code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # The MPC out of Credit vs the MPC Out of Income
#
# This notebook compares the Marginal Propensity to Consume (MPC) out of an increase in a credit limit, and the MPC out of transitory shock to income.
#
# The notebook is heavily commented to help newcomers, and does some things (like importing modules in the body of the code rather than at the top), that are typically deprecated by Python programmers. This is all to make the code easier to read and understand.
#
# The notebook illustrates one simple way to use HARK: import and solve a model for different parameter values, to see how parameters affect the solution.
#
# The first step is to create the ConsumerType we want to solve the model for.
# %%
from __future__ import division, print_function
# %matplotlib inline
# %% code_folding=[]
## Import the HARK ConsumerType we want
## Here, we bring in an agent making a consumption/savings decision every period, subject
## to transitory and permanent income shocks.
from HARK.ConsumptionSaving.ConsIndShockModel import IndShockConsumerType
# %% code_folding=[]
## Import the default parameter values
import HARK.ConsumptionSaving.ConsumerParameters as Params
# %% code_folding=[]
## Now, create an instance of the consumer type using the default parameter values
## We create the instance of the consumer type by calling IndShockConsumerType()
## We use the default parameter values by passing **Params.init_idiosyncratic_shocks as an argument
BaselineExample = IndShockConsumerType(**Params.init_idiosyncratic_shocks)
# %% code_folding=[]
# Note: we've created an instance of a very standard consumer type, and many assumptions go
# into making this kind of consumer. As with any structural model, these assumptions matter.
# For example, this consumer pays the same interest rate on
# debt as she earns on savings. If instead we wanted to solve the problem of a consumer
# who pays a higher interest rate on debt than she earns on savings, this would be really easy,
# since this is a model that is also solved in HARK. All we would have to do is import that model
# and instantiate an instance of that ConsumerType instead. As a homework assignment, we leave it
# to you to uncomment the two lines of code below, and see how the results change!
from HARK.ConsumptionSaving.ConsIndShockModel import KinkedRconsumerType
BaselineExample = KinkedRconsumerType(**Params.init_kinked_R)
# %% [markdown]
r"""
The next step is to change the values of parameters as we want.
To see all the parameters used in the model, along with their default values, see $\texttt{ConsumerParameters.py}$
Parameter values are stored as attributes of the $\texttt{ConsumerType}$ the values are used for. For example, the risk-free interest rate $\texttt{Rfree}$ is stored as $\texttt{BaselineExample.Rfree}$. Because we created $\texttt{BaselineExample}$ using the default parameters values at the moment $\texttt{BaselineExample.Rfree}$ is set to the default value of $\texttt{Rfree}$ (which, at the time this demo was written, was 1.03). Therefore, to change the risk-free interest rate used in $\texttt{BaselineExample}$ to (say) 1.02, all we need to do is:
"""
# %% code_folding=[0]
# Change the Default Riskfree Interest Rate
BaselineExample.Rfree = 1.02
# %% code_folding=[0]
## Change some parameter values
BaselineExample.Rfree = 1.02 #change the risk-free interest rate
BaselineExample.CRRA = 2. # change the coefficient of relative risk aversion
BaselineExample.BoroCnstArt = -.3 # change the artificial borrowing constraint
BaselineExample.DiscFac = .5 #chosen so that target debt-to-permanent-income_ratio is about .1
# i.e. BaselineExample.solution[0].cFunc(.9) ROUGHLY = 1.
## There is one more parameter value we need to change. This one is more complicated than the rest.
## We could solve the problem for a consumer with an infinite horizon of periods that (ex-ante)
## are all identical. We could also solve the problem for a consumer with a fininite lifecycle,
## or for a consumer who faces an infinite horizon of periods that cycle (e.g., a ski instructor
## facing an infinite series of winters, with lots of income, and summers, with very little income.)
## The way to differentiate is through the "cycles" attribute, which indicates how often the
## sequence of periods needs to be solved. The default value is 1, for a consumer with a finite
## lifecycle that is only experienced 1 time. A consumer who lived that life twice in a row, and
## then died, would have cycles = 2. But neither is what we want. Here, we need to set cycles = 0,
## to tell HARK that we are solving the model for an infinite horizon consumer.
## Note that another complication with the cycles attribute is that it does not come from
## Params.init_idiosyncratic_shocks. Instead it is a keyword argument to the __init__() method of
## IndShockConsumerType.
BaselineExample.cycles = 0
# %%
# Now, create another consumer to compare the BaselineExample to.
# %% code_folding=[0]
# The easiest way to begin creating the comparison example is to just copy the baseline example.
# We can change the parameters we want to change later.
from copy import deepcopy
XtraCreditExample = deepcopy(BaselineExample)
# Now, change whatever parameters we want.
# Here, we want to see what happens if we give the consumer access to more credit.
# Remember, parameters are stored as attributes of the consumer they are used for.
# So, to give the consumer more credit, we just need to relax their borrowing constraint a tiny bit.
# Declare how much we want to increase credit by
credit_change = .0001
# Now increase the consumer's credit limit.
# We do this by decreasing the artificial borrowing constraint.
XtraCreditExample.BoroCnstArt = BaselineExample.BoroCnstArt - credit_change
# %% [markdown]
"""
Now we are ready to solve the consumers' problems.
In HARK, this is done by calling the solve() method of the ConsumerType.
"""
# %% code_folding=[0]
### First solve the baseline example.
BaselineExample.solve()
### Now solve the comparison example of the consumer with a bit more credit
XtraCreditExample.solve()
# %% [markdown]
r"""
Now that we have the solutions to the 2 different problems, we can compare them.
We are going to compare the consumption functions for the two different consumers.
Policy functions (including consumption functions) in HARK are stored as attributes
of the *solution* of the ConsumerType. The solution, in turn, is a list, indexed by the time
period the solution is for. Since in this demo we are working with infinite-horizon models
where every period is the same, there is only one time period and hence only one solution.
e.g. BaselineExample.solution[0] is the solution for the BaselineExample. If BaselineExample
had 10 time periods, we could access the 5th with BaselineExample.solution[4] (remember, Python
counts from 0!) Therefore, the consumption function cFunc from the solution to the
BaselineExample is $\texttt{BaselineExample.solution[0].cFunc}$
"""
# %% code_folding=[0]
## First, declare useful functions to plot later
def FirstDiffMPC_Income(x):
# Approximate the MPC out of income by giving the agent a tiny bit more income,
# and plotting the proportion of the change that is reflected in increased consumption
# First, declare how much we want to increase income by
# Change income by the same amount we change credit, so that the two MPC
# approximations are comparable
income_change = credit_change
# Now, calculate the approximate MPC out of income
return (BaselineExample.solution[0].cFunc(x + income_change) -
BaselineExample.solution[0].cFunc(x)) / income_change
def FirstDiffMPC_Credit(x):
# Approximate the MPC out of credit by plotting how much more of the increased credit the agent
# with higher credit spends
return (XtraCreditExample.solution[0].cFunc(x) -
BaselineExample.solution[0].cFunc(x)) / credit_change
# %% code_folding=[0]
## Now, plot the functions we want
# Import a useful plotting function from HARK.utilities
from HARK.utilities import plotFuncs
import pylab as plt # We need this module to change the y-axis on the graphs
# Declare the upper limit for the graph
x_max = 10.
# Note that plotFuncs takes four arguments: (1) a list of the arguments to plot,
# (2) the lower bound for the plots, (3) the upper bound for the plots, and (4) keywords to pass
# to the legend for the plot.
# Plot the consumption functions to compare them
# The only difference is that the XtraCredit function has a credit limit that is looser
# by a tiny amount
print('The XtraCredit consumption function allows the consumer to spend a tiny bit more')
print('The difference is so small that the baseline is obscured by the XtraCredit solution')
plotFuncs([BaselineExample.solution[0].cFunc,XtraCreditExample.solution[0].cFunc],
BaselineExample.solution[0].mNrmMin,x_max,
legend_kwds = {'loc': 'upper left', 'labels': ["Baseline","XtraCredit"]})
# Plot the MPCs to compare them
print('MPC out of Credit v MPC out of Income')
plt.ylim([0.,1.2])
plotFuncs([FirstDiffMPC_Credit,FirstDiffMPC_Income],
BaselineExample.solution[0].mNrmMin,x_max,
legend_kwds = {'labels': ["MPC out of Credit","MPC out of Income"]})
| notebooks/MPC-Out-of-Credit-vs-MPC-Out-of-Income.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import platform
print (platform.python_version())
"""
Straightforward table converter to convert Excel tables into ontology files.
See the inline documentation in the notebook.
7-19-18:
1. Start with Chris' i2b2 Hierarchy View
2. Last column can optionally be comments
3. File is "*i2b2 Hierarchy View.xslx"
4. By default process all sheets in a file
5. There will be a "ready for i2b2" folder
"""
# +
# Example code to set a keyring password for use below
keyring.set_password(password="**<PASSWORD>**",service_name="db.concern_columbia",username="i2b2u")
# +
# Import and set paths
import glob
import pandas as pd
import numpy as np
import keyring
basepath="/Users/jeffklann/Dropbox (Partners HealthCare)/CONCERN All Team Work/Data Elements/Data Structures/Ready/For i2b2/"
outpath="/Users/jeffklann/Dropbox (Partners HealthCare)/CONCERN All Team Work/Data Elements/Data Structures/Ready/For i2b2/i2b2_output/"
password_columbia = keyring.get_password(service_name='db.concern_columbia',username='i2b2u') # You need to previously have set it with set_password
password = keyring.get_password(service_name='db.concern_phs',username='concern_user') # You need to previously have set it with set_password
# +
# Connect to SQL for persistence
# %load_ext sql
connect = "mssql+pymssql://concern_user:%s@phssql2193.partners.org/CONCERN_DEV?charset=utf8" % password
#connect = "mssql+pymssql://i2b2u:%s@10.171.30.160/CONCERN_DEV?charset=utf8" % password_columbia
# %sql $connect
# %sql USE CONCERN_DEV
import sqlalchemy
engine = sqlalchemy.create_engine(connect)
# -
# (Re)create the target ontology table
sql = """
CREATE TABLE [dbo].[autoprocessed_i2b2ontology] (
[index] int NOT NULL,
[C_HLEVEL] int NOT NULL,
[C_FULLNAME] varchar(4000) NOT NULL,
[C_NAME] varchar(2000) NOT NULL,
[C_SYNONYM_CD] char(1) NOT NULL,
[C_VISUALATTRIBUTES] char(3) NOT NULL,
[C_TOTALNUM] int NULL,
[C_BASECODE] varchar(250) NULL,
[C_METADATAXML] varchar(max) NULL,
[C_FACTTABLECOLUMN] varchar(50) NOT NULL,
[C_TABLENAME] varchar(50) NOT NULL,
[C_COLUMNNAME] varchar(50) NOT NULL,
[C_COLUMNDATATYPE] varchar(50) NOT NULL,
[C_OPERATOR] varchar(10) NOT NULL,
[C_DIMCODE] varchar(700) NOT NULL,
[C_COMMENT] varchar(max) NULL,
[C_TOOLTIP] varchar(900) NULL,
[M_APPLIED_PATH] varchar(700) NOT NULL,
[UPDATE_DATE] datetime NULL,
[DOWNLOAD_DATE] datetime NULL,
[IMPORT_DATE] datetime NULL,
[SOURCESYSTEM_CD] varchar(50) NULL,
[VALUETYPE_CD] varchar(50) NULL,
[M_EXCLUSION_CD] varchar(25) NULL,
[C_PATH] varchar(300) NULL,
[C_SYMBOL] varchar(100) NULL
)
ON [PRIMARY]
TEXTIMAGE_ON [PRIMARY]
WITH (
DATA_COMPRESSION = NONE
)
"""
engine.execute("drop table autoprocessed_i2b2ontology")
engine.execute(sql)
# +
""" Input a df with columns (minimally): Name, Code, [Ancestor_Code]*, [Modifier]
If Modifier column is included, the legal values are "" or "Y"
Will add additional columns: tooltip, h_level, fullname
rootName is prepended to the path for non-modifiers
rootModName is prepended to the path for modifiers
Derived from ontology_gen_flowsheet.py
"""
def OntProcess(rootName, df,rootModName='MOD'):
# Little inner function that renames the fullname (replaces rootName with rootName+'_MOD') if modifier is 'Y'
def fullmod(fullname,modifier):
ret=fullname.replace('\\'+rootName,'\\'+rootModName) if modifier=='Y' else fullname
return ret
ancestors=[1,-6] if 'Modifier' in df.columns else [1,-5]
df['fullname']=''
df['tooltip']=''
df['path']=''
df['h_level']=np.nan
df['has_children']=0
df=doNonrecursive(df,ancestors)
df['fullname']=df['fullname'].map(lambda x: x.lstrip(':\\')).map(lambda x: x.rstrip(':\\'))
df['fullname']='\\'+rootName+'\\'+df['fullname'].map(str)+"\\"
df['h_level']=df['fullname'].str.count('\\\\')-2
if ('Modifier' in df.columns):
# If modifier subtract 1 from hlevel and change the root to root+'_MOD'
df['h_level']=df['h_level']-df['Modifier'].fillna('').str.len()
df['fullname']=df[['fullname','Modifier']].apply(lambda x: fullmod(*x), axis=1) #sjm on stackoverflow
# Parent check! Will report has_children is true if the Code in each row is ever used as a parent code
if (len(df.columns)>7):
mydf = df
mymerge = mydf.merge(mydf,left_on=mydf.columns[1],right_on=mydf.columns[2],how='inner',suffixes=['','_r']).groupby('Code').size().reset_index().rename(lambda x: 'size' if x==0 else x,axis='columns')
mymerge=mydf.merge(mymerge,
left_on='Code',right_on='Code',how='left')
df['has_children'] = (mymerge['size'] > 0)
else:
df['has_children'] = False
#old bad code
#df['has_children'] = df['h_level']-len(df.columns[1:-5])-2 - This old version just checked to see if this element was at the max depth, which tells us nothing!
#df['has_children'] = df['has_children'].replace({-1:'Y',0:'N'})
#df['Code'].join(df.ix(3))
df=df.append({'fullname':'\\'+rootName+'\\','Name':rootName.replace('\\',' '),'Code':'toplevel|'+rootName.replace('\\',' '),'h_level':1,'has_children':True},ignore_index=True) # Add root node
return df
def doNonrecursive(df,ancestors):
cols=df.columns[ancestors[0]:ancestors[1]][::-1] # Go from column 5 before the end (we added a bunch of columns) backward to first column
print(cols)
for col in cols:
# doesn't work - mycol = df[col].to_string(na_rep='')
mycol = df[col].apply(lambda x: x if isinstance(x, str) else "{:.0f}".format(x)).astype('str').replace('nan','')
df.fullname = df.fullname.str.cat(mycol,sep='\\',na_rep='')
return df
""" Input a df with (minimally): Name, Code, [Ancestor_Code]*, [Modifier], fullname, path, h_level
Optionally input an applied path for modifiers (only one is supported per ontology at present)
Outputs an i2b2 ontology compatible df.
"""
def OntBuild(df,appliedName=''):
odf = pd.DataFrame()
odf['c_hlevel']=df['h_level']
odf['c_fullname']=df['fullname']
odf['c_visualattributes']=df['has_children'].apply(lambda x: 'FAE' if x==True else 'LAE')
odf['m_applied_path']='@'
if 'Modifier' in df.columns:
odf['c_visualattributes']=odf['c_visualattributes']+df.Modifier.fillna('')
odf['c_visualattributes'].replace(to_replace={'FAEY':'OAE','LAEY':'RAE'},inplace=True)
odf['m_applied_path']=df.Modifier.apply(lambda x: '\\'+appliedName+'\\%' if x=='Y' else '@')
odf['c_name']=df['Name']
odf['c_path']=df['path']
odf['c_basecode']=df['Code'] # Assume here leafs are unique, not dependent on parent code (unlike flowsheets)
odf['c_symbol']=odf['c_basecode']
odf['c_synonym_cd']='N'
odf['c_facttablecolumn']='concept_cd'
odf['c_tablename']='concept_dimension'
odf['c_columnname']='concept_path'
odf['c_columndatatype']='T' #this is not the tval/nval switch - 2/20/18 - df['vtype'].apply(lambda x: 'T' if x==2 else 'N')
odf['c_totalnum']=''
odf['c_operator']='LIKE'
odf['c_dimcode']=df['fullname']
odf['c_comment']=None
odf['c_tooltip']=df['fullname'] # Tooltip right now is just the fullname again
#odf['c_metadataxml']=df[['vtype','Label']].apply(lambda x: mdx.genXML(mdx.mapper(x[0]),x[1]),axis=1)
return odf
# -
# Loop through each sheet in an Excel file to generate an ontology .csv
# f is the filename (with path and extension)
# returns a list of dataframes in ontology format
def ontExcelConverter(f):
dfs = []
dfd = pd.read_excel(f,sheet_name=None)
for i,s in enumerate(dfd.keys()): # Now take all sheets, not just 'Sheet1'
df=dfd[s].dropna(axis='columns',how='all')
if len(df.columns)>1:
# Prettyprint the root node name from file and sheet name
shortf = f[f.rfind('/')+1:] # Remove path, get file name only
shortf = shortf[:shortf.find("i2b2")].strip(' ') # Stop at 'i2b2', bc files should be named *i2b2 hierarchy view.xlsx
shortf=shortf+('' if s=='Sheet1' else '_'+str(i))
print('---'+shortf)
# Clean up df
df = df.rename(columns={'Code (concept_CD/inpatient note type CD)':'Code'}) # Hack bc one file has wrong col name
df = df.drop(['Definition','definition','Comment','Comments'],axis=1,errors='ignore') # Drop occasional definition and comment columns
print(df.columns)
# Process df and add to superframe (dfs)
df = OntProcess('CONCERN\\'+shortf,df,'CONCERN_MOD\\'+shortf)
ndf = OntBuild(df,'CONCERN\\'+shortf).fillna('None')
dfs.append(ndf)
return dfs
# Main loop to process all files in a directory, export to csv, and upload the concatenated version to a database
dfs = []
for f in glob.iglob(basepath+"*.xlsx"): # the old place, multi-directory - now all in one dir"**/*i2b2 Hierarchy View*.xlsx"):
if ('~$' in f): continue # Work around Mac temp files
df = ontExcelConverter(f)
dfs = dfs + df
#ndf.to_csv(outpath+shortf+s+"_autoprocessed.csv")
outdf = pd.concat(dfs)
outdf = outdf.append({'c_hlevel':0,'c_fullname':'\\CONCERN\\','c_name':'CONCERN Root','c_basecode':'.dummy','c_visualattributes':'CAE','c_synonym_cd':'N','c_facttablecolumn':'concept_cd','c_tablename':'concept_dimension','c_columnname':'concept_path','c_columndatatype':'T','c_operator':'LIKE','c_dimcode':'\\CONCERN\\','m_applied_path':'@'},ignore_index=True)
outdf.to_csv(outpath+"autoprocessed_i2b2ontology.csv")
engine.execute("delete from autoprocessed_i2b2ontology") # if we use SQLMagic in the same cell as SQLAlchemy, it seems to hang
outdf.to_sql('autoprocessed_i2b2ontology',con=engine,if_exists='append')
# Perform check to make sure no codes are used twice. This is not necessarily an error so only output a warning.
dups = outdf.groupby('c_basecode').size()
dups = dups[dups>1]
if len(dups)>0:
print("Warning: codes used multiple times, check to verify this is intentional:")
print(dups)
# + [markdown] slideshow={"slide_type": "-"}
# # End of main code...
# ------------------------
# -
# Special hacked code for the weird ADT table file format
# DEPRECATED
dfs = []
dfd=pd.read_excel(basepath+"ADT/ADTEventHierarchy AND LocationHierarchy for Each site i2b2 June 21 2018_update.xlsx",
sheet_name=None)
for k,v in dfd.items():
shortf=k[0:k.find(' ',k.find(' ')+1)].replace(' ','_')
print(shortf)
df=v.dropna(axis='columns',how='all')
df = df.drop(['C_TOOLTIP','c_tooltip'],axis=1,errors='ignore')
print(df.columns)
df = OntProcess('CONCERN\\'+shortf,df)
ndf = OntBuild(df)
dfs.append(ndf)
ndf.to_csv(outpath+shortf+"_autoprocessed.csv")
#tname = 'out_'+shortf
#globals()[tname]=ndf
# #%sql DROP TABLE $tname
# #%sql PERSIST $tname
# One-off a prespecified file
k = "VISIT i2b2 HIERARCHY.xlsx"
dfs = ontExcelConverter("/Users/jeffklann/Downloads/"+k)
outdf = pd.concat(dfs)
outdf.to_csv("/Users/jeffklann/Downloads/VISIT_HIERARCHY"+"_autoprocessed.csv")
# + slideshow={"slide_type": "-"}
# Example of persisting table with SQL Magic
testdict={"animal":["dog",'cat'],'size':[30,15]}
zoop = pd.DataFrame(testdict)
tname = 'zoop'
# %sql DROP TABLE $tname
# %sql PERSIST $tname
# -
# %sql SELECT * from autoprocessed_i2b2ontology
#engine.execute("SELECT * FROM autoprocessed_i2b2ontology").fetchall()
# +
#Workspace, working on folder check code
mydf = dfs[0]
mymerge = mydf.merge(mydf,left_on=mydf.columns[3],right_on=mydf.columns[4],how='inner',suffixes=['','_r']).groupby('c_fullname').size().reset_index().rename(lambda x: 'size' if x==0 else x,axis='columns')
mymerge=mydf.merge(mymerge,
left_on='c_fullname',right_on='c_fullname',how='left')
print(mymerge['size'] > 0)
# -
| ontology_tools/concern_build_ont.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
# # Summary of lecture 1
# Yesterday I introduced the language Julia, a language that is compiled Just in Time: the first time you run a function, it determines the types of the arguments and compiles it, so the next time is faster.
# This has a list of advantages:
# 1. it is easy to type generic code
# 2. the user does not need to specify the types as he writes the code, making it easier and faster to reuse code
# 3. the second execution of a function is really fast
#
# This puts together the advantages of interpreted languages, as Python and Matlab, with the speed of compiled languages as C++ and Fortran.
#
# Summarizing: Julia is faster than Matlab and Python and simpler to program than C++ and Fortran.
#
#
#
# We used Julia to implement some non-rigorous numerical experiments in Ergodic Theory. While this experiments may be used to gather evidence of some underlying mathematical phenomenon, they are not mathematical proofs.
# The objective of our course is to develop tools that allow us to prove, with the assistance of a computer, Theorems in Computational Ergodic Theory.
#
# To do so, we need to introduce some new tools, both practical and theoretical.
#
# Today we will introduce Interval Arithmetics, a numerical concept that allows a computer to compute rigorous enclosures numerical expressions, i.e., intervals that are guaranteed to contain the __TRUE__ mathematical result of a function evaluation.
# # Introducing Interval Arithmetic
# In this notebook I will introduce one of the main tools of validated numerics: interval arithmetics.
# We will install a ready made package, the IntervalArithmetic.jl package, a package in the shared Julia repository.
import Pkg;
Pkg.add("IntervalArithmetic")
using IntervalArithmetic
# The main idea behind IntervalArithmetic is the following, given a function $f$, Interval Arithmetics allows us to define a function $F$, called the __interval extension__ of $f$, such that for any interval $I = [a,b]$ we have that
# $$
# f(I)\subset F(I).
# $$
# In other words, Interval arithmetics allows the computer to compute numerically an interval that contains the true mathematical result of the function.
# This is called the __range enclosure__ property.
# A reference on the topic is the following book by [<NAME> - Validated Numerics - A Short Introduction to Rigorous computation](https://www.amazon.de/-/en/Warwick-Tucker/dp/0691147817)
# The simplest object of Interval Arithmetics are the intervals.
# The macro @interval defines $x$ as the smallest interval that contains the real point $0.1$.
#
# Remark that $0.1$ has infinite binary expansion, so it is not possible to represent it exactly in binary format, so this is a wide interval, with a lower bound different from the upper bound; the upper and lower bounds are the smallest representable number in Floating Point Arithmetic bigger than $0.1$ and the greatest representable number smaller than $x$
@format standard
x = @interval 0.1
# I will change the output format of the intervals, to a midpoint radius format.
@format midpoint
x
@format standard
y = @interval 0.3
# The operations in interval arithmetic are defined so that the sum of two intervals is an interval that contains the sum of all possible intervals (in some sense, this is similar to absolute error estimate as in the statistics of physical experiments).
x+y
# The same works with multiplication.
x*y
# Please note that wide intervals may be defined.
x = @interval -1 1
x+y
# When an interval contains $0$, the real extended line formalism is used.
# This is called the affine line extension in Tucker's book.
y/x
# Summarizing: Interval arithmetics computes intervals of confidence.
#
# __The result of an operation done with interval arithmetics is guaranteed to contain the true mathematical result of the operation.__
# ## Function evaluation
# Library routines are defined so that the __range enclosure__ property is satisfied. This allows us to get rigorous numerical results on mathematical functions.
f(x) = x*sin(1/x)
x = @interval 0 0.1
f(x)
using Plots
N = 16
X1 = [@interval i/N (i+1)/N for i in 0:N-1]
plot(f, 0, 1)
plot!(IntervalBox.(X1, f.(X1)))
# It is important to observe, in the plot above, that while the graph of the function is computed nonrigorously, and therefore it is not validate, the interval computation is validated, so the rectangles in pink have the strength of theorems.
N = 128
X2 = [@interval i/N (i+1)/N for i in 0:N-1]
plot(f, 0, 1)
plot!(IntervalBox.(X2, f.(X2)))
# The important thing to stress is that the evaluation of a function on a correctly defined interval has the strength of a mathematical theorem, i.e.,
f(@interval 0.5 0.7)
# can be read, in other words as, for any $x$ in $[0.5, 0.7]$ we have that $0.454648\leq f(x)\leq 0.700001$.
# Please remark that this is a totally different scenario from numerical computations, where usually the statements are something as: if the algorithm is stable we get a result which is near the real result (called forward error bound) or if the algorithm is stable it solves a problem which is near our original problem (called backward error bound).
# __Exercise__ Evaluate the function $sin(x)$ on the intervals $[0, 0.125]$, $[0, 0.25]$, $[0, 0.5]$, $[0, \pi]$, $[0, 3\pi]$.
# Remember that you can type \pi+TAB to write the character $\pi$.
sin(@interval 0 2*π)
# ## How to use this
# Due to the range enclosure property we can use this, as an example, to exclude intervals that surely __not contain__ zero. There may be intervals that seem to contain zero due to the overestimates of interval arithmetics, but those we excluded we are sure they do not contain zero.
function exclude_not_contain_zero(f, X)
may_contain_zeros = Interval[]
for x in X
if contains_zero(f(x))
append!(may_contain_zeros, x)
end
end
return may_contain_zeros
end
X = [@interval i/N (i+1)/N for i in 0:N-1]
Xzeros = exclude_not_contain_zero(f, X)
plot(f, 0, 1)
plot!(IntervalBox.(Xzeros, f.(Xzeros)))
# Refining the intervals this gives us a way to identify small intervals that may contain zero; please note that this does not guarantee they contain zero, I will show some methods how to find zeros in a moment.
N = 512
X = [@interval i/N (i+1)/N for i in 0:N-1]
Xzeros = exclude_not_contain_zero(f, X)
plot(f, 0, 0.35)
plot!(IntervalBox.(Xzeros, f.(Xzeros)))
# While this may seem nothing extreme, please remark that this can be used to prove that a function is nonzero in a given interval. This can be used to exclude parameters.
# ## The interval Newton method
# I will introduce now the workhorse of Validated Numerics, the interval Newton method.
# To do so, we remember the [mean value theorem](https://en.wikipedia.org/wiki/Mean_value_theorem):
# if $f$ is a continuous function on $[a, b]$ and differentiable on $(a,b)$ then there exists a point
# $c$ in $(a,b)$ such that
# $$
# f(b)-f(a) = f'(c)(b-a).
# $$
# We want now to find a solution to the equation
# $$
# f(x) = 0.
# $$
# By the mean value theorem, if $\tilde{x}$ is a solution of the equation and $x$ is another, nearby point, we have
# $$
# f(x) = f'(c)(x-\tilde{x}),
# $$
# and, if $f'(c)\neq 0$, this implies that
# $$
# \tilde{x} = x-\frac{f(x)}{f'(c)};
# $$
# this is the motivation behind the Newton iteration method, i.e., building the sequence
# $$
# x_{n+1} = x_{n}-\frac{f(x_n)}{f'(x_n)}.
# $$
# Please remark that under some conditions, this method converges to a root, but we do not know which root (there may be more than one) nor we can bound the distance between $x_{n}$ and $\tilde{x}$.
# __Important__ please remark that if we knew the point $c$, we could compute exactly the root in one step. But the mean value theorem is a result of existence, it does not tell us which point is the point $c$.
# The interval Newton method answers all these issues:
# 1. it proves whether an interval contains a root or not
# 2. it returns us an interval guaranteed to contain a root (this has the strength of a mathematical proof)
# 3. it can prove the fact that the root inside the interval is unique.
# The idea is similar to the one used to build the Newton iteration. Suppose we are looking for a root of a function in the interval $I$; then, if $m$ is the midpoint of the interval and $\tilde{x}$ is the root, we have
# that
# $$
# \tilde{x} = m-\frac{f(m)}{f'(c)}
# $$
# where $c$ is a point in the interval $I$. Now, interval arithmetics allows us to compute an interval that contains $f'(I)$, which in turn contains $f'(c)$.
# So, given an interval $I$ we can compute a new interval
# $$
# N(f, I) = m-\frac{f(m)}{f'(I)}.
# $$
#
# In Tucker book it is proved that :
# - if $I$ contains a root, then $N(f, I)$ contains a root.
# - if $N(f,I)\cap I$ is empty, we know that the interval contains no root. If it is not empty we know it contains at least a root.
# - If $N(f, I)$ is strictly contained in $I$ the root is unique.
#
# So, we have now a tool that allows us to rigorously enclose the roots of a function.
Newton(f, fprime, I::Interval{T}) where {T} = intersect(Interval{T}(mid(I))-f(Interval{T}(mid(I)))/fprime(I), I)
x = Newton(sin, cos, @interval 3 4)
# As you can see, the result of the Newton interval step is telling us that the function $sin$ has a unique $0$ in $[3, 4]$ and giving us a tighter interval that contains this $0$.
# We can iterate this process.
for i in 1:10
x = Newton(sin, cos, x)
end
println(x)
println(diam(x))
# So, we have computed an interval that contains $\pi$, which is the $0$ of the function $sin$ in $[3, 4]$, the interval has a diameter of $4.45\cdot e-16$. Please remark again, that this has the strength of a mathematical theorem, i.e., the interval $x$ has been proved, with the aid of a computer, to contain the value of $\pi$.
# If we want tighter bounds, we can use higher precision floating point arithmetics in our intervals.
setprecision(1024)
x = Interval{BigFloat}(x) # we will refine the interval computed in Float64
for i in 1:10
x = Newton(sin, cos, x)
end
println(x)
println(diam(x))
# Again, this is a theorem, we have that $\pi$ is bounded below by
x.lo
# and above by
x.hi
# Using Automatic Differentiation, the code for the Newton method can be written in an even simpler way.
using DualNumbers
Newton(f, I) = Newton(f, x->(f(Dual(x, 1)).epsilon), I)
# We can use this to certify the intervals that may contain the the zeros of the function $f(x) = x \cdot sin(1/x)$.
new_x = [Newton(f, x) for x in Xzeros]
plot(f, 0, 0.35)
plot!(IntervalBox.(new_x, f.(new_x)))
# The last two intervals are so thin that the plotting function is not rendering them exactly, but the Newton method confirmed them. What is striking is that the Newton method allowed us to prove the existence of at least one zero in each one of the intervals above.
#
# We will zoom to $[0.3180, 0.3185]$.
plot(f, 0.3180, 0.3185)
plot!(IntervalBox.(new_x[end], f.(new_x[end])))
# __Exercise__: Modify the Interval Newton Method to solve the equation $f(x)=y$; can you implement it in such a way that $y$ may be an interval? What happens when you use to solve an equation $f(x)=y$ with $y$ a wide interval?
# ## Taylor Models
# I will introduce now another tool of Validated Numerics, used to get rigorous approximations of functions, called Taylor Models.
#
# They are used in the rigorous computation of integrals and the rigorous computation of enclosures of trajectories of ODE.
# We want to be able to approximate a function by a polynomial with rigorous error bound.
import Pkg; Pkg.add("TaylorModels")
using TaylorModels
f(x) = x*(x-1)*(x+2)*(x+3)^2*(x+7)*sin(2*π*x+0.5)
I = @interval -0.5 0.5 # Domain
plot(f, -0.5, 0.5)
# A Taylor model is a polynomial $P$ of degree $k$ and an interval $\Delta$ such that given a function $f$, on an interval $I$, $P$ is the Taylor polynomial of order $k$ at the center of the interval and $\Delta$ is an interval such that
# $$
# f(x)-P(x) \in \Delta
# $$
TM10 = TaylorModel1(10, I) # Taylor model of order 10
TM15 = TaylorModel1(15, I) #Taylor model of order 15
# Plese note that the Taylor model is centered at the midpoint of $I$.
?TaylorModel1
FM10 = f(TM10)
# So, on $[-0.5, 0.5]$ the error between the value of $f$ and the value of the Taylor Model is contained in the interval $[-1.50153, 2.61226]$
# On the same interval the Taylor model of order $15$ is the following.
FM15 = f(TM15)
# As expected, passing from a linear approximation With higher order, we get smaller $\Delta$.
plot(f, -0.5, 0.5)
plot!(FM10, color=:red)
plot!(FM15, color = :green)
# __Exercise__ : Find the Taylor models of order $5$, $10$ and $15$ of the function $f$ on the interval $[-0.125, 0.125]$. What happens to $\Delta$?
# ## Rigorous Integration
# We are interested in computing rigorously the value of an integral. To do so, we will use Taylor Models.
f(x) = sin(exp(1/x))
plot(f, 0.125, 1)
# In this section I will show a tecnique to find intervals that enclose the true value of an integral. To show the power of the method I will compute the integral of an oscillating integral.
# Having a Taylor expansion centered at the center of the interval allows us to easily compute the integral of the function over the integral.
# Let $m$ be the midpoint of the interval, $r = x-m$ and $a_i$ the coefficients of $P$:
# $$
# \int_I f dx = \sum_{i=0}^n a_i \int_I (x-m)^i dx +\int_I f(x)-P(x) dx,
# $$
# so the integral is contained in the interval with center
# $$
# 2\sum_{i=0}^{\lfloor n/2 \rfloor} a_{2i}\frac{r^{2i+1}}{2i+1}
# $$
# and radius $\Delta\cdot |I|$ (where $|I|$ is the length of I).
function integrate1(f, I; steps = 1024, degree = 6)
lo = I.lo
hi = I.hi
l = diam(I)
int_center = Interval(0.0)
int_error = Interval(0.0)
for i in 1:steps
left = lo+(i-1)*(l/steps)
right = lo+i*l/steps
r = (1/2)*(l/steps)
J = interval(left, right)
TM = TaylorModel1(degree, J)
FM = f(TM)
#@info FM
for i in 0:Int64(floor(degree/2))
int_center+=2*(FM.pol[2*i]*r^(2*i+1))/(2*i+1)
int_error +=2*FM.rem*r
end
end
return int_center+int_error
end
integrate1(f, @interval 0.25 1; steps = 1024, degree = 6)
# This integral method can be made adaptive, both in the size of the interval $J$ and the degree of the Taylor expansion. This has the strength of a mathematical proof.
function adaptive_integration(f, I::Interval; tol = 2^-10, steps = 8, degree = 6) # tol 2^-10, steps = 8 are default values
lo = I.lo
hi = I.hi
l = diam(I)
int_value = Interval(0.0)
for i in 1:steps
left = lo+(i-1)*(l/steps)
right = lo+i*l/steps
Istep = Interval(left, right)
val = integrate1(f, Istep)
if radius(val)<tol
int_value += val
else
I₁, I₂ = bisect(I)
val₁ = adaptive_integration(f, I₁; tol = tol/2, steps = steps, degree = degree+2)
val₂ = adaptive_integration(f, I₁; tol = tol/2, steps = steps, degree = degree+2)
int_value +=val₁+val₂
end
end
return int_value
end
@time adaptive_integration(f, @interval 0.125 1)
# __Exercise__: Try to push the computation of the integral further near $0$.
# # Summary of the lecture
# In this lecture I introduced some tools that allow us to prove Theorems with the help of a computer.
# 1. Interval Arithmetic, that allows us to rigorously enclose the range of a function on an interval $I$
# 2. Interval Newton Method, that allows us to find an interval that is proved to contain the solution of the equation $f(x)=y$, allowing us to rigorously find inverse images
# 3. Taylor Models that allow us to approximate functions by polynomials with rigorous and explicit error bounds
# 4. Rigorous integration that allows us to compute rigorously the value of an integral over an Interval.
#
# All these tools are the engine of our implementation of the Ulam method: we use the Interval Newton method to compute preimages, computing an Interval of entries which contains the entries of the true abstract matrix.
#
# This is going to allow us to approximate the invariant density $h$ of the physical measure of a system.
#
# Then are going to use rigorous integration to compute the Birkhoff averages of observables in a rigorous way, through the identity
# $$
# \lim_{n\to +\infty}\frac{1}{n}\sum_{i=0}^{n-1} \phi(T^i(x))=\int \phi h dx.
# $$
| Lecture 2 - Interval Arithmetics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°10
#
#
# <a id='p1'></a>
# ## I.- Problema 01
#
#
# <img src="https://www.goodnewsnetwork.org/wp-content/uploads/2019/07/immunotherapy-vaccine-attacks-cancer-cells-immune-blood-Fotolia_purchased.jpg" width="360" height="360" align="center"/>
#
#
# El **cáncer de mama** es una proliferación maligna de las células epiteliales que revisten los conductos o lobulillos mamarios. Es una enfermedad clonal; donde una célula individual producto de una serie de mutaciones somáticas o de línea germinal adquiere la capacidad de dividirse sin control ni orden, haciendo que se reproduzca hasta formar un tumor. El tumor resultante, que comienza como anomalía leve, pasa a ser grave, invade tejidos vecinos y, finalmente, se propaga a otras partes del cuerpo.
#
# El conjunto de datos se denomina `BC.csv`, el cual contine la información de distintos pacientes con tumosres (benignos o malignos) y algunas características del mismo.
#
#
# Las características se calculan a partir de una imagen digitalizada de un aspirado con aguja fina (FNA) de una masa mamaria. Describen las características de los núcleos celulares presentes en la imagen.
# Los detalles se puede encontrar en [K. <NAME> and <NAME>: "Robust Linear Programming Discrimination of Two Linearly Inseparable Sets", Optimization Methods and Software 1, 1992, 23-34].
#
#
# Lo primero será cargar el conjunto de datos:
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
from sklearn.datasets import load_digits
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
from sklearn.dummy import DummyClassifier
from sklearn.cluster import KMeans
# %matplotlib inline
sns.set_palette("deep", desat=.6)
sns.set(rc={'figure.figsize':(11.7,8.27)})
# -
# cargar datos
df = pd.read_csv(os.path.join("data","BC.csv"), sep=",")
df['diagnosis'] = df['diagnosis'] .replace({'M':1,'B':0}) # target
df.head()
# Basado en la información presentada responda las siguientes preguntas:
#
# 1. Realice un análisis exploratorio del conjunto de datos.
# 1. Normalizar las variables numéricas con el método **StandardScaler**.
# 3. Realizar un método de reducción de dimensionalidad visto en clases.
# 4. Aplique al menos tres modelos de clasificación distintos. Para cada uno de los modelos escogidos, realice una optimización de los hiperparámetros. además, calcule las respectivas métricas. Concluya.
#
#
#
# ### Análisis exploratorio del conjunto de datos:
print('----------------------')
print('Media de cada variable')
print('----------------------')
df.mean(axis=0)
print('-------------------------')
print('Varianza de cada variable')
print('-------------------------')
df.var(axis=0)
df.describe()
# Veamos cuantos tumores son malignos y cuantos beningnos:
B = df[df["diagnosis"]==0]
M = df[df["diagnosis"]==1]
B
M
# Se concluye que 357 tumores son benignos (B) contra 212 malignos (M).
# ### Normalizar las variables numéricas con el método StandardScaler:
# +
scaler = StandardScaler()
df[df.columns.drop(["id","diagnosis"])] = scaler.fit_transform(df[df.columns.drop(["id","diagnosis"])])
df.head()
# -
# ### Realizar un método de reducción de dimensionalidad visto en clases:
# +
# Entrenamiento modelo PCA con escalado de los datos
# ==============================================================================
pca_pipe = make_pipeline(StandardScaler(), PCA())
pca_pipe.fit(df)
# Se extrae el modelo entrenado del pipeline
modelo_pca = pca_pipe.named_steps['pca']
# -
# Se combierte el array a dataframe para añadir nombres a los ejes.
pd.DataFrame(
data = modelo_pca.components_,
columns = df.columns,
index = ['PC1', 'PC2', 'PC3', 'PC4',
'PC5', 'PC6', 'PC7', 'PC8',
'PC9', 'PC10', 'PC11', 'PC12',
'PC13', 'PC14', 'PC15', 'PC16',
'PC17', 'PC18', 'PC19', 'PC20',
'PC21', 'PC22', 'PC23', 'PC24',
'PC25', 'PC26', 'PC27', 'PC28',
'PC29', 'PC30', 'PC31', 'PC32']
)
# Heatmap componentes
# ==============================================================================
plt.figure(figsize=(12,14))
componentes = modelo_pca.components_
plt.imshow(componentes.T, cmap='viridis', aspect='auto')
plt.yticks(range(len(df.columns)), df.columns)
plt.xticks(range(len(df.columns)), np.arange(modelo_pca.n_components_) + 1)
plt.grid(False)
plt.colorbar();
# +
# graficar varianza por componente
percent_variance = np.round(modelo_pca.explained_variance_ratio_* 100, decimals =2)
columns = ['PC1', 'PC2', 'PC3', 'PC4',
'PC5', 'PC6', 'PC7', 'PC8',
'PC9', 'PC10', 'PC11', 'PC12',
'PC13', 'PC14', 'PC15', 'PC16',
'PC17', 'PC18', 'PC19', 'PC20',
'PC21', 'PC22', 'PC23', 'PC24',
'PC25', 'PC26', 'PC27', 'PC28',
'PC29', 'PC30', 'PC31', 'PC32']
plt.figure(figsize=(20,10))
plt.bar(x= range(1,33), height=percent_variance, tick_label=columns)
plt.xticks(np.arange(modelo_pca.n_components_) + 1)
plt.ylabel('Componente principal')
plt.xlabel('Por. varianza explicada')
plt.title('Porcentaje de varianza explicada por cada componente')
plt.show()
# +
# graficar varianza por la suma acumulada de los componente
percent_variance_cum = np.cumsum(percent_variance)
#columns = ['PC1', 'PC1+PC2', 'PC1+PC2+PC3', 'PC1+PC2+PC3+PC4',.....]
plt.figure(figsize=(12,4))
plt.bar(x= range(1,33), height=percent_variance_cum, #tick_label=columns
)
plt.ylabel('Percentate of Variance Explained')
plt.xlabel('Principal Component Cumsum')
plt.title('PCA Scree Plot')
plt.show()
# -
# Proyección de las observaciones de entrenamiento
# ==============================================================================
proyecciones = pca_pipe.transform(X=df)
proyecciones = pd.DataFrame(
proyecciones,
columns = ['PC1', 'PC2', 'PC3', 'PC4',
'PC5', 'PC6', 'PC7', 'PC8',
'PC9', 'PC10', 'PC11', 'PC12',
'PC13', 'PC14', 'PC15', 'PC16',
'PC17', 'PC18', 'PC19', 'PC20',
'PC21', 'PC22', 'PC23', 'PC24',
'PC25', 'PC26', 'PC27', 'PC28',
'PC29', 'PC30', 'PC31', 'PC32'],
index = df.index
)
proyecciones.head()
# ### Aplique al menos tres modelos de clasificación distintos. Para cada uno de los modelos escogidos, realice una optimización de los hiperparámetros. además, calcule las respectivas métricas. Concluya.
# +
df3 = pd.get_dummies(df)
df3.head()
# +
X = np.array(df3)
kmeans = KMeans(n_clusters=8,n_init=25, random_state=123)
kmeans.fit(X)
centroids = kmeans.cluster_centers_ # centros
clusters = kmeans.labels_ # clusters
# -
# etiquetar los datos con los clusters encontrados
df["cluster"] = clusters
df["cluster"] = df["cluster"].astype('category')
centroids_df = pd.DataFrame(centroids)
centroids_df["cluster"] = [1,2,3,4,5,6,7,8]
# +
# implementación de la regla del codo
Nc = [5,10,20,30,50,75,100,200,300]
kmeans = [KMeans(n_clusters=i) for i in Nc]
score = [kmeans[i].fit(df).inertia_ for i in range(len(kmeans))]
df_Elbow = pd.DataFrame({'Number of Clusters':Nc,
'Score':score})
df_Elbow.head()
# -
# graficar los datos etiquetados con k-means
fig, ax = plt.subplots(figsize=(11, 8.5))
plt.title('Elbow Curve')
sns.lineplot(x="Number of Clusters",
y="Score",
data=df_Elbow)
sns.scatterplot(x="Number of Clusters",
y="Score",
data=df_Elbow)
plt.show()
# +
# PCA
#scaler = StandardScaler()
X = df.drop(columns=["id","diagnosis"])
y = df['diagnosis']
embedding = PCA(n_components=2)
X_transform = embedding.fit_transform(X)
df_pca = pd.DataFrame(X_transform,columns = ['Score1','Score2'])
df_pca['diagnosis'] = y
# +
# Plot Digits PCA
# Set style of scatterplot
sns.set_context("notebook", font_scale=1.1)
sns.set_style("ticks")
# Create scatterplot of dataframe
sns.lmplot(x='Score1',
y='Score2',
data=df_pca,
fit_reg=False,
legend=True,
height=9,
hue='diagnosis',
scatter_kws={"s":200, "alpha":0.3})
plt.title('PCA Results: BC', weight='bold').set_fontsize('14')
plt.xlabel('Prin Comp 1', weight='bold').set_fontsize('10')
plt.ylabel('Prin Comp 2', weight='bold').set_fontsize('10')
# +
from sklearn.datasets import make_moons, make_circles, make_classification
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from matplotlib.colors import ListedColormap
# +
X = df_pca.drop(columns='diagnosis')
y = df_pca['diagnosis']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
h = .02 # step size in the mesh
plt.figure(figsize=(12,12))
names = ["Logistic",
"RBF SVM",
"Decision Tree",
"Random Forest"
]
classifiers = [
LogisticRegression(),
SVC(gamma=2, C=1),
DecisionTreeClassifier(max_depth=5),
RandomForestClassifier(max_depth=5, n_estimators=10, max_features=1),
]
X, y = make_classification(n_features=2, n_redundant=0, n_informative=2,
random_state=1, n_clusters_per_class=1)
rng = np.random.RandomState(2)
X += 2 * rng.uniform(size=X.shape)
linearly_separable = (X, y)
datasets = [make_moons(noise=0.3, random_state=0),
make_circles(noise=0.2, factor=0.5, random_state=1),
linearly_separable
]
figure = plt.figure(figsize=(27, 9))
i = 1
# iterate over datasets
for ds_cnt, ds in enumerate(datasets):
# preprocess dataset, split into training and test part
X, y = ds
X = StandardScaler().fit_transform(X)
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# just plot the dataset first
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
if ds_cnt == 0:
ax.set_title("Input data")
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6,
edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
i += 1
# iterate over classifiers
for name, clf in zip(names, classifiers):
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
ax.contourf(xx, yy, Z, cmap=cm, alpha=.8)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
edgecolors='k', alpha=0.6)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
if ds_cnt == 0:
ax.set_title(name)
ax.text(xx.max() - .3, yy.min() + .3, ('%.2f' % score).lstrip('0'),
size=15, horizontalalignment='right')
i += 1
plt.tight_layout()
plt.show()
# +
from metrics_classification import *
class SklearnClassificationModels:
def __init__(self,model,name_model):
self.model = model
self.name_model = name_model
@staticmethod
def test_train_model(X,y,n_size):
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size=n_size , random_state=42)
return X_train, X_test, y_train, y_test
def fit_model(self,X,y,test_size):
X_train, X_test, y_train, y_test = self.test_train_model(X,y,test_size )
return self.model.fit(X_train, y_train)
def df_testig(self,X,y,test_size):
X_train, X_test, y_train, y_test = self.test_train_model(X,y,test_size )
model_fit = self.model.fit(X_train, y_train)
preds = model_fit.predict(X_test)
df_temp = pd.DataFrame(
{
'y':y_test,
'yhat': model_fit.predict(X_test)
}
)
return df_temp
def metrics(self,X,y,test_size):
df_temp = self.df_testig(X,y,test_size)
df_metrics = summary_metrics(df_temp)
df_metrics['model'] = self.name_model
return df_metrics
# +
# metrics
import itertools
# nombre modelos
names_models = ["Logistic",
"RBF SVM",
"Decision Tree",
"Random Forest"
]
# modelos
classifiers = [
LogisticRegression(),
SVC(gamma=2, C=1),
DecisionTreeClassifier(max_depth=5),
RandomForestClassifier(max_depth=5, n_estimators=10, max_features=1),
]
datasets
names_dataset = ['make_moons',
'make_circles',
'linearly_separable'
]
X, y = make_classification(n_features=2, n_redundant=0, n_informative=2,
random_state=1, n_clusters_per_class=1)
rng = np.random.RandomState(2)
X += 2 * rng.uniform(size=X.shape)
linearly_separable = (X, y)
datasets = [make_moons(noise=0.3, random_state=0),
make_circles(noise=0.2, factor=0.5, random_state=1),
linearly_separable
]
# juntar informacion
list_models = list(zip(names_models,classifiers))
list_dataset = list(zip(names_dataset,datasets))
frames = []
for x in itertools.product(list_models, list_dataset):
name_model = x[0][0]
classifier = x[0][1]
name_dataset = x[1][0]
dataset = x[1][1]
X = dataset[0]
Y = dataset[1]
fit_model = SklearnClassificationModels( classifier,name_model)
df = fit_model.metrics(X,Y,0.2)
df['dataset'] = name_dataset
frames.append(df)
# -
# juntar resultados
pd.concat(frames)
| labs/lab_10_Moreno.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import Libraries and Data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
import seaborn as sns
# %matplotlib inline
data = pd.read_csv(r'C:\Users\somendra.singh\Desktop\Data Science codes\Stats and ML july\Machine Learning\diabetes.csv')
data.head()
# ## Correlation Check
correlations = data.corr()
correlations
# ## Visualizing the data for any Relations
# +
def visualise(data):
fig, ax = plt.subplots()
ax.scatter(data.iloc[:,1].values, data.iloc[:,5].values)
ax.set_title('Highly Correlated Features')
ax.set_xlabel('Plasma glucose concentration')
ax.set_ylabel('Body mass index')
visualise(data)
# -
# ## Replacing the Zeros with Null values
data[['Glucose','BMI']] = data[['Glucose','BMI']].replace(0, np.NaN)
data.dropna(inplace=True)
visualise(data)
# ## Feature Selection
X = data[['Glucose','BMI','Pregnancies','BloodPressure','SkinThickness','Insulin',
'DiabetesPedigreeFunction','Age']].values
y = data[['Outcome']].values
# ## Standardization & Scaling of Features
sc = StandardScaler()
X = sc.fit_transform(X)
mean = np.mean(X, axis=0)
print('Mean: (%d, %d)' % (mean[0], mean[1]))
standard_deviation = np.std(X, axis=0)
print('Standard deviation: (%d, %d)' % (standard_deviation[0], standard_deviation[1]))
print(X[0:10,:])
# ## Train-Test Split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# ## Logistic Regression Model
# instantiate the model (using the default parameters)
logreg = LogisticRegression()
# fit the model with data
logreg.fit(X_train,y_train)
# ## Predictions
y_pred=logreg.predict(X_test)
y_pred
# ## Performance & Accuracy
cnf_matrix = metrics.confusion_matrix(y_test, y_pred)
cnf_matrix
class_names=[0,1] # name of classes
fig, ax = plt.subplots()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
# create heatmap
sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu" ,fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
print("Precision:",metrics.precision_score(y_test, y_pred))
print("Recall:",metrics.recall_score(y_test, y_pred))
y_pred_proba = logreg.predict_proba(X_test)[::,1]
fpr, tpr, _ = metrics.roc_curve(y_test, y_pred_proba)
auc = metrics.roc_auc_score(y_test, y_pred_proba)
plt.plot(fpr,tpr,label="data 1, auc="+str(auc))
plt.legend(loc=4)
plt.show()
| Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### You can write code in abstract methods
# +
from abc import ABC, abstractmethod
class Automobile(ABC):
def __init__(self, no_of_wheels):
self.no_of_wheels = no_of_wheels
print("Automobile Created")
@abstractmethod
def start(self):
print("start of Automobile called")
@abstractmethod
def stop(self):
pass
@abstractmethod
def drive(self):
pass
@abstractmethod
def get_no_of_wheels(self):
return self.no_of_wheels
class Car(Automobile):
def start(self):
super().start()
print("start of Car called")
def stop(self):
pass
def drive(self):
pass
def get_no_of_wheels(self):
return super().get_no_of_wheels()
class Bus(Automobile):
def start(self):
pass
def stop(self):
pass
def drive(self):
pass
def get_no_of_wheels(self):
return super().get_no_of_wheels()
c = Car(4)
b = Bus(6)
print(c.get_no_of_wheels())
print(b.get_no_of_wheels())
# +
#Predict the Output:
from abc import ABC, abstractmethod
class A(ABC):
@abstractmethod
def fun1(self):
print("function of class A called")
@abstractmethod
def fun2(self):
pass
class B(A):
def fun1(self):
print("function 1 called")
def fun2(self):
print("function 2 called")
o = B()
o.fun1()
# +
#Predict the Output:
from abc import ABC, abstractmethod
class A(ABC):
@abstractmethod
def fun1(self):
print("function of class A called")
@abstractmethod
def fun2(self):
pass
class B(A):
def fun1(self):
super().fun1()
def fun2(self):
print(" function 2 called")
o = B()
o.fun1()
# -
| 05 OOPS-3/5.2 Abstract class cont.....ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python3]
# language: python
# name: conda-env-python3-py
# ---
# + [markdown] deletable=true editable=true
# # Examples
#
# Last updated: July 12, 2017
#
# Below are a few examples to get you started.
#
# + deletable=true editable=true
import numpy as np
import matplotlib.pyplot as plt
from ftperiodogram.modeler import FastTemplatePeriodogram, FastMultiTemplatePeriodogram, TemplateModel
from ftperiodogram.template import Template
from ftperiodogram.utils import ModelFitParams
rand = np.random
# + [markdown] deletable=true editable=true
# ## The `Template` class
#
# There are two ways of instantiating the `Template` class, which is needed by both the `FastTemplatePeriodogram` and `FastMultiTemplatePeriodogram` classes.
#
# ### If you have a template defined by `phase`, `y_phase`, use the `.from_sampled` class method to instantiate:
# + deletable=true editable=true
# Make a template
phase = np.linspace(0, 1, 100)
amps = [ 0.5, 0.2, 0.3 ]
phases = [ -1., 1.3, 0.2 ]
amps = np.array(amps) / np.sqrt(sum(np.power(amps, 2)))
y_phase = sum([ a * np.cos(2 * (n+1) * np.pi * phase - p) \
for n, (a,p) in enumerate(zip(amps, phases)) ])
# Instantiate the `Template` class
our_template = Template.from_sampled(y_phase, nharmonics=3)
# Plot a comparison
f, ax = plt.subplots()
ax.plot(phase, y_phase, color='k', lw=2, alpha=0.5)
ax.plot(phase, our_template(phase), color='r')
ax.set_xlabel('phase')
ax.set_xlim(0,1)
plt.show()
# -
# ### If you know the Fourier coefficients of the template:
# +
# Obtain the Fourier coefficients from the last template fit
c_n, s_n = our_template.c_n, our_template.s_n
# Instantiate a new `Template` using coefficients
new_template = Template(c_n, s_n)
# Check that both templates are equivalent
print(all(new_template(phase) == our_template(phase)))
# -
# ## The `TemplateModel` class
#
# The `TemplateModel` class stores a template and a set of fit parameters. When called, the model will return
#
# $\hat{y}(t) = aM(\omega(t - \tau)) + c$
#
# where $M(\phi)$ is the template, and $a, \omega, \tau, c$ are the fit parameters. The parameters must be an instance of `ModelFitParams`, which is simply an ordered dictionary with the attributes `a`, `b`, `c`, `sgn`. The `b` and `sgn` variables represent $\cos(\omega \tau)$ and ${\rm sign}(sin(\omega\tau))$, respectively.
# +
# set some model parameters (arbitrary)
params = ModelFitParams(a=1., b=0.3, c=0.2, sgn=1)
#
model = TemplateModel(our_template, frequency=1.0, parameters=params)
t = np.linspace(0, 3, 300)
y0 = model(t)
f, ax = plt.subplots()
ax.plot(t, y0)
plt.show()
# -
# ## Template periodogram, single template -- `FastTemplatePeriodogram`
#
# The `FastTemplatePeriodogram` can be used to fit a (single) periodic template to irregularly sampled time-series data at a number of frequencies.
#
# ### Accessing the best-fit model
#
# After running `power` or `autopower`, the `FastTemplatePeriodogram` will keep the best-fit model in the form of a `TemplateModel` instance accessible via the `.best_model` attribute.
# +
# Instantiate periodogram with template
ftp = FastTemplatePeriodogram(template=our_template)
# Generate random data (using template)
ndata = 30
sigma = 0.2
freq = 20.
t = np.sort(rand.rand(ndata))
phi = (t * freq) % 1.0
y = ftp.template(phi) + sigma * rand.randn(ndata)
yerr = sigma * np.ones_like(y)
# Provide periodogram with data
ftp.fit(t, y, yerr)
# Produce a periodogram over an automatically-determined
# set of frequencies for the given template
freqs, powers = ftp.autopower(samples_per_peak=10)
f, (axper, axfit) = plt.subplots(1, 2, figsize=(12, 3))
best_freq = freqs[np.argmax(powers)]
# Plot periodogram
axper.plot(freqs, powers)
axper.axvline(freq, ls=':', color='0.5')
axper.set_xlabel('frequency')
axper.set_ylabel('power')
axper.set_xlim(min(freqs), max(freqs))
# Plot data
axfit.errorbar((t * best_freq) % 1.0, y, yerr, fmt='o', c='0.5')
axfit.plot(phase, ftp.best_model(phase / best_freq), color='r')
plt.show()
# -
# ## Template periodogram, multiple templates -- `FastMultiTemplatePeriodogram`
#
# If you would like to fit from a selection of templates, you may do so via the `FastMultiTemplatePeriodogram`. At each trial frequency, the `FastMultiTemplatePeriodogram` will pick the template that best fits the data. This is useful for detecting classes of objects that exhibit multiple kinds of periodic fluctuations (for example, RR Lyrae).
# +
# Make two templates
cn1, sn1 = zip(*[ (0.2, 0.2), (0.0, 0.2), (0.2, 0.0) ])
cn2, sn2 = zip(*[ (0.2, 0.1), (0.2, 0.0), (0.0, 0.2) ])
template1 = Template(cn1, sn1)
template2 = Template(cn2, sn2)
# Now, generate some linear combinations of the templates
y_temp1 = template1(phase)
y_temp2 = template2(phase)
signal = lambda x, q=0.5 : q * template1(x) + (1 - q) * template2(x)
templates = []
ftemp, axtemp = plt.subplots()
colormap = plt.get_cmap('viridis')
axtemp.set_title('Templates used for fitting')
axtemp.set_xlabel('Phase')
for q in [ 0.0, 0.5, 1.0 ]:
y_temp = signal(phase, q=q)
templates.append(Template.from_sampled(y_temp, nharmonics=4))
axtemp.plot(phase, templates[-1](phase), color=colormap(q))
axtemp.set_xlim(0, 1)
# generate some new data
q_data = 0.25
y_new = signal(phi, q=q_data) + sigma * rand.rand(len(t))
# instantiate (multi-template) periodogram
ftp_multi = FastMultiTemplatePeriodogram(templates=templates)
# give periodogram data
ftp_multi.fit(t, y_new, yerr)
# evaluate periodogram at a set of automatically determined
# frequencies
freqs_multi, powers_multi = ftp_multi.autopower(samples_per_peak = 10)
best_freq = freqs_multi[np.argmax(powers_multi)]
f2, (axper, axfit) = plt.subplots(1, 2, figsize=(12, 3))
axper.plot(freqs_multi, powers_multi)
axper.axvline(freq, ls=':', color='0.5')
axfit.plot(phase, ftp_multi.best_model(phase/best_freq), color='r')
axfit.plot(phase, signal(phase, q=q_data), color='k', lw=2)
axfit.errorbar((t * best_freq) % 1.0, y_new, yerr, fmt='o', c='0.5')
plt.show()
# -
# ## Single-frequency fits
#
# You may also use either the `FastTemplatePeriodogram` or `FastMultiTemplatePeriodogram` to fit a template at a given frequency using the `fit_model` method.
# +
correct_template = Template.from_sampled(signal(phase, q=q_data), nharmonics=4)
another_model = FastTemplatePeriodogram(template=correct_template).fit(t, y_new, yerr).fit_model(freq)
f,ax = plt.subplots()
ax.plot(phase, another_model(phase/freq), color='r')
ax.plot(phase, signal(phase, q=q_data), color='k', lw=2)
ax.errorbar((t * freq) % 1.0, y_new, yerr, fmt='o', c='0.5')
plt.show()
# -
| notebooks/Examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 課程目標:
#
# 利用神經網路的加法減法數學式來說明梯度下降
# # 範例重點:
#
# 透過網路參數(w, b)的更新可以更容易理解梯度下降的求值過程
# matplotlib: 載入繪圖的工具包
# random, numpy: 載入數學運算的工具包
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
#適用於 Jupyter Notebook, 宣告直接在cell 內印出執行結果
import random as random
import numpy as np
import csv
# # ydata = b + w * xdata
# 給定曲線的曲線範圍
# 給定初始的data
x_data = [ 338., 333., 328., 207., 226., 25., 179., 60., 208., 606.]
y_data = [ 640., 633., 619., 393., 428., 27., 193., 66., 226., 1591.]
#給定神經網路參數:bias 跟weight
x = np.arange(-200,-100,1) #給定bias
y = np.arange(-5,5,0.1) #給定weight
Z = np.zeros((len(x), len(y)))
#meshgrid返回的兩個矩陣X、Y必定是 column 數、row 數相等的,且X、Y的 column 數都等
#meshgrid函數用兩個坐標軸上的點在平面上畫格。
X, Y = np.meshgrid(x, y)
for i in range(len(x)):
for j in range(len(y)):
b = x[i]
w = y[j]
Z[j][i] = 0
for n in range(len(x_data)):
Z[j][i] = Z[j][i] + (y_data[n] - b - w*x_data[n])**2
Z[j][i] = Z[j][i]/len(x_data)
# +
# ydata = b + w * xdata
b = -120 # initial b
w = -4 # initial w
lr = 0.000001 # learning rate
iteration = 100000
# Store initial values for plotting.
b_history = [b]
w_history = [w]
#給定初始值
lr_b = 0.0
lr_w = 0.0
# -
# 在微積分裡面,對多元函數的參數求∂偏導數,把求得的各個參數的偏導數以向量的形式寫出來,就是梯度。
# 比如函數f(x), 對x求偏導數,求得的梯度向量就是(∂f/∂x),簡稱grad f(x)或者▽f (x)。
#
# +
'''
Loss = (實際ydata – 預測ydata)
Gradient = -2*input * Loss
調整後的權重 = 原權重 – Learning * Gradient
'''
# Iterations
for i in range(iteration):
b_grad = 0.0
w_grad = 0.0
for n in range(len(x_data)):
b_grad = b_grad - 2.0*(y_data[n] - b - w*x_data[n])*1.0
w_grad = w_grad - 2.0*(y_data[n] - b - w*x_data[n])*x_data[n]
lr_b = lr_b + b_grad ** 2
lr_w = lr_w + w_grad ** 2
# Update parameters.
b = b - lr * b_grad
w = w - lr * w_grad
# Store parameters for plotting
b_history.append(b)
w_history.append(w)
# -
# plot the figure
plt.contourf(x,y,Z, 50, alpha=0.5, cmap=plt.get_cmap('jet'))
plt.plot([-188.4], [2.67], 'x', ms=12, markeredgewidth=3, color='orange')
plt.plot(b_history, w_history, 'o-', ms=3, lw=1.5, color='black')
plt.xlim(-200,-100)
plt.ylim(-5,5)
plt.xlabel(r'$b$', fontsize=16)
plt.ylabel(r'$w$', fontsize=16)
plt.show()
| D74_Gradient Descent_數學原理/Day74-Gradient Descent_Math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Slicing and Dicing Dataframes
#
# You have seen how to do indexing of dataframes using ```df.iloc``` and ```df.loc```. Now, let's see how to subset dataframes based on certain conditions.
#
# +
# loading libraries and reading the data
import numpy as np
import pandas as pd
df = pd.read_csv("../global_sales_data/market_fact.csv")
df.head()
# -
# ### Subsetting Rows Based on Conditions
#
# Often, you want to select rows which satisfy some given conditions. For e.g., select all the orders where the ```Sales > 3000```, or all the orders where ```2000 < Sales < 3000``` and ```Profit < 100```.
#
# Arguably, the best way to do these operations is using ```df.loc[]```, since ```df.iloc[]``` would require you to remember the integer column indices, which is tedious.
#
# Let's see some examples.
# Select all rows where Sales > 3000
# First, we get a boolean array where True corresponds to rows having Sales > 3000
df.Sales > 3000
# Then, we pass this boolean array inside df.loc
df.loc[df.Sales > 3000]
# An alternative to df.Sales is df['Sales]
# You may want to put the : to indicate that you want all columns
# It is more explicit
df.loc[df['Sales'] > 3000, :]
# We combine multiple conditions using the & operator
# E.g. all orders having 2000 < Sales < 3000 and Profit > 100
df.loc[(df.Sales > 2000) & (df.Sales < 3000) & (df.Profit > 100), :]
# The 'OR' operator is represented by a | (Note that 'or' doesn't work with pandas)
# E.g. all orders having 2000 < Sales OR Profit > 100
df.loc[(df.Sales > 2000) | (df.Profit > 100), :]
# E.g. all orders having 2000 < Sales < 3000 and Profit > 100
# Also, this time, you only need the Cust_id, Sales and Profit columns
df.loc[(df.Sales > 2000) & (df.Sales < 3000) & (df.Profit > 100), ['Cust_id', 'Sales', 'Profit']]
# You can use the == and != operators
df.loc[(df.Sales == 4233.15), :]
df.loc[(df.Sales != 1000), :]
# +
# You may want to select rows whose column value is in an iterable
# For instance, say a colleague gives you a list of customer_ids from a certain region
customers_in_bangalore = ['Cust_1798', 'Cust_1519', 'Cust_637', 'Cust_851']
# To get all the orders from these customers, use the isin() function
# It returns a boolean, which you can use to select rows
df.loc[df['Cust_id'].isin(customers_in_bangalore), :]
| 4_Slicing_Dicing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import csv
import pylab as pp
import matplotlib
# +
os.chdir("D:\PEM article\V_I")
fileName = "V_I.png"
# +
x1 = []
y1 = []
x2 = []
y2 = []
x3 = []
y3 = []
x4 = []
y4 = []
# +
with open('Base.txt','r') as csvfile:
Data1 = csv.reader(csvfile, delimiter=',')
for row in Data1:
x1.append(row[0])
y1.append(row[1])
for i in range(len(x1)):
x1[i] = float(x1[i])
y1[i] = float(y1[i])
#----------------------------------------------------------#
with open('A.txt','r') as csvfile:
Data2 = csv.reader(csvfile, delimiter=',')
for row in Data2:
y2.append(row[0])
x2.append(row[1])
for i in range(len(x2)):
y2[i] = float(y2[i])
x2[i] = float(x2[i])
#----------------------------------------------------------#
with open('B.txt','r') as csvfile:
Data3 = csv.reader(csvfile, delimiter=',')
for row in Data3:
y3.append(row[0])
x3.append(row[1])
for i in range(len(x3)):
y3[i] = float(y3[i])
x3[i] = float(x3[i])
#----------------------------------------------------------#
with open('C - Copy.txt','r') as csvfile:
Data4 = csv.reader(csvfile, delimiter=',')
for row in Data4:
y4.append(row[0])
x4.append(row[1])
for i in range(len(x4)):
y4[i] = float(y4[i])
x4[i] = float(x4[i])
# +
# %matplotlib notebook
pp.autoscale(enable=True, axis='x', tight=True)
pp.plot(x1, y1, marker='o', markerfacecolor='blue', markersize=3, color='skyblue', linewidth=2.5, label = "Base model")
pp.plot(x2, y2, marker='*', color='red', linewidth=1.5, label = "Model A")
pp.plot(x3, y3, marker='', color='green', linewidth=2, linestyle='dashed', label = "Model B")
pp.plot(x4, y4, marker='*', color='olive', linewidth=1.5, label = "Model C")
#pp.legend("Numerical results (single phase)", "Numerical results (two phase)", "Ahmadi et al.", "Wang et al.")
pp.legend()
pp.xlabel(r"Current density $\left(\frac{A}{cm^2}\right)$")
pp.ylabel(r"Cell voltage $(V)$")
fig = matplotlib.pyplot.gcf()
fig.set_size_inches(6, 5)
# -
pp.savefig(fileName, dpi=1200)
| PEM/.ipynb_checkpoints/Plot-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp cycleGAN
# -
#export
from ModernArchitecturesFromPyTorch.nb_XResNet import *
from nbdev.showdoc import show_doc
# # CycleGAN
# > Implementing cycleGAN for unsupervised image to image domain translation. See https://junyanz.github.io/CycleGAN/.
# 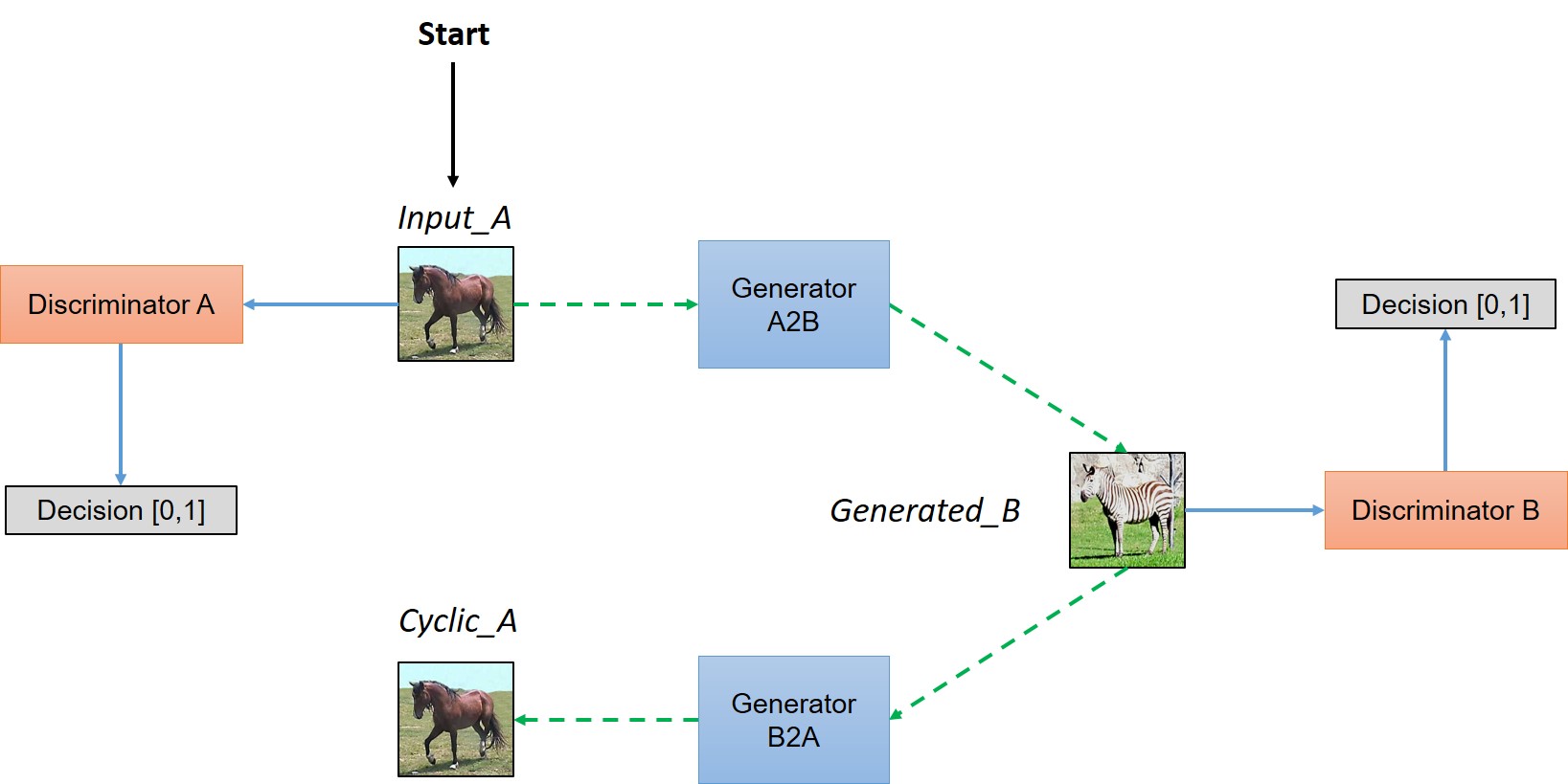
# ## Building Blocks
#export
class AutoTransConv(nn.Module):
"Automatic padding of transpose convolution for input-output feature size"
def __init__(self, n_in, n_out, ks, stride, bias=True):
super().__init__()
padding = ks // 2
self.conv = nn.ConvTranspose2d(n_in, n_out, ks, stride, padding=padding, output_padding=padding, bias=bias)
#export
def trans_conv_norm_relu(n_in, n_out, norm_layer, bias, ks=3, stride=2):
"Transpose convolutional layer"
return [AutoTransConv(n_in, n_out, ks=ks, stride=stride, bias=bias),
norm_layer(n_out),
nn.ReLU()
]
#export
def pad_conv_norm_relu(n_in, n_out, norm_layer, padding_mode="zeros", pad=1, ks=3, stride=1, activ=True, bias=True):
"Adding ability to specify different paddings to convolutional layer"
layers = []
if padding_mode != "zeros":
pad = 0
if padding_mode == "reflection": layers.append(nn.ReflectionPad2d(pad))
elif padding_mode == "border": layers.append(nn.ReplicationPad2d(pad))
layers.append(AutoConv(n_in, n_out, ks, stride=stride, padding_mode=padding_mode, bias=bias))
layers.append(norm_layer(n_out))
if activ: layers.append(nn.ReLU(inplace=True))
return layers
#export
def conv_norm_relu(n_in, n_out, norm_layer=None, ks=3, bias:bool=True, pad=1, stride=1, activ=True, a=0.2):
"Convolutional layer"
layers = []
layers.append(nn.Conv2d(n_in, n_out, ks, stride=stride, padding=pad))
if norm_layer != None: layers.append(norm_layer(n_out))
if activ: layers.append(nn.LeakyReLU(a, True))
return nn.Sequential(*layers)
#export
class ResBlock(nn.Module):
def __init__(self, dim, padding_mode, bias, dropout, norm_layer=nn.InstanceNorm2d):
"Residual connections for middle section of generator"
super().__init__()
layers = []
layers += pad_conv_norm_relu(dim, dim, norm_layer, padding_mode, bias=bias)
if dropout > 0: layers.append(nn.Dropout(dropout))
layers += (pad_conv_norm_relu(dim, dim, norm_layer, padding_mode, bias=bias, activ=False))
self.xb = nn.Sequential(*layers)
def forward(self, xb): return xb + self.conv(xb)
# ## Generator
#export
def generator(n_in, n_out, n_f=64, norm_layer=None, dropout=0., n_blocks=6, pad_mode="reflection"):
"Generator that maps an input of one domain to the other"
norm_layer = norm_layer if norm_layer is not None else nn.InstanceNorm2d
bias = (norm_layer == nn.InstanceNorm2d)
layers = []
layers += pad_conv_norm_relu(n_in, n_f, norm_layer, pad_mode, pad=3, ks=7, bias=bias)
for i in range(2):
layers += pad_conv_norm_relu(n_f, n_f*2, norm_layer, 'zeros', stride=2, bias=bias)
n_f*=2
layers += [ResBlock(n_f, pad_mode, bias, dropout, norm_layer) for _ in range(n_blocks)]
for i in range(2):
layers += trans_conv_norm_relu(n_f, n_f//2, norm_layer, bias=bias)
n_f //= 2
layers.append(nn.ReflectionPad2d(3))
layers.append(nn.Conv2d(n_f, n_out, kernel_size=7, padding=0))
layers.append(nn.Tanh())
return nn.Sequential(*layers)
generator(3,10)
# ## Discriminator
#export
def discriminator(c_in, n_f, n_layers, norm_layer=None, sigmoid=False):
"Discrminator to classify input as belonging to one class or the other"
norm_layer = nn.InstanceNorm2d if norm_layer is None else norm_layer
bias = (norm_layer == nn.InstanceNorm2d)
layers = []
layers += (conv_norm_relu(c_in, n_f, ks=4, stride=2, pad=1))
for i in range(n_layers-1):
new_f = 2*n_f if i <= 3 else n_f
layers += (conv_norm_relu(n_f, new_f, norm_layer, ks=4, stride=2, pad=1, bias=bias))
n_f = new_f
new_f = 2*n_f if n_layers <= 3 else n_f
layers += (conv_norm_relu(n_f, new_f, norm_layer, ks=4, stride=1, pad=1, bias=bias))
layers.append(nn.Conv2d(new_f, 1, kernel_size=4, stride=1, padding=1))
if sigmoid: layers.append(nn.Sigmoid())
return nn.Sequential(*layers)
discriminator(3, 6, 3)
# ## Loss and Trainer
#export
class cycleGAN(nn.Module):
def __init__(self, c_in, c_out, n_f=64, disc_layers=3, gen_blocks=6, drop=0., norm_layer=None, sigmoid=False):
super().__init__()
self.a_discriminator = discriminator(c_in, n_f, disc_layers, norm_layer, sigmoid)
self.b_discriminator = discriminator(c_in, n_f, disc_layers, norm_layer, sigmoid)
self.generate_a = generator(c_in, c_out, n_f, norm_layer, drop, gen_blocks)
self.generate_b = generator(c_in, c_out, n_f, norm_layer, drop, gen_blocks)
def forward(self, real_A, real_B):
generated_a, generated_B = self.generate_a(real_B), self.generate_b(real_A)
if not self.training: return generated_a, generated_b
id_a, id_b = self.generate_a(real_A), self.generate_b(real_B)
return [generated_a, generated_b, id_a, id_b]
#export
class DynamicLoss(nn.Module):
def __init__(self, loss_fn):
"Loss allowing for dynamic resizing of prediction based on shape of output"
super().__init__()
self.loss_fn = loss_fn
def forward(self, pred, targ, **kwargs):
targ = output.new_ones(*pred.shape) if targ == 1 else output.new_zeros(*pred.shape)
return self.loss_fn(pred, targ, **kwargs)
#export
class CycleGANLoss(nn.Module):
def __init__(self, model, loss_fn=F.mse_loss, la=10., lb=10, lid=0.5):
"CycleGAN loss"
super().__init__()
self.model,self.la,self.lb,self.lid = model,la,lb,lid
self.loss_fn = DynamicLoss(loss_fn)
def store_inputs(self, inputs):
self.reala,self.realb = inputs
def forward(self, pred, target):
gena, genb, ida, idb = pred
self.id_loss = self.lid * (self.la * F.l1_loss(ida, self.reala) + self.lb * F.l1_loss(idb,self.realb))
self.gen_loss = self.crit(self.model.a_discriminator(gena), True) + self.crit(self.model.b_discriminator(genb), True)
self.cyc_loss = self.la* F.l1_loss(self.model.generate_a(genb), self.reala) + self.lb*F.l1_loss(self.model.generate_b(gena), self.realb)
return self.id_loss+ self.gen_loss + self.cyc_loss
# CycleGAN loss is composed of 3 parts:
# 1. Identity, an image that has gone through its own domain should remain the same
# 2. Generator, the output images should fool the discriminator into thinking they belong to that class
# 3. Cyclical loss, and image that has been mapped to the other domain then mapped back to itself should resemble the original input
#export
class cycleGANTrainer(Callback):
"Trainer to sequence timing of training both the discriminator as well as the generator's"
_order = -20
def set_grad(self, da=False, db=False):
in_gen = (not da) and (not db)
requires_grad(self.learn.model.generate_a, in_gen)
requires_grad(self.learn.generate_b, in_gen)
requires_grad(self.learn.a_discriminator, da)
requires_grad(self.learn.b_discriminator, db)
if not gen:
self.opt_D_A.lr, self.opt_D_A.mom = self.learn.opt.lr, self.learn.opt.mom
self.opt_D_A.wd, self.opt_D_A.beta = self.learn.opt.wd, self.learn.opt.beta
self.opt_D_B.lr, self.opt_D_B.mom = self.learn.opt.lr, self.learn.opt.mom
self.opt_D_B.wd, self.opt_D_B.beta = self.learn.opt.wd, self.learn.opt.beta
def before_fit(self, **kwargs):
self.ga = self.learn.model.generate_a
self.gb = self.learn.generate_b
self.da = self.learn.a_discriminator
self.db = self.learn.b_discriminator
self.loss_fn = self.learn.loss_func.loss_func
if not getattr(self,'opt_gen',None):
self.opt_gen = self.learn.opt.new([nn.Sequential(*flatten_model(self.ga), *flatten_model(self.gb))])
else:
self.opt_gen.lr,self.opt_gen.wd = self.opt.lr,self.opt.wd
self.opt_gen.mom,self.opt_gen.beta = self.opt.mom,self.opt.beta
if not getattr(self,'opt_da',None):
self.opt_da = self.learn.opt.new([nn.Sequential(*flatten_model(self.da))])
if not getattr(self,'opt_db',None):
self.opt_db = self.learn.opt.new([nn.Sequential(*flatten_model(self.db))])
self.learn.opt.opt = self.opt_gen
self.set_grad()
def before_batch(self, last_input, **kwargs):
self.learn.loss_func.store_inputs(last_input)
def after_batch(self, last_input, last_output, **kwags):
#Discriminator loss
self.ga.zero_grad(), self.gb.zero_grad()
fakea, fakeb = last_output[0].detach(), last_output[1].detach()
reala,realb = last_input
self.set_grad(da=True)
self.da.zero_grad()
lossda = 0.5 * (self.loss_fn(self.da(reala), True) + self.loss_fn(self.da(fakea), False))
lossda.backward()
self.opt_da.step()
self.set_grad(db=True)
self.opt_db.zero_grad()
lossdb = 0.5 * (self.loss_fn(self.db(realb), True) + self.loss_fn(self.da(fakeb), False))
lossdb.backward()
self.opt_db.step()
self.set_grad()
# # Training
cgan = cycleGAN(3, 3, gen_blocks=9)
learn = get_learner(cgan, loss=CycleGANLoss(cgan))
run = get_runner(learn, [cycleGANTrainer()])
cgan
| cycleGAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pymedphys-master
# language: python
# name: pymedphys-master
# ---
import ast
import_with_from = """
from boo import hoo
"""
import_string = """
import attr
import dateutil
import dbfread
import keyring
import libjpeg
import packaging
import pymssql
import pynetdicom
import requests
import shapely
import shapely.affinity
import shapely.geometry
import shapely.ops
import streamlit
import timeago
import tkinter
import tkinter.filedialog
import toml
import tqdm
import watchdog
import watchdog.events
import watchdog.observers
import watchdog.observers.polling
import yaml
import numpy
import pandas
import scipy
import scipy.interpolate
import scipy.ndimage
import scipy.ndimage.measurements
import scipy.optimize
import scipy.signal
import scipy.special
import matplotlib
import matplotlib.patches
import matplotlib.path
import matplotlib.pyplot
import matplotlib.pyplot as plt
import matplotlib.transforms
import mpl_toolkits
import mpl_toolkits.mplot3d.art3d
import imageio
import PIL
import skimage
import skimage.draw
import skimage.measure
import pydicom
import pydicom.dataset
import pydicom.filebase
import pydicom.sequence
import pydicom.uid
"""
ast.Import
ast.alias
for node in ast.parse(import_with_from).body:
if not isinstance(node, ast.Import):
raise ValueError("Only direct import statements are supported")
print(ast.dump(node))
# +
imports_for_apipkg = {}
for node in ast.parse(import_string).body:
if not isinstance(node, ast.Import):
raise ValueError("Only direct import statements are supported")
aliased = list(node.names)
if len(aliased) != 1:
raise ValueError("Only one alias per import supported")
alias = aliased[0]
asname = alias.asname
if asname is None:
asname = alias.name
imports_for_apipkg[asname] = alias.name
imports_for_apipkg
# -
| prototyping/imports/import parsing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rMRWvtRMbmjt" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import h5py
import statistics,stats
import itertools
import time
# + id="BwaRHWiEcOXg" colab_type="code" colab={}
url='https://archive.ics.uci.edu/ml/machine-learning-databases/00279/SUSY.csv.gz'
# + id="P3fQp2WucQXr" colab_type="code" colab={}
coloumn_names=['Class_Label','lepton 1 pT', 'lepton 1 eta', 'lepton 1 phi', 'lepton 2 pT', 'lepton 2 eta', 'lepton 2 phi', 'missing energy magnitude', 'missing energy phi', 'MET_rel', 'axial MET',' M_R',
'M_TR_2', 'R', 'MT2', 'S_R', 'M_Delta_R', 'dPhi_r_b', 'cos(theta_r1)']
# + id="7VIMhqnucSjm" colab_type="code" colab={}
PM_DF=pd.read_csv(url,header=None,index_col=False,names=coloumn_names)
# + id="lhJVf8c3ONqu" colab_type="code" outputId="1eb9873c-5a62-4ed5-dfad-f58694f7e5fe" colab={"base_uri": "https://localhost:8080/", "height": 220}
PM_DF.head()
# + id="lK-yGguocd58" colab_type="code" colab={}
Features=PM_DF.drop('Class_Label',axis=1)
# + id="nhNS_hLDTfRp" colab_type="code" outputId="b709eb23-390f-4062-aeee-987c4b7ca7b3" colab={"base_uri": "https://localhost:8080/", "height": 220}
Features.head()
# + id="3wAcSKi7cg1_" colab_type="code" colab={}
Labels=PM_DF['Class_Label']
# + id="tc6YsZezTnc1" colab_type="code" outputId="a91fe523-9ae8-472d-b472-6fb6904bd05f" colab={"base_uri": "https://localhost:8080/", "height": 121}
Labels.head()
# + id="PxCBb9CqckFA" colab_type="code" outputId="4ac8664f-a58a-4411-8eeb-260a94ddb84e" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(Features.shape , Labels.shape)
# + id="xcU69qv3cmdD" colab_type="code" colab={}
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split,StratifiedKFold,cross_validate
from sklearn.metrics import confusion_matrix, accuracy_score, precision_score, recall_score, f1_score, roc_auc_score, make_scorer
# + id="SrdZ_dGtc-T0" colab_type="code" colab={}
x_train,x_test,y_train,y_test=train_test_split(Features,Labels,test_size=0.333,random_state=1,shuffle=True)
# + id="MtP5fRlXdGF5" colab_type="code" outputId="08a17bca-a137-41a0-d721-50ecced2f3ef" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(x_train.shape,x_test.shape,y_train.shape,y_test.shape)
# + id="wysle6BPdIaL" colab_type="code" outputId="f56edb6f-a882-4aa4-c8fc-83d598c2913c" colab={"base_uri": "https://localhost:8080/", "height": 136}
print(RandomForestClassifier())
# + id="brxuUqEAdO9Y" colab_type="code" outputId="77913e33-5476-473a-c682-5e06056b94b0" colab={"base_uri": "https://localhost:8080/", "height": 163}
RF_model=RandomForestClassifier(random_state=1,criterion='gini',n_estimators=100)
# + id="R-HcBXMgdbpv" colab_type="code" outputId="b9c83967-40ea-471a-a9e5-f2d5f10b8c7b" colab={"base_uri": "https://localhost:8080/", "height": 136}
RF_model.fit(x_train,y_train)
# + id="rciePORAdiDr" colab_type="code" colab={}
y_pred=RF_model.predict(x_test)
# + id="785TN8Mug4UA" colab_type="code" colab={}
def plot_confusion_matrix(cm, classes,
normalize=True,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
# + id="m2Sqb1wYhCGb" colab_type="code" colab={}
cnf_matrix = confusion_matrix(y_test, y_pred)
# + id="XZET4jwRhEKZ" colab_type="code" outputId="9730337f-76d6-4181-ac90-ddb79c0d11ba" colab={"base_uri": "https://localhost:8080/", "height": 347}
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['MBP','BN'],
title='Confusion matrix, without normalization')
# + id="UsrphRw0hGse" colab_type="code" outputId="391112b7-c8b1-471a-bdaf-098362709d60" colab={"base_uri": "https://localhost:8080/", "height": 85}
tn, fp, fn, tp = confusion_matrix(y_test, y_pred).ravel()
print("True Negatives: ",tn)
print("False Positives: ",fp)
print("False Negatives: ",fn)
print("True Positives: ",tp)
# + id="vMckiU8FhKME" colab_type="code" outputId="a82ca79d-e555-4cab-bd4f-94c1a9bc8965" colab={"base_uri": "https://localhost:8080/", "height": 85}
Accuracy = (tn+tp)*100/(tp+tn+fp+fn)
print("Accuracy {:0.2f}%:".format(Accuracy))
Precision = tp/(tp+fp)
print("Precision {:0.2f}".format(Precision))
Recall = tp/(tp+fn)
print("Recall {:0.2f}".format(Recall))
f1 = (2*Precision*Recall)/(Precision + Recall)
print("F1 Score {:0.2f}".format(f1))
# + id="qQd5YFwDhMoQ" colab_type="code" outputId="4f650847-e0f6-4865-c818-c8ca0559b8a6" colab={"base_uri": "https://localhost:8080/", "height": 34}
RF_model.score(x_test,y_test)
# + id="HfyvtrO7hP_E" colab_type="code" colab={}
F_imp=pd.Series(RF_model.feature_importances_,index=Features.columns)
# + id="qHhOhUXUhU6n" colab_type="code" colab={}
nfolds = 10
def tn(y_true, y_pred):
return confusion_matrix(y_true, y_pred)[0, 0]
def fp(y_true, y_pred):
return confusion_matrix(y_true, y_pred)[0, 1]
def fn(y_true, y_pred):
return confusion_matrix(y_true, y_pred)[1, 0]
def tp(y_true, y_pred):
return confusion_matrix(y_true, y_pred)[1, 1]
# + id="k9RaLV6hhaGL" colab_type="code" colab={}
scoring = {'tp': make_scorer(tp), 'tn': make_scorer(tn),
'fp': make_scorer(fp), 'fn': make_scorer(fn),
'ac' : make_scorer(accuracy_score),
're' : make_scorer(recall_score),
'pr' : make_scorer(precision_score),
'f1' : make_scorer(f1_score),
'auc' : make_scorer(roc_auc_score),
}
# + id="Z2EwohDDhcO4" colab_type="code" colab={}
cv_results = cross_validate(RF_model, x_train, y_train, scoring=scoring, cv=StratifiedKFold(n_splits=nfolds, random_state=1))
# + id="h-JpdGNZhiZt" colab_type="code" colab={}
print('Cross Validation scores (nfolds = %d):'% nfolds)
print('tp: ', cv_results['test_tp'], '; mean:', cv_results['test_tp'].mean())
print('fn: ', cv_results['test_fn'], '; mean:', cv_results['test_fn'].mean())
print('fp: ', cv_results['test_fp'], '; mean:', cv_results['test_fp'].mean())
print('tn: ', cv_results['test_tn'], '; mean:', cv_results['test_tn'].mean())
print('ac: ', cv_results['test_ac'], '; mean:', cv_results['test_ac'].mean())
print('re: ', cv_results['test_re'], '; mean:', cv_results['test_re'].mean())
print('pr: ', cv_results['test_pr'], '; mean:', cv_results['test_pr'].mean())
print('f1: ', cv_results['test_f1'], '; mean:', cv_results['test_f1'].mean())
print('auc: ', cv_results['test_auc'], '; mean:', cv_results['test_auc'].mean())
# + id="o5A8zWfnwjAR" colab_type="code" colab={}
def evaluate_model(model, features, labels):
pred = model.predict(features)
accuracy = round(accuracy_score(labels, pred), 3)
precision = round(precision_score(labels, pred), 3)
recall = round(recall_score(labels, pred), 3)
print('Accuracy: {} / Precision: {} / Recall: {}'.format(
accuracy,
precision,
recall
))
# + id="-Muan1hkwljk" colab_type="code" outputId="14cbc337-a7d6-4f45-82ac-7ab20a282ef1" colab={"base_uri": "https://localhost:8080/", "height": 163}
evaluate_model(RF_model,x_test,y_test)
# + id="dLPyXpU9wpx8" colab_type="code" colab={}
import sklearn.metrics as metrics
probs = RF_model.predict_proba(x_test)
# + id="bN-IqTNHyeQY" colab_type="code" colab={}
preds = probs[:,1]
# + id="JlZdsHPTyg0i" colab_type="code" colab={}
fpr, tpr, threshold = metrics.roc_curve(y_test, preds)
# + id="VrvXzQM-yi9e" colab_type="code" colab={}
roc_auc = metrics.auc(fpr, tpr)
# + id="SynZuMdEykr-" colab_type="code" colab={}
import matplotlib.pyplot as plt
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# + id="isg5H7Jfyp5Q" colab_type="code" colab={}
# + id="fQSoZCtPHO6X" colab_type="code" colab={}
# + id="k06m6YU7Hh9u" colab_type="code" colab={}
| Random_Forest_on_All_Attributes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# *Note: This is not yet ready, but shows the direction I'm leaning in for Fourth Edition Search.*
#
# # State-Space Search
#
# This notebook describes several state-space search algorithms, and how they can be used to solve a variety of problems. We start with a simple algorithm and a simple domain: finding a route from city to city. Later we will explore other algorithms and domains.
#
# ## The Route-Finding Domain
#
# Like all state-space search problems, in a route-finding problem you will be given:
# - A start state (for example, `'A'` for the city Arad).
# - A goal state (for example, `'B'` for the city Bucharest).
# - Actions that can change state (for example, driving from `'A'` to `'S'`).
#
# You will be asked to find:
# - A path from the start state, through intermediate states, to the goal state.
#
# We'll use this map:
#
# <img src="http://robotics.cs.tamu.edu/dshell/cs625/images/map.jpg" height="366" width="603">
#
# A state-space search problem can be represented by a *graph*, where the vertexes of the graph are the states of the problem (in this case, cities) and the edges of the graph are the actions (in this case, driving along a road).
#
# We'll represent a city by its single initial letter.
# We'll represent the graph of connections as a `dict` that maps each city to a list of the neighboring cities (connected by a road). For now we don't explicitly represent the actions, nor the distances
# between cities.
# + button=false new_sheet=false run_control={"read_only": false}
romania = {
'A': ['Z', 'T', 'S'],
'B': ['F', 'P', 'G', 'U'],
'C': ['D', 'R', 'P'],
'D': ['M', 'C'],
'E': ['H'],
'F': ['S', 'B'],
'G': ['B'],
'H': ['U', 'E'],
'I': ['N', 'V'],
'L': ['T', 'M'],
'M': ['L', 'D'],
'N': ['I'],
'O': ['Z', 'S'],
'P': ['R', 'C', 'B'],
'R': ['S', 'C', 'P'],
'S': ['A', 'O', 'F', 'R'],
'T': ['A', 'L'],
'U': ['B', 'V', 'H'],
'V': ['U', 'I'],
'Z': ['O', 'A']}
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Suppose we want to get from `A` to `B`. Where can we go from the start state, `A`?
# + button=false new_sheet=false run_control={"read_only": false}
romania['A']
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We see that from `A` we can get to any of the three cities `['Z', 'T', 'S']`. Which should we choose? *We don't know.* That's the whole point of *search*: we don't know which immediate action is best, so we'll have to explore, until we find a *path* that leads to the goal.
#
# How do we explore? We'll start with a simple algorithm that will get us from `A` to `B`. We'll keep a *frontier*—a collection of not-yet-explored states—and expand the frontier outward until it reaches the goal. To be more precise:
#
# - Initially, the only state in the frontier is the start state, `'A'`.
# - Until we reach the goal, or run out of states in the frontier to explore, do the following:
# - Remove the first state from the frontier. Call it `s`.
# - If `s` is the goal, we're done. Return the path to `s`.
# - Otherwise, consider all the neighboring states of `s`. For each one:
# - If we have not previously explored the state, add it to the end of the frontier.
# - Also keep track of the previous state that led to this new neighboring state; we'll need this to reconstruct the path to the goal, and to keep us from re-visiting previously explored states.
#
# # A Simple Search Algorithm: `breadth_first`
#
# The function `breadth_first` implements this strategy:
# + button=false new_sheet=false run_control={"read_only": false}
from collections import deque # Doubly-ended queue: pop from left, append to right.
def breadth_first(start, goal, neighbors):
"Find a shortest sequence of states from start to the goal."
frontier = deque([start]) # A queue of states
previous = {start: None} # start has no previous state; other states will
while frontier:
s = frontier.popleft()
if s == goal:
return path(previous, s)
for s2 in neighbors[s]:
if s2 not in previous:
frontier.append(s2)
previous[s2] = s
def path(previous, s):
"Return a list of states that lead to state s, according to the previous dict."
return [] if (s is None) else path(previous, previous[s]) + [s]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# A couple of things to note:
#
# 1. We always add new states to the end of the frontier queue. That means that all the states that are adjacent to the start state will come first in the queue, then all the states that are two steps away, then three steps, etc.
# That's what we mean by *breadth-first* search.
# 2. We recover the path to an `end` state by following the trail of `previous[end]` pointers, all the way back to `start`.
# The dict `previous` is a map of `{state: previous_state}`.
# 3. When we finally get an `s` that is the goal state, we know we have found a shortest path, because any other state in the queue must correspond to a path that is as long or longer.
# 3. Note that `previous` contains all the states that are currently in `frontier` as well as all the states that were in `frontier` in the past.
# 4. If no path to the goal is found, then `breadth_first` returns `None`. If a path is found, it returns the sequence of states on the path.
#
# Some examples:
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('A', 'B', romania)
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('L', 'N', romania)
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('N', 'L', romania)
# -
breadth_first('E', 'E', romania)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now let's try a different kind of problem that can be solved with the same search function.
#
# ## Word Ladders Problem
#
# A *word ladder* problem is this: given a start word and a goal word, find the shortest way to transform the start word into the goal word by changing one letter at a time, such that each change results in a word. For example starting with `green` we can reach `grass` in 7 steps:
#
# `green` → `greed` → `treed` → `trees` → `tress` → `cress` → `crass` → `grass`
#
# We will need a dictionary of words. We'll use 5-letter words from the [Stanford GraphBase](http://www-cs-faculty.stanford.edu/~uno/sgb.html) project for this purpose. Let's get that file from aimadata.
# + button=false new_sheet=false run_control={"read_only": false}
from search import *
sgb_words = open_data("EN-text/sgb-words.txt")
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We can assign `WORDS` to be the set of all the words in this file:
# + button=false new_sheet=false run_control={"read_only": false}
WORDS = set(sgb_words.read().split())
len(WORDS)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# And define `neighboring_words` to return the set of all words that are a one-letter change away from a given `word`:
# + button=false new_sheet=false run_control={"read_only": false}
def neighboring_words(word):
"All words that are one letter away from this word."
neighbors = {word[:i] + c + word[i+1:]
for i in range(len(word))
for c in 'abcdefghijklmnopqrstuvwxyz'
if c != word[i]}
return neighbors & WORDS
# -
# For example:
# + button=false new_sheet=false run_control={"read_only": false}
neighboring_words('hello')
# -
neighboring_words('world')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now we can create `word_neighbors` as a dict of `{word: {neighboring_word, ...}}`:
# + button=false new_sheet=false run_control={"read_only": false}
word_neighbors = {word: neighboring_words(word)
for word in WORDS}
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now the `breadth_first` function can be used to solve a word ladder problem:
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('green', 'grass', word_neighbors)
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('smart', 'brain', word_neighbors)
# + button=false new_sheet=false run_control={"read_only": false}
breadth_first('frown', 'smile', word_neighbors)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # More General Search Algorithms
#
# Now we'll embelish the `breadth_first` algorithm to make a family of search algorithms with more capabilities:
#
# 1. We distinguish between an *action* and the *result* of an action.
# 3. We allow different measures of the cost of a solution (not just the number of steps in the sequence).
# 4. We search through the state space in an order that is more likely to lead to an optimal solution quickly.
#
# Here's how we do these things:
#
# 1. Instead of having a graph of neighboring states, we instead have an object of type *Problem*. A Problem
# has one method, `Problem.actions(state)` to return a collection of the actions that are allowed in a state,
# and another method, `Problem.result(state, action)` that says what happens when you take an action.
# 2. We keep a set, `explored` of states that have already been explored. We also have a class, `Frontier`, that makes it efficient to ask if a state is on the frontier.
# 3. Each action has a cost associated with it (in fact, the cost can vary with both the state and the action).
# 4. The `Frontier` class acts as a priority queue, allowing the "best" state to be explored next.
# We represent a sequence of actions and resulting states as a linked list of `Node` objects.
#
# The algorithm `breadth_first_search` is basically the same as `breadth_first`, but using our new conventions:
# + button=false new_sheet=false run_control={"read_only": false}
def breadth_first_search(problem):
"Search for goal; paths with least number of steps first."
if problem.is_goal(problem.initial):
return Node(problem.initial)
frontier = FrontierQ(Node(problem.initial), LIFO=False)
explored = set()
while frontier:
node = frontier.pop()
explored.add(node.state)
for action in problem.actions(node.state):
child = node.child(problem, action)
if child.state not in explored and child.state not in frontier:
if problem.is_goal(child.state):
return child
frontier.add(child)
# -
# Next is `uniform_cost_search`, in which each step can have a different cost, and we still consider first one os the states with minimum cost so far.
# + button=false new_sheet=false run_control={"read_only": false}
def uniform_cost_search(problem, costfn=lambda node: node.path_cost):
frontier = FrontierPQ(Node(problem.initial), costfn)
explored = set()
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
explored.add(node.state)
for action in problem.actions(node.state):
child = node.child(problem, action)
if child.state not in explored and child not in frontier:
frontier.add(child)
elif child in frontier and frontier.cost[child] < child.path_cost:
frontier.replace(child)
# -
# Finally, `astar_search` in which the cost includes an estimate of the distance to the goal as well as the distance travelled so far.
# + button=false new_sheet=false run_control={"read_only": false}
def astar_search(problem, heuristic):
costfn = lambda node: node.path_cost + heuristic(node.state)
return uniform_cost_search(problem, costfn)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Search Tree Nodes
#
# The solution to a search problem is now a linked list of `Node`s, where each `Node`
# includes a `state` and the `path_cost` of getting to the state. In addition, for every `Node` except for the first (root) `Node`, there is a previous `Node` (indicating the state that lead to this `Node`) and an `action` (indicating the action taken to get here).
# + button=false new_sheet=false run_control={"read_only": false}
class Node(object):
"""A node in a search tree. A search tree is spanning tree over states.
A Node contains a state, the previous node in the tree, the action that
takes us from the previous state to this state, and the path cost to get to
this state. If a state is arrived at by two paths, then there are two nodes
with the same state."""
def __init__(self, state, previous=None, action=None, step_cost=1):
"Create a search tree Node, derived from a previous Node by an action."
self.state = state
self.previous = previous
self.action = action
self.path_cost = 0 if previous is None else (previous.path_cost + step_cost)
def __repr__(self): return "<Node {}: {}>".format(self.state, self.path_cost)
def __lt__(self, other): return self.path_cost < other.path_cost
def child(self, problem, action):
"The Node you get by taking an action from this Node."
result = problem.result(self.state, action)
return Node(result, self, action,
problem.step_cost(self.state, action, result))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Frontiers
#
# A frontier is a collection of Nodes that acts like both a Queue and a Set. A frontier, `f`, supports these operations:
#
# * `f.add(node)`: Add a node to the Frontier.
#
# * `f.pop()`: Remove and return the "best" node from the frontier.
#
# * `f.replace(node)`: add this node and remove a previous node with the same state.
#
# * `state in f`: Test if some node in the frontier has arrived at state.
#
# * `f[state]`: returns the node corresponding to this state in frontier.
#
# * `len(f)`: The number of Nodes in the frontier. When the frontier is empty, `f` is *false*.
#
# We provide two kinds of frontiers: One for "regular" queues, either first-in-first-out (for breadth-first search) or last-in-first-out (for depth-first search), and one for priority queues, where you can specify what cost function on nodes you are trying to minimize.
# + button=false new_sheet=false run_control={"read_only": false}
from collections import OrderedDict
import heapq
class FrontierQ(OrderedDict):
"A Frontier that supports FIFO or LIFO Queue ordering."
def __init__(self, initial, LIFO=False):
"""Initialize Frontier with an initial Node.
If LIFO is True, pop from the end first; otherwise from front first."""
super(FrontierQ, self).__init__()
self.LIFO = LIFO
self.add(initial)
def add(self, node):
"Add a node to the frontier."
self[node.state] = node
def pop(self):
"Remove and return the next Node in the frontier."
(state, node) = self.popitem(self.LIFO)
return node
def replace(self, node):
"Make this node replace the nold node with the same state."
del self[node.state]
self.add(node)
# + button=false new_sheet=false run_control={"read_only": false}
class FrontierPQ:
"A Frontier ordered by a cost function; a Priority Queue."
def __init__(self, initial, costfn=lambda node: node.path_cost):
"Initialize Frontier with an initial Node, and specify a cost function."
self.heap = []
self.states = {}
self.costfn = costfn
self.add(initial)
def add(self, node):
"Add node to the frontier."
cost = self.costfn(node)
heapq.heappush(self.heap, (cost, node))
self.states[node.state] = node
def pop(self):
"Remove and return the Node with minimum cost."
(cost, node) = heapq.heappop(self.heap)
self.states.pop(node.state, None) # remove state
return node
def replace(self, node):
"Make this node replace a previous node with the same state."
if node.state not in self:
raise ValueError('{} not there to replace'.format(node.state))
for (i, (cost, old_node)) in enumerate(self.heap):
if old_node.state == node.state:
self.heap[i] = (self.costfn(node), node)
heapq._siftdown(self.heap, 0, i)
return
def __contains__(self, state): return state in self.states
def __len__(self): return len(self.heap)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Search Problems
#
# `Problem` is the abstract class for all search problems. You can define your own class of problems as a subclass of `Problem`. You will need to override the `actions` and `result` method to describe how your problem works. You will also have to either override `is_goal` or pass a collection of goal states to the initialization method. If actions have different costs, you should override the `step_cost` method.
# + button=false new_sheet=false run_control={"read_only": false}
class Problem(object):
"""The abstract class for a search problem."""
def __init__(self, initial=None, goals=(), **additional_keywords):
"""Provide an initial state and optional goal states.
A subclass can have additional keyword arguments."""
self.initial = initial # The initial state of the problem.
self.goals = goals # A collection of possibe goal states.
self.__dict__.update(**additional_keywords)
def actions(self, state):
"Return a list of actions executable in this state."
raise NotImplementedError # Override this!
def result(self, state, action):
"The state that results from executing this action in this state."
raise NotImplementedError # Override this!
def is_goal(self, state):
"True if the state is a goal."
return state in self.goals # Optionally override this!
def step_cost(self, state, action, result=None):
"The cost of taking this action from this state."
return 1 # Override this if actions have different costs
# +
def action_sequence(node):
"The sequence of actions to get to this node."
actions = []
while node.previous:
actions.append(node.action)
node = node.previous
return actions[::-1]
def state_sequence(node):
"The sequence of states to get to this node."
states = [node.state]
while node.previous:
node = node.previous
states.append(node.state)
return states[::-1]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Two Location Vacuum World
# + button=false new_sheet=false run_control={"read_only": false}
dirt = '*'
clean = ' '
class TwoLocationVacuumProblem(Problem):
"""A Vacuum in a world with two locations, and dirt.
Each state is a tuple of (location, dirt_in_W, dirt_in_E)."""
def actions(self, state): return ('W', 'E', 'Suck')
def is_goal(self, state): return dirt not in state
def result(self, state, action):
"The state that results from executing this action in this state."
(loc, dirtW, dirtE) = state
if action == 'W': return ('W', dirtW, dirtE)
elif action == 'E': return ('E', dirtW, dirtE)
elif action == 'Suck' and loc == 'W': return (loc, clean, dirtE)
elif action == 'Suck' and loc == 'E': return (loc, dirtW, clean)
else: raise ValueError('unknown action: ' + action)
# + button=false new_sheet=false run_control={"read_only": false}
problem = TwoLocationVacuumProblem(initial=('W', dirt, dirt))
result = uniform_cost_search(problem)
result
# -
action_sequence(result)
state_sequence(result)
# + button=false new_sheet=false run_control={"read_only": false}
problem = TwoLocationVacuumProblem(initial=('E', clean, dirt))
result = uniform_cost_search(problem)
action_sequence(result)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Water Pouring Problem
#
# Here is another problem domain, to show you how to define one. The idea is that we have a number of water jugs and a water tap and the goal is to measure out a specific amount of water (in, say, ounces or liters). You can completely fill or empty a jug, but because the jugs don't have markings on them, you can't partially fill them with a specific amount. You can, however, pour one jug into another, stopping when the seconfd is full or the first is empty.
# + button=false new_sheet=false run_control={"read_only": false}
class PourProblem(Problem):
"""Problem about pouring water between jugs to achieve some water level.
Each state is a tuples of levels. In the initialization, provide a tuple of
capacities, e.g. PourProblem(capacities=(8, 16, 32), initial=(2, 4, 3), goals={7}),
which means three jugs of capacity 8, 16, 32, currently filled with 2, 4, 3 units of
water, respectively, and the goal is to get a level of 7 in any one of the jugs."""
def actions(self, state):
"""The actions executable in this state."""
jugs = range(len(state))
return ([('Fill', i) for i in jugs if state[i] != self.capacities[i]] +
[('Dump', i) for i in jugs if state[i] != 0] +
[('Pour', i, j) for i in jugs for j in jugs if i != j])
def result(self, state, action):
"""The state that results from executing this action in this state."""
result = list(state)
act, i, j = action[0], action[1], action[-1]
if act == 'Fill': # Fill i to capacity
result[i] = self.capacities[i]
elif act == 'Dump': # Empty i
result[i] = 0
elif act == 'Pour':
a, b = state[i], state[j]
result[i], result[j] = ((0, a + b)
if (a + b <= self.capacities[j]) else
(a + b - self.capacities[j], self.capacities[j]))
else:
raise ValueError('unknown action', action)
return tuple(result)
def is_goal(self, state):
"""True if any of the jugs has a level equal to one of the goal levels."""
return any(level in self.goals for level in state)
# + button=false new_sheet=false run_control={"read_only": false}
p7 = PourProblem(initial=(2, 0), capacities=(5, 13), goals={7})
p7.result((2, 0), ('Fill', 1))
# + button=false new_sheet=false run_control={"read_only": false}
result = uniform_cost_search(p7)
action_sequence(result)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Visualization Output
# + button=false new_sheet=false run_control={"read_only": false}
def showpath(searcher, problem):
"Show what happens when searcvher solves problem."
problem = Instrumented(problem)
print('\n{}:'.format(searcher.__name__))
result = searcher(problem)
if result:
actions = action_sequence(result)
state = problem.initial
path_cost = 0
for steps, action in enumerate(actions, 1):
path_cost += problem.step_cost(state, action, 0)
result = problem.result(state, action)
print(' {} =={}==> {}; cost {} after {} steps'
.format(state, action, result, path_cost, steps,
'; GOAL!' if problem.is_goal(result) else ''))
state = result
msg = 'GOAL FOUND' if result else 'no solution'
print('{} after {} results and {} goal checks'
.format(msg, problem._counter['result'], problem._counter['is_goal']))
from collections import Counter
class Instrumented:
"Instrument an object to count all the attribute accesses in _counter."
def __init__(self, obj):
self._object = obj
self._counter = Counter()
def __getattr__(self, attr):
self._counter[attr] += 1
return getattr(self._object, attr)
# + button=false new_sheet=false run_control={"read_only": false}
showpath(uniform_cost_search, p7)
# -
p = PourProblem(initial=(0, 0), capacities=(7, 13), goals={2})
showpath(uniform_cost_search, p)
class GreenPourProblem(PourProblem):
def step_cost(self, state, action, result=None):
"The cost is the amount of water used in a fill."
if action[0] == 'Fill':
i = action[1]
return self.capacities[i] - state[i]
return 0
p = GreenPourProblem(initial=(0, 0), capacities=(7, 13), goals={2})
showpath(uniform_cost_search, p)
# + button=false new_sheet=false run_control={"read_only": false}
def compare_searchers(problem, searchers=None):
"Apply each of the search algorithms to the problem, and show results"
if searchers is None:
searchers = (breadth_first_search, uniform_cost_search)
for searcher in searchers:
showpath(searcher, problem)
# + button=false new_sheet=false run_control={"read_only": false}
compare_searchers(p)
# -
# # Random Grid
#
# An environment where you can move in any of 4 directions, unless there is an obstacle there.
#
#
#
#
# +
import random
N, S, E, W = DIRECTIONS = [(0, 1), (0, -1), (1, 0), (-1, 0)]
def Grid(width, height, obstacles=0.1):
"""A 2-D grid, width x height, with obstacles that are either a collection of points,
or a fraction between 0 and 1 indicating the density of obstacles, chosen at random."""
grid = {(x, y) for x in range(width) for y in range(height)}
if isinstance(obstacles, (float, int)):
obstacles = random.sample(grid, int(width * height * obstacles))
def neighbors(x, y):
for (dx, dy) in DIRECTIONS:
(nx, ny) = (x + dx, y + dy)
if (nx, ny) not in obstacles and 0 <= nx < width and 0 <= ny < height:
yield (nx, ny)
return {(x, y): list(neighbors(x, y))
for x in range(width) for y in range(height)}
Grid(5, 5)
# -
class GridProblem(Problem):
"Create with a call like GridProblem(grid=Grid(10, 10), initial=(0, 0), goal=(9, 9))"
def actions(self, state): return DIRECTIONS
def result(self, state, action):
#print('ask for result of', state, action)
(x, y) = state
(dx, dy) = action
r = (x + dx, y + dy)
return r if r in self.grid[state] else state
gp = GridProblem(grid=Grid(5, 5, 0.3), initial=(0, 0), goals={(4, 4)})
showpath(uniform_cost_search, gp)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Finding a hard PourProblem
#
# What solvable two-jug PourProblem requires the most steps? We can define the hardness as the number of steps, and then iterate over all PourProblems with capacities up to size M, keeping the hardest one.
# + button=false new_sheet=false run_control={"read_only": false}
def hardness(problem):
L = breadth_first_search(problem)
#print('hardness', problem.initial, problem.capacities, problem.goals, L)
return len(action_sequence(L)) if (L is not None) else 0
# + button=false new_sheet=false run_control={"read_only": false}
hardness(p7)
# -
action_sequence(breadth_first_search(p7))
# + button=false new_sheet=false run_control={"read_only": false}
C = 9 # Maximum capacity to consider
phard = max((PourProblem(initial=(a, b), capacities=(A, B), goals={goal})
for A in range(C+1) for B in range(C+1)
for a in range(A) for b in range(B)
for goal in range(max(A, B))),
key=hardness)
phard.initial, phard.capacities, phard.goals
# -
showpath(breadth_first_search, PourProblem(initial=(0, 0), capacities=(7, 9), goals={8}))
# + button=false new_sheet=false run_control={"read_only": false}
showpath(uniform_cost_search, phard)
# + button=false new_sheet=false run_control={"read_only": false}
class GridProblem(Problem):
"""A Grid."""
def actions(self, state): return ['N', 'S', 'E', 'W']
def result(self, state, action):
"""The state that results from executing this action in this state."""
