code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Q1
my_name= 'molly'
print (my_name)
# # Q2
my_id= 12345
print (my_id)
# # Q3
my_id_str= '12345'
print (my_id_str)
# # Q4
# ##### No, because my_name is a string and my_id is a number.
# # Q5
print (my_name + my_id_str)
# ##### Yes, you can add them because they are both strings.
# # Q6
print ('hello, world. This is my first python string.'.split('.'))
# # Q7
str_list = ['a','b','c','e','d']
str_list.sort()
print (str_list)
# ##### Yes, this list can be sorted into alphabetical order.
# # Q8
str_list.append ('f')
print (str_list)
# # Q9
str_list.remove ('d')
print (str_list)
# # Q10
print (str_list[2])
# # Q11
num_list = [12,32,43,35]
print (str_list + num_list)
# ##### Yes, you can add them together.
# # Q12
my_dict = {'name':'molly',
'id':1234}
print (my_dict)
# # Q13
my_dict['name']= 'bruce'
print (my_dict)
# # Q14
print (my_dict['id'])
# # Q15
my_dict['id']=4321
print (my_dict)
# # Q16
my_dict ['num_list']= num_list
print (my_dict)
# ##### It created a new key.
| Lab2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
# #### Import error
#
# If import fails clone both repos and add them to sys path:
#
# ```
# # !pip install EXCAT-Sync
# ```
from exact_sync.v1.api.annotations_api import AnnotationsApi
from exact_sync.v1.api.images_api import ImagesApi
from exact_sync.v1.api.image_sets_api import ImageSetsApi
from exact_sync.v1.api.annotation_types_api import AnnotationTypesApi
from exact_sync.v1.api.products_api import ProductsApi
from exact_sync.v1.api.teams_api import TeamsApi
from exact_sync.v1.models import ImageSet, Team, Product, AnnotationType, Image, Annotation, AnnotationMediaFile
from exact_sync.v1.rest import ApiException
from exact_sync.v1.configuration import Configuration
from exact_sync.v1.api_client import ApiClient
from pathlib import Path
import numpy as np
from glob import glob
import pickle
import torch
from random import randint
import pyvips
# +
from sklearn.neighbors import KDTree
import cv2
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from fastai import *
from fastai.vision import *
from torchvision import datasets, models, transforms
# -
from fastai.utils.collect_env import show_install
show_install()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
# # Load Data
path = Path('D:/Datasets/AstmaPferd/Patches/')
image_folder = Path('Patches')
tfms = get_transforms(flip_vert=False)
size=64
data = (ImageList.from_folder(path)
.split_by_rand_pct()
.label_from_folder()
.transform(tfms, size=size)).databunch().normalize()
data.show_batch(rows=4)
# # Train Model
learn = cnn_learner(data, models.resnet18, metrics=[accuracy], callback_fns=[])
learn.fit_one_cycle(3,1e-3)
# +
def pil2tensor(image, dtype:np.dtype):
"Convert PIL style `image` array to torch style image tensor."
a = np.asarray(image)
if a.ndim==2 : a = np.expand_dims(a,2)
a = np.transpose(a, (1, 0, 2))
a = np.transpose(a, (2, 1, 0))
return torch.from_numpy(a.astype(dtype, copy=False) )
def open_image(fn:Path, div:bool=True, convert_mode:str='RGB'):
"Return `Image` object created from image in file `fn`."
with warnings.catch_warnings():
warnings.simplefilter("ignore", UserWarning) # EXIF warning from TiffPlugin
x = PIL.Image.open(fn).convert(convert_mode).resize((size,size))
x = pil2tensor(x,np.float32)
if div: x.div_(255)
return x
# -
# ## Last feature layer bevor classifier
#
# TODO: https://github.com/KevinMusgrave/pytorch-metric-learning
trained_model = nn.Sequential(learn.model[0], learn.model[1][:7]).to(device)
learn.model[1][:7]
# ## Extract features for each image
mean, std = to_np(data.stats[0]), to_np(data.stats[1])
paths = []
batch_size = 2
results = []
list(Path('D:/Datasets/AstmaPferd/Patches/').glob("*/*.png"))[:5]
with torch.no_grad():
image_batch, paths_batch = [], []
for path in tqdm(Path('D:/Datasets/AstmaPferd/Patches/').glob("*/*.png")):
image = open_image(str(path))
image = transforms.Normalize(mean, std)(image)
image_batch.append(image[None, :, :, :])
paths_batch.append(path)
if len(image_batch) == batch_size:
features_batch = trained_model(torch.cat(image_batch).to(device))
for path, features in zip(paths_batch, features_batch):
results.append({
'Path': str(path),
'Label': str(path.parent.stem),
'Features': np.array(features_batch.cpu())[0]
})
image_batch, paths_batch = [], []
# ## Scale and PCA / TSNE / UMAP
#
# TODO: https://github.com/lmcinnes/umap
# +
# Scale
scaler = StandardScaler()
features = np.array([result['Features'] for result in results])
scaler.fit(features)
norm_features = scaler.transform(features)
# +
import umap
pca_features = umap.UMAP().fit_transform(norm_features)
# +
# Result image size
size = 64
x_total_size = 10000
y_total_size = 10000
image_size = size
# -
# ### Scale features in that range
# +
min_x = pca_features[:, 0].min()
min_y = pca_features[:, 1].min()
pca_features[:, 0] += abs(min_x)
pca_features[:, 1] += abs(min_y)
max_x = pca_features[:, 0].max()
max_y = pca_features[:, 1].max()
pca_features[:, 0] *= ((x_total_size-(2*image_size)) / max_x)
pca_features[:, 1] *= ((y_total_size-(2*image_size)) / max_y)
# -
new_results = []
for entry, norm_feature, pca_feature in zip(results, norm_features, pca_features):
entry['norm_feature'] = norm_feature
entry['pca_feature'] = pca_feature
new_results.append(entry)
results = new_results
# +
import matplotlib
colors = {}
for i, name in enumerate(data.classes):
colors[name] = np.array(matplotlib.cm.get_cmap('Spectral')(i / data.c))[:3][None]
fig = plt.figure(figsize=(12, 6))
ax = fig.add_subplot(1, 1, 1)
for result in results:
x,y = result['pca_feature']
ax.scatter(x, y, c=colors[result['Label']], label=result['Label'])
ax.set(ylabel="Features Y", xlabel="Features X", title="PCA-Features")
# -
# ## Create Image
target_file = Path('Asthma_{}.tif'.format(str(randint(0, 1000))))
target_file
image_size = size
image_size
center_x, center_y = [], []
for x in range(0, x_total_size - image_size, image_size):
for y in range(0, x_total_size - image_size, image_size):
center_x.append(x)
center_y.append(y)
# +
Centers = np.dstack((center_x, center_y))[0]
big_image = np.zeros(shape=(y_total_size, x_total_size, 3), dtype=np.uint8)
big_image += 255
big_image.shape
# -
from tqdm import tqdm
annotations = []
for result in tqdm(results):
feature = result['pca_feature']
label = result['Label']
path = result['Path']
image = cv2.resize(cv2.imread(str(path))[:, :, [2, 1, 0]], (image_size, image_size))
min_x, min_y = int(feature[0]), int(feature[1])
dists = np.hypot(Centers[:, 0] - min_x, Centers[:, 1] - min_y)
ind = np.argmin(dists)
min_x, min_y = Centers[ind].flatten()
min_x, min_y = int(min_x), int(min_y)
Centers = np.delete(Centers, ind.flatten(), axis=0)
max_x, max_y = min_x + image_size, min_y + image_size
big_image[min_y:max_y, min_x:max_x] = image
vector = json.dumps({"x1": min_x + 3, "y1": min_y + 3, "x2": max_x - 3, "y2": max_y - 3})
#row = "{0}|{1}|{2}|{3}|\n".format(target_file.name, label, vector, Path(path).stem)
row = "{0}|{1}|{2}|\n".format(target_file.name, label, vector)
annotations.append(row)
height, width, bands = big_image.shape
linear = big_image.reshape(width * height * bands)
vi = pyvips.Image.new_from_memory(linear.data, width, height, bands, 'uchar')
vi.tiffsave(str(target_file), tile=True, compression='lzw', bigtiff=True, pyramid=True)
# +
import openslide
import matplotlib.pyplot as plt
level = 1
slide = openslide.open_slide(str(target_file))
slide.level_dimensions
# +
patch = np.array(slide.read_region(location=(0, 0),
level=2, size=slide.level_dimensions[2]))[:, :, :3]
plt.imshow(patch)
# -
# # Upload to EXACT
#
# 0. Create Team
# 1. Create Imageset
# 2. Create Products
# 3. Create Annotation Types
# 4. Upload Image
# 5. Upload Annotations
# +
configuration = Configuration()
configuration.username = 'exact'
configuration.password = '<PASSWORD>'
configuration.host = "http://127.0.0.1:8000"
client = ApiClient(configuration)
image_sets_api = ImageSetsApi(client)
annotations_api = AnnotationsApi(client)
annotation_types_api = AnnotationTypesApi(client)
images_api = ImagesApi(client)
product_api = ProductsApi(client)
team_api = TeamsApi(client)
# -
# ## Create Team if not exists
teams = team_api.list_teams(name="cluster_asthma_team")
if teams.count == 0:
team = Team(name="cluster_asthma_team")
team = team_api.create_team(body=team)
else:
team = teams.results[0]
# ## Create Imageset if not exists
image_sets = image_sets_api.list_image_sets(name="cluster_asthma_imageset")
if image_sets.count == 0:
image_set = ImageSet(name="cluster_asthma_imageset", team=team.id)
image_set = image_sets_api.create_image_set(body=image_set)
else:
image_set = image_sets.results[0]
# ## Create Product if not exists
products = product_api.list_products(name="cluster_product")
if products.count == 0:
product = Product(name="cluster_product", imagesets=[image_set.id], team=team.id)
product = product_api.create_product(body=product)
else:
product = products.results[0]
# ## Create Annotation Types if not exists
data.classes
annotation_types = {}
for y in np.unique(data.classes):
annotation_type_server = annotation_types_api.list_annotation_types(name=y, product=product.id)
if annotation_type_server.count == 0:
vector_type = int(AnnotationType.VECTOR_TYPE.BOUNDING_BOX)
annotation_type = AnnotationType(name=str(y), product=product.id, vector_type=vector_type)
annotation_type = annotation_types_api.create_annotation_type(body=annotation_type)
annotation_types[y] = annotation_type
else:
annotation_types[y] = annotation_type_server.results[0]
# ## Upload Image
image_type = int(Image.ImageSourceTypes.DEFAULT)
image = images_api.create_image(file_path=target_file, image_type=image_type, image_set=image_set.id).results[0]
# ## Upload Annotations
threads = []
for row in tqdm(annotations):
name, label, vector = row.split('|')[:3]
unique_identifier = str(uuid.uuid4())
annotation_type = annotation_types[label]
annotation = Annotation(annotation_type=annotation_type.id, vector=vector, image=image.id, unique_identifier=unique_identifier)
thread = annotations_api.create_annotation(body=annotation, async_req=True)
threads.append(thread)
# ### Wait that all annotations are uploaded
while (len(threads) > 0):
for thread in threads:
if thread.ready():
data.results += thread.get().results
threads.remove(thread)
sleep(0.25)
| doc/ClusterCells.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# name: python37364bitfe49a76c3e654e4faa5ade7a1331a904
# ---
# ## Clustering with K Means
# +
import pandas as pd, seaborn as sns
from matplotlib import pyplot as plt
from sklearn.datasets import load_iris
from sklearn.cluster import KMeans
from sklearn.preprocessing import MinMaxScaler
# -
iris = load_iris()
df = pd.DataFrame(iris.data[:, 2:], columns=['length', 'width'])
df.head()
df.plot.scatter('length', 'width')
# +
scaler = MinMaxScaler()
scaler.fit(df[['length']])
df['length'] = scaler.transform(df[['length']])
scaler.fit(df[['width']])
df['width'] = scaler.transform(df[['width']])
# -
df.head()
df.plot.scatter('length', 'width')
sse = []
k_rng = range(1,10)
for k in k_rng:
km = KMeans(n_clusters=k)
km.fit(df)
sse.append(km.inertia_)
plt.xlabel('K')
plt.ylabel('Sum of squared error')
plt.plot(k_rng,sse)
# +
km_2 = KMeans(n_clusters=2)
df['div_2'] = km_2.fit_predict(df)
km_3 = KMeans(n_clusters=3)
df['div_3'] = km_3.fit_predict(df)
df.head()
# -
sns.scatterplot('length', 'width', hue='div_2', data=df)
sns.scatterplot('length', 'width', hue='div_3', data=df, palette='Set1')
# ## With Pipeline
from sklearn.pipeline import Pipeline
clf = Pipeline([
('scaler', MinMaxScaler()),
('model', KMeans(n_clusters=3)),
])
iris = load_iris()
df = pd.DataFrame(iris.data[:, 2:], columns=['length', 'width'])
df['div_3'] = clf.fit_predict(df)
df.sample(5)
sns.scatterplot('length', 'width', hue='div_3', data=df, palette='Set1')
| 8_k_means_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import pacakage
# +
import torch
import torch.nn as nn
import torch.utils.data as data
import torchvision
from torch.autograd import Variable
import numpy as np
import matplotlib.pyplot as plt
# -
# # Setting
# +
# Settings
epochs = 10
lr = 0.008
# DataLoader
train_set = torchvision.datasets.MNIST(
root='../mnist',
train=True,
download=True,
transform=torchvision.transforms.ToTensor(),
)
test_set = torchvision.datasets.MNIST(
root='../mnist',
train=False,
download=True,
transform=torchvision.transforms.ToTensor(),
)
train_loader = data.DataLoader(train_set, batch_size= 128, shuffle=True)
test_loader = data.DataLoader(test_set, batch_size=8, shuffle=False)
# -
# # Model structure
# AutoEncoder (Encoder + Decoder)
class AutoEncoder(nn.Module):
'''
MNISR image shape = (1,28,28)
784 = 28*28
'''
def __init__(self):
super(AutoEncoder, self).__init__()
# Encoder
self.encoder = nn.Sequential(
nn.Linear(784, 128),
nn.Tanh(),
nn.Linear(128, 64),
nn.Tanh(),
nn.Linear(64, 16),
nn.Tanh(),
nn.Linear(16, 2),
)
# Decoder
self.decoder = nn.Sequential(
nn.Linear(2, 16),
nn.Tanh(),
nn.Linear(16, 64),
nn.Tanh(),
nn.Linear(64, 128),
nn.Tanh(),
nn.Linear(128, 784),
nn.Sigmoid()
)
def forward(self, inputs):
codes = self.encoder(inputs)
decoded = self.decoder(codes)
return codes, decoded
# # Optimizer and loss function
# +
# use gpu if available
# device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# model = AutoEncoder().to(device)
# Optimizer and loss function
model = AutoEncoder()
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
loss_function = nn.MSELoss()
# -
# # Training
for data, labels in train_loader:
print(data.shape,labels.shape)
break
# Train
for epoch in range(epochs):
for data, labels in train_loader:
inputs = data.view(-1, 784)
# Forward
codes, decoded = model(inputs)
# Backward
optimizer.zero_grad()
loss = loss_function(decoded, inputs)
loss.backward()
optimizer.step()
# Show progress
print('[{}/{}] Loss:'.format(epoch+1, epochs), loss.item())
# # Save Model
# Save
torch.save(model, 'models/autoencoder.pth')
# # Import Model
# Load model
model = torch.load('models/autoencoder.pth')
model.eval()
print(model)
# # Plot
# +
# def images_show(images_lst):
# '''Show images
# auto count the size
# '''
# sqrtn = int(np.ceil(np.sqrt(images_lst.shape[0])))
# for index, image in enumerate(images_lst):
# plt.subplot(sqrtn, sqrtn, index+1)
# plt.imshow(image.reshape(28, 28))
# plt.axis('off')
def images_setting(images_lst):
'''Show images
manual size setting
'''
plt.figure(figsize=(20,10))
for index, image in enumerate(images_lst):
plt.subplot(5, 10, index+1)
plt.imshow(image.reshape(28, 28))
plt.axis('off')
plt.show()
# Test
with torch.no_grad():
for data in test_loader:
# plot input image
print("Original image")
inputs = data[0].view(-1, 28*28)
images_setting(inputs)
# plot output image
print("Decode image")
code, outputs = model(inputs)
images_setting(outputs)
break
# -
# # Visualization with Internal Representaion (bottleneck)
# +
plt.figure(figsize=(8,8))
axis_x = []
axis_y = []
answers = []
with torch.no_grad():
for data in test_loader:
inputs = data[0].view(-1, 28*28)
answers += data[1].tolist()
# code is the Internal Representation
# which is redunction to 2 dimension
code, outputs = model(inputs)
axis_x += code[:, 0].tolist()
axis_y += code[:, 1].tolist()
break
plt.title("Numbers's Pattern", fontsize=18, ha="center", va="top")
plt.figtext(0.5,0.05, "Different colors represent different numbers", ha="center", va="bottom", fontsize=14, color="r")
plt.scatter(axis_x, axis_y, c=answers)
plt.colorbar()
plt.show()
# -
# # Feed a random sample to decoder
# +
code = Variable(torch.FloatTensor([[1.19, -3.36]]))
decode = model.decoder(code)
# transform back to (1,1,28,28) and then squeeze to (1,28,28)
decode_img = decode.view(1, 1, 28, 28)
decode_img = decode_img.squeeze()
# transfrom to numpy format
decode_img = decode_img.data.numpy()
# plot show
plt.imshow(decode_img)
plt.show()
# -
# # References
#
# - Implementing an Autoencoder in PyTorch
# https://medium.com/pytorch/implementing-an-autoencoder-in-pytorch-19baa22647d1
# - Kaggle:Autoencoders with PyTorch
# https://www.kaggle.com/jagadeeshkotra/autoencoders-with-pytorch
# - https://www.kaggle.com/ljlbarbosa/convolution-autoencoder-pytorch
# - https://discuss.pytorch.org/t/autoencoders-in-pytorch/844
#
#
# # Read More
# 使用 PyTorch 搭建 GAN 模型產生 MNIST 圖片
# - https://clay-atlas.com/blog/2020/01/09/pytorch-chinese-tutorial-mnist-generator-discriminator-mnist/
| AutoEncoder/AE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cryptolytic-env
# language: python
# name: cryptolytic-env
# ---
# # Cryptolytic Arbitrage Modeling
#
# This notebook contains the code to create the arbitrage models used in the Cryptolytic project. You can find more information on data processing in this [notebook](link) and model evaluation in this [notebook](link).
#
# #### Background on Arbitrage Models
# Arbitrage models were created with the goal of predicting arbitrage 10 min before it happens in an active crypto market. The models are generated by getting all of the combinations of 2 exchanges that support the same trading pair, engineering technical analysis features, merging that data on 'closing_time', engineering more features, and creating a target that signals an arbitrage opportunity. Arbitrage signals predicted by the model have a direction indicating which direction the arbitrage occurs in. A valid arbitrage signal is when the arbitrage lasts >30 mins because it takes time to move coins from one exchange to the other in order to successfully complete the arbitrage trades.
#
# The models predict whether there will be an arbitrage opportunity that starts 10 mins after the prediction time and lasts for at least 30 mins, giving a user enough times to execute trades.
#
# #### Baseline Logistic Regression
#
# #### Baseline Random Forest with default parameters
#
# #### Feature Selection
#
# #### Random Forest with hyperparameter tuning
#
# More than 6000+ iterations of models were generated in this notebook and the best ones were selected from each possible arbitrage combination based on model selection criteria outlined later in this section. The models were Random Forest Classifier and the best model parameters varied for each dataset. The data was obtained from the respective exchanges via their api, and we did a 70/30 train/test split on 5 min candlestick data that fell anywhere in the range from Jun 2015 - Oct 2019. There was a 2 week gap left between the train and test sets to prevent data leakage. The models return 0 (no arbitrage), 1 (arbitrage from exchange 1 to exchange 2) and -1 (arbitrage from exchange 2 to exchange 1).
#
# The profit calculation incorporated fees like in the real world. We used mean percent profit as the profitability metric which represented the average percent profit per arbitrage trade if one were to act on all trades predicted by the model in the testing period, whether those predictions were correct or not.
#
# From the 6000+ iterations of models trained, the best models were narrowed down based on the following criteria:
# - How often the models predicted arbitrage when it didn't exist (False positives)
# - How many times the models predicted arbitrage correctly (True positives)
# - How profitable the model was in the real world over the period of the test set.
#
# There were 21 models that met the thresholds for model selection critera (details of these models can be found at the end of this nb). The final models were all profitable with gains anywhere from 0.2% - 2.3% within the varied testing time periods (Note: the model with >9% mean percent profit was an outlier). Visualizations for how these models performed can be viewed at https://github.com/Lambda-School-Labs/cryptolytic-ds/blob/master/finalized_notebooks/visualization/arb_performance_visualization.ipynb
#
# \* It is HIGHLY recommended to run this on sagemaker and split the training work onto 4 notebooks. These functions will take over a day to run if not split up. There are 95 total options for models, 75 of those options have enough data to train models, and with different options for parameters around ~6K models will be trained. After selecting for the best models, there were 21 that met the criteria to be included in this project.
#
# \*** There has been some feature selection done in this process where we removed highly correlated features, but not enough. There should be more exploration into whether removing features improves accuracy.
#
# \**** We haven't tried normalizing the dataset to see if it will improve accuracy, but that should be a top priority to anyone continuing this project
# #### Directory Structure
# ```
# ├── cryptolytic/ <-- The top-level directory for all arbitrage work
# │ ├── modeling/ <-- Directory for modeling work
# │ │ ├──data/ <-- Directory with subdirectories containing 5 min candle data
# │ │ │ ├─ arb_data/ <-- Directory for csv files of arbitrage model training data
# │ │ │ │ └── *.csv
# │ │ │ │
# │ │ │ ├─ csv_data/ <-- Directory for csv files after combining datasets and FE pt.2
# │ │ │ │ └── *.csv
# │ │ │ │
# │ │ │ ├─ ta_data/ <-- Directory for csv files after FE pt.1
# │ │ │ │ └── *.csv
# │ │ │ │
# │ │ │ ├─ *.zip <-- ZIP files of all of the data
# │ │ │
# │ │ ├──final_models/ <-- Directory for final models after model selection
# │ │ │ └── *.pkl
# │ │ │
# │ │ ├──model_perf/ <-- Directory for performance csvs after training models
# │ │ │ └── *.json
# │ │ │
# │ │ ├──models/ <-- Directory for all pickle models
# │ │ │ └── *.pkl
# │ │ │
# │ │ ├─arbitrage_data_processing.ipynb <-- Notebook for data processing and creating csvs
# │ │ │
# │ │ ├─arbitrage_modeling.ipynb <-- Notebook for baseline models and hyperparam tuning
# │ │ │
# │ │ ├─arbitrage_model_selection.ipynb <-- Notebook for model selection
# │ │ │
# │ │ ├─arbitrage_model_evaluation.ipynb <-- Notebook for final model evaluation
# │ │ │
# │ │ ├─environment.yml <-- yml file to create conda environment
# │ │ │
# │ │ ├─trade_recommender_models.ipynb <-- Notebook for trade recommender models
#
# ```
# ## Imports
#
# This project uses conda to manage environments.
# +
# to update your conda env from a yml file from terminal
# conda env update --file modeling/environment.yml
# to export yml from terminal
# conda env export > modeling/environment.yml
# +
import glob
import os
import pickle
import json
import itertools
from zipfile import ZipFile
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime as dt
from ta import add_all_ta_features
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_recall_fscore_support
from sklearn.metrics import precision_score, recall_score, classification_report, roc_auc_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
# -
# ## Data
#
# Store all the arbitrage datasets that will be used in modeling into a list variable.
arb_data_paths = glob.glob('data/arb_data/*.csv')
print(len(arb_data_paths)) #95
# Each dataset looks like this and should have 141 columns.
df = pd.read_csv(arb_data_paths[0], index_col=0)
print(df.shape)
df.head()
# ## Modeling Functions
# #### Features
#
# Note: closing_time feature is being removed before modeling
features = ['close_exchange_1','base_volume_exchange_1',
'nan_ohlcv_exchange_1','volume_adi_exchange_1', 'volume_obv_exchange_1',
'volume_cmf_exchange_1', 'volume_fi_exchange_1','volume_em_exchange_1',
'volume_vpt_exchange_1','volume_nvi_exchange_1', 'volatility_atr_exchange_1',
'volatility_bbhi_exchange_1','volatility_bbli_exchange_1',
'volatility_kchi_exchange_1', 'volatility_kcli_exchange_1',
'volatility_dchi_exchange_1','volatility_dcli_exchange_1',
'trend_macd_signal_exchange_1', 'trend_macd_diff_exchange_1',
'trend_adx_exchange_1', 'trend_adx_pos_exchange_1',
'trend_adx_neg_exchange_1', 'trend_vortex_ind_pos_exchange_1',
'trend_vortex_ind_neg_exchange_1', 'trend_vortex_diff_exchange_1',
'trend_trix_exchange_1', 'trend_mass_index_exchange_1',
'trend_cci_exchange_1', 'trend_dpo_exchange_1', 'trend_kst_sig_exchange_1',
'trend_kst_diff_exchange_1', 'trend_aroon_up_exchange_1',
'trend_aroon_down_exchange_1', 'trend_aroon_ind_exchange_1',
'momentum_rsi_exchange_1', 'momentum_mfi_exchange_1',
'momentum_tsi_exchange_1', 'momentum_uo_exchange_1',
'momentum_stoch_signal_exchange_1', 'momentum_wr_exchange_1',
'momentum_ao_exchange_1', 'others_dr_exchange_1', 'close_exchange_2',
'base_volume_exchange_2', 'nan_ohlcv_exchange_2',
'volume_adi_exchange_2', 'volume_obv_exchange_2',
'volume_cmf_exchange_2', 'volume_fi_exchange_2',
'volume_em_exchange_2', 'volume_vpt_exchange_2',
'volume_nvi_exchange_2', 'volatility_atr_exchange_2',
'volatility_bbhi_exchange_2', 'volatility_bbli_exchange_2',
'volatility_kchi_exchange_2', 'volatility_kcli_exchange_2',
'volatility_dchi_exchange_2', 'volatility_dcli_exchange_2',
'trend_macd_signal_exchange_2',
'trend_macd_diff_exchange_2', 'trend_adx_exchange_2',
'trend_adx_pos_exchange_2', 'trend_adx_neg_exchange_2',
'trend_vortex_ind_pos_exchange_2',
'trend_vortex_ind_neg_exchange_2',
'trend_vortex_diff_exchange_2', 'trend_trix_exchange_2',
'trend_mass_index_exchange_2', 'trend_cci_exchange_2',
'trend_dpo_exchange_2', 'trend_kst_sig_exchange_2',
'trend_kst_diff_exchange_2', 'trend_aroon_up_exchange_2',
'trend_aroon_down_exchange_2',
'trend_aroon_ind_exchange_2',
'momentum_rsi_exchange_2', 'momentum_mfi_exchange_2',
'momentum_tsi_exchange_2', 'momentum_uo_exchange_2',
'momentum_stoch_signal_exchange_2',
'momentum_wr_exchange_2', 'momentum_ao_exchange_2',
'others_dr_exchange_2', 'year', 'month', 'day',
'higher_closing_price', 'pct_higher',
'arbitrage_opportunity', 'window_length']
# #### Functions for print statements
# +
line = '-------------'
sp = ' '
def tbl_stats_headings():
"""Prints the headings for the stats table"""
print(sp*2, line*9, '\n',
sp*3, 'Accuracy Score',
# sp, 'True Positive Rate',
# sp, 'False Postitive Rate',
sp, 'Precision',
sp, 'Recall',
sp, 'F1', '\n',
sp*2, line*9, '\n',
)
def tbl_stats_row(test_accuracy, precision, recall, f1):
"""Prints the row of model stats after each param set fold"""
print(
sp*4, f'{test_accuracy:.4f}', # accuracy
# sp*3, f'{tpr:.4f}', # roc auc
# sp*3, f'{fpr:.4f}', # p/r auc
sp*2, f'{precision:.4f}', # p/r auc
sp*1, f'{recall:.4f}', # p/r auc
sp*1, f'{f1:.4f}', # p/r auc
sp*2, line*9
)
def print_model_name(name, i, arb_data_paths):
print(
'\n\n', line*9, '\n\n',
f'Model {i+1}/{len(arb_data_paths)}: {name}', '\n',
line*9
)
def print_model_params(i, params, pg_list):
print(
sp*2, line*5, '\n',
sp*2, f'Model {i+1}/{len(pg_list)}', '\n',
sp*2, f'params={params if params else None}', '\n',
sp*2, line*5
)
# -
# #### Functions for calculating profit
# +
# specifying arbitrage window length to target, in minutes
interval = 30
def get_higher_closing_price(df):
"""
Returns the exchange with the higher closing price
"""
# exchange 1 has higher closing price
if (df['close_exchange_1'] - df['close_exchange_2']) > 0:
return 1
# exchange 2 has higher closing price
elif (df['close_exchange_1'] - df['close_exchange_2']) < 0:
return 2
# closing prices are equivalent
else:
return 0
def get_close_shift(df, interval=interval):
"""
Shifts the closing prices by the selected interval +
10 mins.
Returns a df with new features:
- close_exchange_1_shift
- close_exchange_2_shift
"""
rows_to_shift = int(-1*(interval/5))
df['close_exchange_1_shift'] = df['close_exchange_1'].shift(
rows_to_shift - 2)
df['close_exchange_2_shift'] = df['close_exchange_2'].shift(
rows_to_shift - 2)
return df
def get_profit(df):
"""
Calculates the profit of an arbitrage trade.
Returns df with new profit feature.
"""
# if exchange 1 has the higher closing price
if df['higher_closing_price'] == 1:
# return how much money you would make if you bought
# on exchange 2, sold on exchange 1, and took account
# of 0.55% fees
return (((df['close_exchange_1_shift'] /
df['close_exchange_2'])-1)*100)-.55
# if exchange 2 has the higher closing price
elif df['higher_closing_price'] == 2:
# return how much money you would make if you bought
# on exchange 1, sold on exchange 2, and took account
# of 0.55% fees
return (((df['close_exchange_2_shift'] /
df['close_exchange_1'])-1)*100)-.55
# if the closing prices are the same
else:
return 0 # no arbitrage
def profit(X_test, y_preds):
# creating dataframe from test set to calculate profitability
test_with_preds = X_test.copy()
# add column with higher closing price
test_with_preds['higher_closing_price'] = test_with_preds.apply(
get_higher_closing_price, axis=1)
# add column with shifted closing price
test_with_preds = get_close_shift(test_with_preds)
# adding column with predictions
test_with_preds['pred'] = y_preds
# adding column with profitability of predictions
test_with_preds['pct_profit'] = test_with_preds.apply(
get_profit, axis=1).shift(-2)
# filtering out rows where no arbitrage is predicted
test_with_preds = test_with_preds[test_with_preds['pred'] != 0]
# calculating mean profit where arbitrage predicted...
pct_profit_mean = round(test_with_preds['pct_profit'].mean(), 2)
# calculating median profit where arbitrage predicted...
pct_profit_median = round(test_with_preds['pct_profit'].median(), 2)
return pct_profit_mean, pct_profit_median
# -
# #### Function for modeling
# +
def create_models(arb_data_paths, model_type, features, param_grid):
"""
This function takes in a list of all the arbitrage data paths,
does train/test split, feature selection, trains models,
saves the pickle file, and prints performance stats for each model
Predictions
___________
Models predict whether arbitrage will in 10 mins from the
prediction time, and last for at least 30 mins:
1: arbitrage from exchange 1 to exchange 2
0: no arbitrage
-1: arbitrage from exchange 2 to exchange 1
Evaluation
__________
- Accuracy Score
- Precision
- Recall
- F1 score
- Mean Percent Profit
- Median Percent Profit
Parameters
__________
arb_data_paths: filepaths for all the datasets used in modeling
model_type: scikit-learn model (LogisticRegression() or
RandomForestClassifier())
features: the features for training or empty [] for all features
param_grid: the params used for hyperparameter tuning or empty {}
"""
base_model_name = str(model_type).split('(')[0]
model_name_dict = {
'LogisticRegression':'lr',
'RandomForestClassifier':'rf'
}
# this is part of a check put in the code to allow the function
# to pick up where it previously left off in case of errors
model_paths = glob.glob('models2/*.pkl')
# pick target
target = 'target'
# iterate through the arbitrage csvs
for i, file in enumerate(arb_data_paths):
# define model name
name = file.split('/')[2][:-8]
# print status
print_model_name(name, i, arb_data_paths)
# read csv
df = pd.read_csv(file, index_col=0)
# convert str closing_time to datetime
df['closing_time'] = pd.to_datetime(df['closing_time'])
# this makes the function dynamic for whether you want
# to select features or hyperparameters or not
# selected features and hyperparameters
if features and param_grid:
pg_list = list(ParameterGrid(param_grid))
# if theres features and no parms
elif features and not param_grid:
pg_list = [param_grid]
# if theres params and no features
elif not features and param_grid:
pg_list = [param_grid]
# baseline: no features and no params
else:
features = df.drop(
labels=['target', 'closing_time'],
axis=1
).columns.to_list()
pg_list = [param_grid]
# hyperparameter tuning
for i, params in enumerate(pg_list):
# define model name
if param_grid:
model_name = '_'.join([
name,
str(max_features),
str(max_depth),
str(n_estimators)
])
else:
model_name = name + '_' + model_name_dict[base_model_name]
# define model filename to check if it exists
model_path = f'models/{model_name}.pkl'
# if the model does not exist
if model_path not in model_paths:
# print status
print_model_params(i, params, pg_list)
# remove 2 weeks from train datasets to create a
# two week gap between the data - prevents data leakage
tt_split_row = round(len(df)*.7)
tt_split_time = df['closing_time'][tt_split_row]
cutoff_time = tt_split_time - dt.timedelta(days=14)
# train and test subsets
train = df[df['closing_time'] < cutoff_time]
test = df[df['closing_time'] > tt_split_time]
# X, y matrix
X_train = train[features]
X_test = test[features]
y_train = train[target]
y_test = test[target]
# printing shapes to track progress
print(sp*2, 'train and test shape: ', train.shape, test.shape)
# filter out datasets that are too small
if ((X_train.shape[0] > 1000)
and (X_test.shape[0] > 100)
and len(set(y_train)) > 1):
# instantiate model
model = model_type.set_params(**params)
# there was a weird error caused by two of the datasets which
# is why this try/except is needed to keep the function running
# try:
# fit model
model = model.fit(X_train, y_train)
# make predictions
y_preds = model.predict(X_test)
pct_prof_mean, pct_prof_median = profit(X_test, y_preds)
print(sp*2,'percent profit mean:', pct_prof_mean)
print(sp*2, 'percent profit median:', pct_prof_median, '\n\n')
# classification report
print(classification_report(y_test, y_preds))
# save model
pickle.dump(model, open(f'models/{model_name}.pkl', 'wb'))
# except:
# print(line*3 + '\n' + line + 'ERROR' + line + '\n' + line*3)
# break # break out of for loop if there is an error with modeling
# dataset is too small
else:
print(f'{sp*2}ERROR: dataset too small for {name}')
# the model exists
else:
print(f'{sp*2}{model_path} already exists.')
# -
create_models(arb_data_paths=arb_data_paths, model_type=LogisticRegression(), features=features, param_grid={})
# ## Class Distribution
#
# +
def class_distribution(arb_data_paths):
"""
Returns the class distribution for all arbitrage
datasets in a df
"""
dist_df = pd.DataFrame(columns=[
'arbitrage_combination',
'ex1_to_ex2_arb',
'ex2_to_ex1_arb',
'no_arb'
])
for path in arb_data_paths:
df = pd.read_csv(path, index_col=0)
arbitrage_combination = path.split('/')[2][:-4]
no_arb = round(df.target.value_counts()[0] / df.target.value_counts().sum(), 2)
ex1_to_ex2_arb = round(df.target.value_counts()[1] / df.target.value_counts().sum(), 2)
ex2_to_ex1_arb = round(df.target.value_counts()[-1] / df.target.value_counts().sum(), 2)
dist_dict = {
'arbitrage_combination': arbitrage_combination,
'ex1_to_ex2_arb': ex1_to_ex2_arb,
'ex2_to_ex1_arb': ex2_to_ex1_arb,
'no_arb': no_arb
}
dist_df = dist_df.append(dist_dict, ignore_index=True)
return dist_df
dist_df = class_distribution(arb_data_paths)
dist_df
# -
# ## Baseline
# #### Logistic Regression Models
create_models(arb_data_paths=arb_data_paths, model_type=LogisticRegression(), features=features, param_grid={})
# #### Random Forest Models w/ default parameters
create_models(arb_data_paths=arb_data_paths, model_type=LogisticRegression(), features=features, param_grid={})
# sns.pairplot(df)
# #### Feature Importances
create_models(arb_data_paths=arb_data_paths, model_type=LogisticRegression(), features=features, param_grid={})
# +
importances = pd.Series(xg.feature_importances_, X_train.columns)
n = 25
plt.figure(figsize=(10,n/2))
plt.title(f'Top {n} features')
importances.sort_values()[-n:].plot.barh(color='blue');
# -
# ## Hyperparameter Tuning
# +
param_grid = {
'max_features': ['auto', 42, 44, 46],
'n_estimators': [250, 300],
'max_depth': [30, 35, 45, 50]
}
create_models(
arb_data_paths=arb_data_paths,
model_type=LogisticRegression(),
features=features, param_grid
)
# +
def preformance_metric():
############## Performance metrics ###############
# TODO: put this all in a function and just return the
# metrics we want
performance_list = []
confusion_dict = {}
# labels for confusion matrix
unique_y_test = y_test.unique().tolist()
unique_y_preds = list(set(y_preds))
labels = list(set(unique_y_test + unique_y_preds))
labels.sort()
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
# create confusion matrix
confusion = pd.DataFrame(confusion_matrix(y_test, y_preds),
columns=columns, index=index)
print(model_name + ' confusion matrix:')
print(confusion, '\n')
# append to confusion list
confusion_dict[model_name] = confusion
# creating dataframe from test set to calculate profitability
test_with_preds = X_test.copy()
# add column with higher closing price
test_with_preds['higher_closing_price'] = test_with_preds.apply(
get_higher_closing_price, axis=1)
# add column with shifted closing price
test_with_preds = get_close_shift(test_with_preds)
# adding column with predictions
test_with_preds['pred'] = y_preds
# adding column with profitability of predictions
test_with_preds['pct_profit'] = test_with_preds.apply(
get_profit, axis=1).shift(-2)
# filtering out rows where no arbitrage is predicted
test_with_preds = test_with_preds[test_with_preds['pred'] != 0]
# calculating mean profit where arbitrage predicted...
pct_profit_mean = test_with_preds['pct_profit'].mean()
# calculating median profit where arbitrage predicted...
pct_profit_median = test_with_preds['pct_profit'].median()
print('percent profit mean:', pct_profit_mean)
print('percent profit median:', pct_profit_median, '\n\n')
# save net performance to list
performance_list.append([name, max_features, max_depth, n_estimators,
pct_profit_mean, pct_profit_median])
######################## END OF TODO ###########################
# -
| modeling/arbitrage_modeling2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VacationPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from api_keys import g_key
import json
# -
import json
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# +
# File to Load (Remember to Change These)
weather = "output_data/WeatherData.csv"
# Read School and Student Data File and store into Pandas DataFrames
df_weather = pd.read_csv(weather)
# -
df_weather
df_weather = df_weather.rename(columns={'Latitude_x': 'Latitude'})
df_weather = df_weather.drop(columns=['Latitude_y'])
df_weather.head()
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# Configure gmaps with API key
gmaps.configure(api_key=g_key)
# +
# Store 'Lat' and 'Lng' into locations
locations = df_weather[["Latitude", "Long"]].astype(float)
# Convert Poverty Rate to float and store
# HINT: be sure to handle NaN values
df_weather = df_weather.dropna()
humi = df_weather["Humidity"].astype(float)
# +
# Create a poverty Heatmap layer
fig = gmaps.figure()
heat_layer = gmaps.heatmap_layer(locations, weights=humi,
dissipating=False, max_intensity=100,
point_radius = 1)
fig.add_layer(heat_layer)
fig
# -
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
df_weather
df_weathercriteria = df_weather.loc[(df_weather['Cloudiness']==0) &
(df_weather['WindSpeed'] < 10.0)&
(df_weather['Temp'] <80.0) &
(df_weather['Temp'] > 70.0) ]
df_weathercriteria
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
narrowed_city_df = df_weathercriteria
narrowed_city_df["Hotel Name"] = ""
narrowed_city_df
# +
#TEST
# params dictionary to update each iteration
#params2 = {
# "radius": 5000,
# "types": "lodging",
# "location": "31.05,76.12",
# "key": g_key
#}
#base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
#testrequest = requests.get(base_url, params= params2)
#print(testrequest.json())
#print(json.dumps(testrequest.json(),indent=2))
#testrequest.json()["results"][0]["name"]
#hotelname = testrequest.json()["results"][0]["name"]
# +
# params dictionary to update each iteration
params = {
"radius": 5000,
"types": "lodging",
"location": "location",
"key": g_key
}
# Use the lat/lng we recovered to identify airports
for index, row in narrowed_city_df.iterrows():
lat = row["Latitude"]
lng = row["Long"]
# change location each iteration while leaving original params in place
params["location"] = f"{lat},{lng}" #<=> "31.05,76.12"
# Use the search term: "International Airport" and our lat/lng
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
# make request and print url
name_address = requests.get(base_url, params=params)
#print the name_address url, avoid doing for public github repos in order to avoid exposing key
#print(name_address.url)
# convert to json
name_address = name_address.json()
# print(json.dumps(name_address, indent=4, sort_keys=True))
# Since some data may be missing we incorporate a try-except to skip any that are missing a data point.
try:
narrowed_city_df.loc[index, "Hotel Name"] = name_address["results"][0]["name"]
except (KeyError, IndexError):
print("Missing field/result... skipping.")
# -
narrowed_city_df
# +
# Store 'Lat' and 'Lng' into locations
locations = narrowed_city_df[["Latitude", "Long"]].astype(float)
# Convert Poverty Rate to float and store
# HINT: be sure to handle NaN values
#df_weather = df_weather.dropna()
humi2 = narrowed_city_df["Humidity"].astype(float)
# +
# Create a poverty Heatmap layer
fig = gmaps.figure()
heat_layer = gmaps.heatmap_layer(locations, weights=humi2,
dissipating=False, max_intensity=100,
point_radius = 1)
fig.add_layer(heat_layer)
fig
# -
narrowed_city_df
hotel_df = narrowed_city_df[["Hotel Name","CityName", "Country","Latitude","Long"]]
#hotel_df.dtypes
# +
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{CityName}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in narrowed_city_df.iterrows()]
locations = hotel_df[["Latitude", "Long"]]
# +
# Add marker layer ontop of heat map
marker_layer = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig = gmaps.figure()
fig.add_layer(marker_layer)
#fig
fig.add_layer(heat_layer)
# Display Map
fig
# -
| WeatherPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sqlite3 as sql
import matplotlib as mp
# Create database connection
db = "/Users/bing/CodeLouisville/Python_DB_notebooks/Titanic.db"
# db = 'titanic.db'
conn = sql.connect(db)
valid_class = {1,2,3,'ALL'}
valid_gender = list("M","F", "BOTH")
valid_age = list(range(0, 100))
valid_survived = list(0,1,"NONE")
# +
print("Build query to display Titanic passenger data")
print()
print("Parameters allowed are:")
print(" ")
print(" Class: [1, 2, 3, or ALL] Example: 1,2 or 1,3 or ALL or just hit the 'Enter' key for ALL ")
print(" Gender: [M, F, or BOTH] Example: M, BOTH, or just hit the 'Enter' key for BOTH")
print(" Age: [Enter an age, a string of ages separated by commas, or ALL] or just hit the 'Enter' key for ALL")
print("Survived: [Y, N, or BOTH] Example: Y BOTH or just hit the 'Enter' key for BOTH")
print("")
class_in_string = " Enter Class(s), or press 'Enter' for all Classes: "
gender_in_string = " Enter Gender, or press 'Enter' for both Genders: "
age_in_string = "Enter list of ages separated by a comma, or press 'Enter' for all Ages: "
survived_in_string = "Enter whether you want data from passengers that survived[Y] or not[N] or press 'Enter' for both"
in_class = input(class_in_string)
in_class = upper(in_class)
if in_class is null:
in_class = 'ALL'
in_gender = input(gender_in_string)
if in_gender is null:
in_gender = 'BOTH'
in_age = input(age_in_string)
if upper(in_age) is null:
in_age = 'ALL'
elsif:
upper(in_age
in_survived = input(survived_in_string)
if in_survived is null:
in_survived = 'ALL'
# -
# Build SQL based on the input given
query = '''
SELECT pl_class.*
FROM passenger_list pl_class,
passenger_list pl_gender,
passenger_list pl_age,
passenger_list pl_survived
WHERE pl_class.id = pl_gender.Id
AND pl_class.id = pl_age.Id
AND pl_class.Id = pl_survived.id
AND pl_class.class =
AND pl_gender.gender = 'M'
AND pl_age.age BETWEEN 20 AND 40
AND NOT pl_survived.Survived;'''
| Old/Python_DB_notebooks/Old/input_params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Sending programs over the wire
#
# Logic can be sent remotely to the server
from prologterms import TermGenerator, PrologRenderer, Program, Var, Term
from sparqlprog import SPARQLProg
from rdflib import Namespace
P = TermGenerator()
R = PrologRenderer()
dbont = Namespace("http://dbpedia.org/ontology/")
dbr = Namespace("http://dbpedia.org/resource/")
server = 'http://localhost:9083'
# ## Creating a program
# +
X = Var('X')
Y = Var('Y')
Z = Var('Z')
rules = [
# in-band if bandMember OR formerBandMember
P.in_band(X,Y) <= P.rdf(Y, dbont['bandMember'], X),
P.in_band(X,Y) <= P.rdf(Y, dbont['formerBandMember'], X),
# shared band members, at any point in time
P.has_shared_band_member(X,Y,Z) <= (P.in_band(Z, X), P.in_band(Z,Y), Term('\=', X, Y))
]
# -
S = SPARQLProg(server=server,
rules=rules,
endpoint='dbpedia')
# ### setting up a query
query = P.has_shared_band_member(dbr['Deep_Purple'], X, Z)
#query = P.has_shared_band_member(Y, X, Z)
#query = P.in_band(X,Y)
# ### running the querry
#
res = S.query(query)
for r in res:
print(f"band: {r['X']} connecting member: {r['Z']}")
| Notebook_02_Programs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install --upgrade pip
# !pip install descartes
# !pip install geopandas
# !pip install CString
# !pip install jimutmap
import csv
import json
import sys
import requests
import os
import urllib
from scipy import ndimage
from scipy import misc
import pandas as pd
import numpy as np
import time
import collections
import hashlib
import hmac
import base64
import urllib.parse
import geopandas as gpd
import matplotlib.pyplot as plt
from shapely.geometry import Polygon
# from jimutmap import api
gpd.io.file.fiona.drvsupport.supported_drivers['KML'] = 'rw'
# +
fp = "KML/blocks.kml"
polys = gpd.read_file(fp, driver='KML')
polys
# +
secret = '<KEY>'
key = '<KEY>'
def generateURL(lat1, long1):
url = 'https://maps.googleapis.com/maps/api/staticmap?center=' + str(lat1) + ',' + \
str(long1) + '&zoom=18&scale=2&size=512x512&maptype=satellite&key=' + key
return url
# -
def save_img(url, file_name):
a = urllib.request.urlopen(url).read()
urllib.request.urlretrieve(url, file_name)
return True
polys['geometry'][0].exterior.coords[0]
i = 1
for c in polys.centroid:
url = generateURL(c.y, c.x)
print(url)
save_img(url, 'blocks/block'+str(i)+'.jpg')
break
i += 1
fig = plt.figure()
ax = polys.plot()
cx.add_basemap(ax, crs=block1.crs, source=cx.providers.OpenStreetMap.Mapnik)
plt.show()
| KML_ploygons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://github.com/Harvard-IACS/2018-CS109A/blob/master/content/styles/iacs.png?raw=true"> CS-S109A Introduction to Data Science
#
# ## Lecture 2 (Pandas + Beautiful Soup)
#
# **Harvard University**<br>
# **Summer 2020**<br>
# **Instructors:** <NAME><br>
# **Authors:** <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# ---
## RUN THIS CELL TO GET THE RIGHT FORMATTING
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# # Table of Contents
# <ol start="0">
# <li> Learning Goals </li>
# <li> Data without Pandas </li>
# <li> Loading and Cleaning with Pandas </li>
# <li> Combinng Data Sources </li>
# <li> Basic Scraping with Beautiful Soup </li>
# </ol>
# ## Learning Goals
#
# This Jupyter notebook accompanies Lecture 2. By the end of this lecture, you should be able to:
#
# - Appreciate that base Python is not great for most data handling.
# - Understand why and how Pandas can be useful.
# - Use Pandas to:
# - Load data into a DataFrame
# - Access subsets of data based on column and row values
# - Address missing values (e.g., `NaN`)
# - Use `groupby()` to select sections of data.
# - Plot DataFrames (e.g., barplot())
# - Use Beautiful Soup to download a webpage and all of its links, and begin to learn parse out tables, links, etc.
# ## Part 1: Processing Data without Pandas
#
# `../data/top50.csv` is a dataset found online (Kaggle.com) that contains information about the 50 most popular songs on Spotify in 2019.
#
# Each row represents a distinct song.
# The columns (in order) are:
# ```
# ID: a unique ID (i.e., 1-50)
# TrackName: Name of the Track
# ArtistName: Name of the Artist
# Genre: the genre of the track
# BeatsPerMinute: The tempo of the song.
# Energy: The energy of a song - the higher the value, the more energetic. song
# Danceability: The higher the value, the easier it is to dance to this song.
# Loudness: The higher the value, the louder the song.
# Liveness: The higher the value, the more likely the song is a live recording.
# Valence: The higher the value, the more positive mood for the song.
# Length: The duration of the song (in seconds).
# Acousticness: The higher the value, the more acoustic the song is.
# Speechiness: The higher the value, the more spoken words the song contains.
# Popularity: The higher the value, the more popular the song is.
# ```
from PIL import Image
Image.open("fig/top50_screenshot.png") # sample of the data
# ### Read and store `../data/top50.csv`
#
# **Q1.1:** Read in the `../data/top50.csv` file and store all of its contents into any data structure(s) that make the most sense to you, keeping in mind that you'd want to easily access any row or column. Why does a dictionary make the most sense to use for data storage?
# +
f = open("../data/top50.csv")
column_names = f.readline().strip().split(",")[1:] # puts names in a list
cleaned_column_names = [name for name in column_names] # removes the extraneous quotes
cleaned_column_names.insert(0, "ID")
dataset = []
# iterates through each line of the .csv file
for line in f:
attributes = line.strip().split(",")
# constructs a new dictionary for each line, and
# appends this dictionary to the `dataset`;
# thus, the dataset is a list of dictionaries (1 dictionary per song)
dataset.append(dict(zip(cleaned_column_names, attributes)))
# dataset[0:2]
# -
# **Q1.2:** Write code to print all songs (Artist and Track name) that are longer than 4 minutes (240 seconds):
# +
########
# your code below: uncomment and fill in the ****
########
#for song in ****:
#if int(song[****] > ****) :
# print(****, "-", ****, "is", **** ,"seconds long")
# -
# **Q1.3:** Write code to print the most popular song (or song(s) if there is a tie):
# +
########
# your code below: uncomment and fill in the ****
########
max_score = -1
most_populars = set()
# for ***:
# if int(song["Popularity"]) > max_score:
# most_populars = set([str(song["ArtistName"] + "-" + song["TrackName"])])
# max_score = int(song["Popularity"])
# elif ****:
# most_populars.add(****)
# print(most_populars)
# -
# **Q1.4:** How would you print the songs (and their attributes) in sorted order by their popularity (highest scoring ones first)?
# *your answer here*
# **Q1.5**: How could you check for null/empty entries?
# *your answer here*
# Often times, one dataset doesn't contain all of the information you are interested in -- in which case, you need to combine data from multiple files.
#
# **Q1.6:** Imagine we had another table (i.e., .csv file) below. How could we combine its data with our already-existing *dataset*?
# *your answer here*
# ## Part 2: Processing Data _with_ Pandas
# **Pandas** is an _open-source_ Python library designed for **data analysis and processing.** Being _open-sourced_ means that anyone can contribute to it (don't worry, a team of people vett all official updates to the library). Pandas allows for high-performance, easy-to-use data structures. Namely, instead of using N-dimensional arrays like NumPy (which are extremely fast, though), Pandas provides a 2D-table object calleda **DataFrame**.
#
# As a very gross simplification: **NumPy** is great for performing math operations with matrices, whereas **Pandas** is excellent for wrangling, processing, and understanding 2D data like spreadsheets (2D data like spreadsheets is very common and great).
#
# Let's get started with simple examples of how to use Pandas. We will continue with our ``top50.csv`` Spotify music data.
#
# First, we need to import pandas so that we have access to it. For typing convenience, we choose to rename it as ``pd``, which is common practice.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
# ### Reading in the data
# Pandas allows us to [read in various structured files](https://pandas.pydata.org/pandas-docs/stable/user_guide/io.html) (e.g., .csv, .json, .html, etc) with just one line:
# we don't always need to specify the encoding, but this particular
# file has special characters that we need to handle
top50 = pd.read_csv("../data/top50.csv")
# ### High-level view of the data
# We can view the data frame by simply printing it:
top50
# Recall that we can also inspect the file by looking at just the first N rows or last N rows (instead of printing the entire dataframe).
# top50.head(5) # first 5 rows
top50.tail(3) # last 3 rows
# **Q2.1:** That's cool, but we can't see all of the columns too well. Write code to print out the columns of 'top50' and the code to calculate number of columns in it.
######
# your code here
######
# Fortunately, many of the features in our dataset are numeric. Conveniently, Pandas' `describe()` function calculates basic statistics for our columns. It's pretty amazing, as it allows us a very coarse-grain approach to understanding our data and checking for errors. That is, if we notice any summary statistics that are drastically different than what we deem reasonable, we should dive deeper and figure out why the values are what they are.
top50.describe()
# **Q2.2:** Which of the variable above appears to be the most skewed? Investigate its skew with a histogram.
# +
######
# your code here
######
# -
# *your answer here*
#
#
# Notice, it calculated statistics only for the columns that are of numeric data types. What about the textual ones (e.g., Track name and Artist)? Pandas is smart enough to infer the data types. **Don't forget to inspect the columns that are text-based though, as we need to ensure they are sound, too.**
#
# To view the data type of each column:
top50.dtypes
# **Q2.3:** Write code to obtain the table of frequencies for any categorical variables in the dataset.
#######
# your code here
#######
# ### Exploring the data
# I agree with Pandas' handling of the data. If any column contained floating point numbers, we would expect to see such here, too.
#
# Now that we've viewed our dataset at a high-level, let's actually use and explore it.
#
# Recall: we can **access a column of data** the same way we access dictionary by its keys:
top50["Length"]
# We could have also used this syntax (identical results):
top50.Length
# If we want just the highest or lowest **value** of a given column, we can use the functions ``max()`` and ``min()``, respectively.
top50['Length'].max()
top50['Length'].min()
# If we want the **row index** that corresponds to a column's max or min value, we can use ``idxmax()`` and ``idxmin()``, respectively.
top50['Length'].idxmax()
top50['Length'].idxmin()
# We can also add `conditional statements` (e.g., >, <, ==) for columns, which yields a boolean vector:
top50['Length'] > 240
# This is useful, as it allows us to process only the rows with the True values.
#
# The **`loc()`** function allows us to access data via labels:
# - A single scalar label
# - A list of labels
# - A slice object
# - A Boolean array
#
# A single scalar:
# single scalar label
top50.loc[0] # prints the (unnamed) row that has a label of 0 (the 1st row)
# list of labels
top50.loc[[0,2]] # prints the (unnamed) rows that have the labels of 0 and 2 (the 1st and 3rd rows)
# +
# a slice of the dataframe, based on the passed-in booleans;
# picture it's like a filter overlaying the DataFrame, and the filter
# dictates which values will be emitted/make it through to us
top50.loc[top50['Length'] > 240] # prints all rows that have Length > 240
# -
# Note, this returns a *DataFrame*. Everything we've learned so far concerns how to use DataFrames, so we can tack on additional syntax to this command if we wish to do further processing.
#
# For example, if we want to index just select columns (e.g., ArtistName, TrackName, and Length) of this returned DataFrame:
top50.loc[top50['Length'] > 240][['ArtistName', 'TrackName', 'Length']]
# Note, the above solves our original **Q1.2:** _(Write code to print all songs (Artist and Track name) that are longer than 4 minutes (240 seconds))_
#
# **Q2.4:** Write code to print the most popular song (or song(s) if there is a tie):
# +
#######
# your code here
#######
# -
# We can also sort our data by a single column! This pertains to our original **Q1.4**!
#
# **Q2.4:** Write code to print the songs (and their attributes), if we sorted by their popularity (highest scoring ones first).
# +
# use top50.sort_values() to answer this question
#######
# your code here
#######
# -
# While ``.loc()`` allows us to index based on passed-in labels, ``.iloc()`` allows us to **access data based on 0-based indices.**
#
# The syntax is ``.iloc[<row selection>, <column selection>]``, where <row selection> and <column selection> can be scalars, lists, or slices of indices.
top50.iloc[5:6] # prints all columns for the 6th row
top50.iloc[:,2] # prints all rows for the 3rd column
top50.iloc[[0,2,3], [2,1]] # prints the 1st, 3rd, and 4th rows of the 3rd and 2nd columns (artist and track)
# ### Inspecting/cleaning the data
#
# As mentioned, it is imperative to ensure the data is sound to use:
# 1. Did it come from a trustworthy, authoritative source?
# 2. Is the data a complete sample?
# 3. Does the data seem correct?
# 4. **(optional)** Is the data stored efficiently or does it have redundancies?
#
# Let's walk through each of these points now:
#
# 1. Did it come from a trustworthy, authoritative source?
#
# The data came from Kaggle.com, which anyone can publish to. However, the author claims that he/she used Spotify.com's official API to query songs in 2019. There are no public comments for it so far. It's potentially credible.
#
# 2. Is the data a complete sample?
#
# Pandas has functions named ``isnull()`` and ``notnull()``, which return DataFrames corresponding to any null or non-null entries, respectively.
#
# For example:
top50[top50.ArtistName.isnull()] # returns an empty DataFrame
top50[top50.ArtistName.notnull()] # returns the complete DataFrame since there are no null Artists
# If we run this for all of our features/columns, we will see there are no nulls. Since this dataset is manageable in size, you can also just scroll through it and notice no nulls.
#
# This answers our original **Q1.5**: How could you check for null/empty entries?
#
# Continuing with our data sanity check list:
#
# 3. Does the data seem correct?
#
# A quick scroll through the data, and we see a song by _Maluma_ titled _0.95833333_. This is possibly a song about probability, but I think the chances are slim. The song is 176 seconds long (2m56s). Looking on Spotify, we see **Maluma's most popular song is currently _11PM_ which is 2m56s in length!** Somehow, during the creation of the dataset, 11PM became 0.95833333. _Bonus points if you can figure out where this pointing number could have come from._
Image("fig/maluma.png") # sample of the data
# Since only one song seems obviously wrong, we can manually fix it. And it's worth noting such to ourselves and to whomever else would see our results or receive a copy of our data. If there were many more wrong values, we'd potentialy not fix them, as we'd explore other options.
top50['TrackName'][top50['ArtistName'] == "Maluma"] = "11PM"
# Watch out for the warning.
# ## Part 3: Grouping and Combining Multiple Data Frames
#
# As mentioned, often times one dataset doesn't contain all of the information you are interested in -- in which case, you need to combine data from multiple files. This also means you need to verify the accuracy (per above) of each dataset.
#
# Pandas' ``groupby()`` function splits the DataFrame into different groups, depending on the passed-in variable. For example, we can group our data by the genres:
grouped_df = top50.groupby('Genre')
#for key, item in grouped_df:
# print("Genre:", key, "(", len(grouped_df.get_group(key)), "items):", grouped_df.get_group(key), "\n\n")
# ``../data/spotify_aux.csv`` contains the same 50 songs as ``top50.csv``; however, it only contains 3 columns:
# - Track Name
# - Artist Name
# - Explicit Language (boolean valued)
#
# Note, that 3rd column is just random values, but pretend as if it's correct. The point of this section is to demonstrate how to merge columns together.
#
# Let's load ``../data/spotify_aux.csv`` into a DataFrame:
explicit_lyrics = pd.read_csv("../data/spotify_aux.csv")
#explicit_lyrics
# Let's merge it with our ``top50`` DataFrame.
#
# ``.merge()`` is a Pandas function that stitches together DataFrames by their columns.
#
# ``.concat()`` is a Pandas function that stitches together DataFrames by their rows (if you pass axis=1 as a flag, it will be column-based)
# 'on='' specifies the column used as the shared key
df_combined = pd.merge(explicit_lyrics, top50, on='TrackName')
#df_combined
# We see that all columns from both DataFrames have been added. That's nice, but having duplicate ArtistName and TrackName is unecessary. Since ``merge()`` uses DataFrames as the passed-in objects, we can simply pass merge() a stripped-down copy of _ExplicitLanguage_, which helps merge() not add any redundant fields.
df_combined = pd.merge(explicit_lyrics[['TrackName', 'ExplicitLanguage']], top50, on='TrackName')
#df_combined
# This answers our original **Q1.6:** Imagine we had another table (i.e., .csv file) below. How could we combine its data with our already-existing *dataset*?
# While we do not exhaustively illustrate Pandas' joining/splitting functionality, you may find the following functions useful:
# - ``merge()``
# - ``concat()``
# - ``aggregate()``
# - ``append()``
# ### Plotting DataFrames
# As a very simple example of how one can plot elements of a DataFrame, we turn to Pandas' built-in plotting:
scatter_plot = top50.plot.scatter(x='Danceability', y='Popularity', c='DarkBlue')
# **Q3.1:** Alternatively, use `plt.scatter` to recreate the scatterplot above.
#
# +
######
# your code here
######
# -
# This shows the lack of a correlation between the Danceability of a song and its popularity, based on just the top 50 songs, of course.
#
# Please feel free to experiment with plotting other items of interest, and we recommend using Seaborn.
#
# ## Practice Problems
#
# **Q3.2:** Print the shortest song (all features):
# +
######
# your code here
######
# -
# **Q3.3:** Print the 5 shortest songs (all features):
# +
######
# your code here
######
# -
# **Q3.4:** What is the average length of the 5 shortest songs?
# +
######
# your code here
######
# -
# **Q3.5:** How many distinct genres are present in the top 50 songs?
# +
######
# your code here
######
# -
# **Q3.6:** Print the songs that have a Danceability score above 80 and a popularity above 86. HINT: you can combine conditional statements with the & operator, and each item must be surrounded with ( ) brackets.
######
# your code here
######
# **Q3.7:** Plot a histogram of the Genre counts (x-axis is the Genres, y-axis is the # of songs with that Genre)
# +
######
# your code here
######
# -
# **Q3.8 (open ended):** Think of a _subset_ of the data that you're interested in. Think of an interesting plot that could be shown to illustrate that data. With a partner, discuss whose would be easier to create. Together, create that plot. Then, try to create the harder plot.
# +
######
# your code here
######
# -
# ## Part 4: Beautiful Soup
# Data Engineering, the process of gathering and preparing data for analysis, is a very big part of Data Science.
#
# Datasets might not be formatted in the way you need (e.g. you have categorical features but your algorithm requires numerical features); or you might need to cross-reference some dataset to another that has a different format; or you might be dealing with a dataset that contains missing or invalid data.
#
# These are just a few examples of why data retrieval and cleaning are so important.
#
# ---
# ### `requests`: Retrieving Data from the Web
# In HW1, you will be asked to retrieve some data from the Internet. `Python` has many built-in libraries that were developed over the years to do exactly that (e.g. `urllib`, `urllib2`, `urllib3`).
#
# However, these libraries are very low-level and somewhat hard to use. They become especially cumbersome when you need to issue POST requests or authenticate against a web service.
#
# Luckly, as with most tasks in `Python`, someone has developed a library that simplifies these tasks. In reality, the requests made both on this lab and on HW1 are fairly simple, and could easily be done using one of the built-in libraries. However, it is better to get acquainted to `requests` as soon as possible, since you will probably need it in the future.
# You tell Python that you want to use a library with the import statement.
import requests
# Now that the requests library was imported into our namespace, we can use the functions offered by it.
#
# In this case we'll use the appropriately named `get` function to issue a *GET* request. This is equivalent to typing a URL into your browser and hitting enter.
# Get the HU Wikipedia page
req = requests.get("https://en.wikipedia.org/wiki/Harvard_University")
# Python is an Object Oriented language, and everything on it is an object. Even built-in functions such as `len` are just syntactic sugar for acting on object properties.
#
# We will not dwell too long on OO concepts, but some of Python's idiosyncrasies will be easier to understand if we spend a few minutes on this subject.
#
# When you evaluate an object itself, such as the `req` object we created above, Python will automatially call the `__str__()` or `__repr__()` method of that object. The default values for these methods are usually very simple and boring. The `req` object however has a custom implementation that shows the object type (i.e. `Response`) and the HTTP status number (200 means the request was successful).
req
# Just to confirm, we will call the `type` function on the object to make sure it agrees with the value above.
type(req)
# Another very nifty Python function is `dir`. You can use it to list all the properties of an object.
#
# By the way, properties starting with a single and double underscores are usually not meant to be called directly.
dir(req)
# Right now `req` holds a reference to a *Request* object; but we are interested in the text associated with the web page, not the object itself.
#
# So the next step is to assign the value of the `text` property of this `Request` object to a variable.
page = req.text
page[20000:30000]
# Great! Now we have the text of the Harvard University Wikipedia page. But this mess of HTML tags would be a pain to parse manually. Which is why we will use another very cool Python library called `BeautifulSoup`.
# ### `BeautifulSoup`
#
# Parsing data would be a breeze if we could always use well formatted data sources, such as CSV, JSON, or XML; but some formats such as HTML are at the same time a very popular and a pain to parse.
#
# One of the problems with HTML is that over the years browsers have evolved to be very forgiving of "malformed" syntax. Your browser is smart enough to detect some common problems, such as open tags, and correct them on the fly.
#
# Unfortunately, we do not have the time or patience to implement all the different corner cases, so we'll let BeautifulSoup do that for us.
#
# You'll notice that the `import` statement bellow is different from what we used for `requests`. The _from library import thing_ pattern is useful when you don't want to reference a function byt its full name (like we did with `requests.get`), but you also don't want to import every single thing on that library into your namespace.
from bs4 import BeautifulSoup
# `BeautifulSoup` can deal with `HTML` or `XML` data, so the next line parses the contents of the `page` variable using its `HTML` parser, and assigns the result of that to the `soup` variable.
soup = BeautifulSoup(page, 'html.parser')
type(soup)
# Doesn't look much different from the `page` object representation. Let's make sure the two are different types.
type(page)
# Looks like they are indeed different.
#
# `BeautifulSoup` objects have a cool little method that allows you to see the `HTML` content in a nice, indented way.
print(soup.prettify()[:1000])
# Looks like it's our page!
#
# We can now reference elements of the `HTML` document in different ways. One very convenient way is by using the dot notation, which allows us to access the elements as if they were properties of the object.
soup.title
# This is nice for `HTML` elements that only appear once per page, such the the `title` tag. But what about elements that can appear multiple times?
# Be careful with elements that show up multiple times.
soup.p
# Uh Oh. Turns out the attribute syntax in `Beautiful` soup is what is called *syntactic sugar*. That's why it is safer to use the explicit commands behind that syntactic sugar I mentioned. These are:
# * `BeautifulSoup.find` for getting single elements, and
# * `BeautifulSoup.find_all` for retrieving multiple elements.
len(soup.find_all("p"))
# If you look at the Wikipedia page on your browser, you'll notice that it has a couple of tables in it. We will be working with the "Demographics" table, but first we need to find it.
#
# One of the `HTML` attributes that will be very useful to us is the `class` attribute.
#
# Getting the class of a single element is easy!
soup.table["class"]
# Next we will use a *list comprehension* to see all the tables that have a `class` attribute.
# the classes of all tables that have a class attribute set on them
[t["class"] for t in soup.find_all("table") if t.get("class")]
# As already mentioned, we will be using the Demographics table for this lab. The next cell contains the `HTML` elements of said table. We will render it in different parts of the notebook to make it easier to follow along the parsing steps.
table_demographics = soup.find_all("table", "wikitable")[2]
from IPython.core.display import HTML
HTML(str(table_demographics))
# First we'll use a list comprehension to extract the rows (*tr*) elements.
rows = [row for row in table_demographics.find_all("tr")]
print(rows)
header_row = rows[0]
HTML(str(header_row))
# We will then use a `lambda` expression to replace new line characters with spaces. `Lambda` expressions are to functions what list comprehensions are to lists: namely a more concise way to achieve the same thing.
#
# In reality, both lambda expressions and list comprehensions are a little different from their function and loop counterparts. But for the purposes of this class we can ignore those differences.
# Lambda expressions return the value of the expression inside it.
# In this case, it will return a string with new line characters replaced by spaces.
rem_nl = lambda s: s.replace("\n", " ")
# #### Splitting the data
# Next we extract the text value of the columns. If you look at the table above, you'll see that we have three columns and six rows.
#
# Here we're doing the following:
# * Taking the first element (`Python` indices start at zero)
# * Iterating over the *th* elements inside it
# * Taking the text value of those elements
#
# We should end up with a list of column names.
#
# But there is one little caveat: the first column of the table is actually an empty string (look at the cell right above the row names). We could add it to our list and then remove it afterwards; but instead we will use the `if` statement inside the list comprehension to filter that out.
#
# In the following cell, `get_text` will return an empty string for the first cell of the table, which means that the test will fail and the value will not be added to the list.
# the if col.get_text() takes care of no-text in the upper left
columns = [rem_nl(col.get_text()) for col in header_row.find_all("th") if col.get_text()]
columns
# Now let's do the same for the rows. Notice that since we have already parsed the header row, we will continue from the second row.
indexes = [row.find("th").get_text() for row in rows[1:]]
indexes
# Now we want to transform the string on the cells to integers. To do this, we follow a very common `python` pattern:
# 1. Check if the last character of the string is a percent sign
# 2. If it is, then convert the characters before the percent sign to integers
# 3. If one of the prior checks fails, return a value of `None`
#
# These steps can be conveniently packaged into a function using `if-else` statements.
def to_num(s):
if s[-1] == "%":
return int(s[:-1])
else:
return None
# Notice the `Python` slices are open on the upper bound. So the `[:-1]` construct will return all elements of the string, except for the last.
# Another nice way to write our `to_num` function would be
# ```python
# def to_num(s):
# return int(s[:-1]) if s[-1] == "%" else None
# ```
# Notice that we only had to write `return` one time and everything conveniently fits on one line. I'll leave it up to you to decide if it's readable or not.
# Now we use the `to_num` function in a list comprehension to parse the table values.
#
# Notice that we have two `for ... in ...` in this list comprehension. That is perfectly valid and somewhat common.
#
# Although there is no real limit to how many iterations you can perform at once, having more than two can be visually unpleasant, at which point either regular nested loops or saving intermediate comprehensions might be a better solution.
values = [to_num(value.get_text()) for row in rows[1:] for value in row.find_all("td")]
values
# The problem with the list above is that the values lost their grouping.
#
# The `zip` function is used to combine two sequences element wise. So
# ```python
# zip([1,2,3], [4,5,6])
# ```
# would return
# ```python
# [(1, 4), (2, 5), (3, 6)]
# ```
#
# Next we create three arrays corresponding to the three columns by putting every three values in each list.
stacked_values_lists = [values[i::3] for i in range(len(columns))]
stacked_values_lists
# We then use `zip`.
stacked_values = zip(*stacked_values_lists)
list(stacked_values)
# Notice the use of the `*` in front: that converts the list of lists to a set of arguments to `zip`. See the ASIDE below.
# Here's the original HTML table for visual understanding
HTML(str(table_demographics))
# **Q4.1:** Use the tables in `soup` to determine how Harvard's Computer Science program ranks both Nationally and Globally.
# +
######
# your code here
######
| content/lectures/lecture2/notebook/Lecture2_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Valid customer IDs
#
# Simon runs a pharmaceutical factory. Only customers who have been pre-approved can enter the manufacturing facility and do a quality check for their order.
# The pre-approved customers get a customer ID which needs to be validated at entrance.
#
# A valid customer ID:
# -It will have groups of three characters separated by "_".
# -Can only contain numbers 0-9 and characters a-z and A-Z.(other than "_")
# -Contains 12 characters excluding the "_"
# -A character cannot be repeated more than thrice.
#
# You have to write Python code to check for valid customer IDs.
# The input will have a customer ID and the output should say "valid" if the entered ID is valid otherwise it should say "invalid".
#
# **Sample Input:**
# abc_123_2ac_Adf
#
# **Sample Output:**
# valid
# +
def validitycheck(cid):
#req1: a-z and 0-9 only
if not (cid.replace("_","").isalnum()):
return ("invalid")
#req2 & 3 :4 groups, each group has three characters with a-z, A-Z and 0-9
groups=cid.split("_")
if len(groups)>4:
return ("invalid")
for group in groups:
if len(group)>3:
return ("invalid")
#Req 5: repeating characters check
from collections import Counter
counts=dict(Counter(cid))
for key in counts.keys():
if counts.get(key)>3:
return("invalid")
return("valid")
cid=input()
print(validitycheck(cid))
# -
# ## Password Validation
#
# Your company requires employees to set a strong password for their email accounts.
#
# The characteristics of a strong password include:
#
# 1. it should be at least 8 characters long
#
# 2. it should have one lowercase alphabet.
#
# 3. it should have one uppercase alphabet
#
# 4. it should have at least one number(0-9)
#
# 5. it should have one special character( a special character is considered among the following: [@%$*])
#
# If the input is a valid password then print "valid" or else print "invalid".
#
#
#
# **Sample input:**
#
# DataScience123
#
#
# **Sample Output:**
#
# invalid
pwd=input()
import re
if ((len(pwd)>=8) and (re.search("[a-z]",pwd)) and (re.search("[A-Z]",pwd)) and (re.search("[0-9]",pwd)) and (re.search("[@%$*]",pwd))):
print("valid")
else:
print("invalid")
# ## Divide the dataframe
#
# Write a Python program to slice a dataframe in a given ratio.
# For example: if the dataframe has 160 rows and you have to slice it in a ratio of 1:3, then the first part will have the first
# 40 rows(0-39) and the second part will have the next 120 rows(40-159).
#
# The input will have two lines with the ratio to separate the dataframe in. For example, for 1:3, the input will be as follows:
# 1
# 3
#
# The output should contain the summary statistics(df.describe()) of both resulting dataframes in the respective order.
# Note: You can assume that the given ratio will split the dataframe exactly into two non-fractional parts.
#
# **Sample Input:**
# 10
# 30
#
import pandas as pd
df=pd.read_csv("https://media-doselect.s3.amazonaws.com/generic/A0zOxQvk78ONwRgLZ1WYJOxWq/titaniMod2.csv")
a=int(input())
b=int(input())
c=int(a/(a+b)*len(df))
print(df[:c].describe())
print(df[c:].describe())
# ## Flatten a dictionary
#
# Consider a nested dictionary as follows:
# {'Fruit': 1, 'Vegetable': {'Cabbage': 2, 'Cauliflower': 3}, 'Spices': 4}
# Your task is to flatten a nested dictionary and join the nested keys with the "_" character. For the above dictionary, the flattened dictionary would be as follows:
# {'Fruit': 1, 'Vegetable_Cabbage': 2, 'Vegetable_Cauliflower': 3, 'Spices': 4}
#
# The input will have a nested dictionary.
# The output should have two lists. The first list will have keys and the second list should have values. Both lists should be sorted.
#
# **Sample Input:**
# {'Fruit': 1, 'Vegetable': {'Cabbage': 2, 'Cauliflower': 3}, 'Spices': 4}
#
# **Sample Output:**
# ['Fruit', 'Spices', 'Vegetable_Cabbage', 'Vegetable_Cauliflower']
# [1, 2, 3, 4]
# +
#import ast,sys
#input_str = sys.stdin.read()
#input_dict = dict(ast.literal_eval(input_str))
input_dict = {'Fruit': 1, 'Vegetable': {'Cabbage': 2, 'Cauliflower': 3}, 'Spices': 4}
def flatten_dict(d):
def expand(key, value):
if isinstance(value, dict):
return [ (key + '_' + k, v) for k, v in flatten_dict(value).items() ]
else:
return [ (key, value) ]
items = [ item for k, v in d.items() for item in expand(k, v) ]
return dict(items)
out1=list(flatten_dict(input_dict).keys())
out2=list(flatten_dict(input_dict).values())
out1.sort()
out2.sort()
print(out1)
print(out2)
# -
# ## 2-Sample t-test
#
# Perform 2-sample t-tests on given columns of the dataframe.
# The input will contain the names of two columns to test in two lines and the output should have the p-value obtained from the paired two sample tests.
#
# **Sample Input:**
# city-mpg
# highway-mpg
#
# **Sample Output:**
# 1.9665445899143185e-113
import pandas as pd
import scipy.stats
df=pd.read_csv("https://media-doselect.s3.amazonaws.com/generic/K9WgyRZ75q4Pkdp38AQabgE0X/Automobile_data.csv")
col1=input()
col2=input()
print(scipy.stats.ttest_rel(df[col1],df[col2]).pvalue)
# ## t-test on a column
# Perform a t-test on a given column of a given data frame.
# The input will contain the column name and the value of x, the mean value to test in two lines respectively. The output should contain the p-value obtained.
#
# **Sample Input:**
# CGPA
# 8
#
# **Sample Output:**
# 1.6070878523226457e-62
import pandas as pd
import scipy.stats
df=pd.read_csv("https://media-doselect.s3.amazonaws.com/generic/5J7nrZegVWOORbGr4M7KVPXE5/Admission_Predict.csv")
col=input()
x=float(input())
print(scipy.stats.ttest_1samp(df[col], x).pvalue)
# ## List Overlap
#
# Write Python code to find elements common between the two lists.
# The output list should exclude duplicate elements. i.e. if both lists have 1 twice then the output list should have 1 only once.
# The input will contain two lines with two lists.
# The output should contain a list of common elements between the two input lists.
#
# **Sample Input:**
# [1,2,3,4,5]
# [4,5,6,7,8]
#
# **Sample Output:**
# [4, 5]
# +
#import ast,sys
#input_str = sys.stdin.read()
#inp = ast.literal_eval(input_str)
list1=[1,2,3,4,5]
#first list
list2= [4,5,6,7,8]
#second list
print(list(set(list1).intersection(set(list2))))
# -
# ## Sorting based on one column
#
# Given a dataframe, you have to sort the rows based on values of one column.
# Note: Sorting should be in descending order of values of given column
#
# The input will contain a column name. The output should contain the first five rows of the dataframe.
# The output will contain the first n rows of the sorted dataframe.
#
# **Sample Input:**
# TOEFL Score
import pandas as pd
col=input()
df=pd.read_csv("https://media-doselect.s3.amazonaws.com/generic/RM8r5NBrJdA4QeVZXvwbjokwv/Admission_Predict.csv")
print(df.sort_values(by=[col],ascending=False).head())
# ## Survival of?
#
# Given is a dataframe with data of passengers of the ship Titanic.
# Here, the "Survived" column has "1" if the passenger has survived. Otherwise, it contains "0". The Pclass column indicates the class the passenger was travelling in(1st class, 2nd class and so on).
#
# Write a Pandas program to create a Pivot table and find survival rate by the given column on various classes according to Pclass.
#
# **Sample Input:**
# Sex
import pandas as pd
df=pd.read_csv("https://media-doselect.s3.amazonaws.com/generic/pLMXoA0GZNAPMRNrdnn88pOxb/train.csv")
col=input()
print(df.pivot_table('Survived', index=[col], columns='Pclass'))
| Hypothesis Coding Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 
#
# # Unit 3 Lab: Object-Oriented Programming
#
# ## Overview
#
# Welcome to the Unit 3 lab! Now that we've learned some new data structures, let's put them to use in our weather forecasting application.
#
# ### Goals
#
# In this lab, you will:
#
# - Write and use complex data types to organize weather data efficiently.
# - Write classes and methods to calculate and display the weather forecast.
#
#
# ---
# ### Restructuring Data
#
# Suppose we want to offer the user the opportunity to select the temperature scale in which they view their weather forecast. Create a variable, `temperature_scale`, that is a `set` with the following strings:
#
# - "Fahrenheit"
# - "Celsius"
# - "Kelvin"
#
# Print the variable `temperature_scale`.
# Enter your code below:
temperature_scale = {"Farenheit", "Celsius", "Kelvin"}
print(temperature_scale)
# Weather data is typically taken and recorded every hour. Let's combine the following lists into a single, table-like structure of data we can review.
#
# Create a list called `weather_data`. This list will be composed of dictionary objects for each hour over a 24-hour span. Each of the dictionaries should have the following four keys: `"hour"`, `"temperature"`, `"humidity"`, and `"rainfall"`. Begin counting at hour `0` and end at hour `23`.
#
# Then, iterate through `weather_data` and print out each dictionary — one dictionary per line.
#
# +
one_day_of_hourly_temperatures = [67,67,68,69,71,73,75,76,79,81,81,80,82,81,81,80,78,75,72,70,67,65,66,66]
one_day_of_hourly_humidity = [60,65,65,70,70,70,70,75,75,75,75,80,80,85,85,85,85,80,80,80,80,80,80,80]
one_day_of_hourly_rainfall = [0,0,0,0.1,0.1,0.05,0.1,0.15,0.2,0.3,0.3,0.5,0,0,0,0,0,0,0,0,0,0,0,0]
# Enter your code below:
weather_data = []
for i in range(len(one_day_of_hourly_temperatures)):
current_hour = {
"hour": i,
"temperature": one_day_of_hourly_temperatures[i],
"humidity": one_day_of_hourly_humidity[i],
"rainfall": one_day_of_hourly_rainfall[i]
}
weather_data.append(current_hour)
for hour in weather_data:
print(hour)
# -
# Using your `weather_data` list, print the following with one line of code for each `print` statement:
# - Temperature at 2 p.m.
# - Humidity at 11 p.m.
# - Rainfall at 9 a.m.
# Enter your code below:
print(weather_data[14]["temperature"])
print(weather_data[23]["humidity"])
print(weather_data[9]["rainfall"])
# ---
# ### Creating a `Forecast` Class
#
# Start by creating a `Forecast` class that accepts one argument, `location`, and assigns it as an instance attribute to `self.location`.
# Enter your code below:
class Forecast():
def __init__(self, location):
self.location = location
# ---
# ### Calculation Methods
#
# Create two methods within the `Forecast` class:
#
# - `get_daily_high()`
# - `get_daily_low()`
#
# Use the built-in `max()` and `min()` functions on `one_day_of_hourly_temperatures` to return the daily high and low temperatures.
#
# Create a third method within the `Forecast` class called `get_daily_chance_of_rain()`. This method should:
# - Create a variable named `number_of_years_of_data` and set it to `10`.
# - Create a variable named `times_it_has_rained` and set it to `0`.
# - Calculate the sum of rainfall for all 24 hours of `one_day_of_hourly_temperatures`.
# - If the sum of all 24 hours is greater than 0, increase `times_it_has_rained` by 1.
# - Convert `times_it_has_rained` to a percentage by dividing it by `number_of_years_of_data` and multiplying it by 100.
# - Return the final value.
#
# _Note: We'll modify these three methods to process live data in future labs. For now, we're setting up the basic logic on test data._
# Copy and paste the Forecast class you built in the previous cell and add the new features to it below:
class Forecast():
def __init__(self, location):
self.location = location
def get_daily_high(self):
return max(one_day_of_hourly_temperatures)
def get_daily_low(self):
return min(one_day_of_hourly_temperatures)
def get_daily_chance_of_rain(self):
number_of_years_of_data = 10
times_it_has_rained = 0
if sum(one_day_of_hourly_rainfall) > 0:
times_it_has_rained += 1
return times_it_has_rained / number_of_years_of_data * 100
# Run the following cell after you're finished to test your output:
test = Forecast("Austin,TX")
print("High:", test.get_daily_high())
print("Low:", test.get_daily_low())
print("Chance of Rain:", test.get_daily_chance_of_rain(),'%')
# ---
# ### Display Methods
#
# Back in Lab 1, we printed `"The weather forecast for today is: High of 85, low of 69, with a 15.0% chance of precipitation."` using the four following variables and the values we assigned to them:
#
# - `message_to_user`
# - `todays_high`
# - `todays_low`
# - `chance_of_precipitation`
#
# Now, we're going to display this same message but with the calculated values from the methods we just created.
#
# Create a `display_daily_forecast()` method within the `Forecast` class that, when called, will print out a string matching the one above.
#
# This method will insert the high temperature, low temperature, and chance of rain by calling the following internal class methods, respectively:
#
# - `get_daily_high()`
# - `get_daily_low()`
# - `get_daily_chance_of_rain()`
#
#
# Next, create a `display_weekly_forecast()` method within the `Forecast` class:
# - When called, this method will print out a message in the format shown in the code block below.
# - _Note: We only have one day of data we're currently using, so `High`, `Low`, and `Rain` will all be the same for now._
#
# ```
# This week's weather forecast:
# Monday: High 82, Low 65, Rain 10.0%
# Tuesday: High 82, Low 65, Rain 10.0%
# Wednesday: High 82, Low 65, Rain 10.0%
# Thursday: High 82, Low 65, Rain 10.0%
# Friday: High 82, Low 65, Rain 10.0%
# Saturday: High 82, Low 65, Rain 10.0%
# Sunday: High 82, Low 65, Rain 10.0%
# ```
#
# _Hint: Use `\n` to create new lines in your `print` statement (i.e., `print("First line\nSecond line")`)._
#
# _Hint: Use `\t` to simulate a tab character in your `print` statement (i.e., `print("First Column\tSecond Column")`._
#
# _Bonus: Convert class methods that are internally called to private methods via the use of the `'underscore'`, `__method_name`._
# Copy and paste the Forecast class you built in the previous cell and add the new features to it below:
class Forecast():
def __init__(self, location):
self.location = location
def __get_daily_high(self):
return max(one_day_of_hourly_temperatures)
def __get_daily_low(self):
return min(one_day_of_hourly_temperatures)
def __get_daily_chance_of_rain(self):
number_of_years_of_data = 10
times_it_has_rained = 0
if sum(one_day_of_hourly_rainfall):
times_it_has_rained += 1
return times_it_has_rained / number_of_years_of_data * 100
def display_daily_forecast(self):
print(f"The weather forecast for today in {self.location}"
f" is: High of {self.__get_daily_high()}, Low of "
f"{self.__get_daily_low()}, with a "
f"{self.__get_daily_chance_of_rain()}% chance of rain.")
def display_weekly_forecast(self):
print(f"The week's weather forecast for: "
f"\n\tMonday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tTuesday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tWednesday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tThursday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tFriday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tSaturday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}"
f"\n\tSunday: High {self.__get_daily_high()}, Low {self.__get_daily_low()}, Rain {self.__get_daily_chance_of_rain()}")
# Run the following cell after you're finished to test your output:
test = Forecast("Austin,TX")
test.display_daily_forecast()
test.display_weekly_forecast()
# ### Nice work!
| unit-3-oop/instructor-resources/14-unit-lab-3/pyth621-day3-lab-solutions.ipynb |
# Transformers installation
# ! pip install transformers
# To install from source instead of the last release, comment the command above and uncomment the following one.
# # ! pip install git+https://github.com/huggingface/transformers.git
# # Fine-tuning with custom datasets
# > **NOTE:** The datasets used in this tutorial are available and can be more easily accessed using the [🤗 NLP library](https://github.com/huggingface/nlp). We do not use this library to access the datasets here since this tutorial
# > meant to illustrate how to work with your own data. A brief of introduction can be found at the end of the tutorial
# > in the section "[nlplib](#nlplib)".
# This tutorial will take you through several examples of using 🤗 Transformers models with your own datasets. The guide
# shows one of many valid workflows for using these models and is meant to be illustrative rather than definitive. We
# show examples of reading in several data formats, preprocessing the data for several types of tasks, and then preparing
# the data into PyTorch/TensorFlow `Dataset` objects which can easily be used either with
# `Trainer`/`TFTrainer` or with native PyTorch/TensorFlow.
#
# We include several examples, each of which demonstrates a different type of common downstream task:
#
# - [seq_imdb](#seq_imdb)
# - [tok_ner](#tok_ner)
# - [qa_squad](#qa_squad)
# - [resources](#resources)
# <a id='seq_imdb'></a>
# ## Sequence Classification with IMDb Reviews
# > **NOTE:** This dataset can be explored in the Hugging Face model hub ([IMDb](https://huggingface.co/datasets/imdb)), and
# > can be alternatively downloaded with the 🤗 NLP library with `load_dataset("imdb")`.
# In this example, we'll show how to download, tokenize, and train a model on the IMDb reviews dataset. This task takes
# the text of a review and requires the model to predict whether the sentiment of the review is positive or negative.
# Let's start by downloading the dataset from the [Large Movie Review Dataset](http://ai.stanford.edu/~amaas/data/sentiment/) webpage.
# ! wget http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
# ! tar -xf aclImdb_v1.tar.gz
# This data is organized into `pos` and `neg` folders with one text file per example. Let's write a function that can
# read this in.
# +
from pathlib import Path
def read_imdb_split(split_dir):
split_dir = Path(split_dir)
texts = []
labels = []
for label_dir in ["pos", "neg"]:
for text_file in (split_dir/label_dir).iterdir():
texts.append(text_file.read_text())
labels.append(0 if label_dir is "neg" else 1)
return texts, labels
train_texts, train_labels = read_imdb_split('aclImdb/train')
test_texts, test_labels = read_imdb_split('aclImdb/test')
# -
# We now have a train and test dataset, but let's also also create a validation set which we can use for for evaluation
# and tuning without training our test set results. Sklearn has a convenient utility for creating such splits:
from sklearn.model_selection import train_test_split
train_texts, val_texts, train_labels, val_labels = train_test_split(train_texts, train_labels, test_size=.2)
# Alright, we've read in our dataset. Now let's tackle tokenization. We'll eventually train a classifier using
# pre-trained DistilBert, so let's use the DistilBert tokenizer.
from transformers import DistilBertTokenizerFast
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-uncased')
# Now we can simply pass our texts to the tokenizer. We'll pass `truncation=True` and `padding=True`, which will
# ensure that all of our sequences are padded to the same length and are truncated to be no longer model's maximum input
# length. This will allow us to feed batches of sequences into the model at the same time.
train_encodings = tokenizer(train_texts, truncation=True, padding=True)
val_encodings = tokenizer(val_texts, truncation=True, padding=True)
test_encodings = tokenizer(test_texts, truncation=True, padding=True)
# Now, let's turn our labels and encodings into a Dataset object. In PyTorch, this is done by subclassing a
# `torch.utils.data.Dataset` object and implementing `__len__` and `__getitem__`. In TensorFlow, we pass our input
# encodings and labels to the `from_tensor_slices` constructor method. We put the data in this format so that the data
# can be easily batched such that each key in the batch encoding corresponds to a named parameter of the
# `DistilBertForSequenceClassification.forward` method of the model we will train.
# +
import tensorflow as tf
train_dataset = tf.data.Dataset.from_tensor_slices((
dict(train_encodings),
train_labels
))
val_dataset = tf.data.Dataset.from_tensor_slices((
dict(val_encodings),
val_labels
))
test_dataset = tf.data.Dataset.from_tensor_slices((
dict(test_encodings),
test_labels
))
# -
# Now that our datasets our ready, we can fine-tune a model either with the 🤗
# `Trainer`/`TFTrainer` or with native PyTorch/TensorFlow. See [training](https://huggingface.co/transformers/training.html).
# <a id='ft_trainer'></a>
# ### Fine-tuning with Trainer
# The steps above prepared the datasets in the way that the trainer is expected. Now all we need to do is create a model
# to fine-tune, define the `TrainingArguments`/`TFTrainingArguments` and
# instantiate a `Trainer`/`TFTrainer`.
# +
from transformers import TFDistilBertForSequenceClassification, TFTrainer, TFTrainingArguments
training_args = TFTrainingArguments(
output_dir='./results', # output directory
num_train_epochs=3, # total number of training epochs
per_device_train_batch_size=16, # batch size per device during training
per_device_eval_batch_size=64, # batch size for evaluation
warmup_steps=500, # number of warmup steps for learning rate scheduler
weight_decay=0.01, # strength of weight decay
logging_dir='./logs', # directory for storing logs
logging_steps=10,
)
with training_args.strategy.scope():
model = TFDistilBertForSequenceClassification.from_pretrained("distilbert-base-uncased")
trainer = TFTrainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_dataset, # training dataset
eval_dataset=val_dataset # evaluation dataset
)
trainer.train()
# -
# <a id='ft_native'></a>
# ### Fine-tuning with native PyTorch/TensorFlow
# We can also train use native PyTorch or TensorFlow:
# +
from transformers import TFDistilBertForSequenceClassification
model = TFDistilBertForSequenceClassification.from_pretrained('distilbert-base-uncased')
optimizer = tf.keras.optimizers.Adam(learning_rate=5e-5)
model.compile(optimizer=optimizer, loss=model.compute_loss) # can also use any keras loss fn
model.fit(train_dataset.shuffle(1000).batch(16), epochs=3, batch_size=16)
# -
# <a id='tok_ner'></a>
# ## Token Classification with W-NUT Emerging Entities
# > **NOTE:** This dataset can be explored in the Hugging Face model hub ([WNUT-17](https://huggingface.co/datasets/wnut_17)),
# > and can be alternatively downloaded with the 🤗 NLP library with `load_dataset("wnut_17")`.
# Next we will look at token classification. Rather than classifying an entire sequence, this task classifies token by
# token. We'll demonstrate how to do this with [Named Entity Recognition](http://nlpprogress.com/english/named_entity_recognition.html), which involves identifying tokens which correspond to
# a predefined set of "entities". Specifically, we'll use the [W-NUT Emerging and Rare entities](http://noisy-text.github.io/2017/emerging-rare-entities.html) corpus. The data is given as a collection of
# pre-tokenized documents where each token is assigned a tag.
#
# Let's start by downloading the data.
# ! wget http://noisy-text.github.io/2017/files/wnut17train.conll
# In this case, we'll just download the train set, which is a single text file. Each line of the file contains either (1)
# a word and tag separated by a tab, or (2) a blank line indicating the end of a document. Let's write a function to read
# this in. We'll take in the file path and return `token_docs` which is a list of lists of token strings, and
# `token_tags` which is a list of lists of tag strings.
# +
from pathlib import Path
import re
def read_wnut(file_path):
file_path = Path(file_path)
raw_text = file_path.read_text().strip()
raw_docs = re.split(r'\n\t?\n', raw_text)
token_docs = []
tag_docs = []
for doc in raw_docs:
tokens = []
tags = []
for line in doc.split('\n'):
token, tag = line.split('\t')
tokens.append(token)
tags.append(tag)
token_docs.append(tokens)
tag_docs.append(tags)
return token_docs, tag_docs
texts, tags = read_wnut('wnut17train.conll')
# -
# Just to see what this data looks like, let's take a look at a segment of the first document.
print(texts[0][10:17], tags[0][10:17], sep='\n')
# `location` is an entity type, `B-` indicates the beginning of an entity, and `I-` indicates consecutive positions
# of the same entity ("Empire State Building" is considered one entity). `O` indicates the token does not correspond to
# any entity.
#
# Now that we've read the data in, let's create a train/validation split:
from sklearn.model_selection import train_test_split
train_texts, val_texts, train_tags, val_tags = train_test_split(texts, tags, test_size=.2)
# Next, let's create encodings for our tokens and tags. For the tags, we can start by just create a simple mapping which
# we'll use in a moment:
unique_tags = set(tag for doc in tags for tag in doc)
tag2id = {tag: id for id, tag in enumerate(unique_tags)}
id2tag = {id: tag for tag, id in tag2id.items()}
# To encode the tokens, we'll use a pre-trained DistilBert tokenizer. We can tell the tokenizer that we're dealing with
# ready-split tokens rather than full sentence strings by passing `is_split_into_words=True`. We'll also pass
# `padding=True` and `truncation=True` to pad the sequences to be the same length. Lastly, we can tell the model to
# return information about the tokens which are split by the wordpiece tokenization process, which we will need in a
# moment.
from transformers import DistilBertTokenizerFast
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-cased')
train_encodings = tokenizer(train_texts, is_split_into_words=True, return_offsets_mapping=True, padding=True, truncation=True)
val_encodings = tokenizer(val_texts, is_split_into_words=True, return_offsets_mapping=True, padding=True, truncation=True)
# Great, so now our tokens are nicely encoded in the format that they need to be in to feed them into our DistilBert
# model below.
#
# Now we arrive at a common obstacle with using pre-trained models for token-level classification: many of the tokens in
# the W-NUT corpus are not in DistilBert's vocabulary. Bert and many models like it use a method called WordPiece
# Tokenization, meaning that single words are split into multiple tokens such that each token is likely to be in the
# vocabulary. For example, DistilBert's tokenizer would split the Twitter handle `@huggingface` into the tokens `['@',
# 'hugging', '##face']`. This is a problem for us because we have exactly one tag per token. If the tokenizer splits a
# token into multiple sub-tokens, then we will end up with a mismatch between our tokens and our labels.
#
# One way to handle this is to only train on the tag labels for the first subtoken of a split token. We can do this in 🤗
# Transformers by setting the labels we wish to ignore to `-100`. In the example above, if the label for
# `@HuggingFace` is `3` (indexing `B-corporation`), we would set the labels of `['@', 'hugging', '##face']` to
# `[3, -100, -100]`.
#
# Let's write a function to do this. This is where we will use the `offset_mapping` from the tokenizer as mentioned
# above. For each sub-token returned by the tokenizer, the offset mapping gives us a tuple indicating the sub-token's
# start position and end position relative to the original token it was split from. That means that if the first position
# in the tuple is anything other than `0`, we will set its corresponding label to `-100`. While we're at it, we can
# also set labels to `-100` if the second position of the offset mapping is `0`, since this means it must be a
# special token like `[PAD]` or `[CLS]`.
# > **NOTE:** Due to a recently fixed bug, -1 must be used instead of -100 when using TensorFlow in 🤗 Transformers <= 3.02.
# +
import numpy as np
def encode_tags(tags, encodings):
labels = [[tag2id[tag] for tag in doc] for doc in tags]
encoded_labels = []
for doc_labels, doc_offset in zip(labels, encodings.offset_mapping):
# create an empty array of -100
doc_enc_labels = np.ones(len(doc_offset),dtype=int) * -100
arr_offset = np.array(doc_offset)
# set labels whose first offset position is 0 and the second is not 0
doc_enc_labels[(arr_offset[:,0] == 0) & (arr_offset[:,1] != 0)] = doc_labels
encoded_labels.append(doc_enc_labels.tolist())
return encoded_labels
train_labels = encode_tags(train_tags, train_encodings)
val_labels = encode_tags(val_tags, val_encodings)
# -
# The hard part is now done. Just as in the sequence classification example above, we can create a dataset object:
# +
import tensorflow as tf
train_encodings.pop("offset_mapping") # we don't want to pass this to the model
val_encodings.pop("offset_mapping")
train_dataset = tf.data.Dataset.from_tensor_slices((
dict(train_encodings),
train_labels
))
val_dataset = tf.data.Dataset.from_tensor_slices((
dict(val_encodings),
val_labels
))
# -
# Now load in a token classification model and specify the number of labels:
from transformers import TFDistilBertForTokenClassification
model = TFDistilBertForTokenClassification.from_pretrained('distilbert-base-cased', num_labels=len(unique_tags))
# The data and model are both ready to go. You can train the model either with
# `Trainer`/`TFTrainer` or with native PyTorch/TensorFlow, exactly as in the
# sequence classification example above.
#
# - [ft_trainer](#ft_trainer)
# - [ft_native](#ft_native)
# <a id='qa_squad'></a>
# ## Question Answering with SQuAD 2.0
# > **NOTE:** This dataset can be explored in the Hugging Face model hub ([SQuAD V2](https://huggingface.co/datasets/squad_v2)), and can be alternatively downloaded with the 🤗 NLP library with
# > `load_dataset("squad_v2")`.
# Question answering comes in many forms. In this example, we'll look at the particular type of extractive QA that
# involves answering a question about a passage by highlighting the segment of the passage that answers the question.
# This involves fine-tuning a model which predicts a start position and an end position in the passage. We will use the
# [Stanford Question Answering Dataset (SQuAD) 2.0](https://rajpurkar.github.io/SQuAD-explorer/).
#
# We will start by downloading the data:
# ! mkdir squad
# ! wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v2.0.json -O squad/train-v2.0.json
# ! wget https://rajpurkar.github.io/SQuAD-explorer/dataset/dev-v2.0.json -O squad/dev-v2.0.json
# Each split is in a structured json file with a number of questions and answers for each passage (or context). We'll
# take this apart into parallel lists of contexts, questions, and answers (note that the contexts here are repeated since
# there are multiple questions per context):
# +
import json
from pathlib import Path
def read_squad(path):
path = Path(path)
with open(path, 'rb') as f:
squad_dict = json.load(f)
contexts = []
questions = []
answers = []
for group in squad_dict['data']:
for passage in group['paragraphs']:
context = passage['context']
for qa in passage['qas']:
question = qa['question']
for answer in qa['answers']:
contexts.append(context)
questions.append(question)
answers.append(answer)
return contexts, questions, answers
train_contexts, train_questions, train_answers = read_squad('squad/train-v2.0.json')
val_contexts, val_questions, val_answers = read_squad('squad/dev-v2.0.json')
# -
# The contexts and questions are just strings. The answers are dicts containing the subsequence of the passage with the
# correct answer as well as an integer indicating the character at which the answer begins. In order to train a model on
# this data we need (1) the tokenized context/question pairs, and (2) integers indicating at which **token** positions the
# answer begins and ends.
#
# First, let's get the **character** position at which the answer ends in the passage (we are given the starting position).
# Sometimes SQuAD answers are off by one or two characters, so we will also adjust for that.
# +
def add_end_idx(answers, contexts):
for answer, context in zip(answers, contexts):
gold_text = answer['text']
start_idx = answer['answer_start']
end_idx = start_idx + len(gold_text)
# sometimes squad answers are off by a character or two – fix this
if context[start_idx:end_idx] == gold_text:
answer['answer_end'] = end_idx
elif context[start_idx-1:end_idx-1] == gold_text:
answer['answer_start'] = start_idx - 1
answer['answer_end'] = end_idx - 1 # When the gold label is off by one character
elif context[start_idx-2:end_idx-2] == gold_text:
answer['answer_start'] = start_idx - 2
answer['answer_end'] = end_idx - 2 # When the gold label is off by two characters
add_end_idx(train_answers, train_contexts)
add_end_idx(val_answers, val_contexts)
# -
# Now `train_answers` and `val_answers` include the character end positions and the corrected start positions. Next,
# let's tokenize our context/question pairs. 🤗 Tokenizers can accept parallel lists of sequences and encode them together
# as sequence pairs.
# +
from transformers import DistilBertTokenizerFast
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-uncased')
train_encodings = tokenizer(train_contexts, train_questions, truncation=True, padding=True)
val_encodings = tokenizer(val_contexts, val_questions, truncation=True, padding=True)
# -
# Next we need to convert our character start/end positions to token start/end positions. When using 🤗 Fast Tokenizers,
# we can use the built in `BatchEncoding.char_to_token` method.
# +
def add_token_positions(encodings, answers):
start_positions = []
end_positions = []
for i in range(len(answers)):
start_positions.append(encodings.char_to_token(i, answers[i]['answer_start']))
end_positions.append(encodings.char_to_token(i, answers[i]['answer_end'] - 1))
# if None, the answer passage has been truncated
if start_positions[-1] is None:
start_positions[-1] = tokenizer.model_max_length
if end_positions[-1] is None:
end_positions[-1] = tokenizer.model_max_length
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
add_token_positions(train_encodings, train_answers)
add_token_positions(val_encodings, val_answers)
# -
# Our data is ready. Let's just put it in a PyTorch/TensorFlow dataset so that we can easily use it for training. In
# PyTorch, we define a custom `Dataset` class. In TensorFlow, we pass a tuple of `(inputs_dict, labels_dict)` to the
# `from_tensor_slices` method.
# +
import tensorflow as tf
train_dataset = tf.data.Dataset.from_tensor_slices((
{key: train_encodings[key] for key in ['input_ids', 'attention_mask']},
{key: train_encodings[key] for key in ['start_positions', 'end_positions']}
))
val_dataset = tf.data.Dataset.from_tensor_slices((
{key: val_encodings[key] for key in ['input_ids', 'attention_mask']},
{key: val_encodings[key] for key in ['start_positions', 'end_positions']}
))
# -
# Now we can use a DistilBert model with a QA head for training:
from transformers import TFDistilBertForQuestionAnswering
model = TFDistilBertForQuestionAnswering.from_pretrained("distilbert-base-uncased")
# The data and model are both ready to go. You can train the model with
# `Trainer`/`TFTrainer` exactly as in the sequence classification example
# above. If using native PyTorch, replace `labels` with `start_positions` and `end_positions` in the training
# example. If using Keras's `fit`, we need to make a minor modification to handle this example since it involves
# multiple model outputs.
#
# - [ft_trainer](#ft_trainer)
# +
# Keras will expect a tuple when dealing with labels
train_dataset = train_dataset.map(lambda x, y: (x, (y['start_positions'], y['end_positions'])))
# Keras will assign a separate loss for each output and add them together. So we'll just use the standard CE loss
# instead of using the built-in model.compute_loss, which expects a dict of outputs and averages the two terms.
# Note that this means the loss will be 2x of when using TFTrainer since we're adding instead of averaging them.
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
model.distilbert.return_dict = False # if using 🤗 Transformers >3.02, make sure outputs are tuples
optimizer = tf.keras.optimizers.Adam(learning_rate=5e-5)
model.compile(optimizer=optimizer, loss=loss) # can also use any keras loss fn
model.fit(train_dataset.shuffle(1000).batch(16), epochs=3, batch_size=16)
# -
# <a id='resources'></a>
# ## Additional Resources
# - [How to train a new language model from scratch using Transformers and Tokenizers](https://huggingface.co/blog/how-to-train). Blog post showing the steps to load in Esperanto data and train a
# masked language model from scratch.
# - [Preprocessing](https://huggingface.co/transformers/preprocessing.html). Docs page on data preprocessing.
# - [Training](https://huggingface.co/transformers/training.html). Docs page on training and fine-tuning.
# <a id='nlplib'></a>
# ### Using the 🤗 NLP Datasets & Metrics library
# This tutorial demonstrates how to read in datasets from various raw text formats and prepare them for training with 🤗
# Transformers so that you can do the same thing with your own custom datasets. However, we recommend users use the [🤗
# NLP library](https://github.com/huggingface/nlp) for working with the 150+ datasets included in the [hub](https://huggingface.co/datasets), including the three datasets used in this tutorial. As a very brief overview, we
# will show how to use the NLP library to download and prepare the IMDb dataset from the first example, [seq_imdb](#seq_imdb).
#
# Start by downloading the dataset:
from nlp import load_dataset
train = load_dataset("imdb", split="train")
# Each dataset has multiple columns corresponding to different features. Let's see what our columns are.
print(train.column_names)
# Great. Now let's tokenize the text. We can do this using the `map` method. We'll also rename the `label` column to
# `labels` to match the model's input arguments.
train = train.map(lambda batch: tokenizer(batch["text"], truncation=True, padding=True), batched=True)
train.rename_column_("label", "labels")
# Lastly, we can use the `set_format` method to determine which columns and in what data format we want to access
# dataset elements.
train.set_format("tensorflow", columns=["input_ids", "attention_mask", "labels"])
{key: val.shape for key, val in train[0].items()})
# We now have a fully-prepared dataset. Check out [the 🤗 NLP docs](https://huggingface.co/nlp/processing.html) for a
# more thorough introduction.
| transformers_doc/tensorflow/custom_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## Common plotting pitfalls that get worse with large data
#
# When working with large datasets, visualizations are often the only way available to understand the properties of that dataset -- there are simply too many data points to examine each one! Thus it is very important to be aware of some common plotting problems that are minor inconveniences with small datasets but very serious problems with larger ones.
#
# We'll cover:
#
# 1. [Overplotting](#1.-Overplotting)
# 2. [Oversaturation](#2.-Oversaturation)
# 3. [Undersampling](#3.-Undersampling)
# 4. [Undersaturation](#4.-Undersaturation)
# 5. [Underutilized range](#5.-Underutilized-range)
# 6. [Nonuniform colormapping](#6.-Nonuniform-colormapping)
#
# You can [skip to the end](#Summary) if you just want to see an illustration of these problems.
#
# This notebook requires [HoloViews](http://holoviews.org), [colorcet](https://github.com/bokeh/colorcet), and matplotlib, and optionally scikit-image, which can be installed with:
#
# ```
# conda install holoviews colorcet matplotlib scikit-image
# ```
#
# We'll first load the plotting libraries and set up some defaults:
# +
import numpy as np
np.random.seed(42)
import holoviews as hv
from holoviews.operation.datashader import datashade
from holoviews import opts, dim
hv.extension('matplotlib')
from colorcet import fire
datashade.cmap=fire[50:]
# -
opts.defaults(
opts.Image(cmap="gray_r", axiswise=True),
opts.Points(cmap="bwr", edgecolors='k', s=50, alpha=1.0), # Remove color_index=2
opts.RGB(bgcolor="black", show_grid=False),
opts.Scatter3D(color=dim('c'), fig_size=250, cmap='bwr', edgecolor='k', s=50, alpha=1.0)) #color_index=3
# ### 1. Overplotting
#
# Let's consider plotting some 2D data points that come from two separate categories, here plotted as blue and red in **A** and **B** below. When the two categories are overlaid, the appearance of the result can be very different depending on which one is plotted first:
# +
def blue_points(offset=0.5,pts=300):
blues = (np.random.normal( offset,size=pts), np.random.normal( offset,size=pts), -1 * np.ones((pts)))
return hv.Points(blues, vdims=['c']).opts(color=dim('c'))
def red_points(offset=0.5,pts=300):
reds = (np.random.normal(-offset,size=pts), np.random.normal(-offset,size=pts), 1*np.ones((pts)))
return hv.Points(reds, vdims=['c']).opts(color=dim('c'))
blues, reds = blue_points(), red_points()
blues + reds + (reds * blues) + (blues * reds)
# -
# Plots **C** and **D** shown the same distribution of points, yet they give a very different impression of which category is more common, which can lead to incorrect decisions based on this data. Of course, both are equally common in this case, so neither **C** nor **D** accurately reflects the data. The cause for this problem is simply occlusion:
hmap = hv.HoloMap({0:blues,0.000001:reds,1:blues,2:reds}, kdims=['level'])
hv.Scatter3D(hmap.table(), kdims=['x','y','level'], vdims=['c'])
# Occlusion of data by other data is called **overplotting** or **overdrawing**, and it occurs whenever a datapoint or curve is plotted on top of another datapoint or curve, obscuring it. It's thus a problem not just for scatterplots, as here, but for curve plots, 3D surface plots, 3D bar graphs, and any other plot type where data can be obscured.
