code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
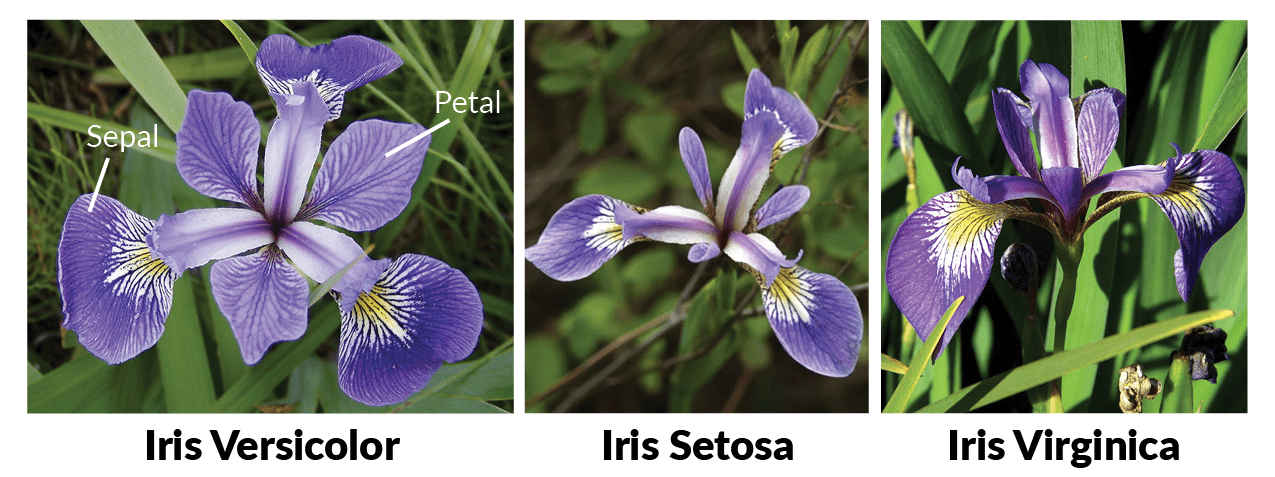
# # The Iris dataset and pandas
#
# 
#
# 
#
# ***
#
# **[Python Data Analysis Library](https://pandas.pydata.org/)**
#
# *[https://pandas.pydata.org/](https://pandas.pydata.org/)*
#
# The pandas website.
#
# ***
#
# **[<NAME>: pandas in 10 minutes | Walkthrough](https://www.youtube.com/watch?foo=bar&v=_T8LGqJtuGc?)**
#
# *[https://www.youtube.com/watch?v=_T8LGqJtuGc](https://www.youtube.com/watch?foo=bar&v=_T8LGqJtuGc)*
#
# Video by the creator of pandas.
#
# ***
#
# **[Python for Data Analysis notebooks](https://github.com/wesm/pydata-book)**
#
# *[https://github.com/wesm/pydata-book](https://github.com/wesm/pydata-book)*
#
# Materials and IPython notebooks for "Python for Data Analysis" by <NAME>, published by O'Reilly Media
#
# ***
#
# **[10 Minutes to pandas](http://pandas.pydata.org/pandas-docs/stable/10min.html)**
#
# *[http://pandas.pydata.org/pandas-docs/stable/10min.html](http://pandas.pydata.org/pandas-docs/stable/10min.html)*
#
# Official pandas tutorial.
#
# ***
#
# **[UC Irvine Machine Learning Repository: Iris Data Set](https://archive.ics.uci.edu/ml/datasets/iris)**
#
# *[https://archive.ics.uci.edu/ml/datasets/iris](https://archive.ics.uci.edu/ml/datasets/iris)*
#
# About the Iris data set from UC Irvine's machine learning repository.
# ## Loading data
# Import pandas.
import pandas as pd
# Load the iris data set from a URL.
df = pd.read_csv("https://raw.githubusercontent.com/uiuc-cse/data-fa14/gh-pages/data/iris.csv")
df
# ***
#
# ## Selecting rows and columns
df['species']
df[['petal_length', 'species']]
df[2:6]
df[['petal_length', 'species']][2:6]
df.loc[2:6]
df.loc[:, 'species']
df.loc[:, ['sepal_length', 'species']]
df.loc[2:6, ['sepal_length', 'species']]
df.iloc[2]
df.iloc[2:4, 1]
df.at[3, 'species']
df.iloc[1:10:2]
# ***
#
# ## Boolean selects
df.loc[:, 'species'] == 'setosa'
df.loc[df.loc[:, 'species'] == 'versicolor']
x = df.loc[df.loc[:, 'species'] == 'versicolor']
x.loc[51]
x.iloc[1]
# ***
#
# ## Summary statictics
df.head()
df.tail()
df.describe()
(df.loc[df.loc[:, 'species'] == 'versicolor']).describe()
(df.loc[df.loc[:, 'species'] == 'setosa']).describe()
(df.loc[df.loc[:, 'species'] == 'virginica']).describe()
df.mean()
# ***
#
# ## Plots
import seaborn as sns
sns.pairplot(df, hue='species')
# ***
#
# ## End
| pandas-with-iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-exploratory-data-analysis
# language: python
# name: venv-exploratory-data-analysis
# ---
# # Grid search
#
# technique used for model tuning that tries several hyperparameters combinations and reports the best based on a metric
# +
# %matplotlib inline
from sklearn import svm
from sklearn.model_selection import GridSearchCV, train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.datasets import make_multilabel_classification
import pandas as pd
import numpy as np
import category_encoders as ce
import multiprocessing
random_state = 42
n_cpu = multiprocessing.cpu_count()
n_cpu
# -
def get_data():
return make_multilabel_classification(n_samples=1_000, n_features=15, n_classes=5, n_labels=2, allow_unlabeled=False, sparse=False, return_distributions=False, random_state=random_state)
data = get_data()
X = data[0]
X[:5,:]
y = np.argmax(data[1], axis=1)
y[:20]
X_train, X_test, y_train, y_test = train_test_split(X, y)
svc = svm.SVC()
svc.fit(X_train, y_train)
preds = svc.predict(X_test)
print(classification_report(y_test, preds))
# ## Try grid search
parameters = {
'kernel': ['linear', 'rbf', 'poly', 'sigmoid'],
'C': [0.1, 0.5, 1.0, 10],
'degree': [1, 3, 5]
}
gs = GridSearchCV(svc, parameters, n_jobs=n_cpu)
gs.fit(X_train, y_train)
gs.best_estimator_
gs.best_score_
gs.best_params_
svc = svm.SVC(kernel='poly', degree=1)
svc.fit(X_train, y_train)
preds = svc.predict(X_test)
print(classification_report(y_test, preds))
| notebooks/012. grid search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from deeptables.models import deeptable,deepnets
from deeptables.utils import consts,dt_logging,batch_trainer
from deeptables.datasets import dsutils
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score, roc_curve
import pandas as pd
import numpy as np
data = dsutils.load_adult()
# +
conf1 = deeptable.ModelConfig(
name='conf1',
apply_gbm_features=True,
gbm_feature_type='dense',
auto_discrete=True,
metrics=['AUC'],
)
conf2 = deeptable.ModelConfig(
name='conf2',
fixed_embedding_dim=False,
embeddings_output_dim=0,
apply_gbm_features=False, #*
gbm_feature_type='dense',
auto_discrete=False, #*
metrics=['AUC'],
)
bt = batch_trainer.BatchTrainer(data,'x_14',
eval_size=0.2,
validation_size=0.2,
eval_metrics=['AUC','accuracy','recall','precision','f1'], #auc/recall/precision/f1/mse/mae/msle/rmse/r2
verbose=1,
dt_epochs=15,
dt_config=[conf1,conf2],
dt_nets=[['dnn_nets'],['cross_nets']],
)
ms = bt.start()
# -
ms.leaderboard()
| examples/batch_trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# \# Developer: <NAME> (<EMAIL>) <br>
# \# 18th December 2018
import pandas as pd
bugs = pd.read_csv('../datasets/lexical_semantic_preprocessed_mantis_bugs_less_columns_with_class.csv')
bug_notes = pd.read_csv('../datasets/lexical_semantic_preprocessed_mantis_bugnotes.csv', index_col='bug_id')
bugs_list = []
no_bug_notes = []
for index, row in bugs.iterrows():
pos = e_pos = neg = e_neg = neutral = 0
bug_features = {'id': row['id'], 'pos': 0, 'neg': 0, 'neu': 0, 'priority': row['priority'],
'severity': row['severity'], 'pm_ticket': row['pm_ticket']}
for col in ['additional_information_vader_polarity', 'summary_vader_polarity', 'description_vader_polarity']:
bug_features[row[col]] += row[col+'_weight']
try:
bug_notes_for_id = bug_notes.loc[row['id'], ['bug_note_vader_polarity', 'bug_note_vader_polarity_weight']]
if isinstance(bug_notes_for_id, pd.core.frame.DataFrame):
for index, row in bug_notes_for_id.iterrows():
bug_features[row['bug_note_vader_polarity']] += row['bug_note_vader_polarity_weight']
else:
bug_features[bug_notes_for_id['bug_note_vader_polarity']] += bug_notes_for_id['bug_note_vader_polarity_weight']
except Exception as e:
no_bug_notes.append(row['id'])
bugs_list.append(bug_features)
vector_df = pd.DataFrame(bugs_list)
vector_df['total_sentiments'] = vector_df['pos'].abs() + vector_df['neg'].abs() + vector_df['neu'].abs()
for col in ['pos', 'neg', 'neu']:
vector_df[col+'_normalized'] = vector_df[col].abs() / vector_df['total_sentiments']
vector_df.to_csv('../datasets/mantis_bugs_vector.csv', encoding='utf-8', index=False)
result = pd.merge(bugs, df_bug_note_table, how='left', left_on='id', right_on='bug_id')
| 2) class_expansion/.ipynb_checkpoints/3.bug_vector_preperation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# ask the user to insert a sentence
# print each element of the sentence on a different row
# -
sentence = input("tell me something")
for word in sentence.split(" "):
print(word)
| What we did in class/ExercisesSolution/SolutionExercise6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sqlite_db import thesisDB, corpusDB, enclose, docData
from pynlpl.clients.frogclient import FrogClient
from lexisnexisparse import LexisParser
import json
import pandas as pd
from ipywidgets import FloatProgress
from IPython.display import display
import re
import os
#settings_file = 'D:/thesis/settings - nl_final.json'
frogip = '192.168.1.126'
frogport = 9772
#Read settings
#settings = json.loads(open(settings_file).read())["settings"]
#dbAddress = settings['db_file']
#datasets = [settings['json_folder']+fname for fname in os.listdir(settings['json_folder']) if fname.lower().endswith(".json.gz")]
dbAddress = 'D:/Thesis/data.db'
datasets = ['D:/Thesis/jsons/trouw.json.gz']
db = thesisDB(dbAddress)
# -
##Connect to frog client
frog = FrogClient(frogip,frogport,returnall = True)
# +
#Progressbar!
f = FloatProgress(min = 0, max = 1, bar_style = 'success')
display(f) # display the bar
for dataset in datasets:
df = pd.read_json(dataset, compression = 'gzip')
df.sort_index(inplace = True)
f.description = dataset[54:-8]
f.value = 0
f.max = len(df)
counter = 0 #commit every 5 documents
for index,row in df.iterrows():
counter += 1
##First input document, save its rowid for cross-reference
document = {'date':str(row['DATE_dt']),
'medium':enclose(row['MEDIUM']),
'headline':enclose(row['HEADLINE']),
'length':str(row['LENGTH'])}
if row['BYLINE']: #sometimes byline is None
document['byline'] = enclose(row['BYLINE'])
if row['SECTION']: #sometimes sections is None
document['section'] = enclose(row['SECTION'])
if counter % 10 == 0:
lastRow = db.insertRow('documents',document)
else:
lastRow = db.insertRow('documents',document,False)
paragraph_no = 1
entities = []
entity = ['','']
for paragraph in row['TEXT']:
res = frog.process(paragraph)
position = 1
for row in res:
if row[0] is None:
continue
if row[0] == '"':
row = list(row)
row[0] = 'DOUBLE_QUOTE'
row[1] = 'DOUBLE_QUOTE'
row = tuple(row)
data = {
'token':enclose(row[0]),
'lemma':enclose(row[1]),
'paragraph_no':str(paragraph_no),
'position':str(position),
'docid':str(lastRow),
'pos':enclose(re.search('^[A-Z]+',row[3])[0])#, Exclude pos_long because it takes storage and is not needed
#'pos_long':enclose(row[3])
}
db.insertRow('tokens',data,False)
if row[4] != 'O': #Found an entity
if re.match('^B-',row[4]) is not None:
#Entity is new. Save old entity if something is stored
if entity[0] != '':
entities.append(entity)
entity = ['','']
entity[0] = row[0]
entity[1] = re.search('[A-Z]+$',row[4])[0]
else:
entity[0] += ' '+row[0] #append next term of entity
position += 1
paragraph_no += 1
if entity[0] != '': #append last entity if present
entities.append(entity)
for ent,t in entities:
data = {'entity':enclose(ent),
'category':enclose(t),
'docid':str(lastRow)}
db.insertRow('entities',data,False)
f.value += 1
db.commit()
# -
frog.close()
| Pipeline frog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: diagnosis
# language: python
# name: diagnosis
# ---
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.model_selection import KFold, StratifiedKFold, train_test_split
from sklearn.metrics import accuracy_score
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.externals import joblib
import lightgbm as lgb
import scipy
from scipy import sparse
from pandas.core.common import SettingWithCopyWarning
import scipy.stats as sp
import pandas as pd
import numpy as np
from collections import Counter
import warnings
import time
import sys
import random
import os
import gc
import datetime
# +
path = '../data/'
data_path = '../trainTestData/'
middle_path = '../model/'
train_y = pd.read_csv(path + 'age_train.csv', names=['uid', 'label'])
sub = pd.read_csv(path + 'age_test.csv', names=['uid'])
train_csr = sparse.load_npz(data_path + 'trainData15112.npz')
test_csr = sparse.load_npz(data_path + 'testData15112.npz')
train_y = train_y["label"].values
# train_csr = train_csr[:200]
# train_y = train_y['label'].values[:200]
# print(train_csr.shape, test_csr.shape)
lgb_model = lgb.LGBMClassifier(
boosting_type='gbdt',
objective='multiclass',
metrics='multi_error',
num_class=6,
n_estimators=20000,
learning_rate=0.1,
num_leaves=512,
max_depth=-1,
subsample=0.95,
colsample_bytree=0.5,
subsample_freq=1,
reg_alpha=1,
reg_lambda=1,
random_state=42,
n_jobs=10
)
oof = np.zeros((train_csr.shape[0], 6))
sub_preds = np.zeros((test_csr.shape[0], 6))
skf = StratifiedKFold(n_splits=5, random_state=812, shuffle=True)
t = time.time()
for index, (train_index, test_index) in enumerate(skf.split(train_csr, train_y)):
print('Fold {}'.format(index + 1))
lgb_model.fit(train_csr[train_index], train_y[train_index],
eval_set=[(train_csr[train_index], train_y[train_index]),
(train_csr[test_index], train_y[test_index])],
eval_names=['train', 'valid'],
early_stopping_rounds=200, verbose=10)
oof[test_index] = lgb_model.predict_proba(train_csr[test_index], num_iteration=lgb_model.best_iteration_)
sub_preds += lgb_model.predict_proba(test_csr, num_iteration=lgb_model.best_iteration_) / skf.n_splits
# lgb_model.savetxt(middle_path+'model/lgb_zl'+str(index)+'_model.txt')
joblib.dump(lgb_model, '../model/lgb_zl_15112_2'+str(index)+'_model.pkl')
print(oof.shape, train_y.shape)
cv_final = accuracy_score(train_y, np.argmax(oof, axis=1)+1)
print('\ncv acc:', cv_final)
sub['label'] = np.argmax(sub_preds, axis=1) + 1
# sub.to_csv(middle_path + 'sub_{}.csv'.format(cv_final), index=False)
oof = np.zeros((train_csr.shape[0], 6))
sub_preds = np.zeros((test_csr.shape[0], 6))
skf = StratifiedKFold(n_splits=5, random_state=812, shuffle=True)
t = time.time()
for index, (train_index, test_index) in enumerate(skf.split(train_csr, train_y)):
print('Fold {}'.format(index + 1))
lgb_model = joblib.load('../model/lgb_zl_15112_2'+str(index)+'_model.pkl')
oof[test_index] = lgb_model.predict_proba(train_csr[test_index], num_iteration=lgb_model.best_iteration_)
sub_preds += lgb_model.predict_proba(test_csr, num_iteration=lgb_model.best_iteration_) / skf.n_splits
cv_final = accuracy_score(train_y, np.argmax(oof, axis=1)+1)
print('\ncv acc:', cv_final)
np.savetxt('../processed/lgboost_val_15112.txt', oof, fmt='%s', delimiter=',', newline='\n')
np.savetxt('../processed/lgboost_test_15112.txt', sub_preds, fmt='%s', delimiter=',', newline='\n')
sub['label'] = np.argmax(sub_preds, axis=1) + 1
# sub.to_csv(middle_path + 'sub_{}.csv'.format(cv_final), index=False)
# -
| src/LightGBM_15112.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
proj = pd.read_csv('./Projects.csv')
# +
# proj.to_pickle("./projects.pkl")
# -
print(proj.columns)
print(proj.head)
proj_cleaned = proj.drop(['Project Essay', 'Project Short Description', 'Project Need Statement'], axis=1)
print(proj_cleaned.columns)
proj_cleaned.to_pickle("./projects_cleaned.pkl")
| data/DataProcessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iam-abbas/Reddit-Stock-Trends/blob/main/Reddit_Stock_Trends.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YTwciulmQ9q2"
# # Trending Stocks on Sub-Reddits
# + [markdown] id="8j6mzW3_Q0WX"
# ### Installing Reddit and Yahoo Finance API wrappers
# + id="nv7x3u7GzoC8" colab={"base_uri": "https://localhost:8080/"} outputId="a169723f-9f0c-4e2f-b7c3-30132fa740b0"
pip install yfinance praw==7.0
# + [markdown] id="mK6yAb7PRFF4"
# ### Imports
# + id="igjxSo-6cw8_"
import numpy as np
import pandas as pd
import yfinance as yf
import re
import praw
# + [markdown] id="4iCXDsOURV4-"
# ### Stop words and Blacklist containing jargon/acronyms
# I added GME and AMC in there lmao, tired of seeing those
# + id="5xTgoasKHigE"
stop_words = ['i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've", "you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', 'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their', 'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', 'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', 'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', 'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after', 'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further', 'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more', 'most', 'other', 'some', 'such', 'no', 'nor', 'not', 'only', 'own', 'same', 'so', 'than', 'too', 'very', 's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', 've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn', "hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'must', 'mustn', "mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", 'won', "won't", 'wouldn', "wouldn't"]
# + id="j2yODV10FMLi"
block_words = ["DIP", "", "$", "RH", "YOLO", "PORN", "BEST", "MOON", "HOLD", "FAKE", "WISH", "USD", "EV", "MARK", "RELAX", "LOL", "LMAO", "LMFAO", "EPS", "DCF", "NYSE", "FTSE", "APE", "CEO", "CTO", "FUD", "DD", "AM", "PM", "FDD", "EDIT", "TA", "UK", "AMC", "GME"]
# + [markdown] id="wdteNrESRYoJ"
# ### Simple Regex to get tickers from text
# + id="TC-aZcvEdPwZ"
def extract_ticker(body):
ticks = re.findall("[$][A-Za-z]*|[A-Z][A-Z]{1,}", str(body))
res = set()
for item in ticks:
if item not in block_words and item.lower() not in stop_words and item:
try:
tic = item.replace("$", "").upper()
res.add(tic)
except:
pass
return res
# + [markdown] id="pYbo-KUjRgvx"
# ### Reddit API setup
# + id="SZy2Ql_W1crv"
reddit = praw.Reddit(
client_id="9Aq-wTeGLJBKsQ",
client_secret="<KEY>",
user_agent="ScrapeStocks"
)
# + [markdown] id="M79xPC9cRl9_"
# ### Scrape subreddits `r/robinhoodpennystocks` and `r/pennystocks`
# Current it does fetch a lot of additional data like upvotes, comments, awards etc but not using anything apart from title for now
# + id="sqHNK1q86Qiv"
_posts = []
new_bets = reddit.subreddit("robinhoodpennystocks+pennystocks").new(limit=None)
new_bets
for post in new_bets:
_posts.append(
[
post.id,
post.title,
post.score,
post.num_comments,
post.upvote_ratio,
post.total_awards_received,
]
)
# create a dataframe
_posts = pd.DataFrame(
_posts,
columns=[
"id",
"title",
"score",
"comments",
"upvote_ratio",
"total_awards",
],
)
# + id="j2IwLMtDL7ao" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="59baed7b-eee1-41d6-cc97-f90b9f770938"
_posts
# + [markdown] id="OY0wSH7nR4tO"
# ### Extract tickers from all titles and creae a new columns
# + id="lqLo3rvFVqjH"
_posts["Tickers"] = _posts.title.apply(extract_ticker)
# + id="KZTR2SWv_idY"
ticker_sets = _posts.Tickers.to_list()
# + [markdown] id="tDZMN4UMR9Kp"
# ### Count number of occurances of the Ticker and verify id the Ticker exists
# This kinda works like a hashmap
# + id="JEidqwnNDDxJ"
counts ={}
# + id="qXjeqn8pNTK3"
for s in ticker_sets:
for tic in s:
if tic in counts:
counts[tic] += 1
else:
counts[tic] = 1
# + id="6UmWCKL9XsJS"
verified_tics = {}
# + colab={"base_uri": "https://localhost:8080/"} id="ZboCwlBwXfjG" outputId="8c67260e-f3ac-46d0-a5ea-91cdaa0ae7c1"
for tic in counts:
try:
if counts[tic] > 3:
print(tic, end=", ")
yf.Ticker(tic).info
verified_tics[tic] = counts[tic]
else:
continue
except:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="pPTVFuTOWtUn" outputId="6c9301ee-3320-4a58-f782-b8acdbb3c459"
verified_tics
# + [markdown] id="ClL4RoY0SPLs"
# ### Create Datable of just mentions
# + id="HTFGjj6zaqXu"
tick_df = pd.DataFrame(verified_tics.items(), columns=["Ticker", "Mentions"])
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="O9stuq8DcyW0" outputId="5e71d3c0-06bc-46a9-a93f-f72c4ab4104e"
tick_df
# + [markdown] id="9l83_x5RSV-F"
# ### Sort
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="3oY2bLuucy9u" outputId="a409cfcd-435a-4391-f629-7f422642be18"
tick_df.sort_values(by=["Mentions"], inplace=True, ascending=False)
tick_df.reset_index(inplace=True, drop=True)
tick_df
# + [markdown] id="HdvynA8LSXx0"
# ### Use Yahoo Finance API to get the relavent data
# + id="JJI4GGUfgfnZ"
def calculate_change(start, end):
return round(((end - start)/start)*100, 2)
# + id="FvnQCArshKwG"
def get_change(ticker, period="1d"):
return calculate_change(yf.Ticker(ticker).history(period)["Open"].to_list()[0], yf.Ticker(ticker).history(period)["Close"].to_list()[-1])
# + id="WiBcQD2miilm"
tick_df.dropna(axis=1)
n25_df = tick_df.head(25)
# + colab={"base_uri": "https://localhost:8080/"} id="qFvdVT3rg-Cr" outputId="b59e72ec-7be0-4eb8-e057-99c8b95d3b48"
n25_df["Name"] = n25_df.Ticker.apply(lambda x: yf.Ticker(x).info["longName"])
n25_df["Bid"] = n25_df.Ticker.apply(lambda x: yf.Ticker(x).info["previousClose"])
n25_df["5d Low"] = n25_df.Ticker.apply(lambda x: min(yf.Ticker(x).history(period="5d")['Low'].to_list()))
n25_df["5d High"] = n25_df.Ticker.apply(lambda x: max(yf.Ticker(x).history(period="5d")['High'].to_list()))
n25_df["1d Change (%)"] = n25_df.Ticker.apply(lambda x: get_change(x))
n25_df["5d Change (%)"] = n25_df.Ticker.apply(lambda x: get_change(x, "5d"))
# + colab={"base_uri": "https://localhost:8080/"} id="66kcExXIpwp1" outputId="3f8245b7-bf7d-44da-8aa9-b8b5c0c7ccab"
n25_df["1mo Change (%)"] = n25_df.Ticker.apply(lambda x: get_change(x, "1mo"))
# + colab={"base_uri": "https://localhost:8080/", "height": 824} id="u1mvvIoSjXC9" outputId="f252b06b-57af-4b70-c5b6-c99b043eb890"
n25_df
# + colab={"base_uri": "https://localhost:8080/"} id="_6zvqkcrDzWo" outputId="428d3b02-b9a5-4151-c10f-7d3c1d8bfd17"
n25_df.rename(columns={'Bid': 'Price - 2/5'}, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="nyBSbceSFonB" outputId="2906c819-f508-4932-e8d7-dc4ffc901b4d"
n25_df["Industry"] = n25_df.Ticker.apply(lambda x: yf.Ticker(x).info['industry'])
# + colab={"base_uri": "https://localhost:8080/", "height": 824} id="evApezERGIep" outputId="041665b0-d373-458e-e2ba-107667b6e466"
n25_df
# + id="c1X35SLwMJDt"
| Reddit_Stock_Trends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pickle
from sklearn.preprocessing import StandardScaler
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pe_data import *
from hashing_vectorizer import *
# +
default_data_dir_path = "./data"
default_result_dir_path = "./result"
train_df_dir_path = "./data/v1"
test_df_dir_path = "./data/v1_test"
std_scaler_file_path = os.path.join(default_result_dir_path, "std_scaler_numeric_features.pkl")
l1_extracted_feature_file_path = os.path.join(default_result_dir_path, "l1_svm_extracted_numeric_features.pkl")
xgb_extracted_feature_file_path = os.path.join(default_result_dir_path, "xgb_extracted_numeric_features.pkl")
l1_svm_model_file_path = os.path.join(default_result_dir_path, "numeric_l1_svm_model.pkl")
xgb_model_file_path = os.path.join(default_result_dir_path, "numeric_xgb_model.pkl")
hashing_vectorizer_dir_path = os.path.join(default_result_dir_path, "hashing_vectorizer")
xgb_clf_file_path = os.path.join(default_result_dir_path, "xgb_clf.pkl")
# -
train_df_file_path_list = [os.path.join(train_df_dir_path, df_file_name) for df_file_name in os.listdir(train_df_dir_path)]
test_df_file_path_list = [os.path.join(test_df_dir_path, df_file_name) for df_file_name in os.listdir(test_df_dir_path)]
test_df_file_path_list
# +
train_df_list = list()
test_df_list = list()
for train_df_file_path in train_df_file_path_list:
train_df_list.append(pd.read_csv(train_df_file_path))
for test_df_file_path in test_df_file_path_list:
test_df_list.append(pd.read_csv(test_df_file_path))
train_df = pd.concat(train_df_list, ignore_index=True)
test_df = pd.concat(test_df_list, ignore_index=True)
# -
# # Train
# ## Numeric Features
train_pfe = PeFeatureExtractor(df=train_df)
train_pfe.preprocessing_label()
train_pfe.preprocessing_type()
numeric_df, str_df = train_pfe.get_numeric_str_df()
numeric_df
# Get extracted feature list
with open(l1_extracted_feature_file_path, "rb") as f:
selected_feature_name_list = pickle.load(f)
selected_numeric_df = numeric_df[selected_feature_name_list]
selected_numeric_df
# Standard scaling
scaler = StandardScaler()
# +
# scaler.fit(X=selected_numeric_df)
# scaled_numeric_data = scaler.transform(X=selected_numeric_df)
# with open(std_scaler_file_path, "wb") as f:
# pickle.dump(scaler, f)
# +
# scaled_numeric_data = scaler.transform(X=selected_numeric_df)
# scaled_numeric_data
# +
with open(std_scaler_file_path, "rb") as f:
scaler = pickle.load(f)
scaled_numeric_data = scaler.transform(X=selected_numeric_df)
# -
scaled_numeric_data
# ## String Features
str_df
vectorizer_object = CustomHashingVectorizer(df=str_df)
column_name_list = str_df.columns
vectorizer_object.load(
column_name_list=column_name_list,
save_dir_path=hashing_vectorizer_dir_path
)
vectorizer_dict = vectorizer_object.vectorizer_dict
# +
str_vector_arr = list()
for column_name in column_name_list:
vectorizer = vectorizer_dict[column_name]
sample_data = str_df[column_name].fillna("")
sample_data = sample_data.map(lambda x : x.replace("/", " "))
sample_data = sample_data.map(lambda x : x.strip())
vector = vectorizer_object.transform(
column_name=column_name,
data=sample_data
)
str_vector_arr.append(vector)
# -
str_vector_arr = np.concatenate(str_vector_arr, axis=1)
scaled_numeric_data.shape
str_vector_arr.shape
train_features = np.concatenate([scaled_numeric_data, str_vector_arr], axis=1)
train_features.shape
# ## Dataset
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(
train_features,
train_pfe.label_list,
test_size=0.2,
random_state=777
)
print(f"Train X shape : {X_train.shape}")
print(f"Val X shape : {X_val.shape}")
print(f"Train y shape : {y_train.shape}")
print(f"Val y shape : {y_val.shape}")
# ## Training (Train / Val)
# ### XGBoost
from xgboost import XGBClassifier
xgb_clf = XGBClassifier(
booster="gbtree",
objective="binary:logistic",
importance_type="gain"
)
xgb_clf.fit(X=X_train, y=y_train)
xgb_pred_list = xgb_clf.predict(X_val)
xgb_pred_list
# ## Evaluation
from sklearn.metrics import classification_report
xgb_result_dict = classification_report(
y_true=y_val,
y_pred=xgb_pred_list,
output_dict=True
)
xgb_result_dict
# ### SVM (rbf)
from sklearn.svm import SVC
svm_clf = SVC(
C=1.0,
kernel="rbf"
)
svm_clf.fit(X=X_train, y=y_train)
svm_pred_list = svm_clf.predict(X_val)
svm_pred_list
# ## Evaluation
svm_result_dict = classification_report(
y_true=y_val,
y_pred=svm_pred_list,
output_dict=True
)
svm_result_dict
# ## Training (Entire)
xgb_entire_clf = XGBClassifier(
booster="gbtree",
objective="binary:logistic",
importance_type="gain"
)
xgb_entire_clf.fit(X=train_features, y=train_pfe.label_list)
# Save
with open(xgb_clf_file_path, "wb") as f:
pickle.dump(xgb_entire_clf, f)
| train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
from tensorflow.examples.tutorials.mnist import input_data
tf.reset_default_graph()
mnist = input_data.read_data_sets("../mnist/data/", one_hot=True)
total_epoch = 15
batch_size = 100
n_hidden = 256
n_input = 28 * 28
n_noise = 128
n_class = 10
X = tf.placeholder(tf.float32, [None, n_input])
Y = tf.placeholder(tf.float32, [None, n_class])
Z = tf.placeholder(tf.float32, [None, n_noise])
def generator(noise, labels):
with tf.variable_scope('generator'):
inputs = tf.concat([noise, labels], 1)
print(inputs)
hidden = tf.layers.dense(inputs, n_hidden, activation=tf.nn.relu, name="gen_hidden")
output = tf.layers.dense(hidden, n_input, activation=tf.nn.sigmoid)
return output
def discriminator(inputs, labels, reuse=None):
with tf.variable_scope('discriminator') as scope:
if reuse:
scope.reuse_variables()
inputs = tf.concat([inputs, labels], 1)
hidden = tf.layers.dense(inputs, n_hidden, activation=tf.nn.relu, name="dis_hidden")
output = tf.layers.dense(hidden, 1, activation=None)
return output
def get_noise(batch_size, n_noise):
return np.random.uniform(-1., 1., size=[batch_size, n_noise])
G = generator(Z, Y)
D_real = discriminator(X, Y)
D_gene = discriminator(G, Y, True)
loss_D_real = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(
logits=D_real, labels=tf.ones_like(D_real)))
loss_D_gene = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(
logits=D_gene, labels=tf.zeros_like(D_gene)))
loss_D = loss_D_real + loss_D_gene
loss_G = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(
logits=D_gene, labels=tf.ones_like(D_gene)))
tf.summary.scalar('costD', loss_D)
tf.summary.scalar('costG', loss_G)
vars_D = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope='discriminator')
vars_G = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope='generator')
train_D = tf.train.AdamOptimizer().minimize(loss_D,
var_list=vars_D)
train_G = tf.train.AdamOptimizer().minimize(loss_G,
var_list=vars_G)
tf.summary.histogram('vars_D', vars_D[0])
tf.summary.histogram('vars_G', vars_G[0])
sess = tf.Session()
merged = tf.summary.merge_all()
writer = tf.summary.FileWriter('./logs', sess.graph)
sess.run(tf.global_variables_initializer())
total_batch = int(mnist.train.num_examples/batch_size)
loss_val_D, loss_val_G = 0, 0
for epoch in range(total_epoch):
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
noise = get_noise(batch_size, n_noise)
m, loss_val_D, _ = sess.run([merged, loss_D, train_D],
feed_dict={X: batch_xs, Y: batch_ys, Z: noise})
loss_val_G, _ = sess.run([loss_G, train_G],
feed_dict={Y: batch_ys, Z: noise})
writer.add_summary(m, i + epoch * total_batch)
# _, _, loss_val_G, loss_val_D = sess.run([train_G, train_D, loss_G, loss_D],
# feed_dict={X: batch_xs, Y: batch_ys, Z: noise})
print('Epoch:', '%04d' % epoch,
'D loss: {:.4}'.format(loss_val_D),
'G loss: {:.4}'.format(loss_val_G))
if epoch == 0 or (epoch + 1) % 10 == 0:
sample_size = 10
noise = get_noise(sample_size, n_noise)
samples = sess.run(G,
feed_dict={Y: mnist.test.labels[:sample_size],
Z: noise})
fig, ax = plt.subplots(2, sample_size, figsize=(sample_size, 2))
for i in range(sample_size):
ax[0][i].set_axis_off()
ax[1][i].set_axis_off()
ax[0][i].imshow(np.reshape(mnist.test.images[i], (28, 28)))
ax[1][i].imshow(np.reshape(samples[i], (28, 28)))
plt.savefig('samples2/{}.png'.format(str(epoch).zfill(3)), bbox_inches='tight')
plt.close(fig)
print('최적화 완료!')
# -
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
| 3minTensorflow/08. GAN/08. GAN2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch-0.4
# language: python
# name: pytorch-0.4
# ---
# +
import numpy as np
import random
import torch as pt
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
# %load_ext autoreload
# %autoreload 2
pt.set_printoptions(linewidth=200)
# -
device = pt.device("cuda:0" if pt.cuda.is_available() else "cpu")
hidden_size = 50
# + code_folding=[]
class DinosDataset(Dataset):
def __init__(self):
super().__init__()
with open('dinos.txt') as f:
content = f.read().lower()
self.vocab = sorted(set(content))
self.vocab_size = len(self.vocab)
self.lines = content.splitlines()
self.ch_to_idx = {c:i for i, c in enumerate(self.vocab)}
self.idx_to_ch = {i:c for i, c in enumerate(self.vocab)}
def __getitem__(self, index):
line = self.lines[index]
x_str = ' ' + line #add a space at the beginning, which indicates a vector of zeros.
y_str = line + '\n'
x = pt.zeros([len(x_str), self.vocab_size], dtype=pt.float)
y = pt.empty(len(x_str), dtype=pt.long)
y[0] = self.ch_to_idx[y_str[0]]
#we start from the second character because the first character of x was nothing(vector of zeros).
for i, (x_ch, y_ch) in enumerate(zip(x_str[1:], y_str[1:]), 1):
x[i][self.ch_to_idx[x_ch]] = 1
y[i] = self.ch_to_idx[y_ch]
return x, y
def __len__(self):
return len(self.lines)
# -
trn_ds = DinosDataset()
trn_dl = DataLoader(trn_ds, batch_size=1, shuffle=True)
# + code_folding=[]
class LSTM(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(LSTM, self).__init__()
self.linear_f = nn.Linear(input_size + hidden_size, hidden_size)
self.linear_u = nn.Linear(input_size + hidden_size, hidden_size)
self.linear_c = nn.Linear(input_size + hidden_size, hidden_size)
self.linear_o = nn.Linear(input_size + hidden_size, hidden_size)
self.i2o = nn.Linear(hidden_size, output_size)
def forward(self, c_prev, h_prev, x):
combined = pt.cat([x, h_prev], 1)
f = pt.sigmoid(self.linear_f(combined))
u = pt.sigmoid(self.linear_u(combined))
c_tilde = pt.tanh(self.linear_c(combined))
c = f*c_prev + u*c_tilde
o = pt.sigmoid(self.linear_o(combined))
h = o*pt.tanh(c)
y = self.i2o(h)
return c, h, y
# -
model = LSTM(trn_ds.vocab_size, hidden_size, trn_ds.vocab_size).to(device)
loss_fn = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-2)
# + code_folding=[]
def print_sample(sample_idxs):
print(trn_ds.idx_to_ch[sample_idxs[0]].upper(), end='')
[print(trn_ds.idx_to_ch[x], end='') for x in sample_idxs[1:]]
# + code_folding=[]
def sample(model):
model.eval()
c_prev = pt.zeros([1, hidden_size], dtype=pt.float, device=device)
h_prev = pt.zeros_like(c_prev)
x = c_prev.new_zeros([1, trn_ds.vocab_size])
sampled_indexes = []
idx = -1
n_chars = 1
newline_char_idx = trn_ds.ch_to_idx['\n']
with pt.no_grad():
while n_chars != 50 and idx != newline_char_idx:
c_prev, h_prev, y_pred = model(c_prev, h_prev, x)
softmax_scores = pt.softmax(y_pred, 1).cpu().numpy().ravel()
np.random.seed(np.random.randint(1, 5000))
idx = np.random.choice(np.arange(trn_ds.vocab_size), p=softmax_scores)
sampled_indexes.append(idx)
x = (y_pred == y_pred.max(1)[0]).float()
n_chars += 1
if n_chars == 50:
sampled_indexes.append(newline_char_idx)
model.train()
return sampled_indexes
# + code_folding=[]
def train_one_epoch(model, loss_fn, optimizer):
model.train()
for line_num, (x, y) in enumerate(trn_dl):
loss = 0
optimizer.zero_grad()
c_prev = pt.zeros([1, hidden_size], dtype=pt.float, device=device)
h_prev = pt.zeros_like(c_prev)
x, y = x.to(device), y.to(device)
for i in range(x.shape[1]):
c_prev, h_prev, y_pred = model(c_prev, h_prev, x[:, i])
loss += loss_fn(y_pred, y[:, i])
if (line_num+1) % 100 == 0:
print_sample(sample(model))
loss.backward()
optimizer.step()
# + code_folding=[]
def train(model, loss_fn, optimizer, dataset='dinos', epochs=1):
for e in range(1, epochs+1):
print(f'{"-"*20} Epoch {e} {"-"*20}')
train_one_epoch(model, loss_fn, optimizer)
# -
train(model, loss_fn, optimizer, epochs=3)
| 5- Sequence Models/Week 1/Dinosaur Island -- Character-level language model/Dinosaur Island LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/The-Gupta/Time/blob/main/01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Kz_A59-oqFV8"
# 20JUN21, SUN
# [Public] Location: https://drive.google.com/drive/folders/1fFoFxY4i3YU-KNkz84_AG9Nftxr4Alnb
# + colab={"base_uri": "https://localhost:8080/"} id="3tHOP6aBqJQG" outputId="b9dd1749-1fc7-4475-d030-8c5ead8910ef"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="4HUYwKvfEuQb"
# + colab={"base_uri": "https://localhost:8080/", "height": 894} id="nONIWmjoEa9n" outputId="451fac69-debb-4830-a079-3f3ecc52c5ef"
# !pip install pyLDAvis
# + [markdown] id="6GiyflCaEw1D"
# Click 'RESTART RUNTIME', and then 'Yes'
# + colab={"base_uri": "https://localhost:8080/"} id="GL_LF6LuqJSD" outputId="0680596f-bb36-4cf9-90a4-8afdc2272820"
# cd "gdrive/MyDrive/Time Magazine"
# + id="WwQVPbeUqJWa"
# + colab={"base_uri": "https://localhost:8080/"} id="_l8H4fDlw3pW" outputId="ec7fe15c-aa45-4acd-d4e6-8966d6ab2832"
import re, os, ast, time, requests, urllib.request
from datetime import datetime
from pytz import timezone
# !sudo apt install tesseract-ocr
# !pip install pytesseract
import pytesseract
pytesseract.pytesseract.tesseract_cmd = (r'/usr/bin/tesseract')
try:
from PIL import Image
except ImportError:
import Image
import pytesseract, cv2, os
import cv2, random, itertools
from google.colab.patches import cv2_imshow
import numpy as np
from sklearn.cluster import DBSCAN
# !python -m spacy download en_core_web_sm
import spacy
nlp = spacy.load('en_core_web_sm')
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
stopword_set = set(stopwords.words('english'))
from __future__ import print_function
import pyLDAvis
import pyLDAvis.sklearn
pyLDAvis.enable_notebook()
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.decomposition import LatentDirichletAllocation
# + id="ooFrIFrxEKuY"
# + [markdown] id="6yOCc6-8rGWk"
# # Scraping [Time Magazine](https://time.com/vault/)
# 1923 - 2015
# + id="R8SQnOciqJYV"
def get_issue_dates(year):
url = 'https://time.com/vault/year/{}/'.format(year)
page = requests.get(url)
issue_dates = re.findall('\"coverDate\":\"(\d\d\d\d-\d\d-\d\d)\"', page.text)
return issue_dates
def download_page_image(url, path):
try:
urllib.request.urlretrieve(url, path)
return True
except Exception as e:
print('\n\tURL:', url, '\n\tPath:', path)
print('\t', e)
return False
def download_issue(issue_date):
year, month, day = issue_date.split('-')
# Find Number of Pages
url = 'https://time.com/vault/issue/{}/page/1/'.format(issue_date)
page = requests.get(url)
page_count = len(ast.literal_eval(re.findall('\"pagePDFs\":(\[[^\[\]]*\])', page.text)[0]))
# Create Issue Directory
if not os.path.exists(year+'/'+issue_date): os.mkdir(year+'/'+issue_date)
# Download Pages
for page_number in range(1, page_count+1):
page_url = 'https://content.time.com/time/subscriber/vault/{}/{}/{}/{}/1550.jpg'.format(year, month, year+month+day, page_number)
# urllib.request.urlretrieve(page_url, '{}/{}/{}.jpg'.format(year, issue_date, page_number))
count = 0
success = download_page_image(page_url, '{}/{}/{}.jpg'.format(year, issue_date, page_number))
while not success and count < 3:
success = download_page_image(page_url, '{}/{}/{}.jpg'.format(year, issue_date, page_number))
time.sleep(1)
count += 1
def get_timestamp():
now_utc = datetime.now(timezone('UTC'))
return now_utc.astimezone(timezone('Asia/Kolkata')).strftime("[%d-%m-%Y %H:%M:%S]")
# + id="V71Hxn3YqJbD"
# %%time
for year in list(range(1923, 2021+1)):
year = str(year)
# Create year Directory
if not os.path.exists(year): os.mkdir(year)
issue_dates = get_issue_dates(year)
for issue_date in issue_dates:
download_issue(issue_date)
print('\n{} Completed: {}.'.format(get_timestamp(), year))
# + id="vjNl8HOCqJeE"
# + [markdown] id="B9dlKXepr5P0"
# # Text Extracation (2000-2015)
# + id="Uk4d_EmkqJje" colab={"base_uri": "https://localhost:8080/"} outputId="31512116-016e-4dae-f905-6123a5f4b3fb"
for year in range(2000, 2016):
year = str(year)
for path,subdirs,files in os.walk(year):
dir_files = os.listdir(path)
print(path)
for name in files:
if name.endswith('.jpg') and name[:-4]+'.txt' not in dir_files:
img_path = os.path.join(path, name)
image = cv2.imread(img_path)
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY) # Convert to Black and White
gray = cv2.resize(gray, None, fx = 3, fy = 3, interpolation = cv2.INTER_CUBIC) # Zoom 3X
text = pytesseract.image_to_string(gray) # OCR
# Save text
f = open(img_path[:-4]+'.txt', 'w')
f.write(text)
f.close()
# + id="hjlWRRmKqJmI"
# + [markdown] id="sYqOxGMr9tLe"
# # What is on the cover page?
# The most Frequent Terms
# + colab={"base_uri": "https://localhost:8080/"} id="tJJE2-WdqJrK" outputId="05ebcfbf-99a1-45c2-bc0a-672cfade0f67"
try: from unidecode import unidecode
except:
# !pip install unidecode
from unidecode import unidecode
def read_text_file(filename):
with open(filename) as f:
content = ' '.join([unidecode(i.strip()) for i in f.readlines() if i.strip()]).replace('- ', '')
return content
def get_magazine_text(path):
pages = sorted([os.path.join(path, file) for file in os.listdir(path) if file.endswith('.txt')], key=lambda x: int(x.split('/')[-1][:-4]))
texts = [read_text_file(page) for page in pages]
return texts
# + id="dCzBALwYyObI"
release_dates = sum([[os.path.join(str(year), dir_) for dir_ in os.listdir(str(year)) if '.' not in dir_] for year in range(2000, 2016)], [])
# + id="EEDFs-oW0E7H"
texts = []
for release_date in release_dates:
# print('\n\n\n', release_date)
text = read_text_file(release_date+'/1.txt')
texts.append(text)
# print(text)
# doc = nlp(text)
# for entity in doc.ents:
# print(entity.label_, ' | ', entity.text)
# + id="EpHDI5QO7gEq" colab={"base_uri": "https://localhost:8080/"} outputId="3678b235-527b-4af1-d98b-b016742b4e52"
from collections import Counter
l = ' '.join(texts).lower().split()
l = [i for i in l if len(i)>=3 and i not in stopword_set]
c = Counter(l)
c.most_common()
# + [markdown] id="bcek5cWK-VOA"
# ### Looks like it's mostly about the US and Geopolitics.
# + colab={"background_save": true} id="LkDnXVs3qJ7c"
# + colab={"base_uri": "https://localhost:8080/"} id="HwZ4bj5mV7VB" outputId="cd035ddd-3806-483e-9b59-2ec779eb77a4"
# Save all texts in a Pickle File (for quick retreval).
# texts = {}
# count = 0
# for release_date in release_dates:
# print(release_date)
# for file_ in os.listdir(release_date):
# count += 1
# texts[os.path.join(release_date, file_)] = (count, get_magazine_text(release_date))
# import pickle
# with open('time_magazine_texts.pickle', 'wb') as f:
# pickle.dump(texts, f, protocol=pickle.HIGHEST_PROTOCOL)
# with open('time_magazine_texts.pickle', 'rb') as f:
# texts = pickle.load(f)
# + [markdown] id="cYeG9KN1-qmp"
# # Let's try to find some topics and their dominant terms
# + colab={"base_uri": "https://localhost:8080/"} id="XUJzBBkgCIM-" outputId="6407a37f-684f-4a67-9b2f-3dacc47e8d23"
len(release_dates)
# + id="QTbp9tUrCIPv"
# + id="2ckHX0JP4uls"
# Let's see for a year first.
# + id="8LEx-hq8BSNH" colab={"base_uri": "https://localhost:8080/"} outputId="27e76475-0c8a-4014-9ec7-b43b8f452f27"
# %%time
l = [get_magazine_text(release_date) for release_date in release_dates[:1]]
pages = [item for sublist in l for item in sublist]
# + id="HEV90fdKAjeX"
n_components = 5 # Number of topics
# + colab={"base_uri": "https://localhost:8080/", "height": 878} id="usvsVV5BqJ-F" outputId="5d39dea7-d80e-4f27-fa90-a7ca92169635"
tf_vectorizer = CountVectorizer(strip_accents = 'unicode',
stop_words = 'english',
lowercase = True,
token_pattern = r'\b[a-zA-Z]{3,}\b',
max_df = 0.5,
min_df = 10)
dtm_tf = tf_vectorizer.fit_transform(pages)
print(dtm_tf.shape)
# for TF DTM
lda_tf = LatentDirichletAllocation(n_components=n_components, random_state=0)
lda_tf.fit(dtm_tf)
pyLDAvis.sklearn.prepare(lda_tf, dtm_tf, tf_vectorizer)
# + colab={"base_uri": "https://localhost:8080/", "height": 878} id="g01NHeccqKAf" outputId="8590ecc7-ef52-4384-853f-25f55215f9e2"
tfidf_vectorizer = TfidfVectorizer(strip_accents = 'unicode',
stop_words = 'english',
lowercase = True,
token_pattern = r'\b[a-zA-Z]{3,}\b',
max_df = 0.5,
min_df = 10)
dtm_tfidf = tfidf_vectorizer.fit_transform(pages)
print(dtm_tfidf.shape)
# for TFIDF DTM
lda_tfidf = LatentDirichletAllocation(n_components=n_components, random_state=0)
lda_tfidf.fit(dtm_tfidf)
pyLDAvis.sklearn.prepare(lda_tfidf, dtm_tfidf, tfidf_vectorizer)
# + id="iB-Ayy2FqKDF"
# TODO: For all the issues, and interpret
# + id="BtNwJGCHqKF2"
# + id="TO0-N0HHDY2c"
# + id="Op1Q7L4EDY5O"
| 01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.3
# language: julia
# name: julia-1.7
# ---
# Before running this, please make sure to activate and instantiate the
# tutorial-specific package environment, using this
# [`Project.toml`](https://raw.githubusercontent.com/juliaai/DataScienceTutorials.jl/gh-pages/__generated/A-composing-models/Project.toml) and
# [this `Manifest.toml`](https://raw.githubusercontent.com/juliaai/DataScienceTutorials.jl/gh-pages/__generated/A-composing-models/Manifest.toml), or by following
# [these](https://juliaai.github.io/DataScienceTutorials.jl/#learning_by_doing) detailed instructions.
# ## Generating dummy data
# Let's start by generating some dummy data with both numerical values and categorical values:
# +
using MLJ
using PrettyPrinting
KNNRegressor = @load KNNRegressor
# -
# input
X = (age = [23, 45, 34, 25, 67],
gender = categorical(['m', 'm', 'f', 'm', 'f']))
# target
height = [178, 194, 165, 173, 168];
# Note that the scientific type of `age` is `Count` here:
scitype(X.age)
# We will want to coerce that to `Continuous` so that it can be given to a regressor that expects such values.
# ## Declaring a pipeline
# A typical workflow for such data is to one-hot-encode the categorical data and then apply some regression model on the data.
