code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Loading and Manipulation
# ## Kindly load the las file of F02-1_logs.las well from the data folder
# ## Answer the following questions:
#
# >1. How many well logs in the file
# 2. How many data points (observations) in the welllogs.
# 3. Is there any null values? how much (%)/ well-log
# 4. Is there a relationship between DT and RHOB?
# ### You can use the following liberaries for your assignment:
# > Numpy, Pandas, Matplotlib, seaborn, LASIO
# !pip install lasio
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
import lasio
from IPython.display import display
# -
F02 = lasio.read("C:/Users/HP/Documents/GitHub/GeoML-2.0/10DaysChallenge/Dutch_F3_Logs/F02-1_logs.las")
type(F02)
F02.keys()
logs=F02.keys()
#No. of well logs
len(logs)
F02.data
df = F02.df()
df.head(7)
df.count()
#check the null values in the dataframe
df.isnull().sum()
Per=df.isnull().sum()/df.count()*100
print(Per, "%")
# +
plt.figure(figsize=(12,10))
plt.scatter(df.DT, df.RHOB, label= "DT vs. RHOB")
plt.xlabel("DT")
plt.ylabel("RHOB")
plt.title(" DT vs RHOB")
plt.legend()
| RY Day1 of 10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:probaprog]
# language: python
# name: conda-env-probaprog-py
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as pl
import pymc3 as pm
import arviz as az
from sklearn.preprocessing import scale
# -
df = pd.read_csv('./dataJar/WIMADISO.txt', delim_whitespace=True,
names=['day','month', 'year', 'temp'],
parse_dates=[['day', 'month', 'year']], na_values=-99)
df.rename(columns={'day_month_year': 'date'}, inplace=True)
df.head()
df['temp_s'] = scale(df.temp)
df.describe()
_, axs = pl.subplots(ncols=2, figsize=(12, 5))
df.plot(y=['temp', 'temp_s'], marker='.', ls='', ax=ax)
df.iloc[:8400].tail()
X = df.iloc[8401].index.values
df.loc[df.date.dt.month]
last_index = df.loc[((df.date.dt.month==12) & (df.date.dt.day==31))].index.values[-1]
ys.size
X = df.loc[:last_index].index.values
ys = df.loc[:last_index, 'temp_s'].values
X[-1], ys.size, X.size
daysperyear = 365.24219
wl = daysperyear / 2
ampPriorMax = 2
with pm.Model() as m1:
ρ = pm.Normal('ρ', 0, 1)
β0 = pm.Normal('β0', 0, 1)
β1 = pm.Normal('β1', 0, 1)
ar1 = pm.AR('ar1', ρ, sigma=1.0, shape=1, constant=False,
init=pm.Normal.dist(mu=0, sd=1))
amp = pm.Uniform('amp', 0, 2)
phase = pm.Uniform('phase', -183, 183)
ν = pm.Exponential('ν', 1/30)
σ = pm.Exponential('σ', 1)
trend_i = β0 + β1 * X + amp * pm.math.cos((X-phase) / wl)
μ_i = trend_i + ar1 + ϵ
y_i = pm.StudentT('y_i', mu=μ_i, sigma=σ, nu=ν, observed=ys)
with m1:
trace = pm.sample(tune=2000, cores=1)
| Exploring Kruschke'WMADISO data with trend and cycle and AR1 noise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Appendix C: True signal and FDR threshold adaptability
#
#
# This Appendix explores concepts associated with 'true signal' and how the FDR threshold adapts to different types of true signal. Let's first import the necessary packages.
import numpy as np
import scipy.stats
from matplotlib import pyplot
import spm1d #https://github.com/0todd0000/spm1d
import power1d #https://github.com/0todd0000/power1d
import fdr1d #https://github.com/0todd0000/fdr1d
# ___
#
# ### True signal model
#
# Next let's define use a Gaussian pulse as our (arbitrary) true signal:
# +
Q = 101 #number of continuum nodes
q = 60 #center of Gaussian pulse
sigma = 20 #standard deviation of the pulse
amp = 3.0 #amplitude of the pulse
signal = power1d.geom.GaussianPulse(Q=Q, q=q, sigma=sigma, amp=amp).toarray()
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
ax.plot( signal )
ax.axhline(0, color='k', ls=':')
ax.axvline(q, color='k', ls=':')
ax.set_xlabel('Continuum position')
ax.set_ylabel('DV value')
ax.set_title('True signal model')
pyplot.show()
# -
# Let's add this signal to some noise:
# +
np.random.seed(200)
J = 8 #sample size
FWHM = 20 #smoothness
noise = spm1d.rft1d.randn1d(J, Q, FWHM, pad=True) #random 1D residuals
y = signal + noise #random sample containing true signal
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
ax.plot( noise.T)
ax.axhline(0, color='k', ls=':')
ax.axvline(q, color='k', ls=':')
ax.set_xlabel('Continuum position')
ax.set_ylabel('DV value')
ax.set_title('Noise (without signal)')
pyplot.show()
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
ax.plot( y.T)
ax.axhline(0, color='k', ls=':')
ax.axvline(q, color='k', ls=':')
ax.set_xlabel('Continuum position')
ax.set_ylabel('DV value')
ax.set_title('Noise (with signal)')
pyplot.show()
# -
# ___
#
# ### Effect of true signal amplitude FDR threshold
#
# Next let's systematically vary the true signal amplitude, compute the t statistic separately for each magnitude, then check the effect on the computed FDR threshold. For simplicity we'll use the same noise each time. We'll also compute the RFT threshold for comparison.
# +
alpha = 0.05 #Type I error rate
df = J - 1 #degrees of freedom
amps = np.linspace(0, 3, 16)
tvalue = []
thresh = []
for a in amps:
y = (a * signal) + noise
t = y.mean(axis=0) / ( y.std(ddof=1, axis=0)/ (J**0.5) )
tstar = fdr1d.inference(t, df, alpha=alpha, stat='T')
tvalue.append(t)
thresh.append(tstar)
thresh_rft = spm1d.rft1d.t.isf(alpha, df, Q, FWHM)
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
ax.plot(amps, thresh, 'o-', color='b')
ax.axhline(thresh_rft, color='r', ls='--')
ax.text(1, 2.6, 'FDR threshold', color='b')
ax.text(2, 3.8, 'RFT threshold', color='r')
ax.set_ylim(2, 4)
ax.set_xlabel('True signal amplitude')
ax.set_ylabel('t value')
pyplot.show()
# -
# We can see the following:
#
# * The FDR threshold decreases as true signal amplitude increases
# * The RFT threshold is constant because it depends only on the smoothness of the noise
#
# Plotting the 1D t statistic clarifies why FDR decreases:
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
colors = ['r', 'g', 'b']
for i,ind in enumerate([2, 3, 5]):
ax.plot( tvalue[ind], color=colors[i], label='Amp = %.1f'%amps[ind] )
ax.axhline( thresh[ind], color=colors[i], ls='--')
ax.text(5, 3.5, 'FDR thresholds', color='k')
ax.axhline(0, color='k', ls=':')
ax.set_xlabel('Continuum position')
ax.set_ylabel('t value')
ax.legend()
pyplot.show()
# As the signal amplitude increases, an increasing number of continuum nodes exceed a given threshold. In order to maintain the **proportion of false positive nodes** at $\alpha$=0.05, FDR must reduce the threshold so that roughly 5% of the nodes are part of the noise (and not the signal). In other words, FDR regards a greater number of suprathreshold nodes as better evidence of the existence of a true signal, so adjusts the threshold downward in attempts to capture approximately 95% of the signal.
#
# However, this interpretation is valid only over a large number of experiments. FDR does not aim to yield false positive nodes at a rate of 5% for each 1D t continuum. Instead it aims to yield false positive nodes at a rate of 5% for the **infinite set of 1D t continua**.
#
# Let's run more simulations to check the long-term behavior of the FDR threshold.
# +
np.random.seed(0)
nIter = 1000
THRESH = []
for i in range(nIter):
thresh = []
for a in amps:
noise = spm1d.rft1d.randn1d(J, Q, FWHM) #random 1D residuals
y = (a * signal) + noise
t = y.mean(axis=0) / ( y.std(ddof=1, axis=0)/ (J**0.5) )
tstar = fdr1d.inference(t, df, alpha=alpha, stat='T')
if tstar is None:
tstar = np.nan
thresh.append( tstar )
THRESH.append( thresh )
#compute long-term average threshold
THRESH = np.array(THRESH)
thresh = THRESH.mean(axis=0)
pyplot.figure()
ax = pyplot.axes( [0, 0, 1, 1] )
ax.plot(amps, thresh, 'o-', color='b', label='FDR threshold')
ax.set_xlabel('True signal amplitude')
ax.set_ylabel('t value')
ax.legend()
pyplot.show()
# -
# We can see that, like the single datasat above, the FDR threshold reduces systematically with signal amplitude over a large number of datasets. However, this threshold reduction over the long term (from about $t$=2.3 to $t$=2.15) is not as pronounced as for the single dataset above (from about $t$=3.1 to $t$=2.15).
| Appendix/ipynb/AppendixC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Tutorial: Azure Machine Learning Quickstart
#
# In this tutorial, you learn how to quickly get started with Azure Machine Learning. Using a *compute instance* - a fully managed cloud-based VM that is pre-configured with the latest data science tools - you will train an image classification model using the CIFAR10 dataset.
#
# In this tutorial you will learn how to:
#
# * Create a compute instance and attach to a notebook
# * Train an image classification model and log metrics
# * Deploy the model
#
# ## Prerequisites
#
# 1. An Azure Machine Learning workspace
# 1. Familiar with the Python language and machine learning workflows.
#
#
# ## Create compute & attach to notebook
#
# To run this notebook you will need to create an Azure Machine Learning _compute instance_. The benefits of a compute instance over a local machine (e.g. laptop) or cloud VM are as follows:
#
# * It is a pre-configured with all the latest data science libaries (e.g. panads, scikit, TensorFlow, PyTorch) and tools (Jupyter, RStudio). In this tutorial we make extensive use of PyTorch, AzureML SDK, matplotlib and we do not need to install these components on a compute instance.
# * Notebooks are seperate from the compute instance - this means that you can develop your notebook on a small VM size, and then seamlessly scale up (and/or use a GPU-enabled) the machine when needed to train a model.
# * You can easily turn on/off the instance to control costs.
#
# To create compute, click on the + button at the top of the notebook viewer in Azure Machine Learning Studio:
#
# <img src="https://dsvmamlstorage127a5f726f.blob.core.windows.net/images/ci-create.PNG" width="500"/>
#
# This will pop up the __New compute instance__ blade, provide a valid __Compute name__ (valid characters are upper and lower case letters, digits, and the - character). Then click on __Create__.
#
# It will take approximately 3 minutes for the compute to be ready. When the compute is ready you will see a green light next to the compute name at the top of the notebook viewer:
#
# <img src="https://dsvmamlstorage127a5f726f.blob.core.windows.net/images/ci-create2.PNG" width="500"/>
#
# You will also notice that the notebook is attached to the __Python 3.6 - AzureML__ jupyter Kernel. Other kernels can be selected such as R. In addition, if you did have other instances you can switch to them by simply using the dropdown menu next to the Compute label.
#
# ## Import Data
#
# For this tutorial, you will use the CIFAR10 dataset. It has the classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck. The images in CIFAR-10 three-channel color images of 32x32 pixels in size.
#
# The code cell below uses the PyTorch API to download the data to your compute instance, which should be quick (around 15 seconds). The data is divided into training and test sets.
#
# * **NOTE: The data is downloaded to the compute instance (in the `/tmp` directory) and not a durable cloud-based store like Azure Blob Storage or Azure Data Lake. This means if you delete the compute instance the data will be lost. The [getting started with Azure Machine Learning tutorial series](https://docs.microsoft.com/azure/machine-learning/tutorial-1st-experiment-sdk-setup-local) shows how to create an Azure Machine Learning *dataset*, which aids durability, versioning, and collaboration.**
# + gather={"logged": 1600881820920}
import torch
import torch.optim as optim
import torchvision
import torchvision.transforms as transforms
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='/tmp/data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='/tmp/data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# -
# ## Take a look at the data
# In the following cell, you have some python code that displays the first batch of 4 CIFAR10 images:
# + gather={"logged": 1600882160868}
import matplotlib.pyplot as plt
import numpy as np
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
print(' '.join('%5s' % classes[labels[j]] for j in range(4)))
# -
# ## Train model and log metrics
#
# In the directory `model` you will see a file called [model.py](./model/model.py) that defines the neural network architecture. The model is trained using the code below.
#
# * **Note: The model training take around 4 minutes to complete. The benefit of a compute instance is that the notebooks are separate from the compute - therefore you can easily switch to a different size/type of instance. For example, you could switch to run this training on a GPU-based compute instance if you had one provisioned. In the code below you can see that we have included `torch.device("cuda:0" if torch.cuda.is_available() else "cpu")`, which detects whether you are using a CPU or GPU machine.**
# + gather={"logged": 1600882387754} tags=["local run"]
from model.model import Net
from azureml.core import Experiment
from azureml.core import Workspace
ws = Workspace.from_config()
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
exp = Experiment(workspace=ws, name="cifar10-experiment")
run = exp.start_logging(snapshot_directory=None)
# define convolutional network
net = Net()
net.to(device)
# set up pytorch loss / optimizer
criterion = torch.nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
run.log("learning rate", 0.001)
run.log("momentum", 0.9)
# train the network
for epoch in range(1):
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# unpack the data
inputs, labels = data[0].to(device), data[1].to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999:
loss = running_loss / 2000
run.log("loss", loss)
print(f'epoch={epoch + 1}, batch={i + 1:5}: loss {loss:.2f}')
running_loss = 0.0
print('Finished Training')
# -
# Once you have executed the cell below you can view the metrics updating in real time in the Azure Machine Learning studio:
#
# 1. Select **Experiments** (left-hand menu)
# 1. Select **cifar10-experiment**
# 1. Select **Run 1**
# 1. Select the **Metrics** Tab
#
# The metrics tab will display the following graph:
#
# <img src="https://dsvmamlstorage127a5f726f.blob.core.windows.net/images/metrics-capture.PNG" alt="dataset details" width="500"/>
# #### Understand the code
#
# The code is based on the [Pytorch 60minute Blitz](https://pytorch.org/tutorials/beginner/blitz/cifar10_tutorial.html#sphx-glr-beginner-blitz-cifar10-tutorial-py) where we have also added a few additional lines of code to track the loss metric as the neural network trains.
#
# | Code | Description |
# | ------------- | ---------- |
# | `experiment = Experiment( ... )` | [Experiment](https://docs.microsoft.com/python/api/azureml-core/azureml.core.experiment.experiment?view=azure-ml-py&preserve-view=true) provides a simple way to organize multiple runs under a single name. Later you can see how experiments make it easy to compare metrics between dozens of runs. |
# | `run.log()` | This will log the metrics to Azure Machine Learning. |
# ## Version control models with the Model Registry
#
# You can use model registration to store and version your models in your workspace. Registered models are identified by name and version. Each time you register a model with the same name as an existing one, the registry increments the version. Azure Machine Learning supports any model that can be loaded through Python 3.
#
# The code below does:
#
# 1. Saves the model on the compute instance
# 1. Uploads the model file to the run (if you look in the experiment on Azure Machine Learning studio you should see on the **Outputs + logs** tab the model has been saved in the run)
# 1. Registers the uploaded model file
# 1. Transitions the run to a completed state
# + gather={"logged": 1600888071066} tags=["register model from file"]
from azureml.core import Model
PATH = 'cifar_net.pth'
torch.save(net.state_dict(), PATH)
run.upload_file(name=PATH, path_or_stream=PATH)
model = run.register_model(model_name='cifar10-model',
model_path=PATH,
model_framework=Model.Framework.PYTORCH,
description='cifar10 model')
run.complete()
# -
# ### View model in the model registry
#
# You can see the stored model by navigating to **Models** in the left-hand menu bar of Azure Machine Learning Studio. Click on the **cifar10-model** and you can see the details of the model like the experiement run id that created the model.
# ## Deploy the model
#
# The next cell deploys the model to an Azure Container Instance so that you can score data in real-time (Azure Machine Learning also provides mechanisms to do batch scoring). A real-time endpoint allows application developers to integrate machine learning into their apps.
#
# * **Note: The deployment takes around 3 minutes to complete.**
# + tags=["deploy service", "aci"]
from azureml.core import Environment, Model
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice
environment = Environment.get(ws, "AzureML-PyTorch-1.6-CPU")
model = Model(ws, "cifar10-model")
service_name = 'cifar-service'
inference_config = InferenceConfig(entry_script='score.py', environment=environment)
aci_config = AciWebservice.deploy_configuration(cpu_cores=1, memory_gb=1)
service = Model.deploy(workspace=ws,
name=service_name,
models=[model],
inference_config=inference_config,
deployment_config=aci_config,
overwrite=True)
service.wait_for_deployment(show_output=True)
# -
# ### Understand the code
#
# | Code | Description |
# | ------------- | ---------- |
# | `environment = Environment.get()` | [Environment](https://docs.microsoft.com/python/api/overview/azure/ml/?view=azure-ml-py#environment) specify the Python packages, environment variables, and software settings around your training and scoring scripts. In this case, you are using a *curated environment* that has all the packages to run PyTorch. |
# | `inference_config = InferenceConfig()` | This specifies the inference (scoring) configuration for the deployment such as the script to use when scoring (see below) and on what environment. |
# | `service = Model.deploy()` | Deploy the model. |
#
# The [*scoring script*](score.py) file is has two functions:
#
# 1. an `init` function that executes once when the service starts - in this function you normally get the model from the registry and set global variables
# 1. a `run(data)` function that executes each time a call is made to the service. In this function, you normally deserialize the json, run a prediction and output the predicted result.
#
#
# ## Test the model service
#
# In the next cell, you get some unseen data from the test loader:
# +
dataiter = iter(testloader)
images, labels = dataiter.next()
# print images
imshow(torchvision.utils.make_grid(images))
print('GroundTruth: ', ' '.join('%5s' % classes[labels[j]] for j in range(4)))
# -
# Finally, the next cell runs scores the above images using the deployed model service.
# +
import json
input_payload = json.dumps({
'data': images.tolist()
})
output = service.run(input_payload)
print(output)
# -
# ## Clean up resources
#
# To clean up the resources after this quickstart, firstly delete the Model service using:
service.delete()
# Next stop the compute instance by following these steps:
#
# 1. Go to **Compute** in the left-hand menu of the Azure Machine Learning studio
# 1. Select your compute instance
# 1. Select **Stop**
#
#
# **Important: The resources you created can be used as prerequisites to other Azure Machine Learning tutorials and how-to articles.** If you don't plan to use the resources you created, delete them, so you don't incur any charges:
#
# 1. In the Azure portal, select **Resource groups** on the far left.
# 1. From the list, select the resource group you created.
# 1. Select **Delete resource group**.
# 1. Enter the resource group name. Then select **Delete**.
#
# You can also keep the resource group but delete a single workspace. Display the workspace properties and select **Delete**.
# ## Next Steps
#
# In this tutorial, you have seen how to run your machine learning code on a fully managed, pre-configured cloud-based VM called a *compute instance*. Having a compute instance for your development environment removes the burden of installing data science tooling and libraries (for example, Jupyter, PyTorch, TensorFlow, Scikit) and allows you to easily scale up/down the compute power (RAM, cores) since the notebooks are separated from the VM.
#
# It is often the case that once you have your machine learning code working in a development environment that you want to productionize this by running as a **_job_** - ideally on a schedule or trigger (for example, arrival of new data). To this end, we recommend that you follow [**the day 1 getting started with Azure Machine Learning tutorial**](https://docs.microsoft.com/azure/machine-learning/tutorial-1st-experiment-sdk-setup-local). This day 1 tutorial is focussed on running jobs-based machine learning code in the cloud.
| 2. Microsoft AI Classroom Series!/tutorials/quickstart/azureml-quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
import pandas as pd
pypoll = "election_data.csv"
pypoll_df = pd.read_csv(pypoll)
pypoll_df.head()
pypoll_df["Candidate"].value_counts()
pypoll_df.count
# total, candidate total
total = 3521001
k = 2218231
c = 704200
l = 492940
o = 105630
# print results
print("Election Results")
print("-----------------------------------")
print("Total Votes: " + str(total))
print("-----------------------------------")
print(("Kahn: ") + str(k) + (" ") + str(kk))
print(("Correy: ") + str(c) + (" ") + str(cc))
print(("Li: ") + str(l) + (" ") + str(ll))
print(("O'Tooley: ") + str(o) + (" ") + str(oo))
print("-----------------------------------")
print("Winner: Kahn")
print("-----------------------------------")
with open('pypoll_analysis.txt', 'w') as text:
text.write("Election Resulst\n")
text.write("-----------------------------------\n")
text.write("Total Votes: " + str(total) + "\n")
text.write("-----------------------------------\n")
text.write(("Kahn: ") + str(k) + (" ") + str(kk) + "\n")
text.write(("Correy: ") + str(c) + (" ") + str(cc) + "\n")
text.write(("Li: ") + str(l) + (" ") + str(ll) + "\n")
text.write(("O'Tooley: ") + str(o) + (" ") + str(oo) + "\n")
text.write("-----------------------------------\n")
text.write("Winner: Kahn" + "\n")
text.write("-----------------------------------\n")
| pypoll.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import requests
import seaborn as sns
from dotenv import load_dotenv
import os
load_dotenv()
API_KEY = os.getenv('BING_API_KEY')
# %config InlineBackend.figure_format = 'retina'
plt.rcParams = plt.rcParamsOrig
# -
cities = [
"Jakarta",
"Bogor",
"Bandung",
"Cikampek",
"Cirebon",
"Tasikmalaya",
"Tegal",
"Purwokerto",
"Semarang",
"Magelang",
"Yogyakarta",
"Tulungagung",
"Kediri",
"Malang",
"Surakarta",
"Surabaya"
]
# +
def get_coordinates(city):
query = f"{city}, Indonesia"
url = f"http://dev.virtualearth.net/REST/v1/Locations/{query}?key={API_KEY}"
resp = requests.get(url)
data = resp.json()
obj = data['resourceSets'][0]['resources'][0]
return obj['name'], obj['point']['coordinates']
def get_distances(locs):
city_coords = eval(locs.drop('city', axis=1).to_json(orient='records'))
payload = {
'origins': city_coords,
'destinations': city_coords,
'travelMode': 'driving'
}
headers = {
'Content-Length': '450',
'Content-Type': 'application/json'
}
url = f"https://dev.virtualearth.net/REST/v1/Routes/DistanceMatrix?key={API_KEY}"
resp = requests.post(url, json=payload, headers=headers)
return pd.DataFrame(
resp.json()['resourceSets'][0]['resources'][0]['results']
)
# +
from tqdm.notebook import tqdm
locations = []
for city in tqdm(cities):
locations.append(get_coordinates(city))
# -
locs = pd.DataFrame(
locations,
columns=['city', 'coordinates']
)
locs = pd.concat([
locs[['city']],
locs['coordinates'].apply(
pd.Series
).rename(
{
0: 'latitude',
1: 'longitude'
},
axis=1
)
], axis=1)
locs['city_name'] = locs['city'].apply(lambda x: x.split(',')[0].strip())
distances = get_distances(locs)
distances['source'] = distances['originIndex'].map(locs.city_name.to_dict())
distances['target'] = distances['destinationIndex'].map(locs.city_name.to_dict())
distances['weight'] = distances['travelDistance']
locs.to_csv('../datasets/java-coordinates.csv', index=False)
distances.to_csv('../datasets/java-dist-matrix.csv', index=False)
dist_matrix = distances.pivot_table(
index='source',
columns='target',
values='travelDistance'
)
# +
from sklearn.metrics.pairwise import haversine_distances
results = haversine_distances(
np.radians(locs[['longitude', 'latitude']])
)
heuristics = pd.DataFrame(
results * 6371000/1000, # multiply by Earth radius to get kilometer
columns=locs.city_name,
index=locs.city_name
)['Surabaya']
heuristics.sort_index().apply(np.round).astype(int)
# -
edges = {
"Jakarta": ["Bogor", "Cikampek"],
"Bogor": ["Bandung"],
"Bandung": ["Tasikmalaya", "Cirebon"],
"Cikampek": ["Bandung", "Cirebon"],
"Cirebon": ["Tegal"],
"Tasikmalaya": ["Purwokerto"],
"Tegal": ["Semarang", "Purwokerto"],
"Purwokerto": ["Magelang", "Yogyakarta"],
"Semarang": ["Surabaya", "Surakarta"],
"Magelang": ["Semarang", "Yogyakarta"],
"Yogyakarta": ["Surakarta", "Tulungagung"],
"Tulungagung": ["Malang", "Kediri"],
"Kediri": ["Surabaya"],
"Malang": ["Surabaya"],
"Surakarta": ["Surabaya"],
"Surabaya": []
}
routes = []
for source in edges:
for target in edges[source]:
routes.append((source, target))
# +
import networkx as nx
G = nx.from_pandas_edgelist(
distances.merge(
pd.DataFrame(
routes,
columns=['source', 'target']
),
how='inner',
on=['source', 'target']
),
edge_attr='weight'
)
# -
pos = locs.set_index('city_name').apply(
lambda x: (x['longitude'], x['latitude']),
axis=1
).to_dict()
# +
import geopandas
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
# -
gdf = geopandas.GeoDataFrame(
locs,
geometry=geopandas.points_from_xy(
locs.longitude,
locs.latitude)
)
# +
ax = plt.gca()
nx.draw(
G, pos,
with_labels=True,
edge_color='#DDDDDD',
node_color='#A0CBE2',
node_size=300,
font_size=10,
ax=ax
)
labels = nx.get_edge_attributes(G, 'weight')
labels = {k: np.round(v).astype(int) for k, v in labels.items()}
nx.draw_networkx_edge_labels(
G, pos,
edge_labels=labels,
ax=ax
);
# plt.savefig('../datasets/java.png', bbox_inches='tight', transparent=True, dpi=200)
# -
# %load_ext watermark
# %watermark --iversions
| scripts/lab4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: dca0ade3e726a953b501b15e8e990130d2b7799f14cfd9f4271676035ebe5511
# name: python3
# ---
import pandas as pd
df = pd.read_csv('adult.data.csv')
# df.set_index('race', drop=False)
df.head()
df['age'].size
race_count = df['race'].value_counts()
race_count
# average age of men
df[df['sex'] == 'Male']
df.loc[df['sex'] == 'Male', 'age'].mean()
df.loc[df['education'] == 'Bachelors'].count()
5355 / 32561
# percentage of people w/ Bachelor's degree?
len(df.loc[df['education'] == 'Bachelors'].index) / len(df.index) * 100
# percentage (num) of people w/ adv edu
higher_education = len(df.loc[((df['education'] == 'Bachelors') | (df['education'] == 'Masters') | (df['education'] == 'Doctorate'))])
higher_education
# now trying to take the w/ education grid and apply a >50k filter
df.loc[((df['education'] == 'Bachelors') | (df['education'] == 'Masters') | (df['education'] == 'Doctorate')) & (df['salary'] == '>50K')]
# +
# percent of people w/ higher ed degrees who make > 50k
len(df.loc[((df['education'] == 'Bachelors') | (df['education'] == 'Masters') | (df['education'] == 'Doctorate')) & (df['salary'] == '>50K')]) / higher_education * 100
# -
lower_education = len(df.loc[((df['education'] != 'Bachelors') & (df['education'] != 'Masters') & (df['education'] != 'Doctorate'))])
lower_education
# +
# people w/o higher ed
df.loc[((df['education'] != 'Bachelors') & (df['education'] != 'Masters') & (df['education'] != 'Doctorate')) & (df['salary'] == '>50K')]
# -
# percent of those w/o higher ed that make > 50k
len(df.loc[((df['education'] != 'Bachelors') & (df['education'] != 'Masters') & (df['education'] != 'Doctorate')) & (df['salary'] == '>50K')]) / lower_education * 100
# min # of hours worked per week
df.sort_values('hours-per-week').iloc[0, 12]
# +
# percent of people w/ min hours who make >50k
# first of all, assign the above # into the var min_work_hours
min_work_hours = df.sort_values('hours-per-week').iloc[0, 12]
# second, calc the # of peole who work 1 hr per week
num_min_workers = len(df.loc[df['hours-per-week'] == min_work_hours])
# then, calc the percentage of those who work the min_work_hours who make >50k
len(df.loc[(df['hours-per-week'] == min_work_hours) & (df['salary'] == '>50K')]) / num_min_workers * 100
# -
len(df.loc[df['hours-per-week'] == min_work_hours]) # test cell
df['native-country'].unique()
df['native-country'].value_counts()
df.info()
df.loc[df['salary'] == '>50K']
df[['native-country', 'salary']]
# returns highest earning country
highest_earning_country = [df.loc[(df['salary'] == '>50K'), 'native-country'].value_counts(), df[['native-country']].value_counts()]
highest_earning_country
# greater than 50k series
gt50ks = df.loc[(df['salary'] == '>50K'), 'native-country'].value_counts()
country_series = df[['native-country']].value_counts()
# greater than 50k salary percentages (by country)
gt50ks_percents = pd.Series()
for i, v in gt50ks.iteritems():
gt50ks_percents[i] = (gt50ks[i] / country_series[i]) * 100
gt50ks_percents = gt50ks_percents.max(level=0) # used to clean up the series so .idxmax() works
highest_earning_country = gt50ks_percents.idxmax()
# new solution to highest_earning_country_percentage
highest_earning_country_percentage = round((len(df.loc[((df['native-country'] == highest_earning_country) & (df['salary'] == '>50K'))]) / len(df.loc[(df['native-country'] == highest_earning_country)]) * 100), 1)
highest_earning_country_percentage
highest_earning_country_percentage = round((len(df.loc[(df['salary'] == '>50K') & (df['native-country'] == highest_earning_country)]) / len(df.index) * 100), 1)
highest_earning_country_percentage
# most pop. occupation for those who earn > 50, in IN
df.loc[(df['native-country'] == 'India') & (df['salary'] == '>50K')]['occupation'].value_counts().index[0]
| sketches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Discrete parameters
#
# In this notebook, we discuss pyABC's abilities to deal with discrete parameters.
# In its Sequential Monte Carlo algorithm, pyABC uses transition kernels $g_t$ that propose parameters for generation $t$ by slightly perturbing parameters that were accepted in generation $t-1$. Thus gradually sampling from a better and better approximation of the posterior, instead of the prior, we can reduce the acceptance threshold $\varepsilon$ while maintaining high acceptance rates. To account for no longer sampling from the prior $\pi$, we need to weight particles by $\pi(\theta) / g_t(\theta)$.
#
# A formal requirement for any transition is that the prior is absolutely continuous with respect to it, i.e. $g_t\gg\pi$. This means that any value that has positive mass under the prior can still be sampled under the transition. Other than that, the transition can be arbitrary. Commonly, it should be close to the underlying posterior in order to achieve high acceptance rates. pyABC offers various transitions kernels that adjust automatically to the problem structure and thus deliver high-quality proposals. The default transition kernel `pyabc.MultivariateNormalTransition` as well as the `pyabc.LocalTransition` kernel both employ (localized/recscaled) multivariate normal kernels with adaptive covariance matrices. Therefore, they can learn about parameter scales and correlations.
#
# However, they can only deal with continuous parameters.
# Sometimes, parameters can only take discrete values and should thus also be modeled as such, e.g. via a discrete integer-valued prior. In that case, the above transitions cannot be applied, as they propose non-integer parameters. The, as far as implementation is concerned, most straightforward way to deal with this is to just round the suggested parameters to the next integer. However, to then obtain statistically correct weights, one would need to integrate the transition density over all values that are rounded to the same value. This can be computationally non-trivial and is also not implemented in pyABC, where one would instead just take the density value at the actually sampled parameters as an approximation, which is not strictly correct.
#
# To efficiently and correctly deal with discrete parameters, pyABC also implements discrete transitions as classes derived from `pyabc.transition.DiscreteTransition`. Let us show an example:
# install if not done yet
# !pip install pyabc --quiet
import numpy as np
import matplotlib.pyplot as plt
import pyabc
# %matplotlib inline
pyabc.settings.set_figure_params('pyabc') # for beautified plots
# We assume that we have a model with two parameters: A continuous variance parameter, and a mean parameter that can for some reason only take integer values from 0 to 6. To spice things up a little and get a non-trivial posterior distribution, the model randomly perturbs the mean by an offset of $\pm 2$.
# +
n_data = 100
def model(p):
return {'y': p['p_discrete'] + np.random.choice([-2, 0, 2], p=[0.2, 0.5, 0.3])
+ p['p_continuous'] * np.random.normal(size=n_data)}
distance = lambda x, x0: sum((x['y'] - x0['y'])**2)
p_true = {'p_discrete': 2, 'p_continuous': 0.5}
obs = model(p_true)
# -
# plot the data
ax = plt.hist(obs['y'])
plt.xlabel("Observable y_i")
plt.ylabel("Frequency")
plt.title("Data")
plt.show()
# Currently, there are two classes offered. First, there is the `pyabc.DiscreteRandomWalkTransition`, which is similarly also implemented in other tools. First it samples a parameter value from the weighted last generation and then performs a random walk of a fixed number of steps. While it can deal with unbounded discrete domains, it has the disadvantage, that it violates the absolute contiunuity condition and will thus not be further considered here. Let us know if you would like to discuss further details on this.
#
# Here, we will in the following focus on the second implemented discrete transition, the `pyabc.DiscreteJumpTransition`, which requires a finite domain. It also first samples a parameter value from the weighted last generation, and ensures absolute positivity by with a positive probability `1 - p_stay` move to an arbitrary other value within the domain.
#
# As we here also have another, continuous, parameter (for which we just use a `pyabc.MultivariateNormalTransition`), we use a `pyabc.AggregatedTransition` to combine both.
# +
# domain of possible discrete values
discrete_domain = np.arange(7)
# priors
prior = pyabc.Distribution(
p_discrete=pyabc.RV('rv_discrete', values=(discrete_domain, [1/7] * 7)),
p_continuous=pyabc.RV('uniform', 0, 2))
# transition kernels
transition = pyabc.AggregatedTransition(mapping={
'p_discrete': pyabc.DiscreteJumpTransition(domain=discrete_domain, p_stay=0.7),
'p_continuous': pyabc.MultivariateNormalTransition()})
# -
# The transition kernels adjust to the problem structure via `fit()` functions that are called after each generation.
#
# Let's run it:
abc = pyabc.ABCSMC(model, prior, distance, transitions=transition, population_size=1000)
abc.new(pyabc.create_sqlite_db_id(), obs)
history = abc.run(max_nr_populations=5)
# We see in the result that we perfectly reproduce the 3 likely mean parameters with their respective probabilities. Over time, the posterior approximation get gradually more accurate.
# some visualizations
fig, axes = plt.subplots(history.max_t+1, 2, figsize=(6, 12))
for t in range(history.max_t+1):
pyabc.visualization.plot_kde_1d_highlevel(
history, 'p_continuous', t=t, refval=p_true, xmin=0, xmax=2,
ax=axes[t, 0])
pyabc.visualization.plot_histogram_1d(
history, 'p_discrete', t=t, bins=np.arange(8), align='left',
ax=axes[t, 1])
axes[t, 0].set_ylabel(f"Posterior t={t}")
fig.tight_layout()
| doc/examples/discrete_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # License
#
# Copyright 2018 <NAME>
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# # Automatic feature engineering using deep learning and Bayesian inference: Application to computer vision and synthetic financial transactions data
# ## Author: <NAME>
#
# We will explore the use of autoencoders for automatic feature engineering. The idea is to automatically learn a set of features from raw data that can be useful in supervised learning tasks such as in computer vision and insurance.
#
# ## Computer Vision
#
# We will use the MNIST dataset for this purpose where the raw data is a 2 dimensional tensor of pixel intensities per image. The image is our unit of analysis: We will predict the probability of each class for each image. This is a multiclass classification task and we will use the accuracy score to assess model performance on the test fold.
#
# 
#
# ## Insurance
#
# We will use a synthetic dataset where the raw data is a 2 dimensional tensor of historical policy level information per policy-period combination: Per unit this will be $\mathbb{R}^{4\times3}$, i.e., 4 historical time periods and 3 transactions types. The policy-period combination is our unit of analysis: We will predict the probability of loss for time period 5 in the future - think of this as a potential renewal of the policy for which we need to predict whether it would make a loss for us or not hence affecting whether we decided to renew the policy and / or adjust the renewal premium to take into account the additional risk. This is a binary class classification task and we will use the AUROC score to assess model performance.
#
# 
# +
import os
import math
import sys
import importlib
import numpy as np
import pandas as pd
from sklearn import linear_model
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import MinMaxScaler, LabelBinarizer, RobustScaler, StandardScaler
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from scipy.stats import norm
import keras
from keras import backend as bkend
from keras.datasets import cifar10, mnist
from keras.layers import Dense, BatchNormalization, Dropout, Flatten, convolutional, pooling
from keras import metrics
from autoencoders_keras.get_session import get_session
import keras.backend.tensorflow_backend as KTF
KTF.set_session(get_session(gpu_fraction=0.75, allow_soft_placement=True, log_device_placement=False))
import tensorflow as tf
from tensorflow.python.client import device_lib
from plotnine import *
import matplotlib.pyplot as plt
from autoencoders_keras.vanilla_autoencoder import VanillaAutoencoder
from autoencoders_keras.convolutional_autoencoder import ConvolutionalAutoencoder
from autoencoders_keras.convolutional2D_autoencoder import Convolutional2DAutoencoder
from autoencoders_keras.seq2seq_autoencoder import Seq2SeqAutoencoder
from autoencoders_keras.variational_autoencoder import VariationalAutoencoder
# %matplotlib inline
np.set_printoptions(suppress=True)
os.environ["KERAS_BACKEND"] = "tensorflow"
importlib.reload(bkend)
print(device_lib.list_local_devices())
mnist = mnist.load_data()
(X_train, y_train), (X_test, y_test) = mnist
X_train = np.reshape(X_train, [X_train.shape[0], X_train.shape[1] * X_train.shape[1]])
X_test = np.reshape(X_test, [X_test.shape[0], X_test.shape[1] * X_test.shape[1]])
y_train = y_train.ravel()
y_test = y_test.ravel()
X_train = X_train.astype("float32")
X_test = X_test.astype("float32")
X_train /= 255.0
X_test /= 255.0
# -
# ## Scikit-learn
#
# We will use the Python machine learning library scikit-learn for data transformation and the classification task. Note that we will code the autoencoders as scikit-learn transformers such that they can be readily used by scikit-learn pipelines.
scaler_classifier = MinMaxScaler(feature_range=(0.0, 1.0))
logistic = linear_model.LogisticRegression(random_state=666)
linear_mod = linear_model.ElasticNetCV()
lb = LabelBinarizer()
lb = lb.fit(y_train.reshape(y_train.shape[0], 1))
# ## MNIST: No Autoencoders
#
# We run the MNIST dataset without using an autoencoder. The 2 dimensional tensor of pixel intensities per image for MNIST images are of dimension $\mathbb{R}^{28 \times 28}$. We reshape them as a 1 dimensional tensor of dimension $\mathbb{R}^{784}$ per image. Therefore we have 784, i.e., $28 \times 28 = 784$, features for this supervised learning task per image.
#
# ### Results
#
# The accuracy score for the MNIST classification task without autoencoders: 92.000000%.
# +
pipe_base = Pipeline(steps=[("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_base = pipe_base.fit(X_train, y_train)
acc_base = pipe_base.score(X_test, y_test)
print("The accuracy score for the MNIST classification task without autoencoders: %.6f%%." % (acc_base * 100))
# -
# ## MNIST: PCA
#
# We use a PCA filter that picks the number of components that explain $99\%$ of the variation.
#
# ### Results
#
# The accuracy score for the MNIST classification task with PCA: 91.430000%.
# +
pipe_pca = Pipeline(steps=[("PCA", PCA(n_components=0.99)),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_pca = pipe_base.fit(X_train, y_train)
acc_pca = pipe_pca.score(X_test, y_test)
print("The accuracy score for the MNIST classification task with PCA: %.6f%%." % (acc_pca * 100))
# -
# ## MNIST: Vanilla Autoencoders
#
# An autoencoder is an unsupervised learning technique where the objective is to learn a set of features that can be used to reconstruct the input data.
#
# Our input data is $X \in \mathbb{R}^{N \times 784}$. An encoder function $E$ maps this to a set of $K$ features such that $E: \mathbb{R}^{N \times 784} \rightarrow \mathbb{R}^{N \times K}$. A decoder function $D$ uses the set of $K$ features to reconstruct the input data such that $D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 784}$.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 784} \\
# &E: \mathbb{R}^{N \times 784} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 784}
# \end{align*}
#
# Lets denote the reconstructed data as $\tilde{X} = D(E(X))$. The goal is to learn the encoding and decoding functions such that we minimize the difference between the input data and the reconstructed data. An example for an objective function for this task can be the Mean Squared Error (MSE) such that $\frac{1}{N}||\tilde{X} - X||^{2}_{2}$.
#
# We learn the encoding and decoding functions by minimizing the MSE using the parameters that define the encoding and decoding functions: The gradient of the MSE with respect to the parameters are calculated using the chain rule, i.e., backpropagation, and used to update the parameters via an optimization algorithm such as Stochastic Gradient Descent (SGD).
#
# Lets assume we have a single layer autoencoder using the Exponential Linear Unit (ELU) activation function, batch normalization, dropout and the Adaptive Moment (Adam) optimization algorithm. $B$ is the batch size, $K$ is the number of features.
#
# * **Exponential Linear Unit:** The activation function is smooth everywhere and avoids the vanishing gradient problem as the output takes on negative values when the input is negative. $\alpha$ is taken to be $1.0$.
#
# \begin{align*}
# H_{\alpha}(z) &=
# \begin{cases}
# &\alpha\left(\exp(z) - 1\right) \quad \text{if} \quad z < 0 \\
# &z \quad \text{if} \quad z \geq 0
# \end{cases} \\
# \frac{dH_{\alpha}(z)}{dz} &=
# \begin{cases}
# &\alpha\left(\exp(z)\right) \quad \text{if} \quad z < 0 \\
# &1 \quad \text{if} \quad z \geq 0
# \end{cases}
# \end{align*}
#
# * **Batch Normalization:** The idea is to transform the inputs into a hidden layer's activation functions. We standardize or normalize first using the mean and variance parameters on a per feature basis and then learn a set of scaling and shifting parameters on a per feature basis that transforms the data. The following equations describe this layer succintly: The parameters we learn in this layer are $\left(\mu_{j}, \sigma_{j}^2, \beta_{j}, \gamma_{j}\right) \quad \forall j \in \{1, \dots, K\}$.
#
# \begin{align*}
# \mu_{j} &= \frac{1}{B} \sum_{i=1}^{B} X_{i,j} \quad &\forall j \in \{1, \dots, K\} \\
# \sigma_{j}^2 &= \frac{1}{B} \sum_{i=1}^{B} \left(X_{i,j} - \mu_{j}\right)^2 \quad &\forall j \in \{1, \dots, K\} \\
# \hat{X}_{:,j} &= \frac{X_{:,j} - \mu_{j}}{\sqrt{\sigma_{j}^2 + \epsilon}} \quad &\forall j \in \{1, \dots, K\} \\
# Z_{:,j} &= \gamma_{j}\hat{X}_{:,j} + \beta_{j} \quad &\forall j \in \{1, \dots, K\}
# \end{align*}
#
# * **Dropout:** This regularization technique simply drops the outputs from input and hidden units with a certain probability say $50\%$.
#
# * **Adam Optimization Algorithm:** This adaptive algorithm combines ideas from the Momentum and RMSProp optimization algorithms. The goal is to have some memory of past gradients which can guide future parameters updates. The following equations for the algorithm succintly describe this method assuming $\theta$ is our set of parameters to be learnt and $\eta$ is the learning rate.
#
# \begin{align*}
# m &\leftarrow \beta_{1}m + \left[\left(1 - \beta_{1}\right)\left(\nabla_{\theta}\text{MSE}\right)\right] \\
# s &\leftarrow \beta_{2}s + \left[\left(1 - \beta_{2}\right)\left(\nabla_{\theta}\text{MSE} \otimes \nabla_{\theta}\text{MSE} \right)\right] \\
# \theta &\leftarrow \theta - \eta m \oslash \sqrt{s + \epsilon}
# \end{align*}
#
# ### Results
#
# The accuracy score for the MNIST classification task with an autoencoder: 96.940000%.
# +
autoencoder = VanillaAutoencoder(n_feat=X_train.shape[1],
n_epoch=50,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=1000,
encoding_dim=500,
denoising=None)
print(autoencoder.autoencoder.summary())
pipe_autoencoder = Pipeline(steps=[("autoencoder", autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_autoencoder = pipe_autoencoder.fit(X_train, y_train)
acc_autoencoder = pipe_autoencoder.score(X_test, y_test)
print("The accuracy score for the MNIST classification task with an autoencoder: %.6f%%." % (acc_autoencoder * 100))
# -
# ## MNIST: Denoising Autoencoders
#
# The idea here is to add some noise to the data and try to learn a set of robust features that can reconstruct the non-noisy data from the noisy data. The MSE objective functions is as follows, $\frac{1}{N}||D(E(X + \epsilon)) - X||^{2}_{2}$, where $\epsilon$ is some noise term.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 784} \\
# &E: \mathbb{R}^{N \times 784} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 784}
# \end{align*}
#
# ### Results
#
# The accuracy score for the MNIST classification task with a denoising autoencoder: 96.930000%.
# +
noise = 0.10 * np.reshape(np.random.uniform(low=0.0,
high=1.0,
size=X_train.shape[0] * X_train.shape[1]),
[X_train.shape[0], X_train.shape[1]])
denoising_autoencoder = VanillaAutoencoder(n_feat=X_train.shape[1],
n_epoch=50,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=1000,
encoding_dim=500,
denoising=noise)
print(denoising_autoencoder.autoencoder.summary())
pipe_denoising_autoencoder = Pipeline(steps=[("autoencoder", denoising_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_denoising_autoencoder = pipe_denoising_autoencoder.fit(X_train, y_train)
acc_denoising_autoencoder = pipe_denoising_autoencoder.score(X_test, y_test)
print("The accuracy score for the MNIST classification task with a denoising autoencoder: %.6f%%." % (acc_denoising_autoencoder * 100))
# -
# ## MNIST: 1 Dimensional Convolutional Autoencoders
#
# So far we have used flattened or reshaped raw data. Such a 1 dimensional tensor of pixel intensities per image, $\mathbb{R}^{784}$, might not take into account useful spatial features that the 2 dimensional tensor, $\mathbb{R}^{28\times28}$, might contain. To overcome this problem, we introduce the concept of convolution filters, considering first their 1 dimensional version and then their 2 dimensional version.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 28 \times 28} \\
# &E: \mathbb{R}^{N \times 28 \times 28} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 28 \times 28}
# \end{align*}
#
# The ideas behind convolution filters are closely related to handcrafted feature engineering: One can view the handcrafted features as simply the result of a predefined convolution filter, i.e., a convolution filter that has not been learnt based on the raw data at hand.
#
# Suppose we have raw transactions data per some unit of analysis, i.e., mortgages, that will potentially help us in classifying a unit as either defaulted or not defaulted. We will keep this example simple by only allowing the transaction values to be either \$100 or \$0. The raw data per unit spans 5 time periods while the defaulted label is for the next period, i.e., period 6. Here is an example of a raw data for a particular unit:
#
# \begin{align*}
# x =
# \begin{array}
# {l}
# \text{Period 1} \\ \text{Period 2} \\ \text{Period 3} \\ \text{Period 4} \\ \text{Period 5}
# \end{array}
# \left[
# \begin{array}
# {c}
# \$0 \\ \$0 \\ \$100 \\ \$0 \\ \$0
# \end{array}
# \right]
# \end{align*}
#
# Suppose further that if the average transaction value is \$20 then we will see a default in period 6 for this particular mortgage unit. Otherwise we do not see a default in period 6. The average transaction value is an example of a handcrafted feature: A predefined handcrafted feature that has not been learnt in any manner. It has been arrived at via domain knowledge of credit risk. Denote this as $\mathbf{H}(x)$.
#
# The idea of learning such a feature is an example of a 1 dimensional convolution filter. As follows:
#
# \begin{align*}
# \mathbf{C}(x|\alpha) = \alpha_1 x_1 + \alpha_2 x_2 + \alpha_3 x_3 + \alpha_4 x_4 + \alpha_5 x_5
# \end{align*}
#
# Assuming that $\mathbf{H}(x)$ is the correct representation of the raw data for this supervised learning task then the optimal set of parameters learnt via supervised learning, or perhaps unsupervised learning and then transferred to the supervised learning task, i.e., transfer learning, for $\mathbf{C}(x|\alpha)$ is as follows where $\alpha$ is $\left[0.2, 0.2, 0.2, 0.2, 0.2\right]$:
#
# \begin{align*}
# \mathbf{C}(x|\alpha) = 0.2 x_1 + 0.2 x_2 + 0.2 x_3 + 0.2 x_4 + 0.2 x_5
# \end{align*}
#
# This is a simple example however this clearly illusrates the principle behind using deep learning for automatic feature engineering or representation learning. One of the main benefits of learning such a representation in an unsupervised manner is that the same representation can then be used for multiple supervised learning tasks: Transfer learning. This is a principled manner of learning a representation from raw data.
#
# To summarize the 1 dimensional convolution filter for our simple example is defined as:
#
# \begin{align*}
# \mathbf{C}(x|\alpha)&= x * \alpha \\
# &= \sum_{t=1}^{5} x_t \alpha_t
# \end{align*}
#
# * $x$ is the input.
# * $\alpha$ is the kernel.
# * The output $x * \alpha$ is called a feature map and $*$ is the convolution operator or filter. This is the main difference between a vanilla neural network and a convolution neural network: We replace the matrix multiplication operator by the convolution operator.
# * Depending on the task at hand we can have different types of convolution filters.
# * Kernel size can be altered. In our example the kernel size is 5.
# * Stride size can be altered. In our example we had no stride size however suppose that stride size was 1 and kernel size was 2, i.e., $\alpha = \left[\alpha_1, \alpha_2\right]$, then we would apply the kernel $\alpha$ at the start of the input, i.e., $\left[x_1, x_2\right] * \left[\alpha_1, \alpha_2\right]$, and move the kernel over the next area of the input, i.e., $\left[x_2, x_3\right] * \left[\alpha_1, \alpha_2\right]$, and so on and so forth until we arrive at a feature map that consists of 4 real values. This is called a valid convolution while a padded, i.e., say padded with zero values, convolution would give us a feature map that is the same size as the input, i.e., 5 real values in our example.
# * We can apply an activation function to the feature maps such as ELU mentioned earlier.
# * Finally we can summarize the information contained in feature maps by taking a maximum or average value over a defined portion of the feature map. For instance, if after using a valid convolution we arrive at a feature map of size 4 and then apply a max pooling operation with size 4 then we will be taking the maximum value of this feature map. The result is another feature map.
#
# This automates feature engineering however introduces architecture engineering where different architectures consisting of various convolution filters, activation functions, batch normalization layers, dropout layers and pooling operators can be stacked together in a pipeline in order to learn a good representation of the raw data. One usually creates an ensemble of such architectures.
#
# The goal behind convolutional autoencoders is to use convolution filters, activation functions, batch normalization layers, dropout layers and pooling operators to create an encoder function which will learn a good representation of our raw data. The decoder will also use a similar set of layers as the encoder to reconstruct the raw data with one exception: Instead of using a pooling operator it will use an upsampling operator. The basic idea behind the upsampling operator is to repeat an element a certain number of times say size 4: One can view this as the inverse operator to the pooling operator. The pooling operator is essentially a downsampling operator and the upsampling operator is simply the inverse of that in some sense.
#
# ### Results
#
# The accuracy score for the MNIST classification task with a 1 dimensional convolutional autoencoder: 97.570000%.
# +
convolutional_autoencoder = ConvolutionalAutoencoder(input_shape=(int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))),
n_epoch=50,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
filters=100,
kernel_size=8,
strides=1,
pool_size=4,
denoising=None)
print(convolutional_autoencoder.autoencoder.summary())
pipe_convolutional_autoencoder = Pipeline(steps=[("autoencoder", convolutional_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_convolutional_autoencoder = pipe_convolutional_autoencoder.fit(np.reshape(X_train, [X_train.shape[0], int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))]),
y_train)
acc_convolutional_autoencoder = pipe_convolutional_autoencoder.score(np.reshape(X_test, [X_test.shape[0], int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))]), y_test)
print("The accuracy score for the MNIST classification task with a 1 dimensional convolutional autoencoder: %.6f%%." % (acc_convolutional_autoencoder * 100))
# -
# ## MNIST: Sequence to Sequence Autoencoders
#
# Given our mortgage default example a potentially more useful deep learning architecture might be the Recurrent Neural Network (RNN), specifically their state of the art variant the Long Short Term Memory (LSTM) network. The goal is to explicitly take into account the sequential nature of the raw data.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 28 \times 28} \\
# &E: \mathbb{R}^{N \times 28 \times 28} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 28 \times 28}
# \end{align*}
#
# The gradients in a RNN depend on the parameter matrices defined for the model. Simply put these parameter matrices can end up being multiplied many times over and hence cause two major problems for learning: Exploding and vanishing gradients. If the spectral radius of the parameter matrices, i.e., the maximum absolute value of the eigenvalues of a matrix, is more than 1 then gradients can become large enough, i.e., explode in value, such that learning diverges and similarly if the spectral radius is less than 1 then gradients can become small, i.e., vanish in value, such that the next best transition for the parameters cannot be reliably calculated. Appropriate calculation of the gradient is important for estimating the optimal set of parameters that define a machine learning method and the LSTM network overcomes these problems in a vanilla RNN. We now define the LSTM network for 1 time step, i.e., 1 memory cell.
#
# We calculate the value of the input gate, the value of the memory cell state at time period $t$ where $f(x)$ is some activation function and the value of the forget gate:
#
# \begin{align*}
# i_{t} &= \sigma(W_{i}x_{t} + U_{i}h_{t-1} + b_{i}) \\
# \tilde{c_{t}} &= f(W_{c}x_{t} + U_{c}h_{t-1} + b_{c}) \\
# f_{t} &= \sigma(W_{f}x_{t} + U_{f}h_{t-1} + b_{f})
# \end{align*}
#
# The forget gate controls the amount the LSTM remembers, i.e., the value of the memory cell state at time period $t-1$ where $\otimes$ is the hadamard product:
#
# \begin{align*}
# c_{t} = i_{t} \otimes \tilde{c_{t}} + f_{t} \otimes c_{t-1}
# \end{align*}
#
# With the updated state of the memory cell we calculate the value of the outputs gate and finally the output value itself:
#
# \begin{align*}
# o_{t} &= \sigma(W_{o}x_{t} + U_{o}h_{t-1} + b_{o}) \\
# h_{t} &= o_{t} \otimes f(c_{t})
# \end{align*}
#
# We can have a wide variety of LSTM architectures such as the convolutional LSTM where note that we replace the matrix multiplication operators in the input gate, the initial estimate $\tilde{c_{t}}$ of the memory cell state, the forget gate and the output gate by the convolution operator $*$:
#
# \begin{align*}
# i_{t} &= \sigma(W_{i} * x_{t} + U_{i} * h_{t-1} + b_{i}) \\
# \tilde{c_{t}} &= f(W_{c} * x_{t} + U_{c} * h_{t-1} + b_{c}) \\
# f_{t} &= \sigma(W_{f} * x_{t} + U_{f} * h_{t-1} + b_{f}) \\
# c_{t} &= i_{t} \otimes \tilde{c_{t}} + f_{t} \otimes c_{t-1} \\
# o_{t} &= \sigma(W_{o} * x_{t} + U_{o} * h_{t-1} + b_{o}) \\
# h_{t} &= o_{t} \otimes f(c_{t})
# \end{align*}
#
# Another popular variant is the peephole LSTM where the gates are allowed to peep at the memory cell state:
#
# \begin{align*}
# i_{t} &= \sigma(W_{i}x_{t} + U_{i}h_{t-1} + V_{i}c_{t-1} + b_{i}) \\
# \tilde{c_{t}} &= f(W_{c}x_{t} + U_{c}h_{t-1} + V_{c}c_{t-1} + b_{c}) \\
# f_{t} &= \sigma(W_{f}x_{t} + U_{f}h_{t-1} + V_{f}c_{t-1} + b_{f}) \\
# c_{t} &= i_{t} \otimes \tilde{c_{t}} + f_{t} \otimes c_{t-1} \\
# o_{t} &= \sigma(W_{o}x_{t} + U_{o}h_{t-1} + V_{o}c_{t} + b_{o}) \\
# h_{t} &= o_{t} \otimes f(c_{t})
# \end{align*}
#
# The goal for the sequence to sequence autoencoder is to create a representation of the raw data using a LSTM as an encoder. This representation will be a sequence of vectors say, $h_{1}, \dots, h_{T}$, learnt from a sequence of raw data vectors say, $x_{1}, \dots, x_{T}$. The final vector of the representation, $h_{T}$, is our encoded representation, also called a context vector. This context vector is repeated as many times as the length of the sequence such that it can be used as an input to a decoder which is yet another LSTM. The decoder LSTM will use this context vector to recontruct the sequence of raw data vectors, $\tilde{x_{1}}, \dots, \tilde{x_{T}}$. If the context vector is useful in the recontruction task then it can be further used for other tasks such as predicting default risk as given in our example.
#
# ### Results
#
# The accuracy score for the MNIST classification task with a sequence to sequence autoencoder: 97.600000%.
# +
seq2seq_autoencoder = Seq2SeqAutoencoder(input_shape=(int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))),
n_epoch=50,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=200,
encoding_dim=200,
stateful=False,
denoising=None)
print(seq2seq_autoencoder.autoencoder.summary())
pipe_seq2seq_autoencoder = Pipeline(steps=[("autoencoder", seq2seq_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_seq2seq_autoencoder = pipe_seq2seq_autoencoder.fit(np.reshape(X_train, [X_train.shape[0], int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))]),
y_train)
acc_seq2seq_autoencoder = pipe_seq2seq_autoencoder.score(np.reshape(X_test, [X_test.shape[0], int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5))]), y_test)
print("The accuracy score for the MNIST classification task with a sequence to sequence autoencoder: %.6f%%." % (acc_seq2seq_autoencoder * 100))
# -
# ## MNIST: Variational Autoencoders
#
# We now combine Bayesian inference with deep learning by using variational inference to train a vanilla autoencoder. This moves us towards generative modelling which can have further use cases in semi-supervised learning. The other benefit of training using Bayesian inference is that we can be more robust to higher capacity deep learners, i.e., avoid overfitting.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 784} \\
# &E: \mathbb{R}^{N \times 784} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 784}
# \end{align*}
#
# * Assume $X$ is our raw data while $Z$ is our learnt representation.
# * We have a prior belief on our learnt representation:
#
# \begin{align*}
# p(Z)
# \end{align*}
#
# * The posterior distribution for our learnt representation is:
#
# \begin{align*}
# p(Z|X)=\frac{p(X|Z)p(Z)}{p(X)}
# \end{align*}
#
# * The marginal likelihood, $p(X)$, is often intractable causing the posterior distribution, $p(Z|X)$, to be intractable:
#
# \begin{align*}
# p(X)=\int_{Z}p(X|Z)p(Z)dZ
# \end{align*}
#
# * We therefore need an approximate posterior distribution via variational inference that can deal with the intractability. This additionally also provides the benefit of dealing with large scale datasets as generally Markov Chain Monte Carlo (MCMC) methods are not well suited for large scale datasets. One might also consider Laplace approximation for the approximate posterior distribution however we will stick with variational inference as it allows a richer set of approximations compared to Laplace approximation. Laplace approximation simply amounts to finding the Maximum A Posteriori (MAP) estimate to an augmented likelihood optimization, taking the negative of the inverse of the Hessian at the MAP estimate to estimate the variance-covariance matrix and finally use the variance-covariance matrix with a multivariate Gaussian distribution or some other appropriate multivariate distribution.
#
# * Assume that our approximate posterior distribution, which is also our probabilistic encoder, is given as:
#
# \begin{align*}
# q(Z|X)
# \end{align*}
#
# * Our probabilistic decoder is given by:
#
# \begin{align*}
# p(X|Z)
# \end{align*}
#
# * Given our setup above with regards to an encoder and a decoder let us now write down the optimization problem where $\theta$ are the generative model parameters while $\phi$ are the variational parameters:
#
# \begin{align*}
# \log{p(X)}= \underbrace{D_{KL}(q(Z|X)||p(Z|X))}_\text{Intractable as p(Z|X) is intractable} + \underbrace{\mathcal{L}(\theta, \phi|X)}_\text{Evidence Lower Bound or ELBO}
# \end{align*}
#
# * Note that $D_{KL}(q(Z|X)||p(Z|X))$ is non-negative therefore that makes the ELBO a lower bound on $\log{p(X)}$:
#
# \begin{align*}
# \log{p(X)}\geq \mathcal{L}(\theta, \phi|X) \quad \text{as} \quad D_{KL}(q(Z|X)||p(Z|X)) \geq 0
# \end{align*}
#
# * Therefore we can alter our optimization problem to look only at the ELBO:
#
# \begin{align*}
# \mathcal{L}(\theta, \phi|X) &= \mathbb{E}_{q(Z|X)}\left[\log{p(X,Z)} - \log{q(Z|X)}\right] \\
# &= \mathbb{E}_{q(Z|X)}\left[\underbrace{\log{p(X|Z)}}_\text{Reconstruction error} + \log{p(Z)} - \log{q(Z|X)}\right] \\
# &= \mathbb{E}_{q(Z|X)}\left[\underbrace{\log{p(X|Z)}}_\text{Reconstruction error} - \underbrace{D_{KL}(q(Z|X)||p(Z))}_\text{Regularization}\right] \\
# &= \int_{Z} \left[\log{p(X|Z)} - D_{KL}(q(Z|X)||p(Z))\right] q(Z|X) dZ
# \end{align*}
#
# * The above integration problem can be solved via Monte Carlo integration as $D_{KL}(q(Z|X)||p(Z))$ is not intractable. Assuming that the probabilistic encoder $q(Z|X)$ is a multivariate Gaussian with a diagonal variance-covariance matrix we use the reparameterization trick to sample from this distribution say $M$ times in order to calculate the expectation term in the ELBO optimization problem. The reparameterization trick in this particular case amounts to sampling $M$ times from the standard Gaussian distribution, multiplying the samples by $\sigma$ and adding $\mu$ to the samples.
#
# * $\mu$ is our learnt representation used for the reconstruction of the raw data. If the learnt representation is useful it can then be used for other tasks as well.
#
# * This is a powerful manner of combining Bayesian inference with deep learning. Variational inference used in this manner can be applied to various deep learning architectures and has further links with the Generative Adversarial Network (GAN). We explore the use of adversarial learning in representation learning in another repo/paper.
#
# ### Results
#
# The accuracy score for the MNIST classification task with a variational autoencoder: 96.520000%.
# +
encoding_dim = 500
variational_autoencoder = VariationalAutoencoder(n_feat=X_train.shape[1],
n_epoch=50,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=1000,
encoding_dim=encoding_dim,
denoising=None)
print(variational_autoencoder.autoencoder.summary())
pipe_variational_autoencoder = Pipeline(steps=[("autoencoder", variational_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_variational_autoencoder = pipe_variational_autoencoder.fit(X_train, y_train)
acc_variational_autoencoder = pipe_variational_autoencoder.score(X_test, y_test)
print("The accuracy score for the MNIST classification task with a variational autoencoder: %.6f%%." % (acc_variational_autoencoder * 100))
if encoding_dim == 2:
test_encoded_df = pd.DataFrame(pipe_variational_autoencoder.named_steps["autoencoder"].encoder.predict(X_test))
test_encoded_df["Target"] = y_test
test_encoded_df.columns.values[0:2] = ["Encoding_1", "Encoding_2"]
scaler_plot = MinMaxScaler(feature_range=(0.25, 0.75))
scaler_plot = scaler_plot.fit(test_encoded_df[["Encoding_1", "Encoding_2"]])
test_encoded_df[["Encoding_1", "Encoding_2"]] = scaler_plot.transform(test_encoded_df[["Encoding_1", "Encoding_2"]])
cluster_plot = ggplot(test_encoded_df) + \
geom_point(aes(x="Encoding_1",
y="Encoding_2",
fill="factor(Target)"),
size=1,
color = "black") + \
xlab("Encoding dimension 1") + \
ylab("Encoding dimension 2") + \
ggtitle("Variational autoencoder with 2-dimensional encoding") + \
theme_matplotlib()
print(cluster_plot)
n = 30
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
grid_x = norm.ppf(np.linspace(0.05, 0.95, n))
grid_y = norm.ppf(np.linspace(0.05, 0.95, n))
for i, xi in enumerate(grid_x):
for j, yi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
x_decoded = pipe_variational_autoencoder.named_steps["autoencoder"].generator.predict(z_sample)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size, j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(20, 20))
plt.imshow(figure, cmap="Greys_r")
plt.title("Variational Autoencoder (VAE) with 2-dimensional encoding\nGenerating new images")
plt.xlabel("Encoding dimension 1")
plt.ylabel("Encoding dimension 2")
plt.savefig(fname="VAE_Generated_Images.png")
plt.show()
# -
# ## MNIST: 2 Dimensional Convolutional Autoencoders
#
# For 2 dimensional convolution filters the idea is similar as for the 1 dimensional convolution filters. We will stick to our previously mentioned banking example to illustrate this point.
#
# \begin{align*}
# x =
# \begin{array}
# {l}
# \text{Period 1} \\ \text{Period 2} \\ \text{Period 3} \\ \text{Period 4} \\ \text{Period 5}
# \end{array}
# \left[
# \begin{array}
# {ccc}
# \$0 & \$0 & \$0 \\
# \$0 & \$200 & \$0 \\
# \$100 & \$0 & \$0 \\
# \$0 & \$0 & \$300 \\
# \$0 & \$0 & \$0
# \end{array}
# \right]
# \end{align*}
#
# In the 2 dimensional tensor of raw transactions data now we have 5 historical time periods, i.e., the rows, and 3 different transaction types, i.e., the columns. We will use a kernel, $\alpha \in \mathbb{R}^{2\times3}$, to extract useful features from the raw data. The choice of such a kernel means that we are interested in finding a feature map across all 3 transaction types and 2 historical time periods. We will use a stride length of 1 and a valid convolution to extract features over different patches of the raw data. The following will illustrate this point where $x_{\text{patch}} \subset x$:
#
# \begin{align*}
# \alpha &=
# \left[
# \begin{array}
# {ccc}
# \alpha_{1,1} & \alpha_{1,2} & \alpha_{1,3} \\
# \alpha_{2,1} & \alpha_{2,2} & \alpha_{2,3}
# \end{array}
# \right] \\
# x_{\text{patch}} &=
# \left[
# \begin{array}
# {ccc}
# \$0 & \$0 & \$0 \\
# \$0 & \$200 & \$0
# \end{array}
# \right] \\
# \mathbf{C}(x=x_{\text{patch}}|\alpha) &= x * \alpha \\
# &= \sum_{t=1}^{2} \sum_{k=1}^{3} x_{t,k} \alpha_{t,k}
# \end{align*}
#
# The principles and ideas apply to 2 dimensional convolution filters as they do for their 1 dimensional counterparts there we will not repeat them here.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 28 \times 28} \\
# &E: \mathbb{R}^{N \times 28 \times 28} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 28 \times 28}
# \end{align*}
#
# ### Results
#
# The accuracy score for the MNIST classification task with a 2 dimensional convolutional autoencoder: 98.860000%.
# +
convolutional2D_autoencoder = Convolutional2DAutoencoder(input_shape=(int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5)), 1),
n_epoch=5,
batch_size=100,
encoder_layers=3,
decoder_layers=3,
filters=100,
kernel_size=(8, 8),
strides=(1, 1),
pool_size=(4, 4),
denoising=None)
print(convolutional2D_autoencoder.autoencoder.summary())
pipe_convolutional2D_autoencoder = Pipeline(steps=[("autoencoder", convolutional2D_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_convolutional2D_autoencoder = pipe_convolutional2D_autoencoder.fit(np.reshape(X_train, [X_train.shape[0], int(math.pow(X_train.shape[1], 0.5)), int(math.pow(X_train.shape[1], 0.5)), 1]),
y_train)
acc_convolutional2D_autoencoder = pipe_convolutional2D_autoencoder.score(np.reshape(X_test, [X_test.shape[0], int(math.pow(X_test.shape[1], 0.5)), int(math.pow(X_test.shape[1], 0.5)), 1]), y_test)
print("The accuracy score for the MNIST classification task with a 2 dimensional convolutional autoencoder: %.6f%%." % (acc_convolutional2D_autoencoder * 100))
# -
# ## Insurance: No Autoencoders
#
# We now proceed to run the insurance model without any handcrafted or deep learning based feature engineering.
#
# ### Results
#
# The AUROC score for the insurance classification task without autoencoders: 92.206261%.
# +
claim_risk = pd.read_csv(filepath_or_buffer="../R/data/claim_risk.csv")
claim_risk.drop(columns="policy.id", axis=1, inplace=True)
claim_risk = np.asarray(claim_risk).ravel()
transactions = pd.read_csv(filepath_or_buffer="../R/data/transactions.csv")
transactions.drop(columns="policy.id", axis=1, inplace=True)
n_policies = 1000
n_transaction_types = 3
n_time_periods = 4
transactions = np.reshape(np.asarray(transactions), (n_policies, n_time_periods * n_transaction_types))
X_train, X_test, y_train, y_test = train_test_split(transactions, claim_risk, test_size=0.3, random_state=666)
min_X_train = np.apply_along_axis(func1d=np.min, axis=0, arr=X_train)
max_X_train = np.apply_along_axis(func1d=np.max, axis=0, arr=X_train)
range_X_train = max_X_train - min_X_train + sys.float_info.epsilon
X_train = (X_train - min_X_train) / range_X_train
X_test = (X_test - min_X_train) / range_X_train
transactions = (transactions - min_X_train) / range_X_train
pipe_base = Pipeline(steps=[("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_base = pipe_base.fit(X_train, y_train)
auroc_base = roc_auc_score(y_true=y_test,
y_score=pipe_base.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task without autoencoders: %.6f%%." % (auroc_base * 100))
# -
# ## Insurance: PCA
#
# We now proceed to run the insurance model without any handcrafted or deep learning based feature engineering however with a PCA filter that picks the number of components that explain $99\%$ of the variation.
#
# ### Results
#
# The AUROC score for the insurance classification task with PCA: 91.128859%.
# +
pipe_pca = Pipeline(steps=[("PCA", PCA(n_components=0.99)),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_pca = pipe_pca.fit(X_train, y_train)
auroc_pca = roc_auc_score(y_true=y_test,
y_score=pipe_pca.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with PCA: %.6f%%." % (auroc_pca * 100))
# -
# ## Insurance: Handcrafted Features
#
# In this case we have created some handcrafted features which we believe provide a useful representation of the raw data for the insurance model.
#
# ### Results
#
# The AUROC score for the insurance classification task with handcrafted features: 93.610635%.
# +
claim_risk = pd.read_csv(filepath_or_buffer="../R/data/claim_risk.csv")
claim_risk.drop(columns="policy.id", axis=1, inplace=True)
claim_risk = np.asarray(claim_risk).ravel()
handcrafted_features = pd.read_csv(filepath_or_buffer="../R/data/handcrafted_features.csv")
handcrafted_features = np.asarray(handcrafted_features)
n_policies = 1000
n_feat = 12
X_train, X_test, y_train, y_test = train_test_split(handcrafted_features, claim_risk, test_size=0.3, random_state=666)
min_X_train = np.apply_along_axis(func1d=np.min, axis=0, arr=X_train)
max_X_train = np.apply_along_axis(func1d=np.max, axis=0, arr=X_train)
range_X_train = max_X_train - min_X_train + sys.float_info.epsilon
X_train = (X_train - min_X_train) / range_X_train
X_test = (X_test - min_X_train) / range_X_train
pipe_hcfe = Pipeline(steps=[("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_hcfe = pipe_hcfe.fit(X_train, y_train)
auroc_hcfe = roc_auc_score(y_true=y_test,
y_score=pipe_hcfe.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with handcrafted features: %.6f%%." % (auroc_hcfe * 100))
# -
# ## Insurance: Handcrafted Features and PCA
#
# In this case we have created some handcrafted features which we believe provide a useful representation of the raw data for the insurance model. We also use a PCA filter.
#
# ### Results
#
# The AUROC score for the insurance classification task with handcrafted features and PCA: 93.160377%.
# +
pipe_hcfe_pca = Pipeline(steps=[("PCA", PCA(n_components=0.99)),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_hcfe_pca = pipe_hcfe_pca.fit(X_train, y_train)
auroc_hcfe_pca = roc_auc_score(y_true=y_test,
y_score=pipe_hcfe_pca.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with handcrafted features and PCA: %.6f%%." % (auroc_hcfe_pca * 100))
# -
# ## Insurance: Vanilla Autoencoders
#
# In this case we use vanilla autoencoders to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 12} \\
# &E: \mathbb{R}^{N \times 12} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 12}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with an autoencoder: 93.932247%.
# +
autoencoder = VanillaAutoencoder(n_feat=X_train.shape[1],
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=100,
encoding_dim=50,
denoising=None)
print(autoencoder.autoencoder.summary())
pipe_autoencoder = Pipeline(steps=[("autoencoder", autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_autoencoder = pipe_autoencoder.fit(X_train, y_train)
auroc_autoencoder = roc_auc_score(y_true=y_test,
y_score=pipe_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with an autoencoder: %.6f%%." % (auroc_autoencoder * 100))
# -
# ## Insurance: Denoising Autoencoders
#
# In this case we use denoising autoencoders to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 12} \\
# &E: \mathbb{R}^{N \times 12} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 12}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with a denoising autoencoder: 93.712479%.
# +
noise = 0.10 * np.reshape(np.random.uniform(low=0.0,
high=1.0,
size=X_train.shape[0] * X_train.shape[1]),
[X_train.shape[0], X_train.shape[1]])
denoising_autoencoder = VanillaAutoencoder(n_feat=X_train.shape[1],
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=100,
encoding_dim=50,
denoising=noise)
print(denoising_autoencoder.autoencoder.summary())
pipe_denoising_autoencoder = Pipeline(steps=[("autoencoder", denoising_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_denoising_autoencoder = pipe_denoising_autoencoder.fit(X_train, y_train)
auroc_denoising_autoencoder = roc_auc_score(y_true=y_test,
y_score=pipe_denoising_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with a denoising autoencoder: %.6f%%." % (auroc_denoising_autoencoder * 100))
# -
# ## Insurance: Sequence to Sequence Autoencoders
#
# In this case we use sequence to sequence autoencoders, taking into account the time series nature, i.e., sequential nature, of the raw transactions data, to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 4 \times 3} \\
# &E: \mathbb{R}^{N \times 4 \times 3} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 4 \times 3}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with a sequence to sequence autoencoder: 91.418310%.
# +
transactions = np.reshape(np.asarray(transactions), (n_policies, n_time_periods, n_transaction_types))
X_train, X_test, y_train, y_test = train_test_split(transactions, claim_risk, test_size=0.3, random_state=666)
seq2seq_autoencoder = Seq2SeqAutoencoder(input_shape=(X_train.shape[1], X_train.shape[2]),
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=100,
encoding_dim=50,
stateful=False,
denoising=None)
print(seq2seq_autoencoder.autoencoder.summary())
pipe_seq2seq_autoencoder = Pipeline(steps=[("autoencoder", seq2seq_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_seq2seq_autoencoder = pipe_seq2seq_autoencoder.fit(X_train, y_train)
auroc_seq2seq_autoencoder = roc_auc_score(y_test,
pipe_seq2seq_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with a sequence to sequence autoencoder: %.6f%%." % (auroc_seq2seq_autoencoder * 100))
# -
# ## Insurance: 1 Dimensional Convolutional Autoencoders
#
# In this case we use 1 dimensional convolutional autoencoders to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 4 \times 3} \\
# &E: \mathbb{R}^{N \times 4 \times 3} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 4 \times 3}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with a 1 dimensional convolutional autoencoder: 91.509434%.
# +
transactions = np.reshape(np.asarray(transactions), (n_policies, n_time_periods, n_transaction_types))
X_train, X_test, y_train, y_test = train_test_split(transactions, claim_risk, test_size=0.3, random_state=666)
convolutional_autoencoder = ConvolutionalAutoencoder(input_shape=(X_train.shape[1], X_train.shape[2]),
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
filters=50,
kernel_size=2,
strides=1,
pool_size=2,
denoising=None)
print(convolutional_autoencoder.autoencoder.summary())
pipe_convolutional_autoencoder = Pipeline(steps=[("autoencoder", convolutional_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_convolutional_autoencoder = pipe_convolutional_autoencoder.fit(X_train, y_train)
auroc_convolutional_autoencoder = roc_auc_score(y_test,
pipe_convolutional_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with a 1 dimensional convolutional autoencoder: %.6f%%." % (auroc_convolutional_autoencoder * 100))
# -
# ## Insurance: 2 Dimensional Convolutional Autoencoders
#
# In this case we use 2 dimensional convolutional autoencoders to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 4 \times 3} \\
# &E: \mathbb{R}^{N \times 4 \times 3} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 4 \times 3}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with a 2 dimensional convolutional autoencoder: 92.645798%.
# +
transactions = np.reshape(np.asarray(transactions), (n_policies, n_time_periods, n_transaction_types, 1))
X_train, X_test, y_train, y_test = train_test_split(transactions, claim_risk, test_size=0.3, random_state=666)
convolutional2D_autoencoder = Convolutional2DAutoencoder(input_shape=(X_train.shape[1], X_train.shape[2], 1),
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
filters=50,
kernel_size=(2, 3),
strides=(1, 1),
pool_size=(2, 1),
denoising=None)
print(convolutional2D_autoencoder.autoencoder.summary())
pipe_convolutional2D_autoencoder = Pipeline(steps=[("autoencoder", convolutional2D_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_convolutional2D_autoencoder = pipe_convolutional2D_autoencoder.fit(X_train, y_train)
auroc_convolutional2D_autoencoder = roc_auc_score(y_test,
pipe_convolutional2D_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with a 2 dimensional convolutional autoencoder: %.6f%%." % (auroc_convolutional2D_autoencoder * 100))
# -
# ## Insurance: Variational Autoencoders
#
# In this case we use variational autoencoders to learn a good representation of the raw data such that we can obtain an uplift, primarily in terms of AUROC, for the supervised learning task.
#
# \begin{align*}
# &X \in \mathbb{R}^{N \times 12} \\
# &E: \mathbb{R}^{N \times 12} \rightarrow \mathbb{R}^{N \times K} \\
# &D: \mathbb{R}^{N \times K} \rightarrow \mathbb{R}^{N \times 12}
# \end{align*}
#
# ### Results
#
# The AUROC score for the insurance classification task with a variational autoencoder: 90.871569%.
# +
transactions = np.reshape(np.asarray(transactions), (n_policies, n_time_periods * n_transaction_types))
X_train, X_test, y_train, y_test = train_test_split(transactions, claim_risk, test_size=0.3, random_state=666)
variational_autoencoder = VariationalAutoencoder(n_feat=X_train.shape[1],
n_epoch=100,
batch_size=50,
encoder_layers=3,
decoder_layers=3,
n_hidden_units=100,
encoding_dim=50,
denoising=None)
print(variational_autoencoder.autoencoder.summary())
pipe_variational_autoencoder = Pipeline(steps=[("autoencoder", variational_autoencoder),
("scaler_classifier", scaler_classifier),
("classifier", logistic)])
pipe_variational_autoencoder = pipe_variational_autoencoder.fit(X_train, y_train)
auroc_variational_autoencoder = roc_auc_score(y_true=y_test,
y_score=pipe_variational_autoencoder.predict_proba(X_test)[:, 1],
average="weighted")
print("The AUROC score for the insurance classification task with a variational autoencoder: %.6f%%." % (auroc_variational_autoencoder * 100))
# -
# ## Conclusion
#
# We have shown how to use deep learning and Bayesian inference to learn a good representation of raw data $X$, i.e., 1 or 2 or perhaps more dimensional tensors per unit of analysis, that can then perhaps be used for supervised learning tasks in the domain of computer vision and insurance. This moves us away from manual handcrafted feature engineering towards automatic feature engineering, i.e., representation learning. This does introduce architecture engineering however that can be automated as well perhaps by the use of genetic algorithms or reinforcement learning - a topic for another paper perhaps.
#
# Finally, I would like to emphasize that the same code used for solving the computer vision task was used to solve the insurance task: In both tasks automatic feature engineering via deep learning had the best performance despite the fact that we were not explicitly looking for the best state of the art architecture possible.
# ## References
#
# 1. <NAME>., <NAME>. and <NAME>. (2016). Deep Learning (MIT Press).
# 2. <NAME>. (2017). Hands-On Machine Learning with Scikit-Learn & Tensorflow (O'Reilly).
# 3. <NAME>., and <NAME>. (2014). Auto-Encoding Variational Bayes (https://arxiv.org/abs/1312.6114).
# 4. http://scikit-learn.org/stable/#
# 5. https://towardsdatascience.com/learning-rate-schedules-and-adaptive-learning-rate-methods-for-deep-learning-2c8f433990d1
# 6. https://stackoverflow.com/questions/42177658/how-to-switch-backend-with-keras-from-tensorflow-to-theano
# 7. https://blog.keras.io/building-autoencoders-in-keras.html
# 8. https://keras.io
# 9. https://www.cs.cornell.edu/courses/cs1114/2013sp/sections/S06_convolution.pdf
# 10. http://deeplearning.net/tutorial/lstm.html
# 11. <NAME>. (2018). Deep Learning with Python (Manning).
| Python/autoencoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo Notebook For Concentric Transmon Qubit
# Start by importing QisKit Metal
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict #, open_docs
# Load the QisKit Metal graphic interface:
design = designs.DesignPlanar()
gui = MetalGUI(design)
# Now we'll load the concentric transmon component:
from qiskit_metal.qlibrary.qubits.transmon_concentric import TransmonConcentric
design.overwrite_enabled = True
q1 = TransmonConcentric(design, 'qubit1')
gui.rebuild()
gui.autoscale()
| tutorials/Appendix/Quick Topic Tutorials Notebooks/Concentric Transmon Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
import math
import panel as pn
pn.extension()
# The ``RangeSlider`` widget allows selecting a floating-point range using a slider with two handles.
#
# For more information about listening to widget events and laying out widgets refer to the [widgets user guide](../../user_guide/Widgets.ipynb). Alternatively you can learn how to build GUIs by declaring parameters independently of any specific widgets in the [param user guide](../../user_guide/Param.ipynb). To express interactivity entirely using Javascript without the need for a Python server take a look at the [links user guide](../../user_guide/Param.ipynb).
#
# #### Parameters:
#
# For layout and styling related parameters see the [customization user guide](../../user_guide/Customization.ipynb).
#
# ##### Core
#
# * **``start``** (float): The range's lower bound
# * **``end``** (float): The range's upper bound
# * **``step``** (float): The interval between values
# * **``value``** (tuple): Tuple of upper and lower bounds of selected range
# * **``value_throttled``** (tuple): Tuple of upper and lower bounds of selected range where events are throttled by `callback_throttle` value.
#
# ##### Display
#
# * **``bar_color``** (color): Color of the slider bar as a hexadecimal RGB value
# * **``callback_policy``** (str, **DEPRECATED**): Policy to determine when slider events are triggered (one of 'continuous', 'throttle', 'mouseup')
# * **``callback_throttle``** (int): Number of milliseconds to pause between callback calls as the slider is moved
# * **``direction``** (str): Whether the slider should go from left to right ('ltr') or right to left ('rtl')
# * **``disabled``** (boolean): Whether the widget is editable
# * **``name``** (str): The title of the widget
# * **``orientation``** (str): Whether the slider should be displayed in a 'horizontal' or 'vertical' orientation.
# * **``tooltips``** (boolean): Whether to display tooltips on the slider handle
#
# ___
# +
range_slider = pn.widgets.RangeSlider(
name='Range Slider', start=0, end=math.pi, value=(math.pi/4., math.pi/2.), step=0.01)
range_slider
# -
# ``RangeSlider.value`` returns a tuple of float values that can be read out and set like other widgets:
range_slider.value
| examples/reference/widgets/RangeSlider.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Heterogeneous graph attention networks
#
# By <NAME> 15.10.2020
# +
from enum import IntEnum
from math import comb
import numpy as np
import pandas as pd
import networkx as nx
import torch
import torch_geometric as pyg
# -
# ## Download and load data
#
# In this notebook, we'll use two small data sets. One will be the same as in the previous notebook -- interactions between FDA-approved drugs (ChCh-Miner), while the other will be the interactions between the drugs and their corresponding targets (ChG-Miner). Both of these data sets can be obtained from the _Stanford Biomedical Network Dataset Collection_.
# +
# !mkdir -p data
# !wget http://snap.stanford.edu/biodata/datasets/10001/files/ChCh-Miner_durgbank-chem-chem.tsv.gz -O data/ChCh-Miner_durgbank-chem-chem.tsv.gz
# !wget http://snap.stanford.edu/biodata/datasets/10002/files/ChG-Miner_miner-chem-gene.tsv.gz -O data/ChG-Miner_miner-chem-gene.tsv.gz
# !yes | gunzip data/ChCh-Miner_durgbank-chem-chem.tsv.gz
# !yes | gunzip data/ChG-Miner_miner-chem-gene.tsv.gz
# -
# !head data/ChCh-Miner_durgbank-chem-chem.tsv
# This time, we have two different node types: drugs and targets. The targets here are actual genes, so we assign each node and edge type its own id, so we can more easily identify them later on.
# +
class NodeType(IntEnum):
DRUG = 1
GENE = 2
class EdgeType(IntEnum):
DRUG_DRUG = 1
DRUG_GENE = 2
GENE_DRUG = 3
GENE_GENE = 4
@classmethod
def from_nodes(cls, n1, n2):
return {
(NodeType.DRUG, NodeType.DRUG): cls.DRUG_DRUG,
(NodeType.DRUG, NodeType.GENE): cls.DRUG_GENE,
(NodeType.GENE, NodeType.DRUG): cls.GENE_DRUG,
(NodeType.GENE, NodeType.GENE): cls.GENE_GENE,
}[int(n1), int(n2)]
# +
f1_drugs = set()
entity_dict, edge_index = {}, []
entity_count = 0 # we will use this to assign numbers to drugs
entity_type = []
with open("data/ChCh-Miner_durgbank-chem-chem.tsv") as f:
for line in f:
line = line.rstrip()
d1, d2 = line.split("\t")
if d1 not in entity_dict:
entity_dict[d1] = entity_count
entity_type.append(NodeType.DRUG)
entity_count += 1
if d2 not in entity_dict:
entity_dict[d2] = entity_count
entity_type.append(NodeType.DRUG)
entity_count += 1
edge_index.append([entity_dict[d1], entity_dict[d2]])
f1_drugs.add(d1)
f1_drugs.add(d2)
print(f"{len(edge_index):,} interactions")
# -
# !head data/ChG-Miner_miner-chem-gene.tsv
# +
f2_drugs = set()
with open("data/ChG-Miner_miner-chem-gene.tsv") as f:
for line in f:
line = line.rstrip()
d1, d2 = line.split("\t")
if d1 not in entity_dict:
entity_dict[d1] = entity_count
entity_type.append(NodeType.DRUG)
entity_count += 1
if d2 not in entity_dict:
entity_dict[d2] = entity_count
entity_type.append(NodeType.GENE)
entity_count += 1
edge_index.append([entity_dict[d1], entity_dict[d2]])
f2_drugs.add(d1)
print(f"{len(edge_index):,} interactions")
# -
# Let's verify that at least some of the drugs overlap between the two datasets.
len(f1_drugs & f2_drugs)
entity_type = torch.LongTensor(entity_type)
pd.Series.value_counts(entity_type.numpy())
edge_index = torch.LongTensor(edge_index).T
edge_index
# ## Prepare graph
# ### Examine graph
print("Sparsity: %.2f" % (edge_index.size(1) / comb(len(edge_index[0].unique()), 2)))
pyg.utils.is_undirected(edge_index)
# Even though the graph is undirected, we're going to cast it to a directed graph, so this will be one less piece of complexity we will have to worry about.
edge_index = pyg.utils.to_undirected(edge_index)
num_nodes = entity_count
print(f"{num_nodes:,} nodes")
# ### Extract LCC
#
# Many graph-based learning techniques cannot operate on disconnected graphs, so it is standard practice to operate on the largest connected component (LCC). The easiest way to exctract the LCC is to use _networkx_, a popular pure Python library for network analysis.
data = pyg.data.Data(edge_index=edge_index, node_type=entity_type)
nx_data = pyg.utils.to_networkx(data, to_undirected=True, node_attrs=["node_type"])
nx_data
connected_components = list(nx.connected_components(nx_data))
print(f"Graph contains {len(connected_components)} connected components")
lcc_nodes = max(connected_components, key=len)
lcc = nx_data.subgraph(lcc_nodes)
print(f"{lcc.number_of_nodes():,} nodes in LCC")
pyg.utils.from_networkx(lcc)
lcc_data = pyg.utils.from_networkx(lcc)
edge_index = lcc_data.edge_index
node_type = lcc_data.node_type
print(f"{len(edge_index.unique()):,} nodes")
# ### Determine edge types
#
# The edge types here are important, as they each carry a different semantic meaning. Interactions between drugs are not semantically equivalent to interactions between drugs and their targets, so we make sure to differentiate between them.
edge_type = []
for n1, n2 in edge_index.T:
edge_type.append(EdgeType.from_nodes(node_type[n1], node_type[n2]))
pd.Series.value_counts(edge_type)
# Notice that there is the same number of edges of types 2 and 3. This is because one of these corresponds to drug-gene and the other to gene-drug edges. Because we cast our graph to an undirected graph, this makes perfect sense.
edge_type = torch.LongTensor(edge_type)
# ## Prepare data set
#
# As in any machine learning task, we split our data set into a training and validation set. In _torch_geometric_, it is easiest to define boolean masks to indicate which edges belong into each set. Why not use integer indices? Because we can easily attach boolean masks to a _torch_geometric_ `pyg.data.Data` object, which will be indexed properly during mini-batching. Indexing the masks is only possible if their shape matches that of the `edge_index`, so a boolean mask is the way to go.
from sklearn.model_selection import train_test_split
indices = torch.arange(edge_index.shape[1])
train_idx, val_idx = train_test_split(indices, test_size=0.01)
train_mask, val_mask = torch.zeros_like(indices).bool(), torch.zeros_like(indices).bool()
train_mask[train_idx] = 1
val_mask[val_idx] = 1
train_mask, val_mask
# Because our nodes have no additional features, we generally use a one-hot encoding as features to our model. Obviously, creating a dense identity matrix is very memory-inefficient, and we will quickly run out of memory when working with larger graphs, and unfortunately, `torch_geometric` doesn't yet fully support sparse matrices. So, when dealing with larger graphs, we often use random features instead.
data = pyg.data.Data(edge_index=edge_index, edge_type=edge_type, train_mask=train_mask, val_mask=val_mask)
data.x = torch.randn(data.num_nodes, 128)
data
# ## Metapaths
#
# Our task in this notebook will be the same as in the `graph-attention` notebook -- predicting drug-drug interactions. However, this time, we have more information avalable. There may be several different ways as to determine how drugs interact with each other. We can encode these *different ways* into meta-paths: these are relations which we suspect might be useful for the prediction task. We'll use three meta-paths in this particular example: drug-drug, drug-drug-drug, and drug-gene-drug.
#
# Each one of these metapaths defines its own adjacency matrix. Because we are working with a relatively small data set, it is easy to obtain the meta-path adjacency matrix through simple matrix multiplication.
import torch_sparse
metapaths = [
(NodeType.DRUG, NodeType.DRUG),
(NodeType.DRUG, NodeType.DRUG, NodeType.DRUG),
(NodeType.DRUG, NodeType.GENE, NodeType.DRUG),
]
unique_edge_types = edge_type.unique()
unique_edge_types
adjs = {}
for et in unique_edge_types:
mask = edge_type == et
row, col = edge_index[:, mask]
adj = torch_sparse.SparseTensor(row=row, col=col, sparse_sizes=(num_nodes, num_nodes))
adjs[int(et)] = adj
adjs
mp_adjs = []
for metapath in metapaths:
mp_adj = adjs[metapath[0]]
for idx in range(len(metapath) - 1):
mp_adj @= adjs[EdgeType.from_nodes(metapath[idx], metapath[idx + 1])]
mp_adjs.append(mp_adj)
mp_adjs
mp_adjs = [torch.stack([mp_adj.storage.row(), mp_adj.storage.col()]) for mp_adj in mp_adjs]
mp_adjs
# ## Heterogeneous attention mechanism
#
# Our graph attention network that we saw in the previous notebook has no notion of metapaths or different edge types. We'll now take a look at heterogeneous graph attention networks (HAN), which are able to account for different edge types. The HAN builds on the GAT convolutional layers, so we'll copy our implementation from the previous notebook here.
# +
import torch.nn.functional as F
from torch_geometric.nn.inits import glorot, zeros
class GATConv(pyg.nn.MessagePassing):
def __init__(self, in_features, out_features, k_heads=1, concat=True):
super().__init__(aggr="add", node_dim=0)
self.in_features = in_features
self.out_features = out_features
self.k_heads = k_heads
self.concat = concat
self.weight = torch.nn.Parameter(torch.Tensor(in_features, out_features * k_heads))
if concat:
self.bias = torch.nn.Parameter(torch.Tensor(k_heads * out_features))
else:
self.bias = torch.nn.Parameter(torch.Tensor(1, out_features))
self.att_l = torch.nn.Parameter(torch.Tensor(1, k_heads, out_features))
self.att_r = torch.nn.Parameter(torch.Tensor(1, k_heads, out_features))
self.reset_parameters()
def reset_parameters(self):
glorot(self.weight)
glorot(self.att_l)
glorot(self.att_r)
zeros(self.bias)
def forward(self, x, edge_index):
x_lin = (x @ self.weight).view(-1, self.k_heads, self.out_features)
alpha_l = torch.sum(x_lin * self.att_l, dim=2)
alpha_r = torch.sum(x_lin * self.att_r, dim=2)
out = self.propagate(edge_index, x=x_lin, alpha=(alpha_l, alpha_r))
if self.concat:
out = out.view(-1, self.k_heads * self.out_features)
else:
out = out.mean(dim=1)
out += self.bias
return out
def message(self, x_j, alpha_i, alpha_j, index):
alpha = alpha_i + alpha_j
alpha = F.leaky_relu(alpha, 0.2)
alpha = pyg.utils.softmax(alpha, index)
return alpha.unsqueeze(-1) * x_j
# -
# ### Semantic-level attention
# HAN's function in the following way: Each metapath specifies its own adjacency matrix, so the first step of the HAN is to apply GAT convolutions to every metapath adjacency matrix. If we have $r$ metapaths, we then obtain $r$ new representations for each node, as calculated the the GAT. The second step of a HAN convolution is to apply attention to these latent representations. Intuitively, we determine how important each metapath representation is to the overall node representation, and apply attention-weighted sums to them. This way, we aggregate the $r$ representations into a single one. The attention mechanism that we use is identical to the one in the previous notebook, so we won't delve into the implementation details.
gat_convs = [GATConv(data.x.shape[1], 8, k_heads=3) for _ in unique_edge_types]
embeddings = [conv(data.x, adj) for conv, adj in zip(gat_convs, mp_adjs)]
embeddings = torch.stack(embeddings, dim=1)
embeddings.shape
# Apply non-linearity
embeddings = F.relu(embeddings)
embeddings.shape
att_in_features = embeddings.shape[2] # 24
att_out_features = 8
weight = torch.Tensor(torch.rand(att_in_features, att_out_features))
bias = torch.Tensor(torch.rand(1, att_out_features))
torch.sum(weight * embeddings.unsqueeze(-1), dim=2).shape
w = torch.sum(weight * embeddings.unsqueeze(-1), dim=2) + bias
w = torch.tanh(w)
w.shape
q = torch.Tensor(torch.rand(1, 1, att_out_features))
beta = torch.sum(q * w, dim=-1) / num_nodes
beta = torch.softmax(beta, dim=1)
beta.shape
# We now have metapath attention coefficients for every for every node. Now, the only thing that remains is to multiply the node-level features by these coefficients, and sum them up.
z = torch.sum(embeddings * beta.unsqueeze(-1), dim=1)
z.shape
z
# ## Putting it into torch_geometric
#
# Again, it is easy to convert the code above into a reusable torch_geometric layer, which we will use from now on.
# +
class HANConv(torch.nn.Module):
def __init__(self, in_channels, num_relations, out_channels, sem_att_channels, node_heads=3):
super().__init__()
self.in_channels = in_channels
self.num_relations = num_relations
self.out_channels = out_channels
self.sem_att_channels = sem_att_channels
self.node_heads = node_heads
self.node_gats = torch.nn.ModuleList()
for _ in range(num_relations):
self.node_gats.append(GATConv(in_channels, out_channels, node_heads))
self.W = torch.nn.Parameter(torch.Tensor(1, 1, out_channels * node_heads, sem_att_channels))
self.b = torch.nn.Parameter(torch.Tensor(1, 1, sem_att_channels))
self.q = torch.nn.Parameter(torch.Tensor(1, 1, sem_att_channels))
self.reset_parameters()
def reset_parameters(self):
glorot(self.W)
zeros(self.b)
glorot(self.q)
def forward(self, x, metapaths):
# Calculate node-level attention representations
out = [gat(x, metapath) for gat, metapath in zip(self.node_gats, metapaths)]
out = torch.stack(out, dim=1).to(x.device)
# Apply non-linearity
out = F.relu(out)
# Aggregate node-leve representation using semantic level attention
w = torch.sum(self.W * out.unsqueeze(-1), dim=-2) + self.b
w = torch.tanh(w)
beta = torch.sum(self.q * w, dim=-1)
beta = torch.softmax(beta, dim=1)
z = torch.sum(out * beta.unsqueeze(-1), dim=1)
return z
conv = HANConv(data.x.shape[1], len(unique_edge_types), 8, 8, 3)
conv(data.x, mp_adjs).shape
# -
# ## Learning
#
# Having implemented HAN convolutions, we can now build a full model, and fit it on our graph data.
from torch_geometric.utils import negative_sampling
data.val_neg_edge_index = negative_sampling(data.edge_index, num_neg_samples=data.val_mask.sum())
# ### Rebuild metapaths with only training edges
metaf1_drugsths = [
(NodeType.DRUG, NodeType.DRUG),
(NodeType.DRUG, NodeType.DRUG, NodeType.DRUG),
(NodeType.DRUG, NodeType.GENE, NodeType.DRUG),
(NodeType.GENE, NodeType.DRUG, NodeType.GENE),
]
edge_index_train = data.edge_index[:, data.train_mask]
edge_types_train = data.edge_type[data.train_mask]
edge_index_train
unique_edge_types = edge_types_train.unique()
unique_edge_types
adjs = {}
for et in unique_edge_types:
row, col = edge_index_train[:, edge_types_train == et]
adj = torch_sparse.SparseTensor(row=row, col=col, sparse_sizes=(num_nodes, num_nodes))
adjs[int(et)] = adj
adjs
mp_adjs = []
for metapath in metapaths:
mp_adj = adjs[metapath[0]]
for idx in range(len(metapath) - 1):
mp_adj @= adjs[EdgeType.from_nodes(metapath[idx], metapath[idx + 1])]
mp_adjs.append(mp_adj)
mp_adjs
mp_adjs = [torch.stack([mp_adj.storage.row(), mp_adj.storage.col()]) for mp_adj in mp_adjs]
mp_adjs
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
class Encoder(torch.nn.Module):
def __init__(self, in_dim, num_relations, hidden_dim, out_dim, node_heads=4):
super().__init__()
self.conv1 = HANConv(in_dim, num_relations, hidden_dim, sem_att_channels=8, node_heads=node_heads)
self.conv2 = HANConv(hidden_dim * node_heads, num_relations, out_dim, sem_att_channels=8, node_heads=node_heads)
def forward(self, x, metapaths):
x = F.relu(self.conv1(x, metapaths))
return self.conv2(x, metapaths)
for i in range(len(mp_adjs)):
mp_adjs[i] = mp_adjs[i].to(device)
from torch_geometric.nn import GAE
model = GAE(Encoder(data.num_features, len(mp_adjs), hidden_dim=8, out_dim=8, node_heads=8)).to(device)
model
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
# +
training_losses, validation_losses, val_auc, val_ap = [], [], [], []
for epoch in range(150):
model.train()
optimizer.zero_grad()
# As in the GAT notebook, we only care about predicting drug-drug edges
drug_drug_mask = data.edge_type == EdgeType.DRUG_DRUG
train_pos_edge_index = data.edge_index[:, data.train_mask & drug_drug_mask].to(device)
x = data.x.to(device)
# z holds the node embeddings
z = model.encode(x, mp_adjs)
# while the link prediction is performed inside this handy model.recon_loss function
loss = model.recon_loss(z, train_pos_edge_index)
training_loss = loss.item()
loss.backward()
optimizer.step()
model.eval()
# To validate our model, we first compute node embeddings for each node using the training
# set edges.
train_pos_edge_index = data.edge_index[:, data.train_mask & drug_drug_mask].to(device)
# Our model's predictions will then be checked against the true, existing validation edges
# stored in val_pos_edge_index
val_pos_edge_index = data.edge_index[:, data.val_mask & drug_drug_mask].to(device)
with torch.no_grad():
z = model.encode(x, mp_adjs)
validation_loss = model.recon_loss(z, val_pos_edge_index).item()
auc, ap = model.test(z, val_pos_edge_index, data.val_neg_edge_index)
training_losses.append(training_loss)
validation_losses.append(validation_loss)
val_auc.append(auc)
val_ap.append(ap)
if (epoch + 1) % 10 == 0:
print("Epoch: {:03d}, Training loss: {:.4f}, Validation loss: {:.4f}, AUC: {:.4f}, AP: {:.4f}".format(
epoch + 1, training_loss, validation_loss, auc, ap))
# -
import matplotlib.pyplot as plt
# %matplotlib inline
# +
fig, ax = plt.subplots(ncols=3, figsize=(24, 4))
ax[0].set_title("Log loss")
ax[0].plot(training_losses, label="Train")
ax[0].plot(validation_losses, label="Validation")
ax[0].legend()
ax[1].set_title("AUC score")
ax[1].plot(val_auc)
ax[2].set_title("Average precision")
ax[2].plot(val_ap)
# -
# #### Inspect node embeddings
#
# Now that our model is trained, we can visualize the node embeddings. We decided that our node embeddings are 64-dimensional (8 dimensions * 8 heads). We can easily convert this to a 2D visualization using any visualization algorithm. We will not go into any interpretation here, as the point of the notebook is to demonstrate how to actually fit these models.
# +
node_types = {}
for i in range(len(data.edge_type)):
n1, n2 = data.edge_index[:, i]
et = data.edge_type[i]
if et == EdgeType.DRUG_DRUG:
node_types[int(n1)] = NodeType.DRUG
node_types[int(n2)] = NodeType.DRUG
elif et == EdgeType.DRUG_GENE:
node_types[int(n1)] = NodeType.DRUG
node_types[int(n2)] = NodeType.GENE
elif et == EdgeType.GENE_DRUG:
node_types[int(n1)] = NodeType.GENE
node_types[int(n2)] = NodeType.DRUG
len(node_types)
# -
node_types = np.array([node_types[i] for i in range(len(node_types))])
node_types
pd.Series.value_counts(node_types)
embedding = z.cpu().detach().numpy()
embedding = embedding[node_types == NodeType.DRUG]
embedding
embedding.shape
# +
from openTSNE import TSNE
tsne_embedding = TSNE(exaggeration=1, n_jobs=4).fit(embedding)
# -
fig, ax = plt.subplots(figsize=(8, 8))
ax.scatter(tsne_embedding[:, 0], tsne_embedding[:, 1], c="tab:blue", s=3)
| 05-graph-attention/heterogenenous-graph-attention.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0
# language: julia
# name: julia-1.7
# ---
# # Advent of Code: Day 13
# +
using Base.Iterators
using DelimitedFiles
using Match
using Underscores
function parse_data(fn)
lines = readlines(fn)
seperatorIndex = findall(v -> v == "", lines)[1]
marks = @_ view(lines, 1:seperatorIndex - 1) |>
map(split(_, ","), __) |>
map(parse.(Int, _), __) |>
map(map(v -> v + 1, _), __)
folds = @_ view(lines, seperatorIndex + 1:length(lines)) |>
map(split(_, "="), __) |>
map([_[1], parse(Int, _[2]) + 1], __)
(M=marks,F=folds)
end
#fn = "ExampleInput.txt"
fn = "SolutionInput.txt"
Data = parse_data(fn)
Paper = zeros(maximum(map(v -> v[2], Data.M)), maximum(map(v -> v[1], Data.M)))
for (x,y) ∈ Data.M
Paper[CartesianIndex(y,x)] = 1
end
Paper
# -
# ## Part 1
# +
function foldpaper_y(paper, index)
Paper0 = view(copy(paper), 1:index - 1, :)
Paper1 = reverse(view(copy(paper), index + 1:size(paper, 1), :); dims = 1)
offset = 0
if size(Paper1, 1) != size(Paper0, 1)
offset = size(Paper0, 1) - size(Paper1, 1)
end
for index ∈ findall(v -> v > 0, Paper1)
Paper0[index + CartesianIndex(offset, 0)] = 1
end
Paper0
end
function foldpaper_x(paper, index)
Paper0 = view(copy(paper), :, 1:index - 1)
Paper1 = reverse(view(copy(paper), :, index + 1:size(paper, 2)); dims = 2)
offset = 0
if size(Paper1, 2) != size(Paper0, 2)
offset = size(Paper0, 2) - size(Paper1, 2)
end
for index ∈ findall(v -> v > 0, Paper1)
Paper0[index + CartesianIndex(0, offset)] = 1
end
Paper0
end
@_ copy(Paper) |>
reduce((paper, fold) -> begin
@match fold[1] begin
"fold along y" => foldpaper_y(paper, fold[2])
"fold along x" => foldpaper_x(paper, fold[2])
end
end, [Data.F[1]]; init=__) |>
count(v -> v > 0, __)
# -
# ## Part 2
# +
R = @_ copy(Paper) |>
reduce((paper, fold) -> begin
@match fold[1] begin
"fold along y" => foldpaper_y(paper, fold[2])
"fold along x" => foldpaper_x(paper, fold[2])
end
end, Data.F; init=__)
O = fill(".", size(R, 1), size(R, 2))
for index ∈ findall(v -> v > 0, R)
O[index] = "#"
end
open("Solution.txt", "w") do io
DelimitedFiles.writedlm(io, O)
end
| 2021/Day13/Day13Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#This module will apply the astropy Lomb-Scargle algorith to real data from the Transiting Exoplanet Survey Satellite
#First, let us import the relevant modules from the previous demo
import numpy as np
from astropy.timeseries import LombScargle
import matplotlib.pyplot as plt
# +
#Now, we need to learn how to import TESS lightcurves into python. They are storted as fits files, which is a widely used standard format in astronomy.
#Let's import the python modules needed to read fits files:
from astropy.io import fits
#And here, we define a function to import the time series, labelling t as the timestamps, y as the fluxes, and dy as the flux uncertainties.
def load_LC(f):
hdul = fits.open(f)
t=hdul[1].data['TIME']
y=hdul[1].data['PDCSAP_FLUX']
dy=hdul[1].data['PDCSAP_FLUX_ERR']
#The RA and Dec represent the astronomical coordinates of the object. These will be useful later for further investigating the source.
RA=hdul[1].header['RA_OBJ']
Dec=hdul[1].header['DEC_OBJ']
idx=y>0
t=t[idx]
y=y[idx]
dy=dy[idx]
return t, y, dy, RA, Dec
#Now, let's run the function on an example:
t, y, dy, RA, Dec=load_LC('tess2021175071901-s0040-0000000284900652-0211-s_lc.fits')
#And let's make a plot to see what things look like:
#plt.errorbar(t,y,dy)
#we see some outliers, so let us sigma clip the lightcurve:
idx=np.abs(y-np.mean(y))<10*np.std(y)
t=t[idx]
dy=dy[idx]
y=y[idx]
plt.errorbar(t,y,dy)
# +
#Let's take a closer look at the above lightcuve by zooming in:
plt.errorbar(t,y,dy)
plt.xlim(2395,2400)
#There is clearly a signal! Now it's time to see if Lomb-Scargle can identify the period
# +
#As shown in the last tutorial, we feed the times, fluxes, and uncertainties to Lomb-Scargle:
frequency, power = LombScargle(t, y, dy).autopower()
plt.plot(frequency, power)
#We see three huge peaks. Suspiciously, one falls at 720 cycles/day, and the other at 1440 cycles/day. Let's print the timestamps to find out more...
# +
print(t)
#We se large numbers for the timestamps because they are recorded in terms of a modified barycentric julian date. Let us look at the delta times between timestamps.
print(t[1:]-t[:-1])
#And since this is in units of days, let's convert to minutes:
print((t[1:]-t[:-1])*1440)
#Ah! The timestamps are taken every 2 minutes, since TESS follows equispaced sampling. That means we have a nyquist limit of 4 minutes!
# +
#Now, we modify Lomb-Scargle to respect this nyquist limit (4 mins corresponds to 360 cycles/day):
frequency, power = LombScargle(t, y, dy).autopower(maximum_frequency=360)
plt.plot(frequency, power)
#Much better! We have removed the two aliased peaks, and are left with one remaining peak. Let's find the period from this.
period=1/frequency[np.argmax(power)]
print(period)
#The period corresponds to about a third of a day!
# +
#Now, let us phase-fold the lightcurve by arranging the measurements according to the phase they were observed at.
phase=(t%period)/period
plt.errorbar(phase,y,dy)
#This is a nice looking phase fold, with the signal clearly visible, but we can make a nicer plot by binning.
# +
#Let us define a function for binning data:
def binning(phases,y,dy,N=100):
binned_LC=[]
mean_phases=np.linspace(0,1-1/N,N)
lightcurve=np.array((phases,y,dy)).T
lightcurve=lightcurve[np.argsort(lightcurve[:,0])]
for i in mean_phases:
lightcurve_bin=lightcurve[lightcurve[:,0]>i]
lightcurve_bin=lightcurve_bin[lightcurve_bin[:,0]<i+1/N]
weights=1/(lightcurve_bin[:,2]**2)
weighted_mean_flux=np.sum(lightcurve_bin[:,1]*weights)/np.sum(weights)
weighted_mean_flux_error=np.sqrt(1/np.sum(weights))
binned_LC.append((i+0.5/N,weighted_mean_flux,weighted_mean_flux_error))
binned_LC=np.array(binned_LC)
return binned_LC
binned_LC=binning(phase,y,dy,N=100)
plt.errorbar(binned_LC[:,0],binned_LC[:,1],binned_LC[:,2],ls=' ')
#A much cleaner plot than the unbinned version! This is the power of beating down noise with sqrt(N) statistics.
# +
#You are now ready to unleash this algorithm on the full TESS dataset. As a test, let us check how long it actually takes the execute.
import time
t1=time.perf_counter()
frequency, power = LombScargle(t, y, dy).autopower(maximum_frequency=360)
print(1/frequency[np.argmax(power)])
print(np.max(power))
t2=time.perf_counter()
print(t2-t1)
#You can extrapolate runtime from this number. Note that the astropy code only uses a single CPU core, so by multithreading or using a different algorithm which is already multithreaded, the code can be significantly accelerated.
# -
| Applying Lomb-Scargle to a TESS lightcurve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Script for training a denoiser
import os
os.environ['XLA_FLAGS']='--xla_gpu_cuda_data_dir=/gpfslocalsys/cuda/11.2'
# -
# %pylab inline
# +
import haiku as hk
import jax
import optax
import jax.numpy as jnp
import numpy as onp
import pickle
from functools import partial
import tensorflow as tf
import tensorflow_datasets as tfds
from jax_lensing.inversion import ks93inv, ks93
from jax_lensing.models import UResNet18
from jax_lensing.models.normalization import SNParamsTree
from jax_lensing.spectral import measure_power_spectrum, make_power_map
from astropy.io import fits
# +
dataset = "kappatng"
weight_dir='/gpfswork/rech/xdy/commun/Remy2021/score_sn1.0_std0.2'
b_mode = False
batch_size = 32
gaussian_prior = True
gaussian_path = "../../data/ktng/ktng_PS_theory.npy"
ps_data_th = onp.load(gaussian_path).astype('float32')
#sigma_gamma = 0.148
map_size = 360
resolution = 0.29
pixel_size = jnp.pi * resolution / 180. / 60. #rad/pixel
# -
ps_data = onp.load(gaussian_path).astype('float32')
ell = jnp.array(ps_data[0,:])
# massivenu: channel 4
ps_halofit = jnp.array(ps_data[1,:] / pixel_size**2) # normalisation by pixel size
# convert to pixel units of our simple power spectrum calculator
#kell = ell / (360/3.5/0.5) / float(FLAGS.map_size)
kell = ell /2/jnp.pi * 360 * pixel_size / map_size
# Interpolate the Power Spectrum in Fourier Space
power_map_th = jnp.array(make_power_map(ps_halofit, map_size, kps=kell))
# +
def forward_fn(x, s, is_training=False):
denoiser = UResNet18(n_output_channels=1)
return denoiser(x, s, is_training=is_training)
model = hk.without_apply_rng(hk.transform_with_state(forward_fn))
# +
model_name = "model-final.pckl"
with open(weight_dir+'/'+ model_name, 'rb') as file:
params, state, sn_state = pickle.load(file)
# -
def log_gaussian_prior(map_data, sigma, ps_map):
data_ft = jnp.fft.fft2(map_data) / float(map_size)
return -0.5*jnp.sum(jnp.real(data_ft*jnp.conj(data_ft)) / (ps_map+sigma[0]**2))
gaussian_prior_score = jax.vmap(jax.grad(log_gaussian_prior), in_axes=[0,0, None])
# +
def log_gaussian_prior_b(map_data, sigma):
data_ft = jnp.fft.fft2(map_data) / float(map_size)
return -0.5*jnp.sum(jnp.real(data_ft*jnp.conj(data_ft)) / (sigma[0]**2))
gaussian_prior_score_b = jax.vmap(jax.grad(log_gaussian_prior_b), in_axes=[0,0])
# +
def score_fn(params, state, x, sigma, is_training=False):
if b_mode:
x = x.reshape((-1,360,360,2))
ke = x[...,0]
kb = x[...,1]
else:
ke = x.reshape((-1,360,360))
if gaussian_prior:
# If requested, first compute the Gaussian prior
gs = gaussian_prior_score(ke, sigma.reshape((-1,1,1)), power_map_th)
gs = jnp.expand_dims(gs, axis=-1)
#print((jnp.abs(sigma.reshape((-1,1,1,1)))**2).shape, (gs).shape)
net_input = jnp.concatenate([ke.reshape((-1,360,360,1)), jnp.abs(sigma.reshape((-1,1,1,1)))**2 * gs],axis=-1)
res, state = model.apply(params, state, net_input, sigma.reshape((-1,1,1,1)), is_training=is_training)
if b_mode:
gsb = gaussian_prior_score_b(kb, sigma.reshape((-1,1,1)))
gsb = jnp.expand_dims(gsb, axis=-1)
else:
gsb = jnp.zeros_like(res)
else:
res, state = model.apply(params, state, ke.reshape((-1,360,360,1)), sigma.reshape((-1,1,1,1)), is_training=is_training)
gs = jnp.zeros_like(res)
gsb = jnp.zeros_like(res)
return _, res, gs, gsb
score_fn = partial(score_fn, params, state)
# +
noise = 0.2
start_and_end_times = jnp.logspace(log10(0.99*noise**2),-5)
plot(start_and_end_times)
yscale('log')
# +
mask = fits.getdata('../../data/COSMOS/cosmos_full_mask_0.29arcmin360copy.fits').astype('float32')
convergence = fits.getdata('../../data/ktng/ktng_kappa360v2.fits').astype('float32')
meas_shear = jnp.stack(ks93inv(convergence, jnp.zeros_like(convergence)), axis=-1)
std1 = fits.getdata('../../data/COSMOS/std1.fits').astype('float32').reshape((360,360,1))
std2 = fits.getdata('../../data/COSMOS/std2.fits').astype('float32').reshape((360,360,1))
sigma_gamma = jnp.concatenate([std1, std2], axis=-1)
#sigma_gamma = 0.148
#meas_shear += sigma_gamma*np.random.randn(360,360,2)
meas_shear += sigma_gamma * jax.random.normal(jax.random.PRNGKey(0), meas_shear.shape)
#g1 = fits.getdata('../data/COSMOS/cosmos_full_e1_0.29arcmin360.fits').astype('float32').reshape([map_size, map_size, 1])
#g2 = fits.getdata('../data/COSMOS/cosmos_full_e2_0.29arcmin360.fits').astype('float32').reshape([map_size, map_size, 1])
#meas_shear = onp.concatenate([g1, g2], axis=-1)
mask = jnp.expand_dims(mask, -1)
sigma_mask = (1-mask)*1*10**3
masked_true_shear = meas_shear*mask
figure(figsize(14,5))
subplot(141)
imshow(masked_true_shear[..., 0])
subplot(142)
imshow(masked_true_shear[..., 1])
subplot(143)
imshow(convergence)
subplot(144)
imshow(mask[...,0])
# +
def log_likelihood(x, sigma, meas_shear, mask):
""" Likelihood function at the level of the measured shear
"""
if b_mode:
x = x.reshape((360, 360,2))
ke = x[...,0]
kb = x[...,1]
else:
ke = x.reshape((360, 360))
kb = jnp.zeros(ke.shape)
model_shear = jnp.stack(ks93inv(ke, kb), axis=-1)
return - jnp.sum((model_shear - masked_true_shear)**2/((sigma_gamma)**2 + sigma**2 + sigma_mask) )/2.