#
#
# ### 2. Oversaturation
#
# You can reduce problems with overplotting by using transparency/opacity, via the alpha parameter provided to control opacity in most plotting programs. E.g. if alpha is 0.1, full color saturation will be achieved only when 10 points overlap, reducing the effects of plot ordering but making it harder to see individual points:
layout = blues + reds + (reds * blues) + (blues * reds)
layout.opts(opts.Points(s=50, alpha=0.1))
# Here **C **and **D **look very similar (as they should, since the distributions are identical), but there are still a few locations with **oversaturation**, a problem that will occur when more than 10 points overlap. In this example the oversaturated points are located near the middle of the plot, but the only way to know whether they are there would be to plot both versions and compare, or to examine the pixel values to see if any have reached full saturation (a necessary but not sufficient condition for oversaturation). Locations where saturation has been reached have problems similar to overplotting, because only the last 10 points plotted will affect the final color (for alpha of 0.1).
#
# Worse, even if one has set the alpha value to approximately or usually avoid oversaturation, as in the plot above, the correct value depends on the dataset. If there are more points overlapping in that particular region, a manually adjusted alpha setting that worked well for a previous dataset will systematically misrepresent the new dataset:
blues, reds = blue_points(pts=600), red_points(pts=600)
layout = blues + reds + (reds * blues) + (blues * reds)
layout.opts(opts.Points(alpha=0.1))
# Here **C **and **D **again look qualitatively different, yet still represent the same distributions. Since we're assuming that the point of the visualization is to reveal the underlying dataset, having to tune visualization parameters manually based on the properties of the dataset itself is a serious problem.
#
# To make it even more complicated, the correct alpha also depends on the dot size, because smaller dots have less overlap for the same dataset. With smaller dots, **C **and **D **look more similar, but the color of the dots is now difficult to see in all cases because the dots are too transparent for this size:
layout = blues + reds + (reds * blues) + (blues * reds)
layout.opts(opts.Points(s=10, alpha=0.1, edgecolor=None))
# As you can see, it is very difficult to find settings for the dotsize and alpha parameters that correctly reveal the data, even for relatively small and obvious datasets like these. With larger datasets with unknown contents, it is difficult to detect that such problems are occuring, leading to false conclusions based on inappropriately visualized data.
#
# ### 3. Undersampling
#
# With a single category instead of the multiple categories shown above, oversaturation simply obscures spatial differences in density. For instance, 10, 20, and 2000 single-category points overlapping will all look the same visually, for alpha=0.1. Let's again consider an example that has a sum of two normal distributions slightly offset from one another, but no longer using color to separate them into categories:
# +
def gaussians(specs=[(1.5,0,1.0),(-1.5,0,1.0)],num=100):
"""
A concatenated list of points taken from 2D Gaussian distributions.
Each distribution is specified as a tuple (x,y,s), where x,y is the mean
and s is the standard deviation. Defaults to two horizontally
offset unit-mean Gaussians.
"""
np.random.seed(1)
dists = [(np.random.normal(x,s,num), np.random.normal(y,s,num)) for x,y,s in specs]
return np.hstack([d[0] for d in dists]), np.hstack([d[1] for d in dists])
points = (hv.Points(gaussians(num=600), label="600 points", group="Small dots") +
hv.Points(gaussians(num=60000), label="60000 points", group="Small dots") +
hv.Points(gaussians(num=600), label="600 points", group="Tiny dots") +
hv.Points(gaussians(num=60000), label="60000 points", group="Tiny dots"))
points.opts(
opts.Points('Small_dots', s=1, alpha=1),
opts.Points('Tiny_dots', s=0.1, alpha=0.1))
# -
# Just as shown for the multiple-category case above, finding settings to avoid overplotting and oversaturation is difficult. The "Small dots" setting (size 0.1, full alpha) works fairly well for a sample of 600 points **A,** but it has serious overplotting issues for larger datasets, obscuring the shape and density of the distribution **B.** Using the "Tiny dots" setting (10 times smaller dots, alpha 0.1) works well for the larger dataset **D,** but not at all for the 600-point dataset **C.** Clearly, not all of these settings are accurately conveying the underlying distribution, as they all appear quite different from one another. Similar problems occur for the same size of dataset, but with greater or lesser levels of overlap between points, which of course varies with every new dataset.
#
# In any case, as dataset size increases, at some point plotting a full scatterplot like any of these will become impractical with current plotting software. At this point, people often simply subsample their dataset, plotting 10,000 or perhaps 100,000 randomly selected datapoints. But as panel **A **shows, the shape of an **undersampled** distribution can be very difficult or impossible to make out, leading to incorrect conclusions about the distribution. Such problems can occur even when taking very large numbers of samples, if examining sparsely populated regions of the space, which will approximate panel **A **for some plot settings and panel **C **for others. The actual shape of the distribution is only visible if sufficient datapoints are available in that region *and* appropriate plot settings are used, as in **D,** but ensuring that both conditions are true is a quite difficult process of trial and error, making it very likely that important features of the dataset will be missed.
#
# To avoid undersampling large datasets, researchers often use 2D histograms visualized as heatmaps, rather than scatterplots showing individual points. A heatmap has a fixed-size grid regardless of the dataset size, so that they can make use of all the data. Heatmaps effectively approximate a probability density function over the specified space, with coarser heatmaps averaging out noise or irrelevant variations to reveal an underlying distribution, and finer heatmaps able to represent more details in the distribution.
#
# Let's look at some heatmaps with different numbers of bins for the same two-Gaussians distribution:
# +
def heatmap(coords,bins=10,offset=0.0,transform=lambda d,m:d, label=None):
"""
Given a set of coordinates, bins them into a 2d histogram grid
of the specified size, and optionally transforms the counts
and/or compresses them into a visible range starting at a
specified offset between 0 and 1.0.
"""
hist,xs,ys = np.histogram2d(coords[0], coords[1], bins=bins)
counts = hist[:,::-1].T
transformed = transform(counts,counts!=0)
span = transformed.max()-transformed.min()
compressed = np.where(counts!=0,offset+(1.0-offset)*transformed/span,0)
args = dict(label=label) if label else {}
return hv.Image(compressed,bounds=(xs[-1],ys[-1],xs[1],ys[1]),**args)
hv.Layout([heatmap(gaussians(num=60000),bins) for bins in [8,20,200]])
# -
# As you can see, a too-coarse binning grid **A **cannot represent this distribution faithfully, but with enough bins **C,** the heatmap will approximate a tiny-dot scatterplot like plot **D **in the previous figure. For intermediate grid sizes **B **the heatmap can average out the effects of undersampling; **B **is actually a more faithful representation of the *distribution* than **C **is (which we know is two offset 2D Gaussians), while **C **more faithfully represents the *sampling* (i.e., the individual points drawn from this distribution). Thus choosing a good binning grid size for a heatmap does take some expertise and knowledge of the goals of the visualization, and it's always useful to look at multiple binning-grid spacings for comparison. Still, at least the binning parameter is something meaningful at the data level (how coarse a view of the data is desired?) rather than just a plotting detail (what size and transparency should I use for the points?) that must be determined arbitrarily.
#
# In any case, at least in principle, the heatmap approach can entirely avoid the first three problems above: **overplotting** (since multiple data points sum arithmetically into the grid cell, without obscuring one another), **oversaturation** (because the minimum and maximum counts observed can automatically be mapped to the two ends of a visible color range), and **undersampling** (since the resulting plot size is independent of the number of data points, allowing it to use an unbounded amount of incoming data).
#
#
#
# ### 4. Undersaturation
#
# Of course, heatmaps come with their own plotting pitfalls. One rarely appreciated issue common to both heatmaps and alpha-based scatterplots is **undersaturation**, where large numbers of data points can be missed entirely because they are spread over many different heatmap bins or many nearly transparent scatter points. To look at this problem, let's again consider a set of multiple 2D Gaussians, but this time with different amounts of spread (standard deviation):
dist = gaussians(specs=[(2,2,0.02), (2,-2,0.1), (-2,-2,0.5), (-2,2,1.0), (0,0,3)],num=10000)
hv.Points(dist) + hv.Points(dist).opts(s=0.1) + hv.Points(dist).opts(s=0.01, alpha=0.05)
# Plots **A,** **B,** and **C **are all scatterplots for the same data, which is a sum of 5 Gaussian distributions at different locations and with different standard deviations:
#
# 1. Location (2,2): very narrow spread
# 2. Location (2,-2): narrow spread
# 3. Location (-2,-2): medium spread
# 4. Location (-2,2): large spread
# 5. Location (0,0): very large spread
#
# In plot **A,** of course, the very large spread covers up everything else, completely obscuring the structure of this dataset by overplotting. Plots **B **and **C **reveal the structure better, but they required hand tuning and neither one is particularly satisfactory. In **B **there are four clearly visible Gaussians, but all but the largest appear to have the same density of points per pixel, which we know is not the case from how the dataset was constructed, and the smallest is nearly invisible. Each of the five Gaussians has the same number of data points (10000), but the second-largest looks like it has more than the others, and the narrowest one is likely to be overlooked altogether, which is thus a clear example of oversaturation obscuring important features. Yet if we try to combat the oversaturation by using transparency in **C,** we now get a clear problem with **undersaturation** -- the "very large spread" Gaussian is now essentially invisible. Again, there are just as many datapoints in that category, but we'd never even know they were there if only looking at **C.**
#
# Similar problems occur for a heatmap view of the same data:
hv.Layout([heatmap(dist,bins) for bins in [8,20,200]])
# Here the narrow-spread distributions lead to pixels with a very high count, and if the other pixels are linearly ramped into the available color range, from zero to that high count value, then the wider-spread values are obscured (as in **B **) or entirely invisible (as in **C **).
#
# To avoid undersaturation, you can add an offset to ensure that low-count (but nonzero) bins are mapped into a visible color, with the remaining intensity scale used to indicate differences in counts:
hv.Layout([heatmap(dist,bins,offset=0.2) for bins in [8,20,200]]).cols(4)
# Such mapping entirely avoids undersaturation, since all pixels are either clearly zero (in the background color, i.e. white in this case), or a non-background color taken from the colormap. The widest-spread Gaussian is now clearly visible in all cases.
#
# However, the actual structure (5 Gaussians of different spreads) is still not visible. In **A **the problem is clearly too-coarse binning, but in **B **the binning is also somewhat too coarse for this data, since the "very narrow spread" and "narrow spread" Gaussians show up identically, each mapping entirely into a single bin (the two black pixels). **C **shouldn't suffer from too-coarse binning, yet it still looks more like a plot of the "very large spread" distribution alone, than a plot of these five distributions of different spreads, and it is thus still highly misleading despite the correction for undersaturation.
#
#
# ### 5. Underutilized range
#
# So, what is the problem in plot **C **above? By construction, we've avoided the first four pitfalls: **overplotting**, **oversaturation**, **undersampling**, and **undersaturation**. But the problem is now more subtle: differences in datapoint density are not visible between the five Gaussians, because all or nearly all pixels end up being mapped into either the bottom end of the visible range (light gray), or the top end (black, used only for the single pixel holding the "very narrow spread" distribution). The entire rest of the visible colors in this gray colormap are unused, conveying no information to the viewer about the rich structure that we know this distribution contains. If the data were uniformly distributed over the range from minimum to maximum counts per pixel (0 to 10,000, in this case), then the above plot would work well, but that's not the case for this dataset or for most real-world datasets.
#
# So, let's try transforming the data from its default linear representation (integer count values) into something that preserves relative differences in count values but maps them into visually distinct colors. A logarithmic transformation is one common choice:
hv.Layout([heatmap(dist,bins,offset=0.2,transform=lambda d,m: np.where(m,np.log1p(d),0)) for bins in [8,20,200]])
# Aha! We can now see the full structure of the dataset, with all five Gaussians clearly visible in **B **and **C,** and the relative spreads also clearly visible in **C.**
#
# We still have a problem, though. The choice of a logarithmic transform was fairly arbitrary, and it mainly works well because we happened to have used an approximately geometric progression of spread sizes when constructing the example. For large datasets with truly unknown structure, can we have a more principled approach to mapping the dataset values into a visible range?
#
# Yes, if we think of the visualization problem in a different way. The underlying difficulty in plotting this dataset (as for very many real-world datasets) is that the values in each bin are numerically very different (ranging from 10,000, in the bin for the "very narrow spread" Gaussian, to 1 (for single datapoints from the "very large spread" Gaussian)). Given the 256 gray levels available in a normal monitor (and the similarly limited human ability to detect differences in gray values), numerically mapping the data values into the visible range is not going to work well. But given that we are already backing off from a direct numerical mapping in the above approaches for correcting undersaturation and for doing log transformations, what if we entirely abandon the numerical mapping approach, using the numbers only to form a partial ordering of the data values? Such an approach would be a rank-order plot, preserving order and not magnitudes. For 100 gray values, you can think of it as a percentile-based plot, with the lowest 1% of the data values mapping to the first visible gray value, the next 1% mapping to the next visible gray value, and so on to the top 1% of the data values mapping to the gray value 255 (black in this case). The actual data values would be ignored in such plots, but their relative magnitudes would still determine how they map onto colors on the screen, preserving the structure of the distribution rather than the numerical values.
#
# We can approximate such a rank-order or percentile encoding using the histogram equalization function from an image-processing package, which makes sure that each gray level is used for about the same number of pixels in the plot:
# +
try:
from skimage.exposure import equalize_hist
eq_hist = lambda d,m: equalize_hist(1000*d,nbins=100000,mask=m)
except ImportError:
eq_hist = lambda d,m: d
print("scikit-image not installed; skipping histogram equalization")
hv.Layout([heatmap(dist,bins,transform=eq_hist) for bins in [8,20,200]])
# -
# Plot **C** now reveals the full structure that we know was in this dataset, i.e. five Gaussians with different spreads, with no arbitrary parameter choices. (Well, there is a "number of bins" parameter for building the histogram for equalizing, but for integer data like this even that parameter can be eliminated entirely.) The differences in counts between pixels are now very clearly visible, across the full (and very wide) range of counts in the original data.
#
# Of course, we've lost the actual counts themselves, and so we can no longer tell just how many datapoints are in the "very narrow spread" pixel in this case. So plot **C** is accurately conveying the structure, but additional information would need to be provided to show the actual counts, by adding a color key mapping from the visible gray values into the actual counts and/or by providing hovering value information.
#
# At this point, one could also consider explicitly highlighting hotspots so that they cannot be overlooked. In plots B and C above, the two highest-density pixels are mapped to the two darkest pixel colors, which can reveal problems with your monitor settings if they were adjusted to make dark text appear blacker. Thus on those monitors, the highest values may not be clearly distinguishable from each other or from nearby grey values, which is a possible downside to fully utilizing the dynamic range available. But once the data is reliably and automatically mapped into a repeatable, reliable, fully utilized range for display, making explicit adjustments (e.g. based on wanting to make hotspots particularly clear) can be done in a principled way that doesn't depend on the actual data distribution (e.g. by just making the top few pixel values into a different color, or by stretching out those portions of the color map to show the extremes more safely across different monitors). Before getting into such specialized manipulations, there's a big pitfall to avoid first:
#
# ### 6. Nonuniform colormapping
#
# Let's say you've managed avoid pitfalls 1-5 somehow. However, there is one more problem waiting to catch you at the last stage, ruining all of your work eliminating the other issues: using a perceptually non-uniform colormap. A heatmap requires a colormap before it can be visualized, i.e., a lookup table from a data value (typically a normalized magnitude in the range 0 to 1) to a pixel color. The goal of a scientific visualization is to reveal the underlying properties of the data to your visual system, and to do so it is necessary to choose colors for each pixel that lead the viewer to perceive that data faithfully. Unfortunately, most of the colormaps in common use in plotting programs are highly *non*uniform.
#
# For instance, in "jet" (the default colormap for matlab and matplotlib until 2015), a large range of data values will all appear in shades of green that are perceptually indistinguishable, and similarly for the yellow regions of their "hot" colormaps:
#
# 
#
# In this image, a good colormap would have "teeth" equally visible at all data values, as for the perceptually uniform equivalents from the [colorcet](https://github.com/bokeh/colorcet) package:
#
# 
#
# We can easily see these effects if we look at our example dataset after histogram equalization, where all the different data levels are known to be distributed evenly in the array of normalized magnitudes:
hv.Layout([heatmap(dist,200,transform=eq_hist,label=cmap).opts(cmap=cmap) for cmap in ["hot","fire"]]).cols(2)
# Comparing **A ** to **B **it should be clear that the "fire" colormap is revealing much more of the data, accurately rendering the density differences between each of the different blobs. The unsuitable "hot" colormap is mapping all of the high density regions to perceptually indistinguishable shades of bright yellow/white, giving an "oversaturated" appearance even though we know the underlying heatmap array is *not* oversaturated (by construction). Luckily it is easy to avoid this problem; just use one of the 50 perceptually uniform colormaps available in the [colorcet](https://github.com/bokeh/colorcet) package, one of the four shipped with matplotlib [(viridis, plasma, inferno, or magma)](https://bids.github.io/colormap), or the Parula colormap shipped with Matlab.
#
#
# ## Summary
#
# Starting with plots of specific datapoints, we showed how typical visualization techniques will systematically misrepresent the distribution of those points. Here's an example of each of those six problems, all for the same distribution:
# +
layout = (hv.Points(dist,label="1. Overplotting") +
hv.Points(dist,label="2. Oversaturation").opts(s=0.1,alpha=0.5) +
hv.Points((dist[0][::200],dist[1][::200]),label="3. Undersampling").opts(s=2,alpha=0.5) +
hv.Points(dist,label="4. Undersaturation").opts(s=0.01,alpha=0.05) +
heatmap(dist,200,offset=0.2,label="5. Underutilized dynamic range") +
heatmap(dist,200,transform=eq_hist,label="6. Nonuniform colormapping").opts(cmap="hot"))
layout.opts(
opts.Points(axiswise=False),
opts.Layout(sublabel_format="", tight=True)).cols(3)
# -
# Here we could avoid each of these problems by hand, using trial and error based on our knowledge about the underlying dataset, since we created it. But for big data in general, these issues are major problems, because you don't know what the data *should* look like. Thus:
#
# #### For big data, you don't know when the viz is lying
#
# I.e., visualization is supposed to help you explore and understand your data, but if your visualizations are systematically misrepresenting your data because of **overplotting**, **oversaturation**, **undersampling**, **undersaturation**, **underutilized range**, and **nonuniform colormapping**, then you won't be able to discover the real qualities of your data and will be unable to make the right decisions.
#
# Luckily, using the systematic approach outlined in this discussion, you can avoid *all* of these pitfalls, allowing you to render your data faithfully without requiring *any* "magic parameters" that depend on your dataset:
heatmap(dist,200,transform=eq_hist).opts(cmap="fire")
# ### [Datashader](https://github.com/bokeh/datashader)
#
# The steps above show how to avoid the six main plotting pitfalls by hand, but it can be awkward and relatively slow to do so. Luckily there is a new Python library available to automate and optimize these steps, named [Datashader](https://github.com/bokeh/datashader). Datashader avoids users having to make dataset-dependent decisions and parameter settings when visualizing a new dataset. Datashader makes it practical to create accurate visualizations of datasets too large to understand directly, up to a billion points on a normal laptop and larger datasets on a compute cluster. As a simple teaser, the above steps can be expressed very concisely using the Datashader interface provided by [HoloViews](http://holoviews.org):
hv.output(size=200)
datashade(hv.Points(dist))
# Without any change to the settings, the same command will work with dataset sizes too large for most plotting programs, like this 50-million-point version of the distribution:
dist = gaussians(specs=[(2,2,0.02), (2,-2,0.1), (-2,-2,0.5), (-2,2,1.0), (0,0,3)], num=10000000)
datashade(hv.Points(dist))
# See the [Datashader web site](https://raw.githubusercontent.com/bokeh/datashader/master/examples/README.md) for details and examples to help you get started.
| examples/user_guide/1_Plotting_Pitfalls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
import wrangle
# -
flood = wrangle.clean_flood()
flood.sample(1)
flood.info()
# +
#flood.sensor_to_ground_feet.replace(to_replace=-999, value=13.500656)
# -
flood_train, flood_validate, flood_test = wrangle.split_data(flood)
flood.sensor_to_ground_feet.value_counts()
flood.sensor_to_ground_meters.value_counts()
# ___
# # Explore Flood
# ____
# def flood_alert(c):
# if 0 < c['flood_depth_meters'] < 10:
# return 'No Risk'
# elif 10 < c['flood_depth_meters'] < 11:
# return 'Minor Risk'
# elif 11 < c['flood_depth_meters'] < 12:
# return 'Moderate Risk'
# elif 12 < c['flood_depth_meters']:
# return 'Major Risk Risk'
# else:
# return 'No Alert'
# flood['flood_alert'] = flood.apply(flood_alert, axis=1)
flood.head()
flood.flood_alert.value_counts()
| workbooks/caitlyn/down_explore_flood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scalability
#
# The solution and estimation of finite-horizon discrete choice dynamic programming model appears straightforward. However, it entails a considerable computational burden due to the well known curse of dimensionality (Bellman and Dreyfus, 1962). The figure below illustrates how the total number of states increases exponentially with each period. The size of the state space is shown for Keane and Wolpin (1994) (all models have the same state space) and the base and extended model of Keane and Wolpin (1997). The latter two models are different because the state space of the base parameterization does not include information on the previous activity which significantly reduces the complexity of the model. Note that the y-axis us log-scaled to compensate for the loss in readibility due to exponential growth.
# + nbsphinx="hidden"
# %matplotlib agg
import matplotlib.pyplot as plt
import pandas as pd
import respy as rp
# + nbsphinx="hidden"
plt.style.use("docs/_static/respy.mplstyle")
# + nbsphinx="hidden"
def _get_states_per_period(model):
params, options, _ = rp.get_example_model(model)
# This is not the official API, but we do only need the state space.
state_space = rp.state_space.StateSpace(params, options)
states_per_period = [
len(range(slice_.start, slice_.stop))
for slice_ in state_space.slices_by_periods
]
return states_per_period
# + nbsphinx="hidden"
models = ["kw_94_one", "kw_97_base", "kw_97_extended"]
container = {}
for model in models:
container[model] = _get_states_per_period(model)
# + nbsphinx="hidden"
labels = [
"Keane and Wolpin (1994)",
"Keane and Wolpin (1997) - Base",
"Keane and Wolpin (1997) - Extended"
]
fig, ax = plt.subplots()
for label, states_per_period in zip(labels, container.values()):
n_states = sum(states_per_period)
ax.semilogy(range(0, len(states_per_period)), states_per_period, label=label)
ax.set_ylim(1e0, 1e7)
ax.grid(which="both", axis="y")
ax.set_xlabel("Period")
ax.set_ylabel("Number of states")
ax.legend();
# -
fig
# In total there are 317,367 states in Keane and Wolpin (1994), 12,991,208 states in Keane and Wolpin (1997) - Base and 59,306,140 states in Keane and Wolpin (1997) - Extended.
#
# During an estimation, thousands of different candidate parameterizations of the model are appraised with respect to the sample likelihood. For each evaluation of the likelihood the $n$-dimensional integral of $E\max$ (where $n$ is the number of choices) needs to be approximated at all states. Below, we show the total computation time required for 1,000 evaluations of the criterion function as we increase the number of threads from two to twelve.
# + nbsphinx="hidden"
data = """
{"model": "kw_94_one", "maxfun": 50, "n_threads": 2, "start": "2019-08-26 17:38:29.779205", "end": "2019-08-26 17:39:50.248813", "duration": "0:01:20.469608"}
{"model": "kw_94_one", "maxfun": 50, "n_threads": 4, "start": "2019-08-26 17:40:12.377112", "end": "2019-08-26 17:41:07.376235", "duration": "0:00:54.999123"}
{"model": "kw_94_one", "maxfun": 50, "n_threads": 6, "start": "2019-08-26 17:41:29.312806", "end": "2019-08-26 17:42:20.699365", "duration": "0:00:51.386559"}
{"model": "kw_94_one", "maxfun": 50, "n_threads": 8, "start": "2019-08-26 17:42:42.730035", "end": "2019-08-26 17:43:29.568749", "duration": "0:00:46.838714"}
{"model": "kw_94_one", "maxfun": 50, "n_threads": 10, "start": "2019-08-26 17:43:52.496852", "end": "2019-08-26 17:44:39.377107", "duration": "0:00:46.880255"}
{"model": "kw_94_one", "maxfun": 50, "n_threads": 12, "start": "2019-08-26 17:45:01.743654", "end": "2019-08-26 17:45:44.754300", "duration": "0:00:43.010646"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 2, "start": "2019-08-26 17:53:53.999918", "end": "2019-08-26 17:56:09.303005", "duration": "0:02:15.303087"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 4, "start": "2019-08-26 17:59:03.115069", "end": "2019-08-26 18:00:26.045647", "duration": "0:01:22.930578"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 6, "start": "2019-08-26 18:03:18.388000", "end": "2019-08-26 18:04:26.720110", "duration": "0:01:08.332110"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 8, "start": "2019-08-26 18:07:14.577545", "end": "2019-08-26 18:08:12.402948", "duration": "0:00:57.825403"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 10, "start": "2019-08-26 18:10:57.492809", "end": "2019-08-26 18:11:49.910569", "duration": "0:00:52.417760"}
{"model": "kw_97_base", "maxfun": 3, "n_threads": 12, "start": "2019-08-26 18:14:32.824227", "end": "2019-08-26 18:15:22.644604", "duration": "0:00:49.820377"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 2, "start": "2019-08-26 18:38:02.674008", "end": "2019-08-26 18:48:22.217274", "duration": "0:10:19.543266"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 4, "start": "2019-08-26 19:05:13.122038", "end": "2019-08-26 19:11:25.247955", "duration": "0:06:12.125917"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 6, "start": "2019-08-26 19:27:25.682803", "end": "2019-08-26 19:32:28.615169", "duration": "0:05:02.932366"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 8, "start": "2019-08-26 19:48:37.839972", "end": "2019-08-26 19:53:05.588380", "duration": "0:04:27.748408"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 10, "start": "2019-08-26 20:08:56.899445", "end": "2019-08-26 20:12:43.769137", "duration": "0:03:46.869692"}
{"model": "kw_97_extended", "maxfun": 3, "n_threads": 12, "start": "2019-08-26 20:27:47.747498", "end": "2019-08-26 20:31:26.824969", "duration": "0:03:39.077471"}
"""
# + nbsphinx="hidden"
df = pd.read_json(data, lines=True, convert_dates=["start", "end"])
# Get duration for 1000 evaluations.
df["duration"] = df.end - df.start
df["duration_1000_eval"] = 1000 / df["maxfun"] * df["duration"]
# + nbsphinx="hidden"
fig, ax = plt.subplots()
for label, group in df.groupby("model"):
ax.semilogy(group.n_threads, group.duration_1000_eval.dt.total_seconds() / 60, label=label)
ax.set_ylim(1e1, 1e4)
ax.grid(which="both", axis="y")
ax.set_xlabel("Number of threads")
ax.set_ylabel("Runtime (in Minutes)")
ax.legend([
"Keane and Wolpin (1994)",
"Keane and Wolpin (1997) - Base",
"Keane and Wolpin (1997) - Extended",
]);
# -
fig
# Adding even more threads, however, does not lead to any further improvements.
#
# For more details, see the script [online](https://github.com/OpenSourceEconomics/respy/blob/main/development/documentation/scalability/scalability_setup.py) or the complete Jupyter notebook in the repository under ``docs/software/scalability.ipynb``.
| docs/reference_guides/scalability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# # Understanding parameters of the first-level model
#
# In this tutorial, we study how first-level models are parametrized for fMRI
# data analysis and clarify the impact of these parameters on the results of the
# analysis.
#
# We use an exploratory approach, in which we incrementally include some new
# features in the analysis and inspect the outcome, i.e. the resulting brain
# maps.
#
# Readers without prior experience in fMRI data analysis should first run the
# `sphx_glr_auto_examples_plot_single_subject_single_run.py` tutorial to get
# a bit more familiar with the base concepts, and only then run this tutorial
# example.
#
# To run this example, you must launch IPython via ``ipython --matplotlib`` in a
# terminal, or use ``jupyter-notebook``.
# :depth: 1
#
# ## Retrieving the data
#
# We use a so-called localizer dataset, which consists in a 5-minutes
# acquisition of a fast event-related dataset.
#
#
#
from nilearn.datasets import func
data = func.fetch_localizer_first_level()
fmri_img = data.epi_img
# Define the paradigm that will be used. Here, we just need to get the provided
# file.
#
# This task, described in Pinel et al., BMC neuroscience 2007 probes basic
# functions, such as button presses with the left or right hand, viewing
# horizontal and vertical checkerboards, reading and listening to short
# sentences, and mental computations (subractions).
#
# Visual stimuli were displayed in four 250-ms epochs, separated by 100ms
# intervals (i.e., 1.3s in total). Auditory stimuli were drawn from a recorded
# male voice (i.e., a total of 1.6s for motor instructions, 1.2-1.7s for
# sentences, and 1.2-1.3s for subtractions). The auditory or visual stimuli
# were shown to the participants for passive listening or viewing or responses
# via button presses in event-related paradigms. Post-scan questions verified
# that the experimental tasks were understood and followed correctly.
#
# This task comprises 10 conditions:
#
# * audio_left_hand_button_press: Left-hand three-times button press, indicated by auditory instruction
# * audio_right_hand_button_press: Right-hand three-times button press, indicated by auditory instruction
# * visual_left_hand_button_press: Left-hand three-times button press, indicated by visual instruction
# * visual_right_hand_button_press: Right-hand three-times button press, indicated by visual instruction
# * horizontal_checkerboard: Visualization of flashing horizontal checkerboards
# * vertical_checkerboard: Visualization of flashing vertical checkerboards
# * sentence_listening: Listen to narrative sentences
# * sentence_reading: Read narrative sentences
# * audio_computation: Mental subtraction, indicated by auditory instruction
# * visual_computation: Mental subtraction, indicated by visual instruction
#
#
#
t_r = 2.4
events_file = data['events']
import pandas as pd
events = pd.read_table(events_file)
events
# ## Running a basic model
#
# First we specify a linear model.
# The .fit() functionality of FirstLevelModel function creates the design
# matrix and the beta maps.
#
#
#
from nilearn.glm.first_level import FirstLevelModel
first_level_model = FirstLevelModel(t_r)
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
# Let us take a look at the design matrix: it has 10 main columns corresponding
# to 10 experimental conditions, followed by 3 columns describing low-frequency
# signals (drifts) and a constant regressor.
#
#
from nilearn.plotting import plot_design_matrix
plot_design_matrix(design_matrix)
import matplotlib.pyplot as plt
plt.show()
# Specification of the contrasts.
#
# For this, let's create a function that, given the design matrix, generates
# the corresponding contrasts. This will be useful to repeat contrast
# specification when we change the design matrix.
#
#
# +
import numpy as np
def make_localizer_contrasts(design_matrix):
""" returns a dictionary of four contrasts, given the design matrix"""
# first generate canonical contrasts
contrast_matrix = np.eye(design_matrix.shape[1])
contrasts = dict([(column, contrast_matrix[i])
for i, column in enumerate(design_matrix.columns)])
contrasts['audio'] = (
contrasts['audio_left_hand_button_press']
+ contrasts['audio_right_hand_button_press']
+ contrasts['audio_computation']
+ contrasts['sentence_listening'])
# one contrast adding all conditions involving instructions reading
contrasts['visual'] = (
contrasts['visual_left_hand_button_press']
+ contrasts['visual_right_hand_button_press']
+ contrasts['visual_computation']
+ contrasts['sentence_reading'])
# one contrast adding all conditions involving computation
contrasts['computation'] = (contrasts['visual_computation']
+ contrasts['audio_computation'])
# one contrast adding all conditions involving sentences
contrasts['sentences'] = (contrasts['sentence_listening']
+ contrasts['sentence_reading'])
# Short dictionary of more relevant contrasts
contrasts = {
'left - right button press': (
contrasts['audio_left_hand_button_press']
- contrasts['audio_right_hand_button_press']
+ contrasts['visual_left_hand_button_press']
- contrasts['visual_right_hand_button_press']
),
'audio - visual': contrasts['audio'] - contrasts['visual'],
'computation - sentences': (contrasts['computation'] -
contrasts['sentences']
),
'horizontal-vertical': (contrasts['horizontal_checkerboard'] -
contrasts['vertical_checkerboard'])
}
return contrasts
# -
# Let's look at these computed contrasts:
#
# * 'left - right button press': probes motor activity in left versus right button presses
# * 'horizontal-vertical': probes the differential activity in viewing a horizontal vs vertical checkerboard
# * 'audio - visual': probes the difference of activity between listening to some content or reading the same type of content (instructions, stories)
# * 'computation - sentences': looks at the activity when performing a mental comptation task versus simply reading sentences.
#
#
#
# +
contrasts = make_localizer_contrasts(design_matrix)
from nilearn.plotting import plot_contrast_matrix
for key, values in contrasts.items():
plot_contrast_matrix(values, design_matrix=design_matrix)
plt.suptitle(key)
plt.show()
# -
# ## A first contrast estimation and plotting
#
# As this script will be repeated several times, we encapsulate model
# fitting and plotting in a function that we call when needed.
#
#
#
# +
from nilearn import plotting
def plot_contrast(first_level_model):
""" Given a first model, specify, estimate and plot the main contrasts"""
design_matrix = first_level_model.design_matrices_[0]
# Call the contrast specification within the function
contrasts = make_localizer_contrasts(design_matrix)
fig = plt.figure(figsize=(11, 3))
# compute the per-contrast z-map
for index, (contrast_id, contrast_val) in enumerate(contrasts.items()):
ax = plt.subplot(1, len(contrasts), 1 + index)
z_map = first_level_model.compute_contrast(
contrast_val, output_type='z_score')
plotting.plot_stat_map(
z_map, display_mode='z', threshold=3.0, title=contrast_id,
axes=ax, cut_coords=1)
# -
# Let's run the model and look at the outcome.
#
#
plot_contrast(first_level_model)
plt.show()
# ## Changing the drift model
#
# The drift model is a set of slow oscillating functions
# (Discrete Cosine transform) with a cut-off frequency. To remove
# spurious low-frequency effects related to heart rate, breathing and
# slow drifts in the scanner signal, the standard cutoff frequency
# is 1/128 Hz ~ 0.01Hz. This is the default value set in the FirstLevelModel
# function. Depending on the design of the experiment, the user may want to
# change this value. The cutoff period (1/high_pass) should be set as the
# longest period between two trials of the same condition multiplied by 2.
# For instance, if the longest period is 32s, the high_pass frequency shall be
# 1/64 Hz ~ 0.016 Hz. Note that the design matrix has more columns to model
# drifts in the data.
#
#
first_level_model = FirstLevelModel(t_r, high_pass=.016)
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
# Does the model perform worse or better ?
#
#
plot_contrast(first_level_model)
plt.show()
# We notice however that this model performs rather poorly.
#
# Another solution is to remove these drift terms. Maybe they're simply
# useless. This is done by setting drift_model to None.
#
#
first_level_model = FirstLevelModel(t_r, drift_model=None)
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Is it better than the original? No!
#
# Note that the design matrix has changed with no drift columns.
# the event columns, on the other hand, haven't changed.
#
# Another alternative to get a drift model is to specify a set of polynomials.
# Let's take a basis of 5 polynomials.
#
#
first_level_model = FirstLevelModel(t_r, drift_model='polynomial',
drift_order=5)
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Is it good? No better, no worse. Let's turn to another parameter.
#
#
# ## Changing the hemodynamic response model
#
# This is the filter used to convert the event sequence into a
# reference BOLD signal for the design matrix.
#
# The first thing that we can do is to change the default model (the
# so-called Glover hrf) for the so-called canonical model of SPM
# --which has a slightly weaker undershoot component.
#
#
first_level_model = FirstLevelModel(t_r, hrf_model='spm')
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# No strong --positive or negative-- effect.
#
#
# ### Adding a time derivative to the design
#
# There seems to be something here. Maybe we could try to go one step further:
# using not only the so-called canonical hrf, but also its time derivative.
# Note that in that case, we still perform the contrasts and obtain statistical
# significance for the main effect --- not the time derivative. This means that
# the inclusion of a time derivative in the design matrix has the sole effect
# of discounting timing misspecification from the error term, which would
# decrease the estimated variance and enhance the statistical significance of
# the effect. Is that the case?
#
#
first_level_model = FirstLevelModel(t_r, hrf_model='spm + derivative')
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Not a huge effect, but rather positive overall. We could keep that one.
#
# Note that a benefit of this approach is that we can test which voxels are
# well explained by the derivative term, hinting at misfit regions, a
# possibly valuable information. This is implemented by an F-test on
# the time derivative regressors.
#
#
# +
contrast_val = np.eye(design_matrix.shape[1])[1:21:2]
plot_contrast_matrix(contrast_val, design_matrix)
z_map = first_level_model.compute_contrast(
contrast_val, output_type='z_score')
plotting.plot_stat_map(
z_map, display_mode='z', threshold=3.0, title='effect of time derivatives')
plt.show()
# -
# There seems to be something here. Maybe we could adjust the
# timing, by increasing the slice_time_ref parameter from 0 to 0.5. Now the
# reference for model sampling is not the beginning of the volume
# acquisition, but the middle of it.
#
#
first_level_model = FirstLevelModel(t_r, hrf_model='spm + derivative',
slice_time_ref=0.5)
first_level_model = first_level_model.fit(fmri_img, events=events)
z_map = first_level_model.compute_contrast(
contrast_val, output_type='z_score')
plotting.plot_stat_map(
z_map, display_mode='z', threshold=3.0,
title='effect of time derivatives after model shift')
plt.show()
# The time derivatives regressors capture less signal: it's better like that.
#
#
# We can also consider adding the so-called dispersion derivative to
# capture some mis-specification in the shape of the hrf.
#
# This is done by specifying `hrf_model='spm + derivative + dispersion'`.
#
#
#
first_level_model = FirstLevelModel(t_r, slice_time_ref=0.5,
hrf_model='spm + derivative + dispersion')
first_level_model = first_level_model.fit(fmri_img, events=events)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Not a huge effect. For the sake of simplicity and readibility, we
# can drop that one.
#
#
# ## The noise model: ar(1), ols, or higher order ar?
#
# So far,we have implicitly used a lag-1 autoregressive model---aka
# ar(1)---for the temporal structure of the noise. An alternative
# choice is to use an ordinary least squares model (ols) that assumes
# no temporal structure (time-independent noise) or
# to use an autoregressive model with a higher order,
# for example a third order autoregressive model---aka ar(3).
#
# First we recompute using the `spm + derivative` hrf model, the
# slice_time_ref parameter chosen above, and explicitly set
# the noise model to be ar(1).
#
#
first_level_model = FirstLevelModel(t_r, slice_time_ref=0.5,
hrf_model='spm + derivative',
noise_model='ar1')
first_level_model = first_level_model.fit(fmri_img, events=events)
plot_contrast(first_level_model)
plt.show()
# Next we change the noise model to ols and observe the difference
# relative to the ar(1) model.
#
#
first_level_model = FirstLevelModel(t_r, slice_time_ref=0.5,
hrf_model='spm + derivative',
noise_model='ols')
first_level_model = first_level_model.fit(fmri_img, events=events)
plot_contrast(first_level_model)
plt.show()
# While the difference is not obvious you should rather stick to the
# ar(1) model, which is arguably more accurate.
#
# Alternatively we can include more terms in the autoregressive model to
# account for greater temporal complexity in the noise structure.
#
#
first_level_model = FirstLevelModel(t_r, slice_time_ref=0.5,
hrf_model='spm + derivative',
noise_model='ar3')
first_level_model = first_level_model.fit(fmri_img, events=events)
plot_contrast(first_level_model)
plt.show()
# This noise model arguably reduces the amount of spurious activity.
# However, as the difference is not obvious you may wish to stick to the
# ar(1) model, which is computationally more efficient.
#
#
# ## Removing confounds
#
# A problematic feature of fMRI is the presence of uncontrolled
# confounds in the data, due to scanner instabilities (spikes) or
# physiological phenomena, such as motion, heart and
# respiration-related blood oxygenation fluctuations. Side
# measurements are sometimes acquired to characterize these
# effects. Here we don't have access to those. What we can do instead
# is to estimate confounding effects from the data themselves, using
# the CompCor approach, and take those into account in the model.
#
# For this we rely on the so-called `high_variance_confounds`_
# routine of Nilearn.
#
#
#
from nilearn.image import high_variance_confounds
confounds = pd.DataFrame(high_variance_confounds(fmri_img, percentile=1))
first_level_model = FirstLevelModel(t_r, hrf_model='spm + derivative',
slice_time_ref=0.5)
first_level_model = first_level_model.fit(fmri_img, events=events,
confounds=confounds)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Note the five additional columns in the design matrix.
#
# The effect on the activation maps is complex: auditory/visual effects are
# killed, probably because they were somewhat colinear to the confounds. On the
# other hand, some of the maps become cleaner (horizontal-vertical,
# computation) after this addition.
#
#
# ## Smoothing
#
# Smoothing is a regularization of the model. It has two benefits:
# decrease the noise level in images, and reduce the discrepancy
# between individuals. The drawback is that it biases the shape and
# position of activation. Here, we simply illustrate the statistical
# gains. We use a mild smoothing of 5mm full-width at half maximum
# (fwhm).
#
#
first_level_model = FirstLevelModel(
t_r, hrf_model='spm + derivative', smoothing_fwhm=5,
slice_time_ref=0.5).fit(fmri_img, events=events, confounds=confounds)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# The design is unchanged but the maps are smoother and more contrasted.
#
#
#
# ## Masking
#
# Masking consists in selecting the region of the image on which the
# model is run: it is useless to run it outside of the brain.
#
# The approach taken by FirstLeveModel is to estimate it from the fMRI
# data itself when no mask is explicitly provided. Since the data
# have been resampled into MNI space, we can use instead a mask of the
# grey matter in MNI space. The benefit is that it makes voxel-level
# comparisons easier across subjects and datasets, and removes
# non-grey matter regions, in which no BOLD signal is expected. The
# downside is that the mask may not fit very well this particular
# data.
#
#
# +
data_mask = first_level_model.masker_.mask_img_
from nilearn.datasets import fetch_icbm152_brain_gm_mask
icbm_mask = fetch_icbm152_brain_gm_mask()
from nilearn.plotting import plot_roi
plt.figure(figsize=(16, 4))
ax = plt.subplot(121)
plot_roi(icbm_mask, title='ICBM mask', axes=ax)
ax = plt.subplot(122)
plot_roi(data_mask, title='Data-driven mask', axes=ax)
plt.show()
# -
# For the sake of time saving, we resample icbm_mask to our data.
# For this we call the resample_to_img routine of Nilearn.
# We use interpolation = 'nearest' to keep the mask as a binary image.
#
#
from nilearn.image import resample_to_img
resampled_icbm_mask = resample_to_img(icbm_mask, data_mask,
interpolation='nearest')
# Impact on the first-level model.
#
#
first_level_model = FirstLevelModel(
t_r, hrf_model='spm + derivative', smoothing_fwhm=5, slice_time_ref=0.5,
mask_img=resampled_icbm_mask).fit(
fmri_img, events=events, confounds=confounds)
design_matrix = first_level_model.design_matrices_[0]
plot_design_matrix(design_matrix)
plot_contrast(first_level_model)
plt.show()
# Note that it removed spurious spots in the white matter.
#
#
# ## Conclusion
#
# Interestingly, the model used here seems quite resilient to
# manipulation of modeling parameters: this is reassuring. It shows
# that Nilearn defaults ('cosine' drift, cutoff=128s, 'glover' hrf,
# ar(1) model) are actually reasonable. Note that these conclusions
# are specific to this dataset and may vary with other ones.
#
#
| plot_first_level_details.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Water usage
# +
# Import statements
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Use pandas to read the data located in the subfolder data.
# Load dataset
waterUsage = pd.read_csv('./data/water_usage.csv', index_col=0)
waterUsage = waterUsage.sort_values(by='Percentage')
# Use a pie chart to visualize the water usage. Highlight one usage of your choice using the explode parameter. Show the percentages for each slice and add a title.
# Create figure
plt.figure(figsize=(8, 8), dpi=300)
# Create pie plot
plt.pie('Percentage', labels='Usage', data=waterUsage,
colors=['#BFBFBF','#A6A6A6','#595959','#3D3D3D','#262626','#0D0D0D'])
# Add title
plt.title('Water Usage')
# Show plot
plt.show()
| Lesson03/Exercise04/exercise04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="d89dfe26" toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#extra-one-hot-features" data-toc-modified-id="extra-one-hot-features-0.1"><span class="toc-item-num">0.1 </span>extra one-hot features</a></span></li><li><span><a href="#extra-keywords" data-toc-modified-id="extra-keywords-0.2"><span class="toc-item-num">0.2 </span>extra keywords</a></span></li></ul></li><li><span><a href="#Dense" data-toc-modified-id="Dense-1"><span class="toc-item-num">1 </span>Dense</a></span></li></ul></div>
# + [markdown] id="05c43715"
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#TextCNN" data-toc-modified-id="TextCNN-1"><span class="toc-item-num">1 </span>TextCNN</a></span><ul class="toc-item"><li><span><a href="#notes:" data-toc-modified-id="notes:-1.1"><span class="toc-item-num">1.1 </span>notes:</a></span></li></ul></li><li><span><a href="#LSTM" data-toc-modified-id="LSTM-2"><span class="toc-item-num">2 </span>LSTM</a></span></li></ul></div>
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8023, "status": "ok", "timestamp": 1639116555162, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="85be328c" outputId="1e584967-0e3b-4708-e699-98b989f0a5a0"
from google.colab import drive
drive.mount('/content/drive')
import os
os.chdir("/content/drive/MyDrive/Text-Classification/code")
# !pip install pyLDAvis
# !pip install gensim
# !pip install pandas==1.3.0
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5024, "status": "ok", "timestamp": 1639116560177, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="7ea8c845" outputId="189aa339-bd8f-4d36-c4b2-e739773192cb"
from classification_utils import *
from clustering_utils import *
from eda_utils import *
from nn_utils_keras import *
from feature_engineering_utils import *
from data_utils import *
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 6659, "status": "ok", "timestamp": 1639116566826, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="f88921e6" outputId="43a0d66c-1544-4648-a52e-cf3838ee1781"
train, test = load_data(only_stem_voc=True)
# train, test = load_data(sample50=True)
# train, upsampling_info = upsampling_train(train)
train_text, train_label = train_augmentation(train, select_comb=None)
test_text, test_label = test['text'], test['label']
# test_text = test_text.apply(lambda x: extract_stem_voc(x))
# train_text = train_text.apply(lambda x: extract_stem_voc(x))
# train_text.to_csv("stem_voc_train.csv")
# test_text.to_csv("stem_voc_test.csv")
# train_text, test_text = load_stem_voc()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 12, "status": "ok", "timestamp": 1639116566827, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="9c089c98" outputId="1862e5a8-f2d3-4505-d76f-94fa9572f125"
####################################
### label mapper
####################################
labels = sorted(train_label.unique())
label_mapper = dict(zip(labels, range(len(labels))))
train_label = train_label.map(label_mapper)
test_label = test_label.map(label_mapper)
y_train = train_label
y_test = test_label
print(train_text.shape)
print(test_text.shape)
print(train_label.shape)
print(test_label.shape)
print(labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 4085, "status": "ok", "timestamp": 1639116570902, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="859a125b" outputId="8745f4ca-04bb-4665-ff5f-df4d0f91ac80"
####################################
### hyper params
####################################
filters = '"#$%&()*+,-/:;<=>@[\\]^_`{|}~\t\n0123465789!.?\''
MAX_NB_WORDS_ratio = 0.95
MAX_DOC_LEN_ratio = 0.99
MAX_NB_WORDS = eda_MAX_NB_WORDS(train_text, ratio=MAX_NB_WORDS_ratio, char_level=False, filters=filters)
MAX_DOC_LEN = eda_MAX_DOC_LEN(train_text, ratio=MAX_DOC_LEN_ratio, char_level=False, filters=filters)
# + executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1639116570903, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="ef858a91"
from tensorflow.keras import optimizers
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.layers import Embedding, Dense, Conv1D, MaxPooling1D, Dropout, Activation, Input, Flatten, Concatenate, Lambda
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.utils import to_categorical
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from tensorflow import keras
import numpy as np
import pandas as pd
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
import os
# + [markdown] id="fd1183b7"
# ## extra one-hot features
# + executionInfo={"elapsed": 10, "status": "ok", "timestamp": 1639116570903, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="b51f5083"
# one_hot_X_train, one_hot_X_test, one_hot_word_to_idx, one_hot_count_vect = count_vectorizer(
# train['Subject']+" " + train['Organization'], test['Subject']+" " + test['Organization'], stop_words=True, binary=False, min_df=3, max_df=0.001)
# + [markdown] id="269d56d2"
# ## extra keywords
# + executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1639116570904, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="a50842dc"
# label_docs = train.groupby('label')['text'].apply(lambda x: " ".join(x)) # 要去除标点符号
# dtm, _, label_word_to_idx, _ = count_vectorizer(label_docs, [''], stop_words=True, min_df=1, binary=True)
# label_idx_to_word = dict([val, key] for key, val in label_word_to_idx.items())
# keywords_threshold = 1
# keywords_idx = np.where(dtm.sum(axis=0)<=keywords_threshold)[0]
# print(" keywords_idx shape: ")
# voc = [label_idx_to_word[idx] for idx in keywords_idx]
# keywords_X_train, keywords_X_test, keywords_word_to_idx, keywords_count_vect = count_vectorizer(
# train['text'], test['text'], voc=voc, stop_words=True, min_df=1, binary=True)
# + [markdown] id="48bd5244"
# # Dense
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 199798, "status": "ok", "timestamp": 1639116770691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="7bda3ee7" outputId="9113c6c0-70ca-48ef-fd67-19a2e28874b2"
X_train, X_test, word_to_idx, tfidf_vect = tfidf_vectorizer(train_text, test_text, stop_words=True, binary=True, min_df=5, ngram_range=(2,3))
X_train, transform_mapper = dimension_reduction(X_train, out_dim=500) # not allow negative
X_test = transform_mapper.transform(X_test)
# + executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1639116770691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="6QMAH7LjxV6D"
# stop
# del X_train
# del X_test
# + [markdown] id="hWRCAMaD3S-u"
#
# + executionInfo={"elapsed": 374, "status": "ok", "timestamp": 1639116771057, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="b9a938ba"
_X_train = np.hstack([X_train])
_X_test = np.hstack([X_test])
# _X_train = np.hstack([X_train, keywords_X_train])
# _X_test = np.hstack([X_test, keywords_X_test])
# _X_train = np.hstack([X_train, one_hot_X_train, keywords_X_train])
# _X_test = np.hstack([X_test, one_hot_X_test, keywords_X_test])
# + executionInfo={"elapsed": 2640, "status": "ok", "timestamp": 1639116773696, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="e156bf89"
opt = optimizers.Adam(learning_rate=0.005)
model = Sequential()
model.add(Input(shape=_X_train.shape[1],)) # input cannot connect with dropout directly
# model.add(Dense(1024, activation='relu'))
# model.add(Dropout(0.5))
# model.add(Dense(256, activation='relu'))
# model.add(Dropout(0.5))
# model.add(Dense(128, activation='relu'))
# model.add(Dropout(0.5))
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(20, activation='softmax'))
model.compile(optimizer=opt,
loss=keras.losses.SparseCategoricalCrossentropy(),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
# model.compile(optimizer=opt,
# loss=keras.losses.CategoricalHinge(),
# metrics=[keras.metrics.CategoricalAccuracy()])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 65442, "status": "ok", "timestamp": 1639118329657, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="761b02dd" outputId="192c545d-e56a-4a1a-d94f-01fa294b4049"
BATCH_SIZE = 64 # 16 # 先在小的batch上train, 容易找到全局最优部分, 然后再到 大 batch 上train, 快速收敛到局部最优
NUM_EPOCHES = 100 # 20步以上
patience = 80
file_name = 'test'
BestModel_Name = file_name + 'Best_GS_6'
BEST_MODEL_FILEPATH = BestModel_Name
earlyStopping = EarlyStopping(monitor='val_sparse_categorical_accuracy', patience=patience, verbose=1, mode='max') # patience: number of epochs with no improvement on monitor : val_loss
checkpoint = ModelCheckpoint(BEST_MODEL_FILEPATH, monitor='val_sparse_categorical_accuracy', verbose=1, save_best_only=True, mode='max')
# history = model.fit(_X_train, y_train, validation_data=(_X_test,y_test), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
history = model.fit(_X_train, y_train, validation_data=(_X_test,y_test), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=None, verbose=1)
# history = model.fit(_X_train, to_categorical(y_train), validation_data=(_X_test,to_categorical(y_test)), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=None, verbose=1)
# history = model.fit(X_train, y_train, validation_data=(X_val, y_val), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 1131, "status": "ok", "timestamp": 1639118331229, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKFRGCPHBXOIreYY1GgMyIUPCGnToBiaki3_u3=s64", "userId": "18154280958911480081"}, "user_tz": 300} id="228d045a" outputId="b65bf720-f9bf-4c07-d87e-2f175c591e39"
#### classification Report
history_plot(history)
y_pred = model.predict(_X_test)
classification_report = evaluation_report(y_test, np.argmax(y_pred, axis=1), labels=labels)
scores = model.evaluate(_X_test, y_test, verbose=2)
roc_auc(y_test, y_pred)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
print( "\n\n\n")
# + [markdown] id="BaRiaoW6Ssng"
#
# + id="4ojr9MZ6I5ne"
# + id="5BhA5R7_I5qA"
| code/history_backup/NN_based_models_Deep_tfidf+onehot+Ngram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Property calculation with cubic EoS
#
# First it is necessary to import the ``component`` class and the equation of state (EoS). For these example we will use Peng-Robinson EoS (``preos``).
import numpy as np
from phasepy import component, preos
# ---
# ### Pure Fluids
#
# A fluid is created (``water``) and then the EoS object is created. The ``eosp`` object includes the methods to evaluate properties from the equation of state, such as, densities, pressure, fugacity coefficients, chemical potential and some thermal derived properties (residual entropy, residual enthalpy, residual heat capacities and speed of sound).
#
# **warning:** thermal derived properties are computed with numerical derivatives using $O(h^4)$ approximation.
# +
water = component(name = 'water', Tc = 647.13, Pc = 220.55, Zc = 0.229, Vc = 55.948, w = 0.344861,
GC = {'H2O':1}, Mw = 18.04)
eosp = preos(water)
# -
# The density of the fluid is computed with the ``eosp.density`` method. It requires the temperature, pressure and the aggregation state.
T = 340. # K
P = 1. # bar
# computed densities in mol/cm3
eosp.density(T, P, 'L'), eosp.density(T, P, 'V')
# Similarly the pressure of the fluid can be computed at given molar volume and temperature using the ``eosp.pressure`` method.
# +
rhol = 0.045697369422639834
vl = 1./rhol
rhov = 3.5769380018112745e-05
vv = 1./rhov
eosp.pressure(T, vl), eosp.pressure(T, vv)
# -
# For pure fluids, the ``eos.psat`` method allows to compute the saturation pressure at given temperature. It returns the equilibrium pressure and molar volumes of the liquid and vapor phase. Similarly, the ``eos.tsat`` method allows to compute the saturation temperarute at given pressure.
#
# The phase equilibria can be verified through fugacity coefficients using the ``eos.logfug`` method or by using chemical potentials with the ``eos.muad`` method. The chemical potentials require that dimensionless density and temperature.
# +
Psat, vlsat, vvsat = eosp.psat(T)
# checking fugacity coefficients
logfugl, vlsat = eosp.logfug(T, Psat, 'L')
logfugv, vvsat = eosp.logfug(T, Psat, 'V')
# checking chemical potentials
Tfactor, Pfactor, rofactor, tenfactor, zfactor = eosp.sgt_adim(T)
Tad = T*Tfactor
rholad = 1/vlsat * rofactor
rhovad = 1/vvsat * rofactor
mul = eosp.muad(rholad, Tad)
muv = eosp.muad(rhovad, Tad)
print('Fugacity coefficients:', np.allclose(logfugl, logfugv ))
print('Chemical potential:', np.allclose(mul, muv ))
# +
# computing saturation temperature
P = 1.01325 # bar
Tsat, vlsat2, vvsat2 = eosp.tsat(P, T0=350.)
# checking fugacity coefficients
logfugl, vlsat = eosp.logfug(Tsat, P, 'L')
logfugv, vvsat = eosp.logfug(Tsat, P, 'V')
# checking chemical potentials
Tfactor, Pfactor, rofactor, tenfactor, zfactor = eosp.sgt_adim(Tsat)
Tad2 = Tsat*Tfactor
rholad2 = 1/vlsat2 * rofactor
rhovad2 = 1/vvsat2 * rofactor
mul = eosp.muad(rholad2, Tad2)
muv = eosp.muad(rhovad2, Tad2)
print('Fugacity coefficients:', np.allclose(logfugl, logfugv ))
print('Chemical potential:', np.allclose(mul, muv ))
# -
# The ``eosp`` object also includes the calculation of some thermal derived properties such as residual entropy (``eosp.EntropyR``), residual enthalpy (``eosp.EnthalpyR``), residual isochoric heat capacity (``eosp.CvR``), , residual isobaric heat capacity (``eosp.CpR``).
#
# For the speed of sound calculation (``eosp.speed_sound``) the ideal gas heat capacities are required, in the example the isochoric and isobaric ideal gas contribution are set to $3R/2$ and $5R/2$, respectively. Better values of ideal gas heat capacities contribution can be found from empirical correlations, such as the provided by DIPPR 801.
# +
# vaporization entropy in J/mol K
Svap = eosp.EntropyR(T, Psat, 'V') - eosp.EntropyR(T, Psat, 'L')
# vaporization enthalpy in J/mol
Hvap = eosp.EnthalpyR(T, Psat, 'V') - eosp.EnthalpyR(T, Psat, 'L')
# isochoric and isobaric residual heats capacities in J / mol K
cvr = eosp.CvR(T, P, 'L')
cpr = eosp.CpR(T, P, 'L')
# ideal gas heat capacities, better values can be obtained with DIPPR 801 correlations
r = 8.314 # J / mol K
CvId = 3*r/2
CpId = 5*r/2
w = eosp.speed_sound(T, P, 'V', CvId=CvId, CpId=CpId)
print('Vaporization Entropy : ', Svap, 'J / mol K')
print('Vaporization Enthalpy : ', Hvap, 'J / mol')
print('Residual isochoric heat capacity : ', cvr, 'J / mol K')
print('Residual isobaric heat capacity : ', cpr, 'J / mol K')
print('Speed of sound : ', w, 'm / s')
# -
# ---
# ### Fluid mixtures
#
# The EoS can be used for mixtures. The mixture is first created from two pure fluids using the ``mixture`` class.
# In this example the mixture is modeled with the Peng-Robinson EoS using the MHV mixing rule and the UNIFAC activity coefficient model.
#
# The ``eos`` object includes the methods to evaluate properties from the equation of state, such as, densities, pressure, fugacity coefficients, chemical potential and some thermal derived properties (residual entropy, residual enthalpy, residual heat capacities and speed of sound).
#
# **again:** thermal derived properties are computed with numerical derivatives using $O(h^4)$ approximation.
# +
from phasepy import mixture
water = component(name = 'water', Tc = 647.13, Pc = 220.55, Zc = 0.229, Vc = 55.948, w = 0.344861,
GC = {'H2O':1}, Mw = 18.04)
ethanol = component(name = 'ethanol', Tc = 514.0, Pc = 61.37, Zc = 0.241, Vc = 168.0, w = 0.643558,
GC = {'CH3':1, 'CH2':1,'OH(P)':1}, Mw = 46.07)
mix = mixture(water, ethanol)
# or
mix = water + ethanol
mix.unifac()
eos = preos(mix, 'mhv_unifac')
# -
# The density of the fluid is computed with the ``eos.density`` method. It requires the composition, temperature, pressure and the aggregation state.
T = 340. # K
P = 1. # bar
x = np.array([0.3, 0.7])
eos.density(x, T, P, 'L'), eos.density(x, T, P, 'V')
# The pressure of the fluid mixture can be computed at given composition, molar volume and temperature using the ``eos.pressure`` method.
# +
rhol = eos.density(x, T, P, 'L')
vl = 1. / rhol
rhov = eos.density(x, T, P, 'V')
vv = 1. / rhov
eos.pressure(x, vl, T), eos.pressure(x, vv, T)
# -
# The effective fugacity coefficients can be computed at given composition, temperature, pressure and aggregation state using the ``eos.logfugef`` method. This functions returns the natural logarithm of the fugacity coefficients and the computed volume root.
#
# The fugacity of the mixture can be computed with the ``eos.logfugmix`` method. It works similarly as the previous method.
#
# The chemical potential are computed at given dimensionless density vector ($\rho_i = x_i \rho b_0$) and temperature.
#
# +
lnphi, v = eos.logfugef(x, T, P, 'L')
lnphimix, v = eos.logfugmix(x, T, P, 'L')
print('Partial molar property : ', np.allclose(lnphimix, np.dot(lnphi, x)))
rhofactor = eos.b[0]
rhoad = 1/v * rofactor
rhovector = x*rhoad
eos.muad(rhovector, T)
# -
# The ``eos`` object also includes the calculation of some thermal derived properties such as residual entropy (``eos.EntropyR``), residual enthalpy (``eos.EnthalpyR``), residual isochoric heat capacity (``eos.CvR``), residual isobaric heat capacity (``eos.CpR``).
#
# For the speed of sound calculation (``eos.speed_sound``) the ideal gas heat capacities are required, in the example, the isochoric and isobaric ideal gas contributions are set to $3R/2$ and $5R/2$, respectively. Better values of ideal gas heat capacities contribution can be found from empirical correlations, such as the provided by DIPPR 801.
# +
# Thermal derived properties
Sr = eos.EntropyR(x, T, P, 'L')
Hr = eos.EnthalpyR(x, T, P, 'L')
Cvr = eos.CvR(x, T, P, 'L')
Cpr = eos.CpR(x, T, P, 'L')
# ideal gas heat capacities, better values can be obtained with DIPPR 801 correlations
r = 8.314 # J / mol K
CvId = 3*r/2
CpId = 5*r/2
w = eos.speed_sound(x, T, P, 'V', CvId=CvId, CpId=CpId)
print('Residual Entropy : ', Sr, 'J / mol K')
print('Residual Enthalpy : ', Hr, 'J / mol')
print('Residual isochoric heat capacity : ', Cvr, 'J / mol K')
print('Residual isobaric heat capacity : ', Cpr, 'J / mol K')
print('Speed of sound : ', w, 'm / s')
# -
# ---
# For further information about each method check out the documentation running: ``eos.function?``
| examples/2. Property calculation from Cubic EoS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="TvpF7wM34qK4"
# # ORF307 Precept 9
#
# + [markdown] id="RV9lH1Ce49rm"
# ## Optimality Conditions Review
# primal and dual solutions are optimal if and only if the following are true:
#
#
# 1. $x$ is primal feasible
# 2. $y$ is dual feasible
# 3. duality gap is zero
#
#
# + [markdown] id="7tiTvyfL5bYS"
# ## Primal and Dual Simplex Takeaways
#
#
#
# * Primal simplex maintains primal feasibility and duality gap of zero, dual becomes feasible when converges
# * Dual simplex maintains dual feasibility and dualitgy gap of zero, primal becomes feasible when converges
#
#
# + [markdown] id="Fzilee-z52jQ"
# ## Sensitivity Analysis - Adding a New Variable
#
#
# 1. $(x^*,0)$ is still primal feasible
# 2. Dual problem has an extra constraint, is $A^T_{n+1}y^* + c_{n+1} \ge 0$ still dual feasible?
# 3. If yes, $(x^*,0)$ is still optimal; If no, run primal simplex.
#
#
# + [markdown] id="TPUQA6Xi5_TE"
# ## Sensitivity Analysis - Adding a New Constraint
#
# 1. Translates to having extra variable in dual problem, thus $(y^*,0)$ is still dual feasible
# 2. Is $a^T_{m+1} x^* = b_{m+1}$ still primal feasible?
# 3. If yes, $(y^*,0)$ is still optimal; If no, run dual simplex.
# + [markdown] id="kMnP6RzM6BuL"
# ## Global/Local Sensitivity Analysis - Changing the Data
# + [markdown] id="uyYE98Gn2NAL"
# ## Example 1
# + [markdown] id="herOAa7E2R5s"
# Consider the problem
#
# $$
# \begin{array}{ll}
# \mbox{minimize} &-5x_1 - x_2 + 12x_3\\
# \text{subject to} &3x_1 + 2x_2 + x_3 = 10\\
# &5x_1 + 3x_2 + x_4 = 16\\
# &x_1, \dots, x_4 \ge 0.
# \end{array}
# $$
#
# An optimal solution to this problem is given by $x=(2,2,0,0)$. Suppose we change $a_{11}$ from 3 to $3 + \delta$. Keeping $x_1$ and $x_2$ as the basic variables, and let $A_B(\delta)$ be the corresponding basis matrix, as a function of $\delta$.
#
# + [markdown] id="TfdR1xfF8IfQ"
# (a) Compute $A_B(\delta)^{-1}b$. For which values of $\delta$ is $A_B(\delta)$ a feasible basis?
#
# (b) Compute $c_B^TA_B(\delta)^{-1}$. For which values of $\delta$ is $A_B(\delta)$ an optimal basis?
# + [markdown] id="KX-V_cAJ6PmM"
# ## Network Flows - Arc-Node Incidence Matrix
#
# $m$ x $n$ matrix $A$ with entries
#
# $$
# \begin{align}
# A_{ij} = \begin{cases}
# 1 &\mbox{if arc $j$ starts at node $i$}\\
# -1 &\mbox{if arc $j$ starts at node $i$}\\
# 0 &\mbox{otherwise}
# \end{cases}
# \end{align}.
# $$
#
# Each column has one -1 and one 1
# + [markdown] id="DJ1AgY_X8HwA"
# ## Network Flows - Total Unimodularity
#
# * A matrix is totally unimodular if all its minors are -1, 0 or 1
# * Key property: the inverse of an nonsingular square submatrix of A has entires +1, -1, or 0
# * Above property, along with $b$ being an integer vector implies all the extreme points of the polyhedron $P = \{x \in \mathbf{R}^n \vert Ax=b, \mkern9mu x\ge0\}$ are integer vectors
# -
#
# + [markdown] id="UrCEJ2-N3nGp"
# ## Example 2
# + [markdown] id="oJE6TX4k3pYt"
# A catering company must provide to a client $r_i$ tablecloths on each of $N$ consecutive days. The catering company can buy new tablecloths at a price of $p$ dollars each, or launder the used ones. Laundering can be done at a fast service facility that makes the tablecloths unavailable for the next $n$ days and cost $f$ dollars per tablecloth, or at a slower facility that makes tablecloths unavailable for the next $m$ days (with $m > n$) at a cost of $g$ dollars per tablecloth ($g<f$). The caterer's problem is to decide how to meet the client's demand at minimum cost, starting with no tablecloths and under the assumption that any leftover tablecloths have no value. Show that the problem can be formulated as a network flow problem.
#
| precepts/09_precept/09_precept.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"} tags=[]
# # Grammar Coverage
#
# [Producing inputs from grammars](GrammarFuzzer.ipynb) gives all possible expansions of a rule the same likelihood. For producing a comprehensive test suite, however, it makes more sense to maximize _variety_ – for instance, by not repeating the same expansions over and over again. In this chapter, we explore how to systematically _cover_ elements of a grammar such that we maximize variety and do not miss out individual elements.
# -
from bookutils import YouTubeVideo
YouTubeVideo('yq1orQJF6ys')
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# **Prerequisites**
#
# * You should have read the [chapter on grammars](Grammars.ipynb).
# * You should have read the [chapter on efficient grammar fuzzing](GrammarFuzzer.ipynb).
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from fuzzingbook.GrammarCoverageFuzzer import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter introduces `GrammarCoverageFuzzer`, an efficient grammar fuzzer extending `GrammarFuzzer` from the [chapter on efficient grammar fuzzing](GrammarFuzzer.ipynb). It strives to _cover all expansions at least once,_ thus ensuring coverage of functionality.
#
# In the following example, for instance, we use `GrammarCoverageFuzzer` to produce an expression. We see that the resulting expression covers all digits and all operators in a single expression.
#
# ```python
# >>> from Grammars import EXPR_GRAMMAR
# >>> expr_fuzzer = GrammarCoverageFuzzer(EXPR_GRAMMAR)
# >>> expr_fuzzer.fuzz()
# ```
# After fuzzing, the `expansion_coverage()` method returns a mapping of grammar expansions covered.
#
# ```python
# >>> expr_fuzzer.expansion_coverage()
# ```
# Subsequent calls to `fuzz()` will go for further coverage (i.e., covering the other area code digits, for example); a call to `reset()` clears the recored coverage, starting anew.
#
# Since such coverage in inputs also yields higher code coverage, `GrammarCoverageFuzzer` is a recommended extension to `GrammarFuzzer`.
#
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Covering Grammar Elements
#
# The aim of test generation is to cover all functionality of a program – hopefully including the failing functionality, of course. This functionality, however, is tied to the _structure of the input_: If we fail to produce certain input elements, then the associated code and functionality will not be triggered either, nixing our chances to find a bug in there.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# As an example, consider our expression grammar `EXPR_GRAMMAR` from the [chapter on grammars.](Grammars.ipynb):
#
# * If we do not produce negative numbers, then negative numbers will not be tested.
# * If we do not produce floating-point numbers, then floating-point numbers will not be tested.
#
# Our aim must thus be to _cover all possible expansions_ – and not only by chance, but _by design_.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# One way to maximize such variety is to _track_ the expansions that occur during grammar production: If we already have seen some expansion, we can prefer other possible expansion candidates out of the set of possible expansions. Consider the following rule in our expression grammar:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# -
from bookutils import quiz
from Fuzzer import Fuzzer
from typing import Dict, List, Set, Union, Optional
# + slideshow={"slide_type": "skip"}
from Grammars import EXPR_GRAMMAR, CGI_GRAMMAR, URL_GRAMMAR, START_SYMBOL
from Grammars import is_valid_grammar, extend_grammar, Grammar
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
EXPR_GRAMMAR["<factor>"]
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# Let us assume we have already produced an `<integer>` in the first expansion of `<factor>`. As it comes to expand the next factor, we would mark the `<integer>` expansion as already covered, and choose one of the yet uncovered alternatives such as `-<factor>` (a negative number) or `<integer>.<integer>` (a floating-point number). Only when we have covered all alternatives would we go back and reconsider expansions covered before.
# -
quiz("Which expansions of `EXPR_GRAMMAR` does the expression `1 + 2` cover?",
[
"`<start> -> <expr>`",
"`<integer> -> <digit><integer>`",
"`<integer> -> <digit>`",
"`<factor> -> +<factor>`"
], [1, 3])
# Indeed! The expression has expansions from `<start>` and into individual digits.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# ### Tracking Grammar Coverage
#
# This concept of _grammar coverage_ is easy to implement. We introduce a class `TrackingGrammarCoverageFuzzer` that keeps track of the current grammar coverage achieved:
# + slideshow={"slide_type": "skip"}
from Grammars import Grammar, Expansion
from GrammarFuzzer import GrammarFuzzer, all_terminals, nonterminals, \
display_tree, DerivationTree
# + slideshow={"slide_type": "skip"}
import random
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class TrackingGrammarCoverageFuzzer(GrammarFuzzer):
"""Track grammar coverage during production"""
def __init__(self, *args, **kwargs) -> None:
# invoke superclass __init__(), passing all arguments
super().__init__(*args, **kwargs)
self.reset_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Keeping Track of Expansions
#
# In the set `covered_expansions`, we store individual expansions seen.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class TrackingGrammarCoverageFuzzer(TrackingGrammarCoverageFuzzer):
def expansion_coverage(self) -> Set[str]:
"""Return the set of covered expansions as strings SYMBOL -> EXPANSION"""
return self.covered_expansions
def reset_coverage(self) -> None:
"""Clear coverage info tracked so far"""
self.covered_expansions: Set[str] = set()
# + [markdown] slideshow={"slide_type": "subslide"}
# We save them the expansions as strings "_symbol_ -> _expansion_", using the function `expansion_key()` to generate a string representation for the (_symbol_, _expansion_) pair.
# + slideshow={"slide_type": "fragment"}
def expansion_key(symbol: str,
expansion: Union[Expansion,
DerivationTree,
List[DerivationTree]]) -> str:
"""Convert (symbol, `expansion`) into a key "SYMBOL -> EXPRESSION".
`expansion` can be an expansion string, a derivation tree,
or a list of derivation trees."""
if isinstance(expansion, tuple):
# Expansion or single derivation tree
expansion, _ = expansion
if not isinstance(expansion, str):
# Derivation tree
children = expansion
expansion = all_terminals((symbol, children))
assert isinstance(expansion, str)
return symbol + " -> " + expansion
# -
# Here's an example:
# + slideshow={"slide_type": "fragment"}
expansion_key(START_SYMBOL, EXPR_GRAMMAR[START_SYMBOL][0])
# + [markdown] slideshow={"slide_type": "subslide"}
# Instead of _expansion_, we can also pass a list of children as argument, which will then automatically be converted into a string.
# + slideshow={"slide_type": "fragment"}
children: List[DerivationTree] = [("<expr>", None), (" + ", []), ("<term>", None)]
expansion_key("<expr>", children)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Computing Possible Expansions
#
# We can compute the set of possible expansions in a grammar by enumerating all expansions. The method `max_expansion_coverage()` traverses the grammar recursively starting from the given symbol (by default: the grammar start symbol) and accumulates all expansions in the set `expansions`. With the `max_depth` parameter (default: $\infty$), we can control how deep the grammar exploration should go; we will need this later in the chapter.
# + slideshow={"slide_type": "subslide"}
class TrackingGrammarCoverageFuzzer(TrackingGrammarCoverageFuzzer):
def _max_expansion_coverage(self, symbol: str,
max_depth: Union[int, float]) -> Set[str]:
if max_depth <= 0:
return set()
self._symbols_seen.add(symbol)
expansions = set()
for expansion in self.grammar[symbol]:
expansions.add(expansion_key(symbol, expansion))
for nonterminal in nonterminals(expansion):
if nonterminal not in self._symbols_seen:
expansions |= self._max_expansion_coverage(
nonterminal, max_depth - 1)
return expansions
def max_expansion_coverage(self, symbol: Optional[str] = None,
max_depth: Union[int, float] = float('inf')) \
-> Set[str]:
"""Return set of all expansions in a grammar
starting with `symbol` (default: start symbol).
If `max_depth` is given, expand only to that depth."""
if symbol is None:
symbol = self.start_symbol
self._symbols_seen: Set[str] = set()
cov = self._max_expansion_coverage(symbol, max_depth)
if symbol == START_SYMBOL:
assert len(self._symbols_seen) == len(self.grammar)
return cov
# + [markdown] slideshow={"slide_type": "subslide"}
# We can use `max_expansion_coverage()` to compute all the expansions within the expression grammar:
# + slideshow={"slide_type": "fragment"}
expr_fuzzer = TrackingGrammarCoverageFuzzer(EXPR_GRAMMAR)
expr_fuzzer.max_expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Tracking Expansions while Fuzzing
#
# During expansion, we can keep track of expansions seen. To do so, we hook into the method `choose_node_expansion()`, expanding a single node in our [Grammar fuzzer](GrammarFuzzer.ipynb).
# + slideshow={"slide_type": "fragment"}
class TrackingGrammarCoverageFuzzer(TrackingGrammarCoverageFuzzer):
def add_coverage(self, symbol: str,
new_child: Union[Expansion, List[DerivationTree]]) -> None:
key = expansion_key(symbol, new_child)
if self.log and key not in self.covered_expansions:
print("Now covered:", key)
self.covered_expansions.add(key)
def choose_node_expansion(self, node: DerivationTree,
children_alternatives:
List[List[DerivationTree]]) -> int:
(symbol, children) = node
index = super().choose_node_expansion(node, children_alternatives)
self.add_coverage(symbol, children_alternatives[index])
return index
# + [markdown] slideshow={"slide_type": "fragment"}
# The method `missing_expansion_coverage()` is a helper method that returns the expansions that still have to be covered:
# + slideshow={"slide_type": "fragment"}
class TrackingGrammarCoverageFuzzer(TrackingGrammarCoverageFuzzer):
def missing_expansion_coverage(self) -> Set[str]:
"""Return expansions not covered yet"""
return self.max_expansion_coverage() - self.expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Putting Things Together
#
# Let us show how tracking works. To keep things simple, let us focus on `<digit>` expansions only.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
digit_fuzzer = TrackingGrammarCoverageFuzzer(
EXPR_GRAMMAR, start_symbol="<digit>", log=True)
digit_fuzzer.fuzz()
# + slideshow={"slide_type": "fragment"}
digit_fuzzer.fuzz()
# + slideshow={"slide_type": "subslide"}
digit_fuzzer.fuzz()
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the set of covered expansions so far:
# + slideshow={"slide_type": "fragment"}
digit_fuzzer.expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# This is the set of all expansions we can cover:
# + slideshow={"slide_type": "fragment"}
digit_fuzzer.max_expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# This is the missing coverage:
# + slideshow={"slide_type": "fragment"}
digit_fuzzer.missing_expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# On average, how many characters do we have to produce until all expansions are covered?
# + slideshow={"slide_type": "fragment"}
def average_length_until_full_coverage(fuzzer: TrackingGrammarCoverageFuzzer) -> float:
trials = 50
sum = 0
for trial in range(trials):
# print(trial, end=" ")
fuzzer.reset_coverage()
while len(fuzzer.missing_expansion_coverage()) > 0:
s = fuzzer.fuzz()
sum += len(s)
return sum / trials
# + slideshow={"slide_type": "fragment"}
digit_fuzzer.log = False
average_length_until_full_coverage(digit_fuzzer)
# + [markdown] slideshow={"slide_type": "subslide"}
# For full expressions, this takes a bit longer:
# + slideshow={"slide_type": "fragment"}
expr_fuzzer = TrackingGrammarCoverageFuzzer(EXPR_GRAMMAR)
average_length_until_full_coverage(expr_fuzzer)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Covering Grammar Expansions
#
# Let us now not only track coverage, but actually _produce_ coverage. The idea is as follows:
#
# 1. We determine children yet uncovered (in `uncovered_children`)
# 2. If all children are covered, we fall back to the original method (i.e., choosing one expansion randomly)
# 3. Otherwise, we select a child from the uncovered children and mark it as covered.
#
# To this end, we introduce a new fuzzer `SimpleGrammarCoverageFuzzer` that implements this strategy in the `choose_node_expansion()` method – the method [the `GrammarFuzzer` superclass uses to select the child to be expanded](GrammarFuzzer.ipynb).
# + slideshow={"slide_type": "subslide"}
class SimpleGrammarCoverageFuzzer(TrackingGrammarCoverageFuzzer):
"""When choosing expansions, prefer expansions not covered."""
def choose_node_expansion(self,
node: DerivationTree,
children_alternatives: List[List[DerivationTree]]) -> int:
"""Return index of expansion in `children_alternatives` to be selected.
Picks uncovered expansions, if any."""
# Prefer uncovered expansions
(symbol, children) = node
uncovered_children = [c for (i, c) in enumerate(children_alternatives)
if expansion_key(symbol, c)
not in self.covered_expansions]
index_map = [i for (i, c) in enumerate(children_alternatives)
if c in uncovered_children]
if len(uncovered_children) == 0:
# All expansions covered - use superclass method
return self.choose_covered_node_expansion(node, children_alternatives)
# Select from uncovered nodes
index = self.choose_uncovered_node_expansion(node, uncovered_children)
return index_map[index]
# + [markdown] slideshow={"slide_type": "subslide"}
# The two methods `choose_covered_node_expansion()` and `choose_uncovered_node_expansion()` are provided for subclasses to hook in:
# + slideshow={"slide_type": "fragment"}
class SimpleGrammarCoverageFuzzer(SimpleGrammarCoverageFuzzer):
def choose_uncovered_node_expansion(self,
node: DerivationTree,
children_alternatives: List[List[DerivationTree]]) \
-> int:
"""Return index of expansion in _uncovered_ `children_alternatives`
to be selected.
To be overloaded in subclasses."""
return TrackingGrammarCoverageFuzzer.choose_node_expansion(
self, node, children_alternatives)
def choose_covered_node_expansion(self,
node: DerivationTree,
children_alternatives: List[List[DerivationTree]]) \
-> int:
"""Return index of expansion in _covered_ `children_alternatives`
to be selected.
To be overloaded in subclasses."""
return TrackingGrammarCoverageFuzzer.choose_node_expansion(
self, node, children_alternatives)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# By returning the set of expansions covered so far, we can invoke the fuzzer multiple times, each time adding to the grammar coverage. Using the `EXPR_GRAMMAR` grammar to produce digits, for instance, the fuzzer produces one digit after the other:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
f = SimpleGrammarCoverageFuzzer(EXPR_GRAMMAR, start_symbol="<digit>")
f.fuzz()
# + slideshow={"slide_type": "fragment"}
f.fuzz()
# + slideshow={"slide_type": "fragment"}
f.fuzz()
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's the set of covered expansions so far:
# + slideshow={"slide_type": "fragment"}
f.expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us fuzz some more. We see that with each iteration, we cover another expansion:
# + slideshow={"slide_type": "fragment"}
for i in range(7):
print(f.fuzz(), end=" ")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# At the end, all expansions are covered:
# + slideshow={"slide_type": "fragment"}
f.missing_expansion_coverage()
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let us apply this on a more complex grammar – e.g., the full expression grammar. We see that after a few iterations, we cover each and every digit, operator, and expansion:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
f = SimpleGrammarCoverageFuzzer(EXPR_GRAMMAR)
for i in range(10):
print(f.fuzz())
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# Again, all expansions are covered:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
f.missing_expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# We see that our strategy is much more effective in achieving coverage than the random approach:
# + slideshow={"slide_type": "fragment"}
average_length_until_full_coverage(SimpleGrammarCoverageFuzzer(EXPR_GRAMMAR))
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=false
# ## Deep Foresight
#
# Selecting expansions for individual rules is a good start; however, it is not sufficient, as the following example shows. We apply our coverage fuzzer on the CGI grammar from the [chapter on grammars](Grammars.ipynb):
# + slideshow={"slide_type": "subslide"}
CGI_GRAMMAR
# + slideshow={"slide_type": "subslide"}
f = SimpleGrammarCoverageFuzzer(CGI_GRAMMAR)
for i in range(10):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# After 10 iterations, we still have a number of expansions uncovered:
# + slideshow={"slide_type": "fragment"}
f.missing_expansion_coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# Why is that so? The problem is that in the CGI grammar, the largest number of variations to be covered occurs in the `hexdigit` rule. However, we first need to _reach_ this expansion. When expanding a `<letter>` symbol, we have the choice between three possible expansions:
# + slideshow={"slide_type": "fragment"}
CGI_GRAMMAR["<letter>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# If all three expansions are covered already, then `choose_node_expansion()` above will choose one randomly – even if there may be more expansions to cover when choosing `<percent>`.
# + [markdown] slideshow={"slide_type": "subslide"}
# What we need is a better strategy that will pick `<percent>` if there are more uncovered expansions following – even if `<percent>` is covered. Such a strategy was first discussed by <NAME> \cite{Burkhardt1967} under the name of "Shortest Path Selection":
#
# > This version selects, from several alternatives for development, that syntactic unit under which there is still an unused unit available, starting with the shortest path.
#
# This is what we will implement in the next steps.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Determining Maximum per-Symbol Coverage
#
# To address this problem, we introduce a new class `GrammarCoverageFuzzer` that builds on `SimpleGrammarCoverageFuzzer`, but with a _better strategy_. First, we need to compute the _maximum set of expansions_ that can be reached from a particular symbol, as we already have implemented in `max_expansion_coverage()`. The idea is to later compute the _intersection_ of this set and the expansions already covered, such that we can favor those expansions with a non-empty intersection.
# + [markdown] slideshow={"slide_type": "fragment"}
# The first step – computing the maximum set of expansions that can be reached from a symbol – is already implemented. By passing a `symbol` parameter to `max_expansion_coverage()`, we can compute the possible expansions for every symbol:
# + slideshow={"slide_type": "subslide"}
f = SimpleGrammarCoverageFuzzer(EXPR_GRAMMAR)
f.max_expansion_coverage('<integer>')
# -
# We see that by expanding `<integer>`, we can cover a total of 12 productions.
quiz("How many productions would `f.max_expansion_coverage('<digit>')` return?",
[
"10",
"11",
"12",
"13"
], "100 / 100")
# Indeed. Here are all the possible expansions for `<digit>`:
# + slideshow={"slide_type": "subslide"}
f.max_expansion_coverage('<digit>')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Determining yet Uncovered Children
#
# We can now start to implement `GrammarCoverageFuzzer`. Our idea is to determine the _missing coverage_ for each child.
#
# Given a list of children, we can use `max_expansion_coverage()` to compute the maximum coverage for each child. From this, we _subtract_ the coverage already seen (`expansion_coverage()`). This results in the coverage we can still obtain.
# + slideshow={"slide_type": "subslide"}
class GrammarCoverageFuzzer(SimpleGrammarCoverageFuzzer):
"""Produce from grammars, aiming for coverage of all expansions."""
def new_child_coverage(self,
symbol: str,
children: List[DerivationTree],
max_depth: Union[int, float] = float('inf')) -> Set[str]:
"""Return new coverage that would be obtained
by expanding (`symbol`, `children`)"""
new_cov = self._new_child_coverage(children, max_depth)
new_cov.add(expansion_key(symbol, children))
new_cov -= self.expansion_coverage() # -= is set subtraction
return new_cov
def _new_child_coverage(self, children: List[DerivationTree],
max_depth: Union[int, float]) -> Set[str]:
new_cov: Set[str] = set()
for (c_symbol, _) in children:
if c_symbol in self.grammar:
new_cov |= self.max_expansion_coverage(c_symbol, max_depth)
return new_cov
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us illustrate `new_child_coverage()`. We again start fuzzing, choosing expansions randomly.
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(EXPR_GRAMMAR, start_symbol="<digit>", log=True)
f.fuzz()
# + [markdown] slideshow={"slide_type": "fragment"}
# This is our current coverage:
# + slideshow={"slide_type": "fragment"}
f.expansion_coverage()
# -
# docassert
assert f.expansion_coverage() == {'<digit> -> 2'}
# If we want to expand `<digit>` into `0`, that would yield us new coverage:
f.new_child_coverage("<digit>", [('0', [])])
# If we want to expand `<digit>` into `2` again, that would yield us _no_ new coverage:
f.new_child_coverage("<digit>", [('2', [])])
# + [markdown] slideshow={"slide_type": "subslide"}
# When we go through the individual expansion possibilities for `<digit>`, we see that all expansions offer additional coverage, _except_ for the `2` we have already covered.
# + slideshow={"slide_type": "fragment"}
for expansion in EXPR_GRAMMAR["<digit>"]:
children = f.expansion_to_children(expansion)
print(expansion, f.new_child_coverage("<digit>", children))
# + [markdown] slideshow={"slide_type": "fragment"}
# This means that whenever choosing an expansion, we can make use of `new_child_coverage()` and choose among the expansions that offer the greatest new (unseen) coverage.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Adaptive Lookahead
# + [markdown] slideshow={"slide_type": "subslide"}
# When choosing a child, we do not look out for the maximum overall coverage to be obtained, as this would have expansions with many uncovered possibilities totally dominate other expansions. Instead, we aim for a _breadth-first_ strategy, first covering all expansions up to a given depth, and only then looking for a greater depth.
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `new_coverages()` is at the heart of this strategy: Starting with a maximum depth (`max_depth`) of zero, it increases the depth until it finds at least one uncovered expansion.
# -
# #### Excursion: Implementing `new_coverage()`
# + slideshow={"slide_type": "subslide"}
class GrammarCoverageFuzzer(GrammarCoverageFuzzer):
def new_coverages(self, node: DerivationTree,
children_alternatives: List[List[DerivationTree]]) \
-> Optional[List[Set[str]]]:
"""Return coverage to be obtained for each child at minimum depth"""
(symbol, children) = node
for max_depth in range(len(self.grammar)):
new_coverages = [
self.new_child_coverage(
symbol, c, max_depth) for c in children_alternatives]
max_new_coverage = max(len(new_coverage)
for new_coverage in new_coverages)
if max_new_coverage > 0:
# Uncovered node found
return new_coverages
# All covered
return None
# -
# #### End of Excursion
# + [markdown] slideshow={"slide_type": "subslide"}
# ### All Together
#
# We can now define `choose_node_expansion()` to make use of this strategy:
# 1. We determine the possible coverages to be obtained (using `new_coverages()`)
# 2. We (randomly) select among the children which sport the maximum coverage (using `choose_uncovered_node_expansion()`).
# -
# #### Excursion: Implementing `choose_node_expansion()`
# + slideshow={"slide_type": "subslide"}
class GrammarCoverageFuzzer(GrammarCoverageFuzzer):
def choose_node_expansion(self, node: DerivationTree,
children_alternatives: List[List[DerivationTree]]) -> int:
"""Choose an expansion of `node` among `children_alternatives`.
Return `n` such that expanding `children_alternatives[n]`
yields the highest additional coverage."""
(symbol, children) = node
new_coverages = self.new_coverages(node, children_alternatives)
if new_coverages is None:
# All expansions covered - use superclass method
return self.choose_covered_node_expansion(node, children_alternatives)
max_new_coverage = max(len(cov) for cov in new_coverages)
children_with_max_new_coverage = [c for (i, c) in enumerate(children_alternatives)
if len(new_coverages[i]) == max_new_coverage]
index_map = [i for (i, c) in enumerate(children_alternatives)
if len(new_coverages[i]) == max_new_coverage]
# Select a random expansion
new_children_index = self.choose_uncovered_node_expansion(
node, children_with_max_new_coverage)
new_children = children_with_max_new_coverage[new_children_index]
# Save the expansion as covered
key = expansion_key(symbol, new_children)
if self.log:
print("Now covered:", key)
self.covered_expansions.add(key)
return index_map[new_children_index]
# -
# #### End of Excursion
# + [markdown] slideshow={"slide_type": "subslide"}
# With this, our `GrammarCoverageFuzzer` is now complete! Let us apply it on a series of examples. On expressions, it quickly covers all digits and operators:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(EXPR_GRAMMAR, min_nonterminals=3)
f.fuzz()
# + slideshow={"slide_type": "fragment"}
f.max_expansion_coverage() - f.expansion_coverage()
# + [markdown] slideshow={"slide_type": "fragment"}
# On average, it is again faster than the simple strategy:
# + slideshow={"slide_type": "fragment"}
average_length_until_full_coverage(GrammarCoverageFuzzer(EXPR_GRAMMAR))
# + [markdown] slideshow={"slide_type": "subslide"}
# On the CGI grammar, it takes but a few iterations to cover all letters and digits:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(CGI_GRAMMAR, min_nonterminals=5)
while len(f.max_expansion_coverage() - f.expansion_coverage()) > 0:
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# This improvement can also be seen in comparing the random, expansion-only, and deep foresight strategies on the CGI grammar:
# + slideshow={"slide_type": "fragment"}
average_length_until_full_coverage(TrackingGrammarCoverageFuzzer(CGI_GRAMMAR))
# + slideshow={"slide_type": "fragment"}
average_length_until_full_coverage(SimpleGrammarCoverageFuzzer(CGI_GRAMMAR))
# + slideshow={"slide_type": "fragment"}
average_length_until_full_coverage(GrammarCoverageFuzzer(CGI_GRAMMAR))
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true toc-nb-collapsed=true
# ## Coverage in Context
#
# Sometimes, grammar elements are used in more than just one place. In our expression grammar, for instance, the `<integer>` symbol is used for integer numbers as well as for floating point numbers:
# + slideshow={"slide_type": "fragment"}
EXPR_GRAMMAR["<factor>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# Our coverage production, as defined above, will ensure that all `<integer>` expansions (i.e., all `<digit>` expansions) are covered. However, the individual digits would be _distributed_ across all occurrences of `<integer>` in the grammar. If our coverage-based fuzzer produces, say, `1234.56` and `7890`, we would have full coverage of all digit expansions. However, `<integer>.<integer>` and `<integer>` in the `<factor>` expansions above would individually cover only a fraction of the digits. If floating-point numbers and whole numbers have different functions that read them in, we would like each of these functions to be tested with all digits; maybe we would also like the whole and fractional part of a floating-point number to be tested with all digits each.
# + [markdown] slideshow={"slide_type": "subslide"}
# Ignoring the context in which a symbol is used (in our case, the various uses of `<integer>` and `<digit>` in the `<factor>` context) can be useful if we can assume that all occurrences of this symbol are treated alike anyway. If not, though, one way to ensure that an occurrence of a symbol is systematically covered independently of other occurrences is to assign the occurrence to a new symbol which is a _duplicate_ of the old symbol. We will first show how to _manually_ create such duplicates, and then a dedicated function which does it automatically.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Extending Grammars for Context Coverage Manually
#
# As stated above, one simple way to achieve coverage in context is by _duplicating_ symbols as well as the rules they reference to. For instance, we could replace `<integer>.<integer>` by `<integer-1>.<integer-2>` and give `<integer-1>` and `<integer-2>` the same definitions as the original `<integer>`. This would mean that not only all expansions of `<integer>`, but also all expansions of `<integer-1>` and `<integer-2>` would be covered.
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us illustrate this with actual code:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar = extend_grammar(EXPR_GRAMMAR,
{
"<factor>": ["+<factor>", "-<factor>", "(<expr>)", "<integer-1>.<integer-2>", "<integer>"],
"<integer-1>": ["<digit-1><integer-1>", "<digit-1>"],
"<integer-2>": ["<digit-2><integer-2>", "<digit-2>"],
"<digit-1>":
["0", "1", "2", "3", "4",
"5", "6", "7", "8", "9"],
"<digit-2>":
["0", "1", "2", "3", "4",
"5", "6", "7", "8", "9"]
}
)
# + slideshow={"slide_type": "skip"}
assert is_valid_grammar(dup_expr_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# If we now run our coverage-based fuzzer on the extended grammar, we will cover all digits both of regular integers, as well as all digits in the whole and fraction part of floating-point numbers:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(dup_expr_grammar, start_symbol="<factor>")
for i in range(10):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "fragment"}
# We see how our "foresighted" coverage fuzzer specifically generates floating-point numbers that cover all digits both in the whole and fractional parts.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Extending Grammars for Context Coverage Programmatically
#
# If we want to enhance coverage in context, manually adapting our grammars may not be the perfect choice, since any change to the grammar will have to be replicated in all duplicates. Instead, we introduce a function that will do the duplication for us.
# + [markdown] slideshow={"slide_type": "subslide"}
# The function `duplicate_context()` takes a grammar, a symbol in the grammar, and an expansion of this symbol (`None` or not given: all expansions of symbol), and it changes the expansion to refer to a duplicate of all originally referenced rules. The idea is that we invoke it as
#
# ```python
# dup_expr_grammar = extend_grammar(EXPR_GRAMMAR)
# duplicate_context(dup_expr_grammar, "<factor>", "<integer>.<integer>")
# ```
#
# and get a similar result as with our manual changes, above.
# + [markdown] slideshow={"slide_type": "subslide"}
# Here is the code:
# + slideshow={"slide_type": "skip"}
from Grammars import new_symbol, unreachable_nonterminals
from GrammarFuzzer import expansion_to_children
# + slideshow={"slide_type": "fragment"}
def duplicate_context(grammar: Grammar,
symbol: str,
expansion: Optional[Expansion] = None,
depth: Union[float, int] = float('inf')):
"""Duplicate an expansion within a grammar.
In the given grammar, take the given expansion of the given `symbol`
(if `expansion` is omitted: all symbols), and replace it with a
new expansion referring to a duplicate of all originally referenced rules.
If `depth` is given, limit duplication to `depth` references
(default: unlimited)
"""
orig_grammar = extend_grammar(grammar)
_duplicate_context(grammar, orig_grammar, symbol,
expansion, depth, seen={})
# After duplication, we may have unreachable rules; delete them
for nonterminal in unreachable_nonterminals(grammar):
del grammar[nonterminal]
# -
# #### Excursion: Implementing `_duplicate_context()`
# + [markdown] slideshow={"slide_type": "subslide"}
# The bulk of the work takes place in this helper function. The additional parameter `seen` keeps track of symbols already expanded and avoids infinite recursion.
# -
import copy
# + slideshow={"slide_type": "fragment"}
def _duplicate_context(grammar: Grammar,
orig_grammar: Grammar,
symbol: str,
expansion: Optional[Expansion],
depth: Union[float, int],
seen: Dict[str, str]) -> None:
"""Helper function for `duplicate_context()`"""
for i in range(len(grammar[symbol])):
if expansion is None or grammar[symbol][i] == expansion:
new_expansion = ""
for (s, c) in expansion_to_children(grammar[symbol][i]):
if s in seen: # Duplicated already
new_expansion += seen[s]
elif c == [] or depth == 0: # Terminal symbol or end of recursion
new_expansion += s
else: # Nonterminal symbol - duplicate
# Add new symbol with copy of rule
new_s = new_symbol(grammar, s)
grammar[new_s] = copy.deepcopy(orig_grammar[s])
# Duplicate its expansions recursively
# {**seen, **{s: new_s}} is seen + {s: new_s}
_duplicate_context(grammar, orig_grammar, new_s, expansion=None,
depth=depth - 1, seen={**seen, **{s: new_s}})
new_expansion += new_s
grammar[symbol][i] = new_expansion
# -
# #### End of Excursion
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's our above example of how `duplicate_context()` works, now with results. We let it duplicate the `<integer>.<integer>` expansion in our expression grammar, and obtain a new grammar with an `<integer-1>.<integer-2>` expansion where both `<integer-1>` and `<integer-2>` refer to copies of the original rules:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar = extend_grammar(EXPR_GRAMMAR)
duplicate_context(dup_expr_grammar, "<factor>", "<integer>.<integer>")
dup_expr_grammar
# + [markdown] slideshow={"slide_type": "subslide"}
# Just like above, using such a grammar for coverage fuzzing will now cover digits in a number of contexts. To be precise, there are five contexts: Regular integers, as well as single-digit and multi-digit whole and fractional parts of floating-point numbers.
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(dup_expr_grammar, start_symbol="<factor>")
for i in range(10):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# The `depth` parameter controls how deep the duplication should go. Setting `depth` to 1 will duplicate only the next rule:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar = extend_grammar(EXPR_GRAMMAR)
duplicate_context(dup_expr_grammar, "<factor>", "<integer>.<integer>", depth=1)
dup_expr_grammar
# + slideshow={"slide_type": "skip"}
assert is_valid_grammar(dup_expr_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# By default, `depth` is set to $\infty$, indicating unlimited duplication. True unbounded duplication could lead to problems for a recursive grammar such as `EXPR_GRAMMAR`, so `duplicate_context()` is set to no longer duplicate symbols once duplicated. Still, if we apply it to duplicate _all_ `<expr>` expansions, we obtain a grammar with no less than 292 rules:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar = extend_grammar(EXPR_GRAMMAR)
duplicate_context(dup_expr_grammar, "<expr>")
# + slideshow={"slide_type": "fragment"}
assert is_valid_grammar(dup_expr_grammar)
len(dup_expr_grammar)
# + [markdown] slideshow={"slide_type": "fragment"}
# This gives us almost 2000 expansions to cover:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(dup_expr_grammar)
len(f.max_expansion_coverage())
# + [markdown] slideshow={"slide_type": "subslide"}
# Duplicating one more time keeps on both growing the grammar and the coverage requirements:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar = extend_grammar(EXPR_GRAMMAR)
duplicate_context(dup_expr_grammar, "<expr>")
duplicate_context(dup_expr_grammar, "<expr-1>")
len(dup_expr_grammar)
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(dup_expr_grammar)
len(f.max_expansion_coverage())
# + [markdown] slideshow={"slide_type": "subslide"}
# At this point, plenty of contexts can be covered individually – for instance, multiplications of elements within additions:
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar["<expr>"]
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar["<term-1-1>"]
# + slideshow={"slide_type": "fragment"}
dup_expr_grammar["<factor-1-1>"]
# + [markdown] slideshow={"slide_type": "subslide"}
# The resulting grammars may no longer be useful for human maintenance; but running a coverage-driven fuzzer such as `GrammarCoverageFuzzer()` will then go and cover all these expansions in all contexts. If you want to cover elements in a large number of contexts, then `duplicate_context()` followed by a coverage-driven fuzzer is your friend.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true toc-nb-collapsed=true
# ## Covering Code by Covering Grammars
#
# With or without context: By systematically covering all input elements, we get a larger variety in our inputs – but does this translate into a wider variety of program behaviors? After all, these behaviors are what we want to cover, including the unexpected behaviors.
# + [markdown] slideshow={"slide_type": "fragment"}
# In a grammar, there are elements that directly correspond to program features. A program handling arithmetic expressions will have functionality that is directly triggered by individual elements - say, an addition feature triggered by the presence of `+`, subtraction triggered by the presence of `-`, and floating-point arithmetic triggered by the presence of floating-point numbers in the input.
# + [markdown] slideshow={"slide_type": "fragment"}
# Such a connection between input structure and functionality leads to a strong _correlation between grammar coverage and code coverage_. In other words: If we can achieve a high grammar coverage, this also leads to a high code coverage.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### CGI Grammars
#
# Let us explore this relationship on one of our grammars – say, the CGI decoder from the [chapter on coverage](Coverage.ipynb).
# + [markdown] tags=[]
# #### Excursion: Creating the Plot
# -
# We compute a mapping `coverages` where in `coverages[x]` = `{y_1, y_2, ...}`, `x` is the grammar coverage obtained, and `y_n` is the code coverage obtained for the `n`-th run.
# + [markdown] slideshow={"slide_type": "subslide"}
# We first compute the maximum coverage, as in the the [chapter on coverage](Coverage.ipynb):
# + slideshow={"slide_type": "skip"}
from Coverage import Coverage, cgi_decode
# + slideshow={"slide_type": "fragment"}
with Coverage() as cov_max:
cgi_decode('+')
cgi_decode('%20')
cgi_decode('abc')
try:
cgi_decode('%?a')
except:
pass
# + [markdown] slideshow={"slide_type": "subslide"}
# Now, we run our experiment:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(CGI_GRAMMAR, max_nonterminals=2)
coverages: Dict[float, List[float]] = {}
trials = 100
for trial in range(trials):
f.reset_coverage()
overall_cov = set()
max_cov = 30
for i in range(10):
s = f.fuzz()
with Coverage() as cov:
cgi_decode(s)
overall_cov |= cov.coverage()
x = len(f.expansion_coverage()) * 100 / len(f.max_expansion_coverage())
y = len(overall_cov) * 100 / len(cov_max.coverage())
if x not in coverages:
coverages[x] = []
coverages[x].append(y)
# + [markdown] slideshow={"slide_type": "subslide"}
# We compute the averages for the `y`-values:
# + slideshow={"slide_type": "fragment"}
xs = list(coverages.keys())
ys = [sum(coverages[x]) / len(coverages[x]) for x in coverages]
# + [markdown] slideshow={"slide_type": "fragment"}
# and create a scatter plot:
# + slideshow={"slide_type": "skip"}
# %matplotlib inline
# + slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt # type: ignore
# + slideshow={"slide_type": "skip"}
import matplotlib.ticker as mtick # type: ignore
# + slideshow={"slide_type": "subslide"}
ax = plt.axes(label="CGI coverage")
ax.yaxis.set_major_formatter(mtick.PercentFormatter())
ax.xaxis.set_major_formatter(mtick.PercentFormatter())
plt.xlim(0, max(xs))
plt.ylim(0, max(ys))
plt.title('Coverage of cgi_decode() vs. grammar coverage')
plt.xlabel('grammar coverage (expansions)')
plt.ylabel('code coverage (lines)')
# + [markdown] tags=[]
# #### End of Excursion
# -
# This scatter plot shows the relationship between grammar coverage (X axis) and code coverage (Y axis).
# + slideshow={"slide_type": "subslide"}
# ignore
plt.scatter(xs, ys);
# + [markdown] slideshow={"slide_type": "fragment"}
# We see that the higher the grammar coverage, the higher the code coverage.
# + [markdown] slideshow={"slide_type": "subslide"}
# This also translates into a correlation coefficient of about 0.9, indicating a strong correlation:
# + slideshow={"slide_type": "skip"}
import numpy as np
# + slideshow={"slide_type": "fragment"}
np.corrcoef(xs, ys)
# + [markdown] slideshow={"slide_type": "fragment"}
# This is also confirmed by the Spearman rank correlation:
# + slideshow={"slide_type": "skip"}
from scipy.stats import spearmanr # type: ignore
# + slideshow={"slide_type": "fragment"}
spearmanr(xs, ys)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### URL Grammars
#
# Let us repeat this experiment on URL grammars. We use the same code as above, except for exchanging the grammars and the function in place:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from urllib.parse import urlparse
# + [markdown] tags=[]
# #### Excursion: Creating the Plot
# + [markdown] slideshow={"slide_type": "fragment"}
# Again, we first compute the maximum coverage, making an educated guess as in the [chapter on coverage](Coverage.ipynb):
# + slideshow={"slide_type": "fragment"}
with Coverage() as cov_max:
urlparse("http://foo.bar/path")
urlparse("https://foo.bar#fragment")
urlparse("ftp://user:password@foo.bar?query=value")
urlparse("ftps://127.0.0.1/?x=1&y=2")
# + [markdown] slideshow={"slide_type": "subslide"}
# Here comes the actual experiment:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(URL_GRAMMAR, max_nonterminals=2)
coverages: Dict[float, List[float]] = {}
trials = 100
for trial in range(trials):
f.reset_coverage()
overall_cov = set()
for i in range(20):
s = f.fuzz()
with Coverage() as cov:
urlparse(s)
overall_cov |= cov.coverage()
x = len(f.expansion_coverage()) * 100 / len(f.max_expansion_coverage())
y = len(overall_cov) * 100 / len(cov_max.coverage())
if x not in coverages:
coverages[x] = []
coverages[x].append(y)
# + slideshow={"slide_type": "subslide"}
xs = list(coverages.keys())
ys = [sum(coverages[x]) / len(coverages[x]) for x in coverages]
# + slideshow={"slide_type": "subslide"}
ax = plt.axes(label="URL coverage")
ax.yaxis.set_major_formatter(mtick.PercentFormatter())
ax.xaxis.set_major_formatter(mtick.PercentFormatter())
plt.xlim(0, max(xs))
plt.ylim(0, max(ys))
plt.title('Coverage of urlparse() vs. grammar coverage')
plt.xlabel('grammar coverage (expansions)')
plt.ylabel('code coverage (lines)')
# -
# #### End of Excursion
# This scatter plot shows the relationship between grammar coverage (X axis) and code coverage (Y axis).
# + slideshow={"slide_type": "subslide"}
plt.scatter(xs, ys);
# + [markdown] slideshow={"slide_type": "subslide"}
# Here, we have an even stronger correlation of more than .95:
# + slideshow={"slide_type": "fragment"}
np.corrcoef(xs, ys)
# + [markdown] slideshow={"slide_type": "fragment"}
# This is also confirmed by the Spearman rank correlation:
# + slideshow={"slide_type": "fragment"}
spearmanr(xs, ys)
# + [markdown] slideshow={"slide_type": "fragment"}
# We conclude: If one wants to obtain high code coverage, it is a good idea to strive for high grammar coverage first.
# + [markdown] slideshow={"slide_type": "subslide"} toc-hr-collapsed=true
# ### Will this always work?
#
# The correlation observed for the CGI and URL examples will not hold for every program and every structure.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Equivalent Elements
#
# First, some grammar elements are treated uniformly by a program even though the grammar sees them as different symbols. In the host name of a URL, for instance, we can have many different characters, although a URL-handling program treats them all the same. Likewise, individual digits, once composed into a number, make less of a difference than the value of the number itself. Hence, achieving variety in digits or characters will not necessarily yield a large difference in functionality.
# + [markdown] slideshow={"slide_type": "fragment"}
# This problem can be addressed by _differentiating elements dependent on their context_, and covering alternatives for each context, as discussed above. The key is to identify the contexts in which variety is required, and those where it is not.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Deep Data Processing
#
# Second, the way the data is processed can make a large difference. Consider the input to a _media player_, consisting of compressed media data. While processing the media data, the media player will show differences in behavior (notably in its output), but these differences cannot be directly triggered through individual elements of the media data. Likewise, a _machine learner_ that is trained on a large set of inputs typically will not have its behavior controlled by a single syntactic element of the input. (Well, it could, but then, we would not need a machine learner.) In these cases of "deep" data processing, achieving structural coverage in the grammar will not necessarily induce code coverage.
# + [markdown] slideshow={"slide_type": "fragment"}
# One way to address this problem is to achieve not only _syntactic_, but actually _semantic_ variety. In the [chapter on fuzzing with constraints](GeneratorGrammarFuzzer.ipynb), we will see how to specifically generate and filter input values, especially numerical values. Such generators can also be applied in context, such that each and every facet of the input can be controlled individually.
# Also, in the above examples, _some_ parts of the input can still be covered structurally: _Metadata_ (such as author name or composer for the media player) or _configuration data_ (such as settings for the machine learner) can and should be covered systematically; we will see how this is done [in the chapter on "Configuration fuzzing"](ConfigurationFuzzer.ipynb).
# -
# ## Synopsis
#
# This chapter introduces `GrammarCoverageFuzzer`, an efficient grammar fuzzer extending `GrammarFuzzer` from the [chapter on efficient grammar fuzzing](GrammarFuzzer.ipynb). It strives to _cover all expansions at least once,_ thus ensuring coverage of functionality.
#
# In the following example, for instance, we use `GrammarCoverageFuzzer` to produce an expression. We see that the resulting expression covers all digits and all operators in a single expression.
from Grammars import EXPR_GRAMMAR
expr_fuzzer = GrammarCoverageFuzzer(EXPR_GRAMMAR)
# ignore
expr_fuzzer.fuzz();
expr_fuzzer.fuzz()
# After fuzzing, the `expansion_coverage()` method returns a mapping of grammar expansions covered.
expr_fuzzer.expansion_coverage()
# Subsequent calls to `fuzz()` will go for further coverage (i.e., covering the other area code digits, for example); a call to `reset()` clears the recored coverage, starting anew.
# Since such coverage in inputs also yields higher code coverage, `GrammarCoverageFuzzer` is a recommended extension to `GrammarFuzzer`.