(W, H) = self.size
if action == 'N' and state > W: return state - W
if action == 'S' and state + W < W * W: return state + W
if action == 'E' and (state + 1) % W !=0: return state + 1
if action == 'W' and state % W != 0: return state - 1
return state
# + button=false new_sheet=false run_control={"read_only": false}
compare_searchers(GridProblem(initial=0, goals={44}, size=(10, 10)))
# + button=false new_sheet=false run_control={"read_only": false}
def test_frontier():
#### Breadth-first search with FIFO Q
f = FrontierQ(Node(1), LIFO=False)
assert 1 in f and len(f) == 1
f.add(Node(2))
f.add(Node(3))
assert 1 in f and 2 in f and 3 in f and len(f) == 3
assert f.pop().state == 1
assert 1 not in f and 2 in f and 3 in f and len(f) == 2
assert f
assert f.pop().state == 2
assert f.pop().state == 3
assert not f
#### Depth-first search with LIFO Q
f = FrontierQ(Node('a'), LIFO=True)
for s in 'bcdef': f.add(Node(s))
assert len(f) == 6 and 'a' in f and 'c' in f and 'f' in f
for s in 'fedcba': assert f.pop().state == s
assert not f
#### Best-first search with Priority Q
f = FrontierPQ(Node(''), lambda node: len(node.state))
assert '' in f and len(f) == 1 and f
for s in ['book', 'boo', 'bookie', 'bookies', 'cook', 'look', 'b']:
assert s not in f
f.add(Node(s))
assert s in f
assert f.pop().state == ''
assert f.pop().state == 'b'
assert f.pop().state == 'boo'
assert {f.pop().state for _ in '123'} == {'book', 'cook', 'look'}
assert f.pop().state == 'bookie'
#### Romania: Two paths to Bucharest; cheapest one found first
S = Node('S')
SF = Node('F', S, 'S->F', 99)
SFB = Node('B', SF, 'F->B', 211)
SR = Node('R', S, 'S->R', 80)
SRP = Node('P', SR, 'R->P', 97)
SRPB = Node('B', SRP, 'P->B', 101)
f = FrontierPQ(S)
f.add(SF); f.add(SR), f.add(SRP), f.add(SRPB); f.add(SFB)
def cs(n): return (n.path_cost, n.state) # cs: cost and state
assert cs(f.pop()) == (0, 'S')
assert cs(f.pop()) == (80, 'R')
assert cs(f.pop()) == (99, 'F')
assert cs(f.pop()) == (177, 'P')
assert cs(f.pop()) == (278, 'B')
return 'test_frontier ok'
test_frontier()
# + button=false new_sheet=false run_control={"read_only": false}
# # %matplotlib inline
import matplotlib.pyplot as plt
p = plt.plot([i**2 for i in range(10)])
plt.savefig('destination_path.eps', format='eps', dpi=1200)
# + button=false new_sheet=false run_control={"read_only": false}
import itertools
import random
# http://stackoverflow.com/questions/10194482/custom-matplotlib-plot-chess-board-like-table-with-colored-cells
from matplotlib.table import Table
def main():
grid_table(8, 8)
plt.axis('scaled')
plt.show()
def grid_table(nrows, ncols):
fig, ax = plt.subplots()
ax.set_axis_off()
colors = ['white', 'lightgrey', 'dimgrey']
tb = Table(ax, bbox=[0,0,2,2])
for i,j in itertools.product(range(ncols), range(nrows)):
tb.add_cell(i, j, 2./ncols, 2./nrows, text='{:0.2f}'.format(0.1234),
loc='center', facecolor=random.choice(colors), edgecolor='grey') # facecolors=
ax.add_table(tb)
#ax.plot([0, .3], [.2, .2])
#ax.add_line(plt.Line2D([0.3, 0.5], [0.7, 0.7], linewidth=2, color='blue'))
return fig
main()
# -
import collections
class defaultkeydict(collections.defaultdict):
"""Like defaultdict, but the default_factory is a function of the key.
>>> d = defaultkeydict(abs); d[-42]
42
"""
def __missing__(self, key):
self[key] = self.default_factory(key)
return self[key]
# # Simulated Annealing visualisation using TSP
#
# Applying simulated annealing in traveling salesman problem to find the shortest tour to travel all cities in Romania. Distance between two cities is taken as the euclidean distance.
class TSP_problem(Problem):
'''
subclass of Problem to define various functions
'''
def two_opt(self, state):
'''
Neighbour generating function for Traveling Salesman Problem
'''
state2 = state[:]
l = random.randint(0, len(state2) - 1)
r = random.randint(0, len(state2) - 1)
if l > r:
l, r = r,l
state2[l : r + 1] = reversed(state2[l : r + 1])
return state2
def actions(self, state):
'''
action that can be excuted in given state
'''
return [self.two_opt]
def result(self, state, action):
'''
result after applying the given action on the given state
'''
return action(state)
def path_cost(self, c, state1, action, state2):
'''
total distance for the Traveling Salesman to be covered if in state2
'''
cost = 0
for i in range(len(state2) - 1):
cost += distances[state2[i]][state2[i + 1]]
cost += distances[state2[0]][state2[-1]]
return cost
def value(self, state):
'''
value of path cost given negative for the given state
'''
return -1 * self.path_cost(None, None, None, state)
# +
def init():
'''
Initialisation function for matplotlib animation
'''
line.set_data([], [])
for name, coordinates in romania_map.locations.items():
ax.annotate(
name,
xy=coordinates, xytext=(-10, 5), textcoords='offset points', size = 10)
text.set_text("Cost = 0 i = 0" )
return line,
def animate(i):
'''
Animation function to set next path and print its cost.
'''
x, y = [], []
for name in states[i]:
x.append(romania_map.locations[name][0])
y.append(romania_map.locations[name][1])
x.append(romania_map.locations[states[i][0]][0])
y.append(romania_map.locations[states[i][0]][1])
line.set_data(x,y)
text.set_text("Cost = " + str('{:.2f}'.format(TSP_problem.path_cost(None, None, None, None, states[i]))))
return line,
# +
# %matplotlib notebook
import matplotlib.pyplot as plt
from matplotlib import animation
import numpy as np
font = {'family': 'roboto',
'color': 'darkred',
'weight': 'normal',
'size': 12,
}
cities = []
distances ={}
states = []
# creating plotting area
fig = plt.figure(figsize = (8,6))
ax = plt.axes(xlim=(60, 600), ylim=(245, 600))
line, = ax.plot([], [], c="b",linewidth = 1.5, marker = 'o', markerfacecolor = 'r', markeredgecolor = 'r',markersize = 10)
text = ax.text(450, 565, "", fontdict = font)
# creating initial path
for name in romania_map.locations.keys():
distances[name] = {}
cities.append(name)
# distances['city1']['city2'] contains euclidean distance between their coordinates
for name_1,coordinates_1 in romania_map.locations.items():
for name_2,coordinates_2 in romania_map.locations.items():
distances[name_1][name_2] = np.linalg.norm([coordinates_1[0] - coordinates_2[0], coordinates_1[1] - coordinates_2[1]])
distances[name_2][name_1] = np.linalg.norm([coordinates_1[0] - coordinates_2[0], coordinates_1[1] - coordinates_2[1]])
# creating the problem
tsp_problem = TSP_problem(cities)
# all the states as a 2-D list of paths
states = simulated_annealing_full(tsp_problem)
# calling the matplotlib animation function
anim = animation.FuncAnimation(fig, animate, init_func = init,
frames = len(states), interval = len(states), blit = True, repeat = False)
plt.show()
# -
# ### Iterative Simulated Annealing
#
# Providing the output of the previous run as input to the next run to give better performance.
# +
next_state = cities
states = []
# creating plotting area
fig = plt.figure(figsize = (8,6))
ax = plt.axes(xlim=(60, 600), ylim=(245, 600))
line, = ax.plot([], [], c="b",linewidth = 1.5, marker = 'o', markerfacecolor = 'r', markeredgecolor = 'r',markersize = 10)
text = ax.text(450, 565, "", fontdict = font)
# to plot only the final states of every simulated annealing iteration
for iterations in range(100):
tsp_problem = TSP_problem(next_state)
states.append(simulated_annealing(tsp_problem))
next_state = states[-1]
anim = animation.FuncAnimation(fig, animate, init_func=init,
frames=len(states),interval=len(states), blit=True, repeat = False)
plt.show()
# -
| search-4e.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python3
# name: python3
# ---
# The Marginal Value of Adaptive Gradient Methods in Machine Learning
# https://arxiv.org/abs/1705.08292
#
# Asynchronous Stochastic Gradient Descent with Delay Compensation for Distributed Deep Learning
# https://arxiv.org/abs/1609.08326
#
# Asynchronous Stochastic Gradient Descent with Variance Reduction for Non-Convex Optimization
# https://arxiv.org/abs/1604.03584
#
# Large Scale Distributed Deep Networks
# https://static.googleusercontent.com/media/research.google.com/en//archive/large_deep_networks_nips2012.pdf
#
# Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift
# <NAME>, <NAME>
# https://arxiv.org/abs/1502.03167
#
# Xavier (Glorot) Normal Initializer
# http://jmlr.org/proceedings/papers/v9/glorot10a/glorot10a.pdf
#
# He Normal Initializer
# http://arxiv.org/abs/1502.01852
#
# For understanding Nesterov Momentum:
# Advances in optimizing Recurrent Networks by <NAME>, Section 3.5
# http://arxiv.org/pdf/1212.0901v2.pdf
| Deep_Learning/archived/Appendix - Extra Reading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: synchro
# language: python
# name: synchro
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Synchro-Project" data-toc-modified-id="Synchro-Project-1">Synchro Project</a></span><ul class="toc-item"><li><span><a href="#TOC" data-toc-modified-id="TOC-1.1">TOC</a></span></li><li><span><a href="#1.-setup" data-toc-modified-id="1.-setup-1.2">1. setup</a></span></li><li><span><a href="#2.-get-data" data-toc-modified-id="2.-get-data-1.3">2. get data</a></span><ul class="toc-item"><li><span><a href="#2.A.-nan-remover" data-toc-modified-id="2.A.-nan-remover-1.3.1">2.A. nan remover</a></span></li><li><span><a href="#2.B.-add-features" data-toc-modified-id="2.B.-add-features-1.3.2">2.B. add features</a></span></li><li><span><a href="#2.C.-pandas-report" data-toc-modified-id="2.C.-pandas-report-1.3.3">2.C. pandas report</a></span></li><li><span><a href="#2.D.-save-data" data-toc-modified-id="2.D.-save-data-1.3.4">2.D. save data</a></span></li><li><span><a href="#2.E.-split-data" data-toc-modified-id="2.E.-split-data-1.3.5">2.E. split data</a></span></li></ul></li><li><span><a href="#3.-Pipeline" data-toc-modified-id="3.-Pipeline-1.4">3. Pipeline</a></span></li><li><span><a href="#4.-optimization" data-toc-modified-id="4.-optimization-1.5">4. optimization</a></span></li><li><span><a href="#5.-feature-importance" data-toc-modified-id="5.-feature-importance-1.6">5. feature importance</a></span></li><li><span><a href="#6.-diagnostic" data-toc-modified-id="6.-diagnostic-1.7">6. diagnostic</a></span></li><li><span><a href="#7.-evaluation" data-toc-modified-id="7.-evaluation-1.8">7. evaluation</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Hotelling" data-toc-modified-id="Hotelling-1.8.0.1">Hotelling</a></span></li></ul></li></ul></li></ul></li></ul></div>
# -
# # Synchro Project
# - [github link](https://github.com/romainmartinez/envergo)
# ## TOC
# 1. setup
# 2. get data
# 1. nan remover
# 2. add features
# 3. pandas report
# 4. save data
# 5. split data
# 3. pipeline
# 4. optimization
# 5. features importance
# 6. diagnostic
# 7. evaluation
# ## 1. setup
# +
# Common imports
import scipy.io as sio
import pandas as pd
import numpy as np
import os
# Figures
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
sns.set_context("notebook", font_scale=1.1)
sns.set_style("ticks")
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42
# to make this notebook's output stable across runs
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
# -
# ## 2. get data
# +
# load from mat
DATA_PATH = './data/'
X_import = sio.loadmat(os.path.join(DATA_PATH, 'X.mat'))['TableauForces']
y_import = sio.loadmat(os.path.join(DATA_PATH, 'y.mat'))['TestData']
X_description = np.array(['AddL', 'AddR', 'AbdL', 'AbdR', 'ErL', 'ErR', 'IrL', 'IrR', 'ExtL', 'ExtR', 'FlexL', 'FlexR'])
y_description = np.array(['Dyn', 'BodyBoost', 'MeanEggBeater', 'MaxEggBeater'])
# -
# ### 2.A. nan remover
nan_id = np.argwhere(np.isnan(X_import))
n_nans = np.sum(np.isnan(X_import).sum(axis=1))
for i in nan_id:
print(f'\tparticipant n: {i[0]}')
print(f'\ttest: {X_description[i[1]]}')
# if left take right, left otherwise
if X_description[i[1]][-1] == 'L':
replacer = i[1] + 1
elif X_description[i[1]][-1] == 'R':
replacer = i[1] - 1
print(f'\t\t"{X_import[i[0], i[1]]}" replace by "{X_import[i[0], replacer]}"')
X_import[i[0], i[1]] = X_import[i[0], replacer]
print('\t', '-' * 5)
# ### 2.B. add features
# load height + weight
anthropo = sio.loadmat(os.path.join(DATA_PATH, 'heightweight.mat'))['HeightWeight']
# replace nan
from sklearn.preprocessing import Imputer
anthropo = Imputer(strategy='median').fit_transform(anthropo)
# add IMC
anthropo = np.c_[anthropo, anthropo[:, 1] / (anthropo[:, 0])**2]
# compute imbalance
imbalance = None
for i in range(0, X_import.shape[1], 2):
if imbalance is None:
imbalance = np.abs((X_import[:, i] - X_import[:, i + 1]) / X_import[:, i]) * 100
else:
imbalance = np.c_[imbalance, np.abs((X_import[:, i] - X_import[:, i + 1]) / X_import[:, i]) * 100]
imbalance = np.mean(imbalance, axis=1)
X_mat = np.c_[X_import, anthropo, imbalance]
X_description = np.append(X_description, ['height', 'weight', 'IMC', 'imbalance'])
df = pd.DataFrame(
data=np.c_[X_mat, y_import],
columns=np.append(X_description, y_description)
)
X_cols = {
'test': np.arange(12),
'height': np.array([12]),
'weight': np.array([13]),
'IMC': np.array([14]),
'imbalance': np.array([15])
}
# ### 2.C. pandas report
REPORT_FILENAME = './pandas_report.html'
if not os.path.isfile(REPORT_FILENAME):
import pandas_profiling
report = pandas_profiling.ProfileReport(df)
report.to_file('./pandas_report.html')
# ### 2.D. save data
FILENAME = './data/dataframe.hdf5'
df.to_hdf(FILENAME, 'SYNCHRO', format='table')
# ### 2.E. split data
# +
# split data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_mat, y_import, test_size=0.2, random_state=RANDOM_SEED)
# -
# ## 3. Pipeline
# +
# custom class
from sklearn.base import BaseEstimator, TransformerMixin
class Normalize(BaseEstimator, TransformerMixin):
"""Normalize a given array with weight, height or IMC"""
def __init__(self, X_cols=X_cols, strategy='IMC'):
self.strategy = strategy
self.X_cols = X_cols
def fit(self, X, y=None):
return self
def transform(self, X):
X_copy = X.copy()
if self.strategy == 'height':
normalizer = X_copy[:, X_cols['height']].reshape(-1, 1)
elif self.strategy == 'weight':
normalizer = X_copy[:, X_cols['weight']].reshape(-1, 1)
elif self.strategy == 'IMC':
normalizer = X_copy[:, X_cols['IMC']].reshape(-1, 1)
else:
raise ValueError(f'please choose one of the following: height, weight, IMC. You have chosen {self.strategy}')
X_copy[:, X_cols['test']] = X_copy[:, X_cols['test']] / normalizer
return X_copy
class TestSide(BaseEstimator, TransformerMixin):
"""Return the mean between left & right or both"""
def __init__(self, X_cols=X_cols, strategy='mean'):
self.strategy = strategy
self.X_cols = X_cols
def fit(self, X, y=None):
return self
def transform(self, X):
if self.strategy is 'mean' or 'Fscore':
output = X.copy()
for i in range(X_cols['test'][-1] + 1):
if i % 2 == 0:
if self.strategy is 'mean':
output[:, i] = np.mean([X[:, i], X[:, i + 1]], axis=0)
else:
output[:, i] = 2 * (X[:, i] * X[:, i + 1]) / (X[:, i] + X[:, i + 1])
else:
output[:, i] = np.nan
else:
raise ValueError('please choose one of the following: mean, Fscore')
return output
class FeaturesAdder(BaseEstimator, TransformerMixin):
"""Add features based on the list `new_features`
Possible `new_features` are: IMC, imbalance
"""
def __init__(self, X_cols=X_cols, new_features='None'):
self.new_features = new_features
self.X_cols = X_cols
def fit(self, X, y=None):
return self
def transform(self, X):
X_copy = X.copy()
for key, cols in X_cols.items():
if not key in self.new_features:
X_copy[:, cols] = np.nan
return X_copy[:, ~np.all(np.isnan(X_copy), axis=0)] # remove nan columns
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PolynomialFeatures
preprocessing = make_pipeline(
Normalize(strategy='IMC'),
TestSide(strategy='Fscore'),
FeaturesAdder(new_features=['test', 'height', 'weight', 'IMC', 'imbalance']),
PolynomialFeatures(degree=1)
)
# +
from sklearn.multioutput import MultiOutputRegressor
from xgboost import XGBRegressor
full_pipeline = make_pipeline(
preprocessing,
MultiOutputRegressor(XGBRegressor())
)
# -
# ## 4. optimization
stuff = ['test', 'height', 'weight', 'IMC', 'imbalance']
# +
import itertools
all_possible_features = ['test', 'height', 'weight', 'IMC', 'imbalance']
all_features_combinations = []
for L in range(1, len(all_possible_features)+1):
for subset in itertools.combinations(all_possible_features, L):
all_features_combinations.append(list(subset))
all_features_combinations
# -
xgb_param = {
'multioutputregressor__estimator__max_depth': [3, 5, 7, 9, 12, 15, 17, 25],
'multioutputregressor__estimator__learning_rate': [0.01, 0.015, 0.025, 0.05, 0.1],
'multioutputregressor__estimator__n_estimators': [10, 25, 50, 150, 200],
'multioutputregressor__estimator__gamma': [0.05, 0.1, 0.3, 0.5, 0.7, 0.9, 0.10],
'multioutputregressor__estimator__min_child_weight': [1, 3, 5, 7],
'multioutputregressor__estimator__subsample': [0.6, 0.7, 0.8, 0.9, 1.0],
'multioutputregressor__estimator__colsample_bytree': [0.6, 0.7, 0.8, 0.9, 1.0],
'multioutputregressor__estimator__reg_lambda': [0.01, 0.1, 1.0],
'multioutputregressor__estimator__reg_alpha': [0, 0.1, 0.5, 1.0]
}
# +
from sklearn.model_selection import GridSearchCV
from sklearn.externals import joblib
optimize = True
MODEL_FILENAME = 'xgboost_model.pkl'
if optimize:
param_grid = [{
'pipeline__normalize': [None, Normalize(strategy='height'), Normalize('weight'), Normalize('IMC')],
'pipeline__testside': [None, TestSide(strategy='mean'), TestSide(strategy='Fscore')],
'pipeline__featuresadder__new_features': all_features_combinations,
'pipeline__polynomialfeatures': [None],
}]
grid_search_prep = GridSearchCV(full_pipeline, param_grid, cv=5, scoring='neg_mean_squared_error',
verbose=1)
grid_search_prep.fit(X_train, y_train)
model = grid_search_prep
joblib.dump(model, MODEL_FILENAME)
else:
model = joblib.load(MODEL_FILENAME)
# -
model.best_score_
model.best_params_
from sklearn.base import clone
xi = clone(full_pipeline)
xi.set_params(**model.best_params_)
xi.fit(X_train, y_train.ravel())
# ---
grid_search_prep.best_params_
grid_search_prep.best_score_
# without optimization: -0.97362502907769455
# ## 5. feature importance
regressors = model.best_estimator_.named_steps["multioutputregressor"]
for ilabel in range(y_test.shape[1]):
fscore = regressors.estimators_[ilabel].booster().get_fscore()
if ilabel is 0:
importance = pd.DataFrame(data={y_description[ilabel]: list(fscore.values())},
index=list(fscore.keys()))
else:
temp = pd.DataFrame(data={y_description[ilabel]: list(fscore.values())},
index=list(fscore.keys()))
importance = importance.join(temp)
importance.sort_index(inplace=True)
relative_importance = (importance.div(importance.sum(axis=1), axis=0)) * 100
relative_importance['features'] = relative_importance.index
# to tidy
relative_importance = pd.melt(relative_importance, id_vars='features',
var_name='label', value_name='importance')
selected_features = ['Add', 'Abd', 'Er', 'Ir', 'Ext', 'Flex', 'weight']
g = sns.factorplot(data=relative_importance, x='features', y='importance', col='label', kind='bar',
saturation=.5)
g.set_xticklabels(selected_features)
sns.despine(offset=10, trim=True)
plt.show()
# ## 6. diagnostic
# +
from sklearn.model_selection import learning_curve
def plot_learning_curve(estimator, X, y, scoring, cv=None, title=None, ylim=None,
n_jobs=1, train_sizes=np.linspace(.1, 1.0, 5)):
plt.figure()
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(estimator, X, y, cv=cv, scoring=scoring,
n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
return train_scores
# -
a = full_pipeline
a.set_params(**grid_search_prep.best_params_)
# +
from sklearn.model_selection import learning_curve
title = 'coucou'
scoring = 'neg_mean_squared_error'
t = plot_learning_curve(a, X_train, y_train, scoring, cv=5)
plt.show()
# -
train_sizes, train_scores, test_scores = learning_curve(grid_search_prep.best_estimator_, X_train, y_train, cv=5,
scoring='neg_mean_squared_error')
# The complex pipeline is certainly not suffering from high bias, as that would imply a higher error on the training set. From the gap between training and test error it rather seems like the model may exhibit too much variance, i.e. overfitting on the training folds. This makes sense both because our model is rather complex, and also because the size of the whole training data is relatively small (less than 8000 documents, compare that to the number of features produced by the tf-df, which can run into several tens of thousands without dimensionality reduction). Collection of more data would thus be one way to try and improve performance here (and it might also be useful to investigate different forms of regularization to avoid overfitting. Interestingly though, grid-search of the logistic regression led to best results without regularization). On the other hand, test error does not seem to be decreasing much with increasing size of the training set, indicating perhaps some inherent unpredictability in the data (some comments in the forum e.g. indicate that the class labels seem to have been assigned somewhat inconsistently).
# ## 7. evaluation
y_pred = grid_search_prep.best_estimator_.predict(X_test)
# +
from sklearn.metrics import mean_squared_error
import spm1d
def mape(y_test, y_pred):
val = (np.abs((y_test - y_pred) / y_test)) * 100
return np.mean(val), np.std(val)
def mse(y_test, y_pred):
val = ((y_test - y_pred) ** 2)
return np.mean(val), np.std(val)
def rmse(y_test, y_pred):
val = np.sqrt(((y_test - y_pred) ** 2))
return np.mean(val), np.std(val)
def t_test(y_test, y_pred, alpha=0.05, iterations=10000):
spm = spm1d.stats.nonparam.ttest2(y_test, y_pred)
spmi = spm.inference(alpha, two_tailed=True,
iterations=iterations, force_iterations=True)
if spmi.h0reject:
output = f'null hypothesis rejected, T2={spmi.z:.3f}, p={spmi.p:.5f}'
else:
output = f'null hypothesis not rejected (p={spmi.p:.5f})'
print(f'\tt-test = {output}')
def evaluate(y_test, y_pred, y_description):
for i in range(y_pred.shape[1]):
print(y_description[i])
mse_mu, mse_std = mse(y_test[:, i], y_pred[:, i])
rmse_mu, rmse_std = rmse(y_test[:, i], y_pred[:, i])
mape_mu, mape_std = mape(y_test[:, i], y_pred[:, i])
print(f'\tmse = {mse_mu:.3f} ({mse_std:.3f})')
print(f'\trmse = {rmse_mu:.3f} ({rmse_std:.3f})')
print(f'\tmape = {mape_mu:.3f}% ({mape_std:.3f})')
t_test(y_test[:, i], y_pred[:, i])
print('-' * 10)
# -
evaluate(y_test, y_pred, y_description)
# #### Hotelling
# +
t_spm = spm1d.stats.nonparam.hotellings2(y_test, y_pred)
t_spmi = t_spm.inference(alpha=0.05, iterations=10000)
print(t_spmi)
| sandbox/0_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import os
from config import population_frame
from initialise_parameters import params,categories
import numpy as np
import pandas as pd
from math import floor
from tqdm import tqdm
import seaborn as sns
cwd = os.getcwd()
sols = pickle.load(open(os.path.join(os.path.dirname(cwd),'saved_runs/' + 'Solution_Camp_2_10_100_No control_0_0_False_all'), 'rb'))
data=sols[2.0000000000000004]
category_map = { '0': 'S',
'1': 'E',
'2': 'I',
'3': 'A',
'4': 'R',
'5': 'H',
'6': 'C',
'7': 'D',
'8': 'NE',
'9': 'ND'
}
# +
csv_sol = np.transpose(data['y']) # age structured
solution_csv = pd.DataFrame(csv_sol)
# setup column names
col_names = []
number_categories_with_age = csv_sol.shape[1]
for i in range(number_categories_with_age):
ii = i % params.number_compartments
jj = floor(i/params.number_compartments)
col_names.append(categories[category_map[str(ii)]]['longname'] + ': ' + str(np.asarray(population_frame.Age)[jj]) )
solution_csv.columns = col_names
solution_csv['Time'] = data['t']
for j in range(len(categories.keys())): # params.number_compartments
solution_csv[categories[category_map[str(j)]]['longname']] = data['y_plot'][j] # summary/non age-structured
# this is our dataframe to be saved
# -
solution_csv['R0']=[2.0000000000000004]*solution_csv.shape[0]
solution_csv
final_frame=pd.DataFrame()
for key, value in tqdm(sols.items()):
csv_sol = np.transpose(value['y']) # age structured
solution_csv = pd.DataFrame(csv_sol)
# setup column names
col_names = []
number_categories_with_age = csv_sol.shape[1]
for i in range(number_categories_with_age):
ii = i % params.number_compartments
jj = floor(i/params.number_compartments)
col_names.append(categories[category_map[str(ii)]]['longname'] + ': ' + str(np.asarray(population_frame.Age)[jj]) )
solution_csv.columns = col_names
solution_csv['Time'] = value['t']
for j in range(len(categories.keys())): # params.number_compartments
solution_csv[categories[category_map[str(j)]]['longname']] = value['y_plot'][j] # summary/non age-structured
solution_csv['R0']=[key]*solution_csv.shape[0]
final_frame=pd.concat([final_frame, solution_csv], ignore_index=True)
final_frame.columns
final_frame.to_csv('Baseline.csv')
sns.relplot(x="Time", y="Exposed",
hue="R0", kind="line", data=final_frame);
sns.relplot(x="Time", y="Exposed", kind="line", data=final_frame);
| Archive/Scratch0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/choderalab/pinot/blob/master/scripts/adlala_mol_graph.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rhlRxaENyQ_F" colab_type="text"
# # import
# + id="5BOqiB_qyS29" colab_type="code" outputId="5e0f1260-2404-490d-dd51-5850f8db89f9" colab={"base_uri": "https://localhost:8080/", "height": 153}
# ! rm -rf pinot
# ! git clone https://github.com/choderalab/pinot.git
# + id="XUeCoVY1Mgwf" colab_type="code" outputId="9073b752-3646-4c8e-dbe6-b77f131a01e2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# ! pip install dgl
# ! wget -c https://repo.continuum.io/miniconda/Miniconda3-latest-Linux-x86_64.sh
# ! chmod +x Miniconda3-latest-Linux-x86_64.sh
# ! time bash ./Miniconda3-latest-Linux-x86_64.sh -b -f -p /usr/local
# ! time conda install -q -y -c conda-forge rdkit
# + id="z69rS92vyVI8" colab_type="code" colab={}
import sys
sys.path.append('/usr/local/lib/python3.7/site-packages/')
sys.path.append('/content/pinot/')
# + [markdown] id="Rg-8ac5GyOVY" colab_type="text"
# # data
#
#
# + id="DLApz255yIBK" colab_type="code" outputId="34c1b441-62fc-4300-baea-fd06e6bb50df" colab={"base_uri": "https://localhost:8080/", "height": 411}
import pinot
dir(pinot)
# + id="UUH_sq01yiym" colab_type="code" colab={}
ds = pinot.data.esol()
ds = pinot.data.utils.batch(ds, 32)
# + id="W_LgcjYXbPg-" colab_type="code" colab={}
ds_tr, ds_te = pinot.data.utils.split(ds, [4, 1])
# + [markdown] id="GxiSR0-heZvU" colab_type="text"
# # network
# + id="LLdJeaUeeZJj" colab_type="code" colab={}
net = pinot.representation.Sequential(
lambda in_feat, out_feat: pinot.representation.dgl_legacy.GN(in_feat, out_feat, 'SAGEConv'),
[32, 'tanh', 32, 'tanh', 32, 'tanh', 1])
# + [markdown] id="KTAVTGXhcGU0" colab_type="text"
# # Adam
# + id="zjvfKMeXcFwI" colab_type="code" colab={}
import torch
import numpy as np
opt = torch.optim.Adam(net.parameters(), 1e-3)
loss_fn = torch.nn.functional.mse_loss
# + id="uJKXM-9ReVhQ" colab_type="code" outputId="f4baeeef-5ae1-4bd1-f43a-37c5ccaf04a9" colab={"base_uri": "https://localhost:8080/", "height": 173}
rmse_tr = []
rmse_te = []
for _ in range(100):
for g, y in ds_tr:
opt.zero_grad()
y_hat = net(g)
loss = loss_fn(y, y_hat)
loss.backward()
opt.step()
rmse_tr.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_tr]))
rmse_te.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_te]))
# + id="bhPu3xilfpd1" colab_type="code" outputId="a4a515ab-8370-419c-81b6-b9dcc22bcd52" colab={"base_uri": "https://localhost:8080/", "height": 306}
import matplotlib
from matplotlib import pyplot as plt
plt.rc('font', size=16)
plt.plot(rmse_tr, label='training $RMSE$', linewidth=5, alpha=0.8)
plt.plot(rmse_te, label='test $RMSE$', linewidth=5, alpha=0.8)
plt.xlabel('epochs')
plt.ylabel('$RMSE (\log (\mathtt{mol/L}))$')
plt.legend()
# + [markdown] id="Xh_jemUKmBfU" colab_type="text"
# # Langevin
# + id="1Uqx6pzcmBFs" colab_type="code" colab={}
net = pinot.representation.Sequential(
lambda in_feat, out_feat: pinot.representation.dgl_legacy.GN(in_feat, out_feat, 'SAGEConv'),
[32, 'tanh', 32, 'tanh', 32, 'tanh', 1])
# + id="UBTl57q3iHy3" colab_type="code" colab={}
opt = pinot.inference.adlala.AdLaLa(net.parameters(), partition='La', h=1e-3)
# + id="xRGYdd6qmOOR" colab_type="code" colab={}
rmse_tr = []
rmse_te = []
for _ in range(100):
for g, y in ds_tr:
def l():
opt.zero_grad()
y_hat = net(g)
loss = loss_fn(y, y_hat)
loss.backward()
print(loss)
return loss
opt.step(l)
rmse_tr.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_tr]))
rmse_te.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_te]))
# + id="BWruLqwD5iDO" colab_type="code" outputId="fec0378f-330d-4935-d956-60ac05e37952" colab={"base_uri": "https://localhost:8080/", "height": 306}
import matplotlib
from matplotlib import pyplot as plt
plt.rc('font', size=16)
plt.plot(rmse_tr, label='training $RMSE$', linewidth=5, alpha=0.8)
plt.plot(rmse_te, label='test $RMSE$', linewidth=5, alpha=0.8)
plt.xlabel('epochs')
plt.ylabel('$RMSE (\log (\mathtt{mol/L}))$')
plt.legend()
# + [markdown] id="a2NPtAHb7T9E" colab_type="text"
# # Adaptive Langevin
# + id="WUBRkKlt7M44" colab_type="code" colab={}
net = pinot.representation.Sequential(
lambda in_feat, out_feat: pinot.representation.dgl_legacy.GN(in_feat, out_feat, 'SAGEConv'),
[32, 'tanh', 32, 'tanh', 32, 'tanh', 1])
# + id="VITA-kEr7XP7" colab_type="code" colab={}
opt = pinot.inference.adlala.AdLaLa(net.parameters(), partition='AdLa', h=1e-3)
# + id="LZlNJ-qF7ZfV" colab_type="code" colab={}
rmse_tr = []
rmse_te = []
for _ in range(100):
for g, y in ds_tr:
def l():
opt.zero_grad()
y_hat = net(g)
loss = loss_fn(y, y_hat)
loss.backward()
print(loss)
return loss
opt.step(l)
rmse_tr.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_tr]))
rmse_te.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_te]))
# + id="WpHzw53O7cCq" colab_type="code" outputId="e80c9c21-bd26-4a41-fe8d-820881f80741" colab={"base_uri": "https://localhost:8080/", "height": 306}
import matplotlib
from matplotlib import pyplot as plt
plt.rc('font', size=16)
plt.plot(rmse_tr, label='training $RMSE$', linewidth=5, alpha=0.8)
plt.plot(rmse_te, label='test $RMSE$', linewidth=5, alpha=0.8)
plt.xlabel('epochs')
plt.ylabel('$RMSE (\log (\mathtt{mol/L}))$')
plt.legend()
# + [markdown] id="ohBKFGiA8Ha4" colab_type="text"
# # AdLaLa: AdLa for GN, La for last layer
# + id="TF_G0pIL8G6V" colab_type="code" colab={}
net = pinot.representation.Sequential(
lambda in_feat, out_feat: pinot.representation.dgl_legacy.GN(in_feat, out_feat, 'SAGEConv'),
[32, 'tanh', 32, 'tanh', 32, 'tanh', 1])
# + id="fPF_QwPP8CKN" colab_type="code" outputId="88254198-792d-4f79-cd5d-87f024199181" colab={"base_uri": "https://localhost:8080/", "height": 612}
net
# + id="E5NM7akf8Ozk" colab_type="code" colab={}
opt = pinot.inference.adlala.AdLaLa(
[
{'params': list(net.f_in.parameters())\
+ list(net.d0.parameters())\
+ list(net.d2.parameters())\
+ list(net.d4.parameters()), 'partition': 'AdLa', 'h': torch.tensor(1e-3)},
{
'params': list(net.d6.parameters()) + list(net.f_out.parameters()),
'partition': 'La', 'h': torch.tensor(1e-3)
}
])
# + id="_E6jgO7L9Oup" colab_type="code" colab={}
rmse_tr = []
rmse_te = []
for _ in range(100):
for g, y in ds_tr:
def l():
opt.zero_grad()
y_hat = net(g)
loss = loss_fn(y, y_hat)
loss.backward()
print(loss)
return loss
opt.step(l)
rmse_tr.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_tr]))
rmse_te.append(np.mean([np.sqrt(loss_fn(y, net(g)).detach().numpy()) for g, y in ds_te]))
# + id="k1Wv0whA9TVb" colab_type="code" outputId="6a3c3bd9-af54-4a01-8cba-54c958dad5ea" colab={"base_uri": "https://localhost:8080/", "height": 306}
import matplotlib
from matplotlib import pyplot as plt
plt.rc('font', size=16)
plt.plot(rmse_tr, label='training $RMSE$', linewidth=5, alpha=0.8)
plt.plot(rmse_te, label='test $RMSE$', linewidth=5, alpha=0.8)
plt.xlabel('epochs')
plt.ylabel('$RMSE (\log (\mathtt{mol/L}))$')
plt.legend()
# + id="bdO1Dgb4_-is" colab_type="code" colab={}
| scripts/adlala_experiments/adlala_mol_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:lincs-complimentarity-figures]
# language: R
# name: conda-env-lincs-complimentarity-figures-r
# ---
# ## Compile supplementary table 1
#
# Per compound and MOA median pairwise Pearson correlations
suppressPackageStartupMessages(library(dplyr))
# +
# Load scores
compound_cols <- readr::cols(
compound = readr::col_character(),
no_of_compounds = readr::col_double(),
well = readr::col_character(),
dose_recode = readr::col_double(),
median_score = readr::col_double(),
p_value = readr::col_double(),
assay = readr::col_character(),
normalization = readr::col_character(),
category = readr::col_character(),
pass_thresh = readr::col_logical(),
neg_log_10_p_val = readr::col_double(),
dose = readr::col_character()
)
compound_df <- readr::read_tsv(file.path("results", "compound_scores.tsv"), col_types = compound_cols) %>%
dplyr::select(compound, dose, no_of_compounds, well, median_score, p_value, assay) %>%
dplyr::rename(
no_of_replicates_per_compound = no_of_compounds,
median_replicate_correlation = median_score
)
print(dim(compound_df))
head(compound_df, 3)
# +
moa_cols <- readr::cols(
moa = readr::col_character(),
no_of_replicates = readr::col_double(),
dose = readr::col_character(),
matching_score = readr::col_double(),
assay = readr::col_character(),
p_value = readr::col_double(),
pass_thresh = readr::col_logical(),
neg_log_10_p_val = readr::col_double()
)
moa_df <- readr::read_tsv(file.path("results", "moa_scores.tsv"), col_types = moa_cols) %>%
dplyr::select(moa, dose, no_of_replicates, matching_score, p_value, assay) %>%
dplyr::rename(
no_of_compounds_per_moa = no_of_replicates,
median_replicate_correlation = matching_score
)
print(dim(moa_df))
head(moa_df, 3)
# +
# Load compound to moa map
file <- file.path(
"..",
"1.Data-exploration",
"Consensus",
"cell_painting",
"moa_sizes_consensus_datasets",
"cell_painting_moa_analytical_set_profiles.tsv.gz"
)
df_cols <- readr::cols(
.default = readr::col_double(),
Metadata_Plate_Map_Name = readr::col_character(),
Metadata_cell_id = readr::col_character(),
Metadata_broad_sample = readr::col_character(),
Metadata_pert_well = readr::col_character(),
Metadata_time_point = readr::col_character(),
Metadata_moa = readr::col_character(),
Metadata_target = readr::col_character(),
broad_id = readr::col_character(),
pert_iname = readr::col_character(),
moa = readr::col_character()
)
df <- readr::read_tsv(file, col_types = df_cols) %>%
dplyr::select(pert_iname, moa) %>%
dplyr::distinct()
df$pert_iname <- tolower(df$pert_iname)
df$moa <- tolower(df$moa)
print(dim(df))
head(df, 3)
# +
total_score_df <- compound_df %>%
dplyr::left_join(df, by = c("compound" = "pert_iname")) %>%
dplyr::left_join(moa_df, by = c("moa", "dose", "assay"), suffix = c("_compound", "_moa"))
print(dim(total_score_df))
head(total_score_df, 3)
# -
# Output sup table 1
output_file <- file.path("results", "supplementary_table1.tsv")
total_score_df %>%
dplyr::select(
assay,
compound,
moa,
dose,
well,
no_of_replicates_per_compound,
median_replicate_correlation_compound,
p_value_compound,
no_of_compounds_per_moa,
median_replicate_correlation_moa,
p_value_moa
) %>%
readr::write_tsv(output_file)
| 6.paper_figures/supplementary-table1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# In these exercises we'll apply groupwise analysis to our dataset.
#
# Run the code cell below to load the data before running the exercises.
# +
import pandas as pd
reviews = pd.read_csv("../input/wine-reviews/winemag-data-130k-v2.csv", index_col=0)
#pd.set_option("display.max_rows", 5)
from learntools.core import binder; binder.bind(globals())
from learntools.pandas.grouping_and_sorting import *
print("Setup complete.")
# -
# # Exercises
# ## 1.
# Who are the most common wine reviewers in the dataset? Create a `Series` whose index is the `taster_twitter_handle` category from the dataset, and whose values count how many reviews each person wrote.
# +
# Your code here
reviews_written = ____
# Check your answer
q1.check()
# +
# #%%RM_IF(PROD)%%
reviews_written = reviews.groupby('taster_twitter_handle').size()
q1.assert_check_passed()
# +
# #%%RM_IF(PROD)%%
reviews_written = reviews.groupby('taster_twitter_handle').count()
q1.assert_check_failed()
# +
# #%%RM_IF(PROD)%%
reviews_written = reviews.groupby('taster_twitter_handle')['taster_twitter_handle'].count()
q1.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q1.hint()
#_COMMENT_IF(PROD)_
q1.solution()
# ## 2.
# What is the best wine I can buy for a given amount of money? Create a `Series` whose index is wine prices and whose values is the maximum number of points a wine costing that much was given in a review. Sort the values by price, ascending (so that `4.0` dollars is at the top and `3300.0` dollars is at the bottom).
# +
best_rating_per_price = ____
# Check your answer
q2.check()
# +
# #%%RM_IF(PROD)%%
best_rating_per_price = reviews.groupby('price')['points'].max().sort_index()
q2.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q2.hint()
#_COMMENT_IF(PROD)_
q2.solution()
# ## 3.
# What are the minimum and maximum prices for each `variety` of wine? Create a `DataFrame` whose index is the `variety` category from the dataset and whose values are the `min` and `max` values thereof.
# +
price_extremes = ____
# Check your answer
q3.check()
# +
# #%%RM_IF(PROD)%%
price_extremes = reviews.groupby('variety').price.agg([min, max])
q3.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q3.hint()
#_COMMENT_IF(PROD)_
q3.solution()
# ## 4.
# What are the most expensive wine varieties? Create a variable `sorted_varieties` containing a copy of the dataframe from the previous question where varieties are sorted in descending order based on minimum price, then on maximum price (to break ties).
# +
sorted_varieties = ____
# Check your answer
q4.check()
# +
# #%%RM_IF(PROD)%%
sorted_varieties = price_extremes.sort_values(by=['min', 'max'], ascending=False)
q4.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q4.hint()
#_COMMENT_IF(PROD)_
q4.solution()
# ## 5.
# Create a `Series` whose index is reviewers and whose values is the average review score given out by that reviewer. Hint: you will need the `taster_name` and `points` columns.
# +
reviewer_mean_ratings = ____
# Check your answer
q5.check()
# +
# #%%RM_IF(PROD)%%
reviewer_mean_ratings = reviews.groupby('taster_name').points.mean()
q5.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q5.hint()
#_COMMENT_IF(PROD)_
q5.solution()
# Are there significant differences in the average scores assigned by the various reviewers? Run the cell below to use the `describe()` method to see a summary of the range of values.
reviewer_mean_ratings.describe()
# ## 6.
# What combination of countries and varieties are most common? Create a `Series` whose index is a `MultiIndex`of `{country, variety}` pairs. For example, a pinot noir produced in the US should map to `{"US", "Pinot Noir"}`. Sort the values in the `Series` in descending order based on wine count.
# +
country_variety_counts = ____
# Check your answer
q6.check()
# +
# #%%RM_IF(PROD)%%
country_variety_counts = reviews.groupby(['country', 'variety']).size().sort_values(ascending=False)
q6.assert_check_passed()
# -
#_COMMENT_IF(PROD)_
q6.hint()
#_COMMENT_IF(PROD)_
q6.solution()
# # Keep going
#
# Move on to the [**data types and missing data**](#$NEXT_NOTEBOOK_URL$).
| notebooks/pandas/raw/ex_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Writing Custom Dataset Importers
#
# This recipe demonstrates how to write a [custom DatasetImporter](https://voxel51.com/docs/fiftyone/user_guide/dataset_creation/datasets.html#custom-formats) and use it to load a dataset from disk in your custom format into FiftyOne.
# ## Setup
#
# If you haven't already, install FiftyOne:
# !pip install fiftyone
# In this recipe we'll use the [FiftyOne Dataset Zoo](https://voxel51.com/docs/fiftyone/user_guide/dataset_creation/zoo_datasets.html) to download the [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html) to use as sample data to feed our custom importer.
#
# Behind the scenes, FiftyOne either the
# [TensorFlow Datasets](https://www.tensorflow.org/datasets) or
# [TorchVision Datasets](https://pytorch.org/docs/stable/torchvision/datasets.html) libraries to wrangle the datasets, depending on which ML library you have installed.
#
# You can, for example, install PyTorch as follows:
# !pip install torch torchvision
# ## Writing a DatasetImporter
#
# FiftyOne provides a [DatasetImporter](https://voxel51.com/docs/fiftyone/api/fiftyone.utils.data.html#fiftyone.utils.data.importers.DatasetImporter) interface that defines how it imports datasets from disk when methods such as [Dataset.from_importer()](https://voxel51.com/docs/fiftyone/api/fiftyone.core.html#fiftyone.core.dataset.Dataset.from_importer) are used.
#
# `DatasetImporter` itself is an abstract interface; the concrete interface that you should implement is determined by the type of dataset that you are importing. See [writing a custom DatasetImporter](https://voxel51.com/docs/fiftyone/user_guide/dataset_creation/datasets.html#custom-formats) for full details.
#
# In this recipe, we'll write a custom [LabeledImageDatasetImporter](https://voxel51.com/docs/fiftyone/api/fiftyone.utils.data.html#fiftyone.utils.data.importers.LabeledImageDatasetImporter) that can import an image classification dataset whose image metadata and labels are stored in a `labels.csv` file in the dataset directory with the following format:
#
# ```
# filepath,size_bytes,mime_type,width,height,num_channels,label
# <filepath>,<size_bytes>,<mime_type>,<width>,<height>,<num_channels>,<label>
# <filepath>,<size_bytes>,<mime_type>,<width>,<height>,<num_channels>,<label>
# ...
# ```
# Here's the complete definition of the `DatasetImporter`:
# +
import csv
import os
import fiftyone as fo
import fiftyone.utils.data as foud
class CSVImageClassificationDatasetImporter(foud.LabeledImageDatasetImporter):
"""Importer for image classification datasets whose filepaths and labels
are stored on disk in a CSV file.
Datasets of this type should contain a ``labels.csv`` file in their
dataset directories in the following format::
filepath,size_bytes,mime_type,width,height,num_channels,label
<filepath>,<size_bytes>,<mime_type>,<width>,<height>,<num_channels>,<label>
<filepath>,<size_bytes>,<mime_type>,<width>,<height>,<num_channels>,<label>
...
Args:
dataset_dir: the dataset directory
skip_unlabeled (False): whether to skip unlabeled images when importing
shuffle (False): whether to randomly shuffle the order in which the
samples are imported
seed (None): a random seed to use when shuffling
max_samples (None): a maximum number of samples to import. By default,
all samples are imported
"""
def __init__(
self,
dataset_dir,
skip_unlabeled=False,
shuffle=False,
seed=None,
max_samples=None,
):
super().__init__(
dataset_dir=dataset_dir,
skip_unlabeled=skip_unlabeled,
shuffle=shuffle,
seed=seed,
max_samples=max_samples
)
self._labels_file = None
self._labels = None
self._iter_labels = None
def __iter__(self):
self._iter_labels = iter(self._labels)
return self
def __next__(self):
"""Returns information about the next sample in the dataset.
Returns:
an ``(image_path, image_metadata, label)`` tuple, where
- ``image_path``: the path to the image on disk
- ``image_metadata``: an
:class:`fiftyone.core.metadata.ImageMetadata` instances for the
image, or ``None`` if :meth:`has_image_metadata` is ``False``
- ``label``: an instance of :meth:`label_cls`, or a dictionary
mapping field names to :class:`fiftyone.core.labels.Label`
instances, or ``None`` if the sample is unlabeled
Raises:
StopIteration: if there are no more samples to import
"""
(
filepath,
size_bytes,
mime_type,
width,
height,
num_channels,
label,
) = next(self._iter_labels)
image_metadata = fo.ImageMetadata(
size_bytes=size_bytes,
mime_type=mime_type,
width=width,
height=height,
num_channels=num_channels,
)
label = fo.Classification(label=label)
return filepath, image_metadata, label
def __len__(self):
"""The total number of samples that will be imported.
Raises:
TypeError: if the total number is not known
"""
return len(self._labels)
@property
def has_dataset_info(self):
"""Whether this importer produces a dataset info dictionary."""
return False
@property
def has_image_metadata(self):
"""Whether this importer produces
:class:`fiftyone.core.metadata.ImageMetadata` instances for each image.
"""
return True
@property
def label_cls(self):
"""The :class:`fiftyone.core.labels.Label` class(es) returned by this
importer.
This can be any of the following:
- a :class:`fiftyone.core.labels.Label` class. In this case, the
importer is guaranteed to return labels of this type
- a dict mapping keys to :class:`fiftyone.core.labels.Label` classes.
In this case, the importer will return label dictionaries with keys
and value-types specified by this dictionary. Not all keys need be
present in the imported labels
- ``None``. In this case, the importer makes no guarantees about the
labels that it may return
"""
return fo.Classification
def setup(self):
"""Performs any necessary setup before importing the first sample in
the dataset.
This method is called when the importer's context manager interface is
entered, :func:`DatasetImporter.__enter__`.
"""
labels_path = os.path.join(self.dataset_dir, "labels.csv")
labels = []
with open(labels_path, "r") as f:
reader = csv.DictReader(f)
for row in reader:
if self.skip_unlabeled and not row["label"]:
continue
labels.append((
row["filepath"],
row["size_bytes"],
row["mime_type"],
row["width"],
row["height"],
row["num_channels"],
row["label"],
))
# The `_preprocess_list()` function is provided by the base class
# and handles shuffling/max sample limits
self._labels = self._preprocess_list(labels)
def close(self, *args):
"""Performs any necessary actions after the last sample has been
imported.
This method is called when the importer's context manager interface is
exited, :func:`DatasetImporter.__exit__`.
Args:
*args: the arguments to :func:`DatasetImporter.__exit__`
"""
pass
# -
# ## Generating a sample dataset
# In order to use `CSVImageClassificationDatasetImporter`, we need to generate a sample dataset in the required format.
#
# Let's first write a small utility to populate a `labels.csv` file in the required format.
def write_csv_labels(samples, csv_path, label_field="ground_truth"):
"""Writes a labels CSV format for the given samples in the format expected
by :class:`CSVImageClassificationDatasetImporter`.
Args:
samples: an iterable of :class:`fiftyone.core.sample.Sample` instances
csv_path: the path to write the CSV file
label_field ("ground_truth"): the label field of the samples to write
"""
# Ensure base directory exists
basedir = os.path.dirname(csv_path)
if basedir and not os.path.isdir(basedir):
os.makedirs(basedir)
# Write the labels
with open(csv_path, "w") as f:
writer = csv.writer(f)
writer.writerow([
"filepath",
"size_bytes",
"mime_type",
"width",
"height",
"num_channels",
"label",
])
for sample in samples:
filepath = sample.filepath
metadata = sample.metadata
if metadata is None:
metadata = fo.ImageMetadata.build_for(filepath)
label = sample[label_field].label
writer.writerow([
filepath,
metadata.size_bytes,
metadata.mime_type,
metadata.width,
metadata.height,
metadata.num_channels,
label,
])
# Now let's populate a directory with a `labels.csv` file in the format required by `CSVImageClassificationDatasetImporter` with some samples from the test split of CIFAR-10:
# +
import fiftyone.zoo as foz
dataset_dir = "/tmp/fiftyone/custom-dataset-importer"
num_samples = 1000
#
# Load `num_samples` from CIFAR-10
#
# This command will download the test split of CIFAR-10 from the web the first
# time it is executed, if necessary
#
cifar10_test = foz.load_zoo_dataset("cifar10", split="test")
samples = cifar10_test.limit(num_samples)
# This dataset format requires samples to have their `metadata` fields populated
print("Computing metadata for samples")
samples.compute_metadata()
# Write labels to disk in CSV format
csv_path = os.path.join(dataset_dir, "labels.csv")
print("Writing labels for %d samples to '%s'" % (num_samples, csv_path))
write_csv_labels(samples, csv_path)
# -
# Let's inspect the contents of the labels CSV to ensure they're in the correct format:
# !head -n 10 /tmp/fiftyone/custom-dataset-importer/labels.csv
# ## Importing a dataset
# With our dataset and `DatasetImporter` in-hand, loading the data as a FiftyOne dataset is as simple as follows:
# Import the dataset
print("Importing dataset from '%s'" % dataset_dir)
importer = CSVImageClassificationDatasetImporter(dataset_dir)
dataset = fo.Dataset.from_importer(importer)
# Print summary information about the dataset
print(dataset)
# Print a sample
print(dataset.first())
# ## Cleanup
#
# You can cleanup the files generated by this recipe by running:
# !rm -rf /tmp/fiftyone
| docs/source/recipes/custom_importer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementation details: deriving expected moment dynamics
# $$
# \def\n{\mathbf{n}}
# \def\x{\mathbf{x}}
# \def\N{\mathbb{\mathbb{N}}}
# \def\X{\mathbb{X}}
# \def\NX{\mathbb{\N_0^\X}}
# \def\C{\mathcal{C}}
# \def\Jc{\mathcal{J}_c}
# \def\DM{\Delta M_{c,j}}
# \newcommand\diff{\mathop{}\!\mathrm{d}}
# \def\Xc{\mathbf{X}_c}
# \newcommand{\muset}[1]{\dot{\{}#1\dot{\}}}
# $$
# This notebook walks through what happens inside `compute_moment_equations()`.
# We restate the algorithm outline, adding code snippets for each step.
# This should help to track down issues when, unavoidably, something fails inside `compute_moment_equations()`.
# +
# initialize sympy printing (for latex output)
from sympy import init_printing
init_printing()
# import functions and classes for compartment models
from compartor import *
from compartor.compartments import ito, decomposeMomentsPolynomial, getCompartments, getDeltaM, subsDeltaM, get_dfMdt_contrib
# -
# We only need one transition class.
# We use "coagulation" from the coagulation-fragmentation example
# +
D = 1 # number of species
x = Content('x')
y = Content('y')
transition_C = Transition(Compartment(x) + Compartment(y), Compartment(x + y), name = 'C')
k_C = Constant('k_C')
g_C = 1
Coagulation = TransitionClass(transition_C, k_C, g_C)
transition_classes = [Coagulation]
display_transition_classes(transition_classes)
# -
# $$
# \def\n{\mathbf{n}}
# \def\x{\mathbf{x}}
# \def\N{\mathbb{\mathbb{N}}}
# \def\X{\mathbb{X}}
# \def\NX{\mathbb{\N_0^\X}}
# \def\C{\mathcal{C}}
# \def\Jc{\mathcal{J}_c}
# \def\DM{\Delta M_{c,j}}
# \newcommand\diff{\mathop{}\!\mathrm{d}}
# \def\Xc{\mathbf{X}_c}
# \newcommand{\muset}[1]{\dot{\{}#1\dot{\}}}
# $$
#
#
# For a compartment population $\n \in \NX$ evolving stochastically according to stoichiometric equations from transition classes $\C$, we want to find an expression for
# $$
# \frac{\diff}{\diff t}\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>
# $$
# in terms of expectations of population moments $M^\alpha, M^{\beta}, \ldots$
fM = Moment(0)**2
display(fM)
# ### (1)
# From the definition of the compartment dynamics, we have
# $$
# \diff M^\gamma = \sum_{c \in \C} \sum_{j \in \Jc} \DM^\gamma \diff R_{c,j}
# $$
# We apply Ito's rule to derive
# $$
# \diff f(M^\gamma, M^{\gamma'}, \ldots) = \sum_{c \in \C} \sum_{j \in \Jc}
# \left(
# f(M^\gamma + \DM^\gamma, M^{\gamma'} + \DM^{\gamma'}, \ldots)
# - f(M^\gamma, M^{\gamma'}, \ldots)
# \right) \diff R_{c,j}
# $$
# Assume, that $f(M^\gamma, M^{\gamma'}, \ldots)$ is a polynomial in $M^{\gamma^i}$ with $\gamma^i \in \N_0^D$.
#
# Then $\diff f(M^\gamma, M^{\gamma'}, \ldots)$ is a polynomial in $M^{\gamma^k}, \DM^{\gamma^l}$ with $\gamma^k, \gamma^l \in \N_0^D$, that is,
# $$
# \diff f(M^\gamma, M^{\gamma'}, \ldots) = \sum_{c \in \C} \sum_{j \in \Jc}
# \sum_{q=1}^{n_q} Q_q(M^{\gamma^k}, \DM^{\gamma^l})
# \diff R_{c,j}
# $$
# where $Q_q(M^{\gamma^k}, \DM^{\gamma^l})$ are monomials in $M^{\gamma^k}, \DM^{\gamma^l}$.
dfM = ito(fM)
dfM
# ### (2)
# Let's write $Q_q(M^{\gamma^k}, \DM^{\gamma^l})$ as
# $$
# Q_q(M^{\gamma^k}, \DM^{\gamma^l}) = k_q \cdot \Pi M^{\gamma^k} \cdot \Pi M^{\gamma^k}
# $$
# where $k_q$ is a constant,
# $\Pi M^{\gamma^k}$ is a product of powers of $M^{\gamma^k}$, and
# $\Pi \DM^{\gamma^l}$ is a product of powers of $\DM^{\gamma^l}$.
#
# Analogous to the derivation in SI Appendix S.3, we arrive at the expected moment dynamics
# $$
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t} =
# \sum_{c \in \C} \sum_{q=1}^{n_q} \left<
# \sum_{j \in \Jc} k_q \cdot \Pi M^{\gamma^k} \cdot \Pi \DM^{\gamma^k} \cdot h_{c,j}(\n)
# \right>
# $$
monomials = decomposeMomentsPolynomial(dfM)
monomials
# ### (3)
# Analogous to SI Appendix S.4, the contribution of class $c$, monomial $q$ to the expected dynamics of $f(M^\gamma, M^{\gamma'}, \ldots)$ is
# $$
# \begin{align}
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t}
# &= \left<
# {\large\sum_{j \in \Jc}} k_q \cdot \Pi M^{\gamma^k} \cdot \Pi \DM^{\gamma^l} \cdot h_{c,j}(\n)
# \right>
# \\
# &= \left<
# {\large\sum_{\Xc}} w(\n; \Xc) \cdot k_c \cdot k_q \cdot \Pi M^{\gamma^k} \cdot g_c(\Xc) \cdot
# \left<
# \Pi \DM^{\gamma^l} \;\big|\; \Xc
# \right>
# \right>
# \end{align}
# $$
#
# +
c = 0 # take the first transition class
q = 1 # ... and the second monomial
tc = transition_classes[c]
transition, k_c, g_c, pi_c = tc.transition, tc.k, tc.g, tc.pi
(k_q, pM, pDM) = monomials[q]
# -
# First we compute the expression
# $$
# l(\n; \Xc) = k_c \cdot k_q \cdot \Pi(M^{\gamma^k}) \cdot g_c(\Xc) \cdot
# \left<
# \Pi \DM^{\gamma^l} \;\big|\; \Xc
# \right>
# $$
# We start by computing the $\DM^{\gamma^l}$ from reactants and products of the transition ...
reactants = getCompartments(transition.lhs)
products = getCompartments(transition.rhs)
DM_cj = getDeltaM(reactants, products, D)
DM_cj
# ... and then substituting this expression into every occurence of $\DM^\gamma$ in `pDM` (with the $\gamma$ in `DM_cj` set appropriately).
pDMcj = subsDeltaM(pDM, DM_cj)
print('pDM = ')
display(pDM)
print('pDMcj = ')
display(pDMcj)
# Then we compute the conditional expectation of the result.
cexp = pi_c.conditional_expectation(pDMcj)
cexp
# Finally we multiply the conditional expectation with the rest of the terms:
# * $k_c$, and $g_c(\Xc)$ from the specification of `transition[c]`, and
# * $k_q$, and $\Pi(M^{\gamma^k})$ from `monomials[q]`.
l_n_Xc = k_c * k_q * pM * g_c * cexp
l_n_Xc
# ### (4)
# Let's consider the expression $A = \sum_{\Xc} w(\n; \Xc) \cdot l(\n; \Xc)$ for the following cases of reactant compartments:
# $\Xc = \emptyset$,
# $\Xc = \muset{\x}$, and
# $\Xc = \muset{\x, \x'}$.
#
# (1) $\Xc = \emptyset$:
#
# Then $w(\n; \Xc) = 1$, and
# $$
# A = l(\n)
# $$
#
# (2) $\Xc = \muset{\x}$:
#
# Then $w(\n; \Xc) = \n(\x)$, and
# $$
# A = \sum_{\x \in \X} \n(\x) \cdot l(\n; \muset{\x})
# $$
#
# (3) $\Xc = \muset{\x, \x'}$:
#
# Then
# $$
# w(\n; \Xc) = \frac{\n(\x)\cdot(\n(\x')-\delta_{\x,\x'})}
# {1+\delta_{\x,\x'}},
# $$
# and
# $$
# \begin{align}
# A &= \sum_{\x \in \X} \sum_{\x' \in \X}
# \frac{1}{2-\delta_{\x,\x'}}
# \cdot w(\n; \Xc) \cdot l(\n; \muset{\x, \x'}) \\
# &= \sum_{\x \in \X} \sum_{\x' \in \X}
# \frac{\n(\x)\cdot(\n(\x')-\delta_{\x,\x'})}{2}
# \cdot l(\n; \muset{\x, \x'}) \\
# &= \sum_{\x \in \X} \sum_{\x' \in \X}
# \n(\x)\cdot\n(\x') \cdot \frac{1}{2}l(\n; \muset{\x, \x'})
# \: - \:
# \sum_{\x \in \X}
# \n(\x) \cdot \frac{1}{2}l(\n; \muset{\x, \x})
# \end{align}
# $$
# ### (5)
# Now let
# $$
# l(\n; \Xc) = k_c \cdot k_q \cdot \Pi(M^{\gamma^k}) \cdot g_c(\Xc) \cdot
# \left<
# \Pi \DM^{\gamma^l} \;\big|\; \Xc
# \right>
# $$
#
# Plugging in the concrete $\gamma^l$ and expanding, $l(\n; \Xc)$ is a polynomial in $\Xc$.
#
# Monomials are of the form $k \x^\alpha$ or $k \x^\alpha \x'^\beta$ with $\alpha, \beta \in \N_0^D$.
# (Note that occurences of $\Pi M^{\gamma^k}$ are part of the constants $k$.)
#
# Consider again the different cases of reactant compartments $\Xc$:
#
# (1) $\Xc = \emptyset$:
# $$
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t}
# = \left<l(\n)\right>
# $$
#
# (2) $\Xc = \muset{\x}$:
# $$
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t}
# = \left<R(l(\n; \muset{\x})\right>
# $$
# where $R$ replaces all $k \x^\alpha$ by $k M^\alpha$.
#
# (3) $\Xc = \muset{\x, \x'}$:
# $$
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t}
# = \frac{1}{2}\left<R'(l(\n; \muset{\x, \x'})\right>
# \: - \:
# \frac{1}{2}\left<R(l(\n; \muset{\x, \x})\right>
# $$
# where $R'$ replaces all $k \x^\alpha \X'^\beta$ by $k M^\alpha M^\beta$,
# and again $R$ replaces all $k \x^\alpha$ by $k M^\alpha$.
# All this (the case destinction and replacements) is done in the function `get_dfMdt_contrib()`.
dfMdt = get_dfMdt_contrib(reactants, l_n_Xc, D)
dfMdt
# ### (6)
# Finally, sum over contributions from all $c$, $q$ for the total
# $$
# \frac{\diff\left< f(M^\gamma, M^{\gamma'}, \ldots) \right>}{\diff t}
# $$
| (EXTRA) Implementation details.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:metis] *
# language: python
# name: conda-env-metis-py
# ---
import pandas as pd
from bs4 import BeautifulSoup
import requests
import os
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import re
import time
import numpy as np
# +
# driver setup
chromedriver = "/Applications/chromedriver" # path to the chromedriver executable
os.environ["webdriver.chrome.driver"] = chromedriver
driver = webdriver.Chrome(chromedriver)
# -
# # <NAME>
# +
def get_attr(node, attr):
if node == None:
return None
if attr == 'text':
return node.text
return node.get(attr)
def scrape_page(source_text):
soup = BeautifulSoup(source_text)
data = []
for row in soup.find('div', class_='member_directory').find_all('tr', class_='list member'):
name = get_attr(row.find('div', class_='title').find('a'), 'text')
chamber_profile_url = get_attr(row.find('div', class_='title').find('a'), 'href')
info_table = row.find('table', class_='info')
phone = get_attr(info_table.find('td'), 'text')
email = get_attr(info_table.find('a', href=re.compile("mailto:")), 'href')
website = get_attr(info_table.find('a', target='_blank'), 'href')
fb = get_attr(info_table.find('a', class_='facebook'), 'href')
twitter = get_attr(info_table.find('a', class_='twitter'), 'href')
insta = get_attr(info_table.find('a', class_='instagram'), 'href')
data.append([name, chamber_profile_url, phone, email, website, fb, twitter, insta])
return data
# -
def scrape_lincoln_park(drv):
drv.get('https://www.lincolnparkchamber.com/directory/')
button = drv.find_element_by_css_selector('button.list-display')
button.click()
time.sleep(3)
more_pages = True
data = []
while more_pages:
new_data = scrape_page(drv.page_source)
data.append(new_data)
try:
next_button = drv.find_element_by_css_selector('span.right-arrow')
next_button.click()
time.sleep(3)
except:
more_pages = False
return data
results = scrape_lincoln_park(driver)
flattened = [row for page in results for row in page]
len(flattened)
lincoln_park = pd.DataFrame(flattened, columns=['name', 'chamber_profile_url', 'phone', 'email', 'website', 'fb', 'twitter', 'insta'])
lincoln_park.phone.value_counts()
# replace Email with nan
# cut 'mailto:' out of email
lincoln_park.email = lincoln_park.email.str.replace('mailto:', '')
lincoln_park.phone = lincoln_park.phone.replace(to_replace='Email', value=np.nan)
lincoln_park.to_csv('../data/processed/lincoln_park_chamber_scraped_0116.csv')
# # Edgewater
def scrape_edgewater_directory(drv):
drv.get('https://www.edgewater.org/membership-directory/')
for _ in range(0, 5):
driver.execute_script(
"window.scrollTo(0, document.documentElement.scrollHeight);" #Alternatively, document.body.scrollHeight
)
time.sleep(3)
soup = BeautifulSoup(drv.page_source)
data = []
for tile in soup.find_all('div', class_='article_box'):
profile_page = tile.find('a').get('href')
business_name = tile.find('div', class_='desc_wrapper').find('h4').text
data.append([business_name, profile_page])
return data
data = scrape_edgewater_directory(driver)
data
def add_edgewater_contact_info(row, drv):
name, url = row
drv.get(url)
soup = BeautifulSoup(drv.page_source)
card = soup.find('div', class_='business')
address = get_attr(card.find('p', id='address'), 'text')
email = get_attr(card.find('p', id='email'), 'text')
link = card.find('a')
website = get_attr(link, 'href')
fb = get_attr(card.find('a', attrs={'title': 'Facebook'}), 'href')
twitter = get_attr(card.find('a', attrs={'title': 'Twitter'}), 'href')
if link:
desc = get_attr(link.find_parent(), 'text')
match = re.search(r'\(\d{3}\) \d{3}-\d{4}', desc)
if match:
return [address, email, website, match.group(0), fb, twitter]
return [address, email, website, None, fb, twitter]
# +
enriched_data = []
for row in data:
try:
new_data = add_edgewater_contact_info(row, driver)
print(new_data)
enriched_data.append(row + new_data)
time.sleep(1)
except:
enriched_data.append(row + [None, None, None, None, None, None])
# -
df = pd.DataFrame(enriched_data, columns=['name', 'chamber_profile_url', 'address', 'email', 'website', 'phone', 'fb', 'twitter'])
df.to_csv('../data/processed/edgewater_chamber_scraped_0116.csv')
# # <NAME>
def scrape_wicker():
rows = []
for pos in range(0,271,30):
url = f'http://www.wickerparkbucktown.com/index.php?src=membership&srctype=membership_lister_alpha&pos={pos},30,296'
new_rows = scrape_wicker_directory_page(url)
rows = rows + new_rows
return rows
# +
def scrape_wicker_directory_page(url):
rows = []
directory_request = requests.get(url)
soup = BeautifulSoup(directory_request.text)
for title_link in soup.find_all('a', class_='title'):
company_profile_url = title_link.get('href')
row = [title_link.text, company_profile_url]
other_fields = scrape_wicker_profile(company_profile_url)
rows.append(row + other_fields)
time.sleep(1)
return rows
def scrape_wicker_profile(url):
profile_req = requests.get(f'http://www.wickerparkbucktown.com/{url}')
profile_soup = BeautifulSoup(profile_req.text)
address = get_attr(profile_soup.find('div', class_='address'), 'text')
contact_title = get_attr(profile_soup.find('div', class_='jobTitle'), 'text')
phone = get_attr(profile_soup.find('div', class_='phone'), 'text')
email = get_attr(profile_soup.find('div', class_='email'), 'text')
company_website = get_attr(profile_soup.find('div', class_='website'), 'text')
contact_name = get_attr(profile_soup.find('h2'), 'text')
return [address, contact_name, contact_title, phone, email]
# -
scrape_wicker_directory_page('http://www.wickerparkbucktown.com/index.php?src=membership&srctype=membership_lister_alpha&pos=0,30,296')
results = scrape_wicker()
results
len(results)
columns = ['company_name', 'profile_url', 'address', 'contact_name', 'contact_title', 'phone', 'email']
wicker = pd.DataFrame(results, columns = columns)
# +
def clean_wicker_columns(df):
df.address = df.address.map(lambda v: v.split(':')[-1].strip() if v else None)
df.contact_title = df.contact_title.map(lambda v: v.split(':')[-1].strip() if v else None)
df.phone = df.phone.map(lambda v: v.split(':')[-1].strip() if v else None)
df.email = df.email.str.replace("Email: document.write\(|\+|'|\);|/s", '')
# -
clean_wicker_columns(wicker)
wicker.to_csv('../data/processed/wicker_chamber_scraped_0117.csv')
| notebooks/scrapeChambersOfCommerce.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp models.base
# +
# export
import torch
from torch import nn
from uberduck_ml_dev.text.symbols import SYMBOL_SETS
class TTSModel(nn.Module):
def __init__(self, hparams):
super().__init__()
self.symbol_set = hparams.symbol_set
self.n_symbols = len(SYMBOL_SETS[self.symbol_set])
self.n_speakers = hparams.n_speakers
# symbols = __import__('uberduck_ml_dev.text.' + hparams.symbols)
def infer(self):
raise NotImplemented
def forward(self):
raise NotImplemented
def from_pretrained(
self, warm_start_path=None, device="cpu", ignore_layers=None, model_dict=None
):
model_dict = model_dict or dict()
if warm_start_path is None and model_dict is None:
raise Exception(
"TTSModel.from_pretrained requires a warm_start_path or state_dict"
)
if warm_start_path is not None:
checkpoint = torch.load(warm_start_path, map_location=device)
if (
"state_dict" in checkpoint.keys()
): # TODO: remove state_dict once off nvidia
model_dict = checkpoint["state_dict"]
if "model" in checkpoint.keys():
model_dict = checkpoint["model"]
if ignore_layers:
model_dict = {k: v for k, v in model_dict.items() if k not in ignore_layers}
dummy_dict = self.state_dict()
for k in self.state_dict().keys():
if k not in model_dict.keys():
print(
f"WARNING! Attempting to load a model with out the {k} layer. This could lead to unexpected results during evaluation."
)
dummy_dict.update(model_dict)
model_dict = dummy_dict
self.load_state_dict(model_dict)
if device == "cuda":
self.cuda()
def to_checkpoint(self):
return dict(model=self.state_dict())
@classmethod
def create(cls, name, opts, folders, all_speakers=True):
pass
# +
# export
from uberduck_ml_dev.vendor.tfcompat.hparam import HParams
DEFAULTS = HParams(
p_arpabet=1.0,
seed=1234,
)
| nbs/models.base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Explore data from Digital Language Death
#
# ## Input
#
# ### Digital Language Death (DLD)
#
# Paper (2012): https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0077056
# Data: http://hlt.sztaki.hu/resources/
# Data description: http://hlt.sztaki.hu/resources/dld-header.pdf
# Online version: https://hlt.bme.hu/en/dld/
#
# ### Crúbadán project
#
# DLD data uses the Crúbadán crawls. The aim of this project is the creation of text corpora for a large number of under-resourced languages by crawling the web. It supports more than 2000 languages.
#
# Paper (2007): https://cs.slu.edu/~scannell/pub/wac3.pdf
# Project website: http://crubadan.org/
#
# ## Output
#
# `dld_2012.csv`
#
# - non-explicit index: lang code
# - for all European languages
# - status of endangerment
# - availability of digital tools
#
# -
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import Normalizer
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.metrics import explained_variance_score
# ## Read Digital Language Death (DLD) data
dld = pd.read_csv("../../data/digital/dld_joined.tsv", sep="\t", skiprows=2, low_memory=False)
for i, col in enumerate(dld.columns):
print(f"{i}\t{col}")
pd.unique(dld["cru1_ISO-639-3"].values.ravel())
display(dld)
# ### Filter columns of interest
# +
# Ids & Names
PRINT_NAME = "Print_Name" # standardized English language name
LANG_LOC = "language (local)" # the local name of the language
SIL = "SIL code" # SIL code
# Endangerment status
ETHN = "Eth_Language Status" # the EGIDS status of the language according to the Ethnologue.
ELP = "end_class" # classification according to the Endangered Languages Project (the higher, the more endangered)
# Speakers
L1 = "L1" # number of people speaking the language natively (L1 speakers)
L2 = "L2" # number of people speaking the language as second language (L2 speakers)
# Wikipedia
WP_NAME = "wikiname" # the two-letter (sometimes longer) code used in Wikipedia
WP_ART = "articles" # number of articles in wikipedia (2012 May)
WP_SIZE_ADJ = "adjusted WP size" # character count of ‘real’ wikipedia pages
WP_SIZE_CH = "WP size in chars" # raw (unadjusted) character count of wikipedia
WP_INCUB = "WP_incubator_new" # whether Wikipedia had an incubator for the language in March 2013
WP_AVG_LEN = "avg good page length" # average length of ‘real’ wikipedia pages
WP_REAL_TOT = "real/ total ratio" # proportion of ‘real’ and 'total' wikipedia pages
# Windows
MS_IN = "MS-if-pack" # Microsoft input-level support
MS_OS = "MS-pack" # OS-level support in Windows 7
# MAC
MAC_IN = "MAC-input" # input-level support by Apple
MAC_OS = "MAC-supp" # OS-level support in MacOS 10.6.8
# Spellchecker
HUN_STAT = "hunspell status" # whether a hunspell checker exists
HUN_COV = "hunspell coverage" # the percentage coverage Hunspell has on the wikipedia dump
# Crúbadán
CR1_ID = "cru1_ISO-639-3" # iso-code from first Crúbadán summary
CR2_ID = "cru2_ISO-639-3" # iso-code from second Crúbadán summary
CR1_CH = "cru1_Characters" # number of characters found by the first Crúbadán crawl
CR2_CH = "cru2_Characters" # number of characters found by the second Crúbadán crawl
CR1_WORDS = "cru1_Words" # number of words found by the first Crúbadán crawl
CR2_WORDS = "cru2_Words" # number of words found by the second Crúbadán crawl
CR1_DOCS = "cru1_Docs" # number of documents found by the first Crúbadán crawl
CR2_DOCS = "cru2_Docs" # number of documents found by the second Crúbadán crawl
CR1_UDHR = "cru1_UDHR" # translation of the Universal Declaration of Human rights exists according to the first Crúbadán summary
CR2_UDHR = "cru2_UDHR" # translation of the Universal Declaration of Human rights exists according to the second Crúbadán summary
CR1_FLOSS = "cru1_FLOSS SplChk" # whether a FLOSS spellchecker exists according to the first Crúbadán summary
CR2_FLOSS = "cru2_FLOSS SplChk" # whether a FLOSS spellchecker exists according to the second Crúbadán summary
CR1_BIB = "cru1_WT" # whether an online Bible exists at watchtower.org according to the first Crúbadán summary
CR2_BIB = "cru2_WT" # whether an online Bible exists at watchtower.org according to the second Crúbadán summary
# Other
TLD = "TLDs(%)" # whether a national level Top Level Domain (not .com, .org, .edu) appeared in the top three domains that the crawl found language data in
OLAC = "la_Primary texts_Online" # the number of OLAC primary texts online
# -
dld_fil = dld[[
# Ids & names
PRINT_NAME,
LANG_LOC,
SIL,
# Endangerment status
ETHN,
ELP,
# Speakers
L1,
L2,
# Wikipedia
WP_NAME,
WP_ART,
WP_SIZE_ADJ,
WP_SIZE_CH,
WP_INCUB,
WP_AVG_LEN,
WP_REAL_TOT,
# Windows
MS_IN,
MS_OS,
# MAC
MAC_IN,
MAC_OS,
# Spellchecker
HUN_STAT,
HUN_COV,
# Crúbadán
CR1_ID,
CR2_ID,
CR1_CH,
CR2_CH,
CR1_WORDS,
CR2_WORDS,
CR1_DOCS,
CR2_DOCS,
CR1_UDHR,
CR2_UDHR,
CR1_FLOSS,
CR2_FLOSS,
CR1_BIB,
CR2_BIB,
# Other
TLD,
OLAC
]]
dld_fil.loc[dld_fil[CR1_ID] != dld_fil[CR2_ID]][[CR1_ID, CR2_ID]]
# ## Read European languages data
LANG_ID = "LangID"
eur_lang = pd.read_csv("../../data/general/european_languages.csv")
eur_lang
# +
# Language status codes
# L(iving)
# (e)X(tinct)
pd.unique(eur_lang["LangStatus"].ravel())
# +
# Number of extinct languages
eur_lang.loc[eur_lang["LangStatus"] == "X"]
# -
# ## Merge DLD with European languages
print(dld_fil.shape)
print(eur_lang.shape)
eur_dld = eur_lang.join(dld_fil.set_index(CR2_ID), on="LangID", how="inner")
print(eur_dld.shape)
# ## Clean data
eur_dld[[L1, PRINT_NAME, SIL, WP_INCUB, MAC_IN, MAC_OS, MS_IN, MS_OS, HUN_STAT, CR1_CH, CR1_FLOSS, CR1_UDHR, CR2_UDHR, CR1_BIB, OLAC]]
eur_dld_clean = eur_dld.copy()
# ### L1
# L1 is interpreted as object -> convert to int
eur_dld_clean[L1] = eur_dld[L1].astype(int)
# +
# Languages that no one speaks as mother tongue.
eur_dld_clean.loc[eur_dld_clean[L1] == 0].iloc[:, :15]
# -
# ### OS support
# +
# MAC OS support is given textual -> convert it to 0/1 (no data is also 0)
def apply_mac(x):
if x == "TRUE":
return 1
return 0
eur_dld_clean[MAC_OS] = eur_dld[MAC_OS].apply(lambda x: apply_mac(x))
# +
# Microsoft OS support is given textual -> convert it to 0/1 (no data is also 0)
def apply_win(x):
if x == "P" or x == "F":
return 1
return 0
eur_dld_clean[MS_OS] = eur_dld[MS_OS].apply(lambda x: apply_win(x))
# -
# ### UHHR & Bible
# +
# UDHR and Bible availability is given textual -> convert it to 0/1 (if there's any it's 1)
def true_id_not_empty(x):
if x == "-":
return 0
return 1
eur_dld_clean[CR1_UDHR] = eur_dld[CR1_UDHR].apply(lambda x: true_id_not_empty(x))
eur_dld_clean[CR2_UDHR] = eur_dld[CR2_UDHR].apply(lambda x: true_id_not_empty(x))
eur_dld_clean[CR1_BIB] = eur_dld[CR1_BIB].apply(lambda x: true_id_not_empty(x))
eur_dld_clean[CR2_BIB] = eur_dld[CR2_BIB].apply(lambda x: true_id_not_empty(x))
# -
# ### FLOSS spellchecker
# +
# FLOSS availability is given textual -> convert it to 0/1 (no data is also 0)
def apply_floss(x):
if x == "yes":
return 1
return 0
eur_dld_clean[CR1_FLOSS] = eur_dld[CR1_FLOSS].apply(lambda x: apply_floss(x))
eur_dld_clean[CR2_FLOSS] = eur_dld[CR2_FLOSS].apply(lambda x: apply_floss(x))
# -
# ### Fill NaN-s
missing = eur_dld.columns[eur_dld.isnull().any()]
print(missing)
pd.unique(eur_dld[MS_IN].values.ravel())
pd.unique(eur_dld[HUN_COV].values.ravel())
# Fill NaN with 0
eur_dld_clean[MS_IN] = eur_dld[MS_IN].fillna(0)
eur_dld_clean[HUN_COV] = eur_dld[HUN_COV].fillna(0)
missing = eur_dld_clean.columns[eur_dld_clean.isnull().any()]
print(missing)
# ### Type check
eur_dld.dtypes
eur_dld_clean.dtypes
# ## Save output
eur_dld_clean.to_csv("../../data/digital/dld_2012.csv", index=False)
# ## Investigate correlation
# +
NUM_COLS = [
L1,
L2,
WP_ART,
WP_SIZE_ADJ,
WP_SIZE_CH,
WP_INCUB,
WP_AVG_LEN,
WP_REAL_TOT,
MS_OS,
MS_IN,
MAC_OS,
MAC_IN,
HUN_COV,
CR1_CH,
CR2_CH,
CR1_WORDS,
CR2_WORDS,
CR1_DOCS,
CR2_DOCS,
CR1_UDHR,
CR2_UDHR,
CR1_FLOSS,
CR2_FLOSS,
CR1_BIB,
CR2_BIB,
OLAC
]
STATUS_COLS = [
ETHN,
ELP,
]
print(len(NUM_COLS + STATUS_COLS))
# -
eur_dld_clean[NUM_COLS]
# ### Correlation matrix of numerical values
f, ax = plt.subplots(figsize=(13, 10))
ax.set_title("Correlation between numeric variables")
corr = eur_dld_clean[NUM_COLS + STATUS_COLS].corr()
sns.heatmap(corr,
xticklabels=corr.columns,
yticklabels=corr.columns)
corr.style.background_gradient(cmap='coolwarm').format(precision=2)
# ### Correlation with endangerment status of Ethnologue
eur_dld_clean.set_index(LANG_ID).sort_index().plot(y=ETHN, figsize=(22, 10), title="Enthologue status", kind="bar")
# +
f, ax = plt.subplots(figsize=(13, 10))
ax.set_title("Correlation with degree and availability of digital tools - Ethnologue")
corr_ethn = corr[[ETHN]].sort_values(by=ETHN)
display(corr_ethn)
sns.heatmap(corr_ethn,
xticklabels=corr_ethn.columns,
yticklabels=corr_ethn.index)
# -
# ### Correlation with endangerment status of Endangered Languages Project
eur_dld_clean.set_index(LANG_ID).sort_index().plot(y=ELP, figsize=(22, 10), title="ELP status", kind="bar")
# +
f, ax = plt.subplots(figsize=(13, 10))
ax.set_title("Correlation with degree and availability of digital tools - ELP")
corr_elp = corr[[ELP]].sort_values(by=ELP)
display(corr_elp)
sns.heatmap(corr_elp,
xticklabels=corr_elp.columns,
yticklabels=corr_elp.index)
# -
# ## Fit model to find out most important factors
# +
def create_dataset(data, label_to_predict, test_size=0.2, random_state=0):
train, test = train_test_split(data, test_size=test_size, random_state=random_state)
y = train[label_to_predict]
y_true = test[label_to_predict]
del train[label_to_predict]
del test[label_to_predict]
return train, test, y, y_true
def print_pred(y_true, y_pred, title):
df = pd.DataFrame()
df["y_true"] = y_true
df["y_pred"] = y_pred
df.sort_index().plot(y=["y_true", "y_pred"], figsize=(22, 10), title=title, kind="bar")
def print_coefs(cols, coefs):
df = pd.DataFrame()
df["Features"] = cols
df["coefs"] = coefs
df["coefs_abs"] = abs(coefs)
display(df.sort_values(by="coefs_abs", ascending=False))
def print_score(y_true, y_pred, method):
print(method(y_true, y_pred))
def fit(scaler, model, train, y, test, y_true, coefs=False, title=None):
reg = make_pipeline(scaler, model)
reg.fit(train, y)
y_pred = reg.predict(test)
print_score(y_true, y_pred, explained_variance_score)
if coefs:
print_coefs(train.columns, model.coef_)
if title:
print_pred(y_true, y_pred, title)
return y_pred
# -
# ### Create dataset
# #### Predict Ethnologue status based on digitalization factors.
data = eur_dld_clean[NUM_COLS + [ETHN, LANG_ID]]
data.set_index("LangID", inplace=True)
train, test, y, y_true = create_dataset(data, ETHN, test_size=0.2, random_state=0)
print(train.shape)
print(test.shape)
lr = LinearRegression()
y_pred_lr = fit(Normalizer(), lr, train, y, test, y_true, coefs=True, title="Ethnologue status from digital advances")
data.loc["ykg"]
data.loc["nor"]
data.loc["kjh"]
# #### Predict Endangered Languages Project status based on digitalization factors.
data = eur_dld_clean[NUM_COLS + [ELP, LANG_ID]]
data.set_index(LANG_ID, inplace=True)
train, test, y, y_true = create_dataset(data, ELP, test_size=0.2, random_state=0)
print(train.shape)
print(test.shape)
lr = LinearRegression()
y_pred_lr = fit(Normalizer(), lr, train, y, test, y_true, coefs=True, title="Ethnologue status from digital advances")
pd.unique(eur_dld_clean[ELP].ravel())
eur_dld_clean.loc[eur_dld_clean[LANG_ID] == "gld"].iloc[:, :15]
eur_dld_clean.loc[eur_dld_clean[LANG_ID] == "eng"].iloc[:, :15]
# ## Conclusion
# Most important digital factors for determining endangerment status (for both metrics):
#
# - MAC and Windows OS level & input support
# - The existence of a spellchecker and their word coverage.
# - Ratio of real/total wikipedia pages.
# - Availability of Bible and Universal Declaration of Human rights translations
#
#
# Not so important featrues (for both metrics):
#
# - Number of speakers.
# - Character size of available resources.
| notebooks/Eszti/dld_european_lang.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# Please find jax implementation of this notebook here: https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/book1/15/gru_jax.ipynb
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/Nirzu97/pyprobml/blob/gru-torch/notebooks/gru_torch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7jysFtXPFE7E"
# # Gated Recurrent Units
#
# We show how to implement GRUs from scratch.
# Based on sec 9.1 of http://d2l.ai/chapter_recurrent-modern/gru.html
# This uses code from the [basic RNN colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/rnn_torch.ipynb).
#
# + colab={"base_uri": "https://localhost:8080/"} id="o6VNlv_FYTbS" outputId="373bc48e-0908-4e92-f286-136277d64065"
import numpy as np
import matplotlib.pyplot as plt
import math
from IPython import display
try:
import torch
except ModuleNotFoundError:
# %pip install torch
import torch
from torch import nn
from torch.nn import functional as F
from torch.utils import data
import collections
import re
import random
import os
import requests
import hashlib
import time
np.random.seed(seed=1)
torch.manual_seed(1)
# !mkdir figures # for saving plots
# + [markdown] id="phjRpyDNFT14"
# # Data
#
# As data, we use the book "The Time Machine" by <NAME>,
# preprocessed using the code in [this colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/text_preproc_torch.ipynb).
# + id="tLIKRJomvBV5"
class SeqDataLoader:
"""An iterator to load sequence data."""
def __init__(self, batch_size, num_steps, use_random_iter, max_tokens):
if use_random_iter:
self.data_iter_fn = seq_data_iter_random
else:
self.data_iter_fn = seq_data_iter_sequential
self.corpus, self.vocab = load_corpus_time_machine(max_tokens)
self.batch_size, self.num_steps = batch_size, num_steps
def __iter__(self):
return self.data_iter_fn(self.corpus, self.batch_size, self.num_steps)
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[1], reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ["<unk>"] + reserved_tokens
uniq_tokens += [token for token, freq in self.token_freqs if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
# + id="-TvZsL4gtHQ2"
def tokenize(lines, token="word"):
"""Split text lines into word or character tokens."""
if token == "word":
return [line.split() for line in lines]
elif token == "char":
return [list(line) for line in lines]
else:
print("ERROR: unknown token type: " + token)
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
def seq_data_iter_random(corpus, batch_size, num_steps):
"""Generate a minibatch of subsequences using random sampling."""
# Start with a random offset (inclusive of `num_steps - 1`) to partition a
# sequence
corpus = corpus[random.randint(0, num_steps - 1) :]
# Subtract 1 since we need to account for labels
num_subseqs = (len(corpus) - 1) // num_steps
# The starting indices for subsequences of length `num_steps`
initial_indices = list(range(0, num_subseqs * num_steps, num_steps))
# In random sampling, the subsequences from two adjacent random
# minibatches during iteration are not necessarily adjacent on the
# original sequence
random.shuffle(initial_indices)
def data(pos):
# Return a sequence of length `num_steps` starting from `pos`
return corpus[pos : pos + num_steps]
num_batches = num_subseqs // batch_size
for i in range(0, batch_size * num_batches, batch_size):
# Here, `initial_indices` contains randomized starting indices for
# subsequences
initial_indices_per_batch = initial_indices[i : i + batch_size]
X = [data(j) for j in initial_indices_per_batch]
Y = [data(j + 1) for j in initial_indices_per_batch]
yield torch.tensor(X), torch.tensor(Y)
def seq_data_iter_sequential(corpus, batch_size, num_steps):
"""Generate a minibatch of subsequences using sequential partitioning."""
# Start with a random offset to partition a sequence
offset = random.randint(0, num_steps)
num_tokens = ((len(corpus) - offset - 1) // batch_size) * batch_size
Xs = torch.tensor(corpus[offset : offset + num_tokens])
Ys = torch.tensor(corpus[offset + 1 : offset + 1 + num_tokens])
Xs, Ys = Xs.reshape(batch_size, -1), Ys.reshape(batch_size, -1)
num_batches = Xs.shape[1] // num_steps
for i in range(0, num_steps * num_batches, num_steps):
X = Xs[:, i : i + num_steps]
Y = Ys[:, i : i + num_steps]
yield X, Y
# + id="yRp-MH0Nv7rN"
def download(name, cache_dir=os.path.join("..", "data")):
"""Download a file inserted into DATA_HUB, return the local filename."""
assert name in DATA_HUB, f"{name} does not exist in {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split("/")[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, "rb") as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # Hit cache
print(f"Downloading {fname} from {url}...")
r = requests.get(url, stream=True, verify=True)
with open(fname, "wb") as f:
f.write(r.content)
return fname
def read_time_machine():
"""Load the time machine dataset into a list of text lines."""
with open(download("time_machine"), "r") as f:
lines = f.readlines()
return [re.sub("[^A-Za-z]+", " ", line).strip().lower() for line in lines]
def load_corpus_time_machine(max_tokens=-1):
"""Return token indices and the vocabulary of the time machine dataset."""
lines = read_time_machine()
tokens = tokenize(lines, "char")
vocab = Vocab(tokens)
# Since each text line in the time machine dataset is not necessarily a
# sentence or a paragraph, flatten all the text lines into a single list
corpus = [vocab[token] for line in tokens for token in line]
if max_tokens > 0:
corpus = corpus[:max_tokens]
return corpus, vocab
def load_data_time_machine(batch_size, num_steps, use_random_iter=False, max_tokens=10000):
"""Return the iterator and the vocabulary of the time machine dataset."""
data_iter = SeqDataLoader(batch_size, num_steps, use_random_iter, max_tokens)
return data_iter, data_iter.vocab
# + id="nG2T4maNYdsG"
DATA_HUB = dict()
DATA_URL = "http://d2l-data.s3-accelerate.amazonaws.com/"
DATA_HUB["time_machine"] = (DATA_URL + "timemachine.txt", "090b5e7e70c295757f55df93cb0a180b9691891a")
batch_size, num_steps = 32, 35
train_iter, vocab = load_data_time_machine(batch_size, num_steps)
# + [markdown] id="fKa87dOCFZ8H"
# # Creating the model from scratch
# + [markdown] id="gq3jJ8IQFjz3"
# Initialize the parameters.
# + id="WnwqslgzFW7E"
def get_params(vocab_size, num_hiddens, device):
num_inputs = num_outputs = vocab_size
def normal(shape):
return torch.randn(size=shape, device=device) * 0.01
def three():
return (
normal((num_inputs, num_hiddens)),
normal((num_hiddens, num_hiddens)),
torch.zeros(num_hiddens, device=device),
)
W_xz, W_hz, b_z = three() # Update gate parameters
W_xr, W_hr, b_r = three() # Reset gate parameters
W_xh, W_hh, b_h = three() # Candidate hidden state parameters
# Output layer parameters
W_hq = normal((num_hiddens, num_outputs))
b_q = torch.zeros(num_outputs, device=device)
# Attach gradients
params = [W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q]
for param in params:
param.requires_grad_(True)
return params
# + [markdown] id="xiMRAVr2Ftir"
# Initial state is a tensor of zeros of size (batch-size, num-hiddens)
# + id="rbMB5SnMFvkZ"
def init_gru_state(batch_size, num_hiddens, device):
return (torch.zeros((batch_size, num_hiddens), device=device),)
# + [markdown] id="9LcxOAdiF1kd"
# Forward function
# + id="SfC5HhBbFv6t"
def gru(inputs, state, params):
W_xz, W_hz, b_z, W_xr, W_hr, b_r, W_xh, W_hh, b_h, W_hq, b_q = params
(H,) = state
outputs = []
for X in inputs:
Z = torch.sigmoid((X @ W_xz) + (H @ W_hz) + b_z)
R = torch.sigmoid((X @ W_xr) + (H @ W_hr) + b_r)
H_tilda = torch.tanh((X @ W_xh) + ((R * H) @ W_hh) + b_h)
H = Z * H + (1 - Z) * H_tilda
Y = H @ W_hq + b_q
outputs.append(Y)
return torch.cat(outputs, dim=0), (H,)
# + id="Qw3vhoRKdVxt"
# Make the model class
# Input X to call is (B,T) matrix of integers (from vocab encoding).
# We transpse this to (T,B) then one-hot encode to (T,B,V), where V is vocab.
# The result is passed to the forward function.
# (We define the forward function as an argument, so we can change it later.)
class RNNModelScratch:
"""A RNN Model implemented from scratch."""
def __init__(self, vocab_size, num_hiddens, device, get_params, init_state, forward_fn):
self.vocab_size, self.num_hiddens = vocab_size, num_hiddens
self.params = get_params(vocab_size, num_hiddens, device)
self.init_state, self.forward_fn = init_state, forward_fn
def __call__(self, X, state):
X = F.one_hot(X.T, self.vocab_size).type(torch.float32)
return self.forward_fn(X, state, self.params)
def begin_state(self, batch_size, device):
return self.init_state(batch_size, self.num_hiddens, device)
# + id="bY_gAWKLw0vR"
def try_gpu(i=0):
"""Return gpu(i) if exists, otherwise return cpu()."""
if torch.cuda.device_count() >= i + 1:
return torch.device(f"cuda:{i}")
return torch.device("cpu")
# + [markdown] id="WtoCsGZ2F8G0"
# # Training and prediction
# + id="y1SRYmSN_cmx"
# + id="LDuvumDhgpvh"
def grad_clipping(net, theta):
"""Clip the gradient."""
if isinstance(net, nn.Module):
params = [p for p in net.parameters() if p.requires_grad]
else:
params = net.params
norm = torch.sqrt(sum(torch.sum((p.grad**2)) for p in params))
if norm > theta:
for param in params:
param.grad[:] *= theta / norm
# + id="tA_QeZH5rp1o"
class Animator:
"""For plotting data in animation."""
def __init__(
self,
xlabel=None,
ylabel=None,
legend=None,
xlim=None,
ylim=None,
xscale="linear",
yscale="linear",
fmts=("-", "m--", "g-.", "r:"),
nrows=1,
ncols=1,
figsize=(3.5, 2.5),
):
# Incrementally plot multiple lines
if legend is None:
legend = []
display.set_matplotlib_formats("svg")
self.fig, self.axes = plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [
self.axes,
]
# Use a lambda function to capture arguments
self.config_axes = lambda: set_axes(self.axes[0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# Add multiple data points into the figure
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return np.array(self.times).cumsum().tolist()
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
def set_axes(axes, xlabel, ylabel, xlim, ylim, xscale, yscale, legend):
"""Set the axes for matplotlib."""
axes.set_xlabel(xlabel)
axes.set_ylabel(ylabel)
axes.set_xscale(xscale)
axes.set_yscale(yscale)
axes.set_xlim(xlim)
axes.set_ylim(ylim)
if legend:
axes.legend(legend)
axes.grid()
def sgd(params, lr, batch_size):
"""Minibatch stochastic gradient descent."""
with torch.no_grad():
for param in params:
param -= lr * param.grad / batch_size
param.grad.zero_()
# + id="Eq_QMLUKhw3k"
def train_epoch(net, train_iter, loss, updater, device, use_random_iter):
state, timer = None, Timer()
metric = Accumulator(2) # Sum of training loss, no. of tokens
for X, Y in train_iter:
if state is None or use_random_iter:
# Initialize `state` when either it is the first iteration or
# using random sampling
state = net.begin_state(batch_size=X.shape[0], device=device)
else:
if isinstance(net, nn.Module) and not isinstance(state, tuple):
# `state` is a tensor for `nn.GRU`
state.detach_()
else:
# `state` is a tuple of tensors for `nn.LSTM` and
# for our custom scratch implementation
for s in state:
s.detach_()
y = Y.T.reshape(-1) # (B,T) -> (T,B)
X, y = X.to(device), y.to(device)
y_hat, state = net(X, state)
l = loss(y_hat, y.long()).mean()
if isinstance(updater, torch.optim.Optimizer):
updater.zero_grad()
l.backward()
grad_clipping(net, 1)
updater.step()
else:
l.backward()
grad_clipping(net, 1)
# batch_size=1 since the `mean` function has been invoked
updater(batch_size=1)
metric.add(l * y.numel(), y.numel())
return math.exp(metric[0] / metric[1]), metric[1] / timer.stop()
# + id="0tPGxQDWiqfl"
def train(net, train_iter, vocab, lr, num_epochs, device, use_random_iter=False):
loss = nn.CrossEntropyLoss()
animator = Animator(xlabel="epoch", ylabel="perplexity", legend=["train"], xlim=[10, num_epochs])
# Initialize
if isinstance(net, nn.Module):
updater = torch.optim.SGD(net.parameters(), lr)
else:
updater = lambda batch_size: sgd(net.params, lr, batch_size)
num_preds = 50
predict_ = lambda prefix: predict(prefix, num_preds, net, vocab, device)
# Train and predict
for epoch in range(num_epochs):
ppl, speed = train_epoch(net, train_iter, loss, updater, device, use_random_iter)
if (epoch + 1) % 10 == 0:
print(predict_("time traveller"))
animator.add(epoch + 1, [ppl])
print(f"perplexity {ppl:.1f}, {speed:.1f} tokens/sec on {str(device)}")
print(predict_("time traveller"))
print(predict_("traveller"))
# + id="MvGhZUIGd3F3"
def predict(prefix, num_preds, net, vocab, device):
"""Generate new characters following the `prefix`."""
state = net.begin_state(batch_size=1, device=device)
outputs = [vocab[prefix[0]]]
get_input = lambda: torch.tensor([outputs[-1]], device=device).reshape((1, 1))
for y in prefix[1:]: # Warm-up period
_, state = net(get_input(), state)
outputs.append(vocab[y])
for _ in range(num_preds): # Predict `num_preds` steps
y, state = net(get_input(), state)
outputs.append(int(y.argmax(dim=1).reshape(1)))
return "".join([vocab.idx_to_token[i] for i in outputs])
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="rw0uiJ4aF3jI" outputId="3161eba2-8ce7-47b3-c9c0-a25280a80316"
vocab_size, num_hiddens, device = len(vocab), 256, try_gpu()
num_epochs, lr = 500, 1
model = RNNModelScratch(len(vocab), num_hiddens, device, get_params, init_gru_state, gru)
train(model, train_iter, vocab, lr, num_epochs, device)
# + id="89IqhaoSwiRT"
class RNNModel(nn.Module):
"""The RNN model."""
def __init__(self, rnn_layer, vocab_size, **kwargs):
super(RNNModel, self).__init__(**kwargs)
self.rnn = rnn_layer
self.vocab_size = vocab_size
self.num_hiddens = self.rnn.hidden_size
# If the RNN is bidirectional (to be introduced later),
# `num_directions` should be 2, else it should be 1.
if not self.rnn.bidirectional:
self.num_directions = 1
self.linear = nn.Linear(self.num_hiddens, self.vocab_size)
else:
self.num_directions = 2
self.linear = nn.Linear(self.num_hiddens * 2, self.vocab_size)
def forward(self, inputs, state):
X = F.one_hot(inputs.T.long(), self.vocab_size)
X = X.to(torch.float32)
Y, state = self.rnn(X, state)
# The fully connected layer will first change the shape of `Y` to
# (`num_steps` * `batch_size`, `num_hiddens`). Its output shape is
# (`num_steps` * `batch_size`, `vocab_size`).
output = self.linear(Y.reshape((-1, Y.shape[-1])))
return output, state
def begin_state(self, device, batch_size=1):
if not isinstance(self.rnn, nn.LSTM):
# `nn.GRU` takes a tensor as hidden state
return torch.zeros((self.num_directions * self.rnn.num_layers, batch_size, self.num_hiddens), device=device)
else:
# `nn.LSTM` takes a tuple of hidden states
return (
torch.zeros((self.num_directions * self.rnn.num_layers, batch_size, self.num_hiddens), device=device),
torch.zeros((self.num_directions * self.rnn.num_layers, batch_size, self.num_hiddens), device=device),
)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="yDw7Ss_0F9MD" outputId="ba88ca30-63ea-457b-88a5-e602a0ff3c0a"
num_inputs = vocab_size
gru_layer = nn.GRU(num_inputs, num_hiddens)
model = RNNModel(gru_layer, len(vocab))
model = model.to(device)
train(model, train_iter, vocab, lr, num_epochs, device)
| notebooks/book1/15/gru_torch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="MBZ61NLJXKLY" outputId="c44c17fe-b150-4c92-edb1-b9e57e9c1b8e"
# !git clone https://github.com/onosyoono/Programming-Language-Classifier.git
# + id="ipBqMraZXXEl" colab={"base_uri": "https://localhost:8080/"} outputId="116c4b9e-571c-44bb-d22f-7d9e41283e2a"
# !ls
# + [markdown] id="nQpj0gxPZ-D7"
# Imports
# + id="b-yaOP9AYEZ4"
import numpy as np
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.neighbors import KNeighborsClassifier
# + [markdown] id="pujI5x9ma_da"
# Loading
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="KIlIChcAa-aL" outputId="511785b5-c681-4c86-e280-398e382291c7"
df = pd.read_csv("./Programming-Language-Classifier/dataset.csv")
df.head()
# + id="UkSRaNSqbHg4"
df = df.values
# + id="neCs7nqbbJxZ"
X, Y = df.T[0], df.T[1]
# + [markdown] id="cmo6KtwKbPnp"
# Vectorizing
# + colab={"base_uri": "https://localhost:8080/"} id="4Q3h7w_0bUJt" outputId="5be23673-d023-42e0-b1eb-564086267d3c"
vectorizer = CountVectorizer()
X_vec = vectorizer.fit_transform(X)
X_vec = X_vec.toarray()
print(X_vec.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="TJ7ismYTbZCn" outputId="1568041e-72d2-4ba2-92fe-8cb5f8abdb6d"
print("Dic Size:", len(vectorizer.get_feature_names()))
# + [markdown] id="UU8wRlYvbgil"
# Creating a KNN Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="sEmOsuTubfRE" outputId="f9a25ac9-355d-4a12-b438-b7df02652360"
neigh = KNeighborsClassifier(n_neighbors=5)
neigh.fit(X_vec[:int(0.8*len(Y))], Y[:int(0.8*len(Y))])
# + [markdown] id="4tRCqTt5cBDi"
# # Measuring Performance
# + [markdown] id="leeaKU14cH33"
# Training
# + colab={"base_uri": "https://localhost:8080/"} id="EdGtOqoNcTfp" outputId="c312e59d-356c-4387-ae54-7aae014487db"
neigh.score(X_vec[:int(0.8*len(Y))], Y[:int(0.8*len(Y))])
# + [markdown] id="avpK_u_ScM_2"
# Valdidation
# + colab={"base_uri": "https://localhost:8080/"} id="8k8XSWJAcUAR" outputId="d2077c1a-b549-4374-9a5a-a6518b4fa196"
neigh.score(X_vec[int(0.8*len(Y)):], Y[int(0.8*len(Y)):])
# + id="BdOHfZ7mczzL"
| knn_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
from sklearn_utils import *
from tensorflow_utils import *
import numpy as np
import pandas as pd
import tensorflow as tf
import nltk
import sklearn
from sklearn.cross_validation import train_test_split
from sklearn.pipeline import Pipeline
import matplotlib.pyplot as plt
import scipy
import math
import joblib
# -
# # Load and Split Kaggle Data
data_filename = '../data/train.csv'
data_df = pd.read_csv(data_filename)
corpus = data_df['Comment']
labels = data_df['Insult']
train_corpus, test_corpus, train_labels, test_labels = \
sklearn.cross_validation.train_test_split(corpus, labels, test_size=0.33)
# ## Build baseline text classification model in Sklearn
pipeline = Pipeline([
('vect', sklearn.feature_extraction.text.CountVectorizer()),
('tfidf', sklearn.feature_extraction.text.TfidfTransformer(sublinear_tf=True,norm='l2')),
('clf', sklearn.linear_model.LogisticRegression()),
])
# +
param_grid = {
#'vect__max_df': (0.5, 0.75, 1.0),
#'vect__max_features': (None, 5000, 10000, 50000),
'vect__ngram_range': ((1, 1), (2, 2), (1,4)), # unigrams or bigrams
#'vect_lowercase': (True, False),
'vect__analyzer' : ('char',), #('word', 'char')
#'tfidf__use_idf': (True, False),
#'tfidf__norm': ('l1', 'l2'),
#'clf__penalty': ('l2', 'elasticnet'),
#'clf__n_iter': (10, 50, 80),
'clf__C': [0.1, 1, 5, 50, 100, 1000, 5000],
}
model = cv (train_corpus, train_labels.values, 5, pipeline, param_grid, 'roc_auc', False, n_jobs=8)
# -
# Hold out set Perf
auc(test_labels.values,get_scores(model, test_corpus))
# This is about as good as the best Kagglers report they did.
joblib.dump(model, '../models/kaggle_ngram.pkl')
# # Score Random Wikipedia User Talk Comments
#
# Lets take a random sample of user talk comments, apply the insult model trained on kaggle and see what we find.
d_wiki = pd.read_csv('../../wikipedia/data/100k_user_talk_comments.tsv', sep = '\t').dropna()[:10000]
d_wiki['prob'] = model.predict_proba(d_wiki['diff'])[:,1]
d_wiki.sort('prob', ascending=False, inplace = True)
_ = plt.hist(d_wiki['prob'].values)
plt.xlabel('Insult Prob')
plt.title('Wikipedia Score Distribution')
_ = plt.hist(model.predict_proba(train_corpus)[:, 1])
plt.xlabel('Insult Prob')
plt.title('Kaggle Score Distribution')
# The distribution over insult probabilities in the two datasets is radically different. Insults in the Wikipedia dataset are much rarer
"%0.2f%% of random wiki comments are predicted to be insults" % ((d_wiki['prob'] > 0.5).mean() * 100)
# ### Check High Scoring Comments
for i in range(5):
print(d_wiki.iloc[i]['prob'], d_wiki.iloc[i]['diff'], '\n')
for i in range(50, 55):
print(d_wiki.iloc[i]['prob'], d_wiki.iloc[i]['diff'], '\n')
for i in range(100, 105):
print(d_wiki.iloc[i]['prob'], d_wiki.iloc[i]['diff'], '\n')
# # Score Blocked Users' User Talk Comments
d_wiki_blocked = pd.read_csv('../../wikipedia/data/blocked_users_user_talk_page_comments.tsv', sep = '\t').dropna()[:10000]
d_wiki_blocked['prob'] = model.predict_proba(d_wiki_blocked['diff'])[:,1]
d_wiki_blocked.sort('prob', ascending=False, inplace = True)
"%0.2f%% of random wiki comments are predicted to be insults" % ((d_wiki_blocked['prob'] > 0.5).mean() * 100)
# ### Check High Scoring Comments
for i in range(5):
print(d_wiki_blocked.iloc[i]['prob'], d_wiki_blocked.iloc[i]['diff'], '\n')
for i in range(50, 55):
print(d_wiki_blocked.iloc[i]['prob'], d_wiki.iloc[i]['diff'], '\n')
for i in range(100, 105):
print(d_wiki_blocked.iloc[i]['prob'], d_wiki.iloc[i]['diff'], '\n')
# # Scratch: Do not keep reading :)
# #### Tensorflow MPL
isinstance(y_train, np.ndarray)
y_train = np.array([y_train, 1- y_train]).T
y_test = np.array([y_test, 1- y_test]).T
# +
# Parameters
learning_rate = 0.001
training_epochs = 60
batch_size = 200
display_step = 5
# Network Parameters
n_hidden_1 = 100 # 1st layer num features
n_hidden_2 = 100 # 2nd layer num features
n_hidden_3 = 100 # 2nd layer num features
n_input = X_train.shape[1]
n_classes = 2
# tf Graph input
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
# Create model
def LG(_X, _weights, _biases):
return tf.matmul(_X, _weights['out']) + _biases['out']
# Store layers weight & bias
weights = {
'out': tf.Variable(tf.random_normal([n_input, n_classes]))
}
biases = {
'out': tf.Variable(tf.random_normal([n_classes]))
}
# Construct model
pred = LG(x, weights, biases)
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(pred, y)) # Softmax loss
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost) # Adam Optimizer
# Initializing the variables
init = tf.initialize_all_variables()
# Launch the graph
sess = tf.Session()
sess.run(init)
# Training cycle
for epoch in range(training_epochs):
avg_cost = 0.
m = 0
batches = batch_iter(X_train.toarray(), y_train, batch_size)
# Loop over all batches
for batch_xs, batch_ys in batches:
batch_m = len(batch_ys)
m += batch_m
# Fit training using batch data
sess.run(optimizer, feed_dict={x: batch_xs, y: batch_ys})
# Compute average loss
avg_cost += sess.run(cost, feed_dict={x: batch_xs, y: batch_ys}) * batch_m
# Display logs per epoch step
if epoch % display_step == 0:
print ("Epoch:", '%04d' % (epoch+1), "cost=", "{:.9f}".format(avg_cost/m))
correct_prediction = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
# Calculate accuracy
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print ("Accuracy:", accuracy.eval({x: X_train.toarray(), y: y_train}, session=sess))
print ("Accuracy:", accuracy.eval({x: X_test.toarray(), y: y_test}, session=sess))
print ("Optimization Finished!")
# Test model
# -
| misc/kaggle/src/n-grams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: limo
# language: python
# name: limo
# ---
# ## Functions as first-class objects
# In Python, functions are first-class objects.
#
# First-Class functions:
# "A programming language is said to have first class functions if it treats functions as first-class citizens."
#
# First-Class Citizen (Programming):
# "A first-class citizen (sometimes calles first-class object) is an entity which supports all the operations generally available to other entities, such as being being passes as an argument, returned from a function, and assigned to a variable."
# Assign function to variables
# +
def square(x):
return x * x
f = square(5) # Assign function *output* to variable
f = square # Assign function to variable (no (), as this triggers execution)
print(square)
print(f) # f points to same object as square
print(f.__name__)
f(5)
# -
# Use function as argument in function
print(map(lambda x: x ** 2, [1, 2, 3]))
filter(lambda x: x % 2 == 0, [1, 2, 3])
# +
def do_twice(fn, *args):
fn(*args)
fn(*args)
do_twice(print, "Hello world!")
# -
# Return function from function
# +
def create_adder(n):
def adder(m):
return n + m
return adder
add_5 = create_adder(5)
print(add_5.__name__)
add_5(10)
# -
# ## Attributes of user-defined functions
# +
def greeter(text="Hello"):
"""Print greeting text."""
print(text)
def printer(func):
print("Docstring:", func.__doc__)
print("Name:", func.__name__)
print("Attributes:", func.__dict__)
print("Code:", func.__code__)
print("Defaults:", func.__defaults__)
print("Globals:", func.__globals__.keys())
print("Closures:", func.__closure__)
printer(greeter)
# -
# - The global attribute points to the global namespace in which the function was defined.
# Some of these attributes return objects that have attributes themselves:
print(greeter.__doc__.upper())
print(greeter.__doc__.splitlines())
print(greeter.__code__.co_argcount)
print(greeter.__code__.co_code)
# ## Function scoping rules
#
# - Each time a function executes, a new local namespace is created that contans the function arguments and all variables that are assigned inside the function body.
# - Variables that are assigned inside the function body are local variables; those assigned outside the function body are global variables.
# - To resolve names, the interpreter first checks the local namespace, then the global namespace, and then the build-in namespace.
# - To change the value of a global variable inside the (local) function context, use the `global` declaration.
# - For nested functions, the interpreter resolves names by checking the namespaces of all enclosing functions from the innermost to the outermost, and then then global and built-in namespaces. To reassign the value of a local variable defined in an enclosing function use the `nonlocal` declaration.
# ### Use of `global`
#
# Because `a` is assigned inside the function body, it is a local variable in the local namespace of the `f` function and a completely separate object from the variable `a` defined in the global namespace.
# +
a = 5
def f():
a = 10
f()
a
# -
# `global` declares the `a` variable created inside the function body to be a global variable, making it reassing the previously declared global variable of the same name.
# +
a = 5
def f():
global a
a = 10
f()
a
# -
# ### Use of `nonlocal`:
# Because `n` reassigned inside `decrement`, it is a local variable of that function's namespace. But because `n -= 1` is the same as `n = n - 1` and Python first runs the right hand side of assignment statements, when running `n - 1` throws an error because there it is asked to reference the variable `n` that hasn't been assigned yet.
# +
def countdown(n):
def display():
print(n)
def decrement():
n -= 1
while n:
display()
decrement()
countdown(3)
# -
# `nonlocal` declares `n` to be a variable in the scope of an enclosing function of `decrement`.
# +
def countdown(n):
def display():
print(n)
def decrement():
nonlocal n
n -= 1
while n:
display()
decrement()
countdown(3)
# -
# ## Closures
#
# > A closure is the object that results from packaging up the statements that make up a function with the environment in which it executes. <NAME>, Python Essential Reference
#
# > A closure is a function that has access to variables that are neither global nor local (i.e. not defined in the function's body). <NAME>, Fluent Python
#
# > A closure is an inner function that remembers and has access to variables in the local scope in which it was created, even after the outer function has finished executing. - 'A closure closes over the free variables from the environment in which they were creates'. - A free variable is a variable defined inside the scope of the outer function that is not an argument in the inner function. Below, the 'free variable' is message. Stanford CS course
#
#
# How I think of them (not sure that's correct):
#
# - The closure attribute of a user-defined function is non-empty if the function is defined inside the scope of an outer function and references a local variable defined in the scope of the outer function. From the point of view of the inner function, said variable is a nonlocal variable.
#
# - A user-defined function is a closure if its closure attribute is non-empty.
#
#
#
#
#
# In the first case below, the inner function doesn't reference a nonlocal variable and thus isn't a closure. In the second case, it does reference a nonlocal variable, which gets stored in its `closure` attribute, which makes `inner` a closure.
# +
def outer():
x = 5
def inner():
print("Hello")
return inner
def outer2():
x = 5
def inner():
print(x)
return inner
g = outer()
print(g.__closure__)
g = outer2()
print(g.__closure__[0].cell_contents)
# -
# ## Mutable function parameters
# *Notes based on chapter 8 in [Fluent Python](https://www.oreilly.com/library/view/fluent-python/9781491946237/).*
# Functions that take mutable objects as arguments require caution, because function arguments are aliases for the passed arguments (i.e. they refer to the original object). This can cause unintended behaviour in two types of situations:
#
# - When setting a mutable object as default
# - When aliasing a mutable object passed to the constructor
# ### Setting a mutable object as default
# +
class HauntedBus:
def __init__(self, passengers=[]):
self.passengers = passengers
def pick(self, name):
self.passengers.append(name)
def drop(self, name):
self.passengers.remove(name)
bus1 = HauntedBus(["pete", "lara", "nick"])
bus1.drop("nick")
print(bus1.passengers)
bus2 = HauntedBus()
bus2.pick("heather")
print(bus2.passengers)
bus3 = HauntedBus()
bus3.pick("katy")
print(bus3.passengers)
# -
# Between bus 1 and 2, all works as intended, since we passed our own list when creating bus 1. Then things get a bit weird, though: how did Heather get into bus 3? When we define the `HauntedBus` class, we create a single empty list that is kept in the background and will be used whenever we instantiate a new bus without a custom passenger list. Importantly, all such buses will operate on the same list. We can see this by checking the object ids of the three buses' passenger lists:
assert bus1.passengers is not bus2.passengers
assert bus2.passengers is bus3.passengers
# This shows that while the passenger list of bus 1 and 2 are not the same object, the lists of bus 2 and 3 are. Once we know that, the above behaviour makes sense: all passenger list operations on buses without a custom list operate on the same list. Anohter way of seeing this by inspecting the default dict of `HauntedBus` after our operations abve.
HauntedBus.__init__.__defaults__
# The above shows that after the `bus3.pick('katy')` call above, the default list is now changed, and will be inherited by future instances of `HauntedBus`.
bus4 = HauntedBus()
bus4.passengers
# This behaviour is an example of why it matters whether we think of variables as boxes or labels. If we think that variables are boxes, then the above bevaviour doesn't make sense, since each passenger list would be its own box with its own content. But when we think of variables as labels -- the correct way to think about them in Python -- then the behaviour makes complete sense: each time we instantiate a bus without a custom passenger list, we create a new label -- of the form `name-of-bus.passengers` -- for the empty list we created when we loaded or created `HauntedBus`.
# What to do to avoid the unwanted behaviour? The solution is to create a new empty list each time no list is provided.
# +
class Bus:
def __init__(self, passengers=None):
if passengers is None:
self.passengers = []
else:
self.passengers = list(passengers)
def pick(self, name):
self.passengers.append(name)
def drop(self, name):
self.passengers.remove(name)
bus1 = Bus()
bus1.pick("tim")
bus2 = Bus()
bus2.passengers
# -
# ### Aliasing a mutable object argument inside the function
# The `init` method of the above class copies the passed passenger list by calling `list(passengers)`. This is critical. If, instead of copying we alias the passed list, we change lists defined outside the function that are passed as arguments, which is probably not what we want.
# +
class Bus:
def __init__(self, passengers=None):
if passengers is None:
self.passengers = []
else:
self.passengers = passengers
def pick(self, name):
self.passengers.append(name)
def drop(self, name):
self.passengers.remove(name)
team = ["hugh", "lisa", "gerd", "adam", "emily"]
bus = Bus(team)
bus.drop("hugh")
team
# -
# Again, the reason for this is that `self.passengers` is an alias for `passengers`, which is itself an alias for `team`, so that all operations we perfom on `self.passengers` are actually performed on `team`. The identity check below shows what the passengers attribute of `bus` is indeed the same object as the team list.
bus.passengers is team
# **To summarise:** unless there is a good reason for an exception, for functions that take mutable objects as arguments do the following:
#
# 1. Create a new object each time a class is instantiated by using None as the default parameter, rather than creating the object at the time of the function definition.