#return - jnp.sum(mask*(model_shear - meas_shear)**2/((sigma_gamma)**2 + sigma**2) )/2.
likelihood_score = jax.vmap(jax.grad(log_likelihood), in_axes=[0,0, None, None])
# -
def score_prior(x, sigma):
if b_mode:
_, res, gaussian_score, gsb = score_fn(x.reshape(-1,360, 360,2), sigma.reshape(-1,1,1,1))
else:
_, res, gaussian_score, gsb = score_fn(x.reshape(-1,360, 360), sigma.reshape(-1,1,1))
ke = (res[..., 0:1] + gaussian_score).reshape(-1, 360*360)
kb = gsb[...,0].reshape(-1, 360*360)
if b_mode:
return jnp.stack([ke, kb],axis=-1)
else:
return ke
def total_score_fn(x, sigma):
if b_mode:
sl = likelihood_score(x, sigma, masked_true_shear, mask).reshape(-1, 360*360,2)
else:
sl = likelihood_score(x, sigma, masked_true_shear, mask).reshape(-1, 360*360)
sp = score_prior(x, sigma)
if b_mode:
return (sl + sp).reshape(-1, 360*360*2)
else:
return (sl + sp).reshape(-1, 360*360)
#return (sp).reshape(-1, 360*360,2)
# + active=""
# def log_gaussian_prior(map_data, sigma, ps_map):
# map_data = map_data.reshape(360, 360)
# data_ft = jnp.fft.fft2(map_data) / float(map_size)
# return -0.5*jnp.sum(jnp.real(data_ft*jnp.conj(data_ft)) / (ps_map+sigma**2))
#
# gaussian_prior_score = jax.vmap(jax.grad(log_gaussian_prior), in_axes=[0,0, None])
#
# def log_likelihood(x, sigma, meas_shear, mask):
# """ Likelihood function at the level of the measured shear
# """
# ke = x.reshape((map_size, map_size))
# kb = jnp.zeros(ke.shape)
# model_shear = jnp.stack(ks93inv(ke, kb), axis=-1)
#
# return - jnp.sum(mask*(model_shear - meas_shear)**2/((sigma_gamma)**2 + sigma**2 + sigma_mask) )/2.
#
# likelihood_score = jax.vmap(jax.grad(log_likelihood), in_axes=[0,0, None, None])
#
#
# +
import numpy as onp
batch_size = 100
initial_temperature = 1.3
if b_mode:
init_image = np.stack([initial_temperature*np.random.randn(batch_size,360*360),
initial_temperature*np.random.randn(batch_size,360*360)], axis=-1)
imshow(init_image[0,...,0].reshape(360, 360))
else:
init_image = initial_temperature*np.random.randn(batch_size,360*360)
imshow(init_image[0,...].reshape(360, 360))
#init_image = initial_temperature*np.random.randn(batch_size,360*360)
print(init_image.shape)
colorbar()
# +
from scipy import integrate
@jax.jit
def dynamics(t, x):
if b_mode:
x = x.reshape([-1,360,360,2])
return - 0.5*total_score_fn(x, sigma=jnp.ones((batch_size,1,1,1))*jnp.sqrt(t)).reshape([-1])
else:
x = x.reshape([-1,360,360])
return - 0.5*total_score_fn(x, sigma=jnp.ones((batch_size,1,1))*jnp.sqrt(t)).reshape([-1])
# +
# %%time
noise = initial_temperature
start_and_end_times = jnp.logspace(log10(0.99*noise**2),-5, num=50)
solution = integrate.solve_ivp(dynamics,
[noise**2,(1e-5)],
init_image.flatten(),
t_eval=start_and_end_times)
# -
# +
if b_mode:
sol = solution.y[:,-1].reshape([batch_size,360,360,2])[...,0]
else:
sol = solution.y[:,-1].reshape([batch_size,360,360])
kwargs = dict(vmin=-0.05, vmax=0.2, cmap='magma')
subplot(131)
title('target')
imshow(convergence, **kwargs)
subplot(132)
title('sample')
imshow(sol[0,...], **kwargs)
subplot(133)
title('mean')
imshow(sol.mean(axis=0), **kwargs)
# -
figure(figsize=[20,20])
kwargs=dict(cmap='magma', vmin=-0.05, vmax=0.2)
subplot(331)
imshow(sol[0,...], **kwargs)
subplot(332)
imshow(sol[1,...], **kwargs)
subplot(333)
imshow(sol[2,...], **kwargs)
subplot(334)
imshow(sol[3,...], **kwargs)
subplot(335)
imshow(sol[4,...], **kwargs)
subplot(336)
imshow(sol[5,...], **kwargs)
subplot(337)
imshow(sol[6,...], **kwargs)
subplot(338)
imshow(sol[7,...], **kwargs)
subplot(339)
imshow(sol[8,...], **kwargs)
# +
#jnp.save('res/no_mode_b_full/samples4.npy', sol)
# + active=""
# sol_B = solution.y[:,-1].reshape([batch_size,360,360,2])[...,1]
# + active=""
# subplot(221)
# imshow(sol_B[0,...])
# colorbar()
# subplot(222)
# imshow(sol_B[1,...])
# colorbar()
# subplot(223)
# imshow(sol_B[2,...])
# colorbar()
# subplot(224)
# imshow(sol_B[3,...])
# colorbar()
# +
def rmse(a, b, mask):
a = a - np.mean(a)
b = b - np.mean(b)
return(np.sqrt(np.sum(mask*(a-b)*(a-b))/np.sum(mask)))
print(rmse(convergence, sol.mean(axis=0), mask[...,0]))
# +
l = [rmse(convergence, sol[0,...].reshape(360,360), mask[...,0])]
for i in range(1,sol.shape[0]):
l.append(rmse(convergence, sol[:i,...].mean(0).reshape(360,360), mask[...,0]))
plot(l)
# -
# ## Power Spectrum
# Compare to fiducial power spectrum, Gaussian posterior sample and target $\kappa$ map
mps = jax.vmap(measure_power_spectrum, in_axes=[0,None])
ell_post, cl_post = mps(sol, pixel_size)
cl_post_mean = cl_post.mean(0)
cl_post_std = cl_post.std(0)
# +
figure(figsize=(7,5), dpi=100)
kwargs = dict(linewidth=1.5)
gaussian_path = "../../data/ktng/ktng_PS_theory.npy"
ps_data_th = np.load(gaussian_path).astype('float32')
plot(ps_data_th[0,:], ps_data_th[1,:], color='pink', label='Fiducial', **kwargs)
l_true, C_true = measure_power_spectrum(convergence, pixel_size)
loglog(l_true, C_true, color='tab:red', label=r'Target $\kappa$', **kwargs)
wiener = np.load('../../notebooks/results/gaussian_samples.npy')
l_true, C_true = measure_power_spectrum(wiener[0,...], pixel_size)
loglog(l_true, C_true, color='tab:orange', label='Wiener posterior sample', **kwargs)
plot(ell_post[0,...], cl_post_mean, color='tab:blue', **kwargs)
fill_between(ell_post[0,...], cl_post_mean-cl_post_std,
cl_post_mean+cl_post_std, color='tab:blue' ,alpha=0.3, label='Our posterior samples')
legend()
ylim(5.e-13,6e-9)
xlim(300,4e4)
xscale('log')
yscale('log')
ylabel(r'$C_\ell$', fontsize=18)
xlabel(r'$\ell$', fontsize=18)
savefig('plots/ps-ODE-comparison-1.pdf', bbox_inches='tight')
# +
figure(figsize=(7,3), dpi=100)
kwargs = dict(linewidth=1.5)
hlines(y=1., xmin=300,xmax=4e4, linestyle='--', color='k')
l_true, C_true = measure_power_spectrum(convergence, pixel_size)
l_true, C_wiener = measure_power_spectrum(wiener[0,...], pixel_size)
plot(l_true, C_wiener/C_true, color='tab:orange', label='Wiener posterior sample', **kwargs)
ell_post, cl_post = mps(sol, pixel_size)
mean = (cl_post/C_true).mean(0)
std = (cl_post/C_true).std(0)
plot(l_true, mean, color='tab:blue', **kwargs)
fill_between(l_true, mean-std,
mean+std, color='tab:blue' ,alpha=0.3, label='Our posterior samples')
legend()
xlim(300,4e4)
xscale('log')
yscale('log')
ylabel(r'$C_\ell/C_\ell^\mathrm{target}$', fontsize=18)
xlabel(r'$\ell$', fontsize=18)
savefig('plots/ps-ODE-comparison-2.pdf', bbox_inches='tight')
# -
# !ls res
1+1
| papers/Remy2021/PosteriorSampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (.qpy)
# language: python
# name: qpy
# ---
# # Unnecessary Complexity Through Layer Proliferation
# In this blog, we continue to elaborate on our loosely `transformer`-based approach to training our computer to "understand" elementary symbolic arithmetic. Just as previously mentioned, a quick and fun intro to the venerable transformer is [here](http://jalammar.github.io/illustrated-transformer/).
#
# Our objective is to expand our layers' repertoire to include the `encoder` and the `decoder` stacks, as well as to ultimately arrive at a model representable by the [graph](./layers.pdf).
#
# We need to prep our environment to run any meaningful code:
import numpy as np
import tensorflow as tf
import dataset as qd
import ragged as qr
ks = tf.keras
kl = ks.layers
# Before we start to focus on our stacks, an important feature of encoding textual inputs needs to be considered. To aid in making sense of a text, we need to include not just the information carried by the tokens themselves but also their position in the input sequence.
#
# Following the "positional encoding" approach from [here](https://github.com/tensorflow/examples/blob/master/community/en/position_encoding.ipynb), we can define our `pos_timing` function as follows.
#
# The quick graphical plot helps us confirm the correctness of our code as the concatenated `sin` and `cos` timing signals give us finely graded and rich positional embeddings:
# +
def pos_timing(width, depth):
assert depth % 2 == 0
d = np.arange(depth)[np.newaxis, :]
d = 1 / np.power(10000, (2 * (d // 2)) / np.float32(depth))
t = np.arange(width)[:, np.newaxis] * d
t = [np.sin(t[:, 0::2]), np.cos(t[:, 1::2])]
t = np.concatenate(t, axis=-1)[np.newaxis, ...]
return t
pos = pos_timing(50, 512)
import matplotlib.pyplot as plt
plt.pcolormesh(pos[0], cmap='RdBu')
plt.xlabel('Depth')
plt.xlim((0, 512))
plt.ylabel('Position')
plt.colorbar()
plt.show()
# -
# Loading our already created meta data from the source files gives us:
print(qd.vocab)
# We continue with defining our own shared Keras "base" layer.
#
# At this time, we only store a reference to our parameters instance in it, as all our layers will need to use this resource:
class Layer(kl.Layer):
def __init__(self, ps, **kw):
kw.setdefault('dtype', tf.float32)
super().__init__(**kw)
self.ps = ps
# The `Embed` layer is taken directly from our previous blog.
#
# As we need to add the above mentioned positional "timing signals" to our embeddings, we first create just such a `constant` tensor.
#
# Then, using the previously mentioned `RaggedTensor`-technique of extracting the ragged-shaped values from a dense tensor, we simply add the positional info to the already determined embedding:
class Embed(Layer):
def __init__(self, ps):
super().__init__(ps)
s = (ps.dim_vocab, ps.dim_hidden)
self.emb = self.add_weight(name='emb', shape=s)
p = pos_timing(ps.len_max_input, ps.dim_hidden)
p = tf.constant(p, dtype=tf.float32)
self.pos = tf.broadcast_to(p, [ps.dim_batch] + p.shape[1:])
def call(self, x):
fv, rs = x
x = tf.RaggedTensor.from_row_splits(fv, rs)
y = tf.ragged.map_flat_values(tf.nn.embedding_lookup, self.emb, x)
y += tf.RaggedTensor.from_tensor(self.pos, lengths=y.row_lengths())
return y
# The next layers to write are the `Encode` and `Decode` stacks.
#
# We implement them as simple lists of the respective `Encoder` and `Decoder` components. When calling the stacks, the code simply loops through the component lists and calls the components.
#
# In order to "chain" the stacks, every component is given the output of the previous component as its input.
#
# In the case of the `Decoder` components, and in addition to their regular inputs, we also supply the previously encoded output as their `ctx` argument:
# +
class Encode(Layer):
def __init__(self, ps):
super().__init__(ps)
self.encs = [Encoder(self, f'enc_{i}') for i in range(ps.dim_stacks)]
def call(self, x):
y = x
for e in self.encs:
y, ctx = e(y)
return [y, ctx]
class Decode(Layer):
def __init__(self, ps):
super().__init__(ps)
self.decs = [Decoder(self, f'dec_{i}') for i in range(ps.dim_stacks)]
def call(self, x):
y, ctx = x
for d in self.decs:
y = d([y, ctx])
return y
# -
# The mirror "image" of the `Embed` layer is our `Debed` layer.
#
# While the embedding step maps `int` tokens to higher-dimensional, learned `float` values, the "debedding" step does the opposite. It takes the higher-dimensional values and maps them to learned `one-hot` vectors, corresponding to approximate output tokens.
#
# As debedding is implemented using a `Dense` component, it also requires a fixed width. Just as in the previous blog, we simply pad our ragged tensors with `0`s to our `len_max_input` parameter, as our calculations are complete and the raggedness is not needed any longer:
class Debed(Layer):
def __init__(self, ps):
super().__init__(ps)
self.max_len = u = ps.len_max_input
s = [u * ps.dim_hidden, ps.dim_vocab]
self.dbd = Dense(self, 'dbd', s)
def call(self, x):
y = x.to_tensor()
s = tf.shape(y)
y = tf.pad(y, [[0, 0], [0, self.max_len - s[-2]], [0, 0]])
y = tf.reshape(y, [-1, self.max_len * s[-1]])
y = self.dbd(y)
return y
# We have now completed the definition of our top layers as Keras layers, but we still need to define the inner components.
#
# We could continue using the seemingly "heavy" Keras layers and nest them deeper. Instead, as presented in a previous blog, we switch over to using the much lighter-weight `Module` as the base class for our inner components.
#
# Our `Encoder` thus becomes a simple module containing the self-attention followed by the feed-forward mechanisms. We fittingly call the inner modules `reflect` and `conclude`.
#
# Our `Decoder` also adds the attention layer looking at the previously encoded "context". Hence, the module encapsulating this attention component is called `consider`:
# +
class Encoder(tf.Module):
def __init__(self, layer, name=None):
super().__init__(name=name)
with self.name_scope:
self.reflect = Attention(layer, 'refl')
self.conclude = Conclusion(layer, 'conc')
@tf.Module.with_name_scope
def __call__(self, x):
y, ctx = self.reflect([x, None])
y = self.conclude(y)
return [y, ctx]
class Decoder(tf.Module):
def __init__(self, layer, name=None):
super().__init__(name=name)
with self.name_scope:
self.reflect = Attention(layer, 'refl')
self.consider = Attention(layer, 'cnsd')
self.conclude = Conclusion(layer, 'conc')
@tf.Module.with_name_scope
def __call__(self, x):
x, ctx = x
y, _ = self.reflect([x, None])
y, _ = self.consider([y, ctx])
y = self.conclude(y)
return y
# -
# Our `Attention` component, again based on `Module`, is taken directly from the previous blog. As explained there, it relies on and takes advantage of the new `RaggedTensor`s:
class Attention(tf.Module):
def __init__(self, layer, name):
super().__init__(name=name)
h = layer.ps.dim_hidden
self.scale = 1 / (h**0.5)
with self.name_scope:
self.q = layer.add_weight('q', shape=(h, h))
self.k = layer.add_weight('k', shape=(h, h))
self.v = layer.add_weight('v', shape=(h, h))
@tf.Module.with_name_scope
def __call__(self, x):
x, ctx = x
q = x.with_values(tf.einsum('ni,ij->nj', x.flat_values, self.q))
k = x.with_values(tf.einsum('ni,ij->nj', x.flat_values, self.k))
v = x.with_values(tf.einsum('ni,ij->nj', x.flat_values, self.v))
y = tf.einsum('bsi,bzi->bsz', q.to_tensor(), k.to_tensor())
y = tf.nn.softmax(y * self.scale)
y = tf.einsum('bsz,bzi->bsi', y, v.to_tensor())
y = tf.RaggedTensor.from_tensor(y, lengths=x.row_lengths())
return [y, tf.constant(1)]
# A new component is our `Conclusion` module. It implements the "feed-forward" functionality of the transformer.
#
# In simple terms, it takes the attention-enhanced, higher-dimensional, element-wise mapping of the token sequence and it first `inflates` it to an even higher dimension with a non-linearity, or `activation`, at the end as its "concluding" step.
#
# Then it `deflates` the activated mapping back to our hidden dimension, making it available for the next level in the stack.
#
# The same `RaggedTensor` trick, as the one we used in the `Attention` module, applies at the end:
class Conclusion(tf.Module):
def __init__(self, layer, name):
super().__init__(name=name)
ps = layer.ps
self.max_len = w = ps.len_max_input
w *= ps.dim_hidden
with self.name_scope:
s = [w, ps.dim_dense]
self.inflate = Dense(layer, 'infl', s, activation='relu')
s = [ps.dim_dense, w]
self.deflate = Dense(layer, 'defl', s, bias=False)
@tf.Module.with_name_scope
def __call__(self, x):
y = x.to_tensor()
s = tf.shape(y)
y = tf.pad(y, [[0, 0], [0, self.max_len - s[-2]], [0, 0]])
y = tf.reshape(y, [-1, self.max_len * s[-1]])
y = self.inflate(y)
y = self.deflate(y)
y = tf.reshape(y, [-1, self.max_len, s[-1]])
y = tf.RaggedTensor.from_tensor(y, lengths=x.row_lengths())
return y
# Our last component is the `Dense` module.
#
# It simply re-implements the Keras layer with the same name, yet with more focused, streamlined functionality and minimal configurability.
#
# The interesting aspect of this module, just as our `Attention` module above, is that the necessarily created Keras weights are added using the enclosing Keras layer, however, topologically, they are directly listed as part of their respective modules:
class Dense(tf.Module):
bias = None
activation = None
def __init__(self, layer, name, shape, activation=None, bias=True):
super().__init__(name=name)
with self.name_scope:
kw = dict(shape=shape, initializer='glorot_uniform')
self.kern = layer.add_weight('kern', **kw)
if bias:
kw.update(shape=shape[1:], initializer='zeros')
self.bias = layer.add_weight('bias', **kw)
self.activation = ks.activations.get(activation)
@tf.Module.with_name_scope
def __call__(self, x):
y = tf.einsum('bi,ij->bj', x, self.kern)
if self.bias is not None:
y = tf.nn.bias_add(y, self.bias)
if self.activation:
y = self.activation(y)
return y
# And now we are ready to define our model.
#
# We have the two inputs, the two components of our input `RaggedTensor`.
#
# We also use our new `Embed`, `Encode`, `Decode` and `Debed` Keras layers, with all the internal, light-weight modules hidden at this level.
#
# The rest of the model is simply carried over from the previous blog:
def model_for(ps):
x = [ks.Input(shape=(), dtype='int32'), ks.Input(shape=(), dtype='int64')]
y = Embed(ps)(x)
y = Encode(ps)(y)
y = Decode(ps)(y)
y = Debed(ps)(y)
m = ks.Model(inputs=x, outputs=y)
m.compile(optimizer=ps.optimizer, loss=ps.loss, metrics=[ps.metric])
print(m.summary())
return m
# Our parameters need to be adjusted to provide parametric values for our stacks:
params = dict(
dim_batch=2,
dim_dense=150,
dim_hidden=6,
dim_stacks=2,
dim_vocab=len(qd.vocab),
len_max_input=20,
loss=ks.losses.SparseCategoricalCrossentropy(from_logits=True),
metric=ks.metrics.SparseCategoricalAccuracy(),
num_epochs=5,
num_shards=2,
optimizer=ks.optimizers.Adam(),
)
# By firing up our training session, we can confirm the model's layers and connections. The listing of a short session follows.
#
# We can easily adjust the parameters to tailor the length of the sessions to our objectives. However, at this point the results are still largely meaningless and extending the trainings is not yet warranted.
ps = qd.Params(**params)
import masking as qm
qm.main_graph(ps, qr.dset_for(ps), model_for(ps))
# With our TensorBoard `callback` in place, the model's `fit` method will generate the standard summaries that TB can conveniently visualize.
#
# If you haven't run the code below, an already generated graph is [here](./layers.pdf).
# +
# #%load_ext tensorboard
# #%tensorboard --logdir /tmp/q/logs
# -
# We can also switch over to the new `eager` execution mode.
#
# This is particularly convenient for experimentation, as all ops are immediately executed. Here is a much shortened `eager` session:
ps.num_epochs = 1
qr.main_eager(ps, qr.dset_for(ps).take(100), model_for(ps))
# This concludes our blog, please see how to further customize our model by clicking on the next blog.
| qnarre.com/static/pybooks/layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Inverse covariance matrices
#
# In other notebooks we see that the emulator does well in predicting covariance matrices. However, parameter constraints are largely driven by $\chi^2$, which depends on the precision matrix or inverse covariance matrix $C^{-1}$. Since precision matrices are by definition invertable, we should be able to emulate them without issue. Here we will attempt that.
import numpy as np
from scipy import stats
import covariance_emulator
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rc("font", size=14, family="serif")
#plt.rc("text", usetex=True)
#Read in the domain locations, or locations in parameter space
parameters = np.loadtxt("cosmo_parameters.txt")[:, :]
#covs = np.load("tomo0_sub_covs.npy")
covs = np.load("gaussian_tomo_sub_covs.npy")
#covs = np.load("gaussian_full_covs.npy")
#Take only a subset
covs = covs[:, 0:200, 0:200]
icovs = np.array([np.linalg.inv(covs[i]) for i in range(len(covs))])
# +
#View the correlation matrix of the first
def corr_from_cov(cov):
D = np.diag(np.sqrt(cov.diagonal()))
Di = np.linalg.inv(D)
return np.dot(Di, np.dot(cov, Di))
def view_corr(cov, lncov=False):
R = corr_from_cov(cov)
fig, ax = plt.subplots()
if lncov:
R = np.log(np.fabs(cov))
im = ax.imshow(R, interpolation="nearest", origin="lower")
plt.colorbar(im)
return
# -
#Split off the last covariance matrix
test_cov = covs[-1]
test_icov = icovs[-1]
test_parameters = parameters[-1]
covs = covs[:-1]
icovs = icovs[:-1]
parameters = parameters[:-1]
#Create an emulator
Emu = covariance_emulator.CovEmu(parameters, icovs, NPC_D=6, NPC_L=6)
iCpredicted = Emu.predict(test_parameters)
#Given a covariance matrix, make realizations of the noise, and then find the optimal kernel set up
def best_kernel_for_C(C, N_samples=1000):
dof = len(C)
means = np.zeros(dof)
chi2s = np.zeros(N_samples)
noise_realizations = np.array([np.random.multivariate_normal(means, C) for i in range(N_samples)])
import george.kernels as kernels
kerns = [kernels.ExpSquaredKernel,
kernels.Matern52Kernel,
kernels.Matern32Kernel]
names = ["Exp2", "Mat52", "Mat32"]
Npars = len(parameters[0])
metric_guess = np.std(parameters, 0)
#Loop over kernel combinations and compute the chi2 shift
best_shift = 1e99
best_kernels = None
for nameD, kd in zip(names, kerns):
kernel_D = 1.*kd(metric=metric_guess, ndim=Npars)
for nameL, kl in zip(names, kerns):
kernel_L = 1.*kl(metric=metric_guess, ndim=Npars)
Emu = covariance_emulator.CovEmu(parameters, icovs, NPC_D=6, NPC_L=6,
kernel_D = kernel_D, kernel_lp = kernel_L)
shift = 0
try:
iCpredicted = Emu.predict(test_parameters)
except np.linalg.LinAlgError:
shift = 1e99
else:
for i in range(N_samples):
chi2s[i] = np.dot(noise_realizations[i], np.dot(iCpredicted, noise_realizations[i]))
shift = np.mean(chi2s) - dof
if shift < best_shift and shift > 0:
best_shift = shift
best_name = "%s %s"%(nameD, nameL)
best_kernels = [kernel_D, kernel_L]
print("%s %s: %e / %d"%(nameD, nameL, shift, dof))
print("Best combination: %s"%best_name)
print("\tshift/dof = %e / %d"%(best_shift, dof))
return best_kernels
best_kernels = best_kernel_for_C(test_cov)
kernel_D, kernel_L = best_kernels
Emu = covariance_emulator.CovEmu(parameters, icovs, NPC_D=6, NPC_L=6,
kernel_D = kernel_D, kernel_lp = kernel_L)
iCpredicted = Emu.predict(test_parameters)
# ## Assessing the emulator performance
#
# One of the best ways to assess the performance of the emulator is to directly compare the true covariance to the emulated covariance. In the next cell, I will draw realizations of the noise from the true covariance, and compute $\chi^2$ values of these noises compared agains the emulated covariance. Then, by checking this against the expected distribution, we can see the performance of the emulator.
# +
#Define a function where we input two covariances, and get back out a list of chi2s
def get_chi2s_between_C_and_iC(C1, iC2, N_samples=10000):
means = np.zeros(len(C1))
chi2s = np.zeros(N_samples)
for i in range(N_samples):
x = np.random.multivariate_normal(means, C1)
chi2s[i] = np.dot(x, np.dot(iC2, x))
return chi2s
dof = len(test_cov)
x = np.linspace(dof*.7, dof*1.4, 1000)
# -
chi2s = get_chi2s_between_C_and_iC(test_cov, test_icov)
plt.hist(chi2s, density=True, bins=100)
plt.plot(x, stats.chi2.pdf(x, dof))
plt.title(r"$C_{\rm true}$ vs $C_{\rm true}^{-1}$")
chi2s = get_chi2s_between_C_and_iC(test_cov, icovs[0], 2000)
plt.hist(chi2s, density=True, bins=100)
plt.plot(x, stats.chi2.pdf(x, dof))
plt.title(r"$C_{\rm true}$ vs $C_{\rm 0}^{-1}$")
plt.xlabel(r"$\chi^2$")
plt.axvline(dof, color="k", ls="--")
ax = plt.gca()
print("Chi2/dof shift = %.2f / %d"%(np.mean(chi2s) - dof, dof))
chi2s = get_chi2s_between_C_and_iC(test_cov, iCpredicted, 10000)
plt.hist(chi2s, density=True, bins=100)
plt.plot(x, stats.chi2.pdf(x, dof))
plt.title(r"$C_{\rm true}$ vs $C_{\rm emu}^{-1}$")
plt.xlabel(r"$\chi^2$")
plt.axvline(dof, color="k", ls="--")
ax = plt.gca()
print("Chi2/dof shift = %.2f / %d"%(np.mean(chi2s) - dof, dof))
#plt.savefig("chi2_realizations.png", dpi=300, bbox_inches="tight")
print chi2s[:4]
print np.mean(chi2s)
print np.max(chi2s), np.min(chi2s)
| notebooks/Emulating inverse matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="mlfU2M0xPdAh"
# # **Выводы в конце ноутбука (построена модель, также реализована собственная модель; loss function и training loop сделаны, не все работает, как запланировано():**
# + [markdown] id="Q9LbFlgP5nrh"
# 1. https://kishorepv.github.io/DSSM/
# 2. https://github.com/v-mk-s/dssm
#
# чекнуть позже
# + [markdown] id="UNxseYLzzvCv"
# # Seminar: simple question answering
# 
#
# Today we're going to build a retrieval-based question answering model with metric learning models.
#
# _this seminar is based on original notebook by [<NAME>](https://github.com/Omrigan/)_
#
#
# + id="eUxx5lPpzvCx"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import pandas as pd
import torch
from sklearn.metrics import log_loss, roc_auc_score
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from keras.preprocessing.sequence import pad_sequences
# + id="k5tlHnRNz1gh" outputId="bd7b6c50-f4f5-4036-e59d-84d86fe4eb01" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472235922, "user_tz": -180, "elapsed": 465, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
# !wget https://raw.githubusercontent.com/yandexdataschool/Practical_DL/fall18/week11_dssm/utils.py
# + id="L-Rp5Msv0BNf" outputId="544af2c5-0c10-41b8-ebc4-f1cc0836eff5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472237612, "user_tz": -180, "elapsed": 1698, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
import nltk
nltk.download('punkt')
# + [markdown] id="OxgOGXMbzvC2"
# ### Dataset
#
# Today's data is Stanford Question Answering Dataset (SQuAD). Given a paragraph of text and a question, our model's task is to select a snippet that answers the question.
#
# We are not going to solve the full task today. Instead, we'll train a model to __select the sentence containing answer__ among several options.
#
# As usual, you are given an utility module with data reader and some helper functions
# + id="US39zXJlzvC2"
import utils
# !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v2.0.json -O squad-v2.0.json 2> log
# backup download link: https://www.dropbox.com/s/q4fuihaerqr0itj/squad.tar.gz?dl=1
train, test = utils.build_dataset('./squad-v2.0.json', tokenized=True)
# + id="pF7j-vCpzvC6" outputId="558f641c-d82e-404f-d4ef-8533a3a0d810" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1644472252218, "user_tz": -180, "elapsed": 58, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
# the data comes pre-tokenized with this simple tokenizer:
utils.tokenize("I... I'm the monument to all your sins.")
# + id="MNr15FbFzvC9" outputId="aff4d098-710c-4290-d1f4-9a960565aec0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472252220, "user_tz": -180, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
pid, question, options, correct_indices, wrong_indices = train.iloc[40]
print('QUESTION', question, '\n') # !! где-то должен быть еще текст
for i, cand in enumerate(options):
print(['[ ]', '[v]'][i in correct_indices], cand)
# + id="cyeFnX6eDuEJ" outputId="fdc18547-4196-47b5-c679-290c4853bc57" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472252221, "user_tz": -180, "elapsed": 42, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
train['question'][:5]
# + id="3vvGDZJUGoZK"
words_o = [w.split() for s in train['options'].values for w in s]
words_o = [w for v in words_o for w in v if w not in '?\'&,.;']
# + id="8dQ3ufZlFC18"
words_q = [s.split() for s in train['question'].values]
words_q = [w for v in words_q for w in v if w not in '?\'&,.;']
# + [markdown] id="sKnrxjsKzvDA"
# ### Tokens & vocabularies
#
# The procedure here is very similar to previous nlp weeks: preprocess text into tokens, create dictionaries, etc.
# + id="7Oh7zJoJzvDC"
from tqdm import tqdm, trange
from collections import Counter, defaultdict
#Dictionary of {token : count}
words = []
for s in train['question']:
for w in utils.tokenize(s).split():
words.append(w)
for b in train['options']:
for s in b:
for w in utils.tokenize(s).split():
words.append(w)
# compute counts for each token; use token_counts;
# count BOTH in train['question'] and in train['options']
# + id="WUt1CTEGIhBt"
token_counts = Counter(words)
# + id="miZzEh1UzvDG" outputId="f8fc0a7d-4cd8-4edc-83b3-39fa8932c7de" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472260250, "user_tz": -180, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
print("Total tokens:", sum(token_counts.values()))
print("Most common:", token_counts.most_common(5))
assert 9000000 < sum(token_counts.values()) < 9100000, "are you sure you counted all unique tokens in questions and options?"
# + [markdown] id="mlw19UFFzvDJ"
# We shall only keep tokens that are present at least 4 times
# + id="aNvC_4tDzvDK" outputId="f7390e88-c3b2-49ae-c4f7-ff804aa4561f" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472260251, "user_tz": -180, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
MIN_COUNT = 5
tokens = [w for w, c in token_counts.items() if c >= MIN_COUNT]
tokens = ["_PAD_", "_UNK_"] + tokens
print("Tokens left:", len(tokens))
# + id="A9NK0qpRzvDM"
# a dictionary from token to it's index in tokens
token_to_id = {t:i for i,t in enumerate(tokens)}
# + id="bAWawL47zvDP"
assert token_to_id['me'] != token_to_id['woods']
assert token_to_id[tokens[42]]==42
assert len(token_to_id)==len(tokens)
# + id="5Xn5NzZ0zvDS"
PAD_ix = token_to_id["_PAD_"]
UNK_ix = token_to_id['_UNK_']
#good old as_matrix for the third time
def as_matrix(sequences, max_len=None):
if isinstance(sequences[0], (str, bytes)):
sequences = [utils.tokenize(s).split() for s in sequences]
max_len = max_len or max(map(len,sequences))
matrix = np.zeros((len(sequences), max_len), dtype='int32') + PAD_ix
for i, seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_ix) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
# + id="wK25QWBNzvDV" outputId="943820dd-2198-4893-ab26-76e7a77601d9" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472260537, "user_tz": -180, "elapsed": 309, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
test = as_matrix(["Definitely, thOsE tokens areN'T LowerCASE!!", "I'm the monument to all your sins."])
print(test)
# assert test.shape==(2,8)
print("Correct!")
# + [markdown] id="T058hedOzvDX"
# ### Data sampler
#
# Our model trains on triplets: $<query, answer^+, answer^->$
#
# For your convenience, we've implemented a function that samples such triplets from data
# + id="Eg4xlH_6zvDX"
import random
import torch
lines_to_tensor = lambda lines, max_len=None: torch.tensor(
as_matrix(lines, max_len=max_len), dtype=torch.int64)
def iterate_minibatches(data, batch_size, shuffle=True, cycle=False):
"""
Generates minibatches of triples: {questions, correct answers, wrong answers}
If there are several wrong (or correct) answers, picks one at random.
"""
indices = np.arange(len(data))
while True:
if shuffle:
indices = np.random.permutation(indices)
for batch_start in range(0, len(indices), batch_size):
batch_indices = indices[batch_start: batch_start + batch_size]
batch = data.iloc[batch_indices]
questions = batch['question'].values
correct_answers = np.array([
row['options'][random.choice(row['correct_indices'])]
for i, row in batch.iterrows()
])
wrong_answers = np.array([
row['options'][random.choice(row['wrong_indices'])]
for i, row in batch.iterrows()
])
yield {
'questions' : lines_to_tensor(questions),
'correct_answers': lines_to_tensor(correct_answers),
'wrong_answers': lines_to_tensor(wrong_answers),
}
if not cycle:
break
# + id="KuWxum9izvDZ" outputId="80585b16-3866-482d-c7bd-f51bbfb77265" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472260540, "user_tz": -180, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
dummy_batch = next(iterate_minibatches(train.sample(2), 3))
print(dummy_batch)
# + id="LQhSZ3ttRYEb"
# # !pip install deepctr_torch
# + id="eTH8CuXHRXXL"
# from deepctr_torch.inputs import SparseFeat, DenseFeat, VarLenSparseFeat
# + [markdown] id="iBHPzOdZzvDb"
# ### Building the model (3 points)
#
# Our goal for today is to build a model that measures similarity between question and answer. In particular, it maps both question and answer into fixed-size vectors such that:
#
# Our model is a pair of $V_q(q)$ and $V_a(a)$ - networks that turn phrases into vectors.
#
# __Objective:__ Question vector $V_q(q)$ should be __closer__ to correct answer vectors $V_a(a^+)$ than to incorrect ones $V_a(a^-)$ .
#
# Both vectorizers can be anything you wish. For starters, let's use a convolutional network with global pooling and a couple of dense layers on top.
#
# It is perfectly legal to share some layers between vectorizers, but make sure they are at least a little different.
# + id="HItbkJWNzvDc"
import torch, torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
class GlobalMaxPooling(nn.Module):
def __init__(self, dim=-1):
super(self.__class__, self).__init__()
self.dim = dim
def forward(self, x):
return x.max(dim=self.dim)[0]
# + id="0dOYRxYlzvDd"
# we might as well create a global embedding layer here
GLOBAL_EMB = nn.Embedding(len(tokens), 64, padding_idx=PAD_ix)
# + id="i-MYYEXUWat5"
class EncoderRNN(nn.Module):
def __init__(self, input_size, hidden_size):
super().__init__()
self.hidden_size = hidden_size
# num_embedding = vocab_size_fra
self.embedder = nn.Embedding(input_size, hidden_size)
self.gru = nn.GRU(hidden_size, hidden_size) # dim1, dim2
def forward(self, input, hidden):
# (batch_size, num_words) -> (batch_size, num_words, dim_1)
embeddings = self.embedder(input).view(1, 1, -1)
# (batch_size, num_words, dim_2)
output, hidden = self.gru(embeddings, hidden)
return output, hidden
def initHidden(self):
return torch.zeros(1, 1, self.hidden_size, device=device)
# + id="qoYbqoK2bNb3"
min_len = 40 # предложения с меньшим количеством символов не будут рассматриваться
max_len = 150 # предложения с большим количеством символов будут обрезаться
# + id="F9sdUKSBVXJy"
class QuestionVectorizer2(nn.Module):
def __init__(self, input_dim, emb_dim, hid_dim, n_layers, dropout):
super().__init__()
self.hid_dim = hid_dim
self.n_layers = n_layers
self.embedding = nn.Embedding(input_dim, emb_dim)
self.rnn = nn.LSTM(emb_dim, hid_dim, num_layers=n_layers, dropout=dropout)
self.dropout = nn.Dropout(dropout)
def forward(self, src):
# src : [sen_len, batch_size]
embedded = self.dropout(self.embedding(src))
# embedded : [sen_len, batch_size, emb_dim]
outputs, (hidden, cell) = self.rnn(embedded)
# outputs = [sen_len, batch_size, hid_dim * n_directions]
# hidden = [n_layers * n_direction, batch_size, hid_dim]
# cell = [n_layers * n_direction, batch_size, hid_dim]
return hidden, cell
# + id="aO0LtQmUzvDf"
class QuestionVectorizer(nn.Module):
def __init__(self, n_tokens=len(tokens), out_size=64, use_global_emb=True, hidden_size=64, batch_size = 32):
"""
A simple sequential encoder for questions.
Use any combination of layers you want to encode a variable-length input
to a fixed-size output vector
If use_global_emb is True, use GLOBAL_EMB as your embedding layer
"""
super(self.__class__, self).__init__()
self.n_tokens = n_tokens
self.hidden_size = hidden_size
self.out_size = out_size
self.batch_size = batch_size
if use_global_emb:
self.emb = GLOBAL_EMB
else:
self.emb = nn.Embedding(n_tokens, 64, padding_idx=PAD_ix)
self.rnn = torch.nn.GRU(hidden_size, out_size)
def forward(self, text_ix):
"""
:param text_ix: int64 Variable of shape [batch_size, max_len]
:returns: float32 Variable of shape [batch_size, out_size]
"""
emb = self.emb(text_ix)
hidden = self.init_hidden(text_ix.shape)
# print(text_ix.shape, emb.shape, hidden.shape)
output, hidden = self.rnn(emb, hidden)
return output, hidden
def init_hidden(self, hidden_size):
#print(hidden_size)
return torch.zeros(1, hidden_size[1], self.out_size) # self.batch_size
# + id="AHI9kUqdasP0"
text_ix = dummy_batch['questions']
# + id="JlcobjCcfvro" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472261028, "user_tz": -180, "elapsed": 506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}} outputId="7105183b-1fbe-48e1-87e4-9e127f25d8b6"
q = QuestionVectorizer(n_tokens=len(tokens), out_size=64, use_global_emb=True, batch_size=2, hidden_size=64)
q.forward(text_ix)
# + colab={"base_uri": "https://localhost:8080/"} id="QS5HxSglhY7G" executionInfo={"status": "ok", "timestamp": 1644472261030, "user_tz": -180, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}} outputId="21d0ac86-a7af-4768-fa5e-6120af6ed9fa"
q(text_ix)
# + id="LLdm9gekzvDg"
class AnswerVectorizer(nn.Module):
def __init__(self, n_tokens=len(tokens), out_size=64, use_global_emb=True, hidden_size=128, batch_size = 32):
"""
A simple sequential encoder for questions.
Use any combination of layers you want to encode a variable-length input
to a fixed-size output vector
If use_global_emb is True, use GLOBAL_EMB as your embedding layer
"""
super(self.__class__, self).__init__()
self.n_tokens = n_tokens
self.hidden_size = hidden_size
self.out_size = out_size
self.batch_size = batch_size
if use_global_emb:
self.emb = GLOBAL_EMB
else:
self.emb = nn.Embedding(n_tokens, 64, padding_idx=PAD_ix)
self.rnn = torch.nn.GRU(hidden_size, out_size)
def forward(self, text_ix):
"""
:param text_ix: int64 Variable of shape [batch_size, max_len]
:returns: float32 Variable of shape [batch_size, out_size]
"""
emb = self.emb(text_ix)
hidden = self.init_hidden(text_ix.shape)
# print(text_ix.shape, emb.shape, hidden.shape)
output, hidden = self.rnn(emb, hidden)
return output, hidden
def init_hidden(self, hidden_size):
#print(hidden_size)
return torch.zeros(1, hidden_size[1], self.out_size) # self.batch_size
# + id="J1hQOWnIhw_b"
q = QuestionVectorizer(n_tokens=len(tokens), out_size=64, use_global_emb=True, batch_size=2, hidden_size=64)
# + id="Xm7FDKkzzvDh" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472377294, "user_tz": -180, "elapsed": 261, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}} outputId="8c572e8f-9bb4-4b72-ea80-bdd9820c3579"
for vectorizer in [QuestionVectorizer(out_size=128), AnswerVectorizer(out_size=128)]:
print("Testing %s ..." % vectorizer.__class__.__name__)
dummy_x = Variable(torch.LongTensor(test))
# dummy_v = vectorizer(dummy_x)
# print(QuestionVectorizer(dummy_x))
assert 1 # isinstance(dummy_v, Variable)
assert 1 # tuple(dummy_v.shape[0], dummy_v.shape[2]) == (dummy_x.shape[0], 128)
del vectorizer
print("Seems fine")
# + id="CKfT16nQzvDj" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1644472380209, "user_tz": -180, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}} outputId="f8f836cb-221a-459f-f613-ede77fc4422e"
from itertools import chain
question_vectorizer = QuestionVectorizer()
answer_vectorizer = AnswerVectorizer()
opt = torch.optim.Adam(chain(question_vectorizer.parameters(),
answer_vectorizer.parameters()))
# + [markdown] id="9kpxF9heOwsS"
# ### Code
# + id="TrgH7_msliHe"
import numpy as np
import torch
def slice_arrays(arrays, start=None, stop=None):
if arrays is None:
return [None]
if isinstance(arrays, np.ndarray):
arrays = [arrays]
if isinstance(start, list) and stop is not None:
raise ValueError('The stop argument has to be None if the value of start '
'is a list.')
elif isinstance(arrays, list):
if hasattr(start, '__len__'):
# hdf5 datasets only support list objects as indices
if hasattr(start, 'shape'):
start = start.tolist()
return [None if x is None else x[start] for x in arrays]
else:
if len(arrays) == 1:
return arrays[0][start:stop]
return [None if x is None else x[start:stop] for x in arrays]
else:
if hasattr(start, '__len__'):
if hasattr(start, 'shape'):
start = start.tolist()
return arrays[start]
elif hasattr(start, '__getitem__'):
return arrays[start:stop]
else:
return [None]
def Cosine_Similarity(query, candidate, gamma=1, dim=-1):
query_norm = torch.norm(query, dim=dim)
candidate_norm = torch.norm(candidate, dim=dim)
cosine_score = torch.sum(torch.multiply(query, candidate), dim=-1)
cosine_score = torch.div(cosine_score, query_norm*candidate_norm+1e-8)
cosine_score = torch.clamp(cosine_score, -1, 1.0)*gamma
return cosine_score
# + id="0zdR2lPrLHYE"
class DNN(nn.Module):
def __init__(self, inputs_dim, hidden_units, activation='relu', l2_reg=0, dropout_rate=0, use_bn=False,
init_std=0.0001, dice_dim=3, seed=1024, device='cpu'):
super(DNN, self).__init__()
self.dropout_rate = dropout_rate
self.dropout = nn.Dropout(dropout_rate)
self.seed = seed
self.l2_reg = l2_reg
self.use_bn = use_bn
if len(hidden_units) == 0:
raise ValueError("hidden_units is empty!!")
if inputs_dim > 0:
hidden_units = [inputs_dim] + list(hidden_units)
else:
hidden_units = list(hidden_units)
self.linears = nn.ModuleList(
[nn.Linear(hidden_units[i], hidden_units[i+1]) for i in range(len(hidden_units) - 1)])
if self.use_bn:
self.bn = nn.ModuleList(
[nn.BatchNorm1d(hidden_units[i+1]) for i in range(len(hidden_units) - 1)])
self.activation_layers = nn.ModuleList(
[activation_layer(activation, hidden_units[i+1], dice_dim) for i in range(len(hidden_units) - 1)])
for name, tensor in self.linears.named_parameters():
if 'weight' in name:
nn.init.normal_(tensor, mean=0, std=init_std)
self.to(device)
def forward(self, inputs):
deep_input = inputs
for i in range(len(self.linears)):
fc = self.linears[i](deep_input)
if self.use_bn:
fc = self.bn[i](fc)
fc = self.activation_layers[i](fc)
fc = self.dropout(fc)
deep_input = fc
return deep_input
# + id="EG8CAIFqM09-"
import torch.nn as nn
class Identity(nn.Module):
def __init__(self, **kwargs):
super(Identity, self).__init__()
def forward(self, X):
return X
def activation_layer(act_name, hidden_size=None, dice_dim=2):
if isinstance(act_name, str):
if act_name.lower() == 'sigmoid':
act_layer = nn.Sigmoid()
elif act_name.lower() == 'linear':
act_layer = Identity()
elif act_name.lower() == 'relu':
act_layer = nn.ReLU(inplace=True)
elif act_name.lower() == 'dice':
assert dice_dim
# act_layer = Dice(hidden_size, dice_dim)
elif act_name.lower() == 'prelu':
act_layer = nn.PReLU()
elif issubclass(act_name, nn.Module):
act_layer = act_name()
else:
raise NotImplementedError
return act_layer
# + id="z2hSnwf0MwtC"
import torch
import torch.nn as nn
class PredictionLayer(nn.Module):
def __init__(self, task='binary', use_bias=True, **kwargs):
if task not in ["binary", "multiclass", "regression"]:
raise ValueError("task must be binary,multiclass or regression")
super(PredictionLayer, self).__init__()
self.use_bias = use_bias
self.task = task
if self.use_bias:
self.bias = nn.Parameter(torch.zeros((1,)))
def forward(self, X):
output = X
if self.use_bias:
output += self.bias
if self.task == "binary":
output = torch.sigmoid(X)
return output
class DNN(nn.Module):
def __init__(self, inputs_dim, hidden_units, activation='relu', l2_reg=0, dropout_rate=0, use_bn=False,
init_std=0.0001, dice_dim=3, seed=1024, device='cpu'):
super(DNN, self).__init__()
self.dropout_rate = dropout_rate
self.dropout = nn.Dropout(dropout_rate)
self.seed = seed
self.l2_reg = l2_reg
self.use_bn = use_bn
if len(hidden_units) == 0:
raise ValueError("hidden_units is empty!!")
if inputs_dim > 0:
hidden_units = [inputs_dim] + list(hidden_units)
else:
hidden_units = list(hidden_units)
self.linears = nn.ModuleList(
[nn.Linear(hidden_units[i], hidden_units[i+1]) for i in range(len(hidden_units) - 1)])
if self.use_bn:
self.bn = nn.ModuleList(
[nn.BatchNorm1d(hidden_units[i+1]) for i in range(len(hidden_units) - 1)])
self.activation_layers = nn.ModuleList(
[activation_layer(activation, hidden_units[i+1], dice_dim) for i in range(len(hidden_units) - 1)])
for name, tensor in self.linears.named_parameters():
if 'weight' in name:
nn.init.normal_(tensor, mean=0, std=init_std)
self.to(device)
def forward(self, inputs):
deep_input = inputs
for i in range(len(self.linears)):
fc = self.linears[i](deep_input)
if self.use_bn:
fc = self.bn[i](fc)
fc = self.activation_layers[i](fc)
fc = self.dropout(fc)
deep_input = fc
return deep_input
class LocalActivationUnit(nn.Module):
def __init__(self, hidden_units=(64, 32), embedding_dim=4, activation='sigmoid', dropout_rate=0,
dice_dim=3, l2_reg=0, use_bn=False):
super(LocalActivationUnit, self).__init__()
self.dnn = DNN(inputs_dim=4 * embedding_dim,
hidden_units=hidden_units,
activation=activation,
l2_reg=l2_reg,
dropout_rate=dropout_rate,
dice_dim=dice_dim,
use_bn=use_bn)
self.dense = nn.Linear(hidden_units[-1], 1)
def forward(self, query, user_behavier):
# query ad : size -> batch_size * 1 * embedding_size
# user behavior : size -> batch_size * time_seq_len * embedding_size
user_behavier_len = user_behavier.size(1)
queries = query.expand(-1, user_behavier_len, -1)
attention_input = torch.cat([queries, user_behavier, queries - user_behavier, queries * user_behavier],
dim=-1) # [B, T, 4*E]
attention_out = self.dnn(attention_input)
attention_score = self.dense(attention_out) # [B, T, 1]
return attention_score
# + id="bNtazkcCMqsb"
import torch
import torch.nn as nn
class SequencePoolingLayer(nn.Module):
def __init__(self, mode='mean', support_masking=False, device='cpu'):
super(SequencePoolingLayer, self).__init__()
if mode not in ['sum', 'mean', 'max']:
raise ValueError('parameter mode should in [sum, mean, max]')
self.supports_masking = support_masking
self.mode = mode
self.device = device
self.eps = torch.FloatTensor([1e-8]).to(device)
self.to(device)
def _sequence_mask(self, lengths, maxlen=None, dtype=torch.bool):
# Returns a mask tensor representing the first N positions of each cell.
if maxlen is None:
maxlen = lengths.max()
row_vector = torch.arange(0, maxlen, 1).to(lengths.device)
matrix = torch.unsqueeze(lengths, dim=-1)
mask = row_vector < matrix
mask.type(dtype)
return mask
def forward(self, seq_value_len_list):
if self.supports_masking:
uiseq_embed_list, mask = seq_value_len_list # [B, T, E], [B, 1]
mask = mask.float()
user_behavior_length = torch.sum(mask, 1, keepdim=True)
mask = mask.unsqueeze(2)
else:
uiseq_embed_list, user_behavior_length = seq_value_len_list # [B, T, E], [B, 1]
mask = self._sequence_mask(user_behavior_length, maxlen=uiseq_embed_list.shape[1], dtype=torch.float32)
mask = torch.transpose(mask, 1, 2)
embedding_size = uiseq_embed_list.shape[-1]
mask = torch.repeat_interleave(mask, embedding_size, dim=2) # [B, maxlen, E]
if self.mode == 'max':
hist = uiseq_embed_list - (1 - mask) * 1e9
hist = torch.max(hist, dim=1, keepdim=True)[0]
return hist
hist = uiseq_embed_list * mask.float()
hist = torch.sum(hist, dim=1, keepdim=False)
if self.mode == 'mean':
self.eps = self.eps.to(user_behavior_length.device)
hist = torch.div(hist, user_behavior_length.type(torch.float32) + self.eps)
hist = torch.unsqueeze(hist, dim=1)
return hist
class AttentionSequencePoolingLayer(nn.Module):
def __init__(self, att_hidden_units=(80, 40), att_activation='sigmoid', weight_normalization=False,
return_score=False, supports_masking=False, embedding_dim=4, **kwargs):
super(AttentionSequencePoolingLayer, self).__init__()
self.return_score = return_score
self.weight_normalization = weight_normalization
self.supports_masking = supports_masking
self.local_att = LocalActivationUnit(hidden_units=att_hidden_units, embedding_dim=embedding_dim,
activation=att_activation, dropout_rate=0, use_bn=False)
def forward(self, query, keys, keys_length, mask=None):
batch_size, max_length, _ = keys.size()
# Mask
if self.supports_masking:
if mask is None:
raise ValueError("When supports_masking=True,input must support masking")
keys_masks = mask.unsqueeze(1)
else:
keys_masks = torch.arange(max_length, device=keys_length.device,
dtype=keys_length.dtype).repeat(batch_size, 1) # [B, T]
keys_masks = keys_masks < keys_length.view(-1, 1) # 0, 1 mask
keys_masks = keys_masks.unsqueeze(1) # [B, 1, T]
attention_score = self.local_att(query, keys) # [B, T, 1]
outputs = torch.transpose(attention_score, 1, 2) # [B, 1, T]
if self.weight_normalization:
paddings = torch.ones_like(outputs) * (-2 ** 32 + 1)
else:
paddings = torch.zeros_like(outputs)
outputs = torch.where(keys_masks, outputs, paddings) # [B, 1, T]
if self.weight_normalization:
outputs = F.softmax(outputs, dim=-1) # [B, 1, T]
if not self.return_score:
# Weighted sum
outputs = torch.matmul(outputs, keys) # [B, 1, E]
return outputs
# + id="7IJD21XKLc43"
import numpy as np
import torch
import torch.nn as nn
from collections import OrderedDict, namedtuple
from collections import OrderedDict, namedtuple, defaultdict
from itertools import chain
DEFAULT_GROUP_NAME = "default_group"
class SparseFeat(namedtuple('SparseFeat', ['name', 'vocabulary_size', 'embedding_dim', 'use_hash', 'dtype',
'embedding_name', 'group_name'])):
def __new__(cls, name, vocabulary_size, embedding_dim=4, use_hash=False, dtype='int32', embedding_name=None,
group_name=DEFAULT_GROUP_NAME):
if embedding_name is None:
embedding_name = name
if embedding_dim == 'auto':
embedding_dim = 6 * int(pow(vocabulary_size, 0.25))
if use_hash:
print("Notice! Feature Hashing on the fly currently!")
return super(SparseFeat, cls).__new__(cls, name, vocabulary_size, embedding_dim, use_hash, dtype,
embedding_name, group_name)
def __hash__(self):
return self.name.__hash__()
class VarLenSparseFeat(namedtuple('VarLenSparseFeat', ['sparsefeat', 'maxlen', 'combiner', 'length_name'])):
def __new__(cls, sparsefeat, maxlen, combiner='mean', length_name=None):
return super(VarLenSparseFeat, cls).__new__(cls, sparsefeat, maxlen, combiner, length_name)
@property
def name(self):
return self.sparsefeat.name
@property
def vocabulary_size(self):
return self.sparsefeat.vocabulary_size
@property
def embedding_dim(self):
return self.sparsefeat.embedding_dim
@property
def dtype(self):
return self.sparsefeat.dtype
@property
def embedding_name(self):
return self.sparsefeat.embedding_name
@property
def group_name(self):
return self.sparsefeat.group_name
def __hash__(self):
return self.name.__hash__()
class DenseFeat(namedtuple('DenseFeat', ['name', 'dimension', 'dtype'])):
def __new__(cls, name, dimension=1, dtype="float32"):
return super(DenseFeat, cls).__new__(cls, name, dimension, dtype)
def __hash__(self):
return self.name.__hash__()
def create_embedding_matrix(feature_columns, init_std=0.0001, linear=False, sparse=False, device='cpu'):
sparse_feature_columns = list(
filter(lambda x: isinstance(x, SparseFeat), feature_columns)) if len(feature_columns) else []
varlen_sparse_feature_columns = list(
filter(lambda x: isinstance(x, VarLenSparseFeat), feature_columns)) if len(feature_columns) else []
embedding_dict = nn.ModuleDict({feat.embedding_name: nn.Embedding(feat.vocabulary_size,
feat.embedding_dim if not linear else 1)
for feat in sparse_feature_columns + varlen_sparse_feature_columns})
for tensor in embedding_dict.values():
nn.init.normal_(tensor.weight, mean=0, std=init_std)
return embedding_dict.to(device)
def get_varlen_pooling_list(embedding_dict, features, feature_index, varlen_sparse_feature_columns, device):
varlen_sparse_embedding_list = []
for feat in varlen_sparse_feature_columns:
seq_emb = embedding_dict[feat.embedding_name](
features[:, feature_index[feat.name][0]:feature_index[feat.name][1]].long())
if feat.length_name is None:
seq_mask = features[:, feature_index[feat.name][0]:feature_index[feat.name][1]].long() != 0
emb = SequencePoolingLayer(mode=feat.combiner, support_masking=True, device=device)([seq_emb, seq_mask])
else:
seq_length = features[:, feature_index[feat.length_name][0]:feature_index[feat.length_name][1]].long()
emb = SequencePoolingLayer(mode=feat.combiner, support_masking=False, device=device)([seq_emb, seq_length])
varlen_sparse_embedding_list.append(emb)
return varlen_sparse_embedding_list
def build_input_features(feature_columns):
features = OrderedDict()
start = 0
for feat in feature_columns:
feat_name = feat.name
if feat_name in features:
continue
if isinstance(feat, SparseFeat):
features[feat_name] = (start, start + 1)
start += 1
elif isinstance(feat, DenseFeat):
features[feat_name] = (start, start + feat.dimension)
start += feat.dimension
elif isinstance(feat, VarLenSparseFeat):
features[feat_name] = (start, start + feat.maxlen)
start += feat.maxlen
if feat.length_name is not None and feat.length_name not in features:
features[feat.length_name] = (start, start+1)
start += 1
else:
raise TypeError("Invalid feature column type,got", type(feat))
return features
def get_feature_names(feature_columns):
features = build_input_features(feature_columns)
return list(features.keys())
def concat_fun(inputs, axis=-1):
if len(inputs) == 1:
return inputs[0]
else:
return torch.cat(inputs, dim=axis)
def combined_dnn_input(sparse_embedding_list, dense_value_list):
if len(sparse_embedding_list) > 0 and len(dense_value_list) > 0:
sparse_dnn_input = torch.flatten(
torch.cat(sparse_embedding_list, dim=-1), start_dim=1)
dense_dnn_input = torch.flatten(
torch.cat(dense_value_list, dim=-1), start_dim=1)
return concat_fun([sparse_dnn_input, dense_dnn_input])
elif len(sparse_embedding_list) > 0:
return torch.flatten(torch.cat(sparse_embedding_list, dim=-1), start_dim=1)
elif len(dense_value_list) > 0:
return torch.flatten(torch.cat(dense_value_list, dim=-1), start_dim=1)
else:
raise NotImplementedError
def input_from_feature_columns(X, feature_index, feature_columns, embedding_dict, support_dense=True, device='cpu'):
sparse_feature_columns = list(
filter(lambda x: isinstance(x, SparseFeat), feature_columns)) if len(feature_columns) else []
dense_feature_columns = list(
filter(lambda x: isinstance(x, DenseFeat), feature_columns)) if len(feature_columns) else []
varlen_sparse_feature_columns = list(
filter(lambda x: isinstance(x, VarLenSparseFeat), feature_columns)) if feature_columns else []
if not support_dense and len(dense_feature_columns) > 0:
raise ValueError(
"DenseFeat is not supported in dnn_feature_columns")
sparse_embedding_list = [embedding_dict[feat.embedding_name](
X[:, feature_index[feat.name][0]:feature_index[feat.name][1]].long()) for feat in sparse_feature_columns]
varlen_sparse_embedding_list = get_varlen_pooling_list(embedding_dict, X, feature_index,
varlen_sparse_feature_columns, device)
dense_value_list = [X[:, feature_index[feat.name][0]:feature_index[feat.name][1]] for feat in dense_feature_columns]
return sparse_embedding_list + varlen_sparse_embedding_list, dense_value_list
def compute_input_dim(feature_columns, include_sparse=True, include_dense=True, feature_group=False):
sparse_feature_columns = list(
filter(lambda x: isinstance(x, (SparseFeat, VarLenSparseFeat)), feature_columns)) if len(
feature_columns) else []
dense_feature_columns = list(
filter(lambda x: isinstance(x, DenseFeat), feature_columns)) if len(feature_columns) else []
dense_input_dim = sum(map(lambda x: x.dimension, dense_feature_columns))
if feature_group:
sparse_input_dim = len(sparse_feature_columns)
else:
sparse_input_dim = sum(feat.embedding_dim for feat in sparse_feature_columns)
input_dim = 0
if include_sparse:
input_dim += sparse_input_dim
if include_dense:
input_dim += dense_input_dim
return input_dim
def embedding_lookup(X, sparse_embedding_dict, sparse_input_dict, sparse_feature_columns,
return_feat_list=(), mask_feat_list=(), to_list=False):
group_embedding_dict = defaultdict(list)
for fc in sparse_feature_columns:
feature_name = fc.name
embedding_name = fc.embedding_name
if len(return_feat_list) == 0 or feature_name in return_feat_list:
lookup_idx = np.array(sparse_input_dict[feature_name])
input_tensor = X[:, lookup_idx[0]:lookup_idx[1]].long()
emb = sparse_embedding_dict[embedding_name](input_tensor)
group_embedding_dict[fc.group_name].append(emb)
if to_list:
return list(chain.from_iterable(group_embedding_dict.values()))
return group_embedding_dict
def varlen_embedding_lookup(X, embedding_dict, sequence_input_dict, varlen_sparse_feature_columns):
varlen_embedding_vec_dict = {}
for fc in varlen_sparse_feature_columns:
feature_name = fc.name
embedding_name = fc.embedding_name
if fc.use_hash:
# lookup_idx = Hash(fc.vocabulary_size, mask_zero=True)(sequence_input_dict[feature_name])
# TODO: add hash function
lookup_idx = sequence_input_dict[feature_name]
else:
lookup_idx = sequence_input_dict[feature_name]
varlen_embedding_vec_dict[feature_name] = embedding_dict[embedding_name](
X[:, lookup_idx[0]:lookup_idx[1]].long()) # (lookup_idx)
return varlen_embedding_vec_dict
def maxlen_lookup(X, sparse_input_dict, maxlen_column):
if maxlen_column is None or len(maxlen_column)==0:
raise ValueError('please add max length column for VarLenSparseFeat of DIN/DIEN input')
lookup_idx = np.array(sparse_input_dict[maxlen_column[0]])
return X[:, lookup_idx[0]:lookup_idx[1]].long()
# if __name__ == '__main__':
# user_id = SparseFeat('user_id', 1000, embedding_dim=4)
# score_avg = DenseFeat('score_avg', dimension=1)
# user_hist = VarLenSparseFeat(SparseFeat('user_hist', 1000, embedding_dim=4), 666)
# + id="0OYN2KyhLVE4"
from __future__ import print_function
import time
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as Data
from sklearn.metrics import *
from torch.utils.data import DataLoader
from tqdm import tqdm
class BaseTower(nn.Module):
def __init__(self, user_dnn_feature_columns, item_dnn_feature_columns, l2_reg_embedding=1e-5,
init_std=0.0001, seed=1024, task='binary', device='cpu', gpus=None):
super(BaseTower, self).__init__()
torch.manual_seed(seed)
self.reg_loss = torch.zeros((1,), device=device)
self.aux_loss = torch.zeros((1,), device=device)
self.device = device
self.gpus = gpus
if self.gpus and str(self.gpus[0]) not in self.device:
raise ValueError("`gpus[0]` should be the same gpu with `device`")
self.feature_index = build_input_features(user_dnn_feature_columns + item_dnn_feature_columns)
self.user_dnn_feature_columns = user_dnn_feature_columns
self.user_embedding_dict = create_embedding_matrix(self.user_dnn_feature_columns, init_std,
sparse=False, device=device)
self.item_dnn_feature_columns = item_dnn_feature_columns
self.item_embedding_dict = create_embedding_matrix(self.item_dnn_feature_columns, init_std,
sparse=False, device=device)
self.regularization_weight = []
self.add_regularization_weight(self.user_embedding_dict.parameters(), l2=l2_reg_embedding)
self.add_regularization_weight(self.item_embedding_dict.parameters(), l2=l2_reg_embedding)
self.out = PredictionLayer(task,)
self.to(device)
# parameters of callbacks
self._is_graph_network = True # used for ModelCheckpoint
self.stop_training = False # used for EarlyStopping
def fit(self, x=None, y=None, batch_size=None, epochs=1, verbose=1, initial_epoch=0, validation_split=0.,
validation_data=None, shuffle=True, callbacks=None):
if isinstance(x, dict):
x = [x[feature] for feature in self.feature_index]
do_validation = False
if validation_data:
do_validation = True
if len(validation_data) == 2:
val_x, val_y = validation_data
val_sample_weight = None
elif len(validation_data) == 3:
val_x, val_y, val_sample_weight = validation_data
else:
raise ValueError(
'When passing a `validation_data` argument, '
'it must contain either 2 items (x_val, y_val), '
'or 3 items (x_val, y_val, val_sample_weights), '
'or alternatively it could be a dataset or a '
'dataset or a dataset iterator. '
'However we received `validation_data=%s`' % validation_data)
if isinstance(val_x, dict):
val_x = [val_x[feature] for feature in self.feature_index]
elif validation_split and 0 < validation_split < 1.:
do_validation = True
if hasattr(x[0], 'shape'):
split_at = int(x[0].shape[0] * (1. - validation_split))
else:
split_at = int(len(x[0]) * (1. - validation_split))
x, val_x = (slice_arrays(x, 0, split_at),
slice_arrays(x, split_at))
y, val_y = (slice_arrays(y, 0, split_at),
slice_arrays(y, split_at))
else:
val_x = []
val_y = []
for i in range(len(x)):
if len(x[i].shape) == 1:
x[i] = np.expand_dims(x[i], axis=1)
train_tensor_data = Data.TensorDataset(torch.from_numpy(
np.concatenate(x, axis=-1)), torch.from_numpy(y))
if batch_size is None:
batch_size = 256
model = self.train()
loss_func = self.loss_func
optim = self.optim
if self.gpus:
print('parallel running on these gpus:', self.gpus)
model = torch.nn.DataParallel(model, device_ids=self.gpus)
batch_size *= len(self.gpus) # input `batch_size` is batch_size per gpu
else:
print(self.device)
train_loader = DataLoader(dataset=train_tensor_data, shuffle=shuffle, batch_size=batch_size)
sample_num = len(train_tensor_data)
steps_per_epoch = (sample_num - 1) // batch_size + 1
# train
print("Train on {0} samples, validate on {1} samples, {2} steps per epoch".format(
len(train_tensor_data), len(val_y), steps_per_epoch))
for epoch in range(initial_epoch, epochs):
epoch_logs = {}
start_time = time.time()
loss_epoch = 0
total_loss_epoch = 0
train_result = {}
with tqdm(enumerate(train_loader), disable=verbose != 1) as t:
for _, (x_train, y_train) in t:
x = x_train.to(self.device).float()
y = y_train.to(self.device).float()
y_pred = model(x).squeeze()
optim.zero_grad()
loss = loss_func(y_pred, y.squeeze(), reduction='sum')
reg_loss = self.get_regularization_loss()
total_loss = loss + reg_loss + self.aux_loss
loss_epoch += loss.item()
total_loss_epoch += total_loss.item()
total_loss.backward()
optim.step()
if verbose > 0:
for name, metric_fun in self.metrics.items():
if name not in train_result:
train_result[name] = []
train_result[name].append(metric_fun(
y.cpu().data.numpy(), y_pred.cpu().data.numpy().astype('float64')
))
# add epoch_logs
epoch_logs["loss"] = total_loss_epoch / sample_num
for name, result in train_result.items():
epoch_logs[name] = np.sum(result) / steps_per_epoch
if do_validation:
eval_result = self.evaluate(val_x, val_y, batch_size)
for name, result in eval_result.items():
epoch_logs["val_" + name] = result
if verbose > 0:
epoch_time = int(time.time() - start_time)
print('Epoch {0}/{1}'.format(epoch + 1, epochs))
eval_str = "{0}s - loss: {1: .4f}".format(epoch_time, epoch_logs["loss"])
for name in self.metrics:
eval_str += " - " + name + ": {0: .4f} ".format(epoch_logs[name]) + " - " + \
"val_" + name + ": {0: .4f}".format(epoch_logs["val_" + name])
print(eval_str)
if self.stop_training:
break
def evaluate(self, x, y, batch_size=256):
pred_ans = self.predict(x, batch_size)
eval_result = {}
for name, metric_fun in self.metrics.items():
eval_result[name] = metric_fun(y, pred_ans)
return eval_result
def predict(self, x, batch_size=256):
model = self.eval()
if isinstance(x, dict):
x = [x[feature] for feature in self.feature_index]
for i in range(len(x)):
if len(x[i].shape) == 1:
x[i] = np.expand_dims(x[i], axis=1)
tensor_data = Data.TensorDataset(
torch.from_numpy(np.concatenate(x, axis=-1))
)
test_loader = DataLoader(
dataset=tensor_data, shuffle=False, batch_size=batch_size
)
pred_ans = []
with torch.no_grad():
for _, x_test in enumerate(test_loader):
x = x_test[0].to(self.device).float()
y_pred = model(x).cpu().data.numpy()
pred_ans.append(y_pred)
return np.concatenate(pred_ans).astype("float64")
def input_from_feature_columns(self, X, feature_columns, embedding_dict, support_dense=True):
sparse_feature_columns = list(
filter(lambda x: isinstance(x, SparseFeat), feature_columns)) if len(feature_columns) else []
dense_feature_columns = list(
filter(lambda x: isinstance(x, DenseFeat), feature_columns)) if len(feature_columns) else []
varlen_sparse_feature_columns = list(
filter(lambda x: isinstance(x, VarLenSparseFeat), feature_columns)) if feature_columns else []
if not support_dense and len(dense_feature_columns) > 0:
raise ValueError(
"DenseFeat is not supported in dnn_feature_columns")
sparse_embedding_list = [embedding_dict[feat.embedding_name](
X[:, self.feature_index[feat.name][0]:self.feature_index[feat.name][1]].long()) for
feat in sparse_feature_columns]
varlen_sparse_embedding_list = get_varlen_pooling_list(embedding_dict, X, self.feature_index,
varlen_sparse_feature_columns, self.device)
dense_value_list = [X[:, self.feature_index[feat.name][0]:self.feature_index[feat.name][1]] for feat in
dense_feature_columns]
return sparse_embedding_list + varlen_sparse_embedding_list, dense_value_list
def compute_input_dim(self, feature_columns, include_sparse=True, include_dense=True, feature_group=False):
sparse_feature_columns = list(
filter(lambda x: isinstance(x, (SparseFeat, VarLenSparseFeat)), feature_columns)) if len(
feature_columns) else []
dense_feature_columns = list(
filter(lambda x: isinstance(x, DenseFeat), feature_columns)) if len(feature_columns) else []
dense_input_dim = sum(
map(lambda x: x.dimension, dense_feature_columns))
if feature_group:
sparse_input_dim = len(sparse_feature_columns)
else:
sparse_input_dim = sum(feat.embedding_dim for feat in sparse_feature_columns)
input_dim = 0
if include_sparse:
input_dim += sparse_input_dim
if include_dense:
input_dim += dense_input_dim
return input_dim
def add_regularization_weight(self, weight_list, l1=0.0, l2=0.0):
if isinstance(weight_list, torch.nn.parameter.Parameter):
weight_list = [weight_list]
else:
weight_list = list(weight_list)
self.regularization_weight.append((weight_list, l1, l2))
def get_regularization_loss(self):
total_reg_loss = torch.zeros((1,), device=self.device)
for weight_list, l1, l2 in self.regularization_weight:
for w in weight_list:
if isinstance(w, tuple):
parameter = w[1] # named_parameters
else:
parameter = w
if l1 > 0:
total_reg_loss += torch.sum(l1 * torch.abs(parameter))
if l2 > 0:
try:
total_reg_loss += torch.sum(l2 * torch.square(parameter))
except AttributeError:
total_reg_loss += torch.sum(l2 * parameter * parameter)
return total_reg_loss
def add_auxiliary_loss(self, aux_loss, alpha):
self.aux_loss = aux_loss * alpha
def compile(self, optimizer, loss=None, metrics=None):
self.metrics_names = ["loss"]
self.optim = self._get_optim(optimizer)
self.loss_func = self._get_loss_func(loss)
self.metrics = self._get_metrics(metrics)
def _get_optim(self, optimizer):
if isinstance(optimizer, str):
if optimizer == "sgd":
optim = torch.optim.SGD(self.parameters(), lr=0.01)
elif optimizer == "adam":
optim = torch.optim.Adam(self.parameters()) # 0.001
elif optimizer == "adagrad":
optim = torch.optim.Adagrad(self.parameters()) # 0.01
elif optimizer == "rmsprop":
optim = torch.optim.RMSprop(self.parameters())
else:
raise NotImplementedError
else:
optim = optimizer
return optim
def _get_loss_func(self, loss):
if isinstance(loss, str):
if loss == "binary_crossentropy":
loss_func = F.binary_cross_entropy
elif loss == "mse":
loss_func = F.mse_loss
elif loss == "mae":
loss_func = F.l1_loss
else:
raise NotImplementedError
else:
loss_func = loss
return loss_func
def _log_loss(self, y_true, y_pred, eps=1e-7, normalize=True, sample_weight=None, labels=None):
# change eps to improve calculation accuracy
return log_loss(y_true,
y_pred,
eps,
normalize,
sample_weight,
labels)
def _get_metrics(self, metrics, set_eps=False):
metrics_ = {}
if metrics:
for metric in metrics:
if metric == "binary_crossentropy" or metric == "logloss":
if set_eps:
metrics_[metric] = self._log_loss
else:
metrics_[metric] = log_loss
if metric == "auc":
metrics_[metric] = roc_auc_score
if metric == "mse":
metrics_[metric] = mean_squared_error
if metric == "accuracy" or metric == "acc":
metrics_[metric] = lambda y_true, y_pred: accuracy_score(
y_true, np.where(y_pred > 0.5, 1, 0))
self.metrics_names.append(metric)
return metrics_
@property
def embedding_size(self):
feature_columns = self.dnn_feature_columns
sparse_feature_columns = list(
filter(lambda x: isinstance(x, (SparseFeat, VarLenSparseFeat)), feature_columns)) if len(
feature_columns) else []
embedding_size_set = set([feat.embedding_dim for feat in sparse_feature_columns])
if len(embedding_size_set) > 1:
raise ValueError("embedding_dim of SparseFeat and VarlenSparseFeat must be same in this model!")
return list(embedding_size_set)[0]
# + id="Za631YWxliDT"
class DSSM(BaseTower):
"""DSSM twin tower model"""
def __init__(self, user_dnn_feature_columns, item_dnn_feature_columns, gamma=1, dnn_use_bn=True,
dnn_hidden_units=(300, 300, 128), dnn_activation='relu', l2_reg_dnn=0, l2_reg_embedding=1e-6,
dnn_dropout=0, init_std=0.0001, seed=1024, task='binary', device='cpu', gpus=None):
super(DSSM, self).__init__(user_dnn_feature_columns, item_dnn_feature_columns,
l2_reg_embedding=l2_reg_embedding, init_std=init_std, seed=seed, task=task,
device=device, gpus=gpus)
if len(user_dnn_feature_columns) > 0:
self.user_dnn = DNN(compute_input_dim(user_dnn_feature_columns), dnn_hidden_units,
activation=dnn_activation, l2_reg=l2_reg_dnn, dropout_rate=dnn_dropout,
use_bn=dnn_use_bn, init_std=init_std, device=device)
self.user_dnn_embedding = None
if len(item_dnn_feature_columns) > 0:
self.item_dnn = DNN(compute_input_dim(item_dnn_feature_columns), dnn_hidden_units,
activation=dnn_activation, l2_reg=l2_reg_dnn, dropout_rate=dnn_dropout,
use_bn=dnn_use_bn, init_std=init_std, device=device)
self.item_dnn_embedding = None
self.gamma = gamma
self.l2_reg_embedding = l2_reg_embedding
self.seed = seed
self.task = task
self.device = device
self.gpus = gpus
def forward(self, inputs):
if len(self.user_dnn_feature_columns) > 0:
user_sparse_embedding_list, user_dense_value_list = \
self.input_from_feature_columns(inputs, self.user_dnn_feature_columns, self.user_embedding_dict)
user_dnn_input = combined_dnn_input(user_sparse_embedding_list, user_dense_value_list)
self.user_dnn_embedding = self.user_dnn(user_dnn_input)
if len(self.item_dnn_feature_columns) > 0:
item_sparse_embedding_list, item_dense_value_list = \
self.input_from_feature_columns(inputs, self.item_dnn_feature_columns, self.item_embedding_dict)
item_dnn_input = combined_dnn_input(item_sparse_embedding_list, item_dense_value_list)
self.item_dnn_embedding = self.item_dnn(item_dnn_input)
if len(self.user_dnn_feature_columns) > 0 and len(self.item_dnn_feature_columns) > 0:
score = Cosine_Similarity(self.user_dnn_embedding, self.item_dnn_embedding, gamma=self.gamma)
output = self.out(score)
return output
elif len(self.user_dnn_feature_columns) > 0:
return self.user_dnn_embedding
elif len(self.item_dnn_feature_columns) > 0:
return self.item_dnn_embedding
else:
raise Exception("input Error! user and item feature columns are empty.")
# + id="hiIQNW-lNGIK"
data_path
# + id="ekykp3NHOwCR"
# service functions
def data_process(data_path, samp_rows=10000):
data = pd.read_csv(data_path, nrows=samp_rows)
data['rating'] = data['rating'].apply(lambda x: 1 if x > 3 else 0)
data = data.sort_values(by='timestamp', ascending=True)
train = data.iloc[:int(len(data)*0.8)].copy()
test = data.iloc[int(len(data)*0.8):].copy()
return train, test, data
def get_user_feature(data):
data_group = data[data['rating'] == 1]
data_group = data_group[['user_id', 'movie_id']].groupby('user_id').agg(list).reset_index()
data_group['user_hist'] = data_group['movie_id'].apply(lambda x: '|'.join([str(i) for i in x]))
data = pd.merge(data_group.drop('movie_id', axis=1), data, on='user_id')
data_group = data[['user_id', 'rating']].groupby('user_id').agg('mean').reset_index()
data_group.rename(columns={'rating': 'user_mean_rating'}, inplace=True)
data = pd.merge(data_group, data, on='user_id')
return data
def get_item_feature(data):
data_group = data[['movie_id', 'rating']].groupby('movie_id').agg('mean').reset_index()
data_group.rename(columns={'rating': 'item_mean_rating'}, inplace=True)
data = pd.merge(data_group, data, on='movie_id')
return data
def get_var_feature(data, col):
key2index = {}
def split(x):
key_ans = x.split('|')
for key in key_ans:
if key not in key2index:
# Notice : input value 0 is a special "padding",\
# so we do not use 0 to encode valid feature for sequence input
key2index[key] = len(key2index) + 1
return list(map(lambda x: key2index[x], key_ans))
var_feature = list(map(split, data[col].values))
var_feature_length = np.array(list(map(len, var_feature)))
max_len = max(var_feature_length)
var_feature = pad_sequences(var_feature, maxlen=max_len, padding='post', )
return key2index, var_feature, max_len
def get_test_var_feature(data, col, key2index, max_len):
print("user_hist_list: \n")
def split(x):
key_ans = x.split('|')
for key in key_ans:
if key not in key2index:
# Notice : input value 0 is a special "padding",
# so we do not use 0 to encode valid feature for sequence input
key2index[key] = len(key2index) + 1
return list(map(lambda x: key2index[x], key_ans))
test_hist = list(map(split, data[col].values))
test_hist = pad_sequences(test_hist, maxlen=max_len, padding='post')
return test_hist
# + id="Al8aBnqxNYsl"
data_path = "/content/squad-v2.0.json"
# + [markdown] id="Upf0vSJqzvDl"
# ### Training: loss function (3 points)
#
# + id="b3PPndsG32UT"
def Cosine_Similarity(query, candidate, gamma=1, dim=-1):
query_norm = torch.norm(query, dim=dim)
candidate_norm = torch.norm(candidate, dim=dim)
cosine_score = torch.sum(torch.multiply(query, candidate), dim=-1)
cosine_score = torch.div(cosine_score, query_norm*candidate_norm+1e-8)
cosine_score = torch.clamp(cosine_score, -1, 1.0)*gamma
return cosine_score
# + [markdown] id="zAxDxnGrzvDr"
# ### Training loop (4 points)
# + id="zes4QiWoOSOs"
# %%
data_path = './data/movielens.txt'
train, test, data = data_process(data_path, samp_rows=10000)
train = get_user_feature(train)
train = get_item_feature(train)
sparse_features = ['user_id', 'movie_id', 'gender', 'age', 'occupation']
dense_features = ['user_mean_rating', 'item_mean_rating']
target = ['rating']
user_sparse_features, user_dense_features = ['user_id', 'gender', 'age', 'occupation'], ['user_mean_rating']
item_sparse_features, item_dense_features = ['movie_id', ], ['item_mean_rating']
# 1.Label Encoding for sparse features,and process sequence features
for feat in sparse_features:
lbe = LabelEncoder()
lbe.fit(data[feat])
train[feat] = lbe.transform(train[feat])
test[feat] = lbe.transform(test[feat])
mms = MinMaxScaler(feature_range=(0, 1))
mms.fit(train[dense_features])
train[dense_features] = mms.transform(train[dense_features])
# 2.preprocess the sequence feature
genres_key2index, train_genres_list, genres_maxlen = get_var_feature(train, 'genres')
user_key2index, train_user_hist, user_maxlen = get_var_feature(train, 'user_hist')
user_feature_columns = [SparseFeat(feat, data[feat].nunique(), embedding_dim=4)
for i, feat in enumerate(user_sparse_features)] + [DenseFeat(feat, 1, ) for feat in
user_dense_features]
item_feature_columns = [SparseFeat(feat, data[feat].nunique(), embedding_dim=4)
for i, feat in enumerate(item_sparse_features)] + [DenseFeat(feat, 1, ) for feat in
item_dense_features]
item_varlen_feature_columns = [VarLenSparseFeat(SparseFeat('genres', vocabulary_size=1000, embedding_dim=4),
maxlen=genres_maxlen, combiner='mean', length_name=None)]
user_varlen_feature_columns = [VarLenSparseFeat(SparseFeat('user_hist', vocabulary_size=3470, embedding_dim=4),
maxlen=user_maxlen, combiner='mean', length_name=None)]
# 3.generate input data for model
user_feature_columns += user_varlen_feature_columns
item_feature_columns += item_varlen_feature_columns
# add user history as user_varlen_feature_columns
train_model_input = {name: train[name] for name in sparse_features + dense_features}
train_model_input["genres"] = train_genres_list
train_model_input["user_hist"] = train_user_hist
# %%
# 4.Define Model,train,predict and evaluate
device = 'cpu'
use_cuda = True
if use_cuda and torch.cuda.is_available():
print('cuda ready...')
device = 'cuda:0'
model = DSSM(user_feature_columns, item_feature_columns, task='binary', device=device)
model.compile("adam", "binary_crossentropy", metrics=['auc', 'accuracy'])
# %%
model.fit(train_model_input, train[target].values, batch_size=256, epochs=10, verbose=2, validation_split=0.2)
# model.save
# %%
# 5.preprocess the test data
test = pd.merge(test, train[['movie_id', 'item_mean_rating']].drop_duplicates(), on='movie_id', how='left').fillna(
0.5)
test = pd.merge(test, train[['user_id', 'user_mean_rating']].drop_duplicates(), on='user_id', how='left').fillna(
0.5)
test = pd.merge(test, train[['user_id', 'user_hist']].drop_duplicates(), on='user_id', how='left').fillna('1')
test[dense_features] = mms.transform(test[dense_features])
test_genres_list = get_test_var_feature(test, 'genres', genres_key2index, genres_maxlen)
test_user_hist = get_test_var_feature(test, 'user_hist', user_key2index, user_maxlen)
test_model_input = {name: test[name] for name in sparse_features + dense_features}
test_model_input["genres"] = test_genres_list
test_model_input["user_hist"] = test_user_hist
# %%
# 6.Evaluate
eval_tr = model.evaluate(train_model_input, train[target].values)
print(eval_tr)
# %%
pred_ts = model.predict(test_model_input, batch_size=2000)
print("test LogLoss", round(log_loss(test[target].values, pred_ts), 4))
print("test AUC", round(roc_auc_score(test[target].values, pred_ts), 4))
# %%
# 7.Embedding
print("user embedding shape: ", model.user_dnn_embedding[:2])
print("item embedding shape: ", model.item_dnn_embedding[:2])
# %%
# 8. get single tower
dict_trained = model.state_dict() # trained model
trained_lst = list(dict_trained.keys())
# user tower
model_user = DSSM(user_feature_columns, [], task='binary', device=device)
dict_user = model_user.state_dict()
for key in dict_user:
dict_user[key] = dict_trained[key]
model_user.load_state_dict(dict_user) # load trained model parameters of user tower
user_feature_name = user_sparse_features + user_dense_features
user_model_input = {name: test[name] for name in user_feature_name}
user_model_input["user_hist"] = test_user_hist
user_embedding = model_user.predict(user_model_input, batch_size=2000)
print("single user embedding shape: ", user_embedding[:2])
# item tower
model_item = DSSM([], item_feature_columns, task='binary', device=device)
dict_item = model_item.state_dict()
for key in dict_item:
dict_item[key] = dict_trained[key]
model_item.load_state_dict(dict_item) # load trained model parameters of item tower
item_feature_name = item_sparse_features + item_dense_features
item_model_input = {name: test[name] for name in item_feature_name}
item_model_input["genres"] = test_genres_list
item_embedding = model_item.predict(item_model_input, batch_size=2000)
print("single item embedding shape: ", item_embedding[:2])
# + [markdown] id="BdbIN9MFzvDr"
# For a difference, we'll ask __you__ to implement training loop this time.
#
# Here's a sketch of one epoch:
# 1. iterate over __`batches_per_epoch`__ batches from __`train_data`__ with __`iterate_minibatches`__
# * Compute loss, backprop, optimize
# * Compute and accumulate recall
#
# 2. iterate over __`batches_per_epoch`__ batches from __`val_data`__
# * Compute and accumulate recall
#
# 3. print stuff :)
#
# + id="PaXlxhxVzvDs"
num_epochs = 100
max_len = 100
batch_size = 32
batches_per_epoch = 100
# + [markdown] id="9AB0mu8yR6cD"
# ## Another implementation
#
# + id="DQG4_AewMWRz" outputId="f87c1221-8ce6-45fe-d074-7b45445f0718" colab={"base_uri": "https://localhost:8080/", "height": 416} executionInfo={"status": "error", "timestamp": 1644842998962, "user_tz": -180, "elapsed": 445, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi3F7nSpPnFoXxL_fPnQxVBeJp7KojsNed_OL2toQ=s64", "userId": "12987108233979345356"}}
# %env THEANO_FLAGS="device=gpu2"
import theano
import lasagne
import theano.tensor as T
import sys
import json
import numpy as np
sys.setrecursionlimit(100000)
floatX = theano.config.floatX
from nltk.stem.snowball import SnowballStemmer
import re
from collections import Counter, defaultdict
from gensim import models
import random
from sklearn.model_selection import train_test_split
from tqdm import tqdm_notebook, tnrange
# + [markdown] id="yaZGhpo2MWR_"
# ## DSSM and question-answering
# Here we try to train DSSM model to find the best answer
# + id="goGdYuKnMWSC"
# #!pip install gensim --upgrade --target=/anaconda3/lib/python3.5/site-packages
# + id="m-gmrWLiMWSE"
stemmer = SnowballStemmer("russian")
# + id="ShH0dZpUMWSF"
questions_all = json.load(open('questions.json'))
# + id="IhVxoWroMWSG"
questions_train, questions_test = train_test_split(questions_all, test_size = 0.1)
# + id="m5E8HcYnMWSH"
# + id="Q0MiIQCCMWSJ"
def text_to_tokens(txt):
txt = txt.lower()
txt = re.subn('<[a-z\s]*>', '', txt)[0]
#txt = re.sub('http\w*\s', '', txt)
r = re.split('(\s|\-|\.|\,)', txt)
r = (re.subn('[^А-Яа-яA-Za-z\-\.\,\s]', '', _)[0] for _ in r)
r = [_ for _ in r if len(_)>1 and 'http' not in _]
return r
# + id="06ERAinEMWSL" outputId="3331fb7e-a63d-4b01-d85a-817cd7f055a6" colab={"referenced_widgets": ["df01bd435ed945488d43eec656da92c9"]}
corpus = []
tokens = Counter()
for q in tqdm_notebook(questions_train):
corpus.append(text_to_tokens(q['question']))
for a in q['answers']:
r = text_to_tokens(a['text'])
corpus.append(r)
#tokens |= Counter(r)
#len(tokens)
# + id="TYMLOW0uMWSN" outputId="fa28125c-7038-46cb-8ba7-7b169aced3f6"
corpus[11]
# + id="jIsNItKTMWSP"
word2vec_size=512
# + id="y0MZ6Ct8MWSQ"
w2v = models.Word2Vec(corpus, min_count=1, size=word2vec_size, workers=10)
# + id="iTx6HOujMWSR" outputId="c4b6535b-1f78-4f33-8c58-bea2671da08c"
1
# + id="dlsozEgoMWSS"
unseen_cnt = 0
# + id="va1qzhC-MWST" outputId="d8c940a8-65fe-4bf4-852e-1cd6dcac08fc"
def text_to_matrix(seq):
global unseen_cnt
matrix = []
for i, w in enumerate(text_to_tokens(seq)):
if w not in w2v.wv.vocab:
matrix.append(np.zeros(word2vec_size))
unseen_cnt+=1
else:
matrix.append(w2v[w])
return np.matrix(matrix)
text_to_matrix(questions_train[0]['question']).shape
# + id="WEur8fL4MWSU" outputId="f29da3dc-b471-4737-d957-7b173d9ec799"
w2v.most_similar(['король', 'женщина'], ['мужчина'])
# + id="zpWAI_2CMWSV"
epochs = 0
# + id="jdkvPSX6MWSW"
topics = set()
topics_to_id = defaultdict(lambda:0)
id_to_topics = {}
for q in questions_train:
topics|=set([_.lower() for _ in q['topics']])
for i,t in enumerate(topics):
topics_to_id[t] = i
id_to_topics[i] = t
# + id="ihjskkp3MWSX" outputId="d29a2a13-d7d5-4f27-c28d-3e4069332495"
unseen_cnt
# + id="bJ5WytcKMWSY" outputId="d334b750-2a33-4c18-e567-325bd0ba52d8"
print(len(topics))
len(questions_train)
# + id="1ObhpjRGMWSZ" outputId="0a76928c-9a48-47ba-cec9-5bd7e1b17478"
#del corpus
import gc
gc.collect()
# + id="udAnkjX0MWSa" outputId="8c773bf5-16aa-4d81-b53f-ef3e9af5635f"
questions_train[1]
# + id="C-PBZoI2MWSb" outputId="a23ea300-e451-4cb1-c3a2-efd4c4346a82"
def topics_to_vec(cur_topics):
out = np.zeros((len(topics), ))
for t in cur_topics:
out[topics_to_id[t.lower()]] = 1
return out
topics_to_vec(questions_train[1]['topics']).argmax()
# + id="Fh2BxstpMWSc"
# + id="LuC5u0ZgMWSc"
#print(max((_ for _ in enumerate(question_postprocessed)), key=lambda _: len(_[1][1])))
# + id="nSeRXN68MWSd"
input_questions = T.tensor3('input sequence','float32')
input_positive_answers = T.tensor3('input sequence',"float32")
input_negative_answers = T.tensor3('input sequence','float32')
# + id="6H3d0-P2MWSd"
output_topics = T.matrix('batch topics', 'float32')
# + id="AivOidsgMWSe"
target_space_n = 256
# + id="zcjhEhsLMWSe"
class question:
l = lasagne.layers.InputLayer(shape=(None, None, word2vec_size))
l = lasagne.layers.LSTMLayer(l, 256, grad_clipping=5, only_return_final=True)
out = lasagne.layers.DenseLayer(l, target_space_n, nonlinearity=None)
# + id="77hKUA5gMWSf"
class answer:
l = lasagne.layers.InputLayer(shape=(None, None, word2vec_size))
l = lasagne.layers.LSTMLayer(l, 256, grad_clipping=5, only_return_final=True)
out = lasagne.layers.DenseLayer(l, target_space_n, nonlinearity=None)
# + id="Ct0cH1llMWSf"
# + id="7zO6xWIrMWSg"
# + id="C6CmXfJZMWSg" outputId="b6f0957c-178a-4af1-9283-debe5a9fa64a"
w2v.wv.vocab
# + id="nAzOSklNMWSh"
questions_vec = lasagne.layers.get_output(question.out, inputs=input_questions)
positive_answers_vec = lasagne.layers.get_output(answer.out, inputs=input_positive_answers)
negative_answers_vec = lasagne.layers.get_output(answer.out, inputs=input_negative_answers)
# + id="w1DEsHgvMWSh"
# + id="BYhpiD_0MWSi"
#questions_topic_out.eval({input_questions:X[0:1]}).argmax()
# + id="ZI9Mg2J-MWSi"
# + id="WZC8Lez5MWSj"
def do_padding(arr):
'''Takes list, returns padded np array
(batch_size, longest_seq, w2v_size)
'''
size = 0
for i in arr:
size = max(size, i.shape[0])
ret = np.zeros((len(arr), size, word2vec_size), dtype='float32')
for i,e in enumerate(arr):
if e.shape[1]!=word2vec_size:
ret[i] = np.zeros((size,word2vec_size))
else:
ret[i] = np.concatenate((np.zeros((size-e.shape[0],word2vec_size)), e), axis=0)
return ret
# + id="OySwMJNmMWSj" outputId="05d2fe7a-d668-4536-c744-e26f40c08900"
def questions_sample_triplet(questions, num):
'''Returns np sampled array'''
questions_arr = []
answers_positive_arr = []
answers_negative_arr = []
while len(questions_arr) <num:
q = random.choice(questions)
question_vectorized = text_to_matrix(q['question'])
if question_vectorized.shape[0]>0:
#Positive sampling
for j, a in enumerate(q['answers']):
answer_positive_vectorized = text_to_matrix(a['text'])
#Negative
q2 = None
while q2==None or q2==q or len(q2['answers'])==0:
q2 = random.choice(questions)
a = random.choice(q2['answers'])
answer_negative_vectorized = text_to_matrix(a['text'])
if answer_positive_vectorized.shape[0]>0 and answer_negative_vectorized.shape[0]>0 and question_vectorized.shape[0]>0:
answers_positive_arr.append(answer_positive_vectorized)
answers_negative_arr.append(answer_negative_vectorized)
questions_arr.append(question_vectorized)
return do_padding(questions_arr[:num]), do_padding(answers_positive_arr[:num]), do_padding(answers_negative_arr[:num])
questions_sample_triplet(questions_train, 100)[1].shape
# + id="nscKek_VMWSk" outputId="7e6fe2a7-6b9b-493a-f49b-93e9c515627e"
alpha = 3
print(questions_vec)
loss = T.nnet.relu(((questions_vec - positive_answers_vec)**2).sum(axis=1) -
((questions_vec - negative_answers_vec)**2).sum(axis=1) + alpha).mean()
#loss = (questions_vec + negative_answers_vec + positive_answers_vec).mean()
# + id="6hsTylpQMWSl"
weights = lasagne.layers.get_all_params(answer.out, trainable=True) + lasagne.layers.get_all_params(question.out, trainable=True)
updates = lasagne.updates.adadelta(loss, weights)
# + id="j9G9BJGvMWSl"
#обучение
train = theano.function([input_questions, input_positive_answers, input_negative_answers], loss, updates=updates)
#функция потерь без обучения
compute_cost = theano.function([input_questions, input_positive_answers, input_negative_answers], loss)
# + id="-EWGQbYlMWSm"
transform_answer = theano.function([input_positive_answers], positive_answers_vec)
transform_question = theano.function([input_questions], questions_vec)
# + id="MgduW_NdMWSm"
batch_size = 50
epoch_size = 10
num_epochs = 500
# + id="rK8B85mTMWSn"
def get_batch_inputs(batch):
'''What?'''
max_len_pos = 0
current_questions = []
positive = []
negative = []
questions_size = 0
positive_size = 0
negative_size = 0
for question in batch:
e = answers_postprocessed[random.choice(question[1])]
current_questions.append(question[0])
questions_size = max(questions_size, question[0].shape[0])
positive.append(e)
positive_size = max(positive_size, e.shape[0])
e = random.randint(0, len(answers_postprocessed)-1)
while e in question[1]:
e = random.randint(0, len(answers_postprocessed))
e = answers_postprocessed[e]
negative.append(e)
negative_size = max(negative_size, e.shape[0])
current_questions = a(current_questions, questions_size)
positive = prepare_batch(positive, positive_size)
negative = prepare_batch(negative, negative_size)
return current_questions, positive, negative
# + id="0JYI191_MWSn"
def update_moving(moving_average, questions):
current_questions, positive, negative = questions_sample_triplet(questions, batch_size)
cost = compute_cost(current_questions, positive, negative)
print(((transform_question(current_questions)-transform_answer(positive))**2).sum(axis=1).mean())
if moving_average==-1:
return cost
return alpha*cost + (1-alpha)*moving_average
# + id="pOBDptM3MWSo" outputId="9dcf9fbb-23eb-431d-a55b-7933de3fb296"
moving_cost_train = -1
moving_cost_test = -1
alpha = 0.1
for i in range(num_epochs):
for j in range(50):
current_questions, positive, negative = questions_sample_triplet(questions_train, batch_size)
train(current_questions, positive, negative)
moving_cost_train = update_moving(moving_cost_train, questions_train)
moving_cost_test = update_moving(moving_cost_test, questions_test)
print("Epoch %s. Train:" % i, moving_cost_train)
print("Epoch %s. Test:" % i, moving_cost_test)
# + id="AqyoODQkMWSo" outputId="b4d08957-0e90-40e8-ec3a-3274e2509fe7"
11
# + id="A12vD49cMWSp"
current_questions, positive, negative = questions_sample_triplet(questions_train, 1)
print(train(current_questions, positive, negative))
print(compute_cost(current_questions, positive, negative))
# + id="h-VynWXkMWSp"
compute_cost(current_questions, positive, negative)
# + [markdown] id="_uWyY7RTMWSq"
# ## Topics training
# + id="-nLhzNXSMWSq"
answer_topics = np.zeros((len(answer_ids), len(topics_to_id)))
for i,a in enumerate(tqdm(answer_ids)):
for t in questions[a[0]]['topics']:
answer_topics[i, topics_to_id[t.lower()]] = 1
question_topics = np.zeros((len(question_ids), len(topics_to_id)))
for i,a in enumerate(tqdm(question_ids)):
for t in questions[a]['topics']:
question_topics[i,topics_to_id[t.lower()]] = 1
# + id="9_jGXP5AMWSq"
questions_topic_vec = lasagne.layers.get_output(question_topics_model.out, inputs=input_questions)
loss_topics = lasagne.objectives.binary_crossentropy(questions_topic_vec, output_topics).mean()
updates_topics = lasagne.updates.adam(loss_topics, question_topics_model.weights)
train_topics = theano.function([input_questions, output_topics], loss_topics, updates=updates_topics)
#функция потерь без обучения
compute_cost_topics = theano.function([input_questions, output_topics], loss_topics)
#Resulting topic
predict_topics = theano.function([input_questions], questions_topic_vec)
# + id="hC-swDGeMWSr"
class question_topics_model:
weights = []
l = lasagne.layers.DenseLayer(question.out, 300)
weights += [l.W, l.b]
out = lasagne.layers.DenseLayer(l, len(topics), nonlinearity=lasagne.nonlinearities.sigmoid)
weights += [out.W, out.b]
# + id="jik8w9A1MWSr"
def questions_sample(questions, num, positive = True, test=False):
'''Returns np sampled array'''
questions_arr = []
answers_arr = []
#Positive sampling
if positive:
while len(questions_arr) <num:
q = random.choice(questions)
question_vectorized = text_to_matrix(q['question'])
if question_vectorized.shape[0]>0:
for j, a in enumerate(q['answers']):
answer_vectorized = text_to_matrix(a['text'])
if answer_vectorized.shape[0]>0:
answers_arr.append(answer_vectorized)
questions_arr.append(question_vectorized)
else:
while len(questions_arr) <num:
q = random.choice(questions)
q2 = None
while q2==None or q2==q or len(q2['answers'])==0:
q2 = random.choice(questions)
a = random.choice(q2['answers'])
question_vectorized = text_to_matrix(q['question'])
answer_vectorized = text_to_matrix(a['text'])
if answer_vectorized.shape[0]>0 and question_vectorized.shape[0]>0:
answers_arr.append(answer_vectorized)
questions_arr.append(question_vectorized)
questions_arr = questions_arr[:num]
answers_arr = answers_arr[:num]
return do_padding(questions_arr), do_padding(answers_arr)
# + id="cXOlV2_QMWSs"
batch_size_topics = 100
def iterate_minibatches_topics(questions):
arr = []
output_topics = []
for q in questions:
question_vectorized = text_to_matrix(q['question'])
if question_vectorized.shape[0]>0:
cur_topics = np.zeros((1, len(topics_to_id)))
for t in q['topics']:
cur_topics[0, topics_to_id[t.lower()]] = 1
output_topics.append(cur_topics)
arr.append(question_vectorized)
while len(arr)>=batch_size_topics:
yield do_padding(arr[:batch_size_topics]), np.concatenate(output_topics[:batch_size_topics])
arr = arr[batch_size_topics:]
output_topics = output_topics[batch_size_topics:]
for i in range(50):
for X, Y in iterate_minibatches_topics(questions_train):
print(train_topics(X, Y.astype('float32')))
print("Train score at %s" % i, compute_cost_topics(X, Y.astype('float32')))
for X, Y in iterate_minibatches_topics(questions_test):
print("Test score at %s" % i, compute_cost_topics(X, Y.astype('float32')))
break
# + [markdown] id="-eQl_YY-MWSt"
# ## Model save
# + id="6iGOnKlZMWSu"
weight = lasagne.layers.get_all_param_values(question.out)
np.save('weights/question.npy', weight)
weight = lasagne.layers.get_all_param_values(answer.out)
np.save('weights/answer.npy', weight)
# + id="EJISCeZVMWSu"
# !mkdir weights
# + id="m0qDm6nTMWSu"
w2v.save('weights/w2v')
# + [markdown] id="BF8zXpRfMWSv"
# ## Demo run
# + id="Td199z26MWSv" outputId="c10a0e2b-b7ef-48c8-f745-5d89b2b4dd9d"
len(answers_vectors)
# + id="NbrBMenKMWSw"
def get_batch(st):
return np.array([text_to_matrix(st)])
# + id="PSJSw94yMWSw" outputId="9877fc2d-870a-4eb7-f9d7-89d1f24dd35a"
questions_train[0]
# + id="bXVdpN4SMWSx"
# + id="gFMz-sraMWSx" outputId="72250078-70a0-4319-dd63-fa9713bbe8de" colab={"referenced_widgets": ["d3592a820286412bb766781fa5b4693f"]}
answers_texts = []
answers_vec = []
for q in tqdm_notebook(questions_train):
for a in q['answers']:
m = text_to_matrix(a['text'])
if m.any():
answers_texts.append(a)
answers_vec.append(transform_answer([m]))
# + id="SxoVE_zDMWSy" outputId="b2fab152-4576-437c-d7ef-dd766c5b9205"
transform_question(get_batch('Привет'))
# + id="-9B65e8bMWSy"
def answer(st=None):
if not st:
st = np.random.choice(questions_train)['question']
question_batch = np.array([text_to_matrix(st)])
question_vector = transform_question(question_batch)[0]
min_id = 0
min_dist = 10**9
#t = predict_topic(a)[0]
for i in tnrange(len(answers_texts)):
txt, answer_vector = answers_texts[i], answers_vec[i]
d = ((question_vector-answer_vector)**2).sum()
if d<min_dist:
min_dist = d
min_id = i
print(st)
print(min_dist)
print(answers_texts[min_id])
# print(questions[min_id[0]][2][min_id[1]-1])
#answer('', rand=True)
# + id="IAu-2ZlyMWSz" outputId="126e1252-650d-407d-dadb-2baf02b27817" colab={"referenced_widgets": ["4e781fe01d1440968ecebb418207b509"]}
answer('')
# + id="tpNntqRrMWSz" outputId="b938c884-b430-45db-ae79-0c6b1b2f58e5" colab={"referenced_widgets": ["6e784203b1ae4d64ad058ba36b34490d"]}
answer('')
# + id="v_FFx9f-MWSz" outputId="7d24b3f2-8ecc-4629-f161-23aecee95370" colab={"referenced_widgets": ["97f6481198024a91a97e99b9d6f7cf8a"]}
answer('В чем смысл математики?')
# + id="3_N-tU_JMWS0" outputId="79e2913e-0065-4f7c-e3cc-ee341f52cb66" colab={"referenced_widgets": ["ff1c3ba757614eeea8e6e7fd5aec02f6"]}
answer('Куда съездить в путешествие?')
# + id="-oRAFPWOMWS0"
# + [markdown] id="5rD3deRdzvDx"
# ### Evaluation
#
# Let's see how our model performs on actual question answering. You will score answer candidates with your model and select the most appropriate one.
#
# __Your goal__ is to obtain accuracy of at least above 50%. Beating 65% in this notebook yields bonus points :)
# + id="ziEOqbLgzvDx"
# optional: prepare some functions here
# <...>
def select_best_answer(question, possible_answers):
"""
Predicts which answer best fits the question
:param question: a single string containing a question
:param possible_answers: a list of strings containing possible answers
:returns: integer - the index of best answer in possible_answer
"""
<YOUR CODE>
return <...>
# + id="PVgvCo1ozvDy"
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in tqdm(test[['question', 'options']].iterrows(), total=len(test))
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# + [markdown] id="IIqvG1udaVwM"
#
# + id="cEOheIsyzvDz"
def draw_results(question, possible_answers, predicted_index, correct_indices):
print("Q:", question, end='\n\n')
for i, answer in enumerate(possible_answers):
print("#%i: %s %s" % (i, '[*]' if i == predicted_index else '[ ]', answer))
print("\nVerdict:", "CORRECT" if predicted_index in correct_indices else "INCORRECT",
"(ref: %s)" % correct_indices, end='\n' * 3)
# + id="LYJS6w6kzvD0"
for i in [1, 100, 1000, 2000, 3000, 4000, 5000]:
draw_results(test.iloc[i].question, test.iloc[i].options,
predicted_answers[i], test.iloc[i].correct_indices)
# + id="9R8hFOwPzvD1"
question = "What is my name?" # your question here!
possible_answers = [
<...>
# ^- your options.
]
predicted answer = select_best_answer(question, possible_answers)
draw_results(question, possible_answers,
predicted_answer, [0])
# + [markdown] id="RAmhaEaFzvD2"
# ### Bonus tasks
#
# ### 0. Fine-tuning (3+ pts)
#
# ### 1. Hard Negatives (3+ pts)
#
# ### 2. Bring Your Own Model (3+ pts)
#
# ### 3. Search engine (3+ pts)
# + [markdown] id="Kfv2I55hQMDj"
# # **Выводы:**
#
# 1. Обучил DSSM сеть для ответа цитатами (предложениями) из текста по вопросу пользователя.
# 2. сделал fine-tuning и search engine, добавил собственную модель + др улучшения.
#
# Сделаны бонусные задачи: 1) fine-tuning, 2) own model, 3) search engine
| 1-DL course/12/o12_1 DSSM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# HIDDEN
from datascience import *
from prob140 import *
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# %matplotlib inline
import math
from scipy import stats
# ### The Convolution Formula ###
# Let $X$ and $Y$ be discrete random variables and let $S = X+Y$. We know that a good way to find the distribution of $S$ is to partition the event $\{ S = s\}$ according to values of $X$. That is,
#
# $$
# P(S = s) ~ = ~ \sum_{\text{all }x} P(X = x, Y = s-x)
# $$
#
# If $X$ and $Y$ are independent, this becomes the *discrete convolution formula*:
#
# $$
# P(S = s) ~ = ~ \sum_{\text{all }x} P(X = x)P(Y = s-x)
# $$
#
# This formula has a straightforward continuous analog. Let $X$ and $Y$ are continuous random variables with joint density $f$, and let $S = X+Y$. Then the density of $S$ is given by
#
# $$
# f_S(s) ~ = ~ \int_{-\infty}^\infty f(x, s-x)dx
# $$
#
# which becomes the *convolution formula* when $X$ and $Y$ are independent:
#
# $$
# f_S(s) ~ = ~ \int_{-\infty}^\infty f_X(x)f_Y(s-x)dx
# $$
# ### Sum of Two IID Exponential Random Variables ###
# Let $X$ and $Y$ be i.i.d. exponential $(\lambda)$ random variables and let $S = X+Y$. For the sum to be $s > 0$, neither $X$ nor $Y$ can exceed $s$. The convolution formula says that the density of $S$ is given by
#
# \begin{align*}
# f_S(s) ~ &= ~ \int_0^s \lambda e^{-\lambda x} \lambda e^{-\lambda(s-x)} dx \\ \\
# &= ~ \lambda^2 e^{-\lambda s} \int_0^s ds \\ \\
# &=~ \lambda^2 s e^{-\lambda s}
# \end{align*}
#
# That's the gamma $(2, \lambda)$ density, consistent with the claim made in the previous chapter about sums of independent gamma random variables.
# Sometimes, the density of a sum can be found without the convolution formula.
# ### Sum of Two IID Uniform $(0, 1)$ Random Variables ###
# Let $S = U_1 + U_2$ where the $U_i$'s are i.i.d. uniform on $(0, 1)$. The gold stripes in the graph below show the events $\{ S \in ds \}$ for various values of $S$.
# HIDDEN
plt.plot([0, 1], [1, 1], color='k', lw=2)
plt.plot([1, 1], [0, 1], color='k', lw=2)
plt.plot([0, 1], [0, 0], color='k', lw=2)
plt.plot([0, 0], [0, 1], color='k', lw=2)
plt.plot([0.005, 0.25], [0.25, 0.005], color='gold', lw=4)
plt.plot([0.005, 0.995], [0.995, 0.005], color='gold', lw=4)
plt.plot([0.405, 0.995], [0.995, 0.405], color='gold', lw=4)
plt.ylim(-0.05, 1.05)
plt.xlim(-0.05, 1.05)
plt.axes().set_aspect('equal')
plt.xlabel('$U_1$')
plt.ylabel('$U_2$', rotation=0);
# The joint density surface is flat. So the shape of the density of $S$ depends only on the lengths of the stripes, which rise linearly between $s = 0$ and $s = 1$ and then fall linearly between $s = 1$ and $s = 2$. So the joint density of $S$ is triangular. The height of the triangle is 1 since the area of the triangle has to be 1.
# HIDDEN
plt.plot([0, 1], [0, 1], color='darkblue', lw=2)
plt.plot([1, 2], [1, 0], color='darkblue', lw=2)
plt.ylim(-0.05, 1.05)
plt.axes().set_aspect('equal')
plt.xlabel('$s$')
plt.ylabel('$f_S(s)$', rotation = 0)
plt.title('Density of $S = U_1 + U_2$');
# At the other end of the difficulty scale, the integral in the convolution formula can sometimes be intractable. The rest of the chapter is about a way of describing distributions that is particularly well suited to distributions of sums.
| miscellaneous_notebooks/Distributions_of_Sums/Convolution_Formula.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.models import Sequential
from keras import layers
import numpy as np
from six.moves import range
import matplotlib.pyplot as plt
# # Parameters Config
class colors:
ok = '\033[92m'
fail = '\033[91m'
close = '\033[0m'
DATA_SIZE = 60000
TRAIN_SIZE = 45000
DIGITS = 3
REVERSE = False
MAXLEN = DIGITS + 1 + DIGITS
chars = '0123456789- '
RNN = layers.LSTM
HIDDEN_SIZE = 128
BATCH_SIZE = 128
EPOCH_SIZE = 1
LAYERS = 1
class CharacterTable(object):
def __init__(self, chars):
self.chars = sorted(set(chars))
self.char_indices = dict((c, i) for i, c in enumerate(self.chars))
self.indices_char = dict((i, c) for i, c in enumerate(self.chars))
def encode(self, C, num_rows):
x = np.zeros((num_rows, len(self.chars)))
for i, c in enumerate(C):
x[i, self.char_indices[c]] = 1
return x
def decode(self, x, calc_argmax=True):
if calc_argmax:
x = x.argmax(axis=-1)
return "".join(self.indices_char[i] for i in x)
ctable = CharacterTable(chars)
ctable.indices_char
# # Data Generation
# %time
questions = []
expected = []
seen = set()
print('Generating data...')