# Let's say that we want to apply the following steps:
# 1. One hot encode the categorical features in `X`
# 1. Standardize the target variable (`:height`)
# 1. Train a KNN regression model on the one hot encoded data and the Standardized target.
# The `Pipeline` constructor helps you define such a simple (non-branching) pipeline of steps to be applied in order:
pipe = Pipeline(
coercer = X -> coerce(X, :age=>Continuous),
one_hot_encoder = OneHotEncoder(),
transformed_target_model = TransformedTargetModel(
model = KNNRegressor(K=3);
target=UnivariateStandardizer()
)
)
# Note the coercion of the `:age` variable to Continuous since `KNNRegressor` expects `Continuous` input.
# Note also the `TransformedTargetModel` which allows one to learn a transformation (in this case Standardization) of the
# target variable to be passed to the `KNNRegressor`.
# Hyperparameters of this pipeline can be accessed (and set) using dot syntax:
pipe.transformed_target_model.model.K = 2
pipe.one_hot_encoder.drop_last = true;
# Evaluation for a pipe can be done with the `evaluate!` method; implicitly it will construct machines that will contain the fitted parameters etc:
evaluate(
pipe,
X,
height,
resampling=Holdout(),
measure=rms
) |> pprint
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| __site/__generated/A-composing-models/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
from astropy.coordinates import SkyCoord
from astropy import units as u
# Read in Issacson 2017 target table:
f = pd.read_csv("../../bl/Isaacson2017.csv")
#k=f.loc[np.where(f['Dist']<=5.0)[0]]
#k
missing=('HIP443' ,'HIP1242', 'HIP2081', 'HIP2762', 'HIP8903', 'HIP11452', 'HIP12351', \
'HIP14576', 'HIP16134', 'HIP16852', 'HIP23875', 'HIP35296', 'GJ273', 'HIP36349',\
'HIP42913', 'HIP46706', 'HIP48659', 'HIP49973', 'HIP51986', 'HIP52190',\
'HIP54182', 'HIP54872', 'HIP65378', 'HIP67301', 'GJ559B', 'GJ559A', 'HIP72622',\
'HIP77516', 'HIP81919', 'HIP82817', 'HIP84123', 'HIP88601', 'GJ702A', 'GJ702B',\
'HIP95501', 'HIP98204', 'HIP101955', 'HIP108752', 'HIP109268', 'HIP109821',\
'HIP113963', 'HIP89937')
k=pd.DataFrame(columns=list(f))
k
for i in range(len(missing)):
k=k.append(f.loc[np.where(f['SimbadName']==missing[i])[0]])
k=k.reset_index(drop=True)
k
# Pick the ones in the MeerKAT FOV
k=k.loc[np.where(k['Dec'].values<45.)[0]]
k=k.reset_index(drop=True)
k
# Add a note for interesting targets:
# Most of these don't have Gaia id's because they're too bright
p=f.rename(columns={'SimbadName':'simbadname'})
p['note'] = pd.Series('NaN', index=p.index)
p.loc[np.where(p['simbadname']=='GJ244A')[0],'note']='Sirius A'
p.loc[np.where(p['simbadname']=='HIP36850')[0],'note']='Castor AB'
p.loc[np.where(p['simbadname']=='GJ280')[0],'note']='Procyon'
p.loc[np.where(p['simbadname']=='HIP37826')[0],'note']='Pollox'
p.loc[np.where(p['simbadname']=='HIP54061')[0],'note']='Dubhe'
p.loc[np.where(p['simbadname']=='HIP62956')[0],'note']='Alioth'
p.loc[np.where(p['simbadname']=='HIP76267')[0],'note']='Alphecca'
p.loc[np.where(p['simbadname']=='HIP84012')[0],'note']='eta Oph'
p.loc[np.where(p['simbadname']=='HIP86032')[0],'note']='alp Oph'
p.loc[np.where(p['simbadname']=='HIP91262')[0],'note']='alp Lyr'
p.loc[np.where(p['simbadname']=='HIP93506')[0],'note']='zet Sgr'
p.loc[np.where(p['simbadname']=='HIP97649')[0],'note']='Altair'
p.loc[np.where(p['simbadname']=='GJ768')[0],'note']='Altair'
p.loc[np.where(p['simbadname']=='HIP105199')[0],'note']='Alderamin'
p.loc[np.where(p['simbadname']=='HIP113368')[0],'note']='Fomalhaut'
p.loc[np.where(p['simbadname']=='GJ551')[0],'note']='Proxima Centauri'
p.loc[np.where(p['simbadname']=='GJ406')[0],'note']='Wolf 359'
p.loc[np.where(p['simbadname']=='GJ699')[0],'note']='Barnards star'
p.loc[np.where(p['simbadname']=='GJ411')[0],'note']='Lalande 21185'
#p.loc[np.where(p['source_id']==4075141768785646848)[0][0],'note']='Ross 154'
p.loc[np.where(p['simbadname']=='GJ905')[0],'note']='Ross 248'
p.loc[np.where(p['simbadname']=='GJ144')[0],'note']='Epsilon Eridani'
p=p.loc[np.where(p['note']!='NaN')[0]]
p=p.reset_index(drop=True)
p=p.rename(columns={'simbadname':'SimbadName'})
p=p.loc[np.where(p['Dec']<45.)[0]]
p=p.reset_index(drop=True)
p
k['note']='NaN'
tot=pd.concat([p,k],sort=False)
tot=tot.reset_index(drop=True)
tot
dupes = tot.duplicated(subset='SimbadName',keep='first')
tot = tot.loc[np.where(dupes==False)[0]]
tot=tot.reset_index(drop=True)
tot
# Update to J2015.5 ref epoch
# Convert the target list proper motions to degrees/yr
pmRA_deg,pmDE_deg = (tot['pmRA'].values*u.arcsec).to(u.deg).value,(tot['pmDE'].values*u.arcsec).to(u.deg).value
# Compute the difference between the ref epoch and updated epoch:
deltat = 2015.5-2000.
# Add the delta RA/Dec to the target list RA/Dec:
RA_J20155 = tot['RA'].values+(pmRA_deg*deltat)
Dec_J20155 = tot['Dec'].values+(pmDE_deg*deltat)
tot['ra'],tot['decl']=RA_J20155,Dec_J20155
tot['ep']='2015.5'
tot
l=pd.read_csv('Our_targets/fornax.csv')
cols=list(l)
print list(tot)
print cols
df = pd.DataFrame(index=range(len(tot)),columns=cols)
df['dist.c'],df['name']=tot['Dist'],tot['SimbadName']
df['ra'],df['decl'],df['ref_epoch']=tot['ra'],tot['decl'],tot['ep']
df['pmra'],df['pmdec']=tot['pmRA'],tot['pmDE']
df['sptype.c']=tot['SpType']
df['phot_g_mean_mag']=tot['Vmag']
df['project']='Not in Gaia'
df
df.to_csv('Our_targets/not_in_gaia.csv',index=False)
| MeerKAT_LSPs/Add close sources not in Gaia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Iterative methods for solving Ax = b</h2>
# The conjugate gradient method (when A is symmetric positive definite, or SPD) and the Jacobi method.
#
# +
# These are the standard imports for CS 111.
# This list may change as the quarter goes on.
import os
import time
import math
import numpy as np
import numpy.linalg as npla
import scipy
from scipy import sparse
from scipy import linalg
import scipy.sparse.linalg as spla
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import axes3d
import cs111
# %matplotlib tk
# -
def Jsolve(A, b, tol = 1e-8, max_iters = 1000, callback = None):
"""Solve a linear system Ax = b for x by the Jacobi iterative method.
Parameters:
A: the matrix.
b: the right-hand side vector.
tol = 1e-8: the relative residual at which to stop iterating.
max_iters = 1000: the maximum number of iterations to do.
callback = None: a user function to call at every iteration.
The callback function has arguments 'x', 'iteration', and 'residual'
Outputs (in order):
x: the computed solution
rel_res: list of relative residual norms at each iteration.
The number of iterations actually done is len(rel_res) - 1
"""
# Check the input
m, n = A.shape
assert m == n, "matrix must be square"
bn, = b.shape
assert bn == n, "rhs vector must be same size as matrix"
# Split A into diagonal D plus off-diagonal C
d = A.diagonal() # diagonal elements of A as a vector
C = A.copy() # copy of A ...
C.setdiag(np.zeros(n)) # ... without the diagonal
# Initial guess: x = 0
x = np.zeros(n)
# Vector of relative residuals
# Relative residual is norm(residual)/norm(b)
# Intitial residual is b - Ax for x=0, or b
rel_res = [1.0]
# Call user function if specified
if callback is not None:
callback(x = x, iteration = 0, residual = 1)
# Iterate
for k in range(1, max_iters+1):
# New x
x = (b - C @ x) / d
# Record relative residual
this_rel_res = npla.norm(b - A @ x) / npla.norm(b)
rel_res.append(this_rel_res)
# Call user function if specified
if callback is not None:
callback(x = x, iteration = k, residual = this_rel_res)
# Stop if within tolerance
if this_rel_res <= tol:
break
return (x, rel_res)
def CGsolve(A, b, tol = 1e-8, max_iters = 1000, callback = None):
"""Solve a linear system Ax = b for x by the conjugate gradient iterative method.
Parameters:
A: the matrix.
b: the right-hand side vector.
tol = 1e-8: the relative residual at which to stop iterating.
max_iters = 1000: the maximum number of iterations to do.
callback = None: a user function to call at every iteration, with one argument x
Outputs (in order):
x: the computed solution
rel_res: list of relative residual norms at each iteration.
The number of iterations actually done is len(rel_res) - 1
"""
# Check the input
m, n = A.shape
assert m == n, "matrix must be square"
bn, = b.shape
assert bn == n, "rhs vector must be same size as matrix"
# Initial guess: x = 0
x = np.zeros(n)
# Initial residual: r = b - A@0 = b
r = b
# Initial step is in direction of residual.
d = r
# Squared norm of residual
rtr = r.T @ r
# Vector of relative residuals
# Relative residual is norm(residual)/norm(b)
# Intitial residual is b - Ax for x=0, or b
rel_res = [1.0]
# Call user function if specified
if callback is not None:
callback(x = x, iteration = 0, residual = 1)
# Iterate
for k in range(1, max_iters+1):
Ad = A @ d
alpha = rtr / (d.T @ Ad) # Length of step
x = x + alpha * d # Update x to new x
r = r - alpha * Ad # Update r to new residual
rtrold = rtr
rtr = r.T @ r
beta = rtr / rtrold
d = r + beta * d # Update d to new step direction
# Record relative residual
this_rel_res = npla.norm(b - A @ x) / npla.norm(b)
rel_res.append(this_rel_res)
# Call user function if specified
if callback is not None:
callback(x = x, iteration = k, residual = this_rel_res)
# Stop if within tolerance
if this_rel_res <= tol:
break
return (x, rel_res)
| 00.AllPythonCode/CGandJacobiSolve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dlmacedo/starter-academic/blob/master/content/courses/deeplearning/notebooks/pytorch/torchvision_finetuning_instance_segmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DfPPQ6ztJhv4"
# # TorchVision Instance Segmentation Finetuning Tutorial
#
# For this tutorial, we will be finetuning a pre-trained [Mask R-CNN](https://arxiv.org/abs/1703.06870) model in the [*Penn-Fudan Database for Pedestrian Detection and Segmentation*](https://www.cis.upenn.edu/~jshi/ped_html/). It contains 170 images with 345 instances of pedestrians, and we will use it to illustrate how to use the new features in torchvision in order to train an instance segmentation model on a custom dataset.
#
# First, we need to install `pycocotools`. This library will be used for computing the evaluation metrics following the COCO metric for intersection over union.
# + id="DBIoe_tHTQgV" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="55b8ee3b-ba36-4941-e140-768c75fd8e54"
# %%shell
pip install cython
# Install pycocotools, the version by default in Colab
# has a bug fixed in https://github.com/cocodataset/cocoapi/pull/354
pip install -U 'git+https://github.com/cocodataset/cocoapi.git#subdirectory=PythonAPI'
# + [markdown] id="5Sd4jlGp2eLm"
# ## Defining the Dataset
#
# The [torchvision reference scripts for training object detection, instance segmentation and person keypoint detection](https://github.com/pytorch/vision/tree/v0.3.0/references/detection) allows for easily supporting adding new custom datasets.
# The dataset should inherit from the standard `torch.utils.data.Dataset` class, and implement `__len__` and `__getitem__`.
#
# The only specificity that we require is that the dataset `__getitem__` should return:
#
# * image: a PIL Image of size (H, W)
# * target: a dict containing the following fields
# * `boxes` (`FloatTensor[N, 4]`): the coordinates of the `N` bounding boxes in `[x0, y0, x1, y1]` format, ranging from `0` to `W` and `0` to `H`
# * `labels` (`Int64Tensor[N]`): the label for each bounding box
# * `image_id` (`Int64Tensor[1]`): an image identifier. It should be unique between all the images in the dataset, and is used during evaluation
# * `area` (`Tensor[N]`): The area of the bounding box. This is used during evaluation with the COCO metric, to separate the metric scores between small, medium and large boxes.
# * `iscrowd` (`UInt8Tensor[N]`): instances with `iscrowd=True` will be ignored during evaluation.
# * (optionally) `masks` (`UInt8Tensor[N, H, W]`): The segmentation masks for each one of the objects
# * (optionally) `keypoints` (`FloatTensor[N, K, 3]`): For each one of the `N` objects, it contains the `K` keypoints in `[x, y, visibility]` format, defining the object. `visibility=0` means that the keypoint is not visible. Note that for data augmentation, the notion of flipping a keypoint is dependent on the data representation, and you should probably adapt `references/detection/transforms.py` for your new keypoint representation
#
# If your model returns the above methods, they will make it work for both training and evaluation, and will use the evaluation scripts from pycocotools.
#
# Additionally, if you want to use aspect ratio grouping during training (so that each batch only contains images with similar aspect ratio), then it is recommended to also implement a `get_height_and_width` method, which returns the height and the width of the image. If this method is not provided, we query all elements of the dataset via `__getitem__` , which loads the image in memory and is slower than if a custom method is provided.
#
# + [markdown] id="bX0rqK-A3Nbl"
# ### Writing a custom dataset for Penn-Fudan
#
# Let's write a dataset for the Penn-Fudan dataset.
#
# First, let's download and extract the data, present in a zip file at https://www.cis.upenn.edu/~jshi/ped_html/PennFudanPed.zip
# + id="_t4TBwhHTdkd" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1b209271-3c3b-4df9-ce4d-2176edb57666"
# %%shell
# download the Penn-Fudan dataset
wget https://www.cis.upenn.edu/~jshi/ped_html/PennFudanPed.zip .
# extract it in the current folder
unzip PennFudanPed.zip
# + [markdown] id="WfwuU-jI3j93"
# Let's have a look at the dataset and how it is layed down.
#
# The data is structured as follows
# ```
# PennFudanPed/
# PedMasks/
# FudanPed00001_mask.png
# FudanPed00002_mask.png
# FudanPed00003_mask.png
# FudanPed00004_mask.png
# ...
# PNGImages/
# FudanPed00001.png
# FudanPed00002.png
# FudanPed00003.png
# FudanPed00004.png
# ```
#
# Here is one example of an image in the dataset, with its corresponding instance segmentation mask
# + id="LDjuVFgexFfh" colab={"base_uri": "https://localhost:8080/", "height": 553} outputId="23c18cf5-dc2f-43c0-b445-8abed4d005e5"
from PIL import Image
Image.open('PennFudanPed/PNGImages/FudanPed00001.png')
# + id="cFHKCvCTxiff" colab={"base_uri": "https://localhost:8080/", "height": 553} outputId="4f5b1d6c-96da-4a22-da36-515db9c9049d"
mask = Image.open('PennFudanPed/PedMasks/FudanPed00001_mask.png')
# each mask instance has a different color, from zero to N, where
# N is the number of instances. In order to make visualization easier,
# let's adda color palette to the mask.
mask.putpalette([
0, 0, 0, # black background
255, 0, 0, # index 1 is red
255, 255, 0, # index 2 is yellow
255, 153, 0, # index 3 is orange
])
mask
# + [markdown] id="C9Ee5NV54Dmj"
# So each image has a corresponding segmentation mask, where each color correspond to a different instance. Let's write a `torch.utils.data.Dataset` class for this dataset.
# + id="mTgWtixZTs3X"
import os
import numpy as np
import torch
import torch.utils.data
from PIL import Image
class PennFudanDataset(torch.utils.data.Dataset):
def __init__(self, root, transforms=None):
self.root = root
self.transforms = transforms
# load all image files, sorting them to
# ensure that they are aligned
self.imgs = list(sorted(os.listdir(os.path.join(root, "PNGImages"))))
self.masks = list(sorted(os.listdir(os.path.join(root, "PedMasks"))))
def __getitem__(self, idx):
# load images ad masks
img_path = os.path.join(self.root, "PNGImages", self.imgs[idx])
mask_path = os.path.join(self.root, "PedMasks", self.masks[idx])
img = Image.open(img_path).convert("RGB")
# note that we haven't converted the mask to RGB,
# because each color corresponds to a different instance
# with 0 being background
mask = Image.open(mask_path)
mask = np.array(mask)
# instances are encoded as different colors
obj_ids = np.unique(mask)
# first id is the background, so remove it
obj_ids = obj_ids[1:]
# split the color-encoded mask into a set
# of binary masks
masks = mask == obj_ids[:, None, None]
# get bounding box coordinates for each mask
num_objs = len(obj_ids)
boxes = []
for i in range(num_objs):
pos = np.where(masks[i])
xmin = np.min(pos[1])
xmax = np.max(pos[1])
ymin = np.min(pos[0])
ymax = np.max(pos[0])
boxes.append([xmin, ymin, xmax, ymax])
boxes = torch.as_tensor(boxes, dtype=torch.float32)
# there is only one class
labels = torch.ones((num_objs,), dtype=torch.int64)
masks = torch.as_tensor(masks, dtype=torch.uint8)
image_id = torch.tensor([idx])
area = (boxes[:, 3] - boxes[:, 1]) * (boxes[:, 2] - boxes[:, 0])
# suppose all instances are not crowd
iscrowd = torch.zeros((num_objs,), dtype=torch.int64)
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["masks"] = masks
target["image_id"] = image_id
target["area"] = area
target["iscrowd"] = iscrowd
if self.transforms is not None:
img, target = self.transforms(img, target)
return img, target
def __len__(self):
return len(self.imgs)
# + [markdown] id="J6f3ZOTJ4Km9"
# That's all for the dataset. Let's see how the outputs are structured for this dataset
# + id="ZEARO4B_ye0s" colab={"base_uri": "https://localhost:8080/", "height": 331} outputId="006161f7-ddad-4ff4-8fb1-b91d0fd81069"
dataset = PennFudanDataset('PennFudanPed/')
dataset[0]
# + [markdown] id="lWOhcsir9Ahx"
# So we can see that by default, the dataset returns a `PIL.Image` and a dictionary
# containing several fields, including `boxes`, `labels` and `masks`.
# + [markdown] id="RoAEkUgn4uEq"
# ## Defining your model
#
# In this tutorial, we will be using [Mask R-CNN](https://arxiv.org/abs/1703.06870), which is based on top of [Faster R-CNN](https://arxiv.org/abs/1506.01497). Faster R-CNN is a model that predicts both bounding boxes and class scores for potential objects in the image.
#
# 
#
# Mask R-CNN adds an extra branch into Faster R-CNN, which also predicts segmentation masks for each instance.
#
# 
#
# There are two common situations where one might want to modify one of the available models in torchvision modelzoo.
# The first is when we want to start from a pre-trained model, and just finetune the last layer. The other is when we want to replace the backbone of the model with a different one (for faster predictions, for example).
#
# Let's go see how we would do one or another in the following sections.
#
#
# ### 1 - Finetuning from a pretrained model
#
# Let's suppose that you want to start from a model pre-trained on COCO and want to finetune it for your particular classes. Here is a possible way of doing it:
# ```
# import torchvision
# from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
#
# # load a model pre-trained pre-trained on COCO
# model = torchvision.models.detection.fasterrcnn_resnet50_fpn(pretrained=True)
#
# # replace the classifier with a new one, that has
# # num_classes which is user-defined
# num_classes = 2 # 1 class (person) + background
# # get number of input features for the classifier
# in_features = model.roi_heads.box_predictor.cls_score.in_features
# # replace the pre-trained head with a new one
# model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
# ```
#
# ### 2 - Modifying the model to add a different backbone
#
# Another common situation arises when the user wants to replace the backbone of a detection
# model with a different one. For example, the current default backbone (ResNet-50) might be too big for some applications, and smaller models might be necessary.
#
# Here is how we would go into leveraging the functions provided by torchvision to modify a backbone.
#
# ```
# import torchvision
# from torchvision.models.detection import FasterRCNN
# from torchvision.models.detection.rpn import AnchorGenerator
#
# # load a pre-trained model for classification and return
# # only the features
# backbone = torchvision.models.mobilenet_v2(pretrained=True).features
# # FasterRCNN needs to know the number of
# # output channels in a backbone. For mobilenet_v2, it's 1280
# # so we need to add it here
# backbone.out_channels = 1280
#
# # let's make the RPN generate 5 x 3 anchors per spatial
# # location, with 5 different sizes and 3 different aspect
# # ratios. We have a Tuple[Tuple[int]] because each feature
# # map could potentially have different sizes and
# # aspect ratios
# anchor_generator = AnchorGenerator(sizes=((32, 64, 128, 256, 512),),
# aspect_ratios=((0.5, 1.0, 2.0),))
#
# # let's define what are the feature maps that we will
# # use to perform the region of interest cropping, as well as
# # the size of the crop after rescaling.
# # if your backbone returns a Tensor, featmap_names is expected to
# # be [0]. More generally, the backbone should return an
# # OrderedDict[Tensor], and in featmap_names you can choose which
# # feature maps to use.
# roi_pooler = torchvision.ops.MultiScaleRoIAlign(featmap_names=[0],
# output_size=7,
# sampling_ratio=2)
#
# # put the pieces together inside a FasterRCNN model
# model = FasterRCNN(backbone,
# num_classes=2,
# rpn_anchor_generator=anchor_generator,
# box_roi_pool=roi_pooler)
# ```
#
# ### An Instance segmentation model for PennFudan Dataset
#
# In our case, we want to fine-tune from a pre-trained model, given that our dataset is very small. So we will be following approach number 1.
#
# Here we want to also compute the instance segmentation masks, so we will be using Mask R-CNN:
# + id="YjNHjVMOyYlH"
import torchvision
from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
from torchvision.models.detection.mask_rcnn import MaskRCNNPredictor
def get_instance_segmentation_model(num_classes):
# load an instance segmentation model pre-trained on COCO
model = torchvision.models.detection.maskrcnn_resnet50_fpn(pretrained=True)
# get the number of input features for the classifier
in_features = model.roi_heads.box_predictor.cls_score.in_features
# replace the pre-trained head with a new one
model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
# now get the number of input features for the mask classifier
in_features_mask = model.roi_heads.mask_predictor.conv5_mask.in_channels
hidden_layer = 256
# and replace the mask predictor with a new one
model.roi_heads.mask_predictor = MaskRCNNPredictor(in_features_mask,
hidden_layer,
num_classes)
return model
# + [markdown] id="-WXLwePV5ieP"
# That's it, this will make model be ready to be trained and evaluated on our custom dataset.
#
# ## Training and evaluation functions
#
# In `references/detection/,` we have a number of helper functions to simplify training and evaluating detection models.
# Here, we will use `references/detection/engine.py`, `references/detection/utils.py` and `references/detection/transforms.py`.
#
# Let's copy those files (and their dependencies) in here so that they are available in the notebook
# + id="UYDb7PBw55b-" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="fe582468-ef26-46cf-f94e-7066365dd995"
# %%shell
# Download TorchVision repo to use some files from
# references/detection
git clone https://github.com/pytorch/vision.git
# cd vision
git checkout v0.3.0
# cp references/detection/utils.py ../
# cp references/detection/transforms.py ../
# cp references/detection/coco_eval.py ../
# cp references/detection/engine.py ../
# cp references/detection/coco_utils.py ../
# + [markdown] id="2u9e_pdv54nG"
#
#
# Let's write some helper functions for data augmentation / transformation, which leverages the functions in `refereces/detection` that we have just copied:
#
# + id="l79ivkwKy357"
from engine import train_one_epoch, evaluate
import utils
import transforms as T
def get_transform(train):
transforms = []
# converts the image, a PIL image, into a PyTorch Tensor
transforms.append(T.ToTensor())
if train:
# during training, randomly flip the training images
# and ground-truth for data augmentation
transforms.append(T.RandomHorizontalFlip(0.5))
return T.Compose(transforms)
# + [markdown] id="FzCLqiZk-sjf"
# #### Note that we do not need to add a mean/std normalization nor image rescaling in the data transforms, as those are handled internally by the Mask R-CNN model.
# + [markdown] id="3YFJGJxk6XEs"
# ### Putting everything together
#
# We now have the dataset class, the models and the data transforms. Let's instantiate them
# + id="a5dGaIezze3y"
# use our dataset and defined transformations
dataset = PennFudanDataset('PennFudanPed', get_transform(train=True))
dataset_test = PennFudanDataset('PennFudanPed', get_transform(train=False))
# split the dataset in train and test set
torch.manual_seed(1)
indices = torch.randperm(len(dataset)).tolist()
dataset = torch.utils.data.Subset(dataset, indices[:-50])
dataset_test = torch.utils.data.Subset(dataset_test, indices[-50:])
# define training and validation data loaders
data_loader = torch.utils.data.DataLoader(
dataset, batch_size=2, shuffle=True, num_workers=4,
collate_fn=utils.collate_fn)
data_loader_test = torch.utils.data.DataLoader(
dataset_test, batch_size=1, shuffle=False, num_workers=4,
collate_fn=utils.collate_fn)
# + [markdown] id="L5yvZUprj4ZN"
# Now let's instantiate the model and the optimizer
# + id="zoenkCj18C4h" colab={"base_uri": "https://localhost:8080/", "height": 103, "referenced_widgets": ["8ae0425ab601486baf1021fb82f3993f", "<KEY>", "<KEY>", "fef3cafdf7f64c0db94b1747e885d680", "853e12df5db74a7383ab44c64985592c", "8d4fb895c3e446e0b7c801b4dd35cd74", "1d2ab659a33f4f25940b6d34450a52b9", "ffb8a8148ef74bf898602dd334c1b4a9"]} outputId="7b2a5cc3-58ee-43e4-eeef-a39fc76cfe8e"
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# our dataset has two classes only - background and person
num_classes = 2
# get the model using our helper function
model = get_instance_segmentation_model(num_classes)
# move model to the right device
model.to(device)
# construct an optimizer
params = [p for p in model.parameters() if p.requires_grad]
optimizer = torch.optim.SGD(params, lr=0.005,
momentum=0.9, weight_decay=0.0005)
# and a learning rate scheduler which decreases the learning rate by
# 10x every 3 epochs
lr_scheduler = torch.optim.lr_scheduler.StepLR(optimizer,
step_size=3,
gamma=0.1)
# + [markdown] id="XAd56lt4kDxc"
# And now let's train the model for 10 epochs, evaluating at the end of every epoch.
# + id="at-h4OWK0aoc" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="77578320-c07f-4146-ab45-9970f03f6d4e"
# let's train it for 10 epochs
num_epochs = 10
for epoch in range(num_epochs):
# train for one epoch, printing every 10 iterations
train_one_epoch(model, optimizer, data_loader, device, epoch, print_freq=10)
# update the learning rate
lr_scheduler.step()
# evaluate on the test dataset
evaluate(model, data_loader_test, device=device)
# + [markdown] id="Z6mYGFLxkO8F"
# Now that training has finished, let's have a look at what it actually predicts in a test image
# + id="YHwIdxH76uPj"
# pick one image from the test set
img, _ = dataset_test[0]
# put the model in evaluation mode
model.eval()
with torch.no_grad():
prediction = model([img.to(device)])
# + [markdown] id="DmN602iKsuey"
# Printing the prediction shows that we have a list of dictionaries. Each element of the list corresponds to a different image. As we have a single image, there is a single dictionary in the list.
# The dictionary contains the predictions for the image we passed. In this case, we can see that it contains `boxes`, `labels`, `masks` and `scores` as fields.
# + id="Lkmb3qUu6zw3" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="1702bfe2-cdf3-4b59-ad69-205182b3110e"
prediction
# + [markdown] id="RwT21rzotFbH"
# Let's inspect the image and the predicted segmentation masks.
#
# For that, we need to convert the image, which has been rescaled to 0-1 and had the channels flipped so that we have it in `[C, H, W]` format.
# + id="bpqN9t1u7B2J" colab={"base_uri": "https://localhost:8080/", "height": 366} outputId="eff0f296-46b1-40b0-ed32-f342d153930d"
Image.fromarray(img.mul(255).permute(1, 2, 0).byte().numpy())
# + [markdown] id="M58J3O9OtT1G"
# And let's now visualize the top predicted segmentation mask. The masks are predicted as `[N, 1, H, W]`, where `N` is the number of predictions, and are probability maps between 0-1.
# + id="5v5S3bm07SO1" colab={"base_uri": "https://localhost:8080/", "height": 366} outputId="d6b5ba28-1b72-41ca-c935-98544a0204c0"
Image.fromarray(prediction[0]['masks'][0, 0].mul(255).byte().cpu().numpy())
# + [markdown] id="0EZCVtCPunrT"
# Looks pretty good!
#
# ## Wrapping up
#
# In this tutorial, you have learned how to create your own training pipeline for instance segmentation models, on a custom dataset.
# For that, you wrote a `torch.utils.data.Dataset` class that returns the images and the ground truth boxes and segmentation masks. You also leveraged a Mask R-CNN model pre-trained on COCO train2017 in order to perform transfer learning on this new dataset.
#
# For a more complete example, which includes multi-machine / multi-gpu training, check `references/detection/train.py`, which is present in the [torchvision GitHub repo](https://github.com/pytorch/vision/tree/v0.3.0/references/detection).
#
#
| content/courses/deeplearning/notebooks/pytorch/torchvision_finetuning_instance_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="mn00tsObyo9G" outputId="761b2b9f-f55e-412f-9e39-55b57e61fb84"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="frP9tuhBOghy" outputId="59e1c84d-3aaf-4e57-801f-46c12ddada7c"
# %cd '/content/drive/My Drive/DGM'
# !ls -la
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="O-tXbpP5Lx2V" outputId="9838daab-1b4b-4259-900e-7dbbd26b0f2b"
import os
import tensorflow.nn as nn
from tensorflow.keras import Input,Model
from tensorflow.keras.utils import plot_model
from tensorflow.keras import layers, regularizers,activations
from tensorflow.keras.optimizers import SGD,Adam
n_filters = 64
dropout = 0.3
def activation(x):
return nn.swish(x)
def input_block(inp,kernel_resize=5,pad='same'):
# CONV2D + BN + activation
x = layers.Conv2D(n_filters, 3, padding=pad)(inp)
x = layers.BatchNormalization()(x)
x = activation(x)
if not kernel_resize:
return x
# CONV2D (resize) + BN + activation
x1 = layers.Conv2D(n_filters,kernel_resize, padding=pad)(inp)
x1 = layers.BatchNormalization()(x1)
x1 = activation(x1)
x = layers.add([x, x1])
return x
def output_policy_block(x):
policy_head = layers.Conv2D(1, 1, padding='same', use_bias=False)(x)
policy_head = layers.BatchNormalization()(policy_head)
policy_head = activation(policy_head)
policy_head = layers.Flatten()(policy_head)
policy_head = layers.Activation('softmax', name='policy')(policy_head)
return policy_head
def output_value_block(x):
value_head = layers.GlobalAveragePooling2D()(x)
value_head = layers.Dense(n_filters)(value_head)
value_head = layers.BatchNormalization()(value_head)
value_head = activation(value_head)
value_head = layers.Dropout(dropout)(value_head)
value_head = layers.Dense(1, activation='sigmoid', name='value')(value_head)
return value_head
inp = Input(shape=(19,19,31), name='board')
x = input_block(inp)
policy_head = output_policy_block(x)
value_head = output_value_block(x)
model = Model(inputs=inp, outputs=[policy_head, value_head])
file = os.path.join(os.getcwd(),'model_imgs',f'block.png')
plot_model(model,to_file=file,show_shapes=0)
# + [markdown] id="yVRv8OPP8BTU"
# # Some Learning Rate Scheduling
# + [markdown] id="MAM4XHxF5fFW"
#
# + id="cA0G3SZW8AdA" colab={"base_uri": "https://localhost:8080/", "height": 330} outputId="9749ca25-cc66-40b6-9dcb-1e31489cb78d"
import matplotlib.pyplot as plt
from math import cos , pi , floor
lr_min = 3e-7
lr_max = 0.00000575
n_epochs = 12_000
def cosine_annealing(epoch,lr_min=lr_min,lr_max=lr_max,n_epochs=n_epochs, n_cycles=1):
epochs_per_cycle = floor(n_epochs/n_cycles)
cos_inner = (pi * (epoch % epochs_per_cycle)) / (epochs_per_cycle)
lr = lr_min + 0.5 * (lr_max - lr_min) * (1 + cos(cos_inner))
return lr
def cosine_annealing(epoch,lr_min=lr_min,lr_max=lr_max,n_epochs=n_epochs):
return lr_min + 0.5 * (lr_max - lr_min) * (1 + cos(pi*(epoch/n_epochs)))
print(cosine_annealing(10_000))
lr_max_2 = 0.0005
_epochs = range(1,n_epochs+1)
_lr_1 = list(map(cosine_annealing,_epochs))
_lr_2 = list(map(lambda e : cosine_annealing(e,lr_max=lr_max_2),_epochs))
lr_max_3 = 0.000625
_lr_3 = list(map(lambda e : cosine_annealing(e,lr_max=lr_max_3),_epochs))
plt.plot(_epochs,_lr_1,'magenta',label='lr_max = 1e-2')
plt.plot(_epochs,_lr_2,'dodgerblue',label='lr_max = 5e-3')
plt.plot(_epochs,_lr_3,'lime',label='lr_max = 6.25e-3')
plt.xlabel('epochs')
plt.ylabel('lr')
plt.title('LR sheduling for cosine annealing')
plt.legend()
# + id="2L5tKrWNRPM_"
import pickle
import matplotlib.pyplot as plt
LOSS , POLICY_LOSS , VALUE_LOSS , POLICY_ACC , VALUE_MSE = range(5)
def get_metric(d,key):
return [v[key][0] for _,v in d.items()]
def get_metric_validation(val_hist,metric_idx):
return [v[metric_idx] for v in val_hist]
def train_plots(epochs , history):
# metrics = ['loss','policy_loss','value_loss','policy_categorical_accuracy','value_mse']
# Losses
loss = get_metric(history,'loss')
policy_loss = get_metric(history,'policy_loss')
value_loss = get_metric(history,'value_loss')
# Accuracy
policy_acc = get_metric(history,'policy_categorical_accuracy')
# MSE
value_mse = get_metric(history,'value_mse')
fig,(ax_loss,ax_acc,ax_mse) = plt.subplots(1,3)
fig.set_size_inches((16,4))
# Losses
ax_loss.plot(epochs, loss ,'c',label = 'Loss')
ax_loss.plot(epochs, value_loss, 'g' , label = 'Value Loss')
ax_loss.plot(epochs, policy_loss ,'r',label = 'Policy Loss')
ax_loss.set_title('Train Losses History ')
ax_loss.legend()
# Accuracy
ax_acc.plot(epochs, policy_acc , 'r',label = 'Policy Accuracy')
ax_acc.set_title('Train Policy Accuracy History ')
ax_acc.legend()
# MSE
ax_mse.plot(epochs, value_mse , 'g',label = 'Value MSE')
ax_mse.set_title('Train Value MSE History ')
ax_mse.legend()
fig.show()
def validation_plots(epochs , history):
# metrics = ['loss','policy_loss','value_loss','policy_categorical_accuracy','value_mse']
# Losses
loss , policy_loss , value_loss = history['loss'], history['policy_loss'],history['value_loss']
# Accuracy
policy_acc = history['policy_acc']
# MSE
value_mse = history['value_mse']
fig,(ax_loss,ax_acc,ax_mse) = plt.subplots(1,3)
fig.set_size_inches((16,4))
# Losses
ax_loss.plot(epochs, loss ,'c',label = 'Loss')
ax_loss.plot(epochs, value_loss, 'g' , label = 'Value Loss')
ax_loss.plot(epochs, policy_loss ,'r',label = 'Policy Loss')
ax_loss.set_title('Validation Losses History ')
ax_loss.legend()
# Accuracy
ax_acc.plot(epochs, policy_acc , 'r',label = 'Policy Accuracy')
ax_acc.set_title('Validation Policy Accuracy History ')
ax_acc.legend()
# MSE
ax_mse.plot(epochs, value_mse , 'g',label = 'Value MSE')
ax_mse.set_title('Validation Value MSE History ')
ax_mse.legend()
fig.show()
def get_validation_metrics(val_hist):
# Get Metrics for Validation
val_loss = [get_metric_validation(h,LOSS) for h in val_hist]
val_policy_loss = [get_metric_validation(h,POLICY_LOSS) for h in val_hist]
val_value_loss = [get_metric_validation(h,VALUE_LOSS) for h in val_hist]
val_policy_acc = [get_metric_validation(h,POLICY_ACC) for h in val_hist]
val_value_mse = [get_metric_validation(h,VALUE_MSE) for h in val_hist]
return {'loss': val_loss , 'policy_loss': val_policy_loss, 'value_loss': val_value_loss,
'policy_acc': val_policy_acc , 'value_mse': val_value_mse}
def plot_train_val(epochs,train_hist,val_hist,cycle=10):
val_epochs = [epoch for epoch in epochs if epoch % cycle == 0]
train_plots(epochs,train_hist)
validation_plots(val_epochs,val_hist)
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="WQXfFgUYUDqG" outputId="51c6d25a-535e-42b0-9b5c-46b9df665366"
with open('best_models/DGMV3_history_cosine', 'rb') as f_hist:
history = pickle.load(f_hist)
epochs = [ k for k in history.keys() if isinstance(k,int)]
train_hist = { k : history[k] for k in epochs}
val_hist = get_validation_metrics(history['val_history'])
plot_train_val(epochs,train_hist,val_hist)
# + id="NxAkoZKubvh5"
def train_plots(epochs , histories,styles):
# metrics = ['loss','policy_loss','value_loss','policy_categorical_accuracy','value_mse']
titles , styles = list(styles.keys()) , list(styles.values())
# Losses
loss = [get_metric(h,'loss') for h in histories]
policy_loss = [get_metric(h,'policy_loss') for h in histories]
value_loss = [get_metric(h,'value_loss') for h in histories]
# Accuracy
policy_acc = [get_metric(h,'policy_categorical_accuracy') for h in histories]
# MSE
value_mse = [get_metric(h,'value_mse') for h in histories]
fig,(ax_loss,ax_acc,ax_mse) = plt.subplots(1,3)
fig.set_size_inches((16,4))
# Losses
for i in range(len(loss)):
ax_loss.plot(epochs, loss[i] ,styles[i][0],label = f"{titles[i]} loss")
ax_loss.plot(epochs, value_loss[i], styles[i][1] , label = f"{titles[i]} val_loss")
ax_loss.plot(epochs, policy_loss[i],styles[i][2],label = f"{titles[i]} pol_loss")
ax_loss.set_title('Train Losses Histories ')
ax_loss.set_xlabel('epochs')
ax_loss.set_ylabel('Loss')
ax_loss.legend()
# Accuracy
for i in range(len(policy_acc)):
ax_acc.plot(epochs, policy_acc[i] , styles[i][0],label = f"{titles[i]} pol_acc")
ax_acc.set_title('Train Policy Accuracy Histories ')
ax_acc.set_xlabel('epochs')
ax_acc.set_ylabel('Acc')
ax_acc.legend()
# MSE
for i in range(len(value_mse)):
ax_mse.plot(epochs, value_mse[i] , styles[i][1],label = f"{titles[i]} val_mse")
ax_mse.set_title('Train Value MSE Histories ')
ax_mse.set_xlabel('epochs')
ax_mse.set_ylabel('MSE')
ax_mse.legend()
fig.show()
# + id="SD0-mGvFsQvA"
def validation_plots(epochs , histories, styles,limit=200):
# metrics = ['loss','policy_loss','value_loss','policy_categorical_accuracy','value_mse']
titles , styles = list(styles.keys()) , list(styles.values())
# Losses
loss = [h['loss'][:limit] for h in histories]
policy_loss = [h['policy_loss'][:limit] for h in histories]
value_loss = [h['value_loss'][:limit] for h in histories]
# Accuracy
policy_acc = [h['policy_acc'][:limit] for h in histories]
# MSE
value_mse = [h['value_mse'][:limit] for h in histories]
fig,(ax_loss,ax_acc,ax_mse) = plt.subplots(1,3)
fig.set_size_inches((16,4))
# Losses
for i in range(len(loss)):
ax_loss.plot(epochs, loss[i] ,styles[i][0],label = f"{titles[i]} loss")
ax_loss.plot(epochs, value_loss[i], styles[i][1] , label = f"{titles[i]} val_loss")
ax_loss.plot(epochs, policy_loss[i],styles[i][2],label = f"{titles[i]} pol_loss")
ax_loss.set_title('Validation Losses History ')
ax_loss.set_xlabel('epochs')
ax_loss.set_ylabel('Loss')
ax_loss.legend()
# Accuracy
for i in range(len(policy_acc)):
ax_acc.plot(epochs, policy_acc[i] , styles[i][0],label = f"{titles[i]} pol_acc")
ax_acc.set_title('Validation Policy Accuracy History ')
ax_acc.set_xlabel('epochs')
ax_acc.set_ylabel('Acc')
ax_acc.legend()
# MSE
for i in range(len(value_mse)):
ax_mse.plot(epochs, value_mse[i] , styles[i][1],label = f"{titles[i]} val_mse")
ax_mse.set_title('Validation Value MSE History ')
ax_mse.set_xlabel('epochs')
ax_mse.set_ylabel('MSE')
ax_mse.legend()
fig.show()
def get_validation_metrics(val_hist):
# Get Metrics for Validation
val_loss = get_metric_validation(val_hist,LOSS)
val_policy_loss = get_metric_validation(val_hist,POLICY_LOSS)
val_value_loss = get_metric_validation(val_hist,VALUE_LOSS)
val_policy_acc = get_metric_validation(val_hist,POLICY_ACC)
val_value_mse = get_metric_validation(val_hist,VALUE_MSE)
return {'loss': val_loss , 'policy_loss': val_policy_loss, 'value_loss': val_value_loss,
'policy_acc': val_policy_acc , 'value_mse': val_value_mse}
def plot_train_val(epochs,train_hist,val_hist,styles,cycle=10,limit=-1):
val_epochs = [epoch for epoch in epochs if epoch % cycle == 0]
train_plots(epochs,train_hist,styles)
validation_plots(val_epochs,val_hist,styles,limit=limit)
# + colab={"base_uri": "https://localhost:8080/", "height": 876} id="TE3fKMLuL6u1" outputId="6d4285c9-2df1-4ce8-aa9e-fe63a2c2a7e1"
path = 'best_models/histories/mnas/DGMV9_history_mnas_250_15_625_sw_1_to_5000'
with ExitStack() as stack:
files = [stack.enter_context(open(path,'rb'))]
histories = [pickle.load(f) for f in files]
# Plot data (x-axis : epochs , y-axis : histories)
epochs = [ k for k in histories[0].keys() if isinstance(k,int) and k <= 2000]
train_hist = [{ k : history[k] for k in epochs} for history in histories]
val_hist = [get_validation_metrics(history['val_history']) for history in histories]
# The Biggest event ever ! lets plot s#it out !
plot_train_val(epochs,train_hist,val_hist,styles,limit=2000)
# + colab={"base_uri": "https://localhost:8080/", "height": 876} id="ZGzdFK-gfT-9" outputId="c51e5463-bfff-44c7-f9bf-264fc1014163"
from contextlib import ExitStack
# First , let's load history files in a clever way 😁
# Here , are histories for models
models_config = {
'mnet' : {
'path' : 'best_models/DGMV4_history_mnet_se_256_sw_2000',
'style': ['k','r','gold']
}
,
'incp' : {
'path': 'best_models/DGMV8_history_cosine_inception_se_192_sw_2000',
'style': ['springgreen','dodgerblue','indigo']
}
,
'mnas' : {
'path': 'best_models/histories/mnas/DGMV9_history_mnas_250_15_625_sw_1_to_5000',
'style': ['fuchsia','lime','aqua']
}
}
# Here we manage to load this histories from the pickle binaries
with ExitStack() as stack:
files = [stack.enter_context(open(fname['path'],'rb')) for fname in models_config.values()]
histories = [pickle.load(f) for f in files]
# Why not some style ? 🇧🇩 ( always in a clever and a dynamic way ! 😃 )
styles = {name : models_config[name]['style'] for name in models_config.keys()}
# Plot data (x-axis : epochs , y-axis : histories)
epochs = [ k for k in histories[0].keys() if isinstance(k,int) and k <= 2000]
train_hist = [{ k : history[k] for k in epochs} for history in histories]
val_hist = [get_validation_metrics(history['val_history']) for history in histories]
# The Biggest event ever ! lets plot s#it out !
plot_train_val(epochs,train_hist,val_hist,styles)
# + id="7M0yTbIWfd2m" colab={"base_uri": "https://localhost:8080/"} outputId="525bcf8d-df03-47b5-d3e9-597a4ab5feb9"
import os
path = "./best_models/histories/mnet"
# A helper function in order to sort names via epochs order
def rename_hist_name(old_name):
new_name = old_name.split('_')
new_name[-1] = f"{int(new_name[-1]):05d}" # rename start epoch
new_name[-3] = f"{int(new_name[-3]):05d}" # rename end epoch
return "_".join(new_name)
def rename_hist_file(path,old_name=""):
os.rename(f"{path}/{old_name}",f"{path}/{rename_hist_name(old_name)}")
# Walk into histories and get ordered set of files (names)
filenames = next(os.walk(path), (None, None, []))[2]
for f_name in filenames:
rename_hist_file(path,f_name)
filenames = next(os.walk(path), (None, None, []))[2]
filenames.sort()
rename_hist_file(path,filenames[0])
hist_path_1 = f"./best_models/histories/mnet/{filenames[0]}"
hist_path_2 = f"./best_models/histories/mnet/{filenames[1]}"
with open(hist_path_1,"rb") as f_hist_1 , open(hist_path_2,"rb") as f_hist_2:
histories_1 = pickle.load(f_hist_1)
histories_2 = pickle.load(f_hist_2)
histories = [histories_2]
epochs_1 = [ k for k in histories_1.keys() if isinstance(k,int)]
epochs_2 = [ k for k in histories_2.keys() if isinstance(k,int)]
epochs = epochs_1 + epochs_2
train_hist_1 = [{ k : history[k] for k in epochs_1} for history in histories_1]
train_hist_2 = [{ k : history[k] for k in epochs_2[1:]} for history in histories_2]
histories =
# + id="ob30799BfgCu"
| src/train_dgm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''base'': conda)'
# name: python3
# ---
# # Multiphase Systems
#
# Kawin supports the usage of multiple phases. Nucleation and growth rate are handled for each precipitate phase independently. Coupling comes from the mass balance where all precipitates contribute to the overall mass changes in the system.
#
# In the Al-Mg-Si system, several phases can form including: $ \beta' $, $ \beta" $, B', U1 and U2. To model precipitation of these phases, they must be defined in the .tdb file, the Thermodynamics module and the PrecipitateModel module.
#
# When defining the thermodynamics module, the first phase in the list of phases will be the parent phase.
# +
from kawin.Thermodynamics import MulticomponentThermodynamics
phases = ['FCC_A1', 'MGSI_B_P', 'MG5SI6_B_DP', 'B_PRIME_L', 'U1_PHASE', 'U2_PHASE']
therm = MulticomponentThermodynamics('AlMgSi.tdb', ['AL', 'MG', 'SI'], phases, drivingForceMethod='approximate')
# -
# In defining the precipitate model, all precipitate phases must be included. Since we already have our list of phases, we can use that and remove the parent phase.
# +
from kawin.KWNEuler import PrecipitateModel
model = PrecipitateModel(0, 25*3600, 1e4, phases=phases[1:], elements=['MG', 'SI'], linearTimeSpacing=True)
# -
# ## Model inputs
#
# Setting up parameters for the parent phase and overall system is the same as for single phase systems. Here, it is just the composition (Al-0.72Mg-0.57Si in mol. %), molar volume ($1e$-$5\text{ }m^3/mol$).
#
# The temperature will be divided into two stages: a 16 hour temper at $175\text{ }^oC$, followed by a 1 hour ramp up to $250 ^oC$. To do this, there needs to be three time designations: $175\text{ }^oC$ at 0 hours, $175\text{ }^oC$ at 16 hours and $250\text{ }^oC$ at 17 hours. The temperature can be plotted to show the profile over time. Here, a parameter called timeUnits is passed to convert the time from seconds to either minutes or hours.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
model.setInitialComposition([0.0072, 0.0057])
model.setVmAlpha(1e-5, 4)
lowTemp = 175+273.15
highTemp = 250+273.15
model.setTemperatureArray([0, 16, 17], [lowTemp, lowTemp, highTemp])
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
model.plot(ax, 'Temperature', timeUnits='h')
ax.set_ylim([400, 550])
ax.set_xscale('linear')
# -
# Setting parameters for each precipitate phase is similar to single phase systems except that the phase has to be defined when inputting parameters.
# +
gamma = {
'MGSI_B_P': 0.18,
'MG5SI6_B_DP': 0.084,
'B_PRIME_L': 0.18,
'U1_PHASE': 0.18,
'U2_PHASE': 0.18
}
for i in range(len(phases)-1):
model.setInterfacialEnergy(gamma[phases[i+1]], phase=phases[i+1])
model.setVmBeta(1e-5, 4, phase=phases[i+1])
model.setThermodynamics(therm, phase=phases[i+1])
# -
# ## Solving the model
#
# As with single precipitate phase systems, running the model is exactly the same.
model.solve(verbose=True, vIt=5000)
# ## Plotting
#
# Plotting is also the same as with single phase systems. The major difference is each phase will be plotted for the radius, volume fraction, precipitate density, nucleation rate and particle size distribution. In addition, the total amount of some variables, such as the precipitate density and volume fraction, can be plotted.