#
# 2. Make a copy of the mutable object for processing inside the function to leave the original object unchanged.
# ## Functional programming
# A programming paradigm built on the idea of composing programs of functions,
# rather than sequential steps of execution.
#
# Advantages of functional programming:
# - Simplifies debugging
# - Shorter and cleaner code
# - Modular and reusable code
#
# Memory and speed considerations:
# - Map/filter more memory efficient because they compute elements only when called, while list comprehension buffers them all.
# - Call to map/filter if you pass lambda comes with extra function overhead which list comprehension doesn't have, which makes the latter usually faster.
#
# See [here](https://docs.python.org/3/howto/functional.html) for more.
# ### Exercises
# *Mapping* is the process of applying a function elementwise to an array and storing the result. Apply the `len` function to each item in `a` and return a list of lengths.
a = ["a", "ab", "abc", "abcd"]
# Procedural approach.
# +
def lengths(items):
result = []
for item in items:
result.append(len(item))
return result
lengths(a)
# -
# List comprehension.
# +
def lengths(items):
return [len(item) for item in items]
lengths(a)
# -
# Functional approach.
# +
def lengths(items):
return list(map(len, items))
lengths(a)
# -
# *Filtering* is the process of extracting items from an iterable which satisfy a certain condition. Filter all elements of even length from `a`.
# Procedural approach.
# +
def even_lengths(items):
result = []
for item in items:
if len(item) % 2 == 0:
result.append(item)
return result
even_lengths(a)
# -
# List comprehension.
# +
def even_lengths(items):
return [item for item in items if len(item) % 2 == 0]
even_lengths(a)
# -
# Functional approach.
# +
def even_lengths(items):
return list(filter(lambda x: len(x) % 2 == 0, items))
even_lengths(a)
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ## Sources
#
# - [Fluent Python](https://www.oreilly.com/library/view/fluent-python/9781491946237/)
# - [Python Cookbook](https://www.oreilly.com/library/view/python-cookbook-3rd/9781449357337/)
# - [Python Essential Reference](https://www.oreilly.com/library/view/python-essential-reference/9780768687040/)
# - [Learning Python](https://www.oreilly.com/library/view/learning-python-5th/9781449355722/)
# - [Stanford CS41](https://stanfordpython.com/#/)
| content/post/python-functions/python-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="akTAhc5AnbiC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="54913da5-19ab-4615-e2ee-0f362ae5b85c" executionInfo={"status": "ok", "timestamp": 1583007810753, "user_tz": -60, "elapsed": 721, "user": {"displayName": "<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhIMcYMqvIWfwRoxNo9FxdnNmEIVhD-YU4dCMAhEg=s64", "userId": "08898779488445107126"}}
print ("Hello Github")
# + id="S9zLrfEyni9E" colab_type="code" colab={}
| HelloGithub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
sns.set_style('whitegrid')
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import cross_val_predict,cross_val_score
from sklearn.metrics import roc_auc_score,roc_curve
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import mean_squared_error
training_data = pd.read_csv("../Data/aps_failure_training_set.csv",na_values="na")
training_data.head()
# # Preprocessing
plt.figure(figsize=(20,12))
sns.heatmap(training_data.isnull(),yticklabels=False,cbar=False,cmap = 'viridis')
# # Missing value handling
# We are going to use different approches with missing values:
#
# 1. Removing the column having 80% missing values (**Self intuition)
# 2. Keeping all the features
# 3. Later, we will try to implement some feature engineering
#
#
# **For the rest of the missing values, we are replacing them with their mean() for now (**Ref)
# <big><b>Second Approach</b>
# +
sample_training_data = training_data
sample_training_data.fillna(sample_training_data.mean(),inplace=True)
#after replacing with mean()
plt.figure(figsize=(20,12))
sns.heatmap(sample_training_data.isnull(),yticklabels=False,cbar=False,cmap='viridis')
# +
#as all the other values are numerical except Class column so we can replace them with 1 and 0
sample_training_data = sample_training_data.replace('neg',0)
sample_training_data = sample_training_data.replace('pos',1)
sample_training_data.head()
# -
# # Testing Data preprocessing
testing_data = pd.read_csv("../Data/aps_failure_test_set.csv",na_values="na")
testing_data.head()
# +
sample_testing_data = testing_data
sample_testing_data.fillna(sample_testing_data.mean(),inplace=True)
#after replacing with mean()
plt.figure(figsize=(20,12))
sns.heatmap(sample_testing_data.isnull(),yticklabels=False,cbar=False,cmap='viridis')
# +
#as all the other values are numerical except Class column so we can replace them with 1 and 0
sample_testing_data = sample_testing_data.replace('neg',0)
sample_testing_data = sample_testing_data.replace('pos',1)
sample_testing_data.head()
# -
from sklearn.feature_selection import SelectFromModel
from sklearn.ensemble import RandomForestClassifier
rf= RandomForestClassifier(random_state=42)
X = sample_training_data.drop('class',axis=1)
y = sample_training_data['class']
features = X.columns.values
features.tolist()
rf.fit(X,y)
rf.feature_importances_
model = SelectFromModel(rf, prefit=True)
# +
X_new = model.transform(X)
# -
X=X_new
X.shape
#Print the chosen features
features = np.array(features.tolist())
print(features[model.get_support()])
testData_X = sample_testing_data.drop('class',axis=1)
testData_y = sample_testing_data['class']
#Test data transformation
newtestdata=testData_X.loc[:, features[model.get_support()]]
newtestdata.head()
# # Test data implementation
from sklearn.ensemble import RandomForestClassifier
rf= RandomForestClassifier(n_estimators=380,max_features='log2',random_state=42,oob_score=True,warm_start=True)
rf.fit(X,y)
testDataPrediction = rf.predict(newtestdata)
print(classification_report(testData_y,testDataPrediction))
print(metrics.accuracy_score(testData_y, testDataPrediction))
print(metrics.r2_score(testData_y, testDataPrediction))
print(metrics.f1_score(testData_y, testDataPrediction))
#testing error
print(metrics.mean_squared_error(testData_y, testDataPrediction))
#Training error
temp = rf.predict(X)
mean_squared_error(y,temp)
#confusion matrix
print(confusion_matrix(testData_y, testDataPrediction))
tn, fp, fn, tp = confusion_matrix(testData_y, testDataPrediction).ravel()
confusionData = [[tn,fp],[fn,tp]]
pd.DataFrame(confusionData,columns=['FN','FP'],index=['TN','TP'])
#without modified threshold
cost = 10*fp+500*fn
values = {'Score':[cost],'Number of Type 1 faults':[fp],'Number of Type 2 faults':[fn]}
pd.DataFrame(values)
# +
#with different threshold
THRESHOLD = 0.02 #optimal one chosen manually
thresholdPrediction = (rf.predict_proba(newtestdata)[:,1] >= THRESHOLD).astype(bool)
tn, fp, fn, tp = confusion_matrix(testData_y,thresholdPrediction).ravel()
cost = 10*fp+500*fn
values = {'Score':[cost],'Number of Type 1 faults':[fp],'Number of Type 2 faults':[fn]}
pd.DataFrame(values)
# -
# # Final Score is 10810
# # Confusion Matrix
# +
from sklearn.metrics import confusion_matrix
import pandas as pd
import seaborn as sn
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
y_true = ["Pos", "Neg"]
y_pred = ["Pos", "Neg"]
data = confusion_matrix(testData_y, thresholdPrediction)
df_cm = pd.DataFrame(data, columns=np.unique(y_true), index = np.unique(y_true))
df_cm.index.name = 'Actual'
df_cm.columns.name = 'Predicted'
plt.figure(figsize = (10,7))
sn.set(font_scale=1.4)#for label size
sn.heatmap(df_cm, cmap="Blues", annot=True,annot_kws={"size": 16},fmt='g')# font size
# +
#precision Score
preScore= tp/(tp+fp)
preScore
# +
#Recall Score
reScore = tp/(tp+fn)
reScore
# +
#Information (Precision-Recall)
#high precision relates to a low false positive rate,
#and high recall relates to a low false negative rate.
#High scores for both show that the classifier is returning accurate results (high precision),
#as well as returning a majority of all positive results (high recall).
# -
def evaluationScored(y_test,prediction):
acc = metrics.accuracy_score(y_test, prediction)
r2 = metrics.r2_score(y_test, prediction)
f1 = metrics.f1_score(y_test, prediction)
mse = metrics.mean_squared_error(y_test, prediction)
values = {'Accuracy Score':[acc],'R2':[r2],'F1':[f1],'MSE':[mse]}
print("\n\nScores")
print (pd.DataFrame(values))
evaluationScored(testData_y,thresholdPrediction)
| Saraf/FINAL-Part-2-checkpoint -- RF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": true}}}}
# %matplotlib inline
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": true}}}}
import seaborn as sns
from ipywidgets import interact
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": true}}}}
tips = sns.load_dataset('tips')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": false}}}}
tips.head()
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": false}}}}
@interact(hue=['smoker', 'sex', 'time', 'day'])
def plots(hue):
_ = sns.pairplot(tips, hue=hue)
# +
# %load_ext watermark
# python, ipython, packages, and machine characteristics
# %watermark -v -m -p seaborn,ipywidgets,matplotlib
| basics/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" tags=[]
# # Format conversion
# + [markdown] kernel="SoS" tags=[]
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
# * `sos convert file.ipynb file.sos` converts notebook to sos script
# * `sos convert file.sos file.ipynb` converts sos script to sos notebook
#
# + [markdown] kernel="SoS" tags=[]
# ## SoS -> ipynb
# + [markdown] kernel="SoS" tags=[]
# You can convert an existing SoS script to the `.ipynb` format using command
# ```
# $ sos convert myscript.sos myscript.ipynb
# ```
# + [markdown] kernel="SoS" tags=[]
# ## ipynb -> SoS
# + [markdown] kernel="SoS" tags=[]
# A Jupyter notebook can contain markdown cell and code cell with different kernels, and a sos cell might or might not contain a real sos step (with section header).
#
# You can save a Jupyter notebook with SoS kernel to a SoS script using `File -> Download As -> SoS` from the browser, or using command
#
# ```
# $ sos convert myscript.ipynb myscript.sos
# ```
#
# The conversion process will export only the embeded workflow to `.sos` file, ignoring all other content of the notebook.
# + [markdown] kernel="SoS" tags=[]
# ## ipynb -> HTML
#
# Command `sos convert my.ipynb my.html --template` essentially calls `jupyter nbconvert --to html` to convert notebook to HTML format, with additional templates provided by SoS Notebook.
#
# SoS provides the following templates
#
# | template | code highlighting | TOC | Hide Cell | Suit for |
# | -- | -- | -- | -- | --|
# | `sos-full` | jupyter | no | no | static short report |
# | `sos-cm` | SoS CodeMirror | no | no | output similar to notebook interface |
# | `sos-report` | jupyter | no | yes | static report with hidden details |
# | `sos-full-toc` | jupyter | yes | no | static long report |
# | `sos-cm-toc` | SoS CodeMirror | yes | none | output of long notebook with notebook interface |
# | `sos-report-toc` | jupyter | yes | yes | long report with hidden details |
#
# Where:
# * **code highlighting**: Jupyter can hilight sos source code using a static syntax hilighter. The output is lightweight but not as nice as the codemirror highlighter.
# * **TOC**: Automatically generate a table of content to the left of the page.
# * **Hide Cell**: HTML page by default only displays only markdown and selected ouptput cells (cells with `report_cell` tag, output of cells with `report_output` tag). A control panel to the left top corner of the page can be used to show all content
# + [markdown] kernel="SoS" tags=[]
# ## ipynb -> pdf
# This command essentially calls command `jupyter nbconvert --to pdf` to convert notebook to PDF format.
# + [markdown] kernel="SoS" tags=[]
# ## ipynb -> md
# This command essentially calls command `jupyter nbconvert --to markdown` to convert notebook to Markdown format.
# + [markdown] kernel="SoS" tags=[]
# ## ipynb -> ipynb
# + [markdown] kernel="SoS" tags=[]
# This command converts a Jupyter notebook in another kernel to a SoS notebook, with the original kernel language as the language of each code cell.
#
# If the original notebook has kernel `python3`, an option `--python3-to-sos` can be used to convert code cells to `SoS`.
#
# This converter will copy the input notebook to output if the notebook is already a SoS notebook. However, if an option `--inplace` is specified, it will overwrite the original notebook with the converted one.
#
# Note that if you already have a non-SoS notebook opened in Jupyter, you can simply use
#
# `Kernel` -> `Change kernel` -> `sos`
#
# to convert the kernel to SoS. You can then use the global language selector to select the appropriate default langauge for the notebook and re-execute the notebook to set the language to each cell.
# + [markdown] kernel="SoS" tags=[]
# ## SoS -> HTML
# + [markdown] kernel="SoS" tags=[]
# The `sos` to `html` converter converts `.sos` script to HTML format. It can be either written to a HTML file, or to standard output if option `--to html` is specified without a destination filename.
# + [markdown] kernel="SoS" tags=[]
# The converter also accepts a number of parameters (as shown above). The `raw` parameter adds a URL to filename in the HTML file so that you can link to the raw `.sos` file from the `.html` output. The `linenos` adds line numbers, and `style` allows you to choose from a number of pre-specified styles. Finally, the `view` option would open the resulting HTML file in a browser.
#
# For example,
# ```
# sos convert ../examples/update_toc.sos --to html --view --style xcode
# ```
# would show a HTML file as
#
# 
# + [markdown] kernel="SoS" tags=[]
# ## Further reading
#
# * [The `%convert` magic](sos_magics.html)
| src/user_guide/convert.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# + [markdown] id="I1n9GDwre1ZS"
# # Группировка табличных данных
# ---
#
# *Сделали с любовью студенты БЭК181:*
#
# **<NAME>**
#
# **<NAME>**
#
# **<NAME>**
# + [markdown] id="l2fbikCExT_J"
# ## Вместо предисловия
# ---
#
# В данном туториале __для начала__ мы показываем как установить Julia и все необходимые для работы с таблицами пакеты. __В основной части__ демонстрируется ряд операций для работы с группировкой табличных данных, выбора данных в сгруппированных датасетах, а также применение различных функций для преобразования данных.
#
# Помимо этого, __в бонусной части__ мы показали как можно быстро визуализировать данные из сгруппированного датасета.
# + [markdown] id="gROm13IegH4j"
# ## Основные источники
# ---
#
# * https://datatofish.com/add-julia-to-jupyter/ - *Подключение Julia к Jupyter notebook*
# * https://dataframes.juliadata.org/v0.14/man/split_apply_combine.html - _Основной источник и документация_
# * https://dataframes.juliadata.org/stable/man/split_apply_combine/ - *Новая версия тех же операций (по ссылке выше уже нет функции by)*
# * https://github.com/nalimilan/FreqTables.jl - *frequency table*
# * https://github.com/bkamins/DataFrames.jl/blob/main/docs/src/man/reshaping_and_pivoting.md - *pivot table*
# * https://www.queryverse.org/VegaLite.jl/stable/gettingstarted/tutorial/ - **bonus with graphics**
#
# + [markdown] id="qDAZt7-Zfm1R"
# ## Установка Julia и необходимых пакетов
# ---
# + [markdown] id="B8_hp5otShP_"
# ### Google Colaboratory
# ---
# + id="GIeFXS0F0zww"
# %%shell
set -e
#---------------------------------------------------#
JULIA_VERSION="1.5.2" # any version ≥ 0.7.0
JULIA_PACKAGES="IJulia BenchmarkTools Plots CSV Econometrics DataFrames RDatasets"
JULIA_PACKAGES_IF_GPU="CuArrays"
JULIA_NUM_THREADS=2
#---------------------------------------------------#
if [ -n "$COLAB_GPU" ] && [ -z `which julia` ]; then
# Install Julia
JULIA_VER=`cut -d '.' -f -2 <<< "$JULIA_VERSION"`
echo "Installing Julia $JULIA_VERSION on the current Colab Runtime..."
BASE_URL="https://julialang-s3.julialang.org/bin/linux/x64"
URL="$BASE_URL/$JULIA_VER/julia-$JULIA_VERSION-linux-x86_64.tar.gz"
wget -nv $URL -O /tmp/julia.tar.gz # -nv means "not verbose"
tar -x -f /tmp/julia.tar.gz -C /usr/local --strip-components 1
rm /tmp/julia.tar.gz
# Install Packages
if [ "$COLAB_GPU" = "1" ]; then
JULIA_PACKAGES="$JULIA_PACKAGES $JULIA_PACKAGES_IF_GPU"
fi
for PKG in `echo $JULIA_PACKAGES`; do
echo "Installing Julia package $PKG..."
julia -e 'using Pkg; pkg"add '$PKG'; precompile;"'
done
# Install kernel and rename it to "julia"
echo "Installing IJulia kernel..."
julia -e 'using IJulia; IJulia.installkernel("julia", env=Dict(
"JULIA_NUM_THREADS"=>"'"$JULIA_NUM_THREADS"'"))'
KERNEL_DIR=`julia -e "using IJulia; print(IJulia.kerneldir())"`
KERNEL_NAME=`ls -d "$KERNEL_DIR"/julia*`
mv -f $KERNEL_NAME "$KERNEL_DIR"/julia
echo ''
echo "Success! Please reload this page and jump to the next section."
fi
# + [markdown] id="ZuS2LY_MSnXG"
# ### Другие устройства
# ---
# + id="GFUnXLzZStEF"
begin
import Pkg; Pkg.add("RDatasets")
Pkg.add("Econometrics")
Pkg.add("GLM")
Pkg.add("CSV")
Pkg.add("DataFrames")
Pkg.add("Statistics")
Pkg.add("FreqTables")
Pkg.add("VegaLite")
Pkg.add("VegaDatasets")
end
# + [markdown] id="SDVsu5LLTb8p"
# ### Проверка успешной установки
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="EEzvvzCl1i0F" outputId="3c533e77-ee6c-4dca-803f-cf8c0e392d72"
versioninfo()
# + colab={"base_uri": "https://localhost:8080/"} id="yQlpeR9wNOi8" outputId="9050a3ad-a488-4842-bead-7cfad596326f"
exp(1) #проверка на работоспособность среды выполнения
# + [markdown] id="P9Lu1vk4galg"
# ## Датасет
# ---
# + id="0HNqxjjebg-1"
using CSV, Econometrics, Statistics
# + id="3bUX8t5bbWmI"
using RDatasets
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="YfHYE2cEdxUJ" outputId="64b72cd2-62ba-4f4b-d981-7f2af2be39e7"
psid = dataset("Ecdat", "PSID")
first(psid, 5)
# + colab={"base_uri": "https://localhost:8080/", "height": 258} id="R49fmPyZiXS7" outputId="91ef40ce-28ae-4592-f651-c4c3e62a07f4"
describe(psid)
# + [markdown] id="gr7GT9j-j_UH"
# Описание переменных:
#
# * IntNum - номер интервью
# * PersNum - персональный номер
# * Age - возраст
# * Educatn - количество лет обучения (получения образования)
# * Hours - количество часов работы в год
# * Married - семейный статус
#
# Более подробно смотреть здесь: https://www.picostat.com/dataset/r-dataset-package-ecdat-psid
# + [markdown] id="pbPHTf04lCJZ"
# ## Операции с данными
# ---
# + [markdown] id="5WJ3lLqQnrzH"
# ### `groupby(df, :variable)`
#
#
# ---
#
#
#
#
# Делит датасет **df** на несколько датасетов, в каждом из которых категориальный признак **variable** одинаковый
#
# Создается датасет из датасетов, далее в объяснениях __gdf__ — сгруппированный датасет
#
# Сгруппируем наш датасет по переменной __Married__:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="xpZ21vqyihbX" outputId="0b483f97-dc29-4d56-9220-fbedd6ab03be"
groupby_married_psid = groupby(psid, :Married)
# + [markdown] id="X8nKal78oG-D"
# Можно вытащить каждый i-ый датасет, вызвав команду **gdf[i]**
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="hlmDBttGlbuv" outputId="2f43618c-a2c1-40c0-9a01-4bb542f1236a"
married_psid = groupby_married_psid[1]
first(married_psid, 5)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="7JrQEoBKBBjK" outputId="0f5102cc-40d7-4e6a-9bca-22b546a86b90"
widowed_psid = groupby_married_psid[3]
first(widowed_psid, 5)
# + [markdown] id="Or_jdndo-ZFA"
# Или же можно вызвать, прописав условие `gdf[(variable='name'),]`
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="95DiU0W0-T4G" outputId="79877942-65ca-4550-f93e-ef1d7c20dcd6"
first(groupby_married_psid[(Married="married",)], 5)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="XpmYQ7vL-4G9" outputId="6d985d79-ac85-49b1-c4dc-d653c9e6f5b4"
first(groupby_married_psid[(Married="widowed",)], 5)
# + [markdown] id="iklUojmEonWe"
# ### __combine__
# + [markdown] id="XyP7C2xgr3kY"
# #### `combine(gdf, nrow)`
# ---
# + [markdown] id="nx21cz3dp6Ni"
# Можно вывести общее число строк в каждом датасете, используя команду **nrow**
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="EZsmXF0Cpp_w" outputId="10a88417-bee7-4455-8438-7db2ad827e48"
combine(groupby_married_psid, nrow)
# + [markdown] id="DeOVs94Wr9oN"
# #### `combine(gdf, :variable => operation => :name)`
# ---
# + [markdown] id="rd78R1t4pQqW"
# Выдает таблицу, где для каждого датасета **gdf[i]** в столбце с названием **name** показано значение признака **variable** после операции **operation**
# + [markdown] id="dFhl-FdBuaaa"
# Математические операции из пакета **Statistics**:
# * mean - среднее значение
#
# * median - медиана
#
# * sum - сумма значений
#
# * std - стандартное отклонение
# + [markdown] id="hD5KknJVvf9n"
# По дефолту название столбца соотвествует **variable_operation**
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="CzwEEuUooCsg" outputId="81867403-8c8a-4637-f974-944565cfff3e"
combine(groupby_married_psid, :Age => mean)
# + [markdown] id="hqdsYci2zvQ9"
# Однако его можно переименовать, для этого после операции необходимо прописать `=> :new_name`
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="goyFV0Aqp5Tv" outputId="2f21afd1-bb66-4563-f8a1-704828685884"
combine(groupby_married_psid, :Age => mean => :mean, nrow)
# + [markdown] id="eEhcSYTNzP6c"
# Аналогично выводить значения операций над признаками можно делать сразу для нескольких столбцов.
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="dEU0tzE3otdA" outputId="2a9b88dc-c248-4e22-88cd-bdb19c2265eb"
combine(groupby_married_psid, :Age => mean, :Kids =>median)
# + [markdown] id="S5TxX7MHyN-6"
# #### `combine(gdf, x -> operation(operation(vab1), operation(vab2)))`
# ---
# + [markdown] id="ZR-3pPVwyaA_"
# Делает операции над признаками **vab1, vab2** и выводит их значения для каждого датасета **gdf[i]**
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="w6Yrb-cVxIWr" outputId="45666ec0-2254-4270-cf8f-c3360a5ae666"
combine(groupby_married_psid, x -> std(x.Hours) / std(x.Age))
# + [markdown] id="0yICbexpH31y"
# #### `combine(gdf) do df (operations) end`
# ---
# + [markdown] id="HBeIJedyHYmZ"
# Можно использовать оператор **do**:
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="3akYt4kbHcmq" outputId="94ad7546-fe7a-430b-f0ee-89ae41de1cc0"
combine(groupby_married_psid) do df
(m = mean(df.Hours), s² = var(df.Hours))
end
# + [markdown] id="j5YTmRhTIVYv"
# #### `combine(gdf, AsTable([vab1, vab2]) => x ->operations))`
# ---
# + [markdown] id="s7GAk8PCypN4"
# Результат такой же, написано по-другому:
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="I9mk3IG3qsI4" outputId="e0f49e58-9ee8-466a-da4f-63bfe228fa78"
combine(groupby_married_psid,
AsTable([:Hours, :Age]) =>
x -> std(x.Hours) / std(x.Age))
# + [markdown] id="YzLRXHK1v7vZ"
# #### `combine(gdf, [:vab1, :vab2] => ((p,s) ->(a = operations(p,s), b=operations(p,s))) => AsTable)`
# ---
# + [markdown] id="hIrqQA-FvtEf"
# Данная операция применима в том случае, если операции над признаками происходят совместно, тогда:
#
# **vab1, vab2** - признаки
#
# **operations** - математические операции из пакета Statistics
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="yR0iU9XeqR92" outputId="06789cf6-21cb-4e1a-bd08-814ea86a1de9"
combine(groupby_married_psid, [:Age, :Kids] => ((p, s) -> (a=mean(p)/mean(s), b=sum(p))) =>
AsTable)
# + [markdown] id="7v779PTaxiWB"
# #### `combine(gdf, 1:2 => cor, nrow)`
# ---
# + [markdown] id="OBPNvq9yxlS6"
# Выводит корреляции между признаками **vab1, vab2** для каждого из датасетов **gdf[i]**, и показывает количество элементов в них
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="JrMuFo_xxQmJ" outputId="349ab982-0a39-45a7-9d98-7f76e450077d"
combine(groupby_married_psid, 1:2 => cor, nrow)
# + [markdown] id="oHnyNilxzfp6"
# ### __select__
#
# + [markdown] id="MhHSQSa9fG9U"
# #### `select(df, x)`
# ---
# Позволяет выбрать из всего датафрейма одну колонку с номером x. Если необходимо выбрать колонку по ее названию, то можно воспользоваться `select(df, :name)`
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="UFof7F4lfcxs" outputId="5923d0b9-3ee3-411d-9924-1a4118bf71da"
first(select(married_psid, :Earnings), 5)
# + [markdown] id="vhw1tLiBibgM"
# #### `select(df, Not(:name))`
# ---
# Выбор всего датафрейма за исключением определенной колонки
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="MV8EZIYWkPJT" outputId="c6a60626-2efd-47e7-f61d-c4a3daef1f0c"
first(select(married_psid, Not(:Earnings)), 5)
# + [markdown] id="HJA81Zd4ka1_"
# #### `select(gd, :c => operation, nrow)`
# ---
# К сгруппированному датасету применяется операция по колонке
#
# + colab={"base_uri": "https://localhost:8080/", "height": 292} id="7bAz-uIok68h" outputId="96cc3c1e-053d-4cda-afeb-8c951a18c283"
first(select(groupby_married_psid, :Kids => sum), 10) # суммирование количества детей по катеогории Married
# + [markdown] id="tdrQOIGnnGYB"
# Такую же операцию можно применить, разделив сгруппированные датасеты с помощью `ungroup=False`, а затем вывести определенный датасет с помощью [number]
# + colab={"base_uri": "https://localhost:8080/", "height": 722} id="BByUpRZynSCI" outputId="ca715068-6797-4ebc-ba8f-f39a72550d16"
select(groupby_married_psid, :Kids => sum, ungroup=false)[1]
# + [markdown] id="r2OOd5oRpeqG"
# #### `select(df, :a => (x -> operation) => :c, [:b, :f])`
# ---
# Выбор колонки а и применение к ней определенной операции, после чего ее переименование в с. Вывод преобразованной колонки и добавление к ней колонок b и f
#
# _Для примера переведем часы из исходного датасета в дни, поделив значения на 24, соответственно и переименуем колонку_
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="nf2_wb60pfLp" outputId="945e738b-7542-4971-a79c-d9f12027d1f5"
first(select(married_psid, :Hours => (x -> /(x, 24)) => :Days, [:Educatn, :Age]), 5)
# + [markdown] id="6dNsPy5MDtdH"
# #### `select(df, x:y => cor)`
# ---
# Позволяет вывести корреляцию между значениями столбцов x и y в df
#
# Команда `select` возвращает датафрейм с тем же количеством строк и втом же их порядке, что и в исходном датасете. Это работает даже в том случае, если в исходном сгруппированном датасете они были переупорядочены
# + [markdown] id="EYmgDABOE01e"
# Найдем корреляцию между уровнем образования и доходами среди женатых людей, выведем первые 5 строк
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="luAjuX8xqAVY" outputId="6fe9be7e-7858-44eb-c0e2-0e755083a8eb"
first(select(married_psid, 4:5 => cor), 5)
# + [markdown] id="Lzk4bBQrFRib"
# То же можно применить и к сгруппированному датасету, в таком случае помимо порядкового номера на выходе получим колонку с категорией
# + colab={"base_uri": "https://localhost:8080/", "height": 292} id="QaIXFR_FFKmd" outputId="0ac69b84-e9da-4e43-c8e8-4fbd18eeb93e"
select_cor = select(groupby_married_psid, 1:2 => cor)
first(select_cor, 10)
# + [markdown] id="lrxfhZLDqYj5"
# ### `freqtable(gdf, :x, :y)`
# ---
#
#
#
# + [markdown] id="r7N6eF0KvE_n"
# С группированными данными еще можно сделать таблицу частот по 2 признакам
#
# Для этого нужно использовать отдельный пакет **FreqTables**
# + id="Es937-eN_c7G"
import Pkg; Pkg.add("FreqTables")
# + id="HnMLdQec_tFH"
using FreqTables
# + colab={"base_uri": "https://localhost:8080/"} id="vgKirq3s9Fyp" outputId="a7f803d4-b522-4fca-a14e-a5f9a2bad462"
freqtable(psid, :Married, :Kids) #мне не нравится этот датасет
# + colab={"base_uri": "https://localhost:8080/"} id="0XBD2DV7FKuJ" outputId="66cd708b-5279-4b4a-d6fa-9d965d8077de"
cars = dataset("datasets", "mtcars")
freqtable(cars, :Model, :Cyl)
# + [markdown] id="EFT8v8KvGf-g"
# ### `pivot table`
# ---
# + [markdown] id="spiguIniGmss"
# Используя метод **combine** и другие основные операции с таблицами (stack, unstack), можно вывести таблицу со средними величинами по каждому признаку для каждой категории
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="1xmc6BN2Erqj" outputId="b7fc8496-dfe5-4b51-edb5-1f35e9b65ac9"
d = stack(psid, Not(:Married))
x = combine(groupby(d, [:variable, :Married]), :value => mean => :vsum)
unstack(x, :Married, :vsum)
# + [markdown] id="1KcgSTr7J-mh"
# ### *__Бонус__: красивый график с группированными данными
# ---
# + id="K2e7QWjhKYxe"
import Pkg; Pkg.add("VegaLite")
# + id="xgIHBMHfP0x8"
using RDatasets, VegaLite
iris = dataset("datasets", "iris")
# + colab={"base_uri": "https://localhost:8080/", "height": 468} id="2KaMfzRWFRRs" outputId="a677d1ff-a683-461d-9852-65517f2be3b7"
iris |> @vlplot(
:point,
x=:PetalLength,
y=:PetalWidth,
color="Species:n",
width = 400,
height=400
)
# + [markdown] id="inklZ5d5T2_b"
# ### ** __Бонус__: еще более красивые графики
# ---
#
# + [markdown] id="EzxVQ9WjaYeo"
# Так как пакеты **VegaDatasets, RDatasets** конфликтуют, поэтому используем VegaDatasets.dataset
# + id="8Ll_bcs1aMy3"
import Pkg; Pkg.add("VegaLite")
# + id="qU1N6DQLRv7-"
import Pkg; Pkg.add("VegaDatasets")
# + colab={"base_uri": "https://localhost:8080/", "height": 385} id="9ikk6RONOy5X" outputId="e4198ade-71f3-4008-bbd1-f7f655c230ea"
using VegaDatasets, VegaLite
data = VegaDatasets.dataset("cars")
# + colab={"base_uri": "https://localhost:8080/", "height": 263} id="eHV7VbfjK-2Y" outputId="efc914c7-b42b-4f5a-cad3-254af047ce2b"
data |>
@vlplot(:point, x=:Miles_per_Gallon, y=:Horsepower, color="Cylinders:n")
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="vSHMMP52Rpdf" outputId="53a6a075-9c07-40af-e505-182a26db4b77"
data |>
@vlplot(:point, x=:Miles_per_Gallon, y=:Horsepower, column=:Origin, color=:Cylinders)
# + id="2gykxKpIb1Uc"
| Groping data/Grouping_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ** Números primos **
# ## <NAME>
def primo():
c = [datos for datos in range(2,100)
if (datos%2!=0 or datos==2) and (datos%3!=0 or datos==3) and (datos%5!=0 or datos ==5) and (datos%7!=0 or datos ==7)]
print(c)
primo()
| primos 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <table>
# <tr align=left><td><img align=left src="./images/CC-BY.png">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) <NAME></td>
# </table>
#
# Note: This material largely follows the text "Numerical Linear Algebra" by Trefethen and Bau (SIAM, 1997) and is meant as a guide and supplement to the material presented there.
# + init_cell=true slideshow={"slide_type": "skip"}
from __future__ import print_function
# %matplotlib inline
import numpy
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "slide"}
# # Orthogonalization:
# # The QR Factorization, Projections,Least Squares Problems and Applications
# + [markdown] slideshow={"slide_type": "slide"}
# ## Projections
#
# A **projector** is a square matrix $P$ that satisfies
# $$
# P^2 = P.
# $$
#
# Why does this definition make sense? Why do we require it to be square?
# + [markdown] slideshow={"slide_type": "subslide"}
# A projector comes from the idea that we want to project a vector $\mathbf{v}$ onto a lower dimensional subspace. For example, suppose that $\mathbf{v}$ lies completely within the subspace, i.e. $\mathbf{v} \in \text{range}(P)$. If that is the case then $P \mathbf{v}$ should not change, or $P\mathbf{v} = \mathbf{v}$. This motivates the definition above.
# + [markdown] slideshow={"slide_type": "fragment"}
# i.e. if
# $$
# P\mathbf{v} = \mathbf{v}
# $$
# then
# $$
# P( P \mathbf{v} ) = P\mathbf{v} = \mathbf{v}.
# $$
# or $$P^2 = P$$
# + [markdown] slideshow={"slide_type": "subslide"}
# As another example, take a vector $\mathbf{x} \notin \text{range}(P)$ and project it onto the subspace $P\mathbf{x} = \mathbf{v}$. If we apply the projection again to $\mathbf{v}$ we now have
#
# $$\begin{aligned}
# P\mathbf{x} &= \mathbf{v}, \\
# P^2 \mathbf{x} & = P \mathbf{v} = \mathbf{v} \\
# \Rightarrow P^2 &= P.
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# It is also important to keep in mind the following, given again $\mathbf{x} \notin \text{range}(P)$, if we look at the difference between the projection and the original vector $P\mathbf{x} - \mathbf{x}$ and apply the projection again we have
# $$
# P(P\mathbf{x} - \mathbf{x}) = P^2 \mathbf{x} - P\mathbf{x} = 0
# $$
# which means the difference between the projected vector $P\mathbf{x} = \mathbf{v}$ lies in the null space of $P$, $\mathbf{v} \in \text{null}(P)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Complementary Projectors
#
# A projector also has a complement defined to be $I - P$.
#
# Show that this complement is also a projector.
# + [markdown] slideshow={"slide_type": "subslide"}
# We can show that this a projector by examining a repeated application of $(I-P)$:
# $$\begin{aligned}
# (I - P)^2 &= I - IP - IP + P^2 \\
# &= I - 2 P + P^2 \\
# &= I - 2P + P \\
# &= I - P.
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# It turns out that the complement projects exactly onto $\text{null}(P)$.
#
# Take
# $$
# \mathbf{x} \in \text{null}(P),
# $$
#
# then
# $$
# (I - P) \mathbf{x} = \mathbf{x} - P \mathbf{x} = \mathbf{x}
# $$
#
# since $P \mathbf{x} = 0$ implying that $\mathbf{x} \in \text{range}(I - P)$.
# + [markdown] slideshow={"slide_type": "skip"}
# We also know that
# $$
# (I - P) \mathbf{x }\in \text{null}(P)
# $$
# as well.
#
# This shows that the
# $$
# \text{range}(I - P) \subseteq \text{null}(P)
# $$
# and
# $$
# \text{range}(I - P) \supseteq \text{null}(P)
# $$
# implying that
# $$
# \text{range}(I - P) = \text{null}(P)
# $$
# exactly.
#
# Reflect on these subspaces and convince yourself that this all makes sense.
# + [markdown] slideshow={"slide_type": "subslide"}
# This result provides an important property of a projector and its complement, namely that they divide a space into two subspaces whose intersection is
#
# $$
# \text{range}(I - P) \cap \text{range}(P) = \{0\}
# $$
#
# or
#
# $$
# \text{null}(P) \cap \text{range}(P) = \{0\}
# $$
#
# These two spaces are called **complementary subspaces**.
# + [markdown] slideshow={"slide_type": "skip"}
# Given this property we can take any $P \in \mathbb C^{m \times m}$ which will split $\mathbb C^{m \times m}$ into two subspaces $S$ and $V$, assume that $\mathbf{s}\in S = \text{range}(P)$, and $\mathbf{v} \in V = \text{null}(P)$. If we have $\mathbf{x} \in \mathbb C^{m \times m}$ that we can split the vector $\mathbf{x}$ into components in $S$ and $V$ by using the projections
# $$\begin{aligned}
# P \mathbf{x} = \mathbf{x}_S& &\mathbf{x}_s \in S \\
# (I - P) \mathbf{x} = \mathbf{x}_V& &\mathbf{x}_V \in V
# \end{aligned}$$
# which we can also observe adds to the original vector as
# $$
# \mathbf{x}_S + \mathbf{x}_V = P \mathbf{x} + (I - P) \mathbf{x} = \mathbf{x}.
# $$
#
# Try constructing a projection matrix so that $P \in \mathbb R^3$ that projects a vector into one of the coordinate directions ($\mathbb R$).
# - What is the complementary projector?
# - What is the complementary subspace?
# + [markdown] slideshow={"slide_type": "skip"}
# ### Example: A non-orthogonal non-linear projector
#
# Given a vector of mols of $N$ chemical components
#
# $$
# \mathbf{n} = \begin{bmatrix} n_1 \\ n_2 \\ \vdots \\ n_N\end{bmatrix}
# $$
# where (e.g. $n_1$ is the number of moles of component $1$) and $n_i\geq 0$
#
# Then we can define the mol fraction of component $i$ as
# $$
# x_i = \frac{n_i}{\mathbf{n}^T\mathbf{1}}
# $$
# and the "vector" of mole fractions
# $$
# \mathbf{x} = \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_N\end{bmatrix}
# $$
#
# + [markdown] slideshow={"slide_type": "skip"}
# In the homework you will show that
#
# * $\mathbf{x}^T\mathbf{1} = 1$ (the sum of the mole fractions add to 1)
# * mole fractions do not form a vector space or subspace
# * There exists a non Orthogonal projector $f$ such that $f(\mathbf{n})=\mathbf{x}$, $f^2=f$
# * $P$ is singular (like all projection matrices) such that if you know the mole-fractions you don't know how many moles you have.
# * Find $N(P)$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Orthogonal Projectors
#
# An **orthogonal projector** is one that projects onto a subspace $S$ that is orthogonal to the complementary subspace $V$ (this is also phrased that $S$ projects along a space $V$). Note that we are only talking about the subspaces (and their basis), not the projectors! i.e. for orthogonal **subspaces** all the vectors in $S$ are orthogonal to all the vectors in $V$.
# + [markdown] slideshow={"slide_type": "subslide"}
# A **hermitian** matrix is one whose complex conjugate transposed is itself, i.e.
#
# $$
# P = P^\ast.
# $$
#
# With this definition we can then say: *A projector $P$ is orthogonal if and only if $P$ is hermitian.*
# + [markdown] slideshow={"slide_type": "fragment"}
# **Quick Proof**
#
# Show that if $P^2 = P$ and $P^\ast = P$, then
# $$
# \langle P\mathbf{x}, (I-P)\mathbf{x}\rangle=0
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Projection with an Orthonormal Basis
#
# We can also directly construct a projector that uses an orthonormal basis on the subspace $S$. If we define another matrix $Q \in \mathbb C^{m \times n}$ which is unitary (its columns are orthonormal) we can construct an orthogonal projector as
# $$
# P = Q Q^*.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Check that $P^* = P$ and $P^2 = P$ (Remember $Q^*Q=I$)
# + [markdown] slideshow={"slide_type": "skip"}
# Note that the resulting matrix $P$ is in $\mathbb C^{m \times m}$ as we require. This means also that the dimension of the subspace $S$ is $n$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example: Orthonormal projection and Least-Squares Problems... A review
#
# Consider the overdetermined problem $A\mathbf{x}=\mathbf{b}$ where $A\in\mathbb{R}^{3\times2}$ and $\mathbf{b}\in\mathbb{R}^3$ i.e.
#
# $$
# \begin{bmatrix} | & | \\
# \mathbf{a}_1 & \mathbf{a}_2\\
# | & | \\
# \end{bmatrix} \begin{bmatrix} x_1 \\ x_2\\ \end{bmatrix}
# = \begin{bmatrix} | \\
# \mathbf{b} \\
# | \\
# \end{bmatrix}
# $$
#
# and $\mathbf{a}_1$, $\mathbf{a}_2$ are linearly independent vectors that span a two-dimensional subspace of $\mathbb{R}^3$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Geometry
#
# Geometrically this problem looks like
#
# <img align=center, src="./images/least_squares_geometry.jpg" alt="Drawing" width=600/>
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# If $\mathbf{b}\notin C(A)$, then there is clearly no solution to $A\mathbf{x}=\mathbf{b}$. However, we can find the point $\mathbf{p}=A\hat{\mathbf{x}}\in C(A)$ that minimizes the length of the the error $\mathbf{e}=\mathbf{b}-\mathbf{p}$. While we could resort to calculus to find the values of $\hat{\mathbf{x}}$ that minimizes $||\mathbf{e}||_2$. It should be clear from the figure that the shortest error (in the $\ell_2$ norm) is the one that is perpendicular to every vector in $C(A)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# But the sub-space of vectors orthogonal to $C(A)$ is the left-Null Space $N(A^T)$, and therefore we simply seek solutions of
#
# \begin{align}
# 0 &= A^T\mathbf{e} \\
# &= A^T(\mathbf{b}-\mathbf{p})\\
# &= A^T(\mathbf{b} - A\hat{\mathbf{x}})\\
# \end{align}
#
# or we just need to solve the "Normal Equations" $A^T A\hat{\mathbf{x}} = A^T\mathbf{b}$ for the least-squares solution
# $$
# \hat{\mathbf{x}} = \left(A^T A\right)^{-1}A^T\mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# if we're actually interested in $\mathbf{p}$ which is the orthogonal projection of $\mathbf{b}$ onto $C(A)$ we get
#
# $$
# \mathbf{p}= A\hat{\mathbf{x}} = A \left(A^T A\right)^{-1}A^T\mathbf{b} = P\mathbf{b}
# $$
# where
# $$
# P = A \left(A^T A\right)^{-1}A^T
# $$
#
# is an orthogonal projection matrix (verify that $P^2 = P$ and $(I - P)\mathbf{b}\perp P\mathbf{b})$
# + [markdown] slideshow={"slide_type": "subslide"}
# For a general matrix $A$, this form of the projection matrix is rather horrid to find, however, if the columns of $A$ formed an orthonormal basis for $C(A)$, i.e. $A=Q$, then the form of the projection matrix is much simpler as $Q^T Q=I$, therefore
#
# $$
# P = QQ^T
# $$
#
# This is actually quite general. Given any orthonormal basis for a vector space $S=\mathrm{span}\langle \mathbf{q}_1,\mathbf{q}_2,\ldots,\mathbf{q}_N\rangle$. If these vectors form the columns of $Q$, then the orthogonal projector onto $S$ is always $QQ^T$ and the complement is always $I-QQ^T$.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Example: Construction of an orthonormal projector**
#
# Take $\mathbb R^3$ and derive a projector that projects onto the x-y plane and is an orthogonal projector.
# + [markdown] slideshow={"slide_type": "subslide"}
# the simplest Orthonormal basis for the $x-y$ plane are the columns of
# $$
# Q = \begin{bmatrix} 1 & 0 \\
# 0 & 1 \\
# 0 & 0
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# and an orthogonal projector onto the $x-y$ plane is simply
#
# $$
# Q Q^\ast = \begin{bmatrix} 1 & 0 \\
# 0 & 1 \\
# 0 & 0
# \end{bmatrix}
# \begin{bmatrix} 1 & 0 & 0 \\
# 0 & 1 & 0
# \end{bmatrix} = \begin{bmatrix}
# 1 & 0 & 0 \\
# 0 & 1 & 0 \\
# 0 & 0 & 0
# \end{bmatrix}
# $$
# + slideshow={"slide_type": "subslide"}
Q = numpy.array([[1., 0.],[0., 1.],[0., 0.]])
P = numpy.dot(Q, Q.T)
I = numpy.identity(3)
x = numpy.array([3., 4., 5.])
print('x = {}'.format(x))
print('\nQ = \n{}'.format(Q))
print('\nP = \n{}'.format(P))
# + slideshow={"slide_type": "fragment"}
x_S = numpy.dot(P, x)
x_V = numpy.dot(I - P, x)
print('x_S = {}'.format(x_S))
print('x_V = {}\n'.format(x_V))
print('x_S + x_V = {}'.format(x_S+x_V))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A numerically more sensible approach
#
# The previous problem calculated the projection matrix first, $P=QQ^T$ then calculated the projection and its complement by matrix vector multiplication, i.e.
#
# $$
# \mathbf{x}_S = P\mathbf{x} = (QQ^T)\mathbf{x}, \quad \mathbf{x}_V = (I - P)\mathbf{x}
# $$
#
# A mathematically equivalent, but numerically more efficient method is to calculate the following
# $$
# \mathbf{x}_S = Q(Q^T\mathbf{x}),\quad \mathbf{x}_V = \mathbf{x} - \mathbf{x}_S
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# How much more efficient is the latter than the former?
# + slideshow={"slide_type": "subslide"}
# Check
Q = numpy.array([[1, 0],[0, 1],[0, 0]])
x = numpy.array([3., 4., 5.])
print('x = {}'.format(x))
# + slideshow={"slide_type": "fragment"}
x_S = Q.dot(Q.T.dot(x))
x_V = x - x_S
print('x_S = {}'.format(x_S))
print('x_V = {}\n'.format(x_V))
print('x_S + x_V = {}'.format(x_S+x_V))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Construction of a projector that eliminates a direction
#
# Goal: Eliminate the component of a vector in the direction $\mathbf{q}$.
# + [markdown] slideshow={"slide_type": "fragment"}
#
#
# <img align=center, src="./images/least_squares_geometry.jpg" alt="Drawing" width=600/>
#
# e.g. We can calculate the projection $\mathbf{p}$ in two equivalent ways
#
# 1. find the matrix $P=QQ^T$ that projects $\mathbf{b}$ onto $C(A)$
# 2. find the matrix $P'=\mathbf{q}\mathbf{q}^T$ that projects onto the unit vector $\mathbf{q}$ normal to the plane (i.e. parallel to $\mathbf{e}$), and then take its Complement projection $\mathbf{p} = (I - P')\mathbf{b} = \mathbf{b} - \mathbf{q}(\mathbf{q}^T\mathbf{b})$
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# Form the projector $P = \mathbf{q} \mathbf{q}^\ast \in \mathbb C^{m \times m}$. The complement $I - P$ will then include everything **BUT** that direction.
#
# If $||\mathbf{q}|| = 1$ we can then simply use $I - \mathbf{q} \mathbf{q}^\ast$. If not we can write the projector in terms of the arbitrary vector $\mathbf{a}$ as
# $$
# I - \frac{\mathbf{a} \mathbf{a}^\ast}{||\mathbf{a}||^2} = I - \frac{\mathbf{a} \mathbf{a}^\ast}{\mathbf{a}^\ast \mathbf{a}}
# $$
# Note that differences in the resulting dimensions between the two values in the fraction. Also note that as we saw with the outer product, the resulting $\text{rank}(\mathbf{a} \mathbf{a}^\ast) = 1$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Now again try to construct a projector in $\mathbb R^3$ that projects onto the $x$-$y$ plane.
# + slideshow={"slide_type": "-"}
q = numpy.array([0, 0, 1])
P = numpy.outer(q, q.conjugate())
P_comp = numpy.identity(3) - P
print(P,'\n')
print(P_comp)
# + slideshow={"slide_type": "fragment"}
x = numpy.array([3, 4, 5])
print(numpy.dot(P, x), q*(q.dot(x)))
print(numpy.dot(P_comp, x), x - q*(q.dot(x)))
# + slideshow={"slide_type": "skip"}
a = numpy.array([0, 0, 3])
P = numpy.outer(a, a.conjugate()) / (numpy.dot(a, a.conjugate()))
P_comp = numpy.identity(3) - P
print(numpy.dot(P, x))
print(numpy.dot(P_comp, x))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quick Review
#
# * A projection matrix is any square matrix $P$ such that $P^2=P$
# * $P$ projects onto a subspace $S=\mathrm{range}(P)$
# * The complementary projection matrix $I-P$ projects onto $V=\mathrm{null}(P)$
# * An *orthogonal* projector can always be constructed as $P=QQ^T$ where the columns of $Q$ form an *orthonormal* basis for $S$ and $S\perp V$
# * Solutions of Least-squares problems $A\mathbf{x}=\mathbf{b}$ are essentially projection problems where we seek to solve $A\mathbf{x}=\mathbf{b}$ where $\mathbf{b}\notin C(A)$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution of Least Squares problems by the Normal Equations
#
# given $A\mathbf{x}=\mathbf{b}$ where $\mathbf{b}\notin C(A)$ we can always solve them using the Normal Equations
#
# $$
# A^T A\hat{\mathbf{x}} = A^T\mathbf{b}
# $$
#
# which actually solves $A\hat{\mathbf{x}} =\mathbf{p}$ where $\mathbf{p}$ is the orthogonal projection of $\mathbf{b}$ onto $C(A)$
# + [markdown] slideshow={"slide_type": "fragment"}
# However...as you will show in the homework, this can be numerically innaccurate because the condition number
#
# $$ \kappa(A^T A) = \kappa(A)^2 $$
#
# but there is a better way
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution of Least Squares problems by the QR factorization
#
# given any matrix $A$ that is full column rank, we will show that we can always factor it as
#
# $$
# A=QR
# $$
#
# where $Q$ is a unitary matrix whose columns form an *orthonormal* basis for $C(A)$, and $R$ is an upper triangular matrix that says how to reconstruct the columns of $A$ from the columns of $Q$
# + [markdown] slideshow={"slide_type": "subslide"}
# $$
# \begin{bmatrix} & & \\ & & \\ \mathbf{a}_1 & \cdots & \mathbf{a}_n \\ & & \\ & & \end{bmatrix} =
# \begin{bmatrix} & & \\ & & \\ \mathbf{q}_1 & \cdots & \mathbf{q}_n \\ & & \\ & & \end{bmatrix}
# \begin{bmatrix} r_{11} & r_{12} & \cdots & r_{1n} \\ & r_{22} & & \\ & & \ddots & \vdots \\ & & & r_{nn} \end{bmatrix}.
# $$
# If we write this out as a matrix multiplication we have
# $$\begin{aligned}
# \mathbf{a}_1 &= r_{11} \mathbf{q}_1 \\
# \mathbf{a}_2 &= r_{22} \mathbf{q}_2 + r_{12} \mathbf{q}_1 \\
# \mathbf{a}_3 &= r_{33} \mathbf{q}_3 + r_{23} \mathbf{q}_2 + r_{13} \mathbf{q}_1 \\
# &\vdots
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# i.e we can construct the columns of $A$ from linear combinations of the columns of $Q$. (And those specific combinations come from $R=Q^TA$)
# + [markdown] slideshow={"slide_type": "subslide"}
# given $A=QR$, then solving
#
# $$
# A\mathbf{x} = \mathbf{b}
# $$
#
# becomes
#
# $$
# QR\mathbf{x} = \mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# or since $Q^T Q=I$
#
# $$
# R\mathbf{x} = Q^T\mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# which can be solve quickly by back-substitution as it is a triangular matrix.
#
# Moreover, this problem is much better conditioned as it can be shown that $\kappa(R)=\kappa(A)$ not $\kappa(A)^2$
# + [markdown] slideshow={"slide_type": "subslide"}
# Multiplying both sides by $Q$ again shows that this problem is equivalent to
#
# $$
# QR\mathbf{x} = QQ^T\mathbf{b}
# $$
#
# or $A\mathbf{x} = QQ^T\mathbf{b}$ i.e. we are just solving $A\mathbf{x}=\mathbf{p}$ where $\mathbf{p}$ is the orthogonal projection of $\mathbf{b}$ onto $C(A)$
# + [markdown] slideshow={"slide_type": "fragment"}
# **Question**... how to find $QR$?
# + [markdown] slideshow={"slide_type": "slide"}
# ## QR Factorization
#
# One of the most important ideas in linear algebra is the concept of factorizing an original matrix into different constituents that may have useful properties. These properties can help us understand the matrix better and lead to numerical methods. In numerical linear algebra one of the most important factorizations is the **QR factorization**.
# + [markdown] slideshow={"slide_type": "fragment"}
# There are actually multiple algorithm's that accomplish the same factorization but with very different methods and numerical stability. Here we will discuss three of the algorithms
#
# 1. **Classical Gram-Schmidt Orthogonalization**: Transform $A\rightarrow Q$ by a succession of projections and calculate $R$ as a by-product of the algorithm. (Unfortunately, prone to numerical floating point error)
# 2. **Modified Gram-Schmidt Orthogonalization**: Transform $A\rightarrow Q$ by a different set of projections. Yields the same $QR$ but more numerically stable.
# 3. **Householder Triangularization**: Transform $A\rightarrow R$ by a series of Unitary transformations, can solve least squares problems directly without accumulating $Q$, or can build $Q$ on the fly.
#
# And want to find a sequence of orthonormal vectors $\mathbf{q}_j$ that span the sequence of subspaces
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Classical Gram-Schmidt
#
# We begin with a matrix $A$ with $n$ linearly independent columns.
# $$
# A = \begin{bmatrix} & & \\ & & \\ \mathbf{a}_1 & \cdots & \mathbf{a}_n \\ & & \\ & & \end{bmatrix}
# $$
# And want to find a sequence of orthonormal vectors $\mathbf{q}_j$ that span the sequence of subspaces
#
# $$
# \text{span}(\mathbf{a}_1) \subseteq \text{span}(\mathbf{a}_1, \mathbf{a}_2) \subseteq \text{span}(\mathbf{a}_1, \mathbf{a}_2, \mathbf{a}_3) \subseteq \cdots \subseteq \text{span}(\mathbf{a}_1, \mathbf{a}_2, \ldots , \mathbf{a}_n)
# $$
#
# where here $\text{span}(\mathbf{v}_1,\mathbf{v}_2,\ldots,\mathbf{v}_m)$ indicates the subspace spanned by the vectors $\mathbf{v}_1, \mathbf{v}_2, \ldots, \mathbf{v}_m$.
# + [markdown] slideshow={"slide_type": "fragment"}
# The individual $\mathbf{q}_j$ will form the columns of the matrix $Q$ such that $C(A_k)=C(Q_k)$ where the $A_k$ is a matrix with the first $k$ columns of A (or Q)
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Starting with the first vector $\mathbf{a}_1$, this forms the basis for a 1-dimensional subspace of $\mathbb{R}^m$ (i.e. a line). Thus $\mathbf{q}_1$ is a unit vector in that line or
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \mathbf{q}_1 = \frac{\mathbf{a}_1}{||\mathbf{a}_1||}.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# For $\text{span}(\mathbf{a}_1, \mathbf{a}_2)$ (which is a plane in $\mathbb{R}^m$ we already have $\mathbf{q}_1$ so we need to find a vector $\mathbf{q}_2$ that is orthogonal to $\mathbf{q}_1$, but still in the plane.
#
# An obvious option is to find the component of $\mathbf{a}_2$ that is orthogonal to $\mathbf{q}_1$ but this is just
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \begin{align}
# \mathbf{v}_2 &= (I - \mathbf{q}_1\mathbf{q}_1^T)\mathbf{a}_2 \\
# &= \mathbf{a}_2 - \mathbf{q}_1(\mathbf{q}_1^T\mathbf{a}_2)\\
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# By construction it's easy to show that $\mathbf{v_2}$ is orthogonal to $\mathbf{q}_1$ but not necessarily a unit vector but we can find $\mathbf{q}_2$ by normalizing
#
# $$
# \mathbf{q}_2 = \frac{\mathbf{v}_2}{||\mathbf{v}_2||}.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Classical Gram-Schmidt as a series of projections
# If we define the unitary matrix
# $$
# Q_k = \begin{bmatrix} & & \\ & & \\ \mathbf{q}_1 & \cdots & \mathbf{q}_k \\ & & \\ & & \end{bmatrix}
# $$
# as the first $k$ columns of Q, then we could also rewrite the first two steps of Classical Gram-Schmidt as
#
#
# set $\mathbf{v}_1 = \mathbf{a}_1$, then
# $$
# \begin{align}
# \mathbf{q}_1 &= \frac{\mathbf{v}_1}{||\mathbf{v}_1||}\\
# \mathbf{v_2} &= (I - Q_1Q_1^T)\mathbf{a}_2 = \mathbf{a}_2 - Q_1(Q_1^T\mathbf{a}_2)\\
# \mathbf{q}_2 &= \frac{\mathbf{v}_2}{||\mathbf{v}_2||}\\
# \end{align}
# $$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# and we can continue
# $$
# \begin{align}
# \mathbf{v_3} &= (I - Q_2Q_2^T)\mathbf{a}_3 = \mathbf{a}_3 - Q_2(Q_2^T\mathbf{a}_3)\\
# \mathbf{q}_3 &= \frac{\mathbf{v}_3}{||\mathbf{v}_3||}\\
# & \vdots \\
# \mathbf{v_k} &= (I - Q_{k-1}Q_{k-1}^T)\mathbf{a}_k = \mathbf{a}_k - Q_{k-1}(Q_{k-1}^T\mathbf{a}_k)\\
# \mathbf{q}_k &= \frac{\mathbf{v}_k}{||\mathbf{v}_k||}\\
# \end{align}
# $$
#
# With each step finding the component of $a_k$ that is orthogonal to all the other vectors before it, and normalizing it.
# + [markdown] slideshow={"slide_type": "subslide"}
# A picture is probably useful here...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### But what about $R$?
#
# The above algorithm appears to transform $A$ directly to $Q$ without calculating $R$ (which we need for least-squares problems).
#
# But actually that's not true as $R$ is hiding in the algorithm (much like $L$ hides in the $LU$ factorizations $A\rightarrow U$)
# + [markdown] slideshow={"slide_type": "fragment"}
# If we consider the full factorization $A = QR$
#
# $$
# \begin{bmatrix} & & \\ & & \\ \mathbf{a}_1 & \cdots & \mathbf{a}_n \\ & & \\ & & \end{bmatrix} =
# \begin{bmatrix} & & \\ & & \\ \mathbf{q}_1 & \cdots & \mathbf{q}_n \\ & & \\ & & \end{bmatrix}
# \begin{bmatrix} r_{11} & r_{12} & \cdots & r_{1n} \\ & r_{22} & & \\ & & \ddots & \vdots \\ & & & r_{nn} \end{bmatrix}.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# it follows that $R=Q^T A$ or
# $$ \begin{bmatrix} r_{11} & r_{12} & \cdots & r_{1n} \\ & r_{22} & & \\ & & \ddots & \vdots \\ & & & r_{nn}\end{bmatrix} =
# \begin{bmatrix} & -\mathbf{q}_1^T- & \\ & -\mathbf{q}_2^T- & \\ & \vdots & \\ & -\mathbf{q}_n^T- & \\ \end{bmatrix}
# \begin{bmatrix} & & \\ & & \\ \mathbf{a}_1 & \cdots & \mathbf{a}_n \\ & & \\ & & \end{bmatrix}
# .
# $$
# Which with a little bit of work, you can show that the $j$th column of $R$
# $$
# \mathbf{r}_j = Q_j^T\mathbf{a}_j
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# So the full Classical Gram-Schmidt algorithm looks something like
#
# Set $\mathbf{v}_1 = \mathbf{a}_1$, $R_{11}=||\mathbf{v}_1||$, $\mathbf{q}_1 = \mathbf{v}_1/R_{11}$
#
# Loop over columns for $j=2,\ldots,n$
#
# * Find $\mathbf{r}_j = Q^T_{j-1}\mathbf{a_j}$
# * $\mathbf{v}_j = \mathbf{a}_j - Q_{j-1}\mathbf{r}_j$
# * $R_{jj} = ||\mathbf{v}_j||_2$
# * $\mathbf{q}_j = \mathbf{v}_j/R_{jj}$
#
# Which builds up both $Q$ and $R$
# + [markdown] slideshow={"slide_type": "skip"}
# Or unrolled
# $$\begin{aligned}
# \mathbf{q}_1 &= \frac{\mathbf{a}_1}{r_{11}} \\
# \mathbf{q}_2 &= \frac{\mathbf{a}_2 - r_{12} \mathbf{q}_1}{r_{22}} \\
# \mathbf{q}_3 &= \frac{\mathbf{a}_3 - r_{13} \mathbf{q}_1 - r_{23} \mathbf{q}_2}{r_{33}} \\
# &\vdots \\
# \mathbf{q}_n &= \frac{\mathbf{a}_n - \sum^{n-1}_{i=1} r_{in} \mathbf{q}_i}{r_{nn}}
# \end{aligned}$$
# leading us to define
# $$
# r_{ij} = \left \{ \begin{aligned}
# &\langle \mathbf{q}_i, \mathbf{a}_j \rangle & &i \neq j \\
# &\left \Vert \mathbf{a}_j - \sum^{j-1}_{i=1} r_{ij} \mathbf{q}_i \right \Vert & &i = j
# \end{aligned} \right .
# $$
#
# This is called the **classical Gram-Schmidt** iteration. Turns out that the procedure above is unstable because of rounding errors introduced.
# + [markdown] slideshow={"slide_type": "subslide"}
# Which can be easily coded up in python as
# + slideshow={"slide_type": "fragment"}
# Implement Classical Gram-Schmidt Iteration
def classic_GS(A):
m, n = A.shape
Q = numpy.empty((m, n))
R = numpy.zeros((n, n))
# loop over columns
for j in range(n):
v = A[:, j]
for i in range(j):
R[i, j] = numpy.dot(Q[:, i].conjugate(), A[:, j])
v = v - R[i, j] * Q[:, i]
R[j, j] = numpy.linalg.norm(v, ord=2)
Q[:, j] = v / R[j, j]
return Q, R
# + [markdown] slideshow={"slide_type": "subslide"}
# #### And check
# + slideshow={"slide_type": "-"}
A = numpy.array([[12, -51, 4], [6, 167, -68], [-4, 24, -41]], dtype=float)
print(A)
# + slideshow={"slide_type": "fragment"}
Q, R = classic_GS(A)
print('Q=\n{}\n'.format(Q))
print('Q^TQ=\n{}'.format(numpy.dot(Q.transpose(), Q)))
# + slideshow={"slide_type": "fragment"}
print('R=\n{}\n'.format(R))
print('QR - A=\n{}'.format(numpy.dot(Q, R) - A))
# + [markdown] slideshow={"slide_type": "skip"}
# #### Full vs. Reduced QR
#
# If the original matrix $A \in \mathbb C^{m \times n}$ where $m \ge n$ then we can still define a QR factorization, called the **full QR factorization**, which appends columns full of zeros to $R$ to reproduce the full matrix.
# $$
# A = Q R = \begin{bmatrix} Q_1 & Q_2 \end{bmatrix}
# \begin{bmatrix} R_1 \\
# 0
# \end{bmatrix} = Q_1 R_1
# $$
# The factorization $Q_1 R_1$ is called the **reduced** or **thin QR factorization** of $A$.
#
# We require that the additional columns added $Q_2$ are an orthonormal basis that is orthogonal itself to $\text{range}(A)$. If $A$ is full ranked then $Q_1$ and $Q_2$ provide a basis for $\text{range}(A)$ and $\text{null}(A^\ast)$ respectively.
# + [markdown] slideshow={"slide_type": "skip"}
# #### QR Existence and Uniqueness
# Two important theorems exist regarding this algorithm which we state without proof:
#
# *Every $A \in \mathbb C^{m \times n}$ with $m \geq n$ has a full QR factorization and therefore a reduced QR factorization.*
#
# *Each $A \in \mathbb C^{m \times n}$ with $m \geq n$ of full rank has a unique reduced QR factorization $A = QR$ with $r_{jj} > 0$.*
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Modified Gram-Schmidt
#
# Unfortunately the classical Gram-Schmidt algorithm is is not stable numerically. Instead we can derive a modified method that is more numerically stable but calculates the same $Q$ and $R$ with just a different order of projections.
# + [markdown] slideshow={"slide_type": "skip"}
# Recall that the basic piece of the original algorithm was to take the inner product of $\mathbf{a}_j$ and all the relevant $\mathbf{q}_i$. Using the rewritten version of Gram-Schmidt in terms of projections we then have
#
# $$
# \mathbf{v}_j = (I - Q_{j-1}Q^T_{j-1}) \mathbf{a}_j.
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# Each of these projections is of different rank $m - (j - 1)$ although we know that the resulting $\mathbf{v}_j$ are linearly independent by construction. The modified version of Gram-Schmidt instead uses projections that are all of rank $m-1$. To construct this projection remember that we can again construct the complement to a projection and perform the following sequence of projections
#
# $$
# P_j = \hat{\!P}_{\mathbf{q}_{j-1}} \hat{\!P}_{\mathbf{q}_{j-2}} \cdots \hat{\!P}_{\mathbf{q}_{2}} \hat{\!P}_{\mathbf{q}_{1}}
# $$
#
# where
# $$\hat{\!P}_{\mathbf{q}_{j}} = I - \mathbf{q}_i\mathbf{q_i}^T$$
#
# which projects onto the complementary space orthogonal to $\mathbf{q}_i$.
#
# Note that this performs mathematically the same job as $P_i \mathbf{a}_i$ however each of these projectors are of rank $m - 1$. The reason why this approach is more stable is that we are not projecting with a possibly arbitrarily low-rank projector, instead we only take projectors that are high-rank.
# + [markdown] slideshow={"slide_type": "skip"}
# again...a picture is probably worth a lot here.
# + [markdown] slideshow={"slide_type": "skip"}
# This leads to the following set of calculations:
#
# $$\begin{aligned}
# 1.\quad \mathbf{v}^{(1)}_i &= \mathbf{a}_i \\
# 2.\quad \mathbf{v}^{(2)}_i &= \hat{\!P}_{\mathbf{q}_1} \mathbf{v}_i^{(1)} = \mathbf{v}^{(1)}_i - \mathbf{q}_1 q_1^\ast \mathbf{v}^{(1)}_i \\
# 3.\quad \mathbf{v}^{(3)}_i &= \hat{\!P}_{\mathbf{q}_2} \mathbf{v}_i^{(2)} = \mathbf{v}^{(2)}_i - \mathbf{q}_2 \mathbf{q}_2^\ast \mathbf{v}^{(2)}_i \\
# & \text{ } \vdots & &\\
# i.\quad \mathbf{v}^{(i)}_i &= \hat{\!P}_{\mathbf{q}_{i-1}} \mathbf{v}_i^{(i-1)} = \mathbf{v}_i^{(i-1)} - \mathbf{q}_{i-1} \mathbf{q}_{i-1}^\ast \mathbf{v}^{(i-1)}_i
# \end{aligned}$$
#
# The reason why this approach is more stable is that we are not projecting with a possibly arbitrarily low-rank projector, instead we only take projectors that are high-rank.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Example: Implementation of modified Gram-Schmidt**
# Implement the modified Gram-Schmidt algorithm checking to make sure the resulting factorization has the required properties.
# + slideshow={"slide_type": "fragment"}
# Implement Modified Gram-Schmidt Iteration
def mod_GS(A):
m, n = A.shape
Q = numpy.empty((m, n))
R = numpy.zeros((n, n))
v = A.copy()
for i in range(n):
R[i, i] = numpy.linalg.norm(v[:, i], ord=2)
Q[:, i] = v[:, i] / R[i, i]
for j in range(i + 1, n):
R[i, j] = numpy.dot(Q[:, i].conjugate(), v[:, j])
v[:, j] -= R[i, j] * Q[:, i]
return Q, R
# + [markdown] slideshow={"slide_type": "skip"}
# #### And check
# + slideshow={"slide_type": "skip"}
A = numpy.array([[12, -51, 4], [6, 167, -68], [-4, 24, -41]], dtype=float)
print(A)
# + slideshow={"slide_type": "skip"}
Q, R = mod_GS(A)
print('Q=\n{}\n'.format(Q))
print('Q^TQ=\n{}'.format(numpy.dot(Q.transpose(), Q)))
# + slideshow={"slide_type": "skip"}
print('R=\n{}\n'.format(R))
print('QR - A=\n{}'.format(numpy.dot(Q, R) - A))
# + [markdown] slideshow={"slide_type": "skip"}
# ### Householder Triangularization
#
# One way to also interpret Gram-Schmidt orthogonalization is as a series of right multiplications by upper triangular matrices of the matrix A. For instance the first step in performing the modified algorithm is to divide through by the norm $r_{11} = ||v_1||$ to give $q_1$:
#
# $$
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{v}_1 & \mathbf{v}_2 & \mathbf{v}_3 & \cdots & \mathbf{v}_n \\
# & & & \\
# & & &
# \end{bmatrix}
# \begin{bmatrix}
# \frac{1}{r_{11}} & &\cdots & \\
# & 1 & \\
# & & \ddots & \\
# & & & 1
# \end{bmatrix} =
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{q}_1 & \mathbf{v}_2 & \mathbf{v}_3 & \cdots & \mathbf{v}_n \\
# & & & \\
# & & &
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# We can also perform all the step (2) evaluations by also combining the step that projects onto the complement of $q_1$ by add the appropriate values to the entire first row:
#
# $$
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{v}_1 & \mathbf{v}_2 & \mathbf{v}_3 & \cdots & \mathbf{v}_n \\
# & & & \\
# & & &
# \end{bmatrix}
# \begin{bmatrix}
# \frac{1}{r_{11}} & -\frac{r_{12}}{r_{11}} & -\frac{r_{13}}{r_{11}} & \cdots \\
# & 1 & \\
# & & \ddots & \\
# & & & 1
# \end{bmatrix} =
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{q}_1 & \mathbf{v}_2^{(2)} & \mathbf{v}_3^{(2)} & \cdots & \mathbf{v}_n^{(2)} \\
# & & & \\
# & & &
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# The next step can then be placed into the second row:
# $$
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{v}_1 & \mathbf{v}_2 & \mathbf{v}_3 & \cdots & \mathbf{v}_n \\
# & & & \\
# & & &
# \end{bmatrix}
# \cdot R_1 \cdot
# \begin{bmatrix}
# 1 & & & & \\
# & \frac{1}{r_{22}} & -\frac{r_{23}}{r_{22}} & -\frac{r_{25}}{r_{22}} & \cdots \\
# & & 1 & \\
# & & & \ddots & \\
# & & & & 1
# \end{bmatrix} =
# \begin{bmatrix}
# & & & \\
# & & & \\
# \mathbf{q}_1 & \mathbf{q}_2 & \mathbf{v}_3^{(3)} & \cdots & \mathbf{v}_n^{(3)} \\
# & & & \\
# & & &
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# If we identify the matrices as $R_1$ for the first case, $R_2$ for the second case and so on we can write the algorithm as
#
# $$
# A \underbrace{R_1R_2 \quad \cdots \quad R_n}_{\hat{R}^{-1}} = \hat{\!Q}.
# $$
#
# This view of Gram-Schmidt is called Gram-Schmidt triangularization.
# + [markdown] slideshow={"slide_type": "skip"}
# Householder triangularization is similar in spirit. Instead of multiplying $A$ on the right Householder multiplies $A$ on the left by unitary matrices $Q_k$. Remember that a unitary matrix (or an orthogonal matrix when strictly real) has as its inverse its adjoint (transpose when real) $Q^* = Q^{-1}$ so that $Q^* Q = I$. We therefore have
#
# $$
# Q_n Q_{n-1} \quad \cdots \quad Q_2 Q_1 A = R
# $$
#
# which if we identify $Q_n Q_{n-1} \text{ } \cdots \text{ } Q_2 Q_1 = Q^*$ and note that if $Q = Q^\ast_n Q^\ast_{n-1} \text{ } \cdots \text{ } Q^\ast_2 Q^\ast_1$ then $Q$ is also unitary.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Householder Triangularization
#
# While Gram-Schmidt and its variants transform $A\rightarrow Q$ and calculate $R$ on the way. Householder triangularization is a rather different algorithm that transforms $A\rightarrow R$ by multiplying $A$ by a series of Unitary matrices that systematically put zeros in the subdiagonal (similar to the LU factorization)
#
# $$
# Q_n Q_{n-1} \quad \cdots \quad Q_2 Q_1 A = R
# $$
#
# which if we identify $Q_n Q_{n-1} \text{ } \cdots \text{ } Q_2 Q_1 = Q^*$ and note that if $Q = Q^\ast_n Q^\ast_{n-1} \text{ } \cdots \text{ } Q^\ast_2 Q^\ast_1$ then $Q$ is also unitary.
# + [markdown] slideshow={"slide_type": "subslide"}
# We can then write this as
# $$\begin{aligned}
# Q_n Q_{n-1} \text{ } \cdots \text{ } Q_2 Q_1 A &= R \\
# Q_{n-1} \cdots Q_2 Q_1 A &= Q^\ast_n R \\
# & \text{ } \vdots \\
# A &= Q^\ast_1 Q^\ast_2 \text{ } \cdots \text{ } Q^\ast_{n-1} Q^\ast_n R \\
# A &= Q R
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# This was we can think of Householder triangularization as one of introducing zeros into $A$ via orthogonal matrices.
#
# $$
# \begin{bmatrix}
# \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} \\
# \end{bmatrix} \overset{Q_1}{\rightarrow}
# \begin{bmatrix}
# \text{+} & \text{+} & \text{+} \\
# & \text{+} & \text{+} \\
# & \text{+} & \text{+} \\
# & \text{+} & \text{+} \\
# \end{bmatrix} \overset{Q_2}{\rightarrow}
# \begin{bmatrix}
# \text{+} & \text{+} & \text{+} \\
# & \text{-} & \text{-} \\
# & & \text{-} \\
# & & \text{-} \\
# \end{bmatrix} \overset{Q_3}{\rightarrow}
# \begin{bmatrix}
# \text{+} & \text{+} & \text{+} \\
# & \text{-} & \text{-} \\
# & & \text{*} \\
# & & \\
# \end{bmatrix} = R
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# Now the question is how do we construct the $Q_k$. The construction is usually broken down into a matrix of the form
#
# $$
# Q_k = \begin{bmatrix} I & 0 \\ 0 & H \end{bmatrix}
# $$
#
# where $I \in \mathbb C^{k-1 \times k-1}$ is the identity matrix that preserves the top $k-1$ rows
#
# and $H \in \mathbb C^{m - (k - 1) \times m - (k-1)}$ is a unitary matrix that just modifies the lower $m-k-1$ rows.
#
# Note that this will leave the rows and columns we have already worked on alone and be unitary.
# + [markdown] slideshow={"slide_type": "subslide"}
# We now turn to the task of constructing the matrix $H$. Note that the definition of $Q_k$ implies that
# $$
# \begin{align}
# Q_k \mathbf{x} &= \begin{bmatrix} I & 0 \\ 0 & H \end{bmatrix} \mathbf{x}
# &= \begin{bmatrix} I & 0 \\ 0 & H \end{bmatrix}
# \begin{bmatrix} \mathbf{x}_{top} \\ \mathbf{x}_{bottom} \end{bmatrix} \\
# &= \begin{bmatrix} \mathbf{x}_{top} \\ H\mathbf{x}_{bottom} \end{bmatrix}. \\
# \end{align}
# $$
# The vectors $\mathbf{x}_{top}$ and $\mathbf{x}_{bottom}$ are defined to be
# $$
# \begin{align}
# \mathbf{x}_{top} &= \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_{k-1}\end{bmatrix} \\
# \mathbf{x}_{bottom} &= \begin{bmatrix} x_k \\ x_{k+1} \\ \vdots \\ x_m \end{bmatrix} \\
# \end{align}
# $$
# The task is to construct the matrix $H$ so that after multiplying, $H\mathbf{x}_{bottom}$ results in a vector that has some nonzero number in the top position and the rest of the numbers are all zeroes.
# + [markdown] slideshow={"slide_type": "subslide"}
# or
#
# $$H\mathbf{x}_{bottom} = \begin{bmatrix} \alpha \\ 0 \\ \vdots \\ 0 \end{bmatrix}$$
#
# But if $H$ is unitary, it can't change the length of a vector which implies
# + [markdown] slideshow={"slide_type": "fragment"}
# $$ \alpha = ||\mathbf{x}_{bottom}||$$
# + [markdown] slideshow={"slide_type": "subslide"}
# One unitary operation that does this is the *Householder Reflection* that reflects the vector $\mathbf{x}$ over the hyper plane $H$ so that $H \mathbf{x} = ||x|| \hat{\mathbf{e}}_1$:
# <table>
# <tr align='center'>
# <img align=center, src="./images/householder.png" alt="Drawing" width=1000/>
# <tr\>
# <table\>
# + [markdown] slideshow={"slide_type": "fragment"}
# or mathematically
# $$
# \mathbf{x} = \begin{bmatrix}
# x_1 \\
# x_2 \\
# \vdots \\
# x_{n}
# \end{bmatrix}, \quad
# H\mathbf{x} = \begin{bmatrix}
# ||\mathbf{x}|| \\
# 0 \\
# \vdots \\
# 0
# \end{bmatrix} = ||\mathbf{x}|| \mathbf{e}_1.\quad\text{where}\quad\mathbf{e}_1 = \begin{bmatrix}
# 1 \\
# 0 \\
# \vdots \\
# 0
# \end{bmatrix}.
# $$
# is the first unit vector
# + [markdown] slideshow={"slide_type": "subslide"}
# This is of course the effect on only one vector. Any other vector will be reflected across $H$ (technically a hyperplane) which is orthogonal to
#
# $$\mathbf{v} = ||\mathbf{x}|| \hat{e}_1 - \mathbf{x}.$$
#
# This has a similar construction as to the projector complements we were working with before. Consider the projector defined as
#
# $$
# P x = \left (I - \mathbf{q} \mathbf{q}^T\right)\mathbf{x} = \mathbf{x} - \mathbf{q}(\mathbf{q}^T\mathbf{x})
# $$
#
# where $$\mathbf{q} = \frac{\mathbf{v}}{||\mathbf{v}||}.$$
#
# This vector $P\mathbf{x}$ is now orthogonal to $\mathbf{v}$. i.e. lies in the plane $H$
# + [markdown] slideshow={"slide_type": "subslide"}
# Since we actually want to transform $\mathbf{x}$ to lie in the direction of $\hat{e}_1$ we need to go twice as far as which allows us to identify the matrix $H$ as
#
# $$
# H = I - 2 \mathbf{q} \mathbf{q}^T.
# $$
#
# <table>
# <tr align='center'>
# <img align=center, src="./images/householder.png" alt="Drawing" width=1000/>
# <tr\>
# <table\>
# + [markdown] slideshow={"slide_type": "subslide"}
# There is actually a non-uniqueness to which direction we reflect over since another definition of $\hat{H}$ which is orthogonal to the one we originally choose is available. For numerical stability purposes we will choose the reflector that is the most different from $\mathbf{x}$. This comes back to having difficulties numerically when the vector $\mathbf{x}$ is nearly aligned with $\hat{e}_1$ and therefore one of the $H$ specification. By convention the $\mathbf{v}$ chosen is defined by
#
# $$
# \mathbf{v} = \text{sign}(x_1)||\mathbf{x}|| \hat{e}_1 + \mathbf{x}.
# $$
# + slideshow={"slide_type": "subslide"}
# Implementation of Householder QR Factorization
def householder_QR(A, verbose=False):
R = A.copy()
m, n = A.shape
QT = numpy.eye(m)
for k in range(n):
x = numpy.zeros(m)
e = numpy.zeros(m)
x[k:] = R[k:, k]
e[k] = 1.0
# simplest version v = ||x||e - x
#v = numpy.linalg.norm(x, ord=2) * e - x
# alternate version
v = numpy.sign(x[k]) * numpy.linalg.norm(x, ord=2) * e + x
v = v / numpy.linalg.norm(v, ord=2)
R -= 2.0 * numpy.outer(v,numpy.dot(v.T,R))
QT -= 2.0 * numpy.outer(v,numpy.dot(v.T,QT))
Q = QT.T[:,:n]
return Q, R
# + slideshow={"slide_type": "subslide"}
A = numpy.array([[12, -51, 4], [6, 167, -68], [-4, 24, -41]], dtype=float)
print("Matrix A = ")
print(A)
# + slideshow={"slide_type": "fragment"}
# %precision 6
Q, R = householder_QR(A, verbose=False)
print("Householder (reduced) Q =\n{}\n".format(Q))
print("Householder (full) R = ")
print(R)
# + slideshow={"slide_type": "subslide"}
m, n = A.shape
print("Check to see if factorization worked...||A -QR|| = {}".format(numpy.linalg.norm(A - numpy.dot(Q, R[:n, :n]))))
print(A - numpy.dot(Q, R[:n, :n]))
print("\nCheck if Q is unitary...||Q^TQ -I|| = {}".format(numpy.linalg.norm(Q.T.dot(Q)-numpy.eye(n))))
print(Q.T.dot(Q))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Comparison of accuracy of the different algorithms
#
# As it turns out, not all $QR$ algorithms produce the same quality of orthogonalization. Here we provide a few examples that compare the behavior of the 3 different algorithms and `numpy.linalg.qr`
# + [markdown] slideshow={"slide_type": "skip"}
# #### Example 1: Random Matrix QR
#
# Here we construct a large matrix $A$ with a random eigenspace and widely varying eigenvalues. The values along the diagonal of $R$ gives us some idea of the size of the projections as we go, i.e. the larger the values the less effective we are in constructing orthogonal directions.
# + hide_input=true slideshow={"slide_type": "skip"}
N = 80
# construct a random matrix with known singular values
U, X = numpy.linalg.qr(numpy.random.random((N, N)))
V, X = numpy.linalg.qr(numpy.random.random((N, N)))
S = numpy.diag(2.0**numpy.arange(-1.0, -(N + 1), -1.0))
A = numpy.dot(U, numpy.dot(S, V))
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1, 1, 1)
Q, R = classic_GS(A)
axes.semilogy(numpy.diag(R), 'bo', label="Classic")
Q, R = mod_GS(A)
axes.semilogy(numpy.diag(R), 'ro', label="Modified")
Q, R = householder_QR(A)
axes.semilogy(numpy.diag(R), 'ko', label="Householder")
Q, R = numpy.linalg.qr(A)
axes.semilogy(numpy.diag(R), 'go', label="numpy")
axes.set_xlabel("Index", fontsize=16)
axes.set_ylabel("$R_{ii}$", fontsize=16)
axes.legend(loc=3, fontsize=14)
axes.plot(numpy.arange(0, N), numpy.ones(N) * numpy.sqrt(numpy.finfo(float).eps), 'k--')
axes.plot(numpy.arange(0, N), numpy.ones(N) * numpy.finfo(float).eps, 'k--')
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example 2: Comparing Orthogonality
#
# Consider the $QR$ factorization of the ill-conditioned matrix
# $$
# A = \begin{bmatrix}
# 0.70000 & 0.70711 \\ 0.70001 & 0.70711
# \end{bmatrix}.
# $$
# + hide_input=true slideshow={"slide_type": "-"}
# %precision 16
A = numpy.array([[0.7, 0.70711], [0.70001, 0.70711]])
print('\ncond(A) = {}'.format(numpy.linalg.cond(A)))
# + [markdown] slideshow={"slide_type": "subslide"}
# To check that the matrix $Q$ is really unitary, we compute $A=QR$ with the different algorithms and compare
#
# $$
# ||Q^TQ - I||
# $$
# + hide_input=true slideshow={"slide_type": "-"}
Q_c, R = classic_GS(A)
r_c = numpy.linalg.norm(numpy.dot(Q_c.transpose(), Q_c) - numpy.eye(2))
print("Classic: ", r_c )
Q, R = mod_GS(A)
print("Modified: ", numpy.linalg.norm(numpy.dot(Q.transpose(), Q) - numpy.eye(2)))
Q_h, R = householder_QR(A)
r_h = numpy.linalg.norm(numpy.dot(Q_h.transpose(), Q_h) - numpy.eye(2))
print("Householder:", r_h)
Q, R = numpy.linalg.qr(A)
r = numpy.linalg.norm(numpy.dot(Q.transpose(), Q) - numpy.eye(2))
print("Numpy: ", r)
print('\ncond(A) = {}'.format(numpy.linalg.cond(A)))
print('r_classic/r_householder = {}'.format(r_c/r_h))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Applications of the QR
#
# The $QR$ factorization and unitary transformation such as Householder reflections, play important roles in a wide range of algorithms for Numerical Algorithms
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Least-Squares problems: Solving $A\mathbf{x} = \mathbf{b}$ with QR
#
# We have already discussed solving overdetermined least-squares problems using the $QR$ factorization. In general, if we seek least-squares solutions to $A\mathbf{x}=\mathbf{b}$ and $A=QR$, the problem reduces to solving the triangular problem
# $$
# R\mathbf{x} = Q^T\mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# If we use Householder triangularization to transform $A\rightarrow R$ directly, we do not need to explicitly form the matrix $Q$ and can save memory and computation. If we consider the augmented system $\begin{bmatrix} A & \mathbf{b}\end{bmatrix}
# $ and Apply the same sequence of unitary, rank 1 tranformations we used in the Householder algorithm, the sequence becomes
#
# $$
# Q_n\ldots Q_2Q_1\begin{bmatrix} A & \mathbf{b}\end{bmatrix} = \begin{bmatrix} Q^TA & Q^T\mathbf{b}\end{bmatrix} =\begin{bmatrix} R & \mathbf{c}\end{bmatrix}
# $$
#
# Thus we just need to solve
# $$
# R\mathbf{x} = \mathbf{c}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Alternatively, during the $QR$ factorization by Householder, we can just store the vectors $\mathbf{q}_1\ldots\mathbf{q}_n$ and reconstruct $Q^T\mathbf{b}$ by rank-1 updates as neccessary.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Finding Eigenvalues
#
# As it turns out, the $QR$ factorization and Householder transformation are also extremely useful for finding eigenvalues of a matrix $A$. There is an entire notebook `13_LA_eigen.ipynb` that develops these ideas in detail. Here we will just discuss the parts that relate to the $QR$ and orthogonalization algorithms.
# + [markdown] slideshow={"slide_type": "skip"}
# ### The basics
#
# The eigenproblem
# $$
# A \mathbf{x} = \lambda \mathbf{x}
# $$
# can be rewritten as
#
# $$
# ( A - \lambda I)\mathbf{x} = \mathbf{0}
# $$
#
# which implies that the eigenvectors are in the Null space of $A-\lambda I$.
#
# However for this matrix to have a non-trivial Null space, requires that $A-\lambda I$ is singular.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Characteristic Polynomial
#
# If $A-\lambda I$ is singular, it follows that
#
# $$
# \det( A - \lambda I) = {\cal P}_A(\lambda) = 0
# $$
#
# where ${\cal P}_A(\lambda)$ can be shown to be a $m$th order polynomial in $\lambda$ known as the **characteristic polynomial** of a matrix $A$
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# ## Computing Eigenvalues
#
# In basic linear algebra classes we usually find the eigenvalues by directly calculating the roots of ${\cal P}_A(\lambda)$ which can work for low-degree polynomials. Unfortunately the following theorem (due to Galois) suggests this is not a good way to compute eigenvalues:
#
# **Theorem:** For an $m \geq 5$ there is a polynomial $\mathcal{P}(z)$ of degree $m$ with rational coefficients that has a real root $\mathcal{P}(z_0) = 0$ with the property that $z_0$ cannot be written using any expression involving rational numbers, addition, subtraction, multiplication, division, and $k$th roots.
#
# I.e., there is no way to find the roots of a polynomial of degree $>4$ in a deterministic, fixed number of steps.
# + [markdown] slideshow={"slide_type": "skip"}
# #### Not all is lost however!
#
# We just must use an iterative approach where we construct a sequence that converges to the eigenvalues.
#
# **Some Questions**
# * How does this relate to how we found roots previously?
# * Why will it still be difficult to use our rootfinding routines to find Eigenvalues?
#
# We will return to how we actually find Eigenvalues (and roots of polynomials) after a bit more review
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Similarity Transformations
#
# Generally, we say any two matrices $A$ and $B$ are **similar** if they can be related through an invertible matrix $M$ as
#
# $$
# A = M^{-1} B M
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# **Example**
#
# The general Eigen problem
# $$
# A\mathbf{x}=\lambda\mathbf{x}
# $$
#
# is really $n$ problems for each eigenvalue, eigenvector pair i.e.
#
# $$
# A\mathbf{x}_i = \lambda_i\mathbf{x}_i\quad\text{for}\quad i=1,\ldots,n
# $$
#
# which can be written concisely in matrix form as
#
# $$
# AX =X\Lambda
# $$
#
# where $X$ is a matrix whose columns contain the eigenvectors and $\Lambda$ is a diagonal matrix of corresponding eigenvalues. This form is always true
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example Diagonalizable matrices
#
# If a matrix $A\in\mathbb{R}^{n\times n}$ has $n$ linearly independent eigenvectors, then we say $A$ is *diagonalizable* and we can factor it as
#
# $$
# A = X\Lambda X^{-1}
# $$
# Where $X$ is a matrix of eigenvectors, and $\Lambda$ is a diagonal matrix of corresponding Eigenvalues.
#
# which says $A$ is similar to $\Lambda$ with similarity transform $X$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Theorem:
# If $A$ and $B$ are similar matrices, they have the same eigenvalues and their eigenvectors are related through an invertible matrix $M$
# + [markdown] slideshow={"slide_type": "fragment"}
# **Proof**: Let
#
# $$
# B = M A M^{-1}
# $$
# or
# $$
# BM = MA
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# if $A\mathbf{x} = \lambda\mathbf{x}$ then
# $$
# BM\mathbf{x} = M A\mathbf{x} = \lambda M\mathbf{x}
# $$
# or
# $$
# B\mathbf{y} = \lambda\mathbf{y}
# $$
#
# which shows that $\lambda$ is also an eigenvalue of $B$ with corresponding eigenvector $\mathbf{y} = M\mathbf{x}$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Schur Factorization
#
# A **Schur factorization** of a matrix $A$ is defined as
#
# $$
# A = Q T Q^\ast
# $$
#
# where $Q$ is unitary and $T$ is upper-triangular. Because $Q^\ast=Q^{-1}$ (for square unitary matrices). It follows directly that $A$ and $T$ are similar.
#
# * Good News! $T$ is upper triangular so its eigenvalues can just be read of the diagonal
# * Bad News! There is no deterministic way to calculate $T$ as that would violate Galois theory of polynomials
# + [markdown] slideshow={"slide_type": "subslide"}
# **Theorem:** Every matrix $A \in \mathbb C^{m \times m}$ has a Schur factorization.
#
# **Partial Proof** for a diagonalizable matrix. If $A$ is diagonalizable, $A=X\Lambda X^{-1}$. But we know we can always factor $X=QR$ and substitute to show
#
# $$
# A = Q(R\Lambda R^{-1})Q^T
# $$
#
# and it is not hard to show that the product $R\Lambda R^{-1}$ is also an upper triangular matrix (exercise left to the reader).
#
# (For a non-diagonalizable matrix the proof requires showing the existence of the Jordan form $A=MJM^{-1}$)
# + [markdown] slideshow={"slide_type": "subslide"}
# Note that the above results imply the following
# - An eigen-decomposition $A = X \Lambda X^{-1}$ exists if and only if $A$ is non-defective (it has a complete set of eigenvectors)
# - A unitary transformation $A = Q \Lambda Q^\ast$ exists if and only if $A$ is Hermitian ($A^\ast = A$)
# - A Schur factorization always exists
#
# Note that each of these lead to a means for isolating the eigenvalues of a matrix and will be useful when considering algorithms for finding them.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Hessenberg form
#
# The first step to finding the Schur factorization is to try and get $A$ as close to triangular as possible without changing its eigenvalues. This requires a series of similarity transformations.
#
# As it turns out, the closest we can do is to reduce it to Hessenberg form which is upper triangular with one extra subdiagonal. Which can be done with $n$ explicit similarity transformations using Householder Transformations
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \begin{bmatrix}
# \text{x} & \text{x} & \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} & \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x} & \text{x} & \text{x}
# \end{bmatrix} \overset{H_1^\ast A_0 H_1}{\rightarrow}
# \begin{bmatrix}
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & \text{x} & \text{x}& \text{x} & \text{x}
# \end{bmatrix} \overset{H_2^\ast A_1H_2}{\rightarrow}
# \begin{bmatrix}
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & 0 & \text{x}& \text{x} & \text{x} \\
# 0 & 0 & \text{x}& \text{x} & \text{x}
# \end{bmatrix} \overset{H_3^\ast A_2H_3}{\rightarrow}
# \begin{bmatrix}
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# \text{x} & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & \text{x} & \text{x}& \text{x} & \text{x} \\
# 0 & 0 & \text{x}& \text{x} & \text{x} \\
# 0 & 0 & 0 & \text{x} & \text{x}
# \end{bmatrix}
# $$
#
# so we have the sequence $H = Q^\ast A Q$ which has the same eigenvalues as the original matrix $A$.
#
# **Question**? Why can't we just use Householder to take $A\rightarrow T$ like we did for the $QR$?
# + [markdown] slideshow={"slide_type": "slide"}
# ## QR/RQ Algorithm
#
# Given a matrix in Hessenberg form, it turns out we can use repeated $QR$ factorizations to reduce the size of the subdiagonal and iterate towards the Schur factorization to find all the eigenvalues simultaneously.
#
# The simplest algorithm just iterates
# ```python
# while not converged:
# Q, R = numpy.linalg.qr(A)
# A = R.dot(Q)
# ```
# calculating the $QR$ factorization of $A$, then forming a new $A=RQ$, This sequence will eventually converge to the Schur decomposition of the matrix $A$.
#
# Code this up and see what happens.
# + slideshow={"slide_type": "subslide"}
# %precision 6
m = 3
A0 = numpy.array([[2, 1, 1], [1, 3, 1], [1, 1, 4]])
# + slideshow={"slide_type": "-"}
MAX_STEPS = 10
A=A0
print('A=')
print(A)
for i in range(MAX_STEPS):
Q, R = numpy.linalg.qr(A)
A = numpy.dot(R, Q)
print()
print("A({}) =".format(i))
print(A)
# + slideshow={"slide_type": "subslide"}
print()
print("True eigenvalues: ")
print(numpy.sort(numpy.linalg.eigvals(A0)))
print()
print("Computed eigenvalues: ")
print(numpy.sort(numpy.diag(A)))
# + [markdown] slideshow={"slide_type": "subslide"}
# So why does this work? The first step is to find the $QR$ factorization
#
# $$A^{(k-1)} = Q^{(k)}R^{(k)}$$
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# which is equivalent to finding
#
# $$
# (Q^{(k)})^T A^{(k-1)} = R^{(k)}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# and multiplying on the right leads to
#
# $$
# (Q^{(k)})^T A^{(k-1)} Q^{(k)} = R^{(k)} Q^{(k)} = A^{(k)}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# In this way we can see that this is a similarity transformation of the matrix $A^{(k-1)}$ since the $Q^{(k)}$ is an orthogonal matrix ($Q^{-1} = Q^T$). This of course is not a great idea to do directly but works great in this case as we iterate to find the upper triangular matrix $R^{(k)}$ which is exactly where the eigenvalues appear.
# + [markdown] slideshow={"slide_type": "subslide"}
# In practice this basic algorithm is modified to include a few additions:
#
# 1. Before starting the iteration $A$ is reduced to tridiagonal or Hessenberg form.
# 1. Motivated by the inverse power iteration we observed we instead consider a shifted matrix $A^{(k)} - \mu^{(k)} I$ for factoring. The $\mu$ picked is related to the estimate given by the Rayleigh quotient. Here we have
#
# $$
# \mu^{(k)} = \frac{(q_m^{(k)})^T A q_m^{(k)}}{(q_m^{(k)})^T q_m^{(k)}} = (q_m^{(k)})^T A q_m^{(k)}.
# $$
#
# 1. Deflation is used to reduce the matrix $A^{(k)}$ into smaller matrices once (or when we are close to) finding an eigenvalue to simplify the problem.
#
# This has been the standard approach until recently for finding eigenvalues of a matrix.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Application: Finding the roots of a polynomial
#
# Numpy has a nice function called roots which returns the $n$ roots of a $n$th degree polynomial
#
# $$
# p(x) = p_0 x^n + p_1 x^{n-1} + p_2 x^{n-2} + \ldots + p_n
# $$
#
# described by a $n+1$ vector of coefficients $\mathbf{p}$
# + slideshow={"slide_type": "subslide"}
p = numpy.array([ 1, 1, -1])
r = numpy.roots(p)
print(r)
# + slideshow={"slide_type": "fragment"}
p = numpy.random.rand(8)
r = numpy.roots(p)
print(r)
# + [markdown] slideshow={"slide_type": "subslide"}
# This routine, does not try and actually find the roots of a high-order polynomial, instead it actually calculates the eigenvalues of a **companion matrix** $C$ whose characteristic polynomial $P_C(\lambda)$ is the **monic** polynomial
#
# $$c(x) = c_0 + c_1 x + c_2 x^2 + \ldots + c_{n-1} x^{n-1} + x^n $$
#
# It can be shown that this matrix can be constructed as ([see e.g.](https://en.wikipedia.org/wiki/Companion_matrix))
# $$
# C(p)=\begin{bmatrix}
# 0 & 0 & \dots & 0 & -c_0 \\
# 1 & 0 & \dots & 0 & -c_1 \\
# 0 & 1 & \dots & 0 & -c_2 \\
# \vdots & \vdots & \ddots & \vdots & \vdots \\
# 0 & 0 & \dots & 1 & -c_{n-1}
# \end{bmatrix}.
# $$
#
# + slideshow={"slide_type": "subslide"}
def myroots(p, verbose=False):
''' Calculate the roots of a polynomial described by coefficient vector
in numpy.roots order
p(x) = p_0 x^n + p_1 x^{n-1} + p_2 x^{n-2} + \ldots + p_n
by finding the eigenvalues of the companion matrix
returns:
--------
eigenvalues sorted by |\lambda|
'''
# construct the companion matrix of the coefficient vector c
# make p monic and reverse the order for this definition of the companion matrix
c = numpy.flip(p/p[0])
if verbose:
print(c)
m = len(c) - 1
C = numpy.zeros((m,m))
C[:,-1] = -c[:-1]
C[1:,:-1] = numpy.eye(m-1)
if verbose:
print('C = \n{}'.format(C))
# calculate the eigenvalues of the companion matrix, then sort by |lambda|
eigs = numpy.linalg.eigvals(C)
index = numpy.flip(numpy.argsort(numpy.abs(eigs)))
return eigs[index]
# + slideshow={"slide_type": "subslide"}
p = numpy.array([ 1, 1, -1])
r = numpy.roots(p)
print(r)
mr = myroots(p)
print
print(mr)
# + slideshow={"slide_type": "fragment"}
# %precision 4
p = numpy.random.rand(5)
r = numpy.roots(p)
print('nproots: {}'.format(r))
mr = myroots(p)
print()
print('myroots: {}'.format(mr))
| 11_LA_QR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
from salishsea_tools import places
# %matplotlib inline
# -
def get_the_data(year):
evenlessink = []
thestring = f'/data/sallen/results/MEOPAR/202007/sink_evenslower/SalishSea_1d_{year}03*ptrc_T*.nc'
for filename in sorted(glob.glob(thestring)):
print (filename)
data = xr.open_dataset(filename)
data_extract = data.isel(deptht = 1, y = places.PLACES['S3']['NEMO grid ji'][0],
x = places.PLACES['S3']['NEMO grid ji'][1] )
evenlessink.append(data_extract)
data.close()
thestring = f'/data/sallen/results/MEOPAR/202007/sink_evenslower/SalishSea_1d_{year}04*ptrc_T*.nc'
for filename in sorted(glob.glob(thestring)):
print (filename)
data = xr.open_dataset(filename)
data_extract = data.isel(deptht = 1, y = places.PLACES['S3']['NEMO grid ji'][0],
x = places.PLACES['S3']['NEMO grid ji'][1] )
evenlessink.append(data_extract)
data.close()
timeseries = xr.concat(evenlessink, dim='time_counter')
return timeseries
timeseries2015 = get_the_data('2015')
timeseries2015.diatoms.plot();
timeseries2016 = get_the_data('2016')
timeseries2016.diatoms.plot();
timeseries2017 = get_the_data('2017');
timeseries2017.diatoms.plot();
| notebooks/Tuning/Bloom_Timing_TVD_evenlesssinking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Standard imports
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# Load mavenn
import mavenn
print(mavenn.__path__)
# +
# Load example data
data_df = mavenn.load_example_dataset('gb1')
# Separate test from data_df
ix_test = data_df['set']=='test'
test_df = data_df[ix_test].reset_index(drop=True)
print(f'test N: {len(test_df):,}')
# Remove test data from data_df
data_df = data_df[~ix_test].reset_index(drop=True)
print(f'training + validation N: {len(data_df):,}')
data_df.head(10)
# +
# Get sequence length
L = len(data_df['x'][0])
# Define model
model = mavenn.Model(regression_type='GE',
L=L,
alphabet='protein',
gpmap_type='additive',
ge_noise_model_type='Gaussian',
ge_heteroskedasticity_order=0)
# -
# Set training data
model.set_data(x=data_df['x'],
y=data_df['y'],
validation_flags=(data_df['set']=='validation'),
shuffle=True)
# Fit model to data
history = model.fit(learning_rate=.0005,
epochs=1000,
batch_size=1000,
early_stopping=True,
early_stopping_patience=5,
linear_initialization=False)
# Save model
model.save('gb1_ge_additive_homogaussian')
# Load model
model = mavenn.load('gb1_ge_additive_homogaussian')
# +
# Subsample indices for easy plotting and information estimation
N_test = len(test_df)
ix = np.random.rand(N_test) < .1
# Get x and y
x = test_df['x'].values[ix]
y = test_df['y'].values[ix]
# +
# Show training history
print('On test data:')
# Compute likelihood information
I_var, dI_var = model.I_variational(x=x, y=y)
print(f'I_var_test: {I_var:.3f} +- {dI_var:.3f} bits')
# Compute predictive information
I_pred, dI_pred = model.I_predictive(x=x, y=y)
print(f'I_pred_test: {I_pred:.3f} +- {dI_pred:.3f} bits')
I_var_hist = model.history['I_var']
val_I_var_hist = model.history['val_I_var']
fig, ax = plt.subplots(1,1,figsize=[4,4])
ax.plot(I_var_hist, label='I_var_train')
ax.plot(val_I_var_hist, label='I_var_val')
ax.axhline(I_var, color='C2', linestyle=':', label='I_var_test')
ax.axhline(I_pred, color='C3', linestyle=':', label='I_pred_test')
ax.legend()
ax.set_xlabel('epochs')
ax.set_ylabel('bits')
ax.set_title('training hisotry')
ax.set_ylim([0, I_pred*1.2]);
# +
# Compute phi and yhat values
phi = model.x_to_phi(x)
yhat = model.phi_to_yhat(phi)
# Create grid for plotting yhat and yqs
phi_lim = [-5, 2.5]
phi_grid = np.linspace(phi_lim[0], phi_lim[1], 1000)
yhat_grid = model.phi_to_yhat(phi_grid)
yqs_grid = model.yhat_to_yq(yhat_grid, q=[.16,.84])
# Create two panels
fig, ax = plt.subplots(1, 1, figsize=[4, 4])
# Illustrate measurement process with GE curve
ax.scatter(phi, y, color='C0', s=5, alpha=.2, label='test data')
ax.plot(phi_grid, yhat_grid, linewidth=2, color='C1',
label='$\hat{y} = g(\phi)$')
ax.plot(phi_grid, yqs_grid[:, 0], linestyle='--', color='C1',
label='68% CI')
ax.plot(phi_grid, yqs_grid[:, 1], linestyle='--', color='C1')
ax.set_xlim(phi_lim)
ax.set_xlabel('latent phenotype ($\phi$)')
ax.set_ylabel('measurement ($y$)')
ax.set_title('measurement process')
ax.legend()
# Fix up plot
fig.tight_layout()
plt.show()
# +
# Set wild-type sequence
gb1_seq = model.x_stats['consensus_seq']
# Get effects of all single-point mutations on phi
theta_dict = model.get_theta(gauge='user',
x_wt=gb1_seq)
# Create two panels
fig, ax = plt.subplots(1, 1, figsize=[10, 5])
# Left panel: draw heatmap illustrating 1pt mutation effects
ax, cb = mavenn.heatmap(theta_dict['theta_lc'],
alphabet=theta_dict['alphabet'],
seq=gb1_seq,
cmap='PiYG',
ccenter=0,
ax=ax)
ax.set_xlabel('position ($l$)')
ax.set_ylabel('amino acid ($c$)')
cb.set_label('effect ($\Delta\phi$)', rotation=-90, va="bottom")
ax.set_title('mutation effects')
# Fix up plot
fig.tight_layout()
plt.show()
# -
# Test simulate_data
sim_df = model.simulate_dataset(N=1000)
sim_df.head()
| mavenn/examples/models/train_gb1_ge_additive_homogaussian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 32-bit
# name: python374jvsc74a57bd0b39ab04170b6c60b52a31d4a7ab4d204539d261a537423dd50ae2a4297d5cc93
# ---
# # [Problem 26: Reciprocal cycles](https://projecteuler.net/problem=26)
#
# ## Key Idea
#
# using the special relation between fraction and remainders
#
# avoiding brute forcing using mathematical properties
# ### Initial idea
#
# O(n³) complexity.
#
# Brute force through every possible patterns (repeating decimals) with string comparison and check whether the pattern is found multiple times in the decimals.
# + tags=[]
from decimal import Decimal as d, getcontext
getcontext().prec = 5000
def get_pattern(s):
l = len(s)
for start_pos in range(l):
for pattern_length in range(1, l // 2 + 1):
# maximum pattern length is l // 2 because of the precision limitation
# can't determine whether it is repeating or not if it's longer than l // 2
pattern = s[start_pos:start_pos + pattern_length]
# print(f'\nstart_pos = {start_pos}, pattern = \'{pattern}\'')
# compare the pattern to the rest of the string
found = None
for find_pos in range(start_pos + pattern_length, l, pattern_length):
compare = s[find_pos:find_pos + pattern_length]
# print(f' comparing \'{pattern}\' and \'{compare}\'')
if pattern != compare:
if found is not None and len(pattern) > len(compare):
# pattern does match but goes over the precision limit
found = pattern
else:
# pattern does not match
found = None
break
else:
found = pattern
if found is not None:
found = pattern
return found # pattern match found
# no pattern is found in the fraction
return ''
max = [('', 0)] # pattern, n (of 1/n)
for i in range(1, 1000):
# d(1) / d(6) = 0.1666666666666666666666666667
# └ string stored in f ┘
# f = '166666666666666666666666666'
# trim first 2 characters and last 1 character if exists
f = str(d(1) / d(i))[2:-1]
if len(f) < 20:
continue # not reciprocal
result = get_pattern(f)
if len(result) > len(max[0][0]):
max = [(result, i, f)]
elif len(result) == len(max[0][0]):
max.append((result, i, f))
print(max[0][1])
# -
# **NOTE**: Python's `count()` function could've been used in this case. No need to write a custom string comparator.
# ### Remainder method (from *<NAME>* in solution thread)
#
# O(n²) complexity. Key idea explained in [Repeating decimal from Wikipedia](https://en.wikipedia.org/wiki/Repeating_decimal#A_shortcut).
#
# > It follows that any repeating decimal with period n, and k digits after the decimal point that do not belong to the repeating part, can be written as a (not necessarily reduced) fraction whose denominator is (10^n − 1)10^k.
#
# > Conversely **the period of the repeating decimal of a fraction c/d will be (at most) the smallest number n such that 10ⁿ − 1 is divisible by d.**
#
# > For example, the fraction 2/7 has d = 7, and the smallest k that makes 10^k − 1 divisible by 7 is k = 6, because 999999 = 7 × 142857. The period of the fraction 2/7 is therefore 6.
#
# Using the fact the problem *only* requires the period of the repeating decimal (length of the pattern), we can derive the period ***without calculating the actual decimals***.
#
# ***NOTE***: Look for *shortcuts* to the answer.
#
# By finding the smallest `p` for the given `i` with the following code, we can efficiently calculate the period of the repeating decimal.
#
# ```python
# while p < i and (10 ** p) % i != 1:
# p += 1
# ```
# + tags=[]
n = 0 # the n with the longest period
max_period = 0
for i in range(1, 1000, 2): # step of 2 to skip every even number
if i % 5 != 0: # skip every multiple of 5
p = 1 # period for current i
while p < i and (10 ** p) % i != 1:
p += 1
if p > max_period:
max_period = p
n = i
print(ans)
| p26.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Step 3 - Prepare Data - Task 4 - Transform, 5 - Sort, 6 - Feature Engineer Data - CLASS ASSIGNMENT
#
# This notebook provides the Python code for taking Covid total cases at county level and creating state level aggregation. It demonstrates how you can transform the data, sort the data, and engineer new features. Its results are stored in an intermediate file for rest of exercises.
#
# Students will be developing a similar notebook for total deaths. The corresponding notebook is included in the answer section.