# ignore
from ClassDiagram import display_class_hierarchy
# ignore
display_class_hierarchy([GrammarCoverageFuzzer],
public_methods=[
Fuzzer.run,
Fuzzer.runs,
GrammarFuzzer.__init__,
GrammarFuzzer.fuzz,
GrammarFuzzer.fuzz_tree,
TrackingGrammarCoverageFuzzer.max_expansion_coverage,
TrackingGrammarCoverageFuzzer.missing_expansion_coverage,
TrackingGrammarCoverageFuzzer.reset_coverage,
GrammarCoverageFuzzer.__init__,
GrammarCoverageFuzzer.fuzz,
GrammarCoverageFuzzer.expansion_coverage,
],
types={
'DerivationTree': DerivationTree,
'Expansion': Expansion,
'Grammar': Grammar
},
project='fuzzingbook')
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * Achieving _grammar coverage_ quickly results in a large variety of inputs.
# * Duplicating grammar rules allows to cover elements in specific _contexts_.
# * Achieving grammar coverage can help in obtaining _code coverage_.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# ## Next Steps
#
# From here, you can learn how to
#
# * [use grammar coverage to systematically test configurations](ConfigurationFuzzer.ipynb).
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# The idea of ensuring that each expansion in the grammar is used at least once goes back to Burkhardt \cite{Burkhardt1967}, to be later rediscovered by <NAME> \cite{Purdom1972}. The relation between grammar coverage and code coverage was discovered by <NAME>, who explores it in his PhD thesis.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 1: Testing ls
#
# Consider the Unix `ls` program, used to list the contents of a directory. Create a grammar for invoking `ls`:
# + cell_style="center" slideshow={"slide_type": "fragment"}
LS_EBNF_GRAMMAR: Grammar = {
'<start>': ['-<options>'],
'<options>': ['<option>*'],
'<option>': ['1', 'A', '@',
# many more
]
}
# + cell_style="center" slideshow={"slide_type": "skip"}
assert is_valid_grammar(LS_EBNF_GRAMMAR)
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
# Use `GrammarCoverageFuzzer` to test all options. Be sure to invoke `ls` with each option set.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** We can copy the set of option characters right from the manual page:
# + slideshow={"slide_type": "skip"} solution2="hidden"
from Grammars import convert_ebnf_grammar, srange
# + slideshow={"slide_type": "skip"} solution2="hidden"
LS_EBNF_GRAMMAR: Grammar = {
'<start>': ['-<options>'],
'<options>': ['<option>*'],
'<option>': srange("ABCFGHLOPRSTUW@abcdefghiklmnopqrstuwx1")
}
# + slideshow={"slide_type": "skip"} solution2="hidden"
assert is_valid_grammar(LS_EBNF_GRAMMAR)
# + slideshow={"slide_type": "skip"} solution2="hidden"
LS_GRAMMAR: Grammar = convert_ebnf_grammar(LS_EBNF_GRAMMAR)
# + slideshow={"slide_type": "skip"} solution2="hidden"
from Fuzzer import ProgramRunner
# + slideshow={"slide_type": "skip"} solution2="hidden"
f = GrammarCoverageFuzzer(LS_GRAMMAR, max_nonterminals=3)
while len(f.max_expansion_coverage() - f.expansion_coverage()) > 0:
invocation = f.fuzz()
print("ls", invocation, end="; ")
args = invocation.split()
ls = ProgramRunner(["ls"] + args)
ls.run()
print()
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# By setting `max_nonterminals` to other values, you can control how many options `ls` should be invoked with. We will see more of such examples in the [chapter on configuration testing](ConfigurationFuzzer.ipynb).
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 2: Caching
#
# The value of `max_expansion_coverage()` depends on the grammar only. Change the implementation such that the values are precomputed for each symbol and depth upon initialization (`__init__()`); this way, `max_expansion_coverage()` can simply lookup the value in the table.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** This is like exercise 1 and 2 [in the chapter on efficient grammar fuzzing](GrammarFuzzer.ipynb); you can implement a similar solution here.
| notebooks/GrammarCoverageFuzzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis]
# language: python
# name: conda-env-analysis-py
# ---
import dask.dataframe as dd
df = dd.read_csv('results/*.csv')
df.head()
df = df.drop(['Unnamed: 0'], axis=1)
df = df.compute()
df.head()
df.to_csv('clean_esmlab_scaling_results.csv', index=False)
# %load_ext watermark
# %watermark --iversion -g -m -v -u -d -h
| benchmarks/data-cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xl
from utils import *
import tensorflow as tf
from sklearn.cross_validation import train_test_split
import time
import random
import os
trainset = sklearn.datasets.load_files(container_path = 'data', encoding = 'UTF-8')
trainset.data, trainset.target = separate_dataset(trainset,1.0)
print (trainset.target_names)
print (len(trainset.data))
print (len(trainset.target))
concat = ' '.join(trainset.data).split()
vocabulary_size = len(list(set(concat)))
data, count, dictionary, rev_dictionary = build_dataset(concat, vocabulary_size)
print('vocab from size: %d'%(vocabulary_size))
print('Most common words', count[4:10])
print('Sample data', data[:10], [rev_dictionary[i] for i in data[:10]])
GO = dictionary['GO']
PAD = dictionary['PAD']
EOS = dictionary['EOS']
UNK = dictionary['UNK']
learning_rate = 1e-4
batch_size = 32
maxlen = 100
n_layer = 3
d_model = 256
d_embed = 256
n_head = 10
d_head = 50
d_inner = 512
class Model:
def __init__(self):
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
self.memory = tf.fill([n_layer,
tf.shape(self.X)[0],
tf.shape(self.X)[1],
d_model], PAD)
self.memory = tf.cast(self.memory, tf.float32)
initializer = tf.initializers.random_normal(stddev = 0.1)
logits, self.next_memory = xl.transformer(
self.X,
self.memory,
len(dictionary),
n_layer,
d_model,
d_embed,
n_head,
d_head,
d_inner,
initializer
)
logits = tf.reduce_mean(logits,axis=1)
self.logits = tf.layers.dense(logits, 2)
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(
self.cost
)
correct_pred = tf.equal(
tf.argmax(self.logits, 1, output_type = tf.int32), self.Y
)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model()
sess.run(tf.global_variables_initializer())
vectors = str_idx(trainset.data,dictionary, maxlen)
train_X, test_X, train_Y, test_Y = train_test_split(vectors, trainset.target,test_size = 0.2)
# +
from tqdm import tqdm
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 3, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
pbar = tqdm(
range(0, len(train_X), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
batch_x = train_X[i : min(i + batch_size, train_X.shape[0])]
batch_y = train_Y[i : min(i + batch_size, train_X.shape[0])]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x
},
)
assert not np.isnan(cost)
train_loss += cost
train_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_X), batch_size), desc = 'test minibatch loop')
for i in pbar:
batch_x = test_X[i : min(i + batch_size, test_X.shape[0])]
batch_y = test_Y[i : min(i + batch_size, test_X.shape[0])]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x
},
)
test_loss += cost
test_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss /= len(train_X) / batch_size
train_acc /= len(train_X) / batch_size
test_loss /= len(test_X) / batch_size
test_acc /= len(test_X) / batch_size
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
EPOCH += 1
# +
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
batch_x = test_X[i : min(i + batch_size, test_X.shape[0])]
batch_y = test_Y[i : min(i + batch_size, test_X.shape[0])]
batch_memory = np.ones(([n_layer, batch_size, maxlen, d_model]))
predict_Y += np.argmax(
sess.run(
model.logits, feed_dict = {model.X: batch_x, model.Y: batch_y,
model.memory:batch_memory}
),
1,
).tolist()
real_Y += batch_y
# -
print(metrics.classification_report(real_Y, predict_Y, target_names = trainset.target_names))
batch_x.shape
| text-classification/64.transformer-xl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Write Number in Expanded Form
#
# ### Problem Statement
#
# You will be given a number and you will need to return it as a string in Expanded Form. For example:
#
# expanded_form(12) # Should return '10 + 2'
# expanded_form(42) # Should return '40 + 2'
# expanded_form(70304) # Should return '70000 + 300 + 4'
#
# NOTE: All numbers will be whole numbers greater than 0.
def expanded_form(num):
# convert number to a string
num_str = str(num)
# create a list with each digit raised to its respective power
power = [str(int(num_str[i])*10**(len(num_str) - 1 - i)) for i in range(len(num_str))]
# combine list into a string
nozeros = ' + '.join(filter(lambda a: a != '0', power))
return nozeros
# ### Test
print('{0}: {1}'.format('expanded form of 12', expanded_form(12)))
print('{0}: {1}'.format('expanded form of 42', expanded_form(42)))
print('{0}: {1}'.format('expanded form of 70304', expanded_form(70304)))
| Write Number in Expanded Form.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, time
import numpy as np
import logging
import torch
import torch.optim as optim
import torch.nn as nn
from torch.utils.data import DataLoader
from model import *
from dataset import *
import matplotlib.pyplot as plt
import torch.nn.functional as F
class FeatureExtractor(nn.Module):
def __init__(self, submodule, extracted_layers):
super(FeatureExtractor,self).__init__()
self.submodule = submodule
self.extracted_layers= extracted_layers
def forward(self, x):
outputs = []
for name, module in self.submodule._modules.items():
if name is "fc": x = x.view(x.size(0), -1)
x = module(x)
#print(name)
if name in self.extracted_layers:
outputs.append(x)
return outputs
def evaluate(dataloader, model, device):
c = 0
total_num = 0
with torch.no_grad():
for data, target in dataloader:
data, target = data.to(device), target.to(device).squeeze()
total_num += len(data)
out = model(data)
predicted = torch.max(out, 1)[1]
c += (predicted == target).sum().item()
return c * 100.0 / total_num
def score(**kwargs):
data_dir = kwargs.get('data_dir', './data')
model_dir = kwargs.get('model_dir', 'models')
batch_size = kwargs.get('batch_size', 1)
device = torch.device("cpu")
eval_dataset = DockDataset(featdir=os.path.join(data_dir, 'test'), is_train=False, shuffle=False)
eval_loader = DataLoader(
eval_dataset,
batch_size=batch_size,
shuffle=False,
drop_last=False,
)
# model = resnet18()
model = resnet18_lsoftmax(device=device)
model.load_state_dict(torch.load(os.path.join(model_dir, "model_best.pth")))
model.to(device)
model.eval()
#######################################################################
# conv1 bn1 relu maxpool layer1 layer2 layer3 layer4 avgpool fc
exact_list = ["conv1"]#["conv1","layer1","layer2","layer3","layer4","avgpool"]
extractor = FeatureExtractor(model,exact_list)
with torch.no_grad():
for data, target in eval_loader:
isDocks = target[0].data.numpy()[0]
if( isDocks): continue
x = extractor(data)
#print (x[0].data)
for i in range(1):
ax = plt.subplot(1, 1, i + 1)
ax.set_title('Sample #{}'.format(i))
ax.axis('off')
plt.imshow(x[0].data.numpy()[0,i,:,:],cmap='jet')
plt.show()
break
#######################################################################
# eval_acc = evaluate(eval_loader, model, device)
# print(f"Test Accuracy is: {eval_acc:.2f}%")
score()
# +
import os, time
import numpy as np
import logging
import torch
import torch.optim as optim
import torch.nn as nn
from torch.utils.data import DataLoader
from model import *
from dataset import *
import matplotlib.pyplot as plt
import torch.nn.functional as F
class FeatureExtractor(nn.Module):
def __init__(self, submodule, extracted_layers):
super(FeatureExtractor,self).__init__()
self.submodule = submodule
self.extracted_layers= extracted_layers
def forward(self, x):
outputs = []
for name, module in self.submodule._modules.items():
if name is "fc": x = x.view(x.size(0), -1)
x = module(x)
#print(name)
if name in self.extracted_layers:
outputs.append(x)
print(name)
return outputs
def evaluate(dataloader, model, device):
c = 0
total_num = 0
with torch.no_grad():
for data, target in dataloader:
data, target = data.to(device), target.to(device).squeeze()
total_num += len(data)
out = model(data)
predicted = torch.max(out, 1)[1]
c += (predicted == target).sum().item()
return c * 100.0 / total_num
def score(**kwargs):
data_dir = kwargs.get('data_dir', './data')
model_dir = kwargs.get('model_dir', 'models')
batch_size = kwargs.get('batch_size', 1)
device = torch.device("cpu")
eval_dataset = DockDataset(featdir=os.path.join(data_dir, 'test'), is_train=False, shuffle=False)
eval_loader = DataLoader(
eval_dataset,
batch_size=batch_size,
shuffle=False,
drop_last=False,
)
# model = resnet18()
model = resnet18_lsoftmax(device=device)
model.load_state_dict(torch.load(os.path.join(model_dir, "model_best.pth")))
model.to(device)
model.eval()
#######################################################################
# conv1 bn1 relu maxpool layer1 layer2 layer3 layer4 avgpool fc
exact_list = ["conv1","layer1","layer2","layer4","avgpool"]#["conv1","layer1","layer2","layer3","layer4","avgpool"]
extractor = FeatureExtractor(model,exact_list)
with torch.no_grad():
for data, target in eval_loader:
isDocks = target[0].data.numpy()[0]
if( not isDocks): continue
x = extractor(data)
#print (x[0].data)
for j in range(len(exact_list)):
ax = plt.subplot(2, 2, j + 1)
ax.set_title('Sample #{}'.format(j))
ax.axis('off')
plt.imshow(x[j].data.numpy()[0,1,:,:],cmap='jet')
plt.show()
#######################################################################
# eval_acc = evaluate(eval_loader, model, device)
# print(f"Test Accuracy is: {eval_acc:.2f}%")
score()
# -
| lu/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
from sqlalchemy import text
from sqlalchemy.engine import create_engine
# # Initialize
engine = create_engine("sqlite+pysqlite:///:memory:", echo=True, future=True)
# # Main
| spark/notebooks/Template SQLAlchemy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Celebrity Quote Analysis with The Cognitive Services on Spark
# <img src="https://mmlspark.blob.core.windows.net/graphics/SparkSummit2/cog_services.png" width="800" style="float: center;"/>
# +
from mmlspark.cognitive import *
from pyspark.ml import PipelineModel
from pyspark.sql.functions import col, udf
from pyspark.ml.feature import SQLTransformer
import os
#put your service keys here
TEXT_API_KEY = os.environ["TEXT_API_KEY"]
VISION_API_KEY = os.environ["VISION_API_KEY"]
BING_IMAGE_SEARCH_KEY = os.environ["BING_IMAGE_SEARCH_KEY"]
# -
# ### Extracting celebrity quote images using Bing Image Search on Spark
#
# Here we define two Transformers to extract celebrity quote images.
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/Cog%20Service%20NB/step%201.png" width="600" style="float: center;"/>
# +
imgsPerBatch = 10 #the number of images Bing will return for each query
offsets = [(i*imgsPerBatch,) for i in range(100)] # A list of offsets, used to page into the search results
bingParameters = spark.createDataFrame(offsets, ["offset"])
bingSearch = BingImageSearch()\
.setSubscriptionKey(BING_IMAGE_SEARCH_KEY)\
.setOffsetCol("offset")\
.setQuery("celebrity quotes")\
.setCount(imgsPerBatch)\
.setOutputCol("images")
#Transformer to that extracts and flattens the richly structured output of Bing Image Search into a simple URL column
getUrls = BingImageSearch.getUrlTransformer("images", "url")
# -
# ### Recognizing Images of Celebrities
# This block identifies the name of the celebrities for each of the images returned by the Bing Image Search.
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/Cog%20Service%20NB/step%202.png" width="600" style="float: center;"/>
# +
celebs = RecognizeDomainSpecificContent()\
.setSubscriptionKey(VISION_API_KEY)\
.setModel("celebrities")\
.setUrl("https://eastus.api.cognitive.microsoft.com/vision/v2.0/")\
.setImageUrlCol("url")\
.setOutputCol("celebs")
#Extract the first celebrity we see from the structured response
firstCeleb = SQLTransformer(statement="SELECT *, celebs.result.celebrities[0].name as firstCeleb FROM __THIS__")
# -
# ### Reading the quote from the image.
# This stage performs OCR on the images to recognize the quotes.
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/Cog%20Service%20NB/step%203.png" width="600" style="float: center;"/>
# +
from mmlspark.stages import UDFTransformer
recognizeText = RecognizeText()\
.setSubscriptionKey(VISION_API_KEY)\
.setUrl("https://eastus.api.cognitive.microsoft.com/vision/v2.0/recognizeText")\
.setImageUrlCol("url")\
.setMode("Printed")\
.setOutputCol("ocr")\
.setConcurrency(5)
def getTextFunction(ocrRow):
if ocrRow is None: return None
return "\n".join([line.text for line in ocrRow.recognitionResult.lines])
# this transformer wil extract a simpler string from the structured output of recognize text
getText = UDFTransformer().setUDF(udf(getTextFunction)).setInputCol("ocr").setOutputCol("text")
# -
# ### Understanding the Sentiment of the Quote
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/Cog%20Service%20NB/step4.jpg" width="600" style="float: center;"/>
# +
sentimentTransformer = TextSentiment()\
.setTextCol("text")\
.setUrl("https://eastus.api.cognitive.microsoft.com/text/analytics/v3.0/sentiment")\
.setSubscriptionKey(TEXT_API_KEY)\
.setOutputCol("sentiment")
#Extract the sentiment score from the API response body
getSentiment = SQLTransformer(statement="SELECT *, sentiment[0].sentiment as sentimentLabel FROM __THIS__")
# -
# ### Tying it all together
#
# Now that we have built the stages of our pipeline its time to chain them together into a single model that can be used to process batches of incoming data
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/Cog%20Service%20NB/full_pipe_2.jpg" width="800" style="float: center;"/>
# +
from mmlspark.stages import SelectColumns
# Select the final coulmns
cleanupColumns = SelectColumns().setCols(["url", "firstCeleb", "text", "sentimentLabel"])
celebrityQuoteAnalysis = PipelineModel(stages=[
bingSearch, getUrls, celebs, firstCeleb, recognizeText, getText, sentimentTransformer, getSentiment, cleanupColumns])
celebrityQuoteAnalysis.transform(bingParameters).show(5)
| notebooks/samples/CognitiveServices - Celebrity Quote Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HAAR DWT2
from jax.config import config
config.update("jax_enable_x64", True)
from jax import random
import numpy as np
import jax.numpy as jnp
import pylops
from cr.sparse import lop
import cr.sparse as crs
n = 4000
m =crs.next_pow_of_2(n)
shape = (n, n)
x_jax = random.randint(random.PRNGKey(0), shape, -10, 10)
x_np = np.array(x_jax)
print(m, n)
level = 8
op_np = pylops.signalprocessing.DWT2D(shape, level=level)
op_jax = lop.jit(lop.dwt2D(shape, level=level))
print(op_jax.shape)
y_np = (op_np @ x_np.flatten()).reshape((m, m))
y_jax = op_jax.times(x_jax)
jnp.allclose(y_np, y_jax)
# np_time = %timeit -o (op_np @ x_np.flatten()).reshape((m, m))
# jax_time = %timeit -o op_jax.times(x_jax).block_until_ready()
gain = np_time.average / jax_time.average
print(gain)
z_np = op_np.rmatvec(y_np.flatten()).reshape(shape)
z_jax = op_jax.trans(y_jax)
jnp.allclose(z_np, z_jax)
# np_time = %timeit -o op_np.rmatvec(y_np.flatten()).reshape(shape)
# jax_time = %timeit -o op_jax.trans(y_jax)
gain = np_time.average / jax_time.average
print(gain)
| comparison/pylops/wavelets/haar_dwt2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rolling Update Tests
#
# Check rolling updates function as expected.
import json
import time
# Before we get started we'd like to make sure that we're making all the changes in a new blank namespace of the name `seldon`
# !kubectl create namespace seldon
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# ## Change Image
# We'll want to try modifying an image and seeing how the rolling update performs the changes.
#
# We'll first create the following model:
# %%writefile resources/fixed_v1.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 3
# Now we can run that model and wait until it's released
# !kubectl apply -f resources/fixed_v1.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# Let's confirm that the state of the model is Available
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can modify the model by providing a new image name, using the following config file:
# %%writefile resources/fixed_v2.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 3
# !kubectl apply -f resources/fixed_v2.yaml
# Now let's actually send a couple of requests to make sure that there are no failed requests as the rolling update is performed
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 3:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v1.yaml
# ## Change Replicas (no rolling update)
# We'll want to try modifying number of replicas and no rolling update is needed.
#
# We'll first create the following model:
# %%writefile resources/fixed_v1_rep2.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 2
# Now we can run that model and wait until it's released
# !kubectl apply -f resources/fixed_v1_rep2.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# Let's confirm that the state of the model is Available
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can modify the model by providing a new image name, using the following config file:
# %%writefile resources/fixed_v1_rep4.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 4
# !kubectl apply -f resources/fixed_v1_rep4.yaml
# Now let's actually send a couple of requests to make sure that there are no failed requests as the rolling update is performed
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 4:
break
time.sleep(1)
print("Rollout Success")
# Now downsize back to 2
# !kubectl apply -f resources/fixed_v1_rep2.yaml
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 2:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v1_rep2.yaml
# ## Separate Service Orchestrator
#
# We can test that the rolling update works when we use the annotation that allows us to have the service orchestrator on a separate pod, namely `seldon.io/engine-separate-pod: "true"`, as per the config file below. Though in this case both the service orchestrator and model pod will be recreated.
# %%writefile resources/fixed_v1_sep.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
annotations:
seldon.io/engine-separate-pod: "true"
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 1
# !kubectl apply -f resources/fixed_v1_sep.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# We can wait until the pod is available before starting the rolling update.
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can make a rolling update by changing the version of the docker image we will be updating it for.
# %%writefile resources/fixed_v2_sep.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
annotations:
seldon.io/engine-separate-pod: "true"
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 1
# !kubectl apply -f resources/fixed_v2_sep.yaml
# And we can send requests to confirm that the rolling update is performed without interruptions
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 1:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v1_sep.yaml
# ## Two PodSpecs
# We can test that the rolling update works when we have multiple podSpecs in our deployment and only does a rolling update the first pod (which also contains the service orchestrator)
# %%writefile resources/fixed_v1_2podspecs.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier1
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier2
graph:
name: classifier1
type: MODEL
children:
- name: classifier2
type: MODEL
name: default
replicas: 1
# !kubectl apply -f resources/fixed_v1_2podspecs.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# We can wait until the pod is available before starting the rolling update.
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can make a rolling update by changing the version of the docker image we will be updating it for.
# %%writefile resources/fixed_v2_2podspecs.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier1
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier2
graph:
name: classifier1
type: MODEL
children:
- name: classifier2
type: MODEL
name: default
replicas: 1
# !kubectl apply -f resources/fixed_v2_2podspecs.yaml
# And we can send requests to confirm that the rolling update is performed without interruptions
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 1:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v1_2podspecs.yaml
# ## Two Models
#
# We can test that the rolling update works when we have two predictors / models in our deployment.
# %%writefile resources/fixed_v1_2models.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
- image: seldonio/fixed-model:0.1
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: default
replicas: 3
# !kubectl apply -f resources/fixed_v1_2models.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# We can wait until the pod is available before starting the rolling update.
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can make a rolling update by changing the version of the docker image we will be updating it for.
# %%writefile resources/fixed_v2_2models.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier
- image: seldonio/fixed-model:0.2
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: default
replicas: 3
# !kubectl apply -f resources/fixed_v2_2models.yaml
# And we can send requests to confirm that the rolling update is performed without interruptions
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 3:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v2_2models.yaml
# ## Two Predictors
#
# We can test that the rolling update works when we have two predictors in our deployment.
# %%writefile resources/fixed_v1_2predictors.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
- image: seldonio/fixed-model:0.1
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: a
replicas: 3
traffic: 50
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
- image: seldonio/fixed-model:0.1
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: b
replicas: 1
traffic: 50
# !kubectl apply -f resources/fixed_v1_2predictors.yaml
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
# We can wait until the pod is available before starting the rolling update.
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now we can make a rolling update by changing the version of the docker image we will be updating it for.
# %%writefile resources/fixed_v2_2predictors.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier
- image: seldonio/fixed-model:0.2
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: a
replicas: 3
traffic: 50
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
- image: seldonio/fixed-model:0.1
name: classifier2
graph:
name: classifier
type: MODEL
children:
- name: classifier2
type: MODEL
name: b
replicas: 1
traffic: 50
# !kubectl apply -f resources/fixed_v2_2predictors.yaml
# And we can send requests to confirm that the rolling update is performed without interruptions
time.sleep(5) # To allow operator to start the update
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numReplicas = int(resources["items"][0]["status"]["replicas"])
if numReplicas == 3:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v2_2predictors.yaml
# ## Model name changes
#
# This will not do a rolling update but create a new deployment.
# %%writefile resources/fixed_v1.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.1
name: classifier
graph:
name: classifier
type: MODEL
name: default
replicas: 3
# !kubectl apply -f resources/fixed_v1.yaml
# We can wait until the pod is available.
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=fixed \
# -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
# state = !kubectl get sdep fixed -o jsonpath='{.status.state}'
state = state[0]
print(state)
if state == "Available":
break
time.sleep(1)
assert state == "Available"
# !curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions \
# -H "Content-Type: application/json"
# Now when we apply the update, we should see the change taking place, but there should not be an actual full rolling update triggered.
# %%writefile resources/fixed_v2_new_name.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: fixed
spec:
name: fixed
protocol: seldon
transport: rest
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/fixed-model:0.2
name: classifier2
graph:
name: classifier2
type: MODEL
name: default
replicas: 3
# !kubectl apply -f resources/fixed_v2_new_name.yaml
time.sleep(5)
for i in range(120):
# responseRaw = !curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' -X POST http://localhost:8003/seldon/seldon/fixed/api/v1.0/predictions -H "Content-Type: application/json"
try:
response = json.loads(responseRaw[0])
except:
print("Failed to parse json", responseRaw)
continue
assert response["data"]["ndarray"][0] == 1 or response["data"]["ndarray"][0] == 5
# jsonRaw = !kubectl get deploy -l seldon-deployment-id=fixed -o json
data = "".join(jsonRaw)
resources = json.loads(data)
numItems = len(resources["items"])
if numItems == 1:
break
time.sleep(1)
print("Rollout Success")
# !kubectl delete -f resources/fixed_v2_new_name.yaml
| notebooks/rolling_updates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison of PyBaMM and COMSOL Discharge Curves
# In this notebook we compare the discharge curves obatined by solving the DFN model both in PyBaMM and COMSOL. Results are presented for a range of C-rates, and we see an excellent agreement between the two implementations. If you would like to compare internal varibles please see the script [compare_comsol_DFN](https://github.com/pybamm-team/PyBaMM/blob/comsol-voltage-compare/examples/scripts/compare_comsol/compare_comsol_DFN.py) which creates a slider plot comparing potentials and concentrations as functions of time and space for a given C-rate. For more information on the DFN model, see the [DFN notebook](https://github.com/pybamm-team/PyBaMM/blob/master/examples/notebooks/models/DFN.ipynb).
# First we need to import pybamm, and then change our working directory to the root of the pybamm folder.
import pybamm
import numpy as np
import os
import pickle
import matplotlib.pyplot as plt
os.chdir(pybamm.__path__[0] + "/..")
# We then create a dictionary of the C-rates we would like to solve for and compare. Note that the repository currently only contains COMSOL results for the C-rates listed below.
C_rates = {"01": 0.1, "05": 0.5, "1": 1, "2": 2, "3": 3}
# We get the DFN model equations, geometry, and default parameters. Before processign the model, we adjust the electrode height and depth to be 1 m, to match the one-dimensional model we solved in COMSOL. The model is then processed using the default geometry and updated paramters. Finally, we create a mesh and discretise the model.
# +
# load model and geometry
model = pybamm.lithium_ion.DFN()
geometry = model.default_geometry
# load parameters and process model and geometry
param = model.default_parameter_values
param["Electrode width [m]"] = 1
param["Electrode height [m]"] = 1
param.process_model(model)
param.process_geometry(geometry)
# create mesh
var = pybamm.standard_spatial_vars
var_pts = {var.x_n: 31, var.x_s: 11, var.x_p: 31, var.r_n: 11, var.r_p: 11}
mesh = pybamm.Mesh(geometry, model.default_submesh_types, var_pts)
# discretise model
disc = pybamm.Discretisation(mesh, model.default_spatial_methods)
disc.process_model(model)
# -
# We create the figure by looping over the dictionary of C-rates. In each step of the loop we load the COMSOL results from a .csv file and solve the DFN model in pybamm. The output variables are then processed, allowing us to plot the discharges curve computed using pybamm and COMSOL, and their absolute difference.
# +
# create figure
fig, ax = plt.subplots(figsize=(15, 8))
plt.tight_layout()
plt.subplots_adjust(left=-0.1)
discharge_curve = plt.subplot(211)
plt.xlim([0, 26])
plt.ylim([3.2, 3.9])
plt.xlabel(r"Discharge Capacity (Ah)")
plt.ylabel("Voltage (V)")
plt.title(r"Comsol $\cdots$ PyBaMM $-$")
voltage_difference_plot = plt.subplot(212)
plt.xlim([0, 26])
plt.yscale("log")
plt.grid(True)
plt.xlabel(r"Discharge Capacity (Ah)")
plt.ylabel(r"$\vert V - V_{comsol} \vert$")
# loop over C_rates dict to create plot
for key, C_rate in C_rates.items():
# load the comsol results
comsol_variables = pickle.load(open("input/comsol_results/comsol_{}C.pickle".format(key), 'rb'))
comsol_time = comsol_variables["time"]
comsol_voltage = comsol_variables["voltage"]
# update current density
param["Typical current [A]"] = 24 * C_rate
param.update_model(model, disc)
# discharge timescale
tau = param.process_symbol(
pybamm.standard_parameters_lithium_ion.tau_discharge
).evaluate(0, 0)
# solve model at comsol times
solver = model.default_solver
t = comsol_time / tau
solution = solver.solve(model, t)
# discharge capacity
discharge_capacity = pybamm.ProcessedVariable(
model.variables["Discharge capacity [A.h]"], solution.t, solution.y, mesh=mesh
)
discharge_capacity_sol = discharge_capacity(solution.t)
comsol_discharge_capacity = comsol_time * param["Typical current [A]"] / 3600
# extract the voltage
voltage = pybamm.ProcessedVariable(
model.variables["Terminal voltage [V]"], solution.t, solution.y, mesh=mesh
)
voltage_sol = voltage(solution.t)
# calculate the difference between the two solution methods
end_index = min(len(solution.t), len(comsol_time))
voltage_difference = np.abs(voltage_sol[0:end_index] - comsol_voltage[0:end_index])
# plot discharge curves and absolute voltage_difference
color = next(ax._get_lines.prop_cycler)["color"]
discharge_curve.plot(
comsol_discharge_capacity, comsol_voltage, color=color, linestyle=":"
)
discharge_curve.plot(
discharge_capacity_sol,
voltage_sol,
color=color,
linestyle="-",
label="{} C".format(C_rate),
)
voltage_difference_plot.plot(
discharge_capacity_sol[0:end_index], voltage_difference, color=color
)
discharge_curve.legend(loc="best")
plt.subplots_adjust(
top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.35
)
plt.show()
# -
| examples/notebooks/compare-comsol-discharge-curve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GPR
# Lead Developer: <NAME>
#
# Co-developers: <NAME> & <NAME>
#
# **Overview**
#
# In this tutorial we will request the snow pit 1S1 location and density, ground-penetrating radar (GPR) two-way travel-times (TWT) and geolocation data, and Magnaprobe snow depths and locations to make a quick comparison of the Magnaprobe snow depth measurements and the GPR estimated snow depths.
#
# We will calculate the average density from the pit and visualize a set of GPR travel-times around Pit 1S1. Given the average dry snow density of 1S1, we will then use an empirically derived expression from Kovacs et. al (1995) to estimate the radar wave speed. The wave speed allows us to convert the radar two-way travel-time to snow depth.
#
# Lastly we will use a few summary statistics to compare the GPR and Magnaprobe snow depths, and we will discuss the various potential sources of error that arise naturally, systematically, and/or algorithmically.
#
# **Slides**
#
# https://docs.google.com/presentation/d/1Hh2CdCCvhWzcWzjzHi9WPum5NmOQt67B1vi1AwOzXCQ/edit?usp=sharing
# ## Retrieve density, GPR, and Magnaprobe data from Pit 1S1
#
# **Goal**: Compare the Magnaprobe snow depth to the GPR estimated snow depth from SnowEx 2020 Grand Mesa IOP Pit 1S1
#
#
#
# **Approach**:
#
# 1. Retrieve the pit location from the Layer Data table
# 2. Build a circle of 50 m radius around the pit location
# 3. Request the pit data to get density layers and calculate the average
# 4. Request all the GPR data within a 50 m distance of our pit
# 5. Plot GPR TWT
# 6. Convert TWT to depth using snow density
# 7. Request the Magnaprobe depths around Pit 1S1
# 8. Interpolate GPR depths to the locations of the Magnaprobe depths
# 9. Compare statistics of GPR and Magnaprobe depths
#
# ## Process
# ### Step 1: Get the pit/site coordinates
# We must first import the necessary libraries for operating with the SnowEx SQL database.
# We then import the Point Data (e.g. GPR) and Layer Data (e.g. snow pit) and GeoPandas, PostGIS, and Python functionality. We also establish the Pit Site ID (1S1) and the buffer radius around the pit (50 m).
# +
## Import our DB access function
from snowexsql.db import get_db
# Import the two tables we need GPR ---> PointData, Density (Pits) --> LayerData
from snowexsql.data import PointData, LayerData
from snowexsql.conversions import query_to_geopandas
# Import to make use of the postgis functions on the db that are not necessarily in python
from sqlalchemy import func, Float
# Import datetime module to filter by a date
import datetime
# use numpy to calculate the average of the density results
import numpy as np
# Import matplotlib
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format='retina'
# PIT Site Identifier
site_id = '1S1'
# Distance around the pit to collect data in meters
buffer_dist = 50
# Connect to the database we made.
db_name = 'snow:<EMAIL>/snowex'
engine, session = get_db(db_name)
# Grab our pit geometry (position) object by provided site id from the site details table, Since there is multiple layers and dates we limit the request to 1
q = session.query(LayerData.geom).filter(LayerData.site_id == site_id).limit(1)
site = q.all()
# -
# ### Step 2: Build a buffered circle around our pit
# +
# Cast the geometry point into text to be used by Postgis function ST_Buffer
point = session.query(site[0].geom.ST_AsText()).all()
print(point)
# Create a polygon buffered by our distance centered on the pit using postgis ST_Buffer
q = session.query(func.ST_Buffer(point[0][0], buffer_dist))
buffered_pit = q.all()[0][0]
# -
# ### Step 3: Grab Density Profiles
#
# We query all Layer Data, cast these values to a float, and then compute the average. Then the query is filtered to only the data type 'density'. The output rho_avg_all is the average density of all snow pits, we also filter the query again by site_id to extract the average density of pit 1S1. These density values are then printed to the screen for comparison.
# +
# Request the average (avg) of Layer data casted as a float. We have to cast to a float in the layer table because all main values are stored as a string to
# ...accommodate the hand hardness.
qry = session.query(func.avg(LayerData.value.cast(Float)))
# Filter our query only to density
qry = qry.filter(LayerData.type=='density')
# Request the data
rho_avg_all = qry.all()
# Request the Average Density of Just 1S1
rho_avg_1s1 = qry.filter(LayerData.site_id == site_id).limit(1)
# This is a gotcha. The data in layer data only is stored as a string to accommodate the hand hardness values
print(f"Average density of all pits is {rho_avg_all[0][0]:0.0f} kg/m3")
print(f"Average density of pit 1S1 is {rho_avg_1s1[0][0]:0.0f} kg/m3")
# Cast Densities to float
rho_avg_all = float(rho_avg_all[0][0])
rho_avg_1s1 = float(rho_avg_1s1[0][0])
# -
# ### Step 4: Request all GPR TWT measured inside the buffer
# In this step, we first print all of the instruments and data types contained in PointData. Doing so let's us know that in order to query the GPR two-way travel-times we use the identifier 'two_way_travel'. We then apply a filter for the date January 29, 2020, and refine this query further with the filter for TWT only within our buffered region. Using geopandas, the query is cast into a dataframe. By default the queried PointData type is given the name 'value'. To be more explicit we rename the variable as 'twt' within the dataframe.
# +
# Collect all Point Data where the instrument string contains the GPR in its name
#qry = session.query(PointData).filter(PointData.instrument.contains('GPR'))
# Print all types of PointData in the query
tmp = session.query(PointData.instrument).distinct().all()
print(tmp)
# Print all types of PointData in the query
tmp = session.query(PointData.type).distinct().all()
print(tmp)
qry = session.query(PointData).filter(PointData.type == 'two_way_travel')
# Additionally Filter by a date
qry = qry.filter(PointData.date==datetime.date(2020, 1, 29))
# See upload details at https://github.com/SnowEx/snowexsql/blob/087b382b8d5098f09db67310efd49f777525c0c8/scripts/upload/add_gpr.py#L27
# Grab all the point data in the buffer using the POSTGIS ST_Within, anytime using the postgis functions we typically have to convert to text
qry = qry.filter(func.ST_Within(PointData.geom.ST_AsText(), buffered_pit.ST_AsText()))
# Use our handy dandy function to execute the query and make it a geopandas dataframe
dfGPR = query_to_geopandas(qry, engine)
# rename 'value' in dataframe as 'twt'
dfGPR.rename(columns={'value': 'twt'},inplace=True )
# -
# ### Step 5: Plot it!
# + tags=["nbsphinx-gallery", "nbsphinx-thumbnail"]
# Get the Matplotlib Axes object from the dataframe object, color points by snow depth value
ax = dfGPR.plot(column='twt', legend=True, cmap='PuBu')
# Use non-scientific notation for x and y ticks
ax.ticklabel_format(style='plain', useOffset=False)
# Set the various plots x/y labels and title.
ax.set_title('Grand Mesa GPR Travel-times w/in {}m of site {}'.format(buffer_dist, site_id))
ax.set_xlabel('Easting [m]')
ax.set_ylabel('Northing [m]')
# -
# ### Step 6: Convert TWT to Depth Using Snow Density
# We will relate the dry snow density to the electromagnetic wave speed using the Kovacs et. al (1995) formula
#
# $$
# \epsilon_{\mathrm{r}}^{\prime}=(1+0.845 \rho)^{2} \quad .
# $$ (permitivity)
#
# Equation {eq}`permitivity` calculates the dielectric constant $\epsilon_{\mathrm{r}}^{\prime}$, provided the snow density $\rho$. We must then relate the dielectric constant to the electromagnetic wave speed $(v)$
#
# $$
# v = \frac{c}{\sqrt{\epsilon_{\mathrm{r}}^{\prime}}} \quad .
# $$ (wavespeed)
# In equation {eq}`wavespeed` $c$ is the universal constant $0.3~m/ns$.
#
# We then calculate the depth of the snow
#
# $$
# z = \frac{vt}{2} \quad ,
# $$ (depthconversion)
#
# using the two-way travel-time $(t)$ and the electromagnetic velocity.
#
# We add the GPR estimated snow depths to the dataframe, and print the head of the dataframe to confirm this addition.
# Average Snow Density
# all pits
rho = rho_avg_all
# 1s1
rho = rho_avg_1s1
# convert density to specific gravity
rho = rho/1000
# Calculate Dielectric Permittivity of Snow
epsilon = (1+0.845*rho)**2
c = 0.3 # m/ns
v = c/np.sqrt(epsilon) # m/ns
t = dfGPR.twt
z = v*t/2*100
# Add the GPR depths to the datafram
dfGPR['depth'] = z
dfGPR.head()
# ### Step 7: Get Depth Probes
# We can recall the PointData types from above in Step 4, and we choose 'depth' as the PointData type filter. Again to ensure we are only considering data that was acquired on the same day as the GPR, we filter the depth data by the date January 29, 2020. We further refine this search to the instrument type 'magnaprobe' and of those data only query the points within our buffer. Lastly, we send this query to a new dataframe using the geopandas functionality, and rename the 'value' column as 'depth'.
# +
# Filter by the dataset type depth
qry = session.query(PointData).filter(PointData.type == 'depth')
# Additionally Filter by a date
qry = qry.filter(PointData.date==datetime.date(2020, 1, 29))
# Additionally Filter by instrument
qry = qry.filter(PointData.instrument=='magnaprobe')
# Grab all the point data in the buffer
qry = qry.filter(func.ST_Within(PointData.geom.ST_AsText(), buffered_pit.ST_AsText()))
# Execute the query
# Use our handy dandy function to execute the query and make it a geopandas dataframe
dfProbe = query_to_geopandas(qry, engine)
# rename Probed Depths to dataframe
dfProbe.rename(columns={'value': 'depth'},inplace=True )
dfProbe.head()
# -
# ### Step 8: Average GPR Depths to Compare with Probed Depths
#
# In this step we will compare the GPR estimated depths to the Magnaprobe depths. In order to accomplish this, we must interpolate the GPR locations to the locations of the probe. We will use inverse distance weighting as our interpolation method.
# Example Code taken from https://stackoverflow.com/questions/3104781/inverse-distance-weighted-idw-interpolation-with-python
#
# Inverse distance weighting is a weighted average interpolant. The weights are computed as the inverse of the distance between the GPR locations $(x,y)$ and the depth probe locations (the interpolated locations $(x_i,y_i)$)
#
# $$
# d = {\sqrt{(x-x_i)^{2}+(y-y_i)^2}} \quad , \\
# w = \frac{1}{d} \quad .
# $$ (weights)
#
# Equation {eq}`weights` is then normalized
#
# $$
# w = \frac{w}{\sum{w}} \quad ,
# $$ (idweights)
#
# to sum to one. For the $i^{th}$ location these weights are multiplied by the GPR depths
#
# $$
# z_i = w_i*z \quad ,
# $$ (applyweights)
#
# to compute a weighted average.
#
# In the following code implementation of the inverse distance weighting algorithm, the `subtract.outer` method of the universal functions (ufunc) within numpy is called which computes the distances in equation {eq}`weights` by looping through the interpolation points. The dot product (inner product) is then used to multiply the weights with the GPR depths. This algorithm relies on the use of a for loop within the `ufunc.outer` call, yet it seems quite efficient! A notable caveat of this algorithm, and a source of error, is that the interpolation considers all points in the domain, rather than a localized interpolation. An interpolation scheme that employs a search radius of three meters, rather than 50 meters (as established by the buffer distance in step one), would be preferable.
#
# We then assign the GPR estimated depths to the probe dataframe, and compute the error between the probed depths and the GPR depths.
#
# The head of the depth probe dataframe is printed to show that we have added the interpolated GPR depths and the error.
# +
# Inverse Distance Weighting Interpolation
def simple_idw(x, y, z, xi, yi):
dist = distance_matrix(x,y, xi,yi)
# In IDW, weights are 1 / distance
weights = 1.0 / dist
# Make weights sum to one
weights /= weights.sum(axis=0)
# Multiply the weights for each interpolated point by all observed Z-values
zi = np.dot(weights.T, z)
return zi
def distance_matrix(x0, y0, x1, y1):
obs = np.vstack((x0, y0)).T
interp = np.vstack((x1, y1)).T
# Make a distance matrix between pairwise observations
# Note: from <http://stackoverflow.com/questions/1871536>
# (Yay for ufuncs!)
d0 = np.subtract.outer(obs[:,0], interp[:,0])
d1 = np.subtract.outer(obs[:,1], interp[:,1])
return np.hypot(d0, d1)
# Estimate the GPR Depths at the Probe Locations
z = simple_idw(dfGPR.easting, dfGPR.northing, dfGPR.depth, dfProbe.easting, dfProbe.northing)
# Assign the GPR depths to the Probe dataframe
dfProbe['depthGPR'] = z
# Calculate the Error
err = dfProbe.depth-dfProbe.depthGPR
# Assign the Error to the Probe Dataframe
dfProbe['error'] = err
dfProbe.head()
# -
# ### Step 9. Plot the Depths, Correlation, and Errors
#
# In this final step, we will compare the GPR depths and probed depths. We compute the Pearson correlation
#
# $$
# r=\frac{\sum\left(x_{i}-\bar{x}\right)\left(y_{i}-\bar{y}\right)}{\sqrt{\sum\left(x_{i}-\bar{x}\right)^{2} \sum\left(y_{i}-\bar{y}\right)^{2}}} \quad ,
# $$ (pearson)
#
# where $x$ represents the probed depths and $y$ represents the GPR depths.
#
# We calculate the bias of the GPR estimated depths
#
# $$
# \mathrm{ME}=\frac{\sum_{i=1}^{N} \left( x_{i}-y_{i} \right) }{N} \quad ,
# $$ (me)
#
# as the mean error ($\mathrm{ME}$). Our example at pit 1S1 is relatively unbiased with a $\mathrm{ME}=1.3~cm$. This indicates that the sources of error are uncorrelated and that systematic errors are small.
# The root-mean-square error
#
# $$
# \mathrm{RMSE}=\sqrt{\frac{\sum_{i=1}^{N}\left(x_{i}-y_{i}\right)^{2}}{N}}
# $$ (rmse)
#
# is a measurement of the standard deviation of the errors, if we assume that the errors are normally distributed and independent. In this example the $\mathrm{RMSE}=11~cm$, which is approximately $1/2$ of the L-band GPR wavelength.
#
# **Potential Sources of Error**
# 1. Incorrect density used in depth conversion
# 2. Depth probe entering the soil or air-gap beneath vegitation
# 3. Geolocation errors
# 4. Sample "footprint" size mismatch
# 5. Interpolation
#
# Data biases can be caused by using the incorrect density value. A lower density value will bias the GPR estimated depths to larger values, whereas, a higher density value will bias the GPR depths to lower values. It has also been shown that the point of the probe can enter the soil, which biases the observed depths positively (McGrath et al., 2019). Similarly, vegitation beneath the snow cover can be a source of error. It is possible that the GPR signal is reflected from the air gap beneath snow that is not grounded. In this case the depth probe may contact the ground, though the GPR travel-time does not, leading to a positive bias. Co-location of the GPR and depth probe presents a third possible source of error. Inaccuracy of GPS measurements or probes not coincident with the GPR transect are likely sources of geolocation error. A fourth possible source of error in the comparison of these depths is the disagreement between the size of the GPR "footprint" (known as the fresnel zone radius), which is on the order of one meter, and the area of the probe tip which is about one centimeter. Because these instruments do not sample the same place on the ground, localized variability in the ground topography can lead to errors between the measured and estimated depths. As mentioned in the previous section, the choice of interpolation scheme will affect the accuracy of the co-located depths. It is important to understand the pros and cons of various interpolation algorithms and to document the choice of algorithm used and it's parameters.
#
# In the code ection we, first, display these summary statistics. Then we view this information graphically. The first plot is the scatter of the observed depths versus the estimated depths with the regression line. The second plot is a histogram of the errors (observed - estimated). The histogram shows a slight positive bias, which indicates that a combination of the errors discussed above resulted in the probe measuring snow depths $1~cm$ greater than the GPR on average. The final plot displays the errors in depth spatially. Roughly, by eye, it appears that areas with low travel-time ($\sim4~ns$, southeast quadrant) overestimate the depth, perhaps due to smearing introduced by the non-localized inverse distance weighting algorithm.
#
# +
# Calculate the Correlation
r = np.corrcoef(dfProbe.depth,dfProbe.depthGPR)
print('The correlation is', round(r[0,1],2))
# Calculate the Mean Error
bias = np.mean(dfProbe.error)
print('The bias is', round(bias,2), 'cm')
# Calculate the Mean Absolute Error
mae = np.mean(np.abs(dfProbe.error))
# Calculate the Root Mean Squared Error
rmse = np.sqrt(np.mean((dfProbe.error)**2))
print('The rmse is', round(rmse,2), 'cm')
# Compute the Regression Line
m, b = np. polyfit(dfProbe.depth,dfProbe.depthGPR, 1)
# Plot the Correlation
plt.figure(0)
plt.plot(dfProbe.depth,dfProbe.depthGPR,'o')
plt.plot(dfProbe.depth, m*dfProbe.depth + b,'k')
plt.xlabel('Probe Depth [cm]')
plt.ylabel('GPR Depth [cm]')
# Plot a Histogram of the Errors
plt.figure(1)
plt.hist(dfProbe.error, density=True, bins=10, edgecolor='black') # density=False would make counts
plt.ylabel('PDF')
plt.xlabel('Error [cm]');
# Get the Matplotlib Axes object from the dataframe object, color points by snow depth value
ax = dfProbe.plot(column='error', legend=True, cmap='PuBu')
# Use non-scientific notation for x and y ticks
ax.ticklabel_format(style='plain', useOffset=False)
# Set the various plots x/y labels and title.
ax.set_title('Error [cm] (Probed Depth - GPR Depth)')
ax.set_xlabel('Easting [m]')
ax.set_ylabel('Northing [m]')
# -
# Close the session to avoid hanging transactions
session.close()
| book/tutorials/gpr/gpr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p align="center">
# <img src="https://github.com/GeostatsGuy/GeostatsPy/blob/master/TCG_color_logo.png?raw=true" width="220" height="240" />
#
# </p>
#
# ## Subsurface Data Analytics
#
# ### The Lasso for Subsurface Data Analytics in Python
#
#
# #### <NAME>, Associate Professor, University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
#
# ### PGE 383 Exercise: The Lasso for Subsurface Modeling in Python
#
# Here's a simple workflow, demonstration of the lasso for regression for subsurface modeling workflows. This should help you get started with building subsurface models with data analytics and machine learning. Here's some basic details about the lasso.
#
# The lasso is an extension of linear regresion, and is closely related to ridge regression so let's review some basic details about linear regression first and then related the lasso to ridge regression.
#
# Here's complete workflows and more details on these:
#
# [Linear Regression](https://github.com/GeostatsGuy/PythonNumericalDemos/blob/master/SubsurfaceDataAnalytics_LinearRegression.ipynb)
#
# [Ridge Regression](https://github.com/GeostatsGuy/PythonNumericalDemos/blob/master/SubsurfaceDataAnalytics_RidgeRegression.ipynb)
#
#
# #### Linear Regression
#
# Linear regression for prediction. Here are some key aspects of linear regression:
#
# **Parametric Model**
#
# * the fit model is a simple weighted linear additive model based on all the available features, $x_1,\ldots,x_m$.
#
# * the parametric model takes the form of:
#
# \begin{equation}
# y = \sum_{\alpha = 1}^m b_{\alpha} x_{\alpha} + b_0
# \end{equation}
#
# **Least Squares**
#
# * least squares optimization is applied to select the model parameters, $b_1,\ldots,b_m,b_0$
#
# * we minize the error, residual sum of squares (RSS) over the training data:
#
# \begin{equation}
# RSS = \sum_{i=1}^n (y_i - \left(\sum_{\alpha = 1}^m b_{\alpha} x_{\alpha} + b_0)\right)^2
# \end{equation}
#
# * this could be simplified as the sum of square error over the training data,
#
# \begin{equation}
# \sum_{i=1}^n (\Delta y_i)^2
# \end{equation}
#
# **Assumptions**
#
# * **Error-free** - predictor variables are error free, not random variables
# * **Linearity** - response is linear combination of feature(s)
# * **Constant Variance** - error in response is constant over predictor(s) value
# * **Independence of Error** - error in response are uncorrelated with each other
# * **No multicollinearity** - none of the features are redundant with other features
#
# #### Other Resources
#
# In $Python$, the $SciPy$ package, specifically the $Stats$ functions (https://docs.scipy.org/doc/scipy/reference/stats.html) provide excellent tools for efficient use of statistics.
# I have previously provided linear regression demonstration in R:
#
# 1. [R](https://github.com/GeostatsGuy/geostatsr/blob/master/linear_regression_demo_v2.R)
# 2. [R Markdown](https://github.com/GeostatsGuy/geostatsr/blob/master/linear_regression_demo_v2.Rmd)
# 3. [knit as an HTML document](https://github.com/GeostatsGuy/geostatsr/blob/master/linear_regression_demo_v2.html)
#
# #### The Lasso
#
# With the lasso we add a hyperparameter, $\lambda$, to our minimization, with a shrinkage penalty term.
#
# \begin{equation}
# \sum_{i=1}^n \left(y_i - \left(\sum_{\alpha = 1}^m b_{\alpha} x_{\alpha} + b_0 \right) \right)^2 + \lambda \sum_{j=1}^m |b_{\alpha}|
# \end{equation}
#
# As a result the lasso has 2 criteria:
#
# 1. set the model parameters to minimize the error with training data
#
# 2. shrink the estimates of the slope parameters towards zero. Note: the intercept is not affected by the lambda, $\lambda$, hyperparameter.
#
# Note the only difference between the lasso and ridge regression is:
#
# * for the lasso the shrinkage term is posed as an $\ell_1$ penalty ($\lambda \sum_{\alpha=1}^m |b_{\alpha}|$)
#
# * for ridge regression the shrinkage term is posed as an $\ell_2$ penalty ($\lambda \sum_{\alpha=1}^m \left(b_{\alpha}\right)^2$).
#
# While both ridge regression and the lasso shrink the model parameters ($b_{\alpha}, \alpha = 1,\ldots,m$) towards zero:
#
# * the lasso parameters reach zero at different rates for each predictor feature as the lambda, $\lambda$, hyperparameter increases.
#
# * as a result the lasso provides a method for feature ranking and selection!
#
# The lambda, $\lambda$, hyperparameter controls the degree of fit of the model and may be related to the model variance and bias trade-off.
#
# * for $\lambda \rightarrow 0$ the prediction model approaches linear regression, there is lower model bias, but the model variance is higher
#
# * as $\lambda$ increases the model variance decreases and the model bias increases
#
# * for $\lambda \rightarrow \infty$ the coefficients all become 0.0 and the model is the global mean
#
# #### Workflow Goals
#
# Learn the basics of the lasso in Python to for analysis, modeling and prediction of porosity from density. This includes:
#
# * Basic Python workflows and data preparation
#
# * Training / fitting a the lasso model and comparison to ridge regression
#
# * Checking the model and learning about the impact of hyperparameters
#
# #### Objective
#
# In the PGE 383: Stochastic Subsurface Modeling class I want to provide hands-on experience with building subsurface modeling workflows. Python provides an excellent vehicle to accomplish this. I have coded a package called GeostatsPy with GSLIB: Geostatistical Library (Deutsch and Journel, 1998) functionality that provides basic building blocks for building subsurface modeling workflows.
#
# The objective is to remove the hurdles of subsurface modeling workflow construction by providing building blocks and sufficient examples. This is not a coding class per se, but we need the ability to 'script' workflows working with numerical methods.
#
# #### Getting Started
#
# Here's the steps to get setup in Python with the GeostatsPy package:
#
# 1. Install Anaconda 3 on your machine (https://www.anaconda.com/download/).
# 2. From Anaconda Navigator (within Anaconda3 group), go to the environment tab, click on base (root) green arrow and open a terminal.
# 3. In the terminal type: pip install geostatspy.
# 4. Open Jupyter and in the top block get started by copy and pasting the code block below from this Jupyter Notebook to start using the geostatspy functionality.
#
# There are examples below with these functions. You can go here to see a list of the available functions, https://git.io/fh4eX, other example workflows and source code.
#
# #### Import Required Packages
#
# Let's import the GeostatsPy package.
import os # to set current working directory
import numpy as np # arrays and matrix math
import scipy.stats as st # statistical methods
import pandas as pd # DataFrames
import matplotlib.pyplot as plt # for plotting
from sklearn.metrics import mean_squared_error, r2_score # specific measures to check our models
from sklearn.linear_model import Ridge # ridge regression implemented in scikit learn
from sklearn.linear_model import Lasso # the lasso implemented in scikit learn
from sklearn.model_selection import cross_val_score # multi-processor K-fold crossvalidation
from sklearn.model_selection import train_test_split # train and test split
from sklearn.preprocessing import StandardScaler # standardize the features
from sklearn import metrics # measures to check our models
from sklearn.linear_model import LinearRegression # linear regression implemented in scikit learn
# If you get a package import error, you may have to first install some of these packages. This can usually be accomplished by opening up a command window on Windows and then typing 'python -m pip install [package-name]'. More assistance is available with the respective package docs.
# #### Set the working directory
#
# I always like to do this so I don't lose files and to simplify subsequent read and writes (avoid including the full address each time). Also, in this case make sure to place the required (see below) data file in this working directory.
os.chdir("C:\PGE383") # set the working directory
# #### Loading Data
#
# Let's load the provided dataset. 'Density_Por_data.csv' is available at https://github.com/GeostatsGuy/GeoDataSets. It is a comma delimited file with 20 density ($\frac{g}{cm^3}$) and porosity (as a fraction) measures from the subsurface. We load the data file with the pandas 'read_csv' function into a data frame we called 'df' and then separate it into train and test datasets. The smaples are in random order so we just split the dataset at the 80th sample. We preview each with the head function from Pandas DataFrames.
df = pd.read_csv("Density_Por_data.csv") # read a .csv file in as a DataFrame
df_mv = pd.read_csv("unconv_MV.csv") # load a multivariate dataset that we will use later
df_train = df.iloc[0:80,:] # extract a training set, note samples are random ordered
df_train.head() # preview the DataFrame
df_test = df.iloc[80:] # extract a testing set, note samples are random ordered
df_test.head() # preview the DataFrame
# It is useful to review the summary statistics of our loaded DataFrame. That can be accomplished with the 'describe' DataFrame member function. We transpose to switch the axes for ease of visualization. We will summarize over the training and testing subsets separately.
df_train.describe().transpose()
df_test.describe().transpose()
# Here we extract the ndarrays with porsity and density, training and testing datasets separate arrays for convenience.
por_train = df_train['Porosity'].values # make a shallow copy of the features for convenvience
den_train = df_train['Density'].values
por_test = df_test['Porosity'].values
den_test = df_test['Density'].values
n_train = len(df_train); n_test = len(df_test) # get the number of data in training and testing
# #### Linear Regression Model
#
# Let's first calculate the linear regression model. We use scikit learn and then extend the same workflow to ridge regression.
# +
# Step 1. Instantiate the Model
linear_reg = LinearRegression() # instantiate the linear regression model
# Step 2: Fit the Data on Training Data
linear_reg.fit(df_train["Density"].values.reshape(n_train,1), df_train["Porosity"]) # fit model
density_model = np.linspace(1.2,2.4,10)
# Print the model parameters
porosity_model = linear_reg.predict(density_model.reshape(10,1)) # predict with the fit model
print('Coefficients: ', str(round(linear_reg.coef_[0],3)) + ', Intercept: ', str(round(linear_reg.intercept_,3)))
# Plot model fit
plt.subplot(111)
plt.scatter(df_train["Density"].values, df_train["Porosity"], color='black', s = 20, alpha = 0.3)
plt.plot(density_model,porosity_model, color='red', linewidth=1)
plt.title('Linear Regression Porosity from Density with Training Data'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.,2.6)#; plt.ylim(0,1500000)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.show()
# -
# Let's run some quick model checks. Much more could be done, but I limit this for breviety here.
# +
# Step 3: - Make predictions using the testing dataset
y_pred = linear_reg.predict(df_test['Density'].values.reshape(n_test,1))
# Report the goodness of fit
print('Variance explained: %.2f' % r2_score(df_test['Porosity'].values, y_pred))
# Plot testing diagnostics
plt.subplot(121)
plt.scatter(df_test['Density'].values, df_test['Porosity'].values, color='black', s = 20, alpha = 0.3)
plt.scatter(df_test['Density'], y_pred, color='blue', s = 20, alpha = 0.3)
plt.title('Linear Regression Model Testing - Production from Porosity'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.0,2.6); plt.ylim(5,24)
y_res = y_pred - df_test['Porosity'].values
print('Residual: mean = ' + str(round(np.average(y_res),2)) + ', standard deviation = ' + str(round(np.var(y_res),2)))
plt.subplot(122)
plt.hist(y_res, alpha = 0.2, color = 'red', edgecolor = 'black', bins=20)
plt.title('Linear Regression Model Prediction Error - Porosity form Density'); plt.xlabel('Porosity Estimation Error (%) (Estimate - Truth)'); plt.ylabel('Frequency')
plt.xlim(-4,4)#; plt.ylim(0,1500000)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.2, top=1.2, wspace=0.3, hspace=0.2)
plt.show()
# -
# #### Ridge Regression
#
# Let's replace the scikit learn linear regression method with the scikit learn ridge regression method. Note, we must now set the lambda hyperparameter.
#
# * the hyperparameter is set with the instantiation of the model
# +
lam = 1.0
# Step 1. Instantiate the Model
ridge_reg = Ridge(alpha=lam)
# Step 2: Fit the Data on Training Data
ridge_reg.fit(df_train["Density"].values.reshape(n_train,1), df_train["Porosity"]) # fit model
density_model = np.linspace(1.2,2.4,10)
# Print the model parameters
porosity_ridge_model = ridge_reg.predict(density_model.reshape(10,1)) # predict with the fit model
print('Coefficients: ', str(round(linear_reg.coef_[0],3)) + ', Intercept: ', str(round(linear_reg.intercept_,3)))
# Plot model fit
plt.subplot(111)
plt.scatter(df_train["Density"].values, df_train["Porosity"], color='black', s = 20, alpha = 0.3)
plt.plot(density_model,porosity_ridge_model, color='red', linewidth=1)
plt.title('Ridge Regression Porosity from Density with Training Data'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.,2.6); plt.ylim(8,17)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.show()
# -
# Let's repeat the simple model checks that we applied with our linear regression model.
# +
# Step 3: - Make predictions using the testing dataset
y_pred = ridge_reg.predict(df_test['Density'].values.reshape(n_test,1))
# Report the goodness of fit
print('Variance explained: %.2f' % r2_score(df_test['Porosity'].values, y_pred))
# Plot testing diagnostics
plt.subplot(121)
plt.scatter(df_test['Density'].values, df_test['Porosity'].values, color='black', s = 20, alpha = 0.3)
plt.scatter(df_test['Density'], y_pred, color='blue', s = 20, alpha = 0.3)
plt.title('Ridge Regression Model Testing - Production from Porosity'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.0,2.6); plt.ylim(5,24)
y_res = y_pred - df_test['Porosity'].values
print('Residual: mean = ' + str(round(np.average(y_res),2)) + ', standard deviation = ' + str(round(np.var(y_res),2)))
plt.subplot(122)
plt.hist(y_res, alpha = 0.2, color = 'red', edgecolor = 'black', bins=20)
plt.title('Ridge Regression Model Prediction Error - Porosity form Density'); plt.xlabel('Porosity Estimation Error (%) (Estimate - Truth)'); plt.ylabel('Frequency')
plt.xlim(-4,4)#; plt.ylim(0,1500000)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.2, top=1.2, wspace=0.3, hspace=0.2)
plt.show()
# -
# Interesting, we explained less variance and have a larger residual standard deviation (more error).
#
# * we reduced both testing variance explained and accuracy in this case!
#
# #### the Lasso
#
# Let's replace the scikit learn linear regression and ridge regression methods with the scikit learn the lasso method. Note, we must now set the lambda hyperparameter.
#
# * the lambda hyperparameter, $\lambda$, is set with the instantiation of the model
# +
lam = 1.0
# Step 1. Instantiate the Model
lasso_reg = Lasso(alpha=lam)
# Step 2: Fit the Data on Training Data
lasso_reg.fit(df_train["Density"].values.reshape(n_train,1), df_train["Porosity"]) # fit model
density_model = np.linspace(1.2,2.4,10)
# Print the model parameters
porosity_lasso_model = lasso_reg.predict(density_model.reshape(10,1)) # predict with the fit model
print('Coefficients: ', str(round(linear_reg.coef_[0],3)) + ', Intercept: ', str(round(linear_reg.intercept_,3)))
# Plot model fit
plt.subplot(111)
plt.scatter(df_train["Density"].values, df_train["Porosity"], color='black', s = 20, alpha = 0.3)
plt.plot(density_model,porosity_lasso_model, color='red', linewidth=1)
plt.title('the Lasso Porosity from Density with Training Data'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.,2.6); plt.ylim(8,17)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.show()
# -
# Let's repeat the simple model checks that we applied with our linear regression model.
# +
# Step 3: - Make predictions using the testing dataset
y_pred = lasso_reg.predict(df_test['Density'].values.reshape(n_test,1))
# Report the goodness of fit
print('Variance explained: %.2f' % r2_score(df_test['Porosity'].values, y_pred))
# Plot testing diagnostics
plt.subplot(121)
plt.scatter(df_test['Density'].values, df_test['Porosity'].values, color='black', s = 20, alpha = 0.3)
plt.scatter(df_test['Density'], y_pred, color='blue', s = 20, alpha = 0.3)
plt.title('Linear Regression Model Testing - Production from Porosity'); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.0,2.6); plt.ylim(5,24)
y_res = y_pred - df_test['Porosity'].values
print('Residual: mean = ' + str(round(np.average(y_res),2)) + ', standard deviation = ' + str(round(np.var(y_res),2)))
plt.subplot(122)
plt.hist(y_res, alpha = 0.2, color = 'red', edgecolor = 'black', bins=20)
plt.title('the Lasso Model Prediction Error - Porosity form Density'); plt.xlabel('Porosity Estimation Error (%) (Estimate - Truth)'); plt.ylabel('Frequency')
plt.xlim(-4,4)#; plt.ylim(0,1500000)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.2, top=1.2, wspace=0.3, hspace=0.2)
plt.show()
# -
# What happenned? With our set lambda level
#
# ```python
# lam = 1.0
# ```
#
# the lasso model removed the density feature and estimated with the global mean. The lasso attempts to remove features as a function of the lambda level, in other words, the lasso performs feature selection! Let's investigate this model behavoir.
#
# #### Investigating the Lambda Hyperparameter
#
# Let's look at the multivariate dataset that we already loaded. This way we can observe the model behavoir over a range of features, for a range of lambda hyperparameter values.
#
# We will first remove the well index and preview the multivariate unconventional dataset.
df_mv = df_mv.drop('WellIndex',axis = 1) # remove the well index feature
df_mv.head() # load the comma delimited data file
# Let's calculate the summary statistics for our data.
df_mv.describe().transpose()
# Let's standardize the feature to have:
#
# * mean = 0.0
# * variance = standard deviation = 1.0
#
# To do this we:
#
# 1. instantiate the StandardScaler from scikit learn. We assign it as 'scaler' so we can use it to conveniently reverse the transformation if we like. We will need to do that to get our predictions back into regular production units.
#
# ```python
# scaler = StandardScaler()
# ```
#
# 2. we then extract all the values from our DataFrame and apply the by-column standardization. The result is a 2D ndarray
#
# ```python
# sfeatures = scaler.fit_transform(df_mv.values)
# ```
# 3. we make an new empty DataFrame
#
# ```python
# df_nmv = pd.DataFrame()
# ```
#
# 4. then we add the transformed value to the new DataFrame while keeping the sample index and feature names from the old DataFramae
#
# ```python
# df_nmv = pd.DataFrame(sfeatures, index=df_mv.index, columns=df_mv.columns)
# ```
#
scaler = StandardScaler() # instantiate the scaler
sfeatures = scaler.fit_transform(df_mv.values) # standardize all the values extracted from the DataFrame
df_nmv = pd.DataFrame() # instantiate a new DataFrame
df_nmv = pd.DataFrame(sfeatures, index=df_mv.index, columns=df_mv.columns) # copy the standardized values into the new DataFrame
df_nmv.head() # preview the the new DataFrame
# Let's check the summary statistics.
df_nmv.describe().transpose() # summary statistics from the new DataFrame
# Success, we have all features standardized. We are ready to build our model. Let's extract training and testing datasets.
X_train, X_test, y_train, y_test = train_test_split(df_nmv.iloc[:,:6], pd.DataFrame({'Production':df_nmv['Production']}), test_size=0.33, random_state=73073)
print('Number of training data = ' + str(len(X_train)) + ' and number of testing data = ' + str(len(X_test)))
# Now let's observe the model coefficients ($b_{\alpha}, \alpha = 1,\ldots,m$) for a range of $\lambda$ hyperparameter values.
# +
nbins = 1000 # number of bins to explore the hyperparameter
df_nmv.describe().transpose() # summary statistics from the new DataFrame
lams = np.linspace(0.001,1.0,nbins) # make a list of lambda values
coefs = np.ndarray((nbins,6))
index = 0
for lam in lams:
lasso_reg = Lasso(alpha=lam) # instantiate the model
lasso_reg.fit(X_train, y_train) # fit model
coefs[index,:] = lasso_reg.coef_ # retreive the coefficients
index = index + 1
color = ['black','blue','green','red','orange','grey']
plt.subplot(111) # plot the results
for ifeature in range(0,6):
plt.semilogx(lams,coefs[:,ifeature], label = df_mv.columns[ifeature], c = color[ifeature], linewidth = 3.0)
plt.title('Standardized Model Coefficients vs. Lambda Hyperparameter'); plt.xlabel('Lambda Hyperparameter'); plt.ylabel('Standardized Model Coefficients')
plt.xlim(0.001,1); plt.ylim(-1.0,1.0); plt.grid(); plt.legend(loc = 'lower right')
plt.subplots_adjust(left=0.0, bottom=0.0, right=1.5, top=1., wspace=0.2, hspace=0.2)
plt.show()
# -
# What do we see?
#
# * for a very low lambda value, all features are included
#
# * as we increase the lambda hyperparameter, total organic carbon is the first predictor feature to be removed
#
# * then acoustic impedance, vitrinite reflectance, brittleness, log perm and finally porosity.
#
# * at $\lambda \ge 0.8$ all features are removed.
#
# Let's repeat this workflow with ridge regression for contrast.
# +
nbins = 5000 # number of bins to explore the hyperparameter
lams = np.logspace(-10,7,nbins)
ridge_coefs = np.ndarray((nbins,6))
index = 0
for lam in lams:
ridge_reg = Ridge(alpha=lam)
ridge_reg.fit(X_train, y_train) # fit model
ridge_coefs[index,:] = ridge_reg.coef_
index = index + 1
color = ['black','blue','green','red','orange','grey']
plt.subplot(111)
for ifeature in range(0,6):
plt.semilogx(lams,ridge_coefs[:,ifeature], label = df_mv.columns[ifeature], c = color[ifeature], linewidth = 3.0)
plt.title('Standardized Model Coefficients vs. Lambda Hyperparameter'); plt.xlabel('Lambda Hyperparameter'); plt.ylabel('Standardized Model Coefficients')
plt.xlim(1.0e-10,1.0e7); plt.ylim(-1.0,1.0); plt.grid(); plt.legend(loc = 'lower right')
plt.subplots_adjust(left=0.0, bottom=0.0, right=1.5, top=1., wspace=0.2, hspace=0.2)
plt.show()
# -
# Ridge regression is quite different in the response of predictor feature to change in the lambda hyperparameter.
#
# * there is no selective removal of predictor features as the lambda hyperparameter increases
#
# * a major component is uniform shrinkage of all coefficients towards zero for $\lambda \in [10^1, 10^5]$
#
# Let's check the mode performance for both models over the hyperparameters. We check the mean squared error and the variance explained for:
#
# * linear regression
#
# * ridge regression
#
# * the lasso
# +
nbins = 1000 # number of bins
lams = np.logspace(-14,6,nbins) # assign equal bins in log space
ridge_error = []; lasso_error = []; linear_error = []
ridge_r2 = []; lasso_r2 = []; linear_r2 = []
linear_reg = LinearRegression()
linear_reg.fit(X_train, y_train) # fit model
linear_predict = linear_reg.predict(X_test)
linear_err = metrics.mean_squared_error(y_test["Production"],linear_predict)
linear_r = metrics.r2_score(y_test["Production"],linear_predict)
index = 0
for lam in lams:
ridge_reg = Ridge(alpha=lam)
ridge_reg.fit(X_train, y_train) # fit model
ridge_predict = ridge_reg.predict(X_test)
ridge_error.append(metrics.mean_squared_error(y_test["Production"],ridge_predict))
ridge_r2.append(metrics.r2_score(y_test["Production"],ridge_predict))
lasso_reg = Lasso(alpha=lam)
lasso_reg.fit(X_train, y_train) # fit model
lasso_predict = lasso_reg.predict(X_test)
lasso_error.append(metrics.mean_squared_error(y_test["Production"],lasso_predict))
lasso_r2.append(metrics.r2_score(y_test["Production"],lasso_predict))
linear_error.append(linear_err)
linear_r2.append(linear_r)
index = index + 1
color = ['black','blue','green','red','orange','grey']
plt.subplot(121)
plt.semilogx(lams,linear_error, label = 'Linear Regression', c = 'black', linewidth = 3.0)
plt.semilogx(lams,ridge_error, label = 'Ridge Regression', c = 'blue', linewidth = 3.0)
plt.semilogx(lams,lasso_error, label = 'the Lasso', c = 'red', linewidth = 3.0)
plt.title('Testing MSE vs. Lambda Hyperparameter'); plt.xlabel('Lambda Hyperparameter'); plt.ylabel('Mean Square Error')
plt.xlim(1.0e-14,1.0e6); plt.ylim(0.0,1.0); plt.grid(); plt.legend(loc = 'lower right')
plt.subplot(122)
plt.semilogx(lams,linear_r2, label = 'Linear Regression', c = 'black', linewidth = 3.0)
plt.semilogx(lams,ridge_r2, label = 'Ridge Regression', c = 'blue', linewidth = 3.0)
plt.semilogx(lams,lasso_r2, label = 'the Lasso', c = 'red', linewidth = 3.0)
plt.title('Testing Variance Explained vs. Lambda Hyperparameter'); plt.xlabel('Lambda Hyperparameter'); plt.ylabel('Variance Explained (R2)')
plt.xlim(1.0e-14,1.0e6); plt.ylim(0.0,1.0); plt.grid(); plt.legend(loc = 'upper right')
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.5, wspace=0.2, hspace=0.2)
plt.show()
# -
# This is quite interesting. Let's make some observations.
#
# * linear regression does not have the shrinkage term, nor the lambda parameter so the performance is constant
#
# * for this problem linear regression is the best performer, lowest testing mean square error and higher testing variance explained
#
# * at a low lambda value, ridge regression and the lasso approach linear regression
#
# #### Making Predictions with Our Model
#
# Here's the method to make predictions with our model for a specific lambda hyperparameter.
#
# 1. Instantiate and fit the model.
lam = 0.05
lasso_reg = Lasso(alpha=lam)
lasso_reg.fit(X_train, y_train) # fit model
print(lasso_reg.coef_)
# 2. Standardize the predictor features with our StandardScalar object.
por = 23.2; logperm = 1.2; AI = -0.4; Brittle = 45.2; TOC = 1.3; VR = 2.2; production = -9999.9
pred_features = np.array([por, logperm, AI, Brittle, TOC, VR, production]).reshape(1, -1)
spred_features = scaler.transform(pred_features)
# 3. Predict with the standardize predictor features.
spred_features[0][6] = lasso_reg.predict(spred_features[0][:6].reshape(1, -1))
predict = scaler.inverse_transform(spred_features[0])
predict
# 2. Now we can use the predictor features in our model (we will remove the production dummy).
pred_features = np.array([por, logperm, AI, Brittle, TOC, VR, production]).reshape(1, -1)
scaler.transform(pred_features)
np.array([por, logperm, AI, Brittle, TOC, VR]).reshape(1, -1)
scaler
df_mv.describe().transpose()
# +
# Arrays to store the results
ncases = 100
lamd_mat = np.linspace(0.0,100.0,ncases)
density_model = np.linspace(1.2,2.4,10)
var_explained_train = np.zeros(ncases); var_explained_test = np.zeros(ncases)
mse_train = np.zeros(ncases); mse_test = np.zeros(ncases)
for ilam in range(0,len(lamd_mat)): # Loop over all lambda values
ridge_reg = Ridge(alpha=lamd_mat[ilam])
ridge_reg.fit(df_train["Density"].values.reshape(n_train,1), df_train["Porosity"]) # fit model
porosity_model = ridge_reg.predict(density_model.reshape(10,1)) # predict with the fit model
porosity_pred_train = ridge_reg.predict(df_train['Density'].values.reshape(n_train,1)) # predict with the fit model
var_explained_train[ilam] = r2_score(df_train['Porosity'].values, porosity_pred_train)
mse_train[ilam] = mean_squared_error(df_train['Porosity'].values, porosity_pred_train)
porosity_pred_test = ridge_reg.predict(df_test['Density'].values.reshape(n_test,1))
var_explained_test[ilam] = r2_score(df_test['Porosity'].values, porosity_pred_test)
mse_test[ilam] = mean_squared_error(df_test['Porosity'].values, porosity_pred_test)
if ilam <= 7:
plt.subplot(4,2,ilam+1)
plt.scatter(df_train["Density"].values, df_train["Porosity"], color='black', s = 20, alpha = 0.3)
plt.plot(density_model,porosity_model, color='red', linewidth=1)
plt.title('Ridge Regression Porosity from Density with Training Data - Lambda = ' + str(round(lamd_mat[ilam],2))); plt.xlabel('Density (g/cm^3)'); plt.ylabel('Porosity (%)')
plt.xlim(1.,2.6); plt.ylim(5,24)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=4.2, wspace=0.2, hspace=0.3)
plt.show()
# -
# We can observed from the first 8 cases above of ridge regression model fit that increase in the lambda hyper parameter decreases the slope of the linear fit.
#
# Let's plot the MSE and variance explained over training and testing datasets.
# +
plt.subplot(121)
plt.plot(lamd_mat, var_explained_train, color='blue', linewidth = 2, label = 'Training')
plt.plot(lamd_mat, var_explained_test, color='red', linewidth = 2, label = 'Test')
plt.title('Variance Explained vs. Lambda'); plt.xlabel('Lambda'); plt.ylabel('Variance Explained')
plt.xlim(0.,100.); plt.ylim(0,1.0)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.legend()
plt.subplot(122)
plt.plot(lamd_mat, mse_train, color='blue', linewidth = 2, label = 'Training')
plt.plot(lamd_mat, mse_test, color='red', linewidth = 2, label = 'Test')
plt.title('MSE vs. Lambda'); plt.xlabel('Lambda'); plt.ylabel('Mean Square Error')
plt.xlim(0.,100.); plt.ylim(0,10.0)
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.legend()
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.2, wspace=0.2, hspace=0.3)
plt.show()
# -
# We observe that as we increase the lambda parameter the variance explained decreases and the mean square error increases.
#
# * this makes sense as the data has a consistent linear trend and as the slope 'shrinks' to zero the error increases and the variance explained decreases
#
# * there could be other cases where the reduced slope actually performs better in testing. For example with sparce and noisy data.
#
# #### Model Variance
#
# Now let's explore the concept of model variance, an important part of machine learning accuracy in testing.
#
# * the sensitivity of the model to the specfic training data
#
# * as lambda increases the sensitivity to the training data, model variance decreases
#
# Let's demonstrate this with this workflow:
#
# * loop over multiple lambda values
# * loop over multiple bootstrap samples of the data
# * calculate the ridge regression fit (slope)
# * calculate the variance of these bootstrap results
#
# +
L = 200 # the number of bootstrap realizations
nsamples = 20 # the number of samples in each bootstrap realization
nlambda = 100 # number of lambda values to evaluate
coef_mat = np.zeros(L) # declare arrays to store the results
variance_coef = np.zeros(nlambda)
lamd_mat = np.linspace(0.0,100.0,nlambda)
df = pd.read_csv("Density_Por_data.csv")
for ilam in range(0,len(lamd_mat)): # loop over all lambda values
for l in range(0, L): # loop over all bootstrap realizations
df_sample = df.sample(n = nsamples) # random sample (1 bootstrap)
ridge_reg = Ridge(alpha=lamd_mat[ilam]) # instatiate model
ridge_reg.fit(df_sample["Density"].values.reshape(nsamples,1), df_sample["Porosity"]) # fit model
coef_mat[l] = ridge_reg.coef_[0] # get the slope parameter
variance_coef[ilam] = np.var(coef_mat) # calculate the variance of the slopes over the L bootstraps
# -
# Now let's plot the result.
plt.subplot(111)
plt.plot(lamd_mat, variance_coef, color='black', linewidth = 2, label = 'Slope Variance')
plt.title('Model Fit Variance vs. Lambda'); plt.xlabel('Lambda'); plt.ylabel('Model Fit Variance')
plt.xlim(0.,100.); plt.ylim(0.001,10.0); plt.yscale('log')
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.0, wspace=0.2, hspace=0.2)
plt.legend()
# The result is as expected, with increase in lambda hyperparameter the sensitivity of the model to the training data is decreased.
#
# #### k-fold Cross Validation
#
# It would be useful to conduct a complete k-fold validation to evaluate the testing error vs. the hyperparameter lambda for model tuning.
#
# * the following code should do this
#
# * but with a single feature as input for fitting the fit function requires a reshape
#
# ```python
# my_array.reshape((nsample,1))
# ```
#
# * this is not included in the scikit learn function 'cross_val_score' so we will skip this for now
#
# I have left the code commented out below for reference:
# +
#score = [] # code modified from StackOverFlow by Dimosthenis
#nlambda = 1
#lambd_mat = np.linspace(0.0,100.0,nlambda)
#for ilam in range(0,nlambda):
# ridge_reg = Ridge(alpha=lambd_mat[ilam])
# scores = cross_val_score(estimator=ridge_reg, X= df['Density'].values, y=df['Porosity'].values, cv=10, n_jobs=4, scoring = "neg_mean_squared_error") # Perform 10-fold cross validation
# score.append(abs(scores.mean()))
# -
# #### Comments
#
# Ridge regression is a variant of linear regression that includes a hyperparameter to constrain the degree of model fit. This allow us to tune the variance-bias trade-off of our model. I hope this was helpful,
#
# *Michael*
#
# <NAME>, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
# On twitter I'm the @GeostatsGuy.
#
#
# ***
#
# #### More on <NAME> and the Texas Center for Geostatistics:
#
# ### <NAME>, Associate Professor, University of Texas at Austin
# *Novel Data Analytics, Geostatistics and Machine Learning Subsurface Solutions*
#
# With over 17 years of experience in subsurface consulting, research and development, Michael has returned to academia driven by his passion for teaching and enthusiasm for enhancing engineers' and geoscientists' impact in subsurface resource development.
#
# For more about Michael check out these links:
#
# #### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
# #### Want to Work Together?
#
# I hope this content is helpful to those that want to learn more about subsurface modeling, data analytics and machine learning. Students and working professionals are welcome to participate.
#
# * Want to invite me to visit your company for training, mentoring, project review, workflow design and / or consulting? I'd be happy to drop by and work with you!
#
# * Interested in partnering, supporting my graduate student research or my Subsurface Data Analytics and Machine Learning consortium (co-PIs including Profs. Foster, Torres-Verdin and van Oort)? My research combines data analytics, stochastic modeling and machine learning theory with practice to develop novel methods and workflows to add value. We are solving challenging subsurface problems!
#
# * I can be reached at <EMAIL>.
#
# I'm always happy to discuss,
#
# *Michael*
#
# <NAME>, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
#
# #### More Resources Available at: [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
| SubsurfaceDataAnalytics_Lasso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Meta Models
#
# Certain models in scikit-lego are "meta". Meta models are
# models that depend on other estimators that go in and these
# models will add features to the input model. One way of thinking
# of a meta model is to consider it to be a way to "decorate" a
# model.
#
# This part of the documentation will highlight a few of them.
#
# ## Thresholder
#
# The thresholder can help tweak recall and precision of a model
# by moving the threshold value of `predict_proba`. Commonly this
# threshold is set at 0.5 for two classes. This meta-model can
# decorate an estimator with two classes such that the threshold
# moves.
#
# We demonstrate the working below. First we'll generate a skewed dataset.
#
# +
import numpy as np
import pandas as pd
import matplotlib.pylab as plt
from sklearn.pipeline import Pipeline
from sklearn.datasets import make_blobs
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_score, recall_score, accuracy_score, make_scorer
from sklego.meta import Thresholder
# -
X, y = make_blobs(1000, centers=[(0, 0), (1.5, 1.5)], cluster_std=[1, 0.5])
plt.scatter(X[:, 0], X[:, 1], c=y, s=5);
# Next we'll make a cross validation pipeline to try out this thresholder.
# +
pipe = Pipeline([
("model", Thresholder(LogisticRegression(solver='lbfgs'), threshold=0.1))
])
mod = GridSearchCV(estimator=pipe,
param_grid = {"model__threshold": np.linspace(0.1, 0.9, 50)},
scoring={"precision": make_scorer(precision_score),
"recall": make_scorer(recall_score),
"accuracy": make_scorer(accuracy_score)},
refit="precision",
cv=5)
mod.fit(X, y);
# -
# With this cross validation trained, we'll make a chart to show the
# effect of changing the threshold value.
(pd.DataFrame(mod.cv_results_)
.set_index("param_model__threshold")
[['mean_test_precision', 'mean_test_recall', 'mean_test_accuracy']]
.plot(figsize=(16, 4)));
# Increasing the threshold will increase the precision but as expected this is at the
# cost of recall (and accuracy).
#
# ## Grouped Estimation
#
# <img src="_static/grouped-model.png" width="50%" alt="img1">
#
# To help explain what it can do we'll consider three methods to predict
# the chicken weight. The chicken data has 578 rows and 4 columns
# from an experiment on the effect of diet on early growth of chicks.
# The body weights of the chicks were measured at birth and every second
# day thereafter until day 20. They were also measured on day 21.
# There were four groups on chicks on different protein diets.
#
# ### Setup
#
# Let's first load a bunch of things to do this.
# +
import numpy as np
import pandas as pd
import matplotlib.pylab as plt
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.metrics import mean_absolute_error, mean_squared_error
from sklego.datasets import load_chicken
from sklego.preprocessing import ColumnSelector
df = load_chicken(give_pandas=True)
def plot_model(model):
df = load_chicken(give_pandas=True)
model.fit(df[['diet', 'time']], df['weight'])
metric_df = df[['diet', 'time', 'weight']].assign(pred=lambda d: model.predict(d[['diet', 'time']]))
metric = mean_absolute_error(metric_df['weight'], metric_df['pred'])
plt.figure(figsize=(12, 4))
plt.scatter(df['time'], df['weight'])
for i in [1, 2, 3, 4]:
pltr = metric_df[['time', 'diet', 'pred']].drop_duplicates().loc[lambda d: d['diet'] == i]
plt.plot(pltr['time'], pltr['pred'], color='.rbgy'[i])
plt.title(f"linear model per group, MAE: {np.round(metric, 2)}");
# -
# This code will be used to explain the steps below.
#
# ### Model 1: Linear Regression with Dummies
#
# First we start with a baseline.
# +
feature_pipeline = Pipeline([
("datagrab", FeatureUnion([
("discrete", Pipeline([
("grab", ColumnSelector("diet")),
("encode", OneHotEncoder(categories="auto", sparse=False))
])),
("continous", Pipeline([
("grab", ColumnSelector("time")),
("standardize", StandardScaler())
]))
]))
])
pipe = Pipeline([
("transform", feature_pipeline),
("model", LinearRegression())
])
plot_model(pipe)
# -
# Because the model is linear the dummy variable causes the intercept
# to change but leaves the gradient untouched. This might not be what
# we want from a model. So let's see how the grouped model can adress
# this.
#
# ### Model 2: Linear Regression in GroupedEstimation
#
# The goal of the grouped estimator is to allow us to split up our data.
# The image below demonstrates what will happen.
#
# <img src="_static/grouped-df.png" width="70%" alt="img2">
#
# We train 5 models in total because the model will also train a
# fallback automatically (you can turn this off via `use_fallback=False`).
# The idea behind the fallback is that we can predict something if
# the group does not appear in the prediction.
#
# Each model will accept features that are in `X` that are not
# part of the grouping variables. In this case each group will
# model based on the `time` since `weight` is what we're trying
# to predict.
#
# Applying this model to the dataframe is easy.
from sklego.meta import GroupedEstimator
mod = GroupedEstimator(LinearRegression(), groups=["diet"])
plot_model(mod)
# And the model looks a bit better.
#
# ### Model 3: Dummy Regression in GroupedEstimation
#
# We could go a step further and train a [DummyRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.dummy.DummyRegressor.html) per diet
# per timestep. The code below works similar as the previous example
# but one difference is that the grouped model does not receive a
# dataframe but a numpy array.
#
# <img src="_static/grouped-np.png" width="70%" alt="img3">
#
# Note that we're also grouping over more than one column here.
# The code that does this is listed below.
# +
from sklearn.dummy import DummyRegressor
feature_pipeline = Pipeline([
("datagrab", FeatureUnion([
("discrete", Pipeline([
("grab", ColumnSelector("diet")),
])),
("continous", Pipeline([
("grab", ColumnSelector("time")),
]))
]))
])
pipe = Pipeline([
("transform", feature_pipeline),
("model", GroupedEstimator(DummyRegressor(strategy="mean"), groups=[0, 1]))
])
plot_model(pipe)
# -
# Note that these predictions seems to yield the lowest error but take it
# with a grain of salt since these errors are only based on the train set.
#
# ## Decayed Estimation
#
# Often you are interested in predicting the future. You use the data from
# the past in an attempt to achieve this and it could be said that perhaps
# data from the far history is less relevant than data from the recent past.
#
# This is the idea behind the `DecayEstimator` meta-model. It looks at the
# order of data going in and it will assign a higher importance to recent rows
# that occurred recently and a lower importance to older rows. Recency is based
# on the order so it is imporant that the dataset that you pass in is correctly
# ordered beforehand.
#
# We'll demonstrate how it works by applying it on a simulated timeseries problem.
#
# +
from sklearn.dummy import DummyRegressor
from sklego.meta import GroupedEstimator, DecayEstimator
from sklego.datasets import make_simpleseries
yt = make_simpleseries(seed=1)
df = (pd.DataFrame({"yt": yt,
"date": pd.date_range("2000-01-01", periods=len(yt))})
.assign(m=lambda d: d.date.dt.month)
.reset_index())
plt.figure(figsize=(12, 3))
plt.plot(make_simpleseries(seed=1));
# -
# We will create two models on this dataset. One model calculates the average
# value per month in our timeseries and the other does the same thing but will
# decay the importance of making accurate predictions for the far history.
#
# +
mod1 = (GroupedEstimator(DummyRegressor(), groups=["m"])
.fit(df[['m']], df['yt']))
mod2 = (GroupedEstimator(DecayEstimator(DummyRegressor(), decay=0.9), groups=["m"])
.fit(df[['index', 'm']], df['yt']))
plt.figure(figsize=(12, 3))
plt.plot(df['yt'], alpha=0.5);
plt.plot(mod1.predict(df[['m']]), label="grouped")
plt.plot(mod2.predict(df[['index', 'm']]), label="decayed")
plt.legend();
# -
# The decay parameter has a lot of influence on the effect of the model but one
# can clearly see that we shift focus to the more recent data.
#
# # Confusion Balancer
#
# **Disclaimer**: This is an experimental feature.
#
# We added an experimental feature to the meta estimators that can be used to force balance in the confusion matrix of an estimator. The approach works
n1, n2, n3 = 100, 500, 50
np.random.seed(42)
X = np.concatenate([np.random.normal(0, 1, (n1, 2)),
np.random.normal(2, 1, (n2, 2)),
np.random.normal(3, 1, (n3, 2))],
axis=0)
y = np.concatenate([np.zeros((n1, 1)),
np.ones((n2, 1)),
np.zeros((n3, 1))],
axis=0).reshape(-1)
plt.scatter(X[:, 0], X[:, 1], c=y);
# Let's take this dataset and train a simple classifier against it.
from sklearn.metrics import confusion_matrix
mod = LogisticRegression(solver='lbfgs', multi_class='multinomial', max_iter=10000)
cfm = confusion_matrix(y, mod.fit(X, y).predict(X))
cfm
# The confusion matrix is not ideal. This is in part because the dataset is slightly inbalanced but in general it is also because of the way the algorithm works. Let's see if we can learn something else from this confusion matrix. I might transform the counts into probabilities.
cfm.T / cfm.T.sum(axis=1).reshape(-1, 1)
# Let's consider the number 0.2346 in the lower left corner. This number represents the probability that the actually class 0 while the model predicts class 1. In math we might write this as $P(C_1 | M_1)$ where $C_i$ denotes the actual label while $M_i$ denotes the label given by the algorithm.
#
# The idea now is that we might rebalance our original predictions $P(M_i)$ by multiplying them;
#
# $$ P_{\text{corrected}}(C_1) = P(C_1|M_0) p(M_0) + P(C_1|M_1) p(M_1) $$
#
# In general this can be written as;
#
# $$ P_{\text{corrected}}(C_i) = \sum_j P(C_i|M_j) p(M_j) $$
#
# In laymens terms; we might be able to use the confusion matrix to learn from our mistakes. By how much we correct is something that we can tune with a hyperparameter.
#
# $$ P_{\text{corrected}}(C_i) = \alpha \sum_j P(C_i|M_j) p(M_j) + (1-\alpha) p(M_j) $$
#
# We'll perform an optimistic demonstration below.
# +
def false_positives(mod, x, y):
return (mod.predict(x) != y)[y == 1].sum()
def false_negatives(mod, x, y):
return (mod.predict(x) != y)[y == 0].sum()
# -
from sklego.meta import ConfusionBalancer
# +
cf_mod = ConfusionBalancer(LogisticRegression(solver='lbfgs', max_iter=1000), alpha=1.0)
grid = GridSearchCV(cf_mod,
param_grid={'alpha': np.linspace(-1.0, 3.0, 31)},
scoring={
"accuracy": make_scorer(accuracy_score),
"positives": false_positives,
"negatives": false_negatives
},
n_jobs=-1,
iid=True,
return_train_score=True,
refit="negatives",
cv=5)
# -
df = pd.DataFrame(grid.fit(X, y).cv_results_)
plt.figure(figsize=(12, 3))
plt.subplot(121)
plt.plot(df['param_alpha'], df['mean_test_positives'], label="false positives")
plt.plot(df['param_alpha'], df['mean_test_negatives'], label="false negatives")
plt.legend()
plt.subplot(122)
plt.plot(df['param_alpha'], df['mean_test_accuracy'], label="test accurracy")
plt.plot(df['param_alpha'], df['mean_train_accuracy'], label="train accurracy")
plt.legend();
# It seems that we can pick a value for $\alpha$ such that the confusion matrix is balanced. There's also a modest increase in accuracy for this balancing moment.
#
# It should be emphesized though that this feature is **experimental**. There have been dataset/model combinations where this effect seems to work very well while there have also been situations where this trick does not work at all. It also deserves mentioning that there might be alternative to your problem. If your dataset is suffering from a huge class imbalance then you might be better off by having a look at the [imbalanced-learn](https://imbalanced-learn.readthedocs.io/en/stable/) project.
| doc/meta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from pathlib import Path
dPath = Path("../docs/dumps")
import pickle
with open(dPath / "train_data.pkl", 'rb') as filename:
train_data = pickle.load(filename)
with open(dPath / "valid_data.pkl", 'rb') as filename:
valid_data = pickle.load(filename)
X_train = train_data.drop("Detected", axis=1)
y_train = train_data.Detected
X_valid = valid_data.drop("Detected", axis=1)
y_valid = valid_data.Detected
with open(dPath / "rf_exp_04_names.pkl", 'rb') as filename:
names = pickle.load(filename)
X_train = X_train[names]
X_valid = X_valid[names]
X_train.head()
from imblearn.over_sampling import ADASYN
sm = ADASYN(random_state=42, n_jobs=-1, n_neighbors=5)
# %time X_train, y_train = sm.fit_resample(X_train, y_train)
import lightgbm
lgbm = lightgbm.LGBMClassifier(
seed=42,
n_estimators=3000,
objective='binary',
n_jobs=-1)
# %time lgbm.fit(X_train,y_train)
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score
def conf_matr(m):
y_pred = m.predict(X_valid)
print(classification_report(y_valid, y_pred))
def print_score(m):
res = [m.score(X_train, y_train), m.score(X_valid, y_valid), roc_auc_score(y_valid, m.predict(X_valid))]
if hasattr(m, 'oob_score_'): res.append(m.oob_score_)
print(res)
print_score(lgbm)
conf_matr(lgbm)
with open(dPath / "lgbm_exp_01.pkl", 'wb') as filename:
pickle.dump(lgbm,filename)
| notebooks/lgbm_experiment_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import kutils
from kutils.model_helper import ModelHelper
from kutils import applications as apps
from kutils import image_utils as iu
from kutils import tensor_ops as ops
from kutils import generic as gen
import resnet101
import pandas as pd
import keras
import numpy as np
from keras.losses import categorical_crossentropy
from keras.models import Model
from keras import backend as K
# -
# ### Initialize dataset meta
# +
aux_root = '/mnt/home/research/data/'
data_root = '/mnt/home/research/koniq/'
ids = pd.read_csv(data_root + 'metadata/koniq10k_distributions_sets.csv')
# define 5 classes
mos_class = np.int32(ids.MOS/20.)
ids.loc[:,'class'] = mos_class
x = keras.utils.to_categorical(mos_class)
classes = pd.DataFrame(x.tolist())
output_classes = ['class0', 'class1', 'class2', 'class3', 'class4']
classes.columns = output_classes
ids = pd.concat([ids, classes], axis=1, verify_integrity=True)
from sklearn.utils import class_weight
class_weights = class_weight.compute_class_weight('balanced',
np.unique(mos_class),
mos_class)
# -
# ### DeepRN base model
# 224x224 crops from 1024x768 + rotation and horizontal flips
# +
input_shape = (224,224,3)
loss = categorical_crossentropy
data_path = data_root + 'images/1024x768'
model_name = 'DeepRN'
model = resnet101.resnet101_model(input_shape = input_shape,
weights_root = 'cnn_finetune/imagenet_models',
include_top = True,
num_classes = 5)
pre = resnet101.preprocess_input_resnet101
# the performance is higher without the rotation augmentation
process_fn = lambda im: pre(iu.ImageAugmenter(im, remap=False).crop((224,224)).fliplr().rotate(5).result)
gen_params = dict(batch_size = 64,
data_path = data_path,
fixed_batches = True,
input_shape = input_shape,
process_fn = process_fn,
outputs = output_classes)
helper = ModelHelper(model, model_name, ids, verbose = False,
loss = categorical_crossentropy,
optimizer = keras.optimizers.SGD(lr = 0.01,
momentum = 0.9,
decay = 5e-4),
metrics = ["MAE",'accuracy'],
monitor_metric = 'val_loss', monitor_mode = 'min',
early_stop_patience = 20,
class_weights = class_weights,
multiproc = True, workers = 3,
logs_root = aux_root + 'logs/koniq',
models_root = data_root + 'models/',
gen_params = gen_params)
helper.set_trainable(index=81)
print 'First trainable layer:', helper.model.layers[81].name
print 'Model name:', helper.model_name()
# +
# x,_ = helper.test_generator()
# iu.view_stack(gen.mapmm(x[0]), figsize=(5,5))
# -
LR = 0.0001 # initial
for i in range(3):
print 'Iteration', i
print 'LR =', LR
helper.train(lr=LR, epochs=100)
LR /= 10
helper.load_model()
valid_gen = helper.make_generator(ids[ids.set=='validation'],
deterministic=True)
print 'Accuracy:', helper.model.evaluate_generator(valid_gen)[2]
apps.test_rating_model(helper, groups=1);
# ### Change model to extract features
# * Huber-loss
# * allow any resolution input
# * only horizontal flip augmentation
# +
input_shape = (768,1024,3)
loss = K.tf.losses.huber_loss
data_path = data_root + 'images/1024x768/'
model = resnet101.resnet101_model(input_shape = (None, None, 3),
weights_root = 'cnn_finetune/imagenet_models',
include_top = False,
num_classes = 5)
pre = resnet101.preprocess_input_resnet101
process_fn = lambda im: pre(iu.ImageAugmenter(im).fliplr().result)
gen_params = dict(batch_size = 2,
data_path = data_path,
fixed_batches = True,
input_shape = input_shape,
process_fn = process_fn,
inputs = 'image_name',
outputs = ('c1','c2','c3','c4','c5'))
helper = ModelHelper(model, model_name, ids,
verbose = True,
logs_root = aux_root + 'logs/koniq',
models_root = data_root + 'models/',
features_root= data_root + 'features/',
gen_params = gen_params)
name = 'DeepRN/bsz64_i1[224,224,3]_lcategori_o1[5]'
helper.load_model(name, by_name=True)
# +
# x,_ = helper.test_generator()
# iu.view_stack(gen.mapmm(x[0]))
# -
# ### Change model head from GAP to SPP
# (for feature extraction)
# +
from layers.SpatialPyramidPooling import SpatialPyramidPooling
# helper.model.summary()
gap_input = helper.model.layers[-2].output
feats = SpatialPyramidPooling([3], name='SPP')(gap_input)
model_spp = Model(inputs=helper.model.input, outputs=feats)
helper.model = model_spp
# -
helper.save_activations(ids=ids, verbose=True, output_layer='SPP',
groups=2, over_write=True)
# ### Train on features
# +
input_shape = (768,1024)
input_size = 2048 * 9
features_path = data_root + 'features/DeepRN/i1[768,1024,3]_lSPP_o1[2048]_r2.h5'
fc1_size = 4096
input_feats = keras.layers.Input(shape=(input_size,), dtype='float32')
# works better if NOT normalizing the features
# norm_feats = Lambda(lambda x: K.tf.nn.l2_normalize(x,1))(input_feats)
pred = apps.fc_layers(input_feats,
name = 'main',
fc_sizes = [4096,4096,4096, 5],
dropout_rates = [0.5, 0.5, 0.5, 0],
batch_norm = 0,
out_activation = 'softmax')
model = keras.models.Model(inputs=input_feats, outputs=pred)
root_name = 'DeepRN_final'
gen_params = dict(batch_size = 128,
data_path = features_path,
process_fn = None,
input_shape = (input_size,),
inputs = ('image_name',),
outputs = ('c1','c2','c3','c4','c5'),
random_group= True)
loss = ops.make_loss(K.tf.losses.huber_loss, delta=1./9)
# works better if using the Huber loss with default delta=1
# loss = K.tf.losses.huber_loss
helper = ModelHelper(model, root_name, ids,
loss = loss,
optimizer = keras.optimizers.SGD(lr = 0.001,
momentum = 0.9,
decay = 4e-4),
metrics=['MAE'],
monitor_metric='val_loss', monitor_mode='min',
early_stop_patience = 20, workers = 10,
logs_root = aux_root+'/logs/koniq',
models_root= data_root+'/models',
gen_params =gen_params)
helper.model_name.update(imsz=list(input_shape))
print helper.model_name()
# -
LR = 0.01
for i in range(4):
print 'Iteration', i
print 'LR =', LR
helper.train(lr=LR, epochs=200)
LR /= 10
helper.load_model()
apps.test_rating_model(helper, groups=2);
| koniq/train_deeprn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **CSE 7324 Lab 3: Extending Logistic Regression**
# ### *<NAME>, <NAME>, <NAME> and <NAME>*
# ------
# ### **1. Preparation and Overview**
# ------
# #### 1.1 Business Understanding
# ---
# Austin Animal Center is the largest no-kill shelter in the United States and provides shelter to more than 16,000 animals each year. As a no-kill shelter they refuse to euthanize any animal unless the animal has a terminal medical issue and is in pain or if the animal is a danger to the public or to the shelter staff. Although the shelter’s primary goal is to find ‘forever homes’ for each and every animal that comes through their doors, many animals end up staying in the shelter for a long time if they are not considered as desirable for adoption as other animals. In addition to adopting out animals, the Austin Animal Center partners with various other rescues and animal sanctuaries to try to find homes for their animals.
#
# The average annual cost per animal at the Austin Animal Center is approximately $715 [3] and with many animals staying at the facility for long periods of time, some for several years, the cost can add up quickly. The shelter has fixed financial support via legislation to cover costs for staffing the shelters and a few grants to cover veterinary staff and services, but the shelter primarily relies on donations to provide for food, bedding and toys for the animals. The shelter must try to minimize costs associated with each animal and try to have the animals leave the shelter through adoption or transfer to a sanctuary as quickly as possible.
#
# The Austin Animal Center keeps track of each animal that comes through their doors and keeps a record of the animal’s outcome; that is whether they were adopted, transferred to a partner shelter or sanctuary or one of many other outcomes. If the shelter could predict an animal’s outcome based on the animal’s characteristics, they could be much more efficient with having animals leave the shelter by knowing which animals they should be able to adopt out and which animals they should transfer to other shelters or sanctuaries. This added efficiency would result in the shelter’s ability to take in more animals which in return would lower the average cost per animal.
#
# This lab examines the Austin Animal Center animal outcome data set to specifically look at cats and the outcome of each cat and attempts to build an accurate model of predicting the outcome. If accurate, this model could serve the Austin Animal Center as well as other cities that are looking at issuing a no-kill ordinance for their shelters.
# #### 1.2 Data Preparation
# ---
# dependencies
import pandas as pd
import numpy as np
import missingno as msno
import matplotlib.pyplot as plt
import re
from sklearn.model_selection import train_test_split
pd.set_option('display.max_columns', 500)
from textwrap import wrap
from sklearn.preprocessing import StandardScaler
import warnings
warnings.filterwarnings("ignore")
import math
# %matplotlib inline
# +
# import data
shelter_outcomes = pd.read_csv("C:/Users/w47518657u/OneDrive/SMU Spring 2019/CSE 7318/Labs/Lab Three/aac_shelter_outcomes.csv")
#shelter_outcomes = pd.read_csv("C:/Users/w47518657u/OneDrive/SMU Spring 2019/CSE 7318/Labs/Lab Three/aac_shelter_outcomes.csv")
# filter animal type for just cats
cats = shelter_outcomes[shelter_outcomes['animal_type'] == 'Cat']
#print(cats.head())
# remove age_upon_outcome and recalculate to standard units (days)
age = cats.loc[:,['datetime', 'date_of_birth']]
# convert to datetime
age.loc[:,'datetime'] = pd.to_datetime(age['datetime'])
age.loc[:,'date_of_birth'] = pd.to_datetime(age['date_of_birth'])
# calculate cat age in days
cats.loc[:,'age'] = (age.loc[:,'datetime'] - age.loc[:,'date_of_birth']).dt.days
# get dob info
cats['dob_month'] = age.loc[:, 'date_of_birth'].dt.month
cats['dob_day'] = age.loc[:, 'date_of_birth'].dt.day
cats['dob_dayofweek'] = age.loc[:, 'date_of_birth'].dt.dayofweek
# get month from datetime
cats['month'] = age.loc[:,'datetime'].dt.month
# get day of month
cats['day'] = age.loc[:,'datetime'].dt.day
# get day of week
cats['dayofweek'] = age.loc[:, 'datetime'].dt.dayofweek
# get hour of day
cats['hour'] = age.loc[:, 'datetime'].dt.hour
# get quarter
cats['quarter'] = age.loc[:, 'datetime'].dt.quarter
# clean up breed attribute
# get breed attribute for processing
# convert to lowercase, remove mix and strip whitespace
# remove space in 'medium hair' to match 'longhair' and 'shorthair'
# split on either space or '/'
breed = cats.loc[:, 'breed'].str.lower().str.replace('mix', '').str.replace('medium hair', 'mediumhair').str.strip().str.split('/', expand=True)
cats['breed'] = breed[0]
cats['breed1'] = breed[1]
# clean up color attribute
# convert to lowercase
# strip spaces
# split on '/'
color = cats.loc[:, 'color'].str.lower().str.strip().str.split('/', expand=True)
cats['color'] = color[0]
cats['color1'] = color[1]
# clean up sex_upon_outcome
sex = cats['sex_upon_outcome'].str.lower().str.strip().str.split(' ', expand=True)
sex[0].replace('spayed', True, inplace=True)
sex[0].replace('neutered', True, inplace=True)
sex[0].replace('intact', False, inplace=True)
sex[1].replace(np.nan, 'unknown', inplace=True)
cats['spayed_neutered'] = sex[0]
cats['sex'] = sex[1]
# add in domesticated attribute
cats['domestic'] = np.where(cats['breed'].str.contains('domestic'), 1, 0)
# combine outcome and outcome subtype into a single attribute
cats['outcome_subtype'] = cats['outcome_subtype'].str.lower().str.replace(' ', '-').fillna('unknown')
cats['outcome_type'] = cats['outcome_type'].str.lower().str.replace(' ', '-').fillna('unknown')
cats['outcome'] = cats['outcome_type'] + '_' + cats['outcome_subtype']
# drop unnecessary columns
cats.drop(columns=['animal_id', 'name', 'animal_type', 'age_upon_outcome', 'date_of_birth', 'datetime', 'monthyear', 'sex_upon_outcome', 'outcome_subtype', 'outcome_type'], inplace=True)
#print(cats['outcome'].value_counts())
cats.head()
# -
# Not all information included in this data set is necessary to the targeted prediction of outcome type. Some animals that were adopted were returned to the shelter as runaways before being returned to their owners. These instances have no impact on trying to predict outcome and will be removed from the data set.
# #### 1.3 Data Description
# ---
# +
print("Default datatypes of shelter cat outcomes:\n")
print(cats.dtypes)
print("\nBelow is a description of the attributes in the cats dataframe:\n")
# -
# Attribute | Description | Scale | Datatype
# --- | --- | --- | ---
# Breed | Primary breed of the cat | Nominal | Object
# Color | Primary color of the cat | Nominal | Object
# Age | Age of cat in days | Ordinal | int64
# DOB_Month | Date of birth month (1-12) for the cat | Ordinal | int64
# DOB_Day | Date of birth day (1-31) for the cat | Ordinal | int64
# DOB_DayOfWeek | Date of birth day of week (1-7) for the cat | Ordinal | int64
# Month | Month (1-12) of the outcome | Ordinal | int64
# Day | Day of month (1-31) of the outcome | Ordinal | int64
# DayOfWeek | Day of week (1-7) of the outcome | Ordinal | int64
# Hour | Hour during the day (0-23) of the outcome | Ordinal | int64
# Quarter | Quarter during the year (1-4) of the outcome | Ordinal | int64
# Breed1 | Secondary breed of the cat | Nominal | Object
# Color1 | Secondary color of the cat | Nominal | Object
# Spayed_Neutered | Is the cat spayed/netured or not | Nominal | bool
# Sex | Sex of the cat | Nominal | bool
# Domestic | Is the cat domesticated | Nominal | bool
# Outcome | The outcome of the animal | nominal | object
print('Below is a listing of the target classes and their distributions:')
cats['outcome'].value_counts()
# Each feature has a different count, a low count per feature decrease the accuracy and the efficiency of the logistic regression method used, so all features with low count was not taken into account in traning the classfier
# examine missing data
msno.matrix(cats)
# Since the missing data shows that breed1 will have little impact on the prediction since there are only two records that have a value, it will be removed from the data set. The missing data in color1 should be handled when one hot encoding is performed on it.