# +
fig, axes = plt.subplots(2, 2, figsize=(12, 10))
model.plot(axes[0,0], 'Total Precipitate Density', timeUnits='h', label='Total', color='k', linestyle=':', zorder=6)
model.plot(axes[0,0], 'Precipitate Density', timeUnits='h')
axes[0,0].set_ylim([1e5, 1e25])
axes[0,0].set_xscale('linear')
axes[0,0].set_yscale('log')
model.plot(axes[0,1], 'Total Volume Fraction', timeUnits='h', label='Total', color='k', linestyle=':', zorder=6)
model.plot(axes[0,1], 'Volume Fraction', timeUnits='h')
axes[0,1].set_xscale('linear')
model.plot(axes[1,0], 'Average Radius', timeUnits='h')
axes[1,0].set_xscale('linear')
model.plot(axes[1,1], 'Composition', timeUnits='h')
axes[1,1].set_xscale('linear')
fig.tight_layout()
# -
# ## References
#
# 1. <NAME> et al, "Calphad modeling of metastable phases in the Al-Mg-Si system" *Calphad* 43 (2013) p. 94
# 2. Q. Du et al, "Modeling over-ageing in Al-Mg-Si alloys by a multi-phase Calphad-coupled Kampmann-Wagner Numerical model" *Acta Materialia* 122 (2017) p. 178
| examples/Multiphase Precipitation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# ### Variational Autoencoders Explained
# First, the images are generated off some arbitrary noise. If you wanted to generate a picture with specific features, there's no way of determining which initial noise values would produce that picture, other than searching over the entire distribution.
# ### What is a variational autoencoder?
# To get an understanding of a VAE, we'll first start from a simple network and add parts step by step. An common way of describing a neural network is an approximation of some function we wish to model. However, they can also be thought of as a data structure that holds information.
#
# Let's say we had a network comprised of a few deconvolution layers. We set the input to always be a vector of ones. Then, we can train the network to reduce the mean squared error between itself and one target image. The "data" for that image is now contained within the network's parameters.
| 04_Autoencoder/04_AE_09_VarAE_ae-var_theory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="t09eeeR5prIJ"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="GCCk8_dHpuNf"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="ovpZyIhNIgoq"
# # Text generation with an RNN
# + [markdown] id="hcD2nPQvPOFM"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/text/tutorials/text_generation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/text/blob/master/docs/tutorials/text_generation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/text/blob/master/docs/tutorials/text_generation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/text/docs/tutorials/text_generation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="BwpJ5IffzRG6"
# This tutorial demonstrates how to generate text using a character-based RNN. You will work with a dataset of Shakespeare's writing from <NAME>'s [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/). Given a sequence of characters from this data ("Shakespear"), train a model to predict the next character in the sequence ("e"). Longer sequences of text can be generated by calling the model repeatedly.
#
# Note: Enable GPU acceleration to execute this notebook faster. In Colab: *Runtime > Change runtime type > Hardware accelerator > GPU*.
#
# This tutorial includes runnable code implemented using [tf.keras](https://www.tensorflow.org/guide/keras/sequential_model) and [eager execution](https://www.tensorflow.org/guide/eager). The following is the sample output when the model in this tutorial trained for 30 epochs, and started with the prompt "Q":
# + [markdown] id="HcygKkEVZBaa"
# <pre>
# QUEENE:
# I had thought thou hadst a Roman; for the oracle,
# Thus by All bids the man against the word,
# Which are so weak of care, by old care done;
# Your children were in your holy love,
# And the precipitation through the bleeding throne.
#
# BISHOP OF ELY:
# Marry, and will, my lord, to weep in such a one were prettiest;
# Yet now I was adopted heir
# Of the world's lamentable day,
# To watch the next way with his father with his face?
#
# ESCALUS:
# The cause why then we are all resolved more sons.
#
# VOLUMNIA:
# O, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, it is no sin it should be dead,
# And love and pale as any will to that word.
#
# QUEEN ELIZABETH:
# But how long have I heard the soul for this world,
# And show his hands of life be proved to stand.
#
# PETRUCHIO:
# I say he look'd on, if I must be content
# To stay him from the fatal of our country's bliss.
# His lordship pluck'd from this sentence then for prey,
# And then let us twain, being the moon,
# were she such a case as fills m
# </pre>
# + [markdown] id="_bGsCP9DZFQ5"
# While some of the sentences are grammatical, most do not make sense. The model has not learned the meaning of words, but consider:
#
# * The model is character-based. When training started, the model did not know how to spell an English word, or that words were even a unit of text.
#
# * The structure of the output resembles a play—blocks of text generally begin with a speaker name, in all capital letters similar to the dataset.
#
# * As demonstrated below, the model is trained on small batches of text (100 characters each), and is still able to generate a longer sequence of text with coherent structure.
# + [markdown] id="srXC6pLGLwS6"
# ## Setup
# + [markdown] id="WGyKZj3bzf9p"
# ### Import TensorFlow and other libraries
# + id="yG_n40gFzf9s"
import tensorflow as tf
from tensorflow.keras.layers.experimental import preprocessing
import numpy as np
import os
import time
# + [markdown] id="EHDoRoc5PKWz"
# ### Download the Shakespeare dataset
#
# Change the following line to run this code on your own data.
# + id="pD_55cOxLkAb"
path_to_file = tf.keras.utils.get_file('shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
# + [markdown] id="UHjdCjDuSvX_"
# ### Read the data
#
# First, look in the text:
# + id="aavnuByVymwK"
# Read, then decode for py2 compat.
text = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# length of text is the number of characters in it
print(f'Length of text: {len(text)} characters')
# + id="Duhg9NrUymwO"
# Take a look at the first 250 characters in text
print(text[:250])
# + id="IlCgQBRVymwR"
# The unique characters in the file
vocab = sorted(set(text))
print(f'{len(vocab)} unique characters')
# + [markdown] id="rNnrKn_lL-IJ"
# ## Process the text
# + [markdown] id="LFjSVAlWzf-N"
# ### Vectorize the text
#
# Before training, you need to convert the strings to a numerical representation.
#
# The `preprocessing.StringLookup` layer can convert each character into a numeric ID. It just needs the text to be split into tokens first.
# + id="a86OoYtO01go"
example_texts = ['abcdefg', 'xyz']
chars = tf.strings.unicode_split(example_texts, input_encoding='UTF-8')
chars
# + [markdown] id="1s4f1q3iqY8f"
# Now create the `preprocessing.StringLookup` layer:
# + id="6GMlCe3qzaL9"
ids_from_chars = preprocessing.StringLookup(
vocabulary=list(vocab), mask_token=None)
# + [markdown] id="ZmX_jbgQqfOi"
# It converts form tokens to character IDs:
# + id="WLv5Q_2TC2pc"
ids = ids_from_chars(chars)
ids
# + [markdown] id="tZfqhkYCymwX"
# Since the goal of this tutorial is to generate text, it will also be important to invert this representation and recover human-readable strings from it. For this you can use `preprocessing.StringLookup(..., invert=True)`.
# + [markdown] id="uenivzwqsDhp"
# Note: Here instead of passing the original vocabulary generated with `sorted(set(text))` use the `get_vocabulary()` method of the `preprocessing.StringLookup` layer so that the `[UNK]` tokens is set the same way.
# + id="Wd2m3mqkDjRj"
chars_from_ids = tf.keras.layers.experimental.preprocessing.StringLookup(
vocabulary=ids_from_chars.get_vocabulary(), invert=True, mask_token=None)
# + [markdown] id="pqTDDxS-s-H8"
# This layer recovers the characters from the vectors of IDs, and returns them as a `tf.RaggedTensor` of characters:
# + id="c2GCh0ySD44s"
chars = chars_from_ids(ids)
chars
# + [markdown] id="-FeW5gqutT3o"
# You can `tf.strings.reduce_join` to join the characters back into strings.
# + id="zxYI-PeltqKP"
tf.strings.reduce_join(chars, axis=-1).numpy()
# + id="w5apvBDn9Ind"
def text_from_ids(ids):
return tf.strings.reduce_join(chars_from_ids(ids), axis=-1)
# + [markdown] id="bbmsf23Bymwe"
# ### The prediction task
# + [markdown] id="wssHQ1oGymwe"
# Given a character, or a sequence of characters, what is the most probable next character? This is the task you're training the model to perform. The input to the model will be a sequence of characters, and you train the model to predict the output—the following character at each time step.
#
# Since RNNs maintain an internal state that depends on the previously seen elements, given all the characters computed until this moment, what is the next character?
#
# + [markdown] id="hgsVvVxnymwf"
# ### Create training examples and targets
#
# Next divide the text into example sequences. Each input sequence will contain `seq_length` characters from the text.
#
# For each input sequence, the corresponding targets contain the same length of text, except shifted one character to the right.
#
# So break the text into chunks of `seq_length+1`. For example, say `seq_length` is 4 and our text is "Hello". The input sequence would be "Hell", and the target sequence "ello".
#
# To do this first use the `tf.data.Dataset.from_tensor_slices` function to convert the text vector into a stream of character indices.
# + id="UopbsKi88tm5"
all_ids = ids_from_chars(tf.strings.unicode_split(text, 'UTF-8'))
all_ids
# + id="qmxrYDCTy-eL"
ids_dataset = tf.data.Dataset.from_tensor_slices(all_ids)
# + id="cjH5v45-yqqH"
for ids in ids_dataset.take(10):
print(chars_from_ids(ids).numpy().decode('utf-8'))
# + id="C-G2oaTxy6km"
seq_length = 100
examples_per_epoch = len(text)//(seq_length+1)
# + [markdown] id="-ZSYAcQV8OGP"
# The `batch` method lets you easily convert these individual characters to sequences of the desired size.
# + id="BpdjRO2CzOfZ"
sequences = ids_dataset.batch(seq_length+1, drop_remainder=True)
for seq in sequences.take(1):
print(chars_from_ids(seq))
# + [markdown] id="5PHW902-4oZt"
# It's easier to see what this is doing if you join the tokens back into strings:
# + id="QO32cMWu4a06"
for seq in sequences.take(5):
print(text_from_ids(seq).numpy())
# + [markdown] id="UbLcIPBj_mWZ"
# For training you'll need a dataset of `(input, label)` pairs. Where `input` and
# `label` are sequences. At each time step the input is the current character and the label is the next character.
#
# Here's a function that takes a sequence as input, duplicates, and shifts it to align the input and label for each timestep:
# + id="9NGu-FkO_kYU"
def split_input_target(sequence):
input_text = sequence[:-1]
target_text = sequence[1:]
return input_text, target_text
# + id="WxbDTJTw5u_P"
split_input_target(list("Tensorflow"))
# + id="B9iKPXkw5xwa"
dataset = sequences.map(split_input_target)
# + id="GNbw-iR0ymwj"
for input_example, target_example in dataset.take(1):
print("Input :", text_from_ids(input_example).numpy())
print("Target:", text_from_ids(target_example).numpy())
# + [markdown] id="MJdfPmdqzf-R"
# ### Create training batches
#
# You used `tf.data` to split the text into manageable sequences. But before feeding this data into the model, you need to shuffle the data and pack it into batches.
# + id="p2pGotuNzf-S"
# Batch size
BATCH_SIZE = 64
# Buffer size to shuffle the dataset
# (TF data is designed to work with possibly infinite sequences,
# so it doesn't attempt to shuffle the entire sequence in memory. Instead,
# it maintains a buffer in which it shuffles elements).
BUFFER_SIZE = 10000
dataset = (
dataset
.shuffle(BUFFER_SIZE)
.batch(BATCH_SIZE, drop_remainder=True)
.prefetch(tf.data.experimental.AUTOTUNE))
dataset
# + [markdown] id="r6oUuElIMgVx"
# ## Build The Model
# + [markdown] id="m8gPwEjRzf-Z"
# This section defines the model as a `keras.Model` subclass (For details see [Making new Layers and Models via subclassing](https://www.tensorflow.org/guide/keras/custom_layers_and_models)).
#
# This model has three layers:
#
# * `tf.keras.layers.Embedding`: The input layer. A trainable lookup table that will map each character-ID to a vector with `embedding_dim` dimensions;
# * `tf.keras.layers.GRU`: A type of RNN with size `units=rnn_units` (You can also use an LSTM layer here.)
# * `tf.keras.layers.Dense`: The output layer, with `vocab_size` outputs. It outputs one logit for each character in the vocabulary. These are the log-likelihood of each character according to the model.
# + id="zHT8cLh7EAsg"
# Length of the vocabulary in chars
vocab_size = len(vocab)
# The embedding dimension
embedding_dim = 256
# Number of RNN units
rnn_units = 1024
# + id="wj8HQ2w8z4iO"
class MyModel(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, rnn_units):
super().__init__(self)
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(rnn_units,
return_sequences=True,
return_state=True)
self.dense = tf.keras.layers.Dense(vocab_size)
def call(self, inputs, states=None, return_state=False, training=False):
x = inputs
x = self.embedding(x, training=training)
if states is None:
states = self.gru.get_initial_state(x)
x, states = self.gru(x, initial_state=states, training=training)
x = self.dense(x, training=training)
if return_state:
return x, states
else:
return x
# + id="IX58Xj9z47Aw"
model = MyModel(
# Be sure the vocabulary size matches the `StringLookup` layers.
vocab_size=len(ids_from_chars.get_vocabulary()),
embedding_dim=embedding_dim,
rnn_units=rnn_units)
# + [markdown] id="RkA5upJIJ7W7"
# For each character the model looks up the embedding, runs the GRU one timestep with the embedding as input, and applies the dense layer to generate logits predicting the log-likelihood of the next character:
#
# 
# + [markdown] id="gKbfm04amhXk"
# Note: For training you could use a `keras.Sequential` model here. To generate text later you'll need to manage the RNN's internal state. It's simpler to include the state input and output options upfront, than it is to rearrange the model architecture later. For more details see the [Keras RNN guide](https://www.tensorflow.org/guide/keras/rnn#rnn_state_reuse).
# + [markdown] id="-ubPo0_9Prjb"
# ## Try the model
#
# Now run the model to see that it behaves as expected.
#
# First check the shape of the output:
# + id="C-_70kKAPrPU"
for input_example_batch, target_example_batch in dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(example_batch_predictions.shape, "# (batch_size, sequence_length, vocab_size)")
# + [markdown] id="Q6NzLBi4VM4o"
# In the above example the sequence length of the input is `100` but the model can be run on inputs of any length:
# + id="vPGmAAXmVLGC"
model.summary()
# + [markdown] id="uwv0gEkURfx1"
# To get actual predictions from the model you need to sample from the output distribution, to get actual character indices. This distribution is defined by the logits over the character vocabulary.
#
# Note: It is important to _sample_ from this distribution as taking the _argmax_ of the distribution can easily get the model stuck in a loop.
#
# Try it for the first example in the batch:
# + id="4V4MfFg0RQJg"
sampled_indices = tf.random.categorical(example_batch_predictions[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices, axis=-1).numpy()
# + [markdown] id="QM1Vbxs_URw5"
# This gives us, at each timestep, a prediction of the next character index:
# + id="YqFMUQc_UFgM"
sampled_indices
# + [markdown] id="LfLtsP3mUhCG"
# Decode these to see the text predicted by this untrained model:
# + id="xWcFwPwLSo05"
print("Input:\n", text_from_ids(input_example_batch[0]).numpy())
print()
print("Next Char Predictions:\n", text_from_ids(sampled_indices).numpy())
# + [markdown] id="LJL0Q0YPY6Ee"
# ## Train the model
# + [markdown] id="YCbHQHiaa4Ic"
# At this point the problem can be treated as a standard classification problem. Given the previous RNN state, and the input this time step, predict the class of the next character.
# + [markdown] id="trpqTWyvk0nr"
# ### Attach an optimizer, and a loss function
# + [markdown] id="UAjbjY03eiQ4"
# The standard `tf.keras.losses.sparse_categorical_crossentropy` loss function works in this case because it is applied across the last dimension of the predictions.
#
# Because your model returns logits, you need to set the `from_logits` flag.
#
# + id="ZOeWdgxNFDXq"
loss = tf.losses.SparseCategoricalCrossentropy(from_logits=True)
# + id="4HrXTACTdzY-"
example_batch_loss = loss(target_example_batch, example_batch_predictions)
mean_loss = example_batch_loss.numpy().mean()
print("Prediction shape: ", example_batch_predictions.shape, " # (batch_size, sequence_length, vocab_size)")
print("Mean loss: ", mean_loss)
# + [markdown] id="vkvUIneTFiow"
# A newly initialized model shouldn't be too sure of itself, the output logits should all have similar magnitudes. To confirm this you can check that the exponential of the mean loss is approximately equal to the vocabulary size. A much higher loss means the model is sure of its wrong answers, and is badly initialized:
# + id="MAJfS5YoFiHf"
tf.exp(mean_loss).numpy()
# + [markdown] id="jeOXriLcymww"
# Configure the training procedure using the `tf.keras.Model.compile` method. Use `tf.keras.optimizers.Adam` with default arguments and the loss function.
# + id="DDl1_Een6rL0"
model.compile(optimizer='adam', loss=loss)
# + [markdown] id="ieSJdchZggUj"
# ### Configure checkpoints
# + [markdown] id="C6XBUUavgF56"
# Use a `tf.keras.callbacks.ModelCheckpoint` to ensure that checkpoints are saved during training:
# + id="W6fWTriUZP-n"
# Directory where the checkpoints will be saved
checkpoint_dir = './training_checkpoints'
# Name of the checkpoint files
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
# + [markdown] id="3Ky3F_BhgkTW"
# ### Execute the training
# + [markdown] id="IxdOA-rgyGvs"
# To keep training time reasonable, use 10 epochs to train the model. In Colab, set the runtime to GPU for faster training.
# + id="7yGBE2zxMMHs"
EPOCHS = 20
# + id="UK-hmKjYVoll"
history = model.fit(dataset, epochs=EPOCHS, callbacks=[checkpoint_callback])
# + [markdown] id="kKkD5M6eoSiN"
# ## Generate text
# + [markdown] id="oIdQ8c8NvMzV"
# The simplest way to generate text with this model is to run it in a loop, and keep track of the model's internal state as you execute it.
#
# 
#
# Each time you call the model you pass in some text and an internal state. The model returns a prediction for the next character and its new state. Pass the prediction and state back in to continue generating text.
#
# + [markdown] id="DjGz1tDkzf-u"
# The following makes a single step prediction:
# + id="iSBU1tHmlUSs"
class OneStep(tf.keras.Model):
def __init__(self, model, chars_from_ids, ids_from_chars, temperature=1.0):
super().__init__()
self.temperature = temperature
self.model = model
self.chars_from_ids = chars_from_ids
self.ids_from_chars = ids_from_chars
# Create a mask to prevent "[UNK]" from being generated.
skip_ids = self.ids_from_chars(['[UNK]'])[:, None]
sparse_mask = tf.SparseTensor(
# Put a -inf at each bad index.
values=[-float('inf')]*len(skip_ids),
indices=skip_ids,
# Match the shape to the vocabulary
dense_shape=[len(ids_from_chars.get_vocabulary())])
self.prediction_mask = tf.sparse.to_dense(sparse_mask)
@tf.function
def generate_one_step(self, inputs, states=None):
# Convert strings to token IDs.
input_chars = tf.strings.unicode_split(inputs, 'UTF-8')
input_ids = self.ids_from_chars(input_chars).to_tensor()
# Run the model.
# predicted_logits.shape is [batch, char, next_char_logits]
predicted_logits, states = self.model(inputs=input_ids, states=states,
return_state=True)
# Only use the last prediction.
predicted_logits = predicted_logits[:, -1, :]
predicted_logits = predicted_logits/self.temperature
# Apply the prediction mask: prevent "[UNK]" from being generated.
predicted_logits = predicted_logits + self.prediction_mask
# Sample the output logits to generate token IDs.
predicted_ids = tf.random.categorical(predicted_logits, num_samples=1)
predicted_ids = tf.squeeze(predicted_ids, axis=-1)
# Convert from token ids to characters
predicted_chars = self.chars_from_ids(predicted_ids)
# Return the characters and model state.
return predicted_chars, states
# + id="fqMOuDutnOxK"
one_step_model = OneStep(model, chars_from_ids, ids_from_chars)
# + [markdown] id="p9yDoa0G3IgQ"
# Run it in a loop to generate some text. Looking at the generated text, you'll see the model knows when to capitalize, make paragraphs and imitates a Shakespeare-like writing vocabulary. With the small number of training epochs, it has not yet learned to form coherent sentences.
# + id="ST7PSyk9t1mT"
start = time.time()
states = None
next_char = tf.constant(['ROMEO:'])
result = [next_char]
for n in range(1000):
next_char, states = one_step_model.generate_one_step(next_char, states=states)
result.append(next_char)
result = tf.strings.join(result)
end = time.time()
print(result[0].numpy().decode('utf-8'), '\n\n' + '_'*80)
print('\nRun time:', end - start)
# + [markdown] id="AM2Uma_-yVIq"
# The easiest thing you can do to improve the results is to train it for longer (try `EPOCHS = 30`).
#
# You can also experiment with a different start string, try adding another RNN layer to improve the model's accuracy, or adjust the temperature parameter to generate more or less random predictions.
# + [markdown] id="_OfbI4aULmuj"
# If you want the model to generate text *faster* the easiest thing you can do is batch the text generation. In the example below the model generates 5 outputs in about the same time it took to generate 1 above.
# + id="ZkLu7Y8UCMT7"
start = time.time()
states = None
next_char = tf.constant(['ROMEO:', 'ROMEO:', 'ROMEO:', 'ROMEO:', 'ROMEO:'])
result = [next_char]
for n in range(1000):
next_char, states = one_step_model.generate_one_step(next_char, states=states)
result.append(next_char)
result = tf.strings.join(result)
end = time.time()
print(result, '\n\n' + '_'*80)
print('\nRun time:', end - start)
# + [markdown] id="UlUQzwu6EXam"
# ## Export the generator
#
# This single-step model can easily be [saved and restored](https://www.tensorflow.org/guide/saved_model), allowing you to use it anywhere a `tf.saved_model` is accepted.
# + id="3Grk32H_CzsC"
tf.saved_model.save(one_step_model, 'one_step')
one_step_reloaded = tf.saved_model.load('one_step')
# + id="_Z9bb_wX6Uuu"
states = None
next_char = tf.constant(['ROMEO:'])
result = [next_char]
for n in range(100):
next_char, states = one_step_reloaded.generate_one_step(next_char, states=states)
result.append(next_char)
print(tf.strings.join(result)[0].numpy().decode("utf-8"))
# + [markdown] id="Y4QwTjAM6A2O"
# ## Advanced: Customized Training
#
# The above training procedure is simple, but does not give you much control.
# It uses teacher-forcing which prevents bad predictions from being fed back to the model, so the model never learns to recover from mistakes.
#
# So now that you've seen how to run the model manually next you'll implement the training loop. This gives a starting point if, for example, you want to implement _curriculum learning_ to help stabilize the model's open-loop output.
#
# The most important part of a custom training loop is the train step function.
#
# Use `tf.GradientTape` to track the gradients. You can learn more about this approach by reading the [eager execution guide](https://www.tensorflow.org/guide/eager).
#
# The basic procedure is:
#
# 1. Execute the model and calculate the loss under a `tf.GradientTape`.
# 2. Calculate the updates and apply them to the model using the optimizer.
# + id="x0pZ101hjwW0"
class CustomTraining(MyModel):
@tf.function
def train_step(self, inputs):
inputs, labels = inputs
with tf.GradientTape() as tape:
predictions = self(inputs, training=True)
loss = self.loss(labels, predictions)
grads = tape.gradient(loss, model.trainable_variables)
self.optimizer.apply_gradients(zip(grads, model.trainable_variables))
return {'loss': loss}
# + [markdown] id="4Oc-eJALcK8B"
# The above implementation of the `train_step` method follows [Keras' `train_step` conventions](https://www.tensorflow.org/guide/keras/customizing_what_happens_in_fit). This is optional, but it allows you to change the behavior of the train step and still use keras' `Model.compile` and `Model.fit` methods.
# + id="XKyWiZ_Lj7w5"
model = CustomTraining(
vocab_size=len(ids_from_chars.get_vocabulary()),
embedding_dim=embedding_dim,
rnn_units=rnn_units)
# + id="U817KUm7knlm"
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True))
# + id="o694aoBPnEi9"
model.fit(dataset, epochs=1)
# + [markdown] id="W8nAtKHVoInR"
# Or if you need more control, you can write your own complete custom training loop:
# + id="d4tSNwymzf-q"
EPOCHS = 10
mean = tf.metrics.Mean()
for epoch in range(EPOCHS):
start = time.time()
mean.reset_states()
for (batch_n, (inp, target)) in enumerate(dataset):
logs = model.train_step([inp, target])
mean.update_state(logs['loss'])
if batch_n % 50 == 0:
template = f"Epoch {epoch+1} Batch {batch_n} Loss {logs['loss']:.4f}"
print(template)
# saving (checkpoint) the model every 5 epochs
if (epoch + 1) % 5 == 0:
model.save_weights(checkpoint_prefix.format(epoch=epoch))
print()
print(f'Epoch {epoch+1} Loss: {mean.result().numpy():.4f}')
print(f'Time taken for 1 epoch {time.time() - start:.2f} sec')
print("_"*80)
model.save_weights(checkpoint_prefix.format(epoch=epoch))
| src/text/text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
using PlotlyJS, WebIO
#WebIO.install_jupyter_nbextension()
# +
l = Layout(
title="Penguins",
xaxis=attr(title="fish", showgrid=true),
yaxis=attr(title="Weight", showgrid=true),
legend=attr(x=0.1, y=0.1),
width = 700,
height = 500,
)
trace1 = scatter(;x=1:4, y=[10, 15, 13, 17], mode="markers")
trace2 = scatter(;x=2:5, y=[16, 5, 11, 9], mode="lines")
trace3 = scatter(;x=1:4, y=[12, 9, 10, 12], mode="lines+markers")
# Display in-place
plt = plot([trace1, trace2, trace3], l)
display(plt)
# Display in-place
#p = Plot([trace1, trace2, trace3], l) # for display in-place
#display(p)
# -
# ## PlotlyJS works
| learn/learn_Jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# +
# !pip install gcsfs
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import glob
import os
import gcsfs
import tensorflow as tf
from tensorflow.python.lib.io import file_io
# -
#
#
# +
# path =r'C:/Users/depri/Desktop/OPS813 cloud computing/Project/case1/amex-nyse-nasdaq-stock-histories/fh_20190217/full_history'
fs = gcsfs.GCSFileSystem(project='Case1-TTPRSD')
# allFiles = glob.glob(path + "/*.csv")
allFiles = fs.ls('stockdatagrp21/full_history')
# allFiles
alldf = []
i=0
for file_ in allFiles:
i+=1
if i%100 == 0:
print(i)
# if i==400:
# break
with fs.open(file_) as f:
dftemp = pd.read_csv(f,index_col=None, header=0)
# dftemp = pd.read_csv(file_,index_col=None, header=0)
# dftemp = df_npd.compute()
#Figure out a way to invert the data frame
dftemp = dftemp[::-1]
#Calculating the 200 day moving average before merging all data
dftemp["200d"] = np.round(dftemp["adjclose"].rolling(window = 200).mean(), 2)
#Calculating the 50 day moving average before merging all data
dftemp["50d"] = np.round(dftemp["adjclose"].rolling(window = 50).mean(), 2)
#Adding a new colmn stock name to identify the stock in question
dftemp["StockName"]=os.path.basename(file_).split('.')[0]
alldf.append(dftemp)
df = pd.concat(alldf, axis = 0, ignore_index = True)
df.columns = ["Date", "Volume","Open", "Close", "High", "Low", "AdjustedClose","200d","50d","StockName"]
# print(df)
# +
#Finding stocks in the last month with golden cross and visualizing it
allgoldencrossings = []
alldeathcrossings = []
for key, grp in df.groupby('StockName'): # handling data by each stock.
previous_50 = grp['50d'].shift(1) # shifting data by 1.
previous_200 = grp['200d'].shift(1) # shifting data by 1.
deathcrossing = ((grp['50d'] <= grp['200d']) & (previous_50 > previous_200))
goldencrossing = ((grp['50d'] >= grp['200d']) & (previous_50 < previous_200))
allgoldencrossings.append(goldencrossing) # Comparing current value of 50d and 200d and comparing previous value of 50d and 200d to see if the golden cross is achieved.
# figuring out whether I am having golden cross or not.
alldeathcrossings.append(deathcrossing)
df['GoldenCross'] = pd.concat(allgoldencrossings)
df['DeathCross'] = pd.concat(alldeathcrossings)
df['WeekChange'] = df['AdjustedClose'].pct_change(periods=7)*100
df['TwoWeekChange'] = df['AdjustedClose'].pct_change(periods=14)*100
df['ThreeWeekChange'] = df['AdjustedClose'].pct_change(periods=21)*100
df['MonthChange'] = df['AdjustedClose'].pct_change(periods=30)*100
df['TwoMonthChange'] = df['AdjustedClose'].pct_change(periods=60)*100
df['ThreeMonthChange'] = df['AdjustedClose'].pct_change(periods=90)*100
df['FourMonthChange'] = df['AdjustedClose'].pct_change(periods=120)*100
df['FiveMonthChange'] = df['AdjustedClose'].pct_change(periods=150)*100
df['SixMonthChange'] = df['AdjustedClose'].pct_change(periods=180)*100
# Count of golden crosses
goldenCount = df[df.GoldenCross].GoldenCross.count()
# Count of death crosses
deathCount = df[df.DeathCross].DeathCross.count()
# df
# Percentage of change of 1% in Golden Cross and Death Cross
postiveOnePercentChange = (df[df.GoldenCross & (df.WeekChange > 1)].GoldenCross.count()/goldenCount)*100
negativeOnePercentChange = (df[df.DeathCross & (df.WeekChange < 1)].DeathCross.count()/deathCount)*100
print(postiveOnePercentChange)
print(negativeOnePercentChange)
# Percentage of change of 2% in Golden Cross and Death Cross
postiveTwoPercentChange = (df[df.GoldenCross & (df.TwoWeekChange > 2)].GoldenCross.count()/goldenCount)*100
negativeTwoPercentChange = (df[df.DeathCross & (df.TwoWeekChange < 2)].DeathCross.count()/deathCount)*100
print(postiveTwoPercentChange)
print(negativeTwoPercentChange)
# Percentage of change of 3% in Golden Cross and Death Cross
postiveThreePercentChange = (df[df.GoldenCross & (df.ThreeWeekChange > 3)].GoldenCross.count()/goldenCount)*100
negativeThreePercentChange = (df[df.DeathCross & (df.ThreeWeekChange < 3)].DeathCross.count()/deathCount)*100
print(postiveThreePercentChange)
print(negativeThreePercentChange)
# Percentage of change of 4% in Golden Cross and Death Cross
postiveFourPercentChange = (df[df.GoldenCross & (df.MonthChange > 4)].GoldenCross.count()/goldenCount)*100
negativeFourPercentChange = (df[df.DeathCross & (df.MonthChange < 4)].DeathCross.count()/deathCount)*100
print(postiveFourPercentChange)
print(negativeFourPercentChange)
# Percentage of change of 5% in Golden Cross and Death Cross
postiveFivePercentChange = (df[df.GoldenCross & (df.TwoMonthChange > 5)].GoldenCross.count()/goldenCount)*100
negativeFivePercentChange = (df[df.DeathCross & (df.TwoMonthChange < 5)].DeathCross.count()/deathCount)*100
print(postiveFivePercentChange)
print(negativeFivePercentChange)
# Percentage of change of 10% in Golden Cross and Death Cross
postiveTenPercentChange = (df[df.GoldenCross & (df.ThreeMonthChange > 10)].GoldenCross.count()/goldenCount)*100
negativeTenPercentChange = (df[df.DeathCross & (df.ThreeMonthChange < 10)].DeathCross.count()/deathCount)*100
print(postiveTenPercentChange)
print(negativeTenPercentChange)
# Percentage of change of 15% in Golden Cross and Death Cross
postiveFifPercentChange = (df[df.GoldenCross & (df.FourMonthChange > 15)].GoldenCross.count()/goldenCount)*100
negativeFifPercentChange = (df[df.DeathCross & (df.FourMonthChange < 15)].DeathCross.count()/deathCount)*100
print(postiveFifPercentChange)
print(negativeFifPercentChange)
# Percentage of change of 20% in Golden Cross and Death Cross
postiveTwenPercentChange = (df[df.GoldenCross & (df.FiveMonthChange > 20)].GoldenCross.count()/goldenCount)*100
negativeTwenPercentChange = (df[df.DeathCross & (df.FiveMonthChange < 20)].DeathCross.count()/deathCount)*100
print(postiveTwenPercentChange)
print(negativeTwenPercentChange)
# -
| case3/case3-priya(new).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
# %matplotlib inline
iris= sns.load_dataset('iris')
iris.head()
sns.scatterplot(data= iris, x= "sepal_length", y= "petal_length", hue= "species");
numeric_cols= ["sepal_length", "sepal_width", "petal_length", "petal_width"]
x= iris[numeric_cols]
# ### K Means Clustering
#
# The K-means algorithm attempts to classify objects into a pre-determined number of clusters by finding optimal central points (called centroids) for each cluster. Each object is classifed as belonging the cluster represented by the closest centroid.
#
# <img src="https://miro.medium.com/max/1400/1*rw8IUza1dbffBhiA4i0GNQ.png" width="640">
from sklearn.cluster import KMeans
model= KMeans(n_clusters= 3, random_state= 42)
model.fit(x)
model.cluster_centers_
x
preds= model.predict(x)
preds
sns.scatterplot(data= x, x= "sepal_length", y= "petal_length", hue= preds)
centers_x, centers_y= model.cluster_centers_[:,0], model.cluster_centers_[:,2]
plt.plot(centers_x, centers_y, "xb");
# As you can see, K-means algorithm was able to classify (for the most part) different specifies of flowers into separate clusters. Note that we did not provide the "species" column as an input to `KMeans`.
# We can check the "goodness" of the fit by looking at `model.inertia_`, which contains the sum of squared distances of samples to their closest cluster center. Lower the inertia, better the fit.
model.inertia_
# Let's try creating 6 clusters.
model= KMeans(n_clusters= 6, random_state= 42).fit(x)
preds= model.predict(x)
preds
sns.scatterplot(data= x, x= "sepal_length", y= "petal_length", hue= preds);
# In most real-world scenarios, there's no predetermined number of clusters. In such a case, you can create a plot of "No. of clusters" vs "Inertia" to pick the right number of clusters.
# +
options = range(2,11)
inertias= []
for n_clusters in options:
model= KMeans(n_clusters, random_state= 42).fit(x)
inertias.append(model.inertia_)
plt.title("No. of clusters vs. Inertia")
plt.plot(options, inertias, "-o")
plt.xlabel('No. of clusters (K)')
plt.ylabel('Inertia');
# -
# The chart is creates an "elbow" plot, and you can pick the number of clusters beyond which the reduction in inertia decreases sharply.
# **Mini Batch K Means**: The K-means algorithm can be quite slow for really large dataset. Mini-batch K-means is an iterative alternative to K-means that works well for large datasets. Learn more about it here: https://scikit-learn.org/stable/modules/clustering.html#mini-batch-kmeans
# ### DBSCAN
#
# Density-based spatial clustering of applications with noise (DBSCAN) uses the density of points in a region to form clusters. It has two main parameters: "epsilon" and "min samples" using which it classifies each point as a core point, reachable point or noise point (outlier).
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/a/af/DBSCAN-Illustration.svg/800px-DBSCAN-Illustration.svg.png" width="400">
#
# Here's a video explaining how the DBSCAN algorithm works: https://www.youtube.com/watch?v=C3r7tGRe2eI
from sklearn.cluster import DBSCAN
model= DBSCAN(eps= 1.1, min_samples= 4)
model = DBSCAN(eps= 1.1, min_samples= 4)
model.fit(x)
# In DBSCAN, there's no prediction step. It directly assigns labels to all the inputs.
model.labels_
sns.scatterplot(data= x, x= "sepal_length", y= "petal_length", hue= model.labels_);
model= DBSCAN(eps= 2.2, min_samples= 8)
model.fit(x)
model.labels_
sns.scatterplot(data= x, x= "sepal_length", y= "petal_length", hue= model.labels_);
model= DBSCAN(eps= 0.5, min_samples= 2)
model.fit(x)
model.labels_
sns.scatterplot(data= x, x= "sepal_length", y= "petal_length", hue= model.labels_);
# Here's how the results of DBSCAN and K Means differ:
#
# <img src="https://miro.medium.com/max/1339/0*xu3GYMsWu9QiKNOo.png" width="640">
# ### Hierarchical Clustering
#
# Hierarchical clustering, as the name suggests, creates a hierarchy or a tree of clusters.
#
# <img src="https://dashee87.github.io/images/hierarch.gif" width="640">
#
# While there are several approaches to hierarchical clustering, the most common approach works as follows:
#
# 1. Mark each point in the dataset as a cluster.
# 2. Pick the two closest cluster centers without a parent and combine them into a new cluster.
# 3. The new cluster is the parent cluster of the two clusters, and its center is the mean of all the points in the cluster.
# 3. Repeat steps 2 and 3 till there's just one cluster left.
#
# Watch this video for a visual explanation of hierarchical clustering: https://www.youtube.com/watch?v=7xHsRkOdVwo
# ## Dimensionality Reduction and Manifold Learning
#
# In machine learning problems, we often encounter datasets with a very large number of dimensions (features or columns). Dimensionality reduction techniques are used to reduce the number of dimensions or features within the data to a manageable or convenient number.
#
#
# Applications of dimensionality reduction:
#
# * Reducing size of data without loss of information
# * Training machine learning models efficiently
# * Visualizing high-dimensional data in 2/3 dimensions
# ### Principal Component Analysis (PCA)
#
# Principal component is a dimensionality reduction technique that uses linear projections of data to reduce their dimensions, while attempting to maximize the variance of data in the projection. Watch this video to learn how PCA works: https://www.youtube.com/watch?v=FgakZw6K1QQ
#
# Here's an example of PCA to reduce 2D data to 1D:
#
# <img src="https://i.imgur.com/ZJ7utlo.png" width="480">
#
# Here's an example of PCA to reduce 3D data to 2D:
#
# <img src="https://lihan.me/assets/images/pca-illustration.png" width="640">
#
#
# Let's apply Principal Component Analysis to the Iris dataset.
iris= sns.load_dataset("iris")
iris
numeric_cols
from sklearn.decomposition import PCA
pca= PCA(n_components= 2)
pca.fit(iris[numeric_cols])
pca
transformed= pca.transform(iris[numeric_cols])
sns.scatterplot(x= transformed[:,0], y= transformed[:,1], hue= iris["species"]);
# As you can see, the PCA algorithm has done a very good job of separating different species of flowers using just 2 measures.
# ### t-Distributed Stochastic Neighbor Embedding (t-SNE)
#
# Manifold learning is an approach to non-linear dimensionality reduction. Algorithms for this task are based on the idea that the dimensionality of many data sets is only artificially high. Scikit-learn provides many algorithms for manifold learning: https://scikit-learn.org/stable/modules/manifold.html . A commonly-used manifold learning technique is t-Distributed Stochastic Neighbor Embedding or t-SNE, used to visualize high dimensional data in one, two or three dimensions.
#
# Here's a visual representation of t-SNE applied to visualize 2 dimensional data in 1 dimension:
#
# <img src="https://i.imgur.com/rVMAaix.png" width="360">
#
#
# Here's a visual representation of t-SNE applied to the MNIST dataset, which contains 28px x 28px images of handrwritten digits 0 to 9, a reduction from 784 dimensions to 2 dimensions ([source](https://colah.github.io/posts/2014-10-Visualizing-MNIST/)):
#
# <img src="https://indico.io/wp-content/uploads/2015/08/mnist-1024x607-1.jpg" width="640">
#
# Here's a video explaning how t-SNE works: https://www.youtube.com/watch?v=NEaUSP4YerM
from sklearn.manifold import TSNE
tsne= TSNE(n_components= 2)
transformed= tsne.fit_transform(iris[numeric_cols])
sns.scatterplot(x= transformed[:,0], y= transformed[:,1], hue= iris["species"]);
| Unsupervised Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Let's use the Trapezoid Method, Simpson's Rule and Romberg Integration
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# ### Define a function for taking an integral
def func(x):
a = 1.01
b = -3.04
c = 2.07
return a*x**2 + b*x + c
# ### Define it's integral so we know the right answer
def func_integral(x):
a = 1.01
b = -3.04
c = 2.07
return (a*x**3)/3. + (b*x**2)/2. + c*x
# ## Define the core of the trapezoid method
def trapezoid_core(f,x,h):
return 0.5*h*(f(x+h) + f(x))
# ### Define a wrapper function to perform trapezoid method
def trapezoid_method(f,a,b,N):
#f == function to integrate
#a == lower limit of integration
#b == upper limit of integration
#N == number of functions
#note; number of chunks will be N-1
#so if the N is odd, then we don't need to adjust the
#last segment
#define x values to perform simpsons rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
#define the value of the integral
Fint = 0.0
#perform the integral using the trapezoid method
for i in range(0,len(x)-1,1):
Fint += trapezoid_core(f,x[i],h)
#print(i,i+1,x[i]+h,x[-2],x[-1])
#return the answer
return Fint
# ### Define the core of the Simpson's Method
def simpson_core(f,x,h):
return h*( f(x) + 4*f(x+h) + f(x+2*h))/3
# ### Define a wrapper function to perform Simpson's Method
def simpsons_method(f,a,b,N):
#f == function to integrate
#a == lower lim of integration
#b == upper lim of integration
#N == number of function evaluations to use
#NOTE: the number of chunks will be N-1
#so if N is odd, then we don't need to adjust the last segment
#define x values to perform simpsons rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
#def the value of the integral
Fint = 0.0
#perform the integral using the simpson's method
for i in range(0,len(x)-2,2):
Fint += simpson_core(f,x[i],h)
#apply simpson's rule over the last interval
#if N is even
if((N%2)==0):
Fint += simpson_core(f,x[-2],0.5*h)
return Fint
# ## Define the Romberg integration core
def romberg_core(f,a,b,i):
#we need the difference b-a
h = b-a
#and the increment between new func evals
dh = h/2.**(i)
#we need a cofactor
K = h/2.**(i+1)
#and the function evaluations
M = 0.0
for j in range(2**i):
M += f(a + 0.5*dh + j*dh)
#return the answer
return K*M
# ## Define a wrapper function to perform Romberg Integration
def romberg_integration(f,a,b,tol):
#define an iteration variable
i = 0
#define a maximum number of iterations
imax = 1000
#define an error estimate, set to a large value
delta = 100.0*np.fabs(tol)
#set an array of integral answer
I = np.zeros(imax,dtype=float)
#get the zeroth romberg iteration
I[0] = 0.5*(b-a)*(f(a)+ f(b))
#iterate by 1
i += 1
while(delta>tol):
#find this romberg iteration
I[i] = 0.5*I[i-1] + romberg_core(f,a,b,i)
#compute the new fractional error
delta = np.fabs( (I[i]-I[i-1])/I[i])
print(i,I[i],I[i-1],delta)
if (delta>tol):
#iterate
i+=1
#check max iterations
if(i>imax):
print("Max iterations reached.")
raise StopIteration('Stopping iterations after ',i)
#return the answer
return I[i]
# ## Find out the answer
Answer = func_integral(1)-func_integral(0)
print("Answer")
print(Answer)
print("Trapezoid")
print(trapezoid_method(func,0,1,10))
print("Simpson's Method")
print(simpsons_method(func,0,1,10))
print("Romberg")
tolerance = 1.0e-6
RI = romberg_integration(func,0,1,tolerance)
print(RI, (RI-Answer)/Answer, tolerance)
| integration_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
from matplotlib import style
# line graphs
# plotting to our canvas, generated our own values
plt.plot([1, 2, 3], [4, 5, 1])
# showing what we plotted
plt.show()
# adding title and labels to the graph
x = [5, 8, 10]
y = [12, 16, 6]
plt.plot(x, y)
plt.title('Info')
plt.ylabel('Y axis')
plt.xlabel('X axis')
plt.show()
# adding style to our graph
style.use("ggplot")
x = [5, 8, 10]
y = [12, 16, 6]
x2 = [6, 9, 11]
y2 = [6, 15, 7]
plt.plot(x,y,'g',label='line one',linewidth=5)
plt.plot(x2,y2,'c',label="line two",linewidth=5)
plt.title("Epic Info")
plt.ylabel("Y axis")
plt.xlabel("X axis")
plt.legend()
plt.grid(True,color='k')
plt.show()
# bar graphs
# categorical variables - EXAMPLE: sex, race, cities, etc
plt.bar([1,3,5,7,9], [5,2,7,8,2], label="Example one")
plt.bar([2,4,6,8,10], [8,6,2,5,6], label="Example two", color='g')
plt.legend()
plt.xlabel('bar number')
plt.ylabel('bar height')
plt.title('Bar Graph!')
plt.show()
# histograms
# quantitative variables - EXAMPLE: age group, how age group contributes to population growth
population_ages = [22,55,62,45,21,22,34,42,42,4,99,102,110,120,121,122,130,111,115,112,80,75,65,54,44,43,42,48]
bins = [0,10,20,30,40,50,60,70,80,90,100,110,120,130]
plt.hist(population_ages, bins, rwidth=0.8)
plt.xlabel('X axis')
plt.ylabel('Y axis')
plt.title('Histogram')
plt.legend()
plt.show()
# scatterplots
# used to compare 2+ variables, looking for correlations or outgroups
x = [1,2,3,4,5,6,7,8]
y = [5,2,4,2,1,4,5,2]
plt.scatter(x,y,label='skitscat', color='k')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Scatter Plot')
plt.legend()
plt.show()
| code/descriptive_stats_demo/matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import myutils as my
import pandas as pd
import seaborn as sns
import numpy as np
from sklearn.manifold import TSNE
# %matplotlib inline
# %load_ext rpy2.ipython
df = pd.read_table("./data/glove.6B.100d.txt", delimiter=" ", header=None, index_col=0, quoting=3)
df = pd.DataFrame(pd.read_csv("./data/minimal.50d.3f.csv")).set_index("0")
lens = (df**2).sum(axis=1).sort_values()
dfn = df.div(np.sqrt(lens), axis='index')
godsies = my.similar(dfn, dfn.loc["god"], 40).drop(["god"])
godplot = my.get_with_axes(dfn, godsies.index, dfn.loc["god"], dfn.loc["good"]-dfn.loc["evil"]).rename(columns={"a":"god", "b":"evil___good"})
# + magic_args="-i godplot" language="R"
# library(ggplot2)
#
# ggplot(godplot, aes(god, evil___good, label=name)) +
# geom_text() +
# scale_x_log10()
# + magic_args="-i godplot" language="R"
# library(ggplot2)
#
# ggplot(godplot, aes(god, evil___good, label=name)) +
# geom_text() +
# scale_x_log10()
# -
| notebooks_julia/5_poniedzialek.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Vectors
# * links
# ## Helpers
# +
import math
def col2row(M, col):
return [M[i][col] for i in range(len(M))]
def row2col(M, row):
return [[num] for num in M[row]]
def get_shape(arr):
shape = []
while type(arr) in [list, np.ndarray]:
shape.append(len(arr))
arr = arr[0]
return tuple(shape)
def dotV(v1, v2):
# expecting two 1D vectors
assert len(v1) == len(v2)
return sum(a*b for a,b in zip(v1,v2))
def dotM(M1, M2):
# Columns of M1 == Rows of M2
assert len(M1[0]) == len(M2)
out = [ [0 for col in M2[0]] for row in M1]
for row in range(len(M1)):
for col in range(len(M2[0])):
v1 = M1[row]
v2 = col2row(M2, col)
out[row][col] = dotV(v1, v2)
return out
# -
# ## Vector Class
class Vector:
def __init__(self, coords):
self.coords = [float(c) for c in coords]
@property
def x(self):
return self.coords[0]
@property
def y(self):
return self.coords[1]
def dot(self, v):
assert len(v) == len(self.coords)
return sum(a*b for a,b in zip(self.coords, v.coords))
def cross(self, v):
pass
def __mul__(self, v):
if isinstance(v, (int, float)):
return Vector([a*v for a in self.coords])
return Vector([a*b for a,b in zip(self.coords, v.coords)])
def __rmul__(self, v):
return self.__mul__(v)
def __truediv__(self, v):
if isinstance(v, (int, float)):
return Vector([a/v for a in self.coords])
return Vector([a/b for a,b in zip(self.coords, v.coords)])
def __floordiv__(self, v):
if isinstance(v, (int, float)):
return Vector([a//v for a in self.coords])
return Vector([a//b for a,b in zip(self.coords, v.coords)])
def __add__(self, v):
if isinstance(v, (int, float)):
return Vector([a+v for a in self.coords])
return Vector([a+b for a,b in zip(self.coords, v.coords)])
def __sub__(self, v):
if isinstance(v, (int, float)):
return Vector([a-v for a in self.coords])
return Vector([a-b for a,b in zip(self.coords, v.coords)])
def __len__(self):
return len(self.coords)
def magnitude(self):
return math.sqrt(sum(c**2 for c in self.coords))
def normalize(self):
return Vector([c/w for c in self.coords])
def display(self):
return [round(c,2) for c in self.coords]
def __repr__(self):
return str(tuple(self.display()))
v = Vector((1,1))
v2 = Vector((2,2))
v.dot(v2)
len(v)
# ## Operations
def distancePoints(self, x1, y1, x2, y2):
pass
| theory/Vectors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Least Square
# +
import numpy as np
x = np.array([1, 2, 3])
y = np.array([23, 24, 27])
A = np.vstack([x, np.ones(len(x))]).T
b1, b2 = np.linalg.lstsq(A, y)[0]
print(b1,b2)
# -
import matplotlib.pyplot as plt
plt.plot(x, y, 'o', label='Original data', markersize=10)
plt.plot(x, b1*x + b2, 'r', label='Fitted line')
plt.legend()
plt.show()
# # ยกกำลังสองพจน์ $(x+y+c)^{2}$
# +
y1 = np.array([[-1,-1,23]])
y1sq = np.kron(y1,y1)
y2 = np.array([[-2,-1,24]])
y2sq = np.kron(y2,y2)
y3 = np.array([[-3,-1,27]])
y3sq = np.kron(y3,y3)
deltaS = y1sq+y2sq+y3sq
deltaS
# -
# # Solve 2 Variable Equation
#
# *Solve
# Solve the system of equations 3 * x0 + x1 = 9 and x0 + 2 * x1 = 8:*
a = np.array([[28,12], [12,6]])
b = np.array([304,148])
x = np.linalg.solve(a, b)
x
| 2 - Least square for linear model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import logging
import glob
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
import torch
from torch.utils import data
from torchvision import transforms
import lmdb
import pickle
import string
import random
# + [markdown] tags=[]
# # Data Loader
# +
from PIL import Image
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
class ImagesDataset(data.Dataset):
def __init__(self, dataset_folder="data/comprehensive_cars/images/*.jpg", size=64):
self.size = size
"Resize Image and only apply Horizontal Flip augmentation method"
self.transform = transforms.Compose([
transforms.Resize((size, size)),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
])
self.data_type = os.path.basename(dataset_folder).split(".")[-1]
import time
t0 = time.time()
print('Start loading file addresses ...')
images = glob.glob(dataset_folder)
random.shuffle(images)
t = time.time() - t0
print('done! time:', t)
print("Number of images found: %d" % len(images))
self.images = images
self.length = len(images)
def __getitem__(self, idx):
try:
buf = self.images[idx]
if self.data_type == 'npy':
img = np.load(buf)[0].transpose(1, 2, 0)
img = Image.fromarray(img).convert("RGB")
else:
img = Image.open(buf).convert('RGB')
img = self.transform(img)
data = {
'image': img
}
return data
except Exception as e:
print(e)
print("Warning: Error occurred when loading file %s" % buf)
return self.__getitem__(np.random.randint(self.length))
def __len__(self):
return self.length
# +
## PLOT IMAGES FROM DATASET
batch_size = 64
n_workers = 8
cars_dataset = ImagesDataset()
cars_loader = torch.utils.data.DataLoader(
cars_dataset, batch_size=batch_size, num_workers=n_workers, shuffle=True,
pin_memory=True, drop_last=True,
)
from torchvision.utils import make_grid
from torchvision.utils import save_image
def show_images(images, nmax=64):
fig, ax = plt.subplots(figsize=(8, 8))
ax.set_xticks([]); ax.set_yticks([])
ax.imshow(make_grid((images.detach()[:nmax]), nrow=8).permute(1, 2, 0))
for batch in cars_loader:
temp_batch = batch["image"]
show_images(temp_batch)
break
# + [markdown] tags=[]
# # Decoder
# Predicts volume density and color from 3D location, viewing direction, and latent code z
#
# Parameterize the object and background feature fields with multi-layer perceptrons (MLPs) which take as input a 3D point x and viewing direction d together with latent shape and appearance codes $z_s, z_a$ and output a density $\sigma$ and feature $f$. More specifically, we apply the positional encoding $\gamma (x)$ and the latent sahape code $z_s$/ This is followed by blocks of fully-connected layes with ReLU activation. We use 8 blocks with a hidden dimenstion of 128 and one skip connection to the fourth layer for the object feature field, and 4 blocks with a hidden dimension of 64 for the background feature field. We then project this to the first output, the one-dimensional density output $\sigma$/ In a second branch, we apply the positional encoding $\gamma$ to the viewing direction $d$, concatenate $\gamma(d)$ to thee latent appearence code $z_a$, and add it to the previous hidden features. We pass it through a single fully-coinnected layer with ReLU activation and project it to the second output $f$.