# ## Import Libraries
# + id="65vQsMRLuVcf"
import pandas as pd
from datetime import date
# -
# ## Set up Environment Flag
# + id="m1mRgcvpUyry"
using_Google_colab = False
using_Anaconda_on_Mac_or_Linux = True
using_Anaconda_on_windows = False
# -
# ## Connect to Google Drive
#
# This step will only be executed if you have set environment flag using_Google_colab to True
# + colab={"base_uri": "https://localhost:8080/"} id="xfu1XYInUyry" outputId="b0beb99f-f2ee-485e-b816-94b8afe5a1fe"
if using_Google_colab:
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="OCcLGXoHqyqH"
# ## PD4.4 - Read file in your chosen environment
# + colab={"base_uri": "https://localhost:8080/", "height": 626} id="WwqKGnxSvTia" outputId="a0ad66ba-e58d-47a6-d081-8169105919cb"
if using_Google_colab:
df_total_deaths = pd.read_csv('/content/drive/MyDrive/COVID_Project/input/USA_Facts/covid_deaths_usafacts.csv')
if using_Anaconda_on_Mac_or_Linux:
df_total_deaths = pd.read_csv('../input/USA_Facts/covid_deaths_usafacts.csv')
if using_Anaconda_on_windows:
df_total_deaths = pd.read_csv(r'..\input\USA_Facts\covid_deaths_usafacts.csv')
df_total_deaths = df_total_deaths.astype({'countyFIPS': str}).astype({'stateFIPS': str})
df_total_deaths
# -
# ## PD 4.4 - Select data for LA County
df_total_deaths_LA = df_total_deaths[df_total_deaths['County Name'] == 'Los Angeles County']
df_total_deaths_LA
# ## PD 4.4 - Transform LA County data to total cases by date
df_total_deaths_LA_by_date = df_total_deaths_LA.melt(id_vars=['State',
'stateFIPS',
'County Name',
'countyFIPS'],
var_name='Date',
value_name='Total Deaths')
df_total_deaths_LA_by_date
# ## PD 4.4 - Transform all County data to total cases by date
df_total_county_deaths_by_date = df_total_deaths.melt(id_vars=['State',
'stateFIPS',
'County Name',
'countyFIPS'],
var_name='Date',
value_name='Total Deaths')
df_total_county_deaths_by_date
# + [markdown] id="WH-pyk7-zUm6"
# ## PD4.5 - Group total deaths by state
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="jUk52W1Azbrk" outputId="6b84b169-8608-4d58-8b78-472ac1b9e0da"
df_total_deaths_by_state = df_total_deaths.groupby(['State', 'stateFIPS']).sum().reset_index()
df_total_deaths_by_state
# + [markdown] id="v8QA3C4e6lX6"
# ## PD 4.5 - Transform state total deaths to total death by date
#
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="9WknN0H96q94" outputId="9ec409cb-c762-4597-e005-29d7c578ea0c"
df_total_deaths_by_state_by_date = df_total_deaths_by_state.melt(id_vars=['State','stateFIPS'],
var_name='Date',
value_name='Total Deaths')
df_total_deaths_by_state_by_date
# -
# ## PD 5.4 - Sort LA County Total Deaths
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="w4teS2bAUyr2" outputId="121b0db4-1d94-4c3e-8e52-dbcf8c48788e"
df_total_deaths_LA_by_date = df_total_deaths_LA_by_date.astype({'Date': 'datetime64[ns]'})
df_sorted_LA_county_total_deaths = df_total_deaths_LA_by_date.sort_values(by=['Date'])
df_sorted_LA_county_total_deaths
# -
# ## PD 5.4 - Sort County Total Deaths
df_total_county_deaths_by_date = df_total_county_deaths_by_date.astype({'Date': 'datetime64[ns]'})
df_sorted_county_total_deaths = df_total_county_deaths_by_date.sort_values(by=['stateFIPS', 'countyFIPS', 'Date'])
df_sorted_county_total_deaths
# ## PD 5.5 - Sort State Total Deaths
df_total_deaths_by_state_by_date = df_total_deaths_by_state_by_date.astype({'Date': 'datetime64[ns]'})
df_sorted_state_total_deaths = df_total_deaths_by_state_by_date.sort_values(by=['stateFIPS', 'Date'])
df_sorted_state_total_deaths
# + [markdown] id="yaaqgwJTP3Dd"
# ## PD 6.5 - Find Incremental Deaths for LA County
# -
df_sorted_LA_county_total_deaths['Incremental Deaths'] = df_sorted_LA_county_total_deaths.groupby(['stateFIPS', 'countyFIPS'])['Total Deaths'].apply(lambda x: x - x.shift(1))
df_sorted_LA_county_total_deaths
# ## PD 6.5 - Find 7 days moving average using Shift
df_sorted_LA_county_total_deaths['deaths moving_avg'] = df_sorted_LA_county_total_deaths.groupby(['stateFIPS', 'countyFIPS'])['Incremental Deaths'].apply(
lambda x: (x + x.shift(1) + x.shift(2) + x.shift(3) + x.shift(4) + x.shift(5) + x.shift(6))/7)
df_sorted_LA_county_total_deaths[40:60]
# ## PD 6.5 - Find Incremental Deaths for all counties
df_sorted_county_total_deaths['Incremental Deaths'] = df_sorted_county_total_deaths.groupby(['stateFIPS', 'countyFIPS'])['Total Deaths'].apply(lambda x: x - x.shift(1))
df_sorted_county_total_deaths
# ## PD 6.5 - Find 7 days moving average for all counties
df_sorted_county_total_deaths['deaths moving_avg'] = df_sorted_county_total_deaths.groupby(['stateFIPS', 'countyFIPS'])['Incremental Deaths'].apply(
lambda x: (x + x.shift(1) + x.shift(2) + x.shift(3) + x.shift(4) + x.shift(5) + x.shift(6))/7)
df_sorted_county_total_deaths
# ## PD 6.5 - Save County data
if using_Google_colab:
df_sorted_county_total_deaths.to_csv('/content/drive/MyDrive/COVID_Project/output/confirmed_deaths_by_state.csv')
if using_Anaconda_on_Mac_or_Linux:
df_sorted_county_total_deaths.to_csv('../output/confirmed_deaths_by_state.csv')
if using_Anaconda_on_windows:
df_sorted_county_total_deaths.to_csv(r'..\output\confirmed_deaths_by_state.csv')
# ## PD 6.6 - Find Incremental Deaths for state
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="DL7Ujc5TQB0B" outputId="952e6ffd-0465-4d66-e5b6-7ddb33caf0f6"
df_sorted_state_total_deaths['Incremental Deaths'] = df_sorted_state_total_deaths.groupby('State')['Total Deaths'].apply(
lambda x: x - x.shift(1))
df_sorted_state_total_deaths
# -
# ## PD 6.6 - Find 7-days moving average for state
# + id="HFoC8cPAjmmQ"
df_sorted_state_total_deaths['deaths moving_avg'] = df_sorted_state_total_deaths.groupby('State')['Incremental Deaths'].apply(
lambda x: (x + x.shift(1) + x.shift(2) + x.shift(3) + x.shift(4) + x.shift(5) + x.shift(6))/7)
df_sorted_state_total_deaths
# + [markdown] id="1PlBLvm0jnaR"
# ## PD 6.6 - Save Final Results for State
# + id="XHMwj-XkRYW7"
if using_Google_colab:
df_sorted_state_total_deaths.to_csv('/content/drive/MyDrive/COVID_Project/output/confirmed_deaths_by_state.csv')
if using_Anaconda_on_Mac_or_Linux:
df_sorted_state_total_deaths.to_csv('../output/confirmed_deaths_by_state.csv')
if using_Anaconda_on_windows:
df_sorted_state_total_deaths.to_csv(r'..\output\confirmed_deaths_by_state.csv')
# + id="pw7eKleQUyr7"
| Notebook-Class-Assignment-Answers/Step-3-Prepare-Data-Task-6-Engineer-Features-Class-Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
with open("time1.txt","r") as f:
time = f.read()
row_list = time.split("\n")
pat = re.compile("time: (\d+.\d+)")
total_time = 0
for i in row_list:
total_time += float(pat.findall(i)[0])
total_time += 71.17
(total_time / 3600)
# +
import pickle as pkl
import sys
import os
sys.path.append("..")
from datatool import train_test_val_split
import numpy as np
import torch
import torch.nn as nn
from torch.utils.data import DataLoader,TensorDataset
from model import BertEncoder, BertPooler,BertConfig,Encoder
from lc_model import LinearRegression
from lc_tool import setup_logger,setup_seed,getGrad,adjust_learning_rate
import time
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.model_selection import train_test_split
import argparse
import logging
import matplotlib.pyplot as plt
def fit(x,encoder,decoder,device,hist_len=6):
encoderInput = x[:,:,:hist_len,:].float().to(device)
encoderOutput,lastEncoderHiddenState,attention = encoder(encoderInput)
decoderInput = lastEncoderHiddenState.squeeze(0)
decoderOutput = decoder(decoderInput)
return decoderOutput, attention
def predict(sample,encoder,decoder,device):
encoder.eval()
decoder.eval()
sample = sample.view(1,*sample.shape)
decoderOutput,attention = fit(sample,encoder,decoder,device)
predict_result = torch.max(decoderOutput, 1)[1].data.cpu().numpy().squeeze()
return predict_result, attention.reshape(6,attention.shape[2],attention.shape[3])
def plot_attention(att):
fig = plt.figure(figsize=(3,3))
plt.imshow(att)
plt.yticks(np.arange(att.shape[0]))
plt.xticks(np.arange(att.shape[1]))
return fig
def plot_batch(one_batch,figsize=(10,10),alpha=0.5):
fig = plt.figure(figsize=figsize)
for i, seq in enumerate(one_batch):
mask = (seq!=0)
if mask.sum()==0:
continue
hist_seq,fut_seq = seq[:6],seq[6:]
if i == 0:
plt.scatter(hist_seq[:,0],hist_seq[:,1],c="orange",label="hist",lw=3)
plt.scatter(fut_seq[:,0],fut_seq[:,1],c="green",label="future",lw=3,ls='--')
# plt.gca().add_patch(patches.Rectangle((seq[5,0]-10, seq[5,1]-0.05), 10, 0.1,edgecolor="black",facecolor="orange"))
plt.text(x= seq[5,0]-5, y = seq[5,1]-0.5,s=f"{i}",fontsize=12,fontweight="bold", horizontalalignment='center')
elif i == 1:
hist_mask,fut_mask = (hist_seq!=0),(fut_seq!=0)
mhist_seq = hist_seq[hist_mask].reshape(-1,2)
mfut_seq = fut_seq[fut_mask].reshape(-1,2)
# plot hist
if hist_mask.sum()==0:
pass
else:
plt.scatter(mhist_seq[:,0],mhist_seq[:,1],c="blue",label="neighbor hist",alpha=alpha+0.3,lw=3)
# plot future
if fut_mask.sum()==0:
pass
else:
plt.scatter(mfut_seq[:,0],mfut_seq[:,1],c="blue",label="neighbor future",alpha=alpha,lw=3,ls='--')
plt.text(x= mfut_seq[0,0]-5, y = mfut_seq[0,1]-0.5,s=f"{i}",fontsize=12,fontweight="bold", horizontalalignment='center')
#plot current
if (seq[5]!=0).all():
pass
# plt.gca().add_patch(patches.Rectangle((seq[5,0]-10, seq[5,1]-0.05), 10, 0.1,edgecolor="black",facecolor="blue"))
# plt.text(x= seq[5,0]-5, y = seq[5,1]-0.5,s=f"{i}",fontsize=12,fontweight="bold", horizontalalignment='center')
else:
hist_mask,fut_mask = (hist_seq!=0).any(axis=1),(fut_seq!=0).any(axis=1)
mhist_seq = hist_seq[hist_mask].reshape(-1,2)
mfut_seq = fut_seq[fut_mask].reshape(-1,2)
# plot hist
if hist_mask.sum()==0:
pass
else:
plt.scatter(mhist_seq[:,0],mhist_seq[:,1],c="blue",alpha=alpha+0.3,lw=3)
# plot future
if fut_mask.sum()==0:
pass
else:
plt.scatter(mfut_seq[:,0],mfut_seq[:,1],c="blue",alpha=alpha,lw=3,ls='--')
plt.text(x= mfut_seq[0,0]-5, y = mfut_seq[0,1]-0.5,s=f"{i}",fontsize=12,fontweight="bold", horizontalalignment='center')
#plot current
if (seq[5]!=0).all():
pass
# plt.gca().add_patch(patches.Rectangle((seq[5,0]-10, seq[5,1]-0.05), 10, 0.1,edgecolor="black",facecolor="blue"))
# plt.text(x= seq[5,0]-5, y = seq[5,1]-0.5,s=f"{i}",fontsize=12,fontweight="bold", horizontalalignment='center')
plt.legend(loc="best")
return fig
# +
# xy-version
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
BS = 256
SEED = 10
setup_seed(SEED)
conf_json = {
"hidden_size":32,
"num_hidden_layers":1,
"num_attention_heads":1,
"intermediate_size":32*4,
"hidden_act":"gelu",
"hidden_dropout_prob":0,
"attention_probs_dropout_prob":0
}
config = BertConfig()
config = config.from_dict(conf_json)
encoder = Encoder(config).to(DEVICE)
decoder = LinearRegression(conf_json['hidden_size']).to(DEVICE)
encoder.load_state_dict(torch.load('model/encoder_sd-10_lr-0.0001_wd-5e-05_maxep-100_bs-256_opt-adam_clip-5.0202118234833480.pkl'))
decoder.load_state_dict(torch.load('model/decoder_sd-10_lr-0.0001_wd-5e-05_maxep-100_bs-256_opt-adam_clip-5.0202118234833480.pkl'))
with open("new_data/total.pkl","rb") as f:
data = pkl.load(f)
right_data, left_data, keep_data = data['right'], data['left'], data['keep']
keep_label = torch.zeros(keep_data.shape[0])
keepDataset = TensorDataset(torch.from_numpy(keep_data),keep_label)
i= 15500
sample_data = keepDataset[i][0]
sample_label = keepDataset[i][1]
plot_batch(sample_data,figsize=(5,5))
predict_result, attention = predict(sample_data,encoder,decoder,DEVICE)
att = attention[:,0,:]
plot_attention(att)
plt.show()
# +
import h5py
class Encoder(nn.Module):
def __init__(self,config):
super(Encoder, self).__init__()
self.config = config
# self.bn = Mask_BN()
self.embedding = nn.Linear(12,self.config.hidden_size, bias=False)
self.encoder = BertEncoder(self.config)
self.pooler = BertPooler(self.config)
self.gru = nn.GRU(self.config.hidden_size,self.config.hidden_size,batch_first=True)
self.output = nn.Linear(self.config.hidden_size,10)
def forward(self,x):
batchsize,vnum,seqlen,featsize = x.shape[0],x.shape[1],x.shape[2],x.shape[3]
# m =(x.transpose(1, 2).reshape(-1,vnum,featsize)!=0).any(axis=2)*1
# x = self.bn(x)
x_3d = x.transpose(1, 2)
x_3d = x_3d.reshape(-1,vnum,featsize)
x_centralization = x_3d - (x_3d!=0)*(x_3d[:,0,:].unsqueeze(1))
x_emb = self.embedding(x_3d)
m = (x_emb!=0).any(axis=2)*1
attention_mask = m.unsqueeze(1).unsqueeze(2)# wzz
attention_mask = (1.0 - attention_mask) * -10000.0
encoded_outputs,attention = self.encoder(x_emb,attention_mask,get_attention_matrices=True)
encoded_output = self.pooler(encoded_outputs[-1])
encoded_output = encoded_output.reshape(batchsize,seqlen,self.config.hidden_size)
encoded_output,last_encoded_output = self.gru(encoded_output)
return encoded_output,last_encoded_output,attention[0].data.cpu().numpy()
def predict(sample,encoder,decoder,device):
encoder.eval()
decoder.eval()
sample = sample.view(1,*sample.shape)
decoderOutput,attention = fit(sample,encoder,decoder,device)
predict_result = torch.max(decoderOutput, 1)[1].data.cpu().numpy().squeeze()
return predict_result, attention.reshape(6,attention.shape[2],attention.shape[3])
def plot_attention(att):
fig = plt.figure(figsize=(3,3))
plt.imshow(att)
plt.yticks(np.arange(att.shape[0]))
plt.xticks(np.arange(att.shape[1]))
return fig
with h5py.File("../pickle_data/salstm_combine_feature_12_dim.hdf5","r") as f:
right_data = f['right_data'][()]
left_data = f['left_data'][()]
keep_data = f['keep_data'][()]
keep_label = torch.zeros(keep_data.shape[0])
keepDataset = TensorDataset(torch.from_numpy(keep_data),keep_label)
i= 15500
sample_data = keepDataset[i][0][:,:,:2]
print(sample_data.shape)
sample_label = keepDataset[i][1]
plot_batch(sample_data,figsize=(5,5))
predict_result, attention = predict(sample_data,encoder,decoder,DEVICE)
att = attention[:,0,:]
plot_attention(att)
plt.show()
# -
for i in range(0,right_data.shape[0],40):
sample_data = rightDataset[i][0]
sample_label = rightDataset[i][1]
plot_batch(sample_data,figsize=(5,5))
plt.savefig(f"temp/right_pic/{i}_track.png")
predict_result, attention = predict(sample_data,encoder,decoder,DEVICE)
att1 = attention[:,0,:]
plot_attention(att1)
plt.savefig(f"temp/right_pic/{i}_att_1.png")
# att2 = attention[:,:,0]
# plot_attention(att2)
# plt.savefig(f"temp/right_pic/{i}_att_2.png")
plt.close("all")
| new_neighbor/lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
r = 7
s = list()
for x in range(r+1):
for y in range(r+1):
for z in range(r+1):
if x ** 2 + y ** 2 + z ** 2 == r**2:
if z == 6:
s.append([x,y,z])
s = sorted(s, reverse=True, key=lambda x: x[2])
print(s)
def stations(r):
r1 = r + 1
rSquare = r ** 2
s = set()
for x in range(-r, r1):
for y in range(-r, r1):
for z in range(-r, r1):
if x ** 2 + y ** 2 + z ** 2 == rSquare:
s.add((x, y, z))
return s
stations(7)
def nextStations(s1, r):
stations = []
x, y, z = s1
if z - 1 == -r:
return (0, 0, -r)
for zNext in range(z-1, -r, -1):
chosen = next_station_in_z(x, y, zNext, r)
stations.append(chosen)
return stations
def risk(s1, s2, r):
x1, y1, z1 = s1
x2, y2, z2 = s2
d = math.sqrt( (x2 - x1) ** 2 + (y2 - y1) ** 2 + (z2 - z1) ** 2)
return (d/ (r * math.pi)) ** 2
def real_risk(s1, s2, r):
x1, y1, z1 = s1
x2, y2, z2 = s2
Δ = math.sqrt((x2-x1)**2+(y2-y1)**2+(z2-z1)**2)
φ = math.asin((Δ/2/r))
d = 2*φ*r
return (d/ (r * math.pi)) ** 2
# +
direct = risk((0, 0, 7), (0, 0, -7), 7)
withOneStop = risk((0, 0, 7), (0, 7, 0), 7) + risk((0, 7, 0), (0, 0, -7), 7)
r_direct = real_risk((0, 0, 7), (0, 0, -7), 7)
r_withOneStop = real_risk((0, 0, 7), (0, 7, 0), 7) + real_risk((0, 7, 0), (0, 0, -7), 7)
# -
print(direct, withOneStop, direct - withOneStop)
print(r_direct, r_withOneStop, r_direct - r_withOneStop)
def is_station(s, r):
x, y, z = s
def valid(val):
return val <= r and val >= -r
return valid(x) and valid(y) and valid(z) and x ** 2 + y ** 2 + z ** 2 == r**2
def next_station_in_z(x, y, z, r):
station = (x, y, z)
for incr_x in range(r * 2):
for incr_y in range(incr_x):
potential_candidates = set([
(x+incr_x, y, z),
(x-incr_x, y, z),
(x, y+incr_y, z),
(x, y-incr_y, z),
(x+incr_x, y+incr_y, z),
(x+incr_x, y-incr_y, z),
(x-incr_x, y+incr_y, z),
(x-incr_x, y-incr_y, z),
(x+incr_y, y, z),
(x-incr_y, y, z),
(x, y+incr_x, z),
(x, y-incr_x, z),
(x+incr_y, y+incr_x, z),
(x+incr_y, y-incr_x, z),
(x-incr_y, y+incr_x, z),
(x-incr_y, y-incr_x, z),
])
candidates = list(filter(lambda x: is_station(x, r), potential_candidates))
if candidates:
return min([(risk(station, c, r), c) for c in candidates])[1]
return None
def nextStation(s1, r):
best_station = None
min_risk = 2 ** 32
x, y, z = s1
if z - 1 == -r:
return (0, 0, -r)
for zNext in range(z-1, -r, -1):
chosen = next_station_in_z(x, y, zNext, r)
if chosen:
new_risk = risk(s1, chosen, r)
if new_risk > min_risk:
return best_station
best_station = chosen
min_risk = new_risk
return best_station
radius = 7
nextStation((0, 0, radius), radius)
sumrisk = 0
start, end = (0, 0, radius), (0, 0, -radius)
while start != end:
# print(start, end)
ns = nextStation(start, radius)
# print(ns)
sumrisk += real_risk(start, ns, radius)
start = ns
print(sumrisk)
def min_risk(r):
stations = nextStations((0, 0, r), r)
stations = [(0, 0, r)] + list(filter(lambda x: x is not None, stations)) + [(0, 0, -r)]
srisks = []
for skip in range(1, len(stations)-1):
lskip = skip
rskip = len(stations) - skip - 1
stations_ = stations[0:lskip] + stations[lskip+1:rskip] + stations[rskip+1:]
srisk = 0
for i in range(len(stations_) - 1):
s = stations_[i]
snext = stations_[i+1]
rr= real_risk(s, snext, r)
#print(s, snext, rr)
srisk += rr
srisks.append(srisk)
return min(srisks)
min_risk(1023)
| 2020-02-25.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a target="_blank" href="https://colab.research.google.com/github/ati-ozgur/kurs-neural-networks-deep-learning/blob/master/notebooks/hello-world-MNIST.ipynb">Run in Google Colab
# </a>
# This notebook modified from following links:
#
# - [tensorflow quickstart beginner](https://www.tensorflow.org/tutorials/quickstart/beginner)
# - [keras cnn](https://github.com/keras-team/keras/blob/master/examples/mnist_cnn.py)
#
import tensorflow as tf
tf.__version__
# +
mnist = tf.keras.datasets.mnist
# +
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# -
print("x_train.shape",x_train.shape)
print("y_train.shape",y_train.shape)
print("x_test.shape",y_test.shape)
print("y_test.shape",y_test.shape)
x_train, x_test = x_train / 255.0, x_test / 255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
model.compile(loss=loss_fn,
optimizer='adam',
metrics=['accuracy'])
model.fit(x_train,
y_train,
batch_size=64,
epochs=10)
model.evaluate(x_test, y_test, verbose=2)
| notebooks/hello-world-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import MultinomialNB
from sklearn import svm
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from skimage.transform import pyramid_gaussian
from sklearn.model_selection import train_test_split
from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_recall_curve
from sklearn.model_selection import GridSearchCV
# import matplotlib.pyplot as plt
from inspect import signature
# print(glob.glob("../ILIYAN Master Thesis/Dataset/*"))
# %matplotlib inline
# %precision 2
# -
df = pd.read_csv('dataset/word2vec/tfidf_stem_3.csv', index_col=0)
df_t1 = pd.read_csv('dataset/w2v_fs/df_top_1_stem_3.csv', index_col=0)
df_t1t2 = pd.read_csv('dataset/w2v_fs/df_top_1_and_2_stem_3.csv', index_col=0)
df_class = pd.read_csv('dataset/features_norm.csv', index_col=0)
df['Class'] = df_class['Class']
df_t1['Class'] = df_class['Class']
df_t1t2['Class'] = df_class['Class']
def draw_confusionmatrix(y_test, y_hat):
plt.figure(figsize=(10,7))
cm = confusion_matrix(y_test, y_hat)
ax = sns.heatmap(cm, annot=True, fmt="d")
plt.ylabel('True label')
plt.xlabel('Predicted label')
acc = accuracy_score(y_test, y_hat)
print(f"Sum Axis-1 as Classification accuracy: {acc}")
print('\n')
print(classification_report(y_test, y_hat))
print('\n')
def gen_train_and_test(df, test_size=0.20, random_state=42):
X = df.loc[:, df.columns != 'Class']
y = df.Class
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=random_state)
return X_train, X_test, y_train, y_test
def split_train_and_test(X, y, test_size=0.20, random_state=42):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=random_state)
return X_train, X_test, y_train, y_test
def start_model(df, model):
X_train, X_test, y_train, y_test = gen_train_and_test(df)
model.fit(X_train, y_train)
y_hat = model.predict(X_test)
draw_confusionmatrix(y_test, y_hat)
# ### GaussianNB
model = GaussianNB()
from sklearn.cross_validation import cross_val_score, cross_val_predict
from sklearn import metrics
def run_cross_validation(df, model, scoring='accuracy', cv=10):
X = df.loc[:, df.columns != 'Class']
y = df.Class
print("avg accuracy:"+str(np.average(cross_val_score(model, X, y, scoring='accuracy', cv=cv))))
print("avg f1:"+str(np.average(cross_val_score(model, X, y, scoring='f1', cv=cv))))
print("avg precision:"+str(np.average(cross_val_score(model, X, y, scoring='precision', cv=cv))))
print("avg recall:"+str(np.average(cross_val_score(model, X, y, scoring='recall', cv=cv))))
return None
run_cross_validation(df, model)
run_cross_validation(df_t1, model)
run_cross_validation(df_t1t2, model)
| NB Optimization Stem 3 - AVERAGE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 알테어를 활용한 탐색적 데이터 시각화
# ===
#
# (Explorative Data Visualization with Altair)
# ===
#
# 
#
# <div style="page-break-after: always;"></div>
# + [markdown] colab_type="text" id="IyWDKt1wJhjq" slideshow={"slide_type": "slide"}
# # 2장. 데이터 유형, 마크 및 인코딩 채널
#
# - 데이터 시각화는 _그래픽 마크_(graphical mark)의 집합체로 완성된다.
# - 마크의 속성에는 위치(position), 모양(shape), 크기(size), 색상(color) 등이 포함된다.
# - 이러한 마크는 _채널_(channel)을 통하여 특정 데이터 값에 인코드(encode)된다.
# + [markdown] slideshow={"slide_type": "fragment"}
# - 다음과 같은 기본적 뼈대를 가지고, 다양한 데이터 시각화 결과를 완성할 수 있다.
# - _데이터 유형_
# - _마크_
# - _인코딩 채널_
# - 이들에 대해서 살펴보고, 이들에 대한 사용 방법을 공부하자.
# - _이 노트북은 [data visualization curriculum](https://github.com/uwdata/visualization-curriculum)의 일부이다._
# + colab={} colab_type="code" id="Y5K12kFBKE6v" slideshow={"slide_type": "fragment"}
import pandas as pd
import altair as alt
# + [markdown] colab_type="text" id="dkfRE2_2KOZ9" slideshow={"slide_type": "slide"}
# ## 1. 세계 개발 데이터
#
# + [markdown] colab_type="text" id="LbfvKf2tzCmb" slideshow={"slide_type": "fragment"}
# - 세계 개발 데이터는 세계 각국의 건강과 인구에 관한 데이터이다.
# - `year`(연도): 1955년부터 2005년까지
# - `country`(국가): 63개 국가
# - `pop`(인구), `life_expect`(기대 수명), `fertility`(출산율)
# - 수집 주체: [Gapminder Foundation](https://www.gapminder.org/)
# - 강연: [<NAME>'s popular TED talk](https://www.youtube.com/watch?v=hVimVzgtD6w)
# + pycharm={"name": "#%%\n"}
# import IPython.display.YouTubeVideo class.
from IPython.display import YouTubeVideo
# create an instance of YouTubeVideo class with provided youtube video id.
youtube_video = YouTubeVideo('hVimVzgtD6w')
# display youtube video
display(youtube_video)
# -
# - 세계 개발 데이터를 시각화하기 위하여,
# [vega-datasets](https://github.com/vega/vega-datasets) 컬렉션으로부터
# 판다스 데이터프레임으로 적재하자.
# + colab={} colab_type="code" id="cMgPRofkJHJP" slideshow={"slide_type": "fragment"}
from vega_datasets import data as vega_data
data = pd.read_json(vega_data.gapminder.url)
# + [markdown] colab_type="text" id="ioAtaF9P0IMX" slideshow={"slide_type": "subslide"}
# - 데이터 세트의 규모를 파악해 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="oM-Lodk_KK8T" outputId="134a2f4a-c15d-4fcd-fd93-49b26a4f5e30" slideshow={"slide_type": "fragment"}
data.shape
# + [markdown] colab_type="text" id="spJtZFQN0KCQ" slideshow={"slide_type": "fragment"}
# - 693행 및 6 열! 데이터 내용을 들여다 보자:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="SpB2dH0JKVJe" outputId="ec103505-4d8f-4310-a6e1-7eea3d789c28" slideshow={"slide_type": "fragment"}
data.head(5)
# + [markdown] slideshow={"slide_type": "subslide"}
# - 데이터에 관한 주요 통계량을 살펴보자.
# + slideshow={"slide_type": "fragment"}
data.describe(include='all')
# + [markdown] colab_type="text" id="ZpJ2OTZbY3Kf" slideshow={"slide_type": "subslide"}
# - 국가마다 5년 주기로 데이터가 저장되어 있다.
# - 인구 규모는 최소 5.4만명에서 최대 13억명 수준이고, 중앙값은 1.2억명 수준이다.
# - 기대수명은 최소 23.6세에서 최대 82.6세이고, 중앙값은 69.5세 수준이다.
# - 출산율은 최소 0.9명에서 최대 8.5명이고, 중앙값은 2.9명 수준이다.
# - `cluster` 필드에는 정수 코드가 보이는데, 이것이 무엇인지는 나중에 알아보자.
# + [markdown] colab_type="text" id="nTSCOwsEvo15" slideshow={"slide_type": "fragment"}
# - 데이터프레임 규모를 줄이기 위하여, 2000년도 데이터만 추출하자:
# + colab={} colab_type="code" id="kCVTdUK2Le_M" slideshow={"slide_type": "fragment"}
data2000 = data.loc[data['year'] == 2000]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="mOLR5Nh-MLWm" outputId="8629f998-e723-4b92-9a95-f0df3c9843a8" slideshow={"slide_type": "fragment"}
data2000.head(5)
# + [markdown] colab_type="text" id="xyIb1bzLJgG-" slideshow={"slide_type": "slide"}
# ## 2. 데이터 유형
#
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "fragment"}
# - 효과적인 시각화를 위한 첫번째 요소는 입력 데이터이다.
# - 데이터 값은 다양한 측정 형태를 표현한다.
# - 이들 측정치에서 어떤 종류의 비교가 가능할까?
# - 이러한 비교를 위해서는 어떤 종류의 시각적 인코딩이 필요할까?
# - 알테어에서 (시각적 인코딩 선택 사항을 알리기 위해) 사용하는 기본적 *데이터 유형*부터 살펴보자.
# - 데이터 유형에 근거하여 우리가 사용할 비교 유형이 결정된다.
# - 이를 통하여 데이터 시각화 설계를 위한 의사결정이 가능하다.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# ### 2.1 명목형
#
# - *명목형*(nominal: N) 데이터는 범주(category) 이름으로 구성되고, *범주형* 데이터라고도 한다.
# - 명목형 데이터에서는 값이 동등한지 비교할 수 있다.
# - 명목형 값 A와 명목형 값 B는 같은가, 아니면 다른가? (A = B) 아니면 (A != B).
# - 위 데이터 세트에서 `country` 필드는 명목형이다.
# - 명목형 데이터를 시각화할 때, 값이 같은지 아니면 다른지를 판단할 수 있어야 한다.
# - 위치, 색상(파랑, 빨강, 초록, *기타*), 모양 등의 채널을 활용할 수 있다.
# - 크기 채널은 사용하기는 어렵다.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# ### 2.2 서수형
#
# - *서수형*(ordinal: O) 데이터는 특정 순서를 가진 값으로 구성된다.
#
# - 서수형 데이터를 사용하여 값의 순위 순서를 비교할 수 있다.
# - 값 A가 값 B보다 앞에 오는가, 아나면 뒤에 오는가? (A < B) 아니면 (A >= B).
# - A가 B보다 작은가, 아니면 A가 B보다 크거나 같은가?
# - 위 데이터 세트에서 `year` 필드는 서수형이다.
# - 서수형 데이터를 시각화할 때, 값의 순위 순서를 판단할 수 있어야 한다.
# - 위치, 크기, 색상 값의 밝기 등의 채널을 활용할 수 있다.
# - (순서를 지정할 수 없는) 색상 채널은 사용하기 어렵다.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# ### 2.3 정량형
#
# - *정량형*(quantitative: Q) 데이터는 수치적 차이를 구분할 수 있는 값들로 구성된다.
# - 정량형 데이터를 구간(interval) 척도와 비율(ratio) 척도로 구분할 수 있다.
# - 구간 척도에 해당하는 데이터에서는 두 값 사이의 거리를 측정할 수 있다.
# - 값 A와 값 B 사이의 거리는 얼마인가? (A - B).
# - 예를 들자면, A는 B에서 12 단위 떨어져 있다.
# - 비율 척도에에서는 영점(zero-point)이 의미를 가지며,
# 비율이나 축척 인자를 측정할 수 있다.
# - 값 A는 값 B의 몇 배인가?(A / B)
# - 예를 들자면, A는 B의 10%에 해당한다. B는 A의 10배이다.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# - 위 데이터 세트에서, `year` 필드는 정량형 구간 척도에 해당하는 필드이다.
# - 구간 척도에서 영점은 의미가 없다. 연도, 온도 등은 구간 척도의 예이다.
# - 연도에서는 서기, 단기, 불기, 공기, ... 등과 같이 영점에 절대적인 의미가 없으며,
# 서기 0년이라는 시점이 시간이 없다는 의미가 아니다.
# - 온도에서는 섭씨, 화씨 등과 같이 영점에 절대적인 의미가 없으며,
# 섭씨 0도라는 점이 온도가 없다는 의미가 아니다.
# - 구간 척도에서 영점은 편의상 기준점을 설정한 것일 뿐이다.
# - 위 데이터 세트에서, `fertility`(출산율) 및 `life_expect`(기대 수명)은 정량형 비율 척도에 해당하는 필드이다.
# - 비율척도에서는 영점이 의미를 가진다.
# 무게, 길이, 넓이, 출산율, 기대 수명 등은 비율 척도의 예이다.
# - 무게나 길이에서 영점은 무게나 길이가 없다는 의미이다.
# - 베가-라이트는 정량형 데이터를 표시할 수 있지만, 구간 척도와 비율 척도를 구별하지는 않는다.
# - 정량형 데이터를 사각화할 때,
# - 위치, 크기, 색상 값 등의 채널을 활용할 수 있다.
# - 비율 척도 값의 비례 관계를 비교할 때는, 축에서 *영점의 표시가 필수적*이다.
# - 구간 척도 값의 비교에서는 *영점 표시를 생략*할 수 있다.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# ### 2.4 시간형
#
# - 시간형(temporal: T) 데이터는 시점이나 시간 구간을 측정하는 값들로 구성된다.
# - 이는 정량형 데이터의 특별한 경우로서 타임스탬프(timestamp)로 부른다.
# - 시간을 표현하는 다양한 표현 방식을 가진다.
# 예를 들어서 [그레고리 달력](https://ko.wikipedia.org/wiki/%EA%B7%B8%EB%A0%88%EA%B3%A0%EB%A6%AC%EB%A0%A5)을 참고하라.
# - 베가-라이트에서 시간 유형은 (연, 월, 일, 시간 등의) 다양한 시간 단위에 대한 처리가 가능하며,
# 특정 시간 단위를 처리할 수 있다.
# - 시간에 관한 값의 예는
# - `“2019-01-04T17:50:35.643Z”`와 같은 세계 표준 [ISO 날짜-시간 형식](https://ko.wikipedia.org/wiki/ISO_8601)과
# - `“2019-01-04”`나 `“Jan 04 2019”`와 같은 날짜를 표시하는 문자열을 포괄한다.
# - 우리가 살펴보고 있는 세계 개발 데이터 세트에는 완전한 시간형 데이터가 포함되지 않았다.
# - 대신에 `year` 필드에는 정수로 표현된 연도 값이 들어있다.
# - 알테어에서 시간 데이터를 쓰는 자세한 방법에 관해서는 [Times and Dates documentation](https://altair-viz.github.io/user_guide/times_and_dates.html)를 참고하라.
# - 날짜/시간에 관한 국제 표준을 이해하려면 [UTC 와 표기법, 그리고 ISO 8601, RFC 3339 표준](https://ohgyun.com/416)를 참고하라.
# + [markdown] colab_type="text" id="vyj1xBF2NTY6" slideshow={"slide_type": "subslide"}
# ### 2.5 요약
#
# - 이들 데이터 유형은 상호배타적이지 않으며, 오히려 계층적 구조라고 할 수 있다.
# - 서수형 데이터에서도 (이름의 동등성에 관한) 명목형 비교가 가능하다.
# - 정량형 데이터에서도 (순위 순서에 관한) 서수형 비교가 가능하다.
# - 더군다나 이들 데이터 유형이 고정된 범주로만 취급되지 않는다.
# - 숫자로 표시된 데이터 필드라고 해서 반드시 정량형 유형으로만 처리해야 하는 것은 아니다.
# - 예를 들어서, (10대, 20대, ...)와 같은 연령 집합을 가정하면, 이를
# 명목형으로 처리하여 이름만 구별할 수도 있고,
# 서수형으로 처리하여 순위를 인정할 수도 있고,
# 정량형으로 처리하여 평균값을 계산할 수도 있다.
# - 이러한 데이터를 어떻게 시각적으로 인코드할 수 있는지 살펴 보자.
# + [markdown] colab_type="text" id="yaz3tS2RKf03" slideshow={"slide_type": "slide"}
# ## 3. 인코딩 채널
# + [markdown] colab_type="text" id="yaz3tS2RKf03" slideshow={"slide_type": "slide"}
# - 알테어의 핵심에는 인코딩의 활용이 있는데,
# 이를 통해 데이터 유형에 따라서 구별된 데이터 필드를
# 선정된 *마크* 유형의 인코딩 *채널*에 연결시킬 수 있다.
# - 사용 가능한 채널을 모두 확인하려면, [Altair encoding documentation](https://altair-viz.github.io/user_guide/encoding.html)를 참고하라.
# + [markdown] slideshow={"slide_type": "subslide"}
# - 중요한 인코딩 채널은 다음과 같다.
# - `x`: 마크의 수평 축 또는 x-축
# - `y`: 마크의 수직 축 또는 y-축
# - `size`: 마크의 크기, 마크 유형에 따라서 마크의 면적이나 길이를 의미함
# - `color`: 마크의 색상, [legal CSS color](https://developer.mozilla.org/en-US/docs/Web/CSS/color_value)로 지정함
# - `opacity`: 마크의 투명도, 0(완전 투명)부터 1(완전 불투명)까지로 지정함
# - `shape`: `point` 마크를 위한 기호의 모양
# - `tooltip`: 마크 위로 마우스를 올릴(hover) 때 출력할 툴팁 텍스트
# - `order`: 마크 표시 순서
# - `column`: 데이터를 수평 정렬하는 하위 플롯으로 배치하는 열
# - `row`: 데이터를 수직 정렬하는 하위 플롯으로 배치하는 행
# + [markdown] colab_type="text" id="vujZA4b6OCRC" slideshow={"slide_type": "slide"}
# ### 3.1 X
#
# - `x` 인코딩 채널로 마크의 수평 위치, x 좌표를 설정한다.
# - 축의 제목은 자동적으로 지정된다.
# - 아래 차트에서 데이터 유형을 `Q`로 지정하였으므로, 연속적인 선형 축 스케일로 자동 처리되었다.
# + colab={"base_uri": "https://localhost:8080/", "height": 87} colab_type="code" id="nswya-ToLaRP" outputId="2a958755-7f70-4a86-ab6f-2cfb24ad7231" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q')
)
# + [markdown] colab_type="text" id="WwQuuyKUOElr" slideshow={"slide_type": "slide"}
# ### 3.2 Y
#
# - `y` 인코딩 채널로 마크의 수직 위치, y 좌표를 설정한다.
# - `cluster` 필드를 서수형(`O`)으로 지정하였으므로, 이산형 축으로 처리되었다.
# - 이산형 축에서 값 간의 간격은 기본값으로 처리된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="eP38vNrdNrHF" outputId="9ff5b4a8-143c-4595-845f-026ed2e747a8" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('cluster:O')
)
# + [markdown] colab_type="text" id="2BQetZgvc0f_" slideshow={"slide_type": "subslide"}
# - 직전 차트에서 서수형으로 지정되었던 y 축을 정량형으로 변경하면 어떻게 될까?
# - y 축을 기대 수명 `life_expect` 필드로, 데이터 유형은 `Q`로 지정하자.
# - 결과는 두 축 모두 선형 스케일을 가지는 산점도(scatter plot)로 시각화된다.
# - 출산율과 기대 수명 사이에 강력한 상관관계가 확인된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="n8PJhH6jN68w" outputId="ac41eaeb-4872-48db-ddf1-ab6b5c84f387" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q')
)
# + [markdown] colab_type="text" id="GUKc1DEDTbA0" slideshow={"slide_type": "subslide"}
# - 기본적으로, 선형 정량형 스케일로 지정된 축에는 원점이 포함되는데,
# 이는 비율 척도 데이터의 비교를 위한 기준선을 제공하기 위한 것이다.
# - 그렇지만 어떤 경우에는 원점 기준선이 의미가 없거나
# 비교를 위하여 특정 구간에만 초점을 맞추고 싶을 수 있다.
# - 원점 기준선이 자동적으로 포함되는 것을 방지하려면,
# 인코딩 `scale` 매개변수를 사용해서 스케일 맵핑을 설정해야 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="-6klI5DhSdik" outputId="8cb0ca55-176b-4d2f-8891-2c124276e429" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q', scale=alt.Scale(zero=False)), # 스케일 맵핑 설정
alt.Y('life_expect:Q', scale=alt.Scale(zero=False)) # 스케일 맵핑 설정
)
# + [markdown] colab_type="text" id="oUKf6WQmSzqC" slideshow={"slide_type": "subslide"}
# - 이렇게 하면, 축 스케일에 원점이 포함되지 않는다.
# - 그래도 일부 내부 여백(padding)이 여전히 포함되는데,
# 5 또는 10 등의 배수 수치로 눈금선을 멋지게(nice) 처리하기 위한 것이다.
# - `nice=False` 인자를 추가로 지정할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="-6klI5DhSdik" outputId="8cb0ca55-176b-4d2f-8891-2c124276e429" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q', scale=alt.Scale(zero=False, nice=False)), # 스케일 맵핑 설정
alt.Y('life_expect:Q', scale=alt.Scale(zero=False, nice=False)) # 스케일 맵핑 설정
)
# + [markdown] colab_type="text" id="hKCsQ0iIPMTV" slideshow={"slide_type": "slide"}
# ### 3.3 크기
# + [markdown] colab_type="text" id="G7YtT-HNdYT9" slideshow={"slide_type": "fragment"}
# - `size` 인코딩 채널은 마크의 크기를 설정한다.
# - 채널의 의미는 마크 유형에 따라 달라진다.
# - `point` 마크라면, `size` 채널은 플롯팅 심볼의 픽셀 영역을 의미한다.
# - 점의 지름은 `size` 값의 제곱근으로 설정된다.
# - 앞서 작성했던 산점도에 대하여 인구 규모 `pop` 필드를 `size` 채널로 인코딩해 보자.
# 차트에는 크기에 대한 범례가 추가된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="sGENuXRfOIPz" outputId="4f76557c-4e35-41fc-9e06-513be34969ae" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q')
)
# + [markdown] colab_type="text" id="g5Cu5F2Ad9xG" slideshow={"slide_type": "subslide"}
# - 어떤 상황에서는 기본적으로 지정되는 크기가 마음에 들지 않을 수 있다.
# - 크기를 맞춤형으로 지정하려면,
# `scale` 속성에 대하여 `range` 매개변수를
# 크기에 대한 최소값과 최대값의 배열로 지정할 수 있다.
# - 아래에서는 점의 크기를 최소값은 0 픽셀로, 최대값은 1,000 픽셀로 지정하였다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="cALgZSQfQzi0" outputId="0d88f561-9737-43fa-a84a-7ad6dcdfbf0c" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0, 1000]))
)
# + [markdown] colab_type="text" id="yF__0zZ0PJyT" slideshow={"slide_type": "slide"}
# ### 3.4 색상과 투명도
# + [markdown] colab_type="text" id="u5QXPLxJeohH" slideshow={"slide_type": "subslide"}
# - `color` 인코딩 채널은 마크 색상을 설정한다.
# 색상 인코딩의 스타일은 데이터 유형에 의존적이다.
# - 명목형 데이터에 대한 색상은 다채로운 색상의 질적 색채도로 처리된다.
# - 서수형 및 정량형 데이터에 대한 색상은 지각 가능도록 순서화된 색상 그래디언트(gradient)로 처리된다.
# - 여기서는 `color` 채널을 명목형(`N`) 데이터 유형의 `cluster` 필드로 지정하였다.
# - 클러스터에 대한 범례가 추가되었다.
# - 클러스터 필드의 의미를 짐작할 수 있는가?
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="bI73XTt5OTmA" outputId="f12aeb57-07c5-422e-8cb1-62253b545af8" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N')
)
# + [markdown] colab_type="text" id="dNOkUJfNfFac" slideshow={"slide_type": "subslide"}
# - 원의 내부가 색상으로 채워지기를 바란다면,
# `mark_point()` 메소드에서 `filled=True` 속성을 지정하면 된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="HH34VMCAQO6s" outputId="8aba4f13-bfe3-43b1-e3ea-554a8d20b7f5" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N')
)
# + [markdown] colab_type="text" id="YY96rt7zfRYG" slideshow={"slide_type": "subslide"}
# - 기본적으로, 알테어는 과도한 겹침 현상을 완화하기 위하여 투명도를 활용한다.
# 투명도를 맞춤형으로 조절하려면,
# - `mark_*` 메소드에 대하여 기본값을 지정하거나,
# - 인코딩 메소드에서 `OpacityValue` 값을 설정해야 한다.
# - 아래에서는 인코드 메소드에서 `OpacityValue` 값을 데이터 필드가 아닌 상수로 지정하였다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="42kM5nuDOfGm" outputId="2dd14842-8fd5-4997-ea47-91cd489e0193" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5)
)
# + [markdown] colab_type="text" id="zW2Siu5sPaiE" slideshow={"slide_type": "slide"}
# ### 3.5 모양
# + [markdown] colab_type="text" id="MGM2hdOqf3Tu" slideshow={"slide_type": "fragment"}
# - `shape` 인코딩 채널은 `point` 마크에 대한 기하학적 모양을 설정한다.
# - 이제껏 공부한 다른 채널과 달리, `shape` 채널은 다른 마크 유형에 대해서는 사용할 수 없다.
# - 모양 인코딩 채널은 명목형 데이터에 대해서만 사용 가능하며,
# 서수형 및 정량형 비교에 대해서는 적용이 불가능하다.
# - `cluster` 필드를 `shape` 및 `color` 채널에 지정해 보자.
# - 단일 필드에 대하여 복수 채널을 지정하는 방식을 *다중 인코딩*(redundant encoding)이라고 한다.
# - 시각화 결과에는 색상과 모양이 결합된 단일 심볼이 범례에 추가된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="czAIeWjSPS5x" outputId="e418f006-988c-42a9-9a81-664d1d52d93c" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="ramTzfIgUtEi" slideshow={"slide_type": "slide"}
# ### 3.6 툴팁 및 순서
# + [markdown] colab_type="text" id="VE-29r8dg1Us" slideshow={"slide_type": "fragment"}
# - 아직까지 시각화된 점이 어느 나라인지를 짐작하기는 쉽지 않다.
# 상호작용 방식으로 볼 수 있는 툴팁으로 나라 이름을 확인할 수 있도록 해보자.
# - `tooltip` 인코딩 채널은 사옹자가 마우스 커서를 특정 마크에 올릴 때 보여줄 툴팁 텍스트를 지정한다.
# `country` 필드에 대한 툴팁 인코딩을 추가하자.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="61evgaeFUyU0" outputId="f13ddd14-3b6f-4f96-ca08-8d3b2ba94a34" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5),
alt.Tooltip('country')
)
# + [markdown] colab_type="text" id="IRexn5IAhgeW" slideshow={"slide_type": "subslide"}
# - 마우스로 탐사해보면, 특정 점에 대한 툴팁을 확인할 수 없는 경우가 있다.
# - 예를 들어서 다크 블루 색상에서 제일 큰 원이 인도이다.
# - 이 원이 인구가 더 작은 어느 나라의 원을 가리고 있고,
# 해당 국가의 정보를 확인할 수가 없다.
# - `order` 인코딩 채널로 이 문제를 해결할 수 있다.
# - `order` 인코딩 채널은 데이터 점의 화면 표시 순서를 결정한다.
# 이 채널은 `line` 및 `area` 마크의 표시 순서를 동시에 결정한다.
# - 차트에서 원이 표시되는 순서를 인구 `pop` 열의 내림차순으로 설정해서,
# 작은 원이 큰 원보다 나중에 그려지도록 조정하자.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="DywIzwtgVD3Z" outputId="3b7254de-09f3-4850-f066-ea8318ec5479" slideshow={"slide_type": "fragment"}
chart = alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5),
alt.Tooltip('country:N'),
alt.Order('pop:Q', sort='descending')
)
# + [markdown] colab_type="text" id="Ei93j-daiG6U" slideshow={"slide_type": "subslide"}
# - 이제 작은 원으로 표시된 국가 이름을 확인할 수 있다.
# 인도 앞의 작은 원으로 표시된 국가는 방글라데시이다.
# - 이제 나라 이름을 확인해보면, `cluster`에 의하여 구분된 색상의 의미를 짐작할 수 있다.
# + slideshow={"slide_type": "fragment"}
chart
# + [markdown] colab_type="text" id="9bsu1tn_idGt" slideshow={"slide_type": "subslide"}
# - 현재 툴팁은 데이터 레코드에 대한 단 한 가지 속성만을 표시하고 있다.
# - 여러 값을 동시에 보여주려면, `tooltip` 채널에 대하여 배열로 원하는 여러 필드를 지정할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 372} colab_type="code" id="450DUk-mXzzQ" outputId="ee47eb3b-146d-4c6b-dfa7-94ddeb0fc5d3" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5),
alt.Order('pop:Q', sort='descending'),
tooltip = [
alt.Tooltip('country:N'),
alt.Tooltip('fertility:Q'),
alt.Tooltip('life_expect:Q')
]
)
# + [markdown] colab_type="text" id="HLQjhEdwiyR2" slideshow={"slide_type": "fragment"}
# - 이제 마우스를 데이터 포인트에 올려서 여러 데이터 필드의 내용을 확인할 수 있다.
# + [markdown] colab_type="text" id="5ViS2LDKRmil" slideshow={"slide_type": "slide"}
# ### 3.7 열/행 다면 구성
# + [markdown] colab_type="text" id="apPtlcDoi3yT" slideshow={"slide_type": "fragment"}
# - 공간적 위치야말로 시각적 인코딩을 위한 가장 강력하고도 유연한 채널이다.
# - 그런데 이미 `x` 및 `y` 채널에 데이터 필드를 할당한 상태라면,
# 데이터의 부분 집합마다 하위 플롯을 별도로 시각화하는 *격자 플롯*(trellis plot)을 생성할 수 있다.
# - 격자 플롯은 뷰를 작은 여러 개로([small multiples](https://en.wikipedia.org/wiki/Small_multiple)) 구성하는 기법의 한 사례이다.
# - `column` 및 `row` 인코딩 채널은 수평적이거나 수직적인 하위 플롯의 집합을 생성하도록 설정해 준다.
# - 지정한 데이터 필드 값에 근거하여 데이터를 분할하여 하위 플롯을 다수 생성한다.
# - 여기서는 `cluster` 값마다 열을 구분하여 하위 플롯을 격자 플롯 형태로 만들어 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 441} colab_type="code" id="gszjktorPmRc" outputId="b62aff8e-a2aa-409a-be1d-4cf3708f9ab3" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000])),
alt.Color('cluster:N'),
alt.OpacityValue(0.5),
alt.Tooltip('country:N'),
alt.Order('pop:Q', sort='descending'),
alt.Column('cluster:N')
)
# + [markdown] colab_type="text" id="8J0dxQoFj9jw" slideshow={"slide_type": "subslide"}
# - 직전 플롯은 너무 작아서 알아보기 어렵다.
# - 차트의 `width` 및 `height` 속성을 `properties()` 메소드에서 설정할 수 있다.
# - 열 헤더에 `cluster` 값이 표시되기 때문에 이에 대한 범례를 제거하기 위해서
# `Color` 인코딩 채널에서 `legend=None`으로 지정한다.
# - 또한 `Size` 인코딩 채널에서 `legend` 속성에 대하여
# `orient='bottom'` 및 `titleOrient='left'`로 지정한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 321} colab_type="code" id="PvKBL59lRrc_" outputId="3eb84b63-0c69-4985-ee9b-d0d69e5ffb33" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True).encode(
alt.X('fertility:Q'),
alt.Y('life_expect:Q'),
alt.Size('pop:Q', scale=alt.Scale(range=[0,1000]),
legend=alt.Legend(orient='bottom', titleOrient='left')),
alt.Color('cluster:N', legend=None),
alt.OpacityValue(0.5),
alt.Tooltip('country:N'),
alt.Order('pop:Q', sort='descending'),
alt.Column('cluster:N')
).properties(width=135, height=135)
# + [markdown] colab_type="text" id="OmQmca9WkKOU" slideshow={"slide_type": "subslide"}
# - `column` 및 `row` 인코딩을 지정하면, 배후에서 이들이 `facet` 뷰 구성을 사용하는 새로운 명세로 변환된다.
# `facet`에 대해서는 나중에 더 자세히 공부할 기회를 가지겠다.
#
# + [markdown] colab_type="text" id="-tZXDmLjbAu5" slideshow={"slide_type": "slide"}
# ### 3.8 상호작용적 필터링
#
# - 데이터 탐색을 위한 상호작용성에 대해서는 나중에 더 자세히 공부할 예정이다.
# - `year` 필드에 범위 슬라이더를 연결하여 각 연도에 대한 상호작용적 문질러 닦기(scrubbing)를 구현할 수 있다.
# - 아래 코드가 다소 복잡해 보이겠지만, 나중에 상호작용성을 자세히 공부할 예정이므로 너무 걱정할 필요는 없다.
# - 연도 슬라이더를 앞뒤로 드래그하여 시간에 따라서 변화하는 데이터 값을 감상하라!
# + colab={"base_uri": "https://localhost:8080/", "height": 393} colab_type="code" id="6kQLe1wqZPub" outputId="3c934853-9167-4ad1-e560-b366250eb40e" slideshow={"slide_type": "subslide"}
select_year = alt.selection_single(
name='select', fields=['year'], init={'year': 1955},
bind=alt.binding_range(min=1955, max=2005, step=5)
)
alt.Chart(data).mark_point(filled=True).encode(
alt.X('fertility:Q', scale=alt.Scale(domain=[0,9])),
alt.Y('life_expect:Q', scale=alt.Scale(domain=[0,90])),
alt.Size('pop:Q', scale=alt.Scale(domain=[0, 1200000000], range=[0,1000])),
alt.Color('cluster:N', legend=None),
alt.OpacityValue(0.5),
alt.Tooltip('country:N'),
alt.Order('pop:Q', sort='descending')
).add_selection(select_year).transform_filter(select_year)
# + [markdown] colab_type="text" id="UPdDrmm9YK5E" slideshow={"slide_type": "slide"}
# ## 4. 그래픽 마크
#
# - 지금까지 인코딩 채널에 대해 공부하면서 `point` 마크에만 집중했었다.
# 이 외에도 다양한 기하학적 모양의 마크를 사용할 수 있다.
# - `mark_area()` - 상단 선과 기저 선으로 정의되는 채워진 영역
# - `mark_bar()` - 사각형 막대
# - `mark_circle()` - 채워진 원으로 표시되는 산점도
# - `mark_line()` - 연결된 꺽어진 선
# - `mark_point()` - 모양을 설정할 수 있는 산점도
# - `mark_rect()` - 열지도(heatmap)등에 사용되는 채워진 사각형
# - `mark_rule()` - 축을 가로지르는 수직선 또는 수평선
# - `mark_square()` - 사각형 점으로 표시되는 산점도
# - `mark_text()` - 텍스트로 표신하는 산점도
# - `mark_tick()` - 수직 또는 수평 눈금
# - 그래픽 마크에 대한 전체 목록과 예제를 확인하려면 [Altair marks documentation](https://altair-viz.github.io/user_guide/marks.html)를 보라.
# - 아래에서 통계적 시각화에서 자주 사용하는 마크 유형을 공부하자.
# + [markdown] colab_type="text" id="jVp7cC-BbYF5" slideshow={"slide_type": "slide"}
# ### 4.1 점 마크
#
# - `point` 마크 유형은 *산점도 scatter plots* 및 *점 플롯 dot plots*과 같이 점을 시각화 한다.
# - 점 마크에서는 기본적으로 2차원 위치를 표시하기 위하여 `x`와 `y` 인코딩 채널을 지정한다.
# - 아울러 색상 `color`, 크기 `size` 및 모양 `shape` 인코딩 채널에 대하여 추가적인 데이터 필드를 지정할 수 있다.
# - 아래는 출산율 `fertility`을 x-축 인코딩 채널로,
# 지역을 의미하는 `cluster` 필드를 y-축 및 모양 `shape` 인코딩 채널로 지정한 점 플롯(dot plot)이다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="3jrFfaKCtbyL" outputId="6332a348-f8f0-4dd8-9c4a-63687e286d4a" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point().encode(
alt.X('fertility:Q'),
alt.Y('cluster:N'),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="bONM4Ccrt4Sw" slideshow={"slide_type": "subslide"}
# - 인코딩 채널 외에도,
# `mark_*()` 메소드에서 여러 옵션 값을 지정하여 마크 스타일을 지정할 수 있다.
# 점 마크는 기본적으로 외곽선으로 그려지는데,
# - `filled` 옵션으로 색상이 채워진 모습으로 도형을 표시하거나,
# - `size` 옵션으로 도형의 크기를 지정할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="3-HgJ1PjvKdH" outputId="dcf26fa9-281b-4297-da50-84c7f5ccd318" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_point(filled=True, size=200).encode(
alt.X('fertility:Q'),
alt.Y('cluster:N'),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="CGMm4gyNcLzB" slideshow={"slide_type": "slide"}
# ### 4.2 원 마크
#
# - `circle` 마크 유형은 내부가 채워진 원 형태로 시각화하기 위하여 `point` 마크 대신에 사용된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="R7JgFhB-vn5X" outputId="80cd9d0f-c126-4673-bfcf-eaf5b34e0058" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_circle(size=100).encode(
alt.X('fertility:Q'),
alt.Y('cluster:N'),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="IAZSJlWhcOr_" slideshow={"slide_type": "slide"}
# ### 4.3 정사각형 마크
#
# - `square` 마크 유형은 내부가 채워진 정사각형 형태로 시각화하기 위하여 `point` 마크 대신에 사용된다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="rfby5R-Mvuwx" outputId="8ff4b0d4-f3ce-45e5-f597-1a6fd296ba47" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_square(size=100).encode(
alt.X('fertility:Q'),
alt.Y('cluster:N'),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="SJF0vNqGcRdv" slideshow={"slide_type": "slide"}
# ### 4.4 눈금 마크
#
# - `tick` 마크 유형은 데이터 포인트를 짧은 선분 형태의 눈금(tick)으로 시각화한다.
# - 이러한 시각화는 일차원 직선 상에서 겹치는 부분을 최소화 한 형태로 값을 비교할 때 유용하다.
# - 눈금 마크로 그려진 점 플롯을 때로 *스트립 플롯 strip plot*으로 부르기도 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="Thvp_VJ2v-xd" outputId="c6133250-333f-4c29-e217-221634dd8337" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_tick().encode(
alt.X('fertility:Q'),
alt.Y('cluster:N'),
alt.Shape('cluster:N')
)
# + [markdown] colab_type="text" id="x8j5XVZJcB7l" slideshow={"slide_type": "slide"}
# ### 4.5 막대 마크
#
# - `bar` 마크 유형은 직사각형의 긴 막대 모양으로 시각화한다.
# - 아래에서는 국가별 인구 `pop` 필드에 대한 단순한 막대 차트를 시각화 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 445} colab_type="code" id="eFN9Kw85xFQK" outputId="8da431d3-9ba7-4ecf-de55-e478c4527a39" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_bar().encode(
alt.X('country:N'),
alt.Y('pop:Q')
)
# + [markdown] slideshow={"slide_type": "subslide"}
# - 수직 막대 차트를 수평 막대 차트로 변경하려면,
# x-축과 y-축에 대한 인코딩 필드를 맞바꾸어 준다.
# + colab={"base_uri": "https://localhost:8080/", "height": 445} colab_type="code" id="eFN9Kw85xFQK" outputId="8da431d3-9ba7-4ecf-de55-e478c4527a39" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_bar().encode(
alt.Y('country:N'),
alt.X('pop:Q')
)
# + [markdown] colab_type="text" id="rHX8bKgbzW3I" slideshow={"slide_type": "subslide"}
# - 막대 폭은 기본값으로 자동 설정된다.
# 이를 변경하려면, `x` 채널에 대한 `scale` 속성의 `rangeStep` 옵션을 설정한다.
# 축, 스케일 및 범례에 대한 설정에 대해서는 나중에 자세히 공부할 예정이다.
# - 막대를 적층식으로 쌓아서 시각화 할 수 있다.
# `x` 인코딩을 지역 `cluster` 필드로 수정하고, 국가 `country` 필드를 `color` 채널로 인코딩하여 보자.
# 또한 국가에 대한 범례는 생략하도록 지정하고, 국가 이름을 툴팁으로 처리하자.
# + colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="code" id="x9maAWhOxsrq" outputId="7960be03-eefd-4c3f-a114-8405395c772c" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_bar().encode(
alt.X('cluster:N'),
alt.Y('pop:Q'),
alt.Color('country:N', legend=None),
alt.Tooltip('country:N')
)
# + [markdown] colab_type="text" id="de4181jhzgUx" slideshow={"slide_type": "fragment"}
# - 위 차트에서, `color` 인코딩 채널이 알테어/베가-라이트로 하여금 막대 마크를 자동으로 쌓아 올리도록 만들어 준다.
# - 만일 적층식이 싫다면, 아래와 같이
# - `y` 인코딩 채널에 대해서 `stack=None` 옵션을 추가 지정하라.
# 이렇게 하면, 모든 국가별 막대가 같은 위치에 그려질 것이다.
# - 이때 `order` 인코딩 채널에서 인구의 오름차순으로 순서를 지정해야 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="code" id="x9maAWhOxsrq" outputId="7960be03-eefd-4c3f-a114-8405395c772c" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_bar().encode(
alt.Order('pop:Q', sort='ascending'), # 주석 제거
alt.X('cluster:N'),
alt.Y('pop:Q', stack=None),
alt.Color('country:N', legend=None),
alt.Tooltip('country:N'),
)
# + [markdown] colab_type="text" id="mLSz2vHb2qiV" slideshow={"slide_type": "subslide"}
# - 이전 막대 차트는 영점 기저 선(zero-baseline)을 기준으로 시각화 되었다.
# - `y` 채널은 영 아닌(non-zero) 값만을 막대로 인코드 한다.
# - 원한다면, 막대 마크에서 막대의 시작 및 끝 점을 범위로 지정할 수 있다.
# - 아래 차트는 `x` (시작 점) 및 `x2` (종료 점) 채널을 사용하여 지역별 기대 수명을 보여준다.
# - 여기서는 `min()` 및 `max()` 집계 함수를 사용하여 기대 수명 범위의 시작 값과 끝 값을 처리하였다.
# - 집계 함수에 대해서는 나중에 자세히 공부할 예정이다.
# - 또 다른 방법으로, `x` 및 `width`를 활용하여
# `x2 = x + width`와 같이 시작 점과 오프셋을 지정할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 192} colab_type="code" id="NxEVEQLj0l9t" outputId="2fa67514-b66c-4449-a1e3-c0b1b377ed3a" slideshow={"slide_type": "fragment"}
alt.Chart(data2000).mark_bar().encode(
alt.X('min(life_expect):Q'),
alt.X2('max(life_expect):Q'),
alt.Y('cluster:N')
)
# + [markdown] colab_type="text" id="bTnEQk1NcD7y" slideshow={"slide_type": "slide"}
# ### 4.6 선 마크
#
# - `line` 마크 유형은 그려진 점들을 선분으로 연결한다.
# 선의 기울기로 변화율 정보를 보여줄 수 있다.
# - 국가별 출산율을 연도에 따라서 꺽은선 차트로 시각화하자.
# - 필터링되지 않은 세계 개발 데이터프레임 전체를 사용하자.
# - 범례는 숨기고 툴팁을 활용하자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="lTWmDwv1Jhzt" outputId="7bbd15a9-4b8f-4f66-fe7e-6a55f2c38449" slideshow={"slide_type": "fragment"}
alt.Chart(data).mark_line().encode(
alt.X('year:O'),
alt.Y('fertility:Q'),
alt.Color('country:N', legend=None),
alt.Tooltip('country:N')
).properties(
width=600
)
# + [markdown] colab_type="text" id="qmV54UhSMPxU" slideshow={"slide_type": "subslide"}
# - 국가마다 보여주는 변화가 흥미롭다.
# - 하지만 전반적인 추세는 시간이 갈수록 출산율이 낮아진다는 점이다.
# - 차트 폭을 변경하였는데, 이 부분을 제거하면 어떻게 되는지 확인해 보라.
# - 맞춤형 시각화를 위하여 마크 매개변수에 대한 기본값을 변경해 보자.
# - `strokeWidth`를 설정하여 선의 굵기를 조절할 수 있다.
# - `opacity`를 설정하여 투명도를 조절할 수 있다.
# - `line` 마크는 기본적으로 꺽은선을 사용하는데, 부드러운 곡선으 사용하도록 조절할 수 있다.
# - `interpolate` 마크 매개변수를 설정하여 보간법을 적용할 수 있다.
# - `'monotone'` 보간법을 적용해 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="PbdSZMs43rgW" outputId="07007338-a2c7-4dc2-9f36-bb344e4103ec" slideshow={"slide_type": "fragment"}
alt.Chart(data).mark_line(
strokeWidth=3, # 선 굵기
opacity=0.5, # 투명도
interpolate='monotone' # 보간법
).encode(
alt.X('year:O'),
alt.Y('fertility:Q'),
alt.Color('country:N', legend=None),
alt.Tooltip('country:N')
).properties(
width=600
)
# + [markdown] colab_type="text" id="_PZcXCF6OZm2" slideshow={"slide_type": "subslide"}
# - `line` 마크를 사용하여 *기울기 그래프 slope graphs*를 시각화 할 수 있다.
# 이를 통해 두 비교 점 간의 변화를 기울기로 보여주는 시각화가 가능하다.
# - 모든 국가의 인구 변화를 기울기로 보여주는 기울기 그래프를 시각화 해 보자.
# - 먼저 판다스 데이터프레임에서 1955년과 2005년에 해당하는 데이터 만을 추출하자.
# - 기본값으로 그리면, 두 연도에 해당하는 x-축 눈금이 너무 근접하게 되므로,
# 폭과 관련한 속성을 `width={"step": 100}`으로 조정하자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="glipk3XoKUcc" outputId="edd1292a-97a7-44d1-b2ec-c5b8d755aa86" slideshow={"slide_type": "fragment"}
dataTime = data.loc[(data['year'] == 1955) | (data['year'] == 2005)]
alt.Chart(dataTime).mark_line(opacity=0.5).encode(
alt.X('year:O'),
alt.Y('pop:Q'),
alt.Color('country:N', legend=None),
alt.Tooltip('country:N')
).properties(
width={"step": 100} # adjust the step parameter
)
# + [markdown] colab_type="text" id="LLo0NmuncGAa" slideshow={"slide_type": "slide"}
# ### 4.7 면적 마크
#
# - `area` 마크 유형은 꺽은선 `line` 마크와 막대 `bar` 마크를 결합한 형태이다.
# 데이터 포인트 간의 기울기와 채워진 영역을 함께 보여준다.
# - 아래 차트는 우리 나라에 대해서, 연도에 따른 출산율의 변화를 면적 차트로 보여준다.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="cXdOREmBHiaH" outputId="37cd899d-2ad1-42ea-fbe0-12e591a28326" slideshow={"slide_type": "fragment"}
dataUS = data.loc[data['country'] == 'South Korea']
alt.Chart(dataUS).mark_area().encode(
alt.X('year:O'),
alt.Y('fertility:Q')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="aDPZ6bWEPv2_" slideshow={"slide_type": "subslide"}
# - `line` 마크와 마찬가지로 `area` 마크도 `interpolate` 매개변수를 지원한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="-ZpfjqMxJvXF" outputId="eba5baab-44be-46ac-c9ae-0ad3b094573a" slideshow={"slide_type": "fragment"}
alt.Chart(dataUS).mark_area(interpolate='monotone').encode(
alt.X('year:O'),
alt.Y('fertility:Q')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="_YolZaDbP7jh" slideshow={"slide_type": "subslide"}
# - `bar` 마크와 유사하게, `area` 마크도 쌓기(stacking)를 지원한다.
# - 북미 3국에 대한 데이터프레임을 새롭게 생성한다.
# - `area` 마크와 `color` 인코딩 채널을 이용하여 나라별로 쌓아 올린 시각화를 완성해 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="tYOS0Il_H8h3" outputId="70e210aa-4771-4ed9-e5a3-0af587d03c60" slideshow={"slide_type": "fragment"}
dataNA = data.loc[
(data['country'] == 'United States') |
(data['country'] == 'Canada') |
(data['country'] == 'Mexico')
]
alt.Chart(dataNA).mark_area().encode(
alt.X('year:O'),
alt.Y('pop:Q'),
alt.Color('country:N')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="oVinPfTZpNtY" slideshow={"slide_type": "subslide"}
# - 기본적으로, 쌓아올리기는 영점 기저선(zeo baseline)에 대한 상대값으로 처리되는데,
# 다른 `stack` 옵션을 지정할 수 있다.
# * `center` - 차트의 중앙을 기저선으로 지정하여, *스트림 그래프 streamgraph*로 시각화 할 수 있다.
# * `normalize` - 데이터의 합계 값을 정규화하여 전체가 100%가 되면서, 각 요소의 비중을 백분율로 시각화 할 수 있다.
# - 아래에서는 `y` 인코딩에 `stack` 속성을 `center`로 지정하여 시각화 해 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="6SQpyJzqpB5G" outputId="3a7cb75a-3acf-4b0e-d0e8-cdc7c85fd5ee" slideshow={"slide_type": "fragment"}
alt.Chart(dataNA).mark_area().encode(
alt.X('year:O'),
alt.Y('pop:Q', stack='center'),
alt.Color('country:N')
).properties(
width={"step": 50}
)
# + [markdown] slideshow={"slide_type": "subslide"}
# - 이번에는 y 인코딩에 `stack` 속성을 `normalize`로 지정하여 시각화 해 보자.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="6SQpyJzqpB5G" outputId="3a7cb75a-3acf-4b0e-d0e8-cdc7c85fd5ee" slideshow={"slide_type": "fragment"}
alt.Chart(dataNA).mark_area().encode(
alt.X('year:O'),
alt.Y('pop:Q', stack='normalize'),
alt.Color('country:N')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="fQN5a3gfQFBW" slideshow={"slide_type": "subslide"}
# - 쌓아올리기를 모두 취소하려면, `stack` 속성을 `None`으로 설정하라.
# - 이렇게 되면 모든 면적 요소가 영점 기저선에서부터 그려진다.
# - 또한 `opacity`를 `area` 마크에 대한 기본 매개변수로 지정하여, 겹쳐진 영역을 확인할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="MY8xDiqkKDMc" outputId="41345e53-eeaa-4c69-969d-738c1479ba26" slideshow={"slide_type": "fragment"}
alt.Chart(dataNA).mark_area(opacity=0.5).encode(
alt.X('year:O'),
alt.Y('pop:Q', stack=None),
alt.Color('country:N')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="ro21p2rsQMl3" slideshow={"slide_type": "subslide"}
# - `area` 마크 유형도 데이터 중심의 기저선을 지원한다.
# - 데이터 필드에 의하여 계산한 상한 및 하한 시리즈를 적용할 수 있다.
# - `bar` 마크에서 처럼, `x` 및 `x2` (또는 `y` 및 `y2`) 채널을 통하여 `area` 마크의 종료 점을 표시할 수 있다.
# - 아래 차트는 북미 3국에 대하여 연도별 출산율의 최소값과 최대값 범위를 시각화 한 결과이다.
# + colab={"base_uri": "https://localhost:8080/", "height": 385} colab_type="code" id="I084oc4aQeMw" outputId="fb6d0868-79f4-4f96-8f5d-aff6d504b3fe" slideshow={"slide_type": "fragment"}
alt.Chart(dataNA).mark_area().encode(
alt.X('year:O'),
alt.Y('min(fertility):Q'),
alt.Y2('max(fertility):Q')
).properties(
width={"step": 50}
)
# + [markdown] colab_type="text" id="msL27DW2qoqJ" slideshow={"slide_type": "fragment"}
# - 1955년에는 4에 육박하는 최소값과 7에 근접하는 최대값을 가질 정도로 출산율의 범위가 넓었다.
# - 그러나 시간이 지날수록 출산율이 낮아져서, 2005년에는 가족당 2 자녀 수준이 되었다.
# + [markdown] colab_type="text" id="5BoLjQCgrHZL" slideshow={"slide_type": "subslide"}
# - 지금까지 살펴 본 모든 `area` 마크 예제에서는 수직으로 범위를 가지는 면적을 시각화 하였다.
# - 알테어 및 베가-라이트는 수평으로 범위를 가지는 면적의 시각화도 가능하다.
# - 앞서 그린 차트에서 `x`와 `y` 채널을 교환함으로써 차트를 전치(transpose)할 수 있다.
# + colab={"base_uri": "https://localhost:8080/", "height": 506} colab_type="code" id="l_TDR4SNrStr" outputId="e7bf35e5-c94b-4316-9f8d-0dc68f242763" slideshow={"slide_type": "fragment"}
alt.Chart(dataNA).mark_area().encode(
alt.Y('year:O'),
alt.X('min(fertility):Q'),
alt.X2('max(fertility):Q')
).properties(
height=400,
width=600
)
# + [markdown] colab_type="text" id="OAmIxLRarios" slideshow={"slide_type": "slide"}
# ## 5. 요약
#
# - 지금까지 데이터 유형, 인코딩 채널, 그리고 그래픽 마크를 공부했다.
# - 이제부터는 스스로 인코딩, 마크 유형 및 마크 매개변수에 대한 공부를 진행할 수 있을 것이다.
# - 여기에서 생략한 전체 내용을 확인하기 위해서는 다음 자료를 확인하라.
# - [marks](https://altair-viz.github.io/user_guide/marks.html)
# - [encoding](https://altair-viz.github.io/user_guide/encoding.html)
# - 다음 장에서는 데이터 변환(transformations)에 관하여 공부한다.
# - 데이터 전체를 요약하는 차트를 시각화하는 방법
# - 기존 데이터에 대한 계산을 통하여 유도한 새로운 데이터 필드로 시각화하는 방법
# - 이후에는 스케일, 축 및 범례를 맞춤형으로 수정하는 방법을 공부한다.
# + [markdown] colab_type="text" id="OAmIxLRarios" slideshow={"slide_type": "subslide"}
# - 시각적 인코딩에 대하여 흥미가 커졌길 기대한다.
# + [markdown] slideshow={"slide_type": "fragment"}
# <img title="Bertin's Taxonomy of Visual Encoding Channels" src="https://cdn-images-1.medium.com/max/2000/1*jsb78Rr2cDy6zrE7j2IKig.png" style="max-width: 650px;"><br/>
#
# <small><a href="https://books.google.com/books/about/Semiology_of_Graphics.html?id=X5caQwAACAAJ"><em>Sémiologie Graphique</em></a>에서 발췌한 Bertin'의 시각적 인코딩에 대한 분류를 <a href="https://bost.ocks.org/mike/">Mike Bostock</a>가 편집한 내용</small>
# + [markdown] slideshow={"slide_type": "subslide"}
# - 마크, 시각적 인코딩 및 데이터 유형에 관한 체계적 연구는 [Jacques Bertin](https://en.wikipedia.org/wiki/Jacques_Bertin)에 의하여, 선구자적이라 할 수 있는 1967년의 [_Sémiologie Graphique (The Semiology of Graphics)_](https://books.google.com/books/about/Semiology_of_Graphics.html?id=X5caQwAACAAJ) 작업에서 시작되었다. 앞에서 제시한 이미지는 위치, 크기, 값(명도), 질감, 색조, 방향성 및 모양이라는 채널의 개념을 이들이 지원하는 데이터 유형에 대하여 Bertin이 추천한 내용과 함께 정리한 것이다.
# - 데이터 유형, 마크 및 채널이라는 체계는 또한 자동화된 시각화 설계 도구에 대하여 지침을 제공한다. 이러한 내용은 1986년에 제시된 [Mackinlay's APT (A Presentation Tool)](https://scholar.google.com/scholar?cluster=10191273548472217907)에서 시작되어, [Voyager](http://idl.cs.washington.edu/papers/voyager/) 및 [Draco](http://idl.cs.washington.edu/papers/draco/)와 같은 최근의 시스템으로 이어지고 있다.
# - 명목형, 서수형, 구간 및 비율 척도에 대한 이론은 최소한 S. S. Steven의 1947년 논문인 [_On the theory of scales of measurement_](https://scholar.google.com/scholar?cluster=14356809180080326415)까지 그 기원을 거슬러 올라갈 수 있다.
| .ipynb_checkpoints/02_altair_marks_encoding-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
### IMPORTS ###
import numpy as np
###########INTEGRATOR FRAMEWORK############
#1 Constraint Checks
def check_type(y,t): # Ensure Input is Correct
return y.dtype == np.floating and t.dtype == np.floating
class _Integrator():
def __init__(self,n_,F_b):
self.n_ = n_
self.F_b = F_b
def integrate(self,func,y0,t):
time_delta_grid = t[1:] - t[:-1]
y = np.zeros((y0.shape[0],t.shape[0]))
y[:,0] = y0
for i in range(time_delta_grid.shape[0]):
k1 = func(y[:,i], t[i]) # RK4 Integration Steps
half_step = t[i] + time_delta_grid[i] / 2
k2 = func(y[:,i] + time_delta_grid[i] * k1 / 2, half_step)
k3 = func(y[:,i] + time_delta_grid[i] * k2 / 2, half_step)
k4 = func(y[:,i] + time_delta_grid[i] * k3, t + time_delta_grid[i])
dy = (k1 + 2 * k2 + 2 * k3 + k4) * (time_delta_grid[i] / 6)
out = dy + y[:,i]
ft = y[-n_:,i]
l = np.zeros(ft.shape)
l_ = t[i]-ft
z = y[:n_,i] < F_b
z_ = out[:n_] > F_b
df = np.where(np.logical_and(z,z_),[l_,l])
ft_ = ft+df
y[:,i+1] = np.concatenate([out[:-n_],ft_],0)
return y
def odeint_fixed(func,y0,t):
y0 = np.array(y0)
t = np.array(t)
if check_type(y0,t):
return _Integrator().integrate(func,y0,t)
else:
print("error encountered")
# +
import numpy as np
import nerveflow as nv
import time
import sys
###########SIMULATION FRAMEWORK############
scale = int(sys.argv[2])/120 # simulation scale
sim_time = 1000000 # max simulation time
sim_res = float(sys.argv[3]) # simulation resolution
n_n = int(120*scale) # number of neurons
p_n = int(90*scale) # number of PNs
l_n = int(30*scale) # number of LNs
#t = np.arange(0.0, sim_time, 0.01) # duration of simulation
t = np.load("time.npy",allow_pickle=True)[int(sys.argv[1])] # duration of simulation
C_m = [1.0]*n_n # Capacitance
# Common Current Parameters #
g_K = [10.0]*n_n # K conductance
g_L = [0.15]*n_n # Leak conductance
g_KL = [0.05]*p_n + [0.02]*l_n # K leak conductance
E_K = [-95.0]*n_n # K Potential
E_L = [-55.0]*p_n + [-50.0]*l_n # Leak Potential
E_KL = [-95.0]*n_n # K Leak Potential
# Type Specific Current Parameters #
## PNs
g_Na = [100.0]*p_n # Na conductance
g_A = [10.0]*p_n # Transient K conductance
E_Na = [50.0]*p_n # Na Potential
E_A = [-95.0]*p_n # Transient K Potential
## LNs
g_Ca = [3.0]*l_n # Ca conductance
g_KCa = [0.3]*l_n # Ca dependent K conductance
E_Ca = [140.0]*l_n # Ca Potential
E_KCa = [-90]*l_n # Ca dependent K Potential
A_Ca = 2*(10**(-4)) # Ca outflow rate
Ca0 = 2.4*(10**(-4)) # Equilibrium Calcium Concentration
t_Ca = 150 # Ca recovery time constant
# Synaptic Current Parameters #
## Acetylcholine
if sys.argv[1] == '0':
ach_mat = np.zeros((n_n,n_n)) # Ach Synapse Connectivity Matrix
ach_mat[p_n:,:p_n] = np.random.choice([0.,1.],size=(l_n,p_n)) # 50% probability of PN -> LN
np.fill_diagonal(ach_mat,0.) # No self connection
np.save("ach_mat",ach_mat)
else:
ach_mat = np.load("ach_mat.npy")
n_syn_ach = int(np.sum(ach_mat)) # Number of Acetylcholine (Ach) Synapses
alp_ach = [10.0]*n_syn_ach # Alpha for Ach Synapse
bet_ach = [0.2]*n_syn_ach # Beta for Ach Synapse
t_max = 0.3 # Maximum Time for Synapse
t_delay = 0 # Axonal Transmission Delay
A = [0.5]*n_n # Synaptic Response Strength
g_ach = [0.35]*p_n+[0.3]*l_n # Ach Conductance
E_ach = [0.0]*n_n # Ach Potential
## GABAa (fast GABA)
if sys.argv[1] == '0':
fgaba_mat = np.zeros((n_n,n_n)) # GABAa Synapse Connectivity Matrix
fgaba_mat[:,p_n:] = np.random.choice([0.,1.],size=(n_n,l_n)) # 50% probability of LN -> LN/PN
np.fill_diagonal(fgaba_mat,0.) # No self connection
np.save("fgaba_mat",fgaba_mat)
else:
fgaba_mat = np.load("fgaba_mat.npy")
n_syn_fgaba = int(np.sum(fgaba_mat)) # Number of GABAa (fGABA) Synapses
alp_fgaba = [10.0]*n_syn_fgaba # Alpha for fGABA Synapse
bet_fgaba = [0.16]*n_syn_fgaba # Beta for fGABA Synapse
V0 = [-20.0]*n_n # Decay Potential
sigma = [1.5]*n_n # Decay Time Constant
g_fgaba = [0.8]*p_n+[0.8]*l_n # fGABA Conductance
E_fgaba = [-70.0]*n_n # fGABA Potential
# Other Parameters #
F_b = [0.0]*n_n # Fire potential
# Property Dynamics #
def K_prop(V):
T = 22
phi = 3.0**((T-36.0)/10)
V_ = V-(-50)
alpha_n = 0.02*(15.0 - V_)/(np.exp((15.0 - V_)/5.0) - 1.0)
beta_n = 0.5*np.exp((10.0 - V_)/40.0)
t_n = 1.0/((alpha_n+beta_n)*phi)
n_inf = alpha_n/(alpha_n+beta_n)
return n_inf, t_n
def Na_prop(V):
T = 22
phi = 3.0**((T-36)/10)
V_ = V-(-50)
alpha_m = 0.32*(13.0 - V_)/(np.exp((13.0 - V_)/4.0) - 1.0)
beta_m = 0.28*(V_ - 40.0)/(np.exp((V_ - 40.0)/5.0) - 1.0)
alpha_h = 0.128*np.exp((17.0 - V_)/18.0)
beta_h = 4.0/(np.exp((40.0 - V_)/5.0) + 1.0)
t_m = 1.0/((alpha_m+beta_m)*phi)
t_h = 1.0/((alpha_h+beta_h)*phi)
m_inf = alpha_m/(alpha_m+beta_m)
h_inf = alpha_h/(alpha_h+beta_h)
return m_inf, t_m, h_inf, t_h
def A_prop(V):
T = 36
phi = 3.0**((T-23.5)/10)
m_inf = 1/(1+np.exp(-(V+60.0)/8.5))
h_inf = 1/(1+np.exp((V+78.0)/6.0))
tau_m = 1/(np.exp((V+35.82)/19.69) + np.exp(-(V+79.69)/12.7) + 0.37) / phi
t1 = 1/(np.exp((V+46.05)/5.0) + np.exp(-(V+238.4)/37.45)) / phi
t2 = (19.0/phi) * np.ones(np.shape(V))
tau_h = np.where(np.less(V,-63.0),t1,t2)
return m_inf, tau_m, h_inf, tau_h
def Ca_prop(V):
m_inf = 1/(1+np.exp(-(V+20.0)/6.5))
h_inf = 1/(1+np.exp((V+25.0)/12))
tau_m = 1.5
tau_h = 0.3*np.exp((V-40.0)/13.0) + 0.002*np.exp((60.0-V)/29)
return m_inf, tau_m, h_inf, tau_h
def KCa_prop(Ca):
T = 26
phi = 2.3**((T-23.0)/10)
alpha = 0.01*Ca
beta = 0.02
tau = 1/((alpha+beta)*phi)
return alpha*tau*phi, tau
# NEURONAL CURRENTS
# Common Currents #
def I_K(V, n):
return g_K * n**4 * (V - E_K)
def I_L(V):
return g_L * (V - E_L)
def I_KL(V):
return g_KL * (V - E_KL)
# PN Currents #
def I_Na(V, m, h):
return g_Na * m**3 * h * (V - E_Na)
def I_A(V, m, h):
return g_A * m**4 * h * (V - E_A)
# LN Currents #
def I_Ca(V, m, h):
return g_Ca * m**2 * h * (V - E_Ca)
def I_KCa(V, m):
T = 26
phi = 2.3**((T-23.0)/10)
return g_KCa * m * phi * (V - E_KCa)
# SYNAPTIC CURRENTS
def I_ach(o,V):
o_ = np.array([0.0]*n_n**2)
ind = np.arange(n_n**2)[ach_mat.reshape(-1) == 1]
o_[ind] = o
o_ = np.transpose(np.reshape(o_,(n_n,n_n)))
return np.reduce_sum(np.transpose((o_*(V-E_ach))*g_ach),1)
def I_fgaba(o,V):
o_ = np.array([0.0]*n_n**2)
ind = np.arange(n_n**2)[fgaba_mat.reshape(-1) == 1]
o_[ind] = o
o_ = np.transpose(np.reshape(o_,(n_n,n_n)))
return np.reduce_sum(np.transpose((o_*(V-E_fgaba))*g_fgaba),1)
def I_inj_t(t):
return current_input.T[np.cast(t*100,np.int32)]
# DIFFERENTIAL EQUATION FORM
def dAdt(X, t): # X is the state vector
# Assign Current Values
V_p = X[0 : p_n]
V_l = X[p_n : n_n]
n_K = X[n_n : 2*n_n]
m_Na = X[2*n_n : 2*n_n + p_n]
h_Na = X[2*n_n + p_n : 2*n_n + 2*p_n]
m_A = X[2*n_n + 2*p_n : 2*n_n + 3*p_n]
h_A = X[2*n_n + 3*p_n : 2*n_n + 4*p_n]
m_Ca = X[2*n_n + 4*p_n : 2*n_n + 4*p_n + l_n]
h_Ca = X[2*n_n + 4*p_n + l_n: 2*n_n + 4*p_n + 2*l_n]
m_KCa = X[2*n_n + 4*p_n + 2*l_n : 2*n_n + 4*p_n + 3*l_n]
Ca = X[2*n_n + 4*p_n + 3*l_n: 2*n_n + 4*p_n + 4*l_n]
o_ach = X[6*n_n : 6*n_n + n_syn_ach]
o_fgaba = X[6*n_n + n_syn_ach : 6*n_n + n_syn_ach + n_syn_fgaba]
fire_t = X[-n_n:]
V = X[:n_n]
# Evaluate Differentials
n0,tn = K_prop(V)
dn_k = - (1.0/tn)*(n_K-n0)
m0,tm,h0,th = Na_prop(V_p)
dm_Na = - (1.0/tm)*(m_Na-m0)
dh_Na = - (1.0/th)*(h_Na-h0)
m0,tm,h0,th = A_prop(V_p)
dm_A = - (1.0/tm)*(m_A-m0)
dh_A = - (1.0/th)*(h_A-h0)
m0,tm,h0,th = Ca_prop(V_l)
dm_Ca = - (1.0/tm)*(m_Ca-m0)
dh_Ca = - (1.0/th)*(h_Ca-h0)
m0,tm = KCa_prop(Ca)
dm_KCa = - (1.0/tm)*(m_KCa-m0)
dCa = - A_Ca*I_Ca(V_l,m_Ca,h_Ca) - (Ca - Ca0)/t_Ca
CmdV_p = - I_Na(V_p, m_Na, h_Na) - I_A(V_p, m_A, h_A)
CmdV_l = - I_Ca(V_l, m_Ca, h_Ca) - I_KCa(V_l, m_KCa)
CmdV = np.concat([CmdV_p,CmdV_l],0)
dV = (I_inj_t(t) + CmdV - I_K(V, n_K) - I_L(V) - I_KL(V) - I_ach(o_ach,V) - I_fgaba(o_fgaba,V)) / C_m
T_ach = np.where(np.logical_and(np.greater(t,fire_t+t_delay),np.less(t,fire_t+t_max+t_delay)),A,np.zeros(A.shape))
T_ach = np.multiply(ach_mat,T_ach)
T_ach = np.reshape(T_ach,(-1,))[ach_mat.reshape(-1) == 1]
do_achdt = alp_ach*(1.0-o_ach)*T_ach - bet_ach*o_ach
T_fgaba = 1.0/(1.0+np.exp(-(V-V0)/sigma))
T_fgaba = np.multiply(fgaba_mat,T_fgaba)
T_fgaba = np.reshape(T_fgaba,(-1,))[fgaba_mat.reshape(-1) == 1]
do_fgabadt = alp_fgaba*(1.0-o_fgaba)*T_fgaba - bet_fgaba*o_fgaba
dfdt = np.zeros(fire_t.shape)
out = np.concat([dV, dn_k,
dm_Na, dh_Na,
dm_A, dh_A,
dm_Ca, dh_Ca,
dm_KCa,
dCa, do_achdt,
do_fgabadt, dfdt ],0)
return out
current_input = np.load("current.npy")
if sys.argv[1] == '0':
state_vector = [-70]* n_n + [0.0]* n_n + [0.0]* (4*p_n) + [0.0]* (3*l_n) + [2.4*(10**(-4))]*l_n + [0]*(n_syn_ach) + [0]*(n_syn_fgaba) + [-(sim_time+1)]*n_n
state_vector = np.array(state_vector)
state_vector = state_vector + 0.01*state_vector*np.random.normal(size=state_vector.shape)
np.save("state_vector",state_vector)
else:
state_vector = np.load("state_vector.npy")
#print("Number of Neurons:",n_n)
#print("Number of Synapses:",(n_syn_ach+n_syn_fgaba))
#print(n_n,(n_syn_ach+n_syn_fgaba))
n_batch = 1
t_batch = np.array_split(t,n_batch)
t_ = time.time()
for n,i in enumerate(t_batch):
#print("Batch",(n+1),"Running...",end="")
t0 = time.time()
if n>0:
i = np.append(i[0]-0.01,i)
init_state = state_vector
state = nv.odeint_fixed(dAdt, init_state, i, n_n, F_b)
t1 = time.time()
#print("Finished in",np.round(t1-t0,2),"secs...Saving...",end="")
state_vector = state[-1,:]
#np.save("batch"+str(int(sys.argv[1])+1)+"_part_"+str(n+1),state)
state=None
t2 = time.time()
#print("Saved ( Execution Time:",np.round(t2-t0,3),"secs )")
np.save("state_vector",state_vector)
print(int(sys.argv[1])+1,"/",int(sys.argv[4]),"Completed.",np.round(time.time()-t_,3),"secs")
| interactive/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Running attribute inference attacks on the Nursery data
# In this tutorial we will show how to run both black-box and white-box inference attacks. This will be demonstrated on the Nursery dataset (original dataset can be found here: https://archive.ics.uci.edu/ml/datasets/nursery).
# ## Preliminaries
# In order to mount a successful attribute inference attack, the attacked feature must be categorical, and with a relatively small number of possible values (preferably binary, but should at least be less then the number of label classes).
#
# In the case of the nursery dataset, the sensitive feature we want to infer is the 'social' feature. In the original dataset this is a categorical feature with 3 possible values. To make the attack more successful, we reduced this to two possible feature values by assigning the original value 'problematic' the new value 1, and the other original values were assigned the new value 0.
#
# We have also already preprocessed the dataset such that all categorical features are one-hot encoded, and the data was scaled using sklearn's StandardScaler.
# ## Load data
# +
import os
import sys
sys.path.insert(0, os.path.abspath('..'))
from art.utils import load_nursery
(x_train, y_train), (x_test, y_test), _, _ = load_nursery(test_set=0.2, transform_social=True)
# -
# ## Train decision tree model
# +
from sklearn.tree import DecisionTreeClassifier
from art.estimators.classification.scikitlearn import ScikitlearnDecisionTreeClassifier
model = DecisionTreeClassifier()
model.fit(x_train, y_train)
art_classifier = ScikitlearnDecisionTreeClassifier(model)
print('Base model accuracy: ', model.score(x_test, y_test))
# -
# ## Attack
# ### Black-box attack
# The black-box attack basically trains an additional classifier (called the attack model) to predict the attacked feature's value from the remaining n-1 features as well as the original (attacked) model's predictions.
# #### Train attack model
# +
import numpy as np
from art.attacks.inference.attribute_inference import AttributeInferenceBlackBox
attack_train_ratio = 0.5
attack_train_size = int(len(x_train) * attack_train_ratio)
attack_x_train = x_train[:attack_train_size]
attack_y_train = y_train[:attack_train_size]
attack_x_test = x_train[attack_train_size:]
attack_y_test = y_train[attack_train_size:]
attack_feature = 1 # social
# get original model's predictions
attack_x_test_predictions = np.array([np.argmax(arr) for arr in art_classifier.predict(attack_x_test)]).reshape(-1,1)
# only attacked feature
attack_x_test_feature = attack_x_test[:, attack_feature].copy().reshape(-1, 1)
# training data without attacked feature
attack_x_test = np.delete(attack_x_test, attack_feature, 1)
bb_attack = AttributeInferenceBlackBox(art_classifier, attack_feature=attack_feature)
# train attack model
bb_attack.fit(attack_x_train)
# -
# #### Infer sensitive feature and check accuracy
# get inferred values
values = [-0.70718864, 1.41404987]
inferred_train_bb = bb_attack.infer(attack_x_test, pred=attack_x_test_predictions, values=values)
# check accuracy
train_acc = np.sum(inferred_train_bb == np.around(attack_x_test_feature, decimals=8).reshape(1,-1)) / len(inferred_train_bb)
print(train_acc)
# This means that for 59% of the training set, the attacked feature is inferred correctly using this attack.
# ## Whitebox attacks
# These two attacks do not train any additional model, they simply use additional information coded within the attacked decision tree model to compute the probability of each value of the attacked feature and outputs the value with the highest probability.
# ### First attack
# +
from art.attacks.inference.attribute_inference import AttributeInferenceWhiteBoxLifestyleDecisionTree
wb_attack = AttributeInferenceWhiteBoxLifestyleDecisionTree(art_classifier, attack_feature=attack_feature)
priors = [3465 / 5183, 1718 / 5183]
# get inferred values
inferred_train_wb1 = wb_attack.infer(attack_x_test, attack_x_test_predictions, values=values, priors=priors)
# check accuracy
train_acc = np.sum(inferred_train_wb1 == np.around(attack_x_test_feature, decimals=8).reshape(1,-1)) / len(inferred_train_wb1)
print(train_acc)
# -
# ### Second attack
# +
from art.attacks.inference.attribute_inference import AttributeInferenceWhiteBoxDecisionTree
wb2_attack = AttributeInferenceWhiteBoxDecisionTree(art_classifier, attack_feature=attack_feature)
# get inferred values
inferred_train_wb2 = wb2_attack.infer(attack_x_test, attack_x_test_predictions, values=values, priors=priors)
# check accuracy
train_acc = np.sum(inferred_train_wb2 == np.around(attack_x_test_feature, decimals=8).reshape(1,-1)) / len(inferred_train_wb2)
print(train_acc)
# -
# The white-box attacks are able to correctly infer the attacked feature value in 62% and 70% of the training set respectively.
#
# Now let's check the precision and recall:
# +
def calc_precision_recall(predicted, actual, positive_value=1):
score = 0 # both predicted and actual are positive
num_positive_predicted = 0 # predicted positive
num_positive_actual = 0 # actual positive
for i in range(len(predicted)):
if predicted[i] == positive_value:
num_positive_predicted += 1
if actual[i] == positive_value:
num_positive_actual += 1
if predicted[i] == actual[i]:
if predicted[i] == positive_value:
score += 1
if num_positive_predicted == 0:
precision = 1
else:
precision = score / num_positive_predicted # the fraction of predicted “Yes” responses that are correct
if num_positive_actual == 0:
recall = 1
else:
recall = score / num_positive_actual # the fraction of “Yes” responses that are predicted correctly
return precision, recall
# black-box
print(calc_precision_recall(inferred_train_bb, np.around(attack_x_test_feature, decimals=8), positive_value=1.41404987))
# white-box 1
print(calc_precision_recall(inferred_train_wb1, np.around(attack_x_test_feature, decimals=8), positive_value=1.41404987))
# white-box 2
print(calc_precision_recall(inferred_train_wb2, np.around(attack_x_test_feature, decimals=8), positive_value=1.41404987))
# -
# To verify the significance of these results, we now run a baseline attack that uses only the remaining features to try to predict the value of the attacked feature, with no use of the model itself.
# +
from art.attacks.inference.attribute_inference import AttributeInferenceBaseline
baseline_attack = AttributeInferenceBaseline(attack_feature=attack_feature)
# train attack model
baseline_attack.fit(attack_x_train)
# infer values
inferred_train_baseline = baseline_attack.infer(attack_x_test, values=values)
# check accuracy
baseline_train_acc = np.sum(inferred_train_baseline == np.around(attack_x_test_feature, decimals=8).reshape(1,-1)) / len(inferred_train_baseline)
print(baseline_train_acc)
# -
# We can see that both the black-box and white-box attacks do better than the baseline.
# ## Membership based attack
# In this attack the idea is to find the target feature value that maximizes the membership attack confidence, indicating that this is the most probable value for member samples. It can be based on any membership attack (either black-box or white-box) as long as it supports the given model.
#
# ### Train membership attack
# +
from art.attacks.inference.membership_inference import MembershipInferenceBlackBox
mem_attack = MembershipInferenceBlackBox(art_classifier)
mem_attack.fit(x_train[:attack_train_size], y_train[:attack_train_size], x_test, y_test)
# -
# ### Apply attribute attack
# +
from art.attacks.inference.attribute_inference import AttributeInferenceMembership
attack = AttributeInferenceMembership(art_classifier, mem_attack, attack_feature=attack_feature)
# infer values
inferred_train = attack.infer(attack_x_test, attack_y_test, values=values)
# check accuracy
train_acc = np.sum(inferred_train == np.around(attack_x_test_feature, decimals=8).reshape(1,-1)) / len(inferred_train)
print(train_acc)
# -
# We can see that this attack does slightly better than the regular black-box attack, even though it still assumes only black-box access to the model (employs a black-box membership attack). But it is not as good as the white-box attacks.
| notebooks/attack_attribute_inference.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1207 - Rogue Domain Controller
# Adversaries may register a rogue Domain Controller to enable manipulation of Active Directory data. DCShadow may be used to create a rogue Domain Controller (DC). DCShadow is a method of manipulating Active Directory (AD) data, including objects and schemas, by registering (or reusing an inactive registration) and simulating the behavior of a DC. (Citation: DCShadow Blog) Once registered, a rogue DC may be able to inject and replicate changes into AD infrastructure for any domain object, including credentials and keys.
#
# Registering a rogue DC involves creating a new server and nTDSDSA objects in the Configuration partition of the AD schema, which requires Administrator privileges (either Domain or local to the DC) or the KRBTGT hash. (Citation: Adsecurity Mimikatz Guide)
#
# This technique may bypass system logging and security monitors such as security information and event management (SIEM) products (since actions taken on a rogue DC may not be reported to these sensors). (Citation: DCShadow Blog) The technique may also be used to alter and delete replication and other associated metadata to obstruct forensic analysis. Adversaries may also utilize this technique to perform [SID-History Injection](https://attack.mitre.org/techniques/T1178) and/or manipulate AD objects (such as accounts, access control lists, schemas) to establish backdoors for Persistence. (Citation: DCShadow Blog)
# ## Atomic Tests
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
# ### Atomic Test #1 - DCShadow - Mimikatz
# Utilize Mimikatz DCShadow method to simulate behavior of a Domain Controller
#
# [DCShadow](https://www.dcshadow.com/)
# [Additional Reference](http://www.labofapenetrationtester.com/2018/04/dcshadow.html)
#
# **Supported Platforms:** windows
# Run it with these steps!
# 1. Start Mimikatz and use !processtoken (and not token::elevate - as it elevates a thread) to escalate to SYSTEM.
# 2. Start another mimikatz with DA privileges. This is the instance which registers a DC and is used to "push" the attributes.
# 3. lsadump::dcshadow /object:ops-user19$ /attribute:userAccountControl /value:532480
# 4. lsadump::dcshadow /push
#
# ## Detection
# Monitor and analyze network traffic associated with data replication (such as calls to DrsAddEntry, DrsReplicaAdd, and especially GetNCChanges) between DCs as well as to/from non DC hosts. (Citation: GitHub DCSYNCMonitor) (Citation: DCShadow Blog) DC replication will naturally take place every 15 minutes but can be triggered by an attacker or by legitimate urgent changes (ex: passwords). Also consider monitoring and alerting on the replication of AD objects (Audit Detailed Directory Service Replication Events 4928 and 4929). (Citation: DCShadow Blog)
#
# Leverage AD directory synchronization (DirSync) to monitor changes to directory state using AD replication cookies. (Citation: Microsoft DirSync) (Citation: ADDSecurity DCShadow Feb 2018)
#
# Baseline and periodically analyze the Configuration partition of the AD schema and alert on creation of nTDSDSA objects. (Citation: DCShadow Blog)
#
# Investigate usage of Kerberos Service Principal Names (SPNs), especially those associated with services (beginning with “GC/”) by computers not present in the DC organizational unit (OU). The SPN associated with the Directory Replication Service (DRS) Remote Protocol interface (GUID E3514235–4B06–11D1-AB04–00C04FC2DCD2) can be set without logging. (Citation: ADDSecurity DCShadow Feb 2018) A rogue DC must authenticate as a service using these two SPNs for the replication process to successfully complete.
# ## Shield Active Defense
# ### Behavioral Analytics
# Deploy tools that detect unusual system or user behavior.
#
# Instrument a system to collect detailed information about process execution and user activity, develop a sense of normal or expected behaviors, and alert on abnormal or unexpected activity. This can be accomplished either onboard the target system or by shipping data to a centralized analysis and alerting system.
# #### Opportunity
# There is an opportunity to detect the presence of an adversary by identifying and alerting on anomalous behaviors.
# #### Use Case
# A defender can implement behavioral analytics which would indicate activity on or against a domain controller. Activity which is out of sync with scheduled domain tasks, or results in an uptick in traffic with a particular system on the network could indicate malicious activity.
# #### Procedures
# Use behavioral analytics to detect Living Off The Land Binaries (LOLBins) being used to download and execute a file.
# Use behavioral analytics to identify a system running development tools, but is not used by someone who does development.
# Use behavioral analytics to identify abnormal system processes being used to launch a different process.
| playbook/tactics/defense-evasion/T1207.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2. Feature extraction
# ### References:
# 1. https://github.com/ypwhs/dogs_vs_cats
# 2. https://www.kaggle.com/yangpeiwen/keras-inception-xception-0-47
# ### Import pkgs
import cv2
import numpy as np
from tqdm import tqdm
import pandas as pd
import os
import random
import matplotlib.pyplot as plt
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
# +
# from keras.preprocessing import image
# from keras.models import Model
# from keras.layers import Dense, GlobalAveragePooling2D
# from keras import backend as K
# from keras.layers import Input
# from keras.layers.core import Lambda
# from keras.applications.vgg16 import VGG16
# from keras.applications.vgg19 import VGG19
# from keras.applications.resnet50 import ResNet50
# from keras.applications.inception_v3 import InceptionV3
# from keras.applications.xception import Xception
# from keras.applications.inception_resnet_v2 import InceptionResNetV2
from keras.layers import *
from keras.models import *
from keras.applications import *
from keras.optimizers import *
from keras.regularizers import *
from keras.applications.inception_v3 import preprocess_input
# -
# ### Load data
cwd = os.getcwd()
df = pd.read_csv(os.path.join(cwd, 'input', 'labels.csv'))
print('lables amount: %d' %len(df))
df.head()
n = len(df)
breed = set(df['breed'])
n_class = len(breed)
class_to_num = dict(zip(breed, range(n_class)))
num_to_class = dict(zip(range(n_class), breed))
width = 299
X = np.zeros((n, width, width, 3), dtype=np.uint8)
y = np.zeros((n, n_class), dtype=np.uint8)
for i in tqdm(range(n)):
X[i] = cv2.resize(cv2.imread('.\\input\\train\\%s.jpg' % df['id'][i]), (width, width))
y[i][class_to_num[df['breed'][i]]] = 1
# ### Preview images
plt.figure(figsize=(12, 6))
for i in range(8):
random_index = random.randint(0, n-1)
plt.subplot(2, 4, i+1)
plt.imshow(X[random_index][:,:,::-1])
plt.title(num_to_class[y[random_index].argmax()])
plt.show()
# ### Export feature
def get_features(MODEL, data=X):
cnn_model = MODEL(include_top=False, input_shape=(width, width, 3), weights='imagenet')
inputs = Input((width, width, 3))
x = inputs
x = Lambda(preprocess_input, name='preprocessing')(x)
x = cnn_model(x)
x = GlobalAveragePooling2D()(x)
cnn_model = Model(inputs, x)
features = cnn_model.predict(data, batch_size=64, verbose=1)
return features
inception_features = get_features(InceptionV3, X)
xception_features = get_features(Xception, X)
features = np.concatenate([inception_features, xception_features], axis=-1)
# ### Train model
inputs = Input(features.shape[1:])
x = inputs
x = Dropout(0.5)(x)
x = Dense(n_class, activation='softmax')(x)
model = Model(inputs, x)
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
h = model.fit(features, y, batch_size=128, epochs=10, validation_split=0.1, verbose=2)
# ### Visualize model
# +
# SVG(model_to_dot(model, show_shapes=True).create(prog='dot', format='svg'))
# -
# ### Virtualize train curve
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
plt.figure(figsize=(10, 4))
plt.subplot(1, 2, 1)
plt.plot(h.history['loss'])
plt.plot(h.history['val_loss'])
plt.legend(['loss', 'val_loss'])
plt.ylabel('loss')
plt.xlabel('epoch')
plt.subplot(1, 2, 2)
plt.plot(h.history['acc'])
plt.plot(h.history['val_acc'])
plt.legend(['acc', 'val_acc'])
plt.ylabel('acc')
plt.xlabel('epoch')
# -
# ### Load test data
df2 = pd.read_csv('.\\input\\sample_submission.csv')
n_test = len(df2)
X_test = np.zeros((n_test, width, width, 3), dtype=np.uint8)
for i in tqdm(range(n_test)):
X_test[i] = cv2.resize(cv2.imread('.\\input\\test\\%s.jpg' % df2['id'][i]), (width, width))
# ### Export test data feature
inception_features = get_features(InceptionV3, X_test)
xception_features = get_features(Xception, X_test)
features_test = np.concatenate([inception_features, xception_features], axis=-1)
# ### Get test data prediction and output
y_pred = model.predict(features_test, batch_size=128)
for b in breed:
df2[b] = y_pred[:,class_to_num[b]]
df2.to_csv('.\\output\\pred.csv', index=None)
print('Done !')
| dog-breed-identification/2. Feature_extraction_InceptionV3+Xception.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### In this notebook
# We showcase how to use the Analyzer class available in `../code/analyzer.py`
#
# We will see how to:
# - load data
# - compute internet state and state duration
# - get summary dataset
# - plot time without internet service
# - plot number of internet service interuptions
# - print some reporting
# +
import sys
sys.path.append("../code")
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from analyzer import Analyzer
mpl.rcParams.update({"font.size": 18})
# -
PATH = "../inputs/time_recorded.csv.gzip"
# - load data
# - compute internet state and state duration
on_off_data = Analyzer.load_data(fp=PATH)
# on_off_data = on_off_data[on_off_data.index<="2020-08-11"] # Filter on date here
on_off_data = Analyzer.compute_state_and_duration(df=on_off_data)
display(on_off_data)
# - get summary dataset
freq = "12H" # "24H"
summary_data = Analyzer.get_summary_data(df=on_off_data, freq=freq)
display(summary_data)
# ### Plot
# - plot time without internet service
Analyzer.plot_time_without_internet(df=summary_data, show=True, save=False)
# - plot number of internet service interuptions
Analyzer.plot_number_of_interruptions(df=summary_data, show=True, save=False)
# ### Reporting
# Most recent time without internet
print("0: Time when internet turned off\n1: Time when internet turned on")
on_off_data[on_off_data["state_cum_duration"]==2]["on_off_ind"].tail(20)
# Percentage of time down
Analyzer.reporting(on_off_data=on_off_data, summary_data=summary_data)
| notebooks/plots-and-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import os
import glob
from ssd_model import SSD300, SSD512
from ssd_utils import PriorUtil
from utils.model import load_weights
# -
# ### Data
# Pascal VOC
from data_voc import GTUtility
gt_util = GTUtility('data/VOC2007test/')
#print(gt_util)
# MS COCO
from data_coco import GTUtility
gt_util = GTUtility('data/COCO/', validation=True)
#gt_util = gt_util.convert_to_voc()
#print(gt_util)
# ### Model
# SDD300
model = SSD300(num_classes=gt_util.num_classes)
weights_path = './models/ssd300_voc_weights_fixed.hdf5'; confidence_threshold = 0.35
#weights_path = './models/ssd300_coco_weights_fixed.hdf5'; confidence_threshold = 0.25
# SSD512
model = SSD512(num_classes=gt_util.num_classes)
weights_path = './models/ssd512_voc_weights_fixed.hdf5'; confidence_threshold = 0.7
#weights_path = './models/ssd512_coco_weights_fixed.hdf5'; confidence_threshold = 0.7
load_weights(model, weights_path)
prior_util = PriorUtil(model)
# ### Predict
_, inputs, images, data = gt_util.sample_random_batch(batch_size=32, input_size=model.image_size)
# plot ground truth
for i in range(len(images)):
break
plt.figure(figsize=[8]*2)
plt.imshow(images[i])
gt_util.plot_gt(data[i])
plt.show()
# plot prior boxes
for m in prior_util.prior_maps:
break
plt.figure(figsize=[8]*2)
plt.imshow(images[0])
m.plot_locations()
m.plot_boxes([0, 10, 100])
plt.show()
preds = model.predict(inputs, batch_size=1, verbose=1)
# +
checkdir = os.path.dirname(weights_path)
for fl in glob.glob('%s/result_*' % (checkdir,)):
#os.remove(fl)
pass
for i in range(2):
#for i in range(len(preds)):
plt.figure(figsize=[8]*2)
plt.imshow(images[i])
res = prior_util.decode(preds[i], confidence_threshold=confidence_threshold, fast_nms=True)
prior_util.plot_results(res, classes=gt_util.classes, show_labels=True, gt_data=data[i])
#prior_util.plot_results(res, classes=gt_util.classes, show_labels=True, gt_data=None)
plt.axis('off')
#plt.savefig('%s/result_%03d.jpg' % (checkdir, i))
plt.show()
# -
# ### Real world images
# +
inputs = []
images = []
img_paths = glob.glob('./data/images/*.jpg')
for img_path in img_paths:
img = cv2.imread(img_path)
inputs.append(preprocess(img, model.image_size))
h, w = model.image_size
img = cv2.resize(img, (w,h), cv2.INTER_LINEAR).astype('float32')
img = img[:, :, (2,1,0)] # BGR to RGB
img /= 255
images.append(img)
inputs = np.asarray(inputs)
preds = model.predict(inputs, batch_size=1, verbose=1)
# -
for i in range(len(images)):
print(img_paths[i])
plt.figure(figsize=[8]*2, frameon=True)
plt.imshow(images[i])
res = prior_util.decode(preds[i], confidence_threshold=link_threshold)
prior_util.plot_results(res, classes=gt_util.classes)
plt.axis('off')
plt.show()
| SSD_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TashreefMuhammad/TwitterSentimentAnalysis-BidirectionalLSTM/blob/main/TwitterSentiment_Bi-LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qkvfgiyhMJIM"
# # CSE 4238 | Assignment 3
# ---
# ---
# + [markdown] id="oMEvxstq3oy2"
# # Sentiment Analysis for Tweets | Bi-LSTM | TensorFlow
# ---
# ---
# + [markdown] id="VYlHeLVLxYi4"
# ## Assigned Task
# ---
#
# My ID $\Rightarrow 170104014$
#
# Last $3$ digits $\Rightarrow 014 \rightarrow 14$
#
# ---
#
# **Dataset Selection**
#
# Now, $14 \% 3 = 2$ and so [Dataset 3](https://drive.google.com/file/d/1hgfcQHIlfnnDTSUAA_3--m-YA7BelKl3/view) was assigned.
#
# The dataset is quite similar to [Sentimental Analysis for Tweets](https://www.kaggle.com/gargmanas/sentimental-analysis-for-tweets)
#
# ---
#
# **Model**
#
# $(14 + 3) \% 5 = 17 \% 5 = 2$
#
# Hence, the required model is use of *Bidirectional LSTM*
#
# + [markdown] id="SDNE7pjCMPEY"
# ## Prepare Dataset
# ---
# Download the [dataset](https://drive.google.com/file/d/1hgfcQHIlfnnDTSUAA_3--m-YA7BelKl3/view).
# + id="E_W1JqdvLpHI" colab={"base_uri": "https://localhost:8080/"} outputId="1749e189-2e30-425f-d0ac-d52f0b8f8e45"
# The dataset => https://drive.google.com/file/d/1hgfcQHIlfnnDTSUAA_3--m-YA7BelKl3/view
# !gdown --id 1hgfcQHIlfnnDTSUAA_3--m-YA7BelKl3
# + [markdown] id="0UpWYZloNo2w"
# ## Import Required Libraries
# ---
# Importing libraries that will be required for the experiment.
# + id="C7RxCMuHNDlx"
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras import layers
from keras import backend as K
from keras.utils.vis_utils import plot_model
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix
# + [markdown] id="26yprpqqPTuQ"
# ## Read the Original Data
# ---
# Read the original data from the CSV file found
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="Qy1qO3NsOYcp" outputId="0dfba7fa-ebd6-4bc8-cbb4-c4700190c189"
data = pd.read_csv('/content/Dataset 3.csv', engine = 'python')
data
# + [markdown] id="1Prb7NIM00CZ"
# ## Understanding Data
# ---
#
# Randomize the data and see what are the labels along with data in CSV
# + colab={"base_uri": "https://localhost:8080/", "height": 477} id="gAT9svnKrMI9" outputId="7e18f080-2360-48d9-b1b9-0a3294491669"
data = data.sample(frac = 1., random_state = 14).reset_index(drop = True)
print(data['sentiment'].value_counts(0))
data
# + [markdown] id="z2h8tQQq1Igx"
# ## Dividing Dataset
# ---
# Dividing dataset into 6:2:2 ratio as Training:Validation:Testing.
#
# Also, *EPOCH* variable is declared for controlling how many epochs for training.
#
# After that, observe the data in Train, Val and Test splits.
#
# ltimatey, convert the data into NumPy array for later usage.
# + id="vXk1PKKmPuNI"
EPOCH = 10
split_val = int(0.2 * data.shape[0])
dataTest = data.iloc[-split_val :]
dataVal = data.iloc[- 2 * split_val : -split_val]
dataTrain = data.iloc[: - 2 * split_val]
# + colab={"base_uri": "https://localhost:8080/", "height": 477} id="aMhe-A-lr4fH" outputId="b5931554-3dc0-4a49-870c-0d6bce27af88"
print(dataTrain['sentiment'].value_counts())
dataTrain
# + colab={"base_uri": "https://localhost:8080/", "height": 477} id="l2tTMKOsr4X0" outputId="0a421cbb-7b97-4951-8572-3b000d56083d"
print(dataVal['sentiment'].value_counts())
dataVal
# + colab={"base_uri": "https://localhost:8080/", "height": 477} id="GdtDG4BLr4Hc" outputId="5b5362b1-831d-4835-aefc-b1f4c38de5b2"
print(dataTest['sentiment'].value_counts())
dataTest
# + id="q9B1E-0HMG6C"
trainX = np.array(dataTrain.iloc[:, 0])
trainY = np.array(dataTrain.iloc[:, 1])
valX = np.array(dataVal.iloc[:, 0])
valY = np.array(dataVal.iloc[:, 1])
testX = np.array(dataTest.iloc[:, 0])
testY = np.array(dataTest.iloc[:, 1])
# # To check the outcome
# print(trainX)
# print(trainY)
# print(valX)
# print(valY)
# print(testX)
# print(testY)
# print(trainX.shape)
# print(trainY.shape)
# print(valX.shape)
# print(valY.shape)
# print(testX.shape)
# print(testY.shape)
# + [markdown] id="tjQoeNp413B4"
# ## Tokenizing Data
# ---
#
# Processing the data. Use train data to fit on the text. Later use it to construct sequences for all of the train, val, test quality.
# + id="Oqe-uYGSwRwS"
top_k = 50000
tokenizer = tf.keras.preprocessing.text.Tokenizer(num_words = top_k,
oov_token = "<unk>",
filters = '!"#$%&()*+.,-/:;=?@[\]^_`{|}~')
tokenizer.fit_on_texts(trainX)
tokenizer.word_index['<pad>'] = 0
tokenizer.index_word[0] = '<pad>'
train_seqs = tokenizer.texts_to_sequences(trainX)
val_seqs = tokenizer.texts_to_sequences(valX)
test_seqs = tokenizer.texts_to_sequences(testX)
train_seqs = tf.keras.preprocessing.sequence.pad_sequences(train_seqs, padding = 'pre')
val_seqs = tf.keras.preprocessing.sequence.pad_sequences(val_seqs, padding = 'pre')
test_seqs = tf.keras.preprocessing.sequence.pad_sequences(test_seqs, padding = 'pre')
# + id="KO367UZUFaXS"
# # To check the outcome
# print(trainX)
# print(valX)
# print(testX)
# print('=====')
# print(train_seqs)
# print(val_seqs)
# print(test_seqs)
# print('=====')
# print(train_seqs.shape)
# print(val_seqs.shape)
# print(test_seqs.shape)
# + [markdown] id="lQozFOh92OQe"
# ## Develop Model
# ---
# Develop the model and print it.
#
# Train the model and observe the values.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="p1alVk9SFqnX" outputId="e6ff858a-2ff0-4034-82de-fe55196d1067"
model = Sequential()
model.add(layers.Embedding(len(tokenizer.word_index), 128))
model.add(layers.Bidirectional(layers.LSTM(256, return_sequences = True, dropout = 0.2)))
model.add(layers.Bidirectional(layers.LSTM(512, return_sequences = True, dropout = 0.2)))
model.add(layers.Bidirectional(layers.LSTM(64, return_sequences = True, dropout = 0.2)))
model.add(layers.Bidirectional(layers.LSTM(8, dropout = 0.2)))
model.add(layers.Dense(1, activation = 'sigmoid'))
model.compile(loss = 'binary_crossentropy', optimizer = 'adam', metrics = ['accuracy'])
model.summary()
plot_model(model, to_file = 'model_plot.png', show_shapes = True, show_layer_names = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="gDJ3y2IXKMXi" outputId="1482dc1e-a32f-<PASSWORD>-de<PASSWORD>"
history = model.fit(train_seqs, trainY, epochs = EPOCH, validation_data = (val_seqs, valY), verbose = 1)
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, 'accuracy')
plot_graphs(history, 'loss')
print('\nEpoch No. Train Accuracy Train Loss Val Accuracy Val Loss')
for i in range(EPOCH):
print('{:8d} {:10f} \t {:10f} \t {:10f} \t {:10f}'.format(i + 1, history.history['accuracy'][i], history.history['loss'][i], history.history['val_accuracy'][i], history.history['val_loss'][i]))
# + [markdown] id="jSPQirEq2nKb"
# ## Find Performance
# ---
# Use the developed model to see how it performs in al the datasets.
#
# Separate performance was calculated for all the Train, Val and Test data.
# + [markdown] id="TyFEiBlM20nr"
# ### Train Data Performance
# + colab={"base_uri": "https://localhost:8080/", "height": 408} id="IlTlpLrtgfMt" outputId="f463316c-0267-4233-939f-1b507bfa3adf"
y_pred = model.predict(train_seqs)
y_pred = np.where(y_pred > 0.5, 1, 0)
y_pred = np.reshape(y_pred, (y_pred.shape[0]))
loss, accuracy = model.evaluate(train_seqs, trainY, verbose = 1)
print('Train Loss:', loss)
print('Train Accuracy:', accuracy, '(Model Metric)')
print('Train Accuracy:', accuracy_score(trainY, y_pred), '(sklearn Metric)')
print('Train Precision:', precision_score(trainY, y_pred))
print('Train Recall:', recall_score(trainY, y_pred))
print('Train F1-score:', f1_score(trainY, y_pred))
sns.heatmap(confusion_matrix(trainY, y_pred), annot = True, fmt = 'g')
# + [markdown] id="2iySVDHT25d6"
# ### Val Data Performance
# + colab={"base_uri": "https://localhost:8080/", "height": 412} id="_D7KdFvHhHOS" outputId="7f6bd9d4-ed39-4184-9b95-f54e93cd994e"
y_pred = model.predict(val_seqs)
y_pred = np.where(y_pred > 0.5, 1, 0)
y_pred = np.reshape(y_pred, (y_pred.shape[0]))
loss, accuracy = model.evaluate(val_seqs, valY, verbose = 1)
print('Validation Loss:', loss)
print('Validation Accuracy:', accuracy, '(Model Metric)')
print('Validation Accuracy:', accuracy_score(valY, y_pred), '(sklearn Metric)')
print('Validation Precision:', precision_score(valY, y_pred))
print('Validation Recall:', recall_score(valY, y_pred))
print('Validation F1-score:', f1_score(valY, y_pred))
sns.heatmap(confusion_matrix(valY, y_pred), annot = True, fmt = 'g')
# + [markdown] id="DHudDMkw28oS"
# ### Test Data Performance
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="SWt3knXRMnJ6" outputId="60bb24d6-35dc-43db-c3a4-1f3c7a857336"
y_pred = model.predict(test_seqs)
y_pred = np.where(y_pred > 0.5, 1, 0)
y_pred = np.reshape(y_pred, (y_pred.shape[0]))
loss, accuracy = model.evaluate(test_seqs, testY, verbose = 1)
print('Test Loss:', loss)
print('Test Accuracy:', accuracy, '(Model Metric)')
print('Test Accuracy:', accuracy_score(testY, y_pred), '(sklearn Metric)')
print('Test Precision:', precision_score(testY, y_pred))
print('Test Recall:', recall_score(testY, y_pred))
print('Test F1-score:', f1_score(testY, y_pred))
sns.heatmap(confusion_matrix(testY, y_pred), annot = True, fmt = 'g')
# + [markdown] id="GTs_zPu9i8BD"
# $\Large\text{Thus, it concludes the full experiment.}$
| TwitterSentiment_Bi-LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/oughtinc/ergo/blob/master/notebooks/covid-19-active.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="4scekT3jMkof" colab_type="text"
# Project: Estimating current Covid-19 infections (including undiagnosed) from multiple sources
#
# # Background
#
# This notebook allows forecasting different COVID-19 related parameters, by using probability distributions, and integrating spreadsheets and the forecasting platform Foretold.io.
