person.
# - P is a "Positive" image--a picture of the same person as the Anchor image.
# - N is a "Negative" image--a picture of a different person than the Anchor image.
#
# These triplets are picked from the training dataset. $(A^{(i)}, P^{(i)}, N^{(i)})$ is used here to denote the $i$-th training example.
#
# You'd like to make sure that an image $A^{(i)}$ of an individual is closer to the Positive $P^{(i)}$ than to the Negative image $N^{(i)}$) by at least a margin $\alpha$:
#
# $$
# || f\left(A^{(i)}\right)-f\left(P^{(i)}\right)||_{2}^{2}+\alpha<|| f\left(A^{(i)}\right)-f\left(N^{(i)}\right)||_{2}^{2}
# $$
#
#
# You would thus like to minimize the following "triplet cost":
#
# $$\mathcal{J} = \sum^{m}_{i=1} \large[ \small \underbrace{\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2}_\text{(1)} - \underbrace{\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2}_\text{(2)} + \alpha \large ] \small_+ \tag{3}$$
# Here, the notation "$[z]_+$" is used to denote $max(z,0)$.
#
# **Notes**:
#
# - The term (1) is the squared distance between the anchor "A" and the positive "P" for a given triplet; you want this to be small.
# - The term (2) is the squared distance between the anchor "A" and the negative "N" for a given triplet, you want this to be relatively large. It has a minus sign preceding it because minimizing the negative of the term is the same as maximizing that term.
# - $\alpha$ is called the margin. It's a hyperparameter that you pick manually. You'll use $\alpha = 0.2$.
#
# Most implementations also rescale the encoding vectors to haven L2 norm equal to one (i.e., $\mid \mid f(img)\mid \mid_2$=1); you won't have to worry about that in this assignment.
#
# <a name='ex-1'></a>
# ### Exercise 1 - triplet_loss
#
# Implement the triplet loss as defined by formula (3). These are the 4 steps:
#
# 1. Compute the distance between the encodings of "anchor" and "positive": $\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2$
# 2. Compute the distance between the encodings of "anchor" and "negative": $\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2$
# 3. Compute the formula per training example: $ \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2 - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2 + \alpha$
# 4. Compute the full formula by taking the max with zero and summing over the training examples:$$\mathcal{J} = \sum^{m}_{i=1} \large[ \small \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2 - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2+ \alpha \large ] \small_+ \tag{3}$$
#
# *Hints*:
#
# - Useful functions: `tf.reduce_sum()`, `tf.square()`, `tf.subtract()`, `tf.add()`, `tf.maximum()`.
#
# - For steps 1 and 2, sum over the entries of $\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2$ and $\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2$.
#
# - For step 4, you will sum over the training examples.
#
# *Additional Hints*:
#
# - Recall that the square of the L2 norm is the sum of the squared differences: $||x - y||_{2}^{2} = \sum_{i=1}^{N}(x_{i} - y_{i})^{2}$
#
# - Note that the anchor, positive and negative encodings are of shape (*m*,128), where *m* is the number of training examples and 128 is the number of elements used to encode a single example.
#
# - For steps 1 and 2, maintain the number of *m* training examples and sum along the 128 values of each encoding. `tf.reduce_sum` has an axis parameter. This chooses along which axis the sums are applied.
#
# - Note that one way to choose the last axis in a tensor is to use negative indexing (axis=-1).
#
# - In step 4, when summing over training examples, the result will be a single scalar value.
#
# - For `tf.reduce_sum` to sum across all axes, keep the default value axis=None.
# [**tf.math.reduce_sum**](https://www.tensorflow.org/api_docs/python/tf/math/reduce_sum): Computes the sum of elements across dimensions of a tensor.<br>
#
# [**tf.math.square**](https://www.tensorflow.org/api_docs/python/tf/math/square): Computes square of `x` element-wise.
#
# [**tf.math.subtract**](https://www.tensorflow.org/api_docs/python/tf/math/subtract): Returns `x` - `y` element-wise.
#
# [**tf.math.add**](https://www.tensorflow.org/api_docs/python/tf/math/add): Returns `x` + `y` element-wise.
#
# [**tf.math.maximum**](https://www.tensorflow.org/api_docs/python/tf/math/maximum): Returns the max of x and y (i.e. `x` > `y` ? `x` : `y`) element-wise.
#
# [What's difference between tf.math.subtract and just minus operation in tensorflow?](https://stackoverflow.com/questions/36110834/whats-difference-between-tf-sub-and-just-minus-operation-in-tensorflow)
# + nbgrader={"grade": false, "grade_id": "cell-f05732f7068382cb", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C1(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: triplet_loss
def triplet_loss(y_true, y_pred, alpha = 0.2):
"""
Implementation of the triplet loss as defined by formula (3)
Arguments:
y_true -- true labels, required when you define a loss in Keras, you don't need it in this function.
y_pred -- python list containing three objects:
anchor -- the encodings for the anchor images, of shape (None, 128)
positive -- the encodings for the positive images, of shape (None, 128)
negative -- the encodings for the negative images, of shape (None, 128)
Returns:
loss -- real number, value of the loss
"""
anchor, positive, negative = y_pred[0], y_pred[1], y_pred[2]
### START CODE HERE
# Step 1: Compute the (encoding) distance between the anchor and the positive
pos_dist = tf.math.reduce_sum(
tf.math.square(
tf.math.subtract(anchor, positive)), axis = -1)
# Step 2: Compute the (encoding) distance between the anchor and the negative
neg_dist = tf.math.reduce_sum(
tf.math.square(
tf.math.subtract(anchor, negative)), axis = -1)
# Step 3: subtract the two previous distances and add alpha.
basic_loss = tf.math.add( tf.math.subtract(pos_dist, neg_dist), alpha )
# Step 4: Take the maximum of basic_loss and 0.0. Sum over the training examples.
loss = tf.math.reduce_sum( tf.math.maximum(basic_loss, 0) )
### END CODE HERE
return loss
# + nbgrader={"grade": true, "grade_id": "cell-440ff81e6bcda96a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# BEGIN UNIT TEST
tf.random.set_seed(1)
y_true = (None, None, None) # It is not used
y_pred = (tf.keras.backend.random_normal([3, 128], mean=6, stddev=0.1, seed = 1),
tf.keras.backend.random_normal([3, 128], mean=1, stddev=1, seed = 1),
tf.keras.backend.random_normal([3, 128], mean=3, stddev=4, seed = 1))
loss = triplet_loss(y_true, y_pred)
assert type(loss) == tf.python.framework.ops.EagerTensor, "Use tensorflow functions"
print("loss = " + str(loss))
y_pred_perfect = ([1., 1.], [1., 1.], [1., 1.,])
loss = triplet_loss(y_true, y_pred_perfect, 5)
assert loss == 5, "Wrong value. Did you add the alpha to basic_loss?"
y_pred_perfect = ([1., 1.],[1., 1.], [0., 0.,])
loss = triplet_loss(y_true, y_pred_perfect, 3)
assert loss == 1., "Wrong value. Check that pos_dist = 0 and neg_dist = 2 in this example"
y_pred_perfect = ([1., 1.],[0., 0.], [1., 1.,])
loss = triplet_loss(y_true, y_pred_perfect, 0)
assert loss == 2., "Wrong value. Check that pos_dist = 2 and neg_dist = 0 in this example"
y_pred_perfect = ([0., 0.],[0., 0.], [0., 0.,])
loss = triplet_loss(y_true, y_pred_perfect, -2)
assert loss == 0, "Wrong value. Are you taking the maximum between basic_loss and 0?"
y_pred_perfect = ([[1., 0.], [1., 0.]],[[1., 0.], [1., 0.]], [[0., 1.], [0., 1.]])
loss = triplet_loss(y_true, y_pred_perfect, 3)
assert loss == 2., "Wrong value. Are you applying tf.reduce_sum to get the loss?"
y_pred_perfect = ([[1., 1.], [2., 0.]], [[0., 3.], [1., 1.]], [[1., 0.], [0., 1.,]])
loss = triplet_loss(y_true, y_pred_perfect, 1)
if (loss == 4.):
raise Exception('Perhaps you are not using axis=-1 in reduce_sum?')
assert loss == 5, "Wrong value. Check your implementation"
# END UNIT TEST
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# <b>loss</b>
# </td>
# <td>
# 527.2598
# </td>
# </tr>
# </table>
# <a name='4'></a>
# ## 4 - Loading the Pre-trained Model
#
# FaceNet is trained by minimizing the triplet loss. But since training requires a lot of data and a lot of computation, you won't train it from scratch here. Instead, you'll load a previously trained model in the following cell; which might take a couple of minutes to run.
# + nbgrader={"grade": false, "grade_id": "cell-953bcab8e9bbba10", "locked": true, "schema_version": 3, "solution": false, "task": false}
FRmodel = model
# -
# Here are some examples of distances between the encodings between three individuals:
#
# <img src="images/distance_matrix.png" style="width:380px;height:200px;"><br>
# <caption><center> <u> <font color='purple'> <b>Figure 4:</b></u> <br> <font color='purple'> Example of distance outputs between three individuals' encodings</center></caption>
#
# Now use this model to perform face verification and face recognition!
# <a name='5'></a>
# ## 5 - Applying the Model
#
# You're building a system for an office building where the building manager would like to offer facial recognition to allow the employees to enter the building.
#
# You'd like to build a face verification system that gives access to a list of people. To be admitted, each person has to swipe an identification card at the entrance. The face recognition system then verifies that they are who they claim to be.
#
# <a name='5-1'></a>
# ### 5.1 - Face Verification
#
# Now you'll build a database containing one encoding vector for each person who is allowed to enter the office. To generate the encoding, you'll use `img_to_encoding(image_path, model)`, which runs the forward propagation of the model on the specified image.
#
# Run the following code to build the database (represented as a Python dictionary). This database maps each person's name to a 128-dimensional encoding of their face.
#tf.keras.backend.set_image_data_format('channels_last')
def img_to_encoding(image_path, model):
img = tf.keras.preprocessing.image.load_img(image_path, target_size=(160, 160))
img = np.around(np.array(img) / 255.0, decimals=12)
x_train = np.expand_dims(img, axis=0)
embedding = model.predict_on_batch(x_train)
return embedding / np.linalg.norm(embedding, ord=2)
database = {}
database["danielle"] = img_to_encoding("images/danielle.png", FRmodel)
database["younes"] = img_to_encoding("images/younes.jpg", FRmodel)
database["tian"] = img_to_encoding("images/tian.jpg", FRmodel)
database["andrew"] = img_to_encoding("images/andrew.jpg", FRmodel)
database["kian"] = img_to_encoding("images/kian.jpg", FRmodel)
database["dan"] = img_to_encoding("images/dan.jpg", FRmodel)
database["sebastiano"] = img_to_encoding("images/sebastiano.jpg", FRmodel)
database["bertrand"] = img_to_encoding("images/bertrand.jpg", FRmodel)
database["kevin"] = img_to_encoding("images/kevin.jpg", FRmodel)
database["felix"] = img_to_encoding("images/felix.jpg", FRmodel)
database["benoit"] = img_to_encoding("images/benoit.jpg", FRmodel)
database["arnaud"] = img_to_encoding("images/arnaud.jpg", FRmodel)
# Load the images of Danielle and Kian:
danielle = tf.keras.preprocessing.image.load_img("images/danielle.png", target_size=(160, 160))
kian = tf.keras.preprocessing.image.load_img("images/kian.jpg", target_size=(160, 160))
np.around(np.array(kian) / 255.0, decimals=12).shape
kian
np.around(np.array(danielle) / 255.0, decimals=12).shape
danielle
# Now, when someone shows up at your front door and swipes their ID card (thus giving you their name), you can look up their encoding in the database, and use it to check if the person standing at the front door matches the name on the ID.
#
# <a name='ex-2'></a>
# ### Exercise 2 - verify
#
# Implement the `verify()` function, which checks if the front-door camera picture (`image_path`) is actually the person called "identity". You will have to go through the following steps:
#
# - Compute the encoding of the image from `image_path`.
# - Compute the distance between this encoding and the encoding of the identity image stored in the database.
# - Open the door if the distance is less than 0.7, else do not open it.
#
# As presented above, you should use the L2 distance `np.linalg.norm`.
#
# **Note**: In this implementation, compare the L2 distance, not the square of the L2 distance, to the threshold 0.7.
#
# *Hints*:
#
# - `identity` is a string that is also a key in the database dictionary.
# - `img_to_encoding` has two parameters: the image_path and model.
# + nbgrader={"grade": false, "grade_id": "cell-ba2f317e79e15a2f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C2(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: verify
def verify(image_path, identity, database, model):
"""
Function that verifies if the person on the "image_path" image is "identity".
Arguments:
image_path -- path to an image
identity -- string, name of the person you'd like to verify the identity. Has to be an employee who works in the office.
database -- python dictionary mapping names of allowed people's names (strings) to their encodings (vectors).
model -- your Inception model instance in Keras
Returns:
dist -- distance between the image_path and the image of "identity" in the database.
door_open -- True, if the door should open. False otherwise.
"""
### START CODE HERE
# Step 1: Compute the encoding for the image. Use img_to_encoding() see example above.
encoding = img_to_encoding(image_path, model)
# Step 2: Compute distance with identity's image
dist = np.linalg.norm( (encoding-database[identity]), ord=2, keepdims=False )
# Step 3: Open the door if dist < 0.7, else don't open
if dist < 0.7:
print("It's " + str(identity) + ", welcome in!")
door_open = True
else:
print("It's not " + str(identity) + ", please go away")
door_open = False
### END CODE HERE
return dist, door_open
# -
# Younes is trying to enter the office and the camera takes a picture of him ("images/camera_0.jpg"). Let's run your verification algorithm on this picture:
#
# <img src="images/camera_0.jpg\" style="width:100px;height:100px;">
# + nbgrader={"grade": true, "grade_id": "cell-014d077254ad7d52", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# BEGIN UNIT TEST
assert(np.allclose(verify("images/camera_1.jpg", "bertrand", database, FRmodel), (0.54364836, True)))
assert(np.allclose(verify("images/camera_3.jpg", "bertrand", database, FRmodel), (0.38616243, True)))
assert(np.allclose(verify("images/camera_1.jpg", "younes", database, FRmodel), (1.3963861, False)))
assert(np.allclose(verify("images/camera_3.jpg", "younes", database, FRmodel), (1.3872949, False)))
verify("images/camera_0.jpg", "younes", database, FRmodel)
# END UNIT TEST
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# <b>It's Younes, welcome in!</b>
# </td>
# <td>
# (0.5992946, True)
# </td>
# </tr>
# </table>
# Benoit, who does not work in the office, stole Kian's ID card and tried to enter the office. Naughty Benoit! The camera took a picture of Benoit ("images/camera_2.jpg).
#
# <img src="images/camera_2.jpg" style="width:100px;height:100px;">
#
# Run the verification algorithm to check if Benoit can enter.
verify("images/camera_2.jpg", "kian", database, FRmodel)
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# <b>It's not Kian, please go away</b>
# </td>
# <td>
# (1.0259346, False)
# </td>
# </tr>
# </table>
# <a name='5-2'></a>
# ### 5.2 - Face Recognition
#
# Your face verification system is mostly working. But since Kian got his ID card stolen, when he came back to the office the next day he couldn't get in!
#
# To solve this, you'd like to change your face verification system to a face recognition system. This way, no one has to carry an ID card anymore. An authorized person can just walk up to the building, and the door will unlock for them!
#
# You'll implement a face recognition system that takes as input an image, and figures out if it is one of the authorized persons (and if so, who). Unlike the previous face verification system, you will no longer get a person's name as one of the inputs.
#
# <a name='ex-3'></a>
# ### Exercise 3 - who_is_it
#
# Implement `who_is_it()` with the following steps:
#
# - Compute the target encoding of the image from `image_path`
# - Find the encoding from the database that has smallest distance with the target encoding.
# - Initialize the `min_dist` variable to a large enough number (100). This helps you keep track of the closest encoding to the input's encoding.
# - Loop over the database dictionary's names and encodings. To loop use for (name, db_enc) in `database.items()`.
# - Compute the L2 distance between the target "encoding" and the current "encoding" from the database. If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
# + nbgrader={"grade": false, "grade_id": "cell-a04ff2b5fd1186f8", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C3(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: who_is_it
def who_is_it(image_path, database, model):
"""
Implements face recognition for the office by finding who is the person on the image_path image.
Arguments:
image_path -- path to an image
database -- database containing image encodings along with the name of the person on the image
model -- your Inception model instance in Keras
Returns:
min_dist -- the minimum distance between image_path encoding and the encodings from the database
identity -- string, the name prediction for the person on image_path
"""
### START CODE HERE
## Step 1: Compute the target "encoding" for the image. Use img_to_encoding() see example above.
encoding = img_to_encoding(image_path, model)
## Step 2: Find the closest encoding ##
# Initialize "min_dist" to a large value, say 100.
min_dist = 100
# Loop over the database dictionary's names and encodings.
for (name, db_enc) in database.items():
# Compute L2 distance between the target "encoding" and the current db_enc from the database.
dist = np.linalg.norm( (encoding-db_enc), ord=2, keepdims=False )
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
if dist < min_dist:
min_dist = dist
identity = name
### END CODE HERE
if min_dist > 0.7:
print("Not in the database.")
else:
print ("it's " + str(identity) + ", the distance is " + str(min_dist))
return min_dist, identity
# -
# Younes is at the front door and the camera takes a picture of him ("images/camera_0.jpg"). Let's see if your `who_it_is()` algorithm identifies Younes.
# + nbgrader={"grade": true, "grade_id": "cell-9c88c8ab87677503", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# BEGIN UNIT TEST
# Test 1 with Younes pictures
who_is_it("images/camera_0.jpg", database, FRmodel)
# Test 2 with Younes pictures
test1 = who_is_it("images/camera_0.jpg", database, FRmodel)
assert np.isclose(test1[0], 0.5992946)
assert test1[1] == 'younes'
# Test 3 with Younes pictures
test2 = who_is_it("images/younes.jpg", database, FRmodel)
assert np.isclose(test2[0], 0.0)
assert test2[1] == 'younes'
# END UNIT TEST
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# <b>it's Younes, the distance is 0.5992946</b>
# </td>
# <td>
# (0.5992946, 'younes')
# </td>
# </tr>
# </table>
#
# You can change "camera_0.jpg" (picture of Younes) to "camera_1.jpg" (picture of Bertrand) and see the result.
# **Congratulations**!
# You've completed this assignment, and your face recognition system is working well! It not only lets in authorized persons, but now people don't need to carry an ID card around anymore!
#
# You've now seen how a state-of-the-art face recognition system works, and can describe the difference between face recognition and face verification. Here's a quick recap of what you've accomplished:
#
# - Posed face recognition as a binary classification problem
# - Implemented one-shot learning for a face recognition problem
# - Applied the triplet loss function to learn a network's parameters in the context of face recognition
# - Mapped face images into 128-dimensional encodings using a pretrained model
# - Performed face verification and face recognition with these encodings
#
# Great work!
# <font color='blue'>
#
# **What you should remember**:
#
# - Face verification solves an easier 1:1 matching problem; face recognition addresses a harder 1:K matching problem.
#
# - Triplet loss is an effective loss function for training a neural network to learn an encoding of a face image.
#
# - The same encoding can be used for verification and recognition. Measuring distances between two images' encodings allows you to determine whether they are pictures of the same person.
# **Ways to improve your facial recognition model**:
#
# Although you won't implement these here, here are some ways to further improve the algorithm:
#
# - Put more images of each person (under different lighting conditions, taken on different days, etc.) into the database. Then, given a new image, compare the new face to multiple pictures of the person. This would increase accuracy.
#
# - Crop the images to contain just the face, and less of the "border" region around the face. This preprocessing removes some of the irrelevant pixels around the face, and also makes the algorithm more robust.
# <a name='6'></a>
# ## 6 - References
# 1. <NAME>, <NAME>, <NAME> (2015). [FaceNet: A Unified Embedding for Face Recognition and Clustering](https://arxiv.org/pdf/1503.03832.pdf)
#
# 2. <NAME>, <NAME>, <NAME>, <NAME> (2014). [DeepFace: Closing the gap to human-level performance in face verification](https://research.fb.com/wp-content/uploads/2016/11/deepface-closing-the-gap-to-human-level-performance-in-face-verification.pdf)
#
# 3. This implementation also took a lot of inspiration from the official FaceNet github repository: https://github.com/davidsandberg/facenet
#
# 4. Further inspiration was found here: https://machinelearningmastery.com/how-to-develop-a-face-recognition-system-using-facenet-in-keras-and-an-svm-classifier/
#
# 5. And here: https://github.com/nyoki-mtl/keras-facenet/blob/master/notebook/tf_to_keras.ipynb
| Convolutional Neural Networks/7_Face_Recognition/Face_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:firm_learning]
# language: python
# name: conda-env-firm_learning-py
# ---
# # GMM error has some dimension error
#
# **Fix it!**
# +
import numpy as np
import dill
import pandas as pd
from scipy import optimize as opt
import time
import sys
sys.path.append('../')
import src
#GMM parameters
maxiters = 50 #120. About 2 minutes per iteration
time_periods = 40 #Maximum spell_t to consider
min_periods = 3 #What
#Check if this parameters still make sense for the current product
β10, β11 = -2, 3.5
β20, β21 = 1.3, -2.
betas = [β10, β11, β20, β21]
#Load policy and value function
#####################
file_n = "2018-10-5vfi_dict.dill" #Personal Macbook
#file_n = "2019-2-16vfi_dict.dill" #Work Macbook
with open('../data/' + file_n, 'rb') as file:
data_d = dill.load(file)
lambdas = src.generate_simplex_3dims(n_per_dim=data_d['n_of_lambdas_per_dim'])
price_grid = np.linspace(data_d['min_price'], data_d['max_price'])
policy = data_d['policy']
valueF = data_d['valueF']
lambdas_ext = src.generate_simplex_3dims(n_per_dim=
data_d['n_of_lambdas_per_dim'])
#Interpolate policy (level price). valueF is already a function
policyF = src.interpolate_wguess(lambdas_ext, policy)
#dataframe and standard deviation
cleaned_data = "../../firm_learning/data/cleaned_data/"
df = pd.read_csv(cleaned_data + "medium_prod_for_gmm.csv")
std_devs = (df.groupby('firm').level_prices.rolling(window=4, min=3)
.std().reset_index()
.rename(columns={'level_1': 't', 'level_prices': 'std_dev_prices'}))
df = pd.merge(df, std_devs, on=['firm', 't'], how='left')
mean_std_observed_prices = df.groupby('t').std_dev_prices.mean()[min_periods:]
xs = df.groupby('firm').xs.first().values
Nfirms = len(xs)
# Just add a zeroes. Makes sense for the gmm estimation
prior_shocks = src.gen_prior_shocks(Nfirms, σerror=0)
# +
from src import from_theta_to_lambda_for_all_firms
θ = [0.1, 2.1, -1, -2.1]
xs_stand = np.abs(0.2*(xs - np.mean(xs))/ (np.std(xs)))
print(np.mean(xs_stand), np.std(xs_stand))
lambdas0 = from_theta_to_lambda_for_all_firms(θ, xs_stand, prior_shocks)
lambdas0[12:18]
# -
mean_std_observed_prices_cl.index
# +
# Fit t to observed_prices
#mean_std_expected_prices
mean_std_observed_prices_cl = mean_std_observed_prices[pd.notnull(mean_std_observed_prices)]
mean_std_expected_prices_cl = mean_std_expected_prices[pd.notnull(mean_std_expected_prices)]
index_inters = np.intersect1d(mean_std_observed_prices_cl.index,
mean_std_expected_prices_cl.index)
mean_std_observed_prices_cl = mean_std_observed_prices_cl.loc[index_inters]
# -
mean_std_observed_prices_cl.head(10)
mean_std_observed_prices.head(10)
# +
w = None
t = len(mean_std_expected_prices)
if w is None:
w = np.identity(t)
g = (1 / t) * (mean_std_expected_prices - mean_std_observed_prices[0:76])[:, np.newaxis]
(g.T @ w @ g)[0, 0]
# +
mean_std_expected_prices = generate_mean_std_pricing_decisions(df, policyF,
lambdas0, min_periods)
try:
assert len(mean_std_observed_prices) == len(mean_std_expected_prices)
except AssertionError as e:
e.args += (len(mean_std_observed_prices), len(mean_std_expected_prices))
raise
t = len(mean_std_expected_prices)
if w is None:
w = np.identity(t)
g = (1 / t) * (mean_std_expected_prices - mean_std_observed_prices)[:, np.newaxis]
return (g.T @ w @ g)[0, 0]
src.gmm_error = gmm_error
# +
def generate_mean_std_pricing_decisions(df, policyF, lambdas_at_0, min_periods=3):
"""
Lambdas0: starting priors for each of the N firms
"""
pricing_decision_dfs = []
for i, firm in enumerate(df.firm.unique()):
prices = src.generate_pricing_decisions(policyF, lambdas_at_0[i],
df[df.firm == firm].log_dmd.values)
pricing_decision_dfs.append(pd.DataFrame({'level_prices': prices,
'firm': np.repeat(firm, len(prices))
}))
pricing_decision_df = pd.concat(pricing_decision_dfs, axis=0)
std_dev_df = (pricing_decision_df.groupby('firm').level_prices.rolling(window=4,
min=min_periods)
.std().reset_index()
.rename(columns={'level_1': 't',
'level_prices': 'std_dev_prices'}))
return std_dev_df.groupby('t').std_dev_prices.mean()[min_periods:]
mean_std_expected_prices = generate_mean_std_pricing_decisions(df, policyF,
lambdas0, min_periods)
# -
len(mean_std_observed_prices), len(mean_std_expected_prices)
mean_std_observed_prices.head(10)
mean_std_observed_prices.tail(10)
# +
# Optimization
######################
maxiters = 2 #120. About 2 minutes per iteration
def error_w_data(θ) -> float:
return src.gmm_error(θ, policyF, xs,
mean_std_observed_prices=mean_std_observed_prices, df=df,
prior_shocks=prior_shocks, min_periods=min_periods)
optimi = opt.differential_evolution(error_w_data, [(-2.5, 0.5), (2.0, 4.0),
(0.5, 2), (-3., 1.)],
maxiter=maxiters)
# -
| Notebooks/debug_gmm_error.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ' Zipline environment'
# language: python
# name: zipline
# ---
# <img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
#
# © Copyright Quantopian Inc.<br>
# © Modifications Copyright QuantRocket LLC<br>
# Licensed under the [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/legalcode).
#
# <a href="https://www.quantrocket.com/disclaimer/">Disclaimer</a>
# # Why You Should Hedge Beta and Sector Exposures
#
# by <NAME> and <NAME>
#
# Whenever we have a trading strategy of any sort, we need to be considering the impact of systematic risk. There needs to be some risk involved in a strategy in order for there to be a return above the risk-free rate, but systematic risk poisons the well, so to speak. By its nature, systematic risk provides a commonality between the many securities in the market that cannot be diversified away. As such, we need to construct a hedge to get rid of it.
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.covariance import LedoitWolf
import seaborn as sns
import statsmodels.api as sm
# # The Fundamental Law of Asset Management
#
# The primary driver of the value of any strategy is whether or not it provides a compelling risk-adjusted return, i.e., the Sharpe Ratio. As expressed in "The Fundamental Law of Active Management", by <NAME>, Sharpe Ratio can be decomposed into two components, skill and breadth, as:
#
# $$IR = IC \sqrt{BR}$$
#
# Technically, this is the definition of the Information Ratio (IR), but for our purposes it is equivalent to the Sharpe Ratio. The IR is the ratio of the excess return of a portfolio over its benchmark per unit active risk, i.e., the excess return of a long-only portfolio less its benchmark per unit tracking error. In the time of Grinold’s publication, however, long/short investing was a rarity. Today, in the world of hedge funds and long/short investing, there is no benchmark. We seek absolute returns so, in this case, the IR is equivalent to the Sharpe ratio.
#
# In this equation, skill is measured by IC (Information Coefficient), calculated with Alphalens. The IC is essentially the Spearman rank correlation, used to correlate your prediction and its realization. Breadth is measured as the number of **independent** bets in the period. The takeaway from this "law" is that, with any strategy, we need to:
#
# 1. Bet well (high IC),
# 2. Bet often (high number of bets), *and*
# 3. **Make independent bets**
#
# If the bets are completely independent, then breadth is the total number of bets we have made for every individual asset, the number of assets times the number of periods. If the bets are not independent then the **effective breadth** can be much much less than the number of assets. Let's see precisely what beta exposure and sector exposure do to **effective breadth**.
# <div class="alert alert-warning">
# <b>TL;DR:</b> Beta exposure and sector exposure lead to a significant increase in correlation among bets. Portfolios with beta and sector bets have very low effective breadth. In order to have high Sharpe then, these portfolios must have very high IC. It is easier to increase effective breadth by hedging beta and sector exposure than it is to increase your IC.
# </div>
# # Forecasts and Bet Correlation
#
# We define a bet as the forecast of the *residual* of a security return. This forecast can be implicit -- i.e., we buy a stock and thus implicity we forecast that the stock will go up. What though do we mean by *residual*? Without any fancy math, this simply means the return **less a hedge**. Let's work through three examples. We use the Ledoit-Wolf covariance estimator to assess our covariance in all cases. For more information on why we use Ledoit-Wolf instead of typical sample covariance, check out the Estimating Covariance Matrices lecture.
#
# ### Example 1: No Hedge!
#
# If we go long on a set of securities, but do not hold any short positions, there is no hedge! So the *residual* is the stock return itself.
#
# $$r_{resid,i} = r_i$$
#
# Let's see what the correlation of our bets are in this case.
# + jupyter={"outputs_hidden": false}
from quantrocket.master import get_securities
from quantrocket import get_prices
tickers = ['WFC', 'JPM', 'USB', 'XOM', 'BHI', 'SLB'] # The securities we want to go long on
securities = get_securities(symbols=tickers, vendors='usstock')
# Obtain prices
historical_prices = get_prices(
'usstock-1d-bundle',
data_frequency='daily',
sids=securities.index.tolist(),
start_date='2015-01-01',
end_date='2017-02-22',
fields='Close')
sids_to_symbols = securities.Symbol.to_dict()
historical_prices = historical_prices.rename(columns=sids_to_symbols)
rets = historical_prices.loc['Close'].pct_change().fillna(0) # Calculate returns
lw_cov = LedoitWolf().fit(rets).covariance_ # Calculate Ledoit-Wolf estimator
def extract_corr_from_cov(cov_matrix):
# Linear algebra result:
# https://math.stackexchange.com/questions/186959/correlation-matrix-from-covariance-matrix
d = np.linalg.inv(np.diag(np.sqrt(np.diag(cov_matrix))))
corr = d.dot(cov_matrix).dot(d)
return corr
# + jupyter={"outputs_hidden": false}
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
# Plot prices
left = historical_prices.loc['Close'].plot(ax=ax1)
# Plot covariance as a heat map
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=tickers, yticklabels=tickers)
# + jupyter={"outputs_hidden": false}
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print('Average pairwise correlation: %.4f' % average_corr)
# -
# The result here is that we have six bets and they are all very highly correlated.
# ### Example 2: Beta Hedge
#
# In this case, we will assume that each bet is hedged against the market (SPY). In this case, the residual is calculated as:
#
# $$ r_{resid,i} = r_i - \beta_i r_i $$
#
# where $\beta_i$ is the beta to the market of security $i$ calculated with the CAPM and $r_i$ is the return of security $i$.
# + jupyter={"outputs_hidden": false}
tickers = ['WFC', 'JPM', 'USB', 'XOM', 'BHI', 'SLB', 'SPY'] # The securities we want to go long on
securities = get_securities(symbols=tickers, vendors='usstock')
# Obtain prices
historical_prices = get_prices(
'usstock-1d-bundle',
data_frequency='daily',
sids=securities.index.tolist(),
start_date='2015-01-01',
end_date='2017-02-22',
fields='Close')
sids_to_symbols = securities.Symbol.to_dict()
historical_prices = historical_prices.rename(columns=sids_to_symbols)
rets = historical_prices.loc['Close'].pct_change().fillna(0) # Calculate returns
market = rets['SPY']
stock_rets = rets.drop('SPY', axis=1)
residuals = stock_rets.copy()*0
for stock in stock_rets.columns:
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
residuals[stock] = results.resid
lw_cov = LedoitWolf().fit(residuals).covariance_ # Calculate Ledoit-Wolf Estimator
# + jupyter={"outputs_hidden": false}
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
left = (1+residuals).cumprod().plot(ax=ax1)
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=stock_rets.columns, yticklabels=stock_rets.columns)
# + jupyter={"outputs_hidden": false}
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print('Average pairwise correlation: %.4f' % average_corr)
# -
# The beta hedge has brought down the average correlation significanty. Theoretically, this should improve our breadth. It is obvious that we are left with two highly correlated clusters however. Let's see what happens when we hedge the sector risk.
# ### Example 3: Sector Hedge
#
# The sector return and the market return are themselves highly correlated. As such, you cannot do a multivariate regression due to multicollinearity, a classic violation of regression assumptions (see the lecture "Violations of Regression Models"). To hedge against both the market and a given security's sector, you first estimate the market beta residuals and then calculate the sector beta on *those* residuals.
#
# $$
# r_{resid,i} = r_i - \beta_i r_i \\
# r_{resid_{SECTOR},i}= r_{resid,i} - \beta_{SECTOR,i}r_{resid,i}
# $$
#
# Here, $r_{resid, i}$ is the residual between the security return and a market beta hedge and $r_{resid_{SECTOR}, i}$ is the residual between *that* residual and a hedge of that residual against the relevant sector.
# + jupyter={"outputs_hidden": false}
tickers = ['WFC', 'JPM', 'USB', 'XLF', 'SPY', 'XOM', 'BHI', 'SLB', 'XLE']
securities = get_securities(symbols=tickers, vendors='usstock')
# Obtain prices
historical_prices = get_prices(
'usstock-1d-bundle',
data_frequency='daily',
sids=securities.index.tolist(),
start_date='2015-01-01',
end_date='2017-02-22',
fields='Close')
sids_to_symbols = securities.Symbol.to_dict()
historical_prices = historical_prices.rename(columns=sids_to_symbols)
rets = historical_prices.loc['Close'].pct_change().fillna(0) # Calculate returns
# Get market hedge ticker
mkt = 'SPY'
# Get sector hedge tickers
sector_1_hedge = 'XLF'
sector_2_hedge = 'XLE'
# Identify securities for each sector
sector_1_stocks = ['WFC', 'JPM', 'USB']
sector_2_stocks = ['XOM', 'BHI', 'SLB']
market_rets = rets[mkt]
sector_1_rets = rets[sector_1_hedge]
sector_2_rets = rets[sector_2_hedge]
stock_rets = rets.drop(['XLF', 'SPY', 'XLE'], axis=1)
residuals_market = stock_rets.copy()*0
residuals = stock_rets.copy()*0
# Calculate market beta of sector 1 benchmark
model = sm.OLS(sector_1_rets.values, market.values)
results = model.fit()
sector_1_excess = results.resid
# Calculate market beta of sector 2 benchmark
model = sm.OLS(sector_2_rets.values, market.values)
results = model.fit()
sector_2_excess = results.resid
for stock in sector_1_stocks:
# Calculate market betas for sector 1 stocks
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
# Calculate residual of security + market hedge
residuals_market[stock] = results.resid
# Calculate sector beta for previous residuals
model = sm.OLS(residuals_market[stock], sector_1_excess)
results = model.fit()
# Get final residual
residuals[stock] = results.resid
for stock in sector_2_stocks:
# Calculate market betas for sector 2 stocks
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
# Calculate residual of security + market hedge
residuals_market[stock] = results.resid
# Calculate sector beta for previous residuals
model = sm.OLS(residuals_market[stock], sector_2_excess)
results = model.fit()
# Get final residual
residuals[stock] = results.resid
# Get covariance of residuals
lw_cov = LedoitWolf().fit(residuals).covariance_
# + jupyter={"outputs_hidden": false}
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
left = (1+residuals).cumprod().plot(ax=ax1)
labels = sector_1_stocks + sector_2_stocks
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=labels, yticklabels=labels)
# + jupyter={"outputs_hidden": false}
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print('Average pairwise correlation: %.4f' % average_corr)
# -
# The sector hedge further brought down the correlation between our bets.
# ## Calculating Effective Breadth
#
# This section is based on "How to calculate breadth: An evolution of the fundamental law of active portfolio management", by <NAME>; Vol. 4, 6, 393-405, 2003, _Journal of Asset Management_. Buckle derives the "semi-generalised fundamental law of active management" under several weak assumptions. The key result of this paper (for us) is a closed-form calculaiton of effective breadth as a function of the correlation between bets. Buckle shows that breadth, $BR$, can be modeled as
#
# $$BR = \frac{N}{1 + \rho(N -1)}$$
#
# where N is the number of stocks in the portfolio and $\rho$ is the assumed single correlation of the expected variation around the forecast.
# + jupyter={"outputs_hidden": false}
def buckle_BR_const(N, rho):
return N/(1 + rho*(N - 1))
corr = np.linspace(start=0, stop=1.0, num=500)
plt.plot(corr, buckle_BR_const(6, corr))
plt.title('Effective Breadth as a function of Forecast Correlation (6 Stocks)')
plt.ylabel('Effective Breadth (Number of Bets)')
plt.xlabel('Forecast Correlation');
# -
# Here we see that in the case of the long-only portfolio, where the average correlation is 0.56, we are *effectively making only approximately 2 bets*. When we hedge beta, with a resulting average correlation of 0.22, things get a little better, *three effective bets*. When we add the sector hedge, we get close to zero correlation, and in this case the number of bets equals the number of assets, 6.
#
# **More independent bets with the same IC leads to higher Sharpe ratio.**
# ## Using this in Practice
#
# Trading costs money due to market impact and commissions. As such, the post hoc implementation of a hedge is almost always suboptimal. In that case, you are trading purely to hedge risk. It is preferable to think about your sector and market exposure *throughout the model development process*. Sector and market risk is naturally hedged in a pairs-style strategy; in a cross-sectional strategy, consider de-meaning the alpha vector by the sector average; with an event-driven strategy, consider adding additional alphas so you can find offsetting bets in the same sector. As a last resort, hedge with a well chosen sector ETF.
#
# ---
#
# **Next Lecture:** [VaR and CVaR](Lecture40-VaR-and-CVaR.ipynb)
#
# [Back to Introduction](Introduction.ipynb)
# ---
#
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian") or QuantRocket LLC ("QuantRocket"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, neither Quantopian nor QuantRocket has taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information believed to be reliable at the time of publication. Neither Quantopian nor QuantRocket makes any guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| quant_finance_lectures/Lecture39-Why-Hedge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Validation
# In the validation step the trained model and pipeline was tested on data that the model has not seen before. The metric used to examine the results were ROC, accuracy, and f1 score. Also, recall and precision for one and zero was examined.
# ### Import pyspark using Docker
import pyspark
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
import matplotlib.pyplot as plt
import numpy as np
from pyspark.ml.classification import LogisticRegression,LogisticRegressionModel
from pyspark.ml.evaluation import BinaryClassificationEvaluator
import warnings
warnings.filterwarnings("ignore")
# ### Start Spark Session
spark = SparkSession.builder.appName('val').getOrCreate()
# ### Load Data
df = spark.read.csv('clean_val/part-00000-2661d739-2781-4738-9b1b-6b4c69096d9d-c000.csv', header = True).select('Text', 'verified')
### View data
df.show(10)
#### look for nan values
print('Null Text:', df.where((df["Text"].isNull())).count())
print('Null verified:', df.where((df["verified"].isNull())).count())
### drop na's
df = df.na.drop()
df.count()
### create a Label column
df = df.withColumn('label', when(df.verified == 'true', 1.0).otherwise(0.0)).select('Text', 'label')
df.show(10)
# ### Load Pipeline & Model
# Pipeline and trained model were imported in to be tested on the validation data.
### Import pipeline
from pyspark.ml import PipelineModel, Pipeline
load_pipline = PipelineModel.read().load('pipline_train')
### import model
model = LogisticRegressionModel.load('LGmodel')
# ### Transform validation data
val = load_pipline.transform(df)
val.show(10)
# ### Predict with validation data
pred = model.transform(val)
pred.select('label', 'prediction', 'probability').show(10)
# ### Metrics
# Metric used were ROC, accuracy, and f1 score. All three metric showed results that were better than the results on the training set.
#### R0C
evaluator = BinaryClassificationEvaluator()
print('Test Area Under ROC', evaluator.evaluate(pred))
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
#### Accuracy
acc = MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label', metricName='accuracy')
print('Accuracy:', acc.evaluate(pred))
#### F1 Score
ff = MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label', metricName='f1')
print('F1 score:', ff.evaluate(pred))
# ### Recall , Precision, F1 score
# When looking at the recall and precision for both zero and one the recall for zero is not great. The model only gets about 0.25 of the actual zero(false) right.
import pandas as pd
from sklearn import metrics as skmetrics
y_true = pred.select(['label']).collect()
y_pred = pred.select(['prediction']).collect()
#### Classification Report
from sklearn.metrics import classification_report
print(classification_report(y_true, y_pred))
| Notebooks/Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Task 1
# ##### Python to take a list as input and to return a dictionary of unique items in the list as keys and the number of times each item appears as values
# ##### I am using my latest golf scorecard as a list of results on each of 18 holes
import numpy as np
# +
def Counts(my_golf_scorecard):
# We have to create an empty dictionary first
count = {}
for i in ['Bogey', 'Bogey','Par','Par', 'Q_Bogey', 'T_Bogey', 'D_Bogey', 'Par', 'Bogey', 'Par', 'Birdie', 'Q_Bogey', 'D_Bogey', 'D_Bogey', 'Par', 'D_Bogey','Par', 'T_Bogey'] :
count[i] = count.get(i, 0) + 1 # using an in-built Python get() function
return count
# Driver function
if __name__ == "__main__":
my_golf_scorecard = ['Bogey', 'Bogey','Par','Par', 'Q_Bogey', 'T_Bogey', 'D_Bogey', 'Par', 'Bogey', 'Par', 'Birdie', 'Q_Bogey', 'D_Bogey', 'D_Bogey', 'Par', 'D_Bogey','Par', 'T_Bogey']
print(Counts(my_golf_scorecard))
# + active=""
#
# References:
#
# Driver function https://www.codegrepper.com/code-examples/delphi/if+__name__%3D%3D+__main__+in+python
# <br><br>
# https://www.geeksforgeeks.org/what-does-the-if-__name__-__main__-do/#:~:text=Python%20files%20can%20act%20as,run%20directly%2C%20and%20not%20imported.
#
#
# -
# ## Task 2
# ##### Write a Python function called dicerolls that simulates rolling dice. The function shoud simulate randomly rolling k dice n times, keeping track of each total face value. It should then return a dictionary with the number of times each possible total face value occured.
# ***
# import necessary libraries
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import random
# +
# number of dice k = 2, times rolled n = 1000
# we need to create a function that will allow us to simulate rolling 2 dice in the loop for 1000 times.
# Then we are to append results to a list.
def dicerolls(n):
roll_results = []
while n < 1001:
dice_1 = np.random.randint(1,6)
dice_2 = np.random.randint(1,6)
dice_total += dice_1 + dice_2
roll_results.append(dice_total)
return roll_results
# -
# #### Converting our array into a list
# Reference: https://www.journaldev.com/32797/python-convert-numpy-array-to-list
dice_total_list = dice_total.tolist()
print(f'Dice_total: {dice_total_list}')
# creating an empty list that will store the 2nd value - amount of times each sum occured
list2 = []
two = dice_total_list.count(2)
three = dice_total_list.count(3)
four = dice_total_list.count(4)
five = dice_total_list.count(5)
six = dice_total_list.count(6)
seven = dice_total_list.count(7)
eight = dice_total_list.count(8)
nine = dice_total_list.count(9)
ten = dice_total_list.count(10)
eleven = dice_total_list.count(11)
twelve = dice_total_list.count(12)
# creating the 1st list that will store the 1st value - all the possible sums of 2 dice
list1 = [2,3,4,5,6,7,8,9,10,11,12]
# +
# using .extend() function to add values to list2
# https://stackoverflow.com/questions/20196159/how-to-append-multiple-values-to-a-list-in-python#:~:text=extend%20to%20extend%20the%20list,provides%20a%20sequence%20of%20values.&text=So%20you%20can%20use%20list,()%20to%20append%20multiple%20values.
list2.extend((two,three,four,five,six,seven,eight,nine,ten,eleven,twelve))
# -
list2
# + active=""
# We combine data from list1 and list2 into one single dictionary
#
# # https://careerkarma.com/blog/python-convert-list-to-dictionary/#:~:text=Converting%20a%20list%20to%20a,the%20Python%20zip()%20function.
# # https://stackoverflow.com/questions/209840/convert-two-lists-into-a-dictionary
# -
dictionary = {list1[i]: list2[i] for i in range(len(list1))}
# dictionary
# <i>Conclusion: we were not able to write the working code, starting again differently</i>
# *****
# *****
# <b>Write a Python function called dicerolls that simulates rolling dice. The function should simulate randomly rolling k dice n times, keeping track of each total face value. It should then return a dictionary with the number of times each possible total face value occured.</b>
# Import necessary libraries
import numpy as np
import random
# First we create a multidimentional array of possible scenarios of dice rolls.<br>
# It will have 1000 rows by 2 columns. Each column will have a randomly generated number between 1 and 6.
# +
dicerolls = np.random.randint(1, 7, size=(1000, 2))
print(dicerolls)
# Reference - Create a random multidimensional array of random integers - https://pynative.com/python-random-randrange/
# +
# Counting sum of two dice together for each of the 1000 rolls
y = np.sum(dicerolls,axis=1)
# Reference - https://numpy.org/doc/stable/reference/generated/numpy.sum.html
# -
y
# +
# Let us look at the data in Pandas.
import pandas as pd
df = pd.value_counts(y)
# Reference - Using Pandas - https://stackoverflow.com/questions/10741346/numpy-most-efficient-frequency-counts-for-unique-values-in-an-array
# -
# Creating DataSeries
df
# +
# Creating DataFrame
df2 = pd.DataFrame(df).reset_index()
df2.columns = ['Dice sum', 'Count']
# Reference DataFrame from DFataSeries, naming columns- https://stackoverflow.com/questions/28503445/assigning-column-names-to-a-pandas-series
# -
df2
# Sorting dataframe by 1st column
dfsort = df2.sort_values(by=['Dice sum']).reset_index()
dfsort
# Let us import matplotlib to do some visualization
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Plotting line plot with matplotlib
dfsort.plot(kind='line',x='Dice sum',y='Count', color='red')
plt.ylabel('Count',color='red',fontsize=16)
plt.xlabel('Sum of two dice',color='red',fontsize=16)
plt.title('Dice sum frequency',fontsize=18,color='red')
plt.show()
# Reference - Pandas dataframe plot examples with matplotlib pyplot
# https://queirozf.com/entries/pandas-dataframe-plot-examples-with-matplotlib-pyplot
# -
# Let us look at the same data on a <b>bar plot</b>
# Plotting bar plot with matplotlib
fig = plt.figure(figsize = (15, 5))
dice_sum = dfsort['Dice sum']
count = dfsort['Count']
plt.bar(dice_sum,count)
# changing frequency of xticks -https://www.kite.com/python/answers/how-to-change-the-frequency-of-ticks-in-a-matplotlib-figure-in-python
x_ticks = np.arange(2, 13, 1)
plt.xticks(x_ticks)
plt.ylabel('Count',color='red',fontsize=16)
plt.xlabel('Sum of two dice',color='red',fontsize=16)
plt.title('Dice sum frequency',fontsize=18,color='red')
plt.show()
# <b>Visualization conclusion</b><br>
# As we can see, the shape of the plot sort of resembles bell-shaped curve, where number 7 is the most popular as it has most number of possible combinations (1+6, 2+5, 3+4, 4+3, 5+2 and 6+1) whereas 2 and 12 are least popular as they have minimum amount of combinations (1+1 and 6+6 respectively).
np.sum(dicerolls,axis=1)
# Now let us finally create a dictionary that will have a particular sum of two dice as keys and amount of times that that sum occured out of 1000 rolls - as values
# importing Counter colection
from collections import Counter
dict = Counter(y)
dict
# ### Task 2 conclusion
# With the help of Numpy random package we created a multidimentional array that simulated rolling two dice of possible value from 1 to 6 each 1000 times.<br>
# Then we transformed our data into a sum of two dice - a DataSeries of 1000 values, simulating 1000 random rolls.<br>
# Then we transformed our DataSeries into a DataFrame and presented our data on a line plot and a bar plot. After that visual representation we saw the clear bell-shaped pattern of frequency of appearance.<br>
# Finally, using Counter collection, we transformed our array into a dictionary with a sum of two dice as keys and amount of times that that sum occured out of 1000 rolls as values.
# ## Task 3
# <b>Write some python code that simulates flipping a coin 100 times. Then run this code 1,000 times, keeping track of the number of heads in each of the 1,000 simulations. Select an appropriate plot to depict the resulting list of 1,000 numbers, showing that it roughly follows a bell-shaped curve. You should explain your work in a Markdown cell above the code.</b>
# <i>Before we start writing actual code, let us do some plotting using binomial() function from the numpy.random package</i>
# Random.binomial() function draws samples from a binomial distribution. It describes outcomes of binary scenarios, so, toss of a coin that will always be heads or tails, will be ideal.
# It has 3 parameters: number of trials, probability of occurence in each one, shape of the returned array.
#
# They must meet the following three criteria:<br>
# 1) The number of observations or trials is fixed. <br>
# 2) Each observation or trial is independent, that is, none of your trials have an effect on the probability of the next trial.<br>
# 3) The probability of success is exactly the same from one trial to another.<br>
# In our case, we meet all 3 parameters ideally.
#
# Reference:
# <i>Random.binomial() function https://www.statisticshowto.com/probability-and-statistics/binomial-theorem/binomial-distribution-formula/#whatis Binomial distribution https://www.w3schools.com/python/numpy_random_binomial.asp</i>
#
# +
# This is a way of simulating a probability test of getting heads in a coin toss
import numpy as np
from numpy import random
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# test - probability of getting heads in a coin toss, tossed 100 times.
# Initializing the parameters 'number of trials' and 'probability of success'
# 1st parameter - number of trials - 100
# 2nd parameter - probability of success - 1/2 = 0.5
# test run 1000 times
bindist = np.random.binomial(100,0.5,1000)
plt.hist(bindist);
# -
# Let us plot the same parameters on the line
bindist = sns.distplot(random.binomial(n=100, p=0.5, size=1000), hist=False)
# As we can see, the resulting curve is very close to the bell-shaped curve indeed.
# *****
# <b>Now let us try and write a code that would simulate flipping a coin chosen amount of time.</b>
# +
import numpy as np # importing numpy module
import random # Importing Numpy's random package
def coin_toss(flips): #Creating a function with an argument "number of flips"
# There are 2 possible scenarios - heads or tails.
heads = 0 # we only need to keep track of number of heads. Creating a running total variable
for i in range(flips): # using for loop to iterate through the range
rand = random.randint(0,1) # generating a variable equal to one of the two equally possible random outcomes.
# 0 is heads and 1 is tails.
if rand == 0: # if outcome of rand is 0 - we increment to the total value of heads
heads += 1
return(heads) # return the result of the total number of heads
# References: Python tutorial: calculating a running total - https://www.youtube.com/watch?v=bkpG5jmPXs4
# Using a loop to keep the running total https://www.youtube.com/watch?v=prNzO_vtPvA
# https://whiscardz.wordpress.com/2015/10/05/python-keep-running-total-in-a-for-loop/
# -
# adding argument to our function - amount of coin flips is 100.
coin_toss(100)
# <b>Now we need to come up with a function that will run this simulation 1000 times</b>
# One of the ways to do it is to create an empty list, iterate our range (1000), run our previously created function <span style="color:blue">coin_toss</span> and append its results to our list.<br>
#
# <i>Reference: Converting printed output of function to a list<br>
# https://stackoverflow.com/questions/35932579/converting-printed-output-of-function-to-a-list </i>
# +
list = []
for i in range(1000):
list.append(coin_toss(100))
# -
list
# Now that we have a list we need to do the counting and save it into a dictionary.<br>
# Dictionary key - amount of heads out of 100 flips.<br>
# Dictionary value - amount of occurences of that particular amount in 1000 runs.
# +
# Counter() function does the counting for us, converting the list into a dictionary
from collections import Counter
list_dict = Counter(list) # list_dict is our dictionary now
# https://stackoverflow.com/questions/2600191/how-can-i-count-the-occurrences-of-a-list-item
# How can I count the occurrences of a list item?
# -
list_dict
# <b>Select an appropriate plot to depict the resulting list of 1,000 numbers, showing that it roughly follows a bell-shaped curve. </b>
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Plotting a <b> bar plot</b> first
# +
plt.bar(*zip(*list_dict.items()),color='green')
plt.xlabel('Number of heads out of 100 flips',fontsize=13)
plt.ylabel('Tests with this result (1000 runs)',fontsize=13)
plt.title('Simulation of 1000 runs',fontsize=18,color='red')
plt.show()
# Plot a bar using matplotlib using a dictionary
# https://stackoverflow.com/questions/16010869/plot-a-bar-using-matplotlib-using-a-dictionary
# https://www.listendata.com/2019/06/matplotlib-tutorial-learn-plot-python.html
# https://www.geeksforgeeks.org/bar-plot-in-matplotlib/
# -
# Plotting a curved line plot.
# +
import matplotlib.pylab as plt
lists = sorted(list_dict.items()) # sorted by key, return a list of tuples
x, y = zip(*lists) # unpack a list of pairs into two tuples
plt.xlabel('Number of heads out of 100 flips',fontsize=13)
plt.ylabel('Tests with this result (1000 runs)',fontsize=13)
plt.title('Simulation of 1000 runs',fontsize=18,color='blue')
plt.plot(x, y)
plt.show()
# Plotting a python dict in order of key values
# https://stackoverflow.com/questions/37266341/plotting-a-python-dict-in-order-of-key-values/37266356
# -
# <b>Conclusion:</b><br>
# <i> So, we have created a code that simulates random coin flip 100 times. Then, we added a code that is able to run that simulation 1000 times. Finally, following that simulation, we plotted the resulting list of 1000 values on a bar plot and line plot.<br>
# We came to the conclusion that the shape of the plots does indeed follow the bell curve very closely, with 47 - 53 values being the top of the curve as most frequent values and the values less than 47 and more than 53 have progressively smaller amount of occurence. It is worth mentioned, though, that in most runs 50 value has lower occurence than neighbouring 49 and 51, so then our bell-curve has a characteristic dip in the middle.</i>
# *****
# ## Task 4
# ##### Simpson’s paradox is a well-known statistical paradox where a trend evident in a number of groups reverses when the groups are combined into one big data set. Use numpy to create four data sets, each with an x array and a corresponding y array, to demonstrate Simpson’s paradox. You might create your x arrays using numpy.linspace and create the y array for each x using notation like y = a * x + b where you choose the a and b for each x , y pair to demonstrate the paradox. You might see the Wikipedia page for Simpson’s paradox for inspiration.
#
# Simpson's paradox, which also goes by several other names, is a phenomenon in probability and statistics, in which a trend appears in several different groups of data but disappears or reverses when these groups are combined. This result is often encountered in social-science and medical-science statistics and is particularly problematic when frequency data is unduly given causal interpretations. The paradox can be resolved when causal relations are appropriately addressed in the statistical modeling. It is also referred to as Simpson's reversal, Yule–Simpson effect, amalgamation paradox, or reversal paradox.
#
# #### Example
#
# ##### Batting averages
#
# A common example of Simpson's paradox involves the batting averages of players in professional baseball. It is possible for one player to have a higher batting average than another player each year for a number of years, but to have a lower batting average across all of those years. This phenomenon can occur when there are large differences in the number of at bats between the years. Mathematician <NAME> demonstrated this using the batting average of two baseball players, <NAME> and <NAME>, during the years 1995 and 1996
#
# 
#
# In both 1995 and 1996, Justice had a higher batting average (in bold type) than Jeter did. However, when the two baseball seasons are combined, Jeter shows a higher batting average than Justice.
#
# <i> Reference: https://en.wikipedia.org/wiki/Simpson%27s_paradox </i>
# *****
# ##### Creating datasets
# We are going to analyze seasonal statyistics of 2 basketball players from "Lakeville Racoons" from Midwestern Basketball League (MWBA) - <NAME> and <NAME>. To be more precise, we will analyze their average scoring (average points per game ) during the two consecutive seasons -1991/92 and 1992/93. After that we will analyze their statistics as total of these two seasons.
# importing libraries
import numpy as np
import random
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Season 1991/92
# ##### <NAME> 1991/92
# Creating DataFrame for <NAME> for season 1991/92
# He played 73 games that season.
np.random.seed(35)
games_kj_92 = np.linspace(1,73,73)
games_kj_92
# Creating scoring data using numpy.triangular() function.
# Scoring stats:
# Worst game - 10 points where he only managed to score 3 times out of 14 from the game (three-point shots 0/4)
# and only scored 4 penalty shots out of 7.
# Maximum score - 37 points where they played at home the worst team in the league and outscored their opponents by 42 points.