# -
class Decoder(nn.Module):
def __init__(self, hidden_size=128, n_blocks=8, n_blocks_view=1,
skips=[4], n_freq_posenc=10, n_freq_posenc_views=4,
z_dim=64, rgb_out_dim=128, downscale_p_by=2, **kwargs):
super().__init__()
self.downscale_p_by = downscale_p_by # downscale factor for input points before positional encdoing
# positional encoding
self.n_freq_posenc = n_freq_posenc # max freq for positional encoding of 3D Location
self.n_freq_posenc_views = n_freq_posenc_views # max freq for positional encoding of viewing direction
dim_embed = 3 * self.n_freq_posenc * 2 # Size of Positional Encoded 3D Location
dim_embed_view = 3 * self.n_freq_posenc_views * 2 # Size of Positional Encoded Viewing Direction
## Density Prediciton Layers
self.n_blocks = n_blocks # Number of Layers
self.hidden_size = hidden_size # hidden size of Decoder Network
self.z_dim = z_dim # Dimension of latent code z
self.fc_in = nn.Linear(dim_embed, hidden_size) # Input r(x) Positional Encoded 3D Location
self.fc_z = nn.Linear(self.z_dim, hidden_size) # Input z_s = latent shape code
self.blocks = nn.ModuleList([
nn.Linear(hidden_size, hidden_size) for i in range(self.n_blocks - 1) # MLP Layers for Density Prediction
])
### Skip Connection
self.skips = skips # where to add a skip connection
n_skips = sum([i in skips for i in range(n_blocks - 1)])
if n_skips > 0:
self.fc_z_skips = nn.ModuleList(
[nn.Linear(z_dim, hidden_size) for i in range(n_skips)]
)
self.fc_p_skips = nn.ModuleList([
nn.Linear(dim_embed, hidden_size) for i in range(n_skips)
])
### Volume Density
self.sigma_out = nn.Linear(hidden_size, 1) # Output sigma (Volume Density) at 8th Layer Before adding Direction
## Feature Prediction Layers
self.n_blocks_view = n_blocks_view # Number of View-dep Layers
self.rgb_out_dim = rgb_out_dim # Output Dimension of Feature Prediction
self.fc_z_view = nn.Linear(self.z_dim, hidden_size) # Input z_a = latent appearance code
self.feat_view = nn.Linear(hidden_size, hidden_size) # Input Density Prediction FC
self.fc_view = nn.Lienar(dim_embed_view, hidden_size) # Input r(d) Positional Encoded Viewing Direction
self.feat_out = nn.Linear(hidden_size, rgb_out_dim) # Output Feature Vector
def transform_points(self, p, views_False, downscale_p_by = 2):
##Positional Encoding of 3D Location and Viewing Direction
p = p / downscale_p_by # Normalize p between [-1, 1]
'''
n_freq_posenc (int), max freq for positional encdoing of 3D Location,
n_freq_posenc_views (int), max freq for positional encoding of Viewing Direction
'''
L = self.n_freq_posenc_views if views else self.n_freq_posenc
p_transformed = torch.cat([torch.cat(
[torch.sin((2 ** i) * np.pi * p),
torch.cos((2 ** i) * np.pi * p)],
dim=-1) for i in range(L)], dim = 1)
return p_transformed
def forward(self, p_in, ray_d, z_shape=None, z_app=None, **kwargs):
a = F.relu
if self.z_dim > 0:
batch_size = p_in.shape[0]
if z_shape is None:
z_shape = torch.randn(batch_size, self.z_dim).to(p_in.device)
if z_app is None:
z_app = torch.randn(batch_size, self.z_dim).to(p_in.device)
p = self.transform_points(p_in)
net = self.fc_in(p)
net = net + self.fc_z(z_shape).unsqueeze(1)
net = a(net)
skip_idx = 0
for idx, layer in enumerate(self.blocks):
net = a(layer(net))
if (idx+1) in self.skips and (idx < len(self.blocks) -1): # add skip connection
net = net + self.fc_z_skips[skip_idx](z_shape).unsqueeze(1)
net = net + self.fc_p_skips[skip_idx](p)
skip_idx += 1
sigma_out = self.sigma_out(net).squeeze(-1)
net = self.feat_view(net)
net = net + self.fc_z_view(z_app).unsqueeze(1)
ray_d = ray_d / torch.norm(ray_d, dim=-1, keepdim=True)
ray_d = self.transform_points(ray_d, views=True)
net = net + self.fc_view(ray_d)
net = a(net)
if self.n_blocks_view > 1:
for layer in self.blocks_view:
net = a(layer(net))
feat_out = self.feat_out(net)
return feat_out, sigma_out
# + [markdown] tags=[]
# # Bounding Box Generator
# Object Detection is used to predict how many objects are there.
# -
import numpy as np
import torch.nn as nn
import torch
from scipy.spatial.transform import Rotation as Rot
from im2scene.camera import get_rotation_matrix
# +
class BoundingBoxGenerator(nn.Module):
''' Bounding box generator class
Args:
n_boxes (int): number of bounding boxes (excluding background)
scale_range_min (list): min scale values for x, y, z
scale_range_max (list): max scale values for x, y, z
translation_range_min (list): min values for x, y, z translation
translation_range_max (list): max values for x, y, z translation
z_level_plane (float): value of z-plane; only relevant if
object_on_plane is set True
rotation_range (list): min and max rotation value (between 0 and 1)
check_collision (bool): whether to check for collisions
collision_padding (float): padding for collision checking
fix_scale_ratio (bool): whether the x/y/z scale ratio should be fixed
object_on_plane (bool): whether the objects should be placed on a plane
with value z_level_plane
prior_npz_file (str): path to prior npz file (used for clevr) to sample
locations from
'''
def __init__(self, n_boxes=1,
scale_range_min=[0.5, 0.5, 0.5],
scale_range_max=[0.5, 0.5, 0.5],
translation_range_min=[-0.75, -0.75, 0.],
translation_range_max=[0.75, 0.75, 0.],
z_level_plane=0., rotation_range=[0., 1.],
check_collison=False, collision_padding=0.1,
fix_scale_ratio=True, object_on_plane=False,
prior_npz_file=None, **kwargs):
super().__init__()
self.n_boxes = n_boxes
self.scale_min = torch.tensor(scale_range_min).reshape(1, 1, 3)
self.scale_range = (torch.tensor(scale_range_max) -
torch.tensor(scale_range_min)).reshape(1, 1, 3)
self.translation_min = torch.tensor(
translation_range_min).reshape(1, 1, 3)
self.translation_range = (torch.tensor(
translation_range_max) - torch.tensor(translation_range_min)
).reshape(1, 1, 3)
self.z_level_plane = z_level_plane
self.rotation_range = rotation_range
self.check_collison = check_collison
self.collision_padding = collision_padding
self.fix_scale_ratio = fix_scale_ratio
self.object_on_plane = object_on_plane
if prior_npz_file is not None:
try:
prior = np.load(prior_npz_file)['coordinates']
# We multiply by ~0.23 as this is multiplier of the original clevr
# world and our world scale
self.prior = torch.from_numpy(prior).float() * 0.2378777237835723
except Exception as e:
print("WARNING: Clevr prior location file could not be loaded!")
print("For rendering, this is fine, but for training, please download the files using the download script.")
self.prior = None
else:
self.prior = None
def check_for_collison(self, s, t):
n_boxes = s.shape[1]
if n_boxes == 1:
is_free = torch.ones_like(s[..., 0]).bool().squeeze(1)
elif n_boxes == 2:
d_t = (t[:, :1] - t[:, 1:2]).abs()
d_s = (s[:, :1] + s[:, 1:2]).abs() + self.collision_padding
is_free = (d_t >= d_s).any(-1).squeeze(1)
elif n_boxes == 3:
is_free_1 = self.check_for_collison(s[:, [0, 1]], t[:, [0, 1]])
is_free_2 = self.check_for_collison(s[:, [0, 2]], t[:, [0, 2]])
is_free_3 = self.check_for_collison(s[:, [1, 2]], t[:, [1, 2]])
is_free = is_free_1 & is_free_2 & is_free_3
else:
print("ERROR: Not implemented")
return is_free
def get_translation(self, batch_size=32, val=[[0.5, 0.5, 0.5]]):
n_boxes = len(val)
t = self.translation_min + \
torch.tensor(val).reshape(1, n_boxes, 3) * self.translation_range # compute translation parameters element-wise(x,y,z)
t = t.repeat(batch_size, 1, 1)
if self.object_on_plane:
t[..., -1] = self.z_level_plane
return t # return translation parameters for each
def get_rotation(self, batch_size=32, val=[0.]):
r_range = self.rotation_range
values = [r_range[0] + v * (r_range[1] - r_range[0]) for v in val]
# compute rotation value for all objects. (0 <= v <= 1, therefore the result is in the range)
r = torch.cat([get_rotation_matrix(
value=v, batch_size=batch_size).unsqueeze(1) for v in values],
dim=1)
r = r.float()
return r # return rotation matrices for each object
def get_scale(self, batch_size=32, val=[[0.5, 0.5, 0.5]]):
n_boxes = len(val)
if self.fix_scale_ratio:
t = self.scale_min + \
torch.tensor(val).reshape(
1, n_boxes, -1)[..., :1] * self.scale_range
else:
t = self.scale_min + \
torch.tensor(val).reshape(1, n_boxes, 3) * self.scale_range
t = t.repeat(batch_size, 1, 1)
return t
def get_random_offset(self, batch_size):
n_boxes = self.n_boxes
# Sample sizes
if self.fix_scale_ratio:
s_rand = torch.rand(batch_size, n_boxes, 1)
else:
s_rand = torch.rand(batch_size, n_boxes, 3)
s = self.scale_min + s_rand * self.scale_range
# Sample translations
if self.prior is not None:
idx = np.random.randint(self.prior.shape[0], size=(batch_size))
t = self.prior[idx]
else:
t = self.translation_min + \
torch.rand(batch_size, n_boxes, 3) * self.translation_range
if self.check_collison:
is_free = self.check_for_collison(s, t)
while not torch.all(is_free):
t_new = self.translation_min + \
torch.rand(batch_size, n_boxes, 3) * \
self.translation_range
t[is_free == 0] = t_new[is_free == 0]
is_free = self.check_for_collison(s, t)
if self.object_on_plane:
t[..., -1] = self.z_level_plane
def r_val(): return self.rotation_range[0] + np.random.rand() * (
self.rotation_range[1] - self.rotation_range[0])
R = [torch.from_numpy(
Rot.from_euler('z', r_val() * 2 * np.pi).as_dcm())
for i in range(batch_size * self.n_boxes)]
R = torch.stack(R, dim=0).reshape(
batch_size, self.n_boxes, -1).cuda().float()
return s, t, R
def forward(self, batch_size=32):
s, t, R = self.get_random_offset(batch_size)
R = R.reshape(batch_size, self.n_boxes, 3, 3)
return s, t, R # return scale parameters for each object(box)
# -
# # Neural Renderer
import torch.nn as nn
import torch
from math import log2
from im2scene.layers import Blur
class NeuralRenderer(nn.Module):
def __init__(
self, n_feat=128, input_dim=128, out_dim=3,
min_feat=32, img_size=64, use_rgb_skip=True,
upsample_feat="nn", upsample_rgb="bilinear",
**kwargs):
super().__init__()
self.input_dim = input_dim # input dimension, should be same as n_features
self.use_rgb_skip = use_rgb_skip # option to use RGB skip connections
n_blocks = int(log2(img_size) - 4)
assert(upsample_feat in ("nn", "bilinear"))
if upsample_feat == "nn":
self.upsample_2 = nn.Upsample(scale_factor=2.)
elif upsample_feat == "bilinear":
self.upsample_2 = nn.Sequential(nn.Upsample(
scale_factor=2, mode='bilinear', align_corners=False), Blur())
assert(upsample_rgb in ("nn", "bilinear"))
if upsample_rgb == "nn":
self.upsample_rgb = nn.Upsample(scale_factor=2.)
elif upsample_rgb == "bilinear":
self.upsample_rgb = nn.Sequential(nn.Upsample(
scale_factor=2, mode='bilinear', align_corners=False), Blur())
if n_feat == input_dim:
self.conv_in = lambda x: x
else:
self.conv_in = nn.Conv2d(input_dim, n_feat, 1, 1, 0)
self.conv_layers = nn.ModuleList(
[nn.Conv2d(n_feat, n_feat // 2, 3, 1, 1)] +
[nn.Conv2d(max(n_feat // (2 ** (i + 1)), min_feat),
max(n_feat // (2 ** (i + 2)), min_feat), 3, 1, 1)
for i in range(0, n_blocks - 1)]
)
if use_rgb_skip:
self.conv_rgb = nn.ModuleList(
[nn.Conv2d(input_dim, out_dim, 3, 1, 1)] +
[nn.Conv2d(max(n_feat // (2 ** (i + 1)), min_feat),
out_dim, 3, 1, 1) for i in range(0, n_blocks)]
)
else:
self.conv_rgb = nn.Conv2d(
max(n_feat // (2 ** (n_blocks)), min_feat), 3, 1, 1)
self.actvn = nn.LeakyReLU(0.2, inplace=True)
def forward(self, x):
net = self.conv_in(x)
if self.use_rgb_skip:
rgb = self.upsample_rgb(self.conv_rgb[0](x))
for idx, layer in enumerate(self.conv_layers):
hid = layer(self.upsample_2(net))
net = self.actvn(hid)
if self.use_rgb_skip:
rgb = rgb + self.conv_rgb[idx + 1](net)
if idx < len(self.conv_layers) - 1:
rgb = self.upsample_rgb(rgb)
if not self.use_rgb_skip:
rgb = self.conv_rgb(net)
rgb = torch.sigmoid(rgb)
return rgb
# # Generator
import torch.nn as nn
import torch.nn.functional as F
import torch
import numpy as np
from scipy.spatial.transform import Rotation as Rot
# +
from im2scene.common import arange_pixels, image_points_to_world, origin_to_world
from im2scene.camera import get_camera_mat, get_random_pose, get_camera_pose
class Generator(nn.Module):
def __init__(self, z_dim=256, z_dim_bg = 128,
range_u=(0, 0), range_v=(0.25, 0.25), n_ray_samples=64,
resolution_vol=16, fov=49.13,
range_radius=(2.732, 2.732), depth_range=[0.5, 6.], background_rotation_range=[0., 0.]):
self.device = "cuda" if torch.cuda.is_available() else "cpu"
self.n_ray_samples = n_ray_samples # Number of samples per ray (= N_s)
self.range_u = range_u # rotation range
self.range_v = range_v # elevation range
self.resolution_vol = resolution_vol # resolution of volume-rendered image
self.fov = fov # field of view
self.z_dim = z_dim # dimension of latent code z
self.z_dim_bg = z_dim_bg # dimension of background latent code z_bg
self.range_radius = range_radius # radius range
self.depth_range = depth_range # near and far depth plane (t_n, t_f)
self.bg_rotation_range = background_rotation_range # background rotation range
self.camera_matrix = get_camera_mat(fov=fov).to(self.device)
'''
camera matrix is a matrix which descirbes the mapping of a pinhole camera
from 3D points in the world to 2D points in an image
'''
self.decoder = Decoder().to(self.device)
self.background_generator = Decoder(hidden_size = 64, n_blocks = 4, downscale_p_by = 12, skips = []).to(self.device)
self.bounding_box_generator = BoundingBoxGenerator().to(self.device)
self.neural_renderer = NeuralRenderer().to(self.device)
def forward(self, batch_size=32, latent_codes=None, camera_matrices=None,
transformations=None, bg_rotation=None, mode="training", it=0,
return_alpha_map=False):
if latent_codes is None:
'''
Latent Codes are sampled from N(0, I)
1) z_shape_obj: z_s for object latent shape code
2) z_app_obj: z_a for object latent appearance code
3) z_shape_bg: z_s for background latent shape code
4) z_app_obj: z_a for object latent appearance code
'''
n_boxex = self.get_n_boxes()
z_shape_obj = torch.randn((batch_size, n_boxes, self.z_dim)).to(self.device)
z_app_obj = torch.randn((batch_size, n_boxes, self.z_dim)).to(self.device)
z_shape_bg = torch.randn((batch_size, self.z_dim_bg)).to(self.device)
z_app_bg = torch.randn((batch_size, self.z_dim_bg)).to(self.device)
latent_codes = (z_shape_obj, z_app_obj, z_shape_bg, z_app_bg)
if camera_matrices is None:
'''
Camera Intrinsics
Camera Pose ξ is sampled from p_ξ: Uniform distributions over dataset-dependent camera elevation angles
define the camera to be on a sphere with radius 2.732, and use t_n=0.5 and t_f=6
'''
camera_mat = self.camera_matrix.repeat(batch_size, 1, 1) # size (32, 4, 4)
world_mat = get_random_pose(self.range_u, self.range_v, self.range_radius, batch_size).to(self.device) # size (32, 4, 4)
camera_matrices = (camera_mat, world_mat)
if transformations is None:
'''
Affine Transformation is used to transform points from object to scene space.
Sampled from p_T: uniform distributions over valid object transformations
s = scale parameter
t = translation parameter
R = rotation matrix
k(x) = R * [s] * x + t
'''
s, t, R = self.bounding_box_generator(batch_size)
s, t, R = s.to(self.device), t.to(self.device), R.to(self.device)
transformations = (s, t, R)
if bg_rotation is None:
'''
Same representation for the background as for objects
but fix the scale and translation parameters s_N and t_N to span the entire scene and centered in origin
'''
if self.backround_rotation_range != [0., 0.]:
bg_r = self.backround_rotation_range
r_random = bg_r[0] + np.random.rand() * (bg_r[1] - bg_r[0])
R_bg = [
torch.from_numpy(Rot.from_euler(
'z', r_random * 2 * np.pi).as_dcm()
) for i in range(batch_size)]
R_bg = torch.stack(R_bg, dim=0).reshape(
batch_size, 3, 3).float()
else:
R_bg = torch.eye(3).unsqueeze(0).repeat(batch_size, 1, 1).float()
R_bg = R_bg.to(self.device)
bg_rotation = R_bg
if return_alpha_map:
rgb_v, alpha_map = self.volume_render_image(
latent_codes, camera_matrices, transformations, bg_rotation,
mode=mode, it=it, return_alpha_map=True,
not_render_background=not_render_background)
return alpha_map
else:
rgb_v = self.volume_render_image(
latent_codes, camera_matrices, transformations, bg_rotation,
mode=mode, it=it, not_render_background=not_render_background,
only_render_background=only_render_background)
if self.neural_renderer is not None:
rgb = self.neural_renderer(rgb_v)
else:
rgb = rgb_v
return rgb
def volume_render_image(self, latent_codes, camera_matrices,
transformations, bg_rotation, mode= "training",
it=0, return_alpha_map=False):
res = self.resolution_vol #
device = self.device
n_steps = self.n_ray_samples
n_points = res * res
depth_range = self.depth_range
batch_size = latent_codes[0].shape[0]
z_shape_obj, z_app_obj, z_shape_bg, z_app_bg = latent_codes
# Arrange Pixels
'''
function: arrange_pixels
Returns the Tupe of [0] Unscaled pixel locations as integers and [1] scaled float values
Then change order of the y axis of each scaled float value.
'''
pixels = arange_pixels((res, res), batch_size, invert_y_axis=False)[1].to(device) # Tensor [batch_size, res**2, 2]
pixels[..., -1] *= -1.
# Project to 3D world
'''
image_points_to_world
Transforms points on image plane to world coordinates. Fixed Depth of 1
origin_to_world
Transforms origin (camera location) to world coordinates.
'''
pixels_world = image_points_to_world(
pixels, camera_mat=camera_matrices[0], world_mat=camera_matrixes[1]
)
camera_world = origin_to_world(
n_points, camera_mat = camera_matrices[0], world_mat=camera_matrices[1]
)
ray_vector = pixels_world - camera_world
# batch_size x n_points x n_steps
di = depth_range[0] + torch.linspace(0., 1., steps=n_steps).reshape(1, 1, -1) * (
depth_range[1] - depth_range[0])
di = di.repeat(batch_size, n_points, 1).to(device)
if mode == "training":
di = self.add_noise_to_interval(di)
n_boxes = letent_codes[0].shape[1]
feat, sigma = [], [] # feature, sigma for each object
n_iter = n_boxes + 1
for i in range(n_iter):
if i < n_boxes: # Object
'''
p_i = input point x = (x, y, z)
r_i = viewing direction d
sigma_i = shape code
feat_i = appearance code
'''
p_i, r_i = self.get_evaluation_points(pixels_world, camera_world, di, transformations, i)
z_shape_i, z_app_i = z_shape_obj[:, i], z_app_obj[:, i]
feat_i, sigma_i = self.decoder(p_i, r_i, z_shape, z_app_i) #Decoder: Obtain feature (128 vector) and density(scalar)
if mode == "training":
sigma_i += torch.randn_like(sigma_i) # add noise during training
# Mask out values outside
padd = 0.1
mask_box = torch.all(p_i <=1. + padd, dim=-1) & torch.all(p_i >= -1.-padd, dim =-1)
sigma_i[mask_box == 0] = 0.
#Reshape
sigma_i = sigma_i.reshape(batch_size, n_points, n_steps) # shape = (32, 256, 64)
feat_i = feat_i.reshape(batch_size, n_points, n_steps, -1) # shape = (32, 256, 64, 128)
else: # Background
p_bg, r_bg = self.get_evaluation_points_bg(
pixels_world, camera_world, di, bg_rotation)
feat_i, sigma_i = self.background_generator(
p_bg, r_bg, z_shape_bg, z_app_bg)
sigma_i = sigma_i.reshape(batch_size, n_points, n_steps)
feat_i = feat_i.reshape(batch_size, n_points, n_steps, -1)
if mode == 'training':
sigma_i += torch.randn_like(sigma_i) # add noise during training
feat.append(feat_i)
sigma.append(sigma_i)
sigma = F.relu(torch.stack(sigma, dim=0))
feat = torch.stack(feat, dim=0)
# Composition Operator
'''
When Combining non-solid objects, sum up individual densites and use the density-weighted mean to combine all features at (x, d)
'''
denom_sigma = torch.sum(sigma, dim=0, keepdim=True)
denom_sigma[denom_sigma == 0] = 1e-4
w_sigma = sigma / denom_sigma
sigma_sum = torch.sum(sigma, dim=0)
feat_weighted = (feat * w_sigma.unsqueeze(-1)).sum(0)
# Get Volume Weights
weights = self.calc_volume_weights(di, ray_vector, sigma_sum)
feat_map = torch.sum(weights.unsqueeze(-1) * feat_weighted, dim=-2) # f = sum(gamma_j * alpha_j * feature_j)
# Reformat Output
feat_map = feat_map.permute(0, 2, 1).reshape(
batch_size, -1, res, res) # Batch x features x h x w
feat_map = feat_map.permute(0, 1, 3, 2) # new to flip x/y
return feat_map
'''
_______________________________________________________________________________________________
_______________________________________________________________________________________________
_______________________________________________________________________________________________
'''
def calc_volume_weights(self, z_vals, ray_vector, sigma, last_dist=1e10):
dists = z_vals[..., 1:] - z_vals[..., :-1] # delta_j: distance between neigboring sample points
dists = torch.cat([dists, torch.ones_like(
z_vals[..., :1]) * last_dist], dim=-1)
dists = dists * torch.norm(ray_vector, dim=-1, keepdim=True)
alpha = 1.-torch.exp(-F.relu(sigma)*dists) # alpha_j = 1 - exp(-sigma_j * delta_j)
weights = alpha * \
torch.cumprod(torch.cat([ # gamma = transmittance
torch.ones_like(alpha[:, :, :1]),
(1. - alpha + 1e-10), ], dim=-1), dim=-1)[..., :-1]
return weights
def add_noise_to_interval(self, di):
di_mid = .5 * (di[..., 1:] + di[..., :-1])
di_high = torch.cat([di_mid, di[..., -1:]], dim=-1)
di_low = torch.cat([di[..., :1], di_mid], dim=-1)
noise = torch.rand_like(di_low)
ti = di_low + (di_high - di_low) * noise
return ti
def get_n_boxes(self):
if self.bounding_box_generator is not None:
n_boxes = self.bounding_box_generator.n_boxes
else:
n_boxes = 1
return n_boxes
def transform_points_to_box(self, p, transformations, box_idx=0,
scale_factor=1.):
bb_s, bb_t, bb_R = transformations
p_box = (bb_R[:, box_idx] @ (p - bb_t[:, box_idx].unsqueeze(1)
).permute(0, 2, 1)).permute(
0, 2, 1) / bb_s[:, box_idx].unsqueeze(1) * scale_factor
return p_box
def get_evaluation_points(self, pixels_world, camera_world, di,
transformations, i):
batch_size = pixels_world.shape[0]
n_steps = di.shape[-1]
pixels_world_i = self.transform_points_to_box(
pixels_world, transformations, i)
camera_world_i = self.transform_points_to_box(
camera_world, transformations, i)
ray_i = pixels_world_i - camera_world_i
p_i = camera_world_i.unsqueeze(-2).contiguous() + \
di.unsqueeze(-1).contiguous() * ray_i.unsqueeze(-2).contiguous()
ray_i = ray_i.unsqueeze(-2).repeat(1, 1, n_steps, 1)
assert(p_i.shape == ray_i.shape)
p_i = p_i.reshape(batch_size, -1, 3)
ray_i = ray_i.reshape(batch_size, -1, 3)
return p_i, ray_i
def get_evaluation_points_bg(self, pixels_world, camera_world, di,
rotation_matrix):
batch_size = pixels_world.shape[0]
n_steps = di.shape[-1]
camera_world = (rotation_matrix @ camera_world.permute(0, 2, 1)).permute(0, 2, 1)
pixels_world = (rotation_matrix @ pixels_world.permute(0, 2, 1)).permute(0, 2, 1)
ray_world = pixels_world - camera_world
p = camera_world.unsqueeze(-2).contiguous() + \
di.unsqueeze(-1).contiguous() * \
ray_world.unsqueeze(-2).contiguous()
r = ray_world.unsqueeze(-2).repeat(1, 1, n_steps, 1)
assert(p.shape == r.shape)
p = p.reshape(batch_size, -1, 3)
r = r.reshape(batch_size, -1, 3)
return p, r
# + [markdown] tags=[]
# # Discriminator
# Discriminator for the GAN
# CNN with leaky ReLU activation
# -
class DCDiscriminator(nn.Module):
def __init__(self, in_dim=3, n_feat=512, img_size=64):
super(DCDiscriminator, self).__init__()
'''
in_dim (int): input dimension
n_feat (int): features of final hidden layer
img_size (int): input image size
'''
self.in_dim = in_dim
n_layers = int(log2(img_size) - 2)
self.blocks = nn.ModuleList(
[nn.Conv2d(
in_dim,
int(n_feat / (2 ** (n_layers - 1))),
4, 2, 1, bias=False)] + [nn.Conv2d(
int(n_feat / (2 ** (n_layers - i))),
int(n_feat / (2 ** (n_layers - 1 - i))),
4, 2, 1, bias=False) for i in range(1, n_layers)])
self.conv_out = nn.Conv2d(n_feat, 1, 4, 1, 0, bias=False)
self.actvn = nn.LeakyReLU(0.2, inplace=True)
def forward(self, x, **kwargs):
batch_size = x.shape[0]
if x.shape[1] != self.in_dim:
x = x[:, :self.in_dim]
for layer in self.blocks:
x = self.actvn(layer(x))
out = self.conv_out(x)
out = out.reshape(batch_size, 1)
return out
# # Training
# +
out_dir = "out/cars"
backup_every: 1000000
lr = 0.0005
lr_d = 0.0001
# -
class GIRAFFE(nn.Module):
''' GIRAFFE model class.
Args:
device (device): torch device
discriminator (nn.Module): discriminator network
generator (nn.Module): generator network
generator_test (nn.Module): generator_test network
'''
def __init__(self, device=None,
discriminator=None, generator=None, generator_test=None,
**kwargs):
super().__init__()
if discriminator is not None:
self.discriminator = discriminator.to(device)
else:
self.discriminator = None
if generator is not None:2
self.generator = generator.to(device)
else:
self.generator = None
if generator_test is not None:
self.generator_test = generator_test.to(device)
else:
self.generator_test = None
def forward(self, batch_size, **kwargs):
gen = self.generator_test
if gen is None:
gen = self.generator
return gen(batch_size=batch_size)
def generate_test_images(self):
gen = self.generator_test
if gen is None:
gen = self.generator
return gen()
def to(self, device):
''' Puts the model to the device.
Args:
device (device): pytorch device
'''
model = super().to(device)
model._device = device
return model
| simple giraffe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Understanding Cluster Analysis with Python
#
# ## Introduction
#
# This notebook uses a simple simuation in Python to illustrate the principles of cluster analysis.his notebook performs the following:
#
# - Generates a synthetic data set comprised of a mixture of 7 Nomal distributions.
# - Plots the data.
# - Computes and plots side by side k-means and hierarchical or agglomerative cluster models for 2, 3, 4, 5, 6, 7 and 8 clusters.
#
# The results of these steps allows you to compare and contrast the results for the two clustering methods.
# ## Create the data set
#
# The code in the cell below computes a two-dimensional data set which is the mixure of 7 Normal distributions. Examine the `create_clusts` function and you can see the vectors of `x` and `y` coordinates of the centroids of each cluster along with a vector of the standard deviations of each Normal.
#
# Execute this code to generate the data set. **Note**, everytime you run this code new realizatons of the 7 Normal distributions will be created.
# +
def sim_clust_data(ox, oy, n, sd, clust):
import pandas as pd
import numpy.random as nr
x = nr.normal(loc = ox, scale = sd, size = n)
y = nr.normal(loc = oy, scale = sd, size = n)
cluster = [clust] * n
out = pd.DataFrame([x, y, cluster]).transpose()
out.columns = ['x', 'y', 'cluster']
return out
def create_clusts(n = 20):
import pandas as pd
oxs = [0, 1, 1, 2, 2, 3, 3]
oys = [0, 1, -1, 2, -2, 1, -1]
sds = [.5, .5, .5, .5, .5, 1, .5]
clust_data = sim_clust_data(oxs[0], oys[0], n, sds[0], 0)
i = 0
for i in range(1, 7):
temp = sim_clust_data(oxs[i], oys[i], n, sds[i], i)
clust_data = pd.concat([clust_data, temp])
return clust_data
clusters = create_clusts()
# -
# ## Plot the data set
#
# The code in the cell below plots the data set. A different color is used to denote each Normal distribution. Run this code and note the overlap in the data samples generated from the Normal distributions.
# %matplotlib inline
def plot_data(df):
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8, 6))
fig.clf()
ax = fig.gca()
clusts = df['cluster'].unique()
cl = ['b', 'r', 'k', 'y', 'g', 'm', 'darkorange', 'hotpink']
for i, cls in enumerate(clusts):
temp = df[df['cluster'] == cls]
temp.plot(kind = 'scatter', x = 'x', y = 'y', s = 40, color = cl[i], ax = ax)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_title('X vs. Y')
return 'Done'
plot_data(clusters)
# ## Compute and plot clusters
#
# With the data set created it is now time to compute and display the clusters. The code in the cell below does the following:
#
# - Loop over the numbers of clusters to be computed and displayed.
# - Compute the hierarchical or agglomerative cluster model for the number of clusters.
# - Compute the k-means model for the number of clusters.
# - Call the plot function to display the result.
#
# Between two and eight clusters is computed and displayed. Two clusters are the minimum a data set can be divided into. The maximum of eight clusters is one more than the seven Normal distributions used to generate the data set.
#
# Execute the code and compare the results for the k-means and hierarchical or agglomerative cluster models. Notice that in each case, the k-means and hierarchical models produce different results. Which result is more useful depends on the application.
# +
def plot_clusters(df, n_clust):
import matplotlib.pyplot as plt
clusts_a = df['assignment_a'].unique()
clusts_k = df['assignment_k'].unique()
cl = ['b', 'r', 'k', 'y', 'g', 'm', 'darkorange', 'hotpink']
fig = plt.figure(figsize=(10, 4))
fig.clf()
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
for clu_a, clu_k, col in zip(clusts_a, clusts_k, cl):
temp = df[df['assignment_a'] == clu_a]
temp.plot(kind = 'scatter', x = 'x', y = 'y', s = 40, color = col, ax = ax1)
temp = df[df['assignment_k'] == clu_k]
temp.plot(kind = 'scatter', x = 'x', y = 'y', s = 40, color = col, ax = ax2)
ax1.set_xlabel('X')
ax1.set_ylabel('Y')
ax1.set_title('Agglomerative Clustering for ' + str(n_clust) + ' clusters')
ax2.set_xlabel('X')
ax2.set_ylabel('Y')
ax2.set_title('K-Means Clustering for ' + str(n_clust) + ' clusters')
return 'Done'
def demo_clusts(clust_data, n = 20, linkage = 'ward'):
from sklearn.cluster import AgglomerativeClustering, KMeans
X = clust_data[['x','y']].as_matrix()
for i in range(2,9):
agg = AgglomerativeClustering(n_clusters = i, linkage = linkage)
clust_data['assignment_a'] = agg.fit_predict(X)
km = KMeans(n_clusters = i)
clust_data['assignment_k'] = km.fit_predict(X)
plot_clusters(clust_data, i)
return 'Done'
demo_clusts(clusters)
# -
| Labs/Module 6/Demo/ClusterDemo-Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 11: Rotation
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# If you want the figures to appear in the notebook,
# and you want to interact with them, use
# # %matplotlib notebook
# If you want the figures to appear in the notebook,
# and you don't want to interact with them, use
# # %matplotlib inline
# If you want the figures to appear in separate windows, use
# # %matplotlib qt5
# tempo switch from one to another, you have to select Kernel->Restart
# %matplotlib inline
from modsim import *
# -
# ### Rolling paper
#
# We'll start by loading the units we need.
radian = UNITS.radian
m = UNITS.meter
s = UNITS.second
# And creating a `Condition` object with the system parameters
condition = Condition(Rmin = 0.02 * m,
Rmax = 0.055 * m,
L = 47 * m,
duration = 130 * s)
# The following function estimates the parameter `k`, which is the increase in the radius of the roll for each radian of rotation.
def estimate_k(condition):
"""Estimates the parameter `k`.
condition: Condition with Rmin, Rmax, and L
returns: k in meters per radian
"""
unpack(condition)
Ravg = (Rmax + Rmin) / 2
Cavg = 2 * pi * Ravg
revs = L / Cavg
rads = 2 * pi * revs
k = (Rmax - Rmin) / rads
return k
# As usual, `make_system` takes a `Condition` object and returns a `System` object.
def make_system(condition):
"""Make a system object.
condition: Condition with Rmin, Rmax, and L
returns: System with init, k, and ts
"""
unpack(condition)
init = State(theta = 0 * radian,
y = 0 * m,
r = Rmin)
k = estimate_k(condition)
ts = linspace(0, duration, 101)
return System(init=init, k=k, ts=ts)
# Testing `make_system`
system = make_system(condition)
system
system.init
# Now we can write a slope function based on the differential equations
#
# $\omega = \frac{d\theta}{dt} = 10$
#
# $\frac{dy}{dt} = r \frac{d\theta}{dt}$
#
# $\frac{dr}{dt} = k \frac{d\theta}{dt}$
#
def slope_func(state, t, system):
"""Computes the derivatives of the state variables.
state: State object with theta, y, r
t: time
system: System object with r, k
returns: sequence of derivatives
"""
theta, y, r = state
unpack(system)
omega = 10 * radian / s
dydt = r * omega
drdt = k * omega
return omega, dydt, drdt
# Testing `slope_func`
slope_func(system.init, 0*s, system)
# Now we can run the simulation.
run_odeint(system, slope_func)
# And look at the results.
system.results.tail()
# Extracting one time series per variable (and converting `r` to radians):
thetas = system.results.theta
ys = system.results.y
rs = system.results.r * 1000
# Plotting `theta`
# +
plot(thetas, label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
# -
# Plotting `y`
# +
plot(ys, color='green', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
# -
# Plotting `r`
# +
plot(rs, color='red', label='r')
decorate(xlabel='Time (s)',
ylabel='Radius (mm)')
# -
# We can also see the relationship between `y` and `r`, which I derive analytically in the book.
# +
plot(rs, ys, color='purple')
decorate(xlabel='Radius (mm)',
ylabel='Length (m)',
legend=False)
# -
# And here's the figure from the book.
# +
subplot(3, 1, 1)
plot(thetas, label='theta')
decorate(ylabel='Angle (rad)')
subplot(3, 1, 2)
plot(ys, color='green', label='y')
decorate(ylabel='Length (m)')
subplot(3, 1, 3)
plot(rs, color='red', label='r')
decorate(xlabel='Time(s)',
ylabel='Radius (mm)')
savefig('chap11-fig01.pdf')
# -
# We can use interpolation to find the time when `y` is 47 meters.
T = interp_inverse(ys, kind='cubic')
t_end = T(47)
t_end
# At that point `r` is 55 mm, which is `Rmax`, as expected.
R = interpolate(rs, kind='cubic')
R(t_end)
# The total amount of rotation is 1253 rad.
THETA = interpolate(thetas, kind='cubic')
THETA(t_end)
# ### Unrolling
# For unrolling the paper, we need more units:
kg = UNITS.kilogram
N = UNITS.newton
# And a few more parameters in the `Condition` object.
condition = Condition(Rmin = 0.02 * m,
Rmax = 0.055 * m,
Mcore = 15e-3 * kg,
Mroll = 215e-3 * kg,
L = 47 * m,
tension = 2e-4 * N,
duration = 180 * s)
# `make_system` computes `rho_h`, which we'll need to compute moment of inertia, and `k`, which we'll use to compute `r`.
def make_system(condition):
"""Make a system object.
condition: Condition with Rmin, Rmax, Mcore, Mroll,
L, tension, and duration
returns: System with init, k, rho_h, Rmin, Rmax,
Mcore, Mroll, ts
"""
unpack(condition)
init = State(theta = 0 * radian,
omega = 0 * radian/s,
y = L)
area = pi * (Rmax**2 - Rmin**2)
rho_h = Mroll / area
k = (Rmax**2 - Rmin**2) / 2 / L / radian
ts = linspace(0, duration, 101)
return System(init=init, k=k, rho_h=rho_h,
Rmin=Rmin, Rmax=Rmax,
Mcore=Mcore, Mroll=Mroll,
ts=ts)
# Testing `make_system`
system = make_system(condition)
system
system.init
# Here's how we compute `I` as a function of `r`:
def moment_of_inertia(r, system):
"""Moment of inertia for a roll of toilet paper.
r: current radius of roll in meters
system: System object with Mcore, rho, Rmin, Rmax
returns: moment of inertia in kg m**2
"""
unpack(system)
Icore = Mcore * Rmin**2
Iroll = pi * rho_h / 2 * (r**4 - Rmin**4)
return Icore + Iroll
# When `r` is `Rmin`, `I` is small.
moment_of_inertia(system.Rmin, system)
# As `r` increases, so does `I`.
moment_of_inertia(system.Rmax, system)
# Here's the slope function.
def slope_func(state, t, system):
"""Computes the derivatives of the state variables.
state: State object with theta, omega, y
t: time
system: System object with Rmin, k, Mcore, rho_h, tension
returns: sequence of derivatives
"""
theta, omega, y = state
unpack(system)
r = sqrt(2*k*y + Rmin**2)
I = moment_of_inertia(r, system)
tau = r * tension
alpha = tau / I
dydt = -r * omega
return omega, alpha, dydt
# Testing `slope_func`
slope_func(system.init, 0*s, system)
# Now we can run the simulation.
run_odeint(system, slope_func)
# And look at the results.
system.results.tail()
# Extrating the time series
thetas = system.results.theta
omegas = system.results.omega
ys = system.results.y
# Plotting `theta`
# +
plot(thetas, label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
# -
# Plotting `omega`
# +
plot(omegas, color='orange', label='omega')
decorate(xlabel='Time (s)',
ylabel='Angular velocity (rad/s)')
# -
# Plotting `y`
# +
plot(ys, color='green', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
# -
# Here's the figure from the book.
# +
subplot(3, 1, 1)
plot(thetas, label='theta')
decorate(ylabel='Angle (rad)')
subplot(3, 1, 2)
plot(omegas, color='orange', label='omega')
decorate(ylabel='Angular velocity (rad/s)')
subplot(3, 1, 3)
plot(ys, color='green', label='y')
decorate(xlabel='Time(s)',
ylabel='Length (m)')
savefig('chap11-fig02.pdf')
# -
# ### Yo-yo
# **Exercise:** Simulate the descent of a yo-yo. How long does it take to reach the end of the string.
#
# I provide a `Condition` object with the system parameters:
#
# * `Rmin` is the radius of the axle. `Rmax` is the radius of the axle plus rolled string.
#
# * `Rout` is the radius of the yo-yo body. `mass` is the total mass of the yo-yo, ignoring the string.
#
# * `L` is the length of the string.
#
# * `g` is the acceleration of gravity.
condition = Condition(Rmin = 8e-3 * m,
Rmax = 16e-3 * m,
Rout = 35e-3 * m,
mass = 50e-3 * kg,
L = 1 * m,
g = 9.8 * m / s**2,
duration = 1 * s)
# Here's a `make_system` function that computes `I` and `k` based on the system parameters.
#
# I estimated `I` by modeling the yo-yo as a solid cylinder with uniform density ([see here](https://en.wikipedia.org/wiki/List_of_moments_of_inertia)). In reality, the distribution of weight in a yo-yo is often designed to achieve desired effects. But we'll keep it simple.
def make_system(condition):
"""Make a system object.
condition: Condition with Rmin, Rmax, Rout,
mass, L, g, duration
returns: System with init, k, Rmin, Rmax, mass,
I, g, ts
"""
unpack(condition)
init = State(theta = 0 * radian,
omega = 0 * radian/s,
y = L,
v = 0 * m / s)
I = mass * Rout**2 / 2
k = (Rmax**2 - Rmin**2) / 2 / L / radian
ts = linspace(0, duration, 101)
return System(init=init, k=k,
Rmin=Rmin, Rmax=Rmax,
mass=mass, I=I, g=g,
ts=ts)
# Testing `make_system`
system = make_system(condition)
system
system.init
# Write a slope function for this system, using these results from the book:
#
# $ r = \sqrt{2 k y + R_{min}^2} $
#
# $ T = m g I / I^* $
#
# $ a = -m g r^2 / I^* $
#
# $ \alpha = m g r / I^* $
#
# where $I^*$ is the augmented moment of inertia, $I + m r^2$.
#
# Hint: If `y` is less than 0, it means you have reached the end of the string, so the equation for `r` is no longer valid. In this case, the simplest thing to do it return the sequence of derivatives `0, 0, 0, 0`
# +
def slope_func(state, t, system):
"""
Calculates slope of variables.
state: state object
t: time
system: system object
returns slopes in a sequence
"""
# I don't know if I agree with this unpacking concept, but it makes for a lot less typing, so...
theta, omega, y, v = state
unpack(system)
# Just spit back the formulas.
r = sqrt(2*k*y + Rmin**2)
alpha = mass * g * r / (I + mass * r**2)
a = -r * alpha
# Quick check for y.
if y < 0:
return 0, 0, 0, 0
return omega, alpha, v, a
# -
# Test your slope function with the initial conditions.
slope_func(system.init, 0*s, system)
# Then run the simulation.
run_odeint(system, slope_func)
# Check the final conditions. If things have gone according to plan, the final value of `y` should be close to 0.
system.results.tail()
# Plot the results.
thetas = system.results.theta
ys = system.results.y
# `theta` should increase and accelerate.
# +
plot(thetas, label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
# -
# `y` should decrease and accelerate down.
# +
plot(ys, color='green', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
# -
| code/chap11mine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jana0601/AA_Summer-school-LMMS/blob/main/Lab_Session_ToyModels.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="j8Gtj00FhwrZ"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import scipy.linalg as scl
# + [markdown] id="sCn7UfcShwrb"
# In this notebook, we will apply the basic EDMD algorithm to analyze data from the linear stochastic differential equation:
# $$ \mathrm{d}X_t = -X_t \mathrm{d}t + \sigma(X_t) \mathrm{d}W_t $$
# + [markdown] id="jWZCfu-dhwrc"
# ### Simulation and Evolution of Densities
# Let us first define a numerical integrator (i.e. the machinery to produce data), and then have a look at the evolution of probability distributions with time.
# + id="lMj5neubhwrc"
# This function realizes the standard Euler scheme
# for a linear stochastic differential equation:
def Euler_Scheme(x0, sigma, dt, m):
# Prepare output:
y = np.zeros(m)
y[0] = x0
# Initialize at x0:
x = x0
# Integrate:
for kk in range(1, m):
# Update:
xn = x - dt * x + sigma * np.sqrt(dt)*np.random.randn()
# Update current state:
y[kk] = xn
x = xn
return y
# + [markdown] id="jyx7KM7uhwrd"
# First, use the above function to produce 1000 simulations, each comprising discrete 1000 steps, at integration time step 1e-2, starting at $x_0 = 2$. Produce a histogram of the data after [10, 20, 50, 100, 200, 500] steps.
# Then, repeat the experiment, but draw the initial condition from a normal distribution with mean zero, and standard deviation 0.5.
# + id="TiD60CCahwrd"
# Settings:
m = 1000
dt = 1e-2
ntraj = 1000
sigma = 1.0
# Generate data:
X = np.zeros((ntraj, m+1))
for l in range(ntraj):
x0 = np.random.randn(1)
X[l,:] = Euler_Scheme(x0,sigma,dt,m+1)
# + colab={"base_uri": "https://localhost:8080/", "height": 353} id="TxhDmWi_laTB" outputId="d2590a3c-a426-40eb-c114-29f74c8ce653"
plt.plot(X[:5,:].T)
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="Fxm29Nr1hwre" outputId="29d49f8c-755a-4e5b-c749-086775e97844"
# Time instances to be used for histogramming:
t_vec = np.array([10, 20, 50, 100, 200, 500])
# Bins for histogram:
xe = np.arange(-2.5, 3.51, 0.1)
xc = 0.5 * (xe[1:] + xe[:-1])
# Histogram the data at different time instances:
plt.figure()
qq = 0
for t in t_vec:
h, _ = np.histogram(X[:,t],bins=xe,density=True)
plt.plot(xc,h,label="%d"%t_vec[qq])
qq += 1
plt.plot(xc, (1.0/np.sqrt(2*np.pi *0.5))*np.exp(-xc**2), "k--")
plt.xlabel("x", fontsize=12)
plt.tick_params(labelsize=12)
plt.ylim([-.5, 1.5])
plt.legend(loc=2)
# + [markdown] id="FPltyWawhwre"
# ### Estimating the Koopman Operator
# + [markdown] id="vuPBwCFihwrf"
# First, write a function to compute a matrix approximation for the Koopman operator. Inputs should the raw data, the time shifted raw data, a callable function to realize the basis set, and the number of basis functions:
# + id="8d0p9VNwhwrf"
def koopman_matrix(X, Y, psi, n):
# Get info on data:
m = X.shape[0]
# Evaluate basis set on full data:
# Compute Koopman matrix:
return K
# + [markdown] id="XCdqDUCThwrg"
# Produce 10,000 pairs $(x_l, y_l)$ by drawing $x_l$ from the invariant measure of our linear SDE. Compute each $y_l$ by running the dynamics over time $t = 0.1$ (10 discrete time steps). Then, estimate the Koopman matrix for the monomial basis of degree 10.
# + id="PX1Hp5UZhwrg"
# Produce the data:
m = 10000
x = np.sqrt(0.5) * np.random.randn(m)
y = np.zeros(m)
# Define basis set:
n = 5
# Compute Koopman matrix:
# + [markdown] id="fEHyzreghwrg"
# ### Koopman-based Prediction
# Diagonalize the Koopman matrix. Use the spectral mapping theorem to predict the eigenvalues at times $[0.1, 0.2, 0.3, ..., 2.0]$. Compare to the analytical values: the $k$-th eigenvalue at lag time $t$ is given by $\exp(-k \cdot t)$.