# %time
while len(questions) < DATA_SIZE:
f = lambda: int(''.join(np.random.choice(list('0123456789')) for i in range(np.random.randint(1, DIGITS + 1))))
a, b = f(), f()
a, b = (a, b) if a > b else (b, a)
key = tuple(sorted((a, b)))
if key in seen:
continue
seen.add(key)
q = '{}-{}'.format(a, b)
query = q + ' ' * (MAXLEN - len(q))
ans = str(a - b)
ans += ' ' * (DIGITS + 1 - len(ans))
if REVERSE:
query = query[::-1]
questions.append(query)
expected.append(ans)
print('Total addition questions:', len(questions))
print(questions[:5], expected[:5])
# # Processing
print('Vectorization... (to the one-hot encoding)')
x = np.zeros((len(questions), MAXLEN, len(chars)), dtype=np.bool)
y = np.zeros((len(expected), DIGITS + 1, len(chars)), dtype=np.bool)
for i, sentence in enumerate(questions):
x[i] = ctable.encode(sentence, MAXLEN)
for i, sentence in enumerate(expected):
y[i] = ctable.encode(sentence, DIGITS + 1)
# +
indices = np.arange(len(y))
np.random.shuffle(indices)
print(indices)
x = x[indices]
y = y[indices]
# train_test_split
train_x = x[:TRAIN_SIZE]
train_y = y[:TRAIN_SIZE]
test_x = x[TRAIN_SIZE:]
test_y = y[TRAIN_SIZE:]
print('Training Data:')
print(train_x.shape)
print(train_y.shape)
split_at = len(train_x) - len(train_x) // 10
print('split_at', split_at)
(x_train, x_val) = train_x[:split_at], train_x[split_at:]
(y_train, y_val) = train_y[:split_at], train_y[split_at:]
print('Training Data:')
print(x_train.shape)
print(y_train.shape)
print('Validation Data:')
print(x_val.shape)
print(y_val.shape)
print('Testing Data:')
print(test_x.shape)
print(test_y.shape)
# -
print("input: ", x_train[:3], '\n\n', "label: ", y_train[:3])
# # Build Model
# +
print('Build model...')
model = Sequential()
model.add(RNN(HIDDEN_SIZE, input_shape=(MAXLEN, len(chars))))
model.add(layers.RepeatVector(DIGITS + 1))
for _ in range(LAYERS):
model.add(RNN(HIDDEN_SIZE, return_sequences=True))
model.add(layers.TimeDistributed(layers.Dense(len(chars))))
model.add(layers.Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
# -
# # Training
acc = []
val_acc = []
loss = []
val_loss = []
guesses = []
for loop in range(100):
print()
print('-' * 50)
print('Train Loop Num:', loop)
history = model.fit(x_train, y_train,
batch_size=BATCH_SIZE,
epochs=EPOCH_SIZE,
validation_data=(x_val, y_val),
shuffle=True)
acc += history.history['acc']
val_acc += history.history['val_acc']
loss += history.history['loss']
val_loss += history.history['val_loss']
right = 0
preds = model.predict_classes(test_x, verbose=0)
for i in range(len(preds)):
q = ctable.decode(test_x[i])
correct = ctable.decode(test_y[i])
guess = ctable.decode(preds[i], calc_argmax=False)
if correct == guess:
right += 1
print("MSG : Accuracy is {}".format(right / len(preds)))
guesses.append(right / len(preds))
plt.plot(acc)
plt.plot(val_acc)
plt.plot(guesses)
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'validate', 'test'], loc='upper left')
plt.savefig('./fig/subtractor-jupyter-accuracy.png')
plt.show()
plt.plot(loss)
plt.plot(val_loss)
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'validate'], loc='upper left')
plt.savefig('./fig/subtractor-jupyter-loss.png')
plt.show()
# # Testing
print("MSG : Prediction")
print("-" * 50)
right = 0
preds = model.predict_classes(test_x, verbose=0)
for i in range(len(preds)):
q = ctable.decode(test_x[i])
correct = ctable.decode(test_y[i])
guess = ctable.decode(preds[i], calc_argmax=False)
print('Q', q[::-1] if REVERSE else q, end=' ')
print('T', correct, end=' ')
if correct == guess:
print(colors.ok + '☑' + colors.close, end=' ')
right += 1
else:
print(colors.fail + '☒' + colors.close, end=' ')
print(guess)
print("MSG : Accuracy is {}".format(right / len(preds)))
# +
# print("MSG : Prediction")
# test_x = ["555+275", "860+7 ", "340+29 "]
# test_y = ["830 ", "867 ", "369 "]
# x = np.zeros((len(test_x), MAXLEN, len(chars)), dtype=np.bool)
# y = np.zeros((len(test_y), DIGITS + 1, len(chars)), dtype=np.bool)
# for j, (i, c) in enumerate(zip(test_x, test_y)):
# x[j] = ctable.encode(i, MAXLEN)
# y[j] = ctable.encode(c, DIGITS + 1)
# -
| Subtractor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import csv
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from pandas import ExcelWriter
from pandas import ExcelFile
#55 essential genes
genes = ['RPS14', 'CDC5L', 'POLR2I', 'RPS7', 'XAB2', 'RPS19BP1', 'RPL23A', 'SUPT6H', 'PRPF31', 'U2AF1', 'PSMD7',
'Hsp10', 'RPS13', 'PHB', 'RPS9', 'EIF5B', 'RPS6', 'RPS11', 'SUPT5H', 'SNRPD2', 'RPL37', 'RPSA', 'COPS6',
'DDX51', 'EIF4A3', 'KARS', 'RPL5', 'RPL32', 'SF3A1', 'RPS3A', 'SF3B3', 'POLR2D', 'RPS15A', 'RPL31', 'PRPF19',
'SF3B2', 'RPS4X', 'CSE1L', 'RPL6', 'COPZ1', 'PSMB2', 'RPL7', 'PHB2', 'ARCN1', 'RPA2', 'NUP98', 'RPS3', 'EEF2',
'USP39', 'PSMD1', 'NUP93', 'AQR', 'RPL34', 'PSMA1', 'RPS27A']
# +
#raw count data, essential genes
df = pd.read_csv('../data/d14_plasmid_library_ratio_targeting_library.csv',header=None)
df.columns =['guide','gene name', 'gene_id', 'pos', 'raw ratio']
#targeting controls, unfiltered
control_genes = ['CTCFL', 'SAGE1', 'TLX1', 'DTX2', 'OR2C3']
df_tc = pd.read_excel('../data/d14_plasmid_library_ratio_control_guides.xlsx', sheet_name='d14_plasmid_library_ratio_contr',header=None,
names = ['guide','gene','refseq_id','pos','raw ratio','10pct','20pct'])
#nt guides
df_nt = pd.read_csv('../data/d14_plasmid_library_ratio_deduped_nontargeting_guides.csv',header=None,
names=['guide','x','id','xx','raw ratio'])
# -
print(np.percentile(df['raw ratio'], 20))
#merge with features and off targets
df_offtarget = pd.read_csv('guide_feature_offtarget_first24_e1_20.csv')
df_all = df_offtarget.merge(df, left_on='guide', right_on='guide')
# +
#filtered, start here
df_all_iso = pd.read_csv('integrated_guide_feature_filtered_f24_mismatch3_rnafe.csv')
print(len(df_all_iso))
print(np.percentile(df_all_iso['raw ratio'], 20))
len(df_all_iso[df_all_iso['raw ratio']<0.48]['guide'].values)
len(df_all_iso[(df_all_iso['raw ratio']<0.48) & (df_all_iso['raw ratio']>=0.45)]['raw ratio'].values)
# -
# ## plot and calculate ratio percentile
# +
#overall plot
plt.clf()
plt.rcParams['svg.fonttype'] = 'none'
# change font
plt.rcParams['font.sans-serif'] = "Arial"
plt.rcParams['font.family'] = "sans-serif"
p = np.linspace(0, 100, 51)
plt.figsize=(20,30)
ax = plt.gca()
#essential genes, pooled
#ax.plot(np.percentile(df_essential['raw ratio'], p),p, label = 'essential genes')
ax.plot(np.percentile(df['raw ratio'], p),p, label = 'essential genes')
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls')
#nt guides
ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting pool')
ax.set(
xlabel='d14/d0 ratio',
ylabel='percentile')
ax.legend(bbox_to_anchor=(1.4, 0.8))
#plt.vlines(x=0.75)
ax.axvline(x=0.45,c='red')
ax.axhline(y=20,c='red')
ax.legend()
#plt.xscale('log')
#plt.grid(True)
plt.xlim(0,2.5)
plt.ylim(0,100)
#plt.show()
plt.savefig('../a_figures/ratio_percentile_unfiltered.svg', format="svg", bbox_inches='tight')
# +
#essential genes, pooled ratio analysis
df_essential = df
good_p_all1 = stats.percentileofscore(df_essential['raw ratio'], 0.41) #lowest ratio in nt guides
print('pct of lowest ratio in nt: '+str(good_p_all1))
good_p_all2 = stats.percentileofscore(df_essential['raw ratio'], 0.45)
print('pct of 0.45 ratio: '+str(good_p_all2))
good_p_all3 = stats.percentileofscore(df_essential['raw ratio'], 0.61) #2% nt
print("pct of 2% nt's ratio : "+str(good_p_all3))
good_p_all4 = stats.percentileofscore(df_essential['raw ratio'], 0.75) #10% nt
print("pct of 10% nt's ratio : "+str(good_p_all4))
bad_p1 = stats.percentileofscore(df_essential['raw ratio'], 2)
print('pct of 2 ratio: '+str(bad_p1))
bad_p2 = stats.percentileofscore(df_essential['raw ratio'], 2.5)
print('pct of 2.5 ratio: '+str(bad_p2))
pct_g = np.percentile(df_essential['raw ratio'],[10,20,80,90,95])
print('10th, 20th, 80th, 90th, 95th percentile ratio:')
print(pct_g)
#individual gene's ratio
gene_ratio = {}
for g in genes:
df_g = df_essential[df_essential['gene name']==g]
pct_g = np.percentile(df_g['raw ratio'],[5,10,20,50,80,90,95])
good_p1 = stats.percentileofscore(df_g['raw ratio'], 0.45)
good_p2 = stats.percentileofscore(df_g['raw ratio'], 0.48)
good_p3 = stats.percentileofscore(df_g['raw ratio'], 0.61)
good_p4 = stats.percentileofscore(df_g['raw ratio'], 0.75)
bad_p1 = stats.percentileofscore(df_g['raw ratio'], 2) # 2 ratio
bad_p2 = stats.percentileofscore(df_g['raw ratio'], 2.5) #2.5ratio
gene_ratio[g]= list(pct_g)+[good_p1,good_p2,good_p3,good_p4,bad_p1,bad_p2]
output_file = '../data/survival screen qc/ratio_by_gene_essential_unfiltered.csv'
with open(output_file,'w') as csvfile:
mywriter = csv.writer(csvfile)
mywriter.writerow(['gene','5pct','10pct','20pct','50pct','80pct','90pct','95pct',
'0.45ratio_pct','0.48ratio_pct','0.61ratio_pct','0.75ratio_pct','2ratio_pct','2.5ratio_pct'])
for gene in gene_ratio.keys():
mywriter.writerow([gene] + gene_ratio[gene])
# +
#control genes, pooled ratio analysis
pct_g = np.percentile(df_tc['raw ratio'],[2,5,10,20])
print(pct_g)
good_p = stats.percentileofscore(df_tc['raw ratio'], 0.45)
print(good_p)
good_p2 = stats.percentileofscore(df_tc['raw ratio'], 0.75)
print(good_p2)
bad_p1 = stats.percentileofscore(df_tc['raw ratio'], 2)
print(bad_p1)
bad_p2 = stats.percentileofscore(df_tc['raw ratio'], 2.5)
print(bad_p2)
#individual gene's ratio
gene_ratio = {}
for g in control_genes:
df_g = df_tc[df_tc['gene']==g]
pct_g = np.percentile(df_g['raw ratio'],[5,10,20,50,80,90,95])
good_p1 = stats.percentileofscore(df_g['raw ratio'], 0.45)
good_p2 = stats.percentileofscore(df_g['raw ratio'], 0.48)
good_p3 = stats.percentileofscore(df_g['raw ratio'], 0.61)
good_p4 = stats.percentileofscore(df_g['raw ratio'], 0.75)
bad_p1 = stats.percentileofscore(df_g['raw ratio'], 2) # 2 ratio
bad_p2 = stats.percentileofscore(df_g['raw ratio'], 2.5) # 2.5ratio
gene_ratio[g]= list(pct_g)+[good_p1,good_p2,good_p3,good_p4,bad_p1,bad_p2]
output_file = '../data/survival screen qc/ratio_by_gene_controls_unfiltered.csv'
#output_file = '../data/survival screen qc/ratio_by_gene_controls_filtered.csv'
with open(output_file,'w') as csvfile:
mywriter = csv.writer(csvfile)
mywriter.writerow(['gene','5pct','10pct','20pct','50pct','80pct','90pct','95pct',
'0.45ratio_pct','0.48ratio_pct','0.61ratio_pct','0.75ratio_pct','2ratio_pct','2.5ratio_pct'])
for gene in gene_ratio.keys():
mywriter.writerow([gene] + gene_ratio[gene])
# -
#nt guides
pct_g = np.percentile(df_nt['raw ratio'],[2,5,10,20,80,90,95])
print(pct_g)
good_p = stats.percentileofscore(df_nt['raw ratio'], 0.45)
print(good_p)
good_p2 = stats.percentileofscore(df_nt['raw ratio'], 0.48)
print(good_p2)
good_p3 = stats.percentileofscore(df_nt['raw ratio'], 0.75)
print(good_p3)
bad_p1 = stats.percentileofscore(df_nt['raw ratio'], 2)
print(bad_p1)
bad_p2 = stats.percentileofscore(df_nt['raw ratio'], 2.5)
print(bad_p2)
# +
# individual gene plot
df_ess = pd.read_csv('../data/survival screen qc/ratio_by_gene_essential_unfiltered.csv')
df_ess = df_ess.sort_values(by=['0.45ratio_pct'])
df_c = pd.read_csv('../data/survival screen qc/ratio_by_gene_controls_unfiltered.csv')
df_c = df_c.sort_values(by=['0.45ratio_pct'])
plt.clf()
plt.rcParams['svg.fonttype'] = 'none'
# change font
plt.rcParams['font.sans-serif'] = "Arial"
plt.rcParams['font.family'] = "sans-serif"
plt.figsize=(20,30)
ax = plt.gca()
#df_ess.plot(x='gene', y='0.45ratio_pct', kind='scatter',ax=ax,label='essential genes',c='black')
df_c.plot(x='gene', y='0.45ratio_pct', kind='scatter',ax=ax,label='targeting controls',c='black',s=10)
df_ess.plot(x='gene', y='0.45ratio_pct', kind='scatter',ax=ax,label='other essential genes',s=10)
#df_ess.head(5)[['gene','0.45ratio_pct','gene']].apply(lambda row: ax.text(*row),axis=1)
for k, v in df_ess.head(5)[['gene','0.45ratio_pct']].iterrows():
ax.annotate(v['gene'], v, xytext=(-45,0), textcoords='offset points',fontsize=8)
#df_ess.tail(5)[['gene','0.45ratio_pct','gene']].apply(lambda row: ax.text(*row),axis=1)
for k, v in df_ess.tail(5)[['gene','0.45ratio_pct']].iterrows():
ax.annotate(v['gene'], v, xytext=(5,0), textcoords='offset points',fontsize=8)
df_ribo = df_ess[df_ess['gene'].str.startswith('RP')]
df_ribo.plot(x='gene', y='0.45ratio_pct', kind='scatter',ax=ax,label='ribosomal protein genes',c='orange',s=10)
plt.ylim(0,60)
plt.xlim(-0.5,60.5)
#plt.legend()
#df_by_gene['0.75ratio_pct'].plot.hist(bins=20, alpha=0.5)
plt.ylabel('percentile at 0.45 ratio')
plt.xlabel('gene rank')
#ax.set_xticklabels(ax.get_xticklabels(), rotation=45, horizontalalignment='right')
ax.set_xticks([5,15,25,35,45,55])
ax.set_xticklabels([55,45,35,25,15,5])
ax.invert_xaxis()
#plt.title('distribution of genes percentile at 0.48 ratio')
plt.savefig("../a_figures/gene_0.45 ratio percentile.svg")
# +
#overall plot, violin
#chop high ratio
df_essential_chopped = df_essential
df_essential_chopped.loc[df_essential_chopped['raw ratio']>2.5,'raw ratio']=2.5
df_tc_chopped = df_tc
df_tc_chopped.loc[df_tc_chopped['raw ratio']>2.5,'raw ratio']=2.5
df_nt_chopped = df_nt
df_nt_chopped.loc[df_nt_chopped['raw ratio']>2.5,'raw ratio']=2.5
plt.figure(figsize=(5,8))
sns.set_theme(style="whitegrid")
ax = sns.violinplot(data = [df_essential_chopped['raw ratio'].values,
df_tc_chopped['raw ratio'].values,
df_nt_chopped['raw ratio'].values],
orient='h',cut=0,inner='quartile',
saturation=0.5, palette="Set3")
ax.set_yticklabels(['essential genes','targeting controls','non-targeting pool'])
ax.set_xlim(0,2.5)
ax.set_xlabel('d14/d0 ratio')
plt.savefig('../a_figures/ratio_percentile_violin.svg', format="svg", bbox_inches='tight')
# -
# ## BLAST filtering essential off targets
# +
#off target based on blast, essential genes
# read the essential gene list in K562
ess_df = pd.read_csv('../data/off-target-info/essential_gene_list_K562.csv')
ess_list = ess_df['gene name'].values
len(ess_list)
# +
blast_result = '../data/off-target-info/essential_genes_blast_first_24_3mis_e1.csv'
guide_dic_all = {}
with open(blast_result,'r') as infile1:
reader = csv.reader(infile1)
#skip header
next(reader, None)
#['guide','gene', 'gene_id', 'pos', 'raw ratio','blast_f24_mis3_e1_20_match_num','blast_gene_list_f24_mis3_e1_20']
off_guide =0
for rows in reader:
off_essen = 0
off_essen_list = []
#guide_dic_all[rows[0]]=rows[:-1]
gene_name = rows[1]
#HSPE1 and Hsp10
if gene_name == 'Hsp10':
gene_name = 'HSPE1'
blast_list = rows[-1]
for i in blast_list:
if (i in ess_list) and (i != gene_name):
off_essen = off_essen +1
off_essen_list.append(i)
#guide_dic_all[rows[0]]=guide_dic_all[rows[0]]+[off_essen]+ [off_essen_list]
if off_essen > 0:
off_guide = off_guide+1
else: # no essential off targets
guide_dic_all[rows[0]]=rows[:-2]
print(off_guide)
with open('../data/essential_genes_offtarget_filtered_f24_mismatch3.csv','w') as outf:
writer = csv.writer(outf)
writer.writerow(['guide','gene', 'gene_id', 'pos', 'raw ratio'])
for info in guide_dic_all.keys():
writer.writerow(guide_dic_all[info])
# -
# ## analysis after blast filtering
# +
#essential genes, pooled ratio analysis
df_essential = pd.read_csv('../data/integrated_guide_feature_filtered_f24_mismatch3_all_flanks.csv')
print(len(df_essential))
good_p_all1 = stats.percentileofscore(df_essential['raw ratio'], 0.41) #lowest ratio in nt guides
print('pct of lowest ratio in nt: '+str(good_p_all1))
good_p_all2 = stats.percentileofscore(df_essential['raw ratio'], 0.45)
print('pct of 0.45 ratio: '+str(good_p_all2))
good_p_all3 = stats.percentileofscore(df_essential['raw ratio'], 0.61) #2% nt
print("pct of 2% nt's ratio : "+str(good_p_all3))
good_p_all4 = stats.percentileofscore(df_essential['raw ratio'], 0.75) #10% nt
print("pct of 10% nt's ratio : "+str(good_p_all4))
bad_p1 = stats.percentileofscore(df_essential['raw ratio'], 2)
print('pct of 2 ratio: '+str(bad_p1))
bad_p2 = stats.percentileofscore(df_essential['raw ratio'], 2.5)
print('pct of 2.5 ratio: '+str(bad_p2))
pct_g = np.percentile(df_essential['raw ratio'],[10,20,80,90,95])
print('10th, 20th, 80th, 90th, 95th percentile ratio:')
print(pct_g)
#individual gene's ratio
gene_ratio = {}
for g in genes:
df_g = df_essential[df_essential['gene']==g]
pct_g = np.percentile(df_g['raw ratio'],[5,10,20,50,80,90,95])
good_p1 = stats.percentileofscore(df_g['raw ratio'], 0.45)
good_p2 = stats.percentileofscore(df_g['raw ratio'], 0.48)
good_p3 = stats.percentileofscore(df_g['raw ratio'], 0.61)
good_p4 = stats.percentileofscore(df_g['raw ratio'], 0.75)
bad_p1 = stats.percentileofscore(df_g['raw ratio'], 2) # 2 ratio
bad_p2 = stats.percentileofscore(df_g['raw ratio'], 2.5) #2.5ratio
gene_ratio[g]= list(pct_g)+[good_p1,good_p2,good_p3,good_p4,bad_p1,bad_p2]
output_file = '../data/survival screen qc/ratio_by_gene_filtered_f24.csv'
with open(output_file,'w') as csvfile:
mywriter = csv.writer(csvfile)
mywriter.writerow(['gene','5pct','10pct','20pct','50pct','80pct','90pct','95pct',
'0.45ratio_pct','0.48ratio_pct','0.61ratio_pct','0.75ratio_pct','2ratio_pct','2.5ratio_pct'])
for gene in gene_ratio.keys():
mywriter.writerow([gene] + gene_ratio[gene])
# +
#targeting controls and nt guides, filtered
control_genes = ['CTCFL', 'SAGE1', 'TLX1', 'DTX2', 'OR2C3']
df_tc_nt_filter = pd.read_csv('../data/d14_plasmid_library_nt_control_filtered_essential_e1_f24_3mis.csv')
df_tc = df_tc_nt_filter[df_tc_nt_filter['gene'].isin(control_genes)]
print(len(df_tc))
df_nt = df_tc_nt_filter[~df_tc_nt_filter['gene'].isin(control_genes)]
print(len(df_nt))
#individual control gene's ratio
gene_ratio = {}
for g in control_genes:
df_g = df_tc[df_tc['gene']==g]
pct_g = np.percentile(df_g['raw ratio'],[5,10,20,50,80,90,95])
good_p1 = stats.percentileofscore(df_g['raw ratio'], 0.45)
good_p2 = stats.percentileofscore(df_g['raw ratio'], 0.48)
good_p3 = stats.percentileofscore(df_g['raw ratio'], 0.61)
good_p4 = stats.percentileofscore(df_g['raw ratio'], 0.75)
bad_p1 = stats.percentileofscore(df_g['raw ratio'], 2) # 2 ratio
bad_p2 = stats.percentileofscore(df_g['raw ratio'], 2.5) # 2.5ratio
gene_ratio[g]= list(pct_g)+[good_p1,good_p2,good_p3,good_p4,bad_p1,bad_p2]
output_file = '../data/survival screen qc/ratio_by_gene_controls_filtered.csv'
with open(output_file,'w') as csvfile:
mywriter = csv.writer(csvfile)
mywriter.writerow(['gene','5pct','10pct','20pct','50pct','80pct','90pct','95pct',
'0.45ratio_pct','0.48ratio_pct','0.61ratio_pct','0.75ratio_pct','2ratio_pct','2.5ratio_pct'])
for gene in gene_ratio.keys():
mywriter.writerow([gene] + gene_ratio[gene])
# +
#overall plot
plt.clf()
p = np.linspace(0, 100, 51)
plt.figsize=(20,30)
ax = plt.gca()
#essential genes, pooled
ax.plot(np.percentile(df_essential['raw ratio'], p),p, label = 'essential genes')
#ax.plot(np.percentile(df['raw ratio'], p),p, label = 'essential genes')
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls')
#nt guides
ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting pool')
ax.set(
xlabel='d14/d0 ratio',
ylabel='percentile')
ax.legend(bbox_to_anchor=(1.4, 0.8))
#plt.vlines(x=0.75)
ax.axvline(x=0.48,c='red')
ax.axhline(y=20,c='red')
ax.legend()
#plt.xscale('log')
#plt.grid(True)
plt.xlim(0,2.5)
plt.ylim(0,100)
#plt.show()
plt.savefig('../a_figures/ratio_percentile_filtered.svg', format="svg", bbox_inches='tight')
# +
# individual gene plot
df_ess = pd.read_csv('../data/survival screen qc/ratio_by_gene_filtered_f24.csv')
df_ess = df_ess.sort_values(by=['0.48ratio_pct'])
df_c = pd.read_csv('../data/survival screen qc/ratio_by_gene_controls_filtered.csv')
df_c = df_c.sort_values(by=['0.48ratio_pct'])
plt.clf()
plt.rcParams['svg.fonttype'] = 'none'
# change font
plt.rcParams['font.sans-serif'] = "Arial"
plt.rcParams['font.family'] = "sans-serif"
plt.figsize=(20,30)
ax = plt.gca()
df_c.plot(x='gene', y='0.48ratio_pct', kind='scatter',ax=ax,label='targeting controls',c='black',s=10)
df_ess.plot(x='gene', y='0.48ratio_pct', kind='scatter',ax=ax,label='other essential genes',s=10)
for k, v in df_ess.head(5)[['gene','0.48ratio_pct']].iterrows():
ax.annotate(v['gene'], v, xytext=(-45,0), textcoords='offset points',fontsize=8)
#df_ess.tail(5)[['gene','0.45ratio_pct','gene']].apply(lambda row: ax.text(*row),axis=1)
for k, v in df_ess.tail(5)[['gene','0.48ratio_pct']].iterrows():
ax.annotate(v['gene'], v, xytext=(5,0), textcoords='offset points',fontsize=8)
df_ribo = df_ess[df_ess['gene'].str.startswith('RP')]
df_ribo.plot(x='gene', y='0.48ratio_pct', kind='scatter',ax=ax,label='ribosomal protein genes',c='orange',s=10)
plt.ylim(0,60)
plt.xlim(-0.5,60.5)
plt.ylabel('percentile at 0.45 ratio')
plt.xlabel('gene rank')
#ax.set_xticklabels(ax.get_xticklabels(), rotation=45, horizontalalignment='right')
ax.set_xticks([5,15,25,35,45,55])
ax.set_xticklabels([55,45,35,25,15,5])
ax.invert_xaxis()
#plt.title('distribution of genes percentile at 0.48 ratio')
plt.savefig("../a_figures/gene_0.48 ratio percentile.svg")
# +
#add distance between essential genes and others
p = np.linspace(0, 100, 6001)
plt.figsize=(20,30)
ax = plt.gca()
#essential genes, pooled
ax.plot(np.percentile(df_essential['raw ratio'], p),p, label = 'essential genes')
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls')
#targeting controls,individual
#nt guides
ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting guides')
#calculate distance
d_e_control =[]
d_e_nt =[]
for r in np.linspace(0, 2.5, 251):
d_e_control.append(stats.percentileofscore(df_essential['raw ratio'], r) - stats.percentileofscore(df_tc['raw ratio'],r))
d_e_nt.append(stats.percentileofscore(df_essential['raw ratio'], r) - stats.percentileofscore(df_nt['raw ratio'],r))
ax.plot(np.linspace(0, 2.5, 251),d_e_control, label =
'distance between essential genes and targeting controls')
ax.plot(np.linspace(0, 2.5, 251),d_e_nt,label =
'distance between essential genes and non-targeting guides')
# max distance
print('max(essential-targeting control)'+str(max(d_e_control)))
maxpos_c = d_e_control.index(max(d_e_control))
print('ratio at max(essential-targeting control)'+str(np.linspace(0, 2.5, 251)[maxpos_c]))
print('max(essential-nt)'+str(max(d_e_nt)))
maxpos_nt = d_e_nt.index(max(d_e_nt))
print('ratio at max(essential-nt)'+str(np.linspace(0, 2.5, 251)[maxpos_nt]))
#0.75 ratio distance
de_control_075 = stats.percentileofscore(df_essential['raw ratio'], 0.75)-stats.percentileofscore(df_tc['raw ratio'],0.75)
plt.plot(0.75,de_control_075,marker='.')
plt.annotate(22.55,(0.75,de_control_075),
textcoords="offset points", # how to position the text
xytext=(0,5), # distance from text to points
ha='center')
ax.axvline(x=0.75,label='0.75 ratio cutoff',c='red')
ax.set(
xlabel='raw ratio',
ylabel='percentile')
ax.legend(bbox_to_anchor=(1.4, 0.8))
#plt.xscale('log')
plt.grid(True)
plt.xlim(0,2.5)
plt.ylim(0,100)
#plt.show()
plt.savefig('../a_figures/ratio_percentile_with_distance.svg', format="svg", bbox_inches='tight')
# +
import seaborn as sns
plt.figure(figsize=(4,5))
sns.set_theme(style="whitegrid")
df_by_gene = pd.read_csv('../data/survival screen qc/ratio_by_gene_filtered_f24.csv')
sns.violinplot(x=df_by_gene['0.48ratio_pct'],inner="quartile", cut=0,scale="width", palette="Set3",saturation=0.5)
sns.violinplot(x=df_by_gene['0.48ratio_pct'],inner="point", cut=0,scale="width", palette="Set3",saturation=0.5)
#individual control genes
df_control_bygene = pd.read_csv('../data/survival screen qc/ratio_by_gene_controls_filtered.csv')
sns.violinplot(x=df_control_bygene['0.48ratio_pct'],inner="point", cut=0, scale="width", palette="Set1",saturation=0.5,alpha=0.4,label = 'control genes')
plt.xlim(0,60)
plt.legend()
#df_by_gene['0.75ratio_pct'].plot.hist(bins=20, alpha=0.5)
plt.xlabel('percentile at 0.48 ratio')
plt.title('distribution of genes percentile at 0.48 ratio')
#plt.show()
plt.savefig("../a_figures/gene_0.48 ratio percentile.pdf")
# +
#good guide percent
import seaborn as sns
plt.figure(figsize=(4,5))
sns.set_theme(style="whitegrid")
df_by_gene = pd.read_csv('ratio_by_gene_filtered_f24.csv')
df_by_gene.loc[df_by_gene['0.75ratio_pct']>20,'0.75ratio_pct']=20
#df_by_gene.loc[df_by_gene['0.75ratio_pct']<=20,'good guide pct']=df_by_gene['0.75ratio_pct']
#df_by_gene['good guide pct'] = max(20,df_by_gene['0.75ratio_pct'])
#plt.violinplot(dataset =df_by_gene['0.75ratio_pct'])
#df_by_gene['0.75ratio_pct'].plot.box()
#plt.ylim(0,21)
#sns.violinplot(y=df_by_gene['0.75ratio_pct'],inner="quartile",orient='v',palette="Set3",saturation=0.7)
#sns.violinplot(y=df_by_gene['0.75ratio_pct'],inner="point",orient='v',palette="Set3",saturation=0.7)
sns.violinplot(x=df_by_gene['0.75ratio_pct'],inner="quartile", cut=0, scale="width", palette="Set3")
sns.violinplot(x=df_by_gene['0.75ratio_pct'],inner="point", cut=0, scale="width", palette="Set3")
#individual control genes
df_control_bygene = pd.read_csv('ratio_by_gene_controls.csv')
df_control_bygene['0.75ratio_pct'].plot.hist(bins=40, alpha=0.5,ax=ax1)
sns.violinplot(x=df_control_bygene['0.75ratio_pct'],inner="point", cut=0, scale="width", palette="Set3",alpha = 0.5,label = 'control genes')
plt.xlim(0,20)
plt.legend()
#df_by_gene['0.75ratio_pct'].plot.hist(bins=20, alpha=0.5)
plt.xlabel('good guide percentage')
plt.title('Top20% guide percent across genes')
#plt.show()
plt.savefig("a_figures/gene_goodguide_percent.svg",format="svg")
# +
#plot bottom genes
bottom_genes = ['CDC5L','RPS19BP1']
p = np.linspace(0, 100, 6001)
plt.figsize=(25,35)
ax = plt.gca()
for g in bottom_genes:
df_g = df_all_iso[df_all_iso['gene']==g]
ax.plot(np.percentile(df_g['raw ratio'], p),p, label = g)
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls,pooled', c='blue')
#targeting controls,individual
for g in tc_genes:
df_g = df_tc[df_tc['gene']==g]
ax.plot(np.percentile(df_g['raw ratio'], p),p, label = g)
#nt guides
#ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting guides', c='black')
ax.set(
xlabel='raw ratio',
ylabel='percentile')
ax.legend(bbox_to_anchor=(1.4, 0.8))
#plt.xscale('log')
plt.grid(True)
plt.xlim(0,2.5)
plt.show()
# +
# plot all genes
p = np.linspace(0, 100, 6001)
plt.figsize=(25,35)
ax = plt.gca()
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls,pooled', c='blue')
#targeting controls,individual
#nt guides
ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting guides', c='black')
for g in genes:
df_g = df_all_iso[df_all_iso['gene']==g]
ax.plot(np.percentile(df_g['raw ratio'], p),p, label = g)
#targeting controls,pooled
ax.plot(np.percentile(df_tc['raw ratio'], p),p, label = 'targeting controls,pooled', c='blue')
#targeting controls,individual
#nt guides
ax.plot(np.percentile(df_nt['raw ratio'], p),p, label = 'non-targeting guides', c='black')
ax.set(
xlabel='raw ratio',
ylabel='percentile')
ax.legend(bbox_to_anchor=(1.4, 0.8))
#plt.xscale('log')
plt.grid(True)
plt.xlim(0,2.5)
plt.show()
# +
# alternative method
input_file = 'd14_plasmid_library_ratio_targeting_library.csv'
gene_dictionary = {}
with open(input_file, mode ='r') as infile:
reader = csv.reader(infile)
#skip header
#next(reader, None)
for row in reader:
if row[1] not in gene_dictionary.keys():
gene_dictionary[row[1]]=[float(row[4])] #dicitonary with gene and guide raw ratio
else:
gene_dictionary[row[1]].append(float(row[4]))
# +
gene_sum_dic = {}
for g in gene_dictionary.keys():
gene_dictionary[g].sort()
pct_g = np.percentile(gene_dictionary[g],[0,2.5 ,5,10,20, 50, 70, 90,95,100])
good_index = np.argmin(np.abs(np.array(gene_dictionary[g])-0.45)) # 0.45 ratio
good_p = good_index/len(gene_dictionary[g]) # percentile of 0.45 ratio
bad_index1 = np.argmin(np.abs(np.array(gene_dictionary[g])-2)) # 2 ratio
bad_p1 = bad_index1/len(gene_dictionary[g])
bad_index2 = np.argmin(np.abs(np.array(gene_dictionary[g])-2.5)) # 2.5 ratio
bad_p2 = bad_index2/len(gene_dictionary[g])
gene_sum = list(pct_g)+[good_p,bad_p1,bad_p2]
gene_sum_dic[g]= gene_sum
#print(gene_sum)
output_file = 'ratio_by_gene.csv'
with open(output_file,'w') as csvfile:
mywriter = csv.writer(csvfile)
mywriter.writerow(['gene','min','2.5pct','5pct','10pct','20pct','50pct','70pct','90pct','95pct','max',
'0.45ratio_pct','2ratio_pct','2.5ratio_pct'])
for gene in gene_sum_dic.keys():
mywriter.writerow([gene] + gene_sum_dic[gene])
# -
import seaborn as sns
plt.figure(figsize=(4,5))
sns.set_theme(style="whitegrid")
sns.violinplot(y=df['0.45ratio_pct'],inner="quartile",orient='v',palette="Set3",saturation=0.7)
sns.violinplot(y=df['0.45ratio_pct'],inner="point",orient='v',palette="Set3",saturation=0.7)
plt.ylabel('good guide percentage')
plt.ylim(-0.1,1)
plt.title('Top20% guide percent across genes')
#plt.show()
plt.savefig("gene_goodguide_percent.svg",format="svg")
# +
# make new filtered dataset with ratio and percentile
guide_info = {}
with open('integrated_guide_feature_filtered_ver3.csv','r') as inf:
reader = csv.reader(inf)
#skip header
next(reader, None)
for rows in reader:
guide_info[rows[0]]=rows
with open('ratio_percentile_bygene.csv','r') as inf2:
reader = csv.reader(inf2)
#skip header
next(reader, None)
for rows in reader:
#print(rows)
if rows[1] in guide_info.keys():
guide_info[rows[1]]= guide_info[rows[1]]+ rows[4:]+[0,0]
#define good guides
if float(rows[5]) <= 0.45: # top 20% in all guides according to ratio
guide_info[rows[1]][-2]=1 #good guide
if float(rows[6]) <= 0.2: # top20% for each gene
guide_info[rows[1]][-1]=1
with open('integrated_guide_feature_filtered_new_ver3.csv','w') as outf:
writer = csv.writer(outf)
writer.writerow(['guide','gene','refseq','chopped ratio','old relative_ratio','binary_relative_ratio','position','is_5UTR','UTR5_position','is_CDS','CDS_position','is_3UTR',
'UTR3_position','RNAseq2','RNAseq3','RNAseq7','RNAseq8','RNAseq2_relative','RNAseq3_relative','RNAseq7_relative','RNAseq8_relative','G_hp','C_hp','A_hp','T_hp',
'np_vivo_ic_has_data','np_vivo_ic_sum', 'pos','ratio','relative_ratio','0.45ratio_good','top 20 pct per gene'])
for guides in guide_info.keys():
writer.writerow(guide_info[guides])
print(len(guide_info))
# -
print(guide_info[rows[1]])
#make new filtered relative ratio and further filter guides with many off targets
df2 = pd.read_csv('integrated_guide_feature_filtered_new_ver3.csv')
df_blast = pd.read_csv('integrated_features.csv')
df_blast_info = df_blast[['guide','guide_specificity_e10_15','guide_specificity_e1_20']]
df_all = df2.merge(df_blast_info, left_on='guide', right_on='guide')
p = np.linspace(0, 100, 3001)
fig, ax = plt.subplots()
ax.plot(-np.percentile(-df_all['guide_specificity_e1_20'].values, p),p)
#plt.xlim(0,20)
plt.xlabel('off target number')
plt.ylabel('percentile')
ax.invert_xaxis()
#df_all['guide_specificity_e1_20'].plot(kind='hist',figsize=(5,5),bins=np.linspace(0, 100, 200),alpha=0.4,xlim=(0,100),xticks=np.arange(0, 100, step=5))
df_all['guide_specificity_e1_20'].describe(percentiles=[0.2, 0.5,0.8,0.9,0.95])
df_all_filter = df_all[df_all['guide_specificity_e1_20']<=4]
df_all_filter['relative_ratio'] = df_all_filter.groupby("gene")["ratio"].rank(pct=True)
df_all
df_all_filter
| scripts/survival screen data preprocessing and QC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/CodeTunisia/PysNum2022/blob/main/jour3/ODE.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ef072325-b4eb-4e08-ba8a-9ff4bd53331d"
# # Résolution des équations différentielles ordinaires
# Nous allons dans un premier temps nous intéresser aux équations différentielles que l'on peut mettre sous la forme :
# $$\dot{x} = f(x(t), t)$$
# où $f$ est une fonction définie sur une partie $U$ de $\mathbb{R}^2$ , à valeurs dans $\mathbb{R}$.
#
# Une solution de cette équation différentielle est une fonction $x$ de classe $C^1$ définie sur un certain intervalle $I$ de $\mathbb{R}$ et à valeurs dans $\mathbb{R}$ vérifiant :
#
# * (i) $\forall t \in I$ ; $(x(t), t) \in U$;
#
# * (ii) $\forall t \in I$ ; $\dot{x} = f(x(t), t)$.
#
# Nous allons adjoindre à cette équation différentielle une *condition initiale* sous la forme d'un couple $(x_0 , t_0 ) \in U$ et chercher à résoudre **le problème de Cauchy** suivant :
#
# $$\left\{
# \begin{array}{ll}
# \dot{x} = f(x(t), t)\\
# x(t_0) = x_0
# \end{array}
# \right.$$
#
# Sous certaines conditions sur $f$ que nous ne détaillerons pas, ce problème admet une unique solution, que nous allons chercher à déterminer numériquement.
#
# ## Méthodes pour la résolution numérique des équations différentielles ordinaires
# ### Méthode d'Euler explicite (progressive)
# Les méthodes que nous allons étudier consistent à subdiviser l'intervalle de temps $[t_0 , t_0 + T]$ en $n + 1$ points $t_0 < t_1 < \dots < t_n = t_0+ T$ puis à approcher la relation :
# $$x(t_{k+1}) - x(t_k) = \int_{t_k}^{t_{k+1}} \dot{x} dt = \int_{t_k}^{t_{k+1}} f(x(t), t) dt $$
# La méthode *d'Euler explicite* consiste à approcher cette intégrale par la méthode du **rectangle gauche**, autrement dit à approcher $\int_{t_k}^{t_{k+1}} f(x(t), t) dt$ par $(t_{k+1} - t_k)f(x(t), t)$.
#
#
# 
#
# En posant $h_k = t_{k+1} - t_k$ , ceci conduit à définir une suite de valeurs $x_0 , x_1,\dots, x_n$ à partir de la condition initiale
# $x_0$ et de la *relation de récurrence* :
# $$\forall k \in [0, n-1], \quad x_{k+1} = x_{k} + h_k f(x_k, t_k)$$
#
# On observera qu'en général, seul le premier point $x_0$ de cette méthode est une valeur exacte ; les autres points sont calculés à partir de l'approximation précédente, ce qui peut conduire la valeur calculée $x_k$ à s'écarter de plus en plus de la valeur exacte $x(t_k)$.
#
# ### Méthode d'Euler implicite (rétrograde)
# La méthode *d'Euler implicite* consiste à approcher l'intégrale $\int_{t_k}^{t_{k+1}} f(x(t), t) dt$ par la méthode du **rectangle droit**emph, ce qui conduit à définir la suite $(x_0 , x_1,\dots, x_n)$ par les relations :
#
# $$\forall k \in [0, n-1], \quad x_{k+1} = x_{k} + h_k f(x_{k+1}, t_{k+1})$$
#
# 
#
# On observe que cette relation *ne procure pas une relation explicite* de $x_{k+1}$ puisque ce terme est aussi présent dans le second membre. Pour calculer ce terme il est souvent nécessaire de coupler cette méthode à une méthode de résolution numérique des équations telle la méthode de Newton-Raphson. Pour cette raison, elle se révèle plus coûteuse à mettre en oeuvre.
#
# Dans la pratique, la méthode d'Euler implicite se révèle souvent plus stable que la méthode explicite : elle est moins précise à court terme, mais diverge moins rapidement de la solution exacte que la méthode explicite.
#
# ### Méthode de Heun
# $y_0 = y(a)$
#
# Pour $k \in[0, n-1]$
#
# $$v_k = y_k + h F(t_k, y_k)$$
#
# La réccurrence pour $k \in[0, n-2]$
#
# $$y_{k+1} = y_k + h \left( \dfrac{F(t_k, y_k)}{2} + \dfrac{F(t_{k+1}, v_k)}{2} \right)$$
#
# Le terme $v_k$ représente la valeur approchée calculée par Euler. Ainsi, la méthode de Heun consiste à faire une moyenne entre la dérivée au temps $t_k$ et celle au temps $t_{k+1}$ en la valeur approchée calculée par Euler.
# + [markdown] id="f5af0d93-f406-44b2-ad30-51467cce54c6"
# ## Programmation en Python
# On va implémenter ces méthodes et les tester avec $F(t,y) =-2.3 y$, par exemple sur $[0,10]$ avec $y_0= 1$. La solution exacte est alors $t \mapsto exp (-2.3 t)$.
#
# Pour la méthode d'Euler implicite, on doit retrouver la relation de récurrence suivante :
# $$y_{i+1} = \dfrac{1}{1 + 2.3h} y_i$$
#
# + id="14a6dae0-08b5-4ebd-b8ed-2d8612f958d5" outputId="86f20a44-6776-4c26-88aa-81f8926a7e03" colab={"base_uri": "https://localhost:8080/", "height": 295}
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
def EulerExplicite(t0, tmax, y0, h, F):
n = int((tmax-t0)/h)
t = np.zeros(n)
y = np.zeros(n)
# conditions initiales
t[0] = t0
y[0] = y0
# Algorithme Euler explicite
for i in range(n-1):
t[i+1] = t[i] + h
y[i+1] = y[i] + h*F(y[i], t[i])
return t, y
def EulerImplicite(t0, tmax, y0, h):
n = int((tmax-t0)/h)
t = np.zeros(n)
y = np.zeros(n)
# conditions initiales
t[0] = t0
y[0] = y0
# Algorithme Euler Implicite
for i in range(n-1):
t[i+1] = t[i] + h
y[i+1] = y[i]/(1+2.3*h)
return t, y
def Heun(t0, tmax, y0, h, F):
n = int((tmax-t0)/h)
t = np.zeros(n)
y = np.zeros(n)
# conditions initiales
t[0] = t0
y[0] = y0
# Algorithme Heun
for i in range(n-1):
t[i+1] = t[i] + h
vi = y[i] + h*F(y[i], t[i])
y[i+1] = y[i] + h*(F(y[i], t[i])/2 + F(vi, t[i+1])/2)
return t, y
def F(y, t):
return -2.3 * y
def Exacte(t):
return np.exp(-2.3*t)
t0, tmax = 0, 10
y0 = 1
h = 0.1
t, yexp = EulerExplicite(t0, tmax, y0,h, F)
t, yimp = EulerImplicite(t0, tmax, y0,h)
t, yheu = Heun(t0, tmax, y0,h, F)
yode = odeint(F, y0, t)
plt.plot(t, yexp, label = "Euler Explicite")
plt.plot(t, yimp, label = "Euler Implicite")
plt.plot(t, yheu, label = "Heun")
plt.plot(t,yode, label = "odeint" )
plt.plot(t,Exacte(t),"--k", label = "Solution Exacte" )
plt.xlabel("t")
plt.ylabel("y(t)")
plt.title("Méthodes numériques : ODE", weight = "bold")
plt.legend()
plt.show()
# + id="ecabec8d-2db3-4e26-83ae-1ca628d0a402"
# + [markdown] id="cc8756fe-fcd9-4f37-83d3-43d287faa901"
# ## Équation différentielles du second ordre
# Une équation différentielle du second ordre s'écrit:
# $$\ddot{y} = F(y, \dot{y}, t)$$
# Nous pouvons transformer l'équation différentielle scalaire du second degré en équation différentielle vectorielle du premier degré :
#
# $\pmb{x}= \left(\begin{array}{c} y \\ \dot{y} \end{array}\right)$, $\pmb{\dot{x}} = G(t, \pmb{x})$
#
# Les méthodes décrites précédemment (en dehors de la méthode Euler Implicite) s'appliquent parfaitement au cas vectoriel.
#
# ### Équation de mouvement : Oscillateur libre amortie
#
# On considère un pendule simple qui va osciller d'arrière en avant à cause du champ de gravité de la Terre $g = 9.8 \ m/s^2$.
#
# 
#
#
# Le pendule a l'équation du mouvement :
#
# \begin{align*}
# \ddot{\theta} &= - sin(\theta) -0,4 \times \dot{\theta}
# \end{align*}
# Nous allons transformer l'équation différentielle d'ordre 2 en deux équations différentielles d'ordre 1 afin de pouvoir utiliser simplement les méthodes décrites précédemment. En posant $\omega(t)~=~\dot{\theta}(t)$ la vitesse angulaire du pendule, on obtient le système de deux fonctions inconnues suivant :
# \begin{align*}
# \dot{\theta} (t) &= \omega (t) \\
# \dot{\omega }(t) &= - sin(\theta (t)) -0,4 \times \omega (t)
# \end{align*}
# d'où
# $$\pmb{x}= \left(\begin{array}{c} \theta \\ \omega \end{array}\right), \ \pmb{\dot{x}} = G(t, \pmb{x}) = \left(\begin{array}{c} \omega \\ - sin(\theta) -0,4 \times \omega \end{array}\right)$$
#
# Pour résoudre ce système nous devons connaître les deux conditions initiales suivantes :
# \begin{align*}
# \theta(t = 0) &= \theta_0 \\
# \omega (t = 0) &= 0
# \end{align*}
#
# ### Programmation en Python
#
#
# + id="06a055a5-1760-402a-a9f1-7f567aa2a623" outputId="78398cb0-dff3-4eeb-bf51-aa565543a8bd" colab={"base_uri": "https://localhost:8080/", "height": 351}
def EulerExplicite(t0, tmax, x0, h, G):
n = int((tmax-t0)/h)
t = np.zeros(n)
x = np.zeros((2,n))
# conditions initiales
t[0] = t0
x[:,0] = x0
# Algorithme Euler explicite
for i in range(n-1):
t[i+1] = t[i] + h
x[:,i+1] = x[:,i] + h*G(x[:,i], t[i])
return t, x
def Heun(t0, tmax, x0, h, G):
n = int((tmax-t0)/h)
t = np.zeros(n)
x = np.zeros((2, n))
# conditions initiales
t[0] = t0
x[:,0] = x0
# Algorithme Heun
for i in range(n-1):
t[i+1] = t[i] + h
vi = x[:,i] + h*G(x[:,i], t[i])
x[:,i+1] = x[:,i] + h*(G(x[:,i], t[i])/2 + G(vi, t[i+1])/2)
return t, x
def G(x, t):
theta, omega = x[0], x[1]
return np.array([omega, -np.sin(theta)-1*omega])
t0, tmax = 0, 20
x0 = np.array([np.pi/2, 0])
h = 0.1
t, xexp = EulerExplicite(t0, tmax, x0, h, G)
t, xHeun = Heun(t0, tmax, x0, h, G)
xode = odeint(G, x0, t)
xode = xode.T
print(xode.shape)
plt.figure(figsize = (10, 5))
plt.subplot(1, 2, 1)
plt.plot(t, xexp[0,:], label = "Euler Explicite")
plt.plot(t, xHeun[0, :], label = "Heun")
plt.plot(t, xode[0, :],"r--", label = "odeint")
plt.legend()
plt.ylabel(r"$\theta (t)$")
plt.xlabel("t")
plt.subplot(1, 2, 2)
plt.plot(xexp[0,:], xexp[1,:], label = "Euler Explicite")
plt.plot(xHeun[0, :], xHeun[1, :], label = "Heun")
plt.plot(xode[0, :], xode[1, :],"r--", label = "odeint")
plt.legend()
plt.show()
# + id="3f192cf0-4f7d-46d6-b9f5-cb807a4970dc"
| jour3/ODE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Drawiin/algoritmos-basicos/blob/master/copa_do_mundo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Bb3yl3VDqL0r"
# #Correlação
# É uma relação de dependencia ou associação entre duas variáveis estatisticas, entretanto essa relação pode ou não ser causal, ou seja correlação não é garantia de de causa e efeito, esse tipo de medida é padronizada, de maneira que assuma um valor sempre entre 1 e -1, onde o valor absolutop 1 sempre indica uma correlação perfeita, ou seja duas variaveis que variam juntas independente de qual proporação da variação, essa vairação pode ser tanto positiva quanto negativa:
# ### Negativa
# > Uma variação positiva em uma variavel acarreta em uma variação negativa em outra, e visse versa.
#
# ### Positiva
# > Uma variação positiva em uma variável acarreta em uma variação positiva em outra, ou uma variação negativa em uma accareta em uma variação negativa em outra, ou seja elas crescen ou aumentam juntas, em uma determinada proporção de uma para outra.
# + id="K9uzn6_9Anfl"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="22lUfBtMAyhG"
games = pd.read_csv('https://raw.githubusercontent.com/edullapa/data/master/FIFAWorldCup2018.csv')
# + id="x-cLDFXUBAjP"
corrs = games.corr()
# + [markdown] id="EEMwuN_MyoIx"
# ### Visualizando
# No dataset abaixo podemos ver a matris de correlação calculada para o dataset de jogos da copa de 2018, no mapa de calor algumas correlações, a maioria é positiva, ignorando as auto correlação, podemos observar por exemplo que a **% de posessão de bola** se correlaciona de maneira positiva com **Gols marcados**, e que escanteios se correlacionam de forma negativa com a posse de bola, ou seja quando há uma maior posse de bola há menos escanteios e visse versa.
#
# + id="bBrzL0-XBTF9" outputId="d91cd76a-0d3b-4eb0-fe63-72e49c8fa61f" colab={"base_uri": "https://localhost:8080/", "height": 1000}
f = plt.figure(figsize=(19, 15))
plt.matshow(corrs, fignum=f.number)
plt.xticks(range(games.shape[1]), games.columns, fontsize=14, rotation=45)
plt.yticks(range(games.shape[1]), games.columns, fontsize=14)
cb = plt.colorbar()
cb.ax.tick_params(labelsize=14)
# + [markdown] id="Duy-1Cjk8c7W"
#
| copa_do_mundo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''SQL'': conda)'
# name: python3
# ---
import pandas as pd
from pathlib import Path
csvpath = Path('../Resources/vehicles_mod.csv')
combined_df = pd.read_csv(
csvpath,
index_col='posting_date',
infer_datetime_format=True
)
combined_df.isnull().mean()
print(combined_df['lat'].isnull().sum())
print(combined_df['long'].isnull().sum())
print(combined_df['price'].isnull().sum())
print(combined_df['model'].isnull().sum())
print(combined_df['condition'].isnull().sum())
print(combined_df['manufacturer'].isnull().sum())
# # Assumptions made / Next steps
# 1. Based upon combination of **year** & **odometer** reading make assupmtion on condition to fill in the ~43% of entries missing a condition
# - Condition Framework
# - If older year and high mileage
# - condition = oldest (worst)
# - If newer year and low mileage
# - condition = newest (best)
# - in between? (good vs fair) (std/variance)
# 2. Both make/manufacturer blank?
# 3. Need to drop non-continental 48
#
# Forrest: parsing/categorizing the odometer for condition
#
# Vish: parsing desc to help fill in missing make values
odometer_range_df = combined_df.loc[
(combined_df['odometer'] >= 75000) &
(combined_df['odometer'] < 100000)
]
grouped_multiple = odometer_range_df.groupby(['region'])
| notebooks/old_data_clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import packages
import pandas as pd
# # Load data
DATASET_FOLDER = '../../datasets/tempetes'
path = DATASET_FOLDER + '/' + 'monthly_hist_temperatures.csv'
hist_temp = pd.read_csv(path,sep = ",")
hist_temp = hist_temp.drop(["Unnamed: 0", "Statistics"], axis = 1)
hist_temp = hist_temp.rename({"Temperature - (Celsius)":"avg_monthly_temperature"}, axis=1)
hist_temp.head()
# # Prepare data s.t. to join it with emdat db
hist_temp_by_year_by_country_agg = hist_temp.groupby(["ISO3", "Year"]).agg({"avg_monthly_temperature": ["mean", "min", "max", "median"]}).reset_index()
column_headers = []
for i in range(len(list(hist_temp_by_year_by_country_agg.columns.get_level_values(0)))):
str0 = list(hist_temp_by_year_by_country_agg.columns.get_level_values(0))[i]
str1 = list(hist_temp_by_year_by_country_agg.columns.get_level_values(1))[i]
header = str0 + "_" + str1
column_headers.append(header)
column_headers
hist_temp_by_year_by_country_agg.columns = column_headers
hist_temp_by_year_by_country_agg.head()
hist_temp_by_year_by_country_agg.shape
# # Join on emdat db
# ### Import emdat db
DATASET_FOLDER = '../../datasets/tempetes'
path = DATASET_FOLDER + '/' + 'wb_disasters_bdd.xlsx'
disasters_df = pd.read_excel(path)
disasters_df.head()
# ### Filter on storms
storms_df = disasters_df[disasters_df["Disaster Type"]=="Storm"]
storms_df.shape
storms_df.columns
# # Join
hist_temp_by_year_by_country_agg.dtypes
storms_df["Start Year"] = storms_df["Start Year"].astype("int64")
storms_by_year_by_country_hist_temp = pd.merge(storms_df, hist_temp_by_year_by_country_agg, how="left", left_on=["Start Year", "ISO"], right_on=["Year_", "ISO3_"])
storms_by_year_by_country_hist_temp.head()
storms_by_year_by_country_hist_temp_sel_cols = storms_by_year_by_country_hist_temp[["Start Year",
"ISO",
"Country",
"Disaster Type",
"No Affected",
"Total Damages ('000 US$)",
"avg_monthly_temperature_mean",
"avg_monthly_temperature_min",
"avg_monthly_temperature_max",
"avg_monthly_temperature_median"
]]
storms_by_year_by_country_hist_temp_sel_cols.shape
storms_by_year_by_country_hist_temp_sel_cols.head()
storms_by_year_by_country_hist_temp_sel_cols.isnull().sum()
test = storms_by_year_by_country_hist_temp_sel_cols[storms_by_year_by_country_hist_temp_sel_cols["avg_monthly_temperature_mean"].isnull()]
test.head(20)
# # Write csv file in datasets > tempetes directory
storms_by_year_by_country_hist_temp_sel_cols.to_csv('../../datasets/tempetes/' + '/' + 'emdat_with_hist_temperatures.csv')
| model_tempetes/notebooks/emdat_enrichment_hist_temp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import the usual libraries
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
#import matplotlib.patches as mpatches
# Enable inline plotting
# %matplotlib inline
# Progress bar
from tqdm.auto import trange, tqdm
# -
import webbpsf_ext, webbpsf
webbpsf_ext.setup_logging('WARN', verbose=False)
inst = webbpsf.NIRCam()
inst.filter='F405N'
inst.include_si_wfe = False
# +
# %%time
inst.detector_position = (1024,1024) # center
psf_cen = inst.calc_psf(fov_pixels=64, oversample=4)
inst.detector_position = (10,10) # bottom left
psf_corn1 = inst.calc_psf(fov_pixels=64, oversample=4)
inst.detector_position = (10,2000) # top left
psf_corn2 = inst.calc_psf(fov_pixels=64, oversample=4)
inst.detector_position = (2000,2000) # top right
psf_corn3 = inst.calc_psf(fov_pixels=64, oversample=4)
inst.detector_position = (2000,10) # bottom right
psf_corn4 = inst.calc_psf(fov_pixels=64, oversample=4)
# +
fig, axes = plt.subplots(2,2, figsize=(10,10))
axes = axes.flatten()
im_cen = psf_cen[2].data
imdiff1 = psf_corn1[2].data - im_cen
imdiff2 = psf_corn2[2].data - im_cen
imdiff3 = psf_corn3[2].data - im_cen
imdiff4 = psf_corn4[2].data - im_cen
imall = [imdiff2, imdiff3, imdiff1, imdiff4]
titles = ['Top Left (10,2000)', 'Top Right (2000,2000)',
'Bottom Left (10,10)', 'Bottom Right (2000,10)']
for i, im in enumerate(imall):
ax = axes[i]
med = np.median(im)
std = np.std(im)
vmin, vmax = 5*std*np.array([-1,1])
ax.imshow(im, interpolation='bicubic', cmap='RdBu', vmin=vmin, vmax=vmax)
ax.set_title(titles[i])
fig.suptitle('F405N PSF Difference vs Center (distortion=`griddata`)', fontsize=14)
fig.tight_layout()
# +
fig, axes = plt.subplots(2,2, figsize=(10,10))
axes = axes.flatten()
im_cen = psf_cen[2].data
imdiff1 = psf_corn1[2].data - im_cen
imdiff2 = psf_corn2[2].data - im_cen
imdiff3 = psf_corn3[2].data - im_cen
imdiff4 = psf_corn4[2].data - im_cen
imall = [imdiff2, imdiff3, imdiff1, imdiff4]
titles = ['Top Left (10,2000)', 'Top Right (2000,2000)',
'Bottom Left (10,10)', 'Bottom Right (2000,10)']
for i, im in enumerate(imall):
ax = axes[i]
med = np.median(im)
std = np.std(im)
vmin, vmax = 5*std*np.array([-1,1])
ax.imshow(im, interpolation='bicubic', cmap='RdBu', vmin=vmin, vmax=vmax)
ax.set_title(titles[i])
fig.suptitle('F405N PSF Difference vs Center (distortion=`RegularGridInterpolator`)', fontsize=14)
fig.tight_layout()
# -
| notebooks/test_distortions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas - 数据处理
# ## 一、简介
# Pandas是一个强大的分析结构化数据的工具集;它的使用基础是Numpy(提供高性能的矩阵运算);用于数据挖掘和数据分析,同时也提供数据清洗功能。
# - Series:一维数组,与Numpy中的一维array类似。二者与Python基本的数据结构List也很相近。Series如今能保存不同种数据类型,字符串、boolean值、数字等都能保存在Series中。
# - Time- Series:以时间为索引的Series。
# - DataFrame:二维的表格型数据结构。很多功能与R中的data.frame类似。可以将DataFrame理解为Series的容器。
# - Panel :三维的数组,可以理解为DataFrame的容器。
# - Panel4D:是像Panel一样的4维数据容器。
# - PanelND:拥有factory集合,可以创建像Panel4D一样N维命名容器的模块。
# ## 二、基本使用
# (1) 生成序列
import pandas as pd
import numpy as np
a1 = pd.Series([1,2,3,np.nan,44,1])
print(a1)
# (2) 生成日期
# Signature: pd.date_range(start=None, end=None, periods=None, freq='D',
# tz=None, normalize=False, name=None, closed=None, **kwargs)
datas = pd.date_range('20190101',periods=6)
print(datas)
# (3)生成数据
# 定义行列
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
print(df)
# 不定义行列
df1 = pd.DataFrame(np.arange(12).reshape(3,4))
print(df1)
print(df1.describe()) #计算
# (4) 基本操作
# 用字典定义
df2 = pd.DataFrame({'A':'aaa','B':pd.Categorical([1,2])}) #缺失的会复制
print(df2)
print(df2.dtypes)
print(df2.index) # 打印行索引
print(df2.columns)# 打印列索引
print(df2.values) # 打印值
print(df2.T) # 转制
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df.sort_index(axis=1,ascending=False)
df.sort_index(axis=0,ascending=False)
# 根据某一参数排序
df.sort_values(by='c')
# ## 三、进阶使用
# (1) 切片取值
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df['a']
# 直接索引
df[0:3]
# 通过下标查找(类似于EXCEL中的行列号)
df.iloc[0:3]
df.iloc[0:3,1:2]
df.iloc[0:1,:]
# 通过标签查找(类似于EXCEL中的标签行列)
df.loc['20190101',:]
# (2)筛选
# 按条件筛选
df[df['a']>0.01]
# (3)修改
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df.iloc[2,2] = 5.5555
df
df[df['a']<0] = 1
df
df.a[df['a']>0.5] = 5
df
# (4) 处理丢失数据
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df.iloc[0,1] = np.nan
df.iloc[1,2] = np.nan
df
df.dropna(axis=0,how='all') # all
df.dropna(axis=0,how='any') # all
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df.iloc[0,1] = np.nan
df.iloc[1,2] = np.nan
df.fillna(value=1000)
df = pd.DataFrame(np.random.randn(6,4),index=datas,columns=['a','b','c','d'])
df.iloc[0,1] = np.nan
df.iloc[1,2] = np.nan
df.isnull()
# (5) 导入导出数据
# pandas可以读取csv、excel、hdf、sql、json、msgpack、html、gbq、stata、clipboard、pickle。读取的格式为read_xxx,写的格式为to_xxx
# 读取CSV
import pandas as pd
data = pd.read_csv('lessons/csvtest1.csv')
print(data)
# 存储为任意格式
data.to_pickle('lessons/csvtest1.pickle')
# (6)合并数据
import pandas as pd
import numpy as np
df0 = pd.DataFrame(np.ones((3,4))*0, columns=['a','b','c','d'])
df1 = pd.DataFrame(np.ones((3,4))*1, columns=['a','b','c','d'])
df2 = pd.DataFrame(np.ones((3,4))*2, columns=['a','b','c','d'])
print(df0)
print(df1)
print(df2)
res = pd.concat([df0,df1,df2], axis=0)
print(res)
res = pd.concat([df0,df1,df2], axis=1)
print(res)
res = pd.concat([df0,df1,df2], axis=0, ignore_index=True)
print(res)
# join,['inner','outer']
import pandas as pd
import numpy as np
df0 = pd.DataFrame(np.ones((3,4))*0, columns=['a','b','c','d'],index=[1,2,3])
df1 = pd.DataFrame(np.ones((3,4))*1, columns=['b','c','d','e'],index=[2,3,4])
print(df0)
print(df1)
res = pd.concat([df0,df1],join='outer',sort=False)
print(res)
res = pd.concat([df0,df1],join='inner',ignore_index=True)
print(res)
# +
# res = pd.merge() merge
# -
# (7) plot画图
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
data = pd.Series(np.random.randn(1000),index=np.arange(1000))
data = data.cumsum()
data.plot()
plt.show()
data = pd.DataFrame(np.random.randn(1000,4),index=np.arange(1000),columns=list('ABCD'))
print(data.head())
data = data.cumsum()
data.plot()
plt.show()
# 参考文献
# 1. [数据分析三剑客之pandas](https://www.cnblogs.com/peng104/p/10398490.html)
| 99 Cache/.ipynb_checkpoints/lesson1-2 Pandas-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="LGwSDVpO288T" outputId="d47fd733-15bc-464a-f265-f006d67366a4"
# !wget "https://storage.googleapis.com/kaggle-data-sets/38019/306654/bundle/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=gcp-kaggle-com%40k<EMAIL>.<EMAIL>.g<EMAIL>account.com%2F20210224%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210224T022521Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=host&X-Goog-Signature=791a652163ff1ba3692db8ad46063bb7048a62edf8e77a82f15994c97731293128ee140366ff06146169104ab38c653831954bfbb433e1f3b1e2c8858f8154ecc54f732a0fd4e6ce497b48662c20b2d07da70b36e1853a37a56e20107520e9fb955e3f2cb652fca0c127316cdc403cc9281a86a681ea84c8bd58baa818fc59b2a615a99a1bc5bf723a065a812263543ab179ac5c9c07b70045085705b939b24d4b8bc054c6e321d5b4e3533ab0243fcf06e36f73b284ca192818d36454a2d1fbe1e1254b731f3d7a7b8bde39b3df3d4130da958b24ca24a8239ad8355a3d836c7d6c38590a6ba99377e809c033b0fef7d3b5ad9e865b14f032fd8fe2977f4906"
# + colab={"base_uri": "https://localhost:8080/"} id="KoPpf4453Goy" outputId="385b4566-260c-4296-e82d-0dc71f53feef"
# !unzip "/content/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=gcp-k<PASSWORD>-com@kaggle-161607.<EMAIL>%2F20210224%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210224T022521Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=hos" -d "/content/dataset"
# + id="POP1Ux373N78"
DIR = "/content/dataset/samples/"
# + id="TkVMIukJ6ml2"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.models import load_model
from keras import layers
from keras.models import Model
import cv2 as cv
import string
import csv
import keras
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
# Any results you write to the current directory are saved as output.
# + colab={"base_uri": "https://localhost:8080/"} id="1_DaFPMr6nWD" outputId="c64836f4-8747-40cb-d190-09c6a055fe8d"
CREAT_MODEL = True
data = []
# Image Parameters
num_chars = 5
img_height = 50
img_width = 200
img_source = DIR
img_count = 0
for dirname, _, filenames in os.walk(img_source):
for filename in filenames:
img=cv.imread(img_source + "/" + filename, cv.IMREAD_GRAYSCALE)
parts = filename.split('.')
data.append([img, parts[0], filename])
img_count += 1
break
print(str(img_count) + " total data points")
# print(data)
# + colab={"base_uri": "https://localhost:8080/", "height": 536} id="R9xm8hpi6y9m" outputId="806529a6-3dc5-4b19-8850-fe0faf826e81"
for filename in data[:3]:
img=mpimg.imread(DIR + filename[2], cv.IMREAD_GRAYSCALE)
plt.figure(figsize=(10,10))
plt.imshow(img, cmap ='gray')
# + colab={"base_uri": "https://localhost:8080/"} id="rdIiAM0F66Km" outputId="10db4e33-fbaf-4462-885a-ee9f90a93f10"
from sklearn.model_selection import train_test_split
data_x = []
data_y = []
for features,label,filename in data:
data_x.append(features)
data_y.append(label)
train_X,rem_X,train_Y,rem_Y = train_test_split(data_x, data_y, train_size=0.66, random_state=52, shuffle=True)
val_X,test_X,val_Y,test_Y = train_test_split(rem_X, rem_Y, train_size=0.5, random_state=22, shuffle=True)
print(str(len(train_X)) + " training samples")
print(str(len(val_X)) + " validation samples")
print(str(len(test_X)) + " test samples")
# + id="UOMW1Ych6-bG"
from keras.utils import to_categorical
#Dictionary to Encode Letters
valid_chars = ['2', '3', '4', '5', '6', '7', '8', 'b', 'c', 'd', 'e', 'f', 'g', 'm', 'n', 'p', 'w', 'x', 'y']
keys = list(range(0,len(valid_chars)))
my_dict = dict(zip(keys, valid_chars))
def target_splitter(Y):
num_entries = len(Y)
num_indexes = len(Y[0])
return_val_index = []
return_val = []
for j in range(num_entries):
return_val_index = []
for i in range(num_indexes):
encoding_index = valid_chars.index(Y[j][i])
# return_val_index.append(to_categorical(encoding_index))
return_val_index.append(encoding_index)
return_val.append(return_val_index)
return return_val
# + id="m8CyYXUh74C3"
#Splitting Into Individual Letters, redimensioning image array
train_Y_split = target_splitter(train_Y)
train_Y_split = np.array(train_Y_split)
train_X_array = np.array(train_X, ndmin=4)
# print(train_X_array.shape)
train_X_array = train_X_array.reshape(len(train_X), img_height, img_width, 1)
val_Y_split = target_splitter(val_Y)
val_Y_split = np.array(val_Y_split)
val_X_array = np.array(val_X, ndmin=4)
val_X_array = val_X_array.reshape(len(val_X), img_height, img_width, 1)
test_Y_split = target_splitter(test_Y)
test_Y_split = np.array(test_Y_split)
test_X_array = np.array(test_X, ndmin=4)
test_X_array = test_X_array.reshape(len(test_X), img_height, img_width, 1)
#Splitting Each Individual Letter into Categorical Data
train_Y_char_1 = train_Y_split[:,0]
train_Y_char_1 = to_categorical(train_Y_char_1)
train_Y_char_2 = train_Y_split[:,1]
train_Y_char_2 = to_categorical(train_Y_char_2)
train_Y_char_3 = train_Y_split[:,2]
train_Y_char_3 = to_categorical(train_Y_char_3)
train_Y_char_4 = train_Y_split[:,3]
train_Y_char_4 = to_categorical(train_Y_char_4)
train_Y_char_5 = train_Y_split[:,4]
train_Y_char_5 = to_categorical(train_Y_char_5)
val_Y_char_1 = val_Y_split[:,0]
val_Y_char_1 = to_categorical(val_Y_char_1)
val_Y_char_2 = val_Y_split[:,1]
val_Y_char_2 = to_categorical(val_Y_char_2)
val_Y_char_3 = val_Y_split[:,2]
val_Y_char_3 = to_categorical(val_Y_char_3)
val_Y_char_4 = val_Y_split[:,3]
val_Y_char_4 = to_categorical(val_Y_char_4)
val_Y_char_5 = val_Y_split[:,4]
val_Y_char_5 = to_categorical(val_Y_char_5)
test_Y_char_1 = test_Y_split[:,0]
test_Y_char_1 = to_categorical(test_Y_char_1)
test_Y_char_2 = test_Y_split[:,1]
test_Y_char_2 = to_categorical(test_Y_char_2)
test_Y_char_3 = test_Y_split[:,2]
test_Y_char_3 = to_categorical(test_Y_char_3)
test_Y_char_4 = test_Y_split[:,3]
test_Y_char_4 = to_categorical(test_Y_char_4)
test_Y_char_5 = test_Y_split[:,4]
test_Y_char_5 = to_categorical(test_Y_char_5)
# + id="92OKdYcX7-AJ"
class CapthaRec(object):
def __init__(self):
self.symbols = string.ascii_uppercase[:19] #only taking 19 chars letters
#self.symbols = 15
def create_model(self, img_shape):
img = layers.Input(shape=img_shape) # Get image as an input and process it through some Convs
conv1 = layers.Conv2D(16, (3, 3), padding='same', activation='relu')(img)
bn1 = layers.BatchNormalization()(conv1)
mp1 = layers.MaxPooling2D(padding='same')(bn1)
conv2 = layers.Conv2D(32, (3, 3), padding='same', activation='relu')(mp1)
bn2 = layers.BatchNormalization()(conv2)
mp2 = layers.MaxPooling2D(padding='same')(bn2)
conv3 = layers.Conv2D(32, (3, 3), padding='same', activation='relu')(mp2)
bn3 = layers.BatchNormalization()(conv3)
mp3 = layers.MaxPooling2D(padding='same')(bn3)
flat = layers.Flatten()(mp3)
outs = list()
for _ in range(num_chars):
dens1 = layers.Dense(len(valid_chars)*2, activation='relu')(flat)
drop = layers.Dropout(rate=0.5)(dens1)
res = layers.Dense(len(valid_chars), activation='sigmoid')(drop)
outs.append(res)
model = Model(inputs=img, outputs=outs)
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=["accuracy"])
return model
def gen_csv_data(self, file_path):
with open(file_path, newline='') as csvfile:
spamreader = csv.reader(csvfile, delimiter=',', quotechar='|')
for row in spamreader:
yield row
def predict(self, model, filepath):
img = cv.imread(filepath, cv.IMREAD_GRAYSCALE)
if img is not None:
if img.shape != (img_height, img_width):
img = cv.resize(img, (img_width, img_height))
img = img / 255.0
else:
print(filepath, "Not detected");
return
img = img[np.newaxis, :, :, np.newaxis]
res = model.predict(img)
res = np.array(res)
ans = np.reshape(res, (num_chars, len(self.symbols)))
l_ind = list()
probs = list()
capt = list()
for a in ans:
l_ind.append(np.argmax(a))
probs.append(np.max(a))
for l in l_ind:
capt.append(self.symbols[l])
return "".join(capt), sum(probs) / num_chars
# + colab={"base_uri": "https://localhost:8080/"} id="ROnMDx_e8Bdj" outputId="f63ba65c-ff59-4566-8786-a5068da6949e"
num_epochs = 100
batch_size = 16
cr = CapthaRec()
# X, Y = cr.preprocess_data("/kaggle/input/captcha-images")
if CREAT_MODEL:
model = cr.create_model((img_height, img_width, 1))
model.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
model.summary()
captcha_trained = model.fit(
train_X_array, [train_Y_char_1, train_Y_char_2, train_Y_char_3, train_Y_char_4, train_Y_char_5],
validation_data=(val_X_array, [val_Y_char_1, val_Y_char_2, val_Y_char_3, val_Y_char_4, val_Y_char_5]),
batch_size=batch_size, epochs=num_epochs, shuffle=True, verbose=1)
model.save('test2.h5')
model = load_model('test2.h5')
# + colab={"base_uri": "https://localhost:8080/", "height": 551} id="9Pu2_0hT8D9E" outputId="3706d86b-aa0d-4eef-c0cd-3bd17b94bb54"
axes = plt.gca()
axes.set_ylim([0,1])
#First Char Accuracies
axes = plt.gca()
axes.set_ylim([0,1])
accuracy = captcha_trained.history['dense_11_accuracy']
val_accuracy = captcha_trained.history['val_dense_11_accuracy']
plt.plot(range(len(accuracy)), accuracy, 'b', label='Training accuracy')
plt.plot(range(len(val_accuracy)), val_accuracy, 'r', label='Validation accuracy')
plt.title('Training and validation accuracy - 1st Char')
plt.legend()
plt.figure()
plt.legend()
plt.show()
# + [markdown] id="6qllnZcQ8Ryg"
# ## Predicting Values
# + colab={"base_uri": "https://localhost:8080/"} id="tLICC8WI8OdR" outputId="a0484bdc-a4c3-4cf6-c091-9d70ae3d90c0"
test_dummy = test_X_array[0]
test_dummy.reshape(1, 50, 200, 1)
test_results = model.predict(test_X_array)
test_results = np.array(test_results)
# print(test_results.shape)
results_array = []
num_correct_preds = 0
num_correct_char_preds = []
per_char_preds_correct = np.zeros(shape=(num_chars, len(valid_chars)))
per_char_preds_total = np.zeros(shape=(num_chars, len(valid_chars)))
for p in range (0, len(test_X)):
temp_string = ""
temp_array = []
for q in range(0, num_chars):
#Per position results
pred_char = my_dict.get(np.argmax(test_results[q,p,:]))
temp_string = temp_string + pred_char
temp_array.append(pred_char == test_Y[p][q])
#Per character results
letter_index = valid_chars.index(test_Y[p][q])
per_char_preds_total[q,letter_index] = per_char_preds_total[q,letter_index] + 1
if pred_char == test_Y[p][q]:
per_char_preds_correct[q,letter_index] = per_char_preds_correct[q,letter_index] + 1
result_string = ""
if temp_string == test_Y[p]:
result_string = "CORRECT"
num_correct_preds = num_correct_preds + 1
else:
result_string = "INCORRECT"
num_correct_char_preds.append(temp_array)
print("True Value: " + test_Y[p] + " Predicted Value: " + temp_string + " Result: " + result_string)
# + colab={"base_uri": "https://localhost:8080/"} id="kWBGcicO8Ub8" outputId="46f87bd1-f9d6-4b96-ffab-14d44387ffeb"
from tabulate import tabulate
print("Total Word Accuracy = " + str(num_correct_preds/len(test_X)))
num_correct_char_preds = np.array(num_correct_char_preds)
print("Per Char Accuracy : " + " ".join(map(str, np.sum(num_correct_char_preds, axis = 0)/len(test_X))))
# print(per_char_preds_total.shape)
print("")
print("Per Char Per Position Accuracy")
letter_array = np.array([my_dict.get(i) for i in range(0, len(valid_chars))], ndmin=2)
letter_array = letter_array.reshape(len(valid_chars), 1)
temp_array = np.transpose(per_char_preds_correct)/np.transpose(per_char_preds_total)
temp_array = np.append(letter_array, temp_array, 1)
print(tabulate(temp_array, floatfmt=".3f", headers=["Char", "Char 1", "Char 2", "Char 3", "Char 4", "Char 5"]))
# + id="4vq79QXl8neG"
| Notebooks/CAPTCHA Recognition/CAPTCHA_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This example shows how to perform GP regression, but using **variational inference** rather than exact inference. There are a few cases where variational inference may be prefereable:
#
# 1) If you have lots of data, and want to perform **stochastic optimization**
#
# 2) If you have a model where you want to use other variational distributions
#
# KISS-GP with SVI was introduced in:
# https://papers.nips.cc/paper/6426-stochastic-variational-deep-kernel-learning.pdf
# +
import math
import torch
import gpytorch
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# Create a training set
# We're going to learn a sine function
train_x = torch.linspace(0, 1, 1000)
train_y = torch.sin(train_x * (4 * math.pi)) + torch.randn(train_x.size()) * 0.2
# ## Performing SGD - the dataloader
#
# Because we want to do stochastic optimization, we have to put the dataset in a pytorch **DataLoader**.
# This creates easy minibatches of the data
from torch.utils.data import TensorDataset, DataLoader
train_dataset = TensorDataset(train_x, train_y)
train_loader = DataLoader(train_dataset, batch_size=64, shuffle=True)
# ## The model
#
# This is pretty similar to a normal regression model, except now we're using a `gpytorch.models.GridInducingVariationalGP` instead of a `gpytorch.models.ExactGP`.
#
# Any of the variational models would work. We're using the `GridInducingVariationalGP` because we have many data points, but only 1 dimensional data.
#
# Similar to exact regression, we use a `GaussianLikelihood`.
# +
class GPRegressionModel(gpytorch.models.GridInducingVariationalGP):
def __init__(self):
super(GPRegressionModel, self).__init__(grid_size=64, grid_bounds=[(-0.05, 1.05)])
self.mean_module = gpytorch.means.ConstantMean(constant_bounds=[-1e-5,1e-5])
self.covar_module = gpytorch.kernels.RBFKernel(log_lengthscale_bounds=(-5, 6))
self.register_parameter('log_outputscale', torch.nn.Parameter(torch.Tensor([0])), bounds=(-5,6))
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
covar_x = covar_x.mul(self.log_outputscale.exp())
return gpytorch.random_variables.GaussianRandomVariable(mean_x, covar_x)
model = GPRegressionModel().cuda()
likelihood = gpytorch.likelihoods.GaussianLikelihood().cuda()
# -
# ## The training loop
#
# This training loop will use **stochastic optimization** rather than batch optimization
# +
model.train()
likelihood.train()
# We'll do 40 iterations of optimization
n_iter = 40
# We use SGD here, rather than Adam
# Emperically, we find that SGD is better for variational regression
optimizer = torch.optim.SGD([
{'params': model.parameters()},
{'params': likelihood.parameters()},
], lr=0.1)
# We use a Learning rate scheduler from PyTorch to lower the learning rate during optimization
# We're going to drop the learning rate by 1/10 after 3/4 of training
# This helps the model converge to a minimum
scheduler = torch.optim.lr_scheduler.MultiStepLR(optimizer, milestones=[0.75 * n_iter], gamma=0.1)
# Our loss object
# We're using the VariationalMarginalLogLikelihood object
mll = gpytorch.mlls.VariationalMarginalLogLikelihood(likelihood, model, n_data=train_y.size(0))
# The training loop
def train():
for i in range(n_iter):
scheduler.step()
# Within each iteration, we will go over each minibatch of data
for x_batch, y_batch in train_loader:
x_batch = torch.autograd.Variable(x_batch.float().cuda())
y_batch = torch.autograd.Variable(y_batch.float().cuda())
optimizer.zero_grad()
# We're going to use two context managers here
# The use_toeplitz flag makes learning faster on the GPU
# See the DKL-MNIST notebook for an explanation
# The diagonal_correction flag improves the approximations we're making for variational inference
# It makes running time a bit slower, but improves the optimization and predictions
with gpytorch.settings.use_toeplitz(False), gpytorch.beta_features.diagonal_correction():
output = model(x_batch)
loss = -mll(output, y_batch)
print('Iter %d/%d - Loss: %.3f (%.3f)' % (i + 1, n_iter, loss.data[0], optimizer.param_groups[0]['lr']))
# The actual optimization step
loss.backward()
optimizer.step()
# %time train()
# -
# ## Testing the model
# +
model.eval()
likelihood.eval()
test_x = torch.autograd.Variable(torch.linspace(0, 1, 51)).cuda()
with gpytorch.settings.max_cg_iterations(2000), gpytorch.settings.use_toeplitz(False), gpytorch.beta_features.diagonal_correction():
observed_pred = likelihood(model(test_x))
lower, upper = observed_pred.confidence_region()
fig, ax = plt.subplots(1, 1, figsize=(4, 3))
ax.plot(train_x.cpu().numpy(), train_y.cpu().numpy(), 'k*')
ax.plot(test_x.data.cpu().numpy(), observed_pred.mean().data.cpu().numpy(), 'b')
ax.fill_between(test_x.data.cpu().numpy(), lower.data.cpu().numpy(), upper.data.cpu().numpy(), alpha=0.5)
ax.set_ylim([-3, 3])
ax.legend(['Observed Data', 'Mean', 'Confidence'])
| examples/kissgp_variational_regression_cuda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
from openpyxl import Workbook
from openpyxl.utils import get_column_letter
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
tracename="WDEV_0"
file = open(tracename)
page_size = 4
update_count = 0
entry = {"lpn": 0, "access_time": 0.0, "update_time": 0.0, "update_count": 0, "update_min": 0, "update_max": 0}
large_lpn = 0
capacity = 32 * 1024 * 1024 # 128G SSD = 32 * 1024* 1024*4KB
#maptable = pd.DataFrame([ [0.0,0.0,0,0.0,0.0,0.0,0] for i in range(0, capacity)],columns=['access_time', 'update_interval', 'update_count', 'update_avg', "update_min", "update_max", "write_flag"])
maptable = [{"access_time": 0.0, "update_interval": 0.0, "update_count": 0, "update_avg": 0.0, "update_min": 1.0 * 10**12, "update_max": 0.0, "write_flag": 0} for i in range(0, capacity)] # write_flag =1 表示已经被写
print("初始化完成 maptable:", len(maptable), sys.getsizeof(maptable) / (1024 * 1024))
#print("初始化完成 maptable:", maptable.index,maptable.columns, sys.getsizeof(maptable) / (1024 * 1024))
#trace df
#tracedf=pd.read_table(tracename,sep=" ",header=None)
#file = open(tracename)
while 1:
lines = file.readlines(100000)
if not lines:
break
for line in lines:
tmp = line.split()
#print(tmp)
if(int(tmp[4]) == 1): # read
first_lpn = int(int(tmp[2]) / page_size) % capacity
while(first_lpn <= int(int(tmp[2]) / page_size + int(tmp[3]) / page_size) % capacity):
#print("first_lpn", first_lpn)
if(maptable[first_lpn]['write_flag'] == 1): # 被写过
# maptable[int(int(tmp[2]) / page_size)]['access_time']=float(tmp[0]) #本次访问的时间
maptable[first_lpn]['update_interval'] = float(tmp[0]) - maptable[first_lpn]['access_time'] # 本次访问的时间
maptable[first_lpn]['update_count'] = maptable[first_lpn]['update_count'] + 1
# if(maptable[first_lpn]['update_interval'] <= 10):
# print(tmp)
if(maptable[first_lpn]['update_min'] > maptable[first_lpn]['update_interval']):
maptable[first_lpn]['update_min'] = maptable[first_lpn]['update_interval']
if(maptable[first_lpn]['update_max'] < maptable[first_lpn]['update_interval']):
maptable[first_lpn]['update_max'] = maptable[first_lpn]['update_interval']
maptable[first_lpn]['update_avg'] = maptable[first_lpn]['update_avg'] + (maptable[first_lpn]['update_interval'] - maptable[first_lpn]['update_avg']) / maptable[first_lpn]['update_count']
first_lpn = first_lpn + 1
elif(int(tmp[4]) == 0):
first_lpn = int(int(tmp[2]) / page_size) % capacity
while(first_lpn <= int(int(tmp[2]) / page_size + int(tmp[3]) / page_size) % capacity):
maptable[first_lpn]['write_flag'] = 1
maptable[first_lpn]['access_time'] = float(tmp[0])
first_lpn = first_lpn + 1
maptable=pd.DataFrame(maptable,columns=['access_time', 'update_interval', 'update_count', 'update_avg', "update_min", "update_max", "write_flag"])
print("trace:%s update_avg:%fns, update_avg / (10 ** 6):%fs,update_min:%fns,update_max:%fns" % (tracename, maptable['update_avg'].mean(), maptable['update_avg'].mean() / (10 ** 6), maptable['update_min'].mean(), maptable['update_max'].mean()))
maptable.sort_values("update_avg",inplace=True)
print("<0的百分比:%f >0和<0的百分比:%f "%(len(maptable[(maptable['update_avg']==0.0)].index)/len(maptable.index),len(maptable[(maptable['update_avg']>0.0) & (maptable['update_avg']<100.0*10**6)].index)/len(maptable.index)))
# +
import numpy as np
updatetable=maptable[(maptable['update_count']!=0)]#排除更新时间为0的
print(len(updatetable.index),updatetable['update_avg'].max())
print(len(updatetable.index),updatetable['update_avg'].min())
dx=0.01
#x=np.arange(0, int(updatetable['update_avg'].max()), dx)
updatetable['update_avg'].plot(kind='kde',style='k--')
import seaborn as sns
sns.set()
#sns.kdeplot(updatetable['update_avg'].tolist())
sns.distplot(updatetable['update_avg'].tolist(),hist=False,kde_kws={'clip': (0.0, int(updatetable['update_avg'].max()))})
#sns.distplot(updatetable['update_avg'].tolist(),kde=False)
# -
updatetable=maptable[(maptable['update_avg']>0.0)]
plt.figure()
updatetable['update_avg'].plot(kind='kde',style='k--')
#updatetable['update_avg'].hist( cumulative = True )
maptable = []
| traces/trace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Part I
# #### Analyze Revenue Data From 2 Airline Companies
airline_1 <- read.csv(url('https://raw.githubusercontent.com/ma010/misc/master/airline1.csv'))
airline_1
airline_2 <- read.csv(url('https://raw.githubusercontent.com/ma010/misc/master/airline2.csv'))
airline_2
# #### Question 1. What is the average revenue for Airline1 in different regions?
# #### Question 2. Where does Airline2 have the largest revenue?
# #### Question 3. For periods and regions the two airlines both served, what is the combined revenue of Airline 1 and Airline 2 for each region in each period? (Join Tables)
| Analytics_R_online.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cryptography
import hashlib
import os
import hmac
import base58
from bitstring import BitArray
from ecdsa import SigningKey, SECP256k1
from ecdsa.util import string_to_number, number_to_string
# +
N_WORDS = 24
PASSPHRASE = ""
INITIAL_ENT = 256
with open("wordlist.txt", mode="r") as wordlist_file:
wordlist = wordlist_file.read().split("\n")
# +
def return_first_n_bits(n, bts):
bits = BitArray(bts)
return bits[:n].uint
def get_word_indices(wordlist_entropy):
indices = []
bits = BitArray(wordlist_entropy)
i = 0
while i < len(bits):
indices.append(bits[i: i + 11].uint)
i += 11
return indices
# -
# # Bitcoin wallet generation demo
#
# This Jupyter Notebook serves as a step-by-step demonstration of how a Bitcoin wallet is generated along with a bunch of public addresses.
#
# __Disclaimer: You could, theoretically, use this as a way to generate your paper wallet, however this is not advised, since the code has not undergone any security review whatsover.__
# ## Generate Initial Entropy
# Source: https://github.com/bitcoin/bips/blob/master/bip-0039.mediawiki
#
# First we generate an initial entropy of `INITIAL_ENT / 8` bits and its checksum. By joining these we get an entropy that'll be used to generate the word list.
#
# __ Notice, that we use `os.urandom` as our source of entropy. The [documentation states](https://docs.python.org/3/library/os.html#os.urandom) that this should be secure enough, but ultimately depends on your system configuration.__
initial_entropy = os.urandom(INITIAL_ENT // 8)
initial_digest = hashlib.sha256(initial_entropy).digest()
initial_checksum = return_first_n_bits(INITIAL_ENT // 32, initial_digest)
wordlist_entropy = initial_digest + bytes([initial_checksum])
print(f"Our wordlist entropy is {wordlist_entropy.hex()}")
word_indexes = get_word_indices(wordlist_entropy)
word_seed = [wordlist[i] for i in word_indexes]
print(f"Word seed is: {word_seed}")
# We use the wordlist and an optional `PASSPHRASE` to generate the seed for the wallet key generation. The seed is generated using PBKDF2 key generation function.
seed = hashlib.pbkdf2_hmac("sha512", ("".join(word_seed) + PASSPHRASE).encode("utf-8"), ("mnemonic" + PASSPHRASE).encode("utf-8"), 2048)
print(f"Our seed is {seed.hex()}")
# ## Generating the wallet
# Primary resource: https://github.com/bitcoin/bips/blob/master/bip-0032.mediawiki
#
# Using the seed, we first generate the master key. We can subsequently use this key to generate child private and public keys for all our accounts.
# +
# Definition of functions as presented in BIP-32. These include key derivation function as well as helper functions.
def serialise_bytes(key, key_type, chain_code, depth=0x00, fingerprint=bytes([0] * 4), child_no=bytes([0] * 4)):
if key_type == "pub":
version = 0x0488B21E.to_bytes(4, "big")
elif key_type == "priv":
version = 0x0488ADE4.to_bytes(4, "big")
key = bytes([0]) + key.to_bytes(32, "big")
return version + bytes([depth]) + fingerprint + child_no + chain_code + key
def key_to_b58(serialised_key):
fingerprint = hashlib.sha256(hashlib.sha256(serialised_key).digest()).digest()[:4]
return base58.b58encode(serialised_key + fingerprint)
def key_fingerprint(key):
key = point_to_pk(key)
ripemd = hashlib.new("ripemd160")
ripemd.update(hashlib.sha256(key).digest())
digest = ripemd.digest()
return digest[:4]
def parse_256(p):
return int.from_bytes(p, "big")
def point_to_pk(point):
x = number_to_string(point.x(), SECP256k1.order)
y = number_to_string(point.y(), SECP256k1.order)
return bytes([((y[31] & 1) + 2)]) + x
def derive_key_and_chain(key, msg):
digest = hmac.new(key=key, msg=msg, digestmod=hashlib.sha512).digest()
return int.from_bytes(digest[:32], "big"), digest[32:]
def print_key_parts(key):
print(key[:8])
print(key[8:10])
print(key[10:18])
print(key[18:26])
print(key[26:90])
print(key[90:156])
print(key[156:])
def point_mul(p):
return SECP256k1.generator * p
def key_mod(key):
return key % SECP256k1.order
# Derivation function definitions
def CKDpriv(key, chain, i):
"""Derives a child private key from a parent private key"""
if i >= 2 ** 31:
# we are generating a hardened key
child_key, child_chain = derive_key_and_chain(chain, bytes([0]) + key.to_bytes(32, "big") + i.to_bytes(4, "big"))
else:
child_key, child_chain = derive_key_and_chain(chain, point_to_pk(point_mul(key)) + i.to_bytes(4, "big"))
child_key = key_mod(child_key + key)
return child_key, child_chain
def CKDpub(key, chain, i):
if i >= 2 ** 31:
raise ValueError("Derivation of pubkey from a parent pubkey is not defined")
child_key, child_chain = derive_key_and_chain(chain, point_to_pk(key) + i.to_bytes(4, "big"))
child_key = point_mul(int.from_bytes(child_key, "big")) + key
return child_key, child_chain
# -
seed = bytes.fromhex("000102030405060708090a0b0c0d0e0f") # this is a seed from the first test vector in BIP-32
master_key, master_chain = derive_key_and_chain(b"Bitcoin seed", seed)
print(f"Your master key is {master_key.to_bytes(32, 'big').hex()} and your master chain code is {master_chain.hex()}")
serialised_master = serialise_bytes(master_key, "priv", master_chain, fingerprint=bytes([0] * 4), child_no=bytes([0] * 4))
master_xpriv = key_to_b58(serialised_master)
print(f"Your master xpriv is {master_xpriv}")
master_public_key = point_mul(master_key)
serialised_master_public = serialise_bytes(point_to_pk(master_public_key), "pub", master_chain, fingerprint=bytes([0] * 4), child_no=bytes([0] * 4))
print(f"Your master xpub is {key_to_b58(serialised_master_public)}")
# This gives us the master public and private keys of our wallet. From these, we can subsequently generate our "sub-accounts". Examples bellow.
i = 2 ** 31
child_privk, child_chain = CKDpriv(master_key, master_chain, i)
serialised_child_priv = serialise_bytes(child_privk, "priv", child_chain, 1, key_fingerprint(master_public_key), i.to_bytes(4, "big"))
print(key_to_b58(serialised_child_priv))
key_to_b58(serialised_child_priv) == "<KEY>"
child_pub = point_mul(child_privk)
serialised_child_public = serialise_bytes(point_to_pk(child_pub), "pub", child_chain, 1, key_fingerprint(master_public_key), i.to_bytes(4, "big"))
print(key_to_b58(serialised_child_public))
# By applying CKDpriv and CKDpub recursively we generate the so-called _key tree_. The root of the tree is the master key, the leaf nodes are the actual wallet keys used for signing and verifying transactions and the inner nodes represent the intermediary parent keys. This structure allows for wallets that hold multiple accounts or even multiple coins (as long as they stick to the same key-derivation algorithms) without the need to generate multiple seeds.
#
# A standard notation for this is of the following form:
#
# ```
# m / i' / j / ... / k
# ```
#
# where `m` is a placeholder for the root key and further components are the `i`th, `j`th (and so on) keys in the hierarchy. The apostrophe denotes a hardened key.
#
# ## Account hierarchy
# Reference:
# * https://github.com/bitcoin/bips/blob/master/bip-0044.mediawiki
# * https://github.com/bitcoin/bips/blob/master/bip-0049.mediawiki
#
# BIP-44 standarises the hierarchy in deterministic wallets.
#
# ```
# m / purpose' / coin_type' / account' / change / address_index
# ```
#
# The semantics behind most of the components are quite obvious from the name, maybe apart from `purpose`. `Purpose` determines the structure of the rest of the path. Usually it is set to `44` (as in BIP-44), however with the introduction of SegWit, constant `49` is used (as in BIP-49). Since we have to use hardened keys, the actual `int` representation will be `2^31 + 49`.
# +
def to_hardened_int(i):
return 2 ** 31 + i
PURPOSE = to_hardened_int(49)
COIN_TYPE = to_hardened_int(0)
ACCOUNT = to_hardened_int(0)
# -
| bitcoin_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started
# ## Overview
#
# MLRun Graphs enable building and running DAGs (directed acyclic graph). Graphs are composed of individual steps.
# The first graph element accepts an `Event` object, transform/process the event and pass the result to the next steps
# in the graph. The final result can be written out to some destination (file, DB, stream, etc.) or returned back to the caller
# (one of the graph steps can be marked with `.respond()`).
#
# The serving graphs can be composed of [pre-defined graph steps](./available-steps.md), block-type elements (model servers, routers, ensembles, data readers and writers, data engineering tasks, validators, etc.), [custom steps](./writing-custom-steps.ipynb), or from native python classes/functions. Graphs can auto-scale and span multiple function
# containers (connected through streaming protocols).
#
# 
#
# Different steps can run on the same local function, or run on a remote function. You can call existing functions from the graph and reuse them from other graphs, as well as scale up and down different components individually.
#
# Graphs can run inside your IDE or Notebook for test and simulation, and can then be deployed
# into production serverless pipeline with a single command. Serving graphs are built on
# top of [Nuclio](https://github.com/nuclio/nuclio) (real-time serverless engine), MLRun Jobs,
# [MLRun Storey](<https://github.com/mlrun/storey>) (native Python async and stream processing engine),
# and other MLRun facilities.
# ## Basic Example
#
# This example uses a custom class and custom function. See [custom steps](./writing-custom-steps.ipynb) for more details.
#
# - [Steps](#steps)
# - [Create a function](#create-a-function)
# - [Build the graph](#build-the-graph)
# - [Visualize the graph](#visualize-the-graph)
# - [Test the function](#test-the-function)
# - [Deploy the function](#deploy-the-function)
# - [Test the deployed function](#test-the-deployed-function)
# ### Steps
#
# The following code defines basic steps that illustrate building a graph. These steps are:
#
# - **`inc`**: increments the value by 1.
# - **`mul`**: multiplies the value by 2.
# - **`WithState`**: class that increments an internal counter, prints an output, and adds the input value to the current counter.
# +
# mlrun: start-code
def inc(x):
return x + 1
def mul(x):
return x * 2
class WithState:
def __init__(self, name, context, init_val=0):
self.name = name
self.context = context
self.counter = init_val
def do(self, x):
self.counter += 1
print(f"Echo: {self.name}, x: {x}, counter: {self.counter}")
return x + self.counter
# mlrun: end-code
# -
# ### Create a function
#
# Now take the code above and create an MLRun function called `serving-graph`.
import mlrun
fn = mlrun.code_to_function("simple-graph", kind="serving", image="mlrun/mlrun")
graph = fn.set_topology("flow")
# ### Build the graph
#
# Use `graph.to()` to chain steps. Use `.respond()` to mark that the output of that step is returned to the caller
# (as an http response). By default the graph is async with no response.
graph.to(name="+1", handler='inc')\
.to(name="*2", handler='mul')\
.to(name="(X+counter)", class_name='WithState').respond()
# ### Visualize the graph
#
# Using the `plot` method, you can visualize the graph.
graph.plot(rankdir='LR')
# ### Test the function
#
# Create a mock server and test the graph locally. Since this graph accepts a numeric value as the input, that value is provided
# in the `body` parameter.
server = fn.to_mock_server()
server.test(body=5)
# Run the function again. This time, the counter should be 2 and the output should be 14.
server.test(body=5)
# ### Deploy the function
#
# Use the `deploy` method to deploy the function.
fn.deploy(project='basic-graph-demo')
# ### Test the deployed function
#
# Use the `invoke` method to call the function.
fn.invoke('', body=5)
fn.invoke('', body=5)
| docs/serving/getting-started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (dataSc)
# language: python
# name: datasc
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Imports" data-toc-modified-id="Imports-1"><span class="toc-item-num">1 </span>Imports</a></span></li><li><span><a href="#Load-data" data-toc-modified-id="Load-data-2"><span class="toc-item-num">2 </span>Load data</a></span></li><li><span><a href="#Numerical-and-Categorical-Plots" data-toc-modified-id="Numerical-and-Categorical-Plots-3"><span class="toc-item-num">3 </span>Numerical and Categorical Plots</a></span></li></ul></div>
# -
# # Imports
# +
import numpy as np
import pandas as pd
import seaborn as sns
import bhishan
from bhishan import bp
import matplotlib.pyplot as plt
# %load_ext autoreload
# %load_ext watermark
# %autoreload 2
# %watermark -a "<NAME>" -d -v -m
# %watermark -iv
# -
# # Load data
titanic = sns.load_dataset('titanic')
titanic.head()
# # Numerical and Categorical Plots
df = titanic
# df.bp.plot_num('age',print_=True,ms='seaborn-darkgrid')
df.bp.plot_num(num='age',print_=True,ms='dark_background')
df.bp.plot_cat('pclass',ms=-1)
df.bp.plot_num_num('age','fare',xticks1=range(0,90,10),ms=-1,
xticks2=range(0,600,50),rot=90,figsize=(12,12))
df.bp.plot_num_cat('pclass','sex',save=True,show=True)
df.bp.plot_cat_num('pclass','age',save=True,show=True,ms='fast')
df.bp.plot_cat_cat('pclass','survived',save=True,show=True,ms=-1)
| examples/pandas_api_num_and_cat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="c7CN1KdyzYZN"
#
#
# * <NAME>
# * November 20th, 2020
# * Final Project Applied Computing
# * Code for the selection of the final model
#
#
#
#
# + [markdown] id="zzj1uhWnHg-r"
# This is code was for the selection of the final model
# + [markdown] id="CaVG0Z2lUNZK"
# Import CSV file
# + id="_8_Ct2i-T2yl" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 72} outputId="771a104c-a08a-411e-ad07-e1c13160bc82"
from google.colab import files
uploaded = files.upload()
# + [markdown] id="iGKCsmk7URFn"
# Import Libraries and Save CSV file data into a pandas Data Frame
# + id="JDYafPBCUg5d" colab={"base_uri": "https://localhost:8080/"} outputId="981cf0e5-a0d1-489c-9dec-e376a15cde23"
import io
import numpy as np #Library to manage arrays
import pandas as pd #Lirary for mange dataframes
import matplotlib.pyplot as plt # Library to make plots.
#Import csv into a dataframe
df = pd.read_csv(io.BytesIO(uploaded['data.csv']))
print("Complete Data shape:", df.shape)
print("Data shape withput NA values",df.dropna().shape)
# + [markdown] id="bCywCoO8Unm5"
# Count the number of NA values on each variable
# + id="GvVcNaICUuXw" colab={"base_uri": "https://localhost:8080/"} outputId="449d53c1-6a02-482f-a474-116552dc3e77"
columns_na = df.isnull().sum(axis = 0)
pd.set_option('display.max_rows', None)
columns_na
# + [markdown] id="8NDYYmFRa7uF"
# Remove NA values from the database
# + id="DmXL9BTNU2_g" colab={"base_uri": "https://localhost:8080/"} outputId="5acd9a7b-dc41-4b5c-a7ea-6abb984c8780"
df_n = df.dropna(axis = 0, how ='any')
print(df_n.shape)
# + id="dfyUmBi1uZGi" colab={"base_uri": "https://localhost:8080/"} outputId="ccdc2b5b-3cad-4634-bfa7-742635959386"
df_n.info()
# + [markdown] id="6g_og7jmbAZf"
# Percentage representation of class in the target variable
# + id="nRn6rhrqU6j4" colab={"base_uri": "https://localhost:8080/", "height": 368} outputId="0e8f1006-f8fc-47d5-8728-ecccf9afc0b0"
classes = df_n['Winner'].value_counts()
print(classes)
print(classes.index.values)
ax = classes.plot.bar(rot = 0, color=["red", "blue", "green"])
for p in ax.patches:
ax.annotate(str(p.get_height()), (p.get_x() * 1.005, p.get_height() * 1.005))
ax.set_ylabel("Counts")
# + id="bqcqIUT8U9gH" colab={"base_uri": "https://localhost:8080/"} outputId="c74e8e7f-1e7f-4777-e6fa-896315e7ad8d"
df_n['Winner'].value_counts(normalize=True)
# + [markdown] id="-X5Uy9yWbXc3"
# Process categorical variables
# + id="QmbvGzPwqlrQ" colab={"base_uri": "https://localhost:8080/"} outputId="66e64544-7805-4615-9202-2b9bba389373"
print('R_fighter numbers:',df_n['R_fighter'].value_counts().shape[0])
print('B_fighter numbers:',df_n['B_fighter'].value_counts().shape[0])
print('Referee numbers:',df_n['Referee'].value_counts().shape[0])
print('date:',df_n['date'].value_counts().shape[0])
print('location:',df_n['location'].value_counts().shape[0])
print('weight_class:',df_n['weight_class'].value_counts().shape[0])
print('R_Stance:',df_n['R_Stance'].value_counts().shape[0])
print('B_Stance:',df_n['B_Stance'].value_counts().shape[0])
# + id="YJ4lhY5CVAp_" colab={"base_uri": "https://localhost:8080/"} outputId="1362e543-4d8f-4144-a287-9ac418685452"
df_n = df_n.drop(['R_fighter', 'B_fighter', 'Referee', 'date', 'location'], axis=1)
#df_n['R_fighter'] = df_n['R_fighter'].astype('category').cat.codes
#df_n['B_fighter'] = df_n['B_fighter'].astype('category').cat.codes
#df_n['location'] = df_n['location'].astype('category').cat.codes
#f_n['Referee'] = df_n['Referee'].astype('category').cat.codes
#df_n['date'] = df_n['date'].astype('category').cat.codes
df_n['title_bout'] = df_n["title_bout"].astype('category').cat.codes
df_n = pd.get_dummies(df_n, columns=['R_Stance', 'B_Stance', 'weight_class'])
print(df_n.head())
print(df_n.shape)
# + [markdown] id="jHOqx6wObdyN"
# Extract target variable from the database
# + id="nc5gYPJZVHww" colab={"base_uri": "https://localhost:8080/"} outputId="07a4957c-6ab0-4e89-b020-ca746ca13a9e"
df_filtered = df_n
# Select input variables and separate target variable
features = df_filtered.loc[:, df_n.columns != 'Winner']
print(features.shape)
# + [markdown] id="5PVHAtOg__sW"
# In this next section, the machine learning techniques for supervised learning were tested with all the input variables of the database.
# + [markdown] id="efQvSz1IVnZA"
# Decision Tree Classifier with all the variables of the database as inputs
# + id="Hld7RcNSVi54" colab={"base_uri": "https://localhost:8080/"} outputId="34df6935-2035-4628-90d1-ead80c15d1f5"
from sklearn.tree import DecisionTreeClassifier # Import Decision Tree Classifier
from sklearn.model_selection import train_test_split # Import train_test_split function
from sklearn.metrics import classification_report #Library for classification report
from sklearn.metrics import confusion_matrix # Library to generate the confusion matrix of the model.
# Features and target variable
X = features.values
y = df_filtered.loc[:,['Winner']].values
print("Shape of input and output variables data", "X:" ,X.shape, "y:" ,y.shape)
# Split data in train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1) # 80% training and 20% test
# Create Decision Tree classifer object
dt = DecisionTreeClassifier( criterion="gini", max_depth= 25)
# Train Decision Tree Classifer
dt = dt.fit(X_train,y_train)
#Predict the response for test dataset
print(X_train.shape, X_test.shape)
print("Accuracy with traning data set for Decision Tree: {:.4f}".format(dt.score(X_train, y_train)))
print("Accuracy with test data set for Desicion Tree: {:.4f}".format(dt.score(X_test, y_test)))
pred = dt.predict(X_test)
print(confusion_matrix(y_test, pred)) # Print confusion matrix
print(classification_report(y_test, pred)) # Print classification report
# Generate Classification Report and save it in CSV file
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_DT_all.csv")
# + [markdown] id="NF1MfbXv3z34"
# Clear input and output variables
# + id="U1S3v6Lo3dKz"
del X, X_test, X_train,y, y_test, y_train, dt, pred, report
# + [markdown] id="bHJaj64xWnQI"
# K NN Classifier using all the variables as predictors
# + id="V_icvzOiV1RI" colab={"base_uri": "https://localhost:8080/"} outputId="4312ba77-4057-4a09-c772-6cd90e90cc1b"
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.neighbors import KNeighborsClassifier
# Input and output variables for the KNN classifier.
X = features.values
y = df_filtered.loc[:,['Winner']].values
print("Shape of input and output variables ", "X:" ,X.shape, "y:" ,y.shape)
#Split data in train and test set.
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0,test_size=0.2)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
#Establish the number of neightbors for the KNN classiier.
n_neighbors = 7
knn = KNeighborsClassifier(n_neighbors)
knn.fit(X_train, y_train.ravel())
print('Accuracy of K-NN classifier on training set: {:.4f}'
.format(knn.score(X_train, y_train)))
print('Accuracy of K-NN classifier on test set: {:.4f}'
.format(knn.score(X_test, y_test)))
pred = knn.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_KNN_all.csv")
# + [markdown] id="uWq85XhPdMpJ"
# Reset variables
# + id="Cj_hWr8m4uZU"
del X, X_test, X_train,y, y_test, y_train, knn, pred, report
# + [markdown] id="21De0LN7WjiI"
# Support Vector Machine with all the variables as inputs
# + id="VjP9YLboV9Xa" colab={"base_uri": "https://localhost:8080/"} outputId="fd38c08a-3bf8-456b-8bdb-01ad0be3aba5"
# Import train_test_split function
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
#Import svm model
from sklearn import svm
X = features.values
y = df_filtered.loc[:,['Winner']].values
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1) # 80% training and 20% test
X_train = StandardScaler().fit_transform(X_train)
X_test = StandardScaler().fit_transform(X_test)
#Create a svm Classifier
svc = svm.SVC(kernel='rbf') # Radial Basis Function Kernel
#Train the model using the training sets
svc.fit(X_train, y_train.ravel())
# Model Accuracy: how often is the classifier correct?
print('Accuracy of SVM classifier on training set: {:.4f}'
.format(svc.score(X_train, y_train)))
print('Accuracy of SVM classifier on test set: {:.4f}'
.format(svc.score(X_test, y_test)))
pred = svc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_SVM_all.csv")
# + id="dazPiOLc6Erq"
del X, X_test, X_train,y, y_test, y_train, svc, pred, report
# + [markdown] id="9a2HghZEWgkP"
# Random Forest Using all the variables as inputs for the model
# + id="5_8l-BMPWPnx" colab={"base_uri": "https://localhost:8080/"} outputId="d43df899-512c-4964-f633-881655a704a6"
#Import Random Forest Model
from sklearn.ensemble import RandomForestClassifier
X = features.values
y = df_filtered.loc[:,['Winner']].values
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,
random_state=1) # 70% training and 30% test
#Create a Gaussian Classifier
rfc=RandomForestClassifier(n_estimators=120, criterion='gini', max_depth=25)
#Train the model using the training sets y_pred=clf.predict(X_test)
rfc.fit(X_train,y_train.ravel())
print('Accuracy of Random classifier on training set: {:.4f}'
.format(rfc.score(X_train, y_train)))
print('Accuracy of Random classifier on test set: {:.4f}'
.format(rfc.score(X_test, y_test)))
#Variables of most importance according to the Gini importance
feature_imp = pd.Series(rfc.feature_importances_,index=features.columns).sort_values(ascending=False)
pred = rfc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_Random_Forest_all.csv")
# + [markdown] id="0BwAR9xrdYr9"
# Reset variables
# + id="MGj3ESS06mnw"
del X, X_test, X_train,y, y_test, y_train, rfc, pred, report
# + [markdown] id="sLXZNZSKVjsI"
# Principal Component Analysis for Feature Selection
# + id="Z7cBJLw1VM8o" colab={"base_uri": "https://localhost:8080/"} outputId="6c06acbb-f38d-4673-a49b-5a0ad85ee4f1"
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
# Separating out the features
x = features
# Separating out the target
y = df_filtered.loc[:,['Winner']].values
# Standardizing the features
x = StandardScaler().fit_transform(x)
Winner = df_filtered.loc[:,['Winner']]
Winner = Winner.reset_index(drop=True)
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(x)
print("Variance percentage of first component:",pca.explained_variance_ratio_[0])
print("Variance percentage of second component:",pca.explained_variance_ratio_[1])
print("Variance percentage of first two components:",sum(pca.explained_variance_ratio_))
principalDf = pd.DataFrame(data = principalComponents,columns = ['principal component 1', 'principal component 2'])
finalDf = pd.concat([principalDf, Winner], axis = 1)
# + id="XlIpUg21VWCo" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="a65dd7cd-74ac-428e-b184-50f773500a59"
import matplotlib.pyplot as plt
plt.plot(pca.explained_variance_ratio_)
plt.title('PCA')
plt.xlabel('Percent of Variance Explained')
plt.ylabel('.')
plt.show()
# + [markdown] id="jUfUkzZZLYvK"
# Plot first two principal components
# + id="yOL_DanJVcr4" colab={"base_uri": "https://localhost:8080/", "height": 522} outputId="39889c87-52f3-4522-8fe6-958af0bdeb5b"
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Principal Component 1', fontsize = 15)
ax.set_ylabel('Principal Component 2', fontsize = 15)
ax.set_title('2 component PCA', fontsize = 20)
targets = ['Red', 'Blue','Draw']
colors = ['r', 'b','g']
for target, color in zip(targets,colors):
indicesToKeep = finalDf['Winner'] == target
ax.scatter(finalDf.loc[indicesToKeep, 'principal component 1']
, finalDf.loc[indicesToKeep, 'principal component 2']
, c = color
, s = 50)
ax.legend(targets)
ax.grid()
# + [markdown] id="je6jj2ldRL5g"
# Compute the loading scores of each varible in the first and second Principal Components
# + id="Z5apD0huBXfu" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="6dda2509-59de-4575-a43c-1e7b8b8c339d"
df_npca = pd.DataFrame(pca.components_, columns=list(features.columns))
df_npca.head(3)
# + [markdown] id="A4YBn5JiRUUO"
# Sort the values of the loading scores and display them in a bar plot from highest to lowest
# + id="2qBFWsXEXELJ" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="b9759532-fbaf-41f9-8838-d4d06fbe7a00"
loading_scores = pd.Series(pca.components_[0], index=features.columns)
soarted_loading_scores = loading_scores.abs().sort_values(ascending = False)
size = 15
ind = soarted_loading_scores[0:size].index.values
print(loading_scores[ind])
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Creating a bar plot
sns.barplot(x=soarted_loading_scores[0:size], y= soarted_loading_scores[0:size].index)
# Add labels to your graph
plt.xlabel('Loading Score')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.legend()
plt.show()
# + [markdown] id="nSriURlJL2BS"
# Extract the variables from the original data base with the highest loading scores
# + id="wDaWt7isXl5Y" colab={"base_uri": "https://localhost:8080/", "height": 324} outputId="d277a744-9cc1-42f9-a74e-0c6176311aa3"
features_nf = df_filtered[ind]
print(features_nf.shape)
features_nf.describe()
# + [markdown] id="_1kUBw80z17q"
# Decision Tree with variables of PCA feature selection
# + id="V7Z5f1P9XWVg" colab={"base_uri": "https://localhost:8080/"} outputId="e7c8beb4-31bb-4225-806b-98df85654515"
X = features_nf.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1234) # 80% training and 20% test
print(X_train.shape, X_test.shape)
# Create Decision Tree classifer object
dt = DecisionTreeClassifier(criterion="gini", max_depth=6)
# Train Decision Tree Classifer
dt = dt.fit(X_train,y_train)
#Predict the response for test dataset
print('Accuracy of Decision Tree classifier on training set: {:.4f}'
.format(dt.score(X_train, y_train)))
print('Accuracy of Decision Tree classifier on test set: {:.4f}'
.format(dt.score(X_test, y_test)))
pred = dt.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_DT_PCA.csv")
# + [markdown] id="0c1WP7b1EInR"
# Erase variables
# + id="THBse4FZ8D0Y"
del X, X_test, X_train,y, y_test, y_train, dt, pred, report
# + [markdown] id="GavnR-Gbz4p4"
# KNN Nearest Neightbor trained with important features according to the loading scores of each variables in the first principal component of PCA.