#
# ## Using the Colab
# - Run the colab by going to Runtime > "Run All"
# - You'll need to follow the instructions in the output of the "Authentication" section to allow the colab to read google sheets
# - If you want to use private foretold predictions, set foretold_token to the token of a bot account on foretold with access to the private channel
# - You can override parameters for individual regions with point estimates in the [Parameter Spreadsheet](https://docs.google.com/spreadsheets/d/1n42Op7sE3vUs4v2oSO19OM7yBH-9LIjPOp2dK_DtsMM/edit?usp=sharing)
# - The priority order for parameters is: 1. foretold distributions from the parameters table [https://www.foretold.io/c/1dd5b83a-075c-4c9f-b896-3172ec899f26/n/245566be-1e01-4dbf-9dcd-f0754b44e99e] 2. point estimates from the parameters google sheet 3. foretold distribution for the parameter for the world from the parameters table
# - The colab notebook will output active_infections_prediction.csv (active infections) and countermeasures.csv (countermeasures). If using Google Chrome they should download automatically, otherwise you can open the File pane on the left side of the screen and right click on the file to download
# - If you want to make changes, it's probably best to make your own copy of the colab
#
# ## Notes
# - Currently, doesn't predict for regions with < 1000 cases or < 10 deaths. If you want to remove this restriction, comment out the cell after "Restrict to countries with > 1000 cases and > 10 deaths"
#
# - Does not yet directly support overriding parameters with a different foretold distribution
# - Does not yet directly support overriding history of the number of cases
#
# If you need to override any of these values for a specific region, you can likely load the data and modify the function that retrieves that parameter in the "Data Retrieval Functions" cell to substitute in your data
#
# + [markdown] id="ULCkjkHtgYXf" colab_type="text"
# # Authentication
# + id="rrqdrJPXqWm6" colab_type="code" outputId="2ff05cca-db90-41a6-bd6b-205aee82b9d5" colab={"base_uri": "https://localhost:8080/", "height": 471}
# !pip install --progress-bar off --quiet typing_extensions
# !pip install --progress-bar off --quiet poetry # Fixes https://github.com/python-poetry/poetry/issues/532
# !pip install --progress-bar off --quiet git+https://github.com/oughtinc/ergo.git@<PASSWORD>
# !pip install --progress-bar off --quiet pendulum seaborn
import ergo
# + id="LftbjNvlR4jI" colab_type="code" colab={}
import google
#Authenticate (needs user interaction, so don't hide this cell)
google.colab.auth.authenticate_user()
foretold_token = "<PASSWORD>"
# + [markdown] id="txvCY3ZnWm7I" colab_type="text"
# # Setup
# + [markdown] id="XDha6c0dWpir" colab_type="text"
# ## General
# + id="R58fPeVfv9qi" colab_type="code" outputId="eccadd59-d26c-4929-bce6-7e1f59825fd1" colab={"base_uri": "https://localhost:8080/", "height": 52}
# !pip install -q pendulum pycountry
# + id="7P_f9L_5Wqdk" colab_type="code" colab={}
# %load_ext google.colab.data_table
# + id="69P3Cn9fWsz_" colab_type="code" colab={}
import pandas as pd
import pendulum
import gspread
import numpy as np
import math
import pycountry
import datetime
import tqdm
import time
import seaborn
import torch
from matplotlib import pyplot as plt
from io import StringIO
from types import SimpleNamespace
from oauth2client.client import GoogleCredentials
# + [markdown] id="8ogcaVqm3JZP" colab_type="text"
# ## Ergo
# + [markdown] id="PLiNYwFYYOkd" colab_type="text"
# ## Epimodel
# + id="ngYXYcrXYVsj" colab_type="code" outputId="dd02433e-e57c-4e7d-f924-52bcbfd71136" colab={"base_uri": "https://localhost:8080/", "height": 104}
# !git clone -q https://github.com/epidemics/epimodel
# %cd /content/epimodel
# !git pull
# !git clone -q https://github.com/epidemics/epimodel-covid-data data
# %cd /content/epimodel/data
# !git pull
# %cd /content
import sys
sys.path.append('epimodel')
# + [markdown] id="vARgWcISYWIy" colab_type="text"
# Epimodel needs unidecode:
# + id="yvZkHOIRYYUW" colab_type="code" outputId="a0716a4b-2c49-4051-b775-4d0c41af4908" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !pip install -q unidecode
# + id="ledQS6OPYkOI" colab_type="code" colab={}
import epimodel
regions = epimodel.RegionDataset.load('epimodel/data/regions.csv')
# + id="-uzQd81iRVns" colab_type="code" colab={}
def name_to_code(name):
try:
return regions.find_one_by_name(name).Code
except KeyError:
return ""
# + [markdown] id="jhPWNnpDRLvS" colab_type="text"
# ## Loading Data
# + id="_ye-pp6jRK0e" colab_type="code" colab={}
# + [markdown] id="iacbEdrVs9id" colab_type="text"
# ## Utility Functions
# + id="NXxTccnUtAvE" colab_type="code" colab={}
from uuid import UUID
def is_uuid(s):
try:
uuid_obj = UUID(s, version=4)
return True
except ValueError:
return False
def load_spreadsheet(url, sheet):
gc = gspread.authorize(GoogleCredentials.get_application_default())
wb = gc.open_by_url(url)
sheet = wb.worksheet(sheet)
values = sheet.get_all_values()
return values
# + [markdown] id="QtTCgPGBN5Ep" colab_type="text"
# # Estimating active infections
#
# In order to get an estimate of active infections, four different models are ensembled.
#
# The first three models are statistical and source their parameters from [this Foretold community](https://www.foretold.io/c/1dd5b83a-075c-4c9f-b896-3172ec899f26/notebooks), if available, and otherwise as point estimates from [this spreadsheet](https://docs.google.com/spreadsheets/d/1n42Op7sE3vUs4v2oSO19OM7yBH-9LIjPOp2dK_DtsMM/edit?usp=sharing).
#
# The fourth model is simply a human-estimated distribution of active infections.
#
# The models are described in more detail [here](https://www.notion.so/Ensemble-model-for-estimating-active-infections-00b72ecf6f674602ad27efb15249dd7e).
# + [markdown] id="SEY97QhQRZtd" colab_type="text"
# ## Load Parameters from [Parameter Spreadsheet](https://docs.google.com/spreadsheets/d/1n42Op7sE3vUs4v2oSO19OM7yBH-9LIjPOp2dK_DtsMM/edit?usp=sharing)
# + id="fN2XAt0RaY69" colab_type="code" colab={}
def get_code(row):
if row["Province/State"]:
return name_to_code(row["Province/State"])
return name_to_code(row["Country/Region"])
def load_parameters(parameters_url, parameters_sheet):
values = load_spreadsheet(parameters_url, parameters_sheet)
d = {}
columns = values[0]
for row in values[1:]:
if row[0]:
code = name_to_code(row[0])
else:
code = name_to_code(row[1])
d[code] = {column: value for column, value in zip(columns, row)}
return d
parameters = load_parameters('https://docs.google.com/spreadsheets/d/1n42Op7sE3vUs4v2oSO19OM7yBH-9LIjPOp2dK_DtsMM/edit', 'Parameters')
# + [markdown] id="Sbe-QOuB9MIz" colab_type="text"
# ## Load Foretold Parameters
# + [markdown] id="xV14LaILePYN" colab_type="text"
# ### Foretold Parameter JSON
# + id="ZEruYh2j87xj" colab_type="code" colab={}
# Need to replace with full json created by Ozzie
foretold_lookup = [
{
"name": "@locations/n-hungary",
"locationData": {
"name": "Hungary",
"isoCode": "HU",
"foretoldName": "@locations/n-hungary"
},
"april19": "fceee746-01f9-42b5-9bb4-f5141196c163",
"april26": "126f6846-90b4-4e44-bc71-cf4918367344",
"may3": "1fdc46e2-c641-45f7-99cf-58469f2d5606",
"may17": "033e3efd-17d7-498a-bc63-28de2cdf225b",
"deathsAttributed": "2dee3f58-c6cd-4503-85f6-3e886f72fb50",
"infectionsAttributed": "0d4f247b-2d25-4289-a3ff-411e44067abf",
"timeToDeath": "9109a7d8-9eab-4094-93ea-15c92348033e",
"infectionFatalityRatio": "0d7bea59-100a-4945-9fcd-f5351e89950b"
},
{
"name": "@locations/n-czech-republic",
"locationData": {
"name": "Czech Republic",
"isoCode": "CZ",
"foretoldName": "@locations/n-czech-republic"
},
"april19": "c08c0ea4-09fe-45fe-9caf-9697a67f3262",
"april26": "46fbf258-70a9-4c39-9f3b-5d6f91930632",
"may3": "d892166d-d14c-4edf-8475-4b1b274c3656",
"may17": "fcab14bf-50e0-4c61-a9fa-cdf1835af007",
"deathsAttributed": "901e86f9-4841-4f9f-b555-d61417d1fb7a",
"infectionsAttributed": "2493fa7f-75f3-4396-bef5-5ae6b4bae324",
"timeToDeath": "7a0b3b4b-2a5c-4a0f-b500-c950d65fc028",
"infectionFatalityRatio": "d465c517-dce7-4d92-b173-26e022e74610"
},
{
"name": "@locations/n-ukraine",
"locationData": {
"name": "Ukraine",
"isoCode": "UA",
"foretoldName": "@locations/n-ukraine"
},
"april19": "a176eb57-39d6-4a9c-97be-58ab9eb9f55a",
"april26": "94750cd8-dcaf-4fad-a483-e46cfbba12e3",
"may3": "465af1bd-90f0-4c0e-bb2a-eec1047aa247",
"may17": "bcba6198-093c-4534-b17a-5815386ac64f",
"deathsAttributed": "d32ad94f-e8c4-4337-9a25-31e91bcfa63d",
"infectionsAttributed": "3b21bbf2-0287-4624-b409-4832f8055b0d",
"timeToDeath": "678bbd80-2864-4cd0-a63d-3ad027600e8b",
"infectionFatalityRatio": "34950965-c488-4b11-8956-c10a20e3d0e6"
},
{
"name": "@locations/n-netherlands",
"locationData": {
"name": "Netherlands",
"isoCode": "NL",
"foretoldName": "@locations/n-netherlands"
},
"april19": "562c5477-5037-45e1-b626-2b0dc33d92f9",
"april26": "b334bfcf-a3ef-49fa-9c4c-26aacd8ff337",
"may3": "c2f72783-a3fc-43d8-b92d-0d2abf52217c",
"may17": "87ae4c8d-5fdf-4961-9087-8639afa12124",
"deathsAttributed": "486f78ed-4c13-4adb-b399-95f9556c5fe3",
"infectionsAttributed": "0902bcff-0253-4eec-9864-d5643057fbae",
"timeToDeath": "7e4e9c8e-307f-4216-8463-9bb5f3fd08a4",
"infectionFatalityRatio": "06e42fc2-eb27-4897-818d-5e954964399c"
},
{
"name": "@locations/n-united-kingdon",
"locationData": {
"name": "United Kingdon",
"isoCode": "GB",
"foretoldName": "@locations/n-united-kingdon"
},
"april19": "42476b18-e386-4c35-9f4b-a8f1cda264a9",
"april26": "7615f40d-f086-49b9-b5b5-f33d0e13a705",
"may3": "2a4336ac-a340-477a-9611-9d8578389304",
"may17": "d1778b1b-d956-414d-8aeb-82d6c3c08110",
"deathsAttributed": "315d095d-ae05-4e9d-80e3-1fb27246f7c2",
"infectionsAttributed": "76261599-422b-463c-ad4d-92962bc6c368",
"timeToDeath": "b88df3fd-8af7-46bf-8144-252703281580",
"infectionFatalityRatio": "70ed6dfb-0e84-462e-a06b-a985800be2d3"
},
{
"name": "@locations/n-russia",
"locationData": {
"name": "Russia",
"isoCode": "RU",
"foretoldName": "@locations/n-russia"
},
"april19": "9cf07291-7d43-4899-b391-bbf6977b5fee",
"april26": "7442037d-e749-4a1b-97a9-716f12363c70",
"may3": "3e919b5d-a049-4090-99b8-0f411f7366d5",
"may17": "a436113f-aa22-449b-951e-9f577d26fbb9",
"deathsAttributed": "57bf9b8b-40ea-43d2-9085-70d12500197d",
"infectionsAttributed": "cdc51237-62ca-4157-9d3f-f1c963add9aa",
"timeToDeath": "a4f58a70-ef24-493a-a835-a1051f23f3fd",
"infectionFatalityRatio": "0f5e66eb-e615-4fcc-a0b9-f9182bb2c03b"
},
{
"name": "@locations/n-france",
"locationData": {
"name": "France",
"isoCode": "FR",
"foretoldName": "@locations/n-france"
},
"april19": "70407aee-ffcf-4466-b352-6b05dc424c10",
"april26": "bd443efd-12f5-4db4-83d4-d65567797c91",
"may3": "4bb96a2d-6103-40af-be6f-2e33f48a4298",
"may17": "44cce87c-649a-4414-ab97-ff24a5fb5167",
"deathsAttributed": "bb4c4a26-4882-4a12-a449-6df4fc1d4b75",
"infectionsAttributed": "855bece8-0364-487d-adf2-0e253fb30475",
"timeToDeath": "dc649a48-f8b1-47cd-ba36-6190e8934108",
"infectionFatalityRatio": "5f8c61ac-00b4-4438-bc12-a4c4444a9676"
},
{
"name": "@locations/n-belarus",
"locationData": {
"name": "Belarus",
"isoCode": "BY",
"foretoldName": "@locations/n-belarus"
},
"april19": "98889cef-85b2-4672-80d4-c6eee321e0c4",
"april26": "e778ebe3-b7c4-4b2b-b210-2015967ef772",
"may3": "b568f648-4542-4487-a540-f91cb571fa22",
"may17": "eaa948ec-be88-4efd-a04f-c5cba0894a0a",
"deathsAttributed": "60c88d73-d8fd-42ca-8259-cafa032fb21d",
"infectionsAttributed": "a294bf30-f589-4b2a-a28b-c50942c26163",
"timeToDeath": "1d90b66b-2c43-42a3-8e2d-b221973a90c8",
"infectionFatalityRatio": "1f9e4a9f-5099-4f74-984e-6f184b0467e5"
},
{
"name": "@locations/n-spain",
"locationData": {
"name": "Spain",
"isoCode": "ES",
"foretoldName": "@locations/n-spain"
},
"april19": "1e17b500-92fe-4ef5-a36c-77182fc5f712",
"april26": "5477c30e-8303-4c5c-93f0-27317226e56f",
"may3": "70a5c96f-1571-4b83-90a3-cc775dd72f6f",
"may17": "0dd70995-b4f0-472a-9095-fbc71d5255ff",
"deathsAttributed": "eff08726-220c-4da5-94fc-a3ba26c44421",
"infectionsAttributed": "230ba580-9681-4379-9fe5-3639d73adcce",
"timeToDeath": "8deae807-bdd3-43c0-a8a4-6eb98c358e83",
"infectionFatalityRatio": "3103be98-d7d5-4335-b279-fde5fe5f670b"
},
{
"name": "@locations/n-italy",
"locationData": {
"name": "Italy",
"isoCode": "IT",
"foretoldName": "@locations/n-italy"
},
"april19": "b6e8caf6-afec-428a-adf3-44b962d99383",
"april26": "7422ee75-a130-4b24-b025-d349df24100e",
"may3": "90e76d42-2685-47ff-b8fe-fb80be829333",
"may17": "a1770f21-55a6-415b-8da2-f71411d1f0ae",
"deathsAttributed": "150b74e0-a92b-41d8-8020-0078bdb5c836",
"infectionsAttributed": "4a3dec41-0688-4605-85ac-cae427e9ca51",
"timeToDeath": "af23f463-4231-493a-897e-9d8b903cac90",
"infectionFatalityRatio": "f2456630-cef3-42dd-8fef-d6e6a95e5d2c"
},
{
"name": "@locations/n-nepal",
"locationData": {
"name": "Nepal",
"foretoldName": "@locations/n-nepal"
},
"april19": "c049c851-a806-4eb7-be7c-fbe988e88121",
"april26": "a9447b87-fd25-49b5-81f1-f89880bc196d",
"may3": "3845ef90-ccb1-418f-a6f3-ee5131fdf167",
"may17": "0b9c19cb-945d-4523-95b1-97d5b578318b",
"deathsAttributed": "a9335e1d-2e85-4bfd-b8b0-4d6915604997",
"infectionsAttributed": "c11e38a3-02ef-4998-99ff-500621f60e5c",
"timeToDeath": "b5eb2dc1-e1d3-4e92-97c5-79d45299f827",
"infectionFatalityRatio": "908ceff9-c8d7-4a07-8984-1f43bde37459"
},
{
"name": "@locations/n-india",
"locationData": {
"name": "India",
"isoCode": "IN",
"foretoldName": "@locations/n-india"
},
"april19": "d5115540-209e-4e3f-9a38-76e45b60574a",
"april26": "c5b8f0f2-b4fa-4bf3-a375-6d0e8cd0bbb0",
"may3": "a3808a82-6424-40c7-bf2e-7459540656b9",
"may17": "a011d2a3-3bc2-4045-9b43-a4381325d5f7",
"deathsAttributed": "0b47a481-72d5-486f-84cb-a2f3ab20c6d0",
"infectionsAttributed": "deecf45a-03c5-457e-ba8e-ed51b63eab01",
"timeToDeath": "c1bfe7db-0bfc-4265-ab5e-388ddf349336",
"infectionFatalityRatio": "ad839ed9-d045-417b-8866-290bec82bdd2"
},
{
"name": "@locations/n-myanmar",
"locationData": {
"name": "Myanmar",
"isoCode": "MM",
"foretoldName": "@locations/n-myanmar"
},
"april19": "cd3e2711-1e35-42bc-88a2-39e23b1dc628",
"april26": "4d5f8a9f-7dd6-419e-aebe-3909ae6121f7",
"may3": "9005b8b6-49db-4368-a670-92f50ce434cf",
"may17": "b7eacba3-5801-4989-93aa-46fdba2c3293",
"deathsAttributed": "3860aaa1-146f-46f3-8fe3-c75e5cac8f6d",
"infectionsAttributed": "ba82bd02-f0e7-42e3-9383-f84dfe6027c7",
"timeToDeath": "73531577-c6de-41cc-8f8e-9d26c0b0aa44",
"infectionFatalityRatio": "63659069-747a-4990-b5f0-9b8bc95aaafa"
},
{
"name": "@locations/n-azerbaijan",
"locationData": {
"name": "Azerbaijan",
"isoCode": "AZ",
"foretoldName": "@locations/n-azerbaijan"
},
"april19": "35244016-cabe-4578-a671-bc4f00f709e7",
"april26": "1e5e52c1-8d4b-4fac-b439-7edd1284fd28",
"may3": "8c037be2-960d-4374-9e96-730d279eac07",
"may17": "4c01b1d0-7617-4246-917b-43c7e3a2d47d",
"deathsAttributed": "d62098b8-3b9a-4413-b2b4-2f883f7d3ead",
"infectionsAttributed": "0eae20cb-6ac7-494c-bef7-558cff55ba67",
"timeToDeath": "a84dae59-2c14-4c5d-bd87-29fda38edb7b",
"infectionFatalityRatio": "aec8bcce-d1b4-44bf-9dac-8293d1acb740"
},
{
"name": "@locations/n-kazakhstan",
"locationData": {
"name": "Kazakhstan",
"isoCode": "KZ",
"foretoldName": "@locations/n-kazakhstan"
},
"april19": "2699f03e-3c2c-4c46-8af0-07c18d82c479",
"april26": "97784799-3d90-4ca7-8cad-0fec3b7d9e9d",
"may3": "798b665a-afeb-4b42-9d5a-dd6819f87388",
"may17": "6b375aff-6ef9-48db-bc98-cc1a99fe28b7",
"deathsAttributed": "874e1d19-f952-41a5-9446-ab7dd562dc30",
"infectionsAttributed": "9c0aafe1-86d4-43c5-a987-0a74769778ac",
"timeToDeath": "8bd6d6a8-f689-4e67-90d3-60e09e9de3ad",
"infectionFatalityRatio": "7e0f9430-1b0b-41e1-9102-5bd481436550"
},
{
"name": "@locations/n-uzbekistan",
"locationData": {
"name": "Uzbekistan",
"isoCode": "UZ",
"foretoldName": "@locations/n-uzbekistan"
},
"april19": "2d483a20-b992-4c4b-9c22-3f69764580f7",
"april26": "bc401ff6-f32a-41dd-a4d0-4bbcad3785b6",
"may3": "29bd5c23-1b07-4b98-bb6f-8835ee68e0e5",
"may17": "a1b9b08b-0e67-446c-b9c9-7836752c4c83",
"deathsAttributed": "8014e845-1977-4da6-af79-ed43ac402674",
"infectionsAttributed": "e1db4a03-85c1-4256-bf7a-63dea8001248",
"timeToDeath": "c15df4ff-6a75-4206-8f17-2c1c3a81dd8a",
"infectionFatalityRatio": "91de9fbf-8ac0-4cf7-b123-1db4b634e71f"
},
{
"name": "@locations/n-saudi-arabia",
"locationData": {
"name": "Saudi Arabia",
"isoCode": "SA",
"foretoldName": "@locations/n-saudi-arabia"
},
"april19": "d8255ff3-a729-4e27-bb8f-c41eba8293ac",
"april26": "4aaa2b3d-bed7-4915-9b14-9db3e99e85b5",
"may3": "e5e9ce79-cb2b-4683-8bc2-da1125176303",
"may17": "d6b36a43-07cd-4406-8fee-a0043d11a23b",
"deathsAttributed": "823bd318-bea9-436d-afdb-bdeef07af01c",
"infectionsAttributed": "c729ad9d-12cb-4d09-9989-a500c382085e",
"timeToDeath": "8ac56ba8-598a-4780-9f80-be32d438467d",
"infectionFatalityRatio": "9480970b-ea5d-4528-8dd8-50290125432c"
},
{
"name": "@locations/n-maldives",
"locationData": {
"name": "Maldives",
"isoCode": "MV",
"foretoldName": "@locations/n-maldives"
},
"april19": "9b40c1b5-614b-4bd5-bf1a-5d517cefe9bc",
"april26": "9a8b8053-38c9-42c0-8572-ae9515ed3f78",
"may3": "ba603b63-d0b5-464b-8a2a-c7c50ac5fa9e",
"may17": "af970a0f-79b2-4f79-bd78-2a5b4e5586d0",
"deathsAttributed": "1666a2ac-22c1-4f30-9154-ec1314e726fa",
"infectionsAttributed": "d7c78665-a563-4467-be49-ec9e80538ce3",
"timeToDeath": "3be92417-597a-495b-b883-def91c7f82dd",
"infectionFatalityRatio": "5de1f32e-f3a8-4085-9c94-0b7b2b4d2c8b"
},
{
"name": "@locations/n-iran",
"locationData": {
"name": "Iran",
"isoCode": "IR",
"foretoldName": "@locations/n-iran"
},
"april19": "99d1aa7c-cd70-41c9-a998-fc3dc57b2d79",
"april26": "b41007b0-2256-4f06-b671-f6de0cca61e6",
"may3": "d4f2fa4d-3aa4-4f85-b6c6-78ac15cdc2ff",
"may17": "de892638-2d8d-4364-b6a7-3ca9ce7eb899",
"deathsAttributed": "5957205a-c3d9-4d24-a8ec-bbdd25a8a86b",
"infectionsAttributed": "d9f7b01c-c70d-4035-bb88-88bc1ba7626c",
"timeToDeath": "459ccf8b-1fa9-4b63-aaa2-78a98ba4051b",
"infectionFatalityRatio": "9454b639-0bb4-4e90-b784-3cf2388348a0"
},
{
"name": "@locations/n-pakistan",
"locationData": {
"name": "Pakistan",
"isoCode": "PK",
"foretoldName": "@locations/n-pakistan"
},
"april19": "61743ebe-5a7f-4375-bd4a-0ab8ea4fc2c6",
"april26": "77b73b79-3fe8-412a-802a-298697ce6e66",
"may3": "ba756e89-4cfa-4188-bc6a-f22a943d5cc4",
"may17": "f578bf44-0d11-474a-83db-145082e3de35",
"deathsAttributed": "7ad878e6-5175-44a2-86ee-777c27c55877",
"infectionsAttributed": "762222b6-aedc-4b61-82ba-5340fd984d92",
"timeToDeath": "5e4830c0-69e7-44de-91b1-51c2a5f0a4a0",
"infectionFatalityRatio": "682befc6-3c19-48b0-98a0-bf52c5221c06"
},
{
"name": "@locations/n-pakistan-balochistan",
"locationData": {
"name": "Balochistan (Pakistan)",
"foretoldName": "@locations/n-pakistan-balochistan"
},
"april19": "8683b39d-2e9b-4994-891d-cdec9d9e0fa1",
"april26": "1b21f58e-5de9-42e0-b107-a4ae8b271c92",
"may3": "d81b580a-08c7-4377-8485-1ac355abf58a",
"may17": "be924708-f253-4d64-91f9-6835473d9db3",
"deathsAttributed": "4a5246fd-20af-47b6-9c60-86954aea21b6",
"infectionsAttributed": "a75bb2fc-9333-4b35-b012-4a0c332aa02c",
"timeToDeath": "2c1e7918-a2ec-459f-88be-5ccaa076f7e3",
"infectionFatalityRatio": "28925e55-336a-4556-b642-3ae6ee5ee1b3"
},
{
"name": "@locations/n-mumbai",
"locationData": {
"name": "Mumbai (India)",
"foretoldName": "@locations/n-mumbai"
},
"april19": "e5567c34-8384-4670-9497-f987b65ef693",
"april26": "985f7858-4886-497f-9fb1-f82b051239b7",
"may3": "247de3c8-38d5-459d-be3a-73c2d2c258f6",
"may17": "0a9b2314-c44a-4b74-8999-83f8df5ce322",
"deathsAttributed": "d070f4ac-6834-4d65-a8b8-af0b10ebefc5",
"infectionsAttributed": "56eb2739-4c26-4700-a9ed-330d56e9d054",
"timeToDeath": "cd1f9036-3360-4838-bead-a2d72022451d",
"infectionFatalityRatio": "51f88e70-b1b2-430c-aff7-47cf5ba70987"
},
{
"name": "@locations/n-bangularu",
"locationData": {
"name": "Bangaluru (India)",
"foretoldName": "@locations/n-bangularu"
},
"april19": "d346fd04-9f23-4261-a47e-e8c088500a7b",
"april26": "056db717-02fd-4c42-91da-447469d50a61",
"may3": "dc049c11-d016-4ce4-93d5-700f4bc70937",
"may17": "0bb99ce6-1e3c-4b8b-bcbd-890ca382676b",
"deathsAttributed": "9969f2b2-6471-4599-9a6e-63107bbe153b",
"infectionsAttributed": "bad11dd7-0af6-46c4-9282-a24500fc5687",
"timeToDeath": "3bbe376f-2710-4d7c-95a2-e514af70b96f",
"infectionFatalityRatio": "121bd923-d349-4c99-8d82-8027d49bd969"
},
{
"name": "@locations/n-chennai",
"locationData": {
"name": "Chennai (India)",
"foretoldName": "@locations/n-chennai"
},
"april19": "84200f66-f029-49a5-bebe-1d197f7543d8",
"april26": "5ea46403-0f15-466e-b6fe-3e69b4fabb1a",
"may3": "31ff9cfd-5f87-4981-ae48-169b83cbb099",
"may17": "17fba54d-fb08-4fd0-bdd6-5219555f329c",
"deathsAttributed": "3251bf68-cc41-4a1f-8a82-ef63fefe2165",
"infectionsAttributed": "34eaaf36-1749-4339-9434-64b1036ac9f8",
"timeToDeath": "2d983268-6a7a-44a0-a7a4-071df77a2201",
"infectionFatalityRatio": "412482d9-8317-496e-8236-7288486b5e82"
},
{
"name": "@locations/n-nigeria",
"locationData": {
"name": "Nigeria",
"isoCode": "NG",
"foretoldName": "@locations/n-nigeria"
},
"april19": "8ba68b08-8d4b-4427-ab37-c6753b49b4e6",
"april26": "753cf457-693d-4c4a-81d9-f4afea52007e",
"may3": "670aae22-358c-4a93-8fde-78478360bb32",
"may17": "9f49e806-1be7-472f-bb68-256d7716dbad",
"deathsAttributed": "0babfbc1-2443-471d-a91e-ee56d380e25a",
"infectionsAttributed": "b2bea67c-0c4f-4330-837c-0aea84970d75",
"timeToDeath": "336d982d-9d18-49c7-8f5f-49b14263db32",
"infectionFatalityRatio": "17288b4b-e416-4c37-8018-e5d623343484"
},
{
"name": "@locations/n-ethiopia",
"locationData": {
"name": "Ethiopia",
"isoCode": "ET",
"foretoldName": "@locations/n-ethiopia"
},
"april19": "a3df96e4-b5cd-4aff-bb01-968729d528c6",
"april26": "a26733b9-7404-46a6-ab74-9b84f79305f0",
"may3": "b89d66f4-80db-4bc2-ba45-5961a3d1db07",
"may17": "fd379538-915c-4e06-9256-b5086d862c35",
"deathsAttributed": "2edcdc5b-d8b3-4204-bf9d-67dc34a5b4c6",
"infectionsAttributed": "7313162c-e096-479a-a1e8-9467db9c1663",
"timeToDeath": "102796b0-5466-4d1c-b497-0928b6326443",
"infectionFatalityRatio": "5002ea92-08c3-4359-8871-8735d6898cc5"
},
{
"name": "@locations/n-central-african-republic",
"locationData": {
"name": "Central African Republic",
"isoCode": "CF",
"foretoldName": "@locations/n-central-african-republic"
},
"april19": "45932c74-7f88-449c-bcca-ef15e2fe4156",
"april26": "1361ea67-22ee-491e-8fd1-8c771740ba63",
"may3": "38395987-83ac-4758-9404-15e128b80cc9",
"may17": "ab0332dc-6624-4548-b6ed-7d8002efafa8",
"deathsAttributed": "8d40e5f0-d2be-4143-a0ca-d3239ff64e1c",
"infectionsAttributed": "230e3edd-fb04-4b60-b67f-ca56e6d53ec0",
"timeToDeath": "2187857a-02a4-4992-bbb6-e8b60bcdaaf9",
"infectionFatalityRatio": "bd1a383e-aeae-4aa8-a50b-9cce1eec26d4"
},
{
"name": "@locations/n-south-africa",
"locationData": {
"name": "South Africa",
"isoCode": "ZA",
"foretoldName": "@locations/n-south-africa"
},
"april19": "06b3f1e7-75ac-4521-9869-daf23d78b9cb",
"april26": "326e6249-c8f0-42ba-9344-0b523dd55369",
"may3": "8e23b00d-13c8-477f-9fec-60a9709fa8c8",
"may17": "c335eb3e-ce57-41bb-b42b-a6ff47d59f5f",
"deathsAttributed": "141b8696-1e10-4ff2-b4af-67c78ba03216",
"infectionsAttributed": "abe98728-bf21-4a2f-a95b-28f6ccd58926",
"timeToDeath": "c5ec06c9-593a-4fe1-9dab-d292e31d3c38",
"infectionFatalityRatio": "eee32571-b22b-46af-bb1f-1f6b468b9e10"
},
{
"name": "@locations/n-ghana",
"locationData": {
"name": "Ghana",
"isoCode": "GH",
"foretoldName": "@locations/n-ghana"
},
"april19": "2363e08b-a632-457b-a190-f0a8f3e7c3a2",
"april26": "00a47809-a88a-47d4-86e5-99096fc98030",
"may3": "323f0a91-de83-4353-92ae-00471eb096d1",
"may17": "c4537d29-212e-464f-8341-094a9c32d425",
"deathsAttributed": "f699c231-c333-44dc-b7f8-1b04c5a11454",
"infectionsAttributed": "f40c7f8f-ed00-4bb0-99bf-e2e43b46ed72",
"timeToDeath": "3af6bbe7-ff5a-4bcb-bc77-5f43104d7883",
"infectionFatalityRatio": "994f13ba-a70e-4364-92ee-ca60289215e1"
},
{
"name": "@locations/n-botswana",
"locationData": {
"name": "Botswana",
"isoCode": "BW",
"foretoldName": "@locations/n-botswana"
},
"april19": "6f19c848-98d0-4795-8e56-d704968ac782",
"april26": "68b939f6-0615-4b83-8612-1803f41053c3",
"may3": "e5d74463-b35b-4973-a3d4-efd4a34c148c",
"may17": "edf8458a-9d6d-4482-b791-e8919267006c",
"deathsAttributed": "02031e71-71a9-4ddd-af77-9564617e6885",
"infectionsAttributed": "4d8f6285-d776-456a-a9fa-651c9b1737d1",
"timeToDeath": "9bda5fd9-8a24-4a30-a44d-16cd112f620c",
"infectionFatalityRatio": "ad9a6300-72fd-456c-966f-c815ec3bdb39"
},
{
"name": "@locations/n-egypt",
"locationData": {
"name": "Egypt",
"isoCode": "EG",
"foretoldName": "@locations/n-egypt"
},
"april19": "0d16ad1e-1471-4f79-81b5-744801421678",
"april26": "f5cd38e9-86e6-4bd3-9d5f-1c3ffc48bf91",
"may3": "e49c026f-5c4b-433c-ad7b-1d8a73123b99",
"may17": "9342ddd7-b003-496c-b0a6-cea0a14d1ed2",
"deathsAttributed": "03997a16-1bc9-474a-81f6-de48961c706c",
"infectionsAttributed": "03c6ade8-cf39-4225-b0e0-fb7cf62390df",
"timeToDeath": "ca33ba7f-6482-4e3a-8469-7a21c6ff13ad",
"infectionFatalityRatio": "9ed98b6d-ca46-41e6-8636-6678fd941b2d"
},
{
"name": "@locations/n-africa",
"locationData": {
"name": "Africa",
"isoCode": "W-AF",
"foretoldName": "@locations/n-africa"
},
"april19": "d58bb5e6-cb38-4f24-a82d-59831d7d3aab",
"april26": "39ecd14f-e434-4397-a2f8-84c34fd24149",
"may3": "e3443eb1-a15b-453a-b3e8-59e076937359",
"may17": "e1a5c8cb-9616-4a70-83a1-7b910c4c7c2c",
"deathsAttributed": "d10b810a-c8c9-4747-917d-fc483b69f11a",
"infectionsAttributed": "c9bf1f21-f6dd-41b3-a8a0-bc25c2f5f483",
"timeToDeath": "70e2665e-0290-43c6-814c-9e1e4f942b9b",
"infectionFatalityRatio": "d9a5f7a9-12d8-4a67-8ff2-6083daefb069"
},
{
"name": "@locations/n-tanzania",
"locationData": {
"name": "Tanzania",
"isoCode": "TZ",
"foretoldName": "@locations/n-tanzania"
},
"april19": "1f805578-533d-4cb9-8036-4dd832ca4144",
"april26": "c18a2ae0-e0fb-437d-a41c-6d2eda05679c",
"may3": "39145127-488d-4ef3-9771-aa45c3e907cd",
"may17": "de1bf5fe-d6fe-48b6-8649-491f7605eccd",
"deathsAttributed": "a508bd09-630d-49e1-a0ec-5c3e80df24ee",
"infectionsAttributed": "d603f1bf-701a-4166-a62e-1f0383ff8a7e",
"timeToDeath": "f7ed278a-f2ea-4452-b8cc-8831efc14933",
"infectionFatalityRatio": "34692218-a2f7-478c-9859-0467c50dabd8"
},
{
"name": "@locations/n-dar-es-salaam",
"locationData": {
"name": "<NAME> (Tanzania)",
"foretoldName": "@locations/n-dar-es-salaam"
},
"april19": "f7f872e0-935a-4ce6-ba6c-31b8aff8ab61",
"april26": "ebe7d72a-148e-42c7-9014-3f7e3bbbb31f",
"may3": "339af780-79bb-4265-bfa0-339ef53938c3",
"may17": "2cd62057-5938-4b01-ae0a-4b674dbb44cb",
"deathsAttributed": "ad61b46c-7cf8-4ea5-b4a4-9e1cb24c0d40",
"infectionsAttributed": "76c68e6a-ffbf-42ec-89d5-f66bf4d88a41",
"timeToDeath": "2fb00922-2401-4ccb-8e51-90b612f6a954",
"infectionFatalityRatio": "ea12b45e-7622-4c35-8c2a-613f1383b7d5"
},
{
"name": "@locations/n-bangui",
"locationData": {
"name": "Bangui (Central African Republic)",
"foretoldName": "@locations/n-bangui"
},
"april19": "674fe066-7904-4979-83aa-52ad4ef26598",
"april26": "ef621706-aa58-45b7-9cac-3aa47100ebcd",
"may3": "cd842f3d-8147-491a-ab4a-5f74a6bf91fb",
"may17": "d46eb996-5629-4ef7-b255-d1f04d24a92b",
"deathsAttributed": "67117b33-8678-4cbe-9ec5-3e5eb88dba9b",
"infectionsAttributed": "28fda8e1-e198-4f87-998e-9f1a01a96541",
"timeToDeath": "36b53810-c094-4697-9d4a-0410f4c92d3b",
"infectionFatalityRatio": "35eac892-190f-4dcd-9fcc-2da1cf7fd21c"
},
{
"name": "@locations/n-argentina",
"locationData": {
"name": "Argentina",
"isoCode": "AR",
"foretoldName": "@locations/n-argentina"
},
"april19": "372f63db-8ac8-432c-951a-9ab03499942f",
"april26": "6bebe133-f437-45a8-b107-b6d5356a9e26",
"may3": "30e6abeb-46d7-49c9-bab3-b5aa259c3755",
"may17": "14732518-00f5-4a4e-a056-29cadd2143d6",
"deathsAttributed": "ea2c876a-6939-47f8-884d-eb6b737df0ff",
"infectionsAttributed": "8882e3f5-62a7-45fc-929c-1d529e2b4cf5",
"timeToDeath": "3374dfa2-1354-40a5-85fd-2763de9fd7af",
"infectionFatalityRatio": "d1f8a394-15d3-4dce-a19e-362e5b6e4dbe"
},
{
"name": "@locations/n-canada",
"locationData": {
"name": "Canada",
"isoCode": "CA",
"foretoldName": "@locations/n-canada"
},
"april19": "6be2450d-2f7a-4658-8d35-4a9d64096cf3",
"april26": "0abd65b7-0d76-4706-87c5-890b9887e9df",
"may3": "ccb7d692-6e35-48a6-b62a-dd263659779d",
"may17": "7cc94a5e-1a94-4c80-a08d-cc4aeb39e9bf",
"deathsAttributed": "bb17604e-7b49-4a4f-ba48-a203ad7bdf13",
"infectionsAttributed": "214e34c0-4d9d-4623-9c0c-8a74a5601ab6",
"timeToDeath": "b098b562-f43f-4f08-b5ed-51eb0fcfbdfb",
"infectionFatalityRatio": "5fe6afd4-4bb1-421e-b5fb-5b31e59fd8ec"
},
{
"name": "@locations/n-brazil",
"locationData": {
"name": "Brazil",
"isoCode": "BR",
"foretoldName": "@locations/n-brazil"
},
"april19": "75c2446f-1c46-4243-acf2-d22cafc68edb",
"april26": "0d1d9368-2e1d-4ce9-81fa-14cf397fe8a4",
"may3": "5aa4a5aa-bcbb-4eef-91a1-b15b9221e804",
"may17": "e9cbf9db-0474-440e-bdea-51c94fedbfc2",
"deathsAttributed": "a1eb540f-47cf-4fb5-98c9-9942753f938a",
"infectionsAttributed": "fb4d7053-8128-43c9-b0f0-4a9d3789d3cd",
"timeToDeath": "0401a033-ca6e-40e1-a3de-709f677fc980",
"infectionFatalityRatio": "b41b0c74-6db1-43cc-8430-1fd13c319263"
},
{
"name": "@locations/n-dominica",
"locationData": {
"name": "Dominica (Caribbean)",
"isoCode": "DM",
"foretoldName": "@locations/n-dominica"
},
"april19": "75b5b52b-75da-4162-bdd5-a50f4cb12e81",
"april26": "9e67b467-361d-4661-a2de-c59cc8e101a1",
"may3": "021ccbeb-1ed4-4f09-acc7-3c9fda9fc295",
"may17": "e3db1040-9662-4b44-af01-542c3b1db4dd",
"deathsAttributed": "9270f742-0977-42ea-9b7e-5f12b83bc21b",
"infectionsAttributed": "41f67dd5-b58b-44a6-a312-1936bdc30ed5",
"timeToDeath": "3c5ca22e-26d6-490f-8279-9bbf0d1876a6",
"infectionFatalityRatio": "d0cd2283-ac11-49d1-8cfb-ff5b470be9b7"
},
{
"name": "@locations/n-united-states-of-america",
"locationData": {
"name": "United States of America",
"isoCode": "US ",
"foretoldName": "@locations/n-united-states-of-america"
},
"april19": "06deab80-6610-4bff-a75d-30a6028790dd",
"april26": "38471260-a79a-4807-942d-0784aa19c2c2",
"may3": "92c2f2f3-91aa-48b8-aaa3-c9bb402c0b08",
"may17": "96040d3b-afcb-425a-a1ab-8fed5f5ea861",
"deathsAttributed": "288302d4-28d4-4678-aac7-43e9d41b3ec5",
"infectionsAttributed": "2a85dc03-5983-46e1-8178-46e9c1e4e812",
"timeToDeath": "6e812b42-4dd2-43f3-8f4c-a51ba5c53bae",
"infectionFatalityRatio": "39e4af70-3df0-4a23-819e-e5c427ff02d3"
},
{
"name": "@locations/n-panama",
"locationData": {
"name": "Panama",
"isoCode": "PA",
"foretoldName": "@locations/n-panama"
},
"april19": "e1f5a738-16de-4579-8e93-3d97a952b168",
"april26": "460676be-f8b8-4c3e-bda7-344b9bc50a4e",
"may3": "7297b5d5-385b-4257-ac3e-857fea582cb9",
"may17": "546aa698-b11d-4192-8403-6981c48c3084",
"deathsAttributed": "f4d2cd4a-2530-4ba5-a6a5-f05125851657",
"infectionsAttributed": "3e16f071-b9b5-43a4-a67d-9fe0fa491880",
"timeToDeath": "b1d282bc-d827-4b82-a49d-3aa9e0bb2d77",
"infectionFatalityRatio": "8ae2f9b6-3260-4067-9c67-078061a4d04b"
},
{
"name": "@locations/n-sao-paulo",
"locationData": {
"name": "São Paulo (Brazil)",
"foretoldName": "@locations/n-sao-paulo"
},
"april19": "21d1c054-f46a-4051-9150-d039872ac089",
"april26": "2d00c7ed-9fc3-4766-bd28-cafe950b02f4",
"may3": "c3b31fc8-043c-477f-b520-49d9f306ac8f",
"may17": "979b748f-c3a4-4f01-958c-98e1ce94dff5",
"deathsAttributed": "3fb64298-51df-4e15-ab83-10e18dcc4d7a",
"infectionsAttributed": "e61704de-064a-446b-8356-f6eec1b79326",
"timeToDeath": "7c067ce3-1677-492b-ba9f-f29af81c7f4e",
"infectionFatalityRatio": "95b74135-cf4d-4445-8581-76ee56c8e476"
},
{
"name": "@locations/n-rio-de-janeiro",
"locationData": {
"name": "Rio de Janeiro (Brazil)",
"foretoldName": "@locations/n-rio-de-janeiro"
},
"april19": "ef29c88e-5fdb-4e23-bc10-0e62d636dab8",
"april26": "5e299e50-4d29-44c9-ba46-bc0e71145c8a",
"may3": "0da9c2e3-866b-4655-9a02-6d1f99ed4210",
"may17": "2f414197-09f1-47ec-8719-5df3e0cb66d5",
"deathsAttributed": "96f105d7-c178-48fc-8a11-297d620df1bf",
"infectionsAttributed": "8c3e3c08-72f8-4367-8b80-e555e7cb775a",
"timeToDeath": "44a86f6b-bd9d-4ad6-8c55-4754813d6231",
"infectionFatalityRatio": "41f351c0-7413-411e-bdce-d94bb44fb791"
},
{
"name": "@locations/n-belo-horizonte",
"locationData": {
"name": "<NAME> (Brazil)",
"foretoldName": "@locations/n-belo-horizonte"
},
"april19": "6883c7dc-0390-46a0-af99-9707aafee1c2",
"april26": "7778285b-84fa-4bf3-b560-44b6735c58fa",
"may3": "3098e88c-ee72-47a0-9cef-b398cd0e9243",
"may17": "79a13cfa-f751-4df2-b8d7-a0c5e1a36382",
"deathsAttributed": "a362f036-6535-4f7d-89e0-e9d7aeaf2d90",
"infectionsAttributed": "3968fdc8-ecae-49de-b93c-56c346d10db4",
"timeToDeath": "fb6224a6-9ccf-4e9e-92e3-843d9d4af4eb",
"infectionFatalityRatio": "cbadb95e-dd71-41c0-9061-a479334c2977"
},
{
"name": "@locations/n-papua-new-guinea",
"locationData": {
"name": "Papua New Guinea",
"isoCode": "PG",
"foretoldName": "@locations/n-papua-new-guinea"
},
"april19": "dde72ef6-4219-47a1-8714-e676af9bd8d1",
"april26": "1c9f44cd-ddb9-4400-8f5d-73081446739d",
"may3": "03838e80-64ef-493e-abcb-962d2527d17e",
"may17": "fc216a87-a28c-43ef-ad3e-9c6f705724ad",
"deathsAttributed": "a79095f8-9178-4ce6-bcdf-880057c728dd",
"infectionsAttributed": "288bca95-b216-4931-adf1-e6646293d103",
"timeToDeath": "c8f599d7-c681-43a4-82e8-742b46412d33",
"infectionFatalityRatio": "c85001d2-bd6f-4ecc-94ca-959077e7d92b"
},
{
"name": "@locations/n-earth",
"locationData": {
"name": "Earth",
"isoCode": "W",
"foretoldName": "@locations/n-earth"
},
"may3": "e04ca67a-5b2c-4093-89f9-23921a70056e",
"deathsAttributed": "7a448d93-b3ea-4708-9a89-5e821c9d0aad",
"infectionsAttributed": "f39561c8-5d2b-448b-bef5-4557e64cca86",
"timeToDeath": "03fd9e12-01ba-46bf-8a6e-0230a7eab40a",
"infectionFatalityRatio": "fdc1580f-9cc8-4f2c-b3e7-9df81651e196"
}
]
# Dummy data
# foretold_lookup = [
# {
# "name": "@locations/n-earth",
# "locationData": {
# "name": "Earth",
# "isoCode": "W",
# "foretoldName": "@locations/n-earth"
# },
# "may3": "e04ca67a-5b2c-4093-89f9-23921a70056e",
# "deathsAttributed": "77936da2-a581-48c7-add1-8a4ebc647c8c",
# "infectionsAttributed": "5f9eaaae-4d88-4fa4-9b8e-8bdc97613dc2",
# "timeToDeath": "399272c4-7b58-4a96-a68d-5a779ad4ffb3",
# "infectionFatalityRatio": "10ab95e3-f169-4caf-8865-6c8c3987851d"
# },
# {
# "name": "@locations/n-united-states-of-america",
# "locationData": {
# "name": "United States of America",
# "isoCode": "US ",
# "foretoldName": "@locations/n-united-states-of-america"
# },
# "april19": "06deab80-6610-4bff-a75d-30a6028790dd",
# "april26": "38471260-a79a-4807-942d-0784aa19c2c2",
# "may3": "92c2f2f3-91aa-48b8-aaa3-c9bb402c0b08",
# "may17": "96040d3b-afcb-425a-a1ab-8fed5f5ea861",
# # "deathsAttributed": "e3d867f7-48b6-46b9-b4ff-30d03ce358b0",
# # "infectionsAttributed": "e3d867f7-48b6-46b9-b4ff-30d03ce358b0",
# # "timeToDeath": "4154fe1f-df9d-42d6-9f12-e03c21105611",
# # "infectionFatalityRatio": "e3d867f7-48b6-46b9-b4ff-30d03ce358b0"
# },
# ]
# + [markdown] id="JybHyEE_eTeB" colab_type="text"
# ### Load Foretold Parameters
# + id="lr_XvzFqMPZ2" colab_type="code" outputId="7bf65765-856e-44f0-d6c6-1e501ed71999" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["f12fd2045ab44451ac88f5cca5fc9bcf", "31b76edfad774a1da7b6b3dfbee33abf", "1c39097d85ce49f4bfd759c87df2c2d5", "b4c2118ec80a47f4be35b1d16a7f1b7a", "<KEY>", "<KEY>", "85d5af35bed74435ac2a6e84f2f967bd", "a07bb6462edc47faaf31154829f60ac2"]}
foretold = ergo.Foretold(foretold_token)
foretold_distributions = {}
for entry in tqdm.notebook.tqdm(foretold_lookup):
isoCode = entry["locationData"].get("isoCode", None)
if isoCode is not None:
isoCode = isoCode.strip()
d = {}
for key, value in entry.items():
if key not in ["name", "locationData"]:
try:
d[key] = foretold.get_question(value)
except TypeError:
pass
foretold_distributions[isoCode] = d
# + [markdown] id="qVRqlh1CTP5T" colab_type="text"
# ## Cases Data
# + id="nn7r1FjowsjR" colab_type="code" outputId="e2992e72-98e0-4068-aa21-7045b6816c2c" colab={"base_uri": "https://localhost:8080/", "height": 34}
rds = epimodel.RegionDataset.load('epimodel/data/regions.csv')
cases_raw = epimodel.read_csv('epimodel/data/johns-hopkins.csv', rds)
# List of region codes to run predictions for
codes_to_predict = [x for x in cases_raw.index.get_level_values(0).unique().values if isinstance(x, str)]
last_available_date = cases_raw.index.get_level_values(1).unique().values[-1]
print("Most recent available data is from " + str(last_available_date)[:10])
# + id="oai_pNSnxMFn" colab_type="code" colab={}
predict_date = last_available_date
# Can set if you want to predict infections at a past date
cutoff_date = None # datetime.date(2020,4,2)
if cutoff_date is not None:
cases_raw = cases_raw[[(ix[1]<= cutoff_date) for ix in cases_raw.index]]
# + [markdown] id="R96z_VxCLVy4" colab_type="text"
# ## Restrict to countries with > 1000 cases and > 10 deaths
# + id="zWSCXdsPzCqs" colab_type="code" colab={}
cases_with_enough_data = (cases_raw
.groupby("Code").filter(lambda x: x["Confirmed"].max() > 1000)
.groupby("Code").filter(lambda x: x["Deaths"].max() > 10))
codes_to_predict = [x for x in cases_with_enough_data.index.get_level_values(0).unique().values if isinstance(x, str)]
# + [markdown] id="vnKYrrtp3-97" colab_type="text"
# ## Data Retrieval Functions
# + id="zMsKN3_22kES" colab_type="code" colab={}
# Define functions to lookup data and parameters
# If you need to override parametes, you can add if statements to override the
# values for a particular region
# Pandas lookup is pretty slow, so we just convert to dictionaries + numpy array
cases = {}
for code in codes_to_predict:
cases[code] = {
'Deaths': cases_raw.loc[code]['Deaths'].values,
'Recovered': cases_raw.loc[code]['Recovered'].values,
'Confirmed': cases_raw.loc[code]['Confirmed'].values,
}
def latest_deaths(code):
return cases[code]['Deaths'][-1]
def latest_recovered(code):
return cases[code]['Recovered'][-1]
def confirmed_n_days_ago(code, n):
return cases[code]['Confirmed'][-1 -n]
def get_parameter(code, name):
try:
return float(parameters[code]["DeathMultiplier"])
except (KeyError, ValueError):
return np.NaN
def death_multiplier(code):
"""How many more deaths have occurred than were reported"""
if code in foretold_distributions:
if "deathsAttributed" in foretold_distributions[code]:
return 1 / foretold_distributions[code]["deathsAttributed"].sample_community()
v = get_parameter(code,"DeathMultiplier")
if not np.isnan(v):
return v
return 1 / foretold_distributions["W"]["deathsAttributed"].sample_community()
def ascertainment_parameter(code):
"""How many more infections have occurred than were reported"""
if code in foretold_distributions:
if "infectionsAttributed" in foretold_distributions[code]:
return 1 / foretold_distributions[code]["infectionsAttributed"].sample_community()
v = get_parameter(code,"AscertainmentParameter")
if not np.isnan(v):
return v
return 1 / foretold_distributions["W"]["infectionsAttributed"].sample_community()
def days_to_death(code):
"""How many days between someone being infected and dying"""
if code in foretold_distributions:
if "timeToDeath" in foretold_distributions[code]:
return foretold_distributions[code]["timeToDeath"].sample_community()
v = get_parameter(code,"timeToDeath")
if not np.isnan(v):
return v
return foretold_distributions["W"]["timeToDeath"].sample_community()
def date_offset(d1, d2):
"""What is the difference in days between these two dates"""
return (d1- d2)/np.timedelta64(1, 'D')
def sample_foretold(code, predict_date, max_extrapolation_days=3):
"""
Sample predictions from foretold for predict_date.
If we're in the range of dates with available predictions, interpolate in log
space between the nearest predictions.
If we're outside but within max_extrapolation_days of the first/last
prediction, return the first/last prediction (without extrapolating yet).
If we're outside the range, return np.NaN.
If one of the predictions needed for interpolation is missing, return
np.NaN
"""
assert isinstance(predict_date, np.datetime64)
if code not in foretold_distributions:
return np.NaN
dists = foretold_distributions[code]
prediction_dates = np.array(['2020-04-19','2020-04-26','2020-05-03','2020-05-17']).astype(np.datetime64)
prediction_names = ["april19", "april26", "may3", "may17"]
x = date_offset(predict_date, prediction_dates[0])
# Sample the same quantile for each distribution
q = ergo.uniform()
xs = [date_offset(d, prediction_dates[0]) for d in prediction_dates]
# If we're too far outside of the range where we have predictions, return NaN
if x < -max_extrapolation_days or x > xs[-1] + max_extrapolation_days:
return np.NaN
ys = np.array([
dists[n].quantile(q) if n in dists else np.NaN
for n in prediction_names
])
# Interpolate in log space
return np.exp(np.interp(x, xs, np.log(ys)))
# + [markdown] id="YxqosvYtfCgf" colab_type="text"
# ## Model
#
#
# + id="5pom5CFb29ri" colab_type="code" outputId="f586b48b-6055-4953-d227-fcc73f246eb3" colab={"base_uri": "https://localhost:8080/", "height": 34}
def model(codes_to_predict):
growth_window_days = 17 # When estimating growth rate, look at this length of past growth
canonical_mortality_rate = foretold_distributions["W"]["infectionFatalityRatio"].sample_community()
max_mortality_rate = 0.05
max_growth_multiplier = 64
recovery_time = 17 # TODO: improve estimate
def mortality_rate(code):
"""Most recent fraction of deaths out of resolved cases (death and confirmed)"""
if code in foretold_distributions:
if "infectionFatalityRatio" in foretold_distributions[code]:
return foretold_distributions[code]["infectionFatalityRatio"].sample_community()
v = get_parameter(code,"infectionFatalityRatio")
if not np.isnan(v):
return v
return min(latest_deaths(code) / latest_deaths(code) + latest_recovered(code), max_mortality_rate)
def growth_rate(code):
"""How much does the number of cases grow every day?"""
# This might give bad estimates for the growth rate if a country had few cases at days_to_death days ago.
# We could make this estimate better if we look for the first date that a country crosses some # of cases threshold (e.g. 100 cases)
growth_multiplier = min(confirmed_n_days_ago(code, 0) / confirmed_n_days_ago(code, growth_window_days), max_growth_multiplier)
return growth_multiplier ** (1/growth_window_days)
for code in codes_to_predict:
active_estimates = []
def add_estimate(value, n):
if not isinstance(value, torch.Tensor):
value = torch.tensor(value)
if not torch.isnan(value):
active_estimates.append(value)
ergo.tag(value, f"{code} active_estimate_{n}")
# Model 1 - country specific mortality rate
deaths_estimate = death_multiplier(code) * latest_deaths(code)
cumulative_infections = deaths_estimate * (growth_rate(code) ** days_to_death(code)) * (1 / mortality_rate(code))
recovered_infections = cumulative_infections / (growth_rate(code) ** (recovery_time))
active_estimate_1 = cumulative_infections - recovered_infections
add_estimate(active_estimate_1, 1)
# Model 2 - global mortality rate
deaths_estimate = death_multiplier(code) * latest_deaths(code)
cumulative_infections = deaths_estimate * (growth_rate(code) ** days_to_death(code)) * (1 / canonical_mortality_rate)
recovered_infections = cumulative_infections / (growth_rate(code) ** (recovery_time))
active_estimate_2 = cumulative_infections - recovered_infections
add_estimate(active_estimate_2, 2)
# Model 3 - fraction of cases ascertained
recovered_estimate = confirmed_n_days_ago(code, 0) / (growth_rate(code) ** days_to_death(code))
active_estimate_3 = (confirmed_n_days_ago(code, 0) - recovered_estimate) * ascertainment_parameter(code)
add_estimate(active_estimate_3, 3)
# Model 4 - foretold prediction
active_estimate_4 = sample_foretold(code, predict_date)
add_estimate(active_estimate_4, 4)
if active_estimates:
active_estimate_combined = ergo.random_choice(active_estimates)
ergo.tag(active_estimate_combined, f"{code} active_estimate_combined")
# Get samples from model for all variables
samples = ergo.run(lambda: model(codes_to_predict), num_samples=200)
# + id="QD4K1_cZ3wxr" colab_type="code" outputId="4fd82093-c656-4f50-d9e3-7c9b1d2f908a" colab={"base_uri": "https://localhost:8080/", "height": 639}
samples.describe().transpose().round(0)
# + [markdown] id="VI0InMEyd7Sh" colab_type="text"
# ## Output File
# + id="YZyo2q3MAJWQ" colab_type="code" outputId="a39a8216-6b1b-4603-90a6-603193c07a19" colab={"base_uri": "https://localhost:8080/", "height": 579}
# Make the output dataframe
def make_quantiles_df(codes_to_predict, samples):
quantiles =[0.05 * i for i in range(21)]
data = []
codes = []
for code in codes_to_predict:
name = f"{code} active_estimate_combined"
if name in samples:
series = samples[name]
codes.append(code)
data.append({
"Name":regions[code].get_display_name(),
"Date":str(predict_date).split("T")[0],
**{
str(round(q,2)): series.quantile(q) for q in quantiles
}})
df = pd.DataFrame(data, index=codes)
df.index.name = "Code"
return df
quantiles_df = make_quantiles_df(codes_to_predict, samples).round(0)
display(quantiles_df)
quantiles_df.to_csv("active_infections_prediction.csv")
# If this doesn't work, you can find this file in the files pane at the side of
# the screen, right click and download
time.sleep(5) # I think it's failing sometimes because the file isn't available yet?
try:
google.colab.files.download("active_infections_prediction.csv")
except:
pass
# + id="SIBbJYr17yv6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 554} outputId="4b44ffa5-78a0-46c3-ba47-e76ce4cce071"
# Can plot selected distributions for debugging
for code in ["US", "AE"]:
try:
s = samples[f"{code} active_estimate_combined"]
seaborn.distplot(s)
plt.show()
except KeyError:
pass
# + [markdown] id="4qSamXO1f6sS" colab_type="text"
# # Countermeasures
#
# This part of the notebook allows you to do scenario modelling using probability distributions, and then automatically turn those scenarios into GLEAMviz definition files [1].
#
# ### Basic usage
#
# Scenarios are specified in spreadsheets; where each free parameter can either be specified as a number or as the ID of a Foretold distribution.
#
# [This document](https://docs.google.com/document/d/1CJmoL0ypDTKjtpVLdyVTajeqKRp4HKujEduavkwobl8/edit?usp=sharing) explains how to format the spreadsheet, and [here](https://docs.google.com/spreadsheets/d/1kAboAHnu2KK8p-1adM8L4ngrMfr2EmOsVICwfFKCEPE/edit?usp=sharing) is an example spreadsheet. Any Foretold distribution should be added to [this channel](https://www.foretold.io/c/93c557b5-ac8d-4201-a6d7-7ca2d2574304).
#
# Then use the function: `make_countermeasures_csv(countermeasures_url, countermeasures_sheet)`
#
#
# ---
#
# [1] Note: at the moment this outputs a CSV version of the spreadsheet, [this issue](https://github.com/epidemics/covid/issues/406) tracks when it will be able to automatically generate gleam files.
#
# + id="5-QjehJ1gAI0" colab_type="code" outputId="558f6057-fd82-452d-822f-6f686d6c0027" colab={"base_uri": "https://localhost:8080/", "height": 567, "referenced_widgets": ["84bf18327c434c7bab21e7fbae08d4c9", "c556210528f3432d9081f43c91f3080a", "a01e293ee19b4995a6399f62c39f7c3d", "<KEY>", "<KEY>", "bc260f02894e464496fea97da059f11d", "b81e376ba0c34c65a28a0670652104fa", "9afa5dc44e1148faa8812fcdcffa0964"]}
# Make countermeasures CSV
def make_countermeasures_csv(countermeasures_url, countermeasures_sheet):
input_data = load_spreadsheet(countermeasures_url, countermeasures_sheet)
headers = input_data[0]
value_column = headers.index("Value")
output_data = []
num_samples = 100
quantiles =[0.05 * i for i in range(21)]
for row in tqdm.notebook.tqdm(input_data[1:]):
d = {h: v for (h,v) in zip(headers, row)}
value = row[value_column]
if is_uuid(value):
foretold_question = foretold.get_question(value)
d.update({
str(round(q,2)): foretold_question.quantile(q) for q in quantiles
})
else:
d.update({
str(round(q,2)): value for q in quantiles
})
output_data.append(d)
countermeasures_df = pd.DataFrame(output_data)
display(countermeasures_df)
countermeasures_df.to_csv("countermeasures.csv")
# If this doesn't work, you can find this file in the files pane at the side of
# the screen, right click and download
time.sleep(1)
try:
google.colab.files.download("countermeasures.csv")
except:
pass
make_countermeasures_csv("https://docs.google.com/spreadsheets/d/1kAboAHnu2KK8p-1adM8L4ngrMfr2EmOsVICwfFKCEPE/edit#gid=0",
"Sheet1")
# + [markdown] id="OP-86_ROSf81" colab_type="text"
# ## Generate gleam files
#
# In `epimodel.gleam.definition`, there is a class `GleamDefinition` with many methods for manipulating a given definition XML - add seed regions, add exceptions, adjust parameters.
#
# For example of usage, see `generate_simulations`. It generates an entire batch for the website, but the main steps are the same.
#
# One of the steps is to take region estimates (e.g. countries or states) and distribute these down to Gleam "basins" (city-like regions). This is done by our function `algorithms.distribute_down_with_population` in the above.
# The modelers need a starting XML - I would let them provide it if they want to modify the disease params etc.
| notebooks/covid-19-active.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="o0OxG-eQ9MAP"
# **Exemplo 2 Carregando modelo pré-treinado utilizando base propria**
#
# * Primeiro passo, habilite o drive em seu colab ao lado esquerdo da IDE
#
# * Segundo passo, adicione a base CatsAndDogs em sua pasta Meu Drive no seu Drive
#
# * Link para base ->: https://drive.google.com/drive/folders/1srR7tCUnM9MVqB9x8RGaK9SG7BrhToJt?usp=sharing
#
# * Habilite a funcionalidade de GPU ou TPU na plataforma
#
# * Editar - Configurações de notebook - GPU ou TPU
# + [markdown] id="AFvP0gyGacsn"
# # Nova seção
# + id="KVGyvtAtCrHP"
import tensorflow as tf
import numpy as np
import pandas as pd
from sklearn.metrics import classification_report, confusion_matrix
import matplotlib.pyplot as plt
import seaborn as sn
import os
epochs = 5
batch = 32
#carrega o modelo da inception_v3 com os pesos aprendidos no treino da ImageNet sem a camada densa (include_top=False)
base_model = tf.keras.applications.ResNet50(weights='imagenet', include_top=False)
#O restante do modelo e suas camadas são discutidos a seguir
#x recebe o final da inception_v3
x=base_model.output
#Nova configuração para o modelo
#adiciona apos x uma camada AveragePooling2D e atribui este no a x novamente (logo x e o topo novamente)
x=tf.keras.layers.GlobalAveragePooling2D()(x)
#adiciona apos x uma camada densa com 32 neuronios com funcao de ativacao relu. Atribui este no a x novamente
x=tf.keras.layers.Dense(128,activation='relu')(x)
#adiciona apos x uma camada densa com 64 neuronios com funcao de ativacao relu. Atribui este no a x novamente
x=tf.keras.layers.Dense(64,activation='relu')(x)
#adiciona apos x uma camada densa com 128 neuronios com funcao de ativacao relu. Atribui este no a x novamente
x=tf.keras.layers.Dense(32,activation='relu')(x)
#adiciona após x os neurônios que devem ser utilizados, nesse caso foram desligados 20% dos neuronios
x=tf.keras.layers.Dropout(0.5)(x)
#adiciona apos x uma camada densa com 2 neuronios (duas classes) com funcao de ativacao softmax (distribuicao de probabilidade). Atribui este no a preds
preds=tf.keras.layers.Dense(2,activation='softmax')(x)
#definindo modelo final
model=tf.keras.models.Model(inputs=base_model.input,outputs=preds)
#mostrando modelo final e sua estrutura
model.summary()
#congelando os neuronios já treinados na ImageNet, queremos retreinar somente a ultima camada
for l in model.layers:
if l.name.split('_')[0] != 'dense':
l.trainable=False
else:
l.trainable=True
#iniciando objeto que apanhara todas as imagens de treino, processando as imagens com o metodo da InceptionV3
train_data_gen = tf.keras.preprocessing.image.ImageDataGenerator(preprocessing_function=tf.keras.applications.resnet50.preprocess_input) #included in our dependencies
#iniciando objeto que apanhara todas as imagens de teste, processando as imagens com o metodo da InceptionV3
test_data_gen = tf.keras.preprocessing.image.ImageDataGenerator(preprocessing_function=tf.keras.applications.resnet50.preprocess_input)
#CARREGANDO PRÓPRIO DATASET PARA USO
#definindo gerador de imagens de treino
train_generator = train_data_gen.flow_from_directory('CatsAndDogs/Train',
target_size=(224, 224), # tamanho da imagem para o generator
batch_size=batch,
class_mode='categorical',
shuffle=True)
#definindo gerador de imagens de teste
test_generator = train_data_gen.flow_from_directory('CatsAndDogs/Test',
target_size=(224, 224), # tamanho da imagem para o generator
batch_size=batch,
class_mode='categorical',
shuffle=True)
lr = tf.keras.optimizers.Adam(learning_rate=0.0001)#estabelecendo taxa de otimização
model.compile(optimizer=lr, loss='categorical_crossentropy', metrics=['accuracy'])
#definicao dos steps
step_size_train=train_generator.n//train_generator.batch_size
step_size_test = test_generator.n//test_generator.batch_size
#treinando e testando o modelo
history = model.fit_generator(generator=train_generator,
steps_per_epoch=step_size_train,
epochs=epochs,
validation_data=test_generator,
validation_steps=step_size_test)
#Avaliando o modelo
loss_train, train_acc = model.evaluate_generator(train_generator, steps=step_size_train)
loss_test, test_acc = model.evaluate_generator(test_generator, steps=step_size_test)
print('Train: %.3f, Test: %.3f' % (train_acc, test_acc))
#Apresentando resultados em graficos
plt.title('Loss')
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
# Criando graficos para visualização dos resultados
print()
print()
plt.title('Accuracy')
plt.plot(history.history['accuracy'], label='train')
plt.plot(history.history['val_accuracy'], label='test')
plt.legend()
plt.show()
# +
print('Criando classificações..')
labels = os.listdir('CatsAndDogs/Train')
print('Rótulos', labels)
#criando estruturas para métricas de avaliação, processo um pouco mais demorado
Y_pred = model.predict_generator(test_generator)
print('Preds Created')
y_pred = np.argmax(Y_pred, axis=1)
print('Preds 1D created')
classification = classification_report(test_generator.classes, y_pred, target_names=labels)
print('----------------CLASSIFICATION--------------')
print(classification)
matrix = confusion_matrix(test_generator.classes, y_pred)
df_cm = pd.DataFrame(matrix, index = [i for i in range(2)],
columns = [i for i in range(2)])
plt.figure(figsize = (10,7))
sn.heatmap(df_cm, annot=True, linewidths=2.5)
# -
| src/Codigo Suporte/Processamento_de_Imagens_CNNs_Exemplos_2_e_3_pynb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import numpy as np
# + deletable=true editable=true
import matplotlib.pyplot as plt
# + deletable=true editable=true
y = [1, 7, 3, 5, 12]
x = [1, 2, 3, 4, 5]
# + deletable=true editable=true
plt.plot(x, y, marker='o');
# + deletable=true editable=true
y = [1, 7, 3, 5, 12]
x = [1, 2, 3, 4, 5]
plt.plot(x, y, marker='o');
# + deletable=true editable=true
# %matplotlib
# + deletable=true editable=true
y = [1, 7, 3, 5, 12, 1, 3, 5, 7, 9, 0, 2, 5, 2, 1, 6, 8, 12, 15, 2, 9, 5, 7, 2, 1, 4, 7, 8, 7, 12, 15, 8, 6, 7]
x = [1, 2, 3, 4, 5, 2, 4, 6, 8, 10, 11, 12, 21, 4, 7, 1, 8, 7, 6, 5, 4, 2, 9, 8, 7, 19, 18, 22, 21, 19, 17, 23, 16, 5]
# + deletable=true editable=true
plt.scatter(x, y, marker='o');
# -
| TCC 01/.ipynb_checkpoints/Cluster-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime
import pandas as pd
import pywt
from pyspark.sql import SparkSession
from pyspark.sql.types import StructType, StructField, StringType, TimestampType, FloatType, IntegerType, ArrayType
from pyspark.sql import functions as F
from pyspark.ml.linalg import Vectors, VectorUDT
# -
spark = SparkSession.builder.getOrCreate()
schema = StructType([
StructField("a", IntegerType()),
StructField("b", IntegerType()),
StructField("c", IntegerType()),
StructField("d", ArrayType(FloatType()))
])
# +
rows = [
[1, 2, 3, [0.1, 0.2, 0.3, 0.4]],
[1, 2, 3, [0.1, 0.2, 0.3, 0.4]],
[1, 2, 3, [0.1, 0.2, 0.3, 0.4]]
]
df = spark.createDataFrame(data=rows, schema=schema)
df.show()
# +
import numpy as np
@F.udf(returnType=VectorUDT())
def tovector(x):
return Vectors.dense(x)
@F.udf(returnType=ArrayType(FloatType()))
def dwt(x):
cA, _ = pywt.dwt(x, wavelet="haar")
return cA.tolist()
# -
df.select(tovector(dwt("d"))).show(truncate=False)
| notebook/eda/Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DOaVwYVGt2Wl"
# # FordA data Visualization
# - get tsv file from google drive(alraedy get by get_data_from_web.ipynb)
# - time series data
# - columns: label, sensor data 1 ~ 500
# - no column name
# -> add column name in this code
# - 1: normal -> change to 0 (for generalization)
# - -1: failure -> change to 1
# - row: 3600 for train data
# + [markdown] id="aA5SIW56wPic"
# ## mount to Google drive
# + colab={"base_uri": "https://localhost:8080/"} id="Tk3nxtWXdw6N" outputId="572d4e79-11f1-4864-959a-8499216a8b0a"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="On3KubXVd3Sx" outputId="3d6b3df1-c8dd-4d57-91a8-3c635caac1ac"
# cd drive/My\ Drive/Colab\ Notebooks/summer_coop/FordA
# + [markdown] id="XH2KoFVTujm7"
# ## naming the column
# - since in data, there is no column name, add column name manually
# - total 501 columns: class, sensor1 ... sensor500
# + colab={"base_uri": "https://localhost:8080/"} id="zPgQMQOgjqah" outputId="6522f6b9-d62e-4ea2-a8bf-ff4c7109419d"
column_name = []
column_name.append('failure')
for i in range(1, 501):
name = 'sensor' + str(i)
column_name.append(name)
print(column_name[-1:])
# + [markdown] id="PUU08yemwSch"
# ## Get data and check data
# + id="6Zppsh1veM73"
import pandas as pd
# get tsv file
data = pd.read_csv('FordA_TRAIN.tsv', delimiter = '\t', header = None)
test_data = pd.read_csv('FordA_TEST.tsv', delimiter = '\t', header = None)
# + id="wYfUGamIkrs7"
data.columns = column_name # assign column name
data_full = data.to_numpy()
test_full = test_data.to_numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 249} id="cvkYmek0eeM_" outputId="6cb1f6f7-26a9-469c-e04d-17e9eb252f78"
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="mqUrm4gZenD6" outputId="f1353e0e-6e29-481e-a411-352d7b9d6d20"
data.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 341} id="ZkJo1cLien2H" outputId="0c204d1d-7a18-4020-932e-b4646e2bb0ce"
data.describe()
# + [markdown] id="vE4HXf4cv8nd"
# ### check is there any NUll data
# - no NULL data, all columns have 3601 rows
# + colab={"base_uri": "https://localhost:8080/"} id="SZHM-7UDvcFc" outputId="cf1f7d9b-27a0-45b8-cd20-3111c9e151bc"
info = data.describe()
print(info.loc['count', :].min(), info.loc['count', :].max())
# + colab={"base_uri": "https://localhost:8080/"} id="W2BTwZBAeqmd" outputId="0a4ae9c8-2040-44ef-b7e8-9a03bed7d35f"
data.shape
# + [markdown] id="J0f8yP6ewaQQ"
# ## Split features and class
# + id="9bjAcNA1e8Mn"
train_target = data_full[:, 0]
train_input = data_full[:, 1:]
test_target = test_full[:, 0]
test_input = test_full[:, 1:]
# + colab={"base_uri": "https://localhost:8080/"} id="FoS833h7nu31" outputId="6fbd01a1-f951-4fe8-834d-a013f2576bf8"
print(train_target)
print(train_input)
# + [markdown] id="tqKUjcFNwfCW"
# ### change class's value for generalization
# - normal: 1 -> 0
# - failure: -1 -> 1
#
# + colab={"base_uri": "https://localhost:8080/"} id="G1DYzl58hDaH" outputId="3036c120-c459-433a-ff33-945bbb6d8c97"
train_target [train_target == 1] = 0
train_target[train_target == -1] = 1
print(train_target)
# + id="UXO-Val4BsnD"
test_target[test_target == 1] = 0
test_target[test_target == -1] = 1
# + colab={"base_uri": "https://localhost:8080/"} id="Y90lI4SBBK6e" outputId="37431821-7c87-4cb0-d3e2-a0d6a0617b6a"
data_full[:100, 0]
# + colab={"base_uri": "https://localhost:8080/"} id="MP_7Kl2pB6as" outputId="57d6fed7-f263-4772-e12a-1fe7b0c6343d"
test_full[:100]
# + colab={"base_uri": "https://localhost:8080/"} id="uuaKjvxmnZPH" outputId="f999ddc7-86c1-4dd7-a6d2-1181ec96a196"
print("No. of failure: \t" + str(train_target.sum()))
# + id="i4EfH3pT47pL"
import numpy as np
np.save('FordA_modified.npy', data_full)
np.save('FordA_test_modified.npy', test_full)
# + [markdown] id="NcHuN3_fws6t"
# ## Check correlation between columns
# + id="pZKSZocenmzn"
corelation = data.corr()
# + colab={"base_uri": "https://localhost:8080/", "height": 249} id="V9rhcHJ1o-kp" outputId="5e40bff7-8a82-4262-e3fe-45c0dd20b59b"
corelation.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 413} id="k1X-Vh8_pizU" outputId="f26063d0-0cd4-405a-d028-6cd3440c07bb"
corelation.loc[:, ['failure']]
# + [markdown] id="HNPoGqyUwyI-"
# ### Visualize correlation between failture and sensors
# - some sensors have high correlation with failure
# - can be helpful using the data for machine learning
# + colab={"base_uri": "https://localhost:8080/", "height": 386} id="rMBxe4TxodXn" outputId="c3778ab4-772c-4075-db83-c06779dd73b6"
import matplotlib.pyplot as plt
plt.figure(figsize = (50, 10))
plt.stem(range(500),corelation.loc['sensor1':, 'failure'], '-.', use_line_collection= True)
plt.xlabel('sensor no.')
plt.ylabel('correlation')
plt.title('correlation between failure and sensors')
plt.savefig('FordA_Correlation_bw_failure_sensors.png')
plt.show()
# + id="7FFi8U2sovIG"
| FordA_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ["CUDA_VISIBLE_DEVICES"]="0"
os.environ["OMP_NUM_THREADS"]="1"
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
import numpy as np
import math
import sys
import glob
from tensorflow.keras.utils import to_categorical
import time
import pickle
exp_name = 'track_seq4_vin_intsum_predv'
# # DATA
# +
v_max = 0
v_min = 0
def read_data_n_label(data, label):
global v_max, v_min
input_data = np.load(data)[:,[0,1,3,2,9]]
input_data[:,2] = np.log(input_data[:,2])/16.
input_data = input_data[input_data[:,0] > 0]
input_data = input_data[input_data[:,0] < 60]
input_data = input_data[input_data[:,1] > -40]
input_data = input_data[input_data[:,1] < 40]
grid_in = np.zeros((1,600,800,2))
count_in = np.ones((1,600,800,1))
for inz in input_data:
pos_x = int(inz[0]//0.1)
pos_y = int((inz[1] + 40)//0.1)
grid_in[0, pos_x, pos_y, 0] += inz[2]#(grid_in[0, pos_x, pos_y, 0] + inz[2])#/(count_in[0, pos_x, pos_y, 0])
grid_in[0, pos_x, pos_y, 1] = (grid_in[0, pos_x, pos_y, 1]*(count_in[0, pos_x, pos_y, 0]-1) + inz[3])/count_in[0, pos_x, pos_y, 0]
count_in[0, pos_x, pos_y, 0] += 1
with open(label) as f:
content = f.readlines()
content = [x.strip() for x in content]
grid_label = np.zeros((1,300,400,8))
if len(content) == 0:
return grid_in, grid_label, np.array([-1,-1,-1,-1])
velocity = (input_data[:,3][input_data[:,-1]>=0])
velocity = velocity.mean() if velocity.size>0 else np.zeros(1)
velocity = velocity/4
tar_location = np.fromstring(content[1][1:-1], dtype= float, sep = ' ')
tar_WL = np.fromstring(content[2][1:-1], dtype= float, sep = ',')
tar_angle = float(content[3])
pos0 = int((tar_location[0])//0.2)
pos1 = int((tar_location[1] + 40)//0.2)
grid_label[0, pos0, pos1, -1] = 1
grid_label[0, pos0, pos1, -2] = tar_angle
grid_label[0, pos0, pos1, 3] = tar_WL[0]
grid_label[0, pos0, pos1, 4] = tar_WL[1]
if tar_WL[0] < tar_WL[1]:
grid_label[0, pos0, pos1, 3] = tar_WL[1]
grid_label[0, pos0, pos1, 4] = tar_WL[0]
grid_label[0, pos0, pos1, -2] = tar_angle + np.pi/2
# print ('ALARM')
grid_label[0, pos0, pos1, 1] = tar_location[0] - (pos0*0.2) - 0.1
grid_label[0, pos0, pos1, 2] = tar_location[1] - (pos1*0.2 -40) -0.1
grid_label[0, pos0, pos1, 5] = velocity
if content[0] == 'bike':
grid_label[0, pos0, pos1, 0] = 1
elif content[0] == 'pedestrian':
grid_label[0, pos0, pos1, 0] = 2
else:
grid_label[0, pos0, pos1, 0] = 0
if grid_label[0, pos0, pos1, -2]>(v_max):
# v_max=grid_label[0, pos0, pos1, -2]
v_max=grid_in[0, :, :, 0].max()
if grid_label[0, pos0, pos1, -2]<(v_min):
# v_min=grid_label[0, pos0, pos1, -2]
v_min=grid_in[0, :, :, 0].min()
return grid_in, grid_label, np.array([tar_location[0], tar_location[1], tar_WL[0], tar_WL[1]])
# X.shape, y[:,:,:,0].shape
# -
training_folders = glob.glob('training_data/*')
training_labels = glob.glob(training_folders[0] + '/labels/radar_left_np/*')
training_X = glob.glob(training_folders[0] + '/radar_left_np/*')
training_labels.sort()
training_X.sort()
X = []
y = []
for j in range(len(training_folders)):
training_labels = glob.glob(training_folders[j] + '/labels/radar_left_np/*')
training_X = glob.glob(training_folders[j] + '/radar_left_np/*')
# print (j,training_folders[j])
for i in range(len(training_labels)):
X_in, y_in, location = read_data_n_label(training_X[i], training_labels[i])
X.append(X_in)
y.append(y_in)
X = np.array(X)
X = X.reshape(len(X),600,800,2)
y = np.array(y)
y = y.reshape(len(y),300,400,8)
def feature_extractor(image_in):
with tf.variable_scope('feature_extractor'):
net = tf.layers.conv2d(image_in, 8, 3,strides=2, activation=tf.nn.relu, padding='same')
net = tf.layers.conv2d(net, 8, 3, strides=1,activation=tf.nn.relu, padding='same')
net = tf.layers.conv2d(net, 8, 3, strides=1,activation=tf.nn.relu, padding='same')
x1 = net
net = tf.layers.conv2d(net, 16, 3,strides=2, activation=tf.nn.relu, padding='same')
net = tf.layers.conv2d(net, 32, 3, strides=1,activation=tf.nn.relu, padding='same')
net = tf.layers.conv2d(net, 32, 3, strides=1,activation=tf.nn.relu, padding='same')
x2 = net
return x1, x2
# +
images_in = tf.placeholder(tf.float32, (None,600,800,2,4))
y_label = tf.placeholder(tf.int32, (None,300, 400, 4, 1 ))
y_label_one_hot = tf.one_hot(y_label, 3, axis=-1)
y_angle = tf.placeholder(tf.float32, (None,300, 400 , 4, 1))
y_wl = tf.placeholder(tf.float32, (None,300, 400 , 4, 2))
y_loc = tf.placeholder(tf.float32, (None,300, 400 , 4, 2))
y_v = tf.placeholder(tf.float32, (None,300, 400 , 4, 1))
y_mask = tf.placeholder(tf.float32, (None,300, 400 , 4))
with tf.variable_scope("features") as scope:
x11, x12 = feature_extractor(images_in[:,:,:,:,0])
scope.reuse_variables()
x21, x22 = feature_extractor(images_in[:,:,:,:,1])
scope.reuse_variables()
x31, x32 = feature_extractor(images_in[:,:,:,:,2])
scope.reuse_variables()
x41, x42 = feature_extractor(images_in[:,:,:,:,3])
x1 = tf.concat([x11, x21, x31, x41], axis = -1)
x2 = tf.concat([x12, x22, x32, x42], axis = -1)
up1 = tf.layers.conv2d_transpose(x1, 32, 3, activation=tf.nn.relu, padding='same')
up2 = tf.layers.conv2d_transpose(x2, 64, 3,strides=2, activation=tf.nn.relu, padding='same')
concat = tf.concat([up1, up2], axis = -1)
# -
def prediction_head(concat_vec, y_label_one_hot_ts, y_angle_ts, y_loc_ts, y_wl_ts, y_v_ts, y_mask_ts, name):
with tf.variable_scope(name):
y_pred_occ = tf.layers.conv2d(concat_vec, 3 , 1, activation=None, padding= 'same')
y_pred_occ_prob = tf.nn.softmax(y_pred_occ, axis = -1)
y_pred_angle = tf.layers.conv2d(concat_vec, 1 , 1, activation=None, padding= 'same')
y_pred_loc = tf.layers.conv2d(concat_vec, 2 , 1, activation=None, padding= 'same')
y_pred_WL = tf.layers.conv2d(concat_vec, 2 , 1, activation=tf.nn.softplus, padding= 'same')
y_pred_v = tf.layers.conv2d(concat_vec, 1 , 1, activation=None, padding= 'same')
log_y_pred_WL = tf.log(y_pred_WL + 0.1)
log_y_wl = tf.log(y_wl_ts + 0.1)
loss_occ = tf.reduce_mean(-y_label_one_hot_ts[:,:, : , 0] *0.008*((1 - y_pred_occ_prob[:,:,:,0])) * tf.log(y_pred_occ_prob[:,:,:,0] + 0.01 )\
-y_label_one_hot_ts[:,:, : , 1] *0.992*((1 - y_pred_occ_prob[:,:,:,1])) * tf.log(y_pred_occ_prob[:,:,:,1] + 0.01)\
-y_label_one_hot_ts[:,:, : , 2] *0.992*((1 - y_pred_occ_prob[:,:,:,2])) * tf.log(y_pred_occ_prob[:,:,:,2] + 0.01)\
)
loss_angle = tf.reduce_mean(y_mask_ts*tf.abs(tf.sin(y_pred_angle - y_angle_ts)))
loss_loc = tf.reduce_mean(y_mask_ts*tf.abs(y_loc_ts- y_pred_loc))
loss_wl = tf.reduce_mean(y_mask_ts*tf.abs(log_y_wl - log_y_pred_WL))
loss_v = tf.reduce_mean(y_mask_ts*tf.abs(y_v_ts - y_pred_v)*(y_v_ts!=0))
total_loss = 1.0*loss_occ + 1.25*loss_angle + 0.75*loss_loc + loss_wl + 1 * loss_v
return y_pred_occ_prob, y_pred_angle, y_pred_loc, y_pred_WL, y_pred_v, total_loss
# +
y_pred_occ_prob0, y_pred_angle0, y_pred_loc0, y_pred_WL0, y_pred_v0, total_loss0 = prediction_head(concat, y_label_one_hot[:,:,:,0,0,:], y_angle[:,:,:,0,:], \
y_loc[:,:,:,0,:], y_wl[:,:,:,0,:], y_v[:,:,:,0,:], tf.expand_dims(y_mask[:,:,:,0], axis = -1), 'prediction_0')
y_pred_occ_prob1, y_pred_angle1, y_pred_loc1, y_pred_WL1, y_pred_v1, total_loss1 = prediction_head(concat, y_label_one_hot[:,:,:,1,0,:], y_angle[:,:,:,1,:], \
y_loc[:,:,:,1,:], y_wl[:,:,:,1,:], y_v[:,:,:,1,:], tf.expand_dims(y_mask[:,:,:,1], axis = -1), 'prediction_1')
y_pred_occ_prob2, y_pred_angle2, y_pred_loc2, y_pred_WL2, y_pred_v2, total_loss2 = prediction_head(concat, y_label_one_hot[:,:,:,2,0,:], y_angle[:,:,:,2,:], \
y_loc[:,:,:,2,:], y_wl[:,:,:,2,:], y_v[:,:,:,2,:], tf.expand_dims(y_mask[:,:,:,2], axis = -1), 'prediction_2')
y_pred_occ_prob3, y_pred_angle3, y_pred_loc3, y_pred_WL3, y_pred_v3, total_loss3 = prediction_head(concat, y_label_one_hot[:,:,:,3,0,:], y_angle[:,:,:,3,:], \
y_loc[:,:,:,3,:], y_wl[:,:,:,3,:], y_v[:,:,:,3,:], tf.expand_dims(y_mask[:,:,:,3], axis = -1), 'prediction_3')
total_loss = total_loss0 + 0.75*total_loss1 + 0.5*total_loss2 + 0.25*total_loss3
# -
v_min,v_max #angle(-1.5687682499618538, 1.5707963267948966)
#velocity (-4.0950820584830465, 4.674418471324403)
#intensity (0.0, 4.248383107944953)
# +
def processing_seq_data(X, Y):
b_x = []
b_y = []
Y = np.expand_dims(Y, axis = 3)
X = np.expand_dims(X, axis = -1)
for i in range(len(X) -6):
b_x.append(np.concatenate((X[i], X[i+1], X[i+2], X[i+3]), axis = -1))
b_y.append(np.concatenate((Y[i+3], Y[i+4], Y[i+5], Y[i+6]), axis = 2))
b_x = np.asarray(b_x)
b_y = np.asarray(b_y)
return b_x, b_y
def dataset(np_seed, seqlen=10):
Xs = []
labels = []
training_folders = glob.glob('training_data/*')
for j in range(len(training_folders)):
training_labels = glob.glob(training_folders[j] + '/labels/radar_left_np/*')
training_X = glob.glob(training_folders[j] + '/radar_left_np/*')
training_labels.sort()
training_X.sort()
i_begin = np.random.RandomState(np_seed).randint(seqlen)
training_labels = training_labels[i_begin:]
training_X = training_X[i_begin:]
n_seq = len(training_X)//seqlen
i_end = n_seq * seqlen
training_labels = training_labels[:i_end]
training_X = training_X[:i_end]
# print(len(training_labels))
Xs.extend(training_X)
labels.extend(training_labels)
return Xs, labels
# +
#shuffle 1
np_seed = 12345
optimizer = tf.train.AdamOptimizer().minimize(total_loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver(max_to_keep=20)
seqlen = 10
with tf.Session() as sess:
sess.run(init)
for i_epoch in range(30):
batch_X = []
batch_y = []
t_begin = time.time()
Xs, labels = dataset(np_seed+i_epoch,seqlen=seqlen)
n_seq = len(Xs)
print(n_seq)
seq_shuffle = np.random.RandomState(np_seed+i_epoch).permutation(n_seq//seqlen)
for i, i_seq in enumerate(seq_shuffle):
for ii in range(seqlen):
X_in, y_in, _ = read_data_n_label(Xs[i_seq*seqlen+ii], labels[i_seq*seqlen+ii])
# print (i,' ',i_seq,' ',ii,' ',i_seq*seqlen+ii)
# print(Xs[i_seq*seqlen+ii])
batch_X.append(X_in)
batch_y.append(y_in)
batch_X_seq = np.array(batch_X).reshape(seqlen,600,800,2)
batch_y_seq = np.array(batch_y).reshape(seqlen,300,400,8)
batch_X, batch_y = processing_seq_data(batch_X_seq, batch_y_seq)
pred_occ, cost,_= sess.run([y_pred_occ_prob0,total_loss, optimizer], feed_dict = {images_in: batch_X,\
y_label: batch_y[:,:,:,:,0:1], \
y_angle: batch_y[:,:,:,:,-2:-1],\
y_loc: batch_y[:,:,:,:,[1,2]], \
y_wl: batch_y[:,:,:,:,[3,4]], \
y_v: batch_y[:,:,:,:,5:6],\
y_mask: batch_y[:,:,:,:,-1]})
if i_seq % (n_seq//4)==0:
print('epoch {}'.format(i_epoch))
print ('cost: ', cost)
z = pred_occ[-1]
argmax = z.argmax(axis = 2)
print ('--------------------------------------------------------')
print ('Predictions:',np.where(argmax > 0))
print ('labels:', np.where(batch_y[-1,:,:,0,-1] > 0.5))
print ('Pre_ID: ', np.argmax(pred_occ[-1]),', GT_ID: ', np.argmax(batch_y[-1,:,:,-1]))
batch_X = []
batch_y = []
save_path = saver.save(sess, "saved_models/{}/model_e{}.ckpt".format(exp_name, i_epoch))
t_elapsed = time.time()-t_begin
print('Time Elapsed: ', t_elapsed)
save_path = saver.save(sess, "saved_models/{}/model.ckpt".format(exp_name))
# -
# ## Save Testing Predictions
# exp_name='track-predv0'
for i_epoch in range(11,30):
saver = tf.train.Saver()
t_begin = time.time()
with tf.Session() as sess:
saver.restore(sess, "saved_models/{}/model_e{}.ckpt".format(exp_name, i_epoch))
# saver.restore(sess, "saved_models/model_e{}.ckpt".format( i_epoch))
print("Model restored.")
predictions = {'metadata': [],'label_preds':[], 'scores':[], 'box3d_lidar':[],'velocity':[],'velocity_gt':[]}
counter = 0
training_folders = glob.glob('validation_data/*')
training_folders.sort()
for folder in range(len(training_folders)):
training_labels = glob.glob(training_folders[folder] + '/labels/radar_left_np/*')
print(folder)
training_X = glob.glob(training_folders[folder] + '/radar_left_np/*')
training_labels.sort()
training_X.sort()
X = []
y = []
for i in range(len(training_labels)):
X_in, y_in, location = read_data_n_label(training_X[i], training_labels[i])
X.append(X_in)
y.append(y_in)
X = np.concatenate(X, axis=0)
y = np.concatenate(y, axis=0)
X = np.concatenate([np.tile(X[:1],(6,1,1,1)),X],axis=0)
var_names = ['occ','angle','loc','wl', 'v']
for j in range(4):
for var in var_names:
code = "pred_{}X{}s=[]".format(var,j)
exec(code)
for i in range(3,len(X)):
pred_occX0, pred_angleX0, pred_locX0, pred_wlX0,pred_vX0, pred_occX1, pred_angleX1, pred_locX1, pred_wlX1,pred_vX1,\
pred_occX2, pred_angleX2, pred_locX2, pred_wlX2,pred_vX2, pred_occX3, pred_angleX3, pred_locX3, pred_wlX3,pred_vX3 \
= sess.run([y_pred_occ_prob0, y_pred_angle0, y_pred_loc0, y_pred_WL0,y_pred_v0,\
y_pred_occ_prob1, y_pred_angle1, y_pred_loc1, y_pred_WL1,y_pred_v1,\
y_pred_occ_prob2, y_pred_angle2, y_pred_loc2, y_pred_WL2,y_pred_v2,\
y_pred_occ_prob3, y_pred_angle3, y_pred_loc3, y_pred_WL3,y_pred_v3], \
feed_dict = {images_in: np.stack((X[i-3],X[i-2],X[i-1],X[i]),axis=-1)[np.newaxis]})
for j in range(4):
for var in var_names:
code = "pred_{}X{}s.append(pred_{}X{})".format(var,j,var,j)
exec(code)
for i in range(6, len(X)):
for var in var_names:
for j in range(4):
code = "pred_{}X{}=pred_{}X{}s[{}]".format(var,j,var,j,i-3-j)
exec(code)
code = "pred_{} = np.mean(np.array([pred_{}X0, pred_{}X1, pred_{}X2, pred_{}X3]), axis = 0)".format(var,var,var,var,var)
exec(code)
pred_index = np.asarray(np.where(pred_occ[0,:,:,:].argmax(axis = -1)>0)).T
#pred_index = np.asarray(np.where(pred_occ[0,:,:,0] < 0.5)).T
iii = i -6
try:
v_gt = y[iii][...,5].sum()
except:
v_gt = 0
predictions['velocity_gt'].append(v_gt*4)
predictions['metadata'].append(training_X[iii])
predictions['label_preds'].append([])
predictions['box3d_lidar'].append([])
predictions['scores'].append([])
predictions['velocity'].append([])
for index in range(len(pred_index)):
predictions['label_preds'][counter].append(pred_occ[:,pred_index[index,0], pred_index[index,1], [1,2]].argmax(axis = -1)[0])
predictions['scores'][counter].append(pred_occ[:,pred_index[index,0], pred_index[index,1], [1,2]].max(axis = -1)[0])
predictions['velocity'][counter].append(pred_v[:,pred_index[index,0], pred_index[index,1], 0]*4)
pred_location_x = pred_loc[:,pred_index[index,0], pred_index[index,1],0] + (pred_index[index,0]*0.2) + 0.1
pred_location_y = pred_loc[:,pred_index[index,0], pred_index[index,1],1] + (pred_index[index,1]*0.2 -40) + 0.1
W_location_xy = pred_wl[:,pred_index[index,0], pred_index[index,1],0]
L_location_xy = pred_wl[:,pred_index[index,0], pred_index[index,1],1]
angle_xy = pred_angle[:,pred_index[index,0], pred_index[index,1],0]
predictions['box3d_lidar'][counter].append([])
predictions['box3d_lidar'][counter][-1].append(pred_location_x[0])
predictions['box3d_lidar'][counter][-1].append(pred_location_y[0])
predictions['box3d_lidar'][counter][-1].append(-1)
predictions['box3d_lidar'][counter][-1].append(W_location_xy[0])
predictions['box3d_lidar'][counter][-1].append(L_location_xy[0])
predictions['box3d_lidar'][counter][-1].append(-1)
predictions['box3d_lidar'][counter][-1].append(angle_xy[0])
counter += 1
fname = "saved_models/{}/model_e{}.pkl".format(exp_name, i_epoch)
with open(fname, 'wb') as output:
pickle.dump(predictions, output)
print(time.time()-t_begin)
# +
# save NMS prediction
from shapely.geometry import Polygon
def get_rect(filler):
W = filler[3]
L = filler[4]
angle = filler[-1]
m0 = np.array([filler[0], filler[1]])
k1 = np.array([np.cos(angle), np.sin(angle)])*W
k2 = np.array([np.sin(angle), -np.cos(angle)])*L
p0 = m0 - k1 - k2 # m0 + k1 + k2
p1 = m0 +k1 - k2 #m0 - k1 - k2
p2 = m0 + k1 + k2
p3 = m0 - k1 + k2
return Polygon([p0, p1, p2, p3]), [p0, p1, p2, p3]
# exp_name = 'track_seq4_vin_intsum_predv'
nonobject= True #False
for i_epoch in range(12,30):
fname = "saved_models/{}/model_e{}".format(exp_name, i_epoch)
with open(fname+".pkl", 'rb') as file:
object_file = pickle.load(file)
nms_dict = {'metadata': [],'label_preds':[], 'scores':[], 'box3d_lidar':[], 'velocity':[]}
v_loss = []
for i in range(len(object_file['metadata'])):
score = np.array(object_file['scores'][i])
llabel = np.array(object_file['label_preds'][i])
rrect = np.array(object_file['box3d_lidar'][i])
velo = np.array(object_file['velocity'][i])
"""okay = np.where(score> 0.45)[0]
score = score[okay]
llabel = llabel[okay]
rrect = rrect[okay]"""
nms_dict['metadata'].append(object_file['metadata'][i])
nms_dict['label_preds'].append([])
nms_dict['box3d_lidar'].append([])
nms_dict['scores'].append([])
nms_dict['velocity'].append([])
if len(score)==0:
v_diff = object_file['velocity_gt'][i]
else:
v_diff = None
while(len(score) > 0):
nms_dict['box3d_lidar'][i].append([])
under_attention = score.argmax()
max_score = score.max()
# print(max_score)
class_pred = llabel[under_attention]
#score = np.delete(score, under_attention)
if v_diff is None:
if object_file['velocity_gt'][i]==0:
v_diff = 1
else:
v_diff = (velo[under_attention] - object_file['velocity_gt'][i])/(object_file['velocity_gt'][i])
# print (llabel)
# print (rrect.shape)
# print(score.shape)
box_attent, box_arr = get_rect(rrect[under_attention])
box_apc = rrect[under_attention]
#print (object_file['box3d_lidar'][i])
#nms_dict['box3d_lidar'][i][-1].append(box_arr)
box_arr = np.array(box_arr)
deletion_list = []
avg_conf= []
count = 1.0
for mmm in range(len(score)):
boxmm, boxyy = get_rect(rrect[mmm])
boxyy = np.array(boxyy)
if (box_attent.intersection(boxmm).area / box_attent.union(boxmm).area) > 0:
if nonobject or (not nonobject and class_pred == llabel[mmm]):
deletion_list.append(mmm)
nms_dict['label_preds'][i].append(class_pred)
nms_dict['scores'][i].append(max_score)
nms_dict['box3d_lidar'][i][-1].append(box_apc)
nms_dict['velocity'][i].append(velo[under_attention])
score = np.delete(score, deletion_list)
rrect = np.delete(rrect, deletion_list, axis = 0)
llabel = np.delete(llabel, deletion_list)
v_loss.append(v_diff)
v_loss = np.array(v_loss)
v_loss1 = np.abs(v_loss).mean() #/len(object_file['metadata'])
v_loss2 = (v_loss**2).mean() #/len(object_file['metadata'])
print(i_epoch, 'velocity loss: ', v_loss1,' ', v_loss2)
nms_list = []
for i in range(len(nms_dict['metadata'])):
sub_dict = dict()
sub_dict['metadata'] = dict()
sub_dict['metadata']['image_idx']= nms_dict['metadata'][i]
# sub_dict['metadata']['image_shape']= np.array([ 600, 800])
sub_dict['label_preds'] = np.array(nms_dict['label_preds'][i])
sub_dict['scores'] = np.array(nms_dict['scores'][i])
sub_dict['velocity'] = np.array(nms_dict['velocity'][i])
if len(nms_dict['box3d_lidar'][i]) >0:
sub_dict['box3d_lidar'] = np.array(nms_dict['box3d_lidar'][i]).squeeze(axis = 1)
else:
sub_dict['box3d_lidar'] = np.array(nms_dict['box3d_lidar'][i])
nms_list.append(sub_dict)
if nonobject:
fname +='-n'
with open(fname+'-nms.pkl', 'wb') as output:
pickle.dump(nms_list, output)
| tracking-train-eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import torchvision.models as models
from torch.utils.data import Dataset,DataLoader
from torchvision import transforms
torch.manual_seed(0)
# -
# You should load the resnet18 model,
# setting the parameter pretrained to True,
# this means the model has been trained before.
model = models.resnet18(pretrained = True)
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
composed = transforms.Compose([transforms.Resize(224),
transforms.ToTensor(),
transforms.Normalize(mean, std)])
train_dataset = Dataset(transforms=composed,train = True)
validation_data = Dataset(transforms=composed)
model.state_dict
model.fc
for param in model.parameters():
param.requires_grad = False
model.fc=nn.Linear(512,7)
# +
train_loader = torch.utils.data.DataLoader(dataset=train_dateset, batch_size = 15)
validation_loader = torch.utils.data.DataLoader(dataset=validation_dateset, batch_size = 10)
# +
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam([parameters for parameters in model.parameters() if parameters.requires_grad], lr = 0.003)
# -
n_epochs = 20
loss_list = []
accuracy_list = []
correct = 0
n_test = len(validation_dataset)
| IBM_AI_Engineering/Course-4-deep-neural-networks-with-pytorch/Week-6-CNN/9.5Torch-vision-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SVC
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
import pandas as pd
import numpy as np
from os import path
# +
filename = path.join(".", "data", "exoplanet_data.csv")
df = pd.read_csv(filename)
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df.head()
# -
target = df["koi_disposition"]
# Use the seven most important features identified in the Random Forest model
data = df[['koi_fpflag_co', 'koi_fpflag_nt', 'koi_fpflag_ss', 'koi_model_snr', 'koi_prad', 'koi_prad_err2', 'koi_duration_err2']]
data.head()
# Split the data into train/test
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target, train_size=0.8, random_state=12)
# +
# Scale the data
from sklearn.preprocessing import MinMaxScaler
X_scaler = MinMaxScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
# Support vector machine linear classifier
from sklearn.svm import SVC
model = SVC()
# ## Tune model parameters with GridSearch
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [1, 5, 10, 50],
'gamma': [0.0001, 0.0005, 0.001, 0.005]}
grid = GridSearchCV(model, param_grid, verbose=3)
grid.fit(X_train, y_train)
print(grid.best_params_)
print(grid.best_score_)
# +
predictions = grid.predict(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions))
# -
print('Test Acc: %.3f' % grid.score(X_test, y_test))
| svc_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 0
#
# ### In class exercise ( no submission is required)
#
# ---
#
# Welcome to EMSE6992 (https://bsharvey.github.io/). In this class, we will be using a variety of tools that will require some initial configuration. To ensure everything goes smoothly moving forward, we will setup the majority of those tools in this in class activity. While some of this will likely be dull, doing it now will enable us to do more exciting work in the weeks that follow without getting bogged down in further software configuration. This will not be graded, however it is essential that you complete it timely since it will enable us to set up your environments.
#
#
# ## Programming expectations
#
# All the assignments and labs for this class will use Python and, for the most part, the browser-based Jupyter notebook format you are currently viewing. Knowledge of Python is not a prerequisite for this course, **provided you are comfortable learning on your own as needed**. While we have strived to make the programming component of this course straightforward, we will not devote much time to teaching prorgramming or Python syntax. Basically, you should feel comfortable with:
#
# * How to look up Python syntax on Google and StackOverflow.
# * Basic programming concepts like functions, loops, arrays, dictionaries, strings, and if statements.
# * How to learn new libraries by reading documentation.
#
# There are many online tutorials to introduce you to scientific python programming. [Here is one](https://github.com/jrjohansson/scientific-python-lectures) that is very nice.
#
# ## Getting Python
#
# You will be using Python throughout the course, including many popular 3rd party Python libraries for scientific computing. [Anaconda](http://continuum.io/downloads) is an easy-to-install bundle of Python and most of these libraries. We recommend that you use Anaconda for this course.
#
# All the labs and assignments in this course are Python-based. This page provides information about configuring Python on your machine.
#
# Also see: http://docs.continuum.io/anaconda/install
#
# **Step 0**
# The Jupyter notebook runs in the browser, and works best in Google Chrome or Firefox.
#
# ## Recommended Method: Anaconda
#
# The Anaconda Python distribution is an easily-installable bundle of Python and many of the libraries used throughout this class. Unless you have a good reason not to, we recommend that you use Anaconda.
#
# ### Mac/Linux users
# 1. Download the [appropriate version](http://continuum.io/downloads) of Anaconda
# 1. Follow the instructions on that page to run the installer
# 1. Test it out: open a terminal window, and type ``python``, you should see something like
# ```
# Python 2.7.5 |Anaconda 1.6.1 (x86_64)| (default, Jun 28 2013, 22:20:13)
# ```
# If `Anaconda` doesn't appear on the first line, you are using a different version of Python. See the troubleshooting section below.
#
# 1. Test out the IPython notebook: open a Terminal window, and type `ipython notebook`. A new browser window should pop up. p
# 1. Click `New Notebook` to create a new notebook file
# 1. Update IPython to the newest version by typing `conda update ipython` at the command line
# 1. Open the command prompt (type conda list --> pip install{see libraries below})
#
#
# ### Windows Users
# 1. Download the [appropriate version](http://continuum.io/downloads) of Anaconda
# 1. Follow the instructions on that page to run the installer. This will create a directory at `C:\Anaconda`
# 1. Test it out: start the Anaconda launcher, which you can find in `C:\Anaconda` or, in the Start menu. Start the IPython notebook. A new browser window should open.
# 1. Click `New Notebook`, which should open a new page.
# 1. Update IPython to the newest version by opening a command prompt, and typing `conda update ipython`
#
# If you did not add Anaconda to your path, be sure to use the full path to the python and ipython executables, such as `/anaconda/bin/python`.
#
# 1. Open the Anaconda command prompt (type conda list --> pip install{see libraries below})
#
# ## Installing additional libraries
# Anaconda includes most of the libraries we will use in this course, but you will need to install a few extra ones:
#
# 1. [BeautifulSoup](http://www.crummy.com/software/BeautifulSoup/)
# 1. [Pattern](http://www.clips.ua.ac.be/pattern)
# 1. [Seaborn](http://web.stanford.edu/~mwaskom/software/seaborn/)
# 1. [MrJob](http://pythonhosted.org/mrjob/)
# 1. [PyQuery](https://pythonhosted.org/pyquery/)
#
# The recommended way to install these packages is to run `pip install BeautifulSoup mrjob pattern seaborn pyquery` on the command line. If this doesn't work, you can download the source code, and run `python setup.py install` from the source code directory. On Unix machines, either of these commands may require `sudo` (i.e. `sudo pip install...` or `sudo python`)
#
# ## Opening IPython Notebooks
# To view an IPython notebook, you must first start the IPython notebook server in the directory where the file lives. Simply navigate to this directory at the command prompt, and type `ipython notebook`. This will open a browser window, listing all the `ipynb` files in that directory.
#
# ## Updating from older Anaconda versions
# You can easily update to the latest Anaconda version by updating conda, then Anaconda as follows:
#
# ```
# conda update conda
# conda update anaconda
# ```
#
# ## Troubleshooting
#
# **Problem**
# When you start python, you don't see a line like `Python 2.7.5 |Anaconda 1.6.1 (x86_64)|`. You are using a Mac or Linux computer
#
# **Reason**
# You are most likely running a different version of Python, and need to modify your Path (the list of directories your computer looks through to find programs).
#
# **Solution**
# Find a file like `.bash_profile`, `.bashrc`, or `.profile`. Open the file in a text editor, and add a line at this line at the end: `export PATH="$HOME/anaconda/bin:$PATH"`. Close the file, open a new terminal window, type `source ~/.profile` (or whatever file you just edited). Type `which python` -- you should see a path that points to the anaconda directory. If so, running `python` should load the proper version
#
# **Problem**
# You are running the right version of python (see above item), but are unable to import numpy.
#
# **Reason**
# You are probably loading a different copy of numpy that is incompatible with Anaconda
#
# **Solution**
# See the above item to find your `.bash_profile`, `.profile`, or `.bashrc` file. Open it, and add the line `unset PYTHONPATH` at the end. Close the file, open a new terminal window, type `source ~/.profile` (or whatever file you just edited), and try again.
# ***
#
# **Problem**
# Under Windows, you receive an error message similar to the following: "'pip' is not recognized as an internal or external command, operable program or batch file."
#
# **Reason**
# The correct Anaconda paths might not be present in your PATH variable, or Anaconda might not have installed correctly.
#
# **Solution**
# Ensure the Anaconda directories to your path environment variable ("\Anaconda" and "\Anaconda\Scripts"). See [this page](http://superuser.com/questions/284342/what-are-path-and-other-environment-variables-and-how-can-i-set-or-use-them) for details.
#
# If this does not correct the problem, reinstall Anaconda.
#
#
# ## Hello, Python
#
# The Jupyter notebook is an application to build interactive computational notebooks. You'll be using them to complete labs and homework. Once you've set up Python, please <a https://github.com/bsharvey/EMSEDataAnalytics/blob/master/EMSE6992_Assignments/HW0.ipynb download="HW0.ipynb">download this page</a>, and open it with Jupyter by typing
#
# ```
# jupyter notebook <name_of_downloaded_file>
# ```
#
# ## Portfolios and Setup
#
# For the rest of the lab/assignment, use your local copy of this page, running on Jupyter and bring up a shell.
#
# Download portfolio template from Github
# https://github.com/bsharve/bsharve.github.io
#
# Download assignments and labs from Github
# https://github.com/bsharvey/EMSEDataAnalytics
#
# Useful github commands
# https://gist.github.com/hofmannsven/6814451
#
# Create a github account
#
# Create a username.github.io repository
# Upload bsharve.github.io zip file to to username.github.io repository
#
# Create a {assignments_labs} repository
# Upload EMSEDataAnalytics zip file to to {assignments_labs} repository
#
# Download and Install Github Desktop
# https://desktop.github.com/
#
# Download and Install Atom IDE
# https://atom.io/
#
# //useful github command line comands
# git init
# git branch -m master gh-pages
# git add --all
# git status
# git commit -m "first commit"
# git remote add origin https://github.com/bsharve/bsharve.github.io.git
# //git remote remove origin
# git push -u origin gh-pages
#
#
# Notebooks are composed of many "cells", which can contain text (like this one), or code (like the one below). Double click on the cell below, and evaluate it by clicking the "play" button above, for by hitting shift + enter
x = [10, 20, 30, 40, 50]
for item in x:
print ("Item is ", item)
# ## Python Libraries
#
# We will be using a several different libraries throughout this course. If you've successfully completed the [installation instructions](https://github.com/cs109/content/wiki/Installing-Python), all of the following statements should run.
# +
#IPython is what you are using now to run the notebook
import jupyter
#print ("Jupyter version: %6.6s (need at least 1.0)" % jupyter.__version__)
# Numpy is a library for working with Arrays
import numpy as np
print ("Numpy version: %6.6s (need at least 1.7.1)" % np.__version__)
# SciPy implements many different numerical algorithms
import scipy as sp
print ("SciPy version: %6.6s (need at least 0.12.0)" % sp.__version__)
# Pandas makes working with data tables easier
import pandas as pd
print ("Pandas version: %6.6s (need at least 0.11.0)" % pd.__version__)
# Module for plotting
import matplotlib
print ("Mapltolib version: %6.6s (need at least 1.2.1)" % matplotlib.__version__)
# SciKit Learn implements several Machine Learning algorithms
import sklearn
print ("Scikit-Learn version: %6.6s (need at least 0.13.1)" % sklearn.__version__)
# Requests is a library for getting data from the Web
import requests
print ("requests version: %6.6s (need at least 1.2.3)" % requests.__version__)
# Networkx is a library for working with networks
import networkx as nx
print ("NetworkX version: %6.6s (need at least 1.7)" % nx.__version__)
#BeautifulSoup is a library to parse HTML and XML documents
import beautifulSoup
print ("BeautifulSoup version:%6.6s (need at least 3.2)" % BeautifulSoup.__version__)
#MrJob is a library to run map reduce jobs on Amazon's computers
import mrjob
print ("Mr Job version: %6.6s (need at least 0.4)" % mrjob.__version__)
#Pattern has lots of tools for working with data from the internet
import pattern
print ("Pattern version: %6.6s (need at least 2.6)" % pattern.__version__)
# -
# If any of these libraries are missing or out of date, you will need to [install them](https://github.com/cs109/content/wiki/Installing-Python#installing-additional-libraries) and restart IPython
# ## Hello matplotlib
# The notebook integrates nicely with Matplotlib, the primary plotting package for python. This should embed a figure of a sine wave:
# +
#this line prepares IPython for working with matplotlib
# %matplotlib inline
# this actually imports matplotlib
import matplotlib.pyplot as plt
x = np.linspace(0, 10, 30) #array of 30 points from 0 to 10
y = np.sin(x)
z = y + np.random.normal(size=30) * .2
plt.plot(x, y, 'ro-', label='A sine wave')
plt.plot(x, z, 'b-', label='Noisy sine')
plt.legend(loc = 'lower right')
plt.xlabel("X axis")
plt.ylabel("Y axis")
# -
# If that last cell complained about the `%matplotlib` line, you need to update IPython to v1.0, and restart the notebook. See the [installation page](https://github.com/cs109/content/wiki/Installing-Python)
# ## Hello Numpy
#
# The Numpy array processing library is the basis of nearly all numerical computing in Python. Here's a 30 second crash course. For more details, consult Chapter 4 of Python for Data Analysis, or the [Numpy User's Guide](http://docs.scipy.org/doc/numpy-dev/user/index.html)
# +
print ("Make a 3 row x 4 column array of random numbers")
x = np.random.random((3, 4))
print (x)
print
print ("Add 1 to every element")
x = x + 1
print (x)
print
print ("Get the element at row 1, column 2")
print (x[1, 2])
print
# The colon syntax is called "slicing" the array.
print ("Get the first row")
print (x[0, :])
print
print ("Get every 2nd column of the first row")
print (x[0, ::2])
print
# -
# Print the maximum, minimum, and mean of the array. This does **not** require writing a loop. In the code cell below, type `x.m<TAB>`, to find built-in operations for common array statistics like this
#your code here
# Call the `x.max` function again, but use the `axis` keyword to print the maximum of each row in x.
#your code here
# Here's a way to quickly simulate 500 coin "fair" coin tosses (where the probabily of getting Heads is 50%, or 0.5)
x = np.random.binomial(500, .5)
print "number of heads:", x
# Repeat this simulation 500 times, and use the [plt.hist() function](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.hist) to plot a histogram of the number of Heads (1s) in each simulation
#your code here
# ## The Monty Hall Problem
#
#
# Here's a fun and perhaps surprising statistical riddle, and a good way to get some practice writing python functions
#
# In a gameshow, contestants try to guess which of 3 closed doors contain a cash prize (goats are behind the other two doors). Of course, the odds of choosing the correct door are 1 in 3. As a twist, the host of the show occasionally opens a door after a contestant makes his or her choice. This door is always one of the two the contestant did not pick, and is also always one of the goat doors (note that it is always possible to do this, since there are two goat doors). At this point, the contestant has the option of keeping his or her original choice, or swtiching to the other unopened door. The question is: is there any benefit to switching doors? The answer surprises many people who haven't heard the question before.
#
# We can answer the problem by running simulations in Python. We'll do it in several parts.
#
# First, write a function called `simulate_prizedoor`. This function will simulate the location of the prize in many games -- see the detailed specification below:
"""
Function
--------
simulate_prizedoor
Generate a random array of 0s, 1s, and 2s, representing
hiding a prize between door 0, door 1, and door 2
Parameters
----------
nsim : int
The number of simulations to run
Returns
-------
sims : array
Random array of 0s, 1s, and 2s
Example
-------
>>> print simulate_prizedoor(3)
array([0, 0, 2])
"""
def simulate_prizedoor(nsim):
#compute here
return answer
#your code here
# Next, write a function that simulates the contestant's guesses for `nsim` simulations. Call this function `simulate_guess`. The specs:
"""
Function
--------
simulate_guess
Return any strategy for guessing which door a prize is behind. This
could be a random strategy, one that always guesses 2, whatever.
Parameters
----------
nsim : int
The number of simulations to generate guesses for
Returns
-------
guesses : array
An array of guesses. Each guess is a 0, 1, or 2
Example
-------
>>> print simulate_guess(5)
array([0, 0, 0, 0, 0])
"""
#your code here
# Next, write a function, `goat_door`, to simulate randomly revealing one of the goat doors that a contestant didn't pick.
"""
Function
--------
goat_door
Simulate the opening of a "goat door" that doesn't contain the prize,
and is different from the contestants guess
Parameters
----------
prizedoors : array
The door that the prize is behind in each simulation
guesses : array
THe door that the contestant guessed in each simulation
Returns
-------
goats : array
The goat door that is opened for each simulation. Each item is 0, 1, or 2, and is different
from both prizedoors and guesses
Examples
--------
>>> print goat_door(np.array([0, 1, 2]), np.array([1, 1, 1]))
>>> array([2, 2, 0])
"""
#your code here
# Write a function, `switch_guess`, that represents the strategy of always switching a guess after the goat door is opened.
"""
Function
--------
switch_guess
The strategy that always switches a guess after the goat door is opened
Parameters
----------
guesses : array
Array of original guesses, for each simulation
goatdoors : array
Array of revealed goat doors for each simulation
Returns
-------
The new door after switching. Should be different from both guesses and goatdoors
Examples
--------
>>> print switch_guess(np.array([0, 1, 2]), np.array([1, 2, 1]))
>>> array([2, 0, 0])
"""
#your code here
# Last function: write a `win_percentage` function that takes an array of `guesses` and `prizedoors`, and returns the percent of correct guesses
"""
Function
--------
win_percentage
Calculate the percent of times that a simulation of guesses is correct
Parameters
-----------
guesses : array
Guesses for each simulation
prizedoors : array
Location of prize for each simulation
Returns
--------
percentage : number between 0 and 100
The win percentage
Examples
---------
>>> print win_percentage(np.array([0, 1, 2]), np.array([0, 0, 0]))
33.333
"""
#your code here
# Now, put it together. Simulate 10000 games where contestant keeps his original guess, and 10000 games where the contestant switches his door after a goat door is revealed. Compute the percentage of time the contestant wins under either strategy. Is one strategy better than the other?
#your code here
# Many people find this answer counter-intuitive (famously, PhD mathematicians have incorrectly claimed the result must be wrong. Clearly, none of them knew Python).