# + id="8KcK8SGShwrh"
# Diagonalize K:
d, V =
# Sort eigenvalues and eigenvectors:
# Plot eigenvalues at multiple lag times:
lags = nsteps * np.arange(1, 21)
plt.figure()
for k in range(1, 4):
plt.plot(dt*lags, np.exp(- k * dt* lags), "x")
# + [markdown] id="aZKuwvQfhwrh"
# Use the Koopman matrix to predict the variance of the process at times $[0.1, 0.2, 0.3, ..., 2.0]$, if started at $x$, as a function of $x$. The variance is
# $\mathbb{E}^x[(X_t)^2]$, which equals the Koopman operator applied to the function $x^2$. Remember this function is contained in your basis set.
# + id="U_9_wR5whwrh"
# Coefficient vector of x**2 with respect to monomial basis:
b = np.eye(n)[:, 2]
# Prepare output:
lag_coeffs = np.zeros((n, lags.shape[0]))
# Repeatedly apply Koopman matrix to coefficient vector:
# Plot coefficients of the variance as a function of t:
for ii in range(n):
plt.plot(dt*lags, lag_coeffs[ii, :], "o--", label=r"$x^{%d}$"%ii)
plt.legend(loc=1)
# + id="SgfakyXFhwri"
| Data-Driven Modeling of Dynamical Systems/Lab_Session_ToyModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Basic-sequence-types" data-toc-modified-id="Basic-sequence-types-1"><span class="toc-item-num">1 </span>Basic sequence types</a></span><ul class="toc-item"><li><span><a href="#List" data-toc-modified-id="List-1.1"><span class="toc-item-num">1.1 </span>List</a></span></li><li><span><a href="#Tuple" data-toc-modified-id="Tuple-1.2"><span class="toc-item-num">1.2 </span>Tuple</a></span></li><li><span><a href="#Set" data-toc-modified-id="Set-1.3"><span class="toc-item-num">1.3 </span>Set</a></span></li><li><span><a href="#Dict" data-toc-modified-id="Dict-1.4"><span class="toc-item-num">1.4 </span>Dict</a></span></li></ul></li><li><span><a href="#Collections-module" data-toc-modified-id="Collections-module-2"><span class="toc-item-num">2 </span>Collections module</a></span><ul class="toc-item"><li><span><a href="#collections.namedtuple" data-toc-modified-id="collections.namedtuple-2.1"><span class="toc-item-num">2.1 </span>collections.namedtuple</a></span></li><li><span><a href="#collections.deque" data-toc-modified-id="collections.deque-2.2"><span class="toc-item-num">2.2 </span>collections.deque</a></span></li><li><span><a href="#collections.defaultdict" data-toc-modified-id="collections.defaultdict-2.3"><span class="toc-item-num">2.3 </span>collections.defaultdict</a></span></li><li><span><a href="#collections.Counter" data-toc-modified-id="collections.Counter-2.4"><span class="toc-item-num">2.4 </span>collections.Counter</a></span></li></ul></li></ul></div>
# -
# ## Basic sequence types
#
# - list, tuple, set, dict
#
# ### List
#
# - mutable (e.g. append)
# +
l = [4, 2, '4', '3', 4, 5.3, 6]
l[:3] # exclusive
l[3:] # inclusive
l[1:3]
l[1:5:2] # slice every 2nd from 1 to 5
l[:] = [1, 2, 3] # clear list and recreate with elements
first, second, *remainder, last = l # unpack list
l.append(7) # append inplace
l.pop() # list can be used as stack with append() + pop()
l.index(2) # returns index of first matching 2
l.count(4) # no 4's in list
l[:0] = [-2, -1] # prepend elements to list
l += [8, 9] # create new list from concatenated lists
3 * [0] # [0, 0, 0]
2 in l # True
[2, 3] in l # False
[2, 3] in [1, [2,3], 4] # True
[x**2 for x in [1, 2, 3]] # use list comprehension for modifying elements in list
sum(map(int,l))
# -
# ### Tuple
#
# - immutable
# - otherwise behaves similar to list
t = (4, 2, '4', '3', 4, 5.3, 6)
# ### Set
# +
s1 = {1, 2, 3}
s2 = {2, 4, 6}
s1 - s2 # s1.difference(s2)
s1 & s2 # s1.intersection(s2)
s1 | s2 # s1.union(s2)
s1 ^ s2 # XOR: s1.symmetric_difference(s2)
s1 >= s2 # s1.issuperset(s2)x
s1.issubset(s2) # False
# inplace add, remove, discard, pop, clear, ...
s1.add(10)
s1.isdisjoint(s2) # False due to 2
# -
# ### Dict
# +
# dict / hashmap
d = {1:'foo', 2:'bar'}
d = dict(zip([1, 2], ['foo', 'bar']))
d[1]
d.get(3) # None instead of KeyError
d.get(3, 'fallback')
1 in d # 1 in d.keys()
'foo' in d.values()
for (k, v) in d.items(): # unpacking
print(k, v)
# -
# ## Collections module
#
# ### collections.namedtuple
# +
from collections import namedtuple
Point = namedtuple('Point', ['x', 'y'])
p = Point(11, y=22)
p.x == p[0]
t = [1, 2]
p2 = Point(*t)
# -
# ### collections.deque
#
# - pronounced "deck"
# - stands for "double-ended queue"
# - implemented as doubly linked list (but uses list of arrays internally for performance)
# - FIFO: queue
# - LIFO: stack
# - faster than implementations in queue module but not synchronized
# - Java equivalent: `Queue<T> = new ArrayDeque<>();`
# +
from collections import deque
stack = deque([1, 2, 3])
stack.append(4)
last_in = stack.pop() # 4
stack.extend([5, 6, 7])
print(last_in)
print(stack[-1]) # peek right
queue = deque([1, 2, 3])
queue.append(4) # atomic operation
first_in = queue.popleft() # atomic operation
queue.extendleft([5, 4]) # order in queue is reverse of argument since items are inserted left one after another
print(first_in)
print(queue[0]) # peek left
limited_deque = deque(range(3), maxlen=3)
limited_deque.append(3) # since queue is full left element is dropped
limited_deque.appendleft(0) # right element is dropped
print(limited_deque)
# -
# ### collections.defaultdict
# - can be used to conveniently create a multimap
# - Java equivalent: `Map<K, List<V>>` or Guava `Multimap`
from collections import defaultdict
dd = defaultdict(list)
dd['key'].append('val1') # {'key': ['val1']}
dd['key'].append('val2') # {'key': ['val1', 'val2']}
# ### collections.Counter
# - can be used to conveniently create a [counting multiset](https://en.wikipedia.org/wiki/Multiset#Counting_multisets)
# - Java equivalent: `Map<E, Integer>` or Guava `Multiset`
# +
from collections import Counter
word_count = Counter('two times two four'.split()) # {'four': 1, 'times': 1, 'two': 2}
word_count.update(['four']) # {'four': 2, 'times': 1, 'two': 2}
# TODO several missing
| python-collections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day02_1_Classification_excercise01
#
#
#
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#이진-분류기" data-toc-modified-id="이진-분류기-1"><span class="toc-item-num">1 </span>이진 분류기</a></span><ul class="toc-item"><li><span><a href="#설정" data-toc-modified-id="설정-1.1"><span class="toc-item-num">1.1 </span>설정</a></span></li><li><span><a href="#모델링" data-toc-modified-id="모델링-1.2"><span class="toc-item-num">1.2 </span>모델링</a></span></li></ul></li><li><span><a href="#-분류의-성능-평가지표-
# --" data-toc-modified-id="-분류의-성능-평가지표-
# ---2"><span class="toc-item-num">2 </span><i class="fa fa-tasks"></i> 분류의 성능 평가지표
# </a></span></li></ul></div>
# -
# ## 이진 분류기
# ### 설정
# 먼저 몇 개의 모듈을 임포트합니다. 맷플롯립 그래프를 인라인으로 출력하도록 만들고 그림을 저장하는 함수를 준비합니다. 또한 파이썬 버전이 3.5 이상인지 확인합니다(파이썬 2.x에서도 동작하지만 곧 지원이 중단되므로 파이썬 3을 사용하는 것이 좋습니다). 사이킷런 버전이 0.20 이상인지도 확인합니다.
# +
# 파이썬 ≥3.5 필수
import sys
assert sys.version_info >= (3, 5)
# 사이킷런 ≥0.20 필수
import sklearn
assert sklearn.__version__ >= "0.20"
# 공통 모듈 임포트
import numpy as np
import os
# 노트북 실행 결과를 동일하게 유지하기 위해
np.random.seed(42)
# 깔끔한 그래프 출력을 위해
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# 그림을 저장할 위치
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "classification"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("그림 저장:", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# +
# data load
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1)
# X, y로 나누기
X, y = mnist["data"], mnist["target"]
# -
# train / test 나누기
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# 단일 분류를 학습하는 것이므로 5 또는 나머지 번호 두 분류로 나눈다.
y_train_5 = (y_train == 5)
y_test_5 = (y_test == 5)
# ### 모델링
# ```python
#
# from sklearn.neighbors import KNeighborsClassifier
# model_knn = KNeighborsClassifier(n_neighbors = 13) # 모델정의하기
# model_knn.fit(???,???) # 모델학습
# model_knn.score(???,???) # 모델점수보기
# model_knn.predict(???,???) # 모델 학습결과저장
#
# ```
from sklearn.neighbors import KNeighborsClassifier
# 모델정의하기
model = KNeighborsClassifier(n_neighbors = 13)
# 모델학습: X_train, y_train_5
model.fit(X_train,y_train_5)
# 모델의 예측값 저장
y_pred = model.predict(X_test)
# <div class="alert alert-success" data-title="">
# <h2><i class="fa fa-tasks" aria-hidden="true"></i> 분류의 성능 평가지표
# </h2>
# </div>
#
# ```python
# from sklearn.metrics import confusion_matrix # 오차행렬
# TN, FP, FN, TP = confusion_matrix(???, ???).ravel()
# ```
# 오차행렬
from sklearn.metrics import confusion_matrix # 오차행렬
TN, FP, FN, TP = confusion_matrix(y_test, y_pred).ravel()
# 정확도(Accuracy):예측한 결과가 실제 결과와 얼마나 동일한 지를 나타내는 지표
#
# - 직접구현: $\mbox{정확도(Accuracy)}=\frac{TP + TN}{TP+FN+FP+TN}$
#
# - sklearn사용:
# ```python
# from sklearn.metrics import accuracy_score
# accuracy_score(???, ???)
# ```
# [TO_DO] 정확도 구하기
# 1.패키지 사용
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
# 2. 직접 구하기
(TP + TN) / TP+FN+FP+TN
# 정밀도 (Precision): `Positive`로 예측한 결과들 중 실제로 `Positive`인 결과들의 비율
#
# - 직접구현: $\mbox{정밀도}=\frac{\mbox{TP}}{\mbox{TP+FP}}$
# - sklearn사용:
# ```python
# from sklearn.metrics import precision_score
# precision_score(???, ???)
# ```
# [TO_DO] 정밀도 구하기
# 1.패키지 사용
from sklearn.metrics import precision_score
precision_score(y_test, y_pred)
# 2. 직접구하기
TP / (TP+FP)
# 재현률 (Recall) : 실제로 Positive인 결과들 중 Positive로 예측한 결과들의 비율
#
# - 직접구현: $\mbox{재현율}=\frac{\mbox{TP}}{\mbox{TP+FN}}$
# - sklearn사용
# ```python
# from sklearn.metrics import precision_score
# recall_score(???, ???)
# ```
# [TO_DO] 재현률 구하기
# 1.패키지 사용
from sklearn.metrics import precision_score
recall_score(y_test, y_pred)
# 2. 직접구하기
TP / (TP + FN )
# F1 score : 정밀도와 재현율의 조화평균이다.
#
# - 직접구현: $\mbox{F}1=\frac{2}{\frac{1}{\mbox{정밀도}} + \frac{1}{\mbox{재현율}}}$
# - sklearn:
# ```python
# from sklearn.metrics import f1_score
# f1_score(???, ???)
# ```
# [TO_DO] f1 구하기
# 1.패키지 사용
from sklearn.metrics import f1_score
f1_score(y_test, y_pred)
# 2. 직접구현
2/ ((1/(TP / (TP+FP))) + (1/(TP / (TP + FN ))))
| code/Day02_answer/Day02_1_Classification_excercise01(KNN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import pylab as pl
import numpy as np
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
# %matplotlib inline
df = pd.read_csv(r'data\OnlineNewsPopularity.csv' , sep=r'\s*,\s*')
df.head()
cdf = df[["global_subjectivity","rate_positive_words","rate_negative_words",
"avg_positive_polarity","min_positive_polarity","max_positive_polarity","avg_negative_polarity","min_negative_polarity",
"max_negative_polarity","title_subjectivity","abs_title_sentiment_polarity","shares","data_channel_is_lifestyle","data_channel_is_entertainment","data_channel_is_bus",
"data_channel_is_socmed","data_channel_is_tech","data_channel_is_world",
"weekday_is_monday","weekday_is_tuesday","weekday_is_wednesday","weekday_is_thursday","weekday_is_friday",
"weekday_is_saturday","weekday_is_sunday"]]
cdf.head(9)
msk = np.random.rand(len(df)) < 0.8
train = cdf[msk]
test = cdf[~msk]
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn import linear_model
train_x = np.asanyarray(train[["global_subjectivity","rate_positive_words","rate_negative_words",
"avg_positive_polarity","min_positive_polarity","max_positive_polarity","avg_negative_polarity","min_negative_polarity",
"max_negative_polarity","title_subjectivity","abs_title_sentiment_polarity","data_channel_is_lifestyle","data_channel_is_entertainment","data_channel_is_bus",
"data_channel_is_socmed","data_channel_is_tech","data_channel_is_world",
"weekday_is_monday","weekday_is_tuesday","weekday_is_wednesday","weekday_is_thursday","weekday_is_friday",
"weekday_is_saturday","weekday_is_sunday"]])
train_y = np.asanyarray(train[["shares"]])
test_x = np.asanyarray(test[["global_subjectivity","rate_positive_words","rate_negative_words",
"avg_positive_polarity","min_positive_polarity","max_positive_polarity","avg_negative_polarity","min_negative_polarity",
"max_negative_polarity","title_subjectivity","abs_title_sentiment_polarity","data_channel_is_lifestyle","data_channel_is_entertainment","data_channel_is_bus",
"data_channel_is_socmed","data_channel_is_tech","data_channel_is_world",
"weekday_is_monday","weekday_is_tuesday","weekday_is_wednesday","weekday_is_thursday","weekday_is_friday",
"weekday_is_saturday","weekday_is_sunday"]])
test_y = np.asanyarray(test[["shares"]])
poly = PolynomialFeatures(degree=2)
train_x_poly = poly.fit_transform(train_x)
train_x_poly
# -
clf = linear_model.LinearRegression()
train_y_ = clf.fit(train_x_poly, train_y)
# The coefficients
print ('Coefficients: ', clf.coef_)
print ('Intercept: ',clf.intercept_)
# +
from sklearn.metrics import r2_score
test_x_poly = poly.fit_transform(test_x)
test_y_ = clf.predict(test_x_poly)
print("Mean absolute error: %.2f" % np.mean(np.absolute(test_y_ - test_y)))
print("Residual sum of squares (MSE): %.2f" % np.mean((test_y_ - test_y) ** 2))
print("R2-score: %.2f" % r2_score(test_y_ , test_y) )
# -
result = confusion_matrix(test_y, test_y_)
print("Confusion Matrix:")
print(result)
result1 = classification_report(test_y, test_y_)
print("Classification Report:",)
print (result1)
result2 = accuracy_score(test_y,test_y_)
print("Accuracy:",result2)
| PolynomialRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import re
import datefinder
from urllib.parse import urlparse
class FWFParser():
def __init__(self, dmp_title, dmp_description, dmp_created, dmp_modified, dmp_id, pi_name, pi_mail, pi_orcid):
self.setup_dmp(dmp_title, dmp_description, dmp_created, dmp_modified, dmp_id, pi_name, pi_mail, pi_orcid)
def setup_dmp(self, dmp_title, dmp_description, dmp_created, dmp_modified, dmp_id, pi_name, pi_mail, pi_orcid):
self.dmp = {
"dmp":{
"title": dmp_title,
"description":"Abstract::"+dmp_description,
"created":dmp_created,
"modified":dmp_modified,
"dmp_id": {
"dmp_id": dmp_id,
"dmp_id_type": "HTTP-DOI"
},
"contact":{
"name": pi_name,
"mail": pi_mail,
"contact_id": {
"contact_id": pi_orcid,
"contact_id_type": "HTTP-ORCID"
}
},
"project": {}
}
}
def __split_text_based_on_title(self, text):
rep = {}
for ds in self.dmp["dmp"]["dataset"]:
title = ds["title"].lower()
rep[title]=";"+title
rep = dict((re.escape(k), v) for k, v in rep.items())
pattern = re.compile("|".join(rep.keys()))
text = pattern.sub(lambda m: rep[re.escape(m.group(0))], text.lower()).split(";")
return text
def __split_text_based_on_fair(self, text):
rep = ["findable", "accessible"]
rep = dict((re.escape(k), v) for k, v in rep.items())
pattern = re.compile("|".join(rep.keys()))
text = pattern.sub(lambda m: rep[re.escape(m.group(0))], text.lower()).split(";")
return text
def parse_question_1_1(self, text):
rep = {"Title:": ";Title:", "Description:": ";Description:","Type:": ";Type:","Format:": ";Format:", "Source:":";Source:", "Size":";Size:"} # define desired replacements here
order = ['b', 'kb', 'mb', 'gb', 'tb', 'pb']
rep_lower = {}
for k, v in rep.items():
rep_lower[k.lower()]=v
rep.update(rep_lower)
rep = dict((re.escape(k), v) for k, v in rep.items())
pattern = re.compile("|".join(rep.keys()))
text = pattern.sub(lambda m: rep[re.escape(m.group(0))], q1_3_test_1)
datasets = []
current_dataset = None
current_format = None
current_size = None
for line in text.split(";"):
line = line.replace("\n","")
if line.startswith("Title:"):
line = line[7:] if line[6:].startswith(" ") else line[6:]
if current_dataset != None:
if "distribution" not in current_dataset:
current_dataset["distribution"] = []
current_dataset["distribution"].append({"title":"Project"})
else:
if current_format != None:
for ds in current_dataset["distribution"]:
ds["format"] = current_format
current_format = None
if current_size != None:
for ds in current_dataset["distribution"]:
ds["byte_size"] = current_size
current_size = None
current_dataset["metadata"] = []
current_dataset["technical_resource"] = []
datasets.append(current_dataset)
current_dataset = {"title":line}
if line.startswith("Description:"):
if current_dataset != None:
line = line[13:] if line[12:].startswith(" ") else line[12:]
current_dataset["description"]=line
if line.startswith("Type:"):
if current_dataset != None:
line = line[6:] if line[5:].startswith(" ") else line[5:]
current_dataset["type"]=line
if line.startswith("Format:"):
if current_dataset != None:
line = line[8:] if line[7:].startswith(" ") else line[7:]
current_format = line
if line.startswith("Size:"):
if current_dataset != None:
line = line[8:] if line[7:].startswith(" ") else line[7:]
regex1 = re.compile(r'(\d+(?:\.\d+)?)\s*([kmgtp]?b)', re.IGNORECASE)
regex2 = re.compile(r'(\d+(?:\.\d+)?)', re.IGNORECASE)
size1 = regex1.findall(line)
size2 = regex2.findall(line)
if len(size1) >= 1:
size1 = size1[0]
size1 = int(float(size1[0]) * (1024**order.index(size1[1].lower())))
current_size = size1
elif len(size2) >= 1:
current_size = size2
if line.startswith("Source:"):
if current_dataset != None:
line = line[8:] if line[7:].startswith(" ") else line[7:]
if line.lower() == "input":
current_dataset["distribution"] = []
current_dataset["distribution"].append({"title":"Origin"})
current_dataset["distribution"].append({"title":"Project"})
elif line.lower() == "produced":
current_dataset["distribution"] = []
current_dataset["distribution"].append({"title":"Project"})
if current_dataset != None:
if "distribution" not in current_dataset:
current_dataset["distribution"] = []
current_dataset["distribution"].append({"title":"Project"})
else:
if current_format != None:
for ds in current_dataset["distribution"]:
ds["format"] = current_format
current_format = None
if current_size != None:
for ds in current_dataset["distribution"]:
ds["byte_size"] = current_size
current_size = None
current_dataset["metadata"] = []
current_dataset["technical_resource"] = []
datasets.append(current_dataset)
self.dmp["dmp"]["dataset"]=datasets
return datasets
def parse_question_1_1_1(self, text):
ret_val = []
keywords = ["Endevor", "AccuRev SCM", "ClearCase", "Dimensions CM", "IC Manage", "PTC Integrity", "PVCS", "Rational Team Concert", "SCM Anywhere", "StarTeam", "Subversion", "SVN", "Surround SCM", "Vault", "Perforce Helix Core", "Synergy", "Plastic SCM", "Azure DevOps", "BitKeeper", "Code Co-op", "darcs", "Fossil", "Git", "Mercurial", "Monotone", "Pijul", "GNU Bazaar", "Revision Control System", "Source Code Control System", "Team Foundation Server"]
text = text.lower()
for keyword in keywords:
keyword = keyword.lower()
if keyword in text:
for ds in self.dmp["dmp"]["dataset"]:
for dist in ds["distribution"]:
if dist["title"] == "Project":
dist["host"] = {
"title": keyword,
"supports_versioning": "yes"
}
ret_val.append({
"dataset":ds["title"],
"supports_versioning": "yes",
"title": keyword
})
return ret_val
def parse_question_2_1(self, text):
ret_val = []
text = self.__split_text_based_on_title(text)
for ds in self.dmp["dmp"]["dataset"]:
for line in text:
if line.startswith(ds["title"].lower()):
url = re.findall('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', line)
filename = re.findall('[\w\d\-.\/:]+\.\w+', line)
if len(url) >= 1:
url = url[0]
if url.endswith(".") or url.endswith(",") or url.endswith(")"):
url = url[:-1]
ds["metadata"].append({
"description":"Dataset Metadata",
"language":"en",
"metadata_id": {
"metadata_id": url,
"metadata_id_type": "HTTP-URI"
}
})
ret_val.append({
"dataset": ds["title"],
"description":"Dataset Metadata",
"language":"en",
"metadata_id": {
"metadata_id": url,
"metadata_id_type": "HTTP-URI"
}
})
if len(filename) >= 1:
new_names = []
for f in filename:
if not f.startswith("http"):
new_names.append(f)
filename = new_names[0]
ds["metadata"].append({
"description":"Dataset Metadata",
"language":"en",
"metadata_id": {
"metadata_id": filename,
"metadata_id_type": "custom"
}
})
ret_val.append({
"dataset": ds["title"],
"description":"Dataset Metadata",
"language":"en",
"metadata_id": {
"metadata_id": filename,
"metadata_id_type": "custom"
}
})
return ret_val
def parse_question_2_2(self, text):
ret_val = []
urls = re.findall('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', text)
if len(urls) >= 1:
url = urls[0]
if url.endswith(".") or url.endswith(",") or url.endswith(")"):
url = url[:-1]
for ds in self.dmp["dmp"]["dataset"]:
for dist in ds["distribution"]:
if dist["title"] == "Project":
dist["access_url"] = url
tld = re.findall('[^.]*\.[^.]{2,3}(?:\.[^.]{2,3})?$', url)
tld = urlparse(url).netloc
dist["host"]["title"] = tld
ret_val.append({"access_url": url, "host":{"title":tld}})
return ret_val
def parse_question_2_3(self, text):
ret_val = {}
for ds in self.dmp["dmp"]["dataset"]:
ds["data_quality_assurance"] = text
ret_val[ds["title"]] = text
return ret_val
def parse_question_3_1(self, text):
ret_val = {}
rep = {}
for ds in self.dmp["dmp"]["dataset"]:
# check if "input"
bIsInput = False
for dist in ds["distribution"]:
if dist["title"] == "Origin":
bIsInput = True
break
if bIsInput:
title = ds["title"].lower()
rep[title]=";"+title
rep = dict((re.escape(k), v) for k, v in rep.items())
pattern = re.compile("|".join(rep.keys()))
text = pattern.sub(lambda m: rep[re.escape(m.group(0))], text.lower()).split(";")
for ds in self.dmp["dmp"]["dataset"]:
ret_val[ds["title"]] = []
for dist in ds["distribution"]:
if dist["title"] == "Origin":
for line in text:
if line.startswith(ds["title"].lower()):
url = re.findall('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', line)
if len(url) >= 1:
url = url[0]
if url.endswith(".") or url.endswith(",") or url.endswith(")"):
url = url[:-1]
ds["dataset_id"] = {
"dataset_id": url,
"dataset_id_type": "HTTP-URI"
}
dist["access_url"]=url
ret_val[ds["title"]].append({
"dataset_id": url,
"dataset_id_type": "HTTP-URI",
"access_url": url
})
else:
ds["dataset_id"] = {
"dataset_id": ds["title"],
"dataset_id_type": "custom"
}
ret_val[ds["title"]].append({
"dataset_id": ds["title"],
"dataset_id_type": "custom"
})
break
else:
if "dataset_id" not in ds:
ds["dataset_id"] = {
"dataset_id": ds["title"],
"dataset_id_type": "custom"
}
ret_val[ds["title"]].append({
"dataset_id": ds["title"],
"dataset_id_type": "custom"
})
return ret_val
def parse_question_3_2(self, text):
ret_val = []
for ds in self.dmp["dmp"]["dataset"]:
ds["security_and_privacy"] = [{
"title": "Data Security",
"text": text
}]
ret_val.append({
"dataset": ds["title"],
"title": "Data Security",
"text": text
})
return ret_val
def parse_question_4_1(self, text):
ret_val = []
for ds in self.dmp["dmp"]["dataset"]:
ds["security_and_privacy"] = [{
"title": "Data Security",
"text": text
}]
ret_val.append({
"dataset": ds["title"],
"title": "Data Security",
"text": text
})
return ret_val
def parse_question_4_2(self, text):
ret_val = {}
self.dmp["dmp"]["ethical_issues_description"] = text
ret_val["ethical_issues_description"] = text
self.dmp["dmp"]["ethical_issues_exist"] = "no"
ret_val["ethical_issues_exist"] = "no"
url = re.findall('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', text)
if len(url)>=1:
url = url[0]
if url.endswith(".") or url.endswith(",") or url.endswith(")"):
url = url[:-1]
self.dmp["dmp"]["ethical_issues_report"] = url
ret_val["ethical_issues_report"] = url
self.dmp["dmp"]["ethical_issues_exist"] = "yes"
ret_val["ethical_issues_exist"] = "yes"
return ret_val
def generate(self):
return json.dumps(self.dmp, indent=2, sort_keys=False)
# -
# ## Test FWFParser
# +
dmp_title = "Parser Test"
dmp_description = "DMP Abstract is located here"
dmp_created = "2017-01-01"
dmp_modified = "2018-01-01"
dmp_id = "http://validorcid.com/12345"
pi_name = "<NAME>"
pi_mail = "<EMAIL>"
pi_orcid = "https://orcid.org/0000-0001-5305-9063"
fwf_parser = FWFParser(dmp_title, dmp_description, dmp_created, dmp_modified, dmp_id, pi_name, pi_mail, pi_orcid)
# -
# ### Test Question 1.1
# +
q1_1_test_1 = """Title: Ehescheidungen (Statistik Austria)
Description: Divorce statistic of austria
Type: Dataset
Format: csv
Source: Input
Size: 1KB
Title: Divorces by duration of marriage (Eurostat)
Description: Divorce statistic of the EU
Type: Dataset
Format: tsv
Source: Input
Size: 297KB
Title: scatter
Description: Scatterplot
Type: Image
Format: png
Source: Produced
Size: 15KB
Title: time_change
Description: Time Change Plot
Type: Image
Format: png
Source: Produced
Size: 30KB
Title: time_corr
Description: Time Correlation Plot
Type: Image
Format: png
Source: Produced
Size: 15KB
The original release was versioned as v1.0.0. Every further release however will follow a date based versioning pattern <year>.<month>.<day>.<sequence_within_day> (e.g. 2019.04.19.01).
Since the project is hosted as a git repository, keeping track of changes to the data is handled automatically.
"""
fwf_parser.parse_question_1_1(q1_1_test_1)
fwf_parser.parse_question_1_1_1(q1_1_test_1)
# -
# ### Test Question 2.1
# +
q2_1_test_1 = "The metadata description for Ehescheidungen (Statistik Austria) can be found in the jupyter notebook file divorce_analysis.ipynb or can be accessed online at https://www.data.gv.at/katalog/dataset/2d8ad82c-4730-3354-9971-9406f2ccf72c. The metadata description for Divorces by duration of marriage (Eurostat) is stored in the same file \"divorce_analysis.ipynb\" or online at http://data.europa.eu/euodp/en/data/dataset/bRJAS74ZDdIpeU7mnKhMiA."
fwf_parser.parse_question_2_1(q2_1_test_1)
# -
# ### Test Question 2.2
# +
q2_2_test_1 = "The data can be accessed through Github (https://github.com/martinpichler/data_stewardship_ex1). Documentation is available there."
fwf_parser.parse_question_2_2(q2_2_test_1)
# -
# ### Test Question 2.3
# +
q2_3_test_1 = "The project only uses already available data and no new data will be generated in the future. The used data was checked for completeness and correctness."
fwf_parser.parse_question_2_3(q2_3_test_1)
# -
# ### Test Question 3.1
# +
q3_1_test_1 = """Ehescheidungen (Statistik Austria) was downloaded from https://www.data.gv.at/katalog/dataset/2d8ad82c-4730-3354-9971-9406f2ccf72c.
Divorces by duration of marriage (Eurostat) was downloaded from https://ec.europa.eu/eurostat/web/products-datasets/-/demo_ndivdur.
scatter was produced during research.
time_change was produced during research.
time_corr was produced during research."""
fwf_parser.parse_question_3_1(q3_1_test_1)
# -
# ### Test Question 3.2
# +
q3_2_test_1 = "Data is stored in a git repository on Github and can only be modified by the owner or an administrator. If data is lost locally, the original data can be downloaded from the Github repository. No sensitive data is stored and thus the repository does not need to be addressed."
fwf_parser.parse_question_3_2(q3_2_test_1)
# -
# ### Test Question 4.1
# +
q4_1_test_1 = "No legal aspects to mention"
fwf_parser.parse_question_4_1(q4_1_test_1)
# -
# ### Test Question 4.2
# +
q4_2_test_1 = "No ethical issues exist"
fwf_parser.parse_question_4_2(q4_2_test_1)
# -
# ### Test Generate
print(fwf_parser.generate())
| test/fwf_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:zengyuyuan]
# language: python
# name: conda-env-zengyuyuan-py
# ---
# +
import os, sys
import time
import torch
import torch.nn as nn
from torch.autograd import Variable
import numpy as np
from models import ResNet as resnet_cifar
import pandas as pd
import argparse
import csv
from torch.optim.lr_scheduler import MultiStepLR
from dataLoader import DataLoader
from sklearn.metrics import confusion_matrix, classification_report # 生成混淆矩阵函数
import matplotlib.pyplot as plt # 绘图库
print('Init Finished!')
# -
args_depth = 20
num_classes = 10
args_dataset = 'cifar-10'
args_batch_size = 1
PATH = '../tb_dir/test_256bs_150epoch_layer3_MaskafterSenet'
# PATH = '../tb_dir/cifar_exp/test_256bs_200epoch_layer3_1/3epoch'
model_path = os.path.join(PATH, 'saved_model.pt')
# Data Loader
loader = DataLoader(args_dataset,batch_size=args_batch_size)
dataloaders,dataset_sizes = loader.load_data()
model = resnet_cifar(depth=args_depth, num_classes=num_classes)
model = model.cuda()
model = torch.nn.DataParallel(model)
model.load_state_dict(torch.load(model_path))
print('Successfully Load Model: ', os.path.basename(model_path))
def printF(i, total=100):
i = int( i / total * 100) + 1
total = 100
k = i + 1
str_ = '>'*i + '' ''*(total-k)
sys.stdout.write('\r'+str_+'[%s%%]'%(i+1))
sys.stdout.flush()
if(i >= total -1): print()
use_gpu = True
epoch = 1
phase = 'val'
running_corrects = 0.0
data_len = len(dataloaders[phase])
# print(data_len)
true_labels = []
model_preds = []
SEMasks = []
for idx,data in enumerate(dataloaders[phase]):
printF(idx, data_len)
inputs,labels = data
if labels[0] == 0:
if use_gpu:
inputs = Variable(inputs.cuda())
labels = Variable(labels.cuda())
else:
inputs, labels = Variable(inputs), Variable(labels)
#forward
outputs,seMask = model(inputs, labels, epoch)
_, preds = torch.max(outputs.data, 1)
SEMasks.append(seMask.cpu().detach().numpy())
y = labels.data
batch_size = labels.data.shape[0]
running_corrects += torch.sum(preds == y)
epoch_acc = float(running_corrects) /dataset_sizes[phase]
true_labels.extend(y.cpu().numpy())
model_preds.extend(preds.cpu().numpy())
print('%s top1 Acc:%.4f'%(phase,epoch_acc))
# +
print(len(SEMasks))
SEMasks = np.array(SEMasks)
avg_masks = np.mean(np.concatenate(SEMasks,axis=0),axis=0)
print(avg_masks.shape)
avg_masks = np.reshape(avg_masks,(64,))
# avg_masks = np.mean(np.reshape(avg_masks,(64,64)),axis=1)
plt.figure()
plt.bar(range(len(avg_masks)),avg_masks)
plt.show()
# -
| Old_Codes_For_Exploration/SENetAnalysis/SENet_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="clear: both; width: 100%; overflow: auto"><img src="img/yabox.png" style="width: 250px; float: left"/></div>
#
# > Yabox: Yet another black-box optimization library for Python - https://github.com/pablormier/yabox
#
# This notebook compares the performance of the Differential Evolution (DE) algorithm and the DE with parallel evaluation (PDE) implemented in [Yabox](https://github.com/pablormier/yabox) against the default [Scipy's implementation](https://docs.scipy.org/doc/scipy-0.17.0/reference/generated/scipy.optimize.differential_evolution.html) over a collection of common optimization functions.
#
# Author: <NAME>, [@pablormier](https://twitter.com/PabloRMier)
# # Imports & boilerplate code
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import sys
from time import time
# Load Yabox (from local)
sys.path.insert(0, '../')
import yabox as yb
import scipy as sp
import numpy as np
# Import the DE implementations
from yabox.algorithms import DE, PDE
from scipy.optimize import differential_evolution as SDE
# -
print('Yabox version: ', yb.__version__)
print('Scipy version: ', sp.__version__)
# # Default config
#
# The initialization of the population in all cases (Yabox/Scipy) is random and the schema used is *rand/1/bin*
# Runs per method and function, average the final results
runs = 1
# Time limit for each method (in seconds)
stop_after = 30
# Max number of iterations
maxiters = 1000000
# Use a constant mutation factor (0-1)
mutation = 0.5
# Recombination probability (0-1)
recombination = 0.5
# Number of individuals in the population. NOTE: Since Scipy uses num_individuals = dimensions * popsize
# Select a size for popsize and a set of dimensions to test so that popsize / dimensions in every case
# produces an integer number
popsize = 64
# Methods to be tested
methods = ['yabox_de', 'yabox_pde', 'scipy_de']
# # Evaluation
#
# In order to evaluate the performance of each implementation, I used 5 different multi-dimensional functions commonly used for benchmarking black-box optimization algorithms. All tests have been taken on a Intel(R) Core(TM) i7-3770 CPU @ 3.40GHz.
# ## Benchmark 1 - Ackley function
#
# > "The [Ackley function](https://www.sfu.ca/~ssurjano/ackley.html) is widely used for testing optimization algorithms. In its two-dimensional form, as shown in the plot above, it is characterized by a nearly flat outer region, and a large hole at the centre. The function poses a risk for optimization algorithms, particularly hillclimbing algorithms, to be trapped in one of its many local minima."
#
# Global minimum:
#
# $ f(\mathbf{x^*}) = 0, \text{at}~\mathbf{x^*} = (0, \dots, 0) $
from yabox.problems import Ackley
Ackley().plot3d();
# +
# Run the set of benchmarks on Ackley using the config defined at the beginning of this notebook
ackley_data = test(Ackley)
# Plot the performance of each algorithm (execution time vs. fitness)
plot_results(ackley_data)
# -
plot_results(ackley_data, use_time=False)
plot_time_per_iteration(ackley_data)
# ## Benchmark 2 - Rastrigin function
#
# > *"The [Rastrigin function](https://www.sfu.ca/~ssurjano/rastr.html) has several local minima. It is highly multimodal, but locations of the minima are regularly distributed. It is shown in the plot in its two-dimensional form."*
#
# Global minimum:
#
# $ f(\mathbf{x^*}) = 0, \text{at}~\mathbf{x^*} = (0, \dots, 0) $
from yabox.problems import Rastrigin
Rastrigin().plot3d();
rastrigin_data = test(Rastrigin)
plot_results(rastrigin_data)
plot_results(rastrigin_data, use_time=False)
plot_time_per_iteration(rastrigin_data)
# ## Benchmark 3 - Schwefel function
#
# > "The [Schwefel function](https://www.sfu.ca/~ssurjano/schwef.html) is complex, with many local minima. The plot shows the two-dimensional form of the function."
#
# Global minimum:
#
# $ f(\mathbf{x^*}) = 0, \text{at}~\mathbf{x^*} = (420.9687, \dots, 420.9687) $
from yabox.problems import Schwefel
Schwefel().plot3d();
schwefel_data = test(Schwefel)
plot_results(schwefel_data)
plot_results(schwefel_data, use_time=False)
plot_time_per_iteration(schwefel_data)
# ## Benchmark 4 - Michalewicz function
#
# > "The [Michalewicz function](https://www.sfu.ca/~ssurjano/michal.html) has d! local minima, and it is multimodal. The parameter m defines the steepness of they valleys and ridges; a larger m leads to a more difficult search. The recommended value of m is m = 10."
from yabox.problems import Michalewicz
Michalewicz().plot3d();
michalewicz_data = test(Michalewicz)
plot_results(michalewicz_data)
# ## Benchmark 5 - Griewank function
#
# > "The [Griewank function](https://www.sfu.ca/~ssurjano/griewank.html) has many widespread local minima, which are regularly distributed."
#
# Global minimum:
#
# $ f(\mathbf{x^*}) = 0, \text{at}~\mathbf{x^*} = (0, \dots, 0) $
from yabox.problems import Griewank
Griewank().plot3d();
griewank_data = test(Griewank)
plot_results(griewank_data)
| notebooks/yabox-vs-scipy-de.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $\underline{\text{Modèle de Verhulst, partie 2 : apparition des $2^n$-cycles}}$
# Bienvenue dans cette deuxième partie du modèle de Verhulst ! Je te conseille de d'abord commencer par la vidéo ou le fichier PDF associés à cette partie, avant de te lancer dans les joies de la simulation !
#
# Cet outil va te permettre d'exécuter du code Python facilement, à travers ce que j'ai déjà codé, ou de petits exercices où tu dois toi-même coder !
#
# L'objectif est que tu découvres la puissance de l'informatique pour ce genre de calculs et pour l'intuition qu'elle peut t'apporter, mais aussi ses limites. Bonne découverte !
#
# PS : Au début, appuie sur Cell->Run All pour tout lancer. Pour activer une cellule de code particulière (comme après une modification), clique dessus et appuie sur Cells->Run Cells, ou Shift+Enter ;)
# +
import os
import sys
import random as rd
import math
import numpy as np
import skimage.io
import matplotlib
import matplotlib.pyplot as plt
from bokeh.io import show, output_notebook
from bokeh.plotting import figure
from bokeh.layouts import column, row
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
from bokeh.plotting import ColumnDataSource, output_file, show
output_notebook(hide_banner=True)
# -
# $\underline{\text{Premiers essais sur la suite}}$
# Je te propose de commencer par visualiser le comportement de la suite, lorsque $\mu$
# +
#fonction f_µ
def f_mu(mu):
return lambda x:mu*x*(1-x)
def calcul_suivant(mu,x, fig):
f=f_mu(mu)
fig.x(f(x),0,color="green")
fig.line([x,x,f(x)],[x,f(x),f(x)])
fig.x(f(x),0,color="green")
return f(x)
def affichage_evolution(mu,x0,nb_etapes):
print("On note :")
print("- x0 la valeur initiale de la suite")
print("- nb_etapes le nombre d'itérations de la suite")
fig = figure(width=800, height=400, title="Evolution de la suite")
liste=np.array(np.linspace(0,1,num=40000))
fig.line(liste, f_mu(mu)(liste), color='blue')
fig.line([0,1],[0,1], color="black")
x=x0
fig.x(x,0,color="green")
for i in range(nb_etapes):
x=calcul_suivant(mu, x, fig)
show(fig)
# -
interact(affichage_evolution, mu=widgets.FloatSlider(min=0, max=3.4, step=0.05, value=2, continuous_update=False), x0=widgets.FloatSlider(min=0, max=1, step=0.05, value=0.1, continuous_update=False), nb_etapes=widgets.IntSlider(min=0, max=200, step=5, value=40, continuous_update=False))
# Tu peux ainsi découvrir par toi-même le comportement de la suite, lorsque l'on fait varier $\mu$, mais aussi la valeur initiale, ou le nombre d'itérations !
#
# Et une fois que tu penses avoir bien compris ce qui s'est passé lorsque l'on passé cette fameuse limite de 3, n'hésite pas à passer à la deuxième vidéo, pour la suite de ce module !
| Math/MOOC_Verhulst_partie_2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="xDcXTjyAJ7pq"
import pandas as pd
import requests
import io
# + colab={"base_uri": "https://localhost:8080/"} id="F6-TSwwwKwdp" outputId="85a4b90f-026c-43eb-98ec-f17cfc228be4"
# Downloading the csv file from our GitHub repo
url = "https://raw.githubusercontent.com/Jingxue-24/QM2-team9/main/Spotify%20API/merged.csv"
download = requests.get(url).content
# Reading the downloaded content and turning it into a pandas dataframe
df = pd.read_csv(io.StringIO(download.decode('utf-8')))
# Previewing data
print (df.head())
print(df.columns)
# + id="YgAEwJNOLRu9"
# Making decades and adding it to dataframe
year = df['Popular_date']
decade = year - (year%10)
df['Decade'] = decade
df.loc[ : , 'Decade'] = decade
# + id="rGOgwzX0K9Ti"
import statsmodels.api as sm
from statsmodels.formula.api import ols
# + colab={"base_uri": "https://localhost:8080/"} id="GtzmRj6mLESd" outputId="ab6e7ea0-464d-4dac-f2e9-8c0ae7e803cc"
# Tempo changing over decades
tempo = df['Tempo']
lm = ols('decade ~ tempo',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #not significant
# + colab={"base_uri": "https://localhost:8080/"} id="fNszTkuVSLs_" outputId="bf018628-b43a-434f-993c-179550ca997b"
# Energy changing over decades
energy = df['Energy']
lm = ols('decade ~ energy',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + colab={"base_uri": "https://localhost:8080/"} id="R2g9ECR9TG6g" outputId="9b4e5c95-9958-4ff3-fbf9-34a3ccd80076"
# Length of songs changing over decades
length = df['Length']
lm = ols('decade ~ length',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + colab={"base_uri": "https://localhost:8080/"} id="yzKw1gTVThUJ" outputId="a49730af-93b5-4b63-fa84-ffdf5b959434"
# Danceability changing over decades
danceability = df['Danceability']
lm = ols('decade ~ danceability',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + colab={"base_uri": "https://localhost:8080/"} id="RD3eWo-BUYoS" outputId="2a4c1bb2-45c0-4f54-9177-e1c1d9cdcbe3"
# Instrumentalness changing over decades
instrumentalness = df['Instrumentalness']
lm = ols('decade ~ instrumentalness',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + colab={"base_uri": "https://localhost:8080/"} id="it5aUfE0VyBU" outputId="6dc8a2b3-b657-4dc0-88e0-9b354e354b53"
# Loudness changing over decades
loudness = df['Loudness']
lm = ols('decade ~ loudness',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + colab={"base_uri": "https://localhost:8080/"} id="XKc6trSlGUUb" outputId="c4e83c09-fec1-4970-f6b6-e937cb8c94f8"
# Speechiness changing over decades
speechiness = df['Speechiness']
lm = ols('decade ~ speechiness',data=df).fit()
table = sm.stats.anova_lm(lm)
print(table) #is significant
# + [markdown] id="OsBOHk8gAVZF"
# Tutorial(s) followed:
#
#
# * https://towardsdatascience.com/1-way-anova-from-scratch-dissecting-the-anova-table-with-a-worked-example-170f4f2e58ad
#
#
| ANOVA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('Basics').getOrCreate()
# +
from pyspark.ml.feature import PCA
from pyspark.ml.linalg import Vectors
data = [(Vectors.sparse(5, [(1, 1.0), (3, 7.0)]),),
(Vectors.dense([2.0, 0.0, 3.0, 4.0, 5.0]),),
(Vectors.dense([4.0, 0.0, 0.0, 6.0, 7.0]),)]
df = spark.createDataFrame(data, ["features"])
pca = PCA(k=3, inputCol="features", outputCol="pcaFeatures")
model = pca.fit(df)
result = model.transform(df)
result.show(truncate=False)
# -
| 4-Data _pre_processing/PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# URL: http://bokeh.pydata.org/en/latest/docs/gallery/texas.html
#
# Most examples work across multiple plotting backends, this example is also available for:
#
# * [Bokeh - texas choropleth example](../bokeh/texas_choropleth_example.ipynb)
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# %output fig='svg'
# # Declaring data
# +
from bokeh.sampledata.us_counties import data as counties
from bokeh.sampledata.unemployment import data as unemployment
counties = [dict(county, Unemployment=unemployment[cid])
for cid, county in counties.items()
if county["state"] == "tx"]
choropleth = hv.Polygons(counties, ['lons', 'lats'], [('detailed name', 'County'), 'Unemployment'])
# -
# ## Plot
# +
plot_opts = dict(logz=True, xaxis=None, yaxis=None,
show_grid=False, show_frame=False, colorbar=True,
fig_size=200, color_index='Unemployment')
style = dict(edgecolor='white')
choropleth.opts(style=style, plot=plot_opts)
| examples/gallery/demos/matplotlib/texas_choropleth_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import time
import json
import shutil
import h5py
import numpy as np
import matplotlib.pyplot as plt
import sklearn.metrics.cluster
import torch
from torch import nn
# filename = 'hists/hypprop_with_predictor_eeb99_hughtok1_20180921_191657.h5'
# filename = 'hists/hypprop_with_predictor_eeb100_hughtok2_20180921_193244.h5'
# filename = 'hists/hypprop_with_predictor_eeb101_hughtok3_20180921_193516.h5'
# filename = 'hists/hypprop_with_predictor_eeb102_hughtok4_20180921_193310.h5'
# filename = 'hists/hypprop_with_predictor_eeb103_hughtok1_20180921_212947.h5'
# filename = 'hists/hypprop_with_predictor_eeb105_hughtok5_20180921_230949.h5'
# filename = 'hists/hypprop_with_predictor_eeb106_hughtok6_20180921_231010.h5'
# filename = 'hists/hypprop_with_predictor_eeb107_hughtok2_20180922_013551.h5'
filename = 'hists/hypprop_with_predictor_eeb108_hughtok1_20180922_170013.h5'
def run():
shutil.copyfile(f'../{filename}', '/tmp/foo.h5')
# f = h5py.File('../hists/hypprop_with_predictor_eeb95_hughtok2_20180921_144419.h5', 'r')
f = h5py.File('/tmp/foo.h5', 'r')
print('f.keys()', f.keys())
hypotheses_train = torch.from_numpy(f['hypotheses_train'][:])
hypotheses_gnd_train = torch.from_numpy(f['gnd_hypotheses_train'][:].astype(np.uint8)).long()
dsrefs_train = torch.from_numpy(f['dsrefs_train'][:].astype(np.uint8)).long()
resdicts = f['resdicts']
print(hypotheses_train.shape)
print(hypotheses_gnd_train.shape)
print(dsrefs_train.shape)
N = dsrefs_train.size(0)
print('N', N)
num_renders = len(resdicts)
num_samples = 1
for i in range(num_samples):
for j in range(5):
print(hypotheses_train[j, -i - 1])
for i in range(num_samples):
for j in range(5):
print(torch.softmax(hypotheses_train[j, -i - 1], dim=-1))
run()
| utils/print_hyps.ipynb |
# !bash ../setup.sh
# # Model Training
#
# In this notebook, we'll train a LightGBM model using Amazon SageMaker, so
# we have an example trained model to explain.
#
# You can bring also bring your own trained models to explain. See the
# customizing section for more details.
#
# <p align="center">
# <img src="https://github.com/awslabs/sagemaker-explaining-credit-decisions/raw/master/docs/architecture_diagrams/stage_2.png" width="1000px">
# </p>
# We start by importing a variety of packages that will be used throughout
# the notebook. One of the most important packages used throughout this
# solution is the Amazon SageMaker Python SDK (i.e. `import sagemaker`). We
# also import modules from our own custom package that can be found at
# `./package`.
# +
from pathlib import Path
from sagemaker.sklearn import SKLearn
from sagemaker.local import LocalSession
from package import config, utils
from package.sagemaker import containers
# -
# ## Container
# We now build our custom Docker image that will be used for model training
# and deployment. It extends the official Amazon SageMaker framework image
# for Scikit-learn, by adding additional packages such as
# [LightGBM](https://lightgbm.readthedocs.io/en/latest/) and
# [SHAP](https://github.com/slundberg/shap). After building the image, we
# upload it to our solution's Amazon ECR repository.
# +
# %env AWS_SDK_LOAD_CONFIG=true
scikit_learn_image = containers.scikit_learn_image()
custom_image = containers.custom_image()
current_folder = utils.get_current_folder(globals())
dockerfile = Path(current_folder, '../containers/model/Dockerfile')
custom_image.build(
dockerfile=dockerfile,
buildargs={'SCIKIT_LEARN_IMAGE': str(scikit_learn_image)}
)
custom_image.push()
