code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 32-bit
# name: python3
# ---
# +
from Classes import *
fm = clsFormulasMath.Bascara_1('R$90')
print(fm.calc_bas())
| calculos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import thermostat; thermostat.get_version()
# +
import sys
import os
import warnings
import logging
from os.path import expanduser
from thermostat.importers import from_csv
from thermostat.exporters import metrics_to_csv, certification_to_csv
from thermostat.stats import compute_summary_statistics
from thermostat.stats import summary_statistics_to_csv
from thermostat.multiple import multiple_thermostat_calculate_epa_field_savings_metrics
logger = logging.getLogger('epathermostat')
# Set this to 'DEBUG' for more logging messages (default: WARNING)
# See `multi_thermostat_tutorial.py` for how to
# use a logging configuration file which logs to console / file
logger.setLevel(logging.WARNING)
data_dir = os.path.join(expanduser("~"), "Downloads")
# -
# dev specific
data_dir = os.path.join(os.path.join("/", *thermostat.__file__.split('/')[:6]), "tests", "data", "single_stage")
data_dir = os.path.join(os.path.curdir, "..", "tests", "data", "single_stage")
metadata_filename = os.path.join(data_dir, "metadata.csv")
# verbose=True will override logging to display the imported thermostats
# Set verbose to "False" to use the logging level instead
thermostats = from_csv(metadata_filename, verbose=True)
data_dir = os.path.join(expanduser("~"), "Downloads")
# Use this for multi-processing thermostats
metrics = multiple_thermostat_calculate_epa_field_savings_metrics(thermostats)
# Use this to process each thermostat one-at-a-time
'''
metrics = []
ts = []
for thermostat_ in thermostats:
outputs = thermostat_.calculate_epa_field_savings_metrics()
metrics.extend(outputs)
ts.append(thermostat_)
'''
output_filename = os.path.join(data_dir, "thermostat_example_output.csv")
metrics_df = metrics_to_csv(metrics, output_filename)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
# uses the metrics_df created in the quickstart above.
stats = compute_summary_statistics(metrics_df)
stats_advanced = compute_summary_statistics(metrics_df, advanced_filtering=True)
# +
product_id = "test-product"
certification_filepath = os.path.join(data_dir, "thermostat_example_certification.csv")
certification_to_csv(stats, certification_filepath, product_id)
stats_filepath = os.path.join(data_dir, "thermostat_example_stats.csv")
stats_df = summary_statistics_to_csv(stats, stats_filepath, product_id)
stats_advanced_filepath = os.path.join(data_dir, "thermostat_example_stats_advanced.csv")
stats_advanced_df = summary_statistics_to_csv(stats_advanced, stats_advanced_filepath, product_id)
# -
| scripts/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 002
# # Even Fibonacci numbers
#
# Even Fibonacci numbers
#
# Each new term in the Fibonacci sequence is generated by adding the previous two terms.
# By starting with 1 and 2, the first 10 terms will be:
#
# 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, ...
#
# By considering the terms in the Fibonacci sequence whose values do not exceed four million,
# find the sum of the even-valued terms.
#
# +
from scripts import myfunc
fib_seq = [1, 2]
max_seq = 4*1e6
for i in range(1000):
x = fib_seq[-1]+fib_seq[-2]
if x <= max_seq:
fib_seq.append(x)
else:
print('Reached to %d' %(int(max_seq)))
break
print(fib_seq[-1])
fib_even = [i for i in fib_seq if i%2 == 0]
print(sum(fib_even))
# -
| notebooks/problem_solved/problem_002_Even_Fibonacci_numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Experimental Design Simulation
#
# This simulation parallels our 50-cycle MAGE experiment and allows exploring the relationship among experimental design parameters including number of oligos tested and number of clones sampled for genotyping. The simulation also investigated different number and effect-size distributions of causal mutations. For a given combination of parameters, we sample a distribution of underlying mutation effects (capped at a total fitness effect of 50%, comparable to the C321.∆A context) and then performed, in silico, iterations of MAGE separated by competitive expansion and bottlenecking of the population. We sample clonal genotypes from this simulated population and calculate phenotypes using the underlying mutation effects. We then perform predictive modeling with the simulated genotype-phenotype data and evaluate precision and recall relative to the underlying mutation effects. We also compare our linear modeling strategy to alternative modeling strategies of quantitative GWAS (a separate linear model for each variant) or using enrichment of mutations in the final population. We made simplifying assumptions of no de novo mutations, no epistatic interactions among mutations, no measurement noise, and equal recombination efficiency for all mutations.
# + deletable=true editable=true
import os
import random
import sys
import time
import ipyparallel as ipp
from IPython.display import display
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
sys.path.append('../')
from common_code import simulation_util
from common_code.simulation_util import SimulationParams
# + [markdown] deletable=true editable=true
# ## Implementation
#
# We implement each component of the simulation as a separate function and perform a small sanity check / plot for the different steps below. The sanity checks build on the previous data.
#
# NOTE: Most of the functions defintions have been extracted to common_code/simulation_util.py.
# + [markdown] deletable=true editable=true
# ### Generate SNP effects
#
# We'll assume some number of SNPs with an effect distributed according to a Gaussian, and then scaled so that the total effect comes out to < 50% total fitness defect.
#
# We use multiplicative effects, e.g. a single mutant with effect 0.9 has 10% reduced doubling time.
# + deletable=true editable=true
# Sanity check
test_non_zero_snp_effects = simulation_util.sample_effects_by_power_law(30, debug_plot=True)
# + deletable=true editable=true
# Sanity check.
test_snp_effects = simulation_util.generate_snp_effects(
simulation_util.DEFAULT_SNPS_CONSIDERED, simulation_util.DEFAULT_SNPS_WITH_EFFECT)
non_trivial_snp_effects = sorted([e for e in test_snp_effects if e != 1], reverse=True)
print 'Total Fitness', np.prod(non_trivial_snp_effects)
plt.figure()
plt.title('Sanity Check | Fitness effects %d/%d SNPs' % (
simulation_util.DEFAULT_SNPS_WITH_EFFECT, simulation_util.DEFAULT_SNPS_CONSIDERED))
plt.bar(range(len(non_trivial_snp_effects)), non_trivial_snp_effects)
plt.show()
# + [markdown] deletable=true editable=true
# ### Update population with MAGE
#
# We'll need a function that performs a cycle of MAGE. We'll simplify this as updating some fraction of the population according to MAGE_EFFICIENCY, where that fraction of the population gets an additional mutation added on top.
# + deletable=true editable=true
# Here we do a sanity check of the above by running 50 cycles of MAGE
# with 100 SNPs observing the resulting distribution of mutations.
# Note that we ignore selection based on fitness in between rounds.
# Initial population.
test_population = np.zeros(
(simulation_util.DEFAULT_POPULATION_SIZE,
simulation_util.DEFAULT_SNPS_CONSIDERED), dtype=np.bool)
for cycle in range(simulation_util.MAGE_CYCLES):
test_population = simulation_util.update_population_with_mage_mutations(test_population)
plt.figure()
plt.title('Sanity check | Expected mutation distribution after 50 cycles (no selection)')
plt.hist(test_population.sum(axis=1))
plt.show()
# + [markdown] deletable=true editable=true
# ### Calculate doubling times from genotype.
#
# Now we need a way to generate doubling times for a population. This will be a function of the SNP effects with a bit of extra random noise.
# + deletable=true editable=true
# Sanity check.
test_doubling_times = simulation_util.generate_doubling_times(
test_population, test_snp_effects)
plt.figure()
plt.title('Sanity check | Doubling times distribution')
plt.bar(range(len(test_doubling_times)), sorted(test_doubling_times, reverse=True))
plt.show()
# + [markdown] deletable=true editable=true
# ## Apply selection between MAGE cycles
#
# For the model to actually work, we need to add some form of selection, so in between each MAGE cycle we'll do an expansion and pruning step that is weighted by the fitness effects.
#
# We'll do this by allowing each genotype to double at a rate relative to the others proportional to its fitness.
# + deletable=true editable=true
# Sanity check.
# NOTE: We are running this on the final unselected population
# to see what would have been selected. See simulation with
# default params below for real end-to-end.
apply_selection_result = simulation_util.apply_selection_pressure(
test_population, test_snp_effects)
subsampled_clones = apply_selection_result['subsampled_clone_ids']
plt.figure()
plt.hist(subsampled_clones, bins=100)
plt.show()
apply_selection_result['metadata_df'].sort_values('growth_rates', ascending=False)[:10]
# + [markdown] deletable=true editable=true
# ### Run linear modeling and evaluate
# + deletable=true editable=true
# Sanity check.
test_lm_result = simulation_util.run_linear_modeling(
test_population, test_doubling_times)
simulation_data = {
'snp_effect': test_snp_effects,
}
lm_evaluation_results = simulation_util.evaluate_modeling_result(
simulation_data, test_lm_result)
print 'Pearson: %f, p-value: %f' % (
lm_evaluation_results['pearson_r'],
lm_evaluation_results['p_value'])
lm_eval_results_df = lm_evaluation_results['results_df']
lm_eval_results_df[lm_eval_results_df['snp_effect'] != 1]
# + [markdown] deletable=true editable=true
# ### Compare to GWAS
#
# Quantitative GWAS is effectively running a linear model with a single SNP at a time. See `model_fitting#single_snp_lienar_modeling`.
# + deletable=true editable=true
# Sanity check.
gwas_results_df = simulation_util.run_gwas(test_population, test_doubling_times)
gwas_eval_results = simulation_util.evaluate_gwas_result(
gwas_results_df, lm_eval_results_df, show_plot=True)
gwas_eval_results_df = gwas_eval_results['results_df']
print 'GWAS Pearson: %f, p-value: %f' % (
gwas_eval_results['pearson_r'],
gwas_eval_results['p_value'])
print 'GWAS results | SNPs with effect or p < 0.05'
gwas_results_of_interest_df = gwas_eval_results_df[
(gwas_eval_results_df['snp_effect'] < 1) |
(gwas_eval_results_df['gwas_p'] < 0.05)]
gwas_results_of_interest_df = gwas_results_of_interest_df[[
'snp_effect', 'gwas_coef', 'gwas_p']].sort_values('snp_effect')
display(gwas_results_of_interest_df)
# + [markdown] deletable=true editable=true
# Here we define some functions for calculating metrics like true positives, false positives, recall, etc.
# + deletable=true editable=true
print 'Linear modeling'
print simulation_util.calculate_modeling_metrics(lm_eval_results_df, 'linear_model_coef', 'lm_')
print '\nGWAS'
print simulation_util.calculate_modeling_metrics(gwas_eval_results_df, 'gwas_coef', 'gwas_')
# + [markdown] deletable=true editable=true
# Slightly different metrics for enrichment.
# + deletable=true editable=true
# Sanity check
test_simulation_data = {
'sim_params': SimulationParams(),
'final_population': test_population,
'final_doubling_times': test_doubling_times,
'snp_effect': test_snp_effects,
}
test_enrichment_result_df = simulation_util.run_enrichment_analysis(test_simulation_data)
simulation_util.calculate_enrichment_metrics(test_enrichment_result_df)
# + [markdown] deletable=true editable=true
# ## Run simulations
#
# First, test our re-useable function for running simulations.
# + deletable=true editable=true
# Sanity checks.
test_sim_params = SimulationParams()
test_sim_params.population_size = 100
test_run_simulation_result = simulation_util.run_simulation(
simulation_params=test_sim_params)
assert min(test_run_simulation_result['wgs_samples_mage_cycle_list']) == 1
assert max(test_run_simulation_result['wgs_samples_mage_cycle_list']) == 50
test_sim_params = SimulationParams()
test_sim_params.population_size = 100
test_sim_params.num_samples = 5
test_run_simulation_result = simulation_util.run_simulation(
simulation_params=test_sim_params)
np.testing.assert_array_equal(
np.array([10, 20, 30, 40, 50]),
np.array(test_run_simulation_result['wgs_samples_mage_cycle_list']))
# + [markdown] deletable=true editable=true
# ### Simulation with our default experiment parameters.
# + deletable=true editable=true
sim_params = SimulationParams()
default_simulation_result = simulation_util.run_simulation(
simulation_params=sim_params)
simulation_util.visualize_simulation_result(default_simulation_result)
# Perform linear modeling.
print '>>> Linear Model Results'
test_lm_result = simulation_util.run_linear_modeling(
default_simulation_result['wgs_samples'],
default_simulation_result['wgs_samples_doubling_times'],
repeats=10)
lm_eval_results = simulation_util.evaluate_modeling_result(
default_simulation_result, test_lm_result)
lm_eval_results_df = lm_eval_results['results_df']
print 'Pearson: %f, p-value: %f' % (
lm_eval_results['pearson_r'], lm_eval_results['p_value'])
display(lm_eval_results_df[lm_eval_results_df['snp_effect'] != 1])
# Compare to GWAS
print '>>> GWAS results'
gwas_results_df = simulation_util.run_gwas(
default_simulation_result['wgs_samples'],
default_simulation_result['wgs_samples_doubling_times'])
gwas_eval_results = simulation_util.evaluate_gwas_result(
gwas_results_df, lm_eval_results_df, show_plot=True)
gwas_eval_results_df = gwas_eval_results['results_df']
print 'GWAS Pearson: %f, p-value: %f' % (
gwas_eval_results['pearson_r'],
gwas_eval_results['p_value'])
print 'GWAS results | SNPs with effect or p < 0.05'
gwas_results_of_interest_df = gwas_eval_results_df[
(gwas_eval_results_df['snp_effect'] < 1) |
(gwas_eval_results_df['gwas_p'] < 0.05)]
gwas_results_of_interest_df = gwas_results_of_interest_df[[
'snp_effect', 'gwas_coef', 'gwas_p']].sort_values('snp_effect')
display(gwas_results_of_interest_df)
print 'GWAS results | SNPs with effect or p < 0.05'
gwas_results_of_interest_df = gwas_eval_results_df[
(gwas_eval_results_df['snp_effect'] < 1) |
(gwas_eval_results_df['gwas_p'] < 0.05)]
gwas_results_of_interest_df = gwas_results_of_interest_df[[
'snp_effect', 'gwas_coef', 'gwas_p']].sort_values('snp_effect')
display(gwas_results_of_interest_df)
# + [markdown] deletable=true editable=true
# ### Compare with same SNP effects against no selection.
# + deletable=true editable=true
sim_params = SimulationParams()
sim_params.num_snps_with_effect = 10
simulation_result_with_selection = simulation_util.run_simulation(
simulation_params=sim_params)
simulation_result_no_selection = simulation_util.run_simulation(
simulation_params=sim_params,
snp_effects=simulation_result_with_selection['snp_effect'],
should_apply_selection_pressure=False)
print '>>>With Selection'
simulation_util.visualize_simulation_result(simulation_result_with_selection)
# Perform linear modeling.
test_lm_result = simulation_util.run_linear_modeling(
simulation_result_with_selection['wgs_samples'],
simulation_result_with_selection['wgs_samples_doubling_times'],
repeats=10)
# Evaluate results.
evaluation_results = simulation_util.evaluate_modeling_result(
simulation_result_with_selection, test_lm_result)
print 'Pearson: %f, p-value: %f' % (evaluation_results['pearson_r'], evaluation_results['p_value'])
eval_results_df = evaluation_results['results_df']
display(eval_results_df[eval_results_df['snp_effect'] != 1])
with_selection_modeling_metrics = {
'lm_pearson_r': evaluation_results['pearson_r'],
'lm_pearson_p': evaluation_results['p_value'],
}
with_selection_modeling_metrics.update(
simulation_util.calculate_modeling_metrics(
eval_results_df, 'linear_model_coef',
results_prefix='lm_'))
print '>>>No Selection'
simulation_util.visualize_simulation_result(simulation_result_no_selection)
test_lm_result = simulation_util.run_linear_modeling(
simulation_result_no_selection['wgs_samples'],
simulation_result_no_selection['wgs_samples_doubling_times'],
max_iter=300, # This converges poorly so manually limit here for now.
repeats=1) # likewise
# Evaluate results.
evaluation_results = simulation_util.evaluate_modeling_result(simulation_result_no_selection, test_lm_result)
print 'Pearson: %f, p-value: %f' % (evaluation_results['pearson_r'], evaluation_results['p_value'])
eval_results_df = evaluation_results['results_df']
display(eval_results_df[eval_results_df['snp_effect'] != 1])
no_selection_modeling_metrics = {
'lm_pearson_r': evaluation_results['pearson_r'],
'lm_pearson_p': evaluation_results['p_value'],
}
no_selection_modeling_metrics.update(
simulation_util.calculate_modeling_metrics(
eval_results_df, 'linear_model_coef',
results_prefix='lm_'))
# + [markdown] deletable=true editable=true
# Also compare linear modeling result if we had just taken all samples from last timepoint, instead of sampling across different MAGE cycles.
# + deletable=true editable=true
# Sub-sample final population.
final_timepoint_subsample = np.zeros(
(simulation_util.DEFAULT_NUM_SAMPLES, simulation_util.DEFAULT_SNPS_CONSIDERED), dtype=np.bool)
random_indeces_from_final_population = np.random.choice(
range(len(simulation_result_with_selection['final_population'])),
size=simulation_util.DEFAULT_NUM_SAMPLES)
final_timepoint_doubling_times = []
for i, random_idx in enumerate(random_indeces_from_final_population):
final_timepoint_subsample[i,:] = (
simulation_result_with_selection['final_population'][random_idx,:])
final_timepoint_doubling_times.append(
simulation_result_with_selection['final_doubling_times'][random_idx])
assert final_timepoint_subsample.shape == (
simulation_util.DEFAULT_NUM_SAMPLES,
simulation_util.DEFAULT_SNPS_CONSIDERED)
assert len(final_timepoint_doubling_times) == simulation_util.DEFAULT_NUM_SAMPLES
# Run linear modeling.
test_lm_result = simulation_util.run_linear_modeling(
final_timepoint_subsample,
final_timepoint_doubling_times,
repeats=1,
max_iter=300)
# Evaluate results.
evaluation_results = simulation_util.evaluate_modeling_result(
simulation_result_with_selection, test_lm_result)
print 'Pearson: %f, p-value: %f' % (evaluation_results['pearson_r'], evaluation_results['p_value'])
eval_results_df = evaluation_results['results_df']
display(eval_results_df[eval_results_df['snp_effect'] != 1])
final_timepoint_modeling_metrics = {
'lm_pearson_r': evaluation_results['pearson_r'],
'lm_pearson_p': evaluation_results['p_value'],
}
final_timepoint_modeling_metrics.update(
simulation_util.calculate_modeling_metrics(
eval_results_df, 'linear_model_coef',
results_prefix='lm_'))
plt.figure(figsize=(10, 4))
# WGS mutation distribution
plt.subplot(1, 2, 1)
plt.title('WGS samples from final population only | Mutations')
plt.hist(final_timepoint_subsample.sum(axis=1))
# WGS doubling times
plt.subplot(1, 2, 2)
plt.title('WGS samples from final population only | Doubling times')
plt.bar(
range(len(final_timepoint_doubling_times)),
sorted(final_timepoint_doubling_times, reverse=True),
edgecolor="none")
plt.tight_layout()
plt.show()
# + [markdown] deletable=true editable=true
# Output for plotting in R.
# + deletable=true editable=true
IGNORE_KEYS = ['sim_params', 'final_population']
# 4a
supp_fig4a_data_dir = 'results/supp_fig4a_data'
for key in simulation_result_with_selection.keys():
if key in IGNORE_KEYS:
continue
np.savetxt(
os.path.join(supp_fig4a_data_dir, key + '.csv'),
simulation_result_with_selection[key],
delimiter=',')
pd.DataFrame([with_selection_modeling_metrics]).to_csv(
os.path.join(supp_fig4a_data_dir, 'modeling_metrics.csv'),
index=False)
# 4b
supp_fig4b_data_dir = 'results/supp_fig4b_data'
np.savetxt(
os.path.join(supp_fig4b_data_dir, 'wgs_samples.csv'),
final_timepoint_subsample,
delimiter=',')
np.savetxt(
os.path.join(supp_fig4b_data_dir, 'wgs_samples_doubling_times.csv'),
final_timepoint_doubling_times,
delimiter=',')
pd.DataFrame([final_timepoint_modeling_metrics]).to_csv(
os.path.join(supp_fig4b_data_dir, 'modeling_metrics.csv'),
index=False)
# 4c
supp_fig4c_data_dir = 'results/supp_fig4c_data'
for key in simulation_result_no_selection.keys():
if key in IGNORE_KEYS:
continue
np.savetxt(
os.path.join(supp_fig4c_data_dir, key + '.csv'),
simulation_result_no_selection[key],
delimiter=',')
pd.DataFrame([no_selection_modeling_metrics]).to_csv(
os.path.join(supp_fig4c_data_dir, 'modeling_metrics.csv'),
index=False)
# + [markdown] deletable=true editable=true
# Placeholder plot for Fig S4a-c for co-authors review.
# + deletable=true editable=true
plt.figure(figsize=(20, 8))
final_timepoint_doubling_times
final_timepoint_subsample
### S4A
plt.subplot(2, 3, 1)
plt.title('100 samples | default parameters')
plt.bar(
simulation_result_with_selection['wgs_samples_mage_cycle_list'],
simulation_result_with_selection['wgs_samples_doubling_times'],
)
plt.ylabel("Relative doubling time")
plt.xlim((0, 50))
plt.ylim((0, 1.0))
plt.subplot(2, 3, 4)
plt.plot(
simulation_result_with_selection['wgs_samples_mage_cycle_list'],
np.sum(simulation_result_with_selection['wgs_samples'], axis=1),
'.')
plt.ylabel("# Mutations")
plt.xlabel('MAGE cycle')
plt.ylim((0, 10))
### S4B
plt.subplot(2, 3, 2)
plt.title('100 samples | final time point')
plt.bar(
range(len(final_timepoint_doubling_times)),
final_timepoint_doubling_times,
)
plt.ylim((0, 1.0))
plt.subplot(2, 3, 5)
plt.plot(
range(len(final_timepoint_subsample)),
np.sum(final_timepoint_subsample, axis=1),
'.'
)
plt.xlabel('Sample #')
plt.ylim((0, 10))
### S4C
plt.subplot(2, 3, 3)
plt.title('100 samples | no selection')
plt.bar(
simulation_result_no_selection['wgs_samples_mage_cycle_list'],
simulation_result_no_selection['wgs_samples_doubling_times'],
)
plt.xlim((0, 50))
plt.ylim((0, 1.0))
plt.subplot(2, 3, 6)
plt.plot(
simulation_result_no_selection['wgs_samples_mage_cycle_list'],
np.sum(simulation_result_no_selection['wgs_samples'], axis=1),
'.')
plt.xlabel('MAGE cycle')
plt.ylim((0, 10))
plt.tight_layout()
plt.rcParams.update({'font.size': 18})
plt.savefig('results/supp_fig_4abc_ipynb_draft.png', dpi=300)
plt.show()
# + [markdown] deletable=true editable=true
# Also look at what using enrichment at final timepoint would give.
# + deletable=true editable=true
final_timepoint_enrichment_with_selection_df = pd.DataFrame({
'snp_effect': simulation_result_with_selection['snp_effect'],
'enrichment_count': final_timepoint_subsample.sum(axis=0)
})
display(final_timepoint_enrichment_with_selection_df.sort_values(
'enrichment_count', ascending=False)[:10])
# + [markdown] deletable=true editable=true
# To call a SNP by enrichment, let's say its enrichment has to be above the average.
# + deletable=true editable=true
simulation_util.calculate_enrichment_metrics(final_timepoint_enrichment_with_selection_df)
# + [markdown] deletable=true editable=true
# This does decently well at our parameters, but misses at least one and gives at least one false positive. This might be an okay strategy at our design parameters, but we should show below that this is not as good with other design parameters.
# + [markdown] deletable=true editable=true
# ## Perform simulations across different parameters and record performance
# + [markdown] deletable=true editable=true
# We'll use ipyparallel to parallelize. To start a local cluster, run this in a terminal:
#
# ipcluster start -n <num_cores>
# + deletable=true editable=true
ipp_client = ipp.Client()
# + [markdown] deletable=true editable=true
# Use load-balanced view which distributes jobs as you would expect.
# + deletable=true editable=true
lbv = ipp_client.load_balanced_view()
# + [markdown] deletable=true editable=true
# **With more strains tested, would it be possible to identify more variants that affect fitness?**
# + deletable=true editable=true
# Clones sequenced.
# num_samples_range = [24, 48, 72, 96, 144, 192, 240, 288, 336, 384] # For manuscript
num_samples_range = [24, 48] # For test
print '\nnum_samples_range'
print num_samples_range
# Total oligos.
# num_snps_considered_range = (
# list(range(10, 100, 10)) +
# list(range(100, 600, 100)) +
# list(range(600, 1001, 200))
# ) # For manuscript
num_snps_considered_range = [10, 20] # For test
print '\nnum_snps_considered'
print num_snps_considered_range
# Causal SNPs.
# Bounded in the foor loop to be less than num_snps_considered.
# num_snps_with_effect_range = [5] + list(range(10, 101, 10)) # For manuscript
num_snps_with_effect_range = range(5, 11, 1) # For test
print '\nnum_snps_with_effect_range'
print num_snps_with_effect_range
# NUM_SNPS_SAMPLES_GRID_REPLICATES = 10 # For manuscript
NUM_SNPS_SAMPLES_GRID_REPLICATES = 1 # For test
# NUM_LM_REPEATS = 10 # For manuscript
NUM_LM_REPEATS = 1 # For test
# Estimated time
print 'Estimated simulations < %d' % (
len(num_samples_range) * len(num_snps_considered_range) *
len(num_snps_with_effect_range) * NUM_SNPS_SAMPLES_GRID_REPLICATES)
# + deletable=true editable=true
more_strains_simulation_async_results = []
for num_snps_considered in num_snps_considered_range:
for num_samples in num_samples_range:
for num_snps_with_effect in num_snps_with_effect_range:
if num_snps_with_effect >= num_snps_considered:
continue
for rep in range(1, NUM_SNPS_SAMPLES_GRID_REPLICATES + 1):
sim_params = simulation_util.SimulationParams()
sim_params.num_snps_with_effect = num_snps_with_effect
sim_params.num_samples = num_samples
sim_params.num_snps_considered = num_snps_considered
more_strains_simulation_async_results.append(
lbv.apply_async(
simulation_util.run_simulation_with_params,
sim_params, rep, repeats=NUM_LM_REPEATS))
print 'Started %d jobs' % len(more_strains_simulation_async_results)
# Loop to write results that are finished at this point.
while True:
more_strains_simulation_result = []
for ar in more_strains_simulation_async_results:
if ar.ready():
more_strains_simulation_result.append(ar.get())
# Write result so far.
more_strains_simulation_result_df = pd.DataFrame(
more_strains_simulation_result)
if len(more_strains_simulation_result_df):
more_strains_simulation_result_df = more_strains_simulation_result_df[
simulation_util.SIM_RESULT_KEY_ORDER]
more_strains_simulation_result_df.to_csv(
'results/sim_results_num_strains_vs_num_snps_identified_small_test.csv',
index=False)
# Check if all ready.
if len(more_strains_simulation_result) == len(more_strains_simulation_async_results):
break
# Delay 30 seconds until next check.
time.sleep(30)
more_strains_simulation_result_df[:3]
| experiment_design_simulations/simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introdução a Deep Learning
# Deep Learning têm recebido [bastante atenção nos últimos anos](https://trends.google.com/trends/explore?date=all&q=Deep%20Learning), tanto no campo da computação como na mídia em geral.
#
# Técnicas de Deep Learning (DL) se tornaram o estado da arte em várias tarefas de Inteligência Artificial e mudado vários grandes campos da área, na visão computacional (classificação de imagens, segmentação de imagens), NLP (tradução, classificação de textos), reinforcement learning (agentes capazes de jogar jogos complexos como Go, Atari e DOTA).
#
# Esse impacto não fica só na teoria, DL está mudando o campo da Medicina, Ciência, Matemática, Física. Artigos de Deep Learning são rotinamente publicados em grandes revistas científicas como: Nature, Science e JAMA.
#
# Nesta seção iremos discutir:
#
# * O que é Deep Learning?
# * Eu realmente deveria me importar com DL ou é só [hype](https://pt.wikipedia.org/wiki/Hype)?
# * Por que eu como programador(a) deveria me importar?
# ## DL = Muitos Dados + Redes Neurais + Otimização
#
# ### Inteligência Artificial $\supseteq$ Machine Learning $\supseteq$ Deep Learning
#
# 
# IA (Inteligência Artificial), ML (Machine Learning) e DL (Deep Learning) são utilizados na mídia quase como sinônimos.
#
# **Inteligência Artificial** se refere ao campo da ciência da computação que se preocupa com o estudo de máquinas para realização de atividades ditas inteligentes. Esse é um grande campo com diferentes "escolas"/"ramificações". Inteligência é um conceito muito amplo e bastante subjetivo.
#
# Uma grande pergunta é **como** construir máquinas inteligentes?
#
# **Machine Learning** se refere ao subcampo da IA que busca obter máquinas inteligentes através da extração de estrutura e inteligência (padrões) de dados (experiência).
#
# > "Um computador aprende a partir da experiência E com respeito a alguma tarefa T e alguma medida de performance P, se sua performance em T melhora com sua experiência E" - [<NAME> (1998)](http://www.cs.cmu.edu/~tom/)
#
# 
#
# A ideia de ML é que, para realizar uma tarefa, nós (programadoras(es)) não implementemos as regras que definem as saídas para uma dada entrada. Imagine que queremos traduzir **qualquer** frase do inglês para português, quais regras iremos utilizar? Listar todas as traduções e regras possíveis é não trivial.
#
# > *“A complexidade de programas de computadores convencionais está no código (programas que as pessoas escrevem). Em Machine Learning, algoritmos são em princípio simples e a complexidade (estrutura) está nos dados. Existe uma maneira de aprender essa estrutura automaticamente? Esse é o princípio fundamental de Machine Learning.”- [<NAME>](https://en.wikipedia.org/wiki/Andrew_Ng)*
#
# Ao invés de listar as regras e funcionamento do sistema podemos utilizar um **modelo de Machine Learning**, e mostrar para o modelo vários exemplos de frases em inglês traduzidas para português. Padrões são extraídos dos dados e utilizados para que o modelo defina como tomar suas próprias decisões de modo a otimizar a tradução.
#
# Modelos de Machine Learning extraem padrões dos dados, e claro são extremamente dependentes da qualidade dos dados para sucesso em suas tarefas, mas também depende de **como os dados são representados**. Por exemplo, como um modelo de ML deve representar uma frase?
#
# 
#
#
# Seria muito bom não ter que se preocupar com a representação dos dados... Tipo, dar para o modelo uma representação bastante simplificada e ele que se preocupe em encontrar algum sentindo nesse dado **bruto**...
#
# Seria e é possível!! Essa abordagem é conhecida como **Representation Learning**, o modelo deve ser capaz não só de resolver a tarefa (exemplo: tradução), mas também de encontrar representações úteis para os dados de modo a resolver essa tarefa. Afinal, a gente não sabe explicar como resolve o problema, provavelmente também não sabemos qual a melhor representação para tal.
#
# **Deep Learning é um subcampo de ML que utiliza representation learning definindo representações mais complexas a partir de outras representações mais simples.**
#
# 
#
# O nome **Deep** vem das múltiplas representações (**camadas**) que utilizamos para construir os modelos. Então, de maneira geral, quando pensamos em DL pensamos em um modelo com váááárias camadas, e também precisamos de muuuuitos dados. Métodos de DL tem fome de dados. Quer boas representações? Me dê vários exemplos de dados; Quer que eu saiba traduzir bem? Me dê vários exemplos de traduções de boa qualidades que eu consigo extrair os padrões e aprender umas representações bem bacanas e ser um excelente tradutor!
#
# **Importante**: se modelos de DL aprendem com os padrões presentes nos dados, significa que eles podem aprender vieses e preconceitos presente nos dados.
#
#
# # Eu realmente deveria me importar com DL ou é só [hype](https://pt.wikipedia.org/wiki/Hype)?
# ### Popularização de DL
#
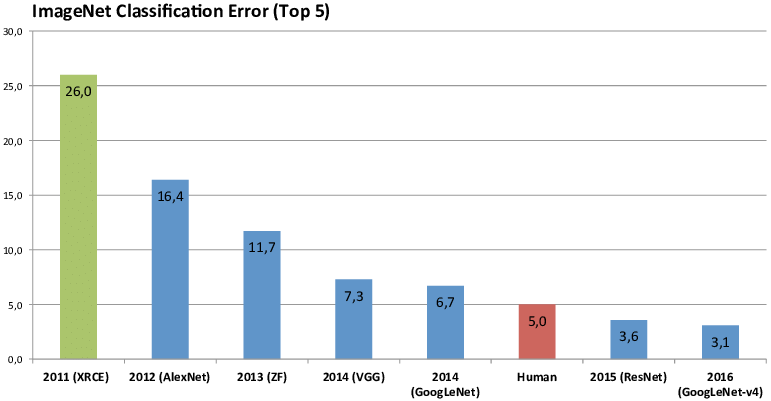
# O termo Deep Learning começou a se popularizar aproximadamente em 2012 quando técnicas de DL foram aplicadas com sucesso para a tarefa de classificação de imagens num desafio chamado "Imaginet Challenge" que consiste em classificar corretamente imagens em 1000 diferentes categorias.
#
# 
#
# Desde então modelos de DL são a melhor coisa para essa tarefa e só ficam melhor. Na imagem acima é mostrado o erro do vencedor do desafio em cada ano. Sendo as barras azuis modelos de DL.
#
# ### Mas... redes neurais (coração do DL) existem desde 1930, por que só agora?
#
# Dois grandes fatores contribuiram para o sucesso de métodos de DL e sua popularização:
#
# 1. Muitos dados disponívels
# 2. Hardware disponível
#
# Com esses fatores foi possível explorar redes neurais com muito mais camadas e novas arquiteturas. Assim DL se tornou imparável e se firmou como estado na arte não só na tarefa de classificação de imagens mas [numa esmagadora gama de tarefas e áreas](https://paperswithcode.com/sota).
#
#
# ### Ta... mas é hype ou não é?
#
# O hype é real, mas o avanço também!
#
# Ainda existem vários problemas em abertos, porém a área de DL está em constante evolução! Cada dia que passa novas técnicas surgem e mais resultados surpreendentes são alcançados.
#
# Aprender sobre DL e seus conceitos básicos é fundamental para acompanhar, entender e contribuir para esse avanço!
# ## Por que eu deveria me importar?
# 1. [ML e DL estão mudando como se constrói e se pensa sobre software!](https://medium.com/@karpathy/software-2-0-a64152b37c35)
# 2. [DL é muito legal, se você entender de DL você se torna mais legal e interessante por tabela :D](https://www.youtube.com/watch?v=PCBTZh41Ris)
#
# Mas sério, a razão 1. é realmente muito relevante, leia [esse blog](https://medium.com/@karpathy/software-2-0-a64152b37c35) e entenda uma forma de ver como DL está mudando como construimos e construiremos software.
# ## Referências
#
# Este conteúdo é baseado nos seguintes materiais:
#
# * [Machine Learning in Formal Verification, FMCAD 2016 tutorial - Manish Pandey](http://www.cs.utexas.edu/users/hunt/FMCAD/FMCAD16/slides/tutorial1.pdf?fbclid=IwAR0RnrpaJzULlF<KEY>)
# * [Deep Learning Book, capítulo 1 - GoodFellow et Al.](https://www.deeplearningbook.org/contents/intro.html)
# * [Deep Learning 101 - Part 1: History and Background - <NAME>](https://beamandrew.github.io/deeplearning/2017/02/23/deep_learning_101_part1.html)
| _notebooks/2021-01-21-intro_a_deep_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 通过DCGAN实现人脸图像生成
#
# 作者:[ZMpursue](https://github.com/ZMpursue)
# 日期:2020.10.26
#
# 本教程将通过一个示例对DCGAN进行介绍。在向其展示许多真实人脸照片(数据集:[Celeb-A Face](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html))后,我们将训练一个生成对抗网络(GAN)来产生新人脸。本文将对该实现进行详尽的解释,并阐明此模型的工作方式和原因。并不需要过多专业知识,但是可能需要新手花一些时间来理解的模型训练的实际情况。为了节省时间,请尽量选择GPU进行训练。
#
# ## 1 简介
# 本项目基于paddlepaddle,结合生成对抗网络(DCGAN),通过弱监督学习的方式,训练生成真实人脸照片
#
# ### 1.1 什么是GAN?
#
# 生成对抗网络(Generative Adversarial Network [1],简称GAN)是非监督式学习的一种方法,通过让两个神经网络相互博弈的方式进行学习。该方法最初由 lan·Goodfellow 等人于2014年提出,原论文见 [Generative Adversarial Network](https://arxiv.org/abs/1406.2661)。
#
# 生成对抗网络由一个生成网络与一个判别网络组成。生成网络从潜在空间(latent space)中随机采样作为输入,其输出结果需要尽量模仿训练集中的真实样本。判别网络的输入为真实样本或生成网络的输出,其目的是将尽可能的分辨输入为真实样本或生成网络的输出。而生成网络则要尽可能地欺骗判别网络。两个网络相互对抗、不断调整参数。
#
# 让$x$是代表图像的数据。$D(x)$是判别器网络,输出的概率为$x$来自训练数据还是生成器。假设$x$为CHW格式,大小为3x64x64的图像数据,D为判别器网络,$D(x)$为$𝑥$来自训练数据还是生成器。当$𝑥$来自训练数据时$𝐷(𝑥)$尽量接近1,$𝑥$来自生成器时$𝐷(𝑥)$尽量接近0。 因此,$𝐷(𝑥)$也可以被认为是传统的二分类器。
#
# 对于生成器网络, 假设$z$为从标准正态分布采样的空间向量。$G(z)$表示生成器网络,该网络将矢量$z$映射到数据空间,$G(z)$表示生成器网络输出的图像。生成器的目标是拟合训练数据($p_{data}$)的分布,以便可以从该估计分布中生成假样本($p_g$)。
#
# 所以,$D(G(z))$是生成器$G$输出是真实的图像的概率。如Goodfellow的论文所述,$D$和$G$玩一个minmax游戏,其中$D$尝试最大化其正确分类真假的可能性$logD(x)$,以及$G$试图最小化以下可能性$D$会预测其输出是假的$log(1-D(G(x)))$。
#
# GAN的损失函数可表示为:
#
# > $\underset{G}{\text{min}} \underset{D}{\text{max}}V(D,G) = \mathbb{E}_{x\sim p_{data}(x)}\big[logD(x)\big] + \mathbb{E}_{z\sim p_{z}(z)}\big[log(1-D(G(z)))\big]$
#
# 从理论上讲,此minmax游戏的解决方案是$p_g = p_{data}$,鉴别者会盲目猜测输入是真实的还是假的。但是,GAN的收敛理论仍在积极研究中,实际上GAN常常会遇到梯度消失/爆炸问题。
# 生成对抗网络常用于生成以假乱真的图片。此外,该方法还被用于生成视频、三维物体模型等。
#
#
# ### 1.2 什么是DCGAN?
#
# DCGAN是深层卷积网络与GAN的结合,其基本原理与GAN相同,只是将生成网络和判别网络用两个卷积网络(CNN)替代。为了提高生成样本的质量和网络的收敛速度,论文中的 DCGAN 在网络结构上进行了一些改进:
#
# * 取消 pooling 层:在网络中,所有的pooling层使用步幅卷积(strided convolutions)(判别器)和微步幅度卷积(fractional-strided convolutions)(生成器)进行替换。
# * 加入batchnorm:在生成器和判别器中均加入batchnorm。
# * 使用全卷积网络:去掉了FC层,以实现更深的网络结构。
# * 激活函数:在生成器(G)中,最后一层使用Tanh函数,其余层采用ReLU函数 ; 判别器(D)中都采用LeakyReLU。
#
#
# ## 2 环境设置及数据集
#
# 环境:paddlepaddle、scikit-image、numpy、matplotlib
#
# 在本教程中,我们将使用[Celeb-A Faces](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html)数据集,该数据集可以在链接的网站或[AI Studio](https://aistudio.baidu.com/aistudio/datasetdetail/39207)中下载。数据集将下载为名为img_align_celeba.zip的文件。下载后,并将zip文件解压缩到该目录中。
# img_align_celeba目录结构应为:
# ```
# /path/to/project
# -> img_align_celeba
# -> 188242.jpg
# -> 173822.jpg
# -> 284702.jpg
# -> 537394.jpg
# ...
# ```
# ### 2.1 数据集预处理
# 多线程处理,以裁切坐标(0,10)和(64,74),将输入网络的图片裁切到64*64。
# +
from PIL import Image
import os.path
import os
import threading
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
'''多线程将图片缩放后再裁切到64*64分辨率'''
#裁切图片宽度
w = 64
#裁切图片高度
h = 64
#裁切点横坐标(以图片左上角为原点)
x = 0
#裁切点纵坐标
y = 20
def cutArray(l, num):
avg = len(l) / float(num)
o = []
last = 0.0
while last < len(l):
o.append(l[int(last):int(last + avg)])
last += avg
return o
def convertjpg(jpgfile,outdir,width=w,height=h):
img=Image.open(jpgfile)
(l,h) = img.size
rate = min(l,h) / width
try:
img = img.resize((int(l // rate),int(h // rate)),Image.BILINEAR)
img = img.crop((x,y,width+x,height+y))
img.save(os.path.join(outdir,os.path.basename(jpgfile)))
except Exception as e:
print(e)
class thread(threading.Thread):
def __init__(self, threadID, inpath, outpath, files):
threading.Thread.__init__(self)
self.threadID = threadID
self.inpath = inpath
self.outpath = outpath
self.files = files
def run(self):
count = 0
try:
for file in self.files:
convertjpg(self.inpath + file,self.outpath)
count = count + 1
except Exception as e:
print(e)
print('已处理图片数量:' + str(count))
if __name__ == '__main__':
inpath = './work/img_align_celeba/'
outpath = './work/imgs/'
if not os.path.exists(outpath):
os.mkdir(outpath)
files = os.listdir(inpath)
files = cutArray(files,8)
T1 = thread(1, inpath, outpath, files[0])
T2 = thread(2, inpath, outpath, files[1])
T3 = thread(3, inpath, outpath, files[2])
T4 = thread(4, inpath, outpath, files[3])
T5 = thread(5, inpath, outpath, files[4])
T6 = thread(6, inpath, outpath, files[5])
T7 = thread(7, inpath, outpath, files[6])
T8 = thread(8, inpath, outpath, files[7])
T1.start()
T2.start()
T3.start()
T4.start()
T5.start()
T6.start()
T7.start()
T8.start()
T1.join()
T2.join()
T3.join()
T4.join()
T5.join()
T6.join()
T7.join()
T8.join()
# -
# ## 3 模型组网
# ### 3.1 定义数据预处理工具-Paddle.io.Dataset
# 具体参考[Paddle.io.Dataset教程](https://www.paddlepaddle.org.cn/documentation/docs/zh/2.0-rc/api/paddle/io/Dataset_cn.html#dataset)
# +
import os
import cv2
import numpy as np
from skimage import io,color,transform
import matplotlib.pyplot as plt
import math
import time
import paddle
from paddle.io import Dataset
import six
from PIL import Image as PilImage
from paddle.static import InputSpec
paddle.enable_static()
img_dim = 64
'''准备数据,定义Reader()'''
PATH = 'work/imgs/'
class DataGenerater(Dataset):
"""
数据集定义
"""
def __init__(self,path=PATH):
"""
构造函数
"""
super(DataGenerater, self).__init__()
self.dir = path
self.datalist = os.listdir(PATH)
self.image_size = (img_dim,img_dim)
# 每次迭代时返回数据和对应的标签
def __getitem__(self, idx):
return self._load_img(self.dir + self.datalist[idx])
# 返回整个数据集的总数
def __len__(self):
return len(self.datalist)
def _load_img(self, path):
"""
统一的图像处理接口封装,用于规整图像大小和通道
"""
try:
img = io.imread(path)
img = transform.resize(img,self.image_size)
img = img.transpose()
img = img.astype('float32')
except Exception as e:
print(e)
return img
# -
# ### 3.2 测试Paddle.io.DataLoader并输出图片
# +
train_dataset = DataGenerater()
img = paddle.static.data(name='img', shape=[None,3,img_dim,img_dim], dtype='float32')
train_loader = paddle.io.DataLoader(
train_dataset,
places=paddle.CPUPlace(),
feed_list = [img],
batch_size=128,
shuffle=True,
num_workers=2,
use_buffer_reader=True,
use_shared_memory=False,
drop_last=True,
)
for batch_id, data in enumerate(train_loader()):
plt.figure(figsize=(15,15))
try:
for i in range(100):
image = np.array(data[0]['img'][i])[0].transpose((2,1,0))
plt.subplot(10, 10, i + 1)
plt.imshow(image, vmin=-1, vmax=1)
plt.axis('off')
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
plt.suptitle('\n Training Images',fontsize=30)
plt.show()
break
except IOError:
print(IOError)
# -
# ### 3.3 权重初始化
# 在 DCGAN 论文中,作者指定所有模型权重应从均值为0、标准差为0.02的正态分布中随机初始化。
# 调用paddle.nn.initializer.Normal实现initialize设置
conv_initializer=paddle.nn.initializer.Normal(mean=0.0, std=0.02)
bn_initializer=paddle.nn.initializer.Normal(mean=1.0, std=0.02)
# ### 3.4 判别器
# 如上文所述,生成器$D$是一个二进制分类网络,它以图像作为输入,输出图像是真实的(相对应$G$生成的假样本)的概率。输入$Shape$为[3,64,64]的RGB图像,通过一系列的$Conv2d$,$BatchNorm2d$和$LeakyReLU$层对其进行处理,然后通过全连接层输出的神经元个数为2,对应两个标签的预测概率。
#
# * 将BatchNorm批归一化中momentum参数设置为0.5
# * 将判别器(D)激活函数leaky_relu的alpha参数设置为0.2
#
# > 输入: 为大小64x64的RGB三通道图片
# > 输出: 经过一层全连接层最后为shape为[batch_size,2]的Tensor
# +
import paddle
import paddle.nn as nn
import paddle.nn.functional as F
class Discriminator(paddle.nn.Layer):
def __init__(self):
super(Discriminator, self).__init__()
self.conv_1 = nn.Conv2D(
3,64,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="d_conv_weight_1_",initializer=conv_initializer)
)
self.conv_2 = nn.Conv2D(
64,128,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="d_conv_weight_2_",initializer=conv_initializer)
)
self.bn_2 = nn.BatchNorm2D(
128,
weight_attr=paddle.ParamAttr(name="d_2_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_3 = nn.Conv2D(
128,256,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="d_conv_weight_3_",initializer=conv_initializer)
)
self.bn_3 = nn.BatchNorm2D(
256,
weight_attr=paddle.ParamAttr(name="d_3_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_4 = nn.Conv2D(
256,512,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="d_conv_weight_4_",initializer=conv_initializer)
)
self.bn_4 = nn.BatchNorm2D(
512,
weight_attr=paddle.ParamAttr(name="d_4_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_5 = nn.Conv2D(
512,1,4,1,0,
bias_attr=False,weight_attr=paddle.ParamAttr(name="d_conv_weight_5_",initializer=conv_initializer)
)
def forward(self, x):
x = self.conv_1(x)
x = F.leaky_relu(x,negative_slope=0.2)
x = self.conv_2(x)
x = self.bn_2(x)
x = F.leaky_relu(x,negative_slope=0.2)
x = self.conv_3(x)
x = self.bn_3(x)
x = F.leaky_relu(x,negative_slope=0.2)
x = self.conv_4(x)
x = self.bn_4(x)
x = F.leaky_relu(x,negative_slope=0.2)
x = self.conv_5(x)
x = F.sigmoid(x)
return x
# -
# ### 3.5 生成器
# 生成器$G$旨在映射潜在空间矢量$z$到数据空间。由于我们的数据是图像,因此转换$z$到数据空间意味着最终创建具有与训练图像相同大小[3,64,64]的RGB图像。在网络设计中,这是通过一系列二维卷积转置层来完成的,每个层都与$BatchNorm$层和$ReLu$激活函数。生成器的输出通过$tanh$函数输出,以使其返回到输入数据范围[−1,1]。值得注意的是,在卷积转置层之后存在$BatchNorm$函数,因为这是DCGAN论文的关键改进。这些层有助于训练过程中的梯度更好地流动。
#
# **生成器网络结构**
# 
#
# * 将$BatchNorm$批归一化中$momentum$参数设置为0.5
#
# > 输入:Tensor的Shape为[batch_size,100]其中每个数值大小为0~1之间的float32随机数
# > 输出:3x64x64RGB三通道图片
#
# +
class Generator(paddle.nn.Layer):
def __init__(self):
super(Generator, self).__init__()
self.conv_1 = nn.Conv2DTranspose(
100,512,4,1,0,
bias_attr=False,weight_attr=paddle.ParamAttr(name="g_dconv_weight_1_",initializer=conv_initializer)
)
self.bn_1 = nn.BatchNorm2D(
512,
weight_attr=paddle.ParamAttr(name="g_1_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_2 = nn.Conv2DTranspose(
512,256,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="g_dconv_weight_2_",initializer=conv_initializer)
)
self.bn_2 = nn.BatchNorm2D(
256,
weight_attr=paddle.ParamAttr(name="g_2_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_3 = nn.Conv2DTranspose(
256,128,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="g_dconv_weight_3_",initializer=conv_initializer)
)
self.bn_3 = nn.BatchNorm2D(
128,
weight_attr=paddle.ParamAttr(name="g_3_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_4 = nn.Conv2DTranspose(
128,64,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="g_dconv_weight_4_",initializer=conv_initializer)
)
self.bn_4 = nn.BatchNorm2D(
64,
weight_attr=paddle.ParamAttr(name="g_4_bn_weight_",initializer=bn_initializer),momentum=0.8
)
self.conv_5 = nn.Conv2DTranspose(
64,3,4,2,1,
bias_attr=False,weight_attr=paddle.ParamAttr(name="g_dconv_weight_5_",initializer=conv_initializer)
)
self.tanh = paddle.nn.Tanh()
def forward(self, x):
x = self.conv_1(x)
x = self.bn_1(x)
x = F.relu(x)
x = self.conv_2(x)
x = self.bn_2(x)
x = F.relu(x)
x = self.conv_3(x)
x = self.bn_3(x)
x = F.relu(x)
x = self.conv_4(x)
x = self.bn_4(x)
x = F.relu(x)
x = self.conv_5(x)
x = self.tanh(x)
return x
# -
# ### 3.6 损失函数
# 选用BCELoss,公式如下:
#
# $Out = -1 * (label * log(input) + (1 - label) * log(1 - input))$
#
###损失函数
loss = paddle.nn.BCELoss()
# ## 4 模型训练
# 设置的超参数为:
# * 学习率:0.0002
# * 输入图片长和宽:64
# * Epoch: 8
# * Mini-Batch:128
# * 输入Tensor长度:100
# * Adam:Beta1:0.5,Beta2:0.999
#
# 训练过程中的每一次迭代,生成器和判别器分别设置自己的迭代次数。为了避免判别器快速收敛到0,本教程默认每迭代一次,训练一次判别器,两次生成器。
# +
import IPython.display as display
import warnings
import paddle.optimizer as optim
warnings.filterwarnings('ignore')
img_dim = 64
lr = 0.0002
epoch = 5
output = "work/Output/"
batch_size = 128
G_DIMENSION = 100
beta1=0.5
beta2=0.999
output_path = 'work/Output'
device = paddle.set_device('gpu')
paddle.disable_static(device)
real_label = 1.
fake_label = 0.
netD = Discriminator()
netG = Generator()
optimizerD = optim.Adam(parameters=netD.parameters(), learning_rate=lr, beta1=beta1, beta2=beta2)
optimizerG = optim.Adam(parameters=netG.parameters(), learning_rate=lr, beta1=beta1, beta2=beta2)
###训练过程
losses = [[], []]
#plt.ion()
now = 0
for pass_id in range(epoch):
# enumerate()函数将一个可遍历的数据对象组合成一个序列列表
for batch_id, data in enumerate(train_loader()):
#训练判别器
optimizerD.clear_grad()
real_cpu = data[0]
label = paddle.full((batch_size,1,1,1),real_label,dtype='float32')
output = netD(real_cpu)
errD_real = loss(output,label)
errD_real.backward()
optimizerD.step()
optimizerD.clear_grad()
noise = paddle.randn([batch_size,G_DIMENSION,1,1],'float32')
fake = netG(noise)
label = paddle.full((batch_size,1,1,1),fake_label,dtype='float32')
output = netD(fake.detach())
errD_fake = loss(output,label)
errD_fake.backward()
optimizerD.step()
optimizerD.clear_grad()
errD = errD_real + errD_fake
losses[0].append(errD.numpy()[0])
###训练生成器
optimizerG.clear_grad()
noise = paddle.randn([batch_size,G_DIMENSION,1,1],'float32')
fake = netG(noise)
label = paddle.full((batch_size,1,1,1),real_label,dtype=np.float32,)
output = netD(fake)
errG = loss(output,label)
errG.backward()
optimizerG.step()
optimizerG.clear_grad()
losses[1].append(errG.numpy()[0])
if batch_id % 100 == 0:
if not os.path.exists(output_path):
os.makedirs(output_path)
# 每轮的生成结果
generated_image = netG(noise).numpy()
imgs = []
plt.figure(figsize=(15,15))
try:
for i in range(100):
image = generated_image[i].transpose()
image = np.where(image > 0, image, 0)
plt.subplot(10, 10, i + 1)
plt.imshow(image, vmin=-1, vmax=1)
plt.axis('off')
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
msg = 'Epoch ID={0} Batch ID={1} \n\n D-Loss={2} G-Loss={3}'.format(pass_id, batch_id, errD.numpy()[0], errG.numpy()[0])
plt.suptitle(msg,fontsize=20)
plt.draw()
plt.savefig('{}/{:04d}_{:04d}.png'.format(output_path, pass_id, batch_id),bbox_inches='tight')
plt.pause(0.01)
display.clear_output(wait=True)
except IOError:
print(IOError)
paddle.save(netG.state_dict(), "work/generator.params")
plt.close()
# -
plt.figure(figsize=(15, 6))
x = np.arange(len(losses[0]))
plt.title('Generator and Discriminator Loss During Training')
plt.xlabel('Number of Batch')
plt.plot(x,np.array(losses[0]),label='D Loss')
plt.plot(x,np.array(losses[1]),label='G Loss')
plt.legend()
plt.savefig('work/Generator and Discriminator Loss During Training.png')
plt.show()
# ## 5 模型评估
# ### 生成器$G$和判别器$D$的损失与迭代变化图
# 
#
# ### 对比真实人脸图像(图一)和生成人脸图像(图二)
# #### 图一
# 
# ### 图二
# 
#
# ## 6 模型预测
# ### 输入随机数让生成器$G$生成随机人脸
# 生成的RGB三通道64*64的图片路径位于“worl/Generate/”
device = paddle.set_device('gpu')
paddle.disable_static(device)
try:
generate = Generator()
state_dict = paddle.load("work/generator.params")
generate.set_state_dict(state_dict)
noise = paddle.randn([100,100,1,1],'float32')
generated_image = generate(noise).numpy()
for j in range(100):
image = generated_image[j].transpose()
plt.figure(figsize=(4,4))
plt.imshow(image)
plt.axis('off')
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
plt.savefig('work/Generate/generated_' + str(j + 1), bbox_inches='tight')
plt.close()
except IOError:
print(IOError)
# ## 7 项目总结
# 简单介绍了一下DCGAN的原理,通过对原项目的改进和优化,一步一步依次对生成器和判别器以及训练过程进行介绍。
# DCGAN采用一个随机噪声向量作为输入,输入通过与CNN类似但是相反的结构,将输入放大成二维数据。采用这种结构的生成模型和CNN结构的判别模型,DCGAN在图片生成上可以达到相当可观的效果。本案例中,我们利用DCGAN生成了人脸照片,您可以尝试更换数据集生成符合个人需求的图片,或尝试修改网络结构观察不一样的生成效果。
# ## 8 参考文献
# [1] Goodfellow, <NAME>.; <NAME>; <NAME>; <NAME>; <NAME>; <NAME>; <NAME>; <NAME>. Generative Adversarial Networks. 2014. arXiv:1406.2661 [stat.ML].
#
# [2] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, And <NAME>, Generative Models, OpenAI, [April 7, 2016]
#
# [3] alimans, Tim; Goodfellow, Ian; <NAME>; <NAME>; <NAME>; <NAME>. Improved Techniques for Training GANs. 2016. arXiv:1606.03498 [cs.LG].
#
# [4] <NAME>, <NAME>, <NAME>. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks[J]. Computer Science, 2015.
#
# [5]<NAME> , Ba J . Adam: A Method for Stochastic Optimization[J]. Computer ence, 2014.
| docs/practices/dcgan_face/dcgan_face.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Mechanics of running regressions
#
# ```{note}
# These next few pages use a classic dataset called "diamonds" to introduce the regression methods. In lectures, we will use finance oriented data.
# ```
#
# ## Objectives
#
# After this page,
#
# 1. You can fit a regression with `statsmodels` or `sklearn` that includes dummy variables, categorical variables, and interaction terms.
# - `statsmodels`: Nicer result tables, usually easier to specifying the regression model
# - `sklearn`: Easier to use within a prediction/ML exercise
# 1. With both modules: You can view the results visually
# 1. With both modules: You can get the coefficients, t-stats, R$^2$, Adj R$^2$, predicted values ($\hat{y}$), and residuals ($\hat{u}$)
#
#
#
# Let's get our hands dirty quickly by loading some data.
# +
# load some data to practice regressions
import seaborn as sns
import numpy as np
diamonds = sns.load_dataset('diamonds')
# this alteration is not strictly necessary to practice a regression
# but we use this in livecoding
diamonds2 = (diamonds.query('carat < 2.5') # censor/remove outliers
.assign(lprice = np.log(diamonds['price'])) # log transform price
.assign(lcarat = np.log(diamonds['carat'])) # log transform carats
.assign(ideal = diamonds['cut'] == 'Ideal')
# some regression packages want you to explicitly provide
# a variable for the constant
.assign(const = 1)
)
# -
#
# ## Our first regression with `statsmodels`
#
# You'll see these steps repeated a lot for the rest of the class:
# 1. Load the module
# 1. Load your data, and set up your y and X variables
# 1. Pick the model, _usually something like:_ `model = <moduleEstimator>`
# 1. Fit the model and store the results, _usually something like:_ `results = model.fit()`
# 1. Get predicted values, _usually something like:_ `predicted = results.predict()`
#
# +
import statsmodels.api as sm # need this
y = diamonds2['lprice'] # pick y
X = diamonds2[['const','lcarat']] # set up all your X vars as a matrix
# NOTICE I EXPLICITLY GIVE X A CONSTANT SO IT FITS AN INTERCEPT
model1 = sm.OLS(y,X) # pick model type (OLS here) and specify model features
results1 = model1.fit() # estimate / fit
print(results1.summary()) # view results (coefs, t-stats, p-vals, R2, Adj R2)
y_predicted1 = results1.predict() # get the predicted results
residuals1 = results1.resid # get the residuals
#residuals1 = y - y_predicted1 # another way to get the residuals
print('\n\nParams:')
print(results1.params) # if you need to access the coefficients (e.g. to save them), results.params
# -
# ## A better way to regress with `statsmodels`
#
# ```{tip}
# This is my preferred way to run a regression in Python unless I _have_ to use sklearn.
#
# [The documentation with tricks and examples for how to write a regression formula with statsmodels is here.](https://www.statsmodels.org/stable/examples/notebooks/generated/formulas.html)
# ```
#
# In the above, replace
# ```python
# y = diamonds2['lprice'] # pick y
# X = diamonds2[['const','lcarat']] # set up all your X vars as a matrix
#
# model1 = sm.OLS(y,X) # pick model type (OLS here) and specify model features
# ```
# with this
# ```python
# model1 = sm.OLS.from_formula('lprice ~ lcarat',data=diamonds2)
# ```
# which I can replace with this (after adding `from statsmodels.formula.api import ols as sm_ols` to my imports)
# ```python
# model1 = sm_ols('lprice ~ lcarat',data=diamonds2)
# ```
#
# **WOW!** This isn't just more convenient (1 line is less than 3), I like this because
#
# 1. You can set up the model (the equation) more naturally. Notice that I didn't set up the y and X variables as explicit variables. Simply tell that $y=a+b*X+c*Z$ by writing out `y ~ X + Z`
# 1. It allows you to **EASILY** include categorical variables (see below)
# 1. It allows you to **EASILY** include interaction effects (see below)
#
# + tags=["hide-output"]
from statsmodels.formula.api import ols as sm_ols # need this
model2 = sm_ols('lprice ~ lcarat', # specify model (you don't need to include the constant!)
data=diamonds2)
results2 = model2.fit() # estimate / fit
print(results2.summary()) # view results ... identical to before
y_predicted2 = results2.predict() # get the predicted results
residuals2 = results2.resid # get the residuals
#residuals1 = y - y_predicted1 # another way to get the residuals
print('\n\nParams:')
print(results2.params) # if you need to access the coefficients (e.g. to save them), results.params
# -
# ```{note}
# We will cover what all these numbers mean later, but this page is focusing on the how-to.
# ```
# ## Regression with `sklearn`
#
# `sklearn` is pretty similar but
# - when setting up the model object, you don't tell it what data to put into the model
# - you call the model object, and then fit it on data, with `model.fit(X,y)`
# - it doesn't have the nice summary tables
# +
from sklearn.linear_model import LinearRegression
y = diamonds2['lprice'] # pick y
X = diamonds2[['const','lcarat']] # set up all your X vars as a matrix
# NOTICE I EXPLICITLY GIVE X A CONSTANT SO IT FITS AN INTERCEPT
model3 = LinearRegression() # set up the model object (but don't tell sklearn what data it gets!)
results3 = model3.fit(X,y) # fit it, and tell it what data to fit on
print('INTERCEPT:', results3.intercept_) # to get the coefficients, you print out the intercept
print('COEFS:', results3.coef_) # and the other coefficients separately (yuck)
y_predicted3 = results3.predict(X) # get predicted y values
residuals3 = y - y_predicted3 # get residuals
# -
# ```{admonition} That's so much uglier. Why use sklearn?
# :class: warning
#
# Because `sklearn` is the go-to for training models using more sophisticated ML ideas (which we will talk about some later in the course!). Two nice walkthroughs:
# - [This guide from the PythonDataScienceHandbook](https://jakevdp.github.io/PythonDataScienceHandbook/05.06-linear-regression.html) (you can use different data though)
# - The "Linear Regression" section [here](https://becominghuman.ai/linear-regression-in-python-with-pandas-scikit-learn-72574a2ec1a5) shows how you can run regressions on training samples and test them out of sample
# ```
# ## Plotting the regression fit
#
# Once you save the predicted values ($\hat{y}$), it's easy to add it to a plot:
# +
import matplotlib.pyplot as plt
# step 1: plot our data as you want
sns.scatterplot(x='lcarat',y='lprice',
data=diamonds2.sample(500,random_state=20)) # sampled just to avoid overplotting
# step 2: add the fitted regression line (the real X values and the predicted y values)
sns.lineplot(x=diamonds2['lcarat'],y=y_predicted1,color='red')
plt.show()
# -
# ```{note}
# `sns.lmplot` and `sns.regplot` will put a regression line on a scatterplot without having to set up and run a regression, but they also overplot the points when you have a lot of data. That's why I used the approach above - scatterplot a subsample of the data and then overlay the regression line.
#
# One other alternative is to use `sns.lmplot` and `sns.regplot`, but use the `x_bins` parameter to report a "binned scatterplot". Check it out if you're curious.
# ```
# ## Including dummy variables
#
# Suppose you started by estimating the price of diamonds as a function of carats
#
# $$
# \log(\text{price})=a+\beta_0 \log(\text{carat}) +u
# $$
#
# but you realize it will be different for ideal cut diamonds. That is, a 1 carat diamond might cost \$1,000, but if it's ideal, it's an extra \$500 dollars.
#
# $$
# \log(\text{price})=
# \begin{cases}
# a+\beta_0 \log(\text{carat}) + \beta_1 +u, & \text{if ideal cut} \\
# a+\beta_0 \log(\text{carat}) +u, & \text{otherwise}
# \end{cases}
# $$
#
# Notice that $\beta_0$ in this model are the same for ideal and non-ideal.
#
# ```{tip}
# **Here is how you run this test: You just add the dummy variable as a new variable in the formula!**
# ```
# + tags=[]
# ideal is a dummy variable = 1 if ideal and 0 if not ideal
print(sm_ols('lprice ~ lcarat + ideal', data=diamonds2).fit().summary())
# -
# <img src=https://media.giphy.com/media/zcCGBRQshGdt6/source.gif width="400">
#
# ## Including categorical variables
#
# Dummy variables take on two values (on/off, True/False, 0/1). Categorical variables can take on many levels. "Industry" and "State" are typical categorical variables in finance applications.
#
# $$
# \log(\text{price})=
# \begin{cases}
# a+\beta_0 \log(\text{carat}) + \beta_1 +u, & \text{if premium cut} \\
# a+\beta_0 \log(\text{carat}) + \beta_2 +u, & \text{if very good cut} \\
# a+\beta_0 \log(\text{carat}) + \beta_3 +u, & \text{if good cut} \\
# a+\beta_0 \log(\text{carat}) + \beta_4 +u, & \text{if fair cut} \\
# a+\beta_0 \log(\text{carat}) +u, & \text{otherwise (i.e. ideal)}
# \end{cases}
# $$
#
# `sm_ols` also processes categorical variables easily!
#
# ```{tip}
# **Here is how you run this test: You just add the categorical variable as a new variable in the formula!**
# ```
#
# ```{warning}
# WARNING 1: A good idea is to **ALWAYS** put your categorical variable inside of "C()" like below. This tells statsmodels that the variable should be treated as a categorical variable EVEN IF it is a number. (Which would otherwise be treated like a number.)
# ```
#
# ```{warning}
# WARNING 2: You don't create a dummy variable for all the categories! As long as you include a constant in the regression ($a$), one of the categories is covered by the constant. Above, "ideal" is captured by the intercept.
#
# And if you look at the results of the next regression, the "ideal" cut level doesn't have a coefficient. **Statsmodels** automatically drops one of the categories when you use the formula approach. Nice!!!
#
# But if you manually set up the regression in statsmodels or sklearn, you have to drop one level yourself!!!
# ```
#
# + tags=[]
print(sm_ols('lprice ~ lcarat + C(cut)', data=diamonds2).fit().summary())
# -
# <img src=https://media.giphy.com/media/NaboQwhxK3gMU/source.gif width="400">
#
# ## Including interaction terms
#
# Suppose that an ideal cut diamond doesn't just add a fixed dollar value to the diamond. Perhaps it also changes the value of having a larger diamond. You might say that
# - A high quality cut is even more valuable for a larger diamond than it is for a small diamond. ("A great cut makes a diamond sparkle, but it's hard to see sparkle on a tiny diamond no matter what.")
# - In other words, the effect of carats depends on the cut and visa versa
# - In other words, "the cut variable **interacts** with the carat variable"
# - So you might say that, "a better cut changes the slope/coefficient of carat"
# - Or equivalently, "a better cut changes the return on a larger carat"
#
# Graphically, it's easy to see, as `sns.lmplot` by default gives each cut a unique slope on carats:
# +
import matplotlib.pyplot as plt
# add reg lines to plot
fig, ax = plt.subplots()
ax = sns.regplot(data=diamonds2.query('cut == "Fair"'),
y='lprice',x='lcarat',scatter=False,ci=None)
ax = sns.regplot(data=diamonds2.query('cut == "Ideal"'),
y='lprice',x='lcarat',scatter=False,ci=None)
# scatter
sns.scatterplot(data=diamonds2.groupby('cut').sample(80,random_state=99).query('cut in ["Ideal","Fair"]'),
x='lcarat',y='lprice',hue='ideal',ax=ax)
plt.show()
# -
# Those two different lines above are estimated by
#
# $$
# \log(\text{price})= a+ \beta_0 \log(\text{carat}) + \beta_1 \text{Ideal} + \beta_2\log(\text{carat})\cdot \text{Ideal}
# $$
#
# If you plug in 1 for $ideal$, you get the line for ideal diamonds as
#
# $$
# \log(\text{price})= a+ \beta_1 +(\beta_0 + \beta_2) \log(\text{carat})
# $$
#
# If you plug in 0 for $ideal$, you get the line for fair diamonds as
#
# $$
# \log(\text{price})= a+ \beta_0 \log(\text{carat})
# $$
#
# So, by including that interaction term, you get that the slope on carats is different for Ideal than Fair diamonds.
#
# ````{tip}
# **Here is how you run this test: You just add the two variables as a new variable in the formula, along with one term where they are both multiplied!**
#
# ````
# + tags=["hide-output"]
# you can include the interaction of x and z by adding "+x*z" in the spec, like:
sm_ols('lprice ~ lcarat + ideal + lcarat*ideal', data=diamonds2.query('cut in ["Fair","Ideal"]')).fit().summary()
# -
# This shows that a 1% increase in carats is associated with a 1.52% increase in price for fair diamonds, but a 1.71% increase for ideal diamonds (1.52+0.18).
#
# Thus: The return on carats is different (and higher) for better cut diamonds!
| content/05/02b_mechanics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Motif Scan
# Transcription Factor binding motifs are commonly found enriched in cis-regulatory elements and can inform the potential regulatory mechanism of the elements. The first step to study these DNA motifs is to scan their occurence in the genome regions.
#
# ## The Default MotifSet
# Currently, the motif scan function uses a default motif dataset that contains >2000 motifs from three databases {cite}`Khan2018,Kulakovskiy2018,Jolma2013`, each motif is also annotated with human and mouse gene names to facilitate further interpretation.
#
# Following the analysis in {cite}`Vierstra2020` (see also [this great blog](https://www.vierstra.org/resources/motif_clustering)), these motifs are clustered into 286 motif-clusters based on their similarity (and some motifs are almost identical). We will scan all individual motifs here, but also aggregate the results to motif-cluster level. It is recommended to perform futher analysis at the motif-cluster level (such as motif enrichment analysis).
from ALLCools.mcds import RegionDS
from ALLCools.motif import MotifSet, get_default_motif_set
# check out the default motif set
default_motif_set = get_default_motif_set()
default_motif_set.n_motifs
# metadata of the motifs
default_motif_set.meta_table
# motif cluster
default_motif_set.motif_cluster
# single motif object
default_motif_set.motif_list[0]
# ### Motif PSSM Cutoffs
# +
# To re-calculate motif thresholds with a different method or parameter
# default_motif_set.calculate_threshold(method='balanced', cpu=1, threshold_value=1000)
# -
# ## Scan Default Motifs
dmr_ds = RegionDS.open('test_HIP', select_dir=['dmr'])
dmr_ds
# ## Default Motif Database
#
# The {func}`scan_motifs <ALLCools.mcds.region_ds.RegionDS.scan_motifs>` method of RegionDS will perform motif scan using the default motif set over all the regions. This is a time consumming step, scanning 2M regions with 40 CPUs take ~3 days.
dmr_ds.scan_motifs(genome_fasta='../../data/genome/mm10.fa',
cpu=45,
standardize_length=500,
motif_set_path=None,
chrom_size_path=None,
combine_cluster=True,
dim='motif')
# ### Motif Values
# After motif scaning, three value for each motif in each region is stored:
# - n_motifs
# - max_score
# - total_score
dmr_ds.get_index('motif_value')
# ### Individual motifs
dmr_ds['dmr_motif_da']
# ### Motif Clusters
dmr_ds['dmr_motif-cluster_da']
# ## Scan Other Motifs
#
# Comming soon...
| docs/allcools/cluster_level/Correlation/04.motif_scan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intel® Low Precision Optimization Tool (iLiT) Sample for Tensorflow
# ## Agenda
# - Train a CNN Model Based on Keras
# - Quantize Keras Model by ilit
# - Compare Quantized Model
# Import python packages and check version.
#
# Make sure the Tensorflow is **2.2** and iLiT, matplotlib are installed.
# +
import tensorflow as tf
print(tf.__version__)
import matplotlib.pyplot as plt
import ilit
print(ilit.__path__)
import numpy as np
# -
# ## Train a CNN Model Based on Keras
#
# We prepare a script '**alexnet.py**' to provide the functions to train a CNN model.
#
# ### Dataset
# Use [MNIST](http://yann.lecun.com/exdb/mnist/) dataset to recognize hand writing numbers.
# Load the dataset.
# +
import alexnet
data = alexnet.read_data()
x_train, y_train, label_train, x_test, y_test, label_test = data
print('train', x_train.shape, y_train.shape, label_train.shape)
print('test', x_test.shape, y_test.shape, label_test.shape)
# -
# ### Build Model
#
# Build a CNN model like Alexnet by Keras API based on Tensorflow.
# Print the model structure by Keras API: summary().
# +
classes = 10
width = 28
channels = 1
model = alexnet.create_model(width ,channels ,classes)
model.summary()
# -
# ### Train the Model with the Dataset
#
# Set the **epochs** to "**3**"
# +
epochs = 3
alexnet.train_mod(model, data, epochs)
# -
# ### Freeze and Save Model to Single PB
#
# Set the input node name is "**x**".
# +
from tensorflow.python.framework.convert_to_constants import convert_variables_to_constants_v2
def save_frezon_pb(model, mod_path):
# Convert Keras model to ConcreteFunction
full_model = tf.function(lambda x: model(x))
concrete_function = full_model.get_concrete_function(
x=tf.TensorSpec(model.inputs[0].shape, model.inputs[0].dtype))
# Get frozen ConcreteFunction
frozen_model = convert_variables_to_constants_v2(concrete_function)
# Generate frozen pb
tf.io.write_graph(graph_or_graph_def=frozen_model.graph,
logdir=".",
name=mod_path,
as_text=False)
fp32_frezon_pb_file = "fp32_frezon.pb"
save_frezon_pb(model, fp32_frezon_pb_file)
# -
# !ls -la fp32_frezon.pb
# ## Quantize FP32 Model by iLiT
#
# iLiT supports to quantize the model with a validation dataset for tuning.
# Finally, it returns an frezon quantized model based on int8.
#
# We prepare a python script "**ilit_quantize_model.py**" to call iLiT to finish the all quantization job.
# Following code sample is used to explain the code.
#
# ### Define Dataloader
#
# The class **Dataloader** provides an iter function to return the image and label as batch size.
# We uses the validation data of MNIST dataset.
# +
import mnist_dataset
import math
class Dataloader(object):
def __init__(self, batch_size):
self.batch_size = batch_size
def __iter__(self):
x_train, y_train, label_train, x_test, y_test,label_test = mnist_dataset.read_data()
batch_nums = math.ceil(len(x_test)/self.batch_size)
for i in range(batch_nums-1):
begin = i*self.batch_size
end = (i+1)*self.batch_size
yield x_test[begin: end], label_test[begin: end]
begin = (batch_nums-1)*self.batch_size
yield x_test[begin:], label_test[begin:]
# -
# ### Define Load FP32 Model
# Load the saved fp32 model in previous step.
#
# It's defined as alexnet.load_pb(in_model)
# !cat alexnet.py
# ### Define Yaml File
#
# We define alexnet.yaml to save the necessary parameters for iLiT.
# In this case, we only need to change the input/output according to the fp32 model.
#
# In this case, the input node name is '**x**'.
#
# Output name is '**Identity**'.
# !cat alexnet.yaml
# ### Define Tuning Function
# We follow the template to create the tuning function. The function will return a frezon quantized model (int8 model).
# +
import ilit
def auto_tune(input_graph_path, yaml_config, batch_size):
fp32_graph = alexnet.load_pb(input_graph_path)
tuner = ilit.Tuner(yaml_config)
assert(tuner)
dataloader = Dataloader(batch_size)
assert(dataloader)
q_model = tuner.tune(
fp32_graph,
q_dataloader=dataloader,
eval_func=None,
eval_dataloader=dataloader)
return q_model
def save_int8_frezon_pb(q_model, path):
from tensorflow.python.platform import gfile
f = gfile.GFile(path, 'wb')
f.write(q_model.as_graph_def().SerializeToString())
print("Save to {}".format(path))
yaml_file = "alexnet.yaml"
batch_size = 200
int8_pb_file = "alexnet_int8_model.pb"
# -
# ### Call Function to Quantize the Model
#
# Show the code in "**ilit_quantize_model.py**".
# !cat ilit_quantize_model.py
# We will execute the "**ilit_quantize_model.py**" to show the whole process of quantizing a model.
# !python ilit_quantize_model.py
# We get a quantized model file "**alexnet_int8_model.pb**"
# ## Compare Quantized Model
#
# Define a function to return validation dataset and calculate the accuracy.
# +
import time
def val_data():
x_train, y_train, label_train, x_test, y_test,label_test = mnist_dataset.read_data()
return x_test, y_test, label_test
def calc_accuracy(predictions, labels):
predictions = np.argmax(predictions, axis=1)
same = 0
for i, x in enumerate(predictions):
if x==labels[i]:
same += 1
if len(predictions)==0:
return 0
else:
return same/len(predictions)
# -
# Define infer function to test the single frezon PB model.
# +
import numpy as np
import tensorflow as tf
def calc_accuracy(predictions, labels):
predictions = np.argmax(predictions, axis=1)
same = 0
for i, x in enumerate(predictions):
if x==labels[i]:
same += 1
if len(predictions)==0:
return 0
else:
return same/len(predictions)
def get_concrete_function(graph_def, inputs, outputs, print_graph=False):
def imports_graph_def():
tf.compat.v1.import_graph_def(graph_def, name="")
wrap_function = tf.compat.v1.wrap_function(imports_graph_def, [])
graph = wrap_function.graph
return wrap_function.prune(
tf.nest.map_structure(graph.as_graph_element, inputs),
tf.nest.map_structure(graph.as_graph_element, outputs))
def infer_perf_pb(pb_model_file, inputs=["x:0"], outputs=["Identity:0"]):
q_model = alexnet.load_pb(pb_model_file)
concrete_function = get_concrete_function(graph_def=q_model.as_graph_def(),
inputs=inputs,
outputs=outputs,
print_graph=True)
x_test, y_test, label_test = val_data()
bt = time.time()
_frozen_graph_predictions = concrete_function(x=tf.constant(x_test))[0]
et = time.time()
accuracy = calc_accuracy(_frozen_graph_predictions, label_test)
print('accuracy:', accuracy)
throughput = x_test.shape[0] / (et - bt)
print('max throughput(fps):', throughput)
#latency when BS=1
bt = time.time()
times = 1000
for i in range(times):
_frozen_graph_predictions = concrete_function(x=tf.constant(x_test[:1]))[0]
et = time.time()
latency = (et - bt) * 1000 / times
print('latency(ms):', latency)
return accuracy, throughput, latency
#warm up
_accuracy32, _throughput32, _latency32 = infer_perf_pb(fp32_frezon_pb_file)
#test
accuracy32, throughput32, latency32 = infer_perf_pb(fp32_frezon_pb_file)
accuracy8, throughput8, latency8 = infer_perf_pb(int8_pb_file)
# -
# Execute the functions to get the performance data.
# +
def autolabel(ax, rects):
"""
Attach a text label above each bar displaying its height
"""
for rect in rects:
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2., 1.05*height,
'%0.2f' % float(height),
ha='center', va='bottom')
def draw_bar(x, t, y, subplot, color, x_lab, y_lab, width=0.2):
plt.subplot(subplot)
plt.xticks(x, t)
ax1 = plt.gca()
ax1.set_xlabel(x_lab)
ax1.set_ylabel(y_lab, color=color)
rects1 = ax1.bar(x, y, color=color, width=width)
ax1.tick_params(axis='y', labelcolor=color)
autolabel(ax1, rects1)
accuracys = [accuracy32, accuracy8]
throughputs = [throughput32, throughput8]
latencys = [latency32, latency8]
print('throughputs', throughputs)
print('latencys', latencys)
print('accuracys', accuracys)
accuracys_perc = [accu*100 for accu in accuracys]
t = ['FP32', 'INT8']
x = [0, 1]
plt.figure(figsize=(16,6))
draw_bar(x, t, throughputs, 131, 'tab:green', 'Throughput(fps)', '', width=0.2)
draw_bar(x, t, latencys, 132, 'tab:blue', 'Latency(s)', '', width=0.2)
draw_bar(x, t, accuracys_perc, 133, '#28a99d', 'Accuracys(%)', '', width=0.2)
plt.show()
# -
# ### FP32 vs INT8
#
# Compare the performance data based on data of FP32 model.
# +
throughputs_times = [1, throughputs[1]/throughputs[0]]
latencys_times = [1, latencys[1]/latencys[0]]
accuracys_times = [0, accuracys_perc[1] - accuracys_perc[0]]
print('throughputs_times', throughputs_times)
print('latencys_times', latencys_times)
print('accuracys_times', accuracys_times)
plt.figure(figsize=(16,6))
draw_bar(x, t, throughputs_times, 131, 'tab:green', 'Throughput Comparison (big is better)', '', width=0.2)
draw_bar(x, t, latencys_times, 132, 'tab:blue', 'Latency Comparison (small is better)', '', width=0.2)
draw_bar(x, t, accuracys_times, 133, '#28a99d', 'Accuracys Loss(%)', '', width=0.2)
plt.show()
# -
# ## Sample Running is Finished
print("[CODE_SAMPLE_COMPLETED_SUCCESFULLY]")
# ### Summary
# Performance Improvement:
#
# - FP32 to INT8.
# - Intel® Deep Learning Boost speed up INT8 if your CPU is the Second Generation Intel® Xeon® Scalable Processors which supports it.
| AI-and-Analytics/Getting-Started-Samples/IntelLowPrecisionOptimization_GettingStarted/iLiT-Sample-for-Tensorflow/ilit_sample_tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_latest_p37
# language: python
# name: conda_pytorch_latest_p37
# ---
# # Module 1. Check Inference Results & Local Mode Deployment
# ---
#
# ## Overview
#
# 본 핸즈온은 AWS AIML Blog의 내용을 기반으로 MNIST 예제 대신 좀 더 실용적인 한국어 자연어 처리 예시를 다루며, 총 3종류(Sentiment Classification, KorSTS, KoBART)의 자연어 처리 모델을 SageMaker 다중 컨테이너 엔드포인트(Multi-container endpoint)로 배포하는 법을 익혀 봅니다.
#
# 이미 SageMaker 기본 개념(로컬 모드, 호스팅 엔드포인트)과 자연어 처리 & Huggingface을 다뤄 보신 분들은 이 섹션을 건너 뛰고 다음 노트북으로 진행하셔도 됩니다.
#
# ### References
# - AWS AIML Blog: https://aws.amazon.com/ko/blogs/machine-learning/deploy-multiple-serving-containers-on-a-single-instance-using-amazon-sagemaker-multi-container-endpoints/
# - Developer Guide: https://docs.aws.amazon.com/sagemaker/latest/dg/multi-container-endpoints.html
# !pip install -qU sagemaker botocore boto3 awscli
# !pip install --ignore-installed PyYAML
# !pip install transformers==4.12.5
# +
import json
import os
import sys
import torch
import boto3
import sagemaker
import datetime
from sagemaker import get_execution_role
from sagemaker.pytorch import PyTorchModel
from src.utils import print_outputs, prepare_model_artifact, NLPPredictor
role = get_execution_role()
boto_session = boto3.session.Session()
sm_session = sagemaker.session.Session()
sm_client = boto_session.client("sagemaker")
sm_runtime = boto_session.client("sagemaker-runtime")
# -
# <br>
#
# ## 1. Check Inference Results & Debugging
# ---
#
# 로컬 엔드포인트나 호스팅 엔드포인트 배포 전, 로컬 환경 상에서 직접 추론을 수행하여 결과를 확인합니다. 참고로, SageMaker에서 TensorFlow를 제외한 머신 러닝 프레임워크 추론 컨테이너는 아래의 인터페이스를 사용합니다.
#
# #### Option 1.
# - `model_fn(model_dir)`: 네트워크 아키텍처를 정의하고 S3의 model_dir에 저장된 모델 아티팩트를 로드합니다.
# - `input_fn(request_body, content_type)`: 입력 데이터를 전처리합니다. (예: request_body로 전송된 bytearray 배열을 PIL.Image로 변환 수 cropping, resizing, normalization등의 전처리 수행). content_type은 입력 데이터 종류에 따라 다양하게 처리 가능합니다. (예: application/x-npy, application/json, application/csv 등)
# - `predict_fn(input_object, model)`: input_fn을 통해 들어온 데이터에 대해 추론을 수행합니다.
# - `output_fn(prediction, accept_type)`: predict_fn에서 받은 추론 결과를 추가 변환을 거쳐 프론트 엔드로 전송합니다.
#
# #### Option 2.
# - `model_fn(model_dir)`: 네트워크 아키텍처를 정의하고 S3의 model_dir에 저장된 모델 아티팩트를 로드합니다.
# - `transform_fn(model, request_body, content_type, accept_type)`: input_fn(), predict_fn(), output_fn()을 transform_fn()으로 통합할 수 있습니다.
#
# 모델, 배포에 초점을 맞추기 위해 Huggingface에 등록된 `KoELECTRA-Small-v3` 모델을 기반으로 네이버 영화 리뷰 데이터셋과 KorSTS (Korean Semantic Textual Similarity) 데이터셋으로 파인 튜닝하였습니다. 파인 튜닝은 온프레미스나 Huggingface on SageMaker로 쉽게 수행 가능합니다.
#
# - KoELECTRA: https://github.com/monologg/KoELECTRA
# - Huggingface on Amazon SageMaker: https://huggingface.co/docs/sagemaker/main
#
#
# ### Model A: Sentiment Classification
#
# 네이버 영화 리뷰 데이터의 긍정/부정 판별 예시입니다.
# - Naver sentiment movie corpus: https://github.com/e9t/nsmc
# - 예시
# - '이 영화는 최고의 영화입니다' => {"predicted_label": "Pos", "score": 0.96}
# - '최악이에요. 배우의 연기력도 좋지 않고 내용도 너무 허접합니다' => {"predicted_label": "Neg", "score": 0.99}
# !pygmentize src/inference_nsmc.py
# +
from src.inference_nsmc import model_fn, input_fn, predict_fn, output_fn
modelA_path = 'model-nsmc'
with open('samples/nsmc.txt', mode='rb') as file:
modelA_input_data = file.read()
modelA = model_fn(modelA_path)
transformed_inputs = input_fn(modelA_input_data)
predicted_classes_jsonlines = predict_fn(transformed_inputs, modelA)
modelA_outputs = output_fn(predicted_classes_jsonlines)
print(modelA_outputs[0])
# -
# ### Model B: Semantic Textual Similarity (STS)
#
# 두 문장간의 유사도를 정량화하는 예시입니다.
# - KorNLI and KorSTS: https://github.com/kakaobrain/KorNLUDatasets
# - 예시
# - ['맛있는 라면을 먹고 싶어요', '후루룩 쩝쩝 후루룩 쩝쩝 맛좋은 라면'] => {"score": 4.78}
# - ['뽀로로는 내친구', '머신러닝은 러닝머신이 아닙니다.'] => {"score": 0.23}
# !pygmentize src/inference_korsts.py
# +
from src.inference_korsts import model_fn, input_fn, predict_fn, output_fn
modelB_path = 'model-korsts'
with open('samples/korsts.txt', mode='rb') as file:
modelB_input_data = file.read()
modelB = model_fn(modelB_path)
transformed_inputs = input_fn(modelB_input_data)
predicted_classes_jsonlines = predict_fn(transformed_inputs, modelB)
modelB_outputs = output_fn(predicted_classes_jsonlines)
print(modelB_outputs[0])
# -
# ### Model C: KoBART (Korean Bidirectional and Auto-Regressive Transformers)
#
# 문서 내용(예: 뉴스 기사)을 요약하는 예시입니다.
#
# - KoBART: https://github.com/SKT-AI/KoBART
# - KoBART Summarization: https://github.com/seujung/KoBART-summarization
# !pygmentize src/inference_kobart.py
# S3로 모델 아티팩트를 복사하는 대신 Huggingface에 등록된 모델을 그대로 사용합니다. model.pth는 0바이트의 빈 파일이며, 추론을 수행하기 위한 소스 코드들만 아카이빙됩니다.
# +
from src.inference_kobart import model_fn, transform_fn
modelC_path = 'model-kobart'
f = open(f"{modelC_path}/model.pth", 'w')
f.close()
with open('samples/kobart.txt', mode='rb') as file:
modelC_input_data = file.read()
modelC = model_fn('./')
outputs = transform_fn(modelC, modelC_input_data)
with open('samples/kobart.txt', mode='rb') as file:
modelC_input_data = file.read()
# -
# 결괏값들을 확인했다면 로컬 모드로 빠르게 배포하여 테스트하는 것을 권장드립니다. 단, SageMaker Studio는 로컬 모드를 지원하지 않기 때문에 아래 섹션은 SageMaker에서 실행해 주세요.
# <br>
#
# ## 2. (SageMaker Only) Local Mode Deployment for Model A
# ---
#
# ### Deploy Model A
# +
modelA_artifact_name = 'modelA.tar.gz'
prepare_model_artifact(modelA_path, model_artifact_name=modelA_artifact_name)
local_model_path = f'file://{os.getcwd()}/{modelA_artifact_name}'
model = PyTorchModel(
model_data=local_model_path,
role=role,
entry_point='inference_nsmc.py',
source_dir='src',
framework_version='1.7.1',
py_version='py3',
predictor_cls=NLPPredictor,
)
predictor = model.deploy(
initial_instance_count=1,
instance_type='local'
)
# -
# ### Invoke using SageMaker Python SDK
# SageMaker SDK `predict()` 메서드로 간단하게 추론을 실행할 수 있습니다.
# +
inputs = [{"text": ["이 영화는 최고의 영화입니다"]},
{"text": ["최악이에요. 배우의 연기력도 좋지 않고 내용도 너무 허접합니다"]}]
predicted_classes = predictor.predict(inputs)
# -
for c in predicted_classes:
print(c)
# ### Invoke using Boto3 API
# 이번에는 boto3의 `invoke_endpoint()` 메서드로 추론을 수행해 보겠습니다.
# Boto3는 서비스 레벨의 low-level SDK로, ML 실험에 초점을 맞춰 일부 기능들이 추상화된 high-level SDK인 SageMaker SDK와 달리 SageMaker API를 완벽하게 제어할 수 있습으며, 프로덕션 및 자동화 작업에 적합합니다.
# +
local_sm_runtime = sagemaker.local.LocalSagemakerRuntimeClient()
endpoint_name = model.endpoint_name
response = local_sm_runtime.invoke_endpoint(
EndpointName=endpoint_name,
ContentType='application/jsonlines',
Accept='application/jsonlines',
Body=modelA_input_data
)
outputs = response['Body'].read().decode()
# -
print_outputs(outputs)
# ### Local Mode Endpoint Clean-up
# 엔드포인트를 계속 사용하지 않는다면, 엔드포인트를 삭제해야 합니다. SageMaker SDK에서는 `delete_endpoint()` 메소드로 간단히 삭제할 수 있습니다.
# 참고로, 노트북 인스턴스에서 추론 컨테이너를 배포했기 때문에 엔드포인트를 띄워 놓아도 별도로 추가 요금이 과금되지는 않습니다.
#
# 로컬 엔드포인트는 도커 컨테이너이기 때문에 `docker rm $(docker ps -a -q)` 으로도 간단히 삭제할 수 있습니다.
predictor.delete_endpoint()
# <br>
#
# ## 3. (SageMaker Only) Local Mode Deployment for Model B
# ---
#
# ### Deploy Model B
# +
modelB_artifact_name = 'modelB.tar.gz'
prepare_model_artifact(modelB_path, model_artifact_name=modelB_artifact_name)
local_model_path = f'file://{os.getcwd()}/{modelB_artifact_name}'
model = PyTorchModel(
model_data=local_model_path,
role=role,
entry_point='inference_korsts.py',
source_dir='src',
framework_version='1.7.1',
py_version='py3',
predictor_cls=NLPPredictor,
)
predictor = model.deploy(
initial_instance_count=1,
instance_type='local'
)
# -
# ### Invoke using SageMaker Python SDK
# SageMaker SDK `predict()` 메서드로 간단하게 추론을 실행할 수 있습니다.
# +
inputs = [{"text": ["맛있는 라면을 먹고 싶어요", "후루룩 쩝쩝 후루룩 쩝쩝 맛좋은 라면"]},
{"text": ["뽀로로는 내친구", "머신러닝은 러닝머신이 아닙니다."]}]
predicted_classes = predictor.predict(inputs)
# -
for c in predicted_classes:
print(c)
# ### Invoke using Boto3 API
# 이번에는 boto3의 `invoke_endpoint()` 메서드로 추론을 수행해 보겠습니다.
# Boto3는 서비스 레벨의 low-level SDK로, ML 실험에 초점을 맞춰 일부 기능들이 추상화된 high-level SDK인 SageMaker SDK와 달리 SageMaker API를 완벽하게 제어할 수 있습으며, 프로덕션 및 자동화 작업에 적합합니다.
# +
local_sm_runtime = sagemaker.local.LocalSagemakerRuntimeClient()
endpoint_name = model.endpoint_name
response = local_sm_runtime.invoke_endpoint(
EndpointName=endpoint_name,
ContentType='application/jsonlines',
Accept='application/jsonlines',
Body=modelB_input_data
)
outputs = response['Body'].read().decode()
# -
print_outputs(outputs)
# ### Local Mode Endpoint Clean-up
# 엔드포인트를 계속 사용하지 않는다면, 엔드포인트를 삭제해야 합니다. SageMaker SDK에서는 `delete_endpoint()` 메소드로 간단히 삭제할 수 있습니다.
# 참고로, 노트북 인스턴스에서 추론 컨테이너를 배포했기 때문에 엔드포인트를 띄워 놓아도 별도로 추가 요금이 과금되지는 않습니다.
#
# 로컬 엔드포인트는 도커 컨테이너이기 때문에 `docker rm $(docker ps -a -q)` 으로도 간단히 삭제할 수 있습니다.
predictor.delete_endpoint()
# <br>
#
# ## 4. (SageMaker Only) Local Mode Deployment for Model C
# ---
#
# ### Deploy Model C
# +
modelC_artifact_name = 'modelC.tar.gz'
prepare_model_artifact(modelC_path, model_artifact_name=modelC_artifact_name)
local_model_path = f'file://{os.getcwd()}/{modelC_artifact_name}'
model = PyTorchModel(
model_data=local_model_path,
role=role,
entry_point='inference_kobart.py',
source_dir='src',
framework_version='1.7.1',
py_version='py3',
predictor_cls=NLPPredictor,
)
predictor = model.deploy(
initial_instance_count=1,
instance_type='local'
)
# -
import time
time.sleep(3)
# ### Invoke using Boto3 API
# **[주의]** BART 모델은 Auto-Regressive 모델로 내부적으로 연산을 많이 수행하여 기본 인스턴스(예: `ml.t2.medium`)를 사용하는 경우, 시간이 상대적으로 오래 소요됩니다.
# +
local_sm_runtime = sagemaker.local.LocalSagemakerRuntimeClient()
endpoint_name = model.endpoint_name
response = local_sm_runtime.invoke_endpoint(
EndpointName=endpoint_name,
ContentType='application/jsonlines',
Accept='application/jsonlines',
Body=modelC_input_data
)
outputs = response['Body'].read().decode()
# -
print_outputs(outputs)
# <br>
#
# ## Local Mode Endpoint Clean up
# ---
#
# 엔드포인트를 계속 사용하지 않는다면, 엔드포인트를 삭제해야 합니다. SageMaker SDK에서는 `delete_endpoint()` 메소드로 간단히 삭제할 수 있습니다.
# 참고로, 노트북 인스턴스에서 추론 컨테이너를 배포했기 때문에 엔드포인트를 띄워 놓아도 별도로 추가 요금이 과금되지는 않습니다.
#
# 로컬 엔드포인트는 도커 컨테이너이기 때문에 `docker rm $(docker ps -a -q)` 으로도 간단히 삭제할 수 있습니다.
predictor.delete_endpoint()
| key_features/ptn_5_multi-container-endpoint/1_local_endpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
#export
from exp.nb_05b import *
torch.set_num_threads(2)
# ## ConvNet
x_train,y_train,x_valid,y_valid = get_data()
# Helper function to quickly normalize with the mean and standard deviation from our training set:
#export
def normalize_to(train, valid):
m,s = train.mean(),train.std()
return normalize(train, m, s), normalize(valid, m, s)
x_train,x_valid = normalize_to(x_train,x_valid)
train_ds,valid_ds = Dataset(x_train, y_train),Dataset(x_valid, y_valid)
# Let's check it behaved properly.
x_train.mean(),x_train.std()
# +
nh,bs = 50,512
c = y_train.max().item()+1
loss_func = F.cross_entropy
data = DataBunch(*get_dls(train_ds, valid_ds, bs), c)
# -
# To refactor layers, it's useful to have a `Lambda` layer that can take a basic function and convert it to a layer you can put in `nn.Sequential`.
#
# NB: if you use a Lambda layer with a lambda function, your model won't pickle so you won't be able to save it with PyTorch. So it's best to give a name to the function you're using inside your Lambda (like flatten below).
# +
#export
class Lambda(nn.Module):
def __init__(self, func):
super().__init__()
self.func = func
def forward(self, x): return self.func(x)
def flatten(x): return x.view(x.shape[0], -1)
# -
# This one takes the flat vector of size `bs x 784` and puts it back as a batch of images of 28 by 28 pixels:
def mnist_resize(x): return x.view(-1, 1, 28, 28)
# We can now define a simple CNN.
def get_cnn_model(data):
return nn.Sequential(
Lambda(mnist_resize),
nn.Conv2d( 1, 8, 5, padding=2,stride=2), nn.ReLU(), #14
nn.Conv2d( 8,16, 3, padding=1,stride=2), nn.ReLU(), # 7
nn.Conv2d(16,32, 3, padding=1,stride=2), nn.ReLU(), # 4
nn.Conv2d(32,32, 3, padding=1,stride=2), nn.ReLU(), # 2
nn.AdaptiveAvgPool2d(1),
Lambda(flatten),
nn.Linear(32,data.c)
)
model = get_cnn_model(data)
# Basic callbacks from the previous notebook:
cbfs = [Recorder, partial(AvgStatsCallback,accuracy)]
opt = optim.SGD(model.parameters(), lr=0.4)
learn = Learner(model, opt, loss_func, data)
run = Runner(cb_funcs=cbfs)
# %time run.fit(1, learn)
# ## CUDA
# This took a long time to run, so it's time to use a GPU. A simple Callback can make sure the model, inputs and targets are all on the same device.