#
# One of the best ways to build intuition about why opening a Goat door affects the odds is to re-run the experiment with 100 doors and one prize. If the game show host opens 98 goat doors after you make your initial selection, would you want to keep your first pick or switch? Can you generalize your simulation code to handle the case of `n` doors?
| Assignments/HW0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: ''
# name: pysparkkernel
# ---
# ## Analysis of Yelp Business Intelligence Data
# We will analyze a subset of Yelp's business, reviews and user data. This dataset comes to us from Kaggle although we have taken steps to pull this data into a publis s3 bucket: s3://sta9760-yelpdataset/yelp-light/*business.json
# ## Installation and Initial Setup
# Begin by installing the necessary libraries that you may need to conduct your analysis. At the very least, you must install pandas and matplotlib
# %%info
sc.install_pypi_package("pandas==1.0.3")
sc.install_pypi_package("matplotlib==3.2.1")
sc.install_pypi_package("seaborn==0.10.1")
sc.list_packages()
# ## Importing
# Now, import the installed packages from the previous block below.
# +
from pyspark.sql.functions import explode
from pyspark.sql.functions import col, split, desc, ltrim, avg, year, when, array_contains, expr,size, array_except, lit, udf
from pyspark.sql.types import StringType
import seaborn as sns
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
# -
# ## Loading Data
# We are finally ready to load data. Using spark load the data from S3 into a dataframe object that we can manipulate further down in our analysis.
df = spark.read.json('s3://9760yelpdata/yelp_academic_dataset_business.json')
# ## Overview of Data
# Display the number of rows and columns in our dataset.
print(f'Total Columns: {len(df.dtypes)}')
print(f'Total Rows: {df.count():,}')
# Display the DataFrame schema below.
df.printSchema()
#
# Display the first 5 rows with the following columns:
#
# *business_id
#
# *name
#
# *city
#
# *state
#
# *categories
cols = ("business_id","name","city","state","categories")
df.select(*cols).show(5)
# ## Analyzing Categories
# Let's now answer this question: how many unique categories are represented in this dataset?
# ### Calculating the association table:
#df1 = df.select(col("business_id"), explode( split(ltrim(col("categories")), ",") ).alias("categories"))
df1 = df.select(col("business_id"), explode( split(col("categories"), ",") ).alias("categories"))
df1 = df1.withColumn('categories', ltrim(df1.categories))
## Display the first 5 rows of your association table below.
df1.show(5)
#
# ## Total Unique Categories
# Finally, we are ready to answer the question: what is the total number of unique categories available?
#
# Below, implement the code necessary to calculate this figure.
print(f'Unique categories:{df1.select("categories").distinct().count()}')
# ## Top Categories By Business
# Now let's find the top categories in this dataset by rolling up categories.
#
# (i've sorted the output by count for easier reading, and later - plotting)
result_df = df1.select('categories').groupby(df1.categories).count().sort(desc("count"))
result_df.show()
# ## Bar Chart of Top Categories
# With this data available, let us now build a barchart of the top 20 categories.
result_pdf = result_df.toPandas()
plt.clf()
ax = result_pdf.head(20).plot.barh(x='categories', y='count', width=.8)
plt.gca().invert_yaxis()
plt.ylabel('Categories')
plt.xlabel('Count')
plt.title('Top Categories by Business')
ax.get_legend().remove()
plt.box(False)
ax.grid(False)
plt.tight_layout()
# %matplot plt
# ## Do Yelp Reviews Skew Negative?
# Oftentimes, it is said that the only people who write a written review are those who are extremely dissatisfied or extremely satisfied with the service received.
#
# How true is this really? Let's try and answer this question.
# ### Do Yelp Reviews Skew Negative?
# Oftentimes, it is said that the only people who write a written review are those who are extremely dissatisfied or extremely satisfied with the service received.
# How true is this really? Let's try and answer this question.
# ## Loading User Data
# Begin by loading the user data set from S3 and printing schema to determine what data is available.
review = spark.read.json('s3://9760yelpdata/yelp_academic_dataset_review.json')
# Total columns
print(f'Total Columns: {len(review.dtypes)}')
print(f'Total Rows: {review.count():,}')
review.printSchema()
# Let's begin by listing the business_id and stars columns together for the user reviews data.
review.select('business_id','stars').show(5)
# Now, let's aggregate along the stars column to get a resultant dataframe that displays average stars per business as accumulated by users who took the time to submit a written review.
# the condition, in this case, is colmn 'text' is not set to Null
rev = review.where(col("text").isNotNull()).groupby(review.business_id).agg(avg(col("stars")))
rev.show(5)
# same thing, for all reviews , not just those that have text
rev_all = review.groupby(review.business_id).agg(avg(col("stars")).alias("actual_stars"))
# ### Now the fun part - let's join our two dataframes (reviews and business data) by business_id
#
joined = df.join(rev, on=['business_id'])
joined_all = df.join(rev_all, on=['business_id'])
# Let's see a few of these:
j = joined.select("avg(stars)","stars","name","city","state").sort("""avg(stars)""",ascending=False)
j.show(5)
j_all = joined_all.select("stars","actual_stars","name","city","state").sort("actual_stars",ascending=False)
#
# Compute a new dataframe that calculates what we will call the skew (for lack of a better word) between the avg stars accumulated from written reviews and the actual star rating of a business (ie: the average of stars given by reviewers who wrote an actual review and reviewers who just provided a star rating).
#
# The formula you can use is something like:
#
# (row['avg(stars)'] - row['stars']) / row['stars']
new_df = j.withColumn("skew_score", ((j["""avg(stars)"""]-j_all["stars"])/j["stars"])).toPandas()
# And finally, graph it!
plt.clf()
plt.figure(figsize=(20,10))
sns.distplot(new_df["skew_score"], hist_kws=dict(alpha=0.2))
plt.title('Top Categories by Business', fontsize=20)
plt.box(False)
# %matplot plt
# So, do Yelp (written) Reviews skew negative? Does this analysis actually prove anything? Expound on implications / interpretations of this graph.
# With a positive skewness score, this histogram has a longer tail to the right and so it is positively skewed, not negatively. The score is <1 , which is usually not considered a "heavy skew"
# ### last part! i'll try to answer the question :
# "how accurate or close are the ratings of an "elite" user (check Users table schema) vs the actual business rating."
# Load user data set, and join with review on user_id
user = spark.read.json('s3://9760yelpdata/yelp_academic_dataset_user.json')
user_rev = review.join(user, on='user_id')
# Add a column where i've taken the year alone from the date of each review:
user_rev = user_rev.withColumn("year",year("date").cast('String'))
# this UDF will check whether a var is contained within another. Using this, we can later check whether a review was given by an 'elite' user , or not. If the year the review was given at is contained within the 'elite' column, the review will be set to "elite review"
# +
def contains(x,y):
z = len(set(x) - set(y))
if z == 0:
return True
else:
return False
contains_udf = udf(contains)
# -
years = user_rev.select('year')
elites = user_rev.select('elite')
user_rev = user_rev.withColumn("elite_rev", contains_udf(*years,*elites))
user_rev = user_rev.join(rev_all.select("business_id","actual_stars"),on=['business_id'])
# Cast to pandas dataframe in order to later plot :
user_rev_pd = user_rev.select('business_id','stars','actual_stars','elite_rev','year').toPandas()
# i'll create a 'diff' column, to represent the difference between stars given in a review and the actual star-count of a business. Later, i will compare the average and variance of this "accuracy" between elite and non-elite reviews
user_rev_pd['diff'] = user_rev_pd['actual_stars']-user_rev_pd['stars']
user_rev_pd['elite_rev'] = np.where(user_rev_pd['elite_rev']=='true','Elite Review','Non-Elite Review')
# # lets graph it!
# i'll create a boxplot, with the median represented as a line in the middle of a box representing the IQR (interquartile range) - 50% of the data
plt.clf()
ax = sns.boxplot(x='elite_rev', y="diff", data=user_rev_pd)
plt.title('mean difference between elite/non reviews and the actual review', fontsize=20)
plt.box(False)
# %matplot plt
# we can see that Elite reviews are not only closer to the actualy score, but have a lower variance too, as in ~50% of scores are closer to eachother and to the actual average than non-elite reviews.
# After doing this analysis, i would definately count on elite reviews more than i have so far
| Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import sqlite3 package
import sqlite3
# create a connection to a database : create the database if not available
connection = sqlite3.connect("classroomDB.db")
connection.close()
connection = sqlite3.connect("classroomDB.db")
cursor = connection.cursor()
create_table = """
CREATE Table classroom (
student_id INTEGER PRIMARY KEY,
name VARCHAR(20),
gender CHAR(1),
physics_marks INTEGER,
chemistry_marks INTEGER,
mathematics_marks INTEGER
);
"""
drop_table = """
DROP TABLE classroom;
"""
cursor.execute(create_table)
# cursor.execute(drop_table)
#cursor.execute("select * from classroom")
connection.commit()
connection.close()
# +
# sample data
classroom_data = [( 1, "Raj","M", 70, 84, 92),
( 2, "Poonam","F", 87, 69, 93),
( 3, "Nik","M", 65, 83, 90),
( 4, "Rahul","F", 83, 76, 89)]
# open connection
connection = sqlite3.connect("classroomDB.db")
# open cursor
cursor = connection.cursor()
# insert each student record
for student in classroom_data:
# formatted query string
insert_statement = """INSERT INTO classroom
(student_id, name, gender, physics_marks, chemistry_marks, mathematics_marks)
VALUES
({0}, "{1}", "{2}", {3}, {4}, {5});""".format(student[0], student[1], student[2],
student[3],student[4], student[5])
# execute insert query
cursor.execute(insert_statement)
# commit the changes
connection.commit()
# close the connection
connection.close()
# -
# open connection
connection = sqlite3.connect("classroomDB.db")
# open cursor
cursor = connection.cursor()
# query
query = "SELECT * FROM classroom"
# execute query
cursor.execute(query)
# fetch results
result = cursor.fetchall()
# print results
for row in result:
print(row)
# close connection
connection.close()
# # My sql database
# ## install package
# !conda install -y -q pymysql
#import package
import pymysql
# ### connect to database
# +
# Connection details
cnx= {'host': 'titanic.cr4famdwl9cg.us-west-2.rds.amazonaws.com',
'username': 'username',
'password': 'password',
'db': 'titanicDB'}
# Connect to the database
connection = pymysql.connect(cnx['host'],cnx['username'],cnx['password'],cnx['db'] )
# close database
connection.close()
# -
# ### create table
# open connection
connection = pymysql.connect(cnx['host'],cnx['username'],cnx['password'],cnx['db'] )
# open cursor
cursor = connection.cursor()
# query for creating table
create_table = """
CREATE TABLE classroom (
student_id INTEGER PRIMARY KEY,
name VARCHAR(20),
gender CHAR(1),
physics_marks INTEGER,
chemistry_marks INTEGER,
mathematics_marks INTEGER
);"""
# execute query
cursor.execute(create_table)
# commit changes
connection.commit()
# close connection
connection.close()
# ### insert data
# +
# sample data
classroom_data = [( 1, "Raj","M", 70, 84, 92),
( 2, "Poonam","F", 87, 69, 93),
( 3, "Nik","M", 65, 83, 90),
( 4, "Rahul","F", 83, 76, 89)]
# open connection
connection = pymysql.connect(cnx['host'],cnx['username'],cnx['password'],cnx['db'] )
# open cursor
cursor = connection.cursor()
# insert each student record
for student in classroom_data:
# formatted query string
insert_statement = """INSERT INTO classroom
(student_id, name, gender, physics_marks, chemistry_marks, mathematics_marks)
VALUES
({0}, "{1}", "{2}", {3}, {4}, {5});""".format(student[0], student[1], student[2],
student[3],student[4], student[5])
# execute insert query
cursor.execute(insert_statement)
# commit the changes
connection.commit()
# close the connection
connection.close()
# -
# ### Extract data
# open connection
connection = pymysql.connect(cnx['host'],cnx['username'],cnx['password'],cnx['db'] )
# open cursor
cursor = connection.cursor()
# query
query = "SELECT * FROM classroom"
# execute query
cursor.execute(query)
# fetch results
result = cursor.fetchall()
# print results
for row in result:
print(row)
# close connection
connection.close()
# ## MS SQL Database
# !conda install -y -q pymssql
import pymssql
# !conda info pymssql
| notebooks/Extracting data - databases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Science Plan
#
# Objective - test hypothesis that weather events can affect the bottom currents that control the bending and rise heights of Hydrothermal plumes
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from mpl_toolkits import mplot3d
import pandas as pd
from datetime import datetime
# ## Step One - Gather Data
#
# Plume Bending Data
df_BD2010 = pd.read_csv('BendData2010Oct.txt', sep=" ", header=None, engine='python')
df_BD2010.columns = ["Direction", "Angle from Vertical", "Julian Day"]
df_BD2010
df_BD2011 = pd.read_csv('BendData2011OctDec.txt', sep=" ", header=None, engine='python')
df_BD2011.columns = ["Direction", "Angle from Vertical", "Julian Day"]
df_BD2011
# Centerline Vertical Flow Rate
df_CVFR2010 = pd.read_csv('CenterlineVerticalFlowRate2010.csv', sep=",", header=None, engine='python', usecols = [6,7,8])
df_CVFR2010.columns = ["Date", "Time", "Flow Rate"]
df_CVFR2010['Date Time'] = df_CVFR2010['Date'] + ' ' + df_CVFR2010['Time']
df_CVFR2010["Date Time"] = pd.to_datetime(df_CVFR2010["Date Time"])
df_CVFR2010 = df_CVFR2010.drop(["Date", "Time"], axis=1)
df_CVFR2010 = df_CVFR2010.set_index('Date Time')
df_CVFR2010
# Ras Data
df_RAS2010 = pd.read_csv('RasData2010.csv', sep=",", header=None, engine='python', usecols = [6,7,8])
df_RAS2010.columns = ["Date", "Time", "Temperature"]
df_RAS2010['Date Time'] = df_RAS2010['Date'] + ' ' + df_RAS2010['Time']
df_RAS2010["Date Time"] = pd.to_datetime(df_RAS2010["Date Time"])
df_RAS2010 = df_RAS2010.drop(["Date", "Time"], axis=1)
df_RAS2010 = df_RAS2010.set_index('Date Time')
df_RAS2010
# Weather Data
df_WD2010C = pd.read_csv('WeatherC2010.csv', sep=",", header=None, engine='python', usecols = [6,7,8,9,10,11,12,13,14,15])
df_WD2010C.columns = ["Date", "Time", "Julian Day", "Wave Height", "Wind Direction", "Wind Speed", "Wind Gust Speed", "Atmospheric Pressure","Air Temperature"]
df_WD2010C['Date Time'] = df_WD2010C['Date'] + ' ' + df_WD2010C['Time']
df_WD2010C["Date Time"] = pd.to_datetime(df_WD2010C["Date Time"])
df_WD2010C = df_WD2010C.drop(["Date", "Time"], axis=1)
df_WD2010C = df_WD2010C.set_index('Date Time')
df_WD2010C
df_WD2011C = pd.read_csv('WeatherC2011.csv', sep=",", header=None, engine='python', usecols = [6,7,8,9,10,11,12,13,14,15])
df_WD2011C.columns = ["Date", "Time", "Julian Day", "Wave Height", "Wind Direction", "Wind Speed", "Wind Gust Speed", "Atmospheric Pressure","Air Temperature"]
df_WD2011C['Date Time'] = df_WD2011C['Date'] + ' ' + df_WD2011C['Time']
df_WD2011C["Date Time"] = pd.to_datetime(df_WD2011C["Date Time"])
df_WD2011C = df_WD2011C.drop(["Date", "Time"], axis=1)
df_WD2011C = df_WD2011C.set_index('Date Time')
df_WD2011C
df_WD2010T = pd.read_csv('WeatherT2010.csv', sep=",", header=None, engine='python', usecols = [6,7,8,9,10,11,12,13,14,15])
df_WD2010T.columns = ["Date", "Time", "Julian Day", "Wave Height", "Wind Direction", "Wind Speed", "Wind Gust Speed", "Atmospheric Pressure","Air Temperature"]
df_WD2010T['Date Time'] = df_WD2010T['Date'] + ' ' + df_WD2010T['Time']
df_WD2010T["Date Time"] = pd.to_datetime(df_WD2010T["Date Time"])
df_WD2010T = df_WD2010T.drop(["Date", "Time"], axis=1)
df_WD2010T = df_WD2010T.set_index('Date Time')
df_WD2010T
df_WD2011T = pd.read_csv('WeatherT2011.csv', sep=",", header=None, engine='python', usecols = [6,7,8,9,10,11,12,13,14,15])
df_WD2011T.columns = ["Date", "Time", "Julian Day", "Wave Height", "Wind Direction", "Wind Speed", "Wind Gust Speed", "Atmospheric Pressure","Air Temperature"]
df_WD2011T['Date Time'] = df_WD2011T['Date'] + ' ' + df_WD2011T['Time']
df_WD2011T["Date Time"] = pd.to_datetime(df_WD2011T["Date Time"])
df_WD2011T = df_WD2011T.drop(["Date", "Time"], axis=1)
df_WD2011T = df_WD2011T.set_index('Date Time')
df_WD2011T
# ## Step Two - Data Plots
# Plume Bending Data 2010
DirPlot2010 = df_BD2010.plot(kind='line',x='Julian Day',y="Direction",color='black')
DirPlot2010.set_title("Direction of Bending")
DirPlot2010.set_xlabel('Julain Day')
DirPlot2010.set_ylabel('Direction (Degrees from North)')
AFVPlot2010 = df_BD2010.plot(kind='line',x='Julian Day',y="Angle from Vertical",color='black')
AFVPlot2010.set_title("Bending Angle from Vertical")
AFVPlot2010.set_xlabel('Julain Day')
AFVPlot2010.set_ylabel('Angle from Vertical (Degress)')
DirAFVPlot2010 = df_BD2010.plot(kind='line',x='Julian Day',y=["Direction", "Angle from Vertical"],color=['blue','red'])
DirAFVPlot2010.set_title("Plume Bending Data 2010")
DirAFVPlot2010.set_xlabel('Julain Day')
DirAFVPlot2010.set_ylabel('Degress')
# Plume Bending Data 2011
DirPlot2011 = df_BD2011.plot(kind='line',x='Julian Day',y="Direction",color='black')
DirPlot2011.set_title("Direction of Bending")
DirPlot2011.set_xlabel('Julain Day')
DirPlot2011.set_ylabel('Direction (Degrees from North)')
AFVPlot2011 = df_BD2011.plot(kind='line',x='Julian Day',y="Angle from Vertical",color='black')
AFVPlot2011.set_title("Bending Angle from Vertical")
AFVPlot2011.set_xlabel('Julain Day')
AFVPlot2011.set_ylabel('Angle from Vertical (Degress)')
DirAFVPlot2011 = df_BD2011.plot(kind='line',x='Julian Day',y=["Direction", "Angle from Vertical"],color=['blue','red'])
DirAFVPlot2011.set_title("Plume Bending Data 2010")
DirAFVPlot2011.set_xlabel('Julain Day')
DirAFVPlot2011.set_ylabel('Degress')
# Centerline Vertical Flow Rate
CVFRPlot = df_CVFR2010.plot(kind='line',color='black')
CVFRPlot.set_title("Centerline Vertical Flow Rate")
CVFRPlot.set_xlabel('Date Time')
CVFRPlot.set_ylabel('Flow Rate (m/s)')
# Ras Data
RASPlot = df_RAS2010.plot(kind='line',color='black')
RASPlot.set_title("RAS Temperature")
RASPlot.set_xlabel('Date Time')
RASPlot.set_ylabel('Temperature (°C)')
# Weather Data 2010 C46036
WH2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Wave Height')
WH2010CPlot.set_title("Wave Height 2010 at C46036")
WH2010CPlot.set_xlabel('Date Time')
WH2010CPlot.set_ylabel('Wave Height')
WDir2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Wind Direction')
WDir2010CPlot.set_title("Wind Direction 2010 at C46036")
WDir2010CPlot.set_xlabel('Date Time')
WDir2010CPlot.set_ylabel('Wind Direction')
WS2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Wind Speed')
WS2010CPlot.set_title("Wind Speed 2010 at C46036")
WS2010CPlot.set_xlabel('Date Time')
WS2010CPlot.set_ylabel('Wind Speed (m/s)')
WGS2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Wind Gust Speed')
WGS2010CPlot.set_title("Wind Gust Speed 2010 at C46036")
WGS2010CPlot.set_xlabel('Date Time')
WGS2010CPlot.set_ylabel('Wind Gust Speed (m/s)')
AP2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Atmospheric Pressure')
AP2010CPlot.set_title("Atmospheric Pressure 2010 at C46036")
AP2010CPlot.set_xlabel('Date Time')
AP2010CPlot.set_ylabel('Atmospheric Pressure')
AT2010CPlot = df_WD2010C.plot(kind='line',color='black', y='Air Temperature')
AT2010CPlot.set_title("Air Temperature 2010 at C46036")
AT2010CPlot.set_xlabel('Date Time')
AT2010CPlot.set_ylabel('Air Temperature')
# Weather Data 2011 C46036
WH2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Wave Height')
WH2011CPlot.set_title("Wave Height 2011 at C46036")
WH2011CPlot.set_xlabel('Date Time')
WH2011CPlot.set_ylabel('Wave Height')
WDir2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Wind Direction')
WDir2011CPlot.set_title("Wind Direction 2011 at C46036")
WDir2011CPlot.set_xlabel('Date Time')
WDir2011CPlot.set_ylabel('Wind Direction')
WS2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Wind Speed')
WS2011CPlot.set_title("Wind Speed 2011 at C46036")
WS2011CPlot.set_xlabel('Date Time')
WS2011CPlot.set_ylabel('Wind Speed (m/s)')
WGS2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Wind Gust Speed')
WGS2011CPlot.set_title("Wind Gust Speed 2011 at C46036")
WGS2011CPlot.set_xlabel('Date Time')
WGS2011CPlot.set_ylabel('Wind Gust Speed (m/s)')
AP2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Atmospheric Pressure')
AP2011CPlot.set_title("Atmospheric Pressure 2011 at C46036")
AP2011CPlot.set_xlabel('Date Time')
AP2011CPlot.set_ylabel('Atmospheric Pressure')
AT2011CPlot = df_WD2011C.plot(kind='line',color='black', y='Air Temperature')
AT2011CPlot.set_title("Air Temperature 2011 at C46036")
AT2011CPlot.set_xlabel('Date Time')
AT2011CPlot.set_ylabel('Air Temperature')
# Weather Data 2010 Tillamook
WDir2010TPlot = df_WD2010T.plot(kind='line',color='black', y='Wind Direction')
WDir2010TPlot.set_title("Wind Direction 2010 at Tillamook")
WDir2010TPlot.set_xlabel('Date Time')
WDir2010TPlot.set_ylabel('Wind Direction')
WS2010TPlot = df_WD2010T.plot(kind='line',color='black', y='Wind Speed')
WS2010TPlot.set_title("Wind Speed 2010 at Tillamook")
WS2010TPlot.set_xlabel('Date Time')
WS2010TPlot.set_ylabel('Wind Speed (m/s)')
WGS2010TPlot = df_WD2010T.plot(kind='line',color='black', y='Wind Gust Speed')
WGS2010TPlot.set_title("Wind Gust Speed 2010 at Tillamook")
WGS2010TPlot.set_xlabel('Date Time')
WGS2010TPlot.set_ylabel('Wind Gust Speed (m/s)')
AT2010TPlot = df_WD2010T.plot(kind='line',color='black', y='Air Temperature')
AT2010TPlot.set_title("Air Temperature 2010 at Tillamook")
AT2010TPlot.set_xlabel('Date Time')
AT2010TPlot.set_ylabel('Air Temperature')
# Weather Data 2011 Tillamook
WDir2011TPlot = df_WD2011T.plot(kind='line',color='black', y='Wind Direction')
WDir2011TPlot.set_title("Wind Direction 2011 at Tillamook")
WDir2011TPlot.set_xlabel('Date Time')
WDir2011TPlot.set_ylabel('Wind Direction')
WS2011TPlot = df_WD2011T.plot(kind='line',color='black', y='Wind Speed')
WS2011TPlot.set_title("Wind Speed 2011 at Tillamook")
WS2011TPlot.set_xlabel('Date Time')
WS2011TPlot.set_ylabel('Wind Speed (m/s)')
WGS2011TPlot = df_WD2011T.plot(kind='line',color='black', y='Wind Gust Speed')
WGS2011TPlot.set_title("Wind Gust Speed 2011 at Tillamook")
WGS2011TPlot.set_xlabel('Date Time')
WGS2011TPlot.set_ylabel('Wind Gust Speed (m/s)')
AT2011TPlot = df_WD2011T.plot(kind='line',color='black', y='Air Temperature')
AT2011TPlot.set_title("Air Temperature 2011 at Tillamook")
AT2011TPlot.set_xlabel('Date Time')
AT2011TPlot.set_ylabel('Air Temperature')
| sacker/SackerSciencePlan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## **問13 if ~ elif ~ else 文の使い方を知る**
# 続いて、if ~ elif ~ else文について学びましょう
# <br>
# <br>
# 問題12から少し問題を変更してプログラムを書いてみましょう。
# <br>
# **0から100の間でランダムに数字を発生させ、その数字を`print()`関数で表示させ、<br>
# その値が80以上なら"クラスA"、60以上なら“クラスB”、30以上なら“クラスC”、それ以外は“クラスD”とします。**
# <br>
# それではプログラムを書いていきましょう。
# <br>
# <br>
# スクリプト名:training13.py
# +
# ランダムに数字を発生させ、その数字が80以上ならクラスA, 60以上80未満ならクラスB, 30以上60未満ならクラスC,30未満ならクラスD
import random
point = random.randint(0,100)
print("ランダムに生成した数字は{0}".format(point))
# 判定
if point >= 80 :
result = "クラスA"
elif point >= 60:
result = "クラスB"
elif point >= 30:
result = "クラスC"
else :
result = "クラスD"
print(result)
# -
# <br>
# **if elif else文について**
# <br>
# if elif else文は、判定条件が3つ以上の時に使用されます。
| source/training13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <a href="https://www.bigdatauniversity.com"><img src="https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width="400" align="center"></a>
#
# <h1 align="center"><font size="5">Classification with Python</font></h1>
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In this notebook we try to practice all the classification algorithms that we learned in this course.
#
# We load a dataset using Pandas library, and apply the following algorithms, and find the best one for this specific dataset by accuracy evaluation methods.
#
# Lets first load required libraries:
# + button=false new_sheet=false run_control={"read_only": false}
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
# %matplotlib inline
import seaborn as sns
from sklearn.model_selection import train_test_split
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### About dataset
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# This dataset is about the performance of basketball teams. The __cbb.csv__ data set includes performance data about five seasons of 354 basketball teams. It includes following fields:
#
# | Field | Description |
# |----------------|---------------------------------------------------------------------------------------|
# |TEAM | The Division I college basketball school|
# |CONF| The Athletic Conference in which the school participates in (A10 = Atlantic 10, ACC = Atlantic Coast Conference, AE = America East, Amer = American, ASun = ASUN, B10 = Big Ten, B12 = Big 12, BE = Big East, BSky = Big Sky, BSth = Big South, BW = Big West, CAA = Colonial Athletic Association, CUSA = Conference USA, Horz = Horizon League, Ivy = Ivy League, MAAC = Metro Atlantic Athletic Conference, MAC = Mid-American Conference, MEAC = Mid-Eastern Athletic Conference, MVC = Missouri Valley Conference, MWC = Mountain West, NEC = Northeast Conference, OVC = Ohio Valley Conference, P12 = Pac-12, Pat = Patriot League, SB = Sun Belt, SC = Southern Conference, SEC = South Eastern Conference, Slnd = Southland Conference, Sum = Summit League, SWAC = Southwestern Athletic Conference, WAC = Western Athletic Conference, WCC = West Coast Conference)|
# |G| Number of games played|
# |W| Number of games won|
# |ADJOE| Adjusted Offensive Efficiency (An estimate of the offensive efficiency (points scored per 100 possessions) a team would have against the average Division I defense)|
# |ADJDE| Adjusted Defensive Efficiency (An estimate of the defensive efficiency (points allowed per 100 possessions) a team would have against the average Division I offense)|
# |BARTHAG| Power Rating (Chance of beating an average Division I team)|
# |EFG_O| Effective Field Goal Percentage Shot|
# |EFG_D| Effective Field Goal Percentage Allowed|
# |TOR| Turnover Percentage Allowed (Turnover Rate)|
# |TORD| Turnover Percentage Committed (Steal Rate)|
# |ORB| Offensive Rebound Percentage|
# |DRB| Defensive Rebound Percentage|
# |FTR| Free Throw Rate (How often the given team shoots Free Throws)|
# |FTRD| Free Throw Rate Allowed|
# |2P_O| Two-Point Shooting Percentage|
# |2P_D| Two-Point Shooting Percentage Allowed|
# |3P_O| Three-Point Shooting Percentage|
# |3P_D| Three-Point Shooting Percentage Allowed|
# |ADJ_T| Adjusted Tempo (An estimate of the tempo (possessions per 40 minutes) a team would have against the team that wants to play at an average Division I tempo)|
# |WAB| Wins Above Bubble (The bubble refers to the cut off between making the NCAA March Madness Tournament and not making it)|
# |POSTSEASON| Round where the given team was eliminated or where their season ended (R68 = First Four, R64 = Round of 64, R32 = Round of 32, S16 = Sweet Sixteen, E8 = Elite Eight, F4 = Final Four, 2ND = Runner-up, Champion = Winner of the NCAA March Madness Tournament for that given year)|
# |SEED| Seed in the NCAA March Madness Tournament|
# |YEAR| Season
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Data From CSV File
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's load the dataset [NB Need to provide link to csv file]
# + button=false new_sheet=false run_control={"read_only": false}
df = pd.read_csv('https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0120ENv3/Dataset/ML0101EN_EDX_skill_up/cbb.csv')
df.head()
# -
df.shape
# ## Add Column
# Next we'll add a column that will contain "true" if the wins above bubble are over 7 and "false" if not. We'll call this column Win Index or "windex" for short.
df['windex'] = np.where(df.WAB > 7, 'True', 'False')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Data visualization and pre-processing
#
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Next we'll filter the data set to the teams that made the Sweet Sixteen, the Elite Eight, and the Final Four in the post season. We'll also create a new dataframe that will hold the values with the new column.
# -
df1 = df.loc[df['POSTSEASON'].str.contains('F4|S16|E8', na=False)]
df1.head()
# + button=false new_sheet=false run_control={"read_only": false}
df1['POSTSEASON'].value_counts()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# 32 teams made it into the Sweet Sixteen, 16 into the Elite Eight, and 8 made it into the Final Four over 5 seasons.
#
# -
# Lets plot some columns to underestand data better:
# notice: installing seaborn might takes a few minutes
# !conda install -c anaconda seaborn -y
# +
bins = np.linspace(df1.BARTHAG.min(), df1.BARTHAG.max(), 10)
g = sns.FacetGrid(df1, col="windex", hue="POSTSEASON", palette="Set1", col_wrap=6)
g.map(plt.hist, 'BARTHAG', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + button=false new_sheet=false run_control={"read_only": false}
bins = np.linspace(df1.ADJOE.min(), df1.ADJOE.max(), 10)
g = sns.FacetGrid(df1, col="windex", hue="POSTSEASON", palette="Set1", col_wrap=2)
g.map(plt.hist, 'ADJOE', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Pre-processing: Feature selection/extraction
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Lets look at how Adjusted Defense Efficiency plots
# + button=false new_sheet=false run_control={"read_only": false}
bins = np.linspace(df1.ADJDE.min(), df1.ADJDE.max(), 10)
g = sns.FacetGrid(df1, col="windex", hue="POSTSEASON", palette="Set1", col_wrap=2)
g.map(plt.hist, 'ADJDE', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We see that this data point doesn't impact the ability of a team to get into the Final Four.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Convert Categorical features to numerical values
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets look at the postseason:
# + button=false new_sheet=false run_control={"read_only": false}
df1.groupby(['windex'])['POSTSEASON'].value_counts(normalize=True)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# 13% of teams with 6 or less wins above bubble make it into the final four while 17% of teams with 7 or more do.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets convert wins above bubble (winindex) under 7 to 0 and over 7 to 1:
#
# + button=false new_sheet=false run_control={"read_only": false}
df1['windex'].replace(to_replace=['False','True'], value=[0,1],inplace=True)
df1.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Feature selection
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets defind feature sets, X:
# + button=false new_sheet=false run_control={"read_only": false}
X = df1[['G', 'W', 'ADJOE', 'ADJDE', 'BARTHAG', 'EFG_O', 'EFG_D',
'TOR', 'TORD', 'ORB', 'DRB', 'FTR', 'FTRD', '2P_O', '2P_D', '3P_O',
'3P_D', 'ADJ_T', 'WAB', 'SEED', 'windex']]
X[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# What are our lables? Round where the given team was eliminated or where their season ended (R68 = First Four, R64 = Round of 64, R32 = Round of 32, S16 = Sweet Sixteen, E8 = Elite Eight, F4 = Final Four, 2ND = Runner-up, Champion = Winner of the NCAA March Madness Tournament for that given year)|
# + button=false new_sheet=false run_control={"read_only": false}
y = df1['POSTSEASON'].values
y[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Normalize Data
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Data Standardization give data zero mean and unit variance (technically should be done after train test split )
# + button=false new_sheet=false run_control={"read_only": false}
X = preprocessing.StandardScaler().fit(X).transform(X)
X[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Training and Validation
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Split the data into Training and Validation data.
# + button=false new_sheet=false run_control={"read_only": false}
# We split the X into train and test to find the best k
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=4)
print ('Train set:', X_train.shape, y_train.shape)
print ('Validation set:', X_val.shape, y_val.shape)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Classification
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now, it is your turn, use the training set to build an accurate model. Then use the validation set to report the accuracy of the model
# You should use the following algorithm:
# - K Nearest Neighbor(KNN)
# - Decision Tree
# - Support Vector Machine
# - Logistic Regression
#
#
# -
# # K Nearest Neighbor(KNN)
#
# <b>Question 1 </b> Build a KNN model using a value of k equals five, find the accuracy on the validation data (X_val and y_val)
# +
from sklearn.metrics import accuracy_score
from sklearn.neighbors import KNeighborsClassifier
# Set k to 5
k = 5
neighbors = KNeighborsClassifier(n_neighbors = k).fit(X_train, y_train)
yhat = neighbors.predict(X_val)
print("Train set Accuracy: ", accuracy_score(y_train, neighbors.predict(X_train)))
print("Test set Accuracy: ", accuracy_score(y_val, yhat))
# -
# <b>Question 2</b> Determine and print the accuracy for the first 15 values of k the on the validation data:
# +
Ks = 15
mean_acc = np.zeros((Ks-1))
std_acc = np.zeros((Ks-1))
for n in range(1, Ks):
#Train Model and Predict
neigh = KNeighborsClassifier(n_neighbors = n).fit(X_train, y_train)
yhat = neigh.predict(X_val)
mean_acc[n-1] = accuracy_score(y_val, yhat)
std_acc[n-1] = np.std(yhat==y_val)/np.sqrt(yhat.shape[0])
print(mean_acc)
# -
# Plot model accuracy
# +
# Plot accuracy
plt.plot(range(1, Ks), mean_acc,'g')
plt.fill_between(range(1, Ks), mean_acc-1*std_acc, mean_acc + 1 * std_acc, alpha=0.10)
plt.legend(('Accuracy ', '+/- 3xstd'))
plt.ylabel('Accuracy ')
plt.xlabel('Number of Neighbors (K)')
plt.tight_layout()
plt.show()
print("Best Accuracy: {}".format(mean_acc.max()))
# -
# # Decision Tree
# The following lines of code fit a <code>DecisionTreeClassifier</code>:
from sklearn.tree import DecisionTreeClassifier
import matplotlib.pyplot as plt
# <b>Question 3</b> Determine the better value for the parameter <code>max_depth</code>
# +
accuracy = []
offset = 1 # Minimum value
for i in range(offset, 11):
decision_tree = DecisionTreeClassifier(criterion="entropy", max_depth=i)
decision_tree.fit(X_train, y_train)
predTree = decision_tree.predict(X_val)
accuracy.append(accuracy_score(y_val, predTree))
print("The most acurate max_depth is {} with an accuracy of {}".format(accuracy.index(max(accuracy))+offset, max(accuracy)))
decision_tree = DecisionTreeClassifier(criterion="entropy", max_depth=accuracy.index(max(accuracy))+offset)
decision_tree.fit(X_train, y_train)
# -
# # Support Vector Machine
# <b>Question 4</b> Train the support vector machine model and determine the accuracy on the validation data for each kernel. Find the kernel (linear, poly, rbf, sigmoid) that provides the best score on the validation data and train a SVM using it.
# +
# SVM
from sklearn import svm, metrics
available_kernels = ['linear', 'rbf', 'sigmoid', 'poly'] # rearranged kernel orders to keep the one with better accuracy
for kernel in available_kernels:
clf = svm.SVC(kernel=kernel)
clf.fit(X_train, y_train)
yhat = clf.predict(X_val)
f1_score = metrics.f1_score(y_val, yhat, average='weighted')
jaccard = metrics.jaccard_similarity_score(y_val, yhat)
print('Kernel: {}\n\tF1 Score: {}\n\tJaccard: {}\n'.format(kernel, f1_score, jaccard))
# -
# # Logistic Regression
# <b>Question 5</b> Train a logistic regression model and determine the accuracy of the validation data (set C=0.01)
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
log_regression = LogisticRegression(C=0.01, solver='liblinear')
log_regression.fit(X_train, y_train)
yhat = log_regression.predict(X_val)
yhat_prob = log_regression.predict_proba(X_val)
print(classification_report(y_val, yhat))
f1_score = metrics.f1_score(y_val, yhat, average='micro')
print('F1 Score: {}'.format(f1_score))
# -
# # Model Evaluation using Test set
from sklearn.metrics import f1_score
# for f1_score please set the average parameter to 'micro'
from sklearn.metrics import log_loss
def jaccard_index(predictions, true):
if (len(predictions) == len(true)):
intersect = 0;
for x,y in zip(predictions, true):
if (x == y):
intersect += 1
return intersect / (len(predictions) + len(true) - intersect)
else:
return -1
# <b>Question 5</b> Calculate the F1 score and Jaccard Similarity score for each model from above. Use the Hyperparameter that performed best on the validation data. **For f1_score please set the average parameter to 'micro'.**
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Test set for evaluation
# + button=false new_sheet=false run_control={"read_only": false}
test_df = pd.read_csv('https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0120ENv3/Dataset/ML0101EN_EDX_skill_up/basketball_train.csv',error_bad_lines=False)
test_df.head()
# -
test_df['windex'] = np.where(test_df.WAB > 7, 'True', 'False')
test_df1 = test_df[test_df['POSTSEASON'].str.contains('F4|S16|E8', na=False)]
test_Feature = test_df1[['G', 'W', 'ADJOE', 'ADJDE', 'BARTHAG', 'EFG_O', 'EFG_D',
'TOR', 'TORD', 'ORB', 'DRB', 'FTR', 'FTRD', '2P_O', '2P_D', '3P_O',
'3P_D', 'ADJ_T', 'WAB', 'SEED', 'windex']]
test_Feature['windex'].replace(to_replace=['False','True'], value=[0,1],inplace=True)
test_X=test_Feature
test_X= preprocessing.StandardScaler().fit(test_X).transform(test_X)
test_X[0:5]
test_y = test_df1['POSTSEASON'].values
test_y[0:5]
# KNN
# +
KNN_pred = neighbors.predict(test_X)
KNN_accuracy = metrics.accuracy_score(test_y, KNN_pred)
KNN_f1_score = metrics.f1_score(test_y, KNN_pred, average='micro')
KNN_jaccard = jaccard_index(test_y, KNN_pred)
print('KNN:\n\tAccuracy Score: {}\n\tF1 Score: {}\n\tJaccard: {}\n'.format(KNN_accuracy, KNN_f1_score, KNN_jaccard))
# -
# Decision Tree
# +
DT_pred = decision_tree.predict(test_X)
DT_accuracy = metrics.accuracy_score(test_y, DT_pred)
DT_f1_score = metrics.f1_score(test_y, DT_pred, average='micro')
DT_jaccard = jaccard_index(test_y, DT_pred)
print('DT:\n\tAccuracy Score: {}\n\tF1 Score: {}\n\tJaccard: {}\n'.format(DT_accuracy, DT_f1_score, DT_jaccard))
# -
# SVM
# +
SVM_pred = clf.predict(test_X)
SVM_accuracy = metrics.accuracy_score(test_y, SVM_pred)
SVM_f1_score = metrics.f1_score(test_y, SVM_pred, average='micro')
SVM_jaccard = jaccard_index(test_y, SVM_pred)
print('SVM:\n\tAccuracy Score: {}\n\tF1 Score: {}\n\tJaccard: {}\n'.format(SVM_accuracy, SVM_f1_score, SVM_jaccard))
# -
# Logistic Regression
# +
LR_pred = log_regression.predict(test_X)
LR_accuracy = metrics.accuracy_score(test_y, LR_pred)
LR_f1_score = metrics.f1_score(test_y, LR_pred, average='micro')
LR_jaccard = jaccard_index(test_y, LR_pred)
print('LR:\n\tAccuracy Score: {}\n\tF1 Score: {}\n\tJaccard: {}\n'.format(LR_accuracy, LR_f1_score, LR_jaccard))
# -
# # Report
# You should be able to report the accuracy of the built model using different evaluation metrics:
# | Algorithm | Accuracy | Jaccard | F1-score | LogLoss |
# |--------------------|----------|----------|-----------|---------|
# | KNN |0.628571 | 0.458333 | 0.628571 | NA |
# | Decision Tree |0.642857 | 0.473684 | 0.642857 | NA |
# | SVM |0.685714 | 0.521739 | 0.685714 | NA |
# | LogisticRegression |0.685714 | 0.521739 | 0.685714 | 1.03719 |
# Something to keep in mind when creating models to predict the results of basketball tournaments or sports in general is that is quite hard due to so many factors influencing the game. Even in sports betting an accuracy of 55% and over is considered good as it indicates profits.
# # My Report
print('| Algorithm | Accuracy | Jaccard | F1-score')
print('|','-'*18, '|', '-'*18, '|', '-'*18, '|', '-'*18)
print('| KNN | {} | {} | {}'.format(KNN_accuracy, KNN_jaccard, KNN_f1_score))
print('| Decision Tree | {} | {} | {}'.format(DT_accuracy, DT_jaccard, DT_f1_score))
print('| SVM | {} | {} | {}'.format(SVM_accuracy, SVM_jaccard, SVM_f1_score))
print('| LogisticRegression | {} | {} | {}'.format(LR_accuracy, LR_jaccard, LR_f1_score))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <h2>Want to learn more?</h2>
#
# Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://cocl.us/ML0101EN_DSX">Watson Studio</a>
#
# <h4>Authors: <a href="https://www.linkedin.com/in/azim-hirjani-691a07179/"><NAME></a> and <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a></h4>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
#
#
# <hr>
#
# <p>Copyright © 2018 <a href="https://cocl.us/DX0108EN_CC">Cognitive Class</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.</p>
| src/Final_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Share Tensor
# ### With Duet Clients
# ## Import libs
# +
import syft as sy
import torch
from sympc.session import Session
from sympc.tensor import MPCTensor
sy.load("sympc")
sy.logger.add(sink="./example.log")
# -
# ## Create the clients
# +
alice = sy.VirtualMachine(name="alice")
bob = sy.VirtualMachine(name="bob")
alice_client = alice.get_client()
bob_client = bob.get_client()
# -
# ## Create a session
# The session is used to send some config information only once between the parties.
# This information can be:
# * the ring size in which we do the computation
# * the precision and base
# * the approximation methods we are using for different functions (TODO)
session = Session(parties=[alice_client, bob_client])
print(session)
# ## Send the session to all the parties
Session.setup_mpc(session)
x = MPCTensor(secret=torch.tensor([1,2,3,4]), session=session)
print(x)
# The values does not make sense for any user, unless they "reconstruct" the original tensor
for val in x.share_ptrs:
print(val.get()) ## Attention! get() destroys the pointer
x_secret = torch.tensor([1,2,3,4])
x = MPCTensor(secret=x_secret, session=session)
print(x.reconstruct()) ## Attention! Also reconstruct, destroys the pointers
# ## Secret is shared by the orchestrator
# - the orchestrator generates shares locally
# - distributes them to the parties
# ### Public Operations
# +
x_secret = torch.tensor([[1,2],[3,4]])
x = MPCTensor(secret=x_secret, session=session)
y = torch.tensor([[5,6],[7,8]])
# -
print("[Priv + Pub] X + Y =\n", (x + y).reconstruct())
print("[Pub + Pub] X + Y =\n", x_secret + y)
print("[Priv - Pub] X - Y =\n", (x - y).reconstruct())
print("[Pub - Pub] X - Y =\n", x_secret - y)
print("[Priv * Pub] X * Y =\n", (x * y).reconstruct())
print("[Pub * Pub] X * Y =\n", x_secret * y)
print("[Priv @ Pub] X * Y =\n", (x @ y).reconstruct())
print("[Pub @ Pub] X * Y =\n", x_secret @ y)
# ### Private Operations
# +
x_secret = torch.tensor([[1,2],[3,4]])
x = MPCTensor(secret=x_secret,session=session)
y_secret = torch.tensor([[5,6],[7,8]])
y = MPCTensor(secret=y_secret, session=session)
# -
print("[Priv + Priv] X + Y =\n", (x + y).reconstruct())
print("[Pub + Pub] X + Y =\n", x_secret + y_secret)
print("[Priv - Priv] X - Y =\n", (x - y).reconstruct())
print("[Pub - Pub] X - Y =\n", x_secret - y_secret)
print("[Priv * Priv] X * Y =\n", (x * y).reconstruct())
print("[Pub * Pub] X * Y =\n", x_secret * y_secret)
print("[Priv @ Pub] X * Y =\n", (x @ y).reconstruct())
print("[Pub @ Pub] X * Y =\n", x_secret @ y_secret)
# ## Private Secret - owned by one party
# - are used more generators to construct PRZS (Pseudo-Random-Zero-Shares)
# - the party that has the secret would add it to their own share
# ### Public Operations
# +
x_secret = alice_client.torch.Tensor([[1,2],[3,4]]) # at Alice
x = MPCTensor(secret=x_secret, shape=(2, 2), session=session)
y = torch.tensor([[5,6],[7,8]], dtype=torch.float)
# -
print("[Priv + Pub] X + Y =\n", (x + y).reconstruct())
print("[Pub + Pub] X + Y =\n", (x_secret + y).get()) # On Alice side
print("[Priv - Pub] X - Y =\n", (x - y).reconstruct())
print("[Pub - Pub] X - Y =\n", (x_secret - y).get()) # On Alice side
print("[Priv * Pub] X * Y =\n", (x * y).reconstruct())
print("[Pub * Pub] X * Y =\n", (x_secret * y).get()) # On Alice side
print("[Priv @ Pub] X * Y =\n", (x @ y).reconstruct())
print("[Pub @ Pub] X * Y =\n", (x_secret @ y).get()) # On Alice side
# ### Private Operations
# +
x_secret = alice_client.torch.Tensor([[1,2],[3,4]]) # at Alice
x = MPCTensor(secret=x_secret, shape=(2, 2), session=session)
y_secret = bob_client.torch.Tensor([[5,6],[7,8]]) # at Bob
y = MPCTensor(secret=y_secret, shape=(2, 2), session=session)
# +
print("[Priv + Priv] X + Y =\n", (x + y).reconstruct())
# We can not simply add them because they are at different locations
# That is why we need to first get them before adding them
print("[Pub + Pub] X + Y =\n", x_secret.get_copy() + y_secret.get_copy())
# -
print("[Priv - Priv] X - Y =\n", (x - y).reconstruct())
print("[Pub - Pub] X - Y =\n", x_secret.get_copy() - y_secret.get_copy())
print("[Priv * Priv] X * Y =\n", (x * y).reconstruct())
print("[Pub * Pub] X * Y =\n", x_secret.get_copy() * y_secret.get_copy())
print("[Priv @ Pub] X * Y =\n", (x @ y).reconstruct())
print("[Pub @ Pub] X * Y =\n", x_secret.get_copy() @ y_secret.get_copy())
| examples/secure-multi-party-computation/VMs/POC - Share Tensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
# %gui qt
from spiketag.mvc.Control import controller
from spiketag.base import probe
mua_filename = './mua.bin'
spk_filename = './spk.bin'
prb_filename = './prb_neuronexus_ff.json'
prb = probe()
prb.load(prb_filename)
# + jupyter={"outputs_hidden": true}
ctrl = controller(
probe = prb,
mua_filename=mua_filename,
spk_filename=spk_filename,
binary_radix=13,
scale=False
# time_segs=[[0,320]]
)
# -
ctrl.show()
| notebooks/template/sorter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import yaml
import pandas as pd
from SPARQLWrapper import SPARQLWrapper, JSON
# + pycharm={"name": "#%%\n"}
with open('data/ontos') as fp:
ontologies = [l.strip() for l in fp.readlines()]
ontologies.append('ontos')
# + pycharm={"name": "#%%\n"}
query = '''
SELECT (COUNT(*) as ?count) where { ?s ?p ?o . }
'''
sizes = dict()
for ontology in ontologies:
e = SPARQLWrapper(f'http://127.0.0.1:9999/blazegraph/namespace/obo-{ontology}/sparql')
e.setRequestMethod('postdirectly')
e.setMethod('POST')
e.setReturnFormat(JSON)
e.setQuery(query)
sizes[ontology] = int(e.query().convert()['results']['bindings'][0]['count']['value'])
# + pycharm={"name": "#%%\n"}
pd.Series(sizes).sort_values()
# + pycharm={"name": "#%%\n"}
sum(list(sizes.values())[:-1])
| obo/summary/ontology_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="peGes_leq5sz" colab_type="text"
# # Clear json for gaze-follow & save .mat
# + [markdown] id="Tt99z49wHARg" colab_type="text"
# ### Import dependencies
# + id="725GjGeQHARh" colab_type="code" colab={}
import numpy as np
from google.colab import files
import io
import pandas as pd
from matplotlib import pyplot as plt
# + id="RsnTdLSyLYfU" colab_type="code" outputId="0a568e37-d05c-44b1-b815-ca54e0522ce3" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 78}
uploaded = files.upload()
# + id="H7XYBhdXeJZR" colab_type="code" colab={}
json = pd.read_json('via_final.json')
# + id="TnzmrodseUzW" colab_type="code" colab={}
df = json.T
# + id="Cb0AdJmieU2I" colab_type="code" outputId="3574e56b-f1b6-4a77-d7f1-f0d2af27505c" colab={"base_uri": "https://localhost:8080/", "height": 138}
df
# + id="MW1NRLkzkkRJ" colab_type="code" outputId="affc873c-23bb-45b1-ccac-5d27df862595" colab={"base_uri": "https://localhost:8080/", "height": 35}
len(df)
# + id="vxgksJO8eU67" colab_type="code" outputId="ec15ac0f-b2ca-4aef-983f-5cd614fed276" colab={"base_uri": "https://localhost:8080/", "height": 126}
df['regions'][0][0]
# + id="U0TDUYf2fL0N" colab_type="code" colab={}
def get_bbox(i):
tmp = df['regions'][i][0]
x = tmp['shape_attributes']['x']
y = tmp['shape_attributes']['y']
height = tmp['shape_attributes']['height']
width = tmp['shape_attributes']['width']
return x, y, height, width
# + id="VjTA_mL6fLsv" colab_type="code" colab={}
def get_eyes(i):
tmp = df['regions'][i][1] # eyes
eyes_x = tmp['shape_attributes']['cx']
eyes_y = tmp['shape_attributes']['cy']
return eyes_x, eyes_y
# + id="AMKA2mY_hBZr" colab_type="code" colab={}
def get_gaze(i):
tmp = df['regions'][i][2] # gaze
gaze_x = tmp['shape_attributes']['cx']
gaze_y = tmp['shape_attributes']['cy']
return gaze_x, gaze_y
# + id="Y0LXbe1_IZmZ" colab_type="code" outputId="d77cc7a9-5585-4b8e-a3da-85dab74427c7" colab={"base_uri": "https://localhost:8080/", "height": 35}
print(get_bbox(0))
# + id="SKJ9VhY7hBiH" colab_type="code" outputId="77bf5f88-04d0-45a4-93d9-ea999642a0af" colab={"base_uri": "https://localhost:8080/", "height": 55}
my_list = []
size = 227
for i in range(len(df)):
name = df['filename'][i]
x, y, h, w = get_bbox(i)
gx, gy = get_eyes(i)
ex, ey = get_gaze(i)
my_list.append(name)
my_list.append(x/size)
my_list.append(y/size)
my_list.append(h/size)
my_list.append(w/size)
my_list.append(gx/size)
my_list.append(gy/size)
my_list.append(ex/size)
my_list.append(ey/size)
print(my_list)
# + id="sTZGn-XShBXN" colab_type="code" outputId="9f7c50be-ca6d-4923-9ad1-8cbe1df22fc1" colab={"base_uri": "https://localhost:8080/", "height": 217}
my_list = np.reshape(my_list,(-1,9))
my_list
# + id="Y-C0M-ISj1Xf" colab_type="code" outputId="515be923-7f03-449f-b124-5f25b803912c" colab={"base_uri": "https://localhost:8080/", "height": 138}
my_df = pd.DataFrame(my_list)
my_df
# + id="6NYf2U21j1e5" colab_type="code" colab={}
my_df.to_csv('ms_annotations.csv', index=False, header=False)
files.download('ms_annotations.csv')
# + id="FEWeUfzcj1lw" colab_type="code" colab={}
# + id="Q3a0V6xCj1jy" colab_type="code" colab={}
# + id="lOUdg6w_j1h1" colab_type="code" colab={}
# + id="ajPhjWT3j1ct" colab_type="code" colab={}
# + id="vH3Lokohj1at" colab_type="code" colab={}
| clear_json_to_gaze_follow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This a jupyter notebook guide on compartment analysis for chr21
#
# by <NAME> and <NAME>
#
# 2020.06.06
#
# ## Import packages
# +
# imports
import sys, os, glob, time, copy
import numpy as np
import scipy
import pickle
sys.path.append(os.path.abspath(r"..\."))
import source as ia
from scipy.signal import find_peaks
from scipy.spatial.distance import cdist,pdist,squareform
print(os.getpid()) # print this so u can terminate through cmd / task-manager
# -
# ## Import plotting
# Required plotting setting
import matplotlib
matplotlib.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
plt.rc('font', family='serif')
plt.rc('font', serif='Arial')
_font_size = 7.5
# Required plotting parameters
from source.figure_tools import _dpi,_single_col_width,_double_col_width,_single_row_height,_ref_bar_length, _ticklabel_size,_ticklabel_width,_font_size
# figure folder
parent_figure_folder = r'\\10.245.74.158\Chromatin_NAS_4\Chromatin_Share\final_figures'
figure_folder = os.path.join(parent_figure_folder, 'Chr21_compartment_figures')
print(figure_folder)
if not os.path.exists(figure_folder):
os.makedirs(figure_folder)
print("generating this folder")
# # 0. Load data
data_folder = r'E:\Users\puzheng\Dropbox\2020 Chromatin Imaging Manuscript\Revision\DataForReviewers'
rep1_filename = os.path.join(data_folder, 'chromosome21.tsv')
rep2_filename = os.path.join(data_folder, 'chromosome21-cell_cycle.tsv')
# ## 0.1 load replicate 1
# load from file and extract info
import csv
rep1_info_dict = {}
with open(rep1_filename, 'r') as _handle:
_reader = csv.reader(_handle, delimiter='\t', quotechar='|')
_headers = next(_reader)
print(_headers)
# create keys for each header
for _h in _headers:
rep1_info_dict[_h] = []
# loop through content
for _contents in _reader:
for _h, _info in zip(_headers,_contents):
rep1_info_dict[_h].append(_info)
# +
from tqdm import tqdm_notebook as tqdm
# clean up infoa
data_rep1 = {'params':{}}
# clean up genomic coordiantes
region_names = np.array([_n for _n in sorted(np.unique(rep1_info_dict['Genomic coordinate']),
key=lambda s:int(s.split(':')[1].split('-')[0]))])
region_starts = np.array([int(_n.split(':')[1].split('-')[0]) for _n in region_names])
region_ends = np.array([int(_n.split(':')[1].split('-')[1]) for _n in region_names])[np.argsort(region_starts)]
region_starts = np.sort(region_starts)
mid_positions = ((region_starts + region_ends)/2).astype(np.int)
mid_positions_Mb = np.round(mid_positions / 1e6, 2)
# clean up chrom copy number
chr_nums = np.array([int(_info) for _info in rep1_info_dict['Chromosome copy number']])
chr_ids, region_cts = np.unique(chr_nums, return_counts=True)
dna_zxys_list = [[[] for _start in region_starts] for _id in chr_ids]
# clean up zxy
for _z,_x,_y,_reg_info, _cid in tqdm(zip(rep1_info_dict['Z(nm)'],rep1_info_dict['X(nm)'],\
rep1_info_dict['Y(nm)'],rep1_info_dict['Genomic coordinate'],\
rep1_info_dict['Chromosome copy number'])):
# get chromosome inds
_cid = int(_cid)
_cind = np.where(chr_ids == _cid)[0][0]
# get region indices
_start = int(_reg_info.split(':')[1].split('-')[0])
_rind = np.where(region_starts==_start)[0][0]
dna_zxys_list[_cind][_rind] = np.array([float(_z),float(_x), float(_y)])
# merge together
dna_zxys_list = np.array(dna_zxys_list)
data_rep1['chrom_ids'] = chr_ids
data_rep1['region_names'] = region_names
data_rep1['mid_position_Mb'] = mid_positions_Mb
data_rep1['dna_zxys'] = dna_zxys_list
# clean up tss and transcription
if 'Gene names' in rep1_info_dict:
import re
# first extract number of genes
gene_names = []
for _gene_info, _trans_info, _tss_coord in zip(rep1_info_dict['Gene names'],
rep1_info_dict['Transcription'],
rep1_info_dict['TSS ZXY(nm)']):
if _gene_info != '':
# split by semicolon
_genes = _gene_info.split(';')[:-1]
for _gene in _genes:
if _gene not in gene_names:
gene_names.append(_gene)
print(f"{len(gene_names)} genes exist in this dataset.")
# initialize gene and transcription
tss_zxys_list = [[[] for _gene in gene_names] for _id in chr_ids]
transcription_profiles = [[[] for _gene in gene_names] for _id in chr_ids]
# loop through to get info
for _cid, _gene_info, _trans_info, _tss_locations in tqdm(zip(rep1_info_dict['Chromosome copy number'],
rep1_info_dict['Gene names'],
rep1_info_dict['Transcription'],
rep1_info_dict['TSS ZXY(nm)'])):
# get chromosome inds
_cid = int(_cid)
_cind = np.where(chr_ids == _cid)[0][0]
# process if there are genes in this region:
if _gene_info != '':
# split by semicolon
_genes = _gene_info.split(';')[:-1]
_transcribes = _trans_info.split(';')[:-1]
_tss_zxys = _tss_locations.split(';')[:-1]
for _gene, _transcribe, _tss_zxy in zip(_genes, _transcribes, _tss_zxys):
# get gene index
_gind = gene_names.index(_gene)
# get transcription profile
if _transcribe == 'on':
transcription_profiles[_cind][_gind] = True
else:
transcription_profiles[_cind][_gind] = False
# get coordinates
_tss_zxy = np.array([np.float(_c) for _c in re.split(r'\s+', _tss_zxy.split('[')[1].split(']')[0]) if _c != ''])
tss_zxys_list[_cind][_gind] = _tss_zxy
tss_zxys_list = np.array(tss_zxys_list)
transcription_profiles = np.array(transcription_profiles)
data_rep1['gene_names'] = gene_names
data_rep1['tss_zxys'] = tss_zxys_list
data_rep1['trans_pfs'] = transcription_profiles
# clean up cell_cycle states
if 'Cell cycle state' in rep1_info_dict:
cell_cycle_types = np.unique(rep1_info_dict['Cell cycle state'])
cell_cycle_flag_dict = {_k:[[] for _id in chr_ids] for _k in cell_cycle_types if _k != 'ND'}
for _cid, _state in tqdm(zip(rep1_info_dict['Chromosome copy number'],rep1_info_dict['Cell cycle state'])):
# get chromosome inds
_cid = int(_cid)
_cind = np.where(chr_ids == _cid)[0][0]
if np.array([_v[_cind]==[] for _k,_v in cell_cycle_flag_dict.items()]).any():
for _k,_v in cell_cycle_flag_dict.items():
if _k == _state:
_v[_cind] = True
else:
_v[_cind] = False
# append to data
for _k, _v in cell_cycle_flag_dict.items():
data_rep1[f'{_k}_flags'] = np.array(_v)
# -
# # Get population averaged maps
# ## imaging-based proximity freq. matrix and median distance matrix
# +
zxys_rep1_list = np.array(data_rep1['dna_zxys'])
distmap_rep1_list = np.array([squareform(pdist(_zxy)) for _zxy in zxys_rep1_list])
# generate median distance map
median_distance_map_rep1 = np.nanmedian(distmap_rep1_list, axis = 0)
# generate contact map
contact_th = 500
contact_rep1_map = np.sum(distmap_rep1_list<contact_th, axis=0) / np.sum(np.isnan(distmap_rep1_list)==False, axis=0)
# -
# ## corresponding Hi-C data from Rao et al.
# +
hic_filename = os.path.join(data_folder, 'Hi-C matrices', 'Hi-C_contacts_chromosome21.tsv')
hic_map = []
with open(hic_filename, 'r') as _handle:
_reader = csv.reader(_handle, delimiter='\t', quotechar='|')
col_regions = next(_reader)[1:]
row_regions = []
# loop through content
for _contents in _reader:
row_regions.append(_contents[0])
hic_map.append([int(_c) for _c in _contents[1:]])
hic_map = np.array(hic_map)
# sort row and col to match tsv dataset
row_order = np.concatenate([np.where(data_rep1['region_names']==_rn)[0] for _rn in row_regions])
col_order = np.concatenate([np.where(data_rep1['region_names']==_cn)[0] for _cn in col_regions])
hic_map = hic_map[row_order][:, col_order]
# -
# ## 0.2 call compartments
# +
## compartment calling by PCA
# Generate correlation map
gaussian_sigma = 2.75
# normalize genomic distance effects
genomic_distance_map = squareform(pdist(data_rep1['mid_position_Mb'][:,np.newaxis]))
genomic_distance_entries = genomic_distance_map[np.triu_indices(len(genomic_distance_map),1)]
median_entries = median_distance_map_rep1[np.triu_indices(len(median_distance_map_rep1),1)]
kept = (genomic_distance_entries > 0) * (median_entries > 0)
median_lr = scipy.stats.linregress(np.log(genomic_distance_entries[kept]), np.log(median_entries[kept]))
print(median_lr)
median_norm_map = np.exp(np.log(genomic_distance_map) * median_lr.slope + median_lr.intercept)
for _i in range(len(median_norm_map)):
median_norm_map[_i,_i] = 1
median_normed_map = median_distance_map_rep1 / median_norm_map
# apply gaussian
from scipy.ndimage import gaussian_filter
median_corr_map_rep1 = np.corrcoef(gaussian_filter(median_normed_map, gaussian_sigma))
# normalize genomic distance effects
contact_entries = contact_rep1_map[np.triu_indices(len(contact_rep1_map),1)]
kept = (genomic_distance_entries > 0) * (contact_entries > 0)
contact_lr = scipy.stats.linregress(np.log(genomic_distance_entries[kept]), np.log(contact_entries[kept]))
print(contact_lr)
contact_norm_map = np.exp(np.log(genomic_distance_map) * contact_lr.slope + contact_lr.intercept)
#for _i in range(len(normalization_mat)):
# contact_norm_map[_i,_i] = 1
contact_normed_map = contact_rep1_map / contact_norm_map
# apply gaussian
from scipy.ndimage import gaussian_filter
contact_corr_map_rep1 = np.corrcoef(gaussian_filter(contact_normed_map, gaussian_sigma))
# normalize genomic distance effects
hic_entries = hic_map[np.triu_indices(len(hic_map),1)]
kept = (genomic_distance_entries > 0) * (hic_entries > 0)
hic_lr = scipy.stats.linregress(np.log(genomic_distance_entries[kept]), np.log(hic_entries[kept]))
print(hic_lr)
hic_norm_map = np.exp(np.log(genomic_distance_map) * hic_lr.slope + hic_lr.intercept)
#for _i in range(len(normalization_mat)):
# hic_norm_map[_i,_i] = 1
hic_normed_map = hic_map / hic_norm_map
# apply gaussian
from scipy.ndimage import gaussian_filter
hic_corr_map = np.corrcoef(gaussian_filter(hic_normed_map, gaussian_sigma))
# Do PCA
from sklearn.decomposition import PCA
median_model = PCA(1)
median_model.fit(median_corr_map_rep1)
median_pc1_rep1 = np.reshape(median_model.fit_transform(median_corr_map_rep1), -1)
contact_model = PCA(1)
contact_model.fit(contact_corr_map_rep1)
contact_pc1_rep1 = np.reshape(contact_model.fit_transform(contact_corr_map_rep1), -1)
hic_model = PCA(1)
hic_model.fit(hic_corr_map)
hic_pc1 = np.reshape(hic_model.fit_transform(hic_corr_map), -1)
# +
# define AB compartment by merging small sub-compartments
temp_AB_dict = {'A':np.where(contact_pc1_rep1 >= 0)[0],
'B':np.where(contact_pc1_rep1 < 0)[0],}
temp_AB_vector = np.ones(len(zxys_rep1_list[0])).astype(np.int) * -1
temp_AB_vector[temp_AB_dict['A']] = 1
temp_AB_vector[temp_AB_dict['B']] = 0
num_small_compartment = np.inf
prev_v = temp_AB_vector[0]
while num_small_compartment > 0:
# find indices for all sub-comaprtments
all_comp_inds = []
_comp_inds = []
prev_v = temp_AB_vector[0] # initialize previous compartment
for _i, _v in enumerate(temp_AB_vector):
if prev_v != _v:
all_comp_inds.append(_comp_inds)
_comp_inds = [_i]
else:
_comp_inds.append(_i)
prev_v = _v
if _comp_inds != []:
all_comp_inds.append(_comp_inds)
# calculate length of each compartment
all_comp_lens = np.array([len(_c) for _c in all_comp_inds])
# update number of small comparment
num_small_compartment = np.sum(all_comp_lens < 4)
print(all_comp_lens, num_small_compartment)
# choose the smallest compartment to flip its AB
flip_ind = np.argmin(all_comp_lens)
temp_AB_vector[np.array(all_comp_inds[flip_ind])] = 1 - temp_AB_vector[np.array(all_comp_inds[flip_ind])]
# based on this cleaned AB_vector, recreate AB_dict
data_rep1['AB_dict'] = {
'A': np.where(temp_AB_vector==1)[0],
'B': np.where(temp_AB_vector==0)[0],
}
# -
# # Plots related to compartment calling
# ## Correlation map with AB calling
# +
lims = [0,len(contact_corr_map_rep1)]
xlims = np.array([min(lims), max(lims)])
ylims = np.array([min(lims), max(lims)])
from mpl_toolkits.axes_grid1 import make_axes_locatable
domain_line_color = [1,1,0,1]
domain_line_width = 1.5
bad_color=[0,0,0,1]
fig, ax1 = plt.subplots(figsize=(_single_col_width, _single_col_width), dpi=600)
# create a color map
current_cmap = matplotlib.cm.get_cmap('seismic')
current_cmap.set_bad(color=[0.5,0.5,0.5,1])
_pf = ax1.imshow(contact_corr_map_rep1, cmap=current_cmap, vmin=-1, vmax=1)
ax1.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=0,
pad=1, labelleft=False, labelbottom=False) # remove bottom ticklabels for ax1
[i[1].set_linewidth(_ticklabel_width) for i in ax1.spines.items()]
# locate ax1
divider = make_axes_locatable(ax1)
# colorbar ax
cax = divider.append_axes('right', size='6%', pad="4%")
cbar = plt.colorbar(_pf,cax=cax, ax=ax1, ticks=[-1,1])
cbar.ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size-1,
pad=1, labelleft=False) # remove bottom ticklabels for ax1
cbar.outline.set_linewidth(_ticklabel_width)
cbar.set_label('Pearson correlation',
fontsize=_font_size, labelpad=0, rotation=270)
# create bottom ax
bot_ax = divider.append_axes('bottom', size='10%', pad="0%",
sharex=ax1, xticks=[])
bot_ax.bar(data_rep1['AB_dict']['A'], height=1, color='r', width=1, label='A')
bot_ax.bar(data_rep1['AB_dict']['B'], height=-1, color='b', width=1, label='B')
bot_ax.set_yticks([])
bot_ax.set_yticklabels([])
bot_ax.set_ylim([-1,1])
_xticks = [0, len(contact_corr_map_rep1)-1]
bot_ax.set_xticks(_xticks)
bot_ax.set_xticklabels(np.round(mid_positions_Mb[_xticks],1))
# tick params
bot_ax.tick_params('both', labelsize=_font_size-0.5,
width=_ticklabel_width, length=_ticklabel_size-1,
pad=1, labelleft=False, labelbottom=True) # remove bottom ticklabels for ax1
[i[1].set_linewidth(_ticklabel_width) for i in bot_ax.spines.items()]
# set labels
bot_ax.set_xlabel(f'Genomic Positions (Mb)', fontsize=_font_size, labelpad=1)
# create left ax
#left_ax = divider.append_axes('left', size='10%', pad="0%", sharey=ax1, xticks=[])
#left_ax.barh(data_rep1['AB_dict']['A'], height=1, color='r', width=1, label='A')
#left_ax.barh(data_rep1['AB_dict']['B'], height=1, left=-1, color='b', width=1, label='B')
#left_ax.set_xticks([-0.5, 0.5])
#left_ax.set_xticklabels(['B', 'A'])
#_yticks = _xticks
#left_ax.set_yticks(_yticks)
#left_ax.set_yticklabels(mid_positions_Mb[_yticks])
#left_ax.tick_params('both', labelsize=_font_size-1,
# width=_ticklabel_width, length=_ticklabel_size-1,
# pad=1, labelleft=True) # remove bottom ticklabels for ax1
#[i[1].set_linewidth(_ticklabel_width) for i in left_ax.spines.items()]
#
#left_ax.set_ylabel(f'Genomic Positions (Mb)', fontsize=_font_size, labelpad=1)
# set limits
bot_ax.set_xlim(xlims-0.5)
#left_ax.set_ylim([max(ylims)-0.5, min(ylims)-0.5])
ax1.set_title(f"Chr21 (~3,500 cells)", fontsize=_font_size+0.5)
# save
plt.gcf().subplots_adjust(bottom=0.15, left=0.16, right=0.88)
plt.savefig(os.path.join(figure_folder, f'Fig2A_chr21_contact_corr_map_rep1.pdf'), transparent=True)
plt.show()
# -
# ## Plot PC1 for chr21 calling
# +
## pc1 barplot
fig, ax = plt.subplots(figsize=(_double_col_width, _single_col_width), dpi=600)
grid = plt.GridSpec(2, 1, height_ratios=[1,1], hspace=0., wspace=0.)
contact_ax = plt.subplot(grid[0])
contact_ax.bar(np.where(contact_pc1_rep1>=0)[0],
contact_pc1_rep1[contact_pc1_rep1>=0],
width=1, color='r', label='A')
contact_ax.bar(np.where(contact_pc1_rep1<0)[0],
contact_pc1_rep1[contact_pc1_rep1<0],
width=1, color='b', label='B')
contact_ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1,labelbottom=False) # remove bottom ticklabels for ax1
[i[1].set_linewidth(_ticklabel_width) for i in contact_ax.spines.items()]
contact_ax.set_ylim([-15,25])
contact_ax.set_yticks([-10,0,10,20])
contact_ax.set_ylabel("Contact PC1", fontsize=_font_size, labelpad=0)
# hic-ax
hic_ax = plt.subplot(grid[1], sharex=contact_ax)
hic_ax.bar(np.where(hic_pc1>=0)[0],
hic_pc1[hic_pc1>=0],
width=1, color='r', label='A')
hic_ax.bar(np.where(hic_pc1<0)[0],
hic_pc1[hic_pc1<0],
width=1, color='b', label='B')
hic_ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1,) # remove bottom ticklabels for ax1
[i[1].set_linewidth(_ticklabel_width) for i in hic_ax.spines.items()]
hic_ax.set_ylim([-15,25])
hic_ax.set_yticks([-10,0,10,20])
hic_ax.set_ylabel("Hi-C PC1", fontsize=_font_size, labelpad=0)
# set x
hic_ax.set_xlim([0, len(contact_pc1_rep1)])
_xticks = [0, len(contact_pc1_rep1)-1]
hic_ax.set_xticks(_xticks)
hic_ax.set_xticklabels(mid_positions_Mb[_xticks])
hic_ax.set_xlabel(f'Genomic Positions (Mb)', fontsize=_font_size, labelpad=1)
plt.gcf().subplots_adjust(bottom=0.15, left=0.1)
plt.savefig(os.path.join(figure_folder, f'FigS2A_chr21_PC1_barplot_rep1.pdf'), transparent=True)
plt.show()
# -
# ### 0.1.3 density scores
# +
import multiprocessing as mp
num_threads=32
density_var = 108 # nm
# density score for 50kb genomic regions
_dna_density_args = [(_zxys,_zxys, data_rep1['AB_dict'], [density_var,density_var,density_var], True)
for _zxys in data_rep1['dna_zxys']]
_dna_density_time = time.time()
print(f"Multiprocessing calculate dna_density_scores", end=' ')
if 'dna_density_scores' not in data_rep1:
with mp.Pool(num_threads) as dna_density_pool:
dna_density_dicts = dna_density_pool.starmap(ia.compartment_tools.scoring.spot_density_scores, _dna_density_args)
dna_density_pool.close()
dna_density_pool.join()
dna_density_pool.terminate()
# save
data_rep1['dna_density_scores'] = dna_density_dicts
print(f"in {time.time()-_dna_density_time:.3f}s.")
# density score for tss
_gene_density_args = [(_gzxys,_zxys, data_rep1['AB_dict'], [density_var,density_var,density_var], True)
for _gzxys, _zxys in zip(data_rep1['tss_zxys'], data_rep1['dna_zxys']) ]
_gene_density_time = time.time()
print(f"Multiprocessing calculate gene_density_scores", end=' ')
if 'gene_density_scores' not in data_rep1:
with mp.Pool(num_threads) as gene_density_pool:
gene_density_dicts = gene_density_pool.starmap(ia.compartment_tools.scoring.spot_density_scores, _gene_density_args)
gene_density_pool.close()
gene_density_pool.join()
gene_density_pool.terminate()
# save
data_rep1['gene_density_scores'] = gene_density_dicts
print(f"in {time.time()-_gene_density_time:.3f}s.")
# -
# ## Mean density for chr21
# +
# Calculate mean A, B density
mean_A_scores = np.nanmedian([_s['A'] for _s in data_rep1['dna_density_scores']], axis=0)
mean_B_scores = np.nanmedian([_s['B'] for _s in data_rep1['dna_density_scores']], axis=0)
# Plot
fig = plt.figure(figsize=(_double_col_width, _single_col_width),dpi=600)
grid = plt.GridSpec(2, 1, height_ratios=[7,1], hspace=0., wspace=0.2)
main_ax = plt.subplot(grid[0], xticklabels=[])
main_ax.plot(mean_A_scores, 'r.--', label='A density', markersize=2, linewidth=1)
main_ax.plot(mean_B_scores, 'b.--', label='B density', markersize=2, linewidth=1)
# ticks
main_ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1, labelbottom=False) # remove bottom ticklabels for ax1
main_ax.tick_params('x', length=0)
[i[1].set_linewidth(_ticklabel_width) for i in main_ax.spines.items()]
main_ax.set_ylabel(f"Mean density scores", fontsize=_font_size, labelpad=1)
handles, labels = main_ax.get_legend_handles_labels()
main_ax.legend(handles[::-1], labels[::-1], fontsize=_font_size, loc='upper right')
main_ax.set_xlim(0,len(mean_A_scores))
comp_ax = plt.subplot(grid[1], xticklabels=[], sharex=main_ax)
comp_ax.eventplot([data_rep1['AB_dict']['A'], data_rep1['AB_dict']['B']], lineoffsets=[0.5,-0.5],linelengths=1, linewidths=1,
colors=np.array([[1, 0, 0],[0, 0, 1]]))
#comp_ax.imshow(comp_vector[np.newaxis,:], cmap='seismic', vmin=-1, vmax=1)
comp_ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1, labelbottom=True) # remove bottom ticklabels for ax1
[i[1].set_linewidth(_ticklabel_width) for i in comp_ax.spines.items()]
comp_ax.set_yticks([0.5,-0.5])
comp_ax.set_yticklabels(["A","B"])
comp_ax.set_ylim([-1,1])
_xticks = [0, len(mean_A_scores)-1]
comp_ax.set_xticks(_xticks)
comp_ax.set_xticklabels(mid_positions_Mb[_xticks])
comp_ax.set_xlabel(f'Genomic Positions (Mb)', fontsize=_font_size, labelpad=1)
plt.gcf().subplots_adjust(bottom=0.1, left=0.05)
plt.savefig(os.path.join(figure_folder, 'FigS2C_chr21_mean_AB_density_rep1.pdf'), transparent=True)
plt.show()
# -
# ### 0.1.4 segregation scores
# +
def randomize_AB_dict(AB_dict):
all_regs = np.sort(np.concatenate(list(AB_dict.values())))
AB_identities = np.zeros(len(all_regs))
AB_identities[np.array([_i for _i,_r in enumerate(all_regs)
if _r in AB_dict['A']])] = 1
# randomize new start
new_start = np.random.randint(0, len(all_regs))
new_AB_identities = np.concatenate([AB_identities[new_start:], AB_identities[:new_start]])
# recreate AB_dict
new_AB_dict = {'A': np.sort(all_regs[np.where(new_AB_identities==1)[0]]),
'B': np.sort(all_regs[np.where(new_AB_identities==0)[0]]),}
return new_AB_dict
# calculate dynamic fraction scores
from scipy.stats import scoreatpercentile
# +
AB_identities_rep1 = np.ones(len(data_rep1['dna_zxys'][0])) * np.nan
AB_identities_rep1[data_rep1['AB_dict']['A']] = 1
AB_identities_rep1[data_rep1['AB_dict']['B']] = 0
from tqdm import tqdm
# calculate re-thresholded fraction scores
A_fracs, B_fracs = [], []
A_ths, B_ths = [], []
cloud_th_per=67
for _sd in tqdm(data_rep1['dna_density_scores']):
# define A,B threshold based on their own densities
_A_th = scoreatpercentile(_sd['A'][data_rep1['AB_dict']['A']], 100-cloud_th_per)
_B_th = scoreatpercentile(_sd['B'][data_rep1['AB_dict']['B']], 100-cloud_th_per)
# calculate purity within A,B clouds
A_fracs.append(np.nanmean(AB_identities_rep1[np.where(_sd['A'] >= _A_th)[0]]))
B_fracs.append(1-np.nanmean(AB_identities_rep1[np.where(_sd['B'] >= _B_th)[0]]))
# store AB thresholds for references
A_ths.append(_A_th)
B_ths.append(_B_th)
# calculate re-thresholded fraction scores
rand_A_fracs, rand_B_fracs = [], []
for _sd in tqdm(data_rep1['dna_density_scores']):
# randomize AB dict
_rand_AB_dict = randomize_AB_dict(data_rep1['AB_dict'])
_rand_A_inds, _rand_B_inds = np.array(_rand_AB_dict['A']), np.array(_rand_AB_dict['B'])
# generate randomized AB_identities_rep1 vector for purity calculation
_rand_AB_identities_rep1 = np.ones(len(data_rep1['dna_zxys'][0])) * np.nan
_rand_AB_identities_rep1[_rand_AB_dict['A']] = 1
_rand_AB_identities_rep1[_rand_AB_dict['B']] = 0
# define A,B threshold based on their own densities
_A_th = scoreatpercentile(_sd['A'][_rand_A_inds], 100-cloud_th_per)
_B_th = scoreatpercentile(_sd['B'][_rand_B_inds], 100-cloud_th_per)
# calculate purity within A,B clouds
rand_A_fracs.append(np.nanmean(_rand_AB_identities_rep1[np.where(_sd['A'] >= _A_th)[0]]))
rand_B_fracs.append(1-np.nanmean(_rand_AB_identities_rep1[np.where(_sd['B'] >= _B_th)[0]]))
# Save
data_rep1['segregation_scores'] = (np.array(A_fracs) + np.array(B_fracs)) / 2
data_rep1['randomized_segregation_scores'] = (np.array(rand_A_fracs) + np.array(rand_B_fracs)) / 2
# -
# ## Segregation score histogram for Chr21
# +
# %matplotlib inline
fig, ax = plt.subplots(figsize=(_single_col_width, _single_col_width),dpi=600)
ax.hist(data_rep1['segregation_scores'], 100, range=(0.,1),
density=True, alpha=0.5,
color=[1,0.5,0], label='Chr21')
ax.hist(data_rep1['randomized_segregation_scores'], 100, range=(0.,1),
density=True, alpha=0.5,
color=[0.3,0.4,0.4], label='randomized control')
ax.legend(fontsize=_font_size-1, loc='upper right')
ax.set_xlabel("Segregation score", fontsize=_font_size, labelpad=1)
ax.set_ylabel("Probability density", fontsize=_font_size, labelpad=1)
ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1, labelleft=True) # remove bottom ticklabels for a_ax
[i[1].set_linewidth(_ticklabel_width) for i in ax.spines.items()]
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.set_xlim([0.4,1])
plt.gcf().subplots_adjust(bottom=0.15, left=0.15)
plt.savefig(os.path.join(figure_folder, 'Fig2C_chr21_segregation_hist_rep1.pdf'), transparent=True)
plt.show()
# -
# ## A/B density ratio difference w/wo transcription
# +
gene_density_dicts = data_rep1['gene_density_scores']
gene_A_scores = np.array([_gsd['A'] for _gsd in gene_density_dicts])
gene_B_scores = np.array([_gsd['B'] for _gsd in gene_density_dicts])
trans_pfs = np.array(data_rep1['trans_pfs'])
on_gene_AB_ratio, off_gene_AB_ratio = [], []
on_gene_total_density = []
off_gene_total_density = []
for _gind in range(gene_A_scores.shape[1]):
# extract AB and transcription
_a_scores = gene_A_scores[:, _gind]
_b_scores = gene_B_scores[:, _gind]
_transcriptions = trans_pfs[:,_gind]
# get AB ratio
_log_ab_ratios = np.log(_a_scores)/np.log(2) - np.log(_b_scores)/np.log(2)
_total_density = _a_scores+_b_scores
# append
on_gene_AB_ratio.append(np.nanmedian(_log_ab_ratios[_transcriptions & (np.isinf(_log_ab_ratios)==False)]))
off_gene_AB_ratio.append(np.nanmedian(_log_ab_ratios[~_transcriptions & (np.isinf(_log_ab_ratios)==False)]))
on_gene_total_density.append(np.nanmedian(_total_density[_transcriptions & (np.isinf(_total_density)==False)]))
off_gene_total_density.append(np.nanmedian(_total_density[~_transcriptions & (np.isinf(_total_density)==False)]))
# convert into arrays
on_gene_AB_ratio = np.array(on_gene_AB_ratio)
off_gene_AB_ratio = np.array(off_gene_AB_ratio)
on_gene_total_density = np.array(on_gene_total_density)
off_gene_total_density = np.array(off_gene_total_density)
gene_ratio_diff = on_gene_AB_ratio - off_gene_AB_ratio
fig, ax = plt.subplots(figsize=(_single_col_width, _single_col_width),dpi=600)
ax.plot(np.sort(gene_ratio_diff), '.', color=[1,0.5,0], label='All genes', markersize=3)
ax.plot(np.zeros(len(gene_ratio_diff)), 'black', label='ref', linewidth=0.75)
ax.tick_params('both', labelsize=_font_size, width=_ticklabel_width, length=_ticklabel_size, pad=1)
# remove bottom ticklabels for ax
[i[1].set_linewidth(_ticklabel_width) for i in ax.spines.items()]
ax.spines["top"].set_visible(False)
ax.spines["right"].set_visible(False)
ax.set_xticks(np.arange(len(gene_ratio_diff)))
ax.set_yticks(np.arange(-6,6,2))
ax.set_xticklabels([])
ax.set_xlabel('Genes', fontsize=_font_size, labelpad=2)
ax.set_ylabel('Log2 difference of \nA/B density ratio', fontsize=_font_size, labelpad=2)
ax.set_xlim([-1, len(gene_ratio_diff)])
ax.set_ylim([-2,4])
#ax.legend(fontsize=_font_size, framealpha=1, loc='upper left')
ax.text(len(gene_ratio_diff), max(ax.get_ylim())/30, f'{np.round(np.mean(gene_ratio_diff>0),2)*100:.0f}%',
fontsize=_font_size-1, verticalalignment='bottom', horizontalalignment='right',)
ax.text(len(gene_ratio_diff), -max(ax.get_ylim())/30, f'{np.round(np.mean(gene_ratio_diff<0),2)*100:.0f}%',
fontsize=_font_size-1, verticalalignment='top', horizontalalignment='right',)
ax.set_title("Transcribing/Silent", fontsize=_font_size)
plt.gcf().subplots_adjust(left=0.2, bottom=0.15)
plt.savefig(os.path.join(figure_folder, 'Fig2H_diff_abratio_w-wo_transcribe_rep1.pdf'), transparent=True)
plt.show()
# -
# ## Firing rate difference with high/low A/B density ratio
# +
gene_density_dicts = data_rep1['gene_density_scores']
gene_A_scores = np.array([_gsd['A'] for _gsd in gene_density_dicts])
gene_B_scores = np.array([_gsd['B'] for _gsd in gene_density_dicts])
trans_pfs = np.array(data_rep1['trans_pfs'])
abratio_ratio = []
for _gid in np.arange(trans_pfs.shape[1]):
# extract info
_ga = gene_A_scores[:,_gid]
_gb = gene_B_scores[:,_gid]
_gratio = np.log(_ga / _gb)
# sel_inds
_inds = np.argsort(_gratio)
_sel_v = np.where((np.isnan(_gratio)==False) * (np.isinf(_gratio)==False))[0]
_sel_inds = np.array([_i for _i in _inds if _i in _sel_v], dtype=np.int)
_li = _sel_inds[:int(len(_sel_inds)/4)]
_hi = _sel_inds[-int(len(_sel_inds)/4):]
#print(len(_sel_inds), np.mean(_gratio[_li]), np.mean(_gratio[_hi]))
_lfr = np.nanmean(trans_pfs[_li, _gid])
_hfr = np.nanmean(trans_pfs[_hi, _gid])
abratio_ratio.append(_hfr / _lfr)
abratio_fr_ratio = np.log(np.array(abratio_ratio)) / np.log(2)
fig, ax = plt.subplots(figsize=(_single_col_width, _single_col_width),dpi=600)
#ax.plot(np.arange(len(sel_A_gene_inds)),
# np.sort(np.log(abratio_ratio[sel_A_gene_inds])),
# 'r.', label='A genes', markersize=3)
#ax.plot(np.arange(len(sel_A_gene_inds),len(sel_A_gene_inds)+len(sel_B_gene_inds)),
# np.sort(np.log(abratio_ratio[sel_B_gene_inds])),
# 'b.', label='B genes', markersize=3)
ax.plot(np.sort(abratio_fr_ratio),
'.', color=[1,0.5,0], label='All genes', markersize=3)
ax.plot(np.zeros(len(abratio_fr_ratio)),
'black', label='reference', linewidth=0.75)
ax.tick_params('both', labelsize=_font_size,
width=_ticklabel_width, length=_ticklabel_size,
pad=1) # remove bottom ticklabels for ax1
ax.set_xticks(np.arange(len(abratio_fr_ratio)))
ax.set_xticklabels([])
ax.set_ylabel('Log odds ratio', fontsize=_font_size, labelpad=1)
ax.set_xlim([-1, len(abratio_fr_ratio)])
ax.set_ylim([-1,2])
[i[1].set_linewidth(_ticklabel_width) for i in ax.spines.items()]
ax.spines["top"].set_visible(False)
ax.spines["right"].set_visible(False)
ax.set_xlabel('Genes', fontsize=_font_size, labelpad=2)
ax.set_ylabel('Log2 difference of firing rate', fontsize=_font_size, labelpad=1)
ax.set_title(f"high/low log(A/B) density ratio",
fontsize=_font_size)
#ax.legend(fontsize=_font_size, framealpha=1, loc='upper left')
ax.text(len(abratio_fr_ratio), max(ax.get_ylim())/30, f'{np.round(np.mean(abratio_fr_ratio>0),2)*100:.0f}%',
fontsize=_font_size-1, verticalalignment='bottom', horizontalalignment='right',)
ax.text(len(abratio_fr_ratio), -max(ax.get_ylim())/30, f'{np.round(np.mean(abratio_fr_ratio<=0),2)*100:.0f}%',
fontsize=_font_size-1, verticalalignment='top', horizontalalignment='right',)
plt.gcf().subplots_adjust(left=0.2, bottom=0.15)
plt.savefig(os.path.join(figure_folder, f"Fig2I_firing-rate_diff_w_high-low_abratio_rep1.pdf"), transparent=True)
plt.show()
# -
| sequential_tracing/PostAnalysis/Part2_Compartment_Analysis_chr21.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dev
# language: python
# name: dev
# ---
import numpy as np
from sklearn.datasets import load_iris
from sklearn.dummy import DummyClassifier as skDummyClassifier
class DummyClassifier():
def __init__(self, strategy="stratified", random_state=0, constant=None):
self.strategy = strategy
self.constant = constant
self.random_state = random_state
def fit(self, X, y):
self.classes_, y_train = np.unique(y, return_inverse=True)
self.n_classes_ = self.classes_.shape[0]
self.class_prior_ = np.bincount(y_train) / y.shape[0]
return self
def predict(self, X):
if self.strategy == "most_frequent" or self.strategy == "prior":
y = np.full(X.shape[0], self.classes_[np.argmax(self.class_prior_)])
elif self.strategy == "constant":
y = np.full(X.shape[0], self.constant)
elif self.strategy == "uniform":
rng = np.random.RandomState(self.random_state)
y = self.classes_[rng.randint(self.n_classes_, size=X.shape[0])]
elif self.strategy == "stratified":
y = self.classes_[np.argmax(self.predict_proba(X), axis=1)]
return y
def predict_proba(self, X):
if self.strategy == "most_frequent":
p = np.zeros((X.shape[0], self.n_classes_))
p[:, np.argmax(self.class_prior_)] = 1
elif self.strategy == "prior":
p = np.tile(self.class_prior_, (X.shape[0], 1))
elif self.strategy == "constant":
p = np.zeros((X.shape[0], self.n_classes_))
p[:, self.classes_ == self.constant] = 1
elif self.strategy == "uniform":
p = np.full((X.shape[0], self.n_classes_), 1 / self.n_classes_)
elif self.strategy == "stratified":
rng = np.random.RandomState(self.random_state)
p = rng.multinomial(1, self.class_prior_, size=X.shape[0])
return p
X, y = load_iris(return_X_y=True)
clf1 = DummyClassifier(strategy="most_frequent").fit(X, y)
clf2 = skDummyClassifier(strategy="most_frequent").fit(X, y)
assert clf1.n_classes_ == clf2.n_classes_
assert np.array_equal(clf1.classes_, clf2.classes_)
assert np.array_equal(clf1.class_prior_, clf2.class_prior_)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
prob1 = clf1.predict_proba(X)
prob2 = clf2.predict_proba(X)
assert np.array_equal(prob1, prob2)
clf1 = DummyClassifier(strategy="prior").fit(X, y)
clf2 = skDummyClassifier(strategy="prior").fit(X, y)
assert clf1.n_classes_ == clf2.n_classes_
assert np.array_equal(clf1.classes_, clf2.classes_)
assert np.array_equal(clf1.class_prior_, clf2.class_prior_)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
prob1 = clf1.predict_proba(X)
prob2 = clf2.predict_proba(X)
assert np.array_equal(prob1, prob2)
clf1 = DummyClassifier(strategy="constant", constant=0).fit(X, y)
clf2 = skDummyClassifier(strategy="constant", constant=0).fit(X, y)
assert clf1.n_classes_ == clf2.n_classes_
assert np.array_equal(clf1.classes_, clf2.classes_)
assert np.array_equal(clf1.class_prior_, clf2.class_prior_)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
prob1 = clf1.predict_proba(X)
prob2 = clf2.predict_proba(X)
assert np.array_equal(prob1, prob2)
clf1 = DummyClassifier(strategy="uniform", random_state=0).fit(X, y)
clf2 = skDummyClassifier(strategy="uniform", random_state=0).fit(X, y)
assert clf1.n_classes_ == clf2.n_classes_
assert np.array_equal(clf1.classes_, clf2.classes_)
assert np.array_equal(clf1.class_prior_, clf2.class_prior_)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
prob1 = clf1.predict_proba(X)
prob2 = clf2.predict_proba(X)
assert np.array_equal(prob1, prob2)
clf1 = DummyClassifier(strategy="stratified", random_state=0).fit(X, y)
clf2 = skDummyClassifier(strategy="stratified", random_state=0).fit(X, y)
assert clf1.n_classes_ == clf2.n_classes_
assert np.array_equal(clf1.classes_, clf2.classes_)
assert np.array_equal(clf1.class_prior_, clf2.class_prior_)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
#assert np.array_equal(pred1, pred2)
prob1 = clf1.predict_proba(X)
prob2 = clf2.predict_proba(X)
assert np.array_equal(prob1, prob2)
| dummy/DummyClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# It was proposed by <NAME> that every odd composite number can be written as the sum of a prime and twice a square.
#
# $$
# 9 = 7 + 2 \cdot 1^2 \\
# 15 = 7 + 2 \cdot 2^2 \\
# 21 = 3 + 2 \cdot 3^2 \\
# 25 = 7 + 2 \cdot 3^2 \\
# 27 = 19 + 2 \cdot 2^2 \\
# 33 = 31 + 2 \cdot 1^2
# $$
#
# It turns out that the conjecture was false.
#
# What is the smallest odd composite that cannot be written as the sum of a prime and twice a square?
# ### Version 1: Brute force
# Formally stated, Goldbach's Other Conjecture says that all odd composite numbers $n$ can be expressed as
#
# $$
# n = 2k^2 + p
# $$
#
# for some integer $k$ and prime number $p$. Let
#
# $$
# S_n = \{ n - 2k^2 : k = 1, 2, \dotsc, \lfloor \sqrt{\frac{n}{2}} \rfloor \}
# $$
#
# If any element $n - 2k^2$ of $S_n$ is prime, then we call $k$ a *witness* to Goldbach's Other Conjecture for $n$. We are asked to find the smallest $n$ that has no witness, i.e. such that *all* elements of $S_n$ are composite.
#
# Let $P_n$ be the set of all prime numbers stricly smaller than $n$, then our algorithm searches for the smallest $n$ such that $P_n \cap S_n = \emptyset$, providing a counterexample to Goldbach's Other Conjecture.
#
# <!-- TEASER_END -->
# +
from IPython.display import Math, display
from collections import defaultdict
from itertools import count
# -
# We modify and augment a very space-efficient implementation (due to <NAME>, UC Irvine) of the Sieve of Erastothenes to generate *composite* numbers and keep track of all prime numbers below it. In the outermost-loop below, `primes` will always be the list of all prime number strictly below $n$, i.e. it is equivalent to $P_n$. Note that if $n$ is odd and composite, then the largest prime below it is no greater than $n-2$, since $n-1$ is even. Since $\max S_n = n - 2$ searching through $P_n$ is sufficient (i.e. if $n-2$ is prime, then $n-2 \in P_n$.) The loop terminates when we encounter an $n$ with no witnesses.
factors = defaultdict(list)
witness = {}
primes = set()
for n in count(2):
if factors[n]:
# n is composite
for m in factors.pop(n):
factors[n+m].append(m)
if n % 2:
# n is odd and composite
for k in range(1, int((n/2)**.5)+1):
p = n - 2*k**2
if p in primes:
# TODO: `in` is O(len(primes))
# could optimize by using set
witness[n] = k
break
if not n in witness:
break
else:
factors[n*n].append(n)
primes.add(n)
n
# The answer is 5777.
# Note that not only is this implementation space-efficient, it is also very time efficient, since we incrementally build our list of primes, composites and witnesses incrementally from bottom-up. If we hadn't augmented the implementation of the prime sieve, we would have had to use the prime sieve to obtain all odd composites, and perform a primality test on all elements of $S_n$, which would (usually) require a prime factorization algorithm, which in turn (usually) requires a prime sieve, not to mention the fact that there would be many overlapping subproblems, i.e. many $n < m$ such that $S_n \cap S_m \ne \emptyset$, so we'd have to use dynamic programming, by, for example memoizing the primality testing function or some other optimization - just a whole mess of redundancies and inefficiencies that we happily avoided with this method :)
# Let's list out the witnesses to the first 100 numbers.
lines = [r'{0} &= {1} + 2 \cdot {2}^2 \\'.format(n, n - 2*witness[n]**2, witness[n]) for n in sorted(witness)]
Math(r"""
\begin{{align}}
{body}
\end{{align}}
""".format(body='\n'.join(lines[:100])))
| problem-46-goldbachs-other-conjecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 2: Intro to Private Training with Remote Execution
#
# In the last section, we learned about PointerTensors, which create the underlying infrastructure we need for privacy preserving Deep Learning. In this section, we're going to see how to use these basic tools to train our first deep learning model using remote execution.
#
# Authors:
# - <NAME> - Twitter: [@YannDupis](https://twitter.com/YannDupis)
# - <NAME> - Twitter: [@iamtrask](https://twitter.com/iamtrask)
#
# ### Why use remote execution?
#
# Let's say you are an AI startup who wants to build a deep learning model to detect [diabetic retinopathy (DR)](https://ai.googleblog.com/2016/11/deep-learning-for-detection-of-diabetic.html), which is the fastest growing cause of blindness. Before training your model, the first step would be to acquire a dataset of retinopathy images with signs of DR. One approach could be to work with a hospital and ask them to send you a copy of this dataset. However because of the sensitivity of the patients' data, the hospital might be exposed to liability risks.
#
#
# That's where remote execution comes into the picture. Instead of bringing training data to the model (a central server), you bring the model to the training data (wherever it may live). In this case, it would be the hospital.
#
# The idea is that this allows whoever is creating the data to own the only permanent copy, and thus maintain control over who ever has access to it. Pretty cool, eh?
# # Section 2.1 - Private Training on MNIST
#
# For this tutorial, we will train a model on the [MNIST dataset](http://yann.lecun.com/exdb/mnist/) to classify digits based on images.
#
# We can assume that we have a remote worker named Bob who owns the data.
# +
import tensorflow as tf
import syft as sy
hook = sy.TensorFlowHook(tf)
bob = sy.VirtualWorker(hook, id="bob")
# -
# Let's download the MNIST data from `tf.keras.datasets`. Note that we are converting the data from numpy to `tf.Tensor` in order to have the PySyft functionalities.
# +
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
x_train, y_train = tf.convert_to_tensor(x_train), tf.convert_to_tensor(y_train)
x_test, y_test = tf.convert_to_tensor(x_test), tf.convert_to_tensor(y_test)
# -
# As decribed in Part 1, we can send this data to Bob with the `send` method on the `tf.Tensor`.
x_train_ptr = x_train.send(bob)
y_train_ptr = y_train.send(bob)
# Excellent! We have everything to start experimenting. To train our model on Bob's machine, we just have to perform the following steps:
#
# - Define a model, including optimizer and loss
# - Send the model to Bob
# - Start the training process
# - Get the trained model back
#
# Let's do it!
# +
# Define the model
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
# Compile with optimizer, loss and metrics
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# -
# Once you have defined your model, you can simply send it to Bob calling the `send` method. It's the exact same process as sending a tensor.
model_ptr = model.send(bob)
model_ptr
# Now, we have a pointer pointing to the model on Bob's machine. We can validate that's the case by inspecting the attribute `_objects` on the virtual worker.
bob._objects[model_ptr.id_at_location]
# Everything is ready to start training our model on this remote dataset. You can call `fit` and pass `x_train_ptr` `y_train_ptr` which are pointing to Bob's data. Note that's the exact same interface as normal `tf.keras`.
model_ptr.fit(x_train_ptr, y_train_ptr, epochs=2, validation_split=0.2)
# Fantastic! you have trained your model acheiving an accuracy greater than 95%.
#
# You can get your trained model back by just calling `get` on it.
# +
model_gotten = model_ptr.get()
model_gotten
# -
# It's good practice to see if your model can generalize by assessing its accuracy on an holdout dataset. You can simply call `evaluate`.
model_gotten.evaluate(x_test, y_test, verbose=2)
# Boom! The model remotely trained on Bob's data is more than 95% accurate on this holdout dataset.
# If your model doesn't fit into the Sequential paradigm, you can use Keras's functional API, or even subclass [tf.keras.Model](https://www.tensorflow.org/guide/keras/custom_layers_and_models#building_models) to create custom models.
# +
class CustomModel(tf.keras.Model):
def __init__(self, num_classes=10):
super(CustomModel, self).__init__(name='custom_model')
self.num_classes = num_classes
self.flatten = tf.keras.layers.Flatten(input_shape=(28, 28))
self.dense_1 = tf.keras.layers.Dense(128, activation='relu')
self.dense_2 = tf.keras.layers.Dense(num_classes, activation='softmax')
def call(self, inputs):
x = self.flatten(inputs)
x = self.dense_1(x)
return self.dense_2(x)
model = CustomModel(10)
# need to call the model on dummy data before sending it
# in order to set the input shape (required when saving to SavedModel)
model.predict(tf.ones([1, 28, 28]))
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model_ptr = model.send(bob)
model_ptr.fit(x_train_ptr, y_train_ptr, epochs=2, validation_split=0.2)
# -
# ## Well Done!
#
# And voilà! We have trained a Deep Learning model on Bob's data by sending the model to him. Never in this process do we ever see or request access to the underlying training data! We preserve the privacy of Bob!!!
# # Congratulations!!! - Time to Join the Community!
#
# Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
#
# ### Star PySyft on GitHub
#
# The easiest way to help our community is just by starring the Repos! This helps raise awareness of the cool tools we're building.
#
# - Star PySyft on GitHub! - [https://github.com/OpenMined/PySyft](https://github.com/OpenMined/PySyft)
# - Star PySyft-TensorFlow on GitHub! - [https://github.com/OpenMined/PySyft-TensorFlow]
#
# ### Join our Slack!
#
# The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
#
# ### Join a Code Project!
#
# The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
#
# - [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
# - [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
#
# ### Donate
#
# If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
#
# [OpenMined's Open Collective Page](https://opencollective.com/openmined)
| examples/Part 02 - Intro to Private Training with Remote Execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="7765UFHoyGx6"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="KsOkK8O69PyT"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="ZS8z-_KeywY9"
# # Creating Keras Models with TFL Layers
# + [markdown] colab_type="text" id="r61fkA2i9Y3_"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/lattice/tutorials/keras_layers"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/lattice/blob/master/docs/tutorials/keras_layers.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/lattice/blob/master/docs/tutorials/keras_layers.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/lattice/docs/tutorials/keras_layers.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="ecLbJCvJSSCd"
# ##Overview
#
# You can use TFL Keras layers to construct Keras models with monotonicity and other shape constraints. This example builds and trains a calibrated lattice model for the UCI heart dataset using TFL layers.
#
# In a calibrated lattice model, each feature is transformed by a `tfl.layers.PWLCalibration` or a `tfl.layers.CategoricalCalibration` layer and the results are nonlinearly fused using a `tfl.layers.Lattice`.
# + [markdown] colab_type="text" id="x769lI12IZXB"
# ## Setup
# + [markdown] colab_type="text" id="fbBVAR6UeRN5"
# Installing TF Lattice package:
# + colab={} colab_type="code" id="bpXjJKpSd3j4"
#@test {"skip": true}
# !pip install tensorflow-lattice pydot
# + [markdown] colab_type="text" id="jSVl9SHTeSGX"
# Importing required packages:
# + cellView="both" colab={} colab_type="code" id="pm0LD8iyIZXF"
import tensorflow as tf
import logging
import numpy as np
import pandas as pd
import sys
import tensorflow_lattice as tfl
from tensorflow import feature_column as fc
logging.disable(sys.maxsize)
# + [markdown] colab_type="text" id="svPuM6QNxlrH"
# Downloading the UCI Statlog (Heart) dataset:
# + cellView="both" colab={} colab_type="code" id="PG3pFtK-IZXM"
# UCI Statlog (Heart) dataset.
csv_file = tf.keras.utils.get_file(
'heart.csv', 'http://storage.googleapis.com/applied-dl/heart.csv')
training_data_df = pd.read_csv(csv_file).sample(
frac=1.0, random_state=41).reset_index(drop=True)
training_data_df.head()
# + [markdown] colab_type="text" id="nKkAw12SxvGG"
# Setting the default values used for training in this guide:
# + cellView="both" colab={} colab_type="code" id="krAJBE-yIZXR"
LEARNING_RATE = 0.1
BATCH_SIZE = 128
NUM_EPOCHS = 100
# + [markdown] colab_type="text" id="0TGfzhPHzpix"
# ## Sequential Keras Model
#
# This example creates a Sequential Keras model and only uses TFL layers.
#
# Lattice layers expect `input[i]` to be within `[0, lattice_sizes[i] - 1.0]`, so we need to define the lattice sizes ahead of the calibration layers so we can properly specify output range of the calibration layers.
#
# + colab={} colab_type="code" id="nOQWqPAbQS3o"
# Lattice layer expects input[i] to be within [0, lattice_sizes[i] - 1.0], so
lattice_sizes = [3, 2, 2, 2, 2, 2, 2]
# + [markdown] colab_type="text" id="W3DnEKWvQYXm"
# We use a `tfl.layers.ParallelCombination` layer to group together calibration layers which have to be executed in paralel in order to be able to create a Sequential model.
#
# + colab={} colab_type="code" id="o_hyk5GkQfl8"
combined_calibrators = tfl.layers.ParallelCombination()
# + [markdown] colab_type="text" id="BPZsSUZiQiwc"
# We create a calibration layer for each feature and add it to the parallel combination layer. For numeric features we use `tfl.layers.PWLCalibration` and for categorical features we use `tfl.layers.CategoricalCalibration`.
# + colab={} colab_type="code" id="DXPc6rSGxzFZ"
# ############### age ###############
calibrator = tfl.layers.PWLCalibration(
# Every PWLCalibration layer must have keypoints of piecewise linear
# function specified. Easiest way to specify them is to uniformly cover
# entire input range by using numpy.linspace().
input_keypoints=np.linspace(
training_data_df['age'].min(), training_data_df['age'].max(), num=5),
# You need to ensure that input keypoints have same dtype as layer input.
# You can do it by setting dtype here or by providing keypoints in such
# format which will be converted to deisred tf.dtype by default.
dtype=tf.float32,
# Output range must correspond to expected lattice input range.
output_min=0.0,
output_max=lattice_sizes[0] - 1.0,
)
combined_calibrators.append(calibrator)
# ############### sex ###############
# For boolean features simply specify CategoricalCalibration layer with 2
# buckets.
calibrator = tfl.layers.CategoricalCalibration(
num_buckets=2,
output_min=0.0,
output_max=lattice_sizes[1] - 1.0,
# Initializes all outputs to (output_min + output_max) / 2.0.
kernel_initializer='constant')
combined_calibrators.append(calibrator)
# # ############### cp ###############
calibrator = tfl.layers.PWLCalibration(
# Here instead of specifying dtype of layer we convert keypoints into
# np.float32.
input_keypoints=np.linspace(1, 4, num=4, dtype=np.float32),
output_min=0.0,
output_max=lattice_sizes[2] - 1.0,
monotonicity='increasing',
# You can specify TFL regularizers as a tuple ('regularizer name', l1, l2).
kernel_regularizer=('hessian', 0.0, 1e-4))
combined_calibrators.append(calibrator)
# ############### trestbps ###############
calibrator = tfl.layers.PWLCalibration(
# Alternatively, you might want to use quantiles as keypoints instead of
# uniform keypoints
input_keypoints=np.quantile(training_data_df['trestbps'],
np.linspace(0.0, 1.0, num=5)),
dtype=tf.float32,
# Together with quantile keypoints you might want to initialize piecewise
# linear function to have 'equal_slopes' in order for output of layer
# after initialization to preserve original distribution.
kernel_initializer='equal_slopes',
output_min=0.0,
output_max=lattice_sizes[3] - 1.0,
# You might consider clamping extreme inputs of the calibrator to output
# bounds.
clamp_min=True,
clamp_max=True,
monotonicity='increasing')
combined_calibrators.append(calibrator)
# ############### chol ###############
calibrator = tfl.layers.PWLCalibration(
# Explicit input keypoint initialization.
input_keypoints=[126.0, 210.0, 247.0, 286.0, 564.0],
dtype=tf.float32,
output_min=0.0,
output_max=lattice_sizes[4] - 1.0,
# Monotonicity of calibrator can be decreasing. Note that corresponding
# lattice dimension must have INCREASING monotonicity regardless of
# monotonicity direction of calibrator.
monotonicity='decreasing',
# Convexity together with decreasing monotonicity result in diminishing
# return constraint.
convexity='convex',
# You can specify list of regularizers. You are not limited to TFL
# regularizrs. Feel free to use any :)
kernel_regularizer=[('laplacian', 0.0, 1e-4),
tf.keras.regularizers.l1_l2(l1=0.001)])
combined_calibrators.append(calibrator)
# ############### fbs ###############
calibrator = tfl.layers.CategoricalCalibration(
num_buckets=2,
output_min=0.0,
output_max=lattice_sizes[5] - 1.0,
# For categorical calibration layer monotonicity is specified for pairs
# of indices of categories. Output for first category in pair will be
# smaller than output for second category.
#
# Don't forget to set monotonicity of corresponding dimension of Lattice
# layer to '1'.
monotonicities=[(0, 1)],
# This initializer is identical to default one('uniform'), but has fixed
# seed in order to simplify experimentation.
kernel_initializer=tf.keras.initializers.RandomUniform(
minval=0.0, maxval=lattice_sizes[5] - 1.0, seed=1))
combined_calibrators.append(calibrator)
# ############### restecg ###############
calibrator = tfl.layers.CategoricalCalibration(
num_buckets=3,
output_min=0.0,
output_max=lattice_sizes[6] - 1.0,
# Categorical monotonicity can be partial order.
monotonicities=[(0, 1), (0, 2)],
# Categorical calibration layer supports standard Keras regularizers.
kernel_regularizer=tf.keras.regularizers.l1_l2(l1=0.001),
kernel_initializer='constant')
combined_calibrators.append(calibrator)
# + [markdown] colab_type="text" id="inyNlSBeQyp7"
# We then create a lattice layer to nonlinearly fuse the outputs of the calibrators.
#
# Note that we need to specify the monotonicity of the lattice to be increasing for required dimensions. The composition with the direction of the monotonicity in the calibration will result in the correct end-to-end direction of monotonicity. This includes partial monotonicity of CategoricalCalibration layer.
# + colab={} colab_type="code" id="DNCc9oBTRo6w"
lattice = tfl.layers.Lattice(
lattice_sizes=lattice_sizes,
monotonicities=[
'increasing', 'none', 'increasing', 'increasing', 'increasing',
'increasing', 'increasing'
],
output_min=0.0,
output_max=1.0)
# + [markdown] colab_type="text" id="T5q2InayRpDr"
# We can then create a sequential model using the combined calibrators and lattice layers.
# + colab={} colab_type="code" id="xX6lroYZQy3L"
model = tf.keras.models.Sequential()
model.add(combined_calibrators)
model.add(lattice)
# + [markdown] colab_type="text" id="W3UFxD3fRzIC"
# Training works the same as any other keras model.
# + colab={} colab_type="code" id="2jz4JvI-RzSj"
features = training_data_df[[
'age', 'sex', 'cp', 'trestbps', 'chol', 'fbs', 'restecg'
]].values.astype(np.float32)
target = training_data_df[['target']].values.astype(np.float32)
model.compile(
loss=tf.keras.losses.mean_squared_error,
optimizer=tf.keras.optimizers.Adagrad(learning_rate=LEARNING_RATE))
model.fit(
features,
target,
batch_size=BATCH_SIZE,
epochs=NUM_EPOCHS,
validation_split=0.2,
shuffle=False,
verbose=0)
model.evaluate(features, target)
# + [markdown] colab_type="text" id="RTHoW_5lxwT5"
# ## Functional Keras Model
#
# This example uses a functional API for Keras model construction.
#
# As mentioned in the previous section, lattice layers expect `input[i]` to be within `[0, lattice_sizes[i] - 1.0]`, so we need to define the lattice sizes ahead of the calibration layers so we can properly specify output range of the calibration layers.
# + colab={} colab_type="code" id="gJjUYvBuW1qE"
# We are going to have 2-d embedding as one of lattice inputs.
lattice_sizes = [3, 2, 2, 3, 3, 2, 2]
# + [markdown] colab_type="text" id="Z03qY5MYW1yT"
# For each feature, we need to create an input layer followed by a calibration layer. For numeric features we use `tfl.layers.PWLCalibration` and for categorical features we use `tfl.layers.CategoricalCalibration`.
# + colab={} colab_type="code" id="DCIUz8apzs0l"
model_inputs = []
lattice_inputs = []
# ############### age ###############
age_input = tf.keras.layers.Input(shape=[1], name='age')
model_inputs.append(age_input)
age_calibrator = tfl.layers.PWLCalibration(
# Every PWLCalibration layer must have keypoints of piecewise linear
# function specified. Easiest way to specify them is to uniformly cover
# entire input range by using numpy.linspace().
input_keypoints=np.linspace(
training_data_df['age'].min(), training_data_df['age'].max(), num=5),
# You need to ensure that input keypoints have same dtype as layer input.
# You can do it by setting dtype here or by providing keypoints in such
# format which will be converted to deisred tf.dtype by default.
dtype=tf.float32,
# Output range must correspond to expected lattice input range.
output_min=0.0,
output_max=lattice_sizes[0] - 1.0,
monotonicity='increasing',
name='age_calib',
)(
age_input)
lattice_inputs.append(age_calibrator)
# ############### sex ###############
# For boolean features simply specify CategoricalCalibration layer with 2
# buckets.
sex_input = tf.keras.layers.Input(shape=[1], name='sex')
model_inputs.append(sex_input)
sex_calibrator = tfl.layers.CategoricalCalibration(
num_buckets=2,
output_min=0.0,
output_max=lattice_sizes[1] - 1.0,
# Initializes all outputs to (output_min + output_max) / 2.0.
kernel_initializer='constant',
name='sex_calib',
)(
sex_input)
lattice_inputs.append(sex_calibrator)
# # ############### cp ###############
cp_input = tf.keras.layers.Input(shape=[1], name='cp')
model_inputs.append(cp_input)
cp_calibrator = tfl.layers.PWLCalibration(
# Here instead of specifying dtype of layer we convert keypoints into
# np.float32.
input_keypoints=np.linspace(1, 4, num=4, dtype=np.float32),
output_min=0.0,
output_max=lattice_sizes[2] - 1.0,
monotonicity='increasing',
# You can specify TFL regularizers as tuple ('regularizer name', l1, l2).
kernel_regularizer=('hessian', 0.0, 1e-4),
name='cp_calib',
)(
cp_input)
lattice_inputs.append(cp_calibrator)
# ############### trestbps ###############
trestbps_input = tf.keras.layers.Input(shape=[1], name='trestbps')
model_inputs.append(trestbps_input)
trestbps_calibrator = tfl.layers.PWLCalibration(
# Alternatively, you might want to use quantiles as keypoints instead of
# uniform keypoints
input_keypoints=np.quantile(training_data_df['trestbps'],
np.linspace(0.0, 1.0, num=5)),
dtype=tf.float32,
# Together with quantile keypoints you might want to initialize piecewise
# linear function to have 'equal_slopes' in order for output of layer
# after initialization to preserve original distribution.
kernel_initializer='equal_slopes',
output_min=0.0,
output_max=lattice_sizes[3] - 1.0,
# You might consider clamping extreme inputs of the calibrator to output
# bounds.
clamp_min=True,
clamp_max=True,
monotonicity='increasing',
name='trestbps_calib',
)(
trestbps_input)
lattice_inputs.append(trestbps_calibrator)
# ############### chol ###############
chol_input = tf.keras.layers.Input(shape=[1], name='chol')
model_inputs.append(chol_input)
chol_calibrator = tfl.layers.PWLCalibration(
# Explicit input keypoint initialization.
input_keypoints=[126.0, 210.0, 247.0, 286.0, 564.0],
output_min=0.0,
output_max=lattice_sizes[4] - 1.0,
# Monotonicity of calibrator can be decreasing. Note that corresponding
# lattice dimension must have INCREASING monotonicity regardless of
# monotonicity direction of calibrator.
monotonicity='decreasing',
# Convexity together with decreasing monotonicity result in diminishing
# return constraint.
convexity='convex',
# You can specify list of regularizers. You are not limited to TFL
# regularizrs. Feel free to use any :)
kernel_regularizer=[('laplacian', 0.0, 1e-4),
tf.keras.regularizers.l1_l2(l1=0.001)],
name='chol_calib',
)(
chol_input)
lattice_inputs.append(chol_calibrator)
# ############### fbs ###############
fbs_input = tf.keras.layers.Input(shape=[1], name='fbs')
model_inputs.append(fbs_input)
fbs_calibrator = tfl.layers.CategoricalCalibration(
num_buckets=2,
output_min=0.0,
output_max=lattice_sizes[5] - 1.0,
# For categorical calibration layer monotonicity is specified for pairs
# of indices of categories. Output for first category in pair will be
# smaller than output for second category.
#
# Don't forget to set monotonicity of corresponding dimension of Lattice
# layer to '1'.
monotonicities=[(0, 1)],
# This initializer is identical to default one ('uniform'), but has fixed
# seed in order to simplify experimentation.
kernel_initializer=tf.keras.initializers.RandomUniform(
minval=0.0, maxval=lattice_sizes[5] - 1.0, seed=1),
name='fbs_calib',
)(
fbs_input)
lattice_inputs.append(fbs_calibrator)
# ############### restecg ###############
restecg_input = tf.keras.layers.Input(shape=[1], name='restecg')
model_inputs.append(restecg_input)
restecg_calibrator = tfl.layers.CategoricalCalibration(
num_buckets=3,
output_min=0.0,
output_max=lattice_sizes[6] - 1.0,
# Categorical monotonicity can be partial order.
monotonicities=[(0, 1), (0, 2)],
# Categorical calibration layer supports standard Keras regularizers.
kernel_regularizer=tf.keras.regularizers.l1_l2(l1=0.001),
kernel_initializer='constant',
name='restecg_calib',
)(
restecg_input)
lattice_inputs.append(restecg_calibrator)
# + [markdown] colab_type="text" id="Fr0k8La_YgQG"
# We then create a lattice layer to nonlinearly fuse the outputs of the calibrators.
#
# Note that we need to specify the monotonicity of the lattice to be increasing for required dimensions. The composition with the direction of the monotonicity in the calibration will result in the correct end-to-end direction of monotonicity. This includes partial monotonicity of `tfl.layers.CategoricalCalibration` layer.