# #### 1.4 One hot encoding of data and splitting into training and testing sets
# +
#cats.drop(columns=['breed1'], inplace=True)
# Breed, Color, Color1, Spayed_Netured and Sex attributes need to be one hot encoded
cats_ohe = pd.get_dummies(cats, columns=['breed', 'color', 'color1', 'spayed_neutered', 'sex'])
cats_ohe.head()
out_t={'euthanasia_suffering' : 0, 'died_in-kennel' : 0, 'return-to-owner_unknown' : 0, 'transfer_partner' : 1, 'euthanasia_at-vet' : 2, 'adoption_foster' : 3, 'died_in-foster' : 0, 'transfer_scrp' : 4, 'euthanasia_medical' : 0, 'transfer_snr' : 0, 'died_enroute' : 0, 'rto-adopt_unknown' : 0, 'missing_in-foster' : 0, 'adoption_offsite' : 0, 'adoption_unknown' :5,'euthanasia_rabies-risk' : 0, 'unknown_unknown' : 0, 'adoption_barn' : 0, 'died_unknown' : 0, 'died_in-surgery' : 0, 'euthanasia_aggressive' : 0, 'euthanasia_unknown' : 0, 'missing_unknown' : 0, 'missing_in-kennel' : 0, 'missing_possible-theft' : 0, 'died_at-vet' : 0, 'disposal_unknown' : 0, 'euthanasia_underage' : 0, 'transfer_barn' : 0}
#output is converted from string to catogries 0 to 5 represent each output
# separate outcome from data
outcome = cats_ohe['outcome']
cats_ohe.drop(columns=['outcome'])
print(cats_ohe.head())
# split the data
X_train, X_test, y_train, y_test = train_test_split(cats_ohe, outcome, test_size=0.2, random_state=0)
X_train.drop(columns=['outcome'], inplace=True)
y_train = [out_t[item] for item in y_train]
#print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
# -
# One hot encoding is used for the cat breed, color, spayed/neutered and sex attributes to convert the categorical variables into a form that should play nicer with logistic regression. Although spayed_neutered and sex are essentially boolean attributes, they had to be converted because there were many unknown values in each.
#
# The data is split with an 80/20 train/test ratio using the train_test_split function in the cross validation functions in Skikit Learn's cross validation package. Although this was an easy method to split the data into training and test sets, it was not a good way to split the data for this dataset. As shown above, the target distribution is skewed and some targets have very few instances. It would have been better to select an 80/20 ratio for each class.
# ### **2. Modeling**
# ------
# +
import numpy as np
pd.set_option('display.max_columns', 5)
class BinaryLogisticRegressionBase:
# private:
def __init__(self, eta, iterations, C,reg):
self.eta = eta
self.iters = iterations
self.C=C
self.reg=reg
# internally we will store the weights as self.w_ to keep with sklearn conventions
def __str__(self):
return 'Base Binary Logistic Regression Object, Not Trainable'
# convenience, private and static:
@staticmethod
def _sigmoid(theta):
return 1/(1+np.exp(-theta))
@staticmethod
def _add_bias(X):
return np.hstack((np.ones((X.shape[0],1)),X)) # add bias term
# public:
def predict_proba(self,X,add_bias=True):
# add bias term if requested
Xb = self._add_bias(X) if add_bias else X
return self._sigmoid(Xb @ self.w_) # return the probability y=1
def predict(self,X):
return (self.predict_proba(X)>0.5) #return the actual prediction
# inherit from base class
class BinaryLogisticRegression(BinaryLogisticRegressionBase):
#private:
def __str__(self):
if(hasattr(self,'w_')):
return 'Binary Logistic Regression Object with coefficients:\n'+ str(self.w_) # is we have trained the object
else:
return 'Untrained Binary Logistic Regression Object'
def _get_gradient(self,X,y):
# programming \sum_i (yi-g(xi))xi
gradient = np.zeros(self.w_.shape) # set gradient to zero
for (xi,yi) in zip(X,y):
# the actual update inside of sum
gradi = (yi - self.predict_proba(xi,add_bias=False))*xi
# reshape to be column vector and add to gradient
gradient += gradi.reshape(self.w_.shape)
return gradient/float(len(y))
# public:
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = self._get_gradient(Xb,y)
self.w_ += gradient*self.eta # multiply by learning rate
import numpy as np
from scipy.special import expit
class VectorBinaryLogisticRegression(BinaryLogisticRegression):
# inherit from our previous class to get same functionality
@staticmethod
def _sigmoid(theta):
# increase stability, redefine sigmoid operation
return expit(theta) #1/(1+np.exp(-theta))
# but overwrite the gradient calculation
def _get_gradient(self,X,y):
ydiff = y-self.predict_proba(X,add_bias=False).ravel() # get y difference
gradient = np.mean(X * ydiff[:,np.newaxis], axis=0) # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
if self.reg=='L2':
gradient[1:] += -2 * self.w_[1:] * self.C
if self.reg=='L1':
gradient[1:] += -self.C # the deravtiv of C*abs(W), which should be dx(abs(w))= w/abs(w)
if self.reg=='L1L2':
gradient[1:] += -self.C-2 * self.w_[1:] * self.C
if self.reg=='none':
gradient[1:]
return gradient
from scipy.optimize import minimize_scalar
import copy
class LineSearchLogisticRegression(VectorBinaryLogisticRegression):
# define custom line search for problem
@staticmethod
def objective_function(eta,X,y,w,grad,C=0.001):
wnew = w - grad*eta
g = expit(X @ wnew)
return -np.sum(np.log(g[y==1]))-np.sum(np.log(1-g[y==0])) + C*sum(wnew**2)
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = -self._get_gradient(Xb,y)
# minimization inopposite direction
# do line search in gradient direction, using scipy function
opts = {'maxiter':self.iters/50} # unclear exactly what this should be
res = minimize_scalar(self.objective_function, # objective function to optimize
bounds=(self.eta/1000,self.eta*10), #bounds to optimize
args=(Xb,y,self.w_,gradient,0.001), # additional argument for objective function
method='bounded', # bounded optimization for speed
options=opts) # set max iterations
eta = res.x # get optimal learning rate
self.w_ -= gradient*eta # set new function values
# subtract to minimize
class StochasticLogisticRegression(BinaryLogisticRegression):
# stochastic gradient calculation
def _get_gradient(self,X,y):
idx = int(np.random.rand()*len(y)) # grab random instance
ydiff = y[idx]-self.predict_proba(X[idx],add_bias=False) # get y difference (now scalar)
gradient = X[idx] * ydiff[:,np.newaxis] # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
if self.reg=='L2':
gradient[1:] += -2 * self.w_[1:] * self.C
if self.reg=='L1':
gradient[1:] += -self.C # the deravtiv of C*abs(W), which should be dx(abs(w))= w/abs(w)
if self.reg=='L1L2':
gradient[1:] += -self.C-(2 * self.w_[1:] * self.C)
if self.reg=='none':
gradient[1:]
return gradient
from scipy.optimize import fmin_bfgs
class BFGSBinaryLogisticRegression(BinaryLogisticRegression):
@staticmethod
def objective_function(w,X,y,C,reg):
g = expit(X @ w)
return -np.sum(np.log(g[y==1]))-np.sum(np.log(1-g[y==0])) + C*sum(w**2) #-np.sum(y*np.log(g)+(1-y)*np.log(1-g))
@staticmethod
def objective_gradient(w,X,y,C,reg):
g = expit(X @ w)
ydiff = y-g # get y difference
gradient = np.mean(X * ydiff[:,np.newaxis], axis=0)
gradient = gradient.reshape(w.shape)
if reg=='L2':
gradient[1:] += -2 * w[1:] * C
if reg=='L1':
gradient[1:] += - C # the deravtiv of C*abs(W), which should be dx(abs(w))= w/abs(w)
if reg=='L1L2':
gradient[1:] +=(-2 * w[1:] * C) - C
if reg=='none':
gradient[1:]
return -gradient
# just overwrite fit function
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = fmin_bfgs(self.objective_function, # what to optimize
np.zeros((num_features,1)), # starting point
fprime=self.objective_gradient, # gradient function
args=(Xb,y,self.C,self.reg), # extra args for gradient and objective function
gtol=1e-03, # stopping criteria for gradient, |v_k|
maxiter=self.iters, # stopping criteria iterations
disp=False)
self.w_ = self.w_.reshape((num_features,1))
from numpy.linalg import pinv
class HessianBinaryLogisticRegression(BinaryLogisticRegression):
# just overwrite gradient function
def _get_gradient(self,X,y):
g = self.predict_proba(X,add_bias=False).ravel() # get sigmoid value for all classes
hessian = X.T @ np.diag(g*(1-g)) @ X - 2 * self.C # calculate the hessian
ydiff = y-g # get y difference
gradient = np.sum(X * ydiff[:,np.newaxis], axis=0) # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
if self.reg=='L2':
gradient[1:] += -2 * self.w_[1:] * self.C
if self.reg=='L1': # the deravtiv of C*abs(W), which should be dx(abs(w))= w/abs(w)
gradient[1:] += -self.C
if self.reg=='L1L2':
gradient[1:] += -self.C-2 * self.w_[1:] * self.C
if self.reg=='none':
gradient[1:]
return pinv(hessian) @ gradient
# +
from scipy.optimize import minimize_scalar
import copy
class LogisticRegression:
def __init__(self, eta, iterations,solver='leaner', C=0.001,reg='L2'):
self.eta = eta
self.iters = iterations
self.slv = solver
self.C=C
self.reg=reg
# internally we will store the weights as self.w_ to keep with sklearn conventions
def __str__(self):
if(hasattr(self,'w_')):
return 'MultiClass Logistic Regression Object with coefficients:\n'+ str(self.w_) # is we have trained the object
else:
return 'Untrained MultiClass Logistic Regression Object'
def fit(self,X,y):
num_samples, num_features = X.shape
self.unique_ = np.sort(np.unique(y)) # get each unique class value
num_unique_classes = len(self.unique_)
self.classifiers_ = [] # will fill this array with binary classifiers
for i,yval in enumerate(self.unique_): # for each unique value
y_binary = (y==yval) # create a binary problem
# train the binary classifier for this class
if self.slv=='stochastic':
slr = StochasticLogisticRegression(self.eta,self.iters,self.C,self.reg)
slr.fit(X,y_binary)
self.classifiers_.append(slr)
if self.slv=='steepest':
mls=LineSearchLogisticRegression(self.eta,self.iters,self.C,self.reg)
mls.fit(X,y_binary)
self.classifiers_.append(mls)
if self.slv=='leaner':
blr = VectorBinaryLogisticRegression(self.eta,self.iters,self.reg)
blr.fit(X,y_binary)
self.classifiers_.append(blr)
if self.slv=='BFGS':
bfgslr = BFGSBinaryLogisticRegression(self.eta,self.iters,self.C,self.reg)
bfgslr.fit(X,y_binary)
self.classifiers_.append(bfgslr)
if self.slv=='newton':
newt = HessianBinaryLogisticRegression(self.eta,self.iters,self.C,self.reg)
newt.fit(X,y_binary)
self.classifiers_.append(newt)
# add the trained classifier to the list
# save all the weights into one matrix, separate column for each class
self.w_ = np.hstack([x.w_ for x in self.classifiers_]).T
def predict_proba(self,X):
probs = []
for blr in self.classifiers_:
probs.append(blr.predict_proba(X)) # get probability for each classifier
return np.hstack(probs) # make into single matrix
def predict(self,X):
return np.argmax(self.predict_proba(X),axis=1) # take argmax along row
# -
pd.set_option('display.max_rows', 10)
pd.set_option('display.max_columns', 10)
pd.set_option('display.width', 100)
pd.set_option('max_colwidth', 300)
pd.set_option('display.expand_frame_repr', True)
np.set_printoptions(threshold=5)
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.01,1000,'stochastic',1,'L1')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
stoc1=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
#y_target_ar=y_train_b.transfer_partner.values
#y_target_ar=y_train_b_v
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.01,1000,'stochastic',.0001,'L1L2')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
stoc2=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.1,50,'steepest',10,'L2')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
steep=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.1,50,'steepest',.0001,'L2')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
steep1=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.1,10,'BFGS',.0001,'L2')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
BFGS1=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.1,10,'BFGS',10,'L2')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
BFGS2=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.1,3,'newton',.0001,'L1')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
newton1=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
# %%time
from sklearn.metrics import accuracy_score
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr = LogisticRegression(.01,3,'newton',.0001,'L1')
lr.fit(x_train_ar,y_target_ar)
print(lr)
yhat = lr.predict(x_train_ar)
newton2=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
import numpy as np
import matplotlib.pyplot as plt
from textwrap import wrap
para=['1-ata=.01,iter=1000, type= stochastic, C=.0001,L1L2 ','2-ata=.1,iter=1000, type= stochastic, C=.01,L1L2 ','ata=.1,iter=50, type= steepest, C=10,L2 ','ata=.1,iter=50, type= steepest, C=.0001,L2 ','ata=.1,iter=10,type=BFGS,C=.001,L2','ata=.1,iter=10,type=BFGS,C=10,L2','ata=.01, iter=3, type=newton, C=.0001, L2','ata=.01,iter=10,type=newton,C=.0001,L1' ]
acc=[stoc1,stoc2,steep,steep1,BFGS1, BFGS2,newton1,newton2]
plt.subplots(figsize=(17, 7))
x=[0,1,2,3,4,5,6,7]
z=np.polyfit(x, acc, 1)
labels = [ '\n'.join(wrap(l, 18)) for l in para ]
labels = [ '\n'.join(wrap(l, 18)) for l in para ]
plt.xlabel('Optimization', fontweight='bold')
plt.ylabel('Accuracy', fontweight='bold')
p = np.poly1d(z)
plt.bar(labels,acc)
# -
# Since the time consumption for newton technique is long (over 2 minutes) and newton technique is not the best optimization technique for logistic regression, we decided not to include it in the following graph.
# +
import numpy as np
import matplotlib.pyplot as plt
from textwrap import wrap
# set width of bar
barWidth = 0.45
# set height of bar
bars1 = [stoc1,stoc2,steep,steep1,BFGS1, BFGS2]
bars2 = [.013,.0129,.27,.21,.05,.32]
x=[0,1,2,3,4,5]
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
#r3 = [x + barWidth for x in r2]
plt.subplots(figsize=(17, 7))
# Make the plot
plt.bar(r1, bars1, color='#7f6d5f', width=barWidth, edgecolor='white', label='accuracy')
plt.bar(r2, bars2, color='#557f2d', width=barWidth, edgecolor='white', label='time (m)')
plt.plot(x,p(x),color='black')
# Add xticks on the middle of the group bars
plt.xlabel('Optimization', fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], labels)
# Create legend & Show graphic
plt.legend()
plt.show()
# -
# ### Observations:
#
# 1-the figures above shows the accuracy and time consumption for various optimization techniques, BFGS with 10 iterations has the highest accuracy with a low delay and L2 regulation.
#
# 2- stochastic has different accuracy values for different runs with the same parameters which mean the logistic regression got stuck in a local minimum depending on the initial value of the gradient and the direction
#
# 3- steepest decent and most of the other optimization techniques returned a better accuracy for a lower C which means stronger regularization prevent from overfitting the data and in return has a better performance
#
# 4- BFGS did better than hessian in terms of accuracy and time consumption which means that hessian is highly computational epically in calculating the second order derivative and inverting the hessian matrix
#
# 5- the output classification had a count per classifier shown in the figure above in the data understanding section all the low count have been given the same class because there is not enough data to train a classifier
#
# 6- the data is preprocessed such that its distribution will have a mean value 0 and standard deviation of 1. Given the distribution of the data, each value in the dataset will have the sample mean value subtracted, and then divided by the standard deviation of the whole dataset.
#
# ### parameters justification
#
# parameters that resulted in the best accuracy and time consuption was chosen useing try and erorr , we started on initial parameters based on professor's suggestions
#
# Using trial and error to obtain optimized parameters for classification does not seem to be "data snooping" of the negative kind in the typical sense. Data snooping is the "misuse of data analysis to find patterns in data that can be presented as statistically significant when in fact there is no real underlying effect." (https://en.wikipedia.org/wiki/Data_dredging) In our case, we are pursuing optimal parameters that enable the highest accuracy classification possible. No matter what methods we use to obtain a model that accurately predicts classes for training data, it does not matter if the same is not also true for new, untrained data. If our classification algorithm is negatively impacted by our tweaking of parameters for new instances of data, then the tweaking of these parameters would be unjustified.
#
# However, since our goal is only to build the best classification tool possible, any means that improve that capability would be permissible, including adjusting parameters such as the regularization term - assuming they contribute to an increase in overall classification performance for untrained data.
#
#
# Based on the observations above, the best best logistic regression optimization technique is BFGS. So we will use BFGS to compare with LBFGS from sklearn.
# +
# %%time
from sklearn.linear_model import LogisticRegression
lr_sk = LogisticRegression(solver='lbfgs',n_jobs=2,C=.0001, max_iter=10)
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
x_train_ar = StandardScaler().fit(x_train_ar).transform(x_train_ar)
lr_sk.fit(x_train_ar,y_target_ar)
print(np.hstack((lr_sk.intercept_[:,np.newaxis],lr_sk.coef_)))
yhat = lr_sk.predict(x_train_ar)
newtsk=accuracy_score(y_target_ar,yhat)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
# +
para=['SK learn, iter=10,type=lbfgs,C=.0001, L2','ata=.1,iter=10,type=BFGS,C=.0001,L2' ]
x=[0,1]
acc=[ newtsk,BFGS1]
time=[1.5, 1.6]
plt.subplots(figsize=(10, 7))
z=np.polyfit(x, acc, 1)
labels = [ '\n'.join(wrap(l, 18)) for l in para ]
plt.bar(labels,acc, .4)
#ax.bar(labels,time,width=0.2,color='g')
plt.xlabel('Optimization', fontweight='bold')
plt.ylabel('Accuracy', fontweight='bold')
p = np.poly1d(z)
plt.plot(x,1.01*p(x),color='black')
# -
# BFGS and steepest decend optimization Techniques resulted in a better accuracy than SK learning BFGS for the same itration, as shown from the blake trend line.
#
# In the next plot, time will be introduced.
#
# +
import numpy as np
import matplotlib.pyplot as plt
# set width of bar
barWidth = 0.25
# set height of bar
bars1 = [newtsk, BFGS1]
bars2 = [.09,.05]
#bars3 = [29, 3, 24, 25, 17]
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
#r3 = [x + barWidth for x in r2]
plt.subplots(figsize=(10, 7))
# Make the plot
plt.bar(r1, bars1, color='#7f6d5f', width=barWidth, edgecolor='white', label='accuracy')
plt.bar(r2, bars2, color='#557f2d', width=barWidth, edgecolor='white', label='time (m)')
plt.plot(x,1.01*p(x),color='black')
# Add xticks on the middle of the group bars
plt.xlabel('Optimization', fontweight='bold')
plt.xticks([r + barWidth for r in range(len(bars1))], labels)
# Create legend & Show graphic
plt.legend()
plt.show()
# -
# ### **3. Deployment**
# ------
#
# Among all the techniques we tested above, BFGS is the best optimization technique for logistic regression. Comparing BFGS to LBFGS from the sklearn, we see that BFGS has higher accuracy with lower time consumption (2.99s vs. 13.9s)
#
# In our opinion,the best method to use is BFGS method since it produces the most accuracy and low time consumption.
#
# Type Markdown and LaTeX: 𝛼2
# ### **4. Optimization Using Mean Squared Error**
# ------
# +
# %%time
# from last time, our logistic regression algorithm is given by (including everything we previously had):
class BinaryLogisticRegressionForMSE:
def __init__(self, eta, iterations=1, C=0.001):
self.eta = eta
self.iters = iterations
self.C = C
# internally we will store the weights as self.w_ to keep with sklearn conventions
def __str__(self):
if(hasattr(self,'w_')):
return 'Binary Logistic Regression Object with coefficients:\n'+ str(self.w_) # is we have trained the object
else:
return 'Untrained Binary Logistic Regression Object'
# convenience, private:
@staticmethod
def _add_bias(X):
return np.hstack((np.ones((X.shape[0],1)),X)) # add bias term
@staticmethod
def _sigmoid(theta):
# increase stability, redefine sigmoid operation
return expit(theta) #1/(1+np.exp(-theta))
# vectorized gradient calculation with regularization using L2 Norm
def _get_gradient(self,X,y):
ydiff = y-self.predict_proba(X,add_bias=False).ravel() # get y difference
gradient = np.mean(X * ydiff[:,np.newaxis], axis=0) # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
gradient[1:] += -2 * self.w_[1:] * self.C
return gradient
# public:
def predict_proba(self,X,add_bias=True):
# add bias term if requested
Xb = self._add_bias(X) if add_bias else X
return self._sigmoid(Xb @ self.w_) # return the probability y=1
def predict(self,X):
return (self.predict_proba(X)>0.5) #return the actual prediction
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = self._get_gradient(Xb,y)
self.w_ += gradient*self.eta # multiply by learning rate
blr = BinaryLogisticRegressionForMSE(eta=0.1,iterations=500,C=0.001)
x_train_ar=X_train.values
y_target_ar=np.asarray(y_train)
blr.fit(x_train_ar,y_target_ar)
print(blr)
yhat = blr.predict(x_train_ar)
print('Accuracy of: ',accuracy_score(y_target_ar,yhat))
class LineSearchLogisticRegressionWithMSE(BinaryLogisticRegressionForMSE):
# define custom line search for problem
@staticmethod
def objective_function(eta,X,y,w,grad,C=0.001):
wnew = w - grad * eta # subtract grad*eta.. from class # 02.21.19 - 10.m4v timestamp: 23:00
yhat = (1/(1+np.exp(-X @ wnew))) >0.5
return np.mean((y-yhat)**2) + C*np.mean(wnew**2) # add regularization term, don't subtract.. from class
# 02.21.19 - 10.m4v timestamp: 17:40
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = -self._get_gradient(Xb,y)
# minimization inopposite direction
# do line search in gradient direction, using scipy function
opts = {'maxiter':self.iters} # unclear exactly what this should be
res = minimize_scalar(self.objective_function, # objective function to optimize
bounds=(self.eta/1000,self.eta*10), #bounds to optimize
args=(Xb,y,self.w_,gradient,0.001), # additional argument for objective function
method='bounded', # bounded optimization for speed
options=opts) # set max iterations
eta = res.x # get optimal learning rate
self.w_ -= gradient*eta # set new function values
# subtract to minimize
# -
# ### **5. References**
# ------
# 1. Austin Animal Center Shelter Outcomes (Kaggle) https://www.kaggle.com/aaronschlegel/austin-animal-center-shelter-outcomes-and/version/1#aac_shelter_outcomes.csv
# 2. Austin Animal Center. (n.d.). Retrieved March 10, 2019, from http://www.austintexas.gov/department/aac
# 3. <NAME>; <NAME>; <NAME>; <NAME>; and <NAME>, "Legislating Components of a Humane City: The Economic Impacts of the Austin, Texas "No Kill" Resolution (City of Austin Resolution 20091105-040)" (2017). Animal Law and Legislation. 1.
# https://animalstudiesrepository.org/anilleg/1
| Project3/Project3.final(modified2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introducing fastpages
# > An easy to use blogging platform with extra features for <a href="https://jupyter.org/">Jupyter Notebooks</a>.
#
# - toc: true
# - badges: true
# - comments: true
# - sticky_rank: 1
# - author: <NAME> & <NAME>
# - image: images/diagram.png
# - categories: [fastpages, jupyter]
# 
#
# We are very pleased to announce the immediate availability of [fastpages](https://github.com/fastai/fastpages). `fastpages` is a platform which allows you to create and host a blog for free, with no ads and many useful features, such as:
#
# - Create posts containing code, outputs of code (which can be interactive), formatted text, etc directly from [Jupyter Notebooks](https://jupyter.org/); for instance see this great [example post](https://drscotthawley.github.io/devblog3/2019/02/08/My-1st-NN-Part-3-Multi-Layer-and-Backprop.html) from <NAME>ley. Notebook posts support features such as:
# - Interactive visualizations made with [Altair](https://altair-viz.github.io/) remain interactive.
# - Hide or show cell input and output.
# - Collapsable code cells that are either open or closed by default.
# - Define the Title, Summary and other metadata via a special markdown cells
# - Ability to add links to [Colab](https://colab.research.google.com/) and GitHub automatically.
# - Create posts, including formatting and images, directly from Microsoft Word documents.
# - Create and edit [Markdown](https://guides.github.com/features/mastering-markdown/) posts entirely online using GitHub's built-in markdown editor.
# - Embed Twitter cards and YouTube videos.
# - Categorization of blog posts by user-supplied tags for discoverability.
# - ... and [much more](https://github.com/fastai/fastpages)
#
# [fastpages](https://github.com/fastai/fastpages) relies on Github pages for hosting, and [Github Actions](https://github.com/features/actions) to automate the creation of your blog. The setup takes around three minutes, and does not require any technical knowledge or expertise. Due to built-in automation of fastpages, you don't have to fuss with conversion scripts. All you have to do is save your Jupyter notebook, Word document or markdown file into a specified directory and the rest happens automatically. Infact, this blog post is written in a Jupyter notebook, which you can see with the "View on GitHub" link above.
#
# [fast.ai](https://www.fast.ai/) have previously released a similar project called [fast_template](https://www.fast.ai/2020/01/16/fast_template/), which is even easier to set up, but does not support automatic creation of posts from Microsoft Word or Jupyter notebooks, including many of the features outlined above.
#
# **Because `fastpages` is more flexible and extensible, we recommend using it where possible.** `fast_template` may be a better option for getting folks blogging who have no technical expertise at all, and will only be creating posts using Github's integrated online editor.
# ## Setting Up Fastpages
#
# [The setup process](https://github.com/fastai/fastpages#setup-instructions) of fastpages is automated with GitHub Actions, too! Upon creating a repo from the fastpages template, a pull request will automatically be opened (after ~ 30 seconds) configuring your blog so it can start working. The automated pull request will greet you with instructions like this:
#
# 
#
# All you have to do is follow these instructions (in the PR you receive) and your new blogging site will be up and running!
# ## Jupyter Notebooks & Fastpages
#
# In this post, we will cover special features that fastpages provides for Jupyter notebooks. You can also write your blog posts with Word documents or markdown in fastpages, which contain many, but not all the same features.
# ### Options via FrontMatter
#
# The first cell in your Jupyter Notebook or markdown blog post contains front matter. Front matter is metadata that can turn on/off options in your Notebook. It is formatted like this:
#
# ```
# # Title
# > Awesome summary
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME> & <NAME>
# - categories: [fastpages, jupyter]
# ```
#
# **All of the above settings are enabled in this post, so you can see what they look like!**
#
# - the summary field (preceeded by `>`) will be displayed under your title, and will also be used by social media to display as the description of your page.
# - `toc`: setting this to `true` will automatically generate a table of contents
# - `badges`: setting this to `true` will display Google Colab and GitHub links on your blog post.
# - `comments`: setting this to `true` will enable comments. See [these instructions](https://github.com/fastai/fastpages#enabling-comments) for more details.
# - `author` this will display the authors names.
# - `categories` will allow your post to be categorized on a "Tags" page, where readers can browse your post by categories.
#
#
# _Markdown front matter is formatted similarly to notebooks. The differences between the two can be [viewed on the fastpages README](https://github.com/fastai/fastpages#front-matter-related-options)._
# ### Code Folding
# put a `#collapse-hide` flag at the top of any cell if you want to **hide** that cell by default, but give the reader the option to show it:
#hide
# !pip install pandas altair
#collapse-hide
import pandas as pd
import altair as alt
# put a `#collapse-show` flag at the top of any cell if you want to **show** that cell by default, but give the reader the option to hide it:
#collapse-show
cars = 'https://vega.github.io/vega-datasets/data/cars.json'
movies = 'https://vega.github.io/vega-datasets/data/movies.json'
sp500 = 'https://vega.github.io/vega-datasets/data/sp500.csv'
stocks = 'https://vega.github.io/vega-datasets/data/stocks.csv'
flights = 'https://vega.github.io/vega-datasets/data/flights-5k.json'
# If you want to completely hide cells (not just collapse them), [read these instructions](https://github.com/fastai/fastpages#hide-inputoutput-cells).
# hide
df = pd.read_json(movies) # load movies data
df.columns = [x.replace(' ', '_') for x in df.columns.values]
genres = df['Major_Genre'].unique() # get unique field values
genres = list(filter(lambda d: d is not None, genres)) # filter out None values
genres.sort() # sort alphabetically
# ### Interactive Charts With Altair
#
# Interactive visualizations made with [Altair](https://altair-viz.github.io/) remain interactive!
#
# We leave this below cell unhidden so you can enjoy a preview of syntax highlighting in fastpages, which uses the [Dracula theme](https://draculatheme.com/).
# +
# select a point for which to provide details-on-demand
label = alt.selection_single(
encodings=['x'], # limit selection to x-axis value
on='mouseover', # select on mouseover events
nearest=True, # select data point nearest the cursor
empty='none' # empty selection includes no data points
)
# define our base line chart of stock prices
base = alt.Chart().mark_line().encode(
alt.X('date:T'),
alt.Y('price:Q', scale=alt.Scale(type='log')),
alt.Color('symbol:N')
)
alt.layer(
base, # base line chart
# add a rule mark to serve as a guide line
alt.Chart().mark_rule(color='#aaa').encode(
x='date:T'
).transform_filter(label),
# add circle marks for selected time points, hide unselected points
base.mark_circle().encode(
opacity=alt.condition(label, alt.value(1), alt.value(0))
).add_selection(label),
# add white stroked text to provide a legible background for labels
base.mark_text(align='left', dx=5, dy=-5, stroke='white', strokeWidth=2).encode(
text='price:Q'
).transform_filter(label),
# add text labels for stock prices
base.mark_text(align='left', dx=5, dy=-5).encode(
text='price:Q'
).transform_filter(label),
data=stocks
).properties(
width=500,
height=400
)
# -
# ### Data Tables
#
# You can display tables per the usual way in your blog:
# display table with pandas
df[['Title', 'Worldwide_Gross',
'Production_Budget', 'IMDB_Rating']].head()
# ## Other Features
# ### GitHub Flavored Emojis
#
# Typing `I give this post two :+1:!` will render this:
#
# I give this post two :+1:!
# ### Images w/Captions
#
# You can include markdown images with captions like this:
#
# ```
# 
# ```
#
#
# 
#
# Of course, the caption is optional.
# ### Tweetcards
#
# Typing `> twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20` will render this:
#
# > twitter: https://twitter.com/jakevdp/status/1204765621767901185?s=20
# ### Youtube Videos
#
# Typing `> youtube: https://youtu.be/XfoYk_Z5AkI` will render this:
#
#
# > youtube: https://youtu.be/XfoYk_Z5AkI
# ### Boxes / Callouts
#
# Typing `> Warning: There will be no second warning!` will render this:
#
#
# > Warning: There will be no second warning!
#
#
#
# Typing `> Important: Pay attention! It's important.` will render this:
#
# > Important: Pay attention! It's important.
#
#
#
# Typing `> Tip: This is my tip.` will render this:
#
# > Tip: This is my tip.
#
#
#
# Typing `> Note: Take note of this.` will render this:
#
# > Note: Take note of this.
#
#
#
# Typing `> Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.` will render in the docs:
#
# > Note: A doc link to [an example website: fast.ai](https://www.fast.ai/) should also work fine.
# ## More Examples
#
# This [tutorial](https://fastpages.fast.ai/jupyter/2020/02/20/test.html) contains more examples of what you can do with notebooks.
# ## How fastpages Converts Notebooks to Blog Posts
# fastpages uses [nbdev](https://nbdev.fast.ai/index.html) to power the conversion process of Jupyter Notebooks to blog posts. When you save a notebook into the `/_notebooks` folder of your repository, GitHub Actions applies `nbdev` against those notebooks automatically. The same process occurs when you save Word documents or markdown files into the `_word` or `_posts` directory, respectively.
#
# We will discuss how GitHub Actions work in a follow up blog post.
# ## Resources & Next Steps
# We highly encourage you to start blogging with `fastpages`! Some resources that may be helpful:
#
# - [fastpages repo](https://github.com/fastai/fastpages) - this is where you can go to create your own fastpages blog!
# - Fastai forums - [nbdev & blogging category](https://forums.fast.ai/c/fastai-users/nbdev/). You can ask questions about fastpages here, as well as suggest new features.
# - [nbdev](https://github.com/fastai/nbdev): this project powers the conversion of Jupyter notebooks to blog posts.
#
# If you end up writing a blog post using fastpages, please let us know on Twitter: [@jeremyphoward](https://twitter.com/jeremyphoward), [@HamelHusain](https://twitter.com/hamelhusain).
| _notebooks/2020-02-21-introducing-fastpages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Export a dynamical decoupling sequence to pyQuil
#
# Q-CTRL Open Controls provides easy-to-use methods to construct dynamical decoupling sequences (DDS) according to well-known dynamical decoupling schemes. Here we show how a DDS from Q-CTRL Open Controls can be exported to a `Program` defined in pyQuil using Q-CTRL pyQuil Adapter.
#
# Note:
# * You need the [pyQuil package](http://docs.rigetti.com/en/stable/start.html) to create a quantum program.
# * You need the [ForestSDK](https://www.rigetti.com/forest) to simulate a quantum program.
# * You can the quantum program on a real quantum computer using [Rigetti QCS](https://qcs.rigetti.com/sign-in).
# ## Imports
# +
#General
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
#Q-CTRL Open Controls
from qctrlopencontrols import new_predefined_dds
#Q-CTRL pyQuil Adapter
from qctrlpyquil import convert_dds_to_pyquil_program
#Q-CTRL Visualizer
from qctrlvisualizer import plot_sequences
#pyQuil
from pyquil.api import get_qc
# -
# ## Running a DDS on a pyQuil Quantum Virtual Machine (QVM)
#
# This section demonstrates how a DDS can be prepared and a corresponding pyQuil program made and executed on a Quantum Virtual Simulator (QVM).
#
# Q-CTRL Open Controls defines a DDS as a set of instantaneous unitary operations performed at specific offset times, see the [technical documentation](https://docs.q-ctrl.com/open-controls/references/qctrl-open-controls/qctrlopencontrols/DynamicDecouplingSequence.html) for mathematical details.
#
# pyQuil implements quantum computation through `Program` that contains a series of [gates](http://docs.rigetti.com/en/stable/apidocs/gates.html). How these gates are physically implemented will depend on the device that it is run on. Rigetti's documentation gives an oversight on Rigetti's [native gates](http://docs.rigetti.com/en/stable/apidocs/gates.html#native-gates-for-rigetti-qpus) and other [physically realizable gates](http://docs.rigetti.com/en/stable/apidocs/gates.html#all-gates-and-instructions).
#
# If a user wants to add pauses (in time) during a computation they can use identity gates. However, executing a quantum program with identity gates cause the compiler to remove the gates before execution to increase efficiency. This can be avoided by using `Pragma PRESERVE` blocks (see [documentation](http://docs.rigetti.com/en/stable/basics.html#pragmas) for more detail and other usages of `Pragma`). All of $I$ (identity gate), $RX$ (X-rotation gates) and $RY$ (Y-rotation gates) take a fixed time (`gate_time`).
#
# Converting a DDS into a pyQuil program is an approximate process where the instantaneous unitaries are replaced with finite duration gates and the pauses in-between unitaries are replaced with the closest integer number of identity gates. The exact algorithm used to make this approximation is documented in the [source code](https://github.com/qctrl/python-pyquil).
#
# In this example we will define a Quadratic DDS and convert it into a program that we can later run on a simulator and on a real device. We also create a Ramsey DDS of the same duration to compare as a benchmark. For both the sequences, we add a $X_{\pi/2}$ rotation on either end of the sequence.
# ### Preparing the sequences
# +
## Quadratic sequence, total duration: 20us
quadratic_sequence = new_predefined_dds(
scheme='quadratic',
duration=5e-6,
number_inner_offsets=2,
number_outer_offsets=2,
pre_post_rotation=True,
name='Quadratic sequence')
print(quadratic_sequence)
## Ramsey sequence, total duration: 20us
ramsey_sequence = new_predefined_dds(
scheme='Ramsey',
duration=5e-6,
pre_post_rotation=True,
name='Ramsey sequence')
print(ramsey_sequence)
# -
# ### Constructing the Program using Q-CTRL Open Controls
#
# To construct a `Program` from a DDS you need to provide:
# * `dynamic_decoupling_sequence` - a DDS.
# * `target_qubits` - the qubit indices on which the DDS will be applied.
# * `gate_time` - the delay (in seconds) introduced by each of the `identity` gates.
# * (optional) `add_measurement` - to add a measurement.
#
# In this example, we will use $1$st qubit and specify the `gate_time` to be $50$ $n$s (see same [specification](http://docs.rigetti.com/en/stable/apidocs/autogen/pyquil.noise.add_decoherence_noise.html#pyquil.noise.add_decoherence_noise)).
# +
## Prepare the pyQuil related parameters
'''
target_qubits : list
A list of integers specifying the target qubits within the set of qubit registers
'''
target_qubits = [1]
'''
gate_time : float
Time delay (in seconds) introduced by identity gate
'''
gate_time = 50e-9
'''
add_measurement : bool
Indicates if the program requires a measurement step.
'''
add_measurement = True
## convert the quadratic sequence to program
quadratic_program = convert_dds_to_pyquil_program(
dynamic_decoupling_sequence=quadratic_sequence,
target_qubits=target_qubits,
gate_time=gate_time,
add_measurement=add_measurement,
)
## convert the ramsey sequence to program
ramsey_program = convert_dds_to_pyquil_program(
dynamic_decoupling_sequence=ramsey_sequence,
target_qubits=target_qubits,
gate_time=gate_time,
add_measurement=add_measurement,
)
# -
# ### Plotting the DDS
#
# We can use the Q-CTRL Python Visualizer package to plot the DDS for comparison against their pyQuil program approximations.
# #### Plotting the Quadratic Sequence
formatted_plot_data = quadratic_sequence.export()
figure = plt.figure()
plt.suptitle('Quadratic Sequence')
plot_sequences(figure, formatted_plot_data)
# #### Plotting the Ramsey sequence
formatted_plot_data = ramsey_sequence.export()
figure = plt.figure()
plt.suptitle('Ramsey Sequence')
plot_sequences(figure, formatted_plot_data)
# #### Printing the Quadratic DDS Program
#
# The Quadratic DDS has $X_{\pi/2}$ rotations at beginning and end to create and remove the superposition state that is preserved by the DDS. The $X_{\pi/2}$ rotations are added to the programs in the form of pre-post-gates that are implemented via pyQuil's $RX(\pi/2)$ gate.
#
# The $RZ(\pi)$ gates are $Z_\pi$ pulses (a $\pi$ rotation around $Z$-axis) and $RX(\pi)$ correspond to $X_{\pi}$ pulses (a $\pi$ rotation around $X$-axis).
#
# The `I` in the program corresponds to the `identity` gates. In the DDS, the first $Z_{\pi}$-pulse is applied at a delay of $0.3125$ $\mu$s. This is approximated by introducing 6-`Id` gates with a delay of $50ns\times 6=0.3$ $\mu s$. Similarly, the second set of 12-`Id` gates introduces a delay of $0.6$ $\mu s$ close to the actual delay of $0.9375-0.3125=0.625\mu s$.
#
# The `Pragma` preserve blocks are added at the start and end of the program so that the compiler preserves the entire program.
#
# At the end of each program, we place a `MEASURE` operator to read out the result.
print(quadratic_program.out())
# #### Printing the Ramsey DDS Program
#
# The Ramsey program has has 99 `I` gates corresponding to the interval $99\times 50ns=4.95\mu$s interval (close to the desired $5\mu$s) between the $X_{\pi/2}$ intervals.
print(ramsey_program.out())
# ### Compiling and running the pograms on Virtual Quantum Device
#
# We can use `pyQuil` to compile the programs generated by Q-CTRL Open Controls.
#
# To execute the program you will need ForestSDK. Start the the Quil Compiler and QVM in server mode, using the following commands in a terminal.
#
# ```
# $ quilc -S
# $ qvm -S
# ```
# NBVAL_SKIP
quantum_device = get_qc("Aspen-4-2Q-A", as_qvm=True)
# #### Compiling the Quadratic DDS Program
# +
# NBVAL_SKIP
# We loop the program in order to run it several times
'''
trials : int
An integer denoting the number of repeats of the program on quantum device
'''
trials=1000
quadratic_program = quadratic_program.wrap_in_numshots_loop(trials)
##Compiling the quadratic program
executable_quadratic_program_for_qvm = quantum_device.compile(quadratic_program)
# +
# NBVAL_SKIP
# We loop the program in order to run it several times
'''
trials : int
An integer denoting the number of repeats of the program on quantum device
'''
trials=1000
ramsey_program = ramsey_program.wrap_in_numshots_loop(trials)
##Compiling the quadratic program
executable_ramsey_program_for_qvm = quantum_device.compile(ramsey_program)
# -
# ### Run the programs using pyQuil's QVM
#
# We can use the quantum device we created earlier to run the program. The experiment consists of a number of `trials`, which repeats the `Program`. Each run measures the state of the qubit as a measurement. The result is displayed as a histogram.
def plot_trial_outcomes(trial_result):
"""Plots the trial result as probabilities. Expects results from runs involving only
single qubit
Parameters
----------
trial_result : numpy.ndarray
An array where each row contains the output trials in computational basis.
"""
qubit_trial_result = trial_result[:, 0]
number_of_trials = qubit_trial_result.shape
outcome = np.array([number_of_trials-np.sum(qubit_trial_result), np.sum(qubit_trial_result)])
outcome_probabilities = outcome / number_of_trials
plt.bar(np.array([0, 1]), outcome_probabilities)
plt.xticks(np.array([0, 1]), [0, 1])
plt.ylabel('Probabilities')
plt.xlabel('States')
# NBVAL_SKIP
quadratic_result = quantum_device.run(executable_quadratic_program_for_qvm)
plot_trial_outcomes(quadratic_result)
# +
# NBVAL_SKIP
## Run the ramsey sequence program, Get the result (counts of state |1> and |0>), plot the histogram
ramsey_result = quantum_device.run(executable_ramsey_program_for_qvm)
plot_trial_outcomes(ramsey_result)
# -
# ### Compiling and running the pograms on Rigetti Quantum Device
#
# If you have an account on [Rigetti QCS](https://qcs.rigetti.com/sign-in) you can run the programs on a real quantum computer. First you need to [reserve a lattice](https://www.rigetti.com/qcs/docs/reservations) and learn how to [run a jupyter notebook](https://www.rigetti.com/qcs/docs/guides#setting-up-a-jupyter-notebook-on-your-qmi). Then the following commands can be used to execute a DDS.
# +
# NBVAL_SKIP
# Get the reserved lattice. In this case we are using 'Aspen-4-2Q-A' - change this to
# the one you reserve
quantum_device = get_qc("Aspen-4-2Q-A", as_qvm=False)
# +
# NBVAL_SKIP
## Compiling the quadratic program for the chosen real lattice
executable_quadratic_program_for_lattice = quantum_device.compile(quadratic_program)
## Run the quadratic sequence program, Get the result (counts of state |1> and |0>), plot the histogram
quadratic_result = quantum_device.run(executable_quadratic_program_for_lattice)
plot_trial_outcomes(quadratic_result)
# +
# NBVAL_SKIP
## Compiling the ramsey program for the chosen real lattice
executable_ramsey_program_for_lattice = quantum_device.compile(ramsey_program)
## Run the ramsey sequence program, Get the result (counts of state |1> and |0>), plot the histogram
ramsey_result = quantum_device.run(executable_ramsey_program_for_lattice)
plot_trial_outcomes(ramsey_result)
# -
# ## Reducing errors by increasing the coherence time
#
# In the ideal noiseless simulator, both the Ramsey DDS and Quadratic DDS produced exactly the same outcome, the $|1 \rangle$ state with probability 1. However, in a real device, we can see a marked difference. The Quadratic DDS produced a probability distribution closer to the expected outcome. This is because the Quadratic DDS is able to cancel the effects of magnetic noise in the environment - extending the [T2 time](https://en.wikipedia.org/wiki/Spin–spin_relaxation), and effectively increasing the coherence of the qubit.
| examples/export-a-dynamical-decoupling-sequence-to-pyquil.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to the course!
# ## 1. Importing entire text files
# In this exercise, you'll be working with the file `moby_dick.txt`. It is a text file that contains the opening sentences of <NAME>, one of the great American novels! Here you'll get experience opening a text file, printing its contents to the shell and, finally, closing it.
#
# **Instructions**
# - Open the file `moby_dick.txt` as read-only and store it in the variable file. Make sure to pass the filename enclosed in quotation marks `''`.
# - Print the contents of the file to the shell using the `print()` function. As Hugo showed in the video, you'll need to apply the method `read()` to the object `file`.
# - Check whether the file is closed by executing `print(file.closed)`.
# - Close the file using the `close()` method.
# - Check again that the file is closed as you did above.
# +
# Open a file: file
file = open('moby_dick.txt','r')
# Print it
print(file.read())
print()
# Check whether file is closed
print(file.closed)
# Close file
file.close()
# Check whether file is closed
print(file.closed)
# -
# ## 1.1 Importing text files line by line
# For large files, we may not want to print all of their content to the shell: you may wish to print only the first few lines. Enter the `readline()` method, which allows you to do this. When a file called `file` is open, you can print out the first line by executing `file.readline()`. If you execute the same command again, the second line will print, and so on.
#
# **Instructions**
# * Open `moby_dick.txt` using the `with` context manager and the variable `file`.
# * Print the first three lines of the file to the shell by using `readline()` three times within the context manager
# Read & print the first 3 lines
with open('moby_dick.txt') as file:
print(file.readline())
print(file.readline())
print(file.readline())
#
# ## 2. The importance of flat files in data science
# ### 2.1 Pop quiz: examples of flat files
#
# You're now well-versed in importing text files and you're about to become a wiz at importing flat files. But can you remember exactly what a flat file is? Test your knowledge by answering the following question: which of these file types below is NOT an example of a flat file?
#
# Answer the question
#
# 1. A .csv file.
# 2. A tab-delimited .txt.
# 3. A relational database (e.g. PostgreSQL).
#
# **Answer:** 3
#
# ### 2.2 Pop quiz: what exactly are flat files?
#
# Which of the following statements about flat files is incorrect?
#
# Answer the question
#
# 1. Flat files consist of rows and each row is called a record.
# 2. Flat files consist of multiple tables with structured relationships between the tables.
# 3. A record in a flat file is composed of fields or attributes, each of which contains at most one item of information.
# 4. Flat files are pervasive in data science.
#
# **Answer:** 2
# ### 2.3 Why we like flat files and the Zen of Python
#
# In PythonLand, there are currently hundreds of Python Enhancement Proposals, commonly referred to as PEPs. [PEP8](https://www.python.org/dev/peps/pep-0008/), for example, is a standard style guide for Python, written by our sensei <NAME> himself. It is the basis for how we here at DataCamp ask our instructors to [style their code](https://www.datacamp.com/teach/documentation#tab_style_guide_python). Another one of my favorites is [PEP20](https://www.python.org/dev/peps/pep-0020/), commonly called the Zen of Python. Its abstract is as follows:
#
# >Long time Pythoneer <NAME> succinctly channels the BDFL's guiding principles for Python's design into 20 aphorisms, only 19 of which have been written down.
#
# If you don't know what the acronym `BDFL` stands for, I suggest that you look [here](https://docs.python.org/3.3/glossary.html#term-bdfl). You can print the Zen of Python in your shell by typing `import this` into it! You're going to do this now and the 5th aphorism (line) will say something of particular interest.
#
#
# The question you need to answer is: **what is the 5th aphorism of the Zen of Python?**
#
# Possible Answers
# 1. Flat is better than nested.
# 2. Flat files are essential for data science.
# 3. The world is representable as a flat file.
# 4. Flatness is in the eye of the beholder.
#
# **Answer:** 1
# ## 3 Importing flat files using NumPy
# ### 3.1 Using NumPy to import flat files
#
# In this exercise, you're now going to load the MNIST digit recognition dataset using the numpy function `loadtxt()` and see just how easy it can be:
#
# - The first argument will be the filename.
# - The second will be the delimiter which, in this case, is a comma.
# You can find more information about the MNIST dataset [here](http://yann.lecun.com/exdb/mnist/) on the webpage of <NAME>, who is currently Director of AI Research at Facebook and Founding Director of the NYU Center for Data Science, among many other things.
#
# **Instructions**
# * Fill in the arguments of `np.loadtxt()` by passing file and a comma `','` for the delimiter.
# * Fill in the argument of `print()` to print the type of the object `digits`. Use the function `type()`.
# * Execute the rest of the code to visualize one of the rows of the data.
# +
# Import package
import numpy as np
import matplotlib.pyplot as plt
# Assign filename to variable: file
file = 'digits.csv'
# Load file as array: digits
digits = np.loadtxt(file, delimiter=',')
# Print datatype of digits
print(type(digits))
# Select and reshape a row
im = digits[21, 1:]
im_sq = np.reshape(im, (28, 28))
# Plot reshaped data (matplotlib.pyplot already loaded as plt)
plt.imshow(im_sq, cmap='Greys', interpolation='nearest')
plt.show()
# -
# ### 3.2 Customizing your NumPy import
#
# What if there are rows, such as a header, that you don't want to import? What if your file has a delimiter other than a comma? What if you only wish to import particular columns?
#
# There are a number of arguments that `np.loadtxt()` takes that you'll find useful: delimiter changes the delimiter that `loadtxt()` is expecting, for example, you can use `','` and `'\t'` for comma-delimited and tab-delimited respectively; `skiprows` allows you to specify how many rows (not indices) you wish to skip; `usecols` takes a list of the indices of the columns you wish to keep.
#
# The file that you'll be importing, `digits_header.txt`,
# - has a header
# - is tab-delimited.
#
# **Instructions**
# - Complete the arguments of `np.loadtxt()`: the file you're importing is tab-delimited, you want to skip the first row and you only want to import the first and third columns.
# - Complete the argument of the `print()` call in order to print the entire array that you just imported.
# +
# Import numpy
import numpy as np
# Assign the filename: file
file = 'digits_header.txt'
# Load the data: data
data = np.loadtxt(file, delimiter="\t",skiprows=1,usecols=[0,2])
# Print data
print(data)
# -
# ### 3.3 Importing different datatypes
#
# The file `seaslug.txt`
# - has a text header, consisting of strings
# - is tab-delimited.
#
# These data consists of percentage of sea slug larvae that had metamorphosed in a given time period. Read more here.
#
# Due to the header, if you tried to import it as-is using `np.loadtxt()`, Python would throw you a `ValueError` and tell you that it could not convert string to float. There are two ways to deal with this: firstly, you can set the data type argument `dtype` equal to `str` (for string).
#
# Alternatively, you can skip the first row as we have seen before, using the `skiprows` argument.
#
# **Instructions**
# - Complete the first call to `np.loadtxt()` by passing file as the first argument.
# - Execute `print(data[0])` to print the first element of data.
# - Complete the second call to `np.loadtxt()`. The file you're importing is tab-delimited, the datatype is `float`, and you want to skip the first row.
# - Print the 10th element of `data_float` by completing the `print()` command. Be guided by the previous `print()` call.
# - Execute the rest of the code to visualize the data.
# +
# Assign filename: file
file = 'seaslug.txt'
# Import file: data
data = np.loadtxt(file, delimiter='\t', dtype=str)
# Print the first element of data
print(data[0])
# Import data as floats and skip the first row: data_float
data_float = np.loadtxt(file, delimiter='\t', dtype=float, skiprows=1)
# Print the 10th element of data_float
print(data_float[9])
# Plot a scatterplot of the data
plt.scatter(data_float[:, 0], data_float[:, 1])
plt.xlabel('time (min.)')
plt.ylabel('percentage of larvae')
plt.show()
# -
# ### 3.4 Working with mixed datatypes (1)
# Much of the time you will need to import datasets which have different datatypes in different columns; one column may contain strings and another floats, for example. The function `np.loadtxt()` will freak at this. There is another function, `np.genfromtxt()`, which can handle such structures. If we pass `dtype=None` to it, it will figure out what types each column should be.
#
# Import 'titanic.csv' using the function `np.genfromtxt()` as follows:
#
# >data = np.genfromtxt('titanic.csv', delimiter=',', names=True, dtype=None)
#
# Here, the first argument is the filename, the second specifies the delimiter `,` and the third argument `names` tells us there is a header. Because the data are of different types, `data` is an object called a [structured array](http://docs.scipy.org/doc/numpy/user/basics.rec.html). Because numpy arrays have to contain elements that are all the same type, the structured array solves this by being a 1D array, where each element of the array is a row of the flat file imported. You can test this by checking out the array's shape in the shell by executing `np.shape(data)`.
#
# Accessing rows and columns of structured arrays is super-intuitive: to get the ith row, merely execute `data[i]` and to get the column with name `'Fare'`, execute `data['Fare']`.
#
# Print the entire column with name `Survived` to the shell. What are the last 4 values of this column?
#
# Possible Answers
# 1. 1,0,0,1.
# 2. 1,2,0,0.
# 3. 1,0,1,0.
# 4. 0,1,1,1.
#
# **Answer** 3
# +
# above question proof
data = np.genfromtxt('titanic.csv', delimiter=',', names=True, dtype=None)
data['Survived'][-4:]
# -
# ### 3.5 Working with mixed datatypes (2)
#
# You have just used `np.genfromtxt()` to import data containing mixed datatypes. There is also another function `np.recfromcsv()` that behaves similarly to `np.genfromtxt()`, except that its default `dtype` is `None`. In this exercise, you'll practice using this to achieve the same result.
#
# **Instructions**
# - Import `titanic.csv` using the function `np.recfromcsv()` and assign it to the variable, `d`. You'll only need to pass `file` to it because it has the defaults `delimiter=','` and names=True in addition to `dtype=None!`
# - Run the remaining code to print the first three entries of the resulting array `d`.
#
# +
# Assign the filename: file
file = 'titanic.csv'
# Import file using : d
d = np.recfromcsv(file)
# Print out first three entries of d
print(d[:3])
# -
# ## 4. Importing flat files using pandas
# ### 4.1 Using pandas to import flat files as DataFrames (1)
#
# In the last exercise, you were able to import flat files containing columns with different datatypes as `numpy` arrays. However, the `DataFrame` object in pandas is a more appropriate structure in which to store such data and, thankfully, we can easily import files of mixed data types as DataFrames using the pandas functions `read_csv()` and `read_table()`.
#
# **Instructions**
# - Import the `pandas` package using the alias `pd`.
# - Read `titanic.csv` into a DataFrame called `df`. The file name is already stored in the file object.
# - In a `print()` call, view the head of the DataFrame.
# +
# Import pandas as pd
import pandas as pd
# Assign the filename: file
file = 'titanic.csv'
# Read the file into a DataFrame: df
df = pd.read_csv(file)
# View the head of the DataFrame
print(df.head())
# -
# ### 4.2 Using pandas to import flat files as DataFrames (2)
#
# In the last exercise, you were able to import flat files into a `pandas` DataFrame. As a bonus, it is then straightforward to retrieve the corresponding `numpy` array using the attribute `values`. You'll now have a chance to do this using the MNIST dataset, which is available as `digits.csv`.
#
# **Instructions**
# - Import the first 5 rows of the file into a DataFrame using the function `pd.read_csv()` and assign the result to `data`. - You'll need to use the arguments `nrows` and `header` (there is no header in this file).
# - Build a numpy array from the resulting DataFrame in data and assign to `data_array`.
# - Execute `print(type(data_array))` to print the datatype of `data_array`.
# +
# Assign the filename: file
file = 'digits.csv'
# Read the first 5 rows of the file into a DataFrame: data
data=pd.read_csv(file,nrows=5,header=None)
# Build a numpy array from the DataFrame: data_array
data_array=data.values
# Print the datatype of data_array to the shell
print(type(data_array))
# -
# ### 4.3 Customizing your pandas import
# The `pandas` package is also great at dealing with many of the issues you will encounter when importing data as a data scientist, such as comments occurring in flat files, empty lines and missing values. Note that missing values are also commonly referred to as `NA` or `NaN`. To wrap up this chapter, you're now going to import a slightly corrupted copy of the Titanic dataset `titanic_corrupt.txt`, which
#
# * contains comments after the character `'#'`
# - is tab-delimited.
# +
# Import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
import pandas as pd
# Assign filename: file
file = 'titanic_corrupt.txt'
# Import file: data
data = pd.read_csv(file, sep='\t', comment='#', na_values=['Nothing'])
# Print the head of the DataFrame
print(data.head())
# Plot 'Age' variable in a histogram
pd.DataFrame.hist(data[['Age']])
plt.xlabel('Age (years)')
plt.ylabel('count')
plt.show()
| 5. Importing Data in Python (Part 1)/Ch_1_Importing_Data_in_Python_(Part 1)/Ch_1_Importing_Data_in_Python_(Part 1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="socSJe925zFv"
# # Tutorial 2: Working With Datasets
#
# Data is central to machine learning. This tutorial introduces the `Dataset` class that DeepChem uses to store and manage data. It provides simple but powerful tools for efficiently working with large amounts of data. It also is designed to easily interact with other popular Python frameworks such as NumPy, Pandas, TensorFlow, and PyTorch.
#
# ## Colab
#
# This tutorial and the rest in this sequence can be done in Google colab. If you'd like to open this notebook in colab, you can use the following link.
#
# [](https://colab.research.google.com/github/deepchem/deepchem/blob/master/examples/tutorials/02_Working_With_Datasets.ipynb)
#
#
# ## Setup
#
# To run DeepChem within Colab, you'll need to run the following installation commands. This will take about 5 minutes to run to completion and install your environment. You can of course run this tutorial locally if you prefer. In that case, don't run these cells since they will download and install Anaconda on your local machine.
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="OyxRVW5X5zF0" outputId="affd23f1-1929-456a-f8a6-e53a874c84b4"
# !curl -Lo conda_installer.py https://raw.githubusercontent.com/deepchem/deepchem/master/scripts/colab_install.py
import conda_installer
conda_installer.install()
# !/root/miniconda/bin/conda info -e
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="CMWAv-Z46nCc" outputId="9ae7cfd0-ebbf-40b0-f6f1-2940cf32a839"
# !pip install --pre deepchem
# + [markdown] colab_type="text" id="Jk47QTZ95zF-"
# We can now import the `deepchem` package to play with.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="PDiY03h35zF_" outputId="cdd7401d-19a0-4476-9297-b04defc67178"
import deepchem as dc
dc.__version__
# + [markdown] colab_type="text" id="B0u7qIZd5zGG"
# # Anatomy of a Dataset
#
# In the last tutorial we loaded the Delaney dataset of molecular solubilities. Let's load it again.
# + colab={} colab_type="code" id="saTaOpXY5zGI"
tasks, datasets, transformers = dc.molnet.load_delaney(featurizer='GraphConv')
train_dataset, valid_dataset, test_dataset = datasets
# + [markdown] colab_type="text" id="F922OPtL5zGM"
# We now have three Dataset objects: the training, validation, and test sets. What information does each of them contain? We can start to get an idea by printing out the string representation of one of them.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="YEDcUsz35zGO" outputId="5a05747f-8b06-407d-9b11-790a1b4d1c8f"
print(test_dataset)
# + [markdown] colab_type="text" id="E8UCFrrN5zGf"
# There's a lot of information there, so let's start at the beginning. It begins with the label "DiskDataset". Dataset is an abstract class. It has a few subclasses that correspond to different ways of storing data.
#
# - `DiskDataset` is a dataset that has been saved to disk. The data is stored in a way that can be efficiently accessed, even if the total amount of data is far larger than your computer's memory.
# - `NumpyDataset` is an in-memory dataset that holds all the data in NumPy arrays. It is a useful tool when manipulating small to medium sized datasets that can fit entirely in memory.
# - `ImageDataset` is a more specialized class that stores some or all of the data in image files on disk. It is useful when working with models that have images as their inputs or outputs.
#
# Now let's consider the contents of the Dataset. Every Dataset stores a list of *samples*. Very roughly speaking, a sample is a single data point. In this case, each sample is a molecule. In other datasets a sample might correspond to an experimental assay, a cell line, an image, or many other things. For every sample the dataset stores the following information.
#
# - The *features*, referred to as `X`. This is the input that should be fed into a model to represent the sample.
# - The *labels*, referred to as `y`. This is the desired output from the model. During training, it tries to make the model's output for each sample as close as possible to `y`.
# - The *weights*, referred to as `w`. This can be used to indicate that some data values are more important than others. In later tutorials we will see examples of how this is useful.
# - An *ID*, which is a unique identifier for the sample. This can be anything as long as it is unique. Sometimes it is just an integer index, but in this dataset the ID is a SMILES string describing the molecule.
#
# Notice that `X`, `y`, and `w` all have 113 as the size of their first dimension. That means this dataset contains 113 samples.
#
# The final piece of information listed in the output is `task_names`. Some datasets contain multiple pieces of information for each sample. For example, if a sample represents a molecule, the dataset might record the results of several different experiments on that molecule. This dataset has only a single task: "measured log solubility in mols per litre". Also notice that `y` and `w` each have shape (113, 1). The second dimension of these arrays usually matches the number of tasks.
#
# # Accessing Data from a Dataset
#
# There are many ways to access the data contained in a dataset. The simplest is just to directly access the `X`, `y`, `w`, and `ids` properties. Each of these returns the corresponding information as a NumPy array.
# + colab={} colab_type="code" id="e5K3rdGV5zGg"
test_dataset.y
# + [markdown] colab_type="text" id="_Zcd7jTd5zGr"
# This is a very easy way to access data, but you should be very careful about using it. This requires the data for all samples to be loaded into memory at once. That's fine for small datasets like this one, but for large datasets it could easily take more memory than you have.
#
# A better approach is to iterate over the dataset. That lets it load just a little data at a time, process it, then free the memory before loading the next bit. You can use the `itersamples()` method to iterate over samples one at a time.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LJc90fs_5zGs" outputId="8c9fd5ab-e23a-40dc-9292-8b4ff3a86890"
for X, y, w, id in test_dataset.itersamples():
print(y, id)
# + [markdown] colab_type="text" id="aQa88cbj5zGw"
# Most deep learning models can process a batch of multiple samples all at once. You can use `iterbatches()` to iterate over batches of samples.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="HSVqeYox5zGx" outputId="270a6a17-6238-4081-b0cf-3f17e23f4bb5"
for X, y, w, ids in test_dataset.iterbatches(batch_size=50):
print(y.shape)
# -
# `iterbatches()` has other features that are useful when training models. For example, `iterbatches(batch_size=100, epochs=10, deterministic=False)` will iterate over the complete dataset ten times, each time with the samples in a different random order.
#
# Datasets can also expose data using the standard interfaces for TensorFlow and PyTorch. To get a `tensorflow.data.Dataset`, call `make_tf_dataset()`. To get a `torch.utils.data.IterableDataset`, call `make_pytorch_dataset()`. See the API documentation for more details.
#
# The final way of accessing data is `to_dataframe()`. This copies the data into a Pandas `DataFrame`. This requires storing all the data in memory at once, so you should only use it with small datasets.
test_dataset.to_dataframe()
# # Creating Datasets
#
# Now let's talk about how you can create your own datasets. Creating a `NumpyDataset` is very simple: just pass the arrays containing the data to the constructor. Let's create some random arrays, then wrap them in a NumpyDataset.
# +
import numpy as np
X = np.random.random((10, 5))
y = np.random.random((10, 2))
dataset = dc.data.NumpyDataset(X=X, y=y)
print(dataset)
# -
# Notice that we did not specify weights or IDs. These are optional, as is `y` for that matter. Only `X` is required. Since we left them out, it automatically built `w` and `ids` arrays for us, setting all weights to 1 and setting the IDs to integer indices.
dataset.to_dataframe()
# What about creating a DiskDataset? If you have the data in NumPy arrays, you can call `DiskDataset.from_numpy()` to save it to disk. Since this is just a tutorial, we will save it to a temporary directory.
# +
import tempfile
with tempfile.TemporaryDirectory() as data_dir:
disk_dataset = dc.data.DiskDataset.from_numpy(X=X, y=y, data_dir=data_dir)
print(disk_dataset)
# -
# What about larger datasets that can't fit in memory? What if you have some huge files on disk containing data on hundreds of millions of molecules? The process for creating a DiskDataset from them is slightly more involved. Fortunately, DeepChem's `DataLoader` framework can automate most of the work for you. That is a larger subject, so we will return to it in a later tutorial.
# + [markdown] colab_type="text" id="MhZxVoVs5zMa"
# # Congratulations! Time to join the Community!
#
# Congratulations on completing this tutorial notebook! If you enjoyed working through the tutorial, and want to continue working with DeepChem, we encourage you to finish the rest of the tutorials in this series. You can also help the DeepChem community in the following ways:
#
# ## Star DeepChem on [GitHub](https://github.com/deepchem/deepchem)
# This helps build awareness of the DeepChem project and the tools for open source drug discovery that we're trying to build.
#
# ## Join the DeepChem Gitter
# The DeepChem [Gitter](https://gitter.im/deepchem/Lobby) hosts a number of scientists, developers, and enthusiasts interested in deep learning for the life sciences. Join the conversation!
| examples/tutorials/02_Working_With_Datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
invcov=np.array([[2,1,0],[1,2,1],[0,1,2]])
covmat=np.linalg.inv(invcov)
samp = np.random.multivariate_normal([0,0,0],covmat,size=(10000,10000))
#sampcov=np.zeros((3,3,1000))
#sampcovinv=np.zeros((3,3,1000))
sampcorr=np.zeros((3,3,10000))
sampcorrinv=np.zeros((3,3,10000))
for i in range(10000):
#sampcov[:,:,i] = np.cov(samp[:,i,:],rowvar=False)
#sampcovinv[:,:,i] = np.linalg.inv(sampcov[:,:,i])
sampcorr[:,:,i] = np.corrcoef(samp[:,i,:],rowvar=False)
sampcorrinv[:,:,i] = np.linalg.inv(sampcorr[:,:,i])
#sampcov[0,1,:] = sampcov[0,1,:] + np.random.normal(0,1,1000)
#fcov = np.corrcoef(sampcov[0,1:3,:])
#finvcov = np.corrcoef(sampcovinv[0,1:3,:])
sampcorrinv[0,1,:] = sampcorrinv[0,1,:] + np.random.normal(0,1,10000)
fcorrinv = np.corrcoef(sampcorrinv[0,1:3,:])
sampcorr[0,1,:] = sampcorr[0,1,:] + np.random.normal(0,1,10000)
fcorr = np.corrcoef(sampcorr[0,1:3,:])
#print(fcov)
#print(finvcov)
print(fcorr)
print(fcorrinv)
# -
def generate_corr_stats(
invcov,
sampsize=100,
sigscale=1,
normvar=1
):
invcov = invcov*sigscale
covmat = np.linalg.inv(invcov)
samp = np.random.multivariate_normal([0,0,0],covmat,size=(sampsize,sampsize))
sampcov=np.zeros((3,3,sampsize))
sampcovinv=np.zeros((3,3,sampsize))
sampcorr=np.zeros((3,3,sampsize))
sampcorrinv=np.zeros((3,3,sampsize))
for i in range(sampsize):
sampcov[:,:,i] = np.cov(samp[:,i,:],rowvar=False)
sampcovinv[:,:,i] = np.linalg.inv(sampcov[:,:,i])
sampcorr[:,:,i] = np.corrcoef(samp[:,i,:],rowvar=False)
sampcorrinv[:,:,i] = np.linalg.inv(sampcorr[:,:,i])
sampcov[0,1,:] = sampcov[0,1,:] + np.random.normal(0,normvar,sampsize)
sampcovinv[0,1,:] = sampcovinv[0,1,:] + np.random.normal(0,normvar,sampsize)
sampcorr[0,1,:] = sampcorr[0,1,:] + np.random.normal(0,normvar,sampsize)
sampcorrinv[0,1,:] = sampcorrinv[0,1,:] + np.random.normal(0,normvar,sampsize)
fcov = np.corrcoef(sampcov[0,1:3,:])
gcov = np.linalg.inv(fcov)[0,1]
fcovinv = np.corrcoef(sampcovinv[0,1:3,:])
gcovinv = np.linalg.inv(fcovinv)[0,1]
fcorr = np.corrcoef(sampcorr[0,1:3,:])
gcorr = np.linalg.inv(fcorr)[0,1]
fcorrinv = np.corrcoef(sampcorrinv[0,1:3,:])
gcorrinv = np.linalg.inv(fcorrinv)[0,1]
return fcov[0,1], fcovinv[0,1], fcorr[0,1], fcorrinv[0,1], gcov, gcovinv, gcorr, gcorrinv
# +
from graspy.plot import heatmap
invcov=np.array([[2,1,0],[1,2,1],[0,1,2]])
resol=100
fcov = np.zeros((resol,resol))
gcov = np.zeros((resol,resol))
fcovinv = np.zeros((resol,resol))
gcovinv = np.zeros((resol,resol))
fcorr = np.zeros((resol,resol))
gcorr = np.zeros((resol,resol))
fcorrinv = np.zeros((resol,resol))
gcorrinv = np.zeros((resol,resol))
for x in range(100):
for y in range(100):
f1,f2,f3,f4,g1,g2,g3,g4 = generate_corr_stats(invcov,sampsize=100,sigscale=.01*(1+x),normvar=.005*(1+y))
fcov[x,y]=f1
gcov[x,y]=g1
fcovinv[x,y]=f2
gcovinv[x,y]=g2
fcorr[x,y]=f3
gcorr[x,y]=g3
fcorrinv[x,y]=f4
gcorrinv[x,y]=g4
#heatmap(fcov,title='fcov')
#heatmap(gcov,title='gcov')
#heatmap(fcovinv,title='fcovinv')
#heatmap(gcovinv,title='gcovinv')
#heatmap(fcorr,title='fcorr')
#heatmap(gcorr,title='gcorr')
#heatmap(fcorrinv,title='fcorrinv')
#heatmap(gcorrinv,title='gcorrinv')
# +
import matplotlib.pyplot as plt
heatmap(np.absolute(fcov))
plt.title('Correlation of Covariances')
plt.ylabel('Scaling Factor of Inverse Covariance (.01-1.01)')
plt.xlabel('Variance of Noise (.005-.505)')
# -
heatmap(np.absolute(gcov))
print(np.amax(gcov))
plt.title('Inverse Correlation of Covariances')
plt.ylabel('Scaling Factor of Inverse Covariance (.01-1.01)')
plt.xlabel('Variance of Noise (.005-.505)')
heatmap(np.absolute(fcovinv))
heatmap(np.absolute(gcovinv))
# +
heatmap(np.absolute(fcorr))
plt.title('Correlation of Correlations')
plt.ylabel('Scaling Factor of Inverse Covariance (.01-1.01)')
plt.xlabel('Variance of Noise (.005-.505)')
# +
heatmap(np.absolute(gcorr))
plt.title('Inverse Correlation of Correlations')
plt.ylabel('Scaling Factor of Inverse Covariance (.01-1.01)')
plt.xlabel('Variance of Noise (.005-.505)')
# -
heatmap(np.absolute(fcorrinv))
heatmap(np.absolute(gcorrinv))
| experiments/experiment_10/Exp10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow.keras.utils
# +
from keras.layers import Conv3D, MaxPooling3D, Flatten,Dense
from keras.models import Sequential
from keras.datasets import mnist
from keras.layers.normalization import BatchNormalization
from config import config
#from helper_tools.generator import Image_Generator
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# -
"""
https://medium.com/@mrgarg.rajat/training-on-large-datasets-that-dont-fit-in-memory-in-keras-60a974785d71
https://www.datacamp.com/community/tutorials/deep-learning-jupyter-aws
https://www.linkedin.com/learning/neural-networks-and-convolutional-neural-networks-essential-training/creating-and-compiling-the-model?u=36492188
"""
X_train_files = np.load(config.LOCAL_TRAIN_SUBSET)
y_train = pd.read_pickle(config.LOCAL_TRAIN_TARGET)
im_gen = Image_Generator(X_train_files, y_train, config.BATCH_SIZE)
# +
cnn = Sequential()
cnn.add(Conv3D(32, kernel_size=(2,5,5), input_shape=(6,123,95), padding='same', activation='relu'))
cnn.add(BatchNormalization())
cnn.add(MaxPooling3D(pool_size=(2,2,2)))
cnn.add(Dropout(0.1))
cnn.add(Conv3D(64, kernel_size=(2,5,5), padding='same', activation='relu'))
cnn.add(BatchNormalization())
cnn.add(MaxPooling3D(pool_size=(2,2,2)))
cnn.add(Dropout(0.1))
cnn.add(Flatten())
cnn.add(Dense(1024, activation='relu'))
cnn.add(Dense(4,activation='softmax'))
# -
cnn.compile(optimizer='adam', loss='categorical_crossentropy',metrics=['accuracy'])
model.summary()
trained_cnn = cnn.fit(im_gen,epochs=10,verbose=1,
use_multiprocessing=True)
| 02 CNN Initial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to make configurations
#
# For FuxiCTR v1.0 only.
#
# This tutorial presents the details of how to use the YAML config files.
# The dataset_config contains the following keys:
#
# + **dataset_id**: the key used to denote a dataset split, e.g., taobao_tiny_data
# + **data_root**: the directory to save or load the h5 dataset files
# + **data_format**: csv | h5
# + **train_data**: training data file path
# + **valid_data**: validation data file path
# + **test_data**: test data file path
# + **min_categr_count**: the default threshold used to filter rare features
# + **feature_cols**: a list of feature columns, each containing the following keys
# - **name**: feature name, i.e., column header name.
# - **active**: True | False, whether to use the feature.
# - **dtype**: the data type of this column.
# - **type**: categorical | numeric | sequence, which type of features.
# - **source**: optional, which feature source, such as user/item/context.
# - **share_embedding**: optional, specify which feature_name to share embedding.
# - **embedding_dim**: optional, embedding dim of a specific field, overriding the default embedding_dim if used.
# - **pretrained_emb**: optional, filepath of pretrained embedding, which should be a h5 file with two columns (id, emb).
# - **freeze_emb**: optional, True | False, whether to freeze embedding is pretrained_emb is used.
# - **encoder**: optional, "MaskedAveragePooling" | "MaskedSumPooling" | "null", specify how to pool the sequence feature. "MaskedAveragePooling" is used by default. "null" means no pooling is required.
# - **splitter**: optional, how to split the sequence feature during preprocessing; the space " " is used by default.
# - **max_len**: optional, the max length set to pad or truncate the sequence features. If not specified, the max length of all the training samples will be used.
# - **padding**: optional, "pre" | "post", either pre padding or post padding the sequence.
# - **na_value**: optinal, what value used to fill the missing entries of a field; "" is used by default.
# + **label_col**: label name, i.e., the column header of the label
# - **name**: the column header name for label
# - **dtype**: the data type
#
# The model_config contains the following keys:
#
# + **expid**: the key used to denote an experiment id, e.g., DeepFM_test. Each expid corresponds to a dataset_id and the model hyper-parameters used for experiment.
# + **model_root**: the directory to save or load the model checkpoints and running logs.
# + **workers**: the number of processes used for data generator.
# + **verbose**: 0 for disabling tqdm progress bar; 1 for enabling tqdm progress bar.
# + **patience**: how many epochs to stop training if no improvments are made.
# + **pickle_feature_encoder**: True | False, whether pickle feature_encoder
# + **use_hdf5**: True | False, whether reuse h5 data if available
# + **save_best_only**: True | False, whether to save the best model weights only.
# + **every_x_epochs**: how many epochs to evaluate the model on valiadtion set, float supported. For example, 0.5 denotes to evaluate every half epoch.
# + **debug**: True | False, whether to enable debug mode. If enabled, every run will generate a new expid to avoid conflicted runs on two code versions.
# + **model**: model name used to load the specific model class
# + **dataset_id**: the dataset_id used for the experiment
# + **loss**: currently support "binary_crossentropy" only.
# + **metrics**: list, currently support ['logloss', 'AUC'] only
# + **task**: currently support "binary_classification" only
# + **optimizer**: "adam" is used by default
# + **learning_rate**: the initial learning rate
# + **batch_size**: the batch size for model training
# + **embedding_dim**: the default embedding dim for all feature fields. It will be ignored if a feature has embedding_dim value.
# + **epochs**: the max number of epochs for model training
# + **shuffle**: True | False, whether to shuffle data for each epoch
# + **seed**: int, fix the random seed for reproduciblity
# + **monitor**: 'AUC' | 'logloss' | {'AUC': 1, 'logloss': -1}, the metric used to determine early stopping. The dict can be used for combine multiple metrics. E.g., {'AUC': 2, 'logloss': -1} means 2 * AUC - logloss and the larger the better.
# + **monitor_mode**: 'max' | 'min', the mode of the metric. E.g., 'max' for AUC and 'min' for logloss.
#
# There are also some model-specific hyper-parameters. E.g., DeepFM has the following specific hyper-parameters:
# + **hidden_units**: list, hidden units of MLP
# + **hidden_activations**: str or list, e.g., 'relu' or ['relu', 'tanh']. When each layer has the same activation, one could use str; otherwise use list to set activations for each layer.
# + **net_regularizer**: regularizaiton weight for MLP, supporting different types such as 1.e-8 | l2(1.e-8) | l1(1.e-8) | l1_l2(1.e-8, 1.e-8). l2 norm is used by default.
# + **embedding_regularizer**: regularizaiton weight for feature embeddings, supporting different types such as 1.e-8 | l2(1.e-8) | l1(1.e-8) | l1_l2(1.e-8, 1.e-8). l2 norm is used by default.
# + **net_dropout**: dropout rate for MLP, e.g., 0.1 denotes that hidden values are dropped randomly with 10% probability.
# + **batch_norm**: False | True, whether to apply batch normalizaiton on MLP.
#
#
# Many config files are available at https://github.com/xue-pai/FuxiCTR/tree/main/config for your reference. Here, we take the config [demo/demo_config](https://github.com/xue-pai/FuxiCTR/tree/main/demo/demo_config) as an example. The dataset_config.yaml and model_config.yaml are as follows.
#
# dataset_config.yaml
taobao_tiny_data: # dataset_id
data_root: ../data/
data_format: csv
train_data: ../data/tiny_data/train_sample.csv
valid_data: ../data/tiny_data/valid_sample.csv
test_data: ../data/tiny_data/test_sample.csv
min_categr_count: 1
feature_cols:
- {name: ["userid","adgroup_id","pid","cate_id","campaign_id","customer","brand","cms_segid",
"cms_group_id","final_gender_code","age_level","pvalue_level","shopping_level","occupation"],
active: True, dtype: str, type: categorical}
label_col: {name: clk, dtype: float}
# Note that we merge the feature_cols with the same config settings for compactness. But we also could expand them as shown below.
taobao_tiny_data:
data_root: ../data/
data_format: csv
train_data: ../data/tiny_data/train_sample.csv
valid_data: ../data/tiny_data/valid_sample.csv
test_data: ../data/tiny_data/test_sample.csv
min_categr_count: 1
feature_cols:
[{name: "userid", active: True, dtype: str, type: categorical},
{name: "adgroup_id", active: True, dtype: str, type: categorical},
{name: "pid", active: True, dtype: str, type: categorical},
{name: "cate_id", active: True, dtype: str, type: categorical},
{name: "campaign_id", active: True, dtype: str, type: categorical},
{name: "customer", active: True, dtype: str, type: categorical},
{name: "brand", active: True, dtype: str, type: categorical},
{name: "cms_segid", active: True, dtype: str, type: categorical},
{name: "cms_group_id", active: True, dtype: str, type: categorical},
{name: "final_gender_code", active: True, dtype: str, type: categorical},
{name: "age_level", active: True, dtype: str, type: categorical},
{name: "pvalue_level", active: True, dtype: str, type: categorical},
{name: "shopping_level", active: True, dtype: str, type: categorical},
{name: "occupation", active: True, dtype: str, type: categorical}]
label_col: {name: clk, dtype: float}
# The following model config contains two parts. When `Base` is available, the base settings will be shared by all expids. The base settings can be also overridden in expid with the same key. This design is for compactness when a large group of model configs are available, as shown in `./config` folder. `Base` and expid `DeepFM_test` can be either put in the same `model_config.yaml` file or the same `model_config` directory. Note that in any case, each expid should be unique among all the expids.
# +
# model_config.yaml
Base:
model_root: '../checkpoints/'
workers: 3
verbose: 1
patience: 2
pickle_feature_encoder: True
use_hdf5: True
save_best_only: True
every_x_epochs: 1
debug: False
DeepFM_test:
model: DeepFM
dataset_id: taobao_tiny_data # each expid corresponds to a dataset_id
loss: 'binary_crossentropy'
metrics: ['logloss', 'AUC']
task: binary_classification
optimizer: adam
hidden_units: [64, 32]
hidden_activations: relu
net_regularizer: 0
embedding_regularizer: 1.e-8
learning_rate: 1.e-3
batch_norm: False
net_dropout: 0
batch_size: 128
embedding_dim: 4
epochs: 1
shuffle: True
seed: 2019
monitor: 'AUC'
monitor_mode: 'max'
# -
# The `load_config` method will automatically merge the above two parts. If you prefer, it is also flexible to remove `Base` and declare all the settings using only one dict as below.
DeepFM_test:
model_root: '../checkpoints/'
workers: 3
verbose: 1
patience: 2
pickle_feature_encoder: True
use_hdf5: True
save_best_only: True
every_x_epochs: 1
debug: False
model: DeepFM
dataset_id: taobao_tiny_data
loss: 'binary_crossentropy'
metrics: ['logloss', 'AUC']
task: binary_classification
optimizer: adam
hidden_units: [64, 32]
hidden_activations: relu
net_regularizer: 0
embedding_regularizer: 1.e-8
learning_rate: 1.e-3
batch_norm: False
net_dropout: 0
batch_size: 128
embedding_dim: 4
epochs: 1
shuffle: True
seed: 2019
monitor: 'AUC'
monitor_mode: 'max'
| tutorials/v1.0/4_how_to_make_configurations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chinese text summarization algorithm
# ### www.KudosData.com
# #### By: <NAME>
# #### March, 2017
# # Imports
# +
# coding=UTF-8
from __future__ import division
import re
# Python2 unicode & float-division support:
# from __future__ import unicode_literals, division
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
# 中文字符和语言处理库
import jieba
# 机器学习库 sklearn 分类学习模型库
#from sklearn import linear_model
from sklearn.feature_extraction import DictVectorizer # 数据结构变换:把 Dict 转换为 稀疏矩阵
# from sklearn.linear_model import LogisticRegression # 逻辑回归分类模型
# from sklearn.pipeline import make_pipeline # 封装机器学习模型流程
# from sklearn.metrics import confusion_matrix, roc_curve, auc
# 中文显示设置
from pylab import *
mpl.rcParams['font.sans-serif'] = ['SimHei'] # 指定默认字体
mpl.rcParams['axes.unicode_minus'] = False # 解决保存图像是负号'-'显示为方块的问题
mpl.rcParams['font.size'] = 14 # 设置字体大小
np.random.seed(88)
# -
# # Define Functions
# Python3
# 中文分词功能小函数, 输出 字符串, 各词组由空格分隔
def KudosData_word_tokenizer(foo):
# remove lead & tail spaces firstly:
foo = foo.strip()
seg_token = jieba.cut(str(foo), cut_all=True)
seg_str = str(' '.join(seg_token)).strip()
return seg_str
# Python2
# 中文分词功能小函数, 输出 字符串, 各词组由空格分隔
# def KudosData_word_tokenizer(foo):
# seg_token = jieba.cut(foo, cut_all=True)
# seg_str = ' '.join(seg_token)
# return seg_str
# Python3
# 中文分词功能小函数, 输出 dictionary: { key 词组, value 计数 }
def KudosData_word_count(foo):
# remove lead & tail spaces firstly:
foo = foo.strip()
seg_token = jieba.cut(str(foo), cut_all=True)
seg_str = str(' '.join(seg_token)).strip()
seg_count = pd.value_counts(str(seg_str).lower().split(' '))
seg_count = seg_count.to_dict()
seg_count.pop('', None) # remove EMPTY dict key: ''
return seg_count
# Python2
# 中文分词功能小函数, 输出 dictionary: { key 词组, value 计数 }
# def KudosData_word_count(foo):
# seg_token = jieba.cut(foo, cut_all=True)
# seg_str = '^'.join(seg_token)
# seg_count = pd.value_counts(seg_str.lower().split('^'))
# return seg_count.to_dict()
# # Input text
# +
# process Unicode text input
with io.open('input_text.txt','r',encoding='utf8') as f:
content = f.read()
title = '''
<Dummy Title>
'''
# +
# content
# -
def format_sentence(text):
# sentence = re.sub(r'\W+', '#', sentence)
text = re.sub(r'\t+', '', text) # remove one or more Tab
return text
def extract_valid_sentence_words(text):
text = re.sub(r'\W+', '#', text.strip())
return text
def linebreak_conversion_win_linux(text):
text = re.sub(r'\r', '', text) # remove one or more Tab
text = re.sub(r'\u3000', ' ', text) # convert white space: \u3000
return text
# +
def clean_some_whitespace(text): # Does not remove normal Space
# sentence = re.sub(r'\W+', '#', sentence)
text = re.sub(r'\t+', '', text) # remove one or more Tab
text = re.sub(r'\f+', '', text) # remove one or more
text = re.sub(r'\v+', '', text) # remove one or more
text = re.sub(r'\n+', '', text) # remove one or more \n, this is to merge sentences within paragraph
text = re.sub(r'(\^\*\#)( +)(\#\*\^)', '^*##*^', text) # remove one or more Spaces between Paragraph-Tags or Sentence-Tags
text = re.sub(r' +', ' ', text) # merge two or more Spaces to 1 Space
# remove lead & tail spaces:
text =text.strip()
# text = re.sub(r'( +)\W', r'\W', text) # remove Spaces before special char
# text = re.sub(r'\W( +)', r'\W', text) # remove Spaces after special char
return text
# +
### Define Paragraph-Tag =
# #*^P^*#
### Define Sentence-Tag =
# #*^S^*#
# -
# add a special tage to end of each paragraph
def tag_paragraph(text):
text = re.sub(r'((\n ) +)+', '#*^P^*#', text) # Tag paragraph, pattern: \n + two or more Spaces
text = re.sub(r'((\n\t) +)+', '#*^P^*#', text) # Tag paragraph, pattern: \n + two or more Tabs
text = re.sub(r'(\n( *)\n)+', '#*^P^*#', text) # Tag paragraph, pattern: \n + zero or more Spaces + \n
text = re.sub(r'(\#\*\^P\^\*\#)+', '#*^P^*#', text) # merge two or more Paragraph-Tags -> 1 Paragraph-Tag
return text
# add a special tage to end of each sentence
def tag_sentence(text):
text = re.sub(r'。+', '。#*^S^*#', text) # Tag sentence - Chinese
text = re.sub(r'!+', '!#*^S^*#', text) # Tag sentence - Chinese
text = re.sub(r'\?+', '?#*^S^*#', text) # Tag sentence - Chinese
text = re.sub(r';+', ';#*^S^*#', text) # Tag sentence - Chinese
text = re.sub(r'!+', '!#*^S^*#', text) # Tag sentence - English
text = re.sub(r'\?+', '?#*^S^*#', text) # Tag sentence - English
text = re.sub(r';+', ';#*^S^*#', text) # Tag sentence - English
# merge two or more sentence-Tags -> 1 Sentence-Tag
text = re.sub(r'(\W?(\#\*\^S\^\*\#))+', '。#*^S^*#', text)
# remove a Sentence-Tag immediately before an ending ”
text = re.sub(r'\#\*\^S\^\*\#”', '”', text)
# remove a Sentence-Tag immediately before a Paragraph-Tag
text = re.sub(r'(\#\*\^S\^\*\#)( *)(\#\*\^P\^\*\#)', '#*^P^*#', text)
text = re.sub(r'(\#\*\^P\^\*\#)+', '#*^P^*#', text) # merge two or more Paragraph-Tags -> 1 Paragraph-Tag
return text
# ### Start tagging:
content_format = linebreak_conversion_win_linux(content)
# content_format
content_format = tag_paragraph(content_format)
# content_format
content_format = clean_some_whitespace(content_format)
# content_format
content_format = tag_sentence(content_format)
# content_format
content_format = clean_some_whitespace(content_format)
content_format
#
# ### Transfer tagged text to Pandas Dataframe
# Split a text into paragraphs
def split_article_to_paragraphs(text):
# text = text.replace("#*^P^*#", "#*^S^*#") # convert Paragraph-Tag
return text.split("#*^P^*#")
# Split a paragraph into sentences
def split_paragraph_to_sentences(text):
# text = text.replace("#*^P^*#", "#*^S^*#") # convert Paragraph-Tag
return text.split("#*^S^*#")
# +
# 1st loop Paragraphs list, 2nd loop Sentences list
# create a few new columns, then write into dataframe, together with original Sentence string
# define empty dataframe:
df_article = pd.DataFrame(columns=('sentence',
'word_count', # sentence word count, including punctuations
'sentence_id', # unique sentence s/n within an article
'sentence_id_paragraph', # sentence s/n within a paragraph
'paragraph_id',
'class_rank',
'score_word', # score based on word tf-idf
'score_sentence', # score based on intersection of sentence pairs
'score_word_norm', # Normalized score
'score_sentence_norm', # Normalized score
'score',
))
df_sentence_id = 0
# split_article_to_paragraphs:
article_paragraphs = split_article_to_paragraphs(content_format)
for i in range(0, len(article_paragraphs)):
# split_paragraph_to_sentences:
article_paragraphs_sentences = split_paragraph_to_sentences(article_paragraphs[i].strip())
for j in range(0, len(article_paragraphs_sentences)):
if article_paragraphs_sentences[j].strip() != '':
df_sentence_id = df_sentence_id + 1
# write to dataframe:
df_article.loc[len(df_article)] = [article_paragraphs_sentences[j].strip(),
len(article_paragraphs_sentences[j].strip()),
df_sentence_id,
j+1,
i+1,
'',
'',
'',
'',
'',
'']
# -
# Make sure no empty sentences:
print('Number of empty sentences in dataframe: %d ' % len(df_article[df_article['sentence'] == '']))
df_article
# +
# assume the 1st sentence as Title of Article
title = df_article['sentence'][0]
print('Title of Article : ', title)
# -
# # Calculate importance score for each sentence
# ### [Optional Reference] word_tokenizer
# KudosData_word_tokenizer
df_article['sentence_tokenized'] = df_article['sentence'].apply(lambda x: KudosData_word_tokenizer(x))
# ### [Optional Reference] Term Frequency
# KudosData_word_count
df_article['sentence_tf'] = df_article['sentence'].apply(lambda x: KudosData_word_count(x))
# ### [Optional Reference] TF-IDF
# +
from sklearn.feature_extraction.text import TfidfVectorizer
corpus = df_article['sentence_tokenized']
vectorizer = TfidfVectorizer()
# my_stopword_list = ['and','to','the','of', 'in']
#vectorizer = TfidfVectorizer(stop_words=my_stopword_list)
# choice of no nomalization of tfidf output (not recommended)
#vectorizer = TfidfVectorizer(norm=None)
# TF-IDF score
tfidf = vectorizer.fit_transform(corpus)
# IDF score
idf_dict = dict(zip(vectorizer.get_feature_names(), vectorizer.idf_))
# TF is in df_article[['sentence_tf']]
# +
### 把TF-iDF数值赋予相对应的词组
tfidf = tfidf.tocsr()
n_docs = tfidf.shape[0]
tfidftables = [{} for _ in range(n_docs)]
terms = vectorizer.get_feature_names()
for i, j in zip(*tfidf.nonzero()):
tfidftables[i][terms[j]] = tfidf[i, j]
# -
# Document-Term-Matrix's TF-IDF matrix size:
print ('This tfidf matrix is a very large table: [ %d rows/docs X %d columns/words ]'
% (tfidf.shape[0], tfidf.shape[1]))
print ('It contains %d eliments: one score per word per document !'
% (tfidf.shape[0] * tfidf.shape[1]))
# Add tfidf score into dataframe
df_article['tfidf'] = tfidftables
# df_article[['sentence', 'sentence_tokenized', 'tfidf']].head()
df_article[['sentence', 'sentence_tokenized', 'sentence_tf', 'tfidf']]
# ### Scoring (1)
# ### Calculate score_word for each sentence, based on sentence word_count tf-idf:
# +
# experiment: use tf-idf and len(sentence_tokenized) to calculate score
# tmp_mean = tmp_sum / len(df_article['sentence_tokenized'][i])
for i in range(0,len(df_article)):
if len(df_article['tfidf'][i]) == 0:
df_article['score_word'][i] = 0
else:
tmp_sum = 0
for key, values in df_article['tfidf'][i].items():
tmp_sum += values
tmp_mean = tmp_sum / len(df_article['sentence_tokenized'][i])
df_article['score_word'][i] = tmp_mean
# -
# ### Scoring (2)
# ### Calculate score_sentence for each sentence, based on pair-wise sentence comparison/intersection:
# Caculate raw intersection score between pair of two sentences, from df_article['sentence_tokenized']
def sentences_intersection(sent1tokenized, sent2tokenized):
# www.KudosData.com - Chinese
# split the sentence into words/tokens
s1 = set(sent1tokenized.split(" "))
s2 = set(sent2tokenized.split(" "))
# If there is not intersection, just return 0
if (len(s1) + len(s2)) == 0:
print('# If there is not intersection, just return 0')
return 0
# Normalize the result by the average number of words
return len(s1.intersection(s2)) / ((len(s1) + len(s2)) / 2)
# ### Below step runs long time... Tuning needed
# +
# Calculate important score of every pair of sentences
n = len(df_article['sentence_tokenized'])
# [Sam python 2.7 -> 3.4] values = [[0 for x in xrange(n)] for x in xrange(n)]
df_score_raw_values = [[0 for x in range(n)] for x in range(n)]
for i in range(0, n):
for j in range(0, n):
df_score_raw_values[i][j] = sentences_intersection(df_article['sentence_tokenized'][i],
df_article['sentence_tokenized'][j])
# The score of a sentence is the sum of all its intersection
sentences_dic = {}
for i in range(0, n):
df_score = 0
for j in range(0, n):
if i == j:
continue
df_score += df_score_raw_values[i][j]
df_article['score_sentence'][i] = df_score
# +
# df_article.head()
# -
# ### Visualize Data
df_article[df_article['word_count'] == None]
# 图表显示:
plt.figure(figsize=(16, 4))
plt.title(u'图')
plt.xlabel(u'X坐标:Sentence word_count')
plt.ylabel(u'Y坐标:Sentence frequency')
# df_article['word_count'].value_counts().sort_values(ascending=False).plot(kind='bar', color='green')
df_article['word_count'].hist(bins=100)
# plt.gca().invert_yaxis()
plt.show()
# 图表显示:
plt.figure(figsize=(16, 4))
plt.title(u'图')
plt.xlabel(u'X坐标:Paragraph_id')
plt.ylabel(u'Y坐标:Sentence frequency')
df_article['paragraph_id'].hist(bins=100)
# plt.gca().invert_yaxis()
plt.show()
# 图表显示:
plt.figure(figsize=(16, 4))
plt.title(u'图')
plt.xlabel(u'X坐标:sentence_id_paragraph')
plt.ylabel(u'Y坐标:Sentence frequency')
df_article['sentence_id_paragraph'].hist(bins=100)
# plt.gca().invert_yaxis()
plt.show()
# 图表显示:
plt.figure(figsize=(16, 4))
plt.title(u'图')
plt.xlabel(u'X坐标:score_word')
plt.ylabel(u'Y坐标:frequency')
df_article['score_word'].hist(bins = 100)
# plt.xscale('log')
# plt.yscale('log')
#plt.xlim(0,0.5)
#plt.ylim(0,0.5)
# plt.gca().invert_yaxis()
plt.show()
# 图表显示:
plt.figure(figsize=(16, 4))
plt.title(u'图')
plt.xlabel(u'X坐标:score_sentence')
plt.ylabel(u'Y坐标:frequency')
df_article['score_sentence'].hist(bins = 100)
# plt.xscale('log')
# plt.yscale('log')
#plt.xlim(0,0.5)
#plt.ylim(0,0.5)
# plt.gca().invert_yaxis()
plt.show()
# df_article[(df_article['score_word'] > 0.15) & (df_article['score_word'] < 0.25)]
# df_article[(df_article['score_word'] > 0.2)].sort_values(by=['score_sentence', 'score_word'], ascending=[False, False,])
# df_article[(df_article['score_sentence'] > 250)].sort_values(by=['score_word', 'score_sentence'], ascending=[False, False,])
# ### Score Normalization
# log(score_word)
df_article['score_word_log'] = np.log(df_article['score_word'].astype('float64') +
df_article[df_article['score_word'] >0 ]['score_word'].min()/2)
# +
# Normalize score_word_log
# df_article['score_word_norm'] = (df_article['score_word'] - df_article['score_word'].mean()) / df_article['score_word'].std()
df_article['score_word_norm'] = (df_article['score_word_log'] - df_article['score_word_log'].mean()) / df_article['score_word_log'].std()
# -
df_article['score_word_norm'].hist(bins=100)
# +
# Normalize score_sentence
df_article['score_sentence_norm'] = (df_article['score_sentence'] - df_article['score_sentence'].mean()) / df_article['score_sentence'].std()
# -
df_article['score_sentence_norm'].hist(bins=100)
# ### Generate class_rank
# +
# Score integration
# df_article['score'] = (df_article['score_sentence_norm'] + df_article['score_word_norm']) / 2
# <NAME>: 23 Mar 2017 - Experiment found that the score_word, which is based on tf-idf, doesn't seem to work well.
# score_word tends to favor short sentences
# score_sentence tends to favor long sentences
# Hence, here we use score_sentence only for final scoring.
df_article['score'] = df_article['score_sentence']
# df_article['score'] = df_article['score_word']
# -
# Min-Max normalization:
df_article['score'] = (df_article['score'] - df_article['score'].min()) / (df_article['score'].max() -df_article['score'].min())
df_article['score'].hist(bins=100)
df_article['score'].hist(bins=100)
# sort firstly
df_article = df_article.sort_values(by=['paragraph_id', 'score'], ascending=[True, False]).reset_index(drop=True)
# ### Below step runs long time... Tuning needed
# +
# Generate Class_Rank
current_class_rank = 0
current_paragraph_id = 0
for i in range(0, len(df_article)):
if df_article['paragraph_id'][i] != current_paragraph_id: # change of Paragraph, thus reset class_rank
current_class_rank = 1
current_paragraph_id = df_article['paragraph_id'][i]
else:
current_class_rank = current_class_rank + 1
df_article['class_rank'][i] = current_class_rank
# -
# sort Dataframe to 'result lookup mode'
df_article = df_article.sort_values(by=['class_rank', 'score', 'paragraph_id', 'sentence_id'],
ascending=[True, False, True, True]).reset_index(drop=True)
df_article[['sentence',
'paragraph_id',
'sentence_id_paragraph',
'class_rank',
'score',
'sentence_tokenized'
]]
# +
# df_article[(df_article['score'] == 0) | (df_article['score'] == 1)]
# -
# # Extract results based on user parameters:
# * Max number of words
# * % of original number of words
# * Max lines of sentences
#
# Make a dataframe copy
df_article_internal = pd.DataFrame.copy(df_article)
# +
total_words_original_article = df_article['sentence'].map(len).sum()
total_words_internal_article = df_article_internal['sentence'].map(len).sum()
# total_words_article_summary = df_article_final['sentence'].map(len).sum()
# print('total_words_original_article : ', total_words_original_article)
# print('total_words_internal_article : ', total_words_internal_article)
# print('total_words_article_summary : ', total_words_article_summary)
# +
# <NAME>: experiment shows no major improvement to use code in this block:
'''
# Heuristic cleaning:
# 1.Remove sentences, which has only one valid word.
# 2.Remove paragraph, which has only single sentence.
# 1.
df_article_internal = df_article_internal[df_article_internal['sentence_tokenized'].map(len) > 1]
print('*** www.KudosData.com *** Removed number of sentences, which has only one valid word : %d'
% (len(df_article) - len(df_article_internal)))
# 2.
df_article_internal_paragraph = df_article_internal['paragraph_id'].value_counts().to_frame(name = 'sentence_count')
df_article_internal_paragraph = df_article_internal_paragraph[df_article_internal_paragraph['sentence_count'] > 1]
valid_paragraph_id = df_article_internal_paragraph.index.tolist()
df_article_internal = df_article_internal[df_article_internal['paragraph_id'].isin(valid_paragraph_id)]
print('*** www.KudosData.com *** Removed number of sentences in total : %d' % (len(df_article) - len(df_article_internal)))
# sort Dataframe to 'result lookup mode'
df_article_internal = df_article_internal.sort_values(by=['class_rank', 'score', 'paragraph_id', 'sentence_id'],
ascending=[True, False, True, True]).reset_index(drop=True)
# Above sort a must sort !!! for below processing:
# Loop Dataframe, accumulate length of sentences, stop when parm_max_word reached, return the index(), cut dataframe to display
'''
# -
# ### Accept user parameters:
# +
# valid range: >= 0
parm_max_word = 200
# valid range: >= 0
parm_max_sentence = 5
# valid range: >=0
parm_max_percent = 0.01
# -
# ### Validation of user parameters:
# +
if (isinstance(parm_max_word, int) | isinstance(parm_max_word, float)):
if parm_max_word >= 0:
print('!1! valid input parm_max_word : ', parm_max_word)
else:
print('!2! Invalid input parm_max_word : ', parm_max_word)
else:
print('!3! Invalid input parm_max_word : ', parm_max_word)
if (isinstance(parm_max_sentence, int) | isinstance(parm_max_sentence, float)):
if parm_max_sentence >= 0:
print('!1! valid input parm_max_sentence : ', parm_max_sentence)
else:
print('!2! Invalid input parm_max_sentence : ', parm_max_sentence)
else:
print('!3! Invalid input parm_max_sentence : ', parm_max_sentence)
if (isinstance(parm_max_percent, int) | isinstance(parm_max_percent, float)):
if parm_max_percent >= 0:
print('!1! valid input parm_max_percent : ', parm_max_percent)
else:
print('!2! Invalid input parm_max_percent : ', parm_max_percent)
else:
print('!3! Invalid input parm_max_percent : ', parm_max_percent)
# +
# cut by parm_max_percent
# Loop Dataframe, accumulate length of sentences, stop when parm_max_word reached, return the index(), cut dataframe to display
sum_current_word = 0
cut_index = len(df_article_internal['sentence'])
# print('Start loop...')
for s in range(0, len(df_article_internal['sentence'])):
# print('s : %d' % s)
if sum_current_word / total_words_original_article <= parm_max_percent:
sum_current_word += len(df_article_internal['sentence'][s])
else:
# stop, return index number
cut_index = s - 1
sum_current_word -= len(df_article_internal['sentence'][s-1])
# print('To break')
break
# print('End loop')
sum_current_percent = sum_current_word / total_words_original_article
print('---------- cut by parm_max_percent :')
print('sum_current_word / total_words_original_article:', sum_current_percent)
print('cut_index : ', cut_index)
# +
# cut by parm_max_word
# Loop Dataframe, accumulate length of sentences, stop when parm_max_word reached, return the index(), cut dataframe to display
sum_current_word = 0
cut_index = len(df_article_internal['sentence'])
# print('Start loop...')
for s in range(0, len(df_article_internal['sentence'])):
# print('s : %d' % s)
if sum_current_word <= parm_max_word:
sum_current_word += len(df_article_internal['sentence'][s])
else:
# stop, return index number
cut_index = s - 1
sum_current_word -= len(df_article_internal['sentence'][s-1])
# print('To break')
break
# print('End loop')
print('---------- cut by parm_max_word :')
print('sum_current_word :', sum_current_word)
print('cut_index : ', cut_index)
# +
# cut by parm_max_sentence
cut_index = parm_max_sentence
print('---------- cut by parm_max_sentence :')
print('cut_index : ', cut_index)
# -
# Extract top number of sentences as summary, based on: cut_index
df_article_final = df_article_internal[0:cut_index]
# sort by original sentence order
df_article_final = df_article_final.sort_values(by=['sentence_id'], ascending=[True])
df_article_final[['sentence', 'paragraph_id', 'sentence_id_paragraph', 'class_rank', 'score']]
# +
# total_words_original_article = df_article['sentence'].map(len).sum()
# total_words_internal_article = df_article_internal['sentence'].map(len).sum()
total_words_article_summary = df_article_final['sentence'].map(len).sum()
print('total_words_original_article : ', total_words_original_article)
print('total_words_internal_article : ', total_words_internal_article)
print('total_words_article_summary : ', total_words_article_summary)
# -
# # Output results to a file
# +
# print('\n'.join(list(df_article_final['sentence'])))
# +
with io.open('output_topic_summary.txt','w',encoding='utf8') as f:
# f.write("Original Length : %s" % total_words_original_article)
f.write("No. Paragraphs : %d" % df_article_internal['paragraph_id'].max())
f.write("\n")
f.write("Original Length : %s" % total_words_internal_article)
f.write("\n")
f.write("Summary Length : %s" % total_words_article_summary)
f.write("\n")
# f.write("Summary Ratio : %s %%" % (100 * (sum_current_word / total_words_original_article)))
f.write("Summary Ratio : %.2f %%" % (100 * (total_words_article_summary / total_words_internal_article)))
f.write("\n")
f.write("\n")
f.write("Title of Article: %s" % title)
f.write("\n")
f.write("\n")
f.write('\n'.join(list(df_article_final['sentence'])))
f.close()
# -
| topic_summary/dated/topic_summary_chn_exp_v007.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # From [Interferometry Primer notebook](https://github.com/seanandrews/DSHARP_CPDs/blob/master/notebooks/Part3_InterferometryPrimer.ipynb)
# ### Detection
# * Interferometer measures electric field induced by the intensity of the source (star, planet, disk, etc.)
# * In the form of an __EM wave__ with voltage:
#
# $$V_i = A_i\cos(2 \pi vt)$$
#
# * If data is also taken from another source at some baseline distance $D_{ij}$, data is captured at different times
# * There is an associated geometric time delay:
# $$V_j= A_j\cos(2\pi v[t+\tau_g])$$
# where $\tau$ is the time difference.
#
#
# ### Receiving/Processing
# * <blockquote> These voltages are passed from the receiver, through a "mixer" (to properly transmit the signals), to a "correlator", which is the machinery that records the measurements. The correlator cross-correlates the signals from antenna pairs, essentially multiplies and integrates over a given integration interval.
# </blockquote>
#
# * Both EM waves can be represented in terms of __visibilities:__
# * A __visibility__ is a complex number: it has an amplitude and a phase, or (more commonly) a real and imaginary component
#
# $$\mathcal{V}_{ij} = \langle V_i V_j \rangle_t = \langle A_i A_j^\ast \cos{(2 \pi \nu t)} \cos{(2 \pi [\nu t + \nu \tau_g])} \rangle_t$$
#
# * which can be expressed as sum of two cosine terms:
# $$\mathcal{V}_{ij} = \langle A_i A_j^\ast \frac{1}{2}\cos{(4 \pi \nu t)} \rangle_t + \langle A_i A_j^\ast \frac{1}{2}\cos{(2 \pi \nu \tau_g)} \rangle_t$$
#
# * __Explanation__
# 1. Visibilities are represented as oscillating waves with phases (__fringe patterns__) that are dependent on the distance between antennas projected onto the sky (geometric delay) and the frequency of the waves
# * First of the terms loosely "averages to zero" over time
# * Frequency of waves are expressed as spatial frequencies $(u, v)$ (disk/celestial object frame) instead of being dependent on $(i, j)$ (earth observer frame)
# * usually packaged in terms of wavelength (kilolambda, megalambda)
# 2. Geometric delay changes over time because of Earth's rotation
# * Allows us to collect more spatial frequencies/reconstruct more emission distribution
# * Visibility at spatial frequencies $(u, v)$ can be represented as the Fourier Transform of the intensity distribution $I_\nu$:
#
# $$\mathcal{V}_\nu(u, v) = \iint I_\nu(l, m) \, e^{-i 2 \pi (ul + vm)} \, dl \, dm$$
#
# where $(l, m)$ are transformed spatial coordinates (see slide 8 of [this presentation](https://drive.google.com/file/d/1snWR17-5llqqoY5BHPW4I9iFJLAdYCK2/view)).
# * This intensity is projected onto the celestial sphere
#
# #### Addendum (9/24 Notes)
# * Mixing phase also involves _local oscillator_, which mixes the signal from radio frequency (~300 GHz) to orders around (1 GHz)
# * preserves the beat frequencies between the two telescope signals
# * Then travels through fiber optic cables to correlator (differences in path length are accounted)
# * Correlator not only multiplies and integrates, but also embeds a _window function_ to the data
# * __Window function__ tells us how the spectrum is distributed into channels
# * Spacing in channels is determined by the geometric time delay $\tau_g$ of the signals
# * Voltage equations are multiplied by multiple window functions = one visibility * autocorrelated window functions
# * These are multiplied by spectral response functions?
# * These are kernels (vector $\vec{v}$ such that $A \cdot \vec{v} = 0$?) that blur the spectra
# * The FWHM of two channels
# ### Review (thus far)
# <blockquote> To re-iterate: the interferometer measures a collection of complex numbers $\{\mathcal{V}_\nu(u, v)\}$, each of which contains some information (at specific spatial frequencies $(u, v)$) about the image of interest, $I_\nu(l, m)$ in the form of its Fourier Transform (FT). If we had an interferometer that was capable of measuring visibilities at *all* spatial frequencies, we could directly measure the image itself through an inverse FT. As you'll learn below, that's not the case!