# Average per game for the season - 23
points_kj_92 = np.random.triangular(10, 23, 34,73).round(0)
points_kj_92
# Total points scored this season
points_kj_92.sum().round(0)
# Creating DataFrame with generated numbers
johnson1992 = pd.DataFrame({'Game number':games_kj_92,'Points':points_kj_92})
johnson1992
# +
# Creating last rows with seasonal totals stats - 'Total games' and 'Total points'
# -
total_games_kj_92 = games_kj_92.max()
total_points_kj_92 = points_kj_92.sum()
# +
johnson1992.loc[73] = ['Total games','Total points']
johnson1992.loc[74] = [total_games_kj_92,total_points_kj_92]
# https://stackoverflow.com/questions/46621712/add-a-new-row-to-a-pandas-dataframe-with-specific-index-name
# -
johnson1992
# #### <NAME> 1991/92
# Creating data for Brent Simpson season 1991/92
# Due to back injury he only played 13 games that season.
np.random.seed(44)
games_bs_92 = np.linspace(1,13,13)
games_bs_92
# Creating scoring data using numpy.triangular() function.
points_bs_92 = np.random.triangular(11, 23,33,13).round(0)
points_bs_92
# Total points scored this season
points_bs_92.sum().round(0)
# Creating DataFrame with generated numbers
simpson1992 = pd.DataFrame({'Game number':games_bs_92,'Points':points_bs_92})
simpson1992
# +
# Creating last rows with seasonal totals stats - 'Total games' and 'Total points'
# -
total_games_bs_92 = games_bs_92.max()
total_points_bs_92 = points_bs_92.sum()
simpson1992.loc[13] = ['Total games','Total points']
simpson1992.loc[14] = [total_games_bs_92,total_points_bs_92]
simpson1992
# ### Season 1992/93
# #### <NAME> 1992/93
# Creating DataFrame for <NAME> for season 1991/92
# Ka'whim missed 7 games due to the grievance and 60 games due to the left achilles injury and only played 8 games out of 75 possible.
np.random.seed(30)
games_kj_93 = np.linspace(1,8,8)
games_kj_93
# Creating scoring data using numpy.triangular() function - defining worst game, best games and average scoring.
points_kj_93 = np.random.triangular(9, 18, 24,8).round(0)
points_kj_93
# Total points scored this season
points_kj_93.sum().round(0)
# Creating DataFrame with generated numbers
johnson1993 = pd.DataFrame({'Game number':games_kj_93,'Points':points_kj_93})
johnson1993
# +
# Creating last rows with seasonal totals stats - 'Total games' and 'Total points'
# -
total_games_kj_93 = games_kj_93.max()
total_points_kj_93 = points_kj_93.sum()
johnson1993.loc[8] = ['Total games','Total points']
johnson1993.loc[9] = [total_games_kj_93,total_points_kj_93]
johnson1993
# #### <NAME> 1992/93
# Creating data for <NAME> season 1992/93
# Still recovering from the injury the injury mid-season and having missed first few months of the season, Brent played 53 games.
np.random.seed(39)
games_bs_93 = np.linspace(1,53,53)
games_bs_93
# Creating scoring data using numpy.triangular() function (defining borders for worst game and best game as well as average)
points_bs_93 = np.random.triangular(9,18,26,53).round(0)
points_bs_93
# Total points scored this season
points_bs_93.sum().round(0)
# Creating DataFrame with generated numbers
simpson1993 = pd.DataFrame({'Game number':games_bs_93,'Points':points_bs_93})
simpson1993
# +
# Creating last rows with seasonal totals stats - 'Total games' and 'Total points'
# -
total_games_bs_93 = games_bs_93.max()
total_points_bs_93 = points_bs_93.sum()
simpson1993.loc[53] = ['Total games','Total points']
simpson1993.loc[54] = [total_games_bs_93,total_points_bs_93]
simpson1993
# So, we now have 4 separate datasets - <NAME> 1991/92, <NAME> 1992/93, <NAME> 1991/92 and <NAME> 1992/93. Each of them has 2 columns - game number and points scored. The last row of each dataset has total amount of games played in particular season and total amount of points scored. <br>
# ##### Creating combined dataset
# Now we will create a separate dataset that will have data for both players and both seasons.<br>
# The data that we are particularly interested in is <b>average scoring record per season</b>.<br>
# Let us pull these data from our datasets again.
# ##### <i><NAME> 1991/92 average scoring record.</i>
# Extracting necessary row from dataset
ave_data_kj_1992 = johnson1992.iloc[74]
ave_data_kj_1992
# averave scoring record calculated
johnson_ave_1992 = ave_data_kj_1992[1] / ave_data_kj_1992[0]
johnson_ave_1992
# ##### <i><NAME> 1992/93 average scoring record.</i>
# Extracting necessary row from dataset
ave_data_kj_1993 = johnson1993.iloc[9]
ave_data_kj_1993
# averave scoring record calculated
johnson_ave_1993 = ave_data_kj_1993[1] / ave_data_kj_1993[0]
johnson_ave_1993
# ##### <i><NAME> 1991/92 average scoring record.</i>
# Extracting necessary row from dataset
ave_data_bs_1992 = simpson1992.iloc[14]
ave_data_bs_1992
# averave scoring record calculated
simpson_ave_1992 = ave_data_bs_1992[1] / ave_data_bs_1992[0]
simpson_ave_1992
# ##### <i><NAME> 1992/93 average scoring record.</i>
# Extracting necessary row from dataset
ave_data_bs_1993 = simpson1993.iloc[54]
ave_data_bs_1993
# averave scoring record calculated
simpson_ave_1993 = ave_data_bs_1993[1] / ave_data_bs_1993[0]
simpson_ave_1993
# ### Combining data from two seasons together
# ##### <i><NAME> combined seasons 1991/92 and 1992/93 average scoring record.</i>
# Total games
Total_games_kj = ave_data_kj_1992[0] + ave_data_kj_1993[0]
Total_games_kj
# Total points
Total_points_kj = ave_data_kj_1992[1] + ave_data_kj_1993[1]
Total_points_kj
# Average scoring record
johnson_total_ave = Total_points_kj / Total_games_kj
johnson_total_ave
# ##### <i><NAME> combined seasons 1991/92 and 1992/93 average scoring record.</i>
# Total games
Total_games_bs = ave_data_bs_1992[0] + ave_data_bs_1993[0]
Total_games_bs
# Total points
Total_points_bs = ave_data_bs_1992[1] + ave_data_bs_1993[1]
Total_points_bs
# Average scoring record
simpson_total_ave = Total_points_bs / Total_games_bs
simpson_total_ave
# ##### Creating DataSet with combined data
d = {'Season':['1991/92','1992/93','Total'],
'<NAME>':[johnson_ave_1992,johnson_ave_1993,johnson_total_ave],
'<NAME>':[simpson_ave_1992,simpson_ave_1993,simpson_total_ave]}
df = pd.DataFrame(data=d)
# +
# styler function
df.style.highlight_max(axis=1)
# <i> Reference: https://pandas.pydata.org/docs/user_guide/style.html#Building-styles </i>
# -
df.round(1)
# Now that we have the data we tried to get so desperately for the last few days, let us have a close look at them.
# <NAME> is the club's top scorer for both seasons (average points scored per game), beating <NAME> on both occasions with a small margin. However, if we calculate the same piece of statistics for both seasons together, we see that <NAME> actually beats his team-mate with a comfortable margin of almost 3 points! Here we witness so called Simpson's Paradox.
# Right was "Lakeville Racoons" head coach <NAME> who used to say that <NAME> was a player full of paradoxes, who could have a splendid game after a wild night out and only few our of sleep , but next time would have a dreadful game after full week's rest and proper training.
# ### Conclusion
# Occurence of Simpson's Paradox shown us importance of analyzing the data carefully and looking for causes. In our case, the paradox occured because of the fact that there was a big discrepancy in number of games played in each season between both players. In season 1991/92 <NAME> played 73 games but <NAME> only 13.<br>
# In season 1992/93 <NAME> played only 8 games but <NAME> 53. So, when we were comparing there 1-season stats, we were not really "comparing like with like", which created the paradox.
# ##### Disclamer
# <i> All characters depicted in the work are fully fictional and never existed. The work is created fully and solely for the educational purpose. Equally, the basketball team as well as the basketball league are totally and fully fictional and were created for the same purpose.</i>
| Tasks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Richter's Predictor: Modeling Earthquake Damage
#
# Based on aspects of building location and construction, the goal is to predict the level of damage to buildings caused by the 2015 Gorkha earthquake in Nepal.
#
# The data was collected through surveys by [Kathmandu Living Labs](http://www.kathmandulivinglabs.org) and the [Central Bureau of Statistics](https://cbs.gov.np), which works under the National Planning Commission Secretariat of Nepal. This survey is **one of the largest post-disaster datasets ever collected**, containing valuable information on earthquake impacts, household conditions, and socio-economic-demographic statistics.
# +
import pandas as pd
import numpy as np
import os
import re
import string
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
# -
# # ! pip install imbalanced-learn
import imblearn
# print(imblearn.__version__)
from imblearn.over_sampling import SMOTE
# +
import sklearn
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV, train_test_split, StratifiedKFold, cross_val_score
from sklearn.pipeline import Pipeline
from sklearn import metrics
from sklearn.metrics import classification_report, f1_score, roc_auc_score, roc_curve, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_selection import RFECV, SelectFromModel
import multiprocessing
# -
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', None)
pd.set_option('display.max_colwidth', None)
# Load data set
train = pd.read_csv(os.path.join('', 'Richters_Predictor_Modeling_Earthquake_Damage_-_Train_Values.csv'))
test = pd.read_csv(os.path.join('', 'Richters_Predictor_Modeling_Earthquake_Damage_-_Test_Values.csv'))
labels = pd.read_csv(os.path.join('', 'Richters_Predictor_Modeling_Earthquake_Damage_-_Train_Labels.csv'))
labels.damage_grade.value_counts()/len(labels.damage_grade)
# **Remarks:**
#
# - According to these results, we can say that there are 56.89% of the building suffered a medium amount of damage, 33.46% of the building were almost completely distructed and 9.64% of the building suffered a low damage.
#
# - Class "low damage" is imbalanced, and might need upsampling. Another option to deal with the imbalance is to choose an appropriate metric, like F1 score or AUC.
train.dtypes.value_counts()
print('Object data types:\n')
#we'll use the function later, without wanting to print anything
def get_obj(train, p = False):
obj_types = []
for column in train.columns:
if train[column].dtype == 'object':
if p: print(column)
obj_types.append(column)
return obj_types
obj_types = get_obj(train, True)
def transform_to_int(train, obj_types):
#Assign dictionaries with current values and replacements for each column
d_lsc = {'n':0, 'o':1, 't':2}
d_ft = {'h':0, 'i':1, 'r':2, 'u':3, 'w':4}
d_rt = {'n':0, 'q':1, 'x':2}
d_gft = {'f':0, 'm':1, 'v':2, 'x':3, 'z':4}
d_oft = {'j':0, 'q':1, 's':2, 'x':3}
d_pos = {'j':0, 'o':1, 's':2, 't':3}
d_pc = {'a':0, 'c':1, 'd':2, 'f':3, 'm':4, 'n':5, 'o':6, 'q':7, 's':8, 'u':9}
d_los = {'a':0, 'r':1, 'v':2, 'w':3}
#Each positional index in replacements corresponds to the column in obj_types
replacements = [d_lsc, d_ft, d_rt, d_gft, d_oft, d_pos, d_pc, d_los]
#Replace using lambda Series.map(lambda)
for i,col in enumerate(obj_types):
train[col] = train[col].map(lambda a: replacements[i][a]).astype('int64')
transform_to_int(train, obj_types)
train.dtypes.value_counts()
train.head()
# ***
# **Data Splitting**
# ***
y = labels.pop('damage_grade')
x = train.drop(["building_id"],axis=1)
print('Original dataset shape:', x.shape)
print('Original labelset shape:', y.shape)
# keep the same random state for reproducibility
RANDOM_STATE = 12
TRAIN_TEST_SPLIT_SIZE = .1
# stratify on damage_grade
x_train, x_test, y_train, y_test = train_test_split(x, y,
test_size = TRAIN_TEST_SPLIT_SIZE,
stratify = y,
random_state = RANDOM_STATE)
print('Training dataset shape:', x_train.shape)
print('Training labelset shape:', y_train.shape)
print('Test labelset shape:', y_test.shape)
print('Test dataset shape:', x_test.shape)
# Scaling is typically done to normalize data so that priority is not given to a particular feature. The role of scaling is mostly important in algorithms that are distance based and require Euclidean Distance. Random Forest is a tree-based model and hence does not require feature scaling.
# ***
# **Feature selection by feature importance of random forest classifier**
# ***
import joblib
# load, no need to initialize the loaded model
clf = joblib.load("./rf.joblib")
sel = SelectFromModel(clf)
sel.fit(x_train, y_train)
sel.get_support()
x_train.columns
features = x_train.columns[sel.get_support()]
features
np.mean(sel.estimator_.feature_importances_)
sel.estimator_.feature_importances_
# ***
# **Recursive Feature Elimination (RFE)**
# ***
def run_randomForest(x_train, x_test, y_train, y_test, clf_rf):
# clf = RandomForestClassifier(n_estimators=600, random_state=0, n_jobs=1)
clf_rf.fit(x_train, y_train)
y_pred = clf_rf.predict(x_test)
score = f1_score(y_test, y_pred, average='micro')
print(f'{score:.4f}')
from sklearn.feature_selection import RFE
sel = RFE(clf, n_features_to_select = 10)
sel.fit(x_train, y_train)
sel.get_support()
features = x_train.columns[sel.get_support()]
features
x_train_rfe = sel.transform(x_train)
x_test_rfe = sel.transform(x_test)
# %%time
run_randomForest(x_train_rfe, x_test_rfe, y_train, y_test, clf)
# +
# importance_rf = pd.DataFrame({"Features":x.columns, "Importance_RF":sel.feature_importances_}).sort_values(by='Importance_RF', ascending = False).head(15)
# RF_styler = importance_rf.style.set_table_attributes("style='display:inline'").set_caption('Top 15 Random Forest importance')
# from IPython.display import display_html
# display_html(RF_styler._repr_html_(), raw=True)
# -
import joblib
# save
joblib.dump(clf, "./rf_rfe.joblib")
# load, no need to initialize the loaded model
loaded_rf = joblib.load("./rf_rfe.joblib")
transform_to_int(test, obj_types)
test = test.drop(["building_id"],axis=1)
predictions = clf.predict(test)
submission_format = pd.read_csv('Richters_Predictor_Modeling_Earthquake_Damage_-_Submission_Format.csv', index_col='building_id')
my_submission = pd.DataFrame(data=predictions,
columns=submission_format.columns,
index=submission_format.index)
my_submission.head()
my_submission.to_csv('submission.csv')
# !head submission.csv
| Random_Forest_Experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Research Question 1
# ## How has COVID-19 affected the cancer waiting times landscape?
#
# This week we will start to try and determine the following:
# > 1. Clean up the data in order to be able to compare different 'stuff'. - Sofie
#
# > 2. The standard of those who meet the standard is relatively unaffected. [Hypothesis testing] - Xell
#
# > 3. There has been a drop in the number of referals. [Hypothesis Testing vs Changepoint] - Chris
#
# > 4. As well as this there has been a change in diagnostic rates due to COVID-19 in line with the change in the number of referals. [High-dimensional Changepoint vs GLM] - Xell/Sofie/Chris
# ### Groupings of Regions
# 1. By HB
# 2. By NOSCAN, WOSCAN, SCAN
# 3. All of Scotland
# ### Groupings of Cancers
# 1. Try completely seperate.
# 2. If data issues group by similar features.
# ### Grouping by Times
# Monthly - try to translate any data into this format where necessary.
# ----------------------------------------
#
# ### Ideas for approaching 1.
#
# Code for 1.
# ----------------------------------------
#
# ### Ideas for approaching 2.
# Code for 2.
# ----------------------------------------
#
# ### Ideas for approaching 3.
# Code for 3.
# ----------------------------------------
#
# ### Ideas for approaching 4.
# Code for 4.
| Week 4 - Research Question 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quem fez o ENEM 2016 apenas para treino
# ___
# Importando as bibliotecas necessárias
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
# +
# Carregando os datasets
treino = pd.read_csv('train.csv')
teste = pd.read_csv('test.csv')
# Selecionando algumas colunas para retirar
retira_colunas = ['NU_INSCRICAO',
# 'IN_TREINEIRO',
# 'CO_PROVA_CH',
# 'CO_PROVA_LC',
# 'CO_PROVA_MT'
]
inscricao_treino = treino[['NU_INSCRICAO']]
inscricao_teste = teste[['NU_INSCRICAO']]
treineiro = treino[['IN_TREINEIRO']]
treino.drop(retira_colunas, axis=1, inplace=True)
teste.drop(retira_colunas, axis=1, inplace=True)
treino.drop('IN_TREINEIRO', axis=1, inplace=True)
# Mostrando o datasets depois da retirada das colunas
treino
# +
# Selecionando as variáveis numéricas do dataset de teste
numeric_features = teste.select_dtypes(include="number").columns.to_list()
# Selecioando as variáveis categóricas do dataset de teste
categoric_feature = [coluna for coluna in teste.columns if coluna not in numeric_features]
# +
# Juntando todas as variáveis escolhidas
all_features = numeric_features + categoric_feature
# Aplicando as colunas escolhidas no dataset de treino
treino = treino[all_features]
treino
# -
# Vendo a quantidade de NaN's nos dados de treino
treino.isna().sum()#.sum()
# Vendo a quantidade de NaN's nos dados de teste
teste.isna().sum()#.sum()
teste['NU_NOTA_LC'].isna().sum()
# +
# Fazendo as trocas dos dados faltantes
for categorica in categoric_feature:
treino[categorica].fillna(method='ffill', inplace=True)
teste[categorica].fillna(method='ffill', inplace=True)
for categorica in categoric_feature:
treino[categorica].fillna(method='bfill', inplace=True)
teste[categorica].fillna(method='bfill', inplace=True)
for numerica in numeric_features:
#ratings_petz_apple[f'{num_stars}-star_diff'].mask(cond=ratings_petz_apple[f'{num_stars}-star_diff'] < 0, other=0)
media_treino = treino[numerica].mean()
treino[numerica].fillna(0, inplace=True)
media_teste = teste[numerica].mean()
teste[numerica].fillna(0, inplace=True)
# -
# Conferindo se deu tudo certo com os dados de treino
treino.isna().sum().sum()
# Conferindo se deu tudo certo com os dados de teste
teste.isna().sum().sum()
# Instanciando um objeto da classe para o StandardScalar
std_scaler = StandardScaler()
# Aplicando o StandardScalar nas variáveis numéricas
treino[numeric_features] = std_scaler.fit_transform(treino[numeric_features])
teste[numeric_features] = std_scaler.fit_transform(teste[numeric_features])
# Conferindo os dados de treino
treino
# +
# Aplicando o get_dummies() nas variáveis categóricas
encoded_columns = pd.get_dummies(treino[categoric_feature])
treino = treino.join(encoded_columns).drop(categoric_feature, axis=1)
encoded_columns = pd.get_dummies(teste[categoric_feature])
teste = teste.join(encoded_columns).drop(categoric_feature, axis=1)
#ohe.fit(treino[categoric_feature])
#ohe.fit(teste[categoric_feature])
# -
# Instanciando um objeto da classe LogisticRegression()
model = LogisticRegression()
treino = treino.join(treineiro)
# Separando X e y de treino
X_train = treino.drop('IN_TREINEIRO', axis=1)
y_train = treino['IN_TREINEIRO']
# + tags=[]
# Treinando o modelo
model.fit(X_train, y_train)
# Testando o modelo
y_test = model.predict(teste)
# -
# Preparando a saída
y_test = pd.DataFrame(y_test)
y_test.rename(columns={0: 'IN_TREINEIRO'}, inplace=True)
y_test
# Juntando o número de inscrição aos dados preditos pelo modelo
my_answer = inscricao_teste.join(y_test)
my_answer
my_answer.to_csv('answer.csv', index=False)
| enem-4/Quem_e_treineiro_no_ENEM_2016.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # KDD Cup 1999 Data
# http://kdd.ics.uci.edu/databases/kddcup99/kddcup99.html
import sklearn
import pandas as pd
from sklearn import preprocessing
from sklearn.utils import resample
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
import numpy as np
from sklearn.decomposition import PCA
from sklearn.neural_network import MLPClassifier
from sklearn.pipeline import Pipeline
import time
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.externals import joblib
from sklearn.utils import resample
print('The scikit-learn version is {}.'.format(sklearn.__version__))
col_names = ["duration","protocol_type","service","flag","src_bytes",
"dst_bytes","land","wrong_fragment","urgent","hot","num_failed_logins",
"logged_in","num_compromised","root_shell","su_attempted","num_root","num_file_creations",
"num_shells","num_access_files","num_outbound_cmds","is_host_login","is_guest_login","count",
"srv_count","serror_rate","srv_serror_rate","rerror_rate","srv_rerror_rate","same_srv_rate",
"diff_srv_rate","srv_diff_host_rate","dst_host_count","dst_host_srv_count",
"dst_host_same_srv_rate","dst_host_diff_srv_rate","dst_host_same_src_port_rate",
"dst_host_srv_diff_host_rate","dst_host_serror_rate","dst_host_srv_serror_rate",
"dst_host_rerror_rate","dst_host_srv_rerror_rate","label"]
data = pd.read_csv("data/corrected", header=None, names = col_names)
data.shape
# # 前処理
# ## カテゴリ化
data.label.value_counts()
data['label2'] = data.label.where(data.label.str.contains('normal'),'atack')
data.label2.value_counts()
data['label3'] = data.label.copy()
data.loc[data.label.str.contains('back|land|neptune|pod|smurf|teardrop|mailbomb|apache2|processtable|udpstorm'),'label3'] = 'DoS'
data.loc[data.label.str.contains('buffer_overflow|loadmodule|perl|rootkit|ps|xterm|sqlattack'),'label3'] = 'U2R'
data.loc[data.label.str.contains('ftp_write|guess_passwd|imap|multihop|phf|spy|warezclient|warezmaster|snmpgetattack|snmpguess|httptunnel|sendmail|named|xlock|xsnoop|worm'),'label3'] = 'R2L'
data.loc[data.label.str.contains('ipsweep|nmap|portsweep|satan|mscan|saint'),'label3'] = 'Probe'
data.label3.value_counts()
# +
#joblib.dump(data,'dump/20171118/corrected.pkl')
# -
# ## サンプリング
# +
#data = resample(data,n_samples=10000,random_state=0)
# +
#data.shape
# -
# ## 数値化
le_protocol_type = preprocessing.LabelEncoder()
le_protocol_type.fit(data.protocol_type)
data.protocol_type=le_protocol_type.transform(data.protocol_type)
le_service = preprocessing.LabelEncoder()
le_service.fit(data.service)
data.service = le_service.transform(data.service)
le_flag = preprocessing.LabelEncoder()
le_flag.fit(data.flag)
data.flag = le_flag.transform(data.flag)
data.describe()
data.shape
# ## ラベルの分離
y_test_1 = data.label.copy()
y_test_2 = data.label2.copy()
y_test_3 = data.label3.copy()
x_test= data.drop(['label','label2','label3'],axis=1)
x_test.shape
y_test_1.shape
y_test_2.shape
y_test_3.shape
# ## 標準化
ss = preprocessing.StandardScaler()
ss.fit(x_test)
x_test = ss.transform(x_test)
col_names2 = ["duration","protocol_type","service","flag","src_bytes",
"dst_bytes","land","wrong_fragment","urgent","hot","num_failed_logins",
"logged_in","num_compromised","root_shell","su_attempted","num_root","num_file_creations",
"num_shells","num_access_files","num_outbound_cmds","is_host_login","is_guest_login","count",
"srv_count","serror_rate","srv_serror_rate","rerror_rate","srv_rerror_rate","same_srv_rate",
"diff_srv_rate","srv_diff_host_rate","dst_host_count","dst_host_srv_count",
"dst_host_same_srv_rate","dst_host_diff_srv_rate","dst_host_same_src_port_rate",
"dst_host_srv_diff_host_rate","dst_host_serror_rate","dst_host_srv_serror_rate",
"dst_host_rerror_rate","dst_host_srv_rerror_rate"]
pd.DataFrame(x_test,columns=col_names2).describe()
# ## 学習
clf = joblib.load('dump/20171118/MLPClassifier10per.pkl')
t1=time.perf_counter()
pred = clf.predict(x_test)
t2=time.perf_counter()
print(t2-t1,"秒")
print(classification_report(y_test_3, pred))
print(confusion_matrix(y_test_3, pred))
# +
#joblib.dump(data,'dump/20171118/MLPClassifier10per.pkl')
# -
| KDDCUP99_18.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np #this library use for frame
import dlib #this library use for detection of face
from tkinter import *
import sys
import datetime #This Library gives current date and time
import time
from PIL import Image, ImageTk
cap = cv2.VideoCapture(0) #this is taking video
detector = dlib.get_frontal_face_detector() #return the face detector / uses for face detection
root = Tk()
#root.geometry("975x585")
root.attributes("-fullscreen", True)
root.title("Face Detection In Real Time")
#def save():
#print ("YOUR DATA IS SAVE")
###------------ FRAME --------###
f1 = Frame(root, bg = "black", borderwidth = 1 , relief = GROOVE)
f1.pack(side = TOP, fill="x")
f2 = Frame(root, bg = "black", borderwidth = 5 , relief = GROOVE)
f2.pack(side = BOTTOM, fill="x")
### FUNCTION ###
def tick():
time_string = time.strftime("%H:%M:%S")
clock.config(text=time_string)
fd()
clock.after(200, tick)
## USE FOR VIDEO FRAME AND CAPTURE#######
def fd():
font=cv2.FONT_HERSHEY_DUPLEX
font_color=(0,255,0)
thickness=2
org=(15,70)
line_type=cv2.LINE_AA
ret,frame=cap.read() #ret give true or false by checking video camera is working or not and frame read video and saved in it.
frame = cv2.flip(frame,1) #here in parameter 1 means flip frame in horizontally and if pass 0 then frame flip in vertically.
gray=cv2.cvtColor(frame,cv2.COLOR_BGR2GRAY) #it will change the color of video from colored to gray.
faces=detector(gray) #this is for detection of gray scale live video.
face_counter=0 #Face Counter
for face in faces: #This loop used for making rectangle around faces one by one.
x,y=face.left(),face.top()
w,h=face.right(),face.bottom()
face_counter+=1 #Face Counter
cv2.rectangle(frame,(x,y),(w,h),(0,225,0),3) #by this Making Rectangular shape around faces.
cv2.putText( frame ,"Detected Faces : "+str(face_counter) , org , font, 1 , font_color , thickness , line_type ) #by Puting Face counter value on real time photage
#For Saving Data in notepad file,I used Filing method.
if(face_counter>0): #Condition
location= open("Data.txt","a") #Data Location
date_now=datetime.datetime.now() #Its Gives Real time
st_datenow=(str(date_now)).replace(" ","\t")
location.write(("\n"+str(st_datenow)+"\t\t"+str(face_counter)+"\n")) #Writing data in notepad file.
location.close() #closing of filing
im1 = Image.fromarray(frame)
photo_root = ImageTk.PhotoImage(im1)
img_root.config(image = photo_root)
img_root.image = photo_root
img_root = Label(root, text = "Live Streaming" , font = ("Arial",30,"bold"))
img_root.pack()
## VIDEO detection
f3 = Frame(root, bg = "black", borderwidth = 1 , relief = GROOVE )
f3.pack(side = TOP, fill="y")
## HEADER ##
l2 = Label(f1, text = " FACE DETECTON ",bg = "black" , fg = "white" , font = ("Arial",30,"bold"))
l2.pack()
## FOOTER ##
l3 = Label(f2, text = "Teachers: <NAME>, <NAME>, <NAME> ", bg = "black", fg="white" , font = ("Arial",10,"bold") )
l3.pack(side=LEFT)
l3 = Label(f2, text = "Members: <NAME>, <NAME>, <NAME>, <NAME>, <NAME> ", bg = "black", fg="white" , font = ("Arial",10,"bold") )
l3.pack(side=RIGHT)
clock=Label(f3, font=("times", 10, "bold"), fg="green", bg="silver")
clock.pack(anchor=S,side=BOTTOM )
## BUTTONS ##
#B3 = Button(root, text ="DATA SAVE", bg = "Black", fg="white", height = 2, width = 10 ,font = ("Arial",10,"bold"), command=save )
#B3.pack(side=RIGHT, anchor="sw", padx=20, pady=20)
B2 = Button(root, text ="CLOSE", bg = "Black", fg="white", height = 2, width = 10 ,font = ("Arial",10,"bold"), command=root.destroy )
B2.pack(side=RIGHT, anchor="sw", padx=20, pady=20)
B1 = Button(root, text ="START", bg = "Black", fg="white", height = 2, width = 10,font = ("Arial",10,"bold"), command=tick )
B1.pack(side=RIGHT, anchor="sw", padx=0, pady=20)
root.mainloop()
cap.release()
# +
#In this code work on OpenCV, numpy and datetime library done by <NAME> and work on dlib done by <NAME>
import cv2
import numpy as np #this library use for frame
import dlib #this library use for detection of face
import datetime #This Library gives current date and time
import sys
cap = cv2.VideoCapture(0) #this is taking video
detector = dlib.get_frontal_face_detector() #return the face detector / uses for face detection
font=cv2.FONT_HERSHEY_DUPLEX
font_color=(0,255,0)
thickness=2
org=(15,70)
line_type=cv2.LINE_AA
while True: #this loop is use to make video contineously until press "s"
ret,frame=cap.read() #ret give true or false by checking video camera is working or not and frame read video and saved in it.
frame = cv2.flip(frame,1) #here in parameter 1 means flip frame in horizontally and if pass 0 then frame flip in vertically.
gray=cv2.cvtColor(frame,cv2.COLOR_BGR2GRAY) #it will change the color of video from colored to gray.
faces=detector(gray)#this is for detection of gray scale live video.
face_counter=0 #Face Counter
for face in faces: #This loop used for making rectangle around faces one by one.
x,y=face.left(),face.top()
w,h=face.right(),face.bottom()
face_counter+=1 #Face Counter
cv2.rectangle(frame,(x,y),(w,h),(0,225,0),3) #by this Making Rectangular shape around faces.
cv2.putText( frame ,"Number of faces : "+str(face_counter) , org , font, 1 , font_color , thickness , line_type ) #by Puting Face counter value on real time photage
cv2.imshow("Face Dectection with real time",frame) #This is showing or displying the video.
#For Saving Data in notepad file,I used Filing method.
if(face_counter>0): #Condition
location= open("Data.txt","a") #Data Location
date_now=datetime.datetime.now() #Its Gives Real time
st_datenow=(str(date_now)).replace(" ","\t")
location.write(("\n"+str(st_datenow)+"\t\t"+str(face_counter)+"\n")) #Writing data in notepad file.
location.close() #closing of filing
if (cv2.waitKey(1) == ord("a")):
break
cap.release() #This is used to realease video or show video
cv2.destroyAllWindows() #this commond will destroy all videos when user press cancel button.
# -
location=open("Data.txt","r")
print(location.read())
| final project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unsupervised Analysis of Days of Week
#
# Treating crossings each day as features to learn about the relationships between various days
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.decomposition import PCA
from sklearn.mixture import GaussianMixture
# plot parameters
FIGSIZE = (12,7)
plt.rcParams['figure.figsize'] = FIGSIZE
plt.style.use('seaborn')
# -
# ## Get Data
from jupyterworkflow.data import get_fremont_data
data = get_fremont_data()
pivoted = data.pivot_table('Total', index=data.index.time, columns=data.index.date)
pivoted.iloc[:,:500].plot(legend=False, alpha=.05, figsize=FIGSIZE);
# ## Principal Component Analysis
X = pivoted.fillna(0).T.values
X.shape
X2 = PCA(2,svd_solver='full').fit_transform(X)
X2.shape
plt.scatter(X2[:,0], X2[:,1]);
# ## Unsupervised Clustering
gmm = GaussianMixture(2)
gmm.fit(X)
labels = gmm.predict(X)
plt.scatter(X2[:,0], X2[:,1], c=labels, cmap='rainbow')
plt.colorbar();
# +
fig, ax = plt.subplots(1, 2, figsize=(15,8))
pivoted.T[labels == 0].T.plot(legend=False, alpha=0.1, ax=ax[0])
pivoted.T[labels == 1].T.plot(legend=False, alpha=0.1, ax=ax[1])
ax[0].set_title("Purple Cluster")
ax[0].set_title("Red Cluster");
# -
# ## Comparing with Day of the week
dayofweek = pd.DatetimeIndex(pivoted.columns).dayofweek
plt.scatter(X2[:,0], X2[:,1], c=dayofweek, cmap='rainbow')
plt.colorbar();
# ## Analyzing outliers
#
# The following points are weekdays with a holiday-like pattern
dates = pd.DatetimeIndex(pivoted.columns)
dates[(labels == 0) & (dayofweek <5)]
| UnsupervisedAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Excercise: use autoencoder (LSTM) for anomaly detection in accelerometer based vibration dataset.
#
# #### The autoencoder tries to reconstruct the input at the output. Hence, for healthy data, it tries to reconstruct healthy data. But it will have a hard time trying to reconstruct faulty data in its neural network bottleneck (LSTM). That's how the anomaly detector works.
# #### Download healthy and faulty dataframes from IBM cloud
# +
# # !pip install tensorflow==2.5.0
# +
# In order to obtain the correct values for "credentias", "bucket_name" and "endpoint"
# please follow the tutorial at https://github.com/IBM/skillsnetwork/wiki/Cloud-Object-Storage-Setup
credentials = {
"apikey": "<KEY>",
"cos_hmac_keys": {
"access_key_id": "<KEY>",
"secret_access_key": "<KEY>"
},
"endpoints": "https://control.cloud-object-storage.cloud.ibm.com/v2/endpoints",
"iam_apikey_description": "Auto-generated for key <KEY>",
"iam_apikey_name": "Service credentials-1",
"iam_role_crn": "crn:v1:bluemix:public:iam::::serviceRole:Writer",
"iam_serviceid_crn": "crn:v1:bluemix:public:iam-identity::a/3dba62a148ab4574867f8eb140c3a44e::serviceid:ServiceId-109769b1-d4d5-4997-93a1-faefc036bfa9",
"resource_instance_id": "crn:v1:bluemix:public:cloud-object-storage:global:a/3dba62a148ab4574867f8eb140c3a44e:643e3143-6265-453a-877a-15ae3947ef9a::"
}
bucket_name = "cloud-object-storage-appliedaideeplearning"
endpoint = "https://s3.eu-de.cloud-object-storage.appdomain.cloud"
# +
import base64
from ibm_botocore.client import Config
import ibm_boto3
import time
# Create client
client = ibm_boto3.client(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url=endpoint
)
client.download_file(bucket_name,'result_healthy_pandas.csv', 'result_healthy_pandas.csv')
client.download_file(bucket_name,'result_faulty_pandas.csv', 'result_faulty_pandas.csv')
# +
import pandas as pd
df_healthy = pd.read_csv('result_healthy_pandas.csv', engine='python', header=None)
print(df_healthy.shape)
df_healthy.head()
# -
df_healthy.loc[df_healthy[1] == 100]
df_faulty = pd.read_csv('result_faulty_pandas.csv', engine='python', header=None)
print(df_faulty.shape)
df_faulty.head()
# #### Import necessary libraries
# +
import numpy as np
from numpy import concatenate
from matplotlib import pyplot
from pandas import read_csv
from pandas import DataFrame
from pandas import concat
import sklearn
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_squared_error
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras.callbacks import Callback
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import LSTM, Dense, Activation
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import time
# %matplotlib inline
# -
# #### Plot data samples
def get_recording(df, file_id):
return np.array(df.sort_values(by=0, ascending=True).loc[df[1] == file_id].drop(0,1).drop(1,1))
# +
healthy_sample = get_recording(df_healthy,100)
faulty_sample = get_recording(df_faulty,105)
print(healthy_sample.shape, faulty_sample.shape)
# -
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_sample)
ax.plot(range(0,size), healthy_sample[:,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,size), healthy_sample[:,1], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(faulty_sample)
ax.plot(range(0,size), faulty_sample[:,1], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,size), faulty_sample[:,0], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
ax.plot(range(0,500), healthy_sample[:500,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,500), healthy_sample[:500,1], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
ax.plot(range(0,500), faulty_sample[:500,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,500), faulty_sample[:500,1], '-', color='blue', animated = True, linewidth=1)
# #### Define and compile autoencoder model
# callback handler, which is called by keras, on the beginning of every training epoch, to record a trajectory of losses during training.
class LossHistory(Callback):
def on_train_begin(self, logs={}):
self.losses = []
def on_batch_end(self, batch, logs={}):
self.losses.append(logs.get('loss'))
# +
timesteps = 100 # we are using 100 past samples to predict 100 future samples
dim = 2 # two accelerometer sensor readings per instance
lossHistory = LossHistory()
# design network
model = Sequential()
model.add(LSTM(50, input_shape=(timesteps, dim), return_sequences=True))
model.add(Dense(2))
model.compile(loss='mae', optimizer='adam')
def train(data):
# we pass data twice, as input and output; this is how an autoencoder works.
model.fit(data, data, epochs=20, batch_size=72, validation_data=(data, data), verbose=1, shuffle=False, callbacks=[lossHistory])
def score(data):
yhat = model.predict(data)
return yhat
# +
# #some learners constantly reported 502 errors in Watson Studio.
# #This is due to the limited resources in the free tier and the heavy resource consumption of Keras.
# #This is a workaround to limit resource consumption
# import os
# # reduce number of threads
# os.environ['TF_NUM_INTEROP_THREADS'] = '1'
# os.environ['TF_NUM_INTRAOP_THREADS'] = '1'
# import tensorflow
# -
# #### Function to create trimmed recordings on which the autoencoder will be trained
def create_trimmed_recording(df, file_id):
recording = get_recording(df, file_id)
# print(recording.shape)
samples = len(recording)
trim = samples % 100
recording_trimmed = recording[:samples-trim]
# print(recording_trimmed.shape)
recording_trimmed.shape = (int((samples-trim)/timesteps), timesteps, dim) # reshape recording array, so it's subdivided in windows of length 'timesteps'.
# print(recording_trimmed.shape)
return recording_trimmed
rec = create_trimmed_recording(df_healthy, 100)
print(rec.shape)
print(rec[0,:5,:])
print(rec[1,:5,:])
# #### Train the autoencoder on healthy data
#pd.unique()
#df_healthy.drop(0,1).drop(2,1).drop(3,1)
pd.unique(df_healthy.iloc[:,1])
# +
file_ids = pd.unique(df_healthy.iloc[:,1])
start = time.time()
for file_id in file_ids:
recording_trimmed = create_trimmed_recording(df_healthy, file_id)
print("Starting training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id, time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
healthy_losses = lossHistory.losses
# -
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_losses)
plt.ylim(0,0.001)
ax.plot(range(0,size), healthy_losses, '-', color='blue', animated = True, linewidth=1)
# +
#file_ids = spark.sql('select distinct _c1 from df_healhty').rdd.map(lambda row : row._c1).collect()
start = time.time()
for file_id in [105]:
recording_trimmed = create_trimmed_recording(df_faulty,file_id)
print("Staring training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id,time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
faulty_losses_105 = lossHistory.losses
# -
pd.unique(df_faulty.iloc[:,1])
# +
file_ids = pd.unique(df_faulty.iloc[:,1])
start = time.time()
for file_id in file_ids:
recording_trimmed = create_trimmed_recording(df_faulty,file_id)
print("Staring training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id,time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
faulty_losses = lossHistory.losses
# -
# #### We append faulty to healthy losses to better identify anomaly
print(len(healthy_losses))
print(len(faulty_losses_105))
print(len(faulty_losses))
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_losses+faulty_losses_105)
plt.ylim(0,0.0006)
ax.plot(range(0,size), healthy_losses+faulty_losses_105, '-', color='blue', animated = True, linewidth=1)
ax.axvline(x=len(healthy_losses), c='magenta', ls='--', lw=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_losses+faulty_losses)
plt.ylim(0,0.0006)
ax.plot(range(0,size), healthy_losses+faulty_losses, '-', color='blue', animated = True, linewidth=1)
ax.axvline(x=len(healthy_losses), c='magenta', ls='--', lw=1)
| 03_Applied_AI_DeepLearning/notebooks/week_3_1_anomaly_detection_keras_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. Load and Explore Dataset
import pandas as pd
import numpy as np
df = pd.read_csv('../data/raw/OnlineNewsPopularity.csv')
df.head()
df.tail()
df.drop(df.tail(1).index,inplace=True)
df.shape
df.info()
df.describe()
# # 4. Prepare Data
# Create a copy of df and save it into a variable called df_cleaned
df_cleaned = df.copy()
# Drop the column url
df_cleaned.drop('url', axis=1, inplace=True)
# Remove leading and trailing space from the column names
df_cleaned.columns = df_cleaned.columns.str.strip()
# Extract the column shares and save it into variable calletarget = df_cleaned.pop('shares')d target
target = df_cleaned.pop('shares')
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
df_cleaned = scaler.fit_transform(df_cleaned)
from joblib import dump
dump(scaler, '../models/scaler.joblib')
# Import train_test_split from sklearn.model_selection
from sklearn.model_selection import train_test_split
# Split randomly the dataset with random_state=8 into 2 different sets: data (80%) and test (20%)
X_data, X_test, y_data, y_test = train_test_split (df_cleaned, target, test_size=0.2, random_state=8)
# Split the remaining data (80%) randomly with random_state=8 into 2 different sets: training (80%) and validation (20%)
X_train, X_val, y_train, y_val = train_test_split(X_data, y_data, test_size=0.2, random_state=8)
# Save the different sets in the folder data/processed
np.save('../data/processed/X_train', X_train)
np.save('../data/processed/X_val', X_val)
np.save('../data/processed/X_test', X_test)
np.save('../data/processed/y_train', y_train)
np.save('../data/processed/y_val', y_val)
np.save('../data/processed/y_test', y_test)
# # 5. Get Baseline Model
# Calculate the average of the target variable for the training set and save it into a variable called y_mean
y_mean = y_train.mean()
# Create a numpy array called y_base of dimensions (len(y_train), 1) filled with this value
y_base = np.full((len(y_train), 1), y_mean)
# Import the MSE and MAE metrics from sklearn
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
# Display the RMSE and MAE scores of this baseline model
print(mse(y_train, y_base, squared=False))
print(mae(y_train, y_base))
# # 6. Train ElasticNet model
# Import the ElasticNet module from sklearn
from sklearn.linear_model import ElasticNet
# instantiate the ElasticNet class into a variable called reg
reg = ElasticNet()
# Fit the model with the prepared data
reg.fit(X_train, y_train)
# Save the fitted model into the folder models as a file called linearelasticnet_default_reg.joblib
dump(reg, '../models/elasticnet_default.joblib')
# Save the predictions from this model for the training and validation sets into 2 variables called y_train_preds and y_val_preds
y_train_preds = reg.predict(X_train)
y_val_preds = reg.predict(X_val)
# Display the RMSE and MAE scores of this model on the training set
print(mse(y_train, y_train_preds, squared=False))
print(mae(y_train, y_train_preds))
# Display the RMSE and MAE scores of this model on the validation set
print(mse(y_val, y_val_preds, squared=False))
print(mae(y_val, y_val_preds))
| adv_dsi_lab_1/notebooks/1_elasticnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Implementation of RAKE
# (Based on: https://www.researchgate.net/publication/227988510_Automatic_Keyword_Extraction_from_Individual_Documents)
# The input text is given below
# +
#Source of text:
#https://www.researchgate.net/publication/227988510_Automatic_Keyword_Extraction_from_Individual_Documents
Text = "Compatibility of systems of linear constraints over the set of natural numbers. \
Criteria of compatibility of a system of linear Diophantine equations, strict inequations, and \
nonstrict inequations are considered. \
Upper bounds for components of a minimal set of solutions and \
algorithms of construction of minimal generating sets of solutions for all \
types of systems are given. \
These criteria and the corresponding algorithms for constructing \
a minimal supporting set of solutions can be used in solving all the \
considered types of systems and systems of mixed types."
# -
# The raw input text is cleaned off non-printable characters (if any) and turned into lower case.
# The processed input text is then tokenized using NLTK library functions.
# +
import nltk
from nltk import word_tokenize
import string
#nltk.download('punkt')
def clean(text):
text = text.lower()
printable = set(string.printable)
text = filter(lambda x: x in printable, text) #filter funny characters, if any.
return text
Cleaned_text = clean(Text)
text = word_tokenize(Cleaned_text)
print "Tokenized Text: \n"
print text
# -
# NLTK is again used for <b>POS tagging</b> the input text.
#
#
# Description of POS tags:
#
#
# http://www.ling.upenn.edu/courses/Fall_2003/ling001/penn_treebank_pos.html
# +
#nltk.download('averaged_perceptron_tagger')
POS_tag = nltk.pos_tag(text)
print "Tokenized Text with POS tags: \n"
print POS_tag
# -
# The tokenized text (mainly the nouns and adjectives) is normalized by <b>lemmatization</b>.
# In lemmatization different grammatical counterparts of a word will be replaced by single
# basic lemma. For example, 'glasses' may be replaced by 'glass'.
#
# Details about lemmatization:
#
# https://nlp.stanford.edu/IR-book/html/htmledition/stemming-and-lemmatization-1.html
# +
#nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
wordnet_lemmatizer = WordNetLemmatizer()
adjective_tags = ['JJ','JJR','JJS']
lemmatized_text = []
for word in POS_tag:
if word[1] in adjective_tags:
lemmatized_text.append(str(wordnet_lemmatizer.lemmatize(word[0],pos="a")))
else:
lemmatized_text.append(str(wordnet_lemmatizer.lemmatize(word[0]))) #default POS = noun
print "Text tokens after lemmatization of adjectives and nouns: \n"
print lemmatized_text
# -
# The <b>lemmatized text</b> is <b>POS tagged</b> here.
# +
POS_tag = nltk.pos_tag(lemmatized_text)
print "Lemmatized text with POS tags: \n"
print POS_tag
# -
# Any word from the lemmatized text, which isn't a noun, adjective, or gerund (or a 'foreign word'), is here
# considered as a <b>stopword</b> (non-content). This is based on the assumption that usually keywords are noun,
# adjectives or gerunds.
#
# Punctuations are added to the stopword list too.
# +
stopwords = []
wanted_POS = ['NN','NNS','NNP','NNPS','JJ','JJR','JJS','VBG','FW']
for word in POS_tag:
if word[1] not in wanted_POS:
stopwords.append(word[0])
punctuations = list(str(string.punctuation))
stopwords = stopwords + punctuations
# -
# Even if we remove the aforementioned stopwords, still some extremely common nouns, adjectives or gerunds may
# remain which are very bad candidates for being keywords (or part of it).
#
# An external file constituting a long list of stopwords is loaded and all the words are added with the previous
# stopwords to create the final list 'stopwords-plus' which is then converted into a set.
#
# (Source of stopwords data: https://www.ranks.nl/stopwords)
#
# Stopwords-plus constitute the sum total of all stopwords and potential phrase-delimiters. The contents of this
# set will be used to partition the lemmatized text into phrases.
#
# Phrases should constitute a group of consecutively occuring words that has no member from stopwords_plus in
# between. Example: "Neural Network".
#
# Each phrase is a <b>keyword candidate</b>.
#
# There are some exceptions, that is, there are some possible cases where a good keyword candidate may contain
# stopword in between. Example: "Word of Mouth".
#
# But, for simplicity's sake I will pretend here that such exceptions do not exist.
# +
stopword_file = open("long_stopwords.txt", "r")
#Source = https://www.ranks.nl/stopwords
lots_of_stopwords = []
for line in stopword_file.readlines():
lots_of_stopwords.append(str(line.strip()))
stopwords_plus = []
stopwords_plus = stopwords + lots_of_stopwords
stopwords_plus = set(stopwords_plus)
#Stopwords_plus contain total set of all stopwords and phrase delimiters that
#will be used for partitioning the text into phrases (candidate keywords).
# -
# Phrases are generated by partitioning the lemmatized text using the members of stopwords_plus
# as delimiters.
# +
phrases = []
phrase = " "
for word in lemmatized_text:
if word in stopwords_plus:
if phrase!= " ":
phrases.append(str(phrase).split())
phrase = " "
elif word not in stopwords_plus:
phrase+=str(word)
phrase+=" "
print "Partitioned Phrases: \n"
print phrases
# -
# Following is the RAKE algorithm.
#
# Frequency of each words in the list of phrases, are calculated here.
#
# The degree of each words are calculating by adding the length of all the
# phrases where the word occurs.
#
# Each word scores are caclulated by dividing degree of the word by its frequency.
#
# +
from collections import defaultdict
from __future__ import division
frequency = defaultdict(int)
degree = defaultdict(int)
word_score = defaultdict(float)
vocabulary = []
for phrase in phrases:
for word in phrase:
frequency[word]+=1
degree[word]+=len(phrase)
if word not in vocabulary:
vocabulary.append(word)
for word in vocabulary:
word_score[word] = degree[word]/frequency[word]
print "Dictionary of degree scores for each words under candidate keywords (phrases): \n"
print degree
print "\nDictionary of frequencies for each words under candidate keywords (phrases): \n"
print frequency
print "\nDictionary of word scores for each words under candidate keywords (phrases): \n"
print word_score
# -
# The phrase scores are calculated by adding individual scores of each of the words
# which form the members of the phrase.
# +
import numpy as np
phrase_scores = []
keywords = []
phrase_vocabulary=[]
for phrase in phrases:
if phrase not in phrase_vocabulary:
phrase_score=0
for word in phrase:
phrase_score+= word_score[word]
phrase_scores.append(phrase_score)
phrase_vocabulary.append(phrase)
phrase_vocabulary = []
j=0
for phrase in phrases:
if phrase not in phrase_vocabulary:
keyword=''
for word in phrase:
keyword += str(word)+" "
phrase_vocabulary.append(phrase)
keyword = keyword.strip()
keywords.append(keyword)
print "Score of candidate keyword '"+keywords[j]+"': "+str(phrase_scores[j])
j+=1
# -
# The index of the phrase score ndarray is then sorted in descending order in terms of
# the score values.
# The index corresponds to the location of the concerned phrase in phrases list.
# So by getting the sorted order of the index, we also get the sorted order of the phrases.
# Each phrase can be considered as a <b>candidate keyword</b>.
# We can then simply choose the top n highest scoring candidate keywords and present them as
# the final exctracted keywords for the system.
# +
sorted_index = np.flip(np.argsort(phrase_scores),0)
keywords_num = 10
print "Keywords:\n"
for i in xrange(0,keywords_num):
print str(keywords[sorted_index[i]])+", ",
# -
# # Input:
#
# Compatibility of systems of linear constraints over the set of natural numbers. Criteria of compatibility of a system of linear Diophantine equations, strict inequations, and nonstrict inequations are considered. Upper bounds for components of a minimal set of solutions and algorithms of construction of minimal generating sets of solutions for all types of systems are given. These criteria and the corresponding algorithms for constructing a minimal supporting set of solutions can be used in solving all the considered types of systems and systems of mixed types.