# + colab={} colab_type="code" id="X15RE0NybNbU"
lattice = tfl.layers.Lattice(
lattice_sizes=lattice_sizes,
monotonicities=[
'increasing', 'none', 'increasing', 'increasing', 'increasing',
'increasing', 'increasing'
],
output_min=0.0,
output_max=1.0,
name='lattice',
)(
lattice_inputs)
# + [markdown] colab_type="text" id="31VzsnMCA9dh"
# To add more flexibility to the model, we add an output calibration layer.
# + colab={} colab_type="code" id="efCP3Yx2A9n7"
model_output = tfl.layers.PWLCalibration(
input_keypoints=np.linspace(0.0, 1.0, 5),
name='output_calib',
)(
lattice)
# + [markdown] colab_type="text" id="1SURnNl8bNgw"
# We can now create a model using the inputs and outputs.
# + colab={} colab_type="code" id="7gY-VXuYbZLa"
model = tf.keras.models.Model(
inputs=model_inputs,
outputs=model_output)
tf.keras.utils.plot_model(model, rankdir='LR')
# + [markdown] colab_type="text" id="tvFJTs94bZXK"
# Training works the same as any other keras model. Note that, with our setup, input features are passed as separate tensors.
# + colab={} colab_type="code" id="vMQTGbFAYgYS"
feature_names = ['age', 'sex', 'cp', 'trestbps', 'chol', 'fbs', 'restecg']
features = np.split(
training_data_df[feature_names].values.astype(np.float32),
indices_or_sections=len(feature_names),
axis=1)
target = training_data_df[['target']].values.astype(np.float32)
model.compile(
loss=tf.keras.losses.mean_squared_error,
optimizer=tf.keras.optimizers.Adagrad(LEARNING_RATE))
model.fit(
features,
target,
batch_size=BATCH_SIZE,
epochs=NUM_EPOCHS,
validation_split=0.2,
shuffle=False,
verbose=0)
model.evaluate(features, target)
| site/en-snapshot/lattice/tutorials/keras_layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Chart Overlays {#chart_overlays_example}
# ==============
#
# This example shows how you can combine multiple charts as overlays in
# the same renderer. For an overview of the different chart types you can
# use, please refer to `chart_basics_example`{.interpreted-text
# role="ref"}. Interaction with a chart can be enabled by a double left
# click on top of it. Note that this will disable interaction with the 3D
# scene. To stop interacting with the chart, perform another double left
# click. This will either enable interaction with another chart (if
# clicked on top of it) or re-enable interaction with the 3D scene.
#
# +
import matplotlib.pyplot as plt
import numpy as np
import pyvista as pv
# -
# Data to display
#
t = np.linspace(0, 5, 50)
h = np.sin(t)
v = np.cos(t)
# Define a Matplotlib figure. Use a tight layout to keep axis labels
# visible on smaller figures.
#
f, ax = plt.subplots(tight_layout=True)
h_line = ax.plot(t[:1], h[:1])[0]
ax.set_ylim([-1, 1])
ax.set_xlabel('Time (s)')
_ = ax.set_ylabel('Height (m)')
# Define plotter, add the created matplotlib figure as the first (left)
# chart to the scene, and define a second (right) chart.
#
# +
p = pv.Plotter()
h_chart = pv.ChartMPL(f, size=(0.46, 0.25), loc=(0.02, 0.06))
h_chart.background_color = (1.0, 1.0, 1.0, 0.4)
p.add_chart(h_chart)
v_chart = pv.Chart2D(
size=(0.46, 0.25), loc=(0.52, 0.06), x_label="Time (s)", y_label="Velocity (m/s)"
)
v_line = v_chart.line(t[:1], v[:1])
v_chart.y_range = (-1, 1)
v_chart.background_color = (1.0, 1.0, 1.0, 0.4)
p.add_chart(v_chart)
p.add_mesh(pv.Sphere(1), name="sphere", render=False)
p.show(auto_close=False, interactive=True, interactive_update=True)
# Method and slider to update all visuals based on the time selection
def update_time(time):
k = np.count_nonzero(t < time)
h_line.set_xdata(t[: k + 1])
h_line.set_ydata(h[: k + 1])
v_line.update(t[: k + 1], v[: k + 1])
p.add_mesh(pv.Sphere(1, center=(0, 0, h[k])), name="sphere", render=False)
p.update()
time_slider = p.add_slider_widget(
update_time, [np.min(t), np.max(t)], 0, "Time", (0.25, 0.9), (0.75, 0.9), event_type='always'
)
# Start incrementing time automatically
for i in range(1, 50):
ax.set_xlim([0, t[i]])
time_slider.GetSliderRepresentation().SetValue(t[i])
update_time(t[i])
p.show() # Keep plotter open to let user play with time slider
| examples/02-plot/chart_overlays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('/Users/keemsunguk/Projects/EssayGrader/')
import spacy
from spacy import displacy
import json
import re
from egrader.preprocess import Preprocess
from egrader.db_util import DBUtil
db_util = DBUtil(local_db=True)
db_util.describe_db()
short_essay = [e['recno'] for e in db_util.remote_ec.find({}) if len(e['essay']) < 2]
db_util.remote_ec.delete_many({'recno':{'$in':short_essay}})
db_util.local_ec.delete_many({'essay': {'$eq':None}})
db_util.remote_ec.find_one({}).keys()
txt = db_util.remote_ec.find_one({})['essay']
tmp = re.finditer('\\r\\n\\r\\n', txt)
pp = Preprocess("")
pp.clean_html()
toefl_df = db_util.get_spacy_labeled_essays('TOEFL', merge_0_1=False, with_topic=True)
# + jupyter={"outputs_hidden": true}
for r, v in toefl_df.iterrows():
toefl_df.iloc[r, 0] = pp.clean_html(v[0])
print(toefl_df.iloc[r, 0])
# -
topic_doc = pp.make_doc_obj(toefl_df.iloc[1, 0])
displacy.render(topic_doc, style="ent", jupyter=True)
noun_chunks = list(topic_doc.noun_chunks)
topic_doc.vector[1]
essay_doc = pp.make_doc_obj(toefl_df.iloc[1, 1])
displacy.render(essay_doc, style="ent", jupyter=True)
essay_doc.similarity(topic_doc)
# + jupyter={"outputs_hidden": true}
topic_doc.to_json()
# -
train_data = [(v[0], v[1]) for k, v in toefl_df.iterrows()]
from collections import Counter
Counter(list(toefl_df[2].values))
a = json.dumps(db_util.remote_ec.find_one({}, {'_id':0}))
a
| notebooks/toefl_grader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:opencv]
# language: python
# name: conda-env-opencv-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Astra Zeneca Case Study:
# ## exploring adverse drug events reported in OpenFDA database
# + [markdown] slideshow={"slide_type": "skip"}
# At interview you should expect to discuss your code, any statistics or visualizations you may have used, limitations of the underlying data, and how your solution could be generalized, extended, and made into a robust product.
#
# + slideshow={"slide_type": "skip"}
import pandas as pd
import json
import requests
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
# + [markdown] slideshow={"slide_type": "slide"}
# ## Are different adverse events reported in different countries?
# + slideshow={"slide_type": "subslide"}
#get all patient reactions
URL = "https://api.fda.gov/drug/event.json?count=patient.reaction.reactionmeddrapt.exact"
data = requests.get(URL).json()
rxn_counts = pd.DataFrame(data.get("results"))
rxn_counts.shape
# + slideshow={"slide_type": "subslide"}
rxn_counts.head(2)
# + slideshow={"slide_type": "subslide"}
#get all country counts
URL = "https://api.fda.gov/drug/event.json?count=occurcountry.exact"
data = requests.get(URL).json()
country = pd.DataFrame(data.get("results"))
country.shape
# + slideshow={"slide_type": "subslide"}
# list of reactions
rxns = []
for i in rxn_counts.term:
rxns.append(i.replace(" ", "+"))
# create list of URLS to get reaction counts for all countries
URL = "https://api.fda.gov/drug/event.json?search=patient.reaction.reactionmeddrapt:XXX&count=occurcountry.exact"
URL_x=[]
for i in rxns:
URL_x.append(URL.replace("XXX", i))
# download data and store in dataframe
df_append = []
for url in URL_x:
data = requests.get(url).json()
df = pd.DataFrame(data.get("results"))
df.set_index("term", drop=True, inplace=True)
df_append.append(df)
# + slideshow={"slide_type": "subslide"}
# concatenate all diseases by country
final_df = pd.concat(df_append, axis = 1)
final_df.columns = rxns
final_df.shape
# + slideshow={"slide_type": "subslide"}
final_df.head(3)
# + slideshow={"slide_type": "subslide"}
final_df.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### NULL VALUES DISTRIBUTION
# + slideshow={"slide_type": "subslide"}
x = (final_df.isnull().sum(axis=0)/final_df.shape[0])
x.plot(kind= "kde", figsize = (25,5), fontsize = 12)
plt.title("null value distribution by adverse event")
plt.show()
# + slideshow={"slide_type": "subslide"}
"""
x = (final_df.isnull().sum(axis=0)/final_df.shape[0])
x.plot(kind= "bar", figsize = (25,5), fontsize = 8)
plt.title("null value distribution by adverse event")
plt.show()
"""
# + slideshow={"slide_type": "subslide"}
x = (final_df.isnull().sum(axis=1)/final_df.shape[1])
x.plot(kind= "kde", figsize = (25,5), fontsize = 12)
plt.title("null value distribution by country")
plt.show()
# + slideshow={"slide_type": "subslide"}
x = (final_df.isnull().sum(axis=1)/final_df.shape[1])
x.sort_values(ascending=True).plot(kind= "bar", figsize = (25,8), fontsize = 10)
plt.title("null value distribution by country")
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# #### Number of adverse events reported (by country)
# + slideshow={"slide_type": "subslide"}
final_df.sum(axis=1).plot(kind="bar", figsize = (30,6))
plt.title("Counts of adverse events reported (by country)")
plt.show()
# the US is definitely OVER REPRESENTED!
# + [markdown] slideshow={"slide_type": "slide"}
# ##### Number of different adverse events reported by country (out of a total of 100 different possible events)
# + slideshow={"slide_type": "subslide"}
final_df.transpose().describe().loc["count"].sort_values().plot(kind="bar", figsize = (32,12), fontsize= 11)
plt.title("Number of different adverse events reported (by country)")
plt.show()
# + slideshow={"slide_type": "subslide"}
final_df.transpose().describe().loc["count"].plot(kind="hist", figsize = (15,3))
plt.title("Number of different adverse events reported (by country)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# grouping countries into continents
continents = pd.read_csv("./data/continents.csv")
continent_dict = {}
for i in continents.columns:
k = list(continents[i].dropna())
continent_dict.update(dict.fromkeys(k, i))
continent_df = final_df.copy()
continent_df["continent"] = [continent_dict.get(i) for i in final_df.index]
continent_df.head(3)
# + slideshow={"slide_type": "subslide"}
continent_df.groupby("continent").agg("sum").sum(axis=1).plot(kind="barh", figsize = (25,8), fontsize= 12)
plt.title("Number of different adverse events reported (by continent)")
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### Most common adverse events
# + slideshow={"slide_type": "subslide"}
common_by_country = final_df.idxmax(axis=1)
top_disease = common_by_country.value_counts()[:25].index
# + slideshow={"slide_type": "subslide"}
common_by_country.value_counts()[:25].plot(kind="barh", figsize=(20,6))
plt.title("Most commonly reported adverse reactions across all countries")
# by number of countries which report the specific adverse event as their most common one
plt.show()
# + slideshow={"slide_type": "subslide"}
final_df.sum().sort_values(ascending=False).plot(kind = "bar", figsize= (25,10), fontsize=12)
plt.title("raw counts of adverse events across all countries")
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# #### Top 10 most common adverse events (by top 25 countries with most reported adverse events)
# + slideshow={"slide_type": "subslide"}
# Top 10 most common adverse events (by top 25 countries with most reported adverse events)
most_reported_events_by_country = final_df.sum(axis=1).sort_values(ascending=False)
top_countries = most_reported_events_by_country.index[:25]
final_df.loc[top_countries][top_disease[:10]].plot(kind="bar", figsize=(25,12), stacked = True)
plt.title("adverse events by top 25 most reporting countries (incl US)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# Top 10 most common adverse events (by top 25 countries with most reported adverse events)
most_reported_events_by_country = final_df.sum(axis=1).sort_values(ascending=False)
top_countries = most_reported_events_by_country.index[:25]
final_df.loc[top_countries][top_disease[:10]].drop("US", axis=0).plot(kind="bar", figsize=(25,12), stacked = True)
plt.title("top 10 adverse events by top 25 most reporting countries (excluding US)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# NORMALISED - Top 10 most common adverse events (by top 25 countries with most reported adverse events)
norm_df = final_df.divide(final_df.sum(axis=1), axis=0)
norm_df.loc[top_countries][top_disease[:10]].plot(kind="bar", figsize=(25,15), stacked = True)
plt.title("normalised counts of adverse events by top 25 most reporting countries (incl US)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# NORMALISED - Top 10 most common adverse events (by continent)
continent_df.groupby("continent").agg("sum")[top_disease[:10]].plot(kind="barh", figsize=(30,15),
stacked = True, fontsize = 15)
plt.title("counts of adverse events by top 25 most reporting countries (incl US)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# NORMALISED - Top 10 most common adverse events (by continent)
continent_counts = continent_df.groupby("continent").agg("sum")
norm_continents = continent_counts.divide(continent_counts.sum(axis=1), axis=0)
norm_continents[top_disease[:10]].plot(kind="barh", figsize=(30,15), stacked = True)
plt.title("normalised counts of adverse events by continent")
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Who is reporting adverse drug reactions?
# + slideshow={"slide_type": "subslide"}
#get reporter data by country
URL = "https://api.fda.gov/drug/event.json?search=primarysource.qualification:XXX&count=occurcountry.exact"
URL_x=[]
for i in range(1,6):
URL_x.append(URL.replace("XXX", str(i)))
# download data and store in dataframe
df_append = []
for url in URL_x:
data = requests.get(url).json()
df = pd.DataFrame(data.get("results"))
df.set_index("term", drop=True, inplace=True)
df_append.append(df)
# + slideshow={"slide_type": "subslide"}
# concatenate
reporter_dict = {1:"Physician", 2:"Pharmacist", 3:"Other health professional", 4:"Lawyer", 5:"Consumer or non-health professional"}
reporter_df = pd.concat(df_append, axis = 1)
reporter_df.columns = [reporter_dict.get(i) for i in range(1,6)]
reporter_df.shape
# + slideshow={"slide_type": "subslide"}
# Reporters of adverse events (incl US)
reporter_df.loc[top_countries].plot(kind="bar", figsize=(25,12), stacked = True)
plt.title("Reporters of adverse events (incl US)")
plt.show()
# + slideshow={"slide_type": "subslide"}
# Reporters of adverse events (excluding US)
reporter_df.loc[top_countries].drop("US", axis=0).plot(kind="bar", figsize=(25,12), stacked = True)
plt.title("Reporters of adverse events (excluding US)")
plt.show()
# +
# NORMALISED - Reporters of adverse events (excluding US)
norm_reporter_df = reporter_df.divide(reporter_df.sum(axis=1), axis=0)
norm_reporter_df.loc[top_countries].plot(kind="bar", figsize=(25,12), stacked = True)
plt.title("NORMALISED - Reporters of adverse events (excluding US)")
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Who is reporting off-label use?
# + slideshow={"slide_type": "subslide"}
#get off label reporter data
URL = "https://api.fda.gov/drug/event.json?search=patient.reaction.reactionmeddrapt.exact:OFF+LABEL+USE&count=primarysource.qualification"
off_label_data = requests.get(URL).json()
# + slideshow={"slide_type": "subslide"}
off_label_df = pd.DataFrame(off_label_data.get("results"))
reporter_dict = {1:"Physician", 2:"Pharmacist", 3:"Other health professional", 4:"Lawyer", 5:"Consumer or non-health professional"}
off_label_df["reporter"] = [reporter_dict.get(i) for i in off_label_df.term]
off_label_df.drop(["term"], axis=1, inplace=True)
# + slideshow={"slide_type": "subslide"}
off_label_df.set_index("reporter", drop=True).plot(kind="barh", figsize = (25,5), fontsize=12)
plt.show()
# -
# ### Serious events
# ##### serious event: adverse event resulted in death, a life threatening condition, hospitalization, disability, congenital anomaly, or other serious condition
# +
# list of reactions
rxns = []
for i in rxn_counts.term:
rxns.append(i.replace(" ", "+"))
# create list of URLS to get reaction counts for all countries
URL = "https://api.fda.gov/drug/event.json?search=patient.reaction.reactionmeddrapt:XXX&count=serious"
URL_x=[]
for i in rxns:
URL_x.append(URL.replace("XXX", i))
# download data and store in dataframe
df_append = []
for url in URL_x:
data = requests.get(url).json()
df = pd.DataFrame(data.get("results"))
df.set_index("term", drop=True, inplace=True)
df_append.append(df)
# -
serious_df = pd.concat(df_append, axis=1)
serious_df.columns = rxns
serious_df.shape
norm_serious_df = serious_df.divide(serious_df.sum(axis=1), axis=0)
norm_serious_df.T.sort_values(by=1, ascending=False).iloc[:25,0].plot(kind="bar", figsize = (25,5))
plt.show()
# ### GDP
# +
gdp = pd.read_csv("./data/gdp_data.csv", usecols = ["Country Code", "IncomeGroup", "TableName"])
cc = pd.read_csv("./data/country_codes.csv")
cc.columns = ["TableName","code"]
cc_df = cc.merge(gdp, on="TableName")
# -
gdp_df = final_df.merge(cc_df, left_index=True, right_on = "code").drop(["Country Code", "TableName"],axis=1)
#.set_index("code", drop=True)
g = gdp_df.groupby(["IncomeGroup"]).sum()[top_disease[:10]]
norm_g = g.divide(g.sum(axis=1), axis=0)
norm_g.plot(kind="bar", figsize=(15, 8), stacked=False)
plt.legend(fontsize=8, loc = 1)
plt.title("top 10 most common adverse events by GDP")
plt.show()
| AZ_case_study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <table>
# <tr align=left><td><img align=left src="./images/CC-BY.png">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) <NAME></td>
# </table>
# + slideshow={"slide_type": "skip"}
from __future__ import print_function
from __future__ import absolute_import
# %matplotlib inline
import numpy
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "slide"}
# # Numerical Differentiation
#
# **GOAL:** Given a set of $N+1$ points $(x_i, y_i)$ compute the derivative of a given order to a specified accuracy.
#
# **Approach:** Find the interpolating polynomial $P_N(x)$ and differentiate that.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Newton's Form
#
# For ease of analysis we will write $P_N(x)$ in Newton's form which looks like
#
# $$P_N(x) = \sum^N_{j=0} a_j n_j(x)$$
#
# where
#
# $$n_j(x) = \prod^{j-1}_{i=0} (x - x_i)$$
# + [markdown] slideshow={"slide_type": "subslide"}
# The $a_j = [y_0, \ldots, y_j]$ are the divided differences defined in general as
#
# $$[y_i] = y_i \quad i \in \{0,\ldots, N+1\}$$
#
# and
#
# $$[y_i, \ldots , y_{i+j}] = \frac{[y_{i+1}, \ldots , y_{i + j}] - [y_{i},\ldots,y_{i+j-1}]}{x_{i+j} - x_{i}} \quad i \in \{0,\ldots,N+1 - j\} \quad j \in \{1,\ldots, N+1\}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# These formulas are recursively defined but not so helpful, here are a few examples to start out with:
#
# $$[y_0] = y_0$$
#
# $$[y_0, y_1] = \frac{y_1 - y_0}{x_1 - x_0}$$
#
# $$[y_0, y_1, y_2] = \frac{[y_1, y_2] - [y_0, y_1]}{x_{2} - x_{0}} = \frac{\frac{y_2 - y_1}{x_2 - x_1} - \frac{y_1 - y_0}{x_1 - x_0}}{x_2 - x_0} = \frac{y_2 - y_1}{(x_2 - x_1)(x_2 - x_0)} - \frac{y_1 - y_0}{(x_1 - x_0)(x_2 - x_0)}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# The benefit of writing a polynomial like this is that it isolates the $x$ dependence (we can easily take derivatives of this form).
#
# In general then $P_N(x)$ can be written in Newton's form as
#
# $$P_N(x) = y_0 + (x-x_0)[y_0, y_1] + (x - x_0) (x - x_1) [y_0, y_1, y_2] + \cdots + (x-x_0) (x-x_1) \cdots (x-x_{N-1}) [y_0, y_1, \ldots, y_{N}]$$
# + [markdown] slideshow={"slide_type": "subslide"}
# As another concrete example consider and quadratic polynomial written in Newton's form
#
# $$P_2(x) = [y_0] + (x - x_0) [y_0, y_1] + (x - x_0)(x - x_1) [y_0, y_1, y_2] \\= y_0 + (x - x_0) \frac{y_1 - y_0}{x_1 - x_0} + (x - x_0)(x - x_1) \left ( \frac{y_2 - y_1}{(x_2 - x_1)(x_2 - x_0)} - \frac{y_1 - y_0}{(x_1 - x_0)(x_2 - x_0)} \right )$$
#
# Recall that the interpolating polynomial of degree $N$ through these points is unique!
# + slideshow={"slide_type": "skip"}
def divided_difference(x, y, N=50):
"""Compute the Nth divided difference using *x* and *y*"""
if N == 0:
raise Exception("Reached recursion limit!")
# Reached the end of the recurssion
if y.shape[0] == 1:
return y[0]
elif y.shape[0] == 2:
return (y[1] - y[0]) / (x[1] - x[0])
else:
return (divided_difference(x[1:], y[1:], N=N-1) - divided_difference(x[:-1], y[:-1], N=N-1)) / (x[-1] - x[0])
# Calculate a polynomial in Newton Form
data = numpy.array([[-2.0, 1.0], [-1.5, -1.0], [-0.5, -3.0], [0.0, -2.0], [1.0, 3.0], [2.0, 1.0]])
N = data.shape[0] - 1
x = numpy.linspace(-2.0, 2.0, 100)
# Construct basis functions
newton_basis = numpy.ones((N + 1, x.shape[0]))
for j in range(N + 1):
for i in range(j):
newton_basis[j, :] *= (x - data[i, 0])
# Construct full polynomial
P = numpy.zeros(x.shape)
for j in range(N + 1):
P += divided_difference(data[:j + 1, 0], data[:j + 1, 1]) * newton_basis[j, :]
# Plot basis and interpolant
fig = plt.figure()
fig.subplots_adjust(hspace=.5)
axes = [None, None]
axes[0] = fig.add_subplot(2, 1, 1)
axes[1] = fig.add_subplot(2, 1, 2)
for j in range(N + 1):
axes[0].plot(x, newton_basis[j, :])
axes[1].plot(data[j, 0], data[j, 1],'ko')
axes[1].plot(x, P)
axes[0].set_title("Newton Polynomial Basis")
axes[0].set_xlabel("x")
axes[0].set_ylabel("$n_j(x)$")
axes[1].set_title("Interpolant $P_%s(x)$" % N)
axes[1].set_xlabel("x")
axes[1].set_ylabel("$P_%s(x)$" % N)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Error Analysis
#
# Given $N + 1$ points we can form an interpolant $P_N(x)$ of degree $N$ where
#
# $$f(x) = P_N(x) + R_N(x)$$
# + [markdown] slideshow={"slide_type": "subslide"}
# We know from Lagrange's Theorem that the remainder term looks like
#
# $$R_N(x) = (x - x_0)(x - x_1)\cdots (x - x_{N})(x - x_{N+1}) \frac{f^{(N+1)}(c)}{(N+1)!}$$
#
# noting that we need to require that $f(x) \in C^{N+1}$ on the interval of interest. Taking the derivative of the interpolant $P_N(x)$ then leads to
#
# $$P_N'(x) = [y_0, y_1] + ((x - x_1) + (x - x_0)) [y_0, y_1, y_2] + \cdots + \left(\sum^{N-1}_{i=0}\left( \prod^{N-1}_{j=0,~j\neq i} (x - x_j) \right )\right ) [y_0, y_1, \ldots, y_N]$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Similarly we can find the derivative of the remainder term $R_N(x)$ as
#
# $$R_N'(x) = \left(\sum^{N}_{i=0} \left( \prod^{N}_{j=0,~j\neq i} (x - x_j) \right )\right ) \frac{f^{(N+1)}(c)}{(N+1)!}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Now if we consider the approximation of the derivative evaluated at one of our data points $(x_k, y_k)$ these expressions simplify such that
#
# $$f'(x_k) = P_N'(x_k) + R_N'(x_k)$$
# + [markdown] slideshow={"slide_type": "subslide"}
# If we let $\Delta x = \max_i |x_k - x_i|$ we then know that the remainder term will be $\mathcal{O}(\Delta x^N)$ as $\Delta x \rightarrow 0$ thus showing that this approach converges and we can find arbitrarily high order approximations (ignoring floating point error).
# + slideshow={"slide_type": "skip"}
# Compute the approximation to the derivative
# data = numpy.array([[-2.0, 1.0], [-1.5, -1.0], [-0.5, -3.0], [0.0, -2.0], [1.0, 3.0], [2.0, 1.0]])
num_points = 4
data = numpy.empty((num_points, 2))
data[:, 0] = numpy.linspace(-2.0, 2.0, num_points)
data[:, 1] = numpy.sin(data[:, 0])
N = data.shape[0] - 1
x_fine = numpy.linspace(-2.0, 2.0, 100)
# General form of derivative of P_N'(x)
P_prime = numpy.zeros(x.shape)
newton_basis_prime = numpy.empty(x.shape)
product = numpy.empty(x.shape)
for n in range(N + 1):
newton_basis_prime = 0.0
for i in range(n):
product = 1.0
for j in range(n):
if j != i:
product *= (x - data[j, 0])
newton_basis_prime += product
P_prime += divided_difference(data[:n+1, 0], data[:n+1, 1]) * newton_basis_prime
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x, numpy.cos(x), 'k')
axes.plot(x, P_prime, 'r--')
axes.set_title("$f'(x)$")
axes.set_xlabel("x")
axes.set_ylabel("$f'(x)$ and $\hat{f}'(x)$")
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Examples
#
# Often in practice we only use a small number of data points to derive a differentiation formula. In the context of differential equations we also often have $f(x)$ so that $f(x_k) = y_k$ and we can approximate the derivative of a known function $f(x)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 1: 1st order Forward and Backward Differences
#
# Using 2 points we can get an approximation that is $\mathcal{O}(\Delta x)$:
#
# $$f'(x) \approx P_1'(x) = [y_0, y_1] = \frac{y_1 - y_0}{x_1 - x_0} = \frac{y_1 - y_0}{\Delta x} = \frac{f(x_1) - f(x_0)}{\Delta x}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# We can also calculate the error as
#
# $$R_1'(x) = -\Delta x \frac{f''(c)}{2}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# We can also derive the "forward" and "backward" formulas by considering the question slightly differently. Say we want $f'(x_n)$, then the "forward" finite-difference can be written as
#
# $$f'(x_n) \approx D_1^+ = \frac{f(x_{n+1}) - f(x_n)}{\Delta x}$$
#
# and the "backward" finite-difference as
#
# $$f'(x_n) \approx D_1^- = \frac{f(x_n) - f(x_{n-1})}{\Delta x}$$
# + [markdown] slideshow={"slide_type": "skip"}
# Note these approximations should be familiar to use as the limit as $\Delta x \rightarrow 0$ these are no longer approximations but equivalent definitions of the derivative at $x_n$.
# + slideshow={"slide_type": "skip"}
f = lambda x: numpy.sin(x)
f_prime = lambda x: numpy.cos(x)
# Use uniform discretization
x = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, 1000)
N = 20
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x = x_hat[1] - x_hat[0]
# Compute forward difference using a loop
f_prime_hat = numpy.empty(x_hat.shape)
for i in range(N - 1):
f_prime_hat[i] = (f(x_hat[i+1]) - f(x_hat[i])) / delta_x
f_prime_hat[-1] = (f(x_hat[-1]) - f(x_hat[-2])) / delta_x
# Vector based calculation
# f_prime_hat[:-1] = (f(x_hat[1:]) - f(x_hat[:-1])) / (delta_x)
# Use first-order differences for points at edge of domain
f_prime_hat[-1] = (f(x_hat[-1]) - f(x_hat[-2])) / delta_x # Backward Difference at x_N
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x, f_prime(x), 'k')
axes.plot(x_hat + 0.5 * delta_x, f_prime_hat, 'ro')
axes.set_xlim((x[0], x[-1]))
axes.set_ylim((-1.1, 1.1))
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Aside: Computing Order of Convergence
#
# Say we had the error $E(\Delta x)$ and we wanted to make a statement about the rate of convergence (note we can replace $E$ here with the $R$ from above). Then we can do the following:
# $$\begin{aligned}
# E(\Delta x) &= C \Delta x^n \\
# \log E(\Delta x) &= \log C + n \log \Delta x
# \end{aligned}$$
#
# The slope of the line is $n$ when modeling the error like this! We can also match the first point by solving for $C$:
#
# $$
# C = e^{\log E(\Delta x) - n \log \Delta x}
# $$
# + slideshow={"slide_type": "skip"}
f = lambda x: numpy.sin(x)
f_prime = lambda x: numpy.cos(x)
# Compute the error as a function of delta_x
N_range = numpy.logspace(1, 4, 10, dtype=int)
delta_x = numpy.empty(N_range.shape)
error = numpy.empty((N_range.shape[0], 4))
for (i, N) in enumerate(N_range):
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x[i] = x_hat[1] - x_hat[0]
# Compute forward difference
f_prime_hat = numpy.empty(x_hat.shape)
f_prime_hat[:-1] = (f(x_hat[1:]) - f(x_hat[:-1])) / (delta_x[i])
# Use first-order differences for points at edge of domain
f_prime_hat[-1] = (f(x_hat[-1]) - f(x_hat[-2])) / delta_x[i] # Backward Difference at x_N
# The differences in error computations is interesting here. Note that the L_\infty norm returns a single
# point-wise like error where as the L_2 and L_1 are adding error up so they exhbit less convergence
error[i, 0] = numpy.linalg.norm(numpy.abs(f_prime(x_hat) - f_prime_hat), ord=numpy.infty)
error[i, 1] = numpy.linalg.norm(numpy.abs(f_prime(x_hat + 0.5 * delta_x[i]) - f_prime_hat), ord=numpy.infty)
error[i, 2] = numpy.linalg.norm(numpy.abs(f_prime(x_hat) - f_prime_hat), ord=1)
error[i, 3] = numpy.linalg.norm(numpy.abs(f_prime(x_hat) - f_prime_hat), ord=2)
error = numpy.array(error)
delta_x = numpy.array(delta_x)
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
fig.set_figheight(fig.get_figheight() * 2)
# plt.rc('legend', fontsize=6)
# plt.rc('font', size=6)
error_type = ['$L^\infty$', 'offset $L^\infty$', '$L^1$', '$L^2$']
for i in range(error.shape[1]):
axes = fig.add_subplot(2, 2, i + 1)
axes.loglog(delta_x, error[:, i], 'ko', label="Error")
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_x, order_C(delta_x[0], error[0, i], 1.0) * delta_x**1.0, 'r--', label="1st Order")
axes.loglog(delta_x, order_C(delta_x[0], error[0, i], 2.0) * delta_x**2.0, 'b--', label="2nd Order")
axes.legend(loc=4)
axes.set_title("Convergence of 1st Order Differences - %s" % error_type[i])
axes.set_xlabel("$\Delta x$")
axes.set_ylabel("$|f'(x) - \hat{f}'(x)|$")
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 2: 2nd Order Centered Difference
#
# Now lets use 3 points to calculate the 2nd order accurate finite-difference. Consider the points $(x_{n}, y_{n})$, $(x_{n-1}, y_{n-1})$, and $(x_{n+1}, y_{n+1})$, from before we have
#
# $$\begin{aligned}
# P_2(x) &= [f(x_0)] + (x - x_0) [f(x_0), f(x_1)] + (x - x_0)(x - x_1) [f(x_0), f(x_1), f(x_2)] \\
# &= f(x_0) + (x - x_0) \frac{f(x_1) - f(x_0)}{x_1 - x_0} + (x - x_0)(x - x_1) \left ( \frac{f(x_2) - f(x_1)}{(x_2 - x_1)(x_2 - x_0)} - \frac{f(x_1) - f(x_0)}{(x_1 - x_0)(x_2 - x_0)} \right )
# \end{aligned}$$
# Compute the formula for the derivative. Assume that the distance between the $x_i$ are equal.
# + [markdown] slideshow={"slide_type": "subslide"}
# $$\begin{aligned}
# P_2'(x) &= [f(x_n), f(x_{n+1})] + ((x - x_n) + (x - x_{n+1})) [f(x_n), f(x_{n+1}), f(x_{n-1})] \\
# &= \frac{f(x_{n+1}) - f(x_n)}{x_{n+1} - x_n} + ((x - x_n) + (x - x_{n+1})) \left ( \frac{f(x_{n-1}) - f(x_{n+1})}{(x_{n-1} - x_{n+1})(x_{n-1} - x_n)} - \frac{f(x_{n+1}) - f(x_n)}{(x_{n+1} - x_n)(x_{n-1} - x_n)} \right )
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Evaluating at $x_n$ and assuming the points $x_{n-1}, x_n, x_{n+1}$ are evenly spaced leads to
#
# $$\begin{aligned}
# P_2'(x_n) &= \frac{f(x_{n+1}) - f(x_n)}{\Delta x} - \Delta x \left ( \frac{f(x_{n-1}) - f(x_{n+1})}{2\Delta x^2} + \frac{f(x_{n+1}) - f(x_n)}{\Delta x^2} \right ) \\
# &=\frac{f(x_{n+1}) - f(x_n)}{\Delta x} - \left ( \frac{f(x_{n+1}) - 2f(x_n) + f(x_{n-1})}{2\Delta x}\right ) \\
# &=\frac{2f(x_{n+1}) - 2f(x_n) - f(x_{n+1}) + 2f(x_n) - f(x_{n-1})}{2 \Delta x} \\
# &=\frac{f(x_{n+1}) - f(x_{n-1})}{2 \Delta x}
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# This finite-difference is second order accurate and is centered about the point it is meant to approximate ($x_n$). We can show that it is second order by again considering the remainder term's derivative
#
# $$\begin{aligned}
# R_2'(x) &= \left(\sum^{2}_{i=0} \left( \prod^{2}_{j=0,~j\neq i} (x - x_j) \right )\right ) \frac{f'''(c)}{3!} \\
# &= \left ( (x - x_{n+1}) (x - x_{n-1}) + (x-x_n) (x-x_{n-1}) + (x-x_n)(x-x_{n+1}) \right ) \frac{f'''(c)}{3!}
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Again evaluating this expression at $x = x_n$ and assuming evenly space points we have
#
# $$R_2'(x_n) = -\Delta x^2 \frac{f'''(c)}{3!}$$
#
# showing that our error is $\mathcal{O}(\Delta x^2)$.
# + slideshow={"slide_type": "skip"}
f = lambda x: numpy.sin(x)
f_prime = lambda x: numpy.cos(x)
# Use uniform discretization
x = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, 1000)
N = 20
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x = x_hat[1] - x_hat[0]
# Compute derivative
f_prime_hat = numpy.empty(x_hat.shape)
f_prime_hat[1:-1] = (f(x_hat[2:]) - f(x_hat[:-2])) / (2 * delta_x)
# Use first-order differences for points at edge of domain
f_prime_hat[0] = (f(x_hat[1]) - f(x_hat[0])) / delta_x # Forward Difference at x_0
f_prime_hat[-1] = (f(x_hat[-1]) - f(x_hat[-2])) / delta_x # Backward Difference at x_N
# f_prime_hat[0] = (-3.0 * f(x_hat[0]) + 4.0 * f(x_hat[1]) - f(x_hat[2])) / (2.0 * delta_x)
# f_prime_hat[-1] = (3.0 * f(x_hat[-1]) - 4.0 * f(x_hat[-2]) + f(x_hat[-3])) / (2.0 * delta_x)
fig = plt.figure()
plt.rcdefaults()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x, f_prime(x), 'k')
axes.plot(x_hat, f_prime_hat, 'ro')
axes.set_xlim((x[0], x[-1]))
# axes.set_ylim((-1.1, 1.1))
plt.show()
# + slideshow={"slide_type": "skip"}
# Compute the error as a function of delta_x
delta_x = []
error = []
# for N in range(2, 101):
for N in range(50, 1000, 50):
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N + 1)
delta_x.append(x_hat[1] - x_hat[0])
# Compute derivative
f_prime_hat = numpy.empty(x_hat.shape)
f_prime_hat[1:-1] = (f(x_hat[2:]) - f(x_hat[:-2])) / (2 * delta_x[-1])
# Use first-order differences for points at edge of domain
f_prime_hat[0] = (f(x_hat[1]) - f(x_hat[0])) / delta_x[-1]
f_prime_hat[-1] = (f(x_hat[-1]) - f(x_hat[-2])) / delta_x[-1]
# Use second-order differences for points at edge of domain
# f_prime_hat[0] = (-3.0 * f(x_hat[0]) + 4.0 * f(x_hat[1]) + - f(x_hat[2])) / (2.0 * delta_x[-1])
# f_prime_hat[-1] = ( 3.0 * f(x_hat[-1]) + -4.0 * f(x_hat[-2]) + f(x_hat[-3])) / (2.0 * delta_x[-1])
# Compute Error
error.append(numpy.linalg.norm(numpy.abs(f_prime(x_hat) - f_prime_hat), ord=numpy.infty))
# error.append(numpy.linalg.norm(numpy.abs(f_prime(x_hat) - f_prime_hat), ord=2))
error = numpy.array(error)
delta_x = numpy.array(delta_x)
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.loglog(delta_x, error, "ro", label="Approx. Derivative")
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_x, order_C(delta_x[0], error[0], 1.0) * delta_x**1.0, 'b--', label="1st Order")
axes.loglog(delta_x, order_C(delta_x[0], error[0], 2.0) * delta_x**2.0, 'r--', label="2nd Order")
axes.legend(loc=4)
axes.set_title("Convergence of 2nd Order Differences")
axes.set_xlabel("$\Delta x$")
axes.set_ylabel("$|f'(x) - \hat{f}'(x)|$")
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 3: Alternative Derivations
#
# An alternative method for finding finite-difference formulas is by using Taylor series expansions about the point we want to approximate. The Taylor series about $x_n$ is
#
# $$f(x) = f(x_n) + (x - x_n) f'(x_n) + \frac{(x - x_n)^2}{2!} f''(x_n) + \frac{(x - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x - x_n)^4)$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Say we want to derive the second order accurate, first derivative approximation that we just did, this requires the values $(x_{n+1}, f(x_{n+1})$ and $(x_{n-1}, f(x_{n-1})$. We can express these values via our Taylor series approximation above as
#
# $$\begin{aligned}
# f(x_{n+1}) &= f(x_n) + (x_{n+1} - x_n) f'(x_n) + \frac{(x_{n+1} - x_n)^2}{2!} f''(x_n) + \frac{(x_{n+1} - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x_{n+1} - x_n)^4) \\
# &= f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)
# \end{aligned}$$
#
# and
#
# $$f(x_{n-1}) = f(x_n) + (x_{n-1} - x_n) f'(x_n) + \frac{(x_{n-1} - x_n)^2}{2!} f''(x_n) + \frac{(x_{n-1} - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x_{n-1} - x_n)^4) $$
#
# $$ = f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4) $$
# + [markdown] slideshow={"slide_type": "subslide"}
# Now to find out how to combine these into an expression for the derivative we assume our approximation looks like
#
# $$
# f'(x_n) + R(x_n) = A f(x_{n+1}) + B f(x_n) + C f(x_{n-1})
# $$
#
# where $R(x_n)$ is our error. Plugging in the Taylor series approximations we find
#
# $$\begin{aligned}
# f'(x_n) + R(x_n) &= A \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right ) \\
# & + B f(x_n) \\
# & + C \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4) \right )
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Since we want $R(x_n) = \mathcal{O}(\Delta x^2)$ we want all terms lower than this to cancel except for those multiplying $f'(x_n)$ as those should sum to 1 to give us our approximation. Collecting the terms with common evaluations of the derivatives on $f(x_n)$ we get a series of expressions for the coefficients $A$, $B$, and $C$ based on the fact we want an approximation to $f'(x_n)$. The $n=0$ terms collected are $A + B + C$ and are set to 0 as we want the $f(x_n)$ term to also cancel.
#
# $$\begin{aligned}
# f(x_n):& &A + B + C &= 0 \\
# f'(x_n): & &A \Delta x - C \Delta x &= 1 \\
# f''(x_n): & &A \frac{\Delta x^2}{2} + C \frac{\Delta x^2}{2} &= 0
# \end{aligned} $$
# + [markdown] slideshow={"slide_type": "subslide"}
# This last equation $\Rightarrow A = -C$, using this in the second equation gives $A = \frac{1}{2 \Delta x}$ and $C = -\frac{1}{2 \Delta x}$. The first equation then leads to $B = 0$. Putting this altogether then gives us our previous expression including an estimate for the error:
#
# $$\begin{aligned}
# f'(x_n) + R(x_n) &= \quad \frac{1}{2 \Delta x} \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right ) \\
# & \quad + 0 \cdot f(x_n) \\
# & \quad - \frac{1}{2 \Delta x} \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4) \right ) \\
# &= f'(x_n) + \frac{1}{2 \Delta x} \left ( \frac{2 \Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right )
# \end{aligned}$$
# so that we find
# $$
# R(x_n) = \frac{\Delta x^2}{3!} f'''(x_n) + \mathcal{O}(\Delta x^3) = \mathcal{O}(\Delta x^2)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Another way...
#
# There is one more way to derive the second order accurate, first order finite-difference formula. Consider the two first order forward and backward finite-differences averaged together:
#
# $$\frac{D_1^+(f(x_n)) + D_1^-(f(x_n))}{2} = \frac{f(x_{n+1}) - f(x_n) + f(x_n) - f(x_{n-1})}{2 \Delta x} = \frac{f(x_{n+1}) - f(x_{n-1})}{2 \Delta x}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 4: Higher Order Derivatives
#
# Using our Taylor series approach lets derive the second order accurate second derivative formula. Again we will use the same points and the Taylor series centered at $x = x_n$ so we end up with the same expression as before:
#
# $$\begin{aligned}
# f''(x_n) + R(x_n) &= \quad A \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5)\right ) \\
# &+ \quad B \cdot f(x_n) \\
# &+ \quad C \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right )
# \end{aligned}$$
#
# except this time we want to leave $f''(x_n)$ on the right hand side.
#
# Try out the same trick as before and see if you can setup the equations that need to be solved.
# + [markdown] slideshow={"slide_type": "subslide"}
# Doing the same trick as before we have the following expressions:
#
# $$\begin{aligned}
# f(x_n): & & A + B + C &= 0\\
# f'(x_n): & & A \Delta x - C \Delta x &= 0\\
# f''(x_n): & & A \frac{\Delta x^2}{2} + C \frac{\Delta x^2}{2} &= 1
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# The second equation implies $A = C$ which combined with the third implies
#
# $$A = C = \frac{1}{\Delta x^2}$$
#
# Finally the first equation gives
#
# $$B = -\frac{2}{\Delta x^2}$$
#
# leading to the final expression
#
# $$
# $$\begin{aligned}
# f''(x_n) + R(x_n) &= \quad \frac{1}{\Delta x^2} \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5)\right ) \\
# &+ \quad -\frac{2}{\Delta x^2} \cdot f(x_n) \\
# &+ \quad \frac{1}{\Delta x^2} \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right ) \\
# &= f''(x_n) + \frac{1}{\Delta x^2} \left(\frac{2 \Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right )
# \end{aligned}$$
# so that
#
# $$
# R(x_n) = \frac{\Delta x^2}{12} f^{(4)}(x_n) + \mathcal{O}(\Delta x^3)
# $$
# + slideshow={"slide_type": "skip"}
f = lambda x: numpy.sin(x)
f_dubl_prime = lambda x: -numpy.sin(x)
# Use uniform discretization
x = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, 1000)
N = 10
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x = x_hat[1] - x_hat[0]
# Compute derivative
f_dubl_prime_hat = numpy.empty(x_hat.shape)
f_dubl_prime_hat[1:-1] = (f(x_hat[2:]) -2.0 * f(x_hat[1:-1]) + f(x_hat[:-2])) / (delta_x**2)
# Use first-order differences for points at edge of domain
f_dubl_prime_hat[0] = (2.0 * f(x_hat[0]) - 5.0 * f(x_hat[1]) + 4.0 * f(x_hat[2]) - f(x_hat[3])) / delta_x**2
f_dubl_prime_hat[-1] = (2.0 * f(x_hat[-1]) - 5.0 * f(x_hat[-2]) + 4.0 * f(x_hat[-3]) - f(x_hat[-4])) / delta_x**2
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
axes.plot(x, f_dubl_prime(x), 'k')
axes.plot(x_hat, f_dubl_prime_hat, 'ro')
axes.set_xlim((x[0], x[-1]))
axes.set_ylim((-1.1, 1.1))
plt.show()
# + slideshow={"slide_type": "skip"}
f = lambda x: numpy.sin(x)
f_dubl_prime = lambda x: -numpy.sin(x)
# Compute the error as a function of delta_x
delta_x = []
error = []
# for N in xrange(2, 101):
for N in range(50, 1000, 50):
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x.append(x_hat[1] - x_hat[0])
# Compute derivative
f_dubl_prime_hat = numpy.empty(x_hat.shape)
f_dubl_prime_hat[1:-1] = (f(x_hat[2:]) -2.0 * f(x_hat[1:-1]) + f(x_hat[:-2])) / (delta_x[-1]**2)
# Use second-order differences for points at edge of domain
f_dubl_prime_hat[0] = (2.0 * f(x_hat[0]) - 5.0 * f(x_hat[1]) + 4.0 * f(x_hat[2]) - f(x_hat[3])) / delta_x[-1]**2
f_dubl_prime_hat[-1] = (2.0 * f(x_hat[-1]) - 5.0 * f(x_hat[-2]) + 4.0 * f(x_hat[-3]) - f(x_hat[-4])) / delta_x[-1]**2
error.append(numpy.linalg.norm(numpy.abs(f_dubl_prime(x_hat) - f_dubl_prime_hat), ord=numpy.infty))
error = numpy.array(error)
delta_x = numpy.array(delta_x)
fig = plt.figure()
axes = fig.add_subplot(1, 1, 1)
# axes.plot(delta_x, error)
axes.loglog(delta_x, error, "ko", label="Approx. Derivative")
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
axes.loglog(delta_x, order_C(delta_x[2], error[2], 1.0) * delta_x**1.0, 'b--', label="1st Order")
axes.loglog(delta_x, order_C(delta_x[2], error[2], 2.0) * delta_x**2.0, 'r--', label="2nd Order")
axes.legend(loc=4)
plt.show()
| 07_differentiation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import os
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import mean_squared_error
# %matplotlib inline
data = pd.read_csv("F:\\Python work\\titanic\\insurance (1).csv")
data.head(5)
data.info()
data.describe()
sex = data.groupby(by = 'sex').size()
print(sex)
smoker = data.groupby(by = 'smoker').size()
print(smoker)
region = data.groupby(by = 'region').size()
print(region)
## check the distribution of charges
distPlot = sns.distplot(data['charges'])
plt.title("Distirbution of Charges")
plt.show(distPlot)
## check charges vs features
meanGender = data.groupby(by = "sex")["charges"].mean()
print(meanGender)
print(meanGender["male"] - meanGender["female"])
boxPlot1 = sns.violinplot(x = "sex", y = "charges", data = data)
## check charges vs features
meanSmoker = data.groupby(by = "smoker")["charges"].mean()
print(meanSmoker)
print(meanSmoker["yes"] - meanSmoker["no"])
boxPlot1 = sns.violinplot(x = "smoker", y = "charges", data = data)
## check charges vs features
meanRegion = data.groupby(by = "region")["charges"].mean()
print(meanRegion)
boxPlot3 = sns.violinplot(x = "region", y = "charges", data = data)
pairPlot = sns.pairplot(data)
##smokers vs non-smokers
sns.set(style = "ticks")
smokerPairs = sns.pairplot(data, hue = "smoker")
## Dummify sex, smoker and region
scaleMinMax = MinMaxScaler()
data[["age", "bmi", "children"]] = scaleMinMax.fit_transform(data[["age", "bmi", "children"]])
data = pd.get_dummies(data, prefix = ["sex", "smoker", "region"])
## retain sex = male, smoker = yes, and remove 1 region = northeast to avoid dummytrap
data = data.drop(data.columns[[4,6,11]], axis = 1)
head = data.head()
print(head)
# +
## Quantifying the effect of the features to the medical charges
##We have already visualized the relationship of variables to charges.
##Now we will further investigate by looking at the relationships using multiple linear regression.
##The aim of this section is to quantify the relationship and not to create the prediction model. Let us first create a training and testing data set to proceed.
##Based on the visualization, we can make a couple of hypothesis about the relationship.
##There is no real difference in charges between gender or regions.
##The charge for smokers are very much higher than the non-smokers.
##The charge gets higher as the individual gets older.
##Lastly, the charge is higher for those who have fewer number of children.
X = data.drop(data.columns[[3]], axis = 1)
Y = data.iloc[:, 3]
X_train, x_test, Y_train, y_test = train_test_split(X, Y, random_state = 0)
# +
import statsmodels.api as sm
from scipy import stats
X_train2 = sm.add_constant(X_train)
linearModel = sm.OLS(Y_train, X_train2)
linear = linearModel.fit()
print(linear.summary())
# +
##There is no real difference in charges between gender or regions.
##Result: The p-value is 0.973 indicating there is no statistical difference between the gender or region group.
##The charge for smokers are very much higher than the non-smokers.
##Result: The p-value is 0.000 which indicates that there is a difference between the group.
##The charge gets higher as the individual gets older.
## Result: The p-value is 0.000 which indicates that the charge is higher as the individual gets older.
##The charge gers higher as the BMI gets higher.
##Result: The p-values is 0.000 which indicates that the charge is higher as the BMI gets higher.
##Lastly, there is significant decrease in charges as the number of children increases.
##Result: The p-value is 0.007. Interestingly, the coefficient is 2,211 which means that the charge gets higher as the individual has more number of childre.
##The initial hypothesis is incorrect. This is essentially the reason why we can't solely rely on visualization in generating conclusions.
# +
##In this section, we will create regression models and try to compare there robustness given the data.
##The models considered are Linear Regression, Ridge, LASSO, and ElasticNet.
# -
## try Linear Regression
from sklearn.linear_model import LinearRegression
linearModel = LinearRegression()
linear = linearModel.fit(X_train, Y_train)
linearPred = linear.predict(x_test)
mseLinear = metrics.mean_squared_error(y_test, linearPred)
rmseLinear = mseLinear**(1/2)
from sklearn.linear_model import Ridge
ridgeModel = Ridge()
ridge = ridgeModel.fit(X_train, Y_train)
ridgePred = ridge.predict(x_test)
mseRidge = metrics.mean_squared_error(y_test, ridgePred)
rmseRidge = mseRidge**(1/2)
from sklearn.linear_model import Lasso
lassoModel = Lasso()
lasso = lassoModel.fit(X_train, Y_train)
lassoPred = lasso.predict(x_test)
mseLasso = metrics.mean_squared_error(y_test, lassoPred)
rmseLasso = mseLasso**(1/2)
from sklearn.linear_model import ElasticNet
elasticNetModel = ElasticNet(alpha = 0.01, l1_ratio = 0.9, max_iter = 20)
ElasticNet = elasticNetModel.fit(X_train, Y_train)
ElasticNetPred = ElasticNet.predict(x_test)
mseElasticNet = metrics.mean_squared_error(y_test, ElasticNetPred)
rmseElasticNet = mseElasticNet**(1/2)
##Comparing the Models
performanceData = pd.DataFrame({"model":["linear", "lasso", "ridge", "elasticnet"], "rmse":[rmseLinear, rmseLasso, rmseRidge, rmseElasticNet]})
print(performanceData)
# +
##Based on the table above, linear regression has a slight edge among the models considered having the least RMSE.
##This is not surprising as the other 3 models are known to be more robust when there are quite a number of features. We only have 8 this time.
| Insurance Charge Multiple Regression 20.05.2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: synapse_pyspark
# kernelspec:
# display_name: Synapse PySpark
# language: Python
# name: synapse_pyspark
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Run Apache Beam on Azure Synpase
#
# Install Apache Beam and dependencies
#
# ```
# apache-beam[interactive]
# pyarrow
# pandas==1.0.0
# ```
#
# +
import apache_beam as beam
with beam.Pipeline() as pipeline:
lines = (
pipeline
| beam.Create([
'To be, or not to be: that is the question: ',
"Whether 'tis nobler in the mind to suffer ",
'The slings and arrows of outrageous fortune, ',
'Or to take arms against a sea of troubles, ',
])
)
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Update Azure Storage Connection string
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
connection_string = ""
os.environ['AZURE_STORAGE_CONNECTION_STRING'] = connection_string
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import os
import re
import apache_beam as beam
from apache_beam.io.azure.blobstoragefilesystem import BlobStorageFileSystem
sampleText = 'azfs://hyundevsynapsestorage/dev-synapse/user/trusted-service-user/beam/sample/sample.txt'
outputPath = 'azfs://hyundevsynapsestorage/dev-synapse/user/trusted-service-user/beam/output/wordcount.txt'
with beam.Pipeline() as pipeline:
lines = (
pipeline
| beam.io.textio.ReadFromText(sampleText)
| beam.FlatMap(lambda line: re.findall(r"[a-zA-Z']+", line))
| beam.Map(lambda word: (word, 1))
| beam.CombinePerKey(sum)
| beam.Map(lambda word_count: str(word_count))
| beam.io.textio.WriteToText(outputPath)
)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
outputPath = 'azfs://hyundevsynapsestorage/dev-synapse/user/trusted-service-user/beam/output/hamlet.txt'
with beam.Pipeline() as pipeline:
lines = (
pipeline
| beam.Create([
'To be, or not to be: that is the question: ',
"Whether 'tis nobler in the mind to suffer ",
'The slings and arrows of outrageous fortune, ',
'Or to take arms against a sea of troubles, ',
])
| beam.io.textio.WriteToText(outputPath)
)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# sampledata_path = 'azfs://hyundevsynapsestorage/dev-synapse/data/bingCOVID19/*'
# with beam.Pipeline() as pipeline:
# lines = (
# pipeline
# | beam.io.parquetio.ReadFromParquet(sampledata_path)
# | beam.Map(lambda table: table.to_pandas())
# )
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import pyarrow
outputdata_path = 'azfs://hyundevsynapsestorage/dev-synapse/data/beamoutput/outputsample.parquet'
with beam.Pipeline() as p:
records = p | 'Read' >> beam.Create(
[{'name': 'foo', 'age': 10}, {'name': 'bar', 'age': 20}]
)
_ = records | 'Write' >> beam.io.WriteToParquet(outputdata_path,
pyarrow.schema(
[('name', pyarrow.binary()), ('age', pyarrow.int64())]
)
)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} microsoft={"language": "python"}
# %%pyspark
df = spark.read.load([
'abfss://dev-synapse@hyundevsynapsestorage.dfs.core.windows.net/data/beamoutput/outputsample.parquet-00000-of-00001'
], format='parquet')
display(df.head(2))
| Apache-Beam-Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: bce_network
# language: python
# name: bce_network
# ---
from hw_demo_estimation import etl, graph_manipulation as gm, data_viz as dv
# # Demography Estimation Homework
#
# This is a markdown cell in a jupyter notebook where I can write analysis about the charts and the statistics that I calculated
nodes, edges = etl.load_and_select_profiles_and_edges()
G= gm.create_graph_from_nodes_and_edges(nodes, edges)
dv.plot_degree_distribution(G)
# descriptive analytics of the nodes
# age distribution by gender
dv.plot_age_distribution_by_gender(nodes)
dv.plot_node_degree_by_gender(nodes, G)
edges_w_features = gm.add_node_features_to_edges(nodes, edges)
dv.plot_age_relations_heatmap(edges_w_features)
# THE END
| POKEC Network Analysis Stub for Students.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
from layer_naive import *
apple = 100
apple_num = 2
tax = 1.1
mul_apple_layer = MulLayer()
mul_tax_layer = MulLayer()
# forward
apple_price = mul_apple_layer.forward(apple, apple_num)
price = mul_tax_layer.forward(apple_price, tax)
# backward
dprice = 1
dapple_price, dtax = mul_tax_layer.backward(dprice)
dapple, dapple_num = mul_apple_layer.backward(dapple_price)
print("price:", int(price))
print("dApple:", dapple)
print("dApple_num:", int(dapple_num))
print("dTax:", dtax)
| ch05/buy_apple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import seaborn
import numpy, scipy, scipy.spatial, matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (14, 3)
# [← Back to Index](index.html)
# # Dynamic Time Warping
# In MIR, we often want to compare two sequences of different lengths. For example, we may want to compute a similarity measure between two versions of the same song. These two signals, $x$ and $y$, may have similar sequences of chord progressions and instrumentations, but there may be timing deviations between the two. Even if we were to express the two audio signals using the same feature space (e.g. chroma or MFCCs), we cannot simply sum their pairwise distances because the signals have different lengths.
#
# As another example, you might want to align two different performances of the same musical work, e.g. so you can hop from one performance to another at any moment in the work. This problem is known as **music synchronization** (FMP, p. 115).
# **Dynamic time warping (DTW)** ([Wikipedia](https://en.wikipedia.org/wiki/Dynamic_time_warping); FMP, p. 131) is an algorithm used to align two sequences of similar content but possibly different lengths.
#
# Given two sequences, $x[n], n \in \{0, ..., N_x - 1\}$, and $y[n], n \in \{0, ..., N_y - 1\}$, DTW produces a set of index coordinate pairs $\{ (i, j) ... \}$ such that $x[i]$ and $y[j]$ are similar.
#
# We will use the same dynamic programming approach described in the notebooks [Dynamic Programming](dp.html) and [Longest Common Subsequence](lcs.html).
# ## Example
# Create two arrays, $x$ and $y$, of lengths $N_x$ and $N_y$, respectively.
x = [0, 4, 4, 0, -4, -4, 0]
y = [1, 3, 4, 3, 1, -1, -2, -1, 0]
nx = len(x)
ny = len(y)
plt.plot(x)
plt.plot(y, c='r')
plt.legend(('x', 'y'))
# In this simple example, there is only one value or "feature" at each time index. However, in practice, you can use sequences of *vectors*, e.g. spectrograms, chromagrams, or MFCC-grams.
# ## Distance Metric
# DTW requires the use of a distance metric between corresponding observations of `x` and `y`. One common choice is the **Euclidean distance** ([Wikipedia](https://en.wikipedia.org/wiki/Euclidean_distance); FMP, p. 454):
scipy.spatial.distance.euclidean(0, [3, 4])
scipy.spatial.distance.euclidean([0, 0], [5, 12])
# Another choice is the **Manhattan or cityblock distance**:
scipy.spatial.distance.cityblock(0, [3, 4])
scipy.spatial.distance.cityblock([0, 0], [5, 12])
# Another choice might be the **cosine distance** ([Wikipedia](https://en.wikipedia.org/wiki/Cosine_similarity); FMP, p. 376) which can be interpreted as the (normalized) angle between two vectors:
scipy.spatial.distance.cosine([1, 0], [100, 0])
scipy.spatial.distance.cosine([1, 0, 0], [0, 0, -1])
scipy.spatial.distance.cosine([1, 0], [-1, 0])
# For more distance metrics, see [`scipy.spatial.distance`](https://docs.scipy.org/doc/scipy/reference/spatial.distance.html).
# ## Step 1: Cost Table Construction
# As described in the notebooks [Dynamic Programming](dp.html) and [Longest Common Subsequence](lcs.html), we will use dynamic programming to solve this problem. First, we create a table which stores the solutions to all subproblems. Then, we will use this table to solve each larger subproblem until the problem is solved for the full original inputs.
# The basic idea of DTW is to find a path of index coordinate pairs the sum of distances along the path $P$ is minimized:
#
# $$ \min \sum_{(i, j) \in P} d(x[i], y[j]) $$
# The path constraint is that, at $(i, j)$, the valid steps are $(i+1, j)$, $(i, j+1)$, and $(i+1, j+1)$. In other words, the alignment always moves forward in time for at least one of the signals. It never goes forward in time for one signal and backward in time for the other signal.
# Here is the optimal substructure. Suppose that the best alignment contains index pair `(i, j)`, i.e., `x[i]` and `y[j]` are part of the optimal DTW path. Then, we prepend to the optimal path
#
# $$ \mathrm{argmin} \ \{ d(x[i-1], y[j]), d(x[i], y[j-1]), d(x[i-1], j-1]) \} $$
# We create a table where cell `(i, j)` stores the optimum cost of `dtw(x[:i], y[:j])`, i.e. the optimum cost from `(0, 0)` to `(i, j)`. First, we solve for the boundary cases, i.e. when either one of the two sequences is empty. Then we populate the table from the top left to the bottom right.
def dtw_table(x, y):
nx = len(x)
ny = len(y)
table = numpy.zeros((nx+1, ny+1))
# Compute left column separately, i.e. j=0.
table[1:, 0] = numpy.inf
# Compute top row separately, i.e. i=0.
table[0, 1:] = numpy.inf
# Fill in the rest.
for i in range(1, nx+1):
for j in range(1, ny+1):
d = scipy.spatial.distance.euclidean(x[i-1], y[j-1])
table[i, j] = d + min(table[i-1, j], table[i, j-1], table[i-1, j-1])
return table
table = dtw_table(x, y)
# Let's visualize this table:
print ' ', ''.join('%4d' % n for n in y)
print ' +' + '----' * (ny+1)
for i, row in enumerate(table):
if i == 0:
z0 = ''
else:
z0 = x[i-1]
print ('%4s |' % z0) + ''.join('%4.0f' % z for z in row)
# The time complexity of this operation is $O(N_x N_y)$. The space complexity is $O(N_x N_y)$.
# ## Step 2: Backtracking
# To assemble the best path, we use **backtracking** (FMP, p. 139). We will start at the end, $(N_x - 1, N_y - 1)$, and backtrack to the beginning, $(0, 0)$.
# Finally, just read off the sequences of time index pairs starting at the end.
def dtw(x, y, table):
i = len(x)
j = len(y)
path = [(i, j)]
while i > 0 or j > 0:
minval = numpy.inf
if table[i-1, j] < minval:
minval = table[i-1, j]
step = (i-1, j)
if table[i][j-1] < minval:
minval = table[i, j-1]
step = (i, j-1)
if table[i-1][j-1] < minval:
minval = table[i-1, j-1]
step = (i-1, j-1)
path.insert(0, step)
i, j = step
return path
path = dtw(x, y, table)
path
# The time complexity of this operation is $O(N_x + N_y)$.
# As a sanity check, compute the total distance of this alignment:
sum(abs(x[i-1] - y[j-1]) for (i, j) in path if i >= 0 and j >= 0)
# Indeed, that is the same as the cumulative distance of the optimal path computed earlier:
table[-1, -1]
# [← Back to Index](index.html)
| dtw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PreFRBLE.convenience import *
from PreFRBLE.estimate_redshift import *
from PreFRBLE.plot import *
from PreFRBLE.likelihood import *
from PreFRBLE.physics import *
import Pshirkov16 ## this contains procedures for Monte-Carlo simulation following Phsirkov e al. 2016
# -
# ### Compare mean(redshift)
# Here we validate the results of the numerical IGM simulation to analytical predictions by comparing the mean DM and RM and their redshift evolution.
# +
## define analytical estimate
from astropy import units as u
from astropy.coordinates import SkyCoord, Distance
from astropy.cosmology import Planck15
from scipy import integrate
rho_crit = Planck15.critical_density0.value # g/cm3
omega_b = Planck15.Ob0
omega_m = Planck15.Om0
omega_L = Planck15.Ode0
m_p = 1.67e-24 # g
mu_e = 1.16
c = 2.99e8 # m/s
H_0 = 1./Planck15.hubble_time.value
n_e = rho_crit * omega_b / m_p / mu_e
def DM(z, f_IGM=0.82):
""" average DM as function of redshift from cosmic parameters (Planck15), assuming homogeneous IGM """
return n_e * f_IGM *c/H_0* integrate.quad( lambda z: (1+z)/( omega_m*(1+z)**3 + omega_L )**0.5, 0, z)[0]
# +
## initialize estimate following Pshirkov et al. 2016
## n_e = 1.8e-7 cm^-3 implicitly assumes ~83% of baryons in ionized IGM
B0 = 1e-4 # muG at n_0=1.8e-7
l_c = 100 # Mpc correlation length
z_max = 3.0
t0 = time()
LoS = Pshirkov16.Sightline( B0, l_c, z_max )
print('initialized in %.0f seconds' % (time() - t0))
# +
## perform Monte-Carlo simulation
t0 = time()
zs = LoS.zarray
N=1000
DMs = np.array([ LoS.DispersionMeasure( *LoS.CreateArrays( z_max, zarray=zs )[::-1] ) for i in range(N)])
print('%.0f seconds, now for the RM' % (time() - t0))
RMs = np.array([ np.abs(LoS.RotationMeasure( *LoS.CreateArrays( z_max, zarray=zs )[::-1])) for i in range(N)])
print('%.0f seconds, now for the plot' % (time() - t0))
# +
## compute results
DM_mean_log = np.mean( np.log10(DMs), axis=0)
DM_mean = 10.**DM_mean_log
DM_std = np.std( np.log10(DMs), axis=0)
DM_err = np.array( [ DM_mean - 10.**(DM_mean_log-DM_std), 10**( DM_mean_log+DM_std) - DM_mean ] )
RM_mean_log = np.mean( np.log10(RMs), axis=0)
RM_mean = 10.**RM_mean_log
#RM_mean = np.mean( RMs, axis=0 )
RM_std = np.std( np.log10(RMs), axis=0)
RM_err = np.array( [ RM_mean - 10.**(RM_mean_log-RM_std), 10**( RM_mean_log+RM_std) - RM_mean ] )
#RM_mean = 10.**np.mean( np.log10(RMs), axis=0)
#RM_std = np.std( RMs, axis=0)
# +
## plot and compare to other models
fig, ax = plt.subplots()
f_IGM = 0.8 ## IGM baryon content implicitly assumed by Pshirkov et al. 2016
scenario_IGM = { 'IGM': ['primordial_C%.0f' % (1000*f_IGM) ]} ## scenario considering only the IGM, using same f_IGM as Pshirkov
## first plot DM
## plot estimate of constrained simulation
PlotAverageEstimate( measure='DM', scenario=scenario_IGM, label='constrained simulation', ax=ax, errorevery=5)
## plot Pshirkov estimate
ax.errorbar( zs[:-1], DM_mean, yerr=DM_err, label='Pshirkov+16', errorevery=500, linestyle='-.')
## plot analytical predictions, see Hackstein et al. 2020
ax.plot( redshift_bins, redshift_bins*1000*f_IGM, label=r"DM$ = f_{\rm IGM} \times z \times 1000$ pc cm$^{-3}$", linestyle='--' )
ax.plot( redshift_bins, [DM(z , f_IGM=f_IGM ) for z in redshift_bins], label=r"DM$(f_{\rm IGM}=%.1f) = \int n_e {\rm d}l$" % f_IGM, linestyle=':' )
ax.set_ylim(1e2,1e4)
ax.legend(fontsize=16, loc=4)
ax.tick_params(axis='both', which='major', labelsize=16)
plt.show()
## plot RM
fig, ax = plt.subplots()
## plot estimate of constrained simulation
PlotAverageEstimate( measure='RM', scenario=scenario_IGM, label='constrained simulation', ax=ax, errorevery=5)
## plot Pshirkov estimate
ax.errorbar( zs[:-1], RM_mean, yerr=RM_err, label='Pshirkov+16', errorevery=500, linestyle='-.')
ax.set_ylim(1e-3,1.5e0)
ax.set_xlim(-.1,3)
ax.legend(fontsize=16, loc=4)
ax.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# The procedure to plot the average estimate of the constrained simulation can of course be used to plot the estimate of any scenario.
# +
Properties = {
'IGM' : ['primordial'],
'Host' : ['Rodrigues18'],
'Inter' : ['Rodrigues18'],
'Local' : ['Piro18_wind'],
'N_inter' : True
}
scenario = Scenario( redshift=0.1, **Properties)
fig, ax = plt.subplots()
PlotAverageEstimate( measure='DM', scenario=scenario, label='intervening', errorevery=5, ax=ax)
plt.show()
| notebooks/RedshiftMean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Customer Churn Prediction with XGBoost
# _**Using Gradient Boosted Trees to Predict Mobile Customer Departure**_
#
# ---
#
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Host](#Host)
# 1. [Evaluate](#Evaluate)
# 1. [Relative cost of errors](#Relative-cost-of-errors)
# 1. [Extensions](#Extensions)
#
# ---
#
# ## Background
#
# _This notebook has been adapted from an [AWS blog post](https://aws.amazon.com/blogs/ai/predicting-customer-churn-with-amazon-machine-learning/)_
#
# Losing customers is costly for any business. Identifying unhappy customers early on gives you a chance to offer them incentives to stay. This notebook describes using machine learning (ML) for the automated identification of unhappy customers, also known as customer churn prediction. ML models rarely give perfect predictions though, so this notebook is also about how to incorporate the relative costs of prediction mistakes when determining the financial outcome of using ML.
#
# We use an example of churn that is familiar to all of us–leaving a mobile phone operator. Seems like I can always find fault with my provider du jour! And if my provider knows that I’m thinking of leaving, it can offer timely incentives–I can always use a phone upgrade or perhaps have a new feature activated–and I might just stick around. Incentives are often much more cost effective than losing and reacquiring a customer.
#
# ---
#
# ## Setup
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
# Define IAM role
import boto3
import re
import sagemaker
role = sagemaker.get_execution_role()
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = 'DEMO-xgboost-churn'
# -
# Next, we'll import the Python libraries we'll need for the remainder of the exercise.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import os
import sys
import time
import json
from IPython.display import display
from time import strftime, gmtime
import sagemaker
from sagemaker.predictor import csv_serializer
# ---
# ## Data
#
# Mobile operators have historical records on which customers ultimately ended up churning and which continued using the service. We can use this historical information to construct an ML model of one mobile operator’s churn using a process called training. After training the model, we can pass the profile information of an arbitrary customer (the same profile information that we used to train the model) to the model, and have the model predict whether this customer is going to churn. Of course, we expect the model to make mistakes–after all, predicting the future is tricky business! But I’ll also show how to deal with prediction errors.
#
# The dataset we use is publicly available and was mentioned in the book [Discovering Knowledge in Data](https://www.amazon.com/dp/0470908742/) by <NAME>. It is attributed by the author to the University of California Irvine Repository of Machine Learning Datasets. Let's download and read that dataset in now:
# !wget http://dataminingconsultant.com/DKD2e_data_sets.zip
# !unzip -o DKD2e_data_sets.zip
churn = pd.read_csv('./Data sets/churn.txt')
pd.set_option('display.max_columns', 500)
churn
# By modern standards, it’s a relatively small dataset, with only 3,333 records, where each record uses 21 attributes to describe the profile of a customer of an unknown US mobile operator. The attributes are:
#
# - `State`: the US state in which the customer resides, indicated by a two-letter abbreviation; for example, OH or NJ
# - `Account Length`: the number of days that this account has been active
# - `Area Code`: the three-digit area code of the corresponding customer’s phone number
# - `Phone`: the remaining seven-digit phone number
# - `Int’l Plan`: whether the customer has an international calling plan: yes/no
# - `VMail Plan`: whether the customer has a voice mail feature: yes/no
# - `VMail Message`: presumably the average number of voice mail messages per month
# - `Day Mins`: the total number of calling minutes used during the day
# - `Day Calls`: the total number of calls placed during the day
# - `Day Charge`: the billed cost of daytime calls
# - `Eve Mins, Eve Calls, Eve Charge`: the billed cost for calls placed during the evening
# - `Night Mins`, `Night Calls`, `Night Charge`: the billed cost for calls placed during nighttime
# - `Intl Mins`, `Intl Calls`, `Intl Charge`: the billed cost for international calls
# - `CustServ Calls`: the number of calls placed to Customer Service
# - `Churn?`: whether the customer left the service: true/false
#
# The last attribute, `Churn?`, is known as the target attribute–the attribute that we want the ML model to predict. Because the target attribute is binary, our model will be performing binary prediction, also known as binary classification.
#
# Let's begin exploring the data:
# +
# Frequency tables for each categorical feature
for column in churn.select_dtypes(include=['object']).columns:
display(pd.crosstab(index=churn[column], columns='% observations', normalize='columns'))
# Histograms for each numeric features
display(churn.describe())
# %matplotlib inline
hist = churn.hist(bins=30, sharey=True, figsize=(10, 10))
# -
# We can see immediately that:
# - `State` appears to be quite evenly distributed
# - `Phone` takes on too many unique values to be of any practical use. It's possible parsing out the prefix could have some value, but without more context on how these are allocated, we should avoid using it.
# - Only 14% of customers churned, so there is some class imabalance, but nothing extreme.
# - Most of the numeric features are surprisingly nicely distributed, with many showing bell-like gaussianity. `VMail Message` being a notable exception (and `Area Code` showing up as a feature we should convert to non-numeric).
churn = churn.drop('Phone', axis=1)
churn['Area Code'] = churn['Area Code'].astype(object)
# Next let's look at the relationship between each of the features and our target variable.
# +
for column in churn.select_dtypes(include=['object']).columns:
if column != 'Churn?':
display(pd.crosstab(index=churn[column], columns=churn['Churn?'], normalize='columns'))
for column in churn.select_dtypes(exclude=['object']).columns:
print(column)
hist = churn[[column, 'Churn?']].hist(by='Churn?', bins=30)
plt.show()
# -
# Interestingly we see that churners appear:
# - Fairly evenly distributed geographically
# - More likely to have an international plan
# - Less likely to have a voicemail plan
# - To exhibit some bimodality in daily minutes (either higher or lower than the average for non-churners)
# - To have a larger number of customer service calls (which makes sense as we'd expect customers who experience lots of problems may be more likely to churn)
#
# In addition, we see that churners take on very similar distributions for features like `Day Mins` and `Day Charge`. That's not surprising as we'd expect minutes spent talking to correlate with charges. Let's dig deeper into the relationships between our features.
display(churn.corr())
pd.plotting.scatter_matrix(churn, figsize=(12, 12))
plt.show()
# We see several features that essentially have 100% correlation with one another. Including these feature pairs in some machine learning algorithms can create catastrophic problems, while in others it will only introduce minor redundancy and bias. Let's remove one feature from each of the highly correlated pairs: Day Charge from the pair with Day Mins, Night Charge from the pair with Night Mins, Intl Charge from the pair with Intl Mins:
churn = churn.drop(['Day Charge', 'Eve Charge', 'Night Charge', 'Intl Charge'], axis=1)
# Now that we've cleaned up our dataset, let's determine which algorithm to use. As mentioned above, there appear to be some variables where both high and low (but not intermediate) values are predictive of churn. In order to accommodate this in an algorithm like linear regression, we'd need to generate polynomial (or bucketed) terms. Instead, let's attempt to model this problem using gradient boosted trees. Amazon SageMaker provides an XGBoost container that we can use to train in a managed, distributed setting, and then host as a real-time prediction endpoint. XGBoost uses gradient boosted trees which naturally account for non-linear relationships between features and the target variable, as well as accommodating complex interactions between features.
#
# Amazon SageMaker XGBoost can train on data in either a CSV or LibSVM format. For this example, we'll stick with CSV. It should:
# - Have the predictor variable in the first column
# - Not have a header row
#
# But first, let's convert our categorical features into numeric features.
model_data = pd.get_dummies(churn)
model_data = pd.concat([model_data['Churn?_True.'], model_data.drop(['Churn?_False.', 'Churn?_True.'], axis=1)], axis=1)
# And now let's split the data into training, validation, and test sets. This will help prevent us from overfitting the model, and allow us to test the models accuracy on data it hasn't already seen.
train_data, validation_data, test_data = np.split(model_data.sample(frac=1, random_state=1729), [int(0.7 * len(model_data)), int(0.9 * len(model_data))])
train_data.to_csv('train.csv', header=False, index=False)
validation_data.to_csv('validation.csv', header=False, index=False)
# Now we'll upload these files to S3.
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation/validation.csv')).upload_file('validation.csv')
# ---
# ## Train
#
# Moving onto training, first we'll need to specify the locations of the XGBoost algorithm containers.
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost')
# Then, because we're training with the CSV file format, we'll create `s3_input`s that our training function can use as a pointer to the files in S3.
s3_input_train = sagemaker.s3_input(s3_data='s3://{}/{}/train'.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data='s3://{}/{}/validation/'.format(bucket, prefix), content_type='csv')
# Now, we can specify a few parameters like what type of training instances we'd like to use and how many, as well as our XGBoost hyperparameters. A few key hyperparameters are:
# - `max_depth` controls how deep each tree within the algorithm can be built. Deeper trees can lead to better fit, but are more computationally expensive and can lead to overfitting. There is typically some trade-off in model performance that needs to be explored between a large number of shallow trees and a smaller number of deeper trees.
# - `subsample` controls sampling of the training data. This technique can help reduce overfitting, but setting it too low can also starve the model of data.
# - `num_round` controls the number of boosting rounds. This is essentially the subsequent models that are trained using the residuals of previous iterations. Again, more rounds should produce a better fit on the training data, but can be computationally expensive or lead to overfitting.
# - `eta` controls how aggressive each round of boosting is. Larger values lead to more conservative boosting.
# - `gamma` controls how aggressively trees are grown. Larger values lead to more conservative models.
#
# More detail on XGBoost's hyperparmeters can be found on their GitHub [page](https://github.com/dmlc/xgboost/blob/master/doc/parameter.md).
# +
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
num_round=100)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ---
# ## Host
#
# Now that we've trained the algorithm, let's create a model and deploy it to a hosted endpoint.
xgb_predictor = xgb.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# ### Evaluate
#
# Now that we have a hosted endpoint running, we can make real-time predictions from our model very easily, simply by making an http POST request. But first, we'll need to setup serializers and deserializers for passing our `test_data` NumPy arrays to the model behind the endpoint.
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
xgb_predictor.deserializer = None
# Now, we'll use a simple function to:
# 1. Loop over our test dataset
# 1. Split it into mini-batches of rows
# 1. Convert those mini-batchs to CSV string payloads
# 1. Retrieve mini-batch predictions by invoking the XGBoost endpoint
# 1. Collect predictions and convert from the CSV output our model provides into a NumPy array
# +
def predict(data, rows=500):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, xgb_predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
predictions = predict(test_data.as_matrix()[:, 1:])
# -
# There are many ways to compare the performance of a machine learning model, but let's start by simply by comparing actual to predicted values. In this case, we're simply predicting whether the customer churned (`1`) or not (`0`), which produces a simple confusion matrix.
pd.crosstab(index=test_data.iloc[:, 0], columns=np.round(predictions), rownames=['actual'], colnames=['predictions'])
# _Note, due to randomized elements of the algorithm, you results may differ slightly._
#
# Of the 48 churners, we've correctly predicted 39 of them (true positives). And, we incorrectly predicted 4 customers would churn who then ended up not doing so (false positives). There are also 9 customers who ended up churning, that we predicted would not (false negatives).
#
# An important point here is that because of the `np.round()` function above we are using a simple threshold (or cutoff) of 0.5. Our predictions from `xgboost` come out as continuous values between 0 and 1 and we force them into the binary classes that we began with. However, because a customer that churns is expected to cost the company more than proactively trying to retain a customer who we think might churn, we should consider adjusting this cutoff. That will almost certainly increase the number of false positives, but it can also be expected to increase the number of true positives and reduce the number of false negatives.
#
# To get a rough intuition here, let's look at the continuous values of our predictions.
plt.hist(predictions)
plt.show()
# The continuous valued predictions coming from our model tend to skew toward 0 or 1, but there is sufficient mass between 0.1 and 0.9 that adjusting the cutoff should indeed shift a number of customers' predictions. For example...
pd.crosstab(index=test_data.iloc[:, 0], columns=np.where(predictions > 0.3, 1, 0))
# We can see that changing the cutoff from 0.5 to 0.3 results in 1 more true positives, 3 more false positives, and 1 fewer false negatives. The numbers are small overall here, but that's 6-10% of customers overall that are shifting because of a change to the cutoff. Was this the right decision? We may end up retaining 3 extra customers, but we also unnecessarily incentivized 5 more customers who would have stayed. Determining optimal cutoffs is a key step in properly applying machine learning in a real-world setting. Let's discuss this more broadly and then apply a specific, hypothetical solution for our current problem.
#
# ### Relative cost of errors
#
# Any practical binary classification problem is likely to produce a similarly sensitive cutoff. That by itself isn’t a problem. After all, if the scores for two classes are really easy to separate, the problem probably isn’t very hard to begin with and might even be solvable with simple rules instead of ML.
#
# More important, if I put an ML model into production, there are costs associated with the model erroneously assigning false positives and false negatives. I also need to look at similar costs associated with correct predictions of true positives and true negatives. Because the choice of the cutoff affects all four of these statistics, I need to consider the relative costs to the business for each of these four outcomes for each prediction.
#
# #### Assigning costs
#
# What are the costs for our problem of mobile operator churn? The costs, of course, depend on the specific actions that the business takes. Let's make some assumptions here.
#
# First, assign the true negatives the cost of \$0. Our model essentially correctly identified a happy customer in this case, and we don’t need to do anything.
#
# False negatives are the most problematic, because they incorrectly predict that a churning customer will stay. We lose the customer and will have to pay all the costs of acquiring a replacement customer, including foregone revenue, advertising costs, administrative costs, point of sale costs, and likely a phone hardware subsidy. A quick search on the Internet reveals that such costs typically run in the hundreds of dollars so, for the purposes of this example, let's assume \$500. This is the cost of false negatives.
#
# Finally, for customers that our model identifies as churning, let's assume a retention incentive in the amount of \\$100. If my provider offered me such a concession, I’d certainly think twice before leaving. This is the cost of both true positive and false positive outcomes. In the case of false positives (the customer is happy, but the model mistakenly predicted churn), we will “waste” the \\$100 concession. We probably could have spent that \\$100 more effectively, but it's possible we increased the loyalty of an already loyal customer, so that’s not so bad.
# #### Finding the optimal cutoff
#
# It’s clear that false negatives are substantially more costly than false positives. Instead of optimizing for error based on the number of customers, we should be minimizing a cost function that looks like this:
#
# ```txt
# $500 * FN(C) + $0 * TN(C) + $100 * FP(C) + $100 * TP(C)
# ```
#
# FN(C) means that the false negative percentage is a function of the cutoff, C, and similar for TN, FP, and TP. We need to find the cutoff, C, where the result of the expression is smallest.
#
# A straightforward way to do this, is to simply run a simulation over a large number of possible cutoffs. We test 100 possible values in the for loop below.
# +
cutoffs = np.arange(0.01, 1, 0.01)
costs = []
for c in cutoffs:
costs.append(np.sum(np.sum(np.array([[0, 100], [500, 100]]) *
pd.crosstab(index=test_data.iloc[:, 0],
columns=np.where(predictions > c, 1, 0)))))
costs = np.array(costs)
plt.plot(cutoffs, costs)
plt.show()
print('Cost is minimized near a cutoff of:', cutoffs[np.argmin(costs)], 'for a cost of:', np.min(costs))
# -
# The above chart shows how picking a threshold too low results in costs skyrocketing as all customers are given a retention incentive. Meanwhile, setting the threshold too high results in too many lost customers, which ultimately grows to be nearly as costly. The overall cost can be minimized at \\$ 8400 by setting the cutoff to 0.46, which is substantially better than the
# \\$ 20k+ I would expect to lose by not taking any action.
# ---
# ## Extensions
#
# This notebook showcased how to build a model that predicts whether a customer is likely to churn, and then how to optimally set a threshold that accounts for the cost of true positives, false positives, and false negatives. There are several means of extending it including:
# - Some customers who receive retention incentives will still churn. Including a probability of churning despite receiving an incentive in our cost function would provide a better ROI on our retention programs.
# - Customers who switch to a lower-priced plan or who deactivate a paid feature represent different kinds of churn that could be modeled separately.
# - Modeling the evolution of customer behavior. If usage is dropping and the number of calls placed to Customer Service is increasing, you are more likely to experience churn then if the trend is the opposite. A customer profile should incorporate behavior trends.
# - Actual training data and monetary cost assignments could be more complex.
# - Multiple models for each type of churn could be needed.
#
# Regardless of additional complexity, similar principles described in this notebook are likely apply.
# ### Optimizing model for prediction using Neo API
# Neo API allows to optimize our model for a specific hardware type. When calling `compile_model()` function, we specify the target instance family (C5) as well as the S3 bucket to which the compiled model would be stored.
#
# **Important. If the following command result in a permission error, scroll up and locate the value of execution role returned by `get_execution_role()`. The role must have access to the S3 bucket specified in ``output_path``.**
output_path = '/'.join(xgb.output_path.split('/')[:-1])
compiled_model = xgb.compile_model(target_instance_family='ml_c5',
input_shape={'data':[1, 69]},
role=role,
framework='xgboost',
framework_version='0.7',
output_path=output_path)
# ### Creating an inference Endpoint
#
# We can deploy this compiled model, note that we need to use the same instance that the target we used for compilation. This creates a SageMaker endpoint that we can use to perform inference.
#
# The arguments to the ``deploy`` function allow us to set the number and type of instances that will be used for the Endpoint. Make sure to choose an instance for which you have compiled your model, so in our case `ml_c5`. Neo API uses a special runtime (DLR runtime), in which our optimzed model will run.
# known issue: need to manually specify endpoint name
compiled_model.name = 'deployed-xgboost-customer-churn'
# There is a known issue where SageMaker SDK locates the incorrect docker image URI for XGBoost
# For now, we manually set Image URI
compiled_model.image = get_image_uri(sess.boto_region_name, 'xgboost-neo', repo_version='latest')
compiled_predictor = compiled_model.deploy(initial_instance_count = 1, instance_type = 'ml.c5.4xlarge')
# ### Making an inference request
# The compiled model accepts CSV content type:
compiled_predictor.content_type = 'text/csv'
compiled_predictor.serializer = csv_serializer
compiled_predictor.deserializer = None
# +
def optimized_predict(data, rows=500):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, compiled_predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
predictions = optimized_predict(test_data.as_matrix()[:, 1:])
# -
# ### (Optional) Clean-up
#
# If you're ready to be done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
sagemaker.Session().delete_endpoint(xgb_predictor.endpoint)
sagemaker.Session().delete_endpoint(compiled_predictor.endpoint)
| sagemaker_neo_compilation_jobs/xgboost_customer_churn/xgboost_customer_churn_neo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def test_exercise_84_1(x) -> bool:
import requests
from bs4 import BeautifulSoup
wiki_home = "https://en.wikipedia.org/wiki/Main_Page"
response = requests.get(wiki_home)
def decode_content(r,encoding):
return (r.content.decode(encoding))
contents = decode_content(response,encoding_check(response))
soup = BeautifulSoup(contents, 'html.parser')
return soup == x
def test_exercise_84_2(x) -> bool:
import requests
from bs4 import BeautifulSoup
wiki_home = "https://en.wikipedia.org/wiki/Main_Page"
response = requests.get(wiki_home)
def decode_content(r,encoding):
return (r.content.decode(encoding))
contents = decode_content(response,encoding_check(response))
soup = BeautifulSoup(contents, 'html.parser')
txt_dump=soup.text
return txt_dump == x
def test_exercise_84_3(x) -> bool:
return "str" == x
def test_exercise_84_4(x) -> bool:
import requests
from bs4 import BeautifulSoup
wiki_home = "https://en.wikipedia.org/wiki/Main_Page"
response = requests.get(wiki_home)
def decode_content(r,encoding):
return (r.content.decode(encoding))
contents = decode_content(response,encoding_check(response))
soup = BeautifulSoup(contents, 'html.parser')
txt_dump=soup.text
return txt_dump[10000:11000] == x
| Chapter07/unit_tests/.ipynb_checkpoints/Exercise 84-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sukrut11/Matplotlib-Tutorial/blob/main/Matplotlib.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="GBoczO9k2QMC"
import matplotlib.pyplot as plt
import numpy as np
# + id="Lua_W71y4h4f"
a = np.array([0, 50])
b = np.array([0, 200])
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="IBYNxOYK4sDt" outputId="caf6c0aa-b031-43c7-d191-44b70dfab878"
plt.plot(a, b)
# + [markdown] id="hsU86H2g48R9"
# ###Draw a line in a diagram from position (1, 3) to position (8, 10)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="ArbTudT44tdt" outputId="de8a6d0a-dd95-4b56-908e-23c304bb7131"
pos1 = np.array([1,8])
pos2 = np.array([3,10])
plt.plot(pos1, pos2)
plt.show()
# + [markdown] id="_DHG_CbAJJTg"
# Draw two points in the diagram, one at position (1, 3) and one in position (8, 10)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="KBPvoX6x5L68" outputId="3c6f43cb-bc87-4425-f983-2ecfcc6b5129"
pos1 = np.array([1,8])
pos2 = np.array([3,10])
plt.plot(pos1, pos2, "o")
plt.show()
# + [markdown] id="nWbHW_tpJpRv"
# Draw a line in a diagram from position (1, 3) to (2, 8) then to (6, 1) and finally to position (8, 10)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="uhZTsV4DJcQg" outputId="01658020-f549-42e9-f9a7-e21076ffdf54"
x = np.array([1,2,6,8])
y = np.array([3,8,1,10])
plt.plot(x , y)
plt.show()
# + [markdown] id="ny-XnipNKP2P"
# Plotting without x-points
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="klvaNb6RKE-P" outputId="e8eb20f5-1549-4f15-bcf9-fb0328fcf8d6"
y = np.array([7,4,2,1,5,8])
plt.plot(y)
plt.show()
# + [markdown] id="ztKaHQfBHMEv"
# ##Matplotlib Markers
# + [markdown] id="mW_fw6hHHWFl"
# You can use the keyword argument marker to emphasize each point with a specified marker
# + id="0d12qolAKhlF" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="f5523e2d-f4e3-4e36-d795-e2dbaab5e69c"
ypoints = np.array([5,6,4,2,1,3])
plt.plot(ypoints, marker = "o")
plt.show()
# + [markdown] id="HcMvGMfNKUES"
# 'o' Circle
# '*' Star
# '.' Point
# ',' Pixel
# 'x' X
# 'X' X (filled)
# '+' Plus
# 'P' Plus (filled)
# 's' Square
# 'D' Diamond
# 'd' Diamond (thin)
# 'p' Pentagon
# 'H' Hexagon
# 'h' Hexagon
# 'v' Triangle Down
# '^' Triangle Up
# '<' Triangle Left
# '>' Triangle Right
# '1' Tri Down
# '2' Tri Up
# '3' Tri Left
# '4' Tri Right
# '|' Vline
# '_' Hline
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="ppDSk-guHyKA" outputId="431b3fca-89e3-4187-eff3-057677d10b76"
plt.plot(ypoints, marker = "*")
plt.show()
# + [markdown] id="ObgQkiWnJBvS"
# ##Format Strings
# You can use also use the shortcut string notation parameter to specify the marker.
#
# This parameter is also called fmt, and is written with this syntax:
#
# marker | line | color
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="zXa1XlhPI6fj" outputId="bd6b0570-5b1d-42b4-acec-f6a4ed7e079a"
y = np.array([3,8,1,10])
plt.plot(y, "o:r")
plt.show()
# + [markdown] id="kudDhYRCKNyq"
# '-' Solid line
# ':' Dotted line
# '--' Dashed line
# '-.' Dashed/dotted line
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="T79T6hrhJ3E0" outputId="026c4e42-c39b-405a-8ead-0224271b0332"
plt.plot(y, "o-.r")
plt.show()
# + [markdown] id="IMcMLmTEKaKt"
# ##Color Reference
# + [markdown] id="OFQK7oRuKaCA"
# 'r' Red
# 'g' Green
# 'b' Blue
# 'c' Cyan
# 'm' Magenta
# 'y' Yellow
# 'k' Black
# 'w' White
# + [markdown] id="9aEP_JoMKoTt"
# ##Marker Size
# You can use the keyword argument markersize or the shorter version, ms to set the size of the markers:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="yBrYMqgSKD4B" outputId="64af09c2-b597-4010-cb4a-2a6555319a47"
ypoints = np.array([3,8,4,9])
plt.plot(ypoints, marker = "o", ms = 20)
plt.show()
# + [markdown] id="VPGN5JHzLPwD"
# ##Marker Color
# You can use the keyword argument markeredgecolor or the shorter mec to set the color of the edge of the markers:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="SpJiXSoFLI1Q" outputId="c37543cf-30a4-4aa7-d384-91fa7c008330"
plt.plot(ypoints, marker = "o", ms = 20, mfc = "r")
plt.show()
# + [markdown] id="I_1FSOZRLiFN"
# Use both the mec and mfc arguments to color of the entire marker:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="hHfoRhYbLZ8n" outputId="07f29f06-82da-4cb2-aeb7-a5903e5969f0"
plt.plot(ypoints, marker = "o", ms = 20, mfc = "r", mec = "g")
plt.show()
# + [markdown] id="jBLZXWmhLymU"
# You can also use Hexadecimal color values:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="w7Sr-pEELsnZ" outputId="b83e606d-84e0-444a-f1c6-65c779b76e7f"
plt.plot(ypoints, marker = "o", ms = 20, mfc = "#fffe00", mec = "#ff0100")
plt.show()
# + [markdown] id="vAIlABAGQRzl"
# Mark each point with the color named "hotpink":
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="R0v45fUwP8zc" outputId="164568f0-330e-426e-c9e4-a0183164af89"
plt.plot(ypoints, marker = "o", ms = 20, mfc = "hotpink", mec = "hotpink")
plt.show()
# + [markdown] id="Q50iWPX5QitR"
# ##Matplotlib Line
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="u-odOAi5QcaA" outputId="3b4349d5-f70d-48b5-dd5f-87856efc69e1"
ypoints = np.array([3, 8, 1, 10])
plt.plot(ypoints, ls = "dotted")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="o0m3-td-Qvvk" outputId="65ac8313-2d9f-4b02-eb5b-040204d97980"
plt.plot(ypoints, ls = "dashed")
plt.show()
# + [markdown] id="bSe4IR8HRL_b"
# 'solid' (default) '-'
# 'dotted' ':'
# 'dashed' '--'
# 'dashdot' '-.'
# 'None' '' or ' '
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="Z1glAD9yQ30H" outputId="7fd852f8-d7c3-4103-9bd0-603b72770aa0"
plt.plot(ypoints, marker = "o", ms = 20, mec = "red", mfc = "yellow", ls = "-.")
plt.plot()
# + [markdown] id="Q6VcKLSQR6Cb"
# ##Line Color
# You can use the keyword argument color or the shorter c to set the color of the line:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="ZZDrEAWoR3uc" outputId="637f9b1e-2481-411f-badb-108840e1d122"
y_points = np.array([6,4,2,7,9])
plt.plot(y_points, color = "r")
plt.show()
# + [markdown] id="SxqurlMoSYkg"
# You can also use Hexadecimal color values:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="oIkWkG1ESK3Z" outputId="b0834156-2d7b-4925-939f-e6adffc7bb5f"
plt.plot(y_points, c = '#4CAF50')
plt.show()
# + [markdown] id="aivQMcQJSVn2"
# Or any of the 140 supported color names.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="7uZO-5U1SRxH" outputId="92abbcdb-02f1-4cce-c87e-24f3e70c91bc"
plt.plot(ypoints, c = 'hotpink')
plt.show()
# + [markdown] id="thekURG_TgHQ"
# ##Line Width
# You can use the keyword argument linewidth or the shorter lw to change the width of the line.
#
# The value is a floating number, in points:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="vnwIHyuVSeY6" outputId="d42bf79d-2764-48fc-aec6-cab66851d16c"
ypoints = np.array([5,6,8,9,1,2,3])
plt.plot(ypoints, lw = "20.5")
plt.show()
# + [markdown] id="41clNyL7UN7K"
# ##Multiple Lines
# You can plot as many lines as you like by simply adding more plt.plot() functions:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="UM_Y6qaWT3Se" outputId="beae3063-eb11-47b4-d8b2-0c3d6b48a686"
y1 = np.array([3, 8, 1, 10])
y2 = np.array([6, 2, 7, 11])
plt.plot(y1)
plt.plot(y2)
plt.show()
# + [markdown] id="OmHrZihNUWIm"
# Draw two lines by specifiyng the x- and y-point values for both lines:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="amR7VG4TUGh6" outputId="33dbda68-3a0a-4469-da0d-1506ba70ca87"
x1 = np.array([0, 1, 2, 3])
y1 = np.array([3, 8, 1, 10])
x2 = np.array([0, 1, 2, 3])
y2 = np.array([6, 2, 7, 11])
plt.plot(x1, y1, x2, y2)
plt.show()
# + [markdown] id="BCnJFV4tzzB7"
# ## Labels
# + [markdown] id="V0Vu8tcVz3_O"
# Add labels to the x- and y-axis
# + id="4Qs0_Pb1UiCV"
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="JY6BgPja0Anx" outputId="0e637b64-906e-43b1-8d58-b799dbb43bc7"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.show()
# + [markdown] id="HHXYbytR0fNl"
# ##Create a Title for a Plot
# With Pyplot, you can use the title() function to set a title for the plot.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="EDQly8Xj0Xvi" outputId="01d09b52-0cc2-4d55-8ac1-36dac54a1e49"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data")
plt.show()
# + [markdown] id="FfW8ZlS51ASd"
# ##Set Font Properties for Title and Labels
# You can use the fontdict parameter in xlabel(), ylabel(), and title() to set font properties for the title and labels.
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="Gbnd3NT30raN" outputId="16cef1b1-7c50-4ceb-8e5d-e10855629188"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
font1 = {"family": "serif", "color": "blue", "size": 20}
font2 = {"family": "serif", "color": "darkred", "size": 15}
plt.title("Sports Watch Data", fontdict = font1)
plt.xlabel("Average Pulse", fontdict = font2)
plt.ylabel("Calorie Burnage", fontdict = font2)
plt.plot(x, y)
plt.show()
# + [markdown] id="9boLuCLC2FRa"
# ##Position the Title
# You can use the loc parameter in title() to position the title.
#
# Legal values are: 'left', 'right', and 'center'. Default value is 'center'.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="iM8L2X7M1_Kd" outputId="57addf42-4d6e-462b-9b12-8a98144a55dd"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data", loc = "left")
plt.show()
# + [markdown] id="9KKmRDdI2bak"
# ## Grid Lines
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="Z0R5t8Pe2TRr" outputId="9f33ab99-d090-4c3e-f892-b445cf93c79a"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data", loc = "center")
plt.grid()
plt.show()
# + [markdown] id="Hew8vyhz2q4K"
# ##Specify Which Grid Lines to Display
# You can use the axis parameter in the grid() function to specify which grid lines to display.
#
# Legal values are: 'x', 'y', and 'both'. Default value is 'both'.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="rzy2L66N2l0S" outputId="64c23a07-24ff-4b61-d279-1f9f663671d0"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data", loc = "center")
plt.grid(axis = "x")
plt.show()
# + [markdown] id="5HjAd-Tl26bA"
# Display only grid lines for the y-axis
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="7DrUcNpl2xo0" outputId="2d89f121-c52d-4a91-8f33-ea274744b211"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data", loc = "center")
plt.grid(axis = "y")
plt.show()
# + [markdown] id="QVt_g98V3EYH"
# ##Set Line Properties for the Grid
# You can also set the line properties of the grid, like this: grid(color = 'color', linestyle = 'linestyle', linewidth = number).
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="3n4hM28J2-eZ" outputId="ac8a4c71-b14a-498d-fbcf-56c5a53146bd"
x = np.array([80, 85, 90, 95, 100, 105, 110, 115, 120, 125])
y = np.array([240, 250, 260, 270, 280, 290, 300, 310, 320, 330])
plt.plot(x, y)
plt.xlabel("Average Pulse")
plt.ylabel("Calorie Burnage")
plt.title("Sports Watch Data", loc = "center")
plt.grid(color = "green", linestyle = "dashdot", linewidth = 0.5)
plt.show()
# + [markdown] id="xhgZ664zGGHZ"
# ##Display Multiple Plots
# With the subplots() function you can draw multiple plots in one figure
# + id="pT6CnjDo3Wkr" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="6e7ebece-546c-4f99-d0fb-e3a30fcf1ec0"
import numpy as np
import matplotlib.pyplot as plt
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([5,2,6,1,8])
plt.subplot(1,2,1) # nrows, ncols, index
plt.plot(x , y)
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([10,20,30,40,50])
plt.subplot(1,2,2) # nrows, ncols, index
plt.plot(x,y)
plt.show()
# + [markdown] id="R1idvRGlHgS5"
# ##The subplots() Function
# The subplots() function takes three arguments that describes the layout of the figure.
#
# The layout is organized in rows and columns, which are represented by the first and second argument.
#
# The third argument represents the index of the current plot.
# + [markdown] id="GvTVwaM9Hm51"
# ##Draw 2 plots on top of each other
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="juMRJLVSHP4l" outputId="65547269-0830-4a2f-c8b8-151d56efe2e9"
import numpy as np
import matplotlib.pyplot as plt
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([5,2,6,1,8])
plt.subplot(2,1,1) # nrows, ncols, index
plt.plot(x , y)
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([10,20,30,40,50])
plt.subplot(2,1,2) # nrows, ncols, index
plt.plot(x,y)
plt.show()
# + [markdown] id="8Ij-jxnHH1G6"
# ## Draw 6 plots
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="4TIbZLA3HtmE" outputId="6335cb2f-47ec-4ba6-d4ee-dc61d823ef11"
import numpy as np
import matplotlib.pyplot as plt
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([5,2,6,1,8])
plt.subplot(2,3,1) # nrows, ncols, index
plt.plot(x , y)
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([10,20,30,40,50])
plt.subplot(2,3,2) # nrows, ncols, index
plt.plot(x,y)
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([1,2,6,7,8])
plt.subplot(2,3,3) # nrows, ncols, index
plt.plot(x , y)
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([5,10,12,14,12])
plt.subplot(2,3,4) # nrows, ncols, index
plt.plot(x,y)
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([6,12,5,8,9])
plt.subplot(2,3,5) # nrows, ncols, index
plt.plot(x , y)
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([20,15,10,9,8])
plt.subplot(2,3,6) # nrows, ncols, index
plt.plot(x,y)
plt.show()
# + [markdown] id="xYohSvJjI2Fw"
# ##Title
# You can add a title to each plot with the title() function
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="9ctKC9uFIn8r" outputId="4eee336d-637f-4d69-cb31-1a85d4538f43"
import numpy as np
import matplotlib.pyplot as plt
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([5,2,6,1,8])
plt.subplot(1,2,1) # nrows, ncols, index
plt.plot(x , y)
plt.title("SALES")
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([10,20,30,40,50])
plt.subplot(1,2,2) # nrows, ncols, index
plt.plot(x,y)
plt.title("INCOME")
plt.show()
# + [markdown] id="h9idyHyqJYmf"
# ##Super Title
# You can add a title to the entire figure with the suptitle() function:
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="tm8Bq9mUJIC1" outputId="82da15fb-c7ef-4ffc-c51d-10ec9d922714"
import numpy as np
import matplotlib.pyplot as plt
# Plot 1
x = np.array([0,1,2,3,4])
y = np.array([5,2,6,1,8])
plt.subplot(1,2,1) # nrows, ncols, index
plt.plot(x , y)
plt.title("SALES")
# Plot 2
x = np.array([0,1,2,3,4])
y = np.array([10,20,30,40,50])
plt.subplot(1,2,2) # nrows, ncols, index
plt.plot(x,y)
plt.title("INCOME")
plt.suptitle("MY SHOP")
plt.show()
# + [markdown] id="f4XDD-siJr53"
# ##Creating Scatter Plots
# With Pyplot, you can use the scatter() function to draw a scatter plot.
#
# The scatter() function plots one dot for each observation. It needs two arrays of the same length, one for the values of the x-axis, and one for values on the y-axis
# + [markdown] id="1mFlGlWfJxCS"
# ##A simple scatter plot
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="AZaphFv5Jih9" outputId="2f517d15-5e41-4f48-efbd-fb2006cede94"
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
plt.scatter(x,y)
plt.show()
# + [markdown] id="x9ND276hKA6Q"
# The observation in the example above is the result of 13 cars passing by.
#
# The X-axis shows how old the car is.
#
# The Y-axis shows the speed of the car when it passes.
#
# Are there any relationships between the observations?
#
# It seems that the newer the car, the faster it drives, but that could be a coincidence, after all we only registered 13 cars.
# + [markdown] id="yS9eV0BNKFcN"
# ##Compare Plots
# In the example above, there seems to be a relationship between speed and age, but what if we plot the observations from another day as well? Will the scatter plot tell us something else?
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="WZFG-C9SJ41U" outputId="695eeb02-2867-4249-b052-cebefe257ed3"
#day one, the age and speed of 13 cars:
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
plt.scatter(x, y)
#day two, the age and speed of 15 cars:
x = np.array([2,2,8,1,15,8,12,9,7,3,11,4,7,14,12])
y = np.array([100,105,84,105,90,99,90,95,94,100,79,112,91,80,85])
plt.scatter(x, y)
plt.show()
# + [markdown] id="2u6qRDftKWPV"
# By comparing the two plots, I think it is safe to say that they both gives us the same conclusion: the newer the car, the faster it drives.
# + [markdown] id="T6qVm6ClLIQq"
# ##Colors
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="Bm3XEyETKOI_" outputId="7c6bf276-652d-4d2a-bc6b-e677037bc1b5"
#day one, the age and speed of 13 cars:
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
plt.scatter(x, y, color = "hotpink")
#day two, the age and speed of 15 cars:
x = np.array([2,2,8,1,15,8,12,9,7,3,11,4,7,14,12])
y = np.array([100,105,84,105,90,99,90,95,94,100,79,112,91,80,85])
plt.scatter(x, y, color = "#88c999")
plt.show()
# + [markdown] id="X0Ivl349LR50"
# ##Color Each Dot
# You can even set a specific color for each dot by using an array of colors as value for the c argument.
#
# **Note**: You cannot use the color argument for this, only the c argument.
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="wsLQ6TXjLD6g" outputId="fcaaac28-d63d-434e-93cf-7bedc8d5f969"
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
colors= np.array(["red","green","blue","yellow","pink","black","orange","purple","beige","brown","gray","cyan","magenta"])
plt.subplot(2,1,1)
plt.scatter(x,y,c= colors)
#day two, the age and speed of 15 cars:
x = np.array([2,2,8,1,15,8,12,9,7,3,11,4,7,14,12])
y = np.array([100,105,84,105,90,99,90,95,94,100,79,112,91,80,85])
colors = np.array(["blue","yellow","red","green","maroon","magenta","silver","gold","hotpink","cyan","brown","purple","tomato","grey","darkblue"])
plt.subplot(2,1,2)
plt.scatter(x,y,c = colors)
plt.suptitle("AGE VS SPEED")
plt.show()
# + [markdown] id="LEYfMKAFNH5t"
# ##ColorMap
# The Matplotlib module has a number of available colormaps.
#
# A colormap is like a list of colors, where each color has a value that ranges from 0 to 100.
#
# Here is an example of a colormap
#
# 
# + [markdown] id="z6c9RZErOfxj"
# This colormap is called 'viridis' and as you can see it ranges from 0, which is a purple color, and up to 100, which is a yellow color.
#
# ##How to Use the ColorMap
# You can specify the colormap with the keyword argument cmap with the value of the colormap, in this case 'viridis' which is one of the built-in colormaps available in Matplotlib.
#
# In addition you have to create an array with values (from 0 to 100), one value for each of the point in the scatter plot
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="9JKXeMh3MBJh" outputId="cc7e6cac-3835-4cb8-fd05-1b35d9030708"
import matplotlib.pyplot as plt
import numpy as np
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
colors = np.array([0, 10, 20, 30, 40, 45, 50, 55, 60, 70, 80, 90, 100])
plt.scatter(x, y, c=colors, cmap='viridis')
plt.show()
# + [markdown] id="PgK9VthZOxnG"
# You can include the colormap in the drawing by including the plt.colorbar() statement
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="EecUzYWlOtX3" outputId="5a711297-0d01-44a4-da63-ecb8226e0703"
import matplotlib.pyplot as plt
import numpy as np
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
colors = np.array([0, 10, 20, 30, 40, 45, 50, 55, 60, 70, 80, 90, 100])
plt.scatter(x, y,c= colors, cmap = "viridis")
plt.colorbar()
plt.show()
# + [markdown] id="3GFt6N22POza"
# ##Size
# You can change the size of the dots with the s argument.
#
# Just like colors, make sure the array for sizes has the same length as the arrays for the x- and y-axis
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="UDkoUvkDO_0D" outputId="94c684b2-4b08-41e3-d2c2-b161435a97fb"
import matplotlib.pyplot as plt
import numpy as np
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
colors = np.array([0, 10, 20, 30, 40, 45, 50, 55, 60, 70, 80, 90, 100])
sizes = np.array([20,50,100,200,500,1000,60,90,10,300,600,800,75])
plt.scatter(x, y, s= sizes, c = colors)
plt.colorbar()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="MAFokg_RQOEc" outputId="9d7fe0ad-e340-4f9e-b82d-0403417fdcb0"
import matplotlib.pyplot as plt
import numpy as np
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
sizes = np.array([20,50,100,200,500,1000,60,90,10,300,600,800,75])
plt.scatter(x, y, s= sizes)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="IH5qpK9PPjXm" outputId="8fc5810d-d495-4914-d451-9baa80e49b52"
import matplotlib.pyplot as plt
import numpy as np
x = np.array([5,7,8,7,2,17,2,9,4,11,12,9,6])
y = np.array([99,86,87,88,111,86,103,87,94,78,77,85,86])
sizes = np.array([20,50,100,200,500,1000,60,90,10,300,600,800,75])
plt.scatter(x, y, s=sizes, alpha = 0.5)
plt.show()
# + [markdown] id="yDofd2EYQbVY"
# ##Combine Color Size and Alpha
# You can combine a colormap with different sizes on the dots. This is best visualized if the dots are transparent
# + [markdown] id="Hrfsh8hfQfWL"
# ####Create random arrays with 100 values for x-points, y-points, colors and sizes
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="lyvQc0oBQIpf" outputId="2d18fc37-17ca-4f1b-e80f-eef17b371258"
x = np.random.randint(100, size= (100))
y = np.random.randint(100, size= (100))
colors = np.random.randint(100, size=(100))
sizes = 10 * np.random.randint(100, size=(100))
plt.scatter(x, y, c=colors, s=sizes, alpha=0.5, cmap='nipy_spectral')
plt.colorbar()
plt.show()
# + [markdown] id="u-iPGG5cD1YN"
# ##Creating Bars
# With Pyplot, you can use the bar() function to draw bar graphs
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="K3jqL5vVDlME" outputId="eff3290f-3321-4299-ef32-f10ec489b605"
import numpy as np
import matplotlib.pyplot as plt
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.bar(x,y)
plt.show()
# + [markdown] id="kAZcJZ1nEYLF"
# The bar() function takes arguments that describes the layout of the bars.
#
# The categories and their values represented by the first and second argument as arrays.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="NQJXw9dHDlJF" outputId="1016d00c-1ccc-4b7a-b352-23331ab6ec96"
x = ["APPLES", "BANANAS"]
y = [450, 516]
plt.bar(x,y)
plt.show()
# + [markdown] id="1TBfMN_SE0at"
# Draw 4 horizontal bars
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="TNXBWhhiDlGe" outputId="6cb96027-d0d7-4488-dc62-f2a72c61361b"
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.barh(x,y)
plt.show()
# + [markdown] id="4w5xfI7JFCOc"
# ##Bar Color
# The bar() and barh() takes the keyword argument color to set the color of the bars
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="2-eNklsdDlDd" outputId="bc7e5f3e-274c-44cd-fb65-01827617e9a6"
import numpy as np
import matplotlib.pyplot as plt
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.bar(x,y,color="red")
plt.show()
# + [markdown] id="BcanQvesFkAk"
# ##Color Names
# You can use any of the [140 supported color names](https://www.w3schools.com/colors/colors_names.asp).
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="27fuzDeHFbbd" outputId="c6338a6c-43ea-4a41-ee3f-f6cfa2328219"
import numpy as np
import matplotlib.pyplot as plt
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.bar(x,y,color="lightgreen")
plt.show()
# + [markdown] id="L1CWJXgGF3GV"
# ##Color Hex
# Or you can use [Hexadecimal color values](https://www.w3schools.com/colors/colors_hexadecimal.asp)
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="fksn8LlrF0vE" outputId="577e0418-38a1-4805-9aeb-5fd95a4ba0bc"
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.barh(x,y,color="#4CAF50")
plt.show()
# + [markdown] id="umKcHS0mGK-l"
# ##Bar Width
# The bar() takes the keyword argument width to set the width of the bars
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="P_9N6oisGHML" outputId="a7196d03-2150-40c7-e591-eb882aa594a5"
import numpy as np
import matplotlib.pyplot as plt
x = np.array(["A","B","C","D","E"])
y = np.array([24,65,32,56,48])
plt.bar(x,y,color="darkgreen",width = 0.1)
plt.show()
# + [markdown] id="75e39S_sGXPu"
# The default width value is 0.8
# + [markdown] id="1LXzXT7bGeKs"
# ##Bar Height
# The barh() takes the keyword argument height to set the height of the bars
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="saKgNMTCGSU-" outputId="fe190111-b719-42a4-b8a3-556e329d0cd3"
x = np.array(["A", "B", "C", "D"])
y = np.array([3, 8, 1, 10])
plt.barh(x,y,height= 0.1)
plt.show()
# + [markdown] id="YhUhmXrJg7OR"
# ##Creating Histograms
# A histogram is a graph showing frequency distributions.
#
# It is a graph showing the number of observations within each given interval.
#
# Example: Say you ask for the height of 250 people, you might end up with a histogram like this
#
# 
# + [markdown] id="bGN_UggJhpq6"
# You can read from the histogram that there are approximately:
#
# 1. 2 people from 140 to 145cm
# 2. 5 people from 145 to 150cm
# 3. 15 people from 151 to 156cm
# 4. 31 people from 157 to 162cm
# 5. 46 people from 163 to 168cm
# 6. 53 people from 168 to 173cm
# 7. 45 people from 173 to 178cm
# 8. 28 people from 179 to 184cm
# 9. 21 people from 185 to 190cm
# 10. 4 people from 190 to 195cm
# + [markdown] id="zp3VfcI9iDuy"
# ##Create Histogram
# In Matplotlib, we use the hist() function to create histograms.
#
# The hist() function will use an array of numbers to create a histogram, the array is sent into the function as an argument.
#
# For simplicity we use NumPy to randomly generate an array with 250 values, where the values will concentrate around 170, and the standard deviation is 10.
# + [markdown] id="RbNuY_S2ikkH"
# A Normal Data Distribution by NumPy
# + id="eK0LggkCGomE" colab={"base_uri": "https://localhost:8080/"} outputId="f9cd0756-83ea-4238-a2f9-e7ad8c9c9d57"
import numpy as np
x = np.random.normal(0, 10, 250)
print(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="KPJXyFy4i3O2" outputId="f6be5893-05c9-46a8-e9fa-5669908f2418"
plt.hist(x)
plt.show()
# + [markdown] id="0OAQ_lhYlk7Q"
# ##Creating Pie Charts
# With Pyplot, you can use the pie() function to draw pie charts
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="wZvs5rvxi_2i" outputId="0b564c88-cb0b-4aae-af71-3942df21222c"
y = np.array([12,23,45,65,32,45])
plt.pie(y)
plt.show()
# + [markdown] id="az8ZNkTJl6F-"
# As you can see the pie chart draws one piece (called a wedge) for each value in the array (in this case [35, 25, 25, 15]).
#
# By default the plotting of the first wedge starts from the x-axis and move counterclockwise
#
# 
# + [markdown] id="0VbI9fZumLna"
# Note: The size of each wedge is determined by comparing the value with all the other values, by using this formula:
#
# The value divided by the sum of all values: x/sum(x)
# + [markdown] id="863FfzcdmdNK"
# ##Labels
# Add labels to the pie chart with the label parameter.
#
# The label parameter must be an array with one label for each wedge
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="a95ju3yMl1WJ" outputId="e9e941b5-fa5c-4311-a4db-cd95413c25de"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
labels = ["Apples","Bananas","WaterMelon","Mangoes"]
plt.pie(y, labels = labels)
plt.show()
# + [markdown] id="sBYuanWcm94K"
# ##Start Angle
# As mentioned the default start angle is at the x-axis, but you can change the start angle by specifying a startangle parameter.