# Somewhat more flexible way
device = torch.device('cuda',0)
class CudaCallback(Callback):
def __init__(self,device): self.device=device
def begin_fit(self): self.model.to(device)
def begin_batch(self): self.run.xb,self.run.yb = self.xb.to(device),self.yb.to(device)
# Somewhat less flexible, but quite convenient
torch.cuda.set_device(device)
#export
class CudaCallback(Callback):
def begin_fit(self): self.model.cuda()
def begin_batch(self): self.run.xb,self.run.yb = self.xb.cuda(),self.yb.cuda()
cbfs.append(CudaCallback)
model = get_cnn_model(data)
opt = optim.SGD(model.parameters(), lr=0.4)
learn = Learner(model, opt, loss_func, data)
run = Runner(cb_funcs=cbfs)
# %time run.fit(3, learn)
# Now, that's definitely faster!
# ## Refactor model
# First we can regroup all the conv/relu in a single function:
def conv2d(ni, nf, ks=3, stride=2):
return nn.Sequential(
nn.Conv2d(ni, nf, ks, padding=ks//2, stride=stride), nn.ReLU())
# Another thing is that we can do the mnist resize in a batch transform, that we can do with a Callback.
# +
#export
class BatchTransformXCallback(Callback):
_order=2
def __init__(self, tfm): self.tfm = tfm
def begin_batch(self): self.run.xb = self.tfm(self.xb)
def view_tfm(*size):
def _inner(x): return x.view(*((-1,)+size))
return _inner
# -
mnist_view = view_tfm(1,28,28)
cbfs.append(partial(BatchTransformXCallback, mnist_view))
# With the `AdaptiveAvgPool`, this model can now work on any size input:
nfs = [8,16,32,32]
# +
def get_cnn_layers(data, nfs):
nfs = [1] + nfs
return [
conv2d(nfs[i], nfs[i+1], 5 if i==0 else 3)
for i in range(len(nfs)-1)
] + [nn.AdaptiveAvgPool2d(1), Lambda(flatten), nn.Linear(nfs[-1], data.c)]
def get_cnn_model(data, nfs): return nn.Sequential(*get_cnn_layers(data, nfs))
# -
# And this helper function will quickly give us everything needed to run the training.
#export
def get_runner(model, data, lr=0.6, cbs=None, opt_func=None, loss_func = F.cross_entropy):
if opt_func is None: opt_func = optim.SGD
opt = opt_func(model.parameters(), lr=lr)
learn = Learner(model, opt, loss_func, data)
return learn, Runner(cb_funcs=listify(cbs))
model = get_cnn_model(data, nfs)
learn,run = get_runner(model, data, lr=0.4, cbs=cbfs)
model
run.fit(3, learn)
# ## Hooks
# ### Manual insertion
# Let's say we want to do some telemetry, and want the mean and standard deviation of each activations in the model. First we can do it manually like this:
class SequentialModel(nn.Module):
def __init__(self, *layers):
super().__init__()
self.layers = nn.ModuleList(layers)
self.act_means = [[] for _ in layers]
self.act_stds = [[] for _ in layers]
def __call__(self, x):
for i,l in enumerate(self.layers):
x = l(x)
if self.training:
self.act_means[i].append(x.data.mean())
self.act_stds [i].append(x.data.std ())
return x
def __iter__(self): return iter(self.layers)
model = SequentialModel(*get_cnn_layers(data, nfs))
learn,run = get_runner(model, data, lr=0.9, cbs=cbfs)
run.fit(2, learn)
# Now we can have a look at the means and stds of the activations at the beginning of training.
for l in model.act_means: plt.plot(l)
plt.legend(range(6));
for l in model.act_stds: plt.plot(l)
plt.legend(range(6));
for l in model.act_means: plt.plot(l[:10])
plt.legend(range(6));
for l in model.act_stds: plt.plot(l[:10])
plt.legend(range(6));
# ### Pytorch hooks
# Hooks are PyTorch object you can add to any nn.Module. A hook will be called when a layer, it is registered to, is executed during the forward pass (forward hook) or the backward pass (backward hook).
#
# Hooks don't require us to rewrite the model.
model = get_cnn_model(data, nfs)
learn,run = get_runner(model, data, lr=0.5, cbs=cbfs)
act_means = [[] for _ in model]
act_stds = [[] for _ in model]
# A hook is attached to a layer, and needs to have a function that takes three arguments: module, input, output. Here we store the mean and std of the output in the correct position of our list.
def append_stats(i, mod, inp, outp):
if mod.training:
act_means[i].append(outp.data.mean())
act_stds [i].append(outp.data.std())
for i,m in enumerate(model): m.register_forward_hook(partial(append_stats, i))
run.fit(1, learn)
for o in act_means: plt.plot(o)
plt.legend(range(5));
# ### Hook class
# We can refactor this in a Hook class. It's very important to remove the hooks when they are deleted, otherwise there will be references kept and the memory won't be properly released when your model is deleted.
# +
#export
def children(m): return list(m.children())
class Hook():
def __init__(self, m, f): self.hook = m.register_forward_hook(partial(f, self))
def remove(self): self.hook.remove()
def __del__(self): self.remove()
def append_stats(hook, mod, inp, outp):
if not hasattr(hook,'stats'): hook.stats = ([],[])
means,stds = hook.stats
if mod.training:
means.append(outp.data.mean())
stds .append(outp.data.std())
# -
# NB: In fastai we use a `bool` param to choose whether to make it a forward or backward hook. In the above version we're only supporting forward hooks.
model = get_cnn_model(data, nfs)
learn,run = get_runner(model, data, lr=0.5, cbs=cbfs)
hooks = [Hook(l, append_stats) for l in children(model[:4])]
run.fit(1, learn)
for h in hooks:
plt.plot(h.stats[0])
h.remove()
plt.legend(range(4));
# ### A Hooks class
# Let's design our own class that can contain a list of objects. It will behave a bit like a numpy array in the sense that we can index into it via:
# - a single index
# - a slice (like 1:5)
# - a list of indices
# - a mask of indices (`[True,False,False,True,...]`)
#
# The `__iter__` method is there to be able to do things like `for x in ...`.
#export
class ListContainer():
def __init__(self, items): self.items = listify(items)
def __getitem__(self, idx):
if isinstance(idx, (int,slice)): return self.items[idx]
if isinstance(idx[0],bool):
assert len(idx)==len(self) # bool mask
return [o for m,o in zip(idx,self.items) if m]
return [self.items[i] for i in idx]
def __len__(self): return len(self.items)
def __iter__(self): return iter(self.items)
def __setitem__(self, i, o): self.items[i] = o
def __delitem__(self, i): del(self.items[i])
def __repr__(self):
res = f'{self.__class__.__name__} ({len(self)} items)\n{self.items[:10]}'
if len(self)>10: res = res[:-1]+ '...]'
return res
ListContainer(range(10))
ListContainer(range(100))
t = ListContainer(range(10))
t[[1,2]], t[[False]*8 + [True,False]]
# We can use it to write a `Hooks` class that contains several hooks. We will also use it in the next notebook as a container for our objects in the data block API.
# +
#export
from torch.nn import init
class Hooks(ListContainer):
def __init__(self, ms, f): super().__init__([Hook(m, f) for m in ms])
def __enter__(self, *args): return self
def __exit__ (self, *args): self.remove()
def __del__(self): self.remove()
def __delitem__(self, i):
self[i].remove()
super().__delitem__(i)
def remove(self):
for h in self: h.remove()
# -
model = get_cnn_model(data, nfs).cuda()
learn,run = get_runner(model, data, lr=0.9, cbs=cbfs)
hooks = Hooks(model, append_stats)
hooks
hooks.remove()
x,y = next(iter(data.train_dl))
x = mnist_resize(x).cuda()
x.mean(),x.std()
p = model[0](x)
p.mean(),p.std()
for l in model:
if isinstance(l, nn.Sequential):
init.kaiming_normal_(l[0].weight)
l[0].bias.data.zero_()
p = model[0](x)
p.mean(),p.std()
# Having given an `__enter__` and `__exit__` method to our `Hooks` class, we can use it as a context manager. This makes sure that onces we are out of the `with` block, all the hooks have been removed and aren't there to pollute our memory.
with Hooks(model, append_stats) as hooks:
run.fit(2, learn)
fig,(ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks:
ms,ss = h.stats
ax0.plot(ms[:10])
ax1.plot(ss[:10])
plt.legend(range(6));
fig,(ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks:
ms,ss = h.stats
ax0.plot(ms)
ax1.plot(ss)
plt.legend(range(6));
# ### Other statistics
# Let's store more than the means and stds and plot histograms of our activations now.
def append_stats(hook, mod, inp, outp):
if not hasattr(hook,'stats'): hook.stats = ([],[],[])
means,stds,hists = hook.stats
if mod.training:
means.append(outp.data.mean().cpu())
stds .append(outp.data.std().cpu())
hists.append(outp.data.cpu().histc(40,0,10)) #histc isn't implemented on the GPU
model = get_cnn_model(data, nfs).cuda()
learn,run = get_runner(model, data, lr=0.9, cbs=cbfs)
for l in model:
if isinstance(l, nn.Sequential):
init.kaiming_normal_(l[0].weight)
l[0].bias.data.zero_()
with Hooks(model, append_stats) as hooks: run.fit(1, learn)
# Thanks to @ste for initial version of histgram plotting code
def get_hist(h): return torch.stack(h.stats[2]).t().float().log1p()
fig,axes = plt.subplots(2,2, figsize=(15,6))
for ax,h in zip(axes.flatten(), hooks[:4]):
ax.imshow(get_hist(h), origin='lower')
ax.axis('off')
plt.tight_layout()
# From the histograms, we can easily get more informations like the min or max of the activations
def get_min(h):
h1 = torch.stack(h.stats[2]).t().float()
return h1[:2].sum(0)/h1.sum(0)
fig,axes = plt.subplots(2,2, figsize=(15,6))
for ax,h in zip(axes.flatten(), hooks[:4]):
ax.plot(get_min(h))
ax.set_ylim(0,1)
plt.tight_layout()
# ## Generalized ReLU
# Now let's use our model with a generalized ReLU that can be shifted and with maximum value.
# +
#export
def get_cnn_layers(data, nfs, layer, **kwargs):
nfs = [1] + nfs
return [layer(nfs[i], nfs[i+1], 5 if i==0 else 3, **kwargs)
for i in range(len(nfs)-1)] + [
nn.AdaptiveAvgPool2d(1), Lambda(flatten), nn.Linear(nfs[-1], data.c)]
def conv_layer(ni, nf, ks=3, stride=2, **kwargs):
return nn.Sequential(
nn.Conv2d(ni, nf, ks, padding=ks//2, stride=stride), GeneralRelu(**kwargs))
class GeneralRelu(nn.Module):
def __init__(self, leak=None, sub=None, maxv=None):
super().__init__()
self.leak,self.sub,self.maxv = leak,sub,maxv
def forward(self, x):
x = F.leaky_relu(x,self.leak) if self.leak is not None else F.relu(x)
if self.sub is not None: x.sub_(self.sub)
if self.maxv is not None: x.clamp_max_(self.maxv)
return x
def init_cnn(m, uniform=False):
f = init.kaiming_uniform_ if uniform else init.kaiming_normal_
for l in m:
if isinstance(l, nn.Sequential):
f(l[0].weight, a=0.1)
l[0].bias.data.zero_()
def get_cnn_model(data, nfs, layer, **kwargs):
return nn.Sequential(*get_cnn_layers(data, nfs, layer, **kwargs))
# -
def append_stats(hook, mod, inp, outp):
if not hasattr(hook,'stats'): hook.stats = ([],[],[])
means,stds,hists = hook.stats
if mod.training:
means.append(outp.data.mean().cpu())
stds .append(outp.data.std().cpu())
hists.append(outp.data.cpu().histc(40,-7,7))
model = get_cnn_model(data, nfs, conv_layer, leak=0.1, sub=0.4, maxv=6.)
init_cnn(model)
learn,run = get_runner(model, data, lr=0.9, cbs=cbfs)
with Hooks(model, append_stats) as hooks:
run.fit(1, learn)
fig,(ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks:
ms,ss,hi = h.stats
ax0.plot(ms[:10])
ax1.plot(ss[:10])
plt.legend(range(5));
fig,(ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks:
ms,ss,hi = h.stats
ax0.plot(ms)
ax1.plot(ss)
plt.legend(range(5));
fig,axes = plt.subplots(2,2, figsize=(15,6))
for ax,h in zip(axes.flatten(), hooks[:4]):
ax.imshow(get_hist(h), origin='lower')
ax.axis('off')
plt.tight_layout()
def get_min(h):
h1 = torch.stack(h.stats[2]).t().float()
return h1[19:22].sum(0)/h1.sum(0)
fig,axes = plt.subplots(2,2, figsize=(15,6))
for ax,h in zip(axes.flatten(), hooks[:4]):
ax.plot(get_min(h))
ax.set_ylim(0,1)
plt.tight_layout()
#export
def get_learn_run(nfs, data, lr, layer, cbs=None, opt_func=None, uniform=False, **kwargs):
model = get_cnn_model(data, nfs, layer, **kwargs)
init_cnn(model, uniform=uniform)
return get_runner(model, data, lr=lr, cbs=cbs, opt_func=opt_func)
sched = combine_scheds([0.5, 0.5], [sched_cos(0.2, 1.), sched_cos(1., 0.1)])
learn,run = get_learn_run(nfs, data, 1., conv_layer, cbs=cbfs+[partial(ParamScheduler,'lr', sched)])
run.fit(8, learn)
# Uniform init may provide more useful initial weights (normal distribution puts a lot of them at 0).
learn,run = get_learn_run(nfs, data, 1., conv_layer, uniform=True,
cbs=cbfs+[partial(ParamScheduler,'lr', sched)])
run.fit(8, learn)
# ## Export
# Here's a handy way to export our module without needing to update the file name - after we define this, we can just use `nb_auto_export()` in the future (h/t <NAME>):
#export
from IPython.display import display, Javascript
def nb_auto_export():
display(Javascript("""{
const ip = IPython.notebook
if (ip) {
ip.save_notebook()
console.log('a')
const s = `!python notebook2script.py ${ip.notebook_name}`
if (ip.kernel) { ip.kernel.execute(s) }
}
}"""))
nb_auto_export()
| dev_course/dl2/06_cuda_cnn_hooks_init.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
import cv2
import numpy as np
import csv
import time
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
from utils import label_map_util
from utils import visualization_utils as vis_util
# -
with open('traffic_measurement.csv', 'w') as f:
writer = csv.writer(f)
csv_line = \
'Vehicle Type/Size, Vehicle Color, Vehicle Movement Direction, Vehicle Speed (km/h)'
writer.writerows([csv_line.split(',')])
cap = cv2.VideoCapture('sub-1504619634606.mp4')
total_passed_vehicle = 0 # using it to count vehicles
MODEL_NAME = 'ssd_mobilenet_v1_coco_2018_01_28'
MODEL_FILE = MODEL_NAME + '.tar.gz'
DOWNLOAD_BASE = \
'http://download.tensorflow.org/models/object_detection/'
PATH_TO_CKPT = MODEL_NAME + '/frozen_inference_graph.pb'
PATH_TO_LABELS = os.path.join('data', 'mscoco_label_map.pbtxt')
NUM_CLASSES = 90
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(PATH_TO_CKPT, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
# +
label_map = label_map_util.load_labelmap(PATH_TO_LABELS)
categories = label_map_util.convert_label_map_to_categories(label_map,
max_num_classes=NUM_CLASSES, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
# Helper code
def load_image_into_numpy_array(image):
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape((im_height, im_width,
3)).astype(np.uint8)
# +
def object_detection_function():
total_passed_vehicle = 0
speed = 'waiting...'
direction = 'waiting...'
size = 'waiting...'
color = 'waiting...'
with detection_graph.as_default():
with tf.Session(graph=detection_graph) as sess: # Definite input and output Tensors for detection_graph
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Each box represents a part of the image where a particular object was detected.
detection_boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Each score represent how level of confidence for each of the objects.
# Score is shown on the result image, together with the class label.
detection_scores = detection_graph.get_tensor_by_name('detection_scores:0')
detection_classes = detection_graph.get_tensor_by_name('detection_classes:0')
num_detections = detection_graph.get_tensor_by_name('num_detections:0')
# for all the frames that are extracted from input video
while cap.isOpened():
(ret, frame) = cap.read()
if not ret:
print ('end of the video file...')
break
input_frame = frame
# Expand dimensions since the model expects images to have shape: [1, None, None, 3]
image_np_expanded = np.expand_dims(input_frame, axis=0)
# Actual detection.
(boxes, scores, classes, num) = \
sess.run([detection_boxes, detection_scores,
detection_classes, num_detections],
feed_dict={image_tensor: image_np_expanded})
# Visualization of the results of a detection.
(counter, csv_line) = \
vis_util.visualize_boxes_and_labels_on_image_array(
cap.get(1),
input_frame,
np.squeeze(boxes),
np.squeeze(classes).astype(np.int32),
np.squeeze(scores),
category_index,
use_normalized_coordinates=True,
line_thickness=2,
)
total_passed_vehicle = total_passed_vehicle + counter
# insert information text to video frame
font = cv2.FONT_HERSHEY_SIMPLEX
cv2.putText(
input_frame,
'Detected Vehicles: ' + str(total_passed_vehicle),
(10, 35),
font,
0.8,
(0, 0xFF, 0xFF),
2,
cv2.FONT_HERSHEY_SIMPLEX,
)
# when the vehicle passed over line and counted, make the color of ROI line green
if counter == 1:
cv2.line(input_frame, (0, 200), (720, 200), (0, 0xFF, 0), 5)
else:
cv2.line(input_frame, (0, 200), (720, 200), (0, 0, 0xFF), 5)
# insert information text to video frame
cv2.rectangle(input_frame, (10, 275), (230, 337), (180, 132, 109), -1)
cv2.putText(
input_frame,
'ROI Line',
(545, 190),
font,
0.6,
(0, 0, 0xFF),
2,
cv2.LINE_AA,
)
cv2.putText(
input_frame,
'LAST PASSED VEHICLE INFO',
(11, 290),
font,
0.5,
(0xFF, 0xFF, 0xFF),
1,
cv2.FONT_HERSHEY_SIMPLEX,
)
cv2.putText(
input_frame,
'-Movement Direction: ' + direction,
(14, 302),
font,
0.4,
(0xFF, 0xFF, 0xFF),
1,
cv2.FONT_HERSHEY_COMPLEX_SMALL,
)
cv2.putText(
input_frame,
'-Speed(km/h): ' + speed,
(14, 312),
font,
0.4,
(0xFF, 0xFF, 0xFF),
1,
cv2.FONT_HERSHEY_COMPLEX_SMALL,
)
cv2.putText(
input_frame,
'-Color: ' + color,
(14, 322),
font,
0.4,
(0xFF, 0xFF, 0xFF),
1,
cv2.FONT_HERSHEY_COMPLEX_SMALL,
)
cv2.putText(
input_frame,
'-Vehicle Size/Type: ' + size,
(14, 332),
font,
0.4,
(0xFF, 0xFF, 0xFF),
1,
cv2.FONT_HERSHEY_COMPLEX_SMALL,
)
cv2.imshow('vehicle detection', input_frame)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
if csv_line != 'not_available':
with open('traffic_measurement.csv', 'a') as f:
writer = csv.writer(f)
(size, color, direction, speed) = \
csv_line.split(',')
writer.writerows([csv_line.split(',')])
cap.release()
cv2.destroyAllWindows()
object_detection_function()
# -
| python_code/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>'s descent
#
# <NAME> (23 March 1882 – 14 April 1935) was a German mathematician who made important contributions to abstract algebra and theoretical physics (https://en.wikipedia.org/wiki/Emmy_Noether).
#
# According to the math genealogy project, <NAME> had 14 doctoral students, who had 76 themselves, ... so until now she has *1365* descendants.
import networkx, json
from francy_widget import FrancyWidget
G = networkx.DiGraph()
data = json.load(open("noether.json"))
nodes = data["nodes"]
#print(len(nodes))
nodes_to_keep = {k:nodes[k] for k in nodes if nodes[k][0]<4}
edges_to_keep = [e for e in data["edges"] if e[1] in nodes_to_keep]
G.add_edges_from(edges_to_keep)
def node_options(n):
options = {}
d = nodes[n]
options["layer"] = d[0]
options["title"] = "%s (%s)" % (d[2].split(",")[0], d[3])
if n in ["6967", "63779", "6982", "29850", "121808", "191816",
"54355", "98035", "44616", "57077", "21851"]:
options["type"] = 'diamond'
else:
options["type"] = 'circle'
return options
FrancyWidget(G, graphType="directed", height=800, zoomToFit=False,
node_options=node_options)
| examples/WomenMathGenealogy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Create 1-Wave and 2-WaveFile and 3-WaveFormat json.
# ## Params define.
class Param:
def __init__(
self,
file_name: str,
num_channels: int,
sample_rate: int,
bits_per_sample: int,
f0: float,
sec: float,
):
self.file_name: str = file_name # FileName
self.chunk_id: str = "RIFF"
self.chunk_size: int = 0
self.wave_format: str = "WAVE"
self.sub_chunk_1_id: str = "fmt "
self.sub_chunk_1_size: int = 16
self.audio_format: int = 1 # BasicPCM
self.num_channels: int = num_channels
self.sample_rate: int = sample_rate # FrameRate
self.byte_rate: int = sample_rate * num_channels * bits_per_sample // 8
self.block_align: int = num_channels * bits_per_sample // 8
self.bits_per_sample: int = bits_per_sample
self.sub_chunk_2_id: str = "data"
self.sub_chunk_2_size: int = 0
self.data: List[int] = None
# Fields above this are required for json
self.dataBytes: bytes = b""
self.f0: float = f0 # FundamentalFrequency(Hz)
self.sec: float = sec
def set_data(self, data):
self.data = data.tolist()
self.dataBytes = data.tobytes()
self.sub_chunk_2_size = len(self.dataBytes)
self.chunk_size = 4 + (8 + self.sub_chunk_1_size) + (
8 + self.sub_chunk_2_size)
params = [
Param("X",
num_channels=1,
sample_rate=8000,
bits_per_sample=16,
f0=440.000,
sec=2),
]
# ## 1. Create wave
import numpy as np
for p in params:
ndarr = np.arange(0, p.sample_rate * p.sec)
sin_wave = 32767 * np.sin(2 * np.pi * p.f0 * ndarr / p.sample_rate)
sin_wave = sin_wave.astype(np.int16)
p.set_data(sin_wave)
# ## 2. Write wav file
SAMPLE_WAVES_DIR = "../internal/samples/waves"
import wave
from os import path
for p in params:
write_path = f"{path.join(SAMPLE_WAVES_DIR, p.file_name)}.wav"
with wave.open(write_path, "wb") as f:
f.setnchannels(p.num_channels)
f.setsampwidth(p.block_align)
f.setframerate(p.sample_rate)
f.writeframes(p.dataBytes)
# ## 3. Write wave format json
SAMPLE_FORMAT_JSON_DIR = "../internal/samples/format_jsons"
import json
unnecessary_keys = ["dataBytes", "f0", "sec"]
for p in params:
d = p.__dict__
for k in unnecessary_keys:
del d[k]
write_path = f"{path.join(SAMPLE_FORMAT_JSON_DIR, p.file_name)}.json"
with open(write_path, "w") as f:
json.dump(d, f, indent=4)
| gen/sample_json_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/samirlenka/pandas_exercises/blob/master/01_Getting_%26_Knowing_Your_Data/Occupation/Exercise_with_Solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qiT6eZHLsjom"
# # Ex3 - Getting and Knowing your Data
#
# Check out [Occupation Exercises Video Tutorial](https://www.youtube.com/watch?v=W8AB5s-L3Rw&list=PLgJhDSE2ZLxaY_DigHeiIDC1cD09rXgJv&index=4) to watch a data scientist go through the exercises
# + [markdown] id="X0jpzAQqsjo0"
# This time we are going to pull data directly from the internet.
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
# + id="Rmpjqkmssjo1"
import pandas as pd
import numpy as np
# + [markdown] id="cth3omU-sjo1"
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/justmarkham/DAT8/master/data/u.user).
# + [markdown] id="sG9BW73xsjo2"
# ### Step 3. Assign it to a variable called users and use the 'user_id' as index
# + id="_VIBvO5gsjo2"
users = pd.read_csv('https://raw.githubusercontent.com/justmarkham/DAT8/master/data/u.user', sep='|', index_col='user_id')
# + [markdown] id="xN_b3fuqsjo3"
# ### Step 4. See the first 25 entries
# + id="wNzhhpWusjo3" outputId="311455a6-7c4d-4d03-c20a-095c3c2b5ba4" colab={"base_uri": "https://localhost:8080/", "height": 855}
users.head(25)
# + [markdown] id="ZPkSromIsjo4"
# ### Step 5. See the last 10 entries
# + id="4f_B8dHksjo5" outputId="32ee7e9c-85c7-4b9a-b4dd-d2aabaf77944"
users.tail(10)
# + [markdown] id="JIv_bat4sjo5"
# ### Step 6. What is the number of observations in the dataset?
# + id="0KcK5VU4sjo6" outputId="19fb7499-bfd3-419a-a8c2-505d07b5097c"
users.shape[0]
# + [markdown] id="crfL-O_-sjo6"
# ### Step 7. What is the number of columns in the dataset?
# + id="ju48FFunsjo7" outputId="ebdf271b-3ebc-432b-d326-f6b2a5d83323"
users.shape[1]
# + [markdown] id="M200gIwZsjo7"
# ### Step 8. Print the name of all the columns.
# + id="xzoADseZsjo7" outputId="fbbb460e-75d5-44bb-81b7-a14beb6ed384"
users.columns
# + [markdown] id="WKpOet0Csjo8"
# ### Step 9. How is the dataset indexed?
# + id="jS4fwwwDsjo8" outputId="939bf86c-b551-41e6-8708-1843931a3628" colab={"base_uri": "https://localhost:8080/"}
# "the index" (aka "the labels")
users.index
# + [markdown] id="d3KHFOwvsjo9"
# ### Step 10. What is the data type of each column?
# + id="qpRxiKlBsjo9" outputId="f6aa2063-d564-4bf5-f219-13705eafe2cd" colab={"base_uri": "https://localhost:8080/"}
users.dtypes
# + [markdown] id="-MLig1E4sjo9"
# ### Step 11. Print only the occupation column
# + id="GgpS74UYsjo-" outputId="150cbf9a-e355-46ba-fc1e-662c9142f8d2" colab={"base_uri": "https://localhost:8080/"}
users.occupation
#or
users['occupation']
# + [markdown] id="msc-gzMFsjo-"
# ### Step 12. How many different occupations are in this dataset?
# + id="Sg0RanDOsjo-" outputId="91e639fa-6094-45ac-f2f7-5ed97d40231d" colab={"base_uri": "https://localhost:8080/"}
users.occupation.nunique()
#or
#users['occupation'].nunique()
#or by using value_counts() which returns the count of unique elements
users.occupation.value_counts().count()
# + [markdown] id="5nqcgVA9sjo_"
# ### Step 13. What is the most frequent occupation?
# + id="Tggtvte4sjo_" outputId="6e1dc275-128c-449e-c951-ca3f9ecc7ba0" colab={"base_uri": "https://localhost:8080/"}
#Because "most" is asked
users.occupation.value_counts().head(1).index[0]
#or
#to have the top 5
users.occupation.value_counts().head()
# + [markdown] id="046aL_X6sjo_"
# ### Step 14. Summarize the DataFrame.
# + id="MR2RV9w0sjpB" outputId="c2dc2d2b-da78-4ea6-bf32-538dac5d6b53"
users.describe() #Notice: by default, only the numeric columns are returned.
# + [markdown] id="769QeZ6jsjpC"
# ### Step 15. Summarize all the columns
# + id="sNIyiZtdsjpD" outputId="5e478586-27c7-44f6-df7a-33fea032ef4f" colab={"base_uri": "https://localhost:8080/", "height": 390}
users.describe(include = "all") #Notice: By default, only the numeric columns are returned.
# + [markdown] id="Lfigw6ftsjpD"
# ### Step 16. Summarize only the occupation column
# + id="rb9kHQsksjpE" outputId="8cfc440d-513b-44da-cfe4-ab463430dd4e" colab={"base_uri": "https://localhost:8080/"}
users.occupation.describe()
# + [markdown] id="v5hZjUZxsjpE"
# ### Step 17. What is the mean age of users?
# + id="omINoFE_sjpF" outputId="7f6598f3-2b59-4924-f2c1-4bfbf2b1f84d" colab={"base_uri": "https://localhost:8080/"}
round(users.age.mean())
# + [markdown] id="p0tOfTDhsjpF"
# ### Step 18. What is the age with least occurrence?
# + id="KhXlF1R0sjpG" outputId="d0545d02-0669-49cb-c61c-ff3978259069" colab={"base_uri": "https://localhost:8080/"}
users.age.value_counts().tail() #7, 10, 11, 66 and 73 years -> only 1 occurrence
# + [markdown] id="9o9F-DBFNtl1"
# ### Step 19. What is the age with most 5 occurrence?
# + id="jVbp40LMNpDg" outputId="41b6acbd-1667-4781-b22a-0e17bb056c7d" colab={"base_uri": "https://localhost:8080/"}
users.age.value_counts().head(5)
| 01_Getting_&_Knowing_Your_Data/Occupation/Exercise_with_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 2 : Finding Gaussian Process hyperparameters
#
# Below some packages to import that will be used for this lesson
#
# Cell bellow is here for avoiding scrolling when plot is create within ipython notebook
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines){
# return false;
# }
# -
# Classical package for manipulating
# array, for plotting and interactiv plots.
import pylab as plt
from matplotlib import gridspec
import numpy as np
import ipywidgets as widgets
from ipywidgets import interact
import itertools
import emcee
import corner
import treecorr
import treegp
from treegp import AnisotropicRBF, eval_kernel
# ## Exercice 5) Maximum Likelihood search of best hyperparameters / kernel (example in 1D):
# +
##########################################################################################
# EXERCICE 5: Maximum Likelihood search of best hyperparameters / kernel (example in 1D) #
##########################################################################################
def log_likelihood(param, kernel_type="RBF"):
if param[1] <=0:
return -np.inf
else:
Kernel = "%f * %s(%f)"%((param[0]**2, kernel_type, param[1]))
#Kernel = eval_kernel(Kernel)
gp = treegp.GPInterpolation(kernel=Kernel, optimizer='none',
normalize=False, white_noise=0., p0=[3000., 0.,0.],
n_neighbors=4, average_fits=None, nbins=20,
min_sep=None, max_sep=None)
gp.initialize(x, y, y_err=y_err)
log_L = gp.return_log_likelihood()
return log_L
def mcmc_hyperparameters_search(run_mcmc=False):
if run_mcmc:
p0 = [1., 0.5]
np.random.seed(42)
ndim, nwalkers = len(p0), 100
pos = [p0 + 1e-4*np.random.randn(ndim) for i in range(nwalkers)]
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_likelihood)
sampler.run_mcmc(pos, 600)
LABEL = ['$\sigma$','$l$']
for j in range(ndim):
plt.figure()
for i in range(nwalkers):
plt.plot(sampler.chain[i,:,j],'k', alpha=0.1)
plt.ylabel(LABEL[j], fontsize=20)
samples = sampler.chain[:, 60:, :].reshape((-1, ndim))
fig = corner.corner(samples, labels=LABEL,
levels=(0.68, 0.95))
return samples
data = np.loadtxt('data/data_1d_grf.txt')
x = data[:,0].reshape((len(data[:,0]),1))
y = data[:,1]
y_err = data[:,2]
def gp_regression(x, new_x, y, kernel, y_err=None):
if y_err is None:
y_err =np.ones_like(y) *1e-10
gp = treegp.GPInterpolation(kernel=kernel, optimizer='none',
normalize=False, white_noise=0., p0=[3000., 0.,0.],
n_neighbors=4, average_fits=None, nbins=20,
min_sep=None, max_sep=None)
gp.initialize(x, y, y_err=y_err)
y_predict, y_cov = gp.predict(new_x, return_cov=True)
y_std = np.sqrt(np.diag(y_cov))
return y_predict, y_std
@interact(sigma = widgets.FloatSlider(value=1.2, min=0.75, max=2.5, step=0.01, description='$\sigma$:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.2f'),
l = widgets.FloatSlider(value=0.6, min=0.4, max=1.5, step=0.01, description='$l$:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.2f'),
kernel = widgets.Dropdown(options=['RBF', 'Matern'],
value='RBF',
description='Kernel:',
disabled=False,))
def plot_samples(sigma, l, kernel):
new_x = np.linspace(-24,24, 400).reshape((400,1))
Kernel = "%f * %s(%f)"%((sigma**2, kernel, l))
y_pred, y_std = gp_regression(x, new_x, y, Kernel, y_err=y_err)
gs = gridspec.GridSpec(1, 2, width_ratios=[1.5, 1])
plt.figure(figsize=(20,8))
plt.subplot(gs[0])
# Data
plt.scatter(x, y, c='b', label = 'data')
plt.errorbar(x, y, linestyle='', yerr=y_err, ecolor='b',
alpha=0.7,marker='.',zorder=0)
# GP prediction
plt.plot(new_x, y_pred, 'r', lw =3, label = 'GP prediction')
plt.fill_between(new_x.T[0], y_pred-y_std, y_pred+y_std, color='r', alpha=0.3)
plt.plot(new_x, np.zeros_like(new_x),'k--')
plt.xlim(-24,24)
plt.ylim(-3.,3.)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('X', fontsize=20)
plt.ylabel('Y', fontsize=20)
plt.legend(fontsize=18)
plt.subplot(gs[1])
distance = np.linspace(0, 2, 40)
coord = np.array([distance, np.zeros_like(distance)]).T
Kernel = eval_kernel(Kernel)
pcf = Kernel.__call__(coord, Y=np.zeros_like(coord))[:,0]
plt.plot(distance, pcf, 'k', lw=3)
plt.ylim(0, 2.5**2)
plt.xlim(0, 2)
plt.ylabel('$\\xi(|x_i-x_j|)$', fontsize=20)
plt.xlabel('$|x_i-x_j|$', fontsize=20)
plt.title('Used correlation function (%s)'%(kernel), fontsize=16)
samples = np.loadtxt('data/data_1d_grf_mcmc_likelihood_sampling_%s.txt'%(kernel))
fig = corner.corner(samples, labels=['$\sigma$','$l$'],
truths=[sigma, l, ], levels=(0.68, 0.95))
fig.suptitle('Kernel type: ' + kernel + ', $\log$ likelihood = %.2f'%(log_likelihood([sigma, l], kernel_type=kernel)))
# -
# ## Exercice 6) 2-point correlation function search of best hyperparameters / kernel (example in 1D) :
# +
##########################################################################################
# EXERCICE 6: Maximum Likelihood search of best hyperparameters / kernel (example in 1D) #
##########################################################################################
data = np.loadtxt('data/data_1d_grf_4000_points.txt')
x = data[:,0].reshape((len(data[:,0]),1))
y = data[:,1]
y_err = data[:,2]
np.random.seed(42)
Filter = np.random.choice([True, False, False, False, False], size=len(y))
cat = treecorr.Catalog(x=x[:,0], y=np.zeros_like(x[:,0]), k=(y-np.mean(y)), w=1./y_err**2)
kk = treecorr.KKCorrelation(min_sep=0.05, max_sep=1.5, nbins=15.)
kk.process(cat)
delta_distance = kk.meanr
xi = kk.xi
def gp_regression(x, new_x, y, kernel, y_err=None):
if y_err is None:
y_err =np.ones_like(y) *1e-10
gp = treegp.GPInterpolation(kernel=kernel, optimizer='none',
normalize=False, white_noise=0., p0=[3000., 0.,0.],
n_neighbors=4, average_fits=None, nbins=20,
min_sep=None, max_sep=None)
gp.initialize(x, y, y_err=y_err)
y_predict, y_cov = gp.predict(new_x, return_cov=True)
y_std = np.sqrt(np.diag(y_cov))
return y_predict, y_std
@interact(sigma = widgets.FloatSlider(value=1.2, min=0.75, max=2.5, step=0.01, description='$\sigma$:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.2f'),
l = widgets.FloatSlider(value=0.6, min=0.4, max=1.5, step=0.01, description='$l$:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.2f'),
kernel = widgets.Dropdown(options=['RBF', 'Matern'],
value='RBF',
description='Kernel:',
disabled=False,))
def plot_samples(sigma, l, kernel):
y_reduce = y[Filter]
x_reduce = x[Filter]
y_err_reduce = y_err[Filter]
new_x = np.linspace(-55, 55, 500).reshape((500,1))
Kernel = "%f * %s(%f)"%((sigma**2, kernel, l))
y_pred, y_std = gp_regression(x_reduce, new_x, y_reduce,
Kernel, y_err=y_err_reduce)
gs = gridspec.GridSpec(1, 2, width_ratios=[1.5, 1])
plt.figure(figsize=(20,8))
plt.subplot(gs[0])
# Data
plt.scatter(x, y, c='b', label = 'data')
plt.errorbar(x, y, linestyle='', yerr=y_err, ecolor='b',
alpha=0.7,marker='.',zorder=0)
# GP prediction
plt.plot(new_x, y_pred, 'r', lw =3, label = 'GP prediction')
plt.fill_between(new_x.T[0], y_pred-y_std, y_pred+y_std, color='r', alpha=0.3)
plt.plot(new_x, np.zeros_like(new_x),'k--')
plt.xlim(-55, 55)
plt.ylim(-3.,3.)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('X', fontsize=20)
plt.ylabel('Y', fontsize=20)
plt.legend(fontsize=18)
plt.subplot(gs[1])
distance = np.linspace(0, 2, 40)
coord = np.array([distance, np.zeros_like(distance)]).T
Kernel = eval_kernel(Kernel)
pcf = Kernel.__call__(coord, Y=np.zeros_like(coord))[:,0]
plt.plot(distance, pcf, 'k', lw=3, label="Used correlation function")
plt.scatter(delta_distance, xi, c='b', s=80, label="Measured 2-point correlation function")
plt.ylim(0, 2.)
plt.xlim(0, 2)
plt.ylabel('$\\xi(|x_i-x_j|)$', fontsize=20)
plt.xlabel('$|x_i-x_j|$', fontsize=20)
plt.legend(fontsize=14)
plt.title('Used correlation function (%s)'%(kernel), fontsize=16)
# -
| notebooks/example_2_hyperparameters_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Hw 18_11_2021
# Проверка целостности матрицы, т.е. количество элементов в строках
# должны совпадать, тип матрицы должен быть список
def CheckMatrixCompletness(matrix):
# проверяем тип всей матрицы
if type(matrix) is not list:
return False
if type(matrix[0]) is not list:
return False
# определим количество столбцов
column_num = len(matrix[0])
# проверяем целостность всех строк
for row in matrix:
if type(row) is not list:
return False
if len(row) != column_num:
return False
# Проверка, что все элементы матрицы - числа
for element in row:
if type(element) is not int and type(element) is not float:
return False
return True
# Проверка корректности вектора
def IsAppropriateVector(arg_list, isRow = True, isPrint = True, dim = None):
if type(arg_list) is not list: # проверка соглашения вида вектора
if isPrint:
print("Argument is not vector!")
return False
if isRow: # вектор-строка состоит только из чисел
for elem in arg_list:
elemType = type(elem)
if elemType is not int and elemType is not float:
if isPrint:
print("Vector has inappropriate element!")
return False
else: # вектор-столбец состоит из одноэлементных списков
for elem in arg_list:
if type(elem) is not list or len(elem) != 1:
if isPrint:
print("Vector has inappropriate element!")
return False
if type(elem[0]) is not int and type(elem[0]) is not float:
if isPrint:
print("Vector has inappropriate element!")
return False
if dim: # проверка длины, если это необходимо
if type(dim) is not int or dim < 1:
if isPrint:
print("Inappropriate vector dimension!")
return False
if len(arg_list) != dim:
if isPrint:
print("Vector dimension isn't correct!")
return False
return True
def AddVectors(vector_a, vector_b):
if len(vector_a) != len(vector_b):
return "Vector lengths aren't equal!"
sum_vector = []
for i in range(len(vector_a)):
sum_vector.append(vector_a[i] + vector_b[i])
return sum_vector
def MultiplyToScalar(vector, scalar):
res_vector = []
for elem in vector:
res_vector.append(scalar * elem)
return res_vector
def GetColumn(matrix, column):
result = []
for i in range(0, len(matrix)):
result += [matrix[i][column]]
return result
def TranspouseMatrix(matrix):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
result = []
for i in range(len(matrix[0])):
result += [[]]
for j in range(len(matrix)):
result[i] += [matrix[j][i]]
return result
'''
[1 2] [1 2 3] = [9 12 15]
[3 4] [4 5 6] = [19 26 33]
1 * [1 3]' + 4 * [2 4]' = [1 3]' + [8 16]' = [9 19]'
2 * [1 3]' + 5 * [2 4]' = [2 6]' + [10 20]' = [12 26]'
3 * [1 3]' + 6 * [2 4]' = [3 9]' + [12 24]' = [15 33]'
'''
def MatrixMultiplation(matrix_a, matrix_b):
# Проверка соответствия на размеров матриц на произведение
matrix_a_shape = CalculateMatrixShape(matrix_a)
matrix_b_shape = CalculateMatrixShape(matrix_b)
if type(matrix_a_shape) is str:
return "Matrix_a is incorrect"
if type(matrix_b_shape) is str:
return "Matrix_b is incorrect"
if matrix_a_shape[1] != matrix_b_shape[0]:
return "Matrices shapes are not insufficient."
# --------------------------------------------------------
#return "Ok"
result = []
for matrix_b_column_id in range(0, len(matrix_b[0])):
#print(GetColumn(matrix_b, matrix_b_column_id))
matrix_b_column = GetColumn(matrix_b, matrix_b_column_id)
vec_res = [0] * len(matrix_a)
for matrix_a_column_id in range(0, len(matrix_a[0])):
vec = MultiplyToScalar(GetColumn(matrix_a, matrix_a_column_id), matrix_b_column[matrix_a_column_id])
vec_res = AddVectors(vec_res, vec)
#print("--", GetColumn(matrix_a, matrix_a_column_id), "--", matrix_b_column[matrix_a_column_id], "--", vec)
#print ("!!!", vec_res)
result += [vec_res]
return TranspouseMatrix(result)
# Вычисление размеров матрицы
def CalculateMatrixShape(matrix):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
# Количетсво строк
row_num = len(matrix)
# Количетсво столбцов
col_num = len(matrix[0])
return (row_num, col_num)
def MatrixMultiplyToScalar(matrix_a, scalar):
# Проверка
matrix_a_shape = CalculateMatrixShape(matrix_a)
if type(matrix_a_shape) is str:
return "Matrix_a is incorrect"
# --------------------------------------------------------
matrix_a = [] + matrix_a
for i in range(matrix_a_shape[0]):
for j in range(matrix_a_shape[1]):
matrix_a[i][j] *= scalar
return matrix_a
# +
from copy import deepcopy
def det(arr):
size_arr = len(arr)
if size_arr == 1: return arr[0][0]
result = 0
for index_x in range(size_arr):
arr_deepcopy = deepcopy(arr)
del (arr_deepcopy[0])
for i in range(len(arr_deepcopy)):
del (arr_deepcopy[i][index_x])
result += arr[0][index_x] * (-1 if index_x & 1 else 1 ) * det(arr_deepcopy)
return result
# +
from copy import deepcopy
#Дополнительный Минор
def AdditionalMinor(matrix, row, col):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
row_num, _ = CalculateMatrixShape(matrix)
matrix = deepcopy(matrix)
del (matrix[row])
for i in range(row_num - 1):
del (matrix[i][col])
return det(matrix)
# -
#Алгебраическое дополнение
def AlgebraiComplement(matrix, row, col):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
return (-1)**(row + col) * AdditionalMinor(matrix, row, col)
def MatrixInverse(matrix):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
matrixDet = det(matrix)
if matrixDet == 0:
return "Det = 0"
row_num, col_num = CalculateMatrixShape(matrix)
if row_num != col_num:
return "row_num != col_num"
result = []
for i in range(row_num):
result += [[]]
for j in range(col_num):
result[i] += [AlgebraiComplement(matrix, j, i)]
return MatrixMultiplyToScalar(result, 1 / matrixDet)
# +
a = [[1, 0, 0],
[0, 2, 0],
[0, 0, 3]]
print (MatrixInverse(a))
# -
# CH 18_11_2021
# +
from copy import deepcopy
def add_row (matrix, k, row_num, row_num2):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
matrix = deepcopy(matrix)
_, col_num = CalculateMatrixShape(matrix)
for c in range(col_num):
matrix[row_num2][c] += matrix[row_num][c] * k
return matrix
a = [[1, 0, 0],
[0, 2, 0],
[0, 0, 3]]
print (add_row(a, 2, 0, 1))
# -
def swap_col(matrix, r1, r2):
matrix[r1], matrix[r2] = matrix[r2], matrix[r1]
# +
a = [[1, 0, 0],
[0, 2, 0],
[0, 0, 3]]
swap_col(a, 0, 1)
print(a)
# -
def getIndexFirstNymberNotZero(vec):
for i in range(len(vec)):
if vec[i] != 0:
return i
return None
vec = [0, 1]
a = getIndexFirstNymberNotZero(vec)
print(a is None)
# +
def step(matrix):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
row_num, col_num = CalculateMatrixShape(matrix)
col = 0
for R in range(row_num - 1):
vec_col = GetColumn(matrix, col)[R:]
if (vec_col[0] == 0):
while True:
index_el = getIndexFirstNymberNotZero(vec_col)
if index_el is None:
col += 1
if col == col_num:
break
vec_col = GetColumn(matrix, col)[R:]
else:
swap_col(matrix, R, R + index_el)
break
if col == col_num:
break
for r in range(R + 1, row_num):
k = (0 - matrix[r][col]) / matrix[R][col]
matrix = add_row(matrix, k, R, r)
col += 1
return matrix
a = [[1, 2, 3],
[1, 2, 3],
[0, 2, 6]]
print (step(a))
# +
def GlueMatrixVector(matrix, vector_values, isVectors = True):
if not IsAppropriateVector(vector_values, True, False) and isVectors:
return "Vector is incorrect"
elif not CheckMatrixCompletness(matrix) and not isVectors:
return "Matrix is incorrect"
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
row_num, col_num = CalculateMatrixShape(matrix)
if row_num != col_num:
return "Matrix row_num != col_num"
matrix = deepcopy(matrix)
for i in range(row_num):
if isVectors:
matrix[i] += [vector_values[i]]
else:
matrix[i] += vector_values[i]
return matrix
a = [[1, 0, 0],
[3, 2, 0],
[0, 0, 3]]
print (GlueMatrixVector(a, [1, 2, 3]))
# -
def GaussMethod(matrix, vector_values):
if not IsAppropriateVector(vector_values):
return "Vector is incorrect"
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
row_num, col_num = CalculateMatrixShape(matrix)
if row_num != col_num:
return "Matrix row_num != col_num"
if len(vector_values) != row_num:
return "len(vector_values) != row_num"
if det(matrix) == 0:
return "Det = 0"
matrix = GlueMatrixVector(matrix, vector_values)
matrix_step = step(matrix)
for i in range(len(matrix_step)):
if not matrix_step[i][i]:
return "бесконечно много решений"
#print(matrix_step)
result = []
for r in range(row_num):
vec_row = matrix_step[-1-r][:len(matrix_step[-1-r]) - 1]
res = matrix_step[-1-r][-1]
#print(vec_row, res)
index_k = getIndexFirstNymberNotZero(vec_row)
result += [res / vec_row[index_k]]
return result
# +
a = [[1, 0, 0],
[0, 0, 0],
[0, 0, 3]]
print (GaussMethod(a, [1, 2, 3]))
# -
def indentity_matrix(size):
arr = [[0 for i in range(0, size)] for i in range(0, size)]
for i in range(0, size):
arr[i][i] = 1
return arr
def MatrixInverseGauss(matrix):
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
if det(matrix) == 0:
return "Det = 0"
row_num, col_num = CalculateMatrixShape(matrix)
if row_num != col_num:
return "row_num != col_num"
I = indentity_matrix(row_num)
M = deepcopy(matrix)
matrix = GlueMatrixVector(M, I, False)
size = row_num
col = 0
for R in range(size):
vec_col = GetColumn(matrix, col)[R:]
if (vec_col[0] == 0):
while True:
index_el = getIndexFirstNymberNotZero(vec_col)
if index_el is None:
col += 1
if col == col_num:
break
vec_col = GetColumn(matrix, col)[R:]
else:
swap_col(matrix, R, R + index_el)
break
if col == col_num:
break
#print(matrix[R][col])
# 1 = el * k
# k = 1 / el
k = 1 / matrix[R][col]
for i in range(R, len(matrix[R])):
#print(matrix[R][i], end=" - ")
matrix[R][i] *= k
for up in range(0, R):
#print("--up", matrix[up][col])
k = (0 - matrix[up][col]) / matrix[R][col]
matrix = add_row(matrix, k, R, up)
for down in range(R + 1, size):
#print("--down", matrix[down][col])
k = (0 - matrix[down][col]) / matrix[R][col]
matrix = add_row(matrix, k, R, down)
col += 1
#print(matrix)
return [matrix[i][size:] for i in range(size)]
# +
A = [[0, 2],
[1, 0]]
a = MatrixInverseGauss(A)
b = MatrixInverse(A)
print(MatrixMultiplation(A, b))
print(MatrixMultiplation(A, a))
# +
def set_col(matrix, vec, col):
matrix = deepcopy(matrix)
for i in range(len(matrix)):
matrix[i][col] = vec[i]
return matrix
A = [[0, 2],
[1, 0]]
set_col(A, [3, 3], 1)
# +
def kramer(matrix, vector_values):
if not IsAppropriateVector(vector_values):
return "Vector is incorrect"
if not CheckMatrixCompletness(matrix):
return "Matrix is incorrect"
row_num, col_num = CalculateMatrixShape(matrix)
for i in range(len(m)):
if not m[i][i]:
print('бесконечно много решений')
return
if len(vector_values) != row_num:
return "len(vector_values) != row_num"
det_matrix = det(matrix)
if det_matrix == 0:
return "Det = 0"
result = []
for col in range(col_num):
result += [det(set_col(matrix, vector_values, col)) / det_matrix]
return result
# +
a = [[5, 6],
[11, 12]]
#print(kramer(a))
print (kramer(a, [1, 2]))
# -
| hw 1_12_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="84c59c4d97b2f0a0eb52a22d74e7939dd51873bf" _cell_guid="48bb1394-d871-4670-812e-9e80ac87a27a"
# ## INTRODUCTION
# - It’s a Python based scientific computing package targeted at two sets of audiences:
# - A replacement for NumPy to use the power of GPUs
# - Deep learning research platform that provides maximum flexibility and speed
# - pros:
# - Iinteractively debugging PyTorch. Many users who have used both frameworks would argue that makes pytorch significantly easier to debug and visualize.
# - Clean support for dynamic graphs
# - Organizational backing from Facebook
# - Blend of high level and low level APIs
# - cons:
# - Much less mature than alternatives
# - Limited references / resources outside of the official documentation
# - I accept you know neural network basics. If you do not know check my tutorial. Because I will not explain neural network concepts detailed, I only explain how to use pytorch for neural network
# - Neural Network tutorial: https://www.kaggle.com/kanncaa1/deep-learning-tutorial-for-beginners
# - The most important parts of this tutorial from matrices to ANN. If you learn these parts very well, implementing remaining parts like CNN or RNN will be very easy.
# <br>
# <br>**Content:**
# 1. [Basics of Pytorch](#1)
# - Matrices
# - Math
# - Variable
# 1. [Linear Regression](#2)
# 1. [Logistic Regression](#3)
# 1. [Artificial Neural Network (ANN)](#4)
# 1. [Concolutional Neural Network (CNN)](#5)
# 1. Recurrent Neural Network (RNN)
# - https://www.kaggle.com/kanncaa1/recurrent-neural-network-with-pytorch
# -
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
# + [markdown] _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
# <a id="1"></a> <br>
# ## Basics of Pytorch
# ### Matrices
# - In pytorch, matrix(array) is called tensors.
# - 3*3 matrix koy. This is 3x3 tensor.
# - Lets look at array example with numpy that we already know.
# - We create numpy array with np.numpy() method
# - Type(): type of the array. In this example it is numpy
# - np.shape(): shape of the array. Row x Column
# + _uuid="d60e9b9f706124117b1d7ffcdeefb45b9aca45e6" _cell_guid="0a70fde1-b9c4-47c5-aed7-1c863b2fd1e1"
# import numpy library
import numpy as np
# numpy array
array = [[1,2,3],[4,5,6]]
first_array = np.array(array) # 2x3 array
print("Array Type: {}".format(type(first_array))) # type
print("Array Shape: {}".format(np.shape(first_array))) # shape
print(first_array)
# + [markdown] _uuid="28a4eb1e95add272552310eacf20740b895e43cd" _cell_guid="042b8e6f-0be6-43a1-96b4-49b32999a208"
# - We looked at numpy array.
# - Now examine how we implement tensor(pytorch array)
# - import pytorch library with import torch
# - We create tensor with torch.Tensor() method
# - type: type of the array. In this example it is tensor
# - shape: shape of the array. Row x Column
# + _uuid="126ea635dff4e2a6bc9a828dc560863e3be6aa74" _cell_guid="b383b085-a18f-4c18-a093-428336b6acf6"
# import pytorch library
import torch
# pytorch array
tensor = torch.Tensor(array)
print("Array Type: {}".format(tensor.type)) # type
print("Array Shape: {}".format(tensor.shape)) # shape
print(tensor)
# + [markdown] _uuid="fa1375fffab5b879eb828de7261e3e436ec15102" _cell_guid="e1f7ac29-8aa8-46ff-929f-94f2cedf7541"
# - Allocation is one of the most used technique in coding. Therefore lets learn how to make it with pytorch.
# - In order to learn, compare numpy and tensor
# - np.ones() = torch.ones()
# - np.random.rand() = torch.rand()
# + _uuid="2d36d68f3b57eef9d8f0ed94885f3376de92b414" _cell_guid="741468a5-5d91-48d7-95b0-6d02180d0c09"
# numpy ones
print("Numpy {}\n".format(np.ones((2,3))))
# pytorch ones
print(torch.ones((2,3)))
# + _uuid="1e6b8ce52af8a26ffc39fcd751a834ea7c870a2d" _cell_guid="a578ff9f-df45-4acd-b5ec-2e26b2690adb"
# numpy random
print("Numpy {}\n".format(np.random.rand(2,3)))
# pytorch random
print(torch.rand(2,3))
# + [markdown] _uuid="22b5e44de713f58261bf1ff0b3a52b22a81ef1ef" _cell_guid="b5177215-45b5-40c1-b838-2d0e3acb48ba"
# - Even if when I use pytorch for neural networks, I feel better if I use numpy. Therefore, usually convert result of neural network that is tensor to numpy array to visualize or examine.
# - Lets look at conversion between tensor and numpy arrays.
# - torch.from_numpy(): from numpy to tensor
# - numpy(): from tensor to numpy
# + _uuid="c6d3a7b8e0e42fcadecb16264b0563f74d01439a" _cell_guid="f2cedc86-bd28-4709-906f-e236f4a4dbbe"
# random numpy array
array = np.random.rand(2,2)
print("{} {}\n".format(type(array),array))
# from numpy to tensor
from_numpy_to_tensor = torch.from_numpy(array)
print("{}\n".format(from_numpy_to_tensor))
# from tensor to numpy
tensor = from_numpy_to_tensor
from_tensor_to_numpy = tensor.numpy()
print("{} {}\n".format(type(from_tensor_to_numpy),from_tensor_to_numpy))
# + [markdown] _uuid="42cbe3900b733ab12867612d484d6fffaccd5e31" _cell_guid="6d7038e6-6aaf-4a1e-9204-406ab21082a2"
# ### Basic Math with Pytorch
# - Resize: view()
# - a and b are tensor.
# - Addition: torch.add(a,b) = a + b
# - Subtraction: a.sub(b) = a - b
# - Element wise multiplication: torch.mul(a,b) = a * b
# - Element wise division: torch.div(a,b) = a / b
# - Mean: a.mean()
# - Standart Deviation (std): a.std()
# + _uuid="66193cb3c790d13b8328c1c1262e1e3c17230bb8" _cell_guid="e43af8e7-53ab-40bc-a4f8-4cea941c6df0"
# create tensor
tensor = torch.ones(3,3)
print("\n",tensor)
# Resize
print("{}{}\n".format(tensor.view(9).shape,tensor.view(9)))
# Addition
print("Addition: {}\n".format(torch.add(tensor,tensor)))
# Subtraction
print("Subtraction: {}\n".format(tensor.sub(tensor)))
# Element wise multiplication
print("Element wise multiplication: {}\n".format(torch.mul(tensor,tensor)))
# Element wise division
print("Element wise division: {}\n".format(torch.div(tensor,tensor)))
# Mean
tensor = torch.Tensor([1,2,3,4,5])
print("Mean: {}".format(tensor.mean()))
# Standart deviation (std)
print("std: {}".format(tensor.std()))
# + [markdown] _uuid="ff85694eebe8d02701e20d7c15b0ad2974175dd3" _cell_guid="9fb8b7d4-848a-4d2c-8436-162fd47a0e11"
# ### Variables
# - It accumulates gradients.
# - We will use pytorch in neural network. And as you know, in neural network we have backpropagation where gradients are calculated. Therefore we need to handle gradients. If you do not know neural network, check my deep learning tutorial first because I will not explain detailed the concepts like optimization, loss function or backpropagation.
# - Deep learning tutorial: https://www.kaggle.com/kanncaa1/deep-learning-tutorial-for-beginners
# - Difference between variables and tensor is variable accumulates gradients.
# - We can make math operations with variables, too.
# - In order to make backward propagation we need variables
# + _uuid="83e3222b53be71e5fc7207da552ce9e9b90486dd" _cell_guid="fd8ceaa3-f1e2-4761-924e-00a6daca4a82"
# import variable from pytorch library
from torch.autograd import Variable
# define variable
var = Variable(torch.ones(3), requires_grad = True)
var
# + [markdown] _uuid="1cc3de04f98fc14624a18cceb5a84034b1dc29c7" _cell_guid="f5d54144-0753-4e2a-bac1-ccfff3084f6e"
# - Assume we have equation y = x^2
# - Define x = [2,4] variable
# - After calculation we find that y = [4,16] (y = x^2)
# - Recap o equation is that o = (1/2)*sum(y) = (1/2)*sum(x^2)
# - deriavative of o = x
# - Result is equal to x so gradients are [2,4]
# - Lets implement
# + _uuid="ff4010e2958a72ce45e43118790f3e20dc2abad6" _cell_guid="cd73c1cf-d250-48e4-bfb7-ffbe8c03c267"
# lets make basic backward propagation
# we have an equation that is y = x^2
array = [2,4]
tensor = torch.Tensor(array)
x = Variable(tensor, requires_grad = True)
y = x**2
print(" y = ",y)
# recap o equation o = 1/2*sum(y)
o = (1/2)*sum(y)
print(" o = ",o)
# backward
o.backward() # calculates gradients
# As I defined, variables accumulates gradients. In this part there is only one variable x.
# Therefore variable x should be have gradients
# Lets look at gradients with x.grad
print("gradients: ",x.grad)
# + [markdown] _uuid="6d8fa48e6e641da312175509aae00fea2760cb2c" _cell_guid="c916b8e5-e078-48de-8bc6-a757022ba65d"
# <a id="2"></a> <br>
# ### Linear Regression
# - Detailed linear regression tutorial is in my machine learning tutorial in part "Regression". I will not explain it in here detailed.
# - Linear Regression tutorial: https://www.kaggle.com/kanncaa1/machine-learning-tutorial-for-beginners
# - y = Ax + B.
# - A = slope of curve
# - B = bias (point that intersect y-axis)
# - For example, we have car company. If the car price is low, we sell more car. If the car price is high, we sell less car. This is the fact that we know and we have data set about this fact.
# - The question is that what will be number of car sell if the car price is 100.
# + _uuid="0bed0a61494fab620e639745f0c48b341f665bf8" _cell_guid="b9a3beb3-9e3c-4502-94c2-fe87ac623ca2"
# As a car company we collect this data from previous selling
# lets define car prices
car_prices_array = [3,4,5,6,7,8,9]
car_price_np = np.array(car_prices_array,dtype=np.float32)
car_price_np = car_price_np.reshape(-1,1)
car_price_tensor = Variable(torch.from_numpy(car_price_np))
# lets define number of car sell
number_of_car_sell_array = [ 7.5, 7, 6.5, 6.0, 5.5, 5.0, 4.5]
number_of_car_sell_np = np.array(number_of_car_sell_array,dtype=np.float32)
number_of_car_sell_np = number_of_car_sell_np.reshape(-1,1)
number_of_car_sell_tensor = Variable(torch.from_numpy(number_of_car_sell_np))
# lets visualize our data
import matplotlib.pyplot as plt
plt.scatter(car_prices_array,number_of_car_sell_array)
plt.xlabel("Car Price $")
plt.ylabel("Number of Car Sell")
plt.title("Car Price$ VS Number of Car Sell")
plt.show()
# + [markdown] _uuid="9e7128ffc3fa1b0e545529d5f56d969d45cab78b" _cell_guid="a6044795-9f61-4d67-a16b-8008903fd482"
# - Now this plot is our collected data
# - We have a question that is what will be number of car sell if the car price is 100$
# - In order to solve this question we need to use linear regression.
# - We need to line fit into this data. Aim is fitting line with minimum error.
# - **Steps of Linear Regression**
# 1. create LinearRegression class
# 1. define model from this LinearRegression class
# 1. MSE: Mean squared error
# 1. Optimization (SGD:stochastic gradient descent)
# 1. Backpropagation
# 1. Prediction
# - Lets implement it with Pytorch
# + _uuid="8040e01e2bdc25d6fdbff800262f3afe8b9dac3a" _cell_guid="2b74a84a-29da-44ed-9b5f-649a5c54b8a9"
# Linear Regression with Pytorch
# libraries
import torch
from torch.autograd import Variable
import torch.nn as nn
import warnings
warnings.filterwarnings("ignore")
# create class
class LinearRegression(nn.Module):
def __init__(self,input_size,output_size):
# super function. It inherits from nn.Module and we can access everythink in nn.Module
super(LinearRegression,self).__init__()
# Linear function.
self.linear = nn.Linear(input_dim,output_dim)
def forward(self,x):
return self.linear(x)
# define model
input_dim = 1
output_dim = 1
model = LinearRegression(input_dim,output_dim) # input and output size are 1
# MSE
mse = nn.MSELoss()
# Optimization (find parameters that minimize error)
learning_rate = 0.02 # how fast we reach best parameters
optimizer = torch.optim.SGD(model.parameters(),lr = learning_rate)
# train model
loss_list = []
iteration_number = 1001
for iteration in range(iteration_number):
# optimization
optimizer.zero_grad()
# Forward to get output
results = model(car_price_tensor)
# Calculate Loss
loss = mse(results, number_of_car_sell_tensor)
# backward propagation
loss.backward()
# Updating parameters
optimizer.step()
# store loss
loss_list.append(loss.data)
# print loss
if(iteration % 50 == 0):
print('epoch {}, loss {}'.format(iteration, loss.data))
plt.plot(range(iteration_number),loss_list)
plt.xlabel("Number of Iterations")
plt.ylabel("Loss")
plt.show()
# + [markdown] _uuid="6f3c764684ae00949ef3f91f5eb75b7bd11c089d" _cell_guid="b86efed7-1e47-44ad-9223-59c370a42560"
# - Number of iteration is 1001.
# - Loss is almost zero that you can see from plot or loss in epoch number 1000.
# - Now we have a trained model.
# - While usign trained model, lets predict car prices.
# + _uuid="ec56085a9c6e91f18042f8a79dede663abf09cf0" _cell_guid="abecf557-4d1f-4e2a-a466-001abb4d27d2"
# predict our car price
predicted = model(car_price_tensor).data.numpy()
plt.scatter(car_prices_array,number_of_car_sell_array,label = "original data",color ="red")
plt.scatter(car_prices_array,predicted,label = "predicted data",color ="blue")
# predict if car price is 10$, what will be the number of car sell
#predicted_10 = model(torch.from_numpy(np.array([10]))).data.numpy()
#plt.scatter(10,predicted_10.data,label = "car price 10$",color ="green")
plt.legend()
plt.xlabel("Car Price $")
plt.ylabel("Number of Car Sell")
plt.title("Original vs Predicted values")
plt.show()
# + [markdown] _uuid="20b4762eb8607ed428703c2156c5aefe8b49ff3f" _cell_guid="49344c72-d0ea-4092-96fe-ed5508ae6e0b"
# <a id="3"></a> <br>
# ### Logistic Regression
# - Linear regression is not good at classification.
# - We use logistic regression for classification.
# - linear regression + logistic function(softmax) = logistic regression
# - Check my deep learning tutorial. There is detailed explanation of logistic regression.
# - **Steps of Logistic Regression**
# 1. Import Libraries
# 1. Prepare Dataset
# - We use MNIST dataset.
# - There are 28*28 images and 10 labels from 0 to 9
# - Data is not normalized so we divide each image to 255 that is basic normalization for images.
# - In order to split data, w use train_test_split method from sklearn library
# - Size of train data is 80% and size of test data is 20%.
# - Create feature and target tensors. At the next parts we create variable from these tensors. As you remember we need to define variable for accumulation of gradients.
# - batch_size = batch size means is that for example we have data and it includes 1000 sample. We can train 1000 sample in a same time or we can divide it 10 groups which include 100 sample and train 10 groups in order. Batch size is the group size. For example, I choose batch_size = 100, that means in order to train all data only once we have 336 groups. We train each groups(336) that have batch_size(quota) 100. Finally we train 33600 sample one time.
# - epoch: 1 epoch means training all samples one time.
# - In our example: we have 33600 sample to train and we decide our batch_size is 100. Also we decide epoch is 29(accuracy achieves almost highest value when epoch is 29). Data is trained 29 times. Question is that how many iteration do I need? Lets calculate:
# - training data 1 times = training 33600 sample (because data includes 33600 sample)
# - But we split our data 336 groups(group_size = batch_size = 100) our data
# - Therefore, 1 epoch(training data only once) takes 336 iteration
# - We have 29 epoch, so total iterarion is 9744(that is almost 10000 which I used)
# - TensorDataset(): Data set wrapping tensors. Each sample is retrieved by indexing tensors along the first dimension.
# - DataLoader(): It combines dataset and sampler. It also provides multi process iterators over the dataset.
# - Visualize one of the images in dataset
# 1. Create Logistic Regression Model
# - Same with linear regression.
# - However as you expect, there should be logistic function in model right?
# - In pytorch, logistic function is in the loss function where we will use at next parts.
# 1. Instantiate Model Class
# - input_dim = 28*28 # size of image px*px
# - output_dim = 10 # labels 0,1,2,3,4,5,6,7,8,9
# - create model
# 1. Instantiate Loss Class
# - Cross entropy loss
# - It calculates loss that is not surprise :)
# - It also has softmax(logistic function) in it.
# 1. Instantiate Optimizer Class
# - SGD Optimizer
# 1. Traning the Model
# 1. Prediction
# - As a result, as you can see from plot, while loss decreasing, accuracy(almost 85%) is increasing and our model is learning(training).
# + _uuid="1382c63fe24710d3b2840e7dcf172cddbf533743" _cell_guid="a0bf0fa7-c527-4fd3-b504-02a88fc94798"
# Import Libraries
import torch
import torch.nn as nn
import torchvision.transforms as transforms
from torch.autograd import Variable
import pandas as pd
from sklearn.model_selection import train_test_split
# + _uuid="c6e0d7d3843719091564a580dbe08f67ee0d93ec" _cell_guid="59cdc9d5-da8f-4d7a-abc5-c62b0008afb0"
# Prepare Dataset
# load data
train = pd.read_csv(r"../input/train.csv",dtype = np.float32)
# split data into features(pixels) and labels(numbers from 0 to 9)
targets_numpy = train.label.values
features_numpy = train.loc[:,train.columns != "label"].values/255 # normalization
# train test split. Size of train data is 80% and size of test data is 20%.
features_train, features_test, targets_train, targets_test = train_test_split(features_numpy,
targets_numpy,
test_size = 0.2,
random_state = 42)
# create feature and targets tensor for train set. As you remember we need variable to accumulate gradients. Therefore first we create tensor, then we will create variable
featuresTrain = torch.from_numpy(features_train)
targetsTrain = torch.from_numpy(targets_train).type(torch.LongTensor) # data type is long
# create feature and targets tensor for test set.
featuresTest = torch.from_numpy(features_test)
targetsTest = torch.from_numpy(targets_test).type(torch.LongTensor) # data type is long
# batch_size, epoch and iteration
batch_size = 100
n_iters = 10000
num_epochs = n_iters / (len(features_train) / batch_size)
num_epochs = int(num_epochs)
# Pytorch train and test sets
train = torch.utils.data.TensorDataset(featuresTrain,targetsTrain)
test = torch.utils.data.TensorDataset(featuresTest,targetsTest)
# data loader
train_loader = torch.utils.data.DataLoader(train, batch_size = batch_size, shuffle = False)
test_loader = torch.utils.data.DataLoader(test, batch_size = batch_size, shuffle = False)
# visualize one of the images in data set
plt.imshow(features_numpy[10].reshape(28,28))
plt.axis("off")
plt.title(str(targets_numpy[10]))
plt.savefig('graph.png')
plt.show()
# + _uuid="7c7a7265a23a8101d5ed0c8826dfec3726d6161d" _cell_guid="03a25584-c567-4b5e-bae1-7bc9e02184fe"
# Create Logistic Regression Model
class LogisticRegressionModel(nn.Module):
def __init__(self, input_dim, output_dim):
super(LogisticRegressionModel, self).__init__()
# Linear part
self.linear = nn.Linear(input_dim, output_dim)
# There should be logistic function right?
# However logistic function in pytorch is in loss function
# So actually we do not forget to put it, it is only at next parts
def forward(self, x):
out = self.linear(x)
return out
# Instantiate Model Class
input_dim = 28*28 # size of image px*px
output_dim = 10 # labels 0,1,2,3,4,5,6,7,8,9
# create logistic regression model
model = LogisticRegressionModel(input_dim, output_dim)
# Cross Entropy Loss
error = nn.CrossEntropyLoss()
# SGD Optimizer
learning_rate = 0.001
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# + _uuid="0cab9c3ec72f73db1b06578fa7a51611141e16da" _cell_guid="82de08d9-7f3c-4eb9-8a99-9d7a8677799c"
# Traning the Model
count = 0
loss_list = []
iteration_list = []
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
# Define variables
train = Variable(images.view(-1, 28*28))
labels = Variable(labels)
# Clear gradients
optimizer.zero_grad()
# Forward propagation
outputs = model(train)
# Calculate softmax and cross entropy loss
loss = error(outputs, labels)
# Calculate gradients
loss.backward()
# Update parameters
optimizer.step()
count += 1
# Prediction
if count % 50 == 0:
# Calculate Accuracy
correct = 0
total = 0
# Predict test dataset
for images, labels in test_loader:
test = Variable(images.view(-1, 28*28))
# Forward propagation
outputs = model(test)
# Get predictions from the maximum value
predicted = torch.max(outputs.data, 1)[1]
# Total number of labels
total += len(labels)
# Total correct predictions
correct += (predicted == labels).sum()
accuracy = 100 * correct / float(total)
# store loss and iteration
loss_list.append(loss.data)
iteration_list.append(count)
if count % 500 == 0:
# Print Loss
print('Iteration: {} Loss: {} Accuracy: {}%'.format(count, loss.data, accuracy))
# + _uuid="db87c03e9d263f07eb75f82a914d3e966895a6c1" _cell_guid="924e9606-e155-4e39-89d3-39f941fd52f8"
# visualization
plt.plot(iteration_list,loss_list)
plt.xlabel("Number of iteration")
plt.ylabel("Loss")
plt.title("Logistic Regression: Loss vs Number of iteration")
plt.show()
# + [markdown] _uuid="ea9eba414f2f0f1e63ef564dc0ee708c753ff51f" _cell_guid="4d38db05-fad0-468c-9000-20caf5465eca"
# <a id="4"></a> <br>
# ### Artificial Neural Network (ANN)
# - Logistic regression is good at classification but when complexity(non linearity) increases, the accuracy of model decreases.
# - Therefore, we need to increase complexity of model.
# - In order to increase complexity of model, we need to add more non linear functions as hidden layer.
# - I am saying again that if you do not know what is artificial neural network check my deep learning tutorial because I will not explain neural network detailed here, only explain pytorch.
# - Artificial Neural Network tutorial: https://www.kaggle.com/kanncaa1/deep-learning-tutorial-for-beginners
# - What we expect from artificial neural network is that when complexity increases, we use more hidden layers and our model can adapt better. As a result accuracy increase.
# - **Steps of ANN:**
# 1. Import Libraries
# - In order to show you, I import again but we actually imported them at previous parts.
# 1. Prepare Dataset
# - Totally same with previous part(logistic regression).
# - We use same dataset so we only need train_loader and test_loader.
# - We use same batch size, epoch and iteration numbers.
# 1. Create ANN Model
# - We add 3 hidden layers.
# - We use ReLU, Tanh and ELU activation functions for diversity.
# 1. Instantiate Model Class
# - input_dim = 28*28 # size of image px*px
# - output_dim = 10 # labels 0,1,2,3,4,5,6,7,8,9
# - Hidden layer dimension is 150. I only choose it as 150 there is no reason. Actually hidden layer dimension is hyperparameter and it should be chosen and tuned. You can try different values for hidden layer dimension and observe the results.
# - create model
# 1. Instantiate Loss Class
# - Cross entropy loss
# - It also has softmax(logistic function) in it.
# 1. Instantiate Optimizer Class
# - SGD Optimizer
# 1. Traning the Model
# 1. Prediction
# - As a result, as you can see from plot, while loss decreasing, accuracy is increasing and our model is learning(training).
# - Thanks to hidden layers model learnt better and accuracy(almost 95%) is better than accuracy of logistic regression model.
# + _uuid="cf25ee4b28129a47bac4c9dc7d932295155b79f7" _cell_guid="6925f8ed-54b7-4d9a-9801-acd65f213bc9"
# Import Libraries
import torch
import torch.nn as nn
import torchvision.transforms as transforms
from torch.autograd import Variable
# + _uuid="cefd0bb2f23b80f30ca65cbb08859ad81ab12e08" _cell_guid="3472f1c1-5888-4abe-822c-3a493a5f8be5"
# Create ANN Model
class ANNModel(nn.Module):
def __init__(self, input_dim, hidden_dim, output_dim):
super(ANNModel, self).__init__()
# Linear function 1: 784 --> 100
self.fc1 = nn.Linear(input_dim, hidden_dim)
# Non-linearity 1
self.relu1 = nn.ReLU()
# Linear function 2: 100 --> 100
self.fc2 = nn.Linear(hidden_dim, hidden_dim)
# Non-linearity 2
self.tanh2 = nn.Tanh()
# Linear function 3: 100 --> 100
self.fc3 = nn.Linear(hidden_dim, hidden_dim)
# Non-linearity 3
self.elu3 = nn.ELU()
# Linear function 4 (readout): 100 --> 10
self.fc4 = nn.Linear(hidden_dim, output_dim)
def forward(self, x):
# Linear function 1
out = self.fc1(x)
# Non-linearity 1
out = self.relu1(out)
# Linear function 2
out = self.fc2(out)
# Non-linearity 2
out = self.tanh2(out)
# Linear function 2
out = self.fc3(out)
# Non-linearity 2
out = self.elu3(out)
# Linear function 4 (readout)
out = self.fc4(out)
return out
# instantiate ANN
input_dim = 28*28
hidden_dim = 150 #hidden layer dim is one of the hyper parameter and it should be chosen and tuned. For now I only say 150 there is no reason.
output_dim = 10
# Create ANN
model = ANNModel(input_dim, hidden_dim, output_dim)
# Cross Entropy Loss
error = nn.CrossEntropyLoss()
# SGD Optimizer
learning_rate = 0.02
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# + _uuid="c91694f3af94e4e1b76ab01489e186718c70ccd3" _cell_guid="7550e98b-5011-4d09-88ee-97b0ecbc6f19"
# ANN model training
count = 0
loss_list = []
iteration_list = []
accuracy_list = []
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
train = Variable(images.view(-1, 28*28))
labels = Variable(labels)
# Clear gradients
optimizer.zero_grad()
# Forward propagation
outputs = model(train)
# Calculate softmax and ross entropy loss
loss = error(outputs, labels)
# Calculating gradients
loss.backward()
# Update parameters
optimizer.step()
count += 1
if count % 50 == 0:
# Calculate Accuracy
correct = 0
total = 0
# Predict test dataset
for images, labels in test_loader:
test = Variable(images.view(-1, 28*28))
# Forward propagation
outputs = model(test)
# Get predictions from the maximum value
predicted = torch.max(outputs.data, 1)[1]
# Total number of labels
total += len(labels)
# Total correct predictions
correct += (predicted == labels).sum()
accuracy = 100 * correct / float(total)
# store loss and iteration
loss_list.append(loss.data)
iteration_list.append(count)
accuracy_list.append(accuracy)
if count % 500 == 0:
# Print Loss
print('Iteration: {} Loss: {} Accuracy: {} %'.format(count, loss.data[0], accuracy))
# + _uuid="c5e2e6da7f1ee801e38358dc28d4c99e32d2b761" _cell_guid="5579a7d6-7766-4d0f-b9d0-584cb4f28321"
# visualization loss
plt.plot(iteration_list,loss_list)
plt.xlabel("Number of iteration")
plt.ylabel("Loss")
plt.title("ANN: Loss vs Number of iteration")
plt.show()
# visualization accuracy
plt.plot(iteration_list,accuracy_list,color = "red")
plt.xlabel("Number of iteration")
plt.ylabel("Accuracy")
plt.title("ANN: Accuracy vs Number of iteration")
plt.show()
# + [markdown] _uuid="cd8f261d231acaccd0f0bc8466fc28c1b0c2f567" _cell_guid="50bbb2e7-15d8-47f5-8f31-25e0d0cb9e29"
# <a id="5"></a> <br>
# ### Convolutional Neural Network (CNN)
# - CNN is well adapted to classify images.
# - You can learn CNN basics and concepts from Pourya's tutorial: https://www.kaggle.com/pouryaayria/convolutional-neural-networks-tutorial-tensorflow
# - **Steps of CNN:**
# 1. Import Libraries
# 1. Prepare Dataset
# - Totally same with previous parts.
# - We use same dataset so we only need train_loader and test_loader.
# 1. Convolutional layer:
# - Create feature maps with filters(kernels).
# - Padding: After applying filter, dimensions of original image decreases. However, we want to preserve as much as information about the original image. We can apply padding to increase dimension of feature map after convolutional layer.
# - We use 2 convolutional layer.
# - Number of feature map is out_channels = 16
# - Filter(kernel) size is 5*5
# 1. Pooling layer:
# - Prepares a condensed feature map from output of convolutional layer(feature map)
# - 2 pooling layer that we will use max pooling.
# - Pooling size is 2*2
# 1. Flattening: Flats the features map
# 1. Fully Connected Layer:
# - Artificial Neural Network that we learnt at previous part.
# - Or it can be only linear like logistic regression but at the end there is always softmax function.
# - We will not use activation function in fully connected layer.
# - You can think that our fully connected layer is logistic regression.
# - We combine convolutional part and logistic regression to create our CNN model.
# 1. Instantiate Model Class
# - create model
# 1. Instantiate Loss Class
# - Cross entropy loss
# - It also has softmax(logistic function) in it.
# 1. Instantiate Optimizer Class
# - SGD Optimizer
# 1. Traning the Model
# 1. Prediction
# - As a result, as you can see from plot, while loss decreasing, accuracy is increasing and our model is learning(training).
# - Thanks to convolutional layer, model learnt better and accuracy(almost 98%) is better than accuracy of ANN. Actually while tuning hyperparameters, increase in iteration and expanding convolutional neural network can increase accuracy but it takes too much running time that we do not want at kaggle.
#
# + _uuid="c78fdf4401cb34db0d963df0249bf92144bc5fbd" _cell_guid="abbc86d1-f677-4d4b-9b7b-09c7f9962288"
# Import Libraries
import torch
import torch.nn as nn
import torchvision.transforms as transforms
from torch.autograd import Variable
# + _uuid="4915535771ffdd33ef480200393216f215b4fc48" _cell_guid="9ca5af9e-6821-4d60-8084-edb523a39c6b"
# Create CNN Model
class CNNModel(nn.Module):
def __init__(self):
super(CNNModel, self).__init__()
# Convolution 1
self.cnn1 = nn.Conv2d(in_channels=1, out_channels=16, kernel_size=5, stride=1, padding=0)
self.relu1 = nn.ReLU()
# Max pool 1
self.maxpool1 = nn.MaxPool2d(kernel_size=2)
# Convolution 2
self.cnn2 = nn.Conv2d(in_channels=16, out_channels=32, kernel_size=5, stride=1, padding=0)
self.relu2 = nn.ReLU()
# Max pool 2
self.maxpool2 = nn.MaxPool2d(kernel_size=2)
# Fully connected 1
self.fc1 = nn.Linear(32 * 4 * 4, 10)
def forward(self, x):
# Convolution 1
out = self.cnn1(x)
out = self.relu1(out)
# Max pool 1
out = self.maxpool1(out)
# Convolution 2
out = self.cnn2(out)
out = self.relu2(out)
# Max pool 2
out = self.maxpool2(out)
out = out.view(out.size(0), -1)
# Linear function (readout)
out = self.fc1(out)
return out
# batch_size, epoch and iteration
batch_size = 100
n_iters = 2500
num_epochs = n_iters / (len(features_train) / batch_size)
num_epochs = int(num_epochs)
# Pytorch train and test sets
train = torch.utils.data.TensorDataset(featuresTrain,targetsTrain)
test = torch.utils.data.TensorDataset(featuresTest,targetsTest)
# data loader
train_loader = torch.utils.data.DataLoader(train, batch_size = batch_size, shuffle = False)
test_loader = torch.utils.data.DataLoader(test, batch_size = batch_size, shuffle = False)
# Create ANN
model = CNNModel()
# Cross Entropy Loss
error = nn.CrossEntropyLoss()
# SGD Optimizer
learning_rate = 0.1
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# + _uuid="f44e02d25698ac1a014795d972a384a3f3003d35" _cell_guid="99a8903c-da15-496c-96b7-f5402c8fc5f0"
# CNN model training
count = 0
loss_list = []
iteration_list = []
accuracy_list = []
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
train = Variable(images.view(100,1,28,28))
labels = Variable(labels)
# Clear gradients
optimizer.zero_grad()
# Forward propagation
outputs = model(train)
# Calculate softmax and ross entropy loss
loss = error(outputs, labels)
# Calculating gradients
loss.backward()
# Update parameters
optimizer.step()
count += 1
if count % 50 == 0:
# Calculate Accuracy
correct = 0
total = 0
# Iterate through test dataset
for images, labels in test_loader:
test = Variable(images.view(100,1,28,28))
# Forward propagation
outputs = model(test)
# Get predictions from the maximum value
predicted = torch.max(outputs.data, 1)[1]
# Total number of labels
total += len(labels)
correct += (predicted == labels).sum()
accuracy = 100 * correct / float(total)
# store loss and iteration
loss_list.append(loss.data)
iteration_list.append(count)
accuracy_list.append(accuracy)
if count % 500 == 0:
# Print Loss
print('Iteration: {} Loss: {} Accuracy: {} %'.format(count, loss.data[0], accuracy))
# + _uuid="44c1ed412d778f3e6b08f11bddc5321f63e408dd" _cell_guid="ac9e4aee-b8af-4641-8794-bad03b650179"
# visualization loss
plt.plot(iteration_list,loss_list)
plt.xlabel("Number of iteration")
plt.ylabel("Loss")
plt.title("CNN: Loss vs Number of iteration")
plt.show()
# visualization accuracy
plt.plot(iteration_list,accuracy_list,color = "red")
plt.xlabel("Number of iteration")
plt.ylabel("Accuracy")
plt.title("CNN: Accuracy vs Number of iteration")
plt.show()
# + [markdown] _uuid="6ce8edd6d5fbf0019d6ab189f8ed1fd9cde0ac85" _cell_guid="a508a4c8-0719-4a8f-847c-bfce1ceef30f"
# ### Conclusion
# In this tutorial, we learn:
# 1. Basics of pytorch
# 1. Linear regression with pytorch
# 1. Logistic regression with pytorch
# 1. Artificial neural network with with pytorch
# 1. Convolutional neural network with pytorch
# 1. Recurrent neural network with pytorch
# - https://www.kaggle.com/kanncaa1/recurrent-neural-network-with-pytorch
#
# <br> **If you have any question or suggest, I will be happy to hear it **
# + _uuid="213619d278384044e3b0dd8577057413a9a7e408" _cell_guid="4c40d91b-8a9c-456c-9300-a6686792c424"
| 2 digit recognizer/pytorch-tutorial-for-deep-learning-lovers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from datetime import datetime
import pandas as pd
from pathlib import Path
import pickle
data_dir = Path('../data')
csvs = [x for x in data_dir.iterdir() if 'timeseries' in str(x)]
counts = []
regions = []
results = pd.DataFrame(columns=['roi', 'new_deaths', 'new_cases', 'new_recover', 'abs_days', 'rel_days'])
i = 1
for csv in csvs:
region = str(csv).split('.')[0].split('_')[-1]
regions.append(region)
df = pd.read_csv(csv)
df = df.iloc[:-1]
df = df[df['cum_cases']>=10]
df['abs_days'] = df['dates2'].apply(lambda x: (datetime.strptime(x, '%m/%d/%y') - datetime.strptime('01/22/20', '%m/%d/%y')).days)
df['rel_days'] = df['dates2'].apply(lambda x: (datetime.strptime(x, '%m/%d/%y') - datetime.strptime(df['dates2'].values[0], '%m/%d/%y')).days)
counts.append(df.shape[0])
subset = ['new_deaths', 'new_cases', 'new_recover']
df[subset] = df[subset].clip(lower = 0)
df['roi'] = i
subset = ['roi', 'new_deaths', 'new_cases', 'new_recover', 'abs_days', 'rel_days']
results = results.append(df[subset])
i += 1
import pickle
with open(data_dir / 'stacked.pkl', 'wb') as f:
pickle.dump({'data': results, 'regions': regions, 'counts': counts}, f)
results
| notebooks/stacked.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch35
# language: python
# name: torch35
# ---
# # Input
category = "Blog"
folder_name = "2020-11-17-jekyll-local-server-start-batch"
title = " 제목 "
excerpt = " 부제 "
toc_label = " toclabel "
pre_post = ""
# # Preparation
if category == "DS":
category_full_name = "Data science"
pre_post_category = "data%20science"
tags = ["Data science"]
elif category == "DL":
category_full_name = "Deep learning"
pre_post_category = "deep%learning"
tags = ["Deep learninig"]
elif category == "ML":
category_full_name = "Machine learning"
pre_post_category = "machine%learning"
tags = ["Machine learninig"]
elif category == "MEDIA":
category_full_name = "Medical image analysis"
pre_post_category = "medical%20image%20analysis"
tags = ["Medical image analysis", "Lecture"]
elif category == "Python":
category_full_name = "Python"
pre_post_category = "python"
tags = ["Python"]
elif category == "Blog":
category_full_name = "Blog"
pre_post_category = "blog"
tags = ["Blog"]
img_dir = "./assets/images/post"
post_dir = "./_posts"
draft_dir = "./_drafts"
# # YAML
# +
data = "---\n"
data+= "title: \"" + title + "\"\n"
data+= "excerpt: \"" + excerpt + "\"\n"
data+= "\n"
data+= "categories:\n"
data+= "- " + category_full_name + "\n"
data+= "\n"
data+= "tags:\n"
for tag in tags:
data+= "- " + tag + "\n"
data+= "\n"
data+= "toc: true\n"
data+= "toc_sticky: true\n"
data+= "toc_label: \"" + toc_label + "\"\n"
data+= "\n"
data+= "use_math: true\n"
data+= "---"
data+= "\n\n"
# -
# # 이전 포스트
if pre_post != "":
f = open(post_dir + "/" + pre_post + ".md", "r", encoding='UTF-8')
f.readline()
pre_post_title = f.readline()
while(1):
line = f.readline()
if line == "---\n":
break
f.readline() # \n
f.readline() # 이전 포스팅
f.readline() # \n
f.readline() # > 이전 포스팅에서는
pre_post_content = f.readline()
f.close()
pre_post_title = pre_post_title[8:-2]
pre_post_content = "> 이전" + pre_post_content[4:-10] + "했습니다 \n"
data+= "이전 포스팅: [" + pre_post_title + "]({{ site.url }}{{ site.baseurl }}/" + pre_post_category + "/" + pre_post[11:] + "/)" + "\n"
data+= "\n"
data+= pre_post_content
data+= "> 이번 포스팅에서는 ** 내용 **을 정리해보고자 합니다.\n"
data+= "\n\n---\n\n"
# # Images
# +
import glob
import os
path = img_dir + "/" + category + "/" + folder_name + "/"
valid_img = [".jpg", ".gif", ".png"]
img_names = glob.glob(path + "*.png")
img_names = [os.path.basename(full_path) for full_path in img_names]
# -
img_names
for img_name in img_names:
data+= "\n\n"
# # 끝
# +
data+= "---\n\n"
data+= "> 다음 포스팅에서는 ** 내용 **을 정리해보고자 합니다\n\n"
data+= "다음 포스팅: [ 작성중 ]({{ site.url }}{{ site.baseurl }}/" + pre_post_category + "/ 작성중 /)"
# -
print(data)
# # md 파일 작성
img_dir + "/" + category + "/" + folder_name + "/" + folder_name + ".md"
f = open(draft_dir + "/" + folder_name + ".md", "w", encoding='UTF-8')
f.write(data)
f.close()
| .ipynb_checkpoints/md-generator-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Booleans
help(bool)
issubclass(bool, int)
type(True), id(True), int(True)
type(False), id(False), int(False)
3 < 4
type(3 < 4)
id(3 < 4)
(3 < 4) == True
(3 < 4) is True
None is False
(1 == 2) == False
1 == 2 == False
1 == 2 and 2 == False
int(True), int(False)
1 + True
100 * False
True > False
(True + True + True) % 2
-True
True and False
True or False
bool(0)
bool(1)
int(True)
bool(True)
bool(False)
bool(100)
bool(-1)
bool(0)
| my_classes/NumericTypes/booleans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# Libraries
import pandas as pd
import time
import datetime as dt
import ipynb.fs.full.market_ingestion_coinmarketcap
from ipynb.fs.full.market_ingestion_coinmarketcap import MarketListener
import ipynb.fs.full.market_register_coinmarketcap
from ipynb.fs.full.market_register_coinmarketcap import MarketRegister
import ipynb.fs.full.market_register_coinmarketcap_s
from ipynb.fs.full.market_register_coinmarketcap_s import SummaryMarketRegister
# +
class MarketProcessing():
"""
The MarketProcessing class is used to create the market data by adding features
"""
def __init__(self,params_market = []):
self.data_market = pd.DataFrame(params_market[0])
self.data_summary_market = pd.DataFrame(params_market[1])
def drop_duplicates_rows(data,colnames):
"""
Drop the duplicates rows
"""
return data.drop_duplicates(subset=colnames)
def reset_index(data):
"""
Reset the index
"""
data = data.reset_index()
#data = data.drop('index',1)
return data
def drop_columns(data,colnames):
"""
Drop the columns
"""
return data.drop(colnames,axis=1)
def rename_columns(data,colnames):
"""
Rename the columns by the colnames
"""
return data.rename(index=str,columns=colnames)
def add_datetime(data):
"""
Add datetime features
"""
data_datetime = pd.DataFrame()
data_datetime['datetime'] = [dt.datetime.utcfromtimestamp(data.iloc[i]["last_updated"]) for i in range(len(data["last_updated"]))]
data_datetime['year'] = [x.year for x in data_datetime['datetime']]
data_datetime['month'] = [x.month for x in data_datetime['datetime']]
data_datetime['day'] = [x.day for x in data_datetime['datetime']]
data_datetime['hour'] = [x.hour for x in data_datetime['datetime']]
data_datetime['minute'] = [x.minute for x in data_datetime['datetime']]
data_datetime['second'] = [x.second for x in data_datetime['datetime']]
return data_datetime
def add_date(data):
"""
Add the date
"""
return [dt.datetime(data.iloc[i]['year'],data.iloc[i]['month'],data.iloc[i]['day'],data.iloc[i]['hour']) for i in range(len(data))]
def add_statistics(data,vargrouped,varnames):
"""
Add the descriptive statistics features
"""
data_stats = pd.DataFrame()
stats = data.groupby(vargrouped).agg(['last','max','mean','median','std','min','first'])
statnames = ['last','max','mean','median','std','min','first']
for var in varnames:
for stat in statnames:
data_stats[stat+'_'+var] = stats[var][stat]
data_stats.reset_index(inplace=True)
return data_stats
def add_percent_currency_marketcap(self):
"""
Add the percent of the currency on the marketcap
"""
return 100 * self.data_market["market_cap_usd"] / self.data_market["total_market_cap_usd"]
def add_indicators():
"""
Add the trading indicators
<TO DO>
"""
return None
def df_aggregate_by_id(data):
"""
Create a list of the dataframe aggregated by id
"""
data_aggregated = []
list_id = list(set(data['id']))
for id in list_id:
data_aggregated.append(data[data['id']==id])
return data_aggregated
def add_duplicated_rows(self):
"""
Add the duplicated rows
"""
list_id_dupplicated = list(set(self.data_market['id'])) * len(self.data_summary_market)
self.data_summary_market = self.data_summary_market.append([self.data_summary_market] * (len(set(self.data_market['id']))-1),ignore_index=True)
self.data_summary_market = self.data_summary_market.sort_values(['date'])
self.data_summary_market['id'] = list_id_dupplicated
return self.data_summary_market
def build_data_market(self):
"""
Build the dataframe with the market
"""
# drop duplicates rows
self.data_market = MarketProcessing.drop_duplicates_rows(self.data_market,["id","last_updated"])
self.data_market = MarketProcessing.reset_index(self.data_market)
# add datetime to the market data
#--
datetime = MarketProcessing.add_datetime(self.data_market)
self.data_market[['datetime','year','month','day','hour','minute','second']] = datetime
# add statistics to the market data
#--
stock = self.data_market[['id','year','month','day','hour','price_btc','price_usd','percent_change_1h','percent_change_7d',
'volume_24h_usd','percent_change_24h','max_supply','total_supply']]
statistics = MarketProcessing.add_statistics(stock,vargrouped=['id','year','month','day','hour'],varnames=['price_btc','price_usd','percent_change_1h','percent_change_7d','volume_24h_usd','percent_change_24h','max_supply','total_supply'])
# drop duplicates rows
self.data_market = MarketProcessing.drop_duplicates_rows(self.data_market,["id","year","month","day","hour"])
self.data_market = MarketProcessing.reset_index(self.data_market)
self.data_market = self.data_market.merge(statistics,on=["id","year","month","day","hour"],how="left")
#--
# add date to the market data
self.data_market['date'] = MarketProcessing.add_date(self.data_market)
colnames = ['level_0','index','last_updated','datetime','minute','second','price_btc','price_usd','percent_change_1h','percent_change_7d','volume_24h_usd','percent_change_24h','max_supply','total_supply']
self.data_market = MarketProcessing.drop_columns(self.data_market,colnames)
return self.data_market
def build_data_summary_market(self):
"""
Build the dataframe with the market summary
"""
# drop duplicates rows
self.data_summary_market = MarketProcessing.drop_duplicates_rows(self.data_summary_market,["last_updated"])
self.data_summary_market = MarketProcessing.reset_index(self.data_summary_market)
# add datetime to the summary market data
#--
datetime = MarketProcessing.add_datetime(self.data_summary_market)
self.data_summary_market[['datetime','year','month','day','hour','minute','second']] = datetime
self.data_summary_market = MarketProcessing.drop_duplicates_rows(self.data_summary_market,['year','month','day','hour'])
self.data_summary_market = MarketProcessing.reset_index(self.data_summary_market)
colnames = ['level_0','index','last_updated','datetime','minute','second','bitcoin_percentage_of_market_cap']
self.data_summary_market = MarketProcessing.drop_columns(self.data_summary_market,colnames)
#--
return self.data_summary_market
def featurize(self):
"""
Create the features
"""
# build the data_market
self.data_market = MarketProcessing.build_data_market(self)
# build the data_summary_market
self.data_summary_market = MarketProcessing.build_data_summary_market(self)
# merge the data_market with data_summary_market
self.data_market = self.data_market.merge(self.data_summary_market,on=["year","month","day","hour"],how="left")
# add percent_currency_market_cap_usd to the features
self.data_market['percent_currency_market_cap_usd'] = MarketProcessing.add_percent_currency_marketcap(self)
colnames = {'cached_x','cached_y','year','month','day','hour'}
self.data_market = MarketProcessing.drop_columns(self.data_market,colnames)
return self.data_market
def put_csv(self,data,filename,type_mode="w"):
"""
Export the data to a csv file
"""
data.to_csv(filename,encoding="utf-8",mode=type_mode)
return
# -
| market_processing_coinmarketcap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Warvito/Normative-modelling-using-deep-autoencoders/blob/master/notebooks/predict.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="YVZp-35UpNAK"
# # **Deviation scores using normative models based on deep autoencoders**
#
# Here in this notebook, we implemented an easy way to you try our normative models trained on the [UK Biobank](https://www.ukbiobank.ac.uk/) dataset.