#
# # Extracted Keywords:
#
# * linear diophantine equation,
# * minimal generating set,
# * minimal supporting set,
# * minimal set,
# * linear constraint,
# * natural number,
# * upper bound,
# * nonstrict inequations
# * strict equations
| Keyword_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This notebook is intented to test, some of the
# results validation of T5 model
import sys
sys.path.append("/home/sidhu/Projects/tf-transformers/src/")
# -
import tensorflow as tf
from tf_transformers.models import T5Model
# +
# Check TF conversion
# !rm -rf /tmp/tf_transformers_cache/t5-base
model_name = 't5-base'
model, config = T5Model.get_model(model_name=model_name, convert_fn_type='tf')
# -
# +
# Chec
# !rm -rf /tmp/tf_transformers_cache/t5-base
model_name = 't5-base'
model, config = T5Model.get_model(model_name=model_name, convert_fn_type='pt')
# -
# +
import numpy as np
from transformers import T5Tokenizer
tokenizer = T5Tokenizer.from_pretrained(model_name)
# +
# T5 text generation without caching
text = "summarize: studies have shown that owning a dog is good for you"
inputs_hf = tokenizer(text, return_tensors='tf')
inputs = {}
inputs['encoder_input_ids'] = inputs_hf['input_ids']
inputs['encoder_input_mask'] = inputs_hf['attention_mask']
inputs['decoder_input_ids'] = tf.constant([[0]])
predictions_non_auto_regressive = []
predictions_prob_non_auto_regressive = []
for i in range(10):
outputs = model(inputs)
predicted_ids = tf.cast(tf.expand_dims(tf.argmax(outputs["last_token_logits"], axis=1), 1), tf.int32)
inputs["decoder_input_ids"] = tf.concat([inputs["decoder_input_ids"], predicted_ids], axis=1)
predictions_non_auto_regressive.append(predicted_ids)
predictions_prob_non_auto_regressive.append(
tf.expand_dims(tf.reduce_max(outputs["last_token_logits"], axis=1), 1)
)
predictions_non_auto_regressive = tf.concat(predictions_non_auto_regressive, axis=1)
predictions_prob_non_auto_regressive = tf.concat(predictions_prob_non_auto_regressive, axis=1)
# Text generation with cache
model, config = T5Model.get_model(model_name=model_name, convert_fn_type='pt', use_auto_regressive=True)
encoder_input_ids = inputs_hf['input_ids']
encoder_input_mask = inputs_hf['attention_mask']
batch_size = tf.shape(encoder_input_ids)[0]
seq_length = tf.shape(encoder_input_ids)[1]
decoder_input_ids = tf.reshape([0] * batch_size, (batch_size,1))
encoder_hidden_dim = config['embedding_size']
num_hidden_layers = config['num_hidden_layers']
num_attention_heads = config['num_attention_heads']
attention_head_size = config['attention_head_size']
encoder_hidden_states = tf.zeros((batch_size, seq_length, encoder_hidden_dim))
decoder_all_cache_key = tf.zeros((num_hidden_layers,
batch_size,
num_attention_heads,
seq_length,
attention_head_size))
decoder_all_cahce_value = tf.zeros((num_hidden_layers,
batch_size,
num_attention_heads,
seq_length,
attention_head_size))
inputs = {}
inputs['encoder_input_ids'] = encoder_input_ids
inputs['encoder_input_mask']= encoder_input_mask
inputs['decoder_input_ids'] = decoder_input_ids
inputs['encoder_hidden_states'] = encoder_hidden_states
inputs['decoder_all_cache_key'] = decoder_all_cache_key
inputs['decoder_all_cache_value'] = decoder_all_cahce_value
predictions_auto_regressive = []
predictions_prob_auto_regressive = []
for i in range(10):
outputs = model(inputs)
predicted_ids = tf.cast(tf.expand_dims(tf.argmax(outputs["last_token_logits"], axis=1), 1), tf.int32)
inputs["decoder_input_ids"] = predicted_ids
inputs["decoder_all_cache_key"] = outputs["decoder_all_cache_key"]
inputs["decoder_all_cache_value"] = outputs["decoder_all_cache_value"]
inputs["encoder_hidden_states"] = outputs["encoder_hidden_states"]
predictions_auto_regressive.append(predicted_ids)
predictions_prob_auto_regressive.append(
tf.expand_dims(tf.reduce_max(outputs["last_token_logits"], axis=1), 1)
)
predictions_auto_regressive = tf.concat(predictions_auto_regressive, axis=1)
predictions_prob_auto_regressive = tf.concat(predictions_prob_auto_regressive, axis=1)
#----------------------------------------------------------------------------------------#
tf.assert_equal(predictions_non_auto_regressive, predictions_auto_regressive)
assert(np.allclose(predictions_prob_non_auto_regressive.numpy(),
predictions_prob_auto_regressive.numpy()) == True)
# -
# +
# Text generation using saved_model with TextDecoder
import tempfile
import shutil
from tf_transformers.text import TextDecoderSeq2Seq
text = "summarize: studies have shown that owning a dog is good for you"
saved_model_dir = tempfile.mkdtemp()
model.save_as_serialize_module(saved_model_dir, overwrite=True)
loaded = tf.saved_model.load(saved_model_dir)
decoder = TextDecoderSeq2Seq(
model = loaded,
decoder_start_token_id = 0 # for t5
)
inputs_hf = tokenizer(text, return_tensors='tf')
inputs = {}
inputs['encoder_input_ids'] = inputs_hf['input_ids']
inputs['encoder_input_mask'] = inputs_hf['attention_mask']
decoder_results = decoder.decode(inputs,
mode='greedy',
max_iterations=10,
eos_id=-100)
expected_ids = [[[ 293, 53, 3, 9, 1782, 19, 207, 21, 25, 6]]]
assert(decoder_results['predicted_ids'].numpy().tolist() == expected_ids)
# -
# +
# Text generation using saved_model with TextDecoderSerializable
import tempfile
import shutil
#from tf_transformers.text import TextDecoderSerializableSeq2Seq
# loaded = tf.saved_model.load(saved_model_dir)
decoder = TextDecoderSerializableSeq2Seq(
model = model,
decoder_start_token_id = 0,
max_iterations=10,
mode="greedy",
do_sample=False,
eos_id=-100
)
# Save
decoder_model = decoder.get_model()
decoder_model.save_serialized(saved_model_dir, overwrite=True)
# Load
loaded_decoder = tf.saved_model.load(saved_model_dir)
model_pb_decoder = loaded_decoder.signatures['serving_default']
text = "summarize: studies have shown that owning a dog is good for you"
inputs_hf = tokenizer(text, return_tensors='tf')
inputs = {}
inputs['encoder_input_ids'] = inputs_hf['input_ids']
inputs['encoder_input_mask'] = inputs_hf['attention_mask']
decoder_results_serialized = model_pb_decoder(**inputs)
np.allclose(decoder_results_serialized['predicted_ids'].numpy(), expected_ids)
# -
| tests/models/t5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monte Carlo A
#
# Simple Monte Carlo Demostration
import numpy as np
np.set_printoptions(precision = 4)
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# generate random variable subject to different distribution
np.random.rand(10)
np.random.normal(loc = 0, scale = 1, size = 10)
np.random.standard_normal(10)
# +
# impose correlation to random variable
corr = np.array([[1, 0.6],
[0.6, 1]])
LT = np.linalg.cholesky(corr).T # cholesky decomposition
dz = np.random.standard_normal((10000,2)) # 10000 * 2 standard normal
dz = np.dot(dz,LT) # impose correlation
print(np.corrcoef(dz[:,0],dz[:,1])) # check the correlation of the new array
# -
# Simulate two stock prices with correlation of 0.6
# +
S0 = 100
r = 0.02
sigma = 0.2
dt = 1/256
path = 100
S = np.zeros((path,2)) # Initiate the array for stock price
S[0] = S0 * np.exp((r-sigma**2/2)*dt + sigma*np.sqrt(dt)*dz[0]) # BSM formula
for i in range(1,path):
S[i] = S[i-1] * np.exp((r-sigma**2/2)*dt + sigma*np.sqrt(dt)*dz[i])
# -
x = np.arange(path)
plt.plot(x, S);
| Code/Monte Carlo A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib
import os
import csv
import time
import math
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import numpy as np
import pandas as pd
from sklearn.metrics import r2_score
from sklearn.preprocessing import scale
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from matplotlib.pyplot import figure
import matplotlib.pyplot as plt
# +
solar_gains = pd.read_csv(r'..\data\energy_demands\solar_gains_testset.csv')
y_test = np.array(solar_gains.loc[:,'Qi(Wh)':].values)
solar_gains = None
inputs_solar = pd.read_csv(r'..\data\inputs\inputs_solar_testset.csv')
X_test = np.array(inputs_solar.loc[:,'G_Dh':].values)
inputs_solar = None
inputs_solar_aux = pd.read_csv(r'..\data\inputs\inputs_solar.csv')
X_aux = np.array(inputs_solar_aux.loc[:,'G_Dh':].values)
inputs_solar_aux = None
# -
#Scaling the data (substracting mean and dividing by the standard deviation)
X_test = np.divide((X_test-X_aux.mean(axis=0)),(X_aux.std(axis=0)))
X_aux = None
class Net(nn.Module):
def __init__(self, input_shape, output_shape):
super(Net, self).__init__()
self.fc1 = nn.Linear(input_shape[1], 20)
self.bn1 = nn.BatchNorm1d(num_features=20)
self.fc2 = nn.Linear(20,10)
self.bn2 = nn.BatchNorm1d(num_features=10)
self.fc3 = nn.Linear(10, output_shape[1])
def forward(self, x):
x = F.relu(self.bn1(self.fc1(x)))
x = F.relu(self.bn2(self.fc2(x)))
return F.relu(self.fc3(x))
device = 'cpu'
net = Net(X_test.shape, y_test.shape).to(device)
criterion = torch.nn.MSELoss()
X_test_torch = torch.tensor(X_test, device='cpu').float()
y_test_torch = torch.tensor(y_test, device='cpu').float()
# +
PATH = r'..\results\dnn_solar'
net.load_state_dict(torch.load(PATH))
net.eval()
start = time.time()
y_pred = net(X_test_torch)
end = time.time()
print(end - start)
# +
loss = criterion(y_pred, y_test_torch)
print(loss.item(), r2_score(y_test_torch.data.numpy(),y_pred.data.numpy()))
# -
mean_squared_error(y_test_torch.data.numpy(),y_pred.data.numpy())
mean_absolute_error(y_test_torch.data.numpy(),y_pred.data.numpy())
# +
figure(num=None, figsize=(16, 8), dpi=80, facecolor='w', edgecolor='k')
matplotlib.rc('xtick', labelsize=22)
matplotlib.rc('ytick', labelsize=22)
plt.plot(list(range(11000,11200)), y_test_torch[11000:11200].data.numpy(),linewidth=3.0)
plt.plot(list(range(11000,11200)), y_pred[11000:11200].data.numpy(),linewidth=3.0)
plt.legend(['Simulation','Neural network prediction'], fontsize=20)
plt.xlabel('hours', fontsize=25)
plt.ylabel('Solar gains (Wh)', fontsize=25)
plt.savefig('solar_gains.png', dpi=600)
# -
| scripts/main-test_solar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Bag of Words Meets Bags of Popcorn
# This is the code from the second part of the tutorial from kaggle.
import pandas as pd
from bs4 import BeautifulSoup
import re
from nltk.corpus import stopwords
import nltk.data
# +
# Read data from files
train = pd.read_csv( "labeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
test = pd.read_csv( "testData.tsv", header=0, delimiter="\t", quoting=3 )
unlabeled_train = pd.read_csv( "unlabeledTrainData.tsv", header=0,
delimiter="\t", quoting=3 )
# Verify the number of reviews that were read (100,000 in total)
print "Read %d labeled train reviews, %d labeled test reviews, " \
"and %d unlabeled reviews\n" % (train["review"].size,
test["review"].size, unlabeled_train["review"].size )
# -
def review_to_wordlist( review, remove_stopwords=False ):
# Function to convert a document to a sequence of words,
# optionally removing stop words. Returns a list of words.
#
# 1. Remove HTML
review_text = BeautifulSoup(review).get_text()
#
# 2. Remove non-letters
review_text = re.sub("[^a-zA-Z]"," ", review_text)
#
# 3. Convert words to lower case and split them
words = review_text.lower().split()
#
# 4. Optionally remove stop words (false by default)
if remove_stopwords:
stops = set(stopwords.words("english"))
words = [w for w in words if not w in stops]
#
# 5. Return a list of words
return(words)
# +
# Load the punkt tokenizer
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
# Define a function to split a review into parsed sentences
def review_to_sentences( review, tokenizer, remove_stopwords=False ):
# Function to split a review into parsed sentences. Returns a
# list of sentences, where each sentence is a list of words
#
# 1. Use the NLTK tokenizer to split the paragraph into sentences
raw_sentences = tokenizer.tokenize(review.strip())
#
# 2. Loop over each sentence
sentences = []
for raw_sentence in raw_sentences:
# If a sentence is empty, skip it
if len(raw_sentence) > 0:
# Otherwise, call review_to_wordlist to get a list of words
sentences.append( review_to_wordlist( raw_sentence, \
remove_stopwords ))
#
# Return the list of sentences (each sentence is a list of words,
# so this returns a list of lists
return sentences
# +
sentences = [] # Initialize an empty list of sentences
print "Parsing sentences from training set"
for review in train["review"]:
sentences += review_to_sentences(review.decode("utf8"), tokenizer)
print "Parsing sentences from unlabeled set"
for review in unlabeled_train["review"]:
sentences += review_to_sentences(review.decode("utf8"), tokenizer)
# +
import logging
from gensim.models import word2vec
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s',\
level=logging.INFO)
# Set values for various parameters
num_features = 300 # Word vector dimensionality
min_word_count = 40 # Minimum word count
num_workers = 4 # Number of threads to run in parallel
context = 10 # Context window size
downsampling = 1e-3 # Downsample setting for frequent words
# Initialize and train the model (this will take some time)
print "Training model..."
model = word2vec.Word2Vec(sentences, workers=num_workers, \
size=num_features, min_count = min_word_count, \
window = context, sample = downsampling)
# If you don't plan to train the model any further, calling
# init_sims will make the model much more memory-efficient.
model.init_sims(replace=True)
# It can be helpful to create a meaningful model name and
# save the model for later use. You can load it later using Word2Vec.load()
model_name = "300features_40minwords_10context"
model.save(model_name)
# -
model.doesnt_match("man woman child kitchen".split())
model.doesnt_match("france england germany berlin".split())
model.most_similar("man")
model.most_similar("queen")
model.most_similar("awful")
model["flower"]
| Word2Vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Now You Code 4: Sentiment v1.0
#
# Let's write a basic sentiment analyzer in Python. Sentiment analysis is the act of extracting mood from text. It has practical applications in analyzing reactions in social media, product opinions, movie reviews and much more.
#
# The 1.0 version of our sentiment analyzer will start with a string of positive and negative words. For any input text and the sentiment score will be calculated by simply adding up the frequency of the positive words, then subtracting out the negative words.
#
# So for example, if:
#
# ```
# positive_text = "happy glad like"
# negative_text = "angry mad hate"
# input_text = "Amazon makes me like so angry and mad"
# score = -1 [ +1 for like, -1 for angry, -1 for mad]
# ```
#
# You will complete this program by first writing the sentiment function, then writing some tests for it.
#
# You will conclude by writing the complete sentiment analyzer to score any sentence.
#
#
# ## Problem Analysis For Sentiment Function
#
#
# You want to write `ScoreSentiment()` as a function:
#
# - Function: `ScoreSentiment()`
# - Arguments (input): `postive_text, negative_text, input_text`
# - Returns (output): `score (int)`
#
# Algorithm (Steps in Program):
#
# ```
# for each word in our tokenized input_text
# if word in positive_text then
# increment seniment score
# else if word in negative_text then
# decrement sentiment score
# ```
#
# ## Step 1: Write the function
#
def ScoreSentiment(positive_text, negative_text, input_text):
#TODO write code here
return score
# ## Step 2: Write tests for the function
#
# With the function complete, we need to test our function. The simplest way to do that is call the function with inputs we expect and verify the output. For example:
#
# ```
# pos_text='happy joy good'
# neg_text ='sad pain bad'
#
# WHEN input_text='I am sad with joy' We EXPECT ScoreSentiment(pos_text, neg_text, input_text) to return 0
# WHEN input_text='I am sad and in pain' We EXPECT ScoreSentiment(pos_text, neg_text, input_text) to return -2
# WHEN input_text='I am happy with joy' We EXPECT ScoreSentiment(pos_text, neg_text, input_text) to return 2
# ```
#
# +
## TODO write tests here.
# -
# ## Step 3: Write final program
#
# Then write a main program that executes like this:
#
# Sample Run
#
# ```
# Sentiment Analyzer 1.0
# Type 'quit' to exit.
# Enter Text: i love a good book from amazon
# 2 positive.
# Enter Text: i hate amazon their service makes me angry
# -2 negative.
# Enter Text: i love to hate amazon
# 0 neutral.
# Enter Text: quit
# ```
#
# NOTE: make up your own strings of positive and negative words to make the sentiment more accurate.
# ### 3.a : Problem Analysis
#
# Inputs:
#
#
# Outputs:
#
#
# Algorithm:
#
#
# +
## TODO: 3b write program
# -
# ## Step 4: Questions
#
# 1. What can be done to make the sentiment more accurate?
#
# Answer:
#
#
# 2. Do you see a problem with the method of scoring?
#
#
# Answer:
#
#
# 3. Does the function improve readability of the final program in 3.b? Why or why not?
#
#
# Answer:
#
# ## Step 5: Reflection
#
# Reflect upon your experience completing this assignment. This should be a personal narrative, in your own voice, and cite specifics relevant to the activity as to help the grader understand how you arrived at the code you submitted. Things to consider touching upon: Elaborate on the process itself. Did your original problem analysis work as designed? How many iterations did you go through before you arrived at the solution? Where did you struggle along the way and how did you overcome it? What did you learn from completing the assignment? What do you need to work on to get better? What was most valuable and least valuable about this exercise? Do you have any suggestions for improvements?
#
# To make a good reflection, you should journal your thoughts, questions and comments while you complete the exercise.
#
# Keep your response to between 100 and 250 words.
#
# `--== Write Your Reflection Below Here ==--`
#
#
| content/lessons/06/Now-You-Code/NYC4-Sentiment-v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification with Scikit-Learn
#
# In this lesson, you will learn the basic functionality of Scikit-Learn, one of the most important Machine Learning packages in Python. We will use an example dataset of Iris flowers.
#
# ### Cheat sheet
# https://s3.amazonaws.com/assets.datacamp.com/blog_assets/Scikit_Learn_Cheat_Sheet_Python.pdf
# ## Concepts
#
# | concept | description |
# |:-----------:|:-----------:|
# | Estimators | how models in Scikit-learn are called |
# | m.fit() | method to train |
# | m.predict() | creates a prediction for unknown data |
# | m.transform() | transforms features (in some models) |
# | train_test_split() |splits data in a training and test portion |
# | random_state | parameter for reproducible random numbers |
#
# ### 1. Load the example data
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data[:,:2]
y = iris.target
# ### 2. Constructing a model in Scikit-Learn
from sklearn import svm
from sklearn.model_selection import train_test_split
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y,
test_size = 0.5,
random_state = 42)
# +
model = svm.SVC(kernel = 'linear', C=1.0)
model.fit(Xtrain, ytrain)
print("Train score: ", model.score(Xtrain, ytrain))
print("Test score: ", model.score(Xtest, ytest))
# -
# ### 3. Predictions for unknown data
# +
import numpy as np
Xnew = np.array([[5.0, 3.3], [4.7, 2.1]])
ypred = model.predict(Xnew)
print("predictions:")
for x, y in zip(Xnew, ypred):
label = iris.target_names[y]
print(x, y, "->", label)
# -
# ## Exercise
#
# 1. Evaluate a SVC classifier on the iris data.
# 2. Use a SVC classifier on the Titanic data.
# 3. Find out what classification methods are there on the Scikit-Learn website. Use one or more of them.
# 4. Use sklearn.dummy.DummyClassifier with the default parameters, resulting in a 33% chance for each type of iris.
# 5. Change the parameters so that sklearn.dummy.DummyClassifier predicts 1 for any data point.
# 6. Evaluate both classifiers on the same dataset.
| 02_ClassificationWithScikit-Learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Given a string of opening and closing parentheses, check whether it's
balanced. (), {}, [] Assume that the string doesn't contain any other
character other than these brackets, no spaces, words, or numbers.
Balanced parentheses require every opening parenthesis to be closed in the
reverse order opened.
Ex. '([])' is balanced but '([)]' is not.
"""
def balance_check(s):
if len(s) % 2 != 0:
return False
opening = set('([{')
matches = set([('(',')'), ('[',']'), ('{','}')])
stack = []
for paren in s:
if paren in opening:
stack.append(paren)
else:
if len(stack) == 0:
return False
last_open = stack.pop()
if (last_open, paren) not in matches:
return False
return len(stack) == 0
balance_check('[]')
balance_check('()[]{}')
balance_check('([)]')
| Python for Algorithms, Data Structures, & Interviews/ Stacks, Queues, and Deques/Balanced Parentheses Check - Interview Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# Compute the x and y coordinates for points on a sine curve
x = np.arange(0, 3 * np.pi, 0.1)
y = np.sin(x)
# Plot the points using matplotlib
plt.plot(x, y)
plt.show() # You must call plt.show() to make graphics appear.
# -
y = 10
x = [0,1,2,3,4,5,6,7]
plt.plot(for i in x,y)
plt.show()
| python/deep_learning/NOTEBOOK/Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IPython 与 RQAlpha
# ## 加载 RQAlpha magic
# %load_ext rqalpha
# ## 查看 RQAlpha magic 帮助
#
# 我们可以通过 `%%rqalpha` 直接在 `cell` 中运行回测代码。 `%%rqalpha` 后面的参数等价于在 CLI 中后面的 `rqalpha run` 的参数
# %%rqalpha -h
""
# ## 使用 %%rqalpha 进行回测
# +
# %%rqalpha -s 20100101 -e 20170505 -p -bm 000001.XSHG --account stock 100000
def init(context):
context.stocks = ['000300.XSHG', '000905.XSHG', '000012.XSHG']
def handle_bar(context, bar_dict):
[hs, zz, gz] = context.stocks
hs_history20 = history_bars(hs, 20, '1d', 'close')
zz_history20 = history_bars(zz, 20, '1d', 'close')
hsIncrease = hs_history20[-1] - hs_history20[0]
zzIncrease = zz_history20[-1] - zz_history20[0]
positions = context.portfolio.positions
[hsQuality, zzQuality, gzQuality] = [positions[hs].quantity, positions[zz].quantity, positions[gz].quantity]
if hsIncrease < 0 and zzIncrease < 0:
if hsQuality > 0: order_target_percent(hs, 0)
if zzQuality > 0: order_target_percent(zz, 0)
order_target_percent(gz, 1)
elif hsIncrease < zzIncrease:
if hsQuality > 0: order_target_percent(hs, 0)
if gzQuality > 0: order_target_percent(gz, 0)
order_target_percent(zz, 1)
else:
if zzQuality > 0: order_target_percent(zz, 0)
if gzQuality > 0: order_target_percent(gz, 0)
order_target_percent(hs, 1)
#logger.info("positions hs300: " + str(hsQuality) + ", zz500: " + str(zzQuality) + ", gz: " + str(gzQuality))
# -
# ## 获取回测报告
#
# 运行完回测后,报告会自动存储到 `report` 变量中。可以直接通过 `report` 变量获取当次回测的结果。
#
# 另外 rqalpha 的 mod 的输出会自动存储在 `results` 变量中。
results.keys()
report.keys()
report.trades[:5]
report.portfolio[:5]
report.stock_positions[:5]
# ## 使用 run_func 运行回测
# +
config = {
"base": {
"start_date": "2010-01-01",
"end_date": "2017-05-05",
"benchmark": "000001.XSHG",
"accounts": {
"stock": 100000
}
},
"extra": {
"log_level": "info",
},
"mod": {
"sys_analyser": {
"enabled": True,
"plot": True,
},
}
}
from rqalpha.api import *
from rqalpha import run_func
def init(context):
context.stocks = ['000300.XSHG', '000905.XSHG', '000012.XSHG']
def handle_bar(context, bar_dict):
[hs, zz, gz] = context.stocks
hs_history20 = history_bars(hs, 20, '1d', 'close')
zz_history20 = history_bars(zz, 20, '1d', 'close')
hsIncrease = hs_history20[-1] - hs_history20[0]
zzIncrease = zz_history20[-1] - zz_history20[0]
positions = context.portfolio.positions
[hsQuality, zzQuality, gzQuality] = [positions[hs].quantity, positions[zz].quantity, positions[gz].quantity]
if hsIncrease < 0 and zzIncrease < 0:
if hsQuality > 0: order_target_percent(hs, 0)
if zzQuality > 0: order_target_percent(zz, 0)
order_target_percent(gz, 1)
elif hsIncrease < zzIncrease:
if hsQuality > 0: order_target_percent(hs, 0)
if gzQuality > 0: order_target_percent(gz, 0)
order_target_percent(zz, 1)
else:
if zzQuality > 0: order_target_percent(zz, 0)
if gzQuality > 0: order_target_percent(gz, 0)
order_target_percent(hs, 1)
results = run_func(init=init, handle_bar=handle_bar, config=config)
# -
report = results["sys_analyser"]
report["trades"][:5]
| docs/source/notebooks/run-rqalpha-in-ipython.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
sns.set_style('darkgrid')
plt.rcParams['font.size'] = 15
plt.rcParams['figure.figsize'] = (10,7)
plt.rcParams['figure.facecolor'] = '#FFE5B4'
data=pd.read_csv("C:\happiness_score_dataset.csv")
data.head()
data_columns =['Country','Region','Happiness Score','Economy (GDP per Capita)','Family','Health (Life Expectancy)','Freedom','Trust (Government Corruption)',
'Generosity']
data = data[data_columns].copy() ## Here i take what is necessary for me.
data.rename(columns= {'Happiness Score':'happiness_score','Happiness Rank':'happiness_rank','Economy (GDP per Capita)':'economy','Health (Life Expectancy)':'health','Trust (Government Corruption)':'Corruption'},inplace = True)
# ##### In Above I Seperated Unused Data.
data
data.isnull().sum() #Checking the null values here.
# +
# Plot between happiness and GDP
plt.rcParams['figure.figsize']= (15,7)
plt.title('plot between Happiness score and GDP')
sns.scatterplot(x =data['happiness_score'], y = data['economy'] , hue = data.Region, s =100);
plt.legend(loc = 'upper left', fontsize = '10')
plt.xlabel('Happiness_score')
plt.ylabel('GDP per capita')
# -
# #### In above Graph the Happiness score and GDP per capita i.E ECONOMY is very low in sub-Saharian region.
# #### Whereareas WESTERN EUROPE has high Happiness score and also some part of LATIN AMERICA AND CARIBBEAN.
# +
# Plot between happiness and GDP
plt.rcParams['figure.figsize']= (15,7)
plt.title('Plot between Family and Generosity')
sns.scatterplot(x =data.Family , y = data.Generosity , hue = data.Region, s =100);
plt.legend(loc = 'upper left', fontsize = '10')
plt.xlabel('Family')
plt.ylabel('Generosity')
# -
gdp_region = data.groupby('Region')['economy'].sum()
gdp_region
gdp_region.plot.pie(autopct = '%1.1f%%') #parameter i.e autopct
plt.title('GDP by region')
plt.ylabel('')
# #### The WESTERN EUROPE AND CENTRAL AND EAST EUROPE is contributing more in GDP.
# #### AUSTRALIA AND NEW ZEALAND is (Contributing)having less GDP by region.
# part of pandas function
# Total countries
total_country = data.groupby('Region')[['Country']].count()
print(total_country)
# +
# correlation Map
cor =data.corr(method = "pearson") ## I use pearson method incorrelation.
f, ax = plt.subplots(figsize =(10,5))
sns.heatmap(cor, mask =np.zeros_like(cor), ## dtype=np.bool
cmap="Blues", square=True, ax=ax)
# -
# #### The dark blue cell colour shows high correlation and light blue greyish show low correlation.
## going to visualise the bar plot
# Corruption in region
Tcorruption = data.groupby('Region')[['Corruption']].mean()
Tcorruption
# #### Australia and New zealand has highest corruption.
# #### Central and eastern Europe has least corruption.
# Barplot
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize']= (12, 8)
plt.title('Corruption in various regions')
plt.xlabel('Region', fontsize = 15)
plt.ylabel('Corruption Index', fontsize =15 )
plt.xticks(rotation =30 , ha='right') ## for horizontal alignment.
plt.bar(Tcorruption.index, Tcorruption.Corruption)
Top_10 = data.head(10)
bottom_10 = data.tail(10)
# +
fig, axes= plt.subplots(1,2, figsize= (16, 6))
plt.tight_layout(pad= 2)
xlabels = Top_10.Country
axes[0].set_title('top 10 happiest country')
axes[0].set_xticklabels(xlabels, rotation=45, ha ='right')
sns.barplot(x= Top_10.Country,y= Top_10.health,ax= axes[0])
axes[0].set_xlabel('Country Name')
axes[0].set_ylabel('Life expectancy(Health)')
xlabels = bottom_10.Country
axes[1].set_title('Bottom 10 Least happiest country')
axes[1].set_xticklabels(xlabels, rotation=45, ha='right')
sns.barplot(x= bottom_10.Country,y= bottom_10.health,ax= axes[1])
axes[1].set_xlabel('Country Name')
axes[1].set_ylabel('Life expectancy(Health)')
# -
data.head(10)
data.tail(10)
plt.rcParams['figure.figsize']= (15, 7)
sns.scatterplot(x =data.Freedom , y = data.happiness_score , hue = data.Region, s=100);
plt.legend(loc ='upper left', fontsize ='12')
plt.xlabel('freedom to make life choices')
plt.ylabel('Happiness score')
# #### Western region has High happiness score and Freedom to make life choices.
country = data.sort_values(by= 'Corruption').head(10)
plt.rcParams['figure.figsize']= (12,6)
plt.title('Country with most corruption')
plt.xlabel('Country', fontsize = 13)
plt.ylabel('Corruption index', fontsize =13)
plt.xticks(rotation =30,ha ='right')
plt.bar(country.Country, country.Corruption)
country = data.sort_values(by= 'Corruption').tail(10)
plt.rcParams['figure.figsize']= (12,6)
plt.title('Country with most corruption')
plt.xlabel('Country', fontsize = 13)
plt.ylabel('Corruption index', fontsize =13)
plt.xticks(rotation =30,ha ='right')
plt.bar(country.Country, country.Corruption)
# +
## Corruption V/S Happiness Score.
plt.rcParams['figure.figsize']= (15,7)
sns.scatterplot(x =data.happiness_score , y = data.Corruption , hue = data.Region, s=100);
plt.legend(loc ='upper left', fontsize ='13')
plt.ylabel('Corruption')
plt.xlabel('Happiness score')
# -
# #### 1)From the above Analysis I observe that Western Europe countries like Switzerland,Denmark,Finland,norway,Swaden has Good #### and high Happiness Score then other Region Apart from that 2) Latin America and Caribbean region Has high happiness
# #### Score And Less
# #### corruption as compare to other region in term of Generiosity also it is High.
#
# #### 3)Sub-Saharian Africa has low Happiness score which consist of 40 countries which is large in number They also have low
# #### Freedom to make choices And corruption is also moderate.
#
# #### 4) In terms of correlation -Economy and Happiness score has high coorelation and Health and Economy,Health and happiness
# #### score also has high correlation.
#
# #### 5)Generiosity and economy has low correlation and curruption and Happiness has moderate correlation.
# #### 6) Western and Eastern Region Contribute more in GDP i.e(20.4%) Whereas Australia and New Zeland contribute ####less(Economy).It contain 2 countries with Highest corruption Rate.
#
#
#
| happines report Final .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 1 for JetSeT v1.2.0-rc3
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
import numpy as np
# -
# ## Basic setup and access to Jet class
import jetset
print(jetset.__version__)
# See for more details:
#
# - https://jetset.readthedocs.io/en/latest/user_guide/jet_model_phys_SSC/Jet_example_phys_SSC.html
#
from jetset.jet_model import Jet
my_jet=Jet(electron_distribution='lppl')
Jet.available_electron_distributions()
my_jet.show_pars()
my_jet.parameters.par_table
my_jet.show_model()
my_jet.set_par('B',val=0.2)
my_jet.set_par('gamma0_log_parab',val=5E3)
my_jet.set_par('gmin',val=1E2)
my_jet.set_par('gmax',val=1E8)
my_jet.set_par('R',val=1E15)
my_jet.set_par('N',val=1E3)
my_jet.parameters.B.val=0.2
my_jet.parameters.r.val=0.4
my_jet.show_electron_distribution()
p=my_jet.electron_distribution.plot()
p=my_jet.electron_distribution.plot(energy_unit='TeV')
p=my_jet.electron_distribution.plot3p()
my_jet.eval()
from jetset.plot_sedfit import PlotSED
my_plot=PlotSED()
my_plot=my_jet.plot_model(plot_obj=my_plot)
#my_plot.rescale(y_max=-12,y_min=-17.5,x_min=8)
my_plot=my_jet.plot_model(frame='src')
my_plot.rescale(y_max=45,y_min=38,x_min=8)
my_plot=my_jet.plot_model(frame='src',density=True)
my_jet.list_spectral_components()
Sync=my_jet.spectral_components.Sync
Sync=my_jet.get_spectral_component_by_name('Sync')
nu_sync=Sync.SED.nu
nuFnu_sync=Sync.SED.nuFnu
nu_sync_src=Sync.SED.nu_src
nuLnu_sync_src=Sync.SED.nuLnu_src
my_jet.spectral_components.build_table(restframe='obs')
t_obs=my_jet.spectral_components.table
t_obs[::10]
my_jet.spectral_components.build_table(restframe='src')
t_src=my_jet.spectral_components.table
t_obs['Sync'][::10].to('GeV/cm2 s')
t_src.write('test_SED.txt',format='ascii.ecsv',overwrite=True)
my_jet.energetic_report()
my_jet.energetic_report_table
my_jet.save_model('test_model.pkl')
my_jet_new=Jet.load_model('test_model.pkl')
my_plot=my_jet_new.plot_model()
my_plot.rescale(y_max=-11,y_min=-17.5,x_min=8)
# ## Define custom emiotters distribution
# See for more details:
#
# - https://jetset.readthedocs.io/en/latest/user_guide/custom_emitters_distr/custom_emitters.html
from jetset.jet_emitters import EmittersDistribution
def distr_func_super_exp(gamma,gamma_cut,s,a):
return np.power(gamma,-s)*np.exp(-(1/a)*(gamma/gamma_cut)**a)
n_e_super_exp=EmittersDistribution('super_exp',spectral_type='plc',normalize=False)
n_e_super_exp.add_par('gamma_cut',par_type='turn-over-energy',val=50000.,vmin=1., vmax=None, unit='lorentz-factor')
n_e_super_exp.add_par('s',par_type='LE_spectral_slope',val=2.3,vmin=-10., vmax=10, unit='')
n_e_super_exp.add_par('a',par_type='spectral_curvature',val=1.8,vmin=0., vmax=100., unit='')
n_e_super_exp.set_distr_func(distr_func_super_exp)
n_e_super_exp.parameters.show_pars()
p=n_e_super_exp.plot()
p=n_e_super_exp.plot(energy_unit='eV')
from jetset.jet_model import Jet
my_jet=Jet(electron_distribution=n_e_super_exp)
n_e_super_exp.normalize
my_jet.electron_distribution.normalize
my_jet.parameters.N.val=5E4
my_jet.show_model()
my_jet.IC_nu_size=100
my_jet.eval()
my_jet.eval()
p=my_jet.plot_model()
p.rescale(y_min=-16,y_max=-13)
my_jet.electron_distribution.normalize=True
# +
my_jet.parameters.N.val=5E4
my_jet.show_model()
my_jet.IC_nu_size=100
my_jet.eval()
# -
my_jet.plot_model(p,comp='Sum',label='Normalized distr')
p.rescale(y_min=-16,y_max=-12)
p.fig
| notebooks/Tutorial_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Library
# +
import pandas as pd
import numpy as np
import re
import pickle
import os
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from fbprophet import Prophet
from joblib import Parallel, delayed
import multiprocessing
# +
def temp_func(func, name, group):
return func(group), name
def applyParallel(dfGrouped, func):
retLst, top_index = zip(
*Parallel(n_jobs=multiprocessing.cpu_count()-1)(delayed(temp_func)(
func, name, group) for name, group in dfGrouped))
return pd.concat(retLst, keys=top_index)
# -
# # Scoring functions
# +
def smape(y_true, y_pred):
"""
Scoring function
"""
denominator = (np.abs(y_true) + np.abs(y_pred)) / 2.0
diff = np.abs(y_true - y_pred) / denominator
diff[denominator == 0] = 0.0
return 100 * np.mean(diff)
def smape_serie(x):
"""
Scoring function on serie
"""
return smape(y_pred=x.Visits, y_true=x.value)
# -
# # Helping functions
# +
def create_train():
if os.path.isfile("../data/work/train.pickle"):
data = pd.read_pickle("../data/work/train.pickle")
else:
data = pd.read_csv('../data/input/train_2.csv')
cols = data.columns[data.columns.str.contains("-")].tolist()
data["Page"] = data["Page"].astype(str)
data = data.set_index("Page").T
data.index = pd.to_datetime(data.index, format="%Y-%m-%d")
data.to_pickle("../data/work/train.pickle")
return data
def create_test():
if os.path.isfile("../data/work/test.pickle"):
df_test = pd.read_pickle("../data/work/test.pickle")
else:
df_test = pd.read_csv("../data/input/key_2.csv")
df_test['date'] = df_test.Page.apply(lambda a: a[-10:])
df_test['Page'] = df_test.Page.apply(lambda a: a[:-11])
df_test['date'] = pd.to_datetime(df_test['date'], format="%Y-%m-%d")
df_test.to_pickle("../data/work/test.pickle")
return df_test
# -
# # Read data
# + code_folding=[]
data = create_train()
print(data.info())
data.head()
# -
# # Train / Test
## Split in train / test to evaluate scoring
train = data.iloc[:-60]
test = data.iloc[-60:]
print(train.shape)
print(test.shape)
print(data.shape)
# # Prophet
def prophet_forecast(df):
return Prophet(
yearly_seasonality=False,
daily_seasonality=False,
weekly_seasonality="auto",
seasonality_prior_scale=5,
changepoint_prior_scale=0.5).fit(df.dropna()).predict(df_predict)[[
"ds", "yhat"
]]
# ## Test
df_predict = pd.DataFrame({"ds": test.index})
df_predict.head()
# +
# page_sample = train.columns[np.random.randint(0, len(train.columns), 10)]
# train_sample = train[page_sample].reset_index().rename(
# columns={"index": "ds"}).melt(id_vars="ds").rename(columns={"value":
# "y"}).dropna()
# test_sample = test[page_sample]
# train_sample.head()
# -
forecast = applyParallel(train.groupby("Page"),
prophet_forecast).reset_index().rename(
columns={"level_0": "Page"}).drop(
"level_1", axis=1)
forecast.head()
forecast = pd.merge(
test_sample.reset_index().rename(columns={"index": "ds"}).melt(
id_vars="ds"),
forecast,
on=["ds", "Page"],
how="inner")
forecast.head()
print("SMAPE is : ")
print(smape(y_pred=forecast["value"], y_true=forecast["yhat"]))
| script/prophet_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ss2020_tvb]
# language: python
# name: conda-env-ss2020_tvb-py
# ---
# + [markdown] colab_type="text" id="KaFIl1Z6r3pz"
# <br>
# <div align="center"><font size="7" face="calibri" color="#000099">Modelling Resting State Brain Dynamics</font></div>
# <br>
# <div align="center"><font size="7" face="calibri" color="#000099">using The Virtual Brain (TVB)</font></div>
# <br><br>
# <div align="center"><span style="font-weight:normal"><font size="4" face="calibri"><b><NAME></b></font></span></div>
#
# <div align="center"><span style="font-weight:normal"><font size="4" face="calibri"><b><NAME></b></font></span></div>
#
# <div align="center"><span style="font-weight:normal"><font size="4" face="calibri"><b><NAME></b></font></span></div>
#
# <div align="center"><span style="font-weight:normal"><font size="4" face="calibri"><b><NAME></b></font></span></div>
# + [markdown] colab_type="text" id="CfpIJUcQr3p1"
# ---
#
# <h2><font size="6" color="#609BC4" face="calibri">Contents</font></h2>
# + [markdown] colab_type="text" id="XMXH0jYSr3p2"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
#
# <a href="#Overview">Overview</a>
# <br>
# <a href="#Setup">Setup</a>
# <br>
# <a href = "#Load-and-prepare-data">Load and prepare data</a>
# <br>
# <a href = "#Computational-model">Computational model</a>
# <br>
# <a href = "#Optimal-working-reigon-of-the-model">Optimal working region of the model</a>
# <br>
# <a href = "#Compute-FC-for-the-best-working-point">Compute FC for the best working point</a>
# <br>
# <a href = "#Conclusions">Conclusions</a>
# <br>
# <a href = "#References">References</a>
#
# </font></div></p>
# + [markdown] colab_type="text" id="2khkeAk3r3p4"
# ---
#
# <h2><font size="6" face="calibri" color="#609BC4">Overview</font></h2>
# + [markdown] colab_type="text" id="_XiIkR2Gr3p5"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
# A current topic in systems neuroscience literature is the presence of brain activity in the absence of a task. These spontaneous fluctuations occur in the so-called <b>resting state</b>. A recurring theme of these fluctuations is that they are not random: instead, the resting brain displays spatial patterns of correlated activity across different brain regions, called <b>resting-state networks</b>.
#
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
# These patterns of <b>resting-state functional connectivity (FC)</b> relate to the underlying anatomical structure of the brain, which can be estimated using diffusion spectrum imaging (DSI). Because The Virtual Brain uses this <b>structural connectivity (SC)</b> as the backbone for simulating spontaneous activity, resting-state activity and its network structure is a prime candidate for modeling in TVB.
#
#
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
#
# <b>In this tutorial, we will:</b>
# <br>
# <ul>
# <li>build a resting brain model using subject-specific structural connectivity (SC), defined using probabilistic tractography, </li>
# <li>generate its resting-state fMRI BOLD signals,</li>
# <li>identify the dynamical working region of the model,</li>
# <li>perform a parameter space exploration to identify regions of improved correlations between simulated and empirical FC.</li>
# </ul>
#
# </font></div></p>
# + [markdown] colab_type="text" id="2wd_TUntr3p6"
# ---
#
# <h2><font size="6" face="calibri" color="#609BC4">Setup</font></h2>
# + [markdown] colab_type="text" id="-XWiJv0MaARv"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
# We will now import the Python packages that we need for the simulations and visualizations </font></div></p>
# +
# If running in google colab, uncomment the install commands and execute this cell:
# TVB scientific library
# #!pip install tvb-library
# TVB datasets
# #!pip install tvb-data
# + colab={} colab_type="code" id="cas5l1wXr3p7"
# imports
import warnings
warnings.filterwarnings('ignore')
import os, sys, scipy.io, numpy as np, seaborn as sns
from pprint import pprint
import timeit, time as tm
from matplotlib import pyplot as plt
from IPython.display import HTML
import zipfile
from scipy.io import loadmat, savemat
# You may need to change these to the correct paths for your system
#tvb_lib_dir = '/scinet/course/ss2019/3/9_brainnetwork/tvb-library'
#tvb_dat_dir = '/scinet/course/ss2019/3/9_brainnetwork/tvb-data'
#sys.path += [tvb_lib_dir,tvb_dat_dir]
from tvb.simulator.lab import *
from tvb.datatypes.time_series import TimeSeriesRegion
#import tvb.analyzers.correlation_coefficient as corr_coeff
# %matplotlib inline
sns.set()
# -
# This is a utility function that gives a syntactically minimal way of writing a = np.array([a])
# (which is needed for many TVB method calls defining scalar parameters etc.)
def __(num): return np.array([num])
# + [markdown] colab_type="text" id="bXGOTOlzG4yS"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
# TVB simulations can take a while to run. In this tutorial we will show you how to run the simulations, but we won't actually run them. Instead, we will load the results from simulations that we ran beforehand. Run the following cell to download the data that we will be using for today's tutorial. </font></div></p>
# + [markdown] colab_type="text" id="fdHZBW_cr3p_"
# ---
#
# <h2><font size="6" face="calibri" color="#609BC4">Load and prepare data</font></h2>
# + [markdown] colab_type="text" id="MJDnnclFr3qA"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
# Here, we use a <b>Structural Connectivity (SC) of 66 regions</b> derived from Diffusion Spectrum Imaging (DSI) and tractography, as previously published in <b>Hagmann et al. (2008)</b> with the modifications introduced by <b>Cabral et al. (2011)</b>. Connections in this SC matrix were defined with a standard parcellation scheme (<b>Desikan et al., 2006</b>), and averaged over 5 healthy right-handed male human subjects.
# </font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
#
# We use a <b>resting-state Functional Connectivity (FC)</b> obtained from the same 5 human subjects and using the same 66 cortical areas adopted for the SC above. The resting-state FC is calculated by measuring the corresponding <b>fMRI BOLD signals</b> during the entire duration of <b>20 min</b>, and then defining FC as the Pearson correlation coefficient between the time series for each pair of the 66 regions.
#
# </font></div></p>
# + colab={} colab_type="code" id="OewLN4hUr3qB"
Hag_con = connectivity.Connectivity.from_file(os.path.abspath('../data/connectivity_HagmannDeco66.zip'))
nregions = len(Hag_con.region_labels) #number of regions
Hag_con.speed = __(np.inf) #set the conduction speed to infinity => no time delays
Hag_con.configure()
Hag_SC = Hag_con.weights
Hag_tract_lengths = Hag_con.tract_lengths
Hag_con.region_labels[33:]=Hag_con.region_labels[33:][::-1]
Hag_FC = np.load('../data/Hagmann_empFC_avg.npy')
# +
# Visualization
fig=plt.figure(figsize=(16,12))
# weights
plt.subplot(221)
plt.imshow((Hag_con.weights), interpolation='nearest', aspect='equal', cmap='magma')
plt.grid('off')
plt.xticks(range(0, nregions), Hag_con.region_labels, fontsize=7, rotation=90)
plt.yticks(range(0, nregions), Hag_con.region_labels, fontsize=7)
cb=plt.colorbar(shrink=0.5)
cb.set_label('weight', fontsize=14)
plt.title('Hagmann SC weights')
#tracts
plt.subplot(222)
plt.imshow(Hag_con.tract_lengths, interpolation='nearest', aspect='equal', cmap='magma')
plt.grid('off');
plt.xticks(range(0, nregions), Hag_con.region_labels, fontsize=7, rotation=90)
plt.yticks(range(0, nregions), Hag_con.region_labels, fontsize=7)
cb=plt.colorbar(shrink=0.5)
cb.set_label('tract length (mm)', fontsize=14)
plt.title('Hagmann SC tract lengths')
# FC
plt.subplot(223)
plt.imshow(Hag_FC, interpolation='nearest', aspect='equal',
cmap='RdBu_r', vmin=-.5, vmax=.5)
plt.grid('off')
plt.xticks(range(0, nregions), Hag_con.region_labels, fontsize=7, rotation=90)
plt.yticks(range(0, nregions), Hag_con.region_labels, fontsize=7)
cb=plt.colorbar(shrink=0.5)
cb.set_label('Pearson Correlation Coefficient', fontsize=14)
plt.title('Hagmann FC', fontsize=14)
fig.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="6_Sp2SAor3qO"
# <h3><font size="5" face="calibri" color="black">SC-FC comparison</font></h3>
# + [markdown] colab_type="text" id="eVjEtpWLr3qP"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">
#
# We compare the SC and FC matrix of the empirical data by adopting as a measure of similarity between the two matrices the Pearson correlation between corresponding elements of the <b>upper (or lower)</b> triangular part of the matrices.
#
# </font></div></p>
# +
# Take upper triangular part of the matrices (excluding the self-connections).
inds = np.triu_indices(66,1)
Hag_SC_triu = Hag_SC[inds]
Hag_FC_triu = Hag_FC[inds]
# non-zero connections from upper triangle
non0 = np.where(Hag_SC_triu!=0)[0]
Hag_SC_non0 = Hag_SC_triu[non0]
Hag_FC_non0 = Hag_FC_triu[non0]
# Compute Pearson correlation coefficients between SC and FC.
pcc = np.corrcoef(Hag_SC_triu, Hag_FC_triu)[0, 1]
print('Correlation between Hagmann SC and FC:', round(pcc,2) )
pcc_non0 = np.corrcoef(Hag_SC_non0, Hag_FC_non0)[0, 1]
print('Correlation between Hagmann SC and FC (non-0 connections):', round(pcc_non0,2) )
fig = plt.figure(figsize=(12,5))
plt.subplot(121)
plt.scatter(Hag_SC_triu, Hag_FC_triu, c='b', alpha=.1)
plt.xlabel('SC'); plt.ylabel('FC'); plt.title('Upper Triangle')
plt.subplot(122)
plt.scatter(Hag_SC_non0, Hag_FC_non0, c='b', alpha=.1)
plt.xlabel('SC'); plt.ylabel('FC'); plt.title('Non-Zero Connections')
plt.show()
# + [markdown] colab_type="text" id="X3XHQlPGr3qV"
# ---
#
# <h2><font size="6" face="calibri" color="#609BC4">Computational model</font></h2>
# + [markdown] colab_type="text" id="Vu-uiUkdr3qW"
# <p><div style="text-align: justify"><font size="4.5" face="calibri">In this tutorial, we will use a computational model of resting-state network dynamics: the <b> dynamic mean field model</b>, previously introduced in <b>(Deco et al., 2013)</b>. The dynamic mean field approach involves approximating the average behaviour of an ensemble of neurons, instead of modeling interactions of individual neurons. This mean field model is a reduction of the model presented in <b>(Wong & Wang, 2006)</b> to a single population model, and is used in modeling studies of resting-state <b>(Deco et al., 2013; Hansen et al., 2015)</b>. The neural activity of each node is given by the following equations:</font></div></p>
#
# \begin{eqnarray}
# \dfrac{\text{d}S_{i}}{\text{d}t} &=& \dfrac{-S_{i}}{\tau_{s}} + \gamma \ (1 - S_{i})\ H(x_{i}) + \sigma\eta_{i}(t)\\
# &\\
# H(x_{i}) &=& \dfrac{ax_{i} - b}{1 - \exp(-d \ (ax_{i} - b))}\\
# &\\
# x_{i} &=& wJ_{N}S_{i} + J_{N}G\sum_{j}C_{ij}S_{j} + I_{0}
# \end{eqnarray}
#
# <br>
# <p><div style="text-align: justify"><font size="4.5" face="calibri">Below is a summary of the model parameters:</font></div></p>
# <br><br>
#
# | Variable | Definition |
# | :------------- |:-------------|
# | $S_{i}$ | average synaptic gating variable at the local area $i$ |
# | $H(x_{i})$ | sigmoid function that converts the input synaptic activity $x_{i}$ into an output population firing rate | |
# | $a = 0.270$ (nA.ms<sup>-1</sup>), $b = 0.108$ (kHz), $d = 154$ (ms) | parameters of the input-output function $H$ |
# | $w = 1.0$ | local excitatory recurrence |
# | $\gamma = 0.641$, $\tau=100$ (ms) | kinetic parameters|
# | $J_{N} = 0.2609$ (nA) | synaptic couplings
# | $I_0 = 0.3$ (nA) | overall effective external input |
# | $C_{ij}$ | entries of the anatomical SC matrix |
# | $G$ | global coupling (reweights the SC) |
# | $\eta_{i}(t)$ | Gaussian white noise |
# | $\sigma = 0.001$ | amplitude of Gaussian white noise |
#
# <br><br>
# <p><div style="text-align: justify"><font size="4.5" face="calibri">We will perform a parameter sweep of $G$ to study the optimal dynamical working region, where the simulated FC maximally fits the empirical FC.</font></div></p>
# <br><br>
# + [markdown] colab_type="text" id="fA07M4SNr3qX"
# <h3><font size="5" face="calibri" color="black">Exploring the model</font></h3>
#
# <p><div style="text-align: justify"><font size="4.5" face="calibri">First, we initialize the model, and display the default parameters.</font></div></p>
# + colab={"base_uri": "https://localhost:8080/", "height": 247} colab_type="code" id="hVzi8huXr3qY" outputId="6c82daf3-6378-41a1-d2ae-0ef2d27325df"
# Initialise the Model.
rww = models.ReducedWongWang()
HTML(rww._repr_html_() + "</table>") # fixes bug with nbconvert->HTML
# + [markdown] colab_type="text" id="2dhtC53dr3qc"
# <h3><font size="4" face="arial" color="black">Effects of the local excitatory recurrence</font></h3>
# + colab={} colab_type="code" id="cLLrvlo0r3qd"
# Initialize the state-variable S
S = np.linspace(0., 1., num=1000).reshape((1, -1, 1))
# Remember: the phase-flow only represents the dynamic behaviour of a disconnected node => SC = 0.
C = S*0.
# + colab={} colab_type="code" id="wFPpDc9tr3qg"
# Parameter sweep
W = np.linspace(0.6, 1.05, num=50)
# Fixed Io value
rww.I_o = __(0.33)
# +
# Visualize phase-flow for different values of w
# make colormap
import matplotlib.colors as mcolors
colors = plt.cm.plasma(np.linspace(0,255,np.shape(W)[0]+10).astype(int))
colors = colors[:-10,:]
mymap = mcolors.LinearSegmentedColormap.from_list('my_colormap', colors)
Z = [[0,0],[0,0]]
levels = np.linspace(min(W), max(W), 50)
CS3 = plt.contourf(Z, levels, cmap=mymap);
plt.clf();
fig = plt.figure(figsize=(12, 5))
for iw, w in enumerate(W):
rww.w = __(w)
dS = rww.dfun(S, C)
plt.plot(S.flat, dS.flat, color=colors[iw,:], alpha=0.5)
rww.w = np.array([1.0])
dS = rww.dfun(S, C)
plt.plot(S.flat, dS.flat, color='black', alpha=0.5)
plt.plot([0, 0] , '--',color='black',linewidth=.6)
plt.title('Phase flow for different values of $w$', fontsize=20)
plt.xlabel('S', fontsize=20); plt.xticks(fontsize=14)
plt.ylabel('dS', fontsize=20); plt.yticks(fontsize=14)
cb=plt.colorbar(CS3,shrink=0.5);
cb.set_label('w', fontsize=14)
plt.show()
# + [markdown] colab_type="text" id="9SGrloupr3qo"
# <h3><font size="4" face="arial" color="black">Effects of the external input</font></h3>
# + colab={} colab_type="code" id="Hg_57K90r3qq"
# Parameter sweep
Io = np.linspace(0.00, 0.40, num=50)
# Fixed w value at 1
rww.w = __(1.0)
# +
# Plot phase-flow for different Io values
rww.w = _(1.0)
colors = plt.cm.plasma(np.linspace(0,255,np.shape(Io)[0]+10).astype(int));
colors = colors[:-10,:];
mymap = mcolors.LinearSegmentedColormap.from_list('my_colormap', colors);
Z = [[0,0],[0,0]];
levels = np.linspace(min(Io), max(Io), 50);
CS3 = plt.contourf(Z, levels, cmap=mymap);
plt.clf();
fig = plt.figure(figsize=(12, 5))
for i, io in enumerate(Io):
rww.I_o = __(io)
dS = rww.dfun(S, C)
plt.plot(S.flat, dS.flat, c = colors[i,:], alpha=0.8, linewidth=.8)
plt.plot([0, 0] ,'--',color= 'black', linewidth=0.6)
rww.I_o = __(0.30); rww.w = __(0.9)
dS = rww.dfun(S, C)
plt.plot(S.flat, dS.flat, c = 'blue', label="Deco 2013: $I_o = 0.30$, $w=0.9$", linewidth=1.)
rww.I_o = __(0.32); rww.w = __(1.0);
dS = rww.dfun(S, C)
plt.plot(S.flat, dS.flat, c = 'green', label="Hansen 2015: $I_o = 0.32$, $w=1.0$", linewidth=1.)
plt.title('Phase flow for different values of $I_o$', fontsize=20)
plt.xlabel('S', fontsize=20); plt.xticks(fontsize=14)
plt.ylabel('dS', fontsize=20); plt.yticks(fontsize=14)
cb=plt.colorbar(CS3,shrink=0.5);
cb.set_label('$I_o$', fontsize=14)
plt.legend(fontsize=12)
plt.show()
zoomplot=False
if zoomplot:
fig = plt.figure(figsize=(12,3))
plt.subplot(121)
plt.plot(S.flat[0:350], dS.flat[0:350]);
plt.title('low')
plt.subplot(122)
plt.plot(S.flat[350:500],dS.flat[350:500]);
plt.title('high')
plt.show()
# + [markdown] colab_type="text" id="PPGRzwy9r3qx"
# <h3><font size="4" face="arial" color="black">Bifurcation diagram</font></h3>
# + [markdown] colab_type="text" id="KFleMp1gr3qz"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
#
# To identify the mechanisms underlying
# resting-state generation, we will first study how the dynamics of the model depends on the global coupling strength
# $G_{coupl}$, describing the scaling or global strength of the coupling between intercortical brain areas. In this case, we will study the fixed points of the local model dynamics in the absence of noise. To this end, we will calculate the <b>bifurcation diagram</b> characterizing the stationarity states of the brain system.</font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">We calculate firing rates $H(x)$ from synaptic activation variable $S$ returned by TVB, using the equation:</font></div></p>
#
# \begin{eqnarray}
# H(x_{i}) &=& \dfrac{ax_{i} - b}{1 - \exp(-d \ (ax_{i} - b))}\\
# &\\
# x_{i} &=& wJ_{N}S_{i} + J_{N}G\sum_{j}C_{ij}S_{j} + I_{0}
# \end{eqnarray}
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# We have to do this manually as TVB doesn't give these numbers by default, although it does calculate them on each integration step using the equation above.
# </font></div></p>
# + colab={} colab_type="code" id="Q0GNmkJFr3q3"
def run_rww_sim_bif(con, G, regime, D, dt, simlen, initconds):
# put regime dict vals into np arrays
regime = {k: __(v) for k,v in regime.items()}
# Initialise Simulator.
sim = simulator.Simulator(
model=MyRWW(**regime), #
connectivity=con, # SC weights matrix
coupling=coupling.Scaling(a=_(G)), # rescale connection strength
integrator=integrators.HeunDeterministic(dt=dt),
monitors=(monitors.TemporalAverage(period=1.),)
)
# Set initial conditions.
if initconds:
if initconds == 'low':
sim.initial_conditions = np.random.uniform(low=0.001, high=0.001, size=((1, 1, nregions, 1)))
elif initconds == 'high':
sim.initial_conditions = np.random.uniform(low=0.8, high=1.0, size=(1, 1, nregions, 1))
sim.configure()
# Launch simulation
H = []
for (t, y), in sim(simulation_length=simlen):
H.append(sim.model.H.copy())
H = np.array(H)
Hmax = np.max(H[14999, :])
return Hmax
# + colab={} colab_type="code" id="rYjdPyxtr3qz"
class MyRWW(models.ReducedWongWang):
def dfun(self, state, coupling, local_coupling=0.0):
# save the x and H value as attribute on object
S = state
c_0 = coupling
lc_0 = local_coupling * S
self.x = self.w * self.J_N * S + self.I_o + self.J_N * c_0 + self.J_N * lc_0
self.H = (self.a*self.x - self.b) / (1 - np.exp(-self.d*(self.a*self.x - self.b)))
# call the default implementation
return super(MyRWW, self).dfun(state, coupling, local_coupling=local_coupling)
# + [markdown] colab_type="text" id="_p72tWfsLHrI"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# The cell below shows you how to run the simulations. We will skip this cell for this tutorial; instead, we will load the results from these simulations.
# </font></div></p>
# +
# %%time
tic = tm.time()
regime = {'a': 270., 'b':108., 'd':0.154, 'gamma':0.641/1000, 'w':0.9, 'I_o':0.3}
# Run G sweep with short runs
Gs = np.arange(0., 3.1, 0.1)
Hmax_low = np.zeros((len(Gs)))
Hmax_high = np.zeros((len(Gs)))
for iG, G in enumerate(Gs):
Hmax_low[iG] = run_rww_sim_bif(Hag_con, __(Gs[iG]), regime, 0.001, 0.1, 15000,'low')
Hmax_high[iG] = run_rww_sim_bif(Hag_con, __(Gs[iG]), regime, 0.001, 0.1, 15000,'high')
#print('simulation required %0.f seconds.' % (tm.time()-tic))
# + [markdown] colab_type="text" id="gJPcpCLRMeXA"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# We will now plot the maximum firing rate activity among all nodesas a function of $G_{coupl}$, for low and high initial conditions.