#
# The startangle parameter is defined with an angle in degrees, default angle is 0
#
# 
# + [markdown] id="VXBg5_vknYK0"
# Start the first wedge at 90 degrees
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="Vnd8gn_Mm4Qp" outputId="f6e49772-0fd9-4565-c4fe-af8f2f4a2019"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
plt.pie(y, labels = mylabels, startangle = 90)
plt.show()
# + [markdown] id="8Lwet4MVnnwa"
# ##Explode
# Maybe you want one of the wedges to stand out? The explode parameter allows you to do that.
#
# The explode parameter, if specified, and not None, must be an array with one value for each wedge.
#
# Each value represents how far from the center each wedge is displayed
# + [markdown] id="RGjr2Nl3nxLo"
# Pull the "Apples" wedge 0.2 from the center of the pie
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="JvTxc0Z9ngi3" outputId="8fa8573f-449d-4cd9-d13d-6af6f69f63af"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
myexplode = [0.2, 0, 0, 0]
plt.pie(y, labels = mylabels, explode = myexplode)
plt.show()
# + [markdown] id="EzwfWD_RoS7E"
# ##Shadow
# Add a shadow to the pie chart by setting the `shadows` parameter to `True`
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="uCSOxpdUoHZk" outputId="15edd676-8f06-4093-bb3e-cc77033b344e"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
myexplode = [0.2, 0, 0, 0]
plt.pie(y, labels = mylabels, explode = myexplode, shadow = True)
plt.show()
# + [markdown] id="MEsHVRGtoyK-"
# ##Colors
# You can set the color of each wedge with the colors parameter.
#
# The colors parameter, if specified, must be an array with one value for each wedge
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="7ehMaDwQoqFB" outputId="43635589-3b66-4ef6-f045-908b112dda06"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
mycolors = ["black", "hotpink", "b", "#4CAF50"]
plt.pie(y, labels = mylabels, colors = mycolors)
plt.show()
# + [markdown] id="2q1RBKgxpH1S"
# ##Legend
# To add a list of explanation for each wedge, use the legend() function
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="q_e62cMQo9t3" outputId="2e9d65f5-a06c-4555-924c-186e3682ec65"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
plt.pie(y, labels = mylabels)
plt.legend()
plt.show()
# + [markdown] id="QFgVxE-qpUuK"
# ##Legend With Header
# To add a header to the legend, add the title parameter to the legend function.
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="0KCs0lQJpP-8" outputId="8e6d8885-ce7e-4328-843c-86827a5cbbaf"
import matplotlib.pyplot as plt
import numpy as np
y = np.array([35, 25, 25, 15])
mylabels = ["Apples", "Bananas", "Cherries", "Dates"]
plt.pie(y, labels = mylabels)
plt.legend(title = "Fruits:")
plt.show()
| Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # microTVM Reference Virtual Machines
#
# **Author**: `<NAME> <<EMAIL>>`_
#
# This tutorial explains how to launch microTVM Reference Virtual Machines. You can use these to
# develop on real physical hardware without needing to individually install the microTVM
# dependencies. These are also particularly useful when trying to reproduce behavior with
# microTVM, such as when filing bug reports.
#
# microTVM is the effort to allow TVM to build and execute models on bare-metal microcontrollers.
# microTVM aims to be compatible with a wide variety of SoCs and runtime environments (i.e. bare metal,
# RTOS, etc). However, some stable software environment is needed to allow developers to share and
# reproduce bugs and results. The microTVM Reference Virtual Machines are intended to provide that
# environment.
#
# How it works
# ============
#
# No Virtual Machines are stored in the TVM repository--instead, the files stored in
# ``apps/microtvm/reference-vm`` describe how to build VMs to the Vagrant_ VM builder tool.
#
# The Reference VMs are split into two parts:
#
# 1. A Vagrant Base Box, which contains all of the stable dependencies for that platform. Build
# scripts are stored in ``apps/microtvm/reference-vm/<platform>/base-box``. TVM committers run
# these when a platform's "stable" dependencies change, and the generated base boxes are stored in
# `Vagrant Cloud`_.
# 2. A per-workspace VM, which users normally build using the Base Box as a starting point. Build
# scripts are stored in ``apps/microtvm/reference-vm/<platform>`` (everything except ``base-box``).
#
#
# Setting up the VM
# =================
#
# Installing prerequisites
# ------------------------
#
# A minimal set of prerequisites are needed:
#
#
# 1. `Vagrant <https://vagrantup.com>`__
# 2. A supported Virtual Machine hypervisor.
# `VirtualBox <https://www.virtualbox.org>`__ is one suggested free hypervisor, but please note
# that the `VirtualBox Extension Pack`_ is required for proper USB forwarding. If using VirtualBox,
# also consider installing the `vbguest <https://github.com/dotless-de/vagrant-vbguest>`_ plugin.
#
#
# First boot
# ----------
#
# The first time you use a reference VM, you need to create the box locally and then provision it.
#
# .. code-block:: bash
#
# # Replace zepyhr with the name of a different platform, if you are not using Zephyr.
# ~/.../tvm $ cd apps/microtvm/reference-vm/zephyr
# # Replace <provider_name> with the name of the hypervisor you wish to use (i.e. virtualbox).
# ~/.../tvm/apps/microtvm/reference-vm/zephyr $ vagrant up --provider=<provider_name>
#
#
# This command will take a couple of minutes to run and will require 4 to 5GB of storage on your
# machine. It does the following:
#
# 1. Downloads the `microTVM base box`_ and clones it to form a new VM specific to this TVM directory.
# 2. Mounts your TVM directory (and, if using ``git-subtree``, the original ``.git`` repo) into the
# VM.
# 3. Builds TVM and installs a Python virtualenv with the dependencies corresponding with your TVM
# build.
#
#
#
# Next, you need to configure USB passthrough to attach your physical development board to the virtual
# machine (rather than directly to your laptop's host OS).
#
# It's suggested you setup a device filter, rather than doing a one-time forward, because often the
# device may reboot during the programming process and you may, at that time, need to enable
# forwarding again. It may not be obvious to the end user when this occurs. Instructions to do that:
#
# * `VirtualBox <https://www.virtualbox.org/manual/ch03.html#usb-support>`__
# * `Parallels <https://kb.parallels.com/122993>`__
# * `VMWare Workstation <https://docs.vmware.com/en/VMware-Workstation-Pro/15.0/com.vmware.ws.using.doc/GUID-E003456F-EB94-4B53-9082-293D9617CB5A.html>`__
#
# Future use
# ----------
#
# After the first boot, you'll need to ensure you keep the build, in ``$TVM_HOME/build-microtvm``,
# up-to-date when you modify the C++ runtime or checkout a different revision. You can either
# re-provision the machine (``vagrant provision`` in the same directory you ran ``vagrant up`` before)
# or manually rebuild TVM yourself.
#
# Remember: the TVM ``.so`` built inside the VM is different from the one you may use on your host
# machine. This is why it's built inside the special directory ``build-microtvm``.
#
# Logging in to the VM
# --------------------
#
# The VM should be available to your host only with the hostname ``microtvm``. You can SSH to the VM
# as follows:
#
# .. code-block:: bash
#
# $ vagrant ssh
#
# Then ``cd`` to the same path used on your host machine for TVM. For example, on Mac:
#
# .. code-block:: bash
#
# $ cd /Users/yourusername/path/to/tvm
#
# Running tests
# =============
#
# Once the VM has been provisioned, tests can executed using ``poetry``:
#
# .. code-block:: bash
#
# $ poetry run python3 tests/micro/qemu/test_zephyr.py --microtvm-platforms=stm32f746xx
#
| _downloads/b953c88e73d1c677901980c0271abd49/micro_reference_vm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ojm_6E9f9Kcf"
# # MLP 108
# + id="hh6XplUvC0j0" outputId="6181ba91-39e6-4b56-b4bc-d0753cd5bcc3" colab={"base_uri": "https://localhost:8080/", "height": 54}
from google.colab import drive
PATH='/content/drive/'
drive.mount(PATH)
DATAPATH=PATH+'My Drive/data/'
PC_FILENAME = DATAPATH+'pcRNA.fasta'
NC_FILENAME = DATAPATH+'ncRNA.fasta'
# + id="VQY7aTj29Kch"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import StratifiedKFold
import tensorflow as tf
from tensorflow import keras
from keras.wrappers.scikit_learn import KerasRegressor
from keras.models import Sequential
from keras.layers import Bidirectional
from keras.layers import GRU
from keras.layers import Dense
from keras.layers import LayerNormalization
import time
dt='float32'
tf.keras.backend.set_floatx(dt)
EPOCHS=200
SPLITS=1
K=4
VOCABULARY_SIZE=4**K+1 # e.g. K=3 => 64 DNA K-mers + 'NNN'
EMBED_DIMEN=16
FILENAME='MLP108'
# + [markdown] id="WV6k-xOm9Kcn"
# ## Load and partition sequences
# + id="1I-O_qzw9Kco"
# Assume file was preprocessed to contain one line per seq.
# Prefer Pandas dataframe but df does not support append.
# For conversion to tensor, must avoid python lists.
def load_fasta(filename,label):
DEFLINE='>'
labels=[]
seqs=[]
lens=[]
nums=[]
num=0
with open (filename,'r') as infile:
for line in infile:
if line[0]!=DEFLINE:
seq=line.rstrip()
num += 1 # first seqnum is 1
seqlen=len(seq)
nums.append(num)
labels.append(label)
seqs.append(seq)
lens.append(seqlen)
df1=pd.DataFrame(nums,columns=['seqnum'])
df2=pd.DataFrame(labels,columns=['class'])
df3=pd.DataFrame(seqs,columns=['sequence'])
df4=pd.DataFrame(lens,columns=['seqlen'])
df=pd.concat((df1,df2,df3,df4),axis=1)
return df
# Split into train/test stratified by sequence length.
def sizebin(df):
return pd.cut(df["seqlen"],
bins=[0,1000,2000,4000,8000,16000,np.inf],
labels=[0,1,2,3,4,5])
def make_train_test(data):
bin_labels= sizebin(data)
from sklearn.model_selection import StratifiedShuffleSplit
splitter = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=37863)
# split(x,y) expects that y is the labels.
# Trick: Instead of y, give it it the bin labels that we generated.
for train_index,test_index in splitter.split(data,bin_labels):
train_set = data.iloc[train_index]
test_set = data.iloc[test_index]
return (train_set,test_set)
def separate_X_and_y(data):
y= data[['class']].copy()
X= data.drop(columns=['class','seqnum','seqlen'])
return (X,y)
def make_slice(data_set,min_len,max_len):
print("original "+str(data_set.shape))
too_short = data_set[ data_set['seqlen'] < min_len ].index
no_short=data_set.drop(too_short)
print("no short "+str(no_short.shape))
too_long = no_short[ no_short['seqlen'] >= max_len ].index
no_long_no_short=no_short.drop(too_long)
print("no long, no short "+str(no_long_no_short.shape))
return no_long_no_short
# + [markdown] id="nRAaO9jP9Kcr"
# ## Make K-mers
# + id="e8xcZ4Mr9Kcs"
def make_kmer_table(K):
npad='N'*K
shorter_kmers=['']
for i in range(K):
longer_kmers=[]
for mer in shorter_kmers:
longer_kmers.append(mer+'A')
longer_kmers.append(mer+'C')
longer_kmers.append(mer+'G')
longer_kmers.append(mer+'T')
shorter_kmers = longer_kmers
all_kmers = shorter_kmers
kmer_dict = {}
kmer_dict[npad]=0
value=1
for mer in all_kmers:
kmer_dict[mer]=value
value += 1
return kmer_dict
KMER_TABLE=make_kmer_table(K)
def strings_to_vectors(data,uniform_len):
all_seqs=[]
for seq in data['sequence']:
i=0
seqlen=len(seq)
kmers=[]
while i < seqlen-K+1 -1: # stop at minus one for spaced seed
#kmer=seq[i:i+2]+seq[i+3:i+5] # SPACED SEED 2/1/2 for K=4
kmer=seq[i:i+K]
i += 1
value=KMER_TABLE[kmer]
kmers.append(value)
pad_val=0
while i < uniform_len:
kmers.append(pad_val)
i += 1
all_seqs.append(kmers)
pd2d=pd.DataFrame(all_seqs)
return pd2d # return 2D dataframe, uniform dimensions
# + id="sEtA0xiV9Kcv"
def make_kmers(MAXLEN,train_set):
(X_train_all,y_train_all)=separate_X_and_y(train_set)
# The returned values are Pandas dataframes.
# print(X_train_all.shape,y_train_all.shape)
# (X_train_all,y_train_all)
# y: Pandas dataframe to Python list.
# y_train_all=y_train_all.values.tolist()
# The sequences lengths are bounded but not uniform.
X_train_all
print(type(X_train_all))
print(X_train_all.shape)
print(X_train_all.iloc[0])
print(len(X_train_all.iloc[0]['sequence']))
# X: List of string to List of uniform-length ordered lists of K-mers.
X_train_kmers=strings_to_vectors(X_train_all,MAXLEN)
# X: true 2D array (no more lists)
X_train_kmers.shape
print("transform...")
# From pandas dataframe to numpy to list to numpy
print(type(X_train_kmers))
num_seqs=len(X_train_kmers)
tmp_seqs=[]
for i in range(num_seqs):
kmer_sequence=X_train_kmers.iloc[i]
tmp_seqs.append(kmer_sequence)
X_train_kmers=np.array(tmp_seqs)
tmp_seqs=None
print(type(X_train_kmers))
print(X_train_kmers)
labels=y_train_all.to_numpy()
return (X_train_kmers,labels)
# + id="jaXyySyO9Kcz"
def make_frequencies(Xin):
# Input: numpy X(numseq,seqlen) list of vectors of kmerval where val0=NNN,val1=AAA,etc.
# Output: numpy X(numseq,65) list of frequencies of 0,1,etc.
Xout=[]
VOCABULARY_SIZE= 4**K + 1 # plus one for 'NNN'
for seq in Xin:
freqs =[0] * VOCABULARY_SIZE
total = 0
for kmerval in seq:
freqs[kmerval] += 1
total += 1
for c in range(VOCABULARY_SIZE):
freqs[c] = freqs[c]/total
Xout.append(freqs)
Xnum = np.asarray(Xout)
return (Xnum)
# + [markdown] id="j7jcg6Wl9Kc2"
# ## Build model
# + id="qLFNO1Xa9Kc3"
def build_model(maxlen,dimen):
act="sigmoid"
embed_layer = keras.layers.Embedding(
VOCABULARY_SIZE,EMBED_DIMEN,input_length=maxlen);
neurons=16
dense1_layer = keras.layers.Dense(neurons, activation=act,dtype=dt,input_dim=VOCABULARY_SIZE)
dense2_layer = keras.layers.Dense(neurons, activation=act,dtype=dt)
dense3_layer = keras.layers.Dense(neurons, activation=act,dtype=dt)
output_layer = keras.layers.Dense(1, activation=act,dtype=dt)
mlp = keras.models.Sequential()
#mlp.add(embed_layer)
mlp.add(dense1_layer)
mlp.add(dense2_layer)
#mlp.add(dense3_layer)
mlp.add(output_layer)
bc=tf.keras.losses.BinaryCrossentropy(from_logits=False)
print("COMPILE...")
mlp.compile(loss=bc, optimizer="Adam",metrics=["accuracy"])
print("...COMPILED")
return mlp
# + [markdown] id="LdIS2utq9Kc9"
# ## Cross validation
# + id="BVo4tbB_9Kc-"
def do_cross_validation(X,y,eps,maxlen,dimen):
model = None
cv_scores = []
fold=0
splitter = ShuffleSplit(n_splits=SPLITS, test_size=0.2, random_state=37863)
for train_index,valid_index in splitter.split(X):
X_train=X[train_index] # use iloc[] for dataframe
y_train=y[train_index]
X_valid=X[valid_index]
y_valid=y[valid_index]
print("BUILD MODEL")
model=build_model(maxlen,dimen)
print("FIT")
start_time=time.time()
# this is complaining about string to float
history=model.fit(X_train, y_train, # batch_size=10, default=32 works nicely
epochs=eps, verbose=1, # verbose=1 for ascii art, verbose=0 for none
validation_data=(X_valid,y_valid) )
end_time=time.time()
elapsed_time=(end_time-start_time)
fold += 1
print("Fold %d, %d epochs, %d sec"%(fold,eps,elapsed_time))
pd.DataFrame(history.history).plot(figsize=(8,5))
plt.grid(True)
plt.gca().set_ylim(0,1)
plt.show()
scores = model.evaluate(X_valid, y_valid, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# What are the other metrics_names?
# Try this from Geron page 505:
# np.mean(keras.losses.mean_squared_error(y_valid,y_pred))
cv_scores.append(scores[1] * 100)
print()
print("Validation core mean %.2f%% (+/- %.2f%%)" % (np.mean(cv_scores), np.std(cv_scores)))
return model
# + [markdown] id="_Q-PEh7D9KdH"
# ## Load
# + id="f8fNo6sn9KdH" outputId="119c729f-af79-45c7-9827-df402a5765dc" colab={"base_uri": "https://localhost:8080/", "height": 452}
print("Load data from files.")
nc_seq=load_fasta(NC_FILENAME,0)
pc_seq=load_fasta(PC_FILENAME,1)
all_seq=pd.concat((nc_seq,pc_seq),axis=0)
print("Put aside the test portion.")
(train_set,test_set)=make_train_test(all_seq)
# Do this later when using the test data:
# (X_test,y_test)=separate_X_and_y(test_set)
nc_seq=None
pc_seq=None
all_seq=None
print("Ready: train_set")
train_set
# + [markdown] id="qd3Wj_vI9KdP"
# ## Len 200-1Kb
# + id="mQ8eW5Rg9KdQ" outputId="7003fb8f-96b2-48ed-8957-585ede8293c5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
MINLEN=200
MAXLEN=1000
print ("Compile the model")
model=build_model(MAXLEN,EMBED_DIMEN)
print ("Summarize the model")
print(model.summary()) # Print this only once
print("Working on full training set, slice by sequence length.")
print("Slice size range [%d - %d)"%(MINLEN,MAXLEN))
subset=make_slice(train_set,MINLEN,MAXLEN)# One array to two: X and y
print ("Sequence to Kmer")
(X_train,y_train)=make_kmers(MAXLEN,subset)
X_train
X_train=make_frequencies(X_train)
X_train
print ("Cross valiation")
model1 = do_cross_validation(X_train,y_train,EPOCHS,MAXLEN,EMBED_DIMEN)
model1.save(FILENAME+'.short.model')
# + [markdown] id="OC68X4zr9KdU"
# ## Len 1Kb-2Kb
# + id="0nm3oU3h9KdV" outputId="1a6f7637-3e90-433b-fa2c-8aa681036705" colab={"base_uri": "https://localhost:8080/", "height": 1000}
MINLEN=1000
MAXLEN=2000
print ("Compile the model")
model=build_model(MAXLEN,EMBED_DIMEN)
print ("Summarize the model")
print(model.summary()) # Print this only once
print("Working on full training set, slice by sequence length.")
print("Slice size range [%d - %d)"%(MINLEN,MAXLEN))
subset=make_slice(train_set,MINLEN,MAXLEN)# One array to two: X and y
print ("Sequence to Kmer")
(X_train,y_train)=make_kmers(MAXLEN,subset)
X_train
X_train=make_frequencies(X_train)
X_train
print ("Cross valiation")
model2 = do_cross_validation(X_train,y_train,EPOCHS,MAXLEN,EMBED_DIMEN)
model2.save(FILENAME+'.medium.model')
# + [markdown] id="vyACRnZx9Kde"
# ## Len 2Kb-3Kb
# + id="kUxmLnQ-9Kde" outputId="0350f287-c865-4c80-92f3-3053b25236c9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
MINLEN=2000
MAXLEN=3000
print ("Compile the model")
model=build_model(MAXLEN,EMBED_DIMEN)
print ("Summarize the model")
print(model.summary()) # Print this only once
print("Working on full training set, slice by sequence length.")
print("Slice size range [%d - %d)"%(MINLEN,MAXLEN))
subset=make_slice(train_set,MINLEN,MAXLEN)# One array to two: X and y
print ("Sequence to Kmer")
(X_train,y_train)=make_kmers(MAXLEN,subset)
X_train
X_train=make_frequencies(X_train)
X_train
print ("Cross valiation")
model3 = do_cross_validation(X_train,y_train,EPOCHS,MAXLEN,EMBED_DIMEN)
model3.save(FILENAME+'.long.model')
| Workshop/MLP_108.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
# ## Hebbian Learner
#linearly separable data
x = [[1.0,2.5],[2.0,-1.0], [1.5,3.0], [0.0,-1.5], [-3.5,1.0],[2.5,0.0],[0.5,1.5],[0.0,-2.0]]
y = [1,0,1,0,1,0,0,0]
# +
def step(x, bi =False): # x is a vector (1 rank input)
ind_less_than_equal_0 = np.where(x<=0)
ind_more_than_0 = np.where(x>0) #using this instead of sets incase of multi rank inputs
z = x
z[ind_more_than_0] = 1
if(bi):
z[ind_less_than_equal_0] = -1
else:
z[ind_less_than_equal_0] = 0
return z
class neuron:
def __init__(self, x_shape, learning_rate = 0.01): #x_shape is number of inputs #x and y are 2 rank: batch_size*dims
self.xshape = x_shape
self.weights = self.init_weights(x_shape)
#self.bias = np.random.randn(1)
self.activation = step #(bi=True)
self.learning_rate = learning_rate
def init_weights(self, n0):
return np.random.randn(1,n0) #n1 is number of input neurons
def output(self, x): #equivalent to forward pass/activation
pred = self.activation(np.dot(x,np.transpose(self.weights)),False)# + self.bias)
return pred
def param_updates(self, preds, x):
#using formula that delta(weight_i) = learning_rate*(X_i*Y_i +|Y_i-X_i|)
#assume preds to be shape batch_size*1
#BELOW formula assumes weights are only updated after entire batch
self.weights = self.weights + self.learning_rate*np.average(preds*x) #on average activations were so and so
#self.bias = self.bias + self.lear
def training(self, x,y): #taking y although it is irrelevant for a pure hebbian learner
x = np.array(x)
preds = self.output(x)
preds = preds.reshape(preds.shape[0],1)
self.param_updates(preds, x)
# -
hebbian_neuron = neuron(2)
hebbian_neuron.training(x,y)
hebbian_neuron.weights
hebbian_neuron.output([-5,-1.5])
np.average(preds*x, axis=0)
preds*x
| Existing Neural net architectures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import utils
# 対象データファイル
DATA_FILES = [
'https://raw.githubusercontent.com/noko-noko-mk2/mgcm-data/main/data/%E3%83%A1%E3%82%A4%E3%83%B3%E3%82%B9%E3%83%88%E3%83%BC%E3%83%AA%E3%83%BC%E7%AC%AC2%E9%83%A8/%E7%AC%AC10%E7%AB%A0.tsv'
]
# 出力カラム(名称, 小数点以下桁数)
COLUMNS = [
('周回数', 0),
('魔法糸中', 3),
('魔法糸大', 3),
('ベル', 1),
('ジュエル', 5),
('プレミアガチャチケット', 7),
('SR以上限定プレミアガチャチケットのかけら', 5),
('1984エリザ', 7),
('2061エリザ', 7),
('クッキングエプロンエリザ', 7),
('ホリデーカジュアルエリザ', 7),
('ラウンジウェアエリザ', 7),
('俊敏の破宝珠', 3),
('精密の破宝珠', 3)]
df = utils.read_data(DATA_FILES)
# -
# ## ステージごとの総ドロップ集計
df.groupby('ステージ').sum()
# ## ステージごとの平均ドロップ
sum_df = df.groupby('ステージ').sum()
stage_avg = sum_df.divide(sum_df['周回数'], axis=0).drop(['周回数', '1周スタミナ'], axis=1)
stage_avg['周回数'] = sum_df['周回数']
stage_avg
with open('../reports/main_story_2_10.md', 'wt') as fp:
fp.write('| chapter | {} |\n'.format( ' | '.join([c for c, f in COLUMNS]) ))
fp.write('| -- | {} |\n'.format( ' | '.join(['--' for c, f in COLUMNS]) ))
for idx, row in stage_avg.iterrows():
fp.write('| {} | {} |\n'.format( idx, ' | '.join([format(row[c], '.%df' % f) for c, f in COLUMNS]) ) )
| notebooks/メインストーリー2章10ドロップ率.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright statement?
# # A New 'Transition'
# Welcome back! At this point in the course you are all likley aces at numerical analysis, expecially after 5 modules ranging from Diffusion, to Convection, to Conduction, in 1 or 2D, with various Neumann and Dirichlet Boundary conditions!
#
# But now it's time for a new "phase". In this lesson we begin looking at what happens when your nice rigid boundary conditions decide to start moving on you, essentially creating a moving boundary interface!
#
# A moving boundary interface is represented by numerous physical behaviors in real-world applications, from the polar ice caps melting, to the phase transformations of metal alloys, to the varying oxygen content of muscles near a clotted bloodvessel [2].
# 
# #### Real World Applications of a Moving Boundary Interface
# ### The Stefan Problem
# This new type of problem is known as the "Stefan Problem" as it was first studied by Slovene physicist <NAME> around 1890 [3]. Though his focus was primarily on the analysis of ice formations, nowadays his name is synonymous with the particular type of boundary value problem for PDEs where the boundary can move with time. Since the classic Stefan problem concentrated on the temperature distribution of a homogeneous material undergoing a phase transition, one of the most commonly studied Stefan problems today is the melting of Ice to Water!
# 
# #### <NAME> pioneered work into phase transitions of materials (ie Ice)
# ## A Review: 1D Heat Conduction
# Recall from both Modules 2 and 4 we took a loook at the Diffusion equation in 1D:
# $$\begin{equation}
# \frac{\partial U}{\partial t} = \alpha \frac{\partial^2U}{\partial x^2}
# \end{equation}$$
# Where we have the temperature distribution $U(x,t)$ and the thermal diffusivity $\alpha$. While before we looked at the conduction of heat through a graphite rod of length 1 meter, in this scenario we will analyze heat conduction though a 1D rod of ice. Let's first list some basic coefficients associated with the new material:
#
# ##### Thermal Properties of Ice at ~$0^{\circ}C$:
# Thermal Conductivity: $k = 2.22 \frac{W}{mK}$
#
# Density: $\rho \approx 1000 \frac{kg}{m^3}$
#
# Specific Heat: $c_{p}= 2.05x10^{-3} \frac{J}{kgK}$
#
# and lastly, Thermal Diffusivity: $\alpha = \frac{k_{ice}}{\rho_{ice}c_{p_{ice}}} = 1.083x10^{-6} \frac{m^2}{sec}$
#
# Melting Temperature of Ice is: $T_{melt}=0^{\circ}C$
#
# Okay! With that out of the way, let's look at the temperature distribution across the a rod of ice of length 1m with some basic initial and boundary conditions. For this first scenario, we will not take into account phase transition if temperatures hit $T_{melt}$, and will therefore assume a static boundary condition.
# ##### Problem Setup
# Governing Equation: $$\begin{equation}
# \frac{\partial U}{\partial t} = \alpha \frac{\partial^2U}{\partial x^2}
# \end{equation}$$
#
# Boundary Conditions:
#
# LHS: $\frac{\partial U}{\partial x} = -e^{\beta t}$, @x=0, t>0, time-dependant (increasing) heat flux in
#
# RHS: $\frac{\partial U}{\partial x} = 0$, @x=L, t>0, Insulated end, no heat flux in/out.
#
#
# Initial Conditions:
#
# $U(x,t) = -10^{\circ}C$, for 0<x<L, t=0
#
# Lets start coding!!
#
import numpy
from matplotlib import pyplot
# %matplotlib inline
from matplotlib import rcParams
rcParams['font.family']= 'serif'
rcParams['font.size']=16
from IPython.display import Image
# Now let's define a function to run the governing equations using the Forward-Time/Central-Difference Method discretization
def FTCS(U, nt, dt, dx, alpha, beta):
for n in range(nt):
Un=U.copy()
U[1:-1] = Un[1:-1] + alpha*(dt/dx**2)*(Un[2:]-2*Un[1:-1]+Un[0:-2])
#Boundary Conditions
U[-1]=U[-2] #RHS Insulated BC
t=n*dt #Increasing time = n*timestep(dt)
U[0]=numpy.exp(beta*t) #LHS Time dependant heat input in, BC
return U
# +
#Basic Parameters and Initialization
# Temperature scale: Celsius
# Length scale: meters
# Mass scale: kg
# Time scale: seconds
# Energy scale: Joules
# Power scale: Watts
L = 1 # Length of my ice rod
nt = 40000 # Number of timesteps
nx = 51 # Number of grid space steps
alpha = 1.083e-6 # Thermal Diffusivity
dx = L/(nx-1) # grid spacing in "x"
Ui = numpy.ones(nx)*(-10) # initialized Temperature array
beta = 0.001 #Growth Factor of my Temperature input
sigma = 1/2 # Stability*
dt=0.1 # Timestep chosen to be 0.1 seconds*
# -
# A word on Stability:
#
# Recall from module 2 we had the Courant-Friedrichs-Lewy (CFL) Stability condition for the FTCS Diffusion equation in the form:
# $$\sigma = \alpha \frac{\Delta t}{\Delta x^{2}} \leq \frac{1}{2} $$
#
# re-arranging to determine the upper limit for a time-step (dt) we have:
# $$\Delta t \leq \frac{\sigma \Delta x^{2}}{\alpha} \approx 923 seconds$$
#
# As you can see, chosing a time-step (dt) equal to $0.1$ seconds, we more than satisfy this CFL Stability condition.
#
x = numpy.linspace(0, 1, nx)
print('Initial ice rod temperature distribution')
pyplot.plot(x, Ui, color = '#003366', ls = '-', lw =4)
pyplot.ylim(-20, 60)
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
U = FTCS(Ui.copy(), nt, dt, dx, alpha, beta)
print('Total elapsed time is ', nt*dt, 'seconds, or', nt*dt/60, 'minutes' )
pyplot.plot(x, U, color = '#003366', ls = '-', lw =3)
pyplot.ylim(-20, 60)
pyplot.xlim(0, 1)
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
# The above figure shows us the temperature distribution of the ice rod at a point in time approximately 1 hour into applying our heat source. Now this would be all well and good if ice didn't melt (and therefore change phase) at zero degrees. But it does!!
#
# Let's build a rudimentary function to see what portion of the rod should be water by now, and which part should still be ice:
def Phase_graph(U, x, nx):
phase=numpy.ones(nx)*(-100)
for n in range(nx):
if U[n]>0:
phase[n]=100
return phase
pyplot.plot(x, U, color = '#003366')
pyplot.ylabel('Temperature', color ='#003366')
Phase1=numpy.ones(nx)
Phase=Phase_graph(U, x, nx)
pyplot.plot(x, Phase, color = '#654321', ls = '-', lw =3)
pyplot.ylim(-12, 10)
pyplot.xlim(0, 0.3)
pyplot.xlabel('Approx Water-Ice Interface', color = '#654321')
# As you can see, the ice SHOULD have melted about 0.07 meters (or 2.75 inches) into our rod. In reality, our boundary interface has moved from x=0 to the right as time elapsed. Not only should our temperature distribution profile change due to the differences in properties ($\rho, k, c_{p}$), but also the feedback from the moving boundary condition.
# ## Solutions to the Dimensionless Stefan problem:
# ### Analytical Solution:
# Before we continue we need to make some simplfications:
#
# 1) No convection, heat transfer is limited to conduction,
#
# 2) Pressure is constant,
#
# 3) Density does not change between the solid and liquid phase (Ice/Water), ie $\rho_{ice}=\rho_{water}\approx 1000 \frac{kg}{m^3}$,
#
# 4) The phase change interface at $s(t)$ has no thickness
#
# Looking closer at the problem at hand, we see that our temperature distribution must conform to the below figure:
# 
# #### Phase Change Domain
# As you can see, our temperature distribution experiences a discontinuity at our solid-liquid interface at x=s(t). Furthermore, this boundary moves to the right as time elapses. If we wish to analyze the distribution in one region or the other, we need to take into account a growing or shrinking domain, and therefore, the boundary of the domain has to be found as part of the solution [2]. In the Melting Problem depicted above, a moving interface separates the liquid and solid phases. The displacement of the boundary interface (denoted as $\dot{s}$) is driven by the heat transport through it [2]. The relationship between the moving interface, s(t), and the temperature distribution through it was developed by and is known as the Stefan Condition (or Stefan Equation) and takes the form of the following boundary conditions:
#
# when x =s(t),
# ###### [1, 3, 2]
# $$\dot{s} = \frac{ds}{dt} = -\frac{\partial U}{\partial x}$$
# $$U=0$$
# There are many solution methods for the Stefan problem, for this lesson we will focus on finding the temperature distribution in the liquid region only (ie 0<x<s(t)). Critical parameters to track are both temperature, U(x,t), and the position of the interface, s(t). Let's first solve this problem analytically:
# To make our lives easier we want a simplified version of the Stefan Problem, In his paper, Crowley [4] presented that both Oleinik and Jerome demonstrated how using an appropriate dimensionless model, and appropriate boundary conditions, and initial conditions, not only could one determine a general solution for the diffusion equation with the Stefan condition, but also that an explicit finite difference scheme converges to this solution [4]. For our dimensionless model we will set $\alpha=1, \beta=1$ to get the following governing equations, and boundary conditions:
# #### Dimensionless Stefan Problem Equations
# $$\frac{\partial U}{\partial t} = \frac{\partial^{2} U}{\partial x^{2}}$$
#
# $$s(t)$$
#
# (1) $\frac{\partial U(x=0, t)}{\partial x} = -e^{t}$, LHS BC (Heat input into the system)
#
# (2) $\frac{\partial U(x=s(t), t)}{\partial x} = -\frac{d s(t)}{d t}$, RHS BC, Stefan Condition
#
# (3) $U(x=s(t), t) = T_{melt}=0$, By definition of the Melting interface
#
# (4) s(t=0)=0, initial condition
#
# These equations set up the new below figure:
# 
# Because we want to be able to judge the accuracy of our numerical analysis (and thanks to Crowley, Oleinik, and Jerome!) we next find the general solution of this problem by means of Separation of Variables:
#
# SOV: $U(x,t)=X(x)T(t)$, gives us a general solution of the form:
#
# $$U(x,t) = c_{1}e^{c_{2}^{2}t-c_{2}x} +c_{3}$$
#
# from BC(1) we get $c_{2}=1, c_{1}=1, yielding: U(x,t) = e^{t-x} +c_{3}$
#
# from BC(2) we get: $\dot{s}=\frac{\partial s(t)}{\partial t} = -\frac{\partial U(x=s(t), t)}{\partial x}\rightarrow -e^{t-s(t)}= \frac{ds(t)}{d t}$, this yields that the only solution fo s(t) is: $$s(t)=t$$
#
# from BC(3) we get: $U(s(t),t) + c_{3} = 0, \rightarrow e^{t-t} +c_{3}=0, \rightarrow c_{3}=-1$,
#
# Finally we get an exact solution for the temperature distribution:
#
# $$U(x,t) = e^{t-x}-1$$, $$s(t) = t$$
#
# and this satisfies the initial condition IC(4) that s(t=0)=0!!
#
# And now you see why we chose the heat input function to be a time-dependant exponential, if you don't believe me that the analytical solution would be much nastier if you had chosesn say, a constant heat input, give SOV a try and let me know how it goes! Let's graph this solution and see if it makes sense.
def Exact_Stefan( nt, dt, x):
U=numpy.ones(nx)
for n in range(nt):
U = numpy.exp(n*dt-x)-1
return U
# +
dt_A=0.002
nt_A=500
ExactS=Exact_Stefan(nt_A, dt_A, x)
Max_tempA=max(ExactS)
pyplot.plot(x, ExactS, color = '#003366', ls = '--', lw =1)
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
print('Analytically, this is our temperature profile after:', nt_A*dt_A,'seconds')
print('The temperature at our LHS boundary is:', Max_tempA, 'degrees')
# -
# Does the solution look right? YES! Remember, this temperature profile is "Dimensionless" so we can't compare it to our previous example (not only is the diffusivity constant vastly different, but we are looking 1 second into the diffusion vs 1 hr!). Also it accounts ONLY for the temperature distribution in the liquid, assuming an ever increasing domain due to an ever expanding RHS boundary. Not only did we expect temperature to be highest at the input side, but our moving boundary interface, s(t), moves to the right with time and always hits our melting temperature ($0^{\circ} c$) at x=1, just as one would expect.
# ### Numerical Solution: The Variable Grid Method
# What makes this problem unique is that we must now track the time-dependant moving boundary. Now, for previous numerical analysis in a 1D domain, you had a constant number of spatial intervals nx (or N) between your two boundaries x=0, and x=L. Thus your spatial grid steps dx were constant and defined as $dx = L/(nx-1)$. But now you have one fixed bounary and one moving boundary, and your domain increases as time passes. [1]
#
# 
#
#
#
#
# One of the key tenements of the Variable Grid Method for solving the Stefan probem is that you keep the NUMBER of grid points (nx) fixed, thus your grid SIZE (dx) will increase as the domain increases. Your grid size now varies dependant on the location of the interaface front, dx = s(t)/N.
#
# $$dx = \frac{L}{N-1} \longrightarrow dx = \frac{s(t)}{N}$$
#
# 
#
# Now, while one might be tempted to view the figure above as the new "Variable Grid" stencil, one must remember that the "Y"-axis is time, and therefore as you move up you are moving forward in time, therefore a more accurate depiction of the FTCS method in stencil form for the Variable Grid method would be:
#
# 
#
# and so it becomes clear - that the our spatial step ($dx$) will be depentant on our time step ($m$)!!
# #### Derivation of the new Governing Equation
# Now let's set up the new governing equations to be discretized for our code. We know that the 1D Diffusion equation must be valid for all points on our spatial grid so we can re-write the LHS of equation:
# $$\frac{\partial U}{\partial t} = \frac{\partial^{2} U}{\partial x^{2}}$$
# to be $$ \frac{\partial U_{i}}{\partial t} = \frac{\partial U_{@t}}{\partial x}\frac{dx_{i}}{dt} + \frac{\partial U_{@x}}{\partial t}$$
#
# we can use the expression: $$\frac{dx_{i}}{dt} = \frac{x_{i}}{s(t)}\frac{ds}{dt}$$ to track the movement of the node $i$.
#
# Substituting these into the diffusion equation (and droping the i,t, and x indices since they are constant) we obtain a new governing equation for diffusion:
#
# $$\frac{\partial U}{\partial t} = \frac{x_{i}}{s}\frac{ds}{dt}\frac{\partial U}{\partial x} + \frac{\partial^{2}U}{\partial x^{2}}$$
#
# This is subject to the boundary and initial conditions (BC1-BC3, IC4) as stated for the 1D dimensionless Stefan problem above.
# #### Discretization
#
# and now we seek to discretize the new governing equations. For this code we will implement an explicit, FTCS scheme, with parameters taking a Taylor expansion centered about the node ($x_{i}^{m}$), and time ($t_{m}$) just like before. In the above equations we discritized the time derivatives of U using forward time, and the spatial derivitaives using centered space. We re-write ds/dt as $\dot{s}$ and leave it a variable for now. (EXERCISE: See if you can discretize this equation form memory!!)
#
# Taylor Expand and Discretize: (moving from left to right)
#
# $\frac{\partial U}{\partial t} \longrightarrow \frac{U_{i}^{m+1}-U_{i}^{m}}{\Delta t}$,
#
# $\frac{x_{i}}{s} \longrightarrow \frac{x_{i}^{m}}{s_{m}}$,
#
# $\frac{ds}{dt} \longrightarrow \dot{s_{m}}$,
#
# $\frac{\partial U }{\partial x} \longrightarrow \frac{U_{i+1}^{m}-U_{i-1}^{m}}{2\Delta x^{m}}$,
#
# $\frac{\partial^{2}U}{\partial x^{2}} \longrightarrow \frac{U_{i+1}^{m}-2U_{i}^{m}+U_{i-1}^{m}}{(\Delta x^{m})^{2}}$
#
# And now we substitute, rearrange and solve for $U_{i}^{m+1}$ to get:
#
# $U_{i}^{m+1}=U_{i}^{m}+\frac{\Delta t x_{i}^{m} \dot{s_{m}}}{2 \Delta x^{m}s_{m}}(U_{i+1}^{m}-U_{i-1}^{m}) + \frac{\Delta t}{(\Delta x^{m})^{2}}(U_{i+1}^{m}-2U_{i}^{m}+U_{i-1}^{m})$
#
#
# Great!! We're almost ready to start coding, but before we begin, do you notice a problem with this expression? What about if i=0? Plug it in and you will see that we have expressions of the form $U_{-1}^{m}$ in both right hand terms, but that can't be right, looking at our stencil we see that $i=-1$ is off our grid! This is where the boundary conditions come in!
# Generate discretization expressions for the boundary conditions at x=0 (LHS) and x=s(t) (RHS). For the RHS and the temperature gradient across the moving boundary interface we will use a three-term backward difference scheme:
#
# LHS: $\frac{\partial U(x=0, t)}{\partial x} = -e^{t} \longrightarrow \frac{U_{i+1}^{m}(0,t)-U_{i-1}^{m}(0,t)}{2\Delta x^{m}} = e^{t_{m}}$, and
#
# RHS: $\frac{\partial U(x=s(t), t)}{\partial x} = -\frac{d s(t)}{d t} \longrightarrow \frac{\partial U(x=s(t),t)}{\partial x} = \frac{3U_{N}^{m}-4U_{N-1}^{m}+U_{N-2}^{m}}{2\Delta x^{m}} $
#
# from our LHS boundary condition expression, if we set $i=0$ and solve for $U_{-1}^{m}$ we get: $$U_{-1}^{m}=U_{1}^{m} + 2\Delta x^{m} e^{t_{m}}$$
# We can now combine this expression and substitute into the governing equation to get expressions for diffusion at $i=0, i=1$ to $(N-1)$, and $i=N$:
#
#
# $U_{i}^{m+1}=(1-2\frac{\Delta t}{(\Delta x^{m})^{2}})U_{i}^{m} + 2\frac{\Delta t}{(\Delta x^{m})^{2}}U_{i+1}^{m} + (2\frac{\Delta t}{\Delta x^{m}}-\frac{\Delta t x_{i}^{m}\dot{s_{m}}}{s_{m}})e^{t_{m}}$
#
#
# $U_{i}^{m+1}=U_{i}^{m}+\frac{\Delta t x_{i}^{m} \dot{s_{m}}}{2 \Delta x^{m}s_{m}}(U_{i+1}^{m}-U_{i-1}^{m}) + \frac{\Delta t}{(\Delta x^{m})^{2}}(U_{i+1}^{m}-2U_{i}^{m}+U_{i-1}^{m})$
#
# $U_{i}^{m}=0$, $i=N$
# Phew!! Now that is one impressive looking set of discretization expressions. But, sorry, we aren't done yet..... In the above expressions what are we supposed to do about $\dot{s_{m}}$ and $s_{m}$?
#
# Well luckily, that is where the Stefan condition comes in, remember:
#
# $\dot{s_{m}} = \frac{ds}{dt}= -\frac{\partial U_{@x=s}}{\partial x} = -\frac{3U_{N}^{m}-4U_{N-1}^{m}+U_{N-2}^{m}}{2\Delta x^{m}} $, and as for $s_{m}$ that is just a Heat Balance equation:
#
# $s_{m+1} = s_{m}+\dot{s_{m}}\Delta t = s_{m} - \frac{\Delta t}{2\Delta x^{m}}(3U_{N}^{m}-4U_{N-1}^{m}+U_{N-2}^{m})$, where $s_{0}=0$
#
# To make our lives easier, instead of inserting these expressions for $\dot{s_{m}}$ and $s_{m}$ into our governing equations, lets keep them as coupled expressions to be calculated during the time loops and calculated at every grid point $i$.
#
# lastly, lets not forget that the updated interface location $s_{m+1}$ and grid size $\Delta x^{m}$ are calculated at every timestep $m$, and have the relationship:
#
# $$\Delta x^{m+1} = \frac{s_{m+1}}{N}$$
#
# Don't worry, we are almost ready to code, I promise! There is just one more expression we need to consider, and that relates to Stability! Following the calculations of Caldwell and Savovic [5], for now, we will use something simple in order to limit the size of the timestep ($\Delta t$), that is: $$\frac{\Delta t}{(\Delta x^{m})^{2}} \leq 1$$, or
#
# $$\Delta t \leq (\Delta x^{m})^{2}$$
# ## Let's Code!!
# To summarize our governing equations we need to code these 7 coupled equations:
#
# (A) $u_{i}^{m+1}=(1-2\frac{\Delta t}{(\Delta x^{m})^{2}})u_{i}^{m} + 2\frac{\Delta t}{(\Delta x^{m})^{2}}u_{i+1}^{m} + (2\frac{\Delta t}{\Delta x^{m}}-\frac{\Delta t x_{i}^{m}\dot{s_{m}}}{s_{m}})e^{t_{m}}$
#
#
# (B) $u_{i}^{m+1}=u_{i}^{m}+\frac{\Delta t x_{i}^{m} \dot{s_{m}}}{2 \Delta x^{m}s_{m}}(u_{i+1}^{m}-u_{i-1}^{m}) + \frac{\Delta t}{(\Delta x^{m})^{2}}(u_{i+1}^{m}-2u_{i}^{m}+u_{i-1}^{m})$
#
# (C) $u_{i}^{m}=0$, $i=N$
#
# (D) $\dot{s_{m}} = \frac{ds}{dt}= -\frac{\partial u_{@x=s}}{\partial x} = -\frac{3u_{N}^{m}-4u_{N-1}^{m}+u_{N-2}^{m}}{2\Delta x^{m}} $
#
# (E) $s_{m+1} = s_{m}+\dot{s_{m}}\Delta t = s_{m} - \frac{\Delta t}{2\Delta x^{m}}(3u_{N}^{m}-4u_{N-1}^{m}+u_{N-2}^{m}), s_{0}=0$
#
# (F) $\Delta x^{m+1} = \frac{s_{m+1}}{N}$
#
# (G) $\Delta t \leq (\Delta x^{m})^{2}$
def VGM(nt, N, U0, dx, s, dt):
Uo=0 #Initial Temperature input
U0 = numpy.ones(N)*(Uo)
s0=0.02 #Initial Interface Position
#(cannot chose zero or our expressions would blow up!)
s=numpy.ones(nt)*(s0)
dx=numpy.ones(nt)*(s[1]/N)
sdot=(s[1]-s[0])/dt
s[1]= s[0] - (dt/(2*dx[0]))*(3*U0[N-1]-4*U0[N-2]+U0[N-3])
dx[0]=(s[1]/N)
for m in range(0, nt-1):
for i in range(0, N-1):
#LHS BC (x=0, i=0)
if(i==0):U0[i]= (1-2*(dt/dx[m]**2))*U0[i]+2*(dt/dx[m]**2)*U0[i+1] + (2*(dt/dx[m])-dt*(dx[m]*i)*sdot/s[m])*numpy.exp(m*dt)
#Governing Equation (B)
else: U0[i]= U0[i]+ ((dt*(dx[m]*i)*sdot)/(2*dx[m]*s[m]))*(U0[i+1]-U0[i-1]) + (dt/(dx[m]**2))*(U0[i+1]-2*U0[i]+U0[i-1])
#RHS BC (x=L, L=dx{m*N})
U0[N-1]=0
s[m+1]= s[m] - (dt/(2*dx[m]))*(3*U0[N-1]-4*U0[N-2]+U0[N-3]) #Heat Balance Equation, (E)
sdot=(s[m+1]-s[m])/dt #Updating Speed of Interface
dx[m+1]=(s[m+1]/N) #Updating dx for each time step m
if (U0[i]>0):
Location=s[m]
return U0, s, dx
# +
# First we set up some new initial parameters.These values differ from our first example since we are now dealing with
# a new non-dimensionalized heat equation, and new governing equations.
#1.0 Numerical Solution Parameters:
dt=2.0e-6 # Size of our Time-step, Set constant, Chosen in accordance with Caldwell and Savovic [5]
nt=500000 # Number of time steps, chosen so that elapsed solution will determine heat diffusion after
# (nt*dt= 1.0 seconds) Just like in our Analytical Solution.
N = 10 # Number of spatial grid points, Chosen, for now, in accordance with Caldwell and Savovic [5]
# -
U_VGM, s_VGM, dx_VGM = VGM(nt, N, U0, dx, s, dt)
print('Initial Position of the interface (So):', s_VGM[0])
print('Initial X_step(dx):', dx_VGM[0])
print('Time Step (dt):', dt)
# ### Results and Discussions
#
# Ok! Now lets see what we get, and more importantly, if it makes any sense:
# +
XX = numpy.linspace(0, 1, N)
Max_tempVGM=max(U_VGM)
pyplot.plot(XX,U_VGM, color = '#654322', ls = '--', lw =4)
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
pyplot.ylim(-.5, 2)
pyplot.xlim(0, 1.0)
print('This is our VGM temperature profile after:', nt*dt, 'seconds')
print('The temperature at our LHS boundary is:', Max_tempVGM, 'degrees')
print('Final position of our interface is:', s_VGM[m])
print('The final speed of our interface is:', sdot)
print('Our grid spaceing (dx) after', nt*dt, 'seconds is:', dx_VGM[m] )
# -
# Hey now! That looks pretty good! Let's compare this result to our earlier Analytical (exact) solution to the 1D Dimensionless problem, for 1 second into the diffusion:
# +
pyplot.figure(figsize=(12,10))
pyplot.plot(XX,U_VGM, color = '#654322', ls = '--', lw =4, label = 'Variable Grid Method')
pyplot.plot(x, ExactS, color = '#003366', ls = '--', lw =1, label='Analytical Solution')
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
pyplot.ylim(0, 1.8)
pyplot.xlim(0, 1.0)
pyplot.legend();
print('Max error (@x=0) is:',abs(Max_tempVGM - Max_tempA)*(100/Max_tempA),'percent')
# -
# Under 2% error is pretty good! What else should we verify? How about the change in the interface location s(t) and the size of our spatial grid (dx) over time?
# +
Time=numpy.linspace(0,dt*nt,nt)
pyplot.figure(figsize=(10,6))
pyplot.plot(Time,dx_VGM, color = '#660033', ls = '--', lw =2, label='Spatial Grid Size')
pyplot.plot(Time,s_VGM,color = '#222222', ls = '-', lw =4, label='Interface Position, s(t)')
pyplot.xlabel('Time (Seconds)')
pyplot.ylabel('Length')
pyplot.legend();
# -
# and so we see that both the spatial grid size, dx, and the interface location, s(t), increase with time over the course of 1 second. This is exactly what we would expect! Infact, remember back to our anayltical solution, we solved the interface function to be: $s(t)=t$, which is exactly the graph that our numerical solution has given us!
#
# Well, then! That's a wrap! Looks like everything is all accounted for, right?. . . . . . . . . . . . . .well, not exactly. There are still quite a few questions left unanswered, and now we must discuss the limitations in the Variable Grid Method of numerical implementation.
# ### Limitations of the Variable Grid Method
#
# A number of questions should be coming to mind, among those are:
#
# 1) Why do we only analyze for 1 second of diffusion?
#
# 2) Why is your VGM timestep (dt) so small at 2.0e-6?
#
# 3) Why is your number of spatial grid points (N) only 10?
#
#
# These are all VERY good questions, do you already know the answer? Here is a hint: it all has to do with stability!
#
# You see, we have a relatively benign stability statement: $\Delta t \leq (\Delta x^{m})^{2}$, but still, it is a crutial aspect of our numerical analysis. This statement limits the size of our timestep, dt, and is the key reason why we chose our initial, constant, dt to be 2.0e-6 (lets not also forget the fact that this dt and N were also chosen by Caldwell and Savovic [5]) . Remember, because we are using a Central Difference scheme, our stability criteria essentially comes from the CFL condition:
#
# $$\frac{\Delta t}{(\Delta x^{m})^{2}} \leq \sigma = 1$$.
#
# The parameters dt and dx are not just arbitrarilty chosen. They determine the speed of your numerical solution. At all times you need the speed at which your solution progresses to be faster than the speed at which the problem (in this case, thermal diffusion) propagates. Afterall, how can you calculate the solution numerically when the solution is faster than the numerical analysis?!
#
#
# Here is a pertinent question for you: Which do you determine first: dt, nt, N, or L? Does it matter which one you chose first?
# The answer is normally, yes, we chose N and L which gives us dx, we then use a stability statement with dx (and $\alpha$) to limit dt, we then choose our nt to determine the elapsed time into our solution in the form of (nt*dt) seconds, easy right?
#
# Well this is where we get to limitation #1 with the Variable Grid Method:
#
# Because the end of our domain "L" is in actuality the position of our interface, s(t), and our interface at time t=0 is approximately at the origin (x=0.02), then if we want a constant number of spatial grids N (say N=10), our initial dx is extremely small!. Remember governing equation (F) says: $\Delta x^{m+1} = \frac{s_{m+1}}{N}$, well at t=0 (m=0) s[0] is 0.02, meaning our initial dx, dx[0] is: 0.002. Plug this back into our stability statement and you see that dt must be smaller than or equal to 4e-6! That is VERY harsh criterion for a timestep dt. In essence, with SUCH a small time step, we would need hundreds of thousands of time loops, nt, in order to get just 1 second of data. Now you see why nt is chosen to be 500,000 in order to get just 1 second of data. If I had wanted more time elapsed, we would have needed iterations in the millions. (This is also why the analytical solution was calculated only for 1 second, we wanted to be able to compare results at the end!)
#
# Now limitation #2 of the Variable Grid Method: Because the interface starts at the origin, the initial spatial step is very small, and forces a very very small timestep, dt. Thus a very very large number of iterations is needed to get into the "Seconds" scale for simulations, this is a slow numerical process...
#
# Not only this, but we have limitation #3 of the Variable Grid Method: If we want a longer dimensional domain, we need to choose a larger N. This will give us a final domain of $L=N*dx_{final}$, but if N is larger, then going back to our governing equation (F), $\Delta x^{m+1} = \frac{s_{m+1}}{N}$ this means that dx[0] is even smaller, which continues to force a much smaller dt. Thus for the Variable Grid Method, if you want a large domain, you need an even smaller timestep, which forces more time in your numerical calculation.
#
#
# Let's test this out for ourselves, lets redo the VGM calculations but increase N to say, 14:
# (The cell below is in Raw-NB Convert since we didn't want it to slow you down the first time you ran this program. Notice the N is now 14, go ahead and run this below cell and see what happens)
# +
dt=2.0e-6
nt=500000
N = 14
U_VGM14, s_VGM14, dx_VGM14 = VGM(nt, N, U0, dx, s, dt)
XX = numpy.linspace(0, 1, N)
Max_tempVGM14=max(U_VGM14)
pyplot.plot(XX,U_VGM14, color = '#654322', ls = '--', lw =4, label = 'VGM')
pyplot.plot(x, ExactS, color = '#003366', ls = '--', lw =1, label='Analytical')
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
pyplot.ylim(-.5, 2)
pyplot.xlim(0, 1.0)
pyplot.legend();
print('Time Step dt is:', dt)
print('Initial spatial step, dx[0]^2 is:', dx_VGM14[0]**2 )
print('Max error (@x=0) is:',abs(Max_tempVGM14-Max_tempA)*(100/Max_tempA),'Percent')
# -
# As you can see, our stability criteria is still met, since $\Delta t \leq (\Delta x_{0})^{2}$. Infact, with a 40% larger N we see that we have less error as well! But what if we set N=15?
# (The cell below is in Raw-NB Convert since we didn't want it to slow you down the first time you ran this program. Notice the N is now 15, go ahead and run this below cell and see what happens THIS time)
# +
dt=2.0e-6
nt=500000
N = 15
U_VGM15, s_VGM15, dx_VGM15 = VGM(nt, N, U0, dx, s, dt)
XX = numpy.linspace(0, 1, N)
Max_tempVGM15=max(U_VGM15)
pyplot.plot(XX,U_VGM15, color = '#654322', ls = '--', lw =4, label = 'VGM')
pyplot.plot(x, ExactS, color = '#003366', ls = '--', lw =1, label='Analytical')
pyplot.xlabel('Length of Domain')
pyplot.ylabel('Temperature')
pyplot.ylim(-.5, 2)
pyplot.xlim(0, 1.0)
pyplot.legend();
print('Time Step dt is:', dt)
print('Initial spatial step, dx[0]^2 is:', dx_VGM15[0]**2 )
print('Max error (@x=0) is:',abs(Max_tempVGM15-Max_tempA)*(100/Max_tempA),'Percent')
# -
# Wow! That blew up! Just as we expect it would now that with N=15 our time step is larger than our initial spatial step!
# #### Final Thoughts
#
# And now we know, that the Stefan Problem, a boundary value PDE with a time-dependant moving boundary, CAN be solved. We have just demonstrated the solution for a 1D, Dimensionless Stefan Problem with specifically chosen (time-dependant) input heat flux in order to give us a simplified exact solution to compare with. You have also seen implementation of the Variable Grid Method, one of many ways inwhich one can numerical simulate the Stefan problem. But we have also seen the limitations, namely in small time-steps ($dt$), smaller numerical domain ($N\cdot dx_{f}$), and large number of iterations ($nt$) for just a small amount of analytical time (t=$nt \cdot dt$ seconds). Perhaps when it comes time for you to model the melting of the Polar Ice caps, you'll choose a more "expedient" method? Just make sure you get the answer BEFORE the caps melt...)
#
# EXERCISE #1: Can you implement the Variable Grid Method discretization governing equations to give us a solution faser? Can it handle millions of time iterations without putting us to sleep?
#
# EXERCISE #2: Can you write a stability condition statement that will maximize our time-step (dt) for a given N and still keep it constant?
# # References
# 1. <NAME>., The numerical solution of one-phase classical Stefan problem, Journal of Computational and Applied Mathematics 81 (1997) 135-144
#
# 2. <NAME>., A Comparison of Numeical Models for one-dimensional Stefan problems, Journal of Compuational and Applied Mathematics 192 (2006) 445-459
#
# 3. <NAME>., "Some historical notes about the Stefan problem". Nieuw Archief voor Wiskunde, 4e serie 11 (2): 157-167 (1993)
#
# 4. <NAME>., Numerical Solution of Stefan Problems, Brunel University, Department of Mathematics, TR/69 December 1976
#
# 5. <NAME>., Numerical Solution of Stefan Problem By Variable Space Grid Method and Boundary Immodbilisation Method, Jour. of Mathematical Sciences, Vol. 13 No.1 (2002) 67-79.
# Execute this cell to load the notebook's style sheet, then ignore it
from IPython.core.display import HTML
css_file ='numericalmoocstyle.css'
HTML(open(css_file, "r").read())
| Chris Tiu Project/Test2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Юнит-тестирование
# <br />
# ##### Основы
# Тестирование по целям:
# * functional
# * performance
# * usability
# * ...
#
# functional-тесты:
# * юнит-тесты
# * интеграционные тесты
# __Вопросы-набросы__:
#
# <details>
# <summary>Зачем нужны юнит-тесты?</summary>
# <p>
#
# * Порверка корректности на этапе написания
# * Предотвращение поломки после рефакторинга
#
# </p>
# </details>
#
# <details>
# <summary>Что тестировать?</summary>
# <p>
#
# * success path
# * failure path
# * граничные значения
# * различные пути выполнения (см. [цикломатическая сложность](https://en.wikipedia.org/wiki/Cyclomatic_complexity))
#
# </p>
# </details>
# Чем тестировать:
#
# Наиболее популярной библиотекой для юнит-тестирования является [google test](https://github.com/google/googletest) и набор вспомогательных утилит для неё [google mock](https://github.com/google/googletest/tree/master/googlemock).
# Правила именования:
#
# ```c++
# TEST(TestSuiteName, TestName)
# ```
# Примеры:
# ```c++
# // sqrt_unittest.cpp
#
# double sqrt(double x);
#
# TEST(sqrt, zero)
# {
# double v = sqrt(0.);
# EXPECT_NEAR(v, 0., 1e-6);
# }
#
# TEST(sqrt, nine)
# {
# EXPECT_NEAR(sqrt(9.), 3., 1e-6);
# }
# ```
# ```c++
# // fabs_unittest.cpp
#
# double fabs(double x);
#
# TEST(fabs, zero)
# {
# EXPECT_EQ(fabs(0.), 0.);
# }
#
# TEST(fabs, positive)
# {
# EXPECT_EQ(fabs(3.), 3.);
# }
#
# TEST(fabs, negative)
# {
# EXPECT_EQ(fabs(-3.), 3.);
# }
# ```
# Библиотека gtest генерирует имена тестов по правилу:
#
# полное имя теста = `TestSuiteName.TestName`
#
# В примере будут сгенерированы тесты:
#
# * `sqrt.zero`
# * `sqrt.nine`
# * `fabs.zero`
# * `fabs.positive`
# * `fabs.negative`
# Макрос `TEST` помимо генерации тела теста ещё регистрирует тело теста и его имя в глобальном контейнере тестов. Затем запускалка перебирает глобальный контейнер и запускает тесты один за другим.
# В файле `sqrt_unittest.o` после компиляции будет сгенерирован код регистрации тестов в глобальном контейнере:
# * `sqrt.zero`
# * `sqrt.nine`
#
# В файле `fabs_unittest.o` после компиляции будет сгенерирован код регистрации тестов в глобальном контейнере:
# * `fabs.zero`
# * `fabs.positive`
# * `fabs.negative`
# Далее линковщик скомпонует `sqrt_unittest.o` и `fabs_unittest.o` вместе в один исполняемый файл, в котором будет код регистрации для каждого теста.
# __Замечание__: Чтобы добавить тесты на функцию `pow` нужно...
#
# <details>
# <summary>...?</summary>
# <p>
#
# создать аналогичный файлик `pow_unittest.cpp` и добавить его в компиляцию и линковку.
#
# </p>
# </details>
#
# <br />
# Запуск:
# `./project_unittest`
# Можно пофильтровать по именам:
# `./project_unittest --gtest_filter=sqrt.*`
# __Вопрос__:
#
# <details>
# <summary>как прогнать тесты sqrt.zero и fabs.zero?</summary>
# <p>
#
# `./project_unittest --gtest_filter=*.zero`
#
# </p>
# </details>
# <br />
# ##### Идеалистическая организация теста
# ```c++
# TEST(function, scenario)
# {
# // настройка окружения
# ...
#
# // выполнение действия, результат которого надо протестировать
# ...
#
# // проверка условий (в идеале - одного-единственного условия)
# ...
# }
# ```
# Пример:
# ```c++
# TEST(read_n, reads_correct_value)
# {
# // prepare
# std::stringstream ss("42");
#
# // run action
# const unsigned n = read_n(ss);
#
# // check assertion
# EXPECT_EQ(n, 42);
# }
# ```
# Для функционала сложнее калькулятора бывает трудно организовать тесты идеальным образом. И это естественно.
# <br />
# ##### EXPECT_* / ASSERT_*
# __Вопрос__:
#
# <details>
# <summary>Чем EXPECT_EQ отличается от ASSERT_EQ?</summary>
# <p>
#
# В случае несовпадения значений
# * `EXPECT_EQ` генерирует нефатальную ошибку и продолжает выполнение теста
# * `ASSERT_EQ` генерирует фатальную ошибку - останавливает выполнение теста
#
# </p>
# </details>
# Для чистоты кода и более адекватных сообщениях об ошибке существуют разные варианты `EXPECT_*` и `ASSERT_*`:
#
# * булевы выражения
# ```c++
# EXPECT_TRUE(v.empty());
# EXPECT_FALSE(v.empty());
# ```
# * равенство (для вещ. чисел см. [статью](https://randomascii.wordpress.com/2012/02/25/comparing-floating-point-numbers-2012-edition/))
# ```c++
# EXPECT_EQ(x, y);
# EXPECT_NE(x, y);
# EXPECT_FLOAT_EQ(x, y);
# EXPECT_DOUBLE_EQ(x, y);
# EXPECT_NEAR(x, y, abs_error); // <-- use this for float/double values
# ```
# * порядок
# ```c++
# EXPECT_LT(x, y);
# EXPECT_LE(x, y);
# EXPECT_GT(x, y);
# EXPECT_GE(x, y);
# ```
# * C-строки
# ```c++
# const char* s = "Get the rack!";
# EXPECT_STREQ(s, "Nobody expects for Spanish Inquisition!");
# EXPECT_STRNE(s, "Nobody expects for Spanish Inquisition!");
# EXPECT_CASEEQ(s, "Nobody expects for Spanish Inquisition!");
# EXPECT_CASENE(s, "Nobody expects for Spanish Inquisition!");
# ```
# * исключения
# ```c++
# EXPECT_THROW(my_function(3., 4.), std::runtime_error);
# EXPECT_ANY_THROW(my_function(3., 4.));
# EXPECT_NO_THROW(my_function(3., 4.));
# EXPECT_NO_THROW({
# const double x = get_number();
# my_function(x);
# });
# ```
# * кастомные условия (можно через `EXPECT_TRUE`, но эти варианты выведут аргументы в значения об ошибке)
# ```c++
# bool isPrime(int n) { ... }
# bool isMutuallyPrime(int m, int n) { ... }
#
# EXPECT_PRED1(isPrime, 5);
# EXPECT_PRED2(isMutuallyPrime, 9, 10);
# ```
# * матчеры из gmock ... (о них в продвинутой части)
# <br />
# ##### ASSERT_NO_FATAL_FAILURE
# `ASSERT_*` - проверки на фатальную ошибку. Делает 3 вещи:
# * печатают сообщение об ошибке
# * добваляют информацию о фатальной ошибке (например, в глобальную (или `thead_local`) переменную)
# * `return` из текущей функции (функция должна быть `void`)
# `ASSERT_NO_FATAL_FAILURE(statement)` - проверка, что `statement` не добавил новых фатальных ошибок, если добавил - `return` из текущей функции (д.б. `void`)
# Зачем?
# ```c++
# void setup_testing_environment()
# {
# const bool network_ok = setup_testing_network();
# ASSERT_TRUE(network_ok);
#
# const bool database_ok = setup_testing_database();
# ASSERT_TRUE(database_ok);
# }
#
# TEST(suite, scenario)
# {
# ASSERT_NO_FATAL_FAILURE(setup_testing_environment());
#
# ...
# }
# ```
# <br />
# ##### Fixtures
# Пример:
# Пишем тесты для функции поиска кратчайшего пути в графе
#
# `Way dijkstra(const Graph& g, const Vertex& v_start, const Vertex& v_final);`
#
# Для серии тестов на `dijkstra` нужно создавать один и тот же граф `G` через вызовы:
#
# ```c++
# auto V = make_special_vertices();
# auto E = make_special_edges(V);
# auto G = make_graph(V, E);
# ```
# Как могли бы выглядеть тесты:
#
# ```c++
# TEST(dijkstra, same_vertex)
# {
# // setup special graph
# const auto V = make_special_vertices();
# const auto E = make_special_edges(V);
# const auto G = make_graph(V, E);
#
# // way v1 -> v1 should have zero len
# const auto v1 = find_vertex(V, "v1");
# const auto way = dijkstra(G, v1, v1);
# EXPECT_EQ(way.length(), 0);
# }
#
# TEST(dijkstra, unreachable_vertex)
# {
# // setup special graph
# const auto V = make_special_vertices();
# const auto E = make_special_edges(V);
# const auto G = make_graph(V, E);
#
# // way between unreachable vertices shouldn't exist
# const auto v1 = find_vertex(V, "v1");
# const auto v9 = find_vertex(V, "v9");
# const auto way = dijkstra(G, v1, v9);
# EXPECT_FALSE(way.is_finite());
# }
#
# TEST(dijkstra, normal_way)
# {
# // setup special graph
# const auto V = make_special_vertices();
# const auto E = make_special_edges(V);
# const auto G = make_graph(V, E);
#
# // normal way should be ok
# const auto v1 = find_vertex(V, "v1");
# const auto v2 = find_vertex(V, "v2");
# const auto way = dijkstra(G, v1, v2);
# EXPECT_TRUE(way.is_finite());
# }
# ```
# __Вопрос__:
#
# <details>
# <summary>В чём проблема? Что должно моментально вызывать аллергическую реакцию программиста?</summary>
# <p>
#
# copy-paste
#
# </p>
# </details>
# Вариант починить проблему - fixtures.
#
# Fixtures позволяют единообразно и разово задать:
# * необходимые действия для инициализации теста
# * необходимые действия для очистки теста
#
# Пример:
# ```c++
# class DijstraSpecialGraph : public Test
# {
# protected:
# void SetUp() override
# {
# V_ = make_special_vertices();
# E_ = make_special_edges(V_);
# G_ = make_graph(V_, E_);
# }
#
# const Vertices& vertices() const noexcept { return V_; }
# const Edges& edges() const noexcept { return E_; }
# const Graph& graph() const noexcept { return G_; }
#
# private:
# Vertices V_;
# Edges E_;
# Graph G_;
# };
#
# TEST_F(DijstraSpecialGraph, same_vertex)
# {
# const auto v1 = find_vertex(vertices(), "v1");
# const auto way = dijkstra(graph(), v1, v1);
# EXPECT_EQ(way.length(), 0);
# }
#
# TEST_F(DijstraSpecialGraph, unreachable_vertex)
# {
# const auto v1 = find_vertex(vertices(), "v1");
# const auto v9 = find_vertex(vertices(), "v9");
# const auto way = dijkstra(graph(), v1, v9);
# EXPECT_FALSE(way.is_finite());
# }
#
# TEST_F(DijstraSpecialGraph, normal_way)
# {
# const auto v1 = find_vertex(vertices(), "v1");
# const auto v2 = find_vertex(vertices(), "v2");
# const auto way = dijkstra(graph(), v1, v2);
# EXPECT_TRUE(way.is_finite());
# }
# ```
# Обратите внимание на:
# * fixture определяется как класс
# * используется макрос `TEST_F`
# * В fixture можно определить методы `void SetUp()` и `void TearDown()`
# * `SetUp` - действия до запуска теста
# * `TearDown` - действия после прогона теста
# __Вопрос__:
#
# <details>
# <summary>Что делать, если нужно 10 тестов на специальный граф и 10 тестов на специальный граф чуть-чуть другого вида, например, с добавлением одного ребра для создания цикла?</summary>
# <p>
#
# Варианты:
# * иерархия классов `DijstraSpecialGraph`: `DijstraSpecialGraphBase` + `DijstraSpecialGraphAcyclic` + `DijstraSpecialGraphCyclic`:
#
# ```c++
# class DijstraSpecialGraphBase : public ::testing::Test
# {
# protected:
# DijstraSpecialGraphBase(bool isCycled) { ... }
# ...
# };
#
# class DijstraSpecialGraphAcyclic : public DijstraSpecialGraphBase
# {
# protected:
# DijstraSpecialGraphAcyclic() : DijstraSpecialGraphBase(false) {}
# };
#
# class DijstraSpecialGraphCyclic : public DijstraSpecialGraphBase
# {
# protected:
# DijstraSpecialGraphCyclic() : DijstraSpecialGraphBase(true) {}
# };
# ```
#
# * шаблонный fixture (почему бы и нет?):
#
# ```c++
# template<bool IsCyclic>
# class DijstraSpecialGraph : public ::testing::Test
# {
# ...
# };
#
# using DijstraSpecialGraphAcyclic = DijstraSpecialGraph<false>;
# using DijstraSpecialGraphCyclic = DijstraSpecialGraph<true>;
# ```
#
# * value-parametrized tests (позже и посложнее)
#
# </p>
# </details>
# __Замечания__:
# * Из-за организации gtest внутри тела теста можно доступаться только до protected данных fixture, т.к. gtest генерирует неявного наследника.
# * В случае иерархии fixture protected поля - сомнительная идея, возможно, лучше защитить их через protected - аксессоры, ещё лучше - const-аксессоры
# <br />
# Как это работает для каждого теста:
# * Сохранить состояния флагов googletest
# * Создать объект fixture
# * Позвать метод `SetUp` у fixture
# * Прогнать тест
# * Позвать `TearDown` у fixture
# * Удалить объект fixture.
# * Восстановить состояния флагов.
#
# ```c++
# save_gtest_flags();
# {
# Fixture f;
# f.SetUp();
# f.RunTest();
# f.TearDown();
# }
# restore_gtest_flags();
# ```
# __Вопрос__: Сколько объектов fixture здесь будет создано?
#
# ```c++
# TEST_F(DijstraSpecialGraph, same_vertex)
# {
# }
#
# TEST_F(DijstraSpecialGraph, unreachable_vertex)
# {
# }
#
# TEST_F(DijstraSpecialGraph, normal_way)
# {
# }
# ```
# <br />
# ##### Отключение тестов
# Если тест падает, но чинить прямо сейчас его некогда (в больших проектах бывает), его можно отключить:
# ```c++
# TEST(DijstraSpecialGraph, DISABLED_unreachable_vertex) { ... }
# ```
# Или отключить всю fixture:
# ```c++
# class DISABLED_DijstraSpecialGraph : public Test { ... };
#
# TEST_F(DISABLED_DijstraSpecialGraph, same_vertex)
# {
# }
#
# TEST_F(DISABLED_DijstraSpecialGraph, unreachable_vertex)
# {
# }
#
# TEST_F(DISABLED_DijstraSpecialGraph, normal_way)
# {
# }
# ```
# __Вопрос__: Вариант решения проблемы - закомментировать тест, но лучше применять DISABLED, почему?
# Как отключить тест для платформы?
# ```c++
# #ifdef __WINDOWS__
# #define MAYBE_normal_way DISABLED_normal_way
# #else
# #define MAYBE_normal_way normal_way
# #endif
#
# TEST_F(DijstraSpecialGraph, MAYBE_normal_way)
# {
# }
# ```
# <br />
# ##### Запуск тестов, доп. опции
# Фильтрация тестов:
#
# * запустить только сюиту `sqrt`:
#
# `./unittests --gtest_filter=sqrt.*`
#
# * запустить `sqrt` && `fabs`:
#
# `./unittests --gtest_filter=*sqrt*:*fabs*`
#
# * запустить `sqrt`, но не `sqrt.zero`:
#
# `./unittests --gtest_filter=sqrt.*-sqrt.zero`
#
# * `sqrt` и `fabs`, но не `zero`:
#
# `./unittests --gtest_filter=sqrt.*:fabs.*-sqrt.zero:fabs.zero`
# Многократный запуск тестов:
#
# `./unittests --gtest_repeat=1000`
#
# __Вопрос__: зачем?
# Остановиться на первом падении:
#
# `./unittests --gtest_break_on_failure`
#
# __Вопрос__: зачем?
# Перемешать порядок тестов:
#
# `./unittest --gtest_shuffle --gtest_random_seed=100500`
#
# __Вопрос__: зачем?
# Изменение формата вывода:
#
# `./unittest --gtest_output=json|xml`
#
# __Вопрос__: зачем?
# <br />
# ##### Что делать, если нужно протестировать состояние приватного поля класса?
# ```c++
# class TriangleShape
# {
# public:
# TriangleShape();
# void draw();
#
# private:
# Color color_; // <-- need to test this value
# Point v1, v2, v3; // <-- need to test this value
# };
# ```
# <details>
# <summary>Ответ</summary>
# <p>
#
# страдать
#
# </p>
# </details>
# Вариант 0: выводить в public всё нужное (совсем плохо)
# Вариант 1: `_for_testing` - методы
# ```c++
# class TriangleShape
# {
# public: // usual interface
# TriangleShape();
#
# void draw();
#
# public: // testing-only methods
# const Color& get_color_for_testing() const { return color_; }
# const Point& get_vertex_1_for_testing() const { return v1; }
# const Point& get_vertex_2_for_testing() const { return v2; }
# const Point& get_vertex_3_for_testing() const { return v3; }
#
# private:
# Color color_;
# Point v1, v2, v3;
# };
# // проще отследить использование "неправильных" методов в боевом коде, но:
# // а) много лишнего шума в классе (читабельность, вреся компиляции)
# // б) лишние методы компилируются в исполняемый файл
# ```
# Вариант 2: `FRIEND_TEST`
# ```c++
# class TriangleShape
# {
# public: // usual interface
# TriangleShape();
#
# void draw();
#
# private:
# // перечислим тесты, которым можно залезать в private:
# FRIEND_TEST(TriangleShape, TestName1);
# FRIEND_TEST(TriangleShape, TestName2);
# FRIEND_TEST(TriangleShape, TestName3);
# FRIEND_TEST(AnotherSuite, AnotherTestName1);
# FRIEND_TEST(AnotherSuite, AnotherTestName2);
#
# Color color_;
# Point v1, v2, v3;
# };
# // в боевом коде лишние методы не компилируется, но:
# // а) много лишнего шума в классе (больше тестов - длиннее текст класса)
# // б) в итоговой программе нужен инклуд из тестовой библиотеки
# ```
# Вариант 3: friend-обёртка над классом для тестов
# ```c++
#
# // production-код:
#
# class TriangleShape
# {
# public: // usual interface
# TriangleShape();
#
# void draw();
#
# private:
# friend class TriangleShapeTestingAccessor;
#
# Color color_;
# Point v1, v2, v3;
# };
#
# // код, компилирующийся только в рамках юнит-тестов:
#
# class TriangleShapeTestingAccessor
# {
# public:
# static const Color& get_color(const TriangleShape& shape) { return shape.color_; }
# };
#
# // сами тесты:
#
# TEST(Suite, Name)
# {
# TriangleShape tri = make_triangle();
# const auto color = TriangleShapeTestingAccessor::get_color(tri);
# }
#
# // в боевом коде почти нет артефактов от тестирования, но:
# // а) приходится писать отдельный TestingAccessor на каждый сложно тестируемый класс
# ```
# <br />
# ##### Матчеры (gmock)
# Матчеры - расширение набора проверок `EXPECT_*`
# https://github.com/google/googletest/blob/master/googlemock/docs/cheat_sheet.md#matchers-matcherlist
# Вызов:
#
# ```c++
# EXPECT_THAT(actual_value, matcher)
# ASSERT_THAT(actual_value, matcher)
# ```
# Пример:
#
# ```c++
# TEST(...)
# {
# std::vector<int> numbers = make_numbers();
# EXPECT_THAT(numbers, IsEmpty());
# }
# ```
# Какие матчеры бывают:
# * аналоги обычных `EXPECT_*/ASSERT_*`
# * `EXPECT_THAT(value, Eq(42));`
# * `EXPECT_THAT(value, Ne(42));`
# * `EXPECT_THAT(value, Ge(42));`
# * `EXPECT_THAT(value, Gt(42));`
# * `EXPECT_THAT(value, Le(42));`
# * `EXPECT_THAT(value, Lt(42));`
# * `EXPECT_THAT(value, IsTrue());`
# * `EXPECT_THAT(value, IsFalse());`
# * `EXPECT_THAT(value, DoubleEq(42.0));`
# * `EXPECT_THAT(value, FloatEq(42.0));`
# * `EXPECT_THAT(value, DoubleNear(42.0, 1e-3));`
# * `EXPECT_THAT(value, FloatNear(42.0, 1e-3));`
# * указатели
# * `EXPECT_THAT(pointer, IsNull());`
# * `EXPECT_THAT(pointer, NotNull());`
# * строки
# * `EXPECT_THAT(s, EndsWith("abc"));`
# * `EXPECT_THAT(s, StartsWith("abc"));`
# * `EXPECT_THAT(s, HasSubstr("abc"));`
# * `EXPECT_THAT(s, StrEq("abc"));`
# * `EXPECT_THAT(s, StrNe("abc"));`
# * `EXPECT_THAT(s, StrCaseEq("abc"));`
# * `EXPECT_THAT(s, StrCaseNe("abc"));`
# * `EXPECT_THAT(s, ContainsRegex("abc*"));`
# * контейнеры
# * `EXPECT_THAT(numbers, IsEmpty());`
# * `EXPECT_THAT(numbers, SizeIs(5));`
# * `EXPECT_THAT(numbers, ContainerEq(another_numbers));`
# * `EXPECT_THAT(numbers, Contains(42));`
# * `EXPECT_THAT(strings, Each(IsEmpty()));`
# * `EXPECT_THAT(numbers, ElementsAre(1, 2, 3, 4, 5));`
# * `EXPECT_THAT(numbers, ElementsAreArray(another_numbers));`
# * `EXPECT_THAT(numbers, UnorderedElementsAre(1, 2, 3, 4, 5));`
# * `EXPECT_THAT(numbers, UnorderedElementsAreArray(another_numbers));`
# * `EXPECT_THAT(numbers, IsSubsetOf({1, 2, 3, 4, 5}));`
# * `EXPECT_THAT(numbers, WhenSorted(ElementsAre(1, 2, 3)));`
# * композиции
# * `AllOf`:
# ```c++
# EXPECT_THAT(string,
# AllOf(EndsWith(".txt"),
# SizeIs(10)));
# ```
# * `AnyOf`:
# ```c++
# EXPECT_THAT(string,
# AnyOf(EndsWith(".txt"),
# SizeIs(10)));
#
# ```
# * `Not`:
# ```c++
# EXPECT_THAT(numbers, Not(IsSubsetOf({1, 2, 3, 4, 5})));
# ```
# * создание собственных матчеров:
#
# ```c++
# MATCHER(IsEven, "") { return (arg % 2) == 0; }
#
# EXPECT_THAT(value, IsEven());
# ```
# __Упражение__:
#
# <details>
# <summary>С помощью матчеров проверить, что в списке имён файлов нет текстовых (.txt)</summary>
# <p>
#
# `EXPECT_THAT(filenames, Each(Not(EnsWith(".txt"))));`
#
# </p>
# </details>
# <br />
# ##### Dependency inversion
# Инверсия зависимостей - техника, позволяющая "развязать" зависимости между компонентами программы через интерфейсы.
# https://en.wikipedia.org/wiki/Dependency_injection
#
# https://en.wikipedia.org/wiki/Dependency_inversion_principle
# Рассмотрим на примере 2 реализации поиска самого молодого студента в БД:
# Вариант 1:
#
# ```c++
# std::optional<Student> find_youngest_student(const Database& db)
# {
# std::optional<Student> rv;
#
# const auto students = db.get_students();
# if (!students.empty())
# {
# rv.emplace(*min_element(begin(students), end(students),
# [](const Student& lhs, const Student& rhs){
# return lhs.age < rhs.age;
# }));
# }
#
# return rv;
# }
# ```
# Вариант 2:
#
# ```c++
#
# // IDatabase.h
# // где-то в районе определения кода БД:
# struct IDatabase
# {
# virtual ~IDatabase() = default;
#
# virtual std::vector<Student> get_students() const = 0;
# };
#
# // Database.h
# // где-то в районе определения кода БД:
# class Database : public IDatabase
# {
# public:
# std::vector<Student> get_students() const override;
# ...
# };
#
# // функция поиска самого молодого студента
# // Обратите внимание на тип принимаемого аргумента
#
# std::optional<Student> find_youngest_student(const IDatabase& db)
# {
# std::optional<Student> rv;
#
# const auto students = db.get_students();
# if (!students.empty())
# {
# auto it = min_element(begin(students), end(students),
# [](const Student& lhs, const Student& rhs){
# return lhs.age < rhs.age;
# });
# rv.emplace(*it);
# }
#
# return rv;
# }
# ```
# __Особенности__:
# * Сложность реализации:
#
# Вариант с интерфейсом посложнее (нужно писать интерфейсы и наследование)
#
#
# * Время выполнения:
#
# Вариант с интерфейсом содежрит доп. расходы на вызов виртуальной функции
#
#
# * Время компиляции:
#
# В среднем по больнице, вариант с интерфейсом компилируется быстрее
#
# <details>
# <summary>Почему? что увеличивает, а что уменьшает время компиляции?</summary>
# <p>
#
# увеличивает время компиляции:
# * парсинг самого интерфейса при компиляции кода БД
# * парсинг с наследованием и накладные расходы на него
# * генерация доп. кода прыжков по вирутальным функциям
#
# уменьшает время компиляции (т.к. внешний код к БД инклудит только IDatabase.h):
# * не нужно парсить и компилировать "..." в классе `Database`
# * из-за этого `IDatabase.h` (как правило) тянет много меньше зависимостей по инклудам
# * не нужно перекомпилировать при изменении `Database.h`
#
# </p>
# </details>
#
# * Тестируемость:
#
# Вариант 1 тестировать нетривиально, вариант 2 - очень легко:
# ```c++
# class EmptyTestingDatabase : public IDatabase
# {
# public:
# std::vector<Student> get_students() const override
# {
# return {};
# }
# };
#
# TEST(find_youngest_student, no_students)
# {
# EmptyTestingDatabase db;
#
# const auto maybe_student = find_youngest_student(db);
#
# EXPECT_FALSE(maybe_student.has_value());
# }
#
# class SingleEinsteinDatabase : public IDatabase
# {
# public:
# std::vector<Students> get_students() const override
# {
# return {Student{"Einstein", 38}};
# }
# };
#
# TEST(find_youngest_student, those_boy_einstein)
# {
# SingleEinsteinDatabase db;
#
# const auto maybe_student = find_youngest_student(db);
# ASSERT_TRUE(maybe_student.has_value());
#
# EXPECT_EQ(maybe_student.value().name == "Einstein");
# }
# ```
# <br />
# ##### Mocks
# Мокирование - механизм, который помогает на лету генерировать вспомогательные тестовые объекты типа `EmptyTestingDatabase`, `SingleEinsteinDatabase`.
# Пример:
# ```c++
# class MockDatabase : public IDatabase
# {
# public:
# MOCK_METHOD(std::vector<Student>, get_students, (), (const, override));
# };
#
#
# TEST(find_youngest_student, no_students)
# {
# MockDatabase db;
# ON_CALL(db, get_students())
# .WillByDefault(Return(std::vector<Student>()));
#
# const auto maybe_student = find_youngest_student(db);
#
# EXPECT_FALSE(maybe_student.has_value());
# }
#
# TEST(find_youngest_student, those_boy_einstein)
# {
# MockDatabase db;
# ON_CALL(db, get_students())
# .WillByDefault(Return(std::vector<Student>{Student{"Einstein", 38}}));
#
# const auto maybe_student = find_youngest_student(db);
# ASSERT_TRUE(maybe_student.has_value());
#
# EXPECT_EQ(maybe_student.value().name == "Einstein");
# }
# ```
# <br />
# 2 варианта работы с mock-объектами:
# * `ON_CALL` - указать, что будет происходить при вызове метода
# * `EXPECT_CALL` - указать ожидания к вызову (expectations) и что
# Примеры с `EXPECT_CALL` от gtest-a:
#
# ```c++
# EXPECT_CALL(turtle, GetX())
# .Times(5)
# .WillOnce(Return(100))
# .WillOnce(Return(150))
# .WillRepeatedly(Return(200));
# // TODO: что произойдёт, если в тесте GetX позвался 4 раза?
#
# // что здесь?
# EXPECT_CALL(turtle, GoTo(50, _))
# .Times(AtLeast(3));
#
# // что здесь?
# EXPECT_CALL(turtle, Forward(Ge(100)))
# .Times(Between(2, 5));
#
# // что ожидается здесь?
# EXPECT_CALL(turtle, GetY())
# .Times(4)
# .WillOnce(Return(100));
# ```
#
# <details>
# <summary>Ответ</summary>
# 100, 0, 0, 0
# </details>
# Mock-объектам можно задать действия, что делать при срабатывании вызова:
# ```c++
# TEST(find_youngest_student, those_boy)
# {
# MockDatabase db;
# ON_CALL(db, get_students())
# .WillOnce(Invoke([]() -> std::vector<Student> {
# std::cout << "get_students() called" << std::endl;
# return {Student{"Einstein", 38}};
# }));
# ...
# }
# ```
# Ещё больше способов задать хитрых действий:
# https://github.com/google/googletest/blob/master/googlemock/docs/cheat_sheet.md#actions-actionlist
# Гайд от gtest-а по работе с mock-объектами:
# 1. создать mock-объект
# 2. если требуется, задать дефолтные actions
# 3. задать expectations
# 4. выполнить код теста
# 5. при уничтожении mock-объекта автоматически проверяется, что все его expectations выполнены
# <br />
# ##### Тестирование последовательности вызовов методов mock-объекта
# Mock-объектам можно указывать, в каком порядке должны вызываться методы.
#
# Для этого используется объект класса `InSequence`.
#
# Пример от gtest-а:
#
# ```c++
# TEST(FooTest, MoveTurtle)
# {
# MockTurtle turtle;
#
# InSequence seq;
# EXPECT_CALL(turtle, PenDown());
# EXPECT_CALL(turtle, Forward(100));
# EXPECT_CALL(turtle, PenUp());
#
# RunScenario(turtle);
# }
# ```
# Пример на псевдокоде, где такое требование может быть логичным:
#
# ```c++
# TEST(LocationService, LocationIsRequested)
# {
# MockLocationService location_service;
#
# InSequence seq;
# EXPECT_CALL(location_service, Initialize());
# EXPECT_CALL(location_service, RequestLocation())
# .WillByDefault(
# Return(
# Location(55, 83)));
# EXPECT_CALL(location_service, Shutdown());
#
# const auto app = LaunchApp();
# DisplayCurrentLocation(app);
# CheckLocationDisplayed(app);
# }
# ```
# Более сложный пример с графом зависимостей:
# ```c++
# Expectation init_x = EXPECT_CALL(obj, InitX());
# Expectation init_y = EXPECT_CALL(obj, InitY());
# EXPECT_CALL(obj, some_method())
# .After(init_x, init_y);
# ```
# Больше примеров в документации:
#
# https://github.com/google/googletest/blob/master/googlemock/docs/cheat_sheet.md#expectation-order
# <br />
# ##### Тесты, параметризированные значениями
# Пример, найденный на просторах сети, на мой субъективный взгляд, исключительно демонстрационный и сверх меры усложняющий альтернативное "тупое" решение:
# Вариант 1:
# ```c++
# class LeapYearTest : public TestWithParam<int> {};
#
# TEST_P(LeapYearTest, LeapYear) {
# const int year = GetParam(); // <-- notice GetParam() call
# EXPECT_TRUE(isLeapYear(year));
# }
#
# INSTANTIATE_TEST_CASE_P(
# LeapYearTestsP,
# LeapYearTest,
# Values(104, 1996, 1960, 2012));
#
#
# class NotLeapYearTest : public TestWithParam<int> {};
#
# TEST_P(NotLeapYearTest, NotLeapYear) {
# const int year = GetParam(); // <-- notice GetParam() call
# EXPECT_FALSE(isLeapYear(year));
# }
#
# INSTANTIATE_TEST_CASE_P(
# NotLeapYearTestsP,
# NotLeapYearTest,
# Values(103, 1995, 1961, 1900));
# ```
# Вариант 2:
# ```c++
# struct LeapYearTestParam
# {
# int year;
# bool leap;
# };
#
# class LeapYearTest : public TestWithParam<LeapYearTestParam> {};
#
# TEST_P(LeapYearTest, ChecksIfLeapYear) {
# EXPECT_EQ(GetParam().leap, isLeapYear(GetParam().year));
# }
#
# INSTANTIATE_TEST_CASE_P(
# LeapYearTests,
# LeapYearTest,
# Values(LeapYearTestParam{7, false},
# LeapYearTestParam{2001, false},
# LeapYearTestParam{1996, true},
# LeapYearTestParam{1700, false},
# LeapYearTestParam{1600, true}));
# ```
# __Замечание__: ещё есть тесты, параметризованные типами (на шаблонах)
# https://github.com/google/googletest/blob/master/googletest/samples/sample6_unittest.cc
# <br />
# ##### шаринг данных в fixture, глобальный шаринг данных
# __Вопрос:__ сколько будет создано объектов класса `DijstraSpecialGraph` при прогоне таких тестов?
#
# ```c++
# class DijstraSpecialGraph : public Test
# {
# ...
# };
#
# TEST_F(DijstraSpecialGraph, same_vertex) { ... }
# TEST_F(DijstraSpecialGraph, unreachable_vertex) { ... }
# TEST_F(DijstraSpecialGraph, normal_way) { ... }
# ```
# Это принцип дизайна gtest-а, который (наиболее вероятно) сэкономил море человекочасов, избежав ненужных зависимостей между соседними тестами.
# В случае, если заполнение данных для fixture слишком дорого чтобы делать это на каждый тест, можно шарить данные между fixture:
# ```c++
# class DijstraSpecialGraphTest : public Test
# {
# protected:
# static void SetUpTestSuite()
# {
# shared_graph_.emplace(...); // setup shared resource
# }
#
# static void TearDownTestSuite()
# {
# shared_graph_.reset(); // cleanup shared resource
# }
#
# void SetUp() override { ... }
# void TearDown() override { ... }
#
# static std::optional<T> shared_graph_;
# };
# ```
# <br />
# Ещё можно делать глобальный setup-teardown:
#
# https://github.com/google/googletest/blob/master/googletest/docs/advanced.md#global-set-up-and-tear-down
#
# Но при более-менее прямой организации тестов не могу представить зачем это может понадобиться.
# <br />
# ##### Ускорение компиляции mock-объектов
# https://github.com/google/googletest/blob/master/googlemock/docs/cook_book.md#making-the-compilation-faster
# Mock-объекты компилируются не быстро.
#
# По утвереждению авторов gtest-а основное время компиляции уходит на автогенерированные конструкторы и деструкторы из-за того, что в них нужно генерировать код инициализации и проверки expectations.
#
# Решение для переиспользуемых mock-объектов - компилировать конструкторы и деструкторы отдельно в cpp-файле.
# Было:
# ```c++
# // MockNetworkService.h
#
# class MockNetworkService : public INetworkService
# {
# public:
# MOCK_METHOD(void, initialize, (), (override));
# MOCK_METHOD(std::string, request, (std::string_view), (override));
# MOCK_METHOD(void, shutdown, (), (override));
# };
# ```
# Стало:
# ```c++
# // MockNetworkService.h
#
# class MockNetworkService : public INetworkService
# {
# public:
# MockNetworkService();
# ~MockNetworkService();
#
# MOCK_METHOD(void, initialize, (), (override));
# MOCK_METHOD(std::string, request, (std::string_view), (override));
# MOCK_METHOD(void, shutdown, (), (override));
# };
#
# // MockNetworkService.cpp
#
# MockNetworkService::MockNetworkService() = default;
# MockNetworkService::~MockNetworkService() = default;
# ```
# __Вопрос:__ сколько раз будет скомпилирован конструктор и деструктор во втором варианте? Почему? А в первом?
# <br />
# Полезный доп.материал для самостоятельного изучения:
# * default actions на вызов мокированного метода
# * nice/strict mocks
# * зачем нужен RetiresOnSaturation
# * тесты, параметризированные типами
# * Death tests
# * gtest listener API
| 2021/sem2/lecture_unit_testing/lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df = pd.read_csv('eda_data.csv')
df['job_simp'].value_counts()
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
df['job_simp']= le.fit_transform(df['job_simp'])
df['job_simp'].value_counts()
df['seniority']= le.fit_transform(df['seniority'])
df['seniority'].value_counts()
df['job_state'].value_counts()
df['job_state']= le.fit_transform(df['job_state'])
df['job_state'].value_counts()
df['Revenue'].value_counts()
df['Revenue']= le.fit_transform(df['Revenue'])
df['Revenue'].value_counts()
df['Sector'].value_counts()
df['Sector']= le.fit_transform(df['Sector'])
df['Sector'].value_counts()
df['Industry'].value_counts()
df['Industry']= le.fit_transform(df['Industry'])
df['Industry'].value_counts()
df['Size'].value_counts()
df['Size']= le.fit_transform(df['Size'])
df['Size'].value_counts()
df['Type of ownership'].value_counts()
df['Type of ownership']= le.fit_transform(df['Type of ownership'])
df['Type of ownership'].value_counts()
df.to_csv('final_data.csv')
df.head()
df_model = df[['avg_salary','Rating','Size','Type of ownership','Industry','Revenue','num_comp','hourly','employer_provided','job_state','same_state','age','python_yn','spark','aws','excel','job_simp','seniority','desc_len']]
df_dum = pd.get_dummies(df_model)
corr_matrix = df.corr()
corr_matrix['avg_salary'].sort_values(ascending=False)
df_final = df_dum[['python_yn','spark','aws','job_state_CA','job_simp_na','Type of ownership_Nonprofit Organization','seniority_na','hourly','job_simp_analyst']]
df_final.head()
| Final Model/.ipynb_checkpoints/label_encoding-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false, "name": "#%%\n"}
import cv2 as cv
import numpy as np
from scipy.io import loadmat
from matplotlib import pyplot as plt
from os.path import basename,join,splitext
from SpectralUtils import saveJPG, demosaic, addNoise, projectToRGBMosaic
# Example noise parameters
# # ! IMPORTANT !
# Actual challenge images will be generated with different parameters
DARK_NOISE = 10
TARGET_NPE = 5000
filePath= "./resources/sample_hs_img_001.mat"
# # ! IMPORTANT !
# Actual challenge images will be generated with a confidential camera response function, the filter
# used below is included as an example
filtersPath = "./resources/example_D40_camera_w_gain.npz"
savePath = "./output/"
# Loading HS image and filters
cube = loadmat(filePath)['cube']
filters = np.load(filtersPath)['filters']
# Apply "Real World" simulated camera pipeline:
# 1. Project to RGGB mosaic image (simulate sensor color filter array)
# 2. Add noise
# 3. Demosaic
im = projectToRGBMosaic(cube,filters)
im = addNoise(im, DARK_NOISE, TARGET_NPE)
im = demosaic(im)
# Save image file
fileName = splitext(basename(filePath))[0]
path = join(savePath, fileName +'_realWorld.jpg')
saveJPG(im,path)
# Display RGB image
img = cv.imread(path)
img = cv.cvtColor(img, cv.COLOR_BGR2RGB)
plt.imshow(img)
plt.title('Example "Real World" Output Image')
plt.show()
| real_world_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''py_37_pytorch'': conda)'
# name: python3
# ---
# ```py
# scales = torch.logspace(0., log(max_freq / 2) / log(base), num_bands, base = base, device = device, dtype = dtype)
# scales = scales[(*((None,) * (len(x.shape) - 1)), Ellipsis)]
#
# x = x * scales * pi
# x = torch.cat([x.sin(), x.cos()], dim=-1)
# x = torch.cat((x, orig_x), dim = -1)
# ```
# $$
# scales[i] = base^{\frac{\log(freq/2) \cdot i}{\log(base) \cdot (band -1 )}} = 10 ^ {\frac{\log(freq/2)}{band - 1}} = (\frac{freq}{2}) ^ {\frac{i}{band - 1}}
# $$
list(map(lambda x: 5**(0.2*x), [0, 1, 2, 3, 4, 5])) # 验证数值一致
# (X, Y) 有244*244个点,X $\in$ \[-1, 1\], Y $\in$ \[-1, 1\] 均匀分布。以X为例,X的encoding为13位:
# $$
# \sin(\pi (freq/2)^0 x), \sin(\pi (freq/2)^{0.2} x),\sin(\pi (freq/2)^{0.4} x),\sin(\pi (freq/2)^{0.6} x),\sin(\pi (freq/2)^{0.8} x), \sin(\pi (freq/2)^{1.0} x),\\
# \cos(\pi (freq/2)^0 x), \cos(\pi (freq/2)^{0.2} x),\cos(\pi (freq/2)^{0.4} x),\cos(\pi (freq/2)^{0.6} x),\cos(\pi (freq/2)^{0.8} x), \cos(\pi (freq/2)^{1.0} x), \\
# x\\
# $$
# Paper 中指出的是这个格式
# $\left[\sin \left(f_{k} \pi x_{d}\right), \cos \left(f_{k} \pi x_{d}\right)\right]$
#
# 这里面的$f_k = (freq/2)^{k/(band-1)}$
#
# 这里的设计很巧妙,在k=0时,是频率最低的时候,这个band下(即encoding的第0位和第6位),$f_k$=1, 周期恰好为2,所有的X在这个band下没有重叠,相当于一个绝对编码,可以绝对地区分每一个X。
#
# 当k越大时,频率约高,可以表征不同粒度(尺度)下的相对位置关系。
#
# 单独的一个X也可以表征绝对位置。可视化如下图:
#
# <img src="figures/Vis_X_Fourier_Encoding.png" width = "300" height = "200" alt="图片名称" align=center />
#
# 这就相当于给了我们一种可行的$f_k$的设计,那就是从恰好一个周期的绝对表征开始,逐渐加大频率,最高频率根据粒度而定。
#
# 在latent transformer部分:
#
# 采用的是256(N) * 512(d)的设置,positional encoding 沿用了和transformer相同的PE:
#
# $$
# \sin\big(\frac{pos}{10000^{\frac{2i}{d}}}\big), \cos\big(\frac{pos}{10000^{\frac{2i}{d}}}\big), i \in [0, d/2)
# $$
#
# 整体上也可以纳入fourier encoding的范畴,$f_k = \frac{1}{\pi\times 10000^{\frac{2k}{d}}}$
#
# 为什么选择10000?
#
# 根据上面的分析,至少要保证encoding的一个band能够表示绝对编码,也就是要求最长的周期要大于256(N), 即$2\pi\times base > 256$, base > 41, 选择base为10000, 则可以保证最长31400的序列都可以有绝对的编码,这种意义上base越大越好(反正最小周期都是2$\pi$)
#
# <img src="figures/Vis_Transformer_PE.png" width = "300" height = "200" alt="图片名称" align=center />
#
| debug.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# WebScrapping data using Beautiful Soup, scrap data from html file we downloaded.
# we will extract table from that and combine data from aqi data and this data
from ipynb.fs.full.Plot_AQI import avg_data_2013,avg_data_2014,avg_data_2015,avg_data_2016
lst=avg_data_2013()
len(lst)
import sys
import requests
import pandas as pd
from bs4 import BeautifulSoup
import os
import csv
def met_data(month,year):
file_html=open('Data/Html_Data/{}/{}.html'.format(year,month),'rb') #rb is read byte mode
plain_text=file_html.read()
#when using beautiful soup , first we have to open html file and read it
tempD=[]
finalD=[]
#initializing beautifulsoup with plain_text anf 'lxml'
soup=BeautifulSoup(plain_text,'lxml')
#we will go to html page and inspect element, seeing the table clause and taking its class
#here class is- 'medias mensuales numspan'
for table in soup.findAll('table',{'class':'medias mensuales numspan'}):
#inside html page we have tbody and tr which have table headings and table rows
for tbody in table:
for tr in tbody: #inside tr we have th(which have specific value) and then we have to pick up text
a=tr.get_text()
tempD.append(a)
#a will have whole row in it and it have 15 features in it
rows=len(tempD)/15
for times in range(round(rows)):
newtempD=[]
#inside each row we have 15 features , so we iterate through each feature now
for i in range(15):
newtempD.append(tempD[0])
tempD.pop(0)
finalD.append(newtempD)
#we will drop features which have no values in it
length=len(finalD)
finalD.pop(length-1) #we not want monthly features in it
finalD.pop(0) #zeroth record is our feature
for a in range(len(finalD)):
finalD[a].pop(14)
finalD[a].pop(13)
finalD[a].pop(12)
finalD[a].pop(11)
finalD[a].pop(10)
finalD[a].pop(6)
finalD[a].pop(0)
return finalD
# +
def data_combine(year, cs):
for a in pd.read_csv('Data/Real-Data/real_' + str(year) + '.csv', chunksize=cs):
df = pd.DataFrame(data=a)
mylist = df.values.tolist()
return mylist
# +
if __name__=='__main__':
if not os.path.exists('Data/real-Data'):
os.makedirs('Data/real-Data')
for year in range(2013,2017):
final_data=[]
with open('Data/real-Data/real_'+str(year)+'.csv','w') as csvfile:
wr=csv.writer(csvfile,dialect='excel')
wr.writerow(
['T', 'TM', 'Tm', 'SLP', 'H', 'VV', 'V', 'VM', 'PM 2.5'])
for month in range(1, 13):
temp = met_data(month, year)
final_data = final_data + temp
#getting dependent features from aqi
pm = getattr(sys.modules[__name__], 'avg_data_{}'.format(year))()
if len(pm) == 364:
pm.insert(364, '-')
for i in range(len(final_data)-1):
# final[i].insert(0, i + 1)
final_data[i].insert(8, pm[i])
with open('Data/Real-Data/real_' + str(year) + '.csv', 'a') as csvfile:
wr = csv.writer(csvfile, dialect='excel')
for row in final_data:
flag = 0
for elem in row:
if elem == "" or elem == "-":
flag = 1
if flag != 1:
wr.writerow(row)
data_2013 = data_combine(2013, 600)
data_2014 = data_combine(2014, 600)
data_2015 = data_combine(2015, 600)
data_2016 = data_combine(2016, 600)
total=data_2013+data_2014+data_2015+data_2016
with open('Data/Real-Data/Real_Combine.csv', 'w') as csvfile:
wr = csv.writer(csvfile, dialect='excel')
wr.writerow(
['T', 'TM', 'Tm', 'SLP', 'H', 'VV', 'V', 'VM', 'PM 2.5'])
wr.writerows(total)
df=pd.read_csv('Data/Real-Data/Real_Combine.csv')
# -
df=pd.read_csv('Data/real-Data/Real_Combine.csv')
df.head(25)
df.shape
| Extract_Combine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ML-Core]
# language: python
# name: conda-env-ML-Core-py
# ---
# +
# default_exp annotation.core
# -
# hide
from nbdev.showdoc import *
# export
import argparse
import logging
import shutil
from abc import ABC, abstractmethod
from enum import Enum
from copy import deepcopy
from os.path import join, isdir, basename
from aiforce import category_tools
from aiforce.core import assign_arg_prefix
from aiforce.io.core import strip_path, scan_files, create_folder
# hide
# %reload_ext autoreload
# %autoreload 2
# export
logger = logging.getLogger(__name__)
# # Annotation
# > Data annotation logic.
# export
class SubsetType(Enum):
"""
The annotation subset type
"""
TRAINVAL = 'trainval'
TRAIN = 'train'
VAL = 'val'
TEST = 'test'
def __str__(self):
return self.value
# export
class RegionShape(Enum):
"""
The supported region shape types
"""
NONE = 'none'
CIRCLE = 'circle'
ELLIPSE = 'ellipse'
POINT = 'point'
POLYGON = 'polygon'
POLYLINE = 'polyline'
RECTANGLE = 'rect'
def __str__(self):
return self.value
# export
def parse_region_shape(shape_str):
"""
Try to parse the region shape from a string representation.
`shape_str`: the shape as string
return: the parsed RegionShape
raises: `ValueError` if unsupported shape parsed
"""
try:
return RegionShape(shape_str)
except ValueError:
raise ValueError("Error, unsupported region shape: {}".format(shape_str))
# export
class Region:
"""
A region
`shape`: the region shape
`points_x`: a list of points x-coordinates
`points_y`: a list of points y-coordinates
`radius_x`: a radius on x-coordinate
`radius_y`: a radius on y-coordinate
`rotation`: a rotation factor
`labels`: a set of region labels
"""
def __init__(self, shape=RegionShape.NONE, points_x=None, points_y=None, radius_x=0, radius_y=0, rotation=0,
labels=None):
self.shape = shape
self.points_x = [] if points_x is None else points_x
self.points_y = [] if points_y is None else points_y
self.radius_x = radius_x
self.radius_y = radius_y
self.rotation = rotation
self.labels = [] if labels is None else labels
# export
class Annotation:
"""
A annotation for a file.
`annotation_id`: a unique annotation identifier
`file_name`: the file
`file_path`: the file path
`regions`: A list of regions
"""
def __init__(self, annotation_id=None, file_path=None, regions=None):
self.annotation_id = annotation_id
self.file_path = file_path
self.regions: [Region] = [] if regions is None else regions
def labels(self):
"""
Returns a list of labels, assigned to the annotation.
return: a list of labels
"""
labels = {}
for region in self.regions:
for label in region.labels:
labels[label] = None
return list(labels.keys())
show_doc(Annotation.labels)
# export
def annotation_filter(annotations, condition):
"""
Filter annotations.
`annotations`: the annotations to filter
`condition`: the filter callback
return: the filtered annotations
"""
filtered = dict()
for (key, value) in annotations.items():
if condition(key, value):
filtered[key] = value
return filtered
# export
def convert_region(region: Region, target_shape: RegionShape):
"""
Convert region to target shape.
`region`: the region to convert
`target_shape`: the target shape to convert to
"""
if target_shape != region.shape:
x_min = min(region.points_x) - region.radius_x if len(region.points_x) else 0
x_max = max(region.points_x) + region.radius_x if len(region.points_x) else 0
y_min = min(region.points_y) - region.radius_y if len(region.points_y) else 0
y_max = max(region.points_y) + region.radius_y if len(region.points_y) else 0
center_x = x_min + x_max - x_min
center_y = y_min + y_max - y_min
region.shape = target_shape
if target_shape == RegionShape.NONE:
region.points_x = []
region.points_y = []
region.radius_x = 0
region.radius_y = 0
region.rotation = 0
elif target_shape == RegionShape.CIRCLE or target_shape == RegionShape.ELLIPSE:
region.points_x = [center_x]
region.points_y = [center_y]
region.radius_x = x_max - center_x
region.radius_y = y_max - center_y
region.rotation = 0
elif target_shape == RegionShape.POINT:
region.points_x = [center_x]
region.points_y = [center_y]
region.radius_x = 0
region.radius_y = 0
region.rotation = 0
elif target_shape == RegionShape.POLYGON or target_shape == RegionShape.POLYLINE:
region.points_x = [x_min, x_min, x_max, x_max, x_min]
region.points_y = [y_min, y_max, y_max, y_min, y_min]
region.radius_x = 0
region.radius_y = 0
region.rotation = 0
elif target_shape == RegionShape.RECTANGLE:
region.points_x = [x_min, x_max]
region.points_y = [y_min, y_max]
region.radius_x = 0
region.radius_y = 0
region.rotation = 0
else:
raise NotImplementedError('unsupported conversion {} -> {}'.format(region.shape, target_shape))
# export
def region_bounding_box(region: Region):
"""
Calculates the region bounding box.
`region`: the region
return: a tuple of points_x and points_y
"""
bbox = deepcopy(region)
convert_region(bbox, RegionShape.RECTANGLE)
return bbox.points_x, bbox.points_y
# export
class AnnotationAdapter(ABC):
"""
Abstract Base Adapter to inherit for writing custom adapters
"""
DEFAULT_CATEGORIES_FILE = 'categories.txt'
def __init__(self, path, categories_file_name=None):
"""
Base Adapter to read and write annotations.
`path`: the folder containing the annotations
`categories_file_name`: the name of the categories file
"""
self.path = strip_path(path)
if categories_file_name is None:
self.categories_file_name = self.DEFAULT_CATEGORIES_FILE
else:
self.categories_file_name = categories_file_name
def list_files(self, subset_type=SubsetType.TRAINVAL):
"""
List all physical files in a sub-set.
`subset_type`: the subset type to list
return: a list of file paths if subset type exist, else an empty list
"""
path = join(self.path, str(subset_type))
return scan_files(path) if isdir(path) else []
def write_files(self, file_paths, subset_type=SubsetType.TRAINVAL):
"""
Write physical files in a sub-set.
`file_paths`: a list of file paths to write
`subset_type`: the subset type to write into
return: a list of written target file paths
"""
path = join(self.path, str(subset_type))
create_folder(path)
copied_files = []
for file_path in file_paths:
file_name = basename(file_path)
target_file = join(path, file_name)
shutil.copy2(file_path, target_file)
copied_files.append(target_file)
return copied_files
@abstractmethod
def read_annotations(self, subset_type=SubsetType.TRAINVAL):
"""
Read annotations.
`subset_type`: the subset type to read
return: the annotations as dictionary
"""
pass
def read_categories(self):
"""
Read categories.
return: a list of category names
"""
path = join(self.path, self.categories_file_name)
logger.info('Read categories from {}'.format(path))
return category_tools.read_categories(path)
@abstractmethod
def write_annotations(self, annotations, subset_type=SubsetType.TRAINVAL):
"""
Write annotations.
`annotations`: the annotations as dictionary
`subset_type`: the subset type to write
return: a list of written target file paths
"""
pass
def write_categories(self, categories):
"""
Write categories.
`categories`: a list of category names
"""
target_folder = create_folder(self.path)
path = join(target_folder, self.categories_file_name)
logger.info('Write categories to {}'.format(path))
category_tools.write_categories(categories, path)
@classmethod
def argparse(cls, prefix=None):
"""
Returns the argument parser containing argument definition for command line use.
`prefix`: a parameter prefix to set, if needed
return: the argument parser
"""
parser = argparse.ArgumentParser()
parser.add_argument(assign_arg_prefix('--path', prefix),
dest="path",
help="the folder containing the annotations.",
required=True)
parser.add_argument(assign_arg_prefix('--categories_file_name', prefix),
dest="categories_file_name",
help="The name of the categories file.",
default=None)
return parser
show_doc(AnnotationAdapter.list_files)
show_doc(AnnotationAdapter.read_annotations)
show_doc(AnnotationAdapter.read_categories)
show_doc(AnnotationAdapter.write_files)
show_doc(AnnotationAdapter.write_annotations)
show_doc(AnnotationAdapter.write_categories)
show_doc(AnnotationAdapter.argparse)
# +
# hide
# for generating scripts from notebook directly
from nbdev.export import notebook2script
notebook2script()
| annotation-core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Matplotlib Exercise 3
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# + [markdown] nbgrader={}
# ## Contour plots of 2d wavefunctions
# + [markdown] nbgrader={}
# The wavefunction of a 2d quantum well is:
#
# $$ \psi_{n_x,n_y}(x,y) = \frac{2}{L}
# \sin{\left( \frac{n_x \pi x}{L} \right)}
# \sin{\left( \frac{n_y \pi y}{L} \right)} $$
#
# This is a scalar field and $n_x$ and $n_y$ are quantum numbers that measure the level of excitation in the x and y directions. $L$ is the size of the well.
#
# Define a function `well2d` that computes this wavefunction for values of `x` and `y` that are NumPy arrays.
# + nbgrader={"checksum": "702bb86154c0c4ec6fbd9e63c7f50289", "solution": true}
def well2d(x, y, nx, ny, L=1.0):
"""Compute the 2d quantum well wave function."""
# YOUR CODE HERE
psi_x_y = (2/L)*np.sin((nx*np.pi*x)/L)*np.sin((ny*np.pi*y)/L)
return psi_x_y
# + deletable=false nbgrader={"checksum": "ee9cdf5b84f1f0d4d545448b2196c9b2", "grade": true, "grade_id": "matplotlibex03a", "points": 2}
psi = well2d(np.linspace(0,1,10), np.linspace(0,1,10), 1, 1)
assert len(psi)==10
assert psi.shape==(10,)
# + [markdown] nbgrader={}
# The `contour`, `contourf`, `pcolor` and `pcolormesh` functions of Matplotlib can be used for effective visualizations of 2d scalar fields. Use the Matplotlib documentation to learn how to use these functions along with the `numpy.meshgrid` function to visualize the above wavefunction:
#
# * Use $n_x=3$, $n_y=2$ and $L=0$.
# * Use the limits $[0,1]$ for the x and y axis.
# * Customize your plot to make it effective and beautiful.
# * Use a non-default colormap.
# * Add a colorbar to you visualization.
#
# First make a plot using one of the contour functions:
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
# YOUR CODE HERE
f = plt.figure(figsize=(10,6))
x = np.linspace(0, 1, 100)
y = np.linspace(0, 1, 100)
w, v = np.meshgrid(x,y)
r = plt.contourf(well2d(w, v, 3, 2), cmap='BuGn')
plt.title('Wavefunction graph')
cbar = r.colorbar(ticks=[-1, 0, 1], orientation='horizontal')
"""
Colormap Possible values are: OrRd, flag, nipy_spectral, coolwarm, hsv_r, gnuplot2, prism, BrBG, afmhot_r, Spectral, Purples, Blues_r, YlGnBu, bone, summer_r, gnuplot2_r, Paired, YlGn, brg, gray_r, binary, ocean_r, spectral, Pastel2, afmhot, BrBG_r, YlGnBu_r, Set3_r, YlGn_r, binary_r, gist_gray, YlOrBr, Dark2_r, PuBuGn_r, Greys_r, winter, RdPu_r, Dark2, Pastel1, PuOr, RdBu, flag_r, GnBu_r, RdBu_r, copper, Paired_r, cool, brg_r, PRGn_r, PuOr_r, Oranges, gnuplot, Greys, hot_r, cool_r, RdYlBu_r, terrain_r, autumn, BuGn, gnuplot_r, bone_r, RdYlBu, Greens, gist_gray_r, spring_r, seismic, coolwarm_r, gist_earth, Set3, jet, RdYlGn_r, terrain, gist_rainbow_r, gist_ncar, PuBu, BuGn_r, Wistia, RdGy_r, summer, rainbow_r, CMRmap, hsv, Reds_r, YlOrRd_r, pink_r, Set2, YlOrBr_r, gray, BuPu_r, PRGn, Set1_r, rainbow, Spectral_r, gist_heat, spectral_r, RdYlGn, bwr, GnBu, CMRmap_r, gist_stern, copper_r, jet_r, gist_rainbow, PuRd, Pastel1_r, PuRd_r, Accent, Wistia_r, Reds, Greens_r, prism_r, BuPu, Pastel2_r, Purples_r, RdGy, Set2_r, Blues, autumn_r, Set1, pink, Oranges_r, gist_stern_r, ocean, gist_yarg, nipy_spectral_r, PuBuGn, Accent_r, gist_earth_r, spring, PiYG_r, RdPu, cubehelix_r, winter_r, seismic_r, bwr_r, PiYG, PuBu_r, gist_ncar_r, OrRd_r, YlOrRd, cubehelix, gist_yarg_r, hot, gist_heat_r
"""
# + deletable=false nbgrader={"checksum": "961e97980ad72d5d3aeace8b9915374a", "grade": true, "grade_id": "matplotlibex03b", "points": 4}
assert True # use this cell for grading the contour plot
# + [markdown] nbgrader={}
# Next make a visualization using one of the pcolor functions:
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
# YOUR CODE HERE
f = plt.figure(figsize=(10,6))
plt.pcolormesh(well2d(w, v, 3, 2), cmap='PuBuGn')
plt.title('Wavefunction graph')
# + deletable=false nbgrader={"checksum": "50b5f6fadc515274f80f58c1694c20ad", "grade": true, "grade_id": "matplotlibex03c", "points": 4}
assert True # use this cell for grading the pcolor plot
| assignments/assignment05/MatplotlibEx03.ipynb |