# -
# ## Model Training
# Amazon SageMaker provides two methods for training and deploying models.
# You can start by quickly testing and debuging models on the Amazon
# SageMaker Notebook instance using local mode. After this, you can scale
# up training with SageMaker mode on dedicated instances and deploy the
# model on dedicated instance too. Since this is a pre-developed solution
# we'll be using SageMaker mode.
# Up next, we configure our SKLearn estimator. We will use it to coordinate
# model training and deployment. We reference our custom container (see
# `image_name`) and our custom code (see `entry_point` and `dependencies`).
# At this stage, we also reference the instance type (and instance count)
# that will be used during training, and the hyperparmeters we wish to use.
# And lastly we set the `output_path` for trained model artifacts and
# `code_location` for a snapshot of the training script that was used.
#
# **Note**: when customizing the solution, you can enable enhanced logging
# by setting the `container_log_level=logging.DEBUG` on the `SKLearn`
# estimator object (after `import logging`).
# +
hyperparameters = {
"tree-n-estimators": 42,
"tree-max-depth": 2,
"tree-min-child-samples": 1,
"tree-boosting-type": "dart"
}
estimator = SKLearn(
image_name=str(custom_image),
entry_point='entry_point.py',
source_dir=str(Path(current_folder, '../containers/model/src').resolve()),
dependencies=[str(Path(current_folder, '../package/package').resolve())],
hyperparameters=hyperparameters,
role=config.SAGEMAKER_IAM_ROLE,
train_instance_count=1,
train_instance_type='ml.c5.xlarge',
output_path='s3://' + str(Path(config.S3_BUCKET, config.OUTPUTS_S3_PREFIX)),
code_location='s3://' + str(Path(config.S3_BUCKET, config.OUTPUTS_S3_PREFIX)),
base_job_name=config.RESOURCE_NAME,
tags=[{'Key': config.TAG_KEY, 'Value': config.RESOURCE_NAME}]
)
# -
# With our estimator now initialized, we can start the Amazon SageMaker
# training job. Since our entry point script expects a number of data
# channels to be defined, we can provide them when calling `fit`. When
# referencing `s3://` folders, the contents of these folders will be
# automatically downloaded from Amazon S3 before the entry point script is
# run. When using local mode, it's possible to avoid this data transfer and
# reference local folder using the `file://` prefix instead: e.g.
# `{'schemas': 'file://' + str(schema_folder)}`
#
# You can expect this step to take approximately 5 minutes.
estimator.fit({
'schemas': 's3://' + str(Path(config.S3_BUCKET, config.SCHEMAS_S3_PREFIX)),
'data_train': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'data_train')),
'label_train': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'label_train')),
'data_test': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'data_test')),
'label_test': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'label_test'))
})
# Our Amazon SageMaker training job has now completed, and we should have a
# number of trained model artifacts that can be deployed and used for
# explanations.
# ## Customization
#
# We have provided an example of model training above, so that we have a
# trained model to explain, but our solution is customizable if you have
# your own models. You can choose to retrain your models on Amazon
# SageMaker or train your models is some other way of your choosing.
#
# When re-training models on Amazon SageMaker you should modify the
# training script found at `./package/sagemaker/estimator_fns.py`. You
# should modify the `train_fn` function as required and change any of the
# other training functions (found in `./package/machine_learning` for
# example). You may need to modify the dependencies too depending on you
# model and these can be adjusted in `./containers/model/requirements.txt`.
#
# When bringing your own trained model, you will need to upload all of the
# model assets to Amazon S3 (in the solution bucket): e.g. trained
# preprocessors, model weights and feature schemas (i.e. data schema after
# feature engineering). Amazon SageMaker expects all of these model assets
# to be packages up as a `model.tar.gz`.
# ## Next Stage
#
# Up next we'll deploy the model explainer to a HTTP endpoint using Amazon
# SageMaker and visualize the explanations.
#
# [Click here to continue.](./3_endpoint.ipynb)
| sagemaker/notebooks/2_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# **Chapter 3 – Classification**
#
# _This notebook contains all the sample code and solutions to the exercices in chapter 3._
# + [markdown] deletable=true editable=true
# # Setup
# + [markdown] deletable=true editable=true
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# + deletable=true editable=true
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import numpy.random as rnd
import os
# to make this notebook's output stable across runs
rnd.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "classification"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# + [markdown] deletable=true editable=true
# # MNIST
# + deletable=true editable=true
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original')
mnist
# + deletable=true editable=true
X, y = mnist["data"], mnist["target"]
X.shape
# + deletable=true editable=true
y.shape
# + deletable=true editable=true
28*28
# + deletable=true editable=true
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
some_digit = X[36000]
some_digit_image = some_digit.reshape(28, 28)
plt.imshow(some_digit_image, cmap = matplotlib.cm.binary,
interpolation="nearest")
plt.axis("off")
save_fig("some_digit_plot")
plt.show()
# + deletable=true editable=true
def plot_digit(data):
image = data.reshape(28, 28)
plt.imshow(image, cmap = matplotlib.cm.binary,
interpolation="nearest")
plt.axis("off")
# + deletable=true editable=true
# EXTRA
def plot_digits(instances, images_per_row=10, **options):
size = 28
images_per_row = min(len(instances), images_per_row)
images = [instance.reshape(size,size) for instance in instances]
n_rows = (len(instances) - 1) // images_per_row + 1
row_images = []
n_empty = n_rows * images_per_row - len(instances)
images.append(np.zeros((size, size * n_empty)))
for row in range(n_rows):
rimages = images[row * images_per_row : (row + 1) * images_per_row]
row_images.append(np.concatenate(rimages, axis=1))
image = np.concatenate(row_images, axis=0)
plt.imshow(image, cmap = matplotlib.cm.binary, **options)
plt.axis("off")
# + deletable=true editable=true
plt.figure(figsize=(9,9))
example_images = np.r_[X[:12000:600], X[13000:30600:600], X[30600:60000:590]]
plot_digits(example_images, images_per_row=10)
save_fig("more_digits_plot")
plt.show()
# + deletable=true editable=true
y[36000]
# + deletable=true editable=true
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# + deletable=true editable=true
import numpy as np
shuffle_index = np.random.permutation(60000)
X_train, y_train = X_train[shuffle_index], y_train[shuffle_index]
# + [markdown] deletable=true editable=true
# # Binary classifier
# + deletable=true editable=true
y_train_5 = (y_train == 5)
y_test_5 = (y_test == 5)
# + deletable=true editable=true
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier(random_state=42)
sgd_clf.fit(X_train, y_train_5)
# + deletable=true editable=true
sgd_clf.predict([some_digit])
# + deletable=true editable=true
from sklearn.model_selection import cross_val_score
cross_val_score(sgd_clf, X_train, y_train_5, cv=3, scoring="accuracy")
# + deletable=true editable=true
from sklearn.model_selection import StratifiedKFold
from sklearn.base import clone
skfolds = StratifiedKFold(n_splits=3, random_state=42)
for train_index, test_index in skfolds.split(X_train, y_train_5):
clone_clf = clone(sgd_clf)
X_train_folds = X_train[train_index]
y_train_folds = (y_train_5[train_index])
X_test_fold = X_train[test_index]
y_test_fold = (y_train_5[test_index])
clone_clf.fit(X_train_folds, y_train_folds)
y_pred = clone_clf.predict(X_test_fold)
n_correct = sum(y_pred == y_test_fold)
print(n_correct / len(y_pred))
# + deletable=true editable=true
from sklearn.base import BaseEstimator
class Never5Classifier(BaseEstimator):
def fit(self, X, y=None):
pass
def predict(self, X):
return np.zeros((len(X), 1), dtype=bool)
# + deletable=true editable=true
never_5_clf = Never5Classifier()
cross_val_score(never_5_clf, X_train, y_train_5, cv=3, scoring="accuracy")
# + deletable=true editable=true
from sklearn.model_selection import cross_val_predict
y_train_pred = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3)
# + deletable=true editable=true
from sklearn.metrics import confusion_matrix
confusion_matrix(y_train_5, y_train_pred)
# + deletable=true editable=true
y_train_perfect_predictions = y_train_5
# + deletable=true editable=true
confusion_matrix(y_train_5, y_train_perfect_predictions)
# + deletable=true editable=true
from sklearn.metrics import precision_score, recall_score
precision_score(y_train_5, y_train_pred)
# + deletable=true editable=true
4344 / (4344 + 1307)
# + deletable=true editable=true
recall_score(y_train_5, y_train_pred)
# + deletable=true editable=true
4344 / (4344 + 1077)
# + deletable=true editable=true
from sklearn.metrics import f1_score
f1_score(y_train_5, y_train_pred)
# + deletable=true editable=true
4344 / (4344 + (1077 + 1307)/2)
# + deletable=true editable=true
y_scores = sgd_clf.decision_function([some_digit])
y_scores
# + deletable=true editable=true
threshold = 0
y_some_digit_pred = (y_scores > threshold)
# + deletable=true editable=true
y_some_digit_pred
# + deletable=true editable=true
threshold = 200000
y_some_digit_pred = (y_scores > threshold)
y_some_digit_pred
# + deletable=true editable=true
y_scores = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3,
method="decision_function")
# + deletable=true editable=true
from sklearn.metrics import precision_recall_curve
precisions, recalls, thresholds = precision_recall_curve(y_train_5, y_scores)
# + deletable=true editable=true
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision", linewidth=2)
plt.plot(thresholds, recalls[:-1], "g-", label="Recall", linewidth=2)
plt.xlabel("Threshold", fontsize=16)
plt.legend(loc="upper left", fontsize=16)
plt.ylim([0, 1])
plt.figure(figsize=(8, 4))
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.xlim([-700000, 700000])
save_fig("precision_recall_vs_threshold_plot")
plt.show()
# + deletable=true editable=true
(y_train_pred == (y_scores > 0)).all()
# + deletable=true editable=true
y_train_pred_90 = (y_scores > 70000)
# + deletable=true editable=true
precision_score(y_train_5, y_train_pred_90)
# + deletable=true editable=true
recall_score(y_train_5, y_train_pred_90)
# + deletable=true editable=true
def plot_precision_vs_recall(precisions, recalls):
plt.plot(recalls, precisions, "b-", linewidth=2)
plt.xlabel("Recall", fontsize=16)
plt.ylabel("Precision", fontsize=16)
plt.axis([0, 1, 0, 1])
plt.figure(figsize=(8, 6))
plot_precision_vs_recall(precisions, recalls)
save_fig("precision_vs_recall_plot")
plt.show()
# + [markdown] deletable=true editable=true
# # ROC curves
# + deletable=true editable=true
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_train_5, y_scores)
# + deletable=true editable=true
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([0, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.figure(figsize=(8, 6))
plot_roc_curve(fpr, tpr)
save_fig("roc_curve_plot")
plt.show()
# + deletable=true editable=true
from sklearn.metrics import roc_auc_score
roc_auc_score(y_train_5, y_scores)
# -
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(random_state=42)
y_probas_forest = cross_val_predict(forest_clf, X_train, y_train_5, cv=3,
method="predict_proba")
y_scores_forest = y_probas_forest[:, 1] # score = proba of positive class
fpr_forest, tpr_forest, thresholds_forest = roc_curve(y_train_5,y_scores_forest)
# + deletable=true editable=true
plt.figure(figsize=(8, 6))
plt.plot(fpr, tpr, "b:", linewidth=2, label="SGD")
plot_roc_curve(fpr_forest, tpr_forest, "Random Forest")
plt.legend(loc="lower right", fontsize=16)
save_fig("roc_curve_comparison_plot")
plt.show()
# + deletable=true editable=true
roc_auc_score(y_train_5, y_scores_forest)
# + deletable=true editable=true
y_train_pred_forest = cross_val_predict(forest_clf, X_train, y_train_5, cv=3)
precision_score(y_train_5, y_train_pred_forest)
# + deletable=true editable=true
recall_score(y_train_5, y_train_pred_forest)
# + [markdown] deletable=true editable=true
# # Multiclass classification
# + deletable=true editable=true
sgd_clf.fit(X_train, y_train)
sgd_clf.predict([some_digit])
# + deletable=true editable=true
some_digit_scores = sgd_clf.decision_function([some_digit])
some_digit_scores
# + deletable=true editable=true
np.argmax(some_digit_scores)
# + deletable=true editable=true
sgd_clf.classes_
# -
sgd_clf.classes_[5]
# + deletable=true editable=true
from sklearn.multiclass import OneVsOneClassifier
ovo_clf = OneVsOneClassifier(SGDClassifier(random_state=42))
ovo_clf.fit(X_train, y_train)
ovo_clf.predict([some_digit])
# + deletable=true editable=true
len(ovo_clf.estimators_)
# + deletable=true editable=true
forest_clf.fit(X_train, y_train)
forest_clf.predict([some_digit])
# + deletable=true editable=true
forest_clf.predict_proba([some_digit])
# + deletable=true editable=true
cross_val_score(sgd_clf, X_train, y_train, cv=3, scoring="accuracy")
# + deletable=true editable=true
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float64))
cross_val_score(sgd_clf, X_train_scaled, y_train, cv=3, scoring="accuracy")
# + deletable=true editable=true
y_train_pred = cross_val_predict(sgd_clf, X_train_scaled, y_train, cv=3)
conf_mx = confusion_matrix(y_train, y_train_pred)
conf_mx
# + deletable=true editable=true
def plot_confusion_matrix(matrix):
"""If you prefer color and a colorbar"""
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
cax = ax.matshow(conf_mx)
fig.colorbar(cax)
# -
plt.matshow(conf_mx, cmap=plt.cm.gray)
save_fig("confusion_matrix_plot", tight_layout=False)
plt.show()
# + deletable=true editable=true
row_sums = conf_mx.sum(axis=1, keepdims=True)
norm_conf_mx = conf_mx / row_sums
# -
np.fill_diagonal(norm_conf_mx, 0)
plt.matshow(norm_conf_mx, cmap=plt.cm.gray)
save_fig("confusion_matrix_errors_plot", tight_layout=False)
plt.show()
# + deletable=true editable=true
cl_a, cl_b = 3, 5
X_aa = X_train[(y_train == cl_a) & (y_train_pred == cl_a)]
X_ab = X_train[(y_train == cl_a) & (y_train_pred == cl_b)]
X_ba = X_train[(y_train == cl_b) & (y_train_pred == cl_a)]
X_bb = X_train[(y_train == cl_b) & (y_train_pred == cl_b)]
plt.figure(figsize=(8,8))
plt.subplot(221); plot_digits(X_aa[:25], images_per_row=5)
plt.subplot(222); plot_digits(X_ab[:25], images_per_row=5)
plt.subplot(223); plot_digits(X_ba[:25], images_per_row=5)
plt.subplot(224); plot_digits(X_bb[:25], images_per_row=5)
save_fig("error_analysis_digits_plot")
plt.show()
# + [markdown] deletable=true editable=true
# # Multilabel classification
# + deletable=true editable=true
from sklearn.neighbors import KNeighborsClassifier
y_train_large = (y_train >= 7)
y_train_odd = (y_train % 2 == 1)
y_multilabel = np.c_[y_train_large, y_train_odd]
knn_clf = KNeighborsClassifier()
knn_clf.fit(X_train, y_multilabel)
# + deletable=true editable=true
knn_clf.predict([some_digit])
# + deletable=true editable=true
y_train_knn_pred = cross_val_predict(knn_clf, X_train, y_train, cv=3)
f1_score(y_train, y_train_knn_pred, average="macro")
# + [markdown] deletable=true editable=true
# # Multioutput classification
# + deletable=true editable=true
noise = rnd.randint(0, 100, (len(X_train), 784))
X_train_mod = X_train + noise
noise = rnd.randint(0, 100, (len(X_test), 784))
X_test_mod = X_test + noise
y_train_mod = X_train
y_test_mod = X_test
# + deletable=true editable=true
some_index = 5500
plt.subplot(121); plot_digit(X_test_mod[some_index])
plt.subplot(122); plot_digit(y_test_mod[some_index])
save_fig("noisy_digit_example_plot")
plt.show()
# + deletable=true editable=true
knn_clf.fit(X_train_mod, y_train_mod)
clean_digit = knn_clf.predict([X_test_mod[some_index]])
plot_digit(clean_digit)
save_fig("cleaned_digit_example_plot")
# + [markdown] deletable=true editable=true
# # Extra material
# + [markdown] deletable=true editable=true
# ## Dummy (ie. random) classifier
# + deletable=true editable=true
from sklearn.dummy import DummyClassifier
dmy_clf = DummyClassifier()
y_probas_dmy = cross_val_predict(dmy_clf, X_train, y_train_5, cv=3, method="predict_proba")
y_scores_dmy = y_probas_dmy[:, 1]
# + deletable=true editable=true
fprr, tprr, thresholdsr = roc_curve(y_train_5, y_scores_dmy)
plot_roc_curve(fprr, tprr)
# + [markdown] deletable=true editable=true
# ## KNN classifier
# + deletable=true editable=true
from sklearn.neighbors import KNeighborsClassifier
knn_clf = KNeighborsClassifier(n_jobs=-1, weights='distance', n_neighbors=4)
knn_clf.fit(X_train, y_train)
# + deletable=true editable=true
y_knn_pred = knn_clf.predict(X_test)
# + deletable=true editable=true
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_knn_pred)
# + deletable=true editable=true
from scipy.ndimage.interpolation import shift
def shift_digit(digit_array, dx, dy, new=0):
return shift(digit_array.reshape(28, 28), [dy, dx], cval=new).reshape(784)
plot_digit(shift_digit(some_digit, 5, 1, new=100))
# + deletable=true editable=true
X_train_expanded = [X_train]
y_train_expanded = [y_train]
for dx, dy in ((1, 0), (-1, 0), (0, 1), (0, -1)):
shifted_images = np.apply_along_axis(shift_digit, axis=1, arr=X_train, dx=dx, dy=dy)
X_train_expanded.append(shifted_images)
y_train_expanded.append(y_train)
X_train_expanded = np.concatenate(X_train_expanded)
y_train_expanded = np.concatenate(y_train_expanded)
X_train_expanded.shape, y_train_expanded.shape
# + deletable=true editable=true
knn_clf.fit(X_train_expanded, y_train_expanded)
# + deletable=true editable=true
y_knn_expanded_pred = knn_clf.predict(X_test)
# + deletable=true editable=true
accuracy_score(y_test, y_knn_expanded_pred)
# + deletable=true editable=true
ambiguous_digit = X_test[2589]
knn_clf.predict_proba([ambiguous_digit])
# + deletable=true editable=true
plot_digit(ambiguous_digit)
# + [markdown] deletable=true editable=true
# # Exercise solutions
# + [markdown] deletable=true editable=true
# **Coming soon**
# + deletable=true editable=true
| 03_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# * This notebook contains the complete code to merge the all data sets used in this application.
#
#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## 1. Plotting the data of sentiment analysis and the stock market prices
df_text = pd.read_pickle("data sets/microsoft_processed_text_with_time_and_sentiments.pkl")
#df_text.head()
plt.style.use('seaborn-dark')
df_text[["sentimental_analysis_split_by_dot_average","sentimental_analysis_complete_text"]].plot(cmap = "viridis",
linestyle='-',figsize = (15,10))
plt.grid()
plt.show()
# +
df_values = pd.read_pickle("data sets/df_dow_jones.pkl")
#df_values.head()
microsoft_df = df_values[["MSFT"]]
microsoft_df.plot(cmap = "viridis",linestyle='-',figsize = (15,10),marker='.')
plt.grid()
plt.show()
# -
# ## 2. Routine to merge the data sets
# +
X = df_text["sentimental_analysis_split_by_dot_average"].copy()
X = X.reset_index(drop= False)
X["time"] = pd.to_datetime(X["time"],errors = 'coerce', format = '%Y-%m-%dT%H:%M',infer_datetime_format = True, cache = True,utc=True)
X["time"] = pd.to_datetime(X["time"])
X = X.set_index(pd.DatetimeIndex(X["time"]))
X.set_index("time")
X.shape
# +
df_text_gensim = pd.read_pickle("data sets/microsoft_processed_text_with_time_and_gensim.pkl")
#df_text_gensim.head()
names = df_text_gensim.columns
df_gensim = df_text_gensim[names[5:]].copy()
df_gensim = df_gensim.set_index(X.index)
X = pd.concat([X,df_gensim],axis= 1)
# -
X = X.drop(["time"],axis= 1)
X = X.resample('1d').first()
X = X.tz_convert(None)
y = microsoft_df.copy()
y.shape
result = pd.concat([X,y], axis=1,join='inner')
result.shape
# +
result = result.fillna(method='ffill', inplace=False)
# -
result.shape
result.head()
result[["sentimental_analysis_split_by_dot_average","amsterdam"]].plot(cmap = "viridis",linestyle='-',figsize = (15,10))
plt.grid()
plt.show()
result.to_pickle("data sets/data_to_paper_microsoft_case.pkl")
| 04 data wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sage
# name: sagemath
# ---
q = 16798108731015832284940804142231733909889187121439069848933715426072753864723
E = EllipticCurve(GF(q), [0,0,0,0,2])
P = E(-1,1); P
P.order()
E.order()
| src/curve_bn254/sage_scripts/G1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# +
import keras
import keras.backend as K
from keras.datasets import mnist
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Conv2DTranspose, Embedding, Multiply, Activation
from functools import partial
from collections import defaultdict
import os
import pickle
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import isolearn.io as isoio
import isolearn.keras as isol
import matplotlib.pyplot as plt
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
import pandas as pd
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class MySequence :
def __init__(self) :
self.dummy = 1
keras.utils.Sequence = MySequence
import isolearn.keras as iso
from sequence_logo_helper_protein import plot_protein_logo, letterAt_protein
class IdentityEncoder(iso.SequenceEncoder) :
def __init__(self, seq_len, channel_map) :
super(IdentityEncoder, self).__init__('identity', (seq_len, len(channel_map)))
self.seq_len = seq_len
self.n_channels = len(channel_map)
self.encode_map = channel_map
self.decode_map = {
val : key for key, val in channel_map.items()
}
def encode(self, seq) :
encoding = np.zeros((self.seq_len, self.n_channels))
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
return encoding
def encode_inplace(self, seq, encoding) :
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
def encode_inplace_sparse(self, seq, encoding_mat, row_index) :
raise NotImplementError()
def decode(self, encoding) :
seq = ''
for pos in range(0, encoding.shape[0]) :
argmax_nt = np.argmax(encoding[pos, :])
max_nt = np.max(encoding[pos, :])
if max_nt == 1 :
seq += self.decode_map[argmax_nt]
else :
seq += "0"
return seq
def decode_sparse(self, encoding_mat, row_index) :
encoding = np.array(encoding_mat[row_index, :].todense()).reshape(-1, 4)
return self.decode(encoding)
class NopTransformer(iso.ValueTransformer) :
def __init__(self, n_classes) :
super(NopTransformer, self).__init__('nop', (n_classes, ))
self.n_classes = n_classes
def transform(self, values) :
return values
def transform_inplace(self, values, transform) :
transform[:] = values
def transform_inplace_sparse(self, values, transform_mat, row_index) :
transform_mat[row_index, :] = np.ravel(values)
# +
#Re-load cached dataframe (shuffled)
dataset_name = "coiled_coil_binders"
experiment = "baker_big_set_5x_negatives"
pair_df = pd.read_csv("pair_df_" + experiment + "_in_shuffled.csv", sep="\t")
print("len(pair_df) = " + str(len(pair_df)))
print(pair_df.head())
#Generate training and test set indexes
valid_set_size = 0.0005
test_set_size = 0.0995
data_index = np.arange(len(pair_df), dtype=np.int)
train_index = data_index[:-int(len(pair_df) * (valid_set_size + test_set_size))]
valid_index = data_index[train_index.shape[0]:-int(len(pair_df) * test_set_size)]
test_index = data_index[train_index.shape[0] + valid_index.shape[0]:]
print('Training set size = ' + str(train_index.shape[0]))
print('Validation set size = ' + str(valid_index.shape[0]))
print('Test set size = ' + str(test_index.shape[0]))
# +
#Calculate sequence lengths
pair_df['amino_seq_1_len'] = pair_df['amino_seq_1'].str.len()
pair_df['amino_seq_2_len'] = pair_df['amino_seq_2'].str.len()
# +
#Initialize sequence encoder
seq_length = 81
residue_map = {'D': 0, 'E': 1, 'V': 2, 'K': 3, 'R': 4, 'L': 5, 'S': 6, 'T': 7, 'N': 8, 'H': 9, 'A': 10, 'I': 11, 'G': 12, 'P': 13, 'Q': 14, 'Y': 15, 'W': 16, 'M': 17, 'F': 18, '#': 19}
encoder = IdentityEncoder(seq_length, residue_map)
# +
#Construct data generators
class CategoricalRandomizer :
def __init__(self, case_range, case_probs) :
self.case_range = case_range
self.case_probs = case_probs
self.cases = 0
def get_random_sample(self, index=None) :
if index is None :
return self.cases
else :
return self.cases[index]
def generate_random_sample(self, batch_size=1, data_ids=None) :
self.cases = np.random.choice(self.case_range, size=batch_size, replace=True, p=self.case_probs)
def get_amino_seq(row, index, flip_randomizer, homodimer_randomizer, max_seq_len=seq_length) :
is_flip = True if flip_randomizer.get_random_sample(index=index) == 1 else False
is_homodimer = True if homodimer_randomizer.get_random_sample(index=index) == 1 else False
amino_seq_1, amino_seq_2 = row['amino_seq_1'], row['amino_seq_2']
if is_flip :
amino_seq_1, amino_seq_2 = row['amino_seq_2'], row['amino_seq_1']
if is_homodimer and row['interacts'] < 0.5 :
amino_seq_2 = amino_seq_1
return amino_seq_1, amino_seq_2
flip_randomizer = CategoricalRandomizer(np.arange(2), np.array([0.5, 0.5]))
homodimer_randomizer = CategoricalRandomizer(np.arange(2), np.array([0.95, 0.05]))
batch_size = 32
data_gens = {
gen_id : iso.DataGenerator(
idx,
{ 'df' : pair_df },
batch_size=(idx.shape[0] // batch_size) * batch_size,
inputs = [
{
'id' : 'amino_seq_1',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: (get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_2',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: (get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_1_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: len(get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[0]),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
},
{
'id' : 'amino_seq_2_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index, flip_randomizer=flip_randomizer, homodimer_randomizer=homodimer_randomizer: len(get_amino_seq(row, index, flip_randomizer, homodimer_randomizer)[1]),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
}
],
outputs = [
{
'id' : 'interacts',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: row['interacts'],
'transformer' : NopTransformer(1),
'dim' : (1,),
'sparsify' : False
}
],
randomizers = [flip_randomizer, homodimer_randomizer],
shuffle = True
) for gen_id, idx in [('train', train_index), ('valid', valid_index), ('test', test_index)]
}
# +
#Load data matrices
[x_1_train, x_2_train, l_1_train, l_2_train], [y_train] = data_gens['train'][0]
[x_1_val, x_2_val, l_1_val, l_2_val], [y_val] = data_gens['test'][0]
print("x_1_train.shape = " + str(x_1_train.shape))
print("x_2_train.shape = " + str(x_2_train.shape))
print("x_1_val.shape = " + str(x_1_val.shape))
print("x_2_val.shape = " + str(x_2_val.shape))
print("l_1_train.shape = " + str(l_1_train.shape))
print("l2_train.shape = " + str(l_2_train.shape))
print("l_1_val.shape = " + str(l_1_val.shape))
print("l2_val.shape = " + str(l_2_val.shape))
print("y_train.shape = " + str(y_train.shape))
print("y_val.shape = " + str(y_val.shape))
# +
#Define sequence templates
sequence_templates = [
'$' * i + '@' * (seq_length - i)
for i in range(seq_length+1)
]
sequence_masks = [
np.array([1 if sequence_templates[i][j] == '$' else 0 for j in range(len(sequence_templates[i]))])
for i in range(seq_length+1)
]
# +
#Load cached dataframe (shuffled)
dataset_name = "coiled_coil_binders"
experiment = "coiled_coil_binders_alyssa"
data_df = pd.read_csv(experiment + ".csv", sep="\t")
print("len(data_df) = " + str(len(data_df)))
test_df = data_df.copy().reset_index(drop=True)
batch_size = 32
test_df = test_df.iloc[:(len(test_df) // batch_size) * batch_size].copy().reset_index(drop=True)
print("len(test_df) = " + str(len(test_df)))
print(test_df.head())
# +
#Construct test data
batch_size = 32
test_gen = iso.DataGenerator(
np.arange(len(test_df), dtype=np.int),
{ 'df' : test_df },
batch_size=(len(test_df) // batch_size) * batch_size,
inputs = [
{
'id' : 'amino_seq_1',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index: (row['amino_seq_1'] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index: row['amino_seq_1'],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_2',
'source_type' : 'dataframe',
'source' : 'df',
#'extractor' : lambda row, index: row['amino_seq_2'] + "#" * seq_length)[:seq_length],
'extractor' : lambda row, index: row['amino_seq_2'],
'encoder' : IdentityEncoder(seq_length, residue_map),
'dim' : (1, seq_length, len(residue_map)),
'sparsify' : False
},
{
'id' : 'amino_seq_1_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: len(row['amino_seq_1']),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
},
{
'id' : 'amino_seq_2_len',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: len(row['amino_seq_2']),
'encoder' : lambda t: t,
'dim' : (1,),
'sparsify' : False
}
],
outputs = [
{
'id' : 'interacts',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : lambda row, index: row['interacts'],
'transformer' : NopTransformer(1),
'dim' : (1,),
'sparsify' : False
}
],
randomizers = [],
shuffle = False
)
#Load data matrices
[x_1_test, x_2_test, l_1_test, l_2_test], [y_test] = test_gen[0]
print("x_1_test.shape = " + str(x_1_test.shape))
print("x_2_test.shape = " + str(x_2_test.shape))
print("l_1_test.shape = " + str(l_1_test.shape))
print("l_2_test.shape = " + str(l_2_test.shape))
print("y_test.shape = " + str(y_test.shape))
# +
#Load predictor model
def get_shared_model() :
gru_1 = Bidirectional(CuDNNGRU(64, return_sequences=False), merge_mode='concat')
drop_1 = Dropout(0.25)
def shared_model(inp) :
gru_1_out = gru_1(inp)
drop_1_out = drop_1(gru_1_out)
return drop_1_out
return shared_model
shared_model = get_shared_model()
#Inputs
res_1 = Input(shape=(seq_length, 19 + 1))
res_2 = Input(shape=(seq_length, 19 + 1))
#Outputs
true_interacts = Input(shape=(1,))
#Interaction model definition
dense_out_1 = shared_model(res_1)
dense_out_2 = shared_model(res_2)
layer_dense_pair_1 = Dense(128, activation='relu')
dense_out_pair = layer_dense_pair_1(Concatenate(axis=-1)([dense_out_1, dense_out_2]))
pred_interacts = Dense(1, activation='sigmoid', kernel_initializer='zeros')(dense_out_pair)
predictor = Model(
inputs=[
res_1,
res_2
],
outputs=pred_interacts
)
predictor.load_weights('saved_models/ppi_rnn_baker_big_set_5x_negatives_classifier_symmetric_drop_25_5x_negatives_balanced_partitioned_data_epoch_10.h5', by_name=True)
predictor.trainable = False
predictor.compile(
optimizer=keras.optimizers.SGD(lr=0.1),
loss='mean_squared_error'
)
# +
#Generate (original) predictions
pred_train = predictor.predict([x_1_train[:, 0, ...], x_2_train[:, 0, ...]], batch_size=32, verbose=True)
pred_val = predictor.predict([x_1_val[:, 0, ...], x_2_val[:, 0, ...]], batch_size=32, verbose=True)
pred_test = predictor.predict([x_1_test[:, 0, ...], x_2_test[:, 0, ...]], batch_size=32, verbose=True)
pred_train = np.concatenate([1. - pred_train, pred_train], axis=1)
pred_val = np.concatenate([1. - pred_val, pred_val], axis=1)
pred_test = np.concatenate([1. - pred_test, pred_test], axis=1)
# +
#Make two-channel targets
y_train = np.concatenate([1. - y_train, y_train], axis=1)
y_val = np.concatenate([1. - y_val, y_val], axis=1)
y_test = np.concatenate([1. - y_test, y_test], axis=1)
# +
from keras.layers import Input, Dense, Multiply, Flatten, Reshape, Conv2D, MaxPooling2D, GlobalMaxPooling2D, Activation
from keras.layers import BatchNormalization
from keras.models import Sequential, Model
from keras.optimizers import Adam
from keras import regularizers
from keras import backend as K
import tensorflow as tf
import numpy as np
from keras.layers import Layer, InputSpec
from keras import initializers, regularizers, constraints
class InstanceNormalization(Layer):
def __init__(self, axes=(1, 2), trainable=True, **kwargs):
super(InstanceNormalization, self).__init__(**kwargs)
self.axes = axes
self.trainable = trainable
def build(self, input_shape):
self.beta = self.add_weight(name='beta',shape=(input_shape[-1],),
initializer='zeros',trainable=self.trainable)
self.gamma = self.add_weight(name='gamma',shape=(input_shape[-1],),
initializer='ones',trainable=self.trainable)
def call(self, inputs):
mean, variance = tf.nn.moments(inputs, self.axes, keep_dims=True)
return tf.nn.batch_normalization(inputs, mean, variance, self.beta, self.gamma, 1e-6)
def bernoulli_sampling (prob):
""" Sampling Bernoulli distribution by given probability.
Args:
- prob: P(Y = 1) in Bernoulli distribution.
Returns:
- samples: samples from Bernoulli distribution
"""
n, x_len, y_len, d = prob.shape
samples = np.random.binomial(1, prob, (n, x_len, y_len, d))
return samples
class INVASE():
"""INVASE class.
Attributes:
- x_train: training features
- y_train: training labels
- model_type: invase or invase_minus
- model_parameters:
- actor_h_dim: hidden state dimensions for actor
- critic_h_dim: hidden state dimensions for critic
- n_layer: the number of layers
- batch_size: the number of samples in mini batch
- iteration: the number of iterations
- activation: activation function of models
- learning_rate: learning rate of model training
- lamda: hyper-parameter of INVASE
"""
def __init__(self, x_train, y_train, model_type, model_parameters):
self.lamda = model_parameters['lamda']
self.actor_h_dim = model_parameters['actor_h_dim']
self.critic_h_dim = model_parameters['critic_h_dim']
self.n_layer = model_parameters['n_layer']
self.batch_size = model_parameters['batch_size']
self.iteration = model_parameters['iteration']
self.activation = model_parameters['activation']
self.learning_rate = model_parameters['learning_rate']
#Modified Code
self.x_len = x_train.shape[1]
self.y_len = x_train.shape[2]
self.dim = x_train.shape[3]
self.label_dim = y_train.shape[1]
self.model_type = model_type
optimizer = Adam(self.learning_rate)
# Build and compile critic
self.critic = self.build_critic()
self.critic.compile(loss='categorical_crossentropy',
optimizer=optimizer, metrics=['acc'])
# Build and compile the actor
self.actor = self.build_actor()
self.actor.compile(loss=self.actor_loss, optimizer=optimizer)
if self.model_type == 'invase':
# Build and compile the baseline
self.baseline = self.build_baseline()
self.baseline.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['acc'])
def actor_loss(self, y_true, y_pred):
"""Custom loss for the actor.
Args:
- y_true:
- actor_out: actor output after sampling
- critic_out: critic output
- baseline_out: baseline output (only for invase)
- y_pred: output of the actor network
Returns:
- loss: actor loss
"""
y_pred = K.reshape(y_pred, (K.shape(y_pred)[0], self.x_len*self.y_len*1))
y_true = y_true[:, 0, 0, :]
# Actor output
actor_out = y_true[:, :self.x_len*self.y_len*1]
# Critic output
critic_out = y_true[:, self.x_len*self.y_len*1:(self.x_len*self.y_len*1+self.label_dim)]
if self.model_type == 'invase':
# Baseline output
baseline_out = \
y_true[:, (self.x_len*self.y_len*1+self.label_dim):(self.x_len*self.y_len*1+2*self.label_dim)]
# Ground truth label
y_out = y_true[:, (self.x_len*self.y_len*1+2*self.label_dim):]
elif self.model_type == 'invase_minus':
# Ground truth label
y_out = y_true[:, (self.x_len*self.y_len*1+self.label_dim):]
# Critic loss
critic_loss = -tf.reduce_sum(y_out * tf.log(critic_out + 1e-8), axis = 1)
if self.model_type == 'invase':
# Baseline loss
baseline_loss = -tf.reduce_sum(y_out * tf.log(baseline_out + 1e-8),
axis = 1)
# Reward
Reward = -(critic_loss - baseline_loss)
elif self.model_type == 'invase_minus':
Reward = -critic_loss
# Policy gradient loss computation.
custom_actor_loss = \
Reward * tf.reduce_sum(actor_out * K.log(y_pred + 1e-8) + \
(1-actor_out) * K.log(1-y_pred + 1e-8), axis = 1) - \
self.lamda * tf.reduce_mean(y_pred, axis = 1)
# custom actor loss
custom_actor_loss = tf.reduce_mean(-custom_actor_loss)
return custom_actor_loss
def build_actor(self):
"""Build actor.
Use feature as the input and output selection probability
"""
actor_model = Sequential()
actor_model.add(Conv2D(self.actor_h_dim, (1, 3), padding='same', activation='linear'))
actor_model.add(InstanceNormalization())
actor_model.add(Activation(self.activation))
for _ in range(self.n_layer - 2):
actor_model.add(Conv2D(self.actor_h_dim, (1, 3), padding='same', activation='linear'))
actor_model.add(InstanceNormalization())
actor_model.add(Activation(self.activation))
actor_model.add(Conv2D(1, (1, 1), padding='same', activation='sigmoid'))
feature = Input(shape=(self.x_len, self.y_len, self.dim), dtype='float32')
selection_probability = actor_model(feature)
return Model(feature, selection_probability)
def build_critic(self):
"""Build critic.
Use selected feature as the input and predict labels
"""
critic_model = Sequential()
critic_model.add(Conv2D(self.critic_h_dim, (1, 3), padding='same', activation='linear'))
critic_model.add(InstanceNormalization())
critic_model.add(Activation(self.activation))
for _ in range(self.n_layer - 2):
critic_model.add(Conv2D(self.critic_h_dim, (1, 3), padding='same', activation='linear'))
critic_model.add(InstanceNormalization())
critic_model.add(Activation(self.activation))
critic_model.add(Flatten())
critic_model.add(Dense(self.critic_h_dim, activation=self.activation))
#critic_model.add(Dropout(0.2))
critic_model.add(Dense(self.label_dim, activation ='softmax'))
## Inputs
# Features
feature = Input(shape=(self.x_len, self.y_len, self.dim), dtype='float32')
# Binary selection
selection = Input(shape=(self.x_len, self.y_len, 1), dtype='float32')
# Element-wise multiplication
critic_model_input = Multiply()([feature, selection])
y_hat = critic_model(critic_model_input)
return Model([feature, selection], y_hat)
def build_baseline(self):
"""Build baseline.
Use the feature as the input and predict labels
"""
baseline_model = Sequential()
baseline_model.add(Conv2D(self.critic_h_dim, (1, 3), padding='same', activation='linear'))
baseline_model.add(InstanceNormalization())
baseline_model.add(Activation(self.activation))
for _ in range(self.n_layer - 2):
baseline_model.add(Conv2D(self.critic_h_dim, (1, 3), padding='same', activation='linear'))
baseline_model.add(InstanceNormalization())
baseline_model.add(Activation(self.activation))
baseline_model.add(Flatten())
baseline_model.add(Dense(self.critic_h_dim, activation=self.activation))
#baseline_model.add(Dropout(0.2))
baseline_model.add(Dense(self.label_dim, activation ='softmax'))
# Input
feature = Input(shape=(self.x_len, self.y_len, self.dim), dtype='float32')
# Output
y_hat = baseline_model(feature)
return Model(feature, y_hat)
def train(self, x_train, y_train):
"""Train INVASE.
Args:
- x_train: training features
- y_train: training labels
"""
for iter_idx in range(self.iteration):
## Train critic
# Select a random batch of samples
idx = np.random.randint(0, x_train.shape[0], self.batch_size)
x_batch = x_train[idx,:]
y_batch = y_train[idx,:]
# Generate a batch of selection probability
selection_probability = self.actor.predict(x_batch)
# Sampling the features based on the selection_probability
selection = bernoulli_sampling(selection_probability)
# Critic loss
critic_loss = self.critic.train_on_batch([x_batch, selection], y_batch)
# Critic output
critic_out = self.critic.predict([x_batch, selection])
# Baseline output
if self.model_type == 'invase':
# Baseline loss
baseline_loss = self.baseline.train_on_batch(x_batch, y_batch)
# Baseline output
baseline_out = self.baseline.predict(x_batch)
## Train actor
# Use multiple things as the y_true:
# - selection, critic_out, baseline_out, and ground truth (y_batch)
if self.model_type == 'invase':
y_batch_final = np.concatenate((np.reshape(selection, (y_batch.shape[0], -1)),
np.asarray(critic_out),
np.asarray(baseline_out),
y_batch), axis = 1)
elif self.model_type == 'invase_minus':
y_batch_final = np.concatenate((np.reshape(selection, (y_batch.shape[0], -1)),
np.asarray(critic_out),
y_batch), axis = 1)
y_batch_final = y_batch_final[:, None, None, :]
# Train the actor
actor_loss = self.actor.train_on_batch(x_batch, y_batch_final)
if self.model_type == 'invase':
# Print the progress
dialog = 'Iterations: ' + str(iter_idx) + \
', critic accuracy: ' + str(critic_loss[1]) + \
', baseline accuracy: ' + str(baseline_loss[1]) + \
', actor loss: ' + str(np.round(actor_loss,4))
elif self.model_type == 'invase_minus':
# Print the progress
dialog = 'Iterations: ' + str(iter_idx) + \
', critic accuracy: ' + str(critic_loss[1]) + \
', actor loss: ' + str(np.round(actor_loss,4))
if iter_idx % 100 == 0:
print(dialog)
def importance_score(self, x):
"""Return featuer importance score.
Args:
- x: feature
Returns:
- feature_importance: instance-wise feature importance for x
"""
feature_importance = self.actor.predict(x)
return np.asarray(feature_importance)
def predict(self, x):
"""Predict outcomes.
Args:
- x: feature
Returns:
- y_hat: predictions
"""
# Generate a batch of selection probability
selection_probability = self.actor.predict(x)
# Sampling the features based on the selection_probability
selection = bernoulli_sampling(selection_probability)
# Prediction
y_hat = self.critic.predict([x, selection])
return np.asarray(y_hat)
# +
#Concatenate input binder pairs
x_train = np.concatenate([x_1_train, x_2_train], axis=2)
x_val = np.concatenate([x_1_val, x_2_val], axis=2)
x_test = np.concatenate([x_1_test, x_2_test], axis=2)
# +
#Execute INVASE
mask_penalty = 0.05
hidden_dims = 32
n_layers = 4
epochs = 25
batch_size = 128
model_parameters = {
'lamda': mask_penalty,
'actor_h_dim': hidden_dims,
'critic_h_dim': hidden_dims,
'n_layer': n_layers,
'batch_size': batch_size,
'iteration': int(x_train.shape[0] * epochs / batch_size),
'activation': 'relu',
'learning_rate': 0.0001
}
invase_model = INVASE(x_train, pred_train, 'invase', model_parameters)
invase_model.train(x_train, pred_train)
importance_scores_test = invase_model.importance_score(x_test)
importance_scores_1_test, importance_scores_2_test = importance_scores_test[:, :, :seq_length, :], importance_scores_test[:, :, seq_length:, :]
# +
#Evaluate INVASE model on train and test data
invase_pred_train = invase_model.predict(x_train)
invase_pred_test = invase_model.predict(x_test)
print("Training Accuracy = " + str(np.sum(np.argmax(invase_pred_train, axis=1) == np.argmax(pred_train, axis=1)) / float(pred_train.shape[0])))
print("Test Accuracy = " + str(np.sum(np.argmax(invase_pred_test, axis=1) == np.argmax(pred_test, axis=1)) / float(pred_test.shape[0])))
# +
#Gradient saliency/backprop visualization
import matplotlib.collections as collections
import operator
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.colors as colors
import matplotlib as mpl
from matplotlib.text import TextPath
from matplotlib.patches import PathPatch, Rectangle
from matplotlib.font_manager import FontProperties
from matplotlib import gridspec
from matplotlib.ticker import FormatStrFormatter
def plot_protein_logo(residue_map, pwm, sequence_template=None, figsize=(12, 3), logo_height=1.0, plot_start=0, plot_end=164) :
inv_residue_map = {
i : sp for sp, i in residue_map.items()
}
#Slice according to seq trim index
pwm = pwm[plot_start: plot_end, :]
sequence_template = sequence_template[plot_start: plot_end]
entropy = np.zeros(pwm.shape)
entropy[pwm > 0] = pwm[pwm > 0] * -np.log2(np.clip(pwm[pwm > 0], 1e-6, 1. - 1e-6))
entropy = np.sum(entropy, axis=1)
conservation = np.log2(len(residue_map)) - entropy#2 - entropy
fig = plt.figure(figsize=figsize)
ax = plt.gca()
height_base = (1.0 - logo_height) / 2.
for j in range(0, pwm.shape[0]) :
sort_index = np.argsort(pwm[j, :])
for ii in range(0, len(residue_map)) :
i = sort_index[ii]
if pwm[j, i] > 0 :
nt_prob = pwm[j, i] * conservation[j]
nt = inv_residue_map[i]
color = None
if sequence_template[j] != '$' :
color = 'black'
if ii == 0 :
letterAt_protein(nt, j + 0.5, height_base, nt_prob * logo_height, ax, color=color)
else :
prev_prob = np.sum(pwm[j, sort_index[:ii]] * conservation[j]) * logo_height
letterAt_protein(nt, j + 0.5, height_base + prev_prob, nt_prob * logo_height, ax, color=color)
plt.xlim((0, plot_end - plot_start))
plt.ylim((0, np.log2(len(residue_map))))
plt.xticks([], [])
plt.yticks([], [])
plt.axis('off')
plt.axhline(y=0.01 + height_base, color='black', linestyle='-', linewidth=2)
for axis in fig.axes :
axis.get_xaxis().set_visible(False)
axis.get_yaxis().set_visible(False)
plt.tight_layout()
plt.show()
def plot_importance_scores(importance_scores, ref_seq, figsize=(12, 2), score_clip=None, sequence_template='', plot_start=0, plot_end=96, save_figs=False, fig_name=None) :
end_pos = ref_seq.find("#")
fig = plt.figure(figsize=figsize)
ax = plt.gca()
if score_clip is not None :
importance_scores = np.clip(np.copy(importance_scores), -score_clip, score_clip)
max_score = np.max(np.sum(importance_scores[:, :], axis=0)) + 0.01
for i in range(0, len(ref_seq)) :
mutability_score = np.sum(importance_scores[:, i])
letterAt_protein(ref_seq[i], i + 0.5, 0, mutability_score, ax, color=None)
plt.sca(ax)
plt.xlim((0, len(ref_seq)))
plt.ylim((0, max_score))
plt.axis('off')
plt.yticks([0.0, max_score], [0.0, max_score], fontsize=16)
for axis in fig.axes :
axis.get_xaxis().set_visible(False)
axis.get_yaxis().set_visible(False)
plt.tight_layout()
if save_figs :
plt.savefig(fig_name + ".png", transparent=True, dpi=300)
plt.savefig(fig_name + ".eps")
plt.show()
# +
np.sum(importance_scores_1_test[0, 0, :, 0] + importance_scores_2_test[0, 0, :, 0])
# +
np.max(importance_scores_1_test[0, 0, :, 0])
# +
#Visualize importance for binder 1
for plot_i in range(0, 5) :
print("Test sequence " + str(plot_i) + ":")
sequence_template = sequence_templates[l_1_test[plot_i, 0]]
plot_protein_logo(residue_map, x_1_test[plot_i, 0, :, :], sequence_template=sequence_template, figsize=(12, 1), plot_start=0, plot_end=81)
plot_importance_scores(importance_scores_1_test[plot_i, 0, :, :].T, encoder.decode(x_1_test[plot_i, 0, :, :]), figsize=(12, 1), score_clip=None, sequence_template=sequence_template, plot_start=0, plot_end=81)
#Visualize importance for binder 2
for plot_i in range(0, 5) :
print("Test sequence " + str(plot_i) + ":")
sequence_template = sequence_templates[l_2_test[plot_i, 0]]
plot_protein_logo(residue_map, x_2_test[plot_i, 0, :, :], sequence_template=sequence_template, figsize=(12, 1), plot_start=0, plot_end=81)
plot_importance_scores(importance_scores_2_test[plot_i, 0, :, :].T, encoder.decode(x_2_test[plot_i, 0, :, :]), figsize=(12, 1), score_clip=None, sequence_template=sequence_template, plot_start=0, plot_end=81)
# +
#Save predicted importance scores
model_name = "invase_" + dataset_name + "_conv" + "_zeropad_no_drop_penalty_005_full_data"
np.save(model_name + "_importance_scores_1_test", importance_scores_1_test)
np.save(model_name + "_importance_scores_2_test", importance_scores_2_test)
# +
#Binder DHD_154
seq_1 = "TAEELLEVHKKSDRVTKEHLRVSEEILKVVEVLTRGEVSSEVLKRVLRKLEELTDKLRRVTEEQRRVVEKLN"[:81]
seq_2 = "DLEDLLRRLRRLVDEQRRLVEELERVSRRLEKAVRDNEDERELARLSREHSDIQDKHDKLAREILEVLKRLLERTE"[:81]
print("Seq 1 = " + seq_1)
print("Seq 2 = " + seq_2)
encoder = IdentityEncoder(81, residue_map)
test_onehot_1 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_1), axis=0), axis=0), (batch_size, 1, 1, 1))
test_onehot_2 = np.tile(np.expand_dims(np.expand_dims(encoder(seq_2), axis=0), axis=0), (batch_size, 1, 1, 1))
test_len_1 = np.tile(np.array([[len(seq_1)]]), (batch_size, 1))
test_len_2 = np.tile(np.array([[len(seq_2)]]), (batch_size, 1))
pred_interacts = predictor.predict(x=[test_onehot_1[:, 0, ...], test_onehot_2[:, 0, ...]])[0, 0]
print("Predicted interaction prob = " + str(round(pred_interacts, 4)))
# +
x_1_test = test_onehot_1[:1]
x_2_test = test_onehot_2[:1]
#Concatenate input binder pairs
x_test = np.concatenate([x_1_test, x_2_test], axis=2)
#Execute INVASE
'''
mask_penalty = 0.05
hidden_dims = 32
n_layers = 4
epochs = 50
batch_size = 128
model_parameters = {
'lamda': mask_penalty,
'actor_h_dim': hidden_dims,
'critic_h_dim': hidden_dims,
'n_layer': n_layers,
'batch_size': batch_size,
'iteration': int(x_train.shape[0] * epochs / batch_size),
'activation': 'relu',
'learning_rate': 0.0001
}
invase_model = INVASE(x_train, pred_train, 'invase', model_parameters)
invase_model.train(x_train, pred_train)
'''
importance_scores_test = invase_model.importance_score(x_test)
importance_scores_1_test, importance_scores_2_test = importance_scores_test[:, :, :seq_length, :], importance_scores_test[:, :, seq_length:, :]
# +
save_figs = True
model_name = "invase_" + dataset_name + "_conv" + "_zeropad_no_drop_penalty_005_full_data"
pair_name = "DHD_154"
#Visualize importance for binder 1
for plot_i in range(0, 1) :
print("Test sequence " + str(plot_i) + ":")
sequence_template = sequence_templates[l_1_test[plot_i, 0]]
plot_protein_logo(residue_map, x_1_test[plot_i, 0, :, :], sequence_template=sequence_template, figsize=(12, 1), plot_start=0, plot_end=81)
plot_importance_scores(importance_scores_1_test[plot_i, 0, :, :].T, encoder.decode(x_1_test[plot_i, 0, :, :]), figsize=(12, 1), score_clip=None, sequence_template=sequence_template, plot_start=0, plot_end=81, save_figs=save_figs, fig_name=model_name + "_scores_" + pair_name + "_binder_1")
#Visualize importance for binder 2
for plot_i in range(0, 1) :
print("Test sequence " + str(plot_i) + ":")
sequence_template = sequence_templates[l_2_test[plot_i, 0]]
plot_protein_logo(residue_map, x_2_test[plot_i, 0, :, :], sequence_template=sequence_template, figsize=(12, 1), plot_start=0, plot_end=81)
plot_importance_scores(importance_scores_2_test[plot_i, 0, :, :].T, encoder.decode(x_2_test[plot_i, 0, :, :]), figsize=(12, 1), score_clip=None, sequence_template=sequence_template, plot_start=0, plot_end=81, save_figs=save_figs, fig_name=model_name + "_scores_" + pair_name + "_binder_2")
# -
| analysis/coiled_coil_binders/invase_coiled_coil_binders_zeropad_full_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gas
# language: python
# name: gas
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error
from sklearn import tree
import lightgbm as lgb
# holiday is added to gas.csv which is supplied by KOGAS
pd.options.mode.chained_assignment = None
gas = pd.read_csv("gas-h.csv", encoding='euc-kr')
groups = gas.groupby(gas.구분)
A_gas = groups.get_group("A")
B_gas = groups.get_group("B")
C_gas = groups.get_group("C")
D_gas = groups.get_group("D")
E_gas = groups.get_group("E")
G_gas = groups.get_group("G")
H_gas = groups.get_group("H")
A_gas['former'] = A_gas['공급량'].shift(periods=1, fill_value=0)
B_gas['former'] = B_gas['공급량'].shift(periods=1, fill_value=0)
C_gas['former'] = C_gas['공급량'].shift(periods=1, fill_value=0)
D_gas['former'] = D_gas['공급량'].shift(periods=1, fill_value=0)
E_gas['former'] = E_gas['공급량'].shift(periods=1, fill_value=0)
G_gas['former'] = G_gas['공급량'].shift(periods=1, fill_value=0)
H_gas['former'] = H_gas['공급량'].shift(periods=1, fill_value=0)
gas2 = pd.concat([A_gas, B_gas, C_gas, D_gas, E_gas, G_gas, H_gas], ignore_index=True)
# former has '공급량' of one hour ago
# weather.csv based on data from data.kma.go.kr
weather = pd.read_csv("weather.csv", encoding='euc-kr')
weather['시간'] = weather['시간'] + 1
total = pd.merge(left=gas2, right=weather, how="inner", on=["연월일", "시간"])
d_map = {}
for i, d in enumerate(total['구분'].unique()):
d_map[d] = i
total['구분'] = total['구분'].map(d_map)
total['연월일'] = pd.to_datetime(total['연월일'])
total['year'] = total['연월일'].dt.year
total['month'] = total['연월일'].dt.month
total['day'] = total['연월일'].dt.day
total['weekday'] = total['연월일'].dt.weekday
train_years = [2013,2014,2015,2016]
val_years = [2017]
test_year = [2018]
train = total[total['year'].isin(train_years)]
val = total[total['year'].isin(val_years)]
test = total[total['year'].isin(test_year)]
features = ['former', '구분', 'month', 'day', 'weekday', '시간', '기온(°C)', '풍속(m/s)', '강수량(mm)', '습도(%)', 'holiday']
train_x = train[features]
train_y = train['공급량']
val_x = val[features]
val_y = val['공급량']
test_x = test[features]
test_y = test['공급량']
d_train = lgb.Dataset(train_x, train_y)
d_val = lgb.Dataset(val_x, val_y)
params = {
'objective': 'regression',
'metric':'mape',
'seed':42
}
model = lgb.train(params, d_train, 500, d_val, verbose_eval=20, early_stopping_rounds=10)
y_pred = model.predict(test_x)
np.mean((np.abs(test_y - y_pred))/test_y)
# -
test_4 = test[test['구분'].isin([4])] # for '구분' E
test_4_x = test_4[features]
test_4_y = test_4['공급량']
y_pred_4 = model.predict(test_4_x)
plt.figure(figsize=(25, 6))
test_4_y_np = test_4_y.to_numpy()
diff = y_pred_4 - test_4_y_np
plt.plot(test_4_y_np)
plt.plot(diff)
| gas-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7fHb6S2zLRwV" colab_type="text"
# ## **Linear Regression**
#
# Using the Infamous Boston Housing Dataset that was available on UCI Repository but is now also included inside scikit learn
# + id="PsseZ5KwJ9Vs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="8887c59b-27cb-4274-b0e1-8f07136579ab"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + id="GG7hCjXQOoYz" colab_type="code" colab={}
#Loading the dataset
from sklearn.datasets import load_boston
boston_dataset = load_boston()
# + id="oNBiYY0XO2iZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4140f9e4-0db0-42cc-8aba-d1c92bc99fe2"
type(boston_dataset)
# + id="ZNIcYBt_Fbyn" colab_type="code" colab={}
boston_dataset.