# </font></div></p>
# +
# Load results
mat = loadmat('../data/sim_outputs/bifurcation_eMFM_Deco.mat')
Hmax_low = mat['Hmax_low'].T
Hmax_high = mat['Hmax_high'].T
Gs = np.arange(0.0, 3.1, 0.1)
# Visualization of the bifurcation diagram
plt.figure(figsize=(15, 5))
# plot low activity
plt.scatter(np.arange(31), Hmax_low, marker='o', facecolors = 'none', edgecolors = 'b',
s=55, linewidth = 1.5, label='low activity')
# plot high activity
plt.scatter(np.arange(31), Hmax_high, marker='o', facecolors = 'red', edgecolors='none',
s=50, label='high activity')
#plt.plot(np.arange(31), Hmax_high, 'ro', markeredgecolor='none', label='high activity')
plt.title('Bifurcation Diagram eMFM', fontsize=20)
plt.xticks(np.arange(len(Gs)), np.round(Gs,2))
plt.xlabel('$G_{coupl}$', fontsize=20); plt.ylabel('max H (spikes.s$^{-1}$)', fontsize=20)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
# critical points
plt.axvline(10, color='k', linestyle='--')
plt.axvline(26, color='k', linestyle='--')
plt.savefig('bifurcation_diagram.png')
plt.show()
#files.download('bifurcation_diagram.png')
# + [markdown] colab_type="text" id="mInr4EZEr3rG"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">The key feature that is shown in the bifurcation diagram, above, is the existence of <b>3 separate regimes</b>:</font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# <ul>
# <li>for <b>small values</b> of the global coupling $G_{coupl}$, only <b>one stable state</b> (i.e., spontaneous state) exists, characterizing by a low firing activity in all cortical areas, </li>
# <li>for a critical value of $G_{coupl}$ = $1.0$, a <b>first bifurcation</b> emerges where a new <b>multistable state</b> of <b>high activity</b> appears, while the state of low activity remains stable,</li>
# <li>for even larger values of $G_{coupl}$, a <b>second bifurcation</b> appears at $G_{coupl} = 2.6$, characterized by a loss of stability in the spontaneous state.</li></font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">In the following, we will seek to identify the parameter regimen in which the emergent model FC matches the empirical one.</font></div></p>
# + [markdown] colab_type="text" id="fs4jE59pr3rG"
# ---
#
# <h2><font size="6" color="#609BC4">Optimal working region of the model</font></h2>
# + [markdown] colab_type="text" id="YA9M_YPar3rH"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">To identify the region of the parameter <i>G</i> where the model best reproduces the empirical functional connectivity, we will convolve the simulated neuronal activity <i>S<sub>i</sub></i> with the <b>canonical hemodynamic response function</b> (implemented with a gamma kernel) with a sampling frequency of <b>0.5 Hz</b> using the <b>BOLD monitor</b> implemented in TVB. Then, we will compute the simulated functional connectivity by calculating the correlation matrix of the BOLD activity between all brain areas. We will then define the "fit" between the simulated and empirical functional connectivity as the Pearson correlation coefficient (PCC) between the simulated and empirical matrices.</font></div></p>
# + colab={} colab_type="code" id="Xuf2gO7kr3rI"
def run_rww_sim_pcc(con, G, regime, D, dt, simlen):
# put regime dict vals into np arrays
regime = {k: __(v) for k,v in regime.items()}
# Initialise Simulator.
sim = simulator.Simulator(
model=models.ReducedWongWang(**regime),
connectivity=con,
coupling=coupling.Scaling(a=__(G)),
integrator=integrators.HeunStochastic(dt=dt, noise=noise.Additive(nsig=__((D**2)/2))),
monitors=(monitors.Bold(period=2000.0),)
)
sim.initial_conditions = (0.001)*np.ones((1, 1, nregions, 1))
sim.configure()
# Launch simulation
res = sim.run(simulation_length=simlen)
(t,B) = res[0]
# Remove transient time
B = B[10:int(simlen/2000),:,:,:]
# Build a TimeSeries Datatype
tsr = TimeSeriesRegion(connectivity=con,
data=B,
sample_period=sim.monitors[0].period)
tsr.configure()
# Compute FC
FC = np.corrcoef(np.squeeze(tsr.data).T)
savemat('FC_' + str(G) + '_' + str(simlen) + '.mat', {'B': B, 'FC': FC})
# Take triangular upper part of connectivity matrices and compute pearson correlations
pcc_FC = np.corrcoef(np.triu(Hag_FC).ravel(), np.triu(FC).ravel())[0, 1]
pcc_SC = np.corrcoef(np.triu(Hag_SC).ravel(), np.triu(FC).ravel())[0, 1]
#return pcc
return pcc_FC, pcc_SC
# + [markdown] colab_type="text" id="ZC7SuddXTPso"
# Again, the below cell illustrates how to run the simulations, but we will skip this cell and load the results directly.
# +
# %%time
#tic = tm.time()
# Run G sweep
Gs = np.arange(0., 3.1, 0.1)
regime = {'a': 270., 'b':108., 'd':0.154, 'gamma':0.641/1000, 'w':1., 'I_o':0.30}
pcc_FC = np.zeros((len(Gs)))
pcc_SC = np.zeros((len(Gs)))
for iG, G in enumerate(Gs):
print(iG)
pcc_FC[iG], pcc_SC[iG] = run_rww_sim_pcc(Hag_con, Gs[iG], regime, 0.001, 0.1, 60000)
#60000 = 1min BOLD, 1230000 = 20.5min BOLD
#'simulation required %0.3f seconds.' % (tm.time()-tic)
# + colab={} colab_type="code" id="N22Ws759-j4l"
Gs = np.arange(0., 3.1, 0.1)
pcc_FC = np.zeros((len(Gs)))
pcc_SC = np.zeros((len(Gs)))
for iG, G in enumerate(Gs):
file2load = '../data/sim_outputs/FC_Deco2013_' + str(np.round(Gs[iG],2)) + '_1230000.mat'
tmp = loadmat(file2load)
B = tmp['B']
B = np.squeeze(B[15:,:,:,:])
FC_sim = np.corrcoef(B.T)
if np.isclose(Gs[iG],2.4):
FC_sim_best = FC_sim
inds = np.triu_indices(66,1)
pcc_FC[iG] = np.corrcoef(FC_sim[inds], Hag_FC[inds])[0,1]
pcc_SC[iG] = np.corrcoef(FC_sim[inds], Hag_SC[inds])[0,1]
# +
# Visualize
plt.figure(figsize=(12,6))
# FC
plt.plot(pcc_FC, '-*', label='FC - FC')
plt.xlabel('$G_{coupl}$', fontsize=20); plt.xticks(np.arange(len(Gs)), np.round(Gs,2))
plt.ylabel('PCC', fontsize=20)
# SC
plt.plot(pcc_SC, '-*g', label='SC - FC')
plt.xlabel('$G_{coupl}$', fontsize=20); #plt.xticks(np.arange(len(Gs)), Gs)
plt.ylabel('PCC', fontsize=20)
plt.title('Correlation Diagram', fontsize=20)
plt.axvline(26, color='k', linestyle='--')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.savefig('sim_emp_corr_diagram.png')
plt.show()
#files.download('sim_emp_corr_diagram.png')
# + [markdown] colab_type="text" id="xv76vz8sr3rQ"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
#
# So, the best fit (maximal correlation) occurs right before the <b>edge of the second bifurcation</b>, where the spontaneous state loses its stability. At this point, the noisy fluctuations of the dynamics are able to explore and reflect the structure of the other attractors that are shaped by the underlying anatomy.
#
# </font></div></p>
# + [markdown] colab_type="text" id="oz6kBVNur3rR"
# ---
#
# <h2><font size="6" color="#609BC4">Visualize FC model for the best working point</font></h2>
# + [markdown] colab_type="text" id="4u293cuwr3rS"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# Here, we will visualize the FC matrix obtained at the best-fit critical point of the model:
# </font></div></p>
# +
# Visualize the FC for the optimal G
plt.figure(figsize=(20,20))
# Hag_SC
plt.subplot(131)
plt.imshow(Hag_SC, interpolation='nearest', cmap='jet')
plt.title('Hag_SC', fontsize=20)
cb=plt.colorbar(shrink=0.23)
cb.set_label('weights', fontsize=15)
# Hag_FC
plt.subplot(132)
plt.imshow(Hag_FC, interpolation='nearest', cmap='jet')
plt.title('Hag_FC', fontsize=20)
cb=plt.colorbar(shrink=0.23, ticks=[-0.1, 0.5])
cb.set_label('PCC', fontsize=15)
plt.clim([-0.1, 0.5])
# FC model
plt.subplot(133)
plt.imshow(FC_sim_best, interpolation='nearest', cmap='jet')
plt.title('Model FC', fontsize=20)
cb=plt.colorbar(shrink=0.23, ticks=[-0.1, 0.5])
cb.set_label('PCC', fontsize=15)
plt.clim([-0.1, 0.5])
plt.show()
# +
# scatterplot of simulated and empirical FC matrices
inds = np.triu_indices(66,1)
fig = plt.figure(figsize=(8,5))
plt.scatter(FC_sim_best[inds], Hag_FC[inds],c='c', alpha=.3)
plt.xlabel('Simulated FC')
plt.ylabel('Empirical FC')
plt.show()
fig = plt.figure(figsize=(8,5))
plt.hist(FC_sim_best[inds],50, alpha=.2, color='#ff0000', label='Simulated FC')
plt.hist(Hag_FC[inds],50, alpha=.2, color='#0000ff', label='Empirical FC')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="BwObIrnhr3rc"
# <h3><font size="4" face="arial" color="black">SC-FC comparisons</font></h3>
# + [markdown] colab_type="text" id="FmHTR9Mcr3rd"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# We will plot the empirical SC matrix, the empirical FC and the model FC between one seed region and all other brain regions at the best operating point (i.e., at the edge of the second bifurcation). We take the <b>left posterior cingulate (lPC)</b> as a seed, which is part of the well-known default-mode network.
# </font></div></p>
# +
roi_ind = 43
print(Hag_con.region_labels[roi_ind])
print(np.corrcoef(Hag_FC[roi_ind, :], FC_sim_best[roi_ind,:])[1,0])
plt.figure(figsize=(10, 10))
plt.subplot(131)
plt.barh(np.arange(nregions), Hag_con.weights[roi_ind, :], align='center')
plt.title('SC', fontsize=15)
plt.xlabel('connection strength', fontsize=15)
plt.xticks([0., 0.05, 0.1])
plt.yticks(np.arange(nregions), Hag_con.region_labels, fontsize=7)
plt.subplot(132)
plt.barh(np.arange(nregions), Hag_FC[roi_ind, :], align='center')
plt.title('FC empirical', fontsize=15)
plt.xlabel('correlation coefficient', fontsize=15)
plt.xticks([-0.2, 0, 0.5])
plt.yticks(np.arange(nregions), Hag_con.region_labels, fontsize=7)
plt.subplot(133)
plt.barh(np.arange(nregions), FC_sim_best[roi_ind, :], align='center')
plt.title('FC model', fontsize=15)
plt.xlabel('correlation coefficient', fontsize=15)
plt.xticks([-0.2, 0, 0.5])
plt.yticks(np.arange(nregions), Hag_con.region_labels, fontsize=7)
plt.show()
# + [markdown] colab_type="text" id="mEm_XSIgr3rh"
# ---
#
# <h2><font size="6" color="#609BC4">Conclusions</font></h2>
# + [markdown] colab_type="text" id="NIY3ndYJr3ri"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# <br>
#
# We hope this has been a useful tutorial and welcome any comments or questions.
# </font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
#
# Further exploration:
#
# </font></div></p>
#
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# <ul>
# <li>Dynamics of Functional Connectivity?</li>
# <ul>
# <li>Can TVB reproduce FC dynamics?</li>
# <li>if yes, is the working region unchanged or not?</li>
# </ul>
# </ul>
# <ul>
# <li>Simulate a lesion?</li>
# <ul>
# <li>Effects of a lesion are not local and are difficult to predict without a simulation</li>
# <li>How long must be the time series to see it?</li>
# </ul>
# </ul>
#
# </font></div></p>
# + [markdown] colab_type="text" id="vuKQgdjJr3ri"
# ---
#
# <h1><font size="6" color="#609BC4">References</font></h1>
# + [markdown] colab_type="text" id="jClvEazFr3rj"
# <p><div style="text-align: justify"><font size="4.5" face="time roman">
# <blockquote>
#
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. (2008) <b>Mapping the structural core of human cerebral cortex.</b> PLoS Biol., 2008, 6, e159. <br />
#
# <br><NAME>., <NAME>., <NAME>., <NAME>. (2011)<b>Role of local network oscillations in resting-state network dynamics.</b> NeuroImage, 57(2011), 130-139.<br />
#
# <br><NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. <NAME>., <NAME>. (2006)<b>An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions on interest.</b> NeuroImage, 2011, 31(3), 968-980.<br />
#
# <br><NAME>. & Wang, X.-J. (2006)<b>A recurrent network mechanism of time integration in perceptual decision.</b> J. Neurosci., 2006, 26, 1314-1328. <br />
#
# <br><NAME>., <NAME>., <NAME>., <NAME>., <NAME>. & <NAME>. (2013)<b>Resting-state functional connectivity emerges from structurally and dynamically shaped slow linear fluctuations.</b> J. Neurosci., 32(27), 11239-11252, 2013.<br />
#
# <br><NAME>., <NAME>., <NAME>., <NAME>. & <NAME>. (2015)<b>Functional connectivity dynamics: modeling the switching behavior of the resting-state.</b> NeuroImage, 105(2015), 525-535.<br />
#
# </blockquote>
# </font></font></div></p>
#
#
| notebooks/modelling_resting_state.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img width="50" src="https://carbonplan-assets.s3.amazonaws.com/monogram/dark-small.png" style="margin-left:0px;margin-top:20px"/>
#
# # Biochar lifetime analysis
#
# _by <NAME> (CarbonPlan), Created May 17, 2020, Last Updated May 24,
# 2021_
#
# Here we present a simple toy model for evaluating the carbon removal and
# permanence of biochar projects. The data and analysis method is based directly
# on two publications
#
# - Spokas (2010) Review of the stability of biochar in soils: predictability of
# O:C molar ratios, Carbon Management, doi: 10.4155/CMT.10.32
#
# - Campbell et al. (2018) Potential carbon storage in biochar made from logging
# residue: Basic principles and Southern Oregon case studies, PLOS One, doi:
# 10.1371/journal.pone.0203475
#
# ### Notebook setup
#
# +
# %matplotlib inline
import logging
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import statsmodels.api as sm
from carbonplan_styles.mpl import set_theme
from carbonplan_styles.colors import colors
set_theme(style='carbonplan_light')
c = colors('carbonplan_light')
# -
# ### The basic model
#
# Campbell et al. (2018) present a simple model for biochar carbon dynamics by
# comparing the carbon content of biomass after biocharing to the carbon content
# that would have resided in the form of the source feedstock (e.g. logging
# residues).
#
# The difference is
#
# $∆ = C_{biochar} - C_{biochar}$
#
# And the mass of carbon in both is modeled using a first-order differential
# equation
#
# $C_t = C_{t-1}e^{-k} + C_{input}$
#
# We'll write a function that generates a complete carbon curve as a function of
# the input and the parameter k over 1000 years
#
def model(t, initial, k):
return initial * np.exp(-k * t)
# And we can now plot carbon curves for both unmodified residue and biochar over a
# fixed duration, assuming an initial carbon content of 20 tC for the residue and
# 12 tC for the biochar (which would be achieved through a pyrolysis process with
# 60% efficiency).
#
t = np.arange(1000)
residue = model(t, 20, 0.03)
biochar = model(t, 12, 0.003)
plt.plot(t, residue)
plt.plot(t, biochar)
plt.xlim([0, 200])
plt.ylim([0, 20])
# This precisely matches Figure 1A from Campbell et al. (2019)
#
# These curves makes clear that biochar is not removing carbon per se, but rather
# avoiding the emissions that would have been associated with the corresponding
# feedstock. For that reason, the appropriate quantity is the difference between
# the two curves.
#
plt.plot(t, biochar - residue)
plt.xlim([0, 200])
plt.ylim([-10, 10])
plt.hlines(0, 0, 200, color=c["secondary"])
# As this curve makes clear, the cumulative effective carbon removal is initially
# negative, quickly reaches a compensation point, and then reaches a point termed
# by Campbell et al. (2018) as "climate parity" where the storage
#
# ### Mapping O:C ratios to half life
#
# A key parameter in the above model is the decay rate (also referred to as the
# biochar's recalcitrance). Campbell et al. (2018) find that this parameter has
# little effect on the time at which climate parity is achieved, so long as it 10
# times greater than the decay rate feedstock. But it is also importantly related
# to the permanence, or time scale over which the carbon stored in the biochar
# remain.
#
# We can use data digitized from a meta-analysis by Spokas (2010) that relate the
# oxygen to carbon (O:C) molar ratio to the predicted half-life of synthetic
# biochar in various laboratory conditions.
#
import pandas as pd
import numpy as np
data = pd.read_csv("biochar.csv")
plt.plot(data.ratio, data.halflife, ".", color=c["primary"])
plt.xlim([0, 0.8])
plt.ylim([1, 10 ** 8])
plt.yscale("log")
# We fit a simple linear model in log space so we can predict half-life as a
# function of ratio. In order to put bounds on our estimates, we use a simple
# bootstrap to fit the model for each of 1000 random samples (with replacement)
# from the data. We store the parameter estimates from each sample, and plot a
# regression line.
#
k = 10000
plt.plot(data.ratio, data.halflife, ".", color=c["primary"])
xhat = np.arange(0, 1, 0.1)
indices = np.arange(34)
alpha = np.zeros(k)
beta = np.zeros(k)
for i in range(k):
samples = np.random.choice(indices, 34)
mod = sm.OLS(
np.log(data.halflife[samples]),
sm.add_constant(data.ratio[samples], prepend=False),
)
res = mod.fit()
alpha[i] = res.params[1]
beta[i] = res.params[0]
yhat = res.predict(sm.add_constant(xhat, prepend=False))
if i % 10 == 0:
plt.plot(xhat, np.exp(yhat), "-", color="red", alpha=0.005)
plt.yscale("log")
plt.xlim([0, 0.8])
plt.ylim([1, 10 ** 8])
# Finally we write a simple function that, for a given ratio, returns a prediction
# from the bootstrapped distribution at a given percentile.
#
def predict(ratio, prctile):
dist = np.exp(alpha + beta * ratio)
return np.percentile(dist, [prctile])[0]
# ## Project evaluation
#
# ### Fixed fraction permanence
# We can now use the above to evaluate some aspects of a biochar projects. If we
# assume a project reports an O:C ratio of 0.08, we can use the simple linear
# model above to we can compute a half-life. We use the 2.5th percentile of the
# posterior predictive distribution as a crude, highly conservative estimate,
# given that permanence is only weakly correlated with composition, and likely
# depends as much or more so on the decay environment, which is often unknown.
#
ratio = 0.09
halflife = predict(ratio, 2.5)
# Still, given the decay kinetics assumed by our toy model, we can compute a decay
# constant from the half-life
#
k = np.log(2) / halflife
# We can now determine the duration after which a fixed percent of the biochar
# remains. For a target of 90% for example, we get the following number of years.
#
fraction = 0.9
years = -np.log(fraction) / k
# We can summarize our parameters
#
print("summary")
print("-------")
print("ratio: " + str(ratio))
print("half-life: " + str(halflife) + " years")
print("fraction: " + str(fraction))
print("k: " + str(k))
print("years: " + str(years))
# And we can plot this on the decay curve from above, assuming a initial volume of carbon storage in the biochar (tC).
#
initial = 100
t = np.arange(0, 20000)
biochar = model(t, initial, k)
plt.plot(t, biochar)
plt.ylim([0, initial])
plt.xlim([0, 2000])
plt.vlines(years, 0, initial)
plt.hlines(initial * fraction, 0, 20000, color=c["secondary"])
# In general, validating the volume and permanence for an actual biochar project
# requires knowing the composition (and thus recalcitrance), but perhaps more
# importantly, also requires knowing the conversion efficiency (the fraction of
# initial feedstock carbon retained in biochar after pyrolysis) and the decay rate
# of the feedstock. That said, simply by knowing the recalcitrance, and making
# some assumptions, we can approximate a permanence over which a fixed fraction of
# volume is likely to remain.
#
# ### Counterfactual feedstock decay
# Per the Campell et al. (2018), biochar acheives carbon storage by decaying more slowly than its feedstock.
#
# Using the approximate permanence horizon calculated in the section above, we can ask how quickly the feedstock would have had to decay for the counterfactual carbon storage in the feedstock to be considered negligible.
# Assuming a pyrolysis efficiency (e.g. 60%), we can estimate the starting carbon storage of the feedstock relative to the biochar.
#
efficiency = 0.6
feedstock_start = initial/efficiency
# By setting a bar for "negligible impact" (e.g. feedstock carbon storage must be <0.5% of biochar carbon storage at the end of the permanence period), we can calculate an upper bound for feedstock carbon storage.
negligible = 0.005
feedstock_end = (initial*fraction) * negligible
# We can now determine a minimum decay constant for a feedstock's counterfactual carbon storage to be considered negligible. (As a reminder, lower decay constant means slower decay!)
k_feedstock = -np.log(feedstock_end/feedstock_start) / years
# We can plot the feedstock and biochar decay curves over the permanence period calculated above.
#
t = np.arange(0, years)
biochar = model(t, initial, k)
feedstock = model(t, feedstock_start, k_feedstock)
plt.plot(t, biochar)
plt.plot(t, feedstock)
plt.ylim([0, initial/efficiency])
plt.xlim([0, years])
plt.vlines(years, 0, initial*fraction, color=c["secondary"])
plt.hlines(initial * 0.90, 0, 20000, color=c["secondary"])
# The net carbon storage at the end of the permanence period is the difference between the mass of biochar carbon storage year and the counterfactual feedstock carbon storage.
NCS = (biochar - feedstock)[years.astype(int)]
print(str(np.round(NCS)) + " tC")
# We can summarize our parameters and outputs:
print("summary")
print("-------")
print("years: " + str(np.round(years)) + " years")
print("efficiency: " + str(efficiency*100) + "%")
print("negligible impact threshold: " + str(negligible*100) + '%')
print("min feedstock k: " + str(np.round(k_feedstock,3)))
# We can compare this feedstock decay rate bound against values found in literature to gain intuition about how important it is to take into account the feedstock counterfactual when crediting biochar.
#
# Publications we have queried for this information include:
#
# - Harmon et al. (2020) Release of coarse woody detritus-related carbon: a synthesis across forest biomes, Carbon Balance Management, doi: 10.1186/s13021-019-0136-6
#
# - Ximenes et al. (2017) The decay of engineered wood products and paper excavated from landfills in Australia, Waste Management, doi: 10.1016/j.wasman.2017.11.035
#
| biochar/biochar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from chembl_webresource_client.new_client import new_client
# +
def chembl_search(search_string):
target = new_client.target
target_query = target.search(search_string)
targets = pd.DataFrame.from_dict(target_query)
return targets
def get_bioactivity(target_chembl_id,standard_type):
activity = new_client.activity
res = activity.filter(target_chembl_id = target_chembl_id).filter(standard_type = standard_type)
df = pd.DataFrame.from_dict(res)
df = df[df.standard_value.notna()]
return df
def main_pipe(target_chembl_id,standard_type):
df = get_bioactivity(target_id,standard_type)
df['bioactivity_inactive'] = df['standard_value'].astype('float')>=10000
df['bioactivity_active'] = df['standard_value'].astype('float')<1000
df['bioactivity_intermediate'] = np.logical_not(np.logical_or(df['bioactivity_inactive'],df['bioactivity_active']))
df = df[['molecule_chembl_id',
'canonical_smiles',
'standard_value',
'bioactivity_inactive',
'bioactivity_active',
'bioactivity_intermediate']]
return df
# -
target_id = 'CHEMBL3927'
standard_type = 'IC50'
main_pipe(target_id,standard_type)
| notebooks/data_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py2env)
# language: python
# name: py2env
# ---
# # McKinsey Data Scientist Hackathon
#
# link: https://datahack.analyticsvidhya.com/contest/mckinsey-analytics-online-hackathon-recommendation/?utm_source=sendinblue&utm_campaign=Download_The_Dataset_McKinsey_Analytics_Online_Hackathon__Recommendation_Design_is_now_Live&utm_medium=email
#
# slack:https://analyticsvidhya.slack.com/messages/C8X88UJ5P/
#
#
# ## Problem Statement ##
#
# Your client is a fast-growing mobile platform, for hosting coding challenges. They have a unique business model, where they crowdsource problems from various creators(authors). These authors create the problem and release it on the client's platform. The users then select the challenges they want to solve. The authors make money based on the level of difficulty of their problems and how many users take up their challenge.
#
# The client, on the other hand makes money when the users can find challenges of their interest and continue to stay on the platform. Till date, the client has relied on its domain expertise, user interface and experience with user behaviour to suggest the problems a user might be interested in. You have now been appointed as the data scientist who needs to come up with the algorithm to keep the users engaged on the platform.
# The client has provided you with history of last 10 challenges the user has solved, and you need to predict which might be the next 3 challenges the user might be interested to solve. Apply your data science skills to help the client make a big mark in their user engagements/revenue.
#
# ### Data Relationships
# Client: problem platform maintainer
# Creators: problem contributors
# Users: people who solve these problems
#
# Question? Given the 10 challenges the user solved, what might be the next 3 challenges user want to solve?
# ## Now let's first look at some raw data
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import sklearn
import pandas
import seaborn
x_data = pandas.read_csv('./train_mddNHeX/train.csv')
y_data = pandas.read_csv('./train_mddNHeX/challenge_data.csv')
x_test = pandas.read_csv('./test.csv')
y_sub_temp = pandas.read_csv('./sample_submission_J0OjXLi_DDt3uQN.csv')
print('shape of submission data = {}, number of users = {}'.format(y_sub_temp.shape, y_sub_temp.shape[0]/13))
y_sub_temp.head()
print('shape of user data = {}, number of users = {}'.format(x_data.shape, x_data.shape[0]/13))
#x_data.sort_values('user_id')
x_data[0:20]
print('shape of user test data = {}, number of users = {}'.format(x_test.shape, x_test.shape[0]/10))
#x_test[0:15]
x_test.head(15)
#x_test.sort_values('user_id').head(15)
print('shape of challenge data = {}'.format(y_data.shape))
y_data[0:10]#.tail()
#print(y_data.loc[:,['challenge_ID','challenge_series_ID']])
#print(y_data.groupby('challenge_series_ID'))
# ## Dirty try
# 1. Need to find a feature vector for a given challenge
# - This is associated with [prog_lang, challenge_series, total submission, publish_time, auth_id, auth_org, categ]
# 2. Create a preference vector for each user
# - This will be randomly initialized
# 3. Use the first 10 samples from each users as ground truth for training the feature vector and the preference vector
# ## Prepare training data
# Let's prepare the challange id as a lookup table to constuct training data
#
def str2ascii(astr):
"""
input:
astr: a string
output:
val: a number which is sum of char's ascii.
"""
val = 0
real = 0
count_val, count_real = 0, 0
for i in list(astr):
num = ord(i)
if 48<= num and num <= 57:
real = real*10 + int(i)
count_real += 1
else:
val += num
count_val += 1
val = val*10**count_real + real
return val
# Retain the original copy of the y_data
ch_table = y_data
orig_y_data = y_data.copy()
print(ch_table.columns)
## Fill NaN with some values
values = {'challenge_series_ID':'SI0000','author_ID':'AI000000','author_gender':'I'
,'author_org_ID':'AOI000000', 'category_id':0.0
,'programming_language':0,'total_submissions':0, 'publish_date':'00-00-0000'}
ch_table = y_data.fillna(value = values)
print(y_data.head(), ch_table.head())
ch_table.iloc[3996]
## Change strings to some encoded values
columns = ['challenge_series_ID','author_ID','author_gender','author_org_ID','publish_date']
#print(ch_table[0:10])
for col in columns:
print(col)
#ch_table[col] = ch_table.apply(lambda x: str2ascii(x[col]),axis=1)
ch_table[col] = ch_table[col].apply(lambda x: str2ascii(x))
ch_table[0:10]
y_data['programming_language'].describe()
# ### Now, we need to normalize the table
## using normalizer
from sklearn import preprocessing
normalizer = preprocessing.Normalizer()
min_max_scaler = preprocessing.MinMaxScaler()
## Decrease the variance between data points in each columns
columns = ch_table.columns
#print(columns[1:],ch_table.loc[:,columns[1:]])
ch_table.loc[:,columns[1:]].head()
minmax_ch_table = min_max_scaler.fit_transform(ch_table.loc[:,columns[1:]])
norm_ch_table = preprocessing.normalize(ch_table.loc[:,columns[1:]],norm='l2')
#ch_table.loc[:,columns[1:]] = norm_ch_table
#ch_table.head()
print(pandas.DataFrame(minmax_ch_table, columns=columns[1:]).head(2))
print(pandas.DataFrame(norm_ch_table, columns=columns[1:]).head(2))
## Finally put the scaled data back
ch_table[columns[1:]] = minmax_ch_table
ch_table.head(10)
# ## Great!, now we have feature vectors for every challenges
#
# Next lets prepare the ground truth matrix for users
#
# Shape of y = (n_c, n_u)
#
# 1. n_c: the number of challenges
# 2. n_u: the number of users
## The ch_features contains
ch_features = ch_table.sort_values('challenge_ID')
ch_features = ch_features.loc[:,columns[1:]].values
#ch_features.head(10)
print('Shape of feature (n_c, n_f) = {}'.format(ch_features.shape))
## Setting up the lookup table
ch_lookup = {}
tmp = ch_table['challenge_ID'].to_dict()
ch_id_lookup=tmp
#for key in tmp.keys
for key in tmp.keys():
#print(key, tmp[key])
ch_lookup[tmp[key]] = key
#ch_lookup
## now lets set up a training y array with shape = (n_c, n_u)
def findChallengeFeatures(challenge_id, table, lookup):
"""
input:
challenge_id: a string of the challenge_id
table: pandas dataframe lookup table
output:
features: numpy array of features
"""
columns = table.columns
return table.loc[lookup[challenge_id], columns[1:]]
# %%time
ch_table.head()
featureVec = findChallengeFeatures(x_data.loc[0,'challenge'],ch_table, ch_lookup)
print(featureVec.shape)
# %%time
from operator import itemgetter
#myvalues = itemgetter(*mykeys)(mydict)
columns = ch_table.columns.values
usr_table = x_data
print(columns[1:])
for i in columns[1:]:\
usr_table[i] = np.nan
nSamples = x_data.shape[0]
## Finding indices
indices = np.array([ch_lookup[i] for i in x_data.loc[:nSamples-1,'challenge']])
print(indices.shape)
usr_table.loc[:nSamples-1, columns[1:]] = ch_table.loc[indices, columns[1:]].values
#print(ch_table.loc[indices,columns[1:3]])
#print(usr_table.loc[:nSamples-1, columns[1:]].shape)
usr_table.head(15)
usr_table.to_csv('train_withFeatureVec_allsamples.csv')
ch_table.to_csv('challenge_featureVecTable_allsamples.csv')
# ## Let's prepare the labels
#
# First, we need an empyty array to hold challenges
ch_emptyVec = np.zeros((ch_table.shape[0]))
ch_emptyVec.shape
x_data.head(13)
# +
## constructing a (n_u, n_c) array for nSamples
# %%time
columns = ch_table.columns
nSamples = int(x_data.shape[0])
x_train = np.zeros((nSamples, 10, len(ch_table.columns)-1)) ## (m, n_i, n_f)
y_train = np.zeros((nSamples, ch_table.shape[0])) ## (m, n_c)
for i in range(nSamples/13):
curpt = i*13
#print(i)
x_train[i] = x_data.loc[curpt:(curpt+9), columns[1:]] ## 0-10, 13-26
#print(x_train[i].shape)
#y_train[i] = ch_emptyVec
#tmp = x_data.loc[(curpt+10):(curpt+12), 'challenge'].values
#tmp = [ch_lookup[tmp[0]],ch_lookup[tmp[1]],ch_lookup[tmp[2]]]
#y_train[i,tmp] = 1 ## 10-13, 26-29
indices = [int(ch_lookup[j]) for j in x_data.loc[(curpt+10):(curpt+12), 'challenge']]
#print(indices, np.ones(3), tmp)
y_train[i, indices] = 1 ## 10-13, 26-29
#break
print('x_train shape = {}, y_train shape = {}'.format(x_train.shape, y_train.shape))
# -
## Flatten the array
x_train = x_train.reshape((x_data.shape[0],-1))
# ## Finally lets dunmp it into a classifier
from sklearn.naive_bayes import GaussianNB
from sklearn import tree
gnb = GaussianNB()
clf = tree.DecisionTreeClassifier()
clf.fit(x_train, y_train)
# +
## A simple NN
from keras.models import Sequential
from keras.layers import Dense, Activation
model = Sequential()
model.add(Dense(64, input_dim=80))
model.add(Activation('relu'))
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dense(1024))
model.add(Activation('relu'))
model.add(Dense(y_train.shape[1]))
model.add(Activation('softmax'))
# -
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=1)
model.save_weights('simpleNN.h5')
#
# ## Running out of time just gonna plug it in and submit
# %%time
nSamples = x_test.shape[0]
columns = ch_table.columns
test_table = x_test
print(columns[1:])
for i in columns[1:]:
test_table[i] = np.nan
indices = np.array([ch_lookup[i] for i in x_test.loc[:nSamples-1,'challenge']])
test_table.loc[:nSamples-1, columns[1:]] = ch_table.loc[indices, columns[1:]].values
print(indices.shape)
# %%time
test_table.to_csv('prepared_test_table_for_prediction.csv')
x_submit = np.zeros((nSamples/10, 10, len(ch_table.columns)-1)) ## (m, n_i, n_f)
y_submit = pandas.DataFrame(columns=['user_sequence','challenge'],
data = np.empty((x_test.shape[0]/10*3,2), dtype=np.str))
#y_submit['user_sequence']
#y_submit.head(15)
# %%time
for i in range(nSamples/10):
curpt = i*10
#print(i)
x_submit[i] = x_test.loc[curpt:(curpt+9), columns[1:]] ## 0-10, 13-26
pred = model.predict(x_submit[i].reshape((1,80)))
ids = np.argsort(pred.reshape(-1))[-3:]
#print(pred, ids, ids.shape)
#print(pred[0,ids])
outpt = i*3
user_id = x_test.loc[curpt,'user_id']
y_submit.iloc[outpt:outpt+3,:] = [[str(user_id)+'_11', ch_id_lookup[ids[0]]],
[str(user_id)+'_12', ch_id_lookup[ids[1]]],
[str(user_id)+'_13', ch_id_lookup[ids[2]]]
]
#print(y_submit.iloc[outpt:outpt+3,:])
y_submit.head()
y_submit.head(15)
y_submit.to_csv('ftl_submission.csv')
| McKinsey-Hackathon/20180310_McKinseyDSHackathon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Submit Training Job Demo
import fml_manager
import json
import time
import requests
import os
# The DSL and Config file can be presented in JOSN format.
# Submitting a job with JSON defined in line, can use ```submit_job(self,dsl, config)```.
# Note: the parameters are dict, the JSON string have to transform to dict with ```json.loads```
# +
dsl='''
{
"components" : {
"secure_add_example_0": {
"module": "SecureAddExample"
}
}
}
'''
config='''
{
"initiator": {
"role": "guest",
"party_id": 9999
},
"job_parameters": {
"work_mode": 1
},
"role": {
"guest": [
9999
],
"host": [
9999
]
},
"role_parameters": {
"guest": {
"secure_add_example_0": {
"seed": [
123
]
}
},
"host": {
"secure_add_example_0": {
"seed": [
321
]
}
}
},
"algorithm_parameters": {
"secure_add_example_0": {
"partition": 10,
"data_num": 1000
}
}
}
'''
manager = fml_manager.FMLManager()
response = manager.submit_job(json.loads(dsl), json.loads(config))
# -
manager.prettify(response, True)
stdout = json.loads(response.content)
jobid = stdout["jobId"]
# Once the job submitted, we can use ```def query_job(self, query_conditions)``` to query the status. the query_condition is a dict, and can add all job's attributes for querying.
query_condition = {
"job_id":jobid
}
job_status = manager.query_job(query_condition)
manager.prettify(job_status, True)
manager.query_job_status(query_condition)
# We can also fetch the logs of job submitted, and save it to working folder.
response = manager.fetch_job_log(jobid)
| fml_manager/Examples/Toy_Example/toy_example_submit_job.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (tensorflow)
# language: python
# name: tensorflow
# ---
# # T81-558: Applications of Deep Neural Networks
# **Module 14: Other Neural Network Techniques**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 14 Video Material
#
# * **Part 14.1: What is AutoML** [[Video]](https://www.youtube.com/watch?v=TFUysIR5AB0&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_14_01_automl.ipynb)
# * Part 14.2: Using Denoising AutoEncoders in Keras [[Video]](https://www.youtube.com/watch?v=4bTSu6_fucc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_14_02_auto_encode.ipynb)
# * Part 14.3: Training an Intrusion Detection System with KDD99 [[Video]](https://www.youtube.com/watch?v=1ySn6h2A68I&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_14_03_anomaly.ipynb)
# * Part 14.4: Anomaly Detection in Keras [[Video]](https://www.youtube.com/watch?v=VgyKQ5MTDFc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_14_04_ids_kdd99.ipynb)
# * Part 14.5: The Deep Learning Technologies I am Excited About [[Video]]() [[Notebook]](t81_558_class_14_05_new_tech.ipynb)
#
#
# # Part 14.1: What is AutoML
#
# Automatic Machine Learning (AutoML) attempts to use machine learning to automate itself. Data is passed to the AutoML application in raw form and models are automatically generated.
#
# ### AutoML from your Local Computer
#
# The following AutoML applications are commercial.
#
# * [Rapid Miner](https://rapidminer.com/educational-program/) - Free student version available.
# * [Dataiku](https://www.dataiku.com/dss/editions/) - Free community version available.
# * [DataRobot](https://www.datarobot.com/) - Commercial
# * [H2O Driverless](https://www.h2o.ai/products/h2o-driverless-ai/) - Commercial
#
# ### AutoML from Google Cloud
#
# * [Google Cloud AutoML Tutorial](https://cloud.google.com/vision/automl/docs/tutorial)
#
#
# ### A Simple AutoML System
#
# The following program is a very simple implementation of AutoML. It is able to take RAW tabular data and construct a neural network.
#
# We begin by defining a class that abstracts the differences between reading CSV over local file system or HTTP/HTTPS.
# +
import requests
import csv
class CSVSource():
def __init__(self, filename):
self.filename = filename
def __enter__(self):
if self.filename.lower().startswith("https:") or \
self.filename.lower().startswith("https:"):
r = requests.get(self.filename, stream=True)
self.infile = (line.decode('utf-8') for line in r.iter_lines())
return csv.reader(self.infile)
else:
self.infile = codecs.open(self.filename, "r", "utf-8")
return csv.reader(self.infile)
def __exit__(self, type, value, traceback):
self.infile.close()
# -
# The following code analyzes the tabular data and determines a way of encoding the feature vector.
# +
import csv
import codecs
import math
import os
import re
from numpy import genfromtxt
MAX_UNIQUES = 200
INPUT_ENCODING = 'latin-1'
CMD_CAT_DUMMY = 'dummy-cat'
CMD_CAT_NUMERIC = 'numeric-cat'
CMD_IGNORE = 'ignore'
CMD_MAP = 'map'
CMD_PASS = 'pass'
CMD_BITS = 'bits'
CONTROL_INDEX = 'index'
CONTROL_NAME = 'name'
CONTROL_COMMAND = 'command'
CONTROL_TYPE = 'type'
CONTROL_LENGTH = 'length'
CONTROL_UNIQUE_COUNT = 'unique_count'
CONTROL_UNIQUE_LIST = 'unique_list'
CONTROL_MISSING = 'missing'
CONTROL_MEAN = 'mean'
CONTROL_SDEV = 'sdev'
MAP_SKIP = True
MISSING_SKIP = False
current_row = 0
def is_number(s):
try:
float(s)
return True
except ValueError:
return False
def isna(s):
return s.upper() == 'NA' or s.upper() == 'N/A' \
or s.upper() == 'NULL' or len(s) < 1 or s.upper() == '?'
def analyze(filename):
fields = []
first_header = None
# Pass 1 (very short. First, look at the first row of each of the
# provided files.
# Build field blocks from the first file, and ensure that other files
# match the first one.
with CSVSource(filename) as reader:
header = next(reader)
if first_header is None:
first_header = header
for idx, field_name in enumerate(header):
fields.append({
'name': field_name,
'command': '?',
'index': idx,
'type': None,
'missing': False,
'unique': {},
'count': 0,
'mean': '',
'sum': 0,
'sdev': '',
'length': 0})
else:
for x, y in zip(header, first_header):
if x != y:
raise ValueError(\
'The headers do not match on the input files')
# Pass 2 over the files
with CSVSource(filename) as reader:
next(reader)
# Determine types and calculate sum
for row in reader:
if len(row) != len(fields):
continue
for data, field_info in zip(row, fields):
data = data.strip()
field_info['length'] = max(len(data),field_info['length'])
if len(data) < 1 or data.upper() == 'NULL' or isna(data):
field_info[CONTROL_MISSING] = True
else:
if not is_number(data):
field_info['type'] = 'text'
# Track the unique values and counts per unique item
cat_map = field_info['unique']
if data in cat_map:
cat_map[data]['count']+=1
else:
cat_map[data] = {'name':data,'count':1}
if field_info['type'] != 'text':
field_info['count'] += 1
field_info['sum'] += float(data)
# Finalize types
for field in fields:
if field['type'] is None:
field['type'] = 'numeric'
field[CONTROL_UNIQUE_COUNT] = len(field['unique'])
# Calculate mean
for field in fields:
if field['type'] == 'numeric' and field['count'] > 0:
field['mean'] = field['sum'] / field['count']
# Pass 3 over the files, calculate standard deviation and
# finailize fields.
sums = [0] * len(fields)
with CSVSource(filename) as reader:
next(reader)
for row in reader:
if len(row) != len(fields):
continue
for data, field_info in zip(row, fields):
data = data.strip()
if field_info['type'] == 'numeric' \
and len(data) > 0 and not isna(data):
sums[field_info['index']] += (float(data) - \
field_info['mean']) ** 2
# Examine fields
for idx, field in enumerate(fields):
if field['type'] == 'numeric' and field['count'] > 0:
field['sdev'] = math.sqrt(sums[field['index']] / field['count'])
# Assign a default command
if field['name'] == 'ID' or field['name'] == 'FOLD':
field['command'] = 'pass'
elif "DATE" in field['name'].upper():
field['command'] = 'date'
elif field['unique_count'] == 2 and field['type'] == 'numeric':
field['command'] = CMD_PASS
elif field['type'] == 'numeric' and field['unique_count'] < 25:
field['command'] = CMD_CAT_DUMMY
elif field['type'] == 'numeric':
field['command'] = 'zscore'
elif field['type'] == 'text' and field['unique_count'] \
<= MAX_UNIQUES:
field['command'] = CMD_CAT_DUMMY
else:
field['command'] = CMD_IGNORE
return fields
def write_control_file(filename, fields):
with codecs.open(filename, "w", "utf-8") as outfile:
writer = csv.writer(outfile,quoting=csv.QUOTE_NONNUMERIC)
writer.writerow([CONTROL_INDEX, CONTROL_NAME, CONTROL_COMMAND,
CONTROL_TYPE, CONTROL_LENGTH, CONTROL_UNIQUE_COUNT,
CONTROL_MISSING, CONTROL_MEAN, CONTROL_SDEV])
for field in fields:
# Write the main row for the field (left-justified)
writer.writerow([field[CONTROL_INDEX], field[CONTROL_NAME],
field[CONTROL_COMMAND], field[CONTROL_TYPE],
field[CONTROL_LENGTH],
field[CONTROL_UNIQUE_COUNT],
field[CONTROL_MISSING], field[CONTROL_MEAN],
field[CONTROL_SDEV]])
# Write out any needed category information
if field[CONTROL_UNIQUE_COUNT] <= MAX_UNIQUES:
sorted_cat = field['unique'].values()
sorted_cat = sorted(sorted_cat, key=lambda k:
k[CONTROL_NAME])
for category in sorted_cat:
writer.writerow(["","",
category[CONTROL_NAME],
category['count']])
else:
catagories = ""
def read_control_file(filename):
with codecs.open(filename, "r", "utf-8") as infile:
reader = csv.reader(infile)
header = next(reader)
lookup = {}
for i, name in enumerate(header):
lookup[name] = i
fields = []
categories = {}
for row in reader:
if row[0] == '':
name = row[2]
mp = '' if len(row)<=4 else row[4]
categories[name] = {'name':name,'count':int(row[3]),
'map':mp}
if len(categories)>0:
field[CONTROL_UNIQUE_LIST] = \
sorted(categories.keys())
else:
# New field
field = {}
categories = {}
field['unique'] = categories
for key in lookup.keys():
value = row[lookup[key]]
if key in ['unique_count', 'count', 'index', 'length']:
value = int(value)
elif key in ['sdev', 'mean', 'sum']:
if len(value) > 0:
value = float(value)
field[key] = value
field['len'] = -1
fields.append(field)
return fields
def header_cat_dummy(field, header):
name = str(field['name'])
for c in field['unique']:
dname = "{}-D:{}".format(name, c)
header.append(dname)
def header_bits(field, header):
for i in range(field['length']):
header.append("{}-B:{}".format(field['name'], i))
def header_other(field, header):
header.append(field['name'])
def column_zscore(field,write_row,value,has_na):
if isna(value) or field['sdev'] == 0:
#write_row.append('NA')
#has_na = True
write_row.append(0)
elif not is_number(value):
raise ValueError("Row {}: Non-numeric for zscore: {}"\
" on field {}".format(current_row,value,field['name']))
else:
value = (float(value) - field['mean']) / field['sdev']
write_row.append(value)
return has_na
def column_cat_numeric(field,write_row,value,has_na):
if CONTROL_UNIQUE_LIST not in field:
raise ValueError("No value list, can't encode {}"\
" to numeric categorical.".format(field[CONTROL_NAME]))
if value not in field[CONTROL_UNIQUE_LIST]:
write_row.append("NA")
has_na = True
else:
idx = field[CONTROL_UNIQUE_LIST].index(value)
write_row.append('class-' + str(idx))
return has_na
def column_map(field,write_row,value,has_na):
if value in field['unique']:
mapping = field['unique'][value]['map']
write_row.append(mapping)
else:
write_row.append("NA")
return True
return has_na
def column_cat_dummy(field,write_row,value,has_na):
for c in field['unique']:
write_row.append(0 if value != c else 1)
return has_na
def column_bits(field,write_row,value,has_na):
if len(value)!=field['length']:
raise ValueError("Invalid bits length: {}, expected: {}".format(
len(value),field['length']))
for c in value:
if c == 'Y':
write_row.append(1)
elif c == 'N':
write_row.append(-1)
else:
write_row.append(0)
return has_na
def transform_file(input_file, output_file, fields):
print("**Transforming to file: {}".format(output_file))
with CSVSource(input_file) as reader, \
codecs.open(output_file, "w", "utf-8") as outfile:
writer = csv.writer(outfile)
next(reader)
header = []
# Write the header
for field in fields:
if field['command'] == CMD_IGNORE:
pass
elif field['command'] == CMD_CAT_DUMMY:
header_cat_dummy(field,header)
elif field['command'] == CMD_BITS:
header_bits(field,header)
else:
header_other(field,header)
print("Columns generated: {}".format(len(header)))
writer.writerow(header)
line_count = 0
lines_skipped = 0
# Process the actual file
current_row = -1
header_len = len(header)
for row in reader:
if len(row) != len(fields):
continue
current_row+=1
has_na = False
write_row = []
for field in fields:
value = row[field['index']].strip()
cmd = field['command']
if cmd == 'zscore':
has_na = column_zscore(field,write_row,value, has_na)
elif cmd == CMD_CAT_NUMERIC:
has_na = column_cat_numeric(field,write_row,value, \
has_na)
elif cmd == CMD_IGNORE:
pass
elif cmd == CMD_MAP:
has_na = column_map(field,write_row,value, has_na)
elif cmd == CMD_PASS:
write_row.append(value)
elif cmd == 'date':
write_row.append(str(value[-4:]))
elif cmd == CMD_CAT_DUMMY:
has_na = column_cat_dummy(field,write_row,value,
has_na)
elif cmd == CMD_BITS:
has_na = column_bits(field,write_row,value,has_na)
else:
raise ValueError(\
"Unknown command: {}, stopping.".format(cmd))
if MISSING_SKIP and has_na:
lines_skipped += 1
pass
else:
line_count += 1
writer.writerow(write_row)
# Double check!
if len(write_row) != header_len:
raise ValueError("Inconsistant column "\
"count near line: {}, only had: {}" \
.format(line_count,len(write_row)))
print("Data rows written: {}, skipped: {}"\
.format(line_count,lines_skipped))
print()
def find_field(control, name):
for field in control:
if field['name'] == name:
return field
return None
def find_transformed_fields(header, name):
y = []
x = []
for idx, field in enumerate(header):
if field.startswith(name + '-') or field==name:
y.append(idx)
else:
x.append(idx)
return x,y
def process_for_fit(control, transformed_file, target):
with CSVSource(transformed_file) as reader:
header = next(reader)
field = find_field(control, target)
if field is None:
raise ValueError(f"Unknown target column specified:{target}")
if field['command'] == 'dummy-cat':
print(f"Performing classification on: {target}")
else:
print(f"Performing regression on: {target}")
x_ids, y_ids = find_transformed_fields(header, target)
x = genfromtxt("transformed.csv", delimiter=',', skip_header=1)
y = x[:,y_ids]
x = x[:,x_ids]
return x,y
# -
# The following code takes the data processed from above and trains a neural network.
# +
import pandas as pd
from scipy.stats import zscore
from sklearn.model_selection import StratifiedKFold
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from sklearn import metrics
from sklearn.model_selection import KFold
def generate_network(x,y,task):
model = Sequential()
model.add(Dense(50, input_dim=x.shape[1], activation='relu')) # Hidden 1
model.add(Dense(25, activation='relu')) # Hidden 2
if task == 'classify':
model.add(Dense(y.shape[1],activation='softmax')) # Output
model.compile(loss='categorical_crossentropy', optimizer='adam')
else:
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
return model
def cross_validate(x,y,folds,task):
if task == 'classify':
cats = y.argmax(axis=1)
kf = StratifiedKFold(folds, shuffle=True, random_state=42).split(\
x,cats)
else:
kf = KFold(folds, shuffle=True, random_state=42).split(x)
oos_y = []
oos_pred = []
fold = 0
for train, test in kf:
fold+=1
print(f"Fold #{fold}")
x_train = x[train]
y_train = y[train]
x_test = x[test]
y_test = y[test]
model = generate_network(x,y,task)
model.fit(x_train,y_train,validation_data=(x_test,y_test),verbose=0,
epochs=500)
pred = model.predict(x_test)
oos_y.append(y_test)
if task == 'classify':
# raw probabilities to chosen class (highest probability)
pred = np.argmax(pred,axis=1)
oos_pred.append(pred)
if task == 'classify':
# Measure this fold's accuracy
y_compare = np.argmax(y_test,axis=1) # For accuracy calculation
score = metrics.accuracy_score(y_compare, pred)
print(f"Fold score (accuracy): {score}")
else:
score = np.sqrt(metrics.mean_squared_error(pred,y_test))
print(f"Fold score (RMSE): {score}")
# Build the oos prediction list and calculate the error.
oos_y = np.concatenate(oos_y)
oos_pred = np.concatenate(oos_pred)
if task == 'classify':
oos_y_compare = np.argmax(oos_y,axis=1) # For accuracy calculation
score = metrics.accuracy_score(oos_y_compare, oos_pred)
print(f"Final score (accuracy): {score}")
else:
score = np.sqrt(metrics.mean_squared_error(oos_y, oos_pred))
print(f"Final score (RMSE): {score}")
# -
# ### Running My Sample AutoML Program
#
# These three variables are all you really need to define.
# +
SOURCE_DATA = \
'https://data.heatonresearch.com/data/t81-558/jh-simple-dataset.csv'
TARGET_FIELD = 'product'
TASK = 'classify'
#SOURCE_DATA = 'https://data.heatonresearch.com/data/t81-558/iris.csv'
#TARGET_FIELD = 'species'
#TASK = 'classify'
#SOURCE_DATA = 'https://data.heatonresearch.com/data/t81-558/auto-mpg.csv'
#TARGET_FIELD = 'mpg'
#TASK = 'reg'
# -
# The following lines of code analyze your source data file and figure out how to encode each column. The result is a control file that you can modify to control how each column is handled. The below code should only be run ONCE to generate a control file as a starting point for you to modify.
# +
import csv
import requests
import codecs
control = analyze(SOURCE_DATA)
write_control_file("control.csv",control)
# -
# If your control file is already create, you can start here (after defining the above constants). Do not rerun the previous section, as it will overwrite your control file. Now transform the data.
control = read_control_file("control.csv")
transform_file(SOURCE_DATA,"transformed.csv",control)
# Load the transformed data into properly preprocessed $x$ and $y$.
x,y = process_for_fit(control, "transformed.csv", TARGET_FIELD)
print(x.shape)
print(y.shape)
# Double check to be sure there are no missing values remaining.
import numpy as np
np.isnan(x).any()
# We are now ready to cross validate and train.
cross_validate(x,y,5,TASK)
| t81_558_class_14_01_automl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# %matplotlib inline
import numpy as np
try:
import cPickle as pickle
except:
import pickle
import pandas as pd
import mxnet as mx
import wget
import time
import os.path
import math
import matplotlib.pyplot as plt
import logging
from tqdm import tqdm
import sys
import queue as Queue
import functools
import threading
import os.path
from mxnet.io import DataBatch
# +
ALPHABET = list("abcdefghijklmnopqrstuvwxyz0123456789-,;.!?:'\"/\\|_@#$%^&*~`+ =<>()[]{}")
FEATURE_LEN = 1014
BATCH_SIZE = 128
NUM_FILTERS = 256
DATA_SHAPE = (BATCH_SIZE, 1, FEATURE_LEN, len(ALPHABET))
ctx = mx.gpu(2)
EPOCHS = 10
SD = 0.05 # std for gaussian distribution
INITY = mx.init.Normal(sigma=SD)
LR = 0.01
MOMENTUM = 0.9
WDECAY = 0.00001
# -
# logging
logger = logging.getLogger()
fhandler = logging.FileHandler(filename='crepe_dbp.log', mode='a')
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s')
fhandler.setFormatter(formatter)
logger.addHandler(fhandler)
logger.setLevel(logging.DEBUG)
def load_file(infile):
print("processing data frame: %s" % infile)
# load data into dataframe
df = pd.read_csv(infile,
header=None,
names=['sentiment', 'summary', 'text'])
# concat summary, review; trim to 1014 char; reverse; lower
df['rev'] = df.apply(lambda x: "%s %s" % (x['summary'], x['text']), axis=1)
df.rev = df.rev.str[:FEATURE_LEN].str[::-1].str.lower()
# store class as nparray
y_split = np.asarray(df.sentiment, dtype='int')
print("finished processing data frame: %s" % infile)
print("data contains %d obs, each epoch will contain %d batches" % (df.shape[0], df.shape[0]//BATCH_SIZE))
return df.rev, y_split
def load_data_frame(X_data, y_data, batch_size=128, shuffle=False):
"""
For low RAM this methods allows us to keep only the original data
in RAM and calculate the features (which are orders of magnitude bigger
on the fly). This keeps only 10 batches worth of features in RAM using
asynchronous programing and yields one DataBatch() at a time.
