code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import plotly
import plotly.graph_objs as go
from IPython.display import display, HTML
plotly.offline.init_notebook_mode(connected=True)
from loadbalanceRL import RAINMAN3
from loadbalanceRL.lib.algorithm.plot_dynamic import *
#
# #### Normal case: Start with 30% Random actions (explore) and rest 70% fetch max actions (exploit)
#
# +
# Server profile: num_ues=120, APs=16, Scale=200.0, explore_radius=1
# If you want to change the num of ues or aps go to reset that in the server.py
# the EPISODES shows here is 100
from collections import OrderedDict
ALGORITHM_CONFIG = OrderedDict(
EPISODES=3,
ALPHA=0.2,
GAMMA=0.9,
EPSILON=0.3,
LAERNING_RATE=0.005,
EPSILON_DECAY=0.99,
EPSILON_MIN=0.01,
VERBOSE=True,
L1_HIDDEN_UNITS=13,
L2_HIDDEN_UNITS=13,
L1_ACTIVATION='relu',
L2_ACTIVATION='relu',
LOSS_FUNCTION='mean_squared_error',
OPTIMIZER='Adam',
)
CELLULAR_MODEL_CONFIG = OrderedDict(
NAME='Cellular',
TYPE='Dev',
SERVER='0.0.0.0',
SERVER_PORT='8000',
VERBOSE=True,
)
# -
RAINMAN3.algorithm_config = ALGORITHM_CONFIG
RAINMAN3.environment_config = CELLULAR_MODEL_CONFIG
result = RAINMAN3.run_experiment("Cellular", "Qlearning", "Naive")
print("Number of states encountered: {}".format(len(result.Q)))
print("Number of q_ap_states encountered: {}".format(len(result.Q_ap)))
print(result.BR_LIST)
# +
# data_values=data["data"]
# print(data_values)
# Including the BR dict
br_list = result.BR_LIST
BR_locations=[]
BR_locations += [x.location for x in br_list]
Rec_dict=[{
'type': 'rect',
'x0': BR_locations[0][0],
'y0': BR_locations[0][2],
'x1': BR_locations[0][1],
'y1': BR_locations[0][3],
'line': {
'color': 'rgba(128, 0, 128, 1)',
'width': 2,
},
'fillcolor': 'rgba(128, 0, 128 0.1)'
}]
Rec_dict.append({
'type': 'rect',
'x0': BR_locations[1][0],
'y0': BR_locations[1][2],
'x1': BR_locations[1][1],
'y1': BR_locations[1][3],
'line': {
'color': 'rgba(128, 0, 128, 1)',
'width': 2,
},
'fillcolor': 'rgba(128, 0, 128 0.1)'
})
Rec_dict.append({
'type': 'rect',
'x0': BR_locations[2][0],
'y0': BR_locations[2][2],
'x1': BR_locations[2][1],
'y1': BR_locations[2][3],
'line': {
'color': 'rgba(128, 0, 128, 1)',
'width': 2,
},
'fillcolor': 'rgba(128, 0, 128 0.1)'
})
Rec_dict.append({
'type': 'rect',
'x0': BR_locations[3][0],
'y0': BR_locations[3][2],
'x1': BR_locations[3][1],
'y1': BR_locations[3][3],
'line': {
'color': 'rgba(128, 0, 128, 1)',
'width': 2,
},
'fillcolor': 'rgba(128, 0, 128 0.1)'
})
# -
# data represents as the fist frame data for plot
data, Frames_data, last_episode, layout_dict = plot_dynamic_data(result, Rec_dict, ALGORITHM_CONFIG['EPISODES'])
# +
data_values=data["data"]
figure={"data":data_values,
"layout": {'xaxis':{'range': [0, 1600], 'title': 'x'},
'yaxis':{'range': [0, 1600], 'title': 'y'},
'title':"UE and AP Distribution in the grid",
'height' : 1000,
'width' :1000,
'showlegend': False,
"shapes":Rec_dict,
'updatemenus': [{'type': 'buttons',
'buttons': [{'label': 'Play',
'method': 'animate',
'args': [None]}]}]
},
'frames':Frames_data}
# print(figure)
plotly.offline.iplot(figure)
# +
# Create rewards graph
rewards = go.Scatter(
x = len(result.Rewards),
y = result.Rewards,
name='Rewards'
)
data = [rewards]
# Plot and embed in ipython notebook!
plotly.offline.iplot(
{"data": data,
"layout": go.Layout(
height=500,
width=800,
title="Rewards over time with UE number of 200",
xaxis=dict(
title='Episodes',
),
yaxis=dict(
title='Rewards',
),
showlegend=True)},
show_link=False,
link_text=None)
# +
# Create handoff graph
handoffs = go.Scatter(
x = len(result.Handoffs),
y = result.Handoffs,
name='Handoffs'
)
data = [handoffs]
# Plot and embed in ipython notebook!
plotly.offline.iplot(
{"data": data,
"layout": go.Layout(
height=500,
width=800,
title='Handoffs over time with UE number of 200',
xaxis=dict(
title='Episodes',
),
yaxis=dict(
title='Handoffs'
),
showlegend=True)},
show_link=False,
link_text=None)
# +
# Create UE graph
ue_sla = go.Scatter(
x = len(result.Meets_SLA),
y = result.Meets_SLA,
name='UE SLA'
)
data = [ue_sla]
# Plot and embed in ipython notebook!
plotly.offline.iplot(
{"data": data,
"layout": go.Layout(
height=500,
width=800,
title="UE's SLA over time with UE number of 200",
xaxis=dict(
title='Episodes',
),
yaxis=dict(
title='UE SLA'
),
showlegend=True)},
show_link=False,
link_text=None)
# -
| loadbalanceRL/notebooks/test_cellular_env.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: autoreview_venv
# language: python
# name: autoreview_venv
# ---
import sys, os, time, pickle
from timeit import default_timer as timer
from humanfriendly import format_timespan
import pandas as pd
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction import DictVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer, TfidfTransformer, CountVectorizer
from sklearn.metrics import classification_report
from dotenv import load_dotenv
load_dotenv('admin.env')
from db_connect_mag import Session, Paper, PaperAuthorAffiliation, db
# +
# test_papers_df = pd.read_pickle('data/collect_haystack_20180409/test_papers.pickle')
# target_papers_df = pd.read_pickle('data/collect_haystack_20180409/target_papers.pickle')
# train_papers_df = pd.read_pickle('data/collect_haystack_20180409/train_papers.pickle')
# -
# this is the data for the fortunato review on Community Detection in Graphs
start = timer()
test_papers_df = pd.read_pickle('data/collect_haystack_20180409_2/test_papers.pickle')
target_papers_df = pd.read_pickle('data/collect_haystack_20180409_2/target_papers.pickle')
train_papers_df = pd.read_pickle('data/collect_haystack_20180409_2/train_papers.pickle')
print(format_timespan(timer()-start))
with open('data/collect_haystack_20180409_2/counter.pickle', 'rb') as f:
c = pickle.load(f)
def get_target_in_test(test, target, id_colname='Paper_ID'):
return set.intersection(set(test[id_colname]), set(target[id_colname]))
len(get_target_in_test(test_papers_df, target_papers_df))
len(target_papers_df)
test_subset = test_papers_df.sample(n=100000)
len(get_target_in_test(test_subset, target_papers_df))
# remove the train (seed) papers from the test set (haystack)
n_before = len(test_subset)
test_subset = test_subset.drop(train_papers_df.index, errors='ignore')
n_after = len(test_subset)
print("removed {} seed papers from the haystack. size of haystack: {}".format(n_before-n_after, n_after))
start = timer()
target_ids = set(target_papers_df.Paper_ID)
test_papers_df['target'] = test_subset.Paper_ID.apply(lambda x: x in target_ids)
print(format_timespan(timer()-start))
# +
# def tree_distance(n1, n2, sep=":"):
# # https://en.wikipedia.org/wiki/Lowest_common_ancestor
# # the distance from v to w can be computed as
# # the distance from the root to v, plus the distance from
# # the root to w, minus twice the distance from
# # the root to their lowest common ancestor
# v, w = [n.split(sep) for n in [n1, n2]]
# distance_root_to_v = len(v)
# distance_root_to_w = len(w)
# distance_root_to_lca = 0
# for i in range(min(distance_root_to_v, distance_root_to_w)):
# if v[i] == w[i]:
# distance_root_to_lca += 1
# else:
# break
# return distance_root_to_v + distance_root_to_w - (2*distance_root_to_lca)
# -
def tree_distance(n1, n2, sep=":"):
# since depth is sort of arbitrary, let's try this
v, w = [n.split(sep) for n in [n1, n2]]
distance_root_to_v = len(v)
distance_root_to_w = len(w)
avg_depth = (distance_root_to_v + distance_root_to_w) * .5
distance_root_to_lca = 0
for i in range(min(distance_root_to_v, distance_root_to_w)):
if v[i] == w[i]:
distance_root_to_lca += 1
else:
break
return (avg_depth - distance_root_to_lca) / avg_depth
def avg_distance(cl, cl_group):
distances = []
for x in cl_group:
distances.append(tree_distance(cl, x))
return sum(distances) / len(distances)
n_before = len(test_subset)
test_subset = test_subset.dropna(subset=['title'])
n_after = len(test_subset)
print("dropped {} rows".format(n_before-n_after))
test_subset = test_subset.reset_index()
# +
start = timer()
vect = CountVectorizer()
data = test_subset.title.append(train_papers_df.title).tolist()
vect.fit(data)
print(format_timespan(timer()-start))
# -
start = timer()
tf_train = vect.transform(train_papers_df.title.tolist())
print(format_timespan(timer()-start))
start = timer()
tf_test = vect.transform(test_subset.title.tolist())
print(format_timespan(timer()-start))
start = timer()
tf_global = vect.transform(data)
print(format_timespan(timer()-start))
tf_transform = TfidfTransformer()
tf_transform.fit(tf_global)
tfidf_train = tf_transform.transform(tf_train)
tfidf_test = tf_transform.transform(tf_test)
tfidf_test.shape
from sklearn.metrics.pairwise import cosine_similarity
csims = cosine_similarity(tfidf_test, tfidf_train.mean(axis=0))
test_subset = test_subset.join(pd.Series(csims.flatten(), name='title_tfidf_cosine_similarity'))
test_subset.sort_values('title_tfidf_cosine_similarity', ascending=False)
start = timer()
test_papers_df['avg_distance_to_train'] = test_papers_df.cl.apply(avg_distance, cl_group=train_papers_df.cl.tolist())
print(format_timespan(timer()-start))
test_papers_df.sort_values(['avg_distance_to_train', 'EF'], ascending=[True, False]).head(50)
test_papers_df.groupby('target')['EF', 'avg_distance_to_train'].describe().T
import matplotlib.pyplot as plt
# %matplotlib inline
# http://scikit-learn.org/stable/auto_examples/hetero_feature_union.html
class ItemSelector(BaseEstimator, TransformerMixin):
"""For data grouped by feature, select subset of data at a provided key.
The data is expected to be stored in a 2D data structure, where the first
index is over features and the second is over samples. i.e.
>> len(data[key]) == n_samples
Please note that this is the opposite convention to scikit-learn feature
matrixes (where the first index corresponds to sample).
ItemSelector only requires that the collection implement getitem
(data[key]). Examples include: a dict of lists, 2D numpy array, Pandas
DataFrame, numpy record array, etc.
>> data = {'a': [1, 5, 2, 5, 2, 8],
'b': [9, 4, 1, 4, 1, 3]}
>> ds = ItemSelector(key='a')
>> data['a'] == ds.transform(data)
ItemSelector is not designed to handle data grouped by sample. (e.g. a
list of dicts). If your data is structured this way, consider a
transformer along the lines of `sklearn.feature_extraction.DictVectorizer`.
Parameters
----------
key : hashable, required
The key corresponding to the desired value in a mappable.
"""
def __init__(self, key):
self.key = key
def fit(self, x, y=None):
return self
def transform(self, data_dict):
return data_dict[self.key]
class ClusterTransformer(BaseEstimator, TransformerMixin):
def __init__(self, colname='cl'):
self.colname = colname
def fit(self, x, y=None):
return self
def transform(self, df):
avg_dist = df[self.colname].apply(avg_distance, cl_group=train_papers_df.cl.tolist())
return avg_dist.as_matrix().reshape(-1, 1)
class DataFrameColumnTransformer(BaseEstimator, TransformerMixin):
def __init__(self, colname):
self.colname = colname
def fit(self, x, y=None):
return self
def transform(self, df):
return df[self.colname].as_matrix().reshape(-1, 1)
pipeline = Pipeline([
('union', FeatureUnion(
transformer_list = [
('avg_distance_to_train', Pipeline([
# ('selector', ItemSelector(key='avg_distance_to_train')),
# ('vect', DictVectorizer(X.avg_distance_to_train.to_dict))
('cl_feat', ClusterTransformer()),
])),
('ef', Pipeline([
# ('selector', ItemSelector(key='avg_distance_to_train')),
# ('vect', DictVectorizer(X.avg_distance_to_train.to_dict))
('ef_feat', DataFrameColumnTransformer('EF')),
])),
# NOTE: this is just to test.
# we probably want features that relate the titles to the seed papers. not just straight features in test set.
('title_bow', Pipeline([
('selector', ItemSelector(key='title')),
('tfidf', TfidfVectorizer(min_df=10)),
]))
],
)),
('svc', SVC(kernel='linear', probability=True))
])
# +
# X = test_papers_df[['EF', 'avg_distance_to_train']]
X = test_papers_df[test_papers_df.title.notnull()]
# Fortunato paper was published in 2010
X = X[X.year<=2010]
# y = test_papers_df['target']
y = X['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=999)
# -
start = timer()
pipeline.fit(X_train, y_train)
print(format_timespan(timer()-start))
start = timer()
# y_pred_proba = model.predict_proba(X_test)[:, 1]
y_pred_proba = pipeline.predict_proba(X)[:, 1]
print(format_timespan(timer()-start))
y_pred_proba
y_pred_proba.shape
pred_ranks = pd.Series(y_pred_proba, index=X.index, name='pred_ranks')
test_papers_df.join(pred_ranks).sort_values('pred_ranks', ascending=False).head()
len(test_papers_df)
len(X)
# top_predictions = test_papers_df.join(pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
top_predictions = X.join(pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
top_predictions.groupby('target')['Paper_ID'].count()
top_predictions.pred_ranks.min()
start = timer()
y_test_pred = pipeline.predict(X_test)
print(format_timespan(timer()-start))
print(classification_report(y_test, y_test_pred))
pipeline = Pipeline([
('union', FeatureUnion(
transformer_list = [
('avg_distance_to_train', Pipeline([
# ('selector', ItemSelector(key='avg_distance_to_train')),
# ('vect', DictVectorizer(X.avg_distance_to_train.to_dict))
('cl_feat', ClusterTransformer()),
])),
('ef', Pipeline([
# ('selector', ItemSelector(key='avg_distance_to_train')),
# ('vect', DictVectorizer(X.avg_distance_to_train.to_dict))
('ef_feat', DataFrameColumnTransformer('EF')),
])),
# NOTE: this is just to test.
# we probably want features that relate the titles to the seed papers. not just straight features in test set.
# ('title_bow', Pipeline([
# ('selector', ItemSelector(key='title')),
# ('tfidf', TfidfVectorizer(min_df=10)),
# ]))
],
)),
('logreg', LogisticRegression())
])
# +
# X = test_papers_df[['EF', 'avg_distance_to_train']]
X = test_papers_df[test_papers_df.title.notnull()]
# Fortunato paper was published in 2010
X = X[X.year<=2010]
# y = test_papers_df['target']
y = X['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=999)
# -
start = timer()
pipeline.fit(X_train, y_train)
print(format_timespan(timer()-start))
start = timer()
# y_pred_proba = model.predict_proba(X_test)[:, 1]
y_pred_proba = pipeline.predict_proba(X)[:, 1]
print(format_timespan(timer()-start))
y_pred_proba
y_pred_proba.shape
pred_ranks = pd.Series(y_pred_proba, index=X.index, name='pred_ranks')
test_papers_df.join(pred_ranks).sort_values('pred_ranks', ascending=False).head()
len(test_papers_df)
len(X)
# top_predictions = test_papers_df.join(pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
top_predictions = X.join(pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
top_predictions.groupby('target')['Paper_ID'].count()
top_predictions.pred_ranks.min()
start = timer()
y_test_pred = pipeline.predict(X_test)
print(format_timespan(timer()-start))
print(classification_report(y_test, y_test_pred))
# +
# what if we only use pagerank?
X = test_papers_df[['EF']]
y = test_papers_df['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=999)
start = timer()
_model = LogisticRegression()
_model.fit(X_train, y_train)
print(format_timespan(timer()-start))
# y_pred_proba = model.predict_proba(X_test)[:, 1]
_y_pred_proba = _model.predict_proba(X)[:, 1]
#y_pred_proba
print(y_pred_proba.shape)
_pred_ranks = pd.Series(_y_pred_proba, index=X.index, name='pred_ranks')
#test_papers_df.join(_pred_ranks).sort_values('pred_ranks', ascending=False).head()
_top_predictions = test_papers_df.join(_pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
_top_predictions.groupby('target')['Paper_ID'].count()
# -
# +
# what if we only use avg distance?
X = test_papers_df[['avg_distance_to_train']]
y = test_papers_df['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=999)
start = timer()
_model = LogisticRegression()
_model.fit(X_train, y_train)
print(format_timespan(timer()-start))
# y_pred_proba = model.predict_proba(X_test)[:, 1]
_y_pred_proba = _model.predict_proba(X)[:, 1]
#y_pred_proba
print(y_pred_proba.shape)
_pred_ranks = pd.Series(_y_pred_proba, index=X.index, name='pred_ranks')
#test_papers_df.join(_pred_ranks).sort_values('pred_ranks', ascending=False).head()
_top_predictions = test_papers_df.join(_pred_ranks).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
_top_predictions.groupby('target')['Paper_ID'].count()
# -
start = timer()
toplevels = test_papers_df.cl.apply(lambda x: x.split(":")[0])
print(format_timespan(timer()-start))
toplevels.name = 'toplevel'
toplevels_set = set(toplevels)
start = timer()
tbl = db.tables['clusters_meta_tree']
sq = tbl.select(tbl.c.toplevel_in_tree.in_(toplevels_set))
# r = db.engine.execute(sq).fetchall()
cl_meta = db.read_sql(sq)
print(format_timespan(timer()-start))
cl_meta = cl_meta.set_index('id')
train_papers_df['toplevel'] = train_papers_df.cl.apply(lambda x: x.split(":")[0]).astype(int)
meta_map = cl_meta.set_index('toplevel_in_tree').meta_cl
train_papers_df['cl_meta'] = train_papers_df.toplevel.map(meta_map)
test_papers_df['toplevel'] = toplevels.astype(int)
test_papers_df['cl_meta'] = test_papers_df.toplevel.map(meta_map)
start = timer()
test_papers_df['meta_avg_distance_to_train'] = test_papers_df.cl_meta.apply(avg_distance, cl_group=train_papers_df.cl_meta.tolist())
print(format_timespan(timer()-start))
# +
# logistic regression including meta cl
X = test_papers_df[['EF', 'avg_distance_to_train', 'meta_avg_distance_to_train']]
y = test_papers_df['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=999)
start = timer()
model_meta = LogisticRegression()
model_meta.fit(X_train, y_train)
print(format_timespan(timer()-start))
# y_pred_proba = model.predict_proba(X_test)[:, 1]
y_pred_proba_meta = model_meta.predict_proba(X)[:, 1]
#y_pred_proba
print(y_pred_proba_meta.shape)
pred_ranks_meta = pd.Series(y_pred_proba_meta, index=X.index, name='pred_ranks')
#test_papers_df.join(_pred_ranks).sort_values('pred_ranks', ascending=False).head()
top_predictions_meta = test_papers_df.join(pred_ranks_meta).sort_values('pred_ranks', ascending=False).head(len(target_papers_df))
top_predictions_meta.groupby('target')['Paper_ID'].count()
# -
from sklearn.metrics import roc_auc_score
print(roc_auc_score(y, y_pred_proba))
print(roc_auc_score(y, y_pred_proba_meta))
print(roc_auc_score(y, _y_pred_proba))
| old/plan_tfidf_cosine_similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
import warnings
warnings.simplefilter('ignore')
# -
# # Seminar 7 - Погружение в глубокое обучение
# В семинаре, будем использовать набор данных `fashion_mnist`, загрузим их
# +
from keras.datasets import fashion_mnist
(x_train, y_train_cat), (x_test, y_test_cat) = fashion_mnist.load_data()
print('Training data shape: ', x_train.shape)
print('Test data shape: ', x_test.shape)
num_classes = 10
class_names = ['T-shirt/top', 'Trouser', 'Pullover',
'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# -
# Сгенерируем случаные примеры для каждого класса и посмотрим на них.
fig = plt.figure(figsize=(15,5))
for i in range(num_classes):
ax = fig.add_subplot(2, 5, 1 + i, xticks=[], yticks=[])
idx = np.where(y_train_cat[:]==i)[0]
features_idx = x_train[idx,::]
img_num = np.random.randint(features_idx.shape[0])
im = features_idx[img_num]
ax.set_title(class_names[i])
plt.imshow(im, cmap='gray_r')
plt.show()
# ## Построим нашу первую нейросеть
# Импорт `Keras`
from keras.models import Sequential # Модель, где все слои соединены друг с другом
from keras.layers import Dense, Flatten, Activation # Слой, где все нйероны предыдущего уровня соединены с нейронами следующего
from keras.utils import np_utils
from keras.optimizers import SGD,Adam,RMSprop
# ### Проведем небольшие предобработки
y_train = np_utils.to_categorical(y_train_cat, num_classes=num_classes)
y_test = np_utils.to_categorical(y_test_cat, num_classes=num_classes)
# ### Построим модель
input_shape = x_train.shape[1:]
# +
# Создаем последовательную модель
model = Sequential()
# Добавляем слои
model.add(Flatten(input_shape=input_shape))
model.add(Dense(200))
model.add(Activation('sigmoid'))
model.add(Dense(100))
model.add(Activation('sigmoid'))
model.add(Dense(10))
model.add(Activation('softmax'))
# Компилируем модель
optimizer = SGD(lr=0.1)
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['accuracy'])
# -
print(model.summary())
# #### Вопрос:
# Почему: $784 \times 200 = 156 800$, а сетка показывает число параметров: $157000$?
# ### Обучим модель, задав параметры
batch_size = 1000 # Выбираем размер Батча
epochs = 100
# Обучаем модель!
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
plt.plot(history.history['acc'], label='train')
plt.plot(history.history['val_acc'], label='val')
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.show()
# ## Что мы можем улучшить?
# - Отнормировать признаки
# - Заменить сигмоиды на ReLu
# - Задать правила инициации весов
# ### Нормирование
# <img src='normalize.png'>
x_train = (x_train/255)*2-1
x_test = (x_test/255)*2-1
# ### Функции активации
# <img src='activations.png'>
# ### Инициациия весов
# __Случайно__
# $ w = a * random$, но тогда если $a \gg 1$, то на выходе $b\gg1$ и если $a \ll 1 $, то $b \approx 0 $
#
# __Xavier__
# $a = \frac{1}{\sqrt{n}}$, где $n$ - кол-во нейронов на входе
#
# __He__
# $a = \frac{1}{\sqrt{\frac{n}{2}}}$, где $n$ - кол-во нейронов на входе
from keras.initializers import he_normal
def create_model(input_shape, optimizer):
model = Sequential()
model.add(Flatten(input_shape=input_shape))
model.add(Dense(200, kernel_initializer=he_normal()))
model.add(Activation('relu'))
model.add(Dense(100, kernel_initializer=he_normal()))
model.add(Activation('relu'))
model.add(Dense(10))
model.add(Activation('softmax'))
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['accuracy'])
return model
optimizer = SGD(lr=0.1)
model = create_model(input_shape, optimizer)
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# +
def viz_history(history):
plt.plot(history.history['acc'], label='train')
plt.plot(history.history['val_acc'], label='val')
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.show()
viz_history(history)
# -
# ## Влияние скорости обучения
# Посмотрим, как влияет параметр `learning_rate` на качество нашей модели на обучающей выборке
# +
for i in tqdm([0.001, 0.01, 0.1, 1]):
optimizer = SGD(lr=i)
model = create_model(input_shape, optimizer)
history = model.fit(x_train, y_train, verbose=0,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
plt.plot(history.history['val_acc'], label='learning rate = {}'.format(i))
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.show()
# -
# ## Влияние метода оптимизации градиентного спуска
# <img src='optimizers7.gif'>
# #### Momentum
# Вместо того, чтобы использовать только градиент текущего шага, мы будем накапливать импульс градиента прошлых шагов для определения направления движения.
# В связи со стохастической природой, обновления градиента происходят "зигзагообразно", с помощью момента мы усиливаем движение вдоль основного направления. На практике коэффициент у момента инициализируется на уровне 0,5 и постепенно увеличивается до 0,9 в течение нескольких эпох.
#
# #### RMSProp (Root Mean Square Propogation)
#
#
# #### Adam (Adaptive moment estimation)
# Cочетает в себе и идею накопления движения и идею более слабого обновления весов для типичных признаков
# +
names = ['SGD with Momentum', 'Adam', 'RMSprop']
optimizers = {'SGD with Momentum': SGD(nesterov=True),
'Adam': Adam(),
'RMSprop': RMSprop()
}
for opt in tqdm(names):
model = create_model(input_shape, optimizers[opt])
history = model.fit(x_train, y_train, verbose=0,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
plt.plot(history.history['val_acc'], label='optimizers = {}'.format(opt))
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.show()
# -
# ## Применим, все полученные знания
# +
optimizer = Adam()
model = create_model(input_shape, optimizer)
history = model.fit(x_train, y_train, verbose=1,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# -
viz_history(history)
# ## Объявляем, борьбу с переобучением
from keras.layers import Dropout, BatchNormalization
model = Sequential()
model.add(Flatten(input_shape=input_shape))
model.add(Dense(200, kernel_initializer=he_normal()))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(100, kernel_initializer=he_normal()))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(10))
model.add(Activation('softmax'))
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(x_train, y_train, verbose=1,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
viz_history(history)
# # Теперь твоя очередь!
# <NAME> 2 в соревновании [Птица или самолет](https://www.kaggle.com/c/bird-or-aircraft/leaderboard)
# # Ссылки
# - [Курс "Deep learning на пальцах", лекция 4](https://youtu.be/tnrbx7V9RbA)
# - [Статья: Оптимизация градиентного спуска](http://ruder.io/optimizing-gradient-descent/)
# - [Статья: Методы оптимизации нейронных сетей](https://habr.com/ru/post/318970/)
| week 07 - deep learning/Seminar 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # IDScalarWaveNRPy: An Einstein Toolkit Initial Data Thorn for the Scalar Wave Equation
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# [comment]: <> (Abstract: TODO)
#
# [comment]: <> (Notebook Status and Validation Notes: TODO)
#
# ### NRPy+ Source Code for this module: [ScalarWave/InitialData_PlaneWave.py](../edit/ScalarWave/InitialData_PlaneWave.py) [\[**tutorial**\]](Tutorial-ScalarWave.ipynb) Contructs the SymPy expressions for plane-wave initial data
#
# ## Introduction:
# In this part of the tutorial, we will construct an Einstein Toolkit (ETK) thorn (module) that will set up *initial data* for the scalar wave initial value problem. In a [previous tutorial notebook](Tutorial-ScalarWave.ipynb), we used NRPy+ to contruct the SymPy expressions for plane-wave initial data. This thorn is largely based on and should function similarly to the $\text{IDScalarWaveC}$ thorn included in the Einstein Toolkit (ETK) $\text{CactusWave}$ arrangement.
#
# We will construct this thorn in two steps.
#
# 1. Call on NRPy+ to convert the SymPy expressions for the initial data into one C-code kernel.
# 1. Write the C code and linkages to the Einstein Toolkit infrastructure (i.e., the .ccl files) to complete this Einstein Toolkit module.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Call on NRPy+ to convert the SymPy expression for the scalar wave initial data into a C-code kernel
# 1. [Step 2](#einstein): Interfacing with the Einstein Toolkit
# 1. [Step 2.a](#einstein_c): Constructing the Einstein Toolkit C-code calling functions that include the C code kernels
# 1. [Step 2.b](#einstein_ccl): CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure
# 1. [Step 2.c](#einstein_list): Add the C code to the Einstein Toolkit compilation list
# 1. [Step 3](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Call on NRPy+ to convert the SymPy expression for the scalar wave initial data into a C-code kernel \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# After importing the core modules, since we are writing an ETK thorn, we'll need to set `"grid::GridFuncMemAccess"` to `"ETK"`. SymPy expressions for plane wave initial data are written inside [ScalarWave/InitialData_PlaneWave.py](../edit/ScalarWave/InitialData_PlaneWave.py), and we simply import them for use here.
# +
# Step 1: Call on NRPy+ to convert the SymPy expression
# for the scalar wave initial data into a C-code kernel
# Step 1a: Import needed NRPy+ core modules:
import NRPy_param_funcs as par
import indexedexp as ixp
import grid as gri
import finite_difference as fin
from outputC import *
import loop
# Step 1b: This is an Einstein Toolkit (ETK) thorn. Here we
# tell NRPy+ that gridfunction memory access will
# therefore be in the "ETK" style.
par.set_parval_from_str("grid::GridFuncMemAccess","ETK")
# Step 1c: Call the InitialData_PlaneWave() function from within the
# ScalarWave/InitialData_PlaneWave.py module.
import ScalarWave.InitialData_PlaneWave as swid
# Step 1d: Within the ETK, the 3D gridfunctions x, y, and z store the
# Cartesian grid coordinates. Setting the gri.xx[] arrays
# to point to these gridfunctions forces NRPy+ to treat
# the Cartesian coordinate gridfunctions properly --
# reading them from memory as needed.
x,y,z = gri.register_gridfunctions("AUX",["x","y","z"])
gri.xx[0] = x
gri.xx[1] = y
gri.xx[2] = z
# Step 1e: Set up the plane wave initial data. This sets uu_ID and vv_ID.
swid.InitialData_PlaneWave()
# Step 1f: Register uu and vv gridfunctions so they can be written to by NRPy.
uu,vv = gri.register_gridfunctions("EVOL",["uu","vv"])
# Step 1g: Set the uu and vv gridfunctions to the uu_ID & vv_ID variables
# defined by InitialData_PlaneWave().
uu = swid.uu_ID
vv = swid.vv_ID
# Step 1h: Create the C code output kernel.
scalar_PWID_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","uu"),rhs=uu),\
lhrh(lhs=gri.gfaccess("out_gfs","vv"),rhs=vv),]
scalar_PWID_CcodeKernel = fin.FD_outputC("returnstring",scalar_PWID_to_print)
scalar_PWID_looped = loop.loop(["i2","i1","i0"],["1","1","1"],["cctk_lsh[2]-1","cctk_lsh[1]-1","cctk_lsh[0]-1"],\
["1","1","1"],["#pragma omp parallel for","",""],"",\
scalar_PWID_CcodeKernel.replace("time","cctk_time"))
# Step 1i: Create directories for the thorn if they don't exist.
# !mkdir IDScalarWaveNRPy 2>/dev/null # 2>/dev/null: Don't throw an error if the directory already exists.
# !mkdir IDScalarWaveNRPy/src 2>/dev/null # 2>/dev/null: Don't throw an error if the directory already exists.
# Step 1j: Write the C code kernel to file.
with open("IDScalarWaveNRPy/src/ScalarWave_PWID.h", "w") as file:
file.write(str(scalar_PWID_looped))
# -
# <a id='einstein'></a>
#
# # Step 2: Interfacing with the Einstein Toolkit \[Back to [top](#toc)\]
# $$\label{einstein}$$
#
# <a id='einstein_c'></a>
#
# ## Step 2.a: Constructing the Einstein Toolkit C-code calling functions that include the C code kernels \[Back to [top](#toc)\]
# $$\label{einstein_c}$$
#
# We will write another C file with the functions we need here.
# +
# %%writefile IDScalarWaveNRPy/src/InitialData.c
#include <math.h>
#include <stdio.h>
#include "cctk.h"
#include "cctk_Parameters.h"
#include "cctk_Arguments.h"
void IDScalarWaveNRPy_param_check(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
if (kk0 == 0 && kk1 == 0 && kk2 == 0) {
CCTK_WARN(0,"kk0==kk1==kk2==0: Zero wave vector cannot be normalized. Set one of the kk's to be != 0.");
}
}
void IDScalarWaveNRPy_InitialData(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS
DECLARE_CCTK_PARAMETERS
const CCTK_REAL *xGF = x;
const CCTK_REAL *yGF = y;
const CCTK_REAL *zGF = z;
#include "ScalarWave_PWID.h"
}
# -
# <a id='einstein_ccl'></a>
#
# ## Step 2. b: CCL files - Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure \[Back to [top](#toc)\]
# $$\label{einstein_ccl}$$
#
# Writing a module ("thorn") within the Einstein Toolkit requires that three "ccl" files be constructed, all in the root directory of the thorn:
#
# 1. `interface.ccl`: defines the gridfunction groups needed, and provides keywords denoting what this thorn provides and what it should inherit from other thorns. Specifically, this file governs the interaction between this thorn and others; more information can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-178000D2.2).
# With "implements", we give our thorn its unique name. By "inheriting" other thorns, we tell the Toolkit that we will rely on variables that exist and are declared "public" within those functions.
# +
# %%writefile IDScalarWaveNRPy/interface.ccl
implements: IDScalarWaveNRPy
inherits: WaveToyNRPy grid
# -
# 2. `param.ccl`: specifies free parameters within the thorn, enabling them to be set at runtime. It is required to provide allowed ranges and default values for each parameter. More information on this file's syntax can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-183000D2.3).
# +
# %%writefile IDScalarWaveNRPy/param.ccl
shares: grid
USES KEYWORD type
shares: WaveToyNRPy
USES REAL wavespeed
restricted:
CCTK_KEYWORD initial_data "Type of initial data"
{
"plane" :: "Plane wave"
} "plane"
restricted:
CCTK_REAL kk0 "The wave number in the x-direction"
{
*:* :: "No restriction"
} 4.0
restricted:
CCTK_REAL kk1 "The wave number in the y-direction"
{
*:* :: "No restriction"
} 0.0
restricted:
CCTK_REAL kk2 "The wave number in the z-direction"
{
*:* :: "No restriction"
} 0.0
# -
# 3. `schedule.ccl`: allocates storage for gridfunctions, defines how the thorn's functions should be scheduled in a broader simulation, and specifies the regions of memory written to or read from gridfunctions. `schedule.ccl`'s official documentation may be found [here](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-186000D2.4).
#
# We specify here the standardized ETK "scheduling bins" in which we want each of our thorn's functions to run.
# +
# %%writefile IDScalarWaveNRPy/schedule.ccl
schedule IDScalarWaveNRPy_param_check at CCTK_PARAMCHECK
{
LANG: C
OPTIONS: global
} "Check sanity of parameters"
schedule IDScalarWaveNRPy_InitialData at CCTK_INITIAL as WaveToy_InitialData
{
STORAGE: WaveToyNRPy::scalar_fields[3]
LANG: C
READS: grid::x(Everywhere)
READS: grid::y(Everywhere)
READS: grid::y(Everywhere)
WRITES: uuGF(Everywhere)
WRITES: vvGF(Everywhere)
} "Initial data for 3D wave equation"
# -
# <a id='einstein_list'></a>
#
# ## Step 2.c: Add the C code to the Einstein Toolkit compilation list \[Back to [top](#toc)\]
# $$\label{einstein_list}$$
#
# We will also need `make.code.defn`, which indicates the list of files that need to be compiled. This thorn only has the one C file to compile.
# +
# %%writefile IDScalarWaveNRPy/src/make.code.defn
SRCS = InitialData.c
# -
# <a id='latex_pdf_output'></a>
#
# # Step 3: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ETK_thorn-IDScalarWaveNRPy.pdf](Tutorial-ETK_thorn-IDScalarWaveNRPy.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-ETK_thorn-IDScalarWaveNRPy.ipynb
# !pdflatex -interaction=batchmode Tutorial-ETK_thorn-IDScalarWaveNRPy.tex
# !pdflatex -interaction=batchmode Tutorial-ETK_thorn-IDScalarWaveNRPy.tex
# !pdflatex -interaction=batchmode Tutorial-ETK_thorn-IDScalarWaveNRPy.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
| Tutorial-ETK_thorn-IDScalarWaveNRPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="a_VmAfWSBJs7"
# <img width=150 src="https://upload.wikimedia.org/wikipedia/commons/thumb/1/1a/NumPy_logo.svg/200px-NumPy_logo.svg.png"></img>
# + [markdown] id="S1pT1uwSqLo1"
# * 範例目標:
# 1. 實做欄位索引之間轉換
# 2. 重新組織資料
# * 範例重點:
# 1. 不管是欄位轉索引或是索引轉欄位,皆由最外層的開始轉換
# 2. 重新組織資料時應注意參數的理解,可以多做嘗試
# + id="cu78CXS6mRpg"
# 載入 NumPy, Pandas 套件
import numpy as np
import pandas as pd
# 檢查正確載入與版本
print(np)
print(np.__version__)
print(pd)
print(pd.__version__)
# -
# # 【基礎15=進階14】
# ## from_product
# + colab={"base_uri": "https://localhost:8080/", "height": 235} executionInfo={"elapsed": 889, "status": "ok", "timestamp": 1608469184981, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="C56aVXSRvRN3" outputId="b1750972-57f8-4a05-82ac-95e44fe45ab8"
index = pd.MultiIndex.from_product([[2013, 2014], [1, 2]],
names=['year', 'visit'])
columns = pd.MultiIndex.from_product([['Bob', 'Guido', 'Sue'], ['HR', 'Temp']],
names=['subject', 'type'])
# mock some data
data = np.round(np.random.randn(4, 6), 1)
df = pd.DataFrame(data, index=index, columns=columns)
df
# -
# ### 欄位轉索引
# * 將一欄位 (column) 轉成一索引 (index),使用 .stack() 即可,可以將 type 這個欄位轉成了索引,所以索引變成了 year、visit、type
# + colab={"base_uri": "https://localhost:8080/", "height": 328} executionInfo={"elapsed": 768, "status": "ok", "timestamp": 1608468894427, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="kJi9Yhbnt0w4" outputId="c79d249f-0ab9-4ea5-bf39-ab67423b3194"
df.stack()
# -
# * 再做一次.stack()索引變成了year、visit、type、subject
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 743, "status": "ok", "timestamp": 1608468905484, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="remuFEB3N6rN" outputId="c73a3343-f3ff-481c-f057-4b337ec0e5c2"
df.stack().stack()
# -
# ### 索引轉欄位
# * 將一索引(index)轉成一欄位(column) ,使用.unstack()即可,可以將visit這個索引轉成了欄位,所以欄位變成了subject、type 、visit
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 803, "status": "ok", "timestamp": 1608469090249, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="GSq8uoPJuJFl" outputId="0b6660b0-10fa-406c-828f-3394f9d86df2"
df.unstack()
# -
# ## from_tuples
# +
# 設定給以下 dataframe 的索引名稱
multicol1 = pd.MultiIndex.from_tuples([('weight', 'kg'),
('weight', 'pounds')])
# 創建我們的範例 dataframe
df_multi_level_cols1 = pd.DataFrame([[1, 2], [2, 4]],
index=['cat', 'dog'],
columns=multicol1)
df_multi_level_cols1
# -
# ### 欄位轉索引
# 使用默認的 .stack() (也就是讓入最下面的欄位)
df_multi_level_cols1.stack()
df_multi_level_cols1.index, df_multi_level_cols1.stack().index
df_multi_level_cols1.stack().unstack()
df_multi_level_cols1.stack(), df_multi_level_cols1.stack().unstack().index
df_multi_level_cols1.stack().unstack().unstack().unstack().unstack().unstack().unstack()
# ## 欄位名稱轉為欄位值
# + colab={"base_uri": "https://localhost:8080/", "height": 142} executionInfo={"elapsed": 883, "status": "ok", "timestamp": 1608466648605, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="8estv2CNEBqf" outputId="75cf5d41-2725-42fe-8f02-0bd6c3a190a7"
df = pd.DataFrame({'Name':{0:'John', 1:'Bob', 2:'Shiela'},
'Course':{0:'Masters', 1:'Graduate', 2:'Graduate'},
'Age':{0:27, 1:23, 2:21}})
df
# -
# ### 保留Name欄位其餘轉成欄位值
# + colab={"base_uri": "https://localhost:8080/", "height": 328} executionInfo={"elapsed": 523, "status": "ok", "timestamp": 1608466655475, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="RuLqccKODP8D" outputId="9017ae36-08a1-47d5-9c14-86c1ddd2237d"
df.melt()
# -
# ### 只轉換Name欄位
# + colab={"base_uri": "https://localhost:8080/", "height": 235} executionInfo={"elapsed": 912, "status": "ok", "timestamp": 1608467012013, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="cjGSPlHIGoXH" outputId="3620133e-6816-48d6-f3f0-0838c364b804"
df.melt(id_vars='Name')
# -
# ### 保留Name欄位其餘轉成欄位值
# 之後再留下value_vars='Name'
# + colab={"base_uri": "https://localhost:8080/", "height": 142} executionInfo={"elapsed": 843, "status": "ok", "timestamp": 1608467266032, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="msqnURPMHpwb" outputId="4b0b065d-4c8f-485d-da71-219bfb952f6a"
df.melt(value_vars='Name')
# -
# ## 重新組織資料
# + colab={"base_uri": "https://localhost:8080/", "height": 235} executionInfo={"elapsed": 915, "status": "ok", "timestamp": 1608472058229, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="MZ_zT1qStT5C" outputId="c35d4936-dab3-4b1d-d8a7-61aff81eea08"
df = pd.DataFrame({'fff': ['one', 'one', 'one', 'two', 'two',
'two'],
'bbb': ['P', 'Q', 'R', 'P', 'Q', 'R'],
'baa': [2, 3, 4, 5, 6, 7],
'zzz': ['h', 'i', 'j', 'k', 'l', 'm']})
df
# -
# ### .pivot() 函數根據給定的索引/列值重新組織給定的DataFrame
# * 參數
# * index:新資料的索引名稱
# * columns:新資料的欄位名稱
# * values:新資料的值名稱
# + colab={"base_uri": "https://localhost:8080/", "height": 142} executionInfo={"elapsed": 680, "status": "ok", "timestamp": 1608472058689, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="wNwoucSEZdAZ" outputId="fa1f79e2-e1bc-4eaf-8748-1a63a2800f2a"
df.pivot(index='fff', columns='bbb', values='baa')
# -
df.pivot(index='fff', columns='bbb')['baa']
df.pivot(index='fff', columns='bbb', values=['baa', 'zzz'])
| Sample/Day_15_Sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Zilleplus/MachineLearning/blob/main/Shakespeare.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YmA9L3g1_ISw"
import tensorflow as tf
import tensorflow.keras as keras
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="sprF6clN-76m" outputId="2940df25-aab4-4fea-83ba-3359d986f0f1"
shakespeare_url = "https://homl.info/shakespeare"
filepath = keras.utils.get_file("shakespear.text", shakespeare_url)
with open(filepath) as f:
shakespeare_text = f.read()
# + id="FKfkTQAQ_ZhA"
tokenizer = keras.preprocessing.text.Tokenizer(char_level=True)
tokenizer.fit_on_texts(shakespeare_text)
# + colab={"base_uri": "https://localhost:8080/"} id="LO0KBBwz_qYj" outputId="d152b425-c20b-479c-e92b-c958c49f1d20"
print(tokenizer.texts_to_sequences(["First"]))
print(tokenizer.sequences_to_texts([[20, 6,9,8,3]]))
# + id="p-kojwk7_9FV"
[encoded] = np.array(tokenizer.texts_to_sequences([shakespeare_text]))
# + id="P1xn6XvP7nNX"
# The windowed dataset, is a dataset of datasets
for e in tf.data.Dataset.from_tensor_slices([1, 2, 3]).window(2):
print("This is a dataset:")
print(e)
print("The dataset contains 2 elements:")
for k in e:
print(k)
break
print("----")
# used flat_map to transforms each window dataset into tensors, batch outputs
# a iterator with one element, flat_map unravels to the single element.
flat_data_example = tf.data.Dataset\
.from_tensor_slices([1, 2, 3])\
.window(2)\
.flat_map(lambda window: window.batch(2))
for e in flat_data_example:
print(e)
print(type(flat_data_example))
# notice the last tensor with [3], that does not have window lenght=2, this is why we enable drop_remainder
# + id="Q_XW98melTv_"
dataset_size = encoded.size
train_size = (dataset_size*90)//100
dataset = tf.data.Dataset.from_tensor_slices(encoded[:train_size])
n_steps = 100
window_length = n_steps + 1 # Input shift 1 character ahead.
dataset = dataset\
.window(window_length, shift=1, drop_remainder=True)\
.flat_map(lambda window: window.batch(window_length))
# + id="oS2weZpi7YDH"
for d in dataset:
print(d)
break
# + id="MmcBzfFt-yie"
# randomize the data
batch_size = 32
dataset = dataset\
.shuffle(10000)\
.batch(batch_size)\
.map(lambda windows: (windows[:, :-1], windows[:, 1:])) # split off the last element, the first dimension is batch, second is series
# + id="WRjNp4IhAkTI"
# use one hot encoding
max_id = len(tokenizer.word_index)
dataset = dataset.map(
lambda X_batch, Y_batch: (tf.one_hot(X_batch, depth=max_id), Y_batch))
# + id="9Hf2tFHrFxrG"
# add prefetch
dataset = dataset.prefetch(buffer_size=1)
# + id="OeKtmaIyGqgu"
model = keras.models.Sequential([
keras.layers.GRU(units=128, return_sequences=True, dropout=0.2, input_shape=[None, max_id]), # recurrent_dropout=0.2
keras.layers.GRU(units=128, return_sequences=True, dropout=0.2), # recurrent_dropout=0.2
keras.layers.TimeDistributed(keras.layers.Dense(max_id, activation='softmax')) # add softmax at the end to get [0,1] intervals
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="adam")
history = model.fit(dataset, epochs=20)
# + id="o4Og8LGpJZ6m" colab={"base_uri": "https://localhost:8080/"} outputId="6ecf840a-2b7d-4afb-bebe-67096a550524"
# find random samples from distribution p(0)=0.7 and p(1)=0.3
tf.random.categorical(tf.math.log([[0.7, 0.3]]), 10)
| HML/chapter16/Shakespeare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow-gpu-test]
# language: python
# name: conda-env-tensorflow-gpu-test-py
# ---
# +
#Question #239 Sliding Window Maximum
#Description:
#Given an array nums, there is a sliding window of size k which is moving from the very left of the array to the very right.
#You can only see the k numbers in the window. Each time the sliding window moves right by one position.
#Return the max sliding window.
#Example:
#Input: nums = [1,3,-1,-3,5,3,6,7], and k = 3
#Output: [3,3,5,5,6,7]
#Explanation
#Window position Max
#--------------- -----
#[1 3 -1] -3 5 3 6 7 3
# 1 [3 -1 -3] 5 3 6 7 3
# 1 3 [-1 -3 5] 3 6 7 5
# 1 3 -1 [-3 5 3] 6 7 5
# 1 3 -1 -3 [5 3 6] 7 6
# 1 3 -1 -3 5 [3 6 7] 7
#import necessary package
import collections
#defines the method for the problem
#input: self, nums, and k
#idea: Use double-ended queue
#steps:
#1. Maintain a deque of the indexes of the largest elements we've seen (candidates)
#2. Deque should never point to elements smaller than current element
#3. Deque should never point to elements outside the sliding window
#4. Keep updating the deque with current element
#5. Keep appending the front of deque to output
def maxSlidingWindow(self, nums, k):
"""
:type nums: List[int]
:type k: int
:rtype: List[int]
"""
d = collections.deque()
out = []
for i, n in enumerate(nums):
#print("i = {}, curr element = {}, d = {} and out = {}".format(i, n, d, out))
while d and nums[d[-1]] < n:
d.pop()
#print("\t Popped from d because d has elements and nums[d.top] < curr element")
d.append(i)
#print("\t Added i to d")
if d[0] == i - k:
d.popleft()
#print("\t Popped left from d because it's outside the window's leftmost (i-k)")
if i>=k-1:
out.append(nums[d[0]])
#print("\t Append nums[d[0]] = {} to out".format(nums[d[0]]))
return out
# +
#test case
#input: nums = [1,3,-1,-3,5,3,6,7], k =3
#exprected output: [3, 3, 5, 5, 6, 7]
nums = [1,3,-1,-3,5,3,6,7]
k = 3
#print output
print(maxSlidingWindow(_, nums, k))
# -
| Sliding_Window_Maximum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt # for 畫圖用
import pandas as pd
# Feature Scaling
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Dropout
from openpyxl import load_workbook
from openpyxl import Workbook
# Import the training set
dataset_train = pd.read_excel('B(半導體).xlsx') # 讀取訓練集
dataset_test = pd.read_excel('B(半導體)_測試.xlsx')
dataset_realtest = pd.read_excel('B(半導體)_50天_ques.xlsx')
data_all = pd.concat((dataset_train,dataset_test), axis = 0)
data_all=data_all.fillna(method="ffill",limit=10)
#dataset_train=dataset_train.dropna(axis=1)
training_set = data_all.iloc[:,[2,5,31,32,33,34,36,41,42,43,44,45,46]]#.values # 取欄位值
training_set1= data_all.iloc[:, 1:2].values
training_set
real_stock_price =data_all.iloc[2100:2232,1].values
train = data_all.iloc[:2100,[2,5,31,32,33,34,36,41,42,43,44,45,46]]
sc = MinMaxScaler(feature_range = (0, 1))
training_set_scaled = sc.fit_transform(training_set)
sc1 = MinMaxScaler(feature_range = (0, 1))
training_set_scaled1 = sc1.fit_transform(training_set1)
# +
X_train = []
y_train = []
for i in range(7, 2100): # 2100 是訓練集總數
X_train.append(training_set_scaled[i-7:i,:])
y_train.append(training_set_scaled1[i,0])
X_train, y_train = np.array(X_train), np.array(y_train) # 轉成numpy array的格式,以利輸入 RNN
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1]*X_train.shape[2], 1))
# Import the Keras libraries and packages
# Initialising the RNN
regressor = Sequential()
# +
# Adding the first LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50, return_sequences = True, input_shape = (X_train.shape[1], 1)))
regressor.add(Dropout(0.2))
# Adding a second LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
# Adding a third LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
# Adding a fourth LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50))
regressor.add(Dropout(0.2))
# Adding the output layer
regressor.add(Dense(units = 1))
# Compiling
regressor.compile(optimizer = 'adam', loss = 'mean_squared_error')
# 進行訓練
regressor.fit(X_train, y_train, epochs = 100, batch_size = 32)
# +
#test = sc.fit_transform(training_set)
# -
X_test=[]
for i in range(2101, 2232): # 1747 是訓練集總數
X_test.append(training_set_scaled[i-7:i,0:])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1]*X_test.shape[2], 1))
# +
#list(X_test)
# -
predicted_stock_price = regressor.predict(X_test)
predicted_stock_price = sc1.inverse_transform(predicted_stock_price)
plt.plot(predicted_stock_price,color='red',label= 'predicted_stock_price')
plt.plot(real_stock_price,color = 'blue',label='real_stock_price')
plt.title('B Stock Price Prediction')
plt.xlabel('Time')
plt.ylabel('B Price')
wb = Workbook()
ws = wb.active
ws['A1'] = '收盤價'
| B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Modeling of bank failures by FDIC
#
# +
import pandas as pd
import numpy as np
import time
import os
import functools
import math
import random
import sys, getopt
import sklearn
sys.path.append("..")
import grading
try:
import matplotlib.pyplot as plt
# %matplotlib inline
except:
pass
print('scikit-learn version:', sklearn.__version__)
# -
### ONLY FOR GRADING. DO NOT EDIT ###
submissions=dict()
assignment_key="7VcH6P8REeeRWA42vRAjYg"
all_parts=["o5YYT", "2cHUA", "Mxrav","JFNf3", "ivHQa"]
### ONLY FOR GRADING. DO NOT EDIT ###
# token expires every 30 min
COURSERA_TOKEN = "<KEY>"# the key provided to the Student under his/her email on submission page
COURSERA_EMAIL = "<EMAIL>"# the email
# +
# common cell - share this across notebooks
state_cols = ['log_TA','NI_to_TA', 'Equity_to_TA', 'NPL_to_TL', 'REO_to_TA',
'ALLL_to_TL', 'core_deposits_to_TA', 'brokered_deposits_to_TA',
'liquid_assets_to_TA', 'loss_provision_to_TL', 'NIM', 'assets_growth']
all_MEVs = np.array(['term_spread',
'stock_mkt_growth',
'real_gdp_growth',
'unemployment_rate_change',
'treasury_yield_3m',
'bbb_spread',
'bbb_spread_change'])
MEV_cols = all_MEVs.tolist()
next_state_cols = ['log_TA_plus_1Q','NI_to_TA_plus_1Q', 'Equity_to_TA_plus_1Q', 'NPL_to_TL_plus_1Q', 'REO_to_TA_plus_1Q',
'ALLL_to_TL_plus_1Q', 'core_deposits_to_TA_plus_1Q', 'brokered_deposits_to_TA_plus_1Q',
'liquid_assets_to_TA_plus_1Q', 'loss_provision_to_TL_plus_1Q',
'ROA_plus_1Q',
'NIM_plus_1Q',
'assets_growth_plus_1Q',
'FDIC_assessment_base_plus_1Q_n']
# -
df_train = pd.read_hdf('../readonly/df_train_FDIC_defaults_1Y.h5', key='df')
df_test = pd.read_hdf('../readonly/df_test_FDIC_defaults_1Y.h5', key='df')
df_data = pd.read_hdf('../readonly/data_adj_FDIC_small.h5', key='df')
df_closure_learn = pd.read_hdf('../readonly/df_FDIC_learn.h5',key='df')
print(df_closure_learn.index.names)
# ## Construct training and testing datasets for logistic regression
df_test.plot(x=state_cols[0], y='defaulter', kind='scatter')
# +
# Plot 4 scatter plots together
# log_TA / NI_to_TA
# log_TA / NPL_to_TL
# log_TA / Equity_to_TA
# log_TA /ROA
first_indx = [0, 0, 0, 0]
second_indx = [1, 3, 2, 10]
X_train = df_train[state_cols].values
y_train = df_train.defaulter.values # .reshape(-1,1)
num_plots = 4
if num_plots % 2 == 0:
f, axs = plt.subplots(num_plots // 2, 2)
else:
f, axs = plt.subplots(num_plots// 2 + 1, 2)
f.subplots_adjust(hspace=.3)
f.set_figheight(10.0)
f.set_figwidth(10.0)
for i in range(num_plots):
if i % 2 == 0:
first_idx = i // 2
second_idx = 0
else:
first_idx = i // 2
second_idx = 1
axs[first_idx,second_idx].plot(X_train[y_train == 1.0, first_indx[i]],
X_train[y_train == 1.0, second_indx[i]], 'r^', label="Failed")
axs[first_idx,second_idx].plot(X_train[y_train == 0.0, first_indx[i]],
X_train[y_train == 0.0, second_indx[i]], 'go',label="Non-failed")
axs[first_idx, second_idx].legend()
axs[first_idx, second_idx].set_xlabel('%s' % state_cols[first_indx[i]])
axs[first_idx, second_idx].set_ylabel('%s' % state_cols[second_indx[i]])
axs[first_idx, second_idx].set_title('Failed banks vs non-failed banks')
axs[first_idx, second_idx].grid(True)
if num_plots % 2 != 0:
f.delaxes(axs[i // 2, 1])
# plt.savefig('Failed_vs_nonfailed_rr_plot.png')
# -
def calc_metrics(model, df_test, y_true, threshold=0.5):
"""
Arguments:
model - trained model such as DecisionTreeClassifier, etc.
df_test - Data Frame of predictors
y_true - True binary labels in range {0, 1} or {-1, 1}. If labels are not binary, pos_label should be explicitly given.
"""
if model is None:
return 0., 0., 0.
# prediction
predicted_sm = model.predict(df_test, linear=False)
predicted_binary = (predicted_sm > threshold).astype(int)
# print(predicted_sm.shape, y_true.shape)
fpr, tpr, _ = metrics.roc_curve(y_true, predicted_sm, pos_label=1)
# compute Area Under the Receiver Operating Characteristic Curve (ROC AUC) from prediction scores
roc_auc = metrics.auc(fpr, tpr)
ks = np.max(tpr - fpr) # Kolmogorov - Smirnov test
# note that here teY[:,0] is the same as df_test.default_within_1Y
accuracy_score = metrics.accuracy_score(y_true, predicted_binary)
# equivalently, Area Under the ROC Curve could be computed as:
# compute Area Under the Receiver Operating Characteristic Curve (ROC AUC) from prediction scores
# auc_score = metrics.roc_auc_score(y_true, predicted_sm)
try:
plt.title('Logistic Regression ROC curve')
plt.plot(fpr, tpr, 'b', label='AUC = %0.2f' % roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1], [0,1], 'r--')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
# plt.savefig('ROC_curve_1.png')
plt.show()
except:
pass
return roc_auc, accuracy_score, ks
def make_test_train(df_train, df_test, choice=0, predict_within_1Y=False):
"""
make the train and test datasets
Arguments:
choice - an integer 0 or -1. Controls selection of predictors.
Add tangible equity and assessment base as predictors
predict_within_1Y - boolean if True, predict defaults within one year
Return:
a tuple of:
- training data set predictors, np.array
- training data set : variable to predict, np.array
- test data set : variable to predict, np.array
- predictor variable names
"""
if choice == -1: # only state cols
predictors = state_cols
elif choice == 0: # original variables
predictors = state_cols + MEV_cols
trX = df_train[predictors].values
teX = df_test[predictors].values
num_features = len(predictors)
num_classes = 2
if predict_within_1Y == True:
trY = df_train[['default_within_1Y','no_default_within_1Y']].values
teY = df_test[['default_within_1Y','no_default_within_1Y']].values
else:
trY = df_train[['defaulter','non_defaulter']].values
teY = df_test[['defaulter','non_defaulter']].values
return trX, trY, teX, teY, predictors
# look at correlations
df_train[MEV_cols].corr()
# ## Logistic regression with statsmodels
#
# ### Part 1
# Perform logistic regression using **cols_to_use** as predictors. Use df_train pandas DataFrame as training data set, and df_test pandas DataDrame as testing data set to perform prediction based on the already trained model. Utilize statsmodels package. The result of fitting logistic regression should be assigned to variable named **model**
# +
import statsmodels.api as sm
from sklearn import metrics
cols_to_use = state_cols + MEV_cols + ['const']
model = None
df_train['const'] = 1
### START CODE HERE ### (≈ 3 lines of code)
# ....
logit = sm.Logit(df_train.defaulter, df_train[cols_to_use])
model = logit.fit()
### END CODE HERE ###
# +
# prediction
predicted_sm = np.array([])
### START CODE HERE ### (≈ 3 lines of code)
predicted_sm = model.predict(df_test[cols_to_use], linear=False) if model is not None else None
### END CODE HERE ###
threshold = 0.5
predicted_binary = (predicted_sm > threshold).astype(int)
auc_score, accuracy_score, ks = calc_metrics(model, df_test[cols_to_use], df_test.defaulter)
print('Accuracy score %f' % accuracy_score)
print('AUC score %f' % auc_score)
print('Kolmogorov-Smirnov statistic %f' % ks)
# note that here teY[:,0] is the same as df_test.default_within_1Y
# +
### GRADED PART (DO NOT EDIT) ###
part_1=[accuracy_score, auc_score, ks]
try:
part1 = " ".join(map(repr, part_1))
except TypeError:
part1 = repr(part_1)
submissions[all_parts[0]]=part1
grading.submit(COURSERA_EMAIL, COURSERA_TOKEN, assignment_key, all_parts[0],all_parts,submissions)
[accuracy_score, auc_score, ks]
### GRADED PART (DO NOT EDIT) ###
# -
# ## Logistic Regression with sklearn
#
# ### Part 2
# In Part 2 you will use scikit-learn to perform logistic regression using the same training and test datasets.
# Once the model is trained using trX, thisTrY, test it using teX, thisTeY and compute logistic regression score.
#
# - Use **"l1"** penalty
# - Set inverse of regularization strength to **1000.0**; must be a positive float. Like in support vector machines, smaller values specify stronger regularization.
# - Set tolerance to **1e-6**
# +
from sklearn import neighbors, linear_model
trX, trY, teX, teY, predictors = make_test_train(df_train, df_test)
lr_score = 0.
thisTrY = trY[:,0]
thisTeY = teY[:,0]
logistic = None # instantiate a model and reference it
result = None # result of fitting the model
### START CODE HERE ### (≈ 3 lines of code)
# .... define random_state argment in logistic regression class. Ininitialize it to 42
# such as this: random_state=42
# the variable name required for grading lr_score
logistic = linear_model.LogisticRegression(penalty='l1', tol=1e-6, C=1000.0)
result = logistic.fit(trX, thisTrY)
lr_score = result.score(teX, thisTeY)
### END CODE HERE ###
print('LogisticRegression score: %f' % lr_score)
# -
### GRADED PART (DO NOT EDIT) ###
part2=str(lr_score)
submissions[all_parts[1]]=part2
grading.submit(COURSERA_EMAIL, COURSERA_TOKEN, assignment_key, all_parts[:2],all_parts,submissions)
lr_score
### GRADED PART (DO NOT EDIT) ###
# **Instructions:**
# In this part you will again use scikit learn logistic regression but with different set of predictors. This will be a smaller set of predictor variables based on the analysis of P-values from the logistic regression. Use cols_to_use as predictors in df_train and df_test data sets. Use **defaulter** column as something to predict.
#
# Initialize reference to the logistic regression model **logistic** with an instance of appropriate class from scikit learn module and let **result** be the result of fitting the model to the training data set.
#
# Just as before initialize the model with the following parameters:
# - Use **"l1"** penalty
# - Set inverse of regularization strength to **1000.0**; must be a positive float. Like in support vector machines, smaller values specify stronger regularization.
# - Set tolerance to **1e-6**
# +
# Do Logistic Regression with a smaller number of predictor, based on analysis of P-values
# for the logistic regression with a full set of variables
# a smaller set is based on the analysis of P-values for the logistic regression
cols_to_use = ['log_TA', 'NI_to_TA', 'Equity_to_TA', 'NPL_to_TL',
'core_deposits_to_TA',
'brokered_deposits_to_TA',
'liquid_assets_to_TA'
] + ['term_spread', 'stock_mkt_growth']
lr_score = 0.
logistic = None
result = None
### START CODE HERE ### (≈ 3 lines of code)
# .... when initializing logistic regression class in 'sklearn', set random_state to 42 like this: random_state=42
# ... like this: random_state=42
# ... for grading, please store the logistic regression model into variable : logistic
random_state=42
trX = df_train[cols_to_use].values
teX = df_test[cols_to_use].values
thisTrY = (df_train.defaulter.values)
thisTeY = (df_test.defaulter.values)
logistic = linear_model.LogisticRegression(penalty='l1', tol=1e-6, C=1000.0)
result = logistic.fit(trX, thisTrY)
lr_score = result.score(teX, thisTeY)
print('LogisticRegression score: %f' % lr_score)
### END CODE HERE ###
# combine results of the Logistic Regression to a small dataframe df_coeffs_LR
df_coeffs_LR = pd.DataFrame({0: np.array([0.] * (len(cols_to_use) + 1), dtype=np.float32)})
if logistic is not None:
model_params = np.hstack((logistic.coef_[0], logistic.intercept_))
df_coeffs_LR = pd.DataFrame(data=model_params, index=cols_to_use + ['const'])
df_coeffs_LR
# -
### GRADED PART (DO NOT EDIT) ###
part_3=list(df_coeffs_LR.values.squeeze())
try:
part3 = " ".join(map(repr, part_3))
except TypeError:
part3 = repr(part_3)
submissions[all_parts[2]]=part3
grading.submit(COURSERA_EMAIL, COURSERA_TOKEN, assignment_key, all_parts[:3],all_parts,submissions)
df_coeffs_LR.values.squeeze()
### GRADED PART (DO NOT EDIT) ###
# ## Logistic Regression with Tensorflow
# +
# Setup inputs and expeced outputs for Logistic Regression using Tensorflow
cols = state_cols + MEV_cols
# inputs to Logistic Regression (via Tensorflow)
X_trainTf = df_train[cols].values
X_testTf = df_test[cols].values
# add constant columns to both
X_trainTf = np.hstack((np.ones((X_trainTf.shape[0], 1)), X_trainTf))
X_testTf = np.hstack((np.ones((X_testTf.shape[0], 1)), X_testTf))
# exepectd outputs:
y_trainTf = df_train.defaulter.astype('int').values.reshape(-1,1)
y_testTf = df_test.defaulter.astype('int').values.reshape(-1,1)
# -
print('Unique values to predict:', np.unique(y_trainTf))
print('Number of samples to train on:', y_trainTf.shape[0])
print('Number of samples to test on:', y_testTf.shape[0])
# to make this notebook's output stable across runs
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
def random_batch(X_train, y_train, batch_size):
np.random.seed(42)
rnd_indices = np.random.randint(0, len(X_train), batch_size)
X_batch = X_train[rnd_indices]
y_batch = y_train[rnd_indices]
return X_batch, y_batch
# ### Build Logistic Regression TF model
#
# **instructions**
#
# in tensorflow create:
# - placeholder for inputs called 'X'
# - placeholder for inputs called 'y'
# - variable for model parameters called 'theta', initialized with theta_init
#
# loss function: use log loss
# optimizer: use Gradient Descent optimizer
# +
import tensorflow as tf
# define the model
reset_graph()
n_inputs = X_trainTf.shape[1]
learning_rate = 0.01
theta_init = tf.random_uniform([n_inputs, 1], -1.0, 1.0, seed=42)
# build Logistic Regression model using Tensorflow
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.float32, shape=(None, 1), name="y")
theta = tf.Variable(theta_init, name="theta")
### START CODE HERE ### (≈ 6-7 lines of code)
### ....
### .... for grading please store probabilities in y_proba
logits = tf.matmul(X, theta)
y_proba = tf.sigmoid(logits) # = 1 / (1 + tf.exp(-logits))
# uses epsilon = 1e-7 by default to regularize the log function
loss = tf.losses.log_loss(y, y_proba, epsilon=1e-07)
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
### END CODE HERE ###
init = tf.global_variables_initializer()
# -
#
# ### Train Logistic Regression TF model
#
# **Instructions**
# - Use random_batch() function to grab batches from X_trainTf and y_trainTf.
# - Once the model is trained evaluate it based on X_testTf and y_testTf.
# - The **y_proba_val** should be assigned the result of the evaluation on test dataset.
# +
n_epochs = 1001
batch_size = 50
num_rec = X_trainTf.shape[0]
n_batches = int(np.ceil(num_rec / batch_size))
y_proba_val = np.array([], dtype=np.float32)
with tf.Session() as sess:
sess.run(init)
### START CODE HERE ### (≈ 6-7 lines of code)
## ....
for epoch in range(n_epochs):
for batch_index in range(n_batches):
X_batch, Y_batch = random_batch(X_trainTf, y_trainTf, batch_size)
_, l = sess.run([optimizer, loss], feed_dict = {X: X_batch, y: Y_batch})
y_proba_val = sess.run(y_proba, feed_dict = {X: X_testTf})
### END CODE HERE ###
# -
# predictions
threshold = 0.5
y_pred = (y_proba_val >= threshold)
print(np.sum(y_pred))
y_pred.squeeze()
# +
# evaluate precision, recall, and AUC
auc_score = 0.
ks = 0.
roc_auc = 0.
recall = 0.
precision = 0.
from sklearn.metrics import precision_score, recall_score
if y_proba_val.shape == y_testTf.shape:
precision = precision_score(y_testTf, y_pred)
recall = recall_score(y_testTf, y_pred)
auc_score = metrics.roc_auc_score(y_testTf, y_proba_val)
fpr, tpr, threshold = metrics.roc_curve(y_testTf, y_proba_val, pos_label=1)
roc_auc = metrics.auc(fpr, tpr)
ks = np.max(tpr - fpr)
print('precision: ', precision)
print('recall: ', recall)
print('AUC score = ', auc_score)
print('roc_auc = ', roc_auc)
print('KS_test = ', ks)
try:
plt.title('ROC_curve')
plt.plot(fpr, tpr, 'b', label='AUC = %0.2f' % roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1], [0,1], 'r--')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.savefig('ROC_curve_TF.png')
plt.show()
except:
pass
# -
### GRADED PART (DO NOT EDIT) ###
part_4=list([precision, recall, roc_auc, ks])
try:
part4 = " ".join(map(repr, part_4))
except TypeError:
part4 = repr(part_4)
submissions[all_parts[3]]=part4
grading.submit(COURSERA_EMAIL, COURSERA_TOKEN, assignment_key, all_parts[:4],all_parts,submissions)
[precision, recall, roc_auc, ks]
### GRADED PART (DO NOT EDIT) ###
# ## Neural Network with Tensorflow
# +
cols = state_cols + MEV_cols
n_inputs = len(cols)
# inputs
X_trainTf = df_train[cols].values
X_testTf = df_test[cols].values
# outputs
y_trainTf = df_train['defaulter'].astype('int').values.reshape(-1,)
y_testTf = df_test['defaulter'].astype('int').values.reshape(-1,)
# -
import numpy as np
def neuron_layer(X, n_neurons, name, activation=None):
with tf.name_scope(name):
tf.set_random_seed(42)
n_inputs = int(X.get_shape()[1])
stddev = 2 / np.sqrt(n_inputs)
init = tf.truncated_normal((n_inputs, n_neurons), stddev=stddev)
W = tf.Variable(init, name="kernel")
b = tf.Variable(tf.zeros([n_neurons]), name="bias")
Z = tf.matmul(X, W) + b
if activation is not None:
return activation(Z)
else:
return Z
# ### Construct Neural Network
#
# **Instructions**
# Implement Neural Network with two hidden layers. The number of nodes in the first and the second hidden layers is **n_hidden1** and **n_hidden2** correspondingly.
# Use neuron_layer() function to construct neural network layers.
#
# - Use ReLU activation function for hidden layers
# - The output layer has **n_outputs** and does not have an activation function
# - Use sparse softmax cross-entropy with logits as a loss function
# +
n_hidden1 = 20
n_hidden2 = 10
n_outputs = 2 # binary classification (defaulted, not defaulted bank)
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
### START CODE HERE ### (≈ 10-15 lines of code)
### ...
layer_1 = neuron_layer(X, n_hidden1, "layer_1", tf.nn.relu)
layer_2 = neuron_layer(layer_1, n_hidden2, "layer_2", tf.nn.relu)
logits = neuron_layer(layer_2, n_outputs, "logits")
entropy = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits, labels=y)
loss = tf.reduce_mean(entropy)
### END CODE HERE ###
init = tf.global_variables_initializer()
# -
# ### Train Neural Network
#
# **Instructions**
# Train neural network passing batches of inputs of size **batch_size**, which predicts bank defaults / non-defaults. Once the network is trained, evaluate accuracy using **X_testTf**, **y_testTf**
#
# +
learning_rate = 0.05
n_epochs = 400
batch_size = 50
num_rec = X_trainTf.shape[0]
n_batches = int(np.ceil(num_rec / batch_size))
acc_test = 0. # assign the result of accuracy testing to this variable
### START CODE HERE ### (≈ 9-10 lines of code)
# ... variable required for testing acc_test
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
with tf.Session() as sess:
sess.run(init)
for i in range(n_epochs):
X_trainTf_batch, y_trainTf_batch = random_batch(X_trainTf, y_trainTf, batch_size)
sess.run([optimizer, loss], feed_dict={X: X_trainTf_batch, y: y_trainTf_batch})
_, loss, logits = sess.run([optimizer, loss, logits], feed_dict={X: X_testTf, y: y_testTf})
preds = tf.nn.softmax(logits)
correct_preds = tf.equal(tf.argmax(preds, 1), y_testTf)
acc_test = sess.run(tf.reduce_sum(tf.cast(correct_preds, tf.float32))) / len(y_testTf)
### END CODE HERE ###
# -
### GRADED PART (DO NOT EDIT) ###
part5=str(acc_test)
submissions[all_parts[4]]=part5
grading.submit(COURSERA_EMAIL, COURSERA_TOKEN, assignment_key, all_parts[:5],all_parts,submissions)
acc_test
### GRADED PART (DO NOT EDIT) ###
| Bank_failure_m1_ex4_v4 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment: Build a Regression Model in Keras
#
# ## Introduction
#
# This assignment was the final project in IBM's [Introduction to Deep Learning & Neural Networks with Keras](https://www.coursera.org/account/accomplishments/certificate/QQEVGBYCHYYP) course on Coursera. The objective: build a regression model to predict the compressive strength of concrete based on its age (in days) and the volumes of the different ingredients used to make it.
#
# Each part of the assignment involved the following steps:
#
# 1. Randomly split the data into training and test sets, holding 30% for testing.
# 2. Build a neural network with hidden layers of 10 nodes each, using ReLU activation, Adam optimization, and a mean squared error loss function.
# 3. Train the model over a certain number of epochs.
# 4. Evaluate the model on the test data and compute the mean squared error between the predicted strength and actual strength.
# 5. Repeat the previous steps 50 times.
# 6. Report the mean and the standard deviation of the 50 mean squared errors.
#
# The four parts differed as follows in terms of whether or not the input data was normalized, the number of hidden layers, and the number of epochs over which the model was trained:
#
# - **Part A**: raw input data, 1 hidden layer, 50 epochs
# - **Part B**: normalized input data, 1 hidden layer, 50 epochs
# - **Part C**: normalized input data, 1 hidden layer, 100 epochs
# - **Part D**: normalized input data, 3 hidden layers, 50 epochs
# ## Part A
#
# ### Download and clean the data
#
# First step: download the dataset and save it as a pandas DataFrame.
# +
import pandas as pd
concrete_data = pd.read_csv("https://cocl.us/concrete_data")
concrete_data.head()
# -
# Looks good. Next up, splitting between predictors (`X`) and target (`y`). Strength is the target variable.
# +
X = concrete_data[concrete_data.columns[concrete_data.columns != "Strength"]]
num_cols = X.shape[1] # Saving for later
y = concrete_data["Strength"]
# -
X.head()
y.head()
# ### Build the neural network
#
# Next I'll build a function to create the neural network, with the number of hidden layers as a parameter, since Part D uses three hidden layers.
# +
import keras
from keras.models import Sequential
from keras.layers import Dense
def regression_model(num_hidden_layers):
model = Sequential()
# Hidden layers
model.add(Dense(10, activation="relu", input_shape=(num_cols,)))
for i in range(num_hidden_layers - 1):
model.add(Dense(10, activation="relu"))
# Output layer
model.add(Dense(1))
model.compile(optimizer="adam", loss="mean_squared_error")
return model
# -
# ### Split, train, evaluate × 50
#
# Now the fun part. I'm going to do the following 50 times:
#
# - Randomly split the data by holding 30% for testing.
# - Train a model on the training data over 50 epochs.
# - Evaluate the model on the test data and compute the mean squared error between predicted concrete strength and actual concrete strength.
#
# I'll save each of the 50 mean squared errors into a list.
#
# And come to think of it, I'm going to define a function for this process, because I'm going to need to repeat it in all the other parts of the assignment, just varying a few key parameters.
# +
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from matplotlib import pyplot
def process_models(X, y, num_hidden_layers, num_epochs):
mean_squared_errors = []
for i in range(50):
# Split the data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# Train the model
model = regression_model(num_hidden_layers)
hist = model.fit(
X_train,
y_train,
epochs=num_epochs,
verbose=0,
validation_data=(X_test, y_test),
)
# Test the model
predictions = model.predict(X_test)
# Find and save the mean squared error
mean_squared_errors.append(mean_squared_error(y_test, predictions))
print("Run #{} complete".format(i + 1))
# Plot the loss from the last run just for fun
pyplot.plot(hist.history["loss"])
pyplot.plot(hist.history["val_loss"])
pyplot.title("Model loss (run #50)")
pyplot.ylabel("Loss")
pyplot.xlabel("Epoch")
pyplot.legend(["Train", "Test"], loc="upper right")
pyplot.show()
return mean_squared_errors
errors_a = process_models(X, y, 1, 50)
# -
# ### Results
#
# I'll make a function for this part, too.
# +
from statistics import mean, stdev
def report_results(mean_squared_errors):
print(
"The mean of the mean squared errors is {}".format(
round(mean(mean_squared_errors), 3)
)
)
print(
"The standard deviation of the mean squared errors is {}".format(
round(stdev(mean_squared_errors), 3)
)
)
report_results(errors_a)
# -
# There you have it. I'll admit, those errors look pretty bad. I'm curious to see how the changes in the next three parts affect that.
# ## Part B
#
# Same as Part A, but using a normalized version of the data. I'll normalize the predictors, then repeat the split/train/evaluate/repeat
# process, then report the results.
# +
X_norm = (X - X.mean()) / X.std()
errors_b = process_models(X_norm, y, 1, 50)
report_results(errors_b)
# -
# The mean of the mean squared errors is just the slightest bit higher than Part A, probably not significantly so. Curious. The standard deviation is much less, though. I assume the inputs vary less after normalizing?
# ## Part C
#
# Same as Part B, but using 100 epochs for training. Here we go:
errors_c = process_models(X_norm, y, 1, 100)
report_results(errors_c)
# That significantly reduced both the mean _and_ the standard deviation of the mean squared errors compared to Part B. Nice.
# ## Part D
#
# Last but not least, this one's the same as Part B but with _three_ hidden layers (each still with 10 nodes and ReLU activation). Sure am glad I made those functions in Part A.
errors_d = process_models(X_norm, y, 3, 50)
report_results(errors_d)
# This is again a significant improvement over Part B (both the mean and standard deviation of the mean squared errors are much less), but not too much better than Part C. Fascinating.
# ## Reflection
#
# What I'm still wondering the most now is how one goes about designing the size and shape of a neural network. I've gotten good at _building_ a network with the Keras API now if I'm told how many layers to use, how many nodes in each of those layers, how many epochs to train it over. But how do you make those decisions for each new dataset and problem?
#
# That's what I intend to learn next.
| complete_writeup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
def deriv(stateTuple, t, k1, k2):
A = stateTuple[0]
B = stateTuple[1]
C = stateTuple[2]
D = stateTuple[3]
dAdt = -1.0 * (2.0 * k1 * A * A + k2 * A * C)
dBdt = k1 * A * A
dCdt = -1.0 * (k2 * A * C)
dDdt = k2 * A * C
return(dAdt, dBdt, dCdt, dDdt)
k1 = 5.0**-8.0
k2 = 10.0**-7.0
A = 10.0**6.0
B = 0
C = 10.0
D = 0
odemax = 100.0
odemin = 0.0
buckets = 1000
step = (odemax - odemin)/buckets
t = list(np.arange(odemin, odemax, step))
ret = odeint(deriv, (A, B, C, D), t, args=(k1, k2))
print("Done")
# -
plt.style.use('ggplot')
fig = plt.figure(figsize=(12,6))
ax = fig.add_subplot(1, 1, 1)
ax.scatter(t, ret.T[2])
| docs-static/Tat_Model/tests/CrystalizationTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Cross correlation of DTOCs and ED Breaches
import numpy as np
import pandas as pd
from scipy import signal
import matplotlib.pyplot as plt
import seaborn as sns
from statsmodels.tsa.stattools import adfuller
# %matplotlib inline
df_dtoc = pd.read_csv('dtoc.csv')
df_dtoc.shape
df_dtoc.head()
df_dtoc.rename(columns={'total_atten_>4hr':'total_breaches'},inplace=True)
# dickey fuller test of stationarity
#
# Notes: both are ns - this means that they are non-stationary
def check_stationarity(series):
result = adfuller(series)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
check_stationarity(df_dtoc['total_breaches'])
check_stationarity(df_dtoc['dtoc_total'])
# Difference time series
df_dtoc.columns
df_dtoc['total_breaches_diff'] = df_dtoc['total_breaches'].diff()
df_dtoc['dtoc_total_diff'] = df_dtoc['dtoc_total'].diff()
df_dtoc['total_admissions_diff'] = df_dtoc['total_admissions'].diff()
df_dtoc['total_atten_diff'] = df_dtoc['total_atten'].diff()
# drop lost row
df_dtoc.dropna(inplace=True)
check_stationarity(df_dtoc.total_breaches_diff.loc[df_dtoc.total_breaches_diff.index > 0])
check_stationarity(df_dtoc.dtoc_total_diff.loc[df_dtoc.dtoc_total_diff.index > 0])
# Still non-stationary!
# Cross correlation
def bk_cov(df1,df2):
'calc cov from two series'
X = df1.values
Xm = df1.values.sum()/len(X)
Y = df2.values
Ym = df2.values.sum()/len(Y)
cov = ( ((X-Xm)*(Y-Ym))/len(Y)).sum()
return(cov)
def bk_cor(df1,df2):
'cal corr'
cor = bk_cov(df1,df2)/np.sqrt(bk_cov(df1,df1) * bk_cov(df2,df2))
return cor
def bk_crosscorr(df1,df2,lag=0):
return bk_cor(df1[lag:],df2.shift(lag)[lag:])
def calc_crosscorr(df,col1,col2,lags):
'calc various cross corr on df with range of lags'
corrs = [bk_crosscorr(df[col1], df[col2], lag=i) for i in range(lags)]
return corrs
cross_corr_scores = pd.DataFrame()
def compute_crosscorr_mult_inputs(df,yvar,xvars,lags,title=''):
''' make df with lagged cross corrs for multiple vars'''
cross_corr_scores = pd.DataFrame()
for i in xvars:
corrs = calc_crosscorr(df,yvar,i,lags)
cross_corr_scores[i] = corrs
ax = plt.subplot()
cross_corr_scores.plot(ax=ax)
ax.set_ylabel('cross-corr coef')
ax.set_xlabel('lags (months)')
ax.set_title(title)
return
df_dtoc.columns
# cross correlation coeff for various lags for undifferenced
xvars = ['dtoc_total','total_admissions','total_atten']
compute_crosscorr_mult_inputs(df_dtoc,'total_breaches',xvars,36,title ='cross-corr coeff of total_breaches vs...')
xvars = ['dtoc_total_diff','total_admissions_diff','total_atten_diff']
compute_crosscorr_mult_inputs(df_dtoc,'total_breaches',xvars,36, title='cross-corr coef of total_breaches vs...')
xvars = ['dtoc_total_diff','total_admissions_diff','total_atten_diff']
compute_crosscorr_mult_inputs(df_dtoc,'total_breaches_diff',xvars,36,title='cross-corr coef for of total_breaches_diff vs...')