# </blockquote>
# ### Fourier Transform Properties
# * Examples are in 1D case, but we can apply it the same in 2D cases too
# * Fourier transform of a function $g(x)$ is $G(s)$, where $x$ is the spatial coordinate (angle on the sky) and $s$ is the spatial frequency
# * $g(x) \,\, ^\mathcal{F}_\rightarrow \,\, G(s)$ is short-hand for $G(s) = \int g(x) \, e^{-i 2\pi x s} \,dx$
# * __Linearity:__
# * Addition
#
# $g(x) + h(x) \,\, ^\mathcal{F}_\rightarrow \,\, G(s) + H(s)$
#
#
# * __Convolution theorem:__
# * Convolution in one plane is multiplication in the other
#
# $g(x) = h(x) \ast k(x) \,\, ^\mathcal{F}_\rightarrow \,\, G(s) = H(s) \times K(s)$
#
# * __Scaling__
# * A larger function in one domain is proportionately smaller in the other
#
# $g(a\,x) \,\, ^\mathcal{F}_\rightarrow \,\, \frac{1}{a} \, G(s / a)$
#
# * __Shifts__
# * An offset in one domain corresponds to a phase shift in the other:
#
# $g(x - x_0) \,\, ^\mathcal{F}_\rightarrow \,\, G(s) \, e^{-i 2\pi x_0 s}$
#
# ### Sampling Function and Imaging
# * The sampling function is a collection of $(u, v)$ points that were recorded in observation
# * __Function:__
#
# $S(u, v) = \sum_k \, \delta(u - u_k, v-v_k)$
#
# (basically a sum of Dirac delta functions at each pair of spatial frequencies)
# * Refresh: delta functions take on a value at particular coordinate, 0 elsewhere
# * Hermitian, so $S(u, v)$ = $S(-u, -v)$
# * The observed visibilites are actually $V_v(u, v)$, such that:
#
# $$V_\nu(u, v) = S(u, v) \, \times \, \mathcal{V}_\nu(u, v)$$
#
# * Inverse Fourier transform of visibilities ($\mathcal{F}^{-1}\{ V_\nu(u, v)\} $) is intensity distribution dirty image $I_\nu^D(l, m)$
# * The dirty image is the convolution of the true image ($\mathcal{V}_\nu(u, v)$, or $I_\nu(l, m)$) with the dirty beam, $s(l, m)$, also referred to as the point-spread function (PSF) or "synthesized" beam, which is the FT of the sampling Function ($S(u, v) \,\, ^\mathcal{F}_\rightarrow \,\, s(l, m)$)
#
# $$I_\nu^D(l, m) = s(l, m) \, \ast \, I_\nu(l, m)$$
# * To reproduce clean image, use a non-linear deconvolution algorithm $\mathtt{clean}$ [(more info here)](https://drive.google.com/file/d/1cB7_B3NtOt2HcnVWbgQ2lvKMkIGJKt_r/view)
# # From [Imaging](https://www.cfa.harvard.edu/~dwilner/videos/SMAIS_2020_Imaging.mp4) Lecture
# * short baseline- wide fringe pattern, low angular resolution
# * long baseline- narrow fringe pattern, high angular resolution
# ### 9/24 Notes
# __Takeaways from Interferometry Primer:__
# * Fourier transform of point source will be constant
# * Fourier transform of an elliptical gaussian distribution will be a rotated elliptical gaussian distribution
# * Fourier transforms of sharp edges (uniform disk) are bessel functions
#
# __(narrow features turn into wide features)__ = things captured at smaller baselines represent large-scale structure and vice versa
# * more angular resolution at larger baselines, but usually need many baselines of varying sizes to fully reconstruct the angular distribution of an object
#
# __Resolution of interferometer:__ $\theta_b \sim \lambda/d$ where d is either telescope diameter, or baseline b/w antennas
# * For small scale data (high baseline), the telescopes are separated by ~16 km, gets you a resolution of ~ 0.03 arcsec
# * For large structure data (low baseline), telescopes are separated by 0.5-2 km
#
#
# * On a plot of visibility vs. baseline, there is less visibility as you go further in baseline b/c there is less emission at a finer resolution
#
# * However, visibility data is limited, you can't measure all (u, v) values
#
# * Baselines within interferometry presents limitations on how large you can go
# * only really a problem for galaxy/stellar research, they combat this by taking interferometer data (multiple telescopes) and add it to the source from one telescope (to get extended view)
#
# Likelihood calculation is possible, the post processing is the difficult part
#
# __Problems to overcome in post processing of likelihood calculation:__
# 1. Because there are window functions and spectral response functions embedded into data, each channel map is not independent
# * channel map i and i+1 are covariant with one another, and the fake data needs to also conform to this using some kind of covariance matrix
# 2. Alma does not record data in an independent fixed reference frame
# * Uses the __LSRK (Kinematic Local Standard of Rest)__
# * Average motion of material in the Milky Way in the neighborhood of the Sun (stars in radius 100 pc from the Sun) [wiki](https://en.wikipedia.org/wiki/Local_standard_of_rest#:~:text=In%20astronomy%2C%20the%20local%20standard,material%20is%20not%20precisely%20circular.)
# * In the range of 202–241 km/s
# * Easy to fix in theory, apply transformation to data
# * __Except__ applying transformations to hundreds of data cubes takes an enormously long time
# * Because of the convolution and interpolation of the transformation, it biases parameters
# * not dynamical mass
# * One way to fix:
# * Bin the observations by one channel map
# * Only transform the halfway point of the observation dataset to standard coordinates
# * covariance matrix will have no covariance, because just single channel map
# * But applying window function makes the matrix invertible (poorly conditioned matrix)
# * Bin the observations by two channel maps (successful)
# * Creates a tri-diagonal covariance matrix, which is invertible
# * __Consequence__: you need 2x better resolution to do this approach
# * This makes posterior widths narrower than they should be, creating precision that you don't have
#
# __Processes that take the longest:__
# 1. Generating cubes
# * can potentially speed this up
# 2. FT of cubes
# 3. Interpolation of spectra
# * Can speed this up using Numba
# * instead of broadcasting arrays, turn them into loops
#
#
# ### Resources:
# [Geometry of Interferometry](https://drive.google.com/file/d/1snWR17-5llqqoY5BHPW4I9iFJLAdYCK2/view)
| notebooks/Justin's Notes/Interferometer Primer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WBk0ZDWY-ff8"
# <table align="center">
# <td align="center"><a target="_blank" href="http://introtodeeplearning.com">
# <img src="https://i.ibb.co/Jr88sn2/mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="https://colab.research.google.com/github/aamini/introtodeeplearning/blob/master/lab1/Part1_TensorFlow.ipynb">
# <img src="https://i.ibb.co/2P3SLwK/colab.png" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/aamini/introtodeeplearning/blob/master/lab1/Part1_TensorFlow.ipynb">
# <img src="https://i.ibb.co/xfJbPmL/github.png" height="70px" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# </table>
#
#
# # Copyright Information
#
# + id="3eI6DUic-6jo"
# Copyright 2021 MIT 6.S191 Introduction to Deep Learning. All Rights Reserved.
#
# Licensed under the MIT License. You may not use this file except in compliance
# with the License. Use and/or modification of this code outside of 6.S191 must
# reference:
#
# © MIT 6.S191: Introduction to Deep Learning
# http://introtode<EMAIL>
#
# + [markdown] id="57knM8jrYZ2t"
# # Lab 1: Intro to TensorFlow and Music Generation with RNNs
#
# In this lab, you'll get exposure to using TensorFlow and learn how it can be used for solving deep learning tasks. Go through the code and run each cell. Along the way, you'll encounter several ***TODO*** blocks -- follow the instructions to fill them out before running those cells and continuing.
#
#
# # Part 1: Intro to TensorFlow
#
# ## 0.1 Install TensorFlow
#
# TensorFlow is a software library extensively used in machine learning. Here we'll learn how computations are represented and how to define a simple neural network in TensorFlow. For all the labs in 6.S191 2021, we'll be using the latest version of TensorFlow, TensorFlow 2, which affords great flexibility and the ability to imperatively execute operations, just like in Python. You'll notice that TensorFlow 2 is quite similar to Python in its syntax and imperative execution. Let's install TensorFlow and a couple of dependencies.
#
# + id="LkaimNJfYZ2w" colab={"base_uri": "https://localhost:8080/"} outputId="abedbd11-ea70-448c-ed60-588ac3a54c8c"
# %tensorflow_version 2.x
import tensorflow as tf
# Download and import the MIT 6.S191 package
# !pip install mitdeeplearning
import mitdeeplearning as mdl
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="2QNMcdP4m3Vs"
# ## 1.1 Why is TensorFlow called TensorFlow?
#
# TensorFlow is called 'TensorFlow' because it handles the flow (node/mathematical operation) of Tensors, which are data structures that you can think of as multi-dimensional arrays. Tensors are represented as n-dimensional arrays of base dataypes such as a string or integer -- they provide a way to generalize vectors and matrices to higher dimensions.
#
# The ```shape``` of a Tensor defines its number of dimensions and the size of each dimension. The ```rank``` of a Tensor provides the number of dimensions (n-dimensions) -- you can also think of this as the Tensor's order or degree.
#
# Let's first look at 0-d Tensors, of which a scalar is an example:
# + id="tFxztZQInlAB" colab={"base_uri": "https://localhost:8080/"} outputId="97086c64-5f66-4787-e353-5cba9e79c442"
sport = tf.constant("Tennis", tf.string)
number = tf.constant(1.41421356237, tf.float64)
print("`sport` is a {}-d Tensor".format(tf.rank(sport).numpy()))
print("`number` is a {}-d Tensor".format(tf.rank(number).numpy()))
# + colab={"base_uri": "https://localhost:8080/"} id="QYQk1SqiRlZL" outputId="66e41a00-6c62-4dca-e323-cc7bc367e922"
print("sport: ", sport)
print()
number
# + id="fDlH5E6fQ6xf"
#rank 0 ~ NO brackets
# + [markdown] id="-dljcPUcoJZ6"
# Vectors and lists can be used to create 1-d Tensors:
# + id="oaHXABe8oPcO" colab={"base_uri": "https://localhost:8080/"} outputId="2dad1911-3e40-4c0c-c332-df6405a47e6f"
sports = tf.constant(["Tennis", "Basketball"], tf.string)
numbers = tf.constant([3.141592, 1.414213, 2.71821], tf.float64)
print("`sports` is a {}-d Tensor with shape: {}".format(tf.rank(sports).numpy(), tf.shape(sports)))
print("`numbers` is a {}-d Tensor with shape: {}".format(tf.rank(numbers).numpy(), tf.shape(numbers)))
# + colab={"base_uri": "https://localhost:8080/"} id="ld6PDRMeR4_X" outputId="94d36998-881d-4fd5-8396-ae0fc0cad957"
print("sports: ", sports)
print()
numbers
# + colab={"base_uri": "https://localhost:8080/"} id="iXt5MSRAFhzr" outputId="0f7cf5e3-7e4c-42ab-e4d9-31f978f4e35c"
#mine 0-d tensor and 1-d tensor:
# example of a 0x0 matriz which is also a 0-d tensor (rank 0 ~ NO brackets: NO arrays, and shape []: 0 arrays, it is just a scalar)
matrix0d = tf.constant(1,tf.int64) # 0 arrays, es un escalar (etiquetado como "1")
# example of a 1x4 matriz which is also a 1-d tensor (rank 1 ~ []: only one array, and shape [4]: 1 arrays of 4 elements)
#1 matriz de 1x4 es realmente un vector fila (de 4 columnas) que contiene escalares como elementos del mismo
matrix1d = tf.constant([1,1,1,1],tf.int64) # 1 array de 4 elementos (cada elemento es un escalar etiquetado como "1")
print("`matrix0d` is a {}-d Tensor with shape: {}".format(tf.rank(matrix0d).numpy(), tf.shape(matrix0d)))
print("`matrix1d` is a {}-d Tensor with shape: {}".format(tf.rank(matrix1d).numpy(), tf.shape(matrix1d)))
# + colab={"base_uri": "https://localhost:8080/"} id="9-x-nhizSK6m" outputId="a2a7c21b-ac47-4002-c9aa-946678bb7d1f"
print("matrix0d: ", matrix0d)
print()
matrix1d
# + [markdown] id="gvffwkvtodLP"
# Next we consider creating 2-d (i.e., matrices) and higher-rank Tensors. For examples, in future labs involving image processing and computer vision, we will use 4-d Tensors. Here the dimensions correspond to the number of example images in our batch, image height, image width, and the number of color channels.
# + id="nSsZr0p5Q2EX"
#rank 1~[]
# + id="tFeBBe1IouS3"
### Defining higher-order Tensors ###
'''TODO: Define a 2-d Tensor'''
# matrix = # TODO
# example of a 3x4 matriz which is also a 2-d tensor (rank 2 ~[[]]: an array containing other arrays, and shape [3 4]: 3 arrays of 4 elements each)
matrix = tf.constant([[1,1,1,1],[2,2,2,2],[3,3,3,3]],tf.int64) # it covers the numbers 1 to 3 (take the multiplication of all "elements" of shape, except the last one, so here it is just 3)
assert isinstance(matrix, tf.Tensor), "matrix must be a tf Tensor object"
assert tf.rank(matrix).numpy() == 2
# + [markdown] id="dX5F5EVEPeGV"
# ***El assert es una instruccion de python que te permite definir condiciones que deban cumplirse siempre. En caso que la expresion booleana sea True assert no hace nada y en caso de False dispara una excepcion.***
# + colab={"base_uri": "https://localhost:8080/"} id="Gh-3PwY-Pa9x" outputId="b7661bb4-1bea-4d04-ff80-99843f4f626a"
isinstance(matrix, tf.Tensor) == True
# + colab={"base_uri": "https://localhost:8080/"} id="eONcElQjO_Oa" outputId="df111852-ff28-41a8-e47a-3bf2e6be6e12"
tf.rank(matrix).numpy() == 2
# + colab={"base_uri": "https://localhost:8080/"} id="qetsENVxO_kO" outputId="d2ac6bfe-6b1f-4787-d37b-4f8b54ab8fb6"
print("`matrix` is a {}-d Tensor with shape: {}".format(tf.rank(matrix).numpy(), tf.shape(matrix)))
# + colab={"base_uri": "https://localhost:8080/"} id="NqKObm1-ST5h" outputId="799f19e7-08c6-4701-8d53-8697562674ae"
matrix
# + [markdown] id="xwZ5-wwwP-rQ"
# **Remember:** The ```shape``` of a Tensor defines its number of dimensions and the size of each dimension. The ```rank``` of a Tensor provides the number of dimensions (n-dimensions) -- you can also think of this as the Tensor's order or degree.
# + id="Ug3YnNE85AsM"
'''Let myself define a 3-d Tensor'''
# example of a 3x4 matriz x depth(=2) which is also a 3-d tensor (rank 3 ~[[[]]]: an array containing other arrays that also contain other ones, and shape [2 3 4]: 2 arrays of 3 arrays of 4 elements each)
matrix3d = tf.constant([ [[1,1,1,1],[2,2,2,2],[3,3,3,3]], [[4,4,4,4],[5,5,5,5],[6,6,6,6]] ],tf.int64) # it covers the numbers 1 to 6 (take the multiplication of all "elements" of shape, except the last one, so here it is 2x3=6)
assert isinstance(matrix3d, tf.Tensor), "matrix must be a tf Tensor object"
assert tf.rank(matrix3d).numpy() == 3
# + colab={"base_uri": "https://localhost:8080/"} id="hJUMKNTe6oun" outputId="43d0fa16-8a3f-4d89-e100-e7705c222082"
isinstance(matrix3d, tf.Tensor) == True
# + colab={"base_uri": "https://localhost:8080/"} id="FV7OomMf6rwJ" outputId="d5e160b4-7650-4e22-ba88-a61f3b0513c8"
tf.rank(matrix3d).numpy() == 3
# + colab={"base_uri": "https://localhost:8080/"} id="kkEeWNX66wae" outputId="ee3cfc46-96ec-4c9b-9492-f5dd586c6b89"
print("`matrix3d` is a {}-d Tensor with shape: {}".format(tf.rank(matrix3d).numpy(), tf.shape(matrix3d)))
# + colab={"base_uri": "https://localhost:8080/"} id="fKJsKoR8ScW3" outputId="3f15c361-f7fb-4192-c6ee-5ef93529c0c8"
matrix3d
# + id="ObbhMz4JBbAx"
'''Let myself define a 4-d Tensor'''
# example of a 3x4 matriz x depth(=2) x another_dim(x5) which is also a 4-d tensor (rank 4 ~[[[[[]]]]]: an array containing other arrays that also contain other ones and so on..., and shape [5 2 3 4]:5 arrays of 2 arrays of 3 arrays of 4 elements each)
# it covers the numbers 1 to 30 (take the multiplication of all "elements" of shape, except the last one, so here it is 5x2x3=30)
matrix4d = tf.constant([ [ [[1,1,1,1],[2,2,2,2],[3,3,3,3]], [[4,4,4,4],[5,5,5,5],[6,6,6,6]] ], [ [[7,7,7,7],[8,8,8,8],[9,9,9,9]], [[10,10,10,10],[11,11,11,11],[12,12,12,12]] ], [ [[13,13,13,13],[14,14,14,14],[15,15,15,15]], [[16,16,16,16],[17,17,17,17],[18,18,18,18]] ], [ [[19,19,19,19],[20,20,20,20],[21,21,21,21]], [[22,22,22,22],[23,23,23,23],[24,24,24,24]] ], [ [[25,25,25,25],[26,26,26,26],[27,27,27,27]], [[28,28,28,28],[29,29,29,29],[30,30,30,30]] ] ],tf.int64)
assert isinstance(matrix4d, tf.Tensor), "matrix must be a tf Tensor object"
assert tf.rank(matrix4d).numpy() == 4
# + colab={"base_uri": "https://localhost:8080/"} id="XDc6CbKMJO86" outputId="2f1b6ef3-5315-4481-db99-cc0863bf0dd2"
isinstance(matrix4d, tf.Tensor) == True
# + colab={"base_uri": "https://localhost:8080/"} id="OMA2Ky73JQ-J" outputId="dc545348-dfa6-464d-e9ee-1a11ac823443"
tf.rank(matrix4d).numpy() == 4
# + colab={"base_uri": "https://localhost:8080/"} id="TE7GNAByJTi4" outputId="1f38a03e-44a3-4473-a8ca-74cb4ae35417"
print("`matrix4d` is a {}-d Tensor with shape: {}".format(tf.rank(matrix4d).numpy(), tf.shape(matrix4d)))
# + colab={"base_uri": "https://localhost:8080/"} id="_mhOo1whSgbd" outputId="6f970016-edc3-4956-e472-3a6fa22f6963"
matrix4d
# + [markdown] id="xOWUmKE_A_01"
# Como ya no es fácil imaginarse algo físicamente en 4D, aparte de haber definido un tensor 4-d por mi propia imaginación, entonces ahora mejor recurro a otro ejemplo más organizado visualmente para entender más, tomado de:
#
# https://www.kaggle.com/omnamahshivai/4d-tensor-numpy-example
# + colab={"base_uri": "https://localhost:8080/"} id="TdY7Iau2BIxg" outputId="d3840745-5a26-4690-c452-4e5ae76b5ab6"
'''
Aug12-2018
<NAME> (https://www.linkedin.com/in/mahesh-babu-mariappan)
Source code for example of a 4d tensor
results:
x.shape
x.ndim
(4, 2, 3, 4)
4
'''
import numpy as np
x = np.array(
[
[
[ [1,2,3,4],
[4,5,6,7],
[7,6,3,2] ],
[ [5,4,5,7],
[4,5,7,4],
[3,6,4,2] ]
],
[
[ [1,2,3,4],
[4,5,6,7],
[7,6,3,2] ],
[ [5,4,5,7],
[4,5,7,4],
[3,6,4,2] ]
],
[
[ [1,2,3,4],
[4,5,6,7],
[7,6,3,2] ],
[ [5,4,5,7],
[4,5,7,4],
[3,6,4,2] ]
],
[
[ [1,2,3,4],
[4,5,6,7],
[7,6,3,2] ],
[ [5,4,5,7],
[4,5,7,4],
[3,6,4,2] ]
]
]
)
print('x.shape: ',x.shape)
print('x.ndim: ',x.ndim)
# + [markdown] id="OjRHsLiBKx5d"
# **Convirtiendo arrays a tensores (utilizando la función tf convert_to_tensor()):**
#
# + colab={"base_uri": "https://localhost:8080/"} id="o4iXlvFPKwu4" outputId="da2e728a-0e50-42fc-96ce-1127bbb720f0"
tensorx = tf.convert_to_tensor(x,tf.int64)
print("`tensorx` is a {}-d Tensor with shape: {}".format(tf.rank(tensorx).numpy(), tf.shape(tensorx)))
print()
print("tensorx:", tensorx)
print()
tensorx
# + id="Zv1fTn_Ya_cz"
'''TODO: Define a 4-d Tensor.'''
# Use tf.zeros to initialize a 4-d Tensor of zeros with size 10 x 256 x 256 x 3.
# You can think of this as 10 images where each image is RGB(3) 256 x 256.
#images = # TODO
# example of a 256x256 (pixelsX,pixelsY) x 3(num_colorsZ:RGB)x 10(num_images) which is also a 4-d tensor (rank 4 ~[[[[]]]]: an array containing other arrays that contain other ones and so on, and shape [10 3 256 256]: 10 images of 3 colors of an square of 256pixels x 256pixels)
#tf.zeros(shape, dtype=tf.dtypes.float32, name=None)
images = tf.zeros([10, 256, 256, 3], dtype=tf.dtypes.int64)
assert isinstance(images, tf.Tensor), "matrix must be a tf Tensor object"
assert tf.rank(images).numpy() == 4, "matrix must be of rank 4"
assert tf.shape(images).numpy().tolist() == [10, 256, 256, 3], "matrix is incorrect shape" # it does not matter where you play the number 3 (before or after 256x256)
# + colab={"base_uri": "https://localhost:8080/"} id="50dHWIXBPGvK" outputId="2b1e64ff-1164-4371-ccdf-2ddf74703a0e"
isinstance(images, tf.Tensor) == True
# + colab={"base_uri": "https://localhost:8080/"} id="3927ibK3PHg3" outputId="b7fb2b88-7749-46fe-fdb0-9c3e5b7b4508"
tf.rank(images).numpy() == 4
# + colab={"base_uri": "https://localhost:8080/"} id="5juRPUOLPI-5" outputId="41a65b81-0041-431d-c076-a29a8bba5b9b"
tf.shape(images).numpy().tolist() == [10, 256, 256, 3]
# + colab={"base_uri": "https://localhost:8080/"} id="AWmu_08XPTaY" outputId="6aebc966-2ccc-4729-936c-86e654dcab48"
print("`images` is a {}-d Tensor with shape: {}".format(tf.rank(images).numpy(), tf.shape(images)))
# + colab={"base_uri": "https://localhost:8080/"} id="3178USF5P2sA" outputId="f9711f44-8f6c-455e-d875-47c5d32df82e"
images
# + colab={"base_uri": "https://localhost:8080/"} id="_csb5rBKQZ1b" outputId="dd71a019-1ba0-4768-95f1-3c5490568f12"
#images is equivalent to images2
images2 = tf.zeros([10, 3, 256, 256], dtype=tf.dtypes.int64) # it does not matter where you play the number 3 (before or after 256x256)
images2
# + [markdown] id="wkaCDOGapMyl"
# As you have seen, the ```shape``` of a Tensor provides the number of elements in each Tensor dimension. The ```shape``` is quite useful, and we'll use it often. You can also use slicing to access subtensors within a higher-rank Tensor:
# + colab={"base_uri": "https://localhost:8080/"} id="0kvCntnDRbE8" outputId="2d40ed85-cbc6-47ac-8340-da85b5337f25"
#RECALLING "matrix":
matrix
# + id="FhaufyObuLEG" colab={"base_uri": "https://localhost:8080/"} outputId="3445b074-ccc9-4a0e-f361-4be849174e05"
row_vector = matrix[1]
column_vector = matrix[:,2]
scalar = matrix[1, 2]
print("`row_vector`: {}".format(row_vector.numpy()))
print("`column_vector`: {}".format(column_vector.numpy()))
print("`scalar`: {}".format(scalar.numpy()))
# + [markdown] id="iD3VO-LZYZ2z"
# ## 1.2 Computations on Tensors
#
# A convenient way to think about and visualize computations in TensorFlow is in terms of graphs. We can define this graph in terms of Tensors, which hold data, and the mathematical operations that act on these Tensors in some order. Let's look at a simple example, and define this computation using TensorFlow:
#
# 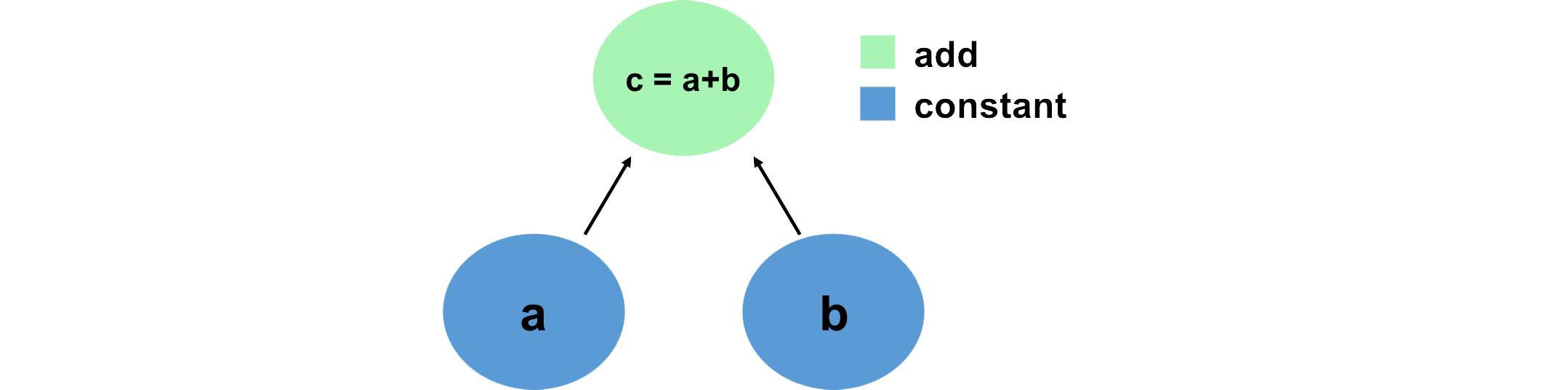
# + id="X_YJrZsxYZ2z" colab={"base_uri": "https://localhost:8080/"} outputId="47202df9-ea9b-4018-c700-47f88fce7cd9"
# Create the nodes in the graph, and initialize values
a = tf.constant(15)
b = tf.constant(61)
# Add them!
c1 = tf.add(a,b)
c2 = a + b # TensorFlow overrides the "+" operation so that it is able to act on Tensors
print(c1)
print(c2)
# + [markdown] id="Mbfv_QOiYZ23"
# Notice how we've created a computation graph consisting of TensorFlow operations, and how the output is a Tensor with value 76 -- we've just created a computation graph consisting of operations, and it's executed them and given us back the result.
#
# Now let's consider a slightly more complicated example:
#
# 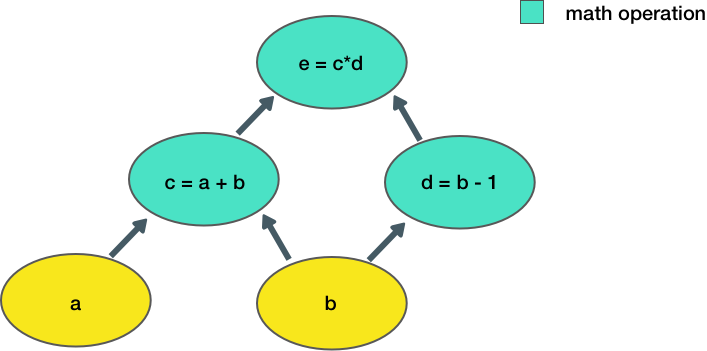
#
# Here, we take two inputs, `a, b`, and compute an output `e`. Each node in the graph represents an operation that takes some input, does some computation, and passes its output to another node.
#
# Let's define a simple function in TensorFlow to construct this computation function:
# + id="PJnfzpWyYZ23"
### Defining Tensor computations ###
# Construct a simple computation function
def func(a,b):
'''TODO: Define the operation for c, d, e (use tf.add, tf.subtract, tf.multiply).'''
#c = # TODO
#d = # TODO
#e = # TODO
c = tf.add(a,b)
uno= tf.constant(1, tf.float64)
d = tf.subtract(b,uno)
e = tf.multiply(c,d)
return e
# + [markdown] id="AwrRfDMS2-oy"
# Now, we can call this function to execute the computation graph given some inputs `a,b`:
# + id="pnwsf8w2uF7p" colab={"base_uri": "https://localhost:8080/"} outputId="7a723f53-bb93-400d-dfc7-787fbf3270c3"
# Consider example values for a,b
a, b = 1.5, 2.5
#turning a, b into tensors
a = tf.constant(a, tf.float64)
b= tf.constant(b, tf.float64)
# Execute the computation
e_out = func(a,b)
print(e_out)
# + colab={"base_uri": "https://localhost:8080/"} id="nEiTopRtGdcu" outputId="9e50e52d-0887-4f9b-a17f-bbe33c260347"
#directly because tensorflow overrides the operators:
(a+b)*(b-1)
# + id="ebR3A3mFIKVl"
#Directly with a function
# Construct a simple computation function
def func1(a,b):
c = a+b
d = b-1
e = c*d
return e
# + colab={"base_uri": "https://localhost:8080/"} id="x-cU2MQgI1QI" outputId="a098eabc-b6a2-40ac-a7d4-1068965fa7d7"
# Consider example values for a,b
a, b = 1.5, 2.5
#turning a, b into tensors
a = tf.constant(a, tf.float64)
b= tf.constant(b, tf.float64)
# Execute the computation
e_out = func1(a,b)
print(e_out)
# + [markdown] id="6HqgUIUhYZ29"
# Notice how our output is a Tensor with value defined by the output of the computation, and that the output has no shape as it is a single scalar value.
# + [markdown] id="1h4o9Bb0YZ29"
# ## 1.3 Neural networks in TensorFlow
# We can also define neural networks in TensorFlow. TensorFlow uses a high-level API called [Keras](https://www.tensorflow.org/guide/keras) that provides a powerful, intuitive framework for building and training deep learning models.
#
# Let's first consider the example of a simple perceptron defined by just one dense layer: $ y = \sigma(xW + b)$, where $W$ represents a matrix of weights, $b$ is a bias, $x$ is the input, $\sigma$ is the sigmoid activation function, and $y$ is the output. We can also visualize this operation using a graph:
#
# 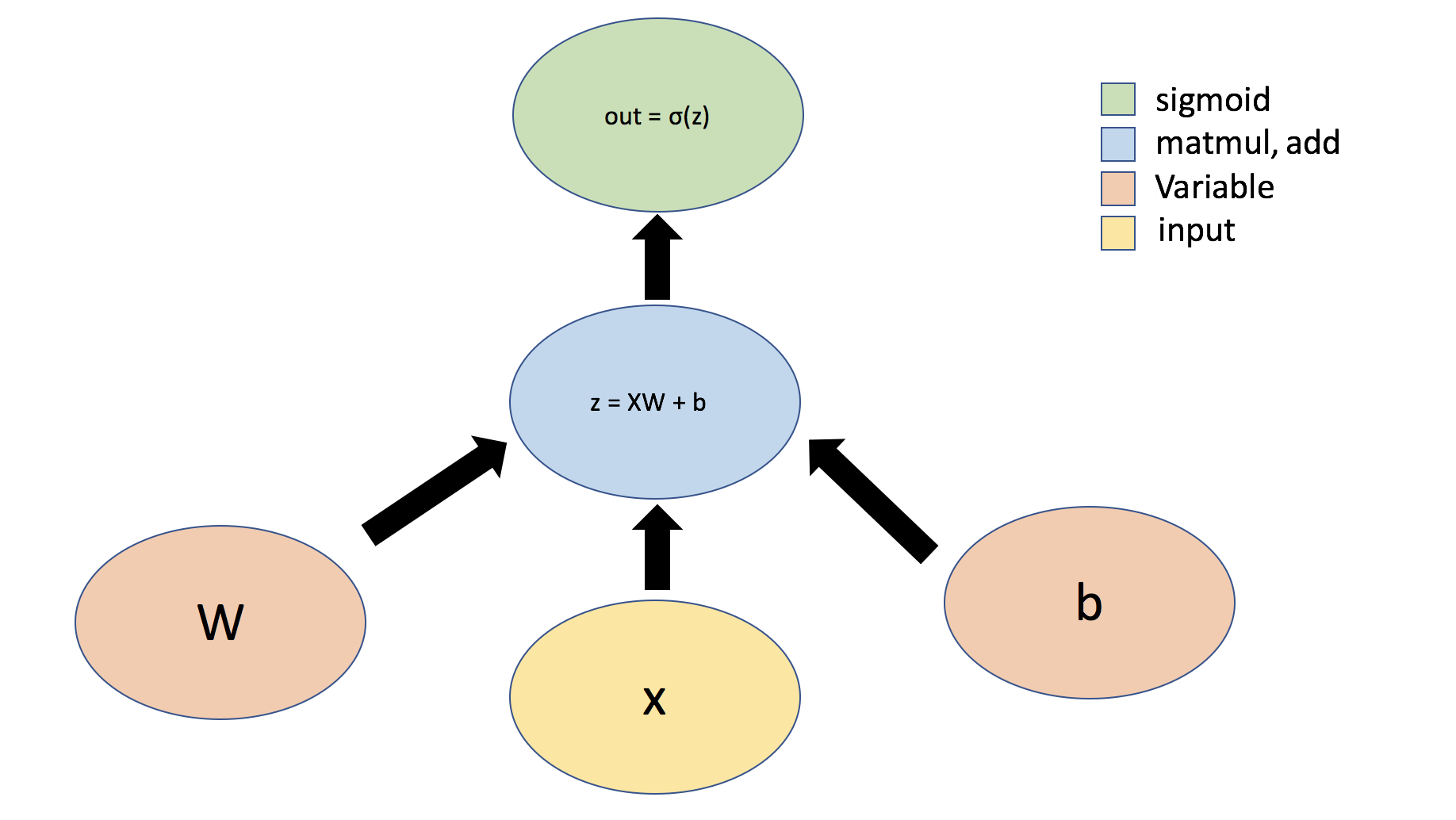
#
# Tensors can flow through abstract types called [```Layers```](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Layer) -- the building blocks of neural networks. ```Layers``` implement common neural networks operations, and are used to update weights, compute losses, and define inter-layer connectivity. We will first define a ```Layer``` to implement the simple perceptron defined above.
# + id="HutbJk-1kHPh" colab={"base_uri": "https://localhost:8080/"} outputId="fe6afe92-1237-4f9d-d925-9f77281f97c7"
### Defining a network Layer ###
# n_output_nodes: number of output nodes
# input_shape: shape of the input
# x: input to the layer
class OurDenseLayer(tf.keras.layers.Layer):
def __init__(self, n_output_nodes):
super(OurDenseLayer, self).__init__()
self.n_output_nodes = n_output_nodes
print("n_output_nodes= ", self.n_output_nodes)
print()
def build(self, input_shape):
print("input_shape= ", input_shape)
print()
d = int(input_shape[-1])
print("d = last element of shape= ",d)
print()
# Define and initialize parameters: a weight matrix W and bias b
# Note that parameter initialization is random!
self.W = self.add_weight("weight", shape=[d, self.n_output_nodes]) # note the dimensionality
print("W= ",self.W) # to know its value
print()
self.b = self.add_weight("bias", shape=[1, self.n_output_nodes]) # note the dimensionality
print("b= ",self.b) # to know its value
print()
def call(self, x):
'''TODO: define the operation for z (hint: use tf.matmul)'''
#z = # TODO
z = tf.add(tf.matmul(x,self.W),self.b) # let it be with tf operations better
print("z= ",z) # to know its value
print()
#z = tf.matmul(x,self.W) + self.b # another way to do it: "+" is an overrode operations for tensors
'''TODO: define the operation for out (hint: use tf.sigmoid)'''
#y = # TODO
y = tf.sigmoid(z)
print("y= ",y) # to know its value
print()
return y
# Since layer parameters are initialized randomly, we will set a random seed for reproducibility (to get the same values when exexuted several times at different moments)
tf.random.set_seed(1) # like in DS4A
layer = OurDenseLayer(3)# n_output_nodes=3 (y13, so it will have 3 values in an array of 1 row and 3 columns)
layer.build((1,2)) #input_shape=(1,2), according to what is inside build(): it means W23 and b13
x_input = tf.constant([[1,2.]], shape=(1,2)) #it means x12=[1,2], so z13=(x12)(W23)+(b13)
print("x= ",x_input) # to know its value
y = layer.call(x_input) #y=sigmoid(z), so y13
print("y=sigmoid(z)= ",y) # to know its value
# test the output!
print("y= ",y.numpy())
mdl.lab1.test_custom_dense_layer_output(y)
# + colab={"base_uri": "https://localhost:8080/"} id="98jHtu9kMskG" outputId="ae9d2513-d27b-446a-d5af-62e179ef6c7e"
W23=np.array([[-0.73366153, 0.8796015 , 0.28695 ],
[-0.14340228, -0.4558388 , 0.3122064 ]])
print("W23= ",W23)
print()
W23
# + colab={"base_uri": "https://localhost:8080/"} id="Uv0H-TcLGRtt" outputId="5ea0472c-50f1-4161-8267-77af0f978c1c"
#according to the above results, let me "prove it" once again using linear algebra:
#given data:
#x is given as input itself:
x12=np.array([[1., 2.]])
print("x12= ",x12)
print()
#firstly initilization of W and b:
W23=np.array([[-0.73366153, 0.8796015 , 0.28695 ],
[-0.14340228, -0.4558388 , 0.3122064 ]])
print("W23= ",W23)
print()
b13=np.array([[ 0.02475715, -0.13831842, -0.2240473 ]])
print("b13= ",b13)
print()
#so now, calculating z:
#z13=(x12)*(W23)+(b13)
z13 = np.dot(x12,W23) + b13
print("z13= ",z13)
print()
#passing z into the non-linear operation of a sigmoid (element by element) to get y:
#y=sigmoid(z)
#I can define myself that fuction in terms of maths
def sigmoid(x):
return 1 / (1 + np.exp(-x))
#logistic.cdf is the sigmoid function as a command y python for arrays
from scipy.stats import logistic
#expit is ALSO the sigmoid function as a command y python for arrays
from scipy.special import expit
#let´s checkout the results using all of those 3 options above:
# %timeit -r 1 sigmoid(z13)
y13_sigmoid = sigmoid(z13) #this one is the best!
print("y13_sigmoid= ",y13_sigmoid)
print()
# %timeit -r 1 logistic.cdf(z13)
y13_logistic_cdf = logistic.cdf(z13) #this one is the best!
print("y13_logistic_cdf= ",y13_logistic_cdf)
print()
# %timeit -r 1 expit(z13)
y13_expit = expit(z13) #this one is the best!
print("y13_expit= ",y13_expit)
print()
print("Compacted:")
print()
print("Dense() implements the operation: output = activation(dot(input, kernel) + bias) where activation is the element-wise activation function passed as the activation argument, kernel is a weights matrix created by the layer, and bias is a bias vector created by the layer (only applicable if use_bias is True)")
print()
print("Here it is: z13 = sigmoid ( np.dot(x12,W23) + b13) = ", sigmoid(np.dot(x12,W23)+b13))
# + [markdown] id="Jt1FgM7qYZ3D"
# Conveniently, TensorFlow has defined a number of ```Layers``` that are commonly used in neural networks, for example a [```Dense```](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense?version=stable). Now, instead of using a single ```Layer``` to define our simple neural network, we'll use the [`Sequential`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/Sequential) model from Keras and a single [`Dense` ](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/Dense) layer to define our network. With the `Sequential` API, you can readily create neural networks by stacking together layers like building blocks.
# + id="7WXTpmoL6TDz"
### Defining a neural network using the Sequential API ###
# Import relevant packages
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense
# Define the number of outputs
n_output_nodes = 3
# First define the model
model = Sequential()
'''TODO: Define a dense (fully connected) layer to compute z'''
# Remember: dense layers are defined by the parameters W and b!
# You can read more about the initialization of W and b in the TF documentation :)
#(son operaciones que se hacen internamente al inicializar con valores aleatorios la primera vez)
# (*) https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense?version=stable
# dense_layer = # TODO
dense_layer = Dense(n_output_nodes, activation='sigmoid')
# Add the dense layer to the model
model.add(dense_layer)
##Taken from(*): https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense?version=stable
## Create a `Sequential` model and add a Dense layer as the first layer.
# model = tf.keras.models.Sequential()
# model.add(tf.keras.Input(shape=(16,)))
# model.add(tf.keras.layers.Dense(32, activation='relu'))
## Now the model will take as input arrays of shape (None, 16)
## and output arrays of shape (None, 32).
## Note that after the first layer, you don't need to specify
## the size of the input anymore:
#model.add(tf.keras.layers.Dense(32))
#model.output_shape
## Answer: (None, 32)
# + [markdown] id="HDGcwYfUyR-U"
# That's it! We've defined our model using the Sequential API. Now, we can test it out using an example input:
# + id="sg23OczByRDb" colab={"base_uri": "https://localhost:8080/"} outputId="d0989035-42d6-4e07-87ac-ac5bef2c61dc"
# Test model with example input
x_input = tf.constant([[1,2.]], shape=(1,2))
'''TODO: feed input into the model and predict the output!'''
#model_output = # TODO
model_output = model(x_input).numpy()# this one gets the final result (output = activation(dot(input, kernel) + bias))
print(model_output) #the output is different from the one we got before, due to the different W and b values which were initialized using Dense
# + [markdown] id="596NvsOOtr9F"
# In addition to defining models using the `Sequential` API, we can also define neural networks by directly subclassing the [`Model`](https://www.tensorflow.org/api_docs/python/tf/keras/Model?version=stable) class, which groups layers together to enable model training and inference. The `Model` class captures what we refer to as a "model" or as a "network". Using Subclassing, we can create a class for our model, and then define the forward pass through the network using the `call` function. Subclassing affords the flexibility to define custom layers, custom training loops, custom activation functions, and custom models. Let's define the same neural network as above now using Subclassing rather than the `Sequential` model.
# + id="K4aCflPVyViD"
### Defining a model using subclassing ###
from tensorflow.keras import Model
from tensorflow.keras.layers import Dense
class SubclassModel(tf.keras.Model):
# In __init__, we define the Model's layers
def __init__(self, n_output_nodes):
super(SubclassModel, self).__init__()
'''TODO: Our model consists of a single Dense layer. Define this layer.'''
#self.dense_layer = '''TODO: Dense Layer'''
self.dense_layer = Dense(n_output_nodes, activation='sigmoid')
# In the call function, we define the Model's forward pass.
def call(self, inputs):
return self.dense_layer(inputs)
# + [markdown] id="U0-lwHDk4irB"
# Just like the model we built using the `Sequential` API, let's test out our `SubclassModel` using an example input.
#
#
# + id="LhB34RA-4gXb" colab={"base_uri": "https://localhost:8080/"} outputId="402732d7-e0be-4b58-809b-6cf631c62424"
n_output_nodes = 3
model = SubclassModel(n_output_nodes)
x_input = tf.constant([[1,2.]], shape=(1,2))
# this one gets the final result (output = activation(dot(input, kernel) + bias)):
print(model.call(x_input))#the output is different from the one we got before, due to the different W and b values which were initialized using Dense
# + [markdown] id="HTIFMJLAzsyE"
# Importantly, Subclassing affords us a lot of flexibility to define custom models. For example, we can use boolean arguments in the `call` function to specify different network behaviors, for example different behaviors during training and inference. Let's suppose under some instances we want our network to simply output the input, without any perturbation. We define a boolean argument `isidentity` to control this behavior:
# + id="P7jzGX5D1xT5"
### Defining a model using subclassing and specifying custom behavior ###
from tensorflow.keras import Model
from tensorflow.keras.layers import Dense
class IdentityModel(tf.keras.Model):
# As before, in __init__ we define the Model's layers
# Since our desired behavior involves the forward pass, this part is unchanged
def __init__(self, n_output_nodes):
super(IdentityModel, self).__init__()
self.dense_layer = tf.keras.layers.Dense(n_output_nodes, activation='sigmoid')
'''TODO: Implement the behavior where the network outputs the input, unchanged,
under control of the isidentity argument.'''
def call(self, inputs, isidentity=False):#default value for isidentity is False
x = self.dense_layer(inputs)
'''TODO: Implement identity behavior'''
if isidentity==True:
return inputs #the network outputs the input, unchanged (y=x)
else:
return x #the network outputs the result y changed (due to Dense: y=sigmoid(xW+b))
# + [markdown] id="Ku4rcCGx5T3y"
# Let's test this behavior:
# + id="NzC0mgbk5dp2" colab={"base_uri": "https://localhost:8080/"} outputId="9df671fe-c1e3-47dd-a067-b07b14996044"
n_output_nodes = 3
model = IdentityModel(n_output_nodes)
x_input = tf.constant([[1,2.]], shape=(1,2))
'''TODO: pass the input into the model and call with and without the input identity option.'''
#out_activate = # TODO
#out_identity = # TODO
out_activate = model.call(x_input) # isidentity=False is by default (no need to put it)
out_identity = model.call(x_input, True) # isidentity=True (necessary to specify it)
print("Network output with activation: {}; network identity output: {}".format(out_activate.numpy(), out_identity.numpy()))
# + [markdown] id="7V1dEqdk6VI5"
# Now that we have learned how to define `Layers` as well as neural networks in TensorFlow using both the `Sequential` and Subclassing APIs, we're ready to turn our attention to how to actually implement network training with backpropagation.
# + [markdown] id="dQwDhKn8kbO2"
# ## 1.4 Automatic differentiation in TensorFlow
#
# [Automatic differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation)
# is one of the most important parts of TensorFlow and is the backbone of training with
# [backpropagation](https://en.wikipedia.org/wiki/Backpropagation). We will use the TensorFlow GradientTape [`tf.GradientTape`](https://www.tensorflow.org/api_docs/python/tf/GradientTape?version=stable) to trace operations for computing gradients later.
#
# When a forward pass is made through the network, all forward-pass operations get recorded to a "tape"; then, to compute the gradient, the tape is played backwards. By default, the tape is discarded after it is played backwards; this means that a particular `tf.GradientTape` can only
# compute one gradient, and subsequent calls throw a runtime error. However, we can compute multiple gradients over the same computation by creating a ```persistent``` gradient tape.
#
# First, we will look at how we can compute gradients using GradientTape and access them for computation. We define the simple function $ y = x^2$ and compute the gradient:
# + id="tdkqk8pw5yJM"
### Gradient computation with GradientTape ###
# y = x^2
# Example: x = 3.0
x = tf.Variable(3.0)
# Initiate the gradient tape
with tf.GradientTape() as tape:
# Define the function
y = x * x
# Access the gradient -- derivative of y with respect to x
dy_dx = tape.gradient(y, x)
assert dy_dx.numpy() == 6.0
# + colab={"base_uri": "https://localhost:8080/"} id="WyPT_hcpmGbf" outputId="564ec0fb-83e4-478a-daf7-dfc0e4492206"
x
# + colab={"base_uri": "https://localhost:8080/"} id="fnVLBrGImA9S" outputId="2794491b-a98c-4a78-8ecd-c4a1e02ed916"
dy_dx
# + colab={"base_uri": "https://localhost:8080/"} id="wugRFWR_l77r" outputId="78459bb2-03dc-42ee-a4cb-3f9cbc2c8829"
dy_dx.numpy() == 6.0
# + [markdown] id="JhU5metS5xF3"
# In training neural networks, we use differentiation and stochastic gradient descent (SGD) to optimize a loss function. Now that we have a sense of how `GradientTape` can be used to compute and access derivatives, we will look at an example where we use automatic differentiation and SGD to find the minimum of $L=(x-x_f)^2$. Here $x_f$ is a variable for a desired value we are trying to optimize for; $L$ represents a loss that we are trying to minimize. While we can clearly solve this problem analytically ($x_{min}=x_f$), considering how we can compute this using `GradientTape` sets us up nicely for future labs where we use gradient descent to optimize entire neural network losses.
# + attributes={"classes": ["py"], "id": ""} id="7g1yWiSXqEf-" colab={"base_uri": "https://localhost:8080/", "height": 421} outputId="8b49ae4b-f4d7-4302-9636-2f24ce060ab0"
### Function minimization with automatic differentiation and SGD ###
# Initialize a random value for our initial x
x = tf.Variable([tf.random.normal([1])])
print("Initializing x={}".format(x.numpy()))
print()
learning_rate = 1e-2 # learning rate for SGD
history = []
# Define the target value
x_f = 4
# We will run SGD for a number of iterations. At each iteration, we compute the loss,
# compute the derivative of the loss with respect to x, and perform the SGD update.
for i in range(500):
with tf.GradientTape() as tape:
'''TODO: define the loss as described above'''
#loss = # TODO
loss = (x-x_f) * (x-x_f)
# loss minimization using gradient tape
grad = tape.gradient(loss, x) # compute the derivative of the loss with respect to x
new_x = x - learning_rate*grad # sgd update
x.assign(new_x) # update the value of x
history.append(x.numpy()[0])
print("History of x as it reaches to the target x_f:")
print()
print(history)
print()
# Plot the evolution of x as we optimize towards x_f!
plt.plot(history)
plt.plot([0, 500],[x_f,x_f])
plt.legend(('Predicted', 'True'))
plt.xlabel('Iteration')
plt.ylabel('x value')
# + id="PXJ-_bdhohRv"
# + [markdown] id="pC7czCwk3ceH"
# `GradientTape` provides an extremely flexible framework for automatic differentiation. In order to back propagate errors through a neural network, we track forward passes on the Tape, use this information to determine the gradients, and then use these gradients for optimization using SGD.
# + [markdown] id="tVNUj0ZnodFf"
# **Finished on March-10th-2021!**
#
# *(Student: <NAME>)*
#
| MIT6S191/Lab1/Copia_de_Part1_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ARCH
# language: python
# name: arch
# ---
# +
import os
import pandas as pd
import numpy as np
from lifelines import CoxPHFitter
import plotly.graph_objects as go
from sklearn.preprocessing import StandardScaler
from plotly.subplots import make_subplots
# -
# # Set export path
# Create path for exporting
path = '../Results/Survival analysis/'
if not os.path.exists(path):
os.makedirs(path)
# # Load the survival dataset
survival_data = pd.read_csv('../Datasets/survival_data.csv')
survival_data
# # Cox hazard analysis using Max initial VAF
# ### LBC21
def survival_analysis(keep_columns, cohort):
# Select cohort and columns
cox_data = survival_data[survival_data.cohort.isin(cohort)][keep_columns + ['days_from_wave1', 'dead']]
# Exclude columns not used as covariates and filter for nan values
cox_data = cox_data.dropna()
# normalise columns used for regression
for column in keep_columns:
data = cox_data[column] - np.mean(cox_data[column])
data = data/np.std(data)
cox_data[column] = data
# Train Cox proportional hazard model
cph = CoxPHFitter()
cph.fit(cox_data, duration_col='days_from_wave1', event_col='dead')
# access the individual results using cph.summary
cph.print_summary()
return cph
def plot_hr_analysis(model, covariate):
fig = make_subplots(rows=1, cols=2, column_widths=[0.3, 0.7],
subplot_titles=(f'Estimated hazard ratio', f'Survival stratification'))
fig.add_trace(
go.Scatter(
y=[model.hazard_ratios_[0]],
x=[covariate],
marker_symbol='diamond',
marker_size=15,
showlegend=False,
error_y=dict(
type='data',
symmetric=False,
array=np.exp(np.array(model.confidence_intervals_)[:,1])-model.hazard_ratios_[0],
arrayminus=model.hazard_ratios_[0]-np.exp(np.array(model.confidence_intervals_)[:,0]))
), row=1, col=1)
# Plot covariate effect
for covariate in model.params_.index:
values =[-2, 0 , 2]
partial_ax = model.plot_partial_effects_on_outcome(covariates=covariate, values=values, cmap='coolwarm')
partial_ax.get_figure()
#add traces to figure
fig.add_trace(
go.Scatter(x=partial_ax.lines[1].get_xdata(),
y=partial_ax.lines[1].get_ydata(),
mode='lines', line=dict(dash='dash', shape='hv'),
name='Mean'), row=1, col=2)
fig.add_trace(
go.Scatter(x=partial_ax.lines[0].get_xdata(),
y=partial_ax.lines[0].get_ydata(),
mode='lines', line=dict(shape='hv'),
name='-2 SD'), row=1, col=2)
fig.add_trace(
go.Scatter(x=partial_ax.lines[2].get_xdata(),
y=partial_ax.lines[2].get_ydata(),
mode='lines', line=dict(shape='hv'),
name='2 SD'), row=1, col=2)
fig.update_layout(template='simple_white',
title=f'Effect of {covariate} on survival',
legend_title_text=f'{covariate}')
y_range_hazards = [np.floor(np.exp(np.array(model.confidence_intervals_)))[0,0], np.ceil(np.exp(np.array(model.confidence_intervals_)))[0,1]]
fig.update_yaxes(title_text="Hazard Ratio (95% CI)", range=y_range_hazards, row=1, col=1, dtick=1)
fig.update_yaxes(title_text="Survivors (proportion)", row=1, col=2, dtick=0.2)
fig.update_xaxes(title_text=covariate, showticklabels=False,tickvals=[0], row=1, col=1)
fig.update_xaxes(title_text="Years", row=1, col=2)
return fig
# # Max initial VAF
# ## LBC21
# +
cohort = [21]
survival_columns = ['Max initial vaf']
cph = survival_analysis(survival_columns, cohort)
fig = plot_hr_analysis(cph, covariate=survival_columns[0])
fig.show()
fig.write_image(path + 'LBC21_init_vaf.svg',width=1000)
# -
# ## LBC36
# +
cohort = [36]
survival_columns = ['Max initial vaf']
cph = survival_analysis(survival_columns, cohort)
fig = plot_hr_analysis(cph, covariate=survival_columns[0])
fig.show()
# -
# # Max gradient
# ## LBC21
cohort = [21]
survival_columns = ['Gradient']
cph = survival_analysis(survival_columns, cohort)
fig = plot_hr_analysis(cph, covariate=survival_columns[0])
fig.show()
# ## LBC36
cohort = [36]
survival_columns = ['Gradient']
cph = survival_analysis(survival_columns, cohort)
fig = plot_hr_analysis(cph, covariate=survival_columns[0])
fig.show()
# ## Combined cohorts
cohort = [21, 36]
survival_columns = ['Gradient']
cph = survival_analysis(survival_columns, cohort)
fig = plot_hr_analysis(cph, covariate=survival_columns[0])
fig.show()
fig.write_image('../Results/Survival/LBC_gradient.svg', width=1000)
| Notebooks/.ipynb_checkpoints/Survival_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Transmission-Distribution Power Flow Co-simulation
# This script runs an iterative transmission-distribution power flow. The network is assumed to consist of a single transmission network connected to distribution feeders at each load bus. The number of distribution feeders connected is determined based on the real power load at the bus and the injection of the distribution feeder. In this example, we use the following:
# + Transmission system: 200-bus network (synthetic network for Illinois system..from TAMU)
# + Distribution feeder: IEEE 8500-node feeder.
#
# The metadata1 supplied has the code for making the connection information for T and D. It is created by the supplied MATLAB function makemetadatafile.m. The metadatafile is required by both the transmission and distribution federates to configure the connection between T and D federates. One can vary the number of federates by setting 'nbdry_selected' variable in the code block below. With nbdry_selected=1, only the first boundary bus will be selected. This boundary bus is set to have three distribution federates (the number of federates is based on the active power load at the bus and the injection at the distribution feeder)
import os
import sys
import subprocess
import shlex
import operator
# +
metadatafile='metadata1'
nbdry_selected = 1
# Metadafile having number of boundary buses
# and the feeder connections at those buses
file = open(metadatafile,"r")
linenum = 1
bdry_buses = []
dist_feeders = {}
for line in file:
if linenum == 1:
nbdry_nfeeders = line.rstrip(' \n').split(',')
nbdry = int(nbdry_nfeeders[0])
nfeeders = int(nbdry_nfeeders[1])
elif linenum < nbdry+2:
if(len(bdry_buses) < nbdry_selected):
bdry_buses.append(line.rstrip(' \n'))
else:
values1 = line.rstrip(' \n')
values = values1.split(',')
if values[0] in bdry_buses:
dist_feeders[values[1]]= int(values[0]) # name:boundary bus
linenum = linenum+1
file.close()
nbdry=len(bdry_buses)
nfeeders=len(dist_feeders)
print 'nbdry=%d ' %nbdry
print 'nfeeders=%d'%nfeeders
# Create another metadatafile with the selected number of boundary buses and feeders
fp = open('metadatafile',"w")
fp.write("%d,%d\n" % (nbdry,nfeeders))
for bus in bdry_buses:
print("%d" % int(bus))
fp.write("%d\n" % int(bus))
for k in dist_feeders:
fp.write("%d,%s\n" % (dist_feeders[k],k))
print("%d,%s" % (dist_feeders[k],k))
fp.close()
nfeds = nfeeders+1
print 'nfederates=%d' %nfeds
metadatafile='metadatafile'
# +
print 'Broker'
broker_args='-nfeds '+str(nfeds)
broker_cmdline='./helicsbroker '+broker_args
broker = shlex.split(broker_cmdline)
print broker_cmdline+'\n'
## Launch broker
subprocess.Popen(broker)
##Launch Transmission federate
print 'Transmission'
netfile='datafiles/case_ACTIVSg200.m'
#print metadatafile
# Launch Transmission federate simulation
pflowhelicst_args_files ='-netfile '+netfile+' -metadatafile '+metadatafile
pflowhelicst_args=pflowhelicst_args_files
pflowhelicst_cmdline='./PFLOWHELICST '+pflowhelicst_args
print pflowhelicst_cmdline+'\n'
pflowhelicst = shlex.split(pflowhelicst_cmdline)
subprocess.Popen(pflowhelicst)
##Launch distribution federates
fednum=0
dnetfile='datafiles/8500-Node/Master.dss'
for k in dist_feeders:
fednum = fednum + 1
print 'D federate '+k
# Dist. federate 1
netfile=dnetfile
dtopic=k
pflowhelicsd_args = '-netfile '+netfile+' -dtopic '+dtopic
pflowhelicsd_cmdline='./PFLOWHELICSD '+pflowhelicsd_args
print pflowhelicsd_cmdline+'\n'
pflowhelicsd = shlex.split(pflowhelicsd_cmdline)
subprocess.Popen(pflowhelicsd)
# -
| ANL-TD-Iterative-Pflow/runtdpflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
file_names = dict()
file_names["a"] = "a_example.txt"
file_names["b"] = "b_lovely_landscapes.txt"
file_names["c"] = "c_memorable_moments.txt"
file_names["d"] = "d_pet_pictures.txt"
file_names["e"] = "e_shiny_selfies.txt"
# get txt file
input_path = "inputs/"
fname = input_path + file_names["b"]
input_file_name = "b_to_input.txt"
output_file_name = "b_output.txt"
with open(fname) as f:
content = f.readlines()
content = [x.strip() for x in content]
print(content)
# +
# prase through input file and put content in a list
N = int(content[0])
# storage is a list of tuples of (orientation, [attributes], [id])
storage = [0] * N
for i in range(1, N+1):
line = content[i].split()
orientation = line[0]
attributes = set(line[2:])
item = (orientation, attributes, [i - 1])
storage[i-1] = item
# -
# give the score of the output list
# ex: [[1], [0,2]]
def score(output_list):
score = 0
M = len(output_list)
for i in range(M-1):
S_left = output_list[i]
S_right = output_list[i+1]
attributes_left = set()
attributes_right = set()
for photo_id in S_left:
photo_attributes = storage[photo_id][1]
attributes_left = attributes_left.union(photo_attributes)
for photo_id in S_right:
photo_attributes = storage[photo_id][1]
attributes_right = attributes_right.union(photo_attributes)
num_intersection = len(attributes_left.intersection(attributes_right))
num_left = len(attributes_left.difference(attributes_right))
num_right = len(attributes_right.difference(attributes_left))
score += min(num_intersection, num_left, num_right)
return score
test_output = [[0], [3], [1, 2]]
print(score(test_output))
# This text cell is for ideas
#
# should use all photos
#
# use gradient method
#
# two ways to influence score:
#
# The order of the slides
#
# How the vertical slides are grouped together
#
# Traveling Saleman Problem
#
# https://github.com/jvkersch/pyconcorde
# helper function ot write the solution list to a txt file
def write_output(filename, solution):
complete_file_name = filename + ".txt"
with open(filename, "w+") as f:
for photo_ids in solution:
line = " ".join([str(num) for num in photo_ids])
line += "\n"
f.write(line)
# test write output
filename = "test_solution"
write_output(filename, test_output)
# import TSPSolver
from concorde.tsp import TSPSolver
# +
# approach 1, randomly assign vertical pairing and then optimize/finefune
vertical_nodes = []
horizontal_nodes = []
for i in range(len(storage)):
photo = storage[i]
if photo[0] == "V":
vertical_nodes.append(photo)
else:
horizontal_nodes.append(photo)
import random
# randomly shuffle the list
random.shuffle(vertical_nodes)
print(vertical_nodes)
merged_nodes = [0] * (len(vertical_nodes) // 2)
for i in range(len(merged_nodes)):
left_vertical_photo = vertical_nodes[2*i]
right_vertical_photo = vertical_nodes[2*i + 1]
orientation = "V"
indexes = left_vertical_photo[2] + right_vertical_photo[2]
tags = left_vertical_photo[1].union(right_vertical_photo[1])
merged_nodes[i] = [orientation, tags, indexes]
# -
print(merged_nodes)
print(horizontal_nodes)
list_for_tsp = merged_nodes + horizontal_nodes
# +
M = len(list_for_tsp)
weights = [ [0]*M for _ in range(M) ]
# set of ints
nodes_used = set()
# list of tuples (start, end)
weights_indexes = []
max_weight = 0
for i in range(M):
left_node = list_for_tsp[i]
attributes_left = left_node[1]
for j in range(i+1, M):
right_node = list_for_tsp[j]
attributes_right = right_node[1]
num_intersection = len(attributes_left.intersection(attributes_right))
num_left = len(attributes_left.difference(attributes_right))
num_right = len(attributes_right.difference(attributes_left))
weight = min(num_intersection, num_left, num_right)
weights[i][j] = weight
if weight > 0:
nodes_used.add(i)
nodes_used.add(j)
weights_indexes.append((i, j))
if max_weight < weight:
max_weight = weight
# -
print(weights)
# +
input_to_pycorde = open(input_file_name, "w+")
num_nodes = len(nodes_used)
num_edges = len(weights_indexes)
input_to_pycorde.write(str(num_nodes) + " " + str(num_edges) + "\n")
for k in range(num_edges):
i, j = weights_indexes[k]
weight = weights[i][j]
input_to_pycorde.write(str(i) + " " + str(j) + " " + str(-weight) + "\n")
# -
from ortools.constraint_solver import pywrapcp
from ortools.constraint_solver import routing_enums_pb2
# Distance callback
def create_distance_callback(dist_matrix):
# Create a callback to calculate distances between cities.
def distance_callback(from_node, to_node):
return int(dist_matrix[from_node][to_node])
return distance_callback
max_weight += 1
for i in range(M):
for j in range(i+1, M):
weights[i][j] = max_weight - weights[i][j]
print(weights)
dist_matrix = weights
city_names = [node[2] for node in list_for_tsp]
print(city_names)
tsp_size = len(city_names)
num_routes = 1
depot = 0
solution_list = []
# Create routing model
if tsp_size > 0:
routing = pywrapcp.RoutingModel(tsp_size, num_routes, depot)
search_parameters = pywrapcp.RoutingModel.DefaultSearchParameters()
# Create the distance callback.
dist_callback = create_distance_callback(dist_matrix)
routing.SetArcCostEvaluatorOfAllVehicles(dist_callback)
# Solve the problem.
assignment = routing.SolveWithParameters(search_parameters)
if assignment:
# Solution distance.
print("Total distance: " + str(assignment.ObjectiveValue()) + " miles\n")
# Display the solution.
# Only one route here; otherwise iterate from 0 to routing.vehicles() - 1
route_number = 0
index = routing.Start(route_number) # Index of the variable for the starting node.
route = ''
while not routing.IsEnd(index):
# Convert variable indices to node indices in the displayed route.
route_i = city_names[routing.IndexToNode(index)]
solution_list.append(route_i)
route += str(route_i) + ' -> '
index = assignment.Value(routing.NextVar(index))
route += str(city_names[routing.IndexToNode(index)])
print("Route:\n\n" + route)
else:
print('No solution found.')
else:
print('Specify an instance greater than 0.')
score(solution_list)
write_output(output_file_name, solution_list)
| submission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: AVES
# language: python
# name: aves
# ---
# +
import sys
from pathlib import Path
AVES_ROOT = Path("..") if not "google.colab" in sys.modules else Path("aves_git")
# -
from aves.data import census
comunas = census.read_census_map('comuna', path=AVES_ROOT / "data" / "external" / "censo_2017_R10")
chiloé = comunas.query('NOM_PROVIN == "CHILOÉ"')
chiloé.plot()
# +
import pandas as pd
ebs = pd.read_stata('../data/external/casen/Base de datos EBS 2021 STATA.dta')
casen = pd.read_stata('../data/external/casen/Casen en Pandemia 2020 STATA.dta').rename(columns={'folio': 'folio_casen', 'id_persona': 'id_persona_casen'})
# -
casen.expc.value_counts().plot()
casen.expc.sum(), casen.expp.sum(), casen.expr.sum()
pd.set_option('display.max_columns', None)
casen.query('expc > 15500')
ebs.query('provincia == "102 Chiloé"').shape
casen.query('provincia == "Chiloé"').shape
casen.query('provincia == "Chiloé"').r2.value_counts()
avecindados = casen.query('provincia == "Chiloé"').query('r2 == "En otra comuna de Chile. Especifique ¿cuál comuna?"')
avecindados.r2_c_cod.value_counts()[:20]
chilotes = casen.query('provincia == "Chiloé"')
chilotes.y1.plot(kind='hist')
import seaborn as sns
from matplotlib import pyplot as plt
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chilotes naive')
sns.histplot(data = chilotes, x = 'y1', stat='density', bins=50)
plt.show()
plt.close()
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chilotes weighted')
sns.histplot(data = chilotes, x = 'y1', stat='density', weights='expp', bins=50)
plt.show()
plt.close()
plt.figure(figsize=(16, 9))
sns.scatterplot(data = chilotes, x = 'y1', y = 'e6a')
plt.figure(figsize=(16, 9))
sns.scatterplot(data = casen, x = 'y1', y = 'e6a')
# +
def is_chiloé(provincia):
if provincia != 'Chiloé':
return 'Resto de Chile'
else:
return 'Chiloé'
casen['chiloe'] = casen.provincia.apply(is_chiloé)
# -
plt.figure(figsize=(16, 9))
sns.scatterplot(data = casen, x = 'y1', y = 'e6a', hue='chiloe')
casen.query('provincia == "Chiloé"').shape[0]
chilotes.shape[0]
continentales = casen.query('provincia != "Chiloé"').sample(chilotes.shape[0])
continentales['donde'] = 'Resto de Chille'
chilotes['donde'] = 'Chiloé'
mix = pd.concat([chilotes, continentales])
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chiloé/Chile naive')
sns.boxplot(data = mix, x = 'y1', y = 'e6a', hue='donde')
def reindex_df(df, weight_col):
"""expand the dataframe to prepare for resampling
result is 1 row per count per sample"""
df = df.reindex(df.index.repeat(df[weight_col]))
df.reset_index(drop=True, inplace=True)
return(df)
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chiloé/Chile weighted')
sns.boxplot(data = reindex_df(mix, 'expp'), x = 'y1', y = 'e6a', hue='donde')
plt.figure(figsize=(16, 9))
sns.boxplot(data = avecindados, x = 'y1', y = 'e6a')
avecindados.oficio1_08.value_counts()
chilotes.comuna.value_counts()[:10]
chilotes['comuna'] = chilotes.comuna.cat.remove_unused_categories()
# +
comunas_interes = ['Castro', 'Ancud', 'Chonchi', 'Quinchao', 'Dalcahue', 'Quellón']
plot_data = chilotes.query('comuna in @comunas_interes')
plot_data['comuna'] = plot_data.comuna.cat.remove_unused_categories()
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chiloé naive')
sns.boxplot(data = plot_data, x = 'y1', y = 'comuna' )
# +
comunas_interes = ['Castro', 'Ancud', 'Chonchi', 'Quinchao', 'Dalcahue', 'Quellón']
plot_data = chilotes.query('comuna in @comunas_interes')
plot_data['comuna'] = plot_data.comuna.cat.remove_unused_categories()
plt.figure(figsize=(16, 9))
plt.title('Ingresos Chiloé weighted')
sns.boxplot(data = reindex_df(plot_data, 'expc'), x = 'y1', y = 'comuna' )
# -
chilotes.groupby(['comuna', 'id_vivienda'], observed=True).count().dropna() #.groupby('comuna').mean() #.mean().id_persona_casen
for group, df in chilotes.groupby('comuna').count().groupby(level=0):
display(group)
display(df)
break
ebs.query('provincia == "Chiloe"')
ebs.provincia.unique()[:100]
ebs_chiloé_exp = reindex_df(ebs.query('provincia == "102 Chiloé"'), 'fexp')
likert2int = {
'1. Pésimo': -2,
'2. Malo': -1,
'3. Regular': 0,
'4. Bueno': 1,
'5. Excelente': 2
}
ebs_chiloé_exp['estado_medio_ambiente'] = ebs_chiloé_exp.g2_3.apply(lambda x: likert2int[x])
ebs_chiloé_exp_mean = ebs_chiloé_exp.groupby('comuna', as_index=False, observed=True).mean()
ebs_chiloé_exp_mean
chiloé['comuna'] = chiloé.NOM_COMUNA.str.title()
# +
geo_ebs_chiloé = chiloé.merge(ebs_chiloé_exp_mean, on='comuna')
geo_ebs_chiloé
# -
geo_ebs_chiloé.plot(column='estado_medio_ambiente', figsize=(16, 9), legend=True)
geo_ebs_chiloé
| notebooks/900-vs-Chilote-avecindados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="gf3sDlIrDfl6"
# # Multi-Class Classification
# + id="np1nF94uDkw_" executionInfo={"status": "ok", "timestamp": 1620492223119, "user_tz": -330, "elapsed": 5776, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
import tensorflow as tf
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="9mI1M6deD-Hu" executionInfo={"status": "ok", "timestamp": 1620492223120, "user_tz": -330, "elapsed": 5770, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="d260ed26-f171-4bd8-c4d1-877000e856d5"
from tensorflow.keras.datasets import fashion_mnist
# The Data is already been sorted to Train and Test Data for us
(train_X, train_y), (test_X, test_y) = fashion_mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/"} id="oZfdskYFEFTQ" executionInfo={"status": "ok", "timestamp": 1620492223120, "user_tz": -330, "elapsed": 5758, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="a8b47727-1652-4a1d-bdeb-1eeaeffdd7a3"
train_X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="KJh5IreWEjAV" executionInfo={"status": "ok", "timestamp": 1620492223121, "user_tz": -330, "elapsed": 5745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="c582e9eb-965a-4a9a-b4ed-effdd9be8911"
train_X[0].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="ig3-sDIyE1yN" executionInfo={"status": "ok", "timestamp": 1620492223121, "user_tz": -330, "elapsed": 5729, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="bbe2ba66-1475-42f9-f903-4463c4448b1f"
plt.imshow(train_X[0])
# + colab={"base_uri": "https://localhost:8080/"} id="218CRtQDFaLG" executionInfo={"status": "ok", "timestamp": 1620492223122, "user_tz": -330, "elapsed": 5714, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="699bed99-e8c5-45b5-fe21-b58c4c041762"
train_y.shape, train_y[0]
# + colab={"base_uri": "https://localhost:8080/"} id="1Tn_nR8dFf0L" executionInfo={"status": "ok", "timestamp": 1620492223122, "user_tz": -330, "elapsed": 5698, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="1c464812-a25c-49ea-a5fb-ecd8ebe3b215"
train_y.min(), train_y.max()
# + id="pm5ZVVIqFw7D" executionInfo={"status": "ok", "timestamp": 1620492223123, "user_tz": -330, "elapsed": 5691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
class_names = ["T-shirt/top", "Trouser", "Pullover", "Dress", "Coat", "Sandal", "Shirt", "Sneaker", "Bag", "Ankle boot"]
# + [markdown] id="pRp3Inb6TB1h"
# ## **Changes**
# * **Input Shape :** (28 x 28)
#
# * **Output Shape :** 10 (10 differesent classes)
#
# * **Loss function :** CatgoricalCrosstentropy() for OneHot Encoded else use sparse
#
# * **Output Layer Activation :** Softmax
#
# Also add a Input Flatten layer
# + colab={"base_uri": "https://localhost:8080/"} id="1r7g26_aGrYC" executionInfo={"status": "ok", "timestamp": 1620492260650, "user_tz": -330, "elapsed": 43210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="099d9eda-5169-4ee6-a0a9-4a2b6d828bc5"
tf.random.set_seed(42)
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(4, activation='relu'),
tf.keras.layers.Dense(4, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax'),
])
model.compile(loss=tf.keras.losses.SparseCategoricalCrossentropy(),
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
history = model.fit(train_X, train_y, epochs = 10)
# + id="qnUk3xqRUp52" executionInfo={"status": "ok", "timestamp": 1620492261244, "user_tz": -330, "elapsed": 43796, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
train_X = train_X / 255.0
test_X = test_X / 255.0
# + colab={"base_uri": "https://localhost:8080/"} id="1bM0hxXxZx5C" executionInfo={"status": "ok", "timestamp": 1620492296290, "user_tz": -330, "elapsed": 78835, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="eb78f487-0b16-4ae7-894b-c06f769cb9aa"
# Set random seed
tf.random.set_seed(42)
# Create the model
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)), # input layer (we had to reshape 28x28 to 784)
tf.keras.layers.Dense(4, activation="relu"),
tf.keras.layers.Dense(4, activation="relu"),
tf.keras.layers.Dense(10, activation="softmax") # output shape is 10, activation is softmax
])
# Compile the model
model.compile(loss=tf.keras.losses.SparseCategoricalCrossentropy(),
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Fit the model (to the normalized data)
norm_history = model.fit(train_X,
train_y,
epochs=10,
validation_data=(test_X, test_y))
# + colab={"base_uri": "https://localhost:8080/", "height": 543} id="qP7gGD5rpNF8" executionInfo={"status": "ok", "timestamp": 1620492308214, "user_tz": -330, "elapsed": 1196, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="a8464fd2-7e03-4384-c07b-8a6a87023098"
pd.DataFrame(history.history).plot(title="Non=Normalized Data")
pd.DataFrame(norm_history.history).plot(title="Normalized Data");
# + colab={"base_uri": "https://localhost:8080/"} id="7Zuvwql5p9W9" executionInfo={"status": "ok", "timestamp": 1620492589254, "user_tz": -330, "elapsed": 140339, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="99a7ecb4-0a43-4730-85dc-ee8567aaa304"
# Set random seed
tf.random.set_seed(42)
# Create the model
model_13 = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)), # input layer (we had to reshape 28x28 to 784)
tf.keras.layers.Dense(4, activation="relu"),
tf.keras.layers.Dense(4, activation="relu"),
tf.keras.layers.Dense(10, activation="softmax") # output shape is 10, activation is softmax
])
# Compile the model
model_13.compile(loss=tf.keras.losses.SparseCategoricalCrossentropy(),
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# Create the learning rate callback
lr_scheduler = tf.keras.callbacks.LearningRateScheduler(lambda epoch: 1e-3 * 10**(epoch/20))
# Fit the model
find_lr_history = model_13.fit(train_X,
train_y,
epochs=40, # model already doing pretty good with current LR, probably don't need 100 epochs
validation_data=(test_X, test_y),
callbacks=[lr_scheduler])
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="PUvZ4CRGp77M" executionInfo={"status": "ok", "timestamp": 1620493309282, "user_tz": -330, "elapsed": 1228, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="ae867679-f325-4039-b46d-d90ffac4825f"
plt.semilogx(find_lr_history.history["lr"], find_lr_history.history["loss"]); # want the x-axis to be log-scale
plt.xlabel("Learning rate")
plt.ylabel("Loss")
plt.title("Finding the ideal learning rate");
# + colab={"base_uri": "https://localhost:8080/"} id="7vOBP3s38J1D" executionInfo={"status": "ok", "timestamp": 1620493864062, "user_tz": -330, "elapsed": 1133, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="3a793690-7d27-455c-de0d-9b0c832567df"
y_preds = (model.predict(test_X))
y_preds = y_preds.argmax(axis=1)
y_preds
# + id="CLgjjeY481OO" executionInfo={"status": "ok", "timestamp": 1620494065276, "user_tz": -330, "elapsed": 804, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(test_y, y_preds)
# + colab={"base_uri": "https://localhost:8080/", "height": 590} id="PZXGQHlg9_mW" executionInfo={"status": "ok", "timestamp": 1620494301519, "user_tz": -330, "elapsed": 1884, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="02cdaa47-a2f5-4eab-a1f5-6d24e3e1959e"
import seaborn as sn
plt.figure(figsize = (10, 10))
sn.heatmap(cm, annot=cm, fmt="", annot_kws={"size": 20}, cmap='Blues');
# + id="6dRFr2iQ-Gpo" executionInfo={"status": "ok", "timestamp": 1620494869344, "user_tz": -330, "elapsed": 819, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
import random
# Create a function for plotting a random image along with its prediction
def plot_random_image(model, images, true_labels, classes):
"""Picks a random image, plots it and labels it with a predicted and truth label.
Args:
model: a trained model (trained on data similar to what's in images).
images: a set of random images (in tensor form).
true_labels: array of ground truth labels for images.
classes: array of class names for images.
Returns:
A plot of a random image from `images` with a predicted class label from `model`
as well as the truth class label from `true_labels`.
"""
# Setup random integer
i = random.randint(0, len(images))
# Create predictions and targets
target_image = images[i]
pred_probs = model.predict(target_image.reshape(1, 28, 28)) # have to reshape to get into right size for model
pred_label = classes[pred_probs.argmax()]
true_label = classes[true_labels[i]]
# Plot the target image
plt.imshow(target_image, cmap=plt.cm.binary)
# Change the color of the titles depending on if the prediction is right or wrong
if pred_label == true_label:
color = "green"
else:
color = "red"
# Add xlabel information (prediction/true label)
plt.xlabel("Pred: {} {:2.0f}% (True: {})".format(pred_label,
100*tf.reduce_max(pred_probs),
true_label),
color=color) # set the color to green or red
# + colab={"base_uri": "https://localhost:8080/", "height": 277} id="2zmTHfIABV_n" executionInfo={"status": "ok", "timestamp": 1620495052643, "user_tz": -330, "elapsed": 1306, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}} outputId="effc39b9-cbb0-403a-d7e7-3c10c77ed9d4"
# Check out a random image as well as its prediction
plot_random_image(model=model,
images=test_X,
true_labels=test_y,
classes=class_names)
# + id="0Ppy5sx9Bbfw" executionInfo={"status": "ok", "timestamp": 1620495010094, "user_tz": -330, "elapsed": 1030, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSL7-vyK3M9O2aew1rAxVHgD8h6RoC4uGUerXZL5s=s64", "userId": "04927215883267902569"}}
| notebooks/mini-projects/keras-fashion_mnist-multi_class-classification_tf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CK+ Results
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
pd.options.display.float_format = "{:,.2f}".format
# -
def extract_metrics_gpu(filename):
f = open(filename, 'r')
lines = f.readlines()
ram_init = None
ram_peak = None
cpu_init = None
cpu_end = None
for line in lines:
if "=> Average precision" in line:
accuracy = float(line.split(" ")[-1])
if "=> Total training time" in line:
training_time = float(line.split(" ")[-2])
if "RAM BEFORE TRAINING:" in line:
ram_init = float(line.split(" ")[-1])
if "PEAK TRAINING RAM:" in line:
ram_peak = float(line.split(" ")[-1])
if "CPU BEFORE TRAINING:" in line:
cpu_init = eval(line.split(":")[-1])
if "CPU BEFORE EVALUATION:" in line:
cpu_end = eval(line.split(":")[-1])
if "GPU BEFORE EVALUATION:" in line:
gpu_u, gpu_mem = eval(line.split(":")[-1])
cpu_u, cpu_k = (cpu_end[0] - cpu_init[0], cpu_end[1] - cpu_init[1])
return (accuracy * 100, training_time, (ram_peak-ram_init) * 1024, cpu_u + cpu_k, gpu_u, gpu_mem)
def calculate_metrics_gpu(files):
metrics = dict()
for file in files:
method, seed = file.split("_")
if method in metrics:
metrics[method].append(extract_metrics_gpu(file))
else:
metrics[method] = [extract_metrics_gpu(file)]
return metrics
def get_means(metrics):
return {method: [sum(x)/len(x) for x in list(zip(*metrics[method]))] for method in metrics}
def get_sds(metrics):
return {method: [np.std(x) for x in list(zip(*metrics[method]))] for method in metrics}
def split_orderings(metrics):
res = {}
for m in metrics:
for i in range(3):
res[m + str(i + 1)] = metrics[m][i]
return res
# ## Task-IL Results
# %cd ~/Desktop/research_project_repo/research_project/experiments/ckplus/2021-05-01-15-30/
# logs = !ls
ckplus_metrics = calculate_metrics_gpu(logs)
ckplus_split_metrics = split_orderings(ckplus_metrics)
df = pd.DataFrame(ckplus_split_metrics, index=['Accuracy (%)',
'Training time (s)',
'RAM Usage (MB)',
'CPU Usage (s)',
'GPU Usage (%)',
'GPU Memory (MB)'])
methods = ["nr", "lr", "gr", "lgr", "grd", "lgrd"]
order = []
for method in methods:
for i in range(1, 4):
order.append(method + str(i))
df.filter(order)
# ## Class-IL Results
# %cd ~/Desktop/research_project_repo/research_project/experiments/ckplus_class/2021-05-10-02-00/
# logs = !ls
ckplus_metrics = calculate_metrics_gpu(logs)
ckplus_split_metrics = split_orderings(ckplus_metrics)
df = pd.DataFrame(ckplus_split_metrics, index=['Accuracy (%)',
'Training time (s)',
'RAM Usage (MB)',
'CPU Usage (s)',
'GPU Usage (%)',
'GPU Memory (MB)'])
methods = ["nr", "lr", "gr", "lgr", "grd", "lgrd"]
order = []
for method in methods:
for i in range(1, 4):
order.append(method + str(i))
df.filter(order)
| analysis/CKPLUS_Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/Mustapha-AJEGHRIR/medical_txt_parser/blob/main/src/notebooks/assertions_nli/ast_nli_scibert.ipynb)
# + [markdown] id="4l_yTcIgxsd_"
# # Relations classification
#
# Based of: https://github.com/huggingface/transformers/blob/master/examples/pytorch/text-classification/run_xnli.py
# -
# !nvidia-smi
# + executionInfo={"elapsed": 4620, "status": "ok", "timestamp": 1642252265905, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg753z6h9fmTPmGyKajJFbNQG48KIqPziiTsxl4Tw=s64", "userId": "11345629174419407363"}, "user_tz": -60} id="LwHxhQx0xseE"
# %%capture
# !pip install seqeval transformers datasets spacy sentence_transformers
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1908, "status": "ok", "timestamp": 1642251232672, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg753z6h9fmTPmGyKajJFbNQG48KIqPziiTsxl4Tw=s64", "userId": "11345629174419407363"}, "user_tz": -60} id="3rCJwcFyyU9J" outputId="f75daf2c-9611-4b10-faf6-dd37b6e411f5"
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive/MyDrive/projects/medical_txt_parser
# + executionInfo={"elapsed": 1358, "status": "ok", "timestamp": 1642251234969, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg753z6h9fmTPmGyKajJFbNQG48KIqPziiTsxl4Tw=s64", "userId": "11345629174419407363"}, "user_tz": -60} id="FyDbgPv1xseF"
# %reload_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings("ignore")
# path = %pwd
while "src" in path:
# %cd ..
# path = %pwd
import glob
import pandas as pd
import os
import numpy as np
from tqdm import tqdm
from pprint import pprint
import matplotlib.pyplot as plt
import random
import logging
import os
import random
import sys
from dataclasses import dataclass, field
from typing import Optional
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
import datasets
import numpy as np
from datasets import load_dataset, load_metric , Dataset
import transformers
from transformers import (
AutoConfig,
AutoModelForSequenceClassification,
AutoTokenizer,
DataCollatorWithPadding,
EvalPrediction,
HfArgumentParser,
Trainer,
TrainingArguments,
default_data_collator,
set_seed,
)
from transformers.trainer_utils import get_last_checkpoint
from transformers.utils import check_min_version
from transformers.utils.versions import require_version
from transformers import pipeline
require_version("datasets>=1.8.0", "To fix: pip install --upgrade datasets")
from src.utils.parse_data import parse_ast, parse_concept, parse_relation
# + executionInfo={"elapsed": 219, "status": "ok", "timestamp": 1642251245139, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg753z6h9fmTPmGyKajJFbNQG48KIqPziiTsxl4Tw=s64", "userId": "11345629174419407363"}, "user_tz": -60} id="Sp1UC7psxseH"
train_data_path = "data/train"
val_data_path = "data/val"
ast_folder_name = "ast"
concept_folder_name = "concept"
rel_folder_name = "rel"
txt_folder_name = "txt"
nli_data_path = "data/nli"
re_data_path = "data/re"
# model args
model_name_or_path = "allenai/scibert_scivocab_uncased" # "gsarti/scibert-nli" "allenai/scibert_scivocab_uncased" "models/scibert_scivocab_uncased-re-1"
cache_dir = None
model_revision = None
tokenizer_name = model_name_or_path
do_lower_case = None
use_fast_tokenizer = True
fp16 = True
# data args
pad_to_max_length = None
max_seq_length = None
set_seed(42)
# -
# ## Model Test - Problem
# + [markdown] id="zP8CizR4xseH"
# ### Import data
# -
re_task = "Te_P"
relations_df = pd.read_csv(re_data_path + os.sep + f"re_scibert_data_{re_task}.tsv", sep="\t", header=None)
relations_df.columns = ["text", "label"]
label2id = {label: i for i, label in enumerate(relations_df["label"].value_counts().index.tolist())}
id2label = {i: label for label, i in label2id.items()}
relations_df["label"] = relations_df.label.map(label2id)
relations_df
# +
# Build HuggingFace Dataset
train_df, val_df = train_test_split(relations_df, train_size=None, shuffle=True, test_size=10, stratify=relations_df["label"], random_state=42)
features = datasets.Features({'text': datasets.Value(dtype='string'),
'label': datasets.ClassLabel(num_classes=len(id2label), names=list(id2label.values()))})
train_dataset = Dataset.from_pandas(train_df, preserve_index=False, features=features)
eval_dataset = Dataset.from_pandas(val_df, preserve_index=False, features=features)
label_list = train_dataset.features["label"].names
num_labels = len(label_list)
label_list = train_dataset.features["label"].names
train_dataset, eval_dataset
# -
# check labels balance
print(f"train_df labels: {train_df['label'].value_counts()}")
print(f"val_df labels: {val_df['label'].value_counts()}")
# Load pretrained model and tokenizer
# In distributed training, the .from_pretrained methods guarantee that only one local process can concurrently
# download model & vocab.
config = AutoConfig.from_pretrained(
model_name_or_path,
num_labels=num_labels,
finetuning_task="re",
cache_dir=cache_dir,
revision=model_revision,
label2id=label2id,
id2label=id2label
)
tokenizer = AutoTokenizer.from_pretrained(
tokenizer_name if tokenizer_name else model_name_or_path,
# do_lower_case=do_lower_case,
cache_dir=cache_dir,
use_fast=use_fast_tokenizer,
revision=model_revision,
)
model = AutoModelForSequenceClassification.from_pretrained(
model_name_or_path,
from_tf=bool(".ckpt" in model_name_or_path),
config=config,
cache_dir=cache_dir,
revision=model_revision
)
# +
# Preprocessing the datasets
# Padding strategy
if pad_to_max_length:
padding = "max_length"
else:
# We will pad later, dynamically at batch creation, to the max sequence length in each batch
padding = False
def preprocess_function(examples):
# Tokenize the texts
return tokenizer(
examples["text"],
padding=padding,
max_length=max_seq_length,
truncation=True,
)
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on train dataset",
)
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on validation dataset",
)
# Log a few random samples from the training set:
for index in random.sample(range(len(train_dataset)), 3):
print(f"Sample {index} of the training set: {train_dataset[index]}.\n")
# +
# Get the metric function
f1_metric = load_metric("f1")
precision_metric = load_metric("precision")
recall_metric = load_metric("recall")
accuracy_metric = load_metric("accuracy")
# You can define your custom compute_metrics function. It takes an `EvalPrediction` object (a namedtuple with a
# predictions and label_ids field) and has to return a dictionary string to float.
def compute_metrics(p: EvalPrediction):
preds = p.predictions[0] if isinstance(p.predictions, tuple) else p.predictions
preds = np.argmax(preds, axis=1)
metrics = {}
metrics.update(f1_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(precision_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(recall_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(accuracy_metric.compute(predictions=preds, references=p.label_ids))
return metrics
# -
# Data collator will default to DataCollatorWithPadding, so we change it if we already did the padding.
if pad_to_max_length:
data_collator = default_data_collator
elif fp16:
data_collator = DataCollatorWithPadding(tokenizer, pad_to_multiple_of=8)
else:
data_collator = None
model_name_or_path
# ### Training
# +
# address class imbalance
import torch
from torch import nn
from transformers import Trainer
from sklearn.utils.class_weight import compute_class_weight
class_weights = compute_class_weight(class_weight='balanced', classes=list(id2label.keys()),y=train_df["label"])
class_weights = torch.tensor(class_weights).log1p()
class CustomTrainer(Trainer):
def compute_loss(self, model, inputs, return_outputs=False):
labels = inputs.get("labels")
# forward pass
outputs = model(**inputs)
logits = outputs.get("logits")
loss_fct = nn.CrossEntropyLoss(weight=torch.tensor(class_weights).float().to(logits.device))
loss = loss_fct(logits.view(-1, self.model.config.num_labels), labels.view(-1))
return (loss, outputs) if return_outputs else loss
# +
# Initialize our Trainer
model_folder_name = f"{model_name_or_path.split('/')[-1]}-re-{re_task}-1"
args = TrainingArguments(
f"training_logs/{model_folder_name}",
evaluation_strategy = "epoch",
save_strategy = "no",
learning_rate=3e-5,
per_device_train_batch_size=32,
per_device_eval_batch_size=32,
num_train_epochs=10,
weight_decay=0.05,
logging_steps=1,
warmup_ratio=0.1,
)
trainer = CustomTrainer(
model=model,
args=args,
train_dataset=train_dataset ,
eval_dataset=eval_dataset ,
compute_metrics=compute_metrics,
tokenizer=tokenizer,
data_collator=data_collator,
)
# -
train_result = trainer.train()
# +
metrics = train_result.metrics
metrics["train_samples"] = len(train_dataset)
trainer.save_model(f"models/{model_folder_name}") # Saves the tokenizer too for easy upload
trainer.log_metrics("train", metrics)
trainer.save_metrics("train", metrics)
# +
print("*** Evaluate ***")
metrics = trainer.evaluate(eval_dataset=eval_dataset)
metrics["eval_samples"] = len(eval_dataset)
trainer.log_metrics("eval", metrics)
trainer.save_metrics("eval", metrics)
# -
predictions, labels, _ = trainer.predict(eval_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
predictions, labels, _ = trainer.predict(train_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
# ## Model Treatment - Problem
# + [markdown] id="zP8CizR4xseH"
# ### Import data
# -
re_task = "Tr_P"
import torch
# empty cuda
torch.cuda.empty_cache()
relations_df = pd.read_csv(re_data_path + os.sep + f"re_scibert_data_{re_task}.tsv", sep="\t", header=None)
relations_df.columns = ["text", "label"]
label2id = {label: i for i, label in enumerate(relations_df["label"].value_counts().index.tolist())}
id2label = {i: label for label, i in label2id.items()}
relations_df["label"] = relations_df.label.map(label2id)
relations_df
# +
# Build HuggingFace Dataset
train_df, val_df = train_test_split(relations_df, train_size=None, shuffle=True, test_size=10, stratify=relations_df["label"], random_state=42)
features = datasets.Features({'text': datasets.Value(dtype='string'),
'label': datasets.ClassLabel(num_classes=len(id2label), names=list(id2label.values()))})
train_dataset = Dataset.from_pandas(train_df, preserve_index=False, features=features)
eval_dataset = Dataset.from_pandas(val_df, preserve_index=False, features=features)
label_list = train_dataset.features["label"].names
num_labels = len(label_list)
label_list = train_dataset.features["label"].names
train_dataset, eval_dataset
# -
# check labels balance
print(f"train_df labels: {train_df['label'].value_counts()}")
print(f"val_df labels: {val_df['label'].value_counts()}")
# Load pretrained model and tokenizer
# In distributed training, the .from_pretrained methods guarantee that only one local process can concurrently
# download model & vocab.
config = AutoConfig.from_pretrained(
model_name_or_path,
num_labels=num_labels,
finetuning_task="re",
cache_dir=cache_dir,
revision=model_revision,
label2id=label2id,
id2label=id2label
)
tokenizer = AutoTokenizer.from_pretrained(
tokenizer_name if tokenizer_name else model_name_or_path,
# do_lower_case=do_lower_case,
cache_dir=cache_dir,
use_fast=use_fast_tokenizer,
revision=model_revision,
)
model = AutoModelForSequenceClassification.from_pretrained(
model_name_or_path,
from_tf=bool(".ckpt" in model_name_or_path),
config=config,
cache_dir=cache_dir,
revision=model_revision,
)
# +
# Preprocessing the datasets
# Padding strategy
if pad_to_max_length:
padding = "max_length"
else:
# We will pad later, dynamically at batch creation, to the max sequence length in each batch
padding = False
def preprocess_function(examples):
# Tokenize the texts
return tokenizer(
examples["text"],
padding=padding,
max_length=max_seq_length,
truncation=True,
)
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on train dataset",
)
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on validation dataset",
)
# Log a few random samples from the training set:
for index in random.sample(range(len(train_dataset)), 3):
print(f"Sample {index} of the training set: {train_dataset[index]}.\n")
# +
# Get the metric function
f1_metric = load_metric("f1")
precision_metric = load_metric("precision")
recall_metric = load_metric("recall")
accuracy_metric = load_metric("accuracy")
# You can define your custom compute_metrics function. It takes an `EvalPrediction` object (a namedtuple with a
# predictions and label_ids field) and has to return a dictionary string to float.
def compute_metrics(p: EvalPrediction):
preds = p.predictions[0] if isinstance(p.predictions, tuple) else p.predictions
preds = np.argmax(preds, axis=1)
metrics = {}
metrics.update(f1_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(precision_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(recall_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(accuracy_metric.compute(predictions=preds, references=p.label_ids))
return metrics
# -
# Data collator will default to DataCollatorWithPadding, so we change it if we already did the padding.
if pad_to_max_length:
data_collator = default_data_collator
elif fp16:
data_collator = DataCollatorWithPadding(tokenizer, pad_to_multiple_of=8)
else:
data_collator = None
model_name_or_path
# ### Training
# +
# address class imbalance
import torch
from torch import nn
from transformers import Trainer
class_weights = [len(train_df)/ (len(train_df[train_df["label"] == i])*len(id2label)) for i in id2label.keys()]
# apply log to weights
class_weights = torch.tensor(class_weights).log1p()
print(class_weights)
class CustomTrainer(Trainer):
def compute_loss(self, model, inputs, return_outputs=False):
labels = inputs.get("labels")
# forward pass
outputs = model(**inputs)
logits = outputs.get("logits")
loss_fct = nn.CrossEntropyLoss(weight=torch.tensor(class_weights).float().to(logits.device))
loss = loss_fct(logits.view(-1, self.model.config.num_labels), labels.view(-1))
return (loss, outputs) if return_outputs else loss
# +
# Initialize our Trainer
model_folder_name = f"{model_name_or_path.split('/')[-1]}-re-{re_task}-1"
args = TrainingArguments(
f"training_logs/{model_folder_name}",
evaluation_strategy = "epoch",
save_strategy = "no",
learning_rate=3e-5,
per_device_train_batch_size=32,
per_device_eval_batch_size=32,
num_train_epochs=10,
weight_decay=0.05,
logging_steps=1,
warmup_ratio=0.1,
)
trainer = CustomTrainer(
model=model,
args=args,
train_dataset=train_dataset ,
eval_dataset=eval_dataset ,
compute_metrics=compute_metrics,
tokenizer=tokenizer,
data_collator=data_collator,
)
# -
train_result = trainer.train()
# +
metrics = train_result.metrics
metrics["train_samples"] = len(train_dataset)
trainer.save_model(f"models/{model_folder_name}") # Saves the tokenizer too for easy upload
trainer.log_metrics("train", metrics)
trainer.save_metrics("train", metrics)
# +
print("*** Evaluate ***")
metrics = trainer.evaluate(eval_dataset=eval_dataset)
metrics["eval_samples"] = len(eval_dataset)
trainer.log_metrics("eval", metrics)
trainer.save_metrics("eval", metrics)
# -
predictions, labels, _ = trainer.predict(eval_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
predictions, labels, _ = trainer.predict(train_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
# empty cuda cache
import torch
torch.cuda.empty_cache()
# !nvidia-smi
# ## Model Problem - Problem
# + [markdown] id="zP8CizR4xseH"
# ### Import data
# -
re_task = "P_P"
relations_df = pd.read_csv(re_data_path + os.sep + f"re_scibert_data_{re_task}.tsv", sep="\t", header=None)
relations_df.columns = ["text", "label"]
label2id = {label: i for i, label in enumerate(relations_df["label"].value_counts().index.tolist())}
id2label = {i: label for label, i in label2id.items()}
relations_df["label"] = relations_df.label.map(label2id)
relations_df
# +
# Build HuggingFace Dataset
train_df, val_df = train_test_split(relations_df, train_size=None, shuffle=True, test_size=10, stratify=relations_df["label"], random_state=42)
features = datasets.Features({'text': datasets.Value(dtype='string'),
'label': datasets.ClassLabel(num_classes=len(id2label), names=list(id2label.values()))})
train_dataset = Dataset.from_pandas(train_df, preserve_index=False, features=features)
eval_dataset = Dataset.from_pandas(val_df, preserve_index=False, features=features)
label_list = train_dataset.features["label"].names
num_labels = len(label_list)
label_list = train_dataset.features["label"].names
train_dataset, eval_dataset
# -
# check labels balance
print(f"train_df labels: {train_df['label'].value_counts()}")
print(f"val_df labels: {val_df['label'].value_counts()}")
# Load pretrained model and tokenizer
# In distributed training, the .from_pretrained methods guarantee that only one local process can concurrently
# download model & vocab.
config = AutoConfig.from_pretrained(
model_name_or_path,
num_labels=num_labels,
finetuning_task="re",
cache_dir=cache_dir,
revision=model_revision,
label2id=label2id,
id2label=id2label
)
tokenizer = AutoTokenizer.from_pretrained(
tokenizer_name if tokenizer_name else model_name_or_path,
# do_lower_case=do_lower_case,
cache_dir=cache_dir,
use_fast=use_fast_tokenizer,
revision=model_revision,
)
model = AutoModelForSequenceClassification.from_pretrained(
model_name_or_path,
from_tf=bool(".ckpt" in model_name_or_path),
config=config,
cache_dir=cache_dir,
revision=model_revision,
)
# +
# Preprocessing the datasets
# Padding strategy
if pad_to_max_length:
padding = "max_length"
else:
# We will pad later, dynamically at batch creation, to the max sequence length in each batch
padding = False
def preprocess_function(examples):
# Tokenize the texts
return tokenizer(
examples["text"],
padding=padding,
max_length=max_seq_length,
truncation=True,
)
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on train dataset",
)
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on validation dataset",
)
# Log a few random samples from the training set:
for index in random.sample(range(len(train_dataset)), 3):
print(f"Sample {index} of the training set: {train_dataset[index]}.\n")
# +
# Get the metric function
f1_metric = load_metric("f1")
precision_metric = load_metric("precision")
recall_metric = load_metric("recall")
accuracy_metric = load_metric("accuracy")
# You can define your custom compute_metrics function. It takes an `EvalPrediction` object (a namedtuple with a
# predictions and label_ids field) and has to return a dictionary string to float.
def compute_metrics(p: EvalPrediction):
preds = p.predictions[0] if isinstance(p.predictions, tuple) else p.predictions
preds = np.argmax(preds, axis=1)
metrics = {}
metrics.update(f1_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(precision_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(recall_metric.compute(predictions=preds, references=p.label_ids, average="macro"))
metrics.update(accuracy_metric.compute(predictions=preds, references=p.label_ids))
return metrics
# -
# Data collator will default to DataCollatorWithPadding, so we change it if we already did the padding.
if pad_to_max_length:
data_collator = default_data_collator
elif fp16:
data_collator = DataCollatorWithPadding(tokenizer, pad_to_multiple_of=8)
else:
data_collator = None
model_name_or_path
# ### Training
# +
# address class imbalance
import torch
from torch import nn
from transformers import Trainer
class_weights = [len(train_df)/ (len(train_df[train_df["label"] == i])*len(id2label)) for i in id2label.keys()]
class_weights = torch.tensor(class_weights).log1p()
print(class_weights)
class CustomTrainer(Trainer):
def compute_loss(self, model, inputs, return_outputs=False):
labels = inputs.get("labels")
# forward pass
outputs = model(**inputs)
logits = outputs.get("logits")
loss_fct = nn.CrossEntropyLoss(weight=torch.tensor(class_weights).float().to(logits.device))
loss = loss_fct(logits.view(-1, self.model.config.num_labels), labels.view(-1))
return (loss, outputs) if return_outputs else loss
# +
# Initialize our Trainer
model_folder_name = f"{model_name_or_path.split('/')[-1]}-re-{re_task}-1"
args = TrainingArguments(
f"training_logs/{model_folder_name}",
evaluation_strategy = "epoch",
save_strategy = "no",
learning_rate=3e-5,
per_device_train_batch_size=32,
per_device_eval_batch_size=32,
num_train_epochs=4,
weight_decay=0.05,
logging_steps=1,
warmup_ratio=0.1,
)
trainer = CustomTrainer(
model=model,
args=args,
train_dataset=train_dataset ,
eval_dataset=eval_dataset ,
compute_metrics=compute_metrics,
tokenizer=tokenizer,
data_collator=data_collator,
)
# -
train_result = trainer.train()
# +
metrics = train_result.metrics
metrics["train_samples"] = len(train_dataset)
trainer.save_model(f"models/{model_folder_name}") # Saves the tokenizer too for easy upload
trainer.log_metrics("train", metrics)
trainer.save_metrics("train", metrics)
# +
print("*** Evaluate ***")
metrics = trainer.evaluate(eval_dataset=eval_dataset)
metrics["eval_samples"] = len(eval_dataset)
trainer.log_metrics("eval", metrics)
trainer.save_metrics("eval", metrics)
# -
predictions, labels, _ = trainer.predict(eval_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
predictions, labels, _ = trainer.predict(train_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
print(classification_report(labels, predictions,target_names=label_list))
# ## Final Predictions
val_data_path = "data/test"
# +
text_files = glob.glob(val_data_path + os.sep + txt_folder_name + os.sep + "*.txt")
filename = ""
df = pd.DataFrame()
for file in tqdm(text_files):
with open(file, 'r') as f:
text = f.read()
# split lines
lines = text.split('\n')
filename =[ file.split("/")[-1].split(".")[0]] * len(lines)
df = df.append(pd.DataFrame({"text": lines, "filename": filename, "line_number": range(len(lines))}), ignore_index=True)
df = df.sort_values(by=["filename", "line_number"])
# remove empty text lines
# df = df[df.text != ""]
# df = df.reset_index(drop=True)
# add concepts
rel_df = pd.DataFrame()
for fname in tqdm(df["filename"].unique()):
concept_dict = parse_concept(val_data_path + os.sep + concept_folder_name + os.sep + fname + ".con")
concept_df = pd.DataFrame(concept_dict).drop(columns=["end_line"])
test_concept_df = concept_df[concept_df["concept_type"] == "test"]
problem_concept_df = concept_df[concept_df["concept_type"] == "problem"]
treatment_concept_df = concept_df[concept_df["concept_type"] == "treatment"]
# class test --> problem
test_problem_df = pd.merge(test_concept_df, problem_concept_df, how="inner", on="start_line")
# class treatment --> problem
treatment_problem_df = pd.merge(treatment_concept_df, problem_concept_df, how="inner", on="start_line")
# class problem --> problem
problem_problem_df = pd.merge(problem_concept_df, problem_concept_df, how="inner", on="start_line")
problem_problem_df = problem_problem_df[problem_problem_df["concept_text_x"] != problem_problem_df["concept_text_y"]] # TODO: remove duplicates ?
tmp = pd.concat([test_problem_df, treatment_problem_df, problem_problem_df], axis=0)
tmp["filename"] = fname
rel_df = rel_df.append(tmp, ignore_index=True)
rel_df = rel_df.sort_values(by=["filename", "start_line"])
rel_df = rel_df.reset_index(drop=True)
rel_df = rel_df[[ "filename", "start_line", "concept_text_x", "concept_text_y", "concept_type_x", "concept_type_y", "start_word_number_x", "end_word_number_x", "start_word_number_y", "end_word_number_y"]]
rel_df
# +
# make predict dataset
def preprocess_text(row):
# find line
line = df[(df["filename"] == row["filename"]) & (df["line_number"] == row["start_line"]-1)]["text"].values[0]
# line = line.lower()
line = " ".join(line.split()) # remove multiple spaces
concept_text_x = "<< "+ " ".join(line.split()[row["start_word_number_x"]:row["end_word_number_x"]+1]) + " >>"
concept_text_y = "[[ " + " ".join(line.split()[row["start_word_number_y"]:row["end_word_number_y"]+1]) + " ]]"
start_word_number_x = row["start_word_number_x"]
end_word_number_x = row["end_word_number_x"]
start_word_number_y = row["start_word_number_y"]
end_word_number_y = row["end_word_number_y"]
if row["start_word_number_x"] > row["start_word_number_y"]:
concept_text_x, concept_text_y = concept_text_y, concept_text_x
start_word_number_x, start_word_number_y = start_word_number_y, start_word_number_x
end_word_number_x, end_word_number_y = end_word_number_y, end_word_number_x
text = " ".join(line.split()[: start_word_number_x] + [concept_text_x] + line.split()[end_word_number_x+1: start_word_number_y] + [concept_text_y] + line.split()[end_word_number_y+1:])
row["text"] = text
return row
predict_df = rel_df.apply(preprocess_text, axis=1)
predict_df
# -
orig_predict_df = predict_df.copy()
# +
re_task = "Tr_P"
if re_task == "P_P":
# problem --> problem
predict_df = orig_predict_df[(orig_predict_df["concept_type_x"] == "problem") & (orig_predict_df["concept_type_y"] == "problem")]
label2id = {'Other': 0, 'PIP': 1}
elif re_task == "Tr_P":
# treatment --> problem
predict_df = orig_predict_df[(orig_predict_df["concept_type_x"] == "treatment") & (orig_predict_df["concept_type_y"] == "problem")]
label2id = {'Other': 0, 'TrAP': 1, 'TrCP': 2, 'TrNAP': 3, 'TrIP': 4, 'TrWP': 5}
elif re_task == "Te_P":
# test --> problem
predict_df = orig_predict_df[(orig_predict_df["concept_type_x"] == "test") & (orig_predict_df["concept_type_y"] == "problem")]
label2id = {'Other': 0, 'TeRP': 1, 'TeCP': 2}
id2label = {v: k for k, v in label2id.items()}
model_folder_name = f"{model_name_or_path.split('/')[-1]}-re-{re_task}-1"
model = AutoModelForSequenceClassification.from_pretrained(f"models/{model_folder_name}", label2id=label2id, id2label=id2label)
tokenizer = AutoTokenizer.from_pretrained(f"models/{model_folder_name}")
# Data collator will default to DataCollatorWithPadding, so we change it if we already did the padding.
if pad_to_max_length:
data_collator = default_data_collator
elif fp16:
data_collator = DataCollatorWithPadding(tokenizer, pad_to_multiple_of=8)
else:
data_collator = None
# Initialize our Trainer
args = TrainingArguments(
f"training_logs/{model_folder_name}",
evaluation_strategy = "epoch",
save_strategy = "no",
learning_rate=1e-5,
per_device_train_batch_size=32,
per_device_eval_batch_size=32,
num_train_epochs=5,
weight_decay=0.05,
logging_steps=1,
warmup_ratio=0.1,
)
trainer = Trainer(
model=model,
# args=args,
# train_dataset=train_dataset ,
# eval_dataset=eval_dataset ,
# compute_metrics=compute_metrics,
tokenizer=tokenizer,
data_collator=data_collator,
)
# +
# Preprocessing the datasets
# Padding strategy
if pad_to_max_length:
padding = "max_length"
else:
# We will pad later, dynamically at batch creation, to the max sequence length in each batch
padding = False
def preprocess_function(examples):
# Tokenize the texts
return tokenizer(
examples["text"],
padding=padding,
max_length=max_seq_length,
truncation=True,
)
predict_dataset = Dataset.from_pandas(predict_df, preserve_index=False)
# predict_dataset = predict_dataset.select(range(10))
predict_dataset = predict_dataset.map(
preprocess_function,
batched=True,
desc="Running tokenizer on prediction dataset",
)
predict_dataset
# -
predictions, labels, metrics = trainer.predict(predict_dataset, metric_key_prefix="predict")
predictions = np.argmax(predictions, axis=1)
len(predictions)
predict_df["prediction"] = [id2label[label] for label in predictions]
rel_df.loc[predict_df.index, "prediction"] = predict_df["prediction"]
rel_df
rel_df["prediction"].value_counts()
# you can now set another re_task
# +
# for each file create <filename>.con
os.makedirs(val_data_path + os.sep + rel_folder_name, exist_ok=True)
# empty folder if exists
files = glob.glob(val_data_path + os.sep + rel_folder_name + os.sep + "*.rel")
for file in files:
os.remove(file)
for i, row in tqdm(rel_df.iterrows()):
filename = row["filename"]
concept_text_x = row["concept_text_x"].lower()
concept_text_y = row["concept_text_y"].lower()
concept_type_x = row["concept_type_x"]
concept_type_y = row["concept_type_y"]
start_word_number_x = row["start_word_number_x"]
end_word_number_x = row["end_word_number_x"]
start_word_number_y = row["start_word_number_y"]
end_word_number_y = row["end_word_number_y"]
line_number = row["start_line"]
prediction = row["prediction"]
if prediction != "Other":
with open(val_data_path + os.sep + rel_folder_name + os.sep + filename + ".rel", "a") as f:
# fill like this c="pefusion imaging" 19:6 19:7||r="TeRP"||c="perfusion defects" 19:12 19:13
f.write(
f"c=\"{concept_text_x}\" {line_number}:{start_word_number_x} {line_number}:{end_word_number_x}||r=\"{prediction}\"||c=\"{concept_text_y}\" {line_number}:{start_word_number_y} {line_number}:{end_word_number_y}\n"
)
# -
rel_files = glob.glob(val_data_path + os.sep + rel_folder_name + os.sep + "*.rel")
rel_files = [f.split(os.sep)[-1][:-4] for f in rel_files]
txt_files = [f.split(os.sep)[-1][:-4] for f in text_files]
# find missing files
missing_files = set(txt_files) - set(rel_files)
missing_files
# create empty files for missing files
for f in missing_files:
with open(val_data_path + os.sep + rel_folder_name + os.sep + f + ".rel", "w") as f:
f.write("")
rel_files = glob.glob(val_data_path + os.sep + rel_folder_name + os.sep + "*.rel")
rel_files = [f.split(os.sep)[-1][:-4] for f in rel_files]
txt_files = [f.split(os.sep)[-1][:-4] for f in text_files]
# find missing files
missing_files = set(txt_files) - set(rel_files)
missing_files
# !cat data/test/rel/0001.rel
# !zip -r scibert-test-rel-2-sep.zip data/test/rel/
| src/notebooks/relations_re/re_clf_scibert_separated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Artificial Intelligence with Python**
#
# # Lab Report 03: PANDAS
#
# * Name: <NAME>
# * CMS ID: 033-16-0017
#
# **Instructor: Dr. <NAME>**
# 1- Load the Titanic Dataset in Kaggle notebook from given link below:
# https://www.kaggle.com/hesh97/titanicdataset-traincsv
# Use the bar plot to show following plots:
#
# *PART_01*
#
# **Plot the Number of people survived and did not survive. Hint: Plot the counts values of**
# **“Survived” column of dataset.**
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
titanic_data = pd.read_csv('../input/titanicdataset-traincsv/train.csv')
sb.countplot(x = 'Survived',data=titanic_data)
titanic_data
# *PART_02*
#
# **Also plot survived comparison by class and by gender**
import pandas as pd
import seaborn as sb
titanic_data = pd.read_csv('../input/titanicdataset-traincsv/train.csv')
sb.countplot(x='Survived',data=titanic_data,palette = "Set2",hue='Sex')
sb.countplot(x='Survived',data=titanic_data,hue='Pclass')
# *PART_03*
#
# **Use the “groupby” function of Pandas to group the mean values (of all features) of**
# **passengers’’ gender and class.**
import pandas as pd
import seaborn as sb
titanic_data = pd.read_csv('../input/titanicdataset-traincsv/train.csv')
titanic_data.groupby('Sex').mean(),titanic_data.groupby('Pclass').mean()
#
#
# **Explore how to add and delete column from a data frame, explain and illustrate with an**
# **example. Also define concept of axis in pandas.**
#
# Answer:
# * Using DataFrame.insert() to add a column
# > df.insert(2, "Age", [21, 23, 24, 21], True),"2" is the position of the column, then column name and its data and set the inplace argument is True. If we change the inplace argument to False then we can't rename it.