#
# **Disclaimer**: this script can not be used for clinical purposes.
#
# Let's start!
#
# ---
# + [markdown] id="S-X8m7t2iuNK" colab_type="text"
# ## Set this notebook's hardware accelerator
#
# First, you'll need to enable the use of Google's [GPUs](https://cloud.google.com/gpu) (graphics processing unit) for this notebook:
#
# - Navigate to Edit→Notebook Settings
# - Select GPU from the Hardware Accelerator drop-down
#
# These GPUs allow us to perform the deep learning model's calculation in a faster way!
# + [markdown] id="O-TAm0sAip7s" colab_type="text"
# ---
#
# + [markdown] colab_type="text" id="rUzvR_q4tFKV"
# ## Download trained models
# Next, we will load the trained normative models based on adversarial autoencoders into this colab environment. During our study, we trained normative models on the UK Biobank using the resampling method called bootstrap method. By using this resampling method, we trained 1,000 different models, each one using a different bootstraped datasets as training set (containing 11,032 brain scans) (check Section 2.4. Normative model training of our paper for more information).
#
# <center><img src="https://github.com/Warvito/Normative-modelling-using-deep-autoencoders/blob/master/notebooks/figures/aae.png?raw=1" height="500">
#
# <small>Structure of the normative model based on adversarial autoencoders. In this configuration, the subject data is inputted into the encoder and then mapped to the latent code. This latent code is fed to the decoder with the demographic data, and then the decoder generates a reconstruction of the original data. During the training of the model, the discriminator is used to shape the distribution of the latent code. Since the model is trained on healthy controls data, it can reconstruct similar data relatively well, yielding to a small reconstruction error. However, the model would generate a high error when processing data affected by unseen underlying mechanisms, e.g. pathological mechanisms.</small></center>
#
# + [markdown] id="QWAMDjEFhgY4" colab_type="text"
#
# For each normative model, we had others auxiliary components, like data scalers and demographic data preprocessors. During training, all these components were stored and are available at https://www.dropbox.com/s/bs89t2davs1p2dm/models_for_normative_paper_2019.zip?dl=0 . This link contains a compressed file that have all files created using the [bootstrap_train_aae_supervised.py](https://github.com/Warvito/Normative-modelling-using-deep-autoencoders/blob/master/bootstrap_train_aae_supervised.py) script. The models files are organized in subdirectories where each one correspond to a bootstrap iteration.
#
# Besides the models, the zipped file contains two templates files (used later in this notebook).
#
# In the following cell, we download the compressed file.
# + colab_type="code" id="xUw80taSTjWW" outputId="8d864df2-9be5-445b-9d7f-6ccf9335bed9" colab={"base_uri": "https://localhost:8080/", "height": 454}
# !wget -O models.zip --no-check-certificate https://www.dropbox.com/s/bs89t2davs1p2dm/models_for_normative_paper_2019.zip?dl=0
# + [markdown] id="oJS0hGA8inYb" colab_type="text"
# ---
#
# + [markdown] colab_type="text" id="eHi0frcUvjWV"
# ## Unzip models files
#
# After downloaded the compressed file, we need to unzip it in our colab enviroment.
# + colab_type="code" id="TOBm7DBZWu-i" colab={}
# !unzip models.zip
# + [markdown] colab_type="text" id="b-FkF6BDvsgV"
# To see the unzipped models, go to “Files” in the Google colab environment. If the Google colab environment is not shown, click in the arrow mark which looks like “>” at the left-hand side of the cells. When you click that you will find a tab with three options, just select “Files” to explore the loaded unzipped models.
#
# <img src="https://github.com/Warvito/Normative-modelling-using-deep-autoencoders/blob/master/notebooks/figures/files.png?raw=1">
#
#
#
#
#
# + [markdown] id="iSowey5Cilh_" colab_type="text"
# ---
#
# + [markdown] colab_type="text" id="hjjJikyJwniF"
# ## Import Python libraries
# Now, we will start to use the necessary Python code to make our predictions. But first let's import all the necessary Python modules for our processing.
# + colab_type="code" id="92qHTRkXYOZS" outputId="9ffbee48-8514-47c7-975b-1b376986cc25" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
from pathlib import Path
import warnings
import joblib
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
from google.colab import files
from tqdm import tqdm
# + [markdown] id="rVLmC4y2jEak" colab_type="text"
#
# ---
#
#
# + [markdown] colab_type="text" id="CLo1uoS8wytB"
# ## Download freesurferData.csv and participants.tsv templates
# In order to make predictions of your data, it is necessary to make it in the format to correctly read by this script. To facilitate this process, we supply the template files to be filled with your data.
#
# As shown below, these template files contain the names of the necessary columns to run the script.
# + colab_type="code" id="QO5bnmgYxi7y" outputId="02042abb-a041-41d1-c885-31dd86994cd3" colab={"base_uri": "https://localhost:8080/", "height": 182}
pd.read_csv('templates/freesurferData.csv')
# + colab_type="code" id="pKl3fXWgxs8q" outputId="fbac724f-925e-4da7-af06-c97e36f10320" colab={"base_uri": "https://localhost:8080/", "height": 81}
pd.read_csv('templates/participants.tsv', sep='\t')
# + [markdown] colab_type="text" id="8vlfd-XWxxZv"
# * Note: The column with gender is codified as 0 = "Female" and 1 = "Male".
#
# The next cells will start the download of the templates.
#
# ---
#
#
# + colab_type="code" id="jUub_LWSZ-FW" colab={}
files.download('templates/freesurferData.csv')
# + colab_type="code" id="CC_YY2N-dJ9G" colab={}
files.download('templates/participants.tsv')
# + [markdown] colab_type="text" id="3oPjKCaPx5qu"
# After filled the templates, upload the files to the Google colab environment.
#
# **Note: You can create the freesurferData.csv file using our colab script on this** [link](https://colab.research.google.com/github/Warvito/Normative-modelling-using-deep-autoencoders/blob/master/notebooks/freesurfer_organizer.ipynb).
#
# Note2: Your data will only be loaded in this runtime of the Google colab. This code is being executed at the Google Cloud Platform by default, and you are not making your data available for our team. If you are concern about uploading your data to the Google Cloud Platform, please, consider executing this notebook in a local runtime in your computer (https://research.google.com/colaboratory/local-runtimes.html).
# + [markdown] colab_type="text" id="UOlTkfiwzg47"
# First, start uploading the freesurferData.csv.
# + colab_type="code" id="gujhqPurZpxn" outputId="2c8b56c8-80af-465b-fbd4-f63ce4388dcf" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 117}
# Remove freesurferData.csv if it exists
# !rm freesurferData.csv
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(name=fn, length=len(uploaded[fn])))
# + id="gVTY79Vf3W_r" colab_type="code" outputId="2a2ed866-84e9-461b-dbf1-b0bffb628ed8" colab={"base_uri": "https://localhost:8080/", "height": 182}
freesurfer_data_df = pd.read_csv(fn)
freesurfer_data_df
# + [markdown] colab_type="text" id="_-l3-DNczoC5"
# Then, upload the participants.tsv file.
# + colab_type="code" id="whliHlezdPVr" outputId="86b8aa5b-9a30-4ded-a0e7-2dff83df1181" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 117}
# Remove participants.tsv if it exists
# !rm participants.tsv
uploaded = files.upload()
for fn2 in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(name=fn2, length=len(uploaded[fn2])))
# + id="3e0FIBOv3hZ1" colab_type="code" outputId="87f6fd5f-7a12-4dd0-d148-71ed665bd6a7" colab={"base_uri": "https://localhost:8080/", "height": 81}
participants_df = pd.read_csv(fn2, sep='\t')
participants_df
# + colab_type="code" id="X3dIqTAdbwTt" outputId="d4931d2b-ee8c-48d2-dc06-ee7d54641325" colab={"base_uri": "https://localhost:8080/", "height": 182}
dataset_df = pd.merge(freesurfer_data_df, participants_df, on='Participant_ID')
dataset_df
# + [markdown] id="uIsvAXKojRPe" colab_type="text"
#
#
# ---
#
#
#
# + [markdown] colab_type="text" id="qPT20CsLzx83"
# ## Predict the deviation scores
# After loading the data, we predict the deviations of the new data based on our trained normative models.
#
# We begin the processing by setting the random seeds.
# + colab_type="code" id="H4FmSLM5aq5q" colab={}
# Set random seed
random_seed = 42
tf.random.set_seed(random_seed)
np.random.seed(random_seed)
# + [markdown] colab_type="text" id="HJScNag1ik3e"
# Next, we define the name of the brain regions in the variable COLUMNS_NAME.
# + colab_type="code" id="N9ymq42YirBj" cellView="form" colab={}
#@title
COLUMNS_NAME = ['Left-Lateral-Ventricle',
'Left-Inf-Lat-Vent',
'Left-Cerebellum-White-Matter',
'Left-Cerebellum-Cortex',
'Left-Thalamus-Proper',
'Left-Caudate',
'Left-Putamen',
'Left-Pallidum',
'3rd-Ventricle',
'4th-Ventricle',
'Brain-Stem',
'Left-Hippocampus',
'Left-Amygdala',
'CSF',
'Left-Accumbens-area',
'Left-VentralDC',
'Right-Lateral-Ventricle',
'Right-Inf-Lat-Vent',
'Right-Cerebellum-White-Matter',
'Right-Cerebellum-Cortex',
'Right-Thalamus-Proper',
'Right-Caudate',
'Right-Putamen',
'Right-Pallidum',
'Right-Hippocampus',
'Right-Amygdala',
'Right-Accumbens-area',
'Right-VentralDC',
'CC_Posterior',
'CC_Mid_Posterior',
'CC_Central',
'CC_Mid_Anterior',
'CC_Anterior',
'lh_bankssts_volume',
'lh_caudalanteriorcingulate_volume',
'lh_caudalmiddlefrontal_volume',
'lh_cuneus_volume',
'lh_entorhinal_volume',
'lh_fusiform_volume',
'lh_inferiorparietal_volume',
'lh_inferiortemporal_volume',
'lh_isthmuscingulate_volume',
'lh_lateraloccipital_volume',
'lh_lateralorbitofrontal_volume',
'lh_lingual_volume',
'lh_medialorbitofrontal_volume',
'lh_middletemporal_volume',
'lh_parahippocampal_volume',
'lh_paracentral_volume',
'lh_parsopercularis_volume',
'lh_parsorbitalis_volume',
'lh_parstriangularis_volume',
'lh_pericalcarine_volume',
'lh_postcentral_volume',
'lh_posteriorcingulate_volume',
'lh_precentral_volume',
'lh_precuneus_volume',
'lh_rostralanteriorcingulate_volume',
'lh_rostralmiddlefrontal_volume',
'lh_superiorfrontal_volume',
'lh_superiorparietal_volume',
'lh_superiortemporal_volume',
'lh_supramarginal_volume',
'lh_frontalpole_volume',
'lh_temporalpole_volume',
'lh_transversetemporal_volume',
'lh_insula_volume',
'rh_bankssts_volume',
'rh_caudalanteriorcingulate_volume',
'rh_caudalmiddlefrontal_volume',
'rh_cuneus_volume',
'rh_entorhinal_volume',
'rh_fusiform_volume',
'rh_inferiorparietal_volume',
'rh_inferiortemporal_volume',
'rh_isthmuscingulate_volume',
'rh_lateraloccipital_volume',
'rh_lateralorbitofrontal_volume',
'rh_lingual_volume',
'rh_medialorbitofrontal_volume',
'rh_middletemporal_volume',
'rh_parahippocampal_volume',
'rh_paracentral_volume',
'rh_parsopercularis_volume',
'rh_parsorbitalis_volume',
'rh_parstriangularis_volume',
'rh_pericalcarine_volume',
'rh_postcentral_volume',
'rh_posteriorcingulate_volume',
'rh_precentral_volume',
'rh_precuneus_volume',
'rh_rostralanteriorcingulate_volume',
'rh_rostralmiddlefrontal_volume',
'rh_superiorfrontal_volume',
'rh_superiorparietal_volume',
'rh_superiortemporal_volume',
'rh_supramarginal_volume',
'rh_frontalpole_volume',
'rh_temporalpole_volume',
'rh_transversetemporal_volume',
'rh_insula_volume']
# + [markdown] colab_type="text" id="bw9sBiGwiBN9"
# Then, we calculate the relative brain region volumes (original volume divided by the total intracranial volume).
# + colab_type="code" id="Ae1dT-5O1Fln" colab={}
# Get the relative brain region volumes
x_dataset = dataset_df[COLUMNS_NAME].values
tiv = dataset_df['EstimatedTotalIntraCranialVol'].values
tiv = tiv[:, np.newaxis]
x_dataset = (np.true_divide(x_dataset, tiv)).astype('float32')
# + [markdown] colab_type="text" id="5F8WZKzc1O-v"
# Next, we iterate over all models performing the calculation of the deviations. In our paper, we define the **deviation score as the mean squared error** between the autoencoder's reconstruction and the inputted data (more details in the Section 2.5 Analysis of the observed deviation).
#
# **Note**: if the age of someone is lower than 47 or higher than 73, the age value will be clipped to be inside the range (47, 73). For example, if someone has age = 40, it will be rounded to 47. We performed this clipping because the age is an important variable for conditioning the predictions of our model.
#
# + colab_type="code" id="eXcs1zxWaw-w" outputId="d47e2f10-b092-4b28-d479-ccef3bccb4cc" colab={"base_uri": "https://localhost:8080/", "height": 34}
warnings.filterwarnings('ignore')
model_dir = Path('models')
N_BOOTSTRAP = 1000
# Create dataframe to store outputs
reconstruction_error_df = pd.DataFrame(columns=['Participant_ID'])
reconstruction_error_df['Participant_ID'] = dataset_df['Participant_ID']
# ----------------------------------------------------------------------------
for i_bootstrap in tqdm(range(N_BOOTSTRAP)):
bootstrap_model_dir = model_dir / '{:03d}'.format(i_bootstrap)
# ----------------------------------------------------------------------------
encoder = keras.models.load_model(bootstrap_model_dir / 'encoder.h5', compile=False)
decoder = keras.models.load_model(bootstrap_model_dir / 'decoder.h5', compile=False)
scaler = joblib.load(bootstrap_model_dir / 'scaler.joblib')
enc_age = joblib.load(bootstrap_model_dir / 'age_encoder.joblib')
enc_gender = joblib.load(bootstrap_model_dir / 'gender_encoder.joblib')
# ----------------------------------------------------------------------------
x_normalized = scaler.transform(x_dataset)
# ----------------------------------------------------------------------------
age = dataset_df['Age'].values
age = np.clip(age, 47, 73)
age = age[:, np.newaxis].astype('float32')
one_hot_age = enc_age.transform(age)
gender = dataset_df['Gender'].values[:, np.newaxis].astype('float32')
one_hot_gender = enc_gender.transform(gender)
y_data = np.concatenate((one_hot_age, one_hot_gender), axis=1).astype('float32')
# ----------------------------------------------------------------------------
encoded = encoder(x_normalized, training=False)
reconstruction = decoder(tf.concat([encoded, y_data], axis=1), training=False)
# ----------------------------------------------------------------------------
reconstruction_error = np.mean((x_normalized - reconstruction) ** 2, axis=1)
reconstruction_error_df[('Reconstruction error {:03d}'.format(i_bootstrap))] = reconstruction_error
# + [markdown] colab_type="text" id="Ah4VJ3R1wqwD"
# Finally, we compute the mean deviation score and save the file with all scores.
# + colab_type="code" id="6AEzjpJ3xvia" outputId="6901d3c6-1890-4df3-d66e-3daaf1b6c4cf" colab={"base_uri": "https://localhost:8080/", "height": 165}
reconstruction_error_df['Mean reconstruction error'] = reconstruction_error_df[reconstruction_error_df.columns[1:]].mean(axis=1)
reconstruction_error_df
# + colab_type="code" id="n5rJsceowHC_" colab={}
reconstruction_error_df.to_csv('reconstruction_error.csv', index=False)
# + [markdown] colab_type="text" id="3xXjXZlo2DB9"
# ## Download predictions
# Finally, you can download the result in the "Files" tab or executing the cell below.
# + colab_type="code" id="T-385ko1v0F4" colab={}
files.download('reconstruction_error.csv')
# + [markdown] id="aafKvn3SjdXk" colab_type="text"
# With the predictions of our 1,000 model, you could use their mean value to have a reliable deviation estimation of the subject. Besides that, the variance (or standard deviation) along the 1,000 predictions could give an ideia of how much our models are certain about the subject's deviation score.
#
| notebooks/predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sqlite3
import pandas as pd
import matplotlib.dates as dates
from datetime import datetime as dt
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import FuncFormatter, PercentFormatter
# -
conn = sqlite3.connect('../data/netflix-data-aggregated.db')
v4 = pd.read_sql_query('select * from success_rate_v4', con=conn, parse_dates=['dtime'])
v6 = pd.read_sql_query('select * from success_rate_v6', con=conn, parse_dates=['dtime'])
conn.close()
v4
v6
for df in [v4, v6]:
df['year'] = df['dtime'].dt.year
df['month'] = df['dtime'].dt.month
# ### Daily medians for each probe
# +
params = {'axes.labelsize' : 14, 'axes.titlesize' : 14,
'font.size' : 14, 'legend.fontsize' : 14,
'xtick.labelsize' : 14, 'ytick.labelsize' : 14}
plt.rcParams.update(params)
ts_fig, (ts_ax_v4, ts_ax_v6) = plt.subplots(figsize = (7, 2.5), nrows=2, gridspec_kw={'height_ratios': [6, 19]})
info = [('IPv4', 'success_rate', ts_ax_v4, v4),
('IPv6', 'success_rate', ts_ax_v6, v6)]
for (version, col, ts_ax, df) in info:
bp = df.boxplot(column = [col], by = ['year', 'month'], ax = ts_ax, sym = "",
medianprops = { 'linewidth' : 2.0 }, return_type = 'dict')
color = 'gray'
if version == 'IPv4':
color = 'blue'
elif version == 'IPv6':
color = 'red'
for key in bp.keys():
for item in bp[key]['medians']:
item.set_color(color)
ts_fig.suptitle('')
ts_ax.set_title('')
ts_ax.set_xlabel('')
ts_ax.set_ylabel('')
if version=='IPv4':
ts_ax.set_ylim([0.925, 1.025])
ts_ax.set_yticks(np.arange(0.95, 1.04, 0.05))
ts_ax.yaxis.set_major_formatter(PercentFormatter(xmax=1))
else:
ts_ax.set_ylim([0.375, 1.025])
ts_ax.set_yticks(np.arange(0.4, 1.05, 0.1))
ts_ax.yaxis.set_major_formatter(PercentFormatter(xmax=1))
major_ticklabels = ts_ax.xaxis.get_majorticklabels()
for ticklabel in major_ticklabels:
if version == 'IPv4':
ticklabel.set_text('')
else:
label = ticklabel.get_text()[1:-1]
try:
y, m = label.split(', ')
except:
pass
if m == "1":
label = 'Jan\n%s' % y
elif m == "10":
label = 'Oct'
elif m == "4":
label = 'Apr'
elif m == "7":
label = 'Jul'
else:
label = ''
ticklabel.set_text(label)
ts_ax.grid(False)
ts_ax.spines['right'].set_color('none')
ts_ax.spines['top'].set_color('none')
ts_ax.yaxis.set_ticks_position('left')
ts_ax.xaxis.set_ticks_position('bottom')
ts_ax.spines['bottom'].set_position(('axes', -0.03))
ts_ax.spines['left'].set_position(('axes', -0.03))
ts_ax.set_xticklabels(major_ticklabels, rotation = 0)
ax1_ = ts_ax.twinx()
ax1_.spines['right'].set_color('none')
ax1_.spines['top'].set_color('none')
ax1_.spines['left'].set_color('none')
ax1_.spines['bottom'].set_color('none')
ax1_.yaxis.set_ticks_position('none')
ax1_.set_ylabel('%s' % version)
plt.setp(ax1_.get_yticklabels(), visible = False)
ts_ax_v4.set_title('Success Rate', y=1.05)
ts_fig.savefig('../plots/success-rate.pdf', bbox_inches = 'tight')
plt.show()
plt.close('all')
| notebooks/fig-3-success-rate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://github.com/PaddlePaddle/PaddleSpeech"><img style="position: absolute; z-index: 999; top: 0; right: 0; border: 0; width: 128px; height: 128px;" src="https://nosir.github.io/cleave.js/images/right-graphite@2x.png" alt="Fork me on GitHub"></a>
#
# # 『听』和『说』
# 人类通过听觉获取的信息大约占所有感知信息的 20% ~ 30%。声音存储了丰富的语义以及时序信息,由专门负责听觉的器官接收信号,产生一系列连锁刺激后,在人类大脑的皮层听区进行处理分析,获取语义和知识。近年来,随着深度学习算法上的进步以及不断丰厚的硬件资源条件,**文本转语音(Text-to-Speech, TTS)** 技术在移动、虚拟娱乐等领域得到了广泛的应用。</font>
# ## "听"书
# 使用 [PaddleOCR](https://github.com/PaddlePaddle/PaddleOCR) 直接获取书籍上的文字。
# download demo sources
# !mkdir download
# !wget -P download https://paddlespeech.bj.bcebos.com/tutorial/tts/ocr_result.jpg
# !wget -P download https://paddlespeech.bj.bcebos.com/tutorial/tts/ocr.wav
# !wget -P download https://paddlespeech.bj.bcebos.com/tutorial/tts/tts_lips.mp4
import IPython.display as dp
from PIL import Image
img_path = 'download/ocr_result.jpg'
im = Image.open(img_path)
dp.display(im)
# 使用 [PaddleSpeech](https://github.com/PaddlePaddle/PaddleSpeech),阅读上一步识别出来的文字。
dp.Audio("download/ocr.wav")
# 具体实现代码详见 [Story Talker](https://github.com/PaddlePaddle/PaddleSpeech/tree/develop/demos/story_talker)
# ## 偶像开口说话
# *元宇宙来袭,构造你的虚拟人!* 看看 [PaddleGAN](https://github.com/PaddlePaddle/PaddleGAN) 怎样合成唇形,让WiFi之母——海蒂·拉玛说话。
from IPython.display import HTML
html_str = '''
<video controls width="600" height="360" src="{}">animation</video>
'''.format("download/tts_lips.mp4")
dp.display(HTML(html_str))
# 具体实现代码请参考 [Metaverse](https://github.com/PaddlePaddle/PaddleSpeech/tree/develop/demos/metaverse)。
#
# 下面让我们来系统地学习语音方面的知识,看看怎样使用 **PaddleSpeech** 实现基本的语音功能,以及怎样结合光学字符识别(Optical Character Recognition,OCR)、自然语言处理(Natural Language Processing,NLP)等技术“听”书、让名人开口说话。
# # 前言
# ## 背景知识
# 为了更好地了解文本转语音任务的要素,我们先简要地回顾一下文本转语音的发展历史。如果你对此已经有所了解,或希望能尽快使用代码实现,请直接跳至[实践](#实践)。
# ### 定义
# <!----
# Note:
# 1.此句抄自 [李沐Dive into Dive Learning](https://zh-v2.d2l.ai/chapter_introduction/index.html)
# 2.修改参考A survey on Neural Speech Sysnthesis.
# --->
# 文本转语音,又称语音合成(Speech Sysnthesis),指的是将一段文本按照一定需求转化成对应的音频,这种特性决定了的输出数据比输入输入长得多。文本转语音是一项包含了语义学、声学、数字信号处理以及机器学习的等多项学科的交叉任务。虽然辨识低质量音频文件的内容对人类来说很容易,但这对计算机来说并非易事。
#
# 按照不同的应用需求,更广义的语音合成研究包括:*语音转换*,例如说话人转换、语音到歌唱转换、语音情感转换、口音转换等;*歌唱合成*,例如歌词到歌唱转换、可视语音合成等。
#
# ### 发展历史
#
# <!--
# 以下摘自维基百科 https://en.wikipedia.org/wiki/Speech_synthesis
# --->
#
# 在第二次工业革命之前,语音的合成主要以机械式的音素合成为主。1779年,德裔丹麦科学家 <NAME> 建造了人类的声道模型,使其可以产生五个长元音。1791年, <NAME> 添加了唇和舌的模型,使其能够发出辅音和元音。贝尔实验室于20世纪30年代发明了声码器(Vocoder),将语音自动分解为音调和共振,此项技术由 <NAME> 改进为键盘式合成器并于 1939年纽约世界博览会展出。
#
# 第一台基于计算机的语音合成系统起源于20世纪50年代。1961年,IBM 的 <NAME>,以及 <NAME> 使用 IBM 704 计算机合成语音,成为贝尔实验室最著名的成就之一。1975年,第一代语音合成系统之一 —— MUSA(MUltichannel Speaking Automation)问世,其由一个独立的硬件和配套的软件组成。1978年发行的第二个版本也可以进行无伴奏演唱。90 年代的主流是采用 MIT 和贝尔实验室的系统,并结合自然语言处理模型。
#
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/55035de353b042cd8c4468819b2d36e2fcc89bffdf2b442fa4c7b0b5499e1592"></center>
#
# ### 主流方法
#
# 当前的主流方法分为**基于统计参数的语音合成**、**波形拼接语音合成**、**混合方法**以及**端到端神经网络语音合成**。基于参数的语音合成包含隐马尔可夫模型(Hidden Markov Model,HMM)以及深度学习网络(Deep Neural Network,DNN)。端到端的方法保函声学模型+声码器以及“完全”端到端方法。
#
#
# ## 基于深度学习的语音合成技术
#
# ### 语音合成基本知识
#
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/10859679d74745ab82fb6f5c9984a95152c25b0e3dce4515b120c8997a6752d8"></center>
# <br></br>
#
# 语音合成流水线包含 <font color="#ff0000">**文本前端(Text Frontend)**</font> 、<font color="#ff0000">**声学模型(Acoustic Model)**</font> 和 <font color="#ff0000">**声码器(Vocoder)**</font> 三个主要模块:
# - 通过文本前端模块将原始文本转换为字符/音素。
# - 通过声学模型将字符/音素转换为声学特征,如线性频谱图、mel 频谱图、LPC 特征等。
# - 通过声码器将声学特征转换为波形。
# # 实践
# ### 安装 paddlespeech
# !pip install --upgrade pip && pip install paddlespeech -U
# 环境安装请参考 [Installation](https://github.com/PaddlePaddle/PaddleSpeech/blob/develop/docs/source/install.md) 教程。
# 下面使用 **PaddleSpeech** 提供的预训练模型合成中文语音。
# ## 数据及模型准备
# ### 获取PaddlePaddle预训练模型
# !wget -P download https://paddlespeech.bj.bcebos.com/Parakeet/released_models/pwgan/pwg_baker_ckpt_0.4.zip
# !unzip -d download download/pwg_baker_ckpt_0.4.zip
# !wget -P download https://paddlespeech.bj.bcebos.com/Parakeet/released_models/fastspeech2/fastspeech2_nosil_baker_ckpt_0.4.zip
# !unzip -d download download/fastspeech2_nosil_baker_ckpt_0.4.zip
# !tree download/pwg_baker_ckpt_0.4
# !tree download/fastspeech2_nosil_baker_ckpt_0.4
# ### 导入 Python 包
# 本项目的依赖需要用到 nltk 包,但是有时会因为网络原因导致不好下载,此处手动下载一下放到百度服务器的包
# !wget https://paddlespeech.bj.bcebos.com/Parakeet/tools/nltk_data.tar.gz
# !tar zxvf nltk_data.tar.gz
# +
# 设置 gpu 环境
# %env CUDA_VISIBLE_DEVICES=0
import logging
import sys
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
import argparse
import os
from pathlib import Path
import IPython.display as dp
import matplotlib.pyplot as plt
import numpy as np
import paddle
import soundfile as sf
import yaml
from paddlespeech.t2s.frontend.zh_frontend import Frontend
from paddlespeech.t2s.models.fastspeech2 import FastSpeech2
from paddlespeech.t2s.models.fastspeech2 import FastSpeech2Inference
from paddlespeech.t2s.models.parallel_wavegan import PWGGenerator
from paddlespeech.t2s.models.parallel_wavegan import PWGInference
from paddlespeech.t2s.modules.normalizer import ZScore
from yacs.config import CfgNode
# ### 设置预训练模型的路径
fastspeech2_config = "download/fastspeech2_nosil_baker_ckpt_0.4/default.yaml"
fastspeech2_checkpoint = "download/fastspeech2_nosil_baker_ckpt_0.4/snapshot_iter_76000.pdz"
fastspeech2_stat = "download/fastspeech2_nosil_baker_ckpt_0.4/speech_stats.npy"
pwg_config = "download/pwg_baker_ckpt_0.4/pwg_default.yaml"
pwg_checkpoint = "download/pwg_baker_ckpt_0.4/pwg_snapshot_iter_400000.pdz"
pwg_stat = "download/pwg_baker_ckpt_0.4/pwg_stats.npy"
phones_dict = "download/fastspeech2_nosil_baker_ckpt_0.4/phone_id_map.txt"
# 读取 conf 配置文件并结构化
with open(fastspeech2_config) as f:
fastspeech2_config = CfgNode(yaml.safe_load(f))
with open(pwg_config) as f:
pwg_config = CfgNode(yaml.safe_load(f))
print("========Config========")
print(fastspeech2_config)
print("---------------------")
print(pwg_config)
# ## 文本前端(Text Frontend)
#
# 一个文本前端模块主要包含:
# - 分段(Text Segmentation)
# - 文本正则化(Text Normalization, TN)
# - 分词(Word Segmentation, 主要是在中文中)
# - 词性标注(Part-of-Speech, PoS)
# - 韵律预测(Prosody)
# - 字音转换(Grapheme-to-Phoneme,G2P)
# <font size=2>(Grapheme: **语言**书写系统的最小有意义单位; Phoneme: 区分单词的最小**语音**单位)</font>
# - 多音字(Polyphone)
# - 变调(Tone Sandhi)
# - “一”、“不”变
# - 三声变调
# - 轻声变调
# - 儿化音
# - 方言
# - ...
#
# (输入给声学模型之前,还需要把音素序列转换为 id)
#
#
# 其中最重要的模块是<font color="#ff0000"> 文本正则化 </font>模块和<font color="#ff0000"> 字音转换(TTS 中更常用 G2P 代指) </font>模块。
#
#
# 各模块输出示例:
# ```text
# • Text: 全国一共有112所211高校
# • Text Normalization: 全国一共有一百一十二所二一一高校
# • Word Segmentation: 全国/一共/有/一百一十二/所/二一一/高校/
# • G2P(注意此句中“一”的读音):
# quan2 guo2 yi2 gong4 you3 yi4 bai3 yi1 shi2 er4 suo3 er4 yao1 yao1 gao1 xiao4
# (可以进一步把声母和韵母分开)
# q uan2 g uo2 y i2 g ong4 y ou3 y i4 b ai3 y i1 sh i2 er4 s uo3 er4 y ao1 y ao1 g ao1 x iao4
# (把音调和声韵母分开)
# q uan g uo y i g ong y ou y i b ai y i sh i er s uo er y ao y ao g ao x iao
# 0 2 0 2 0 2 0 4 0 3 ...
# • Prosody (prosodic words #1, prosodic phrases #2, intonation phrases #3, sentence #4):
# 全国#2一共有#2一百#1一十二所#2二一一#1高校#4
# (分词的结果一般是固定的,但是不同人习惯不同,可能有不同的韵律)
# ```
#
# 文本前端模块的设计需要结合很多专业的语义学知识和经验。人类在读文本的时候可以自然而然地读出正确的发音,但是这些先验知识计算机并不知晓。
# 例如,对于一个句子的分词:
#
# ```text
# 我也想过过过儿过过的生活
# 我也想/过过/过儿/过过的/生活
#
# 货拉拉拉不拉拉布拉多
# 货拉拉/拉不拉/拉布拉多
#
# 南京市长江大桥
# 南京市长/江大桥
# 南京市/长江大桥
# ```
# 或者是词的变调和儿化音:
# ```
# 你要不要和我们一起出去玩?
# 你要不(2声)要和我们一(4声)起出去玩(儿)?
#
# 不好,我要一个人出去。
# 不(4声)好,我要一(2声)个人出去。
#
# (以下每个词的所有字都是三声的,请你读一读,体会一下在读的时候,是否每个字都被读成了三声?)
# 纸老虎、虎骨酒、展览馆、岂有此理、手表厂有五种好产品
# ```
# 又或是多音字,这类情况通常需要先正确分词:
# ```text
# 人要行,干一行行一行,一行行行行行;
# 人要是不行,干一行不行一行,一行不行行行不行。
#
# 佟大为妻子产下一女
#
# 海水朝朝朝朝朝朝朝落
# 浮云长长长长长长长消
# ```
#
# PaddleSpeech Text-to-Speech的文本前端解决方案:
# - [文本正则](https://github.com/PaddlePaddle/PaddleSpeech/tree/develop/examples/other/tn)
# - [G2P](https://github.com/PaddlePaddle/PaddleSpeech/tree/develop/examples/other/g2p)
# - 多音字模块: pypinyin/g2pM
# - 变调模块: 用分词 + 规则
# ### 构造文本前端对象
# 传入`phones_dict`,把相应的`phones`转换成`phone_ids`。
# 传入 phones_dict 会把相应的 phones 转换成 phone_ids
frontend = Frontend(phone_vocab_path=phones_dict)
print("Frontend done!")
# ### 调用文本前端
#
# 文本前端对输入数据进行正则化时会进行分句,若`merge_sentences`设置为`False`,则所有分句的 `phone_ids` 构成一个 `List`;若设置为`True`,`input_ids["phone_ids"][0]`则表示整句的`phone_ids`。
# input = "我每天中午12:00起床"
# input = "我出生于2005/11/08,那天的最低气温达到-10°C"
input = "你好,欢迎使用百度飞桨框架进行深度学习研究!"
input_ids = frontend.get_input_ids(input, merge_sentences=True, print_info=True)
phone_ids = input_ids["phone_ids"][0]
print("phone_ids:%s"%phone_ids)
# ## 用深度学习实现文本前端
#
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/85a5cd8aef1e444cbb980a2f1f184316247bbb7870a34925a77b799802df8ef0"></center>
#
# ## 声学模型(Acoustic Model)
#
# 声学模型将字符/音素转换为声学特征,如线性频谱图、mel 频谱图、LPC 特征等,声学特征以 “帧” 为单位,一般一帧是 10ms 左右,一个音素一般对应 5~20 帧左右, 声学模型需要解决的是 <font color="#ff0000">“不等长序列间的映射问题”</font>,“不等长”是指,同一个人发不同音素的持续时间不同,同一个人在不同时刻说同一句话的语速可能不同,对应各个音素的持续时间不同,不同人说话的特色不同,对应各个音素的持续时间不同。这是一个困难的“一对多”问题。
# ```
# # 卡尔普陪外孙玩滑梯
# 000001|baker_corpus|sil 20 k 12 a2 4 er2 10 p 12 u3 12 p 9 ei2 9 uai4 15 s 11 uen1 12 uan2 14 h 10 ua2 11 t 15 i1 16 sil 20
# ```
#
# 声学模型主要分为自回归模型和非自回归模型,其中自回归模型在 `t` 时刻的预测需要依赖 `t-1` 时刻的输出作为输入,预测时间长,但是音质相对较好,非自回归模型不存在预测上的依赖关系,预测时间快,音质相对较差。
#
# 主流声学模型发展的脉络:
# - 自回归模型:
# - Tacotron
# - Tacotron2
# - Transformer TTS
# - 非自回归模型:
# - FastSpeech
# - SpeedySpeech
# - FastPitch
# - FastSpeech2
# - ...
#
# 在本教程中,我们使用 `FastSpeech2` 作为声学模型。
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/6b6d671713ec4d20a0e60653c7a5d4ae3c35b1d1e58b4cc39e0bc82ad4a341d9"></center>
# <br><center> FastSpeech2 网络结构图</center></br>
#
#
# PaddleSpeech TTS 实现的 FastSpeech2 与论文不同的地方在于,我们使用的的是 phone 级别的 `pitch` 和 `energy`(与 FastPitch 类似),这样的合成结果可以更加**稳定**。
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/862c21456c784c41a83a308b7d9707f0810cc3b3c6f94ed48c60f5d32d0072f0"></center>
# <br><center> FastPitch 网络结构图</center></br>
#
# 更多关于[语音合成模型的发展及改进](https://github.com/PaddlePaddle/PaddleSpeech/blob/develop/docs/source/tts/models_introduction.md)。
# ### 初始化声学模型 FastSpeech2
with open(phones_dict, "r") as f:
phn_id = [line.strip().split() for line in f.readlines()]
vocab_size = len(phn_id)
print("vocab_size:", vocab_size)
odim = fastspeech2_config.n_mels
model = FastSpeech2(
idim=vocab_size, odim=odim, **fastspeech2_config["model"])
# 加载预训练模型参数
model.set_state_dict(paddle.load(fastspeech2_checkpoint)["main_params"])
# 推理阶段不启用 batch norm 和 dropout
model.eval()
stat = np.load(fastspeech2_stat)
# 读取数据预处理阶段数据集的均值和标准差
mu, std = stat
mu, std = paddle.to_tensor(mu), paddle.to_tensor(std)
# 构造归一化的新模型
fastspeech2_normalizer = ZScore(mu, std)
fastspeech2_inference = FastSpeech2Inference(fastspeech2_normalizer, model)
fastspeech2_inference.eval()
print("FastSpeech2 done!")
# ### 调用声学模型
with paddle.no_grad():
mel = fastspeech2_inference(phone_ids)
print("shepe of mel (n_frames x n_mels):")
print(mel.shape)
# 绘制声学模型输出的 mel 频谱
fig, ax = plt.subplots(figsize=(16, 6))
im = ax.imshow(mel.T, aspect='auto',origin='lower')
plt.title('Mel Spectrogram')
plt.xlabel('Time')
plt.ylabel('Frequency')
plt.tight_layout()
# ## 声码器(Vocoder)
# 声码器将声学特征转换为波形。声码器需要解决的是 <font color="#ff0000">“信息缺失的补全问题”</font>。信息缺失是指,在音频波形转换为频谱图的时候,存在**相位信息**的缺失,在频谱图转换为 mel 频谱图的时候,存在**频域压缩**导致的信息缺失;假设音频的采样率是16kHZ, 一帧的音频有 10ms,也就是说,1s 的音频有 16000 个采样点,而 1s 中包含 100 帧,每一帧有 160 个采样点,声码器的作用就是将一个频谱帧变成音频波形的 160 个采样点,所以声码器中一般会包含**上采样**模块。
#
# 与声学模型类似,声码器也分为自回归模型和非自回归模型, 更细致的分类如下:
#
# - Autoregression
# - WaveNet
# - WaveRNN
# - LPCNet
# - Flow
# - <font color="#ff0000">WaveFlow</font>
# - WaveGlow
# - FloWaveNet
# - Parallel WaveNet
# - GAN
# - WaveGAN
# - <font color="#ff0000">Parallel WaveGAN</font>
# - <font color="#ff0000">MelGAN</font>
# - <font color="#ff0000">Style MelGAN</font>
# - <font color="#ff0000">Multi Band MelGAN</font>
# - <font color="#ff0000">HiFi GAN</font>
# - VAE
# - Wave-VAE
# - Diffusion
# - WaveGrad
# - DiffWave
#
# PaddleSpeech TTS 主要实现了百度的 `WaveFlow` 和一些主流的 GAN Vocoder, 在本教程中,我们使用 `Parallel WaveGAN` 作为声码器。
#
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/9eafa4e5642d45309e6e8883bff46380407b3858d0934bf5896868281316ce94" width="700"></center>
# <br><center>图1:Parallel WaveGAN 网络结构图</center></br>
#
# 各 GAN Vocoder 的生成器和判别器的 Loss 的区别如下表格所示:
#
# Model | Generator Loss |Discriminator Loss
# :-------------:| :------------:| :-----
# Mel GAN| adversial loss <br> Feature Matching | Multi-Scale Discriminator |
# Parallel Wave GAN|adversial loss <br> Multi-resolution STFT loss | adversial loss|
# Multi-Band Mel GAN | adversial loss <br> full band Multi-resolution STFT loss <br> sub band Multi-resolution STFT loss |Multi-Scale Discriminator|
# HiFi GAN |adversial loss <br> Feature Matching <br> Mel-Spectrogram Loss | Multi-Scale Discriminator <br> Multi-Period Discriminator|
#
# ### 初始化声码器 Parallel WaveGAN
vocoder = PWGGenerator(**pwg_config["generator_params"])
# 模型加载预训练参数
vocoder.set_state_dict(paddle.load(pwg_checkpoint)["generator_params"])
vocoder.remove_weight_norm()
# 推理阶段不启用 batch norm 和 dropout
vocoder.eval()
# 读取数据预处理阶段数据集的均值和标准差
stat = np.load(pwg_stat)
mu, std = stat
mu, std = paddle.to_tensor(mu), paddle.to_tensor(std)
pwg_normalizer = ZScore(mu, std)
# 构建归一化的模型
pwg_inference = PWGInference(pwg_normalizer, vocoder)
pwg_inference.eval()
print("Parallel WaveGAN done!")
# ### 调用声码器
with paddle.no_grad():
wav = pwg_inference(mel)
print("shepe of wav (time x n_channels):%s"%wav.shape)
# 绘制声码器输出的波形图
wave_data = wav.numpy().T
time = np.arange(0, wave_data.shape[1]) * (1.0 / fastspeech2_config.fs)
fig, ax = plt.subplots(figsize=(16, 6))
plt.plot(time, wave_data[0])
plt.title('Waveform')
plt.xlabel('Time (seconds)')
plt.ylabel('Amplitude (normed)')
plt.tight_layout()
# ### 播放音频
dp.Audio(wav.numpy().T, rate=fastspeech2_config.fs)
# ### 保存音频
# !mkdir output
sf.write(
"output/output.wav",
wav.numpy(),
samplerate=fastspeech2_config.fs)
# ## 进阶 —— 个性化调节
# FastSpeech2 模型可以个性化地调节音素时长、音调和能量,通过一些简单的调节就可以获得一些有意思的效果。
#
# 例如对于以下的原始音频`"凯莫瑞安联合体的经济崩溃,迫在眉睫"`。
# 原始音频
dp.display(dp.Audio(url="https://paddlespeech.bj.bcebos.com/Parakeet/docs/demos/speed/x1_001.wav"))
# speed x 1.2
dp.display(dp.Audio(url="https://paddlespeech.bj.bcebos.com/Parakeet/docs/demos/speed/x1.2_001.wav"))
# speed x 0.8
dp.display(dp.Audio(url="https://paddlespeech.bj.bcebos.com/Parakeet/docs/demos/speed/x0.8_001.wav"))
# pitch x 1.3(童声)
dp.display(dp.Audio(url="https://paddlespeech.bj.bcebos.com/Parakeet/docs/demos/child_voice/001.wav"))
# robot
dp.display(dp.Audio(url="https://paddlespeech.bj.bcebos.com/Parakeet/docs/demos/robot/001.wav"))
# 具体实现代码请参考 [Style FastSpeech2](https://github.com/PaddlePaddle/PaddleSpeech/tree/develop/demos/style_fs2)。
# ## 用 PaddleSpeech 训练 TTS 模型
# PaddleSpeech 的 examples 是按照 数据集/模型 的结构安排的:
# ```text
# examples
# ├── aishell3
# │ ├── README.md
# │ ├── tts3
# │ └── vc0
# ├── csmsc
# │ ├── README.md
# │ ├── tts2
# │ ├── tts3
# │ ├── voc1
# │ └── voc3
# ├── ...
# └── ...
# ```
# 我们在每个数据集的 README.md 介绍了子目录和模型的对应关系, 在 TTS 中有如下对应关系:
# ```text
# tts0 - Tactron2
# tts1 - TransformerTTS
# tts2 - SpeedySpeech
# tts3 - FastSpeech2
# voc0 - WaveFlow
# voc1 - Parallel WaveGAN
# voc2 - MelGAN
# voc3 - MultiBand MelGAN
# ```
# ### 基于 CSMCS 数据集训练 FastSpeech2 模型
# ```bash
# git clone https://github.com/PaddlePaddle/PaddleSpeech.git
# # cd examples/csmsc/tts3
# ```
# 根据 README.md, 下载 CSMCS 数据集和其对应的强制对齐文件, 并放置在对应的位置
# ```bash
# ./run.sh
# ```
# `run.sh` 中包含预处理、训练、合成、静态图推理等步骤:
#
# ```bash
# # #!/bin/bash
# set -e
# source path.sh
# gpus=0,1
# stage=0
# stop_stage=100
# conf_path=conf/default.yaml
# train_output_path=exp/default
# ckpt_name=snapshot_iter_153.pdz
#
# # with the following command, you can choice the stage range you want to run
# # such as `./run.sh --stage 0 --stop-stage 0`
# # this can not be mixed use with `$1`, `$2` ...
# source ${MAIN_ROOT}/utils/parse_options.sh || exit 1
#
# if [ ${stage} -le 0 ] && [ ${stop_stage} -ge 0 ]; then
# # prepare data
# bash ./local/preprocess.sh ${conf_path} || exit -1
# fi
# if [ ${stage} -le 1 ] && [ ${stop_stage} -ge 1 ]; then
# # train model, all `ckpt` under `train_output_path/checkpoints/` dir
# CUDA_VISIBLE_DEVICES=${gpus} ./local/train.sh ${conf_path} ${train_output_path} || exit -1
# fi
# if [ ${stage} -le 2 ] && [ ${stop_stage} -ge 2 ]; then
# # synthesize, vocoder is pwgan
# CUDA_VISIBLE_DEVICES=${gpus} ./local/synthesize.sh ${conf_path} ${train_output_path} ${ckpt_name} || exit -1
# fi
# if [ ${stage} -le 3 ] && [ ${stop_stage} -ge 3 ]; then
# # synthesize_e2e, vocoder is pwgan
# CUDA_VISIBLE_DEVICES=${gpus} ./local/synthesize_e2e.sh ${conf_path} ${train_output_path} ${ckpt_name} || exit -1
# fi
# if [ ${stage} -le 4 ] && [ ${stop_stage} -ge 4 ]; then
# # inference with static model
# CUDA_VISIBLE_DEVICES=${gpus} ./local/inference.sh ${train_output_path} || exit -1
# fi
# ```
#
# ### 基于 CSMCS 数据集训练 Parallel WaveGAN 模型
# ```bash
# git clone https://github.com/PaddlePaddle/PaddleSpeech.git
# # cd examples/csmsc/voc1
# ```
# 根据 README.md, 下载 CSMCS 数据集和其对应的强制对齐文件, 并放置在对应的位置
# ```bash
# ./run.sh
# ```
# `run.sh` 中包含预处理、训练、合成等步骤:
# ```bash
# # #!/bin/bash
# set -e
# source path.sh
# gpus=0,1
# stage=0
# stop_stage=100
# conf_path=conf/default.yaml
# train_output_path=exp/default
# ckpt_name=snapshot_iter_5000.pdz
#
# # with the following command, you can choice the stage range you want to run
# # such as `./run.sh --stage 0 --stop-stage 0`
# # this can not be mixed use with `$1`, `$2` ...
# source ${MAIN_ROOT}/utils/parse_options.sh || exit 1
#
# if [ ${stage} -le 0 ] && [ ${stop_stage} -ge 0 ]; then
# # prepare data
# ./local/preprocess.sh ${conf_path} || exit -1
# fi
# if [ ${stage} -le 1 ] && [ ${stop_stage} -ge 1 ]; then
# # train model, all `ckpt` under `train_output_path/checkpoints/` dir
# CUDA_VISIBLE_DEVICES=${gpus} ./local/train.sh ${conf_path} ${train_output_path} || exit -1
# fi
# if [ ${stage} -le 2 ] && [ ${stop_stage} -ge 2 ]; then
# # synthesize
# CUDA_VISIBLE_DEVICES=${gpus} ./local/synthesize.sh ${conf_path} ${train_output_path} ${ckpt_name} || exit -1
# fi
# ```
# # FAQ
#
# - 需要注意的问题
# - 经验与分享
# - 用户的其他问题
# # 作业
# 在 CSMSC 数据集上利用 FastSpeech2 和 Parallel WaveGAN 实现一个中文 TTS 系统。
# # 关注 PaddleSpeech
# 请关注我们的 [Github Repo](https://github.com/PaddlePaddle/PaddleSpeech/),非常欢迎加入以下微信群参与讨论:
# - 扫描二维码
# - 添加运营小姐姐微信
# - 通过后回复【语音】
# - 系统自动邀请加入技术群
#
# <center><img src="https://ai-studio-static-online.cdn.bcebos.com/bca0bc75dce14b53af44e374e64fc91aeeb13c075c894d6aabed033148f65377" ></center>
#
| docs/tutorial/tts/tts_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp core
# -
# # core
#
# > Functions to split overlapping boundig boxes.
#hide
from nbdev.showdoc import *
from nbdev.export import notebook2script
#export
import numpy as np
import pandas as pd
import geopandas as gpd
import shapely
from shapely.geometry import Polygon, LineString
from tqdm import tqdm
import matplotlib.pyplot as plt
#export
def box2polygon(x):
return Polygon([(x[0], x[1]), (x[0]+x[2], x[1]), (x[0]+x[2], x[1]+x[3]), (x[0], x[1]+x[3])])
df = pd.read_csv('data/train.csv')
boxes = df.groupby('image_id').agg({'bbox' : lambda x : list(x)})
box = boxes.iloc[2]
bbox = np.array([eval(l) for l in box.bbox]).astype(int).tolist()
gdf = gpd.GeoDataFrame({'geometry': [box2polygon(b) for b in bbox]})
gdf.head()
#export
def slice_box(box_A:Polygon, box_B:Polygon, margin=10, line_mult=10):
"Returns box_A sliced according to the distance to box_B."
vec_AB = np.array([box_B.centroid.x - box_A.centroid.x, box_B.centroid.y - box_A.centroid.y])
vec_ABp = np.array([-(box_B.centroid.y - box_A.centroid.y), box_B.centroid.x - box_A.centroid.x])
vec_AB_norm = np.linalg.norm(vec_AB)
split_point = box_A.centroid + vec_AB/2 - (vec_AB/vec_AB_norm)*margin
line = LineString([split_point-line_mult*vec_ABp, split_point+line_mult*vec_ABp])
split_box = shapely.ops.split(box_A, line)
if len(split_box) == 1: return split_box, None, line
is_center = [s.contains(box_A.centroid) for s in split_box]
where_is_center = np.argwhere(is_center).reshape(-1)[0]
where_not_center = np.argwhere(~np.array(is_center)).reshape(-1)[0]
split_box_center = split_box[where_is_center]
split_box_out = split_box[where_not_center]
return split_box_center, split_box_out, line
# +
inter = gdf.loc[gdf.intersects(gdf.iloc[20].geometry)]
box_A = inter.iloc[0].values[0]
box_B = inter.iloc[1].values[0]
polyA, _, lineA = slice_box(box_A, box_B, margin=10, line_mult=1.2)
polyB, _, lineB = slice_box(box_B, box_A, margin=10, line_mult=1.2)
boxes = gpd.GeoDataFrame({'geometry': [box_A, box_B]})
centroids = gpd.GeoDataFrame({'geometry': [box_A.centroid, box_B.centroid]})
splited_boxes = gpd.GeoDataFrame({'geometry': [polyA, polyB]})
lines = gpd.GeoDataFrame({'geometry': [lineA, lineB]})
fig, ax = plt.subplots(dpi=120)
boxes.plot(ax=ax, facecolor='gray', edgecolor='k', alpha=0.5)
centroids.plot(ax=ax, c='k')
ax.axis('off');
fig, ax = plt.subplots(dpi=120)
boxes.plot(ax=ax, facecolor='gray', edgecolor='k', alpha=0.1)
splited_boxes.plot(ax=ax, facecolor='olive', edgecolor='k')
centroids.plot(ax=ax, c='k')
lines.plot(ax=ax, color='k')
ax.axis('off');
# +
#export
def intersection_list(polylist):
r = polylist[0]
for p in polylist:
r = r.intersection(p)
return r
def slice_one(gdf, index):
inter = gdf.loc[gdf.intersects(gdf.iloc[index].geometry)]
if len(inter) == 1: return inter.geometry.values[0]
box_A = inter.loc[index].values[0]
inter = inter.drop(index, axis=0)
polys = []
for i in range(len(inter)):
box_B = inter.iloc[i].values[0]
polyA, *_ = slice_box(box_A, box_B)
polys.append(polyA)
return intersection_list(polys)
def slice_all(gdf):
polys = []
for i in tqdm(range(len(gdf))):
polys.append(slice_one(gdf, i))
return gpd.GeoDataFrame({'geometry': polys})
# +
res_df = slice_all(gdf)
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(10,5), dpi=120)
gdf.plot(ax=ax1, alpha=0.5, color='gray')
res_df.plot(ax=ax2, alpha=0.5, color='olive')
ax1.axis('equal')
ax2.axis('equal')
ax1.set_title('Original bounding boxes')
ax2.set_title('Splited bounding boxes')
fig.tight_layout()
# -
#hide
notebook2script()
| 00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # my first notebook
print ('my first notebook')
1+2
a = 3
print (a)
pip install xlrd
# !pip install xlrd
import xlrd
book = xlrd.open_workbook("Diamonds.xls")
sheet = book.sheet_by_name("Diamonds")
for row_index in range(1,5): # read the first 4 rows, skip the first row
id_, weight, color,_,_,price = sheet.row_values(row_index)
print(id_,weight,color,price)
| Lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.List and its default functions
#
# The list() function creates a list object.
# A list object is a collection which is ordered and changeable.
#
# + active=""
# list append()-add a single elements to the end of the list.
# + active=""
# list clear()-removes all itens from the list.
# + active=""
# list extend()-add iterable elements to the end of the list.
# + active=""
# list pop()-removes element at the given index.
# + active=""
# list reverse()-reverse the list.
# -
# # 2.Dictionary and its default functions
# Dictionary in python is an unordered collection of data values,used to store data values like a map,which unlike other data types that hold ony single value as an elements,Dictionary hold key.
# + active=""
# setdefault()-returns the value of key.
# -
# # 3. Sets and its default functions
#
# A set is an unordered collection of items.Set items are unique and immutable.
#
# copy()-Returns a copy of the set
# + active=""
# difference()-Returns the difference of two or more sets as a new set
# -
# # 4.Tuple and explore default method.
# Tuple are immuatables.i.e.we cannot change items once it is assigned.
# + active=""
# tuple count()-returns count of the elements in the tuple.
# + active=""
# tuple index()- returns the index of the element in the tuple.
# -
# # 5.Strings and explore default method.
# A string is a sequence of characters enclosed in quotation marks.
# + active=""
# string capitalize()-Convert first character to capital letter
#
# + active=""
# string find()-returns the index of first occurrence of substring.
# + active=""
# string index()-returns index of substring
#
# + active=""
# string join()-returns a concatenated string.
# + active=""
# string replace()-replaces substring inside.
| assignment1 day-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pororo
pororo.__version__
from pororo import Pororo
mrc = Pororo(task="mrc", lang="ko")
mrc(
"카카오브레인이 공개한 것은?",
"""카카오 인공지능(AI) 연구개발 자회사 카카오브레인이 AI 솔루션을 첫 상품화했다. 카카오는 카카오브레인 '포즈(pose·자세분석) API'를 유료 공개한다고 24일 밝혔다. 카카오브레인이 AI 기술을 유료 API를 공개하는 것은 처음이다. 공개하자마자 외부 문의가 쇄도한다. 포즈는 AI 비전(VISION, 영상·화면분석) 분야 중 하나다. 카카오브레인 포즈 API는 이미지나 영상을 분석해 사람 자세를 추출하는 기능을 제공한다."""
)
| examples/reading_comprehension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pytorch_tabnet.tab_model import TabNetClassifier
import torch
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
np.random.seed(0)
import os
import wget
from pathlib import Path
import shutil
import gzip
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# # Download ForestCoverType dataset
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/covtype/covtype.data.gz"
dataset_name = 'forest-cover-type'
tmp_out = Path('./data/'+dataset_name+'.gz')
out = Path(os.getcwd()+'/data/'+dataset_name+'.csv')
# +
out.parent.mkdir(parents=True, exist_ok=True)
if out.exists():
print("File already exists.")
else:
print("Downloading file...")
wget.download(url, tmp_out.as_posix())
with gzip.open(tmp_out, 'rb') as f_in:
with open(out, 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
# -
# # Load data and split
# Same split as in original paper
# +
target = "Covertype"
bool_columns = [
"Wilderness_Area1", "Wilderness_Area2", "Wilderness_Area3",
"Wilderness_Area4", "Soil_Type1", "Soil_Type2", "Soil_Type3", "Soil_Type4",
"Soil_Type5", "Soil_Type6", "Soil_Type7", "Soil_Type8", "Soil_Type9",
"Soil_Type10", "Soil_Type11", "Soil_Type12", "Soil_Type13", "Soil_Type14",
"Soil_Type15", "Soil_Type16", "Soil_Type17", "Soil_Type18", "Soil_Type19",
"Soil_Type20", "Soil_Type21", "Soil_Type22", "Soil_Type23", "Soil_Type24",
"Soil_Type25", "Soil_Type26", "Soil_Type27", "Soil_Type28", "Soil_Type29",
"Soil_Type30", "Soil_Type31", "Soil_Type32", "Soil_Type33", "Soil_Type34",
"Soil_Type35", "Soil_Type36", "Soil_Type37", "Soil_Type38", "Soil_Type39",
"Soil_Type40"
]
int_columns = [
"Elevation", "Aspect", "Slope", "Horizontal_Distance_To_Hydrology",
"Vertical_Distance_To_Hydrology", "Horizontal_Distance_To_Roadways",
"Hillshade_9am", "Hillshade_Noon", "Hillshade_3pm",
"Horizontal_Distance_To_Fire_Points"
]
feature_columns = (
int_columns + bool_columns + [target])
# +
train = pd.read_csv(out, header=None, names=feature_columns)
n_total = len(train)
# Train, val and test split follows
# <NAME>, <NAME>, <NAME>, and <NAME>.
# Xgboost: Scalable GPU accelerated learning. arXiv:1806.11248, 2018.
train_val_indices, test_indices = train_test_split(
range(n_total), test_size=0.2, random_state=0)
train_indices, valid_indices = train_test_split(
train_val_indices, test_size=0.2 / 0.6, random_state=0)
# -
# # Simple preprocessing
#
# Label encode categorical features and fill empty cells.
# +
categorical_columns = []
categorical_dims = {}
for col in train.columns[train.dtypes == object]:
print(col, train[col].nunique())
l_enc = LabelEncoder()
train[col] = train[col].fillna("VV_likely")
train[col] = l_enc.fit_transform(train[col].values)
categorical_columns.append(col)
categorical_dims[col] = len(l_enc.classes_)
for col in train.columns[train.dtypes == 'float64']:
train.fillna(train.loc[train_indices, col].mean(), inplace=True)
# -
# # Define categorical features for categorical embeddings
# +
unused_feat = []
features = [ col for col in train.columns if col not in unused_feat+[target]]
cat_idxs = [ i for i, f in enumerate(features) if f in categorical_columns]
cat_dims = [ categorical_dims[f] for i, f in enumerate(features) if f in categorical_columns]
# -
# # Network parameters
clf = TabNetClassifier(
n_d=64, n_a=64, n_steps=5,
gamma=1.5, n_independent=2, n_shared=2,
cat_idxs=cat_idxs,
cat_dims=cat_dims,
cat_emb_dim=1,
lambda_sparse=1e-4, momentum=0.3, clip_value=2.,
optimizer_fn=torch.optim.Adam,
optimizer_params=dict(lr=2e-2),
scheduler_params = {"gamma": 0.95,
"step_size": 20},
scheduler_fn=torch.optim.lr_scheduler.StepLR, epsilon=1e-15
)
# # Training
# +
X_train = train[features].values[train_indices]
y_train = train[target].values[train_indices]
X_valid = train[features].values[valid_indices]
y_valid = train[target].values[valid_indices]
X_test = train[features].values[test_indices]
y_test = train[target].values[test_indices]
# -
max_epochs = 1000 if not os.getenv("CI", False) else 2
clf.fit(
X_train=X_train, y_train=y_train,
X_valid=X_valid, y_valid=y_valid,
max_epochs=max_epochs, patience=100,
batch_size=16384, virtual_batch_size=256
)
# plot losses
plt.plot(clf.history['train']['loss'])
plt.plot(clf.history['valid']['loss'])
# plot accuracies
plt.plot([-x for x in clf.history['train']['metric']])
plt.plot([-x for x in clf.history['valid']['metric']])
# ### Predictions
#
# +
# To get final results you may need to use a mapping for classes
# as you are allowed to use targets like ["yes", "no", "maybe", "I don't know"]
preds_mapper = { idx : class_name for idx, class_name in enumerate(clf.classes_)}
preds = clf.predict_proba(X_test)
y_pred = np.vectorize(preds_mapper.get)(np.argmax(preds, axis=1))
test_acc = accuracy_score(y_pred=y_pred, y_true=y_test)
print(f"BEST VALID SCORE FOR {dataset_name} : {clf.best_cost}")
print(f"FINAL TEST SCORE FOR {dataset_name} : {test_acc}")
# +
# or you can simply use the predict method
y_pred = clf.predict(X_test)
test_acc = accuracy_score(y_pred=y_pred, y_true=y_test)
print(f"FINAL TEST SCORE FOR {dataset_name} : {test_acc}")
# -
# # Global explainability : feat importance summing to 1
clf.feature_importances_
# # Local explainability and masks
explain_matrix, masks = clf.explain(X_test)
# +
fig, axs = plt.subplots(1, 5, figsize=(20,20))
for i in range(5):
axs[i].imshow(masks[i][:50])
axs[i].set_title(f"mask {i}")
# -
# # XGB
n_estimators = 1000 if not os.getenv("CI", False) else 20
# +
from xgboost import XGBClassifier
clf_xgb = XGBClassifier(max_depth=8,
learning_rate=0.1,
n_estimators=n_estimators,
verbosity=0,
silent=None,
objective="multi:softmax",
booster='gbtree',
n_jobs=-1,
nthread=None,
gamma=0,
min_child_weight=1,
max_delta_step=0,
subsample=0.7,
colsample_bytree=1,
colsample_bylevel=1,
colsample_bynode=1,
reg_alpha=0,
reg_lambda=1,
scale_pos_weight=1,
base_score=0.5,
random_state=0,
seed=None,)
clf_xgb.fit(X_train, y_train,
eval_set=[(X_valid, y_valid)],
early_stopping_rounds=40,
verbose=10)
# +
preds_valid = np.array(clf_xgb.predict_proba(X_valid, ))
valid_acc = accuracy_score(y_pred=np.argmax(preds_valid, axis=1) + 1, y_true=y_valid)
print(valid_acc)
preds_test = np.array(clf_xgb.predict_proba(X_test))
test_acc = accuracy_score(y_pred=np.argmax(preds_test, axis=1) + 1, y_true=y_test)
print(test_acc)
# -
| forest_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy.stats import mannwhitneyu
import numpy as np
import pandas as pd
# #### Mann-Whitney u test
# * Non-parametric test
# * Null: two samples are from the same population (not distribution)
population = np.loadtxt('data/finches.csv')
# draw sample 1, 2 from the same population
sample1 = np.random.choice(a=population, size=100)
sample2 = np.random.choice(a=population, size=100)
u, p = mannwhitneyu(sample1, sample2)
p
# draw sample 3 fro ma normal distribution with same mean and variance
sample3 = np.random.normal(np.mean(population), np.std(population), 100)
u, p = mannwhitneyu(sample1, sample3)
p
| statistics_test/mann_whitney.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multivariate Logistic Regression Demo
#
# > ☝Before moving on with this demo you might want to take a look at:
# > - 📗[Math behind the Logistic Regression](https://github.com/trekhleb/homemade-machine-learning/tree/master/homemade/logistic_regression)
# > - ⚙️[Logistic Regression Source Code](https://github.com/trekhleb/homemade-machine-learning/blob/master/homemade/logistic_regression/logistic_regression.py)
#
# **Logistic regression** is the appropriate regression analysis to conduct when the dependent variable is dichotomous (binary). Like all regression analyses, the logistic regression is a predictive analysis. Logistic regression is used to describe data and to explain the relationship between one dependent binary variable and one or more nominal, ordinal, interval or ratio-level independent variables.
#
# Logistic Regression is used when the dependent variable (target) is categorical.
#
# For example:
#
# - To predict whether an email is spam (`1`) or (`0`).
# - Whether online transaction is fraudulent (`1`) or not (`0`).
# - Whether the tumor is malignant (`1`) or not (`0`).
#
# > **Demo Project:** In this example we will train handwritten digits (0-9) classifier.
# +
# To make debugging of logistic_regression module easier we enable imported modules autoreloading feature.
# By doing this you may change the code of logistic_regression library and all these changes will be available here.
# %load_ext autoreload
# %autoreload 2
# Add project root folder to module loading paths.
import sys
sys.path.append('../..')
# -
# ### Import Dependencies
#
# - [pandas](https://pandas.pydata.org/) - library that we will use for loading and displaying the data in a table
# - [numpy](http://www.numpy.org/) - library that we will use for linear algebra operations
# - [matplotlib](https://matplotlib.org/) - library that we will use for plotting the data
# - [math](https://docs.python.org/3/library/math.html) - math library that we will use to calculate sqaure roots etc.
# - [logistic_regression](https://github.com/trekhleb/homemade-machine-learning/blob/master/homemade/logistic_regression/logistic_regression.py) - custom implementation of logistic regression
# +
# Import 3rd party dependencies.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import math
# Import custom logistic regression implementation.
from homemade.logistic_regression import LogisticRegression
# -
# ### Load the Data
#
# In this demo we will be using a sample of [MNIST dataset in a CSV format](https://www.kaggle.com/oddrationale/mnist-in-csv/home). Instead of using full dataset with 60000 training examples we will use cut dataset of just 10000 examples that we will also split into training and testing sets.
#
# Each row in the dataset consists of 785 values: the first value is the label (a number from 0 to 9) and the remaining 784 values (28x28 pixels image) are the pixel values (a number from 0 to 255).
# +
# Load the data.
data = pd.read_csv('../../data/mnist-demo.csv')
# Print the data table.
data.head(10)
# -
# ### Plot the Data
#
# Let's peek first 25 rows of the dataset and display them as an images to have an example of digits we will be working with.
# +
# How many numbers to display.
numbers_to_display = 25
# Calculate the number of cells that will hold all the numbers.
num_cells = math.ceil(math.sqrt(numbers_to_display))
# Make the plot a little bit bigger than default one.
plt.figure(figsize=(10, 10))
# Go through the first numbers in a training set and plot them.
for plot_index in range(numbers_to_display):
# Extrace digit data.
digit = data[plot_index:plot_index + 1].values
digit_label = digit[0][0]
digit_pixels = digit[0][1:]
# Calculate image size (remember that each picture has square proportions).
image_size = int(math.sqrt(digit_pixels.shape[0]))
# Convert image vector into the matrix of pixels.
frame = digit_pixels.reshape((image_size, image_size))
# Plot the number matrix.
plt.subplot(num_cells, num_cells, plot_index + 1)
plt.imshow(frame, cmap='Greys')
plt.title(digit_label)
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
# Plot all subplots.
plt.subplots_adjust(hspace=0.5, wspace=0.5)
plt.show()
# -
# ### Split the Data Into Training and Test Sets
#
# In this step we will split our dataset into _training_ and _testing_ subsets (in proportion 80/20%).
#
# Training data set will be used for training of our model. Testing dataset will be used for validating of the model. All data from testing dataset will be new to model and we may check how accurate are model predictions.
# +
# Split data set on training and test sets with proportions 80/20.
# Function sample() returns a random sample of items.
pd_train_data = data.sample(frac=0.8)
pd_test_data = data.drop(pd_train_data.index)
# Convert training and testing data from Pandas to NumPy format.
train_data = pd_train_data.values
test_data = pd_test_data.values
# Extract training/test labels and features.
num_training_examples = 6000
x_train = train_data[:num_training_examples, 1:]
y_train = train_data[:num_training_examples, [0]]
x_test = test_data[:, 1:]
y_test = test_data[:, [0]]
# -
# ### Init and Train Logistic Regression Model
#
# > ☝🏻This is the place where you might want to play with model configuration.
#
# - `polynomial_degree` - this parameter will allow you to add additional polynomial features of certain degree. More features - more curved the line will be.
# - `max_iterations` - this is the maximum number of iterations that gradient descent algorithm will use to find the minimum of a cost function. Low numbers may prevent gradient descent from reaching the minimum. High numbers will make the algorithm work longer without improving its accuracy.
# - `regularization_param` - parameter that will fight overfitting. The higher the parameter, the simplier is the model will be.
# - `polynomial_degree` - the degree of additional polynomial features (`x1^2 * x2, x1^2 * x2^2, ...`). This will allow you to curve the predictions.
# - `sinusoid_degree` - the degree of sinusoid parameter multipliers of additional features (`sin(x), sin(2*x), ...`). This will allow you to curve the predictions by adding sinusoidal component to the prediction curve.
# - `normalize_data` - boolean flag that indicates whether data normalization is needed or not.
# +
# Set up linear regression parameters.
max_iterations = 10000 # Max number of gradient descent iterations.
regularization_param = 10 # Helps to fight model overfitting.
polynomial_degree = 0 # The degree of additional polynomial features.
sinusoid_degree = 0 # The degree of sinusoid parameter multipliers of additional features.
normalize_data = True # Whether we need to normalize data to make it more unifrom or not.
# Init logistic regression instance.
logistic_regression = LogisticRegression(x_train, y_train, polynomial_degree, sinusoid_degree, normalize_data)
# Train logistic regression.
(thetas, costs) = logistic_regression.train(regularization_param, max_iterations)
# -
# ### Print Training Results
#
# Let's see how model parameters (thetas) look like. For each digit class (from 0 to 9) we've just trained a set of 784 parameters (one theta for each image pixel). These parameters represents the importance of every pixel for specific digit recognition.
# Print thetas table.
pd.DataFrame(thetas)
# +
# How many numbers to display.
numbers_to_display = 9
# Calculate the number of cells that will hold all the numbers.
num_cells = math.ceil(math.sqrt(numbers_to_display))
# Make the plot a little bit bigger than default one.
plt.figure(figsize=(10, 10))
# Go through the thetas and print them.
for plot_index in range(numbers_to_display):
# Extrace digit data.
digit_pixels = thetas[plot_index][1:]
# Calculate image size (remember that each picture has square proportions).
image_size = int(math.sqrt(digit_pixels.shape[0]))
# Convert image vector into the matrix of pixels.
frame = digit_pixels.reshape((image_size, image_size))
# Plot the number matrix.
plt.subplot(num_cells, num_cells, plot_index + 1)
plt.imshow(frame, cmap='Greys')
plt.title(plot_index)
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
# Plot all subplots.
plt.subplots_adjust(hspace=0.5, wspace=0.5)
plt.show()
# -
# ### Analyze Gradient Descent Progress
#
# The plot below illustrates how the cost function value changes over each iteration. You should see it decreasing.
#
# In case if cost function value increases it may mean that gradient descent missed the cost function minimum and with each step it goes further away from it.
#
# From this plot you may also get an understanding of how many iterations you need to get an optimal value of the cost function.
# +
# Draw gradient descent progress for each label.
labels = logistic_regression.unique_labels
for index, label in enumerate(labels):
plt.plot(range(len(costs[index])), costs[index], label=labels[index])
plt.xlabel('Gradient Steps')
plt.ylabel('Cost')
plt.legend()
plt.show()
# -
# ### Calculate Model Training Precision
#
# Calculate how many of training and test examples have been classified correctly. Normally we need test precission to be as high as possible. In case if training precision is high and test precission is low it may mean that our model is overfitted (it works really well with the training data set but it is not good at classifying new unknown data from the test dataset). In this case you may want to play with `regularization_param` parameter to fighth the overfitting.
# +
# Make training set predictions.
y_train_predictions = logistic_regression.predict(x_train)
y_test_predictions = logistic_regression.predict(x_test)
# Check what percentage of them are actually correct.
train_precision = np.sum(y_train_predictions == y_train) / y_train.shape[0] * 100
test_precision = np.sum(y_test_predictions == y_test) / y_test.shape[0] * 100
print('Training Precision: {:5.4f}%'.format(train_precision))
print('Test Precision: {:5.4f}%'.format(test_precision))
# -
# ### Plot Test Dataset Predictions
#
# In order to illustrate how our model classifies unknown examples let's plot first 64 predictions for testing dataset. All green digits on the plot below have been recognized corrctly but all the red digits have not been recognized correctly by our classifier. On top of each digit image you may see the class (the number) that has been recognized on the image.
# +
# How many numbers to display.
numbers_to_display = 64
# Calculate the number of cells that will hold all the numbers.
num_cells = math.ceil(math.sqrt(numbers_to_display))
# Make the plot a little bit bigger than default one.
plt.figure(figsize=(15, 15))
# Go through the first numbers in a test set and plot them.
for plot_index in range(numbers_to_display):
# Extrace digit data.
digit_label = y_test[plot_index, 0]
digit_pixels = x_test[plot_index, :]
# Predicted label.
predicted_label = y_test_predictions[plot_index][0]
# Calculate image size (remember that each picture has square proportions).
image_size = int(math.sqrt(digit_pixels.shape[0]))
# Convert image vector into the matrix of pixels.
frame = digit_pixels.reshape((image_size, image_size))
# Plot the number matrix.
color_map = 'Greens' if predicted_label == digit_label else 'Reds'
plt.subplot(num_cells, num_cells, plot_index + 1)
plt.imshow(frame, cmap=color_map)
plt.title(predicted_label)
plt.tick_params(axis='both', which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
# Plot all subplots.
plt.subplots_adjust(hspace=0.5, wspace=0.5)
plt.show()
| notebooks/logistic_regression/multivariate_logistic_regression_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Verifique se é transformação linear</b>
# $T(x, y) = (x^2, y^2)$
# $v = (x_1, y_1)$
# $u = (x_2, y_2)$
# $u + v = (x_1 + x_2, y_1 + y_2)$
# $T(u + v) = T(u) + T(v)$
# $T(u + v) = ((x_1 + x_2)^2, (y_1 + y_2)^2) = (x_1^2 + 2x_1x_2 + x_2^2, y_1^2 + 2y_1y_2 + y_2^2)$
# $T(u) = (x_2^2, u_2^2)$
# $T(v) = (x_1^2, u_1^2)$
# $(x_1^2 + 2x_1x_2 + x_2^2, y_1^2 + 2y_1y_2 + y_2^2) \neq (x_2^2, u_2^2) + (x_1^2, u_1^2)$
| Grings/10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spatial Joins
#
# A *spatial join* uses [binary predicates](http://shapely.readthedocs.io/en/latest/manual.html#binary-predicates)
# such as `intersects` and `crosses` to combine two `GeoDataFrames` based on the spatial relationship
# between their geometries.
#
# A common use case might be a spatial join between a point layer and a polygon layer where you want to retain the point geometries and grab the attributes of the intersecting polygons.
#
# 
#
# ## Types of spatial joins
#
# We currently support the following methods of spatial joins. We refer to the *left_df* and *right_df* which are the correspond to the two dataframes passed in as args.
#
# ### Left outer join
#
# In a LEFT OUTER JOIN (`how='left'`), we keep *all* rows from the left and duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the right if they intersect and lose right rows that don't intersect. A left outer join implies that we are interested in retaining the geometries of the left.
#
# This is equivalent to the PostGIS query:
# ```
# SELECT pts.geom, pts.id as ptid, polys.id as polyid
# FROM pts
# LEFT OUTER JOIN polys
# ON ST_Intersects(pts.geom, polys.geom);
#
# geom | ptid | polyid
# --------------------------------------------+------+--------
# 010100000040A9FBF2D88AD03F349CD47D796CE9BF | 4 | 10
# 010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 10
# 010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 20
# 0101000000F0D88AA0E1A4EEBF7052F7E5B115E9BF | 2 | 20
# 0101000000818693BA2F8FF7BF4ADD97C75604E9BF | 1 |
# (5 rows)
# ```
#
# ### Right outer join
#
# In a RIGHT OUTER JOIN (`how='right'`), we keep *all* rows from the right and duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the left if they intersect and lose left rows that don't intersect. A right outer join implies that we are interested in retaining the geometries of the right.
#
# This is equivalent to the PostGIS query:
# ```
# SELECT polys.geom, pts.id as ptid, polys.id as polyid
# FROM pts
# RIGHT OUTER JOIN polys
# ON ST_Intersects(pts.geom, polys.geom);
#
# geom | ptid | polyid
# ----------+------+--------
# 01...9BF | 4 | 10
# 01...9BF | 3 | 10
# 02...7BF | 3 | 20
# 02...7BF | 2 | 20
# 00...5BF | | 30
# (5 rows)
# ```
#
# ### Inner join
#
# In an INNER JOIN (`how='inner'`), we keep rows from the right and left only where their binary predicate is `True`. We duplicate them if necessary to represent multiple hits between the two dataframes. We retain attributes of the right and left only if they intersect and lose all rows that do not. An inner join implies that we are interested in retaining the geometries of the left.
#
# This is equivalent to the PostGIS query:
# ```
# SELECT pts.geom, pts.id as ptid, polys.id as polyid
# FROM pts
# INNER JOIN polys
# ON ST_Intersects(pts.geom, polys.geom);
#
# geom | ptid | polyid
# --------------------------------------------+------+--------
# 010100000040A9FBF2D88AD03F349CD47D796CE9BF | 4 | 10
# 010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 10
# 010100000048EABE3CB622D8BFA8FBF2D88AA0E9BF | 3 | 20
# 0101000000F0D88AA0E1A4EEBF7052F7E5B115E9BF | 2 | 20
# (4 rows)
# ```
# ## Spatial Joins between two GeoDataFrames
#
# Let's take a look at how we'd implement these using `GeoPandas`. First, load up the NYC test data into `GeoDataFrames`:
# +
# %matplotlib inline
from shapely.geometry import Point
from geopandas import datasets, GeoDataFrame, read_file
# NYC Boros
zippath = datasets.get_path('nybb')
polydf = read_file(zippath)
# Generate some points
b = [int(x) for x in polydf.total_bounds]
N = 8
pointdf = GeoDataFrame([
{'geometry': Point(x, y), 'value1': x + y, 'value2': x - y}
for x, y in zip(range(b[0], b[2], int((b[2] - b[0]) / N)),
range(b[1], b[3], int((b[3] - b[1]) / N)))])
# Make sure they're using the same projection reference
pointdf.crs = polydf.crs
# -
pointdf
polydf
pointdf.plot()
polydf.plot()
# ## Joins
join_left_df = pointdf.sjoin(polydf, how="left")
join_left_df
# Note the NaNs where the point did not intersect a boro
join_right_df = pointdf.sjoin(polydf, how="right")
join_right_df
# Note Staten Island is repeated
join_inner_df = pointdf.sjoin(polydf, how="inner")
join_inner_df
# Note the lack of NaNs; dropped anything that didn't intersect
# We're not limited to using the `intersection` binary predicate. Any of the `Shapely` geometry methods that return a Boolean can be used by specifying the `op` kwarg.
pointdf.sjoin(polydf, how="left", predicate="within")
# We can also conduct a nearest neighbour join with `sjoin_nearest`.
pointdf.sjoin_nearest(polydf, how="left", distance_col="Distances")
# Note the optional Distances column with computed distances between each point
# and the nearest polydf geometry.
| doc/source/gallery/spatial_joins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import csv
path="C://Users//oorte//Documents//RICE-CLASS-MATERIAL//team3//YK_TX2017-19_Accidents.csv"
traffic_df = pd.read_csv(path)
# -
#import Dependencies
import time
import datetime
import numpy as np
from datetime import date
# Read strings representing Start and End of accident
date_start=traffic_df["Start_Time"]
date_end=traffic_df["End_Time"]
traffic_df['start']=pd.DataFrame(pd.to_datetime(date_start, format='%Y-%m-%d %H:%M:%S'))
traffic_df['end']=pd.DataFrame(pd.to_datetime(date_end, format='%Y-%m-%d %H:%M:%S'))
traffic_df['month'] = pd.DatetimeIndex(traffic_df['start']).month
traffic_df['season']=traffic_df['month']
traffic_df.loc[traffic_df.season == 1, "season"] = "winter"
traffic_df.loc[traffic_df.season == 2, "season"] = "winter"
traffic_df.loc[traffic_df.season == 12, "season"] = "winter"
traffic_df.loc[traffic_df.season == 3, "season"] = "spring"
traffic_df.loc[traffic_df.season == 4, "season"] = "spring"
traffic_df.loc[traffic_df.season == 5, "season"] = "spring"
traffic_df.loc[traffic_df.season == 6, "season"] = "summer"
traffic_df.loc[traffic_df.season == 7, "season"] = "summer"
traffic_df.loc[traffic_df.season == 8, "season"] = "summer"
traffic_df.loc[traffic_df.season == 9, "season"] = "fall"
traffic_df.loc[traffic_df.season == 10, "season"] = "fall"
traffic_df.loc[traffic_df.season == 11, "season"] = "fall"
seasons_accidents_df=pd.DataFrame((pd.DataFrame(traffic_df.groupby(['season']).count())).iloc[:,1])
seasons_accidents_df.columns=['count']
seasons_accidents_df.reset_index()
import matplotlib.pyplot as plt
# +
# Generate a bar plot showing the total number of accidents per season.
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
seasons = ['summer', 'winter']
count = [seasons_accidents_df['count'][2],seasons_accidents_df['count'][3]]
ax.bar(seasons,count)
plt.bar(seasons, count, color='green')
plt.xlabel("Seasons")
plt.ylabel("count")
plt.title("Number of Accidents per Season")
plt.show()
# -
|
| oo_12202020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.1
# language: julia
# name: julia-1.5
# ---
# # loop on the OCs to estimate isochrone and mass
# +
using PyCall
## PopStar modules...
synthetic= pyimport("popstar.synthetic")
evolution= pyimport("popstar.evolution")
atmospheres= pyimport("popstar.atmospheres")
reddening= pyimport("popstar.reddening")
using Distributions, Statistics, Distances
using DataFrames
using Random , Printf, Glob
import PyPlot , CSV
import Distances, StatsBase
rootdir = ENV["GAIA_ROOT"]
push!(LOAD_PATH,"$rootdir/master/src")
using GaiaClustering
## directory
wdir = "$rootdir/products"
plotdir = "$rootdir/products/plotdens"
ocdir= "$wdir/oc2"
isodir= "$wdir/isochrones"
cd(wdir)
# +
### functions
function mass_isochrone(f)
fname= f[1:end-4] ; fvot= f[1:end-7]
ocfile= "$ocdir/$fname.csv" ; voname= "$fvot.vot"
ocmassfile= "$ocdir/$fname-mass.csv"
df= CSV.read(ocfile , delim= ";")
t= split(fvot,"-")
if length(t) == 3
t[1]= t[1]*"-"*t[2]
end
ocname= t[1]
println("## Name: $fname")
logage= df1[(df1.name .== ocname), :log_age]
# println(logage[1])
################################
## ISOCHRONES ##################
################################
agmedian= median(df.ag[.! isnan.(df.ag)])
distance= median(df.distance)
nstar= length(df.distance)
println("## N(star): $nstar")
# Define isochrone parameters
logAge= logage[1] # Age in log(years)
AKs= 0.0 # extinction in mags
dist= distance # distance in parsec
metallicity= 0.0 # Metallicity in [M/H]
println("### log Age: $logAge")
println("### AKs: $AKs")
println("### distance: $dist")
println("### metallicity: $metallicity")
## Define evolution/atmosphere models and extinction law
if logAge>6.5
evo_model = evolution.MergedBaraffePisaEkstromParsec()
else
evo_model= evolution.MISTv1()
end
atm_func = atmospheres.get_merged_atmosphere
red_law = reddening.RedLawHosek18b()
## Filter
filt_list = [ "gaia,dr2_rev,Gbp", "gaia,dr2_rev,G" ,"gaia,dr2_rev,Grp"]
#####################################
println("### Computing isochrones..")
iso = synthetic.IsochronePhot(logAge, AKs, dist, metallicity=metallicity,
evo_model=evo_model, atm_func=atm_func,
red_law=red_law, filters=filt_list, iso_dir= isodir)
println("### Updating the DF and creating the new one with stellar mass..")
data= iso.points
println(data.keys())
ndata= size(data)[1]
mass= zeros(ndata)
Gmag= zeros(ndata)
BPmag= zeros(ndata)
RPmag= zeros(ndata)
color= zeros(ndata)
for i in 1:ndata
mass[i]= data[i][4]
Gmag[i]= data[i][10]
BPmag[i]= data[i][9]
RPmag[i]= data[i][11]
color[i]= BPmag[i]-RPmag[i]
end
## Cluster
BmR= df.bp .- df.rp
GMAG= df.gbar + 5 .* log10.(df.distance) .- 17.
### Stellar mass
################
nxy= length(mass)
A= Array{Float64}(undef,2,nxy)
for i in 1:nxy
A[1,i]= color[i]
A[2,i]= Gmag[i]
end
d= Distances.Euclidean()
mstar= zeros(length(BmR))
for i in 1:length(BmR)
starm= [BmR[i]; GMAG[i]]
r= Distances.colwise(d, A, starm)
idx= argmin(r)
mstar[i]= mass[idx]
end
df.mass= mstar
## Some Logs mass stats...
massmean= mean(mstar)
println("### Mass (mean): $massmean")
### save the OC df
CSV.write(ocmassfile, df, delim=";")
println("\n\n")
end
# +
cd(ocdir)
files= glob("*-oc.csv")
cd(wdir)
sample= rootdir*"/master/notebooks/data/BrowseTargets.18292.1530479692.fake"
fileres= "isolist-done.csv"
df1= CSV.read(sample, delim= "|")
df1.name= strip.(df1.name)
noc= 0
for f in files
filered= CSV.read(fileres,delim=";",copycols= true)
if !(f in filered.fileoc)
println(f)
mass_isochrone(f)
push!(filered,(f,))
CSV.write(fileres,filered,delim=";")
end
end
| notebooks/analysis/loop_isochroneMass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Classification
#
# <NAME> (2016,2018), based particularly on materials from <NAME>, also Ivezic.
#
# Density estimation and clustering are **unsupervised** forms of classification. Let's now move on to **supervised**
# classification. That's where we actually know the "truth" for some of our objects and can use that information to help guide the classification of unknown objects.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Generative vs. Discriminative Classification
#
# We will talk about two different types of classification where each has a slightly different approach. As an example, if you are trying to determine whether your neighbor is speaking Spanish or Portuguese, you could 1) learn both Spanish and Portuguese so that you'd know exactly what they are saying or 2) learn the keys rules about the differences between the languages.
#
# If we find ourselves asking which category is most likely to generate the observed result, then we are using using **density estimation** for classification and this is referred to as **generative classification**. Here we have a full model of the density for each class or we have a model which describes how data could be generated from each class.
#
# If, on the other hand, we don't care about the full distribution, then we are doing something more like clustering, where we don't need to map the distribution, we just need to define boundaries. Classification that finds the **decision boundary** that separates classes is called **discriminative classification**. For high-dimensional data, this may be a better choice.
# + [markdown] slideshow={"slide_type": "slide"}
# For example, in the figure below, to classify a new object with $x=1$, it would suffice to know that either 1) model 1 is a better fit than model 2, or 2) that the decision boundary is at $x=1.4$.
#
# 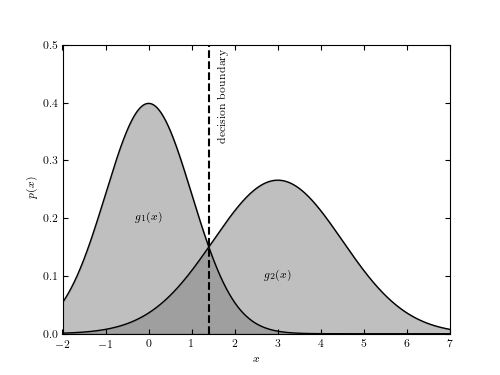
#
# In my work, we actually do both. We first do discriminative classification using a decision boundary based on $K$-$D$ trees and then we do generative classification using density estimation for the class of interest (in order to determine a probability).
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Scoring Your Results
#
# The first question that we need to address is how we score our results (defined the success of our classification).
#
# In the simplest case, there are 2 types of errors:
# * a [False Positive](https://en.wikipedia.org/wiki/False_positives_and_false_negatives#False_positive_error), where we have assigned a *true* class label when it is really false. This is called a "Type-1 error".
# * a [False Negative](https://en.wikipedia.org/wiki/False_positives_and_false_negatives#False_positive_error), where we have assigned a *false* class label when it is really true. This is called a "Type-II error".
#
# All 4 [possibilities](https://en.wikipedia.org/wiki/Sensitivity_and_specificity) are:
# - True Positive = **correctly identified** (apple identified as apple)
# - True Negative = **correctly rejected** (orange rejected as orange)
# - False Positive = **incorrectly identified** (orange identified as apple)
# - False Negative = **incorrectly rejected** (apple rejected as orange)
# + [markdown] slideshow={"slide_type": "slide"}
# Based on these, we usually define either of the following pairs of terms. Which is used is largely a matter of preference in different fields, but we'll see that there are some key differences.
#
# >$ {\rm completeness} = \frac{\rm true\ positives}{\rm true\ positives + false\ negatives}$
#
# >$ {\rm contamination} = \frac{\rm false\ positives}{\rm true\ positives + false\ positives} = {\rm false\ discovery\ rate}$
#
# or
#
# > $ {\rm true\ positive\ rate} = \frac{\rm true\ positives} {\rm true\ positives + false\ negatives}
# $
#
# > $ {\rm false\ positive\ rate} = \frac{\rm false\ positives} {\rm true\ negatives + false\ positives} = {\rm Type1\ error}
# $
#
# where **completeness** = **true positive rate** and this is also called **sensitivity** or **recall**.
#
# Similarly
#
# >$ {\rm efficiency} = 1 - {\rm contamination} = {\rm precision}. $
#
# Scikit-Learn also reports the **F1 score** which is the harmonic mean of precision and sensitivity (efficiency and completeness).
#
# Depending on your goals, you may want to maximize the completeness or the efficiency, or a combination of both.
# + [markdown] slideshow={"slide_type": "slide"}
# For example you might want to minimize voter fraud (contamination), but if doing so reduced voter participation (completeness) by a larger amount, then that wouldn't be such a good thing. So you need to decide what balance you want to strike.
# + [markdown] slideshow={"slide_type": "slide"}
# To better understand the differences between these measures, let's take the needle in a haystack problem. You have 100,000 stars and 1000 quasars. If you correctly identify 900 quasars as such and mistake 1000 stars for quasars, then we have:
# - TP = 900
# - FN = 100
# - TN = 99,000
# - FP = 1000
#
# Which gives
#
# > $ {\rm true\ positive\ rate} = \frac{900}{900 + 100} = 0.9 = {\rm completeness}
# $
#
# > $ {\rm false\ positive\ rate} = \frac{1000}{99000 + 1000} = 0.01
# $
#
# Not bad right? Well, sort of. The FPR isn't bad, but there are *lots* of stars, so the contamination rate isn't so great:
#
# > $ {\rm contamination} = \frac{1000}{900 + 1000} = 0.53
# $
# + [markdown] slideshow={"slide_type": "slide"}
# ## Comparing the performance of classifiers
#
# So, "best" performance is a bit of a subjective topic. We trade contamination as a function of completeness and this is science dependent.
#
# Before we start talking about different classification algorithms, let's first talk about how we can quantify which of the methods is "best". (N.B. We have skipped ahead to Ivezic $\S$ 9.8).
#
# The way that we will do this is with a [**Receiver Operating Characteristic (ROC)**](https://en.wikipedia.org/wiki/Receiver_operating_characteristic) curve. (Apparently this is *yet another* example in statistics/machine learning where the name of something was deliberately chosen to scare people away from the field.) A ROC curve simply plots the true-positive vs. the false-positive rate.
#
# One concern about ROC curves is that they are sensitive to the relative sample sizes. As we already demonstrated above, if there are many more background events than source events small false positive results can dominate a signal. For these cases we can plot efficiency (1 - contamination) vs. completeness.
#
# Indeed, I had never even heard of a ROC curve until I started preparing this class. I have always made "completeness-contamination" plots, which makes a lot more sense to me (both in terms of what can be learned and nomenclature).
# + [markdown] slideshow={"slide_type": "slide"}
# Here is a comparison of the two types of plots:
#
# 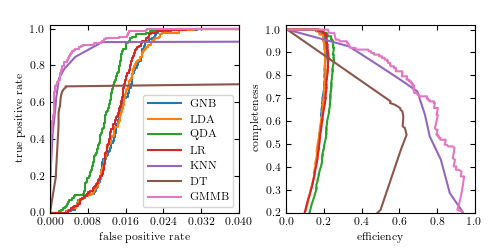
#
# Here we see that to get higher completeness, you could actually suffer significantly in terms of efficiency, but your FPR might not go up that much if there are lots of true negatives. I'll point this out again later when we do a specific example.
# + [markdown] slideshow={"slide_type": "slide"}
# Below is the code that makes these plots. We'll talk about the data that goes into it in a bit. For now, we'll concentrate on how to generate the ROC and completeness-contamination plots.
#
# We'll be comparing 7 different classifiers (with a generic `clf` object), making training and test sets with `split_samples`, then using these tools to generate our plots:
#
# - [sklearn.metrics.roc_curve(y_test, y_prob)](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html)
# - [sklearn.metrics.precision_recall_curve(y_test, y_prob)](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_curve.html)
# - astroML.utils.completeness_contamination(y_pred, y_test)
#
# Note that the [`sklearn.metrics` algorithms](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics) take `y_test`, which are classes, and `y_prob`, which are not class predictions, but rather probabilities, whereas the AstroML algorithm wants `y_pred` (which we get by converting `y_prob` into discrete predictions as a function of the probability).
#
# Generally speaking, you want to chose a decision boundary (see below) that maximizes the area under the curve.
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
# Ivezic, Figure 9.17
# Author: <NAME>
# License: BSD
import numpy as np
from matplotlib import pyplot as plt
from sklearn.naive_bayes import GaussianNB
#from sklearn.lda import LDA
#from sklearn.qda import QDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from astroML.classification import GMMBayes
from sklearn.metrics import precision_recall_curve, roc_curve
from astroML.utils import split_samples, completeness_contamination
from astroML.datasets import fetch_rrlyrae_combined
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined()
y = y.astype(int)
(X_train, X_test), (y_train, y_test) = split_samples(X, y, [0.75, 0.25], random_state=0)
#------------------------------------------------------------
# Fit all the models to the training data
def compute_models(*args):
names = []
probs = []
for classifier, kwargs in args:
clf = classifier(**kwargs)
clf.fit(X_train, y_train)
y_probs = clf.predict_proba(X_test)[:, 1]
names.append(classifier.__name__)
probs.append(y_probs)
return names, probs
names, probs = compute_models((GaussianNB, {}),
(LinearDiscriminantAnalysis, {}),
(QuadraticDiscriminantAnalysis, {}),
(LogisticRegression, dict(class_weight=None)),
(KNeighborsClassifier, dict(n_neighbors=10)),
(DecisionTreeClassifier, dict(random_state=0, max_depth=12, criterion='entropy')),
(GMMBayes, dict(n_components=3, min_covar=1E-5, covariance_type='full')))
#------------------------------------------------------------
# Plot ROC curves and completeness/efficiency
fig = plt.figure(figsize=(18, 6))
fig.subplots_adjust(left=0.1, right=0.95, bottom=0.15, top=0.9, wspace=0.25)
# ax1 will show roc curves
ax1 = plt.subplot(131)
# ax2 will show precision/recall
ax2 = plt.subplot(132)
# ax3 will show completeness/efficiency
ax3 = plt.subplot(133)
labels = dict(GaussianNB='GNB',
LinearDiscriminantAnalysis='LDA',
QuadraticDiscriminantAnalysis='QDA',
KNeighborsClassifier='KNN',
DecisionTreeClassifier='DT',
GMMBayes='GMMB',
LogisticRegression='LR')
thresholds = np.linspace(0, 1, 1001)[:-1]
# iterate through and show results
for name, y_prob in zip(names, probs):
# Note that these take y_prob and not y_pred
fpr, tpr, thresh = roc_curve(y_test, y_prob)
precision, recall, thresh2 = precision_recall_curve(y_test, y_prob)
# add (0, 0) as first point
fpr = np.concatenate([[0], fpr])
tpr = np.concatenate([[0], tpr])
precision = np.concatenate([[0], precision])
recall = np.concatenate([[1], recall])
ax1.plot(fpr, tpr, label=labels[name])
ax2.plot(precision, recall, label=labels[name])
# Whereas this does take y_pred, which we need to compute
# by looping through all possible probability thresholds
comp = np.zeros_like(thresholds)
cont = np.zeros_like(thresholds)
for i, t in enumerate(thresholds):
y_pred = (y_prob >= t)
comp[i], cont[i] = completeness_contamination(y_pred, y_test)
ax3.plot(1-cont, comp, label=labels[name])
ax1.set_xlim(0, 0.04)
ax1.set_ylim(0, 1.02)
ax1.xaxis.set_major_locator(plt.MaxNLocator(5))
ax1.set_xlabel('false positive rate')
ax1.set_ylabel('true positive rate')
ax1.legend(loc=4)
ax2.set_xlabel('precision')
ax2.set_ylabel('recall')
ax2.set_xlim(0, 1.0)
ax2.set_ylim(0.2, 1.02)
ax3.set_xlabel('efficiency')
ax3.set_ylabel('completeness')
ax3.set_xlim(0, 1.0)
ax3.set_ylim(0.2, 1.02)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# Note that I've plotted both recall-precision and completeness-efficiency just to demonstrate that they are the same thing.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Generative Classification
#
# We can use Bayes' theorem to relate the labels to the features in an $N\times D$ data set $X$. The $j$th feature of the $i$th point is $x_{ij}$ and there are $k$ classes giving discrete labels $y_k$. Then we have
#
# $$p(y_k|x_i) = \frac{p(x_i|y_k)p(y_k)}{\sum_i p(x_i|y_k)p(y_k)},$$
#
# where $x_i$ is assumed to be a vector with $j$ components.
#
# $p(y=y_k)$ is the probability of any point having class $k$ (equivalent to the prior probability of the class $k$).
#
# In generative classifiers we model class-conditional densities $p_k(x) = p(x|y=y_k)$ and our goal is to estimate the $p_k$'s.