# Autocorrelations
xvars = ['total_breaches']
compute_crosscorr_mult_inputs(df_dtoc,'total_breaches',xvars,36,title='auto-correlation')
xvars = ['total_breaches_diff']
compute_crosscorr_mult_inputs(df_dtoc,'total_breaches_diff',xvars,36,title='auto-correlation')
xvars = ['dtoc_total']
compute_crosscorr_mult_inputs(df_dtoc,'dtoc_total',xvars,36,title='auto-correlation')
xvars = ['dtoc_total_diff']
compute_crosscorr_mult_inputs(df_dtoc,'dtoc_total_diff',xvars,36,title='auto-correlation')
df_dtoc.columns
df_dtoc.plot.scatter('dtoc_total_diff','total_breaches_diff')
df_dtoc.plot.scatter('total_admissions_diff','total_breaches_diff')
# #### Tom stop reading here
break
cross_corr_scores[i] = corrs
cross_corr_scores.plot()
xcor_monthly = [bk_crosscorr(df_dtoc['total_breaches'], df_dtoc['dtoc_total'], lag=i) for i in range(40)]
pd.DataFrame(xcor_monthly).plot()
xcor_monthly = [bk_crosscorr(df_dtoc['total_breaches_diff'], df_dtoc['dtoc_total_diff'], lag=i) for i in range(40)]
xcor_monthly
pd.DataFrame(xcor_monthly).plot()
corr = signal.correlate(breaches_diff.loc[breaches_diff.index > 0], dtoc_total_diff.loc[dtoc_total_diff.index > 0], mode='same')
#corr = corr/np.max(corr)
corr
fig, (ax_breaches, ax_dtocs) = plt.subplots(2, 1, sharex=True)
ax_breaches.plot(breaches_diff.loc[breaches_diff.index > 0])
ax_breaches.set_title('Breaches')
ax_dtocs.plot(dtoc_total_diff.loc[dtoc_total_diff.index > 0])
ax_dtocs.set_title('DToCs')
ax_breaches.margins(0, 0.1)
fig.tight_layout()
fig, ax_corr = plt.subplots(1, 1, sharex=True)
ax_corr.plot(corr)
ax_corr.set_title('Cross-correlation')
ax_corr.set_xlabel('lag')
ax_corr.axhline(0.0, ls=':')
# Notes:
# Might need to take 2nd difference.
# Note sure what you do about interpretation of significance?
# # Dev
# +
a = [1,2,-2,4,2,3,1,0]
b = [2,3,-2,3,2,4,1,-1]
c = [-2,0,4,0,1,1,0,-2]
data = {'a':a,'b':b,'c':c}
# -
import pandas as pd
# %matplotlib inline
df = pd.DataFrame(data)
df
df.plot()
df.corr()
df.corr('spearman')
len(df.a)
def bk_cov(df1,df2):
'calc cov from two series'
X = df1.values
Xm = df1.values.sum()/len(X)
Y = df2.values
Ym = df2.values.sum()/len(Y)
cov = ( ((X-Xm)*(Y-Ym))/len(Y)).sum()
return(cov)
bk_cov(df.a,df.b)
bk_cov(df.b,df.a)
np.cov(df.a,df.b)
def bk_cor(df1,df2):
'cal corr'
cor = bk_cov(df1,df2)/np.sqrt(bk_cov(df1,df1) * bk_cov(df2,df2))
return cor
bk_cor(df.b,df.c)
bk_cor(df.a,df.c)
np.corrcoef(df,rowvar=0)
def bk_crosscorr(df1,df2,lag=0):
return bk_cor(df1[lag:],df2.shift(lag)[lag:])
df.a.shift(1)[1:]
bk_crosscorr(df.a,df.c)
# on DTOC data
df_dtoc.shape
df_dtoc.dropna().shape
df_dtoc.columns
bk_crosscorr(df_dtoc['total_atten_>4hr'],df_dtoc['dtoc_total'])
xcor_monthly = [bk_crosscorr(df_dtoc['total_atten_>4hr'], df_dtoc['dtoc_total'], lag=i) for i in range(40)]
pd.DataFrame(xcor_monthly).plot()
pd.DataFrame(xcor_monthly).plot()
from scipy import signal
corr = signal.correlate(df.a.values,df.b.values, mode='full',method='direct')
#corr = corr/np.max(corr)
corr
import numpy as np
np.correlate(df.a,df.b,mode='valid')
def bk_cov2(df1,df2):
'calc cov from two series'
X = df1.values
Xm = df1.values.sum()/len(X)
Y = df2.values
Ym = df2.values.sum()/len(Y)
cov = (((X-Xm)*(Y-Ym))/len(Y))#.sum()
return(cov)
bk_cov2(df.a,df.b)
np.corrcoef(df,rowvar=0)
np.corrcoef(df.a,df.b,rowvar=0)
np.cov(df.a,df.b)/(np.sqrt(np.cov(df.a,df.b)*np.cov(df.a,df.a)))
E(XY)−E(X)E(Y)
df.a.values
df.a.values * df.a.values
df.a.sum()
import scipy as sp
sp.corrcoef()
sp.correlate()
# # random noise checks
i = 400
import pandas as pd
from numpy.random import random_sample
# make random noise in 2 time signals
rand = pd.DataFrame({'a': random_sample(i),'b':random_sample(i)})
rand.plot()
rand.corr()
# obivously low correlations
# add trend to each time series
rand['a_trend'] = rand.a + rand.index/100
rand['b_trend'] = rand.b + rand.index/50
rand[['a_trend','b_trend']].plot()
rand.corr()
# correaltions huge!
# difference time series
rand['a_trend_diff'] = rand.a_trend.diff()
rand['b_trend_diff'] = rand.b_trend.diff()
rand.corr()
# correlations back to reasonable level
| dtoc_cross_correlation-bk_version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pokemon = {
'nome': 'Squirtle',
'tipo': 'água',
'hp': 150
}
class Contador():
def __init__(self):
self.valor = 0
def conta(self):
self.valor += 1
return self.valor
def status(self):
return self.valor
def compara(self, outro_contador):
if self.valor > outro_contador.status():
return 'maior'
elif self.valor == outro_contador.status():
return 'igual'
else:
return 'menor'
contador1 = Contador()
contador1
contador1.conta()
contador1.status()
contador2 = Contador()
contador2.conta()
contador1.compara(contador2)
class Pokemon():
def __init__(self, nome_para_construcao, tipo, hp):
self.nome = nome_para_construcao
self.tipo = tipo
self.hp = hp
def aumentar_hp(self, hp):
self.hp += hp
pikachu = Pokemon('Pikachu', 'água', 100)
pikachu
pikachu.set_nome('Charmander')
pikachu.nome
class ArCondicionado():
def __init__(self):
self.temperatura = 18
self.ligado = False
def ligar(self):
if not self.ligado:
self.ligado = True
else:
return 'Já está ligado'
def desligar(self):
if self.ligado:
self.ligado = False
else:
return 'Já está desligado'
sala_4_ac = ArCondicionado()
sala_4_ac.ligado
sala_4_ac.ligar()
sala_4_ac.desligar()
# +
class Pokemon():
def __init__(self, nome_para_construcao, tipo, hp):
self.nome = nome_para_construcao
self.tipo = tipo
self.hp = hp
def aumentar_hp(self, hp):
self.hp += hp
def __repr__(self):
return '<{}, hp: {}>'.format(self.nome, self.hp)
class Treinador():
# Toda vez que usamos argumentos opcionais (com valores default)
# tudo à direita deve ter um valor default também
def __init__(self, nome = 'Ash', cidade = 'Pallet'):
self.nome = nome
self.cidade = cidade
self.pokemons = []
self.storage = []
def capturar(self, pokemon_selvagem):
if not isinstance(pokemon_selvagem, Pokemon):
return 'Não é um pokémon!'
elif len(self.pokemons) == 6:
return 'Não tem mais espaço. Transfira primeiro'
else:
self.pokemons.append(pokemon_selvagem)
return 'Capturado!'
def remover(self, nome_pokemon):
para_transferencia = [] # essa lista guarda todos os pokes para transferir
# Buscando todos os pokemons que serão transferidos
for pokemon in self.pokemons:
if pokemon.nome == nome_pokemon:
para_transferencia.append(pokemon)
# Para cada pokemon que será transferido,
# Adicionamos no storage e removemos da lista principal
for poke_to_transfer in para_transferencia:
self.storage.append(poke_to_transfer)
self.pokemons.remove(poke_to_transfer)
if len(para_transferencia) == 0:
return 'Nenhum pokémon encontrado com esse nome'
else:
return '{} pokemons transferidos!'.format(len(para_transferencia))
# -
ash = Treinador()
ash.nome
# Ele não pode capturar algo que não seja um pokémon!
ash.capturar('Pikachu')
pikachu = Pokemon('Pikachu', 'Elétrico', 150)
ash.capturar(pikachu)
ash.capturar(pikachu)
ash.capturar(pikachu)
ash.capturar(pikachu)
ash.capturar(pikachu)
ash.capturar(pikachu)
ash.pokemons.remove(pikachu)
ash.capturar(pikachu)
ash.remover('Pikachu')
ash.pokemons
ash.storage
ash.pokemons[1].nome = 'Charmander'
ash.pokemons[0].nome
ash.pokemons[1].nome
| Aula 10/Aula 10 - POO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Serbeld/Dataset6C/blob/master/Data_6C.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lxz0Gu3LwQbc" colab_type="text"
# #**Dataset 6C**
# + id="8_msq58NtLmO" colab_type="code" colab={}
#Loading the dataset
# !pip install h5py
import h5py
from google.colab import drive,files
drive.mount('/content/drive')
hdf5_path = '/content/drive/My Drive/Dataset6C/Dataset6C.hdf5'
dataset = h5py.File(hdf5_path, "r")
# + [markdown] id="14tyniVq0FjY" colab_type="text"
# #Exploratory Data Analysis
# + id="wwLgSR6n0Es5" colab_type="code" colab={}
import numpy as np
import matplotlib.pylab as plt
#train
train_img = dataset["train_img"]
xt = np.array(train_img)
yt = np.array(dataset["train_labels"])
#test
xtest = np.array(dataset["test_img"])
ytest = np.array(dataset["test_labels"])
#Validation
xval = np.array(dataset["val_img"])
yval = np.array(dataset["val_labels"])
# + id="V8-EqI6f0o4A" colab_type="code" outputId="cf906d74-226e-48a0-a4a0-25dd35c882db" colab={"base_uri": "https://localhost:8080/", "height": 69}
print("Training Shape: "+ str(xt.shape))
print("Validation Shape: "+ str(xval.shape))
print("Testing Shape: "+ str(xtest.shape))
# + id="o-VEo5xe11X2" colab_type="code" outputId="9fc620c4-8e61-4f54-8c55-8fea65175f65" colab={"base_uri": "https://localhost:8080/", "height": 425}
#Image
num_image = 15
print()
print('Healthy: 0')
print('Pneumonia & Covid-19: 1')
print('Cardiomegaly: 2')
print('Other respiratory disease: 3')
print('Pleural Effusion: 4')
print('Infiltration: 5')
print()
print("Output: "+ str(yt[num_image]))
imagen = train_img[num_image]
plt.imshow(imagen)
plt.show()
# + id="AZLUDOWa0Xul" colab_type="code" outputId="829ef654-6a99-49c9-e8d5-11fa4ef34d14" colab={"base_uri": "https://localhost:8080/", "height": 156}
#Categorical values or OneHot
import keras
num_classes = 6
yt = keras.utils.to_categorical(yt,num_classes)
ytest = keras.utils.to_categorical(ytest,num_classes)
yval = keras.utils.to_categorical(yval,num_classes)
print()
print('Healthy: [1 0 0 0 0 0]')
print('Pneumonia & Covid-19: [0 1 0 0 0 0]')
print('Cardiomegaly: [0 0 1 0 0 0]')
print('Other respiratory disease: [0 0 0 1 0 0]')
print('Pleural Effusion: [0 0 0 0 1 0]')
print('Infiltration: [0 0 0 0 0 1]')
# + id="xT0AJz8688OS" colab_type="code" outputId="18fbecbf-e1dc-4fff-b57b-ba7fc1c565de" colab={"base_uri": "https://localhost:8080/", "height": 425}
#Image
num_image = 15
print()
print('Healthy: [1. 0. 0. 0. 0. 0.]')
print('Pneumonia & Covid-19: [0. 1. 0. 0. 0. 0.]')
print('Cardiomegaly: [0. 0. 1. 0. 0. 0.]')
print('Other respiratory disease: [0. 0. 0. 1. 0. 0.]')
print('Pleural Effusion: [0. 0. 0. 0. 1. 0.]')
print('Infiltration: [0. 0. 0. 0. 0. 1.]')
print()
print("Output: "+ str(yt[num_image]))
imagen = train_img[num_image]
plt.imshow(imagen)
plt.show()
| Data6C/Data_6C.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pymongo
db=pymongo.MongoClient()['crypto']
certs=db['certs']
count_rsa=certs.count_documents({'rsa_PEM':{"$exists":True}})
count_rsa
from collections import Counter
bits_list=[ doc.get('bits',-1) for doc in
certs.find({'rsa_PEM':{"$exists":True}},{"bits":1})]
{k:(v,round(100*v/count_rsa,1)) for k,v in Counter(bits_list).items()}
keyHashes=certs.distinct('keyHash')
len(keyHashes)
certs.estimated_document_count()-len(keyHashes)
certs.find_one().keys()
# %%time
susp=[]
for k in keyHashes:
docs=list(certs.find({'keyHash':k}))
if len(docs)>1:
susp.append(docs)
# +
from itertools import combinations
def is_duplicate_urls(lst):
urls=[d['url'][8:].replace('www.','') for d in lst]
return all([(f in s or s in f) for f,s in combinations(urls,2)])
dedup_url=[s for s in susp if not is_duplicate_urls(s)]
# -
len(dedup_url)
def is_duplicate_urls2(lst):
urls=[set(d['url'][8:].replace('www.','').replace('.pl','').split('.')) for d in lst]
return any([bool(f.intersection(s)) for f,s in combinations(urls,2)])
dedup_url2=[s for s in dedup_url if not is_duplicate_urls2(s)]
len(dedup_url2)
import pandas as pd
df=pd.Series(map(len,dedup_url2))
ax=df.hist(bins=8,range=(2,11),log=True)
ax.set_title('Hist of true duplicates - log')
print('\n\n'.join([' '.join([dd['url'][8:] for dd in d]) for d in dedup_url2 if len(d)>10]))
| Zad3/stats_certs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Imports
import os
import numpy as np
import xarray as xr
# +
def load_ndwi(prod, res):
"""
Load NDWI index (and rename the array)
"""
# Read NDWI index
ndwi = prod.load(NDWI)[NDWI]
ndwi_name = f"{ndwi.attrs['sensor']} NDWI"
return ndwi.rename(ndwi_name)
def extract_water(ndwi):
"""
Extract water from NDWI index (and rename the array)
"""
# Assert water bodies when NDWI index > 0.2
water = xr.where(ndwi > 0.2, 1, 0)
# Set nodata where ndwi is nan.
# WARNING: the function xr.DataArray.where sets by default np.nan where the condition is false !
# See here: http://xarray.pydata.org/en/stable/generated/xarray.DataArray.where.html
water = water.where(~np.isnan(ndwi))
# Plot a subsampled version
water_name = f"{ndwi.attrs['sensor']} WATER"
water = water.rename(water_name)
water.attrs["long_name"] = "Water detection"
return water.rename(water_name)
# +
import logging
logger = logging.getLogger("eoreader")
logger.setLevel(logging.DEBUG)
# create console handler and set level to debug
ch = logging.StreamHandler()
ch.setLevel(logging.DEBUG)
# create formatter
formatter = logging.Formatter('%(message)s')
# add formatter to ch
ch.setFormatter(formatter)
# add ch to logger
logger.addHandler(ch)
# -
prod_folder = os.path.join("/home", "data", "DS3", "CI", "extracteo", "water")
paths = [
# Landsat-8 OLCI collection 2
os.path.join(prod_folder, "LC08_L1TP_200030_20201220_20210310_02_T1.tar"),
# Landsat-5 TM collection 2
os.path.join(prod_folder, "LT05_L1TP_200030_20111110_20200820_02_T1.tar"),
# Sentinel-2 L2A
os.path.join(prod_folder, "S2A_MSIL2A_20191215T110441_N0213_R094_T30TXP_20191215T122756.zip"),
]
# +
from eoreader.reader import Reader
from eoreader.bands import *
# Create the reader
eoreader = Reader()
# Loop on all the products
water_arrays = []
ndwi_arrays = []
extents = []
for path in paths:
logger.info(f"*** {os.path.basename(path)} ***")
# Open the product
prod = eoreader.open(path, remove_tmp=True)
# Get extents
extents.append(prod.extent)
# Read NDWI index
# 6Let's say we want a 60. meters resolution
ndwi = load_ndwi(prod, res=60.)
ndwi_arrays.append(ndwi)
# Extract water
water_arrays.append(extract_water(ndwi))
logger.info("\n")
# +
# Plot the tiles
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import cartopy.crs as ccrs
from shapely.errors import ShapelyDeprecationWarning
import warnings
warnings.filterwarnings("ignore", category=ShapelyDeprecationWarning)
nrows = len(paths)
plt.figure(figsize=(6 * nrows, 6 * nrows))
cmap = ListedColormap(['khaki', 'lightblue'])
for i in range(nrows):
# Compute cartopy projection (EOReader always is in UTM)
extent = extents[i]
str_epsg = str(extent.crs.to_epsg())
zone = str_epsg[-2:]
is_south = str_epsg[2] == 7
proj = ccrs.UTM(zone, is_south)
# Get extent values
# The extents must be defined in the form (min_x, max_x, min_y, max_y)
bounds = extent.bounds
extent_val = [bounds.minx[0], bounds.maxx[0], bounds.miny[0], bounds.maxy[0]]
# Plot NDWI
axes = plt.subplot(nrows, 2, 2*i+1, projection=proj)
axes.set_extent(extent_val, proj)
ndwi_arrays[i][0, fc00:e968:6179::de52:7100, ::10].plot.imshow(origin='upper', extent=extent_val, transform=proj, robust=True)
axes.coastlines(linewidth=1)
axes.set_title(ndwi_arrays[i].name)
# Plot water
axes = plt.subplot(nrows, 2, 2*i+2, projection=proj)
axes.set_extent(extent_val, proj)
water_arrays[i][0, fc00:e968:6179::de52:7100, ::10].plot.imshow(origin='upper', extent=extent_val, transform=proj, cmap=cmap, cbar_kwargs={'ticks': [0, 1]})
axes.coastlines(linewidth=1)
axes.set_title(water_arrays[i].name)
axes.set_title(water_arrays[i].name)
| docs/_build/.jupyter_cache/executed/5a35244f37b8ee189df93f714a0470e9/base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ## 1
# 1. 소수를 작은 순서대로 5개를 뽑아 각 행의 1열로 가지는 5×5 행렬 생성
# 2. 이 행렬의 다음 열들은 1열의 배수(1행:[2, 4, 6, 8, 10], 2행:[3, 6, …, 15])라고 할 때, 이를 Numpy 배열로 나타내고
# 3. 이 행렬의 m행 m열 (m = 1, 2,…, 5)의 원소들의 합과 평균을 구한다.
prime_list = [2, 3, 5, 7, 11]
# 소수(prime number)를 기반으로 2차원 배열 생성
matrix = np.array([[prime_list[x] * i for i in range(1,6)] for x in range(len(prime_list))])
matrix
# ### np.diag
# 배열 matrix의 대각(diagonal) 성분을 추출
matrix_dia = np.diag(matrix)
matrix_dia
# ### np.trace
# 대각 성분의 합과 평균
dia_sum = np.trace(matrix)
dia_mean = np.mean(matrix_dia)
print(f'대각 성분의 합 :{dia_sum}, 대각 성분의 평균 :{dia_mean}')
# ---
# ## 2
# 1. 배열 x 의 1 열을 추출하여 하나의 배열로 만든다.
# 2. 배열 x 의 마지막 행을 추출하여 하나의 배열로 만든다.
# 3. 배열 x의 1열에 대해서 그 평균을 구한다.
# 4. 배열 x 의 마지막 행에 대해서 그 평균을 구한다.
def np_array():
x = np.array([[1, 572, 2, 219, 31, 6, 7, 8, 24, 10],
[12, 222, 33, 1, 2, 3, 99, 24, 1, 42],
[623, 2, 3, 56, 5, 2, 7, 85, 22, 110],
[63, 24, 3, 4, 5, 64, 7, 82, 93, 210],
[48, 28, 3, 24, 57, 63, 7, 8, 9, 102],
[333, 64, 3, 44, 5, 6, 72, 82, 3, 10],
[52, 242, 2, 41, 52, 6, 32, 8, 96, 2],
[33, 223, 52, 4, 35, 62, 7, 8, 9, 10],
[29, 2, 3, 149, 15, 6, 172, 2, 2, 11],
[94, 23, 32, 24, 54, 63, 1, 3, 92, 7]])
# 배열 x 의 1 열을 추출하여 하나의 배열로 만든다.
firstcol_x = x[:,0]
# 배열 x 의 마지막 행을 추출하여 하나의 배열로 만든다.
lastrow_x = x[-1]
# 배열 x의 1열에 대해서 그 평균을 구한다.
mean_firstcol = np.mean(firstcol_x)
# 배열 x 의 마지막 행에 대해서 그 평균을 구한다.
mean_lastrow = np.mean(lastrow_x)
return [firstcol_x, lastrow_x, mean_firstcol, mean_lastrow]
# +
#출력을 위한 함수
def print_output(**kwargs):
for key in kwargs.keys():
print(key,":", kwargs[key])
firstcol_x, lastrow_x, mean_firstcol, mean_lastrow = np_array()
print_output(firstcol_x=firstcol_x, lastrow_x=lastrow_x, mean_firstcol=mean_firstcol, mean_lastrow=mean_lastrow)
# -
| practice/numpy_matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Teapod - Intro to Python Programming
#Dictionaries
# -
# # Setup
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# +
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# -
# # Basics of Python
# +
#A dictionary is a data type similar to arrays, but works with keys and values instead of indexes.
#Each value stored in a dictionary can be accessed using a key, which is any type of object (a string, a number, a list, etc.)
#instead of using its index to address it.
phonebook = {}
phonebook["John"] = 938477566
phonebook["Jack"] = 938377264
phonebook["Jill"] = 947662781
print(phonebook)
# -
phonebook = {
"John" : 938477566,
"Jack" : 938377264,
"Jill" : 947662781
}
print(phonebook)
# +
#iterating over dictionaries
phonebook = {"John" : 938477566,"Jack" : 938377264,"Jill" : 947662781}
for name, number in phonebook.items():
print("Phone number of %s is %d" % (name, number))
# +
#removing a value
phonebook = {
"John" : 938477566,
"Jack" : 938377264,
"Jill" : 947662781
}
del phonebook["John"]
print(phonebook)
# +
#or
phonebook = {
"John" : 938477566,
"Jack" : 938377264,
"Jill" : 947662781
}
phonebook.pop("John")
print(phonebook)
# +
#Exercise
#Add "Jake" to the phonebook with the phone number 938273443, and remove Jill from the phonebook.
phonebook = {
"John" : 938477566,
"Jack" : 938377264,
"Jill" : 947662781
}
# write your code here
# testing code
if "Jake" in phonebook:
print("Jake is listed in the phonebook.")
if "Jill" not in phonebook:
print("Jill is not listed in the phonebook.")
# -
| 6. Python - Dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
#
# ### a) Data Mining Research (2 parts: LA and State of California)
#
# #### Part-I: Total of 10 Datasets for LA County:
#
# 1) LA children and family services (with zipcode, lanlon):
#
# https://geohub.lacity.org/datasets/lacounty::children-and-family-services
#
# 2) LA Health screening and testing (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::health-screening-and-testing-1
#
# 3) LA Health Education and Counseling (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::health-education-and-counseling-1
#
# 4) LA Disability Support Services (with latlon, no zipcode) :
#
# https://geohub.lacity.org/datasets/lacounty::disability-support-services-1
#
# 5) LA Mental Health Counseling (wiht zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::mental-health-counseling-1
#
# 6) LA Substance Abuse Program (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::substance-abuse-programs-1
#
# 7) LA Crime Prevention and Support (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::crime-prevention-and-support-1
#
# 8) LA Crime Reporting and Investigating (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::crime-reporting-and-investigation
#
# 9) LA Domestic Violence (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::domestic-violence-services-1
#
# 10) LA Mental Health Centers (with zipcode, latlon):
#
# https://geohub.lacity.org/datasets/lacounty::mental-health-centers-1
#
#
#
#
# #### Part-II: Total of 6 Datasets for State of California
#
# 1) County Expenditures (with county+zipcode):
#
# https://data.ca.gov/dataset/county-expenditures/resource/45f8ebcb-a759-4c9c-aed4-7e446acb1f8c
#
# 2) Total Construction Cost of Healthcare Projects (with county):
#
# https://data.chhs.ca.gov/dataset/total-construction-cost-of-healthcare-projects
#
# 3) Medically underserved areas (with shapeAreaLen):
#
# https://data.chhs.ca.gov/dataset/medically-underserved-areas/resource/75388087-5bb2-4520-b6c1-dce1832b6441
#
# 4) Medically Underserved Populations (with shapeAreaLen):
#
# https://data.chhs.ca.gov/dataset/medically-underserved-populations/resource/dc197613-da51-4192-bfab-7b67cac43410
#
# 5) Health Professional Shortage Area Primary Care (with county + shapeAreaLen):
#
# https://data.chhs.ca.gov/dataset/health-professional-shortage-area-primary-care/resource/7e693a27-3a40-4804-bd21-5a676ebe1196
#
# 6) Health Professional Shortage Area Mental Health (with county + shapeAreaLen):
#
# https://data.chhs.ca.gov/dataset/health-professional-shortage-area-mental-health/resource/a591c975-f15f-48c9-b9c8-489971a2a1ce
#
#
| dataAnalysis/LAHealthcareProject/2. Healthcare_DataMiningResearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import networkit as nk
from percolate.two_d_percolation import Perc
p = Perc(3, 4)
core_dec = nk.centrality.CoreDecomposition(p.g)
core_dec.run()
set(core_dec.scores())
nk.viztasks.drawGraph(p.g)
# -
nk.viztasks.drawGraph(p.g, node_size=[(k**2)*20 for k in core_dec.scores()])
plt.show()
import matplotlib.pyplot as plt
plt.ion()
p.plot()
| percolate/notebooks/percolation_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
n1 = int(input('Digite o primeiro valor: '))
n2 = int(input('Digite o segundo valor: '))
if n1 > n2:
print('O menor número é o {}\nE o maior número é o {}.' .format(n2, n1))
elif n1 < n2:
print('O menor número é o {}\nE o maior número é o {}.' .format(n1, n2))
else:
print('Os número são iguais.')
| desaf033.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # 6.4 循环神经网络的从零开始实现
# +
import time
import math
import numpy as np
import torch
from torch import nn, optim
import torch.nn.functional as F
import sys
sys.path.append("..")
import d2lzh_pytorch as d2l
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(torch.__version__)
print(device)
# -
(corpus_indices, char_to_idx, idx_to_char, vocab_size) = d2l.load_data_jay_lyrics()
# ## 6.4.1 one-hot向量
# +
def one_hot(x, n_class, dtype=torch.float32):
# X shape: (batch), output shape: (batch, n_class)
x = x.long()
res = torch.zeros(x.shape[0], n_class, dtype=dtype, device=x.device)
res.scatter_(1, x.view(-1, 1), 1)
return res
x = torch.tensor([0, 2])
one_hot(x, vocab_size)
# +
# 本函数已保存在d2lzh_pytorch包中方便以后使用
def to_onehot(X, n_class):
# X shape: (batch, seq_len), output: seq_len elements of (batch, n_class)
return [one_hot(X[:, i], n_class) for i in range(X.shape[1])]
X = torch.arange(10).view(2, 5)
inputs = to_onehot(X, vocab_size)
print(len(inputs), inputs[0].shape)
# -
# ## 6.4.2 初始化模型参数
# +
num_inputs, num_hiddens, num_outputs = vocab_size, 256, vocab_size
print('will use', device)
def get_params():
def _one(shape):
ts = torch.tensor(np.random.normal(0, 0.01, size=shape), device=device, dtype=torch.float32)
return torch.nn.Parameter(ts, requires_grad=True)
# 隐藏层参数
W_xh = _one((num_inputs, num_hiddens))
W_hh = _one((num_hiddens, num_hiddens))
b_h = torch.nn.Parameter(torch.zeros(num_hiddens, device=device, requires_grad=True))
# 输出层参数
W_hq = _one((num_hiddens, num_outputs))
b_q = torch.nn.Parameter(torch.zeros(num_outputs, device=device, requires_grad=True))
return nn.ParameterList([W_xh, W_hh, b_h, W_hq, b_q])
# -
# ## 6.4.3 定义模型
def init_rnn_state(batch_size, num_hiddens, device):
return (torch.zeros((batch_size, num_hiddens), device=device), )
def rnn(inputs, state, params):
# inputs和outputs皆为num_steps个形状为(batch_size, vocab_size)的矩阵
W_xh, W_hh, b_h, W_hq, b_q = params
H, = state
outputs = []
for X in inputs:
H = torch.tanh(torch.matmul(X, W_xh) + torch.matmul(H, W_hh) + b_h)
Y = torch.matmul(H, W_hq) + b_q
outputs.append(Y)
return outputs, (H,)
state = init_rnn_state(X.shape[0], num_hiddens, device)
inputs = to_onehot(X.to(device), vocab_size)
params = get_params()
outputs, state_new = rnn(inputs, state, params)
print(len(outputs), outputs[0].shape, state_new[0].shape)
# ## 6.4.4 定义预测函数
# 本函数已保存在d2lzh_pytorch包中方便以后使用
def predict_rnn(prefix, num_chars, rnn, params, init_rnn_state,
num_hiddens, vocab_size, device, idx_to_char, char_to_idx):
state = init_rnn_state(1, num_hiddens, device)
output = [char_to_idx[prefix[0]]]
for t in range(num_chars + len(prefix) - 1):
# 将上一时间步的输出作为当前时间步的输入
X = to_onehot(torch.tensor([[output[-1]]], device=device), vocab_size)
# 计算输出和更新隐藏状态
(Y, state) = rnn(X, state, params)
# 下一个时间步的输入是prefix里的字符或者当前的最佳预测字符
if t < len(prefix) - 1:
output.append(char_to_idx[prefix[t + 1]])
else:
output.append(int(Y[0].argmax(dim=1).item()))
return ''.join([idx_to_char[i] for i in output])
predict_rnn('分开', 10, rnn, params, init_rnn_state, num_hiddens, vocab_size,
device, idx_to_char, char_to_idx)
# ## 6.4.5 裁剪梯度
# 本函数已保存在d2lzh_pytorch包中方便以后使用
def grad_clipping(params, theta, device):
norm = torch.tensor([0.0], device=device)
for param in params:
norm += (param.grad.data ** 2).sum()
norm = norm.sqrt().item()
if norm > theta:
for param in params:
param.grad.data *= (theta / norm)
# ## 6.4.6 困惑度
# ## 6.4.7 定义模型训练函数
# 本函数已保存在d2lzh_pytorch包中方便以后使用
def train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, device, corpus_indices, idx_to_char,
char_to_idx, is_random_iter, num_epochs, num_steps,
lr, clipping_theta, batch_size, pred_period,
pred_len, prefixes):
if is_random_iter:
data_iter_fn = d2l.data_iter_random
else:
data_iter_fn = d2l.data_iter_consecutive
params = get_params()
loss = nn.CrossEntropyLoss()
for epoch in range(num_epochs):
if not is_random_iter: # 如使用相邻采样,在epoch开始时初始化隐藏状态
state = init_rnn_state(batch_size, num_hiddens, device)
l_sum, n, start = 0.0, 0, time.time()
data_iter = data_iter_fn(corpus_indices, batch_size, num_steps, device)
for X, Y in data_iter:
if is_random_iter: # 如使用随机采样,在每个小批量更新前初始化隐藏状态
state = init_rnn_state(batch_size, num_hiddens, device)
else: # 否则需要使用detach函数从计算图分离隐藏状态
for s in state:
s.detach_()
inputs = to_onehot(X, vocab_size)
# outputs有num_steps个形状为(batch_size, vocab_size)的矩阵
(outputs, state) = rnn(inputs, state, params)
# 拼接之后形状为(num_steps * batch_size, vocab_size)
outputs = torch.cat(outputs, dim=0)
# Y的形状是(batch_size, num_steps),转置后再变成长度为
# batch * num_steps 的向量,这样跟输出的行一一对应
y = torch.transpose(Y, 0, 1).contiguous().view(-1)
# 使用交叉熵损失计算平均分类误差
l = loss(outputs, y.long())
# 梯度清0
if params[0].grad is not None:
for param in params:
param.grad.data.zero_()
l.backward()
grad_clipping(params, clipping_theta, device) # 裁剪梯度
d2l.sgd(params, lr, 1) # 因为误差已经取过均值,梯度不用再做平均
l_sum += l.item() * y.shape[0]
n += y.shape[0]
if (epoch + 1) % pred_period == 0:
print('epoch %d, perplexity %f, time %.2f sec' % (
epoch + 1, math.exp(l_sum / n), time.time() - start))
for prefix in prefixes:
print(' -', predict_rnn(prefix, pred_len, rnn, params, init_rnn_state,
num_hiddens, vocab_size, device, idx_to_char, char_to_idx))
# ## 6.4.8 训练模型并创作歌词
num_epochs, num_steps, batch_size, lr, clipping_theta = 250, 35, 32, 1e2, 1e-2
pred_period, pred_len, prefixes = 50, 50, ['分开', '不分开']
train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, device, corpus_indices, idx_to_char,
char_to_idx, True, num_epochs, num_steps, lr,
clipping_theta, batch_size, pred_period, pred_len,
prefixes)
train_and_predict_rnn(rnn, get_params, init_rnn_state, num_hiddens,
vocab_size, device, corpus_indices, idx_to_char,
char_to_idx, False, num_epochs, num_steps, lr,
clipping_theta, batch_size, pred_period, pred_len,
prefixes)
| code/chapter06_RNN/6.4_rnn-scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import qiskit as q
# from qiskit.visualization import plot_histogram, plot_bloch_multivector
# from matplotlib import style
# import math
# style.use("dark_background")
# # %matplotlib inline
# qasm_sim = q.Aer.get_backend("qasm_simulator")
# statevec_sim = q.Aer.get_backend("statevector_simulator")
# # Uncertain Qubits
# Uncertain qubits don't collapse on same value
c = q.QuantumCircuit(2, 2)
c.ry(math.pi/4, 0)
c.ry(math.pi/4, 1)
orig_statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw()
plot_bloch_multivector(orig_statevec)
orig_counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([orig_counts], legend = ['counts'])
# # Uncertain Qubits, Hadamards at front
c = q.QuantumCircuit(2, 2)
# Putting qubits in superposition
c.h(0)
c.h(1)
c.ry(math.pi/4, 0)
c.ry(math.pi/4, 1)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw()
plot_bloch_multivector(statevec)
counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([counts], legend = ['counts'])
# # Hadamard Sandwich
c = q.QuantumCircuit(2, 2)
# Putting qubits in superposition
c.h(0)
c.h(1)
c.ry(math.pi/4, 0)
c.ry(math.pi/4, 1)
c.h(0)
c.h(1)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw()
plot_bloch_multivector(statevec)
counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([counts], legend = ['counts'])
# # Certain Qubits
c = q.QuantumCircuit(2, 2)
c.x(0)
c.x(1)
orig_statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw()
plot_bloch_multivector(orig_statevec)
# Collapses to |1> always
orig_counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([orig_counts], legend = ['counts'])
# Distribution shows us we always get 11.
# # Hadamard in front of certain qubits
c = q.QuantumCircuit(2, 2)
# Both put to Superposition
c.h(0)
c.h(1)
c.x(0)
c.x(1)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw(output='mpl')
plot_bloch_multivector(statevec)
counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([counts], legend = ['counts'])
# Equal Distribution
# # Hadamard sandwich for certain qubits
c = q.QuantumCircuit(2, 2)
# Both put to Superposition
c.h(0) # hadamard
c.h(1) # hadamard
c.x(0)
c.x(1)
c.h(0) # hadamard
c.h(1) # hadamard
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
c.measure([0, 1], [0, 1])
c.draw(output='mpl')
plot_bloch_multivector(statevec)
# Collapses to |0> always
orig_counts = q.execute(c, backend = qasm_sim, shots = 1024).result().get_counts()
plot_histogram([orig_counts], legend = ['counts'])
# Distribution shows us we always get 00.
# # Deutsch Jozsa Algorithm
# Now given a string of n bits passing through a black box determine whether the black box is constant or balanced.
# Main moto is not finding the nature of the black box but the fact that how can all possible inputs be processed in just 1 pass.
# 0,0 means blackbox is constant 1,1 means blackbox is balanced
# +
def balanced_black_box(c):
c.cx(0, 2)
c.cx(1, 2)
return c
def constant_black_box(c):
return c
# + active=""
# c = q.QuantumCircuit(3, 2) # We don't care about the 3rd qubit
# c = balanced_black_box(c)
# c.draw()
# -
# # Hadamard Sandwich
c = q.QuantumCircuit(3, 2) # We don't care about the 3rd qubit
c.h(0)
c.h(1)
c.h(2)
c = balanced_black_box(c)
c.h(0)
c.h(1)
#c.h(2)
c.draw()
# Creating a balanced blackbox
c = q.QuantumCircuit(3, 2) # We don't care about the 3rd qubit
c.x(2)
c.barrier()
c.h(0)
c.h(1)
c.h(2)
c.barrier()
c = balanced_black_box(c)
c.barrier()
c.h(0)
c.h(1)
#c.h(2)
c.measure([0, 1], [0, 1])
c.draw(output = 'mpl')
counts = q.execute(c, backend = qasm_sim, shots = 1).result().get_counts()
plot_histogram([counts], legend = ['counts'])
# 1,1 means its a balanced circuit
# Creating a constant blackbox
c = q.QuantumCircuit(3, 2) # We don't care about the 3rd qubit
c.x(2)
c.barrier()
c.h(0)
c.h(1)
c.h(2)
c.barrier()
c = constant_black_box(c)
c.barrier()
c.h(0)
c.h(1)
#c.h(2)
c.measure([0, 1], [0, 1])
c.draw(output = 'mpl')
counts = q.execute(c, backend = qasm_sim, shots = 1).result().get_counts()
plot_histogram([counts], legend = ['counts'])
# 0,0 means its a balanced circuit
# + active=""
# It's not the fact that we guessed it in one shot but the fact that how did we do it.
# Considering n bits of input it should atleast take 2 iterations.
# Due to the fact that qubits were in superposition we could account for every single combination of input just in one shot.
# -
c = q.QuantumCircuit(1, 1)
c.h(0)
# hadamard puts qubit in superposition(equal probability of 0 & 1)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
plot_bloch_multivector(statevec)
c = q.QuantumCircuit(1, 1)
c.x(0)
# not gate applied then hadamard so rotates to the negative axis
c.h(0)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
plot_bloch_multivector(statevec)
c = q.QuantumCircuit(1, 1)
c.h(0)
c.x(0)
# Hadamard puts the qubit in superposition (equal chance of 0 & 1).After not gate still in superposition(equal chance of 1 & 0)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
plot_bloch_multivector(statevec)
# Rotating multivector on y axis we could achieve negative axis before or after superposition
c = q.QuantumCircuit(1, 1)
#c.ry(math.pi, 0)
c.h(0)
c.ry(math.pi, 0)
statevec = q.execute(c, backend = statevec_sim).result().get_statevector()
plot_bloch_multivector(statevec)
| Deutsch_Jozsa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true slideshow={"slide_type": "skip"} language="html"
# <style>
# .output_wrapper, .output {
# height:auto !important;
# max-height:300px; /* your desired max-height here */
# }
# .output_scroll {
# box-shadow:none !important;
# webkit-box-shadow:none !important;
# }
# </style>
# + deletable=true editable=true slideshow={"slide_type": "skip"}
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Import Pandas
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
import pandas as pd
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Read in the dataset
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data = pd.read_csv('data-titanic.csv')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Default Index
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data.head()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Set an Index post reading of data
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data.set_index('Name')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Set an Index while reading data
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data = pd.read_csv('data-titanic.csv', index_col=3)
data.head()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Selection using Index
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data.loc['Braund, Mr. <NAME>',:]
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ## Reset Index
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data.reset_index(inplace=True)
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
data.head()
# + deletable=true editable=true slideshow={"slide_type": "skip"}
| Chapter03/Indexing in pandas dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **¿Qué es un portafolio?**
# > Introducción a la materia y descripción de las herramientas computacionales que se van a utilizar a lo largo del curso.
# ___
# ## Portafolio
# - Todos nosotros poseemos un portafolio o un conjunto de activos. Estos portafolios pueden contener activos reales (casa, carro, nevera, celular) o activos financieros (acciones, bonos u otros instrumentos).
#
# - Ahora, la conformación de esos portafolios puede ser el resultado de una serie de decisiones sin relación alguna y ajustadas al momento.
#
# - Por el contrario, puede ser el resultado de una cierta planeación. Escoger un portafolio dados la gran cantidad de activos y las múltiples combinaciones pareciera un panorama abrumador.
# <img style="center" src="http://www.creative-commons-images.com/highway-signs/images/portfolio.jpg" width="500px" height="200px" alt="atom" />
# Aprenderemos los principios básicos para construir portafolios de manera efectiva. Centraremos nuestra atención en activos financieros, aunque mucho de lo que hagamos acá podría ser aplicado a activos reales también.
# **Grandes preguntas:**
# - ¿Cómo distribuimos la riqueza entre los diferentes activos?
# - ¿Cómo elegimos inteligentemente un portafolio de inversión?
# - ¿Cómo maximizamos la rentabilidad asumiendo cierto riesgo?
# - ¿Cómo medir el riesgo?
# - ¿Cómo convencer a tu jefe que siga tu recomendación en cuanto a la distribución del dinero en activos?
# ___
# ## Herramientas computacionales
# ### - [python](https://www.python.org) - [anaconda](https://www.anaconda.com/products/individual) - [jupyter](http://jupyter.org)
# <div>
# <img style="float: left; margin: 0px 0px 15px 15px;" src="https://www.python.org/static/community_logos/python-logo.png" width="200px" height="200px" alt="atom" />
# <img style="float: left; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/en/c/cd/Anaconda_Logo.png" width="120" />
# <img style="float: left; margin: 0px 0px 15px 15px;" src="https://jupyter.org/assets/homepage/main-logo.svg" width="80" />
#
# ¿Porqué Python?
# - https://www.economist.com/graphic-detail/2018/07/26/python-is-becoming-the-worlds-most-popular-coding-language
# - https://www.economist.com/science-and-technology/2018/07/19/python-has-brought-computer-programming-to-a-vast-new-audience
# - https://pypl.github.io/PYPL.html
# ### Instalar ANACONDA... ¡No confundir Anaconda con esto!
from IPython.display import YouTubeVideo
YouTubeVideo('LDZX4ooRsWs')
# ___
# ## Herramientas adicionales
# ### jupyter notebook
# - Aprender a usar el jupyter notebook
# - Los notebooks tienen celdas de entrada y generalmente celdas de salida, así como celdas de texto. Las celdas de texto es lo que estas leyendo ahora. Las celdas de código inician con "In []:" con algún numero en los brackets. Si te colocas sobre una salda de entrada y presionas Shift-Enter, el código correrá en el <Strong> interprete </Strong> de python y el resultado se imprimirá en la celda de salida.
# **Trabajando con el notebook de jupyter**
# Además de poder realizar progrmación, tiene otras ventajas. Como ya se dieron cuenta toda esta presentación esta hecha con el notebook. Además de eso, también se puede incluir directamente dentro de este documento, texto en [Markdown](https://confluence.atlassian.com/bitbucketserver/markdown-syntax-guide-776639995.html).
#
# Uno de los atractivos más relevantes (personalmente) es que puedes escribir ecuaciones estilo $\LaTeX$, esto es gracias al proyecto [MathJax](https://www.mathjax.org) el cual se especializa en que podamos publicar matemáticas en línea. A continuación, se muestra una ejemplo.
# ___
# >Capitalización por *interés compuesto*:
# >$$C_k=C_0(1+i)^k,$$
# >donde:
# >- $C_k$ es el capital al final del $k$-ésimo periodo,
# >- $C_0$ es el capital inicial,
# >- $i$ es la tasa de interés pactada, y
# >- $k$ es el número de periodos.
#
# ___
# https://es.wikipedia.org/wiki/Inter%C3%A9s_compuesto
# #### Archivos de python (script)
# - Estos son simplemente archivos de texto con la extensión .py
# - user $ python miprograma.py
# - Cada linea en el archivo es una declaración de código en python, o parte del código.
# %run welcome.py
# ### [git](https://git-scm.com)
# <img style="float: right; margin: 15px 15px 15px 15px;" src="http://conociendogithub.readthedocs.io/en/latest/_images/Git.png" title="git" width="180" height="50">
#
# `git` es un software (de código abierto) de control de versiones diseñado por *<NAME>*, pensando en la eficiencia y la confiabilidad del mantenimiento de versiones de aplicaciones cuando éstas tienen un gran número de archivos de código fuente.</p>
#
# Se llama control de versiones a la gestión de los diversos cambios que se realizan sobre los elementos de algún producto o una configuración del mismo. Una versión, revisión o edición de un producto, es el estado en el que se encuentra el mismo en un momento dado de su desarrollo o modificación.</p>
#
# `git` es independiente de `GitHub`, y es el programa que se usa para gestionar todos los cambios en las versiones de un proyecto (individual o colaborativo, local o de acceso remoto).
# ### [GitHub](https://github.com)
#
# <img style="float: right; margin: 15px 15px 15px 15px;" src="https://c1.staticflickr.com/3/2238/13158675193_2892abac95_z.jpg" title="git" width="180" height="50">
#
# `GitHub` es una compañía que ofrece servicios de *hosting* para repositorios de `git`, y es la forma más popular para trabajar con el control de versiones que brinda `git` de forma colaborativa. En pocas palabras, permite que tanto tú como otras personas del mundo trabajen juntos en proyectos, haciendo control de versiones con `git`.
#
# De forma que usando `git` y `GitHub` en conjunto se puede trabajar tanto:
# - De forma local: repositorios en tu equipo, sin necesidad de conexión a internet (usando únicamente `git`).
# - De forma remota: repositorios remotos (compartidos) que no están necesariamente en tu máquina.
#
# **Importante**: se recomienda crear una cuenta *seria*, no solo porque la vamos a utilizar continuamente en el curso, sino porque probablemente sea parte importante de su carrera profesional.
# ### [GitKraken](https://www.gitkraken.com/)
#
# <img style="float: right; margin: 15px 15px 15px 15px;" src="https://1v5ymx3zt3y73fq5gy23rtnc-wpengine.netdna-ssl.com/wp-content/uploads/2021/06/gitkraken-keif-mono-teal-sq.svg" title="git" width="180" height="50">
#
# **Usamos `git` desde una interfaz gráfica :)**
#
# Todos los procedimientos con `git` se pueden correr desde la *consola*. Sin embargo, este modo de utilizar `git` puede ser tedioso para algunos (esto no solo ocurre con `git`).
#
# Por ello, `git` viene con algunas herramientas *GUI* (Graphical User Interface) por defecto con su instalación. Aún así, existen varias GUI desarrolladas por otras compañías para brindar una experiencia más cómoda según el sistema operativo.
#
# `GitKraken` es entonces una de estas GUI, **gratuita para el uso no comercial**, que tiene versiones para los sistemas operativos *Windows*, *Mac* y *Linux*. La estaremos usando en el curso por su versatilidad.
# ### Seguir un proyecto de otra persona
# Como ejemplo, van a seguir el proyecto de la asignatura **porinvp2022**.
#
# Los siguientes pasos nos enseñarán como mantener nuestro repositorio local actualizado con el repositorio de la asignatura.
# 1. Entrar al repositorio https://github.com/esjimenezro/porinvp2022.
# 2. En la esquina superior derecha, dar click en *fork* y esperar un momento. Esta acción copia en su cuenta de `GitHub` un repositorio idéntico al de la materia (con el mismo nombre).
# 3. Desde `GitKraken`, clonar el repositorio (el que ya está en su cuenta).
# 4. En la pestaña *REMOTE* dar click en el signo `+`.
# - Picar en `GitHub`.
# - Desplegar la pestaña y elegir esjimenezro/porinvp2022.
# - Picar en *Add remote*.
# 5. <font color=red>Añadiré un nuevo archvo en el repositorio de la materia y ustedes verán qué pasa en `GitKraken`</font>.
# 6. Arrastrar el repositorio remoto ajeno a la rama *master* y dar click en la opción *Merge esjimenezro/master into master*. Ya el repositorio local está actualizado.
# 7. Para actualizar el repositorio remoto propio hacer un *push*.
# > **Tarea 1:** <font color = blue> Realizar una presentación personal haciendo uso de la sintaxis `Markdown`. En ella deben incluir un resumen de uno de los artículos de The Economist. </font>
#
# <img src="https://raw.githubusercontent.com/louim/in-case-of-fire/master/in_case_of_fire.png" title="In case of fire (https://github.com/louim/in-case-of-fire)" width="200" height="50" align="center">
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo1/Clase1_IntroPor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.2 64-bit (''.venv'': venv)'
# name: python37264bitvenvvenvaae2d33867d9478fb568d91a7ec1d408
# ---
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv('accidents_cleanV2.csv')
df.head(2)
# ### 1. How have the number of accidents fluctuated over the past year and a half? Have they increased over the time?
#DATE to datetime format
df["fecha_incidente"] = pd.to_datetime(df["fecha_incidente"])
# +
#prueba = prueba.index.to_period("M")
df["PERIOD"] = df["fecha_incidente"].dt.to_period('M')
#Create WeekDay
df["WEEKDAY"] = df.fecha_incidente.dt.weekday
df = df.set_index('fecha_incidente')
# -
#How have the number of accidents fluctuated over the past two years? Have the increase over the time?
#Line Plot - Last two years.
fig, ax = plt.subplots(figsize=(20,5))
df.loc['2014-01-01':'2019-09-30'].groupby("PERIOD")["radicado"].count().plot(ax=ax)
# ### 2. For any particular day, during which hours are accidents most likely to occur?
# + tags=[]
# %%time
x=df['hora_incidente'].value_counts(dropna=True)
HourlyCount=plt.figure(figsize=(20,5))
ax = HourlyCount.add_subplot(111)
plt.bar(x.index, x.values)
plt.xticks(x.index)
plt.title("Hourly accident count in Medellin")
plt.show()
x.values
# -
# Accidents are most likely to occur in the afternoon.
# 3. Are there more accidents on weekdays than weekends?3. Are there more accidents on weekdays than weekends?
# Los nombres de los días son strings con 9 caracteres
df['dia_nombre']=df['dia_nombre'].apply(lambda x: x.replace(' ', ''))
# + tags=[]
# %%time
x=df['dia_nombre'].value_counts()
OrderedDays = ['LUNES', 'MARTES', 'MIÉRCOLES', 'JUEVES', 'VIERNES', 'SÁBADO', 'DOMINGO']
AccidentCount = [x[i] for i in OrderedDays]
DayOfWeekCount=plt.figure(figsize=(20,5))
ax = DayOfWeekCount.add_subplot(111)
plt.bar(OrderedDays, AccidentCount)
plt.xticks(x.index)
plt.title("Accident count by day of the week in Medellin")
for i in ax.patches:
ax.text(i.get_x()+0.3, i.get_height()+30, i.get_height())
plt.show()
# -
# Accidents are more likely to occur during the weekdays. We can conclude that friday is the highest accidentality weekday.
| backend/src/playground/MJ/EDA_E1_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We’ll start by doing all necessary imports, and we’ll let our Jupyter Notebook know that it should display graphs and images in the notebook itself.
# +
# %matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import urllib
import sys
import os
import zipfile
# -
# The files we're about to use may take five minutes or more to download, so if you're following along by running the program in the corresponding notebook, feel free to start running the next few cells. In the meantime, let’s explore textual entailment in further detail.
import sys
sys.version
print(sys.version)
print(123)
# +
glove_zip_file = "glove.6B.zip"
glove_vectors_file = "glove.6B.50d.txt"
snli_zip_file = "snli_1.0.zip"
snli_dev_file = "snli_1.0_dev.txt"
snli_full_dataset_file = "snli_1.0_train.txt"
# -
# +
def unzip_single_file(zip_file_name, output_file_name):
"""
If the outFile is already created, don't recreate
If the outFile does not exist, create it from the zipFile
"""
if not os.path.isfile(output_file_name):
with open(output_file_name, 'wb') as out_file:
with zipfile.ZipFile(zip_file_name) as zipped:
for info in zipped.infolist():
if output_file_name in info.filename:
with zipped.open(info) as requested_file:
out_file.write(requested_file.read())
return
unzip_single_file(glove_zip_file, glove_vectors_file)
unzip_single_file(snli_zip_file, snli_dev_file)
# unzip_single_file(snli_zip_file, snli_full_dataset_file)
# -
# ---
#
# Now that we have our GloVe vectors downloaded, we can load them into memory, deserializing the space separated format into a Python dictionary:
# The program use the function of numpy to translate the glove_vectors_file into matrix
glove_wordmap = {}
count = 0
with open(glove_vectors_file, "r",encoding='utf-8') as glove:
for line in glove:
name, vector = tuple(line.split(" ", 1))
glove_wordmap[name] = np.fromstring(vector, sep=" ")
count += 1
print(count)
# print(name,glove_wordmap[name])
# ---
#
# Once we have our words, we need our input to contain entire sentences and process it through a neural network. Let's start with making the sequence:
# sentence2sequence 函数通过" "将整个句子进行分词(英文的分词依靠空格)
# 该函数将整个句子分为单独的一个个单词
# 并将整个句子存入一个(n,d)的矩阵之中
# n为句子单词的个数,d为每个单词的长度?最大长度吧
#
# 这是一个简单具有分词功能的函数,它根据句子的语序将整个句子分成
def sentence2sequence(sentence):
"""
- Turns an input sentence into an (n,d) matrix,
where n is the number of tokens in the sentence
and d is the number of dimensions each word vector has.
d是一个定值50,通过glove_wordmap这个从文件中读取的word_map映射,将一个单词映射成为一个50维浮点型数据
n是一个变化的值,根据这个句子中包含的单词的个数决定
该函数将通过“ ”将整个句子进行分词,然后通过golve_map映射,将单个单词转换为一个五十维的浮点向量
之后再放入模型中进行训练
Tensorflow doesn't need to be used here, as simply
turning the sentence into a sequence based off our
mapping does not need the computational power that
Tensorflow provides. Normal Python suffices for this task.
"""
tokens = sentence.lower().split(" ")
rows = []
words = []
#Greedy search for tokens
for token in tokens:
i = len(token)
while len(token) > 0 and i > 0:
# print(token)
word = token[:i]
if word in glove_wordmap:
rows.append(glove_wordmap[word])
words.append(word)
token = token[i:]
else:
i = i-1
#print(rows, words)
# j=0
# for line in rows:
# print(("line %d:" % j) + line)
# print(line)
# j += 1
# print(words)
# for word, row in zip(words, rows):
# print(len(word),len(row))
# for line1, line2 in zip(word, row):
# print(line1 , ":", line2)
return rows, words
# To better visualize the word vectorization process, and to see what the computer sees when it looks at a sentence, we can represent the vectors as images. Each row represents a single word, and the columns represent individual dimensions of the vectorized word. The vectorizations are trained in terms of relationships to other words, and so what the representations actually mean is ambiguous. The computer can understand this vector language, and that’s the most important part to us. Generally speaking, two vectors that contain similar colors in the same positions represent words that are similar in meaning.
#
# "\n"
# visualize函数是为了更好的展示由输入句子产生的矩阵
# 将这个向量矩阵以图片的形式展示了出来
# +
def visualize(sentence):
rows, words = sentence2sequence(sentence)
mat = np.vstack(rows)
fig = plt.figure()
ax = fig.add_subplot(111)
shown = ax.matshow(mat, aspect="auto")
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
fig.colorbar(shown)
ax.set_yticklabels([""]+words)
plt.show()
visualize("The quick brown fox jumped over the lazy dog.")
visualize("The pretty flowers shone in the sunlight.")
# -
# -----
# Recurrent Neural Networks (also known as RNNs) are a sequence-learning tool for neural networks. This type of neural network has only one layer’s worth of hidden inputs, which is re-used for each input from the sequence, along with a “memory” that’s passed ahead to the next input’s calculations. These are calculated using matrix multiplication where the matrix indices are trained weights, just like they are in a fully-connected layer.
#
# The same calculations are repeated for each input in the sequence, meaning that a single “layer” of a recurrent neural network can be unrolled into many layers. In fact, there will be as many layers as there are inputs in the sequence. This allows the network to process a very complex sentence. TensorFlow includes its own implementation of a vanilla RNN cell, BasicRNNCell, which can be added to your TensorFlow graph as follows:
# 通过RNN神经网络对模型进行训练,并且设置RNN模型层数为64
# 并紧接着通过TensorFlow搭建我们的RNN模型
rnn_size = 64
rnn = tf.contrib.rnn.BasicRNNCell(rnn_size)
# ----
#
# # Defining the constants for our network
#
# Since we aren’t going to use a vanilla RNN layer in our network, let's clear out the graph and add an LSTM layer, which TensorFlow also includes by default. Since this is going to be the first part of our actual network, let's also define all the constants we'll need for the network, which we'll talk about as they come up:
# 设置相应的图的参数
# +
#Constants setup
max_hypothesis_length, max_evidence_length = 30, 30
batch_size, vector_size, hidden_size = 128, 50, 64
lstm_size = hidden_size
weight_decay = 0.0001
learning_rate = 1
input_p, output_p = 0.5, 0.5
training_iterations_count = 100000
display_step = 10
def score_setup(row):
convert_dict = {
'entailment': 0,
'neutral': 1,
'contradiction': 2
}
score = np.zeros((3,))
for x in range(1,6):
tag = row["label"+str(x)]
if tag in convert_dict: score[convert_dict[tag]] += 1
return score / (1.0*np.sum(score))
def fit_to_size(matrix, shape):
res = np.zeros(shape)
slices = [slice(0,min(dim,shape[e])) for e, dim in enumerate(matrix.shape)]
res[slices] = matrix[slices]
return res
# -
# snli_1.0_dev.txt中存放了用于训练模型的数据,通过split_data_into_scores函数把这些用于训练的数据转换为向量矩阵生成训练集
# +
def split_data_into_scores():
import csv
with open("snli_1.0_dev.txt","r") as data:
train = csv.DictReader(data, delimiter='\t')
evi_sentences = []
hyp_sentences = []
labels = []
scores = []
for row in train:
hyp_sentences.append(np.vstack(
sentence2sequence(row["sentence1"].lower())[0]))
evi_sentences.append(np.vstack(
sentence2sequence(row["sentence2"].lower())[0]))
labels.append(row["gold_label"])
scores.append(score_setup(row))
hyp_sentences = np.stack([fit_to_size(x, (max_hypothesis_length, vector_size))
for x in hyp_sentences])
evi_sentences = np.stack([fit_to_size(x, (max_evidence_length, vector_size))
for x in evi_sentences])
return (hyp_sentences, evi_sentences), labels, np.array(scores)
data_feature_list, correct_values, correct_scores = split_data_into_scores()
l_h, l_e = max_hypothesis_length, max_evidence_length
N, D, H = batch_size, vector_size, hidden_size
l_seq = l_h + l_e
# -
# We'll also reset the graph to not include the RNN cell we added earlier, since we won't be using that for this network:
#
#
tf.reset_default_graph()
# With both those out of the way, we can define our LSTM using TensorFlow as follows:
# LSTM:长短期记忆网络,是一种特殊的RNN网络类型
lstm = tf.contrib.rnn.BasicLSTMCell(lstm_size)
# The loss of certain pieces of crucial memory means that complicated relationships required for first order logic have a harder time forming with dropout, and so for our LSTM layer we’ll skip using dropout on internal gates, instead using it on everything else. Thankfully, this is the default implementation of Tensorflow’s DropoutWrapper for recurrent layers:
# 使用LSTM的原因
#
lstm_drop = tf.contrib.rnn.DropoutWrapper(lstm, input_p, output_p)
# ----
#
# With all the explanations out of the way, we can finish up our model. The first step is tokenizing and using our GloVe dictionary to turn the two input sentences into a single sequence of vectors. Since we can’t effectively use dropout on information that gets passed within an LSTM, we’ll use dropout on features from words, and on final output instead -- effectively using dropout on the first and last layers from the unrolled LSTM network portions.
#
# The final output from the LSTMs will be passed into a set of fully connected layers, and then from that we’ll get a single real-valued score that indicates how strong each of the kinds of entailment are, which we use to select our final result and our confidence in that result.
# 定义用于训练模型的必要参数以及相应的参数解读
# +
# N: The number of elements in each of our batches,
# which we use to train subsets of data for efficiency's sake.
# l_h: The maximum length of a hypothesis, or the second sentence. This is
# used because training an RNN is extraordinarily difficult without
# rolling it out to a fixed length.
# l_e: The maximum length of evidence, the first sentence. This is used
# because training an RNN is extraordinarily difficult without
# rolling it out to a fixed length.
# D: The size of our used GloVe or other vectors.
hyp = tf.placeholder(tf.float32, [N, l_h, D], 'hypothesis')
evi = tf.placeholder(tf.float32, [N, l_e, D], 'evidence')
y = tf.placeholder(tf.float32, [N, 3], 'label')
# hyp: Where the hypotheses will be stored during training.
# evi: Where the evidences will be stored during training.
# y: Where correct scores will be stored during training.
# lstm_size: the size of the gates in the LSTM,
# as in the first LSTM layer's initialization.
lstm_back = tf.contrib.rnn.BasicLSTMCell(lstm_size)
# lstm_back: The LSTM used for looking backwards
# through the sentences, similar to lstm.
# input_p: the probability that inputs to the LSTM will be retained at each
# iteration of dropout.
# output_p: the probability that outputs from the LSTM will be retained at
# each iteration of dropout.
lstm_drop_back = tf.contrib.rnn.DropoutWrapper(lstm_back, input_p, output_p)
# lstm_drop_back: A dropout wrapper for lstm_back, like lstm_drop.
fc_initializer = tf.random_normal_initializer(stddev=0.1)
# fc_initializer: initial values for the fully connected layer's weights.
# hidden_size: the size of the outputs from each lstm layer.
# Multiplied by 2 to account for the two LSTMs.
fc_weight = tf.get_variable('fc_weight', [2*hidden_size, 3],
initializer = fc_initializer)
# fc_weight: Storage for the fully connected layer's weights.
fc_bias = tf.get_variable('bias', [3])
# fc_bias: Storage for the fully connected layer's bias.
# tf.GraphKeys.REGULARIZATION_LOSSES: A key to a collection in the graph
# designated for losses due to regularization.
# In this case, this portion of loss is regularization on the weights
# for the fully connected layer.
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES,
tf.nn.l2_loss(fc_weight))
x = tf.concat([hyp, evi], 1) # N, (Lh+Le), d
# Permuting batch_size and n_steps
x = tf.transpose(x, [1, 0, 2]) # (Le+Lh), N, d
# Reshaping to (n_steps*batch_size, n_input)
x = tf.reshape(x, [-1, vector_size]) # (Le+Lh)*N, d
# Split to get a list of 'n_steps' tensors of shape (batch_size, n_input)
x = tf.split(x, l_seq,)
# x: the inputs to the bidirectional_rnn
# tf.contrib.rnn.static_bidirectional_rnn: Runs the input through
# two recurrent networks, one that runs the inputs forward and one
# that runs the inputs in reversed order, combining the outputs.
rnn_outputs, _, _ = tf.contrib.rnn.static_bidirectional_rnn(lstm, lstm_back,
x, dtype=tf.float32)
# rnn_outputs: the list of LSTM outputs, as a list.
# What we want is the latest output, rnn_outputs[-1]
classification_scores = tf.matmul(rnn_outputs[-1], fc_weight) + fc_bias
# The scores are relative certainties for how likely the output matches
# a certain entailment:
# 0: Positive entailment
# 1: Neutral entailment
# 2: Negative entailment
# -
# In order to test the accuracy and begin to add in optimization constraints, we need to show TensorFlow how to calculate the accuracy, or -- the percentage of correctly predicted labels.
#
# We also need to determine a loss, to show how poorly the network is doing. Since we have both classification scores and optimal scores, the choice here is using a variation on softmax loss from Tensorflow: tf.nn.softmax_cross_entropy_with_logits. We add in regularization losses to help with overfitting, and then prepare an optimizer to learn how to reduce the loss.
# +
with tf.variable_scope('Accuracy'):
predicts = tf.cast(tf.argmax(classification_scores, 1), 'int32')
y_label = tf.cast(tf.argmax(y, 1), 'int32')
corrects = tf.equal(predicts, y_label)
num_corrects = tf.reduce_sum(tf.cast(corrects, tf.float32))
accuracy = tf.reduce_mean(tf.cast(corrects, tf.float32))
with tf.variable_scope("loss"):
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(
logits = classification_scores, labels = y)
loss = tf.reduce_mean(cross_entropy)
total_loss = loss + weight_decay * tf.add_n(
tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES))
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
opt_op = optimizer.minimize(total_loss)
# -
# Finally, we can train the network! If you installed TQDM, you can use it to keep track of progress as the network trains.
#
# +
# Initialize variables
init = tf.global_variables_initializer()
# Use TQDM if installed
tqdm_installed = False
try:
from tqdm import tqdm
tqdm_installed = True
except:
pass
# Launch the Tensorflow session
# sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
sess.run(init)
# training_iterations_count: The number of data pieces to train on in total
# batch_size: The number of data pieces per batch
training_iterations = range(0,training_iterations_count,batch_size)
if tqdm_installed:
# Add a progress bar if TQDM is installed
training_iterations = tqdm(training_iterations)
for i in training_iterations:
# Select indices for a random data subset
batch = np.random.randint(data_feature_list[0].shape[0], size=batch_size)
# Use the selected subset indices to initialize the graph's
# placeholder values
hyps, evis, ys = (data_feature_list[0][batch,:],
data_feature_list[1][batch,:],
correct_scores[batch])
# Run the optimization with these initialized values
sess.run([opt_op], feed_dict={hyp: hyps, evi: evis, y: ys})
# display_step: how often the accuracy and loss should
# be tested and displayed.
if (i/batch_size) % display_step == 0:
# Calculate batch accuracy
acc = sess.run(accuracy, feed_dict={hyp: hyps, evi: evis, y: ys})
# Calculate batch loss
tmp_loss = sess.run(loss, feed_dict={hyp: hyps, evi: evis, y: ys})
# Display results
print("Iter " + str(i/batch_size) + ", Minibatch Loss= " + \
"{:.6f}".format(tmp_loss) + ", Training Accuracy= " + \
"{:.5f}".format(acc))
# -
# Your network is now trained! You should see accuracies around 50-55%, which can be improved by careful modification of hyperparameters and increasing the dataset size to include the entire training set. Usually, this will correspond with an increase in training time.
#
# Feel free to modify the following code by inserting your own sentences:
# +
evidences = ["I like dogs."]
hypotheses = ["Peopel don't like dogs."]
sentence1 = [fit_to_size(np.vstack(sentence2sequence(evidence)[0]),
(30, 50)) for evidence in evidences]
sentence2 = [fit_to_size(np.vstack(sentence2sequence(hypothesis)[0]),
(30,50)) for hypothesis in hypotheses]
prediction = sess.run(classification_scores, feed_dict={hyp: (sentence1 * N),
evi: (sentence2 * N),
y: [[0,0,0]]*N})
print(["Positive", "Neutral", "Negative"][np.argmax(prediction[0])]+
" entailment")
# -
# Finally, once we're done playing with our model, we'll close the session to free up system resources.
saver = tf.train.Saver()
saver.save(sess,r"C:\Users\86151\Desktop\trained.model")
sess.close()
| Homework/2019/Task1/12/code/Entailment with TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.9 64-bit
# language: python
# name: python3
# ---
# # Generate insights on credit products to avoid legal disputes
# ## Plotting the percentage of disputes lodged per product
# +
## analyze the csv file to determine impact of variables on disputes
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
# read specific columns of csv file using Pandas
df = pd.read_csv("/Users/abhinav_chinta/Code/Abhinav-Chinta-Quant-OA/eda-consumer-finance/consumer_complaints.csv")
## find percentages of case disputed per product
items = df['product'].unique()
result = []
def count_dispute(string, column = 1):
count = 0
for row in df.values:
if (row[column] == string and row[16].lower() == 'yes'):
count += 1
return count
# +
# plotting bar graph for percentage of disputes lodged per product
import numpy as np
import matplotlib.pyplot as plt
data = {}
for item in items:
data[item] = count_dispute(item) / list(df["product"]).count(item) * 100
result = dict(sorted(data.items(), key=lambda item:- item[1]))
# creating the dataset
item = list(result.keys())
values = list(result.values())
fig = plt.figure(figsize = (22, 10))
# creating the bar plot
plt.bar(item, values, color ='maroon',
width = 0.7)
plt.xlabel("products")
plt.ylabel("Disputes percentage (out of 100)")
plt.title("Disputes lodged per product")
plt.show()
# -
# ## Determining states in which disputes are most prevelant in
# +
# states which disputes are most prevalent in
st_abbrv = pd.read_csv("/Users/abhinav_chinta/Code/Abhinav-Chinta-Quant-OA/eda-consumer-finance/state_abbrev.csv")
st_pop = pd.read_csv("/Users/abhinav_chinta/Code/Abhinav-Chinta-Quant-OA/eda-consumer-finance/state_pop.csv")
# merging cosumer_complaints and state population data
new_df = pd.concat([st_pop, st_abbrv], axis=1, ignore_index=True)
new_df = new_df.drop(new_df.columns[[2,3,4,6]], axis=1)
states = df['state'].unique()
print(states)
# +
# generating a dictionary with states and their corresponding percentage of cases as a metric of total
st_perct = {}
for i in range(len(new_df)):
st_perct[states[i]] = count_dispute(states[i], 8) / new_df[1][i] * 100
print(st_perct)
# +
# visualizing states dispute percentages
import matplotlib.pyplot as plt
import numpy as np
y = st_perct.values()
x = st_perct.keys()
plt.pie(y, labels = x, radius=5)
plt.show()
# -
# ## Determining Tags with highest dispute rate
# +
df = pd.read_csv("/Users/abhinav_chinta/Code/Abhinav-Chinta-Quant-OA/eda-consumer-finance/consumer_complaints.csv")
tags = df['tags'].unique()
# generating tag data percentages
tag_data = {}
temp = df.groupby('tags').size()
for tag in tags:
if isinstance(tag, str):
tag_data[tag] = count_dispute(tag, 10) / temp[tag] * 100
print(tag_data)
# +
# visualizing tag dispute percentages using a pie chart
import matplotlib.pyplot as plt
import numpy as np
y = tag_data.values()
x = tag_data.keys()
plt.pie(y, labels = x, radius=5)
plt.show()
# -
# ## Finding the most common language / keywords that companies used in their personal statements
# +
# concatenate all company responses
# creating a string with all company responses
text = ""
for line in df["company_public_response"]:
if isinstance(line, str):
for word in line.split():
if len(word) > 3:
text += word + " "
# +
# finding keywords using spacy
import spacy
nlp = spacy.load("en_core_sci_lg")
# splitting the text into 5 packets
doc = nlp(text[:len(text) // 5])
# +
# Printing all unique keywords
keywords = list(map(str, doc.ents))
print(set(keywords))
# -
# ## Features considered
# - Product
# - Sub_product
# - Issue
# - sub_issue
# - company_response_to_consumer
# - timely_response
# ## Building a model that takes in relevant features and the company's closure method to predict if the consumer is going to dispute it or not
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
rev_df = df[['product','sub_product','issue','sub_issue','company_response_to_consumer','timely_response']]
for column in rev_df:
rev_df[column] = rev_df[column].fillna("N")
# +
# split data in train/test
for feature in rev_df.columns:
le = preprocessing.LabelEncoder()
le = le.fit(rev_df[feature])
rev_df[feature] = le.transform(rev_df[feature])
X = rev_df
y = df['consumer_disputed?']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20)
X_train.head()
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import make_scorer, accuracy_score
from sklearn.model_selection import GridSearchCV
# Choose the type of classifier.
clf = RandomForestClassifier()
# Choose some parameter combinations to try
parameters = {'n_estimators': [4],
'max_features': ['auto'],
'criterion': ['entropy'],
'max_depth': [5],
'min_samples_split': [3],
'min_samples_leaf': [1]
}
# Type of scoring used to compare parameter combinations
acc_scorer = make_scorer(accuracy_score)
# Run the grid search
grid_obj = GridSearchCV(clf, parameters, scoring=acc_scorer, refit=True)
grid_obj = grid_obj.fit(X_train, y_train)
# Set the clf to the best combination of parameters
clf = grid_obj
# Fit the best algorithm to the data.
clf.fit(X_train, y_train)
predictions = clf.predict(X_test)
print(accuracy_score(y_test, predictions))
# -
# ## Prediction accuracy of the RandomForest Classifier
# ### 0.7993650622346932
# ## Future Scope
# - Generate a metric to determine products to sell in each state, miniminzing complaints
# - Determining the best company response to a given consumer complaint
# - Finding the best closure method for the bank
| eda-consumer-finance/abhinav_chinta_Quant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime
# %matplotlib inline
pd.set_option('display.max_rows', 500)
# -
# 
#
pd_JH_data=pd.read_csv('../data/processed/COVID_relational_confirmed.csv',sep=';',parse_dates=[0])
pd_JH_data=pd_JH_data.sort_values('date',ascending=True).reset_index(drop=True).copy()
pd_JH_data.head()
test_data=pd_JH_data[((pd_JH_data['country']=='US')|
(pd_JH_data['country']=='Germany'))&
(pd_JH_data['date']>'2020-03-20')]
| notebooks/Data_Preparation_Large_File_Aayush_Niranjay_Deshpande.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Notebook written by [<NAME>](https://github.com/zhedongzheng)
# +
from tqdm import tqdm
import tensorflow as tf
import numpy as np
import pprint
import os
import sys
sys.path.append(os.path.dirname(os.getcwd()))
from data import WN18
# -
class Config:
seed = 21
n_epochs = 10
batch_size = 100
embed_dim = 200
# +
"""
e: entity
s: subject
p: predicate
o: object
"""
def read_triples(path):
triples = []
with open(path, 'rt') as f:
for line in f.readlines():
s, p, o = line.split()
triples += [(s.strip(), p.strip(), o.strip())]
return triples
def load_triple():
WN18.download()
triples_tr = read_triples('../data/WN18/wn18/train.txt')
triples_va = read_triples('../data/WN18/wn18/valid.txt')
triples_te = read_triples('../data/WN18/wn18/test.txt')
triples_all = triples_tr + triples_va + triples_te
return triples_all, triples_tr, triples_va, triples_te
def build_vocab(triples):
params = {}
e_set = {s for (s, p, o) in triples} | {o for (s, p, o) in triples}
p_set = {p for (s, p, o) in triples}
params['e_vocab_size'] = len(e_set)
params['p_vocab_size'] = len(p_set)
params['embed_dim'] = Config.embed_dim
e2idx = {e: idx for idx, e in enumerate(sorted(e_set))}
p2idx = {p: idx for idx, p in enumerate(sorted(p_set))}
return e2idx, p2idx, params
def build_train_data(triples_tr, e2idx, p2idx):
x_s = np.array([e2idx[s] for (s, p, o) in triples_tr], dtype=np.int32)
x_p = np.array([p2idx[p] for (s, p, o) in triples_tr], dtype=np.int32)
x_o = np.array([e2idx[o] for (s, p, o) in triples_tr], dtype=np.int32)
x = {'s': x_s,
'p': x_p,
'o': x_o}
y = np.ones([len(x_s)], dtype=np.float32)
return x, y
def train_input_fn(triples_tr, e2idx, p2idx, random_state, params):
x, y = build_train_data(triples_tr, e2idx, p2idx)
s, p, o = x['s'], x['p'], x['o']
s_ = random_state.choice(params['e_vocab_size'], s.shape)
o_ = random_state.choice(params['e_vocab_size'], o.shape)
x_ = {
's': np.concatenate([s, s_, s]),
'p': np.concatenate([p, p, p]),
'o': np.concatenate([o, o, o_])}
y_ = np.concatenate([y, np.zeros([2*len(y)], dtype=np.float32)])
return tf.estimator.inputs.numpy_input_fn(x = x_,
y = y_,
batch_size = Config.batch_size,
num_epochs = 1,
shuffle = True)
# +
def s_next_batch(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size):
for (s, p, o) in tqdm(eval_triples, total=len(eval_triples), ncols=70):
s_idx, p_idx, o_idx = e2idx[s], p2idx[p], e2idx[o]
xs = np.arange(nb_entities)
xp = np.full(shape=(nb_entities,), fill_value=p_idx, dtype=np.int32)
xo = np.full(shape=(nb_entities,), fill_value=o_idx, dtype=np.int32)
for i in range(0, len(xs), batch_size):
yield xs[i: i+batch_size], xp[i: i+batch_size], xo[i: i+batch_size]
def o_next_batch(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size):
for (s, p, o) in tqdm(eval_triples, total=len(eval_triples), ncols=70):
s_idx, p_idx, o_idx = e2idx[s], p2idx[p], e2idx[o]
xs = np.full(shape=(nb_entities,), fill_value=s_idx, dtype=np.int32)
xp = np.full(shape=(nb_entities,), fill_value=p_idx, dtype=np.int32)
xo = np.arange(nb_entities)
for i in range(0, len(xs), batch_size):
yield xs[i: i+batch_size], xp[i: i+batch_size], xo[i: i+batch_size]
def s_input_fn(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size):
dataset = tf.data.Dataset.from_generator(
lambda: s_next_batch(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size),
(tf.int32, tf.int32, tf.int32),
(tf.TensorShape([None,]),
tf.TensorShape([None,]),
tf.TensorShape([None,]),))
iterator = dataset.make_one_shot_iterator()
s, p, o = iterator.get_next()
return {'s': s, 'p': p, 'o': o}
def o_input_fn(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size):
dataset = tf.data.Dataset.from_generator(
lambda: o_next_batch(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size),
(tf.int32, tf.int32, tf.int32),
(tf.TensorShape([None,]),
tf.TensorShape([None,]),
tf.TensorShape([None,]),))
iterator = dataset.make_one_shot_iterator()
s, p, o = iterator.get_next()
return {'s': s, 'p': p, 'o': o}
def evaluate_rank(model,
valid_triples,
test_triples,
all_triples,
e2idx,
p2idx,
nb_entities,
batch_size):
#for eval_name, eval_triples in [('valid', valid_triples), ('test', test_triples)]:
for eval_name, eval_triples in [('test', test_triples)]:
_scores_s = list(model.predict(
lambda: s_input_fn(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size), yield_single_examples=False))
_scores_o = list(model.predict(
lambda: o_input_fn(eval_triples,
e2idx,
p2idx,
nb_entities,
batch_size), yield_single_examples=False))
ScoresS = np.concatenate(_scores_s).reshape([len(eval_triples), nb_entities])
ScoresO = np.concatenate(_scores_o).reshape([len(eval_triples), nb_entities])
ranks_s, ranks_o = [], []
filtered_ranks_s, filtered_ranks_o = [], []
for ((s, p, o), scores_s, scores_o) in tqdm(zip(eval_triples,
ScoresS,
ScoresO),
total=len(eval_triples),
ncols=70):
s_idx, p_idx, o_idx = e2idx[s], p2idx[p], e2idx[o]
ranks_s += [1 + np.argsort(np.argsort(- scores_s))[s_idx]]
ranks_o += [1 + np.argsort(np.argsort(- scores_o))[o_idx]]
filtered_scores_s = scores_s.copy()
filtered_scores_o = scores_o.copy()
rm_idx_s = [e2idx[fs] for (fs, fp, fo) in all_triples if fs != s and fp == p and fo == o]
rm_idx_o = [e2idx[fo] for (fs, fp, fo) in all_triples if fs == s and fp == p and fo != o]
filtered_scores_s[rm_idx_s] = - np.inf
filtered_scores_o[rm_idx_o] = - np.inf
filtered_ranks_s += [1 + np.argsort(np.argsort(- filtered_scores_s))[s_idx]]
filtered_ranks_o += [1 + np.argsort(np.argsort(- filtered_scores_o))[o_idx]]
ranks = ranks_s + ranks_o
filtered_ranks = filtered_ranks_s + filtered_ranks_o
for setting_name, setting_ranks in [('Raw', ranks), ('Filtered', filtered_ranks)]:
mean_rank = np.mean(1 / np.asarray(setting_ranks))
print('[{}] {} MRR: {}'.format(eval_name, setting_name, mean_rank))
for k in [1, 3, 5, 10]:
hits_at_k = np.mean(np.asarray(setting_ranks) <= k) * 100
print('[{}] {} Hits@{}: {}'.format(eval_name, setting_name, k, hits_at_k))
# +
def forward(features, params):
e_embed = tf.get_variable('e_embed',
[params['e_vocab_size'], params['embed_dim']],
initializer=tf.contrib.layers.xavier_initializer())
p_embed = tf.get_variable('p_embed',
[params['p_vocab_size'], params['embed_dim']],
initializer=tf.contrib.layers.xavier_initializer())
s = tf.nn.embedding_lookup(e_embed, features['s'])
p = tf.nn.embedding_lookup(p_embed, features['p'])
o = tf.nn.embedding_lookup(e_embed, features['o'])
logits = tf.reduce_sum(s * p * o, axis=1)
return logits
def model_fn(features, labels, mode, params):
logits = forward(features, params)
if mode == tf.estimator.ModeKeys.TRAIN:
tf.logging.info('\n'+pprint.pformat(tf.trainable_variables()))
tf.logging.info('params: %d'%count_train_params())
loss_op = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=logits,
labels=labels))
train_op = tf.train.AdamOptimizer().minimize(
loss_op, global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode = mode,
loss = loss_op,
train_op = train_op)
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode, predictions = tf.sigmoid(logits))
def count_train_params():
return np.sum([np.prod([d.value for d in v.get_shape()]) for v in tf.trainable_variables()])
# +
random_state = np.random.RandomState(Config.seed)
triples_all, triples_tr, triples_va, triples_te = load_triple()
e2idx, p2idx, params = build_vocab(triples_tr)
model = tf.estimator.Estimator(model_fn,
params = params)
for _ in range(Config.n_epochs):
model.train(train_input_fn(triples_tr,
e2idx,
p2idx,
random_state,
params))
evaluate_rank(model,
triples_va,
triples_te,
triples_all,
e2idx,
p2idx,
params['e_vocab_size'],
batch_size = 10*Config.batch_size)
# -
| src_kg/link_prediction/main/wn18_distmult_1v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from splinter import Browser
from bs4 import BeautifulSoup
import pandas as pd
import requests
import shutil
import time
# +
def scrape():
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
# create mars-data dict that we can insert into mongo
marsdata = {}
# visit https://mars.nasa.gov/news/
url='https://mars.nasa.gov/news/'
browser.visit(url)
html=browser.html
soup=BeautifulSoup(html,'html.parser')
news_titles = soup.find_all('div', class_='content_title')
news_ps = soup.find_all('div', class_='article_teaser_body')
# last news title
titles=[]
for news in news_titles:
title=news.find("a").text
titles.append(title)
last_news=titles[0]
# last news paraghragh
paraghraph=[]
for p in news_ps:
par=p.text
paraghraph.append(par)
last_news_p=paraghraph[0]
# add our last news and last paraghraph to to Marse_data
marsdata["news_title"] = last_news
marsdata["news_p"] = last_news_p
#Mars Featured large size Image
# visit https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars
Feature_url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(Feature_url)
# navigate by soliter to site
links_found = browser.find_link_by_partial_text("FULL IMAGE")
links_found.click()
time.sleep(10)
links_found = browser.find_link_by_partial_text("more info")
links_found.click()
browser.is_element_present_by_css("img", wait_time=1)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
url_image= soup.find("img" , class_="main_image")["src"]
if "httss://www.jpl.nasa.gov:" not in url_image:url_image= "https://www.jpl.nasa.gov"+url_image
#add image url in Mars_data dic
marsdata["featured_image_url"] = url_image
# print large picture
# response = requests.get(featured_image_url, stream=True)
# with open('img.png', 'wb') as out_file:
# shutil.copyfileobj(response.raw, out_file)
# from IPython.display import Image
# Image(url='img.png')
#Mars Weather
#visit https://twitter.com/MarsWxReport?lang=en
twitter_url = 'https://twitter.com/MarsWxReport?lang=en'
browser.visit(twitter_url)
time.sleep(5)
html=browser.html
weather_soup=BeautifulSoup(html,'html.parser')
results=weather_soup.find_all('div', class_='js-tweet-text-container')
weather_list=[]
for item in results:
weather=item.find('p',class_='TweetTextSize TweetTextSize--normal js-tweet-text tweet-text').text
if "daylight" in weather:
weather_list.append(weather)
mars_weather=weather_list[0]
# adding last twitt in Mars data dict
marsdata["mars_weather"] = mars_weather
#Mars Fact
# visit http://space-facts.com/mars/ explor data with pandas
fact_url='http://space-facts.com/mars/'
#scraping data from url wiyh pandas data frame
tables = pd.read_html(fact_url)
# convert first table to dataframe
df = tables[0]
df.columns = [' ','Value']
#remove data frame index from html table and justify table
mars_facts_html = df.to_html(na_rep = " ", classes="table table-sm table-striped", justify="left", col_space=0,index=False)
mars_facts_html.replace('\n', '')
# save filr directly as html
df.to_html('Fact.html')
marsdata["Mars_facts"] = mars_facts_html
#Mars Hemispheres
Hemispheres_url='https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(Hemispheres_url)
links = ['Cerberus', 'Schiaparelli', 'Syrtis' , 'Valles']
hemisphere_image_urls=[]
# use for loop to get data for image url and titles and added to list .
for link in links:
hemisphere_image_urls_dic={}
link_click = browser.find_link_by_partial_text(link)
link_click.click()
time.sleep(10)
# browser.is_element_present_by_css("img.wide-image", wait_time=10)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
image_url=soup.find("img",class_="wide-image")["src"]
title=soup.find("h2",class_="title").text
if "https://astrogeology.usgs.gov:" not in image_url: image_url = "https://astrogeology.usgs.gov"+image_url
hemisphere_image_urls_dic['title'] = title
hemisphere_image_urls_dic['image_url']=image_url
hemisphere_image_urls.append(hemisphere_image_urls_dic)
browser.back()
marsdata["hemisphere_title_urls"] = hemisphere_image_urls
browser.quit()
return marsdata
# -
scrape()
| scrape_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exloratory data analysis for google and apple store data
# Including something
# ## Data importing
# Data from the app-store and play-store.
# +
from csv import reader
#Apple
open_apple=open('AppleStore.csv')
read_apple=reader(open_apple)
App_store=list(read_apple)
App_store_header=App_store[0] #variable name line
App_store=App_store[1:]
#print(App_store)
#Android
open_google=open('googleplaystore.csv')
read_google=reader(open_google)
Play_store=list(read_google)
Play_store_header=Play_store[0] #variable name line
Play_store=Play_store[1:]
#print(Play_store)
# -
# ## Create explore_data function
# The function explore_data is a user-defined function that calculates the number of rows and columns.
def explore_data(dataset, start, end, rows_and_columns=False):
dataset_slice =dataset[start:end]
for row in dataset_slice:
print(row)
print('\n')
if rows_and_columns:
print('Number of rows:', len(dataset))
print('Number of columns:',len(dataset[0]))
print(App_store[0:5])
print('\n')
print(Play_store[0:5])
# print(App_store_header)
# explore_data(App_store,0,3,True)
# print('\n')
# print(Play_store_header)
# explore_data(Play_store,0,3,True)
# ## Removing row has an error
# The row index 10472 has problem with value, so remove it.
print(Play_store[10472]) # incorrect row
print('\n')
print(Play_store_header) # header
print('\n')
print(Play_store[0]) # correct row
print(len(Play_store))
del Play_store[10472] # don't run this more than once
print(len(Play_store))
for app in Play_store:
name=app[0]
if name=="Instagram":
print(app)
# ## Removing duplicate data
# We have to remove duplicate rows in the data.
duplicate_apps=[]
unique_apps=[]
for app in Play_store:
name=app[0]
if name in unique_apps:
duplicate_apps.append(name)
else:
unique_apps.append(name)
print('Number of duplicate apps:',len(duplicate_apps))
print('\n')
print('Examples of duplicate apps:',duplicate_apps[:15])
# ## Creating dictionary of app name and highest number of review
# Using dictionary, update to highest number of reviews looping over the dataset.
reviews_max={}
print(reviews_max)
for i in Play_store:
name=i[0]
n_reviews=float(i[3])
if name in reviews_max and reviews_max[name] < n_reviews:
reviews_max[name]=n_reviews
elif name not in reviews_max:
reviews_max[name]=n_reviews
print(reviews_max)
len(reviews_max)
# ## Cleaning data set without duplicate rows
# Remove rows with non-highest reviews in the dataset.
android_clean=[]; already_added=[]
for i in Play_store: #loop over play_store data
name=i[0]
n_reviews=float(i[3])
if (n_reviews ==reviews_max[name]) and (name not in already_added): #two condition should be met.