# * Remove column name 'A', we use the drop() method. i.e. df.drop(['A'], axis = 1)
# * The axis argument is either 0 when it indicates rows and 1 when it is used to drop columns.
# **Generate a random data frame of 20 rows and 04 columns and change their columns**
# **names.**
import pandas as pd
import numpy as np
data = pd.DataFrame(np.random.rand(20,4))
data.columns = ['A','B','C','D']
data
# **Apply logical conditions to print values greater than 0.5 in any one column.**
import pandas as pd
import numpy as np
Data = pd.DataFrame(np.random.rand(20,4))
Data = Data[Data > 0.5]
Data
# **Explore sort function in Pandas, apply ascending and descending sorting according.**
import pandas as pd
import numpy as np
data = pd.DataFrame(np.random.rand(20,4))
data.columns = ['A','B','C','D']
#data of column A is sort in ascending Order and Column B sort in descending order.
data.sort_values(by='A', ascending=True),data.sort_values(by='B',ascending=False)
# **Loc and iloc are two functions of pandas for accessing the data from the data frame.**
# **Define the difference between them and use them each in at least 2 example**
# * loc is label-based, which means that we have to specify the name of the rows and columns that we need to filter out.
# * On the other hand, iloc is integer index-based. So here, we have to specify rows and columns by their integer index.
# > loc[row_label, column_label]
#
# > iloc[row_position, column_position]
#
# **Example 01: LOC[]**
# +
import pandas as pd
Data = pd.DataFrame({'A':[1,23,4,12],'B':[34,14,62,14],'C':["Asif","Sarwar",2,31],'D':[2,44,52,1.2]})
index = ['A','B','C','D']
Data.loc[:,'A']
# -
# **Example 02: LOC[]**
import pandas as pd
data = pd.read_csv('../input/world-happiness/2019.csv')
data.loc[1,'Score'],data
#
# **Example 01: iloc[]**
import pandas as pd
import numpy as np
Data = pd.read_csv('../input/world-happiness/2019.csv')
Data.iloc[:,1]
# **Example 02: iloc[]**
import pandas as pd
Data = pd.read_csv('../input/world-happiness/2019.csv')
Data.iloc[1,2],Data
| lab-03-exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
import sklearn
from pandas import Series, DataFrame
from pylab import rcParams
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
from sklearn import metrics
from sklearn.metrics import classification_report
# %matplotlib inline
rcParams['figure.figsize'] = 10, 8
sb.set_style('whitegrid')
# loading dataset
url = 'https://raw.githubusercontent.com/BigDataGal/Python-for-Data-Science/master/titanic-train.csv'
titanic = pd.read_csv(url)
titanic.columns = ['PassengerId','Survived','Pclass','Name','Sex','Age','SibSp','Parch','Ticket','Fare','Cabin','Embarked']
titanic.head()
sb.countplot(x='Survived',data=titanic, palette=['r','g']) # Pallet = Colors to use for the different levels of the hue variable
titanic.isnull().sum()
titanic.info()
titanic_data = titanic.drop(['PassengerId','Name','Ticket','Cabin'], 1)
titanic_data.head()
sb.boxplot(x='Pclass', y='Age', data=titanic_data, palette='hls')
titanic_data.head()
| 9. Exploratory Data Analysis (EDA).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# <style>
# pre {
# white-space: pre-wrap !important;
# }
# .table-striped > tbody > tr:nth-of-type(odd) {
# background-color: #f9f9f9;
# }
# .table-striped > tbody > tr:nth-of-type(even) {
# background-color: white;
# }
# .table-striped td, .table-striped th, .table-striped tr {
# border: 1px solid black;
# border-collapse: collapse;
# margin: 1em 2em;
# }
# .rendered_html td, .rendered_html th {
# text-align: left;
# vertical-align: middle;
# padding: 4px;
# }
# </style>
# -
# # Machine Learning with vaex.ml
#
# If you want to try out this notebook with a live Python kernel, use mybinder:
#
# <a class="reference external image-reference" href="https://mybinder.org/v2/gh/vaexio/vaex/latest?filepath=docs%2Fsource%2Ftutorial_ml.ipynb"><img alt="https://mybinder.org/badge_logo.svg" src="https://mybinder.org/badge_logo.svg" width="150px"></a>
#
#
# The `vaex.ml` package brings some machine learning algorithms to `vaex`. If you installed the individual subpackages (`vaex-core`, `vaex-hdf5`, ...) instead of the `vaex` metapackage, you may need to install it by running `pip install vaex-ml`, or `conda install -c conda-forge vaex-ml`.
#
# The API of `vaex.ml` stays close to that of [scikit-learn](https://scikit-learn.org/stable/), while providing better performance and the ability to efficiently perform operations on data that is larger than the available RAM. This page is an overview and a brief introduction to the capabilities offered by `vaex.ml`.
# +
import vaex
vaex.multithreading.thread_count_default = 8
import vaex.ml
import numpy as np
import pylab as plt
# -
# We will use the well known [Iris flower](https://en.wikipedia.org/wiki/Iris_flower_data_set) and Titanic passenger list datasets, two classical datasets for machine learning demonstrations.
df = vaex.ml.datasets.load_iris()
df
df.scatter(df.petal_length, df.petal_width, c_expr=df.class_);
# ## Preprocessing
#
# ### Scaling of numerical features
#
# `vaex.ml` packs the common numerical scalers:
#
# * `vaex.ml.StandardScaler` - Scale features by removing their mean and dividing by their variance;
# * `vaex.ml.MinMaxScaler` - Scale features to a given range;
# * `vaex.ml.RobustScaler` - Scale features by removing their median and scaling them according to a given percentile range;
# * `vaex.ml.MaxAbsScaler` - Scale features by their maximum absolute value.
#
# The usage is quite similar to that of `scikit-learn`, in the sense that each transformer implements the `.fit` and `.transform` methods.
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
scaler = vaex.ml.StandardScaler(features=features, prefix='scaled_')
scaler.fit(df)
df_trans = scaler.transform(df)
df_trans
# The output of the `.transform` method of any `vaex.ml` transformer is a _shallow copy_ of a DataFrame that contains the resulting features of the transformations in addition to the original columns. A shallow copy means that this new DataFrame just references the original one, and no extra memory is used. In addition, the resulting features, in this case the scaled numerical features are _virtual columns,_ which do not take any memory but are computed on the fly when needed. This approach is ideal for working with very large datasets.
# ### Encoding of categorical features
#
# `vaex.ml` contains several categorical encoders:
#
# * `vaex.ml.LabelEncoder` - Encoding features with as many integers as categories, startinfg from 0;
# * `vaex.ml.OneHotEncoder` - Encoding features according to the one-hot scheme;
# * `vaex.ml.FrequencyEncoder` - Encode features by the frequency of their respective categories;
# * `vaex.ml.BayesianTargetEncoder` - Encode categories with the mean of their target value;
# * `vaex.ml.WeightOfEvidenceEncoder` - Encode categories their weight of evidence value.
#
# The following is a quick example using the Titanic dataset.
df = vaex.ml.datasets.load_titanic()
df.head(5)
# +
label_encoder = vaex.ml.LabelEncoder(features=['embarked'])
one_hot_encoder = vaex.ml.OneHotEncoder(features=['embarked'])
freq_encoder = vaex.ml.FrequencyEncoder(features=['embarked'])
bayes_encoder = vaex.ml.BayesianTargetEncoder(features=['embarked'], target='survived')
woe_encoder = vaex.ml.WeightOfEvidenceEncoder(features=['embarked'], target='survived')
df = label_encoder.fit_transform(df)
df = one_hot_encoder.fit_transform(df)
df = freq_encoder.fit_transform(df)
df = bayes_encoder.fit_transform(df)
df = woe_encoder.fit_transform(df)
df.head(5)
# -
# Notice that the transformed features are all included in the resulting DataFrame and are appropriately named. This is excellent for the construction of various diagnostic plots, and engineering of more complex features. The fact that the resulting (encoded) features take no memory, allows one to try out or combine a variety of preprocessing steps without spending any extra memory.
# ## Feature Engineering
#
# ### KBinsDiscretizer
#
# With the `KBinsDiscretizer` you can convert a continous into a discrete feature by binning the data into specified intervals. You can specify the number of bins, the strategy on how to determine their size:
#
# * "uniform" - all bins have equal sizes;
# * "quantile" - all bins have (approximately) the same number of samples in them;
# * "kmeans" - values in each bin belong to the same 1D cluster as determined by the `KMeans` algorithm.
kbdisc = vaex.ml.KBinsDiscretizer(features=['age'], n_bins=5, strategy='quantile')
df = kbdisc.fit_transform(df)
df.head(5)
# ### GroupBy Transformer
#
# The `GroupByTransformer` is a handy feature in `vaex-ml` that lets you perform a groupby aggregations on the training data, and then use those aggregations as features in the training and test sets.
gbt = vaex.ml.GroupByTransformer(by='pclass', agg={'age': ['mean', 'std'],
'fare': ['mean', 'std'],
})
df = gbt.fit_transform(df)
df.head(5)
# ### CycleTransformer
#
# The `CycleTransformer` provides a strategy for transforming cyclical features, such as angles or time. This is done by considering each feature to be describing a polar coordinate system, and converting it to Cartesian coorindate system.
# This is shown to help certain ML models to achieve better performance.
df = vaex.from_arrays(days=[0, 1, 2, 3, 4, 5, 6])
cyctrans = vaex.ml.CycleTransformer(n=7, features=['days'])
cyctrans.fit_transform(df)
# ## Dimensionality reduction
#
# ### Principal Component Analysis
#
# The [PCA](https://en.wikipedia.org/wiki/Principal_component_analysis) implemented in `vaex.ml` can scale to a very large number of samples, even if that data we want to transform does not fit into RAM. To demonstrate this, let us do a PCA transformation on the Iris dataset. For this example, we have replicated this dataset thousands of times, such that it contains over **1 billion** samples.
# + tags=["skip-ci"]
df = vaex.ml.datasets.load_iris_1e9()
n_samples = len(df)
print(f'Number of samples in DataFrame: {n_samples:,}')
# + tags=["skip-ci"]
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
pca = vaex.ml.PCA(features=features, n_components=4)
pca.fit(df, progress='widget')
# -
# The PCA transformer implemented in `vaex.ml` can be fit in well under a minute, even when the data comprises 4 columns and 1 billion rows.
# + tags=["skip-ci"]
df_trans = pca.transform(df)
df_trans
# -
# Recall that the transformed DataFrame, which includes the PCA components, takes no extra memory.
# ### Incremental PCA
#
# The PCA implementation in vaex is very fast, but more so for "tall" DataFrames, i.e. DataFrames that have many rows, but not many columns. For DataFrames that have hundreds of columns, it is more efficient to use an Incremental PCA method. `vaex.ml` provides a convenient method that essentialy wraps `sklearn.decomposition.IncrementalPCA`, the fitting of which is more efficient for "wide" DataFrames.
#
# The usage is practically identical to the regular PCA method. Consider the following example:
# +
n_samples = 100_000
n_columns = 50
data_dict = {f'feat_{i}': np.random.normal(0, i+1, size=n_samples) for i in range(n_columns)}
df = vaex.from_dict(data_dict)
features = df.get_column_names()
pca = vaex.ml.PCAIncremental(n_components=10, features=features, batch_size=42_000)
pca.fit(df, progress='widget')
pca.transform(df)
# -
# Note that you need `scikit-learn` installed to only fit the `PCAIncremental` transformer. The the `transform` method does not rely on `scikit-learn` being installed.
#
# ### Random projections
#
# Random projections is another popular way of doing dimensionality reduction, especially when the dimensionality of the data is very high. `vaex.ml` conveniently wraps both `scikit-learn.random_projection.GaussianRandomProjection` and `scikit-learn.random_projection.SparseRandomProjection` in a single `vaex.ml` transformer.
rand_proj = vaex.ml.RandomProjections(features=features, n_components=10)
rand_proj.fit(df)
rand_proj.transform(df)
# ## Clustering
#
# ### K-Means
#
# `vaex.ml` implements a fast and scalable K-Means clustering algorithm. The usage is similar to that of `scikit-learn`.
# +
import vaex.ml.cluster
df = vaex.ml.datasets.load_iris()
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
kmeans = vaex.ml.cluster.KMeans(features=features, n_clusters=3, max_iter=100, verbose=True, random_state=42)
kmeans.fit(df)
df_trans = kmeans.transform(df)
df_trans
# -
# K-Means is an unsupervised algorithm, meaning that the predicted cluster labels in the transformed dataset do not necessarily correspond to the class label. We can map the predicted cluster identifiers to match the class labels, making it easier to construct diagnostic plots.
df_trans['predicted_kmean_map'] = df_trans.prediction_kmeans.map(mapper={0: 1, 1: 2, 2: 0})
df_trans
# Now we can construct simple scatter plots, and see that in the case of the Iris dataset, K-Means does a pretty good job splitting the data into 3 classes.
# +
fig = plt.figure(figsize=(12, 5))
plt.subplot(121)
df_trans.scatter(df_trans.petal_length, df_trans.petal_width, c_expr=df_trans.class_)
plt.title('Original classes')
plt.subplot(122)
df_trans.scatter(df_trans.petal_length, df_trans.petal_width, c_expr=df_trans.predicted_kmean_map)
plt.title('Predicted classes')
plt.tight_layout()
plt.show()
# -
# As with any algorithm implemented in `vaex.ml`, K-Means can be used on billions of samples. Fitting takes **under 2 minutes** when applied on the oversampled Iris dataset, numbering over **1 billion** samples.
# + tags=["skip-ci"]
df = vaex.ml.datasets.load_iris_1e9()
n_samples = len(df)
print(f'Number of samples in DataFrame: {n_samples:,}')
# + tags=["skip-ci"]
# %%time
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
kmeans = vaex.ml.cluster.KMeans(features=features, n_clusters=3, max_iter=100, verbose=True, random_state=31)
kmeans.fit(df)
# -
# ## Supervised learning
#
# While `vaex.ml` does not yet implement any supervised machine learning models, it does provide wrappers to several popular libraries such as [scikit-learn](https://scikit-learn.org/), [XGBoost](https://xgboost.readthedocs.io/), [LightGBM](https://lightgbm.readthedocs.io/) and [CatBoost](https://catboost.ai/).
#
# The main benefit of these wrappers is that they turn the models into `vaex.ml` transformers. This means the models become part of the DataFrame _state_ and thus can be serialized, and their predictions can be returned as _virtual columns_. This is especially useful for creating various diagnostic plots and evaluating performance metrics at no memory cost, as well as building ensembles.
#
# ### `Scikit-Learn` example
#
# The `vaex.ml.sklearn` module provides convenient wrappers to the `scikit-learn` estimators. In fact, these wrappers can be used with any library that follows the API convention established by `scikit-learn`, i.e. implements the `.fit` and `.transform` methods.
#
# Here is an example:
# +
from vaex.ml.sklearn import Predictor
from sklearn.ensemble import GradientBoostingClassifier
df = vaex.ml.datasets.load_iris()
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
target = 'class_'
model = GradientBoostingClassifier(random_state=42)
vaex_model = Predictor(features=features, target=target, model=model, prediction_name='prediction')
vaex_model.fit(df=df)
df = vaex_model.transform(df)
df
# -
# One can still train a predictive model on datasets that are too big to fit into memory by leveraging the on-line learners provided by `scikit-learn`. The `vaex.ml.sklearn.IncrementalPredictor` conveniently wraps these learners and provides control on how the data is passed to them from a `vaex` DataFrame.
#
# Let us train a model on the oversampled Iris dataset which comprises over 1 billion samples.
# + tags=["skip-ci"]
from vaex.ml.sklearn import IncrementalPredictor
from sklearn.linear_model import SGDClassifier
df = vaex.ml.datasets.load_iris_1e9()
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
target = 'class_'
model = SGDClassifier(learning_rate='constant', eta0=0.0001, random_state=42)
vaex_model = IncrementalPredictor(features=features, target=target, model=model,
batch_size=500_000, partial_fit_kwargs={'classes':[0, 1, 2]})
vaex_model.fit(df=df, progress='widget')
df = vaex_model.transform(df)
df
# -
# ### `XGBoost` example
#
# Libraries such as `XGBoost` provide more options such as validation during training and early stopping for example. We provide wrappers that keeps close to the native API of these libraries, in addition to the `scikit-learn` API.
#
# While the following example showcases the `XGBoost` wrapper, `vaex.ml` implements similar wrappers for `LightGBM` and `CatBoost`.
# +
from vaex.ml.xgboost import XGBoostModel
df = vaex.ml.datasets.load_iris_1e5()
df_train, df_test = df.ml.train_test_split(test_size=0.2, verbose=False)
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
target = 'class_'
params = {'learning_rate': 0.1,
'max_depth': 3,
'num_class': 3,
'objective': 'multi:softmax',
'subsample': 1,
'random_state': 42,
'n_jobs': -1}
booster = XGBoostModel(features=features, target=target, num_boost_round=500, params=params)
booster.fit(df=df_train, evals=[(df_train, 'train'), (df_test, 'test')], early_stopping_rounds=5)
df_test = booster.transform(df_train)
df_test
# -
# ### `CatBoost` example
#
# The CatBoost library supports summing up models. With this feature, we can use CatBoost to train a model using data that is otherwise too large to fit in memory. The idea is to train a single CatBoost model per chunk of data, and than sum up the invidiual models to create a master model. To use this feature via `vaex.ml` just specify the `batch_size` argument in the `CatBoostModel` wrapper. One can also specify additional options such as the strategy on how to sum up the individual models, or how they should be weighted.
# + tags=["skip-ci"]
from vaex.ml.catboost import CatBoostModel
df = vaex.ml.datasets.load_iris_1e8()
df_train, df_test = df.ml.train_test_split(test_size=0.2, verbose=False)
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
target = 'class_'
params = {
'leaf_estimation_method': 'Gradient',
'learning_rate': 0.1,
'max_depth': 3,
'bootstrap_type': 'Bernoulli',
'subsample': 0.8,
'sampling_frequency': 'PerTree',
'colsample_bylevel': 0.8,
'reg_lambda': 1,
'objective': 'MultiClass',
'eval_metric': 'MultiClass',
'random_state': 42,
'verbose': 0,
}
booster = CatBoostModel(features=features, target=target, num_boost_round=23,
params=params, prediction_type='Class', batch_size=11_000_000)
booster.fit(df=df_train, progress='widget')
df_test = booster.transform(df_train)
df_test
# -
# ### `River` example
#
# `River` is an up-and-coming library for online learning, and provides a variety of models that can learn incrementally. While most of the `river` models currently support per-sample training, few do support mini-batch training which is extremely fast - a great synergy to do machine learning with vaex.
# + tags=["skip-ci"]
from vaex.ml.incubator.river import RiverModel
from river.linear_model import LinearRegression
from river import optim
df = vaex.ml.datasets.load_iris_1e9()
df_train, df_test = df.ml.train_test_split(test_size=0.2, verbose=False)
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width']
target = 'class_'
river_model = RiverModel(features=features,
target=target,
model=LinearRegression(optimizer=optim.SGD(0.001), intercept_lr=0.001),
prediction_name='prediction_raw',
batch_size=500_000)
river_model.fit(df_train, progress='widget')
river_model.transform(df_test)
# -
# ## State transfer - pipelines made easy
#
# Each `vaex` DataFrame consists of two parts: _data_ and _state_. The _data_ is immutable, and any operation such as filtering, adding new columns, or applying transformers or predictive models just modifies the _state_. This is extremely powerful concept and can completely redefine how we imagine machine learning pipelines.
#
# As an example, let us once again create a model based on the Iris dataset. Here, we will create a couple of new features, do a PCA transformation, and finally train a predictive model.
# +
# Load data and split it in train and test sets
df = vaex.ml.datasets.load_iris()
df_train, df_test = df.ml.train_test_split(test_size=0.2, verbose=False)
# Create new features
df_train['petal_ratio'] = df_train.petal_length / df_train.petal_width
df_train['sepal_ratio'] = df_train.sepal_length / df_train.sepal_width
# Do a PCA transformation
features = ['petal_length', 'petal_width', 'sepal_length', 'sepal_width', 'petal_ratio', 'sepal_ratio']
pca = vaex.ml.PCA(features=features, n_components=6)
df_train = pca.fit_transform(df_train)
# Display the training DataFrame at this stage
df_train
# -
# At this point, we are ready to train a predictive model. In this example, let's use `LightGBM` with its `scikit-learn` API.
# +
import lightgbm
features = df_train.get_column_names(regex='^PCA')
booster = lightgbm.LGBMClassifier()
vaex_model = Predictor(model=booster, features=features, target='class_')
vaex_model.fit(df=df_train)
df_train = vaex_model.transform(df_train)
df_train
# -
# The final `df_train` DataFrame contains all the features we created, including the predictions right at the end. Now, we would like to apply the same transformations to the test set. All we need to do, is to simply extract the _state_ from `df_train` and apply it to `df_test`. This will propagate all the changes that were made to the training set on the test set.
#
# +
state = df_train.state_get()
df_test.state_set(state)
df_test
# -
# And just like that `df_test` contains all the columns, transformations and the prediction we modelled on the training set. The state can be easily serialized to disk in a form of a JSON file. This makes deployment of a machine learning model as trivial as simply copying a JSON file from one environment to another.
# +
df_train.state_write('./iris_model.json')
df_test.state_load('./iris_model.json')
df_test
| docs/source/tutorial_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyscal-test
# language: python
# name: pyscal-test
# ---
# # Disorder variable
#
# In this example, [disorder variable](random link) which was introduced to measure the disorder of a system is explored. We start by importing the necessary modules. We will use :mod:`~pyscal.crystal_structures` to create the necessary crystal structures.
import pyscal as pc
import pyscal.crystal_structures as pcs
import matplotlib.pyplot as plt
import numpy as np
# First an fcc structure with a lattice constant of 4.00 is created.
fcc_atoms, fcc_box = pcs.make_crystal('fcc', lattice_constant=4, repetitions=[4,4,4])
# The created atoms and box are assigned to a :class:`~pyscal.core.System` object.
fcc = pc.System()
fcc.box = fcc_box
fcc.atoms = fcc_atoms
# The next step is find the neighbors, and the calculate the Steinhardt parameter based on which we could calculate the disorder variable.
fcc.find_neighbors(method='cutoff', cutoff='adaptive')
# Once the neighbors are found, we can calculate the Steinhardt parameter value. In this example $q=6$ will be used.
fcc.calculate_q(6)
# Finally, disorder parameter can be calculated.
fcc.calculate_disorder()
# The calculated disorder value can be accessed for each atom using the :attr:`~pyscal.catom.disorder` variable.
atoms = fcc.atoms
disorder = [atom.disorder for atom in atoms]
np.mean(disorder)
# As expected, for a perfect fcc structure, we can see that the disorder is zero. The variation of disorder variable on a distorted lattice can be explored now. We will once again use the `noise` keyword along with :func:`~pyscal.crystal_structures.make_crystal` to create a distorted lattice.
fcc_atoms_d1, fcc_box_d1 = pcs.make_crystal('fcc', lattice_constant=4, repetitions=[4,4,4], noise=0.01)
fcc_d1 = pc.System()
fcc_d1.box = fcc_box_d1
fcc_d1.atoms = fcc_atoms_d1
# Once again, find neighbors and then calculate disorder
fcc_d1.find_neighbors(method='cutoff', cutoff='adaptive')
fcc_d1.calculate_q(6)
fcc_d1.calculate_disorder()
# Check the value of disorder
atoms_d1 = fcc_d1.atoms
disorder = [atom.disorder for atom in atoms_d1]
np.mean(disorder)
# The value of average disorder for the system has increased with noise. Finally trying with a high amount of noise.
fcc_atoms_d2, fcc_box_d2 = pcs.make_crystal('fcc', lattice_constant=4, repetitions=[4,4,4], noise=0.1)
fcc_d2 = pc.System()
fcc_d2.box = fcc_box_d2
fcc_d2.atoms = fcc_atoms_d2
fcc_d2.find_neighbors(method='cutoff', cutoff='adaptive')
fcc_d2.calculate_q(6)
fcc_d2.calculate_disorder()
atoms_d2 = fcc_d2.atoms
disorder = [atom.disorder for atom in atoms_d2]
np.mean(disorder)
# The value of disorder parameter shows an increase with the amount of lattice distortion. An averaged version of disorder parameter, averaged over the neighbors for each atom can also be calculated as shown below.
fcc_d2.calculate_disorder(averaged=True)
atoms_d2 = fcc_d2.atoms
disorder = [atom.avg_disorder for atom in atoms_d2]
np.mean(disorder)
# The disorder parameter can also be calculated for values of Steinhardt parameter other than 6. For example,
fcc_d2.find_neighbors(method='cutoff', cutoff='adaptive')
fcc_d2.calculate_q([4, 6])
fcc_d2.calculate_disorder(q=4, averaged=True)
atoms_d2 = fcc_d2.atoms
disorder = [atom.disorder for atom in atoms_d2]
np.mean(disorder)
# $q=4$, for example, can be useful when measuring disorder in bcc crystals
| examples/04_disorder_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Augmentation - `TensorFlow`
# A technique to increase the diversity of training set by applying random (but realistic) transformations such as image rotation.
#
# #### Imports.
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
from tensorflow import keras
# > Loading the dataset. We are going to use the `tf_flowers` dataset.
#
(train_ds, val_ds, test_ds), metadata = tfds.load(
'tf_flowers',
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
class_names = [metadata.features['label'].int2str(i) for i in range(5)]
class_names
# ### The Keras preprocessing layers
# The Keras preprocessing layers API allows us to build Keras-native input processing pipelines.
# * We can preprocess input and using the `Sequantial` API and then pass them as a layer in a `NN`.
# * [Preprocesing Text, images, etc](https://keras.io/guides/preprocessing_layers/) using keras
#
image, label = next(iter(train_ds))
#image, label
plt.imshow(image)
plt.title(class_names[label])
plt.show()
# Now we want to do transformations on this image.
# +
IMG_SIZE = 180
resize_and_rescale = keras.Sequential([
keras.layers.experimental.preprocessing.Resizing(IMG_SIZE, IMG_SIZE),
keras.layers.experimental.preprocessing.Rescaling(1./255)
])
image_output = resize_and_rescale(image)
# -
plt.imshow(image_output)
plt.title(class_names[label])
plt.show()
data_augmentation = tf.keras.Sequential([
keras.layers.experimental.preprocessing.RandomFlip("horizontal_and_vertical"),
keras.layers.experimental.preprocessing.RandomRotation(0.5),
])
plt.figure(figsize=(10, 10))
for i in range(9):
augmented_image = data_augmentation(tf.expand_dims(image, 0))
ax = plt.subplot(3, 3, i + 1)
plt.imshow(augmented_image[0])
plt.axis("off")
# As we can see from a single image we now have different lokking images of the same flower.
# * [TFTutorial- Docs](https://keras.io/guides/preprocessing_layers/)
# * [Keras Docs](https://www.tensorflow.org/tutorials/images/data_augmentation)
# ### ``Tf.image`` Data argumentation.
#
# * [tf.image](https://www.tensorflow.org/api_docs/python/tf/image)
# This is the recommended way of processing images since the `keras.Sequential` is still under experimenta.
# * You have to define a function that performs transformation on an image and return the processed image.
# * We can use the ``map`` function to appy the transformation to all the images in the dataset.
#
# ```python
# def argument(image, label):
# image = tf.image.resize(image, (224, 224))
# image = tf.image.random_brightness(image, 0.1)
# ....
# return image, label
#
# # All images will be stransformsed to defined arguments in the fn
# transformed_images = train_ds.map(argument)
# ```
# * all transforms that we can apply are found [here](https://www.tensorflow.org/api_docs/python/tf/image)
#
# #### Example:
# > Grayscale image.
# +
def augment(image):
return tf.image.rgb_to_grayscale(image)
plt.imshow(augment(image), cmap="gray")
plt.title(class_names[label])
plt.show()
| tf-data-argumentation/.ipynb_checkpoints/01_Data_Augmentation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:multiplexed_segmentation]
# language: python
# name: conda-env-multiplexed_segmentation-py
# ---
# %%
import tifffile
from pathlib import Path as P
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors as colors
from skimage.feature import match_template
import pandas as pd
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 300
# %%
class C:
fol_path_crops = P('/home/vitoz/Data/SegData/basel_zuri/ilastik_random_combined')
fol_path_full = P('/home/vitoz/Data/SegData/basel_zuri/analysis_stacks')
# %%
file_paths_crop = list(C.fol_path_crops.glob('*_ilastik2_crop250.tiff'))
file_paths_full = [C.fol_path_full / fn.name.replace('_crop250.tiff', '.tiff') for fn in file_paths_crop]
# %%
# %%
file_paths_full[0]
# %%
assert all([fn.exists() for fn in file_paths_full])
# %%
fp_crop = file_paths_crop[1]
fp_full = file_paths_full[1]
# %%
# %%
im_crop = tifffile.imread(str(fp_crop),out = 'memmap')[-1]
im_full = tifffile.imread(str(fp_full),out = 'memmap')[-1]
# %%
# %%
# %%
result = match_template(im_full, im_crop)
ij = np.unravel_index(np.argmax(result), result.shape)
x, y = ij[::-1]
# %%
result.max()
# %%
fig = plt.figure(figsize=(8, 3))
ax1 = plt.subplot(1, 4, 1)
ax2 = plt.subplot(1, 4, 2)
ax4 = plt.subplot(1, 4, 3)
ax3 = plt.subplot(1, 4, 4, sharex=ax2, sharey=ax2)
ax1.imshow(im_crop, cmap=plt.cm.gray,norm=colors.PowerNorm(gamma=1/2))
ax1.set_axis_off()
ax1.set_title('template')
ax2.imshow(im_full, cmap=plt.cm.gray, norm=colors.PowerNorm(gamma=1/2))
#ax2.set_axis_off()
ax2.set_title('image')
# highlight matched region
hcoin, wcoin = im_crop.shape
rect = plt.Rectangle((x, y), wcoin, hcoin, edgecolor='r', facecolor='none')
ax2.add_patch(rect)
ax4.imshow(im_full[y:(y+hcoin), x:(x+wcoin)], cmap=plt.cm.gray, norm=colors.PowerNorm(gamma=1/2))
ax4.set_axis_off()
ax4.set_title('image')
ax3.imshow(result)
ax3.set_axis_off()
ax3.set_title('`match_template`\nresult')
# highlight matched region
ax3.autoscale(False)
ax3.plot(x, y, 'o', markeredgecolor='r', markerfacecolor='none', markersize=10)
plt.show()
# %%
def get_crop_coordinates(img_crop, img_full):
result = match_template(img_full, img_crop)
ij = np.unravel_index(np.argmax(result), result.shape)
x, y = ij[::-1]
w, h = img_crop.shape
return x, y, w, h, result.max()
# %%
# %%time
out_list = []
for fn_crop, fn_full in zip(file_paths_crop, file_paths_full):
im_crop = tifffile.imread(str(fn_crop),out = 'memmap')[-1]
im_full = tifffile.imread(str(fn_full),out = 'memmap')[-1]
x, y, w, h, score = get_crop_coordinates(im_crop, im_full)
out_list.append({
'basename': fn_full.name.replace('_ilastik2.tiff',''),
'x': x,
'y': y,
'w': w,
'h': h,
'score': score
})
print(out_list[-1])
# %%
pd.DataFrame(out_list).to_csv('../../resources/manual_coordinates.csv', index=False)
# %%
| subworkflows/prepare_data/workflow/notebooks/99_find_crop_coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyo import *
import wx
print("Audio host APIS:")
pa_list_host_apis()
pa_list_devices()
print("Default input device: %i" % pa_get_default_input())
print("Default output device: %i" % pa_get_default_output())
s = Server(duplex = 0)
s.setOutputDevice(7)
s.boot()
s.start()
# +
s.amp = 0.25
# Simple infinite sequence.
e = Events(freq=EventSeq([250, 300, 350, 400])).play()
# This will play the same note forever.
e3 = Events(freq=200).play()
s.gui(locals())
# -
s.stop()
| exploring_music_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nitishkiitk/Automatic-Music-Transcription/blob/main/mid_term_implementation_22.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="U702ZjCJVHOn" outputId="07af6119-8547-4f44-fe73-d5bad9bc25dd"
import librosa, librosa.display
import matplotlib.pyplot as plt
import numpy as np
from glob import glob
data_dir="drive/MyDrive/Music_transcription/P2"
audio_file=glob(data_dir +'/*.wav')
n=len(audio_file)
for file in range(0, n,1):
signal, sam_rat=librosa.load(audio_file[file], sr=22050)
fft=np.fft.fft(signal)
spectrum=np.abs(fft)
freq=np.linspace(0, sam_rat, len(spectrum))
l_spectrum=spectrum[:int(len(spectrum)/2)]
l_fres=freq[: int(len(spectrum)/2)]
hope_len=512
n_fft=len(signal)
n_fft_duration=float(n_fft)/sam_rat
hope_len_duration=float(hope_len)/sam_rat
stft=librosa.stft(signal, n_fft=n_fft, hop_length=hope_len )
spectrogram=np.abs(stft)
log_spectrogram=librosa.amplitude_to_db(spectrogram)
fig_size=(12,10)
#fig,ax=plt.subplot(figsize=fig_size)
b=librosa.display.specshow(log_spectrogram, sr=sam_rat,hop_length=hope_len, x_axis='time', y_axis='linear')
plt.xlabel("Time")
plt.ylabel("Frequency")
plt.colorbar(b,format="%+2.0f dB")
plt.title("Spectrogram (dB)")
| mid_term_implementation_22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import torch
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import interp1d
V_2_1 = np.loadtxt('data/data_17_09/V1_V2/data_V1.csv', delimiter=',', skiprows=3)
V_2_2 = np.loadtxt('data/data_17_09/V1_V3/data_V1.csv', delimiter=',', skiprows=3)
V_2_3 = np.loadtxt('data/data_17_09/V1_V4/data_V1.csv', delimiter=',', skiprows=3)
V_2_4 = np.loadtxt('data/data_17_09/V1_V5/data_V1.csv', delimiter=',', skiprows=3)
V_2_5 = np.loadtxt('data/data_17_09/V1_V6/data_V1.csv', delimiter=',', skiprows=3)
V_2_6 = np.loadtxt('data/data_17_09/V1_V7/data_V1.csv', delimiter=',', skiprows=3)
V_2_7 = np.loadtxt('data/data_17_09/V1_V8/data_V1.csv', delimiter=',', skiprows=3)
V_2_8 = np.loadtxt('data/data_17_09/V1_V9/data_V1.csv', delimiter=',', skiprows=3)
V_2_9 = np.loadtxt('data/data_17_09/V1_V10/data_V1.csv', delimiter=',', skiprows=3)
data = np.concatenate((V_2_1[:, 1:], V_2_2[:, 1:], V_2_3[:, 1:], V_2_4[:, 1:], V_2_5[:, 1:], V_2_6[:, 1:], V_2_7[:, 1:], V_2_8[:, 1:], V_2_9[:, 1:]), axis=1)
time = V_2_1[:, 0]
time[0:2]
plt.plot(data)
V2_2 = np.loadtxt('data/data_17_09/V1_V2/data_V2.csv', delimiter=',', skiprows=3)
V2_3 = np.loadtxt('data/data_17_09/V1_V3/data_V3.csv', delimiter=',', skiprows=3)
V2_4 = np.loadtxt('data/data_17_09/V1_V4/data_V4.csv', delimiter=',', skiprows=3)
V2_5 = np.loadtxt('data/data_17_09/V1_V5/data_V5.csv', delimiter=',', skiprows=3)
V2_6 = np.loadtxt('data/data_17_09/V1_V6/data_V6.csv', delimiter=',', skiprows=3)
V2_7 = np.loadtxt('data/data_17_09/V1_V7/data_V7.csv', delimiter=',', skiprows=3)
V2_8 = np.loadtxt('data/data_17_09/V1_V8/data_V8.csv', delimiter=',', skiprows=3)
V2_9 = np.loadtxt('data/data_17_09/V1_V9/data_V9.csv', delimiter=',', skiprows=3)
V2_10 = np.loadtxt('data/data_17_09/V1_V10/data_V10.csv', delimiter=',', skiprows=3)
V2_11 = np.loadtxt('data/data_17_09/V1_V11/data_V11.csv', delimiter=',', skiprows=3)
V2_12 = np.loadtxt('data/data_17_09/V1_V12/data_V12.csv', delimiter=',', skiprows=3)
delta_V = np.concatenate((V2_2[:, 1:], V2_3[:, 1:], V2_4[:, 1:], V2_5[:, 1:], V2_6[:, 1:], V2_7[:, 1:], V2_8[:, 1:], V2_9[:, 1:], V2_10[:, 1:], V2_11[:, 1:], V2_12[:, 1:]), axis=1)
#delta_V_1 = np.concatenate((V10_2[:, 1:], V10_3[:, 1:], V10_4[:, 1:], V10_5[:, 1:], V10_6[:, 1:], V10_7[:, 1:], V10_8[:, 1:], V10_9[:, 1:], V10_10[:, 1:]), axis=1)
delta_V.shape
#plt.figure(figsize=(6, 6))
#plt.subplot(121)
plt.contourf(delta_V[:,:])
plt.plot(delta_V[3000,:10])
plt.plot(delta_V[2000,:10])
plt.plot(delta_V[2500,:10])
a=delta_V[2500,:10]
b=np.linspace(0,9,num=10)
c=np.linspace(0,9,num=15)
f = interp1d(b, a,kind='cubic')
#plt.plot(c,f(c))
plt.plot(f(c))
plt.figure(figsize=(9, 6))
plt.plot(V2_2[540:3000, 1])
plt.plot(V2_3[540:3000, 1])
plt.plot(V2_4[540:3000, 1])
plt.plot(V2_5[540:3000, 1])
plt.plot(V2_6[540:3000, 1])
plt.plot(V2_7[540:3000, 1])
plt.plot(V2_8[540:3000, 1])
plt.plot(V2_9[540:3000, 1])
plt.plot(V2_10[540:3000, 1])
plt.plot(V2_11[540:3000, 1])
plt.plot(V2_12[540:3000, 1])
#plt.plot(time, V2_4[:, 1])
plt.plot(V2_8[560:3000, 1])
plt.plot(V2_9[560:3000, 1])
plt.plot(V2_10[560:3000, 1])
plt.plot(V2_11[560:3000, 1])
plt.plot(V2_12[560:3000, 1])
plt.contourf(output_data)
output_data.shape
# +
# DeepMoD stuff
from deepymod import DeepMoD
from deepymod.model.func_approx import NN, Siren
from deepymod.model.library import Library1D
from deepymod.model.constraint import LeastSquares
from deepymod.model.sparse_estimators import Clustering, Threshold
from deepymod.training import train
from deepymod.training.sparsity_scheduler import TrainTestPeriodic
# Setting cuda
if torch.cuda.is_available():
torch.set_default_tensor_type('torch.cuda.FloatTensor')
# Settings for reproducibility
np.random.seed(42)
torch.manual_seed(0)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# -
output_data = delta_V[560:3000,:].T
output_data.shape
plt.contourf(output_data[:,:2000])
x = np.linspace(0, 1, output_data.shape[0])
t = np.linspace(0, 1, output_data.shape[1])
x
x_grid, t_grid = np.meshgrid(x, t, indexing='ij')
np.max(output_data)
X = np.transpose((t_grid.flatten(), x_grid.flatten()))
y = np.real(output_data).reshape((output_data.size, 1))
y.shape
y = y/np.max(y)
# +
number_of_samples = 8000
idx = np.random.permutation(y.shape[0])
X_train = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y[idx, :][:number_of_samples], dtype=torch.float32)
# -
network = NN(2, [50, 50, 50, 50], 1)
# Configuration of the library function: We select athe library with a 2D spatial input. Note that that the max differential order has been pre-determined here out of convinience. So, for poly_order 1 the library contains the following 12 terms:
# * [$1, u_x, u_{xx}, u_{xxx}, u, u u_{x}, u u_{xx}, u u_{xxx}, u^2, u^2 u_{x}, u^2 u_{xx}, u^2 u_{xxx}$]
library = Library1D(poly_order=2, diff_order=3)
# Configuration of the sparsity estimator and sparsity scheduler used. In this case we use the most basic threshold-based Lasso estimator and a scheduler that asseses the validation loss after a given patience. If that value is smaller than 1e-5, the algorithm is converged.
estimator = Threshold(0.1)
sparsity_scheduler = TrainTestPeriodic(periodicity=50, patience=10, delta=1e-5)
# Configuration of the sparsity estimator
constraint = LeastSquares()
# Configuration of the sparsity scheduler
# Now we instantiate the model and select the optimizer
model = DeepMoD(network, library, estimator, constraint)
# Defining optimizer
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True, lr=2e-3)
# ## Run DeepMoD
# We can now run DeepMoD using all the options we have set and the training data:
# * The directory where the tensorboard file is written (log_dir)
# * The ratio of train/test set used (split)
# * The maximum number of iterations performed (max_iterations)
# * The absolute change in L1 norm considered converged (delta)
# * The amount of epochs over which the absolute change in L1 norm is calculated (patience)
train(model, X_train, y_train, optimizer,sparsity_scheduler, log_dir='runs/Akshay_big/', split=0.8, max_iterations=100000, delta=0.1e-6, patience=10)
# Configuring model
network = NN(2, [30, 30, 30, 30, 30], 1) # Function approximator
library = Library1D(poly_order=1, diff_order=2) # Library function
estimator = Threshold(0.01) # Sparse estimator
constraint = LeastSquares() # How to constrain
model = DeepMoD(network, library, estimator, constraint) # Putting it all in the model
# Running model
sparsity_scheduler = Periodic(periodicity=100) # Defining when to apply sparsity
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True) # Defining optimizer
train(model, X_train, y_train, optimizer, sparsity_scheduler,delta=0.002) # Running
train(model, X_train, y_train, optimizer, sparsity_scheduler,delta=0.0001, max_iterations = 100000) # Running
train(model, X_train, y_train, optimizer, sparsity_scheduler,delta=0.0001, max_iterations = 100000) # Running
train(model, X_train, y_train, optimizer, sparsity_scheduler,delta=0.0001, max_iterations = 100000) # Running
train(model, X_train, y_train, optimizer, sparsity_scheduler,delta=0.0001, max_iterations = 100000) # Running
| paper/Cable_equation/old/cable_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import numpy as np
from bqplot import *
# + [markdown] deletable=true editable=true
# ## Bins Mark
#
# This `Mark` is essentially the same as the `Hist` `Mark` from a user point of view, but is actually a `Bars` instance that bins sample data.
#
# The difference with `Hist` is that the binning is done in the backend, so it will work better for large data as it does not have to ship the whole data back and forth to the frontend.
# + deletable=true editable=true
# Create a sample of Gaussian draws
np.random.seed(0)
x_data = np.random.randn(1000)
# + [markdown] deletable=true editable=true
# Give the `Hist` mark the data you want to perform as the `sample` argument, and also give 'x' and 'y' scales.
# + deletable=true editable=true
x_sc = LinearScale()
y_sc = LinearScale()
hist = Bins(sample=x_data, scales={'x': x_sc, 'y': y_sc}, padding=0,)
ax_x = Axis(scale=x_sc, tick_format='0.2f')
ax_y = Axis(scale=y_sc, orientation='vertical')
Figure(marks=[hist], axes=[ax_x, ax_y], padding_y=0)
# + [markdown] deletable=true editable=true
# The midpoints of the resulting bins and their number of elements can be recovered via the read-only traits `x` and `y`:
# + deletable=true editable=true
hist.x, hist.y
# + [markdown] deletable=true editable=true
# ## Tuning the bins
#
# Under the hood, the `Bins` mark is really a `Bars` mark, with some additional magic to control the binning. The data in `sample` is binned into equal-width bins. The parameters controlling the binning are the following traits:
#
# - `bins` sets the number of bins. It is either a fixed integer (10 by default), or the name of a method to determine the number of bins in a smart way ('auto', 'fd', 'doane', 'scott', 'rice', 'sturges' or 'sqrt').
#
# - `min` and `max` set the range of the data (`sample`) to be binned
#
# - `density`, if set to `True`, normalizes the heights of the bars.
#
# For more information, see the documentation of `numpy`'s `histogram`
# + deletable=true editable=true
x_sc = LinearScale()
y_sc = LinearScale()
hist = Bins(sample=x_data, scales={'x': x_sc, 'y': y_sc}, padding=0,)
ax_x = Axis(scale=x_sc, tick_format='0.2f')
ax_y = Axis(scale=y_sc, orientation='vertical')
Figure(marks=[hist], axes=[ax_x, ax_y], padding_y=0)
# + deletable=true editable=true
# Changing the number of bins
hist.bins = 'sqrt'
# + deletable=true editable=true
# Changing the range
hist.min = 0
# + [markdown] deletable=true editable=true
# ## Histogram Styling
#
# The styling of `Hist` is identical to the one of `Bars`
# + deletable=true editable=true
# Normalizing the count
x_sc = LinearScale()
y_sc = LinearScale()
hist = Bins(sample=x_data, scales={'x': x_sc, 'y': y_sc}, density=True)
ax_x = Axis(scale=x_sc, tick_format='0.2f')
ax_y = Axis(scale=y_sc, orientation='vertical')
Figure(marks=[hist], axes=[ax_x, ax_y], padding_y=0)
# + deletable=true editable=true
# changing the color
hist.colors=['orangered']
# + deletable=true editable=true
# stroke and opacity update
hist.stroke = 'orange'
hist.opacities = [0.5] * len(hist.x)
# + deletable=true editable=true
# Laying the histogram on its side
hist.orientation = 'horizontal'
ax_x.orientation = 'vertical'
ax_y.orientation = 'horizontal'
| examples/Marks/Object Model/Bins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (10.0,20.0)
import sys
sys.path.append('../')
import causalMP
cmp = causalMP.CausalMP(data='../../Data/speech_corpora/TIMIT/')
cmp.thresh = 0.1
cmp.test_inference(length=10000)
cmp.learn_step()
| Notebooks/testing cmp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
from skimage.future import graph
rag = graph.rag.rag_boundary(labels,edges,connectivity=2)
labels_merged = graph.cut_threshold(labels, rag, 0.4, in_place=False) #used 0.88 for large set
graph.show_rag(label_sep[0,:,:], rag, bin_mask_clean[o,:,:])
# Things to discuss:
#
# Post-processing:
# `skimage.segmentation.expand_labels(label_image)`
# `skimage.segmentation.clear_border(labels[, ...])`
#
# edges = segmentation.find_boundaries(labels)
#
#
# +
labels_merged = np.empty(seg_cell_sm.shape, dtype=np.int32) #initialize empty array of right size for ou
for t, labels in enumerate(label_sep):
edges = filters.sobel(seg_cell_sm[t,:,:])
edges = (edges - np.min(edges[:]))/(np.max(edges[:]) - np.min(edges[:]))
rag = graph.rag.rag_boundary(labels,edges.astype(np.float32),connectivity=2)
labels_merged[t,:,:] = graph.cut_threshold(labels, rag, 0.6, in_place=False)
merged_layer = viewer.add_labels(labels_merged, name='after merging')
# +
# Uncomment the following line to see the solution
# # %load ./snippets/solution_exercise.py
# +
# Load config ('2D' or 'mothermachine'):
with open(config_file) as f:
data = json.load(f)
for key, item in data.items():
if isinstance(item, str):
if '~' in item:
data[key] = item.replace('~',str(pathlib.Path.home()))
print(data[key])
new_config_file = str((root / 'config_2D_local.json').resolve())
with open(new_config_file, 'w') as f:
json.dump(data, f)
delta.config.load_config(new_config_file)
# -
grouped = df.groupby('unique_id')
grouped.agg(["first","last"])
# +
# To display results movies:
from IPython.display import HTML
from base64 import b64encode
def display_movie(filename):
mp4 = open(filename,'rb').read()
data_url = "data:video/mp4;base64," + b64encode(mp4).decode()
return HTML("""
<video width=600 controls>
<source src="%s" type="video/mp4">
</video>
""" % data_url
)
| TutorsOnly/code_snippets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 06 Extras
# ### Zusätzliche Basisfunktionen
# * % Gibt den Rest einer Division aus
# * // lässt Kommastellen einer Division weg
#
#
# 1. Teile 2/3 und lasse die Stellen nach dem Komma weg
# 2. Teile 5/3 und gebe nur den Rest aus
# ### Datentypen II
# 3. Ergänze den Code in Zeile 4 so dass die Division keine Fehlermeldung ergibt
# +
c = '162'
d = 3
c/d
# -
# 4. Gib b mal die ziffer a aus. Z.B wenn a = 7 und b = 3 ist das gewünschte Ergebnis die Zahl 777.
#
# *(Tipp: was passiert wenn du einen String mit einer Zahl multiplizierst?)*
#
# +
a = 1
b = 4
# dein Code hier
# -
# ### Listen
# 5. Entferne das Element der Liste welches keine Zahl ist:
# +
list_1 = [1, 4, '162', 3]
# dein code hier
# -
# 6. Wandle die Liste in einen String um, mit jeweils einem Leerschlag zwischen den Elementen:
#
# *Tipp: .join anwenden: https://www.w3schools.com/python/ref_string_join.asp*
# +
list_2 = ['A', 'B', 'C', 'D']
# -
# 7. Speichere deinen Namen als String und wandle ihn in eine Liste bei der jeder Buchstabe ein Listenelement ist
| 02 Python Teil 1/06 Extras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_mxnet_p36)
# language: python
# name: conda_mxnet_p36
# ---
# # Hyperparameter Tuning with Amazon SageMaker and MXNet
# _**Creating a Hyperparameter Tuning Job for an MXNet Network**_
#
# ---
#
# ---
#
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Code](#Code)
# 1. [Tune](#Train)
# 1. [Wrap-up](#Wrap-up)
#
# ---
#
# ## Background
#
# This example notebook focuses on how to create a convolutional neural network model to train the [MNIST dataset](http://yann.lecun.com/exdb/mnist/) using MXNet distributed training. It leverages SageMaker's hyperparameter tuning to kick off multiple training jobs with different hyperparameter combinations, to find the set with best model performance. This is an important step in the machine learning process as hyperparameter settings can have a large impact on model accuracy. In this example, we'll use the [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) to create a hyperparameter tuning job for an MXNet estimator.
#
# ---
#
# ## Setup
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the notebook instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/using-identity-based-policies.html) for more details on creating these. Note, if a role not associated with the current notebook instance, or more than one role is required for training and/or hosting, please replace `sagemaker.get_execution_role()` with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
import sagemaker
role = sagemaker.get_execution_role()
# -
# Now we'll import the Python libraries we'll need.
import sagemaker
import boto3
from sagemaker.mxnet import MXNet
from sagemaker.tuner import IntegerParameter, CategoricalParameter, ContinuousParameter, HyperparameterTuner
# ---
#
# ## Data
#
# The MNIST dataset is widely used for handwritten digit classification, and consists of 70,000 labeled 28x28 pixel grayscale images of hand-written digits. The dataset is split into 60,000 training images and 10,000 test images. There are 10 classes (one for each of the 10 digits). See [here](http://yann.lecun.com/exdb/mnist/) for more details on MNIST.
#
# For this example notebook we'll use a version of the dataset that's already been published in the desired format to a shared S3 bucket. Let's specify that location now.
region = boto3.Session().region_name
train_data_location = 's3://sagemaker-sample-data-{}/mxnet/mnist/train'.format(region)
test_data_location = 's3://sagemaker-sample-data-{}/mxnet/mnist/test'.format(region)
# ---
#
# ## Code
#
# To use SageMaker's pre-built MXNet containers, we need to pass in an MXNet script for the container to run. For our example, we'll define several functions, including:
# - `load_data()` and `find_file()` which help bring in our MNIST dataset as NumPy arrays
# - `build_graph()` which defines our neural network structure
# - `train()` which is the main function that is run during each training job and calls the other functions in order to read in the dataset, create a neural network, and train it.
#
# There are also several functions for hosting which we won't define, like `input_fn()`, `output_fn()`, and `predict_fn()`. These will take on their default values as described [here](https://github.com/aws/sagemaker-python-sdk#model-serving), and are not important for the purpose of showcasing SageMaker's hyperparameter tuning.
# !cat mnist.py
# Once we've specified and tested our training script to ensure it works, we can start our tuning job. Testing can be done in either local mode or using SageMaker training. Please see the [MXNet MNIST example notebooks](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/sagemaker-python-sdk/mxnet_mnist/mxnet_mnist.ipynb) for more detail.
# ---
#
# ## Tune
#
# Similar to training a single MXNet job in SageMaker, we define our MXNet estimator passing in the MXNet script, IAM role, (per job) hardware configuration, and any hyperparameters we're not tuning.
estimator = MXNet(entry_point='mnist.py',
role=role,
instance_count=1,
instance_type='ml.m4.xlarge',
sagemaker_session=sagemaker.Session(),
py_version='py3',
framework_version='1.4.1',
base_job_name='DEMO-hpo-mxnet',
hyperparameters={'batch_size': 100})
# Once we've defined our estimator we can specify the hyperparameters we'd like to tune and their possible values. We have three different types of hyperparameters.
# - Categorical parameters need to take one value from a discrete set. We define this by passing the list of possible values to `CategoricalParameter(list)`
# - Continuous parameters can take any real number value between the minimum and maximum value, defined by `ContinuousParameter(min, max)`
# - Integer parameters can take any integer value between the minimum and maximum value, defined by `IntegerParameter(min, max)`
#
# *Note, if possible, it's almost always best to specify a value as the least restrictive type. For example, tuning `thresh` as a continuous value between 0.01 and 0.2 is likely to yield a better result than tuning as a categorical parameter with possible values of 0.01, 0.1, 0.15, or 0.2.*
hyperparameter_ranges = {'optimizer': CategoricalParameter(['sgd', 'Adam']),
'learning_rate': ContinuousParameter(0.01, 0.2),
'num_epoch': IntegerParameter(1, 5)}
# Next we'll specify the objective metric that we'd like to tune and its definition. This includes the regular expression (Regex) needed to extract that metric from the CloudWatch logs of our training job.
objective_metric_name = 'Validation-accuracy'
metric_definitions = [{'Name': 'Validation-accuracy',
'Regex': 'Validation-accuracy=([0-9\\.]+)'}]
# Now, we'll create a `HyperparameterTuner` object, which we pass:
# - The MXNet estimator we created above
# - Our hyperparameter ranges
# - Objective metric name and definition
# - Number of training jobs to run in total and how many training jobs should be run simultaneously. More parallel jobs will finish tuning sooner, but may sacrifice accuracy. We recommend you set the parallel jobs value to less than 10% of the total number of training jobs (we'll set it higher just for this example to keep it short).
# - Whether we should maximize or minimize our objective metric (we haven't specified here since it defaults to 'Maximize', which is what we want for validation accuracy)
tuner = HyperparameterTuner(estimator,
objective_metric_name,
hyperparameter_ranges,
metric_definitions,
max_jobs=9,
max_parallel_jobs=3)
# And finally, we can start our tuning job by calling `.fit()` and passing in the S3 paths to our train and test datasets.
tuner.fit({'train': train_data_location, 'test': test_data_location})
# Let's just run a quick check of the hyperparameter tuning jobs status to make sure it started successfully and is `InProgress`.
boto3.client('sagemaker').describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name)['HyperParameterTuningJobStatus']
# ---
#
# ## Wrap-up
#
# Now that we've started our hyperparameter tuning job, it will run in the background and we can close this notebook. Once finished, we can use the [HPO Analysis notebook](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/hyperparameter_tuning/analyze_results/HPO_Analyze_TuningJob_Results.ipynb) to determine which set of hyperparameters worked best.
#
# For more detail on Amazon SageMaker's Hyperparameter Tuning, please refer to the AWS documentation.
| hyperparameter_tuning/mxnet_mnist/hpo_mxnet_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="0f-ewnQs55UL"
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual, FloatSlider, IntSlider
import ipywidgets as widgets
from sklearn import datasets
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
from sklearn.decomposition import KernelPCA
from mpl_toolkits.mplot3d import Axes3D
from sklearn.svm import SVC
import numpy as np
import seaborn as sns
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
import warnings
from sklearn.datasets import make_circles
warnings.filterwarnings('ignore')
# + [markdown] id="Ex0Ntnw3tNDl"
# ## Question 2) Kernel PCA
#
# In previous assignments, you've worked with PCA to find a lower dimensional representation of a data matrix, allowing us to perform tasks like classification more easily.
# As we said in the notes, kernels have a wide range of applications. In this problem, we'll take a look at the application of kernels to PCA.
#
# + [markdown] id="hpLooZAsI6yF"
# ### Question 2a)
# First, let's look at the half moon data again.
# + colab={"base_uri": "https://localhost:8080/", "height": 336} id="INaVOdlxI6SG" outputId="5bce54e3-9bf0-4b0b-93e9-ec28c92eae60"
X, y = datasets.make_moons(n_samples = 500, noise = 0.04)
plt.scatter(X[:, 0], X[:, 1], c = y)
# + [markdown] id="DmIcCxvTJ8ks"
# Run PCA on this dataset with 1 and 2 components, and visualize the result. Fill in the code such that `X_red` has the original data projected onto the first 2 principal components. Answer the following questions.
# * **Do you notice anything different about this graph? Why did this change happen?**
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="a1oEQkPDJFse" outputId="6136fd48-53a2-4ba1-f372-87fd44bec675"
fig, axes = plt.subplots(1, 2, figsize=(18,4))
# START TODO
# END TODO
axes[0].scatter(X_red[:,0], np.zeros(X_red.shape[0]), c = y)
axes[1].scatter(X_red[:,0], X_red[:,1], c = y)
# -
# Ridge regression, while it is able to identify the important directions in our data, is confined to a linear feature space. This means that we are still stuck with the problem where our dataset is linearly inseparable. As we know, it often helps to lift our features by mapping each datapoint to a higher dimensional space. Kernels will allow us to lift our features without ever having to explicitly compute the higher dimensional space for our data matrix. Instead, we can simply just perform PCA on the Gram matrix K, which will give us the most important directions in this lifted feature space without having to go through the computational complexity of computing it.
# sk-learn has a built in Kernel PCA implementation that we can use on our half moon dataset here https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.KernelPCA.html
# Complete the following code to finish the function `kernel_pca_poly`, which takes in a value for gamma `g` and computes a 2 component KernelPCA with gamma `g` and the RBF kernel on the data in `X`.
# + id="1eBSd8azLJh7"
def kernel_pca_gamma(gamma):
# START TODO
# END TODO
fig, axes = plt.subplots(1, 2, figsize=(18,4))
axes[0].scatter(X_red_kernel[:,0], np.zeros(X_red_kernel.shape[0]), c = y)
axes[1].scatter(X_red_kernel[:,0], X_red_kernel[:,1], c = y)
plt.show()
# -
# Now run the following code to visualize our results. Play with the gamma parameter and answer the following questions.
# * **What does the graph look like as gamma approaches infinity? What about negative infinity?**
# * **What happens when gamma is 0? Why does this make sense?**
# * **What is the value of gamma that visually seems like it would cause the data to be most separable?**
# * **What method can we use to find an optimal gamma to make this data separable?**
# + id="6rsdMu8yLaGA"
g_widget = FloatSlider(min=-10, max=20.0)
interact(kernel_pca_gamma,gamma=g_widget)
# -
# ### Question 2b)
# Fill in the code to use a polynomial kernel now, and answer the following questions.
# * **Try keeping the degree fixed and changing gamma. What do you notice happens as gamma gets to be a large positive number? What about to be a small negative number? What about 0?**
# * **Now keep the gamma fixed and change the degree. What do you notice happens as the degree takes on even and odd values? What about small? What about 0?**
def kernel_pca_poly(degree, gamma):
# START TODO
# END TODO
fig, axes = plt.subplots(1, 2, figsize=(18,4))
axes[0].scatter(X_red_kernel[:,0], np.zeros(X_red_kernel.shape[0]), c = y)
axes[1].scatter(X_red_kernel[:,0], X_red_kernel[:,1], c = y)
plt.show()
g_widget = FloatSlider(min=-10, max=20.0)
d_widget = IntSlider(min=1, max=10)
interact(kernel_pca_poly,gamma=g_widget,degree=d_widget)
# ### Question 2c)
# Now let's do some classification with Logistic Regression to see how well we can classify the the original dataset, the dataset projected onto the first two principal components, and the dataset projected using the principal components from kernel PCA.
#
# First, let's look at the original dataset. Fill in the code in TODO such that we fit a Logistic Regression model and store the weights in a variable called `w`. Calculate the accuracy of the classifier on the dataset and store that in orig_accuracy.
# +
# START TODO
# END TODO
plt.scatter(X[:, 0], X[:, 1], c = y)
ax = plt.gca()
ax.autoscale(False)
x_vals = np.array(ax.get_xlim())
y_vals = -(x_vals * w[0])/w[1]
plt.plot(x_vals, y_vals, '--', c="red")
print ("Classifier accuracy: ", orig_accuracy)
# -
# Now do the same for the `X_red` dataset. Answer the following questions.
# * **Is the accuracy different?**
# +
# START TODO
# END TODO
plt.scatter(X_red[:,0], X_red[:,1], c = y)
ax = plt.gca()
ax.autoscale(False)
x_vals = np.array(ax.get_xlim())
y_vals = -(x_vals * w[0])/w[1]
plt.plot(x_vals, y_vals, '--', c="red")
print ("Classifier accuracy: ", pca_accuracy)
# -
# Now let's use kernel PCA. Use PCA with an RBF kernel to transform the dataset and use the value for gamma you visually identified in Question 2b.
# * **How's the accuracy this time?**
# +
# START TODO
# END TODO
plt.scatter(X_red_kernel[:,0], X_red_kernel[:,1], c = y)
ax = plt.gca()
ax.autoscale(False)
x_vals = np.array(ax.get_xlim())
y_vals = -(x_vals * w[0])/w[1]
plt.plot(x_vals, y_vals, '--', c="red")
print ("Classifier accuracy: ", kpca_accuracy)
| final_kernels_project_blank/.ipynb_checkpoints/prob2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MNIST training with PyTorch
#
# MNIST is a widely used dataset for handwritten digit classification. It consists of 70,000 labeled 28x28 pixel grayscale images of hand-written digits. The dataset is split into 60,000 training images and 10,000 test images. There are 10 classes (one for each of the 10 digits). This tutorial will show how to train and test an MNIST model on SageMaker using PyTorch.
#
#
# +
import os
import json
import sagemaker
from sagemaker.pytorch import PyTorch
from sagemaker import get_execution_role
sess = sagemaker.Session()
role = get_execution_role()
output_path='s3://' + sess.default_bucket() + '/mnist'
# -
# ## PyTorch Estimator
#
# The `PyTorch` class allows you to run your training script on SageMaker
# infrastracture in a containerized environment. In this notebook, we
# refer to this container as *training container*.
#
# You need to configure
# it with the following parameters to set up the environment:
#
# - entry_point: A user defined python file to be used by the training container as the
# instructions for training. We further discuss this file in the next subsection.
#
# - role: An IAM role to make AWS service requests
#
# - instance_type: The type of SageMaker instance to run your training script.
# Set it to `local` if you want to run the training job on
# the SageMaker instance you are using to run this notebook
#
# - instance count: The number of instances you need to run your training job.
# Multiple instances are needed for distributed training.
#
# - output_path:
# S3 bucket URI to save training output (model artifacts and output files)
#
# - framework_version: The version of PyTorch you need to use.
#
# - py_version: The python version you need to use
#
# For more information, see [the API reference](https://sagemaker.readthedocs.io/en/stable/api/training/estimators.html#sagemaker.estimator.EstimatorBase)
#
#
# ## Implement the entry point for training
#
# The entry point for training is a python script that provides all
# the code for training a PyTorch model. It is used by the SageMaker
# PyTorch Estimator (`PyTorch` class above) as the entry point for running the training job.
#
# Under the hood, SageMaker PyTorch Estimator creates a docker image
# with runtime environemnts
# specified by the parameters you used to initiated the
# estimator class and it injects the training script into the
# docker image to be used as the entry point to run the container.
#
# In the rest of the notebook, we use *training image* to refer to the
# docker image specified by the PyTorch Estimator and *training container*
# to refer to the container that runs the training image.
#
# This means your training script is very similar to a training script
# you might run outside Amazon SageMaker, but it can access the useful environment
# variables provided by the training image. Checkout [the short list of environment variables provided by the SageMaker service](https://sagemaker.readthedocs.io/en/stable/frameworks/mxnet/using_mxnet.html?highlight=entry%20point) to see some common environment
# variables you might used. Checkout [the complete list of environment variables](https://github.com/aws/sagemaker-training-toolkit/blob/master/ENVIRONMENT_VARIABLES.md) for a complete
# description of all environment variables your training script
# can access to.
#
# In this example, we use the training script `code/train.py`
# as the entry point for our PyTorch Estimator.
#
# !pygmentize 'code/train.py'
# ### Set hyperparameters
#
# In addition, PyTorch estimator allows you to parse command line arguments
# to your training script via `hyperparameters`.
#
# <span style="color:red"> Note: local mode is not supported in SageMaker Studio </span>
# +
# set local_mode to be True if you want to run the training script
# on the machine that runs this notebook
local_mode=True
if local_mode:
instance_type='local'
else:
instance_type='ml.c4.xlarge'
est = PyTorch(
entry_point='train.py',
source_dir='code', # directory of your training script
role=role,
framework_version='1.5.0',
py_version='py3',
instance_type=instance_type,
instance_count=1,
output_path=output_path,
hyperparameters={
'batch-size':128,
'epochs':20,
'learning-rate': 1e-3,
'log-interval':100
}
)
# -
# The training container executes your training script like
#
# ```
# python train.py --batch-size 100 --epochs 10 --learning-rate 1e-3 \
# --log-interval 100
# ```
# ## Set up channels for training and testing data
#
# You need to tell `PyTorch` estimator where to find your training and
# testing data. It can be a link to an S3 bucket or it can be a path
# in your local file system if you use local mode. In this example,
# we download the MNIST data from a public S3 bucket and upload it
# to your default bucket.
# +
import logging
import boto3
from botocore.exceptions import ClientError
# Download training and testing data from a public S3 bucket
def download_from_s3(data_dir='/tmp/data', train=True):
"""Download MNIST dataset and convert it to numpy array
Args:
data_dir (str): directory to save the data
train (bool): download training set
Returns:
None
"""
# Get global config
with open('code/config.json', 'r') as f:
CONFIG=json.load(f)
if not os.path.exists(data_dir):
os.makedirs(data_dir)
if train:
images_file = "train-images-idx3-ubyte.gz"
labels_file = "train-labels-idx1-ubyte.gz"
else:
images_file = "t10k-images-idx3-ubyte.gz"
labels_file = "t10k-labels-idx1-ubyte.gz"
# download objects
s3 = boto3.client('s3')
bucket = CONFIG['public_bucket']
for obj in [images_file, labels_file]:
key = os.path.join("datasets/image/MNIST", obj)
dest = os.path.join(data_dir, obj)
if not os.path.exists(dest):
s3.download_file(bucket, key, dest)
return
download_from_s3('/tmp/data', True)
download_from_s3('/tmp/data', False)
# +
# upload to the default bucket
prefix = 'mnist'
bucket = sess.default_bucket()
loc = sess.upload_data(path='/tmp/data', bucket=bucket, key_prefix=prefix)
channels = {
"training": loc,
"testing": loc
}
# -
# The keys of the dictionary `channels` are parsed to the training image
# and it creates the environment variable `SM_CHANNEL_<key name>`.
#
# In this example, `SM_CHANNEL_TRAINING` and `SM_CHANNEL_TESTING` are created in the training image (checkout
# how `code/train.py` access these variables). For more information,
# see: [SM_CHANNEL_{channel_name}](https://github.com/aws/sagemaker-training-toolkit/blob/master/ENVIRONMENT_VARIABLES.md#sm_channel_channel_name)
#
# If you want, you can create a channel for validation:
# ```
# channels = {
# 'training': train_data_loc,
# 'validation': val_data_loc,
# 'test': test_data_loc
# }
# ```
# You can then access this channel within your training script via
# `SM_CHANNEL_VALIDATION`
#
# ## Run the training script on SageMaker
# Now, the training container has everything to execute your training
# script. You can start the container by calling `fit` method.
est.fit(inputs=channels)
# ## Inspect and store model data
#
# Now, the training is finished, the model artifact has been saved in
# the `output_path`. We
pt_mnist_model_data = est.model_data
print("Model artifact saved at:\n", pt_mnist_model_data)
# We store the variable `model_data` in the current notebook kernel.
# In the [next notebook](get_started_with_mnist_deploy.ipynb), you will learn how to retrieve the model artifact and deploy to a SageMaker
# endpoint.
# %store pt_mnist_model_data
# ## Test and debug the entry point before executing the training container
#
# The entry point `code/train.py` provided here has been tested and it can be executed in the training container.
# When you do develop your own training script, it is a good practice to simulate the container environment
# in the local shell and test it before sending it to SageMaker, because debugging in a containerized environment
# is rather cumbersome. The following script shows how you can test your training script:
# !pygmentize code/test_train.py
| frameworks/pytorch/get_started_mnist_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Polygon drill <img align="right" src="../Supplementary_data/dea_logo.jpg">
#
# * [**Sign up to the DEA Sandbox**](https://docs.dea.ga.gov.au/setup/sandbox.html) to run this notebook interactively from a browser
# * **Compatibility:** Notebook currently compatible with both the `NCI` and `DEA Sandbox` environments
# * **Products used:**
# [ga_ls8c_ard_3](https://explorer.sandbox.dea.ga.gov.au/ga_ls8c_ard_3)
# * **Special requirements:**
# A shape file containing the polygon you would like to use for the analysis. Here we use ACT suburb boundaries, available as a [shapefile from data.gov.au](https://www.data.gov.au/dataset/ds-dga-0257a9da-b558-4d86-a987-535c775cf8d8/details?q=).
# ## Description
# A polygon drill can be used to grab a stack of imagery that corresponds to the location of an input polygon.
# It is a useful tool for generating animations, or running analyses over a range of imagery.
#
# This notebook shows you how to:
#
# 1. Use a polygon's geometry to generate a `dc.load` query
# 2. Mask the returned data with the polygon geometry (to remove unwanted pixels)
# 3. Plot a time step from the imagery stack to check it has been loaded as expected
#
# ***
# ## Getting started
#
# To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
# ### Load packages
# Import Python packages that are used for the analysis.
# +
# %matplotlib inline
import sys
import datacube
import geopandas as gpd
from datacube.utils import geometry
sys.path.append('../Scripts')
from dea_plotting import rgb
from dea_spatialtools import xr_rasterize
# -
# ### Connect to the datacube
#
# Connect to the datacube so we can access DEA data.
# The `app` parameter is a unique name for the analysis which is based on the notebook file name.
dc = datacube.Datacube(app='Polygon_drill')
# ### Analysis parameters
#
# An *optional* section to inform the user of any parameters they'll need to configure to run the notebook:
#
# * `polygon_to_drill`: The path containing the polygon to use for the polygon drill.
# If it's a local polygon, then this parameter is the local path to that polygon.
# If it's located online, then this is the path to the online location of the polygon.
# * `time_to_drill`: e.g. `('2016-01-01', '2016-06-30')`.
# The time over which we want to run the polygon drill, entered as a tuple.
#
polygon_to_drill = 'https://data.gov.au/data/dataset/0257a9da-b558-4d86-a987-535c775cf8d8/resource/d9100544-182d-470c-b3b2-75812322c495/download/act_locality_polygon_shp.zip'
time_to_drill = ('2016-02', '2016-03')
# ## Load up the shapefile we want to use for the polygon drill
# Use code comments for low-level documentation of code
polygon_to_drill = gpd.read_file(polygon_to_drill)
# Check that the polygon loaded as expected. We'll just print the first 3 rows to check
polygon_to_drill.head(3)
# ## Query the datacube using the polygon we have loaded
#
# ### Set up the `dc.load` query
#
# We need to grab the geometry from the polygon we want to use for the polygon drill.
# For this example, we'll just grab the first polygon from the file using `.iloc[0]`:
geom = geometry.Geometry(geom=polygon_to_drill.iloc[0].geometry,
crs=polygon_to_drill.crs)
# To set up the query, we need to specify a few parameters:
#
# - `'geopolygon'`: Here we input the geometry we want to use for the drill that we prepared in the cell above
# - `'time'`: Feed in the `time_to_drill` parameter we set earlier
# - `'output_crs'`: We need to specify the coordinate reference scheme of the output.
# We'll use Albers Equal Area projection for Australia
# - `'resolution'`: You can choose the resolution of the output dataset.
# Since Landsat 8 is 30 m resolution, we'll just use that
# - `'measurements'`: Here is where you specify which bands you want to extract.
# We will just be plotting a true colour image, so we just need red, green and blue.
query = {'geopolygon': geom,
'time': time_to_drill,
'output_crs': 'EPSG:3577',
'resolution': (-30, 30),
'measurements': ['nbart_red',
'nbart_green',
'nbart_blue']
}
# ### Use the query to extract data
#
# Here we have hard coded extraction from Landsat 8 by supplying `product='ga_ls8c_ard_3'`, but this can be changed depending on your requirements.
#
# We can verify that the polygon drill has loaded a time series of satellite data by checking the `Dimensions` of the resulting `xarray.Dataset`.
# In this example, we can see that the polygon drill has loaded 7 time steps (i.e. `Dimensions: time: 7`):
# +
# Load data for our polygon and time period
data = dc.load(product='ga_ls8c_ard_3', group_by='solar_day', **query)
# Check we have some data back with multiple timesteps
data
# -
# ## Plot time series of data
# To inspect the satellite data we have loaded using the pixel drill, we can plot an image for each timestep in the data:
rgb(data, col='time', percentile_stretch=(0.05, 0.95))
# ## Mask data using the original polygon
# The data returned from our polygon drill contains data for the bounding box of the extents of the input polygon, not the actual shape of the polygon.
#
# To get rid of the bits of the image located outside the polygon, we need to mask the data using the original polygon.
# Generate a polygon mask to keep only data within the polygon
# We used the first polygon in 'polygon_to_drill', so we need to mask
# with the same one
mask = xr_rasterize(polygon_to_drill.iloc[[0]], data)
# Now apply the mask
data_masked = data.where(mask)
# ## Plot time series of masked data
# When we plot the masked dataset, we can see that the areas located outside of the polygon have now been masked out (i.e. set to `NaN` or white):
rgb(data_masked, col='time', percentile_stretch=(0.05, 0.95))
# ***
#
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Australia data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/GeoscienceAustralia/dea-notebooks).
#
# **Last modified:** October 2020
#
# **Compatible datacube version:**
print(datacube.__version__)
# ## Tags
# Browse all available tags on the DEA User Guide's [Tags Index](https://docs.dea.ga.gov.au/genindex.html)
# + raw_mimetype="text/restructuredtext" active=""
# **Tags**: :index:`NCI compatible`, :index:`sandbox compatible`, :index:`landsat 8`, :index:`dea_plotting`, :index:`rgb`, :index:`polygon drill`, :index:`shapefile`, :index:`GeoPandas`, :index:`datacube.utils.geometry`, :index:`query`, :index:`Scripts`, :index:`xr_rasterize`, :index:`masking`
| Frequently_used_code/Polygon_drill.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## MFEM Example 1
#
# Adapted from [PyMFEM/ex1.py]( https://github.com/mfem/PyMFEM/blob/master/examples/ex1.py).
# Compare with the [original Example 1](https://github.com/mfem/mfem/blob/master/examples/ex1.cpp) in MFEM.
#
# This example code demonstrates the use of MFEM to define a simple finite element discretization of the Laplace problem
#
# \begin{equation*}
# -\Delta x = 1
# \label{laplace}\tag{1}
# \end{equation*}
#
# in a domain $\Omega$ with homogeneous Dirichlet boundary conditions
#
# \begin{equation*}
# x = 0
# \label{laplace2}\tag{2}
# \end{equation*}
#
# on the boundary $\partial \Omega$.
#
# The problme is discretized on a computational mesh in either 2D or 3D using a finite elements space of the specified order (2 by default) resulting in the global sparse linear system
#
# \begin{equation*}
# A X = B.
# \label{laplace3}\tag{3}
# \end{equation*}
#
# The example highlights the use of mesh refinement, finite element grid functions, as well as linear and bilinear forms corresponding to the left-hand side and right-hand side of the
# discrete linear system. We also cover the explicit elimination of essential boundary conditions and using the GLVis tool for visualization.
# Requires PyMFEM, see https://github.com/mfem/PyMFEM
import mfem.ser as mfem
from glvis import glvis, to_stream
# +
# Load the mesh from a local file
# meshfile = '../../mfem/data/star.mesh'
# mesh = mfem.Mesh(meshfile)
# Alternatively, create a simple square mesh and refine it
mesh = mfem.Mesh(5, 5, "TRIANGLE")
mesh.UniformRefinement()
# Create H1 finite element function space
fec = mfem.H1_FECollection(2, mesh.Dimension()) # order=2
fespace = mfem.FiniteElementSpace(mesh, fec)
# Determine essential degrees of freedom (the whole boundary here)
ess_tdof_list = mfem.intArray()
ess_bdr = mfem.intArray([1]*mesh.bdr_attributes.Size())
fespace.GetEssentialTrueDofs(ess_bdr, ess_tdof_list)
# Define Bilinear and Linear forms for the Laplace problem -Δu=1
one = mfem.ConstantCoefficient(1.0)
a = mfem.BilinearForm(fespace)
a.AddDomainIntegrator(mfem.DiffusionIntegrator(one))
a.Assemble()
b = mfem.LinearForm(fespace)
b.AddDomainIntegrator(mfem.DomainLFIntegrator(one))
b.Assemble()
# Create a grid function for the solution and initialize with 0
x = mfem.GridFunction(fespace);
x.Assign(0.0)
# Form the linear system, AX=B, for the FEM discretization
A = mfem.OperatorPtr()
B = mfem.Vector()
X = mfem.Vector()
a.FormLinearSystem(ess_tdof_list, x, b, A, X, B);
print("Size of the linear system: " + str(A.Height()))
# Solve the system using PCG solver and get the solution in x
Asm = mfem.OperatorHandle2SparseMatrix(A)
Msm = mfem.GSSmoother(Asm)
mfem.PCG(Asm, Msm, B, X, 1, 200, 1e-12, 0.0)
a.RecoverFEMSolution(X, b, x)
# -
# ### Plot the Solution with GLVis
# Plot the mesh + solution (all GLVis keys and mouse commands work)
glvis((mesh, x), 400, 400)
# Plot the mesh only
glvis(mesh)
# Visualization with additional GLVis keys
g = glvis(to_stream(mesh,x) + 'keys ARjlmcbp*******')
g.set_size(600, 400)
g
| examples/ex1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Vaishali-Govind/AdminLTE/blob/master/Sentiment_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="VFsf0r6wdhwE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="dd59403e-935f-4ba9-ffd1-656ce1c2eb84"
# !unzip zomato.zip
# + id="UIWtx-gqeNSw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="a5be930c-8687-4e21-e09e-3cea51accd0d"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Importing NLTK library for using stop words method
import nltk
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import CountVectorizer
# %matplotlib inline
# + id="RpZTdad3eohw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="641acc16-357c-4480-c0b5-354ea6b036c5"
sdf = pd.read_csv('zomato/new_processed_reviews.csv')
sdf.head()
# + id="LMcZoE_egBx_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16f16eb8-8a26-4d48-b3a7-98b5220dee56"
print(sdf.shape)
# + id="SPH9BMdXgP3F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="1885cc9e-ac46-4e0d-c243-ee4c728c8b7f"
print(sdf.info())
# + id="m-4WedvPgf2l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 252} outputId="616680b5-738d-40ff-d5c6-233b1b45ee83"
print(sdf.describe)
# + id="nOdvRXDDgvMH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="a5c03501-e6dd-43aa-959a-5856ab34b748"
#creating a new column for length of the text
sdf['text_len'] = sdf['content'].apply(len)
sdf.head()
# + [markdown] id="1kSnYFUCg-_O" colab_type="text"
# # Visualising the data
# + id="GyquKmEThMKj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154} outputId="37a92d9b-7be8-4ebe-f440-d4e15ecdd180"
graph1 = sns.FacetGrid(data=sdf, col='rating')
graph1.map(plt.hist, 'text_len', color = 'green', bins = 50)
# + id="RRkz4Y9chtov" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 324} outputId="ac147d37-c537-48a8-c2ee-fec220311a55"
#Grouping the data using start rating and finding if any correlation
rating = sdf.groupby('rating').mean()
rating.corr()
sns.heatmap(data = rating.corr(), annot = True)
#### This shows that funny is strongly correlated to useful and useful is strongly correlated to text_len
#### Thus, we can say that longer reviews are more funny and useful
# + id="RixqVakUiDk0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="e606f87e-6b87-4f86-a044-b1683b1a9a9e"
#Preparing for classification
sdf_class = sdf[(sdf['rating'] == 1) | (sdf['rating']==5)]
sdf_class.shape
#putting them in seperate variable
x = sdf_class['content']
y = sdf_class['rating']
print(x.head())
print(y.head())
# + id="UbpCi96UiWyr" colab_type="code" colab={}
#Data cleaning by removing stop words and puntuation
import string
def text_process(content):
nopunc = [char for char in content if char not in string.punctuation]
nopunc = ''.join(nopunc)
return [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
# + [markdown] id="RAEWYrz4ilnV" colab_type="text"
# # Performing Vectorization
# + id="Cxe5FFFljo5_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="6fe5d79d-9128-41cc-ccba-36503aa7d468"
nltk.download("stopwords")
# + id="6lJtO2PvirDI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b1b6269a-7180-444a-85a6-38d4347b544d"
#Import countVectorizer and define it with a variable. Along with that we will fit it to our review text stored in x
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(analyzer = text_process).fit(x)
print(len(vectorizer.vocabulary_))
# + id="viAbbp86jmxW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 961} outputId="67cbef19-d52f-476f-eadd-b6db7753d8d3"
review_0 = x[0]
print(review_0)
vocab_0 = vectorizer.transform([review_0])
print(vocab_0)
print("Following Words back")
print(vectorizer.get_feature_names()[1144])
print(vectorizer.get_feature_names()[2027])
# + id="sQSwhnD0moq1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="f1b3b031-305a-4ab7-8ba5-37a2a028c500"
#Now applying vectorization to the ful review set which would check the shape of new x
x = vectorizer.transform(x)
print('Shape of Sparse Matrix: ',x.shape)
print('Amount of Non-zero occurances:',x.nnz)
# + id="B6SQ_eggpc0a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5c1f7cee-53a6-4ec7-c503-7890b1561517"
# Percentage of non-zero values
density = (100.0 * x.nnz / (x.shape[0] * x.shape[1]))
print("Density = ",density)
# + [markdown] id="XQHLN_RKptbD" colab_type="text"
# # Splitting the data into train and test
# + id="Uib-jjB1pxJd" colab_type="code" colab={}
#Splitting the dataset into training data and test data in the proportion of 80:20
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.2,random_state=101)
# + [markdown] id="q5tmTbNkq2SD" colab_type="text"
# # Applying the classification method
# + [markdown] id="M2A0LPvHq8eY" colab_type="text"
# # Multinomial Naive Bayes
# + id="PLaIKsC8rB3A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0666233e-0d49-479a-f694-99c02f22dd6e"
#Building the model
from sklearn.naive_bayes import MultinomialNB
nb = MultinomialNB()
nb.fit(x_train, y_train)
# + id="ctbQC2kyq574" colab_type="code" colab={}
#Testing our model
nb_predict = nb.predict(x_test)
# + id="uN8inpdkrOLY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 252} outputId="b5b34f12-68cc-47d4-d68e-bca1b2852200"
#Creating the confusion matrix
from sklearn.metrics import confusion_matrix, classification_report
print(confusion_matrix(y_test, nb_predict))
print('\n')
#Creating the classification report
print(classification_report(y_test, nb_predict)) ### The model achieved 94% accuracy.
### However, since we know that there are some bias values,
### so let's just test it on a single review.
#positive single review
pos_review = sdf_class['content'][59]
pos_review
pos_review_t = vectorizer.transform([pos_review])
nb.predict(pos_review_t)[0] ### 5 star rating which is good as expected
#Negative single review
neg_review = sdf_class['content'][281]
neg_review
neg_review_t = vectorizer.transform([neg_review])
nb.predict(neg_review_t)[0] ### 1 star rating which is fine as exp
# + [markdown] id="k5ZCFccVr9I0" colab_type="text"
# # K-NN classifier
# + id="jXtd86PPsAXI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="b655a9ca-74be-4550-a4f9-cb84d9971995"
#Building the model
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn.fit(x_train, y_train)
#Testing our model on x_test
knn_predict = knn.predict(x_test)
#Creating the confusion matrix
from sklearn.metrics import confusion_matrix, classification_report
print(confusion_matrix(y_test, knn_predict))
print('\n')
#Creating the classification report
print(classification_report(y_test, knn_predict)) ### The model achieved 83% accuracy
# + [markdown] id="AiyQgMRTsZmB" colab_type="text"
# # Support Vector Machine
# + id="pfv3wWvOsd-p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="5d08dde8-b73f-428f-f890-1a62e9fad61a"
from sklearn.svm import SVC
svm = SVC()
svm.fit(x_train, y_train)
#Testing our model on x_test
svm_predict = svm.predict(x_test)
#Creating the confusion matrix
from sklearn.metrics import confusion_matrix, classification_report
print(confusion_matrix(y_test, svm_predict))
print('\n')
#Creating the classification report
print(classification_report(y_test, svm_predict)) ### The model achieved 90% accuracy
# + [markdown] id="F4ArIPUcsqzq" colab_type="text"
# # Random Forest Classifier
# + id="5Ber_ZqFssZe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="f15a368b-4596-43fc-af68-397b0a414314"
#Building the model
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf.fit(x_train, y_train)
#Testing our model on x_test
rf_predict = rf.predict(x_test)
#Creating the confusion matrix
from sklearn.metrics import confusion_matrix, classification_report
print(confusion_matrix(y_test, rf_predict))
print('\n')
#Creating the classification report
print(classification_report(y_test, rf_predict)) ### The model achieved 93% accuracy
| Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import libraries
import pandas as pd
import numpy as np
import scipy.stats as stats
import os
import random
import statsmodels.api as sm
import statsmodels.stats.multicomp
from statsmodels.formula.api import ols
from statsmodels.stats.anova import anova_lm
import matplotlib.pyplot as plt
from scipy import stats
import seaborn as sns
# -
#Load data
StatewiseTestingDetails=pd.read_csv('./StatewiseTestingDetails.csv')
population_india_census2011=pd.read_csv('./population_india_census2011.csv')
population_india_census2011.head()
StatewiseTestingDetails.head()
StatewiseTestingDetails['Positive'].sort_values().head()
#List down the states which have 0 corona cases
StatewiseTestingDetails['State'][StatewiseTestingDetails['Positive']==0].unique()
#List down the states which have 1 corona cases
StatewiseTestingDetails['State'][StatewiseTestingDetails['Positive']==1].unique()
##We see that there're many entries with 0. That means no case has been detected. So we can add 1 in all entries.
#So while perfroming any sort of Data transformation that involves log in it , won't give error.
StatewiseTestingDetails['Positive']=StatewiseTestingDetails['Positive']+1
StatewiseTestingDetails['Positive'].sort_values()
#Imput missing values by median of each state
stateMedianData=StatewiseTestingDetails.groupby('State')[['Positive']].median().\
reset_index().rename(columns={'Positive':'Median'})
stateMedianData.head()
StatewiseTestingDetails.head()
for index,row in StatewiseTestingDetails.iterrows():
if pd.isnull(row['Positive']):
StatewiseTestingDetails['Positive'][index]=int(stateMedianData['Median'][stateMedianData['State']==row['State']])
StatewiseTestingDetails['Positive'].sort_values()
#Merge StatewiseTestingDetails & population_india_census2011 dataframes
data=pd.merge(StatewiseTestingDetails,population_india_census2011,on='State')
##Sort the Data Frame
data['Positive'].sort_values()
# +
#Write a function to create densityGroup bucket
def densityCheck(data):
data['density_Group']=0
for index,row in data.iterrows():
status=None
i=row['Density'].split('/')[0]
try:
if (',' in i):
i=int(i.split(',')[0]+i.split(',')[1])
elif ('.' in i):
i=round(float(i))
else:
i=int(i)
except ValueError as err:
pass
try:
if (0<i<=300):
status='Dense1'
elif (300<i<=600):
status='Dense2'
elif (600<i<=900):
status='Dense3'
else:
status='Dense4'
except ValueError as err:
pass
data['density_Group'].iloc[index]=status
return data
# -
data.columns
data['Positive'].sort_values()
#Map each state as per its density group
data=densityCheck(data)
#We'll export this data so we can use it for Two - way ANOVA test.
stateDensity=data[['State','density_Group']].drop_duplicates().sort_values(by='State')
data['Positive'].sort_values()
data.to_csv('data.csv',index=False)
stateDensity.to_csv('stateDensity.csv',index=False)
data.head()
data.describe()
# +
#Rearrange dataframe
df=pd.DataFrame({'Dense1':data[data['density_Group']=='Dense1']['Positive'],
'Dense2':data[data['density_Group']=='Dense2']['Positive'],
'Dense3':data[data['density_Group']=='Dense3']['Positive'],
'Dense4':data[data['density_Group']=='Dense4']['Positive']})
# -
data.isna().sum()
data[data['Positive'].isna()]
df.dtypes
# +
####################### Approach 1.##########3
# -
np.random.seed(1234)
dataNew=pd.DataFrame({'Dense1':random.sample(list(data['Positive'][data['density_Group']=='Dense1']), 10),
'Dense2':random.sample(list(data['Positive'][data['density_Group']=='Dense1']), 10),
'Dense3':random.sample(list(data['Positive'][data['density_Group']=='Dense1']), 10),
'Dense4':random.sample(list(data['Positive'][data['density_Group']=='Dense1']), 10)})
# +
# np.random.seed(1234)
# dataNew=pd.DataFrame({'Dense1':data['Positive'][data['density_Group']=='Dense1'],
# 'Dense2':data['Positive'][data['density_Group']=='Dense1'],
# 'Dense3':data['Positive'][data['density_Group']=='Dense1'],
# 'Dense4':data['Positive'][data['density_Group']=='Dense1']})
# -
dataNew.head()
dataNew.describe()
dataNew['Dense1'].sort_values().head()
dataNew.describe()
dataNew['Dense1'].sort_values().head()
# +
#Plot number of Corona cases across different density groups to check their distribution.
fig = plt.figure(figsize=(10,10))
title = fig.suptitle("Corona cases across different density groups", fontsize=14)
fig.subplots_adjust(top=0.85, wspace=0.3)
ax1 = fig.add_subplot(2,2,1)
ax1.set_title("density Group-Dense1 & Corona Cases")
ax1.set_xlabel("density Group -Dense1")
ax1.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense1'], ax=ax1, shade=True,bw=4, color='g')
ax2 = fig.add_subplot(2,2,2)
ax2.set_title("density Group -Dense2 & Corona Cases")
ax2.set_xlabel("density Group -Dense2")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense2'], ax=ax2, shade=True,bw=4, color='y')
ax2 = fig.add_subplot(2,2,3)
ax2.set_title("density Group -Dense2 & Corona Cases")
ax2.set_xlabel("density Group -Dense3")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense3'], ax=ax2, shade=True,bw=4, color='r')
ax2 = fig.add_subplot(2,2,4)
ax2.set_title("density Group -Dense4 & Corona Cases")
ax2.set_xlabel("density Group -Dense4")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense4'], ax=ax2, shade=True,bw=4, color='b')
# -
## Apply BoxCox Transformation to bring the data to close to Gaussian Distribution
dataNew['Dense1'],fitted_lambda = stats.boxcox(dataNew['Dense1'])
dataNew['Dense2'],fitted_lambda = stats.boxcox(dataNew['Dense2'])
dataNew['Dense3'],fitted_lambda = stats.boxcox(dataNew['Dense3'])
dataNew['Dense4'],fitted_lambda = stats.boxcox(dataNew['Dense4'])
# +
##Apply log transformation to treat outliers and to bring to normal distribution
# dataNew = np.log(dataNew + 1)
# -
dataNew.describe()
dataNew.head()
dataNew['Dense1'].describe()
# +
#Plot different density groups
fig = plt.figure(figsize=(10,10))
title = fig.suptitle("Corona cases across different density groups", fontsize=14)
fig.subplots_adjust(top=0.85, wspace=0.3)
ax1 = fig.add_subplot(2,2,1)
ax1.set_title("density Group-Dense1 & Corona Cases")
ax1.set_xlabel("density Group -Dense1")
ax1.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense1'], ax=ax1, shade=True,bw=4, color='g')
ax2 = fig.add_subplot(2,2,2)
ax2.set_title("density Group -Dense2 & Corona Cases")
ax2.set_xlabel("density Group -Dense2")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense2'], ax=ax2, shade=True,bw=4, color='y')
ax2 = fig.add_subplot(2,2,3)
ax2.set_title("density Group -Dense2 & Corona Cases")
ax2.set_xlabel("density Group -Dense3")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense3'], ax=ax2, shade=True,bw=4, color='r')
ax2 = fig.add_subplot(2,2,4)
ax2.set_title("density Group -Dense4 & Corona Cases")
ax2.set_xlabel("density Group -Dense4")
ax2.set_ylabel("Corona Cases")
sns.kdeplot(dataNew['Dense4'], ax=ax2, shade=True,bw=4, color='b')
# -
##############Assumptions check - Normality
stats.shapiro(dataNew['Dense1'])
stats.shapiro(dataNew['Dense2'])
stats.shapiro(dataNew['Dense3'])
stats.shapiro(dataNew['Dense4'])
# Levene variance test
stats.levene(dataNew['Dense1'],dataNew['Dense2'],dataNew['Dense3'],dataNew['Dense4'])
# +
##p-value is more than 0.05 , So we can say that variances among groups are equal.
# -
F, p = stats.f_oneway(dataNew['Dense1'],dataNew['Dense2'],dataNew['Dense3'],dataNew['Dense4'])
print('F statistic =', F, 'p value :',p)
F, p = stats.f_oneway(dataNew['Dense1'],dataNew['Dense2'],dataNew['Dense3'],dataNew['Dense4'])
# Seeing if the overall model is significant
print('F-Statistic=%.3f, p=%.3f' % (F, p))
#Rearrange DataFrame
newDf=dataNew.stack().to_frame().reset_index().rename(columns={'level_1':'density_Group',
0:'Count'})
del newDf['level_0']
################ using Ols Model
model = ols('Count ~ C(density_Group)', newDf).fit()
model.summary()
# Seeing if the overall model is significant
print(f"Overall model F({model.df_model: .0f},{model.df_resid: .0f}) = {model.fvalue: .3f}, p = {model.f_pvalue: .4f}")
# Creates the ANOVA table
res = sm.stats.anova_lm(model, typ= 2)
res
# +
#The F-statistic= 76.48 and the p-value= 8.557817e-33 which is indicating that there is an overall significant
#effect of density_Group on corona positive cases. However, we don’t know where the difference between desnity_groups is yet.
# This is in the post-hoc section.
#So Based on p-value we can reject the H0; that is there's no significant difference as per density of an area
#and number of corona cases
# +
#So what if you find statistical significance? Multiple comparison tests
#When you conduct an ANOVA, you are attempting to determine if there is a statistically significant difference among the groups.
#If you find that there is a difference, you will then need to examine where the group differences lay.
# -
newDf.dtypes
newDf.head()
#Post hoc test
mc = statsmodels.stats.multicomp.MultiComparison(newDf['Count'],newDf['density_Group'])
mc_results = mc.tukeyhsd()
print(mc_results)
# +
#tuckey HSD test clearly says that there's a significant difference between Group1 & Group4
# +
#Above results from Tukey HSD suggests that except Dense1-Dense4 groups, all other pairwise comparisons for number of
#corona cases rejects null hypothesis and indicates statistical significant differences.
# -
### Normality Assumption check
w, pvalue = stats.shapiro(model.resid)
print(w, pvalue)
#Homogeneity of variances Assumption check
w, pvalue = stats.bartlett(newDf['Count'][newDf['density_Group']=='Dense1'], newDf['Count'][newDf['density_Group']=='Dense2']
, newDf['Count'][newDf['density_Group']=='Dense3'], newDf['Count'][newDf['density_Group']=='Dense4'])
print(w, pvalue)
## Q-Q Plot for Normal Distribution check-
#Check the Normal distribution of residuals
res = model.resid
fig = sm.qqplot(res, line='s')
plt.show()
| .ipynb_checkpoints/One Way ANOVA Test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''base'': conda)'
# name: python3
# ---
# # Reading and Writing Files in Python
# In this section, we will learn some basic opearation about reading and writing files. Moreover, as a data scientist, building an accurate machine learning model is not the end of the project. We will showing you how to save and load your machine learning model in Python.This allows you to save your model to file and load it later in order to make predictions.
# ## Read txt file
txt_file_url = "../data/files/Python.txt"
f = open(txt_file_url, "r") #opens file with name of "Python.txt"
# read and print the entire file
print(f.read())
# remember to colse the file
f.close()
# Used the **readline()** method twice, we would get the first 2 lines because of Python's reading process.
f = open(txt_file_url, "r") #opens file with name of "Python.txt"
# read the 1st line
print(f.readline())
# read the next line
print(f.readline())
f.close()
# +
#opens file with name of "Python.txt"
f = open("files/Python.txt", "r")
myList = []
for line in f:
myList.append(line)
f.close()
print(myList)
print(myList[0])
print(myList[1])
# -
# ## Write txt file
# +
# Write file with name of "test.txt"
f = open("../data/files/test.txt","w")
f.write("I love Python.\n")
f.write("I will be a Python master.\n")
f.write("I need to keep learning!")
f.close()
# read and see the test.txt file
f = open("../data/files/test.txt","r")
print(f.read())
f.close()
# -
# ## Read csv file
# +
import csv
csvFile = open("../data/files/test.csv", "r")
reader = csv.reader(csvFile, delimiter=',')
# load the data in a dictionary
result = {}
for item in reader:
# ignore the first line
if reader.line_num == 1:
continue
result[item[0]] = item[1]
csvFile.close()
print(result)
# -
# ## Write csv file
# +
import csv
fileHeader = ["name", "age"]
d1 = ["Chris", "27"]
d2 = ["Ming", "26"]
csvFile = open("../data/files/write.csv", "w")
writer = csv.writer(csvFile)
writer = csv.writer(csvFile)
# write the head and data
writer.writerow(fileHeader)
writer.writerow(d1)
writer.writerow(d2)
# Here is another command
# writer.writerows([fileHeader, d1, d2])
csvFile.close()
# go to see the "write.csv" file.
# -
# You can find more information from the [documentation](https://docs.python.org/3.6/library/csv.html)
# ## Using Pandas to Read CSV file
# +
import pandas as pd
import numpy as np
data = pd.read_csv("../data/files/test.csv")
# data is data
print(data)
# extract the age data
Age = np.array(data.Age, dtype = 'double')
print(Age)
# reshap this age vector
Age = np.reshape(Age, [3,1])
print(Age)
# -
# Find more operation about Pandas in the [documentation](https://pandas.pydata.org/) and [cheatsheet](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf)
# ## Read Matlab file
# The functions --scipy.io.loadmat-- and --scipy.io.savemat-- allow you to read and write MATLAB files. You can read about them in the [documentation](https://docs.scipy.org/doc/scipy/reference/io.html).
# +
import numpy as np
from scipy.io import loadmat, savemat
data = loadmat("../data/files/magic.mat");
print(data);
print(data['magic'])
# -
# ## Write Matlab file
# +
X = np.array(data['magic'])
# Do some calculation
X = X*2
# Dictionary from which to save matfile variables.
data = {'magic2': X}
# save the data
savemat("../data/files/magic2.mat", data)
# Go to matlab and check the data
data = loadmat("../data/files/magic2.mat");
print(data['magic2'])
# -
# ## Save and Load file by Pickle
# The Pickle pacakge is used for serializing and de-serializing a Python object structure. Any object in python can be pickled so that it can be saved on disk and loaded back to continue the work.
# You can read about them in the [documentation](https://docs.python.org/3.6/library/pickle.html?highlight=pickle#module-pickle).
import numpy as np
import pickle
X = np.eye(5)
print(X)
# Save the matirx X
with open('../data/files/X.pickle', 'wb') as f:
pickle.dump(X, f)
# Change the value of the original X
X = X + 4
print(X)
# load the matrix
with open('files/X.pickle', 'rb') as f:
X = pickle.load(f)
print(X)
# # One Example
# In this demonstration, we will use a Logistic Regression Model and the Iris dataset.
# +
from sklearn.linear_model import LogisticRegression
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
# Load and split data
data = load_iris()
Xtrain, Xtest, Ytrain, Ytest = train_test_split(data.data, data.target, test_size=0.3, random_state=4)
# Create a model
model = LogisticRegression(C=0.1,
max_iter=2000,
fit_intercept=True
)
model.fit(Xtrain, Ytrain)
print(model);
# -
# In the following few lines of code, the model which we created in the previous step is saved to file, and then loaded as a new object called pickled_model. The loaded model is then used to calculate the accuracy score and predict outcomes on new unseen (test) data.