"""
if shuffle:
idx = X_data.index
assert len(idx) == len(y_data)
rnd = np.random.permutation(idx)
X_data = X_data.reindex(rnd)
y_data = y_data[rnd]
# Dictionary to create character vectors
hashes = {}
for index, letter in enumerate(ALPHABET):
hashes[letter] = np.zeros(len(ALPHABET), dtype='bool')
hashes[letter][index] = True
# Yield processed batches asynchronously
# Buffy 'batches' at a time
def async_prefetch_wrp(iterable, buffy=1):#buffy=30
poison_pill = object()
def worker(q, it):
for item in it:
q.put(item)
q.put(poison_pill)
queue = Queue.Queue(buffy)
it = iter(iterable)
thread = threading.Thread(target=worker, args=(queue, it))
thread.daemon = True
thread.start()
while True:
item = queue.get()
if item == poison_pill:
return
else:
yield item
# Async wrapper around
def async_prefetch(func):
@functools.wraps(func)
def wrapper(*args, **kwds):
return async_prefetch_wrp(func(*args, **kwds))
return wrapper
@async_prefetch
def feature_extractor(dta, val):
# Yield mini-batch amount of character vectors
X_split = np.zeros([batch_size, 1, FEATURE_LEN, len(ALPHABET)], dtype='bool')
for ti, tx in enumerate(dta):
chars = list(tx)
print(tx)
for ci, ch in enumerate(chars):
if ch in hashes:
X_split[ti % batch_size][0][ci] = hashes[ch].copy()
# No padding -> only complete batches processed
if (ti + 1) % batch_size == 0:
#yield mx.nd.array(X_split), mx.nd.array(val[ti + 1 - batch_size:ti + 1])
yield X_split, val[ti + 1 - batch_size:ti + 1]
X_split = np.zeros([batch_size, 1, FEATURE_LEN, len(ALPHABET)], dtype='bool')
# Yield one mini-batch at a time and asynchronously process to keep 4 in queue
for Xsplit, ysplit in feature_extractor(X_data, y_data):
#yield DataBatch(data=[Xsplit], label=[ysplit])
yield Xsplit, ysplit
def create_crepe():
"""
Replicating: https://github.com/zhangxiangxiao/Crepe/blob/master/train/config.lua
"""
input_x = mx.sym.Variable('data') # placeholder for input
input_y = mx.sym.Variable('softmax_label') # placeholder for output
# 1. alphabet x 1014
conv1 = mx.symbol.Convolution(data=input_x, kernel=(7, 69), num_filter=NUM_FILTERS)
relu1 = mx.symbol.Activation(data=conv1, act_type="relu")
pool1 = mx.symbol.Pooling(data=relu1, pool_type="max", kernel=(3, 1), stride=(3, 1))
# 2. 336 x 256
conv2 = mx.symbol.Convolution(data=pool1, kernel=(7, 1), num_filter=NUM_FILTERS)
relu2 = mx.symbol.Activation(data=conv2, act_type="relu")
pool2 = mx.symbol.Pooling(data=relu2, pool_type="max", kernel=(3, 1), stride=(3, 1))
# 3. 110 x 256
conv3 = mx.symbol.Convolution(data=pool2, kernel=(3, 1), num_filter=NUM_FILTERS)
relu3 = mx.symbol.Activation(data=conv3, act_type="relu")
# 4. 108 x 256
conv4 = mx.symbol.Convolution(data=relu3, kernel=(3, 1), num_filter=NUM_FILTERS)
relu4 = mx.symbol.Activation(data=conv4, act_type="relu")
# 5. 106 x 256
conv5 = mx.symbol.Convolution(data=relu4, kernel=(3, 1), num_filter=NUM_FILTERS)
relu5 = mx.symbol.Activation(data=conv5, act_type="relu")
# 6. 104 x 256
conv6 = mx.symbol.Convolution(data=relu5, kernel=(3, 1), num_filter=NUM_FILTERS)
relu6 = mx.symbol.Activation(data=conv6, act_type="relu")
pool6 = mx.symbol.Pooling(data=relu6, pool_type="max", kernel=(3, 1), stride=(3, 1))
# 34 x 256
flatten = mx.symbol.Flatten(data=pool6)
# 7. 8704
fc1 = mx.symbol.FullyConnected(data=flatten, num_hidden=1024)
act_fc1 = mx.symbol.Activation(data=fc1, act_type="relu")
drop1 = mx.sym.Dropout(act_fc1, p=0.5)
# 8. 1024
fc2 = mx.symbol.FullyConnected(data=drop1, num_hidden=1024)
act_fc2 = mx.symbol.Activation(data=fc2, act_type="relu")
drop2 = mx.sym.Dropout(act_fc2, p=0.5)
# 9. 1024
fc3 = mx.symbol.FullyConnected(data=drop2, num_hidden=NOUTPUT)
crepe = mx.symbol.SoftmaxOutput(data=fc3, label=input_y, name="softmax")
return crepe
# +
# Visualise symbol (for crepe)
crepe = create_crepe()
#a = mx.viz.plot_network(crepe)
#a.render('Crepe Model')
#a
# -
def save_check_point(mod_arg, mod_aux, pre, epoch):
"""
Save model each epoch, load as:
sym, arg_params, aux_params = \
mx.model.load_checkpoint(model_prefix, n_epoch_load)
# assign parameters
mod.set_params(arg_params, aux_params)
OR
mod.fit(..., arg_params=arg_params, aux_params=aux_params,
begin_epoch=n_epoch_load)
"""
save_dict = {('arg:%s' % k): v for k, v in mod_arg.items()}
save_dict.update({('aux:%s' % k): v for k, v in mod_aux.items()})
param_name = '%s-%04d.pk' % (pre, epoch)
pickle.dump(save_dict, open(param_name, "wb"))
print('Saved checkpoint to \"%s\"' % param_name)
print('Saving model with mxnet notation')
mx.callback.do_checkpoint(pre)
def train_model(prefix, filename):
print("Initializing model")
# Create mx.mod.Module()
cnn = create_crepe()
mod = mx.mod.Module(cnn, context=ctx)
# Bind shape
mod.bind(data_shapes=[('data', DATA_SHAPE)],
label_shapes=[('softmax_label', (BATCH_SIZE,))])
# Initialise parameters and optimiser
mod.init_params(mx.init.Normal(sigma=SD))
mod.init_optimizer(optimizer='sgd',
optimizer_params={
"learning_rate": LR,
"momentum": MOMENTUM,
"wd": WDECAY,
"rescale_grad": 1.0/BATCH_SIZE
})
print("Loading file")
# Load Data
X_train, y_train = load_file(filename)
# Train
print("Alphabet %d characters: " % len(ALPHABET), ALPHABET)
print("started training")
tic = time.time()
# Evaluation metric:
metric = mx.metric.Accuracy()
# Train EPOCHS
for epoch in range(EPOCHS):
t = 0
metric.reset()
tic_in = time.time()
for batch in load_data_frame(X_data=X_train,
y_data=y_train,
batch_size=BATCH_SIZE,
shuffle=True):
# Push data forwards and update metric
mod.forward_backward(batch)
mod.update()
mod.update_metric(metric, batch.label)
# For training + testing
#mod.forward(batch, is_train=True)
#mod.update_metric(metric, batch.label)
# Get weights and update
# For training only
#mod.backward()
#mod.update()
# Log every 50 batches = 128*50 = 6400
t += 1
if t % 50 == 0:
train_t = time.time() - tic_in
metric_m, metric_v = metric.get()
print("epoch: %d iter: %d metric(%s): %.4f dur: %.0f" % (epoch, t, metric_m, metric_v, train_t))
# Checkpoint
print("Saving checkpoint")
arg_params, aux_params = mod.get_params()
save_check_point(mod_arg=arg_params,
mod_aux=aux_params,
pre=prefix,
epoch=epoch)
print("Finished epoch %d" % epoch)
print("Done. Finished in %.0f seconds" % (time.time() - tic))
NOUTPUT = 14 # Classes
model_prefix = 'crepe_dbpedia_prefetch'
train_file = '/datadrive/nlp/dbpedia_train.csv'
#test data
X_train, y_train = load_file(train_file)
# +
for batch in load_data_frame(X_data=X_train, y_data=y_train, batch_size=1,shuffle=False):
#print("label = ", np.asarray(batch.label))
#print("data = ", np.asarray(batch.data))
d,l = batch
dint = np.asarray(d,dtype='int32')
break
print("label = ", l)
print("data shape = ", d.shape)
print("data =", dint)
dint = np.reshape(dint, [1014, 69])
print("dint shape = ", dint.shape)
print(type(dint))
#df = pd.DataFrame(dint)
#df.to_csv("file_path.csv")
np.savetxt('first_batch.csv', dint, delimiter=',', fmt='%d',)
# -
def load_check_point(file_name):
# Load file
print(file_name)
save_dict = pickle.load(open(file_name, "rb"))
# Extract data from save
arg_params = {}
aux_params = {}
for k, v in save_dict.items():
tp, name = k.split(':', 1)
if tp == 'arg':
arg_params[name] = v
if tp == 'aux':
aux_params[name] = v
# Recreate model
cnn = create_crepe()
mod = mx.mod.Module(cnn, context=ctx)
# Bind shape
mod.bind(data_shapes=[('data', DATA_SHAPE)],
label_shapes=[('softmax_label', (BATCH_SIZE,))])
# assign parameters from save
mod.set_params(arg_params, aux_params)
print('Model loaded from disk')
return mod
def test_model(pickled_model, filename):
""" This doesn't take too long but still seems it takes longer than
it should be taking ... """
# Load saved model:
mod = load_check_point(pickled_model)
#assert mod.binded and mod.params_initialized
# Load data
X_test, y_test = load_file(filename)
# Score accuracy
metric = mx.metric.Accuracy()
# Test batches
for batch in load_data_frame(X_data=X_test,
y_data=y_test,
batch_size=BATCH_SIZE):
mod.forward(batch, is_train=False)
mod.update_metric(metric, batch.label)
metric_m, metric_v = metric.get()
print("TEST(%s): %.4f" % (metric_m, metric_v))
model_prefix = 'crepe_dbpedia_prefetch'
model_epoch = 9
model_pk = model_prefix + '_000' + str(model_epoch) + '.pk'
test_file = '/datadrive/nlp/dbpedia_test.csv'
test_model(model_pk, test_file)
| Cloud-Scale_Text_Classification_with_CNNs_on_Azure/python/mxnet/crepe_dbpedia_prefetch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
# %matplotlib inline
# # Notes from Chapter 8 of Werner Krauth Stat Mech book
#
# This is our second attempt at modeling Ising systems, except that we will now follow chapter 8 of Statistical Mechanics: Algorithms and Computations by <NAME>, which is a great source of learning about the basic statistical physics and simulations of Ising model.
#
# * An interesting equivalent of Ising model from chemistry is the colloidal particles. They tend to stick together to form clumps, which is equivalent to ferromagnets, where similar spins tend to align in the same direction.
# ## Intro to Ising models and some basic computations.
#
# In an Ising model, we have N spins on a lattice. We use $k$ to denote the index of a spin, hence $k = 1, \cdots, N$. Each spin has direction, given by $\sigma = \pm 1$. Ferromagnet is a type of Ising model, where neighboring spins prefer to align. In other words, it is energetically more favorable for neighboring spins to be in parallel direction. The energy of a particular configuration is given by:
#
# $$
# E = -J\displaystyle\sum_{<k, l>}\sigma_k \sigma_l \tag{5.1}
# $$
# _Note: Equation numbers follow the numbers from textbook._
#
# A very simple implementation of this energy calculation algorithm is provided in https://www.coursera.org/learn/statistical-mechanics/home/week/8 and shown below:
# +
def energy(S, N, nbr):
E = 0.0
for k in range(N):
E -= S[k] * sum(S[nn] for nn in nbr[k])
return 0.5 * E
L = 2
nbr = [[1, 2], [3, 0], [3, 0], [2, 1]]
S = [1, 1, -1, 1]
print S, energy(S, L * L, nbr)
# -
# In the next step, we will learn how to enumerate configurations of a lattice of spins. For N spins, the total number of possible configurations are $2^N$. Each configuration $i$, where $i = 1, \cdots, N$ is related to the binary representation of the number $i - 1$. We can imagine that in the binary representation $0's$ correspond to $-1$. For $N = 10$ and $i = 4$, the corresponding configuration of the system is: $0001$ or $\{-1, -1, -1, 1\}$.
#
# In short, the enumeration of configurations of an $N$ spin lattice is simply the process of counting number from $0$ to $2^N - 1$ and converting each number to its binary representation.
| scratch/krauth_chapter_8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Motif discovery and regulatory analysis - I
# # Table of Contents
# 1. Consensus sequences
# 2. Probability and positional weight matrices
# 3. Information content / entropy
# 4. Motif finding approaches
# ## 1. Consensus sequences
# As you saw in the prelab lecture, there are many ways to represent motifs. In this assignment, we are going to have some more practice with these different representations and the kinds of interesting information contained in each one.
#
# One simple way to represent motifs which is easy for people to actually look at is the <b>exact consensus sequence representation</b>. In this representation, a motif is encoded as the most common base at each position. Say you have the following examples of a given motif:
#
# 1. ACAGGAA
# 2. TGCGGAA
# 3. TGAGGAT
# 4. AGTGGAA
# 5. AACGGAA
# 6. ACAGGAT
# By finding the most common base at each position, what is the exact consensus sequence for this motif?
# + active=""
#
# -
# Although there is a single most common letter at each position in this example, you probably noticed that many of these positions seem to be somewhat flexible, where there is another nucleotide that comes up almost as frequently as the most common base. It is quite common for motifs such as transcription factor binding motifs to include some level of flexibility or degeneracy, and so we also have a human-readable way to encode this, called the <b>degenerate consensus sequence representation</b>.
#
# There are two common ways to encode this. One is related to the concept of regular expressions that we have seen a few times now, where the set of symbols that are possible at each position is contained in brackets, i.e. [AT] means that position can contain either an A or a T. Using this representation, what is the degenerate consensus sequence for this motif?
# + active=""
#
# -
# In this case, we have two positions that seem to be able to contain three different nucleotides. For the sake of clarity, a common convention is to only include a base as a degenerate possibility if more than 25% of the input sequences include that base. In this example, that means that a base that is only present in one of the sequences should not be counted. Rewrite your degenerate representation using this convention:
# + active=""
#
# -
# The other way to represent degenerate consensus sequences is to use specific characters (defined by IUPAC) to represent these sets of possibilities:
# <table width=100%>
# <tr><th>Symbol</th><th>Description</th><th>Bases represented</th><th>Number of bases represented</th></tr>
# <tr><td>A</td><td>Adenine</td><td>A</td><td>1</td></tr>
# <tr><td>C</td><td>Cytosine</td><td>C</td><td>1</td></tr>
# <tr><td>G</td><td>Guanine</td><td>G</td><td>1</td></tr>
# <tr><td>T</td><td>Thymine</td><td>T</td><td>1</td></tr>
# <tr><td>U</td><td>Uracil</td><td>U</td><td>1</td></tr>
# <tr><td>W</td><td>Weak hydrogen bonding</td><td>A,T</td><td>2</td></tr>
# <tr><td>S</td><td>Strong hydrogen bonding</td><td>G,C</td><td>2</td></tr>
# <tr><td>M</td><td>aMino</td><td>A,C</td><td>2</td></tr>
# <tr><td>K</td><td>Keto</td><td>G,T</td><td>2</td></tr>
# <tr><td>R</td><td>puRine</td><td>A,G</td><td>2</td></tr>
# <tr><td>Y</td><td>pYrimidine</td><td>C,T</td><td>2</td></tr>
# <tr><td>B</td><td>not A (B comes after A)</td><td>C,G,T</td><td>3</td></tr>
# <tr><td>D</td><td>not C (D comes after C)</td><td>A,G,T</td><td>3</td></tr>
# <tr><td>H</td><td>not G (H comes after G)</td><td>A,C,T</td><td>3</td></tr>
# <tr><td>V</td><td>not T (V comes after T)</td><td>A,C,G</td><td>3</td></tr>
# <tr><td>N or -</td><td>any Nucleotide (not a gap)</td><td>A,C,G,T</td><td>4</td></tr>
# </table>
# Using this approach, write the representation of the motif with all the possible degenerate positions (don't filter out bases that only appear once in a position):
# + active=""
#
# -
# Now write the representation of the motif with the cleaner definition of degenerate positions (do filter out bases that appear only once in a position):
# + active=""
#
# -
# ## 2. Probability and positional weight matrices
# So far in this lab, we have seen motif representations that are meant to be easily human-readable and interpretable. However, one issue with these representations is that they throw away quantitative information about the probability of each base at each position, and so we cannot use them for any more mathematical approaches to motif interpretation. One very common alternative representation that retains this information is the <b>probability weight matrix (PWM)</b>, which is a matrix with 4 rows, one for each nucleotide, and a number of columns corresponding to the length of the motif. For example, the PWM representation of the six motifs from above (ACAGGAA, TGCGGAA, TGAGGAT, AGTGGAA, AACGGAA, ACAGGAT) is:
# <table width=100%><tr><th>Nucleotide</th><th>Pos. 1 Probability (Observed Counts)</th><th>Pos. 2 Probability (Observed Counts)</th><th>Pos. 3 Probability (Observed Counts)</th><th>Pos. 4 Probability (Observed Counts)</th><th>Pos. 5 Probability (Observed Counts)</th><th>Pos. 6 Probability (Observed Counts)</th><th>Pos. 7 Probability (Observed Counts)</th></tr>
# <tr><td>A</td><td>0.66 (4)</td><td>0.166 (1)</td><td>0.5 (3)</td><td>0.0 (0)</td><td>0.0 (0)</td><td>1.0 (6)</td><td>0.66 (4)</td></tr>
# <tr><td>C</td><td>0.0 (0)</td><td>0.33 (2)</td><td>0.33 (2)</td><td>0.0 (0)</td><td>0.0 (0)</td><td>0.0 (0)</td><td>0.0 (0)</td></tr>
# <tr><td>G</td><td>0.0 (0)</td><td>0.5 (3)</td><td>0.0 (0)</td><td>1.0 (6)</td><td>1.0 (6)</td><td>0.0 (0)</td><td>0.0 (0)</td></tr>
# <tr><td>T</td><td>0.33 (2)</td><td>0.0 (0)</td><td>0.166 (1)</td><td>0.0 (0)</td><td>0.0 (0)</td><td>0.0 (0)</td><td>0.33 (2)</td></tr>
# </table>
# Using this table, we can use a simple approach of finding how well a given putative motif sequence matches what we think the real motif is by just comparing it to this table and multiplying the probability at each base. For example, if we want to quantify how well the motif 'AGAGGAA' (which was our exact consensus sequence) matches, we just go through and multiply 0.66 \* 0.5 \* 0.5 \* 1.0 \* 1.0 \* 1.0 \* 0.66 = .1089. One major issue with using this approach is the fact that some of these cells contain '0.0' as their probability. Consider the motif 'CGAGGAA', which only differs from our exact consensus sequence by a single base pair. If we try to use the same quantification approach, we will compute 0.0 \* 0.5 \* 0.5 \* 1.0 \* 1.0 \* 1.0 \* 0.66 = <b>0.0</b>. In other words, the fact that we had one position containing a nucleotide that was not observed in our reference set means that the probability of that motif, under this PWM, is 0. To avoid this issue, we can add a 'pseudocount' of 1 at every position for every nucleotide, yielding the following PWM:
# <table width=100%><tr><th>Nucleotide</th><th>Pos. 1 Probability (Obs + Pseudocounts)</th><th>Pos. 2 Probability (Obs + Pseudocounts)</th><th>Pos. 3 Probability (Obs + Pseudocounts)</th><th>Pos. 4 Probability (Obs + Pseudocounts)</th><th>Pos. 5 Probability (Obs + Pseudocounts)</th><th>Pos. 6 Probability (Obs + Pseudocounts)</th><th>Pos. 7 Probability
# (Obs + Pseudocounts)</th></tr>
# <tr><td>A</td><td>0.5 (5)</td><td>0.2 (2)</td><td>0.4 (4)</td><td>0.1 (1)</td><td>0.1 (1)</td><td>0.7 (7)</td><td>0.5 (5)</td></tr>
# <tr><td>C</td><td>0.1 (1)</td><td>0.3 (3)</td><td>0.3 (3)</td><td>0.1 (1)</td><td>0.1 (1)</td><td>0.1 (1)</td><td>0.1 (1)</td></tr>
# <tr><td>G</td><td>0.1 (1)</td><td>0.4 (4)</td><td>0.1 (1)</td><td>0.7 (7)</td><td>0.7 (7)</td><td>0.1 (1)</td><td>0.1 (1)</td></tr>
# <tr><td>T</td><td>0.3 (3)</td><td>0.1 (1)</td><td>0.2 (2)</td><td>0.1 (1)</td><td>0.1 (1)</td><td>0.1 (1)</td><td>0.3 (3)</td></tr>
# </table>
# Now if we try to compute the probability of observing 'CGAGGAA', we get 0.1 \* 0.4 \* 0.4 \* 0.7 \* 0.7 \* 0.7 \* 0.5 = 0.0027.
# What is the probability of observing a motif very unlike what we have seen, say 'CTCTTTG'?
# + active=""
#
# -
# ### Generating positional weight matrices
# A further refinement to this idea is to correct these probabilities for the background distribution of bases in the genome you are interested in. Doing this, we can define <b>positional weight matrices</b>. To do this, after we have obtained the matrix of probabilities including pseudocounts (i.e. the table directly above this one), we divide each entry in each row by the background probability of observing the nucleotide corresponding to that row. In the naive case, we just use <i>p(i)</i> = 0.25 for each nucleotide <i>i</i>. This assumes an equal probability of observing any given nucleotide. Finally, a common transformation is to take the natural logarithm (ln, or log base e) of each of these background-corrected quantities (note that these are no longer probabilities). This is done so that in order to compute the score for a given sequence, the entries in each row can be added instead of multiplied together. In our example above, applying these transformations using the naive nucleotide distribution yields the following table:
#
# <table width=100%><tr><th>Nucleotide</th><th>Pos. 1 Log-odds</th><th>Pos. 2 Log-odds</th><th>Pos. 3 Log-odds</th><th>Pos. 4 Log-odds</th><th>Pos. 5 Log-odds</th><th>Pos. 6 Log-odds</th><th>Pos. 7 Log-odds</th></tr>
# <tr><td>A</td><td>0.693</td><td>-0.223</td><td>0.470</td><td>-0.916</td><td>-0.916</td><td>1.030</td><td>0.693</td></tr>
# <tr><td>C</td><td>-0.916</td><td>0.182</td><td>0.182</td><td>-0.916</td><td>-0.916</td><td>-0.916</td><td>-0.916</td></tr>
# <tr><td>G</td><td>-0.916</td><td>0.470</td><td>-0.916</td><td>1.030</td><td>1.030</td><td>-0.916</td><td>-0.916</td></tr>
# <tr><td>T</td><td>0.182</td><td>-0.916</td><td>-0.223</td><td>-0.916</td><td>-0.916</td><td>-0.916</td><td>0.182</td></tr>
# </table>
#
# Now, the corrected probability of any given sequence can be computed by simply adding the entries corresponding to that sequence. If the score is greater than 0, the sequence is more likely to be a functional than a 'random' sequence, and if the score is less than 0, the reverse is true. This is why the column titles refer to the 'log-odds': this model represents the 'odds' or likelihood that a given sequence matches the motif. Compute the score for the exact consensus sequence 'AGAGGAA':
# + active=""
#
# -
# It is worth noting that the human genome does not follow the naive distribution of an equal probability of observing each nucleotide. Instead, the distribution is roughly <i>p(A) = 0.3</i>, <i>p(C) = 0.2</i>, <i>p(G) = 0.2</i>, and <i>p(T) = 0.3</i>. Using this, we can recompute our positional weight matrix:
#
# <table width=100%><tr><th>Nucleotide</th><th>Pos. 1 Log-odds</th><th>Pos. 2 Log-odds</th><th>Pos. 3 Log-odds</th><th>Pos. 4 Log-odds</th><th>Pos. 5 Log-odds</th><th>Pos. 6 Log-odds</th><th>Pos. 7 Log-odds</th></tr>
# <tr><td>A</td><td>0.510</td><td>-0.405</td><td>0.288</td><td>-1.099</td><td>-1.099</td><td>0.847</td><td>0.511</td></tr>
# <tr><td>C</td><td>-0.693</td><td>0.405</td><td>-0.693</td><td>-0.693</td><td>-0.693</td><td>-0.693</td><td>-0.693</td></tr>
# <tr><td>G</td><td>-0.693</td><td>0.693</td><td>-0.693</td><td>1.253</td><td>1.253</td><td>-0.693</td><td>-0.693</td></tr>
# <tr><td>T</td><td>0.000</td><td>-1.099</td><td>-0.405</td><td>-1.099</td><td>-1.099</td><td>-1.099</td><td>0.000</td></tr>
# </table>
#
# Now what is the score for the exact consensus sequence 'AGAGGAA'?
# + active=""
#
# -
# ## 3. Information content and entropy
# One aspect of these PWMs that we have not yet addressed is the concept of how well they actually capture the motif, or how informative they actually are. In other words, we want to know how well a motif, as represented by a PWM, can discriminate between a real signal and background noise. To do so, we can take advantage of a very useful and powerful concept called the <b>information content (IC)</b> of a motif. This is a way of directly quantifying how informative a signal is, and applications of this concept can be found in a wide range of fields from computer encryption to machine learning to physics. In this case, we define the information content of each column $j$ in the PWM (i.e. each position in the motif) as $IC_j = 2 + \sum_{x=A,C,G,T} p_x log_2(p_x)$, where $p_x$ is the entry for nucleotide $x$ in that column. This means that a value of 2.0 is the most informative and a value of 0 is the least informative. Consider the following simple PWM:
#
# <table width=100%><tr><th>Nucleotide</th><th>Pos. 1 Probability</th><th>Pos. 2 Probability</th><th>Pos. 3 Probability</th></tr>
# <tr><td>A</td><td>1.00</td><td>0.25</td><td>0.4</td></tr>
# <tr><td>C</td><td>0.00</td><td>0.25</td><td>0.4</td></tr>
# <tr><td>G</td><td>0.00</td><td>0.25</td><td>0.1</td></tr>
# <tr><td>T</td><td>0.00</td><td>0.25</td><td>0.1</td></tr>
# </table>
# The IC for each column can be calculated:
#
# $IC_1 = 2 + 1.0 * log_2(1.0) + 0.0 + 0.0 + 0.0 = 2$
#
# $IC_2 = 2 + 0.25 * log_2(0.25) + 0.25 * log_2(0.25) + 0.25 * log_2(0.25) + 0.25 * log_2(0.25) = 2 + 0.25 * (-2) + 0.25 * (-2) + 0.25 * (-2) + 0.25 * (-2) = 0$
#
# $IC_3 = 2 + 0.4 * log_2(0.4) + 0.4 * log_2(0.4) + 0.1 * log_2(0.1) + 0.1 * log_2(0.1) = 2 + 0.4 * (-1.32) + 0.4 * (-1.32) + 0.1 * (-3.32) + 0.1 * (-3.32) = 0.27$
#
# So we see that the first position is maximally informative (intuitively, we know that it will always be an A), while the second position is minimally informative (each base has an exactly equal chance of occuring), and the third position is weakly informative (it is more likely to be an A or a C than a G or a T).
#
# Then, the IC for a motif can be calculated as the sum of the information contents of each column, so this motif would have an IC of 2.27.
# Similarly to how we wanted to generate positional weight matrices to correct for the background nucleotide distributions, we may also want to account for the background nucleotide probabilities when we look at the information content in a motif. There is a related concept called <b>relative entropy</b> that allows us to do this. Entropy measures the 'randomness' of a signal, and in that sense is the opposite of information. Relative entropy measures this 'randomness' or 'disorderedness' of a given motif relative to the background distribution. In other words, relative entropy measures how different your motif is from what you would expect given the background distribution; thus, if a motif is very informative, it will have a high relative entropy.
#
# The equation for relative entropy is given as $RE = \sum_{x=A,C,G,T} p_x log_2(p_x/Q_x)$, where $Q_x$ is the background probability of the nucleotide $x$. Thus, if your PWM exactly matches the background probability Q, the relative entropy of your PWM will be 0 (because $p_x / Q_x = 1$ and $log_2(1) = 0$); otherwise, this quantity can be arbitrarily high or low.
# ### Aside: creating motif logos
# A useful way of representing motifs is using what are known as sequence logos, which we saw in the prelab lecture. These logos scale each nucleotide at each position to represent their information content. An easy way to create these logos is to use the website http://weblogo.berkeley.edu/logo.cgi. We will practice this with the set of 6 sequences we were looking at earlier. The general approach is to upload a set of sequences, either by copy and pasting or by uploading the file. These sequences can be provided in fasta format, as we have done here, or as a plain text list, where each line is the same length, as we have in question 4 on the homework. Here, we will just copy and paste the 6 sequences from this box:
# + active=""
# >seq1
# ACAGGAA
# >seq2
# TGCGGAA
# >seq3
# TGAGGAT
# >seq4
# AGTGGAA
# >seq5
# AACGGAA
# >seq6
# ACAGGAT
# -
# Then, navigate to the website and paste those sequences into the box marked 'multiple sequence aligment'. Then, simply press the 'create logo' button, and you should get a sequence logo! Save this file and upload it into the images/ folder of this assignment.
# ## 4. Motif finding approaches
# As we saw in the lecture, there are several different computational approaches that can be used to identify enriched motifs in a given set of sequences, including exact counting, iterative approaches like Gibbs' sampling and expectation maximization, and differential enrichment approaches. For this section of the lab, we will just have some practice using the most common tool for motif enrichment in relatively small datasets, MEME, which is based on expectation maximization.
#
# We will analyze the file called 'selex_seqs.fasta', in the inclass_data/ folder. This fasta-formatted file contains sequences from a SELEX-like experiment, where sequences were pulled down based on their affinity with some transcription factor. We will use the online MEME tool to do this. You can either download this file to your computer (recommended) or copy and paste it to upload it to MEME, but make sure you get the full file if you do this. Navigate to http://meme-suite.org/tools/meme, and under the <b>input the primary sequences</b> header, select whichever approach you are using to upload the sequences.
#
# Under <b>select the site distribution</b>, choose 'one occurrence per sequence', because this file comes from a SELEX-like experiment and so each sequence was experimentally found to bind to some transcription factor. Leave the value of 3 for how many motifs MEME should find, and under advanced options, change the maximum width of the motifs to 20bp to speed up the computation. This will take some time to finish running, so make sure to save the link, or you can provide an email address that they will mail the link to. Make sure to submit this job before starting the homework as some of the questions will be about these results!
# ## Homework problems: motif practice
# <b>Question 1:</b> Consider the following probability weight matrix:
#
# <table width=70%><tr><th>Nucleotide</th><th>Pos. 1</th><th>Pos. 2</th><th>Pos. 3</th><th>Pos. 4</th><th>Pos. 5</th><th>Pos. 6</th><th>Pos. 7</th><th>Pos. 8</th></tr>
# <tr><td>A</td><td>0.01</td><td>0.1</td><td>0.97</td><td>0.95</td><td>0.5</td><td>0.05</td><td>0.8</td><td>0.4</td></tr>
# <tr><td>C</td><td>0.03</td><td>0.05</td><td>0.01</td><td>0.01</td><td>0.1</td><td>0.6</td><td>0.1</td><td>0.08</td></tr>
# <tr><td>G</td><td>0.95</td><td>0.05</td><td>0.01</td><td>0.03</td><td>0.1</td><td>0.05</td><td>0.05</td><td>0.02</td></tr>
# <tr><td>T</td><td>0.01</td><td>0.8</td><td>0.01</td><td>0.01</td><td>0.3</td><td>0.3</td><td>0.05</td><td>0.5</td></tr>
# </table>
#
# What is the information content of positions 3 and 5 in this matrix? <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 2:</b> Using the PWM given above, what is the exact consensus sequence and the degenerate consensus sequence (using either the regular expression or IUPAC characters)? For the degenerate sequence, only count a nucleotide as a degenerate possibility if it has a probability of more than 0.25. <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 3 (short answer):</b> Based on this consensus sequence, do you expect the relative entropy of this probability matrix to be higher when compared to the naive nucleotide distribution (equal probability of any nucleotide) or to the human genome background probability (A and T are more common than G and C)? <b>(1 point)</b>
# + active=""
#
# -
# For the next two questions, we will be using the following set of sequences:
# + active=""
# TGGGAA
# TGGGAA
# TGGGAA
# TGGGAA
# TGGGAA
# TGAGAA
# TGGGAA
# TGGGAA
# TGGGAA
# TGGGAG
# TGAGAA
# TGAGAA
# TGTGAA
# TGGGAA
# TGGGAG
# TGGGAG
# CGGGAA
# TGGGAT
# -
# <b>Question 4:</b> From these sequences, create a positional weight matrix corrected for the human genome background probabilities (p(A) = 0.3, p(C) = 0.2, p(G) = 0.2, and p(T) = 0.3). Recall that this involves counting the nucleotide occurrences at each position, adding a psuedocount of 1, normalizing this into probabilities of each base, and finding the log odds by taking the natural log of each probability divided by the frequency of that nucleotide in the genome. <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 5:</b> Using these sequences, make a logo using the webLogo tool and upload this logo to the 'images/' directory in this assignment <b>(1 point)</b>
# Hopefully your MEME results are now ready, because the rest of the questions will deal with their analysis. On the MEME output page, follow the link to the 'MEME HTML output.' You should see ‘Discovered Motifs’, ‘Motif Locations’, and ‘Program information’ on this result page. ‘Program information’ contains some basic information about what version of MEME you used, your input data, and the parameters. Clicking the “?” links will give you more information on what each output column means, and may help you answer the questions below.
#
# We are going to look at motif 1, which is the 'best' motif identified in the data by MEME. If you press the down arrow under the 'More' column, you can see more information about this motif.
#
# <b>Question 6:</b> Based on the sequence logo, what is the consensus sequence for this motif (either exact or degenerate)? <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 7:</b> Now click on the right-facing arrow, which leads you to the 'submit or download motif' page. Select the 'download motif' tab, where you can download the motif in count matrix format, probability matrix format (useful for finding the degenerate consensus sequence), minimal MEME format, FASTA, and raw formats. Select the minimal MEME format. As you scroll through the window, you should see two familiar matrices, the log-odds matrix and the letter probability matrix. In general, what are some key differences between these two matrices? (PS: we are not looking for answer such as PWM is non-negative) <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 8:</b> What is the E-value of this motif? What does this value represent? Should this be considered to be a significant hit? (Hint: the question mark box next to the E-value contains valuable information.. Hint #2: in the E-value column, a positive number after 'e' means move the decimal point to the right.) <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 9:</b> How many sites contain this motif? What is the information content of this motif? <b>(1 point)</b>
# + active=""
#
# -
# <b>Question 10 (short answer):</b> Of the 3 motifs identified, which, if any, do you think are the true motif? What are you basing this on? Also, if there is only 1 true motif, why do we identify 3? <b>(1 point)</b>
# + active=""
#
| 10_Motif-I/motif_1_inclass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: metacall_jupyter
# language: text
# name: metacall_jupyter
# ---
# # Shell Commands on the MetaCall Jupyter
#
# Using the `!` prefix, you can run Shell commands on the MetaCall Jupyter. Through this, you can install `pip`, `npm` and other dependencies as required for your polyglot applications. You can also use this to interact with external files and run them straight through the Notebook interface.
#
# Note that `!` must be the first character in the cell, otherwise it will not be interpreted as shell prefix character.
!ls
!metacall pip3 install requests==2.20.0
# +
>python
import requests
from urllib.parse import urlencode
import sys
def make_tiny(url):
request_url = ('http://tinyurl.com/api-create.php?' +
urlencode({'url': url}))
result = requests.get(request_url)
return result.text
print(make_tiny("http://harshcasper.hashnode.dev/polyglot-programming-with-metacall"))
| examples/03-Shell-Commands-MetaCall-Jupyter-Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def print_greeting():
print('Hello!')
print_greeting()
def print_date(year, month, day):
"""
prints the date in the format year/month/day
lots of test here
each on it's own line
"""
joined = str(year) + '/' + str(month) + '/' + str(day)
print(joined)
print_date(2019, 8, 27)
print_date(11, 11, 11)
help(print_date)
print_date.__doc__
print_date(day=12, month=4, year=1999)
values=[5,6]
len(values)
if len(values) == 0:
print("empty")
else:
print("not empty")
len(values) = 0
a = 1
if a = 2:
print("nope")
def average(values):
if len(values) == 0:
return None
| functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <table style="width:100%">
# <tr>
# <td style="background-color:#EBF5FB; border: 1px solid #CFCFCF">
# <b>National generation capacity: Check notebook</b>
# <ul>
# <li><a href="main.ipynb">Main notebook</a></li>
# <li><a href="processing.ipynb">Processing notebook</a></li>
# <li>Check notebook (this)</li>
# </ul>
# <br>This Notebook is part of the <a href="http://data.open-power-system-data.org/national_generation_capacity">National Generation Capacity Datapackage</a> of <a href="http://open-power-system-data.org">Open Power System Data</a>.
# </td>
# </tr>
# </table>
# # Table of Contents
# * [1. Introductory notes](#1.-Introductory-notes)
# * [2. Script setup](#2.-Script-setup)
# * [3. Import of processed data](#3.-Import-of-processed-data)
# * [4. Visualisation of results for different energy source levels](#4.-Visualisation-of-results-for-different-energy-source-levels)
# * [4.1 Energy source level 1](#4.1-Energy-source-level-1)
# * [4.1.1 Table](#4.1.1-Table)
# * [4.1.2 Bokeh chart](#4.1.2-Bokeh-chart)
# * [4.2 Energy source level 2](#4.2-Energy-source-level-2)
# * [4.2.1 Table](#4.2.1-Table)
# * [4.2.2 Bokeh chart](#4.2.2-Bokeh-chart)
# * [4.3 Energy source level 3](#4.3-Energy-source-level-3)
# * [4.3.1 Table](#4.3.1-Table)
# * [4.3.2 Bokeh chart](#4.3.2-Bokeh-chart)
# * [4.4 Technology level](#4.4-Technology-level)
# * [4.4.1 Table](#4.4.1-Table)
# * [4.4.2 Bokeh chart](#4.4.2-Bokeh-chart)
# * [5. Comparison of total capacity for energy source levels](#5.-Comparison-of-total-capacity-for-energy-source-levels)
# * [5.1 Calculation of total capacity for energy source levels](#5.1-Calculation-of-total-capacity-for-energy-source-levels)
# * [5.2 Identifcation of capacity differences for energy source levels](#5.2-Identifcation-of-capacity-differences-for-energy-source-levels)
#
# # 1. Introductory notes
# The notebook extends the [processing notebook](processing.ipynb) to make visualisations and perform consistency checks.
# # 2. Script setup
# +
import pandas as pd
import numpy as np
import os.path
import logging
from bokeh.charts import Bar, output_file, show
from bokeh.io import output_notebook
from bokeh.models import HoverTool, NumeralTickFormatter
from bokeh.charts.attributes import color
output_notebook()
# %matplotlib inline
logging.basicConfig(
level=logging.INFO,
format='%(asctime)s - %(name)s - %(levelname)s - %(message)s',
datefmt='%d %b %Y %H:%M:%S'
)
logger = logging.getLogger()
# -
# # 3. Import of processed data
# The processed data is imported at this stage. This requires that the [processing notebook](processing.ipynb) is compiled prior to this step. Otherwise, no or an old dataset is imported.
# +
data_file = 'national_generation_capacity_stacked.csv'
filepath = os.path.join('output', data_file)
data = pd.read_csv(filepath, index_col=0)
data.head()
# -
# # 4. Visualisation of results for different energy source levels
# In the following, national generation capacity is compared for the different energy source levels. Due to the size of the dataset regarding countries and compiled datasources, the following tables and figures tend to be rather confusing. Therefore, we provide the following option to limit the visualisation of the results to a selection of countries and years. If the corresponding subset is empty, all values will be considered.
country_subset = ['DE', 'FR', 'BE', 'NL', 'IT']
year_subset = [2013, 2014, 2015, 2016]
# +
data_selection = pd.DataFrame()
if len(country_subset) == 0:
data_selection = data
else:
for country in country_subset:
for year in year_subset:
if len(data_selection) == 0:
data_selection = data[(data.country == country) &
(data.year == year)]
else:
data_selection = data_selection.append(data[(data.country == country) &
(data.year == year)])
data_selection
# -
# To improve the data visualisation in Bokeh the colors of the defined technologies is specified explicitly. The user is free to adjust or refine the color definition using the following parameter. The color names are defined [here](http://www.w3schools.com/colors/colors_names.asp).
colormap = {
'Fossil fuels': 'Black',
'Lignite': 'SaddleBrown',
'Hard coal': 'Black',
'Oil': 'Violet',
'Natural gas': 'IndianRed',
'Combined cycle': '#d57676',
'Gas turbine': '#e19d9d',
'Other and unknown natural gas': '#c33c3c',
'Differently categorized natural gas': 'IndianRed',
'Non-renewable waste': 'SandyBrown',
'Mixed fossil fuels': 'LightGray',
'Other fossil fuels': 'DarkGray',
'Differently categorized fossil fuels': 'Gray',
'Nuclear': 'Red',
'Renewable energy sources': 'Green',
'Hydro': 'Navy',
'Run-of-river': '#0000b3',
'Reservoir': '#0000e6',
'Reservoir including pumped storage': '#0000e6',
'Pumped storage': '#1a1aff',
'Pumped storage with natural inflow': '#1a1aff',
'Differently categorized hydro': 'Navy',
'Wind': 'SkyBlue',
'Onshore': 'LightSkyBlue',
'Offshore': 'DeepSkyBlue',
'Differently categorized wind': 'SkyBlue',
'Solar': 'Yellow',
'Photovoltaics': '#ffff33',
'Concentrated solar power': '#ffff66',
'Differently categorized solar': 'Yellow',
'Geothermal': 'DarkRed',
'Marine': 'Blue',
'Bioenergy and renewable waste': 'Green',
'Biomass and biogas': '#00b300',
'Sewage and landfill gas': '#00e600',
'Other bioenergy and renewable waste': 'Green',
'Differently categorized renewable energy sources': 'Green',
'Other or unspecified energy sources': 'Orange',
}
# ## 4.1 Energy source level 1
# ### 4.1.1 Table
# +
pivot_capacity_level1 = pd.pivot_table(data_selection[data_selection.energy_source_level_1 == True],
index=('country','year','source'),
columns='technology',
values='capacity',
aggfunc=sum,
margins=False)
pivot_capacity_level1
# -
# ### 4.1.2 Bokeh chart
# Please use the zoom and hover option to inspect the data graphically.
# +
data_energy_level_1 = data_selection[data_selection.energy_source_level_1 == True].copy()
data_energy_level_1['color'] = 'White'
data_energy_level_1['color'] = data_energy_level_1['technology'].map(colormap)
bar = Bar(data_energy_level_1, values='capacity', label=['country', 'year', 'source'], stack='technology',
title="National capacity by type of energy source", tools="pan,wheel_zoom,box_zoom,reset,hover,save",
legend='top_right',
plot_width=1600, plot_height=800,
# color=color(columns='technology', palette=['Black', 'Red', 'Green', 'Orange'], sort=False))
color='color')
bar._yaxis.formatter = NumeralTickFormatter(format="00,000 MW")
hover = bar.select_one(HoverTool)
hover.point_policy = "follow_mouse"
hover.tooltips = [("Country", "@country"),
("Year", "@year"),
("Source", "@source"),
("Category", "@technology"),
("Capacity", "@height{00,000.00} MW"),
]
show(bar)
# -
# ## 4.2 Energy source level 2
# ### 4.2.1 Table
# +
pivot_capacity_level2 = pd.pivot_table(data_selection[data_selection.energy_source_level_2 == True],
index=('country','year','source'),
columns='technology',
values='capacity',
aggfunc=sum,
margins=False)
pivot_capacity_level2
# -
# ### 4.2.2 Bokeh chart
# Please use the zoom and hover option to inspect the data graphically.
# +
data_energy_level_2 = data_selection[data_selection.energy_source_level_2 == True].copy()
data_energy_level_2['color'] = 'White'
data_energy_level_2['color'] = data_energy_level_2['technology'].map(colormap)
bar = Bar(data_energy_level_2, values='capacity', label=['country', 'year', 'source'], stack='technology',
title="National capacity by energy source", tools="pan,wheel_zoom,box_zoom,reset,hover,save",
legend='top_right',
plot_width=1600, plot_height=800,
color='color'
)
bar._yaxis.formatter = NumeralTickFormatter(format="00,000 MW")
hover = bar.select_one(HoverTool)
hover.point_policy = "follow_mouse"
hover.tooltips = [("Country", "@country"),
("Year", "@year"),
("Source", "@source"),
("Category", "@technology"),
("Capacity", "@height{00,000.00} MW"),
]
show(bar)
# -
# ## 4.3 Energy source level 3
# ### 4.3.1 Table
# +
pivot_capacity_level3 = pd.pivot_table(data_selection[data_selection.energy_source_level_3 == True],
index=('country', 'year', 'source'),
columns='technology',
values='capacity',
aggfunc=sum,
margins=False)
pivot_capacity_level3
# -
# ### 4.3.2 Bokeh chart
# Please use the zoom and hover option to inspect the data graphically.
# +
data_energy_level_3 = data_selection[data_selection.energy_source_level_3 == True].copy()
data_energy_level_3['color'] = 'White'
data_energy_level_3['color'] = data_energy_level_3['technology'].map(colormap)
bar = Bar(data_energy_level_3, values='capacity', label=['country', 'year', 'source'], stack='technology',
title="National capacity by energy source", tools="pan,wheel_zoom,box_zoom,reset,hover,save",
# legend='top_right',
plot_width=1600, plot_height=800,
color='color'
)
bar._yaxis.formatter = NumeralTickFormatter(format="00,000 MW")
hover = bar.select_one(HoverTool)
hover.point_policy = "follow_mouse"
hover.tooltips = [("Country", "@country"),
("Year", "@year"),
("Source", "@source"),
("Category", "@technology"),
("Capacity", "@height{00,000.00} MW"),
]
show(bar)
# -
# ## 4.4 Technology level
# ### 4.4.1 Table
# +
pivot_capacity_techlevel = pd.pivot_table(data_selection[data_selection.technology_level == True],
index=('country', 'year', 'source'),
columns='technology',
values='capacity',
aggfunc=sum,
margins=False)
pivot_capacity_techlevel
# -
# ### 4.4.2 Bokeh chart
# Please use the zoom and hover option to inspect the data graphically.
# +
data_technology_level = data_selection[data_selection.technology_level == True].copy()
data_technology_level['color'] = 'White'
data_technology_level['color'] = data_technology_level['technology'].map(colormap)
bar = Bar(data_technology_level, values='capacity', label=['country', 'year', 'source'], stack='technology',
title="National capacity by energy source and technology", tools="pan,wheel_zoom,box_zoom,reset,hover,save",
# legend='top_right',
plot_width=1600, plot_height=800,
color='color'
)
bar._yaxis.formatter = NumeralTickFormatter(format="00,000 MW")
hover = bar.select_one(HoverTool)
hover.point_policy = "follow_mouse"
hover.tooltips = [("Country", "@country"),
("Year", "@year"),
("Source", "@source"),
("Category", "@technology"),
("Capacity", "@height{00,000.00} MW"),
]
show(bar)
# -
# # 5. Comparison of total capacity for energy source levels
# In the following, the installed capacities at the different technology levels are compared to each other. In any case, the total sum of all technologies within a certain technology level should match with other energy source levels. Otherwise the classification of categories to the levels is flawed or the specific data entries are wrong.
#
# Again, the comparison can be done for specific countries, or, if the selection is empty, for all countries.
country_subset = []
#country_subset = ['DE', 'FR', 'IT', 'ES']
# +
data_selection = pd.DataFrame()
if len(country_subset) == 0:
data_selection = data
else:
for country in country_subset:
if len(data_selection) == 0:
data_selection = data[data.country == country]
else:
data_selection = data_selection.append(data[data.country == country])
#data_selection
# -
# ## 5.1 Calculation of total capacity for energy source levels
# +
# Define the columns for grouping
groupby_selection = ['capacity_definition', 'source', 'year', 'type', 'country']
# Calculate the total capacity of all categories within a certain technology level
capacity_total_0 = pd.DataFrame(data_selection[data_selection['energy_source_level_0'] == True]
.groupby(groupby_selection)['capacity'].sum())
capacity_total_1 = pd.DataFrame(data_selection[data_selection['energy_source_level_1'] == True]
.groupby(groupby_selection)['capacity'].sum())
capacity_total_2 = pd.DataFrame(data_selection[data_selection['energy_source_level_2'] == True]
.groupby(groupby_selection)['capacity'].sum())
capacity_total_3 = pd.DataFrame(data_selection[data_selection['energy_source_level_3'] == True]
.groupby(groupby_selection)['capacity'].sum())
capacity_total_tech = pd.DataFrame(data_selection[data_selection['technology_level'] == True]
.groupby(groupby_selection)['capacity'].sum())
# Merge calculated capacity for different technology levels
capacity_total_comparison = pd.DataFrame(capacity_total_0)
capacity_total_comparison = pd.merge(capacity_total_0, capacity_total_1,
left_index=True, right_index=True, how='left')
capacity_total_comparison = capacity_total_comparison.rename(columns={'capacity_x': 'energy source level 0',
'capacity_y': 'energy source level 1'})
capacity_total_comparison = pd.merge(capacity_total_comparison, capacity_total_2,
left_index=True, right_index=True, how='left')
capacity_total_comparison = pd.merge(capacity_total_comparison, capacity_total_3,
left_index=True, right_index=True, how='left')
capacity_total_comparison = capacity_total_comparison.rename(columns={'capacity_x': 'energy source level 2',
'capacity_y': 'energy source level 3'})
capacity_total_comparison = pd.merge(capacity_total_comparison, capacity_total_tech,
left_index=True, right_index=True, how='left')
capacity_total_comparison = capacity_total_comparison.rename(columns={'capacity': 'technology level'})
# Define sorting preferences
capacity_total_comparison = capacity_total_comparison.sortlevel(['country', 'year'])
capacity_total_comparison
# -
# ## 5.2 Identifcation of capacity differences for energy source levels
# Identification of differences between energy source levels for each country, source, and year. The difference is relative to the previous energy source level. Generally, differences between the energy source levels should be zero, but could differ in particular for ENTSO-E data.
capacity_total_difference = capacity_total_comparison.diff(periods=1, axis=1)
capacity_total_difference = capacity_total_difference[(capacity_total_difference['energy source level 1'] > 0.01) |
(capacity_total_difference['energy source level 1'] < -0.01) |
(capacity_total_difference['energy source level 2'] > 0.01) |
(capacity_total_difference['energy source level 2'] < -0.01) |
(capacity_total_difference['energy source level 3'] > 0.01) |
(capacity_total_difference['energy source level 3'] < -0.01)|
(capacity_total_difference['technology level'] > 0.01) |
(capacity_total_difference['technology level'] < -0.01)]
capacity_total_difference
| tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="KhJJX7bF55wA" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from google.colab import drive
# %matplotlib inline
from skimage import io, color
from skimage.transform import resize
from sklearn.model_selection import train_test_split
import tensorflow as tf
from keras.utils import to_categorical
import glob
import re
from tensorflow.python.keras import applications
from tensorflow.python.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.python.keras.models import Sequential, Model, load_model
from tensorflow.python.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, GlobalAveragePooling2D
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import ClusterCentroids
# + id="dneKnRCZ6NbC" colab_type="code" outputId="c484b255-1808-4d2b-a9fa-c7270fcbe077" executionInfo={"status": "ok", "timestamp": 1587355226235, "user_tz": 240, "elapsed": 436, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# mount drive on Google drive to access training data
# Ignore this if you don't use Google Colab
drive.mount("/content/drive")
# + id="QA3llDmf6Z1b" colab_type="code" colab={}
# Access training data in My Drive
train_path = "/content/drive/My Drive/ECE_542/TrainData-C2/"
# get training labels
train_labels = pd.read_csv("/content/drive/My Drive/ECE_542/TrainData-C2/TrainAnnotations.csv")
train_labels.sort_values(by=["file_name"], inplace=True)
train_files = glob.glob(train_path + "*.jpg")
train_files.sort()
# + [markdown] id="D92XpLjw6s09" colab_type="text"
# # Helper functions
# + id="ewodWw1Q6pT7" colab_type="code" colab={}
def extract_data(file_names, labels, size=None):
"""
Extract all images given list of file names and list of labels. Also resize
images according to user-defined size
Inputs:
- filenames: list of file paths to images
- labels: list of label of each image; order based on the order of filenames
Outputs:
- images: list of RGB images
- annotations: list of labels for the images
"""
images = []
annotations = []
for idx, f in enumerate(file_names):
img = io.imread(f)
if size is not None:
img = resize(img, (size, size), anti_aliasing=True)
images.append(img)
annotations.append(labels[idx])
return images, annotations
# + id="3WEn8i6hpn__" colab_type="code" colab={}
def RGB2HSV(images, hue=False):
"""
Convert all RGB images into HSV channel
Input:
- images: list of images of shape (H, W, 3)
"""
hsv = []
for img in images:
if hue:
hsv.append(color.rgb2hsv(img)[:,:,0])
else:
hsv.append(color.rgb2hsv(img))
return hsv
# + [markdown] id="gAdfzWhSpocq" colab_type="text"
# # Preparing data and training model
# + id="qHSRkmv165_V" colab_type="code" colab={}
# This cell may take awhile to run
# extract all data
images, labels = extract_data(train_files, train_labels.annotation, 224)
# convert RGB into HSV
images = RGB2HSV(images, hue=True)
# + id="yD3o17D768YE" colab_type="code" outputId="ba349a38-3d95-4b05-bf2f-b3a98d565b6d" executionInfo={"status": "ok", "timestamp": 1587355597425, "user_tz": 240, "elapsed": 39455, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 241}
# Split training and validation test set
Y = to_categorical(labels)
X = np.array(images)
X_train, X_val, Y_train, Y_val = train_test_split(X, Y, test_size=0.3, random_state=32)
# prepare data for oversampling and undersampling
#n_train, H, W, C = X_train.shape # dimension of training data
n_train, H, W = X_train.shape # dimension of training data in hue space
n_val = len(X_val) # number of samples in validation set
#X_train = np.reshape(X_train, (n_train, H*W*C))
#X_val = np.reshape(X_val, (n_val, H*W*C))
X_train = np.reshape(X_train, (n_train, H*W))
X_val = np.reshape(X_val, (n_val, H*W))
# oversampling the train set
oversample = SMOTE(random_state=32)
X_train, Y_train = oversample.fit_resample(X_train, Y_train)
# undersampling the validation set
cc = ClusterCentroids(random_state=32)
X_val, Y_val = cc.fit_resample(X_val, Y_val)
# reshape training and validation data to prepare for training
#X_val = np.reshape(X_val, (len(X_val), H, W, C))
#X_train = np.reshape(X_train, (len(X_train), H, W, C))
X_val = np.reshape(X_val, (len(X_val), H, W, 1))
X_train = np.reshape(X_train, (len(X_train), H, W, 1))
# Checking the shape of training and validation data
print(X_val.shape)
print(X_train.shape)
# + id="_cEcjo_W7bgL" colab_type="code" outputId="ed0f6dcf-60d8-4607-871b-d41260f9de8b" executionInfo={"status": "ok", "timestamp": 1587355734265, "user_tz": 240, "elapsed": 237, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# class distribution of training and validation sets
print(np.sum(Y_train, axis=0))
print(np.sum(Y_val, axis=0))
# + id="L6BLL-X37hVG" colab_type="code" colab={}
# create, compile and training resNet50 model
base_model = applications.resnet.ResNet50(weights=None,
include_top=False,
input_shape=(224, 224, 1)) # 1 for HUE, 3 for HSV or RGB
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dropout(0.7)(x)
predictions = Dense(5, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=predictions)
# compile model
sgd = SGD(lr=0.01, momentum=0.9)
model.compile(optimizer=sgd,
loss='categorical_crossentropy',
metrics=['accuracy'])
# + id="IEERkeE27ntG" colab_type="code" outputId="de83362c-9b22-499f-9593-790463723580" executionInfo={"status": "ok", "timestamp": 1587355740854, "user_tz": 240, "elapsed": 242, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.summary()
# + id="i9yI6AI47pqe" colab_type="code" outputId="fd554dc1-04e2-4a51-f3d5-b55e71257431" executionInfo={"status": "ok", "timestamp": 1587357656430, "user_tz": 240, "elapsed": 1912176, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 527}
# train model
history4 = model.fit(X_train, Y_train,
epochs=15,
batch_size=32,
validation_data=(X_val, Y_val))
# + id="6r49YX9L7uUG" colab_type="code" outputId="4a8515ae-1b99-468b-93a8-004f239f12be" executionInfo={"status": "ok", "timestamp": 1587357664240, "user_tz": 240, "elapsed": 677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 315}
# plot learning curves
accuracy = history4.history['accuracy']
val_accuracy = history4.history['val_accuracy']
loss = history4.history['loss']
val_loss = history4.history['val_loss']
epochs = range(len(accuracy))
plt.plot(epochs, accuracy, 'b-', label='Training accuracy')
plt.plot(epochs, val_accuracy, 'r-', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
# + id="9WQOG9ZwF-Bi" colab_type="code" outputId="3b67d210-7822-4ec6-d23e-3a1e1eac7a05" executionInfo={"status": "ok", "timestamp": 1587357670987, "user_tz": 240, "elapsed": 349, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_nZvypFCMUCWAJbU1A0LZ3grEq7r-w0UN6Wcn=s64", "userId": "17349355310615593249"}} colab={"base_uri": "https://localhost:8080/", "height": 315}
plt.plot(epochs, loss, 'b-', label='Training loss')
plt.plot(epochs, val_loss, 'r-', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.figure()
# + [markdown] id="XOeA6SR571Lc" colab_type="text"
# # Make prediction on test set
# + id="BRgd3myw7yKu" colab_type="code" colab={}
# Loading the test set
# Access testing data in My Drive
test_path = "/content/drive/My Drive/ECE_542/TestData/"
test_files = glob.glob(test_path + "*.jpg")
test_files.sort()
# + id="nu-gtYAu73Sl" colab_type="code" colab={}
# Load all testing data and convert them into hsv
test_data = []
for f in test_files:
img = io.imread(f)
# rescale images to half original size
img = resize(img, (224, 224), anti_aliasing=True)
#hsv = color.rgb2hsv(img)
hue = color.rgb2hsv(img)[:,:,0]
test_data.append(hue)
# + id="n1s5v2P875rn" colab_type="code" colab={}
# making prediction
P = model.predict(np.array(test_data))
Yhat = np.argmax(P, axis=1)
# + id="rIzcU_Yv7-23" colab_type="code" colab={}
# save result
# Taken from Homework2b
def vectorize_result(nclass, j):
"""
Return a nclass-dimensional unit vector with 1.0 in the j-th position
and zero elsewhere
"""
e = np.zeros((nclass,1))
e[j] = 1.0
return e
# + id="AQtQTH8j8fmt" colab_type="code" colab={}
encode = [vectorize_result(5, Yhat[i]) for i in range(Yhat.shape[0])]
pred_df = pd.DataFrame(np.array(encode).reshape((Yhat.shape[0], 5)).astype(np.uint8))
# Save predictions to csv
pred_df.to_csv("/content/drive/My Drive/ECE_542/prediction_final_hue.csv", header=False, index=False)
# + id="OTUjhTPuHZ1C" colab_type="code" colab={}
model.save("/content/drive/My Drive/ECE_542/resnet_SGD_final_hue.h5")
| DL_classification/sourcecode/ProjC2_resnet_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Kv0jZezPSORM"
# !python -m pip install schedule
# !python -m pip install pystan
# !python -m pip install fbprophet
# !python -m pip install finance-datareader
# + id="wzzOn-ULWESk"
import os
import time
import schedule
import numpy as np
import pandas as pd
import tensorflow as tf
import FinanceDataReader as fdr
import matplotlib.pyplot as plt
from time import sleep
from fbprophet import Prophet
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM, Conv1D, Lambda
from tensorflow.keras.losses import Huber
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
# + id="GH7Gs_GvYir-"
data = pd.read_excel('./samsung.xlsx')
# + id="eLX0LYErn7u9"
def windowed_dataset(series, window_size, batch_size, shuffle):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
if shuffle:
ds = ds.shuffle(1000)
ds = ds.map(lambda w: (w[:-1], w[-1]))
return ds.batch(batch_size).prefetch(1)
# + id="VYhyzKc_xTwP"
def pred_machine(data):
scaler = MinMaxScaler()
scale_cols = list(data.columns[1:])
scaled = scaler.fit_transform(data[scale_cols])
df = pd.DataFrame(scaled, columns=scale_cols)
x_train, x_test, y_train, y_test = train_test_split(df.drop('y', 1), df['y'], test_size=0.2, random_state=0, shuffle=False)
WINDOW_SIZE=120
BATCH_SIZE=32
train_data = windowed_dataset(y_train, WINDOW_SIZE, BATCH_SIZE, True)
test_data = windowed_dataset(y_test, WINDOW_SIZE, BATCH_SIZE, False)
model = Sequential([
Conv1D(filters=32, kernel_size=5,
padding="causal",
activation="relu",
input_shape=[WINDOW_SIZE, 1]),
LSTM(16, activation='tanh'),
Dense(16, activation="relu"),
Dense(1),
])
loss = Huber()
optimizer = Adam(0.0005)
model.compile(loss=Huber(), optimizer=optimizer, metrics=['mse'])
earlystopping = EarlyStopping(monitor='val_loss', patience=100, mode='min')
filename = os.path.join('tmp', 'ckeckpointer.ckpt')
checkpoint = ModelCheckpoint(filename,
save_weights_only=True,
save_best_only=True,
monitor='val_loss',
verbose=1)
history = model.fit(train_data,
validation_data=(test_data),
epochs=500,
callbacks=[checkpoint, earlystopping])
for i in range(10):
merge_data = pd.DataFrame()
for col in data:
if col != 'DATE' and col != 'y':
data_copy = data[['DATE', col, 'DATE']].copy()
data_copy.columns = ['ds', 'y', 'DATE']
data_copy = data_copy.set_index('DATE')
prophet = Prophet(seasonality_mode='multiplicative',
yearly_seasonality=True,
weekly_seasonality=True, daily_seasonality=True,
changepoint_prior_scale=0.5)
prophet.fit(data_copy)
future_data = prophet.make_future_dataframe(periods=1, freq='d')
forecast_data = prophet.predict(future_data)
forecast_copy = pd.DataFrame(forecast_data[['ds', 'yhat']].tail(1))
forecast_copy.columns = ['DATE', col]
merge_data[col] = forecast_copy[col]
merge_data['DATE'] = forecast_copy['DATE']
df_row = pd.concat([data, merge_data])
pred_scaled = scaler.fit_transform(df_row[scale_cols])
pred_df = pd.DataFrame(pred_scaled, columns=scale_cols)
p_x_train, p_x_test, p_y_train, p_y_test = train_test_split(pred_df.drop('y', 1), pred_df['y'], test_size=0.2, random_state=0, shuffle=False)
WINDOW_SIZE=120
BATCH_SIZE=32
pred_train = windowed_dataset(p_y_train, WINDOW_SIZE, BATCH_SIZE, True)
pred_test = windowed_dataset(p_y_test, WINDOW_SIZE, BATCH_SIZE, False)
pred = model.predict(pred_test)
pred_df.iloc[-1]['y'] = pred[-1]
data = scaler.inverse_transform(pred_df)
data = pd.DataFrame(data, columns=scale_cols)
data['DATE'] = df_row['DATE']
data = data[['DATE', '거래량', 'PER', 'PBR', '기관 합계', '기타법인', '개인', '외국인 합계', 'ATR',
'NASDAQ', 'S&P', 'CBOE', 'Exchange rate', 'futures2y', 'futures10y',
'y']]
return(data)
# + id="Fbri8J2QIXyD"
pred_machine(data)
# + id="GllzW2MyZNEV"
schedule.every().day.at("6:00").do(pred_machine, data)
data = pred_machine(data)
print(data)
while True:
schedule.run_pending()
data = data
print(data)
time.sleep(1)
# + id="88Wlf9RYKZ7L"
| modeling/notebook/pred_machine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Plotting
#
# In this quickstart, we'll show all kind of examples. As PySport encourages OpenSource projects, all examples use other sports analytics packages in combination with kloppy. You can find more packages at https://opensource.pysport.org/
#
#
# ## Plotting events using mplsoccer
#
# In this example the [mplsoccer](https://github.com/andrewRowlinson/mplsoccer) package by [<NAME>](https://twitter.com/numberstorm) is used.