android_clean.append(i)
already_added.append(name)
print(already_added)
len(android_clean)
| assets/projects/DQ_App_google/Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Tracing test
import numpy as np
import matplotlib.pyplot as plt
from ttim import *
# Start well with `tsandQ=[(0, -Q), (100, Q), (200, -Q)]`
Q = 100
npor = 0.3
k = 10
H = 10
T = k * H
Ss = 1e-4
S = Ss * H
ml = ModelMaq(kaq=[k], z=[H, 0], Saq=[Ss], tmin=0.01, tmax=1000, M=10)
w = Well(ml, xw=0, yw=0, tsandQ=[(0, -Q), (100, Q), (200, -Q)], rw=0.3)
ml.solve()
trace = timtrace(ml, xstart=0.3, ystart=0, zstart=0,
tstartend=[0, 100, 200, 400], tstartoffset=0.01, deltlist=10,
nstepmax=120, hstepmax=2, correctionstep=True)
xyzt = trace['trace']
plt.plot(xyzt[:, 3], xyzt[:, 0])
plt.grid()
trace['status'], trace['message']
# #### Terminates at well
ml = ModelMaq(kaq=[k], z=[H, 0], Saq=[Ss], tmin=0.01, tmax=1000, M=10)
w = Well(ml, xw=0, yw=0, tsandQ=[(0, -Q), (100, Q)], rw=0.3, label='testwell')
ml.solve()
trace = timtrace(ml, xstart=0.3, ystart=0, zstart=0,
tstartend=[0, 100, 300], tstartoffset=0.01, deltlist=10,
nstepmax=120, hstepmax=2, correctionstep=True)
xyzt = trace['trace']
print(trace['status'], trace['message'])
print(xyzt[-3:])
# #### Compare to numerical integration of velocity from Theis equation
# \begin{equation}
# Q_r = -T\frac{\partial h}{\partial r} = -T\frac{\text{d} h}{\text{d} u}\frac{\partial u}{\partial r} = -\frac{Q}{2\pi} \frac{1}{r}\exp(-u)
# \end{equation}
# \begin{equation}
# u = \frac{S r^2}{4T(t-t_0)}
# \end{equation}
def vxytheis(t, xy):
Q = -100
npor = 0.3
k = 10
H = 10
Ss = 1e-4
T = k * H
S = Ss * H
x, y = xy
r = np.sqrt(x ** 2 + y ** 2)
u = S * r ** 2 / (4 * T * t)
Qr = -Q / (2 * np.pi) / r * np.exp(-u)
vr = Qr / (H * npor)
vx = vr * x / r
vy = vr * y / r
return np.array([vx, vy])
from scipy.integrate import solve_ivp
t = np.linspace(0.01, 100, 100)
path0 = solve_ivp(vxytheis, (0.01, 100), y0=[0.3, 0], t_eval=t)
plt.plot(path0.t, path0.y[0]);
# #### Velocities in multi-layer system
Q = 100
npor = 0.3
k0 = 10
k1 = 40
H = 10
Hstar = 2
c = 100
T = k * H
Ss = 1e-4
S = Ss * H
ml = ModelMaq(kaq=[k0, k1], z=[2 * H + Hstar, H + Hstar, H, 0], c=[c], Saq=[Ss], tmin=0.01, tmax=10000, M=10)
w = Well(ml, xw=0, yw=0, tsandQ=[(0, -Q)], layers=0, rw=0.3)
ml.solve()
for z in np.arange(13, 22):
trace = timtrace(ml, xstart=0.3, ystart=0, zstart=z,
tstartend=[0, 1000], tstartoffset=0.01, deltlist=10,
nstepmax=500, hstepmax=2, correctionstep=True, silent=True)
xyzt = trace['trace']
plt.plot(xyzt[:, 0], xyzt[:, 2])
# #### Model3D
Q = 100
npor = 0.3
k = 10
H = 10
Hstar = 2
c = 1000
T = k * H
#Ss = 1e-4
Ss = 1e-8
S = Ss * H
ml = Model3D(kaq=10, z=[20, 15, 10, 5, 0], tmin=0.01, tmax=1000)
w = Well(ml, xw=0, yw=0, tsandQ=[(0, -Q), (100, Q)], layers=1, rw=0.1)
ml.solve()
zstart = np.linspace(10.01, 14.99, 31)
for zs in zstart:
trace = timtrace(ml, xstart=0.1, ystart=.1, zstart=zs,
tstartend=[0, 100], tstartoffset=0.01, deltlist=5,
nstepmax=40, hstepmax=2, silent=True)
xyzt = trace['trace']
plt.plot(xyzt[:, 0], xyzt[:, 2])
for y in [5, 10, 15]:
plt.axhline(y, color='k')
plt.axis('scaled');
zstart = np.linspace(10.01, 14.99, 31)
for zs in zstart:
trace = timtrace(ml, xstart=0.1, ystart=0.1, zstart=zs,
tstartend=[0, 100, 200], tstartoffset=0.01, deltlist=5,
nstepmax=40, hstepmax=2, silent=True, correctionstep=True)
xyzt = trace['trace']
plt.plot(xyzt[:, 0], xyzt[:, 2])
for y in [5, 10, 15]:
plt.axhline(y, color='k')
plt.axis('scaled');
| notebooks/pathline_trace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Categorical Features
data = [
{'price': 850000, 'rooms': 4, 'neighbourhood': 'Queen Anne'},
{'price': 700000, 'rooms': 3, 'neighborhood': 'Fremont'},
{'price': 650000, 'rooms': 3, 'neighborhood': 'Wallingford'},
{'price': 600000, 'rooms': 2, 'neighborhood': 'Fremont'}
]
# __Using one hot encoding__ When your data comes as a list of dictionaries, Scikit-Learn’s `DictVectorizer` will do this
from sklearn.feature_extraction import DictVectorizer
vec = DictVectorizer(sparse=False, dtype=int)
vec.fit_transform(data)
vec.get_feature_names()
# using sparse for efficient solution
vec = DictVectorizer(sparse=True, dtype=int)
vec.fit_transform(data)
# __Many (though not yet all) of the Scikit-Learn estimators accept such sparse inputs when fitting and evaluating models. sklearn.preprocessing.OneHotEncoder and sklearn.feature_extraction.FeatureHasher are two additional tools that Scikit-Learn includes to support this type of encoding.__
# ### Text Features
sample = ['problem of evil',
'evil queen',
'horizon problem']
# +
from sklearn.feature_extraction.text import CountVectorizer
vec = CountVectorizer()
X = vec.fit_transform(sample)
X
# -
vec.get_feature_names()
import pandas as pd
pd.DataFrame(X.toarray(), columns=vec.get_feature_names())
# __There are some issues with this approach, however: the raw word counts lead to features that put too much weight on words that appear very frequently. The tf-idf value increases proportionally to the number of times a word appears in the document, but is often offset by the frequency of the word in the corpus, which helps to adjust for the fact that some words appear more frequently in general.__
from sklearn.feature_extraction.text import TfidfVectorizer
vec = TfidfVectorizer()
X = vec.fit_transform(sample)
pd.DataFrame(X.toarray(), columns=vec.get_feature_names())
# ### Image Features
# ### Derived Features
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.array([1,2,3,4,5])
y = np.array([4,2,1,3,7])
plt.scatter(x,y);
# -
from sklearn.linear_model import LinearRegression
X = x[:, np.newaxis]
model = LinearRegression().fit(X, y)
yfit = model.predict(X)
plt.scatter(x, y)
plt.plot(x, yfit);
# __adding polynomial features__
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(degree=3, include_bias=False)
X2 = poly.fit_transform(X)
print(X2)
model = LinearRegression().fit(X2, y)
yfit = model.predict(X2)
plt.scatter(x, y)
plt.plot(x, yfit)
# ### Imputation of Missing Data
from numpy import nan
X = np.array([[nan, 0, 3 ],
[ 3, 7, 9 ],
[ 3, 5, 2 ],
[ 4, nan, 6 ],
[ 8, 8, 1 ]])
y = np.array([14, 16, -1, 8, -5])
from sklearn.preprocessing import Imputer
imp = Imputer(strategy='mean')
X2 = imp.fit_transform(X)
X2
model = LinearRegression().fit(X2, y)
model.predict(X2)
# ### Feature Pipeline
# +
from sklearn.pipeline import make_pipeline
model = make_pipeline(Imputer(strategy='mean'),
PolynomialFeatures(degree=2),
LinearRegression())
# -
model.fit(X, y) # note X with missing values
print(y)
print(model.predict(X))
import numpy as np
array = '0123456789'
ints = [1,2,3,4,5,6,7,8,9,0]
a = True
while a == True:
choice = np.random.choice(ints, 5, replace=False)
choice_rev = choice[::-1]
join_choice = (array[choice[0]]+array[choice[1]]+array[choice[2]]+
array[choice[3]]+array[choice[4]])
join_choice_rev = join_choice[::-1]
join_int = np.int32(join_choice)
join_int_rev = np.int32(join_choice_rev)
if join_int * 4 == join_int_rev:
print(join_int)
a = False
choice = np.random.choice(ints, 5, replace=False)
join_choice = (array[choice[0]]+array[choice[1]]+array[choice[2]]+
array[choice[3]]+array[choice[4]])
join_int = np.int32(join_choice)
if
join_choice[::-1]
| Chapter5-Machine Learning/Feature Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import random
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_error
import tensorflow as tf
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
from tensorflow.keras.callbacks import *
from tensorflow.keras.optimizers import *
from tensorflow.keras import backend as K
from kerashypetune import KerasGridSearch
# +
### READ DATA ###
df = pd.read_csv('Punta_Salute_2009.csv', sep=';')
df = df.dropna()
print(df.shape)
df.head()
# +
### PLOT WEEKLY TREND ###
df[:7*24]['Livello P.Salute Canal Grande (cm)'].plot(
y='Livello P.Salute Canal Grande (cm)', x='Ora solare', figsize=(8,6))
# +
### DEFINE T2V LAYER ###
class T2V(Layer):
def __init__(self, output_dim=None, **kwargs):
self.output_dim = output_dim
super(T2V, self).__init__(**kwargs)
def build(self, input_shape):
self.W = self.add_weight(name='W',
shape=(input_shape[-1], self.output_dim),
initializer='uniform',
trainable=True)
self.P = self.add_weight(name='P',
shape=(input_shape[1], self.output_dim),
initializer='uniform',
trainable=True)
self.w = self.add_weight(name='w',
shape=(input_shape[1], 1),
initializer='uniform',
trainable=True)
self.p = self.add_weight(name='p',
shape=(input_shape[1], 1),
initializer='uniform',
trainable=True)
super(T2V, self).build(input_shape)
def call(self, x):
original = self.w * x + self.p
sin_trans = K.sin(K.dot(x, self.W) + self.P)
return K.concatenate([sin_trans, original], -1)
# +
### CREATE GENERATOR FOR LSTM AND T2V ###
sequence_length = 24
def gen_sequence(id_df, seq_length, seq_cols):
data_matrix = id_df[seq_cols].values
num_elements = data_matrix.shape[0]
for start, stop in zip(range(0, num_elements-seq_length), range(seq_length, num_elements)):
yield data_matrix[start:stop, :]
def gen_labels(id_df, seq_length, label):
data_matrix = id_df[label].values
num_elements = data_matrix.shape[0]
return data_matrix[seq_length:num_elements, :]
# +
### DEFINE MODEL STRUCTURES ###
def set_seed(seed):
tf.random.set_seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
random.seed(seed)
def T2V_NN(param, dim):
set_seed(33)
inp = Input(shape=(dim,1))
x = T2V(param['t2v_dim'])(inp)
x = LSTM(param['unit'], activation=param['act'])(x)
x = Dense(1)(x)
m = Model(inp, x)
m.compile(loss='mse', optimizer=Adam(lr=param['lr']))
return m
def NN(param, dim):
set_seed(33)
inp = Input(shape=(dim,1))
x = LSTM(param['unit'], activation=param['act'])(inp)
x = Dense(1)(x)
m = Model(inp, x)
m.compile(loss='mse', optimizer=Adam(lr=param['lr']))
return m
# +
### PREPARE DATA TO FEED MODELS ###
X, Y = [], []
for sequence in gen_sequence(df, sequence_length, ['Livello P.Salute Canal Grande (cm)']):
X.append(sequence)
for sequence in gen_labels(df, sequence_length, ['Livello P.Salute Canal Grande (cm)']):
Y.append(sequence)
X = np.asarray(X)
Y = np.asarray(Y)
# +
### TRAIN TEST SPLIT ###
train_dim = int(0.7*len(df))
X_train, X_test = X[:train_dim], X[train_dim:]
y_train, y_test = Y[:train_dim], Y[train_dim:]
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# +
### DEFINE PARAM GRID FOR HYPERPARM OPTIMIZATION ###
param_grid = {
'unit': [64,32],
't2v_dim': [128,64],
'lr': [1e-2,1e-3],
'act': ['elu','relu'],
'epochs': 200,
'batch_size': [512,1024]
}
# +
### FIT T2V + LSTM ###
es = EarlyStopping(patience=5, verbose=0, min_delta=0.001, monitor='val_loss', mode='auto', restore_best_weights=True)
hypermodel = lambda x: T2V_NN(param=x, dim=sequence_length)
kgs_t2v = KerasGridSearch(hypermodel, param_grid, monitor='val_loss', greater_is_better=False, tuner_verbose=1)
kgs_t2v.search(X_train, y_train, validation_split=0.2, callbacks=[es], shuffle=False)
# -
pred_t2v = kgs_t2v.best_model.predict(X_test).ravel()
mean_absolute_error(y_test.ravel(), pred_t2v)
# +
### VISUALIZE TEST PREDICTIONS ###
plt.figure(figsize=(8,5))
plt.plot(pred_t2v[:365], label='prediction')
plt.plot(y_test.ravel()[:365], label='true')
plt.title('T2V plus LSTM'); plt.legend()
# +
### FIT SIMPLE LSTM ###
del param_grid['t2v_dim']
es = EarlyStopping(patience=5, verbose=0, min_delta=0.001, monitor='val_loss', mode='auto', restore_best_weights=True)
hypermodel = lambda x: NN(param=x, dim=sequence_length)
kgs = KerasGridSearch(hypermodel, param_grid, monitor='val_loss', greater_is_better=False, tuner_verbose=1)
kgs.search(X_train, y_train, validation_split=0.2, callbacks=[es], shuffle=False)
# -
pred_nn = kgs.best_model.predict(X_test).ravel()
mean_absolute_error(y_test.ravel(), pred_nn)
# +
### VISUALIZE TEST PREDICTIONS ###
plt.figure(figsize=(8,5))
plt.plot(pred_nn[:365], label='prediction')
plt.plot(y_test.ravel()[:365], label='true')
plt.title('single LSTM'); plt.legend()
| Time2Vec/Time2Vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 列表List
# - 一个列表可以储存任意大小的数据集合,你可以理解为他是一个容器
def b():
pass
a = [1,1.5,b,'abc',True,[1,2,[1,8]]]#可以放任何东西
a
c = 'adv'#可迭代对象才可以强制转化为list
list(c)
"".join(['a','d','v'])
a = [100,5]
b = [1,2,5,[100,5]]
a in b
a = [100,5]
b = [1,2,5,[a]]
a in b
a = [1,5]
b = [1,2,5,[100,5]]
a + b
# ## 先来一个例子爽一爽
# 
# ## 创建一个列表
# - a = [1,2,3,4,5]
# ## 列表的一般操作
# 
a = [10,5]
a*5
a = [1,5,5,9,4]
a
素雅 = [1,3,5,[3,5]]
len(素雅)
a = [1,5,[8]]
while i<len(a):
type(i)
i+=1
# # 列表索引操作
# - Mylist[index]
# - 正序索引,逆序索引
# - 列表一定注意越界
# - 
a[3:1:-1]
a = [1,2,3,4,5,[100,200]]
a[5][0]#第一个[]是取小列表第二个是取小列表中的元素
a = [1,2,3,4,5,[100,200,[1000,[4000]]]]
a[5][2][1][0]
b = [1,2,3]
b[1]=100
b#索引赋值
# ## 列表切片操作
# - Mylist[start:end]
# - 正序切片,逆序切片
c = [1,5,[5,6],9]
count = 0
for i in c:
if type(i) == list:
for j in i:
count += 1
else:
count += 1
print(count)
b = [1,2,3,4,5,6,7,8,9,10]
for i in range(0,10,2):
b[i]=100
b
# ## 列表 +、*、in 、not in
# ## 使用for循环遍历元素
# - for 循环可以遍历一切可迭代元素
b = [1,2,3,4,5,6,7,8,9,10,11]
for i in range(0,10,3):
print(b[i:i+2])
# ## EP:
# - 使用while 循环遍历列表
# ## 列表的比较
# - \>,<,>=,<=,==,!=
# +
#在列表中进行比大小,求和都必须保证列表中的元素都可进行运算
# -
# ## 列表生成式
# [x for x in range(10)]
# ## 列表的方法**
# 
a = [1,2,3]
b = [11,22]
a.append(b)#()只能加一个元素 添加元素到列表结尾
a
a = [1,2,[5,6],3]
a.count(1)#计算只看第一级
a = [1,2,3]
b = [11,22]
a.extend(b)#合并两个列表,与加不同的是改变了原数据,而加不改变原数据
a
c = [1,2,3]
c.index(1)#元素第一次出现的位置
a = [1,2,3]
a.insert(0,100)#插入元素
a
b = [1,2,3,4,5,6,7,8,9,10,11,12,13]
for i in range(0,len(b)+3,3):
b.insert(i,100)
b
d = [1,2,3,4]
d.reverse()
d
d = [1,2,4,3]
d.sort()
d
d = [1,2,3,4]
d.sort(reverse=True)
d
a = [1,2,3]
a.pop()
a
a = 'a b c d'
a.split(' ')#拆分
a = 'a b c d'
a.split(' ',2)
a = 'ab cd'
a.split(' ')
# ## 将字符串分割成列表
# - split 按照自定义的内容拆分
# ## EP:**********
# 
# 
import random
a = [1,2,4,5]
random.shuffle(a)#随机打乱列表中的数据
a
# ## 列表的复制
# - copy 浅复制
# - deepcopy import copy 深复制
# - http://www.pythontutor.com/visualize.html#mode=edit
import copy
a = [1,2,3]
b = a
b
a[0] = 100
a
c = [1,2]
d = c.copy()
d
e = [1,2,3,[0,1]]
f = e.copy()#复制不会复制子列表的,出来的还是原列表的子列表,所以对原列表改变f也会改变,浅复制
f
e[3][0] = 100
e
f
a = [1,2,3,[0,1]]
b = copy.deepcopy(a)#深复制,可以完全复制,原列表改变,复制后的列表不会因为原列表改变而改变
b
a[3][0] = 100
b
# ## 列表排序
# - sort 会改变原列表
# - sorted 不会改变原列表
# - 列表的多级排序
# - 匿名函数
sorted([])
(lambda x:print(x))(100)#匿名函数,冒号前传入量,冒号后传出量
c = [1,3,4]
c.sort()
c = [['suya',11],['flq',10],['yjy',12]]
c.sort(key=lambda x:x[1])#没有匿名函数前默认列表位置排列,有函数后【】中输入1,则按照数字高低排列,由低到高
c
c = [['syya',100,[20,80]]
,['flq',90[40,50]]
,['wj',98[50,48]]
,['jy',89[56,33]]]
c.sort()
a = [1, 6, 7, 2, 9, 4]
for i in range(len(a)-1, 1, -1):
for j in range(0, i):
if a[j] > a[j+1]:
a[j], a[j+1] = a[j+1], a[j]
print(a)
1504019140092
b = [1,5,0,4,0,1,9,1,4,0,9,2]
length = len(b)
count = 0
for i in range(length):
for j in range(length-i-1):#一次锁定一个最大值。
count += 1
if b[j]>b[j+1]:
b[j],b[j+1] = b[j+1],b[j]
b
print(count)
# ## EP:
# - 手动排序该列表[5,3,8,0,17],以升序或者降序
# - 1
# 
score = [40,55,70,58]
for i in score:
a = max(score)
if i>=(a-10):
print('分数',i,'级别为A')
if i>=(a-20):
print('分数',i,'级别为B')
if i>=(a-30):
print('分数',i,'级别为C')
if i>=(a-40):
print('分数',i,'级别为D')
else:
print('级别为F')
# - 2
# 
a = input(">>")
a[::-1]#切边的使用
# - 3
# 
a = [1,2,5,8,2,45,32,5,3]
for i in a:#i在列表a中循环
print(i,'出现',a.count(i))#打印i和在a中出现的次数
# - 4
# 
score = [51,89,95,65,75,98,25,45,62]
count = 0
for i in score:#i在列表score中循环
b = sum(score)#列表元素求和
c = len(score)#列表长度
if i<(b/c):#如果一个数小于平均数
count += 1#计数一次
print(b/c,count)
# - 5
# 
import random
y = [random.randint(0,9) for i in range(1000)]#列表生成式,产生1000个0到9的随机数
x = set(y)#定义为集合
for j in x:#j在集合x中循环
print(j,y.count(j))#打印j和j在列表y中重复的次数
# - 6
# 
def indexo(a):#定义有参数的函数
c = min(a) #取列表最小值
b = a.index(c)#对最小值返回位置的下标
print(b)
indexo(a=[9,5,6])
# - 7
# 
# 
import random
def shuffle(lst):
a = random.randint(lst)#产生随机数
b = random.randint(lst)#产生随机数
lst.insert(a,b)#在不超出列表长度下随机替换数据
print(lst)
shuffle(lst=[1,5,8,6,23,5,51,58,15,321,56])
# - 8
# 
x = [5,8,9,'r',4,'r','d']
y = list(set(x))#使用集合
print (y)
# - 9
# 
x = [5,1,2]
x.sort()
x
# - 10
# 
a = [1,2,8,6,4,12,7,8,5,3]
for i in range(len(a)-1, 1, -1):
for j in range(0, i):
if a[j] > a[j+1]:
a[j], a[j+1] = a[j+1], a[j]
print(a)
# - 11
# 
# - 12
# 
def isC(num):
count = 0
for i in range(0,len(num)):
if num[i]==num[i+1]:
count += 1
if count>=4:
print(num[i])
else:
count=0
isC(num=[1,2,2,2,2,3,5,7,7,7])
| 7.25.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
sample_data = pd.read_csv('puzzle1(2).csv')
sample_data
sample_data.iloc[0:8,2]
notation = []
for i in range(8):
for j in range(7):
if sample_data.iloc[i,j] == 0:
sample_data.iloc[i,j] = "x"
notation.append(sample_data.iloc[i,j])
sample_data
values = ''.join(str(v) for v in notation)
values
| csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 2</font>
#
# ## Download: http://github.com/dsacademybr
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# ## Variáveis e Operadores
# * usadas para armazenar valores
# * são espaços na memoria para armazenar
# Atribuindo o valor 1 à variável var_teste
var_teste = 1
# Imprimindo o valor da variável
var_teste
# Imprimindo o valor da variável
print(var_teste)
# Não podemos utilizar uma variável que não foi definida. Veja a mensagem de erro.
my_var
var_teste = 2
var_teste
type(var_teste)
var_teste = 9.5
type(var_teste)
x = 1
x
# ## Declaração Múltipla
pessoa1, pessoa2, pessoa3 = "Maria", "José", "Tobias"
pessoa1
pessoa2
pessoa3
fruta1 = fruta2 = fruta3 = "Laranja"
fruta1
fruta2
# Fique atento!!! Python é case-sensitive. Criamos a variável fruta2, mas não a variável Fruta2.
# Letras maiúsculas e minúsculas tem diferença no nome da variável.
Fruta2
# ## Pode-se usar letras, números e underline (mas não se pode começar com números)
x1 = 50
x1
# Mensagem de erro, pois o Python não permite nomes de variáveis que iniciem com números
1x = 50
# ## Não se pode usar palavras reservadas como nome de variável
#
# ## False
# ## class
# ## finally
# ## is
# ## return
# ## None
# ## continue
# ## for
# ## lambda
# ## try
# ## True
# ## def
# ## from
# ## nonlocal
# ## while
# ## and
# ## del
# ## global
# ## not
# ## with
# ## as
# ## elif
# ## if
# ## or
# ## yield
# ## assert
# ## else
# ## import
# ## pass
# ## break
# ## except
# ## in
# ## raise
# Não podemos usar palavras reservadas como nome de variável
break = 1
# ## Variáveis atribuídas a outras variáveis e ordem dos operadores
largura = 2
altura = 4
area = largura * altura
area
perimetro = 2 * largura + 2 * altura
perimetro
# A ordem dos operadores é a mesma seguida na Matemática
perimetro = 2 * (largura + 2) * altura
perimetro
# ## Operações com variáveis
idade1 = 25
idade2 = 35
idade1 + idade2
idade2 - idade1
idade2 * idade1
idade2 / idade1
idade2 % idade1
# ## Concatenação de Variáveis
nome = "Steve"
sobrenome = "Jobs"
fullName = nome + " " + sobrenome
fullName
# # Fim
# ### Obrigado
#
# ### Visite o Blog da Data Science Academy - <a href="http://blog.dsacademy.com.br">Blog DSA</a>
| Data Science Academy/Python Fundamentos/Cap02/Notebooks/DSA-Python-Cap02-02-Variaveis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Configuration
# First, we specify the Slurm server and login information:
# +
import slurmqueen
nots = slurmqueen.SlurmServer('nots.rice.edu', 'jmd11', 'C:/Users/Jeffrey/.ssh/id_rsa')
# -
# `nots.rice.edu` is the Slurm server to connect to. `jmd11` is the account to use on the server. `C:/Users/Jeffrey/.ssh/id_rsa` is an SSH private key used to connect with the server; the corresponding public key should be added to the Slurm server (see [here](https://adamdehaven.com/blog/how-to-generate-an-ssh-key-and-add-your-public-key-to-the-server-for-authentication/) for details on generation and use of SSH keys).
# Next, we choose a partition on the Slurm cluster (e.g. `commons`), a local directory on the current machine (e.g. `C:/Work/Projects/SlurmQueen/example`) and a remote directory on the Slurm cluster (e.g. `/scratch/jmd11/experiments/slurmqueen`). The remote directory should generally be in the scratch filesystem of the Slurm cluster if possible. These directories will be created when a job is started if they do not currently exist.
config = slurmqueen.ExperimentConfig(
server=nots,
partition='scavenge',
local_directory='C:/Work/Projects/SlurmQueen/example',
remote_directory='/scratch/jmd11/experiments/slurmqueen')
# # Defining an experiment
# When running many experiments on the same tool, it is convenient to define a subclass of ```SlurmExperiment``` to hold any additional server setup. The positional arguments to SlurmExperiment are, in order:
# 1. A path (relative to ```local_directory``` and ```remote_directory```) where the input and output files for this experiment should be stored.
# 2. The command to use to run the tool. In this case, ```example_tool.py``` is run through python.
# 3. The list of tasks to execute for this experiment; see below.
# 4. A list of file dependencies for this tool. Note that each can also be a Unix glob to capture multiple files. In this case, our example tool requires a single file to run: ```example_tool.py```.
#
# Our example tool requires Python 3 to be installed on the cluster. We satisfy this dependency by loading the ```Anaconda3/5.0.0``` module on the cluster before running the tool. The string passed to `setup_commands` is copied directly to the [script used to eventually submit the Slurm job](https://github.com/Kasekopf/SlurmQueen/blob/master/example/experiments/slurm_test_1/_run.sh), after pregenerated SBATCH arguments but before any tasks are executed. Custom arguments to `sbatch` can also be provided here, e.g. `#SBATCH --mem=0` to allow full use of the node memory.
#
# By default, everything written to stdout by the tool will be stored in a ```.out``` file and automatically parsed to be queried with SQL, while everything written to stderr by the tool will be stored in a ```.log``` file and not parsed. This behavior can be adjusted by including the optional arguments ```output_argument``` (defaults to ```'>>'``` indicating stdout) and ```log_argument``` (defaults to ```'2>'```, indicating stderr) when initializing the SlurmExperiment.
class SlurmTest(slurmqueen.SlurmExperiment):
def __init__(self, experiment_id, changing_args):
slurmqueen.SlurmExperiment.__init__(self, 'experiments/' + experiment_id,
'python3 example_tool.py',
changing_args,
dependencies=['example_tool.py'],
setup_commands="module load Anaconda3/5.0.0")
# We can then define a single experiment on ```example_tool.py``` by providing a name for this experiment and a list of tasks. Each task is defined by a set of arguments, given by a dictionary of key/value pairs. These arguments are passed as arguments to the tools as ```--key=value``` arguments, with a few exceptions:
# * The key ```''```, if it exists, indicates a list of positional arguments (which are given in the provided order).
# * Keys that contain ```<``` or ```>``` are treated as shell redirections. For example, the pair ```'<': 'path/to/file'``` opens ```path/to/file``` as stdin.
# * Keys that begin with ```'|'``` are not passed to each task. Such keys can be used when processing results.
slurm_test_1 = SlurmTest('slurm_test_1',
[{'': [chr(a+65) + chr(b+65)],
'a': a, 'b': b,
'|desc': '%d + %d' % (a, b)
} for a in range(3) for b in range(3)])
# ## Running an experiment
#
# Once we have defined an experiment and a configuration, we can run the experiment on the provided cluster in a single command. In this case, we run the 9 tasks using 2 Slurm workers (on two separate nodes in the cluster). Each worker is given a timeout of 5 minutes. The following command performs each of the following steps:
# 1. Creates a set of 9 .in files in ```experiments/slurm_test_1/```, each defining a single task.
# 2. Copies all .in files and all dependencies provided to the ```SlurmExperiment``` to the Slurm cluster (in ```remote_directory```)
# 3. Generates an appropriate Slurm script to run all tasks on the provided number of workers (distributed in round-robin).
# 4. Submits the Slurm job, returning the job id.
slurm_test_1.slurm_instance(config).run(2, '5:00')
# `run` also accepts an optional argument not shown here, `cpus_per_worker`, to indicate the number of cpus to request on the Slurm node allocated to each worker (default `1`).
# Once an experiment has finished running, we use a single additional command to download all results back to the local machine (and clean up the files on the cluster).
slurm_test_1.slurm_instance(config).complete()
# We can then use an SQL interface to query the results. All inputs and results appear in a table named `data`. The columns are all the named argument keys (`a`, `b`, and `|desc`), all the output keys (`Repeated Text` and `Sum`), and a column automatically generated by SlurmQueen (`file`, indicating the task id).
slurm_test_1.slurm_instance(config).query('SELECT * FROM data')
# ## (Optional) Generating/Analyzing an experiment without a Slurm cluster.
#
# To aid in reproducability, an experiment can be generated and analyzed even without access to a Slurm cluster. In particular, the ```Experiment``` class (in ```experiment.py```) is sufficient to generate all ```*.in``` files and provide an SQL interface to results without requiring access to a Slurm cluster.
#
# We can define an experiment here purely as the command and set of arguments.
experiment_1 = slurmqueen.Experiment('python3 example_tool.py',
[{'': [chr(a+65) + chr(b+65)],
'a': a, 'b': b,
'|desc': '%d + %d' % (a, b)
} for a in range(3) for b in range(3)])
# We then only need to provide a local directory in order to generate all ```*.in``` files.
experiment_1.instance('experiments/slurm_test_1').setup()
# Each ```*.in``` file is a complete bash script that runs a single task and produces the corresponding ```.out``` file. Once all ```*.out``` files are computated separately or previously provided, the SQL interface can still be used to find the results.
experiment_1.instance('experiments/slurm_test_1').query('SELECT * FROM data WHERE a=1')
| example/example_experimental_setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # Make Photoz test input from VIDEO SXDS data
#
# This notebook uses the HELP python environment to make a eazy-pype input catalogue:
#
# https://github.com/H-E-L-P/herschelhelp_internal
#
# To make the photoz inputs we must merge in the spectroscopic redshifts and make some trivial catalogue changes. We then need to make configuration inputs for the photoz code. This includes downloading filter response curves.
#
# Installation and running of the EAZY based code is described here:
#
# https://github.com/dunkenj/eazy-pype/
#
# This is an early test on the catalogues as part of our validation process.
#
#
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
from herschelhelp.filters import correct_galactic_extinction
from herschelhelp.external import convert_table_for_cigale
from herschelhelp_internal.masterlist import merge_catalogues, nb_merge_dist_plot, specz_merge
from herschelhelp_internal.utils import coords_to_hpidx, ebv, gen_help_id, inMoc
import numpy as np
import os
import time
from astropy.table import Table
import astropy.units as u
from astropy.coordinates import SkyCoord
# -
SUFFIX = os.environ.get('SUFFIX', time.strftime("%Y%m%d"))
SUFFIX
# !mkdir figs
hsc_bands = 'GRIZY'
vista_bands = 'JHK'# removing ZY for now
bands = ['HSC_{}'.format(b) for b in hsc_bands]
bands += ['VISTA_{}'.format(b) for b in vista_bands]
cols = ['id', 'VISTA_Ks_m_coord_ra','VISTA_Ks_m_coord_dec','VISTA_Ks_m_detect_isPrimary']
cols+=['{}_m_slot_ModelFlux_flux'.format(b.replace('K','Ks')) for b in bands]
cols+=['{}_m_slot_ModelFlux_fluxErr'.format(b.replace('K','Ks')) for b in bands]
cat = Table.read('../../dmu5/dmu5_VIDEO/data/full_reduced_cat_SXDS.fits')[cols]
cat['VISTA_Ks_m_coord_ra'].name='ra'
cat['VISTA_Ks_m_coord_dec'].name='dec'
#Some HELP functions require a help_id col
cat['id'].name='help_id'
#Rename to CIGALE/HELP format
for c in cat.colnames:
if c.endswith('m_slot_ModelFlux_flux'):
mask = cat[c]>1e10
cat[c][mask]=np.nan
f_name='{}_{}_flux'.format(
c.split('_')[0].lower(),#.replace('hsc','suprime'),
c.split('_')[1].lower()
)
cat[c].name=f_name
cat[f_name]=cat[f_name].to(u.uJy)
if c.endswith('m_slot_ModelFlux_fluxErr'):
mask = cat[c]>1e10
cat[c][mask]=np.nan
ferr_name='{}_{}_fluxerr'.format(
c.split('_')[0].lower(),#.replace('hsc','suprime'),
c.split('_')[1].lower()
)
cat[c].name=ferr_name
cat[ferr_name]=cat[ferr_name].to(u.uJy)
# +
mask = (cat['ra']<1e10) | (cat['dec']<1e10)
mask&=cat['VISTA_Ks_m_detect_isPrimary']==1
mask&=(cat['hsc_i_flux']/cat['hsc_i_fluxerr'] > 5)
mask&=(cat['vista_ks_flux']/cat['vista_ks_fluxerr'] > 5)
cat = cat[mask]
cat['ra']=cat['ra'].to(u.deg)
cat['dec']=cat['dec'].to(u.deg)
# +
#cat['zspec'].name='z_spec'
# -
# !mkdir data
# !wget http://hedam.lam.fr/HELP/dataproducts/dmu23/dmu23_XMM-LSS/data/XMM-LSS-specz-v2.91.fits.gz -P ./data/
# ## Merge Specz
specz=Table.read('./data/XMM-LSS-specz-v2.91.fits.gz')
for c in specz.colnames:
specz[c].name=c.lower()
specz['objid'].name='specz_id'
len(specz)
nb_merge_dist_plot(
SkyCoord(cat['ra'], cat['dec']),
SkyCoord(specz['ra'] * u.deg, specz['dec'] * u.deg)
)
cat = specz_merge(cat, specz, radius=1. * u.arcsec)
# +
#cat=cat[cat['specz_id']!='']
#cat['redshift']=cat['zspec']
# -
#Replace nans with -99
cat['zspec'].name='z_spec'
cat['z_spec'][np.isnan(cat['z_spec'])]=-99
#remove spaces from specz id
cat['specz_id'] = [s.replace(' ','') for s in cat['specz_id']]
#add int id col
cat['id']=np.arange(len(cat))
len(cat)
plt.hist(cat['hsc_i_flux']/cat['hsc_i_fluxerr'],bins=50,range=[0,30])
try:
z_rat=cat['hsc_z_flux']/cat['vista_z_flux']
z_m=np.isfinite(z_rat)
y_rat=cat['hsc_y_flux']/cat['vista_y_flux']
y_m=np.isfinite(y_rat)
plt.hist(z_rat[z_m],bins=40,range=[0,3],alpha=0.5,label='HSC z/VISTA Z')
plt.hist(y_rat[y_m],bins=40,range=[0,3],alpha=0.5,label='HSC y/VISTA Y')
plt.xlabel('flux ratio')
plt.legend()
plt.savefig('./figs/flux_ratios.png')
except KeyError:
print('No VISTA ZY bands')
try:
m=(y_rat>0) & (y_rat<3)
plt.hexbin(cat[m]['ra'],cat[m]['dec'],C=y_rat[m], gridsize=100)
plt.colorbar()
except NameError:
print('No VISTA ZY bands')
try:
m=(z_rat>0) & (z_rat<3)
plt.hexbin(cat[m]['ra'],cat[m]['dec'],C=z_rat[m], gridsize=100)
plt.colorbar()
except NameError:
print('No VISTA ZY bands')
# ## Add EBV column
#
# This is used to correct for galactic extinction.
cat.add_column(
ebv(cat['ra'], cat['dec'])
)
# ## Write catalogue
for c in cat.colnames:
if 'flux' in c:
m=~np.isfinite(cat[c])
cat[c][m]=-99
cat[0:5]
cat.write('./data/lsst_ir_fusion_sxds_photoz_input_{}.fits'.format(SUFFIX),overwrite=True)
'./data/lsst_ir_fusion_sxds_photoz_input_{}.fits'.format(SUFFIX)
# ## Make res file
#
# Eazy requires the filter transmission curves to be a single res file. Here we download the files from the Spanish Virtual Observatory and put them in a single file. We also label the bands according the order of the filter in the res file
# +
import numpy as np
from urllib.request import urlretrieve
import os
import matplotlib.pyplot as plt
svo_path = 'http://svo2.cab.inta-csic.es/theory/fps3/getdata.php?format=ascii&id=format=ascii&id={0}'
# filter_names = ['BOK/BASS.g', 'BOK/BASS.r', 'KPNO/MzLS.z',
# 'UKIRT/UKIDSS.J', 'UKIRT/UKIDSS.K',
# 'WISE/WISE.W1', 'WISE/WISE.W2']
filter_names = [
'Subaru/HSC.g',
'Subaru/HSC.r',
'Subaru/HSC.i',
'Subaru/HSC.z',
'Subaru/HSC.Y',
# 'Paranal/VISTA.Z',
# 'Paranal/VISTA.Y',
'Paranal/VISTA.J',
'Paranal/VISTA.H',
'Paranal/VISTA.Ks',
]
colors = plt.cm.viridis(np.linspace(0, 1, len(filter_names)))
Fig, Ax = plt.subplots(1,1, figsize=(10, 4))
for i, filt in enumerate(filter_names):
tlscp, name = os.path.split(filt)
print(filt,tlscp,name)
filter_lib, info = urlretrieve(svo_path.format(filt), f'data/{name}.dat')
wave, response = np.loadtxt(f'data/{name}.dat').T
Ax.semilogx(wave, response, color=colors[i], label=f'{name}')
Leg = Ax.legend(loc='upper left', ncol=4)
plt.show()
output_path = 'data/sxds_filters.res'
combined = open(output_path,'w')
combined_info = open(output_path+'.info','w')
translate = open('data/sxds.translate', 'w')
eazy_trnslate='data/testing/eazy/training_subset1_calc_zp.eazy.translate' #Do I also need to write this?
for ifx, filt in enumerate(filter_names):
tlscp, name = os.path.split(filt)
band = name.split('.')[-1]
data = np.loadtxt(f'data/{name}.dat', ndmin=2, skiprows=0)
name_lower=name.lower().replace('.','_')
print(band)
combined.write( '{0:>8d} {1:s} 0 0 0 {2}'.format(int(len(data[:,0])), name_lower, '\n'))
ifx+=1 #Use 1 indexing
translate.write(f'{name_lower:s}_flux F{ifx:.0f}\n')
translate.write(f'{name_lower:s}_fluxerr E{ifx:.0f}\n')
nums = range(1,len(data[:,0])+1)
wave = data[:,0]
resp = data[:,1]
worder = np.argsort(wave)
wave = wave[worder]
resp = resp[worder]
for i in range(len(nums)):
combined.write('{0:>8d} {1:>10.2f} {2:>12.8g} {3}'.format(int(nums[i]), wave[i], resp[i],'\n'))
combined.close()
combined_info.close()
translate.close()
# -
# !ls data
# !mkdir -p data/testing/all_specz/
# !ls data/testing/all_specz/
# !cp data/sxds.translate data/testing/eazy/training_subset1_calc_zp.eazy.translate
# !cp data/sxds.translate data/testing/eazy/zphot.translate
# ## Troubleshooting
#
# Here I am loading some eazy output files to check the runs have progressed properly
inCat=Table.read('./data/testing/test_subset1.cat',format='ascii')
inCat
441 in inCat['id']
outCat=Table.read('./data/testing/eazy/no_zp/test_subset1_no_zp.eazy.zout',format='ascii')
outCat
m=(outCat['z_peak']>0) & (outCat['z_spec']>0)
plt.scatter(outCat[m]['z_spec'],outCat[m]['z_peak']-outCat[m]['z_spec'],s=0.1)
m=(outCat['z_peak']>0) & (outCat['z_spec']>0)
plt.scatter(outCat[m]['z_spec'],outCat[m]['z_peak'],s=0.1)
m2=(outCat['z_peak']>0) & (outCat['z_spec']>0)&(outCat['z_peak']<2) & (outCat['z_spec']<2)
plt.hexbin(outCat[m2]['z_spec'],outCat[m2]['z_peak'], gridsize=70)
plt.xlabel('z_spec')
plt.ylabel('photoz z_peak')
m2=(outCat['z_peak']>0) & (outCat['z_spec']>0)&(outCat['z_peak']<2) & (outCat['z_spec']<2)
plt.hexbin(outCat[m2]['z_spec'],outCat[m2]['z_peak']-outCat[m2]['z_spec'], gridsize=70)
plt.xlabel('z_spec')
plt.ylabel('photoz z_peak - zspec')
| dmu6/dmu6_VIDEO/1_Make_sxds_photoz_input.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// # 99 Scala Exercises (31 to 41)
// ### 31\. Determine whether a given integer number is prime.
// +
import scala.math.sqrt
implicit class IntOps(val x: Int) {
// Simple recursive solution
def isPrime(): Boolean = {
val sqrtx = sqrt(x).toInt
@annotation.tailrec
def inner(i: Int): Boolean = {
if (i > sqrtx)
true
else if (x % i == 0)
false
else
inner(i+1)
}
inner(2)
}
// Concise solution using Streams
def isPrimeLazy(): Boolean =
!(Stream.range(2, sqrt(x).toInt).exists(x % _ == 0))
}
7.isPrime
120.isPrime
157.isPrime
157.isPrimeLazy
time { 550342.isPrime }
time { 550342.isPrimeLazy }
// -
// ### 32\. Determine the greatest common divisor of two positive integer numbers.
// Use Euclid's algorithm.
// +
@annotation.tailrec
def gcd(x: Int, y: Int): Int = {
val rem = x % y
if (rem == 0) y else gcd(y, rem)
}
gcd(36, 63)
// -
// ### 33\. Determine whether two positive integer numbers are coprime.
// Two numbers are coprime if their greatest common divisor equals 1.
// +
implicit class IntOps2(val x: Int) {
def isCoprimeTo(y: Int): Boolean = gcd(x, y) == 1
}
35 isCoprimeTo 64
32 isCoprimeTo 64
// -
// ### 34\. Calculate Euler's totient function phi(m).
// Euler's so-called totient function phi(m) is defined as the number of positive integers r (1 <= r <= m) that are coprime to m.
// +
implicit class IntOps3(val x: Int) {
def totient(): Int = (1 to x).count(x isCoprimeTo _)
}
10 totient
// -
// ### 35\. Determine the prime factors of a given positive integer.
// Construct a flat list containing the prime factors in ascending order.
// +
val primes: Stream[Int] = 2 #:: Stream.from(3).filter { i =>
primes.takeWhile(p => p * p <= i).forall(p => i % p > 0) };
implicit class IntOps4(val x: Int) {
def primeFactors(): List[Int] = {
def inner(i: Int, s: Stream[Int]): List[Int] = s match{
case _ if i isPrime => List(i)
case h#::t if i % h == 0 => h :: inner(i/h, s)
case _#::t => inner(i, t)
}
inner(x, primes)
}
}
315.primeFactors
// -
// ### 36\.Determine the prime factors of a given positive integer (2).
// Construct a list containing the prime factors and their multiplicity.
// Alternately, use a Map for the result.
// +
import scala.collection.immutable.ListMap
implicit class IntOps5(val x: Int) {
def primeFactorMultiplicity(): List[(Int, Int)] =
x.primeFactors.groupBy(identity)
.mapValues { _.length }
.toList
.sortBy(_._1)
def primeFactorMultiplicityMap(): Map[Int, Int] =
ListMap(x.primeFactors
.groupBy(identity)
.mapValues { _.length }
.toSeq
.sortBy(_._1):_*)
}
315.primeFactorMultiplicity
315.primeFactorMultiplicityMap
time { 10324525.primeFactorMultiplicity }
time { 10324525.primeFactorMultiplicityMap }
// -
// ### 37\. Calculate Euler's totient function phi(m) (improved).
// See problem P34 for the definition of Euler's totient function. If the list of the prime factors of a number m is known in the form of problem P36 then the function phi(m>) can be efficiently calculated as follows: Let [[p1, m1], [p2, m2], [p3, m3], ...] be the list of prime factors (and their multiplicities) of a given number m. Then phi(m) can be calculated with the following formula:
// $$ phi(m) = (p1-1)*p1^{(m1-1)} * (p2-1)*p2^{(m2-1)} * (p3-1)*p3^{(m3-1)} * ... $$
// +
implicit class IntOps6(val x: Int) {
def totientImproved(): Int = x.primeFactorMultiplicity.foldLeft(1) {
case (accum, (p, m)) => accum * (p-1) * Math.pow(p, m-1).toInt
}
}
time { 10000000.totientImproved }
time { 10000000.totient }
// -
// ### 38\. Compare the two methods of calculating Euler's totient function.
// Use the solutions of problems P34 and P37 to compare the algorithms. Try to calculate phi(10090) as an example.
(for (_ <- 1 to 10000) yield time(10090 totientImproved)._2.toNanos).sum / 10000
(for (_ <- 1 to 10000) yield time(10090 totient)._2.toNanos).sum / 10000
// ### 39\. A list of prime numbers.
// Given a range of integers by its lower and upper limit, construct a list of all prime numbers in that range.
// +
def listPrimesInRange(range: Seq[Int]): List[Int] =
primes.dropWhile(_ < range.head).takeWhile(_ < range.last).toList
listPrimesInRange(7 to 31)
// -
// ### 40\. Goldbach's conjecture.
// Goldbach's conjecture says that every positive even number greater than 2 is the sum of two prime numbers. E.g. 28 = 5 + 23. It is one of the most famous facts in number theory that has not been proved to be correct in the general case. It has been numerically confirmed up to very large numbers (much larger than Scala's Int can represent). Write a function to find the two prime numbers that sum up to a given even integer.
// +
implicit class IntOps7(val x: Int) {
def goldbach(): Option[(Int, Int)] = {
val p = primes.find(p => (x - p).isPrime)
p map { y => (y, x - y) }
}
}
28.goldbach
// -
// ### 41\. A list of Goldbach compositions.
// Given a range of integers by its lower and upper limit, print a list of all even numbers and their Goldbach composition.
// +
import scalaz.effect.IO._
def printGoldbachList(range: Seq[Int]): IO[Unit] = (for {
n <- range if n % 2 == 0
io = (n goldbach).fold(ioUnit) { case (p1, p2) => putStrLn(s"$n = $p1 + $p2") }
} yield io).foldLeft(ioUnit)(_ |+| _)
printGoldbachList(9 to 20).unsafePerformIO
// -
// In most cases, if an even number is written as the sum of two prime numbers, one of them is very small. Very rarely, the primes are both bigger than, say, 50. Try to find out how many such cases there are in the range 2..3000.
// +
def printGoldbachListLimited(range: Seq[Int], limit: Int): IO[Unit] = (for {
n <- range if n % 2 == 0
io = (n goldbach).fold(ioUnit) {
case (p1, p2) if p1 > limit && p2 > limit => putStrLn(s"$n = $p1 + $p2")
case _ => ioUnit
}
} yield io).foldLeft(ioUnit)(_ |+| _)
printGoldbachListLimited(1 to 2000, 50).unsafePerformIO
| Exercises31-41.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
1 + 2
2 ** 8
x = 6
x
def double(x):
return x * 2
y = double(10)
y
import numpy
numpy.add(3, 7)
numpy.binary_repr(2)
| python/Testing Installation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import sys
sys.path.append('../')
import numpy as np
from tsalib import dim_vars, get_dim_vars
# -
# # Design Principles
# **Dimension Variables** (DVs) are the core abstractions behind tsalib.
# - They allow specifying and modifying shapes of tensors *symbolically*, i.e., using named symbols corresponding to different dimensions of tensor.
# - Making dimension names explicit enables cleaner, DRY code, symbolic shape assertions, and faster debugging.
# - **Symbolic** shapes or **annotations** are *tuples* over DVs and arithmetic expressions over DVs.
#
# The `tsalib` provides a collection of powerful APIs to handle all kinds of shape transformations using explicit dimension variables and shape annotations.
#
#
# - Designed to stay light, easy to incorporate into existing workflow with minimal code changes.
# - The API includes both library-independent and dependent parts, giving developers flexibility in how they choose to incorporate `tsalib` in their workflow.
# - Avoid deeper integration into popular tensor libraries to keep `tsalib` light-weight and avoid backend-inflicted bugs.
#
# Some popular models (resnet, transformer) annotated/re-written with tsalib can be found in the [models](models/) directory.
#
# ## Declare dimension variables
# Dimension variables model both the `name` and the default `size` of a tensor.
# Format: **name(symbol):size** -- `symbol` and `size` are optional
#
# We can declare dimension variables **globally** (Dimensions used in programs are known upfront and programs don't modify dimension names).
# Even better, we can put all these definitions in the Config dictionary.
# globals variables prefixed with underscores
_B, _T, _D, _K = dim_vars('Batch(b):20 SeqLength(t):10 EmbeddingDim(d):100 K(k):1')
_C, _H, _W = dim_vars('Channels(c):3 Height(h):256 Width(w):256')
# +
def test_decls():
print('\nTest declarations ..')
#local declarations
print(f'B, C, D = {_B}, {_C}, {_D}')
#strict=False allows overwriting previous declarations
H, W = dim_vars ('Height(h):256 Width(w):256', exists_ok=True)
print(f'H, W = {H}, {W}')
# Supports arithmetic over a combination of dim vars and other Python variables
def test_arith():
print('\nTest arithmetic ..')
_K, _W, _B, _H = get_dim_vars('k w b h')
_K = _W * 2
h = 4
print((h, _H // h, _K, _B*2))
# Use dimension variables in lieu of constant size values
# note: dim_var declaration must include size of the variable
def test_cast_int():
print('\nTest integer cast ..')
B, C = get_dim_vars('b c')
x = np.zeros((B, C))
print(f'shape of array: ({B},{C}): {x.shape}')
return x
def basic_tests():
test_decls()
test_arith()
x = test_cast_int()
# Test assertions over symbolic shapes
assert x.shape == (_B,_C)
print ('assertions hold')
# -
basic_tests()
# ## Basic tsalib usage
# Can be used to manage tensor shapes with **arbitrary** tensor libraries. Here, examples with *numpy* and *pytorch*.
# - Create new tensors (independent of actual dimension sizes)
# - **Annotate** tensor variables (widely considered best practice, otherwise done using adhoc comments)
# - Check symbolic **assertions** (assertions **do not** change even if dimension size changes)
# +
def test_numpy():
print('\nTest usage with numpy ..')
B, D = get_dim_vars('b d')
import numpy as np
a: (B, D) = np.zeros((B,D))
print(f'original array: {(B,D)}: {a.shape}')
b: (2, B, D) = np.stack([a, a])
print(f'after stack: {(2,B,D)}: {b.shape}')
ax = (2,B,D).index(B)
c: (2, D) = np.mean(b, axis=ax)
print(f'after mean along axis = {ax}: {(2,D)}: {c.shape}')
test_numpy()
# +
def test_pytorch():
print('\nTest usage with pytorch ..')
B, D = get_dim_vars('b d')
B, D = dim_vars('Batch:2 EmbedDim:3', exists_ok=True)
import torch
a = torch.Tensor([[1., 2., 4.], [3., 6., 9.]])
assert a.size() == (B, D)
b = torch.stack([a, a])
print ('Asserting b.size() == (2,B,D)')
assert b.size() == (2, B, D)
c = torch.cat([a, a], dim=1)
print ('Assertion on c.size()')
assert c.size() == (B, D*2)
test_pytorch()
# -
# ## Shape Transformations with Dimensions Variables
# To shape transform without `tsalib`, you either
# - **hard-code** integer constants for each dimension's position in shape transformations, or
# - do shape tuple **surgeries** to compute the 'right' shape (for the general case)
#
# Instead, with `tsalib`, use dimension variables or the shorthand symbols directly.
#
# `tsalib` provides API for common shape transformations: **view** (reshape), **permute** (transpose) and **expand** (tile).
# These are *library-independent*, e.g., shorthand transformation -> target shape tuple -> reshape.
#
# One transformation to rule them all : **warp**. Do a sequence of transformations on a tensor.
# `warp` is implementated for several popular backend libraries.
#
# ## Work with Shorthand Shape Notation
# Writing tuples of shape annotations can get cumbersome.
#
# So, instead of (B, T, D), write 'btd' (each dim gets a single char, concatenated together)
#
# Instead of (B \* T, D // 2, T), write 'b * t, d//2, t' (arbitrary arithmetic expressions, comma-separated)
#
# Anonymous dimension variables : 'b,,d' omits naming dimension t.
# ## Reshapes (view transformations) using dimension variables
# These are library independent: `vt` returns target tensor shapes from shorthand transformation spec.
# +
# without tsalib, this is how we used to do it. See code from BERT.
def test_reshape_old ():
x = np.ones((20, 10, 100))
h = 4
new_shape = x.shape[:2] + (h, x.shape[2]//h) #shape surgery
x = x.reshape(new_shape)
print (x.shape)
from tsalib import view_transform as vt
# with tsalib, simply use dimension vars in-place
def test_reshape():
B, T, D = get_dim_vars('b t d')
x: (B,T,D) = np.ones((B, T, D))
h = 4
x: (B,T,h,D//h) = x.reshape((B, T, h, D//h))
assert x.shape == (B,T,h,D//h)
print ('test_reshape: all assertions hold')
#using shorthand notation, omit dimensions not involved in transformation
def test_reshape_short():
B, T, D = get_dim_vars('b t d')
x: (B,T,D) = np.ones((B, T, D))
h = 4
x = x.reshape(vt(f'btd -> b,t,{h},d//{h}', x.shape))
assert x.shape == (B, T, h, D//h)
x1 = x.reshape(vt('b,t,4,k -> b*t,4,k', x.shape))
assert x1.shape == (B*T, h, D//h)
x1 = x.reshape(vt('b,t,, -> b*t,,', x.shape))
assert x1.shape == (B*T, h, D//h)
print ('test_reshape_short: all assertions hold')
#test_reshape_old()
test_reshape()
test_reshape_short()
# -
# ## Transpose/Permute transformations using dimension variables
# +
from tsalib import permute_transform as pt
from tsalib.transforms import _permute_transform as _pt
# permute using dimension variables (internal, recommended to be not used)
def test_permute():
B, T, D, K = get_dim_vars('b t d k')
x: (B,T,D,K) = np.ones((B, T, D, K))
perm_indices = _pt(src=(B,T,D,K), to=(D,T,B,K))
assert perm_indices == (2,1,0,3)
x = x.transpose(perm_indices)
assert x.shape == (D,T,B,K)
print ('test_permute: all assertions hold')
# shorthand permutes are snazzier (use '_' or ',' as placeholders)
def test_permute_short():
B, T, D, K, C, H, W = get_dim_vars('b t d k c h w')
x: (B,T,D,K) = np.ones((B, T, D, K))
x = x.transpose(pt('btdk -> dtbk')) # (B, T, D, K) -> (D, T, B, K)
assert x.shape == (D,T,B,K)
x = x.transpose(pt('d_b_ -> b_d_')) # (D,T,B,K) -> (B, T, D, K)
assert x.shape == (B,T,D,K)
x: (B, C, H, W) = np.ones((B, C, H, W))
x1 = x.transpose(pt(',c,, -> ,,,c'))
assert x1.shape == (B, H, W, C)
print ('test_permute_short: all assertions hold')
test_permute()
test_permute_short()
# -
# ## Expand transformations
# +
from tsalib import _expand_transform as et
def test_expand():
B, T, D, K = get_dim_vars('b t d k')
x: (B, T, D) = np.ones((B, T, D))
x: (B, K, T, D) = x[:, None]
expand_shape = et(src=(B,K,T,D), expansions=[(K, K*5)], in_shape=x.shape) #(B, K, T, D) -> (B, K*5, T, D)
assert expand_shape == (-1,5,-1,-1)
print ('test_expand: all assertions hold')
def test_expand_short():
B, T, D, K = get_dim_vars('b t d k')
x: 'btd' = np.ones((B, T, D))
x: 'bktd' = x[:, None]
expand_shape = et(src=(B,K,T,D), expansions='k->k*5', in_shape=x.shape)
assert expand_shape == (-1,5,-1,-1)
print ('test_expand_short: all assertions hold')
test_expand()
test_expand_short()
# -
# ## *warp* : generalized shape transformations
#
# Writing a sequence of shape transformations in code can get cumbersome.
# `warp` enables specifying a sequence of transformations together **inline**.
# +
from tsalib import warp
def test_warp():
B, T, D = get_dim_vars('b t d')
x: 'btd' = np.ones((B, T, D))
# two view transformations (reshapes) in sequence
x1 = warp(x, 'btd -> b,t,4,d//4 -> b*t,4,d//4', 'vv', debug=False)
assert(x1.shape == (B*T,4,D//4))
# four reshapes in sequence
x2 = warp(x, 'btd -> b,t,4,d//4 -> b*t,4,d//4 -> b,t,4,d//4 -> btd', 'vvvv', debug=False)
assert(x2.shape == (B,T,D))
# Same reshape sequence in shorthand, specified as list of transformations
x2 = warp(x, ['__d -> ,,4,d//4', 'b,t,, -> b*t,,', 'b*t,, -> b,t,,', ',,4,d//4 -> ,,d'], 'vvvv', debug=True)
assert(x2.shape == (B,T,D))
print ('test_warp: all assertions hold')
def test_warp_pytorch():
B, T, D = get_dim_vars('b t d')
import torch
y: 'btd' = torch.randn(B, T, D)
#a reshape followed by permute
y = warp(y, 'btd -> b,t,4,d//4 -> b,4,t,d//4', 'vp', debug=False)
assert(y.shape == (B,4,T,D//4))
print ('test_warp_pytorch: all assertions hold')
test_warp()
test_warp_pytorch()
# -
# ## Join: unified stack/concatenate for a list of tensors
# Crisp shorthand : `'(b,t,d)* -> b,3*t,d'` (**concat**) or `'(b,t,d)* -> b,^,t,d'` (**stack**)
# +
from tsalib import join, join_transform
def test_join ():
B, T, D = get_dim_vars('b t d')
x1: 'btd' = np.ones((B, T, D))
x2: 'btd' = np.ones((B, T, D))
x3: 'btd' = np.ones((B, T, D))
#concatenate along the (T) dimension: (b,t,d)* -> (b,3*t,d)
x = join([x1, x2, x3], dims=',*,')
assert x.shape == (B, 3*T, D)
#stack: join by adding a new dimension to the front: (b,t,d)* -> (^,b,t,d)
x = join([x1, x2, x3], dims='^')
assert x.shape == (3, B, T, D)
#stack by adding a new dimension at second position: (b,t,d)* -> b,^,t,d)
x = join([x1, x2, x3], dims=',^')
assert x.shape == (B, 3, T, D)
print ('test_join: all assertions passed')
def test_join_transform():
B, T, D = get_dim_vars('b t d')
x1: 'btd' = np.ones((B, T, D))
x2: 'btd' = np.ones((B, T, D))
x3: 'btd' = np.ones((B, T, D))
dims = join_transform([x1,x2,x3], '(b,t,d)* -> b,3*t,d')
assert dims == ',*,'
#now use backend-dependent join
dims = join_transform([x1,x2,x3], '(b,t,d)* -> b,^,t,d')
assert dims == ',^,,'
#now use backend-dependent join
print ('test_join_transform: all assertions passed')
test_join()
test_join_transform()
# -
# ## Align one tensor to another
from tsalib import alignto
def test_align():
B, T, D = dim_vars('Batch(b):20 SeqLength(t):10 EmbeddingDim(d):100', exists_ok=True)
x1 = np.random.randn(D,D)
x2 = np.random.randn(B,D,T,D)
x1_aligned = alignto( (x1, 'dd'), 'bdtd')
assert x1_aligned.shape == (1,D,1,D)
print ('test align: all assertion passed')
test_align()
# ## Dot Product of two tensors (sharing exactly one dimension)
from tsalib import dot
import torch
def test_dot():
B, C, T, D = get_dim_vars('b c t d')
#x = np.random.rand(B, C, T)
#y = np.random.rand(C, D)
x = torch.randn(B, C, T)
y = torch.randn(C, D)
z = dot('_c_.c_', x, y)
assert z.shape == (B, T, D)
print('test_dot: all assertions passed')
test_dot()
# # Reduce ops (min, max, mean, ..) with tsalib
# Reduction operators aggregate values over one or more tensor dimensions.
# `tsalib` provides `reduce_dims` to compute dimension ids using shorthand notation.
# +
from tsalib import reduce_dims as rd
def test_reduce ():
assert rd('2bd->2d') == (1,)
assert rd('2bd->2') == (1,2)
print ('test_reduce: all assertions hold')
test_reduce()
# -
x: 'btd' = np.random.rand(_B, _T, _D)
y = np.mean(x, axis=rd('btd->b'))
assert y.shape == (_B,)
# ## Looong warps
# +
def warp_long1 ():
B, T, D, C = get_dim_vars('b t d c')
x1: 'btd' = np.ones((B, T, D))
x2: 'btd' = np.ones((B, T, D))
x3: 'btd' = np.ones((B, T, D))
y = warp([x1,x2,x3], '(btd)* -> btdc -> bdtc -> b,d//2,t*2,c', 'jpv')
assert y.shape == (B, D//2, T*2, C)
print ('warp_long1: all assertions hold')
def warp_long2 ():
B, T, D, C = get_dim_vars('b t d c')
x1: 'btd' = np.ones((B, T, D))
y = warp(x1, 'btd -> btd1 -> bdt1 -> b,d//2,t*2,1', 'apv')
assert y.shape == (B, D//2, T*2, 1)
print ('warp_long2: all assertions hold')
warp_long1()
warp_long2()
# -
| notebooks/tsalib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="ujaR5QYEIu07"
# ## Importing Libraries
# + colab={} colab_type="code" id="TZ8UU5blhizA"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="xdfcLYxOiRtl" outputId="f895ca87-1a23-4fe2-ed4c-c915c9109dc7"
# %tensorflow_version 2.x
import tensorflow as tf
# + colab={} colab_type="code" id="jMv0atutiUnQ"
import numpy as np
import matplotlib.pyplot as plt
import pathlib
import os
import pandas as pd
import seaborn as sns
import sklearn.metrics as metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="HhMzzi-ribZw" outputId="2c80dd05-987a-4849-f899-97e17253ca45"
tf.__version__
# + [markdown] colab_type="text" id="ny-oI58CJFEi"
# ## Mounting Google Drive with the notebook
# + colab={"base_uri": "https://localhost:8080/", "height": 124} colab_type="code" id="AhJ92g-kid1k" outputId="451b8b02-8659-4568-f4ed-561cb1f03527"
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
root_dir = "/content/gdrive/My Drive/"
base_dir = root_dir + 'DeepLearning/MUSK/'
# + [markdown] colab_type="text" id="QPS6ODQ2JhCL"
# ## Loading the Data in a DataFrame
# + colab={} colab_type="code" id="G3Vll204iiN0"
data_dir = pathlib.Path(base_dir)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="0R37hpHTivuy" outputId="bfffcf56-0f00-4173-d1b3-e887d1699e38"
os.path.exists(data_dir)
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" id="oqMrKiFyjQa4" outputId="06bacfe9-a4ea-4e37-cf20-d4ed75e1b77f"
for name in list(data_dir.glob('*'))[:]:
print(name)
# + colab={} colab_type="code" id="il67lPMcizkF"
data = pd.read_csv(root_dir + 'DeepLearning/MUSK/musk_csv.csv')
# + [markdown] colab_type="text" id="7H2Bzf3pKE31"
# ## Data Exploration & Preprocessing
# + colab={"base_uri": "https://localhost:8080/", "height": 251} colab_type="code" id="9BDfeYIrKCYr" outputId="5dad4552-620a-4a12-f587-df4fd01d4690"
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 338} colab_type="code" id="BjsnJo02kMM7" outputId="0cec46f6-d596-4e72-ba94-e8bd759c632b"
data.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="pvruMsI2jv5X" outputId="d414ab48-2a8d-4b35-e1e5-7dceaa384a1a"
data.shape
# + colab={} colab_type="code" id="fJxJFQ4Qj-MQ"
y = data['class']
x = data.drop(['class','ID','molecule_name','conformation_name'],axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="Hpi5WMDgk3Im" outputId="943e7ed8-1059-434a-da6a-17674e2f591e"
data['conformation_name'].nunique()
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="CaEtvrpjvYNf" outputId="a6c9a4f9-2e3d-4007-9cf9-46d9a45fbcb6"
x.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="IU-1qzcknr73" outputId="1c227fdc-2522-493c-c11f-12e19e6c29b1"
sns.countplot(x='class',data=data)
plt.title("Classes count")
# + colab={} colab_type="code" id="A-jLywkluXbq"
from sklearn.model_selection import train_test_split
np.random.seed(5)
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.2, random_state=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="-LIB3rQf96od" outputId="48222fca-53d5-45a5-fcac-36e324bf1349"
sns.countplot(x=y_train)
plt.title("Classes count")
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="tDA5vY5k-Zih" outputId="9072c919-d122-4d9d-e662-569bf071ce3d"
sns.countplot(x=y_test)
plt.title("Classes count")
# + colab={"base_uri": "https://localhost:8080/", "height": 343} colab_type="code" id="NvZdsMs1Z7c3" outputId="a6cf0043-ab23-4935-ab35-cf76aa85a3e7"
x.describe()
# + [markdown] colab_type="text" id="16EL2nDIKRTm"
# ## Model Creation
# + colab={} colab_type="code" id="mbr4cEg5okb6"
model = tf.keras.models.Sequential()
model.add(tf.keras.layers.Dense(64,input_shape=(x.shape[1],),use_bias=False))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.Dense(64,use_bias=False))
# model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.Activation('relu'))
# model.add(tf.keras.layers.Dense(64,activation='relu',input_shape=(x.shape[1],),kernel_regularizer=tf.keras.regularizers.l2(l=0.001)))
# model.add(tf.keras.layers.Dense(64,activation='relu',kernel_regularizer=tf.keras.regularizers.l2(l=0.001)))
# model.add(tf.keras.layers.Dense(64,activation='relu',kernel_regularizer=tf.keras.regularizers.l2(l=0.001)))
model.add(tf.keras.layers.Dense(1,activation='sigmoid'))
# + colab={"base_uri": "https://localhost:8080/", "height": 364} colab_type="code" id="k7wlijpDv_wZ" outputId="c7f7787c-37e3-4de4-9c48-c91f58e5c35c"
model.summary()
# + colab={} colab_type="code" id="vJbH0HScwBMw"
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss='binary_crossentropy',
metrics=['accuracy'])
# earlyStopping = tf.keras.callbacks.EarlyStopping(monitor='val_accuracy', min_delta=0.0001, patience=6)
LRR = tf.keras.callbacks.ReduceLROnPlateau(monitor='val_accuracy', patience=3, verbose=1, factor=0.5, min_lr=0.0001)
# + [markdown] colab_type="text" id="YXnIPbMWKY6p"
# ## Model Training
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="6BXo3jiAziEd" outputId="d2546e8f-df80-4962-f81b-f183ac4b40d0"
history = model.fit(X_train,y_train,batch_size=32,epochs=100,validation_data=(X_test,y_test))
# history = model.fit(X_train,y_train,batch_size=32,epochs=100,validation_data=(X_test,y_test),callbacks=[LRR])
# + colab={} colab_type="code" id="9mPNJO93g2BS"
history1 = model.fit(X_train,y_train,batch_size=32,epochs=100,validation_data=(X_test,y_test))
# + colab={} colab_type="code" id="IT8guEuQ26ej"
def displayHistory(history,name):
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('loss')
plt.ylim([0,2.0])
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.savefig(name)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 513} colab_type="code" id="tBOrvWLy28MA" outputId="09de59d5-4dc7-4578-cbd8-fe0c65af0d1a"
displayHistory(history,root_dir+'DeepLearning/MUSK/Models/model10.png')
# + [markdown] colab_type="text" id="BCHQ9SwDQMhU"
# ## Evaluation
# + colab={} colab_type="code" id="jKxzyZcOz-e8"
def calculateScores(model):
# predict probabilities for test set
yhat_probs = model.predict(X_test, verbose=0)
# predict crisp classes for test set
yhat_classes = model.predict_classes(X_test, verbose=0)
# reduce to 1d array
yhat_probs = yhat_probs[:, 0]
yhat_classes = yhat_classes[:, 0]
# accuracy: (tp + tn) / (p + n)
accuracy = metrics.accuracy_score(y_test, yhat_classes)
print('Accuracy: %f' % accuracy)
# precision tp / (tp + fp)
precision = metrics.precision_score(y_test, yhat_classes)
print('Precision: %f' % precision)
# recall: tp / (tp + fn)
recall = metrics.recall_score(y_test, yhat_classes)
print('Recall: %f' % recall)
# f1: 2 tp / (2 tp + fp + fn)
f1 = metrics.f1_score(y_test, yhat_classes)
print('F1 score: %f' % f1)
# + colab={"base_uri": "https://localhost:8080/", "height": 87} colab_type="code" id="Oi65iJ_K00Dk" outputId="db720e02-9b4d-44d1-b00f-759264969620"
calculateScores(model)
# + colab={} colab_type="code" id="0FQGWk8B1YRN"
model.save(root_dir + 'DeepLearning/MUSK/Models/model10.h5')
# + colab={} colab_type="code" id="LpMQtmBdjHKN"
model.load_weights(root_dir + 'DeepLearning/MUSK/Models/model10.h5')
# + [markdown] colab_type="text" id="auzoXBaOQR2v"
# ## Fine Tuning
# + colab={} colab_type="code" id="iTnFIQ9sgBAr"
# model.compile(optimizer=tf.keras.optimizers.Adam(),
# loss='binary_crossentropy',
# metrics=['accuracy'])
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if logs.get('val_accuracy') == 1.0:
print("\nReached at Validation accuracy of 1.0000")
self.model.stop_training = True
LRR = tf.keras.callbacks.ReduceLROnPlateau(monitor='val_accuracy', patience=3, verbose=1, factor=0.5, min_lr=0.0001)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="UDjoAO1UivGx" outputId="efd2bbf7-1460-4cc5-ab46-cd0dc945362c"
history2 = model.fit(X_train,y_train,batch_size=32,epochs=150,validation_data=(X_test,y_test),initial_epoch=100,callbacks=[LRR,myCallback()])
# + colab={"base_uri": "https://localhost:8080/", "height": 513} colab_type="code" id="fDfobC2ckYaV" outputId="23519a2a-230d-4b89-e74a-f3051f1b655e"
displayHistory(history2,root_dir+'DeepLearning/MUSK/Models/model10finetune.png')
# + colab={"base_uri": "https://localhost:8080/", "height": 87} colab_type="code" id="md-bXt_mi9YN" outputId="fb500b82-ce09-4f90-cb53-350a6cb6e910"
calculateScores(model)
# + colab={} colab_type="code" id="57PoFmsGj06b"
model.save(root_dir + 'DeepLearning/MUSK/Models/model10finetuned.h5')
| Deep Learning/MUSK & NON-MUSK Compounds Classification/MUSK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# <h1>Model Deployment and Monitoring</h1>
# Once we have built and trained our models for feature engineering (using Amazon SageMaker Processing and SKLearn) and binary classification (using the XGBoost open-source container for Amazon SageMaker), we can choose to deploy them in a pipeline on Amazon SageMaker Hosting, by creating an Inference Pipeline.