# +
import pickle
#
# Create your model here (same as above)
#
# Save to file in the current working directory
pkl_filename = "../data/pickle_model.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(model, file)
# Load from file
with open(pkl_filename, 'rb') as file:
pickle_model = pickle.load(file)
# Calculate the accuracy score and predict target values
score = pickle_model.score(Xtest, Ytest)
print("Test score: {0:.2f} %".format(100 * score))
Ypredict = pickle_model.predict(Xtest)
# -
# Use python to read and write the `yaml` file.
import yaml
yaml_url = "../data/test.yaml"
with open(yaml_url, encoding='utf-8') as file:
data = yaml.safe_load(file)
print(data)
print(data['case1']['json'])
print(data['case1']['json']['username'])
# +
import yaml
content = {
'id': 1,
'text': 'programming languages',
'members': ['java', 'python', 'python', 'c', 'go', 'shell'],
'next': {'a':1,'b':2}
}
save_path = '../data/test_save.yaml'
with open(save_path, 'w', encoding='utf-8') as file:
yaml.dump(content, file, default_flow_style=False, encoding='utf-8', allow_unicode=True)
with open(save_path, encoding='utf-8') as file:
data = yaml.safe_load(file)
print(data)
# +
# import pyyaml module
import yaml
from yaml.loader import SafeLoader
# Open the file and load the file
with open(save_path) as f:
data = yaml.load(f, Loader=SafeLoader)
print(data)
# +
import yaml
from yaml.loader import SafeLoader
with open(save_path, 'r') as f:
data = list(yaml.load_all(f, Loader=SafeLoader))
print(data)
# +
import yaml
# dict object
members = [{'name': 'Zoey', 'occupation': 'Doctor'},
{'name': 'Zaara', 'occupation': 'Dentist'}]
# Convert Python dictionary into a YAML document
print(yaml.dump(members))
# +
import yaml
save_path = '../data/model_config.yaml'
user_details = { 'model_name':'vgg19',
'w1': [1, 2, 3, 4, 5.0],
'AccessKeys': ['EmployeeTable',
'SoftwaresList',
'HardwareList']}
with open(save_path, 'w') as f:
data = yaml.dump(user_details, f, sort_keys=False, default_flow_style=False)
with open(save_path, 'r') as f:
# data = yaml.load_all(f, Loader=SafeLoader)
data = yaml.safe_load(f)
print(data)
# -
type(data['w1'])
| 01.Python/Python_03_Read_and_Write_Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 네이버 플레이스
#
# ## 업체의 가게설명과 리뷰 데이터
#
# [네이버 지도](map.naver.com)에서 첫번째 검색결과로 검색되는 업체의 데이터를 가져옴. 그렇기 때문에 검색결과에 따라 원하는 업체의 데이터가 뽑히지 않을 수 있음.
from naverplacescraper import NaverPlace
store_name = '나폴리회관'
store = NaverPlace(store_name)
store
# ### 간단한 업체 정보
#
# `NaverPlace.info`: 검색된 업체의 간단한 정보
store.info
# ### 가게 설명 텍스트 데이터
# `NaverPlace.get_description()`: 업체에서 등록한 가게 설명 정보를 추출
description = store.get_description()
description
# ### 리뷰 텍스트 데이터
# `NaverPlace.get_reviews(num_of_reviews: int)`: 리뷰 작성 날짜, 글쓴이, 리뷰 데이터만 담고 있음
reviews = store.get_reviews() # num_of_reviews defaults to 100
reviews
# `NaverPlace.raw_review_data`: response 받은 리뷰 데이터가 리스트로 담겨있음
# First review
store.raw_review_data[0]
# ### 네이버 플레이스 검색 결과 리스트 데이터
search_results = store.search_result # list of search results
search_results
# ### 다른 지역 검색
#
# 기본적으로 '서울'을 중심으로 검색하게 되어 있으므로 `location`에 원하는 지역을 설정해주면 해당 지역을 중심으로 검색하게 됨.
store = NaverPlace('애플망고1947', location='제주') # location defaults to '서울'
description = store.get_description()
reviews = store.get_reviews()
description
reviews
# ### 여러 업체 리뷰 데이터 추출
#
# 업체 리스트를 가지고 있고 리스트에 있는 모든 업체들의 리뷰 데이터를 추출함
import pandas as pd
import time
from tqdm.notebook import tqdm_notebook
from IPython.display import display, clear_output
# +
store_list = ['나폴리회관', '애플망고1947', '스타벅스']
real_store_name = []
res = []
for i in tqdm_notebook(store_list):
try:
store = NaverPlace(i)
reviews = store.get_reviews(5)
review_list = reviews['review'].tolist()
name = store.info['name']
real_store_name.append(name)
res.append(review_list)
clear_output(wait=True)
time.sleep(5)
except:
continue
# -
review_df = pd.DataFrame(zip(real_store_name, res), columns=['store', 'review'])
review_df
# ### Http headers 설정
#
# 기본적으로 GET request를 보낼때 User-Agent가 포함된 헤더가 추가됨. 만약 헤더에 내용을 추가 하고 싶다면 `NaverPlace.attach_headers(headers: dict)`를 사용.
print(f'Default headers: {store.headers}')
headers = {
'User-Agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_7) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/98.0.4758.80 Safari/537.36',
}
store.attach_headers(headers=headers)
print(f'New custom headers: {store.headers}')
| example/naverplace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
def unit_sort(ele):
return str(ele)[-1]
test_list = [23,54,91,108,102]
print("The original list is : " + str(test_list))
test_list.sort(key=unit_sort)
print("The unit sorted list : " + str(test_list))
# +
x = [23,54,91,108,102]
a = x[0]
print(a)
b = str(a)
print(b)
m = b[-1]
print(m)
print('in short-->')
unit_digit = str(x[0])[-1]
print(unit_digit)
# -
| notebooks/sort the list on the ascending order of the ones place digit elements .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Exploration of Cedefop Skills Forecast 2030 data set
# <NAME> | 04.05.2021
#
# ## Core Analysis Goal(s)
# 1. Understand structure of dataset
#
# ## Key Insight(s)
# 1.
# 2.
# 3.
# +
import os
import sys
import logging
from pathlib import Path
import numpy as np
import scipy as sp
import statsmodels.api as sm
from statsmodels.formula.api import ols
# %load_ext autoreload
# %autoreload 2
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set_context("poster")
sns.set(rc={'figure.figsize': (16, 9.)})
sns.set_style("ticks")
import pandas as pd
pd.set_option("display.max_rows", 120)
pd.set_option("display.max_columns", 120)
logging.basicConfig(level=logging.INFO, stream=sys.stdout)
# + [markdown] pycharm={"name": "#%% md\n"}
# Define directory structure
# + pycharm={"name": "#%%\n"}
# project directory
abspath = os.path.abspath('')
project_dir = str(Path(abspath).parents[0])
# sub-directories
data_raw = os.path.join(project_dir, "data", "raw")
data_interim = os.path.join(project_dir, "data", "interim")
data_processed = os.path.join(project_dir, "data", "processed")
figure_dir = os.path.join(project_dir, "plots")
# + [markdown] pycharm={"name": "#%% md\n"}
# Cedefop Skills Forecast data set components
# + pycharm={"name": "#%%\n"}
dtypes_supply = {
"popLF": 'category',
"year": 'category',
"country": 'category',
"gender": 'category',
"ageband": 'category',
"qualification": 'category',
"adjustment": 'category',
"weight": "float"
}
dtypes_demand = {
"year": 'category',
"country": 'category',
"industry": 'category',
"occupation": 'category',
"qualification": 'category',
"adjustment": 'category',
"weight": "float"
}
demand = pd.read_csv(os.path.join(data_interim, "cedefop_skills_forecast", "Demand.csv"))
supply = pd.read_csv(os.path.join(data_interim, "cedefop_skills_forecast", "Supply.csv"))
lookups = pd.read_csv(os.path.join(data_interim, "cedefop_skills_forecast", "lookups.csv"))
# + pycharm={"name": "#%%\n"}
supply
demand
# + [markdown] pycharm={"name": "#%% md\n"}
# Decode supply and demand data using lookup table
# + pycharm={"name": "#%%\n"}
supply_decoded = supply.copy()
for col in supply_decoded.columns:
print(col)
renamer = dict(zip(
lookups[lookups.variable == col].drop(columns="variable").value.values,
lookups[lookups.variable == col].drop(columns="variable").name.values
))
supply_decoded[col] = supply_decoded[col].replace(to_replace=renamer)
supply_decoded = supply_decoded.astype(dtypes_supply)
supply_decoded.to_csv(os.path.join(data_processed, "cedefop_skills_forecast", "Supply_decoded.csv"))
supply_decoded.info()
# + pycharm={"name": "#%%\n"}
demand_decoded = demand.copy()
for col in demand_decoded.columns:
print(col)
renamer = dict(zip(
lookups[lookups.variable == col].drop(columns="variable").value.values,
lookups[lookups.variable == col].drop(columns="variable").name.values
))
demand_decoded[col] = demand_decoded[col].replace(to_replace=renamer)
demand_decoded = demand_decoded.astype(dtypes_demand)
demand_decoded.to_csv(os.path.join(data_processed, "cedefop_skills_forecast", "Demand_decoded.csv"))
demand_decoded.info()
# + pycharm={"name": "#%%\n"}
# test = pd.read_csv(os.path.join(data_processed, "cedefop_skills_forecast", "Supply_decoded.csv"), dtype=dtypes_supply, index_col=0)
# test.info()
| notebooks/02-fz-cedefop-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/elenado-1997/DLTK-week-3/blob/master/Week3_ageregression_sexclassification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="XHQUhXv7JYUn" outputId="ba289f21-3c74-40d2-a9cc-baba614ce6bf"
# !git clone https://github.com/DLTK/DLTK.git
# + [markdown] id="vyoiQc8yJuCU"
# Changed line 139 to `na_values=[]).values`
# for /content/DLTK/data/IXI_HH/download_IXI_HH.py
#
# because pandas code is outdated: https://pandas.pydata.org/pandas-docs/version/0.25.1/reference/api/pandas.DataFrame.as_matrix.html
# + colab={"base_uri": "https://localhost:8080/"} id="NbCRINWMJoj2" outputId="e7c62782-7a22-438e-e258-52f7df289b63"
# !pip install SimpleITK
# + colab={"base_uri": "https://localhost:8080/"} id="smfEDtcNJwoh" outputId="d1aa386c-04dd-4f16-c6d3-7cf7d80aff86"
# !python /content/DLTK/data/IXI_HH/download_IXI_HH.py
# + id="HPEQLQBSKiVn"
# !pwd
| Week3_ageregression_sexclassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cadquery as cq
from jupyter_cadquery import set_defaults, set_sidecar, get_defaults
from jupyter_cadquery.cadquery import show, PartGroup, Part, Faces, Edges
from jupyter_cadquery.cad_animation import Animation
from cadquery_massembly import Mate, MAssembly, relocate
# remove "clean" to avoid errors OCP kernel error
cq.occ_impl.shapes.Shape.clean = lambda x: x
set_sidecar("DiskArm", init=True)
set_defaults(axes=True, axes0=True, edge_accuracy=0.01, mate_scale=4, zoom=3.5, bb_factor=1.2)
# +
from math import pi, sin, cos, sqrt, asin, degrees, radians
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
# %matplotlib inline
# -
# # Disk and Arm
# ## Model
# 
# +
r_disk = 100
dist_pivot = 200
def angle_arm(angle_disk):
ra = np.deg2rad(angle_disk)
v = np.array((dist_pivot, 0)) - r_disk * np.array((cos(ra), sin(ra)))
return np.rad2deg(np.arctan2(*v[::-1]))
# -
# ## Visualisation
def diagram(da, aa, i1, i2):
a1, a2 = radians(da), radians(aa)
r1, r2 = r_disk, dist_pivot + r_disk
x1, y1 = (0, r1 * cos(a1)), (0, r1 * sin(a1))
x2, y2 = (dist_pivot, dist_pivot - r2 * cos(a2)), (0, -r2 * sin(a2))
ax = fig.add_subplot(spec[i1, i2])
ax.set_ylim(-r_disk-10, r_disk+10)
ax.set_title(f"a={da}")
ax.plot((0, dist_pivot), (0,0), color="lightgrey")
ax.plot(x1, y1)
ax.plot(x2, y2, ":")
circle = plt.Circle((0,0), 100, fill=False, color="lightgrey", linestyle='--')
ax.add_patch(circle)
# +
disk_angles = range(0, 360, 45)
arm_angles = [angle_arm(d) for d in disk_angles]
fig = plt.figure(constrained_layout=True)
fig.set_size_inches(20, 1.75)
spec = gridspec.GridSpec(ncols=8, nrows=1, figure=fig)
for i, (da, aa) in enumerate(zip(disk_angles, arm_angles)):
diagram(da, aa, 0, i)
# -
# # Assembly
# ## Parts
# +
thickness = 5
nr = 5
disk = cq.Workplane().circle(r_disk + 2 * nr).extrude(thickness)
nipple = cq.Workplane().circle(nr).extrude(thickness)
disk = (
disk
.cut(nipple)
.union(nipple.translate((r_disk, 0, thickness)))
)
pivot_base = cq.Workplane().circle(2*nr).extrude(thickness)
base = (
cq.Workplane()
.rect(6 * nr + dist_pivot, 6 * nr)
.extrude(thickness)
.translate((dist_pivot / 2, 0, 0))
.union(nipple.translate((dist_pivot, 0, thickness)))
.union(pivot_base.translate((0, 0, thickness)))
.union(nipple.translate((0, 0, 2*thickness)))
.edges("|Z").fillet(3)
)
base.faces(">Z[-2]").wires(cq.NearestToPointSelector((dist_pivot + r_disk, 0))).tag("mate")
slot = (cq.Workplane()
.rect(2*r_disk, 2*nr)
.extrude(thickness)
.union(nipple.translate((-r_disk, 0, 0)))
.union(nipple.translate((r_disk, 0, 0)))
.translate((dist_pivot, 0, 0))
)
arm = (
cq.Workplane()
.rect(4 * nr + (r_disk + dist_pivot), 4 * nr)
.extrude(thickness)
.edges("|Z").fillet(3)
.translate(((r_disk + dist_pivot) / 2, 0, 0))
.cut(nipple)
.cut(slot)
)
arm.faces(">Z").wires(cq.NearestToPointSelector((0,0))).tag("mate")
show(
disk,
base.translate((0, -1.5 * r_disk, 0)),
arm.translate((0, 1.5 * r_disk, 0)),
)
# -
# ## Define assembly
def create_disk_arm():
L = lambda *args: cq.Location(cq.Vector(*args))
C = lambda *args: cq.Color(*args)
return (MAssembly(base, name="base", color=C("gray"), loc=L(-dist_pivot/2, 0, 0))
.add(disk, name="disk", color=C("MediumAquaMarine"), loc=L(r_disk, -1.5 * r_disk, 0))
.add(arm, name="arm", color=C("orange"), loc=L(0, 10*nr, 0))
)
# ## Define mates
# +
from collections import OrderedDict as odict
disk_arm = create_disk_arm()
disk_arm.mate("base?mate", name="disk_pivot", origin=True, transforms=odict(rz=180))
disk_arm.mate("base@faces@>Z", name="arm_pivot")
disk_arm.mate("disk@faces@>Z[-2]", name="disk", origin=True)
disk_arm.mate("arm?mate", name="arm", origin=True)
show(disk_arm, render_mates=True)
# -
# ## Relocate and assemble
# +
# ensure all parts are relocated so that the origin mates is the part origin
relocate(disk_arm)
# assemble each part
disk_arm.assemble("arm", "arm_pivot")
disk_arm.assemble("disk", "disk_pivot")
d = show(disk_arm, render_mates=True, axes=False)
# -
# # Animate
# +
animation = Animation(d.root_group)
times = np.linspace(0, 5, 181)
disk_angles = np.linspace(0, 360, 181)
arm_angles = [angle_arm(d) for d in disk_angles]
# move disk
# Note, the selector must follow the path in the CAD view navigation hierarchy
animation.add_track(f"base/disk", "rz", times, disk_angles)
# move arm
animation.add_track(f"base/arm", "rz", times, arm_angles)
animation.animate(speed=2)
# -
| examples/assemblies/1-disk-arm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
PROJECT = "PROJECT" # REPLACE WITH YOUR PROJECT ID
BUCKET = "BUCKET" # REPLACE WITH A BUCKET NAME (PUT YOUR PROJECT ID AND WE CREATE THE BUCKET ITSELF NEXT)
REGION = "us-east1" # REPLACE WITH YOUR REGION e.g. us-central1
# Import os environment variables
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
os.environ["REGION"] = REGION
os.environ["TFVERSION"] = "1.13"
# -
# ## Now write into a python module
# +
# %%writefile lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer/model.py
import tensorflow as tf
# Set logging to be level of INFO
tf.logging.set_verbosity(tf.logging.INFO)
# Determine CSV and label columns
number_of_tags = 5
tag_columns = ["tag_{0}".format(tag) for tag in range(0, number_of_tags)]
UNLABELED_CSV_COLUMNS = tag_columns
LABEL_COLUMN = "anomalous_sequence_flag"
LABELED_CSV_COLUMNS = UNLABELED_CSV_COLUMNS + [LABEL_COLUMN]
# Set default values for each CSV column
UNLABELED_DEFAULTS = [[""] for _ in UNLABELED_CSV_COLUMNS]
LABELED_DEFAULTS = UNLABELED_DEFAULTS + [[0.0]]
# Create an input function reading a file using the Dataset API
# Then provide the results to the Estimator API
def read_dataset(filename, mode, batch_size, params):
def _input_fn():
def decode_csv(value_column, seq_len):
def convert_sequences_from_strings_to_floats(features, column_list):
def split_and_convert_string(string_tensor):
# Split string tensor into a sparse tensor based on delimiter
split_string = tf.string_split(source = tf.expand_dims(
input = string_tensor, axis = 0), delimiter = ",")
# Converts the values of the sparse tensor to floats
converted_tensor = tf.string_to_number(
string_tensor = split_string.values,
out_type = tf.float64)
# Create a new sparse tensor with the new converted values,
# because the original sparse tensor values are immutable
new_sparse_tensor = tf.SparseTensor(
indices = split_string.indices,
values = converted_tensor,
dense_shape = split_string.dense_shape)
# Create a dense tensor of the float values that were converted from text csv
dense_floats = tf.sparse_tensor_to_dense(
sp_input = new_sparse_tensor, default_value = 0.0)
dense_floats_vector = tf.squeeze(input = dense_floats, axis = 0)
return dense_floats_vector
for column in column_list:
features[column] = split_and_convert_string(features[column])
features[column].set_shape([seq_len])
return features
if mode == tf.estimator.ModeKeys.TRAIN or (mode == tf.estimator.ModeKeys.EVAL and params["evaluation_mode"] != "tune_anomaly_thresholds"):
columns = tf.decode_csv(
records = value_column,
record_defaults = UNLABELED_DEFAULTS,
field_delim = ";")
features = dict(zip(UNLABELED_CSV_COLUMNS, columns))
features = convert_sequences_from_strings_to_floats(
features, UNLABELED_CSV_COLUMNS)
return features
else:
columns = tf.decode_csv(
records = value_column,
record_defaults = LABELED_DEFAULTS,
field_delim = ";")
features = dict(zip(LABELED_CSV_COLUMNS, columns))
labels = tf.cast(x = features.pop(LABEL_COLUMN), dtype = tf.float64)
features = convert_sequences_from_strings_to_floats(
features, LABELED_CSV_COLUMNS[0:-1])
return features, labels
# Create list of files that match pattern
file_list = tf.gfile.Glob(filename = filename)
# Create dataset from file list
dataset = tf.data.TextLineDataset(filenames = file_list) # Read text file
# Decode the CSV file into a features dictionary of tensors
dataset = dataset.map(map_func = lambda x: decode_csv(x, params["seq_len"]))
# Determine amount of times to repeat file based on if we are training or evaluating
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # indefinitely
else:
num_epochs = 1 # end-of-input after this
# Repeat files num_epoch times
dataset = dataset.repeat(count = num_epochs)
# Group the data into batches
dataset = dataset.batch(batch_size = batch_size)
# Determine if we should shuffle based on if we are training or evaluating
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(buffer_size = 10 * batch_size)
# Create a iterator and then pull the next batch of features from the example queue
batched_dataset = dataset.make_one_shot_iterator().get_next()
return batched_dataset
return _input_fn
def create_LSTM_stack(lstm_hidden_units, lstm_dropout_output_keep_probs):
# First create a list of LSTM cells using our list of lstm hidden unit sizes
lstm_cells = [tf.contrib.rnn.BasicLSTMCell(
num_units = units,
forget_bias = 1.0,
state_is_tuple = True) for units in lstm_hidden_units] # list of LSTM cells
# Next apply a dropout wrapper to our stack of LSTM cells, in this case just on the outputs
dropout_lstm_cells = [tf.nn.rnn_cell.DropoutWrapper(
cell = lstm_cells[cell_index],
input_keep_prob = 1.0,
output_keep_prob = lstm_dropout_output_keep_probs[cell_index],
state_keep_prob = 1.0) for cell_index in range(len(lstm_cells))]
# Create a stack of layers of LSTM cells
stacked_lstm_cells = tf.contrib.rnn.MultiRNNCell(
cells = dropout_lstm_cells,
state_is_tuple = True) # combines list into MultiRNNCell object
return stacked_lstm_cells
# The rnn_decoder function takes labels during TRAIN/EVAL
# and a start token followed by its previous predictions during PREDICT
# Starts with an intial state of the final encoder states
def rnn_decoder(decoder_inputs, initial_state, cell, inference, dnn_hidden_units, num_features):
# Create the decoder variable scope
with tf.variable_scope("decoder"):
# Load in our initial state from our encoder
state = initial_state # tuple of final encoder c_state and h_state of final encoder layer
# Create an empty list to store our hidden state output for every timestep
outputs = []
# Begin with no previous output
previous_output = None
# Loop over all of our decoder_inputs which will be seq_len long
for index, decoder_input in enumerate(decoder_inputs):
# If there has been a previous output then we will determine the next input
if previous_output is not None:
# Create the input layer to our DNN
network = previous_output # shape = (cur_batch_size, lstm_hidden_units[-1])
# Create our dnn variable scope
with tf.variable_scope(name_or_scope = "dnn", reuse = tf.AUTO_REUSE):
# Add hidden layers with the given number of units/neurons per layer
# shape = (cur_batch_size, dnn_hidden_units[i])
for units in dnn_hidden_units:
network = tf.layers.dense(
inputs = network,
units = units,
activation = tf.nn.relu)
# Connect final hidden layer to linear layer to get the logits
logits = tf.layers.dense(
inputs = network,
units = num_features,
activation = None) # shape = (cur_batch_size, num_features)
# If we are in inference then we will overwrite our next decoder_input
# with the logits we just calculated.
# Otherwise, we leave the decoder_input input as it was from the enumerated list
# We have to calculate the logits even when not using them so that the correct
# dnn subgraph will be generated here and after the encoder-decoder for both
# training and inference
if inference == True:
decoder_input = logits # shape = (cur_batch_size, num_features)
# If this isn"t our first time through the loop, just reuse(share) the same
# variables for each iteration within the current variable scope
if index > 0:
tf.get_variable_scope().reuse_variables()
# Run the decoder input through the decoder stack picking up from the previous state
# output_shape = (cur_batch_size, lstm_hidden_units[-1])
# state_shape = # tuple of final decoder c_state and h_state
output, state = cell(decoder_input, state)
# Append the current decoder hidden state output to the outputs list
# list eventually seq_len long of shape = (cur_batch_size, lstm_hidden_units[-1])
outputs.append(output)
# Set the previous output to the output just calculated
previous_output = output # shape = (cur_batch_size, lstm_hidden_units[-1])
return outputs, state
# This function updates the count of records used
def update_count(count_a, count_b):
return count_a + count_b
# This function updates the mahalanobis distance variables when number_of_rows equals 1
def singleton_batch_mahalanobis_distance_variable_updating(
inner_size,
X,
count_variable,
mean_variable,
cov_variable,
inverse_cov_variable,
eps):
# This function updates the mean vector incrementally
def update_mean_incremental(count_a, mean_a, value_b):
mean_ab = (mean_a * tf.cast(x = count_a, dtype = tf.float64) + \
tf.squeeze(input = value_b, axis = 0)) / tf.cast(x = count_a + 1, dtype = tf.float64)
return mean_ab
# This function updates the covariance matrix incrementally
def update_cov_incremental(count_a, mean_a, cov_a, value_b, mean_ab, sample_cov):
if sample_cov == True:
cov_ab = (cov_a * tf.cast(x = count_a - 1, dtype = tf.float64) + \
tf.matmul(a = value_b - mean_a, b = value_b - mean_ab, transpose_a = True)) \
/ tf.cast(x = count_a, dtype = tf.float64)
else:
cov_ab = (cov_a * tf.cast(x = count_a, dtype = tf.float64) + \
tf.matmul(a = value_b - mean_a, b = value_b - mean_ab, transpose_a = True)) \
/ tf.cast(x = count_a + 1, dtype = tf.float64)
return cov_ab
# Calculate new combined mean to use for incremental covariance matrix calculation
mean_ab = update_mean_incremental(
count_a = count_variable,
mean_a = mean_variable,
value_b = X) # time_shape = (num_features,), features_shape = (sequence_length,)
# Update running variables from single example
count_tensor = update_count(
count_a = count_variable,
count_b = 1) # time_shape = (), features_shape = ()
mean_tensor = mean_ab # time_shape = (num_features,), features_shape = (sequence_length,)
if inner_size == 1:
cov_tensor = tf.zeros_like(
tensor = cov_variable, dtype = tf.float64)
inverse_cov_tensor = tf.eye(
num_rows = tf.shape(
input = cov_tensor)[0],
dtype = tf.float64) / eps
else:
# time_shape = (num_features, num_features)
# features_shape = (sequence_length, sequence_length)
cov_tensor = update_cov_incremental(
count_a = count_variable,
mean_a = mean_variable,
cov_a = cov_variable,
value_b = X,
mean_ab = mean_ab,
sample_cov = True)
# time_shape = (num_features, num_features)
# features_shape = (sequence_length, sequence_length)
inverse_cov_tensor = tf.matrix_inverse(
input = cov_tensor + tf.eye(
num_rows = tf.shape(
input = cov_tensor)[0],
dtype = tf.float64) * eps)
# Assign values to variables, use control dependencies around return to enforce the mahalanobis
# variables to be assigned, the control order matters, hence the separate contexts
with tf.control_dependencies(
control_inputs = [tf.assign(
ref = cov_variable,
value = cov_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(
ref = mean_variable,
value = mean_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(
ref = count_variable,
value = count_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(
ref = inverse_cov_variable,
value = inverse_cov_tensor)]):
return tf.identity(input = cov_variable), tf.identity(input = mean_variable), tf.identity(input = count_variable), tf.identity(input = inverse_cov_variable)
# This function updates the mahalanobis distance variables when number_of_rows does NOT equal 1
def non_singleton_batch_mahalanobis_distance_variable_updating(
cur_batch_size,
inner_size,
X,
count_variable,
mean_variable,
cov_variable,
inverse_cov_variable,
eps):
# This function updates the mean vector using a batch of data
def update_mean_batch(count_a, mean_a, count_b, mean_b):
mean_ab = (mean_a * tf.cast(x = count_a, dtype = tf.float64) + \
mean_b * tf.cast(x = count_b, dtype = tf.float64)) \
/ tf.cast(x = count_a + count_b, dtype = tf.float64)
return mean_ab
# This function updates the covariance matrix using a batch of data
def update_cov_batch(count_a, mean_a, cov_a, count_b, mean_b, cov_b, sample_cov):
mean_diff = tf.expand_dims(input = mean_a - mean_b, axis = 0)
if sample_cov == True:
cov_ab = (cov_a * tf.cast(x = count_a - 1, dtype = tf.float64) + \
cov_b * tf.cast(x = count_b - 1, dtype = tf.float64) + \
tf.matmul(a = mean_diff, b = mean_diff, transpose_a = True) * \
tf.cast(x = count_a * count_b, dtype = tf.float64) \
/ tf.cast(x = count_a + count_b, dtype = tf.float64)) \
/ tf.cast(x = count_a + count_b - 1, dtype = tf.float64)
else:
cov_ab = (cov_a * tf.cast(x = count_a, dtype = tf.float64) + \
cov_b * tf.cast(x = count_b, dtype = tf.float64) + \
tf.matmul(a = mean_diff, b = mean_diff, transpose_a = True) * \
tf.cast(x = count_a * count_b, dtype = tf.float64) \
/ tf.cast(x = count_a + count_b, dtype = tf.float64)) \
/ tf.cast(x = count_a + count_b, dtype = tf.float64)
return cov_ab
# Find statistics of batch
number_of_rows = cur_batch_size * inner_size
# time_shape = (num_features,), features_shape = (sequence_length,)
X_mean = tf.reduce_mean(input_tensor = X, axis = 0)
# time_shape = (cur_batch_size * sequence_length, num_features)
# features_shape = (cur_batch_size * num_features, sequence_length)
X_centered = X - X_mean
if inner_size > 1:
# time_shape = (num_features, num_features)
# features_shape = (sequence_length, sequence_length)
X_cov = tf.matmul(
a = X_centered,
b = X_centered,
transpose_a = True) / tf.cast(x = number_of_rows - 1, dtype = tf.float64)
# Update running variables from batch statistics
count_tensor = update_count(
count_a = count_variable,
count_b = number_of_rows) # time_shape = (), features_shape = ()
mean_tensor = update_mean_batch(
count_a = count_variable,
mean_a = mean_variable,
count_b = number_of_rows,
mean_b = X_mean) # time_shape = (num_features,), features_shape = (sequence_length,)
if inner_size == 1:
cov_tensor = tf.zeros_like(
tensor = cov_variable, dtype = tf.float64)
inverse_cov_tensor = tf.eye(
num_rows = tf.shape(input = cov_tensor)[0], dtype = tf.float64) / eps
else:
# time_shape = (num_features, num_features)
# features_shape = (sequence_length, sequence_length)
cov_tensor = update_cov_batch(
count_a = count_variable,
mean_a = mean_variable,
cov_a = cov_variable,
count_b = number_of_rows,
mean_b = X_mean,
cov_b = X_cov,
sample_cov = True)
# time_shape = (num_features, num_features)
# features_shape = (sequence_length, sequence_length)
inverse_cov_tensor = tf.matrix_inverse(
input = cov_tensor + \
tf.eye(num_rows = tf.shape(input = cov_tensor)[0],
dtype = tf.float64) * eps)
# Assign values to variables, use control dependencies around return to enforce the mahalanobis
# variables to be assigned, the control order matters, hence the separate contexts
with tf.control_dependencies(
control_inputs = [tf.assign(ref = cov_variable, value = cov_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(ref = mean_variable, value = mean_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(ref = count_variable, value = count_tensor)]):
with tf.control_dependencies(
control_inputs = [tf.assign(ref = inverse_cov_variable, value = inverse_cov_tensor)]):
return tf.identity(input = cov_variable), tf.identity(input = mean_variable), tf.identity(input = count_variable), tf.identity(input = inverse_cov_variable)
def mahalanobis_distance(error_vectors_reshaped, mean_vector, inv_covariance, final_shape):
# time_shape = (current_batch_size * seq_len, num_features)
# features_shape = (current_batch_size * num_features, seq_len)
error_vectors_reshaped_centered = error_vectors_reshaped - mean_vector
# time_shape = (num_features, current_batch_size * seq_len)
# features_shape = (seq_len, current_batch_size * num_features)
mahalanobis_right_product = tf.matmul(
a = inv_covariance,
b = error_vectors_reshaped_centered,
transpose_b = True)
# time_shape = (current_batch_size * seq_len, current_batch_size * seq_len)
# features_shape = (current_batch_size * num_features, current_batch_size * num_features)
mahalanobis_distance_vectorized = tf.matmul(
a = error_vectors_reshaped_centered,
b = mahalanobis_right_product)
# time_shape = (current_batch_size * seq_len,)
# features_shape = (current_batch_size * num_features,)
mahalanobis_distance_flat = tf.diag_part(input = mahalanobis_distance_vectorized)
# time_shape = (current_batch_size, seq_len)
# features_shape = (current_batch_size, num_features)
mahalanobis_distance_final_shaped = tf.reshape(
tensor = mahalanobis_distance_flat,
shape = [-1, final_shape])
# time_shape = (current_batch_size, seq_len)
# features_shape = (current_batch_size, num_features)
mahalanobis_distance_final_shaped_abs = tf.abs(x = mahalanobis_distance_final_shaped)
return mahalanobis_distance_final_shaped_abs
def update_anomaly_threshold_variables(
labels_normal_mask,
labels_anomalous_mask,
num_thresholds,
anomaly_thresholds,
mahalanobis_distance,
tp_at_thresholds_variable,
fn_at_thresholds_variable,
fp_at_thresholds_variable,
tn_at_thresholds_variable,
mode):
if mode == tf.estimator.ModeKeys.TRAIN:
# time_shape = (num_time_anomaly_thresholds, current_batch_size, sequence_length)
# features_shape = (num_features_anomaly_thresholds, current_batch_size, number_of_features)
mahalanobis_distance_over_thresholds = tf.map_fn(
fn = lambda anomaly_threshold: mahalanobis_distance > anomaly_threshold,
elems = anomaly_thresholds,
dtype = tf.bool)
else:
# time_shape = (current_batch_size, sequence_length)
# features_shape = (current_batch_size, number_of_features)
mahalanobis_distance_over_thresholds = mahalanobis_distance > anomaly_thresholds
# time_shape = (num_time_anomaly_thresholds, current_batch_size)
# features_shape = (num_features_anomaly_thresholds, current_batch_size)
mahalanobis_distance_any_over_thresholds = tf.reduce_any(
input_tensor = mahalanobis_distance_over_thresholds,
axis = -1)
if mode == tf.estimator.ModeKeys.EVAL:
# time_shape = (1, current_batch_size)
# features_shape = (1, current_batch_size)
mahalanobis_distance_any_over_thresholds = tf.expand_dims(
input = mahalanobis_distance_any_over_thresholds, axis = 0)
# time_shape = (num_time_anomaly_thresholds, current_batch_size)
# features_shape = (num_features_anomaly_thresholds, current_batch_size)
predicted_normals = tf.equal(
x = mahalanobis_distance_any_over_thresholds,
y = False)
# time_shape = (num_time_anomaly_thresholds, current_batch_size)
# features_shape = (num_features_anomaly_thresholds, current_batch_size)
predicted_anomalies = tf.equal(
x = mahalanobis_distance_any_over_thresholds,
y = True)
# Calculate confusion matrix of current batch
# time_shape = (num_time_anomaly_thresholds,)
# features_shape = (num_features_anomaly_thresholds,)
tp = tf.reduce_sum(
input_tensor = tf.cast(
x = tf.map_fn(
fn = lambda threshold: tf.logical_and(
x = labels_anomalous_mask,
y = predicted_anomalies[threshold, :]),
elems = tf.range(start = 0, limit = num_thresholds, dtype = tf.int64),
dtype = tf.bool),
dtype = tf.int64),
axis = 1)
fn = tf.reduce_sum(
input_tensor = tf.cast(
x = tf.map_fn(
fn = lambda threshold: tf.logical_and(
x = labels_anomalous_mask,
y = predicted_normals[threshold, :]),
elems = tf.range(start = 0, limit = num_thresholds, dtype = tf.int64),
dtype = tf.bool),
dtype = tf.int64),
axis = 1)
fp = tf.reduce_sum(
input_tensor = tf.cast(
x = tf.map_fn(
fn = lambda threshold: tf.logical_and(
x = labels_normal_mask,
y = predicted_anomalies[threshold, :]),
elems = tf.range(start = 0, limit = num_thresholds, dtype = tf.int64),
dtype = tf.bool),
dtype = tf.int64),
axis = 1)
tn = tf.reduce_sum(
input_tensor = tf.cast(
x = tf.map_fn(
fn = lambda threshold: tf.logical_and(
x = labels_normal_mask,
y = predicted_normals[threshold, :]),
elems = tf.range(start = 0, limit = num_thresholds, dtype = tf.int64),
dtype = tf.bool),
dtype = tf.int64),
axis = 1)
if mode == tf.estimator.ModeKeys.EVAL:
# shape = ()
tp = tf.squeeze(input = tp)
fn = tf.squeeze(input = fn)
fp = tf.squeeze(input = fp)
tn = tf.squeeze(input = tn)
with tf.control_dependencies(
control_inputs = [tf.assign_add(ref = tp_at_thresholds_variable, value = tp),
tf.assign_add(ref = fn_at_thresholds_variable, value = fn),
tf.assign_add(ref = fp_at_thresholds_variable, value = fp),
tf.assign_add(ref = tn_at_thresholds_variable, value = tn)]):
return tf.identity(input = tp_at_thresholds_variable), tf.identity(input = fn_at_thresholds_variable), tf.identity(input = fp_at_thresholds_variable), tf.identity(input = tn_at_thresholds_variable)
def calculate_composite_classification_metrics(anomaly_thresholds, tp, fn, fp, tn, f_score_beta):
# time_shape = (num_time_anomaly_thresholds,)
# features_shape = (num_features_anomaly_thresholds,)
acc = tf.cast(x = tp + tn, dtype = tf.float64) \
/ tf.cast(x = tp + fn + fp + tn, dtype = tf.float64)
pre = tf.cast(x = tp, dtype = tf.float64) / tf.cast(x = tp + fp, dtype = tf.float64)
rec = tf.cast(x = tp, dtype = tf.float64) / tf.cast(x = tp + fn, dtype = tf.float64)
f_beta_score = (1.0 + f_score_beta ** 2) * (pre * rec) / (f_score_beta ** 2 * pre + rec)
return acc, pre, rec, f_beta_score
def find_best_anomaly_threshold(
anomaly_thresholds, f_beta_score, user_passed_anomaly_threshold, anomaly_threshold_variable):
if user_passed_anomaly_threshold == None:
best_anomaly_threshold = tf.gather(
params = anomaly_thresholds,
indices = tf.argmax(input = f_beta_score,
axis = 0)) # shape = ()
else:
best_anomaly_threshold = user_passed_anomaly_threshold # shape = ()
with tf.control_dependencies(
control_inputs = [
tf.assign(ref = anomaly_threshold_variable, value = best_anomaly_threshold)]):
return tf.identity(input = anomaly_threshold_variable)
# Create our model function to be used in our custom estimator
def lstm_encoder_decoder_autoencoder_anomaly_detection(features, labels, mode, params):
print("\nlstm_encoder_decoder_autoencoder_anomaly_detection: features = \n{}".format(features))
print("lstm_encoder_decoder_autoencoder_anomaly_detection: labels = \n{}".format(labels))
print("lstm_encoder_decoder_autoencoder_anomaly_detection: mode = \n{}".format(mode))
print("lstm_encoder_decoder_autoencoder_anomaly_detection: params = \n{}".format(params))
# 0. Get input sequence tensor into correct shape
# Get dynamic batch size in case there was a partially filled batch
cur_batch_size = tf.shape(input = features[UNLABELED_CSV_COLUMNS[0]], out_type = tf.int64)[0]
# Get the number of features
num_features = len(UNLABELED_CSV_COLUMNS)
# Stack all of the features into a 3-D tensor
# shape = (cur_batch_size, seq_len, num_features)
X = tf.stack(values = [features[key] for key in UNLABELED_CSV_COLUMNS], axis = 2)
# Unstack all of 3-D features tensor into a sequence(list) of 2-D tensors of
# shape = (cur_batch_size, num_features)
X_sequence = tf.unstack(value = X, num = params["seq_len"], axis = 1)
# Since this is an autoencoder, the features are the labels.
# It often works better though to have the labels in reverse order
if params["reverse_labels_sequence"] == True:
Y = tf.reverse_sequence(
input = X, # shape = (cur_batch_size, seq_len, num_features)
seq_lengths = tf.tile(
input = tf.constant(value = [params["seq_len"]], dtype = tf.int64),
multiples = tf.expand_dims(input = cur_batch_size, axis = 0)),
seq_axis = 1,
batch_axis = 0)
else:
Y = X # shape = (cur_batch_size, seq_len, num_features)
################################################################################
# 1. Create encoder of encoder-decoder LSTM stacks
# Create our decoder now
decoder_stacked_lstm_cells = create_LSTM_stack(
params["decoder_lstm_hidden_units"], params["lstm_dropout_output_keep_probs"])
# Create the encoder variable scope
with tf.variable_scope("encoder"):
# Create separate encoder cells with their own weights separate from decoder
encoder_stacked_lstm_cells = create_LSTM_stack(
params["encoder_lstm_hidden_units"], params["lstm_dropout_output_keep_probs"])
# Encode the input sequence using our encoder stack of LSTMs
# encoder_outputs = seq_len long of shape = (cur_batch_size, encoder_lstm_hidden_units[-1])
# encoder_states = tuple of final encoder c_state and h_state for each layer
encoder_outputs, encoder_states = tf.nn.static_rnn(
cell = encoder_stacked_lstm_cells,
inputs = X_sequence,
initial_state = encoder_stacked_lstm_cells.zero_state(
batch_size = tf.cast(x = cur_batch_size, dtype = tf.int32),
dtype = tf.float64),
dtype = tf.float64)
# We just pass on the final c and h states of the encoder"s last layer,
# so extract that and drop the others
# LSTMStateTuple shape = (cur_batch_size, lstm_hidden_units[-1])
encoder_final_states = encoder_states[-1]
# Extract the c and h states from the tuple
# both have shape = (cur_batch_size, lstm_hidden_units[-1])
encoder_final_c, encoder_final_h = encoder_final_states
# In case the decoder"s first layer"s number of units is different than encoder's last
# layer's number of units, use a dense layer to map to the correct shape
encoder_final_c_dense = tf.layers.dense(
inputs = encoder_final_c,
units = params["decoder_lstm_hidden_units"][0],
activation = None) # shape = (cur_batch_size, decoder_lstm_hidden_units[0])
encoder_final_h_dense = tf.layers.dense(
inputs = encoder_final_h,
units = params["decoder_lstm_hidden_units"][0],
activation = None) # shape = (cur_batch_size, decoder_lstm_hidden_units[0])
# The decoder"s first layer"s state comes from the encoder,
# the rest of the layers" initial states are zero
decoder_intial_states = tuple(
[tf.contrib.rnn.LSTMStateTuple(c = encoder_final_c_dense, h = encoder_final_h_dense)] + \
[tf.contrib.rnn.LSTMStateTuple(
c = tf.zeros(shape = [cur_batch_size, units], dtype = tf.float64),
h = tf.zeros(shape = [cur_batch_size, units], dtype = tf.float64))
for units in params["decoder_lstm_hidden_units"][1:]])
################################################################################
# 2. Create decoder of encoder-decoder LSTM stacks
# Train our decoder now
# Encoder-decoders work differently during training/evaluation and inference
# so we will have two separate subgraphs for each
if mode == tf.estimator.ModeKeys.TRAIN and params["evaluation_mode"] == "reconstruction":
# Break 3-D labels tensor into a list of 2-D tensors of shape = (cur_batch_size, num_features)
unstacked_labels = tf.unstack(value = Y, num = params["seq_len"], axis = 1)
# Call our decoder using the labels as our inputs, the encoder final state as our
# initial state, our other LSTM stack as our cells, and inference set to false
decoder_outputs, decoder_states = rnn_decoder(
decoder_inputs = unstacked_labels,
initial_state = decoder_intial_states,
cell = decoder_stacked_lstm_cells,
inference = False,
dnn_hidden_units = params["dnn_hidden_units"],
num_features = num_features)
else:
# Since this is inference create fake labels. The list length needs to be the output
# sequence length even though only the first element is the only one actually used
# (as our go signal)
fake_labels = [tf.zeros(shape = [cur_batch_size, num_features], dtype = tf.float64)
for _ in range(params["seq_len"])]
# Call our decoder using fake labels as our inputs, the encoder final state as our initial
# state, our other LSTM stack as our cells, and inference set to true
# decoder_outputs = seq_len long of shape = (cur_batch_size, decoder_lstm_hidden_units[-1])
# decoder_states = tuple of final decoder c_state and h_state for each layer
decoder_outputs, decoder_states = rnn_decoder(
decoder_inputs = fake_labels,
initial_state = decoder_intial_states,
cell = decoder_stacked_lstm_cells,
inference = True,
dnn_hidden_units = params["dnn_hidden_units"],
num_features = num_features)
# Stack together the list of rank 2 decoder output tensors into one rank 3 tensor of
# shape = (cur_batch_size, seq_len, lstm_hidden_units[-1])
stacked_decoder_outputs = tf.stack(values = decoder_outputs, axis = 1)
# Reshape rank 3 decoder outputs into rank 2 by folding sequence length into batch size
# shape = (cur_batch_size * seq_len, lstm_hidden_units[-1])
reshaped_stacked_decoder_outputs = tf.reshape(
tensor = stacked_decoder_outputs,
shape = [cur_batch_size * params["seq_len"], params["decoder_lstm_hidden_units"][-1]])
################################################################################
# 3. Create the DNN structure now after the encoder-decoder LSTM stack
# Create the input layer to our DNN
# shape = (cur_batch_size * seq_len, lstm_hidden_units[-1])
network = reshaped_stacked_decoder_outputs
# Reuse the same variable scope as we used within our decoder (for inference)
with tf.variable_scope(name_or_scope = "dnn", reuse = tf.AUTO_REUSE):
# Add hidden layers with the given number of units/neurons per layer
for units in params["dnn_hidden_units"]:
network = tf.layers.dense(
inputs = network,
units = units,
activation = tf.nn.relu) # shape = (cur_batch_size * seq_len, dnn_hidden_units[i])
# Connect the final hidden layer to a dense layer with no activation to get the logits
logits = tf.layers.dense(
inputs = network,
units = num_features,
activation = None) # shape = (cur_batch_size * seq_len, num_features)
# Now that we are through the final DNN for each sequence element for each example in the batch,
# reshape the predictions to match our labels.
# shape = (cur_batch_size, seq_len, num_features)
predictions = tf.reshape(
tensor = logits,
shape = [cur_batch_size, params["seq_len"], num_features])
# Variables for calculating error distribution statistics
with tf.variable_scope(
name_or_scope = "mahalanobis_distance_variables", reuse = tf.AUTO_REUSE):
# Time based
abs_err_count_time_variable = tf.get_variable(
name = "abs_err_count_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
abs_err_mean_time_variable = tf.get_variable(
name = "abs_err_mean_time_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [num_features], dtype = tf.float64),
trainable = False) # shape = (num_features,)
abs_err_cov_time_variable = tf.get_variable(
name = "abs_err_cov_time_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [num_features, num_features], dtype = tf.float64),
trainable = False) # shape = (num_features, num_features)
abs_err_inv_cov_time_variable = tf.get_variable(
name = "abs_err_inv_cov_time_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [num_features, num_features], dtype = tf.float64),
trainable = False) # shape = (num_features, num_features)
# Features based
abs_err_count_features_variable = tf.get_variable(
name = "abs_err_count_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
abs_err_mean_features_variable = tf.get_variable(
name = "abs_err_mean_features_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [params["seq_len"]], dtype = tf.float64),
trainable = False) # shape = (seq_len,)
abs_err_cov_features_variable = tf.get_variable(
name = "abs_err_cov_features_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [params["seq_len"], params["seq_len"]], dtype = tf.float64),
trainable = False) # shape = (seq_len, seq_len)
abs_err_inv_cov_features_variable = tf.get_variable(
name = "abs_err_inv_cov_features_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [params["seq_len"], params["seq_len"]], dtype = tf.float64),
trainable = False) # shape = (seq_len, seq_len)
# Variables for automatically tuning anomaly thresholds
with tf.variable_scope(
name_or_scope = "mahalanobis_distance_threshold_variables", reuse = tf.AUTO_REUSE):
# Time based
tp_at_thresholds_time_variable = tf.get_variable(
name = "tp_at_thresholds_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_time_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_time_anomaly_thresholds,)
fn_at_thresholds_time_variable = tf.get_variable(
name = "fn_at_thresholds_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_time_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_time_anomaly_thresholds,)
fp_at_thresholds_time_variable = tf.get_variable(
name = "fp_at_thresholds_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_time_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_time_anomaly_thresholds,)
tn_at_thresholds_time_variable = tf.get_variable(
name = "tn_at_thresholds_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_time_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_time_anomaly_thresholds,)
time_anomaly_threshold_variable = tf.get_variable(
name = "time_anomaly_threshold_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [], dtype = tf.float64),
trainable = False) # shape = ()
# Features based
tp_at_thresholds_features_variable = tf.get_variable(
name = "tp_at_thresholds_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_features_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_features_anomaly_thresholds,)
fn_at_thresholds_features_variable = tf.get_variable(
name = "fn_at_thresholds_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_features_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_features_anomaly_thresholds,)
fp_at_thresholds_features_variable = tf.get_variable(
name = "fp_at_thresholds_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_features_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_features_anomaly_thresholds,)
tn_at_thresholds_features_variable = tf.get_variable(
name = "tn_at_thresholds_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [params["num_features_anomaly_thresholds"]], dtype = tf.int64),
trainable = False) # shape = (num_features_anomaly_thresholds,)
features_anomaly_threshold_variable = tf.get_variable(
name = "features_anomaly_threshold_variable", # shape = ()
dtype = tf.float64,
initializer = tf.zeros(shape = [], dtype = tf.float64),
trainable = False)
# Variables for automatically tuning anomaly thresholds
with tf.variable_scope(
name_or_scope = "anomaly_threshold_eval_variables", reuse = tf.AUTO_REUSE):
# Time based
tp_at_threshold_eval_time_variable = tf.get_variable(
name = "tp_at_threshold_eval_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
fn_at_threshold_eval_time_variable = tf.get_variable(
name = "fn_at_threshold_eval_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
fp_at_threshold_eval_time_variable = tf.get_variable(
name = "fp_at_threshold_eval_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
tn_at_threshold_eval_time_variable = tf.get_variable(
name = "tn_at_threshold_eval_time_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
# Features based
tp_at_threshold_eval_features_variable = tf.get_variable(
name = "tp_at_threshold_eval_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
fn_at_threshold_eval_features_variable = tf.get_variable(
name = "fn_at_threshold_eval_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
fp_at_threshold_eval_features_variable = tf.get_variable(
name = "fp_at_threshold_eval_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
tn_at_threshold_eval_features_variable = tf.get_variable(
name = "tn_at_threshold_eval_features_variable",
dtype = tf.int64,
initializer = tf.zeros(shape = [], dtype = tf.int64),
trainable = False) # shape = ()
dummy_variable = tf.get_variable(
name = "dummy_variable",
dtype = tf.float64,
initializer = tf.zeros(shape = [], dtype = tf.float64),
trainable = True) # shape = ()
# Now branch off based on which mode we are in
predictions_dict = None
loss = None
train_op = None
eval_metric_ops = None
export_outputs = None
# 3. Loss function, training/eval ops
if mode == tf.estimator.ModeKeys.TRAIN and params["evaluation_mode"] != "tune_anomaly_thresholds":
if params["evaluation_mode"] == "reconstruction":
loss = tf.losses.mean_squared_error(labels = Y, predictions = predictions)
train_op = tf.contrib.layers.optimize_loss(
loss = loss,
global_step = tf.train.get_global_step(),
learning_rate = params["learning_rate"],
optimizer = "Adam")
elif params["evaluation_mode"] == "calculate_error_distribution_statistics":
error = Y - predictions # shape = (cur_batch_size, seq_len, num_features)
absolute_error = tf.abs(x = error) # shape = (cur_batch_size, seq_len, num_features)
################################################################################
with tf.variable_scope(
name_or_scope = "mahalanobis_distance_variables", reuse = tf.AUTO_REUSE):
# Check if batch is a singleton or not, very important for covariance math
# Time based ########################################
# shape = (cur_batch_size * seq_len, num_features)
abs_err_reshaped_time = tf.reshape(
tensor = absolute_error,
shape = [cur_batch_size * params["seq_len"], num_features])
singleton_time_condition = tf.equal(
x = cur_batch_size * params["seq_len"], y = 1) # shape = ()
cov_time_update_op, mean_time_update_op, count_time_update_op, inv_time_update_op = tf.cond(
pred = singleton_time_condition,
true_fn = lambda: singleton_batch_mahalanobis_distance_variable_updating(
params["seq_len"],
abs_err_reshaped_time,
abs_err_count_time_variable,
abs_err_mean_time_variable,
abs_err_cov_time_variable,
abs_err_inv_cov_time_variable,
params["eps"]),
false_fn = lambda: non_singleton_batch_mahalanobis_distance_variable_updating(
cur_batch_size,
params["seq_len"],
abs_err_reshaped_time,
abs_err_count_time_variable,
abs_err_mean_time_variable,
abs_err_cov_time_variable,
abs_err_inv_cov_time_variable,
params["eps"]))
# Features based ########################################
# shape = (cur_batch_size, num_features, seq_len)
abs_err_transposed_features = tf.transpose(a = absolute_error, perm = [0, 2, 1])
# shape = (cur_batch_size * num_features, seq_len)
abs_err_reshaped_features = tf.reshape(
tensor = abs_err_transposed_features,
shape = [cur_batch_size * num_features, params["seq_len"]])
# shape = ()
singleton_features_condition = tf.equal(x = cur_batch_size * num_features, y = 1)
cov_features_update_op, mean_features_update_op, count_features_update_op, inv_features_update_op = tf.cond(
pred = singleton_features_condition,
true_fn = lambda: singleton_batch_mahalanobis_distance_variable_updating(
num_features,
abs_err_reshaped_features,
abs_err_count_features_variable,
abs_err_mean_features_variable,
abs_err_cov_features_variable,
abs_err_inv_cov_features_variable,
params["eps"]),
false_fn = lambda: non_singleton_batch_mahalanobis_distance_variable_updating(
cur_batch_size,
num_features,
abs_err_reshaped_features,
abs_err_count_features_variable,
abs_err_mean_features_variable,
abs_err_cov_features_variable,
abs_err_inv_cov_features_variable,
params["eps"]))
# Lastly use control dependencies around loss to enforce the mahalanobis variables to be
# assigned, the control order matters, hence the separate contexts
with tf.control_dependencies(control_inputs = [cov_time_update_op, cov_features_update_op]):
with tf.control_dependencies(control_inputs = [mean_time_update_op, mean_features_update_op]):
with tf.control_dependencies(control_inputs = [count_time_update_op, count_features_update_op]):
with tf.control_dependencies(control_inputs = [inv_time_update_op, inv_features_update_op]):
loss = tf.reduce_sum(input_tensor = tf.zeros(shape = (), dtype = tf.float64) * dummy_variable)
train_op = tf.contrib.layers.optimize_loss(
loss = loss,
global_step = tf.train.get_global_step(),
learning_rate = params["learning_rate"],
optimizer = "SGD")
elif mode == tf.estimator.ModeKeys.EVAL and params["evaluation_mode"] != "tune_anomaly_thresholds":
# Reconstruction loss on evaluation set
loss = tf.losses.mean_squared_error(labels = Y, predictions = predictions)
if params["evaluation_mode"] == "reconstruction":
# Reconstruction eval metrics
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(labels = Y, predictions = predictions),
"mae": tf.metrics.mean_absolute_error(labels = Y, predictions = predictions)
}
elif mode == tf.estimator.ModeKeys.PREDICT or ((mode == tf.estimator.ModeKeys.TRAIN or mode == tf.estimator.ModeKeys.EVAL) and params["evaluation_mode"] == "tune_anomaly_thresholds"):
error = Y - predictions # shape = (cur_batch_size, seq_len, num_features)
absolute_error = tf.abs(x = error) # shape = (cur_batch_size, seq_len, num_features)
with tf.variable_scope(name_or_scope = "mahalanobis_distance_variables", reuse = tf.AUTO_REUSE):
# Time based
# shape = (cur_batch_size * seq_len, num_features)
abs_err_reshaped_time = tf.reshape(
tensor = absolute_error,
shape = [cur_batch_size * params["seq_len"], num_features])
mahalanobis_distance_time = mahalanobis_distance(
error_vectors_reshaped = abs_err_reshaped_time,
mean_vector = abs_err_mean_time_variable,
inv_covariance = abs_err_inv_cov_time_variable,
final_shape = params["seq_len"]) # shape = (cur_batch_size, seq_len)
# Features based
abs_err_mapped_features = tf.map_fn(
fn = lambda x: tf.transpose(a = absolute_error[x, :, :]),
elems = tf.range(start = 0, limit = cur_batch_size, dtype = tf.int64),
dtype = tf.float64) # shape = (cur_batch_size, num_features, seq_len)
# shape = (cur_batch_size * num_features, seq_len)
abs_err_reshaped_features = tf.reshape(
tensor = abs_err_mapped_features,
shape = [cur_batch_size * num_features, params["seq_len"]])
mahalanobis_distance_features = mahalanobis_distance(
error_vectors_reshaped = abs_err_reshaped_features,
mean_vector = abs_err_mean_features_variable,
inv_covariance = abs_err_inv_cov_features_variable,
final_shape = num_features) # shape = (cur_batch_size, num_features)
if mode != tf.estimator.ModeKeys.PREDICT:
labels_normal_mask = tf.equal(x = labels, y = 0)
labels_anomalous_mask = tf.equal(x = labels, y = 1)
if mode == tf.estimator.ModeKeys.TRAIN:
with tf.variable_scope(
name_or_scope = "mahalanobis_distance_variables", reuse = tf.AUTO_REUSE):
# Time based
# shape = (num_time_anomaly_thresholds,)
time_anomaly_thresholds = tf.linspace(
start = tf.constant(value = params["min_time_anomaly_threshold"], dtype = tf.float64),
stop = tf.constant(value = params["max_time_anomaly_threshold"], dtype = tf.float64),
num = params["num_time_anomaly_thresholds"])
tp_time_update_op, fn_time_update_op, fp_time_update_op, tn_time_update_op = \
update_anomaly_threshold_variables(
labels_normal_mask,
labels_anomalous_mask,
params["num_time_anomaly_thresholds"],
time_anomaly_thresholds,
mahalanobis_distance_time,
tp_at_thresholds_time_variable,
fn_at_thresholds_time_variable,
fp_at_thresholds_time_variable,
tn_at_thresholds_time_variable,
mode)
# Features based
# shape = (num_features_anomaly_thresholds,)
features_anomaly_thresholds = tf.linspace(
start = tf.constant(value = params["min_features_anomaly_threshold"], dtype = tf.float64),
stop = tf.constant(value = params["max_features_anomaly_threshold"], dtype = tf.float64),
num = params["num_features_anomaly_thresholds"])
tp_features_update_op, fn_features_update_op, fp_features_update_op, tn_features_update_op = \
update_anomaly_threshold_variables(
labels_normal_mask,
labels_anomalous_mask,
params["num_features_anomaly_thresholds"],
features_anomaly_thresholds,
mahalanobis_distance_features,
tp_at_thresholds_features_variable,
fn_at_thresholds_features_variable,
fp_at_thresholds_features_variable,
tn_at_thresholds_features_variable,
mode)
# Reconstruction loss on evaluation set
with tf.control_dependencies(
control_inputs = [
tp_time_update_op,
fn_time_update_op,
fp_time_update_op,
tn_time_update_op,
tp_features_update_op,
fn_features_update_op,
fp_features_update_op,
tn_features_update_op]):
# Time based
acc_time, pre_time, rec_time, f_beta_score_time = \
calculate_composite_classification_metrics(
time_anomaly_thresholds,
tp_at_thresholds_time_variable,
fn_at_thresholds_time_variable,
fp_at_thresholds_time_variable,
tn_at_thresholds_time_variable,
params["f_score_beta"])
# Features based
acc_features, pre_features, rec_features, f_beta_score_features = \
calculate_composite_classification_metrics(
features_anomaly_thresholds,
tp_at_thresholds_features_variable,
fn_at_thresholds_features_variable,
fp_at_thresholds_features_variable,
tn_at_thresholds_features_variable,
params["f_score_beta"])
with tf.control_dependencies(
control_inputs = [pre_time, pre_features]):
with tf.control_dependencies(
control_inputs = [rec_time, rec_features]):
with tf.control_dependencies(
control_inputs = [f_beta_score_time, f_beta_score_features]):
# Time based
best_anomaly_threshold_time = find_best_anomaly_threshold(
time_anomaly_thresholds,
f_beta_score_time,
params["time_anomaly_threshold"],
time_anomaly_threshold_variable)
# Features based
best_anomaly_threshold_features = find_best_anomaly_threshold(
features_anomaly_thresholds,
f_beta_score_features,
params["features_anomaly_threshold"],
features_anomaly_threshold_variable)
with tf.control_dependencies(
control_inputs = [
tf.assign(
ref = time_anomaly_threshold_variable,
value = best_anomaly_threshold_time),
tf.assign(ref =
features_anomaly_threshold_variable,
value = best_anomaly_threshold_features)]):
loss = tf.reduce_sum(
input_tensor = tf.zeros(shape = (), dtype = tf.float64) * dummy_variable)
train_op = tf.contrib.layers.optimize_loss(
loss = loss,
global_step = tf.train.get_global_step(),
learning_rate = params["learning_rate"],
optimizer = "SGD")
elif mode == tf.estimator.ModeKeys.EVAL:
with tf.variable_scope(
name_or_scope = "anomaly_threshold_eval_variables", reuse = tf.AUTO_REUSE):
# Time based
tp_time_update_op, fn_time_update_op, fp_time_update_op, tn_time_update_op = \
update_anomaly_threshold_variables(
labels_normal_mask,
labels_anomalous_mask,
1,
time_anomaly_threshold_variable,
mahalanobis_distance_time,
tp_at_threshold_eval_time_variable,
fn_at_threshold_eval_time_variable,
fp_at_threshold_eval_time_variable,
tn_at_threshold_eval_time_variable,
mode)
# Features based
tp_features_update_op, fn_features_update_op, fp_features_update_op, tn_features_update_op = \
update_anomaly_threshold_variables(
labels_normal_mask,
labels_anomalous_mask,
1,
features_anomaly_threshold_variable,
mahalanobis_distance_features,
tp_at_threshold_eval_features_variable,
fn_at_threshold_eval_features_variable,
fp_at_threshold_eval_features_variable,
tn_at_threshold_eval_features_variable,
mode)
with tf.variable_scope(
name_or_scope = "anomaly_threshold_eval_variables", reuse = tf.AUTO_REUSE):
# Time based
acc_time_update_op, pre_time_update_op, rec_time_update_op, f_beta_score_time_update_op = \
calculate_composite_classification_metrics(
time_anomaly_threshold_variable,
tp_at_threshold_eval_time_variable,
fn_at_threshold_eval_time_variable,
fp_at_threshold_eval_time_variable,
tn_at_threshold_eval_time_variable,
params["f_score_beta"])
# Features based
acc_features_update_op, pre_features_update_op, rec_features_update_op, f_beta_score_features_update_op = \
calculate_composite_classification_metrics(
features_anomaly_threshold_variable,
tp_at_threshold_eval_features_variable,
fn_at_threshold_eval_features_variable,
fp_at_threshold_eval_features_variable,
tn_at_threshold_eval_features_variable,
params["f_score_beta"])
loss = tf.losses.mean_squared_error(labels = Y, predictions = predictions)
acc_at_threshold_eval_time_variable = (tp_at_threshold_eval_time_variable + tn_at_threshold_eval_time_variable) / (tp_at_threshold_eval_time_variable + fn_at_threshold_eval_time_variable + fp_at_threshold_eval_time_variable + tn_at_threshold_eval_time_variable)
pre_at_threshold_eval_time_variable = tp_at_threshold_eval_time_variable / (tp_at_threshold_eval_time_variable + fp_at_threshold_eval_time_variable)
rec_at_threshold_eval_time_variable = tp_at_threshold_eval_time_variable / (tp_at_threshold_eval_time_variable + fn_at_threshold_eval_time_variable)
f_beta_score_at_threshold_eval_time_variable = (1.0 + params["f_score_beta"] ** 2) * pre_at_threshold_eval_time_variable * rec_at_threshold_eval_time_variable / (params["f_score_beta"] ** 2 * pre_at_threshold_eval_time_variable + rec_at_threshold_eval_time_variable)
acc_at_threshold_eval_features_variable = (tp_at_threshold_eval_features_variable + tn_at_threshold_eval_features_variable) / (tp_at_threshold_eval_features_variable + fn_at_threshold_eval_features_variable + fp_at_threshold_eval_features_variable + tn_at_threshold_eval_features_variable)
pre_at_threshold_eval_features_variable = tp_at_threshold_eval_features_variable / (tp_at_threshold_eval_features_variable + fp_at_threshold_eval_features_variable)
rec_at_threshold_eval_features_variable = tp_at_threshold_eval_features_variable / (tp_at_threshold_eval_features_variable + fn_at_threshold_eval_features_variable)
f_beta_score_at_threshold_eval_features_variable = (1.0 + params["f_score_beta"] ** 2) * pre_at_threshold_eval_features_variable * rec_at_threshold_eval_features_variable / (params["f_score_beta"] ** 2 * pre_at_threshold_eval_features_variable + rec_at_threshold_eval_features_variable)
# Anomaly detection eval metrics
eval_metric_ops = {
# Time based
"time_anomaly_tp": (tp_at_threshold_eval_time_variable, tp_time_update_op),
"time_anomaly_fn": (fn_at_threshold_eval_time_variable, fn_time_update_op),
"time_anomaly_fp": (fp_at_threshold_eval_time_variable, fp_time_update_op),
"time_anomaly_tn": (tn_at_threshold_eval_time_variable, tn_time_update_op),
"time_anomaly_acc": (acc_at_threshold_eval_time_variable, acc_time_update_op),
"time_anomaly_pre": (pre_at_threshold_eval_time_variable, pre_time_update_op),
"time_anomaly_rec": (rec_at_threshold_eval_time_variable, rec_time_update_op),
"time_anomaly_f_beta_score": (f_beta_score_at_threshold_eval_time_variable, f_beta_score_time_update_op),
# Features based
"features_anomaly_tp": (tp_at_threshold_eval_features_variable, tp_features_update_op),
"features_anomaly_fn": (fn_at_threshold_eval_features_variable, fn_features_update_op),
"features_anomaly_fp": (fp_at_threshold_eval_features_variable, fp_features_update_op),
"features_anomaly_tn": (tn_at_threshold_eval_features_variable, tn_features_update_op),
"features_anomaly_acc": (acc_at_threshold_eval_features_variable, acc_features_update_op),
"features_anomaly_pre": (pre_at_threshold_eval_features_variable, pre_features_update_op),
"features_anomaly_rec": (rec_at_threshold_eval_features_variable, rec_features_update_op),
"features_anomaly_f_beta_score": (f_beta_score_at_threshold_eval_features_variable, f_beta_score_features_update_op)
}
else: # mode == tf.estimator.ModeKeys.PREDICT
# Flag predictions as either normal or anomalous
time_anomaly_flags = tf.where(
condition = tf.reduce_any(
input_tensor = tf.greater(
x = tf.abs(x = mahalanobis_distance_time),
y = time_anomaly_threshold_variable),
axis = 1),
x = tf.ones(shape = [cur_batch_size], dtype = tf.int64),
y = tf.zeros(shape = [cur_batch_size], dtype = tf.int64)) # shape = (cur_batch_size,)
features_anomaly_flags = tf.where(
condition = tf.reduce_any(
input_tensor = tf.greater(
x = tf.abs(x = mahalanobis_distance_features),
y = features_anomaly_threshold_variable),
axis = 1),
x = tf.ones(shape = [cur_batch_size], dtype = tf.int64),
y = tf.zeros(shape = [cur_batch_size], dtype = tf.int64)) # shape = (cur_batch_size,)
# Create predictions dictionary
predictions_dict = {
"Y": Y,
"predictions": predictions,
"error": error,
"absolute_error": absolute_error,
"mahalanobis_distance_time": mahalanobis_distance_time,
"mahalanobis_distance_features": mahalanobis_distance_features,
"time_anomaly_flags": time_anomaly_flags,
"features_anomaly_flags": features_anomaly_flags}
# Create export outputs
export_outputs = {
"predict_export_outputs": tf.estimator.export.PredictOutput(
outputs = predictions_dict)}
# Return EstimatorSpec
return tf.estimator.EstimatorSpec(
mode = mode,
predictions = predictions_dict,
loss = loss,
train_op = train_op,
eval_metric_ops = eval_metric_ops,
export_outputs = export_outputs)
# Create our serving input function to accept the data at serving and send it in the
# right format to our custom estimator
def serving_input_fn(seq_len):
# This function fixes the shape and type of our input strings
def fix_shape_and_type_for_serving(placeholder):
current_batch_size = tf.shape(input = placeholder, out_type = tf.int64)[0]
# String split each string in batch and output values from the resulting SparseTensors
split_string = tf.stack(values = tf.map_fn( # shape = (batch_size, seq_len)
fn = lambda x: tf.string_split(source = [placeholder[x]], delimiter = ',').values,
elems = tf.range(start = 0, limit = current_batch_size, dtype = tf.int64),
dtype = tf.string), axis = 0)
# Convert each string in the split tensor to float
# shape = (batch_size, seq_len)
feature_tensor = tf.string_to_number(string_tensor = split_string, out_type = tf.float64)
return feature_tensor
# This function fixes dynamic shape ambiguity of last dimension so that we will be able to
# use it in our DNN (since tf.layers.dense require the last dimension to be known)
def get_shape_and_set_modified_shape_2D(tensor, additional_dimension_sizes):
# Get static shape for tensor and convert it to list
shape = tensor.get_shape().as_list()
# Set outer shape to additional_dimension_sizes[0] since know this is the correct size
shape[1] = additional_dimension_sizes[0]
# Set the shape of tensor to our modified shape
tensor.set_shape(shape = shape) # shape = (batch_size, additional_dimension_sizes[0])
return tensor
# Create placeholders to accept the data sent to the model at serving time
# All features come in as a batch of strings, shape = (batch_size,),
# this was so because of passing the arrays to online ml-engine prediction
feature_placeholders = {
feature: tf.placeholder(
dtype = tf.string, shape = [None]) for feature in UNLABELED_CSV_COLUMNS
}
# Create feature tensors
features = {key: fix_shape_and_type_for_serving(placeholder = tensor)
for key, tensor in feature_placeholders.items()}
# Fix dynamic shape ambiguity of feature tensors for our DNN
features = {key: get_shape_and_set_modified_shape_2D(
tensor = tensor, additional_dimension_sizes = [seq_len])
for key, tensor in features.items()}
return tf.estimator.export.ServingInputReceiver(
features = features, receiver_tensors = feature_placeholders)
# Create estimator to train and evaluate
def train_and_evaluate(args):
# Create our custom estimator using our model function
estimator = tf.estimator.Estimator(
model_fn = lstm_encoder_decoder_autoencoder_anomaly_detection,
model_dir = args["output_dir"],
params = {
"seq_len": args["seq_len"],
"reverse_labels_sequence": args["reverse_labels_sequence"],
"encoder_lstm_hidden_units": args["encoder_lstm_hidden_units"],
"decoder_lstm_hidden_units": args["decoder_lstm_hidden_units"],
"lstm_dropout_output_keep_probs": args["lstm_dropout_output_keep_probs"],
"dnn_hidden_units": args["dnn_hidden_units"],
"learning_rate": args["learning_rate"],
"evaluation_mode": args["evaluation_mode"],
"num_time_anomaly_thresholds": args["num_time_anomaly_thresholds"],
"num_features_anomaly_thresholds": args["num_features_anomaly_thresholds"],
"min_time_anomaly_threshold": args["min_time_anomaly_threshold"],
"max_time_anomaly_threshold": args["max_time_anomaly_threshold"],
"min_features_anomaly_threshold": args["min_features_anomaly_threshold"],
"max_features_anomaly_threshold": args["max_features_anomaly_threshold"],
"time_anomaly_threshold": args["time_anomaly_threshold"],
"features_anomaly_threshold": args["features_anomaly_threshold"],
"eps": args["eps"],
"f_score_beta": args["f_score_beta"]})
if args["evaluation_mode"] == "reconstruction":
early_stopping_hook = tf.contrib.estimator.stop_if_no_decrease_hook(
estimator = estimator,
metric_name = "rmse",
max_steps_without_decrease = 100,
min_steps = 1000,
run_every_secs = 60,
run_every_steps = None)
# Create train spec to read in our training data
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset(
filename = args["train_file_pattern"],
mode = tf.estimator.ModeKeys.TRAIN,
batch_size = args["train_batch_size"],
params = args),
max_steps = args["train_steps"],
hooks = [early_stopping_hook])
# Create eval spec to read in our validation data and export our model
eval_spec = tf.estimator.EvalSpec(
input_fn = read_dataset(
filename = args["eval_file_pattern"],
mode = tf.estimator.ModeKeys.EVAL,
batch_size = args["eval_batch_size"],
params = args),
steps = None,
start_delay_secs = args["start_delay_secs"], # start evaluating after N seconds
throttle_secs = args["throttle_secs"]) # evaluate every N seconds
# Create train and evaluate loop to train and evaluate our estimator
tf.estimator.train_and_evaluate(
estimator = estimator, train_spec = train_spec, eval_spec = eval_spec)
else:
if args["evaluation_mode"] == "calculate_error_distribution_statistics":
# Get final mahalanobis statistics over the entire validation_1 dataset
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset(
filename = args["train_file_pattern"],
mode = tf.estimator.ModeKeys.EVAL, # only read through validation dataset once
batch_size = args["train_batch_size"],
params = args),
max_steps = args["train_steps"])
# Don't create exporter for serving yet since anomaly thresholds aren't trained yet
exporter = None
elif args["evaluation_mode"] == "tune_anomaly_thresholds":
# Tune anomaly thresholds using valdiation_2 and validation_anomaly datasets
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset(
filename = args["train_file_pattern"],
mode = tf.estimator.ModeKeys.EVAL, # only read through validation dataset once
batch_size = args["train_batch_size"],
params = args),
max_steps = args["train_steps"])
# Create exporter that uses serving_input_fn to create saved_model for serving
exporter = tf.estimator.LatestExporter(
name = "exporter", serving_input_receiver_fn = lambda: serving_input_fn(args["sequence_length"]))
# Create eval spec to read in our validation data and export our model
eval_spec = tf.estimator.EvalSpec(
input_fn = read_dataset(
filename = args["eval_file_pattern"],
mode = tf.estimator.ModeKeys.EVAL,
batch_size = args["eval_batch_size"],
params = args),
steps = None,
exporters = exporter,
start_delay_secs = args["start_delay_secs"], # start evaluating after N seconds
throttle_secs = args["throttle_secs"]) # evaluate every N seconds
# Create train and evaluate loop to train and evaluate our estimator
tf.estimator.train_and_evaluate(
estimator = estimator, train_spec = train_spec, eval_spec = eval_spec)
# +
# %%writefile lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer/task.py
import argparse
import json
import os
from . import model
import tensorflow as tf
if __name__ == "__main__":
parser = argparse.ArgumentParser()
# File arguments
parser.add_argument(
"--train_file_pattern",
help = "GCS location to read training data",
required = True
)
parser.add_argument(
"--eval_file_pattern",
help = "GCS location to read evaluation data",
required = True
)
parser.add_argument(
"--output_dir",
help = "GCS location to write checkpoints and export models",
required = True
)
parser.add_argument(
"--job-dir",
help = "this model ignores this field, but it is required by gcloud",
default = "junk"
)
# Sequence shape hyperparameters
parser.add_argument(
"--seq_len",
help = "Number of timesteps to include in each example",
type = int,
default = 32
)
parser.add_argument(
"--horizon",
help = "Number of timesteps to skip into the future",
type = int,
default = 0
)
parser.add_argument(
"--reverse_labels_sequence",
help = "Whether we should reverse the labels sequence dimension or not",
type = bool,
default = True
)
# Architecture hyperparameters
# LSTM hyperparameters
parser.add_argument(
"--encoder_lstm_hidden_units",
help = "Hidden layer sizes to use for LSTM encoder",
default = "64 32 16"
)
parser.add_argument(
"--decoder_lstm_hidden_units",
help = "Hidden layer sizes to use for LSTM decoder",
default = "16 32 64"
)
parser.add_argument(
"--lstm_dropout_output_keep_probs",
help = "Keep probabilties for LSTM outputs",
default = "1.0 1.0 1.0"
)
# DNN hyperparameters
parser.add_argument(
"--dnn_hidden_units",
help = "Hidden layer sizes to use for DNN",
default = "1024 256 64"
)
# Training parameters
parser.add_argument(
"--train_batch_size",
help = "Number of examples in training batch",
type = int,
default = 32
)
parser.add_argument(
"--eval_batch_size",
help = "Number of examples in evaluation batch",
type = int,
default = 32
)
parser.add_argument(
"--train_steps",
help = "Number of batches to train for",
type = int,
default = 2000
)
parser.add_argument(
"--learning_rate",
help = "The learning rate, how quickly or slowly we train our model by scaling the gradient",
type = float,
default = 0.1
)
parser.add_argument(
"--start_delay_secs",
help = "Number of seconds to wait before first evaluation",
type = int,
default = 60
)
parser.add_argument(
"--throttle_secs",
help = "Number of seconds to wait between evaluations",
type = int,
default = 120
)
# Anomaly detection
parser.add_argument(
"--evaluation_mode",
help = "Which evaluation mode we are in (reconstruction, calculate_error_distribution_statistics, tune_anomaly_thresholds)",
type = str,
default = "reconstruction"
)
parser.add_argument(
"--num_time_anomaly_thresholds",
help = "Number of anomaly thresholds to evaluate in the time dimension",
type = int,
default = 120
)
parser.add_argument(
"--num_features_anomaly_thresholds",
help = "Number of anomaly thresholds to evaluate in the features dimension",
type = int,
default = 120
)
parser.add_argument(
"--min_time_anomaly_threshold",
help = "The minimum anomaly threshold to evaluate in the time dimension",
type = float,
default = 100.0
)
parser.add_argument(
"--max_time_anomaly_threshold",
help = "The maximum anomaly threshold to evaluate in the time dimension",
type = float,
default = 2000.0
)
parser.add_argument(
"--min_features_anomaly_threshold",
help = "The minimum anomaly threshold to evaluate in the time dimension",
type = float,
default = 100.0
)
parser.add_argument(
"--max_features_anomaly_threshold",
help = "The maximum anomaly threshold to evaluate in the time dimension",
type = float,
default = 2000.0
)
parser.add_argument(
"--time_anomaly_threshold",
help = "The anomaly threshold in the time dimension",
type = float,
default = None
)
parser.add_argument(
"--features_anomaly_threshold",
help = "The anomaly threshold in the features dimension",
type = float,
default = None
)
parser.add_argument(
"--eps",
help = "The precision value to add to the covariance matrix before inversion to avoid being singular",
type = str,
default = "1e-12"
)
parser.add_argument(
"--f_score_beta",
help = "The value of beta of the f-beta score",
type = float,
default = 0.05
)
# Parse all arguments
args = parser.parse_args()
arguments = args.__dict__
# Unused args provided by service
arguments.pop("job_dir", None)
arguments.pop("job-dir", None)
# Fix list arguments
arguments["encoder_lstm_hidden_units"] = [int(x)
for x in arguments["encoder_lstm_hidden_units"].split(' ')]
arguments["decoder_lstm_hidden_units"] = [int(x)
for x in arguments["decoder_lstm_hidden_units"].split(' ')]
arguments["lstm_dropout_output_keep_probs"] = [float(x)
for x in arguments["lstm_dropout_output_keep_probs"].split(' ')]
arguments["dnn_hidden_units"] = [int(x)
for x in arguments["dnn_hidden_units"].split(' ')]
# Fix eps argument
arguments["eps"] = float(arguments["eps"])
# Append trial_id to path if we are doing hptuning
# This code can be removed if you are not using hyperparameter tuning
arguments["output_dir"] = os.path.join(
arguments["output_dir"],
json.loads(
os.environ.get("TF_CONFIG", "{}")
).get("task", {}).get("trial", "")
)
# Run the training job
model.train_and_evaluate(arguments)
# -
# # Training model module
# ## Locally
# ### Train reconstruction variables
# + language="bash"
# rm -rf trained_model
# export PYTHONPATH=$PYTHONPATH:$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/training_normal_sequences.csv" \
# --eval_file_pattern="data/validation_normal_1_sequences.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --learning_rate=0.1 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --evaluation_mode="reconstruction" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300
# -
# ### Train error distribution statistics variables
# + language="bash"
# export PYTHONPATH=$PYTHONPATH:$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/validation_normal_1_sequences.csv" \
# --eval_file_pattern="data/validation_normal_1_sequences.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2200 \
# --evaluation_mode="calculate_error_distribution_statistics" \
# --eps="1e-12" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300
# -
# ### Tune anomaly thresholds
# + language="bash"
# export PYTHONPATH=$PYTHONPATH:$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module
# python -m trainer.task \
# --train_file_pattern="data/labeled_validation_mixed_sequences.csv" \
# --eval_file_pattern="data/labeled_validation_mixed_sequences.csv" \
# --output_dir=$PWD/trained_model \
# --job-dir=./tmp \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2400 \
# --evaluation_mode="tune_anomaly_thresholds" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300 \
# --min_time_anomaly_threshold=1.0 \
# --max_time_anomaly_threshold=20.0 \
# --min_features_anomaly_threshold=20.0 \
# --max_features_anomaly_threshold=80.0 \
# --f_score_beta=0.05
# -
# ## GCloud
# Copy data over to bucket
# + language="bash"
# gsutil -m cp -r data/* gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data
# -
# ### Train reconstruction variables
# + language="bash"
# OUTDIR=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/trained_model
# JOBNAME=job_lstm_encoder_decoder_autoencoder_anomaly_detection_reconstruction_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gsutil -m rm -rf $OUTDIR
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/training_normal_sequences.csv \
# --eval_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/validation_normal_1_sequences.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --learning_rate=0.1 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --evaluation_mode="reconstruction" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300
# -
# ### Hyperparameter tuning of reconstruction hyperparameters
# %%writefile hyperparam_reconstruction.yaml
trainingInput:
scaleTier: STANDARD_1
hyperparameters:
hyperparameterMetricTag: rmse
goal: MINIMIZE
maxTrials: 30
maxParallelTrials: 1
params:
- parameterName: encoder_lstm_hidden_units
type: CATEGORICAL
categoricalValues: ["64 32 16", "256 128 16", "64 64 64"]
- parameterName: decoder_lstm_hidden_units
type: CATEGORICAL
categoricalValues: ["16 32 64", "16 128 256", "64 64 64"]
- parameterName: lstm_dropout_output_keep_probs
type: CATEGORICAL
categoricalValues: ["0.9 1.0 1.0", "0.95 0.95 1.0", "0.95 0.95 0.95"]
- parameterName: dnn_hidden_units
type: CATEGORICAL
categoricalValues: ["256 128 64", "256 128 16", "64 64 64"]
- parameterName: train_batch_size
type: INTEGER
minValue: 8
maxValue: 512
scaleType: UNIT_LOG_SCALE
- parameterName: learning_rate
type: DOUBLE
minValue: 0.001
maxValue: 0.1
scaleType: UNIT_LINEAR_SCALE
# + language="bash"
# OUTDIR=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/hyperparam_reconstruction
# JOBNAME=job_lstm_encoder_decoder_autoencoder_anomaly_detection_hyperparam_reconstruction_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gsutil -m rm -rf $OUTDIR
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --config=hyperparam_reconstruction.yaml \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/training_normal_sequences.csv \
# --eval_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/validation_normal_1_sequences.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2000 \
# --start_delay_secs=60 \
# --throttle_secs=120 \
# --evaluation_mode="reconstruction" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300
# -
# ### Train error distribution variables
# + language="bash"
# OUTDIR=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/trained_model
# JOBNAME=job_lstm_encoder_decoder_autoencoder_anomaly_detection_calculate_error_distribution_statistics_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/validation_normal_1_sequences.csv \
# --eval_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/validation_normal_1_sequences.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2200 \
# --evaluation_mode="calculate_error_distribution_statistics" \
# --eps="1e-12" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300
# -
# ### Tune anomaly thresholds
# + language="bash"
# OUTDIR=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/trained_model
# JOBNAME=job_lstm_encoder_decoder_autoencoder_anomaly_detection_tune_anomaly_thresholds_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=$PWD/lstm_encoder_decoder_autoencoder_anomaly_detection_module/trainer \
# --job-dir=$OUTDIR \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=STANDARD_1 \
# --runtime-version=1.13 \
# -- \
# --train_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/labeled_validation_mixed_sequences.csv \
# --eval_file_pattern=gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/data/labeled_validation_mixed_sequences.csv \
# --output_dir=$OUTDIR \
# --job-dir=$OUTDIR \
# --seq_len=30 \
# --horizon=0 \
# --reverse_labels_sequence=True \
# --encoder_lstm_hidden_units="64 32 16" \
# --decoder_lstm_hidden_units="16 32 64" \
# --lstm_dropout_output_keep_probs="0.9 0.95 1.0" \
# --dnn_hidden_units="1024 256 64" \
# --train_batch_size=32 \
# --eval_batch_size=32 \
# --train_steps=2400 \
# --evaluation_mode="tune_anomaly_thresholds" \
# --num_time_anomaly_thresholds=300 \
# --num_features_anomaly_thresholds=300 \
# --min_time_anomaly_threshold=2.0 \
# --max_time_anomaly_threshold=15.0 \
# --min_features_anomaly_threshold=20 \
# --max_features_anomaly_threshold=60 \
# --f_score_beta=0.05
# -
# # Deploy
# + language="bash"
# MODEL_NAME="lstm_autoencoder_anomaly_detection"
# MODEL_VERSION="v1"
# MODEL_LOCATION=$(gsutil ls gs://$BUCKET/lstm_encoder_decoder_autoencoder_anomaly_detection/trained_model/export/exporter/ | tail -1)
# echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION ... this will take a few minutes"
# #gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
# #gcloud ml-engine models delete ${MODEL_NAME}
# gcloud ml-engine models create $MODEL_NAME --regions $REGION
# gcloud ml-engine versions create $MODEL_VERSION --model $MODEL_NAME --origin $MODEL_LOCATION --runtime-version 1.13
# -
# # Prediction
UNLABELED_CSV_COLUMNS = ["tag_{0}".format(tag) for tag in range(0, 5)]
import numpy as np
labeled_test_mixed_sequences_array = np.loadtxt(fname = "data/labeled_test_mixed_sequences.csv", dtype = str, delimiter = ";")
print("labeled_test_mixed_sequences_array.shape = {}".format(labeled_test_mixed_sequences_array.shape))
number_of_prediction_instances = 10
print("labels = {}".format(labeled_test_mixed_sequences_array[0:number_of_prediction_instances, -1]))
# ### Local prediction from local model
with open('test_sequences.json', 'w') as outfile:
test_data_normal_string_list = labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]
json_string = ""
for example in test_data_normal_string_list:
json_string += "{" + ','.join(["{0}: \"{1}\"".format('\"' + UNLABELED_CSV_COLUMNS[i] + '\"', example[i])
for i in range(len(UNLABELED_CSV_COLUMNS))]) + "}\n"
json_string = json_string.replace(' ', '').replace(':', ': ').replace(',', ', ')
print(json_string)
outfile.write("%s" % json_string)
# + language="bash"
# model_dir=$(ls ${PWD}/trained_model/export/exporter | tail -1)
# gcloud ml-engine local predict \
# --model-dir=${PWD}/trained_model/export/exporter/${model_dir} \
# --json-instances=./test_sequences.json
# -
# ### GCloud ML-Engine prediction from deployed model
test_data_normal_string_list = labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]
# Format dataframe to instances list to get sent to ML-Engine
instances = [{UNLABELED_CSV_COLUMNS[i]: example[i]
for i in range(len(UNLABELED_CSV_COLUMNS))}
for example in labeled_test_mixed_sequences_array.tolist()[0:number_of_prediction_instances]]
instances
# +
# Send instance dictionary to receive response from ML-Engine for online prediction
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials = credentials)
request_data = {"instances": instances}
parent = 'projects/%s/models/%s/versions/%s' % (PROJECT, 'lstm_autoencoder_anomaly_detection', 'v1')
response = api.projects().predict(body = request_data, name = parent).execute()
print("response = {}".format(response))
# -
| courses/machine_learning/asl/open_project/time_series_anomaly_detection/tf_lstm_encoder_decoder_autoencoder/lstm_encoder_decoder_autoencoder_anomaly_detection_gcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/3_image_processing_deep_learning_roadmap/3_deep_learning_advanced/1_Blocks%20in%20Deep%20Learning%20Networks/9)%20Resnext%20Block%20(Type%20-%201).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Goals
#
# ### 1. Learn to implement Resnext Block (Type - 1) using monk
# - Monk's Keras
# - Monk's Pytorch
# - Monk's Mxnet
#
# ### 2. Use network Monk's debugger to create complex blocks
#
#
# ### 3. Understand how syntactically different it is to implement the same using
# - Traditional Keras
# - Traditional Pytorch
# - Traditional Mxnet
# # Resnext Block - Type 1
#
# - Note: The block structure can have variations too, this is just an example
from IPython.display import Image
Image(filename='imgs/resnext_with_downsample.png')
# # Table of contents
#
# [1. Install Monk](#1)
#
#
# [2. Block basic Information](#2)
#
# - [2.1) Visual structure](#2-1)
#
# - [2.2) Layers in Branches](#2-2)
#
#
# [3) Creating Block using monk visual debugger](#3)
#
# - [3.1) Create the first branch](#3-1)
#
# - [3.2) Create the second branch](#3-2)
#
# - [3.3) Merge the branches](#3-3)
#
# - [3.4) Debug the merged network](#3-4)
#
# - [3.5) Compile the network](#3-5)
#
# - [3.6) Visualize the network](#3-6)
#
# - [3.7) Run data through the network](#3-7)
#
#
# [4) Creating Block Using MONK one line API call](#4)
#
# - [Mxnet Backend](#4-1)
#
# - [Pytorch Backend](#4-2)
#
# - [Keras Backend](#4-3)
#
#
#
# [5) Appendix](#5)
#
# - [Study Material](#5-1)
#
# - [Creating block using traditional Mxnet](#5-2)
#
# - [Creating block using traditional Pytorch](#5-3)
#
# - [Creating block using traditional Keras](#5-4)
#
# <a id='0'></a>
# # Install Monk
# ## Using pip (Recommended)
#
# - colab (gpu)
# - All bakcends: `pip install -U monk-colab`
#
#
# - kaggle (gpu)
# - All backends: `pip install -U monk-kaggle`
#
#
# - cuda 10.2
# - All backends: `pip install -U monk-cuda102`
# - Gluon bakcned: `pip install -U monk-gluon-cuda102`
# - Pytorch backend: `pip install -U monk-pytorch-cuda102`
# - Keras backend: `pip install -U monk-keras-cuda102`
#
#
# - cuda 10.1
# - All backend: `pip install -U monk-cuda101`
# - Gluon bakcned: `pip install -U monk-gluon-cuda101`
# - Pytorch backend: `pip install -U monk-pytorch-cuda101`
# - Keras backend: `pip install -U monk-keras-cuda101`
#
#
# - cuda 10.0
# - All backend: `pip install -U monk-cuda100`
# - Gluon bakcned: `pip install -U monk-gluon-cuda100`
# - Pytorch backend: `pip install -U monk-pytorch-cuda100`
# - Keras backend: `pip install -U monk-keras-cuda100`
#
#
# - cuda 9.2
# - All backend: `pip install -U monk-cuda92`
# - Gluon bakcned: `pip install -U monk-gluon-cuda92`
# - Pytorch backend: `pip install -U monk-pytorch-cuda92`
# - Keras backend: `pip install -U monk-keras-cuda92`
#
#
# - cuda 9.0
# - All backend: `pip install -U monk-cuda90`
# - Gluon bakcned: `pip install -U monk-gluon-cuda90`
# - Pytorch backend: `pip install -U monk-pytorch-cuda90`
# - Keras backend: `pip install -U monk-keras-cuda90`
#
#
# - cpu
# - All backend: `pip install -U monk-cpu`
# - Gluon bakcned: `pip install -U monk-gluon-cpu`
# - Pytorch backend: `pip install -U monk-pytorch-cpu`
# - Keras backend: `pip install -U monk-keras-cpu`
# ## Install Monk Manually (Not recommended)
#
# ### Step 1: Clone the library
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
#
#
#
# ### Step 2: Install requirements
# - Linux
# - Cuda 9.0
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu90.txt`
# - Cuda 9.2
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu92.txt`
# - Cuda 10.0
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu100.txt`
# - Cuda 10.1
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu101.txt`
# - Cuda 10.2
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu102.txt`
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Linux && pip install -r requirements_cpu.txt`
#
#
# - Windows
# - Cuda 9.0 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu90.txt`
# - Cuda 9.2 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu92.txt`
# - Cuda 10.0 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu100.txt`
# - Cuda 10.1 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu101.txt`
# - Cuda 10.2 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu102.txt`
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cpu.txt`
#
#
# - Mac
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Mac && pip install -r requirements_cpu.txt`
#
#
# - Misc
# - Colab (GPU)
# - `cd monk_v1/installation/Misc && pip install -r requirements_colab.txt`
# - Kaggle (GPU)
# - `cd monk_v1/installation/Misc && pip install -r requirements_kaggle.txt`
#
#
#
# ### Step 3: Add to system path (Required for every terminal or kernel run)
# - `import sys`
# - `sys.path.append("monk_v1/");`
# # Imports
# Common
import numpy as np
import math
import netron
from collections import OrderedDict
from functools import partial
# +
#Using mxnet-gluon backend
# When installed using pip
from monk.gluon_prototype import prototype
# When installed manually (Uncomment the following)
#import os
#import sys
#sys.path.append("monk_v1/");
#sys.path.append("monk_v1/monk/");
#from monk.gluon_prototype import prototype
# -
# <a id='2'></a>
# # Block Information
# <a id='2_1'></a>
# ## Visual structure
from IPython.display import Image
Image(filename='imgs/resnext_with_downsample.png')
# <a id='2_2'></a>
# ## Layers in Branches
#
# - Number of branches: 2
#
#
# - Branch 1
# - conv_1x1 -> batchnorm
#
#
# - Branch 2
# - conv_1x1 -> batchnorm -> relu -> conv_3x3 -> batchnorm -> relu -> conv_1x1 -> batchnorm
#
#
# - Branches merged using
# - Elementwise addition
#
#
# (See Appendix to read blogs on resnexts)
# <a id='3'></a>
# # Creating Block using monk debugger
# +
# Imports and setup a project
# To use pytorch backend - replace gluon_prototype with pytorch_prototype
# To use keras backend - replace gluon_prototype with keras_prototype
from monk.gluon_prototype import prototype
# Create a sample project
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
# -
# <a id='3-1'></a>
# ## Create the first branch
def first_branch(output_channels=128, stride=1):
network = [];
network.append(gtf.convolution(output_channels=output_channels, kernel_size=1, stride=stride));
network.append(gtf.batch_normalization());
return network;
# Debug the branch
branch_1 = first_branch(output_channels=128, stride=1)
network = [];
network.append(branch_1);
gtf.debug_custom_model_design(network);
# <a id='3-2'></a>
# ## Create the second branch
def second_branch(output_channels=256, cardinality=8, bottleneck_width=4, stride=1):
network = [];
channels = output_channels//4;
D = int(math.floor(channels * (bottleneck_width / 64)))
group_width = cardinality * D
network.append(gtf.convolution(output_channels=group_width, kernel_size=1, stride=1));
network.append(gtf.batch_normalization());
network.append(gtf.relu());
network.append(gtf.convolution(output_channels=group_width, kernel_size=3, stride=stride));
network.append(gtf.batch_normalization());
network.append(gtf.relu());
network.append(gtf.convolution(output_channels=output_channels, kernel_size=1, stride=1));
network.append(gtf.batch_normalization());
return network;
# Debug the branch
branch_2 = second_branch(output_channels=256, cardinality=8, bottleneck_width=4, stride=1)
network = [];
network.append(branch_2);
gtf.debug_custom_model_design(network);
# <a id='3-3'></a>
# ## Merge the branches
def final_block(output_channels=256, cardinality=8, bottleneck_width=4, stride=1):
network = [];
#Create subnetwork and add branches
subnetwork = [];
branch_1 = first_branch(output_channels=output_channels, stride=stride)
branch_2 = second_branch(output_channels=output_channels,
cardinality=cardinality,
bottleneck_width=bottleneck_width,
stride=stride)
subnetwork.append(branch_1);
subnetwork.append(branch_2);
# Add merging element
subnetwork.append(gtf.add());
# Add the subnetwork
network.append(subnetwork)
network.append(gtf.relu());
return network;
# <a id='3-4'></a>
# ## Debug the merged network
final = final_block(output_channels=256, stride=1)
network = [];
network.append(final);
gtf.debug_custom_model_design(network);
# <a id='3-5'></a>
# ## Compile the network
gtf.Compile_Network(network, data_shape=(3, 224, 224), use_gpu=False);
# <a id='3-6'></a>
# ## Run data through the network
import mxnet as mx
x = np.zeros((1, 3, 224, 224));
x = mx.nd.array(x);
y = gtf.system_dict["local"]["model"].forward(x);
print(x.shape, y.shape)
# <a id='3-7'></a>
# ## Visualize network using netron
gtf.Visualize_With_Netron(data_shape=(3, 224, 224))
# <a id='4'></a>
# # Creating Using MONK LOW code API
# <a id='4-1'></a>
# ## Mxnet backend
# +
from monk.gluon_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.resnext_block(output_channels=256, cardinality=8, bottleneck_width=4,));
gtf.Compile_Network(network, data_shape=(3, 224, 224), use_gpu=False);
# -
# <a id='4-2'></a>
# ## Pytorch backend
#
# - Only the import changes
# +
#Change gluon_prototype to pytorch_prototype
from monk.pytorch_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.resnext_block(output_channels=256, cardinality=8, bottleneck_width=4));
gtf.Compile_Network(network, data_shape=(3, 224, 224), use_gpu=False);
# -
# <a id='4-3'></a>
# ## Keras backend
#
# - Only the import changes
# +
#Change gluon_prototype to keras_prototype
from monk.keras_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.resnext_block(output_channels=256, cardinality=8, bottleneck_width=4,));
gtf.Compile_Network(network, data_shape=(3, 224, 224), use_gpu=False);
# -
# <a id='5'></a>
# # Appendix
# <a id='5-1'></a>
# ## Study links
# - https://towardsdatascience.com/review-resnext-1st-runner-up-of-ilsvrc-2016-image-classification-15d7f17b42ac
# - https://datascience.stackexchange.com/questions/47559/cardinality-vs-width-in-the-resnext-architecture
# - https://arxiv.org/abs/1611.05431
# - https://research.fb.com/downloads/resnext/
# <a id='5-2'></a>
# ## Creating block using traditional Mxnet
#
# - Code credits - https://mxnet.incubator.apache.org/
# Traditional-Mxnet-gluon
import mxnet as mx
from mxnet.gluon import nn
from mxnet.gluon.nn import HybridBlock, BatchNorm
from mxnet.gluon.contrib.nn import HybridConcurrent, Identity
from mxnet import gluon, init, nd
# +
def _conv3x3(channels, stride, in_channels):
return nn.Conv2D(channels, kernel_size=3, strides=stride, padding=1,
use_bias=False, in_channels=in_channels)
class ResnextBlock(HybridBlock):
def __init__(self, output_channels, stride, cardinality=8, bottleneck_width=4, in_channels=0, **kwargs):
super(ResnextBlock, self).__init__(**kwargs)
channels = output_channels//4;
D = int(math.floor(channels * (bottleneck_width / 64)))
group_width = cardinality * D
#Branch - 1
self.downsample = nn.HybridSequential(prefix='')
self.downsample.add(nn.Conv2D(output_channels, kernel_size=1, strides=stride,
use_bias=False, in_channels=in_channels))
self.downsample.add(nn.BatchNorm())
# Branch - 2
self.body = nn.HybridSequential(prefix='')
self.body.add(nn.Conv2D(group_width, kernel_size=1, strides=1,
use_bias=False, in_channels=in_channels))
self.body.add(nn.BatchNorm())
self.body.add(nn.Activation('relu'))
self.body.add(_conv3x3(group_width, stride, in_channels))
self.body.add(nn.BatchNorm())
self.body.add(nn.Activation('relu'))
self.body.add(nn.Conv2D(output_channels, kernel_size=1, strides=1,
use_bias=False, in_channels=in_channels))
self.body.add(nn.BatchNorm())
def hybrid_forward(self, F, x):
residual = x
x = self.body(x)
residual = self.downsample(residual)
x = F.Activation(residual+x, act_type='relu')
return x
# +
# Invoke the block
block = ResnextBlock(256, 1)
# Initialize network and load block on machine
ctx = [mx.cpu()];
block.initialize(init.Xavier(), ctx = ctx);
block.collect_params().reset_ctx(ctx)
block.hybridize()
# Run data through network
x = np.zeros((1, 3, 224, 224));
x = mx.nd.array(x);
y = block.forward(x);
print(x.shape, y.shape)
# Export Model to Load on Netron
block.export("final", epoch=0);
netron.start("final-symbol.json", port=8082)
# -
# <a id='5-3'></a>
# ## Creating block using traditional Pytorch
#
# - Code credits - https://github.com/soeaver/pytorch-priv/blob/master/models/cifar/resnext.py
# Traiditional-Pytorch
import torch
from torch import nn
from torch.jit.annotations import List
import torch.nn.functional as F
class ResNeXtBottleneck(nn.Module):
"""
RexNeXt bottleneck type C (https://github.com/facebookresearch/ResNeXt/blob/master/models/resnext.lua)
"""
def __init__(self, in_channels, out_channels, stride, cardinality, widen_factor):
""" Constructor
Args:
in_channels: input channel dimensionality
out_channels: output channel dimensionality
stride: conv stride. Replaces pooling layer.
cardinality: num of convolution groups.
widen_factor: factor to reduce the input dimensionality before convolution.
"""
super(ResNeXtBottleneck, self).__init__()
D = cardinality * out_channels // widen_factor
self.conv_reduce = nn.Conv2d(in_channels, D, kernel_size=1, stride=1, padding=0, bias=False)
self.bn_reduce = nn.BatchNorm2d(D)
self.conv_conv = nn.Conv2d(D, D, kernel_size=3, stride=stride, padding=1, groups=cardinality, bias=False)
self.bn = nn.BatchNorm2d(D)
self.conv_expand = nn.Conv2d(D, out_channels, kernel_size=1, stride=1, padding=0, bias=False)
self.bn_expand = nn.BatchNorm2d(out_channels)
self.shortcut = nn.Sequential()
if in_channels != out_channels:
self.shortcut.add_module('shortcut_conv', nn.Conv2d(in_channels, out_channels, kernel_size=1, stride=stride,
padding=0, bias=False))
self.shortcut.add_module('shortcut_bn', nn.BatchNorm2d(out_channels))
def forward(self, x):
bottleneck = self.conv_reduce.forward(x)
bottleneck = F.relu(self.bn_reduce.forward(bottleneck), inplace=True)
bottleneck = self.conv_conv.forward(bottleneck)
bottleneck = F.relu(self.bn.forward(bottleneck), inplace=True)
bottleneck = self.conv_expand.forward(bottleneck)
bottleneck = self.bn_expand.forward(bottleneck)
residual = self.shortcut.forward(x)
return F.relu(residual + bottleneck, inplace=True)
# +
# Invoke the block
block = ResNeXtBottleneck(3, 256, 1, 8, 4);
# Initialize network and load block on machine
layers = []
layers.append(block);
net = nn.Sequential(*layers);
# Run data through network
x = torch.randn(1, 3, 224, 224)
y = net(x)
print(x.shape, y.shape);
# Export Model to Load on Netron
torch.onnx.export(net, # model being run
x, # model input (or a tuple for multiple inputs)
"model.onnx", # where to save the model (can be a file or file-like object)
export_params=True, # store the trained parameter weights inside the model file
opset_version=10, # the ONNX version to export the model to
do_constant_folding=True, # whether to execute constant folding for optimization
input_names = ['input'], # the model's input names
output_names = ['output'], # the model's output names
dynamic_axes={'input' : {0 : 'batch_size'}, # variable lenght axes
'output' : {0 : 'batch_size'}})
netron.start('model.onnx', port=9998);
# -
# <a id='5-4'></a>
# ## Creating block using traditional Keras
#
# - Code credits: https://github.com/titu1994/Keras-ResNeXt/blob/master/resnext.py
# Traditional-Keras
import keras
import keras.layers as kla
import keras.models as kmo
import tensorflow as tf
from keras.models import Model
backend = 'channels_last'
from keras import layers
from keras.layers import *
import keras.backend as K
from keras.regularizers import l2
# +
def __initial_conv_block(input, weight_decay=5e-4):
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
x = Conv2D(64, (3, 3), padding='same', use_bias=False, kernel_initializer='he_normal',
kernel_regularizer=l2(weight_decay))(input)
x = BatchNormalization(axis=channel_axis)(x)
x = Activation('relu')(x)
return x
def __initial_conv_block_imagenet(input, weight_decay=5e-4):
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
x = Conv2D(64, (7, 7), padding='same', use_bias=False, kernel_initializer='he_normal',
kernel_regularizer=l2(weight_decay), strides=(2, 2))(input)
x = BatchNormalization(axis=channel_axis)(x)
x = Activation('relu')(x)
x = MaxPooling2D((3, 3), strides=(2, 2), padding='same')(x)
return x
def __grouped_convolution_block(input, grouped_channels, cardinality, strides, weight_decay=5e-4):
init = input
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
group_list = []
if cardinality == 1:
# with cardinality 1, it is a standard convolution
x = Conv2D(grouped_channels, (3, 3), padding='same', use_bias=False, strides=(strides, strides),
kernel_initializer='he_normal', kernel_regularizer=l2(weight_decay))(init)
x = BatchNormalization(axis=channel_axis)(x)
x = Activation('relu')(x)
return x
for c in range(cardinality):
x = Lambda(lambda z: z[:, :, :, c * grouped_channels:(c + 1) * grouped_channels]
if K.image_data_format() == 'channels_last' else
lambda z: z[:, c * grouped_channels:(c + 1) * grouped_channels, :, :])(input)
x = Conv2D(grouped_channels, (3, 3), padding='same', use_bias=False, strides=(strides, strides),
kernel_initializer='he_normal', kernel_regularizer=l2(weight_decay))(x)
group_list.append(x)
group_merge = concatenate(group_list, axis=channel_axis)
x = BatchNormalization(axis=channel_axis)(group_merge)
x = Activation('relu')(x)
return x
def bottleneck_block(input, filters=64, cardinality=8, strides=1, weight_decay=5e-4):
init = input
grouped_channels = int(filters / cardinality)
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
# Check if input number of filters is same as 16 * k, else create convolution2d for this input
if K.image_data_format() == 'channels_first':
if init._keras_shape[1] != 2 * filters:
init = Conv2D(filters * 2, (1, 1), padding='same', strides=(strides, strides),
use_bias=False, kernel_initializer='he_normal', kernel_regularizer=l2(weight_decay))(init)
init = BatchNormalization(axis=channel_axis)(init)
else:
if init._keras_shape[-1] != 2 * filters:
init = Conv2D(filters * 2, (1, 1), padding='same', strides=(strides, strides),
use_bias=False, kernel_initializer='he_normal', kernel_regularizer=l2(weight_decay))(init)
init = BatchNormalization(axis=channel_axis)(init)
x = Conv2D(filters, (1, 1), padding='same', use_bias=False,
kernel_initializer='he_normal', kernel_regularizer=l2(weight_decay))(input)
x = BatchNormalization(axis=channel_axis)(x)
x = Activation('relu')(x)
x = __grouped_convolution_block(x, grouped_channels, cardinality, strides, weight_decay)
x = Conv2D(filters * 2, (1, 1), padding='same', use_bias=False, kernel_initializer='he_normal',
kernel_regularizer=l2(weight_decay))(x)
x = BatchNormalization(axis=channel_axis)(x)
x = add([init, x])
x = Activation('relu')(x)
return x
def create_model(input_shape, filters=64, cardinality=8, strides=1):
img_input = layers.Input(shape=input_shape);
x = bottleneck_block(img_input, filters=filters,
cardinality=cardinality,
strides=strides)
return Model(img_input, x);
# +
# Invoke the block
filters=256;
input_shape=(224, 224, 3);
model = create_model(input_shape, filters=filters, cardinality=8, strides=1);
# Run data through network
x = tf.placeholder(tf.float32, shape=(1, 224, 224, 3))
y = model(x)
print(x.shape, y.shape)
# Export Model to Load on Netron
model.save("final.h5");
netron.start("final.h5", port=8082)
# -
# # Goals Completed
#
# ### 1. Learn to implement Resnext Block (Type - 1) using monk
# - Monk's Keras
# - Monk's Pytorch
# - Monk's Mxnet
#
# ### 2. Use network Monk's debugger to create complex blocks
#
#
# ### 3. Understand how syntactically different it is to implement the same using
# - Traditional Keras
# - Traditional Pytorch
# - Traditional Mxnet
| study_roadmaps/3_image_processing_deep_learning_roadmap/3_deep_learning_advanced/1_Blocks in Deep Learning Networks/9) Resnext Block (Type - 1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Twitter data mining using Python assignment 14
# *Team Rython: <NAME> and <NAME>*
# *Date: 21st of January, 2016*
# *Apache License 2.0*
# ## Imports
# Make sure you have pysqlite2, tweepy and spatialite installed!
from __future__ import division
import tweepy
import datetime
import json
import os
from pysqlite2 import dbapi2 as sqlite3
# ## Twitter authentication (fill this!)
APP_KEY = ""
APP_SECRET = ""
OAUTH_TOKEN = ""
OAUTH_TOKEN_SECRET = ""
# Using Tweepy instead of Twython (because it's more readily available via apt-get or zypper).
auth = tweepy.OAuthHandler(APP_KEY, APP_SECRET)
auth.set_access_token(OAUTH_TOKEN, OAUTH_TOKEN_SECRET)
api = tweepy.API(auth)
# ## Database file to write to (fill this!)
# The database file has to exist and already have a table defined. An emty ready database file is included in "spatial-backup.sqlite", so you can use that.
databasefile = "spatial-backup.sqlite"
# ## SQLite opening
# **Make sure you load pysqlite2, and give a path for mod_spatialite for SpatiaLite support! This may be distribution-specific! You might need to install libspatialite!**
conn = sqlite3.connect(databasefile)
conn.enable_load_extension(True)
conn.execute('SELECT load_extension("/usr/lib64/mod_spatialite.so.7")')
curs = conn.cursor()
# ## Coordinates to WKT
# Converts Twitter coordinates (two points) into a Well Known Text.
def coordinates_to_wkt(coords):
if coords == None:
return ""
return "POINT("+str(coords["coordinates"][0])+" "+str(coords["coordinates"][1])+")"
# ## Bounding box to WKT
# Calculates the centroid of a bounding box and returns a Well Known Text of that point. Only polygonal bounding boxes are supported (but are there any other kind?)
def bbox_to_wkt(bbox):
if bbox.coordinates == None:
return ""
if bbox.type == "Polygon":
centroid = [0, 0]
centroid[0] = (bbox.coordinates[0][2][0] + bbox.coordinates[0][0][0]) / 2
centroid[1] = (bbox.coordinates[0][2][1] + bbox.coordinates[0][0][1]) / 2
return "POINT("+str(centroid[0])+" "+str(centroid[1])+")"
print "Unknown place type!"
return ""
# ## Process query: main function of the script
# Does the query parsing and output to SpatiaLite. Pass the result of `api.search()` to it.
def process_query(search_results):
for tweet in search_results:
full_place_name = ""
place_type = ""
location = ""
username = tweet.user.screen_name
followers_count = tweet.user.followers_count
tweettext = tweet.text.encode("utf-8")
if tweet.place != None:
full_place_name = tweet.place.full_name
place_type = tweet.place.place_type
coordinates = tweet.coordinates
if (coordinates != None) or (tweet.place != None):
print 'Found a geolocated tweet! By:'
print username
print '==========================='
if coordinates != None:
location = coordinates_to_wkt(coordinates)
else:
if tweet.place != None:
location = bbox_to_wkt(tweet.place.bounding_box)
curs.execute("insert into tweets (username, followers_count, tweettext, full_place_name, place_type, coordinates, geometry) values (?, ?, ?, ?, ?, ?, ST_GeomFromText( ? , 4326));", \
(username, followers_count, tweettext.decode('utf-8'), full_place_name, place_type, location, location))
conn.commit()
# # Example queries
# Write queries in succession (or loops if you like). Their results (if they are geolocated) will be added into the SpatiaLite database.
process_query(api.search(q="Beer", count=100))
process_query(api.search(q="Jorn", count=100))
process_query(api.search(q="cairo", count=100))
process_query(api.search(q="washington", count=100))
# ## Close the database
conn.close()
# ## Visualise data
# Opens QGIS with the database passed as an argument. It should show you all the points. Add a layer of OpenStreetMap or such for a nice visualisation of the points.
os.system("qgis "+databasefile)
# 
| Lesson14/Twitter assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generalized Least Squares
import statsmodels.api as sm
# The Longley dataset is a time series dataset:
data = sm.datasets.longley.load(as_pandas=False)
data.exog = sm.add_constant(data.exog)
print(data.exog[:5])
#
# Let's assume that the data is heteroskedastic and that we know
# the nature of the heteroskedasticity. We can then define
# `sigma` and use it to give us a GLS model
#
# First we will obtain the residuals from an OLS fit
ols_resid = sm.OLS(data.endog, data.exog).fit().resid
# Assume that the error terms follow an AR(1) process with a trend:
#
# $\epsilon_i = \beta_0 + \rho\epsilon_{i-1} + \eta_i$
#
# where $\eta \sim N(0,\Sigma^2)$
#
# and that $\rho$ is simply the correlation of the residual a consistent estimator for rho is to regress the residuals on the lagged residuals
resid_fit = sm.OLS(ols_resid[1:], sm.add_constant(ols_resid[:-1])).fit()
print(resid_fit.tvalues[1])
print(resid_fit.pvalues[1])
# While we do not have strong evidence that the errors follow an AR(1)
# process we continue
rho = resid_fit.params[1]
# As we know, an AR(1) process means that near-neighbors have a stronger
# relation so we can give this structure by using a toeplitz matrix
# +
from scipy.linalg import toeplitz
toeplitz(range(5))
# -
order = toeplitz(range(len(ols_resid)))
# so that our error covariance structure is actually rho**order
# which defines an autocorrelation structure
sigma = rho**order
gls_model = sm.GLS(data.endog, data.exog, sigma=sigma)
gls_results = gls_model.fit()
# Of course, the exact rho in this instance is not known so it it might make more sense to use feasible gls, which currently only has experimental support.