#
import sys
# !{sys.executable} -m pip install mplsoccer matplotlib seaborn
# +
## Load data
from mplsoccer.pitch import Pitch
from kloppy import statsbomb
dataset = statsbomb.load_open_data(
event_types=["pass"],
coordinates="statsbomb"
)
home_team, away_team = dataset.metadata.teams
messi = home_team.players[9]
print(f"Going to show passes of: {messi}")
# +
df = (
dataset
.filter(lambda event: event.player == messi)
.to_pandas()
)
pitch = Pitch(pitch_color='#e7f1fa', line_zorder=1, line_color='black', pitch_type="statsbomb")
fig, ax = pitch.draw()
plot = pitch.kdeplot(
df["coordinates_x"],
df["coordinates_y"],
ax=ax,
shade=True,
n_levels=50,
)
# -
| docs/examples/plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import glob
import copy
import numpy as np
import Bio
import scipy.spatial
import pickle
import matplotlib.pyplot as plt
import pandas as pd
from rnai_scripts import *
import bokeh.io
import bokeh.plotting
# Enable viewing Bokeh plots in the notebook
bokeh.io.output_notebook()
# -
def ecdf_vals(data):
"""Return x and y values for an ECDF."""
return np.sort(data), np.arange(1, len(data)+1) / len(data)
# # RNAi recoding
# ## Reading in the Smed transcriptome
# We read in the Smed_v6 transcriptome orfs that were extracted using orfipy. We then join them all into one string and obtain the codon frequencies.
# +
fname = 'data/dd_Smed_v6_transcripts_orfs_large3.fa' # makes smallest proteins be around 30 amino acids
descriptors, seqs = read_many_fasta(fname)
# join all ORFS into one large transcriptome
transcriptome = ''.join(seqs)
# get aminoacidweights and codon weights
codon_frequencies_dic = get_codon_frequencies(transcriptome)
# -
# Now we get frequencies of doublets
doubletscode = get_codon_frequencies_doublets(transcriptome)
# I also found a published version of amino acid frequencies:
# +
df = pd.read_csv('data/codon_usage_smed.csv')
AAs = df['codon'].values
freqs = df['frequency'].values/1000.
codon_frequencies_dic_published = {}
for i in range(len(AAs)):
codon_frequencies_dic_published[AAs[i]] = freqs[i]
print(sum(freqs))
# -
# Let's calculate the average discrepency between the doublets vs. codon frequencies.
diff_published_vs_me = {}
for a in AAs:
diff_published_vs_me[a] = codon_frequencies_dic_published[a] - codon_frequencies_dic[a]
values = np.array(list(diff_published_vs_me.values()))
print(np.mean(values))
print(np.mean(np.abs(values))) # values usually on order
print(np.sum(np.abs(values)))
# Here we find the discrepencies between the frequencies of each doublet vs. the product frequency of the separate codons.
# +
diff_dic = {}
diff_dic_norm = {}
for pair in doubletscode.keys():
if 'TAA' == pair[:3]:
continue
if 'TAG' == pair[:3]:
continue
if 'TGA' == pair[:3]:
continue
freq1 = codon_frequencies_dic[pair[:3]]
freq2 = codon_frequencies_dic[pair[3:]]
diff_dic_norm[pair] = (doubletscode[pair] - freq1*freq2)/np.max(np.array([freq1, freq2]))
diff_dic[pair] = (doubletscode[pair] - freq1*freq2)
# +
# Make figure
p = bokeh.plotting.figure(
frame_width=400,
frame_height=300,
x_axis_label='diff',
y_axis_label='Dist',
# x_axis_type = 'log'
)
diffs, ecdf_diffs = ecdf_vals(np.array(list(diff_dic.values())))
print(np.sum(np.array(list(doubletscode.values()))))
p.circle(diffs*1e4, ecdf_diffs)
#diffs, ecdf_diffs = ecdf_vals(np.array(list(doublets.values())))
#p.circle(diffs, ecdf_diffs, color = 'orange')
bokeh.io.show(p)
# +
# Make figure
p = bokeh.plotting.figure(
frame_width=400,
frame_height=300,
x_axis_label='diff',
y_axis_label='Dist',
# x_axis_type = 'log'
)
diffs, ecdf_diffs = ecdf_vals(np.array(list(diff_dic_norm.values())))
print(np.sum(np.array(list(doubletscode.values()))))
p.circle(diffs, ecdf_diffs)
#diffs, ecdf_diffs = ecdf_vals(np.array(list(doublets.values())))
#p.circle(diffs, ecdf_diffs, color = 'orange')
bokeh.io.show(p)
# -
values = np.array(list(diff_dic_norm.values()))
inds_sort = np.argsort(values)
keys = np.array(list(diff_dic_norm.keys()))
keys[inds_sort][:100]
values = np.array(list(diff_dic.values()))*1e4
inds_sort = np.argsort(values)
keys = np.array(list(diff_dic.keys()))
keys[inds_sort][:100]
diff_dic['AAAAAA']*1e4
doubletscode['AAAAAA']
codon_frequencies_dic['AAA']*codon_frequencies_dic['AAA']
# We use our codon frequencies dictionary to compute CAI weights (based on the weight definition for the CAI) for all codons
#
# $$w_i = \frac{f_i}{\max (f_j)} i,j \in [ \text{synonymouse codons for amino acid} ]$$
#
# Where $f_i$ is the frequency of codon $i$.
#
# We obtain two dictionaries:
#
#
# aminoacidweights: keys are amino acids, values are arrays of $w_i$ for all synonymous codons. The order of the codons is the as those used in aminoacidcode.
#
# gencodeweights: keys are codons, values are $w_i$ for each codon
aminoacidweights, gencodeweights = get_codon_weights(codon_frequencies_dic)
# We pickle dump everything so we do not have to repeat the above line later.
pickle.dump( aminoacidweights,
open( "data/Smed_transcriptome_aminoacidweights.p", "wb" ) )
pickle.dump( gencodeweights,
open( "data/Smed_transcriptome_gencodeweights.p", "wb" ) )
pickle.dump( aminoacidcode,
open( "data/aminoacidcode.p", "wb" ))
pickle.dump( doubletscode,
open( "data/doubletscode.p", "wb" ))
# We reload everything with pickle because why not.
aminoacidweights = pickle.load( open( "data/Smed_transcriptome_aminoacidweights.p",
"rb" ) )
gencodeweights = pickle.load( open( "data/Smed_transcriptome_gencodeweights.p",
"rb" ) )
aminoacidcode = pickle.load(open("data/aminoacidcode.p", 'rb'))
doubletscode = pickle.load(
open( "data/doubletscode.p", "rb" ))
# ## We recode the luc ORFS!!!!
#
# Since SmedNluc2 is so short we must RNAi the whole thing.
SmedNluc2_ORF = 'ATGGTGTTTACTTTGGAAGATTTTGTTGGAGATTGGAGACAAACTGCTGGTTACAATCTGGATCAGGTACTGGAACAAGGCGGTGTTAGTTCATTATTCCAAAACCTGGGTGTGAGTGTAACTCCGATTCAGCGAATAGTGTTGTCTGGAGAAAATGGGCTGAAGATTGATATACACGTCATAATTCCATACGAAGGCTTAAGCGGTGATCAAATGGGACAAATTGAAAAAATTTTTAAAGTAGTTTACCCAGTTGACGACCATCATTTTAAAGTTATCCTTCATTACGGTACACTGGTTATAGATGGTGTAACTCCAAATATGATCGATTATTTCGGAAGACCTTACGAAGGCATAGCCGTTTTTGATGGAAAAAAGATTACAGTAACAGGTACATTGTGGAACGGAAATAAGATTATTGACGAACGTTTAATTAACCCAGATGGAAGTTTGCTCTTTAGAGTTACAATTAATGGTGTGACAGGATGGAGATTATGCGAACGGATACTCGCGTAA'
SmedNluc2_protein = 'MVFTLEDFVGDWRQTAGYNLDQVLEQGGVSSLFQNLGVSVTPIQRIVLSGENGLKIDIHVIIPYEGLSGDQMGQIEKIFKVVYPVDDHHFKVILHYGTLVIDGVTPNMIDYFGRPYEGIAVFDGKKITVTGTLWNGNKIIDERLINPDGSLLFRVTINGVTGWRLCERILA*'
Hluc_ORF = 'ATGGTCTTCACACTCGAAGATTTCGTTGGGGACTGGCGACAGACAGCCGGCTACAACCTGGACCAAGTCCTTGAACAGGGAGGTGTGTCCAGTTTGTTTCAGAATCTCGGGGTGTCCGTAACTCCGATCCAAAGGATTGTCCTGAGCGGTGAAAATGGGCTGAAGATCGACATCCATGTCATCATCCCGTATGAAGGTCTGAGCGGCGACCAAATGGGCCAGATCGAAAAAATTTTTAAGGTGGTGTACCCTGTGGATGATCATCACTTTAAGGTGATCCTGCACTATGGCACACTGGTAATCGACGGGGTTACGCCGAACATGATCGACTATTTCGGACGGCCGTATGAAGGCATCGCCGTGTTCGACGGCAAAAAGATCACTGTAACAGGGACCCTGTGGAACGGCAACAAAATTATCGACGAGCGCCTGATCAACCCCGACGGCTCCCTGCTGTTCCGAGTAACCATCAACGGAGTGACCGGCTGGCGGCTGTGCGAACGCATTCTGGCGTAA'
# I wonder what the CAI for each ORF is?
print('CAI for SMed Nuc:', get_CAI(SmedNluc2_ORF, gencodeweights))
print('CAI for Human Nuc:', get_CAI(Hluc_ORF, gencodeweights))
print('Hamming Distance vs Smed vs Human Nuc', get_hamming_dist(SmedNluc2_ORF, Hluc_ORF))
# Now we can use the function get_RNAi_seq to randomly sample different recoded Luc proteins.
#
# The function get_RNAi_seq requires the ORF, protein sequence, an aminoacidweights and gencodeweights dictionary. We run 1000 random samples and do not enforce that every codon be different. It returns the list of tested sequences (seqs), scores ($CAI + D$/2) for each sequence, codon adaptation indices (CAIs), and Hamming distances (dists = $D$).
def get_doublest_likelihood(dna_seq, weights_dic):
'''
Obtains Codon Adaptation Index (CAI) for a given DNA_seq calculated using weights_dic
CAI = (w_1*.w_i*..w_N)^(1/N) where w_i is the weight of codon i.
Inputs:
dna_seq: ORF in form of string to evaluate CAI
weights_dic: dictionary of CAI weights for each codon. Values are weights and keys are codons.
'''
if len(dna_seq) % 3 > 0.:
raise ValueError("Length of DNA sequence must be divisble by 3")
ncodons = int(len(dna_seq)//3)
score = 0.
for i in range(ncodons-1):
start = i*3
end = start + 6
codonpair = dna_seq[start:end].upper()
score = score+ np.log(weights_dic[codonpair])
return score
# +
seqs, scores, cais, dists = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1000, enforce_different_codons = False, random = True)
best_seq, best_score, best_cai, best_dist = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1, enforce_different_codons = False, random = False)
best_doublet = get_doublest_likelihood(best_seq[0], doubletscode)
doublets_scores = np.array([get_doublest_likelihood(seq, doubletscode) for seq in seqs])
print(best_cai, best_dist, best_doublet)
# -
# We redo the process but enforce that every codon must be different.
# +
seqs_diff, scores_diff, cais_diff, dists_diff = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1000, enforce_different_codons = True, random = True)
best_seq_diff, best_score_diff, best_cai_diff, best_dist_diff = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1, enforce_different_codons = True, random = False)
best_doublet_diff = get_doublest_likelihood(best_seq_diff[0], doubletscode)
doublets_scores_diff = np.array([get_doublest_likelihood(seq, doubletscode) for seq in seqs_diff])
print(best_cai_diff, best_dist_diff, best_doublet_diff)
# -
# We find the best sequences of our random simulation
print(np.max(cais_diff), np.max(dists_diff))
# We repeat with wiggle.
# +
seqs_diff, scores_diff, cais_wiggle, dists_wiggle = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1000, enforce_different_codons = True, random = True, wiggle = True,)
best_seq_diff, best_score_diff, best_cai_diff_wiggle, best_dist_diff_wiggle = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1, enforce_different_codons = True, random = False, wiggle = True
)
best_doublet_diff_wiggle = get_doublest_likelihood(best_seq_diff[0], doubletscode)
doublets_scores_wiggle = np.array([get_doublest_likelihood(seq, doubletscode) for seq in seqs_diff])
print(best_cai_diff_wiggle, best_dist_diff_wiggle, best_doublet_diff_wiggle)
# -
print(np.max(cais_wiggle), np.max(dists_wiggle))
# Doublets baby
# +
seqs_doub, scores_doub, cais_doub, dists_doub = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1000, enforce_different_codons =True, random = True,
pairs = True, doubletscode = doubletscode)
best_seq_doub, best_score_doub, best_cai_doub, best_dist_doub = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1, enforce_different_codons = True, random = False,
pairs = True, doubletscode = doubletscode,)
best_doublet_doub = get_doublest_likelihood(best_seq_doub[0], doubletscode)
doublets_scores_doub= np.array([get_doublest_likelihood(seq, doubletscode) for seq in seqs_doub])
print(best_cai_doub, best_dist_doub, best_doublet_doub)
# +
seqs_doub, scores_doub, cais_doub_wigg, dists_doub_wigg = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1000, enforce_different_codons =True, random = True, wiggle = True,
pairs = True, doubletscode = doubletscode)
best_seq_doub, best_score_doub, best_cai_doub_wiggle, best_dist_doub_wiggle = get_RNAi_seq(SmedNluc2_ORF, SmedNluc2_protein, aminoacidweights,
gencodeweights, trials = 1, enforce_different_codons = True, random = False, wiggle = True,
pairs = True, doubletscode = doubletscode,)
best_doublet_doub = get_doublest_likelihood(best_seq_doub[0], doubletscode)
doublets_scores_doub_wigg = np.array([get_doublest_likelihood(seq, doubletscode) for seq in seqs_doub])
print(best_cai_doub_wiggle, best_dist_doub_wiggle, best_doublet_doub)
# -
# We define a function to compute ECDFs
# We plot ECDFs of the CAIs.
# +
# Make figure
p = bokeh.plotting.figure(
frame_width=400,
frame_height=300,
x_axis_label='CAI',
y_axis_label='ECDF',
)
cais, ecdf_cais = ecdf_vals(cais)
p.circle(cais, ecdf_cais, legend_label = 'Not all different ')
cais_diff, ecdf_cais_diff = ecdf_vals(cais_diff)
p.circle(cais_diff, ecdf_cais_diff, legend_label = 'all different', color = 'orange')
cais_wiggle, ecdf_cais_wiggle = ecdf_vals(cais_wiggle)
p.circle(cais_wiggle, ecdf_cais_wiggle, legend_label = 'all different wiggle', color = 'green')
cais_doub, ecdf_cais_doub = ecdf_vals(cais_doub)
p.circle(cais_doub, ecdf_cais_doub, legend_label = 'doublets', color = 'red')
cais_doub_wiggle, ecdf_cais_doub_wiggle = ecdf_vals(cais_doub_wigg)
p.circle(cais_doub_wiggle, ecdf_cais_doub_wiggle,
legend_label = 'doublets wig', color = 'pink')
p.legend.location = 'bottom_right'
bokeh.io.show(p)
# -
# We plot ECDFs of the hamming distances
# +
# Make figure
p = bokeh.plotting.figure(
frame_width=400,
frame_height=300,
x_axis_label='Hamming Distance',
y_axis_label='ECDF',
)
dists, ecdf_dists = ecdf_vals(dists)
p.circle(dists, ecdf_dists, legend_label = 'Not all different ')
dists_diff, ecdf_dists_diff = ecdf_vals(dists_diff)
p.circle(dists_diff, ecdf_dists_diff, legend_label = 'all different', color = 'orange')
dists_diff_wiggle, ecdf_dists_diff_wiggle = ecdf_vals(dists_wiggle)
p.circle(dists_diff_wiggle, ecdf_dists_diff_wiggle, legend_label = 'wiggle', color = 'green')
dists_doub, ecdf_dists_doub = ecdf_vals(dists_doub)
p.circle(dists_doub, ecdf_dists_doub, legend_label = 'doublets', color = 'red')
dists_doub_wiggle, ecdf_dists_doub_wiggle = ecdf_vals(dists_doub_wigg)
p.circle(dists_doub_wiggle, ecdf_dists_doub_wiggle,
legend_label = 'doublets wig', color = 'pink')
p.legend.location = 'bottom_right'
p.x_range = bokeh.models.Range1d(.1, .6)
bokeh.io.show(p)
# +
# Make figure
p = bokeh.plotting.figure(
frame_width=400,
frame_height=300,
x_axis_label='Hamming Distance',
y_axis_label='ECDF',
)
dists, ecdf_dists = ecdf_vals(doublets_scores)
p.circle(dists, ecdf_dists, legend_label = 'Not all different ')
dists_diff, ecdf_dists_diff = ecdf_vals(doublets_scores_diff)
p.circle(dists_diff, ecdf_dists_diff, legend_label = 'all different', color = 'orange')
dists_diff_wiggle, ecdf_dists_diff_wiggle = ecdf_vals(doublets_scores_wiggle)
p.circle(dists_diff_wiggle, ecdf_dists_diff_wiggle, legend_label = 'wiggle', color = 'green')
dists_doub, ecdf_dists_doub = ecdf_vals(doublets_scores_doub)
p.circle(dists_doub, ecdf_dists_doub, legend_label = 'doublets', color = 'red')
dists_doub_wiggle, ecdf_dists_doub_wiggle = ecdf_vals(doublets_scores_doub_wigg)
p.circle(dists_doub_wiggle, ecdf_dists_doub_wiggle,
legend_label = 'doublets wig', color = 'pink')
p.legend.location = 'bottom_right'
bokeh.io.show(p)
# -
| recoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py39_spgh_dev]
# language: python
# name: conda-env-py39_spgh_dev-py
# ---
# ---------------
#
# **If any part of this notebook is used in your research, please cite with the reference found in** **[README.md](https://github.com/pysal/spaghetti#bibtex-citation).**
#
#
# ----------------
# # Quickstart
# ## Creating and visualizing a `spaghetti.Network` object
#
# **Author: <NAME>** **<<EMAIL>>**
#
# **This notebook provides an explanation of network creation followed by an emprical example for:**
#
# 1. Instantiating a network
# 2. Allocating observations to a network (snapping points)
# 3. Visualizing the original and network-snapped locations with `geopandas` and `matplotlib`
# %config InlineBackend.figure_format = "retina"
# %load_ext watermark
# %watermark
import geopandas
import libpysal
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import matplotlib_scalebar
from matplotlib_scalebar.scalebar import ScaleBar
import shapely
import spaghetti
# %matplotlib inline
# %watermark -w
# %watermark -iv
# --------------------------------
# ## The basics of `spaghetti`
# ### 1. Creating a network instance
#
# Spatial data science techniques can support many types of statistical analyses of spatial networks themselves, and of events that happen along spatial networks in our daily lives, i.e. locations of trees along foot paths, biking accidents along street networks or locations of coffeeshops along streets. `spaghetti` provides computational tools to support statistical analysis of such events along many different types of networks. Within `spaghetti` network objects can be created from [a variety of objects](https://pysal.org/spaghetti/generated/spaghetti.Network.html#spaghetti-network), the most common being shapefiles (read in as file paths) and `geopandas.GeoDataFrame` objects. However, a network could also be created from `libpysal` geometries, as demonstrated in the [connected components tutorial](https://pysal.org/spaghetti/notebooks/connected-components.html#1.-Instantiate-a-network-from-two-collections-of-libpysal.cg.Chain-objects) or a simply as follows:
#
# ```python
# from libpysal.cg import Point, Chain
# import spaghetti
# # create the network
# ntw = spaghetti.Network(in_data=Chain([Point([1, 1]), Point([2, 1])]))
# ```
#
# This will create a single-segment network, which is simply one single line. Although the chances of a single-segment network existing in reality are rare, it is useful for demonstration purposes.
#
# The stucture and characterstics of the networks, can be quantitatively described with `spaghetti` and are topics of research in many areas. However, networks are also utilized as the study space, containing observations or events of interest, in many applications. In these cases the actual objects of interest that will be analysed in the geographic space a network provides, are "network-based events."
#
#
# ### 2. Snapping events (points) to a network
#
# First, point objects, representing our network-based events, must be snapped to the network for meaningful spatial analysis to be done or models to be constructed. As with `spaghetti.Network` objects, `spaghetti.PointPattern` objects can be [created from](https://pysal.org/spaghetti/generated/spaghetti.PointPattern.html#spaghetti.PointPattern) shapefiles and `geopandas.GeoDataFrame` objects. Furthermore, `spaghetti` can also simply handle a single `libpysal.cg.Point `object. Considering the single-segment network above:
#
# ```python
# # create the point and snap it to the network
# ntw.snapobservations(Point([1.5, 1.1]), "point")
# ```
#
# At this point the point is associated with the network and, as such, is defined in network space.
#
#
# ### 3. Visualizating the data
#
# Visualization is a cornerstone in communicating scientific data. Within the context of `spaghetti` elements of the network must be [extracted](https://pysal.org/spaghetti/generated/spaghetti.element_as_gdf.html#spaghetti.element_as_gdf) as `geopandas.GeoDataFrame` objects prior to being visualized with `matplotlib`. This is shown in the following block of code, along with network creation and point snapping.
# +
from libpysal.cg import Point, Chain
import spaghetti
# create the network
ntw = spaghetti.Network(in_data=Chain([Point([1, 1]), Point([2, 1])]))
# create the point and snap it to the network
ntw.snapobservations(Point([1.5, 1.1]), "point")
# network nodes and edges
vertices_df, arcs_df = spaghetti.element_as_gdf(ntw, vertices=True, arcs=True)
# true and snapped location of points
point_df = spaghetti.element_as_gdf(ntw, pp_name="point", snapped=False)
snapped_point_df = spaghetti.element_as_gdf(ntw, pp_name="point", snapped=True)
# plot the network and point
base = arcs_df.plot(figsize=(10,10), color="k", alpha=0.25, zorder=0)
vertices_df.plot(ax=base, color="k", alpha=1)
kwargs = {"ax":base, "alpha":0.5, "zorder":1}
point_df.plot(color="b", marker="x", **kwargs)
snapped_point_df.plot(color="b", markersize=20, **kwargs)
plt.xlim(.9,2.1); plt.ylim(.8,1.2);
# -
# Network creation, observation snapping, and visualization are further reviewed below for an example with empirical datasets available `libpysal`.
#
#
# ------------------------------
# ## Empirical Example
#
# In the following we will walk through an empirical example, visually comparing school locations with a network to crimes committed within the same network.
#
# ### 1. Instantiating a `spaghetti.Network` object
# #### Instantiate the network from a `.shp` file
ntw = spaghetti.Network(in_data=libpysal.examples.get_path("streets.shp"))
# ------------------------------
# ### 2. Allocating observations to a network:
# #### Schools without attributes
ntw.snapobservations(
libpysal.examples.get_path("schools.shp"), "schools", attribute=False
)
# #### True vs. snapped school coordinates comparison: `spaghetti.Network` attributes
print("observation 1\ntrue coords:\t%s\nsnapped coords:\t%s" % (
ntw.pointpatterns["schools"].points[0]["coordinates"],
ntw.pointpatterns["schools"].snapped_coordinates[0]
))
# #### Crimes with attributes
ntw.snapobservations(
libpysal.examples.get_path("crimes.shp"), "crimes", attribute=True
)
# #### True vs. snapped crime coordinates comparison: `spaghetti.Network` attributes
print("observation 1\ntrue coords:\t%s\nsnapped coords:\t%s" % (
ntw.pointpatterns["crimes"].points[0]["coordinates"],
ntw.pointpatterns["crimes"].snapped_coordinates[0]
))
# ------------------------------
# ### 3. Visualizing original and snapped locations
# #### True and snapped school locations
true_schools_df = spaghetti.element_as_gdf(
ntw, pp_name="schools", snapped=False
)
snapped_schools_df = spaghetti.element_as_gdf(
ntw, pp_name="schools", snapped=True
)
# #### True vs. snapped school coordinates comparison: `geopandas.GeoDataFrame`
# Compare true point coordinates & snapped point coordinates
print("observation 1\ntrue coords:\t%s\nsnapped coords:\t%s" % (
true_schools_df.geometry[0].coords[:][0],
snapped_schools_df.geometry[0].coords[:][0]
))
# #### True and snapped crime locations
true_crimes_df = spaghetti.element_as_gdf(
ntw, pp_name="crimes", snapped=False
)
snapped_crimes_df = spaghetti.element_as_gdf(
ntw, pp_name="crimes", snapped=True
)
# #### True vs. snapped crime coordinates comparison: `geopandas.GeoDataFrame`
print("observation 1\ntrue coords:\t%s\nsnapped coords:\t%s" % (
true_crimes_df.geometry[0].coords[:][0],
snapped_crimes_df.geometry[0].coords[:][0]
))
# #### Create `geopandas.GeoDataFrame` objects of the vertices and arcs
# network nodes and edges
vertices_df, arcs_df = spaghetti.element_as_gdf(ntw, vertices=True, arcs=True)
# #### Create legend patches for the `matplotlib` plot
# +
# create legend arguments and keyword arguments for matplotlib
args = [], []
kwargs = {"c":"k"}
# set arcs legend entry
arcs = mlines.Line2D(*args, **kwargs, label="Network Arcs", alpha=0.5)
# update keyword arguments for matplotlib
kwargs.update({"lw":0})
# set vertices legend entry
vertices = mlines.Line2D(
*args, **kwargs, ms=2.5, marker="o", label="Network Vertices"
)
# -
# set true school locations legend entry
tschools = mlines.Line2D(
*args, **kwargs, ms=25, marker="X", label="School Locations"
)
# set network-snapped school locations legend entry
sschools = mlines.Line2D(
*args, **kwargs, ms=12, marker="o", label="Snapped Schools"
)
# +
# update keyword arguments for matplotlib
kwargs.update({"c":"r", "alpha":0.75})
# set true crimes locations legend entry
tcrimes = mlines.Line2D(
*args, **kwargs, ms=7, marker="x", label="Crime Locations"
)
# set network-snapped crimes locations legend entry
scrimes = mlines.Line2D(
*args, **kwargs, ms=3, marker="o", label="Snapped Crimes"
)
# -
# combine all legend patches
patches = [arcs, vertices, tschools, sschools, tcrimes, scrimes]
# #### Plotting `geopandas.GeoDataFrame` objects
# +
# set the streets as the plot base
base = arcs_df.plot(color="k", alpha=0.25, figsize=(12, 12), zorder=0)
# create vertices keyword arguments for matplotlib
kwargs = {"ax":base}
vertices_df.plot(color="k", markersize=5, alpha=1, **kwargs)
# update crime keyword arguments for matplotlib
kwargs.update({"alpha":0.5, "zorder":1})
true_crimes_df.plot(color="r", marker="x", markersize=50, **kwargs)
snapped_crimes_df.plot(color="r", markersize=20, **kwargs)
# update schools keyword arguments for matplotlib
kwargs.update({"cmap":"tab20", "column":"id", "zorder":2})
true_schools_df.plot(marker="X", markersize=500, **kwargs)
snapped_schools_df.plot(markersize=200, **kwargs)
# add scale bar
kw = {"units":"ft", "dimension":"imperial-length", "fixed_value":1000}
base.add_artist(ScaleBar(1, location="lower left", box_alpha=.75, **kw))
# add legend
plt.legend(
handles=patches,
fancybox=True,
framealpha=0.8,
scatterpoints=1,
fontsize="xx-large",
bbox_to_anchor=(1.04, 0.75),
borderpad=2.,
labelspacing=2.
);
# -
# -----------
| notebooks/quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Employee:
no_empl=0;
raise_amount=1.04
def __init__(self):
self.first="Soumyadip"
self.last="Chowdhury"
self.email="<EMAIL>"
self.pay=100000
Employee.no_empl=Employee.no_empl+1
def print(self):
print("Hello")
return "{} {} {} {}".format(self.first,self.last,self.email,self.pay)
emp_1=Employee()
print(Employee.raise_amount)
print(emp_1.raise_amount)
print(emp_1.first)
emp_1.print()
# -
| Employee Class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
df = pd.read_csv('time_data/walmart_stock.csv')
# Make Date as index column, same effect of convert string to datetime
# df = pd.read_csv('time_data/walmart_stock.csv', index_col='Date', parse_dates=True)
# -
df.head()
# Make Date Column to Index
df.info()
# The return shows Date Column Value is String instead of Datetime
df['Date'] = pd.to_datetime(df['Date']) # Convert String to Datetime
df.head()
df.info()
df['Date'] = df['Date'].apply(pd.to_datetime) # Same effect of Convert String to Datetime
# Make Date Column to the index
df.set_index('Date', inplace=True)
df.head()
df1 = pd.read_csv('time_data/walmart_stock.csv')
df1.info()
df = pd.read_csv('time_data/walmart_stock.csv', index_col='Date', parse_dates=True)
df.info()
df.index
# Acting group by with time series object
df.resample(rule='A') # year end frequently
df.head()
df.resample(rule='A').mean()
df.resample(rule='Q').mean()
df.resample(rule='A').max()
def first_day(entry):
return entry[0]
df.resample('A').apply(first_day)
df['Close'].resample('A')
df['Close'].resample('A').mean()
df['Close'].resample('A').mean().plot()
df['Close'].resample('A').mean().plot(kind='bar')
# A is Year End
df['Close'].resample('M').mean().plot(kind='bar')
# M is Month End
df['Close'].resample('M').mean().plot(kind='bar', figsize=(16,6))
| 05-Pandas-with-Time-Series/TimeSeries - Time Resampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Goal
# The objective of this project is to build one or more regression models to determine the scores for each team using the other columns as features. I will follow data preparation and feature engineering process before I use a regression model to predict the scores.
import pandas as pd #load the data into panda data frames
from sklearn import preprocessing #label encoder
from sklearn.model_selection import train_test_split #split the dataset
from sklearn import datasets, ensemble
from sklearn.inspection import permutation_importance
from sklearn.metrics import mean_squared_error
from sklearn.metrics import accuracy_score #generate accuracy score
from sklearn.ensemble import RandomForestRegressor
import matplotlib.pyplot as plt
from sklearn.metrics import r2_score
# ## Load the dataset
# I am using the [soccer dataset](https://github.com/fivethirtyeight/data/tree/master/soccer-spi) from FiveThirtyEight.
#
#
# When I viewed the full dataset, I noticed that some rows' scores are not provided. These games have not taken place yet and hence we only have access to their Soccer Power Index (SPI), which [ESPN describes](https://www.espn.com/soccer/news/story/_/id/1873765) as the "best possible representation of a team's current overall skill level". This observation will help us to split the data into training data and prediction data later.
#
# The first few entries in past dates have scores.
soccer_stats = pd.read_csv('../data/raw/spi_matches.csv')
soccer_stats.head(2)
# While future dates in 2021 do not have score data.
soccer_stats.tail(2)
soccer_stats.keys()
# I understood what the columns meant through some research on an [rdrr.io dataset page](https://rdrr.io/github/fivethirtyeightdata/fivethirtyeightdata/man/spi_matches.html)
#
# - **season:** Season of the soccer game.
#
# - **date:** The date that the match took place.
#
# - **league_id:** A numerical identifier of the league within which the match was played.
#
# - **league:** League name.
#
# - **team1:** One team that participated in the match.
#
# - **team2:** The other team that participated in the match.
#
# - **spi1:** The SPI score of team1.
#
# - **spi2:** The SPI score of team2.
#
# - **prob1:** The probability that team1 would have won the match.
#
# - **prob2:** The probability that team2 would have won the match.
#
# - **probtie:** The probability that the match would have resulted in a tie.
#
# - **proj_score1:** The predicted number of goals that team1 would have scored.
#
# - **proj_score2:** The predicted number of goals that team2 would have scored.
#
# - **score1:** The number of goals that team1 scored.
#
# - **score2:** The number of goals that team2 scored.
#
# The following columns did not have any description importance1, importance2, xg1, xg2, nsxg1, nsxg2, adj_score1, adj_score2 and seem to have NaN values for some rows.
# ## Data Preparation
# ### Cleaning
# Remove columns without descriptions. Since we do not have enough information about these columns, we cannot make educated decisions about how it affects the dataset. So, we drop them:
soccer_stats = soccer_stats.drop(columns=['importance1', 'importance2', 'xg1', 'xg2', 'nsxg1', 'nsxg2', 'adj_score1', 'adj_score2'])
# This is how it looks:
soccer_stats.head()
# ## Feature Engineering
# ### Extraction
# I separate the date column into three date, month and year. The season and year seem to not always align because a season may span between two years.
soccer_stats[['Year','Month', 'Date']] = soccer_stats.date.str.split("-",expand=True)
soccer_stats = soccer_stats.drop(columns=['date'])
soccer_stats.head()
# ### Transformation
# To perform statistical analysis and to pass the data into the regression models, I encode the strings into integer values as the models do not work with strings.
#
# **League Name**
league_encode = preprocessing.LabelEncoder()
league_encode.fit(soccer_stats['league'])
soccer_stats['league'] = league_encode.transform(soccer_stats['league'])
# **Team names**
# +
team1_encode = preprocessing.LabelEncoder()
team1_encode.fit(soccer_stats['team1'])
soccer_stats['team1'] = team1_encode.transform(soccer_stats['team1'])
team2_encode = preprocessing.LabelEncoder()
team2_encode.fit(soccer_stats['team2'])
soccer_stats['team2'] = team2_encode.transform(soccer_stats['team2'])
# -
# Encoded data looks like this:
soccer_stats
# ## Create Training and Predict Data
# My objective is to predict the scores for future games, by using the scores of the games that have already occured.
# - **Training data:** rows of games that have already occured, which contain the scores data
# - **Prediction data:** rows of future games without the scores date
predict_data = soccer_stats[soccer_stats['score1'].isnull()]
train_data = soccer_stats.dropna()
train_data.shape
predict_data.shape
# To ensure that I separated the data into test and training data, I check if the sum of the number of rows is equal to the rows in the original dataset
(34399+7775) == soccer_stats.shape[0]
# I remove the score columns entirely from the predict_data of the future games
predict_data = predict_data.drop(columns=['score1', 'score2'])
predict_data.head()
# ## Regression Models
# First, I split the data useing train_test_split, where the predicted output classes are the scores of the two teams and all other columns are features.
classes = train_data['score1']
features = train_data[['season', 'league_id', 'league', 'team1', 'team2', 'spi1', 'spi2', 'prob1', 'prob2', 'probtie', 'proj_score1', 'proj_score2', 'Year', 'Month', 'Date']]
X_train, X_test, Y_train, Y_test = train_test_split(features, classes, test_size=0.2, random_state=13)
expected_result = Y_test
# First, I use the random forest regressor which has many decision trees and an arbitary numbers of features for splitting. I use the Y_test data and prediction to compute the R^2 score, which is an accuracy measure of regression models. The following is the array of predicted scores:
forest = RandomForestRegressor(n_estimators=100)
forest.fit(X_train, Y_train)
pred_test_forest = forest.predict(X_test)
r2_score(Y_test.to_numpy(),pred_test_forest)
pred_test_forest
# The accuracy is only at about 8%. Let's try a different model- gradient boosting, which uses weak learners to create a strong learner, by adding a model at each iteration. I tuned it to have 500 boosting stages and get the following accuracy and predictions:
params = {'n_estimators': 500,
'max_depth': 4,
'min_samples_split': 5,
'learning_rate': 0.01,
'loss': 'ls'}
gradient_boost = ensemble.GradientBoostingRegressor(**params)
gradient_boost.fit(X_train, Y_train)
grad_test = gradient_boost.predict(X_test)
r2_score(Y_test.to_numpy(),grad_test)
grad_test
# Now, similar to generating a r^2 value when comparing the test values and predictions, I generate an r^2 value after predicting using the 'predict_data' which was the data of the future games. The R^2 value will be similar, so I use this to predict scores of games that have not happened.
pred_future = gradient_boost.predict(predict_data)
r2_score(Y_test, pred_test)
# **Predicted scores of team 1 of games in 2021:**
pred_future
# **Predicted scores of team 2 of games in 2021:**
# Similar to how I predicted the score for team 1, I predict team 2's score:
X_train, X_test, Y_train, Y_test = train_test_split(features, train_data['score2'], test_size=0.2, random_state=13)
gradient_boost.fit(X_train, Y_train)
grad_test_team2 = gradient_boost.predict(X_test)
grad_test_team2
# ## Visualization
# I plot the expected values and predicted values for both team's scores
# +
plt.scatter(grad_test, expected_result, alpha=0.40)
plt.title('Gradient Boosting Regression')
plt.xlabel('Prediction')
plt.ylabel('Expected')
plt.show()
plt.scatter(pred_test_forest, expected_result, alpha=0.40)
plt.title('Random Forest Regression')
plt.xlabel('Prediction')
plt.ylabel('Expected')
plt.show()
| notebooks/MajorLeagues.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 1: Spiral
# 
def solution(A):
"""O(N*M) solution"""
sum_up = 0
while True:
############
# Pass 1
# top row
for c in range(len(A[0])):
sum_up += A[0][c]
A.pop(0)
if len(A) == 0: break
# bottom row
A.pop(-1)
if len(A) == 0: break
###########
# Pass 2
# top row
for r in range(len(A)):
sum_up += A[r][-1]
A[r].pop(-1)
if len(A[0]) == 0: break
# bottom row
for r in range(len(A)-1, -1, -1):
A[r].pop(0)
if len(A[0]) == 0: break
##########
# Pass 3
# top row
for c in range(len(A[-1])-1, -1, -1):
sum_up += A[-1][c]
A.pop(-1)
if len(A) == 0: break
# bottom row
A.pop(0)
if len(A) == 0: break
##########
# Pass 4
# bottom row
for r in range(len(A)-1, -1, -1):
sum_up += A[r][0]
A[r].pop(0)
if len(A[0]) == 0: break
# top row
for r in range(len(A)):
A[r].pop(-1)
if len(A[0]) == 0: break
if sum_up < -100000000 or sum_up > 100000000:
return -1
else:
return sum_up
A = [[5, 3, 8, 9, 4, 1, 3, -2],
[4, 6, 0, 3, 6, 4, 2, 1],
[4, -5, 3, 1, 9, 5, 6, 6],
[3, 7, 5, 3, 2, 8, 9, 4],
[5, 3, -3, 6, 3, 2, 8, 0],
[5, 7, 5, 3, 3, -9, 2, 2],
[0, 4, 3, 2, 5, 7, 5, 4]]
solution(A)
# ### Unit Test
# +
# you can write to stdout for debugging purposes, e.g.
# print("this is a debug message")
def reference_solution(A):
upper, lower = [], []
while True:
#
# Pass TOP
#
# top row
for c in range(len(A[0])):
#print(A[0][c])
upper.append(A[0][c])
A.pop(0)
if len(A) == 0: break
# lower row
for c in range(len(A[-1])-1, -1, -1):
#print(A[-1][c])
lower.append(A[-1][c])
A.pop(-1)
if len(A) == 0: break
#print(upper)
#print(lower)
#
# Pass RIGHT
#
# top row
for r in range(len(A)):
#print(A[r][-1])
upper.append(A[r][-1])
A[r].pop(-1)
if len(A[0]) == 0: break
# lower row
for r in range(len(A)-1, -1, -1):
#print(A[r][0])
lower.append(A[r][0])
A[r].pop(0)
if len(A[0]) == 0: break
#print(upper)
#print(lower)
#
# Pass BOTTOM
#
# top row
for c in range(len(A[-1])-1, -1, -1):
#print(A[-1][c])
upper.append(A[-1][c])
A.pop(-1)
if len(A) == 0: break
# bottom row
for c in range(len(A[0])):
#print(A[0][c])
lower.append(A[0][c])
A.pop(0)
if len(A) == 0: break
#print(upper)
#print(lower)
#A
#
# Pass LEFT
#
# lower row
for r in range(len(A)-1, -1, -1):
#print(A[r][0])
upper.append(A[r][0])
A[r].pop(0)
if len(A[0]) == 0: break
# top row
for r in range(len(A)):
#print(A[r][-1])
lower.append(A[r][-1])
A[r].pop(-1)
if len(A[0]) == 0: break
sum_up = sum(upper)
if sum_up < -100000000 or sum_up > 100000000:
return -1
else:
return sum_up
# +
import random
import time # for testing
import numpy as np # for testing only
def test():
def create_test_arr(N, M, min_, max_):
A = np.random.randint(min_, max_+1, size=(N, M))
return [list(a) for a in A] # convert to list of lists
N = random.randint(1, 1000)
M = random.randint(1, min(100000 // N, 1000))
A1 = create_test_arr(N, M, -2147483648, 2147483647)
A2 = [a[:] for a in A1] # copy
ts = time.time()
sol1 = reference_solution(A1)
t1 = time.time() - ts
ts = time.time()
sol2 = solution(A2)
t2 = time.time() - ts
#print(f'{abs(t1-t2):.3f}')
assert sol1 == sol2
for i in range(100):
test()
# -
| BasicAlg/OnlineTest_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework for week 5
#
# 1. Scrape table from a single page (NFL)
# 2. Scrape table from a single page
# 3. CHALLENGE: Convert export to csv to a function
# ## 1. Scrape table from single page (NFL)
#
# ### We want to scrape a table that contains NFL player salaries by position for 2019.
#
# The webpage is ```https://sandeepmj.github.io/scrape-example-page/```
## import libraries
import pandas as pd ## to scrape tables
import requests ## to get data from websites
from bs4 import BeautifulSoup ## to process data scraped from websites
##scrape url website
url = "https://sandeepmj.github.io/scrape-example-page"
page = requests.get(url)
print(page.status_code) ## should print 200. checks http response code status
## turn into soup
soup = BeautifulSoup(page.content, "html.parser")
print(type(soup))
## MUST turn html into a string
html = str(soup)
print(type(html))
## use Pandas to read tables on page
tables = pd.read_html(html)
tables
## show type of object
type(tables)
## store it into a copy called nfl_df
nfl_df = tables[1]
nfl_df
# +
## use pandas to write to csv file
filename = "nfl_2019_salaries.csv"
nfl_df.to_csv(filename, encoding='utf-8', index=False)
print(f"{filename} is in your project folder!")
# -
# ## 2. Scrape table from a single page
#
#
# [Scrape this table](https://en.wikipedia.org/wiki/List_of_largest_companies_by_revenue) of largest global companies by revenue.
#
# Export the data into a csv file called ```big_revenue.csv```.
#
# **Note**: You might encounter the column headers appearing twice in your scraped table. Ignore that for now.
## import needed libraries
from bs4 import BeautifulSoup
import pandas as pd
import requests
# url to scrape
url = "https://en.wikipedia.org/wiki/List_of_largest_companies_by_revenue"
## get url and print but hard to read. will do prettify next
page = requests.get(url)
soup = BeautifulSoup(page.content, "html.parser")
print(soup)
## MUST turn html into a string
html = str(soup)
print(type(html))
## use Pandas to read tables on page
tables = pd.read_html(html)
tables
## target the table
## let's look at the first table:
tables[0]
top_companies = tables[0]
# ### Export and dealing with that extra header row
#
# Unlike ```pd.read_csv```, ```pd.to_csv``` has no ```skiprows=``` parameter.
#
# So the big picture is that we write out dataframe to a csv, then read it back in as a pandas dataframe where we skip the first row and then write to csv again.
#
# **Step 1 - Just export your dataframe as normal:**
## use pandas to write to csv file
filename = "big_revenue.csv"
top_companies.to_csv(filename, encoding='utf-8', index=False)
print(f"{filename} is in your project folder!")
# **Step 2 - Read the csv you just exported but skip the first row:**
#
# Confirm that the extra header is gone.
top_companies = pd.read_csv("big_revenue.csv", skiprows = 1)
top_companies.head()
# **Step 3 - Write your new dataframe to csv:**
top_companies.to_csv(filename, encoding='utf-8', index=False)
print(f"{filename} is in your project folder!")
# # Challenge
#
# ## 3. Convert the "write to csv file" code into a function
#
# Notice how you keep having to write the same code everytime you export a dataframe as csv.
# Convert it into a function so you just have to call the function with arguments like filename and which dataframe to convert.
def create_csv(df_name, file_name):
'''
Export your dataframe as a csv
argument 1 = your dataframe name
argument 2 = your file name as a string. Must include .csv
For example "my_data.csv"
'''
df_name.to_csv(file_name, encoding='utf-8', index=False )
print(f"{file_name} is in your local folder!")
## test it out on top_companies nfl
create_csv(top_companies, "tester.csv")
| week_06/homework-for-week-6_SOLUTION.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,.pct.py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Heteroskedastic Likelihood and Multi-Latent GP
# %% [markdown]
# ## Standard (Homoskedastic) Regression
# In standard GP regression, the GP latent function is used to learn the location parameter of a likelihood distribution (usually a Gaussian) as a function of the input $x$, whereas the scale parameter is considered constant. This is a homoskedastic model, which is unable to capture variations of the noise distribution with the input $x$.
#
#
# ## Heteroskedastic Regression
# This notebooks shows how to construct a model which uses multiple (here two) GP latent functions to learn both the location and the scale of the Gaussian likelihood distribution. It does so by connecting a **Multi-Output Kernel**, which generates multiple GP latent functions, to a **Heteroskedastic Likelihood**, which maps the latent GPs into a single likelihood.
#
# The generative model is described as:
#
# $$ f_1(x) \sim \mathcal{GP}(0, k_1(\cdot, \cdot)) $$
# $$ f_2(x) \sim \mathcal{GP}(0, k_2(\cdot, \cdot)) $$
# $$ \text{loc}(x) = f_1(x) $$
# $$ \text{scale}(x) = \text{transform}(f_2(x)) $$
# $$ y_i|f_1, f_2, x_i \sim \mathcal{N}(\text{loc}(x_i),\;\text{scale}(x_i)^2)$$
#
# The function $\text{transform}$ is used to map from the unconstrained GP $f_2$ to **positive-only values**, which is required as it represents the $\text{scale}$ of a Gaussian likelihood. In this notebook, the $\exp$ function will be used as the $\text{transform}$. Other positive transforms such as the $\text{softplus}$ function can also be used.
# %%
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_probability as tfp
import gpflow as gpf
# %% [markdown]
# ## Data Generation
# We generate heteroskedastic data by substituting the random latent functions $f_1$ and $f_2$ of the generative model by deterministic $\sin$ and $\cos$ functions. The input $X$ is built with $N=1001$ uniformly spaced values in the interval $[0, 4\pi]$. The outputs $Y$ are still sampled from a Gaussian likelihood.
#
# $$ x_i \in [0, 4\pi], \quad i = 1,\dots,N $$
# $$ f_1(x) = \sin(x) $$
# $$ f_2(x) = \cos(x) $$
# $$ \text{loc}(x) = f_1(x) $$
# $$ \text{scale}(x) = \exp(f_2(x)) $$
# $$ y_i|x_i \sim \mathcal{N}(\text{loc}(x_i),\;\text{scale}(x_i)^2)$$
# %%
N = 1001
np.random.seed(0)
tf.random.set_seed(0)
# Build inputs X
X = np.linspace(0, 4 * np.pi, N)[:, None] # X must be of shape [N, 1]
# Deterministic functions in place of latent ones
f1 = np.sin
f2 = np.cos
# Use transform = exp to ensure positive-only scale values
transform = np.exp
# Compute loc and scale as functions of input X
loc = f1(X)
scale = transform(f2(X))
# Sample outputs Y from Gaussian Likelihood
Y = np.random.normal(loc, scale)
# %% [markdown]
# ### Plot Data
# Note how the distribution density (shaded area) and the outputs $Y$ both change depending on the input $X$.
# %%
def plot_distribution(X, Y, loc, scale):
plt.figure(figsize=(15, 5))
x = X.squeeze()
for k in (1, 2):
lb = (loc - k * scale).squeeze()
ub = (loc + k * scale).squeeze()
plt.fill_between(x, lb, ub, color="silver", alpha=1 - 0.05 * k ** 3)
plt.plot(x, lb, color="silver")
plt.plot(x, ub, color="silver")
plt.plot(X, loc, color="black")
plt.scatter(X, Y, color="gray", alpha=0.8)
plt.show()
plt.close()
plot_distribution(X, Y, loc, scale)
# %% [markdown]
# ## Build Model
# %% [markdown]
# ### Likelihood
# This implements the following part of the generative model:
# $$ \text{loc}(x) = f_1(x) $$
# $$ \text{scale}(x) = \text{transform}(f_2(x)) $$
# $$ y_i|f_1, f_2, x_i \sim \mathcal{N}(\text{loc}(x_i),\;\text{scale}(x_i)^2)$$
# %%
likelihood = gpf.likelihoods.HeteroskedasticTFPConditional(
distribution_class=tfp.distributions.Normal, # Gaussian Likelihood
scale_transform=tfp.bijectors.Exp(), # Exponential Transform
)
print(f"Likelihood's expected latent_dim: {likelihood.latent_dim}")
# %% [markdown]
# ### Kernel
# This implements the following part of the generative model:
# $$ f_1(x) \sim \mathcal{GP}(0, k_1(\cdot, \cdot)) $$
# $$ f_2(x) \sim \mathcal{GP}(0, k_2(\cdot, \cdot)) $$
#
# with both kernels being modeled as separate and independent $\text{SquaredExponential}$ kernels.
# %%
kernel = gpf.kernels.SeparateIndependent(
[
gpf.kernels.SquaredExponential(), # This is k1, the kernel of f1
gpf.kernels.SquaredExponential(), # this is k2, the kernel of f2
]
)
# The number of kernels contained in gpf.kernels.SeparateIndependent must be the same as likelihood.latent_dim
# %% [markdown]
# ### Inducing Points
# Since we will use the **SVGP** model to perform inference, we need to implement the inducing variables $U_1$ and $U_2$, both with size $M=20$, which are used to approximate $f_1$ and $f_2$ respectively, and initialize the inducing points positions $Z_1$ and $Z_2$. This gives a total of $2M=40$ inducing variables and inducing points.
#
# The inducing variables and their corresponding inputs will be Separate and Independent, but both $Z_1$ and $Z_2$ will be initialized as $Z$, which are placed as $M=20$ equally spaced points in $[\min(X), \max(X)]$.
#
# %%
M = 20 # Number of inducing variables for each f_i
# Initial inducing points position Z
Z = np.linspace(X.min(), X.max(), M)[:, None] # Z must be of shape [M, 1]
inducing_variable = gpf.inducing_variables.SeparateIndependentInducingVariables(
[
gpf.inducing_variables.InducingPoints(Z), # This is U1 = f1(Z1)
gpf.inducing_variables.InducingPoints(Z), # This is U2 = f2(Z2)
]
)
# %% [markdown]
# ### SVGP Model
# Build the **SVGP** model by composing the **Kernel**, the **Likelihood** and the **Inducing Variables**.
#
# Note that the model needs to be instructed about the number of latent GPs by passing `num_latent_gps=likelihood.latent_dim`.