# https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html
#
# This notebook demonstrates how to create a pipeline with the SKLearn model for feature engineering and the XGBoost model for binary classification.
#
# Let's define the variables first.
# +
import sagemaker
import sys
import IPython
# Let's make sure we have the required version of the SM PySDK.
required_version = '2.46.0'
def versiontuple(v):
return tuple(map(int, (v.split("."))))
if versiontuple(sagemaker.__version__) < versiontuple(required_version):
# !{sys.executable} -m pip install -U sagemaker=={required_version}
IPython.Application.instance().kernel.do_shutdown(True)
# -
import sagemaker
print(sagemaker.__version__)
# +
import boto3
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sagemaker_session = sagemaker.Session()
bucket_name = sagemaker_session.default_bucket()
prefix = 'endtoendmlsm'
print(region)
print(role)
print(bucket_name)
# -
# ## Retrieve model artifacts
# First, we need to create two Amazon SageMaker **Model** objects, which associate the artifacts of training (serialized model artifacts in Amazon S3) to the Docker container used for inference. In order to do that, we need to get the paths to our serialized models in Amazon S3.
# <ul>
# <li>For the SKLearn model, in Step 02 (data exploration and feature engineering) we defined the path where the artifacts are saved</li>
# <li>For the XGBoost model, we need to find the path based on Amazon SageMaker's naming convention. We are going to use a utility function to get the model artifacts of the last training job matching a specific base job name.</li>
# </ul>
# +
from notebook_utilities import get_latest_training_job_name, get_training_job_s3_model_artifacts
# SKLearn model artifacts path.
sklearn_model_path = 's3://{0}/{1}/output/sklearn/model.tar.gz'.format(bucket_name, prefix)
# XGBoost model artifacts path.
training_base_job_name = 'end-to-end-ml-sm-xgb'
latest_training_job_name = get_latest_training_job_name(training_base_job_name)
xgboost_model_path = get_training_job_s3_model_artifacts(latest_training_job_name)
print('SKLearn model path: ' + sklearn_model_path)
print('XGBoost model path: ' + xgboost_model_path)
# -
# ## SKLearn Featurizer Model
# Let's build the SKLearn model. For hosting this model we also provide a custom inference script, that is used to process the inputs and outputs and execute the transform.
#
# The inference script is implemented in the `sklearn_source_dir/inference.py` file. The custom script defines:
#
# - a custom `input_fn` for pre-processing inference requests. Our input function accepts only CSV input, loads the input in a Pandas dataframe and assigns feature column names to the dataframe
# - a custom `predict_fn` for running the transform over the inputs
# - a custom `output_fn` for returning either JSON or CSV
# - a custom `model_fn` for deserializing the model
# !pygmentize sklearn_source_dir/inference.py
# Now, let's create the `SKLearnModel` object, by providing the custom script and S3 model artifacts as input.
# +
import time
from sagemaker.sklearn import SKLearnModel
code_location = 's3://{0}/{1}/code'.format(bucket_name, prefix)
sklearn_model = SKLearnModel(name='end-to-end-ml-sm-skl-model-{0}'.format(str(int(time.time()))),
model_data=sklearn_model_path,
entry_point='inference.py',
source_dir='sklearn_source_dir/',
code_location=code_location,
role=role,
sagemaker_session=sagemaker_session,
framework_version='0.20.0',
py_version='py3')
# -
# ## XGBoost Model
# Similarly to the previous steps, we can create an `XGBoost` model object. Also here, we have to provide a custom inference script.
#
# The inference script is implemented in the `xgboost_source_dir/inference.py` file. The custom script defines:
#
# - a custom `input_fn` for pre-processing inference requests. This input function is able to handle JSON requests, plus all content types supported by the default XGBoost container. For additional information please visit: https://github.com/aws/sagemaker-xgboost-container/blob/master/src/sagemaker_xgboost_container/encoder.py. The reason for adding the JSON content type is that the container-to-container default request content type in an inference pipeline is JSON.
# - a custom `model_fn` for deserializing the model
# !pygmentize xgboost_source_dir/inference.py
# Now, let's create the `XGBoostModel` object, by providing the custom script and S3 model artifacts as input.
# +
import time
from sagemaker.xgboost import XGBoostModel
code_location = 's3://{0}/{1}/code'.format(bucket_name, prefix)
xgboost_model = XGBoostModel(name='end-to-end-ml-sm-xgb-model-{0}'.format(str(int(time.time()))),
model_data=xgboost_model_path,
entry_point='inference.py',
source_dir='xgboost_source_dir/',
code_location=code_location,
framework_version='0.90-2',
py_version='py3',
role=role,
sagemaker_session=sagemaker_session)
# -
# ## Pipeline Model
# Once we have models ready, we can deploy them in a pipeline, by building a `PipelineModel` object and calling the `deploy()` method.
# Finally we create an endpoint with data capture enabled, for monitoring the model data quality. Data capture is enabled at enpoint configuration level for the Amazon SageMaker real-time endpoint. You can choose to capture the request payload, the response payload or both and captured data is stored in JSON format.
# +
import sagemaker
import time
from sagemaker.pipeline import PipelineModel
from time import gmtime, strftime
from sagemaker.model_monitor import DataCaptureConfig
s3_capture_upload_path = 's3://{}/{}/monitoring/datacapture'.format(bucket_name, prefix)
print(s3_capture_upload_path)
pipeline_model_name = 'end-to-end-ml-sm-xgb-skl-pipeline-{0}'.format(str(int(time.time())))
pipeline_model = PipelineModel(
name=pipeline_model_name,
role=role,
models=[
sklearn_model,
xgboost_model],
sagemaker_session=sagemaker_session)
endpoint_name = 'end-to-end-ml-sm-pipeline-endpoint-{0}'.format(str(int(time.time())))
print(endpoint_name)
pipeline_model.deploy(initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name=endpoint_name,
data_capture_config=DataCaptureConfig(
enable_capture=True,
sampling_percentage=100,
destination_s3_uri=s3_capture_upload_path))
# -
# <span style="color: red; font-weight:bold">Please take note of the endpoint name, since it will be used in the next workshop module.</span>
# ## Getting inferences
# Finally we can try invoking our pipeline of models and get some inferences:
# +
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import CSVDeserializer
from sagemaker.predictor import Predictor
predictor = Predictor(
endpoint_name=endpoint_name,
sagemaker_session=sagemaker_session,
serializer=CSVSerializer(),
deserializer=CSVDeserializer())
payload = "TID008,HAWT,64,80,46,21,55,55,7,34,SE"
print(predictor.predict(payload))
# -
#
#
# Now let's list the data capture files stored in S3. You should expect to see different files from different time periods organized based on the hour in which the invocation occurred.
#
# Note that the delivery of capture data to Amazon S3 can require a couple of minutes so next cell might error. If this happens, please retry after a minute.
#
# +
s3_client = boto3.Session().client('s3')
current_endpoint_capture_prefix = '{}/monitoring/datacapture/{}'.format(prefix, endpoint_name)
result = s3_client.list_objects(Bucket=bucket_name, Prefix=current_endpoint_capture_prefix)
capture_files = ['s3://{0}/{1}'.format(bucket_name, capture_file.get("Key")) for capture_file in result.get('Contents')]
print("Capture Files: ")
print("\n ".join(capture_files))
# -
# We can also read the contents of one of these files and see how capture records are organized in JSON lines format.
# !aws s3 cp {capture_files[0]} datacapture/captured_data_example.jsonl
# !head datacapture/captured_data_example.jsonl
# In addition, we can better understand the content of each JSON line like follows:
# +
import json
with open ("datacapture/captured_data_example.jsonl", "r") as myfile:
data=myfile.read()
print(json.dumps(json.loads(data.split('\n')[0]), indent=2))
# -
#
#
# For each inference request, we get input data, output data and some metadata like the inference time captured and saved.
# ## Baselining
#
# From our validation dataset let's ask Amazon SageMaker to suggest a set of baseline constraints and generate descriptive statistics for our features.
#
# +
baseline_data_path = 's3://{0}/{1}/data/raw'.format(bucket_name, prefix)
baseline_results_path = 's3://{0}/{1}/monitoring/baselining/results'.format(bucket_name, prefix)
print(baseline_data_path)
print(baseline_results_path)
# -
# Please note that running the baselining job will require 8-10 minutes. In the meantime, you can take a look at the Deequ library, used to execute these analyses with the default Model Monitor container: https://github.com/awslabs/deequ
# +
from sagemaker.model_monitor import DefaultModelMonitor
from sagemaker.model_monitor.dataset_format import DatasetFormat
my_default_monitor = DefaultModelMonitor(
role=role,
instance_count=1,
instance_type='ml.c5.4xlarge',
volume_size_in_gb=20,
max_runtime_in_seconds=3600,
)
# -
my_default_monitor.suggest_baseline(
baseline_dataset=baseline_data_path,
dataset_format=DatasetFormat.csv(header=True),
output_s3_uri=baseline_results_path,
wait=True
)
#
#
# Let's display the statistics that were generated by the baselining job.
#
# +
import pandas as pd
baseline_job = my_default_monitor.latest_baselining_job
schema_df = pd.json_normalize(baseline_job.baseline_statistics().body_dict["features"])
schema_df
# -
# Then, we can also visualize the constraints.
constraints_df = pd.json_normalize(baseline_job.suggested_constraints().body_dict["features"])
constraints_df
# ### Switching order of target variable
#
# Amazon SageMaker Model Monitor expects that the target variable is the first feature of your dataset when comparing CSV captured data with the baseline.
# However, since the dataset we used for baselining had the breakdown variable as last feature, we are going to switch its order in the generated statistics and constraints file.
# +
statistics_path = baseline_results_path + '/statistics.json'
constraints_path = baseline_results_path + '/constraints.json'
# !aws s3 cp {statistics_path} baseline/
# !aws s3 cp {constraints_path} baseline/
# +
import json
with open('baseline/statistics.json', 'r') as statistics_file:
loaded_statistics = json.load(statistics_file)
loaded_statistics['features'].insert(0, loaded_statistics['features'][-1])
del loaded_statistics['features'][-1]
with open('baseline/statistics.json', 'w') as statistics_file:
json.dump(loaded_statistics, statistics_file)
# -
# !aws s3 cp baseline/statistics.json {statistics_path}
# +
with open('baseline/constraints.json', 'r') as constraints_file:
loaded_constraints = json.load(constraints_file)
loaded_constraints['features'].insert(0, loaded_constraints['features'][-1])
del loaded_constraints['features'][-1]
with open('baseline/constraints.json', 'w') as constraints_file:
json.dump(loaded_constraints, constraints_file)
# -
# !aws s3 cp baseline/constraints.json {constraints_path}
# ## Results
#
# The baselining job has inspected the validation dataset and generated constraints and statistics, that will be used to monitor our endpoint.
# ## Generating violations artificially
#
# In order to get some result relevant to monitoring analysis, you can try and generate artificially some inferences with feature values causing specific violations, and then invoke the endpoint with this data.
# +
import time
import numpy as np
dist_values = np.random.normal(1, 0.2, 200)
# wind_speed -> set to float (expected integer)
# rpm_blade -> set to empty (missing value)
# humidity -> sampled from random normal distribution [seventh feature]
#TODO
artificial_values = "TID008,HAWT,65.8,,46,21,55,{0},7,34,SE"
for i in range(200):
predictor.predict(artificial_values.format(str(dist_values[i])))
time.sleep(0.15)
if i > 0 and i % 100 == 0 :
print('Executed {0} inferences.'.format(i))
# -
#
# # Monitoring
#
# Once we have built the baseline for our data, we can enable endpoint monitoring by creating a monitoring schedule. When the schedule fires, a monitoring job will be kicked-off and will inspect the data captured at the endpoint with respect to the baseline; then it will generate some report files that can be used to analyze monitoring results.
# ## Create Monitoring Schedule
#
# Let's create the monitoring schedule for the previously created endpoint. When we create the schedule, we can also specify two scripts that will preprocess the records before the analysis takes place and execute post-processing at the end. For this example, we are not going to use a record preprocessor, and we are just specifying a post-processor that outputs some text for demo purposes.
#
# !pygmentize postprocessor.py
# +
import boto3
monitoring_code_prefix = '{0}/monitoring/code'.format(prefix)
print(monitoring_code_prefix)
boto3.Session().resource('s3').Bucket(bucket_name).Object(monitoring_code_prefix + '/postprocessor.py').upload_file('postprocessor.py')
postprocessor_path = 's3://{0}/{1}/monitoring/code/postprocessor.py'.format(bucket_name, prefix)
print(postprocessor_path)
reports_path = 's3://{0}/{1}/monitoring/reports'.format(bucket_name, prefix)
print(reports_path)
# -
# Finally, we create the monitoring schedule with hourly schedule execution.
# +
from sagemaker.model_monitor import CronExpressionGenerator
from time import gmtime, strftime
endpoint_name = predictor.endpoint_name
mon_schedule_name = 'end-to-end-ml-sm-mon-sch-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
my_default_monitor.create_monitoring_schedule(
monitor_schedule_name=mon_schedule_name,
endpoint_input=endpoint_name,
post_analytics_processor_script=postprocessor_path,
output_s3_uri=reports_path,
statistics=my_default_monitor.baseline_statistics(),
constraints=my_default_monitor.suggested_constraints(),
schedule_cron_expression=CronExpressionGenerator.hourly(),
enable_cloudwatch_metrics=True
)
# -
# ## Describe Monitoring Schedule
desc_schedule_result = my_default_monitor.describe_schedule()
desc_schedule_result
#
# ## Delete Monitoring Schedule
#
# Once the schedule is created, it will kick of jobs at specified intervals. Note that if you are kicking this off after creating the hourly schedule, you might find the executions empty. You might have to wait till you cross the hour boundary (in UTC) to see executions kick off. Since we don't want to wait for the hour in this example we can delete the schedule and use the code in next steps to simulate what will happen when a schedule is triggered, by running an Amazon SageMaker Processing Job.
#
# Note: this is just for the purpose of running this example.
my_default_monitor.delete_monitoring_schedule()
# ## Triggering execution manually
#
# In oder to trigger the execution manually, we first get all paths to data capture, baseline statistics, baseline constraints, etc. Then, we use a utility fuction, defined in monitoringjob_utils.py, to run the processing job.
# +
result = s3_client.list_objects(Bucket=bucket_name, Prefix=current_endpoint_capture_prefix)
capture_files = ['s3://{0}/{1}'.format(bucket_name, capture_file.get("Key")) for capture_file in result.get('Contents')]
print("Capture Files: ")
print("\n ".join(capture_files))
data_capture_path = capture_files[len(capture_files) - 1][: capture_files[len(capture_files) - 1].rfind('/')]
statistics_path = baseline_results_path + '/statistics.json'
constraints_path = baseline_results_path + '/constraints.json'
print(data_capture_path)
print(postprocessor_path)
print(statistics_path)
print(constraints_path)
print(reports_path)
# +
from monitoringjob_utils import run_model_monitor_job_processor
run_model_monitor_job_processor(region, 'ml.m5.xlarge', role, data_capture_path, statistics_path, constraints_path, reports_path,
postprocessor_path=postprocessor_path)
# -
# ## Analysis
#
# When the monitoring job completes, monitoring reports are saved to Amazon S3. Let's list the generated reports.
# +
s3_client = boto3.Session().client('s3')
monitoring_reports_prefix = '{}/monitoring/reports/{}'.format(prefix, predictor.endpoint_name)
result = s3_client.list_objects(Bucket=bucket_name, Prefix=monitoring_reports_prefix)
try:
monitoring_reports = ['s3://{0}/{1}'.format(bucket_name, capture_file.get("Key")) for capture_file in result.get('Contents')]
print("Monitoring Reports Files: ")
print("\n ".join(monitoring_reports))
except:
print('No monitoring reports found.')
# -
# !aws s3 cp {monitoring_reports[0]} monitoring/
# !aws s3 cp {monitoring_reports[1]} monitoring/
# !aws s3 cp {monitoring_reports[2]} monitoring/
# Let's display the violations identified by the monitoring execution.
# +
import pandas as pd
pd.set_option('display.max_colwidth', None)
file = open('monitoring/constraint_violations.json', 'r')
data = file.read()
violations_df = pd.json_normalize(json.loads(data)['violations'])
violations_df
# -
#
# ## Advanced Hints
#
# You might be asking yourself what are the type of violations that are monitored and how drift from the baseline is computed.
#
# The types of violations monitored are listed here: https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor-interpreting-violations.html. Most of them use configurable thresholds, that are specified in the monitoring configuration section of the baseline constraints JSON. Let's take a look at this configuration from the baseline constraints file:
#
# !aws s3 cp {statistics_path} baseline/
# !aws s3 cp {constraints_path} baseline/
# +
import json
with open ("baseline/constraints.json", "r") as myfile:
data=myfile.read()
print(json.dumps(json.loads(data)['monitoring_config'], indent=2))
# -
#
#
# This configuration is intepreted when the monitoring job is executed and used to compare captured data to the baseline. If you want to customize this section, you will have to update the constraints.json file and upload it back to Amazon S3 before launching the monitoring job.
#
# When data distributions are compared to detect potential drift, you can choose to use either a Simple or Robust comparison method, where the latter has to be preferred when dealing with small datasets. Additional info: https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor-byoc-constraints.html.
#
# +
#predictor.delete_endpoint()
#predictor.delete_model()
# -
# Once we have tested the endpoint, we can move to the next workshop module. Please access the module <a href="https://github.com/giuseppeporcelli/end-to-end-ml-sm/tree/master/05_API_Gateway_and_Lambda" target="_blank">05_API_Gateway_and_Lambda</a> on GitHub to continue.
| 04_deploy_model/04_deploy_model_monitor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://maltem.com/wp-content/uploads/2020/04/LOGO_MALTEM.png" style="float: left; margin: 20px; height: 55px">
#
# <br>
# <br>
# <br>
# <br>
#
#
# # Bootstrapping and Bagging
#
# ### Learning Objectives
# - Define ensemble model.
# - Name three advantages of using ensemble models.
# - Define and execute bootstrapping.
# - Fit and evaluate bagged decision trees.
# +
# Import libraries.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# Import Bagging Classifier.
from sklearn.ensemble import BaggingClassifier
# -
# ## Load the Data
#
# We'll be using the `Heart.csv` from the [ISLR Website](https://www.statlearning.com/). There's a copy in this repo under `./datasets/Heart.csv`.
# +
# Read in the Heart .csv data.
df = pd.read_csv('datasets/Heart.csv')
# Check the first few rows.
# -
df.head()
# ## Data Cleaning & Model Prep
# +
# Drop the `Unnamed: 0` column.
df.drop('Unnamed: 0',axis = 1, inplace=True)
# Drop NAs.
df.dropna(inplace = True)
# Create dummies for the `ChestPain`, `Thal`, and `AHD` columns.
# Be sure to set `drop_first=True`.
df = pd.get_dummies(
df,
columns = ['ChestPain', 'Thal', 'AHD'],
drop_first = True
)
# Define X and y.
X = df.drop('AHD_Yes', axis = 'columns')
y = df['AHD_Yes']
# Split data into training and testing sets.
X_train,X_test,y_train,y_test = train_test_split(X,
y,
random_state = 42,
stratify = y)
# -
# Our target column will be `AHD_Yes`:
# - 1 means the patient has heart disease
# - 0 means they aren't diagnosed with heart disease
# What is the accuracy of our baseline model?
y.value_counts(normalize = True)
# <details><summary>What does a false positive mean in this case?</summary>
#
# - A false positive indicates someone **falsely** predict as being in the **positive** class.
# - This is someone we incorrectly think has heart disease.
# - Incorrectly predicting someone to have heart disease is bad... but it _might_ be worse to incorrectly predict that someone is healthy!
# </details>
# Instantiate `DecisionTreeClassifier` object.
tree = DecisionTreeClassifier (random_state = 42)
# ## Note: The role of randomness
#
# The algorithms that fit tree-based methods involve randomness, which means it's important to specify a `random_state` if you want to reproduce your results. This is always a good idea.
# - Changing `random_state` from 42 to 43 reduces our model's test performance by 6%!
# Fit and score on the training data.
tree.fit(X_train, y_train)
# Score on the testing data.
tree.score(X_train, y_train)
tree.score(X_test, y_test)
# <details><summary> Where do decision trees tend to fall on the Bias/Variance spectrum?</summary>
#
# - Decision trees very easily overfit.
# - They tend to suffer from **high error due to variance**.
# </details>
# ## Bootstrapping
#
# Bootstrapping is a powerful idea used frequently across statistics and data science.
# - One common use for bootstrapping is to use computers and random number generation to generate confidence intervals or execute hypothesis tests for us, instead of relying on the Central Limit Theorem and memorized formulas.
# - We'll use it later to improve the performance of our decision tree models!
#
#
# #### What is the motivation behind bootstrapping?
# In a perfect world, we would have access to the full population of data instead of a sample of data.
# <details><summary>Why is it unrealistic to assume we have access to the full population of data?</summary>
#
# - It would take too much time.
# - It would cost too much money.
# - Logistical challenges.
# </details>
# In a few words, bootstrapping is **random resampling with replacement**.
#
# The idea is this:
# - Take your original sample of data, with sample size $n$.
# - Take many sub-samples (say $B$) of size $n$ from your sample **with replacement**. These are called **bootstrapped samples**.
# - You have now generated $B$ bootstrapped samples, where each sample is of size $n$!
#
# <img src="./assets/bootstrap.png" alt="drawing" width="550"/>
#
# - Instead of building one model on our original sample, we will now build one model on each bootstrapped sample, giving us $B$ models in total!
# - Experience tells us that combining the models from our bootstrapped samples will be closer to what we'd see from the population than to just get one model from our original sample.
#
# This sets up the idea of an **ensemble model**.
# <details><summary>Why do you think we want to take a sample of size n?</summary>
#
# - Because we want our estimators to be fit on data of the same size!
# - If our original data had a sample size of 1,000, but we fit decision trees to samples of size 50, the decision trees fit to samples of size 50 will probably look very, very different from decision trees fit on a sample of size 1,000.
# </details>
# <details><summary>Why do you think we want to sample with replacement?</summary>
#
# - If we didn't sample with replacement, we'd just get identical samples of size $n$. (These would be copies of our original data!)
# </details>
# If you are generating one bootstrapped sample in `pandas`:
# +
# Generate one bootstrapped sample
# of size n from X_train.
X_train.sample(n= X_train.shape[0],
replace = True,
random_state = 42)
# -
# Let's visualize some bootstrapped samples.
# Create bootstrapped samples
# & plot cholesterol level for each bootstrapped sample.
for i in range(3):
sns.distplot(X_train.sample(n = X_train.shape[0], replace = True)['Chol']);
# ## Introduction to Ensemble Methods
# We can list out the different types of models we've built thus far:
# - Linear Regression
# - Logistic Regression
# - $k$-Nearest Neighbors
# - Naive Bayes Classification
#
# If we want to use any of these models, we follow the same type of process.
# 1. Based on our problem, we identify which model to use. (Is our problem classification or regression? Do we want an interpretable model?)
# 2. Fit the model using the training data.
# 3. Use the fit model to generate predictions.
# 4. Evaluate our model's performance and, if necessary, return to step 2 and make changes.
#
# So far, we've always had exactly one model. Today, however, we're going to talk about **ensemble methods**. Mentally, you should think about this as if we build multiple models and then aggregate their results in some way.
#
# ## Why would we build an "ensemble model?"
#
# Our goal is to estimate $f$, the true function. (Think about $f$ as the **true process** that dictates Ames housing prices.)
#
# We can come up with different models $m_1$, $m_2$, and so on to get as close to $f$ as possible. (Think about $m_1$ as the model you built to predict $f$, think of $m_2$ as the model your neighbor built to predict $f$, and so on.)
#
# ### (BONUS) Three Benefits: Statistical, Computational, Representational
# - The **statistical** benefit to ensemble methods: By building one model, our predictions are almost certainly going to be wrong. Predictions from one model might overestimate housing prices; predictions from another model might underestimate housing prices. By "averaging" predictions from multiple models, we'll see that we can often cancel our errors out and get closer to the true function $f$.
# - The **computational** benefit to ensemble methods: It might be impossible to develop one model that globally optimizes our objective function. (Remember that CART reach locally-optimal solutions that aren't guaranteed to be the globally-optimal solution.) In these cases, it may be **impossible** for one CART to arrive at the true function $f$. However, generating many different models and averaging their predictions may allow us to get results that are closer to the global optimum than any individual model.
# - The **representational** benefit to ensemble methods: Even if we had all the data and all the computer power in the world, it might be impossible for one model to **exactly** equal $f$. For example, a linear regression model can never model a relationship where a one-unit change in $X$ is associated with some *different* change in $Y$ based on the value of $X$. All models have some shortcomings. (See [the no free lunch theorems](https://en.wikipedia.org/wiki/No_free_lunch_in_search_and_optimization).) While individual models have shortcomings, by creating multiple models and aggregating their predictions, we can actually create predictions that represent something that one model cannot ever represent.
#
# We can summarize this as the **wisdom of the crowd**.
#
# ## Wisdom of the Crowd: Guess the weight of Penelope
#
# 
#
# [*Image source*](https://www.npr.org/sections/money/2015/07/17/422881071/how-much-does-this-cow-weigh)
# ## Ensemble models
#
# We can use the "wisdom of the crowd" idea by creating several models and then aggregating their results in some way.
#
# Types of ensemble models:
# - Bagging
# - Boosting
# - [Stacking](https://www.geeksforgeeks.org/stacking-in-machine-learning/)
# ## Bagging: Bootstrap Aggregating
#
# Decision trees are powerful machine learning models. However, decision trees have some limitations. In particular, trees that are grown very deep tend to learn highly irregular patterns (a.k.a. they overfit their training sets).
#
# Bagging (bootstrap aggregating) mitigates this problem by exposing different trees to different sub-samples of the training set.
#
# The process for creating bagged decision trees is as follows:
# 1. From the original data of size $n$, bootstrap $B$ samples each of size $n$ (with replacement!).
# 2. Build a decision tree on each bootstrapped sample.
# 3. Make predictions by passing a test observation through all $B$ trees and developing one aggregate prediction for that observation.
#
# 
#
# ### What do you mean by "aggregate prediction?"
# As with all of our modeling techniques, we want to make sure that we can come up with one final prediction for each observation.
#
# Suppose we want to predict whether or not a Reddit post is going to go viral, where `1` indicates viral and `0` indicates non-viral. We build 100 decision trees. Given a new Reddit post labeled `X_test`, we pass these features into all 100 decision trees.
# - 70 of the trees predict that the post in `X_test` will go viral.
# - 30 of the trees predict that the post in `X_test` will not go viral.
# <details><summary>What might you expect .predict(X_test) to output?</summary>
#
# - `.predict(X_test)` should output a 1, predicting that the post will go viral.
#
# </details>
# <details><summary>What might you expect .predict_proba(X_test) to output?</summary>
#
# - `.predict_proba(X_test)` should output [0.3 0.7], indicating the probability of the post going viral is 70% and the probability of the post not going viral to be 30%.
# </details>
#
# ## Bagging Classifier using a `for` loop
#
# In the cell below, we'll create an ensemble of trees - we'll train each tree on a separate **bootstrapped** sample of the training data.
pd.DataFrame(index = X_test.index)
# +
# Instantiate dataframe.
predections = pd.DataFrame(index = X_test.index)
# Generate ten decision trees.
for i in range (1,11):
# Bootstrap X data.
# Should we add a random seed?
X_sample = X_train.sample( n = X_train.shape[0],
replace = True)
# Get y data that matches the X data.
y_sample = y_train[X_sample.index]
# Instantiate decision tree.
t = DecisionTreeClassifier()
# Fit to our sample data.
t.fit(X_sample, y_sample)
# Put predictions in dataframe.
predections[f'Tree {i}'] = t.predict(X_test)
# -
predections.head()
# Generate aggregated predicted probabilities.
probs = predections.mean(axis = 'columns')
# What's our accuracy?
accuracy_score(y_test,(probs >.5).astype(int))
# ## Bagging Classifier using `sklearn`
#
# [BaggingClassifier Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.BaggingClassifier.html)
#
# In the cell below, create and score instance of `BaggingClassifier` on the test set.
# +
# Instantiate BaggingClassifier.
bag = BaggingClassifier(random_state = 42)
# Fit BaggingClassifier.
bag.fit(X_train,y_train)
# Score BaggingClassifier.
# -
bag.score(X_test,y_test)
# ## Interview Question
# <details><summary>What is bootstrapping?</summary>
#
# - Bootstrapping is random resampling with replacement.
# - We bootstrap when fitting bagged decision trees so that we can fit multiple decision trees on slightly different sets of data. Bagged decision trees tend to outperform single decision trees.
# - Bootstrapping can also be used to conduct hypothesis tests and generate confidence intervals directly from resampled data.
# </details>
| Notebook/Lesson-bagging/starter-code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Content-Based Filtering Using Neural Networks
# This notebook relies on files created in the [content_based_preproc.ipynb](./content_based_preproc.ipynb) notebook. Be sure to run the code in there before completing this notebook.
# Also, we'll be using the **python3** kernel from here on out so don't forget to change the kernel if it's still Python2.
# This lab illustrates:
# 1. how to build feature columns for a model using tf.feature_column
# 2. how to create custom evaluation metrics and add them to Tensorboard
# 3. how to train a model and make predictions with the saved model
# Tensorflow Hub should already be installed. You can check that it is by using "pip freeze".
# + language="bash"
# pip freeze | grep tensor
# -
# Let's make sure we install the necessary version of tensorflow-hub. After doing the pip install below, click **"Restart the kernel"** on the notebook so that the Python environment picks up the new packages.
# !pip3 install tensorflow-hub==0.7.0
# !pip3 install --upgrade tensorflow==1.15.3
# !pip3 install google-cloud-bigquery==1.10
# #### **Note**: Please ignore any incompatibility warnings and errors and re-run the cell to view the installed tensorflow version.
# +
import os
import tensorflow as tf
import numpy as np
import tensorflow_hub as hub
import shutil
PROJECT = 'cloud-training-demos' # REPLACE WITH YOUR PROJECT ID
BUCKET = 'cloud-training-demos-ml' # REPLACE WITH YOUR BUCKET NAME
REGION = 'us-central1' # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# do not change these
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.15.3'
# + language="bash"
# gcloud config set project $PROJECT
# gcloud config set compute/region $REGION
# -
# ### Build the feature columns for the model.
# To start, we'll load the list of categories, authors and article ids we created in the previous **Create Datasets** notebook.
categories_list = open("categories.txt").read().splitlines()
authors_list = open("authors.txt").read().splitlines()
content_ids_list = open("content_ids.txt").read().splitlines()
mean_months_since_epoch = 523
# In the cell below we'll define the feature columns to use in our model. If necessary, remind yourself the [various feature columns](https://www.tensorflow.org/api_docs/python/tf/feature_column) to use.
# For the embedded_title_column feature column, use a Tensorflow Hub Module to create an embedding of the article title. Since the articles and titles are in German, you'll want to use a German language embedding module.
# Explore the text embedding Tensorflow Hub modules [available here](https://alpha.tfhub.dev/). Filter by setting the language to 'German'. The 50 dimensional embedding should be sufficient for our purposes.
# +
embedded_title_column = hub.text_embedding_column(
key="title",
module_spec="https://tfhub.dev/google/nnlm-de-dim50/1",
trainable=False)
content_id_column = tf.feature_column.categorical_column_with_hash_bucket(
key="content_id",
hash_bucket_size= len(content_ids_list) + 1)
embedded_content_column = tf.feature_column.embedding_column(
categorical_column=content_id_column,
dimension=10)
author_column = tf.feature_column.categorical_column_with_hash_bucket(key="author",
hash_bucket_size=len(authors_list) + 1)
embedded_author_column = tf.feature_column.embedding_column(
categorical_column=author_column,
dimension=3)
category_column_categorical = tf.feature_column.categorical_column_with_vocabulary_list(
key="category",
vocabulary_list=categories_list,
num_oov_buckets=1)
category_column = tf.feature_column.indicator_column(category_column_categorical)
months_since_epoch_boundaries = list(range(400,700,20))
months_since_epoch_column = tf.feature_column.numeric_column(
key="months_since_epoch")
months_since_epoch_bucketized = tf.feature_column.bucketized_column(
source_column = months_since_epoch_column,
boundaries = months_since_epoch_boundaries)
crossed_months_since_category_column = tf.feature_column.indicator_column(tf.feature_column.crossed_column(
keys = [category_column_categorical, months_since_epoch_bucketized],
hash_bucket_size = len(months_since_epoch_boundaries) * (len(categories_list) + 1)))
feature_columns = [embedded_content_column,
embedded_author_column,
category_column,
embedded_title_column,
crossed_months_since_category_column]
# -
# ### Create the input function.
#
# Next we'll create the input function for our model. This input function reads the data from the csv files we created in the previous labs.
record_defaults = [["Unknown"], ["Unknown"],["Unknown"],["Unknown"],["Unknown"],[mean_months_since_epoch],["Unknown"]]
column_keys = ["visitor_id", "content_id", "category", "title", "author", "months_since_epoch", "next_content_id"]
label_key = "next_content_id"
def read_dataset(filename, mode, batch_size = 512):
def _input_fn():
def decode_csv(value_column):
columns = tf.decode_csv(value_column,record_defaults=record_defaults)
features = dict(zip(column_keys, columns))
label = features.pop(label_key)
return features, label
# Create list of files that match pattern
file_list = tf.io.gfile.glob(filename)
# Create dataset from file list
dataset = tf.data.TextLineDataset(file_list).map(decode_csv)
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # indefinitely
dataset = dataset.shuffle(buffer_size = 10 * batch_size)
else:
num_epochs = 1 # end-of-input after this
dataset = dataset.repeat(num_epochs).batch(batch_size)
return dataset.make_one_shot_iterator().get_next()
return _input_fn
# ### Create the model and train/evaluate
#
#
# Next, we'll build our model which recommends an article for a visitor to the Kurier.at website. Look through the code below. We use the input_layer feature column to create the dense input layer to our network. This is just a single layer network where we can adjust the number of hidden units as a parameter.
#
# Currently, we compute the accuracy between our predicted 'next article' and the actual 'next article' read next by the visitor. We'll also add an additional performance metric of top 10 accuracy to assess our model. To accomplish this, we compute the top 10 accuracy metric, add it to the metrics dictionary below and add it to the tf.summary so that this value is reported to Tensorboard as well.
def model_fn(features, labels, mode, params):
net = tf.feature_column.input_layer(features, params['feature_columns'])
for units in params['hidden_units']:
net = tf.layers.dense(net, units=units, activation=tf.nn.relu)
# Compute logits (1 per class).
logits = tf.layers.dense(net, params['n_classes'], activation=None)
predicted_classes = tf.argmax(logits, 1)
from tensorflow.python.lib.io import file_io
with file_io.FileIO('content_ids.txt', mode='r') as ifp:
content = tf.constant([x.rstrip() for x in ifp])
predicted_class_names = tf.gather(content, predicted_classes)
if mode == tf.estimator.ModeKeys.PREDICT:
predictions = {
'class_ids': predicted_classes[:, tf.newaxis],
'class_names' : predicted_class_names[:, tf.newaxis],
'probabilities': tf.nn.softmax(logits),
'logits': logits,
}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
table = tf.contrib.lookup.index_table_from_file(vocabulary_file="content_ids.txt")
labels = table.lookup(labels)
# Compute loss.
loss = tf.losses.sparse_softmax_cross_entropy(labels=labels, logits=logits)
# Compute evaluation metrics.
accuracy = tf.metrics.accuracy(labels=labels,
predictions=predicted_classes,
name='acc_op')
top_10_accuracy = tf.metrics.mean(tf.nn.in_top_k(predictions=logits,
targets=labels,
k=10))
metrics = {
'accuracy': accuracy,
'top_10_accuracy' : top_10_accuracy}
tf.summary.scalar('accuracy', accuracy[1])
tf.summary.scalar('top_10_accuracy', top_10_accuracy[1])
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(
mode, loss=loss, eval_metric_ops=metrics)
# Create training op.
assert mode == tf.estimator.ModeKeys.TRAIN
optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
train_op = optimizer.minimize(loss, global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode, loss=loss, train_op=train_op)
# ### Train and Evaluate
# +
outdir = 'content_based_model_trained'
shutil.rmtree(outdir, ignore_errors = True) # start fresh each time
#tf.summary.FileWriterCache.clear() # ensure filewriter cache is clear for TensorBoard events file
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir = outdir,
params={
'feature_columns': feature_columns,
'hidden_units': [200, 100, 50],
'n_classes': len(content_ids_list)
})
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset("training_set.csv", tf.estimator.ModeKeys.TRAIN),
max_steps = 2000)
eval_spec = tf.estimator.EvalSpec(
input_fn = read_dataset("test_set.csv", tf.estimator.ModeKeys.EVAL),
steps = None,
start_delay_secs = 30,
throttle_secs = 60)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# -
# This takes a while to complete but in the end, I get about **30% top 10 accuracy**.
# ### Make predictions with the trained model.
#
# With the model now trained, we can make predictions by calling the predict method on the estimator. Let's look at how our model predicts on the first five examples of the training set.
# To start, we'll create a new file 'first_5.csv' which contains the first five elements of our training set. We'll also save the target values to a file 'first_5_content_ids' so we can compare our results.
# + language="bash"
# head -5 training_set.csv > first_5.csv
# head first_5.csv
# awk -F "\"*,\"*" '{print $2}' first_5.csv > first_5_content_ids
# -
# Recall, to make predictions on the trained model we pass a list of examples through the input function. Complete the code below to make predictions on the examples contained in the "first_5.csv" file we created above.
output = list(estimator.predict(input_fn=read_dataset("first_5.csv", tf.estimator.ModeKeys.PREDICT)))
import numpy as np
recommended_content_ids = [np.asscalar(d["class_names"]).decode('UTF-8') for d in output]
content_ids = open("first_5_content_ids").read().splitlines()
# Finally, we map the content id back to the article title. Let's compare our model's recommendation for the first example. This can be done in BigQuery. Look through the query below and make sure it is clear what is being returned.
# +
from google.cloud import bigquery
recommended_title_sql="""
#standardSQL
SELECT
(SELECT MAX(IF(index=6, value, NULL)) FROM UNNEST(hits.customDimensions)) AS title
FROM `cloud-training-demos.GA360_test.ga_sessions_sample`,
UNNEST(hits) AS hits
WHERE
# only include hits on pages
hits.type = "PAGE"
AND (SELECT MAX(IF(index=10, value, NULL)) FROM UNNEST(hits.customDimensions)) = \"{}\"
LIMIT 1""".format(recommended_content_ids[0])
current_title_sql="""
#standardSQL
SELECT
(SELECT MAX(IF(index=6, value, NULL)) FROM UNNEST(hits.customDimensions)) AS title
FROM `cloud-training-demos.GA360_test.ga_sessions_sample`,
UNNEST(hits) AS hits
WHERE
# only include hits on pages
hits.type = "PAGE"
AND (SELECT MAX(IF(index=10, value, NULL)) FROM UNNEST(hits.customDimensions)) = \"{}\"
LIMIT 1""".format(content_ids[0])
recommended_title = bigquery.Client().query(recommended_title_sql).to_dataframe()['title'].tolist()[0].encode('utf-8').strip()
current_title = bigquery.Client().query(current_title_sql).to_dataframe()['title'].tolist()[0].encode('utf-8').strip()
print("Current title: {} ".format(current_title))
print("Recommended title: {}".format(recommended_title))
# -
| courses/machine_learning/deepdive/10_recommend/content_based_using_neural_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="589NVNT00LOv"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score, accuracy_score, mean_absolute_error
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 814} id="vJgJ0i-Z0gZY" outputId="bd40ad28-68bb-4e63-8869-38e84f6d99d7"
df = pd.read_csv('/content/listings.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="lg9punW0JtGY" outputId="bddd0c30-ba3a-428a-fb38-06fc983754af"
df.columns
# + id="o5FqXPjAJx3p"
df_n = df.drop(columns=['listing_url', 'scrape_id', 'last_scraped', 'name', 'summary','space', 'experiences_offered',
'neighborhood_overview','notes', 'transit', 'thumbnail_url', 'medium_url', 'picture_url','xl_picture_url',
'host_id', 'host_url', 'host_name', 'host_since','host_location', 'host_about', 'host_response_time',
'host_response_rate', 'host_acceptance_rate', 'host_is_superhost','host_thumbnail_url', 'host_picture_url',
'host_neighbourhood','host_listings_count', 'host_total_listings_count','host_verifications',
'host_has_profile_pic', 'host_identity_verified','street', 'neighbourhood_cleansed','neighbourhood_group_cleansed',
'state', 'zipcode', 'market','smart_location', 'country_code', 'country', 'latitude', 'longitude',
'is_location_exact', 'property_type', 'room_type', 'bed_type', 'square_feet', 'weekly_price', 'monthly_price',
'security_deposit','cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights','maximum_nights',
'calendar_updated', 'has_availability','availability_30', 'availability_60', 'availability_90','availability_365',
'calendar_last_scraped', 'number_of_reviews','first_review', 'last_review','review_scores_accuracy',
'review_scores_cleanliness','review_scores_checkin', 'review_scores_communication','review_scores_location',
'review_scores_value', 'requires_license','license', 'jurisdiction_names', 'instant_bookable','cancellation_policy',
'require_guest_profile_picture','require_guest_phone_verification', 'calculated_host_listings_count',
'reviews_per_month', 'amenities', 'accommodates', 'neighbourhood', 'city', 'review_scores_rating'], inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="rc8pSocGJ0Nj" outputId="421a51bc-b1d3-4419-a8c2-e120d429d2dc"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="zdXNuCi6J35d" outputId="6229fa67-aebe-4260-9cf8-51fcf6a23c35"
#Correct datatype for price from str to int
df['price'] = pd.to_numeric(df['price'].map(lambda x: x.strip('$').replace(',','')))
print("price data type: ", type(df['price'].iloc[0]))
# + colab={"base_uri": "https://localhost:8080/"} id="X0cFuL9eJ7nH" outputId="1bc74631-221d-46ad-84ec-bece98c328ca"
# imports
import nltk
nltk.download(['punkt', 'wordnet', 'stopwords'])
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
import re
# define function tokenize
def tokenize(text):
'''
for text strings to be tokenized (tokens)
removing stop words
Lemmatization
'''
tokens = word_tokenize(text)
tokens = re.sub('[^a-zA-Z 0-9]', '', text)
tokens = tokens.lower().split()
stop_words = set(stopwords.words('english'))
lemmatizer = WordNetLemmatizer()
tokens = [w for w in tokens if not w in stop_words]
tokens = [lemmatizer.lemmatize(w.lower().strip()) for w in tokens]
return tokens
# + colab={"base_uri": "https://localhost:8080/"} id="Kxi2xAR7KAEE" outputId="209e27a1-8453-424b-bc8f-67765fdf7c34"
df['description'].value_counts(normalize=True)[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="I4lOJtgdKDZS" outputId="13960cbe-f369-4c04-8a7a-440b5b992<PASSWORD>"
# Using the tokenize function
df['description'] = df['description'].apply(str)
df['base_tokens'] = df['description'].apply(tokenize)
df['base_tokens'].head()
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="kafpJ9P-KF-J" outputId="89bd2032-b729-49ec-e9d5-f688409cd80d"
df[['description', 'base_tokens']][:10]
# + id="mS12AZA9KNY-"
# Create a function which takes a corpus of document and returns a dataframe of word counts
from collections import Counter
def count(docs):
'''This function takes a list of tokenized documents as input and returns
a dataframe with frequency counts and rank for each token.
# Arguments
docs: list, tokenized list of documents
# Returns
wc: dataframe,
'''
word_counts = Counter()
appears_in = Counter()
total_docs = len(docs)
for doc in docs:
word_counts.update(doc)
appears_in.update(set(doc))
temp = zip(word_counts.keys(), word_counts.values())
wc = pd.DataFrame(temp, columns = ['word', 'count'])
wc['rank'] = wc['count'].rank(method='first', ascending=False)
total = wc['count'].sum()
wc['pct_total'] = wc['count'].apply(lambda x: x / total)
wc = wc.sort_values(by='rank')
wc['cul_pct_total'] = wc['pct_total'].cumsum()
t2 = zip(appears_in.keys(), appears_in.values())
ac = pd.DataFrame(t2, columns=['word', 'appears_in'])
wc = ac.merge(wc, on='word')
wc['appears_in_pct'] = wc['appears_in'].apply(lambda x: x / total_docs)
return wc.sort_values(by='rank')
# + colab={"base_uri": "https://localhost:8080/", "height": 221} id="ZjuenUlzKQJb" outputId="55246558-b6b2-48aa-9f51-e442dcb052ad"
wc = count(df['base_tokens'])
print(wc.shape)
wc.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 751} id="scyvlelZKTzP" outputId="77793425-b040-4a4b-ea38-a027ccf4d4a0"
from wordcloud import WordCloud
comment_words = ''
stop_words = set(stopwords.words('english'))
for val in df.base_tokens:
val = str(val)
tokens = word_tokenize(val)
tokens = re.sub('[^a-zA-Z 0-9]', '', val)
tokens = tokens.lower().split()
lemmatizer = WordNetLemmatizer()
tokens = [w for w in tokens if not w in stop_words]
tokens = [lemmatizer.lemmatize(w.lower().strip()) for w in tokens]
comment_words += " ".join(tokens)+" "
wordcloud = WordCloud(width = 800, height = 800,
background_color='white',
stopwords = stop_words,
min_font_size = 10).generate(comment_words)
plt.figure(figsize = (10, 10), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="k8uMBvRCKXna" outputId="b69036bc-c292-4c56-d482-31fd6a12493a"
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import LinearSVC
from sklearn.pipeline import Pipeline
# initialize pipeline
# Create pipeline components
vect = TfidfVectorizer(stop_words='english', ngram_range=(1,2))
svm = LinearSVC()
# Get sparse dtm
dtm = vect.fit_transform(df.description)
# Convert to dataframe
dtm = pd.DataFrame(dtm.todense(), columns=vect.get_feature_names())
dtm.shape
pipe = Pipeline([
('vect', vect),
('clf', svm)
])
# split data and remove NA values
df.dropna(inplace=True)
X = df['description']
y = df['price']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
# pipeline fit
pipe.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="kGyPBqhoKbNk" outputId="5371f0d3-90cc-4f90-8bae-8297de80d4a8"
y_preds = pipe.predict(X_test)
y_preds
# + colab={"base_uri": "https://localhost:8080/"} id="zGQ6-BhvKe0o" outputId="b66841ff-2f10-4883-87a9-f60684e502c1"
mse = mean_squared_error(y_test, y_preds)
mse
# + colab={"base_uri": "https://localhost:8080/"} id="nQxS6QxLKiBP" outputId="acaef89c-23c1-47da-ed11-b253d2604456"
from math import sqrt
#root-mean-square deviation
rmse = sqrt(mse)
rmse
# + colab={"base_uri": "https://localhost:8080/"} id="ylrDq5_9KkHB" outputId="5e7bb9ca-6d24-43c5-a793-30d559291e51"
y_test.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="Ob-ThpNRKsww" outputId="59a02a53-c2a8-4339-9754-ae5bad22a627"
r2 = r2_score(y_test, y_preds)
r2
# + colab={"base_uri": "https://localhost:8080/"} id="rm5eMw_LKuxR" outputId="ede2e1d6-442e-40d8-9477-0141b33f2f21"
mae = mean_absolute_error(y_test, y_preds)
mae
# + colab={"base_uri": "https://localhost:8080/"} id="OIUFCg8ZKxUA" outputId="8d3db4ae-39f6-4d14-f3a8-fb6ac09c075d"
# GridSearchCV
from sklearn.model_selection import GridSearchCV
target = df['price']
rfc = RandomForestClassifier()
pipe2 = Pipeline([('vect', vect), ('clf', rfc)])
parameters = {
'vect__max_df': (0.7, 1.0),
'vect__min_df': (.02, .05, .10),
'vect__max_features':(500, 1500),
'clf__n_estimators':(5, 10),
'clf__max_depth':(5, 10, 15, 20)
}
grid_search = GridSearchCV(pipe2, parameters, cv=5, n_jobs=4, verbose=1)
grid_search.fit(df['description'], target)
# + colab={"base_uri": "https://localhost:8080/"} id="GelwoL-wKz9z" outputId="b5edf5a3-f278-4ad3-bb95-086f8c182827"
grid_search.best_score_
# + colab={"base_uri": "https://localhost:8080/"} id="jn393VXPK3ab" outputId="f9f08676-d619-49c6-e6d9-050dcf76ca1e"
grid_search.best_params_
# + colab={"base_uri": "https://localhost:8080/"} id="FvzZwBvtK6Bx" outputId="53c8ac40-8a53-4ee9-fa11-1bf21ff7da02"
from sklearn.metrics import accuracy_score
# Evaluate on test data
y_test = grid_search.predict(df.description)
accuracy_score(target, y_test)
# + id="KZl71NuvLC_V"
best_model = grid_search.best_estimator_
vect = best_model.named_steps['vect']
clf = best_model.named_steps['clf']
# + colab={"base_uri": "https://localhost:8080/"} id="xVO5wpowLIZ1" outputId="828ab4d2-cdb2-46d9-91e8-8b91093fb10f"
# !pip install eli5
# + colab={"base_uri": "https://localhost:8080/", "height": 445} id="iEikxcl1LLNH" outputId="6b912c2a-2cbe-494a-dcb7-82cf0185c171"
import eli5
eli5.show_weights(clf, vec=vect, top=20)
# + id="XUrNIwqSLNqb"
# Create dataframe containing price prediction per description
pred = grid_search.predict(df['description'])
# + colab={"base_uri": "https://localhost:8080/", "height": 979} id="Qy4SArEsLP6z" outputId="48d0e12c-7223-42b5-a6b0-c77ce71a2459"
#prediction price using description
df_pred = pd.DataFrame({'id': df['id'], 'description': df['description'], 'bathrooms': df['bathrooms'], 'bedrooms': df['bedrooms'],
'beds': df['beds'], 'price': df['price'], 'pred_price_using_desc': pred})
df_pred.head(30)
# + colab={"base_uri": "https://localhost:8080/"} id="xodFBhbkPRM1" outputId="6a179e84-cd61-4b2d-d614-b6a3304687e8"
import nltk
nltk.download('vader_lexicon')
# + colab={"base_uri": "https://localhost:8080/", "height": 564} id="Sz8oYMw4PRI9" outputId="4db79762-4e91-4653-d016-ce43a188fadc"
from nltk.sentiment.vader import SentimentIntensityAnalyzer
sid = SentimentIntensityAnalyzer()
df_pred['polarity_value']="Default"
df_pred['neg']=0.0
df_pred['pos']=0.0
df_pred['neu']=0.0
df_pred['compound']=0.0
for index,row in df_pred.iterrows():
ss = sid.polarity_scores(row['description'])
df_pred.at[index,'polarity_value'] = ss
df_pred.at[index,'neg'] = ss['neg']
df_pred.at[index,'pos'] = ss['pos']
df_pred.at[index,'neu']= ss['neu']
df_pred.at[index,'compound'] = ss['compound']
df_pred.head()
# + id="0e97H9ZlPREs"
# split data
X = df['description']
y = df['price']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
# + colab={"base_uri": "https://localhost:8080/"} id="4L57yCgbPRAr" outputId="c9930859-25e9-492d-faba-a97870b3dbf4"
# !pip install category_encoders
# + colab={"base_uri": "https://localhost:8080/"} id="EFSExyPEPgFJ" outputId="d19eaead-5a8b-417d-d85a-502cfcbb8d0d"
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(strategy='mean'),
StandardScaler(),
LogisticRegression(multi_class='auto', solver='lbfgs', n_jobs=-1)
)
# Fit on train
pipeline.fit(X_train, y_train)
# Score on validation
print ('Accruacy', pipeline.score(X_test, y_test))
# Predict on test
y_pred = pipeline.predict(X_test)
# + id="ZkSoeLzLQKB3"
# Save Model File
import pickle
filename = 'finalized_model.sav'
pickle.dump(pipeline, open(filename, 'wb'))
| notebooks/Architect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BrunaKuntz/Python-Curso-em-Video/blob/main/Mundo03/Desafio103.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xolHFjDi_Hj4"
#
# # **Desafio 103**
# **Python 3 - 3º Mundo**
#
# Descrição: Faça um programa que tenha uma função chamada ficha(), que receba dois parâmetros opcionais: o nome de um jogador e quantos gols ele marcou. O programa deverá ser capaz de mostrar a ficha do jogador, mesmo que algum dado não tenha sido informado corretamente.
#
# Link: https://www.youtube.com/watch?v=FbOvilKfHMI
# + id="8Lylf4Su-6aB"
def ficha(nome=None, gols='0'):
print('-'*20)
if gols not in '123456789':
gols = '0'
if nome in ' ':
nome = '<desconhecido>'
print(f'O jogador {nome} fez {gols} gol(s) no campeonato.')
# programa principal
nome = input('Nome do Jogador: ')
gols = input('Número de Gols: ')
ficha(nome, gols)
| Mundo03/Desafio103.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# # YahooFinance - Send daily prediction to Email
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/YahooFinance/YahooFinance_Send_daily_prediction_to_Email.ipynb" target="_parent"><img src="https://img.shields.io/badge/-Open%20in%20Naas-success?labelColor=000000&logo=data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iMTAyNHB4IiBoZWlnaHQ9IjEwMjRweCIgdmlld0JveD0iMCAwIDEwMjQgMTAyNCIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayIgdmVyc2lvbj0iMS4xIj4KIDwhLS0gR2VuZXJhdGVkIGJ5IFBpeGVsbWF0b3IgUHJvIDIuMC41IC0tPgogPGRlZnM+CiAgPHRleHQgaWQ9InN0cmluZyIgdHJhbnNmb3JtPSJtYXRyaXgoMS4wIDAuMCAwLjAgMS4wIDIyOC4wIDU0LjUpIiBmb250LWZhbWlseT<KEY>></a>
# + [markdown] papermill={} tags=[]
# **Tags:**
# + [markdown] papermill={} tags=[]
# With this template, you can create daily email prediction bot on any ticker available in [Yahoo finance](https://finance.yahoo.com/quote/TSLA/).<br>
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Import libraries
# + papermill={} tags=[]
import naas
import naas_drivers
import markdown2
import plotly.graph_objects as go
# + [markdown] papermill={} tags=[]
# ### Input ticker and dates
# 👉 Here you can change the ticker and timeframe
# + papermill={} tags=[]
ticker = "TSLA"
date_from = -100 # 1OO days max to feed the naas_driver for prediction
date_to = "today"
# + [markdown] papermill={} tags=[]
# ### Input email parameters
# 👉 Here you can input your sender email and destination email
#
# Note: emails are sent from <EMAIL> by default
# + papermill={} tags=[]
email_to = ["<EMAIL>","<EMAIL>"]
email_from = "<EMAIL>"
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Get dataset from Yahoo Finance
# + papermill={} tags=[]
df = naas_drivers.yahoofinance.get(ticker, date_from=date_from, date_to=date_to)
# clean df
df = df.dropna()
df.reset_index(drop=True)
df.head()
# + [markdown] papermill={} tags=[]
# ### Add prediction columns
# + papermill={} tags=[]
df = naas_drivers.prediction.get(dataset=df, prediction_type="all")
# + papermill={} tags=[]
df = df.sort_values("Date", ascending=False).reset_index(drop=True)
df.head()
# + [markdown] papermill={} tags=[]
# ### Save file
# + papermill={} tags=[]
save_file = df.to_excel(f"{ticker}_TODAY.xlsx")
# + [markdown] papermill={} tags=[]
# ### Build chart
# + papermill={} tags=[]
chart = naas_drivers.plotly.linechart(df, label_x="Date", label_y=["Close","ARIMA","SVR","LINEAR","COMPOUND"])
chart.update_layout(
title=f"{ticker} predictions as of today, for next 20 days.",
title_x=0.5,
showlegend=True,
legend=dict(
y=0.5,
)
)
chart
# + [markdown] papermill={} tags=[]
# ### Save as png and html
# + papermill={} tags=[]
naas_drivers.plotly.export(chart, f"{ticker}.png", css=None)
naas_drivers.plotly.export(chart, f"{ticker}.html", css=None)
# + [markdown] papermill={} tags=[]
# ### Expose chart
# + papermill={} tags=[]
link_image = naas.asset.add(f"{ticker}.png")
link_html = naas.asset.add(f"{ticker}.html", {"inline":True})
# + [markdown] papermill={} tags=[]
# ### Set daily variations values
# + papermill={} tags=[]
DATA = naas_drivers.yahoofinance.get(ticker)
DATA = DATA.sort_values("Date", ascending=False).reset_index(drop=True)
DATANOW = DATA.loc[0, "Close"]
DATANOW
# + papermill={} tags=[]
DATAYESTERDAY = DATA.loc[1, "Close"]
DATAYESTERDAY
# + papermill={} tags=[]
VARV = DATANOW - DATAYESTERDAY
VARV = "{:+,.2f}".format(VARV)
VARV
# + papermill={} tags=[]
VARP = ((DATANOW - DATAYESTERDAY) / DATANOW)*100
VARP = "{:+,.2f}".format(VARP)
VARP
# + [markdown] papermill={} tags=[]
# ### Format values
# + papermill={} tags=[]
ARIMA = df.loc[0, "ARIMA"]
ARIMA = round(ARIMA, 1)
ARIMA = "${:,.2f}".format(ARIMA)
ARIMA
# + papermill={} tags=[]
SVR = df.loc[0, "SVR"]
SVR = round(SVR, 1)
SVR = "${:,.2f}".format(SVR)
SVR
# + papermill={} tags=[]
LINEAR = df.loc[0, "LINEAR"]
LINEAR = round(LINEAR, 1)
LINEAR = "${:,.2f}".format(LINEAR)
LINEAR
# + papermill={} tags=[]
COMPOUND = df.loc[0, "COMPOUND"]
COMPOUND = round(COMPOUND, 1)
COMPOUND = "${:,.2f}".format(COMPOUND)
COMPOUND
# + papermill={} tags=[]
DATANOW = round(DATANOW, 1)
DATANOW = "${:,.2f}".format(DATANOW)
DATANOW
# + papermill={} tags=[]
DATAYESTERDAY = round(DATAYESTERDAY, 1)
DATAYESTERDAY = "${:,.2f}".format(DATAYESTERDAY)
DATAYESTERDAY
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Create markdown template
# + papermill={} tags=[]
# %%writefile message.md
Hello world,
The **TICKER** price is **DATANOW** right now, VARV vs yesterday (VARP%).<br>
Yesterday close : DATAYESTERDAY
In +20 days, basic ML models predict the following prices:
- **arima**: ARIMA
- **svr**: SVR
- **linear**: LINEAR
- **compound**: COMPOUND
<img href=link_html target="_blank" src=link_image style="width:640px; height:360px;" /><br>
[Open dynamic chart](link_html)<br>
Please find attached the data in Excel.<br>
Have a nice day.
<br>
PS: You can [send the email again](link_webhook) if you need a fresh update.<br>
<div><strong>Full Name</strong></div>
<div>Open source lover | <a href="http://www.naas.ai/" target="_blank">Naas</a></div>
<div>+ 33 1 23 45 67 89</div>
<div><small>This is an automated email from my Naas account</small></div>
# + papermill={} tags=[]
markdown_file = "message.md"
content = open(markdown_file, "r").read()
md = markdown2.markdown(content)
md
# + [markdown] papermill={} tags=[]
# ### Add email template as a dependency
# + papermill={} tags=[]
naas.dependency.add("message.md")
# + [markdown] papermill={} tags=[]
# ### Add webhook to run your notebook again
# + papermill={} tags=[]
link_webhook = naas.webhook.add()
# + [markdown] papermill={} tags=[]
# ### Replace values in template
# + papermill={} tags=[]
post = md.replace("DATANOW", str(DATANOW))
post = post.replace("TICKER", str(ticker))
post = post.replace("DATAYESTERDAY", str(DATAYESTERDAY))
post = post.replace("VARV", str(VARV))
post = post.replace("VARP", str(VARP))
post = post.replace("LINEAR", str(LINEAR))
post = post.replace("SVR", str(SVR))
post = post.replace("COMPOUND", str(COMPOUND))
post = post.replace("ARIMA", str(ARIMA))
post = post.replace("link_image", str(link_image))
post = post.replace("link_html", str(link_html))
post = post.replace("link_webhook", str(link_webhook))
post
# + [markdown] papermill={} tags=[]
# ### Send by Email
# + papermill={} tags=[]
subject = f"📈 {ticker} predictions as of today"
content = post
files = [f"{ticker}_TODAY.xlsx"]
naas.notification.send(email_to=email_to, subject=subject, html=content, files=files, email_from=email_from)
# + [markdown] papermill={} tags=[]
# ### Schedule every day
# + papermill={} tags=[]
naas.scheduler.add(cron="0 9 * * *")
#naas.scheduler.delete() #if you want to delete the scheduler
| YahooFinance/YahooFinance_Send_daily_prediction_to_Email.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# ## Sagemaker Pipelines
#
# Amazon SageMaker Pipelines is the purpose-built, easy-to-use continuous integration and continuous delivery (CI/CD) service for machine learning (ML).
#
# - SageMaker Integration: SageMaker Pipelines is a fully managed service, which means that it creates and manages resources for you
# - SageMaker Python SDK Integration: you can create your pipelines programmatically using a high-level Python interface that you might already be familiar with
# - SageMaker Studio Integration: offers an environment to manage the end-to-end SageMaker Pipelines experience
# - Data Lineage Tracking: lets you analyze where the data came from, where it was used as an input, and the outputs that were generated from it
#
# To learn more about SageMaker Pipelines, please check
#
# * Doc https://docs.aws.amazon.com/sagemaker/latest/dg/pipelines.html
# * SDK https://sagemaker.readthedocs.io/en/stable/workflows/pipelines/sagemaker.workflow.pipelines.html
#
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import sagemaker
import json
import boto3
from sagemaker import get_execution_role
sm_client = boto3.client('sagemaker')
print(sm_client)
# -
# Retrieve the default bucket
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket()
region = sagemaker_session.boto_region_name
print(region)
print(bucket)
role = get_execution_role()
# Before executing this step, we assume that you have already created a SageMaker Project.
#
# A SageMaker project is an AWS Service Catalog provisioned product that enables you to easily create an end-to-end ML solution. Each SageMaker project has a unique name and ID that are passed to all SageMaker and AWS resources created in the project. By using the name and ID, you can view all entities associated with your project.
# +
project_name = "Replace with your sagemaker project name here"
# project_name = "mlops-cicd-demo"
project_id = sm_client.describe_project(ProjectName=project_name)['ProjectId']
model_package_group_name = project_name + '-' + project_id
print("Model package group name: %s" % model_package_group_name)
# -
# ### Pipeline input parameters
#
# You can introduce variables into your pipeline definition using parameters. Parameters that you define can be referenced throughout your pipeline definition.
# +
from sagemaker.workflow.parameters import ParameterInteger, ParameterString
training_instance_type = ParameterString(
name="TrainingInstanceType",
default_value="ml.m5.xlarge"
)
training_instance_count = ParameterInteger(
name="TrainingInstanceCount",
default_value=1
)
input_raw_data = ParameterString(
name="InputRawData",
default_value='s3://{}/sagemaker/xgboostcontainer/raw-data'.format(bucket)
)
input_train_data = ParameterString(
name="InputDataTrain",
default_value='s3://{}/sagemaker/xgboostcontainer/processed/train'.format(bucket)
)
input_test_data = ParameterString(
name="InputDataTest",
default_value='s3://{}/sagemaker/xgboostcontainer/processed/test'.format(bucket)
)
# -
# ### Preprocessing Step
#
# Amazon SageMaker Processing allows you to run steps for data pre- or post-processing, feature engineering, data validation, or model evaluation workloads on Amazon SageMaker.
#
# A processing step requires a processor, a Python script that defines the processing code, outputs for processing, and job arguments.
from sagemaker.sklearn.processing import SKLearnProcessor
role = get_execution_role()
sklearn_processor = SKLearnProcessor(framework_version='0.20.0',
role=role,
instance_type=training_instance_type,
instance_count=training_instance_count)
# +
from sagemaker.processing import ProcessingInput, ProcessingOutput
from sagemaker.workflow.steps import ProcessingStep
step_process = ProcessingStep(
name="BostonHousingDropColumns",
processor=sklearn_processor,
inputs=[
ProcessingInput(source=input_raw_data, destination="/opt/ml/processing/input"),
],
outputs=[ProcessingOutput(output_name='xgboost_train_data',
source='/opt/ml/processing/output/train',
destination = input_train_data),
ProcessingOutput(output_name='xgboost_test_data',
source='/opt/ml/processing/output/test',
destination = input_test_data)],
code="preprocessing.py"
)
# -
# ### Training Step
#
# You use a training step to create a training job to train a model.
# +
from sagemaker.image_uris import retrieve
from sagemaker.session import Session
# this line automatically looks for the XGBoost image URI and builds an XGBoost container.
# specify the repo_version depending on your preference.
container = retrieve(region=boto3.Session().region_name,
framework='xgboost',
version='1.0-1')
print(container)
# -
# initialize hyperparameters
hyperparameters = {
"max_depth":"10",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"objective":"reg:squarederror",
"num_round":"200"}
# +
# construct a SageMaker estimator that calls the xgboost-container
estimator = sagemaker.estimator.Estimator(image_uri=container,
hyperparameters=hyperparameters,
role=role,
instance_count=1,
instance_type='ml.m5.2xlarge')
# -
# A training step requires an estimator, and training and validation data inputs.
# +
from sagemaker.inputs import TrainingInput
from sagemaker.workflow.steps import TrainingStep
step_train = TrainingStep(
name="TrainingXgBoost",
estimator=estimator,
inputs={
"train": TrainingInput(s3_data=step_process.properties.ProcessingOutputConfig.Outputs["xgboost_train_data"].S3Output.S3Uri, content_type="text/csv"),
"validation": TrainingInput(s3_data=step_process.properties.ProcessingOutputConfig.Outputs["xgboost_test_data"].S3Output.S3Uri, content_type="text/csv"
)
},
)
# -
# ### The Register step that will add a new version to the Model Registry
#
# You use a RegisterModel step to register a model to a model group.
#
# A RegisterModel step requires an estimator, model data output from training, and a model package group name to associate the model package with.
#
# With this step, you can create a model group that tracks all of the models that you train to solve a particular problem. You can then register each model you train and the model registry adds it to the model group as a new model version.
# +
from sagemaker.workflow.step_collections import RegisterModel
# NOTE: model_approval_status is not available as arg in service dsl currently
step_register = RegisterModel(
name="RegisterXgBoostModel",
estimator=estimator,
model_data=step_train.properties.ModelArtifacts.S3ModelArtifacts,
content_types=["text/csv"],
response_types=["text/csv"],
inference_instances=["ml.t2.medium", "ml.m5.large"],
transform_instances=["ml.m5.large"],
model_package_group_name=model_package_group_name
)
# -
# ### Now, we can create the pipeline
#
# This pipeline definition encodes a pipeline using a directed acyclic graph (DAG). This DAG gives information on the requirements for and relationships between each step of your pipeline.
#
# The structure of a pipeline's DAG is determined by the data dependencies between steps (defined within each step previously). These data dependencies are created when the properties of a step's output are passed as the input to another step.
# +
from botocore.exceptions import ClientError, ValidationError
from sagemaker.workflow.pipeline import Pipeline
# NOTE:
# condition steps have issues in service so we go straight to step_register
pipeline_name = "Replace by your pipeline name"
# pipeline_name = "XgBoost-Pipelines-2"
pipeline = Pipeline(
name=pipeline_name,
parameters=[
training_instance_type,
training_instance_count,
input_raw_data,
input_train_data,
input_test_data
],
steps=[step_process,step_train, step_register],
sagemaker_session=sagemaker_session,
)
try:
response = pipeline.create(role_arn=role)
except ClientError as e:
error = e.response["Error"]
if error["Code"] == "ValidationError" and "Pipeline names must be unique within" in error["Message"]:
print(error["Message"])
response = pipeline.describe()
else:
raise
# -
pipeline_arn = response["PipelineArn"]
print(pipeline_arn)
# ### And then, run it
#
# After you’ve created a pipeline definition using the SageMaker Python SDK, you can submit it to SageMaker to start your execution.
# +
import time
start_response = pipeline.start(parameters={
"TrainingInstanceCount": "1"
})
pipeline_execution_arn = start_response.arn
print(pipeline_execution_arn)
while True:
resp = sm_client.describe_pipeline_execution(PipelineExecutionArn=pipeline_execution_arn)
if resp['PipelineExecutionStatus'] == 'Executing':
print('Running...')
else:
print(resp['PipelineExecutionStatus'], pipeline_execution_arn)
break
time.sleep(15)
# -
# ### Finally, approve the model to kick-off the deployment process
# list all packages and select the latest one
packages = sm_client.list_model_packages(ModelPackageGroupName=model_package_group_name)['ModelPackageSummaryList']
packages = sorted(packages, key=lambda x: x['CreationTime'], reverse=True)
packages
# You can either manually approve the model or using sdk
# +
# latest_model_package_arn = packages[0]['ModelPackageArn']
# model_package_update_response = sm_client.update_model_package(
# ModelPackageArn=latest_model_package_arn,
# ModelApprovalStatus="Approved",
# )
# -
# ## Done! :) Let's open the CodePipeline console and get some popcorn to watch
| run_ml_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decomposition framework of the PySAL *segregation* module
#
# This is a notebook that explains a step-by-step procedure to perform decomposition on comparative segregation measures.
#
# First, let's import all the needed libraries.
# +
import pandas as pd
import pickle
import numpy as np
import matplotlib.pyplot as plt
from pysal.explore import segregation
from pysal.explore.segregation.decomposition import DecomposeSegregation
# -
# In this example, we are going to use census data that the user must download its own copy, following similar guidelines explained in https://github.com/spatialucr/geosnap/tree/master/geosnap/data where you should download the full type file of 2010. The zipped file download will have a name that looks like `LTDB_Std_All_fullcount.zip`. After extracting the zipped content, the filepath of the data should looks like this:
# +
#filepath = '~/LTDB_Std_2010_fullcount.csv'
# -
# Then, we read the data:
df = pd.read_csv(filepath, encoding = "ISO-8859-1", sep = ",")
# We are going to work with the variable of the nonhispanic black people (`nhblk10`) and the total population of each unit (`pop10`). So, let's read the map of all census tracts of US and select some specific columns for the analysis:
# +
# This file can be download here: https://drive.google.com/open?id=1gWF0OCn6xuR_WrEj7Ot2jY6KI2t6taIm
with open('data/tracts_US.pkl', 'rb') as input:
map_gpd = pickle.load(input)
map_gpd['INTGEOID10'] = pd.to_numeric(map_gpd["GEOID10"])
gdf_pre = map_gpd.merge(df, left_on = 'INTGEOID10', right_on = 'tractid')
gdf = gdf_pre[['GEOID10', 'geometry', 'pop10', 'nhblk10']]
# -
# In this notebook, we use the Metropolitan Statistical Area (MSA) of US (we're also using the word 'cities' here to refer them). So, let's read the correspondence table that relates the tract id with the corresponding Metropolitan area...
# You can download this file here: https://drive.google.com/open?id=10HUUJSy9dkZS6m4vCVZ-8GiwH0EXqIau
with open('data/tract_metro_corresp.pkl', 'rb') as input:
tract_metro_corresp = pickle.load(input).drop_duplicates()
# ..and merge them with the previous data.
merged_gdf = gdf.merge(tract_metro_corresp, left_on = 'GEOID10', right_on = 'geoid10')
# We now build the composition variable (`compo`) which is the division of the frequency of the chosen group and total population. Let's inspect the first rows of the data.
merged_gdf['compo'] = np.where(merged_gdf['pop10'] == 0, 0, merged_gdf['nhblk10'] / merged_gdf['pop10'])
merged_gdf.head()
# Now, we chose two different metropolitan areas to compare the degree of segregation.
# ## Map of the composition of the Metropolitan area of Los Angeles
la_2010 = merged_gdf.loc[(merged_gdf.name == "Los Angeles-Long Beach-Anaheim, CA")]
la_2010.plot(column = 'compo', figsize = (10, 10), cmap = 'OrRd', legend = True)
plt.axis('off')
# ## Map of the composition of the Metropolitan area of New York
ny_2010 = merged_gdf.loc[(merged_gdf.name == 'New York-Newark-Jersey City, NY-NJ-PA')]
ny_2010.plot(column = 'compo', figsize = (20, 10), cmap = 'OrRd', legend = True)
plt.axis('off')
# We first compare the Gini index of both cities. Let's import the `Gini_Seg` class from `segregation`, fit both indexes and check the difference in point estimation.
# +
from pysal.explore.segregation.aspatial import GiniSeg
G_la = GiniSeg(la_2010, 'nhblk10', 'pop10')
G_ny = GiniSeg(ny_2010, 'nhblk10', 'pop10')
G_la.statistic - G_ny.statistic
# -
# Let's decompose these difference according to *Rey, S. et al "Comparative Spatial Segregation Analytics". Forthcoming*. You can check the options available in this decomposition below:
help(DecomposeSegregation)
# ## Composition Approach (default)
# The difference of -0.10653 fitted previously, can be decomposed into two components. The Spatial component and the attribute component. Let's estimate both, respectively.
DS_composition = DecomposeSegregation(G_la, G_ny)
DS_composition.c_s
DS_composition.c_a
# So, the first thing to notice is that attribute component, i.e., given by a difference in the population structure (in this case, the composition) plays a more important role in the difference, since it has a higher absolute value.
#
# The difference in the composition can be inspected in the plotting method with the type `cdfs`:
DS_composition.plot(plot_type = 'cdfs')
# If your data is a GeoDataFrame, it is also possible to visualize the counterfactual compositions with the argument `plot_type = 'maps'`
#
# The first and second contexts are Los Angeles and New York, respectively.
DS_composition.plot(plot_type = 'maps')
# *Note that in all plotting methods, the title presents each component of the decomposition performed.*
# ## Share Approach
# The share approach takes into consideration the share of each group in each city. Since this approach takes into consideration the focus group and the complementary group share to build the "counterfactual" total population of each unit, it is of interest to inspect all these four cdf's.
#
# *ps.: The share is the population frequency of each group in each unit over the total population of that respectively group.*
DS_share = DecomposeSegregation(G_la, G_ny, counterfactual_approach = 'share')
DS_share.plot(plot_type = 'cdfs')
# We can see that curve between the contexts are closer to each other which represent a drop in the importance of the population structure (attribute component) to -0.062. However, this attribute still overcomes the spatial component (-0.045) in terms of importance due to both absolute magnitudes.
DS_share.plot(plot_type = 'maps')
# We can see that the counterfactual maps of the composition (outside of the main diagonal), in this case, are slightly different from the previous approach.
# ## Dual Composition Approach
#
# The `dual_composition` approach is similar to the composition approach. However, it uses also the counterfactual composition of the cdf of the complementary group.