#
# We can use the GLSAR model with one lag, to get to a similar result:
glsar_model = sm.GLSAR(data.endog, data.exog, 1)
glsar_results = glsar_model.iterative_fit(1)
print(glsar_results.summary())
# Comparing gls and glsar results, we see that there are some small
# differences in the parameter estimates and the resulting standard
# errors of the parameter estimate. This might be do to the numerical
# differences in the algorithm, e.g. the treatment of initial conditions,
# because of the small number of observations in the longley dataset.
print(gls_results.params)
print(glsar_results.params)
print(gls_results.bse)
print(glsar_results.bse)
| docs/source2/examples/notebooks/generated/gls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# metadata:
# interpreter:
# hash: aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# name: python3
# ---
# +
# # !pip install --quiet climetlab
# -
# # External plugins
# Install the demo external dataset. See https://github.com/ecmwf/climetlab-demo-dataset for more information.
# !pip install --quiet climetlab-demo-dataset
import climetlab as cml
cml.__file__
# Get the demo dataset:
# + tags=[]
ds = cml.load_dataset("demo-dataset")
# -
# Plot it:
cml.plot_map(ds)
# !pip install --quiet climetlab-demo-source
# !test -f test.db || wget https://github.com/ecmwf/climetlab/raw/master/docs/examples/test.db
s = cml.load_source("demo-source", "sqlite:///test.db", "select * from data;", parse_dates=["time"])
df = s.to_pandas()
df
cml.plot_map(df, margins=2)
| docs/examples/12-external-plugins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import re
import bs4
import time
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.desired_capabilities import DesiredCapabilities
from fake_useragent import UserAgent
import EasyWebdriver
from scipy.special import comb
res = requests.get("https://pvpoke.com/team-builder/all/2500/mew-m-8-18-0%2Csnorlax-m-0-4-0%2Cswampert-m-0-1-3")
time.sleep(2)
res.raise_for_status()
soup = bs4.BeautifulSoup(res.text,'lxml')
soup.select('b[class="threat-score"]')[0]
def get_phantomjs():
ua = UserAgent()
caps = DesiredCapabilities.PHANTOMJS
caps["phantomjs.page.settings.userAgent"] = ua.random
browser = webdriver.PhantomJS(desired_capabilities=caps)
return browser
def get_chrome():
return EasyWebdriver.Chrome()
pjs = get_phantomjs()
pjs.get("https://pvpoke.com/team-builder/all/2500/mew-m-8-18-0%2Csnorlax-m-0-4-0%2Cswampert-m-0-1-3")
chrome = get_chrome()
chrome.get("https://pvpoke.com/team-builder/all/2500/mew-m-8-18-0%2Csnorlax-m-0-4-0%2Cswampert-m-0-1-3")
chrome.find_element_by_class_name("threat-score").text
def get_threat_score(browser, pvpoke_url, maxwait = 10):
browser.get(pvpoke_url)
threat_score = ""
for i in range(0, maxwait):
threat_score = browser.find_element_by_class_name("threat-score").text
if threat_score != "":
return int(threat_score)
time.sleep(1)
return None
get_threat_score(chrome, "https://pvpoke.com/team-builder/all/2500/mew-m-8-18-0%2Csnorlax-m-0-4-0%2Cswampert-m-0-1-3")
comb(4,3)
comb(10,3)
| initial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
# read data
original_data = pd.read_csv("puff-episode.csv")
original_data = original_data.drop(['offset'], axis=1)
backup_data = pd.read_csv("puff-episode-backup.csv")
original_data = original_data.dropna()
backup_data = backup_data.dropna()
# -
# check data types
print(original_data.dtypes)
print("")
print(backup_data.dtypes)
# +
# convert data types to be consistent
original_data['participant_id'] = original_data['participant_id'].astype(float)
original_data['event'] = original_data['event'].astype(float)
original_data['hour'] = original_data['hour'].astype(float)
original_data['minute'] = original_data['minute'].astype(float)
backup_data['event'] = backup_data['event'].astype(float)
backup_data['participant_id'] = backup_data['participant_id'].astype(float)
backup_data['hour'] = backup_data['hour'].astype(float)
backup_data['minute'] = backup_data['minute'].astype(float)
print(original_data.dtypes)
print(backup_data.dtypes)
# -
# summary of phone data
print("SMOKING_EPISODE Phone Data Summary:")
print(original_data.describe())
# summary of cloud data
print("PUFF_PROBABILITY Cloud Data Summary:")
print(backup_data.describe())
original_data.shape
backup_data.shape
# count rows per participant
original_data['participant_id'].value_counts().sort_index()
backup_data['participant_id'].value_counts().sort_index()
# now take out participants that do not have backup data
original_data_backup = original_data[~original_data.participant_id.isin([201,203,206,210,221,229])]
original_data_backup['participant_id'].value_counts().sort_index()
# +
s1 = set() # cloud data only
s2 = set() # phone data only
d = {}
lst2 = []
lst1 = []
for index, row in backup_data.iterrows():
participant_id = row['participant_id']
hour = row['hour']
minute = row['minute']
day_of_week = row['day_of_week']
valid_key = (participant_id, hour, minute, day_of_week)
s1.add(valid_key)
d[valid_key] = index
for index, row in original_data.iterrows():
participant_id = row['participant_id']
hour = row['hour']
minute = row['minute']
day_of_week = row['day_of_week']
valid_key = (participant_id, hour, minute, day_of_week)
if valid_key in s1:
s1.remove(valid_key)
else:
# print(index)
lst2.append(index)
lst1 = []
for i in s1:
lst1.append(d[i])
lst1.sort()
difference = backup_data.ix[lst1].dropna()
print(difference.shape)
difference # rows in the backup but not in the original
# -
difference = original_data.ix[lst2]
print(difference.shape)
difference # rows in the original but not in the backup
| data_preprocessing/data_source_comparison/.ipynb_checkpoints/PUFF_EPISODE_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# Source localization with MNE/dSPM/sLORETA/eLORETA
# =================================================
#
# The aim of this tutorial is to teach you how to compute and apply a linear
# inverse method such as MNE/dSPM/sLORETA/eLORETA on evoked/raw/epochs data.
#
# +
# sphinx_gallery_thumbnail_number = 10
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.datasets import sample
from mne.minimum_norm import make_inverse_operator, apply_inverse
# -
# Process MEG data
#
#
# +
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw.fif'
raw = mne.io.read_raw_fif(raw_fname) # already has an average reference
events = mne.find_events(raw, stim_channel='STI 014')
event_id = dict(aud_l=1) # event trigger and conditions
tmin = -0.2 # start of each epoch (200ms before the trigger)
tmax = 0.5 # end of each epoch (500ms after the trigger)
raw.info['bads'] = ['MEG 2443', 'EEG 053']
baseline = (None, 0) # means from the first instant to t = 0
reject = dict(grad=4000e-13, mag=4e-12, eog=150e-6)
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, proj=True,
picks=('meg', 'eog'), baseline=baseline, reject=reject)
# -
# Compute regularized noise covariance
# ------------------------------------
#
# For more details see `tut_compute_covariance`.
#
#
# +
noise_cov = mne.compute_covariance(
epochs, tmax=0., method=['shrunk', 'empirical'], rank=None, verbose=True)
fig_cov, fig_spectra = mne.viz.plot_cov(noise_cov, raw.info)
# -
# Compute the evoked response
# ---------------------------
# Let's just use MEG channels for simplicity.
#
#
# +
evoked = epochs.average().pick('meg')
evoked.plot(time_unit='s')
evoked.plot_topomap(times=np.linspace(0.05, 0.15, 5), ch_type='mag',
time_unit='s')
# Show whitening
evoked.plot_white(noise_cov, time_unit='s')
del epochs # to save memory
# -
# Inverse modeling: MNE/dSPM on evoked and raw data
# -------------------------------------------------
#
#
# +
# Read the forward solution and compute the inverse operator
fname_fwd = data_path + '/MEG/sample/sample_audvis-meg-oct-6-fwd.fif'
fwd = mne.read_forward_solution(fname_fwd)
# make an MEG inverse operator
info = evoked.info
inverse_operator = make_inverse_operator(info, fwd, noise_cov,
loose=0.2, depth=0.8)
del fwd
# You can write it to disk with::
#
# >>> from mne.minimum_norm import write_inverse_operator
# >>> write_inverse_operator('sample_audvis-meg-oct-6-inv.fif',
# inverse_operator)
# -
# Compute inverse solution
# ------------------------
#
#
method = "dSPM"
snr = 3.
lambda2 = 1. / snr ** 2
stc, residual = apply_inverse(evoked, inverse_operator, lambda2,
method=method, pick_ori=None,
return_residual=True, verbose=True)
# Visualization
# -------------
# View activation time-series
#
#
plt.figure()
plt.plot(1e3 * stc.times, stc.data[::100, :].T)
plt.xlabel('time (ms)')
plt.ylabel('%s value' % method)
plt.show()
# Examine the original data and the residual after fitting:
#
#
fig, axes = plt.subplots(2, 1)
evoked.plot(axes=axes)
for ax in axes:
ax.texts = []
for line in ax.lines:
line.set_color('#98df81')
residual.plot(axes=axes)
# Here we use peak getter to move visualization to the time point of the peak
# and draw a marker at the maximum peak vertex.
#
#
# +
vertno_max, time_max = stc.get_peak(hemi='rh')
subjects_dir = data_path + '/subjects'
surfer_kwargs = dict(
hemi='rh', subjects_dir=subjects_dir,
clim=dict(kind='value', lims=[8, 12, 15]), views='lateral',
initial_time=time_max, time_unit='s', size=(800, 800), smoothing_steps=5)
brain = stc.plot(**surfer_kwargs)
brain.add_foci(vertno_max, coords_as_verts=True, hemi='rh', color='blue',
scale_factor=0.6, alpha=0.5)
brain.add_text(0.1, 0.9, 'dSPM (plus location of maximal activation)', 'title',
font_size=14)
# -
# Morph data to average brain
# ---------------------------
#
#
# +
# setup source morph
morph = mne.compute_source_morph(
src=inverse_operator['src'], subject_from=stc.subject,
subject_to='fsaverage', spacing=5, # to ico-5
subjects_dir=subjects_dir)
# morph data
stc_fsaverage = morph.apply(stc)
brain = stc_fsaverage.plot(**surfer_kwargs)
brain.add_text(0.1, 0.9, 'Morphed to fsaverage', 'title', font_size=20)
del stc_fsaverage
# -
# Dipole orientations
# -------------------
# The ``pick_ori`` parameter of the
# :func:`mne.minimum_norm.apply_inverse` function controls
# the orientation of the dipoles. One useful setting is ``pick_ori='vector'``,
# which will return an estimate that does not only contain the source power at
# each dipole, but also the orientation of the dipoles.
#
#
stc_vec = apply_inverse(evoked, inverse_operator, lambda2,
method=method, pick_ori='vector')
brain = stc_vec.plot(**surfer_kwargs)
brain.add_text(0.1, 0.9, 'Vector solution', 'title', font_size=20)
del stc_vec
# Note that there is a relationship between the orientation of the dipoles and
# the surface of the cortex. For this reason, we do not use an inflated
# cortical surface for visualization, but the original surface used to define
# the source space.
#
# For more information about dipole orientations, see
# `tut-dipole-orientations`.
#
#
# Now let's look at each solver:
#
#
for mi, (method, lims) in enumerate((('dSPM', [8, 12, 15]),
('sLORETA', [3, 5, 7]),
('eLORETA', [0.75, 1.25, 1.75]),)):
surfer_kwargs['clim']['lims'] = lims
stc = apply_inverse(evoked, inverse_operator, lambda2,
method=method, pick_ori=None)
brain = stc.plot(figure=mi, **surfer_kwargs)
brain.add_text(0.1, 0.9, method, 'title', font_size=20)
del stc
| stable/_downloads/597e5dfe47820d56eb232e93bcc62128/plot_mne_dspm_source_localization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
#
# _You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-machine-learning/resources/bANLa) course resource._
#
# ---
# ## Applied Machine Learning, Module 1: A simple classification task
# ### Import required modules and load data file
# +
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
fruits = pd.read_table('readonly/fruit_data_with_colors.txt')
# -
fruits.head()
# create a mapping from fruit label value to fruit name to make results easier to interpret
lookup_fruit_name = dict(zip(fruits.fruit_label.unique(), fruits.fruit_name.unique()))
lookup_fruit_name
# The file contains the mass, height, and width of a selection of oranges, lemons and apples. The heights were measured along the core of the fruit. The widths were the widest width perpendicular to the height.
# ### Examining the data
# +
# plotting a scatter matrix
from matplotlib import cm
X = fruits[['height', 'width', 'mass', 'color_score']]
y = fruits['fruit_label']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
cmap = cm.get_cmap('gnuplot')
scatter = pd.scatter_matrix(X_train, c= y_train, marker = 'o', s=40, hist_kwds={'bins':15}, figsize=(9,9), cmap=cmap)
# +
# plotting a 3D scatter plot
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection = '3d')
ax.scatter(X_train['width'], X_train['height'], X_train['color_score'], c = y_train, marker = 'o', s=100)
ax.set_xlabel('width')
ax.set_ylabel('height')
ax.set_zlabel('color_score')
plt.show()
# -
# ### Create train-test split
# +
# For this example, we use the mass, width, and height features of each fruit instance
X = fruits[['mass', 'width', 'height']]
y = fruits['fruit_label']
# default is 75% / 25% train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# -
# ### Create classifier object
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors = 5)
# -
# ### Train the classifier (fit the estimator) using the training data
knn.fit(X_train, y_train)
# ### Estimate the accuracy of the classifier on future data, using the test data
knn.score(X_test, y_test)
# ### Use the trained k-NN classifier model to classify new, previously unseen objects
# first example: a small fruit with mass 20g, width 4.3 cm, height 5.5 cm
fruit_prediction = knn.predict([[20, 4.3, 5.5]])
lookup_fruit_name[fruit_prediction[0]]
# second example: a larger, elongated fruit with mass 100g, width 6.3 cm, height 8.5 cm
fruit_prediction = knn.predict([[100, 6.3, 8.5]])
lookup_fruit_name[fruit_prediction[0]]
# ### Plot the decision boundaries of the k-NN classifier
# +
from adspy_shared_utilities import plot_fruit_knn
plot_fruit_knn(X_train, y_train, 5, 'uniform') # we choose 5 nearest neighbors
# -
# ### How sensitive is k-NN classification accuracy to the choice of the 'k' parameter?
# +
k_range = range(1,20)
scores = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors = k)
knn.fit(X_train, y_train)
scores.append(knn.score(X_test, y_test))
plt.figure()
plt.xlabel('k')
plt.ylabel('accuracy')
plt.scatter(k_range, scores)
plt.xticks([0,5,10,15,20]);
# -
# ### How sensitive is k-NN classification accuracy to the train/test split proportion?
# +
t = [0.8, 0.7, 0.6, 0.5, 0.4, 0.3, 0.2]
knn = KNeighborsClassifier(n_neighbors = 5)
plt.figure()
for s in t:
scores = []
for i in range(1,1000):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 1-s)
knn.fit(X_train, y_train)
scores.append(knn.score(X_test, y_test))
plt.plot(s, np.mean(scores), 'bo')
plt.xlabel('Training set proportion (%)')
plt.ylabel('accuracy');
# -
| Applied Machine Learning in Python/Module+1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Wikigap 2021 report ([source code](https://github.com/wmcz/voila-notebooks/blob/master/wikigap-2021-report.ipynb))
#
# ### License statement
# Copyright 2022 <NAME> (<EMAIL>)
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# +
from IPython.display import display, Markdown, Latex, HTML
import os
import requests
from datetime import datetime
import wmpaws
import pandas as pd
pd.set_option('display.max_rows', None)
# -
# !curl -s 'https://cs.wikipedia.org/w/index.php?title=Wikipedie:WikiGap_2021/Seznam_vyznamenan%C3%BDch&oldid=19752375&action=raw' | grep -o 'user=[-a-zA-Z0-9 ()]*' | cut -d= -f 2 > ~/data/wmcz-wikigap-2021-users.txt
users = open(os.path.expanduser('~/data/wmcz-wikigap-2021-users.txt')).read().split('\n')
users.pop()
usersDf = pd.DataFrame({
'user_name': users
})
usersDf.set_index('user_name', inplace=True)
# ## Účastníci Wikigapu podle data registrace
#
# V této sekci reportu jsou zobrazeni účastníci Wikigapu podle jejich zkušeností na Wikipedii. Za nováčka jsou považováni uživatelé, kteří svůj účet vytvořili po 1. březnu 2022.
tenureDf = wmpaws.run_sql('''
SELECT
user_name,
IF(user_registration > '20210308000000', 'Newcomer', 'Experienced') AS user_role
FROM user
WHERE
user_name IN ({users})
AND user_registration IS NOT NULL
'''.format(
users=', '.join(["'%s'" % x for x in users])
), 'cswiki')
tenureAggDf = tenureDf.groupby('user_role').count().rename(columns={'user_name': 'count'})
tenureAggDf.plot.pie(y='count', figsize=(8,8), title='Účastníci Wikigapu podle wiki-zkušeností');
# ### Data
#
# #### Agregováno
tenureAggDf
# #### Seznam nováčků
tenureDf.loc[tenureDf.user_role == 'Newcomer']
# ## Účastníci Wikigapu podle pohlaví
#
# Pohlaví je použito podle informace uložené v [uživatelských nastaveních na české Wikipedii](https://cs.wikipedia.org/wiki/Special:Preferences). Výrazná většina uživatelů tento parametr nemá vyplněný – toto je třeba při vnímání reportu mít na paměti.
gendersDf = wmpaws.run_sql('''
SELECT user_name, up_value AS gender
FROM user_properties
JOIN user ON up_user=user_id
WHERE
up_property='gender'
AND user_name IN ({users})
'''.format(
users=', '.join(["'%s'" % x for x in users])
), 'cswiki')
gendersDf.set_index('user_name', inplace=True)
gendersDf = usersDf.join(gendersDf).reset_index().fillna('unknown')
gendersAggDf = gendersDf.reset_index(drop=True).groupby('gender').count().rename(columns={'user_name': 'count'})
# ### Graf
gendersAggDf.plot.pie(y='count', title='Účastníci na Wikigapu podle pohlaví', figsize=(8,8));
# ### Data
# #### Agregováno
gendersAggDf
# #### Surová data
gendersDf
| wikigap-2021-report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: CLONE 11
# language: python
# name: arcgispro-py3-clone11
# ---
# <img src="..\..\..\pics\panda2.jpg" width=600/>
# <h2>Why use the spatially enabled dataframe</h2>
#
# <ul>
# <li>In Memory</li>
# <li>Fast computation</li>
# <li>On the fly indexing</li>
# <li>Multi-platform</li>
# <li>All the benefits of pandas and more</li>
#
# </ul>
import pandas as pd
from arcgis.features import GeoAccessor, GeoSeriesAccessor
from arcgis import GIS
gis = GIS('https://arcgis.com', 'bhammersley_tech')
# <h1> Data Wrangling </h1>
df = pd.read_csv(r'C:\Users\bhammersley\OneDrive - ESRI (UK) Ltd\Documents\Presentations\Berlin_2019\data\HH.csv')
df.head()
# +
def f():
pass
df["full_address"] = df.apply(
lambda row: '%s %s %s %s %s' %
(row['Name'],row['Street'],row['Town'],row['County'],row['Postcode'])
if str(row["Name"]) != 'nan'
else ('%s %s %s %s %s' %
(int(row['Number']),row['Street'],row['Town'],row['County'],row['Postcode'])
if str(row["Number"]) != 'nan' else f()), axis=1)
df = df.drop(['Name', 'Number', 'Street', 'Town', 'County', 'Postcode'], axis=1).dropna().reset_index(drop=True)
df.head()
# -
df.type.value_counts()
df.type = df.type.str.lower()
q = df.type == 'semi-detached'
df.loc[q, 'type'] = 'semi detached'
df.type.value_counts()
sdf = pd.DataFrame.spatial.from_df(df, 'full_address')
sdf.head()
# <h1> Visualisation </h1>
#
# <ul>
# <li>Matplotlib syntax</li>
# </ul>
# <img src="..\..\..\pics\VIZ.jpg" width=600/>
m = gis.map()
m.center = {'spatialReference': {'latestWkid': 3857, 'wkid': 102100},'x': 17740.16697718523,'y': 6833320.7425390035}
m.zoom = 11
m
sdf.spatial.plot(map_widget=m,
renderer_type='u', # specify the unique value renderer using its notation 'u'
col='type') # column to get unique values from
bbox = sdf.spatial.bbox
m.draw(shape = bbox)
sdf.spatial.to_featurelayer('Berlin dev summit houses', gis=gis)
# <h1> Geoenrichment </h1>
housing_item = gis.content.get('b92ce1f0a169498f8243056a88564125')
housing_lyr = housing_item.layers[0]
sdf = pd.DataFrame.spatial.from_layer(housing_lyr)
sdf.head()
sdf.spatial.sr
sdf.spatial.project(4326)
# +
analysis_variables = [
'EDUC01_CY', # 2017 POP age 16+ by Education: No qualifications
'EDUC02_CY', # 2017 POP age 16+ by Education: Level 1 qualifications
'EDUC03_CY', # 2017 POP age 16+ by Education: Level 2 qualifications
'EDUC04_CY', # 2017 POP age 16+ by Education: Level 3 qualifications
'EDUC05_CY', # 2017 POP age 16+ by Education: Level 4 qualifications and above
'UNEMP_CY', # 2016 unemployed population
'POPDENS_CY', # 2017 population density
'PPPC_CY', # 2017 Purchasing power by capita
'HINC01_CY', # Total households in lowest quintile (below £19,158)
'HINC01_CY', # Total households in 2nd quintile (£19,158 £28,123)
'HINC01_CY', # Total households in 3rd quintile (£28,124 to £38,084)
'HINC01_CY', # Total households in 4th quintile (£38,085 to £54,646)
'HINC01_CY', # Total households in 5th quintile (£54,646 to £19,158)
'HTYP01A_CY', # Households by type: Single person
'HTYP02A_CY', # Households by type: Married couple with dependent children
'HTYP03A_CY', # Households by type: Married couple with no children
'HTYP04A_CY', # Households by type: Cohabiting couple with dependent children
'HTYP05A_CY', # Households by type: Cohabiting couple with no dependent children
'HTYP06A_CY', # Households by type: Single parent with dependent children
'HTYP07A_CY', # Households by type: Single parent with no dependent children
'HTYP08A_CY' # Households by type: Other household types
]
# +
from arcgis.geoenrichment import enrich
enriched_sdf = enrich(sdf, analysis_variables=analysis_variables)
# -
enriched_sdf.head()
enriched_sdf.spatial.to_featurelayer('enriched houses')
| advanced-scripting-with-the-arcgis-api-for-python-eu-dev-summit-19/SPATIALLY ENABLED DATAFRAME.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (Recommender)
# language: python
# name: reco_base
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Factorization Machine Deep Dive
#
# Factorization machine (FM) is one of the representative algorithms that are used for building content-based recommenders model. The algorithm is powerful in terms of capturing the effects of not just the input features but also their interactions. The algorithm provides better generalization capability and expressiveness compared to other classic algorithms such as SVMs. The most recent research extends the basic FM algorithms by using deep learning techniques, which achieve remarkable improvement in a few practical use cases.
#
# This notebook presents a deep dive into the Factorization Machine algorithm, and demonstrates some best practices of using the contemporary FM implementations like [`xlearn`](https://github.com/aksnzhy/xlearn) for dealing with tasks like click-through rate prediction.
# ## 1 Factorization Machine
# ### 1.1 Factorization Machine
# FM is an algorithm that uses factorization in prediction tasks with data set of high sparsity. The algorithm was original proposed in [\[1\]](https://www.csie.ntu.edu.tw/~b97053/paper/Rendle2010FM.pdf). Traditionally, the algorithms such as SVM do not perform well in dealing with highly sparse data that is usually seen in many contemporary problems, e.g., click-through rate prediction, recommendation, etc. FM handles the problem by modeling not just first-order linear components for predicting the label, but also the cross-product of the feature variables in order to capture more generalized correlation between variables and label.
# In certain occasions, the data that appears in recommendation problems, such as user, item, and feature vectors, can be encoded into a one-hot representation. Under this arrangement, classical algorithms like linear regression and SVM may suffer from the following problems:
# 1. The feature vectors are highly sparse, and thus it makes it hard to optimize the parameters to fit the model efficienly
# 2. Cross-product of features will be sparse as well, and this in turn, reduces the expressiveness of a model if it is designed to capture the high-order interactions between features
# <img src="https://recodatasets.blob.core.windows.net/images/fm_data.png?sanitize=true">
# The FM algorithm is designed to tackle the above two problems by factorizing latent vectors that model the low- and high-order components. The general idea of a FM model is expressed in the following equation:
# $$\hat{y}(\textbf{x})=w_{0}+\sum^{n}_{i=1}w_{i}x_{i}+\sum^{n}_{i=1}\sum^{n}_{j=i+1}<\textbf{v}_{i}, \textbf{v}_{j}>x_{i}x_{j}$$
# where $\hat{y}$ and $\textbf{x}$ are the target to predict and input feature vectors, respectively. $w_{i}$ is the model parameters for the first-order component. $<\textbf{v}_{i}, \textbf{v}_{j}>$ is the dot product of two latent factors for the second-order interaction of feature variables, and it is defined as
# $$<\textbf{v}_{i}, \textbf{v}_{j}>=\sum^{k}_{f=1}v_{i,f}\cdot v_{j,f}$$
# Compared to using fixed parameter for the high-order interaction components, using the factorized vectors increase generalization as well as expressiveness of the model. In addition to this, the computation complexity of the equation (above) is $O(kn)$ where $k$ and $n$ are the dimensionalities of the factorization vector and input feature vector, respectively (see [the paper](https://www.csie.ntu.edu.tw/~b97053/paper/Rendle2010FM.pdf) for detailed discussion). In practice, usually a two-way FM model is used, i.e., only the second-order feature interactions are considered to favor computational efficiency.
# ### 1.2 Field-Aware Factorization Machine
# Field-aware factorization machine (FFM) is an extension to FM. It was originally introduced in [\[2\]](https://www.csie.ntu.edu.tw/~cjlin/papers/ffm.pdf). The advantage of FFM over FM is that, it uses different factorized latent factors for different groups of features. The "group" is called "field" in the context of FFM. Putting features into fields resolves the issue that the latent factors shared by features that intuitively represent different categories of information may not well generalize the correlation.
#
# Different from the formula for the 2-order cross product as can be seen above in the FM equation, in the FFM settings, the equation changes to
# $$\theta_{\text{FFM}}(\textbf{w}\textbf{x})=\sum^{n}_{j1=1}\sum^{n}_{j2=j1+1}<\textbf{v}_{j1,f2}, \textbf{v}_{j2,f1}>x_{j1}x_{j2}$$
# where $f_1$ and $f_2$ are the fields of $j_1$ and $j_2$, respectively.
# Compared to FM, the computational complexity increases to $O(n^2k)$. However, since the latent factors in FFM only need to learn the effect within the field, so the $k$ values in FFM is usually much smaller than that in FM.
# ### 1.3 FM/FFM extensions
# In the recent years, FM/FFM extensions were proposed to enhance the model performance further. The new algorithms leverage the powerful deep learning neural network to improve the generalization capability of the original FM/FFM algorithms. Representatives of the such algorithms are summarized as below. Some of them are implemented and demonstrated in the microsoft/recommenders repository.
#
# |Algorithm|Notes|References|Example in Microsoft/Recommenders|
# |--------------------|---------------------|------------------------|
# |DeepFM|Combination of FM and DNN where DNN handles high-order interactions|[\[3\]](https://arxiv.org/abs/1703.04247)|-|
# |xDeepFM|Combination of FM, DNN, and Compressed Interaction Network, for vectorized feature interactions|[\[4\]](https://dl.acm.org/citation.cfm?id=3220023)|[notebook](https://github.com/microsoft/recommenders/blob/master/notebooks/00_quick_start/xdeepfm_criteo.ipynb) / [utilities](https://github.com/microsoft/recommenders/blob/master/reco_utils/recommender/deeprec/models/xDeepFM.py)|
# |Factorization Machine Supported Neural Network|Use FM user/item weight vectors as input layers for DNN model|[\[5\]](https://link.springer.com/chapter/10.1007/978-3-319-30671-1_4)|-|
# |Product-based Neural Network|An additional product-wise layer between embedding layer and fully connected layer to improve expressiveness of interactions of features across fields|[\[6\]](https://ieeexplore.ieee.org/abstract/document/7837964)|-|
# |Neural Factorization Machines|Improve the factorization part of FM by using stacks of NN layers to improve non-linear expressiveness|[\[7\]](https://dl.acm.org/citation.cfm?id=3080777)|-|
# |Wide and deep|Combination of linear model (wide part) and deep neural network model (deep part) for memorisation and generalization|[\[8\]](https://dl.acm.org/citation.cfm?id=2988454)|[notebook](https://github.com/microsoft/recommenders/blob/master/notebooks/00_quick_start/wide_deep_movielens.ipynb) / [utilities](https://github.com/microsoft/recommenders/tree/master/reco_utils/recommender/wide_deep)|
# ## 2 Factorization Machine Implementation
# ### 2.1 Implementations
# The following table summarizes the implementations of FM/FFM. Some of them (e.g., xDeepFM and VW) are implemented and/or demonstrated in the microsoft/recommenders repository
# |Implementation|Language|Notes|Examples in Microsoft/Recommenders|
# |-----------------|------------------|------------------|---------------------|
# |[libfm](https://github.com/srendle/libfm)|C++|Implementation of FM algorithm|-|
# |[libffm](https://github.com/ycjuan/libffm)|C++|Original implemenation of FFM algorithm. It is handy in model building, but does not support Python interface|-|
# |[xlearn](https://github.com/aksnzhy/xlearn)|C++ with Python interface|More computationally efficient compared to libffm without loss of modeling effectiveness|[notebook](https://github.com/microsoft/recommenders/blob/master/notebooks/02_model/fm_deep_dive.ipynb)|
# |[Vowpal Wabbit FM](https://github.com/VowpalWabbit/vowpal_wabbit/wiki/Matrix-factorization-example)|Online library with estimator API|Easy to use by calling API|[notebook](https://github.com/microsoft/recommenders/blob/master/notebooks/02_model/vowpal_wabbit_deep_dive.ipynb) / [utilities](https://github.com/microsoft/recommenders/tree/master/reco_utils/recommender/vowpal_wabbit)
# |[microsoft/recommenders xDeepFM](https://github.com/microsoft/recommenders/blob/master/reco_utils/recommender/deeprec/models/xDeepFM.py)|Python|Support flexible interface with different configurations of FM and FM extensions, i.e., LR, FM, and/or CIN|[notebook](https://github.com/microsoft/recommenders/blob/master/notebooks/00_quick_start/xdeepfm_criteo.ipynb) / [utilities](https://github.com/microsoft/recommenders/blob/master/reco_utils/recommender/deeprec/models/xDeepFM.py)|
# Other than `libfm` and `libffm`, all the other three can be used in a Python environment.
#
# * A deep dive of using Vowbal Wabbit for FM model can be found [here](https://github.com/microsoft/recommenders/blob/master/notebooks/02_model/vowpal_wabbit_deep_dive.ipynb)
# * A quick start of Microsoft xDeepFM algorithm can be found [here](https://github.com/microsoft/recommenders/blob/master/notebooks/00_quick_start/xdeepfm_criteo.ipynb).
#
# Therefore, in the example below, only code examples and best practices of using `xlearn` are presented.
# ### 2.2 xlearn
# Setups for using `xlearn`.
#
# 1. `xlearn` is implemented in C++ and has Python bindings, so it can be directly installed as a Python package from PyPI. The installation of `xlearn` is enabled in the [Recommenders repo environment setup script](https://github.com/microsoft/recommenders/blob/master/scripts/generate_conda_file.py). One can follow the general setup steps to install the environment as required, in which `xlearn` is installed as well.
# 2. NOTE `xlearn` may require some base libraries installed as prerequisites in the system, e.g., `cmake`.
# After a succesful creation of the environment, one can load the packages to run `xlearn` in a Jupyter notebook or Python script.
# +
import time
import sys
sys.path.append("../../")
import os
import papermill as pm
from tempfile import TemporaryDirectory
import xlearn as xl
from sklearn.metrics import roc_auc_score
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib notebook
from matplotlib import pyplot as plt
from reco_utils.common.constants import SEED
from reco_utils.common.timer import Timer
from reco_utils.recommender.deeprec.deeprec_utils import (
download_deeprec_resources, prepare_hparams
)
from reco_utils.recommender.deeprec.models.xDeepFM import XDeepFMModel
from reco_utils.recommender.deeprec.IO.iterator import FFMTextIterator
from reco_utils.tuning.parameter_sweep import generate_param_grid
from reco_utils.dataset.pandas_df_utils import LibffmConverter
print("System version: {}".format(sys.version))
print("Xlearn version: {}".format(xl.__version__))
# -
# In the FM model building, data is usually represented in the libsvm data format. That is, `label feat1:val1 feat2:val2 ...`, where `label` is the target to predict, and `val` is the value to each feature `feat`.
#
# FFM algorithm requires data to be represented in the libffm format, where each vector is split into several fields with categorical/numerical features inside. That is, `label field1:feat1:val1 field2:feat2:val2 ...`.
# In the Microsoft/Recommenders utility functions, [a libffm converter](https://github.com/microsoft/recommenders/blob/290dd920d4a6a4d3bff71dd9ee7273be0c02dbbc/reco_utils/dataset/pandas_df_utils.py#L86) is provided to achieve the transformation from a tabular feature vectors to the corresponding libffm representation. For example, the following shows how to transform the format of a synthesized data by using the module of `LibffmConverter`.
# +
df_feature_original = pd.DataFrame({
'rating': [1, 0, 0, 1, 1],
'field1': ['xxx1', 'xxx2', 'xxx4', 'xxx4', 'xxx4'],
'field2': [3, 4, 5, 6, 7],
'field3': [1.0, 2.0, 3.0, 4.0, 5.0],
'field4': ['1', '2', '3', '4', '5']
})
converter = LibffmConverter().fit(df_feature_original, col_rating='rating')
df_out = converter.transform(df_feature_original)
df_out
# -
print('There are in total {0} fields and {1} features.'.format(converter.field_count, converter.feature_count))
# To illustrate the use of `xlearn`, the following example uses the [Criteo data set](https://labs.criteo.com/category/dataset/), which has already been processed in the libffm format, for building and evaluating a FFM model built by using `xlearn`. Sometimes, it is important to know the total numbers of fields and features. When building a FFM model, `xlearn` can count these numbers automatically.
# + tags=["parameters"]
# Parameters
YAML_FILE_NAME = "xDeepFM.yaml"
TRAIN_FILE_NAME = "cretio_tiny_train"
VALID_FILE_NAME = "cretio_tiny_valid"
TEST_FILE_NAME = "cretio_tiny_test"
MODEL_FILE_NAME = "model.out"
OUTPUT_FILE_NAME = "output.txt"
LEARNING_RATE = 0.2
LAMBDA = 0.002
# The metrics for binary classification options are "acc", "prec", "f1" and "auc"
# for regression, options are "rmse", "mae", "mape"
METRIC = "auc"
EPOCH = 10
OPT_METHOD = "sgd" # options are "sgd", "adagrad" and "ftrl"
# +
tmpdir = TemporaryDirectory()
data_path = tmpdir.name
yaml_file = os.path.join(data_path, YAML_FILE_NAME)
train_file = os.path.join(data_path, TRAIN_FILE_NAME)
valid_file = os.path.join(data_path, VALID_FILE_NAME)
test_file = os.path.join(data_path, TEST_FILE_NAME)
model_file = os.path.join(data_path, MODEL_FILE_NAME)
output_file = os.path.join(data_path, OUTPUT_FILE_NAME)
if not os.path.exists(yaml_file):
download_deeprec_resources(r'https://recodatasets.blob.core.windows.net/deeprec/', data_path, 'xdeepfmresources.zip')
# -
# The following steps are from the [official documentation of `xlearn`](https://xlearn-doc.readthedocs.io/en/latest/index.html) for building a model. To begin with, we do not modify any training parameter values.
# NOTE, if `xlearn` is run through command line, the training process can be displayed in the console.
# +
# Training task
ffm_model = xl.create_ffm() # Use field-aware factorization machine (ffm)
ffm_model.setTrain(train_file) # Set the path of training dataset
ffm_model.setValidate(valid_file) # Set the path of validation dataset
# Parameters:
# 0. task: binary classification
# 1. learning rate: 0.2
# 2. regular lambda: 0.002
# 3. evaluation metric: auc
# 4. number of epochs: 10
# 5. optimization method: sgd
param = {"task":"binary",
"lr": LEARNING_RATE,
"lambda": LAMBDA,
"metric": METRIC,
"epoch": EPOCH,
"opt": OPT_METHOD
}
# Start to train
# The trained model will be stored in model.out
with Timer() as time_train:
ffm_model.fit(param, model_file)
# Prediction task
ffm_model.setTest(test_file) # Set the path of test dataset
ffm_model.setSigmoid() # Convert output to 0-1
# Start to predict
# The output result will be stored in output.txt
with Timer() as time_predict:
ffm_model.predict(model_file, output_file)
# -
# The output are the predicted labels (i.e., 1 or 0) for the testing data set. AUC score is calculated to evaluate the model performance.
# +
with open(output_file) as f:
predictions = f.readlines()
with open(test_file) as f:
truths = f.readlines()
truths = np.array([float(truth.split(' ')[0]) for truth in truths])
predictions = np.array([float(prediction.strip('')) for prediction in predictions])
auc_score = roc_auc_score(truths, predictions)
# -
auc_score
pm.record('auc_score', auc_score)
print('Training takes {0:.2f}s and predicting takes {1:.2f}s.'.format(time_train.interval, time_predict.interval))
# It can be seen that the model building/scoring process is fast and the model performance is good.
# ### 2.3 Hyperparameter tuning of `xlearn`
# The following presents a naive approach to tune the parameters of `xlearn`, which is using grid-search of parameter values to find the optimal combinations. It is worth noting that the original [FFM paper](https://www.csie.ntu.edu.tw/~cjlin/papers/ffm.pdf) gave some hints in terms of the impact of parameters on the sampled Criteo dataset.
#
# The following are the parameters that can be tuned in the `xlearn` implementation of FM/FFM algorithm.
# |Parameter|Description|Default value|Notes|
# |-------------|-----------------|------------------|-----------------|
# |`lr`|Learning rate|0.2|Higher learning rate helps fit a model more efficiently but may also result in overfitting.|
# |`lambda`|Regularization parameter|0.00002|The value needs to be selected empirically to avoid overfitting.|
# |`k`|Dimensionality of the latent factors|4|In FFM the effect of k is not that significant as the algorithm itself considers field where `k` can be small to capture the effect of features within each of the fields.|
# |`init`|Model initialization|0.66|-|
# |`epoch`|Number of epochs|10|Using a larger epoch size will help converge the model to its optimal point|
# +
param_dict = {
"lr": [0.0001, 0.001, 0.01],
"lambda": [0.001, 0.01, 0.1]
}
param_grid = generate_param_grid(param_dict)
# +
auc_scores = []
with Timer() as time_tune:
for param in param_grid:
ffm_model = xl.create_ffm()
ffm_model.setTrain(train_file)
ffm_model.setValidate(valid_file)
ffm_model.fit(param, model_file)
ffm_model.setTest(test_file)
ffm_model.setSigmoid()
ffm_model.predict(model_file, output_file)
with open(output_file) as f:
predictions = f.readlines()
with open(test_file) as f:
truths = f.readlines()
truths = np.array([float(truth.split(' ')[0]) for truth in truths])
predictions = np.array([float(prediction.strip('')) for prediction in predictions])
auc_scores.append(roc_auc_score(truths, predictions))
# -
print('Tuning by grid search takes {0:.2} min'.format(time_tune.interval / 60))
# +
auc_scores = [float('%.4f' % x) for x in auc_scores]
auc_scores_array = np.reshape(auc_scores, (len(param_dict["lr"]), len(param_dict["lambda"])))
auc_df = pd.DataFrame(
data=auc_scores_array,
index=pd.Index(param_dict["lr"], name="LR"),
columns=pd.Index(param_dict["lambda"], name="Lambda")
)
auc_df
# -
fig, ax = plt.subplots()
sns.heatmap(auc_df, cbar=False, annot=True, fmt=".4g")
# More advanced tuning methods like Bayesian Optimization can be used for searching for the optimal model efficiently. The benefit of using, for example, `HyperDrive` from Azure Machine Learning Services, for tuning the parameters, is that, the tuning tasks can be distributed across nodes of a cluster and the optimization can be run concurrently to save the total cost.
#
# * Details about how to tune hyper parameters by using Azure Machine Learning Services can be found [here](https://github.com/microsoft/recommenders/tree/master/notebooks/04_model_select_and_optimize).
# * Note, to enable the tuning task on Azure Machine Learning Services by using HyperDrive, one needs a Docker image to containerize the environment where `xlearn` can be run. The Docker file provided [here](https://github.com/microsoft/recommenders/tree/master/docker) can be used for such purpose.
# ### 2.4 Clean up
tmpdir.cleanup()
# ## References
# <a id='references'></a>
# 1. <NAME>. "Factorization machines." 2010 IEEE International Conference on Data Mining. IEEE, 2010.
# 2. <NAME>, et al. "Field-aware factorization machines for CTR prediction." Proceedings of the 10th ACM Conference on Recommender Systems. ACM, 2016.
# 3. <NAME>, et al. "DeepFM: a factorization-machine based neural network for CTR prediction." arXiv preprint arXiv:1703.04247 (2017).
# 4. <NAME>, et al. "xdeepfm: Combining explicit and implicit feature interactions for recommender systems." Proceedings of the 24th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining. ACM, 2018.
# 5. <NAME>, et al. "Product-based neural networks for user response prediction." 2016 IEEE 16th International Conference on Data Mining (ICDM). IEEE, 2016.
# 6. Zhang, Weinan, <NAME>, and <NAME>. "Deep learning over multi-field categorical data." European conference on information retrieval. Springer, Cham, 2016.
# 7. He, Xiangnan, and <NAME>. "Neural factorization machines for sparse predictive analytics." Proceedings of the 40th International ACM SIGIR conference on Research and Development in Information Retrieval. ACM, 2017.
# 8. Cheng, Heng-Tze, et al. "Wide & deep learning for recommender systems." Proceedings of the 1st workshop on deep learning for recommender systems. ACM, 2016.
# 9. Langford, John, <NAME>, and <NAME>. "Vowpal wabbit online learning project." (2007).
| notebooks/02_model/fm_deep_dive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" id="4SmD11tDPWEL"
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.metrics import ConfusionMatrixDisplay
import tensorflow_addons as tfa
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten, GlobalAveragePooling2D,BatchNormalization,Conv2D, MaxPooling2D, ZeroPadding2D
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import RMSprop, Adam
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
from sklearn.metrics import classification_report, confusion_matrix
# + id="mdpENY0DPWEP"
train_data_dir = '../input/whitebloodcell/whitebloodcell/train'
validation_data_dir = '../input/whitebloodcell/whitebloodcell/val'
test_Data_dir='../input/whitebloodcell/whitebloodcell/test'
num_classes =10
batch_size = 16
img_rows, img_cols = 224, 224
train_datagen = ImageDataGenerator()
validation_datagen = ImageDataGenerator()
test_datagen = ImageDataGenerator()
# + id="POa-LNvVPWEQ"
train_generator = train_datagen.flow_from_directory(train_data_dir, target_size=(img_rows, img_cols), batch_size=batch_size, class_mode='categorical')
validation_generator = validation_datagen.flow_from_directory( validation_data_dir, target_size=(img_rows, img_cols), batch_size=batch_size, class_mode='categorical')
test_generator=test_datagen.flow_from_directory( test_Data_dir, target_size=(img_rows, img_cols), batch_size=batch_size, class_mode='categorical')
# + id="MkvAS_AmPWEQ"
def modelfunction(pretrained,weight):
for layer in pretrained.layers:
layer.trainable = False
def addTopModel(bottom_model, num_classes):
top_model = bottom_model.output
top_model = Flatten()(top_model)
top_model = Dense(num_classes, activation='softmax')(top_model)
return top_model
FC_Head = addTopModel(pretrained, num_classes)
model = Model(inputs=pretrained.input, outputs=FC_Head)
checkpoint = ModelCheckpoint(weight, monitor='val_loss', mode='min', save_best_only=True, verbose=1)
earlystop = EarlyStopping(monitor='val_loss', min_delta=0, patience=10, verbose=1, restore_best_weights=True)
learning_rate_reduction = ReduceLROnPlateau(monitor='val_accuracy', patience=5, verbose=1, factor=0.2, min_lr=0.0002)
callbacks = [earlystop, checkpoint, learning_rate_reduction]
metrics = ['accuracy', tf.keras.metrics.AUC(), tfa.metrics.CohenKappa(num_classes = 10), tfa.metrics.F1Score(num_classes = 10), tf.keras.metrics.Precision(), tf.keras.metrics.Recall()]
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=metrics)
batch_size = 32
history = model.fit_generator(train_generator, steps_per_epoch=4716 // batch_size, epochs=50, callbacks=callbacks, validation_data=validation_generator,
validation_steps=1571 // batch_size)
#plt.figure(figsize=(10,5))
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model Accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# "Loss"
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
model.evaluate(test_generator)
# + [markdown] id="0Ik15eNuPWER"
# # MobileNet
# + id="dOrl0qJpPWET"
from tensorflow.keras.applications import MobileNet
MobileNet = MobileNet(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(MobileNet,'MobileNet.h5')
# + [markdown] id="5dUFiZ0xPWEU"
# # VGG19
# + id="XwdyBYI9PWEV"
from tensorflow.keras.applications import VGG19
vgg = VGG19(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(vgg,'vgg.h5')
# + [markdown] id="T7VxbfSEPWEW"
# # Xception
# + id="bMTcQSCcPWEW"
from tensorflow.keras.applications import Xception
xception = Xception(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(xception,'xception.h5')
# + [markdown] id="Tl0FXUV3PWEX"
# # Inception V3
# + id="JrfeMvu5PWEX"
from tensorflow.keras.applications import InceptionV3
Inception = InceptionV3(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(Inception,'Inception.h5')
# + [markdown] id="Th6nOccRPWEY"
# # InceptionResNetV2
# + id="ek5AF8KePWEY"
from tensorflow.keras.applications import InceptionResNetV2
InceptionResNet = InceptionResNetV2(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(InceptionResNet,'InceptionResNet.h5')
# + [markdown] id="6X-p45Q-PWEY"
# # DenseNet121
# + id="2dHCNDPDPWEZ"
from tensorflow.keras.applications import DenseNet121
DenseNet = DenseNet121(weights='imagenet', include_top=False, input_shape=(img_rows, img_cols, 3))
modelfunction(DenseNet,'DenseNet.h5')
# + id="YBpHkniWPWEZ"
| Pretrained models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML Pipeline Implementation
# ## 1. Import libraries and download nltk packages
import pandas as pd
import numpy as np
from sqlalchemy import create_engine
import nltk
from nltk.tokenize import word_tokenize
from nltk.stem.porter import PorterStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from sklearn.feature_extraction.text import TfidfVectorizer
from nltk.corpus import stopwords
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
import re
from sklearn.metrics import confusion_matrix
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn import multioutput
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.pipeline import Pipeline
#ML models
from sklearn.multiclass import OneVsRestClassifier
from sklearn.svm import LinearSVC,SVC
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, accuracy_score
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import StandardScaler
import pickle
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
# ## 2. Define Functions
# 1. Define function to load data from database
# 2. Defining Tokenizer function
# 3. Define evaluation metric to find F1-score, Precision, Recall
def load_data(db_name,table_name):
'''
Input
db_name: take the database name that has to be loaded
table_name: table_name in the database that needs to be loaded
Output
X: Feature dataframe that will be given as input to the model
y: Target dataframe for the model
category_names: Multi-Label names for the target
'''
engine = create_engine('sqlite:///' + db_name)
df = pd.read_sql_table(table_name, engine)
X = df['message']
y = df.iloc[:, 4:]
category_names = list(df.columns[4:])
return X, y, category_names
def tokenize(text):
'''
Input
text: take the text as input
Output
words_lemmed: tokenized and lemmatized text with stop words removed
'''
text = re.sub(r"[^a-zA-Z0-9]", " ", text.lower())
stop_words = stopwords.words("english")
#tokenize
words = word_tokenize (text)
words_lemmed = [WordNetLemmatizer().lemmatize(w) for w in words if w not in stop_words]
return words_lemmed
def evaulation_metric(y_true,y_pred):
'''
Input
y_true: ground truth dataframe
y_pred: predicted dataframe
Output
report: dataframe that contains mean f1-score,precision and recall value for each class
'''
report = pd.DataFrame ()
for col in y_true.columns:
class_dict = classification_report (output_dict = True, y_true = y_true.loc [:,col], y_pred = y_pred.loc [:,col])
metric_df = pd.DataFrame (pd.DataFrame.from_dict (class_dict))
metric_df.drop(['macro avg', 'weighted avg'], axis =1, inplace = True)
metric_df.drop(index = 'support', inplace = True)
metric_df = pd.DataFrame (metric_df.transpose ().mean ())
metric_df = metric_df.transpose ()
report = report.append (metric_df, ignore_index = True)
report.index = y_true.columns
return report
# ## 3. Load Data
# 1. Load the data from database
# 2. Split the data into train and test
X, y, category_names = load_data('DisasterResponse.db','message_and_category')
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 22)
# ## 4. Define ML pipeline
# 1. Define ML pipeline
# 2. Define search space
# 3. Define GridSearch
# 4. Train the model
# 5. Predict on Test set
# 6. Get the Evaluation metric
pipeline = Pipeline([('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('scale',StandardScaler(with_mean=False)),
('clf', OneVsRestClassifier(LinearSVC()))])
search_space = [{'clf':[OneVsRestClassifier(LinearSVC())],
'clf__estimator__C': [1, 10, 100]},
{'clf': [OneVsRestClassifier(LogisticRegression(solver='sag'))],
'clf__estimator__C': [1, 10, 100]},
{'clf': [OneVsRestClassifier(MultinomialNB())],
'clf__estimator__alpha': [0.1, 0.5, 1]},
{'clf':[multioutput.MultiOutputClassifier(RandomForestClassifier())]}]
cv = GridSearchCV(pipeline, search_space)
cv.fit(X_train,y_train)
pickle.dump(cv, open('pipeline.sav', 'wb'))
def evaulation_metric(y_true,y_pred):
'''
Input
y_true: ground truth dataframe
y_pred: predicted dataframe
Output
report: dataframe that contains mean f1-score,precision and recall value for each class
'''
report = pd.DataFrame ()
for col in y_true.columns:
class_dict = classification_report (output_dict = True, y_true = y_true.loc [:,col], y_pred = y_pred.loc [:,col])
metric_df = pd.DataFrame (pd.DataFrame.from_dict (class_dict))
metric_df.drop(['macro avg', 'weighted avg'], axis =1, inplace = True)
metric_df.drop(index = 'support', inplace = True)
metric_df = pd.DataFrame (metric_df.transpose ().mean ())
metric_df = metric_df.transpose ()
report = report.append (metric_df, ignore_index = True)
report.index = y_true.columns
return report
y_predict = cv.predict(X_test)
y_predict = pd.DataFrame (y_predict, columns = y_test.columns)
report = evaulation_metric(y_test,y_predict)
print(report)
| ML_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras import layers, models, optimizers, metrics, losses
from keras.datasets import fashion_mnist
import matplotlib.pyplot as plt
# # Load Fashion-MNIST dataset
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# # Visualize Fashion-MNIST dataset
# +
print("train_images.shape", train_images.shape)
print("image shape", train_images[0].shape)
print("train_labels.shape", train_labels.shape)
print("len(train_labels)", len(train_labels))
print("test_images.shape", test_images.shape)
print("image shape", test_images[0].shape)
print("test_labels.shape", test_labels.shape)
print("len(test_labels)", len(test_labels))
# -
train_labels[0:10]
n = 3
for i in range(0, n * n):
plt.subplot(n, n, 1 + i)
plt.imshow(train_images[i], cmap='gray')
plt.show()
# # Format the dataset
# +
from keras.utils import to_categorical
train_images = train_images.reshape((60000, 28, 28, 1))
train_images = train_images.astype('float32') / 255
train_labels = to_categorical(train_labels.copy())
test_images = test_images.reshape((10000, 28, 28, 1))
test_images = test_images.astype('float32') / 255
test_labels = to_categorical(test_labels.copy())
print("train_images.shape", train_images.shape)
print("image shape", train_images[0].shape)
print("train_labels.shape", train_labels.shape)
print("train_labels\n", train_labels)
print("train_labels[0]", train_labels[0])
# -
# # Design the network
# +
net = models.Sequential()
net.add(layers.Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=(28, 28, 1))) # need 4D tensor so we need (batch, 28, 28, 1)
net.add(layers.MaxPool2D((2, 2)))
net.add(layers.Conv2D(64, kernel_size=(3, 3), activation='relu'))
net.add(layers.MaxPooling2D(pool_size=(2, 2)))
net.add(layers.Conv2D(64, kernel_size=(3, 3), activation='relu'))
net.add(layers.Flatten())
net.add(layers.Dense(64, activation='relu'))
net.add(layers.Dense(10, activation='softmax'))
net.summary()
net.compile(optimizer=optimizers.Adam(),
loss=losses.categorical_crossentropy,
metrics=[metrics.categorical_accuracy])
# -
history = net.fit(train_images,
train_labels,
epochs=10,
batch_size=64,
validation_data=(test_images, test_labels))
# `The result is about 94.26% on training set and 90.9% on the test set`
from utils import plot_history
plot_history(history)
# # Using data augmentation
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
rotation_range=20,
width_shift_range=0.05,
height_shift_range=0.05,
shear_range=0.15,
#zoom_range=0.15,
horizontal_flip=True,
fill_mode='nearest')
# ## Visualize the data augmentation
# +
from keras.preprocessing import image
x = train_images[0] # (28, 28, 1)
x = x.reshape((1,) + x.shape) # (1, 28, 28, 1)
n = 3
i = 0
for batch in datagen.flow(x, batch_size=1):
plt.subplot(n, n, 1 + i)
plt.imshow(image.array_to_img(batch[0]), cmap='gray')
i += 1
if i % (n * n) == 0:
break
plt.show()
# -
# # Change the network architecture a bit
# +
net = models.Sequential()
net.add(layers.InputLayer(input_shape=(28, 28, 1)))
net.add(layers.BatchNormalization())
net.add(layers.Conv2D(32, kernel_size=(3, 3), activation='relu'))
#net.add(layers.Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=(28, 28, 1))) # need 4D tensor so we need (batch, 28, 28, 1)
net.add(layers.MaxPool2D((2, 2)))
net.add(layers.Conv2D(64, kernel_size=(3, 3), activation='relu'))
net.add(layers.MaxPooling2D(pool_size=(2, 2)))
net.add(layers.Conv2D(64, kernel_size=(3, 3), activation='relu'))
net.add(layers.Flatten())
net.add(layers.Dropout(0.5))
net.add(layers.Dense(128, activation='relu'))
net.add(layers.Dense(10, activation='softmax'))
net.summary()
net.compile(optimizer=optimizers.Adam(),
loss=losses.categorical_crossentropy,
metrics=[metrics.categorical_accuracy])
# -
history = net.fit_generator(datagen.flow(train_images, train_labels, batch_size=64),
epochs=40,
steps_per_epoch=len(train_images) / 64,
validation_data=(test_images, test_labels))
history = net.fit(train_images,
train_labels,
epochs=10,
batch_size=64,
validation_data=(test_images, test_labels))
| series/getting_started/2. Recognize clothes images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ezGUEDo1vG4F" colab_type="text"
# ## In this notebook, we will train a CNN model on the [MNIST](https://en.wikipedia.org/wiki/MNIST_database) dataset and use *activation maximization* to visualize the features that the trained model has learnt.
# + [markdown] id="WG3N5WW12QoC" colab_type="text"
# ### The MNIST database (Modified National Institute of Standards and Technology database) is a database of handwritten digits (0 to 9). It contains 60,000 training images and 10,000 testing images.
# + [markdown] id="OF5RFj1dwo4e" colab_type="text"
# ### Step 1 Install and import all dependencies. (You can ignore the error messages in the outputs.)<br/>
# + id="E2m0dmAw2Eq-" colab_type="code" colab={}
import warnings
warnings.filterwarnings('ignore')
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.FATAL)
# !pip install --quiet --force-reinstall git+https://github.com/raghakot/keras-vis.git
# !pip install --quiet --force-reinstall scipy==1.2
# + id="Sek3y7-ENweQ" colab_type="code" colab={}
import numpy as np
from matplotlib import pyplot as plt
from vis.utils import utils
from vis.visualization import visualize_activation, visualize_cam
import keras
from keras.datasets import mnist
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Flatten, Activation, Input
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
from keras import activations
# + [markdown] id="wvlnsYYY3cs4" colab_type="text"
# ### **Step 2** Download the data. For this exercise, we are using the MNIST dataset. This is a dataset of handwritten digits from 0 to 9.
# + id="QxhtJqXxNO5S" colab_type="code" colab={}
# input image dimensions
img_rows, img_cols = 28, 28
# Download the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
# convert class vectors to categorical class matrices
num_classes = 10
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + [markdown] id="mkeTWL_sIW4z" colab_type="text"
# ### **Step 3** Visualize a sample in MNIST dataset.
# + id="4Y50MHSDIkOW" colab_type="code" colab={}
fig = plt.figure(figsize=(2, 2))
plt.imshow(x_train[0][:,:,0])
# + [markdown] id="SqgKOmJE4YE4" colab_type="text"
# ### **Step 4** Architect and compile your deep learning model. This is a multi-layer CNN architecture. The last layer consists of 10 filters, where each filter corresponds to a number from 0 to 9. The model classifies the input image of a handwritten digit into a number from 0 to 9, depending on which filter maximizes the output activation.
# ### Model below has been adapted from [Keras MNIST CNN documentation](https://keras.io/examples/mnist_cnn/).
# + id="DOyzSFf3OGJm" colab_type="code" colab={}
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
# + id="fdl0W0qPyFZS" colab_type="code" colab={}
# summary of model's architecture
model.summary()
# + [markdown] id="KjuarLwhSFap" colab_type="text"
# ### Step 5 Train your model with the loaded data.
# + id="bd8WzbPoSL9L" colab_type="code" colab={}
batch_size = 128
epochs = 2
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + [markdown] id="5ud1WdmQSja4" colab_type="text"
# ### Step 6 Evaluate model accuracy with test data.
#
# + id="LKiNKACnUcP_" colab_type="code" colab={}
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# + [markdown] id="q5lSz0q8UXEU" colab_type="text"
# ### Step 7 Use *activation maximization* to visualize what the first filter in the last layer of our model has learnt.
# + id="c70QhniZUXZg" colab_type="code" colab={}
fig = plt.figure(figsize=(2, 2))
# index of last layer
layer_idx = 7
# index of the first filter
filter_idx = 0
# modifying activation of last layer to linear
model.layers[layer_idx].activation = activations.linear
model = utils.apply_modifications(model)
# visualize_activation is a function in keras-vis that runs activation maximization
# on your model and outputs an images producing maximum activation for a filter
img = visualize_activation(model, layer_idx, filter_indices=filter_idx, max_iter=1000, tv_weight=1., lp_norm_weight=0.)
plt.imshow(img[:,:,0])
# + [markdown] id="-16QwD7GS33F" colab_type="text"
# ### Step 8 Use activation maximization to visualize the images that maximize output of each filter in the last layer.
# + id="T7vobERBoXBZ" colab_type="code" colab={}
columns = 5
rows = 2
fig = plt.figure(figsize=(16, 16))
for i in range(0,10):
img = visualize_activation(model, layer_idx, filter_indices=[i], max_iter=1000, tv_weight=1., lp_norm_weight=0.)
fig.add_subplot(rows, columns, i+1)
plt.imshow(img[:,:,0])
plt.show()
# + id="Bj39GhNCfynK" colab_type="code" colab={}
| mnist_interpretation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Web Scraping with BeautifulSoup
# ### But first... what is HTML
#
# **HyperText Markup Language**: it is NOT a programming language. As its name points it is a *markup language* is used to indicate to the browser how to layout content.
#
# HTML is based on tags, which indicates what should be done with the content.
#
# The most basic tag is the `<html>`. Everything inside of it is HTML. **Important:** We need to use tags to delimit the scope, so we use open and close tags, like in the example:
# ```html
# <html>
# ...
# </html>
# ```
# Inside of an `html` tag, we can use other tags. Usually, a HTML page has two other scopes defined by tags: `head` and `body`. The content of the web page goes into the body. The head contains metadata about the page, like the title of the page (it sometimes stores JS, CSSs, etc.)
#
# When scrapping, we usually focus on what is inside of the `<body> <\body>`
# ```html
# <html>
# <head>
# ...
# </head>
# <body>
# ...
# </body>
# </html>
# ```
# There are many possible tags with different roles, for example `<p>` delimits a paragraph `<br>` breaks a line, `<a>` represents links
# <html>
# <head>
# </head>
#
# <body>
# <p>
# Paragraph
# <a href="https://www.github.com">Link to GitHub</a>
# </p>
# <p>
# See the link below:
# <a href="https://www.twitter.<EMAIL>">Twitter</a> </p>
# </body>
# </html>
# In the above example, the `<a>` tag presents an `href` attribute, which determines where the link goes.
#
# Elements (tags) may have multiple attributes to define its layout/behavior. The attribute `class`, for example, indicates the CSS that will be applied there. The attribute `id` is used sometimes to identify a tag
# ### Let's scrape
# First, we need to import the module we are using... BeautifulSoup
import requests
from bs4 import BeautifulSoup
# Let's get a page... using requests
result = requests.get("https://pythonprogramming.net/parsememcparseface/")
# We use the content, to get ready to scrape
#
# And we call/instantiate our BeautifulSoup object, using our response content.
content = result.content
soup=BeautifulSoup(content, "html.parser")
soup
# If we want, we can make it easier to read...
print(soup.prettify())
# We can use multiple attributes/methods depending on what we wanna scrape/get!
print(soup.title)
# We can deal with soup.title (which is a Tag object), getting name, content, parent, etc...
print(soup.title.name)
print(soup.title.string)
print(soup.title.parent.name)
print(soup.p)
soup.p['class']
# We can find all the items with the same Tag, and get them as an iterable object
all_para = soup.find_all('p')
print(type(all_para[1]))
# We can even iterate :-) (+ text vs. string)
for para in all_para:
print(para.string)
print(str(para.text))
print("----")
links = soup.find_all('a')
print(links[2])
for url in links:
print(url.text)
print(url.get('href'))
print(url.get('class'))
print("---")
divs = soup.find_all('div', attrs={"class": "container", "style":"max-width:1500px; min-height:100%"})
len(divs)
body = soup.find('div',attrs={"class":"body", })
print(body.prettify())
footer = soup.find('footer')
print(footer.prettify())
for child in footer.children:
print(child)
print("---")
len(list(body.children))
body.find('a')
for item in body.findAll('a') :
print(item.string + " is a link to " + item.get('href'))
if (item.has_attr('target')):
print("target is: " + item.get("target"))
body.findAll('div')
body.findAll('img')
len(list(body.descendants))
print(body.a)
for a in soup.findAll("a"):
if (a.has_attr("data-delay")):
print("YES: " + a.text)
else:
print("NO: " + a.text)
response = requests.get('https://github.com/igorsteinmacher/')
content = response.content
soup=BeautifulSoup(content, "html.parser")
soup
bio = soup.find('div', attrs={'class':'p-note'})
list(bio.children)[0].text
| notebooks/BeautifulSoup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9dzs83hUZmWt"
# **Business Problem** : You work for a Hedge Fund company in India.Your Employer wants you to Predict the Stock price of **Bajaj Finances** , so that he can choose whether to Invest in it or not.He gives you the Data set to create A Model to predict the Stock price.
# + [markdown] id="wbHTZJEKZlx2"
# **1.Importing Libraries**
# + id="OLqwoBbgYPlJ"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + [markdown] id="Udk_nxWKZpgl"
# **2.Importing Dataset**
# + colab={"base_uri": "https://localhost:8080/", "height": 244} id="E2VQGl7uZqBL" outputId="d5e5be32-d137-4cea-bf27-4d289cd07892"
df=pd.read_csv('BAJFINANCE.csv')
df.head()
# + [markdown] id="4zAza5PyaFt-"
# **3.Preliminary Analysis and Missing value Detection & Rectification**
# + id="ZYZXWxePaGXm"
df.set_index('Date',inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="B5V6aouia5zV" outputId="cdb04b11-6eec-4aff-e801-3d0a9d846c58"
# Lets just see an Overview of how the Stock Price changes in Time #
df['VWAP'].plot()
# + colab={"base_uri": "https://localhost:8080/"} id="syyWtRKLbLrk" outputId="6526118f-1537-45d8-e007-e6e7ec3ff1df"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="vrQYu4ohbOl2" outputId="e5215842-6317-45c3-a12a-6c3a8b2bf66c"
# Null value Detection #
df.isna().sum()
# + id="KsdEu0NsbOi8"
# We drop the column Trades as almost half of it is Missing #
df.drop('Trades',axis=1,inplace=True)
# + id="Uu_c6IaMdx61"
# Lets just drop the rest of the missing values as they are not present from the beginning upto row no. 446 #
# If they were missing inbetween rather that From the beginning , we could have used Imputation methods like Mean/Moving Avg to fill it #
df.dropna(inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="BYM8Oyt5d6Fs" outputId="ee1928b4-9c18-4d5a-c2ee-2fa81e3c3121"
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="9tgNz6XzeFXR" outputId="2aecac84-5ac5-4611-c75d-4d564c0cab4d"
# The problem of missing values is Over and the new Shape is given below #
df.shape
# + id="lDcyqKQueK_9"
data=df.copy()
# + colab={"base_uri": "https://localhost:8080/"} id="ud1AEaQoeMlj" outputId="fc426fd1-db0e-4373-e76a-3fcfa80d728d"
data.dtypes
# + [markdown] id="H9EMwD3ogbjJ"
# **4.Creation of the Rolling Statistic**
# + [markdown] id="Ac7gWX5T2Fhz"
# A rolling analysis of a time series model is often used to assess the model’s stability over time. When analyzing financial time series data using a statistical model, a key assumption is that the parameters of the model are constant over time. However, the economic environment often changes considerably, and it may not be reasonable to assume that a model’s parameters are constant. A common technique to assess the constancy of a model’s parameters is to compute parameter estimates over a rolling window of a fixed size through the sample. If the parameters are truly constant over the entire sample, then the estimates over the rolling windows should not be too different. If the parameters change at some point during the sample, then the rolling estimates should capture this instability.
# + colab={"base_uri": "https://localhost:8080/"} id="vn2EqoyReMiZ" outputId="15746b3d-04b5-4bff-d3fc-8144718b48c1"
data.columns
# + id="kxnMHDcYeMc3"
lag_features=['High','Low','Volume','Turnover']
window1=3
window2=7
# + id="6ZSTG8TueMSt"
for feature in lag_features:
data[feature+'rolling_mean_3']=data[feature].rolling(window=window1).mean()
data[feature+'rolling_mean_7']=data[feature].rolling(window=window2).mean()
# + id="_2e__ZAjgmAA"
for feature in lag_features:
data[feature+'rolling_std_3']=data[feature].rolling(window=window1).std()
data[feature+'rolling_std_7']=data[feature].rolling(window=window2).std()
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="gH-cOfXlgl18" outputId="c0391bd1-190b-4f1a-d5be-e6965067b5e4"
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ymam9xawgt7e" outputId="a9385ef2-ea2b-4f60-d384-ccf569b9b076"
# We can see that there are Null Values in the Newly formed Columns #
data.isna().sum()
# + id="maqNgTyugt5F"
# Since the null values are very low compared to the Dataset , lets drop them #
data.dropna(inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="THRlNyGZgt2Y" outputId="4cf7db44-22e8-48b2-a5dc-8709e0e0425d"
data.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="SOI9js02gtzt" outputId="b6855b4f-9a8f-49e5-9e29-cac97e032dd4"
data.columns
# + colab={"base_uri": "https://localhost:8080/"} id="ZFmBqTzBijHF" outputId="dbe2bc1e-4100-4223-b41f-5d1a9c906634"
data.shape
# + [markdown] id="zKlHnAdbisP8"
# **5.Splitting the Data into Trainset and TestSet**
# + id="DaJZn8ZDgtqm"
# Note: we are not spliting the data Randomly using Test train Split because we need proper chrological order in Timeseries Data#
training_data=data[0:4000]
test_data=data[4000:]
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="Y4pCOswPjDir" outputId="ccfd99d6-96dc-4064-b202-7f76dcf7cd3f"
training_data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="QspYzDyBSVGZ" outputId="1131a271-e7a0-467a-8556-fdcbb98d8ff0"
test_data.head()
# + id="AUNsFzytNWoC"
# These are features created by Rolling Analysis to Minimize Outliers and Unstability in Data #
Independent_Features = ['Highrolling_mean_3', 'Highrolling_mean_7',
'Lowrolling_mean_3', 'Lowrolling_mean_7', 'Volumerolling_mean_3',
'Volumerolling_mean_7', 'Turnoverrolling_mean_3',
'Turnoverrolling_mean_7', 'Highrolling_std_3', 'Highrolling_std_7',
'Lowrolling_std_3', 'Lowrolling_std_7', 'Volumerolling_std_3',
'Volumerolling_std_7', 'Turnoverrolling_std_3',
'Turnoverrolling_std_7']
# + [markdown] id="ZNk2SkVZFh3A"
# **6.Training and Fitting the Model**
# + colab={"base_uri": "https://localhost:8080/"} id="Xy5v82LJOOBq" outputId="573c5ea2-bad1-4926-cae3-2a9f518b5cd7"
# !pip install pmdarima
# + id="ZJEGJAvROj4l"
from pmdarima import auto_arima
# + [markdown] id="YWAnQ-vPPwSG"
# We will be using the Infamous Auto-Arima Machine Learning Technique Specialised for TimeSeries to train the Model
# + colab={"base_uri": "https://localhost:8080/"} id="ITm0JLQuPv5T" outputId="4d1ccdb0-d298-4e44-df81-c56d238c0088"
# We will set the 'trace' parameter to 'True' so that the model can consider all bundles of (p,d,q) to find the best #
model=auto_arima(y=training_data['VWAP'],exogenous=training_data[Independent_Features],trace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="-U4QyQB_RItO" outputId="12f3e791-07aa-40a3-a44e-92fe3a7c69ae"
model.fit(training_data['VWAP'],training_data[Independent_Features])
# + [markdown] id="atA65vg9G5Wa"
# **7.Predicting the Test Set**
# + colab={"base_uri": "https://localhost:8080/"} id="todP2r7uRjim" outputId="85c260d2-4433-4dc0-d319-744fb17f508b"
forecast=model.predict(n_periods=len(test_data), exogenous=test_data[Independent_Features])
# + colab={"base_uri": "https://localhost:8080/"} id="rgkzM2FcRtGv" outputId="154e5e74-4031-4a13-ca75-6add90d31c80"
test_data['Forecast_ARIMA']=forecast
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="9F9-zUycRvFC" outputId="c6f3c036-c27f-4f67-fd1e-8fe736e2a9f0"
# Lets plot the Actual Values to the Predicted values by the Model #
test_data[['VWAP','Forecast_ARIMA']].plot(figsize=(14,7))
# + [markdown] id="fxYMBxMXPAwT"
# Note that : The Model was predicting VWAP with very high Accuracy until March 2020 . Which we know what happened after that : 'The Corona pandemic'.After the pandemic the markets were super Unstable ,leading to such drastic highs and lows in predictions after March 2020.This is Clearly Visualized in the graph.
# + [markdown] id="1lkwLE7IHSgs"
# **8.Accuracy Matrix and R Squared Value**
# + colab={"base_uri": "https://localhost:8080/"} id="D3wWWetwHqpO" outputId="68ccf3d4-8e0c-48b8-8574-7c919037e1ac"
x = test_data.iloc[:,8].values
y = test_data.iloc[:,29].values
x = x.reshape(len(x),1)
y = y.reshape(len(y),1)
am = np.concatenate((x,y),1)
am[:10]
# x is the Actual VWAP and y is the predicted VWAP #
# + colab={"base_uri": "https://localhost:8080/"} id="sq4wqrBDRu40" outputId="d305fe2f-02e5-4815-8c39-5049deb23e9d"
from sklearn.metrics import r2_score,mean_absolute_error, mean_squared_error
r2_score(test_data['VWAP'],test_data['Forecast_ARIMA'])
# An R Squared Value of 0.89 makes this model a Good Fit #
# + colab={"base_uri": "https://localhost:8080/"} id="UKo2_gpwGi6B" outputId="b232c1e5-b37f-43fe-b941-eb8e6a24439f"
np.sqrt(mean_squared_error(test_data['VWAP'],test_data['Forecast_ARIMA']))
# + colab={"base_uri": "https://localhost:8080/"} id="axJ3r2r8Gis8" outputId="f99bb4f4-67a2-40a6-b222-8b1c469511b2"
mean_absolute_error(test_data['VWAP'],test_data['Forecast_ARIMA'])
# + [markdown] id="mahdLP2B5Ugb"
# **9.Lets check Accuracy for Test Set just before March 2020 just for the sake of Curiosity**
# + colab={"base_uri": "https://localhost:8080/"} id="yCOM5YIb5qVQ" outputId="4ca68c00-ceab-4142-e1a3-91a7d4c070d9"
test_data.shape
# + id="PZtigGRr8ikO"
pre_covid_test_data = test_data.iloc[:475,:]
post_covid_test_data = test_data.iloc[475:,:]
# + colab={"base_uri": "https://localhost:8080/"} id="kx36ND2L9y9P" outputId="6a394dda-3108-4aac-8e95-0e0838c39fbd"
forecast_a=model.predict(n_periods=len(pre_covid_test_data), exogenous=pre_covid_test_data[Independent_Features])
# + colab={"base_uri": "https://localhost:8080/"} id="0Dv-Ymh8_lgy" outputId="e7342299-aa78-4aa9-88f2-91ea341326f7"
forecast_b=model.predict(n_periods=len(post_covid_test_data), exogenous=post_covid_test_data[Independent_Features])
# + colab={"base_uri": "https://localhost:8080/"} id="rm16HFtt_tzF" outputId="6bb4e544-0324-4cc4-ef8d-aa5391349346"
pre_covid_test_data['Forecast_ARIMA']=forecast_a
# + colab={"base_uri": "https://localhost:8080/"} id="fpan6jWS_0oh" outputId="281364a1-f35b-4039-ec9f-d90dc78a6ef8"
post_covid_test_data['Forecast_ARIMA']=forecast_b
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="lPWH-ql0AE4Y" outputId="45078e22-e131-4b47-e48e-9c5396997ee0"
pre_covid_test_data[['VWAP','Forecast_ARIMA']].plot(figsize=(14,7))
# + colab={"base_uri": "https://localhost:8080/"} id="vqLxfLguAV_3" outputId="dd2622b6-aed0-4676-98bc-c9fd975ebea1"
r2_score(pre_covid_test_data['VWAP'],pre_covid_test_data['Forecast_ARIMA'])
# + colab={"base_uri": "https://localhost:8080/"} id="PFsC6pHmHKQx" outputId="0a7340da-fc91-4061-93eb-9d89d72005bc"
np.sqrt(mean_squared_error(pre_covid_test_data['VWAP'],pre_covid_test_data['Forecast_ARIMA']))
# + colab={"base_uri": "https://localhost:8080/"} id="cv7qd94EHK-g" outputId="d4924952-7c23-4f50-848e-f3c656c801c2"
mean_absolute_error(pre_covid_test_data['VWAP'],pre_covid_test_data['Forecast_ARIMA'])
# + [markdown] id="ogVzMyXVDioR"
# R Squared Value of Pre-Covid Predictions are pretty Amazing with a whooping Accuracy of almost 98% . Also Note that the forecast by Arima is mostly Under-Predicted except a few exceptions.Which is actually good while buying Stock market shares (ie.) you get more profit than your expectations.All said and done still the values MAE and RMSE looks a little bit higher , That is expected after all.Predicting Super Accurate stock prices is nearly Impossible.These predicted values cannot be used in Intra-day trading , but can still be effectively used in Swing trading and Long term Investmest
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="sk-ewNwqAKFI" outputId="df13ff88-22a9-47bb-9924-0cf8f937d422"
post_covid_test_data[['VWAP','Forecast_ARIMA']].plot(figsize=(14,7))
# + colab={"base_uri": "https://localhost:8080/"} id="_BVNNa77AEoZ" outputId="0a38732a-ba6f-4432-a4d4-b52189aeb58b"
r2_score(post_covid_test_data['VWAP'],post_covid_test_data['Forecast_ARIMA'])
# + [markdown] id="N_Xf6weHDp0n"
# As expected the R Squared Value of Post Covid test data is Disappointing and brought down the accuracy of the model.It is implictly proved that this model is not a good fit for this data from the R Squared Value.No need for MAE and RMSE to prove it again.
# + [markdown] id="0nDi-oRmJ1tV"
# **10.Predicting the Value of VWAP in Realtime**
# + id="b_KmrcdnhUQn"
check = pd.read_csv('07-04-2021-TO-06-05-2021BAJFINANCEEQN.csv')
# + [markdown] id="UdmoPkyeLKWA"
# Note that : The training data stops at August 2018 and the below data is from April 2021 till today (May 07 2021) .
# I know it is not perfectly done in Real time , but this the best I could do,to bring in the essence of realtime in this project . please bear with me
# + [markdown] id="is5p-U-iLhJY"
# ***The same preliminary Analysis , Null value Detection & Rectification , Data Cleaning and Creation of Rolling Variables is done for this small Dataset as well***
# + colab={"base_uri": "https://localhost:8080/", "height": 278} id="BOUB_pK7hT9o" outputId="b40da2ae-ad95-4bb9-cd65-811dfaa77493"
check.head()
# + id="bPQt1exRibza"
check.set_index('Date',inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="EhdvuWO2lGm6" outputId="9ab52293-8ea9-4420-ea9a-f3289ac21a6a"
check['VWAP'].plot()
# + id="VPvWAKL6ij3w"
check.drop('No. of Trades',axis=1,inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="k2DDCbVflzjo" outputId="8a7cd033-5a99-4d3e-e392-1a185bc71b8d"
check.shape
# + colab={"base_uri": "https://localhost:8080/"} id="Bk-oEghfl4Jw" outputId="ecc8fc3c-2f84-4b2b-91e5-7788f19bec0a"
check.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="9UsFSi7YmHwR" outputId="85d7574a-4482-4abc-91dd-7b9a415237c6"
check.columns
# + id="jo5W1_Z5mQCT"
lag_feature = ['High Price','Low Price','Total Traded Quantity', 'Turnover']
window3=3
window4=7
# + id="KAS30TZMmzwV"
for feature in lag_feature:
check[feature+'rolling_mean_3']=check[feature].rolling(window=window3).mean()
check[feature+'rolling_mean_7']=check[feature].rolling(window=window4).mean()
# + id="yfTX5R5TnDiv"
for feature in lag_feature:
check[feature+'rolling_std_3']=check[feature].rolling(window=window3).std()
check[feature+'rolling_std_7']=check[feature].rolling(window=window4).std()
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="uKz973BTnkJg" outputId="939d8b7d-8b51-4582-de43-d03965ebc326"
check.head()
# + colab={"base_uri": "https://localhost:8080/"} id="idweEcXtnpah" outputId="974a9845-f0da-4cee-8c59-a71c08c374d5"
check.isna().sum()
# + id="Pp4rtIgGn2mI"
check.dropna(inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="XtPtUl-voExn" outputId="e90eb2c8-8017-41a8-80a5-35cca805236d"
check.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="sOhYTkCYoIUf" outputId="d8a4b0f4-267b-4308-81a9-ace076d7d57d"
check.shape
# + colab={"base_uri": "https://localhost:8080/"} id="BGUZFLQ4oCOI" outputId="50507f15-3416-4b46-ff89-ccfc30841a5e"
check.columns
# + id="4znics_loQQw"
ind_features = ['High Pricerolling_mean_3',
'High Pricerolling_mean_7', 'Low Pricerolling_mean_3',
'Low Pricerolling_mean_7', 'Total Traded Quantityrolling_mean_3',
'Total Traded Quantityrolling_mean_7', 'Turnoverrolling_mean_3',
'Turnoverrolling_mean_7','High Pricerolling_std_3',
'High Pricerolling_std_7', 'Low Pricerolling_std_3',
'Low Pricerolling_std_7', 'Total Traded Quantityrolling_std_3',
'Total Traded Quantityrolling_std_7', 'Turnoverrolling_std_3',
'Turnoverrolling_std_7']
# + [markdown] id="4GW2n22nMJan"
# **11.Forecasting for the Second Dataset**
# + colab={"base_uri": "https://localhost:8080/"} id="d1M2b9lWpIxA" outputId="35f38959-2507-46be-ed78-85cf17b6ac47"
forecast1=model.predict(n_periods=len(check), exogenous=check[ind_features])
# + id="73tOfMw0qi9Z"
check['Forecast_ARIMA']=forecast1
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="eIft9zI6q2PZ" outputId="8ac45cd4-f0fd-4ccb-debd-0b752215fefa"
# plotting Predicted Values VS Actual Values #
check[['VWAP','Forecast_ARIMA']].plot(figsize=(14,7))
# + colab={"base_uri": "https://localhost:8080/"} id="CiuyLjZXrFNR" outputId="c7c37324-8dfe-4ddd-a519-bf26a12fecf3"
from sklearn.metrics import r2_score
r2_score(check['VWAP'],check['Forecast_ARIMA'])
# + [markdown] id="plGe_FKDJtNH"
# It is implictly proved that this model is not a good fit for this data from the R Squared Value.No need for MAE and RMSE to prove it again
# + [markdown] id="Q3d6jx_XMlTG"
# **We can see that R Squared Value has dropped Significantly from (0.89 to 0.74).The main reason for the fall in accuracy is beacuse of the Time period Gap inbetween August 2018 and April 2021.Many things could have happened in the market in these 3 years, whose effects are not Imbued into the model.**
| StockPrice_Prediction_TimeSeriesAnalysis[RealData]_UsingAuto_Arima.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeRateDogs Twitter Feed
# This project looks at various data sources for Tweets from the [WeRateDogs](https://twitter.com/dog_rates) Twitter account, specifically:
#
# 1. the `twitter-archive-enhanced.csv` which contains the tweet text, as is the core data set
# 1. the Twitter API is used to access the original tweets to retrieve missing fields such as the retweet and favorite counts
# 1. an image prediction file containing the top 3 predictions for each of the (up to 4) dog pictures in the tweet
#
# Having gathered the data, we assess, clean and analyse it.
#
# ---
# ---
#
# ## Gather
# We use a number of data assets including remote files on web servers, and JSON payloads returned by the Twitter API.
WE_RATE_DOGS_TWEETS_PATH = 'data/twitter-archive-enhanced.csv'
DOG_BREED_PREDICTIONS_SOURCE_URL = 'https://d17h27t6h515a5.cloudfront.net/topher/2017/August/599fd2ad_image-predictions/image-predictions.tsv'
# ### Gather the enhanced Tweets data
# Pandas `read_csv()` function is quite versatile when uploading data, and can be configured to handle different date formats, numeric data types, not available (NA) markers, etc. Getting this right upfront can save time, but requires the raw data in files to be eyeballed first. For this we can use command line tools like head & tail, or alternatively Excel, which allows column headings to be frozen, data to be sorted and searched, etc.
# Having looked at the raw data, we make the following observations:
#
# 1. tweet Ids are large integers, we need to select an approriate integer datatype so no accuracy is lost
# 1. some tweet Ids use floats, e.g.: `in_reply_to_status_id`, `in_reply_to_user_id`, with NaNs used as a Not Available marker, as mentioned above these need to be converted to integers
# 1. time stamps are close to ISO 8601 format, and are GMT
# Actions taken to address above observations:
#
# * convert floating point tweets Ids to a 64-bit integer, retaining the Not Available representation
# * specifcally tell Pandas which columns are dates
import yaml
import tweepy
import json
import numpy as np
import pandas as pd
# Load the enhanced Twitter archive, using explicit data types for fields, instead of letting Pandas infer them. The [Twitter API](https://developer.twitter.com/en/docs/twitter-api/v1/data-dictionary/overview/tweet-object) will define the data types for the Twitter sourced fields.
#
# To get around the fact that nullable numeric fields are interpreted by `read_csv()` as floats (thus allowing NaNs to represent null), we will map nullable tweet Ids to the Pandas nullable integer data type (Int64).
# +
feed_data_types = {
'tweet_id': np.int64,
'in_reply_to_status_id': 'Int64',
'in_reply_to_user_id': 'Int64',
'retweeted_status_id': 'Int64',
'retweeted_status_user_id': 'Int64',
'text': 'string',
'expanded_urls': 'string',
'rating_numerator': np.int32,
'rating_denominator': np.int32,
'name': 'string',
'doggo': 'string',
'floofer': 'string',
'pupper': 'string',
'puppo': 'string'
}
feed_date_cols = [
'timestamp',
'retweeted_status_timestamp'
]
# -
enhanced_tweets_df = pd.read_csv(WE_RATE_DOGS_TWEETS_PATH,
index_col=['tweet_id'],
dtype=feed_data_types,
parse_dates=feed_date_cols)
enhanced_tweets_df.shape
# The first discrepancy we note is that, according to the project motivation document, the main "archive contains basic tweet data for all 5000+ of their tweets" however that is clearly not the case as, having loaded it, the number of tweets is less than half that. As this is the master data set we have been provided with, this is the data we have to go with, since it has been previously enhanced.
#
# To sanity check this row count, and make sure we have actually read in all the eprovided data, we will run a line count on the input file, which should roughly match the number of rows in the data frame. Any discrepancy on counts is due to those embeded new line (NL) characters in the tweet text, since the number of NL characters is what `wc` bases its line counts on.
# !wc -l {WE_RATE_DOGS_TWEETS_PATH}
# Now we can double check the column data types, against the data type mapping provided to `read_csv()`.
enhanced_tweets_df.info()
# ### Gather the Twitter API enrichment data
# Next we want to use the Twitter API to retrieve the original tweets, so that we can enrich our enhanced tweets data with the missing attributes previously idientified (`retweet_counts`, `favorite_counts`).
# Having registered with Twitter as a developer, and obtained credentials and keys, we stored these in a private project directory and configuration file (which are excluded from our git repo, and thus won't be visible online in [github](https://github.com/benvens-udacity/wrangle-and-analyze-data/blob/main/wrangle_act.ipynb)).
#
# We now use those credentials to authenticate with Twitter for API access.
def read_creds(conf_path):
with open(conf_path, 'r') as cf:
config = yaml.load(cf, Loader=yaml.FullLoader)
return config
creds = read_creds('./config/private/creds.yaml')
consumer_key = creds['consumer_api']['key']
consumer_secret = creds['consumer_api']['secret']
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
access_token = creds['access_token']['token']
acess_secret = creds['access_token']['secret']
auth.set_access_token(access_token, acess_secret)
# Next we will load the enrichment data in batches, for better performance, as API invocations are subject to significant network latency. Twitter also applies rate limiting to their APIs, so it is necessary to throttle the rate at which we make requests, and to retry any failed requests. Luckily, this can be handled automatically by the Tweepy library, by setting the `wait_on_rate_limit_notify` flag when configuring API connection.
api = tweepy.API(auth, wait_on_rate_limit_notify=True)
def process_batch(batch):
idxs = []
retweet_counts = []
favorite_counts = []
for status in batch:
tweet = status._json
idxs.append(tweet['id'])
retweet_counts.append(tweet['retweet_count'])
favorite_counts.append(tweet['favorite_count'])
return np.array(idxs, dtype=np.int64), np.array([retweet_counts, favorite_counts], dtype=np.int64).T
indices = np.empty((0), dtype=np.int64)
rows = np.empty((0, 2), dtype=np.int64)
batch_size = 100
num_tweets = len(enhanced_tweets_df.index)
# %%time
for batch_start in range(0, num_tweets, batch_size):
batch_end = min(batch_start + batch_size, num_tweets)
batch_tweet_ids = enhanced_tweets_df.iloc[batch_start:batch_end].index.to_numpy().tolist()
statuses = api.statuses_lookup(batch_tweet_ids, include_entities=False, map_=False)
b_indices, b_rows = process_batch(statuses)
indices = np.concatenate((indices, b_indices), axis=0)
rows = np.concatenate((rows, b_rows), axis=0)
tweet_counts_df = pd.DataFrame(index=indices, data=rows,
columns=['retweet_counts', 'favorite_counts'],
dtype='Int32').sort_index()
tweet_counts_df.index.name = 'tweet_id'
tweet_counts_df.shape
# Again, we briefly double check on the expected column data type mapping.
tweet_counts_df.info()
# ### Gather the breed prediction data
# Finally we need to gather the breed prediction data. We will read this data from the CloudFront URL, as opposed to the local filesystem, to ensure we get the most up-to-date version.
img_preds_data_types = {
'tweet_id': 'Int64',
'jpg_url': 'string',
'img_num': np.int32,
'p1': 'string',
'p1_conf': np.float32,
'p1_dog': bool,
'p2': 'string',
'p2_conf': np.float32,
'p2_dog': bool,
'p3': 'string',
'p3_conf': np.float32,
'p3_dog': bool
}
# +
# Load the TSV (not CSV) records, and tell read_csv() to use a tab as the field separator
img_preds_df = pd.read_csv(DOG_BREED_PREDICTIONS_SOURCE_URL,
index_col=['tweet_id'],
sep='\t',
dtype=img_preds_data_types)
img_preds_df.shape
# -
# And finally we check for correct data type mapping.
img_preds_df.info()
# ---
# ---
#
# ## Assess
# Having gathered the data we will now assess it, ideally both visually and programmatically.
#
# Some of this visual assesment has already been done against the raw data in files, to ensure we used appropriate data types when uploading the data. Therefore some data quality issues (large integers stored as floating point, with potential loss of accuracy, which invalidates their meaning as an identifier) have been addressed at upload time.
# ### Visual assessment
# We will inspect the data that has been uploaded into the corresponding dataframes.
# +
# Raise the number of viewable rows and columns
# Retain some kind of row counts, as very large data sets may get loaded into the browser, causing memory issues
pd.set_option('display.max_rows', 10000)
pd.set_option('display.max_columns', None)
pd.set_option('display.max_colwidth', None)
# -
# #### Enhanced tweets
# We assess some tweets that include a dog stage name.
enhanced_tweets_df[(enhanced_tweets_df[['doggo', 'floofer', 'pupper', 'puppo']] != 'None').any(axis=1)].head()
# We observe the following:
#
# 1. HTML in the `source` columns, with a lot of repetition (to be verified programmatically)
# 1. the varios rewteet columns frequently hold null values
# 1. on occasions multiple values appearing in the `expanded_urls` column, including repeating values
# 1. quite often no dog stage can be identified, and occasionally no dog name
# 1. dog stages place the stage name in a column named after the stage, this is redundant information
#
# #### Retweet and favorite counts
tweet_counts_df.head()
# There are no immediate issues observed by assessing a small sample of the tweet counts data visually.
# #### Breed predictions
img_preds_df.head()
# We observe the following:
#
# 1. each row refers to an image
# 1. each image is numbered, as it is selected as the best of up to 4 dog images that may be associated with each tweet
# 1. we then have the top 3 breed predictions for that image
#
# Each prediction consists of the following information:
#
# 1. a predicted label or class (e.g.: the dog breed) that describes the image
# 1. a confidence score associated with the above prediction, in the range 0.0 -> 1.0 (0% to 100% confident)
# 1. a boolean indicator confirming if the predicted label is a dog breed, or some other object
#
# Looking at the confidence score for predictions p1 - p3, they appear to be listed in most confident to least confident order. Therefore we will use the column name numeric suffix to generate a ranking column, which we can later sort by (to preserve this decreasing confidence order).
#
# This last attribute confirms that the image classifier used to generate these prediction was trained on a broad set of images, only a subset of which are dog images labelled with their corresponding dog breed. But on occasions the classifier may have interpreted a dog image as an object other than a dog.
# ### Programmatic assessment
# Programmatic assesment gives us the opportunity to validate observations, and search for anomalies, across the entire dataset. This is very difficult to do visually unless the dataset is small, both in trems of the number of rows and columns.
# #### Enhanced tweets
# Assess level of repetition in the `source` column, which holds an HTML anchor node.
enhanced_tweets_df['source'].value_counts()
# Looking at the above results there appear to be 4 sources corresponding to the related applications: iPhone Twitter app, Vine app, Twitter web client and TweetDeck. This data contains a lot of redundant and messy information.
# Check if there are tweets where more than one dog stage is mentioned.
((enhanced_tweets_df[['doggo', 'floofer', 'pupper', 'puppo']] != 'None').sum(axis=1) > 1).sum()
# #### Retweet and favorite counts
# We will quickly validate that all counts are positive.
(tweet_counts_df >= 0).all()
# We will compare the number of entries in the enriched tweets dataframe to the number of entries in the tweet counts dataframe, to see of we successfully retrieved counts for all tweets from the API. The small difference in counts suggests a small number of tweets can no longer be retrieved.
len(enhanced_tweets_df.index), len(tweet_counts_df.index)
# #### Breed predictions
# We will validate the assumption made earlier that the confidence scores are ordered by the numeric suffix of the column name, which can be used to populate a ranking.
((img_preds_df['p1_conf'] > img_preds_df['p2_conf']) & (img_preds_df['p2_conf'] > img_preds_df['p3_conf'])).all()
# Next we validate that all confidence scores are in the range 0.0 to 1.0.
(img_preds_df['p1_conf'].between(0.0, 1.0) &
img_preds_df['p2_conf'].between(0.0, 1.0) &
img_preds_df['p3_conf'].between(0.0, 1.0)).all()
# ### Quality issues found
# As a result of the visual and programmatic assessments, the following data quality have been found, which will require data content to be cleaned.
# #### Enhanced tweets
# 1. the immediate data quality concern is that the project motivation document states that the "archive contains basic tweet data for all 5000+ of their tweets" but we are loading less than half that number of tweets. **However, given the enhanced tweets dataset is our master dataset, there is nothing that we can do to remedy the much smaller number of rows, beyond highlighting this observation**
# 1. as previously mentioned, the issue with some tweet Id columns being treated as floating point numbers, and the fact that rounding could invalidate these, was resolved at data loading time (without impacting the fact that they are nullable columns)
# 1. the format of the `timestamp` is very close to an ISO 8601 timestamp, however it is missing the 't' character as the separator between the date and time portions. There are definite advantages in following a recognised standard, as this will be understood by tools such as database import utilities, however Pandas has correctly parsed dates
# 1. in the `source` column, extract the source app name from the HTML anchor string, and then map this column to a Pandas categorical
# 1. it is unclear why, in the `expanded_urls` columns, the same URL get repeated, since looking at the tweet text there is only one reference to the corresponding link. Therefore we will remove duplicates
# 1. convert the dog stage columns into boolean datatype, and interpret the constant value 'None' as a missing stage
# 1. since the dog stage column names are the stages, storing that same name as a value is redundant information, following on from the previous observation, where the dog stage appears we will just store a boolean true value
# #### Retweet and favorite counts
# 1. while the intention is to obtain retweet and favorite counts for all the tweets in the enhanced tweets dataset, we cannot guarantee that the Twitter API will always return the original Tweet, e.g.: it may subsequently have been deleted
# 1. where the counts were successfully retrieved for the original tweet (the majority of cases, as proven in the programatic assesment), then there is a one-to-one relationship between the rows in the counts dataframe, and the rows in the enhanced tweets dataframe. Therefore the counts columns can be merged back into the enhanced tweets dataframe, as arguably they are part of that tweet observation. In the few cases where the counts are missing, we will store nulls
# #### Breed predictions
# No obvious data quality issues, beyond the prediction column names being used as variables (the numeric suffix added).
# ### Structural issues
# After looking at data frame structure, column naming, and inspecting values, and then applying the [Tidy Data](https://vita.had.co.nz/papers/tidy-data.pdf) principles, the following structural issues will need to be addressed.
# #### Enhanced tweets
# 1. the `source` column must store a category that represent the application (and possibly device) used to author the tweet
# 1. the `expanded_urls` column can store multiple values per row, depending on the web links embeded in the tweet text, therefore these observations need to be stored in a separate table (however, we will first remove any duplicate values).
# 1. dog stage is a multivalued categorical variable, as a tweet can reference more than one stage. Therefore we retain the existing columns but encode them in the style of one hot encoding
# #### Retweet and favorite counts
# No obvious structural issues here.
# #### Breed predictions
# 1. a variable (prediction number) is embeded in the column names of the prediction columns (predicted breed, prediction confidence, and is-a-dog flag)
# 2. the prediction number ranks the predictions in the order most confident (1st prediction) to least confident (3rd prediction)
# 3. the actual breed predictions should be held in a separate dataframe, and linked back to the tweet and tweet image they are associated with
# ---
# ---
#
# ## Clean
# We will now clean the issues uncovered during assesment using a _define/code/test_ framework, which will be applied to each of the issues.
clean_enhanced_tweets_df = enhanced_tweets_df.copy()
clean_enhanced_tweets_df.shape
# ---
#
# ### Extract tweet application from `source` column
# **Define**
#
# * parse source column which holds an HTML anchor node
# * extract anchor node content, describing the application used
# * convert the column to Pandas categorical, as a more efficient representation that can be used in models
# **Code**
# +
# Extract content from anchor node
clean_enhanced_tweets_df['source'] = \
clean_enhanced_tweets_df['source'].str.extract(r'[^<]*a href="[^"]+" rel="[^"]+">([^<]+)<\/a>')
# Convert column to categorical
clean_enhanced_tweets_df['source'] = clean_enhanced_tweets_df['source'].astype('category')
# -
# **Test**
# We will check that the tweet source column is now a categorical, and the number of categories is that expected.
# +
# Assert column data type is categorical
assert isinstance(clean_enhanced_tweets_df['source'].dtype, pd.CategoricalDtype),'Expect categorical'
# Assert the number of categories is as expected
assert len(clean_enhanced_tweets_df['source'].cat.categories) == 4, 'Expect 4 application categories'
# -
# ---
#
# ### Move `expanded_urls` to a detail dataframe
# **Define**
#
# * split multi-valued string of comma separated URLs, into URL arrays
# * remove any duplicate URLs from the array
# * convert each array into list of tuples, bound to the containing `tweet_id`
# * stores these tuples as rows in a new dataframe
# **Code**
# +
# Pull out rows containing one or more expanded URLs, as some rows have none
expanded_urls_col = \
clean_enhanced_tweets_df.loc[clean_enhanced_tweets_df['expanded_urls'].isna() == False]['expanded_urls']
# Nested list comprehension to split multiple URL strings on comma separator, then create [tweet Id, URL] tuples
expanded_url_tuples = [(ix, url) for ix, urls in expanded_urls_col.iteritems() for url in urls.split(',')]
expanded_url_df = pd.DataFrame(expanded_url_tuples, columns=['tweet_id', 'expanded_url'])
# Now drop duplicates and make 'tweet_id' the index for consistency with other dataframes
expanded_url_df = expanded_url_df.drop_duplicates().set_index('tweet_id')
# Finally drop the original expanded_urls column
clean_enhanced_tweets_df = clean_enhanced_tweets_df.drop(columns='expanded_urls')
# -
# **Test**
# We will count total and unique tweet Ids in the new dataframe holding expanded URLs. The later will be lower, accounting for multiple rows (hence web links in the tweet text) associated with the same tweet.
# +
# Note that the index can contain duplicate entries (whenever a tweet has more than one URL)
# We compare duplicate and non-duplicate counts below
len(expanded_url_df.index), len(expanded_url_df.index.unique())
# -
# ---
#
# ### Convert dog stage columns to boolean
# **Define**
#
# * where the value 'None' is stored, set False, otherwise set True
# **Code**
# +
# Convert dog stage columns into a boolean data type
stage_cols = ['doggo', 'floofer', 'pupper', 'puppo']
clean_enhanced_tweets_df[stage_cols] = clean_enhanced_tweets_df[stage_cols].apply(lambda c: c.to_numpy() != 'None')
# -
# **Test**
# We will check that the dog stage columns are now boolean type.
clean_enhanced_tweets_df.info()
for col in stage_cols:
assert clean_enhanced_tweets_df[col].dtype == 'bool', 'Expect boolean column'
# ---
#
# ### Merge retweet and favorite counts into enhanced tweets dataframe
# **Define**
#
# * merge retweet and favorite count columns into enhanced tweets dataframe, using a left join with nulls for missing count values
# **Code**
clean_enhanced_tweets_df = clean_enhanced_tweets_df.merge(tweet_counts_df, how='left', on='tweet_id')
# **Test**
# Validate number of rows after merge, including count of rows with null retweet or favorite
# +
# Count total rows (should be unchanged), and null retweet and favorite counts (tweets no longer available)
print(len(clean_enhanced_tweets_df.index))
clean_enhanced_tweets_df[['retweet_counts', 'favorite_counts']].isna().sum()
# -
# ---
#
# ### Melt image prediction column headers into detail dataframe
# **Define**
#
# * store `jpg_url` and `img_num` columns in a clean dataframe
# * melt prediction 1 to 3 columns into temporary dataframes, with the prediction rank as a constant value, and the related `tweet_id`
# * stack the above temporary dataframes into a predictions dataframe, with repeated `tweet_id` as the index
# **Code**
clean_img_preds_df = img_preds_df.copy()
clean_img_preds_df.shape
def melt_pred_cols(df, numeric):
preds_df = pd.DataFrame(data={'pred_rank': numeric,
'pred_class': df[f'p{numeric}'],
'pred_confidence': df[f'p{numeric}_conf'],
'pred_is_dog': df[f'p{numeric}_dog']})
return preds_df
# +
preds1_df = melt_pred_cols(clean_img_preds_df, 1)
preds2_df = melt_pred_cols(clean_img_preds_df, 2)
preds3_df = melt_pred_cols(clean_img_preds_df, 3)
clean_predictions_df = pd.concat([preds1_df,
preds2_df,
preds3_df]).sort_values(by=['tweet_id', 'pred_rank'])
# +
# Drop melted prediction columns
clean_img_preds_df = clean_img_preds_df.drop(columns=['p1', 'p1_conf', 'p1_dog', \
'p2', 'p2_conf', 'p2_dog', \
'p3', 'p3_conf', 'p3_dog'])
# -
# **Test**
# Validate dataframe column names and structure as expected.
clean_img_preds_df.info()
clean_predictions_df.info()
# +
# Validate master/detail row counts
assert len(clean_img_preds_df.index) == (len(clean_predictions_df.index) / 3), 'Expect 3x number of detail rows'
# -
# ---
# ---
#
# ## Analyse
# In this section we look at the data and analyse it to obtain some insights. Specifically, we are interested in:
#
# 1. Finding the number of tweets with a score above 10/10, versus tweets with a score under 10/10
# 1. Identify the tweets where more than one dog stage appears
# 1. Finding the number of top breed predictions from the image classifier, with a prediction confidence below 0.5
# **Count number of scores above and below 10/10**
((clean_enhanced_tweets_df['rating_numerator'] / clean_enhanced_tweets_df['rating_denominator']) > 1.0).sum(), \
((clean_enhanced_tweets_df['rating_numerator'] / clean_enhanced_tweets_df['rating_denominator']) <= 1.0).sum()
# **Show tweets with more than one dog stage in the tweet text**
stage_cols = ['doggo', 'floofer', 'pupper', 'puppo']
clean_enhanced_tweets_df.loc[clean_enhanced_tweets_df[stage_cols].sum(axis=1) > 1][['text'] + stage_cols]
# **Count tweets where the top scoring breed prediction is below 0.5**
dog_preds = clean_predictions_df.loc[(clean_predictions_df['pred_rank'] == 1) \
& clean_predictions_df['pred_is_dog']]
len(dog_preds[dog_preds['pred_confidence'] < 0.5].index)
# Now we are going to generate some visualisations:
#
# 1. First, based on the top image prediction, look at the frequency distribution for the top 10 breeds only, based on number of tweets
# 2. Now look at the frequency distribution for the top 10 breeds only, based on aggregate number of favorites
# **Breed prediction distribution by number of tweets**
dog_preds['pred_class'].value_counts(sort=True)[0:10].plot.pie()
# **Breed prediction distribution by number of favorites**
dog_preds.join(clean_enhanced_tweets_df['favorite_counts']).groupby(['pred_class']) \
.sum().sort_values(by='favorite_counts', ascending=False)[0:10]['favorite_counts'].plot.pie()
# ### Generate internal report
# Having cleaned the data, and generated data insights, we can now generate the internal documentation from this notebook's markdown cells.
#
# (you probably want to clear all output previous to the data insights output generated in the last section, and then SAVE the notebook)
# +
# # !jupyter nbconvert --no-input --to pdf wrangle_act.ipynb
# # !mv wrangle_act.pdf wrangle_report.pdf
# -
| wrangle_act.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
#
# This notebook accompanies the paper "Impact of non-parabolic electronic band structure on the optical, defect, and transport properties of photovoltaic materials". It reproduces some of the key results from the paper and includes supplementary information. It makes extensive use of the [effmass](http://github.com/lucydot/effmass) package which is available to download on github.
#
# The effmass package contains the [data files](https://github.com/lucydot/effmass/tree/master/paper/data) needed to reproduce all results in the paper. Here we focus upon a subset of that data to calculate:
# - Effective mass and non-parabolicity of CdTe
# - Burstein Moss shift and optical effective mass in MAPI
#
# In both cases we use data calculated using the HSE06 functional with spin-orbit coupling. However, the notebook can be easily adapted to explore other materials and levels of theory if required.
#
# ## Imports and preamble
# +
# plots displayed within notebook
# %matplotlib inline
# import scientific libraries
import math
import matplotlib.pyplot as plt
import numpy as np
# import modules from the effmass package
from effmass import inputs, analysis, extrema, outputs, dos, ev_to_hartree
# -
# ## Effective mass and non-parabolicity of CdTe
#
# ### Settings
# First we use the `inputs` module to create a `Settings` object. The `extrema_search_depth` attribute tells us how far from the CBM/VBM we would like to search for the bandstructure minima/maxima. The `energy_range` attribute sets the energy range for each band `Segment`.
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.25)
# ### Import bandstructure data
# We now use the `inputs` module to create a `Data` object which automatically imports the vasp data from the files specified. We manually specify how many k-points to ignore at the start of each file. These are the k-points which are included as part of the non-self-consistent bandstructure calculation, but which do not form part of the bandstructure itself.
data = inputs.Data("./data/CdTe/HSE06_SoC/OUTCAR","./data/CdTe/HSE06_SoC/PROCAR", ignore=216)
# ### Generate band segments
# We can now use our `Settings` and `Data` objects to generate a list of `Segment` objects using the `extrema` module.
segments = extrema.generate_segments(settings,data)
# Each Segment has a string method which gives the energy of the Segment extrema (referenced to the VBM) and the start- and end- points of the Segment in reciprocal space.
str(segments[-1])
# ### Visualise band segments
# We can visualise the `Segment` objects created using the `outputs` module. The plot is annotated with the `Segment`'s direction and segments argument index.
outputs.plot_segments(data,settings,segments)
outputs.plot_segments(data,settings,[segments[-1],segments[-3]])
# ### Calculate parabolic effective mass
# There are a number of methods associated with each `Segment` object. We can use these to calculate the different definitions of effective mass, assuming a parabolic dispersion $E= \frac{\hbar^2k^2}{2m^*}$.
segments[-1].five_point_leastsq_effmass()
segments[-1].finite_difference_effmass()
segments[-1].weighted_leastsq_effmass()
# We can use inbuilt documentation to find out more about a particular method.
# +
# analysis.Segment.weighted_leastsq_effmass?
# -
# ### Calculate parameters for the Kane dispersion
# The alpha parameter quantifies the extent of non-parabolicity in the kane dispersion $\frac{\hbar^2k^2}{2m^*_0} = E(1 + \alpha E) $
# where $m^*_0$ is the mass at the band edge ($E=0$).
segments[-1].alpha() # note that atomic units are used (hartree^-1)
segments[-1].kane_mass_band_edge()
# ### Summarise results
#
# The `print_results` function in the `outputs` module summarises the results for a segment.
outputs.print_results(segments[-1], data, settings)
# ## Burstein Moss shift and optical effective mass in MAPI
#
# ### Settings, Data and Segments
# As in the previous example we create a `Settings` object and `Data` object then use these to create a list of `Segment` objects.
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.75)
data = inputs.Data("./data/MAPI/HSE06_SoC/OUTCAR","./data/MAPI/HSE06_SoC/PROCAR",ignore=216)
segments = extrema.generate_segments(settings,data)
# ### Kane dispersion parameters
# To calculate the non-parabolic burstein-moss shift we need to know the electron alpha parameter and bandedge transport effective mass. First, let's see the segments we have generated
outputs.plot_segments(data,settings,segments)
# We want to calculate the kane dispersion parameters for `segments[-4]`, `segments[-5]` and `segments[-6]`. These parameters depend upon the `Settings.energy_range` attribute and the order of the polyfit used for calculating the transport mass. We can adjust the `energy_range` and polyfit order until we get a good fit to data.
# +
# energy_range=0.25, polyfit_order=6 for optimal fitting in 001 direction
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.25)
segments = extrema.generate_segments(settings, data)
outputs.print_results(segments[-4],data,settings,polyfit_order=6)
# +
# energy_range=0.5, polyfit_order=6 for optimal fitting in 101 direction
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.5)
segments = extrema.generate_segments(settings, data)
outputs.print_results(segments[-5],data,settings,polyfit_order=6)
# +
# energy_range=0.4, polyfit_order=4 for optimal fitting in 111 direction
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.4)
segments = extrema.generate_segments(settings, data)
outputs.print_results(segments[-6],data,settings,polyfit_order=4)
# -
# ### Burstein Moss shift
#
# The Burstein Moss shift is calculated using analytic expression $\Delta_{BM} =\frac{\hbar^2}{2m^*}(3\pi^2n_e)^{2/3}$ where the effective mass $m^*$ is constant (in the case of a parabolic dispersion) or takes the form $m^*(E) = m_0^*(1+2 \alpha E)$ (for a Kane dispersion).
# +
def burstein_moss_parabolic(mass,concentration):
# the expression comes from the fermi energy for a given concentration, assuming parabolic dispersion
# the fermi wavevector is simply gotten from considering volume of sphere in reciprocal space / volume of each eigenstate (factor 2 for spin)
concentration = concentration * ((5.29E-9)**3) # convert cm-3 --> bohr-3
return (((3*math.pi*math.pi*concentration)**(2/3))/(2*mass))/ev_to_hartree
def burstein_moss_kane(concentration,mass,alpha):
# this expression is an adaptation of the one above where the mass is now dependant upon the shift m_t = m_o(1+2 \alpha E)
concentration = concentration * ((5.29E-9)**3) # convert cm-3 --> bohr-3
return (analysis._solve_quadratic(2*alpha,1,[-((3*math.pi*math.pi*concentration)**(2/3)/(2*mass))])[0])/ev_to_hartree
# -
# We take a mean average of the alpha value across the three directions
average_alpha = (2.214+1.499+0.16)/(3*(ev_to_hartree)) # convert back to atomic units
# The geometric average of the mass is calculated for electrons and holes. Unlike silicon which has 6 equivalent minima and a degeneracy of 6, the minima here are between $\Gamma-R$ which has a multiplicity of 1. These average values are then combined into a reduced mass.
# +
def dos_average(m1,m2,m3,degeneracy=1):
# density of states mass which is used instead of conductivity effective mass as electron concentration populates entire 3D brillouin zone
# this perhaps needs a weighting factor to account for not mx,my,mz
return ((m1*m2*m3)**(1/3))*(degeneracy**(2/3))
def reduced_mass(m_e,m_h):
return 1/((1/m_e)+(1/m_h))
# effective mass at bandedge calculated from bandstructure
em1 = 0.19
em2 = 0.10
em3 = 0.18
hm1 = 0.23
hm2 = 0.10
hm3 = 0.12
average_mass=reduced_mass(dos_average(em1,em2,em3),dos_average(hm1,hm2,hm3))
# -
# Calculate the Burstein Moss shifts for concentration range $1 \times 10^{16}$ to $ 3 \times 10^{20}$
concentrations = np.logspace(16,np.log10(3E20),100)
parabolic_bandshift = [burstein_moss_parabolic(average_mass,x) for x in concentrations]
kane_bandshift = [burstein_moss_kane(x,average_mass,average_alpha) for x in concentrations]
# ### Import density of states data
# We can compare these values to density of states data. We start by parsing the DOSCAR file.
data.parse_DOSCAR("./data/MAPI/HSE06_SoC/DOSCAR")
# We can use this data and the `dos` module to calculate the band filling level for a given concentration. We also need to supply the volume of the unit cell in $\unicode{x212B} ^3$.
volume = 251.13
dos_bandshift = [dos.electron_fill_level(data, volume, x, dos.find_dos_CBM_index(data))for x in concentrations]
# ### Compare results
#
# Let's plot the three bandshift results:
fig = plt.figure()
plt.plot(concentrations,parabolic_bandshift, "-",label="parabolic",color="black")
plt.plot(concentrations,kane_bandshift,"--",label="Kane",color="black")
plt.plot(concentrations,dos_bandshift, ":",label="DFT DOS",color="black")
plt.legend(prop={'size': 8},loc=2)
plt.xlabel(r"concentration cm$^{-3}$")
plt.ylabel("E (eV)")
plt.tight_layout()
plt.savefig("burstein_moss_MAPI_hybrid_SoC.pdf")
plt.xlim([0,3E20])
plt.ylim([0,2.5])
# ### Calculate optical effective mass as a function of concentration
# Our results show that the Kane dispersion is a good approximation to the density of states data (which makes no assumptions about the band dispersions). We can calculate an optical effective mass:
# $\begin{equation}
# \frac{1}{m_o} = \frac{\sum_{l} \int f(E_k(k),T) \frac{\delta^2 E_k(k)}{\delta k^2} dk}{\sum_{l} \int f(E_k(k),T) dk}
# \end{equation}$
#
# with $E(K)$ set to the Kane dispersion. The Fermi level in the Fermi-Dirac distribution $f(E_k(k),T)$ is set to the burstein-moss shift as calculated using the Kane dispersion.
# +
settings = inputs.Settings(extrema_search_depth=0.075, energy_range=0.25) # kane dispersion valid up to energy_range 0.25eV
segments = extrema.generate_segments(settings, data)
concentrations = np.logspace(16,np.log10(2E19),100) # if we set the concentration higher we exceed energies where the kane dispersion is valid and receive a warning
optical_mass_111 = [segments[-6].optical_effmass_kane_dispersion(fermi_level=data.CBM+burstein_moss_kane(x,average_mass,average_alpha),alpha=0.16/ev_to_hartree,mass_bandedge=0.155,upper_limit=0.1) for x in concentrations]
optical_mass_110 = [segments[-5].optical_effmass_kane_dispersion(fermi_level=data.CBM+burstein_moss_kane(x,average_mass,average_alpha),alpha=1.499/ev_to_hartree,mass_bandedge=0.098,upper_limit=0.1) for x in concentrations]
optical_mass_100 = [segments[-4].optical_effmass_kane_dispersion(fermi_level=data.CBM+burstein_moss_kane(x,average_mass,average_alpha),alpha=2.214/ev_to_hartree,mass_bandedge=0.156,upper_limit=0.1) for x in concentrations]
## Plot results
fig,ax1 = plt.subplots()
ax1.plot(np.log10(concentrations),optical_mass_111,":",label="(111) ",color="black")
ax1.plot(np.log10(concentrations),optical_mass_110,"-.",label="(110) ",color="black")
ax1.plot(np.log10(concentrations),optical_mass_100,"--",label="(100) ",color="black")
ax1.set_xticks([16,17,18,19])
ax1.set_xticklabels([r"$10^{16}$",r"$10^{17}$",r"$10^{18}$",r"$10^{19}$"])
ax1.set_xlim([16,np.log10(2E19)])
ax1.set_ylim([0,0.4])
ax1.set_xlabel(r"concentration (cm$^{-3}$)")
ax1.set_ylabel(r"optical $\frac{m^*}{m_e}$")
# -
| paper/notebook.ipynb |