#
# Before we get into the generative classification algortithms, we'll first discuss 3 general concepts:
# - Discriminant Functions
# - Bayes Classifiers
# - Decision Boundaries
# + [markdown] slideshow={"slide_type": "slide"}
# ### The Discriminant Function
#
# We can relate classification to density estimation and regression.
#
# $\hat{y} = f(y|x)$ represents the best guess of $y$ given $x$. So classification can be thought of as the analog of regression where $y$ is a discrete *category* rather than a continuous variable, for example $y=\{0,1\}$.
#
#
# In classification we refer to $f(y|x)$ as the [**discriminant function**](https://en.wikipedia.org/wiki/Discriminant_function_analysis).
# + [markdown] slideshow={"slide_type": "slide"}
# For a simple 2-class example:
#
# $$\begin{eqnarray}
# g(x) = f(y|x) & = & \int y \, p(y|x) \, dy \\
# % & = & \int y p(y|x) \, dy \\
# & = & 1 \cdot p(y=1 | x) + 0 \cdot p(y=0 | x) = p(y=1 | x).
# % & = & p(y=1 | x)
# \end{eqnarray}
# $$
#
# From Bayes rule:
#
# $$g(x) = \frac{p(x|y=1) \, p(y=1)}{p(x|y=1) \, p(y=1) + p(x|y=0) \, p(y=0)}$$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bayes Classifier
#
# If the discriminant function gives a binary prediction, we call it a **Bayes classifier**, formulated as
#
# $$\begin{eqnarray} \widehat{y} & = & \left\{ \begin{array}{cl} 1 & \mbox{if $g(x) > 1/2$}, \\ 0 & \mbox{otherwise,} \end{array} \right. \\ & = & \left\{
# \begin{array}{cl} 1 & \mbox{if $p(y=1|x) > p(y=0|x)$}, \\ 0 & \mbox{otherwise.} \end{array} \right.\end{eqnarray}$$
#
# Where this can be generalized to any number of classes, $k$, and not just two.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Decision Boundary
#
# A **decision boundary** is just set of $x$ values at which each class is equally likely:
#
# $$
# p(x|y=1)p(y=1) = p(x|y=0)p(y=0);
# $$
#
# $$g_1(x) = g_2(x) \; {\rm or}\; g(x) = 1/2$$
#
# Below is an example of a decision boundary in 1-D. In short, we assign classifications according to which pdf is higher at every given $x$.
#
# 
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Simplest Classifier: Naive Bayes
#
# In practice classification can be very complicated as the data are generally multi-dimensional (that is we don't just have $x$, we have $x_{j=0},x_1,x_2,x_3...x_n$, so we want $p(x_0,x_1,x_2,x_3...x_n|y)$.
#
# However, if we **assume** that all attributes are conditionally independent (which is not always true, but is often close enough), then this simplifies to
#
# $$ p(x_1,x_2|y_k) = p(x_1|y)p(x_2|y_k)$$
#
# which can be written as
#
# $$ p({x_{j=0},x_1,x_2,\ldots,x_N}|y_k) = \prod_j p(x_j|y_k).$$
#
# From Bayes' rule and conditional independence we get
#
# $$
# p(y_k | {x_0,x_1,\ldots,x_N}) =
# \frac{\prod_j p(x_j|y_k) p(y_k)}
# {\sum_l \prod_j p(x_j|y_l) p(y_l)}.
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# We calculate the most likely value of $y$ by maximizing over $y_k$:
#
# $$
# \hat{y} = \arg \max_{y_k} \frac{\prod_j p(x_j|y_k) p(y_k)}
# {\sum_l \prod_j p(x_j|y_l) p(y_l)},
# $$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# From there the process is just estimating densities: $p(x|y=y_k)$ and $p(y=y_k)$ are learned from a set of training data, where
# - $p(y=y_k)$ is just the frequency of the class $k$ in the training set
# - $p(x|y=y_k)$ is just the density (probability) of an object with class $k$ having the attributes $x$
#
# A catch is that if the training set does not cover the full parameter space, then $p(x_i|y=y_k)$ can be $0$ for some value of $y_k$ and $x_i$. The posterior probability is then $p(y_k|\{x_i\}) = 0/0$ which is a problem! A trick called [**Laplace smoothing**](https://en.wikipedia.org/wiki/Laplacian_smoothing) can be implemented to fix it.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Gaussian Naive Bayes
#
# It is totally unclear from the discussion in the book that $x_i$ are discrete measurements. However, one way to handle continuous values for $X$ is to model $p(x_i|y=y_k)$ as one-dimensional normal distributions, with means $\mu_{ik}$ and widths $\sigma_{ik}$. The naive Bayes estimator is then
#
# $$\hat{y} = \arg\max_{y_k}\left[\ln p(y=y_k) - \frac{1}{2}\sum_{i=1}^N\left(2\pi(\sigma_{ik})^2 + \frac{(x_i - \mu_{ik})^2}{(\sigma_{ik})^2} \right) \right]$$
# + [markdown] slideshow={"slide_type": "slide"}
# In Scikit-Learn [`Gaussian Naive Bayes`](http://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html) classification is implemented as follows, with a simple example given in the next cell.
# + slideshow={"slide_type": "slide"}
import numpy as np
from sklearn.naive_bayes import GaussianNB
X = np.random.random((100,2))
y = (X[:,0] + X[:,1] > 1).astype(int)
gnb = GaussianNB()
gnb.fit(X,y)
y_pred = gnb.predict(X)
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
# Ivezic, Figure 9.2
# Author: <NAME>
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import colors
from sklearn.naive_bayes import GaussianNB
#------------------------------------------------------------
# Simulate some data
np.random.seed(0)
mu1 = [1, 1]
cov1 = 0.3 * np.eye(2)
mu2 = [5, 3]
cov2 = np.eye(2) * np.array([0.4, 0.1])
X = np.concatenate([np.random.multivariate_normal(mu1, cov1, 100),
np.random.multivariate_normal(mu2, cov2, 100)])
y = np.zeros(200)
y[100:] = 1
#------------------------------------------------------------
# Fit the Naive Bayes classifier
clf = GaussianNB()
clf.fit(X, y)
# predict the classification probabilities on a grid
xlim = (-1, 8)
ylim = (-1, 5)
xx, yy = np.meshgrid(np.linspace(xlim[0], xlim[1], 71), np.linspace(ylim[0], ylim[1], 81))
xystack = np.vstack([xx.ravel(),yy.ravel()])
Xgrid = xystack.T
Z = clf.predict_proba(Xgrid)
# Gives probability for both class 1 and class 2 for each grid point
# As these are degenerate, take just one and then
# re-shape it to the grid pattern needed for contour plotting
Z = Z[:, 1].reshape(xx.shape)
#------------------------------------------------------------
# Plot the results
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111)
# Plot the points
ax.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.PRGn)
# Add the decision boundary, which is just the contour where
# the probability exceeds some threshold, here 0.5.
ax.contour(xx, yy, Z, [0.5], colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# Why does the boundary look like that?
# + [markdown] slideshow={"slide_type": "slide"}
# And now an example using real data. Here we have a 4-D $X$ and we are going to take them 1-D at a time to see how much improvement comes from adding each new dimension of $X$.
# + slideshow={"slide_type": "slide"}
# Ivezic, Figure 9.3
# Author: <NAME>
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from sklearn.naive_bayes import GaussianNB
from astroML.datasets import fetch_rrlyrae_combined
from astroML.utils import split_samples
from astroML.utils import completeness_contamination
# Added by GTR
from sklearn.metrics import precision_recall_curve, roc_curve
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined() # X is a 4-D color-color-color-color space
X = X[:, [1, 0, 2, 3]] # rearrange columns for better 1-color results
# Split the data into training and test sets
(X_train, X_test), (y_train, y_test) = split_samples(X, y, [0.75, 0.25], random_state=0)
N_tot = len(y)
N_stars = np.sum(y == 0)
N_rrlyrae = N_tot - N_stars
N_train = len(y_train)
N_test = len(y_test)
N_plot = 5000 + N_rrlyrae
#----------------------------------------------------------------------
# perform Naive Bayes
# Create blank arrays to hold the output
y_class = []
y_pred = []
y_probs = []
Ncolors = np.arange(1, X.shape[1] + 1)
order = np.array([1, 0, 2, 3])
for nc in Ncolors:
clf = GaussianNB()
clf.fit(X_train[:, :nc], y_train)
y_pred.append(clf.predict(X_test[:, :nc]))
y_class.append(clf)
# Added by GTR
y_probs.append(clf.predict_proba(X_test[:, :nc])[:,1])
# Use astroML utils code to compute completeness and contamination
completeness, contamination = completeness_contamination(y_pred, y_test)
print "completeness", completeness
print "contamination", contamination
#------------------------------------------------------------
# Compute the decision boundary (for 2 colors)
clf = y_class[1]
xlim = (0.7, 1.35)
ylim = (-0.15, 0.4)
xx, yy = np.meshgrid(np.linspace(xlim[0], xlim[1], 81),
np.linspace(ylim[0], ylim[1], 71))
Z = clf.predict_proba(np.c_[yy.ravel(), xx.ravel()])
Z = Z[:, 1].reshape(xx.shape)
#----------------------------------------------------------------------
# plot the results
fig = plt.figure(figsize=(10, 5))
fig.subplots_adjust(bottom=0.15, top=0.95, hspace=0.0,
left=0.1, right=0.95, wspace=0.2)
# left plot: data and decision boundary
ax = fig.add_subplot(121)
im = ax.scatter(X[-N_plot:, 1], X[-N_plot:, 0], c=y[-N_plot:],
s=4, lw=0, cmap=plt.cm.Oranges, zorder=2)
im.set_clim(-0.5, 1)
im = ax.imshow(Z, origin='lower', aspect='auto',
cmap=plt.cm.binary, zorder=1,
extent=xlim + ylim)
im.set_clim(0, 1.5)
ax.contour(xx, yy, Z, [0.5], colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
# Plot completeness vs Ncolors
ax = plt.subplot(222)
ax.plot(Ncolors, completeness, 'o-k', ms=6)
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.set_ylabel('completeness')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
# Plot contamination vs Ncolors
ax = plt.subplot(224)
ax.plot(Ncolors, contamination, 'o-k', ms=6)
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.FormatStrFormatter('%i'))
ax.set_xlabel('N colors')
ax.set_ylabel('contamination')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# If you shifted the decision boundary "up" by hand, what would happen to the completeness, contamination, and false positive rate?
#
# What happens if you change the fraction of objects in the training set?
# + [markdown] slideshow={"slide_type": "slide"}
# How could we replace contamination panel with the false positive rate using `roc_curve`?
# + [markdown] slideshow={"slide_type": "slide"}
# The "naive" refers to the fact that we are assuming that all of the variable are independent. If we relax that assumption and allow for covariances, then we have a **Gaussian Bayes classifier**. But note that this comes with a large jump in computational cost!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Linear Discriminant Analysis
#
# In [Linear Discriminant Analysis (LDA)](https://en.wikipedia.org/wiki/Linear_discriminant_analysis) we assume that the class distributions have identical
# covariances for all $k$ classes (all classes are a set of shifted Gaussians).
#
# <!--- The optimal classifier is derived from the log of the class posteriors --->
#
# <!--- > $g_k(\vec{x}) = \vec{x}^T \Sigma^{-1} \vec{\mu_k} - \frac{1}{2}\vec{\mu_k}^T \Sigma^{-1} \vec{\mu_k} + \log \pi_k,$ --->
#
# <!--- with $\vec{\mu_k}$ the mean of class $k$ and $\Sigma$ the covariance of the Gaussians. --->
#
# <!--- ** note different from book --->
#
# The class-dependent covariances that would normally give rise to a quadratic dependence on
# $X$ cancel out if they are assumed to be constant. The Bayes classifier is, therefore, linear with respect to $X$, and discriminant boundary between classes is the line that minimizes
# the overlap between Gaussians.
#
# <!--- > $ g_k(\vec{x}) - g_\ell(\vec{x}) = \vec{x}^T \Sigma^{-1} (\mu_k-\mu_\ell) - \frac{1}{2}(\mu_k - \mu_\ell)^T \Sigma^{-1}(\mu_k -\mu_\ell) + \log (\frac{\pi_k}{\pi_\ell}) = 0. $ --->
#
# Relaxing the requirement that the covariances of the
# Gaussians are constant, the discriminant function
# becomes quadratic in $X$.
#
# <!--- > $ g(\vec{x}) = -\frac{1}{2} \log | \Sigma_k | - \frac{1}{2}(\vec{x}-\mu_k)^T C^{-1}(\vec{x}-\mu_k) + \log \pi_k. $ --->
#
# This is sometimes known as [Quadratic Discriminant Analysis (QDA)](https://en.wikipedia.org/wiki/Quadratic_classifier#Quadratic_discriminant_analysis).
#
# [`LDA`](http://scikit-learn.org/0.16/modules/generated/sklearn.lda.LDA.html) and [`QDA`](http://scikit-learn.org/0.16/modules/generated/sklearn.qda.QDA.html#sklearn.qda.QDA) are implemented in Scikit-Learn as follows and an example using the same data as above is given below.
# + slideshow={"slide_type": "slide"}
import numpy as np
#from sklearn.lda import LDA
#from sklearn.qda import QDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
X = np.random.random((100,2))
y = (X[:,0] + X[:,1] > 1).astype(int)
lda = LDA()
lda.fit(X,y)
y_pred = lda.predict(X)
qda = QDA()
qda.fit(X,y)
y_pred = qda.predict(X)
# + slideshow={"slide_type": "slide"}
# Ivezic, Figures 9.4 and 9.5
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import colors
#from sklearn.lda import LDA
#from sklearn.qda import QDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
from astroML.datasets import fetch_rrlyrae_combined
from astroML.utils import split_samples
from astroML.utils import completeness_contamination
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined()
X = X[:, [1, 0, 2, 3]] # rearrange columns for better 1-color results
(X_train, X_test), (y_train, y_test) = split_samples(X, y, [0.75, 0.25], random_state=0)
N_tot = len(y)
N_stars = np.sum(y == 0)
N_rrlyrae = N_tot - N_stars
N_train = len(y_train)
N_test = len(y_test)
N_plot = 5000 + N_rrlyrae
#----------------------------------------------------------------------
# perform LDA
lda = LDA()
lda.fit(X_train[:, :2], y_train)
y_predLDA = lda.predict(X_test[:, :2])
# perform QDA
qda = QDA()
qda.fit(X_train[:, :2], y_train)
y_predQDA = qda.predict(X_test[:, :2])
completenessLDA, contaminationLDA = completeness_contamination(y_predLDA, y_test)
completenessQDA, contaminationQDA = completeness_contamination(y_predQDA, y_test)
print "completeness", completenessLDA, completenessQDA
print "contamination", contaminationLDA, contaminationQDA
#------------------------------------------------------------
# Compute the decision boundary
xlim = (0.7, 1.35)
ylim = (-0.15, 0.4)
xx, yy = np.meshgrid(np.linspace(xlim[0], xlim[1], 71),
np.linspace(ylim[0], ylim[1], 81))
Z_LDA = lda.predict_proba(np.c_[yy.ravel(), xx.ravel()])
Z_LDA = Z_LDA[:, 1].reshape(xx.shape)
Z_QDA = qda.predict_proba(np.c_[yy.ravel(), xx.ravel()])
Z_QDA = Z_QDA[:, 1].reshape(xx.shape)
#----------------------------------------------------------------------
# plot the results
fig = plt.figure(figsize=(10, 5))
fig.subplots_adjust(bottom=0.15, top=0.95, hspace=0.0,
left=0.1, right=0.95, wspace=0.2)
# left plot: data and decision boundary
ax = fig.add_subplot(121)
im = ax.scatter(X[-N_plot:, 1], X[-N_plot:, 0], c=y[-N_plot:],
s=4, lw=0, cmap=plt.cm.Oranges, zorder=2)
im.set_clim(-0.5, 1)
im = ax.imshow(Z_LDA, origin='lower', aspect='auto',
cmap=plt.cm.binary, zorder=1,
extent=xlim + ylim)
im.set_clim(0, 1.5)
ax.contour(xx, yy, Z_LDA, [0.5], linewidths=2., colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
# right plot: qda
ax = fig.add_subplot(122)
im = ax.scatter(X[-N_plot:, 1], X[-N_plot:, 0], c=y[-N_plot:],
s=4, lw=0, cmap=plt.cm.Oranges, zorder=2)
im.set_clim(-0.5, 1)
im = ax.imshow(Z_QDA, origin='lower', aspect='auto',
cmap=plt.cm.binary, zorder=1,
extent=xlim + ylim)
im.set_clim(0, 1.5)
ax.contour(xx, yy, Z_QDA, [0.5], linewidths=2., colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# If it is obvious from looking at your data that a linear or quadratic boundary will work well, then great. But what if that is not the case?
# + [markdown] slideshow={"slide_type": "slide"}
# ## GMM and Bayes Classification
#
# Our classifications so far have made some restrictive assumptions: either that of conditional independence or that the distributions are Gaussian. However, a more flexible model might improve the completeness and efficiency of the classification. For that we can look to the techniques from Chapter 6.
#
# The natural extension of the Gaussian assumptions is to use GMM's to determine the density distribution, i.e., a **GMM Bayes Classifier**.
#
# Note that the number of Gaussian components, $K$, must be chosen for each class, $k$, independently.
#
# astroML implements GMM Bayes classification as:
# + slideshow={"slide_type": "slide"}
import numpy as np
from astroML.classification import GMMBayes
#from astroML.classification import GaussianMixture as GMMBayes
X = np.random.random((100,2))
y = (X[:,0] + X[:,1] > 1).astype(int)
gmmb = GMMBayes(3) # 3 clusters per class
gmmb.fit(X,y)
y_pred = gmmb.predict(X)
# + [markdown] slideshow={"slide_type": "slide"}
# We now apply the GMM Bayes classifier to the real data from above. With just one component, we get results that are similar to those from Naive Bayes. But with 5 components (and all 4 attributes), we do pretty well.
# + slideshow={"slide_type": "slide"}
# Ivezic, Figure 9.6
import numpy as np
from matplotlib import pyplot as plt
from astroML.classification import GMMBayes
#from astroML.decorators import pickle_results
from astroML.datasets import fetch_rrlyrae_combined
from astroML.utils import split_samples
from astroML.utils import completeness_contamination
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined()
X = X[:, [1, 0, 2, 3]] # rearrange columns for better 1-color results
# GMM-bayes takes several minutes to run, and is order[N^2]
# truncating the dataset can be useful for experimentation.
#X = X[::10]
#y = y[::10]
(X_train, X_test), (y_train, y_test) = split_samples(X, y, [0.75, 0.25], random_state=0)
N_tot = len(y)
N_stars = np.sum(y == 0)
N_rrlyrae = N_tot - N_stars
N_train = len(y_train)
N_test = len(y_test)
N_plot = 5000 + N_rrlyrae
#----------------------------------------------------------------------
# perform GMM Bayes
Ncolors = np.arange(1, X.shape[1] + 1)
Ncomp = [1, 5]
def compute_GMMbayes(Ncolors, Ncomp):
classifiers = []
predictions = []
for ncm in Ncomp:
classifiers.append([])
predictions.append([])
for nc in Ncolors:
clf = GMMBayes(ncm, min_covar=1E-5, covariance_type='full')
clf.fit(X_train[:, :nc], y_train)
y_pred = clf.predict(X_test[:, :nc])
classifiers[-1].append(clf)
predictions[-1].append(y_pred)
return classifiers, predictions
classifiers, predictions = compute_GMMbayes(Ncolors, Ncomp)
completeness, contamination = completeness_contamination(predictions, y_test)
print "completeness", completeness[0]
print "contamination", contamination[0]
#------------------------------------------------------------
# Compute the decision boundary
clf = classifiers[1][1]
xlim = (0.7, 1.35)
ylim = (-0.15, 0.4)
xx, yy = np.meshgrid(np.linspace(xlim[0], xlim[1], 71),
np.linspace(ylim[0], ylim[1], 81))
Z = clf.predict_proba(np.c_[yy.ravel(), xx.ravel()])
Z = Z[:, 1].reshape(xx.shape)
#----------------------------------------------------------------------
# plot the results
fig = plt.figure(figsize=(8, 4))
fig.subplots_adjust(bottom=0.15, top=0.95, hspace=0.0,
left=0.1, right=0.95, wspace=0.2)
# left plot: data and decision boundary
ax = fig.add_subplot(121)
im = ax.scatter(X[-N_plot:, 1], X[-N_plot:, 0], c=y[-N_plot:],
s=4, lw=0, cmap=plt.cm.binary, zorder=2)
im.set_clim(-0.5, 1)
im = ax.imshow(Z, origin='lower', aspect='auto',
cmap=plt.cm.Oranges, zorder=1,
extent=xlim + ylim)
im.set_clim(0, 1.5)
ax.contour(xx, yy, Z, [0.5], linewidths=2., colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
# plot completeness vs Ncolors
ax = fig.add_subplot(222)
ax.plot(Ncolors, completeness[0], '^--k', c='k', label='N=%i' % Ncomp[0])
ax.plot(Ncolors, completeness[1], 'o-k', c='k', label='N=%i' % Ncomp[1])
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.set_ylabel('completeness')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
# plot contamination vs Ncolors
ax = fig.add_subplot(224)
ax.plot(Ncolors, contamination[0], '^--k', c='k', label='N=%i' % Ncomp[0])
ax.plot(Ncolors, contamination[1], 'o-k', c='k', label='N=%i' % Ncomp[1])
ax.legend(prop=dict(size=12),
loc='lower right',
bbox_to_anchor=(1.0, 0.78))
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.FormatStrFormatter('%i'))
ax.set_xlabel('N colors')
ax.set_ylabel('contamination')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# We can take this to the extreme by having one mixture component at each training point. We also don't have to restrict ourselves to a Gaussian kernel, we can use any kernel that we like. The resulting *non-parametric* Bayes classifier is referred to as **Kernel Discriminant Analysis (KDA)** . It seems like this would be a *lot* more computationally intensive, but at least now we don't have to optimize the locations of the components, we just need to determine the bandwidth of the kernel. In the end, it can result in better classification.
#
# One of the tricks to speed things up is that we don't need to know the actually class probability, we just need to know which is larger. This is explained in more detail in [Riegel, Gray, & Richards 2008](http://epubs.siam.org/doi/abs/10.1137/1.9781611972788.19), and it is implemented in a series of papers starting with [Richards et al. 2004](http://adsabs.harvard.edu/abs/2004ApJS..155..257R).
#
# If you follow those and you are so inclined, you could apply for jobs at [Skytree](http://www.skytree.net/) or [Wise.io](http://www.wise.io/) which were started by my collaborator, <NAME>, and astronomer, <NAME>, respectively.
#
# It is worth noting that this illustrates one of the downsides of the book, astroML, and Scikit-learn. They teach you about the basics of the algorithms, but if you wanted to use a **truly** big data set, then you really need the next level, such as KDA, but that is merely described here, not implemented.
# + [markdown] slideshow={"slide_type": "slide"}
# ## K-Nearest Neighbor Classifier
#
# If we did KDA with a variable bandwidth that depended only on the distance of the nearest neighbor, then we'd have what we call a **Nearest-Neighbor** classifier. Here if $x$ is close to $x'$, then $p(y|x) \approx p(y|x')$. Note that we have not assumed anything about the conditional density distribution, so it is completely non-parametric.
#
# The number of neighbors, $K$, regulates the complexity of the classification, where a larger $K$ decreases the variance in the classification but leads to an increase in the bias. (N.B., the 3rd different use of $K$ or $k$ in this notebook!)
#
# The distance measure is usually N-D Euclidean. However, if the attributes have very different properties, then normalization, weighting, etc. may be needed.
#
# Scikit-learn implements [`K-Nearest Neighbors`](http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html) classification as
# + slideshow={"slide_type": "slide"}
import numpy as np
from sklearn.neighbors import KNeighborsClassifier
X = np.random.random((100,2))
y = (X[:,0] + X[:,1] > 1).astype(int)
knc = KNeighborsClassifier(5) # use 5 nearest neighbors
knc.fit(X,y)
y_pred = knc.predict(X)
# + [markdown] slideshow={"slide_type": "slide"}
# Implementing it for the same example as above shows that it isn't all that great for this particular case. See below. We probably need more training data to reduce the variance for it to work better.
# + slideshow={"slide_type": "slide"}
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import colors
from sklearn.neighbors import KNeighborsClassifier
from astroML.datasets import fetch_rrlyrae_combined
from astroML.utils import split_samples
from astroML.utils import completeness_contamination
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined()
X = X[:, [1, 0, 2, 3]] # rearrange columns for better 1-color results
(X_train, X_test), (y_train, y_test) = split_samples(X, y, [0.75, 0.25], random_state=0)
N_tot = len(y)
N_st = np.sum(y == 0)
N_rr = N_tot - N_st
N_train = len(y_train)
N_test = len(y_test)
N_plot = 5000 + N_rr
#----------------------------------------------------------------------
# perform Classification
classifiers = []
predictions = []
Ncolors = np.arange(1, X.shape[1] + 1)
kvals = [1, 17]
from sklearn import metrics
for k in kvals:
classifiers.append([])
predictions.append([])
for nc in Ncolors:
clf = KNeighborsClassifier(n_neighbors=k)
clf.fit(X_train[:, :nc], y_train)
y_pred = clf.predict(X_test[:, :nc])
classifiers[-1].append(clf)
predictions[-1].append(y_pred)
completeness, contamination = completeness_contamination(predictions, y_test)
print "completeness (as a fn of neighbors and colors)", completeness
print "contamination", contamination
#------------------------------------------------------------
# Compute the decision boundary
clf = classifiers[1][1]
xlim = (0.7, 1.35)
ylim = (-0.15, 0.4)
xx, yy = np.meshgrid(np.linspace(xlim[0], xlim[1], 71),
np.linspace(ylim[0], ylim[1], 81))
Z = clf.predict(np.c_[yy.ravel(), xx.ravel()])
Z = Z.reshape(xx.shape)
#----------------------------------------------------------------------
# plot the results
fig = plt.figure(figsize=(10, 5))
fig.subplots_adjust(bottom=0.15, top=0.95, hspace=0.0,
left=0.1, right=0.95, wspace=0.2)
# left plot: data and decision boundary
ax = fig.add_subplot(121)
im = ax.scatter(X[-N_plot:, 1], X[-N_plot:, 0], c=y[-N_plot:],
s=4, lw=0, cmap=plt.cm.Oranges, zorder=2)
im.set_clim(-0.5, 1)
im = ax.imshow(Z, origin='lower', aspect='auto',
cmap=plt.cm.binary, zorder=1,
extent=xlim + ylim)
im.set_clim(0, 2)
ax.contour(xx, yy, Z, [0.5], linewidths=2., colors='k')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
ax.text(0.02, 0.02, "k = %i" % kvals[1],
transform=ax.transAxes)
# plot completeness vs Ncolors
ax = fig.add_subplot(222)
ax.plot(Ncolors, completeness[0], 'o-k', label='k=%i' % kvals[0])
ax.plot(Ncolors, completeness[1], '^--k', label='k=%i' % kvals[1])
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.set_ylabel('completeness')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
# plot contamination vs Ncolors
ax = fig.add_subplot(224)
ax.plot(Ncolors, contamination[0], 'o-k', label='k=%i' % kvals[0])
ax.plot(Ncolors, contamination[1], '^--k', label='k=%i' % kvals[1])
ax.legend(prop=dict(size=12),
loc='lower right',
bbox_to_anchor=(1.0, 0.79))
ax.xaxis.set_major_locator(plt.MultipleLocator(1))
ax.yaxis.set_major_locator(plt.MultipleLocator(0.2))
ax.xaxis.set_major_formatter(plt.FormatStrFormatter('%i'))
ax.set_xlabel('N colors')
ax.set_ylabel('contamination')
ax.set_xlim(0.5, 4.5)
ax.set_ylim(-0.1, 1.1)
ax.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# $K=1$ is clearly terrible--50% completeness using even all 4 attributes and 40% contamination.
#
# Where did $K=17$ come from?
#
# Well, regardless of whether this is the best algorithm or not, we can choose $K$ to minimize the classifcation error rate by using cross-validation. See below for how this was computed.
# + slideshow={"slide_type": "slide"}
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import colors
from sklearn.neighbors import KNeighborsClassifier
from astroML.datasets import fetch_rrlyrae_combined
from astroML.utils import completeness_contamination
# New
#from sklearn.cross_validation import cross_val_predict
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import accuracy_score
#----------------------------------------------------------------------
# get data and split into training & testing sets
X, y = fetch_rrlyrae_combined()
X = X[:, [1, 0, 2, 3]] # rearrange columns for better 1-color results
#----------------------------------------------------------------------
# perform Classification
scores = []
kvals = np.arange(1,20)
for k in kvals:
clf = KNeighborsClassifier(n_neighbors=k)
CVpredk = cross_val_predict(clf,X,y)
scores.append(accuracy_score(y, CVpredk))
# + slideshow={"slide_type": "slide"}
print("max score is for k={:d}".format(kvals[np.argmax(scores)])) # Complete
fig = plt.figure(figsize=(10, 5))
# Plot number of neighbors vs score
u = np.arange(len(scores))+1
plt.plot(u,scores)
# + [markdown] slideshow={"slide_type": "slide"}
# Below is an example of another way to implement cross validation using `cross_val_score` instead of `cross_val_predict`. Please see the [Scikit-Learn Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_predict.html#sklearn.model_selection.cross_val_predict) for important differences in the use of these two methods.
# + slideshow={"slide_type": "slide"}
#from sklearn.cross_validation import cross_val_score
from sklearn.model_selection import cross_val_score
scores2 = []
for k in kvals:
# Let's do a 2-fold cross-validation of the SVC estimator
clf2 = KNeighborsClassifier(n_neighbors=k)
CVpredk2 = cross_val_score(clf2,X,y,scoring='precision')
scores2.append(CVpredk2)
#scores2.append(cross_val_score(KNeighborsClassifier(n_neighbors=k), X, y, cv=2, scoring='precision'))
fig = plt.figure(figsize=(10, 5))
u = np.arange(len(scores2))+1
plt.plot(u,scores2)
# + [markdown] slideshow={"slide_type": "slide"}
# We can also use the [`metrics` module](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics) in sklearn to compute some statistics for us. Try inserting the code below into the appropriate place in our nearest neighbors classifier above.
# + slideshow={"slide_type": "slide"}
from sklearn import metrics
print k,nc
print("accuracy:", metrics.accuracy_score(y_test, y_pred))
print("precision:", metrics.precision_score(y_test, y_pred))
print("recall:", metrics.recall_score(y_test, y_pred))
| notebooks/Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import os
import copy
import torch
import random
import torchvision
import pandas as pd
import torch.nn as nn
import seaborn as sns
from PIL import Image
from glob import glob
import torch.optim as optim
import torchvision.models as m
import matplotlib.pyplot as plt
from torchvision import transforms, datasets
from sklearn.metrics import classification_report, confusion_matrix
print("PyTorch Version: ",torch.__version__)
print("Torchvision Version: ",torchvision.__version__)
random.seed = 2020
# -
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
# # Data Augmentation
from Augmentor import Pipeline
DIR = os.getcwd()
p = Pipeline(source_directory=f'{DIR}/dataset/val',output_directory=f'{DIR}/dataset/test')
#Pipeline
p.rotate(probability=0.5, max_left_rotation=15, max_right_rotation=15)
p.zoom(probability=0.7, min_factor=1.1, max_factor=1.2)
p.flip_left_right(probability=0.2)
p.sample(200)
# # Model
# +
# Number of classes in the dataset
num_classes = 2
# Batch size for training (change depending on how much memory you have)
batch_size = 8
# Number of epochs to train for
num_epochs = 15
# Flag for feature extracting. When False, we finetune the whole model,
# when True we only update the reshaped layer params
feature_extract = True
# -
def initialize_model(num_classes, feature_extract, use_pretrained=True):
model_ft = None
input_size = 0
model_ft = m.resnet50(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs, num_classes)
input_size = 224
return model_ft, input_size
def set_parameter_requires_grad(model, feature_extracting):
if feature_extracting:
for param in model.parameters():
param.requires_grad = False
def initialize_model(model_name, num_classes, feature_extract, use_pretrained=True):
# Initialize these variables which will be set in this if statement. Each of these
# variables is model specific.
model_ft = None
input_size = 0
if model_name == "resnet":
""" Resnet50
"""
model_ft = m.resnet50(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs, num_classes)
input_size = 224
elif model_name == "alexnet":
""" Alexnet
"""
model_ft = m.alexnet(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.classifier[6].in_features
model_ft.classifier[6] = nn.Linear(num_ftrs,num_classes)
input_size = 224
elif model_name == "vgg":
""" VGG11_bn
"""
model_ft = m.vgg11_bn(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.classifier[6].in_features
model_ft.classifier[6] = nn.Linear(num_ftrs,num_classes)
input_size = 224
elif model_name == "squeezenet":
""" Squeezenet
"""
model_ft = m.squeezenet1_0(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
model_ft.classifier[1] = nn.Conv2d(512, num_classes, kernel_size=(1,1), stride=(1,1))
model_ft.num_classes = num_classes
input_size = 224
elif model_name == "densenet":
""" Densenet
"""
model_ft = m.densenet121(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.classifier.in_features
model_ft.classifier = nn.Linear(num_ftrs, num_classes)
input_size = 224
elif model_name == "inception":
""" Inception v3
Be careful, expects (299,299) sized images and has auxiliary output
"""
model_ft = m.inception_v3(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
# Handle the auxilary net
num_ftrs = model_ft.AuxLogits.fc.in_features
model_ft.AuxLogits.fc = nn.Linear(num_ftrs, num_classes)
# Handle the primary net
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs,num_classes)
input_size = 299
else:
print("Invalid model name, exiting...")
exit()
return model_ft, input_size
# +
models = ['resnet', 'alexnet', 'vgg', 'squeezenet', 'densenet', 'inception']
model_name = models[0]
# Initialize the model for this run
model_ft, input_size = initialize_model(model_name, num_classes, feature_extract, use_pretrained=True)
# Print the model we just instantiated
print(model_ft)
# -
# # Training
# Data augmentation and normalization for training
# Just normalization for validation
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(input_size),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
'val': transforms.Compose([
transforms.Resize(input_size),
transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
'test': transforms.Compose([
transforms.Resize(input_size),
transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
}
def train_model(model, dataloaders, criterion, optimizer, num_epochs=25, is_inception=False):
since = time.time()
val_acc_history = []
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
# Get model outputs and calculate loss
# Special case for inception because in training it has an auxiliary output. In train
# mode we calculate the loss by summing the final output and the auxiliary output
# but in testing we only consider the final output.
outputs = model(inputs)
loss = criterion(outputs, labels)
_, preds = torch.max(outputs, 1)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
epoch_loss = running_loss / len(dataloaders[phase].dataset)
epoch_acc = running_corrects.double() / len(dataloaders[phase].dataset)
print('{} Loss: {:.4f} Acc: {:.4f}'.format(phase, epoch_loss, epoch_acc))
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
if phase == 'val':
val_acc_history.append(epoch_acc)
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
return model, val_acc_history
# +
data_dir = '/Users/capcarde/Desktop/COVID19/dataset'
# Create training and validation datasets
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x), data_transforms[x]) for x in ['train', 'val', 'test']}
# Create training and validation dataloaders
dataloaders_dict = {x: torch.utils.data.DataLoader(image_datasets[x], batch_size=batch_size, shuffle=True, num_workers=0) for x in ['train', 'val', 'test']}
# Detect if we have a GPU available
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# +
# Send the model to GPU
model_ft = model_ft.to(device)
# Gather the parameters to be optimized/updated in this run. If we are
# finetuning we will be updating all parameters. However, if we are
# doing feature extract method, we will only update the parameters
# that we have just initialized, i.e. the parameters with requires_grad
# is True.
params_to_update = model_ft.parameters()
print("Params to learn:")
if feature_extract:
params_to_update = []
for name,param in model_ft.named_parameters():
if param.requires_grad == True:
params_to_update.append(param)
print("\t",name)
else:
for name,param in model_ft.named_parameters():
if param.requires_grad == True:
print("\t",name)
# Observe that all parameters are being optimized
optimizer_ft = optim.SGD(params_to_update, lr=0.001, momentum=0.9)
# +
criterion = nn.CrossEntropyLoss()
# Train and evaluate
model_ft, hist = train_model(model_ft, dataloaders_dict, criterion, optimizer_ft, num_epochs=num_epochs, is_inception=(model_name=="inception"))
# +
#DIR = '/Users/capcarde/Desktop/COVID19'
#torch.save(model_ft.state_dict(), f'{DIR}/checkpoints/chk_resnet_50_epoch_{14}.pt')
# -
# # Evaluation
# Source of code: https://gist.github.com/shaypal5/94c53d765083101efc0240d776a23823
def print_confusion_matrix(confusion_matrix, class_names, figsize = (10,7), fontsize=14):
"""Prints a confusion matrix, as returned by sklearn.metrics.confusion_matrix, as a heatmap.
Arguments
---------
confusion_matrix: numpy.ndarray
The numpy.ndarray object returned from a call to sklearn.metrics.confusion_matrix.
Similarly constructed ndarrays can also be used.
class_names: list
An ordered list of class names, in the order they index the given confusion matrix.
figsize: tuple
A 2-long tuple, the first value determining the horizontal size of the ouputted figure,
the second determining the vertical size. Defaults to (10,7).
fontsize: int
Font size for axes labels. Defaults to 14.
Returns
-------
matplotlib.figure.Figure
The resulting confusion matrix figure
"""
df_cm = pd.DataFrame(
confusion_matrix, index=class_names, columns=class_names,
)
fig = plt.figure(figsize=figsize)
try:
heatmap = sns.heatmap(df_cm, annot=True, fmt="d")
except ValueError:
raise ValueError("Confusion matrix values must be integers.")
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=fontsize)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=45, ha='right', fontsize=fontsize)
plt.ylabel('True label')
plt.xlabel('Predicted label')
return fig
images, labels = zip(*dataloaders_dict['test'].dataset.imgs)
shape = (len(images), 3, 224,224)
# +
resize = data_transforms['test']
x_test = torch.zeros(shape)
target = torch.Tensor(labels).long()
for idx, image in enumerate(images):
x_test[idx,:,:,:] = resize(Image.open(image).convert('RGB'))
nb_samples = 10
nb_classes = 2
output = model_ft(x_test)
pred = torch.argmax(output, 1)
# -
# compute the confusion matrix and and use it to derive the raw
# accuracy, sensitivity, and specificity
cm = confusion_matrix(target, pred)
total = sum(sum(cm))
acc = (cm[0, 0] + cm[1, 1]) / total
sensitivity = cm[0, 0] / (cm[0, 0] + cm[0, 1])
specificity = cm[1, 1] / (cm[1, 0] + cm[1, 1])
# show the confusion matrix, accuracy, sensitivity, and specificity
print("acc: {:.4f}".format(acc))
print("sensitivity: {:.4f}".format(sensitivity))
print("specificity: {:.4f}".format(specificity))
# make predictions on the testing set
class_names = ['covid','normal']
print("[INFO] evaluating network...")
print(classification_report(target, pred,target_names=class_names))
print_confusion_matrix(cm, class_names, figsize = (10,7), fontsize=14)
| notebooks/Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="GhZOHbwJOvio"
# ##What are Convolutions?
#
# What are convolutions? In this lab you'll explore what they are and how they work. In later lessons, you'll see how to use them in your neural network.
# + [markdown] id="nidI4HtcVQ7i"
# Together with convolutions, you'll use something called 'Pooling', which compresses your image, further emphasising the features. You'll also see how pooling works in this lab.
# + [markdown] id="DdBFQswdO-kX"
# ##Limitations of the previous DNN
# In an earlier exercise you saw how to train an image classifier for fashion items using the Fashion MNIST dataset. This gave you a pretty accuract classifier, but there was an obvious constraint: the images were 28x28, grey scale and the item was centered in the image.
#
# For example here are a couple of the images in Fashion MNIST
# 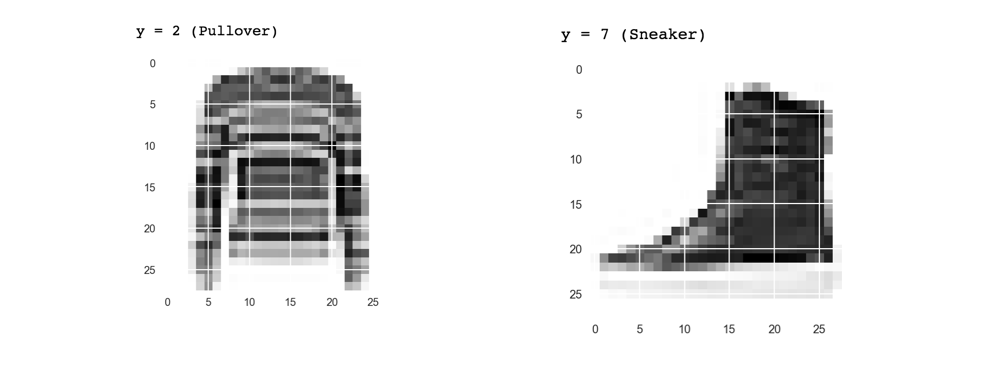
#
# The DNN that you created simply learned from the raw pixels what made up a sweater, and what made up a boot in this context. But consider how it might classify this image?
#
# 
#
# (Image is Public domain CC0 from Pixabay: https://pixabay.com/photos/boots-travel-railroad-tracks-181744/)
#
# While it's clear that there are boots in this image, the classifier would fail for a number of reasons. First, of course, it's not 28x28 greyscale, but more importantly, the classifier was trained on the raw pixels of a left-facing boot, and not the features that make up what a boot is.
#
# That's where Convolutions are very powerful. A convolution is a filter that passes over an image, processing it, and extracting features that show a commonolatity in the image. In this lab you'll see how they work, but processing an image to see if you can extract features from it!
#
#
#
# + [markdown] id="ds0NF5KFVmG2"
# Generating convolutions is very simple -- you simply scan every pixel in the image and then look at it's neighboring pixels. You multiply out the values of these pixels by the equivalent weights in a filter.
#
# So, for example, consider this:
#
# 
#
# In this case a 3x3 Convolution is specified.
#
# The current pixel value is 192, but you can calculate the new one by looking at the neighbor values, and multiplying them out by the values specified in the filter, and making the new pixel value the final amount.
#
#
# + [markdown] id="tJTHvE8Qe5nM"
# Let's explore how convolutions work by creating a basic convolution on a 2D Grey Scale image. First we can load the image by taking the 'ascent' image from scipy. It's a nice, built-in picture with lots of angles and lines.
# + [markdown] id="KTS2sc5nQSCJ"
# Let's start by importing some python libraries.
# + id="DZ5OXYiolCUi"
import cv2
import numpy as np
from scipy import misc
i = misc.ascent()
# + [markdown] id="SRIzxjWWfJjk"
# Next, we can use the pyplot library to draw the image so we know what it looks like.
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="R4p0cfWcfIvi" outputId="b9b14f1b-4afc-4462-cd05-e2664c9014b9"
import matplotlib.pyplot as plt
plt.grid(False)
plt.gray()
plt.axis('off')
plt.imshow(i)
plt.show()
# + [markdown] id="C1mhZ_ZTfPWH"
# We can see that this is an image of a stairwell. There are lots of features in here that we can play with seeing if we can isolate them -- for example there are strong vertical lines.
#
# The image is stored as a numpy array, so we can create the transformed image by just copying that array. Let's also get the dimensions of the image so we can loop over it later.
# + id="o5pxGq1SmJMD"
i_transformed = np.copy(i)
size_x = i_transformed.shape[0]
size_y = i_transformed.shape[1]
# + [markdown] id="Y7PwNkiXfddd"
# Now we can create a filter as a 3x3 array.
# + id="sN3imZannN5J"
# This filter detects edges nicely
# It creates a convolution that only passes through sharp edges and straight
# lines.
#Experiment with different values for fun effects.
#filter = [ [0, 1, 0], [1, -4, 1], [0, 1, 0]]
# A couple more filters to try for fun!
#filter = [ [-1, -2, -1], [0, 0, 0], [1, 2, 1]]
filter = [ [-1, 0, 1], [-2, 0, 2], [-1, 0, 1]]
# If all the digits in the filter don't add up to 0 or 1, you
# should probably do a weight to get it to do so
# so, for example, if your weights are 1,1,1 1,2,1 1,1,1
# They add up to 10, so you would set a weight of .1 if you want to normalize them
weight = 1
# + [markdown] id="JQmm_iBufmCz"
# Now let's create a convolution. We will iterate over the image, leaving a 1 pixel margin, and multiply out each of the neighbors of the current pixel by the value defined in the filter.
#
# i.e. the current pixel's neighbor above it and to the left will be multiplied by the top left item in the filter etc. etc. We'll then multiply the result by the weight, and then ensure the result is in the range 0-255
#
# Finally we'll load the new value into the transformed image.
# + id="299uU2jAr90h"
for x in range(1,size_x-1):
for y in range(1,size_y-1):
convolution = 0.0
convolution = convolution + (i[x - 1, y-1] * filter[0][0])
convolution = convolution + (i[x, y-1] * filter[1][0])
convolution = convolution + (i[x + 1, y-1] * filter[2][0])
convolution = convolution + (i[x-1, y] * filter[0][1])
convolution = convolution + (i[x, y] * filter[1][1])
convolution = convolution + (i[x+1, y] * filter[2][1])
convolution = convolution + (i[x-1, y+1] * filter[0][2])
convolution = convolution + (i[x, y+1] * filter[1][2])
convolution = convolution + (i[x+1, y+1] * filter[2][2])
convolution = convolution * weight
if(convolution<0):
convolution=0
if(convolution>255):
convolution=255
i_transformed[x, y] = convolution
# + [markdown] id="6XA--vgvgDEQ"
# Now we can plot the image to see the effect of the convolution!
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="7oPhUPNhuGWC" outputId="edea2357-55a6-4ff8-b34b-6b8d150dfc6c"
# Plot the image. Note the size of the axes -- they are 512 by 512
plt.gray()
plt.grid(False)
plt.imshow(i_transformed)
#plt.axis('off')
plt.show()
# + [markdown] id="Df7kw1m6XDwz"
# So, consider the following filter values, and their impact on the image.
#
# Using -1,0,1,-2,0,2,-1,0,1 gives us a very strong set of vertical lines:
#
# 
#
# Using -1, -2, -1, 0, 0, 0, 1, 2, 1 gives us horizontal lines:
#
# 
#
# Explore different values for yourself!
# + [markdown] id="xF0FPplsgHNh"
# ## Pooling
#
# As well as using convolutions, pooling helps us greatly in detecting features. The goal is to reduce the overall amount of information in an image, while maintaining the features that are detected as present.
#
# There are a number of different types of pooling, but for this lab we'll use one called MAX pooling.
#
# The idea here is to iterate over the image, and look at the pixel and it's immediate neighbors to the right, beneath, and right-beneath. Take the largest (hence the name MAX pooling) of them and load it into the new image. Thus the new image will be 1/4 the size of the old -- with the dimensions on X and Y being halved by this process. You'll see that the features get maintained despite this compression!
#
# 
#
# This code will show (4, 4) pooling. Run it to see the output, and you'll see that while the image is 1/4 the size of the original in both length and width, the extracted features are maintained!
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="kDHjf-ehaBqm" outputId="bb61962e-0e00-43fd-aee5-87f25ca55f59"
new_x = int(size_x/4)
new_y = int(size_y/4)
newImage = np.zeros((new_x, new_y))
for x in range(0, size_x, 4):
for y in range(0, size_y, 4):
pixels = []
pixels.append(i_transformed[x, y])
pixels.append(i_transformed[x+1, y])
pixels.append(i_transformed[x+2, y])
pixels.append(i_transformed[x+3, y])
pixels.append(i_transformed[x, y+1])
pixels.append(i_transformed[x+1, y+1])
pixels.append(i_transformed[x+2, y+1])
pixels.append(i_transformed[x+3, y+1])
pixels.append(i_transformed[x, y+2])
pixels.append(i_transformed[x+1, y+2])
pixels.append(i_transformed[x+2, y+2])
pixels.append(i_transformed[x+3, y+2])
pixels.append(i_transformed[x, y+3])
pixels.append(i_transformed[x+1, y+3])
pixels.append(i_transformed[x+2, y+3])
pixels.append(i_transformed[x+3, y+3])
pixels.sort(reverse=True)
newImage[int(x/4),int(y/4)] = pixels[0]
# Plot the image. Note the size of the axes -- now 128 pixels instead of 512
plt.gray()
plt.grid(False)
plt.imshow(newImage)
#plt.axis('off')
plt.show()
# + [markdown] id="jZWdU6dVYQm-"
# In the next lab you'll see how to add convolutions to your Fashion MNIST neural network to make it more efficient -- because it will classify based on features, and not on raw pixels.
| ML/Exploring_Convolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Ensemble Learning
#
# ## Initial Imports
# -
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
from pathlib import Path
from collections import Counter
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import balanced_accuracy_score
from sklearn.metrics import confusion_matrix
from imblearn.metrics import classification_report_imbalanced
from imblearn.ensemble import BalancedRandomForestClassifier
from imblearn.ensemble import EasyEnsembleClassifier
from sklearn.preprocessing import LabelEncoder
# ## Read the CSV and Inspect the Data
# +
# Load the data
file_path = Path('Resources/LoanStats_2019Q1.csv')
df = pd.read_csv(file_path)
# Preview the data
df.head()
# -
# Check dataframe for any null or non numeric feature values pre clean
df.info()
# Identify any non numeric feature value columns that need to be treated
# Examine this feature data to determin the appropriate mehtods for cleaning
df.select_dtypes(include='object')
# Return columns from dataframe for any null or non numeric feature values pre clean
df.select_dtypes(include='object').info()
# ### Data Inspection Assessment
#
# It can be seen when examining this data that there are 9 feature data columns (other than our targe column **loan_status**) that contain objects that need to be treated.
#
# At this stage it is noted other than incompatible Dtypes for modelling there are no null rows, values or other basic data integrity issues
# ## Perform Initial Data Cleaning
#
# The feature data will be cleaned and treated using date splitting and conversion along with integer encoding. Before progressing to data set splitting and scaling
# +
# Split and treat the no numeric data in the issue_d and next_pymnt_d columns
# Split year from Month short name into two new columns for issue_d column
df[['issue_month','issue_year']] = df['issue_d'].str.split('-', expand=True)
# Split year from Month short name into two new columns for next_pymnt_d column
df[['next_pymnt_month','next_pymnt_year']] = df['next_pymnt_d'].str.split('-', expand=True)
# Create Months dictionary using month shortname as seen in the data
months_num = {
"Jan": 1,
"Feb": 2,
"Mar": 3,
"Apr": 4,
"May": 5,
"Jun": 6,
"Jul": 7,
"Aug": 8,
"Sep": 9,
"Oct": 10,
"Nov": 11,
"Dec": 12,
}
# Months' names encoded using the dictionary values
df["issue_month_num"] = df["issue_month"].apply(lambda x: months_num[x])
df["next_pymnt_month_num"] = df["next_pymnt_month"].apply(lambda x: months_num[x])
# Convert issue_year and next_pymnt_year column to int data type
df = df.astype({"issue_year": int,"next_pymnt_year": int})
# Drop the date columns as this data has been treated and numerically represented in new columns
df.drop(["issue_d", "issue_month","next_pymnt_d", "next_pymnt_month"], axis=1, inplace=True)
# Create list of all remaining columns that need to be treated by integer encoder
non_integer_columns = ["home_ownership", "verification_status", "pymnt_plan", "initial_list_status", "application_type", "hardship_flag", "debt_settlement_flag"]
#Create loop to integer encode the remaining data
for column in non_integer_columns:
# Creating an instance of label encoder for integer encoding the columns data
label_encoder = LabelEncoder()
# Fitting the label encoder
label_encoder.fit(df[column])
# Encode the data as an integer into new label encoded column
encoded_column_name = column + "_le"
df[encoded_column_name] = label_encoder.transform(df[column])
# Drop the source column
df.drop([column], axis=1, inplace=True)
# Inspect the cleaned data
df.head()
# -
# Return columns from dataframe for any null or non numeric feature values pre clean
df.select_dtypes(include='object').info()
# Check dataframe for any null or non numeric feature values post clean
df.info()
# ### Initial Data Cleaning Assessment
#
# The feature data is ready to split into testing and training sets, scaled and then passed to modelling
# ## Split the Data into Training and Testing
# +
# Create our features
X = df.copy()
# Dropping homeowner and loan_statis columns
X.drop(["loan_status"], axis=1, inplace=True)
# Create our target
y = df["loan_status"]
# -
X.describe()
# Check the balance of our target values
y.value_counts()
# Split the X and y into X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test= train_test_split(X,
y,
random_state=1,
stratify=y)
# Examine X and y testing and training set shapes
#X_train.shape
#X_test.shape
#y_train.shape
y_test.shape
# ## Data Pre-Processing
#
# Scale the training and testing data using the `StandardScaler` from `sklearn`. Remember that when scaling the data, you only scale the features data (`X_train` and `X_testing`).
# Create the StandardScaler instance
scaler = StandardScaler()
# Fit the Standard Scaler with the training data
# When fitting scaling functions, only train on the training dataset
X_scaler = scaler.fit(X_train)
# Scale the training and testing data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# ## Ensemble Learners
#
# In this section, you will compare two ensemble algorithms to determine which algorithm results in the best performance. You will train a Balanced Random Forest Classifier and an Easy Ensemble classifier . For each algorithm, be sure to complete the folliowing steps:
#
# 1. Train the model using the training data.
# 2. Calculate the balanced accuracy score from sklearn.metrics.
# 3. Display the confusion matrix from sklearn.metrics.
# 4. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
# 5. For the Balanced Random Forest Classifier only, print the feature importance sorted in descending order (most important feature to least important) along with the feature score
#
# Note: Use a random state of 1 for each algorithm to ensure consistency between tests
# ### Balanced Random Forest Classifier
# Resample the training data with the BalancedRandomForestClassifier
brf_model = BalancedRandomForestClassifier(n_estimators=100, random_state=1)
brf_model.fit(X_train_scaled, y_train)
y_pred = brf_model.predict(X_test_scaled)
brf_model
# Calculated the balanced accuracy score
balanced_accuracy_score(y_test, y_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred, digits=16))
# + tags=[]
# List the features sorted in descending order by feature importance
importances = brf_model.feature_importances_
sorted(zip(brf_model.feature_importances_, X.columns), reverse=True)
# + [markdown] tags=[]
# ### Easy Ensemble Classifier
# -
# Train the Classifier
ee_model = EasyEnsembleClassifier(n_estimators=100, random_state=1)
ee_model.fit(X_train_scaled, y_train)
y_pred = ee_model.predict(X_test_scaled)
ee_model
# Calculated the balanced accuracy score
balanced_accuracy_score(y_test, y_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred, digits=16))
# ### Final Questions
#
# 1. Which model had the best balanced accuracy score?
#
# - 0.7739584466441858 - Balanced Random Forest Classifier
# - **0.9324304724609305 - Easy Ensemble Classifier**
#
# **ANSWER:** Examining the figures above it can bee seen that the **Easy Ensemble Classifier** model had the best balanced accuracy score.
#
# 2. Which model had the best recall score?
#
# - 0.8915431560592851 - Balanced Random Forest Classifier
# - **0.9451903516419645 - Easy Ensemble Classifier**
#
# **ANSWER:** Examining the figures above it can bee seen that the **Easy Ensemble Classifier** model had the best recall score.
#
# 3. Which model had the best geometric mean score?
#
# - 0.7647885691688799 - Balanced Random Forest Classifier
# - **0.9323413686091855 - Easy Ensemble Classifier**
#
# **ANSWER:** Examining the figures above it can bee seen that the **Easy Ensemble Classifier** model had the best geometric mean score.
#
# 4. What are the top three features?
#
# **ANSWER:** The following lists the top three features
# - 0.0756092393069983, 'total_rec_prncp'
# - 0.06959573158422656, 'total_rec_int'
# - 0.06327652424873491, 'total_pymnt_inv'
| Starter_Code/credit_risk_ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# + language="bash"
# mkdir lab4
# cd lab4
# git clone https://github.com/sophiajwchoi/Lab4-Adding-Discovery-to-Chatbot.git
# + language="bash"
# cd /resources/lab4/Lab4-Adding-Discovery-to-Chatbot
# export npm_config_loglevel=silent
# conda config --set notify_outdated_conda false
# conda install nodejs -y
# rm -f ~/.npmrc
# npm install
# + language="bash"
# #ibmcloud resource groups
# #ibmcloud target -g RESOURCE_GROUP
# ibmcloud account orgs
# #ibmcloud login --apikey <KEY>
# + language="bash"
# ibmcloud target --cf-api 'https://api.us-south.cf.cloud.ibm.com' -r us-south -o <EMAIL>
# ibmcloud account space-create 'lab4'
# + language="bash"
# cd /resources/lab4/Lab4-Adding-Discovery-to-Chatbot
# ibmcloud target -s lab4
# ibmcloud plugin install cloud-functions -f
# export npm_config_loglevel=silent
# npm install -g serverless@1.51.0
# ibmcloud fn --apihost us-south.functions.cloud.ibm.com
# ibmcloud fn list --apihost us-south.functions.cloud.ibm.com
# serverless deploy
# -
# !ibmcloud fn list
| Gus-Primero.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# # Data Visualization -Final Assignment
#
# ## 1. Create dataframe for Topic survey assignment
#
# A survey was conducted to gauge an audience interest in different data science topics, namely:
#
# Big Data (Spark / Hadoop)
# Data Analysis / Statistics
# Data Journalism
# Data Visualization
# Deep Learning
# Machine Learning
# The participants had three options for each topic: Very Interested, Somewhat interested, and Not interested. 2,233 respondents completed the survey.
#
# +
import pandas as pd
import numpy as np
# Read csv and assign it to dataframe as variable df
csv_path = "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork/labs/FinalModule_Coursera/data/Topic_Survey_Assignment.csv"
df = pd.read_csv(csv_path)
df
# -
# ## 2. Create bar chart for survey
#
# Use the artist layer of Matplotlib to replicate the bar chart below to visualize the percentage of the respondents' interest in the different data science topics surveyed.
#
#
# To create this bar chart, you can follow the following steps:
#
# Sort the dataframe in descending order of Very interested.
# Convert the numbers into percentages of the total number of respondents. Recall that 2,233 respondents completed the survey. Round percentages to 2 decimal places.
# As for the chart:
# use a figure size of (20, 8),
# bar width of 0.8,
# use color #5cb85c for the Very interested bars, color #5bc0de for the Somewhat interested bars, and color #d9534f for the Not interested bars,
# use font size 14 for the bar labels, percentages, and legend,
# use font size 16 for the title, and,
# display the percentages above the bars as shown above, and remove the left, top, and right borders.
#
# +
# generate the plots within the browser
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.style.use('ggplot') # ggplot style
# check for latest version of Matplotlib
print ('Matplotlib version: ', mpl.__version__) # >= 2.0.0
# +
#Sort the dataframe in descending order of Very interested
df.sort_values(['Very interested'], ascending=False, axis=0, inplace=True)
df.head()
# +
#Convert the numbers into percentages of the total number of respondents, 2 decimal places
pd.options.display.float_format = '{:.2f}%'.format
df_perc = df.div(df.sum(axis=1), axis=0).multiply(100)
df_perc
# +
#Chart:1.use a figure size of (20, 8)\
# 2.bar width of 0.8\
# 3.use color #5cb85c for the Very interested bars,#color #5bc0de for the Somewhat interested bars, and color #d9534f for the Not interested bars\
# 4.use font size 14 for the bar labels, percentages, and legend\
# 5.use font size 16 for the title\
# 6.display the percentages above the bars\
# 7.remove the left, top, and right borders
ax = df_perc.plot(kind='bar', figsize=(20, 8), alpha = 1.0, width=0.8, color=['#5cb85c','#5bc0de','#d9534f'], fontsize=14)
ax.set_facecolor('xkcd:white')
ax.legend(fontsize=14,facecolor='white')
ax.set_xticklabels(ax.get_xticklabels(), fontsize=14)
ax.set_title("Percentage of Respondents'Interest in Data Science Areas", fontsize=16, loc='center') # add title
ax.grid(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_color('black')
ax.xaxis.set_ticks_position('bottom')
ax.get_yaxis().set_ticks([])
for bar in ax.patches:
ax.annotate(format(bar.get_height(), '.2f') +'%',
(bar.get_x() + bar.get_width() / 2,
bar.get_height()), ha='center', va='center',
size=14, xytext=(0, 10),
textcoords='offset points')
plt.show()
# -
| Final assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 目标检测和边界框
# :label:`sec_bbox`
#
# 在前面的章节(例如 :numref:`sec_alexnet`— :numref:`sec_googlenet`)中,我们介绍了各种图像分类模型。
# 在图像分类任务中,我们假设图像中只有一个主要物体对象,我们只关注如何识别其类别。
# 然而,很多时候图像里有多个我们感兴趣的目标,我们不仅想知道它们的类别,还想得到它们在图像中的具体位置。
# 在计算机视觉里,我们将这类任务称为*目标检测*(object detection)或*目标识别*(object recognition)。
#
# 目标检测在多个领域中被广泛使用。
# 例如,在无人驾驶里,我们需要通过识别拍摄到的视频图像里的车辆、行人、道路和障碍物的位置来规划行进线路。
# 机器人也常通过该任务来检测感兴趣的目标。安防领域则需要检测异常目标,如歹徒或者炸弹。
#
# 在接下来的几节中,我们将介绍几种用于目标检测的深度学习方法。
# 我们将首先介绍目标的*位置*。
#
# + origin_pos=3 tab=["tensorflow"]
# %matplotlib inline
import tensorflow as tf
from d2l import tensorflow as d2l
# + [markdown] origin_pos=4
# 下面加载本节将使用的示例图像。可以看到图像左边是一只狗,右边是一只猫。
# 它们是这张图像里的两个主要目标。
#
# + origin_pos=6 tab=["tensorflow"]
d2l.set_figsize()
img = d2l.plt.imread('../img/catdog.jpg')
d2l.plt.imshow(img);
# + [markdown] origin_pos=7
# ## 边界框
#
# 在目标检测中,我们通常使用*边界框*(bounding box)来描述对象的空间位置。
# 边界框是矩形的,由矩形左上角的以及右下角的$x$和$y$坐标决定。
# 另一种常用的边界框表示方法是边界框中心的$(x, y)$轴坐标以及框的宽度和高度。
#
# 在这里,我们[**定义在这两种表示法之间进行转换的函数**]:`box_corner_to_center`从两角表示法转换为中心宽度表示法,而`box_center_to_corner`反之亦然。
# 输入参数`boxes`可以是长度为4的张量,也可以是形状为($n$,4)的二维张量,其中$n$是边界框的数量。
#
# + origin_pos=8 tab=["tensorflow"]
#@save
def box_corner_to_center(boxes):
"""从(左上,右下)转换到(中间,宽度,高度)"""
x1, y1, x2, y2 = boxes[:, 0], boxes[:, 1], boxes[:, 2], boxes[:, 3]
cx = (x1 + x2) / 2
cy = (y1 + y2) / 2
w = x2 - x1
h = y2 - y1
boxes = tf.stack((cx, cy, w, h), axis=-1)
return boxes
#@save
def box_center_to_corner(boxes):
"""从(中间,宽度,高度)转换到(左上,右下)"""
cx, cy, w, h = boxes[:, 0], boxes[:, 1], boxes[:, 2], boxes[:, 3]
x1 = cx - 0.5 * w
y1 = cy - 0.5 * h
x2 = cx + 0.5 * w
y2 = cy + 0.5 * h
boxes = tf.stack((x1, y1, x2, y2), axis=-1)
return boxes
# + [markdown] origin_pos=9
# 我们将根据坐标信息[**定义图像中狗和猫的边界框**]。
# 图像中坐标的原点是图像的左上角,向右的方向为$x$轴的正方向,向下的方向为$y$轴的正方向。
#
# + origin_pos=10 tab=["tensorflow"]
# bbox是边界框的英文缩写
dog_bbox, cat_bbox = [60.0, 45.0, 378.0, 516.0], [400.0, 112.0, 655.0, 493.0]
# + [markdown] origin_pos=11
# 我们可以通过转换两次来验证边界框转换函数的正确性。
#
# + origin_pos=12 tab=["tensorflow"]
boxes = tf.constant((dog_bbox, cat_bbox))
box_center_to_corner(box_corner_to_center(boxes)) == boxes
# + [markdown] origin_pos=13
# 我们可以[**将边界框在图中画出**],以检查其是否准确。
# 画之前,我们定义一个辅助函数`bbox_to_rect`。
# 它将边界框表示成`matplotlib`的边界框格式。
#
# + origin_pos=14 tab=["tensorflow"]
#@save
def bbox_to_rect(bbox, color):
# 将边界框(左上x,左上y,右下x,右下y)格式转换成matplotlib格式:
# ((左上x,左上y),宽,高)
return d2l.plt.Rectangle(
xy=(bbox[0], bbox[1]), width=bbox[2]-bbox[0], height=bbox[3]-bbox[1],
fill=False, edgecolor=color, linewidth=2)
# + [markdown] origin_pos=15
# 在图像上添加边界框之后,我们可以看到两个物体的主要轮廓基本上在两个框内。
#
# + origin_pos=16 tab=["tensorflow"]
fig = d2l.plt.imshow(img)
fig.axes.add_patch(bbox_to_rect(dog_bbox, 'blue'))
fig.axes.add_patch(bbox_to_rect(cat_bbox, 'red'));
# + [markdown] origin_pos=17
# ## 小结
#
# * 目标检测不仅可以识别图像中所有感兴趣的物体,还能识别它们的位置,该位置通常由矩形边界框表示。
# * 我们可以在两种常用的边界框表示(中间,宽度,高度)和(左上,右下)坐标之间进行转换。
#
# ## 练习
#
# 1. 找到另一张图像,然后尝试标记包含该对象的边界框。比较标注边界框和标注类别哪个需要更长的时间?
# 1. 为什么`box_corner_to_center`和`box_center_to_corner`的输入参数的最内层维度总是4?
#
| tensorflow/chapter_computer-vision/bounding-box.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
from '../calibrate_sense_hat' import SenseHat
from PIL import Image # pillow
import numpy as np
import time
file_path = "calibrate_sense_hat_text.png"
text_file = "calibrate_sense_hat_text.txt"
img = Image.open(file_path).convert('RGB')
text_pixels = list(map(list, img.getdata()))
text_dict = {}
with open(text_file, 'r') as f:
loaded_text = f.read()
for index, s in enumerate(loaded_text):
start = index * 32
end = start + 32
char = text_pixels[start:end]
text_dict[s] = char
text_dict
sense = SenseHat()
pixel_list = sense._get_char_pixels("s")
# +
pix_map0 = np.array([
[0, 1, 2, 3, 4, 5, 6, 7],
[8, 9, 10, 11, 12, 13, 14, 15],
[16, 17, 18, 19, 20, 21, 22, 23],
[24, 25, 26, 27, 28, 29, 30, 31],
[32, 33, 34, 35, 36, 37, 38, 39],
[40, 41, 42, 43, 44, 45, 46, 47],
[48, 49, 50, 51, 52, 53, 54, 55],
[56, 57, 58, 59, 60, 61, 62, 63]
], int)
pix_map90 = np.rot90(pix_map0)
pix_map180 = np.rot90(pix_map90)
pix_map270 = np.rot90(pix_map180)
pix_map = {
0: pix_map0,
90: pix_map90,
180: pix_map180,
270: pix_map270
}
pix_map
# -
def set_pixel(x, y, *args):
pixel = args[0]
print pixel
for index, pix in enumerate(pixel_list):
print index, pix
def get_char_pixels(s):
"""
Internal. Safeguards the character indexed dictionary for the
show_message function below
"""
if len(s) == 1 and s in text_dict.keys():
return list(text_dict[s])
else:
return list(text_dict['?'])
def set_pixels(pixel_list):
map = pix_map[90]
for index, pix in enumerate(pixel_list):
print index, map[index // 8][index % 8] , pix
# + active=""
# map = pix_map[90]
# index = 0
#
# for index in range[0:10]:
# print index // 8, index % 8
# print map[index // 8][index % 8]
#
#
# +
def show_message(text_string,scroll_speed=.1, text_colour=[255, 255, 255], back_colour=[0, 0, 0]):
scroll_pixels = []
string_padding = [[0, 0, 0]] * 64
scroll_pixels.extend(string_padding)
for s in text_string:
scroll_pixels.extend(get_char_pixels(s))
scroll_pixels.extend(string_padding)
scroll_length = len(scroll_pixels) // 8
for i in range(scroll_length - 8):
start = i * 8
end = start + 64
sense.set_pixels(scroll_pixels[start:end])
time.sleep(scroll_speed)
# for s in text_string:
# pixel_list = get_char_pixels(s)
# for index, pix in enumerate(pixel_list):
# sense.set_pixel(index % 8, index // 8, pix)
# print index % 8, index // 8, pix
# -
pix_map[90]
show_message("hello")
text_string = "hello"
for s in text_string:
print s
| notebook/testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
import tensorwatch as tw
train = tw.WatcherClient(port=0)
test = tw.WatcherClient(port=1)
loss_stream = train.create_stream(expr='lambda d:(d.x, d.metrics.batch_loss)', event_name='batch')
loss_plot = tw.Visualizer(loss_stream, vis_type='line',
xtitle='Epoch', ytitle='Train Loss')
loss_plot.show()
acc_stream = train.create_stream(expr='lambda d:(d.x, d.metrics.batch_accuracy)', event_name='batch')
acc_plot = tw.Visualizer(acc_stream, vis_type='line', host=loss_plot,
xtitle='Epoch', ytitle='Train Accuracy', yrange=(0,))
acc_plot.show()
test_loss_stream = test.create_stream(expr='lambda d:(d.x, d.metrics.batch_loss)', event_name='batch')
test_loss_plot = tw.Visualizer(test_loss_stream, vis_type='line', host=loss_plot,
xtitle='Epoch', ytitle='Test Loss', yrange=(0,))
test_loss_plot.show()
test_acc_stream = test.create_stream(expr='lambda d:(d.x, d.metrics.batch_accuracy)', event_name='batch')
test_acc_plot = tw.Visualizer(test_acc_stream, vis_type='line', host=loss_plot,
xtitle='Epoch', ytitle='Test Accuracy', yrange=(0,))
test_acc_plot.show()
grads_stream = train.create_stream(expr='lambda d:agg_params(d.model, lambda p: p.grad.abs().mean().item())',
event_name='batch')
grads_plot = tw.Visualizer(grads_stream, vis_type='line', title="Weight Gradients", clear_after_each=True,
xtitle='Layer', ytitle='Abs Mean Gradient', history_len=20)
grads_plot.show()
rand_pred = train.create_stream(expr="top(l, out_xform=pyt_img_class_out_xform, order='rnd')", event_name='batch', throttle=2)
rand_pred_plot = tw.Visualizer(rand_pred, vis_type='image', title="Random Predictions")
rand_pred_plot.show()
worse_pred = train.create_stream(expr="top(l, out_xform=pyt_img_class_out_xform, order='asc')", event_name='batch', throttle=2)
worse_pred_plot = tw.Visualizer(worse_pred, vis_type='image', title="Worst Predictions")
worse_pred_plot.show()
| notebooks/demo/mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
stu_details = {'stu_id':[1,2,3,4,5],
'stu_name':['A','B','C','D','E'],
'Region':['N','S','W','E','S']}
stu_marks = {'st_id':[1,2,3,4,6],
'sub':['Py','Tab','R','Py','Py'],
'Marks':[35,32,34,37,38]}
# -
stu_details
stu_marks
import pandas as pd
pd.__version__
stu_details = pd.DataFrame(stu_details)
type(stu_details)
stu_marks = pd.DataFrame(stu_marks)
stu_marks
stu_details
stu_marks
pd.merge(stu_details,stu_marks, left_on='stu_id',right_on='st_id', how='inner')
pd.merge(stu_details,stu_marks, on='st_id', how='left')
pd.merge(stu_details,stu_marks, on='st_id', how='right')
pd.merge(stu_details,stu_marks, on='st_id', how='outer')
# +
stu_details_Nag = {'st_id':[1,2,3,4,5],
'stu_name':['A','B','C','D','E'],
'sub':['Py','Tab','Py','Tab','PY']}
stu_details_US = {'st_id':[6,7,8,9,10],
'stu_name':['T','Y','U','I','O'],
'sub':['Tab','Tab','Tab','SQL','PY']}
# -
stu_details_Nag=pd.DataFrame(stu_details_Nag)
stu_details_US=pd.DataFrame(stu_details_US)
stu_details_Nag
stu_details_US
student_details = pd.concat([stu_details_Nag, stu_details_US],axis=0)
student_details.reset_index()
student_details = pd.concat([stu_details_Nag, stu_details_US],axis=1)
student_details
| Pandas/Pandas - Part 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
import intake
cat = intake.open_catalog(r"../catalogs/*.yml")
pcts = cat.pcts2.read()
pcts_cases = pcts["CASE_NBR"]
pcts_cases_split = pcts_cases.str.split('-',expand=True)
# #### note: .fillna() does not change the groupby total rows
pcts_cases_split.drop(columns=pcts_cases_split.columns[4:]).groupby([0,1]).first()
pcts_cases_split.groupby([0,1]).first()
# ### Type the case string parts
def string_type(string):
if string: # ignore None type
if string.isdigit():
return "numeric" + str(len(string))
elif string.isalpha():
return "alpha"
elif string.isalnum():
return "alphanumeric"
pcts_cases_typed = pcts_cases_split.copy()
for c in pcts_cases_split.columns[1:]:
pcts_cases_typed[c] = pcts_cases_typed[c].map(string_type)
pcts_cases_typed.rename(columns={0:'prefix',1:'2',2:'3',3:'4'}, inplace=True)
# ### Groupby to see different combinations; unfortunately this only shows suffix types that appear in first suffix position
pcts_cases_typed.groupby(['prefix','2','3','4'],dropna=False).first()
# ### Alternative to see suffix types in various suffix positions.
group_suffixes = pcts_cases_typed.set_index(
['prefix','2','3']
).stack(
dropna=False
).reset_index(
).groupby(
['prefix','2','3',0], dropna=False
).first(
).reset_index()#.to_csv('pcts_cases_typed_stacked.csv')
group_suffixes.rename(columns={0:'4','level_3':'suffix_position'}, inplace=True)
for c in group_suffixes.columns[1:4]:
print(f'Position {c} contains alpha characters for the following: \n\
{set(group_suffixes[group_suffixes[c].str.contains("alpha").fillna(False)]["prefix"].to_list())}')
# ### Check ZA that showed up with alphanumeric year in second position. Appears to be a year still, but using a convention that must be inquired about with systems or the planners.
pcts.iloc[pcts_cases_split[(
pcts_cases_split[0] == 'ZA') & (
~pcts_cases_split[1].str.isdigit()
)].index]
# ### Note that there are some auto-generated cases with suffixes that have a (n) or (N) attached. The regular expressions won't catch those. Based on the 2010-2019 backup it appears it is only 3 case records associated with DRB suffix. More info [here](https://planning.lacity.org/dcpapi/general/prefixsuffix/active/pdf/).
pcts[pcts.CASE_NBR.str.contains('\(')]
| notebooks/explore-case-strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Long Short-Term Memory (LSTM)
# :label:`sec_lstm`
#
# The challenge to address long-term information preservation and short-term input
# skipping in latent variable models has existed for a long time. One of the
# earliest approaches to address this was the
# long short-term memory (LSTM) :cite:`Hochreiter.Schmidhuber.1997`. It shares many of the properties of the
# GRU.
# Interestingly, LSTMs have a slightly more complex
# design than GRUs but predates GRUs by almost two decades.
#
#
#
# ## Gated Memory Cell
#
# Arguably LSTM's design is inspired
# by logic gates of a computer.
# LSTM introduces a *memory cell* (or *cell* for short)
# that has the same shape as the hidden state
# (some literatures consider the memory cell
# as a special type of the hidden state),
# engineered to record additional information.
# To control the memory cell
# we need a number of gates.
# One gate is needed to read out the entries from the
# cell.
# We will refer to this as the
# *output gate*.
# A second gate is needed to decide when to read data into the
# cell.
# We refer to this as the *input gate*.
# Last, we need a mechanism to reset
# the content of the cell, governed by a *forget gate*.
# The motivation for such a
# design is the same as that of GRUs,
# namely to be able to decide when to remember and
# when to ignore inputs in the hidden state via a dedicated mechanism. Let us see
# how this works in practice.
#
#
# ### Input Gate, Forget Gate, and Output Gate
#
# Just like in GRUs,
# the data feeding into the LSTM gates are
# the input at the current time step and
# the hidden state of the previous time step,
# as illustrated in :numref:`lstm_0`.
# They are processed by
# three fully-connected layers with a sigmoid activation function to compute the values of
# the input, forget. and output gates.
# As a result, values of the three gates
# are in the range of $(0, 1)$.
#
# 
# :label:`lstm_0`
#
# Mathematically,
# suppose that there are $h$ hidden units, the batch size is $n$, and the number of inputs is $d$.
# Thus, the input is $\mathbf{X}_t \in \mathbb{R}^{n \times d}$ and the hidden state of the previous time step is $\mathbf{H}_{t-1} \in \mathbb{R}^{n \times h}$. Correspondingly, the gates at time step $t$
# are defined as follows: the input gate is $\mathbf{I}_t \in \mathbb{R}^{n \times h}$, the forget gate is $\mathbf{F}_t \in \mathbb{R}^{n \times h}$, and the output gate is $\mathbf{O}_t \in \mathbb{R}^{n \times h}$. They are calculated as follows:
#
# $$
# \begin{aligned}
# \mathbf{I}_t &= \sigma(\mathbf{X}_t \mathbf{W}_{xi} + \mathbf{H}_{t-1} \mathbf{W}_{hi} + \mathbf{b}_i),\\
# \mathbf{F}_t &= \sigma(\mathbf{X}_t \mathbf{W}_{xf} + \mathbf{H}_{t-1} \mathbf{W}_{hf} + \mathbf{b}_f),\\
# \mathbf{O}_t &= \sigma(\mathbf{X}_t \mathbf{W}_{xo} + \mathbf{H}_{t-1} \mathbf{W}_{ho} + \mathbf{b}_o),
# \end{aligned}
# $$
#
# where $\mathbf{W}_{xi}, \mathbf{W}_{xf}, \mathbf{W}_{xo} \in \mathbb{R}^{d \times h}$ and $\mathbf{W}_{hi}, \mathbf{W}_{hf}, \mathbf{W}_{ho} \in \mathbb{R}^{h \times h}$ are weight parameters and $\mathbf{b}_i, \mathbf{b}_f, \mathbf{b}_o \in \mathbb{R}^{1 \times h}$ are bias parameters.
#
# ### Candidate Memory Cell
#
# Next we design the memory cell. Since we have not specified the action of the various gates yet, we first introduce the *candidate* memory cell $\tilde{\mathbf{C}}_t \in \mathbb{R}^{n \times h}$. Its computation is similar to that of the three gates described above, but using a $\tanh$ function with a value range for $(-1, 1)$ as the activation function. This leads to the following equation at time step $t$:
#
# $$\tilde{\mathbf{C}}_t = \text{tanh}(\mathbf{X}_t \mathbf{W}_{xc} + \mathbf{H}_{t-1} \mathbf{W}_{hc} + \mathbf{b}_c),$$
#
# where $\mathbf{W}_{xc} \in \mathbb{R}^{d \times h}$ and $\mathbf{W}_{hc} \in \mathbb{R}^{h \times h}$ are weight parameters and $\mathbf{b}_c \in \mathbb{R}^{1 \times h}$ is a bias parameter.
#
# A quick illustration of the candidate memory cell is shown in :numref:`lstm_1`.
#
# 
# :label:`lstm_1`
#
# ### Memory Cell
#
# In GRUs, we have a mechanism to govern input and forgetting (or skipping).
# Similarly,
# in LSTMs we have two dedicated gates for such purposes: the input gate $\mathbf{I}_t$ governs how much we take new data into account via $\tilde{\mathbf{C}}_t$ and the forget gate $\mathbf{F}_t$ addresses how much of the old memory cell content $\mathbf{C}_{t-1} \in \mathbb{R}^{n \times h}$ we retain. Using the same pointwise multiplication trick as before, we arrive at the following update equation:
#
# $$\mathbf{C}_t = \mathbf{F}_t \odot \mathbf{C}_{t-1} + \mathbf{I}_t \odot \tilde{\mathbf{C}}_t.$$
#
# If the forget gate is always approximately 1 and the input gate is always approximately 0, the past memory cells $\mathbf{C}_{t-1}$ will be saved over time and passed to the current time step.
# This design is introduced to alleviate the vanishing gradient problem and to better capture
# long range dependencies within sequences.
#
# We thus arrive at the flow diagram in :numref:`lstm_2`.
#
# 
#
# :label:`lstm_2`
#
#
# ### Hidden State
#
# Last, we need to define how to compute the hidden state $\mathbf{H}_t \in \mathbb{R}^{n \times h}$. This is where the output gate comes into play. In LSTM it is simply a gated version of the $\tanh$ of the memory cell.
# This ensures that the values of $\mathbf{H}_t$ are always in the interval $(-1, 1)$.
#
# $$\mathbf{H}_t = \mathbf{O}_t \odot \tanh(\mathbf{C}_t).$$
#
#
# Whenever the output gate approximates 1 we effectively pass all memory information through to the predictor, whereas for the output gate close to 0 we retain all the information only within the memory cell and perform no further processing.
#
#
#
# :numref:`lstm_3` has a graphical illustration of the data flow.
#
# 
# :label:`lstm_3`
#
#
#
# ## Implementation from Scratch
#
# Now let us implement an LSTM from scratch.
# As same as the experiments in :numref:`sec_rnn_scratch`,
# we first load the time machine dataset.
#
# + origin_pos=2 tab=["pytorch"]
import torch
from torch import nn
from d2l import torch as d2l
batch_size, num_steps = 32, 35
train_iter, vocab = d2l.load_data_time_machine(batch_size, num_steps)
# + [markdown] origin_pos=4
# ### [**Initializing Model Parameters**]
#
# Next we need to define and initialize the model parameters. As previously, the hyperparameter `num_hiddens` defines the number of hidden units. We initialize weights following a Gaussian distribution with 0.01 standard deviation, and we set the biases to 0.
#
# + origin_pos=6 tab=["pytorch"]
def get_lstm_params(vocab_size, num_hiddens, device):
num_inputs = num_outputs = vocab_size
def normal(shape):
return torch.randn(size=shape, device=device)*0.01
def three():
return (normal((num_inputs, num_hiddens)),
normal((num_hiddens, num_hiddens)),
torch.zeros(num_hiddens, device=device))
W_xi, W_hi, b_i = three() # Input gate parameters
W_xf, W_hf, b_f = three() # Forget gate parameters
W_xo, W_ho, b_o = three() # Output gate parameters
W_xc, W_hc, b_c = three() # Candidate memory cell parameters
# Output layer parameters
W_hq = normal((num_hiddens, num_outputs))
b_q = torch.zeros(num_outputs, device=device)
# Attach gradients
params = [W_xi, W_hi, b_i, W_xf, W_hf, b_f, W_xo, W_ho, b_o, W_xc, W_hc,
b_c, W_hq, b_q]
for param in params:
param.requires_grad_(True)
return params
# + [markdown] origin_pos=8
# ### Defining the Model
#
# In [**the initialization function**], the hidden state of the LSTM needs to return an *additional* memory cell with a value of 0 and a shape of (batch size, number of hidden units). Hence we get the following state initialization.
#
# + origin_pos=10 tab=["pytorch"]
def init_lstm_state(batch_size, num_hiddens, device):
return (torch.zeros((batch_size, num_hiddens), device=device),
torch.zeros((batch_size, num_hiddens), device=device))
# + [markdown] origin_pos=12
# [**The actual model**] is defined just like what we discussed before: providing three gates and an auxiliary memory cell. Note that only the hidden state is passed to the output layer. The memory cell $\mathbf{C}_t$ does not directly participate in the output computation.
#
# + origin_pos=14 tab=["pytorch"]
def lstm(inputs, state, params):
[W_xi, W_hi, b_i, W_xf, W_hf, b_f, W_xo, W_ho, b_o, W_xc, W_hc, b_c,
W_hq, b_q] = params
(H, C) = state
outputs = []
for X in inputs:
I = torch.sigmoid((X @ W_xi) + (H @ W_hi) + b_i)
F = torch.sigmoid((X @ W_xf) + (H @ W_hf) + b_f)
O = torch.sigmoid((X @ W_xo) + (H @ W_ho) + b_o)
C_tilda = torch.tanh((X @ W_xc) + (H @ W_hc) + b_c)
C = F * C + I * C_tilda
H = O * torch.tanh(C)
Y = (H @ W_hq) + b_q
outputs.append(Y)
return torch.cat(outputs, dim=0), (H, C)
# + [markdown] origin_pos=16
# ### [**Training**] and Prediction
#
# Let us train an LSTM as same as what we did in :numref:`sec_gru`, by instantiating the `RNNModelScratch` class as introduced in :numref:`sec_rnn_scratch`.
#
# + origin_pos=17 tab=["pytorch"]
vocab_size, num_hiddens, device = len(vocab), 256, d2l.try_gpu()
num_epochs, lr = 500, 1
model = d2l.RNNModelScratch(len(vocab), num_hiddens, device, get_lstm_params,
init_lstm_state, lstm)
d2l.train_ch8(model, train_iter, vocab, lr, num_epochs, device)
# + [markdown] origin_pos=19
# ## [**Concise Implementation**]
#
# Using high-level APIs,
# we can directly instantiate an `LSTM` model.
# This encapsulates all the configuration details that we made explicit above. The code is significantly faster as it uses compiled operators rather than Python for many details that we spelled out in detail before.
#
# + origin_pos=21 tab=["pytorch"]
num_inputs = vocab_size
lstm_layer = nn.LSTM(num_inputs, num_hiddens)
model = d2l.RNNModel(lstm_layer, len(vocab))
model = model.to(device)
d2l.train_ch8(model, train_iter, vocab, lr, num_epochs, device)
# + [markdown] origin_pos=23
# LSTMs are the prototypical latent variable autoregressive model with nontrivial state control.
# Many variants thereof have been proposed over the years, e.g., multiple layers, residual connections, different types of regularization. However, training LSTMs and other sequence models (such as GRUs) are quite costly due to the long range dependency of the sequence.
# Later we will encounter alternative models such as transformers that can be used in some cases.
#
#
# ## Summary
#
# * LSTMs have three types of gates: input gates, forget gates, and output gates that control the flow of information.
# * The hidden layer output of LSTM includes the hidden state and the memory cell. Only the hidden state is passed into the output layer. The memory cell is entirely internal.
# * LSTMs can alleviate vanishing and exploding gradients.
#
#
# ## Exercises
#
# 1. Adjust the hyperparameters and analyze the their influence on running time, perplexity, and the output sequence.
# 1. How would you need to change the model to generate proper words as opposed to sequences of characters?
# 1. Compare the computational cost for GRUs, LSTMs, and regular RNNs for a given hidden dimension. Pay special attention to the training and inference cost.
# 1. Since the candidate memory cell ensures that the value range is between $-1$ and $1$ by using the $\tanh$ function, why does the hidden state need to use the $\tanh$ function again to ensure that the output value range is between $-1$ and $1$?
# 1. Implement an LSTM model for time series prediction rather than character sequence prediction.
#
# + [markdown] origin_pos=25 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1057)
#
| d2l/pytorch/chapter_recurrent-modern/lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pF--zlL9FNaC"
# # Data Types
# + [markdown] id="dIHrvmDnFNaD"
# Every value in Python has a datatype. Since everything is an object in Python programming, data types are actually classes and variables are instance (object) of these classes.
# + [markdown] id="FZUmrAT4FNaE"
# # Numbers
# + [markdown] id="1FED9RJ2FNaE"
# Integers, floating point numbers and complex numbers falls under Python numbers category. They are defined as int, float and complex class in Python.
#
# We can use the type() function to know which class a variable or a value belongs to and the isinstance() function to check if an object belongs to a particular class.
# + id="M1KVhucoFNaG" outputId="327cddc0-8aef-4317-9cc7-71154a30dfa2"
a = 5 #data type is implicitly set to integer
print(a, " is of type", type(a))
# + id="TiBJStWMFNaH" outputId="eed27c37-e182-47a9-87d3-7597af27656c"
a = 2.5 #data type is changed to float
print(a, " is of type", type(a))
# + id="9D1gtN-oFNaH" outputId="3d4d7a16-ed7e-49aa-a6da-4da3b80940b1"
a = 1 + 2j #data type is changed to complex number
print(a, " is complex number?")
print(isinstance(1+2j, complex))
# + [markdown] id="KVmH9kihFNaI"
# # Boolean
# + [markdown] id="uYPK_FXfFNaJ"
# Boolean represents the truth values False and True
# + id="6SoUODOCFNaJ" outputId="337a8de8-1d1f-4463-86c8-cf19577eccb9"
a = True #a is a boolean type
print(type(a))
# + [markdown] id="FTF__TtnFNaJ"
# # Python Strings
# + [markdown] id="EJsqYygyFNaK"
# String is sequence of Unicode characters.
#
# We can use single quotes or double quotes to represent strings.
#
# Multi-line strings can be denoted using triple quotes, ''' or """.
# + [markdown] id="U9HXLh-3FNaL"
# A string in Python consists of a series or sequence of characters - letters, numbers, and special characters.
#
# Strings can be indexed - often synonymously called subscripted as well.
#
# Similar to C, the first character of a string has the index 0.
# + colab={"base_uri": "https://localhost:8080/"} id="yI1n-PvrFNaL" outputId="47f83371-2c1c-4cc0-ab04-27fdf2c6e483"
s = "Hello i am <NAME>"
print(s)
# + colab={"base_uri": "https://localhost:8080/"} id="XSYDbCfqFNaL" outputId="51c07d08-d1a0-4bed-9a30-b8a69fefc8eb"
print(s[0])
#last char s[len(s)-1] or s[-1]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ko_vy0OAFNaM" outputId="a33babab-2144-4eff-beb8-c16837caf0ac"
#slicing
s[5:]
# + [markdown] id="uIG4oLjhMi25"
# ### Mutable vs Imutable
# An object whose internal state can be changed is mutable. On the other hand, immutable doesn't allow any change in the object once it has been created.
# + [markdown] id="W2qNRCA0FNaN"
# # Python List
# + [markdown] id="Bosn4KbRFNaN"
# List is an ordered sequence of items. It is one of the most used datatype in Python and is very flexible. All the items in a list do not need to be of the same type.
#
# Declaring a list is , Items separated by commas are enclosed within brackets [ ].
# + [markdown] id="pxXQ8qyZL0Qe"
# Lists are used to store multiple items in a single variable. Lists are one of 4 built-in data types in Python used to store collections of data, the other 3 are Tuple, Set, and Dictionary, all with different qualities and usage.
# + colab={"base_uri": "https://localhost:8080/"} id="8iWT6WPFFNaN" outputId="791a15b7-bac3-4d21-a3b7-718747651ad6"
a = [10, 20.5, "Hello"]
print(a[1]) #print 1st index element
# + [markdown] id="iv_t3ZoWFNaO"
# Lists are mutable, meaning, value of elements of a list can be altered.
# + colab={"base_uri": "https://localhost:8080/"} id="Eb5aEhNkFNaO" outputId="8f9ba15b-ec2c-4e25-e161-67bbc3e54bfa"
a[1] = 30.7
print(a)
# + [markdown] id="Za9lEe-uFNaO"
# # Python Tuple
# + [markdown] id="me4o-mCSFNaP"
# Tuple is an ordered sequence of items same as list.The only difference is that tuples are immutable. Tuples once created cannot be modified.
# + [markdown] id="5ubG2na4MEJU"
# Use Tuple
# - If your data should or does not need to be changed.
# - Tuples are faster than lists.
# - We should use Tuple instead of a List if we are defining a constant set of values and all we are ever going to do with it is iterate through it.
# - If we need an array of elements to be used as dictionary keys, we can use Tuples.
# + id="YaZY_pyRFNaP"
t = (1, 1.5, "ML")
# + colab={"base_uri": "https://localhost:8080/"} id="pbkx-2V6FNaP" outputId="7f5eb0ed-8d79-4835-c9be-9eee79411fb1"
print(t[1]) #extract particular element
# + colab={"base_uri": "https://localhost:8080/", "height": 165} id="6rNBsEKMFNaQ" outputId="96f4aeb6-9db4-4f4c-b35d-85e432f7c613"
t[1] = 1.25
# + [markdown] id="fTXQpXc0FNaQ"
# # Python Set
# + [markdown] id="ToyQxQPKFNaQ"
# Set is an unordered collection of unique items. Set is defined by values separated by comma inside braces { }. Items in a set are not ordered.
# + colab={"base_uri": "https://localhost:8080/"} id="w7eRy2EtFNaR" outputId="efbb7d84-f37d-46cf-9566-cbcf43b4f1fd"
a = {10, 30, 20, 40, 5}
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="o2sOjgpeFNaR" outputId="e302e094-d977-4344-fe03-433d27cb81d9"
print(type(a)) #print type of a
# + [markdown] id="4Jam-vVOFNaS"
# We can perform set operations like union, intersection on two sets. Set have unique values.
# + colab={"base_uri": "https://localhost:8080/"} id="hRahtOyoFNaS" outputId="2c005473-9918-473a-a4eb-156c565cdd2e"
s = {10, 20, 20, 30, 30, 30}
print(s) #automatically set won't consider duplicate elements
# + colab={"base_uri": "https://localhost:8080/", "height": 182} id="iF947WoOFNaS" outputId="2b7f9f9e-e9c8-4ed4-a2df-c43fcef6a046"
print(s[1]) #we can't print particular element in set because
#it's unorder collections of items
# + [markdown] id="Hdjv0SSeFNaT"
# # Python Dictionary
# + [markdown] id="S6LMol5HFNaT"
# Dictionary is an unordered collection of key-value pairs.
# + [markdown] id="2JPH3d-dFNaT"
# In Python, dictionaries are defined within braces {} with each item being a pair in the form key:value. Key and value can be of any type.
# + colab={"base_uri": "https://localhost:8080/", "height": 130} id="N1bpEdWnFNaU" outputId="845f3a38-1d52-4404-bdce-0172f51a197f"
d = {'a': "apple", 'b': "bat"}
print d['a']
| 4_Datatypes/4_datatypes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to machine learning models
#
#
# ## A quick aside: types of ML
#
# As you get deeper into data science, it might seem like there are a bewildering array of ML algorithms out there. However many you encounter, it can be handy to remember that most ML algorithms fall into three broad categories:
# - **Predictive algorithms**: These analyze current and historical facts to make predictions about unknown events, such as the future or customers’ choices.
# - **Classification algorithms**: These teach a program from a body of data, and the program then uses that learning to classify new observations.
# - **Time-series forecasting algorithms**: While it can argued that these algorithms are a part of predictive algorithms, their techniques are specialized enough that they in many ways functions like a separate category. Time-series forecasting is beyond the scope of this course, but we have more than enough work with focusing here on prediction and classification.
#
# ## Prediction: linear regression
#
# > **Learning goal:** By the end of this subsection, you should be comfortable fitting linear regression models, and you should have some familiarity with interpreting their output.
# ### Data exploration
#
# **Import Libraries**
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# **Dataset Alert**: Boston Housing Dataset
df = pd.read_csv('Data/Housing_Dataset_Sample.csv')
df.head()
# ### Exercise:
# Do you remember the DataFrame method for looking at overall information
# about a DataFrame, such as number of columns and rows? Try it here.
df.describe().T
# **Price Column**
sns.distplot(df['Price'])
# **House Prices vs Average Area Income**
sns.jointplot(df['Avg. Area Income'],df['Price'])
# **All Columns**
sns.pairplot(df)
# **Some observations**
# 1. Blob Data
# 2. Distortions might be a result of data (e.g. no one has 0.3 rooms)
#
#
# ### Fitting the model
#
# **Can We Predict Housing Prices?**
X = df.iloc[:,:5] # First 5 Columns
y = df['Price'] # Price Column
# **Train, Test, Split**
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3,random_state=54)
# **Fit to Linear Regression Model**
# +
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
# -
reg.fit(X_train,y_train)
# ### Evaluating the model
#
# **Predict**
predictions = reg.predict(X_test)
predictions
print(reg.intercept_,reg.coef_)
# **Score**
# +
#Explained variation. A high R2 close to 1 indicates better prediction with less error.
from sklearn.metrics import r2_score
r2_score(y_test,predictions)
# -
# **Visualize Errors**
sns.distplot([y_test-predictions])
# **Visualize Predictions**
# Plot outputs
plt.scatter(y_test,predictions, color='blue')
# ### Exercise:
# Can you think of a way to refine this visualization to make it clearer, particularly if you were explaining the results to someone?
# Hint: Remember to try the plt.scatter parameter alpha=.
# It takes values between 0 and 1.
# > **Takeaway:** In this subsection, you performed prediction using linear regression by exploring your data, then fitting your model, and finally evaluating your model’s performance.
# ## Classification: logistic regression
#
# > **Learning goal:** By the end of this subsection, you should know how logistic regression differs from linear regression, be comfortable fitting logistic regression models, and have some familiarity with interpreting their output.