DS_dual = DecomposeSegregation(G_la, G_ny, counterfactual_approach = 'dual_composition')
DS_dual.plot(plot_type = 'cdfs')
# It is possible to see that the component values are very similar with slight changes from the `composition` approach.
DS_dual.plot(plot_type = 'maps')
# The counterfactual distributions are virtually the same (but not equal) as the one from the `composition` approach.
# ## Inspecting a different index: Relative Concentration
# +
from pysal.explore.segregation.spatial import RelativeConcentration
RCO_la = RelativeConcentration(la_2010, 'nhblk10', 'pop10')
RCO_ny = RelativeConcentration(ny_2010, 'nhblk10', 'pop10')
RCO_la.statistic - RCO_ny.statistic
# -
RCO_DS_composition = DecomposeSegregation(RCO_la, RCO_ny)
RCO_DS_composition.c_s
RCO_DS_composition.c_a
# It is possible to note that, in this case, the spatial component is playing a much more relevant role in the decomposition.
| notebooks/explore/segregation/decomposition_wrapper_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TcEI8iWij_Z3"
# #Import Libraries:
# + id="cWgae7yJpO6v"
import tensorflow
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras import backend as K
from tensorflow.keras import activations
from tensorflow.keras.layers import *
from tensorflow.keras.models import Model, load_model
from tensorflow.keras import models
from tensorflow.keras import layers
import numpy as np
from tqdm import tqdm
import math
import matplotlib.pyplot as plt
# + [markdown] id="Ef6j77WsBhYq"
# # Dataset:
# + id="HWwcWHIKpO_6"
# Load MNIST dataset
(input_train, target_train), (input_test, target_test) = mnist.load_data()
#(input_train, target_train), (input_test, target_test) = tensorflow.keras.datasets.cifar10.load_data()
# + id="Odw8T4J2pqGU" colab={"base_uri": "https://localhost:8080/"} outputId="a49c02e4-ba1c-43aa-dc19-fa35bca6a06b"
input_train = input_train / 255
input_test = input_test / 255
print(f"input_train.shape = {input_train.shape}")
print(f"input_test.shape = {input_test.shape}")
# + colab={"base_uri": "https://localhost:8080/"} id="luezOA85CKuR" outputId="ab3526e7-0b8e-4aa3-d806-234e30fb58aa"
if len(input_test.shape) == 3:
input_train = np.expand_dims(input_train, axis=3)
input_test = np.expand_dims(input_test, axis=3)
print(f"input_train.shape = {input_train.shape}")
print(f"input_test.shape = {input_test.shape}")
else:
print(f"len(input_test.shape) = {len(input_test.shape)}")
# + colab={"base_uri": "https://localhost:8080/"} id="WEzwyCbTgIZe" outputId="4f967dc6-a37e-4efb-94d8-fd8b5cbff116"
num_classes = np.unique(target_test).shape[0]
num_classes
# + id="J4lmZoECfshp"
target_train = tensorflow.keras.utils.to_categorical(target_train, num_classes)
target_test = tensorflow.keras.utils.to_categorical(target_test, num_classes)
# + colab={"base_uri": "https://localhost:8080/"} id="x_-gvBdKgY6C" outputId="fcc6abaa-9292-4790-e10a-bf1abe783df2"
print(f"target_train.shape = {target_train.shape}")
print(f"target_test.shape = {target_test.shape}")
# + colab={"base_uri": "https://localhost:8080/"} id="aPVjov2zetQc" outputId="bf98302f-9d2e-4142-e0b4-c25671925207"
import matplotlib.pyplot as plt
if input_test.shape[-1] == 1:
plt.imshow(input_train[0,:,:,0])
else:
plt.imshow(input_train[0,:,:,:])
plt.show()
# + [markdown] id="041SDwi3qMRE"
# # Model:
# + id="hivPCYiNpO9R"
# Model configuration
batch_size = 250
no_epochs = 1
# + colab={"base_uri": "https://localhost:8080/"} id="b2pSLBOmvjJs" outputId="be52f5a8-a13b-4f72-a165-e7e063462c27"
input_train.shape
# + colab={"base_uri": "https://localhost:8080/"} id="rNA8KbOpvYPa" outputId="bd769a7b-d15f-49d3-e54f-96faa0d98f16"
inputs = Input(input_train.shape[1:])
x = inputs
x = Conv2D(6, kernel_size=(5, 5), activation='relu', name='L1_Conv2D_1')(x)
x = MaxPooling2D(pool_size=(2, 2), name='L2_MaxPooling2D_1')(x)
x = Conv2D(10, kernel_size=(5, 5), activation='relu', name='L3_Conv2D_2')(x)
x = Conv2D(256, kernel_size=(5, 5), activation='relu', name='L4_Conv2D_3')(x)
x = Flatten(name='L4_Flatten_1')(x)
x = Dense(256, activation='relu', name='L5_Dense_1')(x)
outputs = Dense(num_classes, activation='softmax', name='L7_Dense_2')(x)
model=Model(inputs=inputs,outputs=outputs)
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 582} id="slV_u-fgMn7f" outputId="b4f64927-b35e-48af-d9ce-62d38ab5520f"
#'''
tensorflow.keras.utils.plot_model(
model, to_file='model.png', show_shapes=True, show_dtype=False,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=64
)
#'''
# + id="uOTOo_OssYZA"
# Compile the model
model.compile(loss=tensorflow.keras.losses.categorical_crossentropy,
optimizer=tensorflow.keras.optimizers.Adam(),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="-tiMRYxWpqOY" outputId="b68a0664-c01f-4c1f-d105-e4823927bf1e"
# Fit data to model
model.fit(input_train, target_train,
batch_size=batch_size,
epochs=no_epochs,
verbose=1,
#validation_split=validation_split,
validation_data=(input_test, target_test)
)
# + colab={"base_uri": "https://localhost:8080/"} id="RsnL9zGEpqQ0" outputId="784ec1a2-5824-49be-ef8b-a6d2adc3734b"
# Generate generalization metrics
score = model.evaluate(input_test, target_test, verbose=1)
print(f'Test loss: {score[0]} / Test accuracy: {score[1]}')
# + id="dYbgvDJfsdqy"
# + [markdown] id="P5MTfgXP5hbg"
# #Layer Reversing:
# + colab={"base_uri": "https://localhost:8080/"} id="Qf8-3kNo5g0n" outputId="528b170c-51e1-49e3-9e1f-1f5885b3b76f"
layer_outputs = [layer.output for layer in model.layers]
print(f"Total number of layers = {len(activations)}")
print(f"Number of images fed into model = {len(activations[0])}")
# + colab={"base_uri": "https://localhost:8080/"} id="VHlU4-NL5yBx" outputId="971b1e7a-692c-426b-ba40-9c006642d4ab"
layer_names = []
for layer in model.layers:
layer_names.append(layer.name)
print(layer_names)
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="3eaAKsY26jD7" outputId="024f6c33-203b-42b5-c5f3-f5d923f604af"
layer_names[4]
# + colab={"base_uri": "https://localhost:8080/"} id="Hoz1GrKE6vex" outputId="5f36649f-273d-4f0b-c425-9e38fc5a1046"
layer_output = activations[4]
layer_output.shape
# + id="sB3TIi3AD2Z6"
# + colab={"base_uri": "https://localhost:8080/"} id="zab9no5q5yE6" outputId="d1236678-ec0f-454a-e2d8-c1e82710ab22"
inputs = Input(layer_output.shape[1:])
x = inputs
x = Conv2DTranspose(256, kernel_size=(5, 5), activation='relu', name='Transpose_1_L4_Conv2D_3')(x)
x = Conv2DTranspose(10, kernel_size=(5, 5), activation='relu', name='Transpose_2_L3_Conv2D_2')(x)
x = UpSampling2D(size=(2, 2), interpolation="nearest", name='UpSample_1_L2_MaxPooling2D_1')(x)
x = Conv2DTranspose(6, kernel_size=(5, 5), activation='relu', name='Transpose_3_L1_Conv2D_1')(x)
x = Flatten(name='Flatten_1')(x)
outputs = Dense(num_classes, activation='softmax', name='Dense_1')(x)
model_reverse=Model(inputs=inputs,outputs=outputs)
model_reverse.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 509} id="OTyxkreN5yHs" outputId="db6396f2-df7e-437c-990a-080ccf7d6a46"
#'''
tensorflow.keras.utils.plot_model(
model_reverse, to_file='model.png', show_shapes=True, show_dtype=False,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=64
)
#'''
# + id="kyVavKNe5yKY"
# Compile the model
model_reverse.compile(loss=tensorflow.keras.losses.categorical_crossentropy,
optimizer=tensorflow.keras.optimizers.Adam(),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="b0IL14So5yNJ" outputId="23bed531-d547-470f-895f-501cc62012b8"
model_reverse.fit(layer_output, target_test,
batch_size=batch_size,
epochs=no_epochs,
verbose=1
)
# + id="GvH5j59s5yVN"
# + [markdown] id="-Y6GNd6uqzOV"
# # Generating layer output visualizations
# + id="1JAz5fZnbaLS"
reverse_layer_outputs = [layer.output for layer in model_reverse.layers]
activation_model = models.Model(inputs=model_reverse.input, outputs=reverse_layer_outputs)
activations = activation_model.predict(layer_output)
# + colab={"base_uri": "https://localhost:8080/"} id="e-12IFyJbaUs" outputId="6f584035-3f79-4143-dd2e-45b4ef52bdd4"
print(f"Total number of layers = {len(activations)}")
print(f"Number of images fed into model = {len(activations[0])}")
# + id="MkrcIDvYbaag"
# Getting Activations of first layer
first_layer_activation = activations[0]
# + colab={"base_uri": "https://localhost:8080/"} id="0vJ37I08bae1" outputId="a97eaad2-82d3-4b45-d0bf-7b1ed4afaa70"
# shape of first layer activation
print(first_layer_activation.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="sz-x2aHhbah1" outputId="150413c9-7701-4216-f3b4-db1352180ef2"
if input_test.shape[-1] == 3:
print(f"Case-1: input_test.shape[-1] = {input_test.shape[-1]}\n")
plt.matshow(first_layer_activation[0, :, :, :], cmap ='viridis')
else:
print(f"Case-2: input_test.shape[-1] = {input_test.shape[-1]}\n")
plt.matshow(first_layer_activation[0, :, :, 0], cmap ='viridis')
plt.show()
# + [markdown] id="pYpt3Ju0j31V"
# #Model Layers:
# + colab={"base_uri": "https://localhost:8080/"} id="kMPdMN0bba1c" outputId="a1b033ec-a0f4-4110-d6bb-8ff278d52492"
layer_names = []
for layer in model_reverse.layers:
layer_names.append(layer.name)
print(layer_names)
# + [markdown] id="HGuU7zXr_sbh"
# #Visualize Activation Output Images in between a Range:
# + colab={"base_uri": "https://localhost:8080/"} id="W8DdBlUML7OK" outputId="c9a1f845-c91e-4a9e-eb4a-7f256da8a225"
layer_no = -3 # starts from 0
current_layer = activations[layer_no]
print(f"current_layer.shape = {current_layer.shape}")
print(f"image_dimension = {current_layer.shape[1:][:-1]}")
print(f"num_neurons = {current_layer.shape[1:][-1]}\n")
# + colab={"base_uri": "https://localhost:8080/"} id="kobg-FgoNe9A" outputId="9b081ab6-4819-4603-af3f-97c729e0cdce"
current_layer[0].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 754} id="JLWMfAlUadEB" outputId="70503ab3-8f4c-497b-f2db-74d80122cdca"
sub_fig_num_rows = 10 #current_layer.shape[0]
sub_fig_num_cols = current_layer.shape[-1] # current_layer.shape[-1], 10
fig_heigth = 10
fig_width = 10
if len(current_layer.shape) == 4:
fig, axes = plt.subplots(sub_fig_num_rows,sub_fig_num_cols, figsize=(fig_width,fig_heigth))
plt.suptitle(f"Layer {str(layer_no+1)}: {layer_names[layer_no]} {str(current_layer.shape[1:])}", fontsize=20, y=1.1)
for i,ax in tqdm(enumerate(axes.flat)):
row = i//sub_fig_num_cols
col = i%sub_fig_num_cols
ax.imshow(current_layer[row, :, :, col], cmap ='viridis')
ax.set_xticks([])
ax.set_yticks([])
if col == 0:
ax.set_ylabel(f"image {str(row+1)}")
if row == 0:
ax.set_xlabel(f"activation {str(col+1)}", rotation=90, ha='right')
ax.xaxis.set_label_position('top')
ax.set_aspect('auto')
#fig.tight_layout()
plt.subplots_adjust(wspace=0, hspace=0.05)
else:
print(f"Error: len(current_layer.shape) = {len(current_layer.shape)}. However, it has to be 4.")
# + id="hH2Qk9c7adGm"
# + [markdown] id="ktdie3scBFeV"
# #Visualize Certain Activation Output Images:
# + id="DSAjHtSzadJg" colab={"base_uri": "https://localhost:8080/", "height": 719} outputId="68cbc598-4e17-459b-c41b-aa7ed64c8073"
layer_no = 2 # starts from 0
current_layer = activations[layer_no]
print(f"image_dimension = {current_layer.shape[1:][:-1]}\nnum_neurons = {current_layer.shape[1:][-1]}\n")
image_index_list = [0, 9, 99, 999, 9999]
sub_fig_num_rows = len(image_index_list) # current_layer.shape[0]
sub_fig_num_cols = current_layer.shape[-1] # total number of activations in a layer
fig_heigth = 10
fig_width = 12
if len(current_layer.shape) == 4:
fig, axes = plt.subplots(sub_fig_num_rows,sub_fig_num_cols, figsize=(fig_width,fig_heigth))
plt.suptitle(f"Layer {str(layer_no+1)}: {layer_names[layer_no]} {str(current_layer.shape[1:])}", fontsize=20, y=1)
for i,ax in enumerate(axes.flat):
row = i//sub_fig_num_cols
col = i%sub_fig_num_cols
image_index = image_index_list[row]
ax.imshow(current_layer[image_index, :, :, col], cmap ='viridis')
ax.set_xticks([])
ax.set_yticks([])
if col == 0:
ax.set_ylabel(f"image {str(image_index+1)}")
if row == 0:
ax.set_xlabel(f"activation {str(col+1)}", rotation=45, ha='right')
ax.xaxis.set_label_position('top')
ax.set_aspect('auto')
#fig.tight_layout()
plt.subplots_adjust(wspace=0, hspace=0.02)
else:
print(f"Error: len(current_layer.shape) = {len(current_layer.shape)}. However, it has to be 4.")
# + id="ghq902UpadMD"
# + id="4BZF7zEJadO3" colab={"base_uri": "https://localhost:8080/"} outputId="5c410942-3ac7-4b73-b721-b08f95ea9a9c"
layer_no = -2 # starts from 0
current_layer = activations[layer_no]
print(f"image_dimension = {current_layer.shape}")
print(f"image_dimension = {current_layer.shape[1:]}")
print(f"num_neurons = {current_layer.shape[1]}\n")
# + id="6pc67XJFadRq" colab={"base_uri": "https://localhost:8080/"} outputId="c27c5ff3-fdfe-41fd-df45-fe6789dae668"
dense_output = current_layer[0]
print(f"type(dense_output) = {type(dense_output)}")
print(f"dense_output.shape = {dense_output.shape}")
# + id="vj5FNY4madXS" colab={"base_uri": "https://localhost:8080/"} outputId="d6fc73e0-2190-405e-b229-858633c1a2fa"
num_sqrt = math.floor(math.sqrt(dense_output.shape[0]))
img_shape = (num_sqrt,num_sqrt)
print(f"img_shape = {img_shape}")
print(f"Total pixels counted = {num_sqrt*num_sqrt}")
# + colab={"base_uri": "https://localhost:8080/"} id="B0vh0SmlsavR" outputId="1a03ea51-6622-46e5-ca04-538893a7f188"
dense_output_trunc = dense_output[:(num_sqrt*num_sqrt)]
num_pixel_loss = dense_output.shape[0]-len(dense_output_trunc)
print(f"dense_output_trunc.shape = {dense_output_trunc.shape}")
print(f"num_pixel_loss = {num_pixel_loss}")
# + id="jxuLrHA4u6x4" colab={"base_uri": "https://localhost:8080/"} outputId="a89fc038-ed61-4f0d-e246-001057ee05af"
reshaped_output = np.reshape(dense_output_trunc, (img_shape))
reshaped_output.shape
# + id="_D8oMEJRu66c" colab={"base_uri": "https://localhost:8080/", "height": 293} outputId="0137368c-d9a3-4bcc-d615-7eb56bdcf83d"
#plt.matshow(np.array([dense_output]), cmap ='viridis')
plt.matshow(reshaped_output, cmap ='viridis')
# + id="O_074RXdu7Ao"
| Plant_Leaf_MalayaKew_MK_Dataset/Model_Transpose_Input_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install --quiet gTTS playsound
# +
from gtts import gTTS
from playsound import playsound
import os
file= 'speak2.mp3'
os.remove(file)
speak = input("What do you want me to say? ")
tts = gTTS(speak, lang='en')
tts.save(file)
playsound(file)
# -
from gtts import gTTS
from playsound import playsound
import os
file = 'audio.mp3'
tts = gTTS('hello', lang='en')
tts.save(file)
playsound(file)
os.remove(file)
help (gTTS)
| content/lessons/07/In-Class-Text-To-Speech-Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Layerwise Relevance Propagation
# ### CHAPTER 02 - *Model Explainability Methods*
#
# From **Applied Machine Learning Explainability Techniques** by [**<NAME>**](https://www.linkedin.com/in/aditya-bhattacharya-b59155b6/), published by **Packt**
# ### Objective
#
# In this notebook, we will try to implement some of the concepts related to Layerwise Relevance Propagation part of the Influence based explainability methods discussed in Chapter 2 - Model Explainability Methods.
# ### Installing the modules
# Install the following libraries in Google Colab or your local environment, if not already installed.
# !pip install --upgrade numpy matplotlib tensorflow
# ### Loading the modules
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as c_map
from IPython.display import Image, display
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.applications.xception import Xception, preprocess_input, decode_predictions
from tensorflow.keras.preprocessing import image
import warnings
warnings.filterwarnings("ignore")
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# ### Gradient Class Activation Map (Grad-CAM)
# The popular technique of Grad-CAM is used as a visualization technique to explain the working of complex Convolution Neural Networks on unstrutured data like images. We will use tensorflow and keras framework to get a pretrained network on ImageNet dataset and test the approach on a sample open image obtained from the source: https://i.imgur.com/GleAY3f.jpeg. For more examples using Keras and tensorflow please visit: https://keras.io/examples/
model_builder = Xception
preprocess_input = preprocess_input
decode_predictions = decode_predictions
IMG_SIZE = (299, 299)
last_conv_layer = "block14_sepconv2_act"
# ### Loading the data
# +
# The local path to our target image
image_path = keras.utils.get_file(
"tiger.jpg", "https://i.imgur.com/GleAY3f.jpeg"
)
display(Image(image_path))
# -
# ### Preprocessing
def vectorize_image(img_path, size):
'''
Vectorize the given image to get a numpy array
'''
img = image.load_img(img_path, target_size=size)
array = image.img_to_array(img)
array = np.expand_dims(array, axis=0) # Adding dimension to convert array into a batch of size (1,299,299,3)
return array
# +
vectorized_image = preprocess_input(vectorize_image(image_path, size=IMG_SIZE))
model = model_builder(weights="imagenet")
model.layers[-1].activation = None # Removing the last layer as it is the softmax layer used for classification
model_prediction = model.predict(vectorized_image)
print(f"The predicted class is : {decode_predictions(model_prediction, top=1)[0][0][1]}")
# -
# ### Building Grad-CAM Heat-map
# +
def get_heatmap(vectorized_image, model, last_conv_layer, pred_index=None):
'''
Function to visualize grad-cam heatmaps
'''
gradient_model = tf.keras.models.Model(
[model.inputs], [model.get_layer(last_conv_layer).output, model.output]
)
# Gradient Computations
with tf.GradientTape() as tape:
last_conv_layer_output, preds = gradient_model(vectorized_image)
if pred_index is None:
pred_index = tf.argmax(preds[0])
class_channel = preds[:, pred_index]
grads = tape.gradient(class_channel, last_conv_layer_output)
pooled_grads = tf.reduce_mean(grads, axis=(0, 1, 2))
last_conv_layer_output = last_conv_layer_output[0]
heatmap = last_conv_layer_output @ pooled_grads[..., tf.newaxis]
heatmap = tf.squeeze(heatmap)
heatmap = tf.maximum(heatmap, 0) / tf.math.reduce_max(heatmap) # Normalize the heatmap
return heatmap.numpy()
plt.matshow(get_heatmap(vectorized_image, model, last_conv_layer))
plt.show()
# -
# ### Superimpose Grad-CAM Heatmap on image
# +
def superimpose_gradcam(img_path, heatmap, output_path="grad_cam_image.jpg", alpha=0.4):
'''
Superimpose Grad-CAM Heatmap on image
'''
img = image.load_img(img_path)
img = image.img_to_array(img)
heatmap = np.uint8(255 * heatmap) # Back scaling to 0-255 from 0 - 1
jet = c_map.get_cmap("jet") # Colorizing heatmap
jet_colors = jet(np.arange(256))[:, :3] # Using RGB values
jet_heatmap = jet_colors[heatmap]
jet_heatmap = image.array_to_img(jet_heatmap)
jet_heatmap = jet_heatmap.resize((img.shape[1], img.shape[0]))
jet_heatmap = image.img_to_array(jet_heatmap)
superimposed_img = jet_heatmap * alpha + img # Superimposing the heatmap on original image
superimposed_img = image.array_to_img(superimposed_img)
superimposed_img.save(output_path) # Saving the superimposed image
display(Image(output_path)) # Displaying Grad-CAM Superimposed Image
superimpose_gradcam(image_path, get_heatmap(vectorized_image, model, last_conv_layer))
# -
# ### Final Thoughts
# This is a very powerful technique that is used to explain the working of complex Deep Learning algorithms on unstructured data like images. Although this method is difficult to unerstand for beginner learners, but once you get a hang of it, it is a very powerful method and very helpful for model explainability,
# ### Reference
# 1. Keras Tensorflow Tutorial Examples - https://keras.io/examples/
| Chapter02/Layerwise Propagation.ipynb |
# ---
# jupyter:
# jupytext:
# formats: python_scripts//py:percent,notebooks//ipynb
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Introduction to scikit-learn: basic model hyper-parameters tuning
#
# The process of learning a predictive model is driven by a set of internal
# parameters and a set of training data. These internal parameters are called
# hyper-parameters and are specific for each family of models. In addition, a
# specific set of parameters are optimal for a specific dataset and thus they
# need to be optimized.
#
# This notebook shows:
# * the influence of changing model parameters;
# * how to tune these hyper-parameters;
# * how to evaluate the model performance together with hyper-parameter
# tuning.
# %%
import pandas as pd
df = pd.read_csv(
"https://www.openml.org/data/get_csv/1595261/adult-census.csv")
# Or use the local copy:
# df = pd.read_csv(os.path.join("..", "datasets", "adult-census.csv"))
# %%
target_name = "class"
target = df[target_name].to_numpy()
target
# %%
data = df.drop(columns=[target_name, "fnlwgt"])
data.head()
# %% [markdown]
# Once the dataset is loaded, we split it into a training and testing sets.
# %%
from sklearn.model_selection import train_test_split
df_train, df_test, target_train, target_test = train_test_split(
data, target, random_state=42)
# %% [markdown]
# Then, we define the preprocessing pipeline to transform differently
# the numerical and categorical data.
# %%
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OrdinalEncoder
categorical_columns = [
'workclass', 'education', 'marital-status', 'occupation',
'relationship', 'race', 'native-country', 'sex']
categories = [
data[column].unique() for column in data[categorical_columns]]
categorical_preprocessor = OrdinalEncoder(categories=categories)
preprocessor = ColumnTransformer([
('cat-preprocessor', categorical_preprocessor,
categorical_columns),], remainder='passthrough',
sparse_threshold=0)
# %% [markdown]
# Finally, we use a tree-based classifier (i.e. histogram gradient-boosting) to
# predict whether or not a person earns more than 50,000 dollars a year.
# %%
# %%time
# for the moment this line is required to import HistGradientBoostingClassifier
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingClassifier
from sklearn.pipeline import Pipeline
model = Pipeline([
("preprocessor", preprocessor),
("classifier",
HistGradientBoostingClassifier(max_leaf_nodes=16,
learning_rate=0.05,
random_state=42)),])
model.fit(df_train, target_train)
print(
f"The test accuracy score of the gradient boosting pipeline is: "
f"{model.score(df_test, target_test):.2f}")
# %% [markdown]
# ## Quizz
#
# 1. What is the default value of the `learning_rate` parameter of the `HistGradientBoostingClassifier` class? ([link to the API documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.HistGradientBoostingClassifier.html#sklearn-ensemble-histgradientboostingclassifier))
#
# 2. Try to edit the code of the previous cell to set the learning rate parameter to 10. Does this increase the accuracy of the model?
#
# 3. Decrease progressively value of `learning_rate`: can you find a value that yields an accuracy higher than 0.87?
#
# 4. Fix `learning_rate` to 0.05 and try setting the value of `max_leaf_nodes` to the minimum value of 2. Does not improve the accuracy?
#
# 5. Try to progressively increase the value of `max_leaf_nodes` to 256 by taking powers of 2. What do you observe?
# %% [markdown]
# ## The issue of finding the best model parameters
#
# In the previous example, we created an histogram gradient-boosting classifier
# using the default parameters by omitting to explicitely set these parameters.
#
# However, there is no reasons that this set of parameters are optimal for our
# dataset. For instance, fine-tuning the histogram gradient-boosting can be
# achieved by finding the best combination of the following parameters: (i)
# `learning_rate`, (ii) `min_samples_leaf`, and (iii) `max_leaf_nodes`.
# Nevertheless, finding this combination manually will be tedious. Indeed,
# there are relationship between these parameters which are difficult to find
# manually: increasing the depth of trees (increasing `max_samples_leaf`)
# should be associated with a lower learning-rate.
#
# Scikit-learn provides tools to explore and evaluate the parameters
# space.
# %% [markdown]
# ## Finding the best model hyper-parameters via exhaustive parameters search
#
# Our goal is to find the best combination of the parameters stated above.
#
# In short, we will set these parameters with some defined values, train our
# model on some data, and evaluate the model performance on some left out data.
# Ideally, we will select the parameters leading to the optimal performance on
# the testing set.
# %% [markdown]
# The first step is to find the name of the parameters to be set. We use the
# method `get_params()` to get this information. For instance, for a single
# model like the `HistGradientBoostingClassifier`, we can get the list such as:
# %%
print("The hyper-parameters are for a histogram GBDT model are:")
for param_name in HistGradientBoostingClassifier().get_params().keys(
):
print(param_name)
# %% [markdown]
# When the model of interest is a `Pipeline`, i.e. a serie of transformers and
# a predictor, the name of the estimator will be added at the front of the
# parameter name with a double underscore ("dunder") in-between (e.g.
# `estimator__parameters`).
# %%
print("The hyper-parameters are for the full-pipeline are:")
for param_name in model.get_params().keys():
print(param_name)
# %% [markdown]
# The parameters that we want to set are:
# - `'classifier__learning_rate'`: this parameter will
# control the ability of a new tree to correct the error of the previous
# sequence of trees;
# - `'classifier__max_leaf_nodes'`: this parameter will
# control the depth of each tree.
# %% [markdown]
# ## Exercises:
#
# Use the previously defined model (called `model`) and using two nested `for`
# loops, make a search of the best combinations of the `learning_rate` and
# `max_leaf_nodes` parameters. In this regard, you will need to train and test
# the model by setting the parameters. The evaluation of the model should be
# performed using `cross_val_score`. We can propose to define the following
# parameters search:
# - `learning_rate` for the values 0.01, 0.1, and 1;
# - `max_leaf_nodes` for the values 5, 25, 45.
# %% [markdown]
# ## Automated parameter tuning via grid-search
#
# Instead of manually writting the two `for` loops, scikit-learn provides a
# class called `GridSearchCV` which implement the exhaustive search implemented
# during the exercise.
#
# Let see how to use the `GridSearchCV` estimator for doing such search.
# Since the grid-search will be costly, we will only explore the combination
# learning-rate and the maximum number of nodes.
# %%
# %%time
import numpy as np
from sklearn.model_selection import GridSearchCV
param_grid = {
'classifier__learning_rate': (0.05, 0.1, 0.5, 1, 5),
'classifier__max_leaf_nodes': (3, 10, 30, 100),}
model_grid_search = GridSearchCV(model, param_grid=param_grid,
n_jobs=4, cv=2)
model_grid_search.fit(df_train, target_train)
print(f"The test accuracy score of the grid-searched pipeline is: "
f"{model_grid_search.score(df_test, target_test):.2f}")
# %% [markdown]
# The `GridSearchCV` estimator takes a `param_grid` parameter which defines
# all hyper-parameters and their associated values. The grid-search will be in
# charge of creating all possible combinations and test them.
#
# The number of combinations will be equal to the cardesian product of the
# number of values to explore for each parameter (e.g. in our example 3 x 3
# combinations). Thus, adding new parameters with their associated values to be
# explored become rapidly computationally expensive.
#
# Once the grid-search is fitted, it can be used as any other predictor by
# calling `predict` and `predict_proba`. Internally, it will use the model with
# the best parameters found during `fit`.
#
# Get predictions for the 5 first samples using the estimator with the best
# parameters.
# %%
model_grid_search.predict(df_test.iloc[0:5])
# %% [markdown]
# You can know about these parameters by looking at the `best_params_`
# attribute.
# %%
print(f"The best set of parameters is: "
f"{model_grid_search.best_params_}")
# %% [markdown]
# In addition, we can inspect all results which are stored in the attribute
# `cv_results_` of the grid-search. We will filter some specific columns to
# from these results
# %%
cv_results = pd.DataFrame(model_grid_search.cv_results_).sort_values(
"mean_test_score", ascending=False)
cv_results.head()
# %% [markdown]
# Let us focus on the most interesting columns and shorten the parameter names to remove the `"param_classifier__"` prefix for readability:
# %%
# get the parameter names
column_results = [f"param_{name}" for name in param_grid.keys()]
column_results += [
"mean_test_score", "std_test_score", "rank_test_score"]
cv_results = cv_results[column_results]
# %%
def shorten_param(param_name):
if "__" in param_name:
return param_name.rsplit("__", 1)[1]
return param_name
cv_results = cv_results.rename(shorten_param, axis=1)
cv_results
# %% [markdown]
# With only 2 parameters, we might want to visualize the grid-search as a
# heatmap. We need to transform our `cv_results` into a dataframe where the
# rows will correspond to the learning-rate values and the columns will
# correspond to the maximum number of leaf and the content of the dataframe
# will be the mean test scores.
# %%
pivoted_cv_results = cv_results.pivot_table(
values="mean_test_score", index=["learning_rate"],
columns=["max_leaf_nodes"])
pivoted_cv_results
# %%
import matplotlib.pyplot as plt
from seaborn import heatmap
ax = heatmap(pivoted_cv_results, annot=True, cmap="YlGnBu", vmin=0.7,
vmax=0.9)
ax.invert_yaxis()
# %% [markdown]
# The above tables highlights the following things:
#
# - for too high values of the value of `learning_rate`, the performance of the model is degraded and adjusting the value of `max_leaf_nodes` cannot fix that problem;
# - outside of this pathological region, we observe that the optimal choice of `max_leaf_nodes` depends on the value of `learning_rate`;
# - in particular, we observe a "diagonal" of good models with an accuracy close to the maximal of 0.87: when the value of `max_leaf_nodes` is increased, one should increase the value of `learning_rate` accordingly to preserve a good accuracy.
#
# The precise meaning of those two parameters will be explained in a latter notebook.
#
# For now we will note that, in general, **there is no unique optimal parameter setting**: 6 models out of the 16 parameter configuration reach the maximal accuracy (up to smal random fluctuations caused by the sampling of the training set).
# %% [markdown]
# ## Hyper-parameter tuning with Random Search
#
#
# With the `GridSearchCV` estimator, the parameters need to be specified
# explicitely. We mentioned that exploring a large number of values for
# different parameters will be quickly untractable.
#
# Instead, we can randomly generate the parameter candidates. The
# `RandomSearchCV` allows for such stochastic search. It is used similarly to
# the `GridSearchCV` but the sampling distributions need to be specified
# instead of the parameter values. For instance, we will draw candidates using
# a log-uniform distribution also called reciprocal distribution. In addition,
# we will optimize 2 other parameters:
# - `max_iter`: it corresponds to the number of trees in the ensemble;
# - `min_samples_leaf`: it corresponds to the minimum number of samples
# required in a leaf.
# %%
from scipy.stats import reciprocal
from sklearn.model_selection import RandomizedSearchCV
from pprint import pprint
class reciprocal_int:
"""Integer valued version of the log-uniform distribution"""
def __init__(self, a, b):
self._distribution = reciprocal(a, b)
def rvs(self, *args, **kwargs):
"""Random variable sample"""
return self._distribution.rvs(*args, **kwargs).astype(int)
param_distributions = {
'classifier__l2_regularization': reciprocal(1e-6, 1e3),
'classifier__learning_rate': reciprocal(0.001, 10),
'classifier__max_leaf_nodes': reciprocal_int(2, 256),
'classifier__min_samples_leaf': reciprocal_int(1, 100),
'classifier__max_bins': reciprocal_int(2, 255),}
model_random_search = RandomizedSearchCV(
model, param_distributions=param_distributions, n_iter=10,
n_jobs=4, cv=5)
model_random_search.fit(df_train, target_train)
print(f"The test accuracy score of the best model is "
f"{model_random_search.score(df_test, target_test):.2f}")
# %%
print("The best parameters are:")
pprint(model_random_search.best_params_)
# %% [markdown]
# We can inspect the results using the attributes `cv_results` as we previously
# did.
# %%
# get the parameter names
column_results = [
f"param_{name}" for name in param_distributions.keys()]
column_results += [
"mean_test_score", "std_test_score", "rank_test_score"]
cv_results = pd.DataFrame(model_random_search.cv_results_)
cv_results = cv_results[column_results].sort_values(
"mean_test_score", ascending=False)
cv_results = cv_results.rename(shorten_param, axis=1)
cv_results
# %% [markdown]
# In practice, a randomized hyper-parameter search is usually run with a large number of
# iterations. In order to avoid the computation cost and still make a decent
# analysis, we load the results obtained from a similar search with 200
# iterations.
# %%
# model_random_search = RandomizedSearchCV(
# model, param_distributions=param_distributions, n_iter=500,
# n_jobs=4, cv=5)
# model_random_search.fit(df_train, target_train)
# cv_results = pd.DataFrame(model_random_search.cv_results_)
# cv_results.to_csv("../figures/randomized_search_results.csv")
# %%
cv_results = pd.read_csv("../figures/randomized_search_results.csv",
index_col=0)
# %% [markdown]
# As we have more than 2 paramters in our grid-search, we cannot visualize the
# results using a heatmap. However, we can us a parallel coordinates plot.
# %%
(cv_results[column_results].rename(
shorten_param, axis=1).sort_values("mean_test_score"))
# %%
import plotly.express as px
fig = px.parallel_coordinates(
cv_results.rename(shorten_param, axis=1).apply({
"learning_rate": np.log10,
"max_leaf_nodes": np.log2,
"max_bins": np.log2,
"min_samples_leaf": np.log10,
"l2_regularization": np.log10,
"mean_test_score": lambda x: x,}),
color="mean_test_score",
color_continuous_scale=px.colors.sequential.Viridis,
)
fig.show()
# %% [markdown]
# The parallel coordinates plot will display the values of the hyper-parameters
# on different columns while the performance metric is color coded. Thus, we
# are able to quickly inspect if there is a range of hyper-parameters which is
# working or not.
#
# Note that we **transformed most axis values by taking a log10 or log2** to
# spread the active ranges and improve the readability of the plot.
#
# It is possible to **select a range of results by clicking and holding on
# any axis** of the parallel coordinate plot. You can then slide (move)
# the range selection and cross two selections to see the intersections.
# %% [markdown]
# **Quizz**
#
#
# Select the worst performing models (for instance models with a "mean_test_score" lower than 0.7): what do have all these moels in common (choose one):
#
#
# | | |
# |-------------------------------|------|
# | too large `l2_regularization` | |
# | too small `l2_regularization` | |
# | too large `learning_rate` | |
# | too low `learning_rate` | |
# | too large `max_bins` | |
# | too large `max_bins` | |
#
#
# Using the above plot, identify ranges of values for hyperparameter that always prevent the model to reach a test score higher than 0.86, irrespective of the other values:
#
#
# | | True | False |
# |-------------------------------|------|-------|
# | too large `l2_regularization` | | |
# | too small `l2_regularization` | | |
# | too large `learning_rate` | | |
# | too low `learning_rate` | | |
# | too large `max_bins` | | |
# | too large `max_bins` | | |
# %% [markdown]
# ## Exercises:
#
# - Build a machine learning pipeline:
# * preprocess the categorical columns using a `OneHotEncoder` and use
# a `StandardScaler` to normalize the numerical data.
# * use a `LogisticRegression` as a predictive model.
# - Make an hyper-parameters search using `RandomizedSearchCV` and tuning the
# parameters:
# * `C` with values ranging from 0.001 to 10. You can use a reciprocal
# distribution (i.e. `scipy.stats.reciprocal`);
# * `solver` with possible values being `"liblinear"` and `"lbfgs"`;
# * `penalty` with possible values being `"l2"` and `"l1"`;
# * `drop` with possible values being `None` or `"first"`.
#
# You might get some `FitFailedWarning` and try to explain why.
# %% [markdown]
# ## Combining evaluation and hyper-parameters search
#
# Cross-validation was used for searching for the best model parameters. We
# previously evaluated model performance through cross-validation as well. If
# we would like to combine both aspects, we need to perform a **"nested"
# cross-validation**. The "outer" cross-validation is applied to assess the model
# while the "inner" cross-validation sets the hyper-parameters of the model on
# the data set provided by the "outer" cross-validation.
#
#
# In practice, it can be implemented by calling `cross_val_score` or
# `cross_validate` on an instance of `GridSearchCV`, `RandomSearchCV`, or any
# other `EstimatorCV` class.
# %%
from sklearn.model_selection import cross_val_score
# recall the definition of our grid-search
param_distributions = {
'classifier__max_iter': reciprocal_int(10, 50),
'classifier__learning_rate': reciprocal(0.01, 10),
'classifier__max_leaf_nodes': reciprocal_int(2, 16),
'classifier__min_samples_leaf': reciprocal_int(1, 50),}
model_random_search = RandomizedSearchCV(
model, param_distributions=param_distributions, n_iter=10,
n_jobs=4, cv=5)
scores = cross_val_score(model_random_search, data, target, n_jobs=4,
cv=5)
# %%
print(f"The cross-validated accuracy score is:"
f" {scores.mean():.3f} +- {scores.std():.3f}")
# %%
print("The scores obtained for each CV split are:")
print(scores)
# %% [markdown]
# Be aware that the best model found for each split of the outer cross-validation loop might not share the same hyper-parameter values.
#
# When analyzing such model, you should not only look at the
# overall model performance but look at the hyper-parameters variations as
# well.
# %% [markdown]
# ## In this notebook, we have:
#
# * manually tuned the hyper-parameters of a machine-learning pipeline;
# * automatically tuned the hyper-parameters of a machine-learning pipeline by
# by exhaustively searching the best combination of parameters from a defined
# grid;
# * automatically tuned the hyper-parameters of a machine-learning pipeline by
# drawing values candidates from some predefined distributions;
# * nested an hyper-parameters tuning procedure within an cross-validation
# evaluation procedure.
#
# ## Main take-away points
#
# * a grid-search is a costly exhaustive search and does scale with the number of
# parameters to search;
# * a randomized-search will always run with a fixed given budget;
# * when assessing the performance of a model, hyper-parameters search should
# be tuned on the training data of a predifined train test split;
# * alternatively it is possible to nest parameter tuning within a
# cross-validation scheme.
# %%
| notebooks/04_basic_parameters_tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 数据集定义与加载
#
#
# 深度学习模型需要大量的数据来完成训练和评估,这些数据样本可能是图片(image)、文本(text)、语音(audio)等多种类型,而模型训练过程实际是数学计算过程,因此数据样本在送入模型前需要经过一系列处理,如转换数据格式、划分数据集、变换数据形状(shape)、制作数据迭代读取器以备分批训练等。
#
# 在飞桨框架中,可通过如下两个核心步骤完成数据集的定义与加载:
#
# 1. **定义数据集**:将磁盘中保存的原始图片、文字等样本和对应的标签映射到 Dataset,方便后续通过索引(index)读取数据,在 Dataset 中还可以进行一些数据变换、数据增广等预处理操作。在飞桨框架中推荐使用 [paddle.io.Dataset](../../api/paddle/io/Dataset_cn.html#dataset) 自定义数据集,另外在 [paddle.vision.datasets](../../api/paddle/vision/Overview_cn.html#api) 和 [paddle.text](../../api/paddle/text/Overview_cn.html#api) 目录下飞桨内置了一些经典数据集方便直接调用。
#
#
# 2. **迭代读取数据集**:自动将数据集的样本进行分批(batch)、乱序(shuffle)等操作,方便训练时迭代读取,同时还支持多进程异步读取功能可加快数据读取速度。在飞桨框架中可使用 [paddle.io.DataLoader](../../api/paddle/io/DataLoader_cn.html#dataloader) 迭代读取数据集。
#
#
# 本文以图像数据集为例介绍,文本数据集可参考 [NLP 应用实践](../../practices/nlp/index_cn.html)。
# ## 一、定义数据集
#
# ### 1.1 直接加载内置数据集
#
# 飞桨框架在 [paddle.vision.datasets](../../api/paddle/vision/Overview_cn.html#api) 和 [paddle.text](../..//api/paddle/text/Overview_cn.html#api) 目录下内置了一些经典数据集可直接调用,通过以下代码可查看飞桨框架中的内置数据集。
import paddle
print('计算机视觉(CV)相关数据集:', paddle.vision.datasets.__all__)
print('自然语言处理(NLP)相关数据集:', paddle.text.__all__)
# 从打印结果可以看到飞桨内置了 CV 领域的 MNIST、FashionMNIST、Flowers、Cifar10、Cifar100、VOC2012 数据集,以及 NLP 领域的 Conll05st、Imdb、Imikolov、Movielens、UCIHousing、WMT14、WMT16 数据集。
#
#
# 以 [MNIST](../../api/paddle/vision/datasets/MNIST_cn.html) 数据集为例,加载内置数据集的代码示例如下所示。
# +
from paddle.vision.transforms import Normalize
# 定义图像归一化处理方法,这里的CHW指图像格式需为 [C通道数,H图像高度,W图像宽度]
transform = Normalize(mean=[127.5], std=[127.5], data_format='CHW')
# 下载数据集并初始化 DataSet
train_dataset = paddle.vision.datasets.MNIST(mode='train', transform=transform)
test_dataset = paddle.vision.datasets.MNIST(mode='test', transform=transform)
print('train images: ',len(train_dataset),', test images: ',len(test_dataset))
# -
# 内置的 [MNIST](../../api/paddle/vision/datasets/MNIST_cn.html) 数据集已经划分好了训练集和测试集,通过 `mode` 字段传入 `'train'` 或 `'test'` 来区分。
#
# 另外可通过 `transform` 字段传入一些对图像进行变换的操作,飞桨在 [paddle.vision.transforms](../..api/paddle/vision/Overview_cn.html#about-transforms) 下提供了一些常用的图像变换操作,如对图像进行中心裁剪、水平翻转图像和对图像进行归一化等。这里在初始化 MNIST 数据集时传入了 `Normalize` 变换对图像进行归一化,对图像进行归一化可以加快模型训练的收敛速度。
# 完成数据集初始化之后,可以使用下面的代码直接对数据集进行迭代读取。
# +
from matplotlib import pyplot as plt
for data in train_dataset:
image, label = data
print('shape of image: ',image.shape)
plt.title(str(label))
plt.imshow(image[0])
break
# -
# ### 1.2 使用 paddle.io.Dataset 自定义数据集
# 在实际的场景中,一般需要使用自有的数据来定义数据集,这时可以通过 [paddle.io.Dataset](../../api/paddle/io/Dataset_cn.html#dataset) 基类来实现自定义数据集。
#
# 可构建一个子类继承自 `paddle.io.Dataset` ,并且实现下面的三个函数:
#
# 1. `__init__`:完成数据集初始化操作,将磁盘中的样本文件路径和对应标签映射到一个列表中。
# 2. `__getitem__`:定义指定索引(index)时如何获取样本数据,最终返回对应 index 的单条数据(样本数据、对应的标签)。
# 3. `__len__`:返回数据集的样本总数。
#
# 下面介绍下载 MNIST 原始数据集文件后,用 `paddle.io.Dataset` 定义数据集的代码示例。
#
#
# 下载原始的 MNIST 数据集并解压
# ! wget https://paddle-imagenet-models-name.bj.bcebos.com/data/mnist.tar
# ! tar -xf mnist.tar
# +
import os
import cv2
import numpy as np
from paddle.io import Dataset
class MyDataset(Dataset):
"""
步骤一:继承 paddle.io.Dataset 类
"""
def __init__(self, data_dir, label_path, transform=None):
"""
步骤二:实现 __init__ 函数,初始化数据集,将样本和标签映射到列表中
"""
super(MyDataset, self).__init__()
self.data_list = []
with open(label_path,encoding='utf-8') as f:
for line in f.readlines():
image_path, label = line.strip().split('\t')
image_path = os.path.join(data_dir, image_path)
self.data_list.append([image_path, label])
# 传入定义好的数据处理方法,作为自定义数据集类的一个属性
self.transform = transform
def __getitem__(self, index):
"""
步骤三:实现 __getitem__ 函数,定义指定 index 时如何获取数据,并返回单条数据(样本数据、对应的标签)
"""
# 根据索引,从列表中取出一个图像
image_path, label = self.data_list[index]
# 读取灰度图
image = cv2.imread(image_path, cv2.IMREAD_GRAYSCALE)
# 飞桨训练时内部数据格式默认为float32,将图像数据格式转换为 float32
image = image.astype('float32')
# 应用数据处理方法到图像上
if self.transform is not None:
image = self.transform(image)
# CrossEntropyLoss要求label格式为int,将Label格式转换为 int
label = int(label)
# 返回图像和对应标签
return image, label
def __len__(self):
"""
步骤四:实现 __len__ 函数,返回数据集的样本总数
"""
return len(self.data_list)
# 定义图像归一化处理方法,这里的CHW指图像格式需为 [C通道数,H图像高度,W图像宽度]
transform = Normalize(mean=[127.5], std=[127.5], data_format='CHW')
# 打印数据集样本数
train_custom_dataset = MyDataset('mnist/train','mnist/train/label.txt', transform)
test_custom_dataset = MyDataset('mnist/val','mnist/val/label.txt', transform)
print('train_custom_dataset images: ',len(train_custom_dataset), 'test_custom_dataset images: ',len(test_custom_dataset))
# -
# 在上面的代码中,自定义了一个数据集类 `MyDataset`,`MyDataset` 继承自 `paddle.io.Dataset` 基类 ,并且实现了 `__init__`,`__getitem__` 和 `__len__` 三个函数。
# * 在 `__init__` 函数中完成了对标签文件的读取和解析,并将所有的图像路径 `image_path` 和对应的标签 `label` 存放到一个列表 `data_list` 中。
# * 在 `__getitem__` 函数中定义了指定 index 获取对应图像数据的方法,完成了图像的读取、预处理和图像标签格式的转换,最终返回图像和对应标签 `image, label`。
# * 在 `__len__` 函数中返回 `__init__` 函数中初始化好的数据集列表 `data_list` 长度。
#
#
#
#
# 另外,在 `__init__` 函数和 `__getitem__` 函数中还可实现一些数据预处理操作,如对图像的翻转、裁剪、归一化等操作,最终返回处理好的单条数据(样本数据、对应的标签),该操作可增加图像数据多样性,对增强模型的泛化能力带来帮助。飞桨框架在 [paddle.vision.transforms](../..api/paddle/vision/Overview_cn.html#about-transforms) 下内置了几十种图像数据处理方法,详细使用方法可参考 [数据预处理](03_data_preprocessing_cn.html) 章节。
#
# 和内置数据集类似,可以使用下面的代码直接对自定义数据集进行迭代读取。
for data in train_custom_dataset:
image, label = data
print('shape of image: ',image.shape)
plt.title(str(label))
plt.imshow(image[0])
break
# ## 二、迭代读取数据集
#
# ### 2.1 使用 paddle.io.DataLoader 定义数据读取器
#
# 通过前面介绍的直接迭代读取 Dataset 的方式虽然可实现对数据集的访问,但是这种访问方式只能单线程进行并且还需要手动分批次(batch)。在飞桨框架中,推荐使用 [paddle.io.DataLoader](../../api/paddle/io/DataLoader_cn.html#dataloader) API 对数据集进行多进程的读取,并且可自动完成划分 batch 的工作。
# +
# 定义并初始化数据读取器
train_loader = paddle.io.DataLoader(train_custom_dataset, batch_size=64, shuffle=True, num_workers=1, drop_last=True)
# 调用 DataLoader 迭代读取数据
for batch_id, data in enumerate(train_loader()):
images, labels = data
print("batch_id: {}, 训练数据shape: {}, 标签数据shape: {}".format(batch_id, images.shape, labels.shape))
break
# -
# 通过上述方法,初始化了一个数据读取器 `train_loader`,用于加载训练数据集 `custom_dataset`。在数据读取器中几个常用的字段如下:
#
# * `batch_size`:**每批次读取样本数**,示例中 `batch_size=64` 表示每批次读取 64 个样本。
# * `shuffle`:**样本乱序**,示例中 `shuffle=True` 表示在取数据时打乱样本顺序,以减少过拟合发生的可能。
# * `drop_last`:**丢弃不完整的批次样本**,示例中 `drop_last=True` 表示丢弃因数据集样本数不能被 batch_size 整除而产生的最后一个不完整的 batch 样本。
# * `num_workers`:**同步/异步读取数据**,通过 `num_workers` 来设置加载数据的子进程个数,num_workers的值设为大于0时,即开启多进程方式异步加载数据,可提升数据读取速度。
#
#
# 定义好数据读取器之后,便可用 for 循环方便地迭代读取批次数据,用于模型训练了。值得注意的是,如果使用高层 API 的 [paddle.Model.fit](../../api/paddle/Model_cn.html#fit-train-data-none-eval-data-none-batch-size-1-epochs-1-eval-freq-1-log-freq-10-save-dir-none-save-freq-1-verbose-2-drop-last-false-shuffle-true-num-workers-0-callbacks-none) 读取数据集进行训练,则只需定义数据集 Dataset 即可,不需要再单独定义 DataLoader,因为 paddle.Model.fit 中实际已经封装了一部分 DataLoader 的功能,详细可参考 [模型训练、评估与推理](05_train_eval_predict_cn.html) 章节。
#
#
#
# > 注:
# > DataLoader 实际上是通过批采样器 BatchSampler 产生的批次索引列表,并根据索引取得 Dataset 中的对应样本数据,以实现批次数据的加载。DataLoader 中定义了采样的批次大小、顺序等信息,对应字段包括 `batch_size`、`shuffle`、`drop_last`。这三个字段也可以用一个 `batch_sampler` 字段代替,并在 `batch_sampler` 中传入自定义的批采样器实例。以上两种方式二选一即可,可实现相同的效果。下面小节中介绍后一种自定义采样器的使用方法,该用法可以更灵活地定义采样规则。
#
# ### 2.2 (可选)自定义采样器
#
# 采样器定义了从数据集中的采样行为,如顺序采样、批次采样、随机采样、分布式采样等。采样器会根据设定的采样规则,返回数据集中的索引列表,然后数据读取器 Dataloader 即可根据索引列表从数据集中取出对应的样本。
#
# 飞桨框架在 [paddle.io](../../api/paddle/io/Overview_cn.html) 目录下提供了多种采样器,如批采样器 [BatchSampler](../../api/paddle/io/BatchSampler_cn.html)、分布式批采样器 [DistributedBatchSampler](../../api/paddle/io/DistributedBatchSampler_cn.html)、顺序采样器 [SequenceSampler](../../api/paddle/io/SequenceSampler_cn.html)、随机采样器 [RandomSampler](../../api/paddle/io/RandomSampler_cn.html) 等。
#
#
# 下面通过两段示例代码,介绍采样器的用法。
#
# 首先,以 BatchSampler 为例,介绍在 DataLoader 中使用 BatchSampler 获取采样数据的方法。
#
# +
from paddle.io import BatchSampler
# 定义一个批采样器,并设置采样的数据集源、采样批大小、是否乱序等
bs = BatchSampler(train_custom_dataset, batch_size=8, shuffle=True, drop_last=True)
print("BatchSampler 每轮迭代返回一个索引列表")
for batch_indices in bs:
print(batch_indices)
break
# 在 DataLoader 中使用 BatchSampler 获取采样数据
train_loader = paddle.io.DataLoader(train_custom_dataset, batch_sampler=bs, num_workers=1)
print("在 DataLoader 中使用 BatchSampler,返回索引对应的一组样本和标签数据 ")
for batch_id, data in enumerate(train_loader()):
images, labels = data
print("batch_id: {}, 训练数据shape: {}, 标签数据shape: {}".format(batch_id, images.shape, labels.shape))
break
# -
# 以上示例代码中,定义了一个批采样器实例 `bs`,每轮迭代会返回一个 `batch_size` 大小的索引列表(示例中一轮迭代返回 8 个索引值),数据读取器 `train_loader` 通过 `batch_sampler=bs` 字段传入批采样器,即可根据这些索引获取对应的一组样本数据。另外可以看到,`batch_size`、`shuffle`、`drop_last`这三个参数只在 BatchSampler 中设定。
#
#
# 下面再通过一段代码示例,对比几个不同采样器的采样行为。
# +
from paddle.io import SequenceSampler, RandomSampler, BatchSampler, DistributedBatchSampler
class RandomDataset(paddle.io.Dataset):
def __init__(self, num_samples):
self.num_samples = num_samples
def __getitem__(self, idx):
image = np.random.random([784]).astype('float32')
label = np.random.randint(0, 9, (1, )).astype('int64')
return image, label
def __len__(self):
return self.num_samples
train_dataset = RandomDataset(100)
print('-----------------顺序采样----------------')
sampler = SequenceSampler(train_dataset)
batch_sampler = BatchSampler(sampler=sampler, batch_size=10)
for index in batch_sampler:
print(index)
print('-----------------随机采样----------------')
sampler = RandomSampler(train_dataset)
batch_sampler = BatchSampler(sampler=sampler, batch_size=10)
for index in batch_sampler:
print(index)
print('-----------------分布式采样----------------')
batch_sampler = DistributedBatchSampler(train_dataset, num_replicas=2, batch_size=10)
for index in batch_sampler:
print(index)
# -
# 从代码输出结果可以看出:
# * 顺序采样:按照顺序的方式输出各个样本的索引。
# * 随机采样:先将样本顺序打乱,再输出乱序后的样本索引。
# * 分布式采样:常用于分布式训练场景,将样本数据切分成多份,分别放到不同卡上训练。示例中设置了 `num_replicas=2`,样本会被划分到两张卡上,所以这里只输出一半样本的索引。
# ## 三、总结
#
# 本节中介绍了在飞桨框架中将数据送入模型训练之前的处理流程,总结整个流程和用到的关键 API 如下图所示。
#
# 
#
# 图 1:数据集定义和加载流程
#
# 主要包括定义数据集和定义数据读取器两个步骤,另外在数据读取器中可调用采样器实现更灵活地采样。其中,在定义数据集时,本节仅对数据集进行了归一化处理,如需了解更多数据增强相关操作,可以参考 [数据预处理](03_data_preprocessing_cn.html)。
#
# 以上所有数据处理工作完成后,即可进入下一个任务:[模型训练、评估与推理](05_train_eval_predict_cn.html)。
| docs/guides/02_paddle2.0_develop/02_data_load_cn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 7. Calibration of single ended measurement with WLS and confidence intervals
# A single-ended calibration is performed where the unknown parameters are estimated using fiber sections that have a reference temperature. The parameters are estimated with a weighted least squares optimization using Stokes and anti-Stokes measurements from all timesteps. Thus Stokes and anti-Stokes measurements with a large signal to noise ratio contribute more towards estimating the optimal parameter set. But an estimate of the noise variance is required.
#
# Single-ended calibration requires a few steps. Please have a look at [1] for more information:
# 1. Read the raw data files loaded from your DTS machine
# 2. Define the reference sections: fiber sections that have a known temperature.
# 3. Estimate the variance of the noise in the Stokes and anti-Stokes measurements
# 4. Perform the parameter search and compute the temperature along the entire fiber.
# 5. Compute the confidence intervals for the temperature
#
# [1]: des <NAME>., <NAME>., & <NAME>. (2020). Estimation of Temperature and Associated Uncertainty from Fiber-Optic Raman-Spectrum Distributed Temperature Sensing. Sensors, 20(8), 2235. https://doi.org/10.3390/s20082235
# +
import os
import warnings
warnings.simplefilter('ignore') # Hide warnings to avoid clutter in the notebook
from dtscalibration import read_silixa_files
import matplotlib.pyplot as plt
# %matplotlib inline
# +
filepath = os.path.join('..', '..', 'tests', 'data', 'single_ended')
ds = read_silixa_files(
directory=filepath,
timezone_netcdf='UTC',
file_ext='*.xml')
ds = ds.sel(x=slice(-30, 101)) # only calibrate parts of the fiber
sections = {
'probe1Temperature': [slice(20, 25.5)], # warm bath
'probe2Temperature': [slice(5.5, 15.5)], # cold bath
# 'referenceTemperature': [slice(-24., -4)] # The internal coil is not so uniform
}
ds.sections = sections
# -
print(ds.calibration_single_ended.__doc__)
# First calculate the variance in the measured Stokes and anti-Stokes signals, in the forward and backward direction.
#
# The Stokes and anti-Stokes signals should follow a smooth decaying exponential. This function fits a decaying exponential to each reference section for each time step. The variance of the residuals between the measured Stokes and anti-Stokes signals and the fitted signals is used as an estimate of the variance in measured signals.
st_var, resid = ds.variance_stokes_constant(st_label='st')
ast_var, _ = ds.variance_stokes_constant(st_label='ast')
# Similar to the ols procedure, we make a single function call to calibrate the temperature. If the method is `wls` and confidence intervals are passed to `conf_ints`, confidence intervals calculated. As weigths are correctly passed to the least squares procedure, the covariance matrix can be used. This matrix holds the covariances between all the parameters. A large parameter set is generated from this matrix, assuming the parameter space is normally distributed with their mean at the best estimate of the least squares procedure.
#
# The large parameter set is used to calculate a large set of temperatures. By using `percentiles` or `quantile` the 95% confidence interval of the calibrated temperature between 2.5% and 97.5% are calculated.
#
# The confidence intervals differ per time step. If you would like to calculate confidence intervals of temporal averages or of averages of fiber sections see notebook 16.
ds.calibration_single_ended(sections=sections,
st_var=st_var,
ast_var=ast_var,
method='wls')
ds.conf_int_single_ended(
st_var=st_var,
ast_var=ast_var,
conf_ints=[2.5, 97.5],
mc_sample_size=500)
# Lets compare our calibrated values with the device calibration
ds1 = ds.isel(time=0) # take only the first timestep
ds1.tmpf.plot(linewidth=0.8, figsize=(12, 8), label='User calibrated') # plot the temperature calibrated by us
ds1.tmp.plot(linewidth=0.8, label='Device calibrated') # plot the temperature calibrated by the device
ds1.tmpf_mc.plot(linewidth=0.8, hue='CI', label='CI device')
plt.title('Temperature at the first time step')
plt.legend();
ds.tmpf_mc_var.plot(figsize=(12, 8));
ds1.tmpf_mc.sel(CI=2.5).plot(label = '2.5% CI', figsize=(12, 8))
ds1.tmpf_mc.sel(CI=97.5).plot(label = '97.5% CI')
ds1.tmpf.plot(label='User calibrated')
plt.title('User calibrated temperature with 95% confidence interval')
plt.legend();
# We can tell from the graph above that the 95% confidence interval widens furtherdown the cable. Lets have a look at the calculated variance along the cable for a single timestep. According to the device manufacturer this should be around 0.0059 degC.
ds1.tmpf_mc_var.plot(figsize=(12, 8));
# The variance of the temperature measurement appears to be larger than what the manufacturer reports. This is already the case for the internal cable; it is not caused by a dirty connector/bad splice on our side. Maybe the length of the calibration section was not sufficient.
#
# At 30 m the variance sharply increases. There are several possible explanations. E.g., large temperatures or decreased signal strength.
#
# Lets have a look at the Stokes and anti-Stokes signal.
ds1.st.plot(figsize=(12, 8))
ds1.ast.plot();
# Clearly there was a bad splice at 30 m that resulted in the sharp increase of measurement uncertainty for the cable section after the bad splice.
| examples/notebooks/07Calibrate_single_wls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Imports" data-toc-modified-id="Imports-1"><span class="toc-item-num">1 </span>Imports</a></span><ul class="toc-item"><li><span><a href="#Before-vs-After-changepoint" data-toc-modified-id="Before-vs-After-changepoint-1.1"><span class="toc-item-num">1.1 </span>Before vs After changepoint</a></span></li></ul></li></ul></div>
# -
# # Imports
# +
# Data manipulation
import numpy as np
import pandas as pd
# Plotting
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
# %matplotlib inline
# spikelearn
import os
os.chdir('../../')
import sys
sys.path.append('.')
from spikelearn.data import io, SHORTCUTS
from spikelearn.data.selection import select
from spikelearn.visuals.visuals import raster_multiple, raster_plot
# -
# ## Before vs After changepoint
savedir = 'data/results/duration/d_prime'
filename = lambda label: '{}_Dprime_cp_init.csv'.format(label)
loaddata = lambda label: pd.read_csv('{}/{}'.format(savedir, filename(label)) )
cp_d
res.groupby('trial').max().window_d.unstack('trial').transpose().plot()
res.replace([np.inf, -np.inf], np.nan).dropna().groupby('trial').max().window_d.plot()
plt.figure(figsize=(12,10))
for i, label in enumerate(SHORTCUTS['groups']['DRRD']):
res = loaddata(label).replace([np.inf, -np.inf], np.nan).dropna()
res['after_cp'] = 1-res['before_cp']
cp_d = res.drop_duplicates(['unit','time','after_cp']).groupby(['unit','after_cp']).mean()
ax = plt.subplot2grid((4,2), (i, 0))
cp_d.cp_d.unstack('unit').plot(legend=False, ax=ax,marker='o')
sns.violinplot(x='after_cp', y='cp_d', data=cp_d.reset_index())
ax = plt.subplot2grid( (4,2), (i, 1) )
#res.groupby('trial').max().window_d.plot(ax=ax)
mres = res.groupby(['unit','time']).apply(differ).reset_index().groupby('unit').max()
units = mres.sort_values(0,ascending=False).reset_index().unit.values[:4]
res[res.unit.isin(units)].groupby([ 'before_cp','unit', 'time']).mean().fr.unstack('unit').loc[False].plot(ax=ax)
res[res.unit.isin(units)].groupby([ 'before_cp','unit', 'time']).mean().fr.unstack('unit').loc[True].plot(ax=ax)
def differ(df):
if df[df['before_cp']==True].cp_d.values.shape[0] == 0 or df[df['before_cp']==False].cp_d.values.shape[0] == 0:
return 0
d = df[df['before_cp']==True].cp_d.values[0] - df[df['before_cp']==False].cp_d.values[0]
return d
res = loaddata('DRRD 8')
mres = res.groupby(['unit','time']).apply(differ).reset_index().groupby('unit').max()
units = mres.sort_values(0,ascending=False).reset_index().unit.values[:4]
# +
ax=plt.subplot(1,1,1)
res[res.unit.isin(units)].groupby([ 'before_cp','unit', 'time']).mean().fr.unstack('unit').loc[False].plot(ax=ax)
res[res.unit.isin(units)].groupby([ 'before_cp','unit', 'time']).mean().fr.unstack('unit').loc[True].plot(ax=ax)
# -
cp_d.reset_index().groupby(['unit','time']).diff()
| notebooks/exploration/Dprime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import plotly
plotly.tools.set_credentials_file(username='jamespcollins', api_key='eGJeJujrYubjABbnE2Qo')
# +
import plotly.plotly as py
from plotly.graph_objs import *
trace0 = Scatter(
x=[1, 2, 3, 4],
y=[10, 15, 13, 17]
)
trace1 = Scatter(
x=[1, 2, 3, 4],
y=[16, 5, 11, 9]
)
data = Data([trace0, trace1])
py.plot(data, filename = 'basic-line')
# -
| Notebooks/GW viz with plotly.ipynb |
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .clj
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Clojure (clojupyter=0.3.2=1)
;; language: clojure
;; name: python397jvsc74a57bd089a790bea3b9cadff6bf73491f7cf161e4e94bfc015afe6a535623fbe4142b79
;; ---
;; # Def, Symbols, and Vars
;;
;; ### A Global, Stable Place for Your Stuff
;; +
;; 'def' binds a symbol to a value
(def title "Emma")
;; +
;; 'def' is useful to store constants
(def PI 3.14) ; Everyone's favorite universal constant.
(def ISBN-LENGTH 13) ; Length of a standard book ID.
(def COMPANY-NAME "<NAME>") ; Company names are more or less constant.
;; +
;; 'defn' is just 'def' + 'fn'
(defn book-description [book]
(str (:title book)
" Written by "
(:author book)))
;; -
(def book-description
(fn [book]
(str (:title book)
" Written by "
(:author book))))
;; +
;; Bindings from a given 'def' can be used in other bindings
(def ISBN-LENGTH 13) ; Length of a standard book ID.
(def OLD-ISBN-LENGTH 10) ; Before 2007 ISBNs were 10 characters long.
(def isbn-lengths [OLD-ISBN-LENGTH ISBN-LENGTH])
;; +
;; Or inside functions
(defn valid-isbn [isbn]
(or (= (count isbn) OLD-ISBN-LENGTH)
(= (count isbn) ISBN-LENGTH)))
;; -
;; ### Symbols are things
;; +
;; This expression involves 2 values: the string "Austen" and the symbol author
(def author "Austen")
;; +
;; THe single quote prevents the symbol from being evaluated
'author
;; +
;; Symbols can be manipulated as any other value
(str 'author) ; Symbols can be converted to strings
;; -
(= 'title 'title) ;Symbols can be used inside expressions
;; ### Bindings Are Things Too
;; +
;; When invoking 'def', a 'var' is created
(def author "Austen")
;; +
;; A 'var' can be accessed (without evaluating it) as follows
#'author
;; +
;; Vars can be used inside another vars like any other value
(def the-var #'author)
;; +
;; Get the value of the var
(.get the-var) ; Using java methods
;; +
;; Get the symbol of the var
(.-sym the-var) ; Using java methods
;; -
;; ### Varying Your Vars
;; +
;; Given the following var and function
(def PI 3.14)
(defn compute-area [diameter]
(* PI diameter diameter))
(compute-area 4)
;; +
;; Although not recommended in production, they can me modified
;; (bounded to another set of values) as follows
(def PI 3.14159)
(defn compute-area [diameter]
(let [radius (/ diameter 2)]
(* PI radius radius)))
(compute-area 4)
;; +
;; Vars intended to be changed at some point of time
;; are defined with '^dynamic' metadata, as follows
(def ^:dynamic *debug-enabled* false) ; By convention, dynamic vars are enclosed in asterisks
(defn debug [msg]
(if *debug-enabled* (println msg)))
;; +
;; The last var can be dynamically bounded to another value
;; with 'binding', as follows
(defn some-troublesome-function-that-needs-logging []
(println "The darned function"))
(binding [*debug-enabled* true]
(debug "Calling that darned function")
(some-troublesome-function-that-needs-logging)
(debug "Back from that darned function"))
;; -
;; ### Issues with vars
;;
;; +
;; Trying to create vars with 'let' raises an exception
(let [let-bound 42] #'let-bound)
;; -
;; ### Some clojure built-in vars
;;
;; +
;; *print-length* let you print a specific number of elements of a collection
(def books ["Emma" "2001" "Jaws" "<NAME>"])
(set! *print-length* 2)
books
;; +
;; *1, *2, *3 let you print the last, second last and third last element
;; of the REPL, respectively
(+ 2 2)
*1
;; -
"Austen"
"King"
"Orwell"
*3
;; +
;; *e let you print the last exception
(/ 1 0)
;; -
*e
| chapter08/chapter08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kT42ekZXCgVj"
# ### Python if Statement Syntax
# + id="XU5OenBXCpla"
# If the number is positive, we print an appropriate message
# Python if... Statement - 1 option
num = 30
if num > 0:
print(num, "is a positive number.")
print("printed number status.")
# Python if...else Statement - 2 option
if num >= 0:
print("Positive or Zero")
else:
print("Negative number")
# Python if...elif...else Statement - 3 Option
if num > 0:
print("Positive number")
elif num == 0:
print("Zero")
else:
print("Negative number")
# + [markdown] id="NbR8TUc7CtlS"
# ### * Python Nested if statements
#
# If if...elif...else statement is used inside another if...elif...else statement, then it is called nesting in computer programming.
# + id="Db9MJdWECzbM"
num = float(input("Enter a number: "))
if num >= 0:
if num == 0:
print("Zero")
else:
print("Positive number")
else:
print("Negative number")
# + [markdown] id="xfz65qHKC5Yd"
# ## Python for Loop:
# + id="9jHH2_ZMDADA"
# Program to find the sum of all numbers stored in a list
# List of numbers
numbers = [60, 25, 33, 85, 40, 22, 55, 43, 51]
# variable to store the sum
sum = 0
# iterate over the list
for val in numbers:
sum = sum+val
print("The sum is", sum)
# + [markdown] id="-RIkKIP8DIFp"
# ## The range() function
# * We can generate a sequence of numbers using range() function. range(10) will generate numbers from 0 to 9 (10 numbers).
# * We can also define the start, stop and step size as range(start, stop,step_size). step_size defaults to 1 if not provided.
# * This function does not store all the values in memory; it would be inefficient. So it remembers the start, stop, step size and generates the next number on the go.
# * To force this function to output all the items, we can use the function list().
# + id="fDuWxeaeDQhX"
print(range(10))
print(list(range(15)))
print(list(range(2, 10)))
print(list(range(2, 30, 3)))
# + id="AfstgzBWDVaw"
# Program to iterate through a list using indexing
music = ['pop', 'rock', 'jazz','classic']
# iterate over the list using index
for i in range(len(music)):
print("I like", music[i])
#combined with the len() function to iterate through a sequence using indexing.
# + [markdown] id="hsK_PSdnDY1C"
# ## for loop with else
# + id="JKI1oudBDePH"
# program to display student's marks from record
student_name = 'Soyuj'
marks = {'James': 90, 'Jules': 55, 'Arthur': 77}
for student in marks:
if student == student_name:
print(marks[student])
break
else:
print('No entry with that name found.')
# Use of break statement inside the loop
for val in "string":
if val == "i":
break
print(val)
print("The end")
# Program to show the use of continue statement inside loops
for val in "string":
if val == "i":
continue
print(val)
print("The end")
# + [markdown] id="omMqFa4LDho1"
# ## while loop in Python
# * The while loop in Python is used to iterate over a block of code as long as the test expression (condition) is true.
# + id="A2qPDnt2DrZo"
# To take input from the user,
# n = int(input("Enter n: "))
n = 10
# initialize sum and counter
sum = 0
i = 1
while i <= n:
sum = sum + i
i = i+1 # update counter
# print the sum
print("The sum is", sum)
# While loop with else
'''Example to illustrate
the use of else statement
with the while loop'''
counter = 0
while counter < 3:
print("Inside loop")
counter = counter + 1
else:
print("Inside else")
| OOP/Practice Sessions/Python_Control_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Correlated Graph Pairs
import graspologic
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# The correlated Graph Pairs Model means that we generate a graph pair with some correlation. We define the probability of observing the same edge in both graphs by adjusting the correlation values and sampling the graph pairs.
# ## ER correlation graph pair
# There are different ways to define the probability matrix of the first graph. The ER model is a simple example. We assume that each edge for all pairs of vertices is sampled independently from each other edge.
#
# ER_CORR(n, p, r) - this model specifies the number of vertices, $n$, the probability of an edge existing between a given pair of vertices, $p$, and the correlation value between the two, $r$.
#
# Below, we sample a graph pair with the ER model (undirected and no self-loops) $G_1, G_2 \sim ER\_{CORR}\,(50, 0.5, 0.3)$.
from graspologic.simulations import er_corr
# +
n = 50
r = 0.3
p = 0.5
np.random.seed(2)
G1, G2 = er_corr(n, p, r, directed = False, loops = False)
# -
# ## Visualize the graphs using heatmap
from graspologic.plot import heatmap
heatmap(G1, title = 'Correlated ER Simulation Graph 1')
heatmap(G2, title = 'Correlated ER Simulation Graph 2')
# ## SBM correlated graph pair
# Stochastic block models tend to produce graphs containing communities: subsets characterized by being connected with one another with particular edge densities. We can define the partitioned matrix with different probabilities of having edges and the correlation values between the two.
#
# Below, we sample a two-block SBM graph pair (undirected and no self-loops) G1 and G2 with the following parameters:
#
# \begin{align*}
# n &= [20, 30]\\
# p &= \begin{bmatrix}
# 0.5 & 0.2\\
# 0.2 & 0.5
# \end{bmatrix}\\
# r &= 0.3
# \end{align*}
from graspologic.simulations import sbm_corr
# +
np.random.seed(3)
n = [20, 30]
p = [[0.5, 0.2], [0.2, 0.5]]
r = 0.3
G1, G2 = sbm_corr(n, p, r, directed=False, loops=False)
# -
# ## Visualize the graphs using heatmap
heatmap(G1, title = 'Correlated SBM Simulation Graph 1')
heatmap(G2, title = 'Correlated SBM Simulation Graph 2')
| docs/tutorials/simulations/corr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="rQc-wXjqrEuR"
# # Accelerate Inference of NLP models with OpenVINO Post-Training Optimization Tool
# This tutorial demostrates how to apply INT8 quantization to the Natural Language Processing model BERT, using the [Post-Training Optimization Tool API](https://docs.openvinotoolkit.org/latest/pot_compression_api_README.html) (part of the [OpenVINO Toolkit](https://docs.openvinotoolkit.org/)). We will use a fine-tuned [HuggingFace BERT](https://huggingface.co/transformers/model_doc/bert.html) [PyTorch](https://pytorch.org/) model trained for [Microsoft Research Paraphrase Corpus (MRPC)](https://www.microsoft.com/en-us/download/details.aspx?id=52398) task. The code of the tutorial is designed to be extendable to custom models and datasets. It consists of the following steps:
#
# - Download and prepare the MRPC model and dataset
# - Define data loading and accuracy validation functionality
# - Prepare the model for quantization
# - Run optimization pipeline
# - Compare performance of the original and quantized models
# +
import os
import sys
import time
import warnings
from pathlib import Path
import numpy as np
import torch
from addict import Dict
from compression.api import DataLoader as POTDataLoader
from compression.api import Metric
from compression.engines.ie_engine import IEEngine
from compression.graph import load_model, save_model
from compression.graph.model_utils import compress_model_weights
from compression.pipeline.initializer import create_pipeline
from compression.utils.logger import get_logger, init_logger
from torch.utils.data import TensorDataset
from transformers import BertForSequenceClassification, BertTokenizer
from transformers import glue_convert_examples_to_features as convert_examples_to_features
from transformers import glue_output_modes as output_modes
from transformers import glue_processors as processors
# +
DATA_DIR = "data"
MODEL_DIR = "model"
os.makedirs(DATA_DIR, exist_ok=True)
os.makedirs(MODEL_DIR, exist_ok=True)
# + [markdown] id="YytHDzLE0uOJ" pycharm={"name": "#%% md\n"}
# ## Prepare the Model
# Next steps include:
# - Download and unpack pre-trained BERT model for MRPC by PyTorch
# - Convert model to ONNX
# - Run OpenVINO Model Optimizer tool to convert the model from the ONNX representation to the OpenVINO Intermediate Representation (IR)
# + id="f7i6dWUmhloy"
# !curl https://download.pytorch.org/tutorial/MRPC.zip --output $MODEL_DIR/MRPC.zip
# !unzip -n $MODEL_DIR/MRPC.zip -d $MODEL_DIR
# + [markdown] id="ehX7F6KB0uPu"
# Import all dependencies to load the original PyTorch model and convert it to the ONNX representation.
# + id="r5as0_Yg0uQX" pycharm={"name": "#%%\n"}
BATCH_SIZE = 1
MAX_SEQ_LENGTH = 128
def export_model_to_onnx(model, path):
with torch.no_grad():
default_input = torch.ones(1, MAX_SEQ_LENGTH, dtype=torch.int64)
inputs = {
"input_ids": default_input,
"attention_mask": default_input,
"token_type_ids": default_input,
}
outputs = model(**inputs)
symbolic_names = {0: "batch_size", 1: "max_seq_len"}
torch.onnx.export(
model,
(inputs["input_ids"], inputs["attention_mask"], inputs["token_type_ids"]),
path,
opset_version=11,
do_constant_folding=True,
input_names=["input_ids", "input_mask", "segment_ids"],
output_names=["output"],
dynamic_axes={
"input_ids": symbolic_names,
"input_mask": symbolic_names,
"segment_ids": symbolic_names,
},
)
print("ONNX model saved to {}".format(path))
torch_model = BertForSequenceClassification.from_pretrained(os.path.join(MODEL_DIR, "MRPC"))
onnx_model_path = Path(MODEL_DIR) / "bert_mrpc.onnx"
if not onnx_model_path.exists():
export_model_to_onnx(torch_model, onnx_model_path)
# + [markdown] id="sNWDAGGd0uRt"
# Then convert the ONNX model using OpenVINO Model Optimizer with the required parameters.
# + id="-6P0c_960uR5" pycharm={"name": "#%%\n"}
ir_model_xml = onnx_model_path.with_suffix(".xml")
ir_model_bin = onnx_model_path.with_suffix(".bin")
if not ir_model_xml.exists():
# !mo --input_model $onnx_model_path --output_dir $MODEL_DIR --model_name bert_mrpc --input input_ids,input_mask,segment_ids --input_shape [1,128],[1,128],[1,128] --output output --data_type FP16
# + [markdown] id="LBbY7c4NsHzT"
# ## Prepare MRPC Task Dataset
#
# To run this tutorial, you will need to download the GLUE data part for MRPC task from HuggingFace.
# The code below will donwload a script that fetches MRPC dataset.
# + id="NN-qRME1a-Sm"
# !curl https://raw.githubusercontent.com/huggingface/transformers/f98ef14d161d7bcdc9808b5ec399981481411cc1/utils/download_glue_data.py --output download_glue_data.py
# +
from download_glue_data import format_mrpc
os.makedirs(DATA_DIR, exist_ok=True)
format_mrpc(DATA_DIR, "")
# + [markdown] id="E5hsOsj-0uSc"
# ## Define DataLoader for POT
# In this step, we need to define `DataLoader` based on POT API. It will be used to collect statistics for quantization and run model evaluation. We use helper functions from the HuggingFace Transformers to do the data preprocessing. It takes raw text data and encodes sentences and words producing three model inputs. For more details about the data preprocessing and tokenization please refer to this [description](https://medium.com/@dhartidhami/understanding-bert-word-embeddings-7dc4d2ea54ca).
# + id="6xnl2PhM0uSn"
class MRPCDataLoader(POTDataLoader):
# Required methods
def __init__(self, config):
"""Constructor
:param config: data loader specific config
"""
if not isinstance(config, Dict):
config = Dict(config)
super().__init__(config)
self._task = config["task"].lower()
self._model_dir = config["model_dir"]
self._data_dir = config["data_source"]
self._batch_size = config["batch_size"]
self._max_length = config["max_length"]
self._prepare_dataset()
def __len__(self):
"""Returns size of the dataset"""
return len(self.dataset)
def __getitem__(self, index):
"""
Returns annotation, data and metadata at the specified index.
Possible formats:
(index, annotation), data
(index, annotation), data, metadata
"""
if index >= len(self):
raise IndexError
batch = self.dataset[index]
batch = tuple(t.detach().cpu().numpy() for t in batch)
inputs = {"input_ids": batch[0], "input_mask": batch[1], "segment_ids": batch[2]}
labels = batch[3]
return (index, labels), inputs
# Methods specific to the current implementation
def _prepare_dataset(self):
"""Prepare dataset"""
tokenizer = BertTokenizer.from_pretrained(self._model_dir, do_lower_case=True)
processor = processors[self._task]()
output_mode = output_modes[self._task]
label_list = processor.get_labels()
examples = processor.get_dev_examples(self._data_dir)
features = convert_examples_to_features(
examples,
tokenizer,
label_list=label_list,
max_length=self._max_length,
output_mode=output_mode,
)
all_input_ids = torch.tensor([f.input_ids for f in features], dtype=torch.long)
all_attention_mask = torch.tensor([f.attention_mask for f in features], dtype=torch.long)
all_token_type_ids = torch.tensor([f.token_type_ids for f in features], dtype=torch.long)
all_labels = torch.tensor([f.label for f in features], dtype=torch.long)
self.dataset = TensorDataset(
all_input_ids, all_attention_mask, all_token_type_ids, all_labels
)
# + [markdown] id="Re9-YhbBddh3"
# ## Define Accuracy Metric Calculation
# At this step the `Metric` interface for MRPC task metrics is implemented. It is used for validating accuracy of models.
# + id="GB8L492ztZEC"
class Accuracy(Metric):
# Required methods
def __init__(self):
super().__init__()
self._name = "Accuracy"
self._matches = []
@property
def value(self):
"""Returns accuracy metric value for the last model output."""
return {self._name: self._matches[-1]}
@property
def avg_value(self):
"""Returns accuracy metric value for all model outputs."""
return {self._name: np.ravel(self._matches).mean()}
def update(self, output, target):
"""Updates prediction matches.
:param output: model output
:param target: annotations
"""
if len(output) > 1:
raise Exception(
"The accuracy metric cannot be calculated " "for a model with multiple outputs"
)
output = np.argmax(output)
match = output == target[0]
self._matches.append(match)
def reset(self):
"""Resets collected matches"""
self._matches = []
def get_attributes(self):
"""
Returns a dictionary of metric attributes {metric_name: {attribute_name: value}}.
Required attributes: 'direction': 'higher-better' or 'higher-worse'
'type': metric type
"""
return {self._name: {"direction": "higher-better", "type": "accuracy"}}
# + [markdown] id="CclWk-fVd9Wi"
# ## Run Quantization Pipeline
# Here we define a configuration for our quantization pipeline and run it. Please note that we use built-in `IEEngine` implementation of `Engine` interface from the POT API for model inference.
# + id="PiAvrwo0tr6Z"
warnings.filterwarnings("ignore") # Suppress accuracychecker warnings
model_config = Dict({"model_name": "bert_mrpc", "model": ir_model_xml, "weights": ir_model_bin})
engine_config = Dict({"device": "CPU"})
dataset_config = {
"task": "mrpc",
"data_source": os.path.join(DATA_DIR, "MRPC"),
"model_dir": os.path.join(MODEL_DIR, "MRPC"),
"batch_size": BATCH_SIZE,
"max_length": MAX_SEQ_LENGTH,
}
algorithms = [
{
"name": "DefaultQuantization",
"params": {
"target_device": "ANY",
"model_type": "transformer",
"preset": "performance",
"stat_subset_size": 250,
},
}
]
# Step 1: Load the model.
model = load_model(model_config)
# Step 2: Initialize the data loader.
data_loader = MRPCDataLoader(dataset_config)
# Step 3 (Optional. Required for AccuracyAwareQuantization): Initialize the metric.
metric = Accuracy()
# Step 4: Initialize the engine for metric calculation and statistics collection.
engine = IEEngine(config=engine_config, data_loader=data_loader, metric=metric)
# Step 5: Create a pipeline of compression algorithms.
pipeline = create_pipeline(algorithms, engine)
# Step 6 (Optional): Evaluate the original model. Print the results.
fp_results = pipeline.evaluate(model)
if fp_results:
print("FP16 model results:")
for name, value in fp_results.items():
print(f"{name}: {value:.5f}")
# + id="hPj_fcDAG8xG"
# Step 7: Execute the pipeline.
warnings.filterwarnings("ignore") # Suppress accuracychecker warnings
print(
f"Quantizing model with {algorithms[0]['params']['preset']} preset and {algorithms[0]['name']}"
)
start_time = time.perf_counter()
compressed_model = pipeline.run(model)
end_time = time.perf_counter()
print(f"Quantization finished in {end_time - start_time:.2f} seconds")
# Step 8 (Optional): Compress model weights to quantized precision
# in order to reduce the size of final .bin file.
compress_model_weights(compressed_model)
# Step 9: Save the compressed model to the desired path.
compressed_model_paths = save_model(
compressed_model, save_path=MODEL_DIR, model_name="quantized_bert_mrpc"
)
compressed_model_xml = compressed_model_paths[0]["model"]
# + id="hPj_fcDAG8xG"
# Step 10 (Optional): Evaluate the compressed model. Print the results.
int_results = pipeline.evaluate(compressed_model)
if int_results:
print("INT8 model results:")
for name, value in int_results.items():
print(f"{name}: {value:.5f}")
# + [markdown] id="vQACMfAUo52V" tags=[]
# ## Compare Performance of the Original and Quantized Models
# Finally, we will measure the inference performance of the FP32 and INT8 models. To do this, we use [Benchmark Tool](https://docs.openvinotoolkit.org/latest/openvino_inference_engine_tools_benchmark_tool_README.html) - OpenVINO's inference performance measurement tool.
#
# > NOTE: For more accurate performance, we recommended running `benchmark_app` in a terminal/command prompt after closing other applications. Run `benchmark_app -m model.xml -d CPU` to benchmark async inference on CPU for one minute. Change `CPU` to `GPU` to benchmark on GPU. Run `benchmark_app --help` to see an overview of all command line options.