# + id="ICN-RzxEkL8q" colab={"base_uri": "https://localhost:8080/"} outputId="02d892d3-de02-4707-ff7c-5939946163e2"
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
#Feature Importance
X = features_nf.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1)
print(X_test.shape, X_train.shape)
#Standarized data
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
n_neighbors = 7
knn = KNeighborsClassifier(n_neighbors)
knn.fit(X_train, y_train.ravel())
print('Accuracy of K-NN classifier on training set: {:.4f}'
.format(knn.score(X_train, y_train)))
print('Accuracy of K-NN classifier on test set: {:.4f}'
.format(knn.score(X_test, y_test)))
pred = knn.predict(X_test)
print(X_test.shape)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_KNN_PCA.csv")
# + [markdown] id="lrJ5CRAGTUK8"
# Reset variables
# + id="EM4nYyL78bSh"
del X, X_test, X_train,y, y_test, y_train, knn, pred, report
# + [markdown] id="mZLeweVwYLfu"
# SVM with feature reduction of PCA
# + id="ifP4S2ZAX1Qn" colab={"base_uri": "https://localhost:8080/"} outputId="c9cd7ee8-7e00-4c80-802e-3700088c7413"
# Import train_test_split function
from sklearn.model_selection import train_test_split
from sklearn import svm
X = features_nf.values
y = df_filtered.loc[:,['Winner']].values
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 25) # 80% training and 30% test
X_train = StandardScaler().fit_transform(X_train)
X_test = StandardScaler().fit_transform(X_test)
print(X_train.shape, X_test.shape)
#Create a svm Classifier
svc = svm.SVC(kernel='rbf') # Linear Kernel
#Train the model using the training sets
svc.fit(X_train, y_train.ravel())
#Predict the response for test dataset
print('Accuracy of Support Vector classifier on training set: {:.4f}'
.format(svc.score(X_train, y_train)))
print('Accuracy of Support Vector classifier on test set: {:.4f}'
.format(svc.score(X_test, y_test)))
pred = svc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_SVM_PCA.csv")
# + [markdown] id="WvQFpvFJTQiL"
# Reset variables
# + id="MZQ-Ui6eEggE"
del X, X_test, X_train,y, y_test, y_train, svc, pred, report
# + [markdown] id="nTTgwus7TCFA"
# Random Forest trained with important features according to the loading scores of each variables in the first principal component of PCA.
# + id="ckzFhxtaYvcw" colab={"base_uri": "https://localhost:8080/"} outputId="ef2bc5a3-c546-4785-b9c0-e2d04117e447"
#Import Random Forest Model
from sklearn.ensemble import RandomForestClassifier
X = features_nf.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1234)
#Create a Gaussian Classifier
rfc=RandomForestClassifier(n_estimators=120, criterion='gini', max_depth=6)
#Train the model using the training sets y_pred=clf.predict(X_test)
rfc.fit(X_train,y_train.ravel())
print(X_train.shape,X_test.shape)
print('Accuracy of Random classifier on training set: {:.4f}'
.format(rfc.score(X_train, y_train)))
print('Accuracy of Random classifier on test set: {:.4f}'
.format(rfc.score(X_test, y_test)))
pred = rfc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_RF_PCA.csv")
# + [markdown] id="ePfoe93KS1co"
# Reset Variables
# + id="zdoLoAMOKh1Q"
del X, X_test, X_train,y, y_test, y_train, rfc, pred, report
# + [markdown] id="UoZKGDB9S4JV"
# Select features according to the Gini importance
# + id="gJW7I8GwKorP" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="bad4b859-9268-43f3-b8d1-76f4167fbb73"
size = 15
features_rf = df_filtered[feature_imp.index[0:size]]
"""import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# Creating a bar plot
sns.barplot(x=feature_imp[0:size], y= feature_imp[0:size].index)
# Add labels to your graph
plt.xlabel('Feature Importance Score')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.legend()
plt.show()
print(features_rf.shape)
features_rf.describe()
print(features_rf.columns)"""
# + [markdown] id="e2Khfp-wSqlQ"
# Decision Tree trained with important features according to the Gini importance
# + id="D5ISpV-hLFCB" colab={"base_uri": "https://localhost:8080/", "height": 447} outputId="167f3041-f80a-41ea-8674-abcc90da1abe"
inputs = ['R_age', 'R_avg_opp_SIG_STR_pct', 'R_avg_opp_SIG_STR_landed',
'B_avg_DISTANCE_landed', 'B_avg_SIG_STR_att', 'R_avg_opp_HEAD_landed',
'B_age', 'B_avg_opp_TOTAL_STR_landed', 'R_avg_opp_TOTAL_STR_landed',
'B_avg_HEAD_att', 'R_avg_GROUND_att', 'B_avg_opp_SIG_STR_pct',
'B_avg_BODY_att', 'R_avg_GROUND_landed', 'R_avg_opp_DISTANCE_landed']
print(inputs)
dt_final =df_filtered[inputs]
X = dt_final.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1234) # 80% training and 20% test
print(X_train.shape, X_test.shape)
# Create Decision Tree classifer object
dt = DecisionTreeClassifier(criterion="gini", max_depth=6)
# Train Decision Tree Classifer
dt = dt.fit(X_train,y_train)
#Predict the response for test dataset
print('Accuracy of Decision Tree classifier on training set: {:.4f}'
.format(dt.score(X_train, y_train)))
print('Accuracy of Decision Tree classifier on test set: {:.4f}'
.format(dt.score(X_test, y_test)))
pred = dt.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_DT_RFF.csv")
f = dt.predict_proba(X_test)
prob_DF = pd.DataFrame(f, columns=dt.classes_)
prob_DF.head(3)
# + [markdown] id="Sm0fP5k3Sl8W"
# Reset Variables
# + id="eO9MFWOnLN1W"
del X, X_test, X_train,y, y_test, y_train, dt, pred, report
# + [markdown] id="UZ6GFy09SfJ5"
# KNN classifier trained with important features according to the Gini importance
# + id="s3bXhyD6LTqL" colab={"base_uri": "https://localhost:8080/"} outputId="7c67cb4d-59d7-43cb-912c-df399d606ad9"
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
inputs = ['R_age', 'R_avg_opp_SIG_STR_pct', 'R_avg_opp_SIG_STR_landed',
'B_avg_DISTANCE_landed', 'B_avg_SIG_STR_att', 'R_avg_opp_HEAD_landed',
'B_age', 'B_avg_opp_TOTAL_STR_landed', 'R_avg_opp_TOTAL_STR_landed',
'B_avg_HEAD_att', 'R_avg_GROUND_att', 'B_avg_opp_SIG_STR_pct',
'B_avg_BODY_att', 'R_avg_GROUND_landed', 'R_avg_opp_DISTANCE_landed']
print(inputs)
dt_final =df_filtered[inputs]
X = dt_final.values
y = df_filtered.loc[:,['Winner']].values
#Split data in train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1)
print(X_test.shape, X_train.shape)
#Scale the data
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
n_neighbors = 7
knn = KNeighborsClassifier(n_neighbors)
knn.fit(X_train, y_train.ravel())
print('Accuracy of K-NN classifier on training set: {:.4f}'
.format(knn.score(X_train, y_train)))
print('Accuracy of K-NN classifier on test set: {:.4f}'
.format(knn.score(X_test, y_test)))
pred = knn.predict(X_test)
print(X_test.shape)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_KNN_RFC.csv")
# + [markdown] id="McttcJsGSbyM"
# Reset Variables
# + id="VmAB7e3qMOoK"
del X, X_test, X_train,y, y_test, y_train, knn, pred, report
# + [markdown] id="1JqnbMxaSR8z"
# SVM classifier trained with important features according to the Gini importance
# + id="Mk8I0wMaMDQY" colab={"base_uri": "https://localhost:8080/"} outputId="f49e8266-45bf-4504-e99a-4834d708c173"
# Import train_test_split function
from sklearn.model_selection import train_test_split
from sklearn import svm
inputs = ['R_age', 'R_avg_opp_SIG_STR_pct', 'R_avg_opp_SIG_STR_landed',
'B_avg_DISTANCE_landed', 'B_avg_SIG_STR_att', 'R_avg_opp_HEAD_landed',
'B_age', 'B_avg_opp_TOTAL_STR_landed', 'R_avg_opp_TOTAL_STR_landed',
'B_avg_HEAD_att', 'R_avg_GROUND_att', 'B_avg_opp_SIG_STR_pct',
'B_avg_BODY_att', 'R_avg_GROUND_landed', 'R_avg_opp_DISTANCE_landed']
print(inputs)
dt_final =df_filtered[inputs]
X = dt_final.values
y = df_filtered.loc[:,['Winner']].values
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 25) # 80% training and 30% test
X_train = StandardScaler().fit_transform(X_train)
X_test = StandardScaler().fit_transform(X_test)
print(X_train.shape, X_test.shape)
#Create a svm Classifier
svc = svm.SVC(kernel='rbf') # Linear Kernel
#Train the model using the training sets
svc.fit(X_train, y_train.ravel())
#Predict the response for test dataset
print('Accuracy of Random classifier on training set: {:.4f}'
.format(svc.score(X_train, y_train)))
print('Accuracy of Random classifier on test set: {:.4f}'
.format(svc.score(X_test, y_test)))
pred = svc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_SVM_RFC.csv")
# + [markdown] id="6NjJG-b0SPEM"
# Reset variables
# + id="4UbM6mSUwEc8"
del X, X_test, X_train,y, y_test, y_train, svc, pred, report
# + [markdown] id="YntrY9ZGR58y"
# Random Forest with most important features according to the Gini importance
# + id="S45P2GiiwHNR" colab={"base_uri": "https://localhost:8080/"} outputId="cc68ce5f-a80a-4ca3-d212-4b1a4b690cb3"
#Import Random Forest Model
from sklearn.ensemble import RandomForestClassifier
inputs = ['R_age', 'R_avg_opp_SIG_STR_pct', 'R_avg_opp_SIG_STR_landed',
'B_avg_DISTANCE_landed', 'B_avg_SIG_STR_att', 'R_avg_opp_HEAD_landed',
'B_age', 'B_avg_opp_TOTAL_STR_landed', 'R_avg_opp_TOTAL_STR_landed',
'B_avg_HEAD_att', 'R_avg_GROUND_att', 'B_avg_opp_SIG_STR_pct',
'B_avg_BODY_att', 'R_avg_GROUND_landed', 'R_avg_opp_DISTANCE_landed']
print(inputs)
dt_final =df_filtered[inputs]
X = dt_final.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1234)
rfc=RandomForestClassifier(n_estimators=120, criterion='gini', max_depth=6)
rfc.fit(X_train,y_train.ravel())
print(X_train.shape,X_test.shape)
print('Accuracy of Random classifier on training set: {:.4f}'
.format(rfc.score(X_train, y_train)))
print('Accuracy of Random classifier on test set: {:.4f}'
.format(rfc.score(X_test, y_test)))
pred = rfc.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_RF_RFC.csv")
# + id="wjCO46QYys-c"
del X, X_test, X_train,y, y_test, y_train, rfc, pred, report
# + [markdown] id="IZsjz9pwWWEY"
# Final Model Training with variables based on the Gini Index Criteria
# + id="yH6KE4y0f5DL" colab={"base_uri": "https://localhost:8080/", "height": 690} outputId="942f5a62-417a-4358-8ba2-21d1717ffc46"
#This are the final predictors that will be tested for the model
inputs = ['R_age', 'R_avg_opp_SIG_STR_pct', 'R_avg_opp_SIG_STR_landed',
'B_avg_DISTANCE_landed', 'B_avg_SIG_STR_att', 'R_avg_opp_HEAD_landed',
'B_age', 'B_avg_opp_TOTAL_STR_landed', 'R_avg_opp_TOTAL_STR_landed',
'B_avg_HEAD_att', 'R_avg_GROUND_att', 'B_avg_opp_SIG_STR_pct',
'B_avg_BODY_att', 'R_avg_GROUND_landed', 'R_avg_opp_DISTANCE_landed']
print(inputs)
dt_final =df_filtered[inputs]
X = dt_final.values
y = df_filtered.loc[:,['Winner']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 1234) # 80% training and 20% test
print(X_train.shape, X_test.shape)
# Create Decision Tree classifer object
dt = DecisionTreeClassifier(criterion="gini", max_depth=6)
# Train Decision Tree Classifer
dt = dt.fit(X_train,y_train)
#Predict the response for test dataset
print('Accuracy of Decision Tree classifier on training set: {:.4f}'
.format(dt.score(X_train, y_train)))
print('Accuracy of Decision Tree classifier on test set: {:.4f}'
.format(dt.score(X_test, y_test)))
pred = dt.predict(X_test)
print(confusion_matrix(y_test, pred))
print(classification_report(y_test, pred))
report = classification_report(y_test, pred, output_dict=True)
report = pd.DataFrame(report).transpose()
report.to_csv("Report_DT_RFF.csv")
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
dot_data = StringIO()
export_graphviz(dt, out_file=dot_data,
filled=True, rounded=True,
special_characters=True, feature_names = features_rf.columns,class_names=['Blue','Draw', 'Red'])
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
graph.write_png('diabetes.png')
Image(graph.create_png())
f = dt.predict_proba(X_test)
prob_DF = pd.DataFrame(f, columns=dt.classes_)
prob_DF.head(10)
| Final_Project_Development_Code_A01331212_ver_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calculation of control fields for state-to-state transfer of a 2 qubit system using CRAB algorithm
# <NAME> (<EMAIL>)
# Example to demonstrate using the control library to determine control
# pulses using the ctrlpulseoptim.optimize_pulse_unitary function.
# The CRAB algorithm is used to optimize pulse shapes to minimize the fidelity
# error, which is equivalent maximising the fidelity to an optimal value of 1.
#
# The system in this example are two qubits, where the interaction can be
# controlled. The target is to perform a pure state transfer from a down-down
# state to an up-up state.
#
# The user can experiment with the timeslicing, by means of changing the
# number of timeslots and/or total time for the evolution.
# Different initial (starting) pulse types can be tried as well as
# boundaries on the control and a smooth ramping of the pulse when
# switching the control on and off (at the beginning and close to the end).
# The initial and final pulses are displayed in a plot
#
# An in depth discussion of using methods of this type can be found in [1,2]
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import datetime
# +
from qutip import Qobj, identity, sigmax, sigmaz, tensor, mesolve
import random
import qutip.logging_utils as logging
logger = logging.get_logger()
#Set this to None or logging.WARN for 'quiet' execution
log_level = logging.INFO
#QuTiP control modules
import qutip.control.pulseoptim as cpo
example_name = '2qubitInteract'
# -
# ### Defining the physics
# The dynamics of the system are governed by the combined Hamiltonian:
# H(t) = H_d + sum(u1(t)*Hc1 + u2(t)*Hc2 + ....)
# That is the time-dependent Hamiltonian has a constant part (called here the drift) and time vary parts, which are the control Hamiltonians scaled by some functions u_j(t) known as control amplitudes
# In this example we describe an Ising like Hamiltonian, encompassing random coefficients in the drift part and controlling the interaction of the qubits:
#
# $ \hat{H} = \sum_{i=1}^2 \alpha_i \sigma_x^i + \beta_i \sigma_z^i + u(t) \cdot \sigma_z \otimes \sigma_z $
#
# Initial $\newcommand{\ket}[1]{\left|{#1}\right\rangle} \ket{\psi_0} = \text{U_0}$ and target state $\ket{\psi_t} = \text{U_targ}$ are chosen to be:
#
# $ \ket{\psi_0} = \begin{pmatrix} 1 \\ 0 \\ 0 \\ 0 \end{pmatrix}$
#
# $ \ket{\psi_t} = \begin{pmatrix} 0 \\ 0 \\ 0 \\ 1 \end{pmatrix}$
# +
random.seed(20)
alpha = [random.random(),random.random()]
beta = [random.random(),random.random()]
Sx = sigmax()
Sz = sigmaz()
H_d = (alpha[0]*tensor(Sx,identity(2)) +
alpha[1]*tensor(identity(2),Sx) +
beta[0]*tensor(Sz,identity(2)) +
beta[1]*tensor(identity(2),Sz))
H_c = [tensor(Sz,Sz)]
# Number of ctrls
n_ctrls = len(H_c)
q1_0 = q2_0 = Qobj([[1], [0]])
q1_targ = q2_targ = Qobj([[0], [1]])
psi_0 = tensor(q1_0, q2_0)
psi_targ = tensor(q1_targ, q2_targ)
# -
# ### Defining the time evolution parameters
# To solve the evolution the control amplitudes are considered constant within piecewise timeslots, hence the evolution during the timeslot can be calculated using U(t_k) = expm(-i*H(t_k)*dt). Combining these for all the timeslots gives the approximation to the evolution from an initial state $\psi_0$ at t=0 to U(T) at the t=evo_time.
# The number of timeslots and evo_time have to be chosen such that the timeslot durations (dt) are small compared with the dynamics of the system.
# Number of time slots
n_ts = 100
# Time allowed for the evolution
evo_time = 18
# ### Set the conditions which will cause the pulse optimisation to terminate
# At each iteration the fidelity of the evolution is tested by comparaing the calculated evolution U(T) with the target U_targ. For unitary systems such as this one this is typically:
# f = normalise(overlap(U(T), U_targ)). The maximum fidelity (for a unitary system) calculated this way would be 1, and hence the error is calculated as fid_err = 1 - fidelity. As such the optimisation is considered completed when the fid_err falls below such a target value.
#
# In some cases the optimisation either gets stuck in some local minima, or the fid_err_targ is just not achievable, therefore some limits are set to the time/effort allowed to find a solution.
#
# The algorithm uses the CRAB algorithm to determine optimized coefficients that lead to a minimal fidelity error. The underlying optimization procedure is set to be the Nelder-Mead downhill simplex. Therefore, when all vertices shrink together, the algorithm will terminate.
# Fidelity error target
fid_err_targ = 1e-3
# Maximum iterations for the optisation algorithm
max_iter = 600
# Maximum (elapsed) time allowed in seconds
max_wall_time = 120
# ### Set the initial pulse type
# The control amplitudes must be set to some initial values. Typically these are just random values for each control in each timeslot. These do however result in erratic optimised pulses. For this example, a solution will be found for any initial pulse, and so it can be interesting to look at the other initial pulse alternatives.
# pulse type alternatives: RND|ZERO|LIN|SINE|SQUARE|SAW|TRIANGLE|
p_type = 'RND'
# ### Give an extension for output files
#Set to None to suppress output files
f_ext = "{}_n_ts{}_ptype{}.txt".format(example_name, n_ts, p_type)
# ### Run the optimisation
# In this step, the actual optimization is performed. At each iteration the Nelder-Mead algorithm calculates a new set of coefficients that improves the currently worst set among all set of coefficients. For details see [1,2] and a textbook about static search methods. The algorithm continues until one of the termination conditions defined above has been reached. If undesired results are achieved, rerun the algorithm and/or try to change the number of coefficients to be optimized for, as this is a very crucial parameter.
result = cpo.opt_pulse_crab_unitary(H_d, H_c, psi_0, psi_targ, n_ts, evo_time,
fid_err_targ=fid_err_targ,
max_iter=max_iter, max_wall_time=max_wall_time,
init_coeff_scaling=5.0, num_coeffs=5,
method_params={'xtol':1e-3},
guess_pulse_type=p_type, guess_pulse_action='modulate',
out_file_ext=f_ext,
log_level=log_level, gen_stats=True)
# ### Report the results
# Firstly the performace statistics are reported, which gives a breakdown of the processing times. In this example it can be seen that the majority of time is spent calculating the propagators, i.e. exponentiating the combined Hamiltonian.
#
# The optimised U(T) is reported as the 'final evolution', which is essentially the string representation of the Qobj that holds the full time evolution at the point when the optimisation is terminated.
#
# The key information is in the summary (given last). Here the final fidelity is reported and the reason for termination of the algorithm.
result.stats.report()
print("Final evolution\n{}\n".format(result.evo_full_final))
print("********* Summary *****************")
print("Final fidelity error {}".format(result.fid_err))
print("Final gradient normal {}".format(result.grad_norm_final))
print("Terminated due to {}".format(result.termination_reason))
print("Number of iterations {}".format(result.num_iter))
print("Completed in {} HH:MM:SS.US".format(
datetime.timedelta(seconds=result.wall_time)))
# ### Plot the initial and final amplitudes
# Here the (random) starting pulse is plotted along with the pulse (control amplitudes) that was found to produce the target gate evolution to within the specified error.
# +
fig1 = plt.figure()
ax1 = fig1.add_subplot(2, 1, 1)
ax1.set_title("Initial Control amps")
ax1.set_ylabel("Control amplitude")
ax1.step(result.time,
np.hstack((result.initial_amps[:, 0], result.initial_amps[-1, 0])),
where='post')
ax2 = fig1.add_subplot(2, 1, 2)
ax2.set_title("Optimised Control Amplitudes")
ax2.set_xlabel("Time")
ax2.set_ylabel("Control amplitude")
ax2.step(result.time,
np.hstack((result.final_amps[:, 0], result.final_amps[-1, 0])),
where='post')
plt.tight_layout()
plt.show()
# -
# ### Versions
# +
from qutip.ipynbtools import version_table
version_table()
# -
# ### References
# [1] <NAME>., <NAME>. & <NAME>.: Optimal Control Technique for Many-Body Quantum Dynamics. Phys. Rev. Lett. 106, 1–4 (2011).
#
# [2] <NAME>., <NAME>. & <NAME>.: Chopped random-basis quantum optimization. Phys. Rev. A - At. Mol. Opt. Phys. 84, (2011).
tlist = np.linspace(0,evo_time,n_ts)
H = [[H_d, np.ones(n_ts)], [H_c[0],result.final_amps[:,0]]]
sim = mesolve(H,psi_0,tlist,e_ops=[psi_targ*psi_targ.dag()])
plt.plot(tlist,sim.expect[0])
plt.ylim([0,1])
| control-pulseoptim-CRAB-2qubitInerac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Sample 4.1 random drawing points from a uniform distribution
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import time
matplotlib.rc('xtick', labelsize=12)
matplotlib.rc('ytick', labelsize=12)
#generate random number between 1 and 3 following an uniform density
a = 1.
b = 3.
np.random.seed(np.int(time.time()))
x1 = np.random.uniform(a,b,size=1000)
x2 = np.random.uniform(a,b,size=100000)
x3 = np.random.uniform(a,b,size=10000000)
#analyze the random samples with a histogram
xgrid = np.arange(1,3,0.1)
xcenter = (xgrid[1:]+xgrid[:-1])/2.
hx1,xedge = np.histogram(x1,xgrid)
hx2,xedge = np.histogram(x2,xgrid)
hx3,xedge = np.histogram(x3,xgrid)
#draw the histogram
fig = plt.figure(figsize=[10,5])
ax = fig.add_subplot(111)
ax.plot(xcenter,hx1/np.sum(hx1),'ko-')
ax.plot(xcenter,hx2/np.sum(hx2),'ro-')
ax.plot(xcenter,hx3/np.sum(hx3),'b^-',markerfacecolor='none')
#fig.show()
fig.savefig('unifrand_hist.png',bbox_inches='tight')
print('Uniform distribution')
print('a=%(a).2f, b=%(b).2f' %{'a':a,'b':b})
print('E(x)=%(m).3f\tD(x)=%(d).3f' % {'m':np.mean(x1), 'd':np.var(x1)})
print('E(x)=%(m).3f\tD(x)=%(d).3f' % {'m':np.mean(x2), 'd':np.var(x2)})
print('E(x)=%(m).3f\tD(x)=%(d).3f' % {'m':np.mean(x3), 'd':np.var(x3)})
# +
'''
demonstrate how to draw a histogram aligning with different grids
'''
fig = plt.figure(figsize=[6,6])
x = np.array([0.2,0.3,1.2,1.6,1.7,1.8,2.4,2.5,2.9])
xgrid = np.arange(-1,5,1)
xcenter = (xgrid[1:]+xgrid[:-1])/2.
h,xedge = np.histogram(x,bins=xgrid)
ax = fig.add_subplot(111)
ax.step(xedge[1:],h,'k',where='pre')
ax.step(xedge[:-1],h,'r',where='post')
ax.step(xcenter,h,'b',where='mid')
#ax.step(xcenter,h,'r',where='mid')
ax.plot(xcenter,h,'k+-')
# fig.show()
# + jupyter={"outputs_hidden": true}
| sample4.1_uniform_sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.linalg
import matplotlib.pyplot as plt
# # Item II
#
# Let $H_n$ be the $n\times n$ Hilbert matrix whose $ij$-th entry is defined as $1/(i+j-1)$, also, let $\mathbf{1}_n$ be the vector of ones of dimension $n$. *Discuss the following questions.*
# 1. Find, as accurate as possible, the approximate solution $\hat{\mathbf{x}}$ of the linear system $A\mathbf{x}=\mathbf{b}$, where $A=H_n$ and $\mathbf{b}= H_n\mathbf{1}_n$ for $n=3\dots 20$. Notice that we know a priori that the exact solution is just $\mathbf{x} = \mathbf{1}_n$, but (un)fortunately the computer can only give you $\tilde{\mathbf{x}}$.
# 2. What is the relation between $\mathbf{x}$ and $\tilde{\mathbf{x}}$?
# 3. What can we do now?
#
# ---
def hilbert(n):
v = np.arange(1,n+1,dtype='float')
iis = v.reshape((1,n))
jjs = v.reshape((n,1))
return (iis+jjs-1)**-1
# +
NS = np.arange(3,20+1)
errors = []
for n in NS:
H = hilbert(n)
real_x = np.ones(n)
b = np.dot(H,real_x)
x = np.linalg.solve(H,b)# We use np.linalg.solve
err = np.mean(np.abs((x-real_x)/real_x))
errors.append(err)
errors = np.array(errors)
# -
# Plot errors
plt.plot(NS,errors)
plt.grid()
plt.title("Mean relative error between $\\tilde{x}$ and $x$ vs. $n$")
plt.plot()
pass
# We see that as $n$ grows, the relative error between $\tilde{\mathbf{x}}$ and $\mathbf{x}$ grows fast.
#
# This is because $\text{cond}(H_n) = O\left( \left(1+\sqrt{2}\right)^{4n}/\sqrt{n}\right)$, so, given small perturbations on the operations required to solve the problem, the error on the solution grows considerably with $n$.
# ---
#
# To solve the problem we can try to minimize instead of finding the exact solution
# +
lstsq_errors = []
for n in NS:
H = hilbert(n)
real_x = np.ones(n)
b = np.dot(H,real_x)
x,_,_,_ = np.linalg.lstsq(H,b,rcond=None)
err = np.mean(np.abs((x-real_x)/real_x))
lstsq_errors.append(err)
lstsq_errors = np.array(lstsq_errors)
# -
# Plot errors
plt.plot(NS,lstsq_errors,c="#ff0000")
plt.grid()
plt.title("Mean relative error between $\\tilde{x}$ and $x$ vs. $n$ with lstsq")
plt.plot()
pass
# We can see that approaching the problem like a minimization gives far better results, because it can perform more iterations to correct numerical errors.
| t1_questions/item_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # sxs_catalog_download_example
# This notebook demonstrates how to use the `sxs` python library to download data from the SXS Catalog of waveforms hosted on Zenodo (https://github.com/moble/sxs). The catalog is available at https://black-holes.org/waveforms and is described in https://arxiv.org/abs/1904.04831.
#
# You can install `sxs` via `pip install sxs`. You'll need version `2019.4.15.16.32.10` or later.
#
# Note: You should use a recent version of python 3, such as 3.6.5, to run this notebook. I installed python 3.6.5 using anaconda (https://www.anaconda.com).
#
# Note: I found I had to also install tqdm, which you can install similarly. This should be installed when you install the `sxs` library with `pip`, but in case you see errors about being unable to import `tqdm`, `pip install tqdm` solves this.
# ## How to download data
# This section demonstrates how to download simulation data from Zenodo. You can download data from one simulation or from multiple simulations at once.
# +
# For downloading data
import sxs
from sxs import zenodo as zen
# For interacting with the data
import h5py
import numpy as np
from matplotlib import pyplot as plt
import json
# -
# This line attempts to download select files from a specific simulation.
#
# `dry_run = True` means that the function does everything except actually download the data; set this to false to download the data into the same directory as this notebook.
#
# You can download dat for multiple simulations by changing `sxs_ids` to include more simulations, or to include an expression that matches multiple simulations. For instance, `sxs_ids = ['SXS:BBH:']` would download all binary-black-hole waveforms from the catalog.
#
# Set `highest_lev_only = True` to download only the highest resoution of each file that is available at multiple resolutions, instead of downloading files from all resolutions.
zen.download.matching("common-metadata.txt", "metadata.txt", \
"rhOverM_Asymptotic_GeometricUnits_CoM.h5", \
"Horizons.h5", \
sxs_ids = ['SXS:BBH:0444'], \
dry_run = True, \
highest_lev_only = False)
# Print the help text for a function that downloads data from the catalog, for more details on how the function works.
# ?zen.download.matching
# ## Examples of using data from the catalog
# Download the highest resolution Horizons and waveform files for a specific simulation. This will download the data in the same notebook as this notebook, overwriting other files of the same name if they exist.
zen.download.matching("common-metadata.txt", "metadata.txt", "metadata.json", \
"rhOverM_Asymptotic_GeometricUnits_CoM.h5", \
"Horizons.h5", \
sxs_ids = ['SXS:BBH:0444'], \
dry_run = False, \
highest_lev_only = True)
horizons = h5py.File("SXS_BBH_0444/Horizons.h5", 'r')
rhOverM = h5py.File("SXS_BBH_0444/rhOverM_Asymptotic_GeometricUnits_CoM.h5", 'r')
# Keys of the waveform file specify the extrapolation order or a text file giving version history.
print(list(rhOverM.keys()))
print(np.array(rhOverM['VersionHist.ver']))
# Each extrapolation order has some metadata given as attributes.
print(list(rhOverM['Extrapolated_N2.dir'].attrs.keys()))
for key in rhOverM['Extrapolated_N2.dir'].attrs.keys():
print(key + " = " + str(rhOverM['Extrapolated_N2.dir'].attrs[key]))
# Here's an example that plots waveform data from the downloaded H5 files.
timeReIm = np.array(rhOverM['Extrapolated_N2.dir']['Y_l2_m2.dat'])
plt.clf()
plt.plot(timeReIm[:,0], timeReIm[:,1])
plt.xlabel("Time (code units)")
plt.ylabel("Strain")
plt.show()
# Here's a similar example plotting the irreducible mass vs. time.
list(horizons.keys())
list(horizons['AhA.dir'].keys())
plt.clf()
plt.plot(horizons['AhA.dir']['ArealMass.dat'][:,0], horizons['AhA.dir']['ArealMass.dat'][:,1], label='AhA')
plt.plot(horizons['AhB.dir']['ArealMass.dat'][:,0], horizons['AhB.dir']['ArealMass.dat'][:,1], label='AhB')
plt.plot(horizons['AhC.dir']['ArealMass.dat'][:,0], horizons['AhC.dir']['ArealMass.dat'][:,1], label='AhC')
plt.xlabel("Time (code units)")
plt.ylabel("Irreducible mass (code units)")
plt.show()
# ## Example of using metadata in the catalog
with open("SXS_BBH_0444/metadata.json") as file:
metadata_json = json.load(file)
# Here is code to parse the metadata json file `metadata.json`.
list(metadata_json.keys())
metadata_json['remnant_mass']
# And here is an example getting the same metadata from `metadata.txt`.
with open("SXS_BBH_0444/metadata.json") as file:
metadata_txt_raw = file.readlines()
# Simple code to kind of parse the text file. Room for improvement.
metadata_txt = {}
for line in metadata_txt_raw:
if ':' in line:
key = line.split('":')[0].strip().split('"')[1]
value = line.split('":')[-1].split(',')[0].strip()
metadata_txt[key] = value
float(metadata_txt['remnant_mass'])
| sxs_catalog_download_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jjcheung/ctcuc22/blob/main/JJ_Cheung_2022_Python_Programming_Practical_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="jp3BkDml8xiR"
# # **2022 Python Programming Practical 4**
#
# + [markdown] id="-U8w0oQW9P_b"
# **Q1. (Summing series 1)**
# Write a recursive function sum_series1(i) to compute the following series:
# m(i) = 1 + ½ + ⅓ + … + 1/i
#
# + id="D0uTR7B-8qbF" colab={"base_uri": "https://localhost:8080/"} outputId="af071a51-630a-4036-b8b6-8f1f138c616f"
def sum_series1(i):
if i == 1: #terminating case
return 1
else: #add 1/i to the total, while calling recursive function for next smaller i.
return 1/i + sum_series1(i-1)
# main
print(sum_series1(1))
print(sum_series1(5))
# + [markdown] id="7Ui5CuS_9Za1"
# **Q2. (Summing series 2)**
# Write a recursive function sum_series2(i) to compute the following series:
# m(i) = ⅓ + ⅖ + 3/7 + 4/9 + 5/11 + 6/13 + … + i/(2i+1)
#
# + id="CfkRAAWV9cXS" colab={"base_uri": "https://localhost:8080/"} outputId="a3d9148a-6a92-45bb-9aa6-34f45287e0fe"
def sum_series2(i):
if i == 1: #terminating case
return 1 / 3
else: #add i/(2i+1) to total, and calling recursive function for i-1
return (i/(2*i + 1)) + sum_series2(i-1)
# main
print(sum_series2(1))
print(sum_series2(5))
# + [markdown] id="aVhHiT8w9c7d"
# **Q3. (Computing GCD)**
# The greatest common divisor (GCD) of two positive integers m and n, gcd(m, n) can be defined recursively as follows:
#
# If m % n is 0, gcd(m, n) is n.
# Otherwise, gcd(m, n) is gcd(n, m % n).
#
# Write a recursive function gcd(m, n) to find the GCD. Write a test program that computes gcd(24, 16) and gcd(255, 25).
# + id="PwRmweEa9fny" outputId="a0c71cf1-900b-4e33-83b9-d711a4bab80b" colab={"base_uri": "https://localhost:8080/"}
def gcd(m, n):
if m % n == 0: #terminating case
return n
else:
return gcd(n, m % n)
# main
print(gcd(24, 16))
print(gcd(255, 25))
# + [markdown] id="wWe4kRS8ZmwP"
# **Q4. (Reverse the digits in an integer)**
# Write a recursive function reverse_int(n) that reverses the digits of an integer n:
#
# For example, reverse_int(12345) displays 54321.
# + id="aOozTwoR9j9B" outputId="9d9b106f-9996-4305-979b-dd53d4660a4e" colab={"base_uri": "https://localhost:8080/"}
def reverse_int(n):
if n // 10 == 0: #terminating case
return n
else:
digit = n
count = 0
while digit > 10: #while loop to get 1st digit in n
digit = digit // 10
count += 1 #determine power
return digit + 10 * reverse_int(n % (10**count)) #digit is now last digit in number, by calling recursive function for n without the first digit, multiply returned value by 10, and add to digit
# main
print(reverse_int(8)) # test terminating case
print(reverse_int(12345)) # test recursive case
# + colab={"base_uri": "https://localhost:8080/"} id="EaXW2pFOF1zn" outputId="448c04d7-d8b0-4c57-d335-76d08116ac11"
def reverse_int(n):
if n // 10 == 0: #terminating case
return n
else:
digit = int(str(n)[0]) #getting the first digit
rmdr = int(str(n)[1:]) #getting the remaining digits
return digit + 10 * reverse_int(rmdr) #digit is now last digit in number, by calling recursive function for the remaining digits, multiply returned value by 10, and add to digit
# main
print(reverse_int(8)) # test terminating case
print(reverse_int(12345)) # test recursive case
# + colab={"base_uri": "https://localhost:8080/"} id="SOYZcH3IbB-0" outputId="7519d056-8bff-46bc-98e0-fcf7b7e1b4cf"
def reverse_int2(n, rem = 0):
if n == 0:
return rem // 10
else:
return reverse_int2(n //10, (rem+(n%10))*10)
reverse_int2(12345)
# + [markdown] id="lE0ScX9e9kcg"
# **Q5. (Occurrences of a specified character in a string)**
# Write a recursive function count_letter(str, ch) that finds the number of occurrences of a specified letter ch in a string str:
#
# For example, count_letter("Welcome", 'e') returns 2.
# + id="sE_bxwDs9mf6" colab={"base_uri": "https://localhost:8080/"} outputId="8d44df31-b5fa-4dd8-d09c-1c48525bc87f"
def count_letter(string, ch):
if string == '': #terminating case for empty string
return 0
elif string[0] == ch: #checking if first character in string is ch
return 1 + count_letter(string[1:], ch) #add 1 to count, and calls recursive function for the string without the first character
else:
return count_letter(string[1:], ch) #if the character is not ch, calls the recursive function for the string without the first character
#main
count_letter("Welcome", 'e')
# + colab={"base_uri": "https://localhost:8080/"} id="TiD81J3fiaWH" outputId="2f66fddd-ca71-491c-9b0b-77972b7d8ac6"
def count_letter(string, ch):
if len(string) == 1: #terminating case
if string[0] == ch:
return 1
else:
return 0
else:
return count_letter(string[0], ch) + count_letter(string[1:], ch)
#pop first letter to be called recursively to return 1 or 0. rest of letters called recursively to repeat till all letters popped
#main
count_letter("Welcome", 'e')
# + [markdown] id="KSviQY0T9m7Q"
# **Q6. (Summing the digits in an integer)**
# Write a recursive function sum_digits(n) that computes the sum of the digits in an integer n:
#
# For example, sum_digits(234) returns 9.
# + id="YbH9Pbul9rK-" colab={"base_uri": "https://localhost:8080/"} outputId="057f8eae-2af9-4304-99da-11c7b4484a16"
def sum_digits(n):
if n // 10 == 0: #terminating case
return n
else:
return n % 10 + sum_digits(n//10) #adds last digit to total, calls recursive function with the remaining digits
#main
sum_digits(1234)
# + [markdown] id="V9Z95nlg97e4"
# **Q7. (Finding the largest number in an array)**
# Write a recursive function find_largest(alist) that returns the largest integer in an array alist.
#
# For example, given alist = [5, 1, 8, 7, 2], sum_digits(alist) returns 8.
#
#
# + id="1AZvvAW499PK" colab={"base_uri": "https://localhost:8080/"} outputId="14464fd4-2adc-4bfb-a0b2-e9472a155eb7"
def find_largest(alist):
if len(alist) == 1: #terminating case. whatever is left in the list should be the largest value
print(alist[0])
else:
if alist[0]>alist[1]: #checking for first two values in list. Pop the smaller value
alist.pop(1)
else: #if same value, pop one of the values.
alist.pop(0)
find_largest(alist) #calls recursive function with the remaining list
#main
find_largest([5, 1, 10, 7, 10, 2])
# + [markdown] id="KNHIvXFleFFH"
# **Q8. (Finding the number of uppercase letters in a string)**
# Write a recursive function find_num_uppercase(str) to return the number of uppercase letters in a string str.
#
# For example, find_num_uppercase('Good MorninG!') returns 3.
#
#
# + id="-CZ0M7j6eHQe" colab={"base_uri": "https://localhost:8080/"} outputId="f32877a1-b2e9-4853-c360-08bf8be8e65b"
def find_num_uppercase(string):
if string == '': #terminating case
return 0
elif string[0].isupper(): #check if first character in string is uppercase
return 1 + find_num_uppercase(string[1:]) #adds 1 to total, calls recursive function for the remaining characters in string
else:
return find_num_uppercase(string[1:]) #if first character is lowercase, calls recursive functuon for remaining characters in string
#main
find_num_uppercase("Good MorninG!")
# + colab={"base_uri": "https://localhost:8080/"} id="0zE8Xw7ph1jP" outputId="c4da6e0c-adeb-4a0a-8d0f-c81a7f8d11be"
def find_num_uppercase(string):
if len(string) == 1: #terminating case
if string.isupper(): #check if character in string is uppercase
return 1
else:
return 0
else:
return find_num_uppercase(string[0]) + find_num_uppercase(string[1:])
#pop first char in string to be called recursively to return 1 or 0. other char in string called recursively till all chars popped.
#main
find_num_uppercase("Good MorninG!")
# + [markdown] id="0erZpVERHKzF"
# **Fibonnaci**
# + colab={"base_uri": "https://localhost:8080/"} id="s14LVvjJHKR0" outputId="579d1ec2-d134-4832-d636-3c41d632d2ff"
def fibr(n):
if n <= 2:
return 1
else:
return fibr(n-1) + fibr(n-2)
fibr(5)
# + [markdown] id="GDNiYgGgHcyk"
# **Binary search**
# + id="eRr2HzWbPCbG" colab={"base_uri": "https://localhost:8080/"} outputId="51f52cc3-99a8-44bb-968b-8d5f2fdd7298"
def binary_search(target, low, high):
mid = (high+low)//2
if A[mid] == target:
return mid
elif high < low:
return False
elif A[mid] > target:
return binary_search(target, low, mid-1)
else:
return binary_search(target, mid+1, high)
A = [1, 2, 3, 4, 5, 6, 7, 8] #sorted
print (binary_search(3, 0, len(A)-1))
# + [markdown] id="oCq1W_BgK8bI"
# **Factorial**
# + colab={"base_uri": "https://localhost:8080/"} id="HtXyV_B1IsLs" outputId="e39ee9da-f6e8-4578-d309-fe84fddd43d1"
def factr(n):
if n == 1 or n == 0:
return 1
else:
return n * factr(n-1)
print(factr(5))
print(factr(0))
# + id="njLrAtquLPpk"
| JJ_Cheung_2022_Python_Programming_Practical_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://pymt.readthedocs.io"><img style="float: left" src="../media/powered-by-logo-header.png"></a>
# # Exploring Surface Processes using CSDMS Tools: How to Build Coupled Models
#
# ## Day 2: The Basic Model Interface (BMI)
# # Coming Soon... please check back here on Day 2 of the clinic.
| notebooks/bmi/coming_soon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 02 - OLAP Cubes - Slicing and Dicing
# All the databases table in this demo are based on public database samples and transformations
# - `Sakila` is a sample database created by `MySql` [Link](https://dev.mysql.com/doc/sakila/en/sakila-structure.html)
# - The postgresql version of it is called `Pagila` [Link](https://github.com/devrimgunduz/pagila)
# - The facts and dimension tables design is based on O'Reilly's public dimensional modelling tutorial schema [Link](http://archive.oreilly.com/oreillyschool/courses/dba3/index.html)
#
# Start by creating and connecting to the database by running the cells below.
# !PGPASSWORD=student createdb -h 127.0.0.1 -U student pagila_star
# !PGPASSWORD=student psql -q -h 127.0.0.1 -U student -d pagila_star -f Data/pagila-star.sql
# ### Connect to the local database where Pagila is loaded
# +
import sql
# %load_ext sql
DB_ENDPOINT = "127.0.0.1"
DB = 'pagila_star'
DB_USER = 'student'
DB_PASSWORD = '<PASSWORD>'
DB_PORT = '5432'
# postgresql://username:password@host:port/database
conn_string = "postgresql://{}:{}@{}:{}/{}" \
.format(DB_USER, DB_PASSWORD, DB_ENDPOINT, DB_PORT, DB)
print(conn_string)
# %sql $conn_string
# -
# ### Star Schema
# <img src="pagila-star.png" width="50%"/>
# # Start with a simple cube
# TODO: Write a query that calculates the revenue (sales_amount) by day, rating, and city. Remember to join with the appropriate dimension tables to replace the keys with the dimension labels. Sort by revenue in descending order and limit to the first 20 rows. The first few rows of your output should match the table below.
# +
# %%time
# %%sql
SELECT d.day, c.city, m.rating ,sum(f.sales_amount) as revenue
FROM factsales f
join dimdate d on d.date_key =f.date_key
join dimcustomer c on c.customer_key = f.customer_key
join dimmovie m on m.movie_key=f.movie_key
group by (d.day, c.city, m.rating)
order by revenue desc
limit 10
# -
# <div class="p-Widget jp-RenderedHTMLCommon jp-RenderedHTML jp-mod-trusted jp-OutputArea-output jp-OutputArea-executeResult" data-mime-type="text/html"><table>
# <tbody><tr>
# <th>day</th>
# <th>rating</th>
# <th>city</th>
# <th>revenue</th>
# </tr>
# <tr>
# <td>30</td>
# <td>G</td>
# <td>San Bernardino</td>
# <td>24.97</td>
# </tr>
# <tr>
# <td>30</td>
# <td>NC-17</td>
# <td>Apeldoorn</td>
# <td>23.95</td>
# </tr>
# <tr>
# <td>21</td>
# <td>NC-17</td>
# <td>Belm</td>
# <td>22.97</td>
# </tr>
# <tr>
# <td>30</td>
# <td>PG-13</td>
# <td>Zanzibar</td>
# <td>21.97</td>
# </tr>
# <tr>
# <td>28</td>
# <td>R</td>
# <td>Mwanza</td>
# <td>21.97</td>
# </tr>
# </tbody></table></div>
# ## Slicing
#
# Slicing is the reduction of the dimensionality of a cube by 1 e.g. 3 dimensions to 2, fixing one of the dimensions to a single value. In the example above, we have a 3-dimensional cube on day, rating, and country.
#
# TODO: Write a query that reduces the dimensionality of the above example by limiting the results to only include movies with a `rating` of "PG-13". Again, sort by revenue in descending order and limit to the first 20 rows. The first few rows of your output should match the table below.
# +
# %%time
# %%sql
SELECT d.day, c.city, m.rating ,sum(f.sales_amount) as revenue
FROM factsales f
join dimdate d on d.date_key =f.date_key
join dimcustomer c on c.customer_key = f.customer_key
join dimmovie m on m.movie_key=f.movie_key
where m.rating='PG-13'
group by (d.day, c.city, m.rating)
order by revenue desc
limit 10
# -
# <div class="p-Widget jp-RenderedHTMLCommon jp-RenderedHTML jp-mod-trusted jp-OutputArea-output jp-OutputArea-executeResult" data-mime-type="text/html"><table>
# <tbody><tr>
# <th>day</th>
# <th>rating</th>
# <th>city</th>
# <th>revenue</th>
# </tr>
# <tr>
# <td>30</td>
# <td>PG-13</td>
# <td>Zanzibar</td>
# <td>21.97</td>
# </tr>
# <tr>
# <td>28</td>
# <td>PG-13</td>
# <td>Dhaka</td>
# <td>19.97</td>
# </tr>
# <tr>
# <td>29</td>
# <td>PG-13</td>
# <td>Shimoga</td>
# <td>18.97</td>
# </tr>
# <tr>
# <td>30</td>
# <td>PG-13</td>
# <td>Osmaniye</td>
# <td>18.97</td>
# </tr>
# <tr>
# <td>21</td>
# <td>PG-13</td>
# <td>Asuncin</td>
# <td>18.95</td>
# </tr>
# </tbody></table></div>
# ## Dicing
# Dicing is creating a subcube with the same dimensionality but fewer values for two or more dimensions.
#
# TODO: Write a query to create a subcube of the initial cube that includes moves with:
# * ratings of PG or PG-13
# * in the city of Bellevue or Lancaster
# * day equal to 1, 15, or 30
#
# The first few rows of your output should match the table below.
# +
# %%time
# %%sql
SELECT d.day, c.city, m.rating ,sum(f.sales_amount) as revenue
FROM factsales f
join dimdate d on d.date_key =f.date_key
join dimcustomer c on c.customer_key = f.customer_key
join dimmovie m on m.movie_key=f.movie_key
where m.rating in ('PG-13','PG') and c.city in ('Bellevue','Lancaster') and d.day in (1,15,30)
group by (d.day, c.city, m.rating)
order by revenue desc
limit 10
# -
# <div class="p-Widget jp-RenderedHTMLCommon jp-RenderedHTML jp-mod-trusted jp-OutputArea-output jp-OutputArea-executeResult" data-mime-type="text/html"><table>
# <tbody><tr>
# <th>day</th>
# <th>rating</th>
# <th>city</th>
# <th>revenue</th>
# </tr>
# <tr>
# <td>30</td>
# <td>PG</td>
# <td>Lancaster</td>
# <td>12.98</td>
# </tr>
# <tr>
# <td>1</td>
# <td>PG-13</td>
# <td>Lancaster</td>
# <td>5.99</td>
# </tr>
# <tr>
# <td>30</td>
# <td>PG-13</td>
# <td>Bellevue</td>
# <td>3.99</td>
# </tr>
# <tr>
# <td>30</td>
# <td>PG-13</td>
# <td>Lancaster</td>
# <td>2.99</td>
# </tr>
# <tr>
# <td>15</td>
# <td>PG-13</td>
# <td>Bellevue</td>
# <td>1.98</td>
# </tr>
# </tbody></table></div>
| Notebooks_Scripts/L1 E2 - 1 - Slicing and Dicing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:quamd]
# language: python
# name: conda-env-quamd-py
# ---
# +
import os
from skimage import io
import pylab as plt
import wandb
import numpy as np
import time
from ml_utils.model.faster_rcnn import load_model_for_training
from ml_utils.train.dataloader_bbox import get_data_loaders
from ml_utils.train.train_bbox import train
from ml_utils.utils.visualize import draw_bbox
# -
# ### Specify parameters
# +
input_dir = '../example_data/img'
bbox_fn = '../example_data/bboxes.csv'
model_dir = '../outputs/model'
project_name = 'test_project'
batch_size = 2
val_fraction = 0.2
num_workers = 2
config = dict(num_epochs=20,
lr=0.01,
momentum=0.9,
weight_decay=0.0005,
step_size=3,
gamma=0.1,
detection_thr=0.1,
overlap_thr=0.1,
dist_thr=10)
log_progress = True
# -
# ### Show example data
# +
tr_dl, val_dl = get_data_loaders(bbox_fn,
input_dir=input_dir,
val_fraction=val_fraction,
batch_size=batch_size,
num_workers=num_workers)
n = 0
for images, targets, image_ids in tr_dl:
for i in range(len(images)):
boxes = targets[i]['boxes'].cpu().numpy().astype(np.int32)
sample = images[i].permute(1, 2, 0).cpu().numpy()
for box in boxes:
sample = draw_bbox(sample, [box[1], box[0], box[3], box[2]], color=(1, 0, 0))
io.imshow(sample)
plt.show()
n += 1
if n > 3:
break
# -
# ### Load data and model
tr_dl, val_dl = get_data_loaders(bbox_fn,
input_dir=input_dir,
val_fraction=val_fraction,
batch_size=batch_size,
num_workers=num_workers)
model = load_model_for_training()
# ### Train the model
# + tags=[]
# %%time
if log_progress is False:
os.environ['WANDB_MODE'] = 'offline'
wandb.init(project=project_name, config=config)
train(model, tr_dl, val_dl, config=config, log_progress=log_progress, model_dir=model_dir)
wandb.finish()
# -
| notebooks/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp bayes_inference
# -
# # Bayes Inference
#
# > * adding a measure of uncertainty to predictions
#
#
# * useful for detecting out of distribution (OOD) samples in your data
# * works without an OOD sample dataset
# * works with existing trained models
# * tradeoff: slower inference due to sampling over distribution
# * behind the scenes -> uses the MonteCarlo Dropout Callback (MCDropoutCallback)
# * based on the article : [Bayesian deep learning with Fastai : how not to be uncertain about your uncertainty !](https://towardsdatascience.com/bayesian-deep-learning-with-fastai-how-not-to-be-uncertain-about-your-uncertainty-6a99d1aa686e)
# * and on the [github code](https://github.com/dhuynh95/fastai_bayesian) by <NAME>
# * updated for fastai v2
#ci
#hide
# !pip install -Uqq fastai --upgrade
# !pip install -Uqq seaborn
# !pip install -Uqq pandas
#local
#hide
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
#export
from fastai.callback.preds import MCDropoutCallback
from fastai.learner import Learner
from fastcore.foundation import patch, L
from fastcore.basics import tuplify,detuplify
from fastai.torch_core import to_np
from fastai.data.transforms import get_image_files
from fastai.vision.core import PILImage
#export
from collections import Counter
import seaborn as sns
import torch
import pandas as pd
import numpy as np
# #### Bayesian Metrics
#
# This is modified from fastai_bayesian github code by <NAME>
# but modified to use Pytorch tensors instead of Numpy arrays
# +
#export
def entropy(probs):
"""Return the prediction of a T*N*C tensor with :
- T : the number of samples
- N : the batch size
- C : the number of classes
"""
mean_probs = probs.mean(dim=0)
entrop = - (torch.log(mean_probs) * mean_probs).sum(dim=1)
return entrop
def uncertainty_best_probability(probs):
"""Return the standard deviation of the most probable class"""
idx = probs.mean(dim=0).argmax(dim=1)
std = probs[:, torch.arange(len(idx)), idx].std(dim=0)
return std
def BALD(probs):
"""Information Gain, distance between the entropy of averages and average of entropy"""
entrop1 = entropy(probs)
entrop2 = - (torch.log(probs) * probs).sum(dim=2)
entrop2 = entrop2.mean(dim=0)
ig = entrop1 - entrop2
return ig
def top_k_uncertainty(s, k=5, reverse=True):
"""Return the top k indexes"""
sorted_s = sorted(list(zip(torch.arange(len(s)), s)),
key=lambda x: x[1], reverse=reverse)
output = [sorted_s[i][0] for i in range(k)]
def plot_hist_groups(pred,y,metric,bins=None,figsize=(16,16)):
TP = to_np((pred.mean(dim=0).argmax(dim=1) == y) & (y == 1))
TN = to_np((pred.mean(dim=0).argmax(dim=1) == y) & (y == 0))
FP = to_np((pred.mean(dim=0).argmax(dim=1) != y) & (y == 0))
FN = to_np((pred.mean(dim=0).argmax(dim=1) != y) & (y == 1))
result = metric(pred)
TP_result = result[TP]
TN_result = result[TN]
FP_result = result[FP]
FN_result = result[FN]
fig,ax = plt.subplots(2,2,figsize=figsize)
sns.distplot(TP_result,ax=ax[0,0],bins=bins)
ax[0,0].set_title(f"True positive")
sns.distplot(TN_result,ax=ax[0,1],bins=bins)
ax[0,1].set_title(f"True negative")
sns.distplot(FP_result,ax=ax[1,0],bins=bins)
ax[1,0].set_title(f"False positive")
sns.distplot(FN_result,ax=ax[1,1],bins=bins)
ax[1,1].set_title(f"False negative")
return output
# -
# ##### Get predictions for a test set
# This patches a method to learner to make mc dropout predictions
#export
@patch
def bayes_get_preds(self:Learner, ds_idx=1, dl=None, n_sample=10,
act=None,with_loss=False, **kwargs):
"""Get MC Dropout predictions from a learner, and eventually reduce the samples"""
cbs = [MCDropoutCallback()]
if 'cbs' in kwargs:
kw_cbs = kwargs.pop('cbs')
if 'MCDropoutCallback' not in L(kw_cbs).attrgot('name'):
cbs = kw_cbs + cbs
preds = []
with self.no_bar():
for i in range(n_sample):
pred, y = self.get_preds(ds_idx=ds_idx,dl=dl,act=act,
with_loss=with_loss, cbs=cbs, **kwargs)
# pred = n_dl x n_vocab
preds.append(pred)
preds = torch.stack(preds)
ents = entropy(preds)
mean_preds = preds.mean(dim=0)
max_preds = mean_preds.max(dim=1)
best_guess = max_preds.indices
best_prob = max_preds.values
best_cat = L(best_guess,use_list=True).map(lambda o: self.dls.vocab[o.item()])
return preds, mean_preds, ents,best_guess, best_prob, best_cat
# ##### Get predictions for an image item
#export
@patch
def bayes_predict(self:Learner,item, rm_type_tfms=None, with_input=False,
sample_size=10,reduce=True):
"gets a sample distribution of predictions and computes entropy"
dl = self.dls.test_dl([item], rm_type_tfms=rm_type_tfms, num_workers=0)
# modify get_preds to get distributed samples
collect_preds = []
collect_targs = []
collect_dec_preds = []
collect_inp = None
cbs = [MCDropoutCallback()]
with self.no_bar():
for j in range(sample_size):
inp,preds,_,dec_preds = self.get_preds(dl=dl, with_input=True,
with_decoded=True,
cbs=cbs)
i = getattr(self.dls, 'n_inp', -1)
inp = (inp,) if i==1 else tuplify(inp)
dec = self.dls.decode_batch(inp + tuplify(dec_preds))[0]
dec_inp,dec_targ = map(detuplify, [dec[:i],dec[i:]])
# res = dec_targ,dec_preds[0],preds[0]
if with_input and collect_inp is None: # collect inp first iter only
collect_inp = dec_inp
collect_targs.append(dec_targ)
collect_dec_preds.append(dec_preds[0])
collect_preds.append(preds[0])
dist_preds = torch.stack(collect_preds)
dist_dec_preds = L(collect_dec_preds).map(lambda o: o.item())
dist_targs = L(collect_targs)
res1 = (dist_targs, dist_dec_preds, dist_preds)
mean_pred = dist_preds.mean(dim=0)
ent = entropy(dist_preds.unsqueeze(1)).item()
best_guess = torch.argmax(mean_pred).item()
best_prob = mean_pred[best_guess].item()
best_cat = self.dls.vocab[best_guess]
res2 = (ent, best_prob, best_guess, best_cat)
if reduce:
if len(dist_targs.unique()) > 1:
targ = Counter(dist_targs)
else:
targ = dist_targs.unique()[0]
if len(dist_dec_preds.unique()) > 1:
dec_pred = Counter(dist_dec_preds)
else:
dec_pred = dist_dec_preds.unique()[0]
res1 = (targ, dec_pred, mean_pred)
res = res1 + res2
if with_input:
res = (collect_inp,) + res
return res
# ##### Add uncertainty threshold to prediction
#export
@patch
def bayes_predict_with_uncertainty(self:Learner, item, rm_type_tfms=None, with_input=False, threshold_entropy=0.2, sample_size=10, reduce=True):
"gets prediction results plus if prediction passes entropy threshold"
res = self.bayes_predict(item,rm_type_tfms=rm_type_tfms,
with_input=with_input, sample_size=sample_size,
reduce=reduce)
ent = res[4] if with_input else res[3]
return (ent < threshold_entropy,) + res
# ##### Add kitchen sink method to build dataframe, dataloader and predictions
#export
@patch
def bayes_build_inference_dfdlpreds(self:Learner, path, dataset, item_count=100,n_sample=10):
items = get_image_files(path).shuffle()[:item_count]
dl = self.dls.test_dl(items.map(lambda o: PILImage.create(o)), num_workers=0)
res = self.bayes_get_preds(dl=dl,n_sample=n_sample)
ents = res[2]
preds = res[0]
unc = uncertainty_best_probability(preds)
bald = BALD(preds)
df = pd.DataFrame(pd.Series(items,name='image_files'))
df['entropy'] = pd.Series(ents,name='entropy')
df['best_prob_uncertainty'] = pd.Series(unc,name='best_prob_uncertainty')
df['bald'] = pd.Series(bald,name='bald')
df['dataset'] = dataset
return (df,dl, preds)
# ### Test Functions
from fastai.test_utils import synth_dbunch, synth_learner
try:
from contextlib import nullcontext # python 3.7 only
except ImportError as e:
from contextlib import suppress as nullcontext # supported in 3.6 below
dls = synth_dbunch()
dls.vocab = [1,]
learner = synth_learner(data=dls)
learner.no_bar = nullcontext
bears_dl = dls.train
pets_dl = dls.valid
N_SAMPLE = 2
CATEGORIES = 1
BS = 160
# +
#local
from fastai.learner import load_learner
from fastai.data.transforms import get_image_files
from fastai.data.external import Config
from fastai.vision.core import PILImage
import random
# setup objects using local paths
cfg = Config()
learner = load_learner(cfg.model_path/'bears_classifier'/'export.pkl')
bear_path = cfg.data_path/'bears'
pet_path = cfg.data_path/'pets'
bear_img_files = get_image_files(bear_path)
pet_img_files = get_image_files(pet_path)
random.seed(69420) # fix images retrieved
pet_img = PILImage.create(pet_img_files.shuffle()[0])
bear_img = PILImage.create(bear_img_files.shuffle()[0])
pet_items = pet_img_files.shuffle()[:20]
bear_items = bear_img_files.shuffle()[:20]
pet_dset = pet_items.map(lambda o: PILImage.create(o))
bear_dset = bear_items.map(lambda o: PILImage.create(o))
pets_dl = learner.dls.test_dl(pet_dset,num_workers=0)
bears_dl = learner.dls.test_dl(bear_dset,num_workers=0)
# xb.shape = torch.size([20,3,224,224])
N_SAMPLE = 2
CATEGORIES = 3
BS = 20
# -
from fastcore.test import *
# ##### Bayes Prediction for Test Set
bear_res = learner.bayes_get_preds(dl=bears_dl, n_sample=N_SAMPLE)
pet_res = learner.bayes_get_preds(dl=pets_dl, n_sample=N_SAMPLE)
# preds, mean_preds, ents,best_guess, best_prob, best_cat
test_eq(len(bear_res),6)
# ci 6
# local 6
# predictions
test_eq(bear_res[0].shape, [N_SAMPLE,BS,CATEGORIES])
#ci torch.Size([2, 160, 1])
#local torch.Size([5, 20, 3])
# mean predictions
test_eq(bear_res[1].shape, [BS, CATEGORIES])
#ci torch.Size([160, 1])
#local torch.Size([20, 3])
# entropy
test_eq(bear_res[2].shape,[BS])
#ci torch.Size([160])
#local torch.Size([20])
# best guess (index of mean)
test_eq(bear_res[3].shape,[BS])
# ci torch.Size([160])
# local torch.Size([20])
# best probability (mean prediction)
test_eq(bear_res[4].shape,[BS])
#ci torch.Size([160])
#local torch.Size([20])
# best category (mean prediction)
test_eq(len(bear_res[5]),BS)
# ci 160
# local 20
| nbs/05_bayes_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rsskga/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module4-makefeatures/LS_DS_114_Make_Features_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="SnDJqBLi0FYW" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# + [markdown] id="W5GjI1z5yNG4" colab_type="text"
# # ASSIGNMENT
#
# - Replicate the lesson code.
#
# - This means that if you haven't followed along already, type out the things that we did in class. Forcing your fingers to hit each key will help you internalize the syntax of what we're doing.
# - [Lambda Learning Method for DS - By <NAME>](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit?usp=sharing)
# - Convert the `term` column from string to integer.
# - Make a column named `loan_status_is_great`. It should contain the integer 1 if `loan_status` is "Current" or "Fully Paid." Else it should contain the integer 0.
#
# - Make `last_pymnt_d_month` and `last_pymnt_d_year` columns.
# + id="AazB4eFwym2p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="7075a27f-cb70-487b-8cf1-6844196a0ed3"
# !wget https://resources.lendingclub.com/LoanStats_2018Q4.csv.zip
# + id="lSzCb_Zzg6Rx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="96136a93-ce3d-4838-8603-d0aecfa56a33"
# !unzip LoanStats_2018Q4.csv.zip
# + id="D0O-KARJg8az" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 210} outputId="09a55d75-a4fb-4c0c-e5f6-2a1b92482b2a"
# !head LoanStats_2018Q4.csv
# + id="nbUapO70g-NL" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="IshRExRehDhK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="34c58412-04f0-41dd-f343-c2ad73c27226"
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
df = pd.read_csv('LoanStats_2018Q4.csv', header=1, skipfooter=2, engine='python')
df.head()
# + id="9mVt_SIEhcy2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="717c3681-d3ff-4102-cdd7-7fa90d3d5262"
df_preprocessed = pd.read_csv('LoanStats_2018Q4.csv', header=1, skipfooter=2, engine='python')
df_preprocessed.head()
# + id="6YrFnK28hHPe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="d7bf5164-d53d-4022-b2c3-1390a872a97c"
# Convert the term column from string to integer.
def str_to_int(data):
return int(data.strip().strip(' months')) if isinstance(data, str) else data
df_preprocessed['term'] = df_preprocessed['term'].apply(str_to_int)
df_preprocessed['term'].head()
# + id="MOsAYjM7jOEk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="8b7067c6-33a1-4812-cf59-a7231943c1b7"
# Make a column named loan_status_is_great for one-hot encoding
# return 1 if (loan_status == "Current" or "Fully Paid") else 0
df_preprocessed['loan_status'].head(10)
# + id="LnsQHnHliCLt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="5d0bdb14-9311-43bc-9d2c-c1b8e61ba0e0"
def get_boolean(x, **kwargs):
return 1 if x in kwargs['true_conditions'] else 0
df_preprocessed['loan_status_is_great'] = df_preprocessed['loan_status'].apply(get_boolean, true_conditions=('Current|Fully Paid'))
df_preprocessed['loan_status_is_great'].head(10)
# + id="_UvRNrV3cM-I" colab_type="code" colab={}
import math
# + id="AW6OF1O9iH6W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="488b134e-8753-40e9-aaac-06c94bb5a9f1"
# Make last_pymnt_d_month and last_pymnt_d_year columns
# pandas supports NaN floats in dtype='Int64'
# other dtypes could not be mixed (i.e., str and int) based on my efforts
# mixed = dtype('<U21') - see https://github.com/numpy/numpy/issues/7973
# https://pandas.pydata.org/pandas-docs/version/0.24/whatsnew/v0.24.0.html#optional-integer-na-support
def get_datetime_fragment(x, scope):
# def get_datetime_fragment(x, **kwargs):
if isinstance(x, str):
dt = pd.to_datetime(x)
value = int(dt.month) if scope == 'month' else int(dt.year)
# value = int(dt.month) if kwargs['output'] == 'month' else int(dt.year)
# elif isinstance(x, float):
else:
# nan is a float, so convert to str
value = x
return value
# list comprehension results in ints, as desired
lst = df_preprocessed['last_pymnt_d'].tolist()
int_list = [get_datetime_fragment(item, 'month') for item in lst]
df_preprocessed['last_pymnt_d_month'] = pd.Series(int_list, dtype='Int64')
# apply method results in floats, unsure why casting to dtype='Int64' works in one and not the other
# df_preprocessed['last_pymnt_d_month'] = pd.Series([], dtype='Int64')
# df_preprocessed['last_pymnt_d_month'] = df_preprocessed['last_pymnt_d'].apply(get_datetime_fragment, output=('month'))
df_preprocessed['last_pymnt_d_month'].head()
# + id="Qsq4zS1CaPyI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="259b209d-6bc2-400f-e510-a86d12fade89"
lst = df_preprocessed['last_pymnt_d'].tolist()
int_list = [get_datetime_fragment(item, 'year') for item in lst]
df_preprocessed['last_pymnt_d_year'] = pd.Series(int_list, dtype='Int64')
df_preprocessed['last_pymnt_d_year'].head()
# + id="_PI_CNFU1vkG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="652f8088-a3cf-49ed-a7fa-a40395583c1a"
print(type(df_preprocessed['last_pymnt_d_month'][0]))
print(type(df_preprocessed['last_pymnt_d_year'][0]))
print(type(df_preprocessed['last_pymnt_d_month'][128337]))
print(type(df_preprocessed['last_pymnt_d_year'][128337]))
print(df_preprocessed['last_pymnt_d_month'][128337])
print(df_preprocessed['last_pymnt_d_year'][128337])
# + id="PNSsCKOJ4YFf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="e5f97fc5-750a-4be7-ee23-7c5dc8758216"
lst = df_preprocessed['last_pymnt_d'].tolist()
# arr = pd.array([1, 2, np.nan], dtype=pd.Int64Dtype())
print(type(lst))
series = pd.Series(lst)
print(type(series))
# pd.Series([1, 2, 3]).array
# s = pd.Series(['1.0', '2', -3])
# pd.to_numeric(s)
s = pd.Series(['apple', '1.0', '2', -3]) # NaN, 1.0, 2.0, -3.0
# pd.to_numeric(s, errors='ignore')
pd.to_numeric(s, errors='coerce')
print(type(np.nan))
# + [markdown] colab_type="text" id="L8k0LiHmo5EU"
# # STRETCH OPTIONS
#
# You can do more with the LendingClub or Instacart datasets.
#
# LendingClub options:
# - There's one other column in the dataframe with percent signs. Remove them and convert to floats. You'll need to handle missing values.
# - Modify the `emp_title` column to replace titles with 'Other' if the title is not in the top 20.
# - Take initiatve and work on your own ideas!
#
# Instacart options:
# - Read [Instacart Market Basket Analysis, Winner's Interview: 2nd place, Kazuki Onodera](http://blog.kaggle.com/2017/09/21/instacart-market-basket-analysis-winners-interview-2nd-place-kazuki-onodera/), especially the **Feature Engineering** section. (Can you choose one feature from his bulleted lists, and try to engineer it with pandas code?)
# - Read and replicate parts of [Simple Exploration Notebook - Instacart](https://www.kaggle.com/sudalairajkumar/simple-exploration-notebook-instacart). (It's the Python Notebook with the most upvotes for this Kaggle competition.)
# - Take initiative and work on your own ideas!
# + [markdown] colab_type="text" id="0_7PXF7lpEXg"
# You can uncomment and run the cells below to re-download and extract the Instacart data
# + id="urIePNa0yNG6" colab_type="code" colab={}
# # !wget https://s3.amazonaws.com/instacart-datasets/instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="X9zEyu-uyNG8" colab_type="code" colab={}
# # !tar --gunzip --extract --verbose --file=instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="Y3IqrhlpyNG-" colab_type="code" colab={}
# # %cd instacart_2017_05_01
| module4-makefeatures/LS_DS_114_Make_Features_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Run the LFADS algorithm on an RNN that integrates white noise.
#
# The goal of this tutorial is to learn about LFADS by running the algorithm on a simple data generator, a vanilla recurrent neural network (RNN) that was [trained to integrate a white noise input](https://github.com/google-research/computation-thru-dynamics/blob/master/notebooks/Integrator%20RNN%20Tutorial.ipynb). Running LFADS on this integrator RNN will infer two things:
# 1. the underlying hidden state of the integrator RNN
# 2. the white noise input to the integrator RNN.
#
# Doing this will exercise the more complex LFADS architecture that is shown in Figure 5 of the [LFADS paper](https://rdcu.be/6Wji). It's pretty important that you have read at least the introduction of the paper, otherwise, you won't understand *why* we are doing what we are doing.
#
# In this tutorial we do a few things:
# 1. Load the integrator RNN data and "spikify" it by treating the hidden units as nonhomogeneous Poisson processes.
# 2. Explain a bit of the LFADS architecture and highlight some of the relevant hyperparameters.
# 3. Train the LFADS system on the spikified integrator RNN hidden states.
# 4. Plot a whole bunch of training plots and LFADS outputs!
#
# If you make it through this tutorial and understand everything in it, it is *highly* likely you'll be able to run LFADS on your own data.
#
# #### Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ### Import the tutorial code.
#
# If you are going to actually run the tutorial, you have to install JAX, download the computation thru dynamics GitHub repo, and modify a path.
# +
# Numpy, JAX, Matplotlib and h5py should all be correctly installed and on the python path.
from __future__ import print_function, division, absolute_import
import datetime
import h5py
import jax.numpy as np
from jax import random
from jax.experimental import optimizers
from jax.config import config
#config.update("jax_debug_nans", True) # Useful for finding numerical errors
import matplotlib.pyplot as plt
import numpy as onp # original CPU-backed NumPy
import scipy.signal
import scipy.stats
import os
import sys
import time
# +
# You must change this to the location of the computation-thru-dynamics directory.
HOME_DIR = '/home/sussillo/'
sys.path.append(os.path.join(HOME_DIR,'computation-thru-dynamics'))
import lfads_tutorial.lfads as lfads
import lfads_tutorial.plotting as plotting
import lfads_tutorial.utils as utils
from lfads_tutorial.optimize import optimize_lfads, get_kl_warmup_fun
# -
# ### Preliminaries - notes on using JAX
#
# JAX is amazing! It's really, really AMAZING! You program in Numpy/Python and then call a grad on your code, and it'll run speedy on GPUs! It does however have a few quirks and it uses a program deployment model you have to know about. The excited reader should definitely read the [JAX tutorial](https://github.com/google/jax) if they plan on programming with it.
#
# When using JAX for auto diff, auto batching or compiling, you should always have a two-level mental model in your mind:
# 1. At the CPU level, like normal
# 2. at the device level, for example a GPU.
#
# Since JAX compiles your code to device, it is very efficient but creates this split. Thus, for example, we have two NumPY modules kicking around: 'onp' for 'original numpy', which is on the CPU, and np, which is the JAX modified version and runs 'on device'. This latter version of numpy is enabled to compute gradients and run your code quickly.
#
# So the model then is: initialize variables, seeds, etc, at the CPU level, and *dispatch* a JAX based computation to the device. This all happens naturally whenever you call JAX enabled functions.
#
# Thus one of the first things we do initialize the onp random number generator at the CPU level.
onp_rng = onp.random.RandomState(seed=0) # For CPU-based numpy randomness
# ----------
# ### Load the data
#
# You __must__ run through the integrator RNN [tutorial notebook](https://github.com/google-research/computation-thru-dynamics/blob/master/notebooks/Integrator%20RNN%20Tutorial.ipynb) on your machine. Don't worry! It's much simpler than this tutorial! :)
#
# Point to the correct __data__ file for the integrator RNN. Note that the integrator rnn tutorial notebook creates two files, both the parameters file and the data file with examples.
INTEGRATOR_RNN_DATA_FILE = \
'/tmp/vrnn/pure_int/trained_data_vrnn_pure_int_0.00002_2019-06-19_15:12:45.h5'
lfads_dir = '/tmp/lfads/' # where to save lfads data and parameters to
rnn_type = 'lfads'
task_type = 'integrator'
# +
# Make directories
data_dir = os.path.join(lfads_dir, 'data/')
output_dir = os.path.join(lfads_dir, 'output/')
figure_dir = os.path.join(lfads_dir, os.path.join(output_dir, 'figures/'))
if not os.path.exists(output_dir):
os.makedirs(output_dir)
if not os.path.exists(figure_dir):
os.makedirs(figure_dir)
# Load synthetic data
data_dict = utils.read_file(INTEGRATOR_RNN_DATA_FILE)
# -
# ### Plot examples and statistics about the integrator RNN data.
f = plotting.plot_data_pca(data_dict)
# The goal of this tutorial is to infer the hiddens (blue), and input to the integrator RNN (umm... also blue).
f = plotting.plot_data_example(data_dict['inputs'],
data_dict['hiddens'],
data_dict['outputs'],
data_dict['targets'])
# ### Spikify the synthetic data
# The output of the integrator rnn is the continuous inputs,
# hidden states and outputs of the example. LFADS is a tool
# to infer underlying factors in spiking neural data, so we
# are going to "spikify" the integrator rnn example hidden states.
#
# Data was generated w/ VRNN w/ tanh, thus $(\mbox{data}+1) / 2 \rightarrow [0,1]$.
# We put those activations between 0 and 1 here and then convert to spikes.
# +
data_dt = 1.0/25.0 # define our dt in a physiological range
# If data is normed between 0 and 1, then a 1 yields this many
# spikes per second. Pushing this downwards makes the problem harder.
max_firing_rate = 80
train_fraction = 0.9 # Train with 90% of the synthetic data
renormed_fun = lambda x : (x + 1) / 2.0
renormed_data = renormed_fun(data_dict['hiddens'])
# When dimensions are relevant, I use a variable naming scheme like
# name_dim1xdim2x... so below, here is the synthetic data with
# 3 dimensions of batch, time and unit, in that order.
data_bxtxn = utils.spikify_data(renormed_data, onp_rng, data_dt,
max_firing_rate=max_firing_rate)
nexamples, ntimesteps, data_dim = data_bxtxn.shape
train_data, eval_data = utils.split_data(data_bxtxn,
train_fraction=train_fraction)
eval_data_offset = int(train_fraction * data_bxtxn.shape[0])
# -
eval_data.shape
# #### Plot the statistics of the data.
f = plotting.plot_data_stats(data_dict, data_bxtxn, data_dt)
# Let's study this single example of a single neuron's true firing rate (red) and the spikified version in the blue stem plot.
my_example_bidx = eval_data_offset + 0
my_example_hidx = 0
scale = max_firing_rate * data_dt
my_signal = scale*renormed_data[my_example_bidx, :, my_example_hidx]
my_signal_spikified = data_bxtxn[my_example_bidx, :, my_example_hidx]
plt.plot(my_signal, 'r');
plt.stem(my_signal_spikified);
# If you were to increase ```max_firing_rate``` to infinity, the stem plot would approach the red line. This plot gives you an idea of how challenging the data set is, at least on single trials. We can think about this a little bit. If you were to simply filter the spikes, it definitely would not look like the red trace, at this low maximum firing rate. This means that if any technique were to have
nfilt = 3
my_filtered_spikes = scipy.signal.filtfilt(onp.ones(nfilt)/nfilt, 1, my_signal_spikified)
plt.plot(my_signal, 'r');
plt.plot(my_filtered_spikes);
plt.title("This looks terrible");
plt.legend(('True rate', 'Filtered spikes'));
# This would force us to think about ways in which the *population* can be filtered. The first idea is naturally PCA. Perhaps there is a low-d subspace of signal that can be found in the high-variance top PCs. Using the entire trial, it's likely this should do better.
import sklearn
ncomponents = 100
full_pca = sklearn.decomposition.PCA(ncomponents)
full_pca.fit(onp.reshape(data_bxtxn, [-1, data_dim]))
plt.stem(full_pca.explained_variance_)
plt.title('Those top 2 PCs sure look promising!');
ncomponents = 2
pca = sklearn.decomposition.PCA(ncomponents)
pca.fit(onp.reshape(data_bxtxn[0:eval_data_offset,:,:], [-1, data_dim]))
my_example_pca = pca.transform(data_bxtxn[my_example_bidx,:,:])
my_example_ipca = pca.inverse_transform(my_example_pca)
plt.plot(my_signal, 'r')
plt.plot(my_example_ipca[:,my_example_hidx])
plt.legend(('True rate', 'PCA smoothed spikes'))
plt.title('This a bit better.');
# So temporal filtering is not great, and spatial filtering helps only a bit. What to do? The idea LFADS explores is that if you knew the system that generated the data, you would be able to separate signal from noise, the signal being what a system can generate, the noise being the rest.
# ----------------
# ## LFADS - Latent Factor Analysis via Dynamical Systems
#
#
# [Link to paper readcube version of the LFADS Nature Methods 2018 paper](https://rdcu.be/6Wji)
#
#
# ### LFADS architecture with inferred inputs
#
# There are 3 variants of the LFADS architecture in the paper
# 1. autonomous LFADS model (no inferred inputs), Fig. 1a
# 2. stitched LFADS model for data recorded in different sessions, Fig. 4a
# 3. non-autonomous LFADS model (with inferred inputs), Fig. 5a
#
# In this tutorial, we deal with the non-autonomous model, which I believe is conceptually the most interesting, but also the most challenging to understand. This tutorial (and the current code), does **NOT** handle stitched data. Stitching data isn't conceptually hard, but it's a pain to code. The Tensorflow version of the code handles that if you need it.
#
# Here is the non-autonoumous LFADS model architecture: The full description of this model is given in the paper but briefly, the idea is that the data LFADS will 'denoise' or model data generated from a nonlinear, autonoumous system (we call it the data generator and the data generator in this tutorial is the integrator RNN) that receives an input through time. Based on the spiking observations, LFADS will try to pull apart the data into the dynamical system portion, and the input portion, thus the term *inferred inputs*. I.e. we are trying to infer what inputs would drive a high-d nonlinear system to generate the data you've recorded. Doing this allows the system to model the dynamics much better for systems that are input-driven. One final detail is that the model assumes that the spikes are poisson generated from an underlying continuous dynamical system. Of course, this is not true for spiking data from biological neural circuits, but the poisson approximation seems to be ok.
#
# So architecture infers a number of quantities of interest:
# 1. initial state to the generator (also called initial conditions)
# 2. inferred inputs to the generator - e.g. the LFADS component to learn the white noise in the integrator RNN example
# 3. dynamical factors - these are like PCs underlying your data
# 4. rates - a readout from the factors. The rates are really the most intuitive part, which are analogous to filtering your spiking data.
#
# 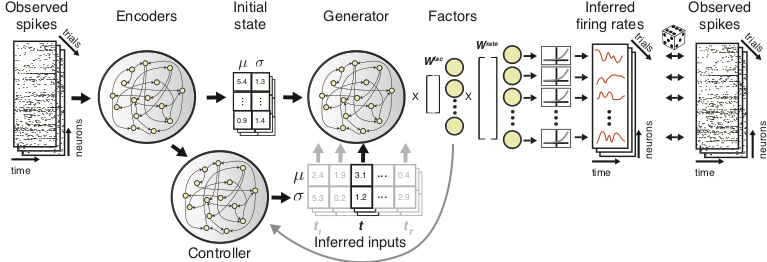
#
#
# To begin, let's focus on the *autonomous* version of the architecture, which *excludes the controller RNN*. The data is put through nonlinear, recurrent **encoders**, and this produces an **initial state distribution**, which is a per-trial mean and variance to produce random vectors to encode that trial. The initial state of the generator is a randomly drawn vector from this distribution. The **generator** marches through time and at each time point produces **factors** and **rates**, ultimately producing outputs that learn to reproduce your data at the rate level.
#
#
# From the perspective on information flow, the autonomous version of LFADS has a *bottleneck* between your data as inputted into LFADS, and the output, which also tries to learn your data. That bottleneck is the initial state of the generator, a potentially very low-bandwidth bottleneck, as a single vector has to encode a high-d time series. Such a system would be adequate for capturing systems that are (in approximation) autonomous. For example, motor cortex dynamics during center-out reaches seem extremely well approximated by autonomous dynamics at the sub-second time scale (e.g. Fig 2). However, if you were to perturb the reach by messing with the cursor the animal was using, e.g perturbing cursor location mid-reach, then the motor cortical dynamics of a corrected reach couldn't possibly be autonomous. In other words, some additional input must have come into the motor cortex and update the system with the information that the cursor had jumped unexpectedly. This is the experimental setting we setup in Fig. 5.
#
# To compensate for such a scenario, we added a **controller** and **inferred inputs** to the generator portion of LFADS. In particular, the controller runs in sync with the generator and receives the output of the generator from the last time step (the only "backward" loop in the architecture, aside from using backprop for training with gradient descent). Thus it knows what the generator output. During training, the system learns that there are patterns in the data that cannot be created by the generator autonomously, so learns to compensate by emitting information from the data, through the encoders, through the controller to the generator. We call this information an inferred input. In our experimental setup, this worked well on two examples: messing with the cursor of an animal making a reach and also for inferring oscillations in the local field potential (LFP).
#
# Please note that the inferred input system is extremely powerful as it provides a leak from your input data to the LFADS output on a per-time point basis. As such, one has to make sure that the system does not pathologically leak all the information from the data trial through LFADS to generate the data trial. LFADS, like all auto-encoders, is at risk of creating a trivial identity function, $x = f(x)$, rather than finding structure in the data. Thus, we utilize many tricks to avoid this (dropout, KL penalties, and even blocking out the information given to the controller from time step t, when decoding time step t.)
# ### Hyperparameters
# +
# LFADS Hyper parameters
data_dim = train_data.shape[2] # input to lfads should have dimensions:
ntimesteps = train_data.shape[1] # (batch_size x ntimesteps x data_dim)
batch_size = 128 # batch size during optimization
# LFADS architecture - The size of the numbers is rather arbitrary,
# but relatively small because we know the integrator RNN isn't too high
# dimensional in its activity.
enc_dim = 128 # encoder dim
con_dim = 128 # controller dim
ii_dim = 1 # inferred input dim, we know there is 1 dim in integrator RNN
gen_dim = 128 # generator dim, should be large enough to generate integrator RNN dynamics
factors_dim = 32 # factors dim, should be large enough to capture most variance of dynamics
# Numerical stability
var_min = 0.001 # Minimal variance any gaussian can become.
# Optimization HPs that percolates into model
l2reg = 0.00002
# -
# ### Hyperparameters for Priors
#
# As was mentioned above, LFADS is an auto-encoder and auto-encoders typically encode data through some kind of information bottleneck. The idea is a lot like PCA, if one gets rid of unimportant variation, then perhaps meaningful and interesting structure in the data will become apparent.
#
# More precisely, LFADS is a **variational auto-encoder (VAE)**, which means that the bottleneck is achieved via probabilistic methods. Namely, each trial initial state is encoded in a per-trial Gaussian distribution called the 'posterior', e.g. initial state parameter's mean and variance are given by $(\mu(\mathbf{x}), \sigma^2(\mathbf{x}))$, where $\mathbf{x}$ is the data. This then is compared to an **uninformative prior** $(\mu_p, \sigma^2_p)$, where uninformative means the prior is independent of the data, including that trial. A type of distance for distributions is used, called the KL-divergence, to force the initial state Gaussian distribution for each trial to be as close to as possible to a Gaussian that doesn't depend on the trial. This is a part of the **ELBO** - Evidence Lower BOund - that is used to train VAEs.
#
# In summary, one way of explaining VAEs is that they are auto-encoders, but they are attempting to limit the information flow from the input to the output using bottleneck based on probability distributions, basically forcing the generator to generate your data from white noise. This is doomed to fail if training works, but in the process, it learns a probabilistic generative model of your data.
#
# In this LFADS architecture, there are two posterior distributions, based on the data, and two prior distributions, unrelated to the data. They are distributions for the initial state and the distributions for the inferred input.
# Initial state prior parameters
# the mean is set to zero in the code
ic_prior_var = 0.1 # this is $\sigma^2_p$ in above paragraph
# ### Hyperparameters for inferred inputs
#
# The inferred inputs are also codes represented by posterior distributions, but now *each time point* is a Gaussian, so each inferred input time series is really a Gaussian process. A natural uninformative prior to comparing the Gaussian process to is the [autoregressive-1](https://en.wikipedia.org/wiki/Autoregressive_model#Example:_An_AR(1)_process) process or AR-1 process for short.
#
# $s_t = c + \phi s_{t-1} + \epsilon_t, \mbox{ with } \epsilon_t \in N(0, \sigma^2_n) $
#
# with c the process mean, $\phi$ giving dependence of process state at time $t-1$ to process state at time $t$ and $\epsilon_t$ is the noise with variance $\sigma^2_n$. In LFADS, $c$ is always $0$.
#
# So if you have 4 inferred inputs, then you have 4 AR-1 process priors. Utilizing an AR-1 process prior *to sequences* allows us to introduce another useful concept, **the auto-correlation** of each sequence. The auto-correlation is the correlation between values in the process at different time points. We are interested in auto-correlation because we may want to penalize very jagged or very smooth inferred inputs on a task by task case, as well as for other technical reasons. As it turns out, the input to the integrator RNN in this tutorial is uncorrelated white noise, so this concept is not too important, but in general it may be very important.
#
# So just like the initial states, which introduced multi-variate Gaussian distributions (the posteriors) for each data trial and an uninformative prior to which the per-trial posteriors are compared, we do the same thing with inferred inputs, now using the KL-divergence to compare the distribution of auto-regressive sequences to uninformative AR-1 priors. In this way, we aim to limit how informative the inferred inputs are by introducing a bottleneck between the encoder and the generator.
# Inferred input autoregressive prior parameters
# Again, these hyper parameters are set "in the ballpark" but otherwise
# pretty randomly.
ar_mean = 0.0 # process mean
ar_autocorrelation_tau = 1.0 # seconds, how correlated each time point is, related to $\phi$ above.
ar_noise_variance = 0.1 # noise variance
lfads_hps = {'data_dim' : data_dim, 'ntimesteps' : ntimesteps,
'enc_dim' : enc_dim, 'con_dim' : con_dim, 'var_min' : var_min,
'ic_prior_var' : ic_prior_var, 'ar_mean' : ar_mean,
'ar_autocorrelation_tau' : ar_autocorrelation_tau,
'ar_noise_variance' : ar_noise_variance,
'ii_dim' : ii_dim, 'gen_dim' : gen_dim,
'factors_dim' : factors_dim,
'l2reg' : l2reg,
'batch_size' : batch_size}
# #### LFADS Optimization hyperparameters
# +
num_batches = 20000 # how many batches do we train
print_every = 100 # give information every so often
# Learning rate HPs
step_size = 0.05 # initial learning rate
decay_factor = 0.9999 # learning rate decay param
decay_steps = 1 # learning rate decay param
# Regularization HPs
keep_rate = 0.98 # dropout keep rate during training
# Numerical stability HPs
max_grad_norm = 10.0 # gradient clipping above this value
# -
# ### Warming up the KL penalties
# The optimization of a VAE optimizes the ELBO, which is
#
# $L(\theta) = -\mathop{\mathbb{E}}_x \left(\log p_\theta(x|z) + KL(q_\theta(z|x) \;\;|| \;\;p(z))\right)$
#
# * $p_\theta(x|z)$ - the reconstruction given the initial state and inferred inputs distributions (collectively denoted $z$ here)
#
# * $q_\theta(z|x)$ - represents the latent variable posterior distributions (the data encoders that ultimately yield the initial state and inferred input codes).
#
# * $p(z)$ - the prior that does not know about the data
#
# where $\theta$ are all the trainable parameters. This is an expectation over all your data, $x$, of the quality of the data generation $p_\theta(x|z)$, plus the KL divergence penalty mentioned above that compares the distributions for the initial state and inferred inputs to uninformative priors.
#
# **All the hacks in hacksville:** It turns out that the KL term can be a lot easier to optimize initially than learning how to reconstruct your data. This results in a pathological stoppage of training where the KL goes to nearly zero and training is broken there on out (as you cannot represent any a given trial from uninformative priors). One way out of this is to warm up the KL penalty, starting it off with a weight term of 0 and then slowly building to 1, giving the reconstruction a chance to train a bit without the KL penalty messing things up.
# The fact that the start and end values are required to be floats is something I need to fix.
kl_warmup_start = 500.0 # batch number to start KL warm-up, explicitly float
kl_warmup_end = 1000.0 # batch number to be finished with KL warm-up, explicitly float
kl_min = 0.01 # The minimum KL value, non-zero to make sure KL doesn't grow crazy before kicking in.
# Note, there is currently a HUGE amount of debate about what the correct parameter value here is for the KL penalty. kl_max = 1 is what creates a lower bound on the (marginal) log likelihood of the data, but folks argue it could be higher or lower than 1. Myself, I have never played around with this HP, but I have the idea that LFADS may benefit from < 1 values, as LFADS is not really being used for random sampling from the distribution of spiking data.
#
# See [$\beta$-VAE: LEARNING BASIC VISUAL CONCEPTS WITH A
# CONSTRAINED VARIATIONAL FRAMEWORK](https://openreview.net/pdf?id=Sy2fzU9gl)
#
# See [Fixing a Broken ELBO](https://arxiv.org/pdf/1711.00464.pdf) as to why you might choose a particular KL maximum value. I found this article pretty clarifying.
kl_max = 1.0
# +
lfads_opt_hps = {'num_batches' : num_batches, 'step_size' : step_size,
'decay_steps' : decay_steps, 'decay_factor' : decay_factor,
'kl_min' : kl_min, 'kl_max' : kl_max, 'kl_warmup_start' : kl_warmup_start,
'kl_warmup_end' : kl_warmup_end, 'keep_rate' : keep_rate,
'max_grad_norm' : max_grad_norm, 'print_every' : print_every,
'adam_b1' : 0.9, 'adam_b2' : 0.999, 'adam_eps' : 1e-1}
assert num_batches >= print_every and num_batches % print_every == 0
# +
# Plot the warmup function and the learning rate decay function.
plt.figure(figsize=(16,4))
plt.subplot(121)
x = onp.arange(0, num_batches, print_every)
kl_warmup_fun = get_kl_warmup_fun(lfads_opt_hps)
plt.plot(x, [kl_warmup_fun(i) for i in onp.arange(1,lfads_opt_hps['num_batches'], print_every)]);
plt.title('KL warmup function')
plt.xlabel('Training batch');
plt.subplot(122)
decay_fun = optimizers.exponential_decay(lfads_opt_hps['step_size'],
lfads_opt_hps['decay_steps'],
lfads_opt_hps['decay_factor'])
plt.plot(x, [decay_fun(i) for i in range(1, lfads_opt_hps['num_batches'], print_every)]);
plt.title('learning rate function')
plt.xlabel('Training batch');
# -
# ### Train the LFADS model
#
# Note that JAX uses its own setup to handle randomness and seeding the pseudo-random number generators. You can read about it [here](https://github.com/google/jax/blob/master/README.md#random-numbers-are-different). If you want to modify the LFADS tutorial you *NEED* to understand this. Otherwise, not so big a deal if you are just messing around with LFADS hyperparameters or applying the tutorial to new data.
# Initialize parameters for LFADS
key = random.PRNGKey(onp.random.randint(0, utils.MAX_SEED_INT))
init_params = lfads.lfads_params(key, lfads_hps)
# Note that the first loop could take a few minutes to run, because the LFADS model is unrolled, and therefor the JIT (just in time) compilation is slow, and happens "just in time", which is the first training loop iteration. On my computer, the JIT compilation takes a few minutes.
#
# You'll see the loss go up when the KL warmup starts turning on.
#
key = random.PRNGKey(onp.random.randint(0, utils.MAX_SEED_INT))
trained_params, opt_details = \
optimize_lfads(key, init_params, lfads_hps, lfads_opt_hps,
train_data, eval_data)
# Plot the training details
x = onp.arange(0, num_batches, print_every)
plt.figure(figsize=(20,6))
plt.subplot(251)
plt.plot(x, opt_details['tlosses']['total'], 'k')
plt.ylabel('Training')
plt.title('Total loss')
plt.subplot(252)
plt.plot(x, opt_details['tlosses']['nlog_p_xgz'], 'b')
plt.title('Negative log p(z|x)')
plt.subplot(253)
plt.plot(x, opt_details['tlosses']['kl_ii'], 'r')
plt.title('KL inferred inputs')
plt.subplot(254)
plt.plot(x, opt_details['tlosses']['kl_g0'], 'g')
plt.title('KL initial state')
plt.subplot(255)
plt.plot(x, opt_details['tlosses']['l2'], 'c')
plt.xlabel('Training batch')
plt.title('L2 loss')
plt.subplot(256)
plt.plot(x, opt_details['elosses']['total'], 'k')
plt.xlabel('Training batch')
plt.ylabel('Evaluation')
plt.subplot(257)
plt.plot(x, opt_details['tlosses']['nlog_p_xgz'], 'b')
plt.xlabel('Training batch')
plt.subplot(258)
plt.plot(x, opt_details['elosses']['kl_ii'], 'r')
plt.xlabel('Training batch')
plt.subplot(259)
plt.plot(x, opt_details['elosses']['kl_g0'], 'g')
plt.xlabel('Training batch');
# See the effect of the KL warmup, which is shown
# by the KL penalities without the warmup scaling.
plt.figure(figsize=(7,4))
plt.subplot(221)
plt.plot(x, opt_details['tlosses']['kl_ii_prescale'], 'r--')
plt.ylabel('Training')
plt.subplot(222)
plt.plot(x, opt_details['tlosses']['kl_g0_prescale'], 'g--')
plt.subplot(223)
plt.plot(x, opt_details['elosses']['kl_ii_prescale'], 'r--')
plt.ylabel('Evaluation')
plt.xlabel('Training batch')
plt.subplot(224)
plt.plot(x, opt_details['elosses']['kl_g0_prescale'], 'g--')
plt.xlabel('Training batch');
# ### Save the LFADS model parameters
# +
fname_uniquifier = datetime.datetime.now().strftime("%Y-%m-%d_%H:%M:%S")
network_fname = ('trained_params_' + rnn_type + '_' + task_type + '_' + \
fname_uniquifier + '.npz')
network_path = os.path.join(output_dir, network_fname)
# Note we are just using numpy save instead of h5 because the LFADS parameter
# is nested dictionaries, something I couldn't get h5 to save down easily.
print("Saving parameters: ", network_path)
onp.savez(network_path, trained_params)
# -
# After training, you can load these up, after locating the save file.
if False:
loaded_params = onp.load(network_path, allow_pickle=True)
trained_params = loaded_params['arr_0'].item()
# ### LFADS Visualization
# To plot the results of LFADS, namely the inferred quantities such as the inferred inputs, factors, or rates, we have to do a sample-and-average operation. Remember, the latent variables for LFADS are the initial state and the inferred inputs, and they are per-trial *stochastic* codes, even for a *single trial*. To get good inference for a given trial, we sample a large number of times from these per-trial stochastic latent variables, run the generator forward, and then average all the quantities of interest over the samples.
#
# If LFADS were linear a linear model, it would be equivalent to do the *much more efficient decode* of the posterior means, that is, just take the mean of the initial state distribution and the mean of the inferred input distribution, and then run the decoder one time. (This, btw, is a great exercise to the tutorial reader: implement posterior-mean decoding in this tutorial.)
#
# Here we use batching and take the 'posterior average' using batch number of samples from the latent variable distributions.
#
# So the main result of this tutorial, the moment you've been waiting for, is the comparison between the true rates of the integrator RNN, and the inferred rates by LFADS, and the true input to the integrator RNN and the inferred inputs given by LFADS. You can see how well we did by generating lots of trials here.
# +
# Plot a bunch of examples of eval trials run through LFADS.
reload(plotting)
#reload(lfads)
def plot_rescale_fun(a):
fac = max_firing_rate * data_dt
return renormed_fun(a) * fac
bidx = my_example_bidx - eval_data_offset
bidx = 0
nexamples_to_save = 1
for eidx in range(nexamples_to_save):
fkey = random.fold_in(key, eidx)
psa_example = eval_data[bidx,:,:].astype(np.float32)
psa_dict = lfads.posterior_sample_and_average_jit(trained_params, lfads_hps, fkey, psa_example)
# The inferred input and true input are rescaled and shifted via
# linear regression to match, as there is an identifiability issue. there.
plotting.plot_lfads(psa_example, psa_dict,
data_dict, eval_data_offset+bidx, plot_rescale_fun)
# -
# And coming back to our example signal, how well does LFADS do on it, compared to the other *much easier to implement* methods? A noticeable improvement on inferring the underlying rate.
# +
plt.figure(figsize=(16,4))
plt.subplot(141)
plt.plot(my_signal, 'r');
plt.stem(my_signal_spikified);
_, _, r2_spike, _, _ = scipy.stats.linregress(my_signal_spikified, my_signal)
plt.title('Raw spikes R^2=%.3f' % (r2_spike))
plt.legend(('True rate', 'Spikes'));
plt.subplot(142)
plt.plot(my_signal, 'r');
plt.plot(my_filtered_spikes);
_, _, c_tfilt, _, _ = scipy.stats.linregress(my_filtered_spikes, my_signal)
plt.title("Temporal filtering R^2=%.3f" % (c_tfilt**2));
plt.legend(('True rate', 'Filtered spikes'));
plt.subplot(143)
plt.plot(my_signal, 'r')
plt.plot(my_example_ipca[:,my_example_hidx])
_, _, c_sfilt, _, _ = scipy.stats.linregress(my_example_ipca[:,my_example_hidx], my_signal)
plt.legend(('True rate', 'PCA smoothed spikes'))
plt.title('Spatial filtering R^2=%.3f' % (c_sfilt**2));
plt.subplot(144)
plt.plot(my_signal, 'r')
my_lfads_rate = onp.exp(psa_dict['lograte_t'][:,my_example_hidx])
plt.plot(my_lfads_rate)
_, _, r2_lfads, _, _ = scipy.stats.linregress(my_lfads_rate, my_signal)
plt.legend(('True rate', 'LFADS rate'))
plt.title('LFADS "filtering" R^2=%.3f' % (r2_lfads));
# -
# That single example can't tell the whole story so let us look at the average. LFADS is much better than spatial averaging across a large set of trials.
#
# Take an average over all the hidden units in 100 evaluation trials.
# +
nexamples = 1000
r2_sfilts = onp.zeros(nexamples*data_dim)
r2_lfadss = onp.zeros(nexamples*data_dim)
eidx = 0
for bidx in range(nexamples):
ebidx = eval_data_offset + bidx
# Get the LFADS decode for this trial.
fkey = random.fold_in(key, bidx)
psa_example = eval_data[bidx,:,:].astype(np.float32)
psa_dict = lfads.posterior_sample_and_average_jit(trained_params, lfads_hps, fkey, psa_example)
# Get the spatially smoothed trial.
trial_rates = scale*renormed_data[ebidx, :, :]
trial_spikes = data_bxtxn[ebidx, :, :]
spikes_pca = pca.transform(trial_spikes)
spikes_ipca = pca.inverse_transform(spikes_pca)
for hidx in range(data_dim):
sig = trial_rates[:, hidx]
ipca_rate = spikes_ipca[:,hidx]
lfads_rate = onp.exp(psa_dict['lograte_t'][:,hidx])
_, _, cc_sfilt, _, _ = scipy.stats.linregress(ipca_rate, sig)
_, _, cc_lfads, _, _ = scipy.stats.linregress(lfads_rate, sig)
r2_sfilts[eidx] = cc_sfilt**2
r2_lfadss[eidx] = cc_lfads**2
eidx += 1
plt.figure(figsize=(8,4))
plt.subplot(121)
plt.hist(r2_sfilts, 50)
plt.title('Spatial filtering, hist of R^2, <%.3f>' % (onp.mean(r2_sfilts)))
plt.xlim([-.5, 1.0])
plt.subplot(122)
plt.hist(r2_lfadss, 50);
plt.title('LFADS filtering, hist of R^2, <%.3f>' % (onp.mean(r2_lfadss)));
plt.xlim([-.5, 1.0]);
# -
# ### Compare the inferred inputs learned by LFADS to the actual inputs to the integrator RNN.
#
# Finally, we can look at the average correlation between the inferred inputs and the true inputs to the integrator RNN. The inferred input can be arbitrarily scaled or rotated, so we first compute a linear regression, to scale the inferred input correctly, and then get the $R^2$.
# +
r2_iis = []
nexamples = 1000
for bidx in range(nexamples):
ebidx = eval_data_offset + bidx
# Get the LFADS decode for this trial.
psa_example = eval_data[bidx,:,:].astype(np.float32)
fkey = random.fold_in(key, bidx)
psa_dict = lfads.posterior_sample_and_average_jit(trained_params, lfads_hps, fkey, psa_example)
# Get the true input and inferred input
true_input = onp.squeeze(data_dict['inputs'][ebidx])
inferred_input = onp.squeeze(psa_dict['ii_t'])
slope, intercept, _, _, _ = scipy.stats.linregress(inferred_input, true_input)
_, _, cc_ii, _, _ = scipy.stats.linregress(slope * inferred_input + intercept, true_input)
r2_iis.append(cc_ii**2)
r2_iis = onp.array(r2_iis)
plt.hist(r2_iis, 20)
plt.title('Correlation between rescaled inferrred inputs and true inputs, hist of R^2, <%.3f>' % (onp.mean(r2_iis)))
plt.xlim([0.0, 1.0]);
# -
# ### Compare the inferred initial state for the LFADS generator to the actual initial state of the integrator RNN.
#
# To finish, we can examine the relationship between the initial condition (h0) of the integrator RNN and the inferred initial condition of the LFADS generator.
# The color we use is the readout of the integrator RNN's initial state, so basically, the state of the line attractor before further information is presented. In the integrator RNN example, we made sure to seed these initial states with various values along the line attractor, so we expect a line of coloration.
ntrials = 1000
true_h0s = onp.zeros([ntrials, data_dim])
ic_means = onp.zeros([ntrials, gen_dim])
colors = onp.zeros(ntrials)
for bidx in range(ntrials):
ebidx = eval_data_offset + bidx
# Get the LFADS decode for this trial.
psa_example = eval_data[bidx,:,:].astype(np.float32)
fkey = random.fold_in(key, bidx)
#psa_dict = lfads.posterior_sample_and_average_jit(trained_params, lfads_hps, fkey, psa_example)
lfads_results = lfads.lfads_jit(trained_params, lfads_hps, fkey, psa_example, 1.0)
# Get the true initial condition (and the readout of the true h0 for color)
# Get the inferred input from LFADS
true_h0s[bidx,:] = data_dict['h0s'][ebidx]
colors[bidx] = data_dict['outputs'][ebidx][0]
ic_means[bidx,:] = lfads_results['ic_mean']
from sklearn.manifold import TSNE
plt.figure(figsize=(16,8))
plt.subplot(121)
h0s_embedded = TSNE(n_components=2).fit_transform(true_h0s)
plt.scatter(h0s_embedded[:,0], h0s_embedded[:,1], c=colors)
plt.title('TSNE visualization of integrator RNN intial state')
plt.subplot(122)
ic_means_embedded = TSNE(n_components=2).fit_transform(ic_means)
plt.scatter(ic_means_embedded[:,0], ic_means_embedded[:,1], c=colors);
plt.title('TSNE visualziation of LFADS inferred intial generator state.')
| notebooks/LFADS Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Entorno de experimentación `HMMLike`
#
# Entorno dónde se utiliza la minima información en la construcción de las *feature lists*. Esto es bias, la letra actual y la letra anterior. Con esto se simula un HMM pero construido con los CRFs
#
# ### Parámetros generales
#
# * Maximo Iteraciones = 50
# * K = 3
#
# ### Parametros por modelo
#
# * `linearCRF_reg.crfsuite`
# * l1 = 0.1
# * l2 = 0.001
# * `HMMLike_baseline.crfsuite`
# * l1 = 0.0
# * l2 = 0.0
# * `linearCRF_l1_zero.crfsuite`
# * l1 = 0
# * l2 = 0.001
# * `linearCRF_l2_zero.crfsuite`
# * l1 = 0.1
# * l2 = 0
#
#
# ### Importando bibliotecas de python
import os
import sys
import random
import time
import pycrfsuite
import numpy as np
from sklearn.model_selection import KFold
from utils import (get_corpus, WordsToLetter, accuracy_score, model_trainer,
model_tester, write_report, eval_labeled_positions,
bio_classification_report)
# ### Funciones auxiliares
# +
def get_feature_lists(sent):
''' Rules that setting up the feature lists for training
:param sent: Data as `[[[[[letter, POS, BIO-label],...],words],sents]]`
:type: list
:return: list of words with characters as features list:
[[[[[letterfeatures],POS,BIO-label],letters],words]]
:rtype: list
'''
featurelist = []
senlen = len(sent)
# each word in a sentence
for i in range(senlen):
word = sent[i]
wordlen = len(word)
lettersequence = ''
# each letter in a word
for j in range(wordlen):
letter = word[j][0]
# gathering previous letters
lettersequence += letter
# ignore digits
if not letter.isdigit():
features = [
'bias',
'letterLowercase=' + letter.lower(),
]
if j >= 1:
features.append('prevletter=' + lettersequence[j-1:j].lower() + '>')
featurelist.append(features)
return featurelist
def get_labels(sent, flag=0):
labels = []
for word in sent:
for letter in word:
labels.append(letter[2])
return labels
def sent2features(data):
return [get_feature_lists(sent) for sent in data]
def sent2labels(data):
return [get_labels(sent) for sent in data]
# -
# ### Funciones de Train y Test
# +
def model_trainer(train_data, hyper):
""" Entrena un modelo y lo guarda en disco
Función encargada de entrenar un modelo con base en los hyperparametro y
lo guarda como un archivo utilizable por `pycrfsuite`
Parameters
----------
train_data : list
models_path : str
hyper : dict
verbose : bool
k : int
Returns
-------
train_time : float
Tiempo de entrenamiento
compositive_name : str
Nombre del modelo entrenado
"""
X_train = sent2features(train_data)
y_train = sent2labels(train_data)
# Train the model
trainer = pycrfsuite.Trainer(verbose=True)
for xseq, yseq in zip(X_train, y_train):
trainer.append(xseq, yseq)
# Set training parameters. L-BFGS is default. Using Elastic Net (L1 + L2)
trainer.set_params({
'c1': hyper['L1'], # coefficient for L1 penalty
'c2': hyper['L2'], # coefficient for L2 penalty
'max_iterations': hyper['max-iter'] # early stopping
})
# The program saves the trained model to a file:
start = time.time()
trainer.train(hyper['path'])
end = time.time()
train_time = end - start
return train_time
def model_tester(test_data, model_path):
""" Prueba un modelo preentrenado
Recibe los datos de prueba y realiza las pruebas con el modelo previo
Parameters
----------
test_data : list
models_path : str
model_name : str
verbose : bool
Returns
-------
y_test : list
Etiquetas reales
y_pred : list
Etiquetas predichas por el modelo
tagger : Object
Objeto que etiqueta con base en el modelo
"""
X_test = sent2features(test_data)
y_test = sent2labels(test_data)
# ### Make Predictions
tagger = pycrfsuite.Tagger()
# Passing model to tagger
tagger.open(model_path)
# Tagging task using the model
y_pred = [tagger.tag(xseq) for xseq in X_test]
# Closing tagger
tagger.close()
return y_test, y_pred
# -
# ## Obteniendo corpus completo
corpus = get_corpus('corpus_otomi_mod', '../corpora/') + \
get_corpus('corpus_otomi_hard', '../corpora/')
letter_corpus = WordsToLetter(corpus)
dataset = np.array(letter_corpus, dtype=object)
# ## Parametros base
models_path = 'models/'
env_name = "HMMLike"
max_iter = 50
k = 3
kf = KFold(n_splits=k, shuffle=True)
# ## Parámetros para `HMMLike_reg.crfsuite`
params = {"L1": 0.1, "L2": 1e-3, "max-iter": max_iter}
variant = "reg"
# ### Entrenamiento y Tests
# %%time
i = 0
full_time = 0
accuracy_set = []
for train_index, test_index in kf.split(dataset):
i += 1
train_data, test_data = dataset[train_index], dataset[test_index]
model_name = f"{env_name}_{variant}_k_{i}.crf"
params['path'] = os.path.join(models_path, env_name, model_name)
print("*"*50)
print(f"Entrenando nuevo modelo '{model_name}' | K = {i}")
print(f"len train: {len(train_data)} len test: {len(test_data)}")
print("*"*50)
train_time = model_trainer(train_data, params)
full_time += train_time
print("*"*50)
print(f"Tiempo de entrenamiento: {train_time}[s] | {train_time / 60}[m]")
print("Test del modelo")
y_test, y_pred = model_tester(test_data, params['path'])
accuracy_set.append(accuracy_score(y_test, y_pred))
print(f"Partial accuracy: {accuracy_set[i - 1]}\n")
# Reports
eval_labeled_positions(y_test, y_pred)
print(bio_classification_report(y_test, y_pred))
print("\n\nAccuracy Set -->", accuracy_set)
train_time_format = str(round(full_time / 60, 2)) + "[m]"
print(f"\nTime>> {train_time_format}")
train_size = len(train_data)
test_size = len(test_data)
params['k-folds'] = k
write_report(model_name, train_size, test_size, accuracy_set, train_time_format,
params)
# ## Parámetros para `HMMLike_l1_zero.crfsuite`
params = {"L1": 0.0, "L2": 1e-3, "max-iter": max_iter}
variant = "l1_zero"
# ### Entrenamiento y Tests
# %%time
i = 0
full_time = 0
accuracy_set = []
for train_index, test_index in kf.split(dataset):
i += 1
train_data, test_data = dataset[train_index], dataset[test_index]
model_name = f"{env_name}_{variant}_k_{i}.crf"
params['path'] = os.path.join(models_path, env_name, model_name)
print("*"*50)
print(f"Entrenando nuevo modelo '{model_name}' | K = {i}")
print(f"len train: {len(train_data)} len test: {len(test_data)}")
print("*"*50)
train_time = model_trainer(train_data, params)
full_time += train_time
print("*"*50)
print(f"Tiempo de entrenamiento: {train_time}[s] | {train_time / 60}[m]")
print("Test del modelo")
y_test, y_pred = model_tester(test_data, params['path'])
accuracy_set.append(accuracy_score(y_test, y_pred))
print(f"Partial accuracy: {accuracy_set[i - 1]}\n")
# Reports
eval_labeled_positions(y_test, y_pred)
print(bio_classification_report(y_test, y_pred))
print("\n\nAccuracy Set -->", accuracy_set)
train_time_format = str(round(full_time / 60, 2)) + "[m]"
print(f"\nTime>> {train_time_format}")
train_size = len(train_data)
test_size = len(test_data)
params['k-folds'] = k
write_report(model_name, train_size, test_size, accuracy_set, train_time_format,
params)
# ## Parámetros para `HMMLike_l2_zero.crfsuite`
params = {"L1": 0.1, "L2": 0.0, "max-iter": max_iter}
variant = "l2_zero"
# ### Entrenamiento y Tests
# %%time
i = 0
full_time = 0
accuracy_set = []
for train_index, test_index in kf.split(dataset):
i += 1
train_data, test_data = dataset[train_index], dataset[test_index]
model_name = f"{env_name}_{variant}_k_{i}.crf"
params['path'] = os.path.join(models_path, env_name, model_name)
print("*"*50)
print(f"Entrenando nuevo modelo '{model_name}' | K = {i}")
print(f"len train: {len(train_data)} len test: {len(test_data)}")
print("*"*50)
train_time = model_trainer(train_data, params)
full_time += train_time
print("*"*50)
print(f"Tiempo de entrenamiento: {train_time}[s] | {train_time / 60}[m]")
print("Test del modelo")
y_test, y_pred = model_tester(test_data, params['path'])
accuracy_set.append(accuracy_score(y_test, y_pred))
print(f"Partial accuracy: {accuracy_set[i - 1]}\n")
# Reports
eval_labeled_positions(y_test, y_pred)
print(bio_classification_report(y_test, y_pred))
print("\n\nAccuracy Set -->", accuracy_set)
train_time_format = str(round(full_time / 60, 2)) + "[m]"
print(f"\nTime>> {train_time_format}")
train_size = len(train_data)
test_size = len(test_data)
params['k-folds'] = k
write_report(model_name, train_size, test_size, accuracy_set, train_time_format,
params)
# ## Parámetros para `baseline.crfsuite`
params = {"L1": 0.0, "L2": 0.0, "max-iter": max_iter}
variant = "baseline"
# ### Entrenamiento y Tests
# %%time
i = 0
full_time = 0
accuracy_set = []
for train_index, test_index in kf.split(dataset):
i += 1
train_data, test_data = dataset[train_index], dataset[test_index]
model_name = f"{env_name}_{variant}_k_{i}.crf"
params['path'] = os.path.join(models_path, env_name, model_name)
print("*"*50)
print(f"Entrenando nuevo modelo '{model_name}' | K = {i}")
print(f"len train: {len(train_data)} len test: {len(test_data)}")
print("*"*50)
train_time = model_trainer(train_data, params)
full_time += train_time
print("*"*50)
print(f"Tiempo de entrenamiento: {train_time}[s] | {train_time / 60}[m]")
print("Test del modelo")
y_test, y_pred = model_tester(test_data, params['path'])
accuracy_set.append(accuracy_score(y_test, y_pred))
print(f"Partial accuracy: {accuracy_set[i - 1]}\n")
# Reports
eval_labeled_positions(y_test, y_pred)
print(bio_classification_report(y_test, y_pred))
print("\n\nAccuracy Set -->", accuracy_set)
train_time_format = str(round(full_time / 60, 2)) + "[m]"
print(f"\nTime>> {train_time_format}")
train_size = len(train_data)
test_size = len(test_data)
params['k-folds'] = k
write_report(model_name, train_size, test_size, accuracy_set, train_time_format,
params)
| notebooks/HMMLike.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TonyRahme/cs480student/blob/main/01/CS480FourierFreq.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="sMCTz9kCLPAw" colab={"base_uri": "https://localhost:8080/"} outputId="6489819f-5c00-4f83-aa7c-40031a5e3761"
#import numpy as np
#import matplotlib.pyplot as plt
# %pylab inline
# + [markdown] id="Bra6c_M5LmNm"
# Populating the interactive namespace from numpy and matplotlib
# + id="eAqbPWn_Lx1m"
#lets create a sine wave
fs = 10000
t = np.arange(0, 1, step = 1. / fs)
frequency = 20
s_t = np.sin(2 * np.pi * frequency * t)
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="jsv7CpvAMNPW" outputId="fea360b3-314a-4cb7-c952-5a5ce2b0be9a"
plt.xlabel('Time [s]')
plt.ylabel('s(t)')
plt.title('Sine Wave with Frequency '+str(frequency))
plt.plot(t, s_t)
# + colab={"base_uri": "https://localhost:8080/", "height": 61} id="LsV0p14PNKAu" outputId="8ee5b5ea-18fc-4890-fc68-9477a67c1bb7"
from IPython.display import Audio
Audio(s_t, rate=fs)
# + id="N6GGQ5lbNcPZ"
#let's make a fourier transform
S = np.fft.fft(s_t)
# + id="uvq1qNiMNkzl"
S_pow = S ** 2
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="KqLHvaWONpbZ" outputId="984df696-12a7-47e7-9424-b8effc54eee9"
plt.xlim(0, 100)
plt.xlabel('Frequency [Hz]')
plt.ylabel('S^2(f)')
plt.plot(np.abs(S_pow))
# + id="gAQjKgy_OtGU"
restored_S = np.sqrt( S_pow )
inv_S = np.fft.ifft( restored_S )
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="ksoG_YqaPCPd" outputId="750252ce-2084-4e69-dd6a-63d041cef281"
plt.plot(inv_S)
# + id="av8CVQQPPPck"
#let's create some random noise
noise = np.random.normal(0., 0.1, s_t.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="TxlLg-MBPj02" outputId="148d42ec-3b95-4792-a4cb-297f94935fca"
plt.plot(noise)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="LNY5-RmjPq2I" outputId="6fd937b0-6005-43a3-dffb-5d763c5eab4b"
#let's add noise to s(t)
s_t_noise = s_t + noise
plt.plot(s_t_noise)
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="46clTg2mQGEF" outputId="cf6cd04e-a16f-417f-8b3d-ad5b88dccf1b"
S_noise = np.fft.fft(s_t_noise)
S_noise_pow = S_noise ** 2
plt.xlim(0, 100)
plt.ylim(0, 1000)
plt.xlabel('Frequency [Hz]')
plt.ylabel('S^2(f)')
plt.plot(np.abs(S_noise_pow))
# + colab={"base_uri": "https://localhost:8080/"} id="vMRcTXVQdxCo" outputId="00e33e49-2d22-435a-eaf5-e8be0e932415"
S_noise_pow[21:] = 0
np.abs(S_noise_pow[15:])
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="-32lMdqcffll" outputId="8a04d9d1-0305-496b-d363-155a1c866171"
plt.xlim(0, 100)
plt.ylim(0, 1000)
plt.xlabel('Frequency [Hz]')
plt.ylabel('S^2(f)')
plt.plot(np.abs(S_noise_pow))
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="10ZDh_a8ghn7" outputId="c24689f9-4ddc-4783-ca43-89c50cea1d8e"
restored_S = np.sqrt( S_noise_pow )
inv_S = np.fft.ifft( restored_S )
plt.plot(inv_S)
# + id="tTG0tcGSgttq"
| 01/CS480FourierFreq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Car Rental 1
#
# ## Objective and Prerequisites
#
# Boost your modeling skills with this example, which will teach you how you can use mathematical optimization to figure out how many cars a car rental company should own and where they should be located every day to maximize weekly profits.