# + id="veou8VEcO4bT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dd6fcef3-890a-4db9-a361-a6372db3bef9"
boston_dataset.keys()
# + [markdown] id="SyrEAcvsPO-S" colab_type="text"
# data: contains the information for various houses
#
# target: prices of the house
#
# feature_names: names of the features
#
# DESCR: describes the dataset
# + id="hdU5vvcgPIbZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 894} outputId="ffed621d-eadf-4c5e-a7ce-fd1fbbaf302a"
print(boston_dataset.DESCR)
# + id="NS-Z9ny6PS1H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="65e38fb7-b4c3-44c0-cd69-e52f26acd35c"
boston = pd.DataFrame(boston_dataset.data, columns=boston_dataset.feature_names)
boston.head()
# + id="gg4JDCDzPVra" colab_type="code" colab={}
boston['MEDV'] = boston_dataset.target
# + id="gQl9O9wWPd9I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="bbca5bc2-77c6-464f-941f-e1a8ede55f9e"
boston.head()
# + [markdown] id="9Z3WAP_-PiDy" colab_type="text"
# Preprocessing
# + id="9V0BJB70PfY8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="8e092a26-0c22-4bcf-ca6a-88799f8fed91"
#To find null values
boston.isnull().sum()
# + [markdown] id="YY_2pL1aPmvh" colab_type="text"
# Exploratory Data Analysis
# + id="tqWf-9HyPkJh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 534} outputId="d1ac5b9a-e5d6-4f8b-806f-e21989c4e3dd"
#distribution of the target variable MEDV
#Represents Normal/Gaussian Distribution
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns.distplot(boston['MEDV'], bins=30)
# + [markdown] id="T14PeV3hQCkx" colab_type="text"
# Correlation Coeff
#
# The correlation coefficient ranges from -1 to 1. If the value is close to 1, it means that there is a strong positive correlation between the two variables. When it is close to -1, the variables have a strong negative correlation.
# + id="FUY3UuPBPs0h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 521} outputId="bd814647-f065-4af7-c9f9-c68646c0d134"
#measuring the linear relationships between the variables
correlation_matrix = boston.corr().round(2)
# annot = True to print the values inside the square
sns.heatmap(data=correlation_matrix, annot=True)
# + [markdown] id="XLmE3Nb6QMsZ" colab_type="text"
# Observations:
#
# To fit a linear regression model, we select those features which have a high correlation with our target variable MEDV. By looking at the correlation matrix we can see that RM has a strong positive correlation with MEDV (0.7) where as LSTAT has a high negative correlation with MEDV(-0.74).
#
# An important point in selecting features for a linear regression model is to check for multi-co-linearity. The features RAD, TAX have a correlation of 0.91. These feature pairs are strongly correlated to each other. We should not select both these features together for training the model.Same goes for the features DIS and AGE which have a correlation of -0.75.
# + id="Uq4E4ritP-in" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 355} outputId="85fe749c-3e36-43f1-caa8-c73366d56257"
plt.figure(figsize=(20, 5))
features = ['LSTAT', 'RM']
target = boston['MEDV']
for i, col in enumerate(features):
plt.subplot(1, len(features) , i+1)
x = boston[col]
y = target
plt.scatter(x, y, marker='o')
plt.title(col)
plt.xlabel(col)
plt.ylabel('MEDV')
# + id="wyfH_0mkQTPv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="e8efbd67-c00a-4633-a178-c97b08130c5a"
#Could have also done this
plt.scatter(boston['LSTAT'],boston['MEDV'])
# + [markdown] id="a9sATMTbQ3Xx" colab_type="text"
# Observations:
#
# The prices increase as the value of RM increases linearly. There are few outliers and the data seems to be capped at 50.
#
# The prices tend to decrease with an increase in LSTAT. Though it doesn’t look to be following exactly a linear line.
# + [markdown] id="Mv16urcAQ9ry" colab_type="text"
# Preparing the data for training the model
# + id="1du8gQmCJ4hd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="c50bcc38-7230-4e40-f57a-d3f02f0195b1"
pd.DataFrame(data = {'LSTAT':boston['LSTAT'],'RM':boston['RM']})
# + id="yIhjogXpQyCW" colab_type="code" colab={}
X = pd.DataFrame(np.c_[boston['LSTAT'], boston['RM']], columns = ['LSTAT','RM'])
Y = boston['MEDV']
# + id="Xqv2BCqFKSNN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="452acb87-1cf7-4da4-bb17-494480b0c7fc"
X
# + id="_6cb5db1Q-IG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="e052c178-836c-496e-e0f3-77efee1b1d59"
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state=101)
print(X_train.shape)
print(X_test.shape)
print(Y_train.shape)
print(Y_test.shape)
# + id="qwyNJYdlRALa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ae39a726-eb2f-494f-9d01-664775716214"
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error,r2_score
lin_model = LinearRegression()
lin_model.fit(X_train, Y_train)
# + id="k2uEM2pvRFAY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="6cfd8be5-dd5b-4508-fe09-e8330145ee32"
# model evaluation for training set
y_train_predict = lin_model.predict(X_train)
rmse = (np.sqrt(mean_squared_error(Y_train, y_train_predict)))
r2 = r2_score(Y_train, y_train_predict)
print("The model performance for training set")
print("--------------------------------------")
#print('RMSE is:',rmse)
print('RMSE is {}'.format(rmse))
print('R2 score is {}'.format(r2))
print("\n")
# model evaluation for testing set
y_test_predict = lin_model.predict(X_test)
rmse = (np.sqrt(mean_squared_error(Y_test, y_test_predict)))
r2 = r2_score(Y_test, y_test_predict)
print("The model performance for testing set")
print("--------------------------------------")
print('RMSE is {}'.format(rmse))
print('R2 score is {}'.format(r2))
# + id="RdjmvVA_Q9Ws" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="04372fd6-900c-4301-e92d-588bef798a8e"
s1 = ['a', 'b', np.nan]
s1
# + id="w7CrQ-8HRB13" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="a510d1dd-36bc-423e-927d-cf02af16e3fc"
pd.get_dummies(s1)
# + [markdown] id="c4OZsO33Rw_h" colab_type="text"
# ## **Logistic Regression**
# + [markdown] id="Ud1zcIyLeVxj" colab_type="text"
# The Dataset can be taken from https://www.kaggle.com/c/titanic/data?select=train.csv
# + id="qlSCPvXvUCc1" colab_type="code" colab={}
#Loading the Dataset
train = pd.read_csv("/content/drive/My Drive/IEEE/MicroHack-02/train.csv")
# + id="tn6ALiuJYEMN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="461f2657-05fc-421d-b4ad-9bbb7f0d3b9f"
train.head()
# + id="_f8kuXfOZjoC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="16da7a15-836e-44c3-c851-0d7921172d2d"
train.count()
# + id="uy3_VPJBSiNH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="0c506beb-104b-4691-a699-b4ce841247ca"
train['Cabin'].value_counts()
# + [markdown] id="XSYZp7jmaA0a" colab_type="text"
# Exploratory Data Analysis
# + id="rNctMv8KZ2h7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 534} outputId="4357277b-a75b-4dbc-d6ca-f6b06c16a443"
sns.countplot(x='Sex',data=train)
# + id="TU8j-wQ9Zn4J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 534} outputId="babde1e2-1d52-4a71-9948-f26e3d75e027"
sns.countplot(x='Survived', hue='Sex', data=train)
# + id="h5S2MJNHZxgH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 534} outputId="8a067ec3-f3dc-4c14-e9be-660bc60de69b"
sns.countplot(x='Survived', hue='Pclass', data=train)
# + [markdown] id="KlulmTVgUT1S" colab_type="text"
# https://towardsdatascience.com/understanding-boxplots-5e2df7bcbd51
# + id="CYiKXaybZ8cQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 465} outputId="79bc5589-837f-4630-e439-0086f97fca9f"
#Plotting a Boxplot
plt.figure(figsize=(10,7))
sns.boxplot(x='Pclass',y='Age',data=train)
# + id="bLJDmB-XU8wQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 465} outputId="71526bcd-1aa8-47f3-a202-09396c41c8a0"
#Plotting a Boxplot
plt.figure(figsize=(10,7))
sns.boxplot(x='Survived',y='Age',data=train)
# + id="vVCf0ijFaJuf" colab_type="code" colab={}
#Let’s impute average age values to null age values:
def add_age(cols):
Age = cols[0]
Pclass = cols[1]
if pd.isnull(Age):
return int(train[train["Pclass"] == Pclass]["Age"].mean())
else:
return Age
# + id="sY-M_ipVaWcW" colab_type="code" colab={}
train["Age"] = train[["Age", "Pclass"]].apply(add_age,axis=1)
# + id="4Em9jqD8aYFs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="6fc281a7-92ca-4a77-b210-27c78b5835a4"
train.head()
# + id="hRjajKqCacks" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="30f611ae-8001-4b57-fe56-295d4ec4c7a4"
train.isnull().sum()
# + id="ZwOX4EzGcsDo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="e0c5425c-9db5-4566-bb8e-fba7354da197"
train.drop("Cabin",inplace=True,axis=1)
# + id="-y9ay4vjcvY0" colab_type="code" colab={}
#removing some rows with Null Values
train.dropna(inplace=True)
# + id="NBu5PjYKWM9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="95b2523c-c8c7-4e66-e63d-969082338ca5"
train.head()
# + id="PkyUrGpRczyU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="26e45f99-d6b5-4ba8-a513-5ea4c1733041"
pd.get_dummies(train["Sex"])
# + id="7Pt6uvGtc1Zo" colab_type="code" colab={}
sex = pd.get_dummies(train["Sex"],drop_first=True)
# + id="HAc0W4_Vc4Y7" colab_type="code" colab={}
embarked = pd.get_dummies(train["Embarked"],drop_first=True)
pclass = pd.get_dummies(train["Pclass"],drop_first=True)
# + id="cuIer9DFc7mD" colab_type="code" colab={}
train = pd.concat([train,pclass,sex,embarked],axis=1)
# + id="3O2foC9CWnFf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="9b072e41-7b36-475d-ec1a-65c3945b5b27"
train.head()
# + id="OZjslhkmdDHR" colab_type="code" colab={}
train.drop(["PassengerId","Pclass","Name","Sex","Ticket","Embarked"],axis=1,inplace=True)
# + id="72gPePKHdGls" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="878ee751-5516-483f-88b9-905246403daa"
train.head()
# + id="FKlVKabxdIdk" colab_type="code" colab={}
X = train.drop("Survived",axis=1)
y = train["Survived"]
# + id="LUM45mBKdpGG" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 101)
# + id="vtSTUaDjdrL5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="950c8cb5-d61d-4452-ba60-1639fac0f8cb"
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression(random_state=101)
logmodel.fit(X_train,y_train)
# + id="TcjHCeg3dx8j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="4a91eb4d-54ca-42d2-96c9-48d7d681d909"
logmodel = LogisticRegression(max_iter=500,random_state=101)
logmodel.fit(X_train,y_train)
# + id="9evJLQMDd5E_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="57dfa1d2-aaca-460f-cdff-da36c3931feb"
predictions = logmodel.predict(X_test)
from sklearn.metrics import classification_report,accuracy_score
print(classification_report(y_test, predictions))
# + id="6-8AdL-RX3gu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ce998c4a-be2d-4d3b-a5a7-238094a910a3"
print(accuracy_score(y_test,predictions))
# + id="xLSPogSpd8a7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="6fa57b04-3faf-4d31-e709-ed998b85c94e"
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, predictions)
# + [markdown] id="pAVmvzlie-Qm" colab_type="text"
# Thanks to everyone.
| MicroHack-02/Project_02_MicroHacks_Linear_and_Logistic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 68} colab_type="code" executionInfo={"elapsed": 1030, "status": "ok", "timestamp": 1528314480195, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="J4ju1kvPpugE" outputId="1f202837-1209-42c4-a56b-08b7b2ade1f2"
# %matplotlib inline
import numpy as np
import scipy
import math
import json
import pprint
import time
import copy
from matplotlib import pyplot as plt
import itertools
import pandas as pd
import cProfile
import csv
import inspect
import sys
sys.path.insert(0, '../../')
sys.path.insert(0, '../')
from mx_sys.power_calcs import power_calcs as makani_FBL
from mx_sys.power_calcs import kite_pose
from mx_sys.power_calcs import kite_loop
from mx_sys.power_calcs import kite_path
import m600_fbl_config_manager as cm
import resource_fbl_manager as rm
reload(makani_FBL)
reload(cm)
reload(rm)
# + [markdown] colab_type="text" id="HJLDKnoPpugL"
# # Setup Kite and Environment
#
# The easiest way to create a kite and resource is to use the managers.
#
# However, they are both just dictionaries. Required and optional elements of the dictionary are specified in the docstrings for the various objects that use them. You can always create, edit, and overwrite the various parts of the configs and resource however you'd like manually.
#
# There are a several options for aero models. There are 2 types:
# 1. Body coefficient models
# - Provide cx, cy, cz as a function of alpha and (optional) beta
# 2. Aero coefficient models
# - Provide cL, cY, cD as a function of alpha and (optional) beta
#
# Either type must also return moment coefficients cl, cm, and cn.
#
# See docstrings for details on how to name things, and error messages will point out missing functions.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 714} colab_type="code" executionInfo={"elapsed": 932, "status": "ok", "timestamp": 1528314481152, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="Qzmk6wvvpugM" outputId="8c6c1652-597d-4a7c-f4de-023e45c037c2"
# using the resource and config managers
resource = rm.GetResourceByName()
other_resource = rm.MakeResourceByShearAndHref(0.2, 80., 1.075, 8.)
base_kite = cm.GetConfigByName()
#M600 does NOT SOLVE high winds with roll limits in place
#removing those limits for a clean example
base_kite.pop('roll_min')
base_kite.pop('roll_max')
print 'Resource:'
pprint.pprint(resource)
print
print 'Other Resource:'
pprint.pprint(resource)
print
print 'Base Kite:'
pprint.pprint(base_kite.keys())
print
# example of resource functions
print inspect.getsource(resource['v_w_at_height'])
print inspect.getsource(other_resource['v_w_at_height'])
# defining a config manually
def rotors_simple(rho, v_a, force):
power_shaft = v_a * -force
out = {'power_shaft': power_shaft}
return out
other_kite = {
'alpha_min': 0.,
'alpha_max': 7.,
'cD_eff_tether': 0.06,
'gs_position': (0.,0.,0.),
'eta_shaft_to_pad': lambda p: 0.82,
'bridle_moment_from_tether_pitch_roll': (
lambda p, r: np.array([0., 0., 0.])),
'l_tether': 430.,
'v_a_min': 30.,
'v_a_max': 70.,
'shaft_power_from_drag_power': rotors_simple,
'rotor_thrust_center': np.array([0., 0., 0.]),
'm_kite': 1200.,
'h_min': 80.,
'm_tether': 300.,
'aero_coeff_from_alpha_beta': lambda a, b, o: {
'cL': 0.11 * a, 'cY': 0.01 * b, 'cD': 0.05 + (0.04 * a)**2,
'cl': 0.1, 'cm': 0.2, 'cn': 0.},
'CG': np.array([0., 0., 0.]),
'tension_max': 250000.,
'beta_min': -5.,
'beta_max': 5.,
'c': 1.28,
'b': 25.66,
's': 32.9,
'inertia': np.array([[3000., 0., 0.],
[0., 3000., 0.],
[0., 0., 3000.]]),
'power_shaft_max': 900000.}
# -
# There are several helper functions to plot things scattered about. As an example, we can inspect the aero database to find where the Loyd limit is.
#
# The Loyd limit is defined as:
# $\zeta_{max} = \frac{4}{27}\frac{C_L^3}{C_D^2}$
#
# $v_{a\_best\_power} \approx v_{k\_best\_power} \approx \frac{2}{3}\frac{L}{D} v_w$
#
# Derivations won't be shown here, but results are below:
# +
zeta, cL, L_over_D, alpha, beta = makani_FBL.calc_loyd_optimums(base_kite)
plots = cm.PlotKiteAero(base_kite, keys=['zeta'])
# + [markdown] colab_type="text" id="-_tB6xHHpugQ"
# # Create Path Object (optional)
#
# Section is optional as you don't need to know how to create a path object as they are usually created and managed by the higher level object, KiteLoop.
# KitePath creates and holds all the path definition needed for the FBL model.
#
# You can create it by manually making the params, or by using the config manager helper function to make an entire set of args for the M600 by specifying a min height and a loop radius.
#
# A path mostly just contains the positions, but also has a few summary values.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 391} colab_type="code" executionInfo={"elapsed": 1011, "status": "ok", "timestamp": 1528314482201, "user": {"displayName": "<NAME>", "photoUrl": "//<KEY>", "userId": "101021799646106665401"}, "user_tz": 240} id="di-morskpugR" outputId="ec5865da-0401-4974-9822-efb09e0b565c"
# using config helper function to get args and splatting it into KitePath
path_args = cm.GetPathArgsByR_LoopAndMinHeight(base_kite, 100., 130.)
print 'Path_Args:'
pprint.pprint(path_args)
print
path = kite_path.KitePath(config=base_kite, **path_args)
# pull off one position to use later
position = path.positions[0]
# print some stuff from the path
print 'Example Position:'
pprint.pprint(position)
print
print 'Example path summary data:'
print 'Half cone angle: %0.2f deg' % math.degrees(path.half_cone_angle)
print('Min height in loop: %0.1f m' % path.h_min)
print 'Virtual Hub Height: %0.2f m' % path.h_hub
# make a path manually
shape_params = {'type': 'circle',
'r_loop': 140.}
location_params = {'azim': 0.,
'incl': 0.}
other_path = kite_path.KitePath(shape_params, location_params, base_kite)
# + [markdown] colab_type="text" id="g3l_sNvYpugT"
# # Creating a KitePose object (optional)
# A KitePose is the base level object to complete a force balance. It is a single point model of a kite. There are many options here, but this section is optional as typical use only uses the higher level objects, which will manage KitePoses automatically.
#
# There are 2 solve options:
# 1. Lift roll angle is not provided
# - It is solved for to make the residual zero, if possible
# 2. Lift roll angle is provided
# - Orientation is fully specified, but the force balance is not likely
#
# For either solution, you must specify a kite speed, either airspeed (v_a) or inertial speed (v_k), and optionally an acceleration along the flight path, body rates, and body accelerations. Gravity and curvature accelerations are applied as a result from the speed and path definition.
#
# Poses can always be solved, but user beware. Just because it solves doesn't mean all the constraints are valid or that the forces were able to be balanced. You need to check it yourself.
#
# Below shows running for a specified lift_roll_angle and without. As a solution fills the pose state, a new pose is needed each time - there is no way to reset a pose.
#
# pose.state is the holder for all the info about the pose. When in doubt about what pose info is available and what keys to use when accessing data from higher level objects, inspect this dictionary.
#
# ## Important Note:
# **FBL will return solutions that aren't valid - ie: either some constraint is violated or the force balance wasn't possible with the parameters given.**
#
# The results are just a function of the state provided. If no lift roll angle is provided, we can solve for the lift roll angle to meet the force balance, which usually works unless you lack enough lift.
#
# The user has information to determine the validity of the solve. Every object has a "valid" attribute that keeps track of this. Violated constraints will show up in KitePose.constraints_violated (also stored in state).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 3315} colab_type="code" executionInfo={"elapsed": 931, "status": "ok", "timestamp": 1528314483201, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="vsm5IuzBpugU" outputId="2836f678-543e-4df7-e23a-8b5ff63c27f2"
# standard solver
# solving with a known aero state
# using kite with body coefficient aero model and v_a
pose = kite_pose.KitePose(position, resource, base_kite,
v_a=50., alpha=5., beta=0.,
v_w_at_h_ref=7.5, verbose=True)
pose.solve(solve_type='unknown_roll')
print 'Known aero state solution power:', pose.state['power']
print 'Pose is valid: ', pose.valid
print
# solving with a known lift_roll_angle
# using kite with aero coefficient aero model and v_k, with accel
pose = kite_pose.KitePose(position, resource, base_kite,
v_a=50., alpha=5., beta=0., lift_roll_angle=-0.1,
v_w_at_h_ref=7.5, verbose=True)
pose.solve(solve_type='full')
print 'Known aero state solution power:', pose.state['power']
print 'Pose is valid: ', pose.valid
print
print 'Example of data stored in pose.state using solution from known tension solution.'
pprint.pprint(pose.state)
# + [markdown] colab_type="text" id="VCsRwl56pugX"
# # Creating KiteLoop objects
#
# KiteLoop objects are a container for a "set" of poses that define an entire loop. The KiteLoop applies accelerations to each pose to make them consistent with the speed strategy applied.
#
# Any necessary variable that isn't specified is determined by an optimizer, with a default seed. Alternatively, you can explictly define a variable to optimize over and set your own seed value and parameterization type. See docstring for parameterization types and usage.
#
# There are also optimization options, under the keyword arg 'opt_params'. Selection of optimization options is the single most finicky part of the process, and the most likely to cause errors or non-optimal results. There are a lot of options for the optimizer. See docstring for options, but defaults should be pretty good.
#
# Two frequently used ones are 'tol' and 'constraint_stiffness'.
# 'tol' is the convergence tolerance. Higher values will take longer to finish, but results will be smoother and make more power. Typical values are ~0.01 - 0.001.
# 'constraint_stiffness' is a weighting factor for constraints. Higher values will make the model more quickly shy away from constraints violations, while lower ones will let the model optimize power first, then try and meet constraints, but can be harder to converge. Typical values are ~1. to 0.01.
#
# Note that at high wind speeds, you many need to provide a better seed, as this space is highly constrained. The KitePowerCurve handles this automatically by seeding loops with the previous loops best v_k strategy, which usually works well for finding a solution.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 2895} colab_type="code" executionInfo={"elapsed": 5626, "status": "ok", "timestamp": 1528314488908, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="o51mIt5YpugY" outputId="033c4283-2623-4392-8305-4884e81b86ee"
# make a loop with some options and solve it
loop = kite_loop.KiteLoop(
resource, base_kite, v_w_at_h_ref=9.,
verbose=True,
opt_params={'tol':0.01,
'constraint_stiffness': 0.01,
'maxiter':700},
vars_to_opt={'v_a': {'param_type': 'spline',
'values': [40.]*6,
'max_step': 5.}})
loop.solve()
# look at some summary data about loop
print
print 'Loop mean power: %0.2f W' % loop.power
print 'Loop average v_a: %0.2f m/s' % loop.data_loop['v_a_avg_time']
print 'Loop valid: ', loop.valid
print
# use Dataframe plotting library
loop.data_poses.plot(y=['v_k', 'v_a', 'power'],
subplots=True, figsize=(15, 12), layout=(2,2))
# use Dataframe feature in jupyter to make table of data
loop.data_poses
# + [markdown] colab_type="text" id="7C7EsdIKpugb"
# ## KiteLoop - Using specific values instead of the optimizer
#
# There are several ways to specify values to hold fixed. If all values are specified, the optimizer isn't used at all, and the solution time is very quick (thousandths of a sec).
#
# See the example below for formats to specify a particular solution, or the docstring. Any single variable can be dropped out and the optimizer will take over only that variable, using defaults optimizer options and seed unless something is specified in the "vars_to_opt" dictionary, as shown in the example above.
#
# This methodology is useful when you just want to locally perturb something to see sensitivities, holding everything else constant. In this example, we sweep out varius azimuths, showing the power variation as the loop is slewed to the right.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 1082} colab_type="code" executionInfo={"elapsed": 2032, "status": "ok", "timestamp": 1528314490975, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="HM52Fkgfpugc" outputId="12ac3e74-9e5c-4ee7-fddf-302b1ee76568"
loops = []
azims = np.linspace(-0.5, 0, 6)
for azim in azims:
temp_loop = kite_loop.KiteLoop(
resource, base_kite, v_w_at_h_ref=7.5, verbose=True,
path_location_params={'azim': azim,
'incl': 0.577},
pose_states_param={'alpha': {'param_type': 'linear_interp', 'values': [4., 3., 3.5, 4.]}},
pose_states={'beta': np.array([-3.]*18),
'v_k': np.array([ 42.18902103, 44.9445889 , 47.92323029, 50.84411908,
53.44207691, 55.55595166, 57.06702792, 57.85671546,
57.80642404, 56.79756345, 54.72297007, 51.73829244,
48.26199128, 44.72395391, 41.55406766, 39.18221986,
38.03770241, 38.34796218])},
path_shape_params={'type': 'circle',
'r_loop': 160.,
'num_pos': 18})
temp_loop.solve()
print
loops.append(temp_loop)
plt.figure(figsize=(10,7))
plt.title('Power vs Normalized Distance Around Loop for Different Azimuths')
plt.ylabel('Power [W]')
for azim, loop in zip(azims, loops):
plt.plot(loop.data_poses['dist_norm'],
loop.data_poses.power_shaft,
label=azim)
plt.legend()
# + [markdown] colab_type="text" id="6zmB5jBSpugg"
# # Using the Plotly Plotting Library
#
# The KiteLoop contains a few tools that output the 3D force solution, as well as variables colored by value around the loop.
#
# The plotting tool can be found at:
# mx_modeling/visualizations/power_calcs_plotting_tool/plotter.html
#
# Open it directly with your browser, and point it to the files generated by the KiteLoop.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 85} colab_type="code" executionInfo={"elapsed": 1170, "status": "ok", "timestamp": 1528314492174, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="_MhmvT1Upugi" outputId="ddd6be17-0f93-4e0a-91fe-6c77d11aa5a6"
# make files for plotly plotter
loop.gen_loop_vec_plot_file('test_forces.json')
loop.gen_loop_positions_plot_file('test_colors_roll_angle.json',
var_to_color='tether_roll_angle')
# + [markdown] colab_type="text" id="g4d8QBVtpugs"
# # Creating a KitePowerCurve object
#
# A KitePowerCurve object creates KiteLoop objects for each wind speed in the range.
#
# All the same optimization parameters, options, etc that were available at the loop level are available here as well (opt_params, vars_to_opt, loop_params, path_shape_params, path_location_params), with the same effect.
#
# Here is an example of not specifying anything and letting the defaults do the job.
#
# Solutions for previous loops are used to seed the solutions for future loops, usually enabling the KitePower curve to more quickly and easily find solutions.
#
# There are three key outputs that trim the data to make a power curve.
# 1. KitePowerCurve.powers is average power for each loop
# 2. KitePowerCurve.powers_valid has invalid loop powers set to None
# 3. KitePowerCurve.powers_final has negative powers, invalid powers, and powers at virtual hub speeds outside of cut in and cut out (if provided in the kite config) set to zero
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 68} colab_type="code" executionInfo={"elapsed": 21648, "status": "ok", "timestamp": 1528314514878, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="rq2sAGu7pugt" outputId="bb49a9ee-1ca2-4912-9422-7c5ba471ea36"
pc = makani_FBL.KitePowerCurve(resource, base_kite,
v_w_at_h_ref_range=(2., 10.), v_w_step=2.)
pc.solve()
print 'Validity of each loop: ', pc.valids
# + [markdown] colab_type="text" id="G_0OrQktpugz"
# ## Multiple Ways to Get Data
#
# There's a ton of data in the KitePowerCurve object, and lot of ways to get it.
# Summary level data is an attribute of the object, and the loop summaries are aggregated into a Dataframe object called:
# self.data_loops
#
# The loops themselves are available in a list at self.loops. You can then pull out a loop and access all the pose data, or use the loop plotting tools.
#
# Below are examples of different ways to get data out.
# 1. Directly access the data and do whatever with it: math, plot it, whatever
#
# Most key data is available as an attribute as well, but the full set is in the data_loops or data_poses Dataframes.
#
# 2. Use the pandas Dataframe to do whatever, including its own plotting library
# 3. Use the plotting helper functions included with some of the objects to plot things . These built in plotting tools do a lot more formatting for you, as shown in the example below.
#
# **Note: The surface plots are way nicer and actually functional if you're using a fully updated version of matplotlib with a local kernel**
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 1917} colab_type="code" executionInfo={"elapsed": 1679, "status": "ok", "timestamp": 1528314516575, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="PpiepahYpug0" outputId="255f1105-f416-4baf-caa9-cfbdd0af81c1"
# 1: access data directly, some as attribute, some in data
plt.plot(pc.v_ws_at_h_hub, pc.data_loops['zeta_padmount_avg_time'])
# 2: use dataframe tools
pc.data_loops.plot(y='zeta_padmount_avg_time')
# 3: use built in plotting helper functions
pc.plot_loop_data(ys=['zeta_padmount_avg_time'])
pc.plot_pose_data_as_surf(keys=['power', 'v_a'])
pc.plot_power_curve()
# + [markdown] colab_type="text" id="RZCqE6zBpuhc"
# # Putting it all together
#
# This is the minimum set of things needed to calculate a power curve:
#
# This example has NOT removed the roll limits, which is why the power curve has a big dip - when invalid solutions are found, the loop inclination is raised until it works, but this is a big performance hit.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "height": 448} colab_type="code" executionInfo={"elapsed": 99700, "status": "ok", "timestamp": 1528314672420, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-SK4pN566-Lo/AAAAAAAAAAI/AAAAAAAACJw/XMl00Uy03gk/s50-c-k-no/photo.jpg", "userId": "101021799646106665401"}, "user_tz": 240} id="kR2Kn9vPpuhd" outputId="700ec086-201e-4ec0-f552-83acc5e2fced"
# we need a kite
m600 = cm.GetConfigByName()
# we need a resource
china_lake = rm.GetResourceByName('CL_nom')
# then we make and solve a power curve
m600pc = makani_FBL.KitePowerCurve(china_lake, m600)
m600pc.solve()
# then we do things with it
m600pc.plot_power_curve()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="nUCNn3VPpuhl"
| analysis/force_balance_loop/tutorial/FBL_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Python imports
from __future__ import division, print_function, unicode_literals
import numpy as np # Matrix and vector computation package
import matplotlib.pyplot as plt # Plotting library
import math
import time
# Allow matplotlib to plot inside this notebook
# %matplotlib inline
# Set the seed of the numpy random number generator so that the tutorial is reproducable
np.random.seed(seed=1)
from sklearn import datasets, metrics # data and evaluation utils
from sklearn.datasets import fetch_openml
from sklearn.model_selection import train_test_split
from matplotlib.colors import colorConverter, ListedColormap # some plotting functions
import itertools
import collections
import csv
# +
# load the data from scikit-learn.
digits = fetch_openml('mnist_784')
# Load the targets.
# Note that the targets are stored as digits, these need to be
# converted to one-hot-encoding for the output sofmax layer.
n = digits.target.shape[0]
T = np.zeros((n,10))
T[np.arange(len(T)), digits.target.astype(int)] += 1
# Divide the data into a train and test set.
X_train, X_test, T_train, T_test = train_test_split(
digits.data, T, test_size=0.4)
# Divide the test set into a validation set and final test set.
X_validation, X_test, T_validation, T_test = train_test_split(
X_test, T_test, test_size=0.5)
print(">>>>>>DATASET FEATURE SPACE<<<<<<")
print(">>>TRAIN<<<")
print(X_train.shape)
print(T_train.shape)
print(">>>VALIDATION<<<")
print(X_validation.shape)
print(T_validation.shape)
print(">>>TEST<<<")
print(X_test.shape)
print(T_test.shape)
#with open("T_TEST_DATA.csv","w") as my_csv:
# csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
# csvWriter.writerows(T_test)
#with open("X_TEST_DATA.csv","w") as my_csv:
# csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
# csvWriter.writerows(X_test)
# +
# Define the non-linear functions used
def logistic(z):
return 1 / (1 + np.exp(-z))
def logistic_deriv(y): # Derivative of logistic function
return np.multiply(y, (1 - y))
def ReLU(x):
return np.maximum(x, 0, x)
def ReLU_Der(out):
out[out<=0] = 0
out[out>0] = 1
return out
def softmax(z):
return np.exp(z) / np.sum(np.exp(z), axis=1, keepdims=True)
# +
class Layer(object):
"""Base class for the different layers.
Defines base methods and documentation of methods."""
def get_params_iter(self):
"""Return an iterator over the parameters (if any).
The iterator has the same order as get_params_grad.
The elements returned by the iterator are editable in-place."""
return []
def get_params_grad(self, X, output_grad):
"""Return a list of gradients over the parameters.
The list has the same order as the get_params_iter iterator.
X is the input.
output_grad is the gradient at the output of this layer.
"""
return []
def get_output(self, X):
"""Perform the forward step linear transformation.
X is the input."""
pass
def get_input_grad(self, Y, output_grad=None, T=None):
"""Return the gradient at the inputs of this layer.
Y is the pre-computed output of this layer (not needed in this case).
output_grad is the gradient at the output of this layer
(gradient at input of next layer).
Output layer uses targets T to compute the gradient based on the
output error instead of output_grad"""
pass
class LinearLayer(Layer):
"""The linear layer performs a linear transformation to its input."""
def __init__(self, n_in, n_out):
"""Initialize hidden layer parameters.
n_in is the number of input variables.
n_out is the number of output variables."""
self.W = np.random.randn(n_in, n_out) * 0.1
self.b = np.zeros(n_out)
def get_params_iter(self):
"""Return an iterator over the parameters."""
return itertools.chain(np.nditer(self.W, op_flags=['readwrite']),
np.nditer(self.b, op_flags=['readwrite']))
def get_params_array(self):
"""Return arrays of the parameters."""
return np.array(self.W), np.array(self.b)
def get_output(self, X):
"""Perform the forward step linear transformation."""
return X.dot(self.W) + self.b
def get_params_grad(self, X, output_grad):
"""Return a list of gradients over the parameters."""
JW = X.T.dot(output_grad)
Jb = np.sum(output_grad, axis=0)
return [g for g in itertools.chain(np.nditer(JW), np.nditer(Jb))]
def get_input_grad(self, Y, output_grad):
"""Return the gradient at the inputs of this layer."""
return output_grad.dot(self.W.T)
class LogisticLayer(Layer):
"""The logistic layer applies the logistic function to its inputs."""
def get_output(self, X):
"""Perform the forward step transformation."""
return logistic(X)
def get_input_grad(self, Y, output_grad):
"""Return the gradient at the inputs of this layer."""
return np.multiply(logistic_deriv(Y), output_grad)
class ReLULayer(Layer):
"""The ReLU layer applies the Rectified Linear Units function to its inputs."""
def get_output(self, X):
"""Perform the forward step transformation."""
return ReLU(X)
def get_input_grad(self, Y, output_grad):
"""Return the gradient at the inputs of this layer."""
return np.multiply(ReLU_Der(Y), output_grad)
class SoftmaxOutputLayer(Layer):
"""The softmax output layer computes the classification propabilities at the output."""
def get_output(self, X):
"""Perform the forward step transformation."""
return softmax(X)
def get_input_grad(self, Y, T):
"""Return the gradient at the inputs of this layer."""
return (Y - T) / Y.shape[0]
def get_cost(self, Y, T):
"""Return the cost at the output of this output layer."""
return - np.multiply(T, np.log(Y)).sum() / Y.shape[0]
# +
# Define a sample model to be trained on the data
hidden_neurons_1 = 500 # Number of neurons in the first hidden-layer
hidden_neurons_2 = 150 # Number of neurons in the second hidden-layer
print("HIDDEN LAYER 1 NEURONS: {}".format(hidden_neurons_1))
print("HIDDEN LAYER 2 NEURONS: {}".format(hidden_neurons_2))
# Create the model
layers_SG = [] # Define a list of layers
#layers_ReLU = []
layers_ReLU_SG = []
# Add first hidden layer
layers_SG.append(LinearLayer(X_train.shape[1], hidden_neurons_1))
layers_SG.append(LogisticLayer())
#DEACTIVATED UNTIL THE DYING ReLU PROBLEM IS FIXED
#layers_ReLU.append(LinearLayer(X_train.shape[1], hidden_neurons_1))
#layers_ReLU.append(ReLULayer())
layers_ReLU_SG.append(LinearLayer(X_train.shape[1], hidden_neurons_1))
layers_ReLU_SG.append(ReLULayer())
# Add second hidden layer
layers_SG.append(LinearLayer(hidden_neurons_1, hidden_neurons_2))
layers_SG.append(LogisticLayer())
#DEACTIVATED UNTIL THE DYING ReLU PROBLEM IS FIXED
#layers_ReLU.append(LinearLayer(hidden_neurons_1, hidden_neurons_2))
#layers_ReLU.append(ReLULayer())
layers_ReLU_SG.append(LinearLayer(hidden_neurons_1, hidden_neurons_2))
layers_ReLU_SG.append(LogisticLayer())
# Add output layer
layers_SG.append(LinearLayer(hidden_neurons_2, T_train.shape[1]))
layers_SG.append(SoftmaxOutputLayer())
#DEACTIVATED UNTIL THE DYING ReLU PROBLEM IS FIXED
#layers_ReLU.append(LinearLayer(hidden_neurons_2, T_train.shape[1]))
#layers_ReLU.append(SoftmaxOutputLayer())
layers_ReLU_SG.append(LinearLayer(hidden_neurons_2, T_train.shape[1]))
layers_ReLU_SG.append(SoftmaxOutputLayer())
# +
# Define the forward propagation step as a method.
def forward_step(input_samples, layers):
"""
Compute and return the forward activation of each layer in layers.
Input:
input_samples: A matrix of input samples (each row is an input vector)
layers: A list of Layers
Output:
A list of activations where the activation at each index i+1 corresponds to
the activation of layer i in layers. activations[0] contains the input samples.
"""
activations = [input_samples] # List of layer activations
# Compute the forward activations for each layer starting from the first
X = input_samples
for layer in layers:
Y = layer.get_output(X) # Get the output of the current layer
activations.append(Y) # Store the output for future processing
X = activations[-1] # Set the current input as the activations of the previous layer
return activations # Return the activations of each layer
# Define the backward propagation step as a method
def backward_step(activations, targets, layers):
"""
Perform the backpropagation step over all the layers and return the parameter gradients.
Input:
activations: A list of forward step activations where the activation at
each index i+1 corresponds to the activation of layer i in layers.
activations[0] contains the input samples.
targets: The output targets of the output layer.
layers: A list of Layers corresponding that generated the outputs in activations.
Output:
A list of parameter gradients where the gradients at each index corresponds to
the parameters gradients of the layer at the same index in layers.
"""
param_grads = collections.deque() # List of parameter gradients for each layer
output_grad = None # The error gradient at the output of the current layer
# Propagate the error backwards through all the layers.
# Use reversed to iterate backwards over the list of layers.
for layer in reversed(layers):
Y = activations.pop() # Get the activations of the last layer on the stack
# Compute the error at the output layer.
# The output layer error is calculated different then hidden layer error.
if output_grad is None:
input_grad = layer.get_input_grad(Y, targets)
else: # output_grad is not None (layer is not output layer)
input_grad = layer.get_input_grad(Y, output_grad)
# Get the input of this layer (activations of the previous layer)
X = activations[-1]
# Compute the layer parameter gradients used to update the parameters
grads = layer.get_params_grad(X, output_grad)
param_grads.appendleft(grads)
# Compute gradient at output of previous layer (input of current layer):
output_grad = input_grad
return list(param_grads) # Return the parameter gradients
# +
# Perform gradient checking
nb_samples_gradientcheck = 10 # Test the gradients on a subset of the data
X_temp = X_train[0:nb_samples_gradientcheck,:]
T_temp = T_train[0:nb_samples_gradientcheck,:]
# Get the parameter gradients with backpropagation
activations_SG = forward_step(X_temp, layers_SG)
param_grads_SG = backward_step(activations_SG, T_temp, layers_SG)
#activations_ReLU = forward_step(X_temp, layers_ReLU)
#param_grads_ReLU = backward_step(activations_ReLU, T_temp, layers_ReLU)
#activations_ReLU_SG = forward_step(X_temp, layers_ReLU_SG)
#param_grads_ReLU_SG = backward_step(activations_ReLU_SG, T_temp, layers_ReLU_SG)
# Set the small change to compute the numerical gradient
eps = 0.0001
# Compute the numerical gradients of the parameters in all layers.
for idx in range(len(layers_SG)):
layer_SG = layers_SG[idx]
# layer_ReLU = layers_ReLU[idx]
# layer_ReLU_SG = layers_ReLU_SG[idx]
layer_backprop_grads_SG = param_grads_SG[idx]
# layer_backprop_grads_ReLU = param_grads_ReLU[idx]
# layer_backprop_grads_ReLU_SG = param_grads_ReLU_SG[idx]
# Compute the numerical gradient for each parameter in the SG layer
for p_idx, param in enumerate(layer_SG.get_params_iter()):
grad_backprop = layer_backprop_grads_SG[p_idx]
# + eps
param += eps
plus_cost = layers_SG[-1].get_cost(forward_step(X_temp, layers_SG)[-1], T_temp)
# - eps
param -= 2 * eps
min_cost = layers_SG[-1].get_cost(forward_step(X_temp, layers_SG)[-1], T_temp)
# reset param value
param += eps
# calculate numerical gradient
grad_num = (plus_cost - min_cost)/(2*eps)
# Raise error if the numerical grade is not close to the backprop gradient
if not math.isclose(grad_num, grad_backprop, rel_tol=1e-04, abs_tol=1e-05):
raise ValueError('SG NN: Numerical gradient of {:.6f} is not close to the backpropagation gradient of {:.6f}!'.format(float(grad_num), float(grad_backprop)))
#DEACTIVATED UNTIL THE DYING ReLU PROBLEM IS FIXED
# Compute the numerical gradient for each parameter in the ReLU layer
# for p_idx, param in enumerate(layer_ReLU.get_params_iter()):
# grad_backprop = layer_backprop_grads_ReLU[p_idx]
# # + eps
# param += eps
# plus_cost = layers_ReLU[-1].get_cost(forward_step(X_temp, layers_ReLU)[-1], T_temp)
# # - eps
# param -= 2 * eps
# min_cost = layers_ReLU[-1].get_cost(forward_step(X_temp, layers_ReLU)[-1], T_temp)
# # reset param value
# param += eps
# # calculate numerical gradient
# grad_num = (plus_cost - min_cost)/(2*eps)
# # Raise error if the numerical grade is not close to the backprop gradient
# if not math.isclose(grad_num, grad_backprop, rel_tol=1e-04, abs_tol=1e-05):
# raise ValueError('ReLU NN: Numerical gradient of {:.6f} is not close to the backpropagation gradient of {:.6f}!'.format(float(grad_num), float(grad_backprop)))
# Compute the numerical gradient for each parameter in the ReLU_SG layer
# for p_idx, param in enumerate(layer_ReLU_SG.get_params_iter()):
# grad_backprop = layer_backprop_grads_ReLU_SG[p_idx]
# # + eps
# param += eps
# plus_cost = layers_ReLU_SG[-1].get_cost(forward_step(X_temp, layers_ReLU_SG)[-1], T_temp)
# # - eps
# param -= 2 * eps
# min_cost = layers_ReLU_SG[-1].get_cost(forward_step(X_temp, layers_ReLU_SG)[-1], T_temp)
# # reset param value
# param += eps
# # calculate numerical gradient
# grad_num = (plus_cost - min_cost)/(2*eps)
# # Raise error if the numerical grade is not close to the backprop gradient
# if not math.isclose(grad_num, grad_backprop, rel_tol=1e-03, abs_tol=1e-04):
# raise ValueError('ReLU_SG NN: Numerical gradient of {:.6f} is not close to the backpropagation gradient of {:.6f}!'.format(float(grad_num), float(grad_backprop)))
print('No gradient errors found')
# +
# Create the minibatches
batch_size = 500 # Approximately 500 samples per batch
nb_of_batches = X_train.shape[0] / batch_size # Number of batches
# Create batches (X,Y) from the training set
XT_batches = list(zip(
np.array_split(X_train, nb_of_batches, axis=0), # X samples
np.array_split(T_train, nb_of_batches, axis=0))) # Y targets
# Define a method to update the parameters
def update_params(layers, param_grads, learning_rate):
"""
Function to update the parameters of the given layers with the given gradients
by gradient descent with the given learning rate.