# +
## compressed_model_xml is defined after quantizing the model.
## Uncomment the lines below to set default values for the model file locations.
# ir_model_xml = "model/bert_mrpc.xml"
# compressed_model_xml = "model/quantized_bert_mrpc.xml"
# -
# Inference FP16 model (IR)
# ! benchmark_app -m $ir_model_xml -d CPU -api async
# Inference INT8 model (IR)
# ! benchmark_app -m $compressed_model_xml -d CPU -api async
| notebooks/105-language-quantize-bert/105-language-quantize-bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import json
# +
from watson_developer_cloud import ToneAnalyzerV3
tone_analyzer = ToneAnalyzerV3(
username='o7VsZkJOk4kw',
password='<PASSWORD>',
version='2016-05-19')
# -
print json.dumps(tone_analyzer.tone(text='A word is dead when it is said, some say. <NAME>'), indent=2)
| files/nb_demo/watson/tone_analyzer-node.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=[]
# #!pip install GitPython
# #!pip install sagemaker
# #!pip install s3fs
# #!pip install aws-mlops
import os
import sys
#sys.path.insert(1, os.getcwd() + '/..')
import pandas as pd
import config as config
from aws_mlops.data_storage import DataStorage
# -
# object for data storage management
studio = DataStorage(config.source_bucket, config.test_data_key)
config_name = 'testing_input'
# load dataframes
[columns_names, target, identifier] = studio.restore_test([config.target, config.identifier])
test = studio.restore(config.test_filename, config.test_path)
score = studio.restore(config.prediction_filename, config.prediction_path)
# join datasets
df = pd.concat([identifier, target, score, test], axis=1)
df.columns = [config.identifier, config.target, config.score] + list(columns_names[0])
# prediction datasets
prediction = df[[config.identifier, config.score]]
# report datasets
report = pd.DataFrame({
'Mean squared error': [metrics.mean_squared_error(df[config.target], df[config.score])],
'Mean absolute error': [metrics.mean_absolute_error(df[config.target], df[config.score])]
})
# output - dataframes: prediction and report
output_dataframes = [prediction, report]
for output in getattr(config, config_name)['outputs']:
dataframe = output_dataframes.pop(0)
ds.local_save(dataframe, output['S3Output']['LocalPath'], output['OutputName'], header = True, index = False)
| example/test_with_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
df = pd.read_csv('taxi.csv')
df.head()
df.describe()
df.sum()
# TO check is the data has null values
df.isnull().sum()
#
x = df.iloc[:,0:-1].values
y = df['Numberofweeklyriders'].values
y
# # Data Visualization
sns.heatmap(df)
# +
plt.subplots(figsize=(12, 10))
plt.hist(df, histtype='bar')
plt.show()
# -
# +
plt.figure(figsize=(12, 12))
plt.hist(df)
# -
sns.boxplot(df)
#spliting the data using train_test_split()
from sklearn.model_selection import train_test_split
x_train, x_test,y_train, y_test = train_test_split(x,y, test_size=0.2)
#importing LinearRegression from sklearn
from sklearn.linear_model import LinearRegression
#creating a model
reg = LinearRegression()
reg.fit(x_train,y_train)
#to cheack the score of model
print('Train score', reg.score(x_train,y_train))
print('Test score', reg.score(x_test,y_test))
#creating a pickle model
pickle.dump(reg,open('taxi.pkl','wb'))
model = pickle.load(open('taxi.pkl','rb'))
model.predict([[80,177000,6000,85]])
| ola_ride.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train_Test_Split
# +
import pandas as pd
import numpy as np
df = pd.read_csv("C:/Users/Student/Desktop/FD2.csv")
# Create arrays for features and target variable
y = df['stars_review']
y = np.asarray(y, dtype="|S6")
X = df.drop(['stars_review'], axis=1)
# -
type(y)
print(df.info())
print(y)
print(X)
# +
# Train, Test, Validation Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.3, random_state=1)
# -
# # LDA Processing
# +
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=2)
X_train_lda = lda.fit_transform(X_train, y_train)
# +
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr = lr.fit(X_train_lda, y_train)
# -
X_test_lda = lda.transform(X_test)
X_train_lda = pd.DataFrame(X_train_lda)
X_train_lda.to_csv("C:/Users/Student/Desktop/X_train_lda.csv", index=False, header=True)
import matplotlib.pyplot as plt
plt.figure(figsize = (10, 10))
plt.scatter(X_train_lda.iloc[:, 0], X_train_lda.iloc[:, 1],marker='o',c=y_train)
plt.show()
| Kaggle/Yelp_2013/DataPreprocessing/LDA.ipynb |
# ---
# jupyter:
# jupytext:
# notebook_metadata_filter: language_info
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# language_info:
# codemirror_mode:
# name: ipython
# version: 3
# file_extension: .py
# mimetype: text/x-python
# name: python
# nbconvert_exporter: python
# pygments_lexer: ipython3
# version: 3.4.5
# ---
# # Brexit and ages
#
# Now we have [for loops](../iteration/iteration) and
# [ranges](../arrays/Ranges), we can solve the problem in
# [population, permutation](population_permutation).
# +
# Array library.
import numpy as np
# Data frame library.
import pandas as pd
# Safe setting for Pandas.
pd.set_option('mode.chained_assignment', 'raise')
# Plotting
import matplotlib.pyplot as plt
# %matplotlib inline
# Fancy plots
plt.style.use('fivethirtyeight')
# -
# We load the Brexit survey data again:
# Load the data frame, and put it in the variable "audit_data"
audit_data = pd.read_csv('audit_of_political_engagement_14_2017.tab', sep='\t')
# Again, we get the ages for the Leavers and the Remainers:
# Drop rows where age is 0
age_not_0 = audit_data['numage'] != 0
good_data = audit_data[age_not_0]
# Get data frames for leavers and remainers
is_remain = good_data['cut15'] == 1
remain_ages = good_data[is_remain]['numage']
is_leave = good_data['cut15'] == 2
leave_ages = good_data[is_leave]['numage']
remain_ages.hist();
leave_ages.hist();
# Here is the number of Remain voters:
n_remain = len(remain_ages)
n_remain
# Here was the actual difference between the means of the two groups:
actual_diff = np.mean(leave_ages) - np.mean(remain_ages)
actual_diff
# We want to know if we have a reasonable chance of seeing a difference of this magnitude, if the two groups are samples from the same underlying population. We don't have the actual population to take samples from, so we need to wing it, by using the data we have.
#
# We asserted we could use permutation to take random samples from the data that we already have:
pooled = np.append(remain_ages, leave_ages)
shuffled = np.random.permutation(pooled)
fake_remainers = shuffled[:n_remain]
fake_leavers = shuffled[n_remain:]
# Those are our samples. Now we get the difference in mean ages, as one example of a difference we might see, if the samples are from the same population:
example_diff = np.mean(fake_leavers) - np.mean(fake_remainers)
example_diff
# Now we know how do to this once, we can use the `for` loop to do the
# permutation operation many times. We collect the results in an array. You will
# recognize the code in the `for` loop from the code in the cells above.
# An array of zeros to store the fake differences
example_diffs = np.zeros(10000)
# Do the shuffle / difference steps 10000 times
for i in np.arange(10000):
shuffled = np.random.permutation(pooled)
fake_remainers = shuffled[:n_remain]
fake_leavers = shuffled[n_remain:]
eg_diff = np.mean(fake_leavers) - np.mean(fake_remainers)
# Collect the results in the results array
example_diffs[i] = eg_diff
# Our results array now has 10000 fake mean differences:
#
# What distribution do these differences have?
plt.hist(example_diffs);
# This is called the *sampling distribution*. In our case, this is the
# sampling distribution of the difference in means. It is the
# *sampling* distribution, because it is the distribution we expect to
# see, when taking random *samples* from the same underlying population.
#
# Our question now is, is the difference we actually saw, a likely value, given the sampling distribution. Let's plot the actual difference, so we can see how similar/different it is to the simulated differences.
# do not worry about the code below, it just plots the sampling distribution, the actual difference in the mean ages,
# and adds some labels to the histogram.
plt.hist(example_diffs, label = 'simulated differences')
fontsize = {'fontsize': 10}
plt.plot(actual_diff, 20 , 'o', markersize = 10,color = 'red', label = 'actual difference')
plt.xlabel('Difference between the mean ages of leavers and remainers', **fontsize)
plt.ylabel('Number of times obtained in simulation', **fontsize)
plt.legend(**fontsize);
# Looking at the distribution above - what do you think?
#
# The blue histogram shows the distribution of differences we would expect to obtain in an ideal world. That is, in a world where there was no difference between the mean age of leavers and remainers. The red dot shows the actual difference between the mean ages of leavers and remainers. Does it look likely that we would obtain the actual difference in the ideal world?
#
# As a first pass, let us check how many of the values from the sampling
# distribution are as large, or larger than the value we actually saw.
are_as_high = example_diffs >= actual_diff
n_as_high = np.count_nonzero(are_as_high)
n_as_high
# The number above is the number of values in the sampling distribution
# that are as high as, or higher than, the value we actually saw. If we
# divide by 10000, we get the proportion of the sampling distribution
# that is as high, or higher.
proportion = n_as_high / 10000
proportion
# We think of this proportion as an estimate of the *probability* that
# we would see a value this high, or higher, *if these were random
# samples from the same underlying population*. We call this a *p
# value*.
| permutation/brexit_ages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import torch
import numpy as np
import k3d
# from libs import tpcpath
from torchpcp.utils import pytorch_tools
pytorch_tools.set_seed(0)
device = pytorch_tools.select_device("cuda")
# -
def visualize_pc(pc, colors=None):
if colors is None:
colors = np.full(len(pc), 0x000000, dtype=np.float64) # white: 0xFFFFFF
plot = k3d.plot()
points = k3d.points(pc, colors.astype(np.float32), point_size=0.04, shader='flat')
plot += points
plot.display()
# ## get point cloud dataset
from libs.dataset import SimpleObjectDataset
file_path = "/home/coder/databox1/datasets/ModelNet/modelnet40_ply_hdf5_2048/ply_data_train0.h5"
dataset = SimpleObjectDataset(file_path=file_path)
pc, label = dataset[0]
visualize_pc(pc)
# ## get FPS points
from torchpcp.modules.functional.sampling import furthest_point_sampling
from torchpcp.modules.functional.other import index2points
t_pc = torch.tensor([pc])
t_pc = t_pc.to(device)
t_pc = t_pc.transpose(1,2)
center_t_idxs = furthest_point_sampling(t_pc, 512)
center_t_pc = index2points(t_pc, center_t_idxs)
center_pc = pytorch_tools.t2n(center_t_pc.transpose(1,2)[0])
visualize_pc(center_pc)
# ## get KNN points
from torchpcp.modules.functional.nns import k_nearest_neighbors as knn
knn_idxs, knn_dists = knn(t_pc, center_t_pc, 5)
print("knn_idxs.shape:", knn_idxs.shape)
# ## compare C++ with python implementation.
from torchpcp.modules.functional.nns import py_k_nearest_neighbors
from torchpcp.utils.monitor import timecheck
t = timecheck()
knn_idxs, knn_dists = knn(center_t_pc, t_pc, 3)
print("c++ impl. shape:", knn_idxs.shape)
t = timecheck(t, "c++ impl. time")
t = timecheck()
knn_idxs_py, knn_dists_py = py_k_nearest_neighbors(center_t_pc, t_pc, 3, True)
print("python impl. shape:", knn_idxs_py.shape)
t = timecheck(t, "python impl. time")
# Check
# np.set_printoptions(threshold=np.Inf)
# torch.set_printoptions(threshold=np.Inf)
print(False in (knn_idxs == knn_idxs_py))
# print(knn_idxs == knn_idxs_py)
# ## compare other implementation.
# +
# PointRCNN impl. from https://github.com/sshaoshuai/PointRCNN
import pointnet2_cuda as pointnet2
from typing import Tuple
class ThreeNN(torch.autograd.Function):
@staticmethod
def forward(ctx, unknown: torch.Tensor, known: torch.Tensor) -> Tuple[torch.Tensor, torch.Tensor]:
"""
Find the three nearest neighbors of unknown in known
:param ctx:
:param unknown: (B, N, 3)
:param known: (B, M, 3)
:return:
dist: (B, N, 3) l2 distance to the three nearest neighbors
idx: (B, N, 3) index of 3 nearest neighbors
"""
assert unknown.is_contiguous()
assert known.is_contiguous()
B, N, _ = unknown.size()
m = known.size(1)
dist2 = torch.cuda.FloatTensor(B, N, 3)
idx = torch.cuda.IntTensor(B, N, 3)
pointnet2.three_nn_wrapper(B, N, m, unknown, known, dist2, idx)
return torch.sqrt(dist2), idx
@staticmethod
def backward(ctx, a=None, b=None):
return None, None
three_nn = ThreeNN.apply
# -
t = timecheck()
c_oth_dists, c_oth_idx = three_nn(center_t_pc.transpose(1,2).contiguous(), t_pc.transpose(1,2).contiguous())
# print(c_oth_idx.shape)
t = timecheck(t, "other impl. time")
print(False in (c_oth_idx == knn_idxs))
# ## compare implementations using scene data batches.
from libs.dataset import SimpleSceneDataset
from torch.utils.data import DataLoader
def speed_test(method, loader):
for i, data in enumerate(loader): pass # for speed processing
# print name
if method == 0:
t_name = "original c++ impl. time"
elif method == 1:
t_name = "original py impl. time"
elif method == 2:
t_name = "other c++ impl. time"
else:
raise NotImplementedError()
# timer start
t = timecheck()
for _ in range(100):
for i, data in enumerate(loader):
point_clouds, sem_labels, ins_labels = data
point_clouds = point_clouds[:, :3].to(device)
center_idxs = furthest_point_sample(point_clouds, 1024)
center_pc = index2points(point_clouds, center_idxs)
if method == 0:
_ = knn(center_points, point_clouds, k=3)
elif method == 1:
_ = py_k_nearest_neighbors(center_pc, point_clouds, k=3, memory_saving=False)
elif method == 2:
_ = three_nn(center_pc.transpose(1,2).contiguous(), point_clouds.transpose(1,2).contiguous())
else:
raise NotImplementedError()
# timer end
timecheck(t, t_name)
dataset = SimpleSceneDataset()
loader = DataLoader(
dataset,
batch_size=32,
num_workers=8,
pin_memory=True,
shuffle=False
)
speed_test(0, loader)
speed_test(1, loader)
speed_test(2, loader)
| examples/function_and_class/neighors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/zhz03/209_project_Blimp_modelling/blob/main/Drag%26moments.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8_jSBx7OxF_C"
# # Useful formula
# $F_{D}=\frac{1}{2} \rho v^{2} C_{D} A$
#
# $C_D,\ \rho $ can be tested once the real model out
# + [markdown] id="_z0hruXe2On_"
# # Input and Output
# Drag force
#
# Input:
# 1. $v$
# 2. $\rho$ , $C_d$...
# 3. volume V
# 4. Angle of attack $\alpha$
# 5. Angle of sideslip $\beta$
#
# Output
# 1. Drag force: $Fd_x,\ Fd_y,\ Fd_z$
# -----------------------------
# Process simulation
#
# Input:
# 1. $F_x,\ F_y,\ F_z$
# 2. $\rho$ , $C_d$
# 3. mass: m
#
# Output
# 1. Ultimate velocity
# 2. plt of process
# + id="S1Mz8CnYPmed"
import math
import matplotlib.pyplot as plt
import numpy as np
# Assume Da 4m , Db 2m, Dc 2m V^(2/3) = 4.125
Da = 4
Db = 2
Dc = 2
# Shape hull + Gondola
cdh0 = 0.025
cdg0 = 0.001
cdch = 1.0
cdcg = 1.0
Ref_Ac = 4.125
Ref_Ag = 0.1
rho = 1.2
l_gz = 1
# + colab={"base_uri": "https://localhost:8080/"} id="pvxmn7O9cyEU" outputId="6148f8e6-7f93-4bae-ac95-6c2675fc3d64"
math.sin(math.pi/2)
abs(1)
# + id="R1l53P7Dv888"
import math
import matplotlib.pyplot as plt
import numpy as np
def calculate_drag(v,alpha,beta):
# CX1 in paper
cdx = -1*(cdh0*Ref_Ac + cdg0*Ref_Ag)
fd_x =0.5*rho*v**2*(cdx*math.cos(alpha)*math.cos(alpha)*math.cos(beta)*math.cos(beta))
# CY3 in paper
cdy = -1*(cdch*1.31*Ref_Ac + cdcg*Ref_Ag)
fd_y =0.5*rho*v**2*(cdy*math.sin(beta)*math.sin(abs(beta)))
# CZ3 in paper
cdz = -1*(cdch*1.31*Ref_Ac)
fd_z =0.5*rho*v**2*(cdz*math.sin(alpha)*math.sin(abs(alpha)))
return fd_x,fd_y,fd_z
def calculate_moment(v,alpha,beta):
# CL2 in paper
cdl = -1*(cdcg*Ref_Ag*l_gz)
L =0.5*rho*v**2*(cdl*math.sin(beta)*math.sin(abs(beta)))
# CM3 in paper
cdm = -1*(cdch*0.53*Ref_Ac*Da)
M =0.5*rho*v**2*(cdm*math.sin(alpha)*math.sin(abs(alpha)))
# CN3 in paper
cdn = -1*cdm
N =0.5*rho*v**2*(cdn*math.sin(beta)*math.sin(abs(beta)))
return L,M,N
# + colab={"base_uri": "https://localhost:8080/"} id="-5UaSJxSfiOf" outputId="ea6127d2-58c3-4940-d865-a26401462664"
alpha = math.pi/10
beta = math.pi/20
v = 10
calculate_drag(v,alpha,beta)
# + colab={"base_uri": "https://localhost:8080/"} id="IcP5gDlBk5KD" outputId="192bed7d-7887-46c8-ac4a-7701cdada8ec"
calculate_moment(v,alpha,beta)
# + [markdown] id="kkNjE6UQC10V"
# # test: calculation of drag force
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="emocWazl-afK" outputId="4868660a-8beb-4c80-f7e1-b482ae73e57c"
import matplotlib.pyplot as plt
# test func: calculate_drag() :
dx = 8
dy = 2
dz = 1
fdxs = []
fdys = []
fdzs = []
vs = range(1,101)
for v in range(1,101):
fd_x,fd_y,fd_z = calculate_drag(v,v,v,dx,dy,dz)
fdxs.append(fd_x)
fdys.append(fd_y)
fdzs.append(fd_z)
plt.plot(vs,fdxs,label='$Fd_x$')
plt.plot(vs,fdys,label='$Fd_y$')
plt.plot(vs,fdzs,label='$Fd_z$')
plt.legend()
plt.xlabel("Velocity(m/s)")
plt.ylabel("force(N)")
# + [markdown] id="xmKveqcnC7Q8"
# # test: simluation with outer force
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="o1ELRigbG9cf" outputId="4d1ffdb5-1768-467d-d43c-0e903bdafb66"
ultimate_velocity(100,100,100,8,2,1,100)
# + [markdown] id="C8TZzjUJWLJ4"
# # Motion Simulation
# try to learn from
# https://www.khanacademy.org/computer-programming/modeling-air-resistance/966875281
| Code/Evaluation_system/old_version/Drag&moments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The groupby method allows you to group rows of data together
# and call aggregate functions.
import numpy as np
import pandas as pd
data = {'Company':['GOOG','GOOG','MSFT','MSFT','FB','FB'],
'Person':['Sam','Charlie','Amy','Vanessa','Carl','Sarah'],
'Sales':[200,120,340,124,243,350]}
df = pd.DataFrame(data)
df
by_company = df.groupby("Company")
#ignores non-numerica data
by_company.sum()
by_company.mean()
by_company.std()
df.groupby('Company').sum().loc['FB']
by_company.min()
by_company.max()
by_company.count()
by_company.describe()
by_company.describe().transpose()
by_company.describe().transpose()['MSFT']
| python-for-data-analysis/pandas/group-by.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# ### Importando os pacotes de funcoes numpy e matplotlib.pyplot
#
# * Usamos o comando "as" para encurtar os caminhos.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Usamos o numpy para carregar os dados do txt
#
# * Definimos que os comentários seriam precedidos por "%".
#
# * Utilizamos ";" ao final da linha para ocultar os dados e não poluir visualmente o código.
dados = np.loadtxt("dados/brazil-TAVG-Trend.txt", comments= "%");
# ### Nomeamos as variáveis
#
# * Para a tarefa, usaremos as anomalias anuais (anom_an).
# * Para o bônus, usaremmos as anomalias decadais (anom_dec) e incertezas decadais(inc_dec)
ano = dados[:,0]
mes = dados [:,1]
anom_an = dados[:,4]
anom_dec = dados[:,8]
inc_dec = dados[:,9]
# ### Calculamos a média e o desvio padrão das anomalias
# * Média das anomalias anuais.
media_an = np.nanmean(anom_an)
print(media_an)
# * Desvio Padrão das anomalias anuais
dp_an = np.nanstd(anom_an)
print(dp_an)
# * Média das anomalias decadais (bônus)
media_dec = np.nanmean(anom_dec)
print(media_dec)
# ### Plotando o gráfico
# +
plt.plot(ano,25 + anom_an, "black", label="Média móvel -12 meses") #Utilizamos 25 como a temperatura média
plt.plot(ano,25 + anom_dec, "red", label="Média móvel -10 anos") #bônus
plt.plot(ano,((25 + anom_dec) + inc_dec), "gray",) #bônus
plt.plot(ano,((25 + anom_dec) - inc_dec), "gray") #bônus
plt.grid(True) #adicionando grade
plt.xlim(1820,2020) #Definindo a escala do gráfico
plt.ylim(23,26.5)
#Formatacão-> legenda , nome dos eixos e título
plt.xlabel ("Anos")
plt.ylabel ("Temperatura °C")
plt.title("Brasil")
plt.legend(loc="upper left")
# -
# * Não conseguimos prencher o espaco entre o limite superior e inferior das incertezas.
# * *Utilizamos 25 para o valor médio de temperatura, logo as anomalias são somadas ou subtraídas desse valor.
| python1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
# Set a seed so that the random numbers will be reproducible
np.random.seed(12)
# -
# # Generate attributes x1, x2 for each of numalts (J) alternatives
# +
# For now, J << 1000 alternatives to speed up runtimes
numalts = 50
def rand(len, min, max):
""" Generate `len` random floats uniformly distributed from `min` to `max` """
return (max - min) * np.random.rand(len) + min
# Attribute x is uniformly distributed over [-2, 1] for half the alternatives
# and over [-1, 2] for the other half, as in Guevara & Ben-Akiva
# X = np.concatenate((rand(numalts/2, -2, 1), rand(numalts/2, -1, 2)))
# Or, attribute x is uniformly distributed over [0, 10] for half the alternatives
# and over [100, 110] for the other half, to induce bias in estimation
X = np.concatenate((rand(int(numalts/2), 0, 10), rand(int(numalts/2), 100, 110)))
# -
X
print(pd.DataFrame(X[:int(numalts/2)]).describe())
print(pd.DataFrame(X[int(numalts/2):]).describe())
# # Generate taste coefficient beta for each of numobs (N) agents
# +
# For regular MNL, use a single value instead of a distribution as
# Guevara & Ben-Akiva used for the mixture model
numobs = 1000 # agents/observations
beta = np.zeros(1000) + 1.5
# beta = 0.8 * np.random.randn(numobs) + 1.5
# -
pd.DataFrame(beta).describe()
# # Simulate a choice from numalts (J) alternatives for each of numobs (N) agents
# +
# Generate a utility matrix for N agents choosing among J alternatives
U = [[beta[n]*x + np.random.gumbel() for x in X] for n in range(numobs)]
len(U), len(U[0])
# -
len(U)
# +
# Each agent chooses the alternative with highest utility
choices = [np.argmax(a) for a in U]
len(choices), choices[:10]
# -
# # 2. Estimate beta without sampling, using PyLogit MNL
import pylogit
from collections import OrderedDict
# +
# Set up the estimation dataset in long format
d = [[n, i, int(choices[n]==i), X[i]] for n in range(numobs) for i in range(numalts)]
df = pd.DataFrame(d, columns=['obs_id', 'alt_id', 'chosen', 'x'])
# -
df.head()
df.alt_id.describe()
df.describe()
# +
# Set up reusable model spec
spec = OrderedDict([('x', 'all_same')])
labels = OrderedDict([('x', 'beta_x')])
# +
# Set up reusable code to estimate a model
def estimate_model(init_val):
"""
Initialize and fit a model, returning it as an object. Will use the
current values of `df`, `spec`, and `labels`.
"""
m = pylogit.create_choice_model(data = df,
alt_id_col = 'alt_id',
obs_id_col = 'obs_id',
choice_col = 'chosen',
specification = spec,
model_type = "MNL",
names = labels)
m.fit_mle(init_vals = np.array([init_val]))
return m
# -
# %%time
m = estimate_model(init_val = 1.2)
m.get_statsmodels_summary()
# # 3a. Estimate beta with random sampling of alternatives
# +
# In the estimation dataset, for each observation include a row for the
# chosen alternative, plus K-1 other alternatives sampled randomly
# without replacement, where K < J.
# Some more notation:
# - true choice set C = range(J)
# - restricted choice set D_n is a subset of C, where len(D_n) = K
# +
# TO DO - rewrite to use sampling weights
def alts(obs_id, C, K):
"""
This function generates a restricted choice set D for a particular
observation. Expects list `C` of alternatives to sample from (either
the full choice set or a stratrum), int `K` alternatives to sample,
and list `choices` of the alt_id chosen for each obs_id. Returns list
of K alt_id's including the chosen one.
"""
chosen = choices[obs_id] # id of chosen alternative
unchosen = [i for i in C if chosen != i] # id's of unchosen alts
sample_unchosen = np.random.choice(unchosen, size=K-1, replace=False).tolist()
return np.sort([chosen] + sample_unchosen)
print(alts(0, range(numalts), 5))
# +
# Set up the estimation dataset, which can use the same spec as earlier
C = range(numalts) # choice set to sample from
K = 10
d = [[n, i, int(choices[n]==i), X[i]] for n in range(numobs) for i in alts(n, C, K)]
df = pd.DataFrame(d, columns=['obs_id', 'alt_id', 'chosen', 'x'])
# -
df.head()
df.shape
df.describe()
# %%time
m = estimate_model(init_val = 1.2)
m.get_statsmodels_summary()
# # Run 1000x with different samples of alternatives
# +
# %%time
# %%capture
beta = []
C = range(numalts)
K = 10
for i in range(100):
d = [[n, i, int(choices[n]==i), X[i]] for n in range(numobs) for i in alts(n, C, K)]
df = pd.DataFrame(d, columns=['obs_id', 'alt_id', 'chosen', 'x'])
m = estimate_model(init_val = 1.2)
beta.append(m.params.beta_x)
# -
pd.Series(beta).describe()
# Looks unbiased, as expected. It's very close to the true beta of 1.5
# # 3b. Estimate beta with over-sampling of irrelevant alternatives
# +
# Recall that half the values of x are in the range [0, 10] and half are
# in the range [100, 110]. The taste coefficient is positive, so the first
# set of alternatives is much less relevant than the second set.
C = range(int(numalts/2)) # alternatives to sample from
K = 10
d = [[n, i, int(choices[n]==i), X[i]] for n in range(numobs) for i in alts(n, C, K)]
df = pd.DataFrame(d, columns=['obs_id', 'alt_id', 'chosen', 'x'])
# -
df.head()
df.describe()
# %%time
m = estimate_model(init_val = 1.5)
m.get_statsmodels_summary()
# # 5. MNL with sampling correction
# Utility of alternative j: $$ V_{j} = \beta x_{j} $$
#
# With sampling, we have to account for the restricted choice set (from Eq 6 in Guevara & Ben-Akiva 2013):
#
# $$ V_j = \beta x_j + \ln \pi(D \mid j) $$
# Where pi is the conditional probability that we would construct the choice set D given that alternative j was chosen. This goes into the likelihood function in both the numerator and denominator.
#
# $$ L_n = \frac {exp(\beta x_i + \ln \pi(D_n \mid i))} {\sum_{j \epsilon D_n} exp(\beta x_j + \ln \pi(D_n \mid j))} $$
# How to calculate pi? From the original formulation of this in McFadden 1978: "Suppose D is comprized of i plus a sample of alternatives from the set C\{i}, obtained by considering each element of this set independently, and including it with probability p. Then, the probability of D will depend solely on the number of elements K it contains."
#
# $$ \pi(D) = p^{K-1} (1 - p)^{J-K} $$
# (?? Without replacement, i think it should be the n-choose-k binomial coefficient, where n=J-1 and k=K-1)
#
# $$ \pi(D) = {n \choose k} = \frac {(K-1)!(J-K)!} {(J-1)!} $$
# +
# Add a column in the estimation data for the constant
N = 1000
d = [[n, i, int(C[n]==i), X[i], 1] for n in range(N) for i in alts(n)]
df = pd.DataFrame(d, columns=['obs_id', 'alt_id', 'choice', 'x', 'const'])
# -
| notebooks/93_mnl_sampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="color:#777777;background-color:#ffffff;font-size:12px;text-align:right;">
# prepared by <NAME> (QuSoft@Riga) | November 07, 2018
# </div>
# <table><tr><td><i> I have some macros here. If there is a problem with displaying mathematical formulas, please run me to load these macros.</i></td></td></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\inner}[2]{\langle #1,#2\rangle} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> <font color="blue"> Solutions for </font>Probabilistic States</h2>
# <a id="task2"></a>
# <h3> Task 2 </h3>
#
# Suppose that Fyodor hiddenly rolls a loaded (tricky) dice with the bias
#
# $$ Pr(1):Pr(2):Pr(3):Pr(4):Pr(5):Pr(6) = 7:5:4:2:6:1 . $$
#
# Represent your information on the result as a column vector. Remark that the size of your column should be 6.
#
# You may use python for your calculations.
# <h3>Solution</h3>
# +
# all portions are stored in a list
all_portions = [7,5,4,2,6,1];
# let's calculate the total portion
total_portion = 0
for i in range(6):
total_portion = total_portion + all_portions[i]
print("total portion is",total_portion)
# find the weight of one portion
one_portion = 1/total_portion
print("the weight of one portion is",one_portion)
print() # print an empty line
# now we can calculate the probabilities of rolling 1,2,3,4,5, and 6
for i in range(6):
print("the probability of rolling",(i+1),"is",(one_portion*all_portions[i]))
# -
# <a id="task4"></a>
# <h3> Task 4 </h3>
#
# For a system with 4 states, randomly create a probabilistic state, and print its entries, e.g., $ 0.16~~0.17~~0.02~~0.65 $.
#
# You may pick your random numbers between 0 and 100 (or 1000), and then divide each by 100 (or 1000) to represent it as a probability.
# <h3>Solution</h3>
# +
# we will randomly create a probabilistic state
#
# we should be careful about two things:
# 1. a probability value must be between 0 and 1
# 2. the total probability must be 1
#
# therefore, we can randomly pick three probability values.
# once we have three probability values, the fourth one is determined automatically
# the fourth one cannot be arbitrary, because the summation of the four values must be 1
# let's use a list of size 4
# initial values are zeros
my_state = [0,0,0,0]
# we pick three random probabilistic values
from random import randrange
# I assume that I have the following total value to distribute to four parts
total = 1000
# I will randomly pick a value, and then continue with the remaining value
for i in range(3): # let's find the three values
pick_a_value = randrange(total)
print("I picked",pick_a_value)
my_state[i] = pick_a_value
total = total - pick_a_value # remaining value for the others
my_state[3] = total # this is the remaining value after three iterations
print("The remaining value is",total)
# let's verify the summation of the elements in my_state
sum = 0
print() # print an empty line
for i in range(len(my_state)):
sum = sum + my_state[i]
print("the summation of the elements in my_state is",sum)
# let's convert the selected values to the probabilities
# we can also call this procedure as **NORMALIZATION**
for i in range(len(my_state)):
my_state[i] = my_state[i]/1000
print() # print an empty line
print("the entries of my probabilistic state:")
# let's print all probabilities
for i in range(len(my_state)): print(my_state[i])
| community/awards/teach_me_quantum_2018/bronze/bronze-solutions/B32_Probabilistic_States_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SweetViz
# +
## Module is a python file with functionality.<br>
## Package is a collection of modules.
## Library is a collection of Packages.
# -
# Hi there. Today, we are going to see how to use SweetViz library in Python which will enable us to perform powerful Exploratory Data Analysis(EDA) on your dataset. So,let's get started.
# First, you will have to pip install this package as it is not an in-built Python package. You can do so from the command prompt or using !pip install sweetviz from jupyter notebook environment.
# I will be using USA Housing data in this example.
# !pip install sweetviz
# Import the necessary libraries
import numpy as np
import pandas as pd
import sweetviz
df = pd.read_csv(r"C:\Users\<NAME>\Desktop\Data science\original\Refactored_Py_DS_ML_Bootcamp-master\11-Linear-Regression\USA_housing.csv")
df.head()
# In this dataset, price column is the target feature or dependent variable.
# # Analyzing a DataFrame
# Use the analysis function from sweetviz module to create a 'DataframeReport' object.
analysis = sweetviz.analyze([df,"EDA"], target_feat='Price')
# Check-out the type of this analysis object.
type(analysis)
# Render the output on a web page.
analysis.show_html('EDA.html')
# This is an amazing visualization library for your data as you instantly get various insights into your data which you could have done manually but would have taken a lot more time. <br>
# For numerical features, you get point plot, histogram, number of value missing, number of distinct values, quartile values and more useful information like skewness of the column.<br>
# For categorical features, along with the number of distinct and missing values, you
# <br>
# Additionally, you also get the the 'Associations' or pair-wise correlations between 2 variables which is helpful for determining feature importance.
# You can also use this library to comapre two DataFrames,say, your Training set and Test set and infer some meaning from the comparison.
train = df[:3000]
test = df[3000:]
# Consider 'train' to be the Training data.
# Consider 'test' to be the Test data.
# The command to perform EDA comparison is:
analysis = sweetviz.compare([train,"Train"],[test,"Test"], "Price") # Price is the target variable common to both tables
# Now you can view your results.
analysis.show_html('EDA2.html')
# Now, you can see comparison between the Train and Test dataset differentiate by different colors for all paramters discussed above.<br>
# Therefore, this is a handy module
| _notebooks/2020-09-19-SweetViz- Automated EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quantum Inspire performance test
#
# We compare performance of the simulator with the circuit from
#
# "Overview and Comparison of Gate Level Quantum Software Platforms", https://arxiv.org/abs/1807.02500
# ## Define the circuit
# +
import time
import os
import numpy as np
from IPython.display import display
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
from qiskit.tools.visualization import plot_histogram, circuit_drawer
from quantuminspire.credentials import get_authentication
from quantuminspire.qiskit import QI
QI_URL = os.getenv('API_URL', 'https://api.quantum-inspire.com/')
# -
# We define the circuit based on the number of qubits and the depth (e.g. the number of iterations of the unit building block).
# +
def pcircuit(nqubits, depth = 10):
""" Circuit to test performance of quantum computer """
q = QuantumRegister(nqubits)
ans = ClassicalRegister(nqubits)
qc = QuantumCircuit(q, ans)
for level in range(depth):
for qidx in range(nqubits):
qc.h( q[qidx] )
qc.barrier()
for qidx in range(nqubits):
qc.rx(np.pi/2, q[qidx])
qc.barrier()
for qidx in range(nqubits):
if qidx!=0:
qc.cx(q[qidx], q[0])
for qidx in range(nqubits):
qc.measure(q[qidx], ans[qidx])
return q, qc
q,qc = pcircuit(4, 1)
qc.draw(output='mpl')
# -
# ## Run the cirquit on the Quantum Inspire simulator
# First we make a connection to the Quantum Inspire website.
authentication = get_authentication()
QI.set_authentication(authentication, QI_URL)
# We create a QisKit backend for the Quantum Inspire interface and execute the circuit generated above.
qi_backend = QI.get_backend('QX single-node simulator')
job = execute(qc, qi_backend)
# We can wait for the results and then print them
result = job.result()
print('Generated histogram:')
print(result.get_counts())
# Visualization can be done with the normal Python plotting routines, or with the QisKit SDK.
plot_histogram(result.get_counts(qc))
# To compare we will run the circuit with 20 qubits and depth 20. This takes:
#
# * QisKit: 3.7 seconds
# * ProjectQ: 2.0 seconds
# Our simulator runs for multiple shots (unless full state projection is used). More details will follow later.
# +
q, qc = pcircuit(10, 10)
start_time = time.time()
job = execute(qc, qi_backend, shots=8)
job.result()
interval = time.time() - start_time
print('time needed: %.1f [s]' % (interval,))
# -
| docs/notebooks/qi-performance-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import sklearn.datasets
X,y = sklearn.datasets.make_moons(200, noise = 0.15)
plt.scatter(X[:, 0], X[:, 1], c = y)
X.shape
input_neurons = 2
output_neurons = 2 # 2 for binary cross entropy, 1 for cross entropy
samples = x.shape[0]
learning_rate = 0.001
lamda = 0.01
# key is input weight from first layer w1, value will be updated
model_dic = {'W1': W1, 'b1': b1, 'W2':W2, 'b2':b2}
def retreive(model_dict):
W1 = model_dict['W1']
b1 = model_dict['b1']
W2 = model_dict['W2']
b2 = model_dict['b2']
return W1, b1, wW, b2
def forward(x, model_dict):
W1, b1, W2, b2 = retreive(model_dict)
z1 = X.dot(W1) + b1
a1 = np.tanh(z1)
z2 = a1.dot(W2) + b2
a2 = np.tanh(z2)
exp_scores = np.exp(z2)
# normalize
softmax = exp_scores / np.sum(exp_scores, axis = 1, keepdims = True)
return z1, a1, softmax
# forward propagation
# calculate the difference between actural and predicted outputs, use cross entropy loss
def loss(softmax, y):
W1, b1, W2, b2 = retreive(model_dict)
m = np.zeros(200)
for i,correct_index in enumerate(y):
predicted = softmax[i][correct_index]
m[i] = predicted
log_prob = -np.log(m) # log_prob = -np.log(predicted)
loss = np.sum(log_prob)
reg_loss = lambda_reg / 2 * (np.sum(np.square(W1)) + np.sum(np.square(W2)))
loss+= reg_loss
return float(loss / y.shape[0])
# prediction
def predict(model_dict, x):
W1, b1, W2, b2 = retreive(model_dict)
z1 = x.dot(W1) + b1
a1 = np.tanh(z1)
z2 = a1.dot(W2) + b2
exp_scores = np.exp(z2)
softmax = exp_scores / np.sum(exp_scores, axis = 1, keepdims = True) # (200,2)
return np.argmax(softmax, axis = 1) # (200,)
| 05_Numpy_neuralNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import time
from agents import Agent, AlphaFour
from collections import namedtuple, deque
from random import choice, sample
import importlib
from connectboard import ConnectBoard
os.environ["KERAS_BACKEND"] = "plaidml.keras.backend"
import keras
# +
def get_game_state(game_board: np.ndarray) -> np.ndarray:
"""Returns the AlphaFour representation of the game board.
When AlphaFour gets the game board, the current player is 1
and the opponent is -1. We translate that to a two layer state
where the current player's pieces are 1's in the first slice,
and the opponent's pieces are 1's in the second slice
Args:
game_board: A numpy array representing the current game state.
Returns:
A numpy array containing the AlphaFour representation of the
given game state.
"""
p1_pieces = np.where(game_board == 1, 1, 0)
p2_pieces = np.where(game_board == -1, 1, 0)
alpha_four_state = np.array([p1_pieces, p2_pieces])
return alpha_four_state
# Named tuple for training data. Stores state, move probabilities, and state value
TrainingSample = namedtuple('TrainingSample', 'state probs value')
# -
def self_play(agent1: AlphaFour, agent2: AlphaFour):
game_board = np.zeros((6,7))
turn = 0
states = []
probs = []
values = []
state = get_game_state(game_board)
while True:
states.append(state.copy()) # Add current state
if state[2].all():
# Get best move, and probability of all moves from current state
move,prob = agent1.get_move_with_prob(state)
state[0] += move
state[2] = np.zeros((6,7))
else:
move,prob = agent2.get_move_with_prob(state)
state[1] += move
state[2] = np.ones((6,7))
probs.append(prob) # Store move probabilities from current state
val = helpers.winner(state[0] - state[1])
if val is not None:
# Add the final state to our arrays
states.append(state.copy())
probs.append(np.zeros((1,7)))
val = -1*abs(val) # If game is over, current player lost unless it's a tie.
break
turn += 1
for i in range(len(probs)):
values.append(val * (-1)**i)
values = values[::-1]
data = [Data(states[i], probs[i], values[i]) for i in range(len(probs))]
return data
D = self_play(AlphaFour('P1'), AlphaFour('P2'))
# +
REPLAY_BUFFER_SIZE = 100000 # Number of past steps to store. This is where our training sample is drawn from
SELF_PLAY_BATCH_SIZE = 100 # How many games to play before updating the buffer
TRAINING_SET_SIZE = 1024 # Size of the training set to sample from the replay buffer
replay_buffer = deque(maxlen=REPLAY_BUFFER_SIZE)
# START OF ONE TRAINING LOOP
# ==========================================================================
players = [AlphaFour('Best'), AlphaFour('New')]
total_count = 0
# Generate new self play games.
for ii in range(SELF_PLAY_BATCH_SIZE):
print(f'\r{ii}', end='')
i = np.random.randint(2) # randomize who plays first
p1 = players[i]
p2 = players[(i+1)%2]
D = self_play(p1,p2)
for d in D:
total_count += 1
replay_buffer.append(d)
train_set = sample(replay_buffer, TRAINING_SET_SIZE)
# Train new bot with updated data
# Play matches between new and old. If new wins more than 55%, replace old with New
# ==========================================================================
# +
state = train_set[123].S
moves = helpers.get_legal_moves(state[0] + state[1])
moves = moves[1:,:,:]
print(moves)
for col in range(7):
print(moves[:,:,col].sum())
# +
state = train_set[123].S
moves = helpers.get_legal_moves(state[0] + state[1])
moves = moves[1:]
col_has_move = moves.sum(axis=0).sum(axis=0)
move_idx = 0
for i in range(7):
if col_has_move[i]:
new_state = state + np.array([moves[move_idx], np.zeros((6,7)), np.zeros((6,7))])
move_idx += 1
print(new_state, '\n\n')
# -
| alpha_four_prototyping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Create a Batch Inferencing Service
#
# Imagine a health clinic takes patient measurements all day, saving the details for each patient in a separate file. Then overnight, the diabetes prediction model can be used to process all of the day's patient data as a batch, generating predictions that will be waiting the following morning so that the clinic can follow up with patients who are predicted to be at risk of diabetes. With Azure Machine Learning, you can accomplish this by creating a *batch inferencing pipeline*; and that's what you'll implement in this exercise.
# ## Connect to your workspace
#
# To get started, connect to your workspace.
#
# > **Note**: If you haven't already established an authenticated session with your Azure subscription, you'll be prompted to authenticate by clicking a link, entering an authentication code, and signing into Azure.
# +
import azureml.core
from azureml.core import Workspace
# Load the workspace from the saved config file
ws = Workspace.from_config()
print('Ready to use Azure ML {} to work with {}'.format(azureml.core.VERSION, ws.name))
# -
# ## Train and register a model
#
# Now let's train and register a model to deploy in a batch inferencing pipeline.
# +
from azureml.core import Experiment
from azureml.core import Model
import pandas as pd
import numpy as np
import joblib
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
# Create an Azure ML experiment in your workspace
experiment = Experiment(workspace=ws, name='mslearn-train-diabetes')
run = experiment.start_logging()
print("Starting experiment:", experiment.name)
# load the diabetes dataset
print("Loading Data...")
diabetes = pd.read_csv('data/diabetes.csv')
# Separate features and labels
X, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values
# Split data into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# Train a decision tree model
print('Training a decision tree model')
model = DecisionTreeClassifier().fit(X_train, y_train)
# calculate accuracy
y_hat = model.predict(X_test)
acc = np.average(y_hat == y_test)
print('Accuracy:', acc)
run.log('Accuracy', np.float(acc))
# calculate AUC
y_scores = model.predict_proba(X_test)
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
run.log('AUC', np.float(auc))
# Save the trained model
model_file = 'diabetes_model.pkl'
joblib.dump(value=model, filename=model_file)
run.upload_file(name = 'outputs/' + model_file, path_or_stream = './' + model_file)
# Complete the run
run.complete()
# Register the model
run.register_model(model_path='outputs/diabetes_model.pkl', model_name='diabetes_model',
tags={'Training context':'Inline Training'},
properties={'AUC': run.get_metrics()['AUC'], 'Accuracy': run.get_metrics()['Accuracy']})
print('Model trained and registered.')
# -
# ## Generate and upload batch data
#
# Since we don't actually have a fully staffed clinic with patients from whom to get new data for this exercise, you'll generate a random sample from our diabetes CSV file, upload that data to a datastore in the Azure Machine Learning workspace, and register a dataset for it.
# +
from azureml.core import Datastore, Dataset
import pandas as pd
import os
# Set default data store
ws.set_default_datastore('workspaceblobstore')
default_ds = ws.get_default_datastore()
# Enumerate all datastores, indicating which is the default
for ds_name in ws.datastores:
print(ds_name, "- Default =", ds_name == default_ds.name)
# Load the diabetes data
diabetes = pd.read_csv('data/diabetes2.csv')
# Get a 100-item sample of the feature columns (not the diabetic label)
sample = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].sample(n=100).values
# Create a folder
batch_folder = './batch-data'
os.makedirs(batch_folder, exist_ok=True)
print("Folder created!")
# Save each sample as a separate file
print("Saving files...")
for i in range(100):
fname = str(i+1) + '.csv'
sample[i].tofile(os.path.join(batch_folder, fname), sep=",")
print("files saved!")
# Upload the files to the default datastore
print("Uploading files to datastore...")
default_ds = ws.get_default_datastore()
default_ds.upload(src_dir="batch-data", target_path="batch-data", overwrite=True, show_progress=True)
# Register a dataset for the input data
batch_data_set = Dataset.File.from_files(path=(default_ds, 'batch-data/'), validate=False)
try:
batch_data_set = batch_data_set.register(workspace=ws,
name='batch-data',
description='batch data',
create_new_version=True)
except Exception as ex:
print(ex)
print("Done!")
# -
# ## Create compute
#
# We'll need a compute context for the pipeline, so we'll use the following code to specify an Azure Machine Learning compute cluster (it will be created if it doesn't already exist).
#
# > **Important**: Change *your-compute-cluster* to the name of your compute cluster in the code below before running it! Cluster names must be globally unique names between 2 to 16 characters in length. Valid characters are letters, digits, and the - character.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
cluster_name = "your-compute-cluster"
try:
# Check for existing compute target
inference_cluster = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
# If it doesn't already exist, create it
try:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS11_V2', max_nodes=2)
inference_cluster = ComputeTarget.create(ws, cluster_name, compute_config)
inference_cluster.wait_for_completion(show_output=True)
except Exception as ex:
print(ex)
# -
# ## Create a pipeline for batch inferencing
#
# Now we're ready to define the pipeline we'll use for batch inferencing. Our pipeline will need Python code to perform the batch inferencing, so let's create a folder where we can keep all the files used by the pipeline:
# +
import os
# Create a folder for the experiment files
experiment_folder = 'batch_pipeline'
os.makedirs(experiment_folder, exist_ok=True)
print(experiment_folder)
# -
# Now we'll create a Python script to do the actual work, and save it in the pipeline folder:
# +
# %%writefile $experiment_folder/batch_diabetes.py
import os
import numpy as np
from azureml.core import Model
import joblib
def init():
# Runs when the pipeline step is initialized
global model
# load the model
model_path = Model.get_model_path('diabetes_model')
model = joblib.load(model_path)
def run(mini_batch):
# This runs for each batch
resultList = []
# process each file in the batch
for f in mini_batch:
# Read the comma-delimited data into an array
data = np.genfromtxt(f, delimiter=',')
# Reshape into a 2-dimensional array for prediction (model expects multiple items)
prediction = model.predict(data.reshape(1, -1))
# Append prediction to results
resultList.append("{}: {}".format(os.path.basename(f), prediction[0]))
return resultList
# -
# Next we'll define a run context that includes the dependencies required by the script
# +
from azureml.core import Environment
from azureml.core.runconfig import DEFAULT_CPU_IMAGE
from azureml.core.runconfig import CondaDependencies
# Add dependencies required by the model
# For scikit-learn models, you need scikit-learn
# For parallel pipeline steps, you need azureml-core and azureml-dataprep[fuse]
cd = CondaDependencies.create(conda_packages=['scikit-learn','pip'],
pip_packages=['azureml-defaults','azureml-core','azureml-dataprep[fuse]'])
batch_env = Environment(name='batch_environment')
batch_env.python.conda_dependencies = cd
batch_env.docker.base_image = DEFAULT_CPU_IMAGE
print('Configuration ready.')
# -
# You're going to use a pipeline to run the batch prediction script, generate predictions from the input data, and save the results as a text file in the output folder. To do this, you can use a **ParallelRunStep**, which enables the batch data to be processed in parallel and the results collated in a single output file named *parallel_run_step.txt*.
#
# > **Note**: An *'enabled' is deprecated* warning may be displayed - you can ignore this.
# +
from azureml.pipeline.steps import ParallelRunConfig, ParallelRunStep
from azureml.pipeline.core import PipelineData
from azureml.core.runconfig import DockerConfiguration
default_ds = ws.get_default_datastore()
output_dir = PipelineData(name='inferences',
datastore=default_ds,
output_path_on_compute='diabetes/results')
parallel_run_config = ParallelRunConfig(
source_directory=experiment_folder,
entry_script="batch_diabetes.py",
mini_batch_size="5",
error_threshold=10,
output_action="append_row",
environment=batch_env,
compute_target=inference_cluster,
node_count=2)
parallelrun_step = ParallelRunStep(
name='batch-score-diabetes',
parallel_run_config=parallel_run_config,
inputs=[batch_data_set.as_named_input('diabetes_batch')],
output=output_dir,
arguments=[],
allow_reuse=True
)
print('Steps defined')
# -
# Now it's time to put the step into a pipeline, and run it.
#
# > **Note**: This may take some time!
# +
from azureml.core import Experiment
from azureml.pipeline.core import Pipeline
pipeline = Pipeline(workspace=ws, steps=[parallelrun_step])
pipeline_run = Experiment(ws, 'mslearn-diabetes-batch').submit(pipeline)
pipeline_run.wait_for_completion(show_output=True)
# -
# When the pipeline has finished running, the resulting predictions will have been saved in the outputs of the experiment associated with the first (and only) step in the pipeline. You can retrieve it as follows:
# +
import pandas as pd
import shutil
# Remove the local results folder if left over from a previous run
shutil.rmtree('diabetes-results', ignore_errors=True)
# Get the run for the first step and download its output
prediction_run = next(pipeline_run.get_children())
prediction_output = prediction_run.get_output_data('inferences')
prediction_output.download(local_path='diabetes-results')
# Traverse the folder hierarchy and find the results file
for root, dirs, files in os.walk('diabetes-results'):
for file in files:
if file.endswith('parallel_run_step.txt'):
result_file = os.path.join(root,file)
# cleanup output format
df = pd.read_csv(result_file, delimiter=":", header=None)
df.columns = ["File", "Prediction"]
# Display the first 20 results
df.head(20)
# -
# ## Publish the Pipeline and use its REST Interface
#
# Now that you have a working pipeline for batch inferencing, you can publish it and use a REST endpoint to run it from an application.
# +
published_pipeline = pipeline_run.publish_pipeline(
name='diabetes-batch-pipeline', description='Batch scoring of diabetes data', version='1.0')
published_pipeline
# -
# Note that the published pipeline has an endpoint, which you can see in the Azure portal. You can also find it as a property of the published pipeline object:
rest_endpoint = published_pipeline.endpoint
print(rest_endpoint)
# To use the endpoint, client applications need to make a REST call over HTTP. This request must be authenticated, so an authorization header is required. To test this out, we'll use the authorization header from your current connection to your Azure workspace, which you can get using the following code:
#
# > **Note**: A real application would require a service principal with which to be authenticated.
# +
from azureml.core.authentication import InteractiveLoginAuthentication
interactive_auth = InteractiveLoginAuthentication()
auth_header = interactive_auth.get_authentication_header()
print('Authentication header ready.')
# -
# Now we're ready to call the REST interface. The pipeline runs asynchronously, so we'll get an identifier back, which we can use to track the pipeline experiment as it runs:
# +
import requests
rest_endpoint = published_pipeline.endpoint
response = requests.post(rest_endpoint,
headers=auth_header,
json={"ExperimentName": "mslearn-diabetes-batch"})
run_id = response.json()["Id"]
run_id
# -
# Since we have the run ID, we can use the **RunDetails** widget to view the experiment as it runs:
# +
from azureml.pipeline.core.run import PipelineRun
from azureml.widgets import RunDetails
published_pipeline_run = PipelineRun(ws.experiments['mslearn-diabetes-batch'], run_id)
# Block until the run completes
published_pipeline_run.wait_for_completion(show_output=True)
# -
# Wait for the pipeline run to complete, and then run the following cell to see the results.
#
# As before, the results are in the output of the first pipeline step:
# +
import pandas as pd
import shutil
# Remove the local results folder if left over from a previous run
shutil.rmtree('diabetes-results', ignore_errors=True)
# Get the run for the first step and download its output
prediction_run = next(pipeline_run.get_children())
prediction_output = prediction_run.get_output_data('inferences')
prediction_output.download(local_path='diabetes-results')
# Traverse the folder hierarchy and find the results file
for root, dirs, files in os.walk('diabetes-results'):
for file in files:
if file.endswith('parallel_run_step.txt'):
result_file = os.path.join(root,file)
# cleanup output format
df = pd.read_csv(result_file, delimiter=":", header=None)
df.columns = ["File", "Prediction"]
# Display the first 20 results
df.head(20)
# -
# Now you have a pipeline that can be used to batch process daily patient data.
#
# **More Information**: For more details about using pipelines for batch inferencing, see the [How to Run Batch Predictions](https://docs.microsoft.com/azure/machine-learning/how-to-run-batch-predictions) in the Azure Machine Learning documentation.
| notebooks/06 - Working with Batch Inferencing Service.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mahfuz978/TECH-I.S.---DATA-PROCESSING/blob/main/Mahfuzur_rahman_VISUALIZATION.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="manGLFZyjduK"
from matplotlib import pyplot as plt
import numpy as np
# %matplotlib inline
# + id="sAnllwQAnsKd"
def create_sample_chart():
x_series = np.arange(10)
y_series = x_series**2
# plotting to our can vas in memory
plt.plot(x_series,y_series)
# title of our canvas
plt.title('Title')
# x-axis label
plt.xlabel('X Axis')
# Y-axis label
plt.ylabel('Y Axis')
# showing what we plotted
plt.show();
# + id="mezZVIJRny51" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="b933b395-87fc-4dc9-ad73-834c3d474517"
create_sample_chart()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="2iLjmsgYRTuk" outputId="6af4e7a3-ca3b-4fc1-a704-381a5e267197"
def sine_wave():
x_series = np.arange(0,11,0.1)
y_series = np.sin(x_series)
# plotting to our can vas in memory
plt.plot(x_series,y_series)
# title of our canvas
plt.title('Title')
# x-axis label
plt.xlabel('X Axis')
# Y-axis label
plt.ylabel('Y Axis')
# showing what we plotted
plt.show();
sine_wave()
# + colab={"base_uri": "https://localhost:8080/", "height": 331} id="aq0prt-uSiAA" outputId="5e6349cb-a548-48cf-a3e0-641e54e6f825"
import pandas as pd
url = 'https://raw.githubusercontent.com/Tech-i-s/data-science-course-wiki/master/Step%201-3%20Data%20Processing/03_EDA%20and%20Visualisation/01_visualisation/data/weather_2012.csv?token=<KEY>'
df1 = pd.read_csv(url, parse_dates=True, index_col='Date/Time')
df1.head(5)
# + colab={"base_uri": "https://localhost:8080/"} id="10YcoRuRd-zs" outputId="51a03f17-cc03-497b-b95e-0f62ca00ea48"
df1.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 442} id="RJ4g6LmNTo_5" outputId="835ff06f-ddeb-4d87-bf7f-759f4b9e9148"
monthly_data = df1.groupby(df1.index.month).mean()
monthly_data
# + colab={"base_uri": "https://localhost:8080/"} id="ISngo02QiU20" outputId="485b1346-d89d-4f79-ca88-4d07863548c8"
monthly_data.index
# + colab={"base_uri": "https://localhost:8080/"} id="dWq9H9wVdI-p" outputId="a62a1de6-f454-48bd-e96d-db847bb80c41"
monthly_data.shape
# + id="jI_DPHs5Ul5v"
x_series = monthly_data.index
y_series = monthly_data['Temp (C)']
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="9D-otHRkU7jS" outputId="e513afb5-68bc-49b5-d2f1-9e339cfdd21c"
plt.plot(x_series, y_series)
plt.title('Temperature Trend, 2012')
plt.xlabel('Month')
plt.ylabel('Temp (C)')
plt.show();
# + colab={"base_uri": "https://localhost:8080/"} id="iswkIwW_U9YU" outputId="962f5545-fd75-494e-ee13-3f8fb4234c97"
# First, get calendar month names
import calendar
calendar_months = calendar.month_name[1:]
print(calendar_months)
# + id="IjIR97UGfd0v"
# def convert(x):
# month = calendar.month_name[x]
# for i in x:
# return month
# + id="2jK9ynKNkYWn"
# def convert():
# for i in x:
# return calendar.month_name[x]
# + id="KsCShirplakh"
# monthly_data = monthly_data.reset_index(convert(monthly_data.index))
# + id="rRbK6iX9rbSb"
monthly_data.index = calendar_months
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="YkvP_chgr8g3" outputId="2b5016bc-8c37-4bf5-beff-6061a7bb23bb"
monthly_data
# + id="s1zFoGPZvAOP"
x_series = monthly_data.index
y_series = monthly_data['Temp (C)']
# + id="HdgXvmdWr-eN" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="6d7d908e-ea66-48b6-868a-39a4dced71d6"
plt.plot(x_series, y_series)
plt.title('Temperature Trend, 2012')
plt.xlabel('Month')
plt.xticks(rotation = 90)
plt.ylabel('Temp (C)')
plt.show();
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="Ymp5w23fujTJ" outputId="9eca78a0-9145-44f2-b2f7-fad9ba94df8b"
def bar_plot():
weekly_data = df1.groupby(df1.index.dayofweek).mean()
plt.bar(weekly_data.index, weekly_data['Visibility (km)'])
plt.title('Visibility by week,2012')
plt.xlabel('Day of week')
plt.ylabel('Visibility (km)')
plt.xticks(weekly_data.index, calendar.day_abbr, rotation = 45)
plt.show();
bar_plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="nWDn_W1gs-nl" outputId="ba3a3064-8166-4390-a549-313236b5dbe2"
# Sample histogram
x = np.arange(0, 10, 0.1)
y1 = (((x - 3) ** 3 ) - 100) + np.random.randint(-20, 20, size=len(x))
plt.hist(y1)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="XOMB62sav4Bh" outputId="31905ecf-5203-4443-9a1a-512db5dffc91"
print(df1['Wind Spd (km/h)'].head())
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="7Z6qt2EgvQGQ" outputId="ea8d177f-5608-4d33-aa0b-3dca6e50a482"
# Find the distribution of values in the Wind Speed column
def hist_plot():
x = df1['Wind Spd (km/h)']
plt.title('Wind Speed Count,2012')
plt.xlabel('Wind Spd (km/h)')
plt.ylabel('Count')
plt.hist(x)
plt.show();
hist_plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="XlKPf8JVwXp7" outputId="64a4e457-ace2-40bd-fdf1-36cad21a9274"
# Sample boxplot
x= np.arange(0, 10, 0.1)
y = np.exp(x)
plt.boxplot(y)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="1ol8oq3oyQj3" outputId="70e06618-fb47-4fb8-b884-84e308be292f"
def box_plot():
x = df1['Wind Spd (km/h)']
plt.title('Wind Speed Count,2012')
plt.ylabel('Wind Spd (km/h)')
plt.boxplot(x)
plt.show();
box_plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="xVDMqkpmYNCQ" outputId="5cfa5aeb-053b-4c66-9962-bf239a5d2718"
# Sample scatter plot
x= np.arange(0, 10, 0.1)
y1 = (((x - 3) ** 3 ) - 100) + np.random.randint(-20, 20, size=len(x))
y2 = (((3 - x) ** 3 ) + 50) + np.random.randint(-20, 20, size=len(x))
plt.scatter(x, y1, c='r')
plt.scatter(x, y2, c='b')
plt.show()
# + id="SCwUnajWdoR_"
# Are the temperature and pressure correlated, according to the data in the
# month of January? Look at it by generating a scatter plot
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="hp7Hz4Q4cLyo" outputId="76606571-78f3-4e6b-b4ef-803d89a7fb9f"
jan_df = df1['2012-01']
y1 = jan_df['Temp (C)']
y2 = jan_df['Stn Press (kPa)']
def scatter_plot():
plt.scatter(y1, y2)
plt.show();
scatter_plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="xOnmw5XUdcN1" outputId="ea7d41d0-48e6-4e0f-e198-57d23f65b426"
fig, ax = plt.subplots()
ax.plot(x, x**2, 'b.-') # blue line with dots
ax.plot(x, x**2.5, 'g--') # green dashed line
ax.plot(x, x**3, c='r') # red line color
fig.show();
# + id="k8VMSvyVdqW3"
def two_plots():
x = np.array([0, 1, 2, 3, 4, 5])
y = x ** 2
# Create Figure (empty canvas)
fig = plt.figure()
# Add set of axes to figure
axes1 = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # main axes
axes2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
# (0.2x left, 0.5x bottom) to (0.4x width, 0.3x height)
# Larger Figure Axes 1
axes1.plot(x, y, 'b')
axes1.set_xlabel('X_label_axes1')
axes1.set_ylabel('Y_label_axes1')
axes1.set_title('Axes 1 Title')
# Insert Figure Axes 2
axes2.plot(y, x, 'r')
axes2.set_xlabel('X_label_axes2')
axes2.set_ylabel('Y_label_axes2')
axes2.set_title('Axes 2 Title');
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="ro6hYVpXhKtB" outputId="c4bfed03-84d9-46c6-89fd-f05654b73a35"
two_plots()
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="RFcI81JyhMOM" outputId="fa8db8ae-64fa-4423-b649-f1a6658d3884"
# Canvas of 2 by 2 subplots
fig, axes = plt.subplots(nrows=2, ncols=2)
# axes is an array of shape (2, 2)
# + id="KZLqzB5vh0qA"
def sub_plots_example():
y = x ** 2
fig, axes = plt.subplots(2, 2)
axes[0,0].plot(y, c='r')
axes[1,1].plot(y, 'g--')
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="w6bwck57iuVR" outputId="9fb79d73-47bf-4afe-d841-74b5255bbb48"
sub_plots_example()
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="yrgBeCx8iwCG" outputId="54404d70-5d62-4a98-abb3-e22149a7da2b"
import seaborn as sns
sns.distplot(df1['Wind Spd (km/h)'], bins = 25);
# + id="HBoqOKZGsi__"
| Mahfuzur_rahman_VISUALIZATION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.7
# language: python
# name: py3.7
# ---
# # Exercise 8.02
# Import the required Libraries
import numpy as np
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.applications.vgg16 import preprocess_input
# Initiate the Model
classifier = VGG16()
classifier.summary()
# Load the Image
new_image = image.load_img('../Data/Prediction/stick_insect.jpg', target_size=(224, 224))
new_image
# Change the image to array
transformed_image = image.img_to_array(new_image)
transformed_image.shape
# Expand the tranfromed image with 4th Dimension
transformed_image = np.expand_dims(transformed_image, axis=0)
transformed_image.shape
# Preprocess the Image
transformed_image = preprocess_input(transformed_image)
transformed_image
# Create a predictor variable
y_pred = classifier.predict(transformed_image)
y_pred
# Check the shape of the array
y_pred.shape
# Make the predictions
from keras.applications.vgg16 import decode_predictions
decode_predictions(y_pred, top=5)
# Make the predictions in readable form
label = decode_predictions(y_pred)
# retrieve the most likely result, i.e. highest probability
decoded_label = label[0][0]
# print the classification
print('%s (%.2f%%)' % (decoded_label[1], decoded_label[2]*100 ))
| Chapter08/Exercise8.02/Exercise8_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib Graph Example
#Library import
import matplotlib.pyplot as plt
import numpy as np
# x and y data
division_name=['Barisal','Chittagong','Dhaka','Khulna','Sylhet','Rangpur','Mymensingh','Rajshahi']
population=[12.32,34.56,33.10,78.00,20.21,10.15,23.43,61.22]
age=[65,54,77,33,23,67,88,33]
index=np.arange(len(division_name))
#First Graph
figure=plt.figure(facecolor='c',figsize=(10,6))
figure.patch.set_alpha(.6)
plt.plot(division_name,population)
plt.title('Popoulation Graph of Bangladesh',style='italic',color='b',fontsize='14',fontweight='bold')
plt.xlabel('<----- Division Name ----->',style='italic',color='r',fontsize='12',fontweight='bold')
plt.ylabel('<---- Population in Million ---->',style='italic',color='g',fontsize='12',fontweight='bold')
plt.xticks(color='r')
plt.yticks(color='g')
plt.grid(linestyle='-',linewidth=.3,color='r')
plt.show()
#Second Graph- Pie
figure=plt.figure(facecolor='c',figsize=(10,6))
figure.patch.set_alpha(.3)
plt.title('Popoulation of Bangladesh',style='italic',color='b',fontsize='14',fontweight='bold')
plt.xlabel('Pie Chart',style='italic',color='r',fontsize='12',fontweight='bold')
plt.pie(population,explode=[0,0,0,0,0,0,0,0],autopct='%1.1f%%', radius=1.1,labels=division_name)
plt.legend(loc='lower left',bbox_to_anchor=(1.2,.3))
plt.show()
#Third Graph- Bar
figure=plt.figure(facecolor='c',figsize=(10,6))
figure.patch.set_alpha(.3)
plt.title('Popoulation of Bangladesh',style='italic',color='b',fontsize='14',fontweight='bold')
plt.xlabel('<----- Division Name ----->',style='italic',color='r',fontsize='12',fontweight='bold')
plt.ylabel('<---- Population in Million ---->',style='italic',color='g',fontsize='12',fontweight='bold')
plt.xticks(color='r')
plt.yticks(color='g')
plt.grid(linestyle='-',linewidth=.2,color='b')
plt.bar(index,age,width=.5,color='g',label='Average Age')
plt.bar(index+.5,population,width=.5,color='r',label='Population')
plt.legend()
plt.show()
#Third Graph- Bar
figure=plt.figure(facecolor='c',figsize=(10,6))
figure.patch.set_alpha(.3)
plt.title('Popoulation of Bangladesh',style='italic',color='b',fontsize='14',fontweight='bold')
plt.xlabel('<----- Division Name ----->',style='italic',color='r',fontsize='12',fontweight='bold')
plt.ylabel('<---- Population in Million ---->',style='italic',color='g',fontsize='12',fontweight='bold')
plt.xticks(color='r')
plt.yticks(color='g')
plt.grid(linestyle='-',linewidth=.2,color='b')
plt.bar(index,population,width=.5,color='g',label='Popupation')
plt.bar(index+.5,age,width=.5,color='y',label='Age')
plt.legend()
plt.show()
# # Data Distribution
# +
import numpy
from scipy import stats
ages = [5,31,43,48,50,41,7,11,15,12,12,12,12,12,39,80,82,32,2,8,6,25,36,27,61,31]
x = numpy.random.normal(5.0,1.0,25000)
mean= numpy.mean(x)
median = numpy.median(x)
mode = stats.mode(ages)
print(mode)
print(median)
print(mean)
print(x)
# -
plt.hist(x,100)
plt.show()
# # Machine Learning - Scatter Plot
# A scatter plot is a diagram where each value in the data set is represented by a dot. The Matplotlib module has a method for drawing scatter plots, it needs two arrays of the same length, one for the values of the x-axis, and one for the values of the y-axis:
x_car_age = [5,7,8,7,2,17,2,9,4,11,12,9,6]
y_car_speed = [99,86,87,88,111,86,103,87,94,78,77,85,86]
plt.scatter(x_car_age,y_car_speed)
plt.xlabel('Car Age', color= 'red')
plt.ylabel('Car Speed',color= 'green')
plt.show()
# What we can read from the diagram is that the two fastest cars were both 2 years old, and the slowest car was 12 years old.
#
# Note: It seems that the newer the car, the faster it drives, but that could be a coincidence, after all we only registered 13 cars.
# ## Random Data Distributions
x_car_age = np.random.normal(5.0,1.0,1000)
y_car_speed = np.random.normal(10.0,2.0,1000)
# mean, Standard Deviation, Range
plt.scatter(x_car_age,y_car_speed)
plt.show()
# We can see that the dots are concentrated around the value 5 on the x-axis, and 10 on the y-axis.
#
# We can also see that the spread is wider on the y-axis than on the x-axis.
# # Machine Learning - Linear Regression
# In Machine Learning, and in statistical modeling, that relationship is used to predict the outcome of future events.Linear regression uses the relationship between the data-points to draw a straight line through all them.
#
# This line can be used to predict future values.In Machine Learning, predicting the future is very important.
x_car_age = [5,7,8,7,2,17,2,9,4,11,12,9,6]
y_car_speed = [99,86,87,88,111,86,103,87,94,78,77,85,86]
slope, intercept, r, p , std_err = stats.linregress(x_car_age,y_car_speed)
def myFunc(x_car_age):
return slope * x_car_age + intercept
model = list(map(myFunc,x_car_age))
plt.scatter(x_car_age, y_car_speed)
plt.plot(x_car_age,model)
plt.show()
# ## R for Relationship
# It is important to know how the relationship between the values of the x-axis and the values of the y-axis is, if there are no relationship the linear regression can not be used to predict anything.
#
# This relationship - the coefficient of correlation - is called r.
#
# The r value ranges from 0 to 1, where 0 means no relationship, and 1 means 100% related.
#
# Python and the Scipy module will compute this value for you, all you have to do is feed it with the x and y values.
print(r)
# Note: The result -0.76 shows that there is a relationship, not perfect, but it indicates that we could use linear regression in future predictions.
# #### Predict Future Values
# Now we can use the information we have gathered to predict future values.
# Example: Let us try to predict the speed of a 10 years old car.
# To do so, we need the same myfunc() function from the example above:
speed = myFunc(10)
print(speed)
plt.show()
# ### Bad Fit?
# Let us create an example where linear regression would not be the best method to predict future values.
x_car_age = [89,43,36,36,95,10,66,34,38,20,26,29,48,64,6,5,36,66,72,40]
y_car_speed = [21,46,3,35,67,95,53,72,58,10,26,34,90,33,38,20,56,2,47,15]
slope, intercept, r, p , std_err = stats.linregress(x_car_age,y_car_speed)
def myFunc(x_car_age):
return slope * x_car_age + intercept
model = list(map(myFunc,x_car_age))
plt.scatter(x_car_age, y_car_speed)
plt.plot(x_car_age,model)
plt.show()
print(r)
# # Machine Learning - Polynomial Regression
# In the example below, we have registered 18 cars as they were passing a certain tollbooth.
# We have registered the car's speed, and the time of day (hour) the passing occurred.
#
# The x-axis represents the hours of the day and the y-axis represents the speed:
x_drive_day = [1,2,3,5,6,7,8,9,10,12,13,14,15,16,18,19,21,22]
y_speed = [100,90,80,60,60,55,60,65,70,70,75,76,78,79,90,99,99,100]
plt.scatter(x_drive_day,y_speed)
plt.show()
model = np.poly1d(np.polyfit(x_drive_day,y_speed,3))
#specify how the line will display, we start at position 1, and end at position 22:
line = np.linspace(1,22,100)
plt.scatter(x_drive_day,y_speed)
plt.plot(line,model(line))
plt.show()
# ## R-Squared
# Python and the Sklearn module will compute this value for you, all you have to do is feed it with the x and y arrays:
from sklearn.metrics import r2_score
print(r2_score(y_speed,model(x_drive_day)))
# Note: The result 0.94 shows that there is a very good relationship, and we can use polynomial regression in future predictions.
# #### Predict Future Values
speed = model(17)
print(speed)
# ## Bad Fit?
# Let us create an example where polynomial regression would not be the best method to predict future values.
x_drive_day = [89,43,36,36,95,10,66,34,38,20,26,29,48,64,6,5,36,66,72,40]
y_speed = [21,46,3,35,67,95,53,72,58,10,26,34,90,33,38,20,56,2,47,15]
model = np.poly1d(np.polyfit(x_drive_day,y_speed,3))
#specify how the line will display, we start at position 1, and end at position 22:
line = np.linspace(2,95,100)
plt.scatter(x_drive_day,y_speed)
plt.plot(line,model(line))
plt.show()
print(r2_score(y_speed,model(x_drive_day)))
# ## Multiple Regression
# Multiple regression is like linear regression, but with more than one independent value, meaning that we try to predict a value based on two or more variables.
#
# We can predict the CO2 emission of a car based on the size of the engine, but with multiple regression we can throw in more variables, like the weight of the car, to make the prediction more accurate.
import pandas as pd
df = pd.read_csv('cars.csv')
#df.head(8)
df
X = df[['Weight', 'Volume']]
y = df['CO2']
from sklearn import linear_model
linreg = linear_model.LinearRegression()
linreg.fit(X,y)
#predict the CO2 emission of a car where the weight is 2300kg, and the volume is 1300cm3:
predCO2 = linreg.predict([[2300,1300]])
print(predCO2)
# We have predicted that a car with 1.3 liter engine, and a weight of 2300 kg, will release approximately 107 grams of CO2 for every kilometer it drives.
# In this case, we can ask for the coefficient value of weight against CO2, and for volume against CO2. The answer(s) we get tells us what would happen if we increase, or decrease, one of the independent values.
print(linreg.coef_)
# ## Result Explained
# The result array represents the coefficient values of weight and volume.
#
# Weight: 0.00755095
# Volume: 0.00780526
#
# These values tell us that if the weight increase by 1kg, the CO2 emission increases by 0.00755095g.
#
# And if the engine size (Volume) increases by 1 cm3, the CO2 emission increases by 0.00780526 g.
#
# I think that is a fair guess, but let test it!
#
# We have already predicted that if a car with a 1300cm3 engine weighs 2300kg, the CO2 emission will be approximately 107g.
#
# What if we increase the weight with 1000kg?
#weight from 2300 to 3300:
predCO2 = linreg.predict([[3300,1300]])
print(predCO2)
# We have predicted that a car with 1.3 liter engine, and a weight of 3300 kg, will release approximately 115 grams of CO2 for every kilometer it drives.
#
# Which shows that the coefficient of 0.00755095 is correct:
#
# 107.2087328 + (1000 * 0.00755095) = 114.75968
# # Machine Learning - Train/Test
# #### What is Train/Test
# Train/Test is a method to measure the accuracy of your model.
#
# It is called Train/Test because you split the the data set into two sets: a training set and a testing set.
#
# 80% for training, and 20% for testing.
#
# You train the model using the training set.
#
# You test the model using the testing set.
#
# Train the model means create the model.
#
# Test the model means test the accuracy of the model.
np.random.seed(2)
x = np.random.normal(3,1,100)
y = np.random.normal(150,40,100)/x
plt.scatter(x,y)
plt.xlabel("number of minutes before making a purchase")
plt.ylabel("amount of money spent on the purchase")
plt.show()
# ### Split Into Train/Test
# The training set should be a random selection of 80% of the original data.
# The testing set should be the remaining 20%.
train_x = x[:80]
train_y = y[:80]
test_x = x[80: ]
test_y = y[80: ]
plt.scatter(train_x, train_y)
plt.show()
plt.scatter(test_x, test_y)
plt.show()
# # Fit the Data Set
model = np.poly1d(np.polyfit(train_x,train_y,4))
line = np.linspace(0,6,100)
plt.scatter(train_x,train_y)
plt.plot(line,model(line))
plt.show()
print(r2_score(train_y,model(train_x)))
# Note: The result 0.799 shows that there is a OK relationship.
print(r2_score(test_y,model(test_x)))
# Note: The result 0.809 shows that the model fits the testing set as well, and we are confident that we can use the model to predict future values.
# ### Predict Values: How much money will a buying customer spend, if she or he stays in the shop for 5 minutes?
print(model(5))
# # Decision Tree
# Luckily our example person has registered every time there was a comedy show in town, and registered some information about the comedian, and also registered if he/she went or not.