#
# This model is example 25 from the fifth edition of Model Building in Mathematical Programming by <NAME> on pages 284-286 and 340-342.
#
# This example is at the intermediate level, where we assume that you know Python and the Gurobi Python API and that you have some knowledge of building mathematical optimization models.
#
# **Download the Repository** <br />
# You can download the repository containing this and other examples by clicking [here](https://github.com/Gurobi/modeling-examples/archive/master.zip).
# ---
# ## Problem Description
#
# A small car rental company (which rents only one type of car) has depots in Glasgow, Manchester, Birmingham and Plymouth. There is an estimated demand for each day of the week except Sunday (when the company is closed). These estimates are given in the following table. It is not necessary to meet all demand.
#
# 
#
#
# Cars can be rented for one, two or three days and returned to either the
# depot from which they were rented or another depot at the start of the next morning. For
# example, a 2-day rental on Thursday means that the car has to be returned on
# Saturday morning; a 3-day rental on Friday means that the car has to be returned
# on Tuesday morning. A 1-day rental on Saturday means that the car has to be
# returned on Monday morning, and a 2-day rental on Tuesday morning.
#
# The rental period is independent of the origin and destination. From historical
# data, the company knows the distribution of rental periods: 55% of cars are hired
# for one day, 20% for two days, and 25% for three days. The current estimates
# of percentages of cars rented from each depot and returned to a given depot
# (independent of the day) are given in the following table.
#
# 
#
# The company's marginal cost of renting out a car (‘wear and tear’, administration, etc.) is estimated as follows:
#
# | Days rented | Marginal cost |
# | --- | --- |
# | 1-day | $\$ 20$ |
# | 2-day | $\$ 25$ |
# | 3-day | $\$ 30$ |
#
# The ‘opportunity cost’ (interest on capital, storage, servicing, etc.) of owning
# a car is $\$ 15$ per week.
#
# It is possible to transfer undamaged cars from one depot to another depot,
# irrespective of distance. Cars cannot be rented out during the day in which they
# are transferred. The costs in USD, per car, of transfer are given in the following table.
#
# 
#
# Ten percent of cars returned by customers are damaged. When this happens,
# the customer is charged an excess of $\$ 100$ (irrespective of the amount of damage
# that the company completely covers by its insurance). In addition, the car has
# to be transferred to a repair depot, where it will be repaired the following day.
# The cost of transferring a damaged car is the same as transferring an undamaged
# one (except when the repair depot is the current depot,
# in which case the cost would be $\$0$). The transfer of a damaged car takes a day, unless it is already at a repair depot.
# Having arrived at a repair depot, all types of repair (or replacement) take a day.
# Only two of the depots have repair capacity. The (cars/day) capacity at each repair depot is as follows:
#
# | Repair depot | Capacity |
# | --- | --- |
# | Manchester | 12 |
# | Birmingham | 20 |
#
# Having been repaired, the car is available for rent at the depot the next day
# or may be transferred to another depot (taking a day). Thus, a car that is returned
# damaged on a Wednesday morning is transferred to a repair depot (if not the
# current depot) on Wednesday, repaired on Thursday, and is available for rent
# at the repair depot on Friday morning.
# The rental price depends on the number of days for which the car is rented
# and whether it is returned to the same depot or not. The prices (in USD) are given in the following table.
#
# 
#
# We assume the following at the beginning of each day:
# 1. Customers return cars that are due that day.
# 2. Damaged cars are sent to the repair depot.
# 3. Cars that were transferred from other depots arrive.
# 4. Transfers are sent out.
# 5. Cars are rented out.
# 6. If it is a repair depot, then the repaired cars are available for rental.
#
# The goal is to determine the numbers of cars the car rental company should own and where should they be located at the start of each week day in order to maximize weekly profit. The company wants a ‘steady state’ solution
# in which the same expected number of cars will be located at the same depot on
# the same day of subsequent weeks.
# ---
# ## Model Formulation
#
# $d,d2 \in \text{Depots}=\{\text{Glasgow}, \text{Manchester}, \text{Birmingham}, \text{Plymouth}\}$
#
# $\text{NRD}=\{\text{Glasgow}, \text{Plymouth}\}$: Depots without repair capacity.
#
# $\text{RD}=\{\text{Manchester}, \text{Birmingham}\}$: Depots with repair capacity.
#
# $t \in \text{Days}=\{\text{Monday},\text{Tuesday},\text{Wednesday},\text{Thursday},\text{Friday},\text{Saturday}\}$
#
# $r \in \text{RentDays}=\{1,2,3\}$: Number of days rented.
#
# ### Parameters
#
# $\text{demand}_{d,t} \in \mathbb{R}^+$: Estimated rental demand at depot $d$ on day $t$.
#
# $\text{pctDepot}_{d,d2} \in \mathbb{R}^+$: Proportion of cars rented at depot $d$ to be returned to depot $d2$.
#
# $\text{cstTransfer}_{d,d2} \in \mathbb{R}^+$: Transfer cost of a car from depot $d$ to depot $d2$.
#
# $\text{pctRent}_{r} \in \mathbb{R}^+$: Proportion of cars rented for $r$ days.
#
# $\text{capRepair}_{d} \in \mathbb{R}^+$: Repair capacity of depot $d$.
#
# $\text{cstSameDepot}_{r} \in \mathbb{R}^+$: Rental cost for $r$ days with return to same depot.
#
# $\text{cstOtherDepot}_{r} \in \mathbb{R}^+$: Rental cost for $r$ days with return to other depot.
#
# $\text{marginalCost}_{r} \in \mathbb{R}^+$: Marginal cost to company of $r$ days rental of a car.
#
# $\text{pctUndamaged } \in [0,1]$: Percent of cars returned by customers that are undamaged.
#
# $\text{pctDamaged } \in [0,1]$: Percent of cars returned by customers that are damaged.
#
# $\text{cstOwn} \in \mathbb{R}^+$: Cost of owning a car.
#
# $\text{damagedFee} = 10$: Damaged car fee. Ten percent of the cars are damaged and the fee for a damaged car is $\$100$.
#
# ### Decision Variables
#
# $\text{xOwned} \in \mathbb{R}^+$: Total number of cars owned.
#
# $\text{xUndamaged}_{d,t} \in \mathbb{R}^+$: Number of undamaged cars available at depot $d$ at the beginning of day $t$.
#
# $\text{xDamaged}_{d,t} \in \mathbb{R}^+$: Number of damaged cars available at depot $d$ at the beginning of day $t$.
#
# $\text{xRented}_{d,t} \in \mathbb{R}^+$: number of cars rented out from depot $d$ at the beginning of day $t$.
#
# $\text{xUDleft}_{d,t} \in \mathbb{R}^+$: Number of undamaged cars available at depot $d$ at the beginning of day $t$.
#
# $\text{xDleft}_{d,t} \in \mathbb{R}^+$: Number of damaged cars left in depo $d$ at the end of day $t$.
#
# $\text{xUDtransfer}_{d,d2,t} \in \mathbb{R}^+$: Number of undamaged cars at depot $d$ at the beginning of day $t$, to be transferred to depot $d2$.
#
# $\text{xDtransfer}_{d,d2,t} \in \mathbb{R}^+$: Number of damaged cars at depot $d$ at the beginning of day $t$, to be transferred to depot $d2$.
#
# $\text{xRepaired}_{d,t} \in \mathbb{R}^+$: Number of damaged cars to be repaired at depot $d$ during day $t$.
# ### Objective function
# The objective is to maximize profit.
#
# \begin{equation}
# \sum_{d \in \text{Depots}}
# \sum_{t \in \text{Days}}
# \sum_{r \in \text{RentDays}}
# \text{pctDepot}_{d,d}*\text{pctRent}_{r}*(\text{cstSameDepot}_{r} - \text{marginalCost}_{r} + \text{damagedFee})*\text{xRented}_{d,t}
# \end{equation}
#
# \begin{equation}
# + \sum_{d \in \text{Depots}} \sum_{d2 \in \text{Depots}}
# \sum_{t \in \text{Days}}
# \sum_{r \in \text{RentDays}}
# \text{pctDepot}_{d,d2}*\text{pctRent}_{r}*(\text{cstOtherDepot}_{r} - \text{marginalCost}_{r} + \text{damagedFee})*\text{xRented}_{d,t}
# \end{equation}
#
# \begin{equation}
# - \sum_{d \in \text{Depots}} \sum_{d2 \in \text{Depots}}
# \sum_{t \in \text{Days}} \text{cstTransfer}_{d,d2}*(\text{xUDtransfer}_{d,d2,t} + \text{xDtransfer}_{d,d2,t} )
# - \text{cstOwn}*\text{xOwned}
# \end{equation}
#
#
# ### Constraints
#
# **Undamaged cars at a non-repair depot** <br />
# Number of undamaged cars available at a non-repair depot $d$ at the beginning of day $t$.
#
# \begin{equation}
# \sum_{d2 \in \text{Depots}}
# \sum_{r \in \text{RentDays}} \text{pctUndamaged}*\text{pctDepot}_{d2,d}*\text{pctRent}_{r}*\text{xRented}_{d2,(t-r)mod(6)}
# \end{equation}
#
# \begin{equation}
# + \sum_{d2 \in \text{Depots}} \text{xUDtransfer}_{d2,d,(t-1)mod(6)} + \text{xUDleft}_{d,(t-1)mod(6)} = \text{xUndamaged}_{d,t}
# \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# Demand of undamaged cars at the non-repair depot $d$ during day $t$.
#
# \begin{equation}
# \text{xUndamaged}_{d,t} = \text{xRented}_{d,t} +
# \sum_{d2 \in \text{Depots}} \text{xUDtransfer}_{d,d2,t} + \text{xUDleft}_{d,t}
# \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# **Undamaged cars at a repair depot** <br />
# Number of undamaged cars available at a repair depot $d$ at the beginning of day $t$.
#
# \begin{equation}
# \sum_{d2 \in \text{Depots}}
# \sum_{r \in \text{RentDays}} \text{pctUndamaged}*\text{pctDepot}_{d2,d}*\text{pctRent}_{r}*\text{xRented}_{d2,(t-r)mod(6)}
# \end{equation}
#
# \begin{equation}
# + \sum_{d2 \in \text{Depots}} \text{xUDtransfer}_{d2,d,(t-1)mod(6)}
# + \text{xRepaired}_{d,(t-1)mod(6)} + \text{xUDleft}_{d,(t-1)mod(6)} = \text{xUndamaged}_{d,t}
# \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# Demand of undamaged cars at the repair depot $d$ during day $t$.
#
# \begin{equation}
# \text{xUndamaged}_{d,t} = \text{xRented}_{d,t} +
# \sum_{d2 \in \text{Depots}} \text{xUDtransfer}_{d,d2,t} + \text{xUDleft}_{d,t}
# \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# **Damaged cars at a non-repair depot** <br />
# Number of damaged cars available at a non-repair depot $d$ at the beginning of day $t$.
#
# \begin{equation}
# \sum_{d2 \in \text{Depots}}
# \sum_{r \in \text{RentDays}} \text{pctDamaged}*\text{pctDepot}_{d2,d}*\text{pctRent}_{r}*\text{xRented}_{d2,(t-r)mod(6)}
# \end{equation}
#
# \begin{equation}
# + \text{xDleft}_{d,(t-1)mod(6)} = \text{xDamaged}_{d,t} \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# Demand of undamaged cars at the non-repair depot $d$ during day $t$.
#
# \begin{equation}
# \text{xDamaged}_{d,t} =
# \sum_{d2 \in \text{Depots} \cap RD} \text{xDtransfer}_{d,d2,t} + \text{xDleft}_{d,t}
# \quad \forall d \in NRD, t \in Days
# \end{equation}
#
# **Damaged cars at a repair depot** <br />
# Number of damaged cars available at a repair depot $d$ at the beginning of day $t$.
#
# \begin{equation}
# \sum_{d2 \in \text{Depots}}
# \sum_{r \in \text{RentDays}} \text{pctDamaged}*\text{pctDepot}_{d2,d}*\text{pctRent}_{r}*\text{xRented}_{d2,(t-r)mod(6)}
# \end{equation}
#
# \begin{equation}
# + \sum_{d2 \in \text{Depots}} \text{xDtransfer}_{d2,d,(t-1)mod(6)}
# + \text{xDleft}_{d,(t-1)mod(6)} = \text{xdamaged}_{d,t}
# \quad \forall d \in RD, t \in Days
# \end{equation}
#
# Demand of undamaged cars at the non-repair depot $d$ during day $t$.
#
# \begin{equation}
# \text{xDamaged}_{d,t} = \text{xRepaired}_{d,t} +
# \sum_{d2 \in \text{Depots} \cap RD} \text{xDtransfer}_{d,d2,t} + \text{xDleft}_{d,t}
# \quad \forall d \in RD, t \in Days
# \end{equation}
#
# **Depot Capacity** <br />
# Repair capacity of depot $d$ for each day $t$.
#
# \begin{equation}
# \text{xRepaired}_{d,t} \leq \text{capRepair}_{d}
# \quad \forall d \in Depots, t \in Days
# \end{equation}
#
# **Depot Demand** <br />
# Demand at depot $d$ for each day $t$.
#
# \begin{equation}
# \text{xRented}_{d,t} \leq \text{demand}_{d,t}
# \quad \forall d \in Depots, t \in Days
# \end{equation}
#
# **Number of cars** <br />
# Total number of cars owned equals number of cars rented out from all depots on Monday for 3 days, plus those on Tuesday for 2 or 3 days, plus all damaged and undamaged cars in depots at the beginning of Wednesday.
# Rationale: Let’s pick a day (Wednesday), count the cars undamaged and damaged that were returned to the depots and that are available on Wednesday morning. Let’s count the cars that have been rented and have not been returned: Cars rented on Monday for 3 days, and cars rented on Tuesday for 2 or 3 days.
#
# \begin{equation}
# \sum_{d \in \text{Depots}} (0.25*\text{xRented}_{d,0} + 0.45*\text{xRented}_{d,1} + \text{xUndamaged}_{d,2} + \text{xdamaged}_{d,2} ) = \text{xOwned}
# \end{equation}
#
# ---
# ## Python Implementation
#
# We import the Gurobi Python Module and other Python libraries.
# %pip install gurobipy
# +
import pandas as pd
from itertools import product
import gurobipy as gp
from gurobipy import GRB
# tested with Python 3.7.0 & Gurobi 9.0
# -
# ## Input data
#
# We define all the input data for the model.
# +
# list of depots and working days of a week
depots = ['Glasgow','Manchester','Birmingham','Plymouth']
NRD = ['Glasgow','Plymouth'] # Non-repair depot
RD =['Manchester','Birmingham'] # Repair depot
days = [0,1,2,3,4,5] # Monday = 0, Tuesday = 1, ... Saturday = 5
rentDays = [1,2,3]
d2w, demand = gp.multidict({
('Glasgow',0): 100,
('Glasgow',1): 150,
('Glasgow',2): 135,
('Glasgow',3): 83,
('Glasgow',4): 120,
('Glasgow',5): 230,
('Manchester',0): 250,
('Manchester',1): 143,
('Manchester',2): 80,
('Manchester',3): 225,
('Manchester',4): 210,
('Manchester',5): 98,
('Birmingham',0): 95,
('Birmingham',1): 195,
('Birmingham',2): 242,
('Birmingham',3): 111,
('Birmingham',4): 70,
('Birmingham',5): 124,
('Plymouth',0): 160,
('Plymouth',1): 99,
('Plymouth',2): 55,
('Plymouth',3): 96,
('Plymouth',4): 115,
('Plymouth',5): 80
})
#repairCap
depots, capacity = gp.multidict({
('Glasgow'): 0,
('Manchester'): 12,
('Birmingham'): 20,
('Plymouth'): 0
})
# Create a dictionary to capture
# pctRent: percentage of cars rented for r days
# cstMarginal: marginal cost for renting a car for r days
# prcSameD: price of renting a car r days and returning to same depot
# prcOtherD: price of renting a car r days and returning to another depot
rentDays, pctRent, costMarginal, priceSameD, priceOtherD = gp.multidict({
(1): [0.55,20,50,70],
(2): [0.20,25,70,100],
(3): [0.25,30,120,150]
})
# Cost of owing a car per week.
cstOwn = 15
# Proportional damaged car fee
damagedFee = 10
# Create a dictionary to capture the proportion of cars rented at depot d to be returned to depot d2
d2d, pctFromToD = gp.multidict({
('Glasgow','Glasgow'): 0.6,
('Glasgow','Manchester'): 0.2,
('Glasgow','Birmingham'): 0.1,
('Glasgow','Plymouth'): 0.1,
('Manchester','Glasgow'): 0.15,
('Manchester','Manchester'): 0.55,
('Manchester','Birmingham'): 0.25,
('Manchester','Plymouth'): 0.05,
('Birmingham','Glasgow'): 0.15,
('Birmingham','Manchester'): 0.2,
('Birmingham','Birmingham'): 0.54,
('Birmingham','Plymouth'): 0.11,
('Plymouth','Glasgow'): 0.08,
('Plymouth','Manchester'): 0.12,
('Plymouth','Birmingham'): 0.27,
('Plymouth','Plymouth'): 0.53
})
# Create a dictionary to capture the transfer costs of cars
d2d, cstFromToD = gp.multidict({
('Glasgow','Glasgow'): 0.001,
('Glasgow','Manchester'): 20,
('Glasgow','Birmingham'): 30,
('Glasgow','Plymouth'): 50,
('Manchester','Glasgow'): 20,
('Manchester','Manchester'): 0.001,
('Manchester','Birmingham'): 15,
('Manchester','Plymouth'): 35,
('Birmingham','Glasgow'): 30,
('Birmingham','Manchester'): 15,
('Birmingham','Birmingham'): 0.001,
('Birmingham','Plymouth'): 25,
('Plymouth','Glasgow'): 50,
('Plymouth','Manchester'): 35,
('Plymouth','Birmingham'): 25,
('Plymouth','Plymouth'): 0.001
})
# Proportion of undamaged and damaged cars returned
pctUndamaged = 0.9
pctDamaged = 0.1
# -
# ### Preprocessing
# We prepare the data structures to build the linear programming model.
# +
# Build a list of tuples (depot, depot2) such that d != d2
list_d2notd = []
for d,d2 in d2d:
if (d != d2):
tp = d,d2
list_d2notd.append(tp)
d2notd = gp.tuplelist(list_d2notd)
# Build a list of tuples (depot, depot2, day)
list_dd2t = []
for d,d2 in d2notd:
for t in days:
tp = d,d2,t
list_dd2t.append(tp)
dd2t = gp.tuplelist(list_dd2t)
# Build a list of tuples (depot, rent_day)
list_dr = []
for d in depots:
for r in rentDays:
tp = d,r
list_dr.append(tp)
dr = gp.tuplelist(list_dr)
# Build a list of tuples (depot, day, rent_days )
list_dtr = []
for d in depots:
for t in days:
for r in rentDays:
tp = d,t,r
list_dtr.append(tp)
dtr = gp.tuplelist(list_dtr)
# Build a list of tuples (depot, depot2, day, rent_days)
list_dd2tr = []
for d,d2 in d2notd:
for t in days:
for r in rentDays:
tp = d,d2,t,r
list_dd2tr.append(tp)
dd2tr = gp.tuplelist(list_dd2tr)
# -
# ## Model Deployment
# We create a model and the variables. The main decision variables are the number of cars to own
# and where should they be located at the start of each day of a week to maximize weekly profits.
# +
model = gp.Model('RentalCar1')
# Number of cars owned
n = model.addVar(name="cars")
# Number of undamaged cars
nu = model.addVars(d2w, name="UDcars")
# Number of damaged cars
nd = model.addVars(d2w, name="Dcars")
# Number of cars hired (rented) cannot exceed their demand
tr = model.addVars(d2w, ub=demand, name="Hcars")
#for d,t in d2w:
#tr[d,t].lb = 1
# End inventory of undamaged cars
eu = model.addVars(d2w, name="EUDcars")
# End inventory of damaged cars
ed = model.addVars(d2w, name="EDcars")
# Number of undamaged cars transferred
tu = model.addVars(dd2t, name="TUDcars")
# Number of damaged cars transferred
td = model.addVars(dd2t, name="TDcars")
# Number of damaged cars repaired
rp = model.addVars(d2w, name="RPcars")
# Number of damaged cars repaired cannot exceed depot capacity
for d,t in d2w:
rp[d,t].ub = capacity[d] #repair capacity
# -
# ### Constraints
# The number of undamaged cars available at a non-repair depot d at the beginning of day t should be equal to the demand of undamaged cars at the non-repair depot d during day t.
# +
# Undamaged cars into a non-repair depot constraints (left hand side of balance equation -availability)
UDcarsNRD_L = model.addConstrs((gp.quicksum(pctUndamaged*pctFromToD[d2,d]*pctRent[r]*tr[d2,(t-r)%6 ] for d2,r in dr )
+ gp.quicksum(tu.select('*',d,(t-1)%6) )
+ eu[d,(t-1)%6 ] == nu[d,t] for d in NRD for t in days ),
name="UDcarsNRD_L")
# Undamaged cars out of a non-repair depot constraints (right hand side of balance equation -requirements)
UDcarsNRD_R = model.addConstrs((tr[d,t]
+ gp.quicksum(tu.select(d,'*',t ))
+ eu[d,t] == nu[d,t] for d in NRD for t in days ), name='UDcarsNRD_R' )
# -
# The number of undamaged cars available at a repair depot d at the beginning of day t should be equal to the demand of undamaged cars at the repair depot d during day t.
# +
# Undamaged cars into a repair depot constraints (left hand side of balance equation -availability)
UDcarsRD_L = model.addConstrs((gp.quicksum(pctUndamaged*pctFromToD[d2,d]*pctRent[r]*tr[d2,(t-r)%6 ] for d2,r in dr )
+ gp.quicksum(tu.select('*',d,(t-1)%6) ) + rp[d, (t-1)%6 ]
+ eu[d,(t-1)%6 ] == nu[d,t] for d in RD for t in days ),
name="UDcarsRD_L")
# Undamaged cars out of a repair depot constraints (right hand side of balance equation -requirements)
UDcarsRD_R = model.addConstrs((tr[d,t]
+ gp.quicksum(tu.select(d,'*',t ) )
+ eu[d,t] == nu[d,t] for d in RD for t in days ), name='UDcarsRD_R' )
# -
# The number of damaged cars available at a non-repair depot d at the beginning of day t should be equal to the demand of damaged cars at the non-repair depot d during day t.
# +
# Damaged cars into a non-repair depot constraints (left hand side of balance equation -availability)
DcarsNRD_L = model.addConstrs((gp.quicksum(pctDamaged*pctFromToD[d2,d]*pctRent[r]*tr[d2,(t-r)%6 ] for d2,r in dr )
+ ed[d,(t-1)%6 ] == nd[d,t] for d in NRD for t in days ),
name="DcarsNRD_L")
# Damaged cars out of a non-repair depot constraints (right hand side of balance equation -requirements)
DcarsNRD_R = model.addConstrs(( gp.quicksum(td[d,d2,t] for d2 in RD )
+ ed[d,t] == nd[d,t] for d in NRD for t in days ), name='DcarsNRD_R' )
# -
# The number of damaged cars available at a repair depot d at the beginning of day t should be equal to the demand of damaged cars at the repair depot d during day t.
# +
# Damaged cars into a repair depot constraints (left hand side of balance equation -availability)
DcarsRD_L = model.addConstrs((gp.quicksum(pctDamaged*pctFromToD[d2,d]*pctRent[r]*tr[d2,(t-r)%6 ] for d2,r in dr )
+ gp.quicksum(td[d2,d,(t-1)%6 ] for d2, dd in d2notd if (dd == d))
+ ed[d,(t-1)%6 ] == nd[d,t] for d in RD for t in days ),
name="DcarsRD_L")
# Damaged cars out of a repair depot constraints (right hand side of balance equation -requirements)
DcarsND_R = model.addConstrs((rp[d,t] + gp.quicksum(td[d,d2,t ] for d2 in NRD )
+ ed[d,t] == nd[d,t] for d in RD for t in days ), name='DcarsND_R' )
# -
# Total number of cars equals the number of cars rented out from all depots on Monday for 3 days, plus those on Tuesday for 2 or 3 days, plus all damaged and undamaged cars in depots at the beginning of Wednesday.
# +
# Total number of cars owned constraint
# Note: 25% of cars are rented for 3 days, and 20% + 25% = 45% of the cars are rented for 2-days or 3-days
carsConstr = model.addConstr((gp.quicksum(0.25*tr[d,0] + 0.45*tr[d,1] + nu[d,2] + nd[d,2] for d in depots )
== n ),name='carsConstr')
# -
# The objective function is to maximize profit.
# +
# Maximize profit objective function
model.setObjective((
gp.quicksum(pctFromToD[d,d]*pctRent[r]*(priceSameD[r] - costMarginal[r] + damagedFee)*tr[d,t] for d,t,r in dtr )
+ gp.quicksum(pctFromToD[d,d2]*pctRent[r]*(priceOtherD[r]-costMarginal[r]+damagedFee)*tr[d,t] for d,d2,t,r in dd2tr)
- gp.quicksum(cstFromToD[d,d2]*tu[d,d2,t] for d,d2,t in dd2t)
- gp.quicksum(cstFromToD[d,d2]*td[d,d2,t] for d,d2,t in dd2t) - cstOwn*n ), GRB.MAXIMIZE)
# +
# Verify model formulation
model.write('CarRental1.lp')
# Run optimization engine
model.optimize()
# -
# ---
# ## Analysis
# +
# Output report
# Total number of cars owned
print(f"The optimal number of cars to be owned is: {round(n.x)}.")
# Optimal profit
print(f"The optimal profit is: {'${:,.2f}'.format(round(model.objVal,2))}.")
# +
# Create a list to translate the number label of each day to the actual name of the day
dayname = ['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday']
# Number of undamaged cars in depot at the beginning of each day.
print("\n\n_________________________________________________________________________________")
print(f"Estimated number of undamaged cars in depot at the beginning of each day: ")
print("_________________________________________________________________________________")
undamaged_cars = pd.DataFrame(columns=['Day','Glasgow','Manchester','Birmingham','Plymouth'])
for t in days:
undamaged_cars = undamaged_cars.append({"Day": dayname[t], "Glasgow": round(nu['Glasgow',t].x), "Manchester": round(nu['Manchester',t].x), 'Birmingham': round(nu['Birmingham',t].x),'Plymouth': round(nu['Plymouth',t].x) }, ignore_index=True)
undamaged_cars.index=[''] * len(undamaged_cars)
undamaged_cars
# +
# Number of Damaged cars in depot at the beginning of each day.
print("_________________________________________________________________________________")
print(f"Estimated number of damaged cars in depot at the beginning of each day: ")
print("_________________________________________________________________________________")
damaged_cars = pd.DataFrame(columns=['Day','Glasgow','Manchester','Birmingham','Plymouth'])
for t in days:
damaged_cars = damaged_cars.append({"Day": dayname[t], "Glasgow": round(nd['Glasgow',t].x), "Manchester": round(nd['Manchester',t].x), 'Birmingham': round(nd['Birmingham',t].x),'Plymouth': round(nd['Plymouth',t].x) }, ignore_index=True)
damaged_cars.index=[''] * len(damaged_cars)
damaged_cars
# +
# Undamaged car rented out from each depot and day.
print("_________________________________________________________________________________")
print(f"Estimated number of undamaged cars rented out from each depot and day: ")
print("_________________________________________________________________________________")
rentedOut = {}
for d in depots:
for t in days:
count = 0
for d2 in depots:
for r in rentDays:
#print(f"Depot {d}, day {t}: cars rented out {tr[d,t].x}")
count += pctUndamaged*pctFromToD[d,d2]*pctRent[r]*tr[d,t].x
rentedOut[d,t] = round(count)
#print(rentedOut)
rentout_cars = pd.DataFrame(columns=['Day','Glasgow','Manchester','Birmingham','Plymouth'])
for t in days:
rentout_cars = rentout_cars.append({"Day": dayname[t], "Glasgow": round(rentedOut['Glasgow',t]), "Manchester": round(rentedOut['Manchester',t]), 'Birmingham': round(rentedOut['Birmingham',t]),'Plymouth': round(rentedOut['Plymouth',t]) }, ignore_index=True)
rentout_cars.index=[''] * len(rentout_cars)
rentout_cars
# -
# ---
# ## References
#
# <NAME>, Model Building in Mathematical Programming, fifth edition.
#
# Copyright © 2020 Gurobi Optimization, LLC
| car_rental_1_2/car_rental_1_gcl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ATOMconsortium/AMPL/blob/master/atomsci/ddm/examples/tutorials/10_Delaney_Solubility_Prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VQYLwMnVDT8l"
# <h1>Predicting Solubility Using AMPL</h1>
#
# The ATOM Modeling PipeLine (AMPL; https://github.com/ATOMconsortium/AMPL) is an open-source, modular, extensible software pipeline for building and sharing models to advance in silico drug discovery.
#
# + [markdown] id="d9wRTs8cHKis"
# ## Notebook execution time: ~ 3 minutes
# + colab={"base_uri": "https://localhost:8080/"} id="YxRoIMr5D0PQ" outputId="86c227bb-314c-4dbf-a0c3-fc0b51ac80e2"
# !date
# + [markdown] id="WlGjgLeOVk2w"
# # Goal: Predict solubility using the ATOM Modeling Pipeline (AMPL) on the public dataset
#
# In this notebook, we describe the following steps using AMPL:
#
# 1. Read a public dataset containing chemical structures and some properties
# 1. Curate the dataset
# 2. Fit a simple model
# 3. Predict solubility for withheld compounds
#
#
# ## Set up
# We first import the AMPL modules for use in this notebook.
#
# The relevant AMPL modules for this example are listed below:
#
# |module|Description|
# |-|-|
# |`atomsci.ddm.pipeline.model_pipeline`|The model pipeline module is used to fit models and load models for prediction.|
# |`atomsci.ddm.pipeline.parameter_parser`|The parameter parser reads through pipeline options for the model pipeline.|
# |`atomsci.ddm.utils.curate_data`|The curate data module is used for data loading and pre-processing.|
# |`atomsci.ddm.utils.struct_utils`|The structure utilities module is used to process loaded structures.|
# |`atomsci.ddm.pipeline.perf_plots`|Perf plots contains a variety of plotting functions.|
# + [markdown] id="Q4ZFqjQVMVXL"
# ## Install AMPL
# + id="RgPw1t5PO-QC"
# ! pip install rdkit-pypi
# ! pip install --pre deepchem
import deepchem
# print(deepchem.__version__)
# ! pip install umap
# ! pip install llvmlite==0.35.0 --ignore-installed
# ! pip install umap-learn
# ! pip install molvs
# ! pip install bravado
# + id="scDiD9K_KKkB"
import deepchem as dc
# get the Install AMPL_GPU_test.sh
# !wget https://raw.githubusercontent.com/ravichas/AMPL-Tutorial/master/config/install_AMPL_GPU_test.sh
# run the script to install AMPL
# ! chmod u+x install_AMPL_GPU_test.sh
# ! ./install_AMPL_GPU_test.sh
# + id="mflA92SEVk2x" colab={"base_uri": "https://localhost:8080/"} outputId="5dd1ec7f-bb13-4b3e-c901-9f5aa4181cc3"
# We temporarily disable warnings for demonstration.
# FutureWarnings and DeprecationWarnings are present from some of the AMPL
# dependency modules.
import warnings
warnings.filterwarnings('ignore')
import json
import numpy as np
import pandas as pd
import os
import requests
import sys
import atomsci.ddm.pipeline.model_pipeline as mp
import atomsci.ddm.pipeline.parameter_parser as parse
import atomsci.ddm.utils.curate_data as curate_data
import atomsci.ddm.utils.struct_utils as struct_utils
from atomsci.ddm.pipeline import perf_plots as pp
# + [markdown] id="oRrd09dCVk22"
# ## Data curation
#
# We then download and do very simple curation to the related dataset.
#
# We need to set the directory we want to save files to. Next we download the dataset.
# + id="11ccpX-onveT"
working_dir = '/content'
# + id="REL1w2ULAH8e"
import io
url = 'https://raw.githubusercontent.com/deepchem/deepchem/master/datasets/delaney-processed.csv'
download = requests.get(url).content
# + id="B524IFKtAOdR"
# Reading the downloaded content and turning it into a pandas dataframe
raw_df = pd.read_csv(io.StringIO(download.decode('utf-8')), sep=',', header=0 )
# + [markdown] id="dCxw6gNmVk29"
# Next, we load the downloaded dataset, and process the compound structures:
# + id="Lk83NzbkwfZG"
data_orig = raw_df
# + colab={"base_uri": "https://localhost:8080/"} id="XoZYHbJK_UA7" outputId="606aa326-ae70-4227-c1d5-0e8f4ad4f219"
raw_df.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="AygEGe71_YSv" outputId="469f6f4b-8ed4-4ee6-ceb0-6c7f02b5891a"
raw_df.head(5)
# + id="0vlfZzbxVk2-"
# base_smiles_from_smiles:
# Generate a standardized SMILES, InChI keys for dataset with curation and structure modules.
# RDkit modules are used to process the SMILES strings
# Default Arg options:
# useIsomericSmiles = True
# removeCharges = False
raw_df['rdkit_smiles'] = raw_df['smiles'].apply(curate_data.base_smiles_from_smiles)
# + [markdown] id="n0OctfHTBclP"
# ## What happened after calling `base_smiles_from_smiles`?
# Also remember the column name of the `rdkit_smiles`, we will use the standardized smiles later as input in our model calculations
# + colab={"base_uri": "https://localhost:8080/", "height": 278} id="E5xPHZptBNB-" outputId="c5e3d9f0-b721-43f2-ed9c-3256b61d0b0d"
raw_df.head(4)
# + [markdown] id="hqRne34HCJNJ"
# ## In the following cell, we will call smiles_to_inchi_key to create InChi Keys
#
# We will create InChi key (https://en.wikipedia.org/wiki/International_Chemical_Identifier) from SMILES string using RDKit. Note if for some reason, the convertion fails, you will see None as output
# + id="lAcU1ThYBFBE"
raw_df['inchi_key'] = raw_df['smiles'].apply(struct_utils.smiles_to_inchi_key)
# + [markdown] id="aA3QdohoCtkV"
# ## Note the addition of new column, inchi_key
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="lhRyu3ssCmWN" outputId="2b02f5c0-f651-49d6-ea48-59b41647c93d"
raw_df.head(3)
# + id="oBqiftr2BITr"
data = raw_df
data['compound_id'] = data['inchi_key']
# + [markdown] id="7FTViEdxV0tL"
# ## Let us review the dataset
# + id="76gx-pYMVqQv" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="79844a7a-f771-46e3-aacc-628d3a7306e2"
data.head(3)
# + [markdown] id="woi0jHBPRWF_"
# ## Let us view some molecules
# + id="e-PXYKNPRc1u" colab={"base_uri": "https://localhost:8080/"} outputId="5fd90cce-4c2a-4243-d23b-e91c3f3d5f45"
data['smiles'][0:10]
# + id="gFVa3jHrTJ6Y" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="dbbd9ecc-5912-480c-fc99-1a21cf519ace"
from rdkit import Chem
from rdkit.Chem import Draw
from itertools import islice
molecules = [Chem.MolFromSmiles(smiles) for smiles in islice(data['smiles'], 9)]
Draw.MolsToGridImage(molecules)
# + [markdown] id="Jmqtyr4bVk3B"
# The next step is to address the case where we have multiple measurements for a single structure (by RDkit canonical SMILEs string). We have a function in the `curate_data()` module to address the processing of compounds. The function parameters are listed below along with an explanation of each parameter:
# + colab={"base_uri": "https://localhost:8080/", "height": 696} id="itthUq-QGRfk" outputId="aca2f71c-23b2-4a39-94b1-90fe59746b8a"
data
# + id="26UUu-vzTTMs"
# del_features = ['VALUE_NUM_mean', 'VALUE_NUM_std', 'Perc_Var', 'Remove_BadDuplicate']
# data.drop(labels=del_features, axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="5ABo-cymSmMJ" outputId="80577988-7621-4112-c4ff-3a4b93b1db32"
# column: Response values column
column = 'measured log solubility in mols per litre'
# tolerance: Percentage of individual respsonse values allowed to be different
# from the average to be included in averaging
tolerance = 10
# list_bad_duplicates: Print structures with bad duplicates
list_bad_duplicates = 'Yes'
# max_std: Maximum allowed standard deviation for computed average response value
# NOTE: In this example, we set this value very high to disable this feature
max_std = 100000
# compound_id: Compound ID column
compound_id = 'compound_id'
# smiles_col: SMILES column
smiles_col = 'rdkit_smiles'
# call the method `average_and_remove_duplicates` which changes the data and returns
# the new object as curated_df, in this case
curated_df = curate_data.average_and_remove_duplicates(column,
tolerance,
list_bad_duplicates,
data,
max_std,
compound_id=compound_id,
smiles_col=smiles_col)
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="tlV94ETHTZDP" outputId="65cf8a83-b399-4866-e410-f09cf405e8e5"
data.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="Yniwt0vbTg4T" outputId="81802be1-698a-4df8-9c1f-5490f79b6054"
curated_df.head(3)
# + id="RWD0PTp_Vk3C"
curated_file = os.path.join(working_dir, 'delaney_curated.csv')
curated_df.to_csv(curated_file, index=False)
# + [markdown] id="2o98pQ8JVk3G"
# Now that we have a curated dataset, we decide what type of featurizer and model we would like. See documentation for all available options. We also set the name of the new averaged response value column.
# + id="YpzrXMrmVk3H"
featurizer = 'ecfp'
model_type = 'RF'
response_cols = ['VALUE_NUM_mean']
# + [markdown] id="VJtQSo3UVk3O"
# Next we set up the parameters for our model. We set datastore and save_results to False to indicate that we are reading the input file and saving the results directly to the file system. There are a wide range of settable parameters; see the documentation for more details.
# + id="TH1aGjcKYOMS" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="11b76d49-0203-41ca-de92-b01503ff49ea"
smiles_col
# + id="1AagWRZkVk3P"
params={"datastore": "False",
"save_results": "False",
"id_col": compound_id,
"smiles_col": smiles_col,
"response_cols": response_cols,
"featurizer": featurizer,
"model_type": model_type,
"result_dir": working_dir,
"dataset_key": curated_file}
# + [markdown] id="_2i5syhgVk3T"
# We use parse.wrapper to process our input configuration. We then build the model pipeline, train the model, and plot the predicted versus true values for our train, valid, test sets.
# + id="f8yt7nFjWwKW"
pparams = parse.wrapper(params)
# + [markdown] id="MNxTSotHicIy"
# ## Details of the following code chunk
# Here we create instances of the class ModelPipeline, called `MP`. The instance `MP` has access to data attributes and methods of the class.
# + id="sGD1DmlkW1eA"
MP = mp.ModelPipeline(pparams)
# + [markdown] id="M3aHIUuyncFo"
# ## Use `dir` function to explore data attributes and functions associated with the class.
#
# The output is a list of data object attributes. Note the attributes that are surrounded by double-underscore (ex., `__le__`) are for internal use
# and you dont have to worry about them. The regular (non underscore attributes) are of your concern. In the example shown below, these attributes start
# with `'create_model_metadata'`
# + id="B3D4pHf4nXON"
# dir(MP)
# + [markdown] id="9SRAFsWMXwfn"
# ### Train_model:
# Build model described by self.params on the training dataset described by self.params.Generate predictions for the training, validation, and test datasets, and save the predictions and performance metrics in the model results DB or in a JSON file.
#
# ## Note `train_model` is a method of class `ModelPipeline`. If you are using COLAB, mouse-over the `MP.train_model()` to view the source code
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="lq9CEnyPY1-f" outputId="659fff13-1399-4dc3-9240-e2edb6f6ee67"
import atomsci.ddm.pipeline.model_pipeline as mp
mp.ampl_version
# + colab={"base_uri": "https://localhost:8080/"} id="LGnlXLCyXff9" outputId="aadf3061-1475-4a7a-a485-c638ad67d90e"
MP.train_model()
# + id="CMb8j-6NVk3T" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="57b0348f-9772-4b93-8472-63ea86cfd517"
pp.plot_pred_vs_actual(MP)
# + id="7OSh60dtNVbc" colab={"base_uri": "https://localhost:8080/"} outputId="fb7f74e0-69c0-4fd4-d5b0-12094f657b10"
pparams
# MP.train_model()
# + id="BcUlli2L_TbR" colab={"base_uri": "https://localhost:8080/"} outputId="e93c3546-8d30-48c4-e312-593aa33ea8f1"
# !date
# + id="X35qLxgVzuN2"
| atomsci/ddm/examples/tutorials/10_Delaney_Solubility_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python385jvsc74a57bd0aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# ---
arr = [[45,25,23,52,26, 0], 5]
def removeLast(lst):
if lst[1] > 0: lst[1] = lst[1] - 1 # aktuelle länge der Liste =-1
else: raise IndexError('list assignment index out of range')
def count(arr, search_el):
c, i = 0, 0
while i < arr[1]:
if arr[0][i] == search_el:
c += 1
i += 1
return c
removeLast(arr)
arr
count(arr, 2)
def index(arr, search_el):
i = 0
while i < arr[1]:
if arr[0][i] == search_el:
return i
i += 1
raise ValueError(str(search_el) + " is not in array")
index(arr, 23243)
def insert(arr, el, index):
if index < 0:
index += arr[1]
if index >= arr[1] or index < 0:
raise IndexError("Index out of range")
else:
i = arr[1]
arr[1] += 1
while i >= index:
arr[0][i] = arr[0][i-1]
i -= 1
arr[0][index] = el
insert(arr, 2, 6)
arr
def remove(arr, el):
i = 0
removed = False
while i < arr[1]:
if removed:
arr[0][i] = arr[0][i+1]
elif arr[0][i] == el:
removed = True
arr[0][i] = arr[0][i+1]
i += 1
if removed:
arr[1] -= 1
else:
raise ValueError("Element to remove is not in array")
arr = [[45,25,23,52,26, 0], 5]
remove(arr, 26)
arr
| LE2/list-operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Logistic Regression
# This notebook tackles "binary classification" where outputs are of 2 labels such as a positive and negative, (1 or 0). To give some examples of what we mean by binary: we could classify email as either spam or not spam, or tumors as either malignant or benign. In both these cases we have a set of data and features, but only two possible outputs. It is possible to have more than just two classes, but for now we will focus on binary classification.
#
# In order to perform this classification we will be using the logistic function to perform logistic regression.
# ### Module import
#
# We need to install a new module we haven't used before: [Statsmodels](http://statsmodels.sourceforge.net/).
#
# You can install it with 'pip install statsmodels' or use your Package Manager to install 'statsmodels' depending on your Python installation. In this lecture we will only be using a dataset from it, but it can do quite a bit, including many statistical computations that SciKit Learn does.
# +
# Data Imports
import numpy as np
import pandas as pd
from pandas import Series,DataFrame
# Math
import math
# Plot imports
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# %matplotlib inline
# Machine Learning Imports
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
# For evaluating our ML results
from sklearn import metrics
# -
# Dataset Import
import statsmodels.api as sm
# ### Mathematical Overview
#
# First, let's take a look at the [Logistic Function](http://en.wikipedia.org/wiki/Logistic_function). The logistic function can take an input from negative to positive infinity and it has always has an output between 0 and 1. The logistic function is defined as:
# $$ \sigma (t)= \frac{1}{1+e^{-t}}$$
#
# A graph of the logistic function looks like this (following code):
# +
# Logistic Function
def logistic(t):
return 1.0 / (1 + math.exp((-1.0)*t) )
# Set t from -10 to 10 ( 501 elements, linearly spaced)
t = np.linspace(-10,10,501)
# Set up y values (using list comprehension)
y = np.array([logistic(ele) for ele in t])
# Plot
plt.plot(t,y)
plt.title(' Logistic Function ')
# -
# When we talked about Linear Regression last week, we could describe a [Linear Regression](http://en.wikipedia.org/wiki/Linear_regression) Function model as:
# $$ y_i = \beta _1 x_{i1} + ... + \beta _i x_{ip}$$
#
# Which was basically an expanded linear equation (y=mx+b) for various x data features. In the case of the above equation, we presume a data set of 'n' number of units, so that the data set would have the form:
# $$ [ y_i, x_{i1},...,x_{ip}]^{n}_{i=1}$$
#
# For our logistic function, if we view *t* as a linear function with a variable *x* and n=1 we could express t as:
# $$ t = \beta _0 + \beta _1 x $$
#
# Here, we've basically just substituted a linear function (form similar to y=mx+b) for t. We could then rewrite our logistic function equation as:
# $$ F(x)= \frac{1}{1+e^{-(\beta _0 + \beta _1 x)}}$$
#
# Now we can interpret F(x) as the probability that the dependent variable is a "success" case, this is a similar style of thinking as in the Binomial Distribution, in which we had successes and failures. So the formula for F(x) that we have here states that the probability of the dependent variable equaling a "success" case is equal to the value of the logistic function of the linear regression expression (the linear equation we used to replace *t* ).
#
# Inputting the linear regression expression into the logistic function allows us to have a linear regression expression value that can vary from positive to negative infinity, but after the transformation due to the logistic expression we will have an output of F(x) that ranges from 0 to 1.
#
# We can now perform a binary classification based on where F(x) lies, either from 0 to 0.5, or 0.5 to 1.
#
# #### Extra Math Resources
#
# This is a very basic overview of binary classification using Logistic Regression, if you're still interested in a deeper dive into the mathematics, check out these sources:
#
# 1.) [<NAME>'s class notes](http://cs229.stanford.edu/notes/cs229-notes1.pdf) on Logistic Regression.
#
# 2.) [CMU notes](http://www.stat.cmu.edu/~cshalizi/uADA/12/lectures/ch12.pdf) Note: Advanced math notation.
#
# 3.) [Wikipedia](http://en.wikipedia.org/wiki/Logistic_regression) has a very extensive look at logistic regression.
#
# -----------
# ### Logistic Regression Dataset Analysis
#
# Let's put Logistic Regression into use for binary classification of this [dataset](http://statsmodels.sourceforge.net/stable/datasets/generated/fair.html)
#
# The dataset is packaged within Statsmodels. It is a data set from a 1974 survey of women by Redbook magazine. Married women were asked if they have had extramarital affairs. The published work on the data set can be found in:
#
# [<NAME>. 1978. “A Theory of Extramarital Affairs,” `Journal of Political Economy`, February, 45-61.](http://fairmodel.econ.yale.edu/rayfair/pdf/1978a200.pdf)
#
# It is important to note that this data comes from a self-reported survey, which can have many issues as far as the accuracy of the data. Also this analysis isn't trying to promote any agenda concerning women or marriage, the data is just interesting but its accuracy should be met with a healthy dose of skepticism.
#
# We'll ignore those issues concerning the data and just worry about the logistic regression aspects to the data.
#
# In this case we will approach this as a classification problem by asking the question:
#
# *Given certain variables for each woman, can we classify them as either having particpated in an affair, or not participated in an affair?*
#
#
# #### DataSet Description
#
# From the [Statsmodels website](http://statsmodels.sourceforge.net/stable/datasets/generated/fair.html) we have the following information about the data:
#
# Number of observations: 6366
# Number of variables: 9
# Variable name definitions:
#
# rate_marriage : How rate marriage, 1 = very poor, 2 = poor, 3 = fair,
# 4 = good, 5 = very good
# age : Age
# yrs_married : No. years married. Interval approximations. See
# original paper for detailed explanation.
# children : No. children
# religious : How relgious, 1 = not, 2 = mildly, 3 = fairly,
# 4 = strongly
# educ : Level of education, 9 = grade school, 12 = high
# school, 14 = some college, 16 = college graduate,
# 17 = some graduate school, 20 = advanced degree
# occupation : 1 = student, 2 = farming, agriculture; semi-skilled,
# or unskilled worker; 3 = white-colloar; 4 = teacher
# counselor social worker, nurse; artist, writers;
# technician, skilled worker, 5 = managerial,
# administrative, business, 6 = professional with
# advanced degree
# occupation_husb : Husband's occupation. Same as occupation.
# affairs : measure of time spent in extramarital affairs
#
# See the original paper for more details.
#
# *Why a Statsmodels data set?* So you can have the option of working through additional example datasets included in SciKit Learn and their own tutorials.
# ### Data Visualization
#
# Now that we've done a quick overview of some math and the data we will be working with, let's go ahead and dive into the code!
#
# We will start with loading the data and visualizing it.
# Standard method of loading Statsmodels datasets into a pandas DataFrame. Note the name fair stands for 'affair' dataset.
df = sm.datasets.fair.load_pandas().data
df.head()
# Great! Let's go ahead and start our classfication by creating a new column called 'Had_Affair'. We will set this column equal to 0 if the affairs column is 0 (meaning no time spent in affairs) otherwise the 'Had_Affair' value will be set as 1, indicating that the woman had an affair.
# +
# Create check function
def affair_check(x):
if x != 0:
return "yes"
else:
return "no"
# Apply to DataFrame
df['Had_Affair'] = df['affairs'].apply(affair_check)
# -
df.head(5)
# Now let's go ahead and groupby the newly created 'Had_Affair' column. We'll do this by grouping by the column and then calling the mean aggregate function.
# Groupby Had Affair column
df.groupby('Had_Affair').mean()
# Looking at this brief glance of the data, it seems that the women who had affairs were slightly older,married longer, and slightly less religious and less educated. However, the mean values of both classes are very close for all variables.
#
# Let's go ahead and try to visualize some of this data.
# Factorplot for age with Had Affair hue
sns.factorplot("age",
hue="Had_Affair",
data=df,
kind="count",
palette='coolwarm')
# This suggests a higher probability of an affair as age increases. Let's check the number of years married.
# Factorplot for years married with Had Affair hue
sns.factorplot('yrs_married',
data=df,
hue='Had_Affair',
kind="count",
palette='coolwarm')
# Looks like probability of having an affair increases with the number of years married. Let's check the number of children.
# Factorplot for number of children with Had Affair hue
sns.factorplot('children',data=df,hue='Had_Affair',kind="count", palette='coolwarm')
# Pretty strong evidence suggesting that increased # children results in an increased probability of an affair. Finally let's check the education level.
# Factorplot for number of children with Had Affair hue
sns.factorplot('educ',data=df,hue='Had_Affair',kind="count", palette='coolwarm')
# There is a slight decrease probability of affairs with education.
# ### Data Preparation
#
# If we look at the data, we'll notice that two columns are unlike the others. Occupation and Husband's Occupation. These columns are in a format know as *Categorical Variables*. Basically they are in set quantity/category, so that 1.0 and 2.0 are seperate variables, not values along a spectrum that goes from 1-2 (e.g. There is no 1.5 for the occupation column). Pandas has a built-in method of getting [dummy variables](http://en.wikipedia.org/wiki/Dummy_variable_%28statistics%29) and creating new columns from them.
# +
# Create new DataFrames for the Categorical Variables
occ_dummies = pd.get_dummies(df['occupation'])
hus_occ_dummies = pd.get_dummies(df['occupation_husb'])
# Let's take a quick look at the results
occ_dummies.head()
# -
# Great! Now let's name the columns something a little more readable.
# Create column names for the new DataFrames
occ_dummies.columns = ['occ1','occ2','occ3','occ4','occ5','occ6']
hus_occ_dummies.columns = ['hocc1','hocc2','hocc3','hocc4','hocc5','hocc6']
# Now we will create the X and Y data sets for out logistic regression!
# +
# Set X as new DataFrame without the occupation columns or the Y target
X = df.drop(['occupation','occupation_husb','Had_Affair'],axis=1)
# Concat the dummy DataFrames Together
dummies = pd.concat([occ_dummies,hus_occ_dummies],axis=1)
# -
# Now we will concatenate all the DataFrames together.
# +
# Now Concat the X DataFrame with the dummy variables
X = pd.concat([X,dummies],axis=1)
# Preview of Result
X.head()
# -
# Now let's go ahead and set up the Y.
# +
# Set Y as Target class, Had Affair
Y = df.Had_Affair
# Preview
Y.head()
# -
# ### Multi-Collinearity
#
# Now we need to get rid of a few columns. We will be dropping the occ1 and hocc1 columns to avoid [multicollinearity](http://en.wikipedia.org/wiki/Multicollinearity#Remedies_for_multicollinearity). Multicollinearity occurs due to the [dummy variables](http://en.wikipedia.org/wiki/Dummy_variable_(statistics)) we created. This is because the dummy variables are highly correlated, our model begins to get distorted because one of the dummy variables can be linearly predicted from the others. We take care of this problem by dropping one of the dummy variables from each set, we do this at the cost of losing a data set point.
#
# The other column we will drop is the affairs column. This is because it is basically a repeat of what will be our Y target, instead of 0 and 1 it just has 0 or a number, so we'll need to drop it for our target to make sense.
# +
# Dropping one column of each dummy variable set to avoid multicollinearity
X = X.drop('occ1',axis=1)
X = X.drop('hocc1',axis=1)
# Drop affairs column so Y target makes sense
X = X.drop('affairs',axis=1)
# PReview
X.head()
# -
# In order to use the Y with SciKit Learn, we need to set it as a 1-D array. This means we need to "flatten" the array. Numpy has a built in method for this called [ravel](http://docs.scipy.org/doc/numpy/reference/generated/numpy.ravel.html). Let's use it!
# +
# Flatten array
Y = np.ravel(Y)
# Check result
Y
# -
# ### Logistic Regression with SciKit Learn
#
# Awesome! Now let's go ahead and run the logistic regression. This is a very similar process to the Linear Regression from the previous lecture. We'll create the model, the fit the data into the model, and check our accuracy score. Then we'll split the data into testing and training sets and see if our results improve.
#
# Let's start by initiating the model!
# +
# Create LogisticRegression model
log_model = LogisticRegression()
# Fit our data
log_model.fit(X,Y)
# Check our accuracy
log_model.score(X,Y)
# -
# Looks like we got a 73% accuracy rating. Let's go ahead and compare this to the original Y data. We can do this by simply taking the mean of the Y data, since it is in the format 1 or 0, we can use the mean to calulate the percentage of women who reported having affairs. This is known as checking the [null error rate](http://en.wikipedia.org/wiki/Type_I_and_type_II_errors).
# Check percentage of women that had affairs
Y.mean()
# This means that if our model just simply guessed "no affair" we would have had 1-0.32=0.68 accuracy (or 68%) accuracy. So while we are doing better than the null error rate, we aren't doing that much better.
#
# Let's go ahead and check the coefficients of our model to check what seemed to be the stronger predictors.
# Use zip to bring the column names and the np.transpose function to bring together the coefficients from the model
coeff_df = DataFrame(zip(X.columns, np.transpose(log_model.coef_)))
# Looking at the coefficients we can see that a positive coeffecient corresponds to increasing the likelihood of having an affair while a negative coefficient means it corresponds to a decreased likelihood of having an affair as the actual data value point increases.
#
# As you might expect, an increased marriage rating corresponded to a decrease in the likelihood of having an affair. Increased religiousness also seems to correspond to a decrease in the likelihood of having an affair.
#
# Since all the dummy variables (the wife and husband occupations) are positive that means the lowest likelihood of having an affair corresponds to the baseline occupation we dropped (1-Student).
# ### Testing and Training Data Sets
#
# Just like we did in the Linear Regression Lecture, we should be splitting our data into training and testing data sets. We'll follow a very similar procedure to the Linear Regression Lecture by using SciKit Learn's built-in train_test_split method.
# +
# Split the data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y)
# Make a new log_model
log_model2 = LogisticRegression()
# Now fit the new model
log_model2.fit(X_train, Y_train)
# -
# Now we can use predict to predict classification labels for the next test set, then we will reevaluate our accuracy score!
# +
# Predict the classes of the testing data set
class_predict = log_model2.predict(X_test)
# Compare the predicted classes to the actual test classes
print metrics.accuracy_score(Y_test,class_predict)
# -
#
# Now we have a 73.35% accuracy score, which is basically the same as our previous accuracy score, 72.58%.
# ### Conclusion and more Resources
#
# So what could we do to try to further improve our Logistic Regression model? We could try some [regularization techniques](http://en.wikipedia.org/wiki/Regularization_%28mathematics%29#Regularization_in_statistics_and_machine_learning) or using a non-linear model.
#
# I'll leave the Logistic Regression topic here for you to explore more possibilites on your own. Here are several more resources and tutorials with other data sets to explore:
#
# 1.) Here's another great post on how to do logistic regression analysis using Statsmodels from [yhat](http://blog.yhathq.com/posts/logistic-regression-and-python.html)!
#
# 2.) The SciKit learn Documentation includes several [examples](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) at the bottom of the page.
#
# 3.) DataRobot has a great overview of [Logistic Regression](http://www.datarobot.com/blog/classification-with-scikit-learn/)
#
# 4.) Fantastic resource from [aimotion.blogspot](http://aimotion.blogspot.com/2011/11/machine-learning-with-python-logistic.html) on the Logistic Regression and the Mathmatics of how it relates to the cost function and gradient!
| lecture08.ML2/logistic-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''torch'': conda)'
# language: python
# name: python3
# ---
import time
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision.transforms import transforms
import numpy as np
import matplotlib.pyplot as plt
import torchvision.utils as vutils
import torch.nn.functional as F
import PIL.Image as Image
# custom weights initialization called on netG and netD
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
nn.init.normal_(m.weight.data, 0.0, 0.02)
elif classname.find('BatchNorm') != -1:
nn.init.normal_(m.weight.data, 1.0, 0.02)
nn.init.constant_(m.bias.data, 0)
# +
class Generator(nn.Module):
def __init__(self, latents):
super(Generator, self).__init__()
self.generator = nn.Sequential(
nn.ConvTranspose2d(20, 64 * 8, 4, 1, 0, bias=False),
nn.BatchNorm2d(64 * 8),
nn.ReLU(True),
# state size. (64*8) x 4 x 4
nn.ConvTranspose2d(64 * 8, 64 * 4, 4, 2, 1, bias=False),
nn.BatchNorm2d(64 * 4),
nn.ReLU(True),
# state size. (64*4) x 8 x 8
nn.ConvTranspose2d(64 * 4, 64 * 2, 4, 2, 1, bias=False),
nn.BatchNorm2d(64 * 2),
nn.ReLU(True),
# state size. (64*2) x 16 x 16
nn.ConvTranspose2d(64 * 2, 64, 4, 2, 1, bias=False),
nn.BatchNorm2d(64),
nn.ReLU(True),
# state size. (64) x 32 x 32
nn.ConvTranspose2d(64, 3, 3, 1, 1, bias=False),
nn.Tanh()
)
def forward(self, x):
x = self.generator(x)
return x
class Discriminator(nn.Module):
def __init__(self,):
super(Discriminator, self).__init__()
self.discriminator = nn.Sequential(
nn.Conv2d(4, 32, 3, 2, 1, bias=False),
nn.BatchNorm2d(32),
nn.ReLU(inplace=True),
nn.Conv2d(32, 64, 3, 1, 1, bias=False),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True),
nn.Conv2d(64, 128, 3, 2, 1, bias=False),
nn.BatchNorm2d(128),
nn.ReLU(inplace=True),
nn.Conv2d(128, 128, 3, 2, 1, bias=False),
nn.BatchNorm2d(128),
nn.ReLU(inplace=True),
nn.Conv2d(128, 64, 3, 2, 1, bias=False),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True),
)
self.fc = nn.Linear(64,2)
def forward(self, x):
x = self.discriminator(x) # [100, 64, 2, 2])
x = nn.functional.adaptive_avg_pool2d(x, 1).reshape(x.shape[0], -1) # [100, 64]
output = self.fc(x)
# print(output.shape)
return output
# +
flag_gpu = 1
# Batch size during training
batch_size = 100
# Number of training epochs
epochs = 50
# Learning rate for optimizers
lr = 0.0002
# GPU
device = 'cuda:0' if (torch.cuda.is_available() & flag_gpu) else 'cpu'
print('GPU State:', device)
# Model
latent_dim = 10
G = Generator(latents=latent_dim).to(device)
D = Discriminator().to(device)
G.apply(weights_init)
D.apply(weights_init)
# Settings
g_optimizer = optim.Adam(G.parameters(), lr=lr, betas=(0.5, 0.999))
d_optimizer = optim.Adam(D.parameters(), lr=lr, betas=(0.5, 0.999))
g_scheduler = torch.optim.lr_scheduler.StepLR(g_optimizer, step_size=5, gamma=0.5)
d_scheduler = torch.optim.lr_scheduler.StepLR(d_optimizer, step_size=5, gamma=0.5)
# Load data
train_set = datasets.CIFAR10('./data', train=True, download=False, transform=transforms.ToTensor())
train_loader = DataLoader(train_set, batch_size=batch_size, shuffle=True)
# +
# Train
adversarial_loss = torch.nn.CrossEntropyLoss().to(device)
# adversarial_loss = torch.nn.BCELoss().to(device)
G.train()
D.train()
loss_g, loss_d = [],[]
start_time= time.time()
for epoch in range(epochs):
epoch += 1
total_loss_g,total_loss_d=0,0
count_d=0
for i_iter, (images, label) in enumerate(train_loader):
i_iter += 1
# ---------Train Generator-------------------------
g_optimizer.zero_grad()
# Sample noise as generator input
noise = torch.randn(images.shape[0], latent_dim, 1, 1) # G input
label_one_hot = torch.reshape(F.one_hot(label,10),(images.shape[0], latent_dim, 1, 1))
noise = torch.cat((noise,label_one_hot),1)
# print(label_one_hot.shape)
# print(noise.shape,label)
noise = noise.to(device)
# fake_label
fake_label = torch.ones(images.shape[0], dtype=torch.long).to(device) # notice: label = 1
# Generate a batch of images
fake_inputs = G(noise)
# print(label)
# print(fake_inputs.shape)
D_label = label[0]*torch.ones(1,1,32,32)
# print(D_label)
for i in range(1,images.shape[0]):
tmp = label[i]*torch.ones(1,1,32,32)
D_label = torch.cat((D_label,tmp),0)
# print(D_label.shape)
D_label = D_label.to(device)
# print(fake_inputs.shape)
fake_inputs = torch.cat((fake_inputs,D_label),1)
# print(fake_inputs.shape)
fake_outputs = D(fake_inputs)
# print(fake_outputs.shape,fake_label.shape)
# fake_outs = torch.reshape(fake_label,(100, 1, 1, 1))
# Loss measures generator's ability to fool the discriminator
loss_g_value = adversarial_loss(fake_outputs, fake_label)
loss_g_value.backward()
g_optimizer.step()
total_loss_g+=loss_g_value
loss_g.append(loss_g_value)
# -------------Train Discriminator----------------
# Zero the parameter gradients
d_optimizer.zero_grad()
# Measure discriminator's ability to classify real from generated samples
# 因為Discriminator希望判斷哪些是真的那些是生成的,所以real_label資料標註用 1,fake_label標註用 0
real_inputs = images.to(device)
real_label = torch.ones(real_inputs.shape[0], dtype=torch.long).to(device)
fake_label = torch.zeros(fake_inputs.shape[0], dtype=torch.long).to(device)
# print(real_inputs.shape,real_label.shape,fake_label.shape)
D_label = label[0]*torch.ones(1,1,32,32)
for i in range(1,images.shape[0]):
tmp = label[i]*torch.ones(1,1,32,32)
D_label = torch.cat((D_label,tmp),0)
# print(tmp)
# print(D_label.shape)
# print(real_inputs.shape)
D_label = D_label.to(device)
real_inputs = torch.cat((real_inputs,D_label),1)
# print(real_inputs.shape)
# print(real_label.shape)
# learning by Discriminator
# print('test',real_inputs.shape)
real_loss = adversarial_loss(D(real_inputs),real_label)
fake_loss = adversarial_loss(D(fake_inputs.detach()),fake_label)
loss_d_value = (real_loss + fake_loss) / 2
loss_d_value.backward()
d_optimizer.step()
total_loss_d+=loss_d_value
loss_d.append(loss_d_value)
total_loss_g/=len(train_loader)
total_loss_d/=len(train_loader)
g_scheduler.step()
d_scheduler.step()
print('[Epoch: {}/{}] D_loss: {:.3f} G_loss: {:.3f}'.format(epoch, epochs, total_loss_d.item(), total_loss_g.item()))
print('Cost Time: {}s'.format(time.time()-start_time))
# plt.show()
# torch.save(G, 'DCGAN_Generator.pth')
# torch.save(D, 'DCGAN_Discriminator.pth')
# print('Model saved.')
print('Training Finished.')
# +
import numpy as np
import matplotlib.pyplot as plt
plt.ion()
import torchvision.utils
G.eval()
# This function takes as an input the images to reconstruct
# and the name of the model with which the reconstructions
# are performed
def to_img(x):
x = x.clamp(0, 1)
return x
def show_image(img):
img = to_img(img)
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
def visualise_output(model):
with torch.no_grad():
noise = torch.randn(20, latent_dim, 1, 1)
noise = noise.to(device)
label = torch.tensor([0,1,2,3,4,5,6,7,8,9,0,1,2,3,4,5,6,7,8,9])
label_one_hot = torch.reshape(F.one_hot(label,10),(20, latent_dim, 1, 1))
label_one_hot = label_one_hot.to(device)
noise = torch.cat((noise,label_one_hot),1)
images = G(noise)
images = images.cpu()
images = to_img(images)
np_imagegrid = torchvision.utils.make_grid(images[0:20], 10, 2).numpy()
plt.imshow(np.transpose(np_imagegrid, (1, 2, 0)))
plt.show()
# plt.show()
visualise_output(G)
| gan/CGAN_cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, gc
import pygrib
import numpy as np
import pandas as pd
import xarray as xr
import multiprocessing as mp
import matplotlib.pyplot as plt
from glob import glob
from functools import partial
from matplotlib import gridspec
from datetime import datetime, timedelta
os.environ['OMP_NUM_THREADS'] = '1'
n_cores = 64
nbm_dir = '/scratch/general/lustre/u1070830/nbm/'
urma_dir = '/scratch/general/lustre/u1070830/urma/'
tmp_dir = '/scratch/general/lustre/u1070830/tmp/'
os.makedirs(tmp_dir, exist_ok=True)
def open_urma(f, cfengine='pynio'):
try:
ds = xr.open_dataset(f, engine=cfengine)
ds['valid'] = datetime.strptime(f.split('/')[-1].split('.')[1], '%Y%m%d%H')
except:
return None
else:
return ds
# -
urma_flist = sorted([f for f in glob(urma_dir + '*.WR.grib2') if 'idx' not in f])
print(len(urma_flist), ' URMA files to read')
# +
print('Producing URMA aggregate')
with mp.get_context('fork').Pool(n_cores) as p:
urma = p.map(open_urma, urma_flist, chunksize=1)
p.close()
p.join()
urma = [f for f in urma if f is not None]
urma = xr.concat(urma, dim='valid').rename({'APCP_P8_L1_GLC0_acc':'apcp6h_mm',
'xgrid_0':'x', 'ygrid_0':'y',
'gridlat_0':'lat', 'gridlon_0':'lon'})
urma = urma['apcp6h_mm']
# urma24 = xr.open_dataset(urma_dir + 'agg/urma_agg.nc')['apcp24h_mm']
# +
date_range = pd.date_range(
datetime(2020, 11, 1, 0),
datetime(2021, 3, 26, 23, 59),
freq='6H')
print(date_range)
# +
urma24 = []
print('Missing 24h period ending: ')
for date in date_range:
date0 = date - timedelta(hours=18)
date_select = urma.sel(valid=slice(date0, date))
n_select = date_select.valid.size
if n_select == 4:
date_select = date_select.sum(dim=['valid'])
date_select['valid'] = date
urma24.append(date_select)
else:
print(date)
urma24 = xr.concat(urma24, dim='valid').rename('apcp24h_mm')
# -
urma24
urma_time = np.array([pd.to_datetime(t) for t in urma24.valid.values])
urma24.isel(valid=np.where((urma_time >= datetime(2020, 11, 1, 0)) & (urma_time <= datetime(2021, 1, 31, 23, 59)))[0])
for threshold in [0.01]:
yes_count = xr.where(urma24 > threshold, 1, 0).sum(dim='valid')
yes_count.plot()
os.makedirs(urma_dir + 'agg/', exist_ok=True)
urma24.to_netcdf(urma_dir + 'agg/urma_agg.new.nc')
# +
time = urma24.valid.values
gap_ends = np.where(((time[1:] - time[:-1]).astype(np.float32)/3.6e12).astype(np.int) > 6)[0]
for ts, te in zip(time[gap_ends], time[gap_ends+1]):
missings = pd.date_range(ts, te, freq='6H')[1:-1]
for missing in missings:
print(missing)
print()
# -
| scraps/notebooks_latest/agg_urma_verif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Synthetic spectra generator
# +
#First reset sys path so we use the correct envoirnment
import sys
sys.path=['', 'C:\\Users\\bryan\\Anaconda3\\envs\\tensorflow_GPU\\python37.zip', 'C:\\Users\\bryan\\Anaconda3\\envs\\tensorflow_GPU\\DLLs', 'C:\\Users\\bryan\\Anaconda3\\envs\\tensorflow_GPU\\lib', 'C:\\Users\\bryan\\Anaconda3\\envs\\tensorflow_GPU', 'C:\\Users\\bryan\\AppData\\Roaming\\Python\\Python37\\site-packages', 'C:\\Users\\bryan\\Anaconda3\\envs\\tensorflow_GPU\\lib\\site-packages']
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
n_points = 640
# -
import tensorflow as tf
import keras.backend as K
from keras.models import Model, Sequential, load_model
from keras.layers import Dense, Conv1D, Flatten, BatchNormalization, Activation, Dropout
from keras import regularizers
from datetime import datetime
physical_devices = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
# +
tf.keras.backend.clear_session()
model = Sequential()
model.add(BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001, center=True, scale=True, beta_initializer='zeros', gamma_initializer='ones', moving_mean_initializer='zeros', moving_variance_initializer='ones', beta_regularizer=None, gamma_regularizer=None, beta_constraint=None, gamma_constraint=None,input_shape = (n_points, 1)))
model.add(Activation('relu'))
model.add(Conv1D(128, activation = 'relu', kernel_size = (32)))
model.add(Conv1D(64, activation = 'relu', kernel_size = (16)))
model.add(Conv1D(16, activation = 'relu', kernel_size = (8)))
model.add(Conv1D(16, activation = 'relu', kernel_size = (8)))
model.add(Conv1D(16, activation = 'relu', kernel_size = (8)))
model.add(Dense(32, activation = 'relu', kernel_regularizer=regularizers.l1_l2(l1 = 0, l2=0.1)))
model.add(Dense(16, activation = 'relu', kernel_regularizer=regularizers.l1_l2(l1 = 0, l2=0.1)))
model.add(Flatten())
model.add(Dropout(.25))
model.add(Dense(n_points, activation='relu'))
model.compile(loss='mse', optimizer='Adam', metrics=['mean_absolute_error','mse','accuracy'])
model.summary()
# -
# ## Training
from Generate_Data import *
for dataset_num in range(1,10):
#X, y = generate_datasets_for_Paper_1(dataset_num,50000)
temp, y = generate_datasets_(dataset_num,50000)
X = np.empty((50000,640,1))
X[:,:,0]=temp
if dataset_num == 1:
a=1
b='a'
elif dataset_num == 2:
a=1
b='b'
elif dataset_num == 3:
a=1
b='c'
elif dataset_num == 4:
a=2
b='a'
elif dataset_num == 5:
a=2
b='b'
elif dataset_num == 6:
a=2
b='c'
elif dataset_num == 7:
a=3
b='a'
elif dataset_num == 8:
a=3
b='b'
else:
a=3
b='c'
history = model.fit(X, y,epochs=10, verbose = 1, validation_split=0.25, batch_size=256)
my_path='./data/'+str(a)+b+'PAPER1'
model.save(my_path)
hist_df = pd.DataFrame(history.history)
with open(my_path+'.csv', mode='w') as f:
hist_df.to_csv(f)
my_path='./data/'+str(a)+b+'PAPER1'
#history = load_model(my_path)
# plt.plot(history.history['loss'])
# plt.plot(history.history['val_loss'])
# plt.title('Model loss')
# plt.ylabel('Loss')
# plt.xlabel('Epoch')
# plt.legend(['Train', 'Test'], loc='upper left')
# plt.show()
size1=100
name1='./data/' +str(a)+b+'Raman_spectrums_valid.csv'
name2= './data/'+str(a)+b+'CARS_spectrums_valid.csv'
RAMAN = pd.read_csv(name1)
BCARS = pd.read_csv(name2)
RAMAN = RAMAN.values[:,1:]
BCARS = BCARS.values[:,1:]
#interpolate 1000 samples down to 640 samples
from scipy import interpolate
min_wavenumber = 0.1
max_wavenumber = 2000
n_points = 1000
step = (max_wavenumber-min_wavenumber)/(n_points)
wavenumber_axis = np.arange(min_wavenumber, max_wavenumber, step)
f_RAMAN = interpolate.interp1d(wavenumber_axis, RAMAN, kind='cubic')
f_BCARS = interpolate.interp1d(wavenumber_axis, BCARS, kind='cubic')
n_points = 640
step = (max_wavenumber-min_wavenumber)/(n_points)
new_axis = np.arange(min_wavenumber, max_wavenumber, step)
RAMAN = f_RAMAN(new_axis)
BCARS = f_BCARS(new_axis)
#end interpolation
RESULTS = RAMAN[:,:]
print(np.shape(BCARS))
X = np.empty((1, n_points,1))
for m in range(size1):
X[0,:,0]=BCARS[m,:]
yhat = model.predict(X, verbose =0)
RESULTS[m,:]=yhat.flatten()
#print(RESULTS[m,:])
# print(np.shape(yhat))
# f, a = plt.subplots(2,1, sharex=True)
# a[0].plot(X.flatten(), label = 'cars')
# a[1].plot(RAMAN[m,:].T+.7, label = 'true',c= 'g' )
# a[1].plot(yhat.flatten()+1.4, label = 'pred.',c='r')
# plt.subplots_adjust(hspace=0)
pd.DataFrame(RESULTS).to_csv('./data/' +str(a)+b+'Raman_spectrums_valid_PAPER1.csv')
# Use this function to test the model on single instances
def predict_and_plot():
temp, y = generate_datasets_(dataset_num,1)
X = np.empty((1,640,1))
X[:,:,0]=temp
yhat = model.predict(X, verbose =0)
f, a = plt.subplots(2,1, sharex=True)
a[0].plot(X.flatten(), label = 'cars')
a[1].plot(y.T+.7, label = 'true',c= 'g' )
a[1].plot(yhat.flatten()+1.4, label = 'pred.',c='r')
plt.subplots_adjust(hspace=0)
#return x, y.flatten(), yhat.flatten(), chi3, NRB
predict_and_plot()
# +
# #predict_and_plot()
# for dataset_num in range(1,2):
# if dataset_num == 1:
# a=1
# b='a'
# elif dataset_num == 2:
# a=1
# b='b'
# elif dataset_num == 3:
# a=1
# b='c'
# elif dataset_num == 4:
# a=2
# b='a'
# elif dataset_num == 5:
# a=2
# b='b'
# elif dataset_num == 6:
# a=2
# b='c'
# elif dataset_num == 7:
# a=3
# b='a'
# elif dataset_num == 8:
# a=3
# b='b'
# else:
# a=3
# b='c'
# my_path='./data/'+str(a)+b+'PAPER1'
# history = load_model(my_path)
# # plt.plot(history.history['loss'])
# # plt.plot(history.history['val_loss'])
# # plt.title('Model loss')
# # plt.ylabel('Loss')
# # plt.xlabel('Epoch')
# # plt.legend(['Train', 'Test'], loc='upper left')
# # plt.show()
# size1=10
# name1='./data/' +str(a)+b+'Raman_spectrums_valid.csv'
# name2= './data/'+str(a)+b+'CARS_spectrums_valid.csv'
# RAMAN = pd.read_csv(name1)
# BCARS = pd.read_csv(name2)
# RAMAN = RAMAN.values[:,1:]
# BCARS = BCARS.values[:,1:]
# from scipy import interpolate
# min_wavenumber = 0.1
# max_wavenumber = 2000
# n_points = 1000
# step = (max_wavenumber-min_wavenumber)/(n_points)
# wavenumber_axis = np.arange(min_wavenumber, max_wavenumber, step)
# f_RAMAN = interpolate.interp1d(wavenumber_axis, RAMAN, kind='cubic')
# f_BCARS = interpolate.interp1d(wavenumber_axis, BCARS, kind='cubic')
# n_points = 640
# step = (max_wavenumber-min_wavenumber)/(n_points)
# new_axis = np.arange(min_wavenumber, max_wavenumber, step)
# RAMAN = f_RAMAN(new_axis)
# BCARS = f_BCARS(new_axis)
# #end interpolation
# RESULTS = RAMAN[:,:]
# #print(np.shape(BCARS))
# X = np.empty((1, n_points,1))
# for m in range(size1):
# X[0,:,0]=BCARS[m,:]
# yhat = model.predict(X, verbose =0)
# RESULTS[m,:]=yhat.flatten()
# #print(RESULTS[m,:])
# # print(np.shape(yhat))
# f, a = plt.subplots(2,1, sharex=True)
# a[0].plot(X.flatten(), label = 'cars')
# a[1].plot(RAMAN[m,:].T+.7, label = 'true',c= 'g' )
# a[1].plot(yhat.flatten()+1.4, label = 'pred.',c='r')
# plt.subplots_adjust(hspace=0)
# #pd.DataFrame(RESULTS).to_csv('./data/' +str(a)+b+'Raman_spectrums_valid_PAPER1.csv')
# -
| Training_SPECNET/Code_to_train_Specnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="-2JD2NccUmab"
# # História das Olimpíadas
# _(créditos ao prof. <NAME>)_
#
# Após um ano de atraso por conta da pandemia de Covid-19, as atenções do mundo todo se voltaram para Tóquio, no Japão, para acompanhar mais uma edição das Olimpíadas.
#
# No Brasil não foi diferente, e muitos se uniram para torcer por nossos atletas em diferentes competições, tanto em esportes onde o Brasil já possui tradição quanto em novos esportes.
#
# Vamos aproveitar o clima para estudar um pouco das Olimpíadas! Utilizaremos um _dataset_ com 120 anos de dados históricos das Olimpíadas, cobrindo desde os jogos de Atenas 1896 até Rio 2016.
#
# Faça o download do _dataset_ em https://www.kaggle.com/heesoo37/120-years-of-olympic-history-athletes-and-results e carregue o arquivo ```athlete_events.csv``` para um DataFrame utilizando Pandas. Aproveite para explorar seu DataFrame e se familiarizar com a sua estrutura.
#
# OBS: Fique à vontade para acrescentar mais células Python conforme necessário em qualquer etapa do exercício.
# + id="LdTmxHO4VGYr"
import pandas as pd
# + [markdown] id="U8QiBQ3KUmai"
# ## 1. O Brasil nas Olimpíadas
#
# *Vamos começar estudando o desempenho do nossos próprio país. Gere um DataFrame novo contendo apenas as informações sobre atletas brasileiros.*
# + id="kgrI2puTUmaj"
atletas = pd.read_csv("C:\\Users\\Fabio\\Desktop\\Data Science\\Módulo 3\\Aula 04\\athlete_events.csv")
atletas
# + id="-dzPmQZ5XRO3"
atletas_brasileiros = atletas[atletas["Team"]== "Brazil"]
atletas_brasileiros
# -
# POR SEXO
atletas_brasil_sexo = atletas_brasileiros["Sex"].value_counts()
atletas_brasil_sexo
# POR IDADE
atletas_brasil_idade = atletas_brasileiros["Age"].value_counts()
atletas_brasil_idade.head(5)
# média de idade por ano
media_idade_ano = atletas_brasileiros.groupby(by=["Year"])["Age"].mean()
df_media_idade_ano = pd.DataFrame(media_idade_ano)
df_media_idade_ano.head(5)
# Média máxima encontrada dentre todos os anos
df_media_idade_ano.max()
# Média mínima encontrada dentre todos os anos
df_media_idade_ano.min()
# Idade mínina encontrada entre atletas brasileiros
idade = atletas_brasileiros["Age"].min()
print(f"{idade} anos")
# Idade máxima encontrada entre atletas brasileiros
idade = atletas_brasileiros["Age"].max()
print(f"{idade} anos")
# + [markdown] id="XJ3hqw3xUmak"
# ### Medalhistas
#
# Vamos focar um pouco nos casos de sucesso do Brasil. Use o seu DataFrame anterior para filtrar apenas informações sobre **medalhistas** brasileiros.
#
# **DICA:** observe como a coluna ```Medal``` é representada quando o atleta não ganhou medalha.
# + colab={"base_uri": "https://localhost:8080/"} id="AMwTCKo6Umal" outputId="7ce31d11-3ccb-4756-a26c-a120c6b4a932"
atletas_brasileiros_medalhistas = atletas_brasileiros.groupby("Medal").size()
df_atletas_brasileiros_medalhistas = pd.DataFrame(atletas_brasileiros_medalhistas)
df_atletas_brasileiros_medalhistas
# + id="XiqA1U_ICyXh"
medalhistas_brasileiros = atletas_brasileiros.dropna(subset=["Medal"])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Tt8FiUOVD-XD" outputId="e2d20ad4-877b-495a-9a3a-faf7569b990c"
medalhistas_brasileiros.groupby(["Medal"]).head(5)
# -
# Atletas brasileiros que ganharam medalhas dividido por sexo
atletas_brasil_sexo = atletas_brasileiros.groupby(by=["Sex"])["Medal"].value_counts()
df_atletas_brasil_sexo = pd.DataFrame(atletas_brasil_sexo)
df_atletas_brasil_sexo
# Todos os medalhistas brasileiros no futebol que ganharam medalhas
atletas_brasil_futebol = medalhistas_brasileiros[medalhistas_brasileiros["Sport"] == "Football"].value_counts()
df_atletas_brasil_futebol = pd.DataFrame(atletas_brasil_futebol)
df_atletas_brasil_futebol.head(5)
# Todos os medalhistas brasileiros no futebol que ganharam medalhas de Ouro
atletas_brasil_futebol_ouro = medalhistas_brasileiros[(medalhistas_brasileiros["Medal"] == "Gold") & (medalhistas_brasileiros["Sport"] == "Football")].value_counts()
df_atletas_brasil_futebol_ouro = pd.DataFrame(atletas_brasil_futebol_ouro)
df_atletas_brasil_futebol_ouro.head(5)
# + [markdown] id="vP6Bhyq_Umam"
# ### Verão vs Inverno
#
# Você deve ter notado que temos duas categorias distintas de jogos olímpicos, representados pela estação: temos os jogos de verão e os jogos de inverno, que ocorrem de maneira intercalada.
#
# Agora que já conhecemos os medalhistas brasileiros, resposta: quantos atletas brasileiros receberam medalha nos jogos de verão e quantos receberam nos jogos de inverno?
# + colab={"base_uri": "https://localhost:8080/"} id="kDjjcU4XUmam" outputId="b603de7a-f938-42bc-dba2-378c5de38f2d"
medalhistas_brasileiros.groupby(["Season"]).size()
# + [markdown] id="oiRYTK9AUman"
# Os jogos de verão são bem mais populares do que os jogos de inverno no Brasil. Portanto, deste ponto em diante iremos focar apenas nos jogos de verão. Descarte de seu DataFrame os dados dos jogos de inverno.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 125} id="IpWlK2ZmUman" outputId="fe662ccb-2b3d-4048-d9df-b1bd09466c0e"
jogos_verao = atletas[atletas["Season"] != "Winter"].value_counts()
df_jogos_verao = pd.DataFrame(jogos_verao)
df_jogos_verao.head(5)
# -
# Participações de jogos em cada cidade (Brasileiros)
verao_medalhistas_jogos_cidade = atletas_brasileiros.groupby(by=["Games"])["City"].value_counts()
df_verao_medalhistas_jogos_cidade = pd.DataFrame(verao_medalhistas_jogos_cidade)
df_verao_medalhistas_jogos_cidade.head(5)
# + [markdown] id="ydqQUemrUmao"
# ### Atletas do Brasil
#
# Vamos conhecer um pouco melhor nossos atletas. Descubra a altura e peso médio de nossos medalhistas.
# + [markdown] id="CZyg-ve-Umao"
# Imaginamos que diferentes esportes podem beneficiar diferentes tipos físicos, certo? Então refaça a análise anterior, mas obtendo os valores médios **por esporte**.
# + id="QYeAJsVfUmap"
colunas_sport_medal = ["Name","Height","Weight","Sport","Medal"]
medalhista_esporte = medalhistas_brasileiros.filter(items= colunas_sport_medal)
colunas_sport_medal
# + colab={"base_uri": "https://localhost:8080/", "height": 383} id="g0m1fqIhLfHI" outputId="4fe7672e-7a5c-45f5-a025-0d0452d02bd8"
medalhista_esporte.groupby(["Sport"]).mean()
# + [markdown] id="bCOvfqUtUmap"
# Será que os dados acima influenciaram no interesse geral dos atletas pelo esporte ou realmente impactaram no desempenho deles? Podemos tentar descobrir se há algum tipo de correlação.
#
# Você ainda possui o dataframe original contendo todos os atletas brasileiros, incluindo os sem medalha? Obtenha os valores médios de peso e altura por esporte daquele dataframe e compare-o com os dos medalhistas. Há alguma diferença significativa em algum esporte?
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="s5JJRIYtUmaq" outputId="3d743dc6-0099-45f2-b6e0-5b2c9b380972"
atletas_brasileiros.head()
# + id="mVUC2Q6GGV5I"
colunas_selecionadas = ["Name","Height","Weight","Sport","Medal,Sex"]
filtro_atletas_brasileiros= atletas_brasileiros.filter(items= colunas_selecionadas)
# + colab={"base_uri": "https://localhost:8080/", "height": 201} id="8jsaxnnoGi_e" outputId="3318fed4-b0b2-43f8-8e01-41a52ab9eabc"
filtro_atletas_brasileiros.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="LqhgEGsvGuEd" outputId="31ca882e-d2ca-45cf-c141-755d5935bf11"
filtro_atletas_brasileiros.groupby(["Sport"]).mean()
# + colab={"base_uri": "https://localhost:8080/", "height": 383} id="bmRoihRFHLY4" outputId="c96289e0-ac9d-4f37-96c5-bebebcd41396"
medalhistas_brasileiros.groupby(["Sport"]).mean().head(5)
# + [markdown] id="zi2uA_lYH6F_"
# *Falta realizar a comparação*
# + colab={"base_uri": "https://localhost:8080/"} id="YAs5MgQLaYkE" outputId="dc48fa69-ffc4-4074-b771-e4db1df56a03"
filtro_medalhistas = medalhista_esporte.groupby(["Sport"]).mean()
filtro_atletasBrasileiros = filtro_atletas_brasileiros.groupby(["Sport"]).mean()
tabela_comparacao= pd.concat([filtro_medalhistas,filtro_atletasBrasileiros], axis=1, keys= ["Medalhistas", "Geral"])
tabela_comparacao_limpa = tabela_comparacao.dropna(thresh=3)
tabela_comparacao_limpa["Comparação Altura %"]= tabela_comparacao_limpa["Medalhistas"]["Height"]/tabela_comparacao_limpa["Geral"]["Height"]
tabela_comparacao_limpa["Comparação Peso %"]= tabela_comparacao_limpa["Medalhistas"]["Weight"]/tabela_comparacao_limpa["Geral"]["Weight"]
tabela_comparacao_limpa.head(5)
# + [markdown] id="7GIRu3SYcyvg"
# **RESPOSTA:** *FALTA FAZER *
# + [markdown] id="ncHladnFUmar"
# Existe um detalhe importante passando batido até agora em nossa análise: as categorias esportivas costumam ser divididas por gênero justamente por conta de diferenças físicas entre homens e mulheres que poderiam influenciar no desempenho. Compare a altura e peso médios de atletas brasileiros por esporte segmentado por sexo.
# + id="Wmd99WGfUmar"
colunas_selecionadas3= ["Sport","Height","Weight","Sex"]
todos_atletas_brasileiros= atletas_brasileiros.filter(items=colunas_selecionadas3)
todos_atletas_brasileiros.head(5)
# + id="PvoOv1oahzTk"
brasileiro_sport_F = todos_atletas_brasileiros[todos_atletas_brasileiros['Sex']=='F']
brasileiro_sport_M = todos_atletas_brasileiros[todos_atletas_brasileiros['Sex']=='M']
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="jxTdcyADe-XQ" outputId="b6ffec41-dca5-42e2-f9ef-4e70c2af6afe"
brasileiro_sport_F.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="wrT304tmez7N" outputId="11bdb40c-93c1-4abd-dfdf-273f34af305e"
brasileiro_sport_F.groupby(["Sport"]).mean().head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="e3Xsg1Hui081" outputId="e6935866-fd44-458d-c4de-53baba286ad2"
brasileiro_sport_M.groupby(["Sport"]).mean().head(5)
# + [markdown] id="UGQaLuZYUmas"
# Qual foi (ou quais foram) o maior medalhista brasileiro em quantidade total de medalhas?
# + id="fwFUu7GeUmau"
colunas_selecionadas2 = ["Name","Medal"]
filtro_medalhistas_brasileiros= medalhistas_brasileiros.filter(items=colunas_selecionadas2)
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="NO-hdkIcwcKN" outputId="a8d4750a-8aeb-4ebc-bc79-28468727d0e2"
filtro_medalhistas_brasileiros.groupby(["Name"]).count().sort_values(by="Medal", ascending= False).head()
# + [markdown] id="v5miQ8VpUmat"
# E o(s) maior(es) em quantidade de medalhas de ouro?
# + id="j1rYqGswNsP9"
medalha_ouro_brasil= filtro_medalhistas_brasileiros[filtro_medalhistas_brasileiros['Medal']=='Gold']
# + colab={"base_uri": "https://localhost:8080/", "height": 475} id="QISnmpf-xH7u" outputId="2953399f-17f6-45d9-80b6-4f779b9f4978"
medalha_ouro_brasil.groupby(['Name']).count().sort_values(by="Medal", ascending= False).head(5)
# + [markdown] id="amLKnFMfUmau"
# Qual esporte rendeu mais medalhas de ouro para o Brasil? E qual rendeu mais medalhas no total?
#
# **DICA:** tome muito cuidado nessa análise: cada **evento esportivo** rende 1 medalha. Por exemplo, quando a equipe de futebol vence, isso é considerado 1 medalha, mesmo tendo cerca de 20 atletas medalhistas na equipe.
# + id="eqpQ2lFhUmav"
colunas_selecionadas3 = ["Sport","Games","Event","Medal"]
medalhas_evento_brasil= medalhistas_brasileiros.filter(items=colunas_selecionadas3)
# + id="qJujJfKY0Kk-"
medalhas_evento_brasil=medalhas_evento_brasil.drop_duplicates()
# + colab={"base_uri": "https://localhost:8080/", "height": 504} id="4Uz2HEJf2eqo" outputId="bc5c9b0e-9e68-4018-b846-8d4966b59231"
medalhas_evento_brasil.groupby(["Sport"]).count().sort_values(by="Medal", ascending= False).filter(items= ["Medal"]).head(5)
# + id="0V-fFQwT3zPJ"
medalhas_sport_ouro_brasil=medalhas_evento_brasil[medalhas_evento_brasil["Medal"]=="Gold"]
medalhas_sport_ouro_brasil.groupby(["Sport"]).count().sort_values(by="Medal", ascending= False).filter(items= ["Medal"]).head(5)
# + [markdown] id="zZ84wFh0Umav"
# Cada "categoria" dentro de um esporte é considerado um evento. Por exemplo, dentro de "atletismo", temos uma competição de 100m masculina, uma de 100m feminino, um revezamento 4 x 100m masculino, um revezamento 4 x 100m feminino, uma competição de 400m masculino, uma de 400m feminino, uma maratona masculina, uma maratona feminina, e assim sucessivamente.
#
# Sabendo disso, qual evento esportivo mais rendeu medalhas de ouro para o Brasil? E total de medalhas?
# + colab={"base_uri": "https://localhost:8080/", "height": 748} id="dzo3bcZfUmaw" outputId="0bfe91b8-74e7-4960-bf13-a29303cb783f"
medalhas_sport_ouro_brasil.groupby(["Event"]).count().sort_values(by="Medal", ascending= False).filter(items= ["Medal"]).head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ylRsuwlB6lZU" outputId="af50d966-48ba-4f9b-8dea-1bf75c241926"
medalhas_evento_brasil.groupby(["Event"]).count().sort_values(by="Medal", ascending= False).filter(items= ["Medal"]).head(5)
# + [markdown] id="bktD1nmTUmax"
# Para finalizar sobre o Brasil: obtenha o total de medalhas de ouro, prata, bronze e total por ano.
# + id="mY6bKCInUmax"
colunas_selecionadas4=["Year","Medal"]
contador_medalhas= medalhistas_brasileiros.filter(items=colunas_selecionadas4)
# + colab={"base_uri": "https://localhost:8080/", "height": 657} id="ihbydDWJ7nfp" outputId="dbf66c0d-4041-40dd-c210-1e92abe167d8"
contador_medalhas.groupby(["Year"]).count().head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="YS_m1BwI8-_g" outputId="b54344a7-aaec-4bd1-a6f3-ba2de47a56c8"
medalha_ouro_contador= contador_medalhas[contador_medalhas["Medal"]=="Gold"].groupby(["Year"]).count()
medalha_ouro_contador.rename(columns={"Medal":"Medalhas de Ouro"}).head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 414} id="VTCd6BGt_Ioa" outputId="1161098a-7ac9-4e46-95ab-38f497cf2d89"
medalha_prata_contador= contador_medalhas[contador_medalhas["Medal"]=="Silver"].groupby(["Year"]).count()
medalha_prata_contador.rename(columns={"Medal":"Medalhas de Prata"}).head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 596} id="4rSpdJCf_oO-" outputId="61558012-6e7b-45f9-8cb6-990885038185"
medalha_bronze_contador= contador_medalhas[contador_medalhas["Medal"]=="Bronze"].groupby(["Year"]).count()
medalha_bronze_contador.rename(columns={"Medal":"Medalhas de Bronze"}).head(5)
# + [markdown] id="GIqomkqOUmay"
# ## 2. O mundo nos jogos de verão
#
# Vamos agora analisar um pouquinho do que aconteceu nas Olimpíadas de verão em todo o mundo.
#
# Retome o DataFrame original e descarte as informações sobre os jogos de inverno.
# + id="L1rBFAIdUmay"
atletas.head(1)
# -
jogos_verao = atletas[atletas["Season"] != "Winter"]
jogos_verao.head(5)
# Verificando se a exclusão foi feita na coluna 'Season'
jogos_verao["Season"].value_counts()
# Verificando se a exclusão foi feita na coluna 'Games'
jogos_verao["Games"].value_counts().head(5)
# + [markdown] id="VXvXFPdvUmay"
# Obtenha a lista de todos os esportes já disputados nas olimpíadas de verão.
# + id="5UOXHoeSUmaz"
pd.DataFrame(jogos_verao["Sport"]).head(5)
# + [markdown] id="20lOsDHbUmaz"
# Obtenha a lista de todas as modalidades esportivas já disputadas nas olimpíadas de verão.
# + id="PpaeLIcFUma0"
pd.DataFrame(jogos_verao["Event"]).head(5)
# + [markdown] id="VUmGHSDLUma1"
# Obtenha a lista de todos os países que já disputaram olimpíadas.
# -
# Excluí da coluna 'Team' os valores duplicados
valores_nao_duplicados = jogos_verao.drop_duplicates(subset=["Team"])
valores_nao_duplicados.head(5)
lista_paises = valores_nao_duplicados["Team"]
lista_paises.head(10)
# + [markdown] id="tb4onXZ9Uma4"
# Qual atleta foi o maior medalhista (em medalhas totais) da história das olimpíadas de verão?
# -
atletas.groupby(["Name"]).count().sort_values(by="Medal", ascending= False).filter(items= ["Medal"]).head(5)
# + [markdown] id="0le8l2KrUma6"
# Qual atleta foi o maior medalhista de ouro da história das olimpíadas de verão?
# + id="KxDErEGRUma7"
grupo_de_atletas = atletas[atletas['Medal']=='Gold'].copy()
maior_medalhista_historia = grupo_de_atletas.groupby(['Name']).count().sort_values(by='Medal', ascending = False)
maior_medalhista_historia.head(5)
# + [markdown] id="0dbGfkcmUma8"
# Qual país foi o maior medalhista de ouro da história das olimpíadas de verão? Lembre-se da questão do evento esportivo, para não considerar múltiplas medalhas para um mesmo evento (ex: uma equipe de futebol fazendo parecer que mais de 20 medalhas foram distribuídas).
# + id="3-Gq62XdUma8"
grupo_de_atletas = atletas[atletas['Medal']=='Gold'].copy()
filtros = ["Team", "Season", "Medal", "Event"]
grupo_de_atletas.drop_duplicates(subset=filtros, inplace=True)
maior_pais_ouro = grupo_de_atletas.groupby(['Team']).count().sort_values(by='Medal', ascending = False)
maior_pais_ouro.head(5)
# + [markdown] id="SjtNfai7Uma9"
# Qual país foi o maior medalhista em medalhas totais na história das olimpíadas de verão?
# + id="FT2y0rY3Uma9"
grupo_de_atletas = atletas.copy()
filtros = ["Team", "Season", "Medal", "Event"]
grupo_de_atletas.drop_duplicates(subset=filtros, inplace=True)
maior_pais_total = grupo_de_atletas.groupby(['Team']).count().sort_values(by='Medal', ascending = False)
maior_pais_total.head(5)
# + [markdown] id="UI5N0bSJUma-"
# Obtenha o total de medalhas de ouro, prata e total por edição das Olimpíadas de verão. Lembre-se da questão do evento esportivo.
# + id="FdbPqttRUmbA"
jogos_verao_filtrado = jogos_verao.filter(items = ["Medal", "Games", "Team", "Event"])
jogos_verao_filtrado.drop_duplicates(inplace=True)
jogos_verao_filtrado.dropna(inplace=True)
outro_prata = jogos_verao_filtrado[jogos_verao_filtrado["Medal"] != "Bronze"]
pd.DataFrame(outro_prata.groupby(by=["Games"])["Medal"].value_counts()).head(5)
# -
jogos_verao_filtrado = jogos_verao.filter(items = ["Medal", "Games", "Team", "Event"])
jogos_verao_filtrado.drop_duplicates(inplace=True)
jogos_verao_filtrado.dropna(inplace=True)
outro_prata_total = jogos_verao_filtrado
pd.DataFrame(outro_prata_total.groupby(by=["Games"]).count()).head(5)
# + [markdown] id="JKwsl8esUmbA"
# ## 3. Brasil vs Mundo
# + [markdown] id="2tznG_rYUmbB"
# Para finalizar, vamos fazer algumas comparações entre Brasil e mundo. Qual o ranking do Brasil em cada edição das olimpíadas? Lembrando que o ranking é ordenado por medalhas de ouro.
# + id="dVyAh7gFUmbB"
# + [markdown] id="y31oeGAVUmbB"
# Compare o maior medalhista em ouros do Brasil com o maior medalhista em ouros do mundo.
# + id="md7_qeOFUmbC"
# MAIOR MEDALHISTA DE OURO DO BRASIL
maior_medalhista_ouro_brasil = atletas[(atletas["Team"] == "Brazil") & (atletas["Medal"] == "Gold")]
maior_medalhista_ouro_brasil = maior_medalhista_ouro_brasil.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_maior_medalhista_ouro_brasil = pd.DataFrame(maior_medalhista_ouro_brasil).sort_values(by=['Medal'], ascending=False).head(1)
df_maior_medalhista_ouro_brasil
# -
# MAIOR MEDALHISTA DE OURO DO MUNDO
maior_medalhista_ouro_mundo = atletas[atletas["Medal"] == "Gold"]
maior_medalhista_ouro_mundo = maior_medalhista_ouro_mundo.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_maior_medalhista_ouro_mundo = pd.DataFrame(maior_medalhista_ouro_mundo).sort_values(by=['Medal'], ascending=False).head(1)
df_maior_medalhista_ouro_mundo
# TABELA COMPARAÇÃO
df_tabela_comparacao_ouro = pd.concat([df_maior_medalhista_ouro_mundo, df_maior_medalhista_ouro_brasil], axis=1)
df_tabela_comparacao_ouro.head(50)
# Conferindo medalhas de Ouro de <NAME>, II - Dados diferentes do Google
df = atletas[(atletas["Name"] == "<NAME>, II") & (atletas["Medal"] == "Gold")].value_counts()
df = pd.DataFrame(df)
df.sum()
# + [markdown] id="GHMsTYE_UmbC"
# Compare o maior medalhista em total de medalhas do Brasil com o maior medalhista em total de medalhas do mundo.
# + id="UghOEEVqUmbD"
# MAIOR MEDALHISTA DO BRASIL NO TOTAL DE MEDALHAS
total_medalhas_brasil = atletas[atletas["Team"] == "Brazil"]
total_medalhas_brasil = total_medalhas_brasil.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_total_medalhas_brasil = pd.DataFrame(total_medalhas_brasil).sort_values(by=['Medal'], ascending=False).head(1)
df_total_medalhas_brasil
# -
# MAIOR MEDALHISTA DO MUNDO NO TOTAL DE MEDALHAS
total_medalhas_mundo = atletas.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_total_medalhas_mundo = pd.DataFrame(total_medalhas_mundo).sort_values(by=['Medal'], ascending=False).head(1)
df_total_medalhas_mundo
# TABELA COMPARAÇÃO
tabela_comparacao_mundo = pd.concat([df_total_medalhas_mundo, df_total_medalhas_brasil], axis=1 )
tabela_comparacao_mundo
# + [markdown] id="fqNhge2NUmbD"
# Compare o maior medalhista em ouros do Brasil com o maior medalhista do mundo no mesmo esporte.
# + id="FVMFIcL7UmbE"
# MAIOR MEDALHISTA DE OURO DO BRASIL
maior_medalhista_ouro_brasil = atletas[(atletas["Team"] == "Brazil") & (atletas["Medal"] == "Gold")]
maior_medalhista_ouro_brasil = maior_medalhista_ouro_brasil.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_maior_medalhista_ouro_brasil = pd.DataFrame(maior_medalhista_ouro_brasil).sort_values(by=['Medal'], ascending=False).head(1)
df_maior_medalhista_ouro_brasil
# -
# MAIOR MEDALHISTA DE OURO DO MUNDO NO MESMO ESPORTE
maior_medalhista_ouro_mundo_esporte = atletas[(atletas["Medal"] == "Gold") & (atletas["Sport"] == "Sailing")]
maior_medalhista_ouro_mundo_esporte = maior_medalhista_ouro_mundo_esporte.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_maior_medalhista_ouro_mundo_esporte = pd.DataFrame(maior_medalhista_ouro_mundo_esporte).sort_values(by=['Medal'], ascending=False).head(1)
df_maior_medalhista_ouro_mundo_esporte
# TABELA COMPARAÇÃO
tabela_comparacao_medalhas_ouro = pd.concat([df_maior_medalhista_ouro_mundo_esporte, df_maior_medalhista_ouro_brasil], axis=1)
tabela_comparacao_medalhas_ouro
# + [markdown] id="rJ02_LOwUmbE"
# Compare o maior medalhista em total de medalhas do Brasil com o maior medalhista do mundo no mesmo esporte.
# + id="ltgUuI6vUmbF"
# MAIOR MEDALHISTA DO BRASIL NO TOTAL DE MEDALHAS
total_medalhas_brasil = atletas[atletas["Team"] == "Brazil"]
total_medalhas_brasil = total_medalhas_brasil.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_total_medalhas_brasil = pd.DataFrame(total_medalhas_brasil).sort_values(by=['Medal'], ascending=False).head(1)
df_total_medalhas_brasil
# -
# MAIOR MEDALHISTA DO MUNDO NO TOTAL DE MEDALHAS NO MESMO ESPORTE
total_medalhas_mundo_mesmoEsporte = atletas[atletas["Sport"] == "Sailing"]
total_medalhas_mundo_mesmoEsporte = total_medalhas_mundo_mesmoEsporte.groupby(by=["Name", "Height", "Weight", "Sport"])["Medal"].count()
df_total_medalhas_mundo_mesmoEsporte = pd.DataFrame(total_medalhas_mundo_mesmoEsporte).sort_values(by=['Medal'], ascending=False).head(1)
df_total_medalhas_mundo_mesmoEsporte
# TABELA COMPARAÇÃO
tabela_comparacao_medalhas_ouro_total = pd.concat([df_total_medalhas_mundo_mesmoEsporte, df_total_medalhas_brasil], axis=1)
tabela_comparacao_medalhas_ouro_total
# + [markdown] id="pvh2BUuBUmbF"
# Calcule o percentual de medalhas de ouro, prata e bronze que o Brasil ganhou em cada olimpíada.
# -
percentual_medalhas_bronze_brasil = atletas[(atletas["Team"] == "Brazil") & (atletas["Medal"] == "Bronze")]
percentual_medalhas_bronze_brasil = percentual_medalhas_bronze_brasil.groupby(by=["Medal"])["Medal"].count().sum()
percentual_bronze = (percentual_medalhas_bronze_brasil/total_medalhas) * 100
print(f"{percentual_medalhas_bronze_brasil} medalhas de bronze, total de {percentual_bronze:.2f} % ")
percentual_medalhas_prata_brasil = atletas[(atletas["Team"] == "Brazil") & (atletas["Medal"] == "Silver")]
percentual_medalhas_prata_brasil = percentual_medalhas_prata_brasil.groupby(by=["Medal"])["Medal"].count().sum()
percentual_prata = (percentual_medalhas_prata_brasil/total_medalhas) * 100
print(f"{percentual_medalhas_prata_brasil} medalhas de prata, total de {percentual_prata:.2f} % ")
percentual_medalhas_ouro_brasil = atletas[(atletas["Team"] == "Brazil") & (atletas["Medal"] == "Gold")]
percentual_medalhas_ouro_brasil = percentual_medalhas_ouro_brasil.groupby(by=["Medal"])["Medal"].count().sum()
percentual_ouro = (percentual_medalhas_ouro_brasil/total_medalhas) * 100
print(f"{percentual_medalhas_ouro_brasil} medalhas de ouro, total de {percentual_ouro:.2f} % ")
total_medalhas = percentual_medalhas_bronze_brasil + percentual_medalhas_prata_brasil + percentual_medalhas_ouro_brasil
print(f"Temos um total de {total_medalhas} medalhas")
# TABELA FINAL
pd.DataFrame(data = [percentual_ouro, percentual_prata, percentual_bronze], index=["Ouro", "Prata", "bronze"], columns=["Porcentagem"])
| Projeto_Olimpiadas_18_10_2021 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7UpxgYfOmzh9"
# # Solution: Convolution and Pooling
#
# [](https://colab.research.google.com/github/ShawnHymel/computer-vision-with-embedded-machine-learning/blob/master/2.1.4%20-%20Project%20-%20Convolution%20and%20Pooling/solution_convolution_and_pooling.ipynb)
#
# Create a convolution filter to filter an image. Then, create a pooling function.
#
# Run each of the cells paying attention to their contents and output. Fill out the necessary parts of the functions where you find the following comment:
#
# ```
# # >>> ENTER YOUR CODE HERE <<<
# ```
#
# Note that you may not use libraries other than those listed in the first cell to perform convolution and pooling.
#
# You can see what the output images are supposed to look like [here](https://github.com/ShawnHymel/computer-vision-with-embedded-machine-learning/raw/master/2.1.4%20-%20Project%20-%20Convolution%20and%20Pooling/output-images.png).
#
# If you get stuck or would like to compare answers, you can find [my solution here](https://colab.research.google.com/github/ShawnHymel/computer-vision-with-embedded-machine-learning/blob/master/2.1.4%20-%20Project%20-%20Convolution%20and%20Pooling/solution_convolution_and_pooling.ipynb).
#
# Author: EdgeImpulse, Inc.<br>
# Date: August 2, 2021<br>
# License: [Apache-2.0](apache.org/licenses/LICENSE-2.0)<br>
# + id="n2_hUewhmyTm"
### Import libraries
import os
import PIL
import requests
import math
import numpy as np
import matplotlib.pyplot as plt
# + id="rnfkQvNYxHCM"
### Download example image
# Image location and path
url = "https://github.com/ShawnHymel/computer-vision-with-embedded-machine-learning/raw/master/2.1.4%20-%20Project%20-%20Convolution%20and%20Pooling/resistor.png"
img_path = os.path.join("/content", "resistor.png")
# Download image
resp = requests.get(url)
# Write image to file
with open(img_path, 'wb') as f:
f.write(resp.content)
# + id="dBV7wfOky1XY"
### Open and view image
# Use PIL to open the image and convert it to grayscale
img = PIL.Image.open(img_path)
img = img.convert('L')
# Convert image to Numpy array
img = np.asarray(img)
# Show dimensions and view array as image
print(img.shape)
plt.imshow(img, cmap='gray', vmin=0, vmax=255)
# + [markdown] id="s97lc17wzo0G"
# ## Part 1: Convolution
#
# Your assignment is to write a function that convolves the image array (stored in the `img` variable) with a given kernel and stride. Assume valid padding (e.g. no padding). Note that the kernel will be given as a 2D Numpy array.
#
# You are welcome to use for loops in this exercise, as it helps to see what's going on in the convolution operation. Bonus points if you do it without for loops (caveat: bonus points don't mean anything in this course).