# **Dataset Alert**: Fates of RMS Titanic Passengers
#
# The dataset has 12 variables:
# - **PassengerId**
# - **Survived:** 0 = No, 1 = Yes
# - **Pclass:** Ticket class 1 = 1st, 2 = 2nd, 3 = 3rd
# - **Sex**
# - **Age**
# - **Sibsp:** Number of siblings or spouses aboard the *Titanic*
# - **Parch:** Number of parents or children aboard the *Titanic*
# - **Ticket:** Passenger ticket number
# - **Fare:** Passenger fare
# - **Cabin:** Cabin number
# - **Embarked:** Port of embarkation; C = Cherbourg, Q = Queenstown, S = Southampton
df = pd.read_csv('Data/train_data_titanic.csv')
df.head()
df.info()
#
#
# ### Remove extraneous variables
df.drop(['Name','Ticket'],axis=1,inplace=True)
#
#
# ### Check for multicollinearity
#
# **Question**: Do any correlations between **Survived** and **Fare** jump out?
sns.pairplot(df[['Survived','Fare']], dropna=True)
# ### Exercise:
# Try running sns.pairplot twice more on some other combinations of columns
# and see if any patterns emerge.
# We can also use `groupby` to look for patterns. Consider the mean values for the various variables when we group by **Survived**:
df.groupby('Survived').mean()
df.head()
df['SibSp'].value_counts()
df['Parch'].value_counts()
df['Sex'].value_counts()
# ### Handle missing values
#
# missing
df.isnull().sum()>(len(df)/2)
df.drop('Cabin',axis=1,inplace=True)
df.info()
df['Age'].isnull().value_counts()
# ### Corelation Exploration
df.groupby('Sex')['Age'].median().plot(kind='bar')
df['Age'] = df.groupby('Sex')['Age'].apply(lambda x: x.fillna(x.median()))
df.isnull().sum()
df['Embarked'].value_counts()
df['Embarked'].fillna(df['Embarked'].value_counts().idxmax(), inplace=True)
df['Embarked'].value_counts()
df = pd.get_dummies(data=df, columns=['Sex', 'Embarked'],drop_first=True)
df.head()
# **Correlation Matrix**
df.corr()
# **Define X and Y**
X = df.drop(['Survived','Pclass'],axis=1)
y = df['Survived']
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3,random_state=67)
# ### Exercise:
#
# We now need to split the training and test data, which you will so as an exercise:
from sklearn.model_selection import train_test_split
# Look up in the portion above on linear regression and use train_test_split here.
# Set test_size = 0.3 and random_state = 67 to get the same results as below when
# you run through the rest of the code example below.
# **Use Logistic Regression Model**
# +
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
# -
lr.fit(X_train,y_train)
predictions = lr.predict(X_test)
# ### Evaluate the model
#
#
# #### Classification report
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
# The classification reports the proportions of both survivors and non-survivors with four scores:
# - **Precision:** The number of true positives divided by the sum of true positives and false positives; closer to 1 is better.
# - **Recall:** The true-positive rate, the number of true positives divided by the sum of the true positives and the false negatives.
# - **F1 score:** The harmonic mean (the average for rates) of precision and recall.
# - **Support:** The number of true instances for each label.
print(classification_report(y_test,predictions))
# #### Confusion matrix
print(confusion_matrix(y_test,predictions))
pd.DataFrame(confusion_matrix(y_test, predictions), columns=['True Survived', 'True Not Survived'], index=['Predicted Survived', 'Predicted Not Survived'])
# #### Accuracy score
print(accuracy_score(y_test,predictions))
# > **Takeaway:** In this subsection, you performed classification using logistic regression by removing extraneous variables, checking for multicollinearity, handling missing values, and fitting and evaluating your model.
# ## Classification: decision trees
#
# > **Learning goal:** By the end of this subsection, you should be comfortable fitting decision-tree models and have some understanding of what they output.
from sklearn import tree
tr = tree.DecisionTreeClassifier()
# ### Exercise:
# Using the same split data as with the logistic regression,
# can you fit the decision tree model?
# Hint: Refer to code snippet for fitting the logistic regression above.
# **Note**: Using the same Titanic Data
tr.fit(X_train, y_train)
tr_predictions = tr.predict(X_test)
pd.DataFrame(confusion_matrix(y_test, tr_predictions),
columns=['True Survived', 'True Not Survived'],
index=['Predicted Survived', 'Predicted Not Survived'])
print(accuracy_score(y_test,tr_predictions))
# **Visualize tree**
# +
import graphviz
dot_file = tree.export_graphviz(tr, out_file=None,
feature_names=X.columns,
class_names='Survived',
filled=True,rounded=True)
graph = graphviz.Source(dot_file)
graph
# -
# > **Takeaway:** In this subsection, you performed classification on previously cleaned data by fitting and evaluating a decision tree.
| Data_Science_2/Full_Day/Instructor Materials/2-MachineLearningModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pathlib import Path
from drone_detector.tiling import *
import os, sys
import geopandas as gpd
import pandas as pd
import rasterio as rio
import math
from shapely.geometry import box
from shapely.geometry import Polygon
def fix_multipolys(multipoly):
temp_poly = None
max_area = 0
for geom in multipoly.geoms:
area = geom.area
if area > max_area:
max_area = area
temp_poly = geom
return Polygon(temp_poly.exterior)
# +
from shapely.geometry import Point
def get_len(geom):
mrr = geom.minimum_rotated_rectangle
x, y = mrr.exterior.coords.xy
edge_len = (Point(x[0], y[0]).distance(Point(x[1], y[1])), Point(x[1], y[1]).distance(Point(x[2], y[2])))
return max(edge_len)
# -
data_path = Path('../data/hiidenportti/resnet50_valsplit/label_nms/eval_results')
# # Ground Truths (TP/FN) -comparison
gts = gpd.read_file(data_path/'gts.geojson')
fns = gts[gts.det_type == 'FN'].copy()
gt_tps = gts[gts.det_type == 'TP'].copy()
# ## False negatives
fns['geometry'] = fns.apply(lambda row: fix_multipolys(row.geometry) if row.geometry.type == 'MultiPolygon'
else Polygon(row.geometry.exterior), axis=1)
fns['area'] = fns.geometry.area
fns['pixel_area'] = fns.area / 0.04**2
fns['bbox_area'] = fns.apply(lambda row: box(*row.geometry.bounds).area, axis=1)
fns['bbox_x_px'] = fns.apply(lambda row: (row.geometry.bounds[2] - row.geometry.bounds[0]) / 0.04, axis=1)
fns['bbox_y_px'] = fns.apply(lambda row: (row.geometry.bounds[3] - row.geometry.bounds[1]) / 0.04, axis=1)
fns['bbox_pixel_area'] = fns.bbox_area / 0.04**2
fns['bbox_aspect_ratio'] = fns.bbox_x_px / fns.bbox_y_px
fns['length'] = fns.apply(lambda row: get_len(row.geometry), axis=1)
pd.pivot_table(fns, index='tree_type', values='area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fns, index='tree_type', values='pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fns, index='tree_type', values='bbox_pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fns, index='tree_type', values='bbox_aspect_ratio', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fns, index='tree_type', values='bbox_x_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fns.plot(kind='hist', y='bbox_x_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(fns, index='tree_type', values='bbox_y_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fns.plot(kind='hist', y='bbox_y_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(fns, index='tree_type', values='length', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fns[fns.tree_type =='Fallen'].plot(kind='hist', y='length', bins=100, grid=True,
title='Distribution of groundwood length in meters',
figsize=(10,6))
fns[fns.tree_type =='Standing'].plot(kind='hist', y='length', bins=10, grid=True,
title='Distribution of uprightwood maximum canopy diameter, in meters',
figsize=(10,6))
# ## True positives
gt_tps['geometry'] = gt_tps.apply(lambda row: fix_multipolys(row.geometry) if row.geometry.type == 'MultiPolygon'
else Polygon(row.geometry.exterior), axis=1)
gt_tps['area'] = gt_tps.geometry.area
gt_tps['pixel_area'] = gt_tps.area / 0.04**2
gt_tps['bbox_area'] = gt_tps.apply(lambda row: box(*row.geometry.bounds).area, axis=1)
gt_tps['bbox_x_px'] = gt_tps.apply(lambda row: (row.geometry.bounds[2] - row.geometry.bounds[0]) / 0.04, axis=1)
gt_tps['bbox_y_px'] = gt_tps.apply(lambda row: (row.geometry.bounds[3] - row.geometry.bounds[1]) / 0.04, axis=1)
gt_tps['bbox_pixel_area'] = gt_tps.bbox_area / 0.04**2
gt_tps['bbox_aspect_ratio'] = gt_tps.bbox_x_px / gt_tps.bbox_y_px
gt_tps['length'] = gt_tps.apply(lambda row: get_len(row.geometry), axis=1)
pd.pivot_table(gt_tps, index='tree_type', values='area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(gt_tps, index='tree_type', values='pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(gt_tps, index='tree_type', values='bbox_pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(gt_tps, index='tree_type', values='bbox_aspect_ratio', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(gt_tps, index='tree_type', values='bbox_x_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
gt_tps.plot(kind='hist', y='bbox_x_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(gt_tps, index='tree_type', values='bbox_y_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
gt_tps.plot(kind='hist', y='bbox_y_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(gt_tps, index='tree_type', values='length', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
gt_tps[gt_tps.tree_type =='Fallen'].plot(kind='hist', y='length', bins=50, grid=True,
title='Distribution of groundwood length in meters',
figsize=(10,6))
gt_tps[gt_tps.tree_type =='Standing'].plot(kind='hist', y='length', bins=20, grid=True,
title='Distribution of uprightwood maximum canopy diameter, in meters',
figsize=(10,6))
# # Detections (TP/FP) -comparison
gts = gpd.read_file(data_path/'dts.geojson')
fps = gts[gts.det_type == 'FP'].copy()
dt_tps = gts[gts.det_type == 'TP'].copy()
# ## False positives
fps['geometry'] = fps.apply(lambda row: fix_multipolys(row.geometry) if row.geometry.type == 'MultiPolygon'
else Polygon(row.geometry.exterior), axis=1)
fps['area'] = fps.geometry.area
fps['pixel_area'] = fps.area / 0.04**2
fps['bbox_area'] = fps.apply(lambda row: box(*row.geometry.bounds).area, axis=1)
fps['bbox_x_px'] = fps.apply(lambda row: (row.geometry.bounds[2] - row.geometry.bounds[0]) / 0.04, axis=1)
fps['bbox_y_px'] = fps.apply(lambda row: (row.geometry.bounds[3] - row.geometry.bounds[1]) / 0.04, axis=1)
fps['bbox_pixel_area'] = fps.bbox_area / 0.04**2
fps['bbox_aspect_ratio'] = fps.bbox_x_px / fps.bbox_y_px
fps['length'] = fps.apply(lambda row: get_len(row.geometry), axis=1)
pd.pivot_table(fps, index='tree_type', values='area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fps, index='tree_type', values='pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fps, index='tree_type', values='bbox_pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fps, index='tree_type', values='bbox_aspect_ratio', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(fps, index='tree_type', values='bbox_x_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fps.plot(kind='hist', y='bbox_x_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(fps, index='tree_type', values='bbox_y_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fps.plot(kind='hist', y='bbox_y_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(fps, index='tree_type', values='length', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
fps[fps.tree_type =='Fallen'].plot(kind='hist', y='length', bins=100, grid=True,
title='Distribution of groundwood length in meters',
figsize=(10,6))
fps[fps.tree_type =='Standing'].plot(kind='hist', y='length', bins=10, grid=True,
title='Distribution of uprightwood maximum canopy diameter, in meters',
figsize=(10,6))
# ## True positives
dt_tps['geometry'] = dt_tps.apply(lambda row: fix_multipolys(row.geometry) if row.geometry.type == 'MultiPolygon'
else Polygon(row.geometry.exterior), axis=1)
dt_tps['area'] = dt_tps.geometry.area
dt_tps['pixel_area'] = dt_tps.area / 0.04**2
dt_tps['bbox_area'] = dt_tps.apply(lambda row: box(*row.geometry.bounds).area, axis=1)
dt_tps['bbox_x_px'] = dt_tps.apply(lambda row: (row.geometry.bounds[2] - row.geometry.bounds[0]) / 0.04, axis=1)
dt_tps['bbox_y_px'] = dt_tps.apply(lambda row: (row.geometry.bounds[3] - row.geometry.bounds[1]) / 0.04, axis=1)
dt_tps['bbox_pixel_area'] = dt_tps.bbox_area / 0.04**2
dt_tps['bbox_aspect_ratio'] = dt_tps.bbox_x_px / dt_tps.bbox_y_px
dt_tps['length'] = dt_tps.apply(lambda row: get_len(row.geometry), axis=1)
pd.pivot_table(dt_tps, index='tree_type', values='area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(dt_tps, index='tree_type', values='pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(dt_tps, index='tree_type', values='bbox_pixel_area', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(dt_tps, index='tree_type', values='bbox_aspect_ratio', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
pd.pivot_table(dt_tps, index='tree_type', values='bbox_x_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
dt_tps.plot(kind='hist', y='bbox_x_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(dt_tps, index='tree_type', values='bbox_y_px', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
dt_tps.plot(kind='hist', y='bbox_y_px', bins=100, grid=True,
title='Distribution of bounding box width, in pixels',
figsize=(10,6))
pd.pivot_table(dt_tps, index='tree_type', values='length', aggfunc=['min', 'max', 'mean', 'median'], margins=True)
dt_tps[dt_tps.tree_type =='Fallen'].plot(kind='hist', y='length', bins=50, grid=True,
title='Distribution of groundwood length in meters',
figsize=(10,6))
dt_tps[dt_tps.tree_type =='Standing'].plot(kind='hist', y='length', bins=20, grid=True,
title='Distribution of uprightwood maximum canopy diameter, in meters',
figsize=(10,6))
| analyses/Result analyses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Fully Bayesian inference for generalized GP models with HMC
# --
#
# *<NAME>, 2015-16*
#
# It's possible to construct a very flexible models with Gaussian processes by combining them with different likelihoods (sometimes called 'families' in the GLM literature). This makes inference of the GP intractable since the likelihoods is not generally conjugate to the Gaussian process. The general form of the model is
# $$\theta \sim p(\theta)\\f \sim \mathcal {GP}(m(x; \theta),\, k(x, x'; \theta))\\y_i \sim p(y | g(f(x_i))\,.$$
#
#
# To perform inference in this model, we'll run MCMC using Hamiltonian Monte Carlo (HMC) over the function-values and the parameters $\theta$ jointly. Key to an effective scheme is rotation of the field using the Cholesky decomposition. We write
#
# $$\theta \sim p(\theta)\\v \sim \mathcal {N}(0,\, I)\\LL^\top = K\\f = m + Lv\\y_i \sim p(y | g(f(x_i))\,.$$
#
# Joint HMC over v and the function values is not widely adopted in the literature becate of the difficulty in differentiating $LL^\top=K$. We've made this derivative available in tensorflow, and so application of HMC is relatively straightforward.
# ### Exponential Regression example
# The first illustration in this notebook is 'Exponential Regression'. The model is
# $$\theta \sim p(\theta)\\f \sim \mathcal {GP}(0, k(x, x'; \theta))\\f_i = f(x_i)\\y_i \sim \mathcal {Exp} (e^{f_i})$$
#
# We'll use MCMC to deal with both the kernel parameters $\theta$ and the latent function values $f$. first, generate a data set.
# +
import gpflow
import numpy as np
from matplotlib import pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
X = np.linspace(-3,3,20)
Y = np.random.exponential(np.sin(X)**2)
# -
# GPflow's model for fully-Bayesian MCMC is called GPMC. It's constructed like any other model, but contains a parameter `V` which represents the centered values of the function.
#build the model
k = gpflow.kernels.Matern32(1,ARD=False) + gpflow.kernels.Bias(1)
l = gpflow.likelihoods.Exponential()
m = gpflow.gpmc.GPMC(X[:,None], Y[:,None], k, l)
m
# The `V` parameter already has a prior applied. We'll add priors to the parameters also (these are rather arbitrary, for illustration).
m.kern.matern32.lengthscales.prior = gpflow.priors.Gamma(1., 1.)
m.kern.matern32.variance.prior = gpflow.priors.Gamma(1.,1.)
m.kern.bias.variance.prior = gpflow.priors.Gamma(1.,1.)
# Running HMC is as easy as hitting m.sample(). GPflow only has HMC sampling for the moment, and it's a relatively vanilla implementation (no NUTS, for example). There are two setting to tune, the step size (epsilon) and the maximum noumber of steps Lmax. Each proposal will take a random number of steps between 1 and Lmax, each of length epsilon.
#
# We'll use the `verbose` setting so that we can see the acceptance rate.
m.optimize(maxiter=15) # start near MAP
samples = m.sample(500, verbose=True, epsilon=0.12, Lmax=15)
xtest = np.linspace(-4,4,100)[:,None]
f_samples = []
for s in samples:
m.set_state(s)
f_samples.append(m.predict_f_samples(xtest, 5))
f_samples = np.vstack(f_samples)
rate_samples = np.exp(f_samples[:,:,0])
plt.figure(figsize=(12, 6))
line, = plt.plot(xtest, np.mean(rate_samples, 0), lw=2)
plt.fill_between(xtest[:,0], np.percentile(rate_samples, 5, axis=0), np.percentile(rate_samples, 95, axis=0), color=line.get_color(), alpha = 0.2)
plt.plot(X, Y, 'kx', mew=2)
plt.ylim(-0.1, np.max(np.percentile(rate_samples, 95, axis=0)))
_ = plt.plot(samples)
kernel_samples = m.kern.get_samples_df(samples)
kernel_samples.head()
kernel_samples.mean()
kernel_samples.std()
| GPflow/doc/source/notebooks/mcmc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iuNWUJKJLY9T"
# %reset -f
# + id="yj4HfL7EcNRq"
# libraries used
# https://stats.stackexchange.com/questions/181/how-to-choose-the-number-of-hidden-layers-and-nodes-in-a-feedforward-neural-netw
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from sklearn import preprocessing
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
import itertools
# + colab={"base_uri": "https://localhost:8080/"} id="WYIp8Qr9ck8Z" outputId="b5df941b-73c3-4385-b9e0-aeff51f4c890"
emotions = pd.read_csv("drive/MyDrive/EEG/emotions.csv")
emotions.replace(['NEGATIVE', 'POSITIVE', 'NEUTRAL'], [2, 1, 0], inplace=True)
emotions['label'].unique()
# + id="PvT5tTotcoS3"
X = emotions.drop('label', axis=1).copy()
y = (emotions['label'].copy())
# + id="GUZOuZK7crNV"
# Splitting data into training and testing as 80-20
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=1)
x = X_train #returns a numpy array
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df = pd.DataFrame(x_scaled)
# + id="BhRoXS4mj6CV"
# resetting the data - https://www.tensorflow.org/api_docs/python/tf/keras/backend/clear_session
tf.keras.backend.clear_session()
# + colab={"base_uri": "https://localhost:8080/"} id="k4a4vwF5cxvU" outputId="09387f81-273d-44d8-9abf-6f0a6358c892"
model = Sequential()
model.add(Dense((2*X_train.shape[1]/3), input_dim=X_train.shape[1], activation='relu'))
model.add(Dense((2*X_train.shape[1]/3), activation='relu'))
model.add(Dense((1*X_train.shape[1]/3), activation='relu'))
model.add(Dense((1*X_train.shape[1]/3), activation='relu'))
model.add(Dense(3, activation='softmax'))
#model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
# + id="A2Q9BOrneeUM"
# for categorical entropy
# https://stackoverflow.com/questions/63211181/error-while-using-categorical-crossentropy
from tensorflow.keras.utils import to_categorical
Y_one_hot=to_categorical(y_train) # convert Y into an one-hot vector
# + id="NEs5ddrJeBeG"
# https://stackoverflow.com/questions/59737875/keras-change-learning-rate
#optimizer = tf.keras.optimizers.Adam(0.001)
#optimizer.learning_rate.assign(0.01)
opt = keras.optimizers.Adadelta(learning_rate=0.001)
model.compile(
optimizer=opt,
loss='sparse_categorical_crossentropy',
metrics=['accuracy']
)
# to be run for categorical cross entropy
# model.compile(loss='categorical_crossentropy', optimizer=tf.keras.optimizers.Adadelta(learning_rate=0.001), metrics=['accuracy'])
# + id="ASdP3qvxes8r" colab={"base_uri": "https://localhost:8080/"} outputId="a3309126-c339-4f34-b018-92744adf4e3d"
# make sure that the input data is shuffled before hand so that the model doesn't notice patterns and generalizes well
# change y_train to y_hot_encoded when using categorical cross entorpy
import time
start_time = time.time()
history = model.fit(
df,
y_train,
validation_split=0.2,
batch_size=32,
epochs=75)
# to be run for categorical cross entropy
#history = model.fit(
# df,
# Y_one_hot,
# validation_split=0.2,
# batch_size=32,
# epochs=75)
# + id="vhwajgGzpvrD" colab={"base_uri": "https://localhost:8080/"} outputId="0dacc101-c086-4548-855b-cd13c8750214"
history.history
# + colab={"base_uri": "https://localhost:8080/"} id="Kyn1dHRRJ5Wn" outputId="9d3de477-542c-43e4-a507-f65517728d9e"
print("--- %s seconds ---" % (time.time() - start_time))
# + id="rOU0i1e6jhr3"
x_test = X_test #returns a numpy array
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled_test = min_max_scaler.fit_transform(x_test)
df_test = pd.DataFrame(x_scaled_test)
# + id="OgeGekJ8lW6S"
predictions = model.predict(x=df_test, batch_size=32)
# + id="1o6zqIsfl2QX"
rounded_predictions = np.argmax(predictions, axis=-1)
# + id="NQTy4F7TmotU"
cm = confusion_matrix(y_true=y_test, y_pred=rounded_predictions)
# + colab={"base_uri": "https://localhost:8080/", "height": 922} id="rieDywHXuA6R" outputId="61ce936e-4e3c-4abf-f4ec-2a0ff5119721"
label_mapping = {'NEGATIVE': 0, 'NEUTRAL': 1, 'POSITIVE': 2}
# for diff dataset
# label_mapping = {'NEGATIVE': 0, 'POSITIVE': 1}
plt.figure(figsize=(8, 8))
sns.heatmap(cm, annot=True, vmin=0, fmt='g', cbar=False, cmap='Blues')
clr = classification_report(y_test, rounded_predictions, target_names=label_mapping.keys())
plt.xticks(np.arange(3) + 0.5, label_mapping.keys())
plt.yticks(np.arange(3) + 0.5, label_mapping.keys())
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Confusion Matrix")
plt.show()
print("Classification Report:\n----------------------\n", clr)
# + id="s5wbr_oav-Kb"
# https://stackoverflow.com/questions/26413185/how-to-recover-matplotlib-defaults-after-setting-stylesheet
import matplotlib as mpl
mpl.rcParams.update(mpl.rcParamsDefault)
# + id="73HZGymlybP2"
training_acc = history.history['accuracy']
validation_acc = history.history['val_accuracy']
training_loss = history.history['loss']
validation_loss = history.history['val_loss']
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="1jgMVKMDyxck" outputId="0e5a71c7-db98-4079-d6d2-8f4ebd9dc876"
epochs = history.epoch
plt.plot(epochs, training_acc, color = '#17e6e6', label='Training Accuracy')
plt.plot(epochs, validation_acc,color = '#e61771', label='Validation Accuracy')
plt.title('Accuracy vs Epochs')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.savefig('AccuracyVsEpochs.png')
plt.show()
| Models/Model9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/unburied/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/David_Martinez_Assignment_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - A First Look at Data
#
#
# + [markdown] id="9dtJETFRhnOG" colab_type="text"
# ## Lecture - let's explore Python DS libraries and examples!
#
# The Python Data Science ecosystem is huge. You've seen some of the big pieces - pandas, scikit-learn, matplotlib. What parts do you want to see more of?
# + [markdown] id="ajmhIxfVcXVc" colab_type="text"
# I'm excited to learn about Theano and all of its high level children. Getting into neural networks feels like it would be the best end goal, especially with Tesla's announcement recently about its adavancements using adavanced neural networks. These are exciting times for this spectrum of data science and I am eager to learn more about it.
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - now it's your turn
#
# Pick at least one Python DS library, and using documentation/examples reproduce in this notebook something cool. It's OK if you don't fully understand it or get it 100% working, but do put in effort and look things up.
# + [markdown] id="qzjN9CJWeqD8" colab_type="text"
# I hope this is ok. I actually worked on this a little after the precourse assignment was done. I feel like it demonstrates something cool and practical(ish) at the same time.
# https://colab.research.google.com/drive/1WMdcl9USQjuDE5n0JKwR0j9l-uqd7zXc
#
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
# 1. Describe in a paragraph of text what you did and why, as if you were writing an email to somebody interested but nontechnical.
#
# In the project I linked above , I was able to make some loose predictions about the company that I currently work for. We receive monthly profit reports from the CEO. I was able to gather almost 2 years worth of data, and feed it to a linear regression model. Some of the cool libraries I was able to use and learn about were google.colab to import files from my computer, and datetime to convert the dates data to useable variables. This was on top of the skilearn library which houses the linear regression model I used to make the predictions. I wanted to see approximately when cd sales where going to become unprofitable, and when our growth product, books, would intersect and surpass it as a product to keep the company afloat. The linear regression model helps with this goal, as it takes the values I submitted, and creates a line of best fit from that data. You can then trace the line past the data you have to make predictions. At the current rate, cd sales and book sales intersect in October next year, and books will steadily climb into a profitable product while cd sales continue to decline significantly.
#
# 2. What was the most challenging part of what you did?
#
# The challenging parts were importing the csv file I created in excel from my computer. It turns out colab can be a little tricky with that, and apprently a library was created to make this easier. Also, converting the dates to useable data was not trivial. I still need to gain a better understanding of what datetime.toordinal acutally does.
#
# 3. What was the most interesting thing you learned?
#
# Besides the programming aspects of overcoming the above challenges, I learned first hand that the linear regression model is very limited. For instance, the prediction is that cd sales bottom out in 5 years. I find that unrealistic, and actually expect it will plateau at some point and hover well above the zero mark for at least the next decade.
#
# 4. What area would you like to explore with more time?
#
# I would like to discover other models that could allow for the predicted data to plateau, or in a sense, output logarithmic regression as opposed to strictly linear. Also, researching whats under the hood of datetime and toordinal.
#
#
#
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub (and since this is the first assignment of the sprint, open a PR as well).
#
# - [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/)
# - [scikit-learn documentation](http://scikit-learn.org/stable/documentation.html)
# - [matplotlib documentation](https://matplotlib.org/contents.html)
# - [Awesome Data Science](https://github.com/bulutyazilim/awesome-datascience) - a list of many types of DS resources
#
# Stretch goals:
#
# - Find and read blogs, walkthroughs, and other examples of people working through cool things with data science - and share with your classmates!
# - Write a blog post (Medium is a popular place to publish) introducing yourself as somebody learning data science, and talking about what you've learned already and what you're excited to learn more about.
# + [markdown] id="YT_drbwdDrvZ" colab_type="text"
# Awesome walkthrough on associatvie analysis:
# https://pbpython.com/market-basket-analysis.html
#
# My first blog post ever:
# https://medium.com/@davi86m/conflicted-science-f767e94a1217
#
| David_Martinez_Assignment_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/UN-GCPDS/python-gcpds.EEG_Tensorflow_models/blob/main/Experimental/DW_LCAM/%5B3%5D_Main_attention_maps_comparison.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Si3756n3xnQh"
# # Load drive
# + id="XUjvvh9Vxm-p"
#-------------------------------------------------------------------------------
from google.colab import drive
drive.mount('/content/drive')
#-------------------------------------------------------------------------------
# + [markdown] id="Plu4I_-dMorV"
# # Install Keras-vis toolbox
#
#
#
# + id="FwCSY1t6J8s8"
#-------------------------------------------------------------------------------
# !pip install tf-keras-vis tensorflow
#-------------------------------------------------------------------------------
# + [markdown] id="zQrwoiJbxqTY"
# # Supporting modules
# + id="qePz_s64pqIj"
#-------------------------------------------------------------------------------
import numpy as np
import scipy.io as sio
import matplotlib.pyplot as plt
import tensorflow as tf
import pickle
from tf_keras_vis.utils.scores import CategoricalScore
from tf_keras_vis.utils.model_modifiers import ReplaceToLinear
from matplotlib import cm
from tf_keras_vis.gradcam import Gradcam
from sklearn.model_selection import ShuffleSplit
from tensorflow import keras
from tensorflow.keras.constraints import max_norm
from tensorflow.keras import backend as K
from tf_keras_vis.saliency import Saliency
from tf_keras_vis.scorecam import Scorecam
from tf_keras_vis.gradcam_plus_plus import GradcamPlusPlus
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn.manifold import TSNE
from sklearn.decomposition import KernelPCA
from sklearn.metrics import pairwise_distances
# %matplotlib inline
#-------------------------------------------------------------------------------
# + [markdown] id="HJXpdxGvxazY"
# # Define load data, normalization and CNN model function
# + id="emU_mxydxafh"
#-------------------------------------------------------------------------------
def TW_data(sbj,time_inf,time_sup):
# Load data/images----------------------------------------------------------
path_cwt = '/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/CWT_CSP_data_mubeta_8_30_Tw_'+str(time_inf)+'s_'+str(time_sup)+'s_subject'+str(sbj)+'_cwt_resized_10.pickle'
with open(path_cwt, 'rb') as f:
X_train_re_cwt, X_test_re_cwt, y_train, y_test = pickle.load(f)
path_csp = '/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/CWT_CSP_data_mubeta_8_30_Tw_'+str(time_inf)+'s_'+str(time_sup)+'s_subject'+str(sbj)+'_csp_resized_10.pickle'
with open(path_csp, 'rb') as f:
X_train_re_csp, X_test_re_csp, y_train, y_test = pickle.load(f)
#---------------------------------------------------------------------------
return X_train_re_cwt, X_train_re_csp, X_test_re_cwt, X_test_re_csp, y_train, y_test
#-------------------------------------------------------------------------------
def norm_data(XF_train_cwt, XF_train_csp, XF_test_cwt, XF_test_csp, n_fb, Ntw, y_train, y_test, fld):
# orden de las inputs:------------------------------------------------------
# [CWT_fb1_TW1, CWT_fb2_TW1 --- CWT_fb1_TW2, CWT_fb2_TW2 --- CWT_fb1_TWN, CWT_fb2_TWN] ... [CSP]
#---------------------------------------------------------------------------
XT_train_csp = []
XT_valid_csp = []
XT_test_csp = []
XT_train_cwt = []
XT_valid_cwt = []
XT_test_cwt = []
for tw in range(Ntw):
for fb in range(n_fb):
X_train_cwt, X_test_cwt = XF_train_cwt[tw][:,fb,:,:].astype(np.uint8), XF_test_cwt[tw][:,fb,:,:].astype(np.uint8)
X_train_csp, X_test_csp = XF_train_csp[tw][:,fb,:,:].astype(np.uint8), XF_test_csp[tw][:,fb,:,:].astype(np.uint8)
#-------------------------------------------------------------------
# train/validation data split
rs = ShuffleSplit(n_splits=1, test_size=.1, random_state=fld)
for train_index, valid_index in rs.split(X_train_cwt):
X_train_cwtf = X_train_cwt[train_index,:,:] # cwt
X_valid_cwtf = X_train_cwt[valid_index,:,:]
X_train_cspf = X_train_csp[train_index,:,:] # csp
X_valid_cspf = X_train_csp[valid_index,:,:]
#-------------------------------------------------------------------
# Normalize data----------------------------------------------------
X_mean_cwt = X_train_cwtf.mean(axis=0, keepdims=True)
X_std_cwt = X_train_cwtf.std(axis=0, keepdims=True) + 1e-7
X_train_cwt = (X_train_cwtf - X_mean_cwt) / X_std_cwt
X_valid_cwt = (X_valid_cwtf - X_mean_cwt) / X_std_cwt
X_test_cwt = (X_test_cwt - X_mean_cwt) / X_std_cwt
X_mean_csp = X_train_cspf.mean(axis=0, keepdims=True)
X_std_csp = X_train_cspf.std(axis=0, keepdims=True) + 1e-7
X_train_csp = (X_train_cspf - X_mean_csp) / X_std_csp
X_valid_csp = (X_valid_cspf - X_mean_csp) / X_std_csp
X_test_csp = (X_test_csp - X_mean_csp) / X_std_csp
#-------------------------------------------------------------------
# set new axis------------------------------------------------------
X_train_cwt = X_train_cwt[..., np.newaxis]
X_valid_cwt = X_valid_cwt[..., np.newaxis]
X_test_cwt = X_test_cwt[..., np.newaxis]
XT_train_cwt.append(X_train_cwt)
XT_valid_cwt.append(X_valid_cwt)
XT_test_cwt.append(X_test_cwt)
X_train_csp = X_train_csp[..., np.newaxis]
X_valid_csp = X_valid_csp[..., np.newaxis]
X_test_csp = X_test_csp[..., np.newaxis]
XT_train_csp.append(X_train_csp)
XT_valid_csp.append(X_valid_csp)
XT_test_csp.append(X_test_csp)
#-------------------------------------------------------------------
y_trainf = y_train[train_index]
y_validf = y_train[valid_index]
y_trainF, y_validF, y_testF = y_trainf.reshape((-1,))-1, y_validf.reshape((-1,))-1, y_test.reshape((-1,))-1
#---------------------------------------------------------------------------
# Convert class vectors to binary class matrices----------------------------
y_train = keras.utils.to_categorical(y_trainF,num_classes)
y_valid = keras.utils.to_categorical(y_validF,num_classes)
y_test = keras.utils.to_categorical(y_testF,num_classes)
#---------------------------------------------------------------------------
XT_train = XT_train_cwt + XT_train_csp
XT_valid = XT_valid_cwt + XT_valid_csp
XT_test = XT_test_cwt + XT_test_csp
#---------------------------------------------------------------------------
return XT_train, XT_valid, XT_test, y_train, y_valid, y_test, train_index, valid_index
#-------------------------------------------------------------------------------
def vis_heatmap(HmapT,Ntw,names_x,norm):
#-----------------------------------------------------------------------------
# normalizing heatmap
if norm == 1:
hmap_max = np.max(np.array(HmapT))
for i in range(20):
HmapT[i] = tf.math.divide_no_nan(HmapT[i],hmap_max)
new_max = np.max(np.array(HmapT))
new_min = np.min(np.array(HmapT))
else:
for i in range(20):
print(np.max(np.array(HmapT[i])),np.min(np.array(HmapT[i])))
HmapT[i] = tf.math.divide_no_nan(HmapT[i],np.max(np.array(HmapT[i])))
new_max = np.max(np.array(HmapT))
new_min = np.min(np.array(HmapT))
#-----------------------------------------------------------------------------
# figure plot setting
fig, axs = plt.subplots(4,5,figsize=(12,7.3))
fig.subplots_adjust(hspace = 0.1, wspace=.0001)
#-----------------------------------------------------------------------------
# creating figure
for tw in range(Ntw):
if tw == 0:
ids_tw = [tw, tw+1, tw+10, tw+10+1]
else:
ids_tw = [tw*2, tw*2+1, tw*2+10, tw*2+10+1]
axs[0,tw].matshow(HmapT[ids_tw[0]],vmin=new_min, vmax=new_max)
axs[1,tw].matshow(HmapT[ids_tw[1]],vmin=new_min, vmax=new_max)
axs[2,tw].matshow(HmapT[ids_tw[2]],vmin=new_min, vmax=new_max)
axs[3,tw].matshow(HmapT[ids_tw[3]],vmin=new_min, vmax=new_max)
axs[3,tw].set(xlabel=names_x[tw])
axs[3,tw].xaxis.get_label().set_fontsize(15)
if tw == 0:
axs[0,tw].set(ylabel=r'$CWT \mu$')
axs[0,tw].yaxis.get_label().set_fontsize(15)
axs[1,tw].set(ylabel=r'$CWT \beta$')
axs[1,tw].yaxis.get_label().set_fontsize(15)
axs[2,tw].set(ylabel=r'$CSP \mu$')
axs[2,tw].yaxis.get_label().set_fontsize(15)
axs[3,tw].set(ylabel=r'$CSP \beta$')
axs[3,tw].yaxis.get_label().set_fontsize(15)
#-----------------------------------------------------------------------------
for ax in axs.flat:
ax.label_outer()
for ax in axs.flat:
ax.set_xticks([])
ax.set_yticks([])
#-------------------------------------------------------------------------------
def vis_render(HmapT,new_input,Ntw):
f, ax = plt.subplots(nrows=4, ncols=5, figsize=(12,7.3))
f.subplots_adjust(hspace = 0.1, wspace=.0001)
for tw in range(Ntw):
if tw == 0:
ids_tw = [tw, tw+1, tw+10, tw+10+1]
else:
ids_tw = [tw*2, tw*2+1, tw*2+10, tw*2+10+1]
heatmap_0 = np.uint8(cm.jet(HmapT[ids_tw[0]])[..., :3] * 255)
heatmap_1 = np.uint8(cm.jet(HmapT[ids_tw[1]])[..., :3] * 255)
heatmap_2 = np.uint8(cm.jet(HmapT[ids_tw[2]])[..., :3] * 255)
heatmap_3 = np.uint8(cm.jet(HmapT[ids_tw[3]])[..., :3] * 255)
ax[0,tw].imshow(np.squeeze(new_input[ids_tw[0]]), cmap='gray',vmin=0,vmax=1)
ax[1,tw].imshow(np.squeeze(new_input[ids_tw[1]]), cmap='gray',vmin=0,vmax=1)
ax[2,tw].imshow(np.squeeze(new_input[ids_tw[2]]), cmap='gray',vmin=0,vmax=1)
ax[3,tw].imshow(np.squeeze(new_input[ids_tw[3]]), cmap='gray',vmin=0,vmax=1)
ax[0,tw].imshow(heatmap_0, cmap='jet', alpha=0.5) # overlay
ax[1,tw].imshow(heatmap_1, cmap='jet', alpha=0.5) # overlay
ax[2,tw].imshow(heatmap_2, cmap='jet', alpha=0.5) # overlay
ax[3,tw].imshow(heatmap_3, cmap='jet', alpha=0.5) # overlay
if tw == 0:
ax[0,tw].set(ylabel=r'$CWT \mu$')
ax[0,tw].yaxis.get_label().set_fontsize(15)
ax[1,tw].set(ylabel=r'$CWT \beta$')
ax[1,tw].yaxis.get_label().set_fontsize(15)
ax[2,tw].set(ylabel=r'$CSP \mu$')
ax[2,tw].yaxis.get_label().set_fontsize(15)
ax[3,tw].set(ylabel=r'$CSP \beta$')
ax[3,tw].yaxis.get_label().set_fontsize(15)
for ax in ax.flat:
ax.set_xticks([])
ax.set_yticks([])
#-------------------------------------------------------------------------------
def cnn_network(n_fb,Nkfeats,Ntw,shape_,n_filt,units,l1p,l2p,lrate,sbj):
#---------------------------------------------------------------------------
keras.backend.clear_session()
np.random.seed(123)
tf.compat.v1.random.set_random_seed(123)
#---------------------------------------------------------------------------
input_ = [None]*Ntw*n_fb*Nkfeats
conv_ = [None]*Ntw*n_fb*Nkfeats
pool_ = [None]*Ntw*n_fb*Nkfeats
batch0_ = [None]*Ntw*n_fb*Nkfeats
batch2_ = [None]*Ntw*n_fb*Nkfeats
for i in range(Ntw*n_fb*Nkfeats):
input_[i] = keras.layers.Input(shape=[shape_,shape_,1])
conv_[i] = keras.layers.Conv2D(filters=n_filt,kernel_size=3,strides=1,activation='relu',padding='SAME',input_shape=[shape_,shape_,1])(input_[i])
#-----------------------------------------------------------------------
batch0_[i] = keras.layers.BatchNormalization()(conv_[i])
#-----------------------------------------------------------------------
pool_[i] = keras.layers.MaxPooling2D(pool_size=2)(batch0_[i])
#-----------------------------------------------------------------------
concat = keras.layers.concatenate(pool_)
flat = keras.layers.Flatten()(concat)
#---------------------------------------------------------------------------
batch1 = keras.layers.BatchNormalization()(flat)
hidden1 = keras.layers.Dense(units=units,activation='relu',kernel_regularizer=keras.regularizers.l1_l2(l1=l1p, l2=l2p), kernel_constraint=max_norm(1.))(batch1)#
batch2 = keras.layers.BatchNormalization()(hidden1)
output = keras.layers.Dense(units=2, activation='softmax', kernel_constraint=max_norm(1.))(batch2)#
model = keras.models.Model(inputs=input_, outputs=[output])
#---------------------------------------------------------------------------
learning_rate_fn = tf.keras.optimizers.schedules.PolynomialDecay(lrate, 4000, power=1.0,cycle=False, name=None)
opt = keras.optimizers.Adam(learning_rate=learning_rate_fn)
model.compile(loss='mean_squared_error', optimizer=opt, metrics=['accuracy'])
return model
#-------------------------------------------------------------------------------
# + [markdown] id="jDpoua7-x7lf"
# # Perform GradCAM
# + id="S66cx-qLHdQ9"
#attention maps wide models
#attention maps
from mpl_toolkits.axes_grid1 import make_axes_locatable
def centroid_(X):
D = pairwise_distances(X, X.mean(axis=0).reshape(1,-1))
inertia_ = D.mean()
return np.argmin(D),inertia_
def plot_attention(tmpr_,rel_model_name,layer_name,list_class,figsize=(10,5), transpose=False):
names_feats = [r'CWT-$\mu$-TW1',r'CWT-$\beta$-TW1',r'CWT-$\mu$-TW2',r'CWT-$\beta$-TW2',r'CWT-$\mu$-TW3',r'CWT-$\beta$-TW3',r'CWT-$\mu$-TW4',r'CWT-$\beta$-TW4',r'CWT-$\mu$-TW5',r'CWT-$\beta$-TW5',
r'CSP-$\mu$-TW1',r'CSP-$\beta$-TW1',r'CSP-$\mu$-TW2',r'CSP-$\beta$-TW2',r'CSP-$\mu$-TW3',r'CSP-$\beta$-TW3',r'CSP-$\mu$-TW4',r'CSP-$\beta$-TW4',r'CSP-$\mu$-TW5',r'CSP-$\beta$-TW5']
if transpose:
x_label_list = layer_name
nC = len(list_class)
nl = len(layer_name)
ncols,nrows = tmpr_.shape
y_label_list = []
for ii in range(nC):
y_label_list += str(list_class[ii])
dw = nrows/nl
list_xticks = []
for ii in range(nl):
list_xticks += [int(dw*(0.5+ii))]
dw = ncols/nC
list_yticks = []
for ii in range(nC):
list_yticks += [int(dw*(0.5+ii))]
else:
y_label_list = layer_name
nC = len(list_class)
nl = len(layer_name)
nrows,ncols = tmpr_.shape
x_label_list = []
for ii in range(nC):
x_label_list += str(list_class[ii])
dw = nrows/nl
list_yticks = []
for ii in range(nl):
list_yticks += [int(dw*(0.5+ii))]
dw = ncols/nC
list_xticks = []
for ii in range(nC):
list_xticks += [int(dw*(0.5+ii))]
plt.figure(figsize=figsize)
ax = plt.gca()
im = ax.imshow(tmpr_)
im = ax.imshow(tmpr_)
ax.set_yticks(list_yticks)
ax.set_yticklabels(y_label_list)
ax.set_xticks(list_xticks)
ax.set_xticklabels(names_feats, rotation='vertical')
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.1)
plt.colorbar(im, cax=cax,extend='both',
ticks=[np.round(tmpr_.min(),3), np.round(0.5*(tmpr_.max()-tmpr_.min()),3), np.round(tmpr_.max(),3)])
plt.xticks(rotation=90)
#plt.savefig('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/results/resulting_attention_maps/attention_map_'+str(n_sbj[sbj])+'_'+rel_model_name+'.svg', format='svg')
plt.tight_layout()
plt.show()
import cv2
def attention_wide(modelw,rel_model_name,layer_name,X_train,y_train,
normalize_cam=False,norm_max_min=False,norm_c=True,
plot_int=False,centroid_=False,smooth_samples=20,
smooth_noise=0.20,transpose=False):
#-------------------------------------------------------------------------------
# define trial sample to visualize
# change activations of last layer by linear
replace2linear = ReplaceToLinear()
#relevance model
if rel_model_name == 'Weights':
#[topo_avg_muT_cwt,topo_avg_beT_cwt,topo_avg_muT_csp,topo_avg_beT_csp]
path='/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/results/matrix_data/WeightsRel_sbj_'+str(n_sbj[sbj])+'_fold_'+str(opt_fld[sbj])+'.pickle'
with open(path, 'rb') as f:
w_data = pickle.load(f)
for i in range(5):
if i ==0:
amw_cwt = cv2.resize(w_data[0][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)
else:
amw_cwt = np.c_[amw_cwt,cv2.resize(w_data[0][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)]
amw_cwt = np.c_[amw_cwt,cv2.resize(w_data[1][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)]
for i in range(5):
if i ==0:
amw_csp = cv2.resize(w_data[2][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)
else:
amw_csp = np.c_[amw_csp,cv2.resize(w_data[2][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)]
amw_csp = np.c_[amw_csp,cv2.resize(w_data[3][i,:,:],(40, 40),interpolation = cv2.INTER_NEAREST)]
amw = np.concatenate((amw_cwt,amw_csp),axis=1)
amw = np.r_[amw,amw]
relM = [None]*len(np.unique(y_train))
#---------------------------------------------------------------------------
tmpr = amw/(1e-8+amw.max())
#---------------------------------------------------------------------------
else:
if rel_model_name == 'Gradcam':
gradcamw = Gradcam(modelw,
model_modifier=replace2linear,
clone=True)
elif rel_model_name == 'Gradcam++':
gradcamw = GradcamPlusPlus(modelw,
model_modifier=replace2linear,
clone=True)
elif rel_model_name == 'Scorecam':
scorecamw = Scorecam(modelw)
elif rel_model_name == 'Saliency':
saliencyw = Saliency(modelw,
model_modifier=replace2linear,
clone=True)
layer_name = [''] #saliency doesn't depend on different layers
nC = len(np.unique(y_train))
relM = [None]*nC
if type(X_train)==list:
n_inputs = len(X_train)
new_input = [None]*n_inputs
for c in range(len(np.unique(y_train))):
id_sample = y_train == np.unique(y_train)[c]
if (type(X_train)==list) and (rel_model_name != 'Saliency'):
relM[c] = np.zeros((sum(id_sample),X_train[0].shape[1],X_train[0].shape[2],len(layer_name)))
#print(1,relM[c].shape)
elif (type(X_train)==list) and (rel_model_name == 'Saliency'):
relM[c] = np.zeros((sum(id_sample),X_train[0].shape[1],X_train[0].shape[2],len(X_train)))
#print(2,relM[c].shape)
else:
relM[c] = np.zeros((sum(id_sample),X_train.shape[1],X_train.shape[2],len(layer_name)))
#print(3,relM[c].shape)
score = CategoricalScore(list(y_train[id_sample])) #-> [0] para probar a una clase diferente
if type(X_train)==list:
for ni in range(n_inputs):
new_input[ni] = X_train[ni][id_sample]
else:
new_input = X_train[id_sample]
#print('rel',rel_model_name,'layer',layer_name[l])
for l in range(len(layer_name)):
#print(rel_model_name,'class', np.unique(y_train)[c],'layer',layer_name[l])
# label score -> target label accoring to the database
#-----------------------------------------------------------------------------
# generate heatmap with GradCAM
if (rel_model_name == 'Gradcam') or (rel_model_name == 'Gradcam++'):
rel = gradcamw(score,
new_input,
penultimate_layer=layer_name[l], #layer to be analized
expand_cam=True,
normalize_cam=normalize_cam)
elif rel_model_name == 'Saliency': #saliency map is too noisy, so let’s remove noise in the saliency map using SmoothGrad!
rel = saliencyw(score, new_input,smooth_samples=smooth_samples,
smooth_noise=smooth_noise,
normalize_map=normalize_cam) #, smooth_samples=20,smooth_noise=0.20) # The number of calculating gradients iterations.
elif rel_model_name == 'Scorecam':
rel = scorecamw(score, new_input, penultimate_layer=layer_name[l], #layer to be analized
expand_cam=True,
normalize_cam=normalize_cam) #max_N=10 -> faster scorecam
#save model
if rel_model_name != 'Saliency':
if type(X_train)==list:
tcc = rel[0]
else:
tcc = rel
dimc = tcc.shape
tccv = tcc.ravel()
tccv[np.isnan(tccv)] = 0
tcc = tccv.reshape(dimc)
if norm_max_min: #normalizing along samples
tcc = MinMaxScaler().fit_transform(tcc.reshape(dimc[0],-1).T).T
tcc = tcc.reshape(dimc)
relM[c][...,l] = tcc
if l==0:
tmp = np.median(relM[c][...,l],axis=0)#relM[c][...,l].mean(axis=0)
else:
if transpose:
tmp = np.c_[tmp,np.median(relM[c][...,l],axis=0)]#np.r_[tmp,relM[c][...,l].mean(axis=0)] #centroid
else:
tmp = np.r_[tmp,np.median(relM[c][...,l],axis=0)]#np.r_[tmp,relM[c][...,l].mean(axis=0)] #centroid
else: #saliency
if type(X_train)==list:
tcc = np.zeros((rel[0].shape[0],rel[0].shape[1],rel[0].shape[2],len(rel)))
for ii in range(len(rel)):
tcc[...,ii] = rel[ii]
else:
tcc = rel
dimc = tcc.shape
tccv = tcc.ravel()
tccv[np.isnan(tccv)] = 0
tcc = tccv.reshape(dimc)
if norm_max_min: #normalizing along samples
tcc = MinMaxScaler().fit_transform(tcc.reshape(dimc[0],-1).T).T
tcc = tcc.reshape(dimc)
relM[c] = tcc
if type(X_train)==list:
tmp = np.median(tcc[...,0],axis=0)
for ii in range(len(rel)-1):
if transpose:
tmp = np.c_[tmp,np.median(tcc[...,ii+1],axis=0)]
else:
tmp = np.r_[tmp,np.median(tcc[...,ii+1],axis=0)]
else:
tmp = np.median(tcc,axis=0)
if norm_c: #normalizing along layers
tmp = tmp/(1e-8+tmp.max())
if c==0:
tmpr = tmp
else:
if transpose:
tmpr = np.r_[tmpr,tmp]
else:
tmpr = np.c_[tmpr,tmp]
#print(tmp.shape,tmp.max())
if plot_int: #plot every class
plt.imshow(tmp)
plt.colorbar(orientation='horizontal')
plt.axis('off')
plt.show()
#---------------------------------------------------------------------------
tmpr = tmpr/(1e-8+tmpr.max())
#---------------------------------------------------------------------------
list_class = np.unique(y_train)
plot_attention(tmpr,rel_model_name,layer_name,list_class,transpose=transpose)
return relM,tmpr
# + id="wE35xUY6sVw8"
#-------------------------------------------------------------------------------
# define parameters
partitions = ['train','valid','test']
names_x = [r'-1.5s-0.5s',r'$-0.5s-1.5s$',r'$0.5s-2.5s$',r'$1.5s-3.5s$',r'$2.5s-4.5s$']
learning_rate = 1e-4
th_name = np.array([[-1.5, 0.5],[-0.5, 1.5],[0.5, 2.5],[1.5, 3.5],[2.5, 4.5]])
n_fb = 2
Ntw = 5
Nkfeats = 2
num_classes = 2
n_filt = 2
n_fld = 3
n_conv_layers = 20
#-------------------------------------------------------------------------------
n_sbj = [41]#[1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,30,31,32,33,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52]
opt_neurons = [200]#[100,200,100,300,200,100,300,200,100,100,200,200,200,300,300,100,100,100,300,200,300,300,200,100,100,300,200,300,300,200,300,200,300,300,100,200,300,300,200,100,200,200,100,300,300,100,100,300,100,300]
opt_l1 = [0.005]#[0.0005,0.0005,0.005,0.005,0.001,0.001,0.0005,0.0005,0.0005,0.005,0.005,0.005,0.005,0.005,0.005,0.0005,0.0005,0.001,0.0005,0.0005,0.0005,0.0005,0.005,0.001,0.001,0.005,0.005,0.0005,0.0005,0.001,0.005,0.001,0.001,0.005,0.005,0.001,0.005,0.005,0.005,0.001,0.005,0.0005,0.005,0.005,0.0005,0.005,0.0005,0.005,0.005,0.005]
opt_l2 = [0.0005]#[0.005,0.001,0.0005,0.005,0.005,0.001,0.0005,0.005,0.005,0.001,0.005,0.005,0.001,0.005,0.001,0.001,0.005,0.0005,0.0005,0.0005,0.0005,0.005,0.0005,0.005,0.005,0.0005,0.001,0.0005,0.0005,0.005,0.0005,0.005,0.0005,0.005,0.005,0.001,0.0005,0.001,0.0005,0.005,0.001,0.0005,0.001,0.0005,0.005,0.001,0.001,0.005,0.0005,0.001]
opt_fld = [1]#[3,1,1,3,3,1,2,2,3,2,1,2,1,2,1,3,2,1,2,1,1,1,3,1,1,2,1,3,3,1,1,2,2,1,1,3,2,1,1,3,2,2,3,1,2,1,1,3,1,1]
#-------------------------------------------------------------------------------
for sbj in range(len(n_sbj)):
print('subject ', n_sbj[sbj])
#-----------------------------------------------------------------------------
# load data train/test trough all tw
XF_train_cwt = []
XF_train_csp = []
XF_test_cwt = []
XF_test_csp = []
for i in range(th_name.shape[0]):
X_train_re_cwt, X_train_re_csp, X_test_re_cwt, X_test_re_csp, y_trainF, y_testF = TW_data(n_sbj[sbj],th_name[i,0],th_name[i,1])
XF_train_cwt.append(X_train_re_cwt)
XF_train_csp.append(X_train_re_csp)
XF_test_cwt.append(X_test_re_cwt)
XF_test_csp.append(X_test_re_csp)
#-----------------------------------------------------------------------------
# partition of data
XT_train, XT_valid, XT_test, y_train, y_valid, y_test, train_index, valid_index = norm_data(XF_train_cwt, XF_train_csp, XF_test_cwt, XF_test_csp, n_fb, Ntw, y_trainF, y_testF, opt_fld[sbj]-1)
#-----------------------------------------------------------------------------
# define model
model = cnn_network(n_fb,Nkfeats,Ntw,40,n_filt,opt_neurons[sbj],opt_l1[sbj],opt_l2[sbj],learning_rate,n_sbj[sbj])
#-----------------------------------------------------------------------------
tf.keras.utils.plot_model(model)
#-----------------------------------------------------------------------------
# loading best model weights
filepath = '/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/results/parameter_setting/weights_sbj_'+str(n_sbj[sbj])+'_filters_2_units_'+str(int(opt_neurons[sbj]))+'_l1_'+str(opt_l1[sbj])+'_l2_'+str(opt_l2[sbj])+'_fld_'+str(opt_fld[sbj])+'.hdf5'
checkpoint_path = filepath
model.load_weights(checkpoint_path)
#-----------------------------------------------------------------------------
rel_model_name = ['Gradcam++','Scorecam','Saliency'] #,'Gradcam++','Scorecam','Saliency'
layer_name = ['conv2d','conv2d_1','conv2d_2','conv2d_3','conv2d_4','conv2d_5','conv2d_6','conv2d_7','conv2d_8','conv2d_9','conv2d_10',
'conv2d_11','conv2d_12','conv2d_13','conv2d_14','conv2d_15','conv2d_16','conv2d_17','conv2d_18','conv2d_19']
#
print('norm_c = False')
relM_ = [None]*len(rel_model_name) #relM[m] -> number classes x input image resolution x number of layers
tmpr_ = [None]*len(rel_model_name)
for m in range(len(rel_model_name)):
relM_[m],tmpr_[m] = attention_wide(model,rel_model_name[m],layer_name,XT_train,np.argmax(y_train,axis=1),
norm_c=False,norm_max_min=False,plot_int=False,transpose=True)
#-----------------------------------------------------------------------------
with open('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/results/resulting_attention_maps/score_attmaps_'+str(n_sbj[sbj])+'.pickle', 'wb') as f:
pickle.dump([relM_, tmpr_], f)
#-----------------------------------------------------------------------------
del model
#-----------------------------------------------------------------------------
| Experimental/DW_LCAM/[3]_Main_attention_maps_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import azureml
from azureml.core import Workspace, Run
# check core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
# -
# # Workspace
# 1. Get workspace associated with folder
# +
# use this code to set up config file
#subscription_id ='<SUB ID>'
#resource_group ='<RESOURCE>'
#workspace_name = '<WS NAME>'
#try:
# ws = Workspace(subscription_id = subscription_id, resource_group = resource_group, workspace_name = workspace_name)
# ws.write_config()
# print('Workspace configuration succeeded. You are all set!')
#except:
# print('Workspace not found. TOO MANY ISSUES!!!')
ws = Workspace.from_config()
# -
# # DataStores
# 1. Getting handle to specific folders on datastores
ds = ws.get_default_datastore()
ml20m = ds.upload(src_dir='data/ml-20m', target_path='ml-20m', show_progress=True)
mllatest = ds.upload(src_dir='data/ml-latest', target_path='ml-latest', show_progress=True)
mllatestsmall = ds.upload(src_dir='data/ml-latest-small', target_path='ml-latest-small', show_progress=True)
# # Experiments
# 1. Running experiments
| control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="eZzc7rgciISm" slideshow={"slide_type": "slide"}
# [](http://introml.analyticsdojo.com)
# <center><h1>Introduction to Python - Null Values</h1></center>
# <center><h3><a href = 'http://introml.analyticsdojo.com'>introml.analyticsdojo.com</a></h3></center>
#
#
#
# -
# # Null Values
# + [markdown] colab_type="text" id="LI0DdIVbiISs"
# ## Running Code using Kaggle Notebooks
# - Kaggle utilizes Docker to create a fully functional environment for hosting competitions in data science.
# - You could download/run this locally or run it online.
# - Kaggle has created an incredible resource for learning analytics. You can view a number of *toy* examples that can be used to understand data science and also compete in real problems faced by top companies.
# + colab={"base_uri": "https://localhost:8080/", "height": 481} colab_type="code" id="RyxIdHyjiP6a" outputId="9af3bb5f-7aa2-4cf2-b17e-6b139f9a8dc4"
# !wget https://raw.githubusercontent.com/rpi-techfundamentals/spring2019-materials/master/input/train.csv
# !wget https://raw.githubusercontent.com/rpi-techfundamentals/spring2019-materials/master/input/test.csv
# + [markdown] colab_type="text" id="A-Fh-RZBiISy"
# ### Null Values Typical When Working with Real Data
# - Null values `NaN` in Pandas
#
# + colab={} colab_type="code" id="4IGd7HVtiIS6"
import numpy as np
import pandas as pd
# Input data files are available in the "../input/" directory.
# Let's input them into a Pandas DataFrame
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 248} colab_type="code" id="Iv4AOq8ziITJ" outputId="d1440faa-df91-449b-8eb7-958c6676e20c"
print(train.dtypes)
# + colab={"base_uri": "https://localhost:8080/", "height": 524} colab_type="code" id="ecKJOpkliITW" outputId="1951a7b0-d700-4d70-c7a5-782309c56125"
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 470} colab_type="code" id="hDczrXrhiITd" outputId="533994ae-55ee-40b7-cc97-70918d2cfcf2"
test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 457} colab_type="code" id="KpnKEoYwiITm" outputId="0d6aab9c-c658-410c-9d42-7ecea4d9dd49"
#Let's get some general s
totalRows=len(train.index)
print("There are ", totalRows, " so totalRows-count is equal to missing variables.")
print(train.describe())
print(train.columns)
# + colab={} colab_type="code" id="q1Q7R1_hiITr" outputId="d65cc20f-bbb1-4c23-c01d-22bb4af0a6fc"
# We are going to do operations on thes to show the number of missing variables.
train.isnull().sum()
# + [markdown] colab_type="text" id="_FPiVaCjiITv"
# ### Dropping NA
# - If we drop all NA values, this can dramatically reduce our dataset.
# - Here while there are 891 rows total, there are only 183 complete rows
# - `dropna()` and `fillna()` are 2 method for dealing with this, but they should be used with caution.
# - [Fillna documentation](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.fillna.html)
# - [Dropna documentation](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.dropna.html)
# + colab={} colab_type="code" id="MGjx1_ZWiITw" outputId="c3860e1f-8a59-4067-f63c-bfc2f956bb88"
# This will drop all rows in which there is any missing values
traindrop=train.dropna()
print(len(traindrop.index))
print(traindrop.isnull().sum())
# + colab={} colab_type="code" id="L0VUTscAiIT2" outputId="fbae90b4-94bf-462f-e5c4-e31b42153087"
# This will drop all rows in which there is any missing values
trainfill=train.fillna(0) #This will just fill all values with nulls. Probably not what we want.
print(len(trainfill.index))
print(traindrop.isnull().sum())
# forward-fill previous value forward.
train.fillna(method='ffill')
# forward-fill previous value forward.
train.fillna(method='bfill')
# + [markdown] colab_type="text" id="obU9f8kAiIT7"
# ### Customized Approach
# - While those approaches
# + colab={} colab_type="code" id="zH7RPahYiIT7" outputId="87e4f9ee-ee46-4537-b9fd-ef2fc5953ed6"
average=train.Age.mean()
print(average)
# + colab={} colab_type="code" id="T1nJxhAkiIUA" outputId="553ae041-916c-486e-c6ed-1e9df27b4e13"
#Let's convert it to an int
average= int(average)
average
# + colab={} colab_type="code" id="W56l1aeIiIUF" outputId="1b537ef9-09ce-4a75-c82d-d6cb11484f44"
#This will select out values that
train.Age.isnull()
# + colab={} colab_type="code" id="pm4cVUJ0iIUW" outputId="fd061961-a4ff-454f-fdc0-e42296d16fb6"
#Now we are selecting out those values
train.loc[train.Age.isnull(),"Age"]=average
train
# + [markdown] colab_type="text" id="AUsTs2IRiIUZ"
# ### More Complex Models - Data Imputation
# - Could be that Age could be inferred from other variables, such as SibSp, Name, Fare, etc.
# - A next step could be to build a more complex regression or tree model that would involve data tat was not null.
# + [markdown] colab_type="text" id="f-fI5Wf3iIUa"
# ### Missing Data - Class Values
# - We have 2 missing data values for the Embarked Class
# - What should we replace them as?
#
# + colab={} colab_type="code" id="7aWsSUzWiIUb" outputId="2327a774-e2b6-4adc-8d17-db06ef117b61"
pd.value_counts(train.Embarked)
# + colab={} colab_type="code" id="5YdotE8hiIUl" outputId="b798e273-4fa8-4193-a976-4dbd4b1f3e8c"
train.Embarked.isnull().sum()
# + colab={} colab_type="code" id="I5vZwZFAiIUp" outputId="4d0e6be0-aacd-4306-94bd-7c2e0b9e3ea6"
train[train.Embarked.isnull()]
# + colab={} colab_type="code" id="i1g2ZO3ciIUr"
train.loc[train.Embarked.isnull(),"Embarked"]="S"
# + [markdown] colab_type="text" id="Rhg68-uSiIUt"
#
#
# This work is licensed under the [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0/) license agreement.
# Adopted from [materials](https://github.com/phelps-sg/python-bigdata) Copyright [<NAME>](http://sphelps.net) 2014
| site/_build/jupyter_execute/notebooks/02-intro-python/03-null-values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.decomposition import NMF
from sklearn.metrics.pairwise import cosine_similarity
from sqlalchemy import create_engine
import pickle
#os.getenv('SQLALCHEMY_URL')
engine = create_engine('postgresql+psycopg2://postgres:postgres@localhost:5432/movies', echo=False)
engine
df_tags=pd.read_csv('tags.csv')
df_tags
df_movies=pd.read_csv('movies.csv')
df_movies2 = df_movies.join(df_movies['genres'].str.split('|', expand=True).add_prefix('genre'))
df_movies2 = pd.DataFrame(df_movies2.set_index(['movieId', 'title', 'genres']).stack())
df_movies2.reset_index(inplace=True)
df_movies2.drop('level_3',1, inplace=True)
df_movies2=df_movies2.rename(columns={0:'genre'})
df_movies2.sort_values(['movieId','genre'], ascending=[True,True])
df_movies2 = df_movies2.join(pd.get_dummies(df_movies2['genre'], drop_first=False))
df_movies2.head()
df_ratings=pd.read_csv('ratings.csv')
df_ratings
df = df_ratings.merge(df_movies, on='movieId', how='left')
df['title'] = df['title'].map(lambda x: str(x)[:-7])
df=df.drop_duplicates(subset='title')
df
df.to_sql('movies', engine, if_exists='replace', method='multi', chunksize=10000)
df = pd.read_sql('movies', engine, index_col=0)
df.tail()
reviews = pd.pivot_table(df, values='rating', index='userId', columns='title')
reviews
mean_values = round(reviews.mean().mean(), 1)
reviews = reviews.fillna(mean_values)
reviews
# **NMF**
# n_components - between 5 and 20. If too much - slower model, overfit
nmf = NMF(n_components = 10)
nmf.fit(reviews)
pickle.dump(nmf, open('nmf_model.pkl', 'wb'))
loaded_model = pickle.load(open('nmf_model.pkl', 'rb'))
nmf_q = loaded_model.components_
Q = nmf_q
P = nmf.transform(reviews)
P.shape, Q.shape
Q = nmf.components_
P = nmf.transform(reviews)
P.shape, Q.shape
Rhat = np.dot(P,Q)
predictions = pd.DataFrame(Rhat, columns=reviews.columns, index = reviews.index)
predictions.head()
nmf.reconstruction_err_
# sample movie
film = reviews.columns[np.random.randint(len(reviews.columns))]
# sample rating
rating = 3
user_input = (film, rating)
user_input
# +
# single movie input
#query = query.fillna(mean_values)
# +
#query = np.zeros(len(reviews.columns))
#list(reviews.columns).index(film)
# +
#query.reshape(-1,1).shape
# +
#query[list(reviews.columns).index(film)] = rating
# -
# multiple movies input
user_movies = 'Toy Story', 'Zulu', 'Titanic'
query = np.zeros(len(reviews.columns))
for i in range(len(user_movies)):
query[list(reviews.columns).index(user_movies_matched[i])] = user_ratings[i]
dfdf=pd.DataFrame()
dfdf['query']=query
dfdf['query'].unique()
new_query=query.reshape(-1,1).T
new_p = nmf.transform(new_query)
new_p.shape
new_prediction = np.dot(new_p,Q)
pd.DataFrame(new_prediction, columns=reviews.columns)
# Movies suggestion
reviews.columns[np.argsort(new_prediction)[0][-5:-1]]
| MNF_movie_recommender/mnf_movie_recommender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
print("hello")
print ("my first python notebook")
from math import log
from __future__ import division
def entropy(x,y):
px = x/(x+y)
py = y/(x+y)
entr = (-px*log(px,2)-py*log(py,2))
print "prob x {},y {}, entr {}".format(px,py,entr)
entropy(3,1)
# +
def cal(pi,h,t,ma,mb):
return [pi* pow(ma,h)*pow(1-ma,t), (1-pi)* pow(mb,h)*pow(1-mb,t)]
def norm(raw):
return [round(float(i)/sum(raw),3) for i in raw]
def ua(a1,a2,a3):
return round((2*a1+4*a2+0*a3)/(4*(a1+a2+a3)),4)
# -
raw = cal(pi=.569,h=0,t=4,ma=.7229,mb=.2057)
print raw
print norm(raw)
print round(.6038,3)
print ua(.335,.005,.981)
| notebooks/CS-6220/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="odNA4QeNLIZ1" colab={"base_uri": "https://localhost:8080/"} outputId="8d8b85c4-d4dd-4d24-ccea-3a67778ac899"
# !unzip Data-Images.zip
# + [markdown] id="cL3rXkEu8-cJ"
# # Imports
# + id="WYuZjnLS8-cK"
import requests
import random
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from PIL import Image
import matplotlib.pyplot as plt
import urllib
import cv2
import json
# + id="tDVaJWZEgQSQ" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b0dce0f5-ad04-4ab6-d344-138a57b211b2"
pwd
# + [markdown] id="MjQXdGWn8-cN"
# # Load the `.json` file containing the download paths of the images
# + id="1KaenEw48-cO" colab={"base_uri": "https://localhost:8080/", "height": 453} outputId="1e3e1e0f-7390-4c90-846a-522425e66088"
# read the data
data = pd.read_json('Indian_Number_plates.json',lines=True)
pd.set_option('display.max_colwidth', -1)
# delete the extras column
del data['extras']
# check the data dataframe
data.head()
# + id="YmpU7vqX8-cS" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="bf4698a2-4ce7-46c0-bb26-2f8798caf9bc"
# Extract the points of the bounding boxes because thats what we want
data['points'] = data.apply(lambda row: row['annotation'][0]['points'], axis=1)
data['height'] = data.apply(lambda row: row['annotation'][0]['imageHeight'], axis=1)
data['width'] = data.apply(lambda row: row['annotation'][0]['imageWidth'], axis=1)
# And drop the rest of the annotation info
del data['annotation']
data.head()
# + id="wXgSwvpz8-cW"
Images = []
Plates = []
Points = []
def downloadTraining(df):
for index, row in df.iterrows():
# Get the image from the URL
resp = urllib.request.urlopen(row[0])
im = np.array(Image.open(resp)) # This is a numpy 3D array
# We append the image to the training input array
Images.append(im)
# Points of rectangle
x_point_top = row[1][0]['x']*im.shape[1]
y_point_top = row[1][0]['y']*im.shape[0]
x_point_bot = row[1][1]['x']*im.shape[1]
y_point_bot = row[1][1]['y']*im.shape[0]
Points.append([int(x_point_top), int(y_point_top),int(x_point_bot), int(y_point_bot)])
# Cut the plate from the image and use it as output
carImage = Image.fromarray(im)
plateImage = carImage.crop((x_point_top, y_point_top, x_point_bot, y_point_bot))
Plates.append(np.array(plateImage))
downloadTraining(data)
# + id="Mw_796_YCp__" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e771cd3c-26de-4dcc-fbdd-40eaa42559c9"
len(Points)
# + [markdown] id="GKPgTJNK8-cY"
# # To check the x_min,y_min,x_max,y_max
# + id="IlctY5ac8-cZ" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="cf580bc9-875e-4210-d781-3beec765e4e2"
# This gives the format of the annotations
Points[:10]
# + [markdown] id="Cp99wvdMLLe7"
# # Dataframe construction
# + id="RmjOXw-yEO4F" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="d673fbf9-cf1c-4052-937d-7fe3e5a70011"
main_df = pd.DataFrame([row for row in Points], columns=['xmin', 'ymin', 'xmax', 'ymax'])
main_df.head()
# + [markdown] id="NC7s4dqILQdp"
# # The great (not really) filename generation hack
# + id="vJeIRFoHF2Dl" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="30f88baf-4a82-45f4-a091-f47033b2da1c"
images_path = 'Data-Images/Cars/'
give_me_236 = [images_path + str(i) + '.jpg' for i in list(range(0, 237))]
give_me_236[:10]
# + id="NX5AGKc2GjVg" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="b2e7590d-66f5-414d-cb0d-4445318af49a"
main_df['filename'] = give_me_236
main_df.head(10)
# + id="ntPjwW7gGtX6" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="e4306d93-4eef-44fe-822f-7201f1ae6a06"
main_df['width'] = main_df['xmax'] - main_df['xmin']
main_df['height'] = main_df['ymax'] - main_df['ymin']
main_df.head()
# + id="PfrihGPvHejR" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="de68d373-c028-45d2-ad27-f45110fcd4d7"
modified_df = pd.DataFrame()
modified_df['filename'] = main_df['filename']
modified_df['width'] = main_df['width']
modified_df['height'] = main_df['height']
modified_df['class'] = 'license_plate'
modified_df['xmin'] = main_df['xmin']
modified_df['ymin'] = main_df['ymin']
modified_df['xmax'] = main_df['xmax']
modified_df['ymax'] = main_df['ymax']
modified_df.head()
# + [markdown] id="bGTewHBILWXf"
# Booh yeah! We have the data in the right format (for TensorFlow Object Detection API).
# + [markdown] id="ko-OkesnLcYQ"
# # Splits
# + id="QnxTRPPzIfUN"
# 95% for train
train_labels = modified_df.sample(frac=0.95)
modified_df.drop(train_labels.index, axis=0, inplace=True)
# 5% for test
test_labels = modified_df
# + id="dk1ZD7T9JJZT" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="10a018af-732c-409e-9b9f-d7da5a19ca89"
train_labels.head()
# + id="c93xQWkSJQig" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="b2b9fb62-0ec1-4da3-dfcc-1c7b6f7a72bb"
test_labels.head()
# + id="dADIiZhZJUs7" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f4c97a0a-4f1b-4897-8e0e-bd18dfab9ef8"
train_labels.shape
# + id="GxefIS_8JYsJ" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="df7c46a4-49fc-48d1-c648-e54d22d6e9f6"
test_labels.shape
# + id="ryqhUe1LhhhC" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a35b0950-521f-41fc-cac6-644befe22952"
pwd
# + [markdown] id="V45vOFb_LfRI"
# # Manual inspection of the bounding boxes
# + id="1kuthQb-JjxJ"
image_1 = cv2.imread('136.jpg')
image_1 = cv2.cvtColor(image_1,cv2.COLOR_BGR2RGB)
# + id="2eUEg46WJyut" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ae6b3bfd-571c-47c6-9d03-c8c1e6e2a682"
# 155 200 328 274
cv2.rectangle(image_1, (155, 200), (328, 274), (255,0,0), 2)
cv2.imwrite('rectangle.png', image_1)
# + id="lRop8P86KH8f" colab={"base_uri": "https://localhost:8080/", "height": 268} outputId="82b36ac7-72ee-41c7-edee-90ea20099fa0"
image = plt.imread('rectangle.png')
plt.imshow(image)
plt.show()
# + [markdown] id="xOGZToutLiTn"
# # Serialize the dataframes
# + id="YEYA_2kdDe_T"
train_labels.to_csv('train_labels.csv', index=False)
test_labels.to_csv('test_labels.csv', index=False)
| Bounding_Box_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
import numpy as np
np.set_printoptions(suppress=True)
digits = load_digits()
X, y = digits.data, digits.target
X_train, X_test, y_train, y_test = train_test_split(X, y)
# -
# Removing mean and scaling variance
# ===================================
from sklearn.preprocessing import StandardScaler
# 1) Instantiate the model
scaler = StandardScaler()
# 2) Fit using only the data.
scaler.fit(X_train)
# 3) `transform` the data (not `predict`).
X_train_scaled = scaler.transform(X_train)
X_train.shape
X_train_scaled.shape
X_train.mean(axis=0)
# The transformed version of the data has the mean removed:
X_train_scaled.mean(axis=0)
X_train_scaled.std(axis=0)
X_test_transformed = scaler.transform(X_test)
X_test_transformed.mean(axis=0)
# Principal Component Analysis
# =============================
# 0) Import the model
from sklearn.decomposition import PCA
# 1) Instantiate the model
pca = PCA(n_components=2)
# 2) Fit to training data
pca.fit(X)
# 3) Transform to lower-dimensional representation
print(X.shape)
X_pca = pca.transform(X)
X_pca.shape
# Visualize
# ----------
# +
import matplotlib.pyplot as plt
# %matplotlib notebook
plt.rcParams["figure.dpi"] = 200
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=plt.cm.Vega10(y))
# -
# Manifold Learning
# ==================
from sklearn.manifold import Isomap
isomap = Isomap()
X_isomap = isomap.fit_transform(X)
plt.figure()
plt.scatter(X_isomap[:, 0], X_isomap[:, 1], c=plt.cm.Vega10(y))
# # Exercises
# Visualize the digits dataset using the TSNE algorithm from the sklearn.manifold module (it runs for a couple of seconds).
#
# +
# # %load solutions/digits_tsne.py
| notebooks/02 - Unsupervised Transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tigress data exclusive processing
# This is only necessary for the CMIP data, which resides on the princeton machine
import numpy as np
import xarray as xr
import xesmf as xe
from glob import glob
from xarrayutils import concat_dim_da
# %matplotlib inline
# +
def fetch_equator_full(ds, field='o2'):
ds_target = xr.Dataset({'lat': (['lat'], np.arange(-5, 5, 0.5)),
'lon': (['lon'], np.arange(-270, -60, 1)),
}
)
ds = ds.copy()
print(ds.attrs['model_id'])
if len(ds.lon.shape) == 1:
lon, lat = np.meshgrid(ds.lon, ds.lat)
ds = ds.rename({'lon':'rlon', 'lat':'rlat'})
llon = xr.DataArray(lon, dims=['rlat', 'rlon'])
llat = xr.DataArray(lat, dims=['rlat', 'rlon'])
ds.coords['lon'] = llon
ds.coords['lat'] = llat
regridder = xe.Regridder(ds, ds_target, 'bilinear')
out = regridder(ds[field])
regridder.clean_weight_file()
return out
# So this is not ideal..I need to rename the coords to dimensions and then average over 1S-1N
# Check if all files are there
def filecheck(path):
li = glob(path)
if len(li) == 1:
return li[0]
elif len(li) == 0:
raise RuntimeError('No files found for %s' %(path))
else:
raise RuntimeError('Multiple files found: %s' %li)
def replace_time(ds, reftime):
ds = ds.copy()
# if len(ds.time)>=len(reftime):
# ds['time'].data = reftime[0:len(reftime)]
# else:
ds['time'].data = reftime[0:len(ds.time)]
# print(ds.time[0])
return ds
# +
path = '/tigress/laurer/CMIP5'
for scenario in ['piControl', 'rcp85', ]:
print('SCENARIO:%s' %scenario)
opath = '../data/processed/CMIP5_tropical_combined_%s.nc' %scenario
filelist = glob(path+'/cmip*')
models = list(set([a.replace(path,'').split('.')[3] for a in filelist]))
# there is no piControl fir MRI
if scenario == 'piControl':
models = [m for m in models if not 'MRI' in m]
# for now I am missing the HadGEM and Miroc
models = [m for m in models if ('HadGEM' not in m) and ('MIROC' not in m)]
uo_files = [filecheck('%s/cmip5.merge.*.%s.%s.yr.*uo*nc' %(path, m, scenario)) for m in models]
o2_files = [filecheck('%s/cmip5.merge.*.%s.%s.yr.*o2*nc' %(path, m, scenario)) for m in models]
raw_uo_datasets = [xr.open_mfdataset(path) for path in uo_files]
raw_o2_datasets = [xr.open_mfdataset(path) for path in o2_files]
############ interpolate all datasets onto reference model (take one 300 yr one)
# time_ref = uo_datasets['IPSL-CM5A-LR'].time
# Ok this is HELL! I will replace all calendars assuming they start in 2006 and represent yearly averages
# I CANNOT SAVE THIS AAAEEERRRHGGGHFGHGHGHFGHGHGHGHG so i will restrict it to a valid
time_ref = xr.cftime_range(start='2006', end='2301', freq='AS-JUL', calendar='standard')#.to_datetimeindex()
# parse to dict
uo_datasets = {k:v for k,v in zip(models, raw_uo_datasets)}
o2_datasets = {k:v for k,v in zip(models, raw_o2_datasets)}
# precut the weird french model grid
if 'IPSL-CM5A-LR' in o2_datasets.keys():
o2_datasets['IPSL-CM5A-LR'] = o2_datasets['IPSL-CM5A-LR'].sel(y=slice(0,85))
uo_datasets['IPSL-CM5A-LR'] = uo_datasets['IPSL-CM5A-LR'].sel(y=slice(0,85))
if 'MRI-ESM1' in o2_datasets.keys():
o2_datasets['MRI-ESM1'] = o2_datasets['MRI-ESM1'].sel(y=slice(0,250))
uo_datasets['MRI-ESM1'] = uo_datasets['MRI-ESM1'].sel(y=slice(0,250))
############# interpolate everything on a common vertical grid
ref_lev = o2_datasets['NorESM1-ME'].lev
o2_datasets = {k:replace_time(v.interp(lev=ref_lev),
time_ref) for k,v in o2_datasets.items()}
uo_datasets = {k:replace_time(v.interp(lev=ref_lev),
time_ref) for k,v in uo_datasets.items()}
# Process the datasets:
uo_datasets_processed = {kk:fetch_equator_full(dd.load(), field='uo') for kk, dd in uo_datasets.items()}
o2_datasets_processed = {kk:fetch_equator_full(dd.load(), field='o2') for kk, dd in o2_datasets.items()}
datasets_combined = {k:xr.Dataset({'o2':o2_datasets_processed[k].copy().interp(lev=ref_lev),
'uo':uo_datasets_processed[k].copy().interp(lev=ref_lev)}) for k in o2_datasets_processed.keys()}
mod_names = list(datasets_combined.keys())
mod_ds = [datasets_combined[k] for k in mod_names]
ds = xr.concat(mod_ds, concat_dim_da(mod_names, 'model'))
print(ds)
ds.attrs['scenario'] = scenario
ds.o2.isel(time=10, lev=12).plot(col='model', col_wrap=4)
ds.to_netcdf(opath)
| notebooks/jbusecke_00a_Buseckeetal_GRL_processing_tigress.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading NeuroArch Database with Hemibrain Dataset v1.1
# This tutorial provides code to load NeuroArch database with Hemibrain Dataset v1.1. Requirement before running the notebook:
# - Installed [NeuroArch](https://github.com/fruitflybrain/neuroarch), [OrientDB Community Version](https://www.orientdb.org/download), and [pyorient](https://github.com/fruitflybrain/pyorient). The [NeuroNLP Docker image](https://hub.docker.com/r/fruitflybrain/neuronlp) and [FlyBrainLab Docker image](https://hub.docker.com/r/fruitflybrain/fbl) all have a copy of the software requirement ready.
# - Installed [PyMeshLab](https://pypi.org/project/pymeshlab/).
# - Installed [neuprint-python](https://github.com/connectome-neuprint/neuprint-python).
# - Download the [Neuprint database dump for the Hemibrain dataset v1.1](https://storage.cloud.google.com/hemibrain/v1.1/exported-traced-adjacencies-v1.1.tar.gz).
# - Have the [token](https://connectome-neuprint.github.io/neuprint-python/docs/client.html#neuprint.client.Client) for Neuprint HTTP access ready.
# - Have more than 60 GB free disk space (for Neuprint dump and NeuroArch database).
#
# A backup of the database created by this notebook can be downloaded [here](https://drive.google.com/file/d/1Y63UpypJ-eMgOdX3bcSRO4Ct3DqmH6-X/view?usp=sharing). To restore it in OrientDB, run
# ```
# /path/to/orientdb/bin/console.sh "create database plocal:../databases/hemibrain admin admin; restore database /path/to/hemibrain1.1_na_v1.0_backup.zip"
# ```
# +
import glob
import os
import subprocess
import csv
import json
import warnings
from requests import HTTPError
import numpy as np
import pandas as pd
from neuprint import Client
from tqdm import tqdm
import h5py
import pymeshlab as ml
import neuroarch.na as na
# -
# ## Define Brain Region
# First define all brain regions in the hemibrain data, and assign them as subsystem, neuropil or subregions.
all_brain_regions = \
{'OL(R)': {'System': 'OL(R)', 'Neuropil': None, 'Subregions': None},
'MB(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': None},
'MB(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': None},
'CX': {'System': 'CX', 'Neuropil': None, 'Subregions': None},
'LX(R)': {'System': 'LX(R)', 'Neuropil': None, 'Subregions': None},
'LX(L)': {'System': 'LX(L)', 'Neuropil': None, 'Subregions': None},
'VLNP(R)': {'System': 'VLNP(R)', 'Neuropil': None, 'Subregions': None},
'LH(R)': {'System': 'LH(R)', 'Neuropil': 'LH(R)', 'Subregions': None},
'SNP(R)': {'System': 'SNP(R)', 'Neuropil': None, 'Subregions': None},
'SNP(L)': {'System': 'SNP(L)', 'Neuropil': None, 'Subregions': None},
'INP': {'System': 'INP', 'Neuropil': None, 'Subregions': None},
'AL(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': None},
'AL(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': None},
'VMNP': {'System': 'VMNP', 'Neuropil': None, 'Subregions': None},
'PENP': {'System': 'PENP', 'Neuropil': None, 'Subregions': None},
'GNG': {'System': 'GNG', 'Neuropil': 'GNG', 'Subregions': None},
'AOT(R)': {'Tract': 'AOT(R)', 'Neuropil': None, 'Subregions': None},
'GC': {'Tract': 'GC', 'Neuropil': None, 'Subregions': None},
'GF(R)': {'Tract': 'GF(R)', 'Neuropil': None, 'Subregions': None},
'mALT(R)': {'Tract': 'mALT(R)', 'Neuropil': None, 'Subregions': None},
'mALT(L)': {'Tract': 'mALT(L)', 'Neuropil': None, 'Subregions': None},
'POC': {'Tract': 'POC', 'Neuropil': None, 'Subregions': None},
'ME(R)': {'System': 'OL(R)', 'Neuropil': 'ME(R)', 'Subregions': None},
'AME(R)': {'System': 'OL(R)', 'Neuropil': 'AME(R)', 'Subregions': None},
'CA(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'CA(R)'},
'CA(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': 'CA(L)'},
'dACA(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'dACA(R)'},
'lACA(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'lACA(R)'},
'vACA(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'vACA(R)'},
'PED(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'PED(R)'},
'CB': {'System': 'CX', 'Neuropil': ['FB', 'EB'], 'Subregions': None},
'PB': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': None},
'NO': {'System': 'CX', 'Neuropil': ['NO(R)', 'NO(L)'], 'Subregions': None},
'BU(R)': {'System': 'LX(R)', 'Neuropil': 'BU(R)', 'Subregions': None},
'BU(L)': {'System': 'LX(L)', 'Neuropil': 'BU(L)', 'Subregions': None},
'LAL(R)': {'System': 'LX(R)', 'Neuropil': 'LAL(R)', 'Subregions': None},
'LAL(L)': {'System': 'LX(L)', 'Neuropil': 'LAL(L)', 'Subregions': None},
'AOTU(R)': {'System': 'VLNP(R)', 'Neuropil': 'AOTU(R)', 'Subregions': None},
'PLP(R)': {'System': 'VLNP(R)', 'Neuropil': 'PLP(R)', 'Subregions': None},
'WED(R)': {'System': 'VLNP(R)', 'Neuropil': 'WED(R)', 'Subregions': None},
'SLP(R)': {'System': 'SNP(R)', 'Neuropil': 'SLP(R)', 'Subregions': None},
'SIP(R)': {'System': 'SNP(R)', 'Neuropil': 'SIP(R)', 'Subregions': None},
'SIP(L)': {'System': 'SNP(L)', 'Neuropil': 'SIP(L)', 'Subregions': None},
'SMP(R)': {'System': 'SNP(R)', 'Neuropil': 'SMP(R)', 'Subregions': None},
'SMP(L)': {'System': 'SNP(L)', 'Neuropil': 'SMP(L)', 'Subregions': None},
'CRE(R)': {'System': 'INP', 'Neuropil': 'CRE(R)', 'Subregions': None},
'CRE(L)': {'System': 'INP', 'Neuropil': 'CRE(L)', 'Subregions': None},
'IB': {'System': 'INP', 'Neuropil': 'IB', 'Subregions': None},
'ATL(R)': {'System': 'INP', 'Neuropil': 'ATL(R)', 'Subregions': None},
'ATL(L)': {'System': 'INP', 'Neuropil': 'ATL(L)', 'Subregions': None},
'AL-DC3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DC3(R)'},
'SAD': {'System': 'PENP', 'Neuropil': 'SAD', 'Subregions': None},
'FLA(R)': {'System': 'PENP', 'Neuropil': 'FLA(R)', 'Subregions': None},
'CAN(R)': {'System': 'PENP', 'Neuropil': 'CAN(R)', 'Subregions': None},
'PRW': {'System': 'PENP', 'Neuropil': 'PRW', 'Subregions': None},
'LO(R)': {'System': 'OL(R)', 'Neuropil': 'LO(R)', 'Subregions': None},
'LOP(R)': {'System': 'OL(R)', 'Neuropil': 'LOP(R)', 'Subregions': None},
"a'L(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "a'L(R)"},
"a'1(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "a'1(R)"},
"a'2(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "a'2(R)"},
"a'3(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "a'3(R)"},
"a'L(L)": {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': "a'L(L)"},
'aL(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'aL(R)'},
'a1(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'a1(R)'},
'a2(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'a2(R)'},
'a3(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'a3(R)'},
'aL(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': 'aL(L)'},
'gL(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'gL(R)'},
'g1(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'g1(R)'},
'g2(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'g2(R)'},
'g3(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'g3(R)'},
'g4(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'g4(R)'},
'g5(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'g5(R)'},
'gL(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': 'gL(L)'},
"b'L(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "b'L(R)"},
"b'1(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "b'1(R)"},
"b'2(R)": {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': "b'2(R)"},
"b'L(L)": {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': "b'L(L)"},
'bL(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'bL(R)'},
'b1(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'b1(R)'},
'b2(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': 'MB(R)', 'Subregions': 'b2(R)'},
'bL(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': 'MB(L)', 'Subregions': 'bL(L)'},
'FB': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': None},
'FBl1': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl1'},
'FBl2': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl2'},
'FBl3': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl3'},
'FBl4': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl4'},
'FBl5': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl5'},
'FBl6': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl6'},
'FBl7': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl7'},
'FBl8': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl8'},
'FBl9': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FBl9'},
'EB': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': None},
'EBr1': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr1'},
'EBr2r4': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr2r4'},
'EBr3am': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr3am'},
'EBr3d': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr3d'},
'EBr3pw': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr3pw'},
'EBr5': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr5'},
'EBr6': {'System': 'CX', 'Neuropil': 'EB', 'Subregions': 'EBr6'},
'PB(R1)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R1)'},
'PB(R2)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R2)'},
'PB(R3)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R3)'},
'PB(R4)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R4)'},
'PB(R5)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R5)'},
'PB(R6)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R6)'},
'PB(R7)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R7)'},
'PB(R8)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R8)'},
'PB(R9)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(R9)'},
'PB(L1)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L1)'},
'PB(L2)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L2)'},
'PB(L3)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L3)'},
'PB(L4)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L4)'},
'PB(L5)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L5)'},
'PB(L6)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L6)'},
'PB(L7)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L7)'},
'PB(L8)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L8)'},
'PB(L9)': {'System': 'CX', 'Neuropil': 'PB', 'Subregions': 'PB(L9)'},
'NO(R)': {'System': 'CX', 'Neuropil': 'NO(R)', 'Subregions': None},
'NO(L)': {'System': 'CX', 'Neuropil': 'NO(L)', 'Subregions': None},
'GA(R)': {'System': 'LX(R)', 'Neuropil': 'LAL(R)', 'Subregions': 'GA(R)'},
'AVLP(R)': {'System': 'VLNP(R)', 'Neuropil': 'AVLP(R)', 'Subregions': None},
'PVLP(R)': {'System': 'VLNP(R)', 'Neuropil': 'PVLP(R)', 'Subregions': None},
'RUB(R)': {'System': 'INP', 'Neuropil': 'CRE(R)', 'Subregions': 'RUB(R)'},
'RUB(L)': {'System': 'INP', 'Neuropil': 'CRE(L)', 'Subregions': 'RUB(L)'},
'ROB(R)': {'System': 'INP', 'Neuropil': 'CRE(R)', 'Subregions': 'ROB(R)'},
'SCL(R)': {'System': 'INP', 'Neuropil': 'SCL(R)', 'Subregions': None},
'SCL(L)': {'System': 'INP', 'Neuropil': 'SCL(L)', 'Subregions': None},
'ICL(R)': {'System': 'INP', 'Neuropil': 'ICL(R)', 'Subregions': None},
'ICL(L)': {'System': 'INP', 'Neuropil': 'ICL(L)', 'Subregions': None},
'VES(R)': {'System': 'VMNP', 'Neuropil': 'VES(R)', 'Subregions': None},
'VES(L)': {'System': 'VMNP', 'Neuropil': 'VES(L)', 'Subregions': None},
'EPA(R)': {'System': 'VMNP', 'Neuropil': 'EPA(R)', 'Subregions': None},
'EPA(L)': {'System': 'VMNP', 'Neuropil': 'EPA(L)', 'Subregions': None},
'GOR(R)': {'System': 'VMNP', 'Neuropil': 'GOR(R)', 'Subregions': None},
'GOR(L)': {'System': 'VMNP', 'Neuropil': 'GOR(L)', 'Subregions': None},
'SPS(R)': {'System': 'VMNP', 'Neuropil': 'SPS(R)', 'Subregions': None},
'SPS(L)': {'System': 'VMNP', 'Neuropil': 'SPS(L)', 'Subregions': None},
'IPS(R)': {'System': 'VMNP', 'Neuropil': 'IPS(R)', 'Subregions': None},
'AMMC': {'System': 'PENP', 'Neuropil': 'SAD', 'Subregions': 'AMMC'},
'AB(R)': {'System': 'CX', 'Neuropil': 'AB(R)', 'Subregions': None},
'AB(L)': {'System': 'CX', 'Neuropil': 'AB(L)', 'Subregions': None},
'FB-column3': {'System': 'CX', 'Neuropil': 'FB', 'Subregions': 'FB-column3'},
'NO1(R)': {'System': 'CX', 'Neuropil': 'NO(R)', 'Subregions': 'NO1(R)'},
'NO1(L)': {'System': 'CX', 'Neuropil': 'NO(L)', 'Subregions': 'NO1(L)'},
'NO2(R)': {'System': 'CX', 'Neuropil': 'NO(R)', 'Subregions': 'NO2(R)'},
'NO2(L)': {'System': 'CX', 'Neuropil': 'NO(L)', 'Subregions': 'NO2(L)'},
'NO3(R)': {'System': 'CX', 'Neuropil': 'NO(R)', 'Subregions': 'NO3(R)'},
'NO3(L)': {'System': 'CX', 'Neuropil': 'NO(L)', 'Subregions': 'NO3(L)'},
'MB(+ACA)(R)': {'System': 'MB(+ACA)(R)', 'Neuropil': None, 'Subregions': None},
'MB(+ACA)(L)': {'System': 'MB(+ACA)(L)', 'Neuropil': None, 'Subregions': None},
'LAL(-GA)(R)': {'System': 'LX(R)', 'Neuropil': 'LAL(R)', 'Subregions': 'LAL(-GA)(R)'},
'SAD(-AMMC)': {'System': 'PENP', 'Neuropil': 'SAD', 'Subregions': 'SAD(-AMMC)'},
'CRE(-RUB)(L)': {'System': 'INP', 'Neuropil': 'CRE(L)', 'Subregions': 'CRE(-RUB)(L)'},
'CRE(-ROB,-RUB)(R)': {'System': 'INP', 'Neuropil': 'CRE(R)', 'Subregions': 'CRE(-ROB,-RUB)(R)'},
'AL-DA1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DA1(R)'},
'AL-DA2(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DA2(L)'},
'AL-DA2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DA2(R)'},
'AL-DA3(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DA3(L)'},
'AL-DA3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DA3(R)'},
'AL-DA4l(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DA4l(R)'},
'AL-DA4m(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DA4m(L)'},
'AL-DA4m(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DA4m(R)'},
'AL-DC1(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DC1(L)'},
'AL-DC1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DC1(R)'},
'AL-DC2(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DC2(L)'},
'AL-DC2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DC2(R)'},
'AL-DC3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DC3(R)'},
'AL-DC4(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DC4(L)'},
'AL-DC4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DC4(R)'},
'AL-DL1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL1(R)'},
'AL-DL2d(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL2d(R)'},
'AL-DL2v(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL2v(R)'},
'AL-DL3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL3(R)'},
'AL-DL4(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DL4(L)'},
'AL-DL4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL4(R)'},
'AL-DL5(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DL5(L)'},
'AL-DL5(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DL5(R)'},
'AL-D(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-D(L)'},
'AL-DM1(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM1(L)'},
'AL-DM1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM1(R)'},
'AL-DM2(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM2(L)'},
'AL-DM2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM2(R)'},
'AL-DM3(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM3(L)'},
'AL-DM3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM3(R)'},
'AL-DM4(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM4(L)'},
'AL-DM4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM4(R)'},
'AL-DM5(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM5(L)'},
'AL-DM5(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM5(R)'},
'AL-DM6(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DM6(L)'},
'AL-DM6(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DM6(R)'},
'AL-DP1l(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DP1l(R)'},
'AL-DP1m(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-DP1m(L)'},
'AL-DP1m(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-DP1m(R)'},
'AL-D(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-D(R)'},
'AL-VA1d(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA1d(R)'},
'AL-VA1v(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA1v(R)'},
'AL-VA2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA2(R)'},
'AL-VA3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA3(R)'},
'AL-VA4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA4(R)'},
'AL-VA5(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA5(R)'},
'AL-VA6(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-VA6(L)'},
'AL-VA6(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA6(R)'},
'AL-VA7l(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA7l(R)'},
'AL-VA7m(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VA7m(R)'},
'AL-VC1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC1(R)'},
'AL-VC2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC2(R)'},
'AL-VC3l(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC3l(R)'},
'AL-VC3m(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC3m(R)'},
'AL-VC4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC4(R)'},
'AL-VC5(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VC5(R)'},
'AL-VL1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VL1(R)'},
'AL-VL2a(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VL2a(R)'},
'AL-VL2p(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VL2p(R)'},
'AL-VM1(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM1(R)'},
'AL-VM2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM2(R)'},
'AL-VM3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM3(R)'},
'AL-VM4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM4(R)'},
'AL-VM5d(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM5d(R)'},
'AL-VM5v(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM5v(R)'},
'AL-VM7d(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-VM7d(L)'},
'AL-VM7d(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM7d(R)'},
'AL-VM7v(L)': {'System': 'AL(L)', 'Neuropil': 'AL(L)', 'Subregions': 'AL-VM7v(L)'},
'AL-VM7v(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VM7v(R)'},
'AL-V(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-V(R)'},
'AL-VP5(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP5(R)'},
'AL-VP4(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP4(R)'},
'AL-VP3(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP3(R)'},
'AL-VP2(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP2(R)'},
'AL-VP1m(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP1m(R)'},
'AL-VP1l(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP1l(R)'},
'AL-VP1d(R)': {'System': 'AL(R)', 'Neuropil': 'AL(R)', 'Subregions': 'AL-VP1d(R)'},
}
# Extract from neuprint server the mesh files defining the boundary of these regions. The subsystems do not have mesh. You will need to put your token here.