# +
import pandas
from sklearn import tree
from sklearn.tree import DecisionTreeClassifier
import matplotlib.pyplot as plt
import matplotlib.image as pltimg
import pydotplus
df = pandas.read_csv("shows.csv")
print(df)
# -
# To make a decision tree, all data has to be numerical.
#
# We have to convert the non numerical columns 'Nationality' and 'Go' into numerical values.
#
# Pandas has a map() method that takes a dictionary with information on how to convert the values.
#
# {'UK': 0, 'USA': 1, 'N': 2}
#
# Means convert the values 'UK' to 0, 'USA' to 1, and 'N' to 2.
d = {'UK' :0, 'USA':1, 'N':2}
df['Nationality'] = df['Nationality'].map(d)
d = {'YES':1, 'NO':0}
df['Go'] = df['Go'].map(d)
print(df)
# Then we have to separate the feature columns from the target column.
#
# The feature columns are the columns that we try to predict from, and the target column is the column with the values we try to predict.
features = ['Age', 'Experience', 'Rank', 'Nationality']
X = df[features]
y=df['Go']
print(X)
print(y)
from IPython.display import Image
dtree = DecisionTreeClassifier()
dtree = dtree.fit(X, y)
data = tree.export_graphviz(dtree, out_file=None, feature_names=features)
graph = pydotplus.graph_from_dot_data(data)
graph.write_png('mydecisiontree.png')
| Matplotlib Graph Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="7Oqedsmht04x" executionInfo={"status": "ok", "timestamp": 1610323105429, "user_tz": 300, "elapsed": 1217, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="b80ffeca-c5d3-47b2-a494-d591b1decdb2"
# %cd /content/drive/My Drive/Colab Notebooks/regaetton_songs_nlp
# + id="hpdLsevet5oY" executionInfo={"status": "ok", "timestamp": 1610323105768, "user_tz": 300, "elapsed": 1546, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
normalized_eval_path = '/content/drive/MyDrive/Colab Notebooks/regaetton_songs_nlp/data/normalized_eval_lyrics.csv'
split_train_path = '/content/drive/MyDrive/Colab Notebooks/regaetton_songs_nlp/data/normalized_train_split.csv'
scores_path = '/content/drive/MyDrive/Colab Notebooks/regaetton_songs_nlp/data/scores.csv'
# + colab={"base_uri": "https://localhost:8080/"} id="LGQZOEVvuAYt" executionInfo={"status": "ok", "timestamp": 1610323106152, "user_tz": 300, "elapsed": 1923, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="1a6876e1-1e8d-4297-9248-1b6fef90ab4b"
import nltk
import pandas as pd
from nltk import word_tokenize
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import accuracy_score, recall_score
nltk.download('punkt')
def add_score(df, model, parameters, accuracy, recall):
row = {'model': model, 'parameters': parameters,
'accuracy': accuracy, 'recall': recall}
df = df.append(row, ignore_index=True)
return df
# + id="lBQbqEVBuF1V" executionInfo={"status": "ok", "timestamp": 1610323162681, "user_tz": 300, "elapsed": 1973, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
eval = pd.read_csv(normalized_eval_path)
train = pd.read_csv(split_train_path)
scores = pd.read_csv(scores_path)
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="qP2dpMWQud_p" executionInfo={"status": "ok", "timestamp": 1610323164179, "user_tz": 300, "elapsed": 1055, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="336127d9-c60e-4364-d405-2c7e5f631d94"
scores.head()
# + [markdown] id="QWAMy_D3uJad"
# # Create model
#
# + id="k1fOH3yruNRc" executionInfo={"status": "ok", "timestamp": 1610323107564, "user_tz": 300, "elapsed": 3312, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
X_train, X_valid, y_train, y_valid = train_test_split(train.lyrics.values, train.sexual_content.values,
stratify=train.sexual_content.values, random_state=10,
test_size=0.15, shuffle=True)
# + id="4oXup8qjuR99" executionInfo={"status": "ok", "timestamp": 1610323296323, "user_tz": 300, "elapsed": 11148, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
count_vect = CountVectorizer(tokenizer=lambda x: word_tokenize(x, language='spanish'), encoding='utf-8', ngram_range=(1, 3))
X_train_ctv = count_vect.fit_transform(X_train)
X_valid_ctv = count_vect.transform(X_valid)
# + colab={"base_uri": "https://localhost:8080/"} id="_wprYqHsvZy4" executionInfo={"status": "ok", "timestamp": 1610323331280, "user_tz": 300, "elapsed": 21031, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="635038e6-fc91-49d7-91d8-62ac50ed3000"
# Fitting a simple Logistic Regression on Counts
lgr_parameters = {'C': 1.0}
clf = LogisticRegression(C=1.0)
clf.fit(X_train_ctv, y_train)
predictions = clf.predict(X_valid_ctv)
print(f'Loggistic regression accuracy {accuracy_score(predictions, y_valid)}')
print(f'Loggistic regression recall {recall_score(predictions, y_valid)}')
# + colab={"base_uri": "https://localhost:8080/"} id="fv6_jR7Tv7ZJ" executionInfo={"status": "ok", "timestamp": 1610323362005, "user_tz": 300, "elapsed": 1378, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="8d2a582e-f700-4fd0-faed-f3bda094adb7"
X_eval_ctv = count_vect.transform(eval.lyrics.values)
eval_preds = clf.predict(X_eval_ctv)
lgr_accuracy = accuracy_score(eval_preds, eval.sexual_content.values)
lgr_recall = recall_score(eval_preds, eval.sexual_content.values)
print(f'Loggistic regression accuracy in hand-labeled lyrics {lgr_accuracy}')
print(f'Loggistic regression recall in hand-labeled lyrics {lgr_recall}')
# + id="uF78kkSSwHsp" executionInfo={"status": "ok", "timestamp": 1610323387541, "user_tz": 300, "elapsed": 971, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
scores = add_score(scores, 'logistic regression - Bag of n-grams', lgr_parameters, lgr_accuracy, lgr_recall)
# + colab={"base_uri": "https://localhost:8080/"} id="mNNToKI9wOBx" executionInfo={"status": "ok", "timestamp": 1610323430382, "user_tz": 300, "elapsed": 1428, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="1605d453-4041-4643-d4ea-932bbf92a91d"
# Fitting a simple Naive Bayes
clf = MultinomialNB()
clf.fit(X_train_ctv, y_train)
predictions = clf.predict(X_valid_ctv)
print(f'Loggistic regression accuracy {accuracy_score(predictions, y_valid)}')
print(f'Loggistic regression recall {recall_score(predictions, y_valid)}')
X_eval_ctv = count_vect.transform(eval.lyrics.values)
eval_preds = clf.predict(X_eval_ctv)
naive_accuracy = accuracy_score(eval_preds, eval.sexual_content.values)
naive_recall = recall_score(eval_preds, eval.sexual_content.values)
print(f'Loggistic regression accuracy in hand-labeled lyrics {naive_accuracy}')
print(f'Loggistic regression recall in hand-labeled lyrics {naive_recall}')
# + id="Lgu_X_drwYX8" executionInfo={"status": "ok", "timestamp": 1610323850761, "user_tz": 300, "elapsed": 1222, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
scores = add_score(scores, 'naive bayes - Bag of ngrams', '', naive_accuracy, naive_recall)
scores.to_csv(scores_path, index=False)
# + [markdown] id="BUtuim98wf1B"
# # Let's use Tfidf instead of count vectorizer
# + id="u0bHLywQwpKw" executionInfo={"status": "ok", "timestamp": 1610323592701, "user_tz": 300, "elapsed": 939, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
tfv = TfidfVectorizer(tokenizer=lambda x: word_tokenize(x, language='spanish'),min_df=3, max_features=None,
ngram_range=(1, 3), use_idf=1,smooth_idf=1,sublinear_tf=1)
# + id="LGqn-uD0xAIX" executionInfo={"status": "ok", "timestamp": 1610323630923, "user_tz": 300, "elapsed": 11026, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
X_train_tfv = tfv.fit_transform(X_train)
X_valid_tfv = tfv.transform(X_valid)
# + colab={"base_uri": "https://localhost:8080/"} id="ow7gLshLxG_3" executionInfo={"status": "ok", "timestamp": 1610323690860, "user_tz": 300, "elapsed": 1379, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="9367bb74-6a83-4ce7-a647-d1ae2d96d47c"
# Fitting a simple Logistic Regression on tfidf
lgr_parameters = {'C': 1.0}
clf = LogisticRegression(C=1.0)
clf.fit(X_train_tfv, y_train)
predictions = clf.predict(X_valid_tfv)
print(f'Loggistic regression accuracy {accuracy_score(predictions, y_valid)}')
print(f'Loggistic regression recall {recall_score(predictions, y_valid)}')
# + colab={"base_uri": "https://localhost:8080/"} id="lb-vQuivxSuv" executionInfo={"status": "ok", "timestamp": 1610323748404, "user_tz": 300, "elapsed": 1311, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="26356bf1-143a-4bca-b291-fb1ffb4b28f8"
X_eval_tfv = tfv.transform(eval.lyrics.values)
eval_preds = clf.predict(X_eval_tfv)
lgr_accuracy = accuracy_score(eval_preds, eval.sexual_content.values)
lgr_recall = recall_score(eval_preds, eval.sexual_content.values)
print(f'Loggistic regression accuracy in hand-labeled lyrics {lgr_accuracy}')
print(f'Loggistic regression recall in hand-labeled lyrics {lgr_recall}')
# + id="Zr0HeeAExi-3" executionInfo={"status": "ok", "timestamp": 1610323785514, "user_tz": 300, "elapsed": 1341, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
scores = add_score(scores, 'logistic regression - Bag of n-grams - tfidf', lgr_parameters, lgr_accuracy, lgr_recall)
# + colab={"base_uri": "https://localhost:8080/"} id="LC62saunxvGG" executionInfo={"status": "ok", "timestamp": 1610323841350, "user_tz": 300, "elapsed": 984, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="1b20f30a-0edb-408b-bf84-106ecce3a43e"
# Fitting a simple Naive Bayes
clf = MultinomialNB()
clf.fit(X_train_tfv, y_train)
predictions = clf.predict(X_valid_tfv)
print(f'Loggistic regression accuracy {accuracy_score(predictions, y_valid)}')
print(f'Loggistic regression recall {recall_score(predictions, y_valid)}')
eval_preds = clf.predict(X_eval_tfv)
naive_accuracy = accuracy_score(eval_preds, eval.sexual_content.values)
naive_recall = recall_score(eval_preds, eval.sexual_content.values)
print(f'Loggistic regression accuracy in hand-labeled lyrics {naive_accuracy}')
print(f'Loggistic regression recall in hand-labeled lyrics {naive_recall}')
# + id="viT6jzTkx80l" executionInfo={"status": "ok", "timestamp": 1610323884756, "user_tz": 300, "elapsed": 1103, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
scores = add_score(scores, 'naive bayes - Bag of ngrams - tfidf', '', naive_accuracy, naive_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="K6Vs38LfyHY9" executionInfo={"status": "ok", "timestamp": 1610323896675, "user_tz": 300, "elapsed": 875, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="5c176a48-6c41-498c-cee7-790da02208ce"
scores.head()
# + [markdown] id="rNGV-rgwyJQs"
# # Trying SVM
# + id="1Z5zs20AyWx8" executionInfo={"status": "ok", "timestamp": 1610324158829, "user_tz": 300, "elapsed": 1112, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
from sklearn.svm import SVC
from sklearn.decomposition import TruncatedSVD
from sklearn.preprocessing import StandardScaler
# + id="NMuuufJzyduU" executionInfo={"status": "ok", "timestamp": 1610324168338, "user_tz": 300, "elapsed": 6469, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
n_components = 120
svd = TruncatedSVD(n_components=n_components)
svd.fit(X_train_tfv)
X_train_svd = svd.transform(X_train_tfv)
X_valid_svd = svd.transform(X_valid_tfv)
# Scale the data obtained from SVD. Renaming variable to reuse without scaling.
scl = StandardScaler()
scl.fit(X_train_svd)
X_train_svd_scaled = scl.transform(X_train_svd)
X_valid_svd_scaled = scl.transform(X_valid_svd)
# + id="_5cyBXXhzCQ8" executionInfo={"status": "ok", "timestamp": 1610324283411, "user_tz": 300, "elapsed": 5389, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
# Fitting a simple SVM
svm_parameters = {'C':1.0}
clf = SVC(C=1.0)
clf.fit(X_train_svd_scaled, y_train)
predictions = clf.predict(X_valid_svd_scaled)
# + colab={"base_uri": "https://localhost:8080/"} id="8XKtotFFzj1r" executionInfo={"status": "ok", "timestamp": 1610324336498, "user_tz": 300, "elapsed": 907, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="4db61e4a-f008-4704-d0bd-b3cfd116bd91"
print(f'SVM accuracy {accuracy_score(predictions, y_valid)}')
print(f'SVM recall {recall_score(predictions, y_valid)}')
# + id="_tM1vRJYz1uC" executionInfo={"status": "ok", "timestamp": 1610324533247, "user_tz": 300, "elapsed": 1168, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
X_eval_svd = svd.transform(X_eval_tfv)
X_eval_svd_scaled = scl.transform(X_eval_svd)
predictions = clf.predict(X_eval_svd_scaled)
svm_accuracy = accuracy_score(predictions, eval.sexual_content.values)
svm_recall = recall_score(predictions, eval.sexual_content.values)
# + colab={"base_uri": "https://localhost:8080/"} id="1WNWZmc10Iyj" executionInfo={"status": "ok", "timestamp": 1610324534846, "user_tz": 300, "elapsed": 929, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="a6910018-b5da-49cc-d476-96f1b71e5b37"
print(f'SVM accuracy in hand-labeled lyrics {naive_accuracy}')
print(f'SVM regression recall in hand-labeled lyrics {naive_recall}')
# + id="QIqxGR6X0mJK" executionInfo={"status": "ok", "timestamp": 1610324612633, "user_tz": 300, "elapsed": 1022, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
scores = add_score(scores, 'svm', svm_parameters, svm_accuracy, svm_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="D_r_6XcR05Gy" executionInfo={"status": "ok", "timestamp": 1610324717167, "user_tz": 300, "elapsed": 991, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="df10e1e5-b1ee-47df-b946-5c5fa3f80ce2"
scores.head(10)
# + [markdown] id="7nbiSwx71SpA"
# # Stronger model
#
# Now that we have an idea of how well can we do in our eval dataset with models that are consider simple, let's try a stronger model now using gridsearch as well to explore more possibilities.
# + id="bGmgfK839uBx" executionInfo={"status": "ok", "timestamp": 1610329009510, "user_tz": 300, "elapsed": 952, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
import numpy as np
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import GridSearchCV
# + id="5OeRaElIBApN" executionInfo={"status": "ok", "timestamp": 1610329161441, "user_tz": 300, "elapsed": 996, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
params = {'learning_rate': [0.0001, 0.001, 0.01, 0.025, 0.05, 0.075, 0.1, 0.15, 0.2],
'n_estimators': [10],
'max_features': ["log2","sqrt"],
'max_depth':[3,5,8, 10, 15],
'criterion':['friedman_mse', 'mae'],
}
clf = RandomizedSearchCV(GradientBoostingClassifier(), params, cv=5, n_jobs=-1)
# + [markdown] id="4XD5bMjeLJtp"
# # Train using the Tfidf features
# + colab={"base_uri": "https://localhost:8080/"} id="RwkPfSoTDGwe" executionInfo={"status": "ok", "timestamp": 1610330912514, "user_tz": 300, "elapsed": 1750404, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="0529bf9d-7614-4409-dfcb-97084bf83a51"
clf.fit(X_train_tfv, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="-epJSfaZDaxX" executionInfo={"status": "ok", "timestamp": 1610330938949, "user_tz": 300, "elapsed": 1030, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="b6f4a452-e34a-4217-b24e-ef0335b72bb2"
print(f'Best params: \n{clf.best_params_}')
best_params = clf.best_params_
predictions = clf.predict(X_valid_tfv)
print(f'Gradient Boosting accuracy {accuracy_score(predictions, y_valid)}')
print(f'Gradient Boosting recall {recall_score(predictions, y_valid)}')
# + id="xXwxKIaSKQC6" executionInfo={"status": "ok", "timestamp": 1610330967862, "user_tz": 300, "elapsed": 893, "user": {"displayName": "<NAME>uti\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
predictions = clf.predict(X_eval_tfv)
gb_accuracy = accuracy_score(predictions, eval.sexual_content.values)
gb_recall = recall_score(predictions, eval.sexual_content.values)
scores = add_score(scores, 'Gradient Boosting on Tfidf', best_params, gb_accuracy, gb_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="NrOygaAZNNl2" executionInfo={"status": "ok", "timestamp": 1610331045106, "user_tz": 300, "elapsed": 898, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="1427725e-0f81-4979-d4c7-a107e20d52e1"
print(f'Gradient Boosting accuracy in hand-labeled lyrics {gb_accuracy}')
print(f'Gradient Boosting recall in hand-labeled lyrics {gb_recall}')
# + [markdown] id="vSpUSz1hLORw"
# # Train using the count vectorizer features
# + colab={"base_uri": "https://localhost:8080/"} id="8ypRTO1SJ93U" executionInfo={"status": "ok", "timestamp": 1610335176359, "user_tz": 300, "elapsed": 4116415, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="2c309f4b-401b-402a-8f7f-f2fa78bb7988"
clf.fit(X_train_ctv, y_train)
print(f'Best params: \n{clf.best_params_}')
best_params = clf.best_params_
predictions = clf.predict(X_valid_ctv)
print(f'Gradient Boosting accuracy {accuracy_score(predictions, y_valid)}')
print(f'Gradient Boosting recall {recall_score(predictions, y_valid)}')
# + id="YR9ymUkSKheh" executionInfo={"status": "ok", "timestamp": 1610335185589, "user_tz": 300, "elapsed": 1001, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
predictions = clf.predict(X_eval_ctv)
gb_accuracy = accuracy_score(predictions, eval.sexual_content.values)
gb_recall = recall_score(predictions, eval.sexual_content.values)
scores = add_score(scores, 'Gradient Boosting on count vectorizer', best_params, gb_accuracy, gb_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="x0wx_yYtOovp" executionInfo={"status": "ok", "timestamp": 1610335186220, "user_tz": 300, "elapsed": 1574, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="c6da2eed-9e7b-4271-8c69-d3e930408678"
print(f'Gradient Boosting accuracy in hand-labeled lyrics {gb_accuracy}')
print(f'Gradient Boosting recall in hand-labeled lyrics {gb_recall}')
# + [markdown] id="Hd_Wh6o4LIHR"
# # Train using the Dimensionality reduction using truncated SVD
# + colab={"base_uri": "https://localhost:8080/"} id="hXvoslQwLIWq" executionInfo={"status": "ok", "timestamp": 1610335973026, "user_tz": 300, "elapsed": 278278, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="7e1e8700-f498-4a3a-a54d-833b87992286"
clf.fit(X_train_svd, y_train)
print(f'Best params: \n{clf.best_params_}')
best_params = clf.best_params_
predictions = clf.predict(X_valid_svd)
print(f'Gradient Boosting accuracy {accuracy_score(predictions, y_valid)}')
print(f'Gradient Boosting recall {recall_score(predictions, y_valid)}')
# + id="4AqLEMjlOykX" executionInfo={"status": "ok", "timestamp": 1610335973031, "user_tz": 300, "elapsed": 277635, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
predictions = clf.predict(X_eval_svd)
gb_accuracy = accuracy_score(predictions, eval.sexual_content.values)
gb_recall = recall_score(predictions, eval.sexual_content.values)
scores = add_score(scores, 'Gradient Boosting on 120 svd', best_params, gb_accuracy, gb_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="eAGcjW-DO24h" executionInfo={"status": "ok", "timestamp": 1610335973043, "user_tz": 300, "elapsed": 276386, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="7a40a378-360d-475f-ce95-e66d97b3c2df"
print(f'Gradient Boosting accuracy in hand-labeled lyrics {gb_accuracy}')
print(f'Gradient Boosting recall in hand-labeled lyrics {gb_recall}')
# + [markdown] id="Dm8jS0sFO5DT"
# # Now using the scaled version
# + colab={"base_uri": "https://localhost:8080/"} id="YjAcL7jAO7vp" executionInfo={"status": "ok", "timestamp": 1610336155480, "user_tz": 300, "elapsed": 457669, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="d1a9e738-6517-48d0-9531-ec00de7206b5"
clf.fit(X_train_svd_scaled, y_train)
print(f'Best params: \n{clf.best_params_}')
best_params = clf.best_params_
predictions = clf.predict(X_valid_svd_scaled)
print(f'Gradient Boosting accuracy {accuracy_score(predictions, y_valid)}')
print(f'Gradient Boosting recall {recall_score(predictions, y_valid)}')
# + id="7RUdeIZLPBn6" executionInfo={"status": "ok", "timestamp": 1610336155491, "user_tz": 300, "elapsed": 456322, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}}
predictions = clf.predict(X_eval_svd_scaled)
gb_accuracy = accuracy_score(predictions, eval.sexual_content.values)
gb_recall = recall_score(predictions, eval.sexual_content.values)
scores = add_score(scores, 'Gradient Boosting on 120 svd scaled', best_params, gb_accuracy, gb_recall)
scores.to_csv(scores_path, index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="o_91HUWtfL6q" executionInfo={"status": "ok", "timestamp": 1610336158834, "user_tz": 300, "elapsed": 1236, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="2f27a3e7-6518-45e8-b1b7-be1abf894d6a"
print(f'Gradient Boosting accuracy in hand-labeled lyrics {gb_accuracy}')
print(f'Gradient Boosting recall in hand-labeled lyrics {gb_recall}')
# + [markdown] id="jYJt7eaXfRWB"
# as we can see, a more powerful model does not mean always a better performance. Although, we could increase the grid search.
# + colab={"base_uri": "https://localhost:8080/", "height": 426} id="I9gWwE-pibts" executionInfo={"status": "ok", "timestamp": 1610336801969, "user_tz": 300, "elapsed": 1033, "user": {"displayName": "<NAME>\u00e9rrez", "photoUrl": "", "userId": "11854874717100226645"}} outputId="19ff3be8-27f5-4b41-993b-3c7e3fd364d5"
scores.head(20)
# + id="4eQ059hdjTiD"
| models/bag_of_ngrams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import fsolve
from scipy.integrate import quad
from scipy.special import eval_laguerre
import itertools
import copy
from image_matrix_helper import compute_master_list, imshow_list, rgb_map, color_to_rgb, list_to_matrix
import random
import time
nb_start = time.time()
# -
# ## Simulation for General Case of Ligand-Receptor Binding
#
# In this notebook, we simulate a thermal system of particles of various colors binding onto a grid. We have $R$ different types of particles and particle of type $j$ has $n_j$ copies in the system. Particles can exist both on and off the grid and particles of type $j$ have a binding affinity of $\gamma_j$ to the grid. Each particle type also has a collection of "correct" locations on the grid. A particle of type $j$ binds to this correct location with an additional optimal binding affinity factor of $\delta_j$ (i.e., its net affinity to such a site is $\gamma_j \delta_j$). Here we want to use simulations of this system to affirm analytical calculations of the average number of bound particles and the average number of correctly bound particles as functions of temperature.
#
# ### Numerical representations of analytical work
# #### Equations of Large $N$ approximation
#
# In the large $N$ limit, the order parameters for the system can be approximated as
#
# \begin{equation}
# \langle k \rangle = \sum_{j=1}^r \frac{n_j }{\bar{z} \gamma_j +1} \left(\bar{z} \gamma_j + \frac{\displaystyle L_{n_j-1} \left( \bar{\phi}_{j}\right)}{\displaystyle L_{n_j} \left( \bar{\phi}_{j} \right)}\right) \qquad \langle m \rangle = \sum_{j=1}^r \frac{n_j \delta_j}{\delta_j-1} \frac{\displaystyle L_{n_j-1} \left( \bar{\phi}_{j}\right)}{\displaystyle L_{n_j} \left( \bar{\phi}_{j} \right)},
# \end{equation}
# where $\bar{z}$ and $\bar{x}$ are defined as
# \begin{equation}
# \bar{z} = \sum_{j=1}^{R} \frac{n_j}{\bar{z} \gamma_j +1} \left(1- \frac{\displaystyle L_{n_j-1} \left( \bar{\phi}_{j}\right)}{\displaystyle L_{n_j} \left( \bar{\phi}_{j} \right)}\right), \qquad \bar{x} = \sum_{j=1}^{R} n_j\left(1- \frac{\displaystyle L_{n_j-1} \left( \bar{\phi}_{j}\right)}{\displaystyle L_{n_j} \left( \bar{\phi}_{j} \right)}\right)
# \end{equation}
# with
# \begin{equation}
# \bar{\phi}_{j}\equiv \frac{\bar{x}}{1-\delta_j}\left(1+ \frac{1}{\bar{z} \gamma_j}\right).
# \label{eq:phi_def}
# \end{equation}
# and $L_n(x)$ the $n$th Laguerre polynomial.
#
# For these simulations we will take $$\gamma_j = (\beta E_V)^{3/2} e^{-\beta E_j}, \qquad \delta_j = e^{\beta \Delta_j}$$ where $E_V$ is a volumetric Boltzmann factor associated with free particles (e.g., for a point-particle $E_V \equiv h^2/2\pi mV^{2/3}$), and $E_j$ is the binding energy for particles to the grid. We also take where $\Delta_j$ is the binding energy advantage for particles binding to their correct locations in the grid.
# #### Parameter function definitions
# helper function definitions
gamma_func = lambda E0, Ev, T: 4*np.sqrt(2)*np.exp(E0/T)*(Ev/T)**(3/2)
delta_func = lambda Del, T: np.exp(Del/T)
phi_func = lambda x, z, gamma, delta: x*(1+ 1/(z*gamma))/(1-delta)
# #### Equilibrium equations
# +
def constr_func(X, T, E0s, Dels, Evs, Ns):
"""
Equations constraint equations that determine zbar and xbar
"""
x = X[0]
z = X[1]
F = np.ones(2)
R = len(Ns)
gammas_ = gamma_func(E0s,Evs, T)
deltas_ = delta_func(Dels, T)
phis_ = phi_func(x, z, gammas_, deltas_)
F[0] = z- np.sum([Ns[j]/(z*gammas_[j]+1)*(1-eval_laguerre(Ns[j]-1, phis_[j])/eval_laguerre(Ns[j], phis_[j])) for j in range(R)])
F[1] = x- np.sum([Ns[j]*(1-eval_laguerre(Ns[j]-1, phis_[j])/eval_laguerre(Ns[j], phis_[j]) ) for j in range(R)])
return F
def m_avg(T, E0s, Dels, Evs, Ns):
"""
Function that computes m_avg
"""
x, z = fsolve(constr_func, x0 = (50,500), args = (T, E0s, Dels, Evs, Ns))
R = len(Ns)
gammas_ = gamma_func(E0s,Evs, T)
deltas_ = delta_func(Dels, T)
phis_ = phi_func(x, z, gammas_, deltas_)
return np.sum([Ns[j]*deltas_[j]/(deltas_[j]-1)*eval_laguerre(Ns[j]-1, phis_[j])/eval_laguerre(Ns[j], phis_[j]) for j in range(R)] )
def k_avg( T, E0s, Dels, Evs, Ns):
"""
Function that computes k_avg
"""
x, z = fsolve(constr_func, x0 = (50,500), args = (T, E0s, Dels, Evs, Ns))
R = len(Ns)
gammas_ = gamma_func(E0s,Evs, T)
deltas_ = delta_func(Dels, T)
phis_ = phi_func(x, z, gammas_, deltas_)
return np.sum([Ns[j]/(z*gammas_[j]+1)*(z*gammas_[j] + eval_laguerre(Ns[j]-1, phis_[j])/eval_laguerre(Ns[j], phis_[j])) for j in range(R)])
# -
# #### Important temperatures
#
# The temperature $k_BT_{\text{crit}}$ at which the system settles into its completely correct configuration is defined by
#
# \begin{equation}
# 1 = \sum_{j=1}^{R}n_j e^{-\beta_{\text{crit}} \Delta_j}\left( 1+ (\beta_{\text{crit}} E_V)^{-3/2} e^{-\beta_{\text{crit}} E_j}\right) + O(e^{-2\beta_{\text{crit}} \Delta_j}).
# \label{eq:master_therm_subs}
# \end{equation}
# +
# general thermal constraint
def master_them_constr(T, E0s, Dels, Evs, Ns):
F = 1-np.sum(Ns*delta_func(Dels, T)**(-1)*(1+gamma_func(E0s, Evs, T)**(-1)))
return F
# critical temperature
kBTcrit_master = lambda E0s, Dels, Evs, Ns: fsolve(master_them_constr, x0 = 100.5, args = (E0s, Dels, Evs, Ns))[0]
# -
# #### Example analytical plot
# +
# temperature vals
Tvals = np.linspace(0.1, 3.0, 50)
# parameters for the integral
np.random.seed(42)
R = 50
E0_bar, sigma_E = 10.0, 2.0
Del_bar, sigma_D = 3.0, 1.0
E0s = np.random.randn(R)*sigma_E+E0_bar
Dels = np.random.randn(R)*sigma_D+Del_bar
Nelems = np.random.randint(1,10,R)
Evs = np.ones(R)*0.001
# computing analytical values of k and m
avg_k_approx_vals = [k_avg(T, E0s, Dels, Evs, Nelems) for T in Tvals]
avg_m_approx_vals = [m_avg(T, E0s, Dels, Evs, Nelems) for T in Tvals]
# +
## plotting order parameters
ax = plt.subplot(111)
ax.plot(Tvals, avg_k_approx_vals/np.sum(Nelems), label = r'$\langle k \rangle$ (large $N$)')
ax.plot(Tvals, avg_m_approx_vals/np.sum(Nelems), label = r'$\langle m \rangle$ (large $N$)')
ax.set_xlabel(r'$k_BT$', fontsize = 18, labelpad = 10.5)
ax.grid(alpha = 0.5)
ax.axvline(x = kBTcrit_master(E0s, Dels, Evs, Nelems), color = 'g', linestyle = '--')
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
# -
# ### Metropolis Hastings simulation code
# #### Microstate transitions
# +
## dissociation operator
def trans_dissoc(free_objs, bound_objs):
# indices of non-empty
indxs = [i for i, x in enumerate(bound_objs) if x != "-"]
# random choice for bound object
random_indx = random.choice(indxs)
## new state vector
free_objs_new = copy.deepcopy(free_objs)
bound_objs_new = copy.deepcopy(bound_objs)
# putting empty slot
bound_objs_new[random_indx] = '-'
# appending previously bound object to free objects
free_objs_new.append(bound_objs[random_indx])
return free_objs_new, bound_objs_new
## association operator
def trans_assoc(free_objs, bound_objs):
# random element to associate
elem = random.choice(free_objs)
# indices of empty spaces
indxs = [i for i, x in enumerate(bound_objs) if x == "-"]
# random choice for empty space
random_indx = random.choice(indxs)
## new state vector
free_objs_new = copy.deepcopy(free_objs)
bound_objs_new = copy.deepcopy(bound_objs)
## state
free_objs_new.remove(elem)
bound_objs_new[random_indx] = elem
return free_objs_new, bound_objs_new
## permutation operator
def trans_perm(free_objs, bound_objs):
Ncomp = len(bound_objs)
i1 = int(random.choice(range(Ncomp)))
i2 = int(random.choice(range(Ncomp)))
## new omega vector
bound_objs_new = copy.deepcopy(bound_objs)
bound_objs_new[i2] = bound_objs[i1]
bound_objs_new[i1] = bound_objs[i2]
return free_objs, bound_objs_new
# -
# #### Logarithm of Botlzmann factor
#
# The logarithm of the Botlzmann factor for a microstate (i.e., the temperature normalized negative energy of the microstate) is defined as
#
# \begin{equation}
# \beta E(\boldsymbol{k}, \boldsymbol{m}) = \sum_{i=1}^R(m_i \ln \delta_i + k_i \ln \gamma_i).
# \label{eq:sim_en}
# \end{equation}
def log_boltz(free_objs, bound_objs, mstr_vec, deltas, gammas, name_key):
elem_set = list(set(mstr_vec))
count_dict = dict()
for elem in elem_set:
count_dict[elem] = bound_objs.count(elem)
bind_log_factor = 0
for elem in elem_set:
key = name_key[elem]
bind_log_factor += count_dict[elem]*np.log(gammas[key])
corr_log_factor = 0
for j in range(len(bound_objs)):
if bound_objs[j] == mstr_vec[j]:
elem = bound_objs[j]
key = name_key[elem]
corr_log_factor+=np.log(deltas[key])
return bind_log_factor+corr_log_factor
# #### Function to count the number of correctly bound particles
def m_calc(bound_objs, mstr_vec):
num = 0
for k in range(len(mstr_vec)):
if mstr_vec[k] == bound_objs[k]:
num += 1
return num
# #### Checking logarithm of Boltzmann factor definition
# +
# defining name key
name_key0 = dict()
key_list = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', ]
for j in range(len(key_list)):
name_key0[key_list[j]] = j
# random energies
np.random.seed(2)
q1 = np.random.rand(10)
q2 = np.random.rand(10)
# sample master list
sample_master = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', ]
bound_init_0 = ['-', '-', '-', '-', '-', '-', 'G', 'H', 'I', 'J', ]
free_init_0 = ['A', 'B', 'C', 'D', 'E', 'F' ]
print('Energy for original ordering:', log_boltz( free_init_0,bound_init_0, sample_master, deltas = np.exp(-q1), gammas = np.exp(-q2), name_key = name_key0))
e1 = -np.sum([q1[k] for k in range(len(sample_master)) if sample_master[k] == bound_init_0[k]])
e2 = -np.sum([q2[k] for k in range(len(sample_master)) if sample_master[k] in bound_init_0])
print('Checking energy value:', e1+e2)
print('Number of correctly placed elements:', m_calc(bound_init_0, sample_master))
print('Number of bound elements:', np.sum([1 for elem in bound_init_0 if elem!='-']))
print('-----')
random.seed(1)
free_init_0, perm_bound = trans_perm(free_init_0, bound_init_0)
free_init_new, bound_init_new = trans_assoc(free_init_0, perm_bound)
print('Energy after permutation and associaation values:', log_boltz(free_init_new, bound_init_new, sample_master, deltas = np.exp(-q1), gammas = np.exp(-q2), name_key = name_key0))
e1 = -np.sum([q1[k] for k in range(len(sample_master)) if sample_master[k] == bound_init_new[k]])
e2 = -np.sum([q2[k] for k in range(len(sample_master)) if sample_master[k] in bound_init_new])
print('Checking energy value:', e1+e2)
print('Number of correctly placed elements:', m_calc(bound_init_new, sample_master))
print('Number of bound elements:', np.sum([1 for elem in bound_init_new if elem!='-']))
# -
# #### Metropolis Hastings algorithm
# +
### Metropolis Monte Carlo Algorithm
## loads uniform random sampling
runif = np.random.rand
def met_assembly_grid(Niter, free_objs, bound_objs, mstr_vec, deltas, gammas, name_key):
'''
#################################################################
# function to sample using Metropolis
#
# n_iter: number of iterations
# initial_state: initial state for the start position for our chain
# gamma: energy cost for incorrect component
# temp: temperature
##################################################################
'''
# Initialize state values
free_objs_vals = [0]*(Niter+1)
bound_objs_vals = [0]*(Niter+1)
# Set initial values
free_objs_vals[0] = free_objs[:]
bound_objs_vals[0] = bound_objs[:]
# Initialize acceptance counts
# We can use this to tune our number of steps
accepted = 0
# debugging code
debug_assoc, debug_dissoc, debug_perm = 0, 0, 0
for i in range(Niter):
# get current monomer and dimer states
current_free_objs = copy.deepcopy(free_objs_vals[i])
current_bound_objs = copy.deepcopy(bound_objs_vals[i])
N_free = len(current_free_objs)
N_bound = len(current_bound_objs)-len(current_free_objs)
u_trans = runif()
if u_trans < 1/3: #first type of transition; monomer association
if N_free < 1:
log_alpha = np.log(1e-15)
else:
# proposed new monomer and dimer states
new_free_objs, new_bound_objs = trans_assoc(current_free_objs, current_bound_objs)
# transition elements
log_init = log_boltz(current_free_objs, current_bound_objs, mstr_vec, deltas, gammas, name_key)
log_final = log_boltz(new_free_objs, new_bound_objs, mstr_vec, deltas, gammas, name_key)
# weight
num = N_free*N_free
den = N_bound+1
# Log-acceptance rate
log_alpha = log_final-log_init+np.log(num/den)
elif 1/3 <= u_trans < 2/3: #second type of transition; bound monomer dissociation
if N_bound <1:
log_alpha = np.log(1e-15)
else:
# proposed new monomer and dimer states
new_free_objs, new_bound_objs = trans_dissoc(current_free_objs, current_bound_objs)
# transition elements
log_init = log_boltz(current_free_objs, current_bound_objs, mstr_vec, deltas, gammas, name_key)
log_final = log_boltz(new_free_objs, new_bound_objs, mstr_vec, deltas, gammas, name_key)
# weight
num = N_bound
den = (N_free+1)*(N_free+1)
# Log-acceptance rate
log_alpha = log_final-log_init+np.log(num/den)
elif 2/3 <= u_trans: #third type of transition; switching bounded elements
if N_bound <2:
log_alpha = np.log(1e-15)
else:
# proposed new monomer and dimer states
new_free_objs, new_bound_objs = trans_perm(current_free_objs, current_bound_objs)
# transition elements
log_init = log_boltz(current_free_objs, current_bound_objs, mstr_vec, deltas, gammas, name_key)
log_final = log_boltz(new_free_objs, new_bound_objs, mstr_vec, deltas, gammas, name_key)
# Log-acceptance rate
log_alpha = log_final-log_init
# Sample a uniform random variate
u = runif()
# Test proposed value
if np.log(u) < log_alpha:
# Accept
free_objs_vals[i+1] = new_free_objs
bound_objs_vals[i+1] = new_bound_objs
#log_current_prob = log_proposed_prob
accepted += 1
else:
# Stay put
free_objs_vals[i+1] = free_objs_vals[i]
bound_objs_vals[i+1] = bound_objs_vals[i]
# return our samples and the number of accepted steps
return free_objs_vals, bound_objs_vals, accepted
# -
# #### Computing microstate averages from simiulations
# +
def avg_k(bound_objs_vals, Nmc):
"""
Microstate average of number of bound objects
We only consider microstates near the end of theh chain to ensure
that the system has equilibrated
"""
length = int(Nmc/50)
ls = [0]*length
ls = np.array(ls)
for k in range(length):
ls[k] = len(bound_objs_vals[Nmc-length+k]) - bound_objs_vals[Nmc-length+k].count('-')
return(np.mean(ls))
# average number of correctly bound objects
def avg_m(bound_objs_vals, mstr_vec, Nmc):
"""
Microstate average of number of correctly bound objects
We only consider microstates near the end of theh chain to ensure
that the system has equilibrated
"""
length = int(Nmc/50)
ls = [0]*length
ls = np.array(ls)
for k in range(length):
ls[k] = np.sum([1 for j in range(len(mstr_vec)) if bound_objs_vals[Nmc-length+k][j]==mstr_vec[j]])
return(np.mean(ls))
# -
# #### Image grid for completely correct configuration
# defining master_list
master_list =compute_master_list()
# testing plot
imshow_list(master_list, title = 'Completely Correct Configuration');
# defining Nelems
Nelems = np.zeros(8)
key_list = list(rgb_map.keys())[:-1]
name_key_ = dict()
for j in range(len(key_list)):
name_key_[key_list[j]] = j
Nelems[j] = master_list.count(key_list[j])
# displaying copy-number counts of the various elements
Nelems
# #### Simulating system
# +
## Generate lf for each temperature from .03 to 2.0 in npoints steps
t0 = time.time()
# number of steps for MC algortihm
Nmc = 30000
# parameter definitions
R = 8
np.random.seed(24)
Del_bar, sigma_D = 3.0, 1.0
Dels = np.random.randn(R)*sigma_D+Del_bar
E0_bar, sigma_E = 14.0, 2.0
E0s = np.random.randn(R)*sigma_E+E0_bar
Evs = np.ones(R)*0.001
# initial monomer and dimer states;
# system in microstate of all correct dimers
random.seed(0)
free_objs_0 = []
bound_objs_0 = random.sample(master_list, len(master_list))
mstr_vec = copy.deepcopy(master_list)
# temperature limits
Tmin = .05
Tmax = 3.0
npoints = 15 #number of temperature values
navg = 5 # number of times we run simulation at each temperature; 50 in paper
temp_vals = np.linspace(Tmin, Tmax, npoints).tolist()
# list of dimer values
sim_k_vals = [0]*npoints
# list of correct dimer values
sim_m_vals = [0]*npoints
# accepted list
accepted_list = [0]*npoints
# saved list for plotting
saved_list = dict()
for k in range(npoints):
fin_k_vals = [0]*navg
fin_m_vals = [0]*navg
fin_accepted = [0]*navg
for j in range(navg):
# make copy of initial monomer and dimer states
free_objs_copy = copy.deepcopy(free_objs_0)
bound_objs_copy = copy.deepcopy(bound_objs_0)
# defining helper functions
gammas_ = gamma_func(E0s, Evs, temp_vals[k])
deltas_ = delta_func(Dels, temp_vals[k])
# metroplois generator
free_list, bound_list, accepted = met_assembly_grid(Nmc,
free_objs_copy,
bound_objs_copy,
mstr_vec,
deltas_,
gammas_,
name_key_)
# averaging final states to compute observables
fin_k_vals[j] = avg_k(bound_list, Nmc)
fin_m_vals[j] = avg_m(bound_list, mstr_vec, Nmc)
fin_accepted[j] = accepted
# saving every 5 temperatures
if (k+1)%5 ==0 or k ==0:
saved_list[k] = ['white' if x=='-' else x for x in bound_list[-1]]
# averaging over computed equilibrium averages
sim_k_vals[k] = np.mean(np.array(fin_k_vals))
sim_m_vals[k] = np.mean(np.array(fin_m_vals))
accepted_list[k] = np.mean(np.array(fin_accepted))
t_prelim = time.time()
print("Temperature Run:",str(k+1),"; Current Time:", round(t_prelim-t0,2),"secs")
t1 = time.time()
print("Total Simulation Run Time:",t1-t0,"secs")
# -
# #### Simulated image grid at various temperatures
# +
# figure parameters
rows, cols, idx = 2, 2, 0
fig, axes = plt.subplots(nrows=rows, ncols=cols, figsize=(9,9))
# list of keys for saved snapshots of image
img_key_list = list(saved_list.keys())
for i in range(rows):
for j in range(cols):
if idx < 4:
axes[i, j].imshow(color_to_rgb(list_to_matrix(saved_list[img_key_list[idx]])))
ax = plt.gca()
# making ticks invisible
axes[i, j].set_xticks([])
axes[i, j].set_yticks([])
# Minor ticks
axes[i, j].set_xticks(np.arange(-0.5, 11, 1), minor=True)
axes[i, j].set_yticks(np.arange(-0.5, 10, 1), minor=True)
axes[i, j].tick_params(axis='y', colors='red')
# labeling images
itimes = 'i'*(1+idx) if idx<3 else 'iv'
# Gridlines based on minor ticks
axes[i, j].grid(which='minor', color='w', linestyle='-', linewidth=3)
axes[i, j].set_title(fr'({itimes}) $k_BT = {round(temp_vals[img_key_list[idx]],2)}$', fontsize = 18, y = -.2)
# making spines invisible
axes[i, j].spines['right'].set_visible(False)
axes[i, j].spines['top'].set_visible(False)
axes[i, j].spines['left'].set_visible(False)
axes[i, j].spines['bottom'].set_visible(False)
idx +=1
# plt.savefig('general_grid_assembly_grid_plots.png', bbox_inches='tight', format = 'png')
plt.show()
# -
# #### Comparing analytical and simulation results
# +
plt.figure(figsize = (7,5))
ax = plt.subplot(111)
# simulation results
plt.plot(temp_vals,np.array(sim_k_vals)/np.sum(Nelems),
label = r'Sim. $\langle k \rangle$/N',
markersize = 7.5,
marker = 'D',
linestyle = '')
plt.plot(temp_vals,np.array(sim_m_vals)/np.sum(Nelems),
label = r'Sim. $\langle m \rangle$/N',
markersize = 7.5,
marker = 's',
linestyle = '')
# large N analytical results
k_avg_approx_vals = [k_avg(T, E0s, Dels, Evs, Nelems)/np.sum(Nelems) for T in Tvals]
m_avg_approx_vals = [m_avg(T, E0s, Dels, Evs, Nelems)/np.sum(Nelems) for T in Tvals]
plt.plot(Tvals, k_avg_approx_vals, label = r'Large $N$ $\langle k \rangle$/N', linestyle= '--', linewidth = 3.0)
plt.plot(Tvals, m_avg_approx_vals, label = r'Large $N$ $\langle m \rangle$/N', linewidth = 2.0 )
ax.axvline(x = kBTcrit_master(E0s, Dels, Evs, Nelems), color = 'k', linestyle = '-.', linewidth = 2)
plt.legend(loc = 'best', fontsize = 12)
# plot formatting
ax.set_xlabel(r'$k_B T$', fontsize = 18)
plt.xlim([-0.01,3.2])
plt.ylim([0,1.1])
plt.grid(alpha = 0.45)
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
# increase label size
ax.tick_params(axis='both', which='major', labelsize=12)
ax.tick_params(axis='both', which='minor', labelsize=12)
ax.text(kBTcrit_master(E0s, Dels, Evs, Nelems)-.2, 0.25, r'$k_BT_{crit}$', color='black', fontsize = 14.5,
bbox=dict(facecolor='white', edgecolor='none', pad=5.0))
for i in range(4):
ax.text(temp_vals[img_key_list[i]], sim_k_vals[img_key_list[i]]/np.sum(Nelems)+.05,'('+'i'*(1+i)+')' if i<3 else '(iv)', fontsize = 14 )
# plt.savefig(f'general_grid_assembly.png', bbox_inches='tight', format = 'png')
# -
print('Total Notebook Runtime: %.3f mins' % ((time.time()-nb_start)/60))
| general_grid_assembly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# + tags=["remove-input", "active-ipynb", "remove-output"]
# try:
# from openmdao.utils.notebook_utils import notebook_mode
# except ImportError:
# !python -m pip install openmdao[notebooks]
# -
# # ScipyKrylov
#
# ScipyKrylov is an iterative linear solver that wraps the methods found in `scipy.sparse.linalg`.
# The default method is "gmres", or the Generalized Minimal RESidual method. Support for other
# `scipy.sparse.linalg` solvers will be added over time. This linear solver is capable of handling any
# system topology very effectively. It also solves all subsystems below it in the hierarchy, so
# assigning different solvers to subsystems will have no effect on the solution at this level.
#
# This is a serial solver, so it should never be used under MPI; use [PETScKrylov](../../../_srcdocs/packages/solvers.linear/petsc_ksp) instead.
#
# Here, we calculate the total derivatives across the Sellar system.
# + tags=["remove-input", "remove-output"]
from openmdao.utils.notebook_utils import get_code
from myst_nb import glue
glue("code_src41", get_code("openmdao.test_suite.components.sellar.SellarDis1withDerivatives"), display=False)
# -
# :::{Admonition} `SellarDis1withDerivatives` class definition
# :class: dropdown
#
# {glue:}`code_src41`
# :::
# + tags=["remove-input", "remove-output"]
from openmdao.utils.notebook_utils import get_code
from myst_nb import glue
glue("code_src42", get_code("openmdao.test_suite.components.sellar.SellarDis2withDerivatives"), display=False)
# -
# :::{Admonition} `SellarDis2withDerivatives` class definition
# :class: dropdown
#
# {glue:}`code_src42`
# :::
# +
import numpy as np
import openmdao.api as om
from openmdao.test_suite.components.sellar import SellarDis1withDerivatives, SellarDis2withDerivatives
prob = om.Problem()
model = prob.model
model.add_subsystem('d1', SellarDis1withDerivatives(), promotes=['x', 'z', 'y1', 'y2'])
model.add_subsystem('d2', SellarDis2withDerivatives(), promotes=['z', 'y1', 'y2'])
model.add_subsystem('obj_cmp', om.ExecComp('obj = x**2 + z[1] + y1 + exp(-y2)',
z=np.array([0.0, 0.0]), x=0.0),
promotes=['obj', 'x', 'z', 'y1', 'y2'])
model.add_subsystem('con_cmp1', om.ExecComp('con1 = 3.16 - y1'), promotes=['con1', 'y1'])
model.add_subsystem('con_cmp2', om.ExecComp('con2 = y2 - 24.0'), promotes=['con2', 'y2'])
model.nonlinear_solver = om.NonlinearBlockGS()
model.linear_solver = om.ScipyKrylov()
prob.setup()
prob.set_val('x', 1.)
prob.set_val('z', np.array([5.0, 2.0]))
prob.run_model()
wrt = ['z']
of = ['obj']
J = prob.compute_totals(of=of, wrt=wrt, return_format='flat_dict')
# + tags=["remove-input", "remove-output"]
from openmdao.utils.assert_utils import assert_near_equal
assert_near_equal(J['obj', 'z'][0][0], 9.61001056, .00001)
assert_near_equal(J['obj', 'z'][0][1], 1.78448534, .00001)
# -
# ## ScipyKrylov Options
# + tags=["remove-input"]
om.show_options_table("openmdao.solvers.linear.scipy_iter_solver.ScipyKrylov")
# -
# ## ScipyKrylov Constructor
#
# The call signature for the `ScipyKrylov` constructor is:
#
# ```{eval-rst}
# .. automethod:: openmdao.solvers.linear.scipy_iter_solver.ScipyKrylov.__init__
# :noindex:
# ```
#
# ## ScipyKrylov Option Examples
#
# **maxiter**
#
# `maxiter` lets you specify the maximum number of GMRES iterations to apply. The default maximum is 1000, which
# is much higher than the other linear solvers because each multiplication by the system Jacobian is considered
# to be an iteration. You may have to decrease this value if you have a coupled system that is converging
# very slowly. (Of course, in such a case, it may be better to add a preconditioner.) Alternatively, you
# may have to raise it if you have an extremely large number of components in your system (a 1000-component
# ring would need 1000 iterations just to make it around once.)
#
# This example shows what happens if you set maxiter too low (the derivatives should be nonzero, but it stops too
# soon.)
# +
from openmdao.test_suite.components.sellar import SellarDis1withDerivatives, SellarDis2withDerivatives
prob = om.Problem()
model = prob.model
model.add_subsystem('d1', SellarDis1withDerivatives(), promotes=['x', 'z', 'y1', 'y2'])
model.add_subsystem('d2', SellarDis2withDerivatives(), promotes=['z', 'y1', 'y2'])
model.add_subsystem('obj_cmp', om.ExecComp('obj = x**2 + z[1] + y1 + exp(-y2)',
z=np.array([0.0, 0.0]), x=0.0),
promotes=['obj', 'x', 'z', 'y1', 'y2'])
model.add_subsystem('con_cmp1', om.ExecComp('con1 = 3.16 - y1'), promotes=['con1', 'y1'])
model.add_subsystem('con_cmp2', om.ExecComp('con2 = y2 - 24.0'), promotes=['con2', 'y2'])
model.nonlinear_solver = om.NonlinearBlockGS()
model.linear_solver = om.ScipyKrylov()
model.linear_solver.options['maxiter'] = 3
prob.setup()
prob.set_val('x', 1.)
prob.set_val('z', np.array([5.0, 2.0]))
prob.run_model()
wrt = ['z']
of = ['obj']
J = prob.compute_totals(of=of, wrt=wrt, return_format='flat_dict')
# -
print(J['obj', 'z'][0][0])
print(J['obj', 'z'][0][1])
# + tags=["remove-input", "remove-output"]
assert_near_equal(J['obj', 'z'][0][0], 0.0, .00001)
assert_near_equal(J['obj', 'z'][0][1], 0.0, .00001)
# -
# **atol**
#
# Here, we set the absolute tolerance to a much tighter value (default is 1.0e-12) to show what happens. In
# practice, the tolerance serves a dual role in GMRES. In addition to being a termination criteria, the tolerance
# also defines what GMRES considers to be tiny. Tiny numbers are replaced by zero when the argument vector is
# normalized at the start of each new matrix-vector product. The end result here is that we iterate longer to get
# a marginally better answer.
#
# You may need to adjust this setting if you have abnormally large or small values in your global Jacobian.
# +
from openmdao.test_suite.components.sellar import SellarDis1withDerivatives, SellarDis2withDerivatives
prob = om.Problem()
model = prob.model
model.add_subsystem('d1', SellarDis1withDerivatives(), promotes=['x', 'z', 'y1', 'y2'])
model.add_subsystem('d2', SellarDis2withDerivatives(), promotes=['z', 'y1', 'y2'])
model.add_subsystem('obj_cmp', om.ExecComp('obj = x**2 + z[1] + y1 + exp(-y2)',
z=np.array([0.0, 0.0]), x=0.0),
promotes=['obj', 'x', 'z', 'y1', 'y2'])
model.add_subsystem('con_cmp1', om.ExecComp('con1 = 3.16 - y1'), promotes=['con1', 'y1'])
model.add_subsystem('con_cmp2', om.ExecComp('con2 = y2 - 24.0'), promotes=['con2', 'y2'])
model.nonlinear_solver = om.NonlinearBlockGS()
model.linear_solver = om.ScipyKrylov()
model.linear_solver.options['atol'] = 1.0e-20
prob.setup()
prob.set_val('x', 1.)
prob.set_val('z', np.array([5.0, 2.0]))
prob.run_model()
wrt = ['z']
of = ['obj']
J = prob.compute_totals(of=of, wrt=wrt, return_format='flat_dict')
# -
print(J['obj', 'z'][0][0])
print(J['obj', 'z'][0][1])
# + tags=["remove-input", "remove-output"]
assert_near_equal(J['obj', 'z'][0][0], 9.61001055699, .00001)
assert_near_equal(J['obj', 'z'][0][1], 1.78448533563, .00001)
# -
# **rtol**
#
# The 'rtol' setting is not supported by Scipy GMRES.
#
# ## Specifying a Preconditioner
#
# You can specify a preconditioner to improve the convergence of the iterative linear solution by setting the `precon` attribute. The
# motivation for using a preconditioner is the observation that iterative methods have better convergence
# properties if the linear system has a smaller condition number, so the goal of the preconditioner is to
# improve the condition number in part or all of the Jacobian.
#
# Here, we add a Gauss-Seidel preconditioner to a problem that contains two subgroups, each with an implicit component that implements a quadratic
# equation. These are solved together by a Newton solver at the top. The goal of the preconditioner here is to solve the smaller linear systems
# for the quadratic components independently, and use that solution to precondition the full system solution. This is accomplished by setting up
# the linear solver hierarchy so that the preconditioner is `LinearBlockGS` and the subsytems `sub1` and `sub2` contain a `DirectSolver`.
#
# Note that the number of GMRES iterations is lower when using the preconditioner.
# + tags=["remove-input", "remove-output"]
from openmdao.utils.notebook_utils import get_code
from myst_nb import glue
glue("code_src43", get_code("openmdao.test_suite.components.quad_implicit.QuadraticComp"), display=False)
# -
# :::{Admonition} `QuadraticComp` class definition
# :class: dropdown
#
# {glue:}`code_src43`
# :::
# + tags=["output_scroll"]
from openmdao.test_suite.components.quad_implicit import QuadraticComp
prob = om.Problem()
model = prob.model
sub1 = model.add_subsystem('sub1', om.Group())
sub1.add_subsystem('q1', QuadraticComp())
sub1.add_subsystem('z1', om.ExecComp('y = -6.0 + .01 * x'))
sub2 = model.add_subsystem('sub2', om.Group())
sub2.add_subsystem('q2', QuadraticComp())
sub2.add_subsystem('z2', om.ExecComp('y = -6.0 + .01 * x'))
model.connect('sub1.q1.x', 'sub1.z1.x')
model.connect('sub1.z1.y', 'sub2.q2.c')
model.connect('sub2.q2.x', 'sub2.z2.x')
model.connect('sub2.z2.y', 'sub1.q1.c')
model.nonlinear_solver = om.NewtonSolver(solve_subsystems=False)
model.linear_solver = om.ScipyKrylov()
prob.setup()
model.sub1.linear_solver = om.DirectSolver()
model.sub2.linear_solver = om.DirectSolver()
model.linear_solver.precon = om.LinearBlockGS()
prob.set_solver_print(level=2)
prob.run_model()
# -
print(prob.get_val('sub1.q1.x'))
print(prob.get_val('sub2.q2.x'))
# + tags=["remove-input", "remove-output"]
assert_near_equal(prob.get_val('sub1.q1.x'), 1.996, .0001)
assert_near_equal(prob.get_val('sub2.q2.x'), 1.996, .0001)
| openmdao/docs/openmdao_book/features/building_blocks/solvers/scipy_iter_solver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/intel-analytics/analytics-zoo/blob/master/docs/docs/colab-notebook/chronos/chronos_autots_nyc_taxi.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xgV_QLGE9Lox"
#
# 
# ---
# + [markdown] id="fDBPZ0_rfBmU"
# ##### Copyright 2018 Analytics Zoo Authors.
# + id="xBWVU_bhfkY7"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# + [markdown] id="voMBntim9bMf"
# ## **Environment Preparation**
# + [markdown] id="I_OS4HKJMNpv"
# **Install Analytics Zoo**
#
# You can install the latest pre-release version with chronos support using `pip install --pre --upgrade analytics-zoo[automl]`.
# + id="3qfT8CaC51hI"
# Install latest pre-release version of Analytics Zoo
# Installing Analytics Zoo from pip will automatically install pyspark, bigdl, and their dependencies.
# !pip install --pre --upgrade analytics-zoo[automl]
exit() # restart the runtime to refresh installed pkg
# + [markdown] id="TsT-0y8w-6N5"
# ### **Step 0: Download & prepare dataset**
# We used NYC taxi passengers dataset in [Numenta Anomaly Benchmark (NAB)](https://github.com/numenta/NAB) for demo, which contains 10320 records, each indicating the total number of taxi passengers in NYC at a corresonponding time spot.
# + id="devxHuDW-0Mb"
# download the dataset
# !wget https://raw.githubusercontent.com/numenta/NAB/v1.0/data/realKnownCause/nyc_taxi.csv
# + id="LVpFKkCX_3WF"
# load the dataset. The downloaded dataframe contains two columns, "timestamp" and "value".
import pandas as pd
df = pd.read_csv("nyc_taxi.csv", parse_dates=["timestamp"])
# -
# ## **Time series forecasting using Chronos Forecaster**
# ### Forecaster Step1. Data transformation and feature engineering using Chronos TSDataset
# [TSDataset](https://analytics-zoo.readthedocs.io/en/latest/doc/PythonAPI/Chronos/tsdataset.html) is our abstract of time series dataset for data transformation and feature engineering. Here we use it to preprocess the data.
from zoo.chronos.data import TSDataset
from sklearn.preprocessing import StandardScaler
# Initialize train, valid and test tsdataset from raw pandas dataframe.
tsdata_train, tsdata_valid, tsdata_test = TSDataset.from_pandas(df, dt_col="timestamp", target_col="value",
with_split=True, val_ratio=0.1, test_ratio=0.1)
# Preprocess the datasets. Here we perform:
# - deduplicate: remove those identical data records
# - impute: fill the missing values
# - gen_dt_feature: generate feature from datetime (e.g. month, day...)
# - scale: scale each feature to standard distribution.
# - roll: sample the data with sliding window.
#
# For forecasting task, we will look back 3 hours' historical data (6 records) and predict the value of next 30 miniutes (1 records).
#
# We perform the same transformation processes on train, valid and test set.
# +
lookback, horizon = 6, 1
scaler = StandardScaler()
for tsdata in [tsdata_train, tsdata_valid, tsdata_test]:
tsdata.deduplicate()\
.impute()\
.gen_dt_feature()\
.scale(scaler, fit=(tsdata is tsdata_train))\
.roll(lookback=lookback, horizon=horizon)
# -
# ### Forecaster Step 2: Time series forecasting using Chronos Forecaster
# After preprocessing the datasets. We can use [Chronos Forecaster](https://analytics-zoo.readthedocs.io/en/latest/doc/PythonAPI/Chronos/forecasters.html) to handle the forecasting tasks.
# set random seeds to help reproduce the same result
import torch, random, numpy
from zoo.chronos.model.forecast.tcn_forecaster import TCNForecaster
torch.manual_seed(0)
numpy.random.seed(0)
random.seed(0)
# Transform TSDataset to sampled numpy ndarray and feed them to forecaster.
# +
x, y = tsdata_train.to_numpy()
# x.shape = (num of sample, lookback, num of input feature)
# y.shape = (num of sample, horizon, num of output feature)
forecaster = TCNForecaster(past_seq_len=lookback, # number of steps to look back
future_seq_len=horizon, # number of steps to predict
input_feature_num=x.shape[-1], # number of feature to use
output_feature_num=y.shape[-1]) # number of feature to predict
res = forecaster.fit(x, y, epochs=3)
# -
# ### Forecaster Step 3: Further deployment with fitted forecaster
# Use fitted forecaster to predict test data and plot the result
x_test, y_test = tsdata_test.to_numpy()
pred = forecaster.predict(x_test)
pred_unscale, groundtruth_unscale = tsdata_test.unscale_numpy(pred), tsdata_test.unscale_numpy(y_test)
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(24,6))
plt.plot(pred_unscale[:,:,0])
plt.plot(groundtruth_unscale[:,:,0])
plt.legend(["prediction", "ground truth"])
# -
# Save & restore the forecaster.
forecaster.save("nyc_taxi.fxt")
forecaster.restore("nyc_taxi.fxt")
| docs/docs/colab-notebook/chronos/chronos_nyc_taxi_tsdataset_forecaster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# **Course website**: https://github.com/leomiquelutti/UFU-geofisica-1
#
# **Note**: This notebook is part of the course "Geofísica 1" of Geology program of the
# [Universidade Federal de Uberlândia](http://www.ufu.br/).
# All content can be freely used and adapted under the terms of the
# [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
#
# 
#
# Agradecimentos especiais ao [<NAME>](www.leouieda.com)
# Esse documento que você está usando é um [Jupyter notebook](http://jupyter.org/). É um documento interativo que mistura texto (como esse), código (como abaixo), e o resultado de executar o código (números, texto, figuras, videos, etc).
# # Gravimetria - Interpolação, mapas e a gravidade da Terra
# ## Objetivos
#
# * Entender a influência da interpolação na geração de mapas de dados geofísicos
# * Visualizar as variações geográficas da gravidade da Terra
# * Entender como a escala de cores utilizada nos mapas influencia nossa interpretação
# * Aprender quais são os fatores que devem ser considerados quando visualizamos um dado em mapa
# ## Instruções
#
# O notebook te fornecerá exemplos interativos que trabalham os temas abordados no questionário. Utilize esses exemplos para responder as perguntas.
#
# As células com números ao lado, como `In [1]:`, são código [Python](http://python.org/). Algumas dessas células não produzem resultado e servem de preparação para os exemplos interativos. Outras, produzem gráficos interativos. **Você deve executar todas as células, uma de cada vez**, mesmo as que não produzem gráficos.
#
# Para executar uma célula, clique em cima dela e aperte `Shift + Enter`. O foco (contorno verde ou cinza em torno da célula) deverá passar para a célula abaixo. Para rodá-la, aperte `Shift + Enter` novamente e assim por diante. Você pode executar células de texto que não acontecerá nada.
# ## Preparação
#
# Exectute as células abaixo para carregar as componentes necessárias para nossa prática. Vamos utilizar várias *bibliotecas*, inclusive uma de geofísica chamada [Fatiando a Terra](http://www.fatiando.org).
# %matplotlib inline
from __future__ import division
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
import ipywidgets as widgets
from IPython.display import display
import seaborn
from fatiando import utils, gridder
import fatiando
from icgem import load_icgem_gdf, down_sample
print("Usando a versão do Fatiando a Terra: {}".format(fatiando.__version__))
# ## Interpolação
# O melhor jeito de entendermos o efeito da interpolação é fabricando alguns dados fictícios (sintéticos).
# Assim, podemos gerar os dados tanto em pontos aleatórios quanto em um grid regular.
# Isso nos permite comparar os resultados da interpolação com o *verdadeiro*. Nosso verdadeiro será um conjunto de dados medidos em um grid regular. Como se tivéssemos ido ao campo e medido em um grid regular.
# Rode a célula abaixo para gerar os dados em pontos aleatórios e em um grid regular.
area = (-5000., 5000., -5000., 5000.)
shape = (100, 100)
xp, yp = gridder.scatter(area, 100, seed=6)
x, y = [i.reshape(shape) for i in gridder.regular(area, shape)]
aletatorio = 50*utils.gaussian2d(xp, yp, 10000, 1000, angle=45)
regular = 50*utils.gaussian2d(x, y, 10000, 1000, angle=45).reshape(shape)
# Rode as duas células abaixo para gerar um gráfico interativo. Nesse gráfico você poderá controlar:
#
# * O número de pontos (em x e y) do grid utilizado na interpolação (`num_pontos`)
# * O método de interpolação utilizado (`metodo`). Pode ser interpolação cúbica ou linear.
# * Mostrar ou não os pontos de medição aleatórios no mapa interpolado.
#
# **Repare no que acontece com as bordas do mapa e onde não há observações**.
def interpolacao(num_pontos, metodo, pontos_medidos):
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
ishape = (num_pontos, num_pontos)
tmp = gridder.interp(yp, xp, aletatorio, ishape, area=area, algorithm=metodo, extrapolate=True)
yi, xi, interp = [i.reshape(ishape) for i in tmp]
ranges = np.abs([interp.min(), interp.max()]).max()
kwargs = dict(cmap="RdBu_r", vmin=-ranges, vmax=ranges)
ax = axes[0]
ax.set_title(u'Pontos medidos')
ax.set_aspect('equal')
tmp = ax.scatter(yp*0.001, xp*0.001, s=80, c=aletatorio, **kwargs)
plt.colorbar(tmp, ax=ax, aspect=50, pad=0.01)
ax.set_xlabel('y (km)')
ax.set_ylabel('x (km)')
ax.set_xlim(-5, 5)
ax.set_ylim(-5, 5)
plt.tight_layout(pad=0)
ax = axes[1]
ax.set_title(u'Interpolado')
ax.set_aspect('equal')
tmp = ax.contourf(yi*0.001, xi*0.001, interp, 40, **kwargs)
plt.colorbar(tmp, ax=ax, aspect=50, pad=0.01)
if pontos_medidos:
ax.plot(yp*0.001, xp*0.001, '.k')
ax.set_xlabel('y (km)')
ax.set_ylabel('x (km)')
ax.set_xlim(-5, 5)
ax.set_ylim(-5, 5)
plt.tight_layout(pad=0)
w = widgets.interactive(interpolacao, num_pontos=(5, 100, 5), metodo=['cubic', 'linear'], pontos_medidos=False)
display(w)
# Vamos verificar se alguma das combinações chegou perto do resultado *verdadeiro*.
#
# Rode a célula abaixo para gerar um gráfico dos dados verdadeiros (gerados em um grid regular). Esse deveria ser o resultado observado se a interpolação fosse perfeita.
fig, ax = plt.subplots(1, 1, figsize=(7, 6))
ranges = np.abs([regular.min(), regular.max()]).max()
kwargs = dict(cmap="RdBu_r", vmin=-ranges, vmax=ranges)
ax.set_title(u'Verdadeiro')
ax.set_aspect('equal')
tmp = ax.contourf(y*0.001, x*0.001, regular, 40, **kwargs)
plt.colorbar(tmp, ax=ax, aspect=50, pad=0.01)
ax.plot(yp*0.001, xp*0.001, '.k')
ax.set_xlabel('y (km)')
ax.set_ylabel('x (km)')
plt.tight_layout(pad=0)
# # Gravidade do mundo
# Vamos visualizar como a gravidade da Terra varia geograficamente. Os dados da gravidade do mundo foram baixados de http://icgem.gfz-potsdam.de/ICGEM/potato/Service.html usando o modelo EIGEN-6c3stat.
#
# **As medições foram feitas em cima da superfície da Terra**, ou seja, acompanhando a topografia.
# Rode as células abaixo para carregar os dados.
dados = load_icgem_gdf('data/eigen-6c3stat-0_5-mundo.gdf')
lat, lon, grav = dados['latitude'], dados['longitude'], dados['gravity_earth']
# Vamos fazer um mapa da gravidade utilizando a [projeção Mollweid](http://en.wikipedia.org/wiki/Map_projection). Esses dados estão em mGal: 1 mGal = 10⁻⁵ m/s².
#
# Rode as duas células abaixo para gerar o gráfico (isso pode demorar um pouco).
bm = Basemap(projection='moll', lon_0=0, resolution='c')
x, y = bm(lon, lat)
plt.figure(figsize=(18, 10))
tmp = bm.contourf(x, y, grav, 100, tri=True, cmap='Reds')
plt.colorbar(orientation='horizontal', pad=0.01, aspect=50, shrink=0.5).set_label('mGal')
plt.title("Gravidade medida na superficie da Terra", fontsize=16)
# ## Escala de cor
#
# A escala de cores que utilizamos para mapear os valores pode ter um impacto grande na nossa interpretação dos resultados. Abaixo, veremos como o nosso dado de gravidade mundial fica quando utilizamos diferentes escalas de cor.
#
# As escalas podem ser divididas em 3 categorias:
#
# * lineares: as cores variam de um tom claro (geralmente branco) a uma cor (por exemplo, vermelho) de maneira linear
# * divergente: as cores variam de uma cor escura, passando por um tom claro (geralmente branco), e depois para outra cor escura.
# * raindow ou qualitativos: as cores variam sem um padrão de intensidade claro. Podem ser as cores do arco-íris ou outra combinação.
#
# Nas escalas lineares e divergentes, as cores sempre variam de baixa intensidade para alta intensidade (e vice-versa para escalas divergentes).
# Rode as células abaixo para gerar um mapa interativo da gravidade mundial. Você poderá controlar qual escala de cor você quer usar. Experimente com elas e veja como elas afetam sua percepção.
#
# **Para pensar**: Como isso pode afetar alguem que é [daltônico](https://pt.wikipedia.org/wiki/Daltonismo)?
def grav_mundial(escala_de_cor):
plt.figure(figsize=(18, 10))
tmp = bm.contourf(x, y, grav, 100, tri=True, cmap=escala_de_cor)
plt.colorbar(orientation='horizontal', pad=0.01, aspect=50, shrink=0.5).set_label('mGal')
plt.title("Escala de cor: {}".format(escala_de_cor), fontsize=16)
escalas = 'Reds Blues Greys YlOrBr RdBu BrBG PRGn Dark2 jet ocean rainbow gnuplot'.split()
w = widgets.interactive(grav_mundial, escala_de_cor=escalas)
display(w)
# []()
| notebooks/GRAV_1-mapas-interpolacao-gravidade.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="introduction"></a>
# ## Introduction to Dask XGBoost
# #### By <NAME>
# -------
#
# In this notebook, we will show how to work with Dask XGBoost in RAPIDS.
#
# **Table of Contents**
#
# * [Introduction to Dask XGBoost](#introduction)
# * [Setup](#setup)
# * [Load Libraries](#libraries)
# * [Create a Cluster and Client](#cluster)
# * [Generate Data](#generate)
# * [Load Data](#load)
# * [Simulate Data](#simulate)
# * [Split Data](#split)
# * [Check Dimensions](#check)
# * [Distribute Data using Dask cuDF](#distribute)
# * [Set Parameters](#parameters)
# * [Train Model](#train)
# * [Generate Predictions](#predict)
# * [Evaluate Model](#evaluate)
# * [Conclusion](#conclusion)
# <a id="setup"></a>
# ## Setup
#
# This notebook was tested using the following Docker containers:
#
# * `rapidsai/rapidsai-nightly:0.8-cuda10.0-devel-ubuntu18.04-gcc7-py3.7` from [DockerHub - rapidsai/rapidsai-nightly](https://hub.docker.com/r/rapidsai/rapidsai-nightly)
#
# This notebook was run on the NVIDIA Tesla V100 GPU. Please be aware that your system may be different and you may need to modify the code or install packages to run the below examples.
#
# If you think you have found a bug or an error, please file an issue here: https://github.com/rapidsai/notebooks/issues
#
# Before we begin, let's check out our hardware setup by running the `nvidia-smi` command.
# !nvidia-smi
# Next, let's see what CUDA version we have.
# !nvcc --version
# <a id="libraries"></a>
# ## Load Libraries
#
# Let's load some of the libraries within the RAPIDs ecosystem and see which versions we have.
import cudf; print('cuDF Version:', cudf.__version__)
import dask; print('Dask Version:', dask.__version__)
import dask_cudf; print('Dask cuDF Version:', dask_cudf.__version__)
import dask_xgboost; print('Dask XGBoost Version:', dask_xgboost.__version__)
import numpy as np; print('numpy Version:', np.__version__)
import pandas as pd; print('pandas Version:', pd.__version__)
import sklearn; print('Scikit-Learn Version:', sklearn.__version__)
# import xgboost as xgb; print('XGBoost Version:', xgb.__version__)
# <a id="cluster"></a>
# ## Create a Cluster and Client
#
# Let's start by creating a local cluster of workers and a client to interact with that cluster.
# +
from dask.distributed import Client
from dask_cuda import LocalCUDACluster
# create a local CUDA cluster
cluster = LocalCUDACluster()
client = Client(cluster)
client
# -
# <a id="generate"></a>
# ## Generate Data
#
# <a id="load"></a>
# ### Load Data
#
# We can load the data using `pandas.read_csv`. We've provided a helper function `load_data` that will load data from a CSV file (and will only read the first 1 billion rows if that file is unreasonably big).
# helper function for loading data
def load_data(filename, n_rows):
if n_rows >= 1e9:
df = pd.read_csv(filename)
else:
df = pd.read_csv(filename, nrows=n_rows)
return df.values.astype(np.float32)
# <a id="simulate"></a>
# ### Simulate Data
#
# Alternatively, we can simulate data for our train and validation datasets. The features will be tabular with `n_rows` and `n_columns` in the training dataset, where each value is either of type `np.float32`. We can simulate data for both classification and regression using the `make_classification` or `make_regression` functions from the Scikit-Learn package.
# +
from sklearn.datasets import make_classification, make_regression
# helper function for simulating data
def simulate_data(m, n, k=2, random_state=None, classification=True):
if classification:
features, labels = make_classification(n_samples=m, n_features=n,
n_informative=int(n/5), n_classes=k,
random_state=random_state)
else:
features, labels = make_regression(n_samples=m, n_features=n,
n_informative=int(n/5), n_targets=1,
random_state=random_state)
return np.c_[labels, features].astype(np.float32)
# -
# settings
simulate = True
classification = True # change this to false to use regression
n_rows = int(1e6) # we'll use 1 millions rows
n_columns = int(100)
n_categories = 2
random_state = np.random.RandomState(43210)
# +
# %%time
if simulate:
dataset = simulate_data(n_rows, n_columns, n_categories,
random_state=random_state,
classification=classification)
else:
dataset = load_data('/tmp', n_rows)
print(dataset.shape)
# -
# <a id="split"></a>
# ### Split Data
#
# We'll split our dataset into a 80% training dataset and a 20% validation dataset.
# +
# identify shape and indices
n_rows, n_columns = dataset.shape
train_size = 0.80
train_index = int(n_rows * train_size)
# split X, y
X, y = dataset[:, 1:], dataset[:, 0]
del dataset
# split train data
X_train, y_train = X[:train_index, :], y[:train_index]
# split validation data
X_validation, y_validation = X[train_index:, :], y[train_index:]
# -
# <a id="check"></a>
# ### Check Dimensions
#
# We can check the dimensions and proportions of our training and validation dataets.
# +
# check dimensions
print('X_train: ', X_train.shape, X_train.dtype, 'y_train: ', y_train.shape, y_train.dtype)
print('X_validation', X_validation.shape, X_validation.dtype, 'y_validation: ', y_validation.shape, y_validation.dtype)
# check the proportions
total = X_train.shape[0] + X_validation.shape[0]
print('X_train proportion:', X_train.shape[0] / total)
print('X_validation proportion:', X_validation.shape[0] / total)
# -
# <a id="distribute"></a>
# ### Distribute Data using Dask cuDF
#
# Next, let's distribute our data across multiple GPUs using Dask cuDF.
# +
# create Pandas DataFrames for X_train and X_validation
n_columns = X_train.shape[1]
X_train_pdf = pd.DataFrame(X_train)
X_train_pdf.columns = ['feature_' + str(i) for i in range(n_columns)]
X_validation_pdf = pd.DataFrame(X_validation)
X_validation_pdf.columns = ['feature_' + str(i) for i in range(n_columns)]
# create Pandas DataFrames for y_train and y_validation
y_train_pdf = pd.DataFrame(y_train)
y_train_pdf.columns = ['y']
y_validation_pdf = pd.DataFrame(y_validation)
y_validation_pdf.columns = ['y']
# +
# Dask settings
npartitions = 8
# create Dask DataFrames for X_train and X_validation
X_train_dask_pdf = dask.dataframe.from_pandas(X_train_pdf, npartitions=npartitions)
X_validation_dask_pdf = dask.dataframe.from_pandas(X_validation_pdf, npartitions=npartitions)
# create Dask cuDF DataFrames for X_train and X_validation
X_train_dask_cudf = dask_cudf.from_dask_dataframe(X_train_dask_pdf)
X_validation_dask_cudf = dask_cudf.from_dask_dataframe(X_validation_dask_pdf)
# create Dask DataFrames for y_train and y_validation
y_train_dask_pdf = dask.dataframe.from_pandas(y_train_pdf, npartitions=npartitions)
y_validation_dask_pdf = dask.dataframe.from_pandas(y_validation_pdf, npartitions=npartitions)
# create Dask cuDF DataFrames for y_train and y_validation
y_train_dask_cudf = dask_cudf.from_dask_dataframe(y_train_dask_pdf)
y_validation_dask_cudf = dask_cudf.from_dask_dataframe(y_validation_dask_pdf)
# -
# Optional: persist training and validation data into memory
X_train_dask_cudf = X_train_dask_cudf.persist()
X_validation_dask_cudf = X_validation_dask_cudf.persist()
y_train_dask_cudf = y_train_dask_cudf.persist()
y_validation_dask_cudf = y_validation_dask_cudf.persist()
# <a id="parameters"></a>
# ## Set Parameters
#
# There are a number of parameters that can be set before XGBoost can be run.
#
# * General parameters relate to which booster we are using to do boosting, commonly tree or linear model
# * Booster parameters depend on which booster you have chosen
# * Learning task parameters decide on the learning scenario. For example, regression tasks may use different parameters with ranking tasks.
# +
# instantiate params
params = {}
# general params
general_params = {'silent': 1}
params.update(general_params)
# booster params
n_gpus = 1
booster_params = {}
booster_params['max_depth'] = 8
booster_params['grow_policy'] = 'lossguide'
booster_params['max_leaves'] = 2**8
booster_params['tree_method'] = 'gpu_hist'
booster_params['n_gpus'] = 1 # keep this at 1, even if using more than 1 GPU - Dask XGBoost uses 1 GPU per worker
params.update(booster_params)
# learning task params
learning_task_params = {}
if classification:
learning_task_params['eval_metric'] = 'auc'
learning_task_params['objective'] = 'binary:logistic'
else:
learning_task_params['eval_metric'] = 'rmse'
learning_task_params['objective'] = 'reg:squarederror'
params.update(learning_task_params)
print(params)
# -
# <a id="train"></a>
# ## Train Model
#
# Now it's time to train our model! We can use the `dask_xgboost.train` function and pass in the parameters, training dataset, the number of boosting iterations, and the list of items to be evaluated during training.
# model training settings
num_round = 100
# +
# %%time
bst = dask_xgboost.train(client, params, X_train_dask_cudf, y_train_dask_cudf, num_boost_round=num_round)
# -
# <a id="predict"></a>
# ## Generate Predictions
#
# We can generated predictions using the `dask_xgboost.predict` method and then using `dask.dataframe.multi.concat` to concatenate the multiple resulting dataframes together.
y_predictions = dask_xgboost.predict(client, bst, X_validation_dask_cudf)
y_predictions = dask.dataframe.multi.concat([y_predictions], axis=1)
# <a id="evaluate"></a>
# ## Evaluate Model
#
# Lastly, we can evaluate our model (depending on classification or regression) and calculate accuracy or rmse, respectively.