#
# You may find some of the following Numpy operations to be helpful:
#
# * [numpy.arange()](https://numpy.org/doc/stable/reference/generated/numpy.arange.html)
#
# + id="0FDjgm5pzZOg"
### Convolution function
def convolve(img, kernel, stride):
# Compute dimensions of output image
out_height = math.floor((img.shape[0] - kernel.shape[0]) / stride) + 1
out_width = math.floor((img.shape[1] - kernel.shape[1]) / stride) + 1
# Create blank output image
convolved_img = np.zeros((out_height, out_width))
# >>> ENTER YOUR CODE HERE <<<
# Loop through each pixel in the output array. Note that this is not the most efficient way of
# doing convolution, but it provides some insights into what's going on.
for i in np.arange(0, out_height):
for j in np.arange(0, out_width):
# Set a temporary variable to 0
accumulator = 0
# Do element-wise multiplication and sum the result over the window/kernel
for m in np.arange(0, kernel.shape[0]):
for n in np.arange(0, kernel.shape[1]):
accumulator += img[(stride * i) + m, (stride * j) + n] * kernel[m, n]
# Set output image pixel to accumulator value
convolved_img[i, j] = accumulator
# Round all elements, convert to integers, and clamp to values between 0 and 255
convolved_img = np.rint(convolved_img).astype(int)
convolved_img = np.clip(convolved_img, 0, 255)
return convolved_img
# + id="IeLJ2y9h0bVH"
### Test 1: Gaussian blur filter
# Define kernel
kernel = np.array([[1/16, 2/16, 1/16],
[2/16, 4/16, 2/16],
[1/16, 2/16, 1/16]])
# Call your convolve function (with a stride of 1)
out_img = convolve(img, kernel, 1)
# Show dimensions and view array as image
print(out_img.shape)
plt.imshow(out_img, cmap='gray', vmin=0, vmax=255)
# + id="OU_dTpq229_D"
### Test 2: Edge detection
# Define kernel
kernel = np.array([[-1, -1, -1],
[-1, 8, -1],
[-1, -1, -1]])
# Call your convolve function (with a stride of 1)
out_img = convolve(img, kernel, 1)
# Show dimensions and view array as image
print(out_img.shape)
plt.imshow(out_img, cmap='gray', vmin=0, vmax=255)
# + id="NJcZaIPs3Ksh"
### Test 3: Sharpen with stride > 1
# Define kernel
kernel = np.array([[0, -1, 0],
[-1, 5, -1],
[0, -1, 0]])
# Call your convolve function (with a stride of 2)
out_img = convolve(img, kernel, 2)
# Show dimensions and view array as image
print(out_img.shape)
plt.imshow(out_img, cmap='gray', vmin=0, vmax=255)
# + [markdown] id="MHAcZ4qK5IAK"
# ## Part 2: Pooling
#
# Your assignment is to write a function that performs max pooling on the input image array (stored in the `img` variable). Assume valid padding (e.g. no padding). Also, assume that the pool width determines the horizontal stride and the pool height determines the vertical stride.
#
# You are welcome to use for loops in this exercise, as it helps to see what's going on in the convolution operation. Bonus points if you do it without for loops (caveat: bonus points don't mean anything in this course).
#
# You may find some of the following Numpy operations to be helpful:
#
# * [numpy.arange()](https://numpy.org/doc/stable/reference/generated/numpy.arange.html)
# + id="yBeCY9Bd4o_b"
### Max pooling function
def maxpooling(img, pool_height, pool_width):
# Set stride amounts
stride_y = pool_height
stride_x = pool_width
# Compute dimensions of output image
out_height = math.floor((img.shape[0] - pool_height) / stride_y) + 1
out_width = math.floor((img.shape[1] - pool_width) / stride_x) + 1
# Create blank output image
pooled_img = np.zeros((out_height, out_width))
# >>> ENTER YOUR CODE HERE <<<
# Loop through each pixel in the output array. Note that this is not the most efficient way of
# doing convolution, but it provides some insights into what's going on.
for i in np.arange(0, out_height):
for j in np.arange(0, out_width):
# Set output to the value of the first element in the window
out_val = img[(stride_y * i), (stride_x * j)]
# Look through each element in the window to find the max value
for m in np.arange(0, pool_height):
for n in np.arange(0, pool_width):
out_val = max(out_val, img[(stride_y * i) + m, (stride_x * j) + n])
# Set element in output array to max value
pooled_img[i, j] = out_val
# Round all elements, convert to integers, and clamp to values between 0 and 255
pooled_img = np.rint(pooled_img).astype(int)
pooled_img = np.clip(pooled_img, 0, 255)
return pooled_img
# + id="4M-Q5Wzl_LK4"
### Test 1: Max pool original image with pool size of (2, 3)
# Call your pooling function (with pool_size=(2, 3))
out_img = maxpooling(img, 2, 3)
# Show dimensions and view array as image
print(out_img.shape)
plt.imshow(out_img, cmap='gray', vmin=0, vmax=255)
# + id="-Wm4R5MS_obx"
### Test 2: Detect edges and pool
# Define kernel
kernel = np.array([[-1, -1, -1],
[-1, 8, -1],
[-1, -1, -1]])
# Call your convolve function (with a stride of 1)
convolved_img = convolve(img, kernel, 1)
# Call your pooling function (with pool_size=(2, 2))
out_img = maxpooling(convolved_img, 2, 2)
# Show dimensions and view array as image
print(out_img.shape)
plt.imshow(out_img, cmap='gray', vmin=0, vmax=255)
# + id="k2rg60rGAXOS"
| 2.1.4 - Project - Convolution and Pooling/solution_convolution_and_pooling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split, GridSearchCV, StratifiedKFold, \
learning_curve, validation_curve
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, confusion_matrix, precision_score, recall_score
from sklearn.decomposition import PCA
from scipy.stats import mannwhitneyu
sns.set(font_scale=1.5)
pd.options.display.max_columns = 50
# -
# ## Project plan
# * 1. Feature and data explanation
# * 2. Primary data analysis
# * 3. Primary visual data analysis
# * 4. Insights and found dependencies
# * 5. Metrics selection
# * 6. Model selection
# * 7. Data preprocessing
# * 8. Cross-validation and adjustment of model hyperparameters
# * 9. Creation of new features and description of this process
# * 10. Plotting training and validation curves
# * 11. Prediction for test or hold-out samples
# * 12. Conclusions
# ## 1. Feature and data explanation
df = pd.read_csv('data.csv')
df.head()
# ### 1.1 Process of collecting data
#
#
# Features are computed from a digitized image of a fine needle aspirate (FNA) of a breast mass. They describe characteristics of the cell nuclei present in the image. n the 3-dimensional space is that described in: [<NAME> and <NAME>: "Robust Linear Programming Discrimination of Two Linearly Inseparable Sets", Optimization Methods and Software 1, 1992, 23-34].
#
#
# The data used is available through https://www.kaggle.com/uciml/breast-cancer-wisconsin-data
# And can be found on UCI Machine Learning Repository: https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+%28Diagnostic%29
# This database is also available through the UW CS ftp server: ftp ftp.cs.wisc.edu cd math-prog/cpo-dataset/machine-learn/WDBC/
# ### 1.2 Detailed explanation of the task
# The task here is to predict whether the cancer is benign or malignant based in 30 real-valued features.
# ### 1.3 Target features
# Attribute Information:
#
# 1) ID number
# 2) Diagnosis (M = malignant, B = benign)
# 3-32)
#
# Ten real-valued features are computed for each cell nucleus:
# a) radius (mean of distances from center to points on the perimeter)
# b) texture (standard deviation of gray-scale values)
# c) perimeter
# d) area
# e) smoothness (local variation in radius lengths)
# f) compactness (perimeter^2 / area - 1.0)
# g) concavity (severity of concave portions of the contour)
# h) concave points (number of concave portions of the contour)
# i) symmetry
# j) fractal dimension ("coastline approximation" - 1)
#
# The mean, standard error and "worst" or largest (mean of the three largest values) of these features were computed for each image, resulting in 30 features. For instance, field 3 is Mean Radius, field 13 is Radius SE, field 23 is Worst Radius.
#
#
# All feature values are recoded with four significant digits.
# Missing attribute values: none
# Class distribution: 357 benign, 212 malignant
# ## 2. Primary data analysis
# ### 2.0 Data preprocessing
target = pd.DataFrame(df['diagnosis'])
data = df.drop(['diagnosis'], axis=1)
# ### 2.1 Constant columns
# General data overview:
data.info()
# Drop constant column ** Unnamed: 32 ** and **id** column which is useless for analize:
data.drop(['Unnamed: 32', 'id'], axis=1, inplace=True)
# ### 2.2 Missing values
# Check data for missing values:
print("Are there missing values:", data.isnull().values.any())
# ### 2.3 Summary statistics
# General data statistics overview:
data.describe()
# **Conclusion:** here we can see vary different min/max values for features, for example *area_mean* and *smoothness_mean*. Thus we should check for outliers (box plot is good option for that).
# ### 2.4 Statistics for different classes
# Check if difference of features mean values is statistically important. We will use Mann Whitney criteria, because of it unsesetive to outliers and different samples distribution.
for column in data.columns:
m = data[column][target['diagnosis']=='M']
b = data[column][target['diagnosis']=='B']
statistic, pvalue = mannwhitneyu(m, b)
print('Column:', column, 'Important:', pvalue < 0.05 )
# **Conclusion:** differences in almost all features are statistically important. So they will contribute more enough information to classification.
# ### 2.5 Target feature
# Number of eamples for each class:
target['diagnosis'].value_counts()
# Let's check the ratio of examples belong to each class:
target['diagnosis'].value_counts() / target['diagnosis'].size
# **Conclusion:** there are a lot more examples for benign class, but not enough for skewed classes problem.
# ## 3. Primary visual data analysis
# For the sake of informative data visualization we need to standardize and scale features, because of some features have very different max/min values.
scaler = StandardScaler()
scaled_data = scaler.fit_transform(data)
data_scaled = pd.DataFrame(scaled_data, columns=data.columns)
data_scaled['diagnosis'] = target['diagnosis']
# ### 3.1 Linear dependecies of the features (correlation matrix):
# Helper function for plotting feature correlations:
def plot_corr(data):
plt.figure(figsize=[40, 40])
ax = sns.heatmap(data.corr(), annot=True, fmt= '.1f', linewidths=.5)
ax.set_xticklabels(ax.get_xticklabels(), size='xx-large')
ax.set_yticklabels(ax.get_yticklabels(), size='xx-large')
plt.show();
# Data correlations:
plot_corr(data)
# **Conclusion:** there are several groups of correlated features:
# - radius_mean, perimeter_mean, area_mean
# - compactness_mean, concavity_mean, concave points_mean
# - radius_se, perimeter_se, area_se
# - radius_worst, perimeter_worst and area_worst
# - compactness_worst, concavity_worst, concave points_worst
# - compactness_se, concavity_se, concave points_se
# - texture_mean, texture_worst
# - area_worst, area_mean
# ### 3.2 Outliers
data_z = pd.melt(data_scaled, id_vars="diagnosis", var_name="features", value_name='value')
plt.figure(figsize=(20, 10));
ax = sns.boxplot(x='features', y='value', hue='diagnosis', data=data_z);
ax.set_xticklabels(ax.get_xticklabels());
plt.xticks(rotation=90);
# **Conclusion:** there are a lot of variable with outliers. So before training we have to handle it.
# ### 3.3 Distribution of classes
plt.figure(figsize=(30, 20));
ax = sns.violinplot(x="features", y="value", hue="diagnosis", data=data_z, split=True, inner="quartile");
ax.set_xticklabels(ax.get_xticklabels(), size='large');
plt.xticks(rotation=90);
# **Conclusion:** in some features, like *radius_mean*, *texture_mean*, median of each class separated, so they can be useful for classification. Other features, like *smoothness_se*, are not so separated and my be less useful for classification. Most all the features have normal-like distribution with long tail.
# ### 3.4 Dimensionality reduction
# Apply pca for dimensionality reduction:
# +
pca = PCA(random_state=24)
pca.fit(scaled_data)
plt.figure(figsize=(10, 10))
plt.plot(pca.explained_variance_ratio_, linewidth=2)
plt.xlabel('Number of components');
plt.ylabel('Explained variance ratio');
# -
# **Conclusion:** according to elbow method 3 components may be choosen.
# Check the number of components for explaining data variance:
components = range(1, pca.n_components_ + 1)
plt.figure(figsize=(15, 5));
plt.bar(components, np.cumsum(pca.explained_variance_ratio_));
plt.hlines(y = .95, xmin=0, xmax=len(components), colors='green');
# **Conclusion:** The two first components explains the 0.6324 of the variance. We need 10 principal components to explain more than 0.95 of the variance and 17 to explain more than 0.99.
# Reduce dimensions of data and plot it:
pca_two_comp = PCA(n_components=2, random_state=24)
two_comp_data = pca_two_comp.fit_transform(scaled_data)
plt.scatter(x=two_comp_data[:, 0], y=two_comp_data[:, 1],
c=target['diagnosis'].map({'M': 'red', 'B': 'green'}))
plt.show()
# **Conclusion:** data is good enough separable using only two components.
# ## 4. Insights and found dependencies
# Data summary:
# - there are a lot of groups with correlated features. Next we have to get rid from multi-collinearity by selectig one feature for each group.
# - ration of examples in each class 0.67/0.27. No skewed classes here, which is important for metric selection;
# - differences in features stitistics (mean) for each class are statisticalli important. So this features will be important for classification.
# - there are outliers in data. It's important to get rid of them for outliers-sensetive models (logistic regression for example) before training;
# - PCA shows thad data is good enough separable using only 3-5 features.
# ## 5. Metrics selection
# Predict whether the cancer is benign or malignant is a **binary classification** task. Here we don't face the probem of skewed classes. So **accuracy** metric will be a good choice for model evaluation. Also this metric is simple enough, thus highly interpretable.
# $$Accuracy=\frac{Number~of~corrected~predictions}{Total~number~of~predictions}$$
# Aslo for the test set we will calculate **precision** and **recall**.
# ## 6. Model selection
# As model was selected **Logistic regression** because:
# - works well with non categorical features (in our data all features are continious);
# - robust to small noise in the data;
# - cases of multi-collinearity can be handled by implementing regularization;
# - works well if there are no missing data;
# - efficient implementation availavle;
# - feature space of current task is not large.
# ## 7. Data preprocessing
# ### 7.1 Drop useless columns
# Drop constant column **Unnamed: 32** and useless folumn **id** for classification.
X = df.drop(['id', 'Unnamed: 32', 'diagnosis'], axis=1)
y = df['diagnosis'].map(lambda x: 1 if x=='M' else 0)
# ### 7.3 Split data into train/test
#
# Split data into train/test with proportional 0.7/0.3 which is common split for such amount of data.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=24)
print('Train size:', X_train.size)
print('Test size:', X_test.size)
# ### 7.2 Feature selection
# First of all we should handle multi-collinearity. From each group of correleted features we will select only by one feature. So here columns to drop:
corr_columns = ['perimeter_mean','radius_mean','compactness_mean',
'concave points_mean','radius_se','perimeter_se',
'radius_worst','perimeter_worst','compactness_worst',
'concave points_worst','compactness_se','concave points_se',
'texture_worst','area_worst',
'concavity_mean']
# Drop correlated columns from train data:
X_train = X_train.drop(corr_columns, axis=1)
# Drop correlated columns from test data:
X_test = X_test.drop(corr_columns, axis=1)
# Check number of features left:
print('Current number of features:', X_train.shape[1])
# ### 7.3 Feature scaling
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
# ## 8. Cross-validation and adjustment of model hyperparameters
# Use 3 splits because of we don't have large amount of training data and shuffle samples in random order.
cv = StratifiedKFold(n_splits=3, random_state=24)
# Model:
model = LogisticRegression(random_state=24)
# Model parameters:
model_parameters = {'penalty': ['l1', 'l2'],
'C': np.linspace(.1, 1, 10)}
# To find best hyperparameters we will use grid search as in is quite simple and efficient enough.
grig_search = GridSearchCV(model, model_parameters, n_jobs=-1, cv=cv, scoring='accuracy')
# %%time
grig_search.fit(X_train_scaled, y_train);
# Best model parameters:
grig_search.best_params_
# Best cv score:
print('Accuracy:', grig_search.best_score_)
# ## 9. Creation of new features
# Helper function for applying map operation to data frame attributes:
def apply_cat_op(data, attrs, operation, prefix):
"""
Apply one operation to data attributes.
"""
series = [data[attr].map(operation) for attr in attrs]
_data = pd.concat(series, axis=1).add_prefix(prefix)
new_attrs = _data.columns.values
return _data, new_attrs
# Creating new features based on medicine requires strong domain knowledge. So we will create them based on mathematics nature of current features. Basic approach for numerical features for regression model is to calculate squares of features in order to capture non-linear dependencies.
# Square function:
sq_operation = lambda x: x**2
# Create squared feature for each columns and test in with model:
for column in X_train.columns:
X_train_sq, sq_attr = apply_cat_op(X_train, [column], sq_operation, 'sq_')
data = pd.concat([X_train, X_train_sq], axis=1)
scaler = StandardScaler()
data_scaled = scaler.fit_transform(data)
grig_search = GridSearchCV(model, model_parameters, n_jobs=-1, cv=cv, scoring='accuracy')
grig_search.fit(data_scaled, y_train);
print('Column:', column, ' ',
'Accuracy:', grig_search.best_score_, ' ',
'Best params:', grig_search.best_params_)
# As we ca see squaring feature *fractal_dimension_mean*, gives score improving with params {'C': 0.2, 'penalty': 'l2'}
# Add new feature to train data:
X_train_sq, atr = apply_cat_op(X_train, ['fractal_dimension_mean'], sq_operation, 'sq_')
X_train = pd.concat([X_train, X_train_sq], axis=1)
# Add new feature to test data:
X_test_sq, atr = apply_cat_op(X_test, ['fractal_dimension_mean'], sq_operation, 'sq_')
X_test = pd.concat([X_test, X_test_sq], axis=1)
# #### Scale the final data:
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
# #### Train model with best parameters on all train data:
final_model = LogisticRegression(penalty='l2', C=0.2)
final_model.fit(X_train_scaled, y_train)
# ## 10. Plotting training and validation curves
# ### 10.1 Training curve
# Plotting [learning curve fuction](https://scikit-learn.org/stable/auto_examples/model_selection/plot_learning_curve.html#sphx-glr-auto-examples-model-selection-plot-learning-curve-py):
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None,
n_jobs=None, train_sizes=np.linspace(.1, 1.0, 5)):
"""
Generate a simple plot of the test and training learning curve.
Parameters
----------
estimator : object type that implements the "fit" and "predict" methods
An object of that type which is cloned for each validation.
title : string
Title for the chart.
X : array-like, shape (n_samples, n_features)
Training vector, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape (n_samples) or (n_samples, n_features), optional
Target relative to X for classification or regression;
None for unsupervised learning.
ylim : tuple, shape (ymin, ymax), optional
Defines minimum and maximum yvalues plotted.
cv : int, cross-validation generator or an iterable, optional
Determines the cross-validation splitting strategy.
Possible inputs for cv are:
- None, to use the default 3-fold cross-validation,
- integer, to specify the number of folds.
- :term:`CV splitter`,
- An iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if ``y`` is binary or multiclass,
:class:`StratifiedKFold` used. If the estimator is not a classifier
or if ``y`` is neither binary nor multiclass, :class:`KFold` is used.
Refer :ref:`User Guide <cross_validation>` for the various
cross-validators that can be used here.
n_jobs : int or None, optional (default=None)
Number of jobs to run in parallel.
``None`` means 1 unless in a :obj:`joblib.parallel_backend` context.
``-1`` means using all processors. See :term:`Glossary <n_jobs>`
for more details.
train_sizes : array-like, shape (n_ticks,), dtype float or int
Relative or absolute numbers of training examples that will be used to
generate the learning curve. If the dtype is float, it is regarded as a
fraction of the maximum size of the training set (that is determined
by the selected validation method), i.e. it has to be within (0, 1].
Otherwise it is interpreted as absolute sizes of the training sets.
Note that for classification the number of samples usually have to
be big enough to contain at least one sample from each class.
(default: np.linspace(0.1, 1.0, 5))
"""
plt.figure()
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
return plt
plot_learning_curve(final_model, 'Logistic regression',
X_train_scaled, y_train, cv=cv);
# **Conclusion:** such gap between training and validating curve indicates overfitting. But we can see that validation curve increasing with increasing amount of training examples, so more data is likely to help beat overfitting.
# ### 10.2 Validation curve
# Plotting validation curve function:
def plot_validation_curve(estimator, title, X, y, param_name, param_range,
cv=None, scoring=None, ylim=None, n_jobs=None):
"""
Generates a simple plot of training and validation scores for different parameter values.
Parameters
----------
estimator : object type that implements the "fit" and "predict" methods
An object of that type which is cloned for each validation.
title : string
Title for the chart.
X : array-like, shape (n_samples, n_features)
Training vector, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape (n_samples) or (n_samples, n_features), optional
Target relative to X for classification or regression;
None for unsupervised learning.
param_name : string
Name of the parameter that will be varied.
param_range : array-like, shape (n_values,)
The values of the parameter that will be evaluated.
cv : int, cross-validation generator or an iterable, optional
Determines the cross-validation splitting strategy.
Possible inputs for cv are:
- None, to use the default 3-fold cross-validation,
- integer, to specify the number of folds.
- :term:`CV splitter`,
- An iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if ``y`` is binary or multiclass,
:class:`StratifiedKFold` used. If the estimator is not a classifier
or if ``y`` is neither binary nor multiclass, :class:`KFold` is used.
Refer :ref:`User Guide <cross_validation>` for the various
cross-validators that can be used here.
scoring : string, callable or None, optional, default: None
A string (see model evaluation documentation) or
a scorer callable object / function with signature
``scorer(estimator, X, y)``.
ylim : tuple, shape (ymin, ymax), optional
Defines minimum and maximum yvalues plotted.
n_jobs : int or None, optional (default=None)
Number of jobs to run in parallel.
``None`` means 1 unless in a :obj:`joblib.parallel_backend` context.
``-1`` means using all processors. See :term:`Glossary <n_jobs>`
for more details.
"""
train_scores, test_scores = validation_curve(
estimator, X, y, param_name, param_range,
cv=cv, scoring=scoring, n_jobs=n_jobs)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.figure()
plt.grid()
plt.title(title)
plt.xlabel(param_name)
plt.ylabel("Score")
if ylim is not None:
plt.ylim(*ylim)
plt.semilogx(param_range, train_scores_mean, 'o-', label="Training score",
color="darkorange")
plt.fill_between(param_range, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.2,
color="darkorange")
plt.semilogx(param_range, test_scores_mean, 'o-', label="Cross-validation score",
color="navy")
plt.fill_between(param_range, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.2,
color="navy")
plt.legend(loc="best")
return plt
# Plot validation curve for model complexity parameter:
plot_validation_curve(final_model, 'Logistic regression', X_train_scaled, y_train,
'C', model_parameters['C'],
cv=cv, scoring='accuracy');
# **Conclusion:** gap between training and validating curve indicates overfitting. The best **C** parameter is 0.2
# ## 11. Prediction for test samples
# Make predictions for test samples:
test_predictions = final_model.predict(X_test_scaled)
# #### Accuracy score:
print('Accuracy test score:', accuracy_score(y_test, test_predictions))
# **Conclusion:** result on the test samples are comparable to the results on cross-validation, even better. Thus our validation scheme is valid.
# #### Confusion matrix:
test_confusion_matrix = confusion_matrix(test_predictions, y_test);
sns.heatmap(test_confusion_matrix, annot=True, fmt='d');
# From confusion matrix we can see that we have made a few wrong predicions.
# #### Precision:
print('Precision:', precision_score(y_test, test_predictions))
# #### Recall:
print('Recall:', recall_score(y_test, test_predictions))
# ## 12. Conclusions
# Although we try simple model, it gives 98% accuracy, 98% precision and 97% recall on the test set. There are several (3-5) most important features for classification, which could indicates that our data is not representable or biased. So, it's a good option to try model on more data. Feature generation based on medicine knowledge for such data is quite challenging, so we build them based on math nature.
# #### Ways of improving:
# - collect more data and re-train model on it, as we can see validation score improvement with data amount increasing on learning curve;
# - dig into domain and generate more features based on medicine;
# - try another models, like neural network (for capturing complex non-linear dependences) or random forest (robust to overfitting);
# - apply PCA for data dimensionality reduction and train model on reduced data;
# - try stacking differet models.
| jupyter_english/projects_indiv/cancer_benign_or_malignant_nikolai_timonin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning used for Asset Allocation: Multi-task Lasso
#
# **<NAME>, CFA**<br>
# *AlphaWave Data*
#
# **September 2021**
# ## Introduction
#
# In this article, we use [machine learning](https://hdonnelly6.medium.com/list/machine-learning-for-investing-7f2690bb1826) to make future returns predictions for equity and fixed income ETFs so that we can create optimized Equity Only, Fixed Income Only, and 60/40 Allocation portfolios versus respective benchmarks. We show that we are able to outperform the benchmarks using the Multi-task Lasso model.
#
# Jupyter Notebooks are available on [Google Colab](https://colab.research.google.com/drive/1HraWoI6I6dHc7YEZuAOgZHMzNgwN8OMZ?usp=sharing) and [Github](https://github.com/AlphaWaveData/Jupyter-Notebooks/blob/master/AlphaWave%20Data%20Machine%20Learning%20used%20for%20Asset%20Allocation%20example.ipynb).
#
# For this project, we use several Python-based scientific computing technologies listed below.
# +
import time
import requests
import numpy as np
import pandas as pd
from tqdm import tqdm
from itertools import product
import plotly.graph_objects as go
from IPython.display import display
from datetime import datetime, timedelta
from scipy import stats
from sklearn.linear_model import MultiTaskLasso
from sklearn.utils.testing import ignore_warnings
from sklearn.exceptions import ConvergenceWarning
# -
# ## Asset Allocation
#
# Let's start with a quick overview of asset allocation. Asset owners are concerned with accumulating and maintaining the wealth needed to meet their needs and aspirations. In that endeavor, investment portfolios—including individuals’ portfolios and institutional funds—play important roles. Asset allocation is a strategic—and often a first or early—decision in portfolio construction. Because it holds that position, it is widely accepted as important and meriting careful attention.
#
# Generally, investment firms manage a group of portfolios and have particular outcomes or target dates assigned to each of these portfolios. To make sure these portfolios meet their assigned goals, there can be a strategic asset allocation associated with them. The strategic asset allocation decision determines return levels in which allocations are invested, irrespective of the degree of active management.
#
# A common example of a portfolio with a strategic asset allocation is a portfolio with defined weightings for equity and fixed income asset classes. The equity weighting may be allocated between U.S. equities and global equities while the fixed income weighting may be divided between Treasuries, corporate bonds, high-yield credit, and emerging market debt. These weightings are usually quite fixed and are based on a long time horizon of historical returns and correlations. There may be other asset classes also considered in a portfolio with a strategic asset allocation, like commodities and derivatives. The portfolios are typically rebalanced periodically in order to maintain the same asset class exposure going forward in time.
#
# However, we live in a world that changes quickly. As a result, some investment firms often employ a tactical asset allocation. This provides the portfolio with short-term tilts. There can be dynamic weightings, or put differently, migrations slightly away from the strategic asset allocation. The portfolio that uses a tactical asset allocation may focus on allocations within sub-asset classes. If you are working within the fixed income asset class, you may move your allocation away from Treasuries and short-term notes and towards a high-yield or emerging market credit allocation. The belief is that the portfolio is better able to outperform the market with these new asset allocations.
#
# To employ a tactical asset allocation, there are a few different approaches that can be taken. One of the simplest approaches is a discretionary one whereby portfolio managers and chief investment officers overweight or underweight particular assets within the portfolio based on their view of the business cycle. They often move within certain risk boundaries when changing portfolio weightings.
#
# Another tactical asset allocation technique used is a momentum, trend following approach. With this, you look to capitalize on an asset class outperforming its peers in the next couple of rebalancing periods.
#
# The Black-Litterman model is yet another tactical asset allocation approach. The model came out of Goldman Sachs research published in 1992 by <NAME> and <NAME>. This model allows you to look at the historical returns, risk, and correlation of the assets in your portfolio. The user is only required to state how her assumptions about expected returns differ from the markets and to state her degree of confidence in the alternative assumptions. From this, the Black–Litterman method computes the desired (mean-variance efficient) asset allocation.
#
# Made famous more recently by the likes of AQR, risk parity has become a popular tactical asset allocation technique. The risk parity approach asserts that when asset allocations are adjusted (leveraged or deleveraged) to the same risk level, the risk parity portfolio can achieve a higher Sharpe ratio and can be more resistant to market downturns than the traditional portfolio. This is a slightly different approach in that you almost forget about forward forecasting returns. Instead, you take the approach that you better understand where the risks lie in the portfolio and, as such, strive for an equal risk weighting among asset classes in the portfolio. Essentially, you are focused on allocation of risk, usually defined as volatility, rather than allocation of capital. For example, if you had a 60% allocation to equities and a 40% allocation to fixed income in your portfolio, risk parity will likely force you to increase your allocation to fixed income as it is likely to have less risk than equities. Risk parity is vulnerable to significant shifts in correlation regimes in practice, such as observed in Q1 2020, which led to the significant underperformance of risk-parity funds in the Covid-19 sell-off.
# ---
# ## Examine 60/40 Base Case Scenario
# Let's first get total returns for equity and fixed income ETFs that will serve as our benchmarks in this asset allocation analysis.
# +
# fetch daily return data for benchmarks: SPY and AGG ETFs
global_eq = 'SPY'
global_fi = 'AGG'
stock_tickers = [global_eq, global_fi]
# -
# We can use the [10 Year Historical Monthly Prices](https://rapidapi.com/alphawave/api/stock-prices2) endpoint from the [AlphaWave Data Stock Prices API](https://rapidapi.com/alphawave/api/stock-prices2/endpoints) to pull in the ten year monthly historical prices so that we can calculate the returns.
#
# To call this API with Python, you can choose one of the supported Python code snippets provided in the API console. The following is an example of how to invoke the API with Python Requests. You will need to insert your own <b>x-rapidapi-host</b> and <b>x-rapidapi-key</b> information in the code block below.
# +
# fetch 10 year monthly return data
url = "https://stock-prices2.p.rapidapi.com/api/v1/resources/stock-prices/10y-1mo-interval"
headers = {
'x-rapidapi-host': "YOUR_X-RAPIDAPI-HOST_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS",
'x-rapidapi-key': "YOUR_X-RAPIDAPI-KEY_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS"
}
stock_frames = []
# for ticker in stock_tickers:
for ticker in tqdm(stock_tickers, position=0, leave=True, desc = "Retrieving AlphaWave Data Benchmark Info"):
querystring = {"ticker":ticker}
stock_daily_price_response = requests.request("GET", url, headers=headers, params=querystring)
# Create Stock Prices DataFrame
stock_daily_price_df = pd.DataFrame.from_dict(stock_daily_price_response.json())
stock_daily_price_df = stock_daily_price_df.transpose()
stock_daily_price_df = stock_daily_price_df.rename(columns={'Close':ticker})
stock_daily_price_df = stock_daily_price_df[{ticker}]
stock_frames.append(stock_daily_price_df)
yf_combined_stock_price_df = pd.concat(stock_frames, axis=1, sort=True)
yf_combined_stock_price_df = yf_combined_stock_price_df.dropna(how='all')
yf_combined_stock_price_df = yf_combined_stock_price_df.fillna("")
periodic_returns = yf_combined_stock_price_df.pct_change().dropna()
periodic_returns
# -
# ### Add a 60/40 Equity/Fixed Income Allocation
# Let's also build a basic benchmark 60/40 Portfolio.
# create benchmark 60/40 Portfolio
periodic_returns['60/40 Portfolio'] = sum([periodic_returns[global_eq] * 0.6, periodic_returns[global_fi] * 0.4])
periodic_returns[["SPY",
"AGG",
"60/40 Portfolio"]] = periodic_returns[["SPY",
"AGG",
"60/40 Portfolio"]].apply(pd.to_numeric)
periodic_returns = periodic_returns.sort_index()
periodic_returns
# ### Plot the Cumulative Returns
#
# Next, we define `make_single_line_chart` and `make_all_line_charts` functions that will help us plot the benchmark returns.
# function to create a single line chart
def make_single_line_chart(column, alt_name=None):
data = cumulative_returns[[column]]
name = column
if alt_name is not None:
name = f'{alt_name} ({column})'
return go.Scatter(x=data.index, y=data[column], name=name)
# function to create a multi line chart
def make_all_line_charts(emphasize=None):
alt_names = {'SPY': '100% Equities', 'AGG': '100% Bonds'}
data = []
for column in cumulative_returns:
alt_name = None
if column in alt_names:
alt_name = alt_names[column]
chart = make_single_line_chart(column, alt_name)
if emphasize is not None:
if type(emphasize) != list:
emphasize = [emphasize]
if column not in emphasize:
chart.line.width = 1
chart.mode = 'lines'
else:
chart.line.width = 3
chart.mode = 'lines+markers'
data.append(chart)
return data
# +
# let's plot the cumulative returns
cumulative_rtns = (periodic_returns+1).cumprod() - 1
cumulative_returns = cumulative_rtns
chart_title = '60/40 Base Case'
emphasize = '60/40 Portfolio'
data = make_all_line_charts(emphasize)
layout = {'template': 'plotly_dark',
'title': chart_title,
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Cumulative Total Return'},
'tickformat': '.0%'}}
figure = go.Figure(data=data, layout=layout)
f2 = go.FigureWidget(figure)
f2
# -
# ### Plot Returns Chart - Logarithmic Scale
# +
# let's plot the returns on a logarithmic scale
log_cumulative_rtns = (periodic_returns+1).cumprod() * 100
cumulative_returns = log_cumulative_rtns
chart_title = '60/40 Base Case'
emphasize = '60/40 Portfolio'
data = make_all_line_charts(emphasize)
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Cumulative Total Return'},
'type': 'log', 'tickformat': '$.3s'},
'title': f'{chart_title} - Logarithmic Scale'})
figure = go.Figure(data=data, layout=layout)
f3 = go.FigureWidget(figure)
f3
# -
# ---
# ## Define Universe of Equity and Fixed Income ETFs
# Our optimized portfolios will be created using ETFs selected from this universe based on predictions from the Multi-task Lasso model made at a later step.
# ### Equity ETFs
# fetch daily return data for equity ETFs
equity_etfs = ['QQQ','VUG','VTV','IWF','IJR','IWM','IJH','VIG','IWD','VO','VGT','VB','XLK','XLF']
len(equity_etfs)
# ### Fixed Income ETFs
# fetch daily return data for fixed income ETFs
fi_etfs = ['VCIT','LQD','VCSH','BSV','TIP','IGSB','MBB','MUB','EMB','HYG','SHY','TLT']
len(fi_etfs)
# ---
# ## Pull Historical Data
# Now, let's get historical returns for our equity ETF universe.
# We can use the [10 Year Historical Monthly Prices](https://rapidapi.com/alphawave/api/stock-prices2) endpoint from the [AlphaWave Data Stock Prices API](https://rapidapi.com/alphawave/api/stock-prices2/endpoints) to pull in the ten year monthly historical prices so that we can calculate the returns.
#
# To call this API with Python, you can choose one of the supported Python code snippets provided in the API console. The following is an example of how to invoke the API with Python Requests. You will need to insert your own <b>x-rapidapi-host</b> and <b>x-rapidapi-key</b> information in the code block below.
# +
# fetch 10 year monthly return data
url = "https://stock-prices2.p.rapidapi.com/api/v1/resources/stock-prices/10y-1mo-interval"
headers = {
'x-rapidapi-host': "YOUR_X-RAPIDAPI-HOST_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS",
'x-rapidapi-key': "YOUR_X-RAPIDAPI-KEY_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS"
}
stock_frames = []
# for ticker in equity_etfs:
for ticker in tqdm(equity_etfs, position=0, leave=True, desc = "Retrieving AlphaWave Data Equity ETF Info"):
querystring = {"ticker":ticker}
stock_daily_price_response = requests.request("GET", url, headers=headers, params=querystring)
# Create Stock Prices DataFrame
stock_daily_price_df = pd.DataFrame.from_dict(stock_daily_price_response.json())
stock_daily_price_df = stock_daily_price_df.transpose()
stock_daily_price_df = stock_daily_price_df.rename(columns={'Close':ticker})
stock_daily_price_df = stock_daily_price_df[{ticker}]
stock_frames.append(stock_daily_price_df)
yf_combined_equity_etfs_df = pd.concat(stock_frames, axis=1, sort=True)
yf_combined_equity_etfs_df = yf_combined_equity_etfs_df.dropna(how='all')
yf_combined_equity_etfs_df = yf_combined_equity_etfs_df.fillna("")
equity_returns = yf_combined_equity_etfs_df.pct_change().dropna()
equity_returns
# -
# We next pull historical returns for our universe of fixed income ETFs.
# We can use the [10 Year Historical Monthly Prices](https://rapidapi.com/alphawave/api/stock-prices2) endpoint from the [AlphaWave Data Stock Prices API](https://rapidapi.com/alphawave/api/stock-prices2/endpoints) to pull in the ten year monthly historical prices so that we can calculate the returns.
#
# To call this API with Python, you can choose one of the supported Python code snippets provided in the API console. The following is an example of how to invoke the API with Python Requests. You will need to insert your own <b>x-rapidapi-host</b> and <b>x-rapidapi-key</b> information in the code block below.
# +
# fetch 10 year monthly return data
url = "https://stock-prices2.p.rapidapi.com/api/v1/resources/stock-prices/10y-1mo-interval"
headers = {
'x-rapidapi-host': "YOUR_X-RAPIDAPI-HOST_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS",
'x-rapidapi-key': "YOUR_X-RAPIDAPI-KEY_WILL_COPY_DIRECTLY_FROM_RAPIDAPI_PYTHON_CODE_SNIPPETS"
}
stock_frames = []
# for ticker in fi_etfs:
for ticker in tqdm(fi_etfs, position=0, leave=True, desc = "Retrieving AlphaWave Data FI ETF Info"):
querystring = {"ticker":ticker}
stock_daily_price_response = requests.request("GET", url, headers=headers, params=querystring)
# Create Stock Prices DataFrame
stock_daily_price_df = pd.DataFrame.from_dict(stock_daily_price_response.json())
stock_daily_price_df = stock_daily_price_df.transpose()
stock_daily_price_df = stock_daily_price_df.rename(columns={'Close':ticker})
stock_daily_price_df = stock_daily_price_df[{ticker}]
stock_frames.append(stock_daily_price_df)
yf_combined_fi_etfs_df = pd.concat(stock_frames, axis=1, sort=True)
yf_combined_fi_etfs_df = yf_combined_fi_etfs_df.dropna(how='all')
yf_combined_fi_etfs_df = yf_combined_fi_etfs_df.fillna("")
fi_returns = yf_combined_fi_etfs_df.pct_change().dropna()
fi_returns
# -
# ---
# ## Construct Time Series Model
#
# The goal of this model is to predict the returns for each of these equity and fixed income ETFs and pick the best ETFs to place in the portfolio. We will not be altering the 60/40 allocation split between equities and fixed income ETFs. We will be rebalancing monthly to target the 60/40 allocation while also changing the composition of equity and fixed income ETFs in the portfolio. This will give the model the ability to choose which equity and fixed income ETFs to invest in each month.
#
# For the equity strategy and fixed income strategy, the model will be trained on past return data only.
# ### Autoregressive Time Series Forecasting
#
# We will be using the Multi-task Lasso model in this analysis. The model looks at return data for equities and fixed income ETFs. It will train on five periods (monthly in this example) of returns in order to make a prediction of returns one month ahead. The Multi-task Lasso model allows us to fit multiple regression problems jointly. This means the model will look at all features at the same time to predict all of the future returns. For example, if we were analyzing stock A and stock B, the Multi-task Lasso model would not only look at the historical returns of stock A to predict the future returns for stock A. Rather, the model looks at the historical returns for both Stock A and stock B in order to predict the future returns for stock A.
#
# The reason we use Lasso is because it has a penalty term, called [regularization](https://medium.datadriveninvestor.com/introduction-to-machine-learning-an-overview-5ed43a37985d), for the betas that tries to shrink the coefficients down toward zero. If a coefficient is not important, it drops out of the model completely.
# <img src="img/autoregressive_gif.gif" >
@ignore_warnings(category=ConvergenceWarning)
def forecast_returns(return_time_series_data, non_return_data=None, window_size=5, num_test_dates=90):
"""
Use a given dataset and the MultiTaskLasso object from sklearn to
generate a DataFrame of predicted returns
Args:
================================
return_time_series_data (pandas.DataFrame):
pandas DataFrame of an actual return time series for a set of given indices.
Must be in the following format:
Period |
Ending | Ticker_1 Ticker_2 ... Ticker_N
----------- | ---------- ---------- ----- ----------
YYYY-MM-DD | 0.01 0.03 ... -0.05
|
YYYY-MM-DD | -0.05 -0.01 ... 0.04
non_return_data (pandas.DataFrame):
pandas DataFrame of an actual time series of non-return data
for a set of given indices. Must be in the same format, same
ticker order, and have the same periodicity as the return_time_series_data above
window_size (int):
Number of periods used to predict the next value.
Example: if window_size = 5, look 5 periods back to predict the next value
Default = 5
num_test_dates (int):
Number of periods for which to generate forecasts
Example: 120 = 10 years of monthly predictions, or 30 years of quarterly predicitons
depending on the periodicity of the input data in return_time_series_data and non_return_data
Default = 120
Returns:
================================
pandas.DataFrame
Output is a DataFrame of expected returns in the same format as return_time_series_data
"""
# descriptive variables for later use
names = list(return_time_series_data.columns)
dates = [f'{date.year}-{date.month}-{date.day}' for date in list(pd.to_datetime(return_time_series_data.index))]
# transform pandas to numpy arrays
X_returns = return_time_series_data.to_numpy()
X_input = X_returns
max_iter = 7500
# concatenate non_return_data if it exists
if non_return_data is not None:
max_iter = 3000
X_non_rtn = non_return_data.to_numpy()
X_input = np.concatenate((X_returns, X_non_rtn), axis=1)
# number of time series (tickers) to model
n_series = X_returns.shape[1]
# number of features at each date; equal to n_series * number of features (return, oas_spread, etc.)
n_features_per_time_point = X_input.shape[1]
num_features = window_size * n_features_per_time_point
num_training_points = X_returns.shape[0] - window_size
X_train = np.zeros((num_training_points, num_features))
Y_train = X_returns[window_size:,:]
for i in range(num_training_points-1):
X_train[i,:] = np.matrix.flatten(X_input[i : window_size + i,:])
# establish empty arrays & variables for use in training each model
mtl_list=[]
alpha= 0.001
Y_pred = np.zeros((num_test_dates, n_series))
delta_Y = np.zeros((num_test_dates, n_series))
dY_percent = np.zeros((num_test_dates, n_series))
mse_pred = np.zeros(num_test_dates)
predict_dates=[]
# loop through dates & predict returns
for i in range(num_test_dates):
X_i = X_train[:num_training_points - num_test_dates + (i-1)]
Y_i = Y_train[:num_training_points - num_test_dates + (i-1)]
print("X shape: ", X_i.shape, "Y shape: ", Y_i.shape)
print("number of points in training data:", X_i.shape[0] )
mtl = MultiTaskLasso(alpha=alpha, max_iter=max_iter, warm_start=True).fit(X_i, Y_i)
mtl_list.append(mtl)
print(f"using X from {dates[num_training_points - num_test_dates + (i-1) + window_size]}\
to predict {dates[num_training_points - num_test_dates + (i-1) + 1 + window_size]}")
predict_dates.append(dates[num_training_points - num_test_dates + (i-1) + window_size])
X_i_plus_1 = X_train[num_training_points - num_test_dates + (i-1) + 1]
Y_pred[i,:] = mtl.predict([X_i_plus_1])
Y_act = Y_train[num_training_points - num_test_dates + (i-1) + 1]
delta_Y[i] = (Y_pred[i,:] - Y_act)
mse_pred[i] = np.sqrt(np.sum((Y_pred[i,:] - Y_act)**2))/len(Y_act)
print("mse", mse_pred[i])
predictions = pd.DataFrame(Y_pred, index=predict_dates, columns=names)
predictions.index = [pd.Timestamp(i).strftime('%Y-%m-%d') for i in predictions.index]
return predictions
# run the model
eq_predictions = forecast_returns(equity_returns)
fi_predictions = forecast_returns(fi_returns)
# view predictions
eq_predictions.head()
# view returns
equity_returns.head()
# calculate the average equity prediction error
average_equity_return_error = eq_predictions.subtract(equity_returns).mean(axis=1).dropna()
equity_avg_error_plot_df = pd.DataFrame({'Avg Error': average_equity_return_error}, index=average_equity_return_error.index)
equity_avg_error_plot_df
# calculate the average fixed income prediction error
average_fi_return_error = fi_predictions.subtract(fi_returns).mean(axis=1).dropna()
fi_avg_error_plot_df = pd.DataFrame({'Avg Error': average_fi_return_error}, index=average_fi_return_error.index)
fi_avg_error_plot_df
# Next, we check if the model introduces any systematic bias by calculating the average prediction error per month. This means we take the average difference between the actual and estimated returns for each month. If the error plotted in the below charts were all negative or all positive, then we would know our model has a systematic bias. According to the charts below, it appears we do not have a systematic bias which means we are clear to proceed.
# +
# check if the model introduces any systematic bias for equity ETFs
def SetColor(y):
if(y < 0):
return "red"
elif(y >= 0):
return "green"
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Avg Error %'}},
'title': f'Average Equity Prediction Error'})
fig = go.Figure(layout=layout)
fig.add_trace(go.Bar(
x=equity_avg_error_plot_df.index,
y=equity_avg_error_plot_df.iloc[:,0],
marker=dict(color = list(map(SetColor, equity_avg_error_plot_df.iloc[:,0])))
))
f4 = go.FigureWidget(fig)
f4
# +
# check if the model introduces any systematic bias for fixed income ETFs
def SetColor(y):
if(y < 0):
return "red"
elif(y >= 0):
return "green"
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Avg Error %'}},
'title': f'Average Fixed Income Prediction Error'})
fig = go.Figure(layout=layout)
fig.add_trace(go.Bar(
x=fi_avg_error_plot_df.index,
y=fi_avg_error_plot_df.iloc[:,0],
marker=dict(color = list(map(SetColor, fi_avg_error_plot_df.iloc[:,0])))
))
f5 = go.FigureWidget(fig)
f5
# -
# ---
# ## Allocate Strategy Portfolio Based on Model Results
# Create three strategies to measure model performance:
# 1. 60/40 Allocation Strategy
# 1. Equity Only Portfolio
# 1. Fixed Income Only Portfolio
#
# Below we define `allocate_portfolio` and `get_historical_portfolio_holdings` functions to create these three strategies. These functions help us identify the equity and fixed income ETFs with the largest expected returns as calculated by our model for each month, which we then place in our optimized portfolios to see if they beat the benchmarks.
def allocate_portfolio(expected_eq_returns,
expected_fi_returns,
actual_eq_returns,
actual_fi_returns,
for_period_ending,
total_equity_weight=0.6,
n_equity_funds=5,
n_bond_funds=5):
"""
Allocate a portfolio by picking the top n_equity_funds & top n_bond_funds for the period
ending on for_period_ending
"""
fi_wgt = 1 - total_equity_weight
eq_fund_wgt = total_equity_weight / n_equity_funds
fi_fund_wgt = fi_wgt / n_bond_funds
for_period_ending = pd.Timestamp(for_period_ending).strftime('%Y-%m-%d')
eq_returns = pd.DataFrame(expected_eq_returns.loc[for_period_ending])
eq_returns.columns = ['Expected Return']
eq_returns['Type'] = ['Equity'] * len(eq_returns)
eq_returns['Weight'] = [eq_fund_wgt] * len(eq_returns)
eq_returns = eq_returns.sort_values(by='Expected Return', ascending=False).head(n_equity_funds)
fi_returns = pd.DataFrame(expected_fi_returns.loc[for_period_ending])
fi_returns.columns = ['Expected Return']
fi_returns['Type'] = ['Fixed Income'] * len(fi_returns)
fi_returns['Weight'] = [fi_fund_wgt] * len(fi_returns)
fi_returns = fi_returns.sort_values(by='Expected Return', ascending=False).head(n_bond_funds)
holdings_df = pd.concat([eq_returns, fi_returns], axis=0)
holdings_df.index.name = 'Index'
actual_returns = []
for i in range(len(holdings_df)):
index_type = holdings_df['Type'].iloc[i]
index_name = holdings_df.index[i]
if index_type == 'Equity':
actual_returns.append(actual_eq_returns[index_name].loc[for_period_ending])
elif index_type == 'Fixed Income':
actual_returns.append(actual_fi_returns[index_name].loc[for_period_ending])
holdings_df['Actual Return'] = actual_returns
holdings_df.index = pd.MultiIndex.from_tuples([(for_period_ending, i) for i in holdings_df.index], names=['For Period Ending', 'Fund Ticker'])
holdings_df = holdings_df[['Type', 'Weight', 'Expected Return', 'Actual Return']]
return holdings_df
def get_historical_portfolio_holdings(expected_eq_returns,
expected_fi_returns,
actual_eq_returns,
actual_fi_returns,
total_equity_weight):
"""
Loop over the time frame given in expected_fi_returns
and run allocate_portfolio at each date
"""
holdings = []
for date in expected_fi_returns.index:
holdings_at_date = allocate_portfolio(expected_eq_returns=expected_eq_returns,
expected_fi_returns=expected_fi_returns,
actual_eq_returns=actual_eq_returns,
actual_fi_returns=actual_fi_returns,
for_period_ending=date,
total_equity_weight=total_equity_weight)
holdings.append(holdings_at_date)
return pd.concat(holdings)
# Run the functions to create our historical ETF holdings.
# +
params = {'expected_eq_returns': eq_predictions,
'expected_fi_returns': fi_predictions,
'actual_eq_returns': equity_returns,
'actual_fi_returns': fi_returns}
portfolio_holdings = get_historical_portfolio_holdings(**params, total_equity_weight=0.6)
bond_only_holdings = get_historical_portfolio_holdings(**params, total_equity_weight=0)
equity_only_holdings = get_historical_portfolio_holdings(**params, total_equity_weight=1)
portfolio_holdings.tail(20)
# -
# ---
# ## Calculate Benchmark & Strategy Portfolio Returns
# Before we begin, let's review the assumptions we apply to this example:
# * Assume 0% slippage and trading fees
# * Risk measures not considered
#
# Below we define `get_excess_return`, `get_excess_return_string`, and `get_portfolio_returns` functions that will help us calculate and compare the returns of the optimized portfolios to those of the benchmarks.
def get_excess_return(strategy, benchmark):
investment_horizon_years = (datetime.strptime(periodic_returns.index[-1], '%Y-%m-%d') - datetime.strptime(periodic_returns.index[0], '%Y-%m-%d')).days / 365
annualized_excess_return = (cumulative_returns[strategy][-1] / cumulative_returns[benchmark][-1]) ** (1/investment_horizon_years) - 1
return annualized_excess_return
def get_excess_return_string(strategy, benchmark):
start_date = periodic_returns.index[0]
end_date = periodic_returns.index[-1]
r = get_excess_return(strategy=strategy, benchmark=benchmark)
qualifier = 'UNDERPERFORMED'
if r > 0:
qualifier = 'OUTPERFORMED'
return f'{strategy} {qualifier} {benchmark} by an annualized rate of {r:.2%} per year for the period between {start_date} and {end_date}.'
def get_portfolio_returns(portfolio_holdings_df, port_name='Optimized Portfolio'):
weighted_returns = portfolio_holdings_df['Actual Return'] * portfolio_holdings_df['Weight']
returns_df = pd.DataFrame(weighted_returns.groupby(level=[0]).sum())
returns_df.columns = [port_name]
return returns_df
# Let's take a look at a dataframe that includes the returns of all the optimized portfolios and benchmarks.
# +
new_60_40_returns = get_portfolio_returns(portfolio_holdings, 'Optimized 60/40')
bond_strategy_rtns = get_portfolio_returns(bond_only_holdings, 'Optimized Bond Strategy')
equity_strategy_rtns = get_portfolio_returns(equity_only_holdings, 'Optimized Equity Strategy')
all_returns = pd.concat([periodic_returns, new_60_40_returns, bond_strategy_rtns, equity_strategy_rtns], axis=1).dropna()
all_returns.head()
# -
# ### Fixed Income Only Strategy
# Let's see if the Optimized Fixed Income Only Strategy beats its benchmark.
# +
# calculate the returns
periodic_returns = all_returns[['AGG', 'Optimized Bond Strategy']]
log_cumulative_rtns = (periodic_returns+1).cumprod() * 100
cumulative_returns = log_cumulative_rtns
cumulative_returns
# +
# let's plot the returns on a logarithmic scale
chart_title = 'Optimized FI Returns vs Bond Index'
emphasize = 'Optimized Bond Strategy'
data = make_all_line_charts(emphasize)
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Cumulative Total Return'},
'type': 'log', 'tickformat': '$.3s'},
'title': f'{chart_title} - Logarithmic Scale'})
figure = go.Figure(data=data, layout=layout)
f6 = go.FigureWidget(figure)
f6
# -
print(get_excess_return_string(strategy='Optimized Bond Strategy', benchmark='AGG'))
# ### Equity Only Strategy
# Let's see if the Optimized Equity Only Strategy beats its benchmark.
# +
# calculate the returns
periodic_returns=all_returns[['SPY', 'Optimized Equity Strategy']]
log_cumulative_rtns = (periodic_returns+1).cumprod() * 100
cumulative_returns = log_cumulative_rtns
cumulative_returns
# +
# let's plot the returns on a logarithmic scale
chart_title = 'Optimized Equity Returns vs Equity Index'
emphasize = 'Optimized Equity Strategy'
data = make_all_line_charts(emphasize)
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Cumulative Total Return'},
'type': 'log', 'tickformat': '$.3s'},
'title': f'{chart_title} - Logarithmic Scale'})
figure = go.Figure(data=data, layout=layout)
f7 = go.FigureWidget(figure)
f7
# -
print(get_excess_return_string(strategy='Optimized Equity Strategy', benchmark='SPY'))
# ### 60/40 Allocation Strategy
# Let's see if the Optimized 60/40 Allocation Strategy beats its benchmark.
# +
# calculate the returns
periodic_returns = all_returns[['60/40 Portfolio', 'Optimized 60/40']]
log_cumulative_rtns = (periodic_returns+1).cumprod() * 100
cumulative_returns = log_cumulative_rtns
cumulative_returns
# +
# let's plot the returns on a logarithmic scale
chart_title = 'Optimized 60/40 Returns vs 60/40 Portfolio Index'
emphasize = 'Optimized 60/40'
data = make_all_line_charts(emphasize)
layout = ({'template': 'plotly_dark',
'xaxis': {'title': {'text': 'Date'}},
'yaxis': {'title': {'text': 'Cumulative Total Return'},
'type': 'log', 'tickformat': '$.3s'},
'title': f'{chart_title} - Logarithmic Scale'})
figure = go.Figure(data=data, layout=layout)
f8 = go.FigureWidget(figure)
f8
# -
print(get_excess_return_string(strategy='Optimized 60/40', benchmark='60/40 Portfolio'))
# As a framework, we see that the model's optimized portfolios outperform their benchmarks. We can make this model more complex by adding more data if we wish, but as a start it appears the model is able to make predictions that result in allocations outperforming their benchmarks.
# ---
# ## Additional Resources
# [Machine Learning for Investing](https://hdonnelly6.medium.com/list/machine-learning-for-investing-7f2690bb1826)
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by AlphaWave Data, Inc. ("AlphaWave Data"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, AlphaWave Data, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to AlphaWave Data, Inc. at the time of publication. AlphaWave Data makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| AlphaWave Data Machine Learning used for Asset Allocation example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 16
# + [markdown] tags=[]
# *Modeling and Simulation in Python*
#
# Copyright 2021 <NAME>
#
# License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=[]
# install Pint if necessary
try:
import pint
except ImportError:
# !pip install pint
# + tags=[]
# download modsim.py if necessary
from os.path import basename, exists
def download(url):
filename = basename(url)
if not exists(filename):
from urllib.request import urlretrieve
local, _ = urlretrieve(url, filename)
print('Downloaded ' + local)
download('https://github.com/AllenDowney/ModSimPy/raw/master/' +
'modsim.py')
# + tags=[]
# import functions from modsim
from modsim import *
# -
# [Click here to run this chapter on Colab](https://colab.research.google.com/github/AllenDowney/ModSimPy/blob/master//chapters/chap16.ipynb)
# + tags=[]
download('https://github.com/AllenDowney/ModSimPy/raw/master/' +
'chap15.py')
# + tags=[]
# import code from previous notebooks
from chap15 import change_func
from chap15 import run_simulation
from chap15 import make_system
# -
# In the previous chapter we wrote a simulation of a cooling cup of
# coffee. Given the initial temperature of the coffee, the temperature of the atmosphere, and the rate parameter, `r`, we can predict how the
# temperature of the coffee will change over time.
# Then we used a root finding algorithm to estimate `r` based on data I invented for the example.
#
# If you did the exercises, you simulated the temperature of the milk as it warmed, and estimated its rate parameter as well.
#
# Now let's put it together.
# In this chapter we'll write a function that simulates mixing the two liquids and use it to answer the question we started with: is it better to mix the coffee and milk at the beginning, the end, or somewhere in the middle?
# ## Mixing liquids
#
# When we mix two liquids, the temperature of the mixture depends on the
# temperatures of the ingredients as well as their volumes, densities, and specific heat capacities (as defined in the previous chapter).
# In this section I'll explain how.
#
# Assuming there are no chemical reactions that either produce or consume heat, the total thermal energy of the system is the same before and after mixing; in other words, thermal energy is **conserved**.
#
# If the temperature of the first liquid is $T_1$, the temperature of the second liquid is $T_2$, and the final temperature of the mixture is $T$, the heat transfer into the first liquid is $C_1 (T - T_1)$ and the heat transfer into the second liquid is $C_2 (T - T_2)$, where $C_1$ and $C_2$ are the thermal masses of the liquids.
#
# In order to conserve energy, these heat transfers must add up to 0:
#
# $$C_1 (T - T_1) + C_2 (T - T_2) = 0$$
#
# We can solve this equation for T:
#
# $$T = \frac{C_1 T_1 + C_2 T_2}{C_1 + C_2}$$
#
# For the coffee cooling problem, we have the volume of each liquid; if we also know the density, $\rho$, and the specific heat capacity, $c_p$, we can compute thermal mass:
#
# $$C = \rho V c_p$$
#
# If the liquids have the same density and heat capacity, they drop out of the equation, and we can write:
#
# $$T = \frac{V_1 T_1 + V_2 T_2}{V_1 + V_2}$$
#
# where $V_1$ and $V_2$ are the volumes of the liquids.
#
# As an approximation, I'll assume that milk and coffee have the same
# density and specific heat. As an exercise, you can look up these
# quantities and see how good this assumption is.
#
# The following function takes two `System` objects, representing the
# coffee and milk, and creates a new `System` to represent the mixture:
def mix(system1, system2):
V1, V2 = system1.volume, system2.volume
T1, T2 = system1.T_final, system2.T_final
V_mix = V1 + V2
T_mix = (V1 * T1 + V2 * T2) / V_mix
return make_system(T_init=T_mix,
volume=V_mix,
r=system1.r,
t_end=30)
# The first two lines extract volume and temperature from the `System` objects. The next two lines compute the volume and temperature of the mixture. Finally, `mix` makes a new `System` object and returns it.
#
# This function uses the value of `r` from `system1` as the value of `r`
# for the mixture. If `system1` represents the coffee, and we are adding
# the milk to the coffee, this is probably a reasonable choice. On the
# other hand, when we increase the amount of liquid in the coffee cup,
# that might change `r`. So this is an assumption we might want to
# revisit.
#
# Now we have everything we need to solve the problem.
# ## Mix first or last?
#
# First I'll create objects to represent the coffee and milk.
# For `r_coffee`, I'll use the value we computed in the previous chapter.
r_coffee = 0.0115
coffee = make_system(T_init=90, volume=300, r=r_coffee, t_end=30)
# For `r_milk`, I'll use the value I estimated in the exercise from the previous chapter.
r_milk = 0.133
milk = make_system(T_init=5, volume=50, r=r_milk, t_end=15)
# Now we can mix them and simulate 30 minutes:
# +
mix_first = mix(coffee, milk)
run_simulation(mix_first, change_func)
mix_first.T_final
# -
# The final temperature is 61.5 °C which is still warm enough to be
# enjoyable. Would we do any better if we added the milk last?
#
# I'll simulate the coffee and milk separately, and then mix them:
run_simulation(coffee, change_func)
run_simulation(milk, change_func)
mix_last = mix(coffee, milk)
mix_last.T_final
# After mixing, the temperature is 62.9 °C, so it looks like adding the
# milk at the end is better.
# But is that the best we can do?
# ## Optimization
#
# Adding the milk after 30 minutes is better than adding it immediately, but maybe there's something in between that's even better. To find out, I'll use the following function, which takes the time to add the milk, `t_add`, as a parameter:
def run_and_mix(t_add, t_total):
coffee.t_end = t_add
coffee_results = run_simulation(coffee, change_func)
milk.t_end = t_add
milk_results = run_simulation(milk, change_func)
mixture = mix(coffee, milk)
mixture.t_end = t_total - t_add
results = run_simulation(mixture, change_func)
return mixture.T_final
# `run_and_mix` simulates both systems for the given time, `t_add`.
# Then it mixes them and simulates the mixture fir the remaining time, `t_total - t_add`.
#
# When `t_add` is`0`, we add the milk immediately; when `t_add` is `30`, we add it at the end. Now we can sweep the range of values in between:
sweep = SweepSeries()
for t_add in linspace(0, 30, 11):
sweep[t_add] = run_and_mix(t_add, 30)
# Here's what the results look like:
# +
sweep.plot(label='mixture', color='C2')
decorate(xlabel='Time until mixing (minutes)',
ylabel='Final emperature (C)')
# -
# Note that this is a parameter sweep, not a time series.
#
# The final temperature is maximized when `t_add=30`, so adding the milk
# at the end is optimal.
# ## Analysis
#
# Simulating Newton's law of cooling isn't really necessary because we can solve the differential equation analytically. If
#
# $$\frac{dT}{dt} = -r (T - T_{env})$$
#
# the general solution is
#
# $$T{\left (t \right )} = C \exp(-r t) + T_{env}$$
#
# and the particular solution where $T(0) = T_{init}$ is
#
# $$T_{env} + \left(- T_{env} + T_{init}\right) \exp(-r t)$$
#
# If you would like to see this solution done by hand, you can watch this video: <http://modsimpy.com/khan3>.
# Now we can use the observed data to estimate the parameter $r$. If we
# observe the that temperature at $t_{end}$ is $T_{final}$, we can plug these values into the particular solution and solve for $r$. The result is:
#
# $$r = \frac{1}{t_{end}} \log{\left (\frac{T_{init} - T_{env}}{T_{final} - T_{env}} \right )}$$
#
# The following function takes a `System` object and computes `r`:
# +
from numpy import log
def compute_r(system):
t_end = system.t_end
T_init = system.T_init
T_final = system.T_final
T_env = system.T_env
r = log((T_init - T_env) / (T_final - T_env)) / t_end
return r
# -
# We can use this function to compute `r` for the coffee, given the parameters of the problem.
coffee2 = make_system(T_init=90, volume=300, r=0, t_end=30)
coffee2.T_final = 70
r_coffee2 = compute_r(coffee2)
r_coffee2
# This value is close to the value of `r` we computed in the previous chapter, `0.115`, but not exactly the same.
# That's because the simulations use discrete time steps, and the analysis uses continuous time.
#
# Nevertheless, the results of the analysis are consistent with the simulation.
# To check, we'll use the following function, which takes a `System` object and uses the analytic result to compute a time series:
# +
from numpy import exp
def run_analysis(system):
T_env, T_init, r = system.T_env, system.T_init, system.r
t_array = linrange(system.t_0, system.t_end, system.dt)
T_array = T_env + (T_init - T_env) * exp(-r * t_array)
system.T_final = T_array[-1]
return make_series(t_array, T_array)
# -
# The first line unpacks the system variables.
# The next two lines compute `t_array`, which is a NumPy array of time stamps, and `T_array`, which is an array of the corresponding temperatures.
#
# The last two lines store the final temperature in the `System` object and use `make_series` to return the results in a Pandas `Series`.
#
# We can run it like this:
coffee2.r = r_coffee2
results2 = run_analysis(coffee2)
coffee2.T_final
# The final temperature is 70 °C, as it should be. In fact, the results
# are identical to what we got by simulation, with a small difference due to rounding.
# + tags=[]
coffee.r = 0.011543
results = run_simulation(coffee, change_func)
# + tags=[]
from numpy import allclose
allclose(results, results2)
# -
# ## Summary
#
# In this chapter we finished the coffee cooling problem from the previous chapter, and found that it is better to add the milk at the end, at least for the version of the problem I posed.
#
# As an exercise you will have a chance to explore a variation of the problem where the answer might be different.
#
# In the next chapter we'll move on to a new example, a model of how glucose and insulin interact to control blood sugar.
# ## Exercises
# **Exercise:** Use `compute_r` to compute `r_milk` according to the analytic solution. Run the analysis with this value of `r_milk` and confirm that the results are consistent with the simulation.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# **Exercise:** Suppose the coffee shop won't let me take milk in a separate container, but I keep a bottle of milk in the refrigerator at my office. In that case is it better to add the milk at the coffee shop, or wait until I get to the office?
#
# Hint: Think about the simplest way to represent the behavior of a refrigerator in this model. The change you make to test this variation of the problem should be very small!
# +
# Solution goes here
| chapters/chap16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro notebook
#
# Some important links to keep open during the workshop – open these tabs **now**!:
#
# - [TF documentation](https://www.tensorflow.org/versions/master/api_guides/python/array_ops) : Use the search box (top right) to get documentation on Tensorflow's rich API
#
# - [solutions/](https://github.com/tensorflow/workshops/tree/master/extras/amld/notebooks/solutions) : Every notebook in the `exercises/` directory has a corresponding notebook in the `solutions/` directory.
import tensorflow as tf
import numpy as np
from matplotlib import pyplot
# %matplotlib inline
# Always make sure you are using running the expected version.
# There are considerable differences between versions...
# Make sure your version is "1.4.X"
tf.__version__
# Important shortcuts – **give it a try!**
#
# If you run this notebook with **Colab**:
#
# - `<CTRL-ENTER>` : executes current cell
# - `<SHIFT-ENTER>` : executes current cell and moves to next cell
# - `<CTRL-SHIFT-P>` : shows searchable command palette
# - `<CTRL-M> <A>` : insert cell above
# - `<CTRL-M> <B>` : append cell below
# - `<CTRL-M> <Y>` : convert cell to code
# - `<CTRL-M> <M>` : convert cell to Markdown
#
# If you run this notebook with **Jupyter**:
#
# - `<ESC>` : goes from edit to "command" mode
# - `<ENTER>` : goes from "command" to edit mode
# - `<CTRL-ENTER>` : executes current cell
# - `<SHIFT-ENTER>` : executes current cell and moves to next cell
# - `<h>` : shows help (works only in "command" mode)
# - `<CTRL-SHIFT-P>` : shows searchable command palette
# - `<m>` : change cell from "code" to "markdown" (works only in "command" mode)
# - `<y>` : change cell from "markdown" to "code" (works only in "command" mode)
#
# Note: On OS X you can use `<COMMAND>` instead of `<CTRL>`
# Check that matplotlib is working.
x = np.random.rand(100).astype(np.float32)
noise = np.random.normal(scale=0.3, size=len(x))
y = np.sin(x * 7) + noise
pyplot.scatter(x, y)
# You can run shell commands directly in Jupyter: simply prepend
# the command with a "!".
# !pwd
# !ls
# Auto-completion: Jupyter shows possible completions of partially typed
# commands -- place the cursor below just "one" and type <TAB>
### YOUR ACTION REQUIRED:
# Complete below line.
tf.one_hot #tf.one
# +
def xor_str(a, b):
return ''.join([chr(ord(a[i % len(a)]) ^ ord(b[i % len(b)]))
for i in range(max(len(a), len(b)))])
### YOUR ACTION REQUIRED:
# Try to find the "correct value" for below variable...
workshop_secret = 'Tensorflow rocks' #workshop_secret = '(replace me!)'
xor_str(workshop_secret, '\x03\x00\x02\x10\x00\x1f\x03L\x1b\x18\x00\x06\x07\x06K2\x19)*S;\x17\x08\x1f\x00\x05F\x1e\x00\x14K\x115\x16\x07\x10\x1cR1\x03\x1d\x1cS\x1a\x00\x13J')
# Hint: You might want to checkout the ../solutions directory
# (you should already have opened this directory in a browser tab :-)
| extras/amld/notebooks/solutions/0_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('ggplot')
import numpy as np
import matplotlib.pyplot as plt
import numpy.ma as ma
from netCDF4 import Dataset, date2index, num2date
from palettable import colorbrewer
from datetime import datetime, timedelta
from bs4 import BeautifulSoup
import requests
def listFD(url, ext=''):
page = requests.get(url).text
#print(page)
soup = BeautifulSoup(page, 'html.parser')
return [url + node.get('href') for node in soup.find_all('a') if (node.get('href').endswith(ext) and node.get('href').startswith('2'))]
# +
from os.path import exists
filename_viirs_proc = 'F:/data/cruise_data/saildrone/baja-2018/viirs_l2p_files.npy'
filename_viirs_sst = 'F:/data/cruise_data/saildrone/baja-2018/viirs_sst_l2p_data.npy'
filename_png = 'F:/data/cruise_data/saildrone/baja-2018/recent_viirs_sst_l2p_.png'
# +
for incr_day in range(0,1):
d = datetime.today() + timedelta(days=incr_day)
day_of_year = d.timetuple().tm_yday
url = 'https://opendap.jpl.nasa.gov/opendap/OceanTemperature/ghrsst/data/GDS2/L2P/VIIRS_NPP/OSPO/v2.41/' \
+ str(d.year) + '/' + str(day_of_year).zfill(3) + '/'
ext = 'nc'
filenames=listFD(url, ext)
ilen=len(filenames)
for ic in range(1,ilen):
file = filenames[ic]
print(file)
#save which files have been processed
idyj=int(file[101:104])
ihr=int(file[113:115])
imin=int(file[115:117])
print(idyj,ihr,imin)
nc = Dataset(file)
lat2 = nc.variables['lat'][:] #[2500:3250]
nc.close()
maxlat = np.ma.max(lat2)
minlat = np.ma.min(lat2)
ilat_check=0
if minlat<40.0 and maxlat>40.0:
ilat_check=1
if minlat>25.0 and maxlat<40.0:
ilat_check=1
if minlat<25.0 and maxlat>25.0:
ilat_check=1
if ilat_check==0:
continue #no data at right latitude
nc = Dataset(file)
lon2 = nc.variables['lon'][:] #[2500:3250]
nc.close()
minlon = np.ma.min(lon2)
print(maxlat,minlat)
if minlon>0:
continue
minlon = np.ma.min(lon2[lon2<0])
maxlon = np.ma.max(lon2[lon2<0])
ilon_check=0
if minlon<-130 and maxlon>-130.0:
ilon_check=1
if minlon>-130 and maxlon<-110.0:
ilon_check=1
if minlon<-110 and maxlon>-110.0:
ilon_check=1
if ilon_check==1 and ilat_check==1:
nc = Dataset(file)
sst = nc.variables['sea_surface_temperature'][:] #:,2500:3250,2500:3500]
nc.close()
print(sst.shape,lat2.shape,lon2.shape)
break
# +
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
import numpy as np
#print(lon2[lon2>0].count())
#print(lon2[lon2<0].count())
print(sst.shape,lon2.shape,lat2.shape)
#if minlon<-110.0 and minlon
#25.03 39.99 -129.97 -110.01
#plt.scatter(lon,lat)
#plt.show()
print('drawing map....') # "Drawing map... ",
m = Basemap(projection='cyl', llcrnrlat=minlat, urcrnrlat=maxlat,\
llcrnrlon=minlon, urcrnrlon=maxlon, resolution='l')
m.drawcoastlines()
m.drawmapboundary(fill_color='coral')
#m.scatter(xlist, ylist, 20, c=confidence, cmap=p.cm.hot, marker='o', edgecolors='none', zorder=10)
#m.imshow(sst)
print('done') # "done !"
# +
#import numpy as np
#from pyresample import image, geometry
#area_def = geometry.AreaDefinition('areaD', 'Europe (3km, HRV, VTC)', 'areaD',
#... {'a': '6378144.0', 'b': '6356759.0',
#... 'lat_0': '50.00', 'lat_ts': '50.00',
#... 'lon_0': '8.00', 'proj': 'stere'},
#... 800, 800,
#... [-1370912.72, -909968.64,
#... 1029087.28, 1490031.36])
#swath_def = geometry.SwathDefinition(lons=lon2, lats=lat2)
#swath_con = image.ImageContainerNearest(sst, swath_def, radius_of_influence=5000)
#area_con = swath_con.resample(area_def)
#result = area_con.image_data
# +
from palettable import colorbrewer
from copy import copy
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.mlab as mlab
palette = copy(plt.cm.jet)
palette.set_over('r', 1.0)
palette.set_under('g', 1.0)
palette.set_bad(alpha = 0.0)
#fig = plt.figure(figsize=(6, 5.4))
# plot using 'continuous' color map
print(sst.shape)
fig, ax = plt.subplots()
im = ax.imshow(sst[0,:,:].T-273.15, interpolation='bilinear',cmap=palette,norm=colors.Normalize(vmin=10, vmax=17.0),aspect='auto',origin='lower')
# We want to show all ticks...
ax.set_xticks(lon2)
ax.set_yticks(lat2)
# ... and label them with the respective list entries
#ax.set_xticklabels(farmers)
#extent=[x0, x1, y0, y1])
#fig.set_title('SST')
#cbar = fig.colorbar(im, extend='both', shrink=0.9, ax=im)
#cbar.set_label('uniform')
#for ticklabel in ax1.xaxis.get_ticklabels():
# ticklabel.set_visible(False)
plt.show()
# +
if inew_data==1:
sst_new = ma.copy(sst)
inew_data = 0
if cnt<5:
continue
print(ic,cnt,file)
mask = sst_new < -10
sst_new[mask] = sst[mask]
if itotal_proc>0:
lat, lon = np.meshgrid(lat, lon)
mask = (sst_new < -10)
sst_new[mask]=-9999.
sst_new2 = sst_new[0,:,:]
sstx = ma.masked_values (sst_new2, -9999.)
print(lon.shape,lat.shape,sstx.shape)
sst3x = np.flip(sstx, 1)
sst4x = np.flip(sst3x, 0)
sst5x = ma.swapaxes(sst4x,0,1)
print(lon.shape,lat.shape,sst5x.shape)
pixels = 1024 * 10
cmap = colorbrewer.get_map('Spectral', 'diverging', 11, reverse=True).mpl_colormap
fig, ax = gearth_fig(llcrnrlon=lon.min(),
llcrnrlat=lat.min(),
urcrnrlon=lon.max(),
urcrnrlat=lat.max(),
pixels=pixels)
cs = ax.pcolormesh(lon, lat, sst5x - 273.15, cmap=cmap, vmin = 10, vmax = 17)
ax.set_axis_off()
fig.savefig(filename_png, transparent=False, format='png')
print('done')
print(lat[1,1],lat[1,-1],lon[1,1],lon[-1,1])
make_kml(llcrnrlon=lon[1,1], llcrnrlat=lat[1,1],
urcrnrlon=lon[-1,1], urcrnrlat=lat[1,-1],
figs= [filename_png], colorbar='legend.png',
kmzfile=filename_kmz, name='VIIRS Sea Surface Temperature')
np.save(filename_viirs_proc, proc_save)
sst_new.dump(filename_viirs_sst)
fig = plt.figure(figsize=(1.0, 4.0), facecolor=None, frameon=False)
ax = fig.add_axes([0.0, 0.05, 0.2, 0.9])
cb = fig.colorbar(cs, cax=ax)
cb.set_label('SST [C]', rotation=-90, color='k', labelpad=20)
fig.savefig('legend.png', transparent=False, format='png') # Change transparent to True if your colorbar is not on space :)
# +
#now just do todays data
from os.path import exists
filename_png = 'F:/data/cruise_data/saildrone/baja-2018/today_viirs_sst.png'
filename_kmz = 'F:/data/cruise_data/saildrone/baja-2018/todat_viirs_sst.kmz'
inew_data=1
for incr_day in range(-1,1):
d = datetime.today() + timedelta(days=incr_day)
day_of_year = d.timetuple().tm_yday
url = 'https://opendap.jpl.nasa.gov/opendap/OceanTemperature/ghrsst/data/GDS2/L3U/VIIRS_NPP/OSPO/v2.41/' \
+ str(d.year) + '/' + str(day_of_year).zfill(3) + '/'
ext = 'nc'
filenames=listFD(url, ext)
ilen=len(filenames)
for ic in range(1,ilen):
file = filenames[ic]
#save which files have been processed
idyj=int(file[101:104])
ihr=int(file[113:115])
imin=int(file[115:117])
nc = Dataset(file)
sst = nc.variables['sea_surface_temperature'][:,2500:3250,2500:3500]
sst = np.flip(sst, 2)
cnt = ma.count(sst)
lat = nc.variables['lat'][2500:3250]
lat = np.flip(lat, 0)
lon = nc.variables['lon'][2500:3500]
nc.close()
if inew_data==1:
sst_new = ma.copy(sst)
inew_data = 0
if cnt<5:
continue
print(ic,cnt,file)
mask = sst_new < -10
sst_new[mask] = sst[mask]
lat, lon = np.meshgrid(lat, lon)
mask = (sst_new < -10)
sst_new[mask]=-9999.
sst_new2 = sst_new[0,:,:]
sstx = ma.masked_values (sst_new2, -9999.)
print(lon.shape,lat.shape,sstx.shape)
sst3x = np.flip(sstx, 1)
sst4x = np.flip(sst3x, 0)
sst5x = ma.swapaxes(sst4x,0,1)
print(lon.shape,lat.shape,sst5x.shape)
pixels = 1024 * 10
cmap = colorbrewer.get_map('Spectral', 'diverging', 11, reverse=True).mpl_colormap
fig, ax = gearth_fig(llcrnrlon=lon.min(),
llcrnrlat=lat.min(),
urcrnrlon=lon.max(),
urcrnrlat=lat.max(),
pixels=pixels)
cs = ax.pcolormesh(lon, lat, sst5x - 273.15, cmap=cmap, vmin = 10, vmax = 17)
ax.set_axis_off()
fig.savefig(filename_png, transparent=False, format='png')
print('done')
print(lat[1,1],lat[1,-1],lon[1,1],lon[-1,1])
make_kml(llcrnrlon=lon[1,1], llcrnrlat=lat[1,1],
urcrnrlon=lon[-1,1], urcrnrlat=lat[1,-1],
figs= [filename_png], colorbar='legend.png',
kmzfile=filename_kmz, name='VIIRS Sea Surface Temperature')
fig = plt.figure(figsize=(1.0, 4.0), facecolor=None, frameon=False)
ax = fig.add_axes([0.0, 0.05, 0.2, 0.9])
cb = fig.colorbar(cs, cax=ax)
cb.set_label('SST [C]', rotation=-90, color='k', labelpad=20)
fig.savefig('legend.png', transparent=False, format='png') # Change transparent to True if your colorbar is not on space :)
# -
pixels = 1024 * 10
cmap = colorbrewer.get_map('Spectral', 'diverging', 11, reverse=True).mpl_colormap
fig, ax = gearth_fig(llcrnrlon=lon.min(),
llcrnrlat=lat.min(),
urcrnrlon=lon.max(),
urcrnrlat=lat.max(),
pixels=pixels)
cs = ax.pcolormesh(lon, lat, sst5x - 273.15, cmap=cmap, vmin = 10, vmax = 17)
ax.set_axis_off()
fig.show()
# +
from copy import copy
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.mlab as mlab
print(sst5x.shape)
print(sst4x.shape)
print(sst3x.shape)
palette = copy(plt.cm.jet)
palette.set_over('r', 1.0)
palette.set_under('g', 1.0)
palette.set_bad(alpha = 0.0)
fig, (ax1, ax2) = plt.subplots(nrows=2, figsize=(6, 5.4))
# plot using 'continuous' color map
im = ax1.imshow(sst5x - 273.15, interpolation='bilinear',
#im = ax1.imshow(sst4x[540:560,280:289] - 273.15, interpolation='bilinear',
cmap=palette,
norm=colors.Normalize(vmin=10.0, vmax=17.0),
aspect='auto',
origin='lower')
#extent=[x0, x1, y0, y1])
ax1.set_title('sst')
plt.show()
print(lon[283,546],lat[283,546])
# -
| sat_collocation/.ipynb_checkpoints/viirs_l2p_image-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Results with Merged Dataset
# #### Q5: For all of the models that were produced in 2008 that are still being produced now, how much has the mpg improved and which vehicle improved the most?
# Remember to use your new dataset, `combined_dataset.csv`. You should've created this data file in the previous section: *Merging Datasets*.
# load dataset
import pandas as pd
df = pd.read_csv('combined_dataset.csv')
# ### 1. Create a new dataframe, `model_mpg`, that contain the mean combined mpg values in 2008 and 2018 for each unique model
#
# To do this, group by `model` and find the mean `cmb_mpg_2008` and mean `cmb_mpg` for each.
model_mpg = df.groupby('model').mean()[['cmb_mpg_2008', 'cmb_mpg']]
model_mpg.head()
# ### 2. Create a new column, `mpg_change`, with the change in mpg
# Subtract the mean mpg in 2008 from that in 2018 to get the change in mpg
model_mpg['mpg_change'] = model_mpg['cmb_mpg'] - model_mpg['cmb_mpg_2008']
model_mpg.head()
# ### 3. Find the vehicle that improved the most
# Find the max mpg change, and then use query or indexing to see what model it is!
max_change = model_mpg['mpg_change'].max()
max_change
model_mpg[model_mpg['mpg_change'] == max_change]
idx = model_mpg.mpg_change.idxmax()
# pandas also has a useful function "idxmax" used to find the index of the row containing a columns maximum value
idx = model_mpg.mpg_change.idxmax()
idx
model_mpg.loc[idx]
| 2_Intro_to_data_analysis/Data_Analysis_Case_Study_2/results_merged.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The time series of a chemical concentration necessarily depends on its initial conditions; i.e. the concentration at time 0. An analogous statement is true for gating variables, etc. How do we specify this?
# <h2>Option 1: NEURON and NMODL defaults</h2>
# If the species corresponds to one with initial conditions specified by NMODL (or in the case of sodium, potassium, or calcium with meaningful NEURON defaults), then omitting the initial argument will tell NEURON to use those rules. e.g.
# +
from neuron import h, rxd
soma = h.Section(name='soma')
cyt = rxd.Region(h.allsec(), name='cyt', nrn_region='i')
ca = rxd.Species(cyt, name='ca', charge=2, atolscale=1e-6)
na = rxd.Species(cyt, name='na', charge=1)
k = rxd.Species(cyt, name='k', charge=1)
unknown = rxd.Species(cyt, name='unknown', charge=-1)
h.finitialize(-65)
print('ca: %g mM' % ca.nodes[0].concentration)
print('na: %g mM' % na.nodes[0].concentration)
print('k: %g mM' % k.nodes[0].concentration)
print('unknown: %g mM' % unknown.nodes[0].concentration)
# -
# As shown here, unknown ions/proteins are by default assigned a concentration by NEURON of 1 mM. The atolscale value for calcium has no effect on the initialized value, but is included here as an example of best practice for working with low concentrations.
# Importantly, the NEURON/NMODL rules only apply if there is a corresponding classical NEURON state variable. That is, <tt>nrn_region</tt> must be set and the Species must have a <tt>name</tt> assigned.
# Running what is otherwise the same code without the <tt>nrn_region</tt> assigned causes everything to default to 0 µM:
# +
from neuron import h, rxd
soma = h.Section(name='soma')
cyt = rxd.Region(h.allsec(), name='cyt')
ca = rxd.Species(cyt, name='ca', charge=2)
na = rxd.Species(cyt, name='na', charge=1)
k = rxd.Species(cyt, name='k', charge=1)
unknown = rxd.Species(cyt, name='unknown', charge=-1)
h.finitialize(-65)
print('ca: %g mM' % ca.nodes[0].concentration)
print('na: %g mM' % na.nodes[0].concentration)
print('k: %g mM' % k.nodes[0].concentration)
print('unknown: %g mM' % unknown.nodes[0].concentration)
# -
# For extracellular species, there is no equivalent traditional NEURON state variable (as those only exist within and along the cell), however NEURON's constant initialization parameters for the <tt>nrn_region='o'</tt> space are used if available; e.g.
# +
from neuron import h, crxd as rxd
# enabled by default in NEURON 7.7+
rxd.options.enable.extracellular = True
ecs = rxd.Extracellular(-100, -100, -100,
100, 100, 100,
dx=20, volume_fraction=0.2, tortuosity=1.6)
# defining calcium on both intra- and extracellular regions
ca = rxd.Species(ecs, name='ca', charge=2)
# global initialization for NEURON extracellular calcium
# in NEURON 7.7+ can just use: h.cao0_ca_ion = 0.42
# but in older versions of NEURON the variable would not have been defined yet
# and is thus unsettable except via a string call to h
h('cao0_ca_ion = 0.42')
h.finitialize(-65)
print('ca: %g mM' % ca.nodes[0].concentration)
# -
# We could do something similar using <tt>cai0_ca_ion</tt> to set the global initial intracellular calcium concentration.
# <h2>Option 2: Uniform initial concentration</tt></h2>
# Setting <tt>initial=</tt> to a Species or State assigns that value every time the system reinitializes. e.g.
# +
from neuron import h, rxd
soma = h.Section(name='soma')
cyt = rxd.Region([soma], name='cyt')
m = rxd.State(cyt, initial=0.47)
h.finitialize(-65)
print('m = %g' % m.nodes[0].value)
# -
# <h2>Option 3: Initializing to a function of position</h2>
# The <tt>initial=</tt> keyword argument also accepts a callable (e.g. a function) that receives a node object. Nodes have certain properties that are useful for assinging based on position, including <tt>.segment</tt> (intracellular nodes only) and <tt>.x3d</tt>, <tt>.y3d</tt>, and <tt>.z3d</tt>:
# <h3>Using <tt>.segment</tt>:</h3>
# Here we use the morphology <a href="http://neuromorpho.org/dableFiles/amaral/CNG%20version/c91662.CNG.swc">c91662.swc</a> from NeuroMorpho.Org and initialize based on <i>path distance</i> from the soma.
# +
from neuron import h, gui, rxd
h.load_file('stdrun.hoc')
h.load_file('import3d.hoc')
# load the morphology and instantiate at the top level (i.e. not in a class)
cell = h.Import3d_SWC_read()
cell.input('c91662.swc')
h.Import3d_GUI(cell, 0)
i3d = h.Import3d_GUI(cell, 0)
i3d.instantiate(None) # pass in a class to instantiate inside the class instead
# increase the number of segments
for sec in h.allsec():
sec.nseg = 1 + 2 * int(sec.L / 20)
def my_initial(node):
# set a reference point
h.distance(0, h.soma[0](0.5))
# compute the distance
distance = h.distance(node.segment)
# return a certain function of the distance
return 2 * h.tanh(distance / 1000.)
cyt = rxd.Region(h.allsec(), name='cyt', nrn_region='i')
ca = rxd.Species(cyt, name='ca', charge=2, initial=my_initial)
h.finitialize(-65)
# -
# <center>
# <img style="height:20em" src="http://neuron.yale.edu/neuron/static/tutorials/images/shapeplot-distance.png"/>
# </center>
# <h3>Using position:</h3>
# We continue the above example adding a new species, that is initialized based on the x-coordinate. This could happen, for example, on a platform with a nutrient or temperature gradient:
# +
def my_initial2(node):
# return a certain function of the x-coordinate
return 1 + h.tanh(node.x3d / 100.)
alpha = rxd.Parameter(cyt, name='alpha', initial=my_initial2)
h.finitialize(-65)
# -
# <center>
# <img style="height:20em" src="http://neuron.yale.edu/neuron/static/tutorials/images/shapeplot-x3d.png"/>
# </center>
#
# <h2>Option 4: to steady state</h2>
# Sometimes one might want to initialize a simulation to steady-state where e.g. diffusion, ion channel currents, and chemical reactions all balance each other out. There may be no such possible initial condition due to the interacting parts.
#
# In principle, such initial conditions could be assigned using a variant of the option 3 approach above. In practice, however, it may be simpler to omit the <tt>initial=</tt> keyword argument, and use an <tt><a href="https://www.neuron.yale.edu/neuron/static/py_doc/simctrl/programmatic.html#FInitializeHandler">h.FInitializeHandler</a></tt> to loop over locations, setting the values for all states at a given location at the same time. A full example is beyond the scope of this tutorial.
| docs/rxd-tutorials/initialization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3-ecpaperenv]
# language: python
# name: conda-env-miniconda3-ecpaperenv-py
# ---
# +
import xarray as xr
import sys
import pandas as pd
import numpy as np
import xesmf as xe
import warnings
import dask
import datetime
warnings.filterwarnings('ignore')
# +
# get the workers going
#ncores = 36
#nmem = str(int(365*ncores/36))+'GB'
#from dask_jobqueue import SLURMCluster
#from dask.distributed import Client
#cluster = SLURMCluster(cores=ncores,
# processes=ncores, memory=nmem,
# project='P04010022',
# walltime='12:00:00')
#cluster.scale(ncores)
#client = Client(cluster)
# -
# get the workers going
ncores = 1
nworkers = 12
nmems="5GB"
from dask.distributed import Client
from dask_jobqueue import SLURMCluster
cluster = SLURMCluster(
cores = ncores, processes = ncores, project="P04010022", walltime="04:00:00")
cluster.scale(nworkers)
client = Client(cluster)
# do this until you see you've got some workers
client
# location of ERA5 data on RDA
filepath="/gpfs/fs1/collections/rda/data/ds633.0/e5.oper.fc.sfc.accumu/"
# output location
outpath="/glade/scratch/islas/processed/era5/STRD_day/"
ystart=1979 ; yend=2019 ; nyears=yend-ystart+1
# open up CESM data to get the output grid.
cesmdat = xr.open_dataset("/glade/campaign/cesm/collections/cesmLE/CESM-CAM5-BGC-LE/atm/proc/tseries/monthly/PHIS/f.e11.F1850C5CNTVSST.f09_f09.002.cam.h0.PHIS.040101-050012.nc")
grid_out = xr.Dataset({'lat': (['lat'], cesmdat.lat)}, {'lon': (['lon'], cesmdat.lon)})
dailydat2.STRD.data
reusewgt=False
wgtfile=outpath+"wgtfile.nc"
for iyear in range(ystart,yend+1,1):
print(iyear)
timeout = pd.date_range(start = str(iyear)+"-01-01", end = str(iyear)+"-12-31")
countdays=0
for imon in range(1,12+1,1):
#for imon in range(2,3,1):
monstr=str(imon).zfill(2)
outfile=outpath+"strd_"+str(iyear)+monstr+".nc"
file=filepath+"/"+str(iyear)+monstr+"/*_str.*.nc"
print(file)
data = xr.open_mfdataset(file,coords="minimal", join="override",
decode_times=True, use_cftime=True )
data = data.STR
data6h = data.loc[{"forecast_initial_time": data.forecast_initial_time.dt.hour.isin([6])}]
data18h = data.loc[{"forecast_initial_time": data.forecast_initial_time.dt.hour.isin([18])}]
middle = data6h.sum(dim="forecast_hour")
end = data18h.loc[dict(forecast_hour=slice(1,6))].sum(dim="forecast_hour")
beginning = data18h.loc[dict(forecast_hour=slice(7,13))].sum(dim="forecast_hour")
dayspermon = middle.forecast_initial_time.dt.daysinmonth.data
dayendstr = str(dayspermon[0])
timeout = pd.date_range(
start=str(iyear)+"-"+monstr+"-01",
end=str(iyear)+"-"+monstr+"-"+dayendstr)
timeout2 = timeout + datetime.timedelta(days=1)
beginning['forecast_initial_time'] = timeout2
middle['forecast_initial_time'] = timeout
end['forecast_initial_time'] = timeout
dailydat = (beginning + middle + end)/86400.
if ((imon == 1) and (iyear == ystart)):
day1dat = (middle.isel(forecast_initial_time=0) + end.isel(forecast_initial_time=0))/64800.
else:
day1dat = (begsaved + middle.isel(forecast_initial_time=0) + end.isel(forecast_initial_time=0))/86400.
dailydat2 = xr.concat([day1dat, dailydat], dim="forecast_initial_time")
begsaved = beginning.isel(forecast_initial_time=dayspermon[0]-1).persist()
#dailydat2 = dailydat2.rename(longitude="lon")
#dailydat2 = dailydat2.rename(latitude="lat")
#dailydat2 = dailydat2.chunk({'forecast_initial_time': -1})
dailydat2 = dailydat2.transpose("forecast_initial_time","latitude","longitude")
regridder = xe.Regridder(dailydat2, grid_out, 'bilinear', periodic=True, reuse_weights=reusewgt, filename=wgtfile)
reusewgt=True
dataday_rg = regridder(dailydat2)
dataday_rg = dataday_rg.rename(forecast_initial_time="time")
dataday_rg = dataday_rg.assign_attrs({'long_name':'Surface net thermal longwave radiation', 'units':'Wm**2'})
dataday_rg.to_netcdf(path=outfile)
| examples/grabera5strd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title"><b>The Knapsack Problem</b></span> by <a xmlns:cc="http://creativecommons.org/ns#" href="http://mate.unipv.it/gualandi" property="cc:attributionName" rel="cc:attributionURL"><NAME></a> is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>.<br />Based on a work at <a xmlns:dct="http://purl.org/dc/terms/" href="https://github.com/mathcoding/opt4ds" rel="dct:source">https://github.com/mathcoding/opt4ds</a>.
# # 3. The Knapsack Problem
# In this first notebook, we show how to solve the Knapsack Problem using Integer Linear Programming.
# ## 3.1 Software Installation
# If you are running this notebook in a Colab, you don't need to install anything else on your computer.
#
# Otherwise, if you have installed the recommended Anaconda Python distribution, you have to run the following two commands:
#
# 1. To install the [Pyomo](http://www.pyomo.org/) optimization modeling language:
#
# ```
# conda install -c conda-forge pyomo
# ```
#
# 2. To install the open source [GLPK](https://www.gnu.org/software/glpk/) solver:
#
# ```
# conda install -c conda-forge glpk
# ```
#
# 3. (Optional) You can install some extra packages of Pyomo using the following command:
#
# ```
# conda install -c conda-forge pyomo.extras
# ```
#
# For details about the Pyomo installation, we refer to the official [Pyomo Documentation](https://pyomo.readthedocs.io/en/stable/).
# The following lines are for running this notebook in a COLAB:
# +
import shutil
import sys
import os.path
if not shutil.which("pyomo"):
# !pip install -q pyomo
assert(shutil.which("pyomo"))
if not (shutil.which("glpk") or os.path.isfile("glpk")):
if "google.colab" in sys.modules:
# !apt-get install -y -qq glpk-utils
else:
try:
# !conda install -c conda-forge glpk
except:
pass
# -
# ## 3.2 Mixed Integer Programming model
# The Knapsack problem can be formulated as follows.
#
# The input data are:
#
# * The index set $I$ referring to the items
# * The profit vector $c$
# * The weight vector $A$
# * The budget value $B$
#
# For each item $i\in I$, we introduce a binary decision variables $x_i \in \{0,1\}$, which is used to define the following **Integer Linear Programming (ILP)**
# problem:
#
# \begin{align}\label{eq:1}
# \max \;\; & c^T x \\
# \mbox{s.t.} \;\; & \sum_{i \in I} A_i x_i \leq B & \\
# & x_i \in \{0,1\},& \forall i \in I.
# \end{align}
#
# Since $x_i=1$ represents the decision of selecting item the $i$-th item, it is clear that the objective function (1) consists of maximizing the dot product $c^T\,x$. The single constraint (2) limits the number of selected item in such a way that the sum of the weights $A_i$ of the selected items does not exceed the available capacity $B$. The constraints (3) impose the domain on the decision variables $x_i$.
# ## 3.3 Pyomo Knapsack Model
# The ILP model (1)-(3) can be expressed using the Pyomo optimization modeling language as shown next.
#
# As a first step, we need to define the input data using the standard Python data structure. The simplest method to define the data is as follows:
I = range(5) # Items
C = [2, 3, 1, 4, 3] # Profits
A = [3, 4, 2, 1, 6] # Weights
B = 9 # Budget
# This snippet of code is defining the following input data:
#
# * The set $I$ as the range of number from 0 to 4, using the [range](https://docs.python.org/3/library/functions.html#func-range) builtin class. You can think of the range as a generator function for the list $[0,1,2,3,4]$.
# * The cost vector $C$ and the weight vector $A$ are defined as two lists, using the standard [list()](https://docs.python.org/3/library/functions.html#func-list) builtin class.
# * The budget parameter is defined as a given integer.
#
# So far, we have not used any **Pyomo** construct. Before we start, we need to import the Pyomo library:
from pyomo.environ import *
# +
# If you want to check all the elements imported from the library uncomment the following line
# who
# -
# ### 3.3.1 Model and Variables
# The first step in defining the model consists in choosing the type of Model we want to use. In Pyomo there are two options, the [ConcreteModel](https://pyomo.readthedocs.io/en/stable/library_reference/aml/index.html#pyomo.environ.ConcreteModel) and the [AbstractModel](https://pyomo.readthedocs.io/en/stable/library_reference/aml/index.html#pyomo.environ.AbstractModel). In this first example, we use the simpler ConcreteModel as follows:
# Create concrete model
model = ConcreteModel()
# The choice of the `model` name is recommended if you are planning to use the Pyomo command line tool.
#
# Once we have defined the model, we can define the binary decision variable by using the [Var](https://pyomo.readthedocs.io/en/stable/library_reference/aml/index.html#pyomo.environ.Var) class. We define an object of type **Var** for each element of the range $I$, with the following command:
# Variables
model.x = Var(I, within=Binary)
# The `Binary` keyword is part of the `pyomo.environ` setting, and it is used to specify the constraint $x_i \in \{0,1\}$. Other possible values for the optional parameter `within` are: `NonNegativeReals`, `PositiveReals`, `PositiveIntegers`.
#
# Note the the choice of $x$ as a name of the variable is arbitrary.
# ### 3.3.2 Objective Function
# The objective function is defined via the [Objective](https://pyomo.readthedocs.io/en/stable/library_reference/aml/index.html#pyomo.environ.Objective.construct) class, as follows:
# Objective Function: Maximize Profit
model.obj = Objective(expr = sum(C[i]*model.x[i] for i in I),
sense = maximize)
# Again, the `obj` name is arbitrary, and you can select the one you prefer. The parameter `epxr` is mandatory, and is used to define the objective function expression. In the following example, we are using the `list comprehension` syntax to define our linear objective $\sum_{i \in I} c_i x_i$. Note that the python notation is very similar to the mathematical notation.
#
# The parameter `sense` is optional, and it is used to define the type of objective function: `maximize` or `minimize`.
#
# With Pyomo, we are not directly restricted to use only linear objective functions. It is the type solver we use that limits that type of problem that we can solve. As long as we use the GLPK solver, we can only define linear objective function.
# ### 3.3.3 Constraints
# Finally, we need to define the budget constraint.
# The constraints are defined using the [Constraint](https://pyomo.readthedocs.io/en/stable/library_reference/aml/index.html#pyomo.environ.Constraint) class. The minimal use of this class requires to define the `expr` input parameter. In the knapsack problem we have to define the budget constraint $\sum_{i \in I} A_i x_i \leq B$ as follows:
# Constraint
model.capacity = Constraint(expr = sum(A[i]*model.x[i] for i in I) <= B)
# Also the constraints can be named, in this case, we named it `capacity`. The name of the constraint can be used to retrieve information about the status of the constraint in a solution, that is, given a solution $\bar x$ to check whether $\sum_{i \in I} A_i \bar x_i < B$ or $\sum_{i \in I} A_i \bar x_i = B$.
# ### 3.3.4 Solving the Pyomo model
# The complete Pyomo model defined so far is as follows:
# Create concrete model
model = ConcreteModel()
# Variables
model.x = Var(I, within=Binary)
# Objective Function: Maximize Profit
model.obj = Objective(expr = sum(C[i]*model.x[i] for i in I),
sense = maximize)
# Constraint
model.capacity = Constraint(expr = sum(A[i]*model.x[i] for i in I) <= B)
# In order to solve this model, we need to use a **solver**, that is a software that binds the data to the model and solve the corresponding instance of the problem. In this notebook, we use the GLPK solver, using a `SolverFactory` as follows:
# Solve the model
sol = SolverFactory('glpk').solve(model)
# Basic info about the solution process
for info in sol['Solver']:
print(info)
# Finally, in order to check the values of the decision variables, we can query the solved model by using the variable, objective function, and constraints names:
# Report solution value
print("Optimal solution value: z =", model.obj())
print("Decision variables:")
for i in I:
print("x_{} = {}".format(i, model.x[i]()))
print("Capacity left in the knapsack:", B-model.capacity())
# In this case, we have found a solution with value equal to 9, given by selecting the three items $[0, 1, 3]$. We still had a unit of capacity left in the knapsack, but since no left items as a weight equal to 1, and we cannot take fractional items, that capacity stay unused.
# ## 3.4 Complete Python Script
# The whole Python script for solving the Knapsack problem is as follows:
# +
# Import the libraries
from pyomo.environ import ConcreteModel, Var, Objective, Constraint, SolverFactory
from pyomo.environ import maximize, Binary
# CONCRETE MODEL: Data First, then model
I = range(5) # Items
C = [2, 3, 1, 4, 3] # Profits
A = [3, 4, 2, 1, 6] # Weights
B = 9 # Budget
# Create concrete model
model = ConcreteModel()
# Variables
model.x = Var(I, within=Binary)
# Objective Function: Maximize Profit
model.obj = Objective(expr = sum(C[i]*model.x[i] for i in I),
sense = maximize)
# Constraint
model.capacity = Constraint(expr = sum(A[i]*model.x[i] for i in I) <= B)
# Solve the model
sol = SolverFactory('glpk').solve(model)
# Basic info about the solution process
for info in sol['Solver']:
print(info)
# Report solution value
print("Optimal solution value: z =", model.obj())
print("Decision variables:")
for i in I:
print("x_{} = {}".format(i, model.x[i]()))
print("Capacity left in the knapsack:", B-model.capacity())
| KnapsackProblem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 2) Training Decoders in Practice
#
# Now that we have discussed the conceptual foundations, strategies and techniques involved, we will provide detailed examples of how train decoders via the procedures discussed. In particular, in this notebook we will walk through a very simple script for training a decoder with a given set of hyper-parameters, providing the foundation for a later discussion concerning how to obtain optimal decoders for a range of error rates through an iterative training procedure involving a hyper-parameter optimization for each error rate (see the companion notebook "Large Scale Iterative Training").
#
# ##### 2a) Requirements
#
# The following packages are required, and can be installed via PIP:
#
# <ol>
# <li> Python 3 (with numpy and scipy)</li>
# <li> tensorflow </li>
# <li> keras </li>
# <li> gym </li>
# </ol>
#
# In addition, a modified version of the Keras-RL package is required, which should be installed from <a href="https://github.com/R-Sweke/keras-rl">this fork</a>
# ##### 2b) A Simple Training Script
#
# We begin by importing all required packages and methods:
# +
import numpy as np
import keras
import tensorflow
import gym
from Function_Library import *
from Environments import *
import rl as rl
from rl.agents.dqn import DQNAgent
from rl.policy import BoltzmannQPolicy, EpsGreedyQPolicy, LinearAnnealedPolicy, GreedyQPolicy
from rl.memory import SequentialMemory
from rl.callbacks import FileLogger
import json
import copy
import sys
import os
import shutil
import datetime
import pickle
# -
# We then proceed by providing all required hyperparameters and physical configuration settings. In order to allow for easier grid searching and incremented training later on we choose to split all hyperparameters into two categories:
#
# - fixed configs: These remain constant during the course of a grid search or incremented training procedure.
# - variable configs: We will later set up training grids over these hyperparameters.
#
# In particular, the fixed parameters one must provide are:
#
# 1. **d**: The lattice width (equal to the lattice height)
# - **use_Y**: If true then the agent can perform Y Pauli flips directly, if False then the agent can only perform X and Z Pauli flips.
# - **train_freq**: The number of agent-environment interaction steps which occur between each updating of the agent's weights.
# - **batch_size**: The size of batches used for calculating loss functions for gradient descent updates of agent weights.
# - **print_freq**: Every print_freq episodes the statistics of the training procedure will be logged.
# - **rolling_average_length**: The number of most recent episodes over which any relevant rolling average will be calculated.
# - **stopping_patience**: The number of episodes after which no improvement will result in the early stopping of the training procedure.
# - **error_model**: A string in ["X", "DP"], specifiying the noise model of the environment as X flips only or depolarizing noise.
# - **c_layers**: A list of lists specifying the structure of the convolutional layers of the agent deepQ network. Each inner list describes a layer and has the form [num_filters, filter_width, stride].
# - **ff_layers**: A list of lists specifying the structure of the feed-forward neural network sitting on top of the convolutional neural network. Each inner list has the form [num_neurons, output_dropout_rate].
# - **max_timesteps**: The maximum number of training timesteps allowed.
# - **volume_depth**: The number of syndrome measurements taken each time a new syndrome extraction is performed - i.e. the depth of the syndrome volume passed to the agent.
# - **testing_length**: The number of episodes uses to evaluate the trained agents performance.
# - **buffer_size**: The maximum number of experience tuples held in the memory from which the update batches for agent updating are drawn.
# - **dueling**: A boolean indicating whether or not a [dueling architecture](https://arxiv.org/abs/1511.06581) should be used.
# - **masked_greedy**: A boolean which indicates whether the agent will only be allowed to choose legal actions (actions next to an anyon or previously flipped qubit) when acting greedily (i.e. when choosing actions via the argmax of the Q-values)
# - **static_decoder**: For training within the fault tolerant setting (multi-cycle decoding) this should always be set to True.
#
# In addition, the parameters which we will later incrementally vary or grid search around are:
#
# 1. **p_phys**: The physical error probability
# 2. **p_meas**: The measurement error probability
# 3. **success_threshold**: The qubit lifetime rolling average at which training has been deemed succesfull and will be stopped.
# 4. **learning_starts**: The number of initial steps taken to contribute experience tuples to memory before any weight updates are made.
# 5. **learning_rate**: The learning rate for gradient descent optimization (via the Adam optimizer)
# 6. **exploration_fraction**: The number of time steps over which epsilon, the parameter controlling the probability of a random explorative action, is annealed.
# 7. **max_eps**: The initial maximum value of epsilon.
# 8. **target_network_update_freq**: In order to achieve stable training, a target network is cloned off from the active deepQ agent every target_network_update_freq interval of steps. This target network is then used to generate the target Q-function over the following interval.
# 9. **gamma**: The discount rate used for calculating the expected discounted cumulative return (the Q-values).
# 10. **final_eps**: The final value at which annealing of epsilon will be stopped.
#
# Furthermore, in addition to all the above parameters one must provide a directory into which results and training progress as logged, as well as the path to a pre-trained referee decoder. Here e provide two pre-trained feed forward classification based referee decoders, one for X noise and one for DP noise. However, in principle any perfect-measurement decoding algorithm (such as MWPM) could be used here.
# +
fixed_configs = {"d": 5,
"use_Y": False,
"train_freq": 1,
"batch_size": 32,
"print_freq": 250,
"rolling_average_length": 500,
"stopping_patience": 500,
"error_model": "X",
"c_layers": [[64,3,2],[32,2,1],[32,2,1]],
"ff_layers": [[512,0.2]],
"max_timesteps": 1000000,
"volume_depth": 5,
"testing_length": 101,
"buffer_size": 50000,
"dueling": True,
"masked_greedy": False,
"static_decoder": True}
variable_configs = {"p_phys": 0.001,
"p_meas": 0.001,
"success_threshold": 10000,
"learning_starts": 1000,
"learning_rate": 0.00001,
"exploration_fraction": 100000,
"max_eps": 1.0,
"target_network_update_freq": 5000,
"gamma": 0.99,
"final_eps": 0.02}
logging_directory = os.path.join(os.getcwd(),"logging_directory/")
static_decoder_path = os.path.join(os.getcwd(),"referee_decoders/nn_d5_X_p5")
all_configs = {}
for key in fixed_configs.keys():
all_configs[key] = fixed_configs[key]
for key in variable_configs.keys():
all_configs[key] = variable_configs[key]
static_decoder = load_model(static_decoder_path)
logging_path = os.path.join(logging_directory,"training_history.json")
logging_callback = FileLogger(filepath = logging_path,interval = all_configs["print_freq"])
# -
# Now that we have specified all the required parameters we can instantiate our environment:
env = Surface_Code_Environment_Multi_Decoding_Cycles(d=all_configs["d"],
p_phys=all_configs["p_phys"],
p_meas=all_configs["p_meas"],
error_model=all_configs["error_model"],
use_Y=all_configs["use_Y"],
volume_depth=all_configs["volume_depth"],
static_decoder=static_decoder)
# The environment class is defined to mirror the environments of [https://gym.openai.com/](openAI gym), and such contains the required "reset" and "step" methods, via which the agent can interact with the environment, in addition to decoding specific methods and attributes whose details can be found in the relevant method docstrings.
# We can now proceed to define the agent. We being by specifying the memory to be used, as well as the exploration and testing policies.
# +
memory = SequentialMemory(limit=all_configs["buffer_size"], window_length=1)
policy = LinearAnnealedPolicy(EpsGreedyQPolicy(masked_greedy=all_configs["masked_greedy"]),
attr='eps', value_max=all_configs["max_eps"],
value_min=all_configs["final_eps"],
value_test=0.0,
nb_steps=all_configs["exploration_fraction"])
test_policy = GreedyQPolicy(masked_greedy=True)
# -
# Finally, we can then build the deep convolutional neural network which will represent our Q-function and compile our agent.
# +
model = build_convolutional_nn(all_configs["c_layers"],
all_configs["ff_layers"],
env.observation_space.shape,
env.num_actions)
dqn = DQNAgent(model=model,
nb_actions=env.num_actions,
memory=memory,
nb_steps_warmup=all_configs["learning_starts"],
target_model_update=all_configs["target_network_update_freq"],
policy=policy,
test_policy = test_policy,
gamma = all_configs["gamma"],
enable_dueling_network=all_configs["dueling"])
dqn.compile(Adam(lr=all_configs["learning_rate"]))
# -
# With both the agent and the environment specified, it is then possible to train the agent by calling the agent's "fit" method. If you want to run this on a single computer, be careful, it may take up to 12 hours!
# +
now = datetime.datetime.now()
started_file = os.path.join(logging_directory,"started_at.p")
pickle.dump(now, open(started_file, "wb" ) )
history = dqn.fit(env,
nb_steps=all_configs["max_timesteps"],
action_repetition=1,
callbacks=[logging_callback],
verbose=2,
visualize=False,
nb_max_start_steps=0,
start_step_policy=None,
log_interval=all_configs["print_freq"],
nb_max_episode_steps=None,
episode_averaging_length=all_configs["rolling_average_length"],
success_threshold=all_configs["success_threshold"],
stopping_patience=all_configs["stopping_patience"],
min_nb_steps=all_configs["exploration_fraction"],
single_cycle=False)
# -
# As you can see above, during the training procedure various statistics are logged, both to stdout and to file in the specified directory. As you may notice above we manually stopped training after approximately 7000 seconds while the agent was still improving, and before it has reached the specified success threshold.