# +
try:
os.mkdir('roi')
except FileExistsError:
warnings.warn('folder roi already exists.')
pass
token = ''
c = Client('neuprint.janelia.org', dataset='hemibrain:v1.1', token=token)
for region in all_brain_regions:
try:
c.fetch_roi_mesh(region, 'roi/{}.obj'.format(region))
except:
print(region)
# -
# ## Extract Neuron Attributes
# In the next two cells, we extract from the database dump all the *Traced* neurons and write them into 'neurons.csv' file.
def process(chunk):
status = np.nonzero(np.array([i == 'Traced' for i in chunk['status:string'].values]))[0]
used = chunk.iloc[status]
neurons = []
for i, row in used.iterrows():
neuropil_list = []
subregion_list = []
tract_list = []
kk = json.loads(row['roiInfo:string'])
for k, v in kk.items():
if k == "None": continue
region = all_brain_regions[k]
if region['Subregions'] is None:
if region['Neuropil'] is None:
if 'Tract' in region:
tract_list.append('{}:{}:{}'.format(
region['Tract'], v.get('pre',0), v.get('post',0)))
else:
continue
elif isinstance(region['Neuropil'], list):
continue
else:
neuropil_list.append('{}:{}:{}'.format(
region['Neuropil'], v.get('pre', 0), v.get('post', 0)))
else:
subregion_list.append('{}:{}:{}'.format(
region['Subregions'], v.get('pre', 0), v.get('post', 0)))
neuropil_list = ';'.join(neuropil_list)
subregion_list = ';'.join(subregion_list)
tract_list = ';'.join(tract_list)
li = [row['bodyId:long'], row['pre:int'], row['post:int'], row['status:string'],\
row['statusLabel:string'], int(row['cropped:boolean']) if not np.isnan(row['cropped:boolean']) else row['cropped:boolean'], row['instance:string'], \
row['type:string'], row['notes:string'], row['cellBodyFiber:string'], row['somaLocation:point{srid:9157}'], \
row['somaRadius:float'], row['size:long'], neuropil_list, subregion_list,tract_list]
neurons.append(li)
return neurons
# The mesh are then downsampled using [MeshLab](https://www.meshlab.net/):
# +
chunksize = 100000
with open('neurons.csv', 'w') as f:
writer = csv.writer(f)
writer.writerow(['bodyID','pre','post','status','statusLabel','cropped','instance','type','notes','cellBodyFiber','somaLocation','somaRadius','size','neuropils','subregions','tracts'])
for chunk in tqdm(pd.read_csv('hemibrain_v1.1_neo4j_inputs/Neuprint_Neurons_20631.csv', chunksize=chunksize)):
neurons = process(chunk)
writer.writerows(neurons)
# -
# To download the SWC files for all the traced neurons into folder 'swc', we can run the code below uncommented, or download it from [here](https://drive.google.com/file/d/12hH2n9Au2UUcStgU52WZg4XX8y7phJ0X).
# +
# try:
# os.mkdir('swc')
# except FileExistsError:
# warnings.warn('folder roi already exists.')
# pass
# neurons = pd.read_csv('neurons.csv')
# for i, row in tqdm.tqdm(neurons.iterrows()):
# bodyID = int(row['bodyID'])
# try:
# s = c.fetch_skeleton(bodyID, format='pandas')
# s.to_csv('swc/{}.swc'.format(bodyID), header=False, index=False, sep=' ')
# except HTTPError:
# print(bodyID)
# -
# ## Extract Synapses
# We only use the neurons that are traced, or roughly traced, or has a name/instance assigned to it. Only synapses between these neurons are extracted below.
# +
neurons = pd.read_csv('neurons.csv')
used = []
for i, row in neurons.iterrows():
if row['statusLabel'] in ['Traced', 'Roughly traced'] or isinstance(row['instance'], str) or isinstance(row['type'], str):
used.append(i)
traced_neuron_id = neurons.iloc[used]['bodyID'].to_numpy()
chunksize = 1000000
pre_syn = np.empty((int(1e8),3), np.int64)
post_syn = np.empty((int(1e8),3), np.int64)
pre_count = 0
post_count = 0
count = 0
for chunk in pd.read_csv('hemibrain_v1.1_neo4j_inputs/Neuprint_SynapseSet_to_Synapses_20631.csv', chunksize=chunksize):
ids = chunk[':START_ID']
pre_site = np.array([[n, int(i.split('_')[0]), int(i.split('_')[1])] \
for n,i in enumerate(ids) if i.split('_')[2] == 'pre'])
post_site = np.array([[n, int(i.split('_')[0]), int(i.split('_')[1])] \
for n,i in enumerate(ids) if i.split('_')[2] == 'post'])
pre_site_known = pre_site[np.logical_and(
np.isin(pre_site[:,1], traced_neuron_id),
np.isin(pre_site[:,2], traced_neuron_id)),0]
post_site_known = post_site[np.logical_and(
np.isin(post_site[:,1], traced_neuron_id),
np.isin(post_site[:,2], traced_neuron_id)),0]
retrieved_pre_site = chunk.iloc[pre_site_known]
pre_site = np.array([[row[':END_ID(Syn-ID)'], int(row[':START_ID'].split('_')[0]), int(row[':START_ID'].split('_')[1])] \
for i, row in retrieved_pre_site.iterrows()])
retrieved_post_site = chunk.iloc[post_site_known]
post_site = np.array([[row[':END_ID(Syn-ID)'], int(row[':START_ID'].split('_')[0]), int(row[':START_ID'].split('_')[1])] \
for i, row in retrieved_post_site.iterrows()])
if pre_site.size:
pre_syn[pre_count:pre_count+pre_site.shape[0], :] = pre_site
pre_count += pre_site.shape[0]
if post_site.size:
post_syn[post_count:post_count+post_site.shape[0], :] = post_site
post_count += post_site.shape[0]
count += chunksize
print(count, pre_count, post_count)
pre_syn = pre_syn[:pre_count,:]
post_syn = post_syn[:post_count,:]
ind = np.argsort(pre_syn[:,0])
pre_syn_sorted = pre_syn[ind, :]
ind = np.argsort(post_syn[:,0])
post_syn_sorted = post_syn[ind, :]
# +
# extract synapse (pre-site) to synapse (post-site) connection
# use only the post synaptic site to get all the synapses because one presynaptic site can have multiple postsynaptic sites
post_syn_index = post_syn_sorted[:,0].copy()
df = pd.read_csv('hemibrain_v1.1_neo4j_inputs/Neuprint_Synapse_Connections_20631.csv')
post_ids = df[':END_ID(Syn-ID)']
used = np.where(post_ids.isin(post_syn_index).to_numpy())[0]
connections = df.iloc[used].to_numpy()
ind = np.argsort(connections[:,1])
connections = connections[ind, :]
# +
# extract synapse details
chunksize = 100000
pre_syn_index = list(set(pre_syn_sorted[:,0].copy()))
pre_syn_index.extend(list(post_syn_sorted[:,0].copy()))
syn_index = np.array(sorted(pre_syn_index))
del pre_syn_index#, pre_syn_sorted, post_syn_sorted
synapse_array = np.empty((len(syn_index), 230+6), np.int64)
synapse_count = 0
count = 0
for chunk in pd.read_csv('hemibrain_v1.1_neo4j_inputs/Neuprint_Synapses_20631.csv', chunksize=chunksize):
ids = chunk[':ID(Syn-ID)']
start_id = ids.iloc[0]
stop_id = ids.iloc[-1]
pre_start = np.searchsorted(syn_index, start_id, side='left')
pre_end = np.searchsorted(syn_index, stop_id, side='right')
if pre_start >= len(syn_index):
pre_index = []
else:
if pre_end >= len(syn_index):
pre_index = syn_index[pre_start:pre_end] #same as syn_index[pre_start:]
else:
pre_index = syn_index[pre_start:pre_end]
pre_used_synapse = chunk.loc[ids.isin(pre_index)]
li = np.empty((pre_index.size, 230+6), np.int64)
i = 0
for _, row in pre_used_synapse.iterrows():
location = eval(row['location:point{srid:9157}'].replace('x', "'x'").replace('y', "'y'").replace('z', "'z'"))
li[i,:6] = [row[':ID(Syn-ID)'], # synpase id
0 if row['type:string'] == 'pre' else 1, #synapse type
int(row['confidence:float']*1000000), #confidence
location['x'], location['y'], location['z']]
li[i,6:] = ~np.isnan(np.asarray(row.values[5:], np.double))
i += 1
synapse_array[synapse_count:synapse_count+pre_index.shape[0],:] = li
synapse_count += pre_index.shape[0]
count += chunksize
print(count, len(pre_used_synapse))
synapse_array = synapse_array[:synapse_count,:]
# +
# reorder synapses
synapse_connections = connections
ids = synapse_array[:,0]
syn_id_dict = {j: i for i, j in enumerate(ids)}
ids = post_syn_sorted[:,0]
post_syn_id_dict = {j: i for i, j in enumerate(ids)} # map syn id to post_syn_sorted
synapse_dict = {}
wrong_synapse = 0
for i, pair in tqdm(enumerate(synapse_connections)):
pre_syn_id = pair[0]
post_syn_id = pair[1]
post_id = post_syn_id_dict[post_syn_id]
post_info = synapse_array[syn_id_dict[post_syn_id]]
post_neuron_id, pre_neuron_id = post_syn_sorted[post_id, 1:]
pre_info = synapse_array[syn_id_dict[pre_syn_id]]
if pre_neuron_id not in synapse_dict:
synapse_dict[pre_neuron_id] = {}
pre_dict = synapse_dict[pre_neuron_id]
if post_neuron_id not in synapse_dict[pre_neuron_id]:
pre_dict[post_neuron_id] = {'pre_synapse_ids': [],
'post_synapse_ids': [],
'pre_confidence': [],
'post_confidence': [],
'pre_x': [],
'pre_y': [],
'pre_z': [],
'post_x': [],
'post_y': [],
'post_z': [],
'regions': np.zeros(230, np.int32)}
info_dict = pre_dict[post_neuron_id]
info_dict['pre_synapse_ids'].append(pre_syn_id)
info_dict['post_synapse_ids'].append(post_syn_id)
info_dict['pre_confidence'].append(pre_info[2])
info_dict['post_confidence'].append(post_info[2])
info_dict['pre_x'].append(pre_info[3])
info_dict['pre_y'].append(pre_info[4])
info_dict['pre_z'].append(pre_info[5])
info_dict['post_x'].append(post_info[3])
info_dict['post_y'].append(post_info[4])
info_dict['post_z'].append(post_info[5])
info_dict['regions'] += post_info[6:]
chunk = pd.read_csv('hemibrain_v1.1_neo4j_inputs/Neuprint_Synapses_20631.csv', chunksize=1).get_chunk()
labels = [i.split(':')[0] for i in chunk.columns.to_list()]
regions = labels[5:]
with open('synapses.csv', 'w') as f:
writer = csv.writer(f)
writer.writerow(['pre_id','post_id','N','pre_confidence','post_confidence',\
'pre_x','pre_y','pre_z','post_x','post_y','post_z',\
'neuropils','subregions','tracts'])
for pre, k in tqdm(synapse_dict.items()):
for post, v in k.items():
reg = {regions[i]: v['regions'][i] for i in np.nonzero(v['regions'])[0]}
neuropil_list = []
subregion_list = []
tract_list = []
for k, n in reg.items():
region = all_brain_regions[k]
if region['Subregions'] is None:
if region['Neuropil'] is None:
if 'Tract' in region:
tract_list.append('{}:{}'.format(
region['Tract'], n))
else:
continue
elif isinstance(region['Neuropil'], list):
continue
else:
neuropil_list.append('{}:{}'.format(
region['Neuropil'], n))
else:
subregion_list.append('{}:{}'.format(
region['Subregions'], n))
neuropil_list = ';'.join(neuropil_list)
subregion_list = ';'.join(subregion_list)
tract_list = ';'.join(tract_list)
writer.writerow([pre, post, len(v['pre_x']), str(v['pre_confidence']), \
str(v['post_confidence']), str(v['pre_x']), str(v['pre_y']), str(v['pre_z']), \
str(v['post_x']), str(v['post_y']), str(v['post_z']), \
neuropil_list, subregion_list, tract_list])
# -
# ## Loading NeuroArch Database
# Create and connect to database. mode 'o' overwrites the entire database.
hemibrain = na.NeuroArch('hemibrain', mode = 'o')
# Create a species
species = hemibrain.add_Species('Drosophila melanogaster', stage = 'adult',
sex = 'female',
synonyms = ['fruit fly', 'common fruit fly', 'vinegar fly'])
# Create a datasource under the species
version = '1.1'
datasource = hemibrain.add_DataSource('Hemibrain', version = version,
url = 'https://www.janelia.org/project-team/flyem/hemibrain',
species = species)
hemibrain.default_DataSource = datasource
# Create subsystems, tracts, neuropils and subregions under the datasource
for k, v in all_brain_regions.items():
if v['Neuropil'] is None and v['Subregions'] is None:
if 'System' in v:
hemibrain.add_Subsystem(k)
elif 'System' in v:
if v['Neuropil'] == v['System'] and v['Subregions'] is None:
hemibrain.add_Subsystem(k)
with open('filter_file_tmp.mlx', 'w') as f:
f.write("""<!DOCTYPE FilterScript>
<FilterScript>
<filter name="Simplification: Quadric Edge Collapse Decimation">
<Param type="RichInt" value="60000" name="TargetFaceNum"/>
<Param type="RichFloat" value="0.05" name="TargetPerc"/>
<Param type="RichFloat" value="1" name="QualityThr"/>
<Param type="RichBool" value="true" name="PreserveBoundary"/>
<Param type="RichFloat" value="1" name="BoundaryWeight"/>
<Param type="RichBool" value="true" name="OptimalPlacement"/>
<Param type="RichBool" value="true" name="PreserveNormal"/>
<Param type="RichBool" value="true" name="PlanarSimplification"/>
</filter>
</FilterScript>""")
for k, v in all_brain_regions.items():
if v['Neuropil'] is None and v['Subregions'] is None:
if 'Tract' in v:
ms = ml.MeshSet()
ms.load_new_mesh("roi/{}.obj".format(k))
ms.load_filter_script('filter_file_tmp.mlx')
ms.apply_filter_script()
current_mesh = ms.current_mesh()
hemibrain.add_Tract(k, morphology = {'type': 'mesh',
"vertices": (current_mesh.vertex_matrix()*0.008).flatten().tolist(),
"faces": current_mesh.face_matrix().flatten().tolist()})
for k, v in all_brain_regions.items():
if v['Neuropil'] is not None and v['Subregions'] is None:
if isinstance(v['Neuropil'], list):
continue
ms = ml.MeshSet()
ms.load_new_mesh("roi/{}.obj".format(k))
ms.load_filter_script('filter_file_tmp.mlx')
ms.apply_filter_script()
current_mesh = ms.current_mesh()
hemibrain.add_Neuropil(k,
morphology = {'type': 'mesh',
"vertices": (current_mesh.vertex_matrix()*0.008).flatten().tolist(),
"faces": current_mesh.face_matrix().flatten().tolist()},
subsystem = v['System'])
for k, v in all_brain_regions.items():
if v['Subregions'] is not None:
if isinstance(v['Neuropil'], list):
continue
if os.path.exists("roi/{}.obj".format(k)):
ms = ml.MeshSet()
ms.load_new_mesh("roi/{}.obj".format(k))
ms.load_filter_script('filter_file_tmp.mlx')
ms.apply_filter_script()
current_mesh = ms.current_mesh()
hemibrain.add_Subregion(k,
morphology = {'type': 'mesh',
"vertices": (current_mesh.vertex_matrix()*0.008).flatten().tolist(),
"faces": current_mesh.face_matrix().flatten().tolist()},
neuropil = v['Neuropil'])
else:
hemibrain.add_Subregion(k,
neuropil = v['Neuropil'])
# Load Neurons
# If you downloaded the swc using neuprint API, use this cell
def load_swc(file_name):
df = pd.read_csv(file_name, sep = ' ', header = None, comment = '#', index_col = False,
names = ['sample', 'x', 'y', 'z', 'r', 'parent'],
skipinitialspace = True)
return df
# +
# If you downloaded the swc from the zip file, use this cell
# def load_swc(file_name):
# df = pd.read_csv(file_name, sep = ' ', header = None, comment = '#', index_col = False,
# names = ['sample', 'identifier', 'x', 'y', 'z', 'r', 'parent'],
# skipinitialspace = True)
# return df
# +
neuron_list = pd.read_csv('neurons.csv')
swc_dir = 'swc'
uname_dict = {}
for i, row in tqdm(neuron_list.iterrows()):
if row['statusLabel'] in ['Traced', 'Roughly traced']:
pass
elif isinstance(row['instance'], str) or isinstance(row['type'], str):
pass
else:
continue
bodyID = row['bodyID']
cell_type = row['type']
name = row['instance']
if not isinstance(name, str):
if isinstance(cell_type, str):
name = '{}_{}'.format(cell_type, bodyID)
else:
cell_type = 'unknown'
name = 'unknown_{}'.format(bodyID)
else:
if not isinstance(cell_type, str):
cell_type = 'unknown'
if not isinstance(name, str):
name = 'unknown_{}'.format(bodyID)
else:
name = '{}_{}'.format(name, bodyID)
else:
if name not in uname_dict:
uname_dict[name] = 0
uname_dict[name] += 1
name = '{}_{}'.format(name, uname_dict[name])
info = {}
if isinstance(row['notes'], str):
info['notes'] = row['notes']
c_neuropils = row['neuropils']
c_subregions = row['subregions']
c_tracts = row['tracts']
arborization = []
if isinstance(c_neuropils, str):
dendrites = {j.split(':')[0]: int(j.split(':')[2]) for j in c_neuropils.split(';') if int(j.split(':')[2]) > 0}
axons = {j.split(':')[0]: int(j.split(':')[1]) for j in c_neuropils.split(';') if int(j.split(':')[1]) > 0}
arborization.append({'dendrites': dendrites, 'axons': axons, 'type': 'neuropil'})
if isinstance(c_subregions, str):
dendrites = {j.split(':')[0]: int(j.split(':')[2]) for j in c_subregions.split(';') if int(j.split(':')[2]) > 0}
axons = {j.split(':')[0]: int(j.split(':')[1]) for j in c_subregions.split(';') if int(j.split(':')[1]) > 0}
arborization.append({'dendrites': dendrites, 'axons': axons, 'type': 'subregion'})
if isinstance(c_tracts, str):
dendrites = {j.split(':')[0]: int(j.split(':')[2]) for j in c_tracts.split(';') if int(j.split(':')[2]) > 0}
axons = {j.split(':')[0]: int(j.split(':')[1]) for j in c_tracts.split(';') if int(j.split(':')[1]) > 0}
arborization.append({'dendrites': dendrites, 'axons': axons, 'type': 'tract'})
df = load_swc('{}/{}.swc'.format(swc_dir, bodyID))
morphology = {'x': (df['x']*0.008).tolist(),
'y': (df['y']*0.008).tolist(),
'z': (df['z']*0.008).tolist(),
'r': (df['r']*0.008).tolist(),
'parent': df['parent'].tolist(),
'identifier': [0]*(len(df['x'])),
'sample': df['sample'].tolist(),
'type': 'swc'}
hemibrain.add_Neuron(name, # uname
cell_type, # name
referenceId = str(bodyID), #referenceId
info = info if len(info) else None,
morphology = morphology,
arborization = arborization)
# +
# If restarting the kernel after loading neurons, start with this
# hemibrain = na.NeuroArch('hemibrain', mode = 'w')
# hemibrain.default_DataSource = hemibrain.find_objs('DataSource', name = 'Hemibrain')[0]
# +
# find all the neurons so they can be keyed by their referenceId.
neurons = hemibrain.sql_query('select from Neuron').nodes_as_objs
# set the cache so there is no need for database access.
for neuron in neurons:
hemibrain.set('Neuron', neuron.uname, neuron, hemibrain.default_DataSource)
neuron_ref_to_obj = {int(neuron.referenceId): neuron for neuron in neurons}
# -
# Load synapses
# +
synapse_df = pd.read_csv('synapses.csv')
for i, row in tqdm(synapse_df.iterrows()):
pre_neuron = neuron_ref_to_obj[row['pre_id']]
post_neuron = neuron_ref_to_obj[row['post_id']]
pre_conf = np.array(eval(row['pre_confidence']))/1e6
post_conf = np.array(eval(row['post_confidence']))/1e6
NHP = np.sum(np.logical_and(post_conf>=0.7, pre_conf>=0.7))
c_neuropils = row['neuropils']
c_subregions = row['subregions']
c_tracts = row['tracts']
arborization = []
neuropils = {}
if isinstance(c_neuropils, str):
arborization.append({'type': 'neuropil',
'synapses': {j.split(':')[0]: int(j.split(':')[1]) \
for j in c_neuropils.split(';') \
if int(j.split(':')[1]) > 0}})
if isinstance(c_subregions, str):
arborization.append({'type': 'subregion',
'synapses': {j.split(':')[0]: int(j.split(':')[1]) \
for j in c_subregions.split(';') \
if int(j.split(':')[1]) > 0}})
if isinstance(c_tracts, str):
arborization.append({'type': 'tract',
'synapses': {j.split(':')[0]: int(j.split(':')[1]) \
for j in c_tracts.split(';') \
if int(j.split(':')[1]) > 0}})
content = {'type': 'swc'}
content['x'] = [round(i, 3) for i in (np.array(eval(row['pre_x'])+eval(row['post_x']))*0.008).tolist()]
content['y'] = [round(i, 3) for i in (np.array(eval(row['pre_y'])+eval(row['post_y']))*0.008).tolist()]
content['z'] = [round(i, 3) for i in (np.array(eval(row['pre_z'])+eval(row['post_z']))*0.008).tolist()]
content['r'] = [0]*len(content['x'])
content['parent'] = [-1]*(len(content['x'])//2) + [i+1 for i in range(len(content['x'])//2)]
content['identifier'] = [7]*(len(content['x'])//2) + [8]*(len(content['x'])//2)
content['sample'] = [i+1 for i in range(len(content['x']))]
content['confidence'] = [round(i, 3) for i in pre_conf.tolist()] + [round(i, 3) for i in post_conf.tolist()]
hemibrain.add_Synapse(pre_neuron, post_neuron, N = row['N'], NHP = NHP,
morphology = content,
arborization = arborization)
# -
| hemibrain/v1.1/Hemibrain_Neuprint_to_NeuroArch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tf2_pcse)
# language: python
# name: conda_tf2_pcse
# ---
import sys
sys.path.append('./')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from SIMPLE.crop import *
# ## Loading parameters
crop_params = pd.read_csv('./SIMPLE/params/crop_params.csv', index_col='Crop')
soil_params = pd.read_csv('./SIMPLE/params/soil_params.csv', index_col='Crop')
crop_params = crop_params.loc['Tomato']
soil_params = soil_params.loc['Tomato']
crop_params
soil_params
crop_params = crop_params.values[0, 1:]
soil_params = soil_params.values[0, 4:]
# ## Loading weather data
weather_df = pd.read_csv('./sample_input/UFGA9601.txt', index_col='DATE', delim_whitespace=True)
weather_df = weather_df.iloc[:-1, :3]
weather_df['CO2'] = 400
weather_df = weather_df[['TMAX', 'TMIN', 'SRAD', 'CO2']]
cultvation_period = int(weather_df.index[-1]) - int(weather_df.index[0])
weather_df.head()
# ## Simulation
test_tomato = Crop(crop_params, soil_params)
yields, TTs, biomass, days, solar = test_tomato.run(weather_df.values)
output_df = pd.DataFrame([days, TTs, biomass, solar]).T
output_df.columns = ['date', 'cum_temp', 'biomass', 'solar']
output_df['date'] = output_df['date'].astype(int)
output_df = output_df.set_index('date')
output_df['yield'] = np.nan
output_df.iloc[-1, -1] = yields
output_df.to_csv('./results/SIMPLE_tomato_out.csv')
# ## Plotting
# +
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(10,8))
for var, ax in zip(['cum_temp', 'biomass', 'solar', 'yield'], axes.flatten()):
if var == 'yield':
ax.plot(output_df.index, output_df[var], 'bo')
ax.set_title(var)
ax.set_xbound([output_df.index[0]-1, output_df.index[-1]+1])
break
ax.plot(output_df.index, output_df[var], 'b-')
ax.set_title(var)
ax.set_xbound([output_df.index[0]-1, output_df.index[-1]+1])
# fig.autofmt_xdate()
# fig.savefig('./results/sugarbeet.png')
| 1_SIMPLE_implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Youtube Comments and Captions
# %pip install youtube_comment_scraper
# %pip install youtube_dl
from youtube_comment_scraper import *
import pandas as pd
youtube.open("https://www.youtube.com/watch?v=cw7FzcqRXqE&ab")
response=youtube.video_comments()
data=response['body']
# #data=[{"Comment": "This planet belongs to all of us ", "UserLink": "https://www.youtube.com/channel/UCpH24tSZnFGAECI_ZCa7jJQ", "user": "
# <NAME>", "Time": "14 hours ago", "Likes": "23"}]
len(data)
df = pd.DataFrame(data)
df
youtube.close()
# + jupyter={"outputs_hidden": true}
data
# -
replies = youtube.get_comment_replies()
dfr = pd.DataFrame.from_dict(replies, orient='index')
dfr = dfr.transpose()
dfr['body'][3]
# # captions
https://www.youtube.com/watch?v=4vlM8vr5Aco&ab
import youtube_dl
# +
def download_subs(url, lang="en"):
opts = {
"skip_download": True,
"writesubtitles": "%(name)s.vtt",
"subtitlelangs": lang
}
with youtube_dl.YoutubeDL(opts) as yt:
yt.download([url])
url = "https://www.youtube.com/watch?v=rtzCPSM0720"
response = download_subs(url)
# -
df = pd.read_vtt
| ETL Youtube.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#---------Test Imports-------------
import numpy as np
import keras
import gensim
import nltk
keras.layers.Dense(20)
# +
#-------Download R packages---------
## these two lines for JupyterHub only
#import os
#os.environ['R_HOME'] = "/usr/share/anaconda3/lib/R"
import rpy2
from rpy2.robjects.packages import importr
r_utils = importr('utils')
package_list = ['aplpack',
'cluster',
'codetools',
'dbscan',
'factoextra',
'gam',
'ggplot2',
'splines',
'TeachingDemos'
]
for name in package_list:
r_utils.install_packages(name)
import rpy2
from rpy2.robjects.packages import importr
r_utils = importr('utils')
package_list = ['aplpack',
'cluster',
'codetools',
'dbscan',
'factoextra',
'gam',
'ggplot2',
'splines',
'TeachingDemos'
]
for name in package_list:
r_utils.install_packages(name)
# -
| content/labs/lab1/cs109b_lab1_Rsetup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
import subprocess
import os
import sys
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# +
d = '/home/jobasha/Documents/Dropbox/Repositories/planningfortransparency/problems/grid-obst/'
output_dir = '../results/'
output_file = 'results.txt'
# +
object_vector = []
fluent_vector = []
time_first_action_vector = []
time_average_vector = []
time_full_vector = []
average_ambiguity_vector = []
threshold_1_v = []
threshold_2_v = []
threshold_3_v = []
ns = []
hs = []
ps = []
ths = []
tis = []
tots = []
cs = []
obs = []
wcds = []
domain = d + "/domain.pddl"
count = 0
dirs = list(sorted(os.listdir(d)))
#dirs.reverse()
#dirs = os.listdir(d)
for problem_n in dirs:#os.listdir(d):
try:
if(not problem_n.startswith("problem")):
print("\t\tReject: " + problem_n)
continue
count = 1 + count
problem_file = d + problem_n
numbers = problem_n.split('_')
n = int(numbers[1])
m = int(numbers[2])
#prob_number = int(numbers[4])
hype_number = int(numbers[3])
prob_number = int(numbers[4].split(".")[0])
objects = n*m
print(str(count) + ". Problem - " + str(n) + "x" +str(m) + "-"+ str(prob_number))
sys.stdout.flush()
hype = d + "hyp_" + str(n) + "_" +str(m) + "_" + str(hype_number) + ".dat"
#print("\t" +problem_file)
problem = problem_file
goal_set = hype
#max_nodes = "10"
#df = "0.5"
#max_s_nodes = "10"
precision = "2"
theta = "0.1"
cmd = ["ulimit -Sv 10000000; /home/jobasha/Documents/Dropbox/Repositories/planningfortransparency/utilities/WCD_w_Belief_s/./WCD" + " " + domain + " " + problem + " " + goal_set] #+ " -mc"]
flags = [ domain, problem, goal_set,
# "-m", max_nodes,
# "-D", df,
# "-s", max_s_nodes,
# "-P", precision,
# "-t", theta,
"-mc"]
pipe = subprocess.Popen(
cmd,
stdin=subprocess.PIPE, stdout=subprocess.PIPE, stderr=subprocess.STDOUT, shell=True)
answer, stderr = pipe.communicate()
wcd = float(answer.decode().split("\n")[1])
#print(str(answer.decode()))
# resf = open(output_dir + problem_n + ".res", "w")
# resf.write(str(answer))
# resf.close()
# #print(domain,problem)
# #print(answer.decode())
# #print(stderr)
# lines = str(answer).split("\\n")[2:-1]
# #print(lines)
# actions = lines[0].split(",")[:-1]
# #print(actions)
# times = list(map(float,lines[1].split(",")[:-1]))
# #print(times)
# distances = list(map(float,lines[2].split(",")[:-1]))
# #print(distances)
# average_distance = sum(distances) / float(len(distances))
# prob_dists = []
# for line in lines[3:-6]:
# prob_dists += [list(map(float,line.split(",")[:-1]))]
# fluents = int(lines[-5])
# goal_index = int(lines[-6])
# threshold_3 = float(lines[-1])
# threshold_2 = float(lines[-2])
# threshold_1 = float(lines[-3])
# threshold_1_v.append(threshold_1)
# threshold_2_v.append(threshold_2)
# threshold_3_v.append(threshold_3)
# #print(threshold_1, threshold_2, threshold_3)
# #prob_dists = prob_dists[:-2]
# n_l = 1/len(prob_dists)
# prob_dists = [[n_l] + prob_dists[x] for x in range(len(prob_dists))]
# actions = ["I"] + actions
# total_time = sum(times)
# average_time_per_action = sum(times) / float(len(times))
# time_first_action = times[0]
print("\t" + str(count) + ". Problem - " + str(n) + "x" + str(prob_number) +
", WCD: " + str(wcd))
# if (threshold_1 > 0.5):
# print(actions)
# ns.append(n)
# hs.append(hype_number)
# obs.append(prob_number)
# ths.append(threshold_1)
# tis.append(time_first_action)
# tots.append(total_time)
wcds.append(wcd)
# object_vector.append(objects)
# time_first_action_vector.append(time_first_action)
# time_average_vector.append(average_time_per_action)
# time_full_vector.append(total_time)
# average_ambiguity_vector.append(average_distance)
# fluent_vector.append(fluents)
except BaseException as e:
print("\t\tERROR FILE: " + str(n) + "_" + str(prob_number))
print(e)
print(answer.decode())
print(stderr)
continue
# average_t_1 = sum(threshold_1_v) / len(threshold_1_v)
# average_t_2 = sum(threshold_2_v) / len(threshold_2_v)
# average_t_3 = sum(threshold_3_v) / len(threshold_3_v)
# print()
# for i in ns:
# print(i)
# print()
# for i in hs:
# print(i)
# print()
# for i in ths:
# print(i)
# print()
# for i in tis:
# print(i)
# print()
for i in wcds:
print(i)
print()
# output_dir = '../results/'
# output_file = 'online_n_blocks.txt'
# file_out = open(output_dir + output_file , "a+")
# file_out.write(problem)
# file_out.write("\t")
# file_out.write(str(total_time))
# file_out.write("\t")
# file_out.write(str(time_first_action))
# file_out.write("\t")
# file_out.write(str(average_time_per_action))
# file_out.write("\t")
# file_out.write(str(average_distance))
# file_out.write("\n")
# file_out.write(str(objects))
# file_out.write("\n")
# file_out.close()
# print(object_vector)
# print(time_first_action_vector)
# file_out = open(output_dir + output_file , "a+")
# file_out.write(problem)
# file_out.write("\t")
# file_out.write(str(total_time))
# file_out.write("\t")
# file_out.write(str(time_first_action))
# file_out.write("\t")
# file_out.write(str(average_time_per_action))
# file_out.write("\t")
# file_out.write(str(average_distance))
# file_out.write("\n")
# file_out.write(objects)
# file_out.write("\n")
# file_out.close()
# print("Actions: ")
# print(actions)
# print()
# print("Times: ")
# print(times)
# print()
# print("Distances: ")
# print(distances)
# print()
# print("Probabilities: ")
# print(prob_dists)
# print(prob_dists)
# plt.figure(2)
# axes = plt.gca()
# axes.set_ylim([0,1])
# axes.set_xlabel('actions')
# axes.set_ylabel('P(G | O)')
# counter = 0
# print(actions)
# print(prob_dists[0])
# for i in prob_dists:
# plt.plot(range(0, len(actions)), i, label=counter)
# counter += 1
# #plt.xticks(range(0, len(actions)-1), actions)
# plt.legend()
# plt.figure(1)
# ax = plt.axes(projection='3d')
# ax.set_ylim([0,1])
# ax.set_xlim([0,1])
# ax.set_zlim([0,1])
# ax.set_xlabel('P(G2 | O)')
# ax.set_ylabel('P(G1 | O)')
# ax.set_zlabel('P(G* | O)')
# G_star = 0
# G1 = 3
# G2 = 4
# r = range(len(prob_dists[0]))
# p0 = [-prob_dists[G_star][x0] + prob_dists[G_star][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# p1 = [-prob_dists[G1][x0] + prob_dists[G1][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# p2 = [-prob_dists[G2][x0] + prob_dists[G2][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# ax.plot(prob_dists[G1], prob_dists[G2], prob_dists[G_star], '-b')
# ax.quiver(prob_dists[G1][:-1], prob_dists[G2][:-1],prob_dists[G_star][:-1], p1, p2,p0, length=0.1, pivot='tail')
# plt.show()
# -
new_wcds = [x for x in wcds if x < 1]
print(str(sum(new_wcds)/len(new_wcds)))
# +
average_t_1 = sum(threshold_1_v) / len(threshold_1_v)
average_t_2 = sum(threshold_2_v) / len(threshold_2_v)
average_t_3 = sum(threshold_3_v) / len(threshold_3_v)
print(average_t_1)
print(average_t_2)
print(average_t_3)
# +
# plt.figure(5)
# plt.ylim([0,25])
# plt.xlabel('Fluents')
# plt.ylabel('Time for first action (s)')
# av = list(map(abs, average_ambiguity_vector))
# ob = object_vector
# tf = time_first_action_vector
# ta = time_average_vector
# tfu = time_full_vector
# print(fluent_vector[4])
# print()
# plt.scatter(fluent_vector, time_first_action_vector)
# +
# domain = d + "domain.pddl"
# count = 0
# object_vector_l = []
# time_first_action_vector_l = []
# for problem_n in os.listdir(d):
# # print(problem_n)
# try:
# if(not problem_n.startswith("problem")):
# continue
# problem_file = d + problem_n
# numbers = problem_n.split('_')
# n = int(numbers[1])
# m = int(numbers[2])
# objects = n
# hype = d + "hype_" + str(n) + "_" + str(m) + ".dat"
# # print("\t" +problem_file)
# problem = problem_file
# cmd = ["time", "python"]
# flags = [ "/home/jobasha/Documents/PhD/Planners/FastDownward/fast-downward.py","--alias", "seq-sat-lama-2011", domain, problem]
# pipe = subprocess.Popen(
# cmd + flags,
# stdin=subprocess.PIPE, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
# #print(''.join(pipe.stderr.readlines()))
# answer, stderr = pipe.communicate()
# answer1 = str(stderr).split("\'")
# answer2 = answer1[1].split("system")
# time_taken = sum(list(map(float, answer2[0].split("user"))))
# file = open("sas_plan.1","r")
# counter = 0
# for lines in file :
# counter = counter + 1
# print("Problem - " + str(n) + "x" + str(m) + ", Cost: " + str(counter) + ", " + str(time_taken) )
# os.remove("sas_plan.1")
# os.remove("output")
# os.remove("output.sas")
# if(counter != 1):
# time_first_action_vector_l.append(time_taken)
# object_vector_l.append(objects)
# except:
# print("\t\tERROR FILE: " + str(n) + "x" + str(m))
# #print(answer)
# continue
# object_vector = []
# time_first_action_vector = []
# time_average_vector = []
# time_full_vector = []
# average_ambiguity_vector = []
# domain = d + "/domain.pddl"
# count = 0
# for problem_n in os.listdir(d):
# #print(problem_n)
# # try:
# if(not problem_n.startswith("problem")):
# continue
# count = 1 + count
# problem_file = d + problem_n
# numbers = problem_n.split('_')
# n = int(numbers[1])
# m = int(numbers[2])
# objects = n*m
# hype = d + "hype_" + str(n) + "_" + str(m) + ".dat"
# #print("\t" +problem_file)
# problem = problem_file
# goal_set = hype
# #max_nodes = "10"
# #df = "0.5"
# #max_s_nodes = "10"
# precision = "2"
# cmd = ["./PFT.x"]
# flags = [ domain, problem, goal_set,
# # "-m", max_nodes,
# # "-D", df,
# # "-s", max_s_nodes,
# # "-P", precision,
# "-cO"]
# pipe = subprocess.Popen(
# cmd + flags,
# stdin=subprocess.PIPE, stdout=subprocess.PIPE, stderr=subprocess.STDOUT)
# answer, stderr = pipe.communicate()
# # print(answer)
# lines = str(answer).split("\\n")[2:-1]
# actions = lines[0].split(",")[:-1]
# times = list(map(float,lines[1].split(",")[:-1]))
# distances = list(map(float,lines[2].split(",")[:-1]))
# average_distance = sum(distances) / float(len(distances))
# prob_dists = []
# for line in lines[3:]:
# prob_dists += [list(map(float,line.split(",")[:-1]))]
# prob_dists = prob_dists[:-1]
# n_l = 1/len(prob_dists)
# prob_dists = [[n_l] + prob_dists[x] for x in range(len(prob_dists))]
# actions = ["I"] + actions
# total_time = sum(times)
# average_time_per_action = sum(times) / float(len(times))
# time_first_action = times[0]
# print(str(count) + ". Problem - " + str(n) + "x" + str(m) + ", Cost: " + str(average_distance) + ", " + str(time_first_action) )
# object_vector.append(objects)
# time_first_action_vector.append(time_first_action)
# time_average_vector.append(average_time_per_action)
# time_full_vector.append(total_time)
# average_ambiguity_vector.append(average_distance)
# output_dir = 'results/'
# output_file = 'online_grid.txt'
# file_out = open(output_dir + output_file , "a+")
# file_out.write(problem)
# file_out.write("\t")
# file_out.write(str(total_time))
# file_out.write("\t")
# file_out.write(str(time_first_action))
# file_out.write("\t")
# file_out.write(str(average_time_per_action))
# file_out.write("\t")
# file_out.write(str(average_distance))
# file_out.write("\n")
# file_out.write(str(objects))
# file_out.write("\n")
# file_out.close()
# # except:
# # print("\t\tERROR FILE: " + str(n) + "x" + str(m))
# # continue
# # print(object_vector)
# # print(time_first_action_vector)
# # file_out = open(output_dir + output_file , "a+")
# # file_out.write(problem)
# # file_out.write("\t")
# # file_out.write(str(total_time))
# # file_out.write("\t")
# # file_out.write(str(time_first_action))
# # file_out.write("\t")
# # file_out.write(str(average_time_per_action))
# # file_out.write("\t")
# # file_out.write(str(average_distance))
# # file_out.write("\n")
# # file_out.write(objects)
# # file_out.write("\n")
# # file_out.close()
# # print("Actions: ")
# # print(actions)
# # print()
# # print("Times: ")
# # print(times)
# # print()
# # print("Distances: ")
# # print(distances)
# # print()
# # print("Probabilities: ")
# # print(prob_dists)
# # print(prob_dists)
# # plt.figure(2)
# # axes = plt.gca()
# # axes.set_ylim([0,1])
# # axes.set_xlabel('actions')
# # axes.set_ylabel('P(G | O)')
# # counter = 0
# # print(actions)
# # print(prob_dists[0])
# # for i in prob_dists:
# # plt.plot(range(0, len(actions)), i, label=counter)
# # counter += 1
# # #plt.xticks(range(0, len(actions)-1), actions)
# # plt.legend()
# # plt.figure(1)
# # ax = plt.axes(projection='3d')
# # ax.set_ylim([0,1])
# # ax.set_xlim([0,1])
# # ax.set_zlim([0,1])
# # ax.set_xlabel('P(G2 | O)')
# # ax.set_ylabel('P(G1 | O)')
# # ax.set_zlabel('P(G* | O)')
# # G_star = 0
# # G1 = 3
# # G2 = 4
# # r = range(len(prob_dists[0]))
# # p0 = [-prob_dists[G_star][x0] + prob_dists[G_star][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# # p1 = [-prob_dists[G1][x0] + prob_dists[G1][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# # p2 = [-prob_dists[G2][x0] + prob_dists[G2][x1] for x0 in list(r) for x1 in list(r) if x0 + 1 == x1]
# # ax.plot(prob_dists[G1], prob_dists[G2], prob_dists[G_star], '-b')
# # ax.quiver(prob_dists[G1][:-1], prob_dists[G2][:-1],prob_dists[G_star][:-1], p1, p2,p0, length=0.1, pivot='tail')
# # plt.show()
# -
# fluents_l = list(map(lambda x: ((x+1)**2), object_vector_l))
# plt.figure(10)
# plt.ylim([0,25])
# plt.xlim([10,90])
# plt.xlabel('Objects')
# plt.ylabel('Time (sec)')
# #av_o = list(map(abs, average_ambiguity_vector))
# plt.scatter(fluents_l, time_first_action_vector_l)
# +
# object_vector_l = []
# time_first_action_vector_l = []
# domain = d + "domain.pddl"
# for problem_n in os.listdir(d):
# # print(problem_n)
# # try:
# if(not problem_n.startswith("problem")):
# continue
# problem_file = d + problem_n
# numbers = problem_n.split('_')
# n = int(numbers[1])
# m = int(numbers[2])
# objects = n*m
# hype = d + "hype_" + str(n) + "_" + str(m) + ".dat"
# # print("\t" +problem_file)
# problem = problem_file
# cmd = ["python"]
# flags = [ "/home/jobasha/Documents/PhD/Planners/FastDownward/fast-downward.py","--alias", "seq-sat-lama-2011", domain, problem]
# start = time.clock()
# subprocess.call(cmd + flags)
# end = time.clock()
# file = open("sas_plan.1","r")
# counter = 0
# for lines in file :
# counter = counter + 1
# print("Problem - " + str(n) + "x" + str(m) + ", Cost: " + str(counter) + ", " + str(end - start) )
# os.remove("sas_plan.1")
# os.remove("output")
# os.remove("output.sas")
# if(counter != 1):
# time_first_action_vector_l.append(end - start)
# object_vector_l.append(objects)
# except:
# continue
# -
# plt.figure(3)
# plt.scatter(object_vector_l, time_first_action_vector_l)
| on_line_planning/pt_paper/eval/backup/graphing_pft-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train and hyperparameter tune with RAPIDS
# ## Prerequisites
# - Create an Azure ML Workspace and setup environmnet on local computer following the steps in [Azure README.md](https://github.com/rapidsai/cloud-ml-examples/blob/master/azure/README.md)
# +
# verify installation and check Azure ML SDK version
import azureml.core
print('SDK version:', azureml.core.VERSION)
# -
# ## Create a FileDataset
# In this example, we will use 20 million rows (samples) of the [airline dataset](http://kt.ijs.si/elena_ikonomovska/data.html). The [FileDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) below references parquet files that have been uploaded to a public [Azure Blob storage](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-overview), you can download to your local computer or mount the files to your AML compute.
# +
from azureml.core.dataset import Dataset
airline_ds = Dataset.File.from_files('https://airlinedataset.blob.core.windows.net/airline-20m/*')
# larger dataset (10 years of airline data) is also available for multi-GPU option
# airline_ds = Dataset.File.from_files('https://airlinedataset.blob.core.windows.net/airline-10years/*')
# -
# download the dataset as local files
airline_ds.download(target_path='/local/path')
# ## Initialize workspace
# Load and initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`
# +
from azureml.core.workspace import Workspace
# if a locally-saved configuration file for the workspace is not available, use the following to load workspace
# ws = Workspace(subscription_id=subscription_id, resource_group=resource_group, workspace_name=workspace_name)
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
datastore = ws.get_default_datastore()
print("Default datastore's name: {}".format(datastore.name))
# -
# ## Upload data
# Upload the dataset to the workspace's default datastore:
# + tags=["outputPrepend"]
path_on_datastore = 'airline_data'
datastore.upload(src_dir='/local/path', target_path=path_on_datastore, overwrite=False, show_progress=True)
# -
ds_data = datastore.path(path_on_datastore)
print(ds_data)
# ## Create AML compute
# You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this notebook, we will use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training using a dynamically scalable pool of compute resources.
#
# This notebook will use 10 nodes for hyperparameter optimization, you can modify `max_node` based on available quota in the desired region. Similar to other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. [This article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) includes details on the default limits and how to request more quota.
# `vm_size` describes the virtual machine type and size that will be used in the cluster. RAPIDS requires NVIDIA Pascal or newer architecture, you will need to specify compute targets from one of `NC_v2`, `NC_v3`, `ND` or `ND_v2` [GPU virtual machines in Azure](https://docs.microsoft.com/en-us/azure/virtual-machines/sizes-gpu); these are VMs that are provisioned with P40 and V100 GPUs. Let's create an `AmlCompute` cluster of `Standard_NC6s_v3` GPU VMs:
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
gpu_cluster_name = 'gpu-cluster'
if gpu_cluster_name in ws.compute_targets:
gpu_cluster = ws.compute_targets[gpu_cluster_name]
if gpu_cluster and type(gpu_cluster) is AmlCompute:
print('Found compute target. Will use {0} '.format(gpu_cluster_name))
else:
print('creating new cluster')
# m_size parameter below could be modified to one of the RAPIDS-supported VM types
provisioning_config = AmlCompute.provisioning_configuration(vm_size = 'Standard_NC6s_v3', max_nodes = 5, idle_seconds_before_scaledown = 300)
# Use VM types with more than one GPU for multi-GPU option, e.g. Standard_NC12s_v3
# create the cluster
gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout
# if no min node count is provided it uses the scale settings for the cluster
gpu_cluster.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# use get_status() to get a detailed status for the current cluster
print(gpu_cluster.get_status().serialize())
# -
# ## Prepare training script
# Create a project directory that will contain code from your local machine that you will need access to on the remote resource. This includes the training script and additional files your training script depends on. In this example, the training script is provided:
# <br>
# `train_rapids.py` - entry script for RAPIDS Estimator that includes loading dataset into cuDF data frame, training with Random Forest and inference using cuML.
# +
import os
project_folder = './train_rapids'
os.makedirs(project_folder, exist_ok=True)
# -
# We will log some metrics by using the `Run` object within the training script:
#
# ```python
# from azureml.core.run import Run
# run = Run.get_context()
# ```
#
# We will also log the parameters and highest accuracy the model achieves:
#
# ```python
# run.log('Accuracy', np.float(accuracy))
# ```
#
# These run metrics will become particularly important when we begin hyperparameter tuning our model in the 'Tune model hyperparameters' section.
# Copy the training script `train_rapids.py` into your project directory:
notebook_path = os.path.realpath('__file__'+'/../../code')
rapids_script = os.path.join(notebook_path, 'train_rapids.py')
azure_script = os.path.join(notebook_path, 'rapids_csp_azure.py')
# +
import shutil
shutil.copy(rapids_script, project_folder)
shutil.copy(azure_script, project_folder)
# -
# ## Train model on the remote compute
# Now that you have your data and training script prepared, you are ready to train on your remote compute.
# ### Create experiment
# Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace.
# +
from azureml.core import Experiment
experiment_name = 'train_rapids'
experiment = Experiment(ws, name=experiment_name)
# -
# ### Create environment
# The [Environment class](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.environment.environment?view=azure-ml-py) allows you to build a Docker image and customize the system that you will use for training. We will build a container image using a RAPIDS container as base image and install necessary packages. This build is necessary only the first time and will take about 15 minutes. The image will be added to your Azure Container Registry and the environment will be cached after the first run, as long as the environment definition remains the same.
# +
from azureml.core import Environment
# create the environment
rapids_env = Environment('rapids_env')
# create the environment inside a Docker container
rapids_env.docker.enabled = True
# specify docker steps as a string. Alternatively, load the string from a file
dockerfile = """
FROM rapidsai/rapidsai:0.14-cuda10.2-runtime-ubuntu18.04-py3.7
RUN source activate rapids && \
pip install azureml-sdk && \
pip install azureml-widgets
"""
#FROM nvcr.io/nvidia/rapidsai/rapidsai:0.12-cuda10.0-runtime-ubuntu18.04
# set base image to None since the image is defined by dockerfile
rapids_env.docker.base_image = None
rapids_env.docker.base_dockerfile = dockerfile
# use rapids environment in the container
rapids_env.python.user_managed_dependencies = True
# +
# from azureml.core.container_registry import ContainerRegistry
# # this is an image available on Docker Hub
# image_name = 'zronaghi/rapidsai-nightly:0.13-cuda10.0-runtime-ubuntu18.04-py3.7-azuresdk-030920'
# # use rapids environment, don't build a new conda environment
# user_managed_dependencies = True
# -
# ### Create a RAPIDS Estimator
# The [Estimator](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.estimator.estimator?view=azure-ml-py) class can be used with machine learning frameworks that do not have a pre-configure estimator.
#
# `script_params` is a dictionary of command-line arguments to pass to the training script.
# +
from azureml.train.estimator import Estimator
script_params = {
'--data_dir': ds_data.as_mount(),
'--n_bins': 32,
}
estimator = Estimator(source_directory=project_folder,
script_params=script_params,
compute_target=gpu_cluster,
entry_script='train_rapids.py',
environment_definition=rapids_env)
# custom_docker_image=image_name,
# user_managed=user_managed_dependencies
# -
# ## Tune model hyperparameters
# We can optimize our model's hyperparameters and improve the accuracy using Azure Machine Learning's hyperparameter tuning capabilities.
# ### Start a hyperparameter sweep
# Let's define the hyperparameter space to sweep over. We will tune `n_estimators`, `max_depth` and `max_features` parameters. In this example we will use random sampling to try different configuration sets of hyperparameters and maximize `Accuracy`.
# +
from azureml.train.hyperdrive.runconfig import HyperDriveConfig
from azureml.train.hyperdrive.sampling import RandomParameterSampling
from azureml.train.hyperdrive.run import PrimaryMetricGoal
from azureml.train.hyperdrive.parameter_expressions import choice, loguniform, uniform
param_sampling = RandomParameterSampling( {
'--n_estimators': choice(range(50, 500)),
'--max_depth': choice(range(5, 19)),
'--max_features': uniform(0.2, 1.0)
}
)
hyperdrive_run_config = HyperDriveConfig(estimator=estimator,
hyperparameter_sampling=param_sampling,
primary_metric_name='Accuracy',
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=10,
max_concurrent_runs=5)
# -
# This will launch the RAPIDS training script with parameters that were specified in the cell above.
# start the HyperDrive run
hyperdrive_run = experiment.submit(hyperdrive_run_config)
# ## Monitor HyperDrive runs
# Monitor and view the progress of the machine learning training run with a [Jupyter widget](https://docs.microsoft.com/en-us/python/api/azureml-widgets/azureml.widgets?view=azure-ml-py).The widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
# +
from azureml.widgets import RunDetails
RunDetails(hyperdrive_run).show()
# + tags=["outputPrepend"]
hyperdrive_run.wait_for_completion(show_output=True)
# +
# hyperdrive_run.cancel()
# -
# ### Find and register best model
best_run = hyperdrive_run.get_best_run_by_primary_metric()
print(best_run.get_details()['runDefinition']['arguments'])
# List the model files uploaded during the run:
print(best_run.get_file_names())
# Register the folder (and all files in it) as a model named `train-rapids` under the workspace for deployment
# +
# model = best_run.register_model(model_name='train-rapids', model_path='outputs/model-rapids.joblib')
# -
# ## Delete cluster
# +
# delete the cluster
# gpu_cluster.delete()
| azure/notebooks/HPO-RAPIDS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: conda-env-anaconda3-py
# ---
# 03 Using Nipype to load fMRI data
# =====================
# #### Date: Feb 12 2018; Author: Farahana
# ## Data Acquisition and Simple Preprocessing
# *Part 1*: Data Acquisition for anatomical and functional NifTi images
#
# *Part 2*: Simple preprocessing
#
# *Part 3*: Getting the output of pre-processing steps (Datasink and connect the workflows.)
# +
# Base packages
# %pylab inline
import numpy as np
#import matplotlib.pyplot as plt
import pandas as pd
# Neuroimaging packages
import nibabel as nib
from nilearn.plotting import plot_img, plot_anat, plot_stat_map
from nipype import SelectFiles, Node, DataSink
from nipype.pipeline.engine import Workflow, MapNode
# We will import the pre-processing process in their parts.
# -
# #### Checking the packages version
import sys
import os
import nilearn
print("Python version:", sys.version.split()[0])
print("Nibabel version: ", nib.__version__)
print("Nilearn version:", nilearn.__version__)
# #### Some helper functions to be used
# +
# Simple viewer
def plot_nii(in_file):
nii = nib.load(in_file)
nii.orthoview()
# Let's create a short helper function to plot 3D NIfTI images
def plot_slice(fname, slice_no):
# Load the image
img = nib.load(fname)
data = img.get_data()
# Cut in the middle of the brain
cut = int(data.shape[-2]/2)
# Plot the data
imshow(np.rot90(data[:, :, cut, slice_no]), cmap="gray")
gca().set_axis_off()
# -
# ### Part 1: Data Acquisition for anatomical and functional NifTi images
# +
# The template string
templates = { 'anat' : 'sub*/anatomy/highres001.nii*',
'func' : 'sub*/BOLD/task001_run{ses_no}/bold.nii*'}
# How to address and import using SelectFiles node
sf = Node(SelectFiles(templates),
name='selectfiles')
sf.inputs.base_directory = '/home/farahana/Documents/dataset/Multi_Subject/ds117'
# -
# Choose the first run for subjects imports
sf.inputs.ses_no = "001"
# To view the import, use
# ~~~
# sf.run().outputs
# ~~~
#
# However, we will run this using the connect/workflow of Nipype together
# with other pre-processing steps.
# Visualizing the input data folder
# !tree /home/farahana/Documents/dataset/Multi_Subject/ds117
# ### Part 2: Simple preprocessing
# *Step 1*: Brain extractions
#
# *Step 2*: Realignment
# +
# Import BET from the FSL interface
from nipype.interfaces.fsl import (BET, IsotropicSmooth, ExtractROI, FAST, FLIRT, ImageMaths,
MCFLIRT, SliceTimer, Threshold)
from nipype.interfaces.spm import Smooth
from nipype.interfaces.utility import IdentityInterface
# Import Retroicor from the SPM interface
from nipype.interfaces.spm import Realign
# -
# #### Let us try with 1 subject 1 task pre-processing
# Define the subject chosen and initialize it
input_file = "/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/BOLD/task001_run001/bold.nii"
# +
skullstrip_sub3 = BET(in_file = input_file,
out_file = "output/sub-03_brain.nii.gz",
functional= True,
mask = True)
# Run this to get the output for the input of next step
# %time skullstrip_sub3.run()
# +
smooth_sub3 = IsotropicSmooth(in_file = "output/sub-03_brain.nii.gz",
out_file = "output/sub-03_smooth.nii.gz",
fwhm=4)
# %time smooth_sub3.run()
# +
motcor_sub3 = MCFLIRT(in_file = "output/sub-03_smooth.nii.gz",
out_file = "output/sub-03_mcflirt.nii.gz",
cost='mutualinfo')
# %time motcor_sub3.run()
# -
f = plt.figure(figsize=(12, 4))
for i, img in enumerate(["bold", "brain","smooth", "mcflirt"]):
f.add_subplot(1, 4, i + 1)
if i == 0:
plot_slice("/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/BOLD/task001_run001/%s.nii" % img, 1)
else:
plot_slice("output/sub-03_%s.nii.gz" % img, 1)
plt.title(img)
# #### Now, let us try with many subjects
# Initiation of a workflow
wf = Workflow(name="preproc", base_dir="/output/working_dir")
# +
# Workflow of pre-processing steps
skullstrip = Node(BET(output_type='NIFTI_GZ',
name="skullstrip"))
smooth = Node(IsotropicSmooth(output_type='NIFTI_GZ',
name="smooth"))
wf.connect([()])
| 03-Nipype.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BZylN3BRIp9S"
# ## Dependency Parsing
# + id="JGRcIQs8GtjT"
import spacy
nlp = spacy.load('en_core_web_sm')
# + id="zrb-zMT8Iuer"
sentence = 'Smith jumps over the lazy dog.'
doc = nlp(sentence)
# + colab={"base_uri": "https://localhost:8080/"} id="7sNLPtBGI6lq" outputId="4918d037-caf7-403e-fa69-c9a63e4843b3"
[(token.text, token.tag_, token.dep_, token.head) for token in doc]
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="pJkEaTajJhjq" outputId="47047e54-a956-4238-ed17-02638d9f15c1"
spacy.displacy.render(doc,style='dep',jupyter=True)
| s8/codes/s8d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Building your own algorithm container
#
# With Amazon SageMaker, you can package your own algorithms that can than be trained and deployed in the SageMaker environment. This notebook will guide you through an example that shows you how to build a Docker container for SageMaker and use it for training and inference.
#
# By packaging an algorithm in a container, you can bring almost any code to the Amazon SageMaker environment, regardless of programming language, environment, framework, or dependencies.
#
# _**Note:**_ SageMaker now includes a [pre-built scikit container](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/sagemaker-python-sdk/scikit_learn_iris/Scikit-learn%20Estimator%20Example%20With%20Batch%20Transform.ipynb). We recommend the pre-built container be used for almost all cases requiring a scikit algorithm. However, this example remains relevant as an outline for bringing in other libraries to SageMaker as your own container.
#
# 1. [Building your own algorithm container](#Building-your-own-algorithm-container)
# 1. [When should I build my own algorithm container?](#When-should-I-build-my-own-algorithm-container%3F)
# 1. [Permissions](#Permissions)
# 1. [The example](#The-example)
# 1. [The presentation](#The-presentation)
# 1. [Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker](#Part-1%3A-Packaging-and-Uploading-your-Algorithm-for-use-with-Amazon-SageMaker)
# 1. [An overview of Docker](#An-overview-of-Docker)
# 1. [How Amazon SageMaker runs your Docker container](#How-Amazon-SageMaker-runs-your-Docker-container)
# 1. [Running your container during training](#Running-your-container-during-training)
# 1. [The input](#The-input)
# 1. [The output](#The-output)
# 1. [Running your container during hosting](#Running-your-container-during-hosting)
# 1. [The parts of the sample container](#The-parts-of-the-sample-container)
# 1. [The Dockerfile](#The-Dockerfile)
# 1. [Building and registering the container](#Building-and-registering-the-container)
# 1. [Testing your algorithm on your local machine or on an Amazon SageMaker notebook instance](#Testing-your-algorithm-on-your-local-machine-or-on-an-Amazon-SageMaker-notebook-instance)
# 1. [Part 2: Using your Algorithm in Amazon SageMaker](#Part-2%3A-Using-your-Algorithm-in-Amazon-SageMaker)
# 1. [Set up the environment](#Set-up-the-environment)
# 1. [Create the session](#Create-the-session)
# 1. [Upload the data for training](#Upload-the-data-for-training)
# 1. [Create an estimator and fit the model](#Create-an-estimator-and-fit-the-model)
# 1. [Hosting your model](#Hosting-your-model)
# 1. [Deploy the model](#Deploy-the-model)
# 2. [Choose some data and use it for a prediction](#Choose-some-data-and-use-it-for-a-prediction)
# 3. [Optional cleanup](#Optional-cleanup)
# 1. [Run Batch Transform Job](#Run-Batch-Transform-Job)
# 1. [Create a Transform Job](#Create-a-Transform-Job)
# 2. [View Output](#View-Output)
#
# _or_ I'm impatient, just [let me see the code](#The-Dockerfile)!
#
# ## When should I build my own algorithm container?
#
# You may not need to create a container to bring your own code to Amazon SageMaker. When you are using a framework (such as Apache MXNet or TensorFlow) that has direct support in SageMaker, you can simply supply the Python code that implements your algorithm using the SDK entry points for that framework. This set of frameworks is continually expanding, so we recommend that you check the current list if your algorithm is written in a common machine learning environment.
#
# Even if there is direct SDK support for your environment or framework, you may find it more effective to build your own container. If the code that implements your algorithm is quite complex on its own or you need special additions to the framework, building your own container may be the right choice.
#
# If there isn't direct SDK support for your environment, don't worry. You'll see in this walk-through that building your own container is quite straightforward.
#
# ## Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` permissions. This is because we'll creating new repositories in Amazon ECR. The easiest way to add these permissions is simply to add the managed policy `AmazonEC2ContainerRegistryFullAccess` to the role that you used to start your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
#
# ## The example
#
# Here, we'll show how to package a simple Python example which showcases the [decision tree][] algorithm from the widely used [scikit-learn][] machine learning package. The example is purposefully fairly trivial since the point is to show the surrounding structure that you'll want to add to your own code so you can train and host it in Amazon SageMaker.
#
# The ideas shown here will work in any language or environment. You'll need to choose the right tools for your environment to serve HTTP requests for inference, but good HTTP environments are available in every language these days.
#
# In this example, we use a single image to support training and hosting. This is easy because it means that we only need to manage one image and we can set it up to do everything. Sometimes you'll want separate images for training and hosting because they have different requirements. Just separate the parts discussed below into separate Dockerfiles and build two images. Choosing whether to have a single image or two images is really a matter of which is more convenient for you to develop and manage.
#
# If you're only using Amazon SageMaker for training or hosting, but not both, there is no need to build the unused functionality into your container.
#
# [scikit-learn]: http://scikit-learn.org/stable/
# [decision tree]: http://scikit-learn.org/stable/modules/tree.html
#
# ## The presentation
#
# This presentation is divided into two parts: _building_ the container and _using_ the container.
# # Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker
#
# ### An overview of Docker
#
# If you're familiar with Docker already, you can skip ahead to the next section.
#
# For many data scientists, Docker containers are a new concept, but they are not difficult, as you'll see here.
#
# Docker provides a simple way to package arbitrary code into an _image_ that is totally self-contained. Once you have an image, you can use Docker to run a _container_ based on that image. Running a container is just like running a program on the machine except that the container creates a fully self-contained environment for the program to run. Containers are isolated from each other and from the host environment, so the way you set up your program is the way it runs, no matter where you run it.
#
# Docker is more powerful than environment managers like conda or virtualenv because (a) it is completely language independent and (b) it comprises your whole operating environment, including startup commands, environment variable, etc.
#
# In some ways, a Docker container is like a virtual machine, but it is much lighter weight. For example, a program running in a container can start in less than a second and many containers can run on the same physical machine or virtual machine instance.
#
# Docker uses a simple file called a `Dockerfile` to specify how the image is assembled. We'll see an example of that below. You can build your Docker images based on Docker images built by yourself or others, which can simplify things quite a bit.
#
# Docker has become very popular in the programming and devops communities for its flexibility and well-defined specification of the code to be run. It is the underpinning of many services built in the past few years, such as [Amazon ECS].
#
# Amazon SageMaker uses Docker to allow users to train and deploy arbitrary algorithms.
#
# In Amazon SageMaker, Docker containers are invoked in a certain way for training and a slightly different way for hosting. The following sections outline how to build containers for the SageMaker environment.
#
# Some helpful links:
#
# * [Docker home page](http://www.docker.com)
# * [Getting started with Docker](https://docs.docker.com/get-started/)
# * [Dockerfile reference](https://docs.docker.com/engine/reference/builder/)
# * [`docker run` reference](https://docs.docker.com/engine/reference/run/)
#
# [Amazon ECS]: https://aws.amazon.com/ecs/
#
# ### How Amazon SageMaker runs your Docker container
#
# Because you can run the same image in training or hosting, Amazon SageMaker runs your container with the argument `train` or `serve`. How your container processes this argument depends on the container:
#
# * In the example here, we don't define an `ENTRYPOINT` in the Dockerfile so Docker will run the command `train` at training time and `serve` at serving time. In this example, we define these as executable Python scripts, but they could be any program that we want to start in that environment.
# * If you specify a program as an `ENTRYPOINT` in the Dockerfile, that program will be run at startup and its first argument will be `train` or `serve`. The program can then look at that argument and decide what to do.
# * If you are building separate containers for training and hosting (or building only for one or the other), you can define a program as an `ENTRYPOINT` in the Dockerfile and ignore (or verify) the first argument passed in.
#
# #### Running your container during training
#
# When Amazon SageMaker runs training, your `train` script is run just like a regular Python program. A number of files are laid out for your use, under the `/opt/ml` directory:
#
# /opt/ml
# ├── input
# │ ├── config
# │ │ ├── hyperparameters.json
# │ │ └── resourceConfig.json
# │ └── data
# │ └── <channel_name>
# │ └── <input data>
# ├── model
# │ └── <model files>
# └── output
# └── failure
#
# ##### The input
#
# * `/opt/ml/input/config` contains information to control how your program runs. `hyperparameters.json` is a JSON-formatted dictionary of hyperparameter names to values. These values will always be strings, so you may need to convert them. `resourceConfig.json` is a JSON-formatted file that describes the network layout used for distributed training. Since scikit-learn doesn't support distributed training, we'll ignore it here.
# * `/opt/ml/input/data/<channel_name>/` (for File mode) contains the input data for that channel. The channels are created based on the call to CreateTrainingJob but it's generally important that channels match what the algorithm expects. The files for each channel will be copied from S3 to this directory, preserving the tree structure indicated by the S3 key structure.
# * `/opt/ml/input/data/<channel_name>_<epoch_number>` (for Pipe mode) is the pipe for a given epoch. Epochs start at zero and go up by one each time you read them. There is no limit to the number of epochs that you can run, but you must close each pipe before reading the next epoch.
#
# ##### The output
#
# * `/opt/ml/model/` is the directory where you write the model that your algorithm generates. Your model can be in any format that you want. It can be a single file or a whole directory tree. SageMaker will package any files in this directory into a compressed tar archive file. This file will be available at the S3 location returned in the `DescribeTrainingJob` result.
# * `/opt/ml/output` is a directory where the algorithm can write a file `failure` that describes why the job failed. The contents of this file will be returned in the `FailureReason` field of the `DescribeTrainingJob` result. For jobs that succeed, there is no reason to write this file as it will be ignored.
#
# #### Running your container during hosting
#
# Hosting has a very different model than training because hosting is reponding to inference requests that come in via HTTP. In this example, we use our recommended Python serving stack to provide robust and scalable serving of inference requests:
#
# 
#
# This stack is implemented in the sample code here and you can mostly just leave it alone.
#
# Amazon SageMaker uses two URLs in the container:
#
# * `/ping` will receive `GET` requests from the infrastructure. Your program returns 200 if the container is up and accepting requests.
# * `/invocations` is the endpoint that receives client inference `POST` requests. The format of the request and the response is up to the algorithm. If the client supplied `ContentType` and `Accept` headers, these will be passed in as well.
#
# The container will have the model files in the same place they were written during training:
#
# /opt/ml
# └── model
# └── <model files>
#
#
# ### The parts of the sample container
#
# In the `container` directory are all the components you need to package the sample algorithm for Amazon SageMager:
#
# .
# ├── Dockerfile
# ├── build_and_push.sh
# └── decision_trees
# ├── nginx.conf
# ├── predictor.py
# ├── serve
# ├── train
# └── wsgi.py
#
# Let's discuss each of these in turn:
#
# * __`Dockerfile`__ describes how to build your Docker container image. More details below.
# * __`build_and_push.sh`__ is a script that uses the Dockerfile to build your container images and then pushes it to ECR. We'll invoke the commands directly later in this notebook, but you can just copy and run the script for your own algorithms.
# * __`decision_trees`__ is the directory which contains the files that will be installed in the container.
# * __`local_test`__ is a directory that shows how to test your new container on any computer that can run Docker, including an Amazon SageMaker notebook instance. Using this method, you can quickly iterate using small datasets to eliminate any structural bugs before you use the container with Amazon SageMaker. We'll walk through local testing later in this notebook.
#
# In this simple application, we only install five files in the container. You may only need that many or, if you have many supporting routines, you may wish to install more. These five show the standard structure of our Python containers, although you are free to choose a different toolset and therefore could have a different layout. If you're writing in a different programming language, you'll certainly have a different layout depending on the frameworks and tools you choose.
#
# The files that we'll put in the container are:
#
# * __`nginx.conf`__ is the configuration file for the nginx front-end. Generally, you should be able to take this file as-is.
# * __`predictor.py`__ is the program that actually implements the Flask web server and the decision tree predictions for this app. You'll want to customize the actual prediction parts to your application. Since this algorithm is simple, we do all the processing here in this file, but you may choose to have separate files for implementing your custom logic.
# * __`serve`__ is the program started when the container is started for hosting. It simply launches the gunicorn server which runs multiple instances of the Flask app defined in `predictor.py`. You should be able to take this file as-is.
# * __`train`__ is the program that is invoked when the container is run for training. You will modify this program to implement your training algorithm.
# * __`wsgi.py`__ is a small wrapper used to invoke the Flask app. You should be able to take this file as-is.
#
# In summary, the two files you will probably want to change for your application are `train` and `predictor.py`.
# ### The Dockerfile
#
# The Dockerfile describes the image that we want to build. You can think of it as describing the complete operating system installation of the system that you want to run. A Docker container running is quite a bit lighter than a full operating system, however, because it takes advantage of Linux on the host machine for the basic operations.
#
# For the Python science stack, we will start from a standard Ubuntu installation and run the normal tools to install the things needed by scikit-learn. Finally, we add the code that implements our specific algorithm to the container and set up the right environment to run under.
#
# Along the way, we clean up extra space. This makes the container smaller and faster to start.
#
# Let's look at the Dockerfile for the example:
# !cat container/Dockerfile
# ### Building and registering the container
#
# The following shell code shows how to build the container image using `docker build` and push the container image to ECR using `docker push`. This code is also available as the shell script `container/build-and-push.sh`, which you can run as `build-and-push.sh decision_trees_sample` to build the image `decision_trees_sample`.
#
# This code looks for an ECR repository in the account you're using and the current default region (if you're using a SageMaker notebook instance, this will be the region where the notebook instance was created). If the repository doesn't exist, the script will create it.
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=sagemaker-word2vec
#
# cd container
#
# chmod +x decision_trees/train
# chmod +x decision_trees/serve
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
# region=${region:-us-east-1}
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email)
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full name.
#
# docker build -t ${algorithm_name} .
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# ## Testing your algorithm on your local machine or on an Amazon SageMaker notebook instance
#
# While you're first packaging an algorithm use with Amazon SageMaker, you probably want to test it yourself to make sure it's working right. In the directory `container/local_test`, there is a framework for doing this. It includes three shell scripts for running and using the container and a directory structure that mimics the one outlined above.
#
# The scripts are:
#
# * `train_local.sh`: Run this with the name of the image and it will run training on the local tree. For example, you can run `$ ./train_local.sh sagemaker-word2vec`. It will generate a model under the `/test_dir/model` directory. You'll want to modify the directory `test_dir/input/data/...` to be set up with the correct channels and data for your algorithm. Also, you'll want to modify the file `input/config/hyperparameters.json` to have the hyperparameter settings that you want to test (as strings).
# * `serve_local.sh`: Run this with the name of the image once you've trained the model and it should serve the model. For example, you can run `$ ./serve_local.sh sagemaker-word2vec`. It will run and wait for requests. Simply use the keyboard interrupt to stop it.
# * `predict.sh`: Run this with the name of a payload file and (optionally) the HTTP content type you want. The content type will default to `text/csv`. For example, you can run `$ ./predict.sh payload.csv text/csv`.
#
# The directories as shipped are set up to test the decision trees sample algorithm presented here.
# # Part 2: Using your Algorithm in Amazon SageMaker
#
# Once you have your container packaged, you can use it to train models and use the model for hosting or batch transforms. Let's do that with the algorithm we made above.
#
# ## Set up the environment
#
# Here we specify a bucket to use and the role that will be used for working with SageMaker.
# +
# S3 prefix
prefix = 'word2vec'
# Define IAM role
import boto3
import re
import os
import numpy as np
import pandas as pd
from sagemaker import get_execution_role
role = get_execution_role()
# -
# ## Create the session
#
# The session remembers our connection parameters to SageMaker. We'll use it to perform all of our SageMaker operations.
# +
import sagemaker as sage
from time import gmtime, strftime
sess = sage.Session()
# -
# ## Upload the data for training
#
# When training large models with huge amounts of data, you'll typically use big data tools, like Amazon Athena, AWS Glue, or Amazon EMR, to create your data in S3. For the purposes of this example, we're using some the classic [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set), which we have included.
#
# We can use use the tools provided by the SageMaker Python SDK to upload the data to a default bucket.
# +
WORK_DIRECTORY = 'data'
data_location = sess.upload_data(WORK_DIRECTORY, key_prefix=prefix)
# -
# tar the model file:
# !tar -czvf model.tar.gz ./word2vec_2.model
# +
# tree = ./model.tar.gz
# -
# ## Create an estimator and fit the model
#
# In order to use SageMaker to fit our algorithm, we'll create an `Estimator` that defines how to use the container to train. This includes the configuration we need to invoke SageMaker training:
#
# * The __container name__. This is constructed as in the shell commands above.
# * The __role__. As defined above.
# * The __instance count__ which is the number of machines to use for training.
# * The __instance type__ which is the type of machine to use for training.
# * The __output path__ determines where the model artifact will be written.
# * The __session__ is the SageMaker session object that we defined above.
#
# Then we use fit() on the estimator to train against the data that we uploaded above.
# +
account = sess.boto_session.client('sts').get_caller_identity()['Account']
region = sess.boto_session.region_name
image = '{}.dkr.ecr.{}.amazonaws.com/sagemaker-word2vec:latest'.format(account, region)
tree = sage.estimator.Estimator(image,
role, 1, 'ml.c4.2xlarge',
output_path="s3://{}/output".format(sess.default_bucket()),
sagemaker_session=sess)
tree.fit(data_location)
# -
# ## Hosting your model
# You can use a trained model to get real time predictions using HTTP endpoint. Follow these steps to walk you through the process.
# ### Deploy the model
#
# Deploying the model to SageMaker hosting just requires a `deploy` call on the fitted model. This call takes an instance count, instance type, and optionally serializer and deserializer functions. These are used when the resulting predictor is created on the endpoint.
from sagemaker.predictor import csv_serializer
predictor = tree.deploy(1, 'ml.m4.xlarge', serializer=csv_serializer)
# ### Choose some data and use it for a prediction
#
# In order to do some predictions, we'll extract some of the data we used for training and do predictions against it. This is, of course, bad statistical practice, but a good way to see how the mechanism works.
df=pd.read_csv("data/word2vec_test_data.csv", header=None)
df.sample(3)
#
df = df['movieid']
# +
import itertools
list = df.values.tolist()
# -
list
# Prediction is as easy as calling predict with the predictor we got back from deploy and the data we want to do predictions with. The serializers take care of doing the data conversions for us.
print(predictor.predict(list.values).decode('utf-8'))
# ### Optional cleanup
# When you're done with the endpoint, you'll want to clean it up.
# +
# sess.delete_endpoint(predictor.endpoint)
# -
# ## Run Batch Transform Job
# You can use a trained model to get inference on large data sets by using [Amazon SageMaker Batch Transform](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-batch.html). A batch transform job takes your input data S3 location and outputs the predictions to the specified S3 output folder. Similar to hosting, you can extract inferences for training data to test batch transform.
# ### Create a Transform Job
# We'll create an `Transformer` that defines how to use the container to get inference results on a data set. This includes the configuration we need to invoke SageMaker batch transform:
#
# * The __instance count__ which is the number of machines to use to extract inferences
# * The __instance type__ which is the type of machine to use to extract inferences
# * The __output path__ determines where the inference results will be written
# +
# transform_output_folder = "batch-transform-output"
# output_path="s3://{}/{}".format(sess.default_bucket(), transform_output_folder)
# transformer = tree.transformer(instance_count=1,
# instance_type='ml.m4.xlarge',
# output_path=output_path,
# assemble_with='Line',
# accept='text/csv')
# -
# We use tranform() on the transfomer to get inference results against the data that we uploaded. You can use these options when invoking the transformer.
#
# * The __data_location__ which is the location of input data
# * The __content_type__ which is the content type set when making HTTP request to container to get prediction
# * The __split_type__ which is the delimiter used for splitting input data
# * The __input_filter__ which indicates the first column (ID) of the input will be dropped before making HTTP request to container
# +
# transformer.transform(data_location, content_type='text/csv', split_type='Line', input_filter='$[1:]')
# transformer.wait()
# -
# For more information on the configuration options, see [CreateTransformJob API](https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateTransformJob.html)
# ### View Output
# Lets read results of above transform job from s3 files and print output
# +
# s3_client = sess.boto_session.client('s3')
# s3_client.download_file(sess.default_bucket(), "{}/iris.csv.out".format(transform_output_folder), '/tmp/iris.csv.out')
# with open('/tmp/iris.csv.out') as f:
# results = f.readlines()
# print("Transform results: \n{}".format(''.join(results)))
| SageMaker/word2vec_bring_my_own.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
import plotly
plotly.__version__
# #### United States Bubble Map
#
# Note about `sizeref`:
#
# To scale the bubble size, use the attribute sizeref. We recommend using the following formula to calculate a sizeref value:
#
# `sizeref = 2. * max(array of size values) / (desired maximum marker size ** 2)`
#
# Note that setting `sizeref` to a value greater than $1$, decreases the rendered marker sizes, while setting `sizeref` to less than $1$, increases the rendered marker sizes.
#
# See https://plot.ly/python/reference/#scatter-marker-sizeref for more information. Additionally, we recommend setting the sizemode attribute: https://plot.ly/python/reference/#scatter-marker-sizemode to area.
# +
import plotly.plotly as py
import plotly.graph_objs as go
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2014_us_cities.csv')
df.head()
df['text'] = df['name'] + '<br>Population ' + (df['pop']/1e6).astype(str)+' million'
limits = [(0,2),(3,10),(11,20),(21,50),(50,3000)]
colors = ["rgb(0,116,217)","rgb(255,65,54)","rgb(133,20,75)","rgb(255,133,27)","lightgrey"]
cities = []
scale = 5000
for i in range(len(limits)):
lim = limits[i]
df_sub = df[lim[0]:lim[1]]
city = go.Scattergeo(
locationmode = 'USA-states',
lon = df_sub['lon'],
lat = df_sub['lat'],
text = df_sub['text'],
marker = go.scattergeo.Marker(
size = df_sub['pop']/scale,
color = colors[i],
line = go.scattergeo.marker.Line(
width=0.5, color='rgb(40,40,40)'
),
sizemode = 'area'
),
name = '{0} - {1}'.format(lim[0],lim[1]) )
cities.append(city)
layout = go.Layout(
title = go.layout.Title(
text = '2014 US city populations<br>(Click legend to toggle traces)'
),
showlegend = True,
geo = go.layout.Geo(
scope = 'usa',
projection = go.layout.geo.Projection(
type='albers usa'
),
showland = True,
landcolor = 'rgb(217, 217, 217)',
subunitwidth=1,
countrywidth=1,
subunitcolor="rgb(255, 255, 255)",
countrycolor="rgb(255, 255, 255)"
)
)
fig = go.Figure(data=cities, layout=layout)
py.iplot(fig, filename='d3-bubble-map-populations')
# -
# #### Ebola Cases in West Africa
# +
import plotly.plotly as py
import plotly.graph_objs as go
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2014_ebola.csv')
df.head()
cases = []
colors = ['rgb(239,243,255)','rgb(189,215,231)','rgb(107,174,214)','rgb(33,113,181)']
months = {6:'June',7:'July',8:'Aug',9:'Sept'}
for i in range(6,10)[::-1]:
cases.append(go.Scattergeo(
lon = df[ df['Month'] == i ]['Lon'], #-(max(range(6,10))-i),
lat = df[ df['Month'] == i ]['Lat'],
text = df[ df['Month'] == i ]['Value'],
name = months[i],
marker = go.scattergeo.Marker(
size = df[ df['Month'] == i ]['Value']/50,
color = colors[i-6],
line = go.scattergeo.marker.Line(width = 0)
)
)
)
cases[0]['text'] = df[ df['Month'] == 9 ]['Value'].map('{:.0f}'.format).astype(str)+' '+\
df[ df['Month'] == 9 ]['Country']
cases[0]['mode'] = 'markers+text'
cases[0]['textposition'] = 'bottom center'
inset = [
go.Choropleth(
locationmode = 'country names',
locations = df[ df['Month'] == 9 ]['Country'],
z = df[ df['Month'] == 9 ]['Value'],
text = df[ df['Month'] == 9 ]['Country'],
colorscale = [[0,'rgb(0, 0, 0)'],[1,'rgb(0, 0, 0)']],
autocolorscale = False,
showscale = False,
geo = 'geo2'
),
go.Scattergeo(
lon = [21.0936],
lat = [7.1881],
text = ['Africa'],
mode = 'text',
showlegend = False,
geo = 'geo2'
)
]
layout = go.Layout(
title = go.layout.Title(
text = 'Ebola cases reported by month in West Africa 2014<br> \
Source: <a href="https://data.hdx.rwlabs.org/dataset/rowca-ebola-cases">\
HDX</a>'),
geo = go.layout.Geo(
resolution = 50,
scope = 'africa',
showframe = False,
showcoastlines = True,
showland = True,
landcolor = "rgb(229, 229, 229)",
countrycolor = "rgb(255, 255, 255)" ,
coastlinecolor = "rgb(255, 255, 255)",
projection = go.layout.geo.Projection(
type = 'mercator'
),
lonaxis = go.layout.geo.Lonaxis(
range= [ -15.0, -5.0 ]
),
lataxis = go.layout.geo.Lataxis(
range= [ 0.0, 12.0 ]
),
domain = go.layout.geo.Domain(
x = [ 0, 1 ],
y = [ 0, 1 ]
)
),
geo2 = go.layout.Geo(
scope = 'africa',
showframe = False,
showland = True,
landcolor = "rgb(229, 229, 229)",
showcountries = False,
domain = go.layout.geo.Domain(
x = [ 0, 0.6 ],
y = [ 0, 0.6 ]
),
bgcolor = 'rgba(255, 255, 255, 0.0)',
),
legend = go.layout.Legend(
traceorder = 'reversed'
)
)
fig = go.Figure(layout=layout, data=cases+inset)
py.iplot(fig, filename='West Africa Ebola cases 2014')
# -
# #### Reference
# See https://plot.ly/python/reference/#choropleth and https://plot.ly/python/reference/#scattergeo for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'bubble-maps.ipynb', 'python/bubble-maps/', 'Bubble Maps',
'How to make bubble maps in Python with Plotly.',
title = 'Python Bubble Maps | Plotly',
has_thumbnail='true', thumbnail='thumbnail/bubble-map.jpg',
language='python',
page_type='example_index',
display_as='maps', order=3,
uses_plotly_offline=False)
# -
| _posts/python-v3/maps/bubble-maps/bubble-maps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zkufh760uvF3"
# 
#
# [](https://colab.research.google.com/github/JohnSnowLabs/nlu/blob/master/examples/colab/Training/binary_text_classification/NLU_training_sentiment_classifier_demo_finanical_news.ipynb)
#
#
#
# # Training a Sentiment Analysis Classifier with NLU
# ## 2 class Finance News sentiment classifier training
# With the [SentimentDL model](https://nlp.johnsnowlabs.com/docs/en/annotators#sentimentdl-multi-class-sentiment-analysis-annotator) from Spark NLP you can achieve State Of the Art results on any multi class text classification problem
#
# This notebook showcases the following features :
#
# - How to train the deep learning classifier
# - How to store a pipeline to disk
# - How to load the pipeline from disk (Enables NLU offline mode)
#
#
# + [markdown] id="dur2drhW5Rvi"
# # 1. Install Java 8 and NLU
# + id="hFGnBCHavltY"
import os
from sklearn.metrics import classification_report
# ! apt-get update -qq > /dev/null
# Install java
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# ! pip install nlu pyspark==2.4.7 > /dev/null
import nlu
# + [markdown] id="f4KkTfnR5Ugg"
# # 2. Download Finanical News Sentiment dataset
# https://www.kaggle.com/ankurzing/sentiment-analysis-for-financial-news
#
# This dataset contains the sentiments for financial news headlines from the perspective of a retail investor. Further details about the dataset can be found in: <NAME>., <NAME>., <NAME>., <NAME>. and <NAME>. (2014): “Good debt or bad debt: Detecting semantic orientations in economic texts.” Journal of the American Society for Information Science and Technology.
# + colab={"base_uri": "https://localhost:8080/"} id="OrVb5ZMvvrQD" executionInfo={"status": "ok", "timestamp": 1610788018304, "user_tz": -300, "elapsed": 2399, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="f494fab0-8f9c-4087-f554-31a21764a207"
# ! wget http://ckl-it.de/wp-content/uploads/2021/01/all-data.csv
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="y4xSRWIhwT28" executionInfo={"status": "ok", "timestamp": 1610788018314, "user_tz": -300, "elapsed": 660, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="e1e2496a-8df8-4e5d-db53-63d62ef1f050"
import pandas as pd
train_path = '/content/all-data.csv'
train_df = pd.read_csv(train_path)
# the text data to use for classification should be in a column named 'text'
# the label column must have name 'y' name be of type str
columns=['text','y']
train_df = train_df[columns]
train_df = train_df[~train_df["y"].isin(["neutral"])]
train_df
# + [markdown] id="0296Om2C5anY"
# # 3. Train Deep Learning Classifier using nlu.load('train.sentiment')
#
# You dataset label column should be named 'y' and the feature column with text data should be named 'text'
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3ZIPkRkWftBG" executionInfo={"status": "ok", "timestamp": 1609264914996, "user_tz": -300, "elapsed": 191025, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="6dc536e4-252e-4324-e070-cd477a79330d"
import nlu
# load a trainable pipeline by specifying the train. prefix and fit it on a datset with label and text columns
# by default the Universal Sentence Encoder (USE) Sentence embeddings are used for generation
trainable_pipe = nlu.load('train.sentiment')
fitted_pipe = trainable_pipe.fit(train_df.iloc[:50])
# predict with the trainable pipeline on dataset and get predictions
preds = fitted_pipe.predict(train_df.iloc[:50],output_level='document')
#sentence detector that is part of the pipe generates sone NaNs. lets drop them first
preds.dropna(inplace=True)
print(classification_report(preds['y'], preds['sentiment']))
preds
# + [markdown] id="lVyOE2wV0fw_"
# # Test the fitted pipe on new example
# + colab={"base_uri": "https://localhost:8080/", "height": 107} id="qdCUg2MR0PD2" executionInfo={"status": "ok", "timestamp": 1609264917602, "user_tz": -300, "elapsed": 193623, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="8fe5b9aa-c87a-42d3-e00d-920e63ca6aa4"
fitted_pipe.predict('According to the most recent update there has been a major decrese in the rate of oil')
# + [markdown] id="xflpwrVjjBVD"
# ## Configure pipe training parameters
# + colab={"base_uri": "https://localhost:8080/"} id="UtsAUGTmOTms" executionInfo={"status": "ok", "timestamp": 1609264917604, "user_tz": -300, "elapsed": 193620, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="ac9c8b1a-7fdd-4a6f-bdfd-1dbb823d9bf4"
trainable_pipe.print_info()
# + [markdown] id="2GJdDNV9jEIe"
# ## Retrain with new parameters
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="mptfvHx-MMMX" executionInfo={"status": "ok", "timestamp": 1609264924472, "user_tz": -300, "elapsed": 200484, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="1dd94bc8-09c8-45db-ab81-bbd64acb8a4b"
# Train longer!
trainable_pipe['sentiment_dl'].setMaxEpochs(5)
fitted_pipe = trainable_pipe.fit(train_df.iloc[:100])
# predict with the trainable pipeline on dataset and get predictions
preds = fitted_pipe.predict(train_df.iloc[:100],output_level='document')
#sentence detector that is part of the pipe generates sone NaNs. lets drop them first
preds.dropna(inplace=True)
print(classification_report(preds['y'], preds['sentiment']))
preds
# + [markdown] id="qFoT-s1MjTSS"
# # Try training with different Embeddings
# + id="nxWFzQOhjWC8" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609264924477, "user_tz": -300, "elapsed": 200483, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="e259763c-470b-4d46-b3d1-28cf545f5dcd"
# We can use nlu.print_components(action='embed_sentence') to see every possibler sentence embedding we could use. Lets use bert!
nlu.print_components(action='embed_sentence')
# + id="IKK_Ii_gjJfF" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609266286092, "user_tz": -300, "elapsed": 1562094, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="4237752f-4fbe-4235-b33d-5d7b8ba29d48"
trainable_pipe = nlu.load('en.embed_sentence.small_bert_L12_768 train.sentiment')
# We need to train longer and user smaller LR for NON-USE based sentence embeddings usually
# We could tune the hyperparameters further with hyperparameter tuning methods like gridsearch
# Also longer training gives more accuracy
trainable_pipe['sentiment_dl'].setMaxEpochs(70)
trainable_pipe['sentiment_dl'].setLr(0.0005)
fitted_pipe = trainable_pipe.fit(train_df)
# predict with the trainable pipeline on dataset and get predictions
preds = fitted_pipe.predict(train_df,output_level='document')
#sentence detector that is part of the pipe generates sone NaNs. lets drop them first
preds.dropna(inplace=True)
print(classification_report(preds['y'], preds['sentiment']))
#preds
# + [markdown] id="2BB-NwZUoHSe"
# # 5. Lets save the model
# + id="eLex095goHwm" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609266449598, "user_tz": -300, "elapsed": 1725594, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="b31b5e1e-3f09-4ab3-e97a-fb32ac87b319"
stored_model_path = './models/classifier_dl_trained'
fitted_pipe.save(stored_model_path)
# + [markdown] id="e_b2DPd4rCiU"
# # 6. Lets load the model from HDD.
# This makes Offlien NLU usage possible!
# You need to call nlu.load(path=path_to_the_pipe) to load a model/pipeline from disk.
# + id="SO4uz45MoRgp" colab={"base_uri": "https://localhost:8080/", "height": 124} executionInfo={"status": "ok", "timestamp": 1609266465229, "user_tz": -300, "elapsed": 1741220, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="5d9cc34a-693c-44d7-e50a-6e0ca5d4e024"
hdd_pipe = nlu.load(path=stored_model_path)
preds = hdd_pipe.predict('According to the most recent update there has been a major decrese in the rate of oil')
preds
# + id="e0CVlkk9v6Qi" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609266465232, "user_tz": -300, "elapsed": 1741218, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02458088882398909889"}} outputId="ec54f7c0-8174-4fd4-9db8-51c1d15be3eb"
hdd_pipe.print_info()
| examples/colab/Training/binary_text_classification/NLU_training_sentiment_classifier_demo_finanical_news.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.3 ('autumn310')
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import os
import warnings
from math import ceil
from summer.utils import ref_times_to_dti
from autumn.core.project import get_project, load_timeseries
from autumn.core.plots.utils import REF_DATE
from autumn.core.runs.managed import ManagedRun
from autumn.core.plots.calibration.plots import plot_prior
# -
warnings.filterwarnings("ignore", category=DeprecationWarning)
# ## Specify the run id and the outputs to plot
# +
run_id = "hierarchical_sir/multi/1653962883/0e2003c"
BURN_IN = 5000
# Outputs requested for full run plots
outputs_for_full_run_plots = (
"incidence",
"incidence_AUS",
"incidence_ITA",
)
# Outputs requested with uncertainty
outputs_to_plot_with_uncertainty = (
"incidence_AUS",
"incidence_ITA",
)
scenarios_to_plot_with_uncertainty = [0]
# +
param_lookup = {
"contact_rate": "infection risk per contact",
"testing_to_detection.assumed_cdr_parameter": "CDR at one test per 1,000 population per day",
"voc_emergence.omicron.new_voc_seed.start_time": "Omicron emergence date",
"voc_emergence.omicron.contact_rate_multiplier": "relative transmissibility Omicron",
"age_stratification.cfr.multiplier": "modification to fatality rate",
"age_stratification.prop_hospital.multiplier": "modification to hospitalisation rate",
}
title_lookup = {
"notifications": "daily notifications",
"infection_deaths": "COVID-19-specific deaths",
"hospital_admissions": "new daily hospital admissions",
"icu_admissions": "new daily admissions to ICU",
"proportion_seropositive": "proportion recovered from COVID-19",
"incidence": "daily new infections",
"prop_incidence_strain_delta": "proportion of cases due to Delta",
"hospital_admissions": "daily hospital admissions",
"hospital_occupancy": "total hospital beds",
"icu_admissions": "daily ICU admissions",
"icu_occupancy": "total ICU beds",
"prop_ever_infected": "ever infected with Delta or Omicron",
"cdr": "case detection rate",
}
# -
# ## Load run outputs and pre-process data
model, region = run_id.split("/")[0:2]
mr = ManagedRun(run_id)
full_run = mr.full_run.get_derived_outputs()
pbi = mr.powerbi.get_db()
targets = pbi.get_targets()
results = pbi.get_uncertainty()
mcmc_params = mr.calibration.get_mcmc_params()
mcmc_runs = mr.calibration.get_mcmc_runs()
# +
project = get_project(model, region, reload=True)
params_list = list(mcmc_params.columns)
chains = mcmc_runs.chain.unique()
mcmc_table = mcmc_params.merge(mcmc_runs, on=["urun"])
post_burnin_uruns = mcmc_runs[mcmc_runs["run"] >= BURN_IN].index
post_burnin_mcmc_table = mcmc_table.filter(items=post_burnin_uruns, axis=0)
# -
# ## Plot posteriors
# +
plt.style.use("ggplot")
n_params = len(params_list)
n_col = 3
n_row = ceil(n_params / n_col)
fig, axes = plt.subplots(n_row, n_col, figsize=(n_col * 4, n_row * 3.5))
for i_ax, axis in enumerate(axes.reshape(-1)):
if i_ax >= n_params:
axis.set_visible(False)
else:
param = mcmc_params.columns[i_ax]
prior_dict = [d for d in project.calibration.all_priors if d['param_name'] == param][0]
if prior_dict['distri_params'] is not None:
plot_prior(0, prior_dict, ax=axis, print_distri=False, alpha=.5)
axis.hist(post_burnin_mcmc_table[param], weights=post_burnin_mcmc_table["weight"], density=True, bins=15, color='coral')
par_name = param if param not in param_lookup else param_lookup[param]
axis.set_title(par_name)
axis.set_xlabel(par_name)
fig.suptitle("parameter posterior histograms", fontsize=15, y=1)
fig.tight_layout()
# -
# ## Plot traces
# +
n_col = 2
n_row = ceil(n_params / n_col)
fig, axes = plt.subplots(n_row, n_col, figsize=(8*n_col, 4*n_row))
for i_ax, axis in enumerate(axes.reshape(-1)):
if i_ax >= n_params:
axis.set_visible(False)
else:
param = params_list[i_ax]
for chain in chains:
chain_filter = mcmc_table["chain"] == chain
axis.plot(mcmc_table[chain_filter]['run'], mcmc_table[chain_filter][param], lw=.5)
par_name = param if param not in param_lookup else param_lookup[param]
axis.set_title(par_name)
# -
# # Model outputs
# ### Get all targets, including those not used in calibration, to use as a validation
all_targets = load_timeseries(os.path.join(project.get_path(), "timeseries.json"))
for target in all_targets:
all_targets[target].index = ref_times_to_dti(REF_DATE, all_targets[target].index)
# ### Calibration fits with individual model runs
n_outputs = len(outputs_for_full_run_plots)
n_col = 2
n_row = ceil(n_outputs / n_col)
fig, axes = plt.subplots(n_row, n_col, figsize=(n_col * 8, n_row * 6), sharex="all")
for i_ax, axis in enumerate(axes.reshape(-1)):
if i_ax >= n_outputs:
axis.set_visible(False)
else:
output = outputs_for_full_run_plots[i_ax]
scenario_chain = (full_run["scenario"] == 0) & (full_run["chain"] == 0)
for i_run in full_run[scenario_chain]["run"].unique():
selection = full_run[(full_run["run"] == i_run) & scenario_chain]
axis.plot(ref_times_to_dti(REF_DATE, selection["times"]), selection[output])
if output in all_targets and len(all_targets[output]) > 0:
all_targets[output].plot.line(ax=axis, linewidth=0., markersize=10., marker="o")
axis.scatter(all_targets[output].index, all_targets[output], color="k", s=5, alpha=0.5, zorder=10)
if output in targets:
axis.scatter(targets.index, targets[output], facecolors="r", edgecolors="k", s=15, zorder=10)
axis.tick_params(axis="x", labelrotation=45)
title = output if output not in title_lookup else title_lookup[output]
axis.set_title(title)
# ### Calibration fits with uncertainty
# +
colours = ((0.2, 0.2, 0.8), (0.8, 0.2, 0.2), (0.2, 0.8, 0.2), (0.8, 0.8, 0.2), (0.8, 0.2, 0.2), (0.2, 0.8, 0.2), (0.8, 0.8, 0.2))
def plot_outputs_with_uncertainty(outputs, scenarios = [0]):
n_outputs = len(outputs)
n_col = 2
n_row = ceil(n_outputs / n_col)
fig, axes = plt.subplots(n_row, n_col, figsize=(n_col * 8, n_row * 6), sharex="all")
for i_ax, axis in enumerate(axes.reshape(-1)):
if i_ax >= n_outputs:
axis.set_visible(False)
else:
output = outputs[i_ax]
for scenario in scenarios:
colour = colours[scenario]
results_df = results[(output, scenario)]
indices = results_df.index
interval_label = "baseline" if scenario == 0 else project.param_set.scenarios[scenario - 1]["description"]
scenario_zorder = 10 if scenario == 0 else scenario
axis.fill_between(
indices,
results_df[0.025], results_df[0.975],
color=colour,
alpha=0.5,
label="_nolegend_",
zorder=scenario_zorder,
)
axis.fill_between(
indices,
results_df[0.25], results_df[0.75],
color=colour, alpha=0.7,
label=interval_label,
zorder=scenario_zorder
)
axis.plot(indices, results_df[0.500], color=colour)
if output in all_targets and len(all_targets[output]) > 0:
all_targets[output].plot.line(
ax=axis,
linewidth=0.,
markersize=8.,
marker="o",
markerfacecolor="w",
markeredgecolor="w",
alpha=0.2,
label="_nolegend_",
zorder=11,
)
if output in targets:
targets[output].plot.line(
ax=axis,
linewidth=0.,
markersize=5.,
marker="o",
markerfacecolor="k",
markeredgecolor="k",
label="_nolegend_",
zorder=12,
)
axis.tick_params(axis="x", labelrotation=45)
title = output if output not in title_lookup else title_lookup[output]
axis.set_title(title)
if i_ax == 0:
axis.legend()
fig.tight_layout()
# -
plot_outputs_with_uncertainty(outputs_to_plot_with_uncertainty, scenarios_to_plot_with_uncertainty)
| notebooks/user/rragonnet/managed_runs_generic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Hyperoxia and Organ Dysfunction
# ## <NAME> for Team 11
# ## II MIT-HIAE Datathon
#
# The overall goal is to assess the association between hyperoxia in the first 24 hours of mechanical ventilation and markers of organ dysfunction which, for this event, will be $\Delta$SOFA.