# %%
model = gpf.models.SVGP(
kernel=kernel,
likelihood=likelihood,
inducing_variable=inducing_variable,
num_latent_gps=likelihood.latent_dim,
)
model
# %% [markdown]
# ## Model Optimization
#
# ### Build Optimizers (NatGrad + Adam)
# %%
data = (X, Y)
loss_fn = model.training_loss_closure(data)
gpf.utilities.set_trainable(model.q_mu, False)
gpf.utilities.set_trainable(model.q_sqrt, False)
variational_vars = [(model.q_mu, model.q_sqrt)]
natgrad_opt = gpf.optimizers.NaturalGradient(gamma=0.1)
adam_vars = model.trainable_variables
adam_opt = tf.optimizers.Adam(0.01)
@tf.function
def optimisation_step():
natgrad_opt.minimize(loss_fn, variational_vars)
adam_opt.minimize(loss_fn, adam_vars)
# %% [markdown]
# ### Run Optimization Loop
# %%
epochs = 100
log_freq = 20
for epoch in range(1, epochs + 1):
optimisation_step()
# For every 'log_freq' epochs, print the epoch and plot the predictions against the data
if epoch % log_freq == 0 and epoch > 0:
print(f"Epoch {epoch} - Loss: {loss_fn().numpy() : .4f}")
Ymean, Yvar = model.predict_y(X)
Ymean = Ymean.numpy().squeeze()
Ystd = tf.sqrt(Yvar).numpy().squeeze()
plot_distribution(X, Y, Ymean, Ystd)
model
# %% [markdown]
# ## Further reading
#
# See [Chained Gaussian Processes](http://proceedings.mlr.press/v51/saul16.html) by Saul et al. (AISTATS 2016).
| doc/source/notebooks/advanced/heteroskedastic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import rclpy
from rclpy.node import Node
from geometry_msgs.msg import Twist
rclpy.init(args=None)
node = Node('talker')
pub = node.create_publisher(Twist, 'cmd_vel', 10)
msg = Twist()
# -
msg.linear.x = -1.0
msg.angular.z = 0.0
pub.publish(msg)
msg.linear.x = 0.0
msg.angular.z = 0.0
pub.publish(msg)
msg.linear.x = 0.0
msg.angular.z = -1.0
pub.publish(msg)
node.destroy_node()
rclpy.shutdown()
| notebooks/ros2_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import turtle
# # Draw simple Square motion
# +
# motion
turtle.forward(100)
turtle.right(90)
turtle.forward(100)
turtle.right(90)
turtle.forward(100)
# clear the drawing and remain turtle
turtle.clear()
# -
# # Drawing Circles with Directionality
# +
# make a turtle object
# and do some drawing
t1 = turtle.Turtle()
t1.up()
t1.setpos(-100, 50)
t1.down()
t1.circle(50)
# make a turtle object
# and do some drawing
t2 = turtle.Turtle()
t2.up()
t2.setpos(50, 50)
t2.down()
t2.circle(50)
# make a turtle object
# and do some drawing
t3 = turtle.Turtle()
t3.up()
t3.setpos(50, -100)
t3.down()
t3.circle(50)
# make a turtle object
# and do some drawing
t4 = turtle.Turtle()
t4.up()
t4.setpos(-100, -100)
t4.down()
t4.circle(50)
# here we clear the work done by turtle
# objects : t1 and t3 only but turtle
# shape remain as it is
t1.clear()
t3.clear()
# -
| Python/7. Applications Fun/animation_fun/simple_turtle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 9: Supervised Machine Learning
# *Use a set of training data to make predictions about new data.*
# ## Instructions
# This tutorial provides step-by-step training divided into numbered sections. The sections often contain embeded exectable code for demonstration. This tutorial is accompanied by a practice notebook: [L09-Supervised_Machine_Learning-Practice.ipynb](./L09-Supervised_Machine_Learning-Practice.ipynb).
#
# Throughout this tutorial sections labeled as "Tasks" are interspersed and indicated with the icon: . You should follow the instructions provided in these sections by performing them in the practice notebook. When the tutorial is completed you can turn in the final practice notebook.
# ## Introduction
#
# For this notebook we will learn about supervised machine learning using the **scikit-learn** (**sklearn**) package. skLearn is a Python library that provides machine clearning capabilities. It is built on Numpy, SciPy and matplotlib.
#
# For machine learning there are three primary purposes:
# - Classification: to predict the outcome "class" to which a sample belongs.
# - Regression: predicting an outcome value on a continuous scale.
# - Clustering: automatic grouping of similar objects into outcome classes when those classes are unknown.
#
# You can read more at these links:
#
# - Classification: https://en.wikipedia.org/wiki/Statistical_classification
# - Prediction: https://en.wikipedia.org/wiki/Predictive_analytics
# - Clustering: https://en.wikipedia.org/wiki/Cluster_analysis
#
# For supervised machine learning, a training set of data is provided to a set of appropriate algorithms (i.e. for classification, regression or clustering). The training data set typically consists of a subset of all available data that includes both independent and dependent variables (i.e. **outcome**) measured across a variety of samples. This training data is provided to one or more algorithms which determine a **model** that can be used to classify or predict outcomes. Typically, a set of data is set aside to test, or validate, the accuracy of the model.
#
# For this notebook we will once again use the Iris dataset and we will use supervised machine learning to create a model for predicting species. Therefore, the outcome variable is `species` and all others (`sepal_width`, `sepal_length`, `petal_width`, `petal_length`) are the independent variables. Therefore, this notebook will demonstrate a "Classification" example.
# ---
# ## 1. Getting Started
# As before, we import any needed packages at the top of our notebook. Let's import Numpy, Pandas, Seaborn, matplotlib and the sklearn machine learning libraries.
# +
# %matplotlib inline
# Data Management
import numpy as np
import pandas as pd
# Visualization
import seaborn as sns
import matplotlib.pyplot as plt
# Machine learning
from sklearn import model_selection
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
# -
# #### Task 1a: Setup
#
# <span style="float:right; margin-left:10px; clear:both;">
# </span>
#
# Import the following package sets:
# + packages for data management
# + pacakges for visualization
# + packages for machine learning
#
# Remember to activate the `%matplotlib inline` magic.
# ---
# ## 2. Data Exploration
# Import the iris dataset
iris = sns.load_dataset('iris')
# ### 2.1 Summarize the dataset
# Just as we learned in the Data Wrangling notebook, we should always summarize the data. Execute the following commands to explore this data.
iris.shape
iris.head(10)
iris.dtypes
iris.describe()
# How many samples do we have for our outcome variable?
iris.groupby('species').size()
# ### 2.2 Check for missing or duplicated data
# Just like we did in the Data Wrangling notebook, we want to check for missing values and duplication
iris.isna().sum()
iris.duplicated().sum()
iris[iris.duplicated(keep= False)]
iris.nunique()
# ### 2.3 Examine data distributions and check for outliers
# Some statistical methods make assumptions about the distribution of samples for each variable and the presence of outliers, so, we need to be aware of how our data appears and any potential outliers. Pandas dataframes have a very convenient `hist` function for printing histograms of all numeric columns. It is based on Matplotlib.
iris.hist()
plt.show()
# Conveniently, Pandas dataframes also have a flexible `plot` function as well, that allows us to print other types of pltos, such as a boxplot. You can learn more about the plot function by viewing the [online documentation](https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.plot.html)
iris.plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False)
plt.show()
# Let's see if there are outliers without our outcome groups. It turns out we can easily do this with a `grouby`. The DataFrameGroupBy object, also has a `boxplot` function that makes this easy to view as well!
iris.groupby(by='species').boxplot(rot=90);
# It does indeed appear that we have outliers.
# ### 2.4 Search for Collinearity
# It is important to identify collinearity prior to creating any machine learning model. There are two types of collinearity:
# - Structurual: this occurs when we create new columns in our data that are derived from other columns. For example, a log transformation of one column.
# - Multicollinearity: this can be natural in our data and occurs when one variable can be used to linearly predict another.
#
# Why can collinearity be bad? For regression models, collinearity can weaken the p-values because it makes the estimated co-efficients too sensitive to changes in the model. For an example of the affect that collinarity can have with a regression model sse the [JMP Multicollinearity](https://www.jmp.com/en_us/statistics-knowledge-portal/what-is-multiple-regression/multicollinearity.html) page.
#
# We can empirically check for collinarity using a simple pairwise scatterplot using Seaborn.
sns.pairplot(iris, hue="species")
# You should consider removing columns that are severly collinear. The more collinear, the more significant the impact on the model.
# #### Task 2a: Data Exploration
#
# <span style="float:right; margin-left:10px; clear:both;">
# </span>
#
# After reviewing the data in sections 2.1, 2.2, 2.3 and 2.4 do you see any problems with this iris dataset? If so, please describe them in the practice notebook. If not, simply indicate that there are no issues.
# #### Task 2b: Make Assumptions
# <span style="float:right; margin-left:10px; clear:both;">
# </span>
#
# After reviewing the data in sections 2.1, 2.2, 2.3 and 2.4 are there any columns that would make poor predictors of species?
#
# **Hint**: columns that are poor predictors are:
# + those with too many missing values
# + those with no difference in variation when grouped by the outcome class
# + variables with high levels of collinearity
# ## 3. Prepare the Data
# ### 3.1 Prepare the Data
# The sklearn package expects that all indpendent variables are numerical and that independent and dependent variables are separated into different data objects. It also expects that data is in Numpy arrays (not Pandas data frames). First, let's separate the variables to only include those that are numeric. Then, we'll create a 2D Numpy array, named `X`, containing the indpendent variables.
# +
X = iris.loc[:,'sepal_length':'petal_width'].values
# Show the contents of X by displaying the first 10 rows.
X[0:10]
# -
# Next we will create a new Numpy 1D array named `Y` that will house the dependent variable (species name).
# +
Y = iris['species'].values
# Show the contents of X by displaying the first 10 elements.
Y[0:10]
# -
# Observe that we have no independent variables that are categorical. All are numeric. If we did have categorical data we must convert those to numeric values. sklearn requires that all data in `X` be numeric. We have several options as defined in the [Preprocessing data](https://scikit-learn.org/stable/modules/preprocessing.html#encoding-categorical-features) section of sklearn:
# 1. Use the sklearn `processing.OrdinalEncoder` function
# - It converts categorical data to ordered numbers.
# - Use this only if the classes are also ordinal.
# 2. Use the `processing.OneHotEncoder` function.
# - It converts all but one class for a single variable to 0 and leaves one category of interest as 1.
# - Use this only if you are predicting only a single class of outcome.
# 3. Pivot the categorical column into multiple new binary columns.
# - The values in the columns are 0 and 1 and indicate if the category applies to the sample row.
# - Use this if there are multiple classes for a single variable and they are not ordinal. Unfortunately, this is **not** tidy. But it is required to handle multiple classes.
#
# As the iris data is all numeric we have no need for any of these options.
# ### 3.2 Normalize the data
# Many machine learning algorithms expect that the quantitative columns are centered at 0 and scaled to unit variance. See the [preprocessing documentation](https://scikit-learn.org/stable/modules/preprocessing.html#preprocessing-scaler) for sklearn.
#
# According to the sklearn documentation:
#
# > In practice we often ignore the shape of the distribution and just transform the data to center it by removing the mean value of each feature, then scale it by dividing non-constant features by their standard deviation... The function `scale` provides a quick and easy way to perform this operation on a single array-like dataset.
#
# > If your data contains many outliers, scaling using the mean and variance of the data is likely to not work very well. In these cases, you can use `robust_scale`... [it uses] more robust estimates for the center and range of your data.
#
#
# We can, therefore, normalize the `X` Numpy array using the [preprocessing.scale](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.scale.html#sklearn.preprocessing.scale) or [preprocessing.robust_scale](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.robust_scale.html#sklearn.preprocessing.robust_scale) function of Sklearn. Choose the method most appropriate given the state of outliers in the data. As it is clear we have outliers in some of the columns of the iris data we should use the`robust_scale` method.
# +
X = preprocessing.robust_scale(X)
# Show the contents of X by displaying the first 10 rows.
# The values should be scaled between -1 to 1
X[0:10]
# -
# ### 3.3 Split the data for testing and validation
# For supervised machine learning, you must provide a testing dataset to create a model. The model is then used on another dataset to validate its accuracy. If the dataset is large enough we split the original dataset into a test and a validation set. We can do this using the sklearn `model_selection.train_test_split` function. The function takes as input the 2D Numpy array of independent variables and the 1D Numpy array with the oucome variable. It also takes the following arguments:
#
# - `test_size`: "should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split." For example, to use 80% of the data for model construction and 20% for testing (i.e. validation) this argument should be set to 0.2. The default is 0.25.
# - `random_state`: rows are randomly selected to include in the split. You can ensure the same random order by providing a seed. This allows for reproducibility.
#
# Lets split the data with 20% used for testing and a random seed of 10:
# +
# Split-out validation dataset
Xt, Xv, Yt, Yv = model_selection.train_test_split(X, Y, test_size=0.2, random_state=10)
# Print the shapes of each dataset
print("The sizes of the training independent and dependent datasets")
print(Xt.size)
print(Yt.size)
print("The sizes of the validation independent and dependent datasets")
print(Xv.size)
print(Yv.size)
# -
# Using the code above `Xt` and `Yt` become the "training" data. and `Xv` and `Yv` become the "testing" data used for validation.
# ## 3. Perform Supervised Machine Learning
#
# ### 3.1 K-Fold Strategy
# Now that we have our data separated into training and validation sets, we should establish a training strategy. To avoid overfitting a model to the data, we should perform a K-fold cross-validation strategy. This type of strategy will further divide our training data into <i>k</i> subsets. The model will be trained <i>k</i> times using <i>k</i>-1 subsets. The <i>k</i>th subset will be set aside for validation. But, for each of the <i>k</i> tests, a different subset will have its turn for validation. An accuracy score is provided for each attempt. You can evaulate the performance of a machine learning algorithm by exploring the distribution (mean and variance) of the <i>k</i> attempts (we will try this later in the notebook).
#
# 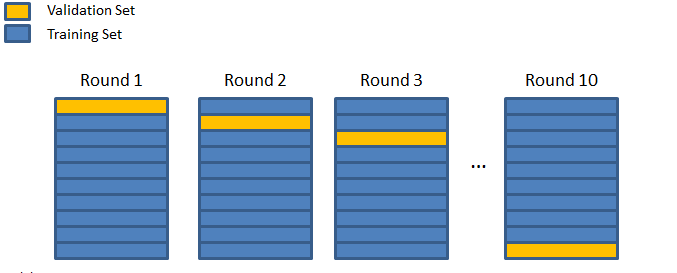
#
# <sup>Image from [TowardsDataScience.com Cross Validation](https://towardsdatascience.com/cross-validation-explained-evaluating-estimator-performance-e51e5430ff85) page.
#
# To establish a K-fold cross-validation strategy we use the `model_selection.KFold` function of sklearn. It takes two important arguments:
#
# - `n_splits`: the number of subsets to split the data.
# - `random_state`: you can ensure the same random order in the subsets by providing a seed. This allows for reproducibility.
#
# Lets create a K-fold strategy that splits the iris data into 10 subsets and a random seed of 10.
kfold = model_selection.KFold(n_splits=10, random_state=10)
# ### 3.2. Evaulate ML Algorithms
# sklearn provides a variety of supervised machine learning aglorithms. Here, we will evaluate six of them:
#
# 1. Logistic Regression (LR)
# 2. Linear Discriminant Analysis (LDA)
# 3. K-Nearest Neighbors (KNN).
# 4. Classification and Regression Trees (CART).
# 5. Gaussian Naive Bayes (NB).
# 6. Support Vector Machines (SVM).
#
# Each of these algorithms is provided by sklearn as an object. Notice, these were imported in Section 1 above:
#
# ```python
# from sklearn.linear_model import LogisticRegression
# from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
# from sklearn.neighbors import KNeighborsClassifier
# from sklearn.tree import DecisionTreeClassifier
# from sklearn.naive_bayes import GaussianNB
# from sklearn.svm import SVC, LinearSVC
# ```
#
# To build a model, we must first create an object for the algorithm. For example, to create an object for Logistic Regression we could use the following code:
# ```python
# alg = LogisticRegression(solver='lbfgs', multi_class="auto")
# ```
#
#
# In the following sections, we will construct an object for each algorithm and use it to create a predictive model, but first, we need a way to store the results of all ML algorithms that we'll be using. To make this easy, we will store results from each method in a dictionary. The following code prepares a dictionary containing, as keys, the names of the 6 methods we will explore. The value is an array of 10 zeros. This is because we will be performing a 10-fold cross-validation and we replace the zeros with results of each of the 10 attempts per method.
results = {
'LogisticRegression' : np.zeros(10),
'LinearDiscriminantAnalysis' : np.zeros(10),
'KNeighborsClassifier' : np.zeros(10),
'DecisionTreeClassifier' : np.zeros(10),
'GaussianNB' : np.zeros(10),
'SVC' : np.zeros(10)
}
results
# After creating the algorithm object (in this example a LogisiticRegression object) we perform the model building and cross-validation in one step using the `model_selection.cross_val_score` funtion:
#
# ```python
# model_selection.cross_val_score(alg, Xt, Yt, cv=kfold, scoring="accuracy", error_score=np.nan)
# ```
# Here we provide the algorithm object, the training data, validation data, k-fold object, the `scoring` accuracy argument and an `error_score` argument. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html) for details about other arguments. We can use the same `cross_val_score` function for any of the 6 algorithms.
#
# In the following sections, six supervised machine learning methods will be briefly introduced. For brevity we will not explore these approaches too deeply. A brief summary is provided followed by links for additional details.
# #### 3.2.1 Logistic Regression
#
# **Brief Summary**
#
# In statistics, the logistic model (or logit model) uses a logistic function to model a binary outcome (i.e. values of 0 and 1) and a set of one or more independent variables: [X<sub>1</sub>, X<sub>2</sub>, X<sub>3</sub>, ... X<sub>n</sub>]:
#
# \begin{equation*}
# logit(p) = b_0 + b_1X_1 + b_2X_2 + b_2X_2 ... b_nX_n
# \end{equation*}
#
# Its goal is to determine the set of coefficients, [b<sub>1</sub>, b<sub>2</sub>, b<sub>3</sub>, ... b<sub>n</sub>], that best fit the relationship between the dependent and independent variables. Alternatively, it is the natural log of the odds ratio:
#
# \begin{equation*}
# odds\ ratio = \frac{p}{1-p} = \frac{probability\ of\ presence}{probability\ of\ absense}
# \end{equation*}
#
# \begin{equation*}
# logit(p) = ln(\frac{p}{1-p})
# \end{equation*}
#
#
# The following image from Wikipedia shows an example where the dependent variable of *"passing an exam"* with a single independent variable of *"hours studied"*.
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/6/6d/Exam_pass_logistic_curve.jpeg" style="height:300px">
#
# Samples can be classified as predicting a passing score or a non-passing score by where they fall (above or below) on the line. As a machine learning approach, we can provide a set of training data to create the model, then use future data to predict an outcome.
#
# **When to use:**
# + Input data is quantitative.
# + No colliniearity.
# + No outliers.
# + Relationship is expected to be linear
#
# **Additional Resources**
# + [skLearn Logisitic Regression Function](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html)
# + [Wikipedia LR](https://en.wikipedia.org/wiki/Logistic_regression)
# + [TowardsDataScience.com LR](https://towardsdatascience.com/building-a-logistic-regression-in-python-step-by-step-becd4d56c9c8)
#
# **Practice**
#
# To perform Logisitc Regression with sklearn, we must first create a LogisiticRegression object. There are two important arguments to provide:
#
# - `solver`: There are several different solvers: ‘newton-cg’, ‘lbfgs’, ‘liblinear’, ‘sag’, ‘saga’. These are meant for different types of data:
# + For small datasets, ‘liblinear’ is a good choice, whereas ‘sag’ and ‘saga’ are faster for large ones.
# + For multiclass problems, only ‘newton-cg’, ‘sag’, ‘saga’ and ‘lbfgs’ handle multinomial loss; ‘liblinear’ is limited to one-versus-rest schemes.
# + ‘newton-cg’, ‘lbfgs’ and ‘sag’ only handle L2 penalty, whereas ‘liblinear’ and ‘saga’ handle L1 penalty.
# - `multi_class`: set this argument when there are more than two classes in the outcome variable. Setting it to `auto` allows the algorithm to select the best approach for working with the data.
#
# Because the iris species is multinomial (multiple categories), we will set the `multi_class` argument to `auto` and the `solver` to `libfgs`
# +
# Create the LogisticRegression object prepared for a multinomial outcome validation set.
alg = LogisticRegression(solver='lbfgs', multi_class="auto")
# Execute the cross-validation strategy
results['LogisticRegression'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['LogisticRegression']
# -
# #### 3.2.2 Linear Discriminant Analysis (LDA)
#
# **Brief Summary**
#
# Linear Discriminant Analysis (LDA) is a classification method that employs a dimensionality reduction technique.
#
# From the [Wikipedia LDA](https://en.wikipedia.org/wiki/Linear_discriminant_analysis) page:
#
# > Linear discriminant analysis (LDA),... [is] a method used in statistics, pattern recognition and machine learning to find a linear combination of features that characterizes or separates two or more classes of objects or events... LDA is closely related to analysis of variance (ANOVA) and regression analysis, which also attempt to express one dependent variable as a linear combination of other features or measurements. However, ANOVA uses categorical independent variables and a continuous dependent variable, whereas discriminant analysis has continuous independent variables and a categorical dependent variable (i.e. the class label)
#
# Assumptions:
# + Outcome classes are normally distributed.
# + Variance between outcome classes is equal.
#
# LDA is similar to Principal Component Analysis (PCA), but….
# + PCA is unsupervised, LDA is supervised
# + PCA tries to maximize the variance.
# + LDA tries to maximize the separation between data classes.
#
# <img src="http://sebastianraschka.com/images/blog/2014/linear-discriminant-analysis/lda_1.png" style="height:250px">
# <sup><i>Image from <a href="http://sebastianraschka.com/Articles/2014_python_lda.html">SebastianRaschka Liniear Discriminant Analysis</a> page</i></sup>
#
# **When to use:**
# + The dependent variable is categorical (qualitative) with "classes" or categories as outcomes
# + The independnet variables are quantitative.
#
# **Additional Resources**
# + [skLearn LDA](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.LinearDiscriminantAnalysis.html)
# + [Wikipedia LDA](https://en.wikipedia.org/wiki/Linear_discriminant_analysis)
# + [TowardsDataSciecne LDA](https://towardsdatascience.com/classification-part-2-linear-discriminant-analysis-ea60c45b9ee5)
#
# **Practice**
#
# Similar to Logistic Regression, we must first create a LinearDiscriminantAnalysis object, then call `cross_val_score`. Here we'll just use the default settings. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.LinearDiscriminantAnalysis.html) for a more thorough description of arguments and flexibility of the algorithm.
# +
# Create the LinearDiscriminantAnalysis object with defaults.
alg = LinearDiscriminantAnalysis()
# Execute the cross-validation strategy
results['LinearDiscriminantAnalysis'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['LinearDiscriminantAnalysis']
# -
# #### 3.2.3 K-Nearest Neighbors (KNN)
#
# **Brief Summary**
#
# From the [Wikipedia KNN](https://en.wikipedia.org/wiki/K-nearest_neighbors_algorithm) page:
#
# > In pattern recognition, the k-nearest neighbors algorithm (k-NN) is a non-parametric method used for classification and regression...
# >
# > <img src="https://upload.wikimedia.org/wikipedia/commons/e/e7/KnnClassification.svg" style="height: 200px">
# >
# > [Above is an] example of k-NN classification. The test sample (green circle) should be classified either to the first class of blue squares or to the second class of red triangles. If k = 3 (solid line circle) it is assigned to the second class because there are 2 triangles and only 1 square inside the inner circle. If k = 5 (dashed line circle) it is assigned to the first class (3 squares vs. 2 triangles inside the outer circle).
#
# For the KNN approach to be useful, the outcome classes must be distinguishable. As an exapmle of distinguishable classes, consider the Iris dataset petal width vs petal length:
sns.relplot(x="petal_width", y="petal_length", hue="species", data=iris);
# **When to Use**
# + Multiple outcomes (not just binary)
# + There are no assumptions for the outcome distribution
# + The outcomes classes are already distinguishable in the problem space.
#
# **Additional Resources**
# + [skLearn KNN](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html)
# + [Wikipedia KNN](https://en.wikipedia.org/wiki/K-nearest_neighbors_algorithm)
# + [TowardsDataScience.com KNN](https://towardsdatascience.com/knn-k-nearest-neighbors-1-a4707b24bd1d)
#
# **Practice**
#
# Similar to other algorithms, we must first create a KNeighborsClassifier object, then call `cross_val_score`. Here we'll just use the default settings. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html) for a more thorough description of arguments and flexibility of the algorithm.
# +
# Create the KNeighborsClassifier object with defaults.
alg = KNeighborsClassifier()
# Execute the cross-validation strategy
results['KNeighborsClassifier'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['KNeighborsClassifier']
# -
# #### 3.2.4 Classification and Regression Trees (Decision Trees)
#
# **Brief Summary**
#
# From the [Wikipedia Decision Tree Learning](https://en.wikipedia.org/wiki/Decision_tree_learning) page:
#
# > In computer science, Decision tree learning uses a decision tree (as a predictive model) to go from observations about an item (represented in the branches) to conclusions about the item's target value (represented in the leaves). It is one of the predictive modeling approaches used in statistics, data mining and machine learning. Tree models where the target variable can take a discrete set of values are called classification trees; in these tree structures, leaves represent class labels and branches represent conjunctions of features that lead to those class labels. Decision trees where the target variable can take continuous values (typically real numbers) are called regression trees.
# >
# >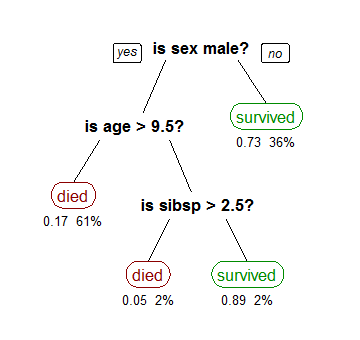
# >
# > [The above] tree [shows] survival of passengers on the Titanic. The figures under the leaves show the probability of survival and the percentage of observations in the leaf.
#
# In short, trees are learned by splitting the source data based on an attribute value and two types of trees are possible based on the outcome data:
#
# Classification Trees
# - When outcome is categorical
#
# Regression Trees
# - When outcome is continuous
#
# **When to Use**
# - For both numerical and categorical outcome data
# - Distribution of outcome is unknown or doesn’t meet other model assumptions.
# - Multiple outcomes.
#
# **Additional Resources**
# - [sklearn Decision Trees](https://scikit-learn.org/stable/modules/tree.html)
# - [sklearn CART](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html)
# - [Wikipediea Decision Trees](https://en.wikipedia.org/wiki/Decision_tree)
# - [TowardsDataScience.com CART](https://towardsdatascience.com/decision-trees-d07e0f420175)
#
#
# **Practice**
#
# Similar to other algorithms, we must first create a DecisionTreeClassifier object, then call `cross_val_score`. Here we'll just use the default settings. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html) for a more thorough description of arguments and flexibility of the algorithm.
# +
# Create the DecisionTreeClassifier object with defaults.
alg = DecisionTreeClassifier()
# Execute the cross-validation strategy
results['DecisionTreeClassifier'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['DecisionTreeClassifier']
# -
# #### 3.2.5 Gaussian Naive Bayes (NB)
#
# **Brief Summary**
#
# From the [Wikipedia Naive Bayes classifier](https://en.wikipedia.org/wiki/Naive_Bayes_classifier) page:
#
# > In machine learning, naive Bayes classifiers are a family of simple "probabilistic classifiers" based on applying Bayes' theorem with strong (naive) independence assumptions between the features.
#
# Naive Bayes methods rely on Bayes theorem, and follows axioms of the conditional probability:
#
# \begin{equation*}
# P(A | B) = \frac{P(B|A)P(A)}{P(B)}
# \end{equation*}
#
# .
# Where:
# - A = outcome class
# - B = input data vector for a sample.
#
# There are many NB algorithms and they differ in assumptions about the distribution of P(A|B). For the Guassian approach the distributions are expected to be Guassian and the input data is quantitative.
#
# **When to Use**
#
# - High independence between input data (no multicollinearity)
# - Multiple outcome classes.
# - Independent variables are quantitative.
#
# **Additional Resources**
# - [skLearn Guassian NB](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html)
# - [Wikipedia NB](https://en.wikipedia.org/wiki/Naive_Bayes_classifier)
# - [TowardsDataScience NB](https://towardsdatascience.com/all-about-naive-bayes-8e13cef044cf)
#
# **Practice**
#
# Similar to other algorithms, we must first create a GaussianNB object, then call `cross_val_score`. Here we'll just use the default settings. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html) for a more thorough description of arguments and flexibility of the algorithm.
# +
# Create the GaussianNB object with defaults.
alg = GaussianNB()
# Execute the cross-validation strategy
results['GaussianNB'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['GaussianNB']
# -
# #### 3.2.6 Support Vector Machines (SVM)
# **Brief Summary**
#
# A support vector machine attempts to find an optimal separation between outcomes by separating them in multi-dimensional space (ℝ<sup>n</sup>). Consider the following figure:
#
# <img src="https://cdn-images-1.medium.com/max/1000/1*ZpkLQf2FNfzfH4HXeMw4MQ.png" style="height:250px">
# <sup><i>Image from <a href="https://towardsdatascience.com/support-vector-machine-introduction-to-machine-learning-algorithms-934a444fca47">TowardsDataScience.com SVM</a> page</i></sup>
#
# Here the line in the first 2-dimensional plot separates the two classes and in the 3-dimensional plot, the plane separates them. For larger dimensions, SVMs find the optimal **hyperplane**. The optimal "hyperplane" (line or plane) is selected as the one which maximizes the **margin** between the outcome classes. Consier the following figure:
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/b/b5/Svm_separating_hyperplanes_%28SVG%29.svg" style="height:250px">
# <sup><i>Image from <a href="https://en.wikipedia.org/wiki/Support-vector_machine">Wikipedia SVM</a> page</i></sup>
#
# In the example figure above:
# - The H1 line does not separate the classes.
# - The H2 line does, but only with a small margin (distance between samples in the classes)
# - The H3 line separates them with the maximal margin.
#
# There are multiple types of SVMs. The SVM that performs classification is known as a Support Vector Classification (SVC) algorithm.
#
# **When to use**
# - Multiple outcome classes.
# - Distribution of outcome is unknown or doesn’t meet other model assumptions.
# - High dimensional data
# - There are more samples than features (otherwise overfitting may occur)
#
# **Additional Resources**
# - [skLearn SVM](https://scikit-learn.org/stable/modules/svm.html)
# - [skLearn SVC](https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html#sklearn.svm.SVC)
# - [Wikipedia SVM](https://en.wikipedia.org/wiki/Support-vector_machine)
# - [TowardsDataScience.com SVM](https://towardsdatascience.com/support-vector-machines-a-brief-overview-37e018ae310f)
#
#
# **Practice**
#
# Similar to other algorithms, we must first create a SVC object, then call `cross_val_score`. Here we'll just use the default settings. See the [online documentation](https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html#sklearn.svm.SVC) for a more thorough description of arguments and flexibility of the algorithm.
# +
# Create the SVC object with defaults.
alg = SVC(gamma='auto')
# Execute the cross-validation strategy
results['SVC'] = model_selection.cross_val_score(alg, Xt, Yt, cv=kfold,
scoring="accuracy", error_score=np.nan)
# Take a look at the scores for each of the 10-fold runs.
results['SVC']
# -
# ### 3.3 Compare Scores
# Now that we have performed training and validation of six different methods we can compare the results to see which performed best. We can do so by converting the results dictionary into a dataframe and using the `plot` function that comes with Panda's data frames to create a box plot of each test.
pd.DataFrame(results).plot(kind="box", rot=45);
# ### 3.4 Test the model (make predictions)
# From the boxplot in the previous section, it seems as if Linear Discriminant Analysis performed best! We will use this algorithm and the trained model to compare predictions made from the 20% of data we set aside for testing match the actual outcomes. To do this we will use two new functions that are members of any of the algorithm objects:
#
# - `fit`: This function employs the algorithm to create the final predictive model. All the training data is provided to this function.
# - `predict`: This function uses the trained model to make predictions using the testing data.
# +
# Create the LinearDiscriminantAnalysis object with defaults.
alg = LinearDiscriminantAnalysis()
# Create a new model using all of the training data.
alg.fit(Xt, Yt)
# Using the testing data, predict the iris species.
predictions = alg.predict(Xv)
# Let's see the predictions
predictions
# -
# Finally, we can evaulate how accurate the model has been by comparing the predictions with the actual species for the test data. The `accuracy_score` function provides this:
accuracy_score(Yv, predictions)
# One way to explore the performance of the algorithm is by way of a **Confusion Matrix** or error matrix. A confusion matrix is an <i>n</i> x <i>n</i> matrix where n is the number of outcome classes. In the case of the Iris data, <i>i</i> = 3. To view the confusion matrix use the sklearn `confusion_matrix` function. We pass in the testing outcome set, the predictions and the order that we want the classes to appear (using the `labels` argument):
labels = ['versicolor', 'virginica', 'setosa']
cm = confusion_matrix(Yv, predictions, labels=labels)
print(cm)
# The elements of the confusion matrix have the following meaning:
# - rows of the confusion matrix represent the predicted classes
# - columns represent the actual classes.
# - elements on the diagnoal of the matrix represent the true positives
# - errors are present when counts above zero are outside of the diagnoal.
#
# ***Note***: Because the result is a Numpy array, there are no row and column labels. However, because we provided the `labels` argument we know the order of the classes in the matrix.
#
# We can use the Seaborn `heatmap` to visualize the matrix:
sns.heatmap(cm, annot=True, xticklabels=labels, yticklabels=labels);
# Finally, the `classification_report` function indicates how well the model was at predicting each class.
cr = classification_report(Yv, predictions)
print(cr)
# Where:
# - `precision`: The precision is the ratio `tp / (tp + fp)` where `tp` is the number of true positives and `fp` the number of false positives. The precision is intuitively the ability of the classifier not to label as positive a sample that is negative.
# - `recall`: The recall is the ratio `tp / (tp + fn)` where `tp` is the number of true positives and `fn` the number of false negatives. The recall is intuitively the ability of the classifier to find all the positive samples.
# - `f1-score`: The F-beta score can be interpreted as a weighted harmonic mean of the precision and recall, where an F-beta score reaches its best value at 1 and worst score at 0.
# - `support`: the number of actual samples with the given outcome.
# For comparison, let's examine a less performant model to see how the confusion matrix and reports indicate error. Let's use the K-neighbors classifier
# +
# Create the LinearDiscriminantAnalysis object with defaults.
alg = KNeighborsClassifier()
# Create a new model using all of the training data.
alg.fit(Xt, Yt)
# Using the testing data, predict the iris species.
predictions = alg.predict(Xv)
# Let's see the predictions
predictions
# -
accuracy_score(Yv, predictions)
labels = ['versicolor', 'virginica', 'setosa']
cm = confusion_matrix(Yv, predictions, labels=labels)
sns.heatmap(cm, annot=True, xticklabels=labels, yticklabels=labels);
cr = classification_report(Yv, predictions)
print(cr)
# Here we have one prediction that is false where `virginica` was predicted to be `versicolor`
# #### Task 3a: Practice with the random forest classifier
#
# <span style="float:right; margin-left:10px; clear:both;">
# </span>
#
# Now that you have learned how to perform supervised machine learning using a variety of algorithms, lets practice using a new algorithm we haven't looked at yet: the Random Forest Classifier. The random forest classifier builds multiple decision trees and merges them together. Review the sklearn [online documentation for the RandomForestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html). For this task:
#
# 1. Perform a 10-fold cross-validation strategy to see how well the random forest classifier performs with the iris data
# 2. Use a boxplot to show the distribution of accuracy
# 3. Use the `fit` and `predict` functions to see how well it performs with the testing data.
# 4. Plot the confusion matrix
# 5. Print the classification report.
#
| .ipynb_checkpoints/L09-Supervised_Machine_Learning-Lesson-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table>
# <tr><td align="right" style="background-color:#ffffff;">
# <img src="../images/logo.jpg" width="20%" align="right">
# </td></tr>
# <tr><td align="right" style="color:#777777;background-color:#ffffff;font-size:12px;">
# <NAME> | April 27, 2019 (updated)
# </td></tr>
# <tr><td align="right" style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;">
# This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros.
# </td></tr>
# </table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> <font color="blue"> Solution for </font>Random Quantum States</h2>
# <a id="task1"></a>
# <h3> Task 1 </h3>
#
# Define a function randomly creating a quantum state based on the given idea.
#
# Randomly create a quantum state by using this function.
#
# Draw the quantum state on the unit circle.
#
# Repeat the task for a few times.
#
# Randomly create 100 quantum states and draw all of them without labeling them.
# <h3>Solution</h3>
# First, we define our function.
# randomly create a 2-dimensional quantum state
from math import cos, sin, pi
from random import randrange
def random_quantum_state2():
angle_degree = randrange(360)
angle_radian = 2*pi*angle/360
return [cos(angle_radian),sin(angle_radian)]
# Second, we test our function with 6 quantum states.
# +
# include our predefined functions
# %run qlatvia.py
# draw the axes
draw_qubit()
for i in range(6):
[x,y]=random_quantum_state2()
draw_quantum_state(x,y,"|v"+str(i)+">")
# -
# Third, we test our function with 100 quantum states.
# +
# include our predefined functions
# %run qlatvia.py
# draw the axes
draw_qubit()
for i in range(100):
[x,y]=random_quantum_state2()
draw_quantum_state(x,y,"")
# -
| bronze/B54_Random_Quantum_States_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="mIJRdeBNDfMb"
from IPython import display
import pandas as pd
import seaborn as sns
from google.cloud import storage
from typing import Any, List, Tuple
# + id="OapbQ6VIEBY_"
from google.colab import auth
auth.authenticate_user()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 12767, "status": "ok", "timestamp": 1647639715463, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="AhYeG9SuDmb_" outputId="d8baeccb-df80-428e-a8e6-0081ec26892a"
def get_file_paths_with_prefix(project, bucket_name, prefix):
storage_client = storage.Client(project)
blobs = storage_client.list_blobs(bucket_name, prefix=prefix)
return ['gs://%s/%s' %(bucket_name, blob.name) for blob in blobs]
file_paths = get_file_paths_with_prefix('YOUR_PROJECT', 'antibiotic-combination-images', 'site_images')
print('Number of matching files = %d' % len(file_paths))
# + colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"elapsed": 2470, "status": "ok", "timestamp": 1647641120946, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="ZL4sB5etDpB6" outputId="0e9a0197-ee69-4c52-d982-429f89d67ef2"
def make_image_metadata_df(file_paths, image_extension):
file_df = pd.DataFrame()
file_df['path'] = file_paths
file_df = file_df[file_df['path'].str.endswith(image_extension)]
image_metadata_df = file_df.path.str.extract(r'.*site_images\/(\w*)\-(\d*)-([A-Z])(\d{2})-(\d)-0-(\w*).*TIF')
image_metadata_df.columns = ['batch', 'plate', 'well_row', 'well_col', 'site', 'stain']
image_metadata_df['well'] = image_metadata_df['well_row'] + image_metadata_df['well_col']
image_metadata_df['image_path'] = file_df['path']
# Order columns
image_metadata_df = image_metadata_df[['batch', 'plate', 'well', 'site', 'stain', 'well_row', 'well_col', 'image_path']]
return image_metadata_df
image_metadata_df = make_image_metadata_df(file_paths, '.TIF')
with pd.option_context('display.max_colwidth', 300):
display.display(image_metadata_df)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 396, "status": "ok", "timestamp": 1647641252640, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="XrCBnXTlFF9L" outputId="29625da6-331a-45c0-8306-e6bd5381be76"
for c in ['batch', 'plate', 'well', 'site', 'stain']:
print(f'"{c}" unique values are:\n{image_metadata_df[c].unique()}\n')
# + id="qta8HRPuEJVv"
if image_metadata_df.isna().any().any():
raise ValueError('There should be no NaNs in image_metadata_df.')
# + colab={"base_uri": "https://localhost:8080/", "height": 143} executionInfo={"elapsed": 166, "status": "ok", "timestamp": 1647641344214, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="qwKLJhqpFLOB" outputId="ead81cc8-127b-46e6-e76d-cf0a3dbf4a52"
# Specify how sites are arranged within a well.
site_df = pd.DataFrame(
columns=pd.Index(['00', '01'], name='site_col'),
index=pd.Index(['00', '01'], name='site_row'),
data=[['1', '2'],
['3', '4']])
site_df
# + colab={"base_uri": "https://localhost:8080/", "height": 280} executionInfo={"elapsed": 364, "status": "ok", "timestamp": 1647641346753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="CNdqLhTgFNuT" outputId="c40665a5-cbfa-4bd5-9eba-2818fa56fe53"
sns.heatmap(site_df.apply(pd.to_numeric), annot=site_df, fmt='', cmap='viridis', cbar=False);
# + colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"elapsed": 306, "status": "ok", "timestamp": 1647641353376, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="ihUELz2SFO2z" outputId="615c0512-2467-4033-85f9-2132569b5f17"
def get_missing_column_values(subset: pd.DataFrame, sett: pd.DataFrame,
columns: List[str]) -> List[Tuple[Any, ...]]:
"""Returns subset column values that are not in sett column values."""
subset_tuples = subset[columns].drop_duplicates().apply(tuple, axis='columns')
set_tuples = sett[columns].apply(tuple, axis='columns')
column_value_in_set = subset_tuples.isin(set_tuples)
missing_values = list(subset_tuples[~column_value_in_set])
return missing_values
def robust_many_to_one_left_join(left: pd.DataFrame, right: pd.DataFrame,
join_cols: List[str]) -> pd.DataFrame:
# Verify left_df[join_cols] is subset of right_df[join_cols]
missing_column_values = get_missing_column_values(left, right,
join_cols)
if missing_column_values:
raise ValueError('There were missing join values. missing: %s' %
str(missing_column_values))
# The many_to_one validation asserts that the right_df join values are unique.
return pd.merge(
left,
right,
how='left',
on=join_cols,
suffixes=(None, None), # Don't create columns if they already exist.
validate='many_to_one')
def join_site_df(image_metadata_df, site_df):
long_site_df = pd.melt(site_df, value_name='site', ignore_index=False).reset_index()
return robust_many_to_one_left_join(image_metadata_df, long_site_df, join_cols=['site'])
full_image_metadata_df = join_site_df(image_metadata_df, site_df)
full_image_metadata_df
# + colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"elapsed": 1487, "status": "ok", "timestamp": 1647641359181, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhPQEmfwU_PxYdM1KocRvCVpTb8sCbgVT1bym6Xjw=s64", "userId": "07410353554973206216"}, "user_tz": 420} id="ZMLAnJX5FpX8" outputId="852e8f87-4aa3-49b2-fd84-72746bb4a226"
local_image_metadata_csv_path = 'image_metadata.csv'
with open(local_image_metadata_csv_path, 'w') as f:
full_image_metadata_df.to_csv(f, header=True, index=False)
# Read it back to inspect that the written csv looks right.
with open(local_image_metadata_csv_path, 'r') as f:
recovered_df = pd.read_csv(f, dtype=str)
recovered_df
# + id="z7Wn6YBvFr9q"
gcs_image_metadata_csv_path = 'gs://YOUR_BUCKET/YOUR_PATH/image_metadata.csv'
# !gsutil cp {local_image_metadata_csv_path} {gcs_image_metadata_csv_path}
| cell_img/image_grid/file_paths_to_image_metadata_csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# Azure Climate Change Analysis
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import LinearRegression
import seaborn as sns; sns.set()
import pandas as pd
data1 = "./5-year-mean-1951-1980.csv"
data2 = "./5-year-mean-1882-2014.csv"
df1 = pd.read_csv(data1, header=None)
df2 = pd.read_csv(data2, header=None)
# The data was imported without headers and need to be defined.
df1.columns = ['yearsBase','meanBase']
df2.columns = ['years','mean']
print(df1,
"\n",
df2)
# For plotting, the x and y variables will need to be in a series and are defined as follows:
yearsBase = df1['yearsBase']
meanBase = df1['meanBase']
years = df2['years']
mean = df2['mean']
plt.scatter(yearsBase, meanBase)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
# +
# Creates a linear regression from the data points
m,b = np.polyfit(yearsBase, meanBase, 1)
# This is a simple y = mx + b line function
def f(x):
return m*x + b
# This generates the same scatter plot as before, but adds a line plot using the function above
plt.scatter(yearsBase, meanBase)
plt.plot(yearsBase, f(yearsBase))
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
# Prints text to the screen showing the computed values of m and b
print(' y = {0} * x + {1}'.format(m, b))
plt.show()
# +
# Pick the Linear Regression model and instantiate it
model = LinearRegression(fit_intercept=True)
# Fit/build the model
model.fit(yearsBase[:, np.newaxis], meanBase)
mean_predicted = model.predict(yearsBase[:, np.newaxis])
# Generate a plot like the one in the previous exercise
plt.scatter(yearsBase, meanBase)
plt.plot(yearsBase, mean_predicted)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
print(' y = {0} * x + {1}'.format(model.coef_[0], model.intercept_))
# -
plt.scatter(years, mean)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
sns.regplot(yearsBase, meanBase)
plt.show()
| projects/intro-to-ml-with-python/climatechange.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
def number_1(m,k,n):
number = random.randint(m, k)
sum = 0
i = 0
while i<n:
sum = number+sum
i=i+1
return
aver = sum/n
print((sqrt(aver))
#原函数
print(number_1(1,1000,20))
# -
import random
def num_1(m,k):
number = random.randint(m, k)
print(math.log(e,number))
print(1/(math.log(e,number))
#原函数
x = int(input('请输入一个数字,并以回车结束。'))
y = int(input('请输入一个数字,并以回车结束。'))
num_1(x,y)
import random
def num_1(m,k):
number = random.randint(m, k)
print(math.log(e,number))
print(1/(math.log(e,number))
#原函数
x = int(input('请输入一个数字,并以回车结束。'))
y = int(input('请输入一个数字,并以回车结束。'))
num_1(x,y)
| chapter2/homework/computer/4-5/201611680388 (2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import tensorflow as tf
graph = tf.get_default_graph()
graph.get_operations()
for op in graph.get_operations():
print(op.name)
sess = tf.Session()
sess.close()
with tf.Session() as sess:
sess.run(f)
a = tf.constant(1.0)
a
print(a)
with tf.Session() as sess:
print(sess.run(a))
b = tf.Variable(2.0, name = "test_var")
b
init_op = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init_op)
print(sess.run(b))
graph = tf.get_default_graph()
for op in graph.get_operations():
print(op.name)
a = tf.placeholder("float")
b = tf.placeholder("float")
y = tf.multiply(a, b)
feed_dict = {a:2, b:3}
with tf.Session() as sess:
print(sess.run(y, feed_dict))
w = tf.Variable(tf.random_normal([784, 10], stddev=0.01))
b = tf.Variable([10,20,30,40,50,60], name='t')
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
print(sess.run(tf.reduce_mean(b)))
a =[[0.1, 0.2, 0.3],
[20, 2, 3]]
b = tf.Variable(a, name='b')
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
print(sess.run(tf.argmax(b, 1)))
| crackingcode/day1/cc_tf_day1_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## String variable
# If we store any string type ina variable then that will be kniwn as string variables.
#Define variables
a='hello'
b="welcome to python"
c='''Here you can do any job you want '''
#print variables
print(a)
print(b)
print(c)
# Here there are three variables named as a,b and c. each are assigned with strings so all are string variables
#
# String variables are maintaining indexing property
# 
# ### Operations in string
#define a string variable
var1 = '<NAME>'
#Get length of the string
print(len(var1))
#Get the index of a alphabet
print(var1.index("o"))
#Count how many times a letter is repeted
print(var1.count("l"))
#prints complete string
print(var1)
#prints first charecter of the string
print(var1[0])
#prints charecters starting from 3rd to 5th
print(var1[2:5])
#prints string starting from 3rd to last
print(var1[2:])
# +
#prints the characters of string from 3 to 7 skipping one character.
#This is extended slice syntax. The general form is [start:stop:step].
print(var1[3:7:1])
# -
#reverse a string
print(var1[::-1])
#prints the string two times
print(var1 * 2)
#prints concatenated string
print(var1 + " Test")
# ## String Formatting
#
# Suppose you want to add a variable value to your string on a perticular place, In that case you can use string formatting
#define a variable
name = "jag"
#add name to the string
"Hi {}, Welcome to Python".format(name)
#you can also use formatted string
f"Hi {name}, Welcome to Python"
#Format using index
print("I like {1} and {0}".format("Java", "Python"))
# +
# unpacking from sequences
t = ("Java", "Python")
print("I like {1} and {0}".format(*t))
l = ["Java", "Python"]
print("I like {} and {}".format(*t))
# -
#Keyword arguments
print("{name} is the {job} of {company}".format(name='<NAME>', job='CEO', company='Apple Inc.'))
#Dictionary as Keyword arguments
d = {"name": "<NAME>", "job": "CEO", "company": "Apple Inc."}
print("{company} {job} is {name}".format(**d))
#integer to different bases
print("int: {0:d}; hex: {0:x}; oct: {0:o}; bin: {0:b}".format(28))
#separating number with comma
print('{:,}'.format(1234567890))
#Percentage, Padding and Rounding
print('Percentage: {:.3%}'.format(19 / 28))
print('{0:7.2f}'.format(2.344))
print('{0:10.2f}'.format(22222.346))
#get the string in lower case
name.lower()
#Get the string as a title
name.title()
#get the string in Upper case
name.upper()
#Check if starts with some letters
print(var1.startswith("Hello"))
##Check if ends with some letters
print(var1.endswith("asdfasdfasdf"))
#check if string have some letters
"ell" in var1
#split a string
var1.split(" ") #Here we are spliting with space
# ## String Templating
#
# In String module, Template Class allows us to create simplified syntax for output specification. The format uses placeholder names formed by \\$ with valid Python identifiers (alphanumeric characters and underscores). Surrounding the placeholder with braces allows it to be followed by more alphanumeric letters with no intervening spaces. Writing $$ creates a single escaped $:
#import Template
from string import Template
# Create a template that has placeholder for value of x
t = Template('$name is the $job of $company')
#print the template
print('Template String =', t.template)
# Substitute value of x in above template
out_string = t.substitute(name='<NAME>', job='CEO', company='Apple Inc.')
print(out_string)
# dictionary as substitute argument
d = {"name": "<NAME>", "job": "CEO", "company": "Apple Inc."}
s = t.substitute(**d)
print(s)
#Safe_substitute for passing parameters less than required
s = t.safe_substitute(name='<NAME>', job='CEO')
print(s)
| Day1/Python-Programming-Tutorials-master/e__string_variable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# %env MKL_NUM_THREADS=16
# %env OMP_NUM_THREADS=16
# +
import numpy as np
import pandas as pd
from ipypb import track
from polara.evaluation import evaluation_engine as ee
from polara.evaluation.pipelines import find_optimal_svd_rank
from polara import (SVDModel,
PopularityModel,
RandomModel)
from polara.recommender.hybrid.models import SimilarityAggregation
from polara.recommender.coldstart.models import (SVDModelItemColdStart,
RandomModelItemColdStart,
PopularityModelItemColdStart,
SimilarityAggregationItemColdStart)
from data_preprocessing import (get_yahoo_music_data,
get_similarity_data,
prepare_data_model,
prepare_cold_start_data_model)
from utils import (report_results, save_results,
apply_config, print_data_stats,
save_training_time, save_cv_training_time)
# %matplotlib inline
# -
from polara.recommender import defaults
defaults.memory_hard_limit = 15 # allowed memory usage during recommendations generation
max_test_workers = 6 # use this manyparallel thread for evaluation each using up to {memory_hard_limit} Gb of RAM
seed = 42
experiment_name = 'baseline'
# # Experiment setup
data_labels = ['YaMus']
ranks_grid = [1, 15, 30, 50, 75, 100, 150, 200, 250, 300, 400,
500, 750, 1000, 1250, 1500, 2000, 2500, 3000]
svd_ranks = {'YaMus': ranks_grid}
topk_values = [1, 3, 10, 20, 30]
target_metric = 'mrr'
data_dict = dict.fromkeys(data_labels)
meta_dict = dict.fromkeys(data_labels)
similarities = dict.fromkeys(data_labels)
sim_indices = dict.fromkeys(data_labels)
feature_idx = dict.fromkeys(data_labels)
all_data = [data_dict, similarities, sim_indices, meta_dict]
# ## Yahoo Music
lbl = 'YaMus'
data_dict[lbl], meta_dict[lbl] = get_yahoo_music_data('/gpfs/gpfs0/e.frolov/recsys/yahoo_music/yamus_train0_rating5.gz',
meta_path='/gpfs/gpfs0/e.frolov/recsys/yahoo_music/yamus_attrs.gz',
implicit=True,
pcore=5,
filter_data={'genreid': [0]}, # filter unknown genre
filter_no_meta=True)
similarities[lbl], sim_indices[lbl], feature_idx[lbl] = get_similarity_data(meta_dict[lbl])
(meta_dict[lbl].applymap(len).sum(axis=1)==0).mean()
# ## Data stats
print_data_stats(data_labels, all_data)
# # Standard experiment
# +
def prepare_recommender_models(data_label, data_models, config):
data_model = data_models[data_label]
models = [SVDModel(data_model),
SimilarityAggregation(data_model),
PopularityModel(data_model),
RandomModel(data_model, seed=seed)]
for model in models:
model.max_test_workers = max_test_workers
apply_config(models, config, data_label)
return models
def fine_tune_svd(model, ranks, label, record_time=False):
model.max_test_workers = max_test_workers
best_svd_rank, svd_scores = find_optimal_svd_rank(model, ranks, target_metric,
return_scores=True,
iterator=lambda x: track(x, label=f'{label} ranks'))
model_config = {model.method: {'rank': best_svd_rank}}
model_scores = {model.method: svd_scores}
try:
if record_time:
save_training_time(experiment_name, model, pd.Index([max(ranks)], name='rank'), label)
finally:
return model_config, model_scores
# -
# ## tuning
config = {}
scores = {}
data_models = {}
for label in track(data_labels):
data_models[label] = prepare_data_model(label, *all_data, seed)
config[label], scores[label] = fine_tune_svd(SVDModel(data_models[label]),
svd_ranks[label],
label, True)
report_results('rank', scores);
config
# ### saving data
save_results(experiment_name, config=config, tuning=scores)
# ## cross-validation
# +
result = {}
for label in track(data_labels):
models = prepare_recommender_models(label, data_models, config)
result[label] = ee.run_cv_experiment(models,
fold_experiment=ee.topk_test,
topk_list=topk_values,
ignore_feedback=True,
iterator=lambda x: track(x, label=f'{label} folds'))
save_cv_training_time(experiment_name, models, label)
# -
report_results('topn', result, target_metric);
# ### saving data
save_results(experiment_name, cv=result)
# # Cold start
# + active=""
# import gc
# gc.collect()
# -
def prepare_cold_start_recommender_models(data_label, data_models, config):
data_model = data_models[data_label]
models = [SVDModelItemColdStart(data_model, item_features=meta_dict[data_label]),
SimilarityAggregationItemColdStart(data_model),
PopularityModelItemColdStart(data_model),
RandomModelItemColdStart(data_model, seed=seed)]
for model in models:
model.max_test_workers = max_test_workers
apply_config(models, config, data_label)
return models
# ## tuning
config_cold = {}
scores_cold = {}
data_models_cold = {}
for label in track(data_labels):
data_models_cold[label] = prepare_cold_start_data_model(label, *all_data, seed)
model = SVDModelItemColdStart(data_models_cold[label],
item_features=meta_dict[label])
model.use_raw_features = True
config_cold[label], scores_cold[label] = fine_tune_svd(model,
svd_ranks[label],
label)
report_results('rank', scores_cold);
config_cold
# ### saving data
save_results(experiment_name+'_coldstart', config=config_cold, tuning=scores_cold)
# ## cross validation
result_cold = {}
for label in track(data_labels):
models_cold = prepare_cold_start_recommender_models(label, data_models_cold, config_cold)
result_cold[label] = ee.run_cv_experiment(models_cold,
fold_experiment=ee.topk_test,
topk_list=topk_values,
ignore_feedback=True,
iterator=lambda x: track(x, label=f'{label} folds'))
report_results('topn', result_cold, target_metric);
report_results('topn', result_cold, 'coverage');
# ### saving data
save_results(experiment_name+'_coldstart', cv=result_cold)
| recsys19_hybridsvd/Baselines_YaMus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.4
# language: julia
# name: julia-1.0
# ---
using Revise # lets you change A2funcs without restarting julia!
includet("A2_src.jl")
using Plots
using Statistics: mean
using Zygote
using Test
using Logging
using .A2funcs: log1pexp # log(1 + exp(x)) stable
using .A2funcs: factorized_gaussian_log_density
using .A2funcs: skillcontour!
using .A2funcs: plot_line_equal_skill!
function log_prior(zs)
return factorized_gaussian_log_density(0, 0, zs)
end
function logp_a_beats_b(za,zb)
return -log1pexp(zb-za)
end
function all_games_log_likelihood(zs,games)
zs_a = zs[games[:,1],:]
zs_b = zs[games[:,2],:]
likelihoods = logp_a_beats_b.(zs_a, zs_b)
return sum(likelihoods, dims=1)
end
function joint_log_density(zs,games)
return log_prior(zs) + all_games_log_likelihood(zs, games)
end
@testset "Test shapes of batches for likelihoods" begin
B = 15 # number of elements in batch
N = 4 # Total Number of Players
test_zs = randn(4,15)
test_games = [1 2; 3 1; 4 2] # 1 beat 2, 3 beat 1, 4 beat 2
@test size(test_zs) == (N,B)
#batch of priors
@test size(log_prior(test_zs)) == (1,B)
# loglikelihood of p1 beat p2 for first sample in batch
@test size(logp_a_beats_b(test_zs[1,1],test_zs[2,1])) == ()
# loglikelihood of p1 beat p2 broadcasted over whole batch
@test size(logp_a_beats_b.(test_zs[1,:],test_zs[2,:])) == (B,)
# batch loglikelihood for evidence
@test size(all_games_log_likelihood(test_zs,test_games)) == (1,B)
# batch loglikelihood under joint of evidence and prior
@test size(joint_log_density(test_zs,test_games)) == (1,B)
end
# +
# Convenience function for producing toy games between two players.
two_player_toy_games(p1_wins, p2_wins) = vcat([repeat([1,2]',p1_wins), repeat([2,1]',p2_wins)]...)