#
# In order to evaluate the agent later on, or apply the agent in a production decoding scenario we can easily save the weights:
weights_file = os.path.join(logging_directory, "dqn_weights.h5f")
dqn.save_weights(weights_file, overwrite=True)
# And finally, in order to evaluate the training procedure we may be interested in viewing any of the metrics which were logged. These are all saved within the history.history dictionary. For example, we are often most interested in analyzing the training procedure by looking at the rolling average of the qubit lifetime, which we can do as follows:
# +
from matplotlib import pyplot as plt
# %matplotlib inline
training_history = history.history["episode_lifetimes_rolling_avg"]
plt.figure(figsize=(12,7))
plt.plot(training_history)
plt.xlabel('Episode')
plt.ylabel('Rolling Average Qubit Lifetime')
_ = plt.title("Training History")
# -
# From the above plot one can see that during the exploration phase the agent was unable to do well, due to constant exploratory random actions, but was able to exploit this knowledge effectively once the exploration probability became sufficiently low. Again, it is also clear that the agent was definitely still learning and improving when we chose to stop the training procedure.
| example_notebooks/2) Training Example.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
function Lgauss2(A,B,Σ,R,T,x0)
# x' = Ax + N(0,Σ) A :dx×dx
# y' = Bx'+ N(0,R) B :dy⨱dx
x = zeros(2,T+1)
y = zeros(2,T)
x[:,1] = x0
for i in 1:T
x[:,i+1] = A*x[:,i] + rand(MvNormal(Σ))
y[:,i] = B*x[:,i+1] + rand(MvNormal(R))
end
return(x[:,2:T+1],y)
end
function KFfilter2(A,B,Σ,R,data,Q0,m0)
# x' = Ax + N(0,Σ) A :dx×dx
# y' = Bx'+ N(0,R) B :dy⨱dx
n = length(data[1,:])
Qc = Q0
mc = m0
fm = []
fQ = []
pm = []
pQ = []
for i in 1:n
y = data[:,i]
predmean = A*mc
predvar = A*Qc*A'+Σ
push!(pm,predmean)
push!(pQ,predvar)
E = A*Qc*A' + Σ
Qn = E*(I+zeros(2,2) - B'*inv(B*E*B' + R)*B*E)
mn = (I+zeros(2,2) - E*B'inv(B*E*B'+R)*B)*A*mc + E*B'*inv(B*E*B'+R)*y
push!(fm,mn)
push!(fQ,Qn)
Qc = copy(Qn)
mc = copy(mn)
end
return(fm,fQ,pm,pQ)
end
# +
using Distributions
using LinearAlgebra
using Plots
using StatsPlots
using StatsBase
theme(:ggplot2)
#set paramaters of the model
A = [0.5 0.4;0.6 0.3]
B = I + zeros(2, 2)
P = [0.9 0.3;0.3 0.9]
Σ = 0.3.*P
R = 0.5.*P
T = 100
x0 = rand(MvNormal(P))
#simulate from LGmodel
x,y = Lgauss2(A,B,Σ,R,T,x0)
# run KF
Q0 = P
m0 = x0
fm,fQ,pm,pQ = KFfilter2(A,B,Σ,R,y,Q0,m0)
#calculate prior, filtering density and predictive density at time t,t-1
x_grid = range(-3.0, 3.0, length = 100)
y_grid = range(-3.0, 3.0, length = 100)
priordist = MvNormal(P)
filterdist = MvNormal(fm[100],fQ[100])
predictiondist = MvNormal(pm[99],pQ[99])
z1 = [pdf(priordist, [i,j]) for i in y_grid, j in x_grid]
z2 = [pdf(predictiondist, [i,j]) for i in y_grid, j in x_grid]
z3 = [pdf(filterdist, [i,j]) for i in y_grid, j in x_grid]
a=contour(x_grid,y_grid, z1,fill = false,color = :viridis, cbar = false,title="Prior")
contour(x_grid,y_grid, z2,fill = false,color = :viridis, cbar = false,title="Predictive density")
b=annotate!(x[1,100], x[2,100], "x", color = :black)
contour(x_grid,y_grid, z3,fill = false,color = :viridis, cbar = false,title="Filtering density")
c=annotate!(x[1,100], x[2,100], "x", color = :black)
plot(a,b,c,layout=(1,3))
#savefig("kalman.png")
# +
#estimation
est_state1 = zeros(100,100)
est_state2 = zeros(100,100)
CIest1 = zeros(100)
CIest2 = zeros(100)
for i in 1:100
est = rand(MvNormal(fm[i],Symmetric(fQ[i])),100)
est_state1[i,:] = est[1,:]
est_state2[i,:] = est[2,:]
CIest1[i] = percentile(est_state1[i,:],97.5) - percentile(est_state1[i,:],2.5)
CIest2[i] = percentile(est_state2[i,:],97.5) - percentile(est_state2[i,:],2.5)
end
est1 = mean(est_state1, dims=2)
est2 = mean(est_state2, dims=2)
plot(est1,ribbon=CIest1,fillalpha=.5,label="Filter",foreground_color_legend = nothing,legend=:topleft)
a=plot!(x[1,:],label="x1")
plot(est2,ribbon=CIest2,fillalpha=.5,label="Filter",foreground_color_legend = nothing,legend=:topleft)
b=plot!(x[2,:],label="x2")
plot(a,b,layout=(1,2))
#savefig("kalman2.png")
# -
| Kalman Filter/Kalman_filter_prac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def lengthOfLIS(self, nums: List[int]) -> int:
if not nums:
return 0
dp = []
for i in range(len(nums)):
dp.append(1)
for j in range(i):
if nums[j]< nums[i]:
dp[i] = max(dp[i],dp[j]+1)
return max(dp)
| medium/longest increasing sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # *Quick, Draw!* GAN
# * code based directly on [Grant Beyleveld's](https://github.com/grantbey/quickdraw-GAN/blob/master/octopus-v1.0.ipynb), which is derived from [<NAME>'s](https://towardsdatascience.com/gan-by-example-using-keras-on-tensorflow-backend-1a6d515a60d0) under [MIT License](https://github.com/roatienza/Deep-Learning-Experiments/blob/master/LICENSE)
# * data provided by [Google](https://github.com/googlecreativelab/quickdraw-dataset) under [Creative Commons Attribution 4.0 license](https://creativecommons.org/licenses/by/4.0/)
# #### Select processing devices
# +
# import os
# os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
# # os.environ["CUDA_VISIBLE_DEVICES"] = ""
# os.environ["CUDA_VISIBLE_DEVICES"] = "1"
# -
# #### Load dependencies
# +
# for data input and output:
import numpy as np
import os
# for deep learning:
import keras
from keras.models import Sequential, Model
from keras.layers import Input, Dense, Conv2D, BatchNormalization, Dropout, Flatten
from keras.layers import Activation, Reshape, Conv2DTranspose, UpSampling2D # new!
from keras.optimizers import RMSprop
# for plotting:
import pandas as pd
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# #### Load data
# NumPy bitmap files are [here](https://console.cloud.google.com/storage/browser/quickdraw_dataset/full/numpy_bitmap) -- pick your own drawing category -- you don't have to pick *apples* :)
input_images = "quickdraw/apple.npy"
data = np.load(input_images) # 28x28 (sound familiar?) grayscale bitmap in numpy .npy format; images are centered
data.shape
data[4242]
data = data/255
data = np.reshape(data,(data.shape[0],28,28,1)) # fourth dimension is color
img_w,img_h = data.shape[1:3]
data.shape
data[4242]
plt.imshow(data[4242,:,:,0], cmap='Greys')
# #### Create discriminator network
def discriminator_builder(depth=64,p=0.4):
# Define inputs
inputs = Input((img_w,img_h,1))
# Convolutional layers
conv1 = Conv2D(depth*1, 5, strides=2, padding='same', activation='relu')(inputs)
conv1 = Dropout(p)(conv1)
conv2 = Conv2D(depth*2, 5, strides=2, padding='same', activation='relu')(conv1)
conv2 = Dropout(p)(conv2)
conv3 = Conv2D(depth*4, 5, strides=2, padding='same', activation='relu')(conv2)
conv3 = Dropout(p)(conv3)
conv4 = Conv2D(depth*8, 5, strides=1, padding='same', activation='relu')(conv3)
conv4 = Flatten()(Dropout(p)(conv4))
# Output layer
output = Dense(1, activation='sigmoid')(conv4)
# Model definition
model = Model(inputs=inputs, outputs=output)
model.summary()
return model
discriminator = discriminator_builder()
discriminator.compile(loss='binary_crossentropy',
optimizer=RMSprop(lr=0.0008, decay=6e-8, clipvalue=1.0),
metrics=['accuracy'])
# #### Create generator network
def generator_builder(z_dim=100,depth=64,p=0.4):
# Define inputs
inputs = Input((z_dim,))
# First dense layer
dense1 = Dense(7*7*64)(inputs)
dense1 = BatchNormalization(momentum=0.9)(dense1) # default momentum for moving average is 0.99
dense1 = Activation(activation='relu')(dense1)
dense1 = Reshape((7,7,64))(dense1)
dense1 = Dropout(p)(dense1)
# De-Convolutional layers
conv1 = UpSampling2D()(dense1)
conv1 = Conv2DTranspose(int(depth/2), kernel_size=5, padding='same', activation=None,)(conv1)
conv1 = BatchNormalization(momentum=0.9)(conv1)
conv1 = Activation(activation='relu')(conv1)
conv2 = UpSampling2D()(conv1)
conv2 = Conv2DTranspose(int(depth/4), kernel_size=5, padding='same', activation=None,)(conv2)
conv2 = BatchNormalization(momentum=0.9)(conv2)
conv2 = Activation(activation='relu')(conv2)
conv3 = Conv2DTranspose(int(depth/8), kernel_size=5, padding='same', activation=None,)(conv2)
conv3 = BatchNormalization(momentum=0.9)(conv3)
conv3 = Activation(activation='relu')(conv3)
# Output layer
output = Conv2D(1, kernel_size=5, padding='same', activation='sigmoid')(conv3)
# Model definition
model = Model(inputs=inputs, outputs=output)
model.summary()
return model
generator = generator_builder()
# #### Create adversarial network
def adversarial_builder(z_dim=100):
model = Sequential()
model.add(generator)
model.add(discriminator)
model.compile(loss='binary_crossentropy',
optimizer=RMSprop(lr=0.0004, decay=3e-8, clipvalue=1.0),
metrics=['accuracy'])
model.summary()
return model
adversarial_model = adversarial_builder()
# #### Train!
def make_trainable(net, val):
net.trainable = val
for l in net.layers:
l.trainable = val
# +
def train(epochs=2000,batch=128):
d_metrics = []
a_metrics = []
running_d_loss = 0
running_d_acc = 0
running_a_loss = 0
running_a_acc = 0
for i in range(epochs):
if i%100 == 0:
print(i)
real_imgs = np.reshape(data[np.random.choice(data.shape[0],batch,replace=False)],(batch,28,28,1))
fake_imgs = generator.predict(np.random.uniform(-1.0, 1.0, size=[batch, 100]))
x = np.concatenate((real_imgs,fake_imgs))
y = np.ones([2*batch,1])
y[batch:,:] = 0
# make_trainable(discriminator, True)
d_metrics.append(discriminator.train_on_batch(x,y))
running_d_loss += d_metrics[-1][0]
running_d_acc += d_metrics[-1][1]
# make_trainable(discriminator, False)
noise = np.random.uniform(-1.0, 1.0, size=[batch, 100])
y = np.ones([batch,1])
a_metrics.append(adversarial_model.train_on_batch(noise,y))
running_a_loss += a_metrics[-1][0]
running_a_acc += a_metrics[-1][1]
if (i+1)% 100 == 0:
print('Epoch #{}'.format(i+1))
log_mesg = "%d: [D loss: %f, acc: %f]" % (i, running_d_loss/i, running_d_acc/i)
log_mesg = "%s [A loss: %f, acc: %f]" % (log_mesg, running_a_loss/i, running_a_acc/i)
print(log_mesg)
noise = np.random.uniform(-1.0, 1.0, size=[16, 100])
gen_imgs = generator.predict(noise)
plt.figure(figsize=(5,5))
for k in range(gen_imgs.shape[0]):
plt.subplot(4, 4, k+1)
plt.imshow(gen_imgs[k, :, :, 0], cmap='gray')
plt.axis('off')
plt.tight_layout()
plt.show()
return a_metrics, d_metrics
# -
a_metrics_complete, d_metrics_complete = train(epochs=3000)
ax = pd.DataFrame(
{
'Generator': [metric[0] for metric in a_metrics_complete],
'Discriminator': [metric[0] for metric in d_metrics_complete],
}
).plot(title='Training Loss', logy=True)
ax.set_xlabel("Epochs")
ax.set_ylabel("Loss")
ax = pd.DataFrame(
{
'Generator': [metric[1] for metric in a_metrics_complete],
'Discriminator': [metric[1] for metric in d_metrics_complete],
}
).plot(title='Training Accuracy')
ax.set_xlabel("Epochs")
ax.set_ylabel("Accuracy")
| notebooks/gans/generative_adversarial_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 12.4.3 Models with increments
#
# The following scripts are dedicated to:
# - the simulation of the paths of a degradation process with exponentially distributed increments
# - the statistical properties of the increments of a degradation process with exponentially distributed increments
# - the simulation of the paths of a homogeneous gamma process
# - the statistical properties of the increments of a degradation process with gamma distributed increments
# - the empirical distribution of the increment of a gamma process
# - the distribution(s) of the hitting time of a gamma process (RUL)
#
# ## Degradation process with exponetially distributed increments
import numpy as np
# %matplotlib notebook
import matplotlib.pyplot as plt
# ### Parameters
# Deterioration rate
lambd = 5
# Window of sampling frequency
# Time parameters
tStart = 0
tEnd = 10
# Sampling frequency window
fs = np.array([1, 1])
# Number of histories
nbHist = 100000
# ### Time vector
#
# In the cell below a time vector with random intervals drawn randomly from an uniform law between lower and upper bounds of the sampling frequency window. To set a constant interval, specify an upper bound equal to the lower one (e.g. `fs = np.array([1, 1])`).
#
# Time:
dt = np.array([tStart])
while np.sum(dt)<tEnd:
dt = np.append(dt, 1/np.random.uniform(low=fs[0], high=fs[1]))
t = np.cumsum(dt)
# ### Drawn of several paths (associated to the same time sampling)
# Rate definition
rate = lambd/(dt[1:].reshape(dt[1:].shape[0], 1)*np.ones((1, nbHist)))
# Drawn of increments
I = np.random.exponential(scale=1/rate)
I = np.concatenate((np.zeros((1, nbHist)), I), axis=0)
# Degradation calculation
Y = np.cumsum(I, axis=0)
# Illustration
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1)
for id in range(np.minimum(75, nbHist)):
ax.plot(t, Y[:, id], '.-', color=(0.8, 0.8, 0.8))
ax.plot(t, np.mean(Y, axis=1), '.-', color=(0, 0, 1))
ax.set_xlabel('Time')
ax.set_ylabel('Degradation')
ax.set_title('Degradation with exponetial increments')
ax.grid(True)
fig.show()
# ### Statistical properties (rather obvious since they correspond to the way how the increments have been defined)
# $\mathrm{E}[I(t_1,t_2)] = (t_2-t_1)/\lambda$
#
idMin = int(np.floor(len(t)*0.2))
idMax = int(np.floor(len(t)*0.8))
print('{:f} ≃ {:f}'.format(np.mean(I[idMin,:]), (t[idMin]-t[idMin-1])/lambd))
print('{:f} ≃ {:f}'.format(np.mean(I[idMax,:]), (t[idMax]-t[idMax-1])/lambd))
# $\mathrm{var}[I(t_1,t_2)] = (t_2-t_1)^2/\lambda^2$
print('{:f} ≃ {:f}'.format(np.var(I[idMin,:]), (t[idMin]-t[idMin-1])**2/lambd**2))
print('{:f} ≃ {:f}'.format(np.var(I[idMax,:]), (t[idMax]-t[idMax-1])**2/lambd**2))
# ---
# ## Homogeneous gamma process
# +
# %reset -f
import numpy as np
# %matplotlib notebook
import matplotlib.pyplot as plt
# -
# ### Parameters
# Deterioration parameters
alpha = 5 # a coefficient of the shape parameter
beta = 6 # the rate parameter (equals to 1/theta, where theta is the scale parameter)
# Time parameters
tStart = 0
tEnd = 10
# Sampling frequency window
fs = np.array([2, 5])
# Number of histories
nbHist = 100000
# ### Time vector
#
# In the cell below a time vector with random intervals drawn randomly from an uniform law between lower and upper bounds of the sampling frequency window. To set a constant interval, specify an upper bound equal to the lower one (e.g. `fs = np.array([1, 1])`).
# Time:
dt = np.array([tStart])
while np.sum(dt)<tEnd:
dt = np.append(dt, 1/np.random.uniform(low=fs[0], high=fs[1]))
t = np.cumsum(dt)
# ### Drawn of several paths (associated to the same time sampling)
# Shape parameter
k = alpha*(dt[1:].reshape(dt[1:].shape[0], 1)*np.ones((1, nbHist)))
# Drawn of increments
I = np.random.gamma(shape=k, scale=1/beta)
I = np.concatenate((np.zeros((1, nbHist)), I), axis=0)
# Degradation calculation
Y = np.cumsum(I, axis=0)
# Illustration
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1)
for id in range(np.minimum(75, nbHist)):
ax.plot(t, Y[:, id], '.-', color=(0.8, 0.8, 0.8))
ax.plot(t, np.mean(Y, axis=1), '.-', color=(0, 0, 1))
ax.plot(t, alpha/beta*t, '.-', color=(1, 0, 0))
ax.set_xlabel('Time')
ax.set_ylabel('Degradation')
ax.set_title('Degradation with homogeneous gamma process')
ax.grid(True)
fig.show()
# ### Statistical properties
# $\mathrm{E}[Y(t_2)-Y(t_1)]=\mathrm{E}[I(t_1, t_2)] = \alpha (t_2-t_1)/{\beta}$
idMin = int(np.floor(len(t)*0.2))
idMax = int(np.floor(len(t)*0.8))
print('{:f} = {:f} ≃ {:f}'.format(np.mean(Y[idMin,:]-Y[idMin-1,:]),
np.mean(I[idMin,:]),
alpha*(t[idMin]-t[idMin-1])/beta))
print('{:f} = {:f} ≃ {:f}'.format(np.mean(Y[idMax,:]-Y[idMax-1,:]),
np.mean(I[idMax,:]),
alpha*(t[idMax]-t[idMax-1])/beta))
# $\mathrm{var}[Y(t_2)-Y(t_1)]=\mathrm{var}[I(t_1, t_2)] = \alpha (t_2-t_1)/\beta^2$
idMin = int(np.floor(len(t)*0.2))
idMax = int(np.floor(len(t)*0.8))
print('{:f} = {:f} ≃ {:f}'.format(np.var(Y[idMin,:]-Y[idMin-1,:]),
np.var(I[idMin,:]),
alpha*(t[idMin]-t[idMin-1])/(beta**2)))
print('{:f} = {:f} ≃ {:f}'.format(np.var(Y[idMax,:]-Y[idMax-1,:]),
np.var(I[idMax,:]),
alpha*(t[idMax]-t[idMax-1])/(beta**2)))
# ---
# ## Homogeneous gamma process: focus on the increment distribution
# +
# %reset -f
import numpy as np
from scipy.stats import gamma
# %matplotlib notebook
import matplotlib.pyplot as plt
# -
# ### Parameters
# Deterioration parameters
alpha = 5 # a coefficient of the shape parameter
beta = 6 # the rate parameter (equals to 1/theta, where theta is the scale parameter)
# Time parameters
tStart = 0
tEnd = 5
# Sampling frequency
fs = 2
# ### Time and a single history
#
# Note that a constant sampling frequency is assumed here.
# Time vector
t = np.linspace(tStart, tEnd, int((tEnd-tStart)*fs)+1)
# Draw degradation increments
I = np.random.gamma(shape=alpha/fs, scale=1/beta, size=(t.shape[0]-1,))
I = np.concatenate(([0], I), axis=0)
# Degradation calculation
Y = np.cumsum(I, axis=0)
yLimS = Y[-1]+2*alpha/(beta**2)
# ### Illustration of the increments distributions
# Figure framwork
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1)
ax.set_xlim([t[0], t[-1]+1/fs])
ax.grid(True)
ax.set_xlabel('Time')
ax.set_ylabel('Degradation')
# Loop illustrating the increments distribution depending on time
for id in range(len(Y)):
ax.plot(t[0:id], Y[0:id], 'o-', color=(1, 0, 0))
ax.plot([t[id], t[id]+1/fs], [Y[id], Y[id]], ':', color=(0, 0, 1))
y = np.linspace(Y[id], yLimS, 100)
z = gamma.pdf(y-Y[id], a=alpha/fs, scale=1/beta)
z = z/z.max()*(t[1]-t[0])*0.75
ax.plot([t[id]+1/fs, t[id]+1/fs], [y[0], y[-1]], '--', color=(0, 0, 1))
ax.plot(z+t[id]+1/fs, y, '-', color=(0, 0, 1))
ax.set_ylim([0, yLimS])
ax.plot(t, Y, 'o-', color=(1, 0, 0))
fig.show()
# ---
# ## Distribution of RUL
# +
# %reset -f
import numpy as np
import numpy.matlib
from scipy.stats import gamma
# %matplotlib notebook
import matplotlib.pyplot as plt
# -
# ### Parameters
# Deterioration parameters
alpha = 2 # a coefficient of the shape parameter
beta = 2 # the rate parameter (equals to 1/theta, where theta is the scale parameter)
# Window of sampling frequency
# Time parameters
tStart = 0
tEnd = 100
# Sampling frequency
fs = 2
# Number of histories
nbHist = 1000
# Failure level
l = 70
# Flag "specific" history
flagSpec = True
tMeas = 55
# ### Time vector and drawn of several paths
#
# Note that a constant sampling frequency is assumed here. Therefore, parameter `alpha` is directly used in the function that draws sample from the Gamma distribution.
# Time vector
t = np.linspace(tStart, tEnd, int((tEnd-tStart)*fs)+1)
# Draw degradation increments
I = np.random.gamma(shape=alpha/fs, scale=1/beta, size=(t.shape[0]-1, nbHist))
I = np.concatenate((np.zeros((1, nbHist)), I), axis=0)
# Degradation calculation
Y = np.cumsum(I, axis=0)
# Same process for the "specific" history
if flagSpec:
ISpec1 = np.random.gamma(shape=alpha/fs, scale=1/beta, size=(np.sum(t<=tMeas)-1, 1))
ISpec2 = np.random.gamma(shape=alpha/fs, scale=1/beta, size=(np.sum(t>tMeas), nbHist))
ISpec = np.concatenate((np.zeros((1, nbHist)), np.matlib.repmat(ISpec1, 1, nbHist), ISpec2), axis=0)
YSpec = np.cumsum(ISpec, axis=0)
# ### Calculation of the failure threshold hitting time
#
# **CAUTION:** for the sake of simplicity, it is assumed that the failure level `l` is lower than the sum of simulated increments, if not, the cell below will raise a message.
#
# The *exact* reaching time is obtained through a linear hypothesis.
if np.sum(Y[-1,:]<l)>0:
print('WARNING: Increase the trajectory duration')
indMax = np.argmax(Y*(Y<l), axis=0)
thTime = np.zeros((nbHist,))
for idh in range(nbHist):
a = (Y[indMax[idh]+1, idh]-Y[indMax[idh], idh])*fs
thTime[idh] = t[indMax[idh]]+(l-Y[indMax[idh], idh])/a
if flagSpec:
indMaxSpec = np.argmax(YSpec*(YSpec<l), axis=0)
thTimeSpec = np.zeros((nbHist,))
for idh in range(nbHist):
aSpec = (YSpec[indMaxSpec[idh]+1, idh]-YSpec[indMaxSpec[idh], idh])*fs
thTimeSpec[idh] = t[indMaxSpec[idh]]+(l-YSpec[indMaxSpec[idh], idh])/aSpec
# ### Illustration of trajectories and hitting time at the level `l`
#
# Distributions of the hitting times are estimated, by simulations, and are given through histograms while true distributions are given through an analytical development relying on equation 12.40 (p. 537).
# Figure framework
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(2, 1, 2)
ax.set_xlabel('Time')
ax.set_ylabel('Degradation')
# Plot trajectories
for idh in range(np.minimum(100, nbHist)):
ax.plot(np.concatenate((t[0:indMax[idh]], np.array([thTime[idh]]))),
np.concatenate((Y[0:indMax[idh], idh], np.array([l]))),
'-', color=(0, 0, 1))
if flagSpec:
for idh in range(np.minimum(100, nbHist)):
ax.plot(np.concatenate((t[0:indMaxSpec[idh]], np.array([thTimeSpec[idh]]))),
np.concatenate((YSpec[0:indMaxSpec[idh], idh], np.array([l]))),
'-', color=(0, 1, 0))
xlim = ax.get_xlim()
ax.plot(xlim, [l, l], 'r-', label='failure level $l$')
ax.set_xlim(xlim)
ax.legend()
ax.grid(True)
# Histogram of the hitting time
ax = fig.add_subplot(2, 1, 1)
ax.hist(thTime, bins=25, density=True, edgecolor=(0, 0, 1), facecolor=(1, 1, 1))
if flagSpec:
ax.hist(thTimeSpec, bins=25, density=True, alpha=0.5, edgecolor=(0, 1, 0),
facecolor=(1, 1, 1))
ax.set_xlabel('Time')
ax.set_ylabel('Hitting time distribution')
ax.set_xlim(xlim)
ax.grid(True)
# Distribution of the hitting time
cumDist = -gamma.cdf(l, a=alpha*t, scale=1/beta)
cumDist = np.concatenate((np.array([0]), np.diff(cumDist)*fs))
ax.plot(t, cumDist, '-', color=(0, 0, 1))
if flagSpec:
cumDist = np.zeros(t.shape);
cumDist[t>=tMeas] = -gamma.cdf(l-np.max(YSpec[t<=tMeas, 0]),
a=alpha*(t[t>=tMeas]-tMeas), scale=1/beta)
cumDist = np.concatenate((np.array([0]), np.diff(cumDist)*fs))
ax.plot(t, cumDist, '-', color=(0, 1, 0))
| 12_PreventiveMaintenance/12_04_DegradationModels/12_04_03_ModelsWithIncrements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''venv'': venv)'
# metadata:
# interpreter:
# hash: ed2973e18a45e46195aea83f71ebc09fe8d3266afb265a31fa421a956b7d8fb0
# name: 'Python 3.7.7 64-bit (''venv'': venv)'
# ---
# # Testing keyword extraction with YAKE
import os
import yake
# ## Extraire les mots clés des documents avec YAKE
# + tags=[]
# Faire une liste des mots à ignorer
ignored = set(["conseil communal", "conseil général"])
ignored
# -
# Instantier l'extracteur de mots clés
kw_extractor = yake.KeywordExtractor(lan="fr", top=20)
kw_extractor
# Lister les PDFs
data_path = "../data/txt/"
files = os.listdir(data_path)
# Imprimer le nombre de PDFs identifiés
len(files)
# Les dix premiers PDFs
files[:20]
# Enlever les fichiers qui ne commencent pas par Bxl_
bxl_files = [f for f in files if f.startswith('Bxl_')]
len(bxl_files)
# Choisir un fichier
this_file = bxl_files[42]
this_file
# Récupérer le texte dans le fichier
text = open(f'{data_path}/{this_file}', encoding='utf-8').read()
text[:500]
# Extraire les mots clés de ce texte
keywords = kw_extractor.extract_keywords(text)
keywords
# Ne garder que les bigrams
kept = []
for score, kw in keywords:
words = kw.split()
if len(words) > 1 and kw not in ignored:
kept.append(kw)
kept
# # Faire la même opération sur tous les documents
for f in sorted(bxl_files)[:10]:
text = open(f'{data_path}/{f}', encoding='utf-8').read()
keywords = kw_extractor.extract_keywords(text)
kept = []
for score, kw in keywords:
words = kw.split()
if len(words) > 1 and kw not in ignored:
kept.append(kw)
print(f"{f} mentions these keywords: {', '.join(kept)}...")
| module3/s1_keywords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install keras
# +
from PIL import Image
import os, glob, sys, numpy as np
from sklearn.model_selection import train_test_split
from keras.utils import np_utils
img_dir = './train1'
categories = ['new_up_2015', 'new_down_2015']
np_classes = len(categories)
image_w = 64
image_h = 64
pixel = image_h * image_w * 3
X = []
y = []
for idx, cat in enumerate(categories):
img_dir_detail = img_dir + "/" + cat
files = glob.glob(img_dir_detail+"/*.png")
for i, f in enumerate(files):
try:
img = Image.open(f)
img = img.convert("RGB")
img = img.resize((image_w, image_h))
data = np.asarray(img)
#Y는 0 아니면 1이니까 idx값으로 넣는다.
X.append(data)
y.append(idx)
if i % 3000 == 0:
print(cat, " : ", f)
except:
print(cat, str(i)+" 번째에서 에러 ")
X = np.array(X)
Y = np.array(y)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.3)
xy = (X_train, X_test, Y_train, Y_test)
np.save("./numpy_data/binary_image_data.npy", xy)
# -
import numpy as np
np.version.version
# +
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Dense, Flatten, Dropout
from keras.callbacks import EarlyStopping, ModelCheckpoint
import matplotlib.pyplot as plt
import keras.backend.tensorflow_backend as K
import numpy as np
X_train, X_test, y_train, y_test = np.load('./numpy_data/binary_image_data.npy')
print(X_train.shape)
print(X_train.shape[0])
print(np.bincount(y_train))
print(np.bincount(y_test))
# +
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# +
image_w = 64
image_h = 64
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
model = Sequential()
model.add(Conv2D(32, (3,3), padding="same", input_shape=X_train.shape[1:], activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(32, (3,3), padding="same", activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(64, (3,3), padding="same", activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3,3), padding="same", activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation="relu"))
model.add(Dropout(0.5))
model.add(Dense(1, activation="sigmoid"))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model_dir = './model'
if not os.path.exists(model_dir):
os.mkdir(model_dir)
model_path = model_dir + "/dog_cat_classify.model"
checkpoint = ModelCheckpoint(filepath=model_path, monitor='val_loss', verbose=1, save_best_only=True)
early_stopping = EarlyStopping(monitor='val_loss', patience=7)
# -
model.summary()
history = model.fit(X_train, y_train, batch_size=64, epochs=100, validation_split=0.15, callbacks=[checkpoint, early_stopping])
print("정확도 : %.3f " %(model.evaluate(X_test, y_test)[1]))
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['loss', 'val_loss', 'acc', 'val_acc'], loc='upper left')
plt.show()
| image_analysis_CNN_binary_2015~2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: base
# language: python
# name: base
# ---
# +
import torch
import numpy as np
import torch.nn as nn
xr = torch.from_numpy(np.array([1,0,2,0,1,0,0,1,2,1,1,2,1,0,1,0,1,1,1,1,1,2,0,1,1]).reshape(-1, 5))
xg = torch.from_numpy(np.array([0,1,2,1,1,2,1,0,1,1,2,1,1,2,0,0,0,1,2,2,0,0,0,1,2]).reshape(-1, 5))
xb = torch.from_numpy(np.array([1,0,0,1,1,1,1,1,0,0,0,2,2,0,2,0,2,0,1,0,2,2,0,1,1]).reshape(-1, 5))
x = torch.cat([xr, xg, xb]).view(3, 5, 5)
w0 = torch.LongTensor([[-1,-1,0,-1,-1,1,-1,0,1], [0,-1,1,1,0,1,1,1,-1],[0,1,0,-1,1,0,-1,-1,1]]).view(3, 3, 3)
xr = xr.numpy().reshape(-1,)
w0_1 = w0[0].numpy().reshape(-1,)
b = [1]
o = np.convolve(xr, w0_1)
print(xr.shape)
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torch.autograd import Variable
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 6, 5)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10, 1)
self.o = nn.Softmin()
def forward(self, x):
x = F.max_pool2d(F.relu(self.conv1(x)), 2)
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
x = self.o(x)
return x
def num_flat_features(self, x):
size = x.size()[1:]
flat_features = 1
for t in size:
flat_features *= t
return flat_features
net = Net()
input = Variable(torch.randn(1, 1, 32, 32))
output = net(input)
# -
from PIL import Image
from torchvision.transforms import ToTensor
import matplotlib.pyplot as plt
print('hi')
img = Image.open('/Users/Shian/ml/deepbi/test_image.tif')
img = ToTensor()(img)
img = img.view(-1, 1).numpy()
print(img.shape)
num = 0
for t in img:
if t != 0:
num += 1
if t == 300:
print(t)
print('end')
print(num)
# +
from __future__ import print_function, division
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
from torch.autograd import Variable
import numpy as np
import torchvision
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import time
import os
import copy
plt.ion() # interactive mode
# +
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
}
data_dir = 'hymenoptera_data'
image_datasets = {
x: datasets.ImageFolder(os.path.join(data_dir, x),
data_transforms[x])
for x in ['train', 'val']
}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x],
batch_size=4,
shuffule=True, num_workers)}
| my-work/deepbi/parse_image_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: analysis
# language: python
# name: analysis
# ---
# # Data profiling
#
# Automatic data profiling using the Python library [pandas-profiling](https://github.com/pandas-profiling/pandas-profiling).
# %matplotlib inline
from sklearn import datasets
import numpy as np
import pandas as pd
from pandas_profiling import ProfileReport
# ### data creation
data = datasets.load_iris()
df = pd.DataFrame(data= np.c_[data['data'], data['target']],columns= list(data['feature_names']) + ['target'])
df['target'] = df['target'].apply(lambda x: data['target_names'][int(x)])
# ### data profiling
# get report
profile = ProfileReport(df)
# display
profile
# report to dictionary
dprofile = profile.description_set
# report to file
profile.to_file("your_report.html")
| notebooks/analysis/profiling/pandas_profiling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise notebook :
# +
import warnings
warnings.simplefilter('ignore', FutureWarning)
import pandas as pd
from datetime import datetime
# -
# # Weather Analysis
#
# You have learned some more about Python and the pandas module and tried it out on a
# fairly small dataset. You are now ready to explore a dataset from the Weather
# Underground.
# ## Weather Data
#
# Will be looking at investigating historic weather data.
# Of course, such data is hugely important for research into the large-scale, long-term shift
# in our planet’s weather patterns and average temperatures – climate change. However,
# such data is also incredibly useful for more mundane planning purposes. To demonstrate
# the learning this week, we will be using historic weather data to try and plan a
# summer holiday. You’ll use the data too and get a chance to work on your own
# project at the end of the week.
# The dataset we’ll use to do this will come from the [Weather Underground](http://www.wunderground.com/), which creates
# weather forecasts from data sent to them by a worldwide network of over 100,000 weather
# enthusiasts who have personal weather stations on their house or in their garden.
# In addition to creating weather forecasts from that data, the Weather Underground also
# keeps that data as historic weather records allowing members of the public to download
# weather datasets for a particular time period and location. These datasets are
# downloaded as CSV files, explained in the next step.
# Datasets are rarely ‘clean’ and fit for purpose, so it will be necessary to clean up the data
# and ‘mould it’ for your purposes. You will then learn how to visualise data by creating
# graphs using the `plot()` function
#
# We have downloaded the file London_2014.csv from our website, it can now be read into a dataframe.
london = pd.read_csv('London_2014.csv')
london.head()
# `Note that the right hand side of the table has been cropped to fit on the page.
# You’ll find out how to remove rogue spaces.`
# ### Removing initial spaces
# One of the problems often encountered with CSV files is rogue spaces before or after data
# values or column names.
# You learned earlier, in What is a CSV file? , that each value or column name is separated
# by a comma. However, if you opened ‘London_2014.csv’ in a text editor, you would see
# that in the row of column names sometimes there are spaces after a comma:
#
# `GMT,Max TemperatureC,Mean TemperatureC,Min TemperatureC,Dew PointC,
# MeanDew PointC,Min DewpointC,Max Humidity, Mean Humidity, Min Humidity,
# Max Sea Level PressurehPa, Mean Sea Level PressurehPa, Min Sea Level
# PressurehPa, Max VisibilityKm, Mean VisibilityKm, Min VisibilitykM, Max Wind
# SpeedKm/h, Mean Wind SpeedKm/h, Max Gust SpeedKm/h,Precipitationmm,
# CloudCover, Events,WindDirDegrees`
#
# For example, there is a space after the comma between Max Humidity and Mean
# Humidity. This means that when read_csv() reads the row of column names it will
# interpret a space after a comma as part of the next column name. So, for example, the
# column name after `'Max Humidity'` will be interpreted as `' Mean Humidity'` rather
# than what was intended, which is `'Mean Humidity'`. The ramification of this is that code
# such as:
#
# `london[['Mean Humidity']]`
#
# will cause a key error (see Selecting a column ), as the column name is confusingly `'
# Mean Humidity '`.
#
# This can easily be rectified by adding another argument to the `read_csv()` function:
# `skipinitialspace=True`
# which will tell `read_csv()` to ignore any spaces after a comma:
# There are too many columns for the dataframe to fit horizontally in this notebook, but they can be displayed separately.
london.columns
# This shows that <code>' Max Wind SpeedKm/h'</code> is prefixed by a space, as are other columm names such as <code>' Mean Humidity'</code> and <code>' Max Sea Level PressurehPa'</code>.
#
# The <code>read_csv()</code> function has interpreted spaces after commas as being part of the next value. This can be rectified easily by adding another argument to the <code>read_csv()</code> function to skip the initial spaces after a comma.
london = pd.read_csv('London_2014.csv', skipinitialspace=True)
# ### Removing extra characters
#
# Another problem shown above is that the final column is called <code>'WindDirDegrees< br />'</code>.
#
# When the dataset was exported from the Weather Underground web site html line breaks were automatically added to each line in the file which <code>read_csv()</code> has interpreted as part of the column name and its values. This can be seen more clearly by looking at more values in the final column:
# In fact, the problem is worse than this, let’s look at some values in the final column:
london['WindDirDegrees<br />'].head()
# It’s seems there is an html line break at the end of each line. If I opened `‘London_2014.
# csv’` in a text editor and looked at the ends of all lines in the file this would be confirmed.
# Once again I’m not going to edit the CSV file but rather fix the problem in the dataframe.
#
# To change `'WindDirDegrees
# '` to `'WindDirDegrees'` all I have to do is use the `rename()` method as follows:
london = london.rename(columns={'WindDirDegrees<br />' : 'WindDirDegrees'})
# Don’t worry about the syntax of the argument for `rename()` , just use this example as a
# template for whenever you need to change the name of a column.
#
# Now I need to get rid of those pesky html line breaks from the ends of the values in the `'WindDirDegrees'` column, so that
# they become something sensible. I can do that using the `string method rstrip()` which
# is used to remove characters from the `end or ‘rear’` of a string, just like this:
london['WindDirDegrees'] = london['WindDirDegrees'].str.rstrip('<br />')
# Again don’t worry too much about the syntax of the code and simply use it as a template
# for whenever you need to process a whole column of values stripping characters from the
# end of each string value.
# Let’s display the first few rows of the `' WindDirDegrees'`to confirm the changes:
london['WindDirDegrees'].head()
# ### Missing values
#
# Missing (also called null or not available) values are marked as NaN (not a number) in dataframes, these are one of the reasons to clean data.
# The `isnull()` method returns `True` for each row in a column that has a null value. The method can be used to select and display those rows. Scroll the table below to the right to check that the events column is only showing missing values.
#
# Finding missing values in a particular column can be done with the column method
# isnull() , like this:
london[london['Events'].isnull()]
# The above code returns a series of Boolean values, where `True` indicates that the
# corresponding row in the `'Events'` column is missing a value and `False` indicates the
# presence of a value.
# One way to deal with missing values is to replace them by some value. The column method `fillna()` fills all not available value cells with the value given as argument. In the example below, each missing event is replaced by the empty string.
# If, as you did with the comparison expressions, you put this code within square brackets
# after the dataframe’s name, it will return a new dataframe consisting of all the rows without
# recorded events **(rain, fog, thunderstorm, etc.):**
london[london['Events'].isnull()]
# This will return a new dataframe with 114 rows, showing that more than one in three days had no particular event recorded.
# If you scroll the table to the right, you will see that all values in the `'Events'` column are
# marked `NaN` , which stands for `‘Not a Number’`, but is also used to mark non-numeric
# missing values, like in this case (events are strings, not numbers).
#
# Once you know how much and where data is missing, you have to decide what to do:
#
# - ignore those rows?
# - Replace with a fixed value?
# - Replace with a computed value, like the mean?
#
# In this case, only the first two options are possible. The method call `london.dropna()`
# will drop (remove) all rows that have a missing (non-available) value somewhere,
# returning a new dataframe. This will therefore also remove rows that have missing values
# in other columns.
# The column method `fillna()` will replace all non-available values with the value given
# as argument. For this case, each NaN could be replaced by the empty string.
london['Events'] = london['Events'].fillna('')
london[london['Events'].isnull()]
# The second line above will now show an empty dataframe, because there are no longer
# missing values in the events column.
# As a final note on missing values, pandas ignores them when computing numeric
# statistics, i.e. you don’t have to remove missing values before applying `sum(),
# median()` and other similar methods.
# The empty dataframe (no rows) confirms there are no more missing event values.
#
# Another way to deal with missing values is to ignore rows with them. The `dropna()` dataframe method returns a new dataframe where all rows with at least one non-available value have been removed.
london.dropna()
# Note that the table above has fewer than 251 of the original 365 rows, so there must be further null values besides the 114 missing events.
# ## Changing the value type of a column
#
# The function `read_csv()` may, for many reasons, wrongly interpret the data type of the
# values in a column, so when cleaning data it’s important to check the data types of each
# column are what is expected, and if necessary change them.
# The type of every column in a dataframe can be determined by looking at the dataframe's `dtypes` attribute, like this:
london.dtypes
# In the above output, you can see the column names to the left and to the right the data
# types of the values in those columns.
# - **int64** is the pandas data type for whole numbers such as `55 or 2356`
# - **float64** is the pandas data type for decimal numbers such as `55.25 or 2356.00`
# - **object** is the pandas data type for strings such as 'hello world' or 'rain'
# Most of the column data types seem fine, however two are of concern, `'GMT'` and
# `'WindDirDegrees'` , both of which are of `type object`. Let’s take a look at
# `'WindDirDegrees'` first.
# **Changing the data type of the `'WindDirDegrees'` column**
#
# The `read_csv()` method has interpreted the values in the `'WindDirDegrees'` column
# as strings `(type object )`. This is because in the CSV file the values in that column had all
# been suffixed with that html line break string
# so `read_csv()` had no alternative but to interpret the values as strings.
# The values in the `'WindDirDegrees'` column are meant to represent wind direction in
# terms of `degrees from true north (360) and meteorologists always define the wind
# direction as the direction the wind is coming from`. So if you stand so that the wind is
# blowing directly into your face, the direction you are facing names the wind, so a westerly
# wind is reported as 270 degrees. The compass rose shown below should make this
# clearer:
# We need to be able to make queries such as ‘Get and display the rows where the wind
# direction is greater than 350 degrees’. To do this we need to change the data type of the
# `‘WindDirDegrees’` column from object to `type int64`.
# The type of all the values in a column can be changed using the <code>astype()</code> method. The following code will change the values in the <code>'WindDirDegrees'</code> column from strings (`object`) to integers (<code>int64</code>).
london['WindDirDegrees'] = london['WindDirDegrees'].astype('int64')
# Now all the values in the `'WindDirDegrees'` column are of `type int64` and we can
# make our query:
london[london['WindDirDegrees'] > 350]
# **Changing the data type of the ‘GMT’ column**
#
# Recall that I noted that the `'GMT'` column was of type object , the type pandas uses for
# strings.
#
# The `'GMT'` column is supposed to represent dates. It would be helpful for the date values
# not to be strings to make it possible to make queries of the data such as `‘Return the row
# where the date is 4 June 2014’`.
#
# Pandas has a function called `to_datetime()` which can convert a column of `object
# (string)` values such as those in the `'GMT'` column into values of a proper date type called
# `datetime64`, just like this:
london['GMT'] = pd.to_datetime(london['GMT'])
london.dtypes
# From the above output, we can confirm that the `'WindDirDegrees'` column type has
# been changed from `object to int64` and that the `'GMT'` column type has been changed
# from `object to datetime64`.
#
# To make queries such as `‘Return the row where the date is 4 June 2014’` you’ll need to be
# able to create a `datetime64 value to represent June 4 2014`. It cannot be:
# `london[london['GMT'] == '2014-1-3']`
# because `‘2014-1-3’` is a string and the values in the `‘GMT’` column are of type
# `datetime64`. Instead you must create a `datetime64 value using thedatetime()`
# function like this:
#
# `datetime(2014, 6, 4)`
#
# In the function call above, the first integer argument is the year, the second the month and
# the third the day.
#
# Let’s try the function out by executing the code to `‘Return the row where the date is 4
# June 2014’`:
london[london['GMT'] == datetime(2014, 6, 4)]
# You can also now make more complex queries involving dates such as 'Return all the rows where the date is between 8 December and 12 December' can be made:
dates = london['GMT']
start = datetime(2014, 12, 8)
end = datetime(2014, 12, 12)
london[(dates >= start) & (dates <= end)]
# ### Tasks
#
# Now that the wind direction is given by a number, write code to select all days that had a northerly wind. Hint: select the rows where the direction is greater than or equal to 350 **or** smaller than or equal to 10, as the compass rose shows.
# In the code cell below, write code to get and display all the rows in the dataframe that are beween 1 April 2014 and
# 11 April 2014.
# In the cell below, write two lines of code to display the first five rows that have a missing value in the `'Max Gust SpeedKm/h'` column. Hint: first select the missing value rows and store them in a new dataframe, then display the first five rows of the new dataframe.
| Caroline WT-043-2021/Data Analysis-Pandas-2/16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 14
# + [markdown] tags=["remove-cell"]
# *Modeling and Simulation in Python*
#
# Copyright 2021 <NAME>
#
# License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=["remove-cell"]
# check if the libraries we need are installed
try:
import pint
except ImportError:
# !pip install pint
try:
import modsim
except ImportError:
# !pip install modsimpy
# -
# ### Code from previous chapters
# +
from modsim import State, System
def make_system(beta, gamma):
"""Make a system object for the SIR model.
beta: contact rate in days
gamma: recovery rate in days
returns: System object
"""
init = State(S=89, I=1, R=0)
init /= sum(init)
t0 = 0
t_end = 7 * 14
return System(init=init, t0=t0, t_end=t_end,
beta=beta, gamma=gamma)
# -
def update_func(state, t, system):
"""Update the SIR model.
state: State with variables S, I, R
t: time step
system: System with beta and gamma
returns: State object
"""
s, i, r = state
infected = system.beta * i * s
recovered = system.gamma * i
s -= infected
i += infected - recovered
r += recovered
return State(S=s, I=i, R=r)
# +
from numpy import arange
from modsim import TimeFrame
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: TimeFrame
"""
frame = TimeFrame(columns=system.init.index)
frame.loc[system.t0] = system.init
for t in arange(system.t0, system.t_end):
frame.loc[t+1] = update_func(frame.loc[t], t, system)
return frame
# -
def calc_total_infected(results, system):
s_0 = results.S[system.t0]
s_end = results.S[system.t_end]
return s_0 - s_end
# +
from modsim import SweepSeries
def sweep_beta(beta_array, gamma):
"""Sweep a range of values for beta.
beta_array: array of beta values
gamma: recovery rate
returns: SweepSeries that maps from beta to total infected
"""
sweep = SweepSeries()
for beta in beta_array:
system = make_system(beta, gamma)
results = run_simulation(system, update_func)
sweep[beta] = calc_total_infected(results, system)
return sweep
# +
from modsim import SweepFrame
def sweep_parameters(beta_array, gamma_array):
"""Sweep a range of values for beta and gamma.
beta_array: array of infection rates
gamma_array: array of recovery rates
returns: SweepFrame with one row for each beta
and one column for each gamma
"""
frame = SweepFrame(columns=gamma_array)
for gamma in gamma_array:
frame[gamma] = sweep_beta(beta_array, gamma)
return frame
# -
beta_array = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0 , 1.1]
gamma_array = [0.2, 0.4, 0.6, 0.8]
frame = sweep_parameters(beta_array, gamma_array)
frame.head()
# In the previous chapters we used simulation to predict the effect of an infectious disease in a susceptible population and to design
# interventions that would minimize the effect.
#
# In this chapter we use analysis to investigate the relationship between the parameters, `beta` and `gamma`, and the outcome of the simulation.
# ## Nondimensionalization
#
# The figures in
# Section [\[sweepframe\]](#sweepframe){reference-type="ref"
# reference="sweepframe"} suggest that there is a relationship between the parameters of the SIR model, `beta` and `gamma`, that determines the outcome of the simulation, the fraction of students infected. Let's think what that relationship might be.
#
# - When `beta` exceeds `gamma`, that means there are more contacts
# (that is, potential infections) than recoveries during each day (or other unit of time). The difference between `beta` and `gamma` might be called the "excess contact rate\", in units of contacts per day.
#
# - As an alternative, we might consider the ratio `beta/gamma`, which
# is the number of contacts per recovery. Because the numerator and
# denominator are in the same units, this ratio is **dimensionless**, which means it has no units.
#
# Describing physical systems using dimensionless parameters is often a
# useful move in the modeling and simulation game. It is so useful, in
# fact, that it has a name: **nondimensionalization** (see
# <http://modsimpy.com/nondim>).
#
# So we'll try the second option first.
# ## Exploring the results
#
# Suppose we have a `SweepFrame` with one row for each value of `beta` and one column for each value of `gamma`. Each element in the `SweepFrame` is the fraction of students infected in a simulation with a given pair of parameters.
#
# We can print the values in the `SweepFrame` like this:
for gamma in frame.columns:
column = frame[gamma]
for beta in column.index:
frac_infected = column[beta]
print(beta, gamma, frac_infected)
# This is the first example we've seen with one `for` loop inside another:
#
# - Each time the outer loop runs, it selects a value of `gamma` from
# the columns of the `DataFrame` and extracts the corresponding
# column.
#
# - Each time the inner loop runs, it selects a value of `beta` from the
# column and selects the corresponding element, which is the fraction
# of students infected.
#
# In the example from the previous chapter, `frame` has 4 columns, one for
# each value of `gamma`, and 11 rows, one for each value of `beta`. So
# these loops print 44 lines, one for each pair of parameters.
#
# The following function encapulates the previous loop and plots the
# fraction infected as a function of the ratio `beta/gamma`:
# +
from matplotlib.pyplot import plot
def plot_sweep_frame(frame):
for gamma in frame.columns:
series = frame[gamma]
for beta in series.index:
frac_infected = series[beta]
plot(beta/gamma, frac_infected, 'o',
color='C1', alpha=0.4)
# +
from modsim import decorate
plot_sweep_frame(frame)
decorate(xlabel='Contact number (beta/gamma)',
ylabel='Fraction infected')
# -
# The results fall on a single curve, at least approximately. That means that we can predict the fraction of students who will be infected based on a single parameter, the ratio `beta/gamma`. We don't need to know the values of `beta` and `gamma` separately.
# ## Contact number
#
# From Section xxx, recall that the number of new infections in a
# given day is $\beta s i N$, and the number of recoveries is
# $\gamma i N$. If we divide these quantities, the result is
# $\beta s / \gamma$, which is the number of new infections per recovery
# (as a fraction of the population).
#
# When a new disease is introduced to a susceptible population, $s$ is
# approximately 1, so the number of people infected by each sick person is $\beta / \gamma$. This ratio is called the "contact number\" or "basic reproduction number\" (see <http://modsimpy.com/contact>). By convention it is usually denoted $R_0$, but in the context of an SIR model, this notation is confusing, so we'll use $c$ instead.
# The results in the previous section suggest that there is a relationship between $c$ and the total number of infections. We can derive this relationship by analyzing the differential equations from
# Section xxx:
#
# $$\begin{aligned}
# \frac{ds}{dt} &= -\beta s i \\
# \frac{di}{dt} &= \beta s i - \gamma i\\
# \frac{dr}{dt} &= \gamma i\end{aligned}$$
#
# In the same way we divided the
# contact rate by the infection rate to get the dimensionless quantity
# $c$, now we'll divide $di/dt$ by $ds/dt$ to get a ratio of rates:
#
# $$\frac{di}{ds} = -1 + \frac{1}{cs}$$
#
# Dividing one differential equation by another is not an obvious move, but in this case it is useful because it gives us a relationship between $i$, $s$ and $c$ that does not depend on time. From that relationship, we can derive an equation that relates $c$ to the final value of $s$. In theory, this equation makes it possible to infer $c$ by observing the course of an epidemic.
# Here's how the derivation goes. We multiply both sides of the previous
# equation by $ds$:
#
# $$di = \left( -1 + \frac{1}{cs} \right) ds$$
#
# And then integrate both sides:
#
# $$i = -s + \frac{1}{c} \log s + q$$
#
# where $q$ is a constant of integration. Rearranging terms yields:
#
# $$q = i + s - \frac{1}{c} \log s$$
#
# Now let's see if we can figure out what $q$ is. At the beginning of an epidemic, if the fraction infected is small and nearly everyone is susceptible, we can use the approximations $i(0) = 0$ and $s(0) = 1$ to compute $q$:
#
# $$q = 0 + 1 + \frac{1}{c} \log 1$$
#
# Since $\log 1 = 0$, we get $q = 1$.
# Now, at the end of the epidemic, let's assume that $i(\infty) = 0$, and $s(\infty)$ is an unknown quantity, $s_{\infty}$. Now we have:
#
# $$q = 1 = 0 + s_{\infty}- \frac{1}{c} \log s_{\infty}$$
#
# Solving for $c$, we get $$c = \frac{\log s_{\infty}}{s_{\infty}- 1}$$ By relating $c$ and $s_{\infty}$, this equation makes it possible to estimate $c$ based on data, and possibly predict the behavior of future epidemics.
# ## Analysis and simulation
#
# Let's compare this analytic result to the results from simulation. I'll create an array of values for $s_{\infty}$
# +
from numpy import linspace
s_inf_array = linspace(0.0001, 0.999, 31)
# -
# And compute the corresponding values of $c$:
# +
from numpy import log
c_array = log(s_inf_array) / (s_inf_array - 1)
# -
# To get the total infected, we compute the difference between $s(0)$ and
# $s(\infty)$, then store the results in a `Series`:
frac_infected = 1 - s_inf_array
# Recall from Section [\[dataframe\]](#dataframe){reference-type="ref"
# reference="dataframe"} that a `Series` object contains an index and a
# corresponding sequence of values. In this case, the index is `c_array`
# and the values are from `frac_infected`.
#
# Now we can plot the results:
# compares the analytic results from this
# section with the simulation results from
# Section [\[nondim\]](#nondim){reference-type="ref" reference="nondim"}.
# +
plot_sweep_frame(frame)
plot(c_array, frac_infected, label='analysis')
decorate(xlabel='Contact number (c)',
ylabel='Fraction infected')
# -
# When the contact number exceeds 1, analysis and simulation agree. When
# the contact number is less than 1, they do not: analysis indicates there should be no infections; in the simulations there are a small number of infections.
#
# The reason for the discrepancy is that the simulation divides time into a discrete series of days, whereas the analysis treats time as a
# continuous quantity. In other words, the two methods are actually based on different models. So which model is better?
#
# Probably neither. When the contact number is small, the early progress
# of the epidemic depends on details of the scenario. If we are lucky, the original infected person, "patient zero", infects no one and there is no epidemic. If we are unlucky, patient zero might have a large number of close friends, or might work in the dining hall (and fail to observe safe food handling procedures).
#
# For contact numbers near or less than 1, we might need a more detailed
# model. But for higher contact numbers the SIR model might be good
# enough.
# ## Estimating contact number
#
# Figure xxx shows that if we know the contact number, we can compute the fraction infected. But we can also read the figure the other way; that is, at the end of an epidemic, if we can estimate the fraction of the population that was ever infected, we can use it to estimate the contact number.
#
# Well, in theory we can. In practice, it might not work very well,
# because of the shape of the curve. When the contact number is near 2,
# the curve is quite steep, which means that small changes in $c$ yield
# big changes in the number of infections. If we observe that the total
# fraction infected is anywhere from 20% to 80%, we would conclude that
# $c$ is near 2.
#
# On the other hand, for larger contact numbers, nearly the entire
# population is infected, so the curve is nearly flat. In that case we
# would not be able to estimate $c$ precisely, because any value greater
# than 3 would yield effectively the same results. Fortunately, this is
# unlikely to happen in the real world; very few epidemics affect anything close to 90% of the population.
#
# So the SIR model has limitations; nevertheless, it provides insight into the behavior of infectious disease, especially the phenomenon of herd immunity. As we saw in Chapter xxx, if we know the parameters of the model, we can use it to evaluate possible interventions. And as we saw in this chapter, we might be able to use data from earlier outbreaks to estimate the parameters.
#
# ## Exercises
# **Exercise:** If we didn't know about contact numbers, we might have explored other possibilities, like the difference between `beta` and `gamma`, rather than their ratio.
#
# Write a version of `plot_sweep_frame`, called `plot_sweep_frame_difference`, that plots the fraction infected versus the difference `beta-gamma`.
#
# What do the results look like, and what does that imply?
# +
# Solution
def plot_sweep_frame_difference(frame):
for gamma in frame.columns:
column = frame[gamma]
for beta in column.index:
frac_infected = column[beta]
plot(beta - gamma, frac_infected, 'ro',
color='C1', alpha=0.4)
# +
# Solution
plot_sweep_frame_difference(frame)
decorate(xlabel='Excess infection rate (infections-recoveries per day)',
ylabel='Fraction infected')
# +
# Solution
# The results don't fall on a line, which means that if we
# know the difference between `beta` and `gamma`,
# but not their ratio, that's not enough to predict
# the fraction infected.
# -
# **Exercise:** Suppose you run a survey at the end of the semester and find that 26% of students had the Freshman Plague at some point.
#
# What is your best estimate of `c`?
#
# Hint: if you print `frac_infected_series`, you can read off the answer.
# +
# Solution
from pandas import Series
Series(frac_infected, index=c_array)
# +
# Solution
# It looks like the fraction infected is 0.26 when the contact
# number is about 1.16
# -
| jupyter/chap14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
from typing import Callable, Dict, Tuple
from collections import namedtuple
import numpy as np
import matplotlib.pyplot as plt
from scipy.linalg import ldl
# -
# ## Carga de Datos
# +
file_name = './yk.txt'
with open(file_name) as f:
Y = [y for line in f for y in map(lambda y: float(y), line.split(','))]
Y = np.array(Y[1:]) # Remove the list lenght
Y.shape
# -
# ## Descenso de Gradiente
# +
def _back_tracking(x_k: np.array,
grad_k: np.array,
f :Callable[[], float],
alpha:float=100.0,
ro:float=0.95,
c1:float=1E-4,
**kwargs) -> float:
""" Search by Backtraking for an step size valid """
f_kwargs = kwargs.get('f_kwargs', {})
_alpha = alpha
while f(x_k - _alpha * grad_k, **f_kwargs) > f(x_k, **f_kwargs) - c1 * _alpha * grad_k @ grad_k:
_alpha = ro * _alpha
return _alpha
def gradient_descent(X: np.array,
f: Callable[[np.array], np.array],
g: Callable[[np.array], np.array],
H: Callable[[np.array], np.array],
mxitr: int=10000,
tol_x: float=1e-12,
tol_f: float=1e-12,
tol_g: float=1e-12,
msg='StepHess',
**kwargs):
""" Gradient descent implementation
Args:
X: Start Point
mxitr: Maximun number of iterations
tol_x: Minimun value to reach the stopping criteria for norm(xk_1 - xk) / max(1, norm(xk)), used by default
tol_g: Minimun value to reach the stopping criteria for norm(grad_k)
tol_f: Minimun value to reach the stopping criteria for norm(f(xk_1) - f(xk)) / max(1, norm(f(xk)))
f: Function to minimize
g: Gradient of f
H: Hessian of f
msg: Type of computation for the step length (alpha)
valid options are 'StepHess', 'Backtracking', 'StepFixed'
alpha(float): Step size, required when 'msg'='StepFixed'
backtraking_options(Dict): Dictionary with the Backtraking options; 'alpha', 'ro', 'c1'
Returns:
A tuple with the [ ||x_k+1 − x_k || ], [ f(x_k) ], [ grad(f(x_k)) ] values
"""
Result = namedtuple('Result', 'x_log f_log stop_x_log stop_f_log stop_g_log')
# History of values computed
x_log = []; f_log = []
stop_x_log = []; stop_f_log = []; stop_g_log = []
# Initials values
alpha_k = kwargs.get('alpha', 0.001)
x_k = X
x_k_next = None
# Stop criterias
stop_x = lambda x_k, x_k_next: np.linalg.norm(x_k_next - x_k) / max(np.linalg.norm(x_k), 1.0)
stop_f = lambda f_x_k, f_x_k_next: np.abs(f_x_k_next - f_x_k) / max(np.abs(f_x_k), 1.0)
stop_g = lambda grad_k: np.linalg.norm(grad_k)
step = 0
while True:
# Compute gradient
grad_k = g(x_k, **kwargs.get('g_kwargs', {}))
hessian_k = H(x_k, **kwargs.get('H_kwargs', {}))
# Compute step size
if msg == 'StepHess':
alpha_k = (grad_k @ grad_k) / (grad_k @ hessian_k @ grad_k)
elif msg == 'Backtracking':
alpha_k = _back_tracking(x_k, grad_k, f, f_kwargs=kwargs.get('g_kwargs', {}), **kwargs.get('backtraking_kwargs', {}))
else:
pass # value from alpha param
# Compute next x
x_k_next = x_k - alpha_k * grad_k
# Save algorithm logs
f_x_k = f(x_k, **kwargs.get('f_kwargs', {}))
f_x_k_next = f(x_k_next, **kwargs.get('f_kwargs', {}))
x_log.append(x_k_next)
f_log.append(f_x_k_next)
stop_x_log.append(stop_x(x_k, x_k_next))
stop_g_log.append(stop_g(grad_k))
stop_f_log.append(stop_f(f_x_k, f_x_k_next))
if kwargs.get('log', False) and step % kwargs.get('step_log', 100) == 0:
print(f_log[-1], stop_x_log[-1], stop_g_log[-1], stop_f_log[-1])
# Stop criteria
if mxitr < step or stop_x_log[-1] < tol_x or stop_g_log[-1] < tol_g or stop_f_log[-1] < tol_f:
break
# Update x_k
x_k = x_k_next
step += 1
return Result(np.array(x_log), np.array(f_log), np.array(stop_x_log), np.array(stop_f_log), np.array(stop_g_log))
# -
# ## Newton Method
def newton_method(X: np.array,
f: Callable[[np.array], np.array],
g: Callable[[np.array], np.array],
H: Callable[[np.array], np.array],
mxitr: int=10000,
tol_x: float=1e-12,
tol_f: float=1e-12,
tol_g: float=1e-12,
**kwargs):
""" Newton Method implementation
Args:
X: Start Point
mxitr: Maximun number of iterations
tol_x: Minimun value to reach the stopping criteria for norm(xk_1 - xk) / max(1, norm(xk)), used by default
tol_g: Minimun value to reach the stopping criteria for norm(grad_k)
tol_f: Minimun value to reach the stopping criteria for norm(f(xk_1) - f(xk)) / max(1, norm(f(xk)))
f: Function to minimize
g: Gradient of f
H: Hessian of f
alpha(float): Step size, required when 'msg'='StepFixed'
Returns:
A tuple with the [ ||x_k+1 − x_k || ], [ f(x_k) ], [ grad(f(x_k)) ] values
"""
Result = namedtuple('Result', 'x_log f_log stop_x_log stop_f_log stop_g_log')
# History of values computed
x_log = []; f_log = []
stop_x_log = []; stop_f_log = []; stop_g_log = []
# Initials values
alpha_k = kwargs.get('alpha', 1.0)
x_k = X
x_k_next = None
# Stop criterias
stop_x = lambda x_k, x_k_next: np.linalg.norm(x_k_next - x_k) / max(np.linalg.norm(x_k), 1.0)
stop_f = lambda f_x_k, f_x_k_next: np.abs(f_x_k_next - f_x_k) / max(np.abs(f_x_k), 1.0)
stop_g = lambda grad_k: np.linalg.norm(grad_k)
step = 0
while True:
# Compute gradient
grad_k = g(x_k, **kwargs.get('g_kwargs', {}))
hessian_k = H(x_k, **kwargs.get('H_kwargs', {}))
# Ensure that the H is positive semidefined
_, D, _ = ldl(hessian_k)
min_l = np.min(np.sum(D, axis=0))
if min_l < 0:
hessian_k = hessian_k - min_l
# Compute next x
x_k_next = x_k - alpha_k * (np.linalg.inv(hessian_k) @ grad_k)
# Save algorithm logs
f_x_k = f(x_k, **kwargs.get('f_kwargs', {}))
f_x_k_next = f(x_k_next, **kwargs.get('f_kwargs', {}))
x_log.append(x_k_next)
f_log.append(f_x_k_next)
stop_x_log.append(stop_x(x_k, x_k_next))
stop_g_log.append(stop_g(grad_k))
stop_f_log.append(stop_f(f_x_k, f_x_k_next))
if kwargs.get('log', False) and step % kwargs.get('step_log', 100) == 0:
print(f_log[-1], stop_x_log[-1], stop_g_log[-1], stop_f_log[-1])
# Stop criteria
if mxitr < step or stop_x_log[-1] < tol_x or stop_g_log[-1] < tol_g or stop_f_log[-1] < tol_f:
break
# Update x_k
x_k = x_k_next
step += 1
return Result(np.array(x_log), np.array(f_log), np.array(stop_x_log), np.array(stop_f_log), np.array(stop_g_log))
# ## Definición de la funcion
def f1(X: np.array=None, Y: np.array=None, _lambda: float=1.0):
""" Returns the evaluation for f """
return np.sum((X - Y)**2 ) + _lambda * np.sum((X[1:] - X[:-1])**2)
def gradient(X: np.array=None, Y: np.array=None, _lambda:float=1.0):
""" Return the evaluation for the gradient of f """
s1 = X - Y
s2 = X[1:] - X[:-1]
return 2 * s1 + 2 * _lambda * (np.concatenate([[0], s2]) - np.concatenate([s2, [0]]))
def hessian(X: np.array=None, _lambda :float=1.0):
""" Return the evaluation for the Hessian of f """
size = X.shape[0]
h = np.zeros((size, size))
for i in range(size):
h[i][i] = 2 * _lambda + 2 if i == 0 or i == size-1 else 4 * _lambda + 2
if i > 0:
h[i][i-1] = h[i-1][i] = - 2 * _lambda
return h
# ## Ejecuciones Para Método de Newton
def plot_xy(x, y, title=''):
plt.figure(figsize=(16, 8))
plt.plot(x)
plt.plot(Y)
plt.title(title)
plt.xlabel('Index i')
plt.ylabel('Values for x_i and y_i')
plt.legend(['X', 'Y'])
plt.grid(True)
plt.show()
X = np.array(range(0, len(Y)))
# ### Lambda = 1
# +
_lambda = 1
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 10000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'alpha': 1,
}
toc = time.time()
results = newton_method(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print(results.stop_x_log)
plot_xy(results.x_log[-1], Y, "Método de Newton con lambda=1")
# -
# ### Lambda = 100
# +
_lambda = 100
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 10000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'alpha': 1,
}
toc = time.time()
results = newton_method(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print(results.stop_x_log)
plot_xy(results.x_log[-1], Y, "Método de Newton con lambda=100")
# -
# ### Lambda = 1000
# +
_lambda = 1000
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 10000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'alpha': 1,
}
toc = time.time()
results = newton_method(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print(results.stop_x_log)
plot_xy(results.x_log[-1], Y, "Método de Newton con lambda=1000")
# -
# ## Ejecuciones para Método de Gradiente
# ### Lambda = 1
# +
_lambda = 1
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 10000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'msg':'StepHess',
# 'msg':'StepFixed',
# 'msg':'Backtracking',
'alpha': 0.17,
'backtraking_kwargs': {
'alpha': 0.17,
'ro': 0.001,
'c1': 1e-14
}
}
toc = time.time()
results = gradient_descent(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print("Iterations %d" % (len(results.stop_f_log)))
print(results.stop_f_log)
plot_xy(results.x_log[-1], Y, "Descenso de Gradiente con lambda=100")
# -
# ### Lambda = 100
# +
_lambda = 100
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 10000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'msg':'StepHess',
# 'msg':'StepFixed',
# 'msg':'Backtracking',
'alpha': 0.0024,
'backtraking_kwargs': {
'alpha': 0.0024,
'ro': 0.001,
'c1': 1e-14
}
}
toc = time.time()
results = gradient_descent(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print("Iterations %d" % (len(results.stop_f_log)))
print(results.stop_f_log)
plot_xy(results.x_log[-1], Y, "Descenso de Gradiente con lambda=100")
# -
# ### Lambda = 1000
# +
_lambda = 1000
params = {
'X': X,
'f': f1,
'f_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'g': gradient,
'g_kwargs': {
'Y': Y,
'_lambda': _lambda
},
'H': hessian,
'H_kwargs': {
'_lambda': _lambda
},
'mxitr': 100000,
'tol_x': 1e-12,
'tol_g': 1e-12,
'tol_f': 1e-12,
'msg':'StepHess',
# 'msg':'StepFixed',
# 'msg':'Backtracking',
'alpha': 0.0001,
'backtraking_kwargs': {
'alpha': 0.0001,
'ro': 0.001,
'c1': 1e-14
}
}
toc = time.time()
results = gradient_descent(**params)
tic = time.time()
print("Time: %ss" % (tic-toc))
print("Iterations %d" % (len(results.stop_f_log)))
print(results.stop_f_log)
plot_xy(results.x_log[-1], Y, "Descenso de Gradiente con lambda=100")
# -
| Tarea 3/src/Tarea3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Container Basics for Research
#
# In this workshop, we will start with a simple application running in a Docker container. We will take a closer look at the key components and environments that are needed.
#
# There are several container technologies available, but Docker container is the most popular once. We will focus on Docker container in theis workshop.
#
# We will also explore diffent ways of running containers in AWS with different services.
#
# Why containers for research
# - Repeatable and sharable tools and applications
# - Portable - run on different environemnts ( develop on laptop, test on-prem, run large scale in the cloud)
# - Stackable - run differnet applications in a pipeline with different OS, e.g.
#
#You only need to do this once per kernel - used in analyzing fastq data. If you don't need to run the last step, you don't need this
# #!pip install bioinfokit
# +
import boto3
import botocore
import json
import time
import os
import base64
import docker
import pandas as pd
import project_path # path to helper methods
from lib import workshop
from botocore.exceptions import ClientError
# create a bucket for the workshop to store output files.
session = boto3.session.Session()
bucket = workshop.create_bucket_name('container-ws-')
session.resource('s3').create_bucket(Bucket=bucket)
print(bucket)
# -
# First of all, let's create a helper magic for us to easily create and save a file from the notebook
# +
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, 'w+') as f:
f.write(cell.format(**globals()))
# -
# ## Running an application in a container locally.
#
# This SageMaker Jupyter notebook runs on an EC2 instance with docker daemon installed. We can build and test docker containers on the same instance.
#
# We are going to build a simple web server container that says "Hello World!". Let's start with the Docker files
#
# ### Let's start the the Dockerfile
# Think about the Dockerfile as the automation script that you usually do on a linux VM. It just run inside an container. You start with a base image (in this case ubuntu:18.04), then you install, configue compile or build the software you need.
#
# +
# %%writetemplate Dockerfile
FROM ubuntu:18.04
# Install dependencies and apache web server
RUN apt-get update && apt-get -y install apache2
# Create the index html
RUN echo 'Hello World!' > /var/www/html/index.html
# Configure apache
RUN echo '. /etc/apache2/envvars' > /root/run_apache.sh && \
echo 'mkdir -p /var/run/apache2' >> /root/run_apache.sh && \
echo 'mkdir -p /var/lock/apache2' >> /root/run_apache.sh && \
echo '/usr/sbin/apache2 -D FOREGROUND' >> /root/run_apache.sh && \
chmod 755 /root/run_apache.sh
EXPOSE 80
CMD /root/run_apache.sh
# -
# ### Now let's build the container.
#
# The server that runs this SageMaker Jupyter notebook happen to have "docker" runtime installed.
# Docker builld will use the "Dockerfile" in the current directory and use "-t" to build and tag the image. The image will be in the local docker image registry.
#
# We will later learn how to use an external image registry (AWS ECR, e.g.) to push the image to.
# !docker build -t simple_server .
# ### Run the container
#
# Run the container locally, we will bind the container port 80 to the localhsot port 8080 ("-d" runs detached/background)
#
# We use curl to access the web server on port 8080
#
# +
# c_id = !docker run -d -p 8080:80 simple_server
# !curl http://localhost:8080
# +
docker_client = docker.from_env()
simple_server_container = docker_client.containers.get(c_id[0])
def list_all_running_containers():
docker_client = docker.from_env()
container_list = docker_client.containers.list()
for c in container_list:
print(c.attrs['Id'], c.attrs['State']['Status'])
return container_list
running_containers = list_all_running_containers()
# Now stop the running container
simple_server_container.stop()
# -
# ## Let's run some real workload
#
# We are going to use The NCBI SRA (Sequence Read Archive) SRA Tool (https://github.com/ncbi/sra-tools) fasterq-dump (https://github.com/ncbi/sra-tools/wiki/HowTo:-fasterq-dump) to extract fastq from SRA-accessions.
#
# The command takes a package name as an argument
# ```
# $ fasterq-dump SRR000001
# ```
#
# The base image is provided by https://hub.docker.com/r/pegi3s/sratoolkit/
#
# The workflow of the contianer:
# 1. Upon start, container runs a script "sratest.sh".
# 3. sratest.sh will "prefetch" the data package, whose name is passed via an environment variable.
# 4. sratest.sh then run fasterq-dump on the dat apackage
# 5. sratest.sh will then upload the result to S3://{bucket}
#
# The output of the fasterq-dump will be stored in s3://{bucket}/data/sra-toolkit/fasterq/{{PACKAGE_NAME}
#
# +
PACKAGE_NAME='SRR000002'
# this is where the output will be stored
sra_prefix = 'data/sra-toolkit/fasterq'
sra_output = f"s3://{bucket}/{sra_prefix}"
# to run the docker container locally, you need the access credtitials inside the container when usign aws cli
# pass the current keys and session token to the container va environment variables
credentials = boto3.session.Session().get_credentials()
current_credentials = credentials.get_frozen_credentials()
# Please don't print those out:
access_key=current_credentials.access_key
secret_key=current_credentials.secret_key
token=current_credentials.token
# +
# %%writetemplate sratest.sh
# #!/bin/bash
set -x
# this is where ncbi/sra-toolkit is installed on the container inside the pegi3s/sratookit image
#export PATH="/opt/sratoolkit.2.9.6-ubuntu64/bin:${{PATH}}"
prefetch $PACKAGE_NAME --output-directory /tmp
fasterq-dump $PACKAGE_NAME -e 18
aws s3 sync . $SRA_OUTPUT/$PACKAGE_NAME
# +
# %%writetemplate Dockerfile.pegi3s
FROM pegi3s/sratoolkit
RUN apt-get update --fix-missing && apt-get install -y unzip python
RUN wget -O "awscli-bundle.zip" -q "https://s3.amazonaws.com/aws-cli/awscli-bundle.zip"
RUN unzip awscli-bundle.zip
RUN ./awscli-bundle/install -i /usr/local/aws -b /usr/local/bin/aws
RUN export PATH=/usr/local/bin/aws/bin:$PATH
ADD sratest.sh /usr/local/bin/sratest.sh
RUN chmod +x /usr/local/bin/sratest.sh
WORKDIR /tmp
ENTRYPOINT ["/usr/local/bin/sratest.sh"]
# -
#
#
# !docker build -t myncbi/sra-tools -f Dockerfile.pegi3s .
# +
PACKAGE_NAME='SRR000002'
# only run this when you need to clean up the registry and storage
# #!docker system prune -a -f
# !docker run --env SRA_OUTPUT=$sra_output --env PACKAGE_NAME=$PACKAGE_NAME --env PACKAGE_NAME=$PACKAGE_NAME --env AWS_ACCESS_KEY_ID=$access_key --env AWS_SECRET_ACCESS_KEY=$secret_key --env AWS_SESSION_TOKEN=$token myncbi/sra-tools:latest
# -
# Now try a differnet package
PACKAGE_NAME = 'SRR000003'
# !docker run --env SRA_OUTPUT=$sra_output --env PACKAGE_NAME=$PACKAGE_NAME --env PACKAGE_NAME=$PACKAGE_NAME --env AWS_ACCESS_KEY_ID=$access_key --env AWS_SECRET_ACCESS_KEY=$secret_key --env AWS_SESSION_TOKEN=$token myncbi/sra-tools:latest
# ### Build your own docker image
#
# So far, we have been using existing pegi3s ncbi/sratools image. Let's build our own image using a ubuntu base image.
#
# 1. Install tzdata - this is a dependency of some of the other packages we need. Normally we do not need to install it specifically, however there is an issue with tzdata requireing an interaction to select timezone during the installation process, which would halt the docker built. so install it separately with -y.
# 2. Install wget and awscli.
# 3. Download sratookit ubuntu binary and unzip into /opt
# 4. set the PATH to include sratoolkit/bin
# 5. USER nobody is needed to set the permission for sratookit configuration.
# 6. use the same sratest.sh script
# +
# %%writetemplate Dockerfile.myown
#FROM ubuntu:18.04
FROM public.ecr.aws/ubuntu/ubuntu:latest
RUN apt-get update
RUN DEBIAN_FRONTEND="noninteractive" apt-get -y install tzdata \
&& apt-get install -y wget libxml-libxml-perl awscli
RUN wget -q https://ftp-trace.ncbi.nlm.nih.gov/sra/sdk/2.10.0/sratoolkit.2.10.0-ubuntu64.tar.gz -O /tmp/sratoolkit.tar.gz \
&& tar zxf /tmp/sratoolkit.tar.gz -C /opt/ && rm /tmp/sratoolkit.tar.gz
ENV PATH="/opt/sratoolkit.2.10.0-ubuntu64/bin/:${{PATH}}"
ADD sratest.sh /usr/local/bin/sratest.sh
RUN chmod +x /usr/local/bin/sratest.sh
WORKDIR /tmp
USER nobody
ENTRYPOINT ["/usr/local/bin/sratest.sh"]
# -
# Build the image
# !docker build -t myownncbi/sra-tools -f Dockerfile.myown .
# +
PACKAGE_NAME='SRR000004'
# !docker run --env SRA_OUTPUT=$sra_output --env PACKAGE_NAME=$PACKAGE_NAME --env AWS_ACCESS_KEY_ID=$access_key --env AWS_SECRET_ACCESS_KEY=$secret_key --env AWS_SESSION_TOKEN=$token myownncbi/sra-tools:latest
# +
# checkou the outfiles on S3
s3_client = session.client('s3')
objs = s3_client.list_objects(Bucket=bucket, Prefix=sra_prefix)
for obj in objs['Contents']:
fn = obj['Key']
p = os.path.dirname(fn)
if not os.path.exists(p):
os.makedirs(p)
s3_client.download_file(bucket, fn , fn)
# +
# you can use interactive python interpreter, jupyter notebook, google colab, spyder or python code
# I am using interactive python interpreter (Python 3.8.2)
from bioinfokit.analys import fastq
fastq_iter = fastq.fastq_reader(file=f"{sra_prefix}/{PACKAGE_NAME}/{PACKAGE_NAME}.fastq")
# read fastq file and print out the first 10,
i = 0
for record in fastq_iter:
# get sequence headers, sequence, and quality values
header_1, sequence, header_2, qual = record
# get sequence length
sequence_len = len(sequence)
# count A bases
a_base = sequence.count('A')
if i < 10:
print(sequence, qual, a_base, sequence_len)
i +=1
print(f"Total number of records for package {PACKAGE_NAME} : {i}")
# -
# !aws s3 rb s3://$bucket --force
# !rm -rf $sra_prefix
# ## Other ways to run the container
#
# We looked at creating and running containers locally in this notebook. Please checkout notebook/hpc/hatch-fastqc notebook for running containers in AWS Batch service.
| notebooks/container/container-basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Survival Data
# +
from sksurv.datasets import load_veterans_lung_cancer
data_x, data_y = load_veterans_lung_cancer()
# -
data_x.head()
data_x.shape
data_y.shape
data_y[:5]
pd.DataFrame.from_records(data_y[[11, 5, 32, 13, 23]], index=range(1, 6))
# +
from sksurv.nonparametric import kaplan_meier_estimator
time, survival_prob = kaplan_meier_estimator(data_y['Status'], data_y['Survival_in_days'])
plt.step(time, survival_prob, where='post')
plt.ylabel('est. probability of survival $\hat{S}(t)$')
plt.xlabel('time $t$');
# -
# ## Survival functions by treatment
data_x['Treatment'].value_counts()
# +
for treatment_type in ('standard', 'test'):
mask_treat = data_x['Treatment'] == treatment_type
time_treatment, survival_prob_treatment = kaplan_meier_estimator(
data_y['Status'][mask_treat],
data_y['Survival_in_days'][mask_treat])
plt.step(time_treatment, survival_prob_treatment, where='post',
label = 'Treatment = %s' % treatment_type)
plt.ylabel('est. probability of survival $\hat{S}(t)$')
plt.xlabel('time $t$')
plt.legend(loc='best');
# -
# ## Survival functions by cell type
# +
for value in data_x['Celltype'].unique():
mask = data_x['Celltype'] == value
time_cell, survival_prob_cell = kaplan_meier_estimator(data_y['Status'][mask],
data_y['Survival_in_days'][mask])
plt.step(time_cell, survival_prob_cell, where='post',
label = '%s (n = %d)' % (value, mask.sum()))
plt.ylabel('est. probability of survival $\hat{S}(t)$')
plt.xlabel('time $t$')
plt.legend(loc='best');
# -
# ## Multivariate Survival Models
# +
from sksurv.preprocessing import OneHotEncoder
data_x_numeric = OneHotEncoder().fit_transform(data_x)
# -
data_x_numeric.head()
# +
from sksurv.linear_model import CoxPHSurvivalAnalysis
estimator = CoxPHSurvivalAnalysis()
estimator.fit(data_x_numeric, data_y)
# -
pd.Series(estimator.coef_, index=data_x_numeric.columns)
x_new = pd.DataFrame.from_dict({
1: [65, 0, 0, 1, 60, 1, 0, 1],
2: [65, 0, 0, 1, 60, 1, 0, 0],
3: [65, 0, 1, 0, 60, 1, 0, 0],
4: [65, 0, 1, 0, 60, 1, 0, 1]},
columns=data_x_numeric.columns, orient='index')
x_new
# +
pred_surv = estimator.predict_survival_function(x_new)
time_points = np.arange(1, 1000)
for i, surv_func in enumerate(pred_surv):
plt.step(time_points, surv_func(time_points), where='post',
label='Sample %d' % (i+1))
plt.ylabel('est. probability of survival $\hat{S}(t)$')
plt.xlabel('time $t$')
plt.legend(loc='best');
# -
# ## Measuring the Performance of Survival Models
# +
from sksurv.metrics import concordance_index_censored
prediction = estimator.predict(data_x_numeric)
result = concordance_index_censored(data_y['Status'], data_y['Survival_in_days'], prediction)
result[0]
# -
estimator.score(data_x_numeric, data_y)
# ## Feature Selection: Which Variable is Most Predictive?
# +
def fit_and_score_features(X, y):
n_features = X.shape[1]
scores = np.empty(n_features)
m = CoxPHSurvivalAnalysis()
for j in range(n_features):
Xj = X[:,j:j+1]
m.fit(Xj, y)
scores[j] = m.score(Xj, y)
return scores
scores = fit_and_score_features(data_x_numeric.values, data_y)
pd.Series(scores, index=data_x_numeric.columns).sort_values(ascending=False)
# +
from sklearn.feature_selection import SelectKBest
from sklearn.pipeline import Pipeline
pipe = Pipeline([('encode', OneHotEncoder()),
('select', SelectKBest(fit_and_score_features, k=3)),
('model', CoxPHSurvivalAnalysis())])
# +
from sklearn.model_selection import GridSearchCV, KFold
param_grid = {'select__k': np.arange(1, data_x_numeric.shape[1] + 1)}
cv = KFold(n_splits=3, random_state=1, shuffle=True)
gcv = GridSearchCV(pipe, param_grid, return_train_score=True, cv=cv)
gcv.fit(data_x, data_y)
results = pd.DataFrame(gcv.cv_results_).sort_values(by='mean_test_score', ascending=False)
results.loc[:, ~results.columns.str.endswith('_time')]
# +
pipe.set_params(**gcv.best_params_)
pipe.fit(data_x, data_y)
encoder, transformer, final_estimator = [s[1] for s in pipe.steps]
pd.Series(final_estimator.coef_, index=encoder.encoded_columns_[transformer.get_support()])
# -
# ## Evaluating Survival Models
# +
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import train_test_split
from sksurv.datasets import load_flchain
from sksurv.linear_model import CoxPHSurvivalAnalysis
from sksurv.preprocessing import OneHotEncoder
from sksurv.util import Surv
from sksurv.metrics import (concordance_index_censored,
concordance_index_ipcw,
cumulative_dynamic_auc)
plt.rcParams['figure.figsize'] = [7.2, 4.8]
# +
import scipy.optimize as opt
def generate_marker(n_samples,
hazard_ratio,
baseline_hazard,
rnd):
# create synthetic risk score
X = rnd.randn(n_samples, 1)
# create linear model
hazard_ratio = np.array([hazard_ratio])
logits = np.dot(X, np.log(hazard_ratio))
# draw actual survival times from exponential distribution,
# refer to Bender et al. (2005), https://doi.org/10.1002/sim.2059
u = rnd.uniform(size = n_samples)
time_event = -np.log(u) / (baseline_hazard * np.exp(logits))
# compute the actual concordance in the absence of censoring
X = np.squeeze(X)
actual = concordance_index_censored(np.ones(n_samples, dtype=bool),
time_event, X)
return X, time_event, actual[0]
def generate_survival_data(n_samples,
hazard_ratio,
baseline_hazard,
percentage_cens,
rnd):
X, time_event, actual_c = generate_marker(n_samples, hazard_ratio,
baseline_hazard, rnd)
def get_observed_time(x):
rnd_cens = np.random.RandomState(0)
# draw censoring times
time_censor = rnd_cens.uniform(high=x, size=n_samples)
event = time_event < time_censor
time = np.where(event, time_event, time_censor)
return event, time
def censoring_amount(x):
event, _ = get_observed_time(x)
cens = 1.0 - event.sum() / event.shape[0]
return (cens - percentage_cens)**2
# search for upper limit to obtain the desired censoring amount
res = opt.minimize_scalar(censoring_amount,
method='bounded',
bounds = (0, time_event.max()))
# compute observed time
event, time = get_observed_time(res.x)
# upper time limit such that the probability
# of being censored is non-zero for `t > tau`
tau = time[event].max()
y = Surv.from_arrays(event=event, time=time)
mask = time < tau
X_test = X[mask]
y_test = y[mask]
return X_test, y_test, y, actual_c
def simulation(n_samples, hazard_ratio, n_repeats=100):
measures = ("censoring", "Harrel's C", "Uno's C",)
data_mean = {}
data_std = {}
for measure in measures:
data_mean[measure] = []
data_std[measure] = []
rnd = np.random.RandomState(seed=987)
# iterate over different amount of censoring
for cens in (.1, .25, .4, .5, .6, .7):
data = {"censoring": [], "Harrel's C": [], "Uno's C": [],}
for _ in range(n_repeats):
# generate data
X_test, y_test, y_train, actual_c = generate_survival_data(
n_samples, hazard_ratio,
baseline_hazard=0.1,
percentage_cens = cens,
rnd = rnd)
# estimate c-index
c_harrell = concordance_index_censored(y_test['event'], y_test['time'], X_test)
c_uno = concordance_index_ipcw(y_train, y_test, X_test)
# save results
data["censoring"].append(100. - y_test['event'].sum() * 100./y_test.shape[0])
data["Harrel's C"].append(actual_c - c_harrell[0])
data["Uno's C"].append(actual_c - c_uno[0])
# aggregate results
for key, values in data.items():
data_mean[key].append(np.mean(data[key]))
data_std[key].append(np.std(data[key], ddof=1))
data_mean = pd.DataFrame.from_dict(data_mean)
data_std = pd.DataFrame.from_dict(data_std)
return data_mean, data_std
def plot_results(data_mean, data_std, **kwargs):
index = pd.Index(data_mean['censoring'].round(3), name='mean percentage censoring')
for df in (data_mean, data_std):
df.drop('censoring', axis=1, inplace=True)
df.index=index
ax = data_mean.plot.bar(yerr=data_std, **kwargs)
ax.set_ylabel('Actual C - Estimated C')
ax.yaxis.grid(True)
ax.axhline(0.0, color='gray');
# -
hazard_ratio = 2.0
ylim = [-0.035, 0.035]
mean_1, std_1 = simulation(100, hazard_ratio)
plot_results(mean_1, std_1, ylim=ylim);
mean_2, std_2 = simulation(1000, hazard_ratio)
plot_results(mean_2, std_2, ylim=ylim);
mean_3, std_3 = simulation(2000, hazard_ratio)
plot_results(mean_3, std_3, ylim=ylim);
# ## Time-dependent Area under the ROC
x, y = load_flchain()
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.2, random_state=0)
# +
num_columns = ['age', 'creatinine', 'kappa', 'lambda']
imputer = SimpleImputer().fit(X_train.loc[:, num_columns])
X_train = imputer.transform(X_train.loc[:, num_columns])
X_test = imputer.transform(X_test.loc[:, num_columns])
# +
y_events = y_train[y_train['death']]
train_min, train_max = y_events['futime'].min(), y_events['futime'].max()
y_events = y_test[y_test['death']]
test_min, test_max = y_events['futime'].min(), y_events['futime'].max()
assert train_min <= test_min < test_max < train_max, \
"time range or test data is not within time range of training data."
# -
times = np.percentile(y['futime'], np.linspace(5, 81, 15))
print(times)
# +
def plot_cumulative_dynamic_auc(risk_score, label, color=None):
auc, mean_auc = cumulative_dynamic_auc(y_train, y_test, risk_score, times)
plt.plot(times, auc, marker='o', color=color, label=label)
plt.xlabel('days from enrollment')
plt.ylabel('time-dependent AUC')
plt.axhline(mean_auc, color=color, linestyle='--')
plt.legend()
for i, col in enumerate(num_columns):
plot_cumulative_dynamic_auc(X_test[:, i], col, color='C{}'.format(i))
ret = concordance_index_ipcw(y_train, y_test, X_test[:, i], tau = time[-1]);
# +
from sksurv.datasets import load_veterans_lung_cancer
va_x, va_y = load_veterans_lung_cancer()
cph = make_pipeline(OneHotEncoder(), CoxPHSurvivalAnalysis())
cph.fit(va_x, va_y)
va_times = np.arange(7, 183, 7)
# estimate performance on training data, thus use `va_y` twice
va_auc, va_mean_auc = cumulative_dynamic_auc(va_y, va_y, cph.predict(va_x), va_times)
plt.plot(va_times, va_auc, marker='o')
plt.axhline(va_mean_auc, linestyle='--')
plt.xlabel('days from enrollment')
plt.ylabel('time-dependent AUC')
plt.grid(True);
# -
# ## Penalized Cox Models
# +
from sksurv.datasets import load_breast_cancer
from sksurv.linear_model import CoxPHSurvivalAnalysis, CoxnetSurvivalAnalysis
from sksurv.preprocessing import OneHotEncoder
from sklearn.model_selection import GridSearchCV, KFold
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
# -
X, y = load_breast_cancer()
Xt = OneHotEncoder().fit_transform(X)
Xt.round(2).head()
# +
alphas = 10. ** np.linspace(-4, 4, 50)
coefficients = {}
cph = CoxPHSurvivalAnalysis()
for alpha in alphas:
cph.set_params(alpha=alpha)
cph.fit(Xt, y)
key = round(alpha, 5)
coefficients[key] = cph.coef_
coefficients = (pd.DataFrame
.from_dict(coefficients)
.rename_axis(index='feature', columns='alpha')
.set_index(Xt.columns))
# -
def plot_coefficients(coefs, n_highlight):
_, ax = plt.subplots(figsize=(9, 6))
n_features = coefs.shape[0]
alphas = coefs.columns
for row in coefs.itertuples():
ax.semilogx(alphas, row[1:], '.-', label=row.Index)
alpha_min = alphas.min()
top_coefs = coefs.loc[:, alpha_min].map(abs).sort_values().tail(n_highlight)
for name in top_coefs.index:
coef = coefs.loc[name, alpha_min]
plt.text(
alpha_min, coef, name + ' ',
horizontalalignment = 'right',
verticalalignment = 'center'
)
ax.yaxis.set_label_position('right')
ax.yaxis.tick_right()
ax.grid(True)
ax.set_xlabel('alpha')
ax.set_ylabel('coefficient')
plot_coefficients(coefficients, n_highlight=5);
# ## LASSO
cox_lasso = CoxnetSurvivalAnalysis(l1_ratio=1.0, alpha_min_ratio=0.01)
cox_lasso.fit(Xt, y)
coefficients_lasso = pd.DataFrame(
cox_lasso.coef_,
index=Xt.columns,
columns=np.round(cox_lasso.alphas_, 5)
)
plot_coefficients(coefficients_lasso, n_highlight=5);
# ## Elastic Net
cox_elastic_net = CoxnetSurvivalAnalysis(l1_ratio=0.9, alpha_min_ratio=0.01)
cox_elastic_net.fit(Xt, y)
coefficients_elastic_net = pd.DataFrame(
cox_elastic_net.coef_,
index=Xt.columns,
columns=np.round(cox_elastic_net.alphas_, 5)
)
plot_coefficients(coefficients_elastic_net, n_highlight=5)
# ## Choosing penalty strength α
# +
import warnings
from sklearn.exceptions import ConvergenceWarning
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
coxnet_pipe = make_pipeline(
StandardScaler(),
CoxnetSurvivalAnalysis(l1_ratio=0.9, alpha_min_ratio=0.01, max_iter=100)
)
warnings.simplefilter('ignore', ConvergenceWarning)
coxnet_pipe.fit(Xt, y)
# +
cv = KFold(n_splits=5, shuffle=True, random_state=0)
gcv = GridSearchCV(
make_pipeline(StandardScaler(), CoxnetSurvivalAnalysis(l1_ratio=0.9)),
param_grid={"coxnetsurvivalanalysis__alphas": [[v] for v in alphas]},
cv=cv,
error_score=0.5,
n_jobs=4).fit(Xt, y)
cv_results = pd.DataFrame(gcv.cv_results_)
# +
alphas = cv_results.param_coxnetsurvivalanalysis__alphas.map(lambda x: x[0])
mean = cv_results.mean_test_score
std = cv_results.std_test_score
fig, ax = plt.subplots(figsize=(9, 6))
ax.plot(alphas, mean)
ax.fill_between(alphas, mean-std, mean+std, alpha=0.15)
ax.set_xscale('log')
ax.set_xlabel('alpha')
ax.set_ylabel('concordance index')
ax.axvline(gcv.best_params_['coxnetsurvivalanalysis__alphas'][0], c='C1')
ax.axhline(0.5, color='grey', linestyle='--')
ax.grid(True);
# +
best_model = gcv.best_estimator_.named_steps['coxnetsurvivalanalysis']
best_coefs = pd.DataFrame(
best_model.coef_,
index=Xt.columns,
columns=['coefficient']
)
non_zero = np.sum(best_coefs.iloc[:, 0] != 0)
print('Number of non-zero coefficients: {}'.format(non_zero))
non_zero_coefs = best_coefs.query('coefficient != 0')
coef_order = non_zero_coefs.abs().sort_values('coefficient').index
_, ax = plt.subplots(figsize=(6, 8))
non_zero_coefs.loc[coef_order].plot.barh(ax=ax, legend=False)
ax.set_xlabel('coefficient')
ax.grid(True);
# -
# ## Survival and Cumulative Hazard Function
coxnet_pred = make_pipeline(
StandardScaler(),
CoxnetSurvivalAnalysis(l1_ratio=0.9, fit_baseline_model=True)
)
coxnet_pred.set_params(**gcv.best_params_)
coxnet_pred.fit(Xt, y)
# +
surv_fns = coxnet_pred.predict_survival_function(Xt)
time_points = np.quantile(y['t.tdm'], np.linspace(0, 0.6, 100))
legend_handles = []
legend_labels = []
_, ax = plt.subplots(figsize=(9, 6))
for fn, label in zip(surv_fns, Xt.loc[:, 'er=positive'].astype(int)):
line, = ax.step(time_points, fn(time_points), where='post',
color='C{:d}'.format(label), alpha=0.5)
if len(legend_handles) <= label:
name = 'positive' if label == 1 else 'negative'
legend_labels.append(name)
legend_handles.append(line)
ax.legend(legend_handles, legend_labels)
ax.set_xlabel('time')
ax.set_ylabel('Survival probability')
ax.grid(True);
# -
# ## Using Random Survival Forests
# +
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OrdinalEncoder
from sksurv.datasets import load_gbsg2
from sksurv.preprocessing import OneHotEncoder
from sksurv.ensemble import RandomSurvivalForest
# -
X, y = load_gbsg2()
# +
grade_str = X.loc[:, 'tgrade'].astype(object).values[:, np.newaxis]
grade_num = OrdinalEncoder(categories=[['I', 'II', 'III']]).fit_transform(grade_str)
X_no_grade = X.drop('tgrade', axis=1)
Xt = OneHotEncoder().fit_transform(X_no_grade)
Xt = np.column_stack((Xt.values, grade_num))
feature_names = X_no_grade.columns.tolist() + ['tgrade']
# +
random_state = 20
X_train, X_test, y_train, y_test = train_test_split(
Xt, y, test_size=0.25, random_state=random_state)
# -
# ## Training
rsf = RandomSurvivalForest(n_estimators=1000,
min_samples_split=10,
min_samples_leaf=15,
max_features='sqrt',
n_jobs=-1,
random_state=random_state)
rsf.fit(X_train, y_train)
rsf.score(X_test, y_test)
# ## Predicting
# +
a = np.empty(X_test.shape[0], dtype=[('age', float), ('pnodes', float)])
a['age'] = X_test[:, 0]
a['pnodes'] = X_test[:, 4]
sort_idx = np.argsort(a, order=['pnodes', 'age'])
X_test_sel = pd.DataFrame(
X_test[np.concatenate((sort_idx[:3], sort_idx[-3:]))],
columns=feature_names)
X_test_sel
# -
pd.Series(rsf.predict(X_test_sel))
# +
surv = rsf.predict_survival_function(X_test_sel, return_array=True)
for i, s in enumerate(surv):
plt.step(rsf.event_times_, s, where='post', label=str(i))
plt.xlabel('Time in days')
plt.ylabel('Survival probability')
plt.legend()
plt.grid(True);
# -
# ## Permutation-based Feature Importance
# +
import eli5
from eli5.sklearn import PermutationImportance
perm = PermutationImportance(rsf, n_iter=15, random_state=random_state)
perm.fit(X_test, y_test)
eli5.show_weights(perm, feature_names=feature_names)
# -
# ## Gradient Boosted Models
#
# ### Base Learners
#
# ### Losses
#
# Cox’s Partial Likelihood
from sklearn.model_selection import train_test_split
from sksurv.datasets import load_breast_cancer
from sksurv.ensemble import ComponentwiseGradientBoostingSurvivalAnalysis
from sksurv.ensemble import GradientBoostingSurvivalAnalysis
from sksurv.preprocessing import OneHotEncoder
# +
X, y = load_breast_cancer()
Xt = OneHotEncoder().fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(Xt, y, test_size=0.25, random_state=0)
# -
est_cph_tree = GradientBoostingSurvivalAnalysis(
n_estimators=100, learning_rate=1.0, max_depth=1, random_state=0
)
est_cph_tree.fit(X_train, y_train)
cindex = est_cph_tree.score(X_test, y_test)
print(round(cindex, 3))
# +
scores_cph_tree = {}
est_cph_tree = GradientBoostingSurvivalAnalysis(
learning_rate=1.0, max_depth=1, random_state=0
)
for i in range(1, 31):
n_estimators = i * 5
est_cph_tree.set_params(n_estimators=n_estimators)
est_cph_tree.fit(X_train, y_train)
scores_cph_tree[n_estimators] = est_cph_tree.score(X_test, y_test)
# -
x, y = zip(*scores_cph_tree.items())
plt.plot(x, y)
plt.xlabel('n_estimator')
plt.ylabel('concordance index')
plt.grid(True);
# +
scores_cph_ls = {}
est_cph_ls = ComponentwiseGradientBoostingSurvivalAnalysis(
learning_rate=1.0, random_state=0
)
for i in range(1, 31):
n_estimators = i * 10
est_cph_ls.set_params(n_estimators=n_estimators)
est_cph_ls.fit(X_train, y_train)
scores_cph_ls[n_estimators] = est_cph_ls.score(X_test, y_test)
# -
x, y = zip(*scores_cph_ls.items())
plt.plot(x, y)
plt.xlabel('n_estimator')
plt.ylabel('concordance index')
plt.grid(True);
# +
coef = pd.Series(est_cph_ls.coef_, ['Intercept'] + Xt.columns.tolist())
print('Number of non-zero coefficients:', (coef != 0).sum())
coef_nz = coef[coef != 0]
coef_order = coef_nz.abs().sort_values(ascending=False).index
coef_nz.loc[coef_order]
# -
# ## Accelerated Failure Time Model
est_aft_ls = ComponentwiseGradientBoostingSurvivalAnalysis(
loss='ipcwls', n_estimators=300, learning_rate=1.0, random_state=0
).fit(X_train, y_train)
cindex = est_aft_ls.score(X_test, y_test)
print(round(cindex, 3))
# ## Regularization
# +
n_estimators = [i * 5 for i in range(1, 21)]
estimators = {
'no regularization': GradientBoostingSurvivalAnalysis(
learning_rate=1.0, max_depth=1, random_state=0
),
'learning rate': GradientBoostingSurvivalAnalysis(
learning_rate=0.1, max_depth=1, random_state=0
),
'dropout': GradientBoostingSurvivalAnalysis(
learning_rate=1.0, dropout_rate=0.1, max_depth=1, random_state=0
),
'subsample': GradientBoostingSurvivalAnalysis(
learning_rate=1.0, subsample=0.5, max_depth=1, random_state=0
),
}
scores_reg = {k: [] for k in estimators.keys()}
for n in n_estimators:
for name, est in estimators.items():
est.set_params(n_estimators=n)
est.fit(X_train, y_train)
cindex=est.score(X_test, y_test)
scores_reg[name].append(cindex)
scores_reg = pd.DataFrame(scores_reg, index=n_estimators)
# -
ax = scores_reg.plot(xlabel='n_estimators', ylabel='concordance index')
ax.grid(True);
# +
class EarlyStoppingMonitor:
def __init__(self, window_size, max_iter_without_improvement):
self.window_size = window_size
self.max_iter_without_improvement = max_iter_without_improvement
self._best_step = -1
def __call__(self, iteration, estimator, args):
# continue training for first self.window_size iterations
if iteration < self.window_size:
return False
# compute average improvement in last self.window_size iterations.
# oob_improvement_ is the different in negative log partial likelihood
# between the previous and current iteration
start = iteration - self.window_size + 1
end = iteration + 1
improvement = np.mean(estimator.oob_improvement_[start:end])
if improvement > 1e-6:
self._best_step = iteration
return False
# stop fitting if there was no improvement
# in last max_iter_without_improvement iterations
diff = iteration - self._best_step
return diff >= self.max_iter_without_improvement
est_early_stopping = GradientBoostingSurvivalAnalysis(
n_estimators=1000, learning_rate=0.05, subsample=0.5,
max_depth=1, random_state=0
)
monitor = EarlyStoppingMonitor(25, 50)
est_early_stopping.fit(X_train, y_train, monitor=monitor)
print('Fitted base learners:', est_early_stopping.n_estimators_)
cindex = est_early_stopping.score(X_test, y_test)
print('Performance on test set', round(cindex, 3))
# +
improvement = pd.Series(
est_early_stopping.oob_improvement_,
index=np.arange(1, 1 + len(est_early_stopping.oob_improvement_))
)
ax = improvement.plot(xlabel='iteration', ylabel='oob improvement')
ax.axhline(0.0, linestyle='--', color='gray')
cutoff = len(improvement) - monitor.max_iter_without_improvement
ax.axvline(cutoff, linestyle='--', color='C3')
_ = improvement.rolling(monitor.window_size).mean().plot(ax=ax, linestyle=':')
# -
# # Linear Survival Support Vector Machine
# +
from sklearn.model_selection import ShuffleSplit, GridSearchCV
from sksurv.datasets import load_veterans_lung_cancer
from sksurv.column import encode_categorical
from sksurv.metrics import concordance_index_censored
from sksurv.svm import FastSurvivalSVM
sns.set_style("whitegrid")
# -
data_x, y = load_veterans_lung_cancer()
x = encode_categorical(data_x)
n_censored = y.shape[0] - y['Status'].sum()
print('%.1f%% of records are censored' % (n_censored / y.shape[0] * 100))
plt.figure(figsize=(9, 6))
val, bins, patches = plt.hist((y['Survival_in_days'][y['Status']],
y['Survival_in_days'][~y['Status']]),
bins=30, stacked=True)
_ = plt.legend(patches, ['Time of Death', 'Time of Censoring']);
estimator = FastSurvivalSVM(max_iter=1000, tol=1e-5, random_state=0)
def score_survival_model(model, X, y):
prediction = model.predict(X)
result = concordance_index_censored(y['Status'], y['Survival_in_days'], prediction)
return result[0]
param_grid = {'alpha': 2. ** np.arange(-12, 13, 2)}
cv = ShuffleSplit(n_splits=100, test_size=0.5, random_state=0)
gcv = GridSearchCV(estimator, param_grid, scoring=score_survival_model,
n_jobs=4, refit=False,
cv=cv)
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
gcv = gcv.fit(x, y)
round(gcv.best_score_, 3), gcv.best_params_
def plot_performance(gcv):
n_splits=gcv.cv.n_splits
cv_scores={'alpha': [], 'test_score': [], 'split': []}
order = []
for i, params in enumerate(gcv.cv_results_['params']):
name = '%.5f' % params['alpha']
order.append(name)
for j in range(n_splits):
vs = gcv.cv_results_['split%d_test_score' % j][i]
cv_scores['alpha'].append(name)
cv_scores['test_score'].append(vs)
cv_scores['split'].append(j)
df = pd.DataFrame.from_dict(cv_scores)
_, ax = plt.subplots(figsize=(11, 6))
sns.boxplot(x='alpha', y='test_score', data=df, order=order, ax=ax)
_, xtext = plt.xticks()
for t in xtext:
t.set_rotation('vertical')
plot_performance(gcv)
estimator.set_params(**gcv.best_params_)
estimator.fit(x, y)
pred = estimator.predict(x.iloc[:2])
print(np.round(pred, 3))
print(y[:2])
# ## Regression Objective
y_log_t = y.copy()
y_log_t['Survival_in_days'] = np.log1p(y['Survival_in_days'])
# +
ref_estimator = FastSurvivalSVM(rank_ratio=0.0, max_iter=1000, tol=1e-5, random_state=0)
ref_estimator.fit(x, y_log_t)
cindex = concordance_index_censored(
y['Status'],
y['Survival_in_days'],
-ref_estimator.predict(x),
)
print(round(cindex[0], 3))
# -
pred_log = ref_estimator.predict(x.iloc[:2])
pred_y = np.expm1(pred_log)
print(np.round(pred_y, 3))
# ## Kernel Survival Support Vector Machine
from sksurv.kernels import clinical_kernel
from sksurv.svm import FastKernelSurvivalSVM
kernel_matrix = clinical_kernel(data_x)
# +
kssvm = FastKernelSurvivalSVM(optimizer='rbtree', kernel='precomputed', random_state=0)
kgcv = GridSearchCV(kssvm, param_grid, scoring=score_survival_model, n_jobs=4, refit=False, cv=cv)
# -
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
kgcv = kgcv.fit(kernel_matrix, y)
round(kgcv.best_score_, 3), kgcv.best_params_
plot_performance(kgcv)
| Scikit-Survival.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, sys
currentdir = os.path.dirname(os.getcwd())
sys.path.append(currentdir)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import utilities.plotlib as pltlib
import itertools
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
# +
DIR = "C:<save folder"
df_nn = pd.read_pickle('results_FFN.pkl')
df_nn = df_nn[df_nn.loc[:, 'test_passed'] == True]
df_rnn = pd.read_pickle('results_RNN.pkl')
df_rnn = df_rnn[df_rnn.loc[:, 'test_passed'] == True]
df_combined = pd.read_pickle('results_combined.pkl')
df_combined = df_combined[df_combined.loc[:, 'test_passed'] == True]
network_results = [df_nn, df_rnn, df_combined]
labels = ['FFN', 'RNN', 'FFN + RNN']
for df in network_results:
df.history_training_data = df.history_training_data.apply(lambda x:np.array(x))
df.history_test_data = df.history_test_data.apply(lambda x:np.array(x))
# +
for name, df in zip(labels, network_results):
params_history = {'title' : ('Model accuracy of {} trained on event data'.format(name)),
'x_axis' : 'Epoch number',
'y_axis' : 'Accuracy',
'legend' : ['training data', 'test data'],
'figsize' : (4, 4),
'dpi' : 200,
'colors' : ['#662E9B', '#F86624'],
'full_y' : False}
history_training_data_mean = np.mean(df.history_training_data, axis=0)
history_test_data_mean = np.mean(df.history_test_data, axis=0)
history_training_data_std = np.std(df.history_training_data.values, axis=0)
history_test_data_std = np.std(df.history_test_data.values, axis=0)
history_error_bars = [history_training_data_std, history_test_data_std]
fig1 = pltlib.training_history_plot(history_training_data_mean,
history_test_data_mean,
params_history,
history_error_bars)
# fig1.savefig(DIR + 'training_history_{}.pdf'.format(name))
# -
for name, df in zip(labels, network_results):
fig2, ax2 = plt.subplots(figsize=(4,4), dpi=700)
background_bin_vals = np.mean(df.background_bin_vals, axis=0)
signal_bin_vals = np.mean(df.signal_bin_vals, axis=0)
background_bin_err = np.std(df.background_bin_vals.values, axis=0)
signal_bin_err = np.std(df.signal_bin_vals.values, axis=0)
bin_edges = df.bins.iloc[0]
bin_width = 0.7 * (bin_edges[1] - bin_edges[0])
bin_centers = (bin_edges[:-1] + bin_edges[1:])/2
colours = ['brown', 'teal']
ax2.bar(bin_centers, background_bin_vals, align='center', width=bin_width, alpha=0.5,
label='ttH (signal)', color=colours[0])
ax2.bar(bin_centers, signal_bin_vals, align='center', width=bin_width, alpha=0.5,
label='tt¯ (background)', color=colours[1])
ax2.set_xlabel('Label prediction')
ax2.set_ylabel('Average number of events')
ax2.set_title(name)
ax2.legend()
fig2.tight_layout()
# fig2.savefig(DIR + '{}_discriminator_plot.pdf'.format(name))
# +
class_names = ['ttH (signal)', 'tt¯ (background)']
cm_list = []
for df in network_results:
cm_list += [np.mean(df.confusion_matrix, axis=0)]
fig3 = pltlib.mulit_confusion_matrix(cm_list, class_names, labels, figsize=(4, 4), dpi=200)
fig3.tight_layout()
# fig3.savefig(DIR + 'confusion_matrices.pdf')
# -
for name, cm in zip(labels, cm_list):
params_cm = {'title' : (''),
'x_axis' : 'Predicted label',
'y_axis' : 'True label',
'class_names' : ['ttH (signal)', 'tt¯ (background)'],
'figsize' : (4, 4),
'dpi' : 200,
'colourbar' : False}
Fig = pltlib.confusion_matrix(cm, params_cm)
Fig.tight_layout()
Fig.savefig(DIR + '{}_confusion_matrix.pdf'.format(name))
# +
fig4, ax4 = plt.subplots(figsize=(4,4), dpi=300)
ax4.plot([0, 1], [0, 1], 'k--')
ax4.set_xlim([0.0, 1.005])
ax4.set_ylim([0.0, 1.005])
ax4.set_xlabel('False Positive Rate')
ax4.set_ylabel('True Positive Rate')
linestyles = ['-', '-', '--']
for name, df, ls in zip(labels, network_results, linestyles):
mean_fpr = np.mean(df.roc_fpr_vals, axis=0)
mean_tpr = np.mean(df.roc_tpr_vals, axis=0)
mean_auc = df.roc_auc.mean()
std_auc = df.roc_auc.std()
label = '{} auc = {:.3f} \u00B1 {:.3f}'.format(name, mean_auc, std_auc)
ax4.plot(mean_fpr, mean_tpr, ls, label=label)
ax4.legend()
# fig4.savefig(DIR + 'roc_curves.pdf')
# +
accuracy_results = [df.accuracy_test.to_list() for df in network_results]
roc_auc_results = [df.roc_auc.to_list() for df in network_results]
labels = ['FFN', 'RNN', 'FFN + RNN']
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
fig5, ax5 = plt.subplots(figsize=(4,4), dpi=700)
ax5.yaxis.grid(True)
boxplot1 = ax5.boxplot(accuracy_results, patch_artist=True, showfliers=False)
ax5.set_xticklabels(labels)
ax5.set_xlabel('Network architecture')
ax5.set_ylabel('Accuracy')
fig6,ax6 = plt.subplots(figsize=(4,4), dpi=300)
ax6.yaxis.grid(True)
boxplot2 = ax6.boxplot(roc_auc_results, patch_artist=True, showfliers=False)
ax6.set_xticklabels(labels)
ax6.set_xlabel('Network architecture')
ax6.set_ylabel('ROC AUC score')
# fill with colors
colors = ['pink', 'lightblue', 'lightgreen']
for bplot in (boxplot1, boxplot2):
for patch, color in zip(bplot['boxes'], colors):
patch.set_facecolor(color)
fig5.tight_layout()
fig6.tight_layout()
fig5.savefig(DIR + 'accuracy_boxplot.pdf')
fig6.savefig(DIR + 'aoc_boxplot.pdf')
# -
for name, df in zip(labels, network_results):
mean_accuracy = df.accuracy_test.mean()
std_accuracy = df.accuracy_test.std()
mean_auc_score = df.roc_auc.mean()
std_auc_score = df.roc_auc.std()
print('----------Accuracy----------')
print('{} : {} \u00B1 {}'.format(name, mean_accuracy, std_accuracy))
print('----------AOC score----------')
print('{} : {} \u00B1 {}'.format(name, mean_auc_score, std_auc_score))
| src/results/results_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Install Required Packages
# # Practical and Consistent Estimation of $f$-Divergence
# !pip install --upgrade pip
# !pip install cvxpy
# !pip install seaborn
# !pip install --upgrade tensorflow-probability
# ## Import Required Libraries
# +
import tensorflow as tf
print(f"Tensorflow version: {tf.__version__}")
gpus = tf.config.list_physical_devices('GPU')
if gpus:
print("Num GPUs Available: ", len(gpus))
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
import tensorflow_probability as tfp
from tensorflow_probability import distributions as tfd
from matplotlib import pyplot as plt
import numpy as np
import cvxpy as cp
import time
import os
import matplotlib.cm as cm
from scipy import stats
from scipy.special import logsumexp
import h5py
import seaborn as sns
from matplotlib import rc
rc('font', **{'family': 'sans-serif', 'sans-serif':['Helvetica']})
rc('text', usetex=False)
# -
# - $\chi^2$-divergence:
#
# $\chi^2(P, Q) = \int \log \Bigl(p(z) / q(z)\Bigr)^2 q(z) dz - 1$.
# ## Closed-Form Divergence Computations
# ### Helper functions
# +
def get_dims(A):
"""Gets input and latent dimensions from matrix (tensor) A.
Input dimension: #columns.
Latent dimension: #rows.
Args:
A: Parameter matrix.
Returns:
dim_latent, dim_input: A tuple containing lists representing the row and column
dimensions of the parameter matrix A.
"""
dim_latent, dim_input = A.get_shape().as_list()
return dim_latent, dim_input
def get_cov(A, std):
"""Constructs the covariance matrix with given matrix and standard deviation.
Args:
A: Parameter matrix determining the covariance matrix.
Returns:
cov: A tf tensor representing the constructed covariance matrix.
"""
dim_latent, _ = get_dims(A)
cov = tf.matmul(A, A, transpose_b=True) + std**2 * tf.eye(dim_latent)
return cov
# -
# ### Compute KL-divergence
#
# KL-divergence:
#
# $D_{KL}(Q, P) = \int \log \Bigl(q(z) / p(z)\Bigr) q(z) dz$
def compute_kl(A, b, std):
"""Computes the KL-divergence between baseline distribution Pz and distribution Qz.
Here the baseline distribution Pz is a unit Multivariate Normal distribution with
mean zero and diag(1) covariance. The distribution Qz is a Multivariate Normal
distribution with mean b and covariance AA^t + (std**2)I.
Args:
A: Parameter matrix determining covariance matrix of Qz.
b: Mean of Qz.
std: Standard deviation parameter determining covariance matrix of Qz.
Returns:
kl_divergence: A numpy array of computed KL-divergence.
"""
dim_latent, _ = get_dims(A)
# Create a Multivariate Normal distribution with a diagonal covariance and mean 0.
# The Multivariate Normal distribution is defined over R^k and parameterized by a
# length-k loc vector (aka 'mu') and a k xk scale matrix; Note that the covariance
# is given by covariance = scale @ scale.T, where @ denotes matrix multiplication
p = tfd.MultivariateNormalDiag(loc=tf.zeros(shape=(dim_latent,)),
scale_diag=tf.ones(dim_latent))
q_cov = get_cov(A, std)
q = tfd.MultivariateNormalTriL(loc=b, scale_tril=tf.linalg.cholesky(q_cov))
kl_divergence = q.kl_divergence(p).numpy()
return kl_divergence
# ### Compute Squared Hellinger distance
#
# Squared Hellinger distance:
#
# $\mathcal{H}^2(P, Q) = \int \Bigl(\sqrt{p(z)} - \sqrt{q(z)}\Bigr)^2 dz$
#
# Squared Hellinger distance is a *metric* to measure the difference between two probability distributions. It is the probabilistic analog of **Euclidean Distance**.
def compute_h2(A, b, std):
"""Computes the squared Hellinger distance between unit Gaussian Pz and Gaussian Qz
with mean b and covariance AA^t + (std**2)I.