#
# We will start by extracting out the base ICU cohort and then identifying the `itemid` that correspond to $SpO_2$ values.
#
# __TODO:__ Change how SOFA day 1 and 2 are pulled in using the pivoted tables.
#
# ## 0 - Envrionment
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import statsmodels.formula.api as smf
import psycopg2
dbname = 'mimic'
schema_name = 'mimiciii'
db_schema = 'SET search_path TO {0};'.format(schema_name)
con = psycopg2.connect(database=dbname)
# -
# ## 1 - Cohort Extraction
#
# A lot of the desired concepts already exist from previous work either directly in the database or via materialized views. We will extract as much as we can this way.
query = db_schema + '''
-- note that this code uses a lot of local materialized veiws
-- the SQL queries for this will be loaded on the repository
-- and the views need to be built
-- TODO: combine this queries into one ventilator parameters query
-- as this is just a sloppy way with copy and pasting.
WITH tidal_volume_24h AS(
WITH tidal_volume AS (
SELECT ce.icustay_id, ce.charttime - ie.intime AS offset
, ce.valuenum
FROM icustays ie
LEFT JOIN chartevents ce
ON ie.icustay_id = ce.icustay_id
WHERE ce.itemid = 224685
)
SELECT tv.icustay_id, MAX(tv.valuenum) AS tv_max_24h
FROM tidal_volume tv
WHERE tv.offset <= interval '24' hour
GROUP BY tv.icustay_id
)
, peep_24h AS (
WITH peep AS (
SELECT ce.icustay_id, ce.charttime - ie.intime AS offset
, ce.valuenum
FROM icustays ie
LEFT JOIN chartevents ce
ON ie.icustay_id = ce.icustay_id
WHERE ce.itemid = 220339
)
SELECT peep.icustay_id, MAX(peep.valuenum) AS peep_max_24h
FROM peep
WHERE peep.offset <= interval '24' hour
GROUP BY peep.icustay_id
)
, fio2_24h AS(
WITH fio2 AS (
SELECT ce.icustay_id, ce.charttime - ie.intime AS offset
, ce.valuenum
FROM icustays ie
LEFT JOIN chartevents ce
ON ie.icustay_id = ce.icustay_id
WHERE ce.itemid = 223835
)
SELECT fio2.icustay_id, MAX(fio2.valuenum) AS fio2_max_24h
FROM fio2
WHERE fio2.offset <= interval '24' hour
GROUP BY fio2.icustay_id
)
SELECT ie.icustay_id, ie.hadm_id, ie.subject_id, ie.dbsource
, ie.first_careunit, ie.intime, ie.outtime, ie.los
, ied.admission_age, ied.gender, ied.ethnicity, v.resprate_max
, tv.tv_max_24h, pp.peep_max_24h, fi.fio2_max_24h -- vent paramaters
, ied.first_icu_stay, oa.oasis AS oasis_score
, s1.sofa AS sofa_day1, s2.sofa AS sofa_day2
, l1.lactate_max AS lactate_max_day1, l2.lactate_max AS lactate_max_day2
, elixdx.congestive_heart_failure, elixdx.cardiac_arrhythmias --- start of comorbidities
, elixdx.valvular_disease, elixdx.pulmonary_circulation
, elixdx.peripheral_vascular, elixdx.hypertension
, elixdx.paralysis, elixdx.other_neurological
, elixdx.chronic_pulmonary, elixdx.diabetes_uncomplicated
, elixdx.diabetes_complicated, elixdx.hypothyroidism
, elixdx.renal_failure, elixdx.liver_disease
, elixdx.peptic_ulcer, elixdx.aids
, elixdx.lymphoma, elixdx.metastatic_cancer
, elixdx.solid_tumor, elixdx.rheumatoid_arthritis
, elixdx.coagulopathy, elixdx.obesity
, elixdx.weight_loss, elixdx.fluid_electrolyte
, elixdx.blood_loss_anemia, elixdx.deficiency_anemias
, elixdx.alcohol_abuse, elixdx.drug_abuse
, elixdx.psychoses, elixdx.depression -- end of comorbidities
, elixscore.elixhauser_vanwalraven AS elixhauser_score -- score for comorbid
, vd.starttime AS vent_start, vd.endtime AS vent_end
, vd.duration_hours AS ventduration, ad.hospital_expire_flag
FROM icustays ie
LEFT JOIN icustay_detail ied
ON ie.icustay_id = ied.icustay_id
LEFT JOIN admissions ad
ON ie.hadm_id = ad.hadm_id
LEFT JOIN vitalsfirstday v
ON ie.icustay_id = v.icustay_id
LEFT JOIN labsfirstday l1
ON ie.icustay_id = l1.icustay_id
LEFT JOIN labssecondday l2
ON ie.icustay_id = l2.icustay_id
LEFT JOIN tidal_volume_24h tv
ON ie.icustay_id = tv.icustay_id
LEFT JOIN peep_24h pp
ON ie.icustay_id = pp.icustay_id
LEFT JOIN fio2_24h fi
ON ie.icustay_id = fi.icustay_id
LEFT JOIN elixhauser_ahrq_score elixscore
ON ie.hadm_id = elixscore.hadm_id
LEFT JOIN elixhauser_ahrq_no_drg_all_icd elixdx
ON ie.hadm_id = elixdx.hadm_id
LEFT JOIN oasis oa
ON ie.icustay_id = oa.icustay_id
LEFT JOIN sofa_firstday s1
ON ie.icustay_id = s1.icustay_id
LEFT JOIN sofa_secondday s2
ON ie.icustay_id = s2.icustay_id
LEFT JOIN ventdurations vd
ON ie.icustay_id = vd.icustay_id
AND vd.ventnum = 1 -- we only take first vent period because we only care about first 24 hour
WHERE ie.dbsource = 'metavision';
'''
cohort_df = pd.read_sql(query, con)
print(cohort_df.shape)
display(cohort_df.head())
# We'll do a quick missingness check for sanity.
sns.heatmap(cohort_df.reset_index().isna(), cbar=False)
# This pattern of missingness seems reasonable; patients missing vent start/end are missing vent parameters because they likely weren't on the ventilator.
# ## 2 - Extracting $SpO_2$ and Feature Engineering Hyperoxic Time
#
# We start by identifying the metavision ID for $SpO_2$.
# +
query = db_schema + '''
SELECT itemid, label, dbsource, linksto
FROM d_items
WHERE LOWER(label) LIKE '%o2%'
AND dbsource='metavision';
'''
d_search = pd.read_sql_query(query, con)
display(d_search)
# -
# It is `itemid` 2202777. We then extract all of these values from `chartevents` in the interval [0, 24h].
# +
query = db_schema + '''
WITH spo2_day1 AS (
SELECT ce.icustay_id, ce.charttime - ie.intime AS offset
, ce.value
FROM icustays ie
LEFT JOIN chartevents ce
ON ie.icustay_id = ce.icustay_id
WHERE ce.itemid = 220277
)
SELECT sp.icustay_id, sp.value AS spo2_24h, sp.offset
FROM spo2_day1 sp
WHERE sp.offset <= interval '24' hour
AND sp.offset >= interval '0' hour
ORDER BY sp.offset ASC;
'''
sp_df = pd.read_sql(query, con)
display(sp_df.head())
# -
# After which we do some brief data cleaning.
sp_df.spo2_24h = sp_df.spo2_24h.astype(float)
sp_df = sp_df.loc[sp_df.spo2_24h <= 100, :]
# We can visualize an example $SpO_2$ trend.
example_patient = sp_df.loc[sp_df.icustay_id == 241249, :].copy()
example_patient.loc[:, 'offset'] = example_patient.offset / np.timedelta64(1, 'ns')
example_patient.loc[:, 'spo2_24h'] = example_patient.spo2_24h.astype(float)
sns.lineplot(x=example_patient.offset, y=example_patient.spo2_24h)
sns.lineplot(x=[example_patient.offset.min(), example_patient.offset.max()], y=[95.99,95.99])
plt.savefig('example1.png')
# We will want to integrate the peaks above the line designating normoxia; we can visualize this first.
example_patient.loc[:, 'spo2_24h'] = example_patient.spo2_24h - 95.99
example_patient.loc[example_patient.spo2_24h < 0, 'spo2_24h'] = 0
sns.lineplot(x=example_patient.offset, y=example_patient.spo2_24h)
plt.savefig('example2.png')
# We will write a function to integrate the area under these peaks for each patient.
# +
def hyperoxic_time(time, spo2):
spo2 = spo2 - 95.99
spo2[spo2 < 0] = 0
if (spo2 > 0).any():
return np.trapz(y=spo2, x=time) / np.timedelta64(1, 'ns') / (3.6*10**12)
else:
return 0
hyperoxia_24h = [(i, hyperoxic_time(sp_df.loc[sp_df.icustay_id == i, 'offset'], sp_df.loc[sp_df.icustay_id == i, 'spo2_24h']))\
for i in sp_df.icustay_id.unique()]
hyperoxia_24h = pd.DataFrame(hyperoxia_24h)
hyperoxia_24h = hyperoxia_24h.rename({0 : 'icustay_id', 1 : 'hyperoxia_24h'}, axis=1)
hyperoxia_24h.head()
# -
# We then visualize the distribution of hyperoxic time in the first 24 hours to see if the result seems reasonable.
hyperoxia_24h.hyperoxia_24h.hist()
# We then add this feature to our cohort data.
cohort_df = cohort_df.merge(right=hyperoxia_24h, how='left', on='icustay_id')
print(cohort_df.shape)
display(cohort_df.head())
# This looks how we would expect. The next steps are joining this feature to the cleaning, exploration, and ultimately model building.
# ## 3 - Exploration and Cleaning
#
# General Plan:
# * Calculate $\Delta$SOFA
# * Exclude patients with no SOFA on the second day / no $\Delta$SOFA (same difference)
# * Exclude everyone younger than 16
# * Exclude $F_iO_2$ $\leq$ 21%
# * Require 24 hours on vent and vent start within 4 hours of ICU admission
# * Use only first ICU stay
# * Fix the ages greater than 89 (which are converted to 300 for HIPPA)
# * Code gender as binary
# * Limit our cohort to only patients who have been mechanically ventilated
# * Build a simple model
#
#
# __Problem:__
# * It is unclear how best to model the $\Delta$SOFA; linear regression is not appropriate
# * Alistair thinks the best approach would be Probit Ordered Regression
# * This is not easily available in `Python` and we'll probably have to use `R`
# * __Conclusion: For the datathon binarize around a positive $\Delta$SOFA and use Logistic Regression__
#
# So we'll knock out the above todos, and then just viusalize the exposure variable (likely to not show much). Then we'll build a few simple models to present at the end.
delta_sofa = cohort_df.sofa_day2 - cohort_df.sofa_day1
cohort_df = cohort_df.assign(delta_sofa=delta_sofa)
eda_df = cohort_df.loc[~pd.isna(cohort_df.delta_sofa), :].copy()
eda_df.gender = (eda_df.gender == 'M').astype(int)
eda_df = eda_df.loc[cohort_df.admission_age >= 16, :]
eda_df.loc[eda_df.admission_age > 90, 'age'] = 91.3
eda_df = eda_df.loc[eda_df.los >= 0.167, :] # redundant because we require SOFA day 2
eda_df = eda_df.loc[eda_df.chronic_pulmonary == 0, :]
eda_df = eda_df.loc[eda_df.ventduration > 0, :]
eda_df = eda_df.loc[eda_df.fio2_max_24h > 21, :]
eda_df = eda_df.loc[eda_df.ventduration >= 24, :]
eda_df = eda_df.loc[eda_df.first_icu_stay, :]
vent_near_intime = (eda_df.vent_start < eda_df.intime + np.timedelta64('240', 'm')) | (eda_df.vent_start > eda_df.intime - np.timedelta64('240', 'm'))
eda_df = eda_df.loc[vent_near_intime, :]
label = (eda_df.delta_sofa > 0).astype(int)
eda_df = eda_df.assign(label=label)
print(eda_df.shape)
sns.violinplot(x='label', y='hyperoxia_24h', data=eda_df)
sns.distplot(a=eda_df.loc[eda_df.label == 0, 'hyperoxia_24h'].dropna())
sns.distplot(a=eda_df.loc[eda_df.label == 1, 'hyperoxia_24h'].dropna())
sns.violinplot(x='label', y='hyperoxia_24h', hue='gender', split=True, data=eda_df)
# ## 4 - Modeling
#
# As said above, to do this for publication we'll need ordinal probit regression. For this event we'll binarize the $\Delta$SOFA outcome and sfit a logistic model for positive change in $\Delta$SOFA. We will also look at max lactate on the second day as a continuous outcome with OLS.
logit = smf.logit(formula='label ~ hyperoxia_24h', data=eda_df)
result = logit.fit()
result.summary()
logit = smf.logit(formula='label ~ hyperoxia_24h + admission_age + gender + oasis_score + elixhauser_score', data=eda_df)
result = logit.fit()
result.summary()
logit = smf.logit(formula='label ~ hyperoxia_24h + admission_age + gender + oasis_score + elixhauser_score + resprate_max + peep_max_24h + tv_max_24h', data=eda_df)
result = logit.fit()
result.summary()
lm = smf.ols(formula='lactate_max_day2 ~ hyperoxia_24h', data=eda_df)
result = lm.fit()
result.summary()
lm = smf.ols(formula='lactate_max_day2 ~ hyperoxia_24h + lactate_max_day1 + oasis_score + elixhauser_score + admission_age + gender', data=eda_df)
result = lm.fit()
result.summary()
| hyperoxia-orgdys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
# # Logistická regresia
#
# Dáta:
#
# https://www.kaggle.com/code/mnassrib/titanic-logistic-regression-with-python/data?select=train.csv
# https://www.kaggle.com/code/mnassrib/titanic-logistic-regression-with-python/data?select=test.csv
# https://www.kaggle.com/code/mnassrib/titanic-logistic-regression-with-python/data?select=gender_submission.csv
#
# Veľmi dobre popísaný celý postup:
#
# https://www.kaggle.com/code/mnassrib/titanic-logistic-regression-with-python/notebook
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
data = pd.read_csv("train.csv")
data.head()
data.isnull()
sns.heatmap(data.isnull(),yticklabels=False,cbar=False,cmap="viridis")
sns.countplot(x="Survived",hue="Pclass",data=data)
data["Age"].plot.hist(bins=30)
data.info()
data.describe()
data["Fare"].plot.hist(bins=30)
sns.boxplot(x="Pclass",y="Age",data=data)
def imput_age(cols):
Age = cols[0]
Pclass = cols[1]
if pd.isnull(Age):
if Pclass == 1:
return 37
elif Pclass == 2:
return 29
else:
return 25
else:
return Age
data["Age"] = data[["Age","Pclass"]].apply(imput_age,axis=1)
sns.heatmap(data.isnull(),yticklabels=False,cbar=False,cmap="viridis")
data.drop("Cabin",axis=1,inplace=True)
data.dropna(inplace=True)
sex = pd.get_dummies(data["Sex"],drop_first=True)
embark = pd.get_dummies(data["Embarked"],drop_first=True)
data = pd.concat([data,sex,embark],axis=1)
data.drop(["PassengerId","Sex","Name","Ticket","Embarked"],axis=1,inplace=True)
data.head()
from sklearn.model_selection import train_test_split
X = data.drop(["Survived"],axis=1)
y = data["Survived"]
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3,random_state=101)
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression(max_iter=3000)
logmodel.fit(X_train,y_train)
predictions = logmodel.predict(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions))
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_test, predictions))
X_test = X_test.reset_index(drop=True)
for i, v in enumerate(y_test):
if v != list(predictions)[i]:
print(X_test.loc[[i]])
novy = pd.DataFrame([{"Pclass":2,"Age":30,"SibSp":0,"Parch":1,"Fare":70,"male":1,"Q":0,"S":1}])
novy
pred = logmodel.predict(novy)
p = logmodel.predict_proba(novy)
print(p)
| logisticka-regresia/logisticka-regresia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from math import ceil
from CIoTS import *
from tqdm import trange
runs = 20
max_p = 22
dimensions = 3
data_length = 10000
alpha = 0.05
ics = ["bic"]
p_estimations = pd.DataFrame(columns=['p' ,'method', 'mean_p', 'std_p'])
scores = pd.DataFrame(columns=['p' ,'method', 'mean_f1', 'std_f1', 'mean_recall', 'std_recall',
'mean_precision', 'std_precision', 'mean_fpr', 'std_fpr'])
for p in trange(2, max_p, 2):
incoming_edges = 3
f1 = {ic: [] for ic in ics}
f1.update({'incremental_' + ic: [] for ic in ics})
f1['real'] = []
precision = {ic: [] for ic in ics}
precision.update({'incremental_' + ic: [] for ic in ics})
precision['real'] = []
recall = {ic: [] for ic in ics}
recall.update({'incremental_' + ic: [] for ic in ics})
recall['real'] = []
fpr = {ic: [] for ic in ics}
fpr.update({'incremental_' + ic: [] for ic in ics})
fpr['real'] = []
p_est = {ic: [] for ic in ics}
p_est.update({'incremental_' + ic: [] for ic in ics})
for i in trange(runs):
generator = CausalTSGenerator(dimensions=dimensions, max_p=p, data_length=data_length,
incoming_edges=incoming_edges)
ts = generator.generate()
predicted_graph = pc_chen_modified(partial_corr_test, ts, p, alpha)
eval_result = evaluate_edges(generator.graph, predicted_graph)
f1['real'].append(eval_result['f1-score'])
recall['real'].append(eval_result['TPR'])
fpr['real'].append(eval_result['FPR'])
precision['real'].append(eval_result['precision'])
# VAR estimation
var_ranking, var_scores = var_order_select(ts, 2*(max_p-2), ics)
for ic in ics:
predicted_graph = pc_chen_modified(partial_corr_test, ts, var_ranking[ic][0], alpha)
eval_result = evaluate_edges(generator.graph, predicted_graph)
p_est[ic].append(var_ranking[ic][0])
f1[ic].append(eval_result['f1-score'])
recall[ic].append(eval_result['TPR'])
fpr[ic].append(eval_result['FPR'])
precision[ic].append(eval_result['precision'])
# incremental
for ic in ics:
predicted_graph = pc_incremental_extensive(partial_corr_test, ts, 0.05, 2*(max_p-2), ic=ic)
eval_result = evaluate_edges(generator.graph, predicted_graph)
p_est['incremental_' + ic].append(len(predicted_graph.nodes())/dimensions - 1)
f1['incremental_' + ic].append(eval_result['f1-score'])
recall['incremental_' + ic].append(eval_result['TPR'])
fpr['incremental_' + ic].append(eval_result['FPR'])
precision['incremental_' + ic].append(eval_result['precision'])
scores = scores.append({'p': p, 'method': 'real',
'mean_f1': np.mean(f1['real']), 'std_f1': np.std(f1['real']),
'mean_recall': np.mean(recall['real']), 'std_recall': np.std(recall['real']),
'mean_precision': np.mean(precision['real']), 'std_precision': np.std(precision['real']),
'mean_fpr': np.mean(fpr['real']), 'std_fpr': np.std(fpr['real'])},
ignore_index=True)
for ic in ics:
p_estimations = p_estimations.append({'p': p, 'method': ic, 'mean_p': np.mean(p_est[ic]),
'std_p': np.std(p_est[ic])},
ignore_index=True)
scores = scores.append({'p': p, 'method': ic,
'mean_f1': np.mean(f1[ic]), 'std_f1': np.std(f1[ic]),
'mean_recall': np.mean(recall[ic]), 'std_recall': np.std(recall[ic]),
'mean_precision': np.mean(precision[ic]), 'std_precision': np.std(precision[ic]),
'mean_fpr': np.mean(fpr[ic]), 'std_fpr': np.std(fpr[ic])},
ignore_index=True)
for ic in ics:
p_estimations = p_estimations.append({'p': p, 'method': 'incremental_' + ic,
'mean_p': np.mean(p_est['incremental_' + ic]),
'std_p': np.std(p_est['incremental_' + ic])},
ignore_index=True)
scores = scores.append({'p': p, 'method': 'incremental_' + ic,
'mean_f1': np.mean(f1['incremental_' + ic]), 'std_f1': np.std(f1['incremental_' + ic]),
'mean_recall': np.mean(recall['incremental_' + ic]),
'std_recall': np.std(recall['incremental_' + ic]),
'mean_precision': np.mean(precision['incremental_' + ic]),
'std_precision': np.std(precision['incremental_' + ic]),
'mean_fpr': np.mean(fpr['incremental_' + ic]),
'std_fpr': np.std(fpr['incremental_' + ic])},
ignore_index=True)
scores.to_csv('results/extensive/scores.csv')
p_estimations.to_csv('results/extensive/p_estimations.csv')
| first_experiments_extensive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# !pip install -q torch==1.4.0
# !pip install -q transformers==2.5.1
# +
from transformers import pipeline
# Allocate a pipeline for sentiment-analysis
nlp = pipeline('sentiment-analysis')
nlp('We are very happy to include pipeline into the transformers repository.')
# -
| 07_train/archive/bert/99_Predict_Reviews_Transformers_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="DEsLktBKRtLF" colab_type="text"
# # BYOL-Pytorch
# Pytorch Implementation of BYOL: Bootstrap Your Own Latent: A New Approach to Self-Supervised Learning (https://arxiv.org/abs/2006.07733).
# Major part of Code is inspired from https://github.com/sthalles/PyTorch-BYOL.
# The Code has more appropriate Naming Convention.
# # Default Training
# * Running the Python File without any changes trains BYOL with **CIFAR10** Dataset.
# * All the Parameters are contained in ___Params Object___ in the script.
# # Custom Training
# * Change the __Dataset Object__.
# * Update the Required Parameters in the ___Params Object___.
#
# + [markdown] id="Iguz4yzHRuOI" colab_type="text"
# ## Import Statement
# + id="JXwnvO2WFr5N" colab_type="code" colab={}
import os
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data.dataloader
from torchvision import datasets
from torchvision.transforms import transforms
import torchvision.models
import cv2
import numpy as np
from tqdm import tqdm
# + id="gcTb54y7Fr5U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3708d645-f203-47a1-cf03-c48b989f87f6"
np.random.seed(0)
torch.manual_seed(42)
# + [markdown] id="OjicKwA5IS4a" colab_type="text"
# # Augmentation Functions
# Given an image, augmentation is applied to create two different views. Augmentation used over here is very similar to that of [SimCLR](https://arxiv.org/abs/2002.05709).
# ### GaussianBlur(Class)
# 
# * **Parameter** = Kernel Size
# * **Output** = Gaussian Blur Transformed Image
# ### Transforms(Function)
# * **Parameter** = Input Dimension of the Image.
# * **Output** = Composes a torchvision.transforms Object with all the Transformation functions intact.
# ### MultiViewDataInjector(Class)
# * **Parameter** = Input Image.
# * **Output** = Applies **Transforms** Function to result two different augmented image.
#
#
# + id="O1ws4HuZFr5d" colab_type="code" colab={}
class GaussianBlur(object):
"""blur a single image on CPU"""
def __init__(self, kernel_size):
radias = kernel_size // 2
kernel_size = radias * 2 + 1
self.blur_h = nn.Conv2d(3, 3, kernel_size=(kernel_size, 1),
stride=1, padding=0, bias=False, groups=3)
self.blur_v = nn.Conv2d(3, 3, kernel_size=(1, kernel_size),
stride=1, padding=0, bias=False, groups=3)
self.k = kernel_size
self.r = radias
self.blur = nn.Sequential(
nn.ReflectionPad2d(radias),
self.blur_h,
self.blur_v
)
self.pil_to_tensor = transforms.ToTensor()
self.tensor_to_pil = transforms.ToPILImage()
def __call__(self, img):
img = self.pil_to_tensor(img).unsqueeze(0)
sigma = np.random.uniform(0.1, 2.0)
x = np.arange(-self.r, self.r + 1)
x = np.exp(-np.power(x, 2) / (2 * sigma * sigma))
x = x / x.sum()
x = torch.from_numpy(x).view(1, -1).repeat(3, 1)
self.blur_h.weight.data.copy_(x.view(3, 1, self.k, 1))
self.blur_v.weight.data.copy_(x.view(3, 1, 1, self.k))
with torch.no_grad():
img = self.blur(img)
img = img.squeeze()
img = self.tensor_to_pil(img)
return img
# + id="zJYmcyhVFr5i" colab_type="code" colab={}
def Transforms(Input_Dim,S=1):
Color_Jitter = transforms.ColorJitter(0.8*S,0.8*S,0.8*S,0.2*S)
Data_Transforms = transforms.Compose([transforms.RandomResizedCrop(size=Input_Dim[0]),
transforms.RandomHorizontalFlip(),
transforms.RandomApply([Color_Jitter],p=0.75),
transforms.RandomGrayscale(p=0.2),
GaussianBlur(int(0.1*Input_Dim[0])),
transforms.ToTensor(),
])
return Data_Transforms
# + id="30KwuU7oFr5m" colab_type="code" colab={}
class MultiViewDataInjector(object):
def __init__(self,Transforms):
self.transforms = Transforms
def __call__(self,Sample,*Consistent_Flip):
if Consistent_Flip:
Sample = torchvision.transforms.RandomHorizontalFlip()
Output = [transforms(Sample) for transforms in self.transforms]
return Output
# + [markdown] id="BDHuM4ssFr5q" colab_type="text"
# # Model
# Contains two basic Neural Networks
# * **MLP_Base** - Creates the **Latent Space** from the Encoder.
# * **Skeleton Net** - Encompases MLP_BASE for **Latent Space** creation and uses **ResNet18** to learn Feature Representations.
#
# + id="LSCHcyc6Fr5r" colab_type="code" colab={}
class MLP_Base(nn.Module):
def __init__(self,Inp,Hidden,Projection):
super(MLP_Base,self).__init__()
self.Linear1 = nn.Linear(Inp,Hidden)
self.BatchNorm = nn.BatchNorm1d(Hidden)
self.Linear2 = nn.Linear(Hidden,Projection)
def forward(self,Input):
Linear_Inp = torch.relu(self.BatchNorm(self.Linear1(Input)))
Linear_Out = self.Linear2(Linear_Inp)
return Linear_Out
# + id="3R2korLYFr5x" colab_type="code" colab={}
class SkeletonNet(nn.Module):
def __init__(self,Hid,Proj):
super(SkeletonNet,self).__init__()
Resnet = torchvision.models.resnet18(pretrained=False)
self.Encoder = torch.nn.Sequential(*list(Resnet.children())[:-1])
self.Proj = MLP_Base(Resnet.fc.in_features,Hid,Proj)
def forward(self,Input):
Enc_Out = self.Encoder(Input)
Enc_Out = Enc_Out.view(Enc_Out.size(0),Enc_Out.size(1))
Final = self.Proj(Enc_Out)
return Final
# + [markdown] id="aCEF9jV8Fr53" colab_type="text"
# # Training Class
# + id="yANwkv3lFr53" colab_type="code" colab={}
class BYOL:
def __init__(self,Online_Net,Target_Net,Predictor,Optim,Params):
self.Online_Net = Online_Net
self.Target_Net = Target_Net
self.Predictor = Predictor
self.Optim = Optim
self.Device = Params['Device']
self.Epochs = Params['Epochs']
self.Moment = Params['M']
self.Batch_Size = Params['Batch_Size']
self.Save_Path = 'Models/BYOL.pth'
@torch.no_grad()
def Update_Target_Params(self):
for Param_Online,Param_Target in zip(self.Online_Net.parameters(),self.Target_Net.parameters()):
Param_Target = Param_Target.data *self.Moment + Param_Online.data*(1-self.Moment)
@staticmethod
def Loss(Rep1,Rep2):
Norm_Rep1 = F.normalize(Rep1,dim=-1,p=2) #L2-Normalized Rep One
Norm_Rep2 = F.normalize(Rep2,dim=-1,p=2) #L2 Normalized Rep Two
Loss = -2 * (Norm_Rep1*Norm_Rep2).sum(dim=-1)
return Loss
def Init_Target_Network(self):
for Param_Online,Param_Target in zip(self.Online_Net.parameters(),self.Target_Net.parameters()):
Param_Target.data.copy_(Param_Online.data) #Init Target with Param_Online
Param_Target.requires_grad = False
def TrainLoop(self,View1,View2):
self.Optim.zero_grad()
Pred1 = self.Predictor(self.Online_Net(View1))
Pred2 = self.Predictor(self.Online_Net(View2))
with torch.no_grad():
Target2 = self.Target_Net(View1)
Target1 = self.Target_Net(View2)
Loss_Calc = self.Loss(Pred1,Target1) + self.Loss(Pred2,Target2)
return Loss_Calc.mean()
def Train(self,Trainset):
TrainLoader = torch.utils.data.DataLoader(Trainset,batch_size=self.Batch_Size,drop_last=False,shuffle=True)
self.Init_Target_Network()
for Epoch in range(self.Epochs):
Loss_Count = 0.0
print("Epoch {}".format(Epoch))
for (View_1,View_2),_ in tqdm(TrainLoader):
View_1 = View_1.to(self.Device)
View_2 = View_2.to(self.Device)
Loss = self.TrainLoop(View_1,View_2)
Loss_Count += Loss.item()
Loss.backward()
self.Optim.step()
self.Update_Target_Params()
Epoch_Loss = Loss_Count/len(TrainLoader)
print("\n Epoch {} Loss:{} : ".format(Epoch,Epoch_Loss))
self.Save(self.Save_Path)
def Save(self,Save):
torch.save({'Online_Net':self.Online_Net.state_dict(),
'Enc_Net':self.Online_Net.Encoder.state_dict(),
'Target_Net':self.Target_Net.state_dict(),
'Optim':self.Optim.state_dict()},Save)
# + [markdown] id="SMEfYm3hFr58" colab_type="text"
# # Main Training
# + id="yV-8hgamFr58" colab_type="code" colab={}
Parameters = {'Epochs':50,'M':0.99,'Batch_Size':64,'Device':'cuda','Hidden':512,'Proj':128,'LR':0.03}
# + id="GHGyBW_QFr6A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e6dfb91-ef0b-4fdb-cd59-534eb044cb11"
Data_Transforms = Transforms((3,32,32))
Dataset = datasets.CIFAR10('./data',download=True,transform=MultiViewDataInjector([Data_Transforms,Data_Transforms]))
# + id="wd4ia_ilFr6E" colab_type="code" colab={}
Online_Network = SkeletonNet(Parameters['Hidden'],Parameters['Proj'])
Predictor = MLP_Base(Online_Network.Proj.Linear2.out_features,Parameters['Hidden'],Parameters['Proj'])
Target_Network = SkeletonNet(Parameters['Hidden'],Parameters['Proj'])
# + id="otHq9L2EFr6K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2ab41072-5d94-4958-a5da-1b5bb1c9b48d"
Online_Network.to(Parameters['Device'])
Predictor.to(Parameters['Device'])
Target_Network.to(Parameters['Device'])
print("Models Made.")
# + id="yrNO-ksrFr6P" colab_type="code" colab={}
Optimizer = torch.optim.SGD(list(Online_Network.parameters())+list(Predictor.parameters()),lr=0.03)
# + id="yTCxCJZjFr6T" colab_type="code" colab={}
Trainer = BYOL(Online_Network,Target_Network,Predictor,Optimizer,Parameters)
# + id="fFaoH14dFr6a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="a98b7a02-8214-4463-f13f-ba1d5823e7eb"
Trainer.Train(Dataset)
# + id="Ioq4CAvRHjgY" colab_type="code" colab={}
| BYOL_Pytorch (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import io
import sys
sys.path.append("../..")
import alkymi as alk
import pandas as pd
import matplotlib.pyplot as plt
import urllib.request
base_url = "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports_us"
url_arg = alk.recipes.args(name="url_arg")
@alk.recipe(ingredients=[url_arg])
def download_csv(url):
return urllib.request.urlopen(url).read()
@alk.recipe(ingredients=[download_csv])
def load_csv(data):
return pd.read_csv(io.BytesIO(data))
# +
# Decide which dataset to download
year = 2020
month = 12
day = 26
url_arg.set_args(f"{base_url}/{month:02d}-{day:02d}-{year}.csv")
# Run pipeline to fetch and load data
df = load_csv.brew()
# Plot the final results
num_to_show = 5
df.sort_values("Active", ascending=False) \
.head(num_to_show) \
.set_index("Province_State")["Active"] \
.plot.bar()
plt.title(f"Top {num_to_show} states by active case count")
plt.show()
| examples/notebook/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importing the Libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# Importing the dataset
dataset = pd.read_csv('../datasets/Social_Network_Ads.csv')
X = dataset.iloc[:, [2, 3]].values
y = dataset.iloc[:, 4].values
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Fitting Logistic Regression to the Training set
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
cm = confusion_matrix(y_test, y_pred)
print(cm) # print confusion_matrix
print(classification_report(y_test, y_pred)) # print classification report
#Visualization
from matplotlib.colors import ListedColormap
X_set,y_set=X_train,y_train
X1,X2=np.meshgrid(np.arange(start=X_set[:,0].min()-1, stop=X_set[:, 0].max()+1, step=0.01),
np.arange(start=X_set[:,1].min()-1, stop=X_set[:,1].max()+1, step=0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(),X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(),X1.max())
plt.ylim(X2.min(),X2.max())
for i,j in enumerate(np. unique(y_set)):
plt.scatter(X_set[y_set==j,0],X_set[y_set==j,1],
c = ListedColormap(('red', 'green'))(i), label=j)
plt.title(' LOGISTIC(Training set)')
plt.xlabel(' Age')
plt.ylabel(' Estimated Salary')
plt.legend()
plt.show()
X_set,y_set=X_test,y_test
X1,X2=np.meshgrid(np.arange(start=X_set[:,0].min()-1, stop=X_set[:, 0].max()+1, step=0.01),
np.arange(start=X_set[:,1].min()-1, stop=X_set[:,1].max()+1, step=0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(),X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(),X1.max())
plt.ylim(X2.min(),X2.max())
for i,j in enumerate(np. unique(y_set)):
plt.scatter(X_set[y_set==j,0],X_set[y_set==j,1],
c = ListedColormap(('red', 'green'))(i), label=j)
plt.title(' LOGISTIC(Test set)')
plt.xlabel(' Age')
plt.ylabel(' Estimated Salary')
plt.legend()
plt.show()
# -
| Code/Day 6_Logistic_Regression_practise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Local Thickness
#
# This example explains how to use the ``local_thicknes`` filter to get information about the pore size distribution from an image. The local thickness is probably the closest you can get to an actual pore size distribution. Unlike porosimetry experiments or simulations it is unaffected by artifacts such as edge effects. The implementation in PoreSpy is slightly different than the common approach done in ImageJ, as will be explained below.
# These notebooks were generated with a specific version of Python, PoreSpy, Numpy, etc. For reference to future viewers, the version information is given below. This notebook may or may not work with later versions, but we can assert it works with the version listed below:
# Import the needed packages from the Scipy stack:
import numpy as np
import porespy as ps
import scipy.ndimage as spim
import matplotlib.pyplot as plt
# ## Generate Test Image
# Start by generating an image. We'll use the RSA generator for fun:
# NBVAL_IGNORE_OUTPUT
im = np.zeros([300, 300])
im = ps.generators.RSA(im, radius=20, volume_fraction=0.2)
im = ps.generators.RSA(im, radius=15, volume_fraction=0.4)
im = ps.generators.RSA(im, radius=10, volume_fraction=0.6)
im = im == 0
plt.figure(figsize=[6, 6])
fig = plt.imshow(im)
# ## Apply Local Thickness Filter
# The local thickness filter is called by simply passing in the image. Like all filters in PoreSpy it is applied to the foreground, indicated by 1's or ``True``:
# NBVAL_IGNORE_OUTPUT
thk = ps.filters.local_thickness(im, mode='dt')
plt.figure(figsize=[6, 6])
fig = plt.imshow(thk, cmap=plt.cm.jet)
# ## Extracting PSD as a Histogram
# Obtaining pore size distribution information from this image requires obtaining a histogram of voxel values. A function in the ``metrics`` module does this for us:
psd = ps.metrics.pore_size_distribution(im=thk)
# The result returned into ``psd`` is a "named-tuple", which is a list of arrays that can be accessed by location (i.e. ``psd[0]``), but has the added benefit of accessing arrays by name so you know what you're getting. You can print a list of available arrays as follows:
print(psd._fields)
# Let's plot a pore-size distribution histogram:
# NBVAL_IGNORE_OUTPUT
# Set figure styles using predefined args in PoreSpy
ps.visualization.set_mpl_style()
plt.figure(figsize=[6, 6])
plt.xlabel('log(Pore Radius) [voxels]')
plt.ylabel('Normalized Volume Fraction')
fig = plt.bar(x=psd.logR, height=psd.pdf, width=psd.bin_widths, edgecolor='k')
# ## PoreSpy Implementation
# The ``local_thickness`` filter in PoreSpy is implemented differently that the *normal* approach such as the ImageJ plugin, though the end result is comparible though not identical.
#
# In our approach, we use a form of image dilation and erosion. We start with a large spherical structuring element, and note all the places where this fits in the pore space. This gives a result like that below for a structuring element of radius R=10:
# NBVAL_IGNORE_OUTPUT
R = 10
strel = ps.tools.ps_disk(R)
im_temp = spim.binary_opening(im, structure=strel)
plt.figure(figsize=[6, 6])
fig = plt.imshow(im_temp*2.0 + ~im)
# The key is to make a *master* array containing the numerical value of the largest sphere that covers each voxel. We'll initialize a new array with the current locations where R=10 fits:
im_result = im_temp*R
# Now this is repeated for a range of decreasing structuring element sizes. For illustration, do R = 8:
R = 8
strel = ps.tools.ps_disk(R)
im_temp = spim.binary_opening(im, structure=strel)
# This new image must be added to the ``im_result`` array, but only in places that were not filled at any larger radius. This is done using boolean logic as follows:
im_result[(im_result == 0)*im_temp] = R
# There are now 2 values in the ``im_results`` array indicating the locations where the structuring element of size 10 fits, and where size 8 fit on the subsequent step:
# NBVAL_IGNORE_OUTPUT
plt.figure(figsize=[6, 6])
fig = plt.imshow(im_result + ~im)
# The procedure is then repeated for smaller structuring elements down to R = 1. It's possible to specify which sizes are used, but by default all integers between $R_{max}$ and 1. This yields the image showed above:
# NBVAL_IGNORE_OUTPUT
plt.figure(figsize=[6, 6])
fig = plt.imshow(thk, cmap=plt.cm.jet)
| examples/filters/local_thickness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="C8KJ6MkAC2d1"
# NNFL ASSIGNMENT
# + colab={"base_uri": "https://localhost:8080/"} id="TNdhGk9VWXcn" outputId="eec5bdc5-bc44-4fb2-c9c5-ac007e3b0349"
# %tensorflow_version 1.x
# + colab={"base_uri": "https://localhost:8080/"} id="WzirspccVNgK" outputId="db666de8-9fb4-4edc-9e07-e0cf4c9738ea"
pip install -r lib/Mask_RCNN/requirements.txt
# + colab={"base_uri": "https://localhost:8080/"} id="I7l3RG7cELxM" outputId="d996b997-6a8d-4b66-b650-6621aa46855f"
# !pip uninstall tensorflow
# !pip install tensorflow==1.14
# !pip uninstall keras
# !pip install keras==2.2.4
# + colab={"base_uri": "https://localhost:8080/"} id="sOUaWPPVEUEY" outputId="15ff5882-0ca5-465f-badf-6edf2b93a108"
# !pip install Keras-Applications
# + colab={"base_uri": "https://localhost:8080/"} id="mDxI6obwSKfF" outputId="c6a9be56-ab1e-4673-9daf-f265d3a675bf"
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# #%load_ext line_profiler
import tensorflow as tf
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.INFO)
sess_config = tf.compat.v1.ConfigProto()
import sys
import os
COCO_DATA = 'data/coco/'
MASK_RCNN_MODEL_PATH = 'lib/Mask_RCNN/'
if MASK_RCNN_MODEL_PATH not in sys.path:
sys.path.append(MASK_RCNN_MODEL_PATH)
from lib.Mask_RCNN.samples.coco import coco
from mrcnn import utils
from mrcnn import model as modellib
from mrcnn import visualize
from lib import utils as siamese_utils
from lib import model as siamese_model
from lib import config as siamese_config
import time
import datetime
import random
import numpy as np
import skimage.io
import imgaug
import pickle
import matplotlib.pyplot as plt
from collections import OrderedDict
# Root directory of the project
ROOT_DIR = os.getcwd()
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# + colab={"base_uri": "https://localhost:8080/"} id="qyX-KprST172" outputId="07d60c05-19c3-4dcc-8e0f-6d58e101468c"
# train_classes = coco_nopascal_classes
train_classes = np.array(range(1,81))
# Load COCO/val dataset
coco_val = siamese_utils.IndexedCocoDataset()
coco_object = coco_val.load_coco(COCO_DATA, "val", year="2017", return_coco=True)
coco_val.prepare()
coco_val.build_indices()
coco_val.ACTIVE_CLASSES = train_classes
# + id="cEMybx3zT5VN"
class SmallEvalConfig(siamese_config.Config):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
NUM_CLASSES = 1 + 1
NAME = 'coco'
EXPERIMENT = 'evaluation'
CHECKPOINT_DIR = 'checkpoints/'
NUM_TARGETS = 1
class LargeEvalConfig(siamese_config.Config):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
NUM_CLASSES = 1 + 1
NAME = 'coco'
EXPERIMENT = 'evaluation'
CHECKPOINT_DIR = 'checkpoints/'
NUM_TARGETS = 1
# Large image sizes
TARGET_MAX_DIM = 192
TARGET_MIN_DIM = 150
IMAGE_MIN_DIM = 800
IMAGE_MAX_DIM = 1024
# Large model size
FPN_CLASSIF_FC_LAYERS_SIZE = 1024
FPN_FEATUREMAPS = 256
# Large number of rois at all stages
RPN_ANCHOR_STRIDE = 1
RPN_TRAIN_ANCHORS_PER_IMAGE = 256
POST_NMS_ROIS_TRAINING = 2000
POST_NMS_ROIS_INFERENCE = 1000
TRAIN_ROIS_PER_IMAGE = 200
DETECTION_MAX_INSTANCES = 100
MAX_GT_INSTANCES = 100
# + colab={"base_uri": "https://localhost:8080/"} id="8EB8IQJMT8fl" outputId="1e25135f-9e29-4a14-c816-4e0a4364a65f"
# The small model trains on a single GPU and runs much faster.
# The large model is the same we used in our experiments but needs multiple GPUs and more time for training.
model_size = 'small' # or 'large'
if model_size == 'small':
config = SmallEvalConfig()
elif model_size == 'large':
config = LargeEvalConfig()
config.display()
# Provide training schedule of the model
# When evaluationg intermediate steps the tranining schedule must be provided
train_schedule = OrderedDict()
if model_size == 'small':
train_schedule[1] = {"learning_rate": config.LEARNING_RATE, "layers": "heads"}
train_schedule[120] = {"learning_rate": config.LEARNING_RATE, "layers": "4+"}
train_schedule[160] = {"learning_rate": config.LEARNING_RATE/10, "layers": "all"}
elif model_size == 'large':
train_schedule[1] = {"learning_rate": config.LEARNING_RATE, "layers": "heads"}
train_schedule[240] = {"learning_rate": config.LEARNING_RATE, "layers": "all"}
train_schedule[320] = {"learning_rate": config.LEARNING_RATE/10, "layers": "all"}
# Select checkpoint
if model_size == 'small':
checkpoint = 'checkpoints/small_siamese_mrcnn_0160.h5'
elif model_size == 'large':
checkpoint = 'checkpoints/large_siamese_mrcnn_coco_full_0320.h5'
# + colab={"base_uri": "https://localhost:8080/", "height": 559} id="b73yjE6zUCaO" outputId="776353ec-bbf0-421c-b168-21ee536445ab"
# Load and evaluate model
# Create model object in inference mode.
model = siamese_model.SiameseMaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
model.load_checkpoint(checkpoint, training_schedule=train_schedule)
# Evaluate only active classes
active_class_idx = np.array(coco_val.ACTIVE_CLASSES) - 1
# + colab={"base_uri": "https://localhost:8080/"} id="HiFUSEjbl-_1" outputId="505ec33f-7426-499d-ab4a-9807992fdf1a"
count = 0
for layer in model.keras_model.layers:
count += 1
if str(layer).find('Model') != -1:
print(count, str(layer))
break
index = count - 1
model.keras_model.layers[index].summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="aMddS9QTtQse" outputId="e6069499-181a-49b7-bf3c-3a076eeeb2f7"
filters, biases = model.keras_model.layers[index].layers[9].get_weights()
filters_min, filters_max = filters.min(), filters.max()
filters = (filters - filters_min)/(filters_max - filters_min)
_, _, n_channels, n_filters = filters.shape
n_channels = 16 #Can't visualise all filters and channels, hence 16 each
n_filters = 16
ind = 1
plt.rcParams["figure.figsize"] = (30,30)
for i in range(n_filters):
f = filters[:, :, :, i]
for j in range(n_channels):
ax = plt.subplot(n_filters, n_channels, ind)
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(f[:, :, j], cmap= 'viridis')
ind += 1
plt.show()
# + id="0wNA7v06gIyA"
plt.rcParams["figure.figsize"] = (10,10)
from keras.models import Model
indexes = [17,27,37,49,59,69,79,91,101,111,121,131,141,153,163,173]
outputs = [model.keras_model.layers[index].layers[i].output for i in indexes]
model_feature_maps = Model(inputs=model.keras_model.layers[index].inputs, outputs=outputs)
# + colab={"base_uri": "https://localhost:8080/", "height": 487} id="NMWPMD-T3QST" outputId="ae809981-0570-4dc1-8aba-c0026e47db4e"
# Select category
category = 1
np.random.seed(7)
image_id = np.random.choice(coco_val.category_image_index[category])
# Load target
target = siamese_utils.get_one_target(category, coco_val, config)
# Load image
image = coco_val.load_image(image_id)
img = np.expand_dims(image, axis=0)
feature_maps = model_feature_maps.predict(img)
plt.imshow(image)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="K-qgqbPfxJJr" outputId="c8a80a11-406f-4314-bfc4-b4e3d9a549eb"
feature_maps = model_feature_maps.predict(img)
# plot the output from each block
square = 2
s = f"""\n"""
for fmap in feature_maps:
# plot 64 maps for first 3 layers
ix = 1
for _ in range(square):
for _ in range(square):
# specify subplot
ax = plt.subplot(square, square, ix)
# plot filter channel
plt.imshow(fmap[0, :, :, ix-1], cmap='viridis')
ix += 1
plt.text(0,64,s)
plt.show()
| visualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
engine = create_engine("sqlite:///hawaii.sqlite")
# +
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# -
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Climate Analysis
# +
# Design a query to retrieve the last 12 months of precipitation data and plot the results.
#Starting from the last data point in the database.
prev_year = dt.date(2017, 8, 23)
# Calculate the date one year from the last date in data set.
prev_year = dt.date(2017, 8, 23) - dt.timedelta(days=365)
# Perform a query to retrieve the data and precipitation scores
results = []
results = session.query(Measurement.date, Measurement.prcp).filter(Measurement.date >= prev_year).all()
# Save the query results as a Pandas DataFrame and set the index to the date column
df = pd.DataFrame(results, columns=['date','precipitation'])
# Sort the dataframe by date
df.set_index(df['date'], inplace=True)
df = df.sort_index()
# Use Pandas Plotting with Matplotlib to plot the data
df.plot()
# -
# Use Pandas to calcualte the summary statistics for the precipitation data
df.describe()
# How many stations are available in this dataset?
session.query(func.count(Station.station)).all()
# What are the most active stations?
# List the stations and the counts in descending order.
session.query(Measurement.station, func.count(Measurement.station)).\
group_by(Measurement.station).order_by(func.count(Measurement.station).desc()).all()
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature most active station?
session.query(func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs)).\
filter(Measurement.station == 'USC00519281').all()
# +
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
results = session.query(Measurement.tobs).\
filter(Measurement.station == 'USC00519281').\
filter(Measurement.date>= prev_year).all()
df = pd.DataFrame(results, columns=['tobs'])
df.plot.hist(bins=12)
plt.tight_layout()
# -
# Write a function called `calc_temps` that will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
# # Challenge
# +
# Set variables for December start and end
december_start = dt.date(2016, 12, 1)
december_end = dt.date(2016, 12, 31)
# Query the Measurements data for the month of December
results = []
results = session.query(Measurement.date, Measurement.prcp, Measurement.tobs, Measurement.station).filter(Measurement.date >= december_start).\
filter(Measurement.date <= december_end).all()
# Add to DataFrame
df = pd.DataFrame(results, columns=['date', 'precipitation', 'tobs', 'station'])
# Set index by the date
df.set_index(df['date'], inplace=True)
df = df.sort_index()
# Find the December statistics using .decribe
df.describe()
# +
# Set variables for June start and end
june_start = dt.date(2016, 6, 1)
june_end = dt.date(2016, 6, 30)
# Query the Measurements data for the month of June
results = []
results = session.query(Measurement.date, Measurement.prcp, Measurement.tobs, Measurement.station).filter(Measurement.date >= june_start).\
filter(Measurement.date <= june_end).all()
# Add to DataFrame
df = pd.DataFrame(results, columns=['date', 'precipitation', 'tobs', 'station'])
# Set index by the date
df.set_index(df['date'], inplace=True)
df = df.sort_index()
# Find the June statistics using .decribe
df.describe()
# -
# # Findings
#
# +
# The June and December statistics are very close to each other.
# December has a higher max of precipitation, but the average for both months is very close. This low average should show that the shop will not have a drastic difference in business between the two months.
# The average temperature for June is only 4 degrees higher than December, showing that ice cream sales should be consistent between the two months.
# Further analysis should be done on the other months as December may not be the height of the rainy season in Oahu and there could be average weather in another month that will negatively affect business more.
| climate_analysis_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory Data Analysis - Sports (Indian Premier League)
# As a sports analysts, find out the most successful teams, players and factors
# contributing win or loss of a team. Suggest teams or players a company should endorse for its products.
# #### Author : <NAME>
# ### IMPORTING REQUIRED LIBRARIES
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
df_m =pd.read_csv("matches.csv")
df_m.head()
# +
df_d= pd.read_csv("deliveries.csv")
df_d.head()
# -
wins_per_season = df_m.groupby("season")["winner"].value_counts()
wins_per_season
# ### Data Preparation and Cleaning
df_m.info()
df_d.info()
df_m.isnull().sum()
# #### The variable 'umpire3' is dropped because it contains too many missing data.
df_m.drop('umpire3',axis = 1, inplace=True)
df_d.isnull().sum()
# ### player_dismissed,dismissal_kind,fielder have lot of missing values
# #### It is normal for these features to contain missing data as dismissal of player does not happen in every ball or over.
df_m["team1"].unique()
df_m["team2"].unique()
df_d["batting_team"].unique()
df_d["bowling_team"].unique()
# #### 'Rising Pune Supergiant' & 'Rising Pune Supergiants' are same
df_m.replace('Rising Pune Supergiant','Rising Pune Supergiants', inplace=True)
df_d.replace('Rising Pune Supergiant','Rising Pune Supergiants', inplace=True)
df_m["city"].unique()
#
# #### 'Bangalore' & 'Bengaluru' are same
df_m.replace('Bangalore','Bengaluru', inplace=True)
# ## Exploratory Analysis and Visualization
# #### Number of Matches played in each IPL season
plt.figure(figsize = (18,10))
sns.countplot('season',data=df_m,palette="winter")
plt.title("Number of Matches played in each IPL season",fontsize=20)
plt.xlabel("season",fontsize=15)
plt.ylabel('Matches',fontsize=15)
plt.show()
# #### Numbers of matches won by team
plt.figure(figsize = (18,10))
sns.countplot(x='winner',data=df_m, palette='cool')
plt.title("Numbers of matches won by team ",fontsize=20)
plt.xticks(rotation=50)
plt.xlabel("Teams",fontsize=15)
plt.ylabel("No of wins",fontsize=15)
plt.show()
# #### Number of Season wins per team:
# "Mumbai Indians is the most successful team with highest wins in 4 seasons
#followed by Chennai Super Kings with 3 seasons"
plt.subplots(figsize=(11,6))
season_winner = df_m.drop_duplicates('season', keep='last')
ax=sns.countplot(x='winner', data=season_winner, palette = 'inferno')
plt.xticks(rotation=90)
for p in ax.patches:
ax.annotate(str(p.get_height()), (p.get_x(), p.get_height()))
plt.show()
#Top 10 Players based on MOM:CH Gayle is the player with the most man of the match award with 21
#followed by <NAME> with 20
plt.subplots(figsize=(16,8))
ax = sns.barplot(x = df_m['player_of_match'].value_counts()[:10].index, y = df_m['player_of_match'].value_counts()[:10],palette = 'bright')
plt.ylabel('Number of MOM won',fontsize=15)
plt.title('Top 10 players based on MOM won',fontsize=20)
plt.xlabel('Players',fontsize=12)
plt.xticks(rotation=90,fontsize=14)
for p in ax.patches:
ax.annotate(np.round(p.get_height(), decimals=0).astype(np.int64), (p.get_x(), p.get_height()))
plt.show()
# ### factors contributing win or loss of a team.
len(df_m)
defending = df_m['win_by_runs'].value_counts()
defending.sort_values(ascending=False)
# +
#There were more matches won by chasing the total(419 matches) i.e. 55% than defending(350 matches) i.e.45%.
df_m['win_by']=np.where(df_m['win_by_runs']>0,'Bat first','Bowl first')
match=df_m.win_by.value_counts()
labels=np.array(match.index)
sizes = match.values
colors = ['green', 'gold']
plt.pie(sizes, labels=labels, colors=colors,autopct='%1.1f%%', shadow=True,startangle=90)
plt.title('Match Result')
plt.axis('equal')
plt.show()
# -
# ## Toss Decision:
# +
# 61% of toss decision is fielding and 39% is batting.
toss=df_m.toss_decision.value_counts()
labels=np.array(toss.index)
sizes = toss.values
colors = ['red', 'orange']
plt.pie(sizes, labels=labels, colors=colors, autopct='%1.1f%%', shadow=True,startangle=90)
plt.title('Toss Result')
plt.axis('equal')
plt.show()
# -
# ## number of matches win is higher when toss_decision is 'field'.
plt.subplots(figsize=(16,9))
toss=df_m[df_m['toss_winner']==df_m['winner']]
ax = sns.countplot("winner", data = toss, hue = 'toss_decision',order = toss['toss_winner'].value_counts().index,palette='inferno')
plt.title("Number of winning teams according to the toss decision",fontsize=15)
plt.xticks(rotation='vertical', ha = 'right',fontsize=15)
plt.ylabel('Number of Matches Win',fontsize=15)
plt.xlabel('Winner',fontsize=15)
for p in ax.patches: ax.annotate(np.round(p.get_height(), decimals=0).astype(np.int64), (p.get_x(), p.get_height()))
plt.show()
# ### To analyse if toss winners are match winners in most seasons.
# +
#In the final matches in each seasons, most of the toss winners are match winners
final_matches=df_m.drop_duplicates(subset=['season'], keep='last')
final_matches[['season','winner', 'toss_winner']].reset_index(drop=True).sort_values('season')
# -
#
# ### Suggest teams or players a company should endorse for its products
# #### Batsman analysis:
# +
#Top 10 Batsman with most number of Fours
#<NAME> has scored highest fours followed by SK Raina
batsman_df = df_d.groupby('batsman')['batsman_runs'].agg(lambda x: (x==4).sum()).reset_index().sort_values(by='batsman_runs', ascending=False).reset_index(drop=True)
batsman_df = batsman_df.iloc[:10,:]
labels = np.array(batsman_df['batsman'])
ind = np.arange(len(labels))
width = 0.9
fig, ax = plt.subplots(figsize=(15,8))
rects = ax.bar(ind, np.array(batsman_df['batsman_runs']), width=width, color='green')
ax.set_xticks(ind+((width)/2.))
ax.set_xticklabels(labels, rotation='vertical')
ax.set_ylabel("Count")
ax.set_title("Top 10 Batsman with most number of Fours",fontsize = 10)
for p in ax.patches:
ax.annotate(np.round(p.get_height(), decimals=0).astype(np.int64), (p.get_x(), p.get_height()))
plt.show()
# +
#Top 10 Batsman with most number of Fours
##CH Gayle has scored highest sixes followed by <NAME>
batsman_df = df_d.groupby('batsman')['batsman_runs'].agg(lambda x: (x==6).sum()).reset_index().sort_values(by='batsman_runs', ascending=False).reset_index(drop=True)
batsman_df = batsman_df.iloc[:10,:]
labels = np.array(batsman_df['batsman'])
ind = np.arange(len(labels))
width = 0.9
fig, ax = plt.subplots(figsize=(15,8))
rects = ax.bar(ind, np.array(batsman_df['batsman_runs']), width=width, color='orange')
ax.set_xticks(ind+((width)/2.))
ax.set_xticklabels(labels, rotation='vertical')
ax.set_ylabel("Count")
ax.set_title("Top 10 Batsman with most number of Sixes",fontsize = 10)
for p in ax.patches:
ax.annotate(np.round(p.get_height(), decimals=0).astype(np.int64), (p.get_x(), p.get_height()))
plt.show()
# +
#Batsman with highest runs in IPL
# <NAME> is the highest run scorer followed by <NAME>
batsman_df = df_d.groupby('batsman')['batsman_runs'].agg('sum').reset_index().sort_values(by='batsman_runs', ascending=False).reset_index(drop=True)
batsman_df = batsman_df.iloc[:10,:]
labels = np.array(batsman_df['batsman'])
ind = np.arange(len(labels))
width = 0.9
fig, ax = plt.subplots(figsize=(15,8))
rects = ax.bar(ind, np.array(batsman_df['batsman_runs']), width=width, color='blue')
ax.set_xticks(ind+((width)/2.))
ax.set_xticklabels(labels, rotation='vertical')
ax.set_ylabel("Count")
ax.set_title("Batsman with highest runs in IPL")
ax.set_xlabel('Batsmane Name')
for p in ax.patches:
ax.annotate(np.round(p.get_height(), decimals=0).astype(np.int64), (p.get_x(), p.get_height()))
plt.show()
# -
# ### Bowler Analysis
# +
#Top 10 bowlers in IPL who bowled maximum balls
#<NAME> is the bowler with highest balls followed by <NAME>
df = pd.merge(df_m, df_d, left_on='id', right_on='match_id')
bowler = df['bowler'].value_counts()[:10]
plt.figure(figsize=(15,7))
plt.bar(x=bowler.index, height=bowler.values,color='purple')
plt.title('Bowlers who bowled maximum balls', fontsize=20)
plt.xlabel('BOWLER',fontsize=15)
plt.ylabel('BALLS',fontsize=15)
for i,v in enumerate(bowler.values):
plt.text(x=i, y=v+1, s=v)
plt.show()
# +
#Top 10 bowlers in IPL with highest wicket.
# SL Malinga is the bowler with highest wicket of 170 followed by A Mishra with 156 wickets
bowling_wickets= df_d[df_d['dismissal_kind']!='run out']
bowling_tot=bowling_wickets.groupby('bowler').apply(lambda x:x['dismissal_kind'].dropna()).reset_index(name='Wickets')
bowling_wick_count=bowling_tot.groupby('bowler').count().reset_index()
bowling_top=bowling_wick_count.sort_values(by='Wickets',ascending=False)
top_bowlers=bowling_top.loc[:,['bowler','Wickets']][0:10]
print('The Top Wicket Takers in the Tournament are:\n',top_bowlers)
plt.bar(top_bowlers['bowler'],top_bowlers['Wickets'],color='r')
plt.plot(top_bowlers['bowler'],top_bowlers['Wickets'],color='g')
plt.xticks(rotation=90)
plt.xlabel('Top 10 Bowlers',size=15)
plt.ylabel('Wickets Taken',size=15)
plt.title('Top 10 Bowlers with highest wickets in IPL',size=20)
plt.show()
# -
# Dismissals in IPL
plt.figure(figsize=(18,10))
ax=sns.countplot(df_d.dismissal_kind)
plt.title("Dismissals in IPL",fontsize=20)
plt.xlabel("Dismissals kind",fontsize=15)
plt.ylabel("count",fontsize=15)
plt.xticks(rotation=90)
plt.show()
# ### Conclusion :
#
# - The highest number of match played in IPL season was 2013,2014,2015.
#
# - The highest number of match won by Mumbai Indians i.e 4 match out of 12 matches.
#
# - Mumbai Indians is the most successful team with highest wins in 4 seasons followed by Chennai Super Kings in 3 seasons
#
# - MOST SUCCESSFUL PLAYERS:
# CH Gayle is the player with the most man of the match award with 21 followed by AB de Villiers with 20.
#
# - FACTORS CONTRIBUTING WIN OR LOSS OF A TEAM.
# There were more matches won by chasing the total (419 matches, 55%) than defending (350 matches, 45%).
#
# - The number of wins is higher when toss_decision is 'field'.
# Toss winners are match winners in most IPL season's final matches.
#
# - SUGGEST TEAMS OR PLAYERS A COMPANY SHOULD ENDORSE FOR ITS PRODUCTS.
# - The teams that can be recommended for companies to endorse are: Mumbai Indians, Chennai Super Kings and Kolkata Knight Riders.
# - The recommended bowler for companies to endorse could be: SL Malinga, Harbajan Singh and A.Mishra.
# - The recommended batman for companies to endorse could be: CH Gayle, AB de Villiers, V Kohli, S Dhawan and SK Raina.
#
# - Dismissals in IPL was most by Catch out .
# # Thank You!!
#
# ### <NAME>
| Exploratory Data Analysis - Sports.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%%\n"}
# AutoML searches for the optimal model and its parameter combination through several trials. At the end of the experiment, in addition to returning the optimal model, **model ensemble** can also be performed on ``topk`` to improve the generalization of the pipeline.
#
# In HyperTS, we introduce a mechanism called GreedyEnsemble for model ensemble. Its specific process is as follows[1]:
# -
# 1. Start with the empty ensemble.
#
# 2. Add to the ensemble the model in the library that maximizes the ensemble’s performance to the error metric on a hillclimb (validation) set.
#
# 3. Repeat Step 2 for a fixed number of iterations or until all the models have been used.
#
# 4. Return the ensemble from the nested set of ensembles that has maximum performance on the hillclimb (validation) set.
# References
#
# [1] Caruana, Rich, et al. "Ensemble selection from libraries of models." in ICML. 2004.
# **Example of use:**
# #### 1. Prepare Dataset
from hyperts.datasets import load_network_traffic
from sklearn.model_selection import train_test_split
df = load_network_traffic(univariate=True)
train_data, test_data = train_test_split(df, test_size=168, shuffle=False)
# #### 2. Create Experiment and Run
from hyperts import make_experiment
# Set parameter ``ensemble_size`` to control the number of ensemble models.
experiment = make_experiment(train_data=train_data.copy(),
task='forecast',
mode='dl',
timestamp='TimeStamp',
covariates=['HourSin', 'WeekCos', 'CBWD'],
forecast_train_data_periods=24*12,
max_trials=10,
ensemble_size=5)
model = experiment.run()
model.get_params
# #### 3. Infer and Evaluation
X_test, y_test = model.split_X_y(test_data.copy())
forecast = model.predict(X_test)
results = model.evaluate(y_true=y_test, y_pred=forecast)
results
| examples/09_ensemble_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sV0U5S_GwKAa"
# # Mini-lab 6: EM for k-means
# + id="zLNlSVQYkc_F" colab={"base_uri": "https://localhost:8080/"} outputId="193728c0-086a-4382-e563-ccc63329a9e0"
import numpy as np # vectors etc
import matplotlib.pyplot as plt # for plotting
np.random.seed(0) # Ensures we get the same 'random' data every time
X = np.random.multivariate_normal([0, 0], [[4, 0],[0, 4]], size=200)
y = [0]*X.shape[0]
X = np.concatenate([X, np.random.multivariate_normal([5, 3], [[1, 0],[0, 1]], size=100)])
y.extend([1]*(X.shape[0]-len(y)))
X = np.concatenate([X, np.random.multivariate_normal([-3, 8], [[1, 0],[0, 1]], size=50)])
y.extend([2]*(X.shape[0]-len(y)))
y = np.asarray(y)
X.shape, y.shape
# + [markdown] id="-WKXo_4-wN0S"
#
# + id="ZAuFFpFhsF6C"
n_clusters = 3
model_data = {'centres': None,
'data': X,
'class_attributions': None}
# + colab={"base_uri": "https://localhost:8080/", "height": 499} id="WHVxlnpowv5G" outputId="731a0d69-7d7d-44fe-f2b5-18f3428df23b"
def plot_model():
global model_data
plt.figure(figsize=(5, 5), dpi=120)
X = model_data['data']
if model_data['class_attributions'] is None:
y = np.zeros(X.shape[0])
else:
y = model_data['class_attributions']
plt.scatter(X[:, 0], X[:, 1], c=y, s=30, edgecolor='k', cmap='tab10')
if model_data['centres'] is not None:
M = model_data['centres']
plt.scatter(M[:, 0], M[:, 1], c=np.arange(M.shape[0]), s=60, marker='x', linewidth=3, cmap='tab10', zorder=1)
plt.xticks([])
plt.yticks([])
plt.show()
plot_model()
# + [markdown] id="KCWyn_IewPgy"
# ## E step
# + id="2ALenkKuvImg"
plot_model()
# + [markdown] id="NwAbZSa_wRZL"
# ## M step
# + id="aOGUGVcywTvH"
plot_model()
| Mini-lab 06 - EM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Predicting Solar Flares with Machine Learning**
# + [markdown] toc=true
# <h1><b>Table of Contents</b><span class="tocSkip"></span> </h1>
# <div class="toc">
# <!-- <ul class="toc-item"> -->
# <li><span><a href="#Template-Notebook-for-EarthCube---Long-Version"
# data-toc-modified-id="Template-Notebook-for-EarthCube---Long-Version-1"><span
# class="toc-item-num">1 </span>YA_Notebook for Predicting Solar Flares with Machine
# Learning</a></span>
# <ul class="toc-item" style='margin-top:0px; margin-bottom:0px'>
# <li><span><a href="#Author(s)" data-toc-modified-id="Author(s)-1.1"><span
# class="toc-item-num">1.1 </span>Author(s)</a></span></li>
# <li><span><a href="#Purpose" data-toc-modified-id="Purpose-1.2"><span
# class="toc-item-num">1.2 </span>Purpose</a></span></li>
# <li><span><a href="#Technical-Contributions" data-toc-modified-id="Technical-Contributions-1.3"><span
# class="toc-item-num">1.3 </span>Technical Contributions</a></span></li>
# <li><span><a href="#Methodology" data-toc-modified-id="Methodology-1.4"><span
# class="toc-item-num">1.4 </span>Methodology</a></span></li>
# <li><span><a href="#Funding" data-toc-modified-id="Funding-1.6"><span
# class="toc-item-num">1.5 </span>Funding</a></span></li>
# <li><span><a href="#Keywords" data-toc-modified-id="Keywords-1.7"><span
# class="toc-item-num">1.6 </span>Keywords</a></span></li>
# <li><span><a href="#Citation" data-toc-modified-id="Citation-1.8"><span
# class="toc-item-num">1.7 </span>Citation</a></span></li>
# <li><span style='display:inline'><a href="#Acknowledgements" data-toc-modified-id="Acknowledgements-1.11"><span
# class="toc-item-num">1.8 </span>Acknowledgements</a></span></li>
# </ul>
# </li>
# <!-- </ul> -->
# <li><span><a href="#Setup" data-toc-modified-id="Setup-2"><span
# class="toc-item-num">2 </span>Setup</a></span>
# </li>
# <li><span><a href="#Data-Processing-and-Analysis" data-toc-modified-id="Data-Processing-and-Analysis-3"><span
# class="toc-item-num">3 </span>Data Processing and Analysis</a></span></li>
# <li><span><a href="#Binder" data-toc-modified-id="Binder-4"><span
# class="toc-item-num">4 </span>Binder</a></span></li>
# <li><span><a href="#FlareML-Workflow" data-toc-modified-id="FlareML-Workflow-5"><span
# class="toc-item-num">5 </span>FlareML Workflow and Results</a></span>
# <ul class="toc-item" style='margin-top:0px;margin-bottom:0px'>
# <li>
# <span><a href="#Data-Preparation-and-Loading"
# data-toc-modified-id="Data-Preparation-and-Loading-5.1"><span
# class="toc-item-num">5.1 </span>Data Preparation and Loading</a></span>
# </li>
# <li>
# <span><a href="#Predicting-with-Pretrained-Models"
# data-toc-modified-id="Predicting-with-Pretrained Models-5.6"><span
# class="toc-item-num">5.2 </span>Predicting with Pretrained Models</a></span>
# <!-- <ul class="toc-item">
# <li style='list-style: square'>
# <span><a href="#Plotting-the-Pretrained-Models-Results"
# data-toc-modified-id="Plotting-the-Pretrained-Models-Results-5.2.3">
# <span class="toc-item-num">5.2.1
# </span>Plotting the Pretrained Models Results</a></span>
# </li>
# </ul> -->
# </li>
# <li>
# <span><a href="#ENS-Model-Training-and-Testing"
# data-toc-modified-id="ENS-Model-Training-and-Testing-5.2"><span
# class="toc-item-num">5.3 </span>ENS Model Training and Testing</a></span>
# <!-- <ul class="toc-item">
# <li style='list-style: square'>
# <span><a href="#ENS-Model-Training-with-Default-Data"
# data-toc-modified-id="ENS Model-Training-with-Default-Data-6">
# <span class="toc-item-num">5.3.1
# </span>ENS Model Training with Default Data</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Predicting-with-Your-ENS-Model"
# data-toc-modified-id="Predicting-with-Your-ENS-Model-5.2.2">
# <span class="toc-item-num">5.3.2
# </span>Predicting with Your ENS Model</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Plotting-the-ENS-Results" data-toc-modified-id="Plotting-the-ENS-Results-5.2.3">
# <span class="toc-item-num">5.3.3
# </span>Plotting the ENS Results</a></span>
# </li>
# </ul> -->
# </li>
# <li>
# <span><a href="#RF-Model-Training-and-Testing"
# data-toc-modified-id="RF-Model-Training-and-Testing-5.2"><span
# class="toc-item-num">5.4 </span>RF Model Training and Testing</a></span>
# <!-- <ul class="toc-item">
# <li style='list-style: square'>
# <span><a href="#RF-Model-Training-with-Default-Data"
# data-toc-modified-id="RF Model-Training-with-Default-Data-6">
# <span class="toc-item-num">5.4.1
# </span>RF Model Training with Default Data</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Predicting-with-Your-RF-Model"
# data-toc-modified-id="Predicting-with-Your-RF-Model-5.2.2">
# <span class="toc-item-num">5.4.2
# </span>Predicting with Your RF Model</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Plotting-the-RF-Results" data-toc-modified-id="Plotting-the-RF-Results-5.2.3">
# <span class="toc-item-num">5.4.3
# </span>Plotting the RF Results</a></span>
# </li>
# </ul> -->
# </li>
# <li>
# <span><a href="#MLP-Model-Training-and-Testing"
# data-toc-modified-id="MLP-Model-Training-and-Testing-5.2"><span
# class="toc-item-num">5.5 </span>MLP Model Training and Testing</a></span>
# <!-- <ul class="toc-item">
# <li style='list-style: square'>
# <span><a href="#MLP-Model-Training-with-Default-Data"
# data-toc-modified-id="MLP Model-Training-with-Default-Data-6">
# <span class="toc-item-num">5.5.1
# </span>MLP Model Training with Default Data</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Predicting-with-Your-MLP-Model"
# data-toc-modified-id="Predicting-with-Your-RF-Model-5.2.2">
# <span class="toc-item-num">5.5.2
# </span>Predicting with Your MLP Model</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Plotting-the-MLP-Results" data-toc-modified-id="Plotting-the-MLP-Results-5.2.3">
# <span class="toc-item-num">5.5.3
# </span>Plotting the MLP Results</a></span>
# </li>
# </ul> -->
# </li>
# <li>
# <span><a href="#ELM-Model-Training-and-Testing"
# data-toc-modified-id="ELM-Model-Training-and-Testing-5.2"><span
# class="toc-item-num">5.6 </span>ELM Model Training and Testing</a></span>
# <!-- <ul class="toc-item">
# <li style='list-style: square'>
# <span><a href="#ELM-Model-Training-with-Default-Data"
# data-toc-modified-id="ELM Model-Training-with-Default-Data-6">
# <span class="toc-item-num">5.6.1
# </span>ELM Model Training with Default Data</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Predicting-with-Your-ELM-Model"
# data-toc-modified-id="Predicting-with-Your-ELM-Model-5.2.2">
# <span class="toc-item-num">5.6.2
# </span>Predicting with Your ELM Model</a></span>
# </li>
# <li style='list-style: square'>
# <span><a href="#Plotting-the-ELM-Results" data-toc-modified-id="Plotting-the-ELM-Results-5.2.3">
# <span class="toc-item-num">5.6.3
# </span>Plotting the ELM Results</a></span>
# </li>
# </ul> -->
# </li>
# <li>
# <span><a href="#Timing"
# data-toc-modified-id="Timing-5.7"><span
# class="toc-item-num">5.7 </span>Timing</a></span>
# </li>
# </ul>
# </li>
# <!-- </ul> -->
# </div>
# </ul>
# </li>
# <li><span><a href="#Conclusions" data-toc-modified-id="Conclusions-6"><span
# class="toc-item-num">6 </span>Conclusions</a></span>
# </li>
# <li><span><a href="#References" data-toc-modified-id="References-7"><span
# class="toc-item-num">7 </span>References</a></span>
# </li>
# </ul>
# </div>
# -
# ### **Author(s)**
# - Author1 = {"name": "<NAME>", "affiliation": "Department of Computer Science, New Jersey Institute of Technology", "email": "<EMAIL>", "orcid": "https://orcid.org/0000-0003-0792-2270"}
# - Author2 = {"name": "<NAME>", "affiliation": "Department of Computer Science, New Jersey Institute of Technology", "email": "<EMAIL>", "orcid": "https://orcid.org/0000-0002-2486-1097"}
# - Author3 = {"name": "<NAME>", "affiliation": "Institute for Space Weather Sciences, New Jersey Institute of Technology", "email": "<EMAIL>", "orcid": "https://orcid.org/0000-0002-5233-565X"}
# ### **Purpose**
# Solar flare prediction plays an important role in understanding and forecasting space weather. The main goal of the Helioseismic and Magnetic Imager (HMI), one of the instruments on NASA's Solar Dynamics Observatory (SDO), is to study the origin of solar variability and characterize the Sun's magnetic activity. HMI provides continuous full-disk observations of the solar vector magnetic field with high cadence data that lead to reliable predictive capability; yet, solar flare prediction effort utilizing these data is still limited.
#
# In this notebook we provide an overview of the FlareML system to demonstrate how to predict solar flares using machine learning (ML) and SDO/HMI vector magnetic data products (SHARP parameters).
#
# ### **Technical Contributions**
#
# - We provide the community with a new tool to predict solar flares.
#
#
# ### **Methodology**
#
# Here we present a flare prediction system, named FlareML, for predicting solar flares using machine learning (ML) based on HMI’s vector magnetic data products. Specifically, we construct training data by utilizing the physical parameters provided by the Space-weather HMI Active Region Patches (SHARP) and categorize solar flares into four classes, namely B, C, M, X, according to the X-ray flare catalogs prepared by the National Centers for Environmental Information (NCEI). Thus, the solar flare prediction problem at hand is essentially a multi-class (i.e., four-class) classification problem. The FlareML system employs four machine learning methods to tackle this multi-class prediction problem. These four methods are: (i) ensemble (ENS), (ii) random forests (RF), (iii) multilayer perceptrons (MLP), and (iv) extreme learning machines (ELM). ENS works by taking the majority vote of the results obtained from RF, MLP and ELM. This notebook leverages python machine learning and visualization packages: matplotlib, numpy, scikit-learn, sklearn-extensions, and pandas. It describes the steps on how to use the FlareML tool to predict solar flare types: B, C, M, and X. The notebook is trained and tested on sample data sets to show flare predictions and their accuracies in graphical bar plots. FlareML is the backend of an online machine-learning-as-a-service system accessible at: https://nature.njit.edu/spacesoft/DeepSun/.
#
# Notes: <br>
# <ul>
# <li>Some models used in FlareML are not deterministic due to the randomness of their processes. Therefore, these models do not make the same prediction after re-training.
# </li>
# <li>
# Detailed information about the parameters used for each model can be found in our published paper: <a href='https://iopscience.iop.org/article/10.1088/1674-4527/21/7/160'>https://iopscience.iop.org/article/10.1088/1674-4527/21/7/160</a>
# </li>
# </ul>
#
#
#
# ### **Funding**
# This work was supported by U.S. NSF grants AGS-1927578 and AGS-1954737.
#
# ### **Keywords**
# keywords=["Flare", "Prediction", "Machine", "Learning", "SHARP"]
#
# ### **Citation**
# To cite this notebook: <NAME>, <NAME>, & <NAME>. Predicting Solar Flares with Machine Learning, available at: <a href='https://github.com/ccsc-tools/FlareML/blob/main/YA_01_PredictingSolarFlareswithMachineLearning.ipynb' target='new'>https://github.com/ccsc-tools/FlareML/blob/main/YA_01_PredictingSolarFlareswithMachineLearning.ipynb</a>.
#
#
#
# ### **Acknowledgements**
#
# We thank the team of SDO/HMI for producing vector magnetic data products. The flare catalogs were prepared by and made available through NOAA NCEI.
# # **Setup**
#
# **Installation on Local Machine**<br>
# Running this notebook in a local machine requires Python version 3.8.x with the following packages and their version:
#
# |Library | Version | Description |
# |:---|---|:---|
# |matplotlib|3.4.2| Graphics and visualization|
# |numpy| 1.19.5| Array manipulation|
# |scikit-learn| 0.24.2| Machine learning|
# | sklearn-extensions | 0.0.2 | Extension for scikit-learn |
# | pandas|1.2.4| Data loading and manipulation|
# You may install the package using Python pip packages manager as follows:
#
# pip install matplotlib==3.4.2 numpy==1.19.5 scikit-learn==0.24.2 sklearn-extensions==0.0.2 pandas==1.2.4
#
#
#
# **Library Import**<br>
# The following libraries need to be imported.
# +
import warnings
warnings.filterwarnings('ignore')
# Data manipulation
import pandas as pd
import numpy as np
# Training the models
# The following libraries are used to train the algorithms: Random Forest, MLP, and ELM.
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn_extensions.extreme_learning_machines.elm import GenELMClassifier
from sklearn_extensions.extreme_learning_machines.random_layer import RBFRandomLayer, MLPRandomLayer
# Visualizations
import matplotlib.pyplot as plt
from flareml_utils import plot_custom_result
# Running the training, testing and prediction.
from flareml_train import train_model
from flareml_test import test_model
# -
# # **Data Processing and Analysis**
# We created and stored 845 data samples in
# our database accessible at <a target='new' href='https://nature.njit.edu/spacesoft/Flare-Predict/'>https://nature.njit.edu/spacesoft/Flare-Predict/</a>, where each data sample
# contains values of 13 physical parameters or features. The two digits following a class label (B,
# C, M, X) are ignored in performing flare prediction. The time
# point of a data sample is the beginning time (00:00:01 early
# morning) of the start date of a flare and the label of the data
# sample is the class which the flare belongs to. These labeled
# data samples are used to train the FlareML system.
#
# For this notebook, we use sample data sets for training and testing.
# # **Binder**
#
# This notebook is Binder enabled and can be run on <a target='new' href='https://mybinder.org/'>mybinder.org</a> by using the image link below:
#
# <p float="left"> <a href='https://mybinder.org/v2/gh/ccsc-tools/FlareML/HEAD?labpath=YA_01_PredictingSolarFlareswithMachineLearning.ipynb' target='new'><img align="left" src='https://mybinder.org/badge_logo.svg'></img></a></p>
#
# <br><br>
# Please note that starting Binder might take some time to create and start the image.
#
#
# # **FlareML Workflow and Results**
#
# ### **Data Preparation and Loading**
# The data folder includes two sub-directories: train_data and test_data.
# * The train_data includes a CSV training data file that is used to train the model.
# * The test_data includes a CSV test data file that is used to predict the included flares.
#
# The files are loaded and used during the testing and training process.
# ### **Predicting with Pretrained Models**
# There are default and pretrained models that can be used to predict without running your own trained model. The modelid is set to default_model which uses all pretrained algorithms.
from flareml_test import test_model
args = {'test_data_file': 'data/test_data/flaringar_simple_random_40.csv',
'modelid': 'default_model'}
result = test_model(args)
# **Plotting the Pretrained Models Results**<br>
from flareml_utils import plot_result
plot_result(result)
# ### **ENS Model Training and Testing**
# You may train the model with your own data or train the model with the default data.
#
# **ENS Model Training with Default Data**<br>
# Here, we show how to train the model with default data.
# To train the model with your own data:
# 1. You should first upload your file to the data directory (in the left hand side file list).
# 2. Edit the args variable in the following code and update the path to the training file:<br> 'train_data_file':'data/train_data/flaringar_training_sample.csv' <br>and replace the value 'data/train_data/flaringar_training_sample.csv' with your new file name.
print('Loading the train_model function...')
from flareml_train import train_model
args = {'train_data_file':'data/train_data/flaringar_training_sample.csv',
'algorithm': 'ENS',
'modelid': 'custom_model_id'
}
train_model(args)
# **Predicting with Your ENS Model**<br>
# To predict the testing data using the model you trained above, make sure the modelid value in the args variable in the following code is set exactly as the one used in the training, for example: 'custom_model_id'.
from flareml_test import test_model
args = {'test_data_file': 'data/test_data/flaringar_simple_random_40.csv',
'algorithm': 'ENS',
'modelid': 'custom_model_id'}
custom_result = test_model(args)
# **Plotting the ENS Results**<br>
# The prediction result can be plotted by passing the result variable to the function plot_custom_result as shown in the following example. The result shows the accuracy (TSS value) your model achieves for each flare class.
from flareml_utils import plot_custom_result
plot_custom_result(custom_result)
# ### **RF Model Training and Testing**
# **RF Model Training with Default Data**<br>
print('Loading the train_model function...')
from flareml_train import train_model
args = {'train_data_file':'data/train_data/flaringar_training_sample.csv',
'algorithm': 'RF',
'modelid': 'custom_model_id'
}
train_model(args)
# **Predicting with Your RF Model**<br>
from flareml_test import test_model
args = {'test_data_file': 'data/test_data/flaringar_simple_random_40.csv',
'algorithm': 'RF',
'modelid': 'custom_model_id'}
custom_result = test_model(args)
# **Plotting the RF Results**<br>
from flareml_utils import plot_custom_result
plot_custom_result(custom_result)
# ### **MLP Model Training and Testing**
# **MLP Model Training with Default Data**<br>
print('Loading the train_model function...')
from flareml_train import train_model
args = {'train_data_file':'data/train_data/flaringar_training_sample.csv',
'algorithm': 'MLP',
'modelid': 'custom_model_id'
}
train_model(args)
# **Predicting with Your MLP Model**<br>
from flareml_test import test_model
args = {'test_data_file': 'data/test_data/flaringar_simple_random_40.csv',
'algorithm': 'MLP',
'modelid': 'custom_model_id'}
custom_result = test_model(args)
# **Plotting the MLP Results**<br>
from flareml_utils import plot_custom_result
plot_custom_result(custom_result)
# ### **ELM Model Training and Testing**
# **ELM Model Training with Default Data**<br>
print('Loading the train_model function...')
from flareml_train import train_model
args = {'train_data_file':'data/train_data/flaringar_training_sample.csv',
'algorithm': 'ELM',
'modelid': 'custom_model_id'
}
train_model(args)
# **Predicting with Your ELM Model**<br>
from flareml_test import test_model
args = {'test_data_file': 'data/test_data/flaringar_simple_random_40.csv',
'algorithm': 'ELM',
'modelid': 'custom_model_id'}
custom_result = test_model(args)
# **Plotting the ELM Resluts**<br>
from flareml_utils import plot_custom_result
plot_custom_result(custom_result)
# ### **Timing**
# Please note that the execution time in mybinder varies based on the availability of resources. The average time to run the notebook is 10-15 minutes, but it could be more.
#
# # **Conclusions**
# <!-- We present a machine-learning-as-a-service framework
# (DeepSun) for solar flare prediction. This framework provides
# two interfaces: a web server where the user enters the information through a graphical interface and a programmable
# interface that can be used by any RESTful client. DeepSun employs three existing machine learning algorithms, namely
# random forests (RF), multilayer perceptrons (MLP), extreme learning machines (ELM), and an ensemble algorithm (ENS) that combines the three machine learning algorithms. Our experimental results demonstrated the good performance of the ensemble algorithm and its superiority over the three existing machine learning algorithms.
# In the current work we focus on data samples composed of SHARP physical parameters. We collect 845 data samples
# belonging to four flare classes: B, C, M, and X across 472 active regions. In addition, the Helioseismic Magnetic
# Imager (HMI) aboard the Solar Dynamics Observatory (SDO) produces continuous full-disk observations (solar images).<br>In
# future work we plan to incorporate these HMI images into our DeepSun framework and extend our previously developed deep learning techniques to directly process the images. -->
#
# We present a machine learning-based system (FlareML) for solar flare prediction. FlareML employs three existing machine learning algorithms, namely random forests (RF), multilayer perceptrons (MLP), extreme learning machines (ELM), and an ensemble algorithm (ENS) that combines the three machine learning algorithms. Our experimental results demonstrated the good performance of the ensemble algorithm and its superiority over the three existing machine learning algorithms. In the current work we focus on data samples composed of SHARP physical parameters. We collect 845 data samples belonging to four flare classes: B, C, M, and X across 472 active regions. In addition, the Helioseismic Magnetic Imager (HMI) aboard the Solar Dynamics Observatory (SDO) produces continuous full-disk observations (solar images). In future work we plan to incorporate these HMI images into our FlareML framework and extend our previously developed deep learning techniques to directly process the images for solar flare prediction.
#
# # **References**
# <ol>
# <li>
# DeepSun: Machine-Learning-as-a-Service for Solar Flare Prediction<br>
# <NAME>, <NAME> and <NAME><br>
# <a target='new' href='https://iopscience.iop.org/article/10.1088/1674-4527/21/7/160'>https://iopscience.iop.org/article/10.1088/1674-4527/21/7/160</a>
# </li>
# <li>
# Predicting Solar Flares Using SDO/HMI Vector Magnetic Data Products and the Random Forest Algorithm<br>
# <NAME>, <NAME>, <NAME> and <NAME><br>
# <a target='new' href='https://iopscience.iop.org/article/10.3847/1538-4357/aa789b'>https://iopscience.iop.org/article/10.3847/1538-4357/aa789b</a>
# </li>
# <li>
# Artificial Neural Networks: An Introduction to ANN Theory and Practice<br>
# <NAME>, <NAME>, <NAME> <br>
# <a target='new' href='https://link.springer.com/book/10.1007/BFb0027019'>https://link.springer.com/book/10.1007/BFb0027019</a>
# </li>
# <li>
# Enhanced Random Search Based Incremental Extreme Learning Machine<br>
# <NAME> and <NAME><br>
# <a target='new' href='https://www.sciencedirect.com/science/article/abs/pii/S0925231207003633?via%3Dihub'>https://www.sciencedirect.com/science/article/abs/pii/S0925231207003633?via%3Dihub</a>
# </li>
# <li>
# Predicting Solar Energetic Particles Using SDO/HMI Vector Magnetic Data Products and a
# Bidirectional LSTM Network<br>
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME><br>
# <a target='new' href='https://iopscience.iop.org/article/10.3847/1538-4365/ac5f56'>https://iopscience.iop.org/article/10.3847/1538-4365/ac5f56</a>
# </li>
# </ol>
| YA_01_PredictingSolarFlareswithMachineLearning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keras - Deep Learning on Scenario A
#
# Here, we will be applying deep neural networks in an to attempt to differentiate between Tor and nonTor data from the ISCXTor2016 dataset.
# +
# DataFrame handling
import pandas as pd
# Confusion matrix function
import numpy as np
import seaborn as sn
import matplotlib.pyplot as plt
# keras Models
from keras.models import Sequential
from keras.layers import Dense
from keras.metrics import CategoricalAccuracy, Precision, Recall
# sklearn Models
from sklearn.dummy import DummyClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
# Split data with stratified cv
from sklearn.model_selection import StratifiedKFold, train_test_split
# Encoding of classifications
from sklearn.preprocessing import LabelEncoder
from keras.utils import to_categorical, normalize
print('Imports complete.')
# -
# Set up a few constants to keep track of
random_state=1
path='../../tor_dataset/Scenario-A/'
dep_var = 'class'
num_classes=0
def getXy(filename='', dep_var='', verbose=0):
"""
This function takes a filename, loads the data into a dataframe, then separates the classification data
args:
filename => str, path to csv file to be loaded
returns:
list(X,y) => data, classifications
"""
df = pd.read_csv(filename)
if verbose == 2:
print('\tBefore encoding and splitting:')
print(df.head())
# Actual data
#X = df.loc[:, df.columns != dep_var]
X = pd.DataFrame()
try:
cols = ['min_fiat',
'std_biat',
'mean_biat',
'max_biat']
X = df[cols]
except:
cols = ['min_fiat',
'total_biat',
'mean_biat',
'max_biat']
X = df[cols]
if verbose == 1 or verbose == 2:
print('\n\tFeatures: {}'.format(X.columns))
# Set number of classes we see
num_classes = df[dep_var].nunique()
# Classifications
encoder = LabelEncoder()
y = encoder.fit_transform(df[dep_var])
if verbose == 2:
print('\tClassification encoding:')
for i in range(len(encoder.classes_)):
print('\t{} => {}'.format(i, encoder.classes_[i]))
print('\tAfter encoding and splitting:')
print('\tX = ')
print(X.head())
print('\n\ty = ')
print(y[:5])
# X holds the data while y holds the classifications
return X, y
# +
def show_conf_matrix(model, X_test, y_test, names):
# Techniques from https://stackoverflow.com/questions/29647749/seaborn-showing-scientific-notation-in-heatmap-for-3-digit-numbers
# and https://stackoverflow.com/questions/35572000/how-can-i-plot-a-confusion-matrix#51163585
predictions = model.predict(X_test)
matrix = [ [ 0 for j in range(len(predictions[0])) ] for i in range(len(predictions[0])) ]
for i in range(len(predictions)):
pred = predictions[i]
test = y_test[i]
guess = np.argmax(pred)
actual = np.argmax(test)
matrix[actual][guess] += 1
df_cm = pd.DataFrame(matrix, range(len(matrix)), range(len(matrix)))
int_cols = df_cm.columns
df_cm.columns = names
df_cm.index = names
plt.figure(figsize=(10,7))
sn.set(font_scale=1.5) # for label size
ax = sn.heatmap(df_cm, annot=True, annot_kws={"size": 16}, fmt='g', cmap=sn.color_palette("Blues")) # font size
ax.set_ylabel('Actual')
ax.set_xlabel('Predicted')
plt.show()
def create_dnn_model(metrics=['accuracy']):
model = Sequential([
Dense(64, input_shape=(4,)),
Dense(32, activation='relu'),
Dense(2, activation='softmax')
])
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=metrics)
return model
# +
# Just run a DNN experiment
# All of the data files
files=['TimeBasedFeatures-10s-TOR-NonTOR.csv',
'TimeBasedFeatures-15s-TOR-NonTOR.csv',
'TimeBasedFeatures-30s-TOR-NonTOR.csv',
'TimeBasedFeatures-60s-TOR-NonTOR.csv',
'TimeBasedFeatures-120s-TOR-NonTOR.csv']
# Gathered from the getXy function
#0 => NONTOR
#1 => TOR
names = ['NONTOR', 'TOR']
for file in files:
print('Training for {}'.format(file), end='')
X, y = getXy(path + file, dep_var, verbose=0)
X = normalize(X)
y = to_categorical(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=random_state)
# Set up the metrics
metrics = [
CategoricalAccuracy(),
Recall(),
Precision()
]
# Deep Neural Network
dnn = create_dnn_model(metrics)
dnn.fit(X_train, y_train, epochs=25, verbose=0)
results = dnn.evaluate(X_test, y_test, verbose=0)
print('\nMetrics:')
print('\tLoss\tAcc\tRecall\tPrecision')
for result in results:
print('\t{:.2f}'.format(result), end='')
# Show a confusion matrix
show_conf_matrix(dnn, X_test, y_test, names)
# -
from math import sqrt
def get_std(x=[], xbar=0):
o2=0
for xi in x:
o2 += (xi - xbar)**2
o2 /= len(x)-1
return sqrt(o2)
# +
# Lists for accuracies collected from models
list_dummy = []
list_dt = []
list_knn = []
list_dnn = []
std_dummy = []
std_dt = []
std_knn = []
std_dnn = []
# All of the data files
files=['TimeBasedFeatures-10s-TOR-NonTOR.csv',
'TimeBasedFeatures-15s-TOR-NonTOR.csv',
'TimeBasedFeatures-30s-TOR-NonTOR.csv',
'TimeBasedFeatures-60s-TOR-NonTOR.csv',
'TimeBasedFeatures-120s-TOR-NonTOR.csv']
for file in files:
print('Training for {}...'.format(file), end='')
# Load in the data
X, y = getXy(path + file, dep_var, verbose=1)
# Mean accuracies for each model
mean_dummy = 0 # This is the worst kind of dummy
mean_dt = 0
mean_knn = 0
mean_dnn = 0
# Keep to calculate std
results_dummy = []
results_dt = []
results_knn = []
results_dnn = []
# 10-fold Stratified Cross-Validation
n_splits = 10
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=random_state)
for train_idxs, test_idxs in skf.split(X, y):
# Define the training and testing sets
X_train, X_test = X.iloc[train_idxs], X.iloc[test_idxs]
y_train, y_test = y[train_idxs], y[test_idxs]
# Create a different version of the y_train and y_test for the Deep Neural Network
y_train_dnn = to_categorical(y_train, num_classes=num_classes)
y_test_dnn = to_categorical(y_test, num_classes=num_classes)
# Initialize the sklearn models
dummy = DummyClassifier(strategy='most_frequent')
dt = DecisionTreeClassifier(random_state=random_state)
knn = KNeighborsClassifier()
# Deep Neural Network
dnn = create_dnn_model()
# Train the models
dummy.fit(X_train, y_train)
dt.fit(X_train, y_train)
knn.fit(X_train, y_train)
dnn.fit(x=X_train, y=y_train_dnn, epochs=10, batch_size=20, verbose=0, validation_data=(X_test, y_test_dnn))
# Evaluate the models
results_dummy.append(dummy.score(X_test, y_test))
results_dt.append(dt.score(X_test, y_test))
results_knn.append(knn.score(X_test, y_test))
results_dnn.append( (dnn.evaluate(X_test, y_test_dnn, verbose=0) )[1] )
#print(results_dummy[-1])
#print(results_dt[-1])
#print(results_knn[-1])
#print(results_dnn[-1])
# Add the results to the running mean
mean_dummy += results_dummy[-1] / (n_splits * 1.0)
mean_dt += results_dt[-1] / (n_splits * 1.0)
mean_knn += results_knn[-1] / (n_splits * 1.0)
mean_dnn += results_dnn[-1] / (n_splits * 1.0)
# Push the mean results from all of the splits to the lists
list_dummy.append(mean_dummy)
list_dt.append(mean_dt)
list_knn.append(mean_knn)
list_dnn.append(mean_dnn)
print('done')
std_dummy.append(get_std(results_dummy, mean_dummy))
std_dt.append(get_std(results_dt, mean_dt))
std_knn.append(get_std(results_knn, mean_knn))
std_dnn.append(get_std(results_dnn, mean_dnn))
print('All trainings complete!')