# Example for how to use contour plotting code
plot(title="Example Gaussian Contour Plot",
xlabel = "Player 1 Skill",
ylabel = "Player 2 Skill"
)
# TODO: plot prior contours
example_gaussian(zs) = exp(factorized_gaussian_log_density([-1.,2.],[0.,0.5],zs))
skillcontour!(example_gaussian)
plot_line_equal_skill!()
savefig(joinpath("plots","prior_contours"))
# -
# TODO: plot likelihood contours
plot(title="Likelihood Contour Plot",
xlabel = "Player 1 Skill",
ylabel = "Player 2 Skill")
likelihood(zs) = exp.(logp_a_beats_b.(zs[1,:], zs[2,:]))
skillcontour!(likelihood)
plot_line_equal_skill!()
savefig(joinpath("plots", "likelihood_contours"))
# TODO: plot joint contours with player A winning 1 game
plot(title="Joint Contour Plot 1 Game",
xlabel = "Player 1 Skill",
ylabel = "Player 2 Skill")
one_game = two_player_toy_games(1, 0)
joint_posterior_1(zs) = exp(joint_log_density(zs, one_game))
skillcontour!(joint_posterior_1)
plot_line_equal_skill!()
savefig(joinpath("plots", "posterior_contours_1"))
# TODO: plot joint contours with player A winning 10 games
plot(title="Joint Contour Plot 10 Games (d)",
xlabel = "Player 1 Skill",
ylabel = "Player 2 Skill")
ten_games = two_player_toy_games(10, 0)
joint_posterior_10d(zs) = exp(joint_log_density(zs, ten_games))
skillcontour!(joint_posterior_10d)
plot_line_equal_skill!()
savefig(joinpath("plots", "posterior_contours_10d"))
#TODO: plot joint contours with player A winning 10 games and player B winning 10 games
plot(title="Joint Contour Plot 20 Games (e)",
xlabel = "Player 1 Skill",
ylabel = "Player 2 Skill")
twenty_games = two_player_toy_games(10, 10)
joint_posterior_20e(zs) = exp(joint_log_density(zs, twenty_games))
skillcontour!(joint_posterior_20e)
plot_line_equal_skill!()
savefig(joinpath("plots", "posterior_contours_20e"))
function elbo(params, logp, num_samples)
mu, logsig = params
samples = mu .+ exp.(logsig) .* randn(size(mu)[1], num_samples)
logp_estimate = logp(samples)
logq_estimate = factorized_gaussian_log_density(mu, logsig, samples)
return mean(logp_estimate .- logq_estimate)#TODO: should return scalar (hint: average over batch)
end
# Conveinence function for taking gradients
function neg_toy_elbo(params; games = two_player_toy_games(1,0), num_samples = 100)
# TODO: Write a function that takes parameters for q,
# evidence as an array of game outcomes,
# and returns the -elbo estimate with num_samples many samples from q
logp(zs) = joint_log_density(zs,games)
return -elbo(params,logp, num_samples)
end
# Toy game
num_players_toy = 2
toy_mu = [-2.,3.] # Initial mu, can initialize randomly!
toy_ls = [0.5,0.] # Initual log_sigma, can initialize randomly!
toy_params_init = (toy_mu, toy_ls)
function fit_toy_variational_dist(init_params, toy_evidence; num_itrs=200, lr= 1e-2, num_q_samples = 10, fp="TvsV")
params_cur = init_params
for i in 1:num_itrs
grad_params = gradient(params_cur -> neg_toy_elbo(params_cur; games=toy_evidence, num_samples=num_q_samples), params_cur)#TODO: gradients of variational objective with respect to parameters
mu, logsig = params_cur
mu -= lr .* grad_params[1][1]
logsig -= lr .* grad_params[1][2]
params_cur = mu, logsig #TODO: update paramters with lr-sized step in descending gradient
e = neg_toy_elbo(params_cur; games=toy_evidence, num_samples=num_q_samples)
@info "ELBO:" e#TODO: report the current elbbo during training
# TODO: plot true posterior in red and variational in blue
# hint: call 'display' on final plot to make it display during training
plot(title="Target Dist vs Variational Approx",
xlabel="Player 1 Skill",
ylabel="Player 2 Skill");
if i == num_itrs - 1
true_dist(zs) = exp(joint_log_density(zs, toy_evidence))
variational_dist(zs) = exp(factorized_gaussian_log_density(mu, logsig, zs))
skillcontour!(true_dist,colour=:red) # plot likelihood contours for target posterior
plot_line_equal_skill!()
display(skillcontour!(variational_dist, colour=:blue)) # plot likelihood contours for variational posterior
#TODO: save final posterior plots
savefig(joinpath("plots", fp))
end
end
return params_cur
end
#TODO: fit q with SVI observing player A winning 1 game
one_game = two_player_toy_games(1, 0)
fp = "Toy_vs_Var_1"
fitted = fit_toy_variational_dist(toy_params_init, one_game; fp=fp)
#TODO: fit q with SVI observing player A winning 10 games
ten_games = two_player_toy_games(10, 0)
fp = "Toy_vs_Var_10"
fitted = fit_toy_variational_dist(toy_params_init, ten_games, fp=fp)
#TODO: save final posterior plots
#TODO: fit q with SVI observing player A winning 10 games and player B winning 10 games
twenty_games = two_player_toy_games(10, 10)
fp = "Toy_vs_Var_20"
fitted = fit_toy_variational_dist(toy_params_init, twenty_games, fp=fp)
#TODO: save final posterior plots
## Question 4
# Load the Data
using MAT
vars = matread("tennis_data.mat")
player_names = vars["W"]
tennis_games = Int.(vars["G"])
num_players = length(player_names)
print("Loaded data for $num_players players")
function fit_variational_dist(init_params, tennis_games; num_itrs=200, lr= 1e-2, num_q_samples = 10)
params_cur = init_params
for i in 1:num_itrs
grad_params = gradient(params_cur -> neg_toy_elbo(params_cur; games=tennis_games, num_samples=num_q_samples), params_cur)#TODO: gradients of variational objective with respect to parameters
mu, logsig = params_cur
mu -= lr .* grad_params[1][1]
logsig -= lr .* grad_params[1][2]
params_cur = mu, logsig #TODO: update paramters with lr-sized step in descending gradient
e = neg_toy_elbo(params_cur; games=tennis_games, num_samples=num_q_samples)
@info "ELBO:" e#TODO: report the current elbbo during training
end
return params_cur
end
# TODO: Initialize variational family
init_mu = randn(num_players)#random initialziation
init_log_sigma = rand(num_players)# random initialziation
init_params = (init_mu, init_log_sigma)
# Train variational distribution
trained_params = fit_variational_dist(init_params, tennis_games)
#TODO: 10 players with highest mean skill under variational model
#hint: use sortperm
means, logstd = trained_params
perm = sortperm(means)
plot(means[perm], yerror=exp.(logstd[perm]),
title="Approximate Mean and Variance",
xlabel = "Player",
ylabel = "Skill", label="Mean")
savefig(joinpath("plots", "apx_mean_var"))
reverse(player_names[perm[num_players-9:end]]) # Top ten
#TODO: joint posterior over "Roger-Federer" and ""Rafael-Nadal""
RF = findall(x -> x == "Roger-Federer", player_names)
RN = findall(x -> x == "Rafael-Nadal", player_names)
mu = means[RF, RN]
logsig = logstd[RF, RN]
variational_dist(zs) = exp(factorized_gaussian_log_density(mu, logsig, zs))
plot(title="Nadal vs Federer",
xlabel = "Federer Skill",
ylabel = "Nadal Skill")
skillcontour!(variational_dist)
plot_line_equal_skill!()
savefig(joinpath("plots", "Fed_v_Nad"))
#hint: findall function to find the index of these players in player_names
using Distributions
# P(Federer has higher skill than Nadal)
# Exact
mu = means[RF][1] - means[RN][1]
var = exp(logstd[RF][1])^2 + exp(logstd[RN][1])^2
D = Normal(mu, sqrt(var))
p = 1 - cdf(D, 0)
# Monte Carlo
count = 0
for i in 1:10000
z = mu + randn()*sqrt(var)
if z > 0
count += 1
end
end
p = count/10000
# Federer vs Worst Player
player = perm[1]
mu = means[RF][1] - means[player][1]
var = exp(logstd[RF][1])^2 + exp(logstd[player][1])^2
D = Normal(mu, sqrt(var))
p = 1 - cdf(D, 0)
# Monte Carlo
count = 0
for i in 1:10000
z = mu + randn()*sqrt(var)
if z > 0
count += 1
end
end
p = count/10000
| STA/SVI/A2_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="JMx3BR_Doatb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 182} outputId="63d9ecba-09ca-46be-912a-70f4ba6432c2" executionInfo={"status": "ok", "timestamp": 1585944122470, "user_tz": -120, "elapsed": 4319, "user": {"displayName": "Kamil", "photoUrl": "", "userId": "13475327348794705332"}}
# !pip install hyperopt
# + id="pbwdvt-QosMf" colab_type="code" colab={}
import pandas as pd
import numpy as np
import os
import datetime
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPool2D, Dense, Flatten, Dropout
from tensorflow.keras.utils import to_categorical
import matplotlib.pyplot as plt
from skimage import color, exposure
from sklearn.metrics import accuracy_score
from hyperopt import hp, STATUS_OK, tpe, Trials, fmin
# + id="eYauYZ5wo652" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="aa935e3e-33a7-4abc-b88b-0cebc35a365d" executionInfo={"status": "ok", "timestamp": 1585944330294, "user_tz": -120, "elapsed": 675, "user": {"displayName": "Kamil", "photoUrl": "", "userId": "13475327348794705332"}}
# cd '/content/drive/My Drive/Colab Notebooks/dw_matrix_three'
# + id="5v1CEt97o8F3" colab_type="code" colab={}
train = pd.read_pickle('data/train.p')
test = pd.read_pickle('data/test.p')
X_train, y_train = train['features'], train['labels']
X_test, y_test = test['features'], test['labels']
# + id="OimaT_tMo_zm" colab_type="code" colab={}
if y_train.ndim == 1: y_train = to_categorical(y_train)
if y_test.ndim == 1: y_test = to_categorical(y_test)
# + id="qcljzdvqpA1e" colab_type="code" colab={}
input_shape = X_train.shape[1:]
num_classes = y_train.shape[1]
# + id="ZRXjePJipEIP" colab_type="code" colab={}
def train_model(model, X_train, y_train, params_fit={}):
model.compile(loss='categorical_crossentropy', optimizer='Adam', metrics=['accuracy'])
logdir = os.path.join("logs", datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
tensorboard_callback = tf.keras.callbacks.TensorBoard(logdir, histogram_freq=1)
model.fit(
X_train,
y_train,
batch_size=params_fit.get('batch_size', 128),
epochs=params_fit.get('epochs', 5),
verbose=params_fit.get('verbose',1),
validation_data=params_fit.get('validation_data', (X_train, y_train)),
callbacks=[tensorboard_callback]
)
return model
# + id="J5Gbg0jTpGE-" colab_type="code" colab={}
def predict(trained_model, X_test, y_test, scoring=accuracy_score):
y_pred_probe = trained_model.predict(X_test)
y_pred = np.argmax(y_pred_probe, axis=1)
y_test_norm = np.argmax(y_test, axis=1)
return scoring(y_test_norm, y_pred)
# + id="N-ONThw-pJ-H" colab_type="code" colab={}
def get_cnn_v5(input_shape, num_classes):
return Sequential([
Conv2D(filters=32, kernel_size=(3,3), activation='relu', input_shape=input_shape),
Conv2D(filters=32, kernel_size=(3,3), activation='relu', padding='same'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3,3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3,3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3,3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3,3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Flatten(),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(num_classes, activation='softmax')
])
# + id="7DoAE2PZpWP4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 219} outputId="ebdc57f7-f6f9-4bf0-e33b-ce0c3a36ea51" executionInfo={"status": "ok", "timestamp": 1585944422389, "user_tz": -120, "elapsed": 33841, "user": {"displayName": "Kamil", "photoUrl": "", "userId": "13475327348794705332"}}
model = get_cnn_v5(input_shape, num_classes)
trained_model = train_model(model, X_train, y_train)
predict(trained_model, X_test, y_test)
# + id="Rin2AnwBqcJh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="a57040f3-9964-4e22-98ec-492f2b28511b" executionInfo={"status": "ok", "timestamp": 1585944593866, "user_tz": -120, "elapsed": 1033, "user": {"displayName": "Kamil", "photoUrl": "", "userId": "13475327348794705332"}}
trained_model.evaluate(X_test, y_test)[0]
# + id="J8KhA4QQsFgx" colab_type="code" colab={}
def get_model(params):
return Sequential([
Conv2D(filters=32, kernel_size=(3,3), activation='relu', input_shape=input_shape),
Conv2D(filters=32, kernel_size=(3,3), activation='relu', padding='same'),
MaxPool2D(),
Dropout(params['dropout_cnn_block_one']),
Conv2D(filters=64, kernel_size=(3,3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3,3), activation='relu'),
MaxPool2D(),
Dropout(params['dropout_cnn_block_two']),
Conv2D(filters=128, kernel_size=(3,3), activation='relu', padding='same'),
Conv2D(filters=128, kernel_size=(3,3), activation='relu'),
MaxPool2D(),
Dropout(params['dropout_cnn_block_three']),
Flatten(),
Dense(1024, activation='relu'),
Dropout(params['dropout_dense_block_one']),
Dense(1024, activation='relu'),
Dropout(params['dropout_dense_block_two']),
Dense(num_classes, activation='softmax')
])
# + id="V2C7YtPHp6_M" colab_type="code" colab={}
def func_obj(params):
model = get_model(params)
model.compile(loss='categorical_crossentropy', optimizer='Adam', metrics=['accuracy'])
model.fit(
X_train,
y_train,
batch_size=int(params.get('batch_size', 128)),
epochs=5,
verbose=0
)
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = score[1]
print(params, 'accuracy={}'.format(accuracy))
return {'loss': -accuracy, 'status': STATUS_OK, 'model': model}
# + id="46wB7EPirRYk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6dd8b64d-e093-4fa9-90d4-ab762eb7c232" executionInfo={"status": "ok", "timestamp": 1585946482400, "user_tz": -120, "elapsed": 315944, "user": {"displayName": "Kamil", "photoUrl": "", "userId": "13475327348794705332"}}
space = {
'batch_size': hp.quniform('batch_size', 100, 200, 10),
'dropout_cnn_block_one': hp.uniform('dropout_cnn_block_one', 0.3, 0.6),
'dropout_cnn_block_two': hp.uniform('dropout_cnn_block_two', 0.3, 0.6),
'dropout_cnn_block_three': hp.uniform('dropout_cnn_block_three', 0.3, 0.6),
'dropout_dense_block_one': hp.uniform('dropout_dense_block_one', 0.3, 0.7),
'dropout_dense_block_two': hp.uniform('dropout_dense_block_two', 0.3, 0.7)
}
best = fmin(
func_obj,
space,
tpe.suggest,
30,
Trials()
)
# + id="NXZ-DTTWrsnW" colab_type="code" colab={}
| day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Custom (test-ml-python)
# language: python
# name: test-ml-python
# ---
# ## Example 4 - Blocking the right paths in the network
#
# Here we investigate the the hypothesised pathway from Barents and Kara sea ice (BK) in autumn to the Stratospheric polar vortex (SPV) in winter via affecting sea level pressure over the Ural Mountains region (URAL). The latter is also assumed to affect BK. Moreover, the El Niño Southern Oscillation (ENSO) and the Madden Julien Oscillation (MJO) influence North Pacific sea level pressure (NP), and thereby both the SPV and BK.
#
#
# <img src="../images/ex4.png" width="500" height="600">
# # Imports
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import os
import iris
import iris.quickplot as qplt
import iris.coord_categorisation as coord_cat
import statsmodels.api as sm
from scipy import signal
from scipy import stats
# ## Step 1) Load the data + Extract regions of interest
#
bk_sic = iris.load_cube('../sample_data/bk_sic.nc', "sic")
nh_spv = iris.load_cube('../sample_data/nh_spv_uwnd.nc', "uwnd")
np_slp = iris.load_cube('../sample_data/np_slp.nc', "slp")
ural_slp = iris.load_cube('../sample_data/ural_slp.nc', "slp")
# +
# make seasonal means
def do_mean_over_months(data_cube, list_months):
# extract months of interest
ond_constraint = iris.Constraint(month=lambda v: v in list_months) # ['Oct','Nov', 'Dec'])
precip_ond = data_cube.extract(ond_constraint)
# create the mean
precip_ond_mean = precip_ond.aggregated_by(['year'],iris.analysis.MEAN)
return precip_ond_mean
# -
bk = do_mean_over_months(bk_sic, ['Oct','Nov', 'Dec'])
spv = do_mean_over_months(nh_spv, ['Jan','Feb', 'Mar'])
ural = do_mean_over_months(ural_slp, ['Oct','Nov', 'Dec'])
pac = do_mean_over_months(np_slp, ['Oct','Nov', 'Dec'])
# ### plot the time-series
# +
fig = plt.figure(figsize=(8, 8))
plt.subplot(411)
qplt.plot(bk)
plt.title('BK-SIC')
plt.subplot(412)
qplt.plot(ural)
plt.title('Ural_slp')
plt.subplot(413)
qplt.plot(pac)
plt.title('NP_slp')
plt.subplot(414)
qplt.plot(spv)
plt.title('NH-SPV')
plt.tight_layout()
# -
# ## Step 2) Data processing
#
# #### standardize
BK = (bk - np.mean(bk.data))/np.std(bk.data)
SPV = (spv - np.mean(spv.data))/np.std(spv.data)
URAL = (ural - np.mean(ural.data))/np.std(ural.data)
NP = (pac - np.mean(pac.data))/np.std(pac.data)
# #### detrend
y0 = 0
BK = signal.detrend(BK[y0:].data)
SPV = signal.detrend(SPV[y0:].data)
URAL = signal.detrend(URAL[y0:].data)
NP = signal.detrend(NP[y0:].data)
# ## Step 3) Data analysis
# +
#================================================================
# Determine the effect of ENSO on CA conditioned on Jet
#================================================================
# note the one-calendar year lag between the autumn drivers BK, URAL, NO and the reponse variable of winter SPV
X = np.stack([BK[:-1], URAL[:-1], NP[:-1]]).T
Y = SPV[1: ]
model = sm.OLS(Y,X)
results = model.fit()
ce_x1 = results.params[0]
ce_x2 = results.params[1]
ce_x3 = results.params[2]
print("The causal effect of BK-SIC on SPV is (cond on URAL , NP)", round(ce_x1,3))
print('\n')
print("The regression coeff. of URAL on SPV is ", round(ce_x2,3))
print("The regression coeff. of NP on SPV is ", round(ce_x3,3))
| causality_paper/notebooks/example4_blocking_paths.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Value at Risk: A simple way to monitor market risk with atoti
#
# Financial institutions all have to find a balance between profit and risk. The more risk taken the higher the profit can be. However if we want to avoid collapses such as that of Lehman Brothers in 2008, risk has to be controlled.
#
# There are several kinds of risk:
# - Shortfall of a counterparty, also known as credit risk: This is the risk that a borrower cannot repay its credit
# - Market risk: This is the risk that certain assets could lose their value. For example one might invest in wine bottle in the hope that they gain value with age while they might not.
#
# Market risk is widely monitored in finance. Institutions have large portfolios with a lot of assets, and forecasting the value of each asset is simply impossible as COVID-19 kindly reminded us recently. The key is then to assess what are the (statistical) chances that the value of certain assets remain in a certain envelope and what the potential losses are. This is where the value at risk – or VaR – comes into action.
#
# There are different approaches to calculating the VaR. The one we will use in this notebook is based on the aggregation of simulated profit & losses, and then calculated using a percentile of the empirical distribution.
#
# We will see how we can compute and aggregate pretty easily this non-linear indicator with atoti, and then perform simulations around it.
#
# <div style="text-align:center"><a href="https://www.atoti.io/?utm_source=gallery&utm_content=value-at-risk" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover.png" alt="atoti" /></a></div>
# ## Importing the necessary libraries
import atoti
# ## Data Loading
# #### Initializing atoti
# tell atoti to load the database containing the UI dashboards
session = atoti.create_session(config={"user_content_storage": "content"})
# #### Loading the data
# Instruments are financial products. In this notebook they are foreign exchange options.
# +
# uncomment this line to install atoti-aws
# # !conda install atoti-aws -y
# -
instruments = session.read_csv(
"s3://data.atoti.io/notebooks/var/instruments.csv",
keys=["instrument_code"],
table_name="Instruments",
)
instruments.head()
# The analytics table gives more information on each instrument, more notably:
# - The PnL (profit and loss) of the previous day
# - A vector of the PnLs of the instrument for the last 372 days. PnLs are typically calculated by complex price engines and such vectors would be their output.
analytics = session.read_csv(
"s3://data.atoti.io/notebooks/var/simulated_pl_vol_depth_150.csv",
keys=["instrument_code"],
table_name="Instruments Analytics",
array_separator=";",
)
analytics.head()
# We will force the type of those two columns so that when using auto mode to create the cube, they will directly create sum and avg measures.
# Since Int columns create hierarchies in auto mode, another solution would have been to create the measures manually.
positions_table_types = {
"quantity": atoti.type.DOUBLE,
"purchase_price": atoti.type.DOUBLE,
}
# Positions give us the quantities of each instrument we currently hold in our portfolio.
# They are grouped into books.
positions = session.read_csv(
"s3://data.atoti.io/notebooks/var/eod_positions.csv",
keys=["instrument_code", "book_id"],
table_name="Positions",
types=positions_table_types,
)
positions.head()
# ### Data model and cube
# We will first join the three previous tables altogether.
positions.join(instruments)
instruments.join(analytics)
# To start our analysis, we create our cube using `Positions` as the base table.
cube = session.create_cube(positions, "Positions")
cube.schema
# In auto mode, atoti creates hierarchies for each column that is not of type float, sum and average measures for each column of type float.
# This can of course be fine-tuned to either switch to full manual mode and create hierarchies/measures yourself, or simply edit what has been created automatically (adding a hierarchy for a numerical column for example). The available cube creation modes are detailed in the [documentation](https://docs.atoti.io).
#
# Below you can explore which measures/levels/hierarchies have been automatically created in our cube.
m, h, lvl = cube.measures, cube.hierarchies, cube.levels
cube
# #### Computing the PnL of the previous day
m["pnl.VALUE"] = atoti.value(analytics["pnl"])
m["pnl_vector.VALUE"] = atoti.value(analytics["pnl_vector"])
# A simple command lets you run atoti's UI directly in the notebook. This is pretty convenient to explore the data you just loaded or make sure the measures defined produce the correct results.
# + atoti={"height": 333, "widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[quantity.SUM]", "[Measures].[pnl.VALUE]", "[Measures].[pnl_vector.VALUE]"], "rows": ["[Positions].[underlying].[underlying]"]}, "name": "", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY Crossjoin({[Measures].[quantity.SUM], [Measures].[pnl.VALUE], [Measures].[pnl_vector.VALUE]}, Hierarchize(DrilldownLevel([Instruments].[option_type].[ALL].[AllMember]))) ON COLUMNS, NON EMPTY Union(Hierarchize(DrilldownLevel([Positions].[underlying].[ALL].[AllMember])), Hierarchize(Descendants({[Positions].[underlying].[AllMember]}, 1, SELF_AND_BEFORE))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# ### Looking at the PnL in various ways
# Compute the PnL for each instrument by multiplying the PnL value against the quantity against the `instrument_code` level.
# Above the scope level, the PnL will be aggregated.
m["PnL"] = atoti.agg.sum(
m["quantity.SUM"] * m["pnl.VALUE"], scope=atoti.scope.origin(lvl["instrument_code"])
)
# Run the following cells to see the atoti visualizations
# + activeviam={"state": {"name": "PnL Pivot Table", "type": "container", "value": {"body": {"configuration": {"tabular": {"addButtonFilter": "numeric", "columns": [], "columnsGroups": [{"captionProducer": "firstColumn", "cellFactory": "kpi-status", "selector": "kpi-status"}, {"captionProducer": "firstColumn", "cellFactory": "lookup", "selector": "lookup"}, {"captionProducer": "expiry", "cellFactory": "expiry", "selector": "kpi-expiry"}, {"captionProducer": "columnMerge", "cellFactory": {"args": {"automaticExpansion": true}, "key": "treeCells"}, "selector": "member"}], "defaultOptions": {}, "hideAddButton": true, "lineNumbers": true, "pinnedHeaderSelector": "member", "sortingMode": "non-breaking", "statisticsShown": true}}, "contextValues": {}, "mdx": "SELECT NON EMPTY Crossjoin(Hierarchize(DrilldownLevel([Hierarchies].[underlying].[ALL].[AllMember])), Hierarchize(DrilldownLevel([Hierarchies].[option_type].[ALL].[AllMember]))) ON ROWS, NON EMPTY {[Measures].[contributors.COUNT], [Measures].[Quantity], [Measures].[PnL]} ON COLUMNS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "ranges": {"column": {"chunkSize": 50, "thresholdPercentage": 0.2}, "row": {"chunkSize": 2000, "thresholdPercentage": 0.1}}, "serverUrl": "", "updateMode": "once"}, "containerKey": "pivot-table", "showTitleBar": false, "style": {}}}} atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[PnL]", "[Measures].[pnl.VALUE]", "[Measures].[quantity.SUM]"], "rows": ["[Positions].[underlying].[underlying]", "[Positions].[instrument_code].[instrument_code]"]}, "name": "PnL Pivot Table", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY {[Measures].[PnL], [Measures].[pnl.VALUE], [Measures].[quantity.SUM]} ON COLUMNS, NON EMPTY Crossjoin(Union([Positions].[underlying].[underlying].Members, Hierarchize(Descendants({[Positions].[underlying].[AllMember]}, 1, SELF_AND_BEFORE))), Hierarchize(Descendants({[Positions].[instrument_code].[AllMember]}, 1, SELF_AND_BEFORE))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}}
session.visualize(name="PnL Pivot Table")
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "values": ["[Measures].[PnL]"], "verticalSubplots": [], "xAxis": ["[Positions].[underlying].[underlying]"]}, "name": "", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY [Measures].[PnL] ON COLUMNS, NON EMPTY [Positions].[underlying].[underlying].Members ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-waterfall-chart"}}
session.visualize()
# -
# ### Collaboration tools
# All the tables/charts created in the notebook can be published and made available in atoti's UI, a user friendly interface where anybody can create dashboards, share them, and drill down the data.
#
# atoti's UI can be reached with a link using command `session.link()`
# Run the cell below to have a look at a dashboard we have prepared using the above chart and pivot table.
session.link(path="#/dashboard/94c")
# ### Customizing hierarchies
#
# In large organizations, books usually belong to business units that are made up of smaller sub-business units and different trading desks.
# atoti lets you add new hierarchies on the fly without having to add columns into existing tables or re-launch time consuming batch computations.
#
# In this example we will import a file containing level information on Business Units, Sub-Business Units, Trading Desks and Book. Since we already have book IDs linked to our instruments, we will simply use this new information to create an additional hierarchy with these levels under it.
# +
trading_desks = session.read_csv(
"s3://data.atoti.io/notebooks/var/trading_desk.csv",
keys=["book_id"],
table_name="Trading Desk",
)
positions.join(trading_desks)
h["Trading Book Hierarchy"] = {
"Business Unit": lvl["business_unit"],
"Sub Business Unit": lvl["sub_business_unit"],
"Trading Desk": lvl["trading_desk"],
"Book": lvl["book"],
}
# -
# The cube structure has been modified on the fly, we can now use the new hierarchy on any visualization. The data model becomes the following:
cube.schema
# + activeviam={"state": {"name": "Business Hierarchy Pivot Table", "type": "container", "value": {"body": {"configuration": {"mapping": {"splitBy": ["[Hierarchies].[book].[book]", "[Hierarchies].[underlying].[underlying]"], "values": ["[Measures].[PnL]"]}, "type": "plotly-tree-map"}, "query": {"mdx": "SELECT NON EMPTY [Measures].[PnL] ON COLUMNS, NON EMPTY Crossjoin([Hierarchies].[book].[book].Members, [Hierarchies].[underlying].[underlying].Members) ON ROWS FROM (SELECT {[Hierarchies].[underlying].[ALL].[AllMember].[EURAUD=X], [Hierarchies].[underlying].[ALL].[AllMember].[EURCHF=X], [Hierarchies].[underlying].[ALL].[AllMember].[EURCNY=X], [Hierarchies].[underlying].[ALL].[AllMember].[EURJPY=X], [Hierarchies].[underlying].[ALL].[AllMember].[EURSEK=X], [Hierarchies].[underlying].[ALL].[AllMember].[EURUSD=X]} ON COLUMNS FROM [Positions]) CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "serverUrl": "", "updateMode": "once"}}, "containerKey": "chart", "showTitleBar": false, "style": {}}}} atoti={"height": 274, "widget": {"columnWidths": {"[Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]": 133}, "mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[PnL]", "[Measures].[quantity.SUM]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Trading Desk]"]}, "name": "Business Hierarchy Pivot Table", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY {[Measures].[PnL], [Measures].[quantity.SUM]} ON COLUMNS, NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex].[Developed Market]}, [Trading Desk].[Trading Book Hierarchy].[Trading Desk]))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}}
session.visualize("Business Hierarchy Pivot Table")
# -
# ### Value at Risk
#
# We have vectors of the PnLs of every instrument for the last 372 days for each instrument.
# First thing we will do is define a "scaled vector" measure that will multiply those PnLs vectors by the quantities we hold in our positions at instrument level, aggregate it as a sum above.
scaled_pnl_vector = m["quantity.SUM"] * m["pnl_vector.VALUE"]
m["Position Vector"] = atoti.agg.sum(
scaled_pnl_vector, scope=atoti.scope.origin(lvl["instrument_code"])
)
# From [Wikipedia](https://en.wikipedia.org/wiki/Value_at_risk):
# Value at risk (VaR) \[...\] estimates how much a set of investments might lose (with a given probability), given normal market conditions, in a set time period such as a day.
# For a given portfolio, time horizon, and probability $\rho$, the $\rho$ VaR can be defined informally as the maximum possible loss during that time after we exclude all worse outcomes whose combined probability is at most $\rho$.
#
# In our notebook, we will rather use a confidence level that is $1 - \rho$, where $\rho$ is a 5% chance that we will make a loss greater than the maximum possible loss calculated.
# The maximum possible loss will be computed based on the past PnLs that we have per instrument in vectors.
m["Confidence Level"] = 0.95
m["VaR"] = atoti.array.quantile(m["Position Vector"], (1 - m["Confidence Level"]))
# + atoti={"height": 259, "widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[PnL]", "[Measures].[VaR]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Trading Desk]"]}, "name": "", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY {[Measures].[PnL], [Measures].[VaR]} ON COLUMNS, NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex].[Developed Market]}, [Trading Desk].[Trading Book Hierarchy].[Trading Desk]))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# The results above show that with a 95% confidence level, we are sure that the maximum loss would be -488k for Forex.
#
# 95% is an arbitrary value, what if the extreme cases are ten times worse than what we have? Or what if chosing a lower confidence level would tremendously decrease the VaR?
#
# This kind of simulation is pretty easy to put in place with atoti.
# Below we setup a simulation on measure `Confidence level` then define what its value should be in various scenarios.
# +
confidence_levels = cube.create_parameter_simulation(
"Confidence Level",
measure_name="Confidence Level",
default_value=0.95,
base_scenario_name="95%",
)
# Creating scenarios programmatically:
confidence_levels += ("90%", 0.90)
confidence_levels += ("98%", 0.98)
# -
# Once the simulation is setup, we can access its different values using the new `Confidence level` hierarchy that has automatically been created
# + activeviam={"state": {"name": "VAR 98% on SBU", "type": "container", "value": {"body": {"configuration": {"featuredValues": {}}, "contextValues": {}, "mdx": "SELECT NON EMPTY Crossjoin(Hierarchize([Epoch].[Epoch].[Branch].Members), Hierarchize([Confidence Level].[Confidence Level].[Confidence Level].Members), [Hierarchies].[Trading Book Hierarchy].[Trading Desk].Members) ON ROWS, NON EMPTY [Measures].[VaR] ON COLUMNS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "ranges": {"column": {"chunkSize": 20, "thresholdPercentage": 0.1}, "row": {"chunkSize": 20, "thresholdPercentage": 0.1}}, "serverUrl": "", "updateMode": "once"}, "containerKey": "featured-values", "showTitleBar": false, "style": {}}}} atoti={"height": 247, "widget": {"mapping": {"columns": ["ALL_MEASURES", "[Confidence Level].[Confidence Level].[Confidence Level]"], "measures": ["[Measures].[VaR]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]"]}, "name": "VaR per scenario", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY Crossjoin([Measures].[VaR], [Confidence Level].[Confidence Level].[Confidence Level].Members) ON COLUMNS, NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize("VaR per scenario")
# -
# ### Marginal VaR
#
# Since the VaR is not additive – the sum of the VaRs of multiple elements is not equal to the VaR of their parent in a hierarchy – contributory measures are used by Risk Managers to analyze the impact of a Sub-Portfolio on the Value at Risk of the total Portfolio. These measures can help to track down individual positions that have significant effects on VaR. Furthermore, contributory measures can be a useful tool in hypothetical analyses of portfolio development versus VaR development.
#
# One of those measures, the marginal VaR, computes the contribution of one element on the VaR of its parent.
#
# Cells below detail how the marginal VaR is defined with atoti.
m["Parent Position Vector Ex"] = atoti.agg.sum(
m["Position Vector"],
scope=atoti.scope.siblings(h["Trading Book Hierarchy"], exclude_self=True),
)
m["Parent VaR Ex"] = atoti.array.quantile(
m["Parent Position Vector Ex"], (1 - m["Confidence Level"])
)
m["Parent VaR"] = atoti.parent_value(m["VaR"], degrees={h["Trading Book Hierarchy"]: 1})
m["Marginal VaR"] = m["Parent VaR"] - m["Parent VaR Ex"]
# That's it, our marginal VaR is computed, let's have a look at where we could reduce the VaR the most now.
# + atoti={"height": 277, "widget": {"columnWidths": {"[Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]": 133}, "mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[VaR]", "[Measures].[Marginal VaR]", "[Measures].[Parent VaR]", "[Measures].[Parent VaR Ex]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Trading Desk]"]}, "name": "", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY {[Measures].[VaR], [Measures].[Marginal VaR], [Measures].[Parent VaR], [Measures].[Parent VaR Ex]} ON COLUMNS, NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex].[Developed Market]}, [Trading Desk].[Trading Book Hierarchy].[Trading Desk]))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# ## PnL Models Comparison
#
# The VaR calculation is heavily based on the PnL vectors that depend on the results of our instruments pricers, and the history that we have.
# What would happen if pricers used a different model, or if we changed the amount of history we use to compute the VaR.
#
# atoti also lets you perform easy simulations on data tables that were loaded.
# We will load this new file in the analytics table, but in a new scenario called "Model short volatility".
analytics.scenarios["Model short Volatility"].load_csv(
"s3://data.atoti.io/notebooks/var/simulated_pl_vol_depth_270.csv",
array_separator=";",
)
# And that's it, there is no need to re-load any of the previous files, re-define measures or perform batch computations. Everything we have previously defined is available in both our previous and this new scenario.
# Let's have a look at it.
# + atoti={"height": 259, "widget": {"columnWidths": {"[Epoch].[Epoch].[Model short Volatility]": 137, "[Measures].[VaR],[Epoch].[Epoch].[Base - Model short Volatility]": 173, "[Measures].[VaR],[Epoch].[Epoch].[Model short Volatility]": 137, "[Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]": 133}, "mapping": {"columns": ["ALL_MEASURES", "[Epoch].[Epoch].[Branch]"], "measures": ["[Measures].[VaR]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]"]}, "query": {"mdx": "WITH Member [Epoch].[Epoch].[Base - Model short Volatility] AS [Epoch].[Epoch].[Base] - [Epoch].[Epoch].[Model short Volatility], CAPTION = \"Base - Model short Volatility\" SELECT NON EMPTY Hierarchize(Union(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]))) ON ROWS, NON EMPTY Crossjoin({[Measures].[VaR]}, Hierarchize(Union([Epoch].[Epoch].[Branch].Members, [Epoch].[Epoch].[Base - Model short Volatility]))) ON COLUMNS FROM [Positions]", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}}
session.visualize("Pivot table comparison Model Short Volatility")
# -
# ## Combined Scenarios
#
# We may also combine scenarios together and answer questions such as "What would be the VaR and Marginal VaR for the Short Volatility model combined with the 95% and 98% confidence level scenarios?"
# + atoti={"widget": {"columnWidths": {"[Measures].[VaR],[Confidence Level].[Confidence Level].[95%],[Epoch].[Epoch].[Model short Volatility]": 137, "[Measures].[VaR],[Confidence Level].[Confidence Level].[98%],[Epoch].[Epoch].[Model short Volatility]": 137, "[Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]": 133, "[Trading Desk].[Trading Book Hierarchy].[Trading Desk]": 92}, "filters": ["{[Confidence Level].[Confidence Level].[98%], [Confidence Level].[Confidence Level].[95%]}"], "mapping": {"columns": ["ALL_MEASURES", "[Confidence Level].[Confidence Level].[Confidence Level]", "[Epoch].[Epoch].[Branch]"], "measures": ["[Measures].[VaR]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Trading Desk]"]}, "name": "Combined Scenarios", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY Crossjoin({[Measures].[VaR]}, [Confidence Level].[Confidence Level].[Confidence Level].Members, [Epoch].[Epoch].[Branch].Members) ON COLUMNS, NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex].[Developed Market]}, [Trading Desk].[Trading Book Hierarchy].[Trading Desk]))) ON ROWS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize("Combined Scenarios")
# -
# # LEstimated VaR
#
# The LEstimated VaR is a contributory measure. It is an additive measure such that the LEstimated VaRs of all Sub-Portfolios add up to the VaR of the parent Portfolio.
#
# The LEstimated VaR shows the simulated PL for the tail scenario, that has been identified as the VaR scenario for the parent Portfolio.
# +
# Compute the rank of the VaR scenario
vectorSize = atoti.array.len(m["Position Vector"])
m["VaR Rank Current Portfolio"] = atoti.math.floor(
(1 - m["Confidence Level"]) * vectorSize
)
# Pick the id of the scenario at the rank
m["Tail Indices"] = atoti.array.n_lowest_indices(
m["Position Vector"], m["VaR Rank Current Portfolio"]
)
m["VaR Scenario Id"] = m["Tail Indices"][m["VaR Rank Current Portfolio"] - 1]
m["VaR Value"] = m["Position Vector"][m["VaR Scenario Id"]]
# Create a measure to access the parent's level Id
m["VaR Scenario Id Parent"] = atoti.parent_value(
m["VaR Scenario Id"], degrees={h["Trading Book Hierarchy"]: 1}
)
# Finally, the LEstimated VaR measure
m["LEstimated VaR"] = m["Position Vector"][m["VaR Scenario Id Parent"]]
# + atoti={"height": 278, "widget": {"columnWidths": {"[Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]": 133, "[Trading Desk].[Trading Book Hierarchy].[Trading Desk]": 92}, "mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[LEstimated VaR]", "[Measures].[VaR Value]", "[Measures].[VaR Scenario Id]", "[Measures].[VaR Scenario Id Parent]"], "rows": ["[Trading Desk].[Trading Book Hierarchy].[Business Unit] => [Trading Desk].[Trading Book Hierarchy].[Trading Desk]"]}, "name": "", "query": {"context": {"queriesResultLimit.intermediateSize": 1000000, "queriesResultLimit.transientSize": 10000000}, "mdx": "SELECT NON EMPTY Hierarchize(Union(Hierarchize(DrilldownLevel([Trading Desk].[Trading Book Hierarchy].[ALL].[AllMember])), Hierarchize(Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember]}, 1, SELF_AND_BEFORE)), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex]}, [Trading Desk].[Trading Book Hierarchy].[Sub Business Unit]), Descendants({[Trading Desk].[Trading Book Hierarchy].[AllMember].[Forex].[Developed Market]}, [Trading Desk].[Trading Book Hierarchy].[Trading Desk]))) ON ROWS, NON EMPTY {[Measures].[LEstimated VaR], [Measures].[VaR Value], [Measures].[VaR Scenario Id], [Measures].[VaR Scenario Id Parent]} ON COLUMNS FROM [Positions] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
session.link()
#
# <div style="text-align:center"><a href="https://www.atoti.io/?utm_source=gallery&utm_content=value-at-risk" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover-try.png" alt="atoti" /></a></div>
| notebooks/value-at-risk/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import scipy.special as sp
import math as ma
import numpy as np
import scipy.stats as st
import numba as nb
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
from scipy.optimize import minimize
import bayessplicedmodels as bsm
from joblib import Parallel, delayed
# # Burr distribution
#
# $X\sim\text{Burr}(\alpha, \beta, \sigma)$ with pdf
#
# $$
# f(x) = \frac{\alpha\beta\sigma^{\alpha\beta}x^{\beta-1}}{(\sigma^\beta +x^\beta)^{\alpha+1}}
# $$
class loss_model:
def __init__(self,name, parm_names):
self.name = name
self.parm_names = parm_names
self.d= len(parm_names)
def set_logp(self, X):
if self.name == "Burr":
def logp(parms):
α, β, σ = parms
if np.all(parms) > 0:
return(len(X)*(np.log(α) + np.log(β) + α * β * np.log(σ)) + \
(β - 1) * np.sum(np.log(X)) - (α + 1) * np.sum(np.log(σ**β + X**β)) )
else:
return(-np.inf)
self.logp = nb.jit(nopython = True)(logp)
def set_logps(self):
if self.name == "Burr":
def logp_body(X, parms, γ):
α, β, σ = parms
F1 = 1 - (1 + (γ/σ)**β)**(-α)
if np.all(parms > 0) and F1 > 0:
return(len(X)*(np.log(α) + np.log(β) + α * β * np.log(σ)) + \
(β - 1) * np.sum(np.log(X)) - (α + 1) * np.sum(np.log(σ**β + X**β)) \
- len(X) * np.log(F1))
else:
return(-np.inf)
def logp_tail(X, parms, γ):
α, β, σ = parms
F2 = 1 - (1 + (γ/σ)**β)**(-α)
if np.all(parms > 0) and F2 < 1:
return(len(X)*(np.log(α) + np.log(β) + α * β * np.log(σ)) + \
(β - 1) * np.sum(np.log(X)) - (α + 1) * np.sum(np.log(σ**β + X**β)) \
- len(X) * np.log(1 - F2))
else:
return(-np.inf)
self.logp_body = nb.jit(nopython = True)(logp_body)
self.logp_tail = nb.jit(nopython = True)(logp_tail)
def set_logd(self, parms):
if self.name == "Burr":
def logd(x):
α, β, σ = parms[:,0], parms[:,1], parms[:,2]
res = np.zeros(len(β))
s = np.logical_and(α >0, np.logical_and(β > 0, σ > 0))
res[np.where(s)] = np.log(α[s]) + np.log(β[s]) + α[s] * β[s] * np.log(σ[s]) +\
(β[s] - 1) * np.log(x) - (α[s] + 1) * np.log(σ[s]**β[s] + x**β[s])
res[np.where(np.invert(s))] = -np.inf
return(res)
self.logd = logd
def set_logds(self):
if self.name == "Burr":
def logd_body(x, parms, γ):
α, β, σ = parms[:,0], parms[:,1], parms[:,2]
F1 = 1 - (1 + (γ/σ)**β)**(-α)
res = np.zeros(len(β))
s = np.logical_and(np.logical_and(α >0, np.logical_and(β > 0, σ > 0)), x < γ)
res[np.where(s)] = np.log(α[s]) + np.log(β[s]) + α[s] * β[s] * np.log(σ[s]) +\
(β[s] - 1) * np.log(x) - (α[s] + 1) * np.log(σ[s]**β[s] + x**β[s]) - np.log(F1[s])
res[np.where(np.invert(s))] = -np.inf
return(res)
def logd_tail(x, parms, γ):
α, β, σ = parms[:,0], parms[:,1], parms[:,2]
F2 = 1 - (1 + (γ/σ)**β)**(-α)
res = np.zeros(len(β))
s = np.logical_and(np.logical_and(α >0, np.logical_and(β > 0, σ > 0)), x > γ)
res[np.where(s)] = np.log(α[s]) + np.log(β[s]) + α[s] * β[s] * np.log(σ[s]) +\
(β[s] - 1) * np.log(x) - (α[s] + 1) * np.log(σ[s]**β[s] + x**β[s]) - np.log(1 - F2[s])
res[np.where(np.invert(s))] = -np.inf
return(res)
self.logd_body = logd_body
self.logd_tail = logd_tail
def set_cdf(self):
if self.name == "Burr":
def cdf(parms, x):
α, β, σ = parms
return(1 - (1 + (x / σ)**β)**(-α))
self.cdf = nb.jit(nopython = True)(cdf)
def set_pdf(self):
if self.name == "Burr":
def pdf(parms, x):
α, β, σ = parms
return(α * β * σ**(α * β) * x**(β - 1) / (σ**β + x**β)**(α + 1))
self.pdf = nb.jit(nopython = True)(pdf)
def set_ppf(self):
if self.name == "Burr":
def ppf(parms, y):
α, β, σ = parms
return( σ * ( (1-y)**(-1 / α) - 1)**(1 / β))
self.ppf = ppf
def sample(self, parms, n):
if self.name == "Burr":
α, β, σ = parms
return(st.burr12( β, α).rvs(size = n) * σ)
burr_dist = loss_model("Burr", ["α", "β", "σ"])
print(burr_dist.name, burr_dist.parm_names, burr_dist.d)
parms = np.array([2, 2, 1])
α, β, σ = parms
x, y = 2, 0.5
burr_dist.set_cdf(), burr_dist.set_pdf(), burr_dist.set_ppf()
burr_dist.cdf(parms, x) - st.burr12( β, α).cdf(x / σ),\
burr_dist.ppf(parms, y)- st.burr12(β, α).ppf(y) * σ,\
burr_dist.pdf(parms, x)- st.burr12(β, α).pdf(x / σ) / σ
X, γ = st.burr12( β, α).rvs(size = 100) * σ, 2
burr_dist.set_logps(), burr_dist.set_logp(X)
print(burr_dist.logp(parms) - np.sum(np.log(st.burr12( β, α).pdf(X / σ) / σ)))
print(burr_dist.logp_body(X, parms, γ) - np.sum(np.log(st.burr12( β, α).pdf(X / σ) / σ / st.burr12( β, α).cdf(γ / σ))))
print(burr_dist.logp_tail(X, parms, γ)- np.sum(np.log(st.burr12( β, α).pdf(X / σ) / σ / (1 - st.burr12( β, α).cdf(γ / σ)))))
X = st.burr12( β, α).rvs(size = 10) * σ
α_prior, β_prior, σ_prior, γ_prior= bsm.prior_model('gamma','α', 1, 1), bsm.prior_model('gamma','β', 1, 1), bsm.prior_model('gamma','σ', 1, 1), bsm.prior_model('gamma','γ', 1, 1)
prior_gamma_model = bsm.independent_priors([α_prior, β_prior, σ_prior, γ_prior])
particle_cloud = prior_gamma_model.sample(20)
burr_dist.set_logds(), burr_dist.set_logd(particle_cloud.values)
α_vec, β_vec, σ_vec, γ_vec = particle_cloud.values[:,0], particle_cloud.values[:,1], \
particle_cloud.values[:,2], particle_cloud.values[:,3]
print(np.array([np.log(st.burr12(β_vec[i], α_vec[i]).pdf(X[1] / σ_vec[i]) / σ_vec[i]) for i in range(len(γ_vec))] -
burr_dist.logd(X[1])))
print(burr_dist.logd_body(X[0], particle_cloud.values, particle_cloud.values[:,-1]) -
np.array([np.sum(np.log(st.burr12(β_vec[i], α_vec[i]).pdf(X[0] / σ_vec[i]) / σ_vec[i] / st.burr12(β_vec[i], α_vec[i]).cdf(γ_vec[i] / σ_vec[i])))
for i in range(len(γ_vec)) ]) )
print(burr_dist.logd_tail(X[0], particle_cloud.values, particle_cloud.values[:,-1]) -
np.array([np.sum(np.log(st.burr12(β_vec[i], α_vec[i]).pdf(X[0] / σ_vec[i]) / σ_vec[i] / (1-st.burr12(β_vec[i], α_vec[i]).cdf(γ_vec[i] / σ_vec[i]))))
for i in range(len(γ_vec))]))
parms_true = np.array([2, 3, 1])
f = loss_model("Burr", ["α", "β", "σ"])
# X= st.burr12(parms_true[1], parms_true[0]).rvs(size = 500) * parms_true[2]
danish = pd.read_csv("Data/danish.csv").x
X = danish.values
plt.hist(X,bins=100)
sns.despine()
α_prior, β_prior, σ_prior = bsm.prior_model('gamma','α', 1, 1), bsm.prior_model('gamma','β', 1, 1), bsm.prior_model('gamma','σ', 1, 1)
prior_single_model = bsm.independent_priors([α_prior, β_prior, σ_prior])
popSize, ρ, c, n_step_max, err, paralell, n_proc, verbose = 2000, 1/2, 0.99, 25, 1e-6, False, 4, True
# %time trace, log_marg, DIC, WAIC = bsm.smc_likelihood_annealing(X, f, popSize, prior_single_model, ρ, c,n_step_max, err, paralell, 4, verbose)
# +
f.set_ppf()
print(log_marg, DIC, WAIC, bsm.compute_Wasserstein(X, f, trace.mean().values, 1))
bsm.posterior_plots(f, trace)
bsm.trace_plots(f, trace)
bsm.qq_plot(X, f, trace.mean().values)
# -
import bayessplicedmodels as bsm
parms_true = np.array([3, 1.5, 1.2, 1, 2, 5, 0.9])
f1, f2 = bsm.loss_model("Weibull", ["μ1", "λ1"]), bsm.loss_model("Burr", ["α2", "β2", "σ2"])
f = bsm.spliced_loss_model(f1 , f2, "continuous")
# X= f.sample(parms_true, 1000)
danish = pd.read_csv("Data/danish.csv").x
X = danish.values
# α1_prior, β1_prior, σ1_prior = bsm.prior_model('gamma','α1', 1, 1), bsm.prior_model('gamma','β1', 1, 1), bsm.prior_model('gamma','σ1', 1, 1)
μ1_prior, λ1_prior = bsm.prior_model('gamma','μ1', 1, 1), bsm.prior_model('gamma','λ1', 1, 1)
α2_prior, β2_prior, σ2_prior = bsm.prior_model('gamma','α2', 1, 1), bsm.prior_model('gamma','β2',1, 1), bsm.prior_model('gamma','σ2', 1, 1)
γ_prior, p_prior = bsm.prior_model('uniform','γ',min(X), max(X)), bsm.prior_model('uniform', 'p', 0, 1)
prior_spliced_model = bsm.independent_priors([μ1_prior, λ1_prior, α2_prior, β2_prior, σ2_prior, γ_prior])
plt.hist(X,bins=200)
sns.despine()
popSize, ρ, c, n_step_max, err, paralell, n_proc, verbose = 10000, 1/2, 0.99, 25, 1e-6, True, 4, True
# %time trace, log_marg, DIC, WAIC = bsm.smc_likelihood_annealing(X, f, popSize, prior_spliced_model, ρ, c,n_step_max, err, paralell, 4, verbose)
# +
f.set_ppf()
print(log_marg, DIC, WAIC, bsm.compute_Wasserstein(X, f, trace.mean().values, 1))
print(trace.mean())
bsm.posterior_plots(f, trace)
bsm.trace_plots(f, trace)
bsm.qq_plot(X, f, trace.mean().values)
# -
# # On the danish fire insurance data set
# +
# The data
danish = pd.read_csv("Data/danish.csv").x
X = danish.values
# Model for the bulk distribution
body_model_names = ["Exp", "Gamma", "Weibull", "Inverse-Gaussian", "Lognormal"]
body_model_param_names = [['λ1'], ["r1", "m1"], ["k1", "β1"], ["μ1", "λ1"], ["μ1", "σ1"]]
# Prior distributions over the parameters of the bulk distribution
body_model_priors= [[bsm.prior_model('gamma',body_model_param_names[0][0], 1, 1)],
[bsm.prior_model('gamma',body_model_param_names[1][0], 1, 1), bsm.prior_model('gamma',body_model_param_names[1][1], 1, 1)],
[bsm.prior_model('gamma',body_model_param_names[2][0], 1, 1), bsm.prior_model('gamma',body_model_param_names[2][1], 1, 1)],
[bsm.prior_model('gamma',body_model_param_names[3][0], 1, 1), bsm.prior_model('gamma',body_model_param_names[3][1], 1, 1)],
[bsm.prior_model('normal',body_model_param_names[4][0], 0, 0.5), bsm.prior_model('gamma',body_model_param_names[4][1], 1, 1)]
]
# Model for the tail of the distribution
tail_model_names = ["Burr"]
tail_model_param_names = [["α2", "β2", "σ2"]]
# Prior distributions over the parameters of the bulk distribution
tail_model_priors= [
[bsm.prior_model('gamma',tail_model_param_names[0][0], 1, 1), bsm.prior_model('gamma',tail_model_param_names[0][1], 1, 1), bsm.prior_model('gamma',tail_model_param_names[0][2], 1, 1)]]
γ_prior, p_prior = bsm.prior_model('uniform', "γ", min(X), max(X)), bsm.prior_model('uniform',"p", 0, 1)
#Splicing model type
splicing_types = ["continuous"]
# Setting the models
fs, f_names, prior_spliced_model = [], [], []
for i in range(len(body_model_names)):
for j in range(len(tail_model_names)):
for splicing_type in splicing_types:
f1, f2 = bsm.loss_model(body_model_names[i], body_model_param_names[i]), bsm.loss_model(tail_model_names[j], tail_model_param_names[j])
fs.append(bsm.spliced_loss_model(f1 , f2, splicing_type))
f_names.append(body_model_names[i] +"-"+ tail_model_names[j]+"-"+splicing_type)
if splicing_type == "disjoint":
prior_spliced_model.append(bsm.independent_priors(body_model_priors[i] + tail_model_priors[j] + [γ_prior, p_prior]))
else:
prior_spliced_model.append(bsm.independent_priors(body_model_priors[i] + tail_model_priors[j] + [γ_prior]))
for f in fs:
f.set_ppf()
fs_dict = dict(zip(f_names, fs))
# -
popSize, ρ, c, n_step_max, err, paralell, n_proc, verbose = 4000, 1/2, 0.99, 25, 1e-6, False, 4, False
def fit_spliced_models(i):
trace, log_marg, DIC, WAIC = bsm.smc_likelihood_annealing(X, fs[i], popSize, prior_spliced_model[i], ρ, c,n_step_max, err, paralell, 4, verbose)
return([trace, log_marg, DIC, WAIC])
# %time res = Parallel(n_jobs=4)(delayed(fit_spliced_models)(i) for i in range(len(f_names)))
# +
fit_spliced_models_dic = dict(zip(f_names, res))
γ_map = np.array([fit_spliced_models_dic[f_names[k]][0]['γ'].mean() for k in range(len(fit_spliced_models_dic))])
spliced_model_df = pd.DataFrame({'model':f_names,
"d": np.array([f.d for f in fs]),
"γ_map": np.array([fit_spliced_models_dic[f_names[k]][0]['γ'].mean() for k in range(len(fit_spliced_models_dic))]),
'log_marg': np.array([fit_spliced_models_dic[f_names[k]][1] for k in range(len(fit_spliced_models_dic))]),
"DIC": np.array([fit_spliced_models_dic[f_names[k]][2] for k in range(len(fit_spliced_models_dic))]),
"WAIC":np.array([fit_spliced_models_dic[f_names[k]][3] for k in range(len(fit_spliced_models_dic))])})
spliced_model_df["posterior_probability"] = np.exp(spliced_model_df["log_marg"] - np.max(spliced_model_df["log_marg"])) / np.sum(np.exp(spliced_model_df["log_marg"] - np.max(spliced_model_df["log_marg"])))
spliced_model_df["Wass_dist"] = np.array([bsm.compute_Wasserstein(X, fs_dict[model_name], fit_spliced_models_dic[model_name][0].mean().values, 1) for model_name in spliced_model_df["model"].values])
spliced_model_df.sort_values(by='DIC', ascending=False)
# -
model_names = spliced_model_df.sort_values(by='log_marg', ascending=False)["model"]
for model_name in model_names:
f, trace = fs_dict[model_name], fit_spliced_models_dic[model_name][0]
# print(trace.mean().values)
bsm.posterior_plots(f, trace)
bsm.trace_plots(f, trace)
bsm.qq_plot(X, f, trace.mean().values)
| tests/distributions/burr_dist.ipynb |