"""
dim_latent, dim_input = get_dims(A)
Sigma1 = tf.eye(dim_latent)
Sigma2 = tf.matmul(A, A, transpose_b=True) + std**2 * tf.eye(dim_latent)
result = tf.linalg.logdet(Sigma1) / 4. + tf.linalg.logdet(Sigma2) / 4.
result -= tf.linalg.logdet(0.5 * Sigma1 + 0.5 * Sigma2) / 2.
result = tf.exp(result)
quad_form = tf.matmul(tf.linalg.inv(0.5 * Sigma1 + 0.5 * Sigma2),
tf.reshape(b, (dim_latent, -1)))
quad_form = tf.matmul(tf.reshape(b, (-1, dim_latent)), quad_form)
result *= tf.exp(-1. / 8 * quad_form)
return (2. - 2. * result[0, 0]).numpy()
# ## Estimators
def compute_ram_mc(n, m, A, b, std, f, num_iters):
"""Estimates Df(Qz, Pz) with RAM-MC estimator where Pz is a unit Gaussian and Qz
is a Gaussian with mean b and covariance AA^t + (std**2)I.
Args:
n: Number of mixture components to approximate Qz.
m: Number of MC samples to use.
A: Parameter determining covariance matrix of Qz.
b: Mean of Qz.
std: Standard deviation parameter determining covariance matrix of Qz.
f: A string representing the f-divergence type, now "KL" only.
num_iters: Number of iterations to perform.
Returns:
estimates: A numpy array of estimates, one per num_iter.
"""
dim_latent, dim_input = get_dims(A)
p = tfd.MultivariateNormalDiag(loc=tf.zeros(shape=(dim_latent,)),
scale_diag=tf.ones(dim_latent))
# Base P(X) distribution, which is a standard normal in d_input.
p_base = tfd.MultivariateNormalDiag(loc=tf.zeros(shape=(dim_input,)),
scale_diag=tf.ones(dim_input))
p_base_samples = p_base.sample(n * num_iters) # Minibatch samples from P(X).
p_base_samples = tf.reshape(p_base_samples, [num_iters, n, dim_input])
A = tf.reshape(A, [1, dim_latent, dim_input])
# Create a new tensor by replicating A num_iters times.
A = tf.tile(A, [num_iters, 1, 1])
p_base_posterior = tfd.MultivariateNormalDiag(
loc=tf.matmul(p_base_samples, A, transpose_b=True) + b,
scale_diag=std * tf.ones(dim_latent)
)
# Compute a mixture distribution. Experiment-specific parameters are indexed with
# the first dimension (num_iters) in p_base_posterior.
mixture = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(probs=[1. / n] * n),
components_distribution=p_base_posterior
)
if f == 'KL':
mc_samples = mixture.sample(m)
log_density_ratios = mixture.log_prob(mc_samples) - p.log_prob(mc_samples)
estimates = (tf.reduce_mean(log_density_ratios, axis=0)).numpy()
elif f == 'H2':
mc_samples = mixture.sample(m)
logratio = -mixture.log_prob(mc_samples) + p.log_prob(mc_samples)
estimates = 2.
estimates -= 2. * tf.exp(tf.reduce_logsumexp(0.5 * logratio, axis=0)) / m
estimates = estimates.numpy()
else:
raise ValueError("f must be one of 'KL', 'H2'.")
return estimates
# ## Plug-in Estimator
def estimator_plugin(n, m, A, b, std, f, num_iters, eps=1e-8):
"""Estimates Df(Qz, Pz) with the plugin estimator. Pz is the unit Gaussian and Qz is
Gaussian with mean b and covariance AA^t +(std**2)I. First perform kernel density
estimation of two densities, then plug in.
"""
def numpy_sample(p, n, d):
points = p.sample(n)
points = tf.reshape(points, [d, -1]).numpy()
return points
dim_latent, dim_input = get_dims(A)
p = tfd.MultivariateNormalDiag(loc=tf.zeros(shape=(dim_latent,)),
scale_diag=tf.ones(dim_latent))
q_cov = get_cov(A, std)
q = tfd.MultivariateNormalTriL(loc=b, scale_tril=tf.linalg.cholesky(q_cov))
# Repeat experiments for num_iters iterations.
results = []
for experiment in range(num_iters):
# Get i.i.d. samples from p and q to perform kernel density estimations.
p_kde_points = numpy_sample(p, n, dim_latent)
q_kde_points = numpy_sample(q, n, dim_latent)
try:
p_hat = stats.gaussian_kde(p_kde_points)
q_hat = stats.gaussian_kde(q_kde_points)
except:
results.append(np.nan)
continue
mc_points = numpy_sample(q, m, dim_latent)
try:
q_vals = q_hat.evaluate(mc_points)
p_vals = p_hat.evaluate(mc_points) + eps
log_q_vals = q_hat.logpdf(mc_points)
log_p_vals = p_hat.logpdf(mc_points) + eps
except:
results.append(np.nan)
continue
if f == 'KL':
results.append(np.mean(log_q_vals - log_p_vals))
elif f == 'H2':
logratio = log_p_vals - log_q_vals
estimate_val = 2.
estimate_val -= 2. * np.exp(logsumexp(0.5 * logratio)) / m
results.append(estimate_val)
else:
raise ValueError("f must be one of 'KL', 'H2'.")
return np.array(results)
# ## Run experiments and make plots
# ### Experiment configurations
# +
N_RANGE = [1, 500] # Sample sizes
MC_NUM = 128 # Number of Monte-Carlo samples for RAM-MC
N_EXP = 10 # Number of iterations to repeat each experiment
K = 20 # Base space dimensionality.
STD = 0.5 # Gaussian covariance noise.
BETA = 0.5 # Scale for base covariance.
D_RANGE = [1, 4, 16] # Latent space dimensionality.
LBD_MAX = 2. # Lambda range.
ROOT_PATH = '/data/'
tf.random.set_seed(345)
# Generating A and b parameters for various dimensions.
BASE_PARAMS = {}
for d in D_RANGE:
b0 = tf.random.normal(shape=(d,))
b0 /= np.linalg.norm(b0)
A0 = tf.random.normal(shape=(d, K))
A0 /= tf.linalg.norm(A0)
BASE_PARAMS[d] = {'b0': b0, 'A0': A0}
# -
# ### Run experiments
# +
RUN_RAM_MC_PLUGIN_EXPERIMENTS = True
def load_figure1_data(file_name):
data = {}
path = os.path.join(ROOT_PATH, file_name)
with h5py.File(path, 'r') as f:
for i in f:
data[int(i)] = {}
for j in f[i]:
data[int(i)][int(j)] = {}
for k in f[i][j]:
data[int(i)][int(j)][k] = list(f[i][j][k])
return data
if RUN_RAM_MC_PLUGIN_EXPERIMENTS:
ram_mc_plugin_results = {}
for d in D_RANGE:
if d not in ram_mc_plugin_results:
ram_mc_plugin_results[d] = {}
for n in N_RANGE:
print(d, n)
if n not in ram_mc_plugin_results[d]:
ram_mc_plugin_results[d][n] = {}
for lbd in np.linspace(-LBD_MAX, LBD_MAX, 51):
# Create Abase with ones on diagonal
Abase = np.zeros((d, K))
np.fill_diagonal(Abase, 1.)
Abase = tf.convert_to_tensor(Abase, tf.dtypes.float32)
Albd = Abase * BETA + lbd * BASE_PARAMS[d]['A0']
blbd = lbd * BASE_PARAMS[d]['b0']
# Compute true closed form values (only once)
if n == N_RANGE[0]:
true_kl = compute_kl(Albd, blbd, STD)
true_h2 = compute_h2(Albd, blbd, STD)
else:
true_kl = None
true_h2 = None
for dvg in ['KL', 'H2']:
if dvg not in ram_mc_plugin_results[d][n]:
ram_mc_plugin_results[d][n][dvg] = []
batch_ram_mc = compute_ram_mc(n, MC_NUM, Albd, blbd, STD,
f=dvg, num_iters=N_EXP)
batch_plugin = estimator_plugin(n, MC_NUM, Albd, blbd, STD,
f=dvg, num_iters=N_EXP)
ram_mc_plugin_results[d][n][dvg].append(
(true_kl, true_h2, batch_ram_mc, batch_plugin))
else:
ram_mc_plugin_results = load_figure1_data('ram_mc_plugin_results.hdf5')
# -
ram_mc_plugin_results[1][1]['H2'][0][3]
# +
def make_plot_figure1(ram_mc_plugin_results):
sns.set_style("white")
fig = plt.figure(figsize = (13, 3))
elinewidth = 0.4 # Width of errorbars
errorevery = 3 # Set spacing of error bars to avoid crowding of figure.
def overflow_std(array):
"""Calculates std of array, but if overflow error would occur returns a
finite number larger than the range of any axes used in plots."""
if (np.inf in array) or (np.nan in array) or any(1e20 < array):
std = 1e20
else:
std = np.std(array)
return std
for i in range(1, 7):
sp = plt.subplot(2, 3, i)
d = D_RANGE[(i - 1) % 3]
dvg = ['KL', 'H2'][int((i - 1) / 3)]
colors = cm.rainbow(np.linspace(0, 1, len(N_RANGE)))
for color, n in zip(colors, N_RANGE):
if n == N_RANGE[0]:
# Plot true values
idx = N_RANGE[0]
true_kl = np.array([el[0] for el in ram_mc_plugin_results[d][idx][dvg]])
true_h2 = np.array([el[1] for el in ram_mc_plugin_results[d][idx][dvg]])
if dvg == 'KL':
plt.plot(true_kl, color='blue', linewidth=3, label='Truth')
plt.yscale('log')
if dvg == 'H2':
plt.plot(true_h2, color='blue', linewidth=3, label='Truth')
# Plot RAM-MC estimates for N=500.
if n == 500:
mean_ram_mc_n500 = np.array(
[np.mean(el[2]) for el in ram_mc_plugin_results[d][n][dvg]])
std_ram_mc_n500 = np.array(
[np.std(el[2]) for el in ram_mc_plugin_results[d][n][dvg]])
color = 'red'
plt.errorbar(range(51),
mean_ram_mc_n500,
errorevery=errorevery,
yerr=std_ram_mc_n500,
elinewidth=elinewidth,
color=color, label='RAM-MC estimator, N=' + str(n),
marker="^", markersize=5, markevery=10)
# Plot plug-in estimates
if n == 500:
mean_plugin = np.array(
[np.mean(el[3]) for el in ram_mc_plugin_results[d][n][dvg]])
std_plugin = np.array(
[overflow_std(el[3]) for el in ram_mc_plugin_results[d][n][dvg]])
color = 'darkorange'
plt.errorbar(range(51),
mean_plugin,
errorevery=errorevery,
yerr=std_plugin,
elinewidth=elinewidth,
color=color, label='Plug-in estimator, N=' + str(n),
marker="s", markersize=5, markevery=10)
# Plot RAM-MC with N=1.
if n == N_RANGE[0]:
color = 'black'
mean_ram_mc1 = np.array(
[np.mean(el[2]) for el in ram_mc_plugin_results[d][n][dvg]])
std_ram_mc1 = np.array(
[np.std(el[2]) for el in ram_mc_plugin_results[d][n][dvg]])
plt.errorbar(range(51) + 0.3 * np.ones(51),
mean_ram_mc1,
errorevery=errorevery,
yerr=std_ram_mc1,
elinewidth=elinewidth,
color=color, label='RAM-MC estimator, N=1',
marker="o", markersize=5, markevery=10)
if dvg == 'KL':
plt.ylim((0.03, 15))
if dvg == 'H2':
plt.ylim((0., 2))
sp.axes.get_xaxis().set_ticklabels([])
if d != 1:
sp.axes.get_yaxis().set_ticklabels([])
else:
sp.axes.tick_params(axis='both', labelsize=15)
if i < 4:
plt.title("d = {}".format(d), fontsize=18)
if i == 1:
plt.ylabel('KL', fontsize=18)
if i == 4:
plt.ylabel(r'$\mathcal{H}^2$', fontsize=18)
ax = fig.axes[1]
handles, labels = ax.get_legend_handles_labels()
labels, handles = zip(*sorted(zip(labels, handles), key=lambda t: t[0]))
fig.legend(handles, labels, loc='lower center', bbox_to_anchor=(0.51, 1.0),
ncol=5, fancybox=True, shadow=True, fontsize=12, frameon=True)
plt.tight_layout()
plt.show()
make_plot_figure1(ram_mc_plugin_results)
# -
| .ipynb_checkpoints/Practical and Consistent Estimation of f-Divergences-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VinceDeilord/CPEN21A-ECE-2-1/blob/main/Demo1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NhpnzEsnj-FD"
# #Intro to Python Programming
# + colab={"base_uri": "https://localhost:8080/"} id="s2GJSCmCj1b5" outputId="78fbe835-57fa-46ea-ff66-763eea140b84"
#Python Indention
if 5>2:
print("Five is greater than two")
# + [markdown] id="dscv47OBj8nA"
# #Python Variable
# + colab={"base_uri": "https://localhost:8080/"} id="yFA5FTWlk7Cy" outputId="cefe3076-b5fb-4f71-bde8-a22a6d6096c5"
x=1
a, b = 0,1
a,b,c="zero","one","two"
print(x)
print(a)
print(b)
print(c)
# + colab={"base_uri": "https://localhost:8080/"} id="0menGkAQlYqt" outputId="b504dd54-ba6c-4d38-ace4-b8834d9ce2a0"
d="Sally" #This is a string
D="Ana"
print(d)
e="John"
print(e)
print(d)
# + colab={"base_uri": "https://localhost:8080/"} id="g1yCC43clwL-" outputId="c4465983-f90b-4ba2-cfe6-09d95794fb5e"
print(type(d))
print(type(x))
# + [markdown] id="09cGPfYyk7e5"
# #Casting
# + colab={"base_uri": "https://localhost:8080/"} id="IvTLMJMek-7U" outputId="1b0b933b-930f-4b2b-86e1-095b76d4deb2"
f=float(4)
print(f)
g=int(5)
print(g)
# + [markdown] id="l2JnWo1ck_j2"
# #Multiple Variables with One Value
# + colab={"base_uri": "https://localhost:8080/"} id="vLWQFZPzlDEn" outputId="d06d74ce-06a4-4ca6-9741-50448326b594"
x = y = z = "four"
print(x)
print(y)
print(z)
# + colab={"base_uri": "https://localhost:8080/"} id="c1UqnCTcmM3e" outputId="f6e3c29b-93c1-40d9-848b-b14e2bbb0e36"
x= "enjoying"
print("Python Programming is"
" " +x)
# + [markdown] id="J9J1pP7dmFr2"
# #Operations in Python
# + colab={"base_uri": "https://localhost:8080/"} id="2lKlgdnomI-1" outputId="79b15842-971b-4371-a495-f3418cc0c1a1"
x=5
y=7
x+=y #This is the same as x = x + y
print(x+y)
print(x*y)
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="FpKhvdLamSGl" outputId="87b450f2-4888-447e-d8bb-a1ac05a44ff3"
x=5
y=7
not(x>y or y==x)
# + colab={"base_uri": "https://localhost:8080/"} id="TlkE7nWLmSb9" outputId="496301c8-b2e5-47b9-b211-0f8847cd1253"
x is y
# + colab={"base_uri": "https://localhost:8080/"} id="UtzLVutYmVlV" outputId="d7c6a33a-0cf8-44de-a5d7-0591cf567bee"
x is not y
| Demo1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Creating a script to parse blast files of custom or standard tabular format
# **Started with standard function to parse blast files of set format**
# function for parsing blast file
def old_blast_parser(blastfile, tab=None):
"""parse tabular blast files to retrieve all information for downstream"""
blast = open(blastfile)
if tab is None: #'standard': # alternative add later
std = {'qseqid':0, 'sseqid':1, 'pident':2, 'length':3, 'mismatch':4, 'gapopen':5,'qstart':6, 'qend':7, 'sstart':8, 'send':9, 'evalue':10, 'bitscore':11}
fmt_list = ['qseqid', 'sseqid', 'pident', 'length', 'mismatch', 'gapopen', 'qstart', 'qend', 'sstart', 'send', 'evalue', 'bitscore']
#else:
integers = ['qlen','slen','qstart','qend','sstart','send','length','nident','mismatch','positive','gapopen','gaps','qcovs','qcovhsp']
floaters = ['bitscore','score','pident','ppos']
ranges = {}
rng_index = {}
#len_min = 100
#min_pct = 30
hit_index = {}
hit_dict = {}
hitlist = []
fld_range = len(fmt_list)
field_dict = {}
for f in range(0,fld_range):
field_dict[fmt_list[f]]=f
ct = 0
for hit in blast:
hit = hit.strip('\n')
parts = hit.split('\t')
## have to change this to be flexible as to input format, start with standard format
for fld in range(0,fld_range):
fld_name = fmt_list[fld]
fld_value = field_dict[fld_name]
ranges.setdefault(hitnum, {})['qseqid']=query
query = parts[std['qseqid']]
#print(query)
subject = parts[std['sseqid']]
pct_ident = float(parts[std['pident']])
aln_length = int(parts[std['length']])
num_mismatch = int(parts[std['mismatch']])
gap_open = int(parts[std['gapopen']])
query_start = int(parts[std['qstart']])
query_end = int(parts[std['qend']])
substart = int(parts[std['sstart']])
subend = int(parts[std['send']])
e_val = parts[std['evalue']]
bit_score = parts[std['bitscore']]
#if pct_ident > min_pct: # filtering will happen outside script
if query in hitlist:
ct += 1
else:
hitlist.append(query)
ct = 0
hitnum = query+';hit'+str(ct)
rng_index.setdefault(query, []).append(hitnum)
ranges.setdefault(hitnum, {})['qseqid']=query
ranges.setdefault(hitnum, {})['sseqid']=subject
ranges.setdefault(hitnum, {})['pident']=pct_ident
ranges.setdefault(hitnum, {})['length']=aln_length
ranges.setdefault(hitnum, {})['mismatch']=num_mismatch
ranges.setdefault(hitnum, {})['gapopen']=gap_open
ranges.setdefault(hitnum, {})['qstart']=query_start
ranges.setdefault(hitnum, {})['qend']=query_end
ranges.setdefault(hitnum, {})['sstart']=substart
ranges.setdefault(hitnum, {})['send']=subend
ranges.setdefault(hitnum, {})['evalue']=e_val
ranges.setdefault(hitnum, {})['bitscore']=bit_score
return(rng_index, ranges)
# **The parsing function below will handle any custom tabular format**
# function for parsing blast file, default is standard tab format
def blast_parser(blastfile, tab='standard'):
"""parse tabular blast files to retrieve all information for downstream"""
blast = open(blastfile)
if tab is 'standard': #'standard': # alternative add later
fmt_list = ['qseqid', 'sseqid', 'pident', 'length', 'mismatch', 'gapopen', 'qstart', 'qend', 'sstart', 'send', 'evalue', 'bitscore']
else:
fmt_list = tab
integers = ['qlen','slen','qstart','qend','sstart','send','length','nident','mismatch','positive','gapopen','gaps','qcovs','qcovhsp']
floaters = ['bitscore','score','pident','ppos','evalue']
ranges = {}
rng_index = {}
hit_index = {}
hit_dict = {}
hitlist = []
# adding list of queries to pass to tuple, to preserve order of queries
query_list = []
hit_order = []
fld_range = len(fmt_list)
ct = 0
for hit in blast:
hit = hit.strip('\n')
parts = hit.split('\t')
for fld in range(0,fld_range):
fld_name = fmt_list[fld]
# differentiate between field types
if fld_name in integers:
fld_value = int(parts[fld])
elif fld_name in floaters:
fld_value = float(parts[fld])
else:
fld_value = parts[fld]
if fld == 0:
if fld_value in hitlist:
ct += 1
else:
hitlist.append(fld_value)
ct = 0
hitnum = str(fld_value)+';hit'+str(ct)
rng_index.setdefault(str(fld_value), []).append(hitnum)
if fld_value not in query_list:
query_list.append(fld_value)
ranges.setdefault(hitnum, {})[fld_name]=fld_value
if hitnum not in hit_order:
hit_order.append(hitnum)
return(rng_index, ranges, query_list, hit_order)
## testing with a nonstandard format
std_fmt = ['qseqid', 'sseqid', 'length', 'pident', 'mismatch', 'qcovs', 'evalue', 'bitscore', 'qstart', 'qend', 'sstart', 'send', 'gapopen']
blast_results = blast_parser('example_blastn_custom_output.outfmt6', std_fmt)
# returned set with two dictionaries: first has a list of hits for each query
blast_results[0]['MA_10046850g0010']
# the second dictionary has all information for each hit
blast_results[1]['MA_10429268g0010;hit0']
query_ls = blast_results[3]
query_ls[1]
# write script to filter blast_parser output to file as table
def get_parameters(parameter_file):
para_file = open(parameter_file, 'r')
para_dict = {}
for line in para_file:
line = line.strip('\n')
if line.startswith('#'):
continue
else:
line_parts = line.split('=')
para_name = line_parts[0]
para_value =line_parts[1]
if para_name == 'FORMAT':
fieldlist = para_value.split(' ')
para_dict['FORMAT']=fieldlist
elif para_name == 'MIN_LENGTH':
para_dict['MIN_LENGTH']=int(para_value)
elif para_name == 'MIN_PCT_ID':
para_dict['MIN_PCT_ID']=int(para_value)
elif para_name == 'QCOV':
para_dict['QCOV']=int(para_value)
elif para_name == 'MAX_MISMATCH':
para_dict['MAX_MISMATCH']=int(para_value)
return para_dict
paras1 = get_parameters('parameters_file_template.txt')
paras1
fmt1 = paras1['FORMAT']
fmt1
# +
# a script that implements the blast parser, filters the result for length and pident, and outputs to summary
blastout = open('blast_results_filtered.txt', 'w')
blastfile = ('example_blastn_custom_output.outfmt6')
parameter_file = ('parameters_file_template.txt')
blast_parameters = get_parameters(parameter_file)
blast_tab_format = blast_parameters['FORMAT']
min_length = blast_parameters['MIN_LENGTH']
min_pct = blast_parameters['MIN_PCT_ID']
blast_results = blast_parser(blastfile, blast_tab_format)
query_index = blast_results[0]
hit_results = blast_results[1]
for key, value in query_index.iteritems():
for hit in value: # now loops through lists of hits for this query
hit_dict = hit_results[hit]
if hit_dict['length'] > min_length:
if hit_dict['pident'] > min_pct:
# if it passes, write all fields to new file
# use blast_tab_format to put values in order
blaststring = ''
for field in blast_tab_format:
#print field
field_value = str(hit_dict[field])
blaststring = blaststring+field_value+'\t'
blastwrite = blaststring+'\n'
blastwrite = blastwrite.replace('\t\n','\n')
blastout.write(blastwrite)
blastout.close()
# -
# have to set it up so all conditions can be tested simultaneously. Not nested 'if' conditions, but test each one, and add them up (if x > minimum: count += 1 or something), then total of all condition tests must add up to number of conditions from parameters
# To have a variable number of conditions to test to filter blast results: create dictionary from parameters file, and can use length of dictionary to set number of conditions
# match the parameter to blast output fields
field_match = {'MIN_LENGTH':'length','MIN_PCT_ID':'pident','MAX_MISMATCH':'mismatch','EVALUE':'evalue','SUBJECT_ID':'sseqid'}
# +
# a script that implements the blast parser, filters the result for length and pident, and outputs to summary
# modified
# establish how to test conditions
mins = ['MIN_LENGTH','MIN_PCT_ID']
maxes = ['MAX_MISMATCH','EVALUE']
matches = ['SUBJECT_ID']
blastout = open('blast_results_filtered_newer.txt', 'w')
blastfile = ('example_blastn_custom_output.outfmt6')
parameter_file = ('parameters_file_template.txt')
blast_parameters = get_parameters(parameter_file)
# get format
blast_tab_format = blast_parameters['FORMAT']
# take out format for filtering
blast_parameters.pop('FORMAT')
# get number of conditions to test
num_conditions = len(blast_parameters)
# get blast results
blast_results = blast_parser(blastfile, blast_tab_format)
query_index = blast_results[0]
hit_results = blast_results[1]
for key, value in query_index.iteritems():
for hit in value: # now loops through lists of hits for this query
conditions_reached = 0
hit_dict = hit_results[hit]
# now test all conditions separately
for k,v in blast_parameters.iteritems():
blast_field = field_match[k]
if k in mins:
if hit_dict[blast_field] > v:
conditions_reached += 1
elif k in maxes:
if hit_dict[blast_field] < v:
conditions_reached += 1
elif k in matches:
if hit_dict[blast_field] == v:
conditions_reached += 1
if conditions_reached == num_conditions:
# use blast_tab_format to put values in order
blaststring = ''
for field in blast_tab_format:
#print field
field_value = str(hit_dict[field])
blaststring = blaststring+field_value+'\t'
blastwrite = blaststring+'\n'
blastwrite = blastwrite.replace('\t\n','\n')
blastout.write(blastwrite)
blastout.close()
print(blast_parameters)
# -
dict1 = {'a':1,'b':2,'c':3}
dict2 = dict1.pop('c')
dict1
a = 1.222
b = 1.223
b > a
# ### Rewrite the parameter filter as a function
def filter_blast(blast_result_dict, blast_parameters):
"""Filter a blast result dictionary based on variable parameters"""
mins = ['MIN_LENGTH','MIN_PCT_ID']
maxes = ['MAX_MISMATCH','EVALUE']
matches = ['SUBJECT_ID']
filtered_blast = {}
blast_parameters.pop('FORMAT')
num_conditions = len(blast_parameters)
query_index = blast_result_dict[0]
hit_results = blast_result_dict[1]
for key, value in query_index.iteritems():
for hit in value: # now loops through lists of hits for this query
conditions_reached = 0
hit_dict = hit_results[hit]
# now test all conditions separately
for k,v in blast_parameters.iteritems():
blast_field = field_match[k]
if k in mins:
if hit_dict[blast_field] > v:
conditions_reached += 1
elif k in maxes:
if hit_dict[blast_field] < v:
conditions_reached += 1
elif k in matches:
if hit_dict[blast_field] == v:
conditions_reached += 1
if conditions_reached == num_conditions:
filtered_blast[hit]=hit_dict
return filtered_blast
# +
# run script from functions
blastout = open('blast_results_filtered_from_function2.txt', 'w')
blastfile = ('example_blastn_custom_output.outfmt6')
parameter_file = ('parameters_file_template.txt')
blast_parameters = get_parameters(parameter_file)
# get format
blast_tab_format = blast_parameters['FORMAT']
# have to put default format parameters here, in case none provided
# get blast results
blast_results = blast_parser(blastfile, blast_tab_format)
hit_order = blast_results[3]
# filter blast results
filt_blast_results = filter_blast(blast_results, blast_parameters)
# write filtered results to file
for hit in hit_order:
if hit in filt_blast_results:
hit_values = filt_blast_results[hit]
#for key,value in filt_blast_results.iteritems():
blaststring = ''
for field in blast_tab_format:
#print field
field_value = str(hit_values[field])
blaststring = blaststring+field_value+'\t'
blastwrite = blaststring+'\n'
blastwrite = blastwrite.replace('\t\n','\n')
blastout.write(blastwrite)
blastout.close()
# -
# just try to order the dictionary, so they all come out
#filt_blast_results
| notebooks/blast_file_parsing_dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# earth engine
import ee
import geemap.eefolium as geemap
#from geemap import geojson_to_ee, ee_to_geojson
# analysis and mapping
import pandas as pd
import numpy as np
import geopandas as gpd
import folium
#import json
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib import rcParams
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
from pprint import pprint
# file management stuff
import datetime
import glob
import urllib
import zipfile
from zipfile import ZipFile
# -
# ee.Authenticate()
ee.Initialize()
# +
# Define a method for displaying Earth Engine image tiles on a folium map.
def add_ee_layer(self, ee_object, vis_params, name):
try:
# display ee.Image()
if isinstance(ee_object, ee.image.Image):
map_id_dict = ee.Image(ee_object).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles = map_id_dict['tile_fetcher'].url_format,
attr = 'Google Earth Engine',
name = name,
overlay = True,
control = True
).add_to(self)
# display ee.ImageCollection()
elif isinstance(ee_object, ee.imagecollection.ImageCollection):
ee_object_new = ee_object.mosaic()
map_id_dict = ee.Image(ee_object_new).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles = map_id_dict['tile_fetcher'].url_format,
attr = 'Google Earth Engine',
name = name,
overlay = True,
control = True
).add_to(self)
# display ee.Geometry()
elif isinstance(ee_object, ee.geometry.Geometry):
folium.GeoJson(
data = ee_object.getInfo(),
name = name,
overlay = True,
control = True
).add_to(self)
# display ee.FeatureCollection()
elif isinstance(ee_object, ee.featurecollection.FeatureCollection):
ee_object_new = ee.Image().paint(ee_object, 0, 2)
map_id_dict = ee.Image(ee_object_new).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles = map_id_dict['tile_fetcher'].url_format,
attr = 'Google Earth Engine',
name = name,
overlay = True,
control = True
).add_to(self)
except:
print("Could not display {}".format(name))
# Add EE drawing method to folium.
folium.Map.add_ee_layer = add_ee_layer
# Add EE drawing method to folium.
#folium.Map.add_ee_layer = add_ee_layer
folium.plugins.DualMap.add_ee_layer = add_ee_layer
# -
# Function to sample values by HOLC zones
def sample_holc_layer(zones, dataset, scale):
resultList = []
for feat in zones.getInfo()['features']:
name = feat['properties']['level_0']
print(name)
region = ee.Feature(zones.filter(ee.Filter.eq('level_0', name)).first())
# Reduce the region. The region parameter is the Feature geometry.
meanDictionary = dataset.reduceRegion(**{'reducer': ee.Reducer.mean(),'geometry': region.geometry(),'scale': scale,'maxPixels': 1e9})
# The result is a Dictionary. Print it.
#resultDict[name]=meanDictionary.getInfo()
resultList.append(meanDictionary.getInfo())
pprint(meanDictionary.getInfo())
datadf = pd.DataFrame(resultList)
datadf.reset_index(inplace=True)
datadf['joinid']=datadf['index'].apply(str)
return datadf
# +
url = r'https://dsl.richmond.edu/panorama/redlining/static/fullshpfile.zip'
dwnfile = url.split('/')[-1]
# Copy a network object to a local file
urllib.request.urlretrieve(url, dwnfile)
# specifying the zip file name
file_name = dwnfile
# opening the zip file in READ mode
with ZipFile(file_name, 'r') as zf:
# printing all the contents of the zip file
zf.printdir()
# extracting all the files
print('Extracting all the files now...')
zf.extractall()
print('Done!')
del zf
# -
from google.colab import drive
drive.mount('/content/drive')
| EarthEngine/HOLC Grades and Environmental Conditions with EarthEngine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="l_O6aR86YcW3"
# # Using SVM to predict Multi-XRF
# + [markdown] id="bQtYXR80rIlH"
# TL;DR, RBF is best kernal. Poly does not work.
# + [markdown] id="KkdddlNSYcW-"
# Put together by <NAME>, <EMAIL>, all errors are mine
# + [markdown] id="UE4eky2QYcXB"
# If you are interested in graident boosting, here is a good place to start: https://xgboost.readthedocs.io/en/latest/tutorials/model.html
#
# This is a supervised machine learning method.
# + id="qC2ECegCYcXD"
# If you have installation questions, please reach out
import pandas as pd # data storage
import numpy as np # math and stuff
import sklearn
import datetime
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score, KFold
from sklearn.metrics import mean_squared_error
from sklearn.utils.class_weight import compute_sample_weight
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.multioutput import MultiOutputRegressor
from sklearn.metrics import median_absolute_error, max_error, mean_squared_error
from sklearn import svm
from sklearn.svm import SVR
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt # plotting utility
# + id="WNiabSVfYjTE" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1613420616857, "user_tz": 420, "elapsed": 275, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="1baaf99b-6b3d-480f-85ae-2cd586486d14"
from google.colab import drive
drive.mount('/content/drive')
# + id="Hk1AsPnSYcXQ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1613420617078, "user_tz": 420, "elapsed": 492, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="b097fc6c-9597-4841-af89-9d6a9f410503"
df = pd.read_csv('drive/My Drive/1_lewis_research/core_to_wl_merge/Merged_dataset_inner_imputed_12_21_2020.csv')
# + id="Ws9xTzdwYzgX" colab={"base_uri": "https://localhost:8080/", "height": 469} executionInfo={"status": "ok", "timestamp": 1613420617286, "user_tz": 420, "elapsed": 694, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="55b40cda-5acb-4fd3-b170-8a2bf5808e16"
df = df.drop(['Unnamed: 0', 'Unnamed: 0.1', 'LiveTime2','ScanTime2', 'LiveTime1','ScanTime1',
'ref_num', 'API', 'well_name', 'sample_num' ], axis=1)
print(df.columns.values)
df.describe()
# + id="fogHW-tlzXH1"
df = df[df['Ca']>0]
# + id="qWpB6BHQ0PY0"
# df = df[df.USGS_ID != 'E997']
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="u1GhriIzzkFx" executionInfo={"status": "ok", "timestamp": 1613420617529, "user_tz": 420, "elapsed": 927, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="4d0480f6-1828-4df6-9676-35711c3c2a02"
df.describe()
# + [markdown] id="rKN-0n34YcXP"
# ## Loading in Dataset
# + id="91nAGubNYcYo"
dataset = df[[
'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID', 'GR_smooth', 'PE_smooth',
'Ca', 'Si', 'Ti', 'Mg', 'Al'
]]
# + id="HhYFK3K6YcYy" colab={"base_uri": "https://localhost:8080/", "height": 163} executionInfo={"status": "ok", "timestamp": 1613420617530, "user_tz": 420, "elapsed": 922, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="ce92d31d-a910-470d-8098-821a2d8baef2"
dataset.head(3)
# + id="MxCYJ2GVYcZA"
X = dataset[[ 'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID', 'GR_smooth', 'PE_smooth']]
Y = dataset[['Ca', 'Si', 'Ti', 'Mg', 'Al']]
# + [markdown] id="rfNwgw_MYcZJ"
# ## Starting to set up the ML model params
# + id="q_Zq4vu_YcZK"
seed = 7 # random seed is only used if you want to compare exact answers with friends
test_size = 0.25 # how much data you want to withold, .15 - 0.3 is a good starting point
X_train, X_test, y_train, y_test = train_test_split(X.values, Y.values, test_size=test_size)
# + colab={"base_uri": "https://localhost:8080/"} id="cdy8A1nA1Nf9" executionInfo={"status": "ok", "timestamp": 1613420618653, "user_tz": 420, "elapsed": 2038, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="9e7fa2ec-1d86-4681-e524-643b54976a1c"
multioutputregressor = MultiOutputRegressor(svm.SVR(kernel='rbf',
C=5,
max_iter=1000,
tol=0.01,
gamma= 0.1,
degree = 2))
multioutputregressor.fit(X_train, y_train)
# + id="D2sM69dN15AI"
preds = multioutputregressor.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="F9IVWgOQ2Bzp" executionInfo={"status": "ok", "timestamp": 1613420618855, "user_tz": 420, "elapsed": 2235, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="2ba57884-f146-4d6f-ec78-78579229bd38"
preds
# + id="RD4T7LkzbawD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1613420618856, "user_tz": 420, "elapsed": 2233, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="5f4b35bf-e467-4ec0-cdd9-2fd679447a58"
rmse2 = mean_squared_error(y_test, preds, squared=False)
print("Mean Squared Error: %f" % (rmse2))
MAE2 = median_absolute_error(y_test, preds)
print("Median Abs Error: %f" % (MAE2))
# + id="juV8amnc3fzq" colab={"base_uri": "https://localhost:8080/", "height": 80} executionInfo={"status": "ok", "timestamp": 1613420618856, "user_tz": 420, "elapsed": 2230, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="1af17a85-9f12-4a5e-b953-3f7f5531e01f"
x = datetime.datetime.now()
d = {'target': ['MultiXRF'],
'MSE': [rmse2],
'MAE': [MAE2],
'day': [x.day],
'month':[x.month ],
'year':[x.year ],
'model':['SVM-RBF'],
'version':[sklearn.__version__]}
results = pd.DataFrame(data=d)
results.to_csv('drive/My Drive/1_lewis_research/analysis/experiments/svm/svm_results/MultiXRF_SVM.csv')
results
| svm/old_notebooks/SVM_regression_Multi_output_XRF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [<NAME>](https://orcid.org/0000-0001-7225-9992),
# Professorship Signal Theory and Digital Signal Processing,
# [Institute of Communications Engineering (INT)](https://www.int.uni-rostock.de/),
# Faculty of Computer Science and Electrical Engineering (IEF),
# [University of Rostock, Germany](https://www.uni-rostock.de/en/)
#
# # Tutorial Signals and Systems (Signal- und Systemtheorie)
#
# Summer Semester 2021 (Bachelor Course #24015)
#
# - lecture: https://github.com/spatialaudio/signals-and-systems-lecture
# - tutorial: https://github.com/spatialaudio/signals-and-systems-exercises
#
# WIP...
# The project is currently under heavy development while adding new material for the summer semester 2021
#
# Feel free to contact lecturer [<EMAIL>](https://orcid.org/0000-0002-3010-0294)
#
# ## Übung / Exercise 5
import numpy as np
import matplotlib as mpl
from mpl_toolkits.mplot3d import Axes3D
#from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
# # Lowpass 2nd Order
# +
N = 2**9
re_s = np.linspace(-5, 5, N)
im_s = np.linspace(-5, 5, N)
sigma, omega = np.meshgrid(re_s, im_s, sparse=False, indexing='xy')
s = sigma+1j*omega
soo1 = -3/4+1j
soo2 = -3/4-1j
x = 1 / (s - soo1) * 1 / (s - soo2)
xft = 1 / (1j*omega - soo1) * 1 / (1j*omega - soo2)
xa = 20*np.log10(np.abs(x))
xfta = 20*np.log10(np.abs(xft))
xa_max = np.max(xa)
xa_min = np.min(xa)
print(xa_max, xa_min)
fig = plt.figure(figsize=(6, 5))
ax = fig.add_subplot(projection='3d')
Ncol = 72//6
col_tick = np.linspace(-36, 36, Ncol, endpoint=False)
cmap = mpl.cm.get_cmap('cividis')
norm = mpl.colors.BoundaryNorm(col_tick, cmap.N)
surf = ax.plot_surface(sigma, omega, xa,
cmap=cmap,
norm=norm,
rstride=5, cstride=5, linewidth=0, alpha=1)
ax.plot3D(sigma[:, N//2], omega[:, N//2],
xfta[:, N//2], 'darkred', lw=3, alpha=1)
cbar = fig.colorbar(surf, ax=ax, ticks=col_tick[::Ncol//10],
label=r'$|H(s)|$ in dB', pad=0.15)
# TBD: location='left' in newer patplotlib version
ax.set_xlabel(r'$\Re(s)$')
ax.set_ylabel(r'$\Im(s)$')
ax.set_zlabel(r'$|H(s)|$ in dB')
ax.view_init(azim=-50, elev=45)
ax.set_xlim(-5, 5)
ax.set_xticks(np.arange(-5, 6, 1))
ax.set_ylim(-5, 5)
ax.set_yticks(np.arange(-5, 6, 1))
ax.set_zlim(-36, 36)
ax.set_zticks(np.arange(-36, 30+6, 6))
ax.set_zticklabels(['-36', ' ', '-24', ' ', '-12', ' ',
'0', ' ', '12', ' ', '24', ' '])
plt.savefig('bodeplot_example_approximations_pzmaps3D_44EB4169E9.pdf')
# -
# ## Copyright
#
# This tutorial is provided as Open Educational Resource (OER), to be found at
# https://github.com/spatialaudio/signals-and-systems-exercises
# accompanying the OER lecture
# https://github.com/spatialaudio/signals-and-systems-lecture.
# Both are licensed under a) the Creative Commons Attribution 4.0 International
# License for text and graphics and b) the MIT License for source code.
# Please attribute material from the tutorial as *<NAME>,
# Continuous- and Discrete-Time Signals and Systems - A Tutorial Featuring
# Computational Examples, University of Rostock* with
# ``main file, github URL, commit number and/or version tag, year``.
| laplace_system_analysis/bodeplot_example_approximations_pzmaps3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing cell states
#
# The purpose of this notebook is to compare the cell states for different examples. By this I mean taking the plots I generate, and instead looking at them numerically.
import numpy as np
from pathlib import Path
import pandas as pd
import csv
import seaborn as sns
from matplotlib import pyplot as plt
# ## Load an example
class Config:
n_terms = 4
n_digits = 2
# +
cell_state_dir = Path(f'../Code/experiments/cell_states/{Config.n_terms}term_{Config.n_digits}dig')
cell_states = np.load(cell_state_dir / Path('cell_states.npy'), allow_pickle=True)
print(f'{cell_states.shape[0]} samples, each with shape {cell_states.shape[1:]}')
input_seqs = []
with open(cell_state_dir / Path('input.csv')) as f:
csv_reader = csv.reader(f)
for row in csv_reader:
# When reading back the file it doesn't record the newline, so include it
input_seqs.append(row[-1] + '\n')
# -
print(f'Working with {input_seqs[0]}')
cell_df = pd.DataFrame(cell_states[0])
cell_df.columns = [f'Cell {x}' for x in cell_df.columns]
cell_df
# +
corr = cell_df.corr()
plt.figure(figsize=(20,20))
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
# -
| Notebooks/Comparing cell states.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Back to the main [Index](../index.ipynb)
# # Ontology Search
# Search in ontologies via the bioservices WebServices.
from __future__ import print_function, division
# ## Query KEGG.Reaction and ChEBI
# Input search term and click **search**. The search can take some time due to the WebServices.
import tellurium as te
from tellurium.notebooks import OntologySearch
form = OntologySearch()
| examples/notebooks/widgets/widgets_ontology_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Visualization
# Data visualization refers to the techniques used to communicate data or information by encoding it as visual objects (e.g., points, lines or bars) contained in graphics. The goal is to communicate information clearly and efficiently to users. See [Wikipedia's Data Visualization Page](https://en.wikipedia.org/wiki/Data_visualization)
# ## Python Plotting Libraries
#
# + [Matplotlib](https://matplotlib.org/)
# + [Pandas Visualization](https://pandas.pydata.org/pandas-docs/stable/user_guide/visualization.html)
# + [Seaborn](https://seaborn.pydata.org/)
# + [ggplot](http://ggplot.yhathq.com/)
# + [plotly](https://plot.ly/python/)
# + [Boken](http://bokeh.pydata.org/en/latest/)
# # Dataset
# 1. [Wine Reviews](https://www.kaggle.com/zynicide/wine-reviews/)
# 2. [Iris Data Set](https://archive.ics.uci.edu/ml/datasets/Iris)
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# +
import re
attributeRE = re.compile(r"7.\s+Attribute Information:")
missingValueRE = re.compile("8. Missing Attribute Values:")
with open("datasets\\Iris\\iris.names", "r") as f:
lines = f.readlines()
start_idx = np.nan
idx = 0
for line in lines:
# print(line)
if attributeRE.search(line):
start_idx = idx + 1
if missingValueRE.search(line):
stop_idx = idx
idx = idx + 1
#print(start_idx, lines[start_idx])
#print(stop_idx, lines[stop_idx])
attributeLineRE = re.compile("^\s*\d+.\s+([:\w\s]+)$")
fields = []
for line in lines[start_idx:stop_idx]:
line = line.rstrip()
m = attributeLineRE.match(line)
if m:
field = m.group(1)
field = field.replace(':', '').replace(' ', '_')
fields.append(field)
print(fields)
with open("datasets\\Iris\\iris.csv", "w") as f:
f.write(",".join(fields))
f.write("\n")
with open("datasets\\Iris\\iris.data", "r") as inf:
lines = inf.readlines()
f.writelines(lines)
# -
iris = pd.read_csv("datasets\\Iris\\iris.csv")
iris.head(3)
cols_map = {c: c.replace("_in_cm", "") for c in iris.columns}
iris.head(3)
iris.rename(columns=cols_map, inplace=True)
iris.drop(columns=["Unnamed: 0"], inplace=True)
iris.set_index(pd.Series(range(0, iris.shape[0])), inplace=True)
iris.to_csv("datasets\\Iris\\iris.csv", index=False)
iris = pd.read_csv("datasets\\Iris\\iris.csv")
iris.head(3)
# ## Draw Scatter
# %matplotlib inline
# ### Basic Scatter
fig, ax = plt.subplots()
ax.scatter(iris['sepal_length'], iris["sepal_width"])
ax.set_title("Iris Dataset")
ax.set_xlabel("sepal_length")
ax.set_ylabel("sepal_width")
# ### Color by Class
dfs = {}
colors = {'Iris-setosa':'r', 'Iris-versicolor':'g', 'Iris-virginica':'b'}
for kind in colors.keys():
dfs[kind] = iris.loc[iris["class"] == kind]
fig, ax = plt.subplots()
for kind in colors.keys():
ax.scatter(dfs[kind]['sepal_length'], dfs[kind]["sepal_width"], color=colors[kind])
ax.set_title("Iris Dataset")
ax.set_xlabel("sepal_length")
ax.set_ylabel("sepal_width")
dfs = {}
colors = {'Iris-setosa':'r', 'Iris-versicolor':'g', 'Iris-virginica':'b'}
for kind in colors.keys():
dfs[kind] = iris.loc[iris["class"] == kind]
fig, ax = plt.subplots()
for kind in colors.keys():
ax.scatter(dfs[kind]['petal_length'], dfs[kind]["petal_width"], color=colors[kind])
ax.set_title("Iris Dataset")
ax.set_xlabel("sepal_length")
ax.set_ylabel("sepal_width")
def scatter_plot(v1, v2):
dfs = {}
colors = {'Iris-setosa':'r', 'Iris-versicolor':'g', 'Iris-virginica':'b'}
for kind in colors.keys():
dfs[kind] = iris.loc[iris["class"] == kind]
fig, ax = plt.subplots()
for kind in colors.keys():
ax.scatter(dfs[kind][v1], dfs[kind][v2], color=colors[kind])
ax.set_title("Iris Dataset")
ax.set_xlabel(v1)
ax.set_ylabel(v2)
for v1 in iris.columns[:-1]:
for v2 in iris.columns[:-1]:
if v1 == v2:
continue
else:
scatter_plot(v1, v2)
| numpy_pandas_matplotlib/data_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="3b7edccd"
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import cm
# %matplotlib inline
import pandas as pd
import tensorflow as tf
from tensorflow import keras
#import numpy as np
#from matplotlib import pyplot as plt
#import seaborn as sns
# #%pylab inline
# + [markdown] id="3hK7LxVXUCG4"
# ## Exercise 10
# + id="_bqTFWiCUA5p" colab={"base_uri": "https://localhost:8080/"} outputId="fb026039-066e-44ea-f40c-59c5e7f75f48"
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/"} id="BP_5mS7QUH_U" outputId="1c3857a8-398b-47ae-91a7-68b6801ccd6a"
X_train_full.shape
# + id="m6C5U-1MUJlx"
X_valid, X_train = X_train_full[:5000] / 255., X_train_full[5000:] / 255.
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
X_test = X_test / 255.
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="wUcw0VpeULwj" outputId="f44435a6-43bb-4ddb-8b3a-93b9a47e22ad"
plt.imshow(X_train[0], cmap="binary")
plt.axis('off')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="CwYsCGePUNME" outputId="40007ea2-0ef3-4d29-fc03-f17a7aed2f85"
X_valid.shape
# + colab={"base_uri": "https://localhost:8080/"} id="gskkILTnUPcx" outputId="715c476c-788d-4f1a-fd61-b50c7f52a33a"
X_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="ZB4vg3CLUQuo" outputId="6583fb3e-3a3a-4ffd-e34d-56ce99b34443"
n_rows = 4
n_cols = 10
plt.figure(figsize=(n_cols * 1.2, n_rows * 1.2))
for row in range(n_rows):
for col in range(n_cols):
index = n_cols * row + col
plt.subplot(n_rows, n_cols, index + 1)
plt.imshow(X_train[index], cmap="binary", interpolation="nearest")
plt.axis('off')
plt.title(y_train[index], fontsize=12)
plt.subplots_adjust(wspace=0.2, hspace=0.5)
plt.show()
# + id="AHlbGMODUV2H"
K = keras.backend
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_batch_end(self, batch, logs):
self.rates.append(K.get_value(self.model.optimizer.learning_rate))
self.losses.append(logs["loss"])
K.set_value(self.model.optimizer.learning_rate, self.model.optimizer.learning_rate * self.factor)
# + id="KeqGo_-NUWVj"
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
# + id="o2GGP4zPUakj"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
expon_lr = ExponentialLearningRate(factor=1.005)
# + colab={"base_uri": "https://localhost:8080/"} id="a2QkD8M9UrJ8" outputId="2410cfe7-b84b-4262-ff40-6e40c756ae05"
history = model.fit(X_train, y_train, epochs=1,
validation_data=(X_valid, y_valid),
callbacks=[expon_lr])
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="QEpf9K7CUro-" outputId="172457d5-f3eb-4ff0-b7c7-09b86d1b76d6"
plt.plot(expon_lr.rates, expon_lr.losses)
plt.gca().set_xscale('log')
plt.hlines(min(expon_lr.losses), min(expon_lr.rates), max(expon_lr.rates))
plt.axis([min(expon_lr.rates), max(expon_lr.rates), 0, expon_lr.losses[0]])
plt.grid()
plt.xlabel("Learning rate")
plt.ylabel("Loss")
# + id="5yrtcIUCUtZG"
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
# + id="-sokvAQVU1Hc"
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=3e-1),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="oC2-hohaU1mW" outputId="6ea50ccf-2257-4225-beb7-fb09b4434345"
import os
run_index = 1 # increment this at every run
run_logdir = os.path.join(os.curdir, "my_mnist_logs", "run_{:03d}".format(run_index))
run_logdir
# + colab={"base_uri": "https://localhost:8080/"} id="OzkYZDt3U3PZ" outputId="126224aa-a9d8-4cc2-874a-e3f3f529f8c8"
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
checkpoint_cb = keras.callbacks.ModelCheckpoint("my_mnist_model.h5", save_best_only=True)
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
history = model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[checkpoint_cb, early_stopping_cb, tensorboard_cb])
# + colab={"base_uri": "https://localhost:8080/"} id="sMArfl96U5UL" outputId="7b57d2c3-7cce-4c23-cc7c-3f761803930b"
model = keras.models.load_model("my_mnist_model.h5") # rollback to best model
model.evaluate(X_test, y_test)
# + [markdown] id="a941fa14"
# ## Hyperparameter Tuning
#
# + id="8aPNHxyv9lrP"
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
housing = fetch_california_housing()
X_train_full, X_test, y_train_full, y_test = train_test_split(housing.data, housing.target, random_state=42)
X_train, X_valid, y_train, y_valid = train_test_split(X_train_full, y_train_full, random_state=42)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_valid = scaler.transform(X_valid)
X_test = scaler.transform(X_test)
# + id="96eb8439"
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
# + id="e03921ba"
def build_model(n_hidden=1, n_neurons=30, learning_rate=3e-3, input_shape=[8]):
model = keras.models.Sequential()
model.add(keras.layers.InputLayer(input_shape=input_shape))
for layer in range(n_hidden):
model.add(keras.layers.Dense(n_neurons, activation="relu"))
model.add(keras.layers.Dense(1))
optimizer = keras.optimizers.SGD(learning_rate=learning_rate)
model.compile(loss="mse", optimizer=optimizer)
return model
# + colab={"base_uri": "https://localhost:8080/"} id="27116836" outputId="38a3ed06-1408-4b25-ddb9-6269fbab60b1"
keras_reg = keras.wrappers.scikit_learn.KerasRegressor(build_model)
# + colab={"base_uri": "https://localhost:8080/"} id="9e76d264" outputId="291a6753-5c91-4863-dadb-1ac5a3a9c0bd"
keras_reg.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[keras.callbacks.EarlyStopping(patience=10)])
# + colab={"base_uri": "https://localhost:8080/"} id="50733161" outputId="f570ae4f-9313-420e-b668-684b71fbd249"
mse_test = keras_reg.score(X_test, y_test)
# + id="d40289cb"
np.random.seed(42)
tf.random.set_seed(42)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5ac8fd18" outputId="e431b722-5dcb-46fe-f258-a877a8652ff0"
from scipy.stats import reciprocal
from sklearn.model_selection import RandomizedSearchCV
param_distribs = {
"n_hidden": [0, 1, 2, 3],
"n_neurons": np.arange(1, 100) .tolist(),
"learning_rate": reciprocal(3e-4, 3e-2) .rvs(1000).tolist(),
}
rnd_search_cv = RandomizedSearchCV(keras_reg, param_distribs, n_iter=10, cv=3, verbose=2)
rnd_search_cv.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[keras.callbacks.EarlyStopping(patience=10)])
# + colab={"base_uri": "https://localhost:8080/"} id="4f47faae" outputId="b916ebd6-5786-40e4-cc31-8fdf75429e85"
rnd_search_cv.best_params_
# + colab={"base_uri": "https://localhost:8080/"} id="b023aaf7" outputId="634b4c42-f54a-4d96-d712-8a74b829851b"
rnd_search_cv.best_score_
# + colab={"base_uri": "https://localhost:8080/"} id="347a3ffe" outputId="44593616-3473-44d1-9fc2-ba061c90677e"
rnd_search_cv.best_estimator_
| Lab9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Dog:
def __init__(self, name, breed):
self.name = name
self.breed = breed
# -
# +
class Vehicle:
def __init__(self, num_wheels, color):
self.num_wheels = num_wheels
self.color = color
self.fuel_percent = 0
def add_fuel(self, fuel_amt):
self.fuel_percent += fuel_amt
def add_fuel_outside(vehicle, amt):
vehicle.add_fuel(amt)
# -
class Truck(Vehicle):
def __init__(self, num_wheels, color):
super().__init__(num_wheels, color)
class Node:
def __init__(self, value):
self.value = value
self.next = None
self.prev = None
| learn/04week/code/sandbox/.ipynb_checkpoints/lecture-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 ('tf_pt')
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # 8.
#
# _문제: 선형적으로 분리되는 데이터셋에 `LinearSVC`를 훈련시켜보세요. 그런 다음 같은 데이터셋에 `SVC`와`SGDClassifier`를 적용해보세요. 거의 비슷한 모델이 만들어지는지 확인해보세요._
# Iris 데이터셋을 사용하겠습니다. Iris Setosa와 Iris Versicolor 클래스는 선형적으로 구분이 가능합니다.
# +
from sklearn import datasets
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # 꽃잎 길이, 꽃잎 너비
y = iris["target"]
setosa_or_versicolor = (y == 0) | (y == 1)
X = X[setosa_or_versicolor]
y = y[setosa_or_versicolor]
# +
from sklearn.svm import SVC, LinearSVC
from sklearn.linear_model import SGDClassifier
from sklearn.preprocessing import StandardScaler
C = 5
alpha = 1 / (C * len(X))
lin_clf = LinearSVC(loss="hinge", C=C, random_state=42)
svm_clf = SVC(kernel="linear", C=C)
sgd_clf = SGDClassifier(loss="hinge", learning_rate="constant", eta0=0.001, alpha=alpha,
max_iter=1000, tol=1e-3, random_state=42)
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
lin_clf.fit(X_scaled, y)
svm_clf.fit(X_scaled, y)
sgd_clf.fit(X_scaled, y)
print("LinearSVC: ", lin_clf.intercept_, lin_clf.coef_)
print("SVC: ", svm_clf.intercept_, svm_clf.coef_)
print("SGDClassifier(alpha={:.5f}):".format(sgd_clf.alpha), sgd_clf.intercept_, sgd_clf.coef_)
# -
# 이 3개 모델의 결정 경계를 그려 보겠습니다:
# +
# 각 결정 경계의 기울기와 편향을 계산합니다
w1 = -lin_clf.coef_[0, 0]/lin_clf.coef_[0, 1]
b1 = -lin_clf.intercept_[0]/lin_clf.coef_[0, 1]
w2 = -svm_clf.coef_[0, 0]/svm_clf.coef_[0, 1]
b2 = -svm_clf.intercept_[0]/svm_clf.coef_[0, 1]
w3 = -sgd_clf.coef_[0, 0]/sgd_clf.coef_[0, 1]
b3 = -sgd_clf.intercept_[0]/sgd_clf.coef_[0, 1]
# 결정 경계를 원본 스케일로 변환합니다
line1 = scaler.inverse_transform([[-10, -10 * w1 + b1], [10, 10 * w1 + b1]])
line2 = scaler.inverse_transform([[-10, -10 * w2 + b2], [10, 10 * w2 + b2]])
line3 = scaler.inverse_transform([[-10, -10 * w3 + b3], [10, 10 * w3 + b3]])
# 세 개의 결정 경계를 모두 그립니다
plt.figure(figsize=(11, 4))
plt.plot(line1[:, 0], line1[:, 1], "k:", label="LinearSVC")
plt.plot(line2[:, 0], line2[:, 1], "b--", linewidth=2, label="SVC")
plt.plot(line3[:, 0], line3[:, 1], "r-", label="SGDClassifier")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs") # label="Iris versicolor"
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo") # label="Iris setosa"
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper center", fontsize=14)
plt.axis([0, 5.5, 0, 2])
plt.show()
# -
# 아주 비슷!
# # 9.
#
# _문제: MNIST 데이터셋에 SVM 분류기를 훈련시켜보세요. SVM 분류기는 이진 분류기라서 OvA 전략을 사용해 10개의 숫자를 분류해야 합니다. 처리 속도를 높이기 위해 작은 검증 세트로 하이퍼파라미터를 조정하는 것이 좋습니다. 어느 정도까지 정확도를 높일 수 있나요?_
# 먼저 데이터셋을 로드하고 훈련 세트와 테스트 세트로 나눕니다. `train_test_split()` 함수를 사용할 수 있지만 보통 처음 60,000개의 샘플을 훈련 세트로 사용하고 나머지는 10,000개를 테스트 세트로 사용합니다(이렇게 하면 다른 사람들의 모델과 성능을 비교하기 좋습니다):
# +
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, cache=True)
X = mnist["data"]
y = mnist["target"].astype(np.uint8)
X_train = X[:60000]
y_train = y[:60000]
X_test = X[60000:]
y_test = y[60000:]
# -
# 많은 훈련 알고리즘은 훈련 샘플의 순서에 민감하므로 먼저 이를 섞는 것이 좋은 습관입니다. 하지만 이 데이터셋은 이미 섞여있으므로 이렇게 할 필요가 없습니다.
# 선형 SVM 분류기부터 시작해보죠. 이 모델은 자동으로 OvA(또는 OvR) 전략을 사용하므로 특별히 처리해 줄 것이 없습니다. 간단하네요!
#
# **경고**: 이 작업은 하드웨어에 따라 몇 분이 걸릴 수 있습니다.
lin_clf = LinearSVC(random_state=42)
lin_clf.fit(X_train, y_train)
# 훈련 세트에 대한 예측을 만들어 정확도를 측정해 보겠습니다(최종 모델을 선택해 훈련시킨 것이 아니기 때문에 아직 테스트 세트를 사용해서는 안됩니다):
# +
from sklearn.metrics import accuracy_score
y_pred = lin_clf.predict(X_train)
accuracy_score(y_train, y_pred)
# -
# MNIST에서 83.5% 정확도면 나쁜 성능입니다. 선형 모델이 MNIST 문제에 너무 단순하기 때문이지만 먼저 데이터의 스케일을 조정할 필요가 있습니다:
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float32))
X_test_scaled = scaler.transform(X_test.astype(np.float32))
# **경고**: 이 작업은 하드웨어에 따라 몇 분이 걸릴 수 있습니다.
lin_clf = LinearSVC(random_state=42)
lin_clf.fit(X_train_scaled, y_train)
y_pred = lin_clf.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
# 훨씬 나아졌지만(에러율을 절반으로 줄였습니다) 여전히 MNIST에서 좋은 성능은 아닙니다. SVM을 사용한다면 커널 함수를 사용해야 합니다. RBF 커널(기본값)로 `SVC`를 적용해 보겠습니다.
# **노트**: 향후 버전을 위해 사이킷런 0.22에서 기본값인 `gamma="scale"`을 지정합니다.
svm_clf = SVC(gamma="scale")
svm_clf.fit(X_train_scaled[:10000], y_train[:10000])
y_pred = svm_clf.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
# 아주 좋네요 6배나 적은 데이터에서 모델을 훈련시켰지만 더 좋은 성능을 얻었습니다. 교차 검증을 사용한 랜덤 서치로 하이퍼파라미터 튜닝을 해보겠습니다. 진행을 빠르게 하기 위해 작은 데이터셋으로 작업하겠습니다:
# +
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(svm_clf, param_distributions, n_iter=10, verbose=2, cv=3)
rnd_search_cv.fit(X_train_scaled[:1000], y_train[:1000])
# -
rnd_search_cv.best_estimator_
rnd_search_cv.best_score_
# 이 점수는 낮지만 1,000개의 샘플만 사용한 것을 기억해야 합니다. 전체 데이터셋으로 최선의 모델을 재훈련시켜 보겠습니다:
#
# **경고**: 사용하는 하드웨어에 따라 다음 셀을 실행하는데 몇 시간이 걸릴 수 있습니다.
rnd_search_cv.best_estimator_.fit(X_train_scaled, y_train)
y_pred = rnd_search_cv.best_estimator_.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
# 아주 훌륭하네요! 이 모델을 선택하겠습니다. 이제 테스트 세트로 모델을 테스트합니다:
y_pred = rnd_search_cv.best_estimator_.predict(X_test_scaled)
accuracy_score(y_test, y_pred)
# 아주 나쁘지 않지만 확실히 모델이 다소 과대적합되었습니다.
#
# 하이퍼파라미터를 조금 더 수정할 수 있지만(가령, `C`와/나 `gamma`를 감소시킵니다)
#
# 그렇게 하면 테스트 세트에 과대적합될 위험이 있습니다. 다른 사람들은
#
# 하이퍼파라미터 `C=5`와 `gamma=0.005`에서 더 나은 성능(98% 이상의 정확도)을 얻었습니다.
#
# 훈련 세트를 더 많이 사용해서 더 오래 랜덤 서치를 수행하면 이런 값을 얻을 수 있을지 모릅니다.
# ## 10. _문제: 캘리포니아 주택 가격 데이터셋에 SVM 회귀를 훈련시켜보세요._
# 사이킷런의 `fetch_california_housing()` 함수를 사용해 데이터셋을 로드합니다:
# +
from sklearn.datasets import fetch_california_housing
housing = fetch_california_housing()
X = housing["data"]
y = housing["target"]
# -
# 훈련 세트와 테스트 세트로 나눕니다:
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
# 데이터의 스케일을 조정하는 것을 잊지 마세요:
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
# -
# 먼저 간단한 `LinearSVR`을 훈련시켜 보죠:
# +
from sklearn.svm import LinearSVR
lin_svr = LinearSVR(random_state=42)
lin_svr.fit(X_train_scaled, y_train)
# -
# 훈련 세트에 대한 성능을 확인해 보겠습니다:
# +
from sklearn.metrics import mean_squared_error
y_pred = lin_svr.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
mse
# -
# RMSE를 확인해 보겠습니다:
np.sqrt(mse)
# 훈련 세트에서 타깃은 만달러 단위입니다. RMSE는 기대할 수 있는 에러의 정도를 대략 가늠하게 도와줍니다
#
# (에러가 클수록 큰 폭으로 증가합니다). 이 모델의 에러가 대략 $10,000 정도로 예상할 수 있습니다.
#
# 썩 훌륭하지 않네요. RBF 커널이 더 나을지 확인해 보겠습니다. 하이퍼파라미터 `C`와 `gamma`의 적절한 값을
#
# 찾기 위해 교차 검증을 사용한 랜덤 서치를 적용하겠습니다:
# +
from sklearn.svm import SVR
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(SVR(), param_distributions, n_iter=10, verbose=2, cv=3, random_state=42)
rnd_search_cv.fit(X_train_scaled, y_train)
# -
rnd_search_cv.best_estimator_
# 이제 훈련 세트에서 RMSE를 측정해 보겠습니다:
y_pred = rnd_search_cv.best_estimator_.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
np.sqrt(mse)
# 선형 모델보다 훨씬 나아졌네요. 이 모델을 선택하고 테스트 세트에서 평가해 보겠습니다:
y_pred = rnd_search_cv.best_estimator_.predict(X_test_scaled)
mse = mean_squared_error(y_test, y_pred)
np.sqrt(mse)
| mytest/MachineLearn/5_prac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loop Condition
# ## Imports
import sys
sys.path.append("../../src")
# %reload_ext autoreload
# %autoreload 2
from helpers import graph
from path_invariants import *
from math import sqrt
# The successor graph for $L=pqrpqrpqrpqr=4(pqr)$ is the following:
L="pqr"*4
successor_G_L = successor_graph(L)
graph(successor_G_L, filename="figures/successor_graph_loop.png")
# The directed graph $G=(V, E, W)$ verifies the **loop condition** if the following conditions are true:
#
# 1. G has a f-cut ($C^U$ upper cut) that verifies the path condition, let $w^+=f$ the weight of such cut.
# 1. The upper cut covers all G: $ |V(C^U)| = |V(G)| $
# 1. $\exists w^- \in \mathbb{N} : \forall (u,v) \in $C^U$ \land u \neq v \implies W(v,u)=w^-$
# 1. $r=\sqrt{ w^+ + w^- } \in \mathbb{N} $
# 1. $w^+ > w^-$ (But I conjecture that this condition is implied by having (1), then $\forall u : W(u,u)=w^+$, that introduces an assimetry.)
#
# Condition (1) implies that G has only one component. Condition (2) is needed to avoid false discovery in subgraphs. Condition (3) ensures the symetry in the inversed path Condition (4) verifies the correct combinatory between $(u, v)$ , $(v, u)$ pairs.
# The graph $G^S_L$ has weights $w^+=10$ , $w^-=6$, and $r = \sqrt{10 + 6} = 4 $, then it verifies the loop condition.
loop_condition(successor_G_L)
# The $f$-layers of $L$ for $f=10$ and $f=6$ are:
graph( f_layer( 10, successor_G_L ), "figures/upper-layer.png" )
graph( f_layer( 6, successor_G_L ), "figures/lower-layer.png" )
# The upper-layer of $L$ verifies the path condition and the loop conditions:
path_condition( f_layer( 10, successor_G_L ) )
# But the lower-layer don't
path_condition( f_layer( 6, successor_G_L ) )
# Examples of traces that are not single loops
a_given_graph = successor_graph("abxyxycd")
loop_condition(a_given_graph)
a_given_graph = successor_graph("aabcdabcd")
loop_condition(a_given_graph)
a_given_graph = successor_graph("abcabcxyzxyz")
loop_condition(a_given_graph)
| notebooks/section-2-paths-invariants/Loop condition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import langevin
from scipy.stats import pearsonr,gaussian_kde, chi2
from scipy.optimize import root
SMALL_SIZE = 16
MEDIUM_SIZE = 18
BIGGER_SIZE = 20
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
#SEED = 35010732 # from random.org
#np.random.seed(SEED)
print(plt.style.available)
plt.style.use('seaborn-white')
# +
def correlated_ts(c,delta_t = 0.1,N=1000):
# parameters for coupled oscillator
K,D = 1.0,1.0
data1 = langevin.time_series(A=1/K, D=D, delta_t=delta_t, N=N)
data2 = langevin.time_series(A=1/(K+np.abs(c)), D=D, delta_t=delta_t, N=N)
x1 = (data1 + data2)/2
if c>0:
x2 = (data1 - data2)/2
else:
x2 = (data2-data1)/2
return x1,x2
def c_rho(rho):
return 2*np.abs(rho)/(1-np.abs(rho))*np.sign(rho)
# -
def calc_fundstats(x):
return x[0]**2+x[-1]**2,np.sum(x[1:-1]**2),np.sum(x[0:-1]*x[1:])
# +
def b(D,A,delta_t):
return np.exp(-D/A*delta_t)
def q(aep,ass,ac,b):
return (aep + (1+b**2)*ass - 2*b*ac)/(1-b**2)
def dqdB(aep,ass,ac,b):
return 2*(b*aep+2*b*ass-(1+b**2)*ac)/(1-b**2)**2
def d2qdB2(aep,ass,ac,b):
return (6*b+2)/(1-b**2)**3*(aep+2*ass)-(4*b**3+12*b)/(1-b**2)**3*ac
def dBdA(b,D,A,delta_t):
return b*D*delta_t/A**2
def dBdD(b,A,delta_t):
return -b*delta_t/A
def d2BdA2(b,D,A,delta_t):
return b*D*delta_t/A**3*(D*delta_t/A-2)
def d2BdD2(b,A,delta_t):
return b*delta_t**2/A**2
def d2BdAdD(b,D,A,delta_t):
return b*delta_t/A**2*(1-D*delta_t/A)
def d2qdD2(aep,ass,ac,b,A,delta_t):
return d2qdB2(aep,ass,ac,b)*dBdD(b,A,delta_t)**2+dqdB(aep,ass,ac,b)*d2BdD2(b,A,delta_t)
def d2qdA2(aep,ass,ac,b,D,A,delta_t):
return d2qdB2(aep,ass,ac,b)*dBdA(b,D,A,delta_t)**2+dqdB(aep,ass,ac,b)*d2BdA2(b,D,A,delta_t)
def d2qdAdD(aep,ass,ac,b,D,A,delta_t):
return d2qdB2(aep,ass,ac,b)*dBdA(b,D,A,delta_t)*dBdD(b,A,delta_t)+dqdB(aep,ass,ac,b)*d2BdAdD(b,D,A,delta_t)
#def d2PdA2(N,aep,ass,ac,b,D,A,delta_t):
# return (N/2/A**2 -
# q(aep,ass,ac,b)/A**3 +
# (N-1)/(1-b**2)*(b*d2BdA2(b,D,A,delta_t) + dBdA(b,D,A,delta_t)**2*(1+b**2)/(1-b**2)) -
# d2qdA2(aep,ass,ac,b,D,A,delta_t)/2/A +
# 1/A**2*dqdB(aep,ass,ac,b)*dBdA(b,D,A,delta_t))
def d2PdA2(N,aep,ass,ac,b,D,A,delta_t):
return (-N/2/A**2 +
(N-1)/(1-b**2)*(b*d2BdA2(b,D,A,delta_t) +
dBdA(b,D,A,delta_t)**2*(1+b**2)/(1-b**2) +
2*b/A*dBdA(b,D,A,delta_t)) -
d2qdA2(aep,ass,ac,b,D,A,delta_t)/2/A)
def d2PdAdD(N,aep,ass,ac,b,D,A,delta_t):
return (dqdB(aep,ass,ac,b)*dBdD(b,A,delta_t)/2/A**2 -
d2qdAdD(aep,ass,ac,b,D,A,delta_t)/2/A +
(N-1)/(1-b**2)*(b*d2BdAdD(b,D,A,delta_t) + dBdA(b,D,A,delta_t)*dBdD(b,A,delta_t)*(1+b**2)/(1-b**2)))
def d2PdD2(N,a1ep,a1ss,a1c,a2ep,a2ss,a2c,b1,b2,D,A1,A2,delta_t):
return ((N-1)/(1-b1**2)*(b1*d2BdD2(b1,A1,delta_t) + dBdD(b1,A1,delta_t)**2*(1+b1**2)/(1-b1**2))+
(N-1)/(1-b2**2)*(b2*d2BdD2(b2,A2,delta_t) + dBdD(b2,A2,delta_t)**2*(1+b2**2)/(1-b2**2))-
d2qdD2(a1ep,a1ss,a1c,b1,A1,delta_t)/2/A1 -
d2qdD2(a2ep,a2ss,a2c,b2,A2,delta_t)/2/A2)
def phi_deriv(x,a1ep,a1ss,a1c,a2ep,a2ss,a2c,delta_t,N):
# x[0] = A1, x[1] = A2, x[2]=D
A1 = x[0]
A2 = x[1]
D = x[2]
b1 = b(D,A1,delta_t)
b2 = b(D,A2,delta_t)
Q1 = q(a1ep,a1ss,a1c,b1)
Q2 = q(a2ep,a2ss,a2c,b2)
dQ1 = dqdB(a1ep,a1ss,a1c,b1)
dQ2 = dqdB(a2ep,a2ss,a2c,b2)
y1 = -N*A1**2/2 + A1*Q1/2 + b1*D*delta_t*(A1*b1*(N-1)/(1-b1**2)-dQ1/2)
y2 = -N*A2**2/2 + A2*Q2/2 + b2*D*delta_t*(A2*b2*(N-1)/(1-b2**2)-dQ2/2)
y3 = (b1*(N-1)/(1-b1**2)-dQ1/A1/2)*b1/A1 + (b2*(N-1)/(1-b2**2)-dQ2/A2/2)*b2/A2
return np.array([y1,y2,y3])
def hessian(x,a1ep,a1ss,a1c,a2ep,a2ss,a2c,delta_t,N):
# x[0] = A1, x[1] = A2, x[2]=D
A1 = x[0]
A2 = x[1]
D = x[2]
b1 = b(D,A1,delta_t)
b2 = b(D,A2,delta_t)
d2PdA2_1m = d2PdA2(N,a1ep,a1ss,a1c,b1,D,A1,delta_t)
d2PdA2_2m = d2PdA2(N,a2ep,a2ss,a2c,b2,D,A2,delta_t)
d2PdD2m = d2PdD2(N,a1ep,a1ss,a1c,a2ep,a2ss,a2c,b1,b2,D,A1,A2,delta_t)
d2PdAdD_1m = d2PdAdD(N,a1ep,a1ss,a1c,b1,D,A1,delta_t)
d2PdAdD_2m = d2PdAdD(N,a2ep,a2ss,a2c,b2,D,A2,delta_t)
return np.array([[d2PdA2_1m,0,d2PdAdD_1m],[0,d2PdA2_2m,d2PdAdD_2m],[d2PdAdD_1m,d2PdAdD_2m,d2PdD2m]])
# +
def d2PdA2N(N,b,A,delta_t):
return -N/2/A - N/(1-b**2)**2*dBdA(b,D,A,delta_t)**2*(1+b**2+6*b/(1+b))+2*N*b/(1-b**2)*dBdA(b,D,A,delta_t)
def d2PdAdDN(N,b,D,A,delta_t):
return N*b/(1-b**2)*dBdD(b,A,delta_t) - N/(1-b**2)**2*dBdA(b,D,A,delta_t)*dBdD(b,A,delta_t)*(1+b**2+6*b/(1+b))
def d2PdD2N(N,b1,b2,D,A1,A2,delta_t):
return (-N/(1-b1**2)**2*dBdD(b1,A1,delta_t)**2*(1+b1**2+6*b1/(1+b1))-
N/(1-b2**2)**2*dBdD(b2,A2,delta_t)**2*(1+b2**2+6*b2/(1+b2)))
# -
corr1k = pd.read_csv("correlations1k01.csv")
corr1k
# +
rho = 0.1
delta_t = 0.3
N = 1000
c = 2*rho/(1-rho)
guessa1 = 1.0
guessa2 = 1.0/(1.0+c)
guessd = 1.0
A1_list = []
A2_list = []
dA1_list = []
dA2_list = []
dA1dA2_list = []
dA1dD_list = []
dA2dD_list = []
C_list = []
dC_list = []
D_list = []
dD_list = []
tau1_list = []
dtau1_list = []
tau2_list = []
dtau2_list = []
print(guessa1,guessa2,guessd,c)
for index, row in corr1k.iterrows():
a1ep,a1ss,a1c = row['a1ep'], row['a1ss'], row['a1c']
a2ep,a2ss,a2c = row['a2ep'], row['a2ss'], row['a2c']
para = (a1ep,a1ss,a1c,a2ep,a2ss,a2c,delta_t,N)
result = root(phi_deriv, [guessa1,guessa2,guessd],jac=hessian,args=para)
A1 = result.x[0]
A2 = result.x[1]
D = result.x[2]
# lets calculate the Hessian
h = hessian([A1,A2,D],a1ep,a1ss,a1c,a2ep,a2ss,a2c,delta_t,N)
var = -np.linalg.inv(h)
dA1 = np.sqrt(var[0,0])
dA2 = np.sqrt(var[1,1])
dD = np.sqrt(var[2,2])
dA1A2 = var[0,1]
dA1dD = var[0,2]
dA2dD = var[1,2]
C = (A1-A2)/A2
dC = np.sqrt(1/A2**2*dA1**2+A1**2/A2**4*dA2**2-A1/A2**4*dA1A2)
tau1 = A1/D
tau2 = A2/D
dtau1 = np.sqrt(1/D**2*dA1+A1**2/D**4*dD-A1/D**3*dA1dD)
dtau2 = np.sqrt(1/D**2*dA1+A1**2/D**4*dD-A1/D**3*dA2dD)
print(A1,dA1,A2,dA2,D,dD,C,dC)
# add results to list
A1_list.append(A1)
A2_list.append(A2)
dA1_list.append(dA1)
dA2_list.append(dA2)
D_list.append(D)
dD_list.append(dD)
dA1dA2_list.append(dA1A2)
dA1dD_list.append(dA1dD)
dA2dD_list.append(dA2dD)
C_list.append(C)
dC_list.append(dC)
tau1_list.append(tau1)
tau2_list.append(tau2)
dtau1_list.append(dtau1)
dtau2_list.append(dtau2)
# -
corr1k['A1'] = A1_list
corr1k['A2'] = A2_list
corr1k['dA1'] = dA1_list
corr1k['dA2'] = dA2_list
corr1k['D'] = D_list
corr1k['dD'] = dD_list
corr1k['dA1dA2'] = dA1dA2_list
corr1k['dA1dD'] = dA1dD_list
corr1k['dA2dD'] = dA2dD_list
corr1k['C'] = C_list
corr1k['dC'] = dC_list
corr1k['tau1'] = tau1_list
corr1k['tau2'] = tau2_list
corr1k['dtau1'] = dtau1_list
corr1k['dtau2'] = dtau2_list
corr1k
# display statistics
print(corr1k['A1'].mean(),corr1k['A1'].std(),corr1k['dA1'].mean(),corr1k['dA1'].std())
print(corr1k['a1'].mean(),corr1k['a1'].std(),corr1k['da1'].mean(),corr1k['da1'].std())
print(corr1k['A2'].mean(),corr1k['A2'].std(),corr1k['dA2'].mean(),corr1k['dA2'].std())
print(corr1k['a2'].mean(),corr1k['a2'].std(),corr1k['da2'].mean(),corr1k['da2'].std())
print(corr1k['C'].mean(),corr1k['C'].std(),corr1k['dC'].mean(),corr1k['dC'].std())
print(corr1k['c'].mean(),corr1k['c'].std(),corr1k['dc'].mean(),corr1k['dc'].std())
plt.figure(facecolor="white")
xs = np.linspace(0.65,1.4,200)
densityA1 = gaussian_kde(corr1k['A1'])
densitya1 = gaussian_kde(corr1k['a1'])
plt.plot(xs,densityA1(xs),"k-",label=r"$A_{1}$ ML")
plt.plot(xs,densitya1(xs),"k:",label=r"$A_{1}$ MCMC")
plt.axvline(x=1.0,color="k")
plt.legend()
plt.xlabel(r"$A_1$")
plt.ylabel(r"$p(A_{1})$")
plt.savefig("A1kde01.png",format="png",dpi=300,bbox_inches='tight',facecolor="white",backgroundcolor="white")
plt.figure(facecolor="white")
xs = np.linspace(0.6,1.1,200)
densityA2 = gaussian_kde(corr1k['A2'])
densitya2 = gaussian_kde(corr1k['a2'])
plt.plot(xs,densityA2(xs),"k-",label=r"$A_{2}$ ML")
plt.plot(xs,densitya2(xs),"k:",label=r"$A_{2}$ MCMC")
plt.axvline(x=0.8,color="k")
plt.legend()
plt.xlabel(r"$A_2$")
plt.ylabel(r"$p(A_{2})$")
plt.savefig("A2kde01.png",format="png",dpi=300,bbox_inches='tight',facecolor="white",backgroundcolor="white")
# display statistics
print(corr1k['dA1dA2'].mean(),corr1k['dA1dA2'].std(),corr1k['dA1dD'].mean(),corr1k['dA2dD'].mean())
print(corr1k['da1da2'].mean(),corr1k['dA1dA2'].std(),corr1k['da1dd'].mean(),corr1k['dasdd'].mean())
plt.hist(corr1k['dA1'],bins=30)
plt.hist(corr1k['da1'],bins=30)
plt.hist(corr1k['dA2'],bins=30)
plt.hist(corr1k['da2'],bins=30)
print(corr1k['A1'].std()/corr1k['dA1'].mean())
print(corr1k['A2'].std()/corr1k['dA2'].mean())
print(corr1k['D'].std()/corr1k['dD'].mean())
print(corr1k['dA1dA2'].mean()/corr1k['da1da2'].mean())
print(corr1k['dA1dD'].mean()/corr1k['da1dd'].mean())
print(corr1k['dA2dD'].mean()/corr1k['dasdd'].mean())
plt.hist(corr1k['dA1dA2'],bins=30)
plt.hist(corr1k['da1da2'],bins=30)
plt.hist(corr1k['dA1dD'],bins=30)
plt.hist(corr1k['da1dd'],bins=30)
plt.hist(corr1k['dA2dD'],bins=30)
plt.hist(corr1k['dasdd'],bins=30)
p1 = np.polyfit(corr1k['da1'],corr1k['dA1'],1)
print(p1)
print("factor of underestimation: ",1/p1[0])
da1 = np.linspace(0.052,0.13,200)
dA1 = p1[0]*da1 + p1[1]
plt.figure(facecolor="white")
plt.scatter(corr1k['da1'],corr1k['dA1'],color="k")
plt.plot(da1,dA1,"k:")
plt.xlabel(r"$dA_1$ MCMC")
plt.ylabel(r"$dA_{1}$ ML")
plt.savefig("dA1corrkde01.png",format="png",dpi=300,bbox_inches='tight',facecolor="white",backgroundcolor="white")
p1 = np.polyfit(corr1k['da2'],corr1k['dA2'],1)
print(p1)
print("factor of underestimation: ",1/p1[0])
da2 = np.linspace(0.04,0.1,200)
dA2 = p1[0]*da2 + p1[1]
plt.figure(facecolor="white")
plt.scatter(corr1k['da2'],corr1k['dA2'],color="k")
plt.plot(da2,dA2,"k:")
plt.xlabel(r"$dA_2$ MCMC")
plt.ylabel(r"$dA_{2}$ ML")
#plt.xlim((0.005,0.008))
#plt.ylim((0.004,0.008))
plt.savefig("dA2corrkde01.png",format="png",dpi=300,bbox_inches='tight',facecolor="white",backgroundcolor="white")
corr1k['dA2'].min()
| Correlated OU parameter estimation from simulation rho=0.1 1k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tempo Multi-Model Introduction
#
# 
#
# In this multi-model introduction we will:
#
# * [Describe the project structure](#Project-Structure)
# * [Train some models](#Train-Models)
# * [Create Tempo artifacts](#Create-Tempo-Artifacts)
# * [Run unit tests](#Unit-Tests)
# * [Save python environment for our classifier](#Save-Classifier-Environment)
# * [Test Locally on Docker](#Test-Locally-on-Docker)
# ## Prerequisites
#
# This notebooks needs to be run in the `tempo-examples` conda environment defined below. Create from project root folder:
#
# ```bash
# conda env create --name tempo-examples --file conda/tempo-examples.yaml
# ```
# ## Project Structure
# !tree -P "*.py" -I "__init__.py|__pycache__" -L 2
# ## Train Models
#
# * This section is where as a data scientist you do your work of training models and creating artfacts.
# * For this example we train sklearn and xgboost classification models for the iris dataset.
import os
from tempo.utils import logger
import logging
import numpy as np
logger.setLevel(logging.ERROR)
logging.basicConfig(level=logging.ERROR)
ARTIFACTS_FOLDER = os.getcwd()+"/artifacts"
# + code_folding=[0]
# # %load src/train.py
import joblib
from sklearn.linear_model import LogisticRegression
from src.data import IrisData
from xgboost import XGBClassifier
SKLearnFolder = "sklearn"
XGBoostFolder = "xgboost"
def train_sklearn(data: IrisData, artifacts_folder: str):
logreg = LogisticRegression(C=1e5)
logreg.fit(data.X, data.y)
with open(f"{artifacts_folder}/{SKLearnFolder}/model.joblib", "wb") as f:
joblib.dump(logreg, f)
def train_xgboost(data: IrisData, artifacts_folder: str):
clf = XGBClassifier()
clf.fit(data.X, data.y)
clf.save_model(f"{artifacts_folder}/{XGBoostFolder}/model.bst")
# -
from src.data import IrisData
from src.train import train_sklearn, train_xgboost
data = IrisData()
train_sklearn(data, ARTIFACTS_FOLDER)
train_xgboost(data, ARTIFACTS_FOLDER)
# ## Create Tempo Artifacts
#
# * Here we create the Tempo models and orchestration Pipeline for our final service using our models.
# * For illustration the final service will call the sklearn model and based on the result will decide to return that prediction or call the xgboost model and return that prediction instead.
from src.tempo import get_tempo_artifacts
classifier, sklearn_model, xgboost_model = get_tempo_artifacts(ARTIFACTS_FOLDER)
# + code_folding=[0]
# # %load src/tempo.py
from typing import Tuple
import numpy as np
from src.train import SKLearnFolder, XGBoostFolder
from tempo.serve.metadata import ModelFramework
from tempo.serve.model import Model
from tempo.serve.pipeline import Pipeline, PipelineModels
from tempo.serve.utils import pipeline
PipelineFolder = "classifier"
SKLearnTag = "sklearn prediction"
XGBoostTag = "xgboost prediction"
def get_tempo_artifacts(artifacts_folder: str) -> Tuple[Pipeline, Model, Model]:
sklearn_model = Model(
name="test-iris-sklearn",
platform=ModelFramework.SKLearn,
local_folder=f"{artifacts_folder}/{SKLearnFolder}",
uri="s3://tempo/basic/sklearn",
description="An SKLearn Iris classification model",
)
xgboost_model = Model(
name="test-iris-xgboost",
platform=ModelFramework.XGBoost,
local_folder=f"{artifacts_folder}/{XGBoostFolder}",
uri="s3://tempo/basic/xgboost",
description="An XGBoost Iris classification model",
)
@pipeline(
name="classifier",
uri="s3://tempo/basic/pipeline",
local_folder=f"{artifacts_folder}/{PipelineFolder}",
models=PipelineModels(sklearn=sklearn_model, xgboost=xgboost_model),
description="A pipeline to use either an sklearn or xgboost model for Iris classification",
)
def classifier(payload: np.ndarray) -> Tuple[np.ndarray, str]:
res1 = classifier.models.sklearn(input=payload)
if res1[0] == 1:
return res1, SKLearnTag
else:
return classifier.models.xgboost(input=payload), XGBoostTag
return classifier, sklearn_model, xgboost_model
# -
# ## Unit Tests
#
# * Here we run our unit tests to ensure the orchestration works before running on the actual models.
# + code_folding=[]
# # %load tests/test_tempo.py
import numpy as np
from src.tempo import SKLearnTag, XGBoostTag, get_tempo_artifacts
def test_sklearn_model_used():
classifier, _, _ = get_tempo_artifacts("")
classifier.models.sklearn = lambda input: np.array([[1]])
res, tag = classifier(np.array([[1, 2, 3, 4]]))
assert res[0][0] == 1
assert tag == SKLearnTag
def test_xgboost_model_used():
classifier, _, _ = get_tempo_artifacts("")
classifier.models.sklearn = lambda input: np.array([[0.2]])
classifier.models.xgboost = lambda input: np.array([[0.1]])
res, tag = classifier(np.array([[1, 2, 3, 4]]))
assert res[0][0] == 0.1
assert tag == XGBoostTag
# -
# !python -m pytest tests/
# ## Save Classifier Environment
#
# * In preparation for running our models we save the Python environment needed for the orchestration to run as defined by a `conda.yaml` in our project.
# !cat artifacts/classifier/conda.yaml
from tempo.serve.loader import save
save(classifier)
# ## Test Locally on Docker
#
# * Here we test our models using production images but running locally on Docker. This allows us to ensure the final production deployed model will behave as expected when deployed.
from tempo import deploy_local
remote_model = deploy_local(classifier)
remote_model.predict(np.array([[1, 2, 3, 4]]))
remote_model.undeploy()
# ## Production Option 1 (Deploy to Kubernetes with Tempo)
#
# * Here we illustrate how to run the final models in "production" on Kubernetes by using Tempo to deploy
#
# ### Prerequisites
#
# Create a Kind Kubernetes cluster with Minio and Seldon Core installed using Ansible as described [here](https://tempo.readthedocs.io/en/latest/overview/quickstart.html#kubernetes-cluster-with-seldon-core).
# !kubectl apply -f k8s/rbac -n production
from tempo.examples.minio import create_minio_rclone
import os
create_minio_rclone(os.getcwd()+"/rclone.conf")
from tempo.serve.loader import upload
upload(sklearn_model)
upload(xgboost_model)
upload(classifier)
from tempo.serve.metadata import SeldonCoreOptions
runtime_options = SeldonCoreOptions(**{
"remote_options": {
"namespace": "production",
"authSecretName": "minio-secret"
}
})
from tempo import deploy_remote
remote_model = deploy_remote(classifier, options=runtime_options)
print(remote_model.predict(payload=np.array([[0, 0, 0, 0]])))
print(remote_model.predict(payload=np.array([[1, 2, 3, 4]])))
# ### Illustrate use of Deployed Model by Remote Client
from tempo.seldon.k8s import SeldonKubernetesRuntime
k8s_runtime = SeldonKubernetesRuntime(runtime_options.remote_options)
models = k8s_runtime.list_models(namespace="production")
print("Name\tDescription")
for model in models:
details = model.get_tempo().model_spec.model_details
print(f"{details.name}\t{details.description}")
models[0].predict(payload=np.array([[1, 2, 3, 4]]))
remote_model.undeploy()
# ###### Production Option 2 (Gitops)
#
# * We create yaml to provide to our DevOps team to deploy to a production cluster
# * We add Kustomize patches to modify the base Kubernetes yaml created by Tempo
k8s_runtime = SeldonKubernetesRuntime(runtime_options.remote_options)
yaml_str = k8s_runtime.manifest(classifier)
with open(os.getcwd()+"/k8s/tempo.yaml","w") as f:
f.write(yaml_str)
# !kustomize build k8s
| docs/examples/multi-model/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Load packages to be used:
import dcurves
from dcurves.dca import dca
from dcurves.load_test_data import load_binary_df, load_survival_df
from dcurves.dca import plot_net_benefit_graphs
import numpy as np
import matplotlib.pyplot as plt
import os
import pandas as pd
import pkg_resources
from os import path
# -
root_test_dir = path.dirname(path.realpath("__file__"))
resources_dir = path.join(root_test_dir, 'data')
root_test_dir
resources_dir
os.getcwd()
df_binary = load_binary_df()
df_surv = load_survival_df()
binary_inputs = {
'data': df_binary,
'outcome': 'cancer',
'predictors': ['cancerpredmarker', 'marker'],
'thresh_lo': 0.01,
'thresh_hi': 0.35,
'thresh_step': 0.01,
'harm': None,
'probabilities': [False, True],
'time': None,
'prevalence': None,
'time_to_outcome_col': None
}
| jupyter_notebooks/Untitled2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# +
dfx = pd.read_csv('weightedX.csv')
dfy = pd.read_csv('weightedY.csv')
X = dfx.values
Y = dfy.values
dfx.head(n=5)
dfx.shape
# +
#Normalize the Data
X = (X - X.mean())/X.std()
plt.figure(0)
plt.scatter(X,Y)
plt.title("Normalized Data Plot")
plt.show()
# +
# Hypothesis Same for both linear and locally weighted regression
def getHypothesis(theta,x):
return np.dot(theta,x)
#Function to compute wi
def getWeight(xi,x,bandwidthParam):
return np.exp(((xi-x).T*(xi-x))/(-2*bandwidthParam*bandwidthParam))
#Helper Function to compute W
def getWeightMatrix(query_x,x,bandwidthParam):
no_of_samples = x.shape[0]
#W is an identity matrix of size MXM
W = np.mat(np.eye(no_of_samples))
X = np.mat(x)
for i in range(no_of_samples):
W[i,i] = getWeight(query_x,x[i],bandwidthParam)
return W
#Locally Weighted Linear Regression, Predict for a Query Point
def makePrediction(x,y,query_x,bandwidthParam=0.8):
X = np.mat(x)
ones = np.ones((x.shape[0],1))
X = np.hstack((x,ones))
X = np.mat(X)
#Using the formula : theta =(X′WX)inv * X′WY
X = np.mat(X)
Y = np.mat(y)
W = getWeightMatrix(query_x,x,bandwidthParam)
XT = X.T
YT = Y.T
firstPart = XT*(W*X)
secondPart = (XT*(W*Y))
theta = np.linalg.pinv(firstPart)*secondPart
hx = theta[0][0]*query_x + theta[1][0]
return hx,theta
#Linear Regression Using Normal Equations
def linearRegression(x,y):
X = np.mat(x)
ones = np.ones((x.shape[0],1))
X = np.hstack((x,ones))
X = np.mat(X)
Y = np.mat(y)
XT = X.T
YT = Y.T
firstPart = XT*X
secondPart = XT*Y
theta = np.linalg.pinv(firstPart)*secondPart
return theta
# +
#Test Function
def LinearRegressionTest():
x_test = np.linspace(-2,1.5,20)
y_test = []
theta = linearRegression(X,Y)
#print(theta.shape)
print(theta)
for i in x_test:
y_predicted = theta[0][0]*i + theta[1][0]
y_test.append(y_predicted[0][0])
x_test = np.asarray(x_test)
y_test = np.asarray(y_test)
plt.figure(0)
plt.scatter(X,Y,label="Data")
y_test = y_test.reshape((20,))
plt.plot(x_test,y_test,c='r',label="Linear Regression Hypothesis")
plt.legend()
plt.title("Original vs Predicted Data(Linear Regression)")
plt.show()
LinearRegressionTest()
# +
#Test Function
def LocallyWeightedTest():
x_test = np.linspace(-2,1.5,20)
y_test = []
for i in x_test:
y_predicted,theta = makePrediction(X,Y,i,0.8)
y_test.append(y_predicted[0][0])
x_test = np.asarray(x_test)
y_test = np.asarray(y_test)
plt.figure(0)
plt.scatter(X,Y,label="Data")
y_test = y_test.reshape((20,))
plt.scatter(x_test,y_test,c='r',label="Loess Model")
plt.title("Original vs Predicted Data \n Bandwidth Param : 0.8")
plt.legend()
plt.show()
LocallyWeightedTest()
# +
def testBandwidthParams(minX,maxX,bandwidthParams,testPointsCount):
j=0
for bandwidthParam in bandwidthParams:
x_test = np.linspace(minX,maxX,testPointsCount)
y_test = []
for i in x_test:
y_predicted,theta = makePrediction(X,Y,i,bandwidthParam)
y_test.append(y_predicted)
y_test = np.asarray(y_test)
plt.figure(j+2)
plt.scatter(X,Y,label="Data")
y_test = y_test.reshape((30,))
plt.scatter(x_test,y_test,c='r',label="Model")
plt.legend()
plt.title("Original vs Predicted Data \n Bandwidth Param : "+str(bandwidthParam))
plt.show()
j = j + 1
testBandwidthParams(-2,2,[0.01,0.1,0.3,2,10],30)
# -
| ml_repo/.ipynb_checkpoints/Locally Weighted Linear Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### x lines of Python
#
# # Read and write CSV files
#
# This notebook accompanies the [agilescientific.com](https://agilescientific.com/) blog post of the same name:
#
# > [**x lines of Python: read and write CSV**](https://agilescientific.com/blog/2017/8/23/x-lines-of-python-read-and-write-csv)
#
# ## Introduction
#
# CSV files are the de facto standard way to store data on the web. They are human-readable, easy to parse with multiple tools, and they compress easily. So you need to know how to read and write them in Python.
#
# Nine times out of ten, the way to read and write CSV files with Python is with [`pandas`](http://pandas.pydata.org/). We'll do that first. But that's not always an option (maybe you don't want the dependency in your code), so we'll do it lots of ways:
#
# - With `pandas`
# - 2 ways with the built-in package `csv`...
# - `csv.reader`
# - `csv.DictReader`
# - With `NumPy`.
#
# We'll also use `pandas` to read a couple of CSV files from the web (as opposed to from flat files on our computer).
# ## Using `pandas`
#
# Using [`pd.read_csv`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html).
import pandas as pd
fname = "../data/periods.csv"
df = pd.read_csv(fname)
df
# We can get the start of the Permian like this:
df[df.name=="Permian"].start
# Let's fix the start of the Cretaceous:
df.loc[df.name=='Cretaceous', 'start'] = 145.0
df.loc[df.name=='Cretaceous', 'start']
# After you have changed or added to a DataFrame, `pandas` also makes it very easy to write a CSV file containing your data.
df.to_csv("../data/pdout.csv")
# ## Using `csv.reader`
#
# [Docs for the ordinary reader.](https://docs.python.org/3.5/library/csv.html#csv.reader)
import csv
with open(fname) as f:
reader = csv.reader(f)
data = [row for row in reader]
data
[d[2] for d in data if d[0]=="Permian"]
# Note that we needed to know the positions of the items in the rows, which we could only get by inspection. We could skip that header row if we wanted to, but there's a better way: use the header as the keys in a dictionary...
# ## Using `csv.DictReader`
#
# [Docs for the DictReader.](https://docs.python.org/3.5/library/csv.html#csv.DictReader)
with open(fname) as f:
reader = csv.DictReader(f)
data = [row for row in reader]
data
[d['start'] for d in data if d['name']=="Permian"]
# There is a corresponding [`DictWriter`](https://docs.python.org/3.5/library/csv.html#csv.DictWriter) class for writing CSVs.
# Note that `pandas` has lots of file readers, including ones for:
#
# - Excel files
# - JSON
# - SAS
# - Stata
#
# It can even read the clipboard!
# ## Bonus: reading a CSV file from the web
#
# For example, [this one](https://github.com/seg/2016-ml-contest/blob/master/training_data.csv) is hosted by GitHub. It's publicly readable, so we don't need to authenticate.
#
# In the [X Lines of Python: Machine Learning](04_Machine_learning.ipynb) notebook, I read the online file into a buffer, but it turns out you don't need to do this — you can just give `pd.read_csv()` a URL!
# +
import requests
import io
df = pd.read_csv('https://raw.githubusercontent.com/agile-geoscience/xlines/master/data/periods.csv')
df.head()
# -
# ## Bonus: using NumPy
#
# `pandas` is perfect for this CSV because it's really a table, containing a mixture of data types (strings and floats).
#
# Nonetheless, we can read it as an array... I'm not really into 'named arrays', so I'll just read the two numeric columns.
#
# We'll use [`np.genfromtxt`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.genfromtxt.html).
import numpy as np
x = np.genfromtxt(fname, delimiter=',', skip_header=1, usecols=[2,3])
x
# We can write a CSV like so:
np.savetxt("../data/npout.csv", x, delimiter=",", header="start,end")
# ## Bonus: reading a CSV file from Google Docs
#
# It used to be easy to anonymously read a public file directly from Google Docs, but now you need an API key. It's not too hard to set up, but you'll need to [read some docs](https://developers.google.com/sheets/api/).
#
# When you have an API key, put it here...
key = "PUT YOUR KEY HERE"
import json
# +
url = "https://sheets.googleapis.com/v4/spreadsheets/{id}/values/{sheet}"
meta = {"id": "1YlnEGT8uHpRllk7rjAgFFl8V6B5-kl02DBie11PjG9Q",
"sheet": "Sheet1"
}
url = url.format(**meta)
params = {"key": key}
r = requests.get(url, params=params)
j = json.loads(r.text)['values']
df = pd.DataFrame(j[1:], columns=j[0])
df.head()
# -
| notebooks/06_Read_and_write_CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "4"} tags=["context"]
# ## 1. Regression discontinuity: banking recovery
# <p>After a debt has been legally declared "uncollectable" by a bank, the account is considered "charged-off." But that doesn't mean the bank <strong><em>walks away</em></strong> from the debt. They still want to collect some of the money they are owed. The bank will score the account to assess the expected recovery amount, that is, the expected amount that the bank may be able to receive from the customer in the future. This amount is a function of the probability of the customer paying, the total debt, and other factors that impact the ability and willingness to pay.</p>
# <p>The bank has implemented different recovery strategies at different thresholds (\$1000, \$2000, etc.) where the greater the expected recovery amount, the more effort the bank puts into contacting the customer. For low recovery amounts (Level 0), the bank just adds the customer's contact information to their automatic dialer and emailing system. For higher recovery strategies, the bank incurs more costs as they leverage human resources in more efforts to obtain payments. Each additional level of recovery strategy requires an additional \$50 per customer so that customers in the Recovery Strategy Level 1 cost the company \$50 more than those in Level 0. Customers in Level 2 cost \$50 more than those in Level 1, etc. </p>
# <p><strong>The big question</strong>: does the extra amount that is recovered at the higher strategy level exceed the extra \$50 in costs? In other words, was there a jump (also called a "discontinuity") of more than \$50 in the amount recovered at the higher strategy level? We'll find out in this notebook.</p>
# <p>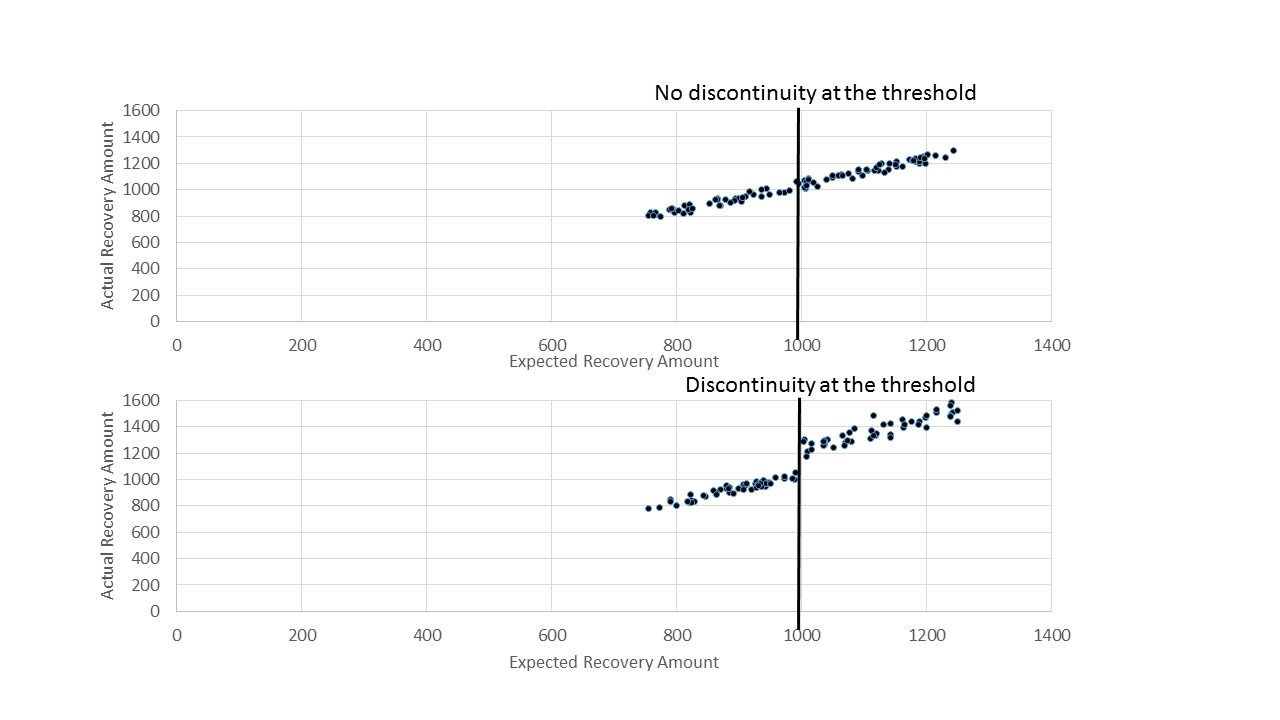</p>
# <p>First, we'll load the banking dataset and look at the first few rows of data. This lets us understand the dataset itself and begin thinking about how to analyze the data.</p>
# + tags=["sample_code"] dc={"key": "4"}
# Import modules
import pandas as pd
import numpy as np
# Read in dataset
df = pd.read_csv("datasets/bank_data.csv")
df.head()
# Print the first few rows of the DataFrame
# ... YOUR CODE FOR TASK 1 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "11"} tags=["context"]
# ## 2. Graphical exploratory data analysis
# <p>The bank has implemented different recovery strategies at different thresholds (\$1000, \$2000, \$3000 and \$5000) where the greater the Expected Recovery Amount, the more effort the bank puts into contacting the customer. Zeroing in on the first transition (between Level 0 and Level 1) means we are focused on the population with Expected Recovery Amounts between \$0 and \$2000 where the transition between Levels occurred at \$1000. We know that the customers in Level 1 (expected recovery amounts between \$1001 and \$2000) received more attention from the bank and, by definition, they had higher Expected Recovery Amounts than the customers in Level 0 (between \$1 and \$1000).</p>
# <p>Here's a quick summary of the Levels and thresholds again:</p>
# <ul>
# <li>Level 0: Expected recovery amounts >\$0 and <=\$1000</li>
# <li>Level 1: Expected recovery amounts >\$1000 and <=\$2000</li>
# <li>The threshold of \$1000 separates Level 0 from Level 1</li>
# </ul>
# <p>A key question is whether there are other factors besides Expected Recovery Amount that also varied systematically across the \$1000 threshold. For example, does the customer age show a jump (discontinuity) at the \$1000 threshold or does that age vary smoothly? We can examine this by first making a scatter plot of the age as a function of Expected Recovery Amount for a small window of Expected Recovery Amount, \$0 to \$2000. This range covers Levels 0 and 1.</p>
# + tags=["sample_code"] dc={"key": "11"}
# Scatter plot of Age vs. Expected Recovery Amount
from matplotlib import pyplot as plt
# %matplotlib inline
plt.scatter(x=df['expected_recovery_amount'], y=df['age'], c="g", s=2)
plt.xlim(0, 2000)
plt.ylim(0, 60)
plt.xlabel("Expected Recovery Amount")
plt.ylabel("Age")
plt.legend(loc=2)
# ... YOUR CODE FOR TASK 2 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "18"} tags=["context"]
# ## 3. Statistical test: age vs. expected recovery amount
# <p>We want to convince ourselves that variables such as age and sex are similar above and below the \$1000 Expected Recovery Amount threshold. This is important because we want to be able to conclude that differences in the actual recovery amount are due to the higher Recovery Strategy and not due to some other difference like age or sex.</p>
# <p>The scatter plot of age versus Expected Recovery Amount did not show an obvious jump around \$1000. We will now do statistical analysis examining the average age of the customers just above and just below the threshold. We can start by exploring the range from \$900 to \$1100.</p>
# <p>For determining if there is a difference in the ages just above and just below the threshold, we will use the Kruskal-Wallis test, a statistical test that makes no distributional assumptions.</p>
# + dc={"key": "18"}
df["expected_recovery_amount"].unique()
# + tags=["sample_code"] dc={"key": "18"}
# Import stats module
from scipy import stats
# Compute average age just below and above the threshold
era_900_1100 = df.loc[(df['expected_recovery_amount']<1100) &
(df['expected_recovery_amount']>=900)]
by_recovery_strategy = era_900_1100.groupby(['recovery_strategy'])
by_recovery_strategy['age'].describe().unstack()
# Perform Kruskal-Wallis test
Level_0_age = era_900_1100.loc[df['recovery_strategy']=="Level 0 Recovery"]['age']
Level_1_age = era_900_1100.loc[df['recovery_strategy']=="Level 1 Recovery"]['age']
stats.kruskal(Level_0_age,Level_1_age)
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "26"} tags=["context"]
# ## 4. Statistical test: sex vs. expected recovery amount
# <p>We have seen that there is no major jump in the average customer age just above and just
# below the \$1000 threshold by doing a statistical test as well as exploring it graphically with a scatter plot. </p>
# <p>We want to also test that the percentage of customers that are male does not jump across the \$1000 threshold. We can start by exploring the range of \$900 to \$1100 and later adjust this range.</p>
# <p>We can examine this question statistically by developing cross-tabs as well as doing chi-square tests of the percentage of customers that are male vs. female.</p>
# + tags=["sample_code"] dc={"key": "26"}
# Number of customers in each category
crosstab = pd.crosstab(df.loc[(df['expected_recovery_amount']<1100) &
(df['expected_recovery_amount']>=900)]['recovery_strategy'],
df['sex'])
# ... YOUR CODE FOR TASK 4 ...
print(crosstab)
# Chi-square test
chi2_stat, p_val, dof, ex = stats.chi2_contingency(crosstab)
print(p_val)
# ... YOUR CODE FOR TASK 4 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "33"} tags=["context"]
# ## 5. Exploratory graphical analysis: recovery amount
# <p>We are now reasonably confident that customers just above and just below the \$1000 threshold are, on average, similar in their average age and the percentage that are male. </p>
# <p>It is now time to focus on the key outcome of interest, the actual recovery amount.</p>
# <p>A first step in examining the relationship between the actual recovery amount and the expected recovery amount is to develop a scatter plot where we want to focus our attention at the range just below and just above the threshold. Specifically, we will develop a scatter plot of Expected Recovery Amount (Y) vs. Actual Recovery Amount (X) for Expected Recovery Amounts between \$900 to \$1100. This range covers Levels 0 and 1. A key question is whether or not we see a discontinuity (jump) around the \$1000 threshold.</p>
# + tags=["sample_code"] dc={"key": "33"}
# Scatter plot of Actual Recovery Amount vs. Expected Recovery Amount
plt.scatter(x=df['expected_recovery_amount'], y=df['actual_recovery_amount'], c="g", s=2)
plt.xlim(900, 1100)
plt.ylim(0, 2000)
plt.xlabel("Expected Recovery Amount")
plt.ylabel( "Actual Recovery Amount")
plt.legend(loc=2)
plt.show()
# ... YOUR CODE FOR TASK 5 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "40"} tags=["context"]
# ## 6. Statistical analysis: recovery amount
# <p>As we did with age, we can perform statistical tests to see if the actual recovery amount has a discontinuity above the \$1000 threshold. We are going to do this for two different windows of the expected recovery amount \$900 to \$1100 and for a narrow range of \$950 to \$1050 to see if our results are consistent.</p>
# <p>Again, we will use the Kruskal-Wallis test.</p>
# <p>We will first compute the average actual recovery amount for those customers just below and just above the threshold using a range from \$900 to \$1100. Then we will perform a Kruskal-Wallis test to see if the actual recovery amounts are different just above and just below the threshold. Once we do that, we will repeat these steps for a smaller window of \$950 to \$1050.</p>
# + tags=["sample_code"] dc={"key": "40"}
# Compute average actual recovery amount just below and above the threshold
by_recovery_strategy['actual_recovery_amount'].describe().unstack()
# Perform Kruskal-Wallis test
Level_0_actual = era_900_1100.loc[df['recovery_strategy']=='Level 0 Recovery']['actual_recovery_amount']
Level_1_actual = era_900_1100.loc[df['recovery_strategy']=='Level 1 Recovery']['actual_recovery_amount']
stats.kruskal(Level_0_actual,Level_1_actual)
# Repeat for a smaller range of $950 to $1050
era_950_1050 = df.loc[(df['expected_recovery_amount']<1050) &
(df['expected_recovery_amount']>=950)]
Level_0_actual = era_950_1050.loc[df['recovery_strategy']=='Level 0 Recovery']['actual_recovery_amount']
Level_1_actual = era_950_1050.loc[df['recovery_strategy']=='Level 1 Recovery']['actual_recovery_amount']
stats.kruskal(Level_0_actual,Level_1_actual)
# ... YOUR CODE FOR TASK 6 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "48"} tags=["context"]
# ## 7. Regression modeling: no threshold
# <p>We now want to take a regression-based approach to estimate the program impact at the \$1000 threshold using data that is just above and below the threshold. </p>
# <p>We will build two models. The first model does not have a threshold while the second will include a threshold.</p>
# <p>The first model predicts the actual recovery amount (dependent variable) as a function of the expected recovery amount (independent variable). We expect that there will be a strong positive relationship between these two variables. </p>
# <p>We will examine the adjusted R-squared to see the percent of variance explained by the model. In this model, we are not representing the threshold but simply seeing how the variable used for assigning the customers (expected recovery amount) relates to the outcome variable (actual recovery amount).</p>
# + tags=["sample_code"] dc={"key": "48"}
# Import statsmodels
import statsmodels.api as sm
# Define X and y
X = era_900_1100['expected_recovery_amount']
y = era_900_1100['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y, X).fit()
predictions = model.predict(X)
model.summary()
# Print out the model summary statistics
# ... YOUR CODE FOR TASK 7 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "55"} tags=["context"]
# ## 8. Regression modeling: adding true threshold
# <p>From the first model, we see that the expected recovery amount's regression coefficient is statistically significant. </p>
# <p>The second model adds an indicator of the true threshold to the model (in this case at \$1000). </p>
# <p>We will create an indicator variable (either a 0 or a 1) that represents whether or not the expected recovery amount was greater than \$1000. When we add the true threshold to the model, the regression coefficient for the true threshold represents the additional amount recovered due to the higher recovery strategy. That is to say, the regression coefficient for the true threshold measures the size of the discontinuity for customers just above and just below the threshold.</p>
# <p>If the higher recovery strategy helped recovery more money, then the regression coefficient of the true threshold will be greater than zero. If the higher recovery strategy did not help recovery more money, then the regression coefficient will not be statistically significant.</p>
# + tags=["sample_code"] dc={"key": "55"}
# Create indicator (0 or 1) for expected recovery amount >= $1000
df['indicator_1000'] = np.where(df['expected_recovery_amount']<1000, 0, 1)
era_900_1100 = df.loc[(df['expected_recovery_amount']<1100) &
(df['expected_recovery_amount']>=900)]
# Define X and y
X = era_900_1100[['expected_recovery_amount','indicator_1000']]
y = era_900_1100['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y,X).fit()
model.summary()
# Print the model summary
# ... YOUR CODE FOR TASK 8 ...
# + run_control={"frozen": true} editable=false deletable=false dc={"key": "62"} tags=["context"]
# ## 9. Regression modeling: adjusting the window
# <p>The regression coefficient for the true threshold was statistically significant with an estimated impact of around \$278. This is much larger than the \$50 per customer needed to run this higher recovery strategy. </p>
# <p>Before showing this to our manager, we want to convince ourselves that this result wasn't due to choosing an expected recovery amount window of \$900 to \$1100. Let's repeat this analysis for the window from \$950 to \$1050 to see if we get similar results.</p>
# <p>The answer? Whether we use a wide (\$900 to \$1100) or narrower window (\$950 to \$1050), the incremental recovery amount at the higher recovery strategy is much greater than the \$50 per customer it costs for the higher recovery strategy. So we conclude that the higher recovery strategy is worth the extra cost of \$50 per customer.</p>
# + tags=["sample_code"] dc={"key": "62"}
# Redefine era_950_1050 so the indicator variable is included
era_950_1050 = df.loc[(df['...']<...) &
(df['...']>=...)]
# Define X and y
X = ...[['expected_recovery_amount','indicator_1000']]
y = ...['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y,X).fit()
# Print the model summary
model.summary()
| Which Debts Are Worth the Bank's Effort_/notebook.ipynb |