# -
print(std_dummy)
# Output results
print('File\tDummy\t\tDecision Tree\t\tk-Nearest Neighbor\t\tDeep Neural Network')
print('-'*99)
for i in range(len(files)):
print('{0}\t{1:.2f}\u00B1{2:.2f}%\t{3:.2f}\u00B1{4:.2f}%\t\t{5:.2f}\u00B1{6:.2f}%\t\t\t{7:.2f}\u00B1{8:.2f}%'.format(files[i][18:-15], 100*list_dummy[i], 100*std_dummy[i], 100*list_dt[i], 100*std_dt[i], 100*list_knn[i], 100*std_knn[i], 100*list_dnn[i], 100*std_dnn[i]))
# +
# Just run a DNN experiment
# All of the data files
files=['downsampled_TimeBasedFeatures-10s-TOR-NonTOR.csv',
'downsampled_TimeBasedFeatures-15s-TOR-NonTOR.csv',
'downsampled_TimeBasedFeatures-30s-TOR-NonTOR.csv',
'downsampled_TimeBasedFeatures-60s-TOR-NonTOR.csv',
'downsampled_TimeBasedFeatures-120s-TOR-NonTOR.csv']
# Gathered from the getXy function
#0 => NONTOR
#1 => TOR
names = ['NONTOR', 'TOR']
for file in files:
print('Training for {}'.format(file), end='')
X, y = getXy(path + file, dep_var, verbose=0)
X = normalize(X)
y = to_categorical(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=random_state)
# Set up the metrics
metrics = [
CategoricalAccuracy(),
Recall(),
Precision()
]
# Deep Neural Network
dnn = create_dnn_model(metrics)
dnn.fit(X_train, y_train, epochs=25, verbose=0)
results = dnn.evaluate(X_test, y_test, verbose=0)
print('\nMetrics:')
print('\tLoss\tAcc\tRecall\tPrecision')
for result in results:
print('\t{:.2f}'.format(result), end='')
# Show a confusion matrix
show_conf_matrix(dnn, X_test, y_test, names)
# +
# Lists for accuracies collected from models
list_dummy = []
list_dt = []
list_knn = []
list_dnn = []
std_dummy = []
std_dt = []
std_knn = []
std_dnn = []
for file in files:
print('Training for {}...'.format(file), end='')
# Load in the data
X, y = getXy(path + file, dep_var, verbose=1)
# Mean accuracies for each model
mean_dummy = 0 # This is the worst kind of dummy
mean_dt = 0
mean_knn = 0
mean_dnn = 0
# Keep to calculate std
results_dummy = []
results_dt = []
results_knn = []
results_dnn = []
# 10-fold Stratified Cross-Validation
n_splits = 10
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=random_state)
for train_idxs, test_idxs in skf.split(X, y):
# Define the training and testing sets
X_train, X_test = X.iloc[train_idxs], X.iloc[test_idxs]
y_train, y_test = y[train_idxs], y[test_idxs]
# Create a different version of the y_train and y_test for the Deep Neural Network
y_train_dnn = to_categorical(y_train, num_classes=num_classes)
y_test_dnn = to_categorical(y_test, num_classes=num_classes)
# Initialize the sklearn models
dummy = DummyClassifier(strategy='most_frequent')
dt = DecisionTreeClassifier(random_state=random_state)
knn = KNeighborsClassifier()
# Deep Neural Network
dnn = create_dnn_model()
# Train the models
dummy.fit(X_train, y_train)
dt.fit(X_train, y_train)
knn.fit(X_train, y_train)
dnn.fit(x=X_train, y=y_train_dnn, epochs=10, batch_size=20, verbose=0, validation_data=(X_test, y_test_dnn))
# Evaluate the models
results_dummy.append(dummy.score(X_test, y_test))
results_dt.append(dt.score(X_test, y_test))
results_knn.append(knn.score(X_test, y_test))
results_dnn.append( (dnn.evaluate(X_test, y_test_dnn, verbose=0) )[1] )
#print(results_dummy[-1])
#print(results_dt[-1])
#print(results_knn[-1])
#print(results_dnn[-1])
# Add the results to the running mean
mean_dummy += results_dummy[-1] / (n_splits * 1.0)
mean_dt += results_dt[-1] / (n_splits * 1.0)
mean_knn += results_knn[-1] / (n_splits * 1.0)
mean_dnn += results_dnn[-1] / (n_splits * 1.0)
# Push the mean results from all of the splits to the lists
list_dummy.append(mean_dummy)
list_dt.append(mean_dt)
list_knn.append(mean_knn)
list_dnn.append(mean_dnn)
print('done')
std_dummy.append(get_std(results_dummy, mean_dummy))
std_dt.append(get_std(results_dt, mean_dt))
std_knn.append(get_std(results_knn, mean_knn))
std_dnn.append(get_std(results_dnn, mean_dnn))
print('All trainings complete!')
# -
# Output results
print('File\tDummy\t\tDecision Tree\t\tk-Nearest Neighbor\t\tDeep Neural Network')
print('-'*99)
for i in range(len(files)):
print('{0}\t{1:.2f}\u00B1{2:.2f}%\t{3:.2f}\u00B1{4:.2f}%\t\t{5:.2f}\u00B1{6:.2f}%\t\t\t{7:.2f}\u00B1{8:.2f}%'.format(files[i][30:-15], 100*list_dummy[i], 100*std_dummy[i], 100*list_dt[i], 100*std_dt[i], 100*list_knn[i], 100*std_knn[i], 100*list_dnn[i], 100*std_dnn[i]))
| notebooks/keras scenario a experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Investigating The Sampling Theorem
# In this section, we investigate the implications of the sampling theorem. Here is the usual statement of the theorem from wikipedia:
#
# *"If a function $x(t)$ contains no frequencies higher than B hertz, it is completely determined by giving its ordinates at a series of points spaced 1/(2B) seconds apart."*
#
# Since a function $x(t)$ is a function from the real line to the real line, there are uncountably many points between any two ordinates, so sampling is a massive reduction of data since it only takes a tiny number of points to completely characterize the function. This is a powerful idea worth exploring. In fact, we have seen this idea of reducing a function to a discrete set of numbers before in Fourier series expansions where (for periodic $x(t)$)
#
# $$ a_n = \frac{1}{T} \int^{T}_0 x(t) \exp (-j \omega_n t )dt $$
#
#
# with corresponding reconstruction as:
#
# $$ x(t) = \sum_k a_n \exp( j \omega_n t) $$
#
#
# But here we are generating discrete points $a_n$ by integrating over the **entire** function $x(t)$, not just evaluating it at a single point. This means we are collecting information about the entire function to compute a single discrete point $a_n$, whereas with sampling we are just taking individual points in isolation.
#
# Let's come at this the other way: suppose we are given a set of samples $[x_1,x_2,..,x_N]$ and we are then told to reconstruct the function. What would we do? This is the kind of question seldom asked because we typically sample, filter, and then do something else without trying to reconstruct the function from the samples directly.
#
# Returning to our reconstruction challenge, perhaps the most natural thing to do is draw a straight line between each of the points as in linear interpolation. The next block of code takes samples of the $sin$ over a single period and draws a line between sampled ordinates.
# +
import numpy as np
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
f = 1.0 # Hz, signal frequency
fs = 20.0 # Hz, sampling rate (ie. >= 2*f)
t = np.arange(-1, 1+1/fs, 1/fs) # sample interval, symmetric for convenience later
x = np.sin(2*np.pi*f*t)
ax.plot(t, x, 'o-')
ax.set_xlabel('time', fontsize=18)
ax.set_ylabel('amplitude', fontsize=18)
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# In this plot, notice how near the extremes of the $sin$ at $t=1/(4f)$ and $t=3/(4 f)$, we are taking the same density of points since the sampling theorem makes no requirement on *where* we should sample as long as we sample at a regular intervals. This means that on the up and down slopes of the $sin$, which are obviously linear-looking and where a linear approximation is a good one, we are taking the same density of samples as near the curvy peaks. Here's a bit of code that zooms in to the first peak to illustrate this.
# +
fig, ax = plt.subplots()
ax.plot(t, x, 'o-')
ax.axis(xmin = 1/(4*f)-1/fs*3, xmax = 1/(4*f)+1/fs*3, ymin = 0, ymax = 1.1 )
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# To drive this point home (and create some cool matplotlib plots), we can construct the piecewise linear interpolant and compare the quality of the approximation using ``numpy.piecewise``:
# +
interval = [] # piecewise domains
apprx = [] # line on domains
# build up points *evenly* inside of intervals
tp = np.hstack([np.linspace(t[i], t[i+1], 20, False) for i in range(len(t)-1) ])
# construct arguments for piecewise
for i in range(len(t)-1):
interval.append(np.logical_and(t[i] <= tp,tp < t[i+1]))
apprx.append((x[i+1]-x[i])/(t[i+1]-t[i])*(tp[interval[-1]]-t[i]) + x[i])
x_hat = np.piecewise(tp, interval, apprx) # piecewise linear approximation
# -
# Now, we can examine the squared errors in the interpolant. The following snippet plots the $sin$ and with the filled-in error of the linear interpolant.
ax1 = plt.figure().add_subplot(111)
ax1.fill_between(tp, x_hat, np.sin(2*np.pi*f*tp), color='black')
ax1.set_xlabel('time', fontsize=18)
ax1.set_ylabel('Amplitude', fontsize=18)
ax1.set_title('Errors with Piecewise Linear Interpolant')
ax2 = ax1.twinx()
sqe = (x_hat - np.sin(2*np.pi*f*tp))**2
ax2.plot(tp, sqe, 'r')
ax2.axis(xmin=-1, ymax=sqe.max() )
ax2.set_ylabel('squared error', color='r', fontsize=18)
# ax1.figure.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# Note: I urge you to change the $fs$ sampling rate in the code above then rerun this notebook to see how these errors change with more/less sampling points.
# Now, we could pursue this line of reasoning with higher-order polynomials instead of just straight lines, but this would all eventually take us to the same conclusion; namely, that all of these approximations improve as the density of sample points increases, which is the *exact* opposite of what the sampling theorem says --- there is *sparse* set of samples points that will retrieve the original function. Furthermore, we observed that the quality of the piecewise linear interpolation is sensitive to *where* the sample points are taken and the sampling theorem is so powerful that it *has no such requirement*.
# ## Reconstruction
# Let's look at this another way by examing the Fourier Transform of a signal that is bandlimited and thus certainly satisfies the hypothesis of the sampling theorem:
#
# $$ X(f) = 0~ where ~ |f|> W $$
#
# Now, the inverse Fourier transform of this is the following:
#
# $$ x(t) = \int_{-W}^W X(f) e^{j 2 \pi f t} df $$
#
# We can take the $X(f)$ and expand it into a Fourier series by pretending that it is periodic with period $2 W$. Thus, we can formally write the following:
#
# $$ X(f) = \sum_k a_k e^{ - j 2 \pi k f/(2 W) } $$
#
# we can compute the coefficients $a_k$ as
#
# $$ a_k = \frac{1}{2 W} \int_{-W}^W X(f) e^{ j 2 \pi k f/(2 W) } df $$
#
# These coefficients bear a striking similarity to the $x(t)$ integral we just computed above. In fact, by lining up terms, we can write:
#
# $$ a_k = \frac{1}{2 W} x \left( t = \frac{k}{2 W} \right) $$
#
# Now, we can write out $X(f)$ in terms of this series and these $a_k$ and then invert the Fourier transform to obtain the following:
#
# $$ x(t) = \int_{-W}^W \sum_k a_k e^{ - j 2 \pi k f/(2 W) } e^{j 2 \pi f t} df $$
#
# substitute for $a_k$
#
# $$ x(t) = \int_{-W}^W \sum_k ( \frac{1}{2 W} x( t = \frac{k}{2 W} ) ) e^{ - j 2 \pi k f/(2 W) } e^{j 2 \pi f t} df $$
#
# switch summation and integration (usually dangerous, but OK here)
#
# $$ x(t) = \sum_k x(t = \frac{k}{2 W}) \frac{1}{2 W} \int_{-W}^W e^{ - j 2 \pi k f/(2 W) +j 2 \pi f t} df $$
#
# which gives finally:
#
# $$ x(t) = \sum_k x(t = \frac{k}{2 W}) \frac{sin(\pi (k-2 t W))} {\pi (k- 2 t W)} $$
#
# And this what we have been seeking! A formula that reconstructs the function from its samples. Let's try it!
# Note that since our samples are spaced at $t= k/f_s $, we'll use $ W= f_s /2 $ to line things up.
# +
ax = plt.figure().add_subplot(111)
t = np.linspace(-1, 1, 100) # redefine this here for convenience
ts = np.arange(-1, 1+1/fs, 1/fs) # sample points
num_coeffs = len(ts)
sm = 0
for k in range(-num_coeffs, num_coeffs): # since function is real, need both sides
sm += np.sin(2*np.pi*(k/fs))*np.sinc(k - fs * t)
ax.plot(t, sm,'--', t, np.sin(2*np.pi*t), ts, np.sin(2*np.pi*ts), 'o')
ax.set_title('sampling rate=%3.2f Hz' % fs)
# ax.figure.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# We can do the same check as we did for the linear interpolant above as
# +
ax1 = plt.figure().add_subplot(111)
ax1.fill_between(t,sm, np.sin(2*np.pi*f*t), color='black')
ax1.set_ylabel('Amplitude',fontsize=18)
ax2 = ax1.twinx()
sqe = (sm - np.sin(2*np.pi*f*t))**2
ax2.plot(t, sqe,'r')
ax2.axis(xmin=0, ymax=sqe.max())
ax2.set_ylabel('squared error', color='r', fontsize=18)
ax1.set_title('Errors with sinc Interpolant')
# ax1.figure.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# These interpolating functions are called the "Whittaker" interpolating functions. Let's examine these functions more closely with the following code
# +
fig = plt.figure()
ax = fig.add_subplot(111) # create axis handle
k = 0
fs = 2 # makes this plot easier to read
ax.plot (t, np.sinc(k - fs * t),
t, np.sinc(k+1 - fs * t), '--', k/fs,1, 'o', (k)/fs, 0,'o',
t, np.sinc(k-1 - fs * t), '--', k/fs,1, 'o', (-k)/fs, 0,'o'
)
ax.hlines(0, -1, 1)
ax.vlines(0, -.2, 1)
ax.annotate('sample value goes here',
xy=(0, 1),
xytext=(-1+.1,1.1),
arrowprops={'facecolor':'red','shrink':0.05},
)
ax.annotate('no interference here',
xy=(0,0),
xytext=(-1+.1,0.5),
arrowprops={'facecolor':'green','shrink':0.05},
)
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# The vertical line in the previous plot shows that where one function has a peak, the other function has a zero. This is why when you put samples at each of the peaks, they match the sampled function exactly at those points. In between those points, the crown shape of the functions fills in the missing values. Furthermore, as the figure above shows, there is no interference between the functions sitting on each of the interpolating functions because the peak of one is perfectly aligned with the zero of the others (dotted lines). Thus, the sampling theorem says that the filled-in values are drawn from the curvature of the sinc functions, not straight lines as we investigated earlier.
#
# As an illustration, the following code shows how the individual Whittaker functions (dashed lines) are assembled into the final approximation (black-line) using the given samples (blue-dots). I urge you to play with the sampling rate to see what happens. Note the heavy use of `numpy` broadcasting in this code instead of the multiple loops we used earlier.
# +
fs = 20.0 # sampling rate
k = np.array(sorted(set((t*fs).astype(int)))) # sorted coefficient list
fig, ax = plt.subplots()
ax.plot(t,(np.sin(2*np.pi*(k[:,None]/fs))*np.sinc(k[:,None]-fs*t)).T,'--', # individual whittaker functions
t,(np.sin(2*np.pi*(k[:,None]/fs))*np.sinc(k[:,None]-fs*t)).sum(axis=0),'k-', # whittaker interpolant
k/fs, np.sin(2*np.pi*k/fs),'ob')# samples
ax.set_xlabel('time',fontsize=14)
ax.axis((-1.1,1.1,-1.1,1.1));
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# However, if you've been following carefully, you should be somewhat uncomfortable with the second to the last plot that shows the errors in the Whittaker interpolation. Namely, *why are there any errors*? Does not the sampling theorem guarantee exact-ness which should mean no error at all? It turns out that answering this question takes us further into the implications of the sampling theorem, but that is the topic of our next post.
#
#
#
# ## Summary
# In this section, we started our investigation of the famous sampling theorem that is the bedrock of the entire field of signal processing and we asked if we could reverse-engineer the consquences of the sampling theorem by reconstructing a sampled function from its discrete samples. This led us to consider the famous *Whittaker interpolator*, whose proof we sketched here. However, after all this work, we came to a disturbing conclusion regarding the exact-ness of the sampling theorem that we will investigate in a subsequent posting. In the meantime, I urge you to start at the top of notebook and play with the sampling frequency, and maybe even the sampled function and see what else you can discover about the sampling theorem.
# ## References
#
# * This is in the [IPython Notebook format](http://ipython.org/) and was converted to HTML using [nbconvert](https://github.com/ipython/nbconvert).
#
# * See [Signal analysis](http://books.google.com/books?id=Re5SAAAAMAAJ) for more detailed mathematical development.
#
# * The IPython Notebook corresponding to this post can be found [here](https://github.com/unpingco/Python-for-Signal-Processing/blob/master/Sampling_Theorem.ipynb).
| notebook/Sampling_Theorem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # scRFE source V3 - kidney split by celltype
# +
# cleaning up code after meeting with Sevahn
# +
# all code broken up into separate arguments for scRFE
# -
# Imports
import numpy as np
import pandas as pd
import scanpy as sc
from anndata import read_h5ad
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import SelectFromModel
from sklearn.metrics import accuracy_score
from sklearn.feature_selection import RFE
from sklearn.feature_selection import RFECV
# read in data
adata = read_h5ad('/Users/madelinepark/Downloads/Kidney_facs.h5ad')
tiss = adata
# +
# tiss.obs['cell_ontology_class']
# -
# split data for training
def loc_split_function(tiss, feature='cell_ontology_class'):
tiss.obs['feature_type_of_interest'] = 'rest'
for c in list(set(tiss.obs[feature])):
feature_of_interest = c
tiss.obs.loc[tiss.obs[tiss.obs[feature] == feature_of_interest].index,'feature_type_of_interest'] = feature_of_interest
feat_labels = tiss.var_names
X = tiss.X
y = tiss.obs['feature_type_of_interest']
return X, y, feature, feat_labels #this is returning only the last thing in the loop
# create random forest and selector, then train
def train_function(X, y, test_size, random_state):
print('training...')
loc_split = loc_split_function(tiss=tiss, feature='cell_ontology_class')
X = loc_split[0]
y = loc_split[1]
feat_labels = loc_split[3] #this should not be hardcoded #genes be argument
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.05, random_state=0)
clf = RandomForestClassifier(n_estimators=1000, random_state=0, n_jobs=-1, oob_score=True)
selector = RFECV(clf, step=0.2, cv=3, n_jobs=4)
clf.fit(X_train, y_train)
selector.fit(X_train, y_train)
feature_selected = feat_labels[selector.support_]
return selector, clf, feat_labels, feature_selected, selector.estimator_.feature_importances_, X_train, X_test, y_train, y_test
# cd /Users/madelinepark/src3/scRFE/scRFE-results
# result write
def result_write (c, feature_selected, tiss, feature='cell_ontology_class',test_size=0.05, random_state=0):
results_df = pd.DataFrame()
print('result writing...')
loc_split = loc_split_function(tiss=tiss, feature= feature)
X = loc_split[0]
y = loc_split[1]
train = train_function(X, y, test_size, random_state)
feat_labels = train[0]
feature_selected = train[3]
gini_scores = train[4]
column_headings = []
column_headings.append(c)
column_headings.append(c + '_gini')
resaux = pd.DataFrame(columns=column_headings)
resaux[c] = feature_selected
resaux[c + '_gini'] = (gini_scores)
results_df = pd.concat([results_df,resaux],axis=1)
tiss.obs['feature_type_of_interest'] = 'rest'
file_name = feature + c + ".csv"
print(results_df)
results_df.to_csv(file_name) #trying to save results as a csv
return results_df
# +
# gini_3_m = result_write (c='3m', feature_selected, tiss, feature='age',test_size=0.05, random_state=0)
# gini_24_m = result_write (c='24m', feature_selected, tiss, feature='age',test_size=0.05, random_state=0)
# -
# combined functions
def scRFE (tiss=tiss, X=tiss.X, feature='cell_ontology_class', n_estimators=1000, random_state=0, n_jobs=-1, oob_score=True, test_size = 0.05, step=0.2, cv=5) :
all_trees = []
for c in list(set(tiss.obs[feature])):
print(c)
loc_split = loc_split_function(tiss=tiss, feature= feature)
X = loc_split[0] #change age to feature
y = loc_split[1]
feature = loc_split[2]
feat_labels = loc_split[3]
train = train_function(X, y, test_size, random_state)
feature_selected = train[1]
X_train = train[2]
X_test = train[3]
y_train = train[4]
y_test = train[5]
all_trees += [result_write(c, feature_selected, tiss, feature=feature,test_size=0.05, random_state=0)]
#returns data frames from SCRFE in order
tiss.obs['age_type_of_interest'] = 'rest'
return all_trees
# results_df.to_csv('scRFE-results.csv') #trying to save results as a csv - this didnt work
scRFE(tiss=tiss, feature='cell_ontology_class', n_estimators=10, random_state=0, n_jobs=-1, oob_score=True, test_size = 0.05, step=0.2, cv=5)
makeAllTrees = scRFE(tiss=tiss, feature='cell_ontology_class', n_estimators=10, random_state=0, n_jobs=-1, oob_score=True, test_size = 0.05, step=0.2, cv=5)
# age3 = makeAllTrees[1]
# age24 = makeAllTrees[0]
# +
# run function
# scRFE(tiss=tiss, feature='age', n_estimators=1000, random_state=0, n_jobs=-1, oob_score=True, test_size = 0.05, step=0.2, cv=5)
# +
# how do i get it to append after so the last dataframe doesnt completely overwrite the previous one
# +
# # combined functions
# def scRFE_save (tiss=tiss, X=tiss.X, feature='age', n_estimators=1000, random_state=0, n_jobs=-1, oob_score=True, test_size = 0.05, step=0.2, cv=5) :
# for c in list(set(tiss.obs[feature])):
# print(c)
# X = loc_split(tiss=tiss, feature= feature)[0] #change age to feature
# y = loc_split(tiss=tiss, feature= feature)[1]
# feature = loc_split(tiss=tiss, feature= feature)[2]
# feat_labels = loc_split(tiss=tiss, feature= feature)[3]
# feature_selected = train(X, y, test_size, random_state)[1]
# X_train = train(X, y, test_size, random_state)[2]
# X_test = train(X, y, test_size, random_state)[3]
# y_train = train(X, y, test_size, random_state)[4]
# y_test = train(X, y, test_size, random_state)[5]
# result_write(c, feature_selected, tiss, feature=feature,test_size=0.05, random_state=0)
# tiss.obs['age_type_of_interest'] = 'rest'
# # not sorted
# -
| scripts/practiceScripts/.ipynb_checkpoints/scRFEsourceV3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Competition Details
#
# Hosted by - HackerEarth - June 2021
#
# Dataset and Problem - https://www.kaggle.com/infernape/fast-furious-and-insured
#
# __notebooks__:
# - https://www.kaggle.com/ashuto7h/fast-furious-crash
#
# - https://www.kaggle.com/ashuto7h/2-fast-furious-regression
#
# My final Score - 46.875
#
# Winner Final Score - 58.359
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" jupyter={"source_hidden": true} papermill={"duration": 0.236356, "end_time": "2021-06-04T04:53:06.787967", "exception": false, "start_time": "2021-06-04T04:53:06.551611", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy # linear algebra
import pandas # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
print(dirname)
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + jupyter={"source_hidden": true} papermill={"duration": 0.066539, "end_time": "2021-06-04T04:53:06.875772", "exception": false, "start_time": "2021-06-04T04:53:06.809233", "status": "completed"} tags=[]
train_df = pandas.read_csv('/kaggle/input/fast-furious-and-insured/Fast_Furious_Insured/train.csv')
train_df
# + jupyter={"source_hidden": true} papermill={"duration": 0.045409, "end_time": "2021-06-04T04:53:06.944469", "exception": false, "start_time": "2021-06-04T04:53:06.899060", "status": "completed"} tags=[] language="bash"
# mkdir /kaggle/working/train/damaged -p
# mkdir /kaggle/working/train/undamaged -p
# mkdir /kaggle/working/test/damaged -p
# mkdir /kaggle/working/test/undamaged -p
# + jupyter={"source_hidden": true} papermill={"duration": 13.898128, "end_time": "2021-06-04T04:53:20.863271", "exception": false, "start_time": "2021-06-04T04:53:06.965143", "status": "completed"} tags=[]
from tqdm.notebook import tqdm
for dirname,_,filenames in tqdm(os.walk('/kaggle/input/fast-furious-and-insured/Fast_Furious_Insured/trainImages')):
for file in filenames:
condition = train_df.loc[train_df['Image_path'] == file, 'Condition']
if condition.iloc[0] == 0:
os.system(f'cp {dirname}/{file} /kaggle/working/train/undamaged')
else:
os.system(f'cp {dirname}/{file} /kaggle/working/train/damaged')
# + jupyter={"source_hidden": true} papermill={"duration": 5.792645, "end_time": "2021-06-04T04:53:26.677540", "exception": false, "start_time": "2021-06-04T04:53:20.884895", "status": "completed"} tags=[]
import matplotlib.pyplot as pyplot
import seaborn
import keras
from keras.models import Sequential
from keras.layers import Dense, Conv2D , MaxPool2D , Flatten , Dropout
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam
from sklearn.metrics import classification_report,confusion_matrix
import tensorflow as tflow
import cv2
import os
import numpy
# + jupyter={"source_hidden": true} papermill={"duration": 10.07615, "end_time": "2021-06-04T04:53:36.775417", "exception": false, "start_time": "2021-06-04T04:53:26.699267", "status": "completed"} tags=[]
labels = ['damaged','undamaged']
img_size = 224
def get_data(data_dir):
data = []
for label in labels:
path = os.path.join(data_dir, label)
class_num = labels.index(label)
for img in os.listdir(path):
try:
img_arr = cv2.imread(os.path.join(path, img))[...,::-1] #convert BGR to RGB format
resized_arr = cv2.resize(img_arr, (img_size, img_size)) # Reshaping images to preferred size
data.append([resized_arr, class_num])
except Exception as e:
print(e)
return numpy.array(data)
train = get_data('/kaggle/working/train')
# + jupyter={"source_hidden": true} papermill={"duration": 0.17061, "end_time": "2021-06-04T04:53:36.969233", "exception": false, "start_time": "2021-06-04T04:53:36.798623", "status": "completed"} tags=[]
l = []
for i in train:
if(i[1] == 0):
l.append("damaged")
else:
l.append("undamaged")
seaborn.set_style('darkgrid')
seaborn.countplot(l)
# + papermill={"duration": 0.622094, "end_time": "2021-06-04T04:53:37.614326", "exception": false, "start_time": "2021-06-04T04:53:36.992232", "status": "completed"} tags=[]
x_train = []
y_train = []
for feature, label in train:
x_train.append(feature)
y_train.append(label)
# Normalize the data
x_train = numpy.array(x_train) / 255
x_train.reshape(-1, img_size, img_size, 1)
y_train = numpy.array(y_train)
# + papermill={"duration": 0.030763, "end_time": "2021-06-04T04:53:37.668321", "exception": false, "start_time": "2021-06-04T04:53:37.637558", "status": "completed"} tags=[]
numpy.unique(y_train)
# + [markdown] papermill={"duration": 0.023356, "end_time": "2021-06-04T04:53:37.714578", "exception": false, "start_time": "2021-06-04T04:53:37.691222", "status": "completed"} tags=[]
# ### Smote Oversampling
# + jupyter={"source_hidden": true} papermill={"duration": 0.030094, "end_time": "2021-06-04T04:53:37.767958", "exception": false, "start_time": "2021-06-04T04:53:37.737864", "status": "completed"} tags=[]
x_train.shape
# + jupyter={"source_hidden": true} papermill={"duration": 0.03121, "end_time": "2021-06-04T04:53:37.822911", "exception": false, "start_time": "2021-06-04T04:53:37.791701", "status": "completed"} tags=[]
ReX_train = x_train.reshape(1399, 224 * 224 * 3)
ReX_train.shape
# + papermill={"duration": 5.203751, "end_time": "2021-06-04T04:53:43.051677", "exception": false, "start_time": "2021-06-04T04:53:37.847926", "status": "completed"} tags=[]
from imblearn.over_sampling import SMOTE
smote = SMOTE(random_state = 42)
x_train2, y_train2 = smote.fit_resample(ReX_train,y_train)
x_train2.shape,y_train2.shape
# + papermill={"duration": 0.036158, "end_time": "2021-06-04T04:53:43.112633", "exception": false, "start_time": "2021-06-04T04:53:43.076475", "status": "completed"} tags=[]
x_train2 = x_train2.reshape(-1,224,224,3)
x_train2.shape
# + papermill={"duration": 0.160619, "end_time": "2021-06-04T04:53:43.297676", "exception": false, "start_time": "2021-06-04T04:53:43.137057", "status": "completed"} tags=[]
l = []
for i in y_train2:
if(i == 0):
l.append("damaged")
else:
l.append("undamaged")
seaborn.set_style('darkgrid')
seaborn.countplot(l)
# + [markdown] papermill={"duration": 0.026188, "end_time": "2021-06-04T04:53:43.350936", "exception": false, "start_time": "2021-06-04T04:53:43.324748", "status": "completed"} tags=[]
# # Models
# + jupyter={"source_hidden": true} papermill={"duration": 0.035159, "end_time": "2021-06-04T04:53:43.412422", "exception": false, "start_time": "2021-06-04T04:53:43.377263", "status": "completed"} tags=[]
def simple_model():
model = Sequential()
model.add(Conv2D(32,3,padding="same", activation="relu", input_shape=(224,224,3)))
model.add(MaxPool2D())
model.add(Conv2D(32, 3, padding="same", activation="relu"))
model.add(MaxPool2D())
model.add(Conv2D(64, 3, padding="same", activation="relu"))
model.add(MaxPool2D())
model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(128,activation="relu"))
model.add(Dense(2, activation="softmax"))
print('Simple Model')
model.summary()
return model
# + papermill={"duration": 0.046837, "end_time": "2021-06-04T04:53:43.484239", "exception": false, "start_time": "2021-06-04T04:53:43.437402", "status": "completed"} tags=[]
# xception model
import tensorflow as tf
from tensorflow.keras.layers import Input,Dense,Conv2D,Add
from tensorflow.keras.layers import SeparableConv2D,ReLU
from tensorflow.keras.layers import BatchNormalization,MaxPool2D
from tensorflow.keras.layers import GlobalAvgPool2D
from tensorflow.keras import Model
# creating the Conv-Batch Norm block
def conv_bn(x, filters, kernel_size, strides=1):
x = Conv2D(filters=filters,
kernel_size = kernel_size,
strides=strides,
padding = 'same',
use_bias = False)(x)
x = BatchNormalization()(x)
return x
# creating separableConv-Batch Norm block
def sep_bn(x, filters, kernel_size, strides=1):
x = SeparableConv2D(filters=filters,
kernel_size = kernel_size,
strides=strides,
padding = 'same',
use_bias = False)(x)
x = BatchNormalization()(x)
return x
# entry flow
def entry_flow(x):
x = conv_bn(x, filters =32, kernel_size =3, strides=2)
x = ReLU()(x)
x = conv_bn(x, filters =64, kernel_size =3, strides=1)
tensor = ReLU()(x)
x = sep_bn(tensor, filters = 128, kernel_size =3)
x = ReLU()(x)
x = sep_bn(x, filters = 128, kernel_size =3)
x = MaxPool2D(pool_size=3, strides=2, padding = 'same')(x)
tensor = conv_bn(tensor, filters=128, kernel_size = 1,strides=2)
x = Add()([tensor,x])
x = ReLU()(x)
x = sep_bn(x, filters =256, kernel_size=3)
x = ReLU()(x)
x = sep_bn(x, filters =256, kernel_size=3)
x = MaxPool2D(pool_size=3, strides=2, padding = 'same')(x)
tensor = conv_bn(tensor, filters=256, kernel_size = 1,strides=2)
x = Add()([tensor,x])
x = ReLU()(x)
x = sep_bn(x, filters =728, kernel_size=3)
x = ReLU()(x)
x = sep_bn(x, filters =728, kernel_size=3)
x = MaxPool2D(pool_size=3, strides=2, padding = 'same')(x)
tensor = conv_bn(tensor, filters=728, kernel_size = 1,strides=2)
x = Add()([tensor,x])
return x
# middle flow
def middle_flow(tensor):
for _ in range(8):
x = ReLU()(tensor)
x = sep_bn(x, filters = 728, kernel_size = 3)
x = ReLU()(x)
x = sep_bn(x, filters = 728, kernel_size = 3)
x = ReLU()(x)
x = sep_bn(x, filters = 728, kernel_size = 3)
x = ReLU()(x)
tensor = Add()([tensor,x])
return tensor
def exit_flow(tensor):
x = ReLU()(tensor)
x = sep_bn(x, filters = 728, kernel_size=3)
x = ReLU()(x)
x = sep_bn(x, filters = 1024, kernel_size=3)
x = MaxPool2D(pool_size = 3, strides = 2, padding ='same')(x)
tensor = conv_bn(tensor, filters =1024, kernel_size=1, strides =2)
x = Add()([tensor,x])
x = sep_bn(x, filters = 1536, kernel_size=3)
x = ReLU()(x)
x = sep_bn(x, filters = 2048, kernel_size=3)
x = GlobalAvgPool2D()(x)
x = Dense (units = 2, activation = 'softmax')(x)
return x
def xception_model():
# shape = 299 preferred
input = Input(shape = (224,224,3))
x = entry_flow(input)
x = middle_flow(x)
output = exit_flow(x)
model = Model (inputs=input, outputs=output)
# model.summary()
return model
# + [markdown] papermill={"duration": 0.024792, "end_time": "2021-06-04T04:53:43.533900", "exception": false, "start_time": "2021-06-04T04:53:43.509108", "status": "completed"} tags=[]
# ## Training
# + papermill={"duration": 5488.866308, "end_time": "2021-06-04T06:25:12.425030", "exception": false, "start_time": "2021-06-04T04:53:43.558722", "status": "completed"} tags=[]
epochs = 200
model = xception_model()
model.compile(optimizer = Adam() , loss = tflow.keras.losses.SparseCategoricalCrossentropy(from_logits=True) , metrics = ['accuracy'])
history = model.fit(x=x_train2, y=y_train2, epochs=epochs, verbose=1, validation_split=0.2, workers=2)
# + papermill={"duration": 3.793112, "end_time": "2021-06-04T06:25:19.695223", "exception": false, "start_time": "2021-06-04T06:25:15.902111", "status": "completed"} tags=[]
acc = history.history['accuracy']
loss = history.history['loss']
epochs_range = range(epochs)
pyplot.figure(figsize=(15, 15))
pyplot.subplot(2, 2, 1)
pyplot.plot(epochs_range, acc, label='Training Accuracy')
pyplot.legend(loc='lower right')
pyplot.title('Training Accuracy')
pyplot.subplot(2, 2, 2)
pyplot.plot(epochs_range, loss, label='Training Loss')
pyplot.legend(loc='upper right')
pyplot.title('Training Loss')
pyplot.show()
# + [markdown] papermill={"duration": 3.412965, "end_time": "2021-06-04T06:25:26.764059", "exception": false, "start_time": "2021-06-04T06:25:23.351094", "status": "completed"} tags=[]
# ## Prediction
# + jupyter={"source_hidden": true} papermill={"duration": 3.771614, "end_time": "2021-06-04T06:25:34.108383", "exception": false, "start_time": "2021-06-04T06:25:30.336769", "status": "completed"} tags=[]
test =[]
import pandas
import os
path = '/kaggle/input/fast-furious-and-insured/Fast_Furious_Insured/testImages'
for img in os.listdir(path):
test.append(img)
test = pandas.DataFrame(test)
test.to_csv('imgs.csv')
# + papermill={"duration": 11.570607, "end_time": "2021-06-04T06:25:49.114817", "exception": false, "start_time": "2021-06-04T06:25:37.544210", "status": "completed"} tags=[]
test =[]
path = '/kaggle/input/fast-furious-and-insured/Fast_Furious_Insured/testImages'
for img in os.listdir(path):
img_arr = cv2.imread(os.path.join(path, img))[...,::-1]
resized_arr = cv2.resize(img_arr, (img_size, img_size)) # Reshaping images to preferred size
test.append(resized_arr)
test = numpy.array(test) / 255
test.reshape(-1, img_size, img_size, 1)
test.shape
# + papermill={"duration": 3.695217, "end_time": "2021-06-04T06:25:56.249651", "exception": false, "start_time": "2021-06-04T06:25:52.554434", "status": "completed"} tags=[]
y_prob.shape
# + papermill={"duration": 6.071898, "end_time": "2021-06-04T06:26:05.759624", "exception": false, "start_time": "2021-06-04T06:25:59.687726", "status": "completed"} tags=[]
y_prob = model.predict(test)
y_classes = y_prob.argmax(axis=-1)
predictions = y_classes.reshape(1,-1)[0]
pred_df = pandas.DataFrame(predictions)
pred_df.to_csv('predict.csv')
# + papermill={"duration": 3.397323, "end_time": "2021-06-04T06:26:12.862406", "exception": false, "start_time": "2021-06-04T06:26:09.465083", "status": "completed"} tags=[]
pred_df
# + papermill={"duration": 3.690358, "end_time": "2021-06-04T06:26:20.415327", "exception": false, "start_time": "2021-06-04T06:26:16.724969", "status": "completed"} tags=[]
test_df = pandas.read_csv('/kaggle/input/fast-furious-and-insured/Fast_Furious_Insured/test.csv')
test_df
# + papermill={"duration": 3.433605, "end_time": "2021-06-04T06:26:27.284179", "exception": false, "start_time": "2021-06-04T06:26:23.850574", "status": "completed"} tags=[]
pred_df.columns = ['Condition']
combo= pandas.concat(objs= [test_df,pred_df], axis = 1)
combo
# + papermill={"duration": 3.428337, "end_time": "2021-06-04T06:26:34.420357", "exception": false, "start_time": "2021-06-04T06:26:30.992020", "status": "completed"} tags=[]
combo.to_csv('test.csv')
train_df.to_csv('train.csv')
| Hackerearth/fast_furious_insured - HE May 2021/fast-furious-crash.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from xml.dom import minidom
from sklearn.model_selection import train_test_split
import numpy as np
# +
data_directory = "./data/pan21-author-profiling-training-2021-03-14/en"
text_list = []
label_list = []
with open(data_directory+"/truth.txt") as f:
for l in f.readlines():
l = l.rstrip().split(":::")
user_id = l[0]
label = int(l[1])
xml_file = minidom.parse(data_directory+"/"+user_id+".xml")
document_list = xml_file.getElementsByTagName('document')
aux = []
for i, doc in enumerate(document_list):
aux.append(doc.firstChild.nodeValue)
aux = " ".join(aux)
text_list.append(aux)
label_list.append(label)
label_list = np.array(label_list)
print(len(text_list), "Total users")
# +
vocab_size = 20000
maxlen = 200
test_split = 0.2
val_split = 0.1
x_train_r, x_test_r, y_train, y_test = train_test_split(text_list, label_list, \
test_size=test_split, random_state=42)
tokenizer = keras.preprocessing.text.Tokenizer(num_words=vocab_size, oov_token="<OOV>")
tokenizer.fit_on_texts(x_train_r)
word_index = tokenizer.word_index
x_train, x_test = tokenizer.texts_to_sequences(x_train_r), tokenizer.texts_to_sequences(x_test_r)
print(len(x_train), "Training sequences")
print(len(x_test), "Testing sequences")
print(vocab_size, "Size of the vocabulary")
x_train = keras.preprocessing.sequence.pad_sequences(x_train)
maxlen = x_train[0].shape[0]
x_test = keras.preprocessing.sequence.pad_sequences(x_test, maxlen=maxlen)
# +
emb_dim = 16
inputs = layers.Input(shape=(maxlen,))
embs = layers.Embedding(vocab_size, emb_dim, input_length=maxlen)(inputs)
convs = embs
for _ in range(1):
convs = layers.Conv1D(filters=32, kernel_size=3, padding="same")(convs)
convs = layers.MaxPool1D(pool_size=2, strides=2, padding="valid")(convs)
convs = layers.Conv1D(filters=32, kernel_size=3, padding="same")(convs)
recs = layers.Bidirectional(layers.LSTM(8))(convs)
denses = layers.Dropout(0.2)(recs)
denses = layers.Dense(16, activation="relu")(denses)
denses = layers.Dropout(0.2)(denses)
outputs = layers.Dense(1, activation="sigmoid")(denses)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile("adam", "binary_crossentropy", metrics=["accuracy"])
model.summary()
# -
history = model.fit(
x_train, y_train, batch_size=16, epochs=50, validation_split=0.2
)
test_loss, test_acc = model.evaluate(x_test, y_test, batch_size=32)
print("%.3f" % test_acc, "Testing accuracy")
# +
from matplotlib import pyplot as plt
# Accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# Loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# -
| simple_bi_rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Biosignal Data Processing
# This notebook serves to process biosingal data collected from Emaptica4.
# ```
# user_id:
# eda.json
# hr.json
#
# # Inside a eda.json
# session_id: 123 #in case we have multiple data by the same user
# user_id: 478
# tags: ["pilot", "bio", "time-series"] # time-series tells us to expect a "data" key
# name: "EDA"
# description: "Electrodermal activity"
# channels: 1
# sampling_frequency: 3 #Hertz
# units: "microsiemens"
# ... # any other metadata
# timestamp: 12301923012390 #unix
# data: [1, 2, 3, 4, ..., 100900]
#
# # Inside video.json
# session_id: 123 #in case we have multiple data sessions with the same user
# user_id: 478
# tags: ["pilot", "video", "resource"] # resource tells us to expect a "url" key
# name: "SC"
# description: "Screencapture of user screen"
# ...
# timestamp: 12301923012390 #unix
# url: "adjadllkf.mp4"
#
# ```
# ## Create and Save json Files
# +
import json
def save_jsonfile(name, data):
#file = name + ".json" #name is a string
with open(name, 'w') as outfile:
json.dump(data, outfile)
print("File saved!", name)
# -
# ## Bio_Data
# .csv files in this archive are in the following format:
# The first row is the initial time of the session expressed as unix timestamp in UTC.
# The second row is the sample rate expressed in Hz.
#
# ### EDA.csv
# Data from the electrodermal activity sensor expressed as microsiemens (μS).
#
# ### HR.csv
# Average heart rate extracted from the BVP signal.The first row is the initial time of the session expressed as unix timestamp in UTC.
# The second row is the sample rate expressed in Hz.
#
# ### Format of csv files
# userID_sessionID_type.csv
# e.g: 25_0_HR.csv
# ## TODO:
#
# * Need to generalize the following code to create specific folder under user folder
# * code should automatically generate sessionID, userID, and name
# +
BIO_ROOT = "data/biosignals/"
USER_ROOT = "data/users/"
import os
import pandas as pd
import numpy as np
import csv
# create .json file from csv files
def create_bio_json(bioName, sessionID, userID, name, sampFreq, timestamp, data):
unit = ""
description = ""
if bioName == "HR":
unit = "bpm"
description = "Heart rate"
if bioName == "EDA":
unit = "microsiemens"
description = "Electrodermal data"
tags = ["bio", "time-series"]
data = {"sessionID": sessionID, \
"userID": userID, \
"tags": tags, \
"name": name, \
"description": description, \
"sampling_frequency": sampFreq, \
"timestamp": timestamp, \
"unit": unit, \
"data": data \
}
filePath = USER_ROOT + bioName + ".json"
# print(filePath)
save_jsonfile(filePath, data)
# save the csv file into a .json file of the same name
def grab_and_save(filename):
data = []
file = BIO_ROOT + filename
name = filename[:-4]
#print(file)
with open(file) as csvfile:
csvreader = csv.reader(csvfile)
timestamp = next(csvreader)[0]
#print(timestamp)
sampling_rate = next(csvreader)[0]
#print(sampling_rate)
for row in csvreader:
data.append(row[0])
# TODO: modify parameters
# sessionID, userID, tags, name, description, sampFreq, data
create_bio_json(name, 0, 25, "cesar",\
sampling_rate, timestamp, data)
#create json file and save data into json file
# Iterate through the csv files in BIO_ROOT and generate json objects
directory = os.fsencode(BIO_ROOT)
for file in os.listdir(directory):
filename = os.fsdecode(file)
if filename.endswith(".csv"):
#print(filename[:-4])
grab_and_save(filename)
# -
# ## Video Data
#
# Video data are uploaded in data/videos. Coded csv files live in data/videos/coded. We will be aligning the video with biosignals using unix timestamp.
# +
VIDEO_ROOT = "data/videos/"
import datetime, platform
# get the timestamp of the video
def get_video_timestamp(userID):
uid = str(userID)
raw_video = VIDEO_ROOT + uid + "H" + ".MOV"
if platform.system() == "Darwin":
posix_time = os.stat(raw_video).st_birthtime
# t = datetime.datetime.fromtimestamp(posix_time).strftime(
# '%Y-%m-%dT%H:%M:%SZ')
# print(t)
# print(posix_time)
# session = os.path.basename(raw_video).split('.')[0]
# url = "/" + raw_video
# print(url)
return posix_time + 77 #just for this time. Adjust for delayed save of screencast
# For this specific subject run
get_video_timestamp(25)
# -
# ## Biosignal + Video Data Analysis
# +
CODED_ROOT = "data/videos/coded/"
from pprint import pprint
# calculates the difference in terms of unix timestamp (seconds) between the video and the biosignals.
# If difference is positive, then video is taken after the biosignal
# If difference is negative, then video is taken before the biosignal
def timestamp_difference(bioname, userID):
videoT = get_video_timestamp(userID)
print(videoT)
filepath = USER_ROOT + bioname + ".json"
with open(filepath) as f:
data = json.load(f)
bioT = float(data["timestamp"])
difference = int(videoT) - int(bioT)
return difference
# print(timestamp_difference("EDA", 25))
# print(timestamp_difference("HR", 25))
# A more general function of the above
def time_difference(videoT, bioname, userID):
filepath = USER_ROOT + bioname + ".json"
with open(filepath) as f:
data = json.load(f)
bioT = float(data["timestamp"])
difference = int(videoT) - int(bioT)
return difference
# Get the sampling frequency of this particular biodata
def get_frequency(bioname):
filepath = USER_ROOT + bioname + ".json"
with open(filepath) as f:
data = json.load(f)
frequency = int(float(data["sampling_frequency"]))
return frequency
# Now calculate how much to index into the data array of biosignal json object to get matching data
def get_beginning_index(bioname, userID):
difference = timestamp_difference(bioname, userID)
frequency = get_frequency(bioname)
return difference * frequency
# A more general function of the above
def get_adjusted_index(videoT, bioname, userID):
difference = time_difference(videoT, bioname, userID)
frequency = get_frequency(bioname)
#print(frequency)
#print(difference)
return difference * frequency
# add spliced data to the biosignal json object
def add_spliced_data_to_json(bioname, startIndex):
filepath = USER_ROOT + bioname + ".json"
with open(filepath) as f:
data = json.load(f)
data["spliced_data"] = data["data"][startIndex:]
# rewrite this dictionary back to json file
save_jsonfile(filepath, data)
# Converts "HH:mm:ss.S" to number of seconds
def elapsed_to_seconds(elapsed):
hour = 0
minute = 0
second = 0
result = 0
if len(elapsed) > 7:
hour = int(elapsed[:2])
minute = int(elapsed[3:5])
second = int(elapsed[6:8])
result = hour * 60 * 60 + minute * 60 + second
else:
minute = int(elapsed[:2])
second = int(elapsed[3:5])
result = minute * 60 + second
return result
# Calculate average biosignal during stage
def average(bioname, stage, userID):
file = CODED_ROOT + "25-" + stage + ".csv"
bio_filepath = USER_ROOT + bioname + ".json"
videoT = get_video_timestamp(userID)
begin = 0
end = 0
spliced_data = []
with open(file) as csvfile:
csvreader = csv.reader(csvfile)
next(csvreader) # pass the headings
for times in csvreader:
begin = times[2]
end = times[3]
begin = elapsed_to_seconds(begin)
end = elapsed_to_seconds(end)
vT_begin = videoT + begin
vT_end = videoT + end
index_begin = get_adjusted_index(vT_begin, bioname, userID)
index_end = get_adjusted_index(vT_end, bioname, userID)
with open(bio_filepath) as bf:
data = json.load(bf)
for item in data["data"][index_begin:index_end]: # possibly need to add 1? Figure out
# need to convert spliced data into a list of ints instead of strings
spliced_data.append(item)
spliced_data = list(map(float, spliced_data))
count = len(spliced_data)
result = sum(spliced_data)/count
return result
# testing for heart rate of getting started
HR_GETSTART = average("HR", "Getting Started", 25)
EDA_GETSTART = average("EDA", "Getting Started", 25)
HR_SUCCESS = average("HR", "Success", 25)
EDA_SUCCESS = average("EDA", "Success", 25)
HR_ENCOUNTERDIFF = average("HR", "Encountering Difficulties", 25)
EDA_ENCOUNTERDIFF = average("EDA", "Encountering Difficulties", 25)
HR_DEALDIFF = average("HR", "Dealing with Difficulties", 25)
EDA_DEALDIFF = average("EDA", "Dealing with Difficulties", 25)
print("Average HR for Getting Started:")
print(HR_GETSTART)
print("Average EDA for Getting Started:")
print(EDA_GETSTART)
print("Average HR for Success:")
print(HR_SUCCESS)
print("Average EDA for Success:")
print(EDA_SUCCESS)
print("Average HR for Encountering Difficulties:")
print(HR_ENCOUNTERDIFF)
print("Average EDA for Encountering Difficulties:")
print(EDA_ENCOUNTERDIFF)
print("Average HR for Dealing with Difficulties:")
print(HR_DEALDIFF)
print("Average EDA for Dealing with Difficulties:")
print(EDA_DEALDIFF)
# -
# ## Finer Biosignal Analysis
# We generate "cropped" biosignal csv files for each biosignal at each stage. There should be 8 biosignal files as of now generated.
#
#
# Naming convention:
#
# GS - Getting Started
#
# S - Success
#
# ED - Encountering Difficulties
#
# DD - Dealing with Difficulties
#
# e.g: ED_HR.csv, ED_EDA.csv
#
# The csv files will have a single column of all the biosignal measures during the specific stage.
# +
CROPPED_ROOT = "data/cropped/"
def generate_csv_at_cropped(filename, mylist):
name = CROPPED_ROOT + filename + ".csv"
with open(name, 'w') as csvfile:
wr = csv.writer(csvfile)
for v in mylist:
wr.writerow([v])
def shorten(stage):
if stage == "Getting Started":
return "GS"
elif stage == "Success":
return "S"
elif stage == "Encountering Difficulties":
return "ED"
elif stage == "Dealing with Difficulties":
return "DD"
else:
print("No match for this stage.")
def splice(bioname, stage, userID):
file = CODED_ROOT + "25-" + stage + ".csv"
bio_filepath = USER_ROOT + bioname + ".json"
videoT = get_video_timestamp(userID)
begin = 0
end = 0
spliced_data = []
with open(file) as csvfile:
csvreader = csv.reader(csvfile)
next(csvreader) # pass the headings
for times in csvreader:
begin = times[2]
end = times[3]
begin = elapsed_to_seconds(begin)
end = elapsed_to_seconds(end)
vT_begin = videoT + begin
vT_end = videoT + end
index_begin = get_adjusted_index(vT_begin, bioname, userID)
index_end = get_adjusted_index(vT_end, bioname, userID)
with open(bio_filepath) as bf:
data = json.load(bf)
for item in data["data"][index_begin:index_end]: # possibly need to add 1? Figure out
# need to convert spliced data into a list of ints instead of strings
spliced_data.append(item)
spliced_data = list(map(float, spliced_data))
#print(spliced_data)
shortened = shorten(stage)
filename = shortened + "_" + bioname
generate_csv_at_cropped(filename, spliced_data)
splice("HR", "Encountering Difficulties", 25)
splice("EDA", "Encountering Difficulties", 25)
splice("HR", "Getting Started", 25)
splice("EDA", "Getting Started", 25)
splice("HR", "Success", 25)
splice("EDA", "Success", 25)
splice("HR", "Dealing with Difficulties", 25)
splice("EDA", "Dealing with Difficulties", 25)
# generate_csv_at_cropped("dog")
# -
# ## Experiment with Tau Value
# +
TAU = 18
# Calculate average biosignal during stage, but delayed by TAU seconds
def average_shifted(bioname, stage, userID):
file = CODED_ROOT + "25-" + stage + ".csv"
bio_filepath = USER_ROOT + bioname + ".json"
videoT = get_video_timestamp(userID)
begin = 0
end = 0
spliced_data = []
with open(file) as csvfile:
csvreader = csv.reader(csvfile)
next(csvreader) # pass the headings
for times in csvreader:
begin = times[2]
end = times[3]
begin = elapsed_to_seconds(begin) + TAU
end = elapsed_to_seconds(end) + TAU
vT_begin = videoT + begin
vT_end = videoT + end
index_begin = get_adjusted_index(vT_begin, bioname, userID)
index_end = get_adjusted_index(vT_end, bioname, userID)
with open(bio_filepath) as bf:
data = json.load(bf)
for item in data["data"][index_begin:index_end]: # possibly need to add 1? Figure out
# need to convert spliced data into a list of ints instead of strings
spliced_data.append(item)
spliced_data = list(map(float, spliced_data))
count = len(spliced_data)
result = sum(spliced_data)/count
return result
HR_GETSTART_S = average_shifted("HR", "Getting Started", 25)
EDA_GETSTART_S = average_shifted("EDA", "Getting Started", 25)
HR_SUCCESS_S = average_shifted("HR", "Success", 25)
EDA_SUCCESS_S = average_shifted("EDA", "Success", 25)
HR_ENCOUNTERDIFF_S = average_shifted("HR", "Encountering Difficulties", 25)
EDA_ENCOUNTERDIFF_S = average_shifted("EDA", "Encountering Difficulties", 25)
HR_DEALDIFF_S = average_shifted("HR", "Dealing with Difficulties", 25)
EDA_DEALDIFF_S = average_shifted("EDA", "Dealing with Difficulties", 25)
print("Average HR for Getting Started:")
print(HR_GETSTART_S)
print("Average EDA for Getting Started:")
print(EDA_GETSTART_S)
print("Average HR for Success:")
print(HR_SUCCESS_S)
print("Average EDA for Success:")
print(EDA_SUCCESS_S)
print("Average HR for Encountering Difficulties:")
print(HR_ENCOUNTERDIFF_S)
print("Average EDA for Encountering Difficulties:")
print(EDA_ENCOUNTERDIFF_S)
print("Average HR for Dealing with Difficulties:")
print(HR_DEALDIFF_S)
print("Average EDA for Dealing with Difficulties:")
print(EDA_DEALDIFF_S)
# -
# ## Tau value experiment 2
#
# +
TAU_ROOT = "data/TAU/"
def generate_csv_at_TAU(filename, mylist):
name = TAU_ROOT + filename + ".csv"
with open(name, 'w') as csvfile:
wr = csv.writer(csvfile)
for v in mylist:
wr.writerow([v])
def splice_TAU(bioname, stage, userID):
file = CODED_ROOT + "25-" + stage + ".csv"
bio_filepath = USER_ROOT + bioname + ".json"
videoT = get_video_timestamp(userID)
begin = 0
end = 0
index = 0
spliced_data = []
with open(file) as csvfile:
csvreader = csv.reader(csvfile)
next(csvreader) # pass the headings
for times in csvreader:
begin = times[2]
end = times[3]
begin = elapsed_to_seconds(begin)
end = elapsed_to_seconds(end)
vT_begin = videoT + begin - 5 # Want to see the previous 5 seconds of the biosignal prior to stimulus
vT_end = videoT + end
index_begin = get_adjusted_index(vT_begin, bioname, userID)
index_end = get_adjusted_index(vT_end, bioname, userID)
with open(bio_filepath) as bf:
data = json.load(bf)
for item in data["data"][index_begin:index_end]: # possibly need to add 1? Figure out
# need to convert spliced data into a list of ints instead of strings
spliced_data.append(item)
# for every occurence of encountering difficulties, make a csv file of biosignals around that timestamp
spliced_data = list(map(float, spliced_data))
shortened = shorten(stage)
filename = shortened + "_" + bioname + "_" + str(index)
generate_csv_at_TAU(filename, spliced_data)
# increment index, restore spliced_data array
index += 1
spliced_data = []
splice_TAU("EDA","Encountering Difficulties", 25 )
splice_TAU("HR","Encountering Difficulties", 25 )
# -
| codewords/BiosignalAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Deep Learning Bootcamp November 2017, GPU Computing for Data Scientists
#
# <img src="../images/bcamp.png" align="center">
#
# ## 31-PyTorch-using-CONV1D-on-one-dimensional-data
#
# Web: https://www.meetup.com/Tel-Aviv-Deep-Learning-Bootcamp/events/241762893/
#
# Notebooks: <a href="https://github.com/QuantScientist/Data-Science-PyCUDA-GPU"> On GitHub</a>
#
# *<NAME>*
#
# <img src="../images/pt.jpg" width="35%" align="center">
#
#
# ### Data
# - Download from https://numer.ai/leaderboard
#
# <img src="../images/Numerai.png" width="35%" align="center">
#
# + [markdown] slideshow={"slide_type": "slide"}
# # PyTorch Imports
#
# + slideshow={"slide_type": "-"}
# # !pip install pycuda
# %reset -f
# # %%timeit
import torch
from torch.autograd import Variable
import numpy as np
import pandas
import numpy as np
import pandas as pd
from sklearn import cross_validation
from sklearn import metrics
from sklearn.metrics import roc_auc_score, log_loss, roc_auc_score, roc_curve, auc
import matplotlib.pyplot as plt
from sklearn import cross_validation
from sklearn import metrics
from sklearn.metrics import roc_auc_score, log_loss, roc_auc_score, roc_curve, auc
from sklearn.cross_validation import StratifiedKFold, ShuffleSplit, cross_val_score, train_test_split
import logging
import numpy
import numpy as np
from __future__ import print_function
from __future__ import division
import math
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
import os
import torch
from torch.utils.data.dataset import Dataset
from torch.utils.data import DataLoader
from torchvision import transforms
from torch import nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
from sklearn.preprocessing import MultiLabelBinarizer
import time
from sklearn.preprocessing import PolynomialFeatures
import pandas as pd
import numpy as np
import scipy
# %matplotlib inline
from pylab import rcParams
rcParams['figure.figsize'] = (6, 6) # setting default size of plots
import tensorflow as tf
print("tensorflow:" + tf.__version__)
# !set "KERAS_BACKEND=tensorflow"
import torch
import sys
print('__Python VERSION:', sys.version)
print('__pyTorch VERSION:', torch.__version__)
print('__CUDA VERSION')
from subprocess import call
print('__CUDNN VERSION:', torch.backends.cudnn.version())
print('__Number CUDA Devices:', torch.cuda.device_count())
print('__Devices')
# # !pip install http://download.pytorch.org/whl/cu75/torch-0.2.0.post1-cp27-cp27mu-manylinux1_x86_64.whl
# # !pip install torchvision
# # ! pip install cv2
# import cv2
print("OS: ", sys.platform)
print("Python: ", sys.version)
print("PyTorch: ", torch.__version__)
print("Numpy: ", np.__version__)
handler=logging.basicConfig(level=logging.INFO)
lgr = logging.getLogger(__name__)
# %matplotlib inline
# # !pip install psutil
import psutil
def cpuStats():
print(sys.version)
print(psutil.cpu_percent())
print(psutil.virtual_memory()) # physical memory usage
pid = os.getpid()
py = psutil.Process(pid)
memoryUse = py.memory_info()[0] / 2. ** 30 # memory use in GB...I think
print('memory GB:', memoryUse)
cpuStats()
# # %%timeit
use_cuda = torch.cuda.is_available()
# use_cuda = False
FloatTensor = torch.cuda.FloatTensor if use_cuda else torch.FloatTensor
LongTensor = torch.cuda.LongTensor if use_cuda else torch.LongTensor
Tensor = FloatTensor
lgr.info("USE CUDA=" + str (use_cuda))
# + [markdown] slideshow={"slide_type": "slide"}
# # Global params
# +
# NN params
LR = 0.005
MOMENTUM= 0.9
# Data params
TARGET_VAR= 'target'
TOURNAMENT_DATA_CSV = 'numerai_tournament_data.csv'
TRAINING_DATA_CSV = 'numerai_training_data.csv'
BASE_FOLDER = 'numerai/'
# fix seed
seed=17*19
np.random.seed(seed)
torch.manual_seed(seed)
if use_cuda:
torch.cuda.manual_seed(seed)
# + [markdown] slideshow={"slide_type": "slide"}
# # Train / Validation / Test Split
# - Numerai provides a data set that is allready split into train, validation and test sets.
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.pipeline import Pipeline
from collections import defaultdict
# Train, Validation, Test Split
def loadDataSplit():
df_train = pd.read_csv(BASE_FOLDER + TRAINING_DATA_CSV)
# TOURNAMENT_DATA_CSV has both validation and test data provided by NumerAI
df_test_valid = pd.read_csv(BASE_FOLDER + TOURNAMENT_DATA_CSV)
answers_1_SINGLE = df_train[TARGET_VAR]
df_train.drop(TARGET_VAR, axis=1,inplace=True)
df_train.drop('id', axis=1,inplace=True)
df_train.drop('era', axis=1,inplace=True)
df_train.drop('data_type', axis=1,inplace=True)
df_train.to_csv(BASE_FOLDER + TRAINING_DATA_CSV + 'clean.csv', header=False, index = False)
df_train= pd.read_csv(BASE_FOLDER + TRAINING_DATA_CSV + 'clean.csv', header=None, dtype=np.float32)
df_train = pd.concat([df_train, answers_1_SINGLE], axis=1)
feature_cols = list(df_train.columns[:-1])
target_col = df_train.columns[-1]
trainX, trainY = df_train[feature_cols], df_train[target_col]
df_validation_set=df_test_valid.loc[df_test_valid['data_type'] == 'validation']
df_validation_set=df_validation_set.copy(deep=True)
answers_1_SINGLE_validation = df_validation_set[TARGET_VAR]
df_validation_set.drop(TARGET_VAR, axis=1,inplace=True)
df_validation_set.drop('id', axis=1,inplace=True)
df_validation_set.drop('era', axis=1,inplace=True)
df_validation_set.drop('data_type', axis=1,inplace=True)
df_validation_set.to_csv(BASE_FOLDER + TRAINING_DATA_CSV + '-validation-clean.csv', header=False, index = False)
df_validation_set= pd.read_csv(BASE_FOLDER + TRAINING_DATA_CSV + '-validation-clean.csv', header=None, dtype=np.float32)
df_validation_set = pd.concat([df_validation_set, answers_1_SINGLE_validation], axis=1)
feature_cols = list(df_validation_set.columns[:-1])
target_col = df_validation_set.columns[-1]
valX, valY = df_validation_set[feature_cols], df_validation_set[target_col]
df_test_set = pd.read_csv(BASE_FOLDER + TOURNAMENT_DATA_CSV)
df_test_set=df_test_set.copy(deep=True)
df_test_set.drop(TARGET_VAR, axis=1,inplace=True)
tid_1_SINGLE = df_test_set['id']
df_test_set.drop('id', axis=1,inplace=True)
df_test_set.drop('era', axis=1,inplace=True)
df_test_set.drop('data_type', axis=1,inplace=True)
feature_cols = list(df_test_set.columns) # must be run here, we dont want the ID
df_test_set = pd.concat([tid_1_SINGLE, df_test_set], axis=1)
testX = df_test_set[feature_cols].values
return trainX, trainY, valX, valY, testX, df_test_set
# +
trainX, trainY, valX, valY, testX, df_test_set = loadDataSplit()
print (trainX.shape)
print (trainY.shape)
print (valX.shape)
print (valY.shape)
print (testX.shape)
print (df_test_set.shape)
# +
# Convert the np arrays into the correct dimention and type
# Note that BCEloss requires Float in X as well as in y
def XnumpyToTensor(x_data_np):
x_data_np = np.array(x_data_np.values, dtype=np.float32)
print(x_data_np.shape)
print(type(x_data_np))
if use_cuda:
lgr.info ("Using the GPU")
X_tensor = Variable(torch.from_numpy(x_data_np).cuda()) # Note the conversion for pytorch
else:
lgr.info ("Using the CPU")
X_tensor = Variable(torch.from_numpy(x_data_np)) # Note the conversion for pytorch
print(type(X_tensor.data)) # should be 'torch.cuda.FloatTensor'
print((X_tensor.data.shape)) # torch.Size([108405, 29])
return X_tensor
# Convert the np arrays into the correct dimention and type
# Note that BCEloss requires Float in X as well as in y
def YnumpyToTensor(y_data_np):
y_data_np=y_data_np.reshape((y_data_np.shape[0],1)) # Must be reshaped for PyTorch!
print(y_data_np.shape)
print(type(y_data_np))
if use_cuda:
lgr.info ("Using the GPU")
# Y = Variable(torch.from_numpy(y_data_np).type(torch.LongTensor).cuda())
Y_tensor = Variable(torch.from_numpy(y_data_np)).type(torch.FloatTensor).cuda() # BCEloss requires Float
else:
lgr.info ("Using the CPU")
# Y = Variable(torch.squeeze (torch.from_numpy(y_data_np).type(torch.LongTensor))) #
Y_tensor = Variable(torch.from_numpy(y_data_np)).type(torch.FloatTensor) # BCEloss requires Float
print(type(Y_tensor.data)) # should be 'torch.cuda.FloatTensor'
print(y_data_np.shape)
print(type(y_data_np))
return Y_tensor
# -
# # CNN Architecture
#
# +
# def myModule(nn.Module):
# def __init__(self):
# # Init stuff here
# self.X = nn.Sequential(
# nn.Linear(num_input_genes, num_tfs),
# nn.ReLU(),
# nn.BatchNorm1d(num_tfs)
# )
# self.C = nn.Sequential(
# nn.Conv1d(num_tfs, num_conv_out_channels, conv_kernel_size),
# nn.ReLU(),
# nn.BatchNorm1d(num_conv_out_channels),
# nn.MaxPool1d(max_pool_kernel_size)
# )
# def forward(self, input, M):
# x_out = self.X(input)
# x_out = M * x_out # With required reshaping, ...
# x_out = self.C(x_out)
# return x_out
# +
# References:
# https://github.com/vinhkhuc/PyTorch-Mini-Tutorials/blob/master/5_convolutional_net.py
# https://gist.github.com/spro/c87cc706625b8a54e604fb1024106556
# Arguments should (by docs) be nn.Conv1d(#input channels, #output channels, kernel size)
X_tensor_train= XnumpyToTensor(trainX) # default order is NBC for a 3d tensor, but we have a 2d tensor
X_shape=X_tensor_train.data.size()
# Dimensions
# Number of features for the input layer
N_FEATURES=trainX.shape[1]
# Number of rows
NUM_ROWS_TRAINNING=trainX.shape[0]
# this number has no meaning except for being divisable by 2
MULT_FACTOR=8
# Size of first linear layer
Layer1Size=N_FEATURES * MULT_FACTOR
# CNN kernel size
CNN_KERNEL=7
MAX_POOL_KERNEL=4
LAST_OP_SIZE=(int(MULT_FACTOR/MAX_POOL_KERNEL))
class Net2(nn.Module):
def __init__(self, n_feature, n_hidden, n_output):
super(Net2, self).__init__()
self.n_feature=n_feature
self.l1 = nn.Sequential(
torch.nn.Linear(n_feature, n_hidden),
torch.nn.Dropout(p=1 -.95),
torch.nn.LeakyReLU (0.1),
torch.nn.BatchNorm1d(n_hidden, eps=1e-05, momentum=0.1, affine=True)
)
self.c1= nn.Sequential(
torch.nn.Conv1d(n_feature, n_feature * MULT_FACTOR, kernel_size=(CNN_KERNEL,), stride=(1,), padding=(1,)),
torch.nn.Dropout(p=1 -.75),
torch.nn.LeakyReLU (0.1),
torch.nn.BatchNorm1d(n_hidden, eps=1e-05, momentum=0.1, affine=True),
# torch.nn.MaxPool1d (MAX_POOL_KERNEL,MAX_POOL_KERNEL) # (x.size() after conv1d:torch.Size([108405, 84, 1])
# after conv1d:torch.Size([108405, 84, 4])
)
self.out = nn.Sequential(
torch.nn.Linear(MULT_FACTOR * N_FEATURES * (MULT_FACTOR - CNN_KERNEL + 3) , n_output),
)
self.sig=nn.Sigmoid()
def forward(self, x):
x=self.l1(x)
# print ('(x.size() after l1:' + str (x.size()))
# for CNN
x = x.view(x.shape[0],self.n_feature,MULT_FACTOR)
# print ('(x.size() after re-shape:' + str (x.size()))
x=self.c1(x)
# print ('(x.size() after conv1d:' + str (x.size()))
# for Linear layer
x = x.view(x.shape[0], self.n_feature * MULT_FACTOR * (MULT_FACTOR - CNN_KERNEL + 3))
# print ('(x.size() after re-shape 2:' + str (x.size()))
x=self.out(x)
# print ('(x.size() after l2:' + str (x.size()))
x=self.sig(x)
return x
net = Net2(n_feature=N_FEATURES, n_hidden=Layer1Size, n_output=1) # define the network
if use_cuda:
net=net.cuda() # very important !!!
lgr.info(net)
b = net(X_tensor_train)
print ('(b.size():' + str (b.size())) # torch.Size([108405, 928])
# +
optimizer = torch.optim.Adam(net.parameters(), lr=LR,weight_decay=5e-4) # L2 regularization
loss_func=torch.nn.BCELoss() # Binary cross entropy: http://pytorch.org/docs/nn.html#bceloss
if use_cuda:
lgr.info ("Using the GPU")
net.cuda()
loss_func.cuda()
lgr.info (optimizer)
lgr.info (loss_func)
# +
import time
start_time = time.time()
epochs=200
all_losses = []
X_tensor_train= XnumpyToTensor(trainX)
Y_tensor_train= YnumpyToTensor(trainY)
print(type(X_tensor_train.data), type(Y_tensor_train.data)) # should be 'torch.cuda.FloatTensor'
# From here onwards, we must only use PyTorch Tensors
for step in range(epochs):
out = net(X_tensor_train) # input x and predict based on x
cost = loss_func(out, Y_tensor_train) # must be (1. nn output, 2. target), the target label is NOT one-hotted
optimizer.zero_grad() # clear gradients for next train
cost.backward() # backpropagation, compute gradients
optimizer.step() # apply gradients
if step % 50 == 0:
loss = cost.data[0]
all_losses.append(loss)
print(step, cost.data.cpu().numpy())
prediction = (net(X_tensor_train).data).float() # probabilities
pred_y = prediction.cpu().numpy().squeeze()
target_y = Y_tensor_train.cpu().data.numpy()
tu = ((pred_y == target_y).mean(),log_loss(target_y, pred_y),roc_auc_score(target_y,pred_y ))
print ('ACC={}, LOG_LOSS={}, ROC_AUC={} '.format(*tu))
end_time = time.time()
print ('{} {:6.3f} seconds'.format('GPU:', end_time-start_time))
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(all_losses)
plt.show()
false_positive_rate, true_positive_rate, thresholds = roc_curve(target_y,pred_y)
roc_auc = auc(false_positive_rate, true_positive_rate)
plt.title('LOG_LOSS=' + str(log_loss(target_y, pred_y)))
plt.plot(false_positive_rate, true_positive_rate, 'b', label='AUC = %0.6f' % roc_auc)
plt.legend(loc='lower right')
plt.plot([0, 1], [0, 1], 'r--')
plt.xlim([-0.1, 1.2])
plt.ylim([-0.1, 1.2])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# -
# # Validation ROC_AUC
# +
net.eval()
# Validation data
print (valX.shape)
print (valY.shape)
X_tensor_val= XnumpyToTensor(valX)
Y_tensor_val= YnumpyToTensor(valY)
print(type(X_tensor_val.data), type(Y_tensor_val.data)) # should be 'torch.cuda.FloatTensor'
predicted_val = (net(X_tensor_val).data).float() # probabilities
# predicted_val = (net(X_tensor_val).data > 0.5).float() # zero or one
pred_y = predicted_val.cpu().numpy()
target_y = Y_tensor_val.cpu().data.numpy()
print (type(pred_y))
print (type(target_y))
tu = (str ((pred_y == target_y).mean()),log_loss(target_y, pred_y),roc_auc_score(target_y,pred_y ))
print ('\n')
print ('acc={} log_loss={} roc_auc={} '.format(*tu))
false_positive_rate, true_positive_rate, thresholds = roc_curve(target_y,pred_y)
roc_auc = auc(false_positive_rate, true_positive_rate)
plt.title('LOG_LOSS=' + str(log_loss(target_y, pred_y)))
plt.plot(false_positive_rate, true_positive_rate, 'b', label='AUC = %0.6f' % roc_auc)
plt.legend(loc='lower right')
plt.plot([0, 1], [0, 1], 'r--')
plt.xlim([-0.1, 1.2])
plt.ylim([-0.1, 1.2])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# print (pred_y)
# -
# # Submision
# +
print (df_test_set.shape)
columns = ['id', 'probability']
df_pred=pd.DataFrame(data=np.zeros((0,len(columns))), columns=columns)
df_pred.id.astype(int)
for index, row in df_test_set.iterrows():
rwo_no_id=row.drop('id')
# print (rwo_no_id.values)
x_data_np = np.array(rwo_no_id.values, dtype=np.float32)
if use_cuda:
X_tensor_test = Variable(torch.from_numpy(x_data_np).cuda()) # Note the conversion for pytorch
else:
X_tensor_test = Variable(torch.from_numpy(x_data_np)) # Note the conversion for pytorch
X_tensor_test=X_tensor_test.view(1, trainX.shape[1]) # does not work with 1d tensors
predicted_val = (net(X_tensor_test).data).float() # probabilities
p_test = predicted_val.cpu().numpy().item() # otherwise we get an array, we need a single float
df_pred = df_pred.append({'id':row['id'].astype(int), 'probability':p_test},ignore_index=True)
df_pred.head(5)
# +
df_pred.id=df_pred.id.astype(int)
def savePred(df_pred, loss):
# csv_path = 'pred/p_{}_{}_{}.csv'.format(loss, name, (str(time.time())))
csv_path = 'pred/pred_{}_{}.csv'.format(loss, (str(time.time())))
df_pred.to_csv(csv_path, columns=('id', 'probability'), index=None)
print (csv_path)
savePred (df_pred, log_loss(target_y, pred_y))
# -
| day 02 PyTORCH and PyCUDA/PyTorch/31-PyTorch-using-CONV1D-on-one-dimensional-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
fb = pd.DataFrame.from_csv('../data/facebook.csv')
ms = pd.DataFrame.from_csv('../data/microsoft.csv')
# ## Create a new column in the DataFrame (1) - Price difference
#Create a new column PriceDiff in the DataFrame fb
fb['PriceDiff'] = fb['Close'].shift(-1) - fb['Close']
#Your turn to create PriceDiff in the DataFrame ms
ms['PriceDiff'] = ms['Close'].shift(-1) - ms['Close']
#Run this code to display the price difference of Microsoft on 2015-01-05
print(ms['PriceDiff'].loc['2015-01-05'])
# ** Expected Output: ** -0.68
# ## Create a new column in the DataFrame (2) - Daily return
#
# Daily Return is calcuated as PriceDiff/Close
#Create a new column Return in the DataFrame fb
fb['Return'] = fb['PriceDiff'] /fb['Close']
#Your turn to create a new column Return in the DataFrame MS
ms['Return'] = ms['PriceDiff'] /ms['Close']
#Run this code to print the return on 2015-01-05
print(ms['Return'].loc['2015-01-05'])
# ** Expected Output: ** -0.0146773142811
# ## Create a new column in the DataFrame using List Comprehension - Direction
# +
#Create a new column Direction.
#The List Comprehension means : if the price difference is larger than 0, denote as 1, otherwise, denote as 0,
#for every record in the DataFrame - fb
fb['Direction'] = [1 if fb['PriceDiff'].loc[ei] > 0 else 0 for ei in fb.index ]
# +
# Your turn to create a new column Direction for MS
ms['Direction'] = [1 if ms['PriceDiff'].loc[ei] > 0 else 0 for ei in ms.index ]
# -
# Run the following code to show the price difference on 2015-01-05
print('Price difference on {} is {}. direction is {}'.format('2015-01-05', ms['PriceDiff'].loc['2015-01-05'], ms['Direction'].loc['2015-01-05']))
# ** Expected Output: ** Price difference on 2015-01-05 is -0.6799999999999997. direction is 0
# ## Create a new column in the DataFrame using Rolling Window calculation (.rolling()) - Moving average
# +
fb['ma50'] = fb['Close'].rolling(50).mean()
#plot the moving average
plt.figure(figsize=(10, 8))
fb['ma50'].loc['2015-01-01':'2015-12-31'].plot(label='MA50')
fb['Close'].loc['2015-01-01':'2015-12-31'].plot(label='Close')
plt.legend()
plt.show()
# +
# You can use .rolling() to calculate any numbers of days' Moving Average. This is your turn to calculate "60 days"
# moving average of Microsoft, rename it as "ma60". And follow the codes above in plotting a graph
ms['ma60'] = ms['Close'].rolling(60).mean()
#plot the moving average
plt.figure(figsize=(10, 8))
ms['ma60'].loc['2015-01-01':'2015-12-31'].plot(label='MA60')
ms['Close'].loc['2015-01-01':'2015-12-31'].plot(label='Close')
plt.legend()
plt.show()
# -
# ** Expected Output: **
# <img src="MA.png">
| HKUST - Python and Statistics for Financial Analysis/module1- Visualizing and Munging Stock Data/Create+new+features+and+columns+in+DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jdvelasq/talleres-presenciales/blob/main/AP_sesion_4_2021_10_30_ejemplo_completo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="rwYUPknQxhoc"
from sklearn.datasets import make_moons
X_full, y_full = make_moons(
n_samples=100,
shuffle=False,
noise=None,
random_state=12345,
)
# + id="CK9Vx8b6x4ur"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X_full,
y_full,
test_size=15,
random_state=12345,
)
# + colab={"base_uri": "https://localhost:8080/"} id="0_vpxfAUx--S" outputId="2e4ea75c-82e1-4614-a9bc-20bfd5faf84f"
from sklearn.preprocessing import MinMaxScaler
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
estimators = [
("minMaxScaler", MinMaxScaler()),
("clf", SVC()),
]
pipeline = Pipeline(
steps=estimators,
verbose=False,
)
pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="foyYVsF2yslY" outputId="28bfef40-076d-4588-f511-23d9de8286ed"
from sklearn.model_selection import GridSearchCV
param_grid = {
"clf__kernel": ["rbf"],
"clf__gamma": [1e-3, 1e-4],
"clf__C": [1, 10, 100, 1000],
}
gridSearchCV = GridSearchCV(
estimator=pipeline,
param_grid=param_grid,
cv=5,
scoring="accuracy",
refit=True,
return_train_score=False,
)
gridSearchCV.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="kk4h9PNizqHx" outputId="e79ab793-ea5d-430b-9295-aa66c6a314b0"
gridSearchCV.best_estimator_
# + colab={"base_uri": "https://localhost:8080/"} id="REFFWtYszsrE" outputId="d9acfd8c-c39a-4680-e2f8-eb08a05d4ad5"
gridSearchCV.predict(X_train)
# + id="gaILOHE2z862"
import pickle
s = pickle.dumps(gridSearchCV)
with open('modelo.pkl', 'wb') as file:
file.write(s)
# + colab={"base_uri": "https://localhost:8080/"} id="DCGyr3ce0Pao" outputId="e1a9786b-4981-40bc-bfd9-1cc9d0f109ca"
import pickle
with open('modelo.pkl', 'rb') as file:
modelo = pickle.load(file)
modelo.predict(X_test) == y_test
# + id="xSw-AD5E0iWH"
| AP_sesion_4_2021_10_30_ejemplo_completo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Using Fugue on Databricks
#
# A lot of Spark users use the `databricks-connect` library to execute Spark commands on a Databricks cluster instead of a local session. `databricks-connect` replaces the local installation of `pyspark` and makes `pyspark` code get executed on the cluster, allowing users to use the cluster directly from their local machine.
#
# In this tutorial, we will go through the following steps:
#
# 1. Setting up a Databricks cluster
# 2. Installing and configuring `databricks-connect`
# 3. Using Fugue with a Spark backend
#
# ## Setup Workspace
#
# Databricks is available across all three major cloud providers (AWS, Azure, GCP). In general, it will involve going to the Marketplace of your cloud vendor, and then signing up for Databricks.
#
# For more information, you can look at the following cloud-specific documentation.
# * https://databricks.com/product/aws
# * https://databricks.com/product/azure
# * https://databricks.com/product/google-cloud
#
# The picture below is what entering a workspace will look like
#
# 
#
# ## Create a Cluster
#
# From here, you can create a cluster by clicking the “New Cluster” link. The cluster will serve as the backend for our Spark commands. With the databricks-connectpackage that we will use later, we can connect our laptop to the cluster automatically just by using Spark commands. The computation graph is built locally and then sent to the cluster for execution.
#
# 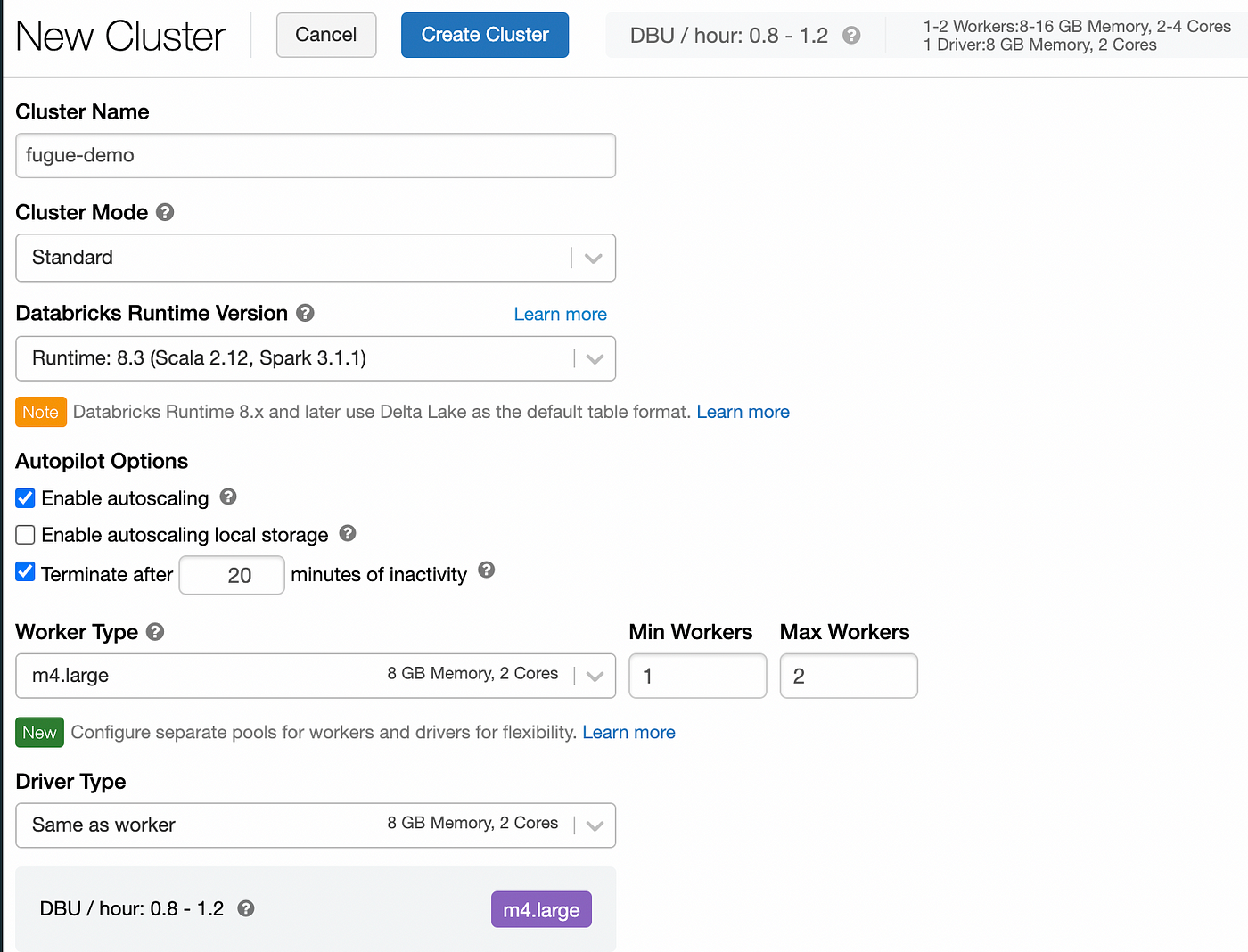
#
# Note when creating a cluster that you can start out by reducing the Worker Type. Clusters can be very expensive! Be sure to lower the worker type to something reasonable for your workload. Also, you can enable autoscaling and terminating the cluster after a certain number of minutes of inactivity.
#
#
# ## Installing Fugue
#
# In order to use Fugue on the cluster from our local environment, we need to install it on the cluster that we created. The easiest way to do this is to navigate the the libraries tab and add the package there. Below is an image of the tab.
#
# 
# ## Databricks-connect
#
# `databricks-connect` is a library that Databricks provides to run Spark commands on the cluster. The content here will just be a summary, but the full guide to installing databricks-connect can be found [here](https://docs.databricks.com/dev-tools/databricks-connect.html).
#
# `databricks-connect` can be installed using `pip`:
#
# ```
# pip uninstall pyspark
# pip install databricks-connect
# ```
#
# Pyspark needs to be uninstalled because it conflicts with `databricks-connect`. databricks-connect will replace your pyspark installation. This means that `import pyspark` will load the databricks-connect version of PySpark and all succeeding Spark commands are sent to the cluster for execution.
#
# Note that the version of `databricks-connect` must match the Databricks Runtime Version of your cluster. Otherwise, you will run into errors and the code will not be able to execute correctly.
#
# ## Configuring the Cluster
#
# Now that you have a cluster created from the first step and `databricks-connect` installed, you can configure the cluster by doing `databricks-connect configure`.
#
# There are more details where to get the relevant pieces of information in their documentation here. For my example on AWS, it looked like this:
#
# 
#
# You can verify if this worked by using `databricks-connect test`, and then you should see a message that all tests passed.
# ## Fugue and databricks-connect
#
# After setting the cluster up and configuring `databricks-connect` to point to the cluster, there is no added effort needed to connect Fugue to your Spark cluster. The `SparkExecutionEngine` imports `pyspark`, meaning that it will import the `databricks-connect` configuration under the hood and use the configured cluster. Fugue works with `databricks-connect` seamlessly, allowing for convenient switching between local development and a remote cluster.
#
# The code below will execute on the Databricks cluster if you followed the steps above. In order to run this locally, simple use the default `NativeExecutionEngine` instead of the `SparkExecutionEngine`
# +
import pandas as pd
from fugue import transform
from fugue_spark import SparkExecutionEngine
data = pd.DataFrame({'numbers':[1,2,3,4], 'words':['hello','world','apple','banana']})
# schema: *, reversed:str
def reverse_word(df: pd.DataFrame) -> pd.DataFrame:
df['reversed'] = df['words'].apply(lambda x: x[::-1])
return df
spark_df = transform(data, reverse_word, engine=SparkExecutionEngine())
spark_df.show()
# -
# ## Additional Configuration
#
# Most `databricks-connect` users add additional Spark configurations on the cluster through the DAtabricks UI. If additional configruation is needed, it can be provided with the following syntax:
# +
from pyspark.sql import SparkSession
from fugue_spark import SparkExecutionEngine
spark_session = (SparkSession
.builder
.config("spark.executor.cores",4)
.config("fugue.dummy","dummy")
.getOrCreate())
engine = SparkExecutionEngine(spark_session, {"additional_conf":"abc"})
# -
# ## Using Fugue-sql on the Cluster
#
# Because Fugue-sql also just uses the `SparkExecutionEngine`, it can also be easily executed on a remote cluster.
from fugue_notebook import setup
setup()
# %%fsql spark
SELECT *
FROM data
TRANSFORM USING reverse_word
PRINT
# ## Conclusion
#
# Here we have shown how to connect Fugue to a Databricks cluster. Using Fugue along with `databricks-connect` provides the best of both worlds, only utilitizing the cluster when needed.
#
# `databricks-connect` can slow down developer productitity and increase compute costs because all the Spark code becomes configured to run on the cluster. Using Fugue, we can toggle between Fugue's default `NativeExecutionEngine` and `SparkExecutionEngine`. The default `NativeExecutionEngine` will run on local without using Spark and Fugue's `SparkExecutionEngine` will seamlessly use whatever `pyspark` is configured for the user.
#
# Fugue also allows for additional configuration of the underlying frameworks. We showed the syntax for passing a `SparkSession` to the `SparkExecutionEngine`.
| tutorials/applications/databricks_connect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.chdir('..')
os.chdir('..')
print(os.getcwd())
import rsnapsim as rss
import numpy as np
os.chdir('rsnapsim')
os.chdir('interactive_notebooks')
import matplotlib.pyplot as plt
import time
poi_strs, poi_objs, tagged_pois,seq = rss.seqmanip.open_seq_file('../gene_files/Bactin_withTags.txt')
Bactin_obj = tagged_pois['1'][0]
rss.solver.protein = Bactin_obj #pass the protein object
t = np.linspace(0,1000,2001)
solution = rss.solver.solve_ssa(Bactin_obj.kelong, t, ki=.033, kt = 10, n_traj=10,low_memory=False,record_stats=True)
solver = rss.solver
# -
plt.style.use('dark_background')
plt.rcParams['figure.dpi'] = 120
plt.rcParams['lines.linewidth'] = 1
plt.rcParams['axes.linewidth'] = 1.5
plt.rcParams['font.size'] = 15
plt.rcParams['axes.grid'] = False
frag_start_t, fragments = rss.inta.get_fragments(solution.ribosome_locations[0].T)
for fragt,frag in zip(frag_start_t,fragments):
print(fragt,frag)
print(solution.time[fragt:fragt+len(frag)])
plt.plot(frag,solution.time[fragt:fragt+len(frag)], )
# +
ft, fa = rss.inta.get_fragments(solution.ribosome_locations[0].T)
for i in range(len(fa)):
frag = fa[i][fa[i] > 0]
timeseg = t[ft[i]: ft[i] + len(frag) ]
plt.plot(frag ,timeseg,'#cccccc')
for i in range(len(solution.col_points[0])):
plt.plot(solution.col_points[0][i,0],solution.col_points[0][i,1],'.',color='#05FF22')
plt.gca().invert_yaxis()
plt.xlabel('codon position')
plt.ylabel('Time (s)')
# -
solution.col_points[0][i,:]
# +
ke = Bactin_obj.kelong
ke[400] = .1
ke[600] = .1
rss.solver.protein = Bactin_obj #pass the protein object
t = np.linspace(0,1000,2001)
solution = rss.solver.solve_ssa(ke, t, ki=.033, kt = 10, n_traj=10,low_memory=False,record_stats=True)
solver = rss.solver
| rsnapsim/interactive_notebooks/.ipynb_checkpoints/Example Kymograph-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Imports
#
# Import things needed for Tensorflow and CoreML
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __builtin__ import any as b_any
import math
import os
os.environ["CUDA_VISIBLE_DEVICES"]=""
import numpy as np
from PIL import Image
import tensorflow as tf
import configuration
import inference_wrapper
import sys
sys.path.insert(0, 'im2txt/inference_utils')
sys.path.insert(0, 'im2txt/ops')
import caption_generator
import image_processing
import vocabulary
import urllib, os, sys, zipfile
from os.path import dirname
from tensorflow.core.framework import graph_pb2
from tensorflow.python.tools.freeze_graph import freeze_graph
from tensorflow.python.tools import strip_unused_lib
from tensorflow.python.framework import dtypes
from tensorflow.python.platform import gfile
import tfcoreml
import configuration
from coremltools.proto import NeuralNetwork_pb2
# -
# Turn on debugging on error
# %pdb off
# ## Create the models
#
# Create the Tensorflow model and strip all unused nodes
# +
checkpoint_file = './trainlogIncNEW/model.ckpt-1000000'
pre_frozen_model_file = './frozen_model_textgenCUSTOM.pb'
frozen_model_file = './frozen_model_textgenCUSTOM.pb'
# Which nodes we want to input for the network
# Use ['image_feed'] for just Memeception
input_node_names = ['seq_embeddings','lstm/state_feed']
# Which nodes we want to output from the network
# Use ['lstm/initial_state'] for just Memeception
output_node_names = ['softmax_T','lstm/state']
# Set the depth of the beam search
beam_size = 2
# +
# Build the inference graph.
g = tf.Graph()
with g.as_default():
model = inference_wrapper.InferenceWrapper()
restore_fn = model.build_graph_from_config(configuration.ModelConfig(),
checkpoint_file)
g.finalize()
# +
# Write the graph
tf_model_path = './log/pre_graph_textgenCUSTOM.pb'
tf.train.write_graph(
g,
'./log',
'pre_graph_textgenCUSTOM.pb',
as_text=False,
)
with open(tf_model_path, 'rb') as f:
serialized = f.read()
tf.reset_default_graph()
original_gdef = tf.GraphDef()
original_gdef.ParseFromString(serialized)
# +
# Strip unused graph elements and serialize the output to file
gdef = strip_unused_lib.strip_unused(
input_graph_def = original_gdef,
input_node_names = input_node_names,
output_node_names = output_node_names,
placeholder_type_enum = dtypes.float32.as_datatype_enum)
# Save it to an output file
with gfile.GFile(pre_frozen_model_file, 'wb') as f:
f.write(gdef.SerializeToString())
# +
# Freeze the graph with checkpoint data inside
freeze_graph(input_graph=pre_frozen_model_file,
input_saver='',
input_binary=True,
input_checkpoint=checkpoint_file,
output_node_names=','.join(output_node_names),
restore_op_name='save/restore_all',
filename_tensor_name='save/Const:0',
output_graph=frozen_model_file,
clear_devices=True,
initializer_nodes='')
# -
# ## Verify the model
#
# Check that it is producing legit captions for *One does not simply*
# +
# Configure the model and load the vocab
config = configuration.ModelConfig()
vocab_file ='vocab4.txt'
vocab = vocabulary.Vocabulary(vocab_file)
# +
# Generate captions on a hard-coded image
with tf.Session(graph=g) as sess:
restore_fn(sess)
generator = caption_generator.CaptionGenerator(
model, vocab, beam_size=beam_size)
for i,filename in enumerate(['memes/advice-god.jpg']):
with tf.gfile.GFile(filename, "rb") as f:
image = Image.open(f)
image = ((np.array(image.resize((299,299)))/255.0)-0.5)*2.0
for k in range(10):
captions = generator.beam_search(sess, image)
for i, caption in enumerate(captions):
sentence = [vocab.id_to_word(w) for w in caption.sentence[1:-1]]
sentence = " ".join(sentence)
print(sentence)
# -
# ## Convert the model to CoreML
#
# Specify output variables from the graph to be used
# +
# Define basic shapes
# If using Memeception, add 'image_feed:0': [299, 299, 3]
input_tensor_shapes = {
'seq_embeddings:0': [1, beam_size, 300],
'lstm/state_feed:0': [1, beam_size, 1024],
}
coreml_model_file = './Textgen_CUSTOM.mlmodel'
# -
output_tensor_names = [node + ':0' for node in output_node_names]
weightLSTM = np.loadtxt('weightLSTM')
weightFully = np.loadtxt('weightFully')
# +
def convert_matmul(**kwargs):
# Only convert this Lambda layer if it is for our swish function.
tf_op = kwargs["op"]
if tf_op.name == 'lstm/basic_lstm_cell/LSTMmatmul2':
W = weightLSTM
print('LSTM')
else:
W = weightFully
print('Fully')
coreml_nn_builder = kwargs["nn_builder"]
constant_inputs = kwargs["constant_inputs"]
params = NeuralNetwork_pb2.CustomLayerParams()
# The name of the Swift or Obj-C class that implements this layer.
params.className = "MatMul"
# The desciption is shown in Xcode's mlmodel viewer.
params.description = "A fancy new matmul"
#W = constant_inputs.get(tf_op.inputs[1].name,[0,100,0,0])
#print(tf_op.inputs[1])
#size = constant_inputs.get(tf_op.inputs[2].name, [0,0,0,0])
# add begin and size as two repeated weight fields
for i,weightvec in enumerate(W):
W_as_weights = params.weights.add()
W_as_weights.floatValue.extend(map(float, weightvec))
#print(W_as_weights)
#size_as_weights = params.weights.add()
#size_as_weights.floatValue.extend(map(float, size))
coreml_nn_builder.add_custom(name=tf_op.name,
input_names=[tf_op.inputs[0].name],
output_names=[tf_op.outputs[0].name],
custom_proto_spec=params)
#return params
coreml_model = tfcoreml.convert(
tf_model_path=frozen_model_file,
mlmodel_path=coreml_model_file,
input_name_shape_dict=input_tensor_shapes,
output_feature_names=output_tensor_names,
add_custom_layers=True,
custom_conversion_functions={ "lstm/basic_lstm_cell/LSTMmatmul2": convert_matmul, "logits/Fullymatmul": convert_matmul}
#custom_conversion_functions={ "MatMuldlskfjslkfj": convert_matmul}
)
# -
# ## Test the model
#
# Run a predictable randomly seeded inputs through and see where the disparities are
# +
seq_rand = np.random.rand(300)
seq_embeddings_tf = np.array([[seq_rand, seq_rand]])
seq_embeddings_ml = np.array([[[sr, sr]] for sr in seq_rand])
state_rand = np.random.rand(1024)
state_feed_tf = np.array([[state_rand, state_rand]])
state_feed_ml = np.array([[[sr, sr]] for sr in state_rand])
# -
coreml_inputs = {
'seq_embeddings__0': seq_embeddings_ml,
'lstm__state_feed__0': state_feed_ml,
}
coreml_output = coreml_model.predict(coreml_inputs, useCPUOnly=True)
# print(coreml_output['lstm__state__0'].shape)
# print(coreml_output['softmax__0'].shape)
# print(coreml_output['softmax__0'].reshape(38521, 1, 2))
# print(coreml_output)
def print_ml(ml):
for key in sorted(ml.keys()):
print(key)
print(ml[key].shape)
print(ml[key])
print_ml(coreml_output)
with tf.Session(graph=g) as sess:
# Load the model from checkpoint.
restore_fn(sess)
input_names = ['lstm/state:0', 'softmax:0', 'softmax_T:0']
output_values = sess.run(
fetches=input_names,
feed_dict={
#"input_feed:0": input_feed,
"lstm/state_feed:0": state_feed_tf,
"seq_embeddings:0": seq_embeddings_tf,
#"seq_embedding/embedding_map:0": self.embedding_map
})
for (index, value) in sorted(enumerate(input_names), key=lambda x: x[1]):
print(value)
print(output_values[index].shape)
print(output_values[index])
np.matmul(np.random.rand(1, 20), np.random.rand(20, 45)).shape
np.random.rand(1, 2, 812)[0,:].shape
| im2txt/model_conversion_debug.ipynb |