# +
from sklearn.metrics import accuracy_score
if classification:
thresholded_predictions = (y_predictions[0] > 0.5).compute().to_array() * 1.0
accuracy = accuracy_score(y_validation, thresholded_predictions)
print('Accuracy:', accuracy)
else:
test['squared_error'] = (y_predictions[0] - y_validation_dask_cudf['y'])**2
rmse = np.sqrt(test.squared_error.mean().compute())
print('Root Mean Squared Error:', rmse)
# -
# <a id="conclusion"></a>
# ## Conclusion
#
# In this notebook, we showed how to work with Dask XGBoost in RAPIDS.
#
# To learn more about RAPIDS, be sure to check out:
#
# * [Open Source Website](http://rapids.ai)
# * [GitHub](https://github.com/rapidsai/)
# * [Press Release](https://nvidianews.nvidia.com/news/nvidia-introduces-rapids-open-source-gpu-acceleration-platform-for-large-scale-data-analytics-and-machine-learning)
# * [NVIDIA Blog](https://blogs.nvidia.com/blog/2018/10/10/rapids-data-science-open-source-community/)
# * [Developer Blog](https://devblogs.nvidia.com/gpu-accelerated-analytics-rapids/)
# * [NVIDIA Data Science Webpage](https://www.nvidia.com/en-us/deep-learning-ai/solutions/data-science/)
| getting_started_materials/intro_tutorials_and_guides/08_Introduction_to_Dask_XGBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# %matplotlib inline
path = 'snb-data-fdiaustabsa-de-all-20171215_0900.xlsx'
df = pd.read_excel(path, na_values='-')
df
df.set_index('Sektoren')
df['Sektoren']
df['Industrie - Total']
df.dtypes
df["Sektoren"] = df['Sektoren'].astype('int')
df.dtypes
df['Beteiligungskapital'] = df.Beteiligungskapital.astype(float)
.astype)(int)
df['Beteiligungskapital'].astype(str).astype(int)
cdf = df.convert_objects(convert_numeric=True)
# +
data = {'Beteiligungskapital'}
df = pd.DataFrame(data, index = ['Kapitalart', 'Selektoren und Branchen','1998', '1999', '2000', '2001', '2001', '2002',
'2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011',
'2012', '2013', '2014', '2015', '2016'])
df
# -
df.Beteiligungskapital.value_counts().sort_index()
# +
data = {'Beteiligungskapital ohne reinvestierte Erträge': [2]}
df = pd.DataFrame(data, index = ['Kapitalart', 'Selektoren und Branchen','1998', '1999', '2000', '2001', '2001', '2002',
'2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011',
'2012', '2013', '2014', '2015', '2016'])
df
df_irn = df[data]
df_irn.head(3)
df_irn.data.value_counts().sort_index().plot()
# -
df.set_index('Jahre')
felderliste = ['Jahre', 'Beteiligungskapital ohne reinvestierte Erträge', 'Beteiligungskapital ohne reinvestierte Erträge.1',
'Beteiligungskapital ohne reinvestierte Erträge.2', 'Beteiligungskapital ohne reinvestierte Erträge.3']
felderliste = ['Jahre', 'Beteiligungskapital ohne reinvestierte Erträge', 'Beteiligungskapital ohne reinvestierte Erträge.1',
'Beteiligungskapital ohne reinvestierte Erträge.2', 'Beteiligungskapital ohne reinvestierte Erträge.3']
df[felderliste].head(5)
# +
df = pd.DataFrame({'Kapitalart': ['Selektoren und Branchen','1998', '1999', '2000', '2001', '2001', '2002',
'2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011',
'2012', '2013', '2014', '2015', '2016'],
'Beteiligungskapital ohne reinvestierte Erträge': ['6374']})
df.columns = ['Jahre',]
df
data = {'name': ['Jason', 'Molly', 'Tina', 'Jake', 'Amy'],
'year': [2012, 2012, 2013, 2014, 2014],
'reports': [4, 24, 31, 2, 3]}
df = pd.DataFrame(data, index = ['Cochice', 'Pima', '<NAME>', 'Maricopa', 'Yuma'])
df
# -
df.drop([1])
df_kapital..value_counts().sort_index().plot()
df_irn = df[df.Beteiligungskapital ohne reinvestierte Erträge.1 == 'IRN']
df_irn.head(3)
data = {'Beteiligungskapital ohne reinvestierte Erträge': ['']}
df = pd.DataFrame(data, index = ['Kapitalart', 'Sektoren und Branchen', 'Leerzeile',
'1998', '1999', '2000', '2001', '2002', '2003', '2004', '2205',
'2006', '2007', '2008', '2009', '2010', '2011', '2012',
'2013', '2014', '2015', '2016'])
df
df.drop(['Leerzeile'])
df.shape
# +
data = {}
df = pd.DataFrame(data, index = ['Kapitalart', 'Sektoren und Branchen', 'Leerzeile',
'1998', '1999', '2000', '2001', '2002', '2003', '2004', '2205',
'2006', '2007', '2008', '2009', '2010', '2011', '2012',
'2013', '2014', '2015', '2016'])
df
'name': ['Jason', 'Molly', 'Tina', 'Jake', 'Amy'],
'year': [2012, 2012, 2013, 2014, 2014],
'reports': [4, 24, 31, 2, 3]
# -
# +
data = {'name': ['Jason', 'Molly', 'Tina', 'Jake', 'Amy'],
'year': [2012, 2012, 2013, 2014, 2014],
'reports': [4, 24, 31, 2, 3]}
df = pd.DataFrame(data, index = ['Cochice', 'Pima', '<NAME>', 'Maricopa', 'Yuma'])
df
# -
df.dtypes
df.set_index('Kapitalart')
df = df.transpose()
df
df.set_index('Kapitalart')
data = ()
# setting first name as index column
data.set_index(["Kapitalart"], inplace = True,
append = True, drop = True)
df.set_index(['1998', '1999', '2000', '2001',
'2002', '2003', '2004', '2005', '2006',
'2007', '2008', '2009', '2010'])
| 000 Projekte_Juerg/00 Direktinvestitionen/Kopien/Direktinvestitionen CH-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Loading data into a dictionary
# <p>Netflix! What started in 1997 as a DVD rental service has since exploded into the largest entertainment/media company by <a href="https://www.marketwatch.com/story/netflix-shares-close-up-8-for-yet-another-record-high-2020-07-10">market capitalization</a>, boasting over 200 million subscribers as of <a href="https://www.cbsnews.com/news/netflix-tops-200-million-subscribers-but-faces-growing-challenge-from-disney-plus">January 2021</a>.</p>
# <p>Given the large number of movies and series available on the platform, it is a perfect opportunity to flex our data manipulation skills and dive into the entertainment industry.</p>
# <p>As evidence of this, they have provided us with the following information. For the years from 2011 to 2020, the average movie durations are 103, 101, 99, 100, 100, 95, 95, 96, 93, and 90, respectively.</p>
# +
# Create the years and durations lists
years = [2011, 2012, 2013, 2014, 2015, 2016, 2017, 2018, 2019, 2020]
durations = [103, 101, 99, 100, 100, 95, 95, 96, 93, 90]
# Create a dictionary with the two lists
movie_dict = {"years" : years, "durations" : durations}
# Print the dictionary
movie_dict
# -
# ## 2. Creating a DataFrame from a dictionary
# <p>To convert our dictionary <code>movie_dict</code> to a <code>pandas</code> DataFrame, we will first need to import the library under its usual alias. We'll also want to inspect our DataFrame to ensure it was created correctly.</p>
# +
# Import pandas under its usual alias
import pandas as pd
# Create a DataFrame from the dictionary
durations_df = pd.DataFrame(movie_dict)
# Print the DataFrame
durations_df
# -
# ## 3. A visual inspection of our data
# <p>Alright, we now have a <code>pandas</code> DataFrame, the most common way to work with tabular data in Python. Now back to the task at hand. We want to follow up on our friend's assertion that movie lengths have been decreasing over time. A great place to start will be a visualization of the data.</p>
# <p>Given that the data is continuous, a line plot would be a good choice, with the dates represented along the x-axis and the average length in minutes along the y-axis. This will allow us to easily spot any trends in movie durations. There are many ways to visualize data in Python, but <code>matploblib.pyplot</code> is one of the most common packages to do so.</p>
# +
# Import matplotlib.pyplot under its usual alias and create a figure
import matplotlib.pyplot as plt
fig = plt.figure()
# Draw a line plot of release_years and durations
plt.plot(years, durations)
# Create a title
plt.title('Netflix Movie Durations 2011-2020')
# Show the plot
plt.show()
# -
# ## 4. Loading the rest of the data from a CSV
# <p>Well, it looks like there is something to the idea that movie lengths have decreased over the past ten years! There are a few questions about this trend that we are currently unable to answer, including:</p>
# <ol>
# <li>What does this trend look like over a longer period of time?</li>
# <li>Is this explainable by something like the genre of entertainment?</li>
# </ol>
# <p>We now have access to the CSV file, available at the path <code>"datasets/netflix_data.csv"</code>. Let's create another DataFrame, this time with all of the data. Given the length of our friend's data, printing the whole DataFrame is probably not a good idea, so we will inspect it by printing only the first five rows.</p>
# +
# Read in the CSV as a DataFrame
netflix_df = pd.read_csv("datasets/netflix_data.csv")
# Print the first five rows of the DataFrame
netflix_df.head()
# -
# ## 5. Filtering for movies!
# <p>Okay, we have our data! Now we can dive in and start looking at movie lengths. </p>
# <p>Or can we? Looking at the first five rows of our new DataFrame, we notice a column <code>type</code>. Scanning the column, it's clear there are also TV shows in the dataset! Moreover, the <code>duration</code> column we planned to use seems to represent different values depending on whether the row is a movie or a show (perhaps the number of minutes versus the number of seasons)?</p>
# <p>Fortunately, a DataFrame allows us to filter data quickly, and we can select rows where <code>type</code> is <code>Movie</code>. While we're at it, we don't need information from all of the columns, so let's create a new DataFrame <code>netflix_movies</code> containing only <code>title</code>, <code>country</code>, <code>genre</code>, <code>release_year</code>, and <code>duration</code>.</p>
# +
# Subset the DataFrame for type "Movie"
netflix_df_movies_only = netflix_df[netflix_df["type"] == "Movie"]
# Select only the columns of interest
netflix_movies_col_subset = netflix_df_movies_only[['title', 'country', 'genre', 'release_year', 'duration']]
# Print the first five rows of the new DataFrame
netflix_movies_col_subset.head()
# -
# ## 6. Creating a scatter plot
# <p>Okay, now we're getting somewhere. We've read in the raw data, selected rows of movies, and have limited our DataFrame to our columns of interest. Let's try visualizing the data again to inspect the data over a longer range of time.</p>
# <p>This time, we are no longer working with aggregates but instead with individual movies. A line plot is no longer a good choice for our data, so let's try a scatter plot instead. We will again plot the year of release on the x-axis and the movie duration on the y-axis.</p>
# +
# Create a figure and increase the figure size
fig = plt.figure(figsize=(12,8))
# Create a scatter plot of duration versus year
plt.scatter(netflix_movies_col_subset['release_year'], netflix_movies_col_subset['duration'])
# Create a title
plt.title("Movie Duration by Year of Release")
# Show the plot
plt.show()
# -
# ## 7. Digging deeper
# <p>This is already much more informative than the simple plot we created when our friend first gave us some data. We can also see that, while newer movies are overrepresented on the platform, many short movies have been released in the past two decades.</p>
# <p>Upon further inspection, something else is going on. Some of these films are under an hour long! Let's filter our DataFrame for movies with a <code>duration</code> under 60 minutes and look at the genres. This might give us some insight into what is dragging down the average.</p>
# +
# Filter for durations shorter than 60 minutes
short_movies = netflix_movies_col_subset[netflix_movies_col_subset["duration"] < 60]
# Print the first 20 rows of short_movies
short_movies.head(20)
# -
# ## 8. Marking non-feature films
# <p>Interesting! It looks as though many of the films that are under 60 minutes fall into genres such as "Children", "Stand-Up", and "Documentaries". This is a logical result, as these types of films are probably often shorter than 90 minute Hollywood blockbuster. </p>
# <p>We could eliminate these rows from our DataFrame and plot the values again. But another interesting way to explore the effect of these genres on our data would be to plot them, but mark them with a different color.</p>
# <p>In Python, there are many ways to do this, but one fun way might be to use a loop to generate a list of colors based on the contents of the <code>genre</code> column. Much as we did in Intermediate Python, we can then pass this list to our plotting function in a later step to color all non-typical genres in a different color!</p>
# +
# Define an empty list
colors = []
# Iterate over rows of netflix_movies_col_subset
for lab, row in netflix_movies_col_subset.iterrows():
if row['genre'] == "Children":
colors.append("red")
elif row['genre'] == 'Documentaries':
colors.append("blue")
elif row['genre'] == 'Stand-Up':
colors.append("green")
else:
colors.append("black")
# Inspect the first 10 values in your list
print(colors[:10])
# -
# ## 9. Plotting with color!
# <p>Lovely looping! We now have a <code>colors</code> list that we can pass to our scatter plot, which should allow us to visually inspect whether these genres might be responsible for the decline in the average duration of movies.</p>
# <p>This time, we'll also spruce up our plot with some additional axis labels and a new theme with <code>plt.style.use()</code>. The latter isn't taught in Intermediate Python, but can be a fun way to add some visual flair to a basic <code>matplotlib</code> plot.</p>
# +
# Set the figure style and initalize a new figure
plt.style.use('fivethirtyeight')
fig = plt.figure(figsize=(12,8))
# Create a scatter plot of duration versus release_year
plt.scatter(netflix_movies_col_subset["release_year"], netflix_movies_col_subset["duration"], c = colors)
# Create a title and axis labels
plt.title("Movie duration by year of release")
plt.xlabel("Release year")
plt.ylabel("Duration (min)")
# Show the plot
plt.show()
# -
# Well, as we suspected, non-typical genres such as children's movies and documentaries are all clustered around the bottom half of the plot. But we can't know for certain until we perform additional analyses.
| notebook1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
'''Handles analytically deconvoving two Gaussians and finding of a common beam.'''
def quadratic2elliptic(A,B,C,D=0,E=0,F=-np.log(2)):
"""Invert:
(A0 cos^2 phi + c0 sin^2 phi)k = A
(A0-C0)sin 2phi k = B
(a0 sin^2 phi + c0 cos^2 phi) k = C
returns bmaj,bmin,bpa[,xc,y if D,E != 0]"""
if (np.isinf(A) and np.isinf(B) and np.isinf(C)):#delta function
return 0., 0., 0.
assert (B**2 - 4*A*C) != 0, "It is parabolic, not elliptic or hyperbolic"
if A!=C:#not a circle so should be able to solve second equation
#A-C = A0 cos^2 phi + c0 sin^2 phi - a0 sin^2 phi - c0 cos^2 phi
#A-C = A0 (cos^2 phi - sin^2 phi)- c0 (-sin^2 phi + c0 cos^2 phi) = (A0 - C0) cos 2phi
#(cos^2 phi - sin^2 phi) = cos 2 phi
phi = np.arctan2(B,(A-C))/2.#choose your own modulus
else:#circle
phi = np.pi/4.#arbitrary, nonzero cos and sin for the rest
phi = 0.
#rotate x,y to phi to x',y' = xcos - ysin, xsin + ycos
#then expand A(x' - xc)^2 + B(x'-xc)(y'-yc) + C(y' - yc)^2 + D(x' - xc) + E(y' - yc) + F = 0
#then now the coord sys is unrotated relative to the quadratic paramters
c = np.cos(phi)
s = np.sin(phi)
c2 = c*c
s2 = s*s
A1 = A*c2 + B*c*s + C*s2
B1 = 2.*(C-A)*s*c+B*(c2-s2)#should be zero since there's no cross term in the unroated
#print "Rotated cross term: {0} =? 0".format(B1)
C1 = A*s2 - B*c*s + C*c2
D1 = D*c + E*s
E1 = -D*s + E*c
assert (A1 != 0) and (C1 != 0), "degenerate between ellipse and hyperbola"
#complete square for x's and y's
#A1(x-xc)^2 + C1(y-yc)^2 + F = A1xx - 2A1xxc + A1xcxc + C1yy - 2C1yyc + C1ycyc = A1() + D1() + C1() + E1() + A1xcxc + C1ycyc + F =0
xc1 = D1/(-2.*A1)
yc1 = E1/(-2.*C1)#solutions (in rotated frame)
#now unrotate them
xc = xc1*c - yc1*s#might be xc1*c + yc1*s
yc = xc1*s + yc1*c#might be -xc1*s + yc1*c
#bring the remainder of completed squares to rhs with F
rhs = -F + D1**2/(4.*A1) + E1**2/(4.*C1)
#(x-xc)^2/(bmaj/2)^2 + (y-yc)^2/(bmin/2)^2 = 1
#A1(x-xc)^2 + C1*y-yc)^2 = rhs
A0 = A1/rhs
C0 = C1/rhs
bmaj = np.sign(A0)*2.*np.sqrt(1./np.abs(A0))
bmin = np.sign(C0)*2.*np.sqrt(1./np.abs(C0))
assert bmaj*bmin > 0, "Hyperbolic solution ;) inversion success but not physical."
#return None,None,None
if bmin > bmaj:
temp = bmin
bmin = bmaj
bmaj = temp
bpa = phi# - np.pi/2.#starts at y
if E==0 and D==0:
return bmaj,bmin,bpa*180./np.pi
return bmaj,bmin,bpa*180./np.pi,xc,yc
def elliptic2quadratic(bmaj,bmin,pa,xc=0,yc=0,k=np.log(2)):
'''a*x**2 + b*x*y + c*y**2 + d*x + e*y + f = 0
pa in deg
return A,B,C[,D,E,F if xc,yc!=0]'''
#unrotated solution
a0 = k/(bmaj/2.)**2
c0 = k/(bmin/2.)**2
theta = (pa + 90.)*np.pi/180.
#Rotated Solution
cos2 = np.cos(theta)**2
sin2 = np.sin(theta)**2
A = (a0*cos2 + c0*sin2)
C = (c0*cos2 + a0*sin2)
B = (a0 - c0 )*np.sin(2.*theta)
#Now move center
D = -2.*A*xc - B*yc
E = -2.*C*yc - B*xc
F = A*xc**2 + B*xc*yc + C*yc**2 - 1./k
if xc==0 and yc==0:
return A,B,C
return A,B,C,D,E,F
def deconvolve(A1,B1,C1,A2,B2,C2):
'''Solves analytically G(A1,B1,C1) = convolution(G(A2,B2,C2), G(Ak,Bk,Ck))
Returns Ak,Bk,Ck
A,B,C are quadratic parametrization.
If you have bmaj,bmin,bpa, then get A,B,C = ecliptic2quadratic(0,0,bmaj,bmin,bpa)
Returns (np.inf,np.inf,np.inf) if solution is delta function'''
D = B1**2 - 2*B1*B2 + B2**2 - 4*A1*C1 + 4* A2* C1 + 4* A1* C2 - 4* A2* C2
if (np.abs(D) < 10*(1-2./3.-1./3.)):
return np.inf,np.inf,np.inf#delta function
if (D<0.):
#print "Inverse Gaussian, discriminant D:",D
#ie. hyperbolic solution, still valid but elliptic representation is impossible instead you get hyperbolic parameters: negative bmaj/bmin
pass
Ak = (-A2* B1**2 + A1* B2**2 + 4* A1* A2* C1 - 4* A1* A2* C2)/D
Bk = (-B1**2 *B2 + B1* B2**2 + 4* A1* B2* C1 - 4* A2* B1* C2)/D
Ck = (B2**2 *C1 - B1**2 *C2 + 4* A1* C1* C2 - 4* A2* C1* C2)/D
assert (Bk*Bk - 4*Ak*Ck) != 0, "Indifinite deconvolution det = 0"
return Ak,Bk,Ck
def convolve(A1,B1,C1,A2,B2,C2):
'''
Convolves two gaussians with quadratic parametrization:
A,B,C are quadratic parametrization.
If you have bmaj,bmin,bpa, then get A,B,C = elliptic2quadratic(0,0,bmaj,bmin,bpa)
Where g = factor*Exp(-A*X**2 - B*X*Y - C*Y**2)
'''
D1 = 4.*A1*C1 - B1**2
D2 = 4.*A2*C2 - B2**2
D3 = -2.*B1 * B2 + 4.*A2*C1 + 4.*A1*C2 + D1+D2
D4 = C2*D1+C1*D2
#Non-solvable cases
if (D1*D2*D3*D4 == 0):
print ("Can't convolve...")
return (None,None,None)
if (D3 < 0):#always imaginary
print ("D3 < 0, Imaginary solution",D3)
return (None,None,None)
factor = 2.*np.pi*np.sqrt(D1 + 0j)*np.sqrt(D2 + 0j)/np.sqrt(D3/D4 + 0j)/np.sqrt(D4/(D1*D2) + 0j)
if np.abs(np.imag(factor)) > 10.*(7./3 - 4./3 - 1.):
print ("Imaginary result somehow...")
return (None,None,None)
factor = np.real(factor)
A = (A2*D1 + A1 * D2)/D3
B = (B2*D1+B1*D2)/D3
C = D4/D3
k = np.log(factor*2.)
return A,B,C#,factor
def findCommonBeam(beams, debugplots=False,confidence=0.005):
'''Given a list `beams` where each element of beams is a list of elliptic beam parameters (bmaj_i,bmin_i, bpa_i)
with bpa in degrees
return the beam parameters of the common beam of minimal area.
Common beam means that all beams can be convolved to the common beam.
`confidence` parameter is basically how confident you want solution. So 0.01 is knowing solution to 1%.
Specifically it's how long to sample so that there are enough statistics to properly sample likelihood with required accuracy.
default is 0.005. Computation time scale inversely with it.'''
def beamArea(bmaj,bmin,bpa=None):
return bmaj*bmin*np.pi/4./np.log(2.)
def isCommonBeam(beamCandQuad,beamsQuad):
for beamQuad in beamsQuad:
try:
Ak,Bk,Ck = deconvolve(*beamCandQuad,*beamQuad)
bmaj,bmin,bpa = quadratic2elliptic(Ak,Bk,Ck)
except:
return False
return True
def samplePrior(beamLast, beamsQuad):
iter = 0
while True:
std = 1.5
beam = [beamLast[0]*np.exp(np.log(std)*np.random.uniform(low=-1,high=1.)),
beamLast[1]*np.exp(np.log(std)*np.random.uniform(low=-1,high=1.)),
beamLast[2] + np.random.uniform(low=-5,high=5)]
#beam[0] = np.abs(beam[0])
#beam[1] = np.abs(beam[1])
if beam[1] > beam[0]:
temp = beam[1]
beam[1] = beam[0]
beam[0] = temp
while beam[2] > 90.:
beam[2] -= 180.
while beam[2] < -90.:
beam[2] += 180.
A,B,C = elliptic2quadratic(*beam)
if isCommonBeam((A,B,C),beamsQuad):
return beam
iter += 1
def misfit(beam,areaLargest):
area = beamArea(*beam)
L2 = (area - areaLargest)**2/2.
return L2
#Get beam areas
N = len(beams)
areas = []
beamsQuad = []
i = 0
while i < N:
areas.append(beamArea(*beams[i]))
beamsQuad.append(elliptic2quadratic(*beams[i]))
i += 1
beam0 = beams[np.argmax(areas)]
areaLargest = np.max(areas)
beam0Quad = elliptic2quadratic(*beam0)
if isCommonBeam(beam0Quad,beamsQuad):
return beam0
else:
bmajMax = np.max(beams,axis=0)[0]
beam0 = [bmajMax,bmajMax,0.]
#MC search, 1/binning = confidence
binning = int(1./confidence)
Nmax = 1e6
beamsMH = np.zeros([binning*binning,3],dtype=np.double)
beamsMul = np.zeros(binning*binning,dtype=np.double)
beamsMH[0,:] = beam0
beamsMul[0] = 1
accepted = 1
Si = misfit(beam0,areaLargest)
Li = np.exp(-Si)
maxL = Li
maxLBeam = beam0
iter = 0
while accepted < binning**2 and iter < Nmax:
beam_j = samplePrior(beamsMH[accepted-1], beamsQuad)
Sj = misfit(beam_j,areaLargest)
Lj = np.exp(-Sj)
#print("Sj = {}".format(Sj))
if Sj < Si or np.log(np.random.uniform()) < Si - Sj:
Si = Sj
beamsMH[accepted,:] = beam_j
beamsMul[accepted] += 1
#print("Accepted")
accepted += 1
else:
beamsMul[accepted-1] += 1
if Lj > maxL:
maxL = Lj
maxLBeam = beam_j
iter += 1
if accepted == binning**2:
pass
#print("Converged in {} steps with an acceptance rate of {}".format(iter,float(accepted)/iter))
else:
beamsMH = beamsMH[:iter,:]
beamsMul = beamsMul[:iter]
if debugplots:
import pylab as plt
# plt.hist(beamsMH[:,0],bins=binning)
# plt.show()
# plt.hist(beamsMH[:,1],bins=binning)
# plt.show()
# plt.hist(beamsMH[:,2],bins=binning)
# plt.show()
from matplotlib.patches import Ellipse
ax = plt.subplot(1,1,1)
ax.add_artist(Ellipse(xy=(0,0), width=maxLBeam[0], height=maxLBeam[1], angle=maxLBeam[2], facecolor="none",edgecolor='red',alpha=1,label='common beam'))
for beam in beams:
ax.add_artist(Ellipse(xy=(0,0), width=beam[0], height=beam[1], angle=beam[2], facecolor="none",edgecolor='black',ls='--',alpha=1))
ax.set_xlim(-0.5,0.5)
ax.set_ylim(-0.5,0.5)
plt.legend(frameon=False)
plt.show()
# meanBeam = np.sum(beamsMH.T*beamsMul,axis=1)/np.sum(beamsMul)
# stdBeam = np.sqrt(np.sum(beamsMH.T**2*beamsMul,axis=1)/np.sum(beamsMul) - meanBeam**2)
# print ("(Gaussian) beam is {} +- {}".format(meanBeam,stdBeam))
# logmeanBmajBmin = np.sum(np.log(beamsMH[:,:2]).T*beamsMul,axis=1)/np.sum(beamsMul)
# logstdBmajBmin = np.sqrt(np.sum(np.log(beamsMH[:,:2]).T**2*beamsMul,axis=1)/np.sum(beamsMul) - logmeanBmajBmin**2)
# logstdBmajBminu = np.exp(logmeanBmajBmin + logstdBmajBmin) - np.exp(logmeanBmajBmin)
# logstdBmajBminl = np.exp(logmeanBmajBmin) - np.exp(logmeanBmajBmin - logstdBmajBmin)
# logmeanBmajBmin = np.exp(logmeanBmajBmin)
# print("(Lognormal) bmaj/bmin is {} + {} - {}".format(logmeanBmajBmin,logstdBmajBminu,logstdBmajBminl))
# print ("Max Likelihood beam is {}".format(maxLBeam))
return maxLBeam
def fftGaussian(A,B,C,X,Y):
D = 4*A*C-B**2
return 2*np.pi/np.sqrt(D)*np.exp(-4*np.pi/D*(-C*X**2 +B*X*Y -A*Y**2))
def gaussian(A,B,C,X,Y):
return np.exp(-A*X**2 - B*X*Y - C*Y**2)
def psfTGSS1(dec):
'''input declination in degrees
return bmaj(arcsec), bmin(arcsec), bpa(degrees)'''
if dec > 19.0836824:
return 25.,25.,0.
else:
return 25.,25./np.cos(np.pi*(dec-19.0836824)/180.),0.
def test_elliptic2quadratic():
for i in range(100):
bpa = np.random.uniform()*180.-90.#deg
bmaj = np.random.uniform()
bmin = np.random.uniform()*bmaj
A,B,C = elliptic2quadratic(bmaj,bmin,bpa)
bmaj2,bmin2,bpa2 = quadratic2elliptic(A,B,C)
assert np.isclose(bmaj,bmaj2) and np.isclose(bmin,bmin2) and np.isclose(bpa,bpa2), "Failed to pass {},{},{} != {},{},{}".format(bmaj,bmin,bpa,bmaj2,bmin2,bpa2)
return True
def test_convolvedeconvolve(N=100):
for i in range(N):
bpa = np.random.uniform()*180.-90.#deg
bmaj = np.random.uniform()
bmin = np.random.uniform()*bmaj
A1,B1,C1 = elliptic2quadratic(bmaj,bmin,bpa)
bpa2 = np.random.uniform()*180.-90.#deg
bmaj2 = np.random.uniform()
bmin2 = np.random.uniform()*bmaj2
A2,B2,C2 = elliptic2quadratic(bmaj2,bmin2,bpa2)
Ac,Bc,Cc = convolve(A1,B1,C1,A2,B2,C2)
Ak,Bk,Ck = deconvolve(Ac,Bc,Cc,A1,B1,C1)
bmaj2_,bmin2_,bpa2_ = quadratic2elliptic(Ak,Bk,Ck)
assert np.isclose(bmaj2_,bmaj2) and np.isclose(bmin2_,bmin2) and np.isclose(bpa2_,bpa2), "Failed to pass {},{},{} != {},{},{}".format(bmaj2_,bmin2_,bpa2_,bmaj2,bmin2,bpa2)
return True
def test_deltaFunctionDeconvolve():
bpa = np.random.uniform()*180.-90.#deg
bmaj = np.random.uniform()
bmin = np.random.uniform()*bmaj
A1,B1,C1 = elliptic2quadratic(bmaj,bmin,bpa)
#deconv same beam
Ak,Bk,Ck = deconvolve(A1,B1,C1,A1,B1,C1)
bmaj_d, bmin_d, bpa_d = quadratic2elliptic(Ak,Bk,Ck)
assert bmaj_d==0 and bmin_d==0 and bpa_d==0,"Supposed to be the delta"
return True
def test_timing():
from time import clock
i = 0
t1 = clock()
for i in range(10000):
bpa = np.random.uniform()*180.-90.#deg
bmaj = np.random.uniform()
bmin = np.random.uniform()*bmaj
A1,B1,C1 = elliptic2quadratic(bmaj,bmin,bpa)
bpa2 = np.random.uniform()*180.-90.#deg
bmaj2 = np.random.uniform()
bmin2 = np.random.uniform()*bmaj2
A2,B2,C2 = elliptic2quadratic(bmaj2,bmin2,bpa2)
Ac,Bc,Cc = convolve(A1,B1,C1,A2,B2,C2)
Ak,Bk,Ck = deconvolve(Ac,Bc,Cc,A1,B1,C1)
bmaj2_,bmin2_,bpa2_ = quadratic2elliptic(Ak,Bk,Ck)
print("Time avg. ~ {} seconds".format((clock()-t1)/10000))
def test_findCommonBeam():
np.random.seed(1234)
for i in range(10):
beams = []
for i in range(3):
bpa = np.random.uniform()*180.-90.#deg
bmaj = np.random.uniform()
bmin = np.random.uniform()*bmaj
beams.append((bmaj,bmin,bpa))
commonBeam = findCommonBeam(beams,debugplots=True)
print("Common beam amongst {} is {}".format(beams,commonBeam))
if __name__ == '__main__':
# test_elliptic2quadratic()
# test_convolvedeconvolve()
# test_deltaFunctionDeconvolve()
# test_timing()
test_findCommonBeam()
# -
| src/rathings/notebooks/BeamDeconvolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys, os
sys.path.insert(0, os.path.abspath('..'))
from scripts import project_functions
csv = "Electric_Vehicle_Population_Data.csv"
df = project_functions.load_and_process(csv)
# -
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df.shape
df.head()
df.columns
df.nunique(axis=0)
corr = df.corr()# plot the heatmap
sns.heatmap(corr, xticklabels=corr.columns, yticklabels=corr.columns, annot=True, cmap=sns.diverging_palette(220, 20, as_cmap=True))
df.plot(kind='scatter', x='Longitude', y='Latitude')
ax = sns.countplot(y = df["EV_Type"], data = df).set_title("Number of Electric Vehicles in Washington by Electric Vehicle Types")
fig, ax1 = plt.subplots()
fig.set_size_inches(10, 9)
ax1 = sns.countplot(y = df["Make"], data = df).set_title("Number of Electric Vehicles in Washington by EV Make")
df1 = df[(df['Latitude'] > 45) & (df['Longitude'] < -115)]
sns.jointplot(x="Latitude", y="Longitude", kind="hex", data=df1, color="#4571c5")
df1.plot(kind='scatter', x='Longitude', y='Latitude')
hist = df.hist(bins=10,figsize =(10,10))
boxplot = sns.boxplot(x = df1['Longitude'])
boxplot1 = sns.boxplot(x = df1['Latitude'])
| Analysis/Michael/Michael_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## simple ResNet
import torch
import numpy as np
from torch import nn
from torch.nn import functional
import matplotlib.pyplot as plt
import ipywidgets
# +
# number of features in
nf = 2
# number of classes
nClass = 2
X = torch.randn(2,1500)
R = torch.sqrt(X[0,:]**2 + X[1,:]**2)
indRed = (R < 0.9).nonzero()
indBlue = (R >= 0.9).nonzero()
C = torch.zeros(1500,dtype = torch.int64)
C[indRed] = 0
C[indBlue] = 1
plt.plot(X.numpy()[0,indRed],X.numpy()[1,indRed],'.C3')
plt.plot(X.numpy()[0,indBlue],X.numpy()[1,indBlue],'.C0')
# +
# Get a validation set
XVal = torch.randn(2,1500)
R = torch.sqrt(XVal[0,:]**2 + XVal[1,:]**2)
indRed = (R < 0.9).nonzero()
indBlue = (R >= 0.9).nonzero()
CVal = torch.zeros(1500,dtype = torch.int64)
CVal[indRed] = 0
CVal[indBlue] = 1
plt.plot(XVal.numpy()[0,indRed],XVal.numpy()[1,indRed],'.C3')
plt.plot(XVal.numpy()[0,indBlue],XVal.numpy()[1,indBlue],'.C0')
# +
lossFunc = nn.CrossEntropyLoss()
def misfit(X, W, bias, C):
n = W.shape
X = X.view(n[1], -1)
S = torch.matmul(W, X)
S += bias.unsqueeze(1)
return lossFunc(S.t(),C), S
# +
class ResNet(nn.Module):
def __init__(self, nt=64, dt=0.1, gamma=1e-3):
super().__init__()
self.nt = nt
self.dt = dt
self.gamma = gamma
def forward(self, x, K, b, K0=None):
if K0 is not None:
x = torch.matmul(K0, x)
# Allocate space for all times
X = torch.zeros(x.shape[0], x.shape[1], self.nt+1)
X[:, :, 0] = x
# forward Euler
for i in range(self.nt):
Ki = self.M_imag_eigs(K[i])
z = torch.matmul(Ki, x)
x = x + self.dt * functional.relu(z+b[i])
ind = i+1
X[:, :, ind] = x
# k1 = functional.relu(torch.matmul(K[i], x) + b[i])
# k2 = functional.relu(torch.matmul(K[i], x+k1/2.0) + b[i])
# k3 = functional.relu(torch.matmul(K[i], x+k2/2.0) + b[i])
# k4 = functional.relu(torch.matmul(K[i], x+k3) + b[i])
# x = x + self.dt/6.0*(k1+k2+k3+k4)
# ind = i+1
# X[:,:,ind] = x
return x, X
def M_imag_eigs(self, K):
return 0.5*(K - K.t() - self.gamma*torch.eye(K.shape[0]))
# +
dt = 0.1
nt = 64
nf_net = 3
net = ResNet(dt=dt, nt=nt, gamma=1e-2)
# initialize K's
K0 = nn.Parameter(torch.randn(nf_net, nf), requires_grad=True)
K = []
for i in range(nt):
Ki = nn.Parameter(torch.randn(nf_net, nf_net)*1e-3, requires_grad=True)
K.append(Ki)
b = nn.Parameter(torch.randn(nt)*1e-3)
# -
# Run the network
xF, Xpath = net(X,K,b,K0)
# +
# weights for linear classifier
W = nn.Parameter(torch.randn(nClass, nf_net)*1e-3, requires_grad=True)
bW = nn.Parameter(torch.randn(nClass)*1e-2, requires_grad=True)
nparout = np.prod(W.shape) + np.prod(bW.shape) + np.prod(K[0].shape)*nt + nt
print('Total number of parameter',nparout)
print('Total number of data',C.shape[0])
# -
# run the network
x, _ = net(X, K, b, K0)
label = C
loss, S = misfit(x, W, bW, label)
print('Initial loss = ', loss.detach().numpy())
print('Sanity Check = ', np.log(2.0))
batchSize = 16
import torch.optim as optim
optimizer = optim.SGD(
[{'params': W}, {'params': K}, {'params': K0}, {'params': bW}, {'params': b}],
lr = 1e0, momentum=0
)
# +
for epoch in range(30): # loop over the dataset multiple times
# zero the parameter gradients
g = 0.0
loss = 0.0
ind = 0
while ind < X.shape[1]:
optimizer.zero_grad()
# get the inputs
inputs = X[:,ind:ind+batchSize]
labels = C[ind:ind+batchSize]
# forward
x,_ = net(inputs, K, b, K0)
lossi, _ = misfit(x, W, bW, labels)
if i==0:
loss = lossi
else:
loss += lossi
lossi.backward()
optimizer.step()
g += torch.norm(W.grad) + torch.norm(bW.grad)
ind += batchSize
print('%d %.3f %.3f'% (epoch+1, loss, g))
print('Finished Training')
# -
S
# +
x,_ = net(XVal, K, b, K0)
S = torch.matmul(W,x)
S += bW.unsqueeze(1)
Cpred = functional.softmax(S.t(), dim=0)
_,Cpred = torch.max(S,dim=0)
label = C
loss, S = misfit(x, W, bW, label)
print('Validation loss = ', loss.detach().numpy())
indRed = (Cpred==0).nonzero()
indBlue = (Cpred==1).nonzero()
plt.plot(XVal[0,indRed].numpy(),XVal[1,indRed].numpy(),'.C3')
plt.plot(XVal[0,indBlue].numpy(),XVal[1,indBlue].numpy(),'.C0')
# +
# plot the path
x, Xpath = net(X, K, b, K0)
indRed = (C==0).nonzero()
indBlue = (C==1).nonzero()
from mpl_toolkits.mplot3d import Axes3D
def plot_path(i):
if nf_net == 3:
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
Xi = Xpath[:, :, i].detach().numpy()
ax.scatter(Xi[0,indRed], Xi[1,indRed], Xi[2,indRed], color='C3')
ax.scatter(Xi[0,indBlue], Xi[1,indBlue], Xi[2,indBlue], color='C0')
elif nf_net == 2:
fig, ax = plt.subplots(1, 1)
Xi = Xpath[:, :, i].detach().numpy()
ax.scatter(Xi[0,indRed], Xi[1,indRed], color='C3')
ax.scatter(Xi[0,indBlue], Xi[1,indBlue], color='C0')
ax.set_title("Time {}".format(i))
ipywidgets.interact(
plot_path, i=ipywidgets.IntSlider(min=0, max=nt, value=0)
)
# +
tmp = x.grad_fn
print(x.grad_fn)
tmp1, tmp2 = tmp.next_functions
print(tmp.next_functions)
print(tmp1[0].next_functions, tmp2[0].next_functions)
print("\n")
print(tmp1[0].next_functions[0][0].next_functions)
print(tmp1[0].next_functions[1][0].next_functions)
print(tmp2[0].next_functions[0][0].next_functions)
# -
x = torch.randn(2, requires_grad=True)
x
f = functional.relu(x)
f
| SimpleResNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3_ModelFitting/student/W1D3_Tutorial6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 6: Model Selection: Cross-validation
# **Week 1, Day 3: Model Fitting**
#
# **By Neuromatch Academy**
#
# **Content creators**: <NAME>, <NAME>, <NAME> with help from <NAME>
#
# **Content reviewers**: <NAME>, <NAME>, <NAME>
#
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Tutorial Objectives
#
# *Estimated timing of tutorial: 25 minutes*
#
# This is Tutorial 6 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
#
# Tutorial objectives:
# * Implement cross-validation and use it to compare polynomial regression model
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/2mkq4/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
# # Setup
# + cellView="both"
# Imports
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
# + cellView="form"
#@title Figure Settings
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form"
# @title Plotting Functions
def plot_cross_validate_MSE(mse_all):
""" Plot the MSE values for the K_fold cross validation
Args:
mse_all (ndarray): an array of size (number of splits, max_order + 1)
"""
plt.figure()
plt.boxplot(mse_all, labels=np.arange(0, max_order + 1))
plt.xlabel('Polynomial Order')
plt.ylabel('Validation MSE')
plt.title(f'Validation MSE over {n_splits} splits of the data')
def plot_AIC(order_list, AIC_list):
""" Plot the AIC value for fitted polynomials of various orders
Args:
order_list (list): list of fitted polynomial orders
AIC_list (list): list of AIC values corresponding to each polynomial model on order_list
"""
plt.bar(order_list, AIC_list)
plt.ylabel('AIC')
plt.xlabel('polynomial order')
plt.title('comparing polynomial fits')
plt.show()
# + cellView="form"
# @title Helper Functions
def ordinary_least_squares(x, y):
"""Ordinary least squares estimator for linear regression.
Args:
x (ndarray): design matrix of shape (n_samples, n_regressors)
y (ndarray): vector of measurements of shape (n_samples)
Returns:
ndarray: estimated parameter values of shape (n_regressors)
"""
return np.linalg.inv(x.T @ x) @ x.T @ y
def make_design_matrix(x, order):
"""Create the design matrix of inputs for use in polynomial regression
Args:
x (ndarray): input vector of shape (n_samples)
order (scalar): polynomial regression order
Returns:
ndarray: design matrix for polynomial regression of shape (samples, order+1)
"""
# Broadcast to shape (n x 1)
if x.ndim == 1:
x = x[:, None]
#if x has more than one feature, we don't want multiple columns of ones so we assign
# x^0 here
design_matrix = np.ones((x.shape[0], 1))
# Loop through rest of degrees and stack columns
for degree in range(1, order + 1):
design_matrix = np.hstack((design_matrix, x**degree))
return design_matrix
def solve_poly_reg(x, y, max_order):
"""Fit a polynomial regression model for each order 0 through max_order.
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
max_order (scalar): max order for polynomial fits
Returns:
dict: fitted weights for each polynomial model (dict key is order)
"""
# Create a dictionary with polynomial order as keys, and np array of theta
# (weights) as the values
theta_hats = {}
# Loop over polynomial orders from 0 through max_order
for order in range(max_order + 1):
X = make_design_matrix(x, order)
this_theta = ordinary_least_squares(X, y)
theta_hats[order] = this_theta
return theta_hats
def evaluate_poly_reg(x, y, theta_hats, max_order):
""" Evaluates MSE of polynomial regression models on data
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
theta_hat (dict): fitted weights for each polynomial model (dict key is order)
max_order (scalar): max order of polynomial fit
Returns
(ndarray): mean squared error for each order, shape (max_order)
"""
mse = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
X_design = make_design_matrix(x, order)
y_hat = np.dot(X_design, theta_hats[order])
residuals = y - y_hat
mse[order] = np.mean(residuals ** 2)
return mse
# -
# ---
# # Section 1: Cross-validation
#
#
# + cellView="form"
# @title Video 1: Cross-Validation
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1mt4y1Q7C4", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="OtKw0rSRxo4", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# We now have multiple choices for which model to use for a given problem: we could use linear regression, order 2 polynomial regression, order 3 polynomial regression, etc. As we saw in Tutorial 5, different models will have different quality of predictions, both on the training data and on the test data.
#
# A commonly used method for model selection is to asks how well the model predicts new data that it hasn't seen yet. But we don't want to use test data to do this, otherwise that would mean using it during the training process! One approach is to use another kind of held-out data which we call **validation data**: we do not fit the model with this data but we use it to select our best model.
#
# We often have a limited amount of data though (especially in neuroscience), so we do not want to further reduce our potential training data by reassigning some as validation. Luckily, we can use **k-fold cross-validation**! In k-fold cross validation, we divide up the training data into k subsets (that are called *folds*, see diagram below), train our model on the first k-1 folds, and then compute error on the last held-out fold. We can then repeat this process k times, once on each k-1 folds of the data. Each of these k instances (which are called *splits*, see diagram below) excludes a different fold from fitting. We then average the error of each of the k trained models on its held-out subset - this is the final measure of performance which we can use to do model selection.
#
# To make this explicit, let's say we have 1000 samples of training data and choose 4-fold cross-validation. Samples 0 - 250 would be subset 1, samples 250 - 500 subset 2, samples 500 - 750 subset 3, and samples 750-1000 subset 4. First, we train an order 3 polynomial regression on subsets 1, 2, 3 and evaluate on subset 4. Next, we train an order 3 polynomial model on subsets 1, 2, 4 and evalute on subset 3. We continue until we have 4 instances of a trained order 3 polynomial regression model, each with a different subset as held-out data, and average the held-out error from each instance.
#
# We can now compare the error of different models to pick a model that generalizes well to held-out data. We can choose the measure of prediction quality to report error on the held-out subsets to suit our purposes. We will use MSE here but we could also use log likelihood of the data and so on.
#
# As a final step, it is common to retrain this model on all of the training data (without subset divisions) to get our final model that we will evaluate on test data. This approach allows us to evaluate the quality of predictions on new data without sacrificing any of our precious training data.
#
# Note that the held-out subsets are called either validation or test subsets. There is not a consensus and may depend on the exact use of k-fold cross validation. Sometimes people use k-fold cross validation to choose between different models/parameters to then apply to held-out test data and sometimes people report the averaged error on the held-out subsets as the model performance. If you are doing the former (using k-fold cross validation for model selection), you must report performance on held-out test data! In this text/code, we will refer to them as validation subsets to differentiate from our completely held-out test data (differing from the video above).
#
# These steps are summarized in this diagram from Scikit-learn (https://scikit-learn.org/stable/modules/cross_validation.html)
#
# 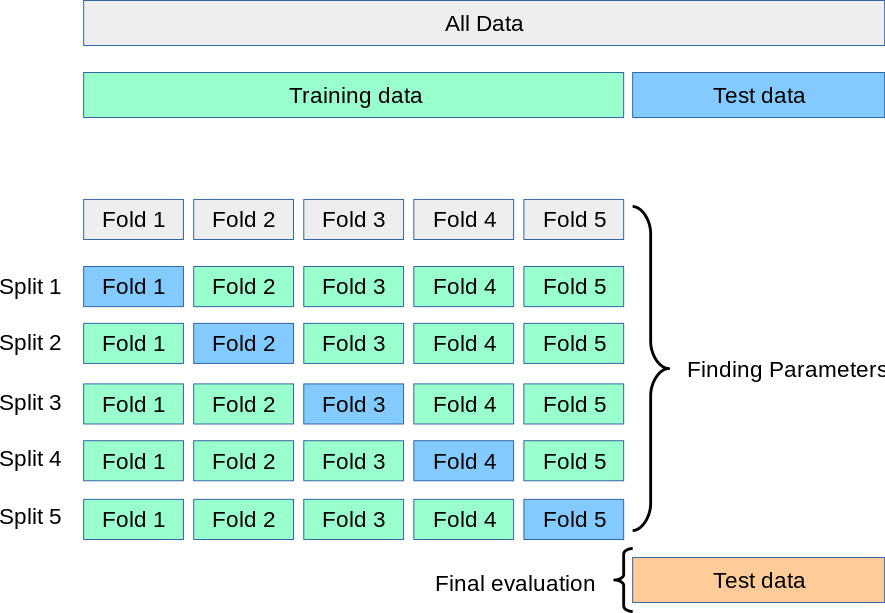
# Importantly, we need to be very careful when dividing the data into subsets. The held-out subset should not be used in any way to fit the model. We should not do any preprocessing (e.g. normalization) before we divide into subsets or the held-out subset could influence the training subsets. A lot of false-positives in cross-validation come from wrongly dividing.
#
# An important consideration in the choice of model selection method are the relevant biases. If we just fit using MSE on training data, we will generally find that fits get better as we add more parameters because the model will overfit the data, as we saw in Tutorial 5. When using cross-validation, the bias is the other way around. Models with more parameters are more affected by variance so cross-validation will generally prefer models with fewer parameters.
#
#
#
#
#
# We will again simulate some train and test data and fit polynomial regression models
#
# + cellView="form"
#@title
#@markdown Execute this cell to simulate data and fit polynomial regression models
### Generate training data
np.random.seed(0)
n_train_samples = 50
x_train = np.random.uniform(-2, 2.5, n_train_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_train_samples) # sample from a standard normal distribution
y_train = x_train**2 - x_train - 2 + noise
### Generate testing data
n_test_samples = 20
x_test = np.random.uniform(-3, 3, n_test_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_test_samples) # sample from a standard normal distribution
y_test = x_test**2 - x_test - 2 + noise
### Fit polynomial regression models
max_order = 5
theta_hats = solve_poly_reg(x_train, y_train, max_order)
# -
# ## Coding Exercise 1: Implement cross-validation
#
# Given our set of models to evaluate (polynomial regression models with orders 0 through 5), we will use cross-validation to determine which model has the best predictions on new data according to MSE.
#
# In this code, we split the data into 10 subsets using `Kfold` (from `sklearn.model_selection`). `KFold` handles cross-validation subset splitting and train/val assignments. In particular, the `Kfold.split` method returns an iterator which we can loop through. On each loop, this iterator assigns a different subset as validation and returns new training and validation indices with which to split the data.
#
# We will loop through the 10 train/validation splits and fit several different polynomial regression models (with different orders) for each split. You will need to use the `solve_poly_reg` method from Tutorial 4 and `evaluate_poly_reg` from Tutorial 5 (already implemented in this notebook).
#
# We will visualize the validation MSE over 10 splits of the data for each polynomial order using box plots.
help(solve_poly_reg)
help(evaluate_poly_reg)
# +
def cross_validate(x_train, y_train, max_order, n_splits):
""" Compute MSE for k-fold validation for each order polynomial
Args:
x_train (ndarray): training data input vector of shape (n_samples)
y_train (ndarray): training vector of measurements of shape (n_samples)
max_order (scalar): max order of polynomial fit
n_split (scalar): number of folds for k-fold validation
Return:
ndarray: MSE over splits for each model order, shape (n_splits, max_order + 1)
"""
# Initialize the split method
kfold_iterator = KFold(n_splits)
# Initialize np array mse values for all models for each split
mse_all = np.zeros((n_splits, max_order + 1))
for i_split, (train_indices, val_indices) in enumerate(kfold_iterator.split(x_train)):
# Split up the overall training data into cross-validation training and validation sets
x_cv_train = x_train[train_indices]
y_cv_train = y_train[train_indices]
x_cv_val = x_train[val_indices]
y_cv_val = y_train[val_indices]
#############################################################################
## TODO for students: Fill in missing ... in code below to choose which data
## to fit to and compute MSE for
# Fill out function and remove
raise NotImplementedError("Student exercise: implement cross-validation")
#############################################################################
# Fit models
theta_hats = ...
# Compute MSE
mse_this_split = ...
mse_all[i_split] = mse_this_split
return mse_all
# Cross-validate
max_order = 5
n_splits = 10
mse_all = cross_validate(x_train, y_train, max_order, n_splits)
# Visualize
plot_cross_validate_MSE(mse_all)
# + [markdown] cellView="both" colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D3_ModelFitting/solutions/W1D3_Tutorial6_Solution_ddce210a.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1116.0 height=828.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D3_ModelFitting/static/W1D3_Tutorial6_Solution_ddce210a_0.png>
#
#
# -
# Which polynomial order do you think is a better model of the data?
# ---
# # Summary
#
# *Estimated timing of tutorial: 25 minutes*
#
# We need to use model selection methods to determine the best model to use for a given problem.
#
# Cross-validation focuses on how well the model predicts new data.
# ---
# # Bonus
# ---
# ## Bonus Section 1: Akaike's Information Criterion (AIC)
#
# In order to choose the best model for a given problem, we can ask how likely the data is under a given model. We want to choose a model that assigns high probability to the data. A commonly used method for model selection that uses this approach is **Akaike’s Information Criterion (AIC)**.
#
# Essentially, AIC estimates how much information would be lost if the model predictions were used instead of the true data (the relative information value of the model). We compute the AIC for each model and choose the model with the lowest AIC. Note that AIC only tells us relative qualities, not absolute - we do not know from AIC how good our model is independent of others.
#
# AIC strives for a good tradeoff between overfitting and underfitting by taking into account the complexity of the model and the information lost. AIC is calculated as:
#
# \begin{align}
# \mathrm{AIC} = 2K - 2 \log(L)
# \end{align}
#
# where $K$ is the number of parameters in your model and $L$ is the likelihood that the model could have produced the output data.
#
# Now we know what AIC is, we want to use it to pick between our polynomial regression models. We haven't been thinking in terms of likelihoods though - so how will we calculate L?
#
# As we saw in Tutorial 2, there is a link between mean squared error and the likelihood estimates for linear regression models that we can take advantage of.
#
# *Derivation time!*
#
# We start with our formula for AIC from above:
#
# \begin{align} AIC = 2k - 2 \log L \end{align}
#
# For a model with normal errors, we can use the log likelihood of the normal distribution:
#
# \begin{align} \log L = -\frac{n}{2} \log(2 \pi) -\frac{n}{2}\log(\sigma^2) - \sum_i^N \frac{1}{2 \sigma^2} (y_i - \tilde y_i)^2
# \end{align}
#
# We can drop the first as it is a constant and we're only assessing relative information with AIC. The last term is actually also a constant: we don't know $\sigma^2$ in advance so we use the empirical estimate from the residual ($\hat{\sigma}^2 = 1/N\sum_i^N (y_i - \tilde y_i)^2$). Once we plug this in, the two $\sum [(y - \tilde y)^2]$ terms (in the numerator and denominator, respectively) cancel out and we are left with the last term as $\frac N 2$.
#
# Once we drop the constant terms and incorporate into the AIC formula we get:
#
# \begin{align}
# \mathrm{AIC} = 2k + n\log(\sigma^2)
# \end{align}
#
# We can replace $\sigma^2$ with the computation for variance (the sum of squared errors divided by number of samples). Thus, we end up with the following formula for AIC for linear and polynomial regression:
#
# \begin{align}
# \mathrm{AIC} = 2K + n \log(\frac{\mathrm{SSE}}{n})
# \end{align}
#
# where k is the number of parameters, n is the number of samples, and SSE is the summed squared error.
#
# ### Bonus Exercise 1: Compute AIC
# +
AIC_list = []
order_list = list(range(max_order + 1))
for order in order_list:
# Compute predictions for this model
X_design = make_design_matrix(x_train, order)
y_hat = np.dot(X_design, theta_hats[order])
#############################################################################
## TODO for students:
## to fit to and compute MSE for
# Fill out function and remove
raise NotImplementedError("Student exercise: implement compute AIC")
# 1) Compute sum of squared errors (SSE) given prediction y_hat and y_train
# 2) Identify number of parameters in this model (K in formula above)
# 3) Compute AIC (call this_AIC) according to formula above
#############################################################################
# Compute SSE
residuals = ...
sse = ...
# Get K
K = len(theta_hats[order])
# Compute AIC
AIC = ...
AIC_list.append(AIC)
# Visualize
plt.bar(order_list, AIC_list)
plt.ylabel('AIC')
plt.xlabel('polynomial order')
plt.title('comparing polynomial fits')
plt.show()
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D3_ModelFitting/solutions/W1D3_Tutorial6_Solution_16748857.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1120.0 height=832.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D3_ModelFitting/static/W1D3_Tutorial6_Solution_16748857_0.png>
#
#
# -
# Which model would we choose based on AIC?
| tutorials/W1D3_ModelFitting/student/W1D3_Tutorial6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: shopee
# language: python
# name: shopee
# ---
import pandas as pd
from tqdm import tqdm_notebook, tqdm
tqdm.pandas()
df_train = pd.read_csv('/Users/cenk/data-sets/shopee/shopee-product-matching/train.csv')
df_train.head()
df_products = df_train.groupby(['label_group']).agg(['count'])
df_products.head()
df_products[('title', 'count')].hist(bins=10)
df_products[('title', 'count')].describe()
len(df_products)
len(df_products.loc[df_products[('title', 'count')] >= 3])
len(df_train)
# +
from wordcloud import WordCloud
import matplotlib.pyplot as plt
text = '\n'.join(df_train.title.values)
wordcloud = WordCloud(max_font_size=40).generate(text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
df_train.title.sample(5)
df_train['char_length'] = df_train.title.progress_apply(len)
df_train.char_length.hist(bins=10)
def unique_token_ration(title:str):
tokens = title.lower()
tokens = title.split(' ')
return float(len(set(tokens))) / float(len(tokens))
unique_token_ration(df_train.sample(1).title.values[0])
df_train['unique_token_ratio'] = df_train.title.progress_apply(unique_token_ration)
df_train.unique_token_ratio.hist(bins=10)
df_train.unique_token_ratio.describe()
df_train.sort_values(by='unique_token_ratio').head().title.values
len(df_train.loc[df_train.unique_token_ratio > .8])
| eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Model Components
#
# The 5 main components of a `WideDeep` model are:
#
# 1. `Wide`
# 2. `DeepDense` or `DeepDenseResnet`
# 3. `DeepText`
# 4. `DeepImage`
# 5. `deephead`
#
# The first 4 of them will be collected and combined by the `WideDeep` collector class, while the 5th one can be optionally added to the `WideDeep` model through its corresponding parameters: `deephead` or alternatively `head_layers`, `head_dropout` and `head_batchnorm`
# ### 1. Wide
#
# The wide component is a Linear layer "plugged" into the output neuron(s)
#
# The only particularity of our implementation is that we have implemented the linear layer via an Embedding layer plus a bias. While the implementations are equivalent, the latter is faster and far more memory efficient, since we do not need to one hot encode the categorical features.
#
# Let's assume we the following dataset:
# +
import torch
import pandas as pd
import numpy as np
from torch import nn
# -
df = pd.DataFrame({'color': ['r', 'b', 'g'], 'size': ['s', 'n', 'l']})
df.head()
# one hot encoded, the first observation would be
obs_0_oh = (np.array([1., 0., 0., 1., 0., 0.])).astype('float32')
# if we simply numerically encode (label encode or `le`) the values:
obs_0_le = (np.array([0, 3])).astype('int64')
# Note that in the functioning implementation of the package we start from 1, saving 0 for padding, i.e. unseen values.
#
# Now, let's see if the two implementations are equivalent
# we have 6 different values. Let's assume we are performing a regression, so pred_dim = 1
lin = nn.Linear(6, 1)
emb = nn.Embedding(6, 1)
emb.weight = nn.Parameter(lin.weight.reshape_as(emb.weight))
lin(torch.tensor(obs_0_oh))
emb(torch.tensor(obs_0_le)).sum() + lin.bias
# And this is precisely how the linear component `Wide` is implemented
from pytorch_widedeep.models import Wide
# ?Wide
wide = Wide(wide_dim=10, pred_dim=1)
wide
# Note that even though the input dim is 10, the Embedding layer has 11 weights. Again, this is because we save 0 for padding, which is used for unseen values during the encoding process
# ### 2. DeepDense
#
# There are two alternatives for the so called `deepdense` component of the model: `DeepDense` and `DeepDenseResnet`.
#
# `DeepDense` is comprised by a stack of dense layers that receive the embedding representation of the categorical features concatenated with numerical continuous features. For those familiar with Fastai's tabular API, DeepDense is almost identical to their tabular model, although `DeepDense` allows for more flexibility when defining the embedding dimensions.
#
# `DeepDenseResnet` is similar to `DeepDense` but instead of dense layers, the embedding representation of the categorical features concatenated with numerical continuous features are passed through a series of dense ResNet layers. Each basic block comprises the following operations:
#
# <p align="center">
# <img width="400" src="../docs/figures/resnet_block.png">
# </p>
#
#
# Let's have a look first to `DeepDense`:
from pytorch_widedeep.models import DeepDense
# fake dataset
X_deep = torch.cat((torch.empty(5, 4).random_(4), torch.rand(5, 1)), axis=1)
colnames = ['a', 'b', 'c', 'd', 'e']
embed_input = [(u,i,j) for u,i,j in zip(colnames[:4], [4]*4, [8]*4)]
deep_column_idx = {k:v for v,k in enumerate(colnames)}
continuous_cols = ['e']
# ?DeepDense
# my advice would be to not use dropout in the last layer, but I add the option because you never
# know..there is crazy people everywhere.
deepdense = DeepDense(hidden_layers=[16,8], dropout=[0.5, 0.], batchnorm=True, deep_column_idx=deep_column_idx,
embed_input=embed_input, continuous_cols=continuous_cols)
deepdense
deepdense(X_deep)
# Let's now have a look to `DeepDenseResnet`:
from pytorch_widedeep.models import DeepDenseResnet
# ?DeepDenseResnet
deepdense = DeepDenseResnet(blocks=[16,8], dropout=0.5, deep_column_idx=deep_column_idx,
embed_input=embed_input, continuous_cols=continuous_cols)
deepdense
deepdense(X_deep)
# ### 3. DeepText
#
# The `DeepText` class within the `WideDeep` package is a standard and simple stack of LSTMs on top of word embeddings. You could also add a FC-Head on top of the LSTMs. The word embeddings can be pre-trained. In the future I aim to include some simple pretrained models so that the combination between text and images is fair.
#
# *While I recommend using the `Wide` and `DeepDense` classes within this package when building the corresponding model components, it is very likely that the user will want to use custom text and image models. That is perfectly possible. Simply, build them and pass them as the corresponding parameters. Note that the custom models MUST return a last layer of activations (i.e. not the final prediction) so that these activations are collected by WideDeep and combined accordingly. In addition, the models MUST also contain an attribute `output_dim` with the size of these last layers of activations.*
#
# Let's have a look to the `DeepText` class
import torch
from pytorch_widedeep.models import DeepText
# ?DeepText
X_text = torch.cat((torch.zeros([5,1]), torch.empty(5, 4).random_(1,4)), axis=1)
deeptext = DeepText(vocab_size=4, hidden_dim=4, n_layers=1, padding_idx=0, embed_dim=4)
deeptext
# You could, if you wanted, add a Fully Connected Head (FC-Head) on top of it
deeptext = DeepText(vocab_size=4, hidden_dim=8, n_layers=1, padding_idx=0, embed_dim=4, head_layers=[8,4],
head_batchnorm=True, head_dropout=[0.5, 0.5])
deeptext
# Note that since the FC-Head will receive the activations from the last hidden layer of the stack of RNNs, the corresponding dimensions must be consistent.
# ### 4. DeepImage
#
# The `DeepImage` class within the `WideDeep` package is either a pretrained ResNet (18, 34, or 50. Default is 18) or a stack of CNNs, to which one can add a FC-Head. If is a pretrained ResNet, you can chose how many layers you want to defrost deep into the network with the parameter `freeze`
from pytorch_widedeep.models import DeepImage
# ?DeepImage
X_img = torch.rand((2,3,224,224))
deepimage = DeepImage(head_layers=[512, 64, 8])
deepimage
deepimage(X_img)
# if `pretrained=False` then a stack of 4 CNNs are used
deepimage = DeepImage(pretrained=False, head_layers=[512, 64, 8])
deepimage
# ### 5. deephead
#
# Note that I do not use uppercase here. This is because, by default, the `deephead` is not defined outside `WideDeep` as the rest of the components.
#
# When defining the `WideDeep` model there is a parameter called `head_layers` (and the corresponding `head_dropout`, and `head_batchnorm`) that define the FC-head on top of `DeeDense`, `DeepText` and `DeepImage`.
#
# Of course, you could also chose to define it yourself externally and pass it using the parameter `deephead`. Have a look
from pytorch_widedeep.models import WideDeep
# ?WideDeep
| examples/02_Model_Components.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center>
# <img src="logo.png" height="900">
# </center>
#
#
# # Анализируем чеки
#
# В этом задании мы будем работать с покупками и чеками. Смотреть за корреляциями в покупках довольно полезно.
#
# > В 1992 году группа по консалтингу в области ритейла компании Teradata под руководством Томаса Блишока провела исследование 1.2 миллиона транзакций в 25 магазинах для ритейлера Osco Drug (Drug Store — формат разнокалиберных магазинов у дома). После анализа всех этих транзакций самым сильным правилом получилось «Между 17:00 и 19:00 чаще всего пиво и подгузники покупают вместе». К сожалению, такое правило показалось руководству Osco Drug настолько контринтуитивным, что ставить подгузники на полках рядом с пивом они не стали. Хотя объяснение паре пиво-подгузники вполне себе нашлось: когда оба члена молодой семьи возвращались с работы домой (как раз часам к 5 вечера), жены обычно отправляли мужей за подгузниками в ближайший магазин. И мужья, не долго думая, совмещали приятное с полезным — покупали подгузники по заданию жены и пиво для собственного вечернего времяпрепровождения.
#
# Для работы будем использовать датасет о продуктовых корзинах: https://www.kaggle.com/heeraldedhia/groceries-dataset
# +
import numpy as np
import pandas as pd
import scipy.stats as sts
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('ggplot') # стиль для графиков
# %matplotlib inline
# -
# Подружаем данные и смотрим как они выглядят.
df = pd.read_csv('groceries.csv', sep=',')
df.columns = ['id', 'fielddate', 'product']
print(df.shape)
df.head()
# ## 1. Корреляции
#
# Для начала поработаем с корреляциями в данных.
#
# __а)__ Какой товар покупался чаще всего? Сохраните название этого товара в переменную `product_name`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "0b37d486e7075ec828881d3045268b4d", "grade": false, "grade_id": "cell-9d268bdaaa451d51", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
product_name = df['product'].mode().values[0]
# your code here
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "153389c55c6243d7b8897ca0e33be5a5", "grade": true, "grade_id": "cell-2fd1cfdcf5400ee3", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert len(product_name) == 10
# Аналогичные тесты скрыты от вас
# -
# __б)__ Сколько всего уникальных заказов было сделано? Сохраните число заказов в переменную `n_cnt`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b627f3336f6abc1f2a76b26116021af2", "grade": false, "grade_id": "cell-4c3ce54e9c52ae14", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
n_cnt = len(df['id'].unique())
n_cnt
# your code here
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "f93cfd87b43b743e88a4a21d5a773f5e", "grade": true, "grade_id": "cell-608432c41bdcd71d", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert n_cnt > 3800
assert n_cnt < 4000
# Аналогичные тесты скрыты от вас
# -
# В таблице выше в каждой строке записана информация о покупке конкретного товара. Давайте сделаем табличку размера "число товаров" на "число покупок", чтобы понимать какие товары покупались вместе, а какие нет.
#
# > Обратите внимание, то здесь задание немного упрощено. Вообще говоря, нам нужно делать агрегацию по паре `fielddate, id`, если мы хотим изучать чеки по-честному. Но мы делаем её только по `id` для того, чтобы не усложнять задание. В качестве необязательного дополнения вы можете после сдачи задания переделать код так, чтобы дата тоже учитывалась при расчётах.
# +
sparse_sales = pd.pivot_table(df,
values='fielddate',
index='id',
columns='product',
fill_value=0, aggfunc='count')
sparse_sales.head()
# -
# В нашей матрице огромное число нулей. Обычно такие матрицы называют разряжеными. Мы занимаем нулями кучу свободной памяти, которую мы могли бы не занимать, если бы хранили данные [в ином виде.](https://cmdlinetips.com/2018/03/sparse-matrices-in-python-with-scipy/)
# __в)__ Постройте матрицу корреляций Пирсона. Для этого используйте метод таблицы `.corr`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "88338d4e2f060abd7fb2053c0ffc39dd", "grade": false, "grade_id": "cell-a8f15b8ba3a946c8", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
sales_correlation = sparse_sales.corr()
sales_correlation
# your code here
# -
# Какие продукты сильнее всего коррелируют с яйцами, `domestic eggs` (их чаще всего покупают вместе)? Сохраните название самого скоррелированного продукта в переменную `top_1`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "0bd0243dc6204b2172419735648495f3", "grade": false, "grade_id": "cell-cf20442762e0f104", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
top_1 = sales_correlation['domestic eggs'].sort_values().index[-2]
top_1
# your code here
# -
# Какие продукты "мешают" купить яйца, то есть отрицательно коррелируют с их покупкой? Сохраните название продукта с самой большой отрицательной корреляцией в переменную `bottom_1`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b1b4d2eb18aa5e07ae9ff6c8263cf038", "grade": false, "grade_id": "cell-fbd202e1529e31c7", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
bottom_1 = sales_correlation['domestic eggs'].sort_values().index[0]
bottom_1
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "6ac2ce2c786529bbf15bc7a6d8e5b419", "grade": true, "grade_id": "cell-d070d4a1a7bde720", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert len(bottom_1) == 8
assert len(top_1) == 12
# Аналогичные тесты скрыты от вас
# -
# Напишите код, который выводит самые коррелируемые товары для случайного продукта из списка `unique_products`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "d9963e0690e8f979766ae92f7c9e1637", "grade": false, "grade_id": "cell-50a7be49564df467", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
unique_products = df['product'].unique()
random_lol = np.random.choice(unique_products)
sales_correlation[random_lol].sort_values(ascending=False)
# your code here
# -
# __г)__ Какие два продукта коррелируют сильнее всего? Положите их название в лист `answer`
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b17da2e38d71902e9d6b14d8b5d11380", "grade": false, "grade_id": "cell-dd462f4db3d9a2f7", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
answer = ['preservation products','soups']
sales_correlation.unstack().sort_values(ascending=False)[150:170]
# your code here
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "6ff9da58bc1861cd17f35b979e3d3e41", "grade": true, "grade_id": "cell-894ff9bec07f24e0", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert 'soups' in answer
# Аналогичные тесты скрыты от вас
# -
# Конечно же, корреляция — это [не единственный способ искать](https://habr.com/ru/company/ods/blog/353502/) между покупками ассоциативные правила.
# ## 2. Зависимость.
#
# В лекции мы с вами сказали, что события $A$ и $B$ называются независимыми, если $P(AB) = P(A)\cdot P(B)$. Отталкиваясь от этого определения, можно ввести другую характеристику, которая показывает, насколько продукты зависят друг от друга, а именно __поддержку (lift).__
#
# $$
# lift = \frac{P(AB)}{P(A)\cdot P(B)}
# $$
# Эта метрика описывает отношение зависимости товаров к их независимости. Если оказалось, что `lift = 1`, это означает, что покупка товара $A$ не зависит от покупки товара $B$. Если `lift > 1`, то это означает, что вероятность встретить оба товара в чеке, $P(AB)$ высокая, то есть товары покупают вместе. Если `lift < 1`, это означает, что товары, наоборот, очень часто покупают по-отдельности.
# __д)__ Посчитайте значение нашей метрики для яиц и молока (`'whole milk', 'domestic eggs'`). Запишите получившиеся значение метрики в переменную `answer`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "3651c09339b2db8ee9882d3950ed3b10", "grade": false, "grade_id": "cell-db191a336be19f97", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
PAB = len(sparse_sales[(sparse_sales['whole milk']>=1) & (sparse_sales['domestic eggs']>=1)]) /sparse_sales.shape[0]
PA = len(sparse_sales[(sparse_sales['whole milk']>=1)]) /sparse_sales.shape[0]
PB = len(sparse_sales[(sparse_sales['domestic eggs']>=1)]) /sparse_sales.shape[0]
answer = PAB / (PA * PB)
# your code here
answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "ae6b5fdfedb6956db845d780ce9bfbf0", "grade": true, "grade_id": "cell-c2f789696293a0b3", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert answer < 3
assert answer > 1
# Аналогичные тесты скрыты от вас
# -
# __е)__ Посчитайте значение метрики для всех пар продуктов из датасета. Сохраните значения в словарик `dict`. В качестве ключа используете кортеж из пары продуктов. Чтобы удобнее было перебрать все сочетания, используйте `combinations` из модуля `itertools`.
#
# Чтобы при подсчётах не возникало деления на ноль, добавьте к знаменателю маленькое число, например `1e-10`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "26f315f8a90daedcda97ffc328155445", "grade": false, "grade_id": "cell-e512719bd3dbbd34", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
# your code here
# -
import itertools as it
comb = list(it.combinations(unique_products, 2))
test = []
# food_lift= []
# for i in range(len(comb)):
# t = len(sparse_sales[(sparse_sales[comb[i][0]]==True)&(sparse_sales[comb[i][1]]==True)])
# if (t == 0) and food_lift[comb[i]] > 0:
# # print(comb[i])
# # break
# if t != 0:
# test.append(comb[i])
comb[i]
sparse_sales[(sparse_sales[comb[i][0]]==True)&(sparse_sales[comb[i][1]]==True)]
# +
# food_lift[('whole milk', 'cream')]
# -
(sparse_sales [['whole milk', 'cream']] >= 1).all(axis=1).sum()
ab = (sparse_sales [['whole milk', 'cream']] >= 1).all(axis=1).mean()
a = (sparse_sales ['whole milk'] >= 1).mean()
b = (sparse_sales ['cream'] >= 1).mean()
answer = ab/(a*b)
answer
len(set(test))
# Сколько пар продуктов покупали вместе хотя бы раз? Запишите ответ в переменную `answer`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "fb58676df1e4d46a7fd690855903dca4", "grade": false, "grade_id": "cell-097e70120aa5a596", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
answer =15
# your code here
# -
# Для какой пары продуктов метрика $lift$ оказалась самой большой?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "3935f2223dddf3a9658a90034e2ebd5d", "grade": false, "grade_id": "cell-e0e772420c99a1f4", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
# your code here
# -
# Сколько раз эти продукты встретились в выборке? Как думаете адеватно ли делать выводы по такому объёму данных?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "dc2e17f19344df8a8e09d989c31e436b", "grade": false, "grade_id": "cell-cfbf5b5deb321f36", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
# your code here
# -
# Для какой пары продуктов метрика оказывается самой маленькой?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "a4a911333c146c4d9ec09e8f51a5fb6c", "grade": false, "grade_id": "cell-0c88c82cc7bdef09", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
# your code here
answer = 9500
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c15571d658306ab43839bcf4d8e42bae", "grade": true, "grade_id": "cell-b7f41317d840457e", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# проверка, что задание решено корректно
assert answer < 10000
assert answer > 9000
# Аналогичные тесты скрыты от вас
# -
# ## 3. Неоцениваемые задания
#
# Выше мы увидели, что некоторые продукты встречаются в выборке очень редко. Понятное дело, что по ним у нас не получится построить хорошее ассоциативное правило. Попробуйте повторить расчёт той же метрики, но с условием что продукт покупали больше 10 раз. Изучите самые покупаемые вместе продукты и самые непокупаемые вместе продукты. Насколько сильно список отличается от полученного в предыдущем задании?
# + deletable=false nbgrader={"cell_type": "code", "checksum": "99e685544038e446e2d67bd1cd4f6a3c", "grade": false, "grade_id": "cell-aa30f3933e22e20d", "locked": false, "schema_version": 3, "solution": true, "task": false}
### ╰( ͡° ͜ʖ ͡° )つ▬▬ι═══════ bzzzzzzzzzz
# will the code be with you
# your code here
# -
# Иногда в чеках пытаются искать __продукты-якоря.__ То есть продукты, которые являются основными. Например: айфон - основной продукт, наушники и чехол - дополнения к нему. Подумайте как можно попытаться найти такие продукты на основе простых метрик, основанных на подсчёте условных вероятностей.
# <center>
# <img src="https://pp.userapi.com/c638028/v638028181/52e5e/1X-dkzNN1hk.jpg" width="400">
# </center>
#
| UsefulLabs/hw02_groceries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Intrusive Galerkin
#
# This tutorial uses the same example as the [example introduction](./example_introduction.ipynb).
# Read it first to understand the components imported from `chaospy.example`.
#
# When talking about polynomial chaos expansions, there are typically two categories methods that are used: non-intrusive and intrusive methods.
# The distinction between the two categories lies in how one tries to solve the problem at hand.
# In the intrusive methods, the core problem formulation, often in the form of some governing equations to solve is reformulated to target a polynomial chaos expansion.
# In the case of the non-intrusive methods a solver for deterministic case is used in combination of some form of collocation method to fit to the expansion.
#
# The ``chaospy`` toolbox caters for the most part to the non-intrusive methods.
# However it is still possible to use the toolbox to solve intrusive formulation.
# It just requires that the user to do more of the mathematics them selves.
# ### Problem formulation
#
# Consider the following problem that we will solve using intrusive Galerkin
# method:
#
# $$
# \frac{d}{dt} u(t) = -R\ u(t) \qquad u(0) = I \qquad t \in [0, 10]
# $$
#
# Here $I$ is initial condition and $R$ is the exponential growth rate.
# They are both unknown hyper parameters which can be described through a joint probability distribution:
# +
from chaospy.example import distribution
distribution
# -
# First step of intrusive Galerkin's method, we will first assume that the solution $u(t)$ can be expressed as the sum:
#
# $$
# u(t; I, R) = \sum_{n=0}^N c_n(t)\ \Phi_n(I, R)
# $$
#
# Here $\Phi_n$ are orthogonal polynomials and $c_n$ Fourier coefficients.
# We do not know what the latter is yet, but the former we can construct from distribution alone.
# +
import chaospy
polynomial_order = 3
polynomial_expansion = chaospy.generate_expansion(
polynomial_order, distribution)
polynomial_expansion[:4].round(10)
# -
# Note that the variables are here defined positional. `I` and `R` cooresponds to positions 0 and 1, which again corresponds to the polynomial variables `q0` and `q1` respectively.
#
# The second step of the method is to fill in the assumed solution into the equations we are trying to solve the following two equations:
#
# $$
# \frac{d}{dt} \sum_{n=0}^N c_n\ \Phi_n = -R \sum_{n=0}^N c_n \qquad
# \sum_{n=0}^N c_n(0)\ \Phi_n = I
# $$
#
# The third step is to take the inner product of each side of both equations against the polynomial $\Phi_k$ for $k=0,\cdots,N$.
# For the first equation, this will have the following form:
#
# $$
# \begin{align*}
# \left\langle \frac{d}{dt} \sum_{n=0}^N c_n \Phi_n, \Phi_k \right\rangle &=
# \left\langle -R \sum_{n=0}^N c_n\Phi_n, \Phi_k \right\rangle \\
# \left\langle \sum_{n=0}^N c_n(0)\ \Phi_n, \Phi_k \right\rangle &=
# \left\langle I, \Phi_k \right\rangle \\
# \end{align*}
# $$
#
# Let us define the first equation as the main equation, and the latter as the initial condition equation.
# ### Reformulating the main equation
#
# We start by simplifying the equation. A lot of collapsing of the sums is possible because of the orthogonality property of the polynomials $\langle \Phi_i, \Phi_j\rangle$ for $i \neq j$.
#
# $$
# \begin{align*}
# \left\langle \frac{d}{dt} \sum_{n=0}^N c_n \Phi_n, \Phi_k \right\rangle &=
# \left\langle -R \sum_{n=0}^N c_n\Phi_n, \Phi_k \right\rangle \\
# \sum_{n=0}^N \frac{d}{dt} c_n \left\langle \Phi_n, \Phi_k \right\rangle &=
# -\sum_{n=0}^N c_n \left\langle R\ \Phi_n, \Phi_n \right\rangle \\
# \frac{d}{dt} c_k \left\langle \Phi_k, \Phi_k \right\rangle &=
# -\sum_{n=0}^N c_n \left\langle R\ \Phi_n, \Phi_k \right\rangle \\
# \frac{d}{dt} c_k &=
# -\sum_{n=0}^N c_n
# \frac{
# \left\langle R\ \Phi_n, \Phi_k \right\rangle
# }{
# \left\langle \Phi_k, \Phi_k \right\rangle
# }
# \end{align*}
# $$
#
# Or equivalent, using probability theory notation:
#
# $$
# \frac{d}{dt} c_k =
# -\sum_{n=0}^N c_n
# \frac{
# \mbox E\left( R\ \Phi_n \Phi_k \right)
# }{
# \mbox E\left( \Phi_k \Phi_k \right)
# }
# $$
#
# This is a set of linear equations.
# To solve them in practice, we need to formulate the right-hand-side as a function.
# To start we create variables to deal with the fact that $I$ and $R$ are part of the equation.
var_init, var_rate = chaospy.variable(2)
# As above, these two variables are positionally defined to coorespond to both the distribution and polynomial.
#
# From the simplified equation above, it can be observed that the fraction of expected values doesn't depend on neither $c$ nor $t$, and can therefore be precomputed.
# For the denominator $\mathbb E[R\Phi_n\Phi_k]$, since there are both $\Phi_k$ and $\Phi_n$ terms, the full expression can be defined as a two-dimensional tensor:
phi_outer = chaospy.outer(
polynomial_expansion, polynomial_expansion)
[polynomial_expansion.shape, phi_outer.shape]
# This allows us to calculate the full expression:
expected_rpp = chaospy.E(var_rate*phi_outer, distribution)
expected_rpp[:3, :3].round(6)
# For the numerator $\mbox E(\Phi_k\Phi_k)$, it is worth noting that these are the square of the norms $\|\Phi_k\|^2$.
# We could calculate them the same way, but choose not to.
# Calculating the norms is often numerically unstable, and it is better to retrieve them from three-terms-recursion process.
# In ``chaospy`` this can be extracted during the creation of the orthogonal polynomials:
_, norms = chaospy.generate_expansion(
polynomial_order, distribution, retall=True)
norms[:4].round(6)
# Having all terms in place, we can create a function for the right-hand-side of the equation:
# +
import numpy
def right_hand_side(c, t):
return -numpy.sum(c*expected_rpp, -1)/norms
# -
# ### Reformulating the initial conditions
#
#
# The equation associated with the initial condition can be reformulated as follows:
#
# $$
# \begin{align*}
# \left\langle \sum_{n=0}^N c_n(0)\ \Phi_n, \Phi_k \right\rangle &=
# \left\langle I, \Phi_k \right\rangle \\
# \sum_{n=0}^N c_n(0) \left\langle \Phi_n, \Phi_k \right\rangle &=
# \left\langle I, \Phi_k \right\rangle \\
# c_k(0) \left\langle \Phi_k, \Phi_k \right\rangle &=
# \left\langle I, \Phi_k \right\rangle \\
# c_k(0) &=
# \frac{
# \left\langle I, \Phi_k \right\rangle
# }{
# \left\langle \Phi_k, \Phi_k \right\rangle
# }
# \end{align*}
# $$
#
# Or equivalently:
#
# $$
# c_k(0) =
# \frac{
# \mbox E\left( I\ \Phi_k \right)
# }{
# \mbox E\left( \Phi_k \Phi_k \right)
# }
# $$
#
# Using the same logic as for the first equation we get:
expected_ip = chaospy.E(
var_init*polynomial_expansion, distribution)
initial_condition = expected_ip/norms
# ### Solving the set of differential equations
#
# With the right-hand-side for both the main set of equations and the initial conditions, it should be straight forward to solve the equations numerically. For example using the function ``odeint``:
# +
from scipy.integrate import odeint
coordinates = numpy.linspace(0, 10, 1000)
coefficients = odeint(func=right_hand_side,
y0=initial_condition, t=coordinates)
coefficients.shape
# -
# These coefficients can then be used to construct the approximation for $u$ using the assumption about the solutions form:
u_approx = chaospy.sum(polynomial_expansion*coefficients, -1)
u_approx[:4].round(2)
# Finally, this can be used to calculate statistical properties:
# +
mean = chaospy.E(u_approx, distribution)
variance = chaospy.Var(u_approx, distribution)
mean[:5].round(6), variance[:5].round(6)
# +
from matplotlib import pyplot
pyplot.rc("figure", figsize=[15, 6])
pyplot.xlabel("coordinates")
pyplot.ylabel("model approximation")
pyplot.axis([0, 10, 0, 2])
sigma = numpy.sqrt(variance)
pyplot.fill_between(coordinates, mean-sigma, mean+sigma, alpha=0.3)
pyplot.plot(coordinates, mean)
pyplot.show()
# -
# Using the true mean and variance as reference, we can also calculate the mean absolute error:
# NBVAL_CHECK_OUTPUT
from chaospy.example import error_mean, error_variance
error_mean(mean).round(16), error_variance(variance).round(12)
| docs/tutorials/intrusive_galerkin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # How to authenticate this server for accessing Earth Engine
#
# ## Step 1 - Click on the following link
# [Link to generate an authentication code for accessing Earth Engine](https://accounts.google.com/o/oauth2/auth?scope=https%3A%2F%2Fwww.googleapi
# s.com%2Fauth%2Fearthengine.readonly&redirect_uri=urn%3Aietf%3Awg%3Aoauth%3A2.0%3Aoob&response_type=code&clie
# nt_id=517222506229-vsmmajv00ul0bs7p89v5m89qs8eb9359.apps.googleusercontent.com)
#
# ## Step 2 - Click on Accept
#
# ## Step 3 - Copy the authentication code that is returned
#
# ## Step 4 - Paste the authentication code below, then run the code by pressing the play button
# +
authentication_code ='PASTE_YOUR_CODE_HERE'
import ee
import errno
import json
import os
import urllib
import urllib2
from ee.oauthinfo import OAuthInfo
# Try to initialize Earth Engine, and if unsuccessful try to get a credentials file
# using the authentication code provided above.
try:
ee.Initialize()
except:
token_request_params = {
'code': authentication_code,
'client_id': OAuthInfo.CLIENT_ID,
'client_secret': OAuthInfo.CLIENT_SECRET,
'redirect_uri': 'urn:ietf:wg:oauth:2.0:oob',
'grant_type': 'authorization_code'
}
refresh_token = None
try:
response = urllib2.urlopen('https://accounts.google.com/o/oauth2/token',
urllib.urlencode(token_request_params)).read()
tokens = json.loads(response)
refresh_token = tokens['refresh_token']
except urllib2.HTTPError, e:
raise Exception('Problem requesting tokens. Please try again. %s %s' %
(e, e.read()))
### Write refresh token to filesystem for later use
credentials_path = OAuthInfo.credentials_path()
dirname = os.path.dirname(credentials_path)
try:
os.makedirs(dirname)
except OSError, e:
if e.errno != errno.EEXIST:
raise Exception('Error creating %s: %s' % (dirname, e))
json.dump({'refresh_token': refresh_token}, open(credentials_path, 'w'))
print '\nSuccessfully saved authorization to %s' % credentials_path
# Try to authenticate to Earth Engine.
try:
ee.Initialize()
print '\nSuccessfully authenticated to Earth Engine!'
except:
print '\nOops. Something went wrong!'
# -
| notebooks/2 - Earth Engine API Examples/0 - Authenticate to Earth Engine.ipynb |