"""
for layer, layer_backprop_grads in zip(layers, param_grads):
for param, grad in zip(layer.get_params_iter(), layer_backprop_grads):
# The parameter returned by the iterator point to the memory space of
# the original layer and can thus be modified inplace.
param -= learning_rate * grad # Update each parameter
def get_accuracy(output_activation, T):
true_pred = 0
for idx, pred in enumerate(output_activation):
pred_digit = np.argmax(pred)
label_digit = np.argmax(T[idx])
if pred_digit == label_digit:
true_pred += 1
accuracy = (true_pred/len(T))*100
return accuracy
def get_certainty(output_activation):
certainty = []
for idx, pred in enumerate(output_activation):
certainty.append((np.amax(pred))*100)
av_certainty = sum(certainty)/float(len(certainty))
return av_certainty, certainty
# +
# Perform backpropagation
max_nb_of_iterations = 300 # Train for a maximum of 300 iterations (Epochs)
learning_rate = 0.1 # Gradient descent learning rate
def train_network(layers):
# initalize some lists to store the cost for future analysis
minibatch_costs = []
training_costs = []
train_accuracy = []
train_certainty = []
validation_costs = []
val_accuracy = []
val_certainty = []
# Train for the maximum number of iterations
start_time = time.time()
for iteration in range(max_nb_of_iterations):
print("Starting Epoch: {}".format(iteration+1))
for X, T in XT_batches: # For each minibatch sub-iteration
activations = forward_step(X, layers) # Get the activations
minibatch_cost = layers[-1].get_cost(activations[-1], T) # Get cost
minibatch_costs.append(minibatch_cost)
param_grads = backward_step(activations, T, layers) # Get the gradients
update_params(layers, param_grads, learning_rate) # Update the parameters
# Get full training cost for future analysis (plots)
activations = forward_step(X_train, layers)
train_accuracy.append(get_accuracy(activations[-1], T_train))
train_certainty.append(get_certainty(activations[-1])[0])
train_cost = layers[-1].get_cost(activations[-1], T_train)
training_costs.append(train_cost)
# Get full validation cost
activations = forward_step(X_validation, layers)
val_accuracy.append(get_accuracy(activations[-1], T_validation))
val_certainty.append(get_certainty(activations[-1])[0])
validation_cost = layers[-1].get_cost(activations[-1], T_validation)
validation_costs.append(validation_cost)
if len(validation_costs) > 3:
# Stop training if the cost on the validation set doesn't decrease
# for 3 iterations
if validation_costs[-1] >= validation_costs[-2] >= validation_costs[-3]:
print("CONVERGENCE ACHIEVED!")
break
end_time = time.time()
t_time = end_time - start_time
nb_of_iterations = iteration + 1 # The number of iterations that have been executed
return layers, nb_of_iterations, minibatch_costs, training_costs, validation_costs, train_accuracy, train_certainty, val_accuracy, val_certainty, t_time
def plot_result(nb_of_iterations, minibatch_costs, training_costs, validation_costs, train_accuracy, train_certainty, val_accuracy, val_certainty, t_time):
# Plot the minibatch, full training set, and validation costs
minibatch_x_inds = np.linspace(0, nb_of_iterations, num=nb_of_iterations*nb_of_batches)
iteration_x_inds = np.linspace(1, nb_of_iterations, num=nb_of_iterations)
# Plot the cost over the iterations
plt.plot(minibatch_x_inds, minibatch_costs, 'k-', linewidth=0.5, label='cost minibatches')
plt.plot(iteration_x_inds, training_costs, 'r-', linewidth=2, label='cost full training set')
plt.plot(iteration_x_inds, validation_costs, 'b-', linewidth=3, label='cost validation set')
# Add labels to the plot
plt.xlabel('iteration')
plt.ylabel('$\\xi$', fontsize=15)
plt.title('Decrease of cost over backprop epochs')
plt.legend()
x1,x2,y1,y2 = plt.axis()
plt.axis((0,nb_of_iterations,0,1.5))
plt.grid()
plt.show()
#Plot accuracy
plt.plot(iteration_x_inds, train_accuracy, 'r-', linewidth=2, label='accuracy full training set')
plt.plot(iteration_x_inds, val_accuracy, 'b-', linewidth=3, label='accuracy validation set')
# Add labels to the plot
plt.xlabel('iteration')
plt.ylabel('Accuracy', fontsize=15)
plt.title('Increase of accuracy over backprop epochs')
plt.legend()
x1,x2,y1,y2 = plt.axis()
plt.axis((0,nb_of_iterations,0,100))
plt.grid()
plt.show()
#Plot certainty
plt.plot(iteration_x_inds, train_certainty, 'r-', linewidth=2, label='certainty full training set')
plt.plot(iteration_x_inds, val_certainty, 'b-', linewidth=3, label='certainty validation set')
# Add labels to the plot
plt.xlabel('iteration')
plt.ylabel('Certainty', fontsize=15)
plt.title('Increase of certainty over backprop epochs')
plt.legend()
x1,x2,y1,y2 = plt.axis()
plt.axis((0,nb_of_iterations,0,100))
plt.grid()
plt.show()
#Print time for training
m, s = divmod(t_time, 60)
h, m = divmod(m, 60)
print("NUMBER OF TRAINING EPOCHS: {}".format(nb_of_iterations))
print("OVERALL TIME FOR TRAINING & VALIDATION: {}h:{}m:{}s".format(h,m,s))
print("LAST THREE VALIDATION COSTS: {} {} {}".format(validation_costs[-3], validation_costs[-2], validation_costs[-1]))
# +
#layers_SG, nb_of_iterations_SG, minibatch_costs_SG, training_costs_SG, validation_costs_SG, train_accuracy_SG, train_certainty_SG, val_accuracy_SG, val_certainty_SG, time_SG = train_network(layers_SG)
layers_ReLU_SG, nb_of_iterations_ReLU_SG, minibatch_costs_ReLU_SG, training_costs_ReLU_SG, validation_costs_ReLU_SG, train_accuracy_ReLU_SG, train_certainty_ReLU_SG, val_accuracy_ReLU_SG, val_certainty_ReLU_SG, time_ReLU_SG = train_network(layers_ReLU_SG)
#plot_result(nb_of_iterations_SG, minibatch_costs_SG, training_costs_SG, validation_costs_SG, train_accuracy_SG, train_certainty_SG, val_accuracy_SG, val_certainty_SG, time_SG)
plot_result(nb_of_iterations_ReLU_SG, minibatch_costs_ReLU_SG, training_costs_ReLU_SG, validation_costs_ReLU_SG, train_accuracy_ReLU_SG, train_certainty_ReLU_SG, val_accuracy_ReLU_SG, val_certainty_ReLU_SG, time_ReLU_SG)
# +
def test_network(layers, T_test):
# Get results of test data
y_true = np.argmax(T_test, axis=1) # Get the target outputs
activations = forward_step(X_test, layers) # Get activation of test samples
y_pred = np.argmax(activations[-1], axis=1) # Get the predictions made by the network
test_accuracy = metrics.accuracy_score(y_true, y_pred) # Test set accuracy
print('The accuracy on the test set is {:.2f}'.format(test_accuracy))
test_accuracy = get_accuracy(activations[-1], T_test)
av_test_certainty, test_certainty = get_certainty(activations[-1])
print("TEST ACCURACY: {}%".format(test_accuracy))
print("AVERAGE TEST CERTAINTY: {}%".format(av_test_certainty))
return y_true, y_pred, test_accuracy, av_test_certainty, test_certainty
def plot_confusion_table(y_true, y_pred):
# Show confusion table
conf_matrix = metrics.confusion_matrix(y_true, y_pred, labels=None) # Get confustion matrix
# Plot the confusion table
class_names = ['${:d}$'.format(x) for x in range(0, 10)] # Digit class names
fig = plt.figure()
ax = fig.add_subplot(111)
# Show class labels on each axis
ax.xaxis.tick_top()
major_ticks = range(0,10)
minor_ticks = [x + 0.5 for x in range(0, 10)]
ax.xaxis.set_ticks(major_ticks, minor=False)
ax.yaxis.set_ticks(major_ticks, minor=False)
ax.xaxis.set_ticks(minor_ticks, minor=True)
ax.yaxis.set_ticks(minor_ticks, minor=True)
ax.xaxis.set_ticklabels(class_names, minor=False, fontsize=15)
ax.yaxis.set_ticklabels(class_names, minor=False, fontsize=15)
# Set plot labels
ax.yaxis.set_label_position("right")
ax.set_xlabel('Predicted label')
ax.set_ylabel('True label')
fig.suptitle('Confusion table', y=1.03, fontsize=15)
# Show a grid to seperate digits
ax.grid(b=True, which=u'minor')
# Color each grid cell according to the number classes predicted
ax.imshow(conf_matrix, interpolation='nearest', cmap='binary')
# Show the number of samples in each cell
for x in range(conf_matrix.shape[0]):
for y in range(conf_matrix.shape[1]):
color = 'w' if x == y else 'k'
ax.text(x, y, conf_matrix[y,x], ha="center", va="center", color=color)
plt.show()
def SHOW_IMAGES(INPUT_FEATURES, LABELS, P_LABELS, CERTAINTY):
print("Would You Like to See Image Prediction?")
USER_I = input()
if (USER_I == 'Yes' or USER_I == 'Y' or USER_I == 'y'):
ROW = 4
COLUMN = 5
n_per_frame = ROW * COLUMN
n_frames = len(INPUT_FEATURES)/(n_per_frame)
for frame in range(int(n_frames)):
for i in range(ROW * COLUMN):
image = INPUT_FEATURES[frame*n_per_frame+i].reshape(28, 28) # not necessary to reshape if ndim is set to 2
plt.subplot(ROW, COLUMN, i+1) # subplot with size (width 3, height 5)
plt.imshow(image, cmap='gray') # cmap='gray' is for black and white picture.
plt.title('P [{}]({}%)'.format(P_LABELS[frame*n_per_frame+i], int(CERTAINTY[frame*n_per_frame+i])))
#plt.set_xlabel('L({})'.format(LABELS[frame*n_per_frame+i]))
plt.axis('off') # do not show axis value
plt.tight_layout() # automatic padding between subplots
#pad=0, w_pad=0, h_pad=2
#plt.savefig('images/mnist_plot.png')
plt.show()
if frame == (n_frames - 1):
print('Showed You All!')
break
print("Would You Like to See Some More?")
USER_I = input()
if (USER_I == 'No' or USER_I == 'N' or USER_I == 'n'):
print("Enough is Enough I Guess!")
break
elif (USER_I == 'No' or USER_I == 'N' or USER_I == 'n'):
print("Well Your Loss!")
else: print("Did not get that, try y/n")
return 1
# +
#labels_output, pred_output_SG, test_accuracy, test_certainty, certainty_vec = test_network(layers_SG, T_test)
labels_output, pred_output_ReLU_SG, test_accuracy, test_certainty, certainty_vec = test_network(layers_ReLU_SG, T_test)
#plot_confusion_table(labels_output, pred_output_SG)
plot_confusion_table(labels_output, pred_output_ReLU_SG)
SHOW_IMAGES(X_test, labels_output, pred_output_ReLU_SG, certainty_vec)
# +
#Write the model into files
w1, b1 = layers_ReLU_SG[0].get_params_array()
w2, b2 = layers_ReLU_SG[2].get_params_array()
w3, b3 = layers_ReLU_SG[4].get_params_array()
with open("ReLU_SG_HL_1_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w1)
with open("ReLU_SG_HL_1_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b1)
with open("ReLU_SG_HL_2_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w2)
with open("ReLU_SG_HL_2_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b2)
with open("ReLU_SG_OL_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w3)
with open("ReLU_SG_OL_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b3)
w1, b1 = layers_SG[0].get_params_array()
w2, b2 = layers_SG[2].get_params_array()
w3, b3 = layers_SG[4].get_params_array()
with open("SG_HL_1_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w1)
with open("SG_HL_1_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b1)
with open("SG_HL_2_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w2)
with open("SG_HL_2_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b2)
with open("SG_OL_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerows(w3)
with open("SG_OL_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator='},\n{')
csvWriter.writerow(b3)
# +
w1, b1 = layers_ReLU_SG[0].get_params_array()
w2, b2 = layers_ReLU_SG[2].get_params_array()
w3, b3 = layers_ReLU_SG[4].get_params_array()
with open("ReLU_SG_HL_1_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerows(w1)
with open("ReLU_SG_HL_1_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerow(b1)
with open("ReLU_SG_HL_2_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerows(w2)
with open("ReLU_SG_HL_2_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerow(b2)
with open("ReLU_SG_OL_W.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerows(w3)
with open("ReLU_SG_OL_b.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerow(b3)
# +
with open("T_TEST_DATA.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerows(T_test)
with open("X_TEST_DATA.csv","w") as my_csv:
csvWriter = csv.writer(my_csv,delimiter=',',lineterminator=',')
csvWriter.writerows(X_test)
# -
| src/3L_NN_MNIST/Python/3L_NN_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from skimage import morphology as skmo
from skimage import io as skio
from skimage.filters import threshold_mean
import numpy as np
from matplotlib import pyplot as plt
dog_path = 'sobaka-400x300.jpeg'
image = skio.imread(dog_path, as_grey=True)
thresh = threshold_mean(image)
binary = image > thresh
plt.imshow(image, cmap=plt.cm.gray)
def skeletonize(img, mask):
skeleton_set = []
while True:
erosion = skmo.erosion(img, mask)
if np.sum(erosion) == 0:
skeleton_set.append(img)
break
dilation = skmo.dilation(erosion, mask)
skeleton_set.append(img ^ (img & dilation))
img = erosion
result = skeleton_set[0]
for skeleton in skeleton_set[1:]:
result |= skeleton
return (result, skeleton_set)
plt.imshow(skeletonize(binary, np.ones((3, 3)))[0], cmap=plt.cm.gray)
def get_histogram(img):
tmpBins = np.linspace(0, 256, 100)
tmpImg = img * 256
plt.hist(tmpImg.reshape(-1), tmpBins)
get_histogram(image)
get_histogram(skeletonize(binary, np.ones((3, 3)))[0])
# # Case 1
# Пройтись по картинке фильтром лапки кота в разных положениях и затем подчиcтить места, где филтр показывает наличие лапки кота.
# # Case 2
# Посчитать градиенты во всех пикселях изображений, а затем подкрасить пиксели с наибольшим градиентом.
# # Case 3
# Нагенерить признаков, запустить knn.
| 6_sem/image_analysis/kr1/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Simple CNN
#
# ## Solving Fashion-MNIST dataset through CNN
#
# In this section, we try to solve the Fashion-MNIST via a CNN and compare it with an MLP network.
# +
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Conv2D, MaxPooling2D
from random import randint
# -
# First, we normalize the input between 0 and 1. This will result in better and lower weights for the out network.
# +
dataset = keras.datasets.fashion_mnist
(x_train, y_train), (x_test, y_test) = dataset.load_data()
x_train = x_train / 255.0
x_test = x_test / 255.0
print "x_train.shape =", x_train.shape
print "x_test.shape =", x_test.shape
# +
# %matplotlib inline
def get_label_name(y):
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
return class_names[y]
rnd_index = randint(0, x_train.shape[0]-1)
plt.imshow(x_train[rnd_index])
plt.title(get_label_name(y_train[rnd_index]))
plt.colorbar()
plt.show()
# -
# ## MLP
#
# Below we have a two-layered MLP network with 128 hidden units. The hidden units have the ReLU activation function. The reason for choosing ReLU the distribution of the input data which are between 0 and 1. The practical tests showed that ReLU works best with images. Another thing to note here is that we have used a 35% dropout connection rate for hidden units which made the performance of the network significantly higher.
# +
model = Sequential()
model.add(Flatten(input_shape=(28, 28)))
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.35))
model.add(Dense(10, activation='softmax'))
print 'Training...'
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(
x_train,
y_train,
epochs=45,
batch_size=128,
# verbose=0,
validation_data=(x_test, y_test)
)
print 'Training Finished!'
train_loss, train_acc = model.evaluate(x_train, y_train, verbose=0)
test_loss, test_acc = model.evaluate(x_test, y_test, verbose=0)
print ''
print 'Train accuracy:', train_acc
print 'Test accuracy:', test_acc
# +
# %matplotlib inline
# Plot training & validation accuracy values
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# ## CNN
#
# Although the MLP worked very good accordingly, the results were not very satisfactory. The accuracy for the MLP as it was showed above was 88% on test which is very low. Thus, we tried to solve the problem with CNN. The network we have used consists of convolutional layers with 32 and 64 units, both have 3x3 kernel size and ReLU activation and both have a 30% dropout rate. Then, these layers are followed by 512 dense hidden units.
# +
x_train_ch = np.reshape(x_train, (x_train.shape[0], 28, 28, 1))
x_test_ch = np.reshape(x_test, (x_test.shape[0], 28, 28, 1))
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=(28, 28, 1)))
model.add(Dropout(0.3))
# model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Conv2D(48, (3, 3), activation='relu'))
# model.add(Dropout(0.3))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(Dropout(0.3))
# model.add(Conv2D(128, (3, 3), activation='relu'))
# model.add(Dropout(0.3))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(10, activation='softmax'))
print 'Training...'
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(
x_train_ch,
y_train,
epochs=40,
batch_size=128,
# verbose=0,
validation_data=(x_test_ch, y_test)
)
print 'Training Finished!'
train_loss, train_acc = model.evaluate(x_train_ch, y_train, verbose=0)
test_loss, test_acc = model.evaluate(x_test_ch, y_test, verbose=0)
print ''
print 'Train accuracy:', train_acc
print 'Test accuracy:', test_acc
# +
# %matplotlib inline
# Plot training & validation accuracy values
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# --------------------------------------
#
# In the next section we solve the CIFAR-10 using CNNs. Please follow up with this [link](https://github.com/ArefMq/simple-nn/blob/master/Day-6.ipynb).
| Day-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize Otter
import otter
grader = otter.Notebook("PS88_lab_week3.ipynb")
# -
# # PS 88 Week 3 Lab: Simulations and Pivotal Voters
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from datascience import Table
from ipywidgets import interact
# %matplotlib inline
# ## Part 1: Plotting expected utility
#
# We can use Python to do expected utility calculations and explore the relationship between parameters in decision models and optimal choices.
#
# In class we showed that the expected utility for voting for a preferred candidate can be written $p_1 b - c$. A nice way to do calculations like this is to first assign values to the variables:
p1=.6
b=100
c=2
p1*b-c
# **Question 1.1. Write code to compute the expected utility to voting when $p_1 = .5$, $b=50$, and $c=.5$ **
#Answer to 1.1 here
p1=.5
b=50
c=.5
p1*b-c
# We don't necessarily care about these expected utilities on their own, but how they compare to the expected utility to abstaining.
#
# **Question 1.2. If $b=50$ and $p_0 = .48$, write code to compute the expected utility to abstaining.**
# Code for 1.2 here
p0=.48
b=50
p0*b
# **Question 1.3. Given 1.1 and 1.2, is voting the expected utility maximizing choice given given these parameters?**
# Answer to 1.3 here
# We can also use the graphic capabilities of Python to learn more about how these models work.
plt.hlines(p0*b, 0,1, label='Abstaining Utility')
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import interact, IntSlider, FloatSlider
def plotEU(b):
plt.hlines(p0*b, 0,2, label='Abstaining Utility')
c = np.arange(0,2, step=.01)
y = p1*b-c
plt.ticklabel_format(style='plain')
plt.xticks(rotation=45)
plt.plot(c,y, label='Voting Expected Utility')
# plt.vlines(no_lobbying+1e7-1e5*p, -2e7, 0, linestyles="dashed")
plt.xlabel('Voting Cost')
plt.ylabel('Expected Utility')
plt.legend()
interact(plotEU, b=IntSlider(min=0,max=300, value=100))
# ## Part 2: Simulating votes
# How can we estimate the probability of a vote mattering? One route is to use probability theory, which in realistic settings (like the electoral college in the US) requires lots of complicated mathematical manipulation. Another way, which will often be faster and uses the tools you are learning in Data 8, is to run simulations.
# As we will see throughout the class, simulation is an incredibly powerful tool that can be used for many purposes. For example, later in the class we will use simulation to see how different causal processes can produce similar data.
#
# For now, we are going to use simulation to simulate the probability a vote matters. The general idea is simple. We will create a large number of "fake electorates" with parameters and randomness that we control, and then see how often an individual vote matters in these simulations.
# Before we get to voting, let's do a simple exercise as warmup. Suppose we want to simulate flipping a coin 10 times. To do this we can use the `random.binomial` function from `numpy` (imported above as `np`). This function takes two arguments: the number of flips (`n`) and the probability that a flip is "heads" (`p`). More generally, we often call $n$ the number of "trials" and $p$ the probability of "success".
#
# The following line of code simulates flipping a "fair" (i.e., $p=.5$) coin 10 times. Run it a few times.
# First number argument is the number of times to flip, the second is the probability of a "heads"
np.random.binomial(n=10, p=.5)
# We can simulate 100 coin flips at a time by changing the `n` argument to 100:
np.random.binomial(n=100, p=.5)
# In the 2020 election, about 158.4 million people voted. This is a big number to have to keep typing, so let's define a variable:
voters2020 = 158400000
# + [markdown] deletable=false editable=false
# **Question 2a. Write a line of code to simulate 158.4 million people flipping a coin and counting how many heads there are.**
# <!--
# BEGIN QUESTION
# name: q2a
# -->
# -
# Code for 2a here
sim = np.random.binomial(n=voters2020, p=.5) #SOLUTION
sim
# + deletable=false editable=false
grader.check("q2a")
# + [markdown] deletable=false editable=false
# Of course, we don't care about coin flipping per se, but we can think about this as the number of "yes" votes if we have n people who vote for a candidate with probability $p$. In the 2020 election, about 51.3% of the voters voted fro <NAME>. Let's do a simulated version of the election: by running `np.random.binomial` with 160 million trials and a probability of "success" of 51.3%.
#
# **Question 2b. Write code for this trial**
# <!--
# BEGIN QUESTION
# name: q2b
# -->
# -
# Code for 2b
joe_count = np.random.binomial(n=voters2020, p=.513) #SOLUTION
joe_count
# + deletable=false editable=false
grader.check("q2b")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# In reality, Biden won 81.3 million votes.
#
# **Question 2c. How close was your answer to the real election? Compare this to the cases where you flipped 10 coins at a time.**
# <!--
# BEGIN QUESTION
# name: q2c
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# **SOLUTION**: Answer to 2c here
# <!-- END QUESTION -->
#
#
#
# ## Part 3. Pivotal votes.
# Suppose that you are a voter in a population with 10 people who are equally likely to vote for candidate A or candidate B, and you prefer candidate A. If you turn out to vote, you will be pivotal if the other 10 are split evenly between the two candidates. How often will this happen?
#
# We can answer this question by running a whole bunch of simulations where we effectively flip 10 coins and count how many heads there are.
#
# The following line runs the code to do 10 coin flips with `p=5` 10,000 times, and stores the results in an array.(Don't worry about the details here: we will cover how to write "loops" like this later.)
# + tags=[]
ntrials=10000
trials10 = [np.random.binomial(n=10, p=.5) for _ in range(ntrials)]
# + [markdown] tags=[]
# Here is the ouput:
# + jupyter={"outputs_hidden": true} tags=[]
trials10
# -
# Let's put these in a table, and then make a histogram to see how often each trial number happens. To make sure we just get a count of how many are at each interval, we need to get the "bins" right.
list(range(11))
# + tags=[]
simtable = Table().with_column("sims10",trials10)
simtable.hist("sims10", bins=range(11))
# -
# Let's see what happens with 20 coin flips. First we create a bunch of simulations:
trials20 = [np.random.binomial(n=20, p=.5) for _ in range(ntrials)]
# And then add the new trials to `simtable` using the `.with_column()` function.
simtable=simtable.with_column("sims20", trials20)
simtable
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 3.1 Make a histogram of the number of heads in the trials with 20 flips. Make sure to set the bins so there each one contains exactly one integer.**
# <!--
# BEGIN QUESTION
# name: q3a
# manual: true
# -->
# -
simtable.hist("sims20", bins=range(21)) # SOLUTION
# <!-- END QUESTION -->
#
#
#
# Let's see what this looks like with a different probability of success. Here is a set of 10 trials with a higher probaility of success ($p = .7$)
trials_high = [np.random.binomial(n=10, p=.7) for _ in range(ntrials)]
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 3.2. Add this array to `simtable`, as a variable called `sims_high`, and create a histogram which shows the frequency of heads in these trials**
# <!--
# BEGIN QUESTION
# name: q3b
# manual: true
# -->
# -
simtable=simtable.with_column("sims_high", trials_high) # SOLUTION
simtable.hist("sims_high", bins=range(11)) # SOLUTION
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
# + tags=[]
simtable
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 3.3. Compare this to the graph where $p=.5$**
# <!--
# BEGIN QUESTION
# name: q3c
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# **SOLUTION**: Answer to 3.3 here
# <!-- END QUESTION -->
#
#
#
# Next we want to figure out exactly how often a voter is pivotal in different situations. To do this, let's create a variable called `pivot10` which is true when there are exactly 5 other voters choosing each candidate.
simtable = simtable.with_column("pivot10", simtable.column("sims10")==5)
simtable
# We can then count the number of trials where a voter was pivotal.
sum(simtable.column("pivot10"))
# Since there were 10,000 trials, we can convert this into a percentage:
sum(simtable.column("pivot10"))/ntrials
# + [markdown] deletable=false editable=false
# **Question 3.4. Write code to determine what proportion of the time a voter is pivotal when $n=20$**
# <!--
# BEGIN QUESTION
# name: q3d
# -->
#
# -
simtable=simtable.with_column("pivot20", simtable.column("sims20")==10) # SOLUTION
pivotal_freq = sum(simtable.column("pivot20"))/ntrials # SOLUTION
pivotal_freq
# + deletable=false editable=false
grader.check("q3d")
# -
# To explore how chaning the size of the electorate and the probabilities of voting affect the probability of being pivotal without having to go through all of these steps, we will define a function which does one simulation and then checks whether a new voter would be pivotal.
def one_pivot(n,p):
return 1*(np.random.binomial(n=n,p=p)==n/2)
# Run this a few times.
one_pivot(n=10, p=.6)
# Let's see how the probability of being pivotal changes with a higher n. To do so, we will use the same looping trick to store 10,000 simulations in an array called piv_trials100. (Note we defined `ntrials=10,000` above)
def pivotal_prob(p):
return sum(one_pivot(n=100, p=.5) for _ in range(ntrials))/ntrials
interact(pivotal_prob, p=(0,1, .1))
# Or a lower p
piv_trials100 = [one_pivot(n=100, p=.4) for _ in range(ntrials)]
sum(piv_trials100)/ntrials
# + [markdown] deletable=false editable=false
# **Question 3.5 Write a line of code to simulate how often a voter will be pivotal in an electorate with 50 voters and $p=.55$**
# <!--
# BEGIN QUESTION
# name: q3e
# -->
#
# -
piv_trials35 = [one_pivot(n=50, p=.55) for _ in range(ntrials)] # SOLUTION
pivotal_freq = sum(piv_trials35)/ntrials # SOLUTION
pivotal_freq
# + deletable=false editable=false
grader.check("q3e")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question 3.6 (Optional) Try running the one_pivot function with an odd number of voters. What happens and why?**
# <!--
# BEGIN QUESTION
# name: q
# manual: true
# -->
#
# -
one_pivot(n=11, p=.5) # SOLUTION
# <!-- END QUESTION -->
#
#
#
# ## Part 4. Pivotal votes with groups
# To learn about situations like the electoral college, let's do a simluation with groups. Imagine there are three groups, who all make a choice by majority vote. The winning candidate is the one who wins a majority vote of the majority of groups, in this case at least two groups.
#
# Questions like this become interesting when the groups vary, maybe in size or in predisposition towards certain candidates. To get started, we will look at an example where all the groups have 50 voters. Group 1 leans against candidate A, group B is split, and group C leans towards group A.
#
# We start by making a table with the number of votes for candidate A in each group. All groups have 50 members, but they have different probabilities of voting for A.
#Group sizes
n1=50
n2=50
n3=50
# Probability of voting for A, by group
p1=.4
p2=.5
p3=.6
np.random.seed(88)
# Creating arrays for simulations for each group
group1 = [np.random.binomial(n=n1, p=p1) for _ in range(ntrials)]
group2 = [np.random.binomial(n=n2, p=p2) for _ in range(ntrials)]
group3 = [np.random.binomial(n=n3, p=p3) for _ in range(ntrials)]
#Putting the arrays into a table
grouptrials = Table().with_columns("votes1",group1,
"votes2", group2,
"votes3",group3)
grouptrials
# Next we create a variable to check whether an individual voter would be pivotal if placed in each group.
grouptrials = grouptrials.with_columns("voter piv1", 1*(grouptrials.column("votes1")==n1/2),
"voter piv2", 1*(grouptrials.column("votes2")==n2/2),
"voter piv3", 1*(grouptrials.column("votes3")==n3/2))
grouptrials
# Let's check how often voters in group 1 are pivotal
sum(grouptrials.column("voter piv1"))/ntrials
# + [markdown] deletable=false editable=false
# **Question 4a. Check how often voters in groups 2 and 3 are pivotal.**
# <!--
# BEGIN QUESTION
# name: q4a
# -->
# -
group2pivotal = sum(grouptrials.column("voter piv2"))/ntrials #SOLUTION
group3pivotal = sum(grouptrials.column("voter piv3"))/ntrials #SOLUTION
group2pivotal, group3pivotal
# + deletable=false editable=false
grader.check("q4a")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# **Question: you should get that two of the groups have a similar probability of being pivotal, but one is different. Which is different any why?**
# <!--
# BEGIN QUESTION
# name: q4b
# manual: true
# -->
# -
# _Type your answer here, replacing this text._
# **SOLUTION**
# <!-- END QUESTION -->
#
#
#
# Now let's check if each group is pivotal, i.e., if the group changing their vote changes which candidate wins the majority of groups. [Note: tricky stuff about ties here is important]
# +
group1piv = 1*((grouptrials.column("votes2") <= n2/2)*(grouptrials.column("votes3") >= n3/2)+
(grouptrials.column("votes2") >= n2/2)*(grouptrials.column("votes3") <= n3/2))
group2piv = 1*((grouptrials.column("votes1") <= n2/2)*(grouptrials.column("votes3") >= n3/2)+
(grouptrials.column("votes1") >= n2/2)*(grouptrials.column("votes3") <= n3/2))
group3piv = 1*((grouptrials.column("votes1") <= n2/2)*(grouptrials.column("votes2") >= n3/2)+
(grouptrials.column("votes1") >= n2/2)*(grouptrials.column("votes2") <= n3/2))
grouptrials = grouptrials.with_columns("group piv1", group1piv,
"group piv2", group2piv,
"group piv3", group3piv)
grouptrials
# + [markdown] deletable=false editable=false
# **How often is each group pivotal?**
# <!--
# BEGIN QUESTION
# name: q4c
# -->
# -
group1_pivotal_rate = sum(grouptrials.column("group piv1"))/ntrials # SOLUTION
group2_pivotal_rate = sum(grouptrials.column("group piv2"))/ntrials # SOLUTION
group3_pivotal_rate = sum(grouptrials.column("group piv3"))/ntrials # SOLUTION
group1_pivotal_rate, group2_pivotal_rate, group3_pivotal_rate
# + deletable=false editable=false
grader.check("q4c")
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# <!--
# BEGIN QUESTION
# name: q4d
# manual: true
# -->
# **Two groups should have similar probabilities, with one group fairly different. Why is the case?**
# -
# _Type your answer here, replacing this text._
# **SOLUTION**: Answer to 2c here
# <!-- END QUESTION -->
#
#
#
# A voter will be pivotal "overall" if they are pivotal within the group and the group is pivotal in the election. We can compute this by multiplying whether a voter is pivotal by whether their group is pivotal: the only voters who will be pivotal (represented by a 1) will have a 1 in both columns.
# +
grouptrials = grouptrials.with_columns("overall piv 1",
grouptrials.column("voter piv1")*grouptrials.column("group piv1"),
"overall piv 2",
grouptrials.column("voter piv2")*grouptrials.column("group piv2"),
"overall piv 3",
grouptrials.column("voter piv3")*grouptrials.column("group piv3"))
grouptrials
# + [markdown] deletable=false editable=false
# **What is the probability of a voter in each group being pivotal?**
# <!--
# BEGIN QUESTION
# name: q4e
# -->
# +
voter_1_pivotal_prob = sum(grouptrials.column("overall piv 1"))/ntrials
voter_2_pivotal_prob = sum(grouptrials.column("overall piv 2"))/ntrials # SOLUTION
voter_3_pivotal_prob = sum(grouptrials.column("overall piv 3"))/ntrials # SOLUTION
voter_1_pivotal_prob, voter_2_pivotal_prob, voter_3_pivotal_prob
# + deletable=false editable=false
grader.check("q4e")
# -
# We can graph the frequency with which a voter in a group is pivotal using `.hist("COLUMNNAME")`. Below, we graph the frequency of a voter in group one being pivotal. You can try graphing the frequency for voters in other groups by changing the column name below.
grouptrials.hist("overall piv 1")
# How frequently do we see the combination of different voter and group pivotal status? In the cells below, we calculate both the absolute frequency as well as percentage of times in which a voter is or is not pivotal within their group as well as their group is or is not. For example, for the cell in which `col_0` and `row_0` equal 0, neither the voter nor the group was pivotal.
pd.crosstab(grouptrials.column("voter piv1"), grouptrials.column("group piv1"))
# This cell mimics the above, except that by normalizing, we see the frequencies as a percentage overall.
pd.crosstab(grouptrials.column("voter piv1"), grouptrials.column("group piv1"), normalize=True)
# Here is a function that ties it all together. It creates a table with group population parameter sizes as well as parameters for the probability of each kind of voter voting for candidate A.
def maketable(n1=50, n2=50, n3=50, p1=.4, p2=.5, p3=.6, ntrials=10000):
group1 = [np.random.binomial(n=n1, p=p1) for _ in range(ntrials)]
group2 = [np.random.binomial(n=n2, p=p2) for _ in range(ntrials)]
group3 = [np.random.binomial(n=n3, p=p3) for _ in range(ntrials)]
grouptrials = Table().with_columns("votes1",group1,
"votes2", group2,
"votes3",group3)
grouptrials = grouptrials.with_columns("voter piv1", 1*(grouptrials.column("votes1")==n1/2),
"voter piv2", 1*(grouptrials.column("votes2")==n2/2),
"voter piv3", 1*(grouptrials.column("votes3")==n3/2))
group1piv = 1*((grouptrials.column("votes2") <= n2/2)*(grouptrials.column("votes3") >= n3/2)+
(grouptrials.column("votes2") >= n2/2)*(grouptrials.column("votes3") <= n3/2))
group2piv = 1*((grouptrials.column("votes1") <= n2/2)*(grouptrials.column("votes3") >= n3/2)+
(grouptrials.column("votes1") >= n2/2)*(grouptrials.column("votes3") <= n3/2))
group3piv = 1*((grouptrials.column("votes1") <= n2/2)*(grouptrials.column("votes2") >= n3/2)+
(grouptrials.column("votes1") >= n2/2)*(grouptrials.column("votes2") <= n3/2))
grouptrials = grouptrials.with_columns("group piv1", group1piv,
"group piv2", group2piv,
"group piv3", group3piv)
grouptrials = grouptrials.with_columns("overall piv1",
grouptrials.column("voter piv1")*grouptrials.column("group piv1"),
"overall piv2",
grouptrials.column("voter piv2")*grouptrials.column("group piv2"),
"overall piv3",
grouptrials.column("voter piv3")*grouptrials.column("group piv3"))
return grouptrials
test = maketable()
test
# What happens as we change the number of voters in each group relative to one another? In the following cell, use the sliders to change the number of voters in each group.
def voter_piv_rate(n1, n2, n3):
sims = maketable(n1, n2, n3)
for i in range(1,4):
print("Voter and Group are Both Pivotal Frequency", sum(sims.column(f"overall piv{i}"))/ntrials)
sims.hist(f"overall piv{i}")
plt.show()
interact(voter_piv_rate, n1=IntSlider(min=0,max=300, value=100), n2=IntSlider(min=0,max=300, value=100), n3=IntSlider(min=0,max=300, value=100))
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# What happens as you change the sliders? Can you make the frequencies the same? How?
# <!--
# BEGIN QUESTION
# name: q4f
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# **SOLUTION**: Answer to 4f here
# <!-- END QUESTION -->
#
#
#
# If we keep the voter populations static, but change their probability of voting for candidate A, what happens?
def voter_piv_rate(p1, p2, p3):
sims = maketable(p1=p1, p2=p2, p3=p3)
for i in range(1,4):
print("Voter and Group are Both Pivotal Frequency", sum(sims.column(f"overall piv{i}"))/ntrials)
sims.hist(f"overall piv{i}")
plt.show()
return sims
interact(voter_piv_rate, p1=FloatSlider(min=0,max=1, value=.4, step=.1), p2=FloatSlider(min=0,max=1, value=.5, step=.1), p3=FloatSlider(min=0,max=1, value=.6, step=.1))
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# What happens as you change the sliders? Can you make the frequencies the same? How?
# <!--
# BEGIN QUESTION
# name: q4g
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# **SOLUTION**: Answer to 4g here
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
# -
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
# + [markdown] deletable=false editable=false
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
#
# These are some submission instructions.
# + deletable=false editable=false
# Save your notebook first, then run this cell to export your submission.
grader.export()
# -
#
| lab/lab3/dist/autograder/PS88_lab_week3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
Image(filename='images/aiayn.png')
# The Transformer from ["Attention is All You Need"](https://arxiv.org/abs/1706.03762) has been on a lot of people's minds over the last year. Besides producing major improvements in translation quality, it provides a new architecture for many other NLP tasks. The paper itself is very clearly written, but the conventional wisdom has been that it is quite difficult to implement correctly.
#
# In this post I present an "annotated" version of the paper in the form of a line-by-line implementation. I have reordered and deleted some sections from the original paper and added comments throughout. This document itself is a working notebook, and should be a completely usable implementation. In total there are 400 lines of library code which can process 27,000 tokens per second on 4 GPUs.
#
# To follow along you will first need to install [PyTorch](http://pytorch.org/). The complete notebook is also available on [github](https://github.com/harvardnlp/annotated-transformer) or on Google [Colab](https://drive.google.com/file/d/1xQXSv6mtAOLXxEMi8RvaW8TW-7bvYBDF/view?usp=sharing).
#
# Note this is merely a starting point for researchers and interested developers. The code here is based heavily on our [OpenNMT](http://opennmt.net) packages. (If helpful feel free to [cite](#conclusion).) For other full-sevice implementations of the model check-out [Tensor2Tensor](https://github.com/tensorflow/tensor2tensor) (tensorflow) and [Sockeye](https://github.com/awslabs/sockeye) (mxnet).
#
# - <NAME> ([@harvardnlp](https://twitter.com/harvardnlp) or <EMAIL>)
#
# # Prelims
# +
# # !pip install http://download.pytorch.org/whl/cu80/torch-0.3.0.post4-cp36-cp36m-linux_x86_64.whl numpy matplotlib spacy torchtext seaborn
# -
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import math, copy, time
from torch.autograd import Variable
import matplotlib.pyplot as plt
import seaborn
seaborn.set_context(context="talk")
# %matplotlib inline
# Table of Contents
#
#
# * Table of Contents
# {:toc}
# > My comments are blockquoted. The main text is all from the paper itself.
# # Background
# The goal of reducing sequential computation also forms the foundation of the Extended Neural GPU, ByteNet and ConvS2S, all of which use convolutional neural networks as basic building block, computing hidden representations in parallel for all input and output positions. In these models, the number of operations required to relate signals from two arbitrary input or output positions grows in the distance between positions, linearly for ConvS2S and logarithmically for ByteNet. This makes it more difficult to learn dependencies between distant positions. In the Transformer this is reduced to a constant number of operations, albeit at the cost of reduced effective resolution due to averaging attention-weighted positions, an effect we counteract with Multi-Head Attention.
#
# Self-attention, sometimes called intra-attention is an attention mechanism relating different positions of a single sequence in order to compute a representation of the sequence. Self-attention has been used successfully in a variety of tasks including reading comprehension, abstractive summarization, textual entailment and learning task-independent sentence representations. End-to-end memory networks are based on a recurrent attention mechanism instead of sequencealigned recurrence and have been shown to perform well on simple-language question answering and
# language modeling tasks.
#
# To the best of our knowledge, however, the Transformer is the first transduction model relying entirely on self-attention to compute representations of its input and output without using sequence aligned RNNs or convolution.
# # Model Architecture
# Most competitive neural sequence transduction models have an encoder-decoder structure [(cite)](https://arxiv.org/abs/1409.0473). Here, the encoder maps an input sequence of symbol representations $(x_1, ..., x_n)$ to a sequence of continuous representations $\mathbf{z} = (z_1, ..., z_n)$. Given $\mathbf{z}$, the decoder then generates an output sequence $(y_1,...,y_m)$ of symbols one element at a time. At each step the model is auto-regressive [(cite)](https://arxiv.org/abs/1308.0850), consuming the previously generated symbols as additional input when generating the next.
class EncoderDecoder(nn.Module):
"""
A standard Encoder-Decoder architecture. Base for this and many
other models.
"""
def __init__(self, encoder, decoder, src_embed, tgt_embed, generator):
super(EncoderDecoder, self).__init__()
self.encoder = encoder
self.decoder = decoder
self.src_embed = src_embed
self.tgt_embed = tgt_embed
self.generator = generator
def forward(self, src, tgt, src_mask, tgt_mask):
"Take in and process masked src and target sequences."
return self.decode(self.encode(src, src_mask), src_mask,
tgt, tgt_mask)
def encode(self, src, src_mask):
return self.encoder(self.src_embed(src), src_mask)
def decode(self, memory, src_mask, tgt, tgt_mask):
return self.decoder(self.tgt_embed(tgt), memory, src_mask, tgt_mask)
class Generator(nn.Module):
"Define standard linear + softmax generation step."
def __init__(self, d_model, vocab):
super(Generator, self).__init__()
self.proj = nn.Linear(d_model, vocab)
def forward(self, x):
return F.log_softmax(self.proj(x), dim=-1)
# The Transformer follows this overall architecture using stacked self-attention and point-wise, fully connected layers for both the encoder and decoder, shown in the left and right halves of Figure 1, respectively.
Image(filename='images/ModalNet-21.png')
# ## Encoder and Decoder Stacks
#
# ### Encoder
#
# The encoder is composed of a stack of $N=6$ identical layers.
def clones(module, N):
"Produce N identical layers."
return nn.ModuleList([copy.deepcopy(module) for _ in range(N)])
class Encoder(nn.Module):
"Core encoder is a stack of N layers"
def __init__(self, layer, N):
super(Encoder, self).__init__()
self.layers = clones(layer, N)
self.norm = LayerNorm(layer.size)
def forward(self, x, mask):
"Pass the input (and mask) through each layer in turn."
for layer in self.layers:
x = layer(x, mask)
return self.norm(x)
# We employ a residual connection [(cite)](https://arxiv.org/abs/1512.03385) around each of the two sub-layers, followed by layer normalization [(cite)](https://arxiv.org/abs/1607.06450).
class LayerNorm(nn.Module):
"Construct a layernorm module (See citation for details)."
def __init__(self, features, eps=1e-6):
super(LayerNorm, self).__init__()
self.a_2 = nn.Parameter(torch.ones(features))
self.b_2 = nn.Parameter(torch.zeros(features))
self.eps = eps
def forward(self, x):
mean = x.mean(-1, keepdim=True)
std = x.std(-1, keepdim=True)
return self.a_2 * (x - mean) / (std + self.eps) + self.b_2
# That is, the output of each sub-layer is $\mathrm{LayerNorm}(x + \mathrm{Sublayer}(x))$, where $\mathrm{Sublayer}(x)$ is the function implemented by the sub-layer itself. We apply dropout [(cite)](http://jmlr.org/papers/v15/srivastava14a.html) to the output of each sub-layer, before it is added to the sub-layer input and normalized.
#
# To facilitate these residual connections, all sub-layers in the model, as well as the embedding layers, produce outputs of dimension $d_{\text{model}}=512$.
class SublayerConnection(nn.Module):
"""
A residual connection followed by a layer norm.
Note for code simplicity the norm is first as opposed to last.
"""
def __init__(self, size, dropout):
super(SublayerConnection, self).__init__()
self.norm = LayerNorm(size)
self.dropout = nn.Dropout(dropout)
def forward(self, x, sublayer):
"Apply residual connection to any sublayer with the same size."
return x + self.dropout(sublayer(self.norm(x)))
# Each layer has two sub-layers. The first is a multi-head self-attention mechanism, and the second is a simple, position-wise fully connected feed-forward network.
class EncoderLayer(nn.Module):
"Encoder is made up of self-attn and feed forward (defined below)"
def __init__(self, size, self_attn, feed_forward, dropout):
super(EncoderLayer, self).__init__()
self.self_attn = self_attn
self.feed_forward = feed_forward
self.sublayer = clones(SublayerConnection(size, dropout), 2)
self.size = size
def forward(self, x, mask):
"Follow Figure 1 (left) for connections."
x = self.sublayer[0](x, lambda x: self.self_attn(x, x, x, mask))
return self.sublayer[1](x, self.feed_forward)
# ### Decoder
#
# The decoder is also composed of a stack of $N=6$ identical layers.
#
class Decoder(nn.Module):
"Generic N layer decoder with masking."
def __init__(self, layer, N):
super(Decoder, self).__init__()
self.layers = clones(layer, N)
self.norm = LayerNorm(layer.size)
def forward(self, x, memory, src_mask, tgt_mask):
for layer in self.layers:
x = layer(x, memory, src_mask, tgt_mask)
return self.norm(x)
# In addition to the two sub-layers in each encoder layer, the decoder inserts a third sub-layer, which performs multi-head attention over the output of the encoder stack. Similar to the encoder, we employ residual connections around each of the sub-layers, followed by layer normalization.
class DecoderLayer(nn.Module):
"Decoder is made of self-attn, src-attn, and feed forward (defined below)"
def __init__(self, size, self_attn, src_attn, feed_forward, dropout):
super(DecoderLayer, self).__init__()
self.size = size
self.self_attn = self_attn
self.src_attn = src_attn
self.feed_forward = feed_forward
self.sublayer = clones(SublayerConnection(size, dropout), 3)
def forward(self, x, memory, src_mask, tgt_mask):
"Follow Figure 1 (right) for connections."
m = memory
x = self.sublayer[0](x, lambda x: self.self_attn(x, x, x, tgt_mask))
x = self.sublayer[1](x, lambda x: self.src_attn(x, m, m, src_mask))
return self.sublayer[2](x, self.feed_forward)
# We also modify the self-attention sub-layer in the decoder stack to prevent positions from attending to subsequent positions. This masking, combined with fact that the output embeddings are offset by one position, ensures that the predictions for position $i$ can depend only on the known outputs at positions less than $i$.
def subsequent_mask(size):
"Mask out subsequent positions."
attn_shape = (1, size, size)
subsequent_mask = np.triu(np.ones(attn_shape), k=1).astype('uint8')
return torch.from_numpy(subsequent_mask) == 0
# > Below the attention mask shows the position each tgt word (row) is allowed to look at (column). Words are blocked for attending to future words during training.
# +
plt.figure(figsize=(5,5))
plt.imshow(subsequent_mask(20)[0])
None
# -
# ### Attention
# An attention function can be described as mapping a query and a set of key-value pairs to an output, where the query, keys, values, and output are all vectors. The output is computed as a weighted sum of the values, where the weight assigned to each value is computed by a compatibility function of the query with the corresponding key.
#
# We call our particular attention "Scaled Dot-Product Attention". The input consists of queries and keys of dimension $d_k$, and values of dimension $d_v$. We compute the dot products of the query with all keys, divide each by $\sqrt{d_k}$, and apply a softmax function to obtain the weights on the values.
#
Image(filename='images/ModalNet-19.png')
#
# In practice, we compute the attention function on a set of queries simultaneously, packed together into a matrix $Q$. The keys and values are also packed together into matrices $K$ and $V$. We compute the matrix of outputs as:
#
# $$
# \mathrm{Attention}(Q, K, V) = \mathrm{softmax}(\frac{QK^T}{\sqrt{d_k}})V
# $$
def attention(query, key, value, mask=None, dropout=None):
"Compute 'Scaled Dot Product Attention'"
d_k = query.size(-1)
scores = torch.matmul(query, key.transpose(-2, -1)) \
/ math.sqrt(d_k)
if mask is not None:
scores = scores.masked_fill(mask == 0, -1e9)
p_attn = F.softmax(scores, dim = -1)
if dropout is not None:
p_attn = dropout(p_attn)
return torch.matmul(p_attn, value), p_attn
# The two most commonly used attention functions are additive attention [(cite)](https://arxiv.org/abs/1409.0473), and dot-product (multiplicative) attention. Dot-product attention is identical to our algorithm, except for the scaling factor of $\frac{1}{\sqrt{d_k}}$. Additive attention computes the compatibility function using a feed-forward network with a single hidden layer. While the two are similar in theoretical complexity, dot-product attention is much faster and more space-efficient in practice, since it can be implemented using highly optimized matrix multiplication code.
#
#
# While for small values of $d_k$ the two mechanisms perform similarly, additive attention outperforms dot product attention without scaling for larger values of $d_k$ [(cite)](https://arxiv.org/abs/1703.03906). We suspect that for large values of $d_k$, the dot products grow large in magnitude, pushing the softmax function into regions where it has extremely small gradients (To illustrate why the dot products get large, assume that the components of $q$ and $k$ are independent random variables with mean $0$ and variance $1$. Then their dot product, $q \cdot k = \sum_{i=1}^{d_k} q_ik_i$, has mean $0$ and variance $d_k$.). To counteract this effect, we scale the dot products by $\frac{1}{\sqrt{d_k}}$.
#
Image(filename='images/ModalNet-20.png')
# Multi-head attention allows the model to jointly attend to information from different representation subspaces at different positions. With a single attention head, averaging inhibits this.
# $$
# \mathrm{MultiHead}(Q, K, V) = \mathrm{Concat}(\mathrm{head_1}, ..., \mathrm{head_h})W^O \\
# \text{where}~\mathrm{head_i} = \mathrm{Attention}(QW^Q_i, KW^K_i, VW^V_i)
# $$
#
# Where the projections are parameter matrices $W^Q_i \in \mathbb{R}^{d_{\text{model}} \times d_k}$, $W^K_i \in \mathbb{R}^{d_{\text{model}} \times d_k}$, $W^V_i \in \mathbb{R}^{d_{\text{model}} \times d_v}$ and $W^O \in \mathbb{R}^{hd_v \times d_{\text{model}}}$. In this work we employ $h=8$ parallel attention layers, or heads. For each of these we use $d_k=d_v=d_{\text{model}}/h=64$. Due to the reduced dimension of each head, the total computational cost is similar to that of single-head attention with full dimensionality.
class MultiHeadedAttention(nn.Module):
def __init__(self, h, d_model, dropout=0.1):
"Take in model size and number of heads."
super(MultiHeadedAttention, self).__init__()
assert d_model % h == 0
# We assume d_v always equals d_k
self.d_k = d_model // h
self.h = h
self.linears = clones(nn.Linear(d_model, d_model), 4)
self.attn = None
self.dropout = nn.Dropout(p=dropout)
def forward(self, query, key, value, mask=None):
"Implements Figure 2"
if mask is not None:
# Same mask applied to all h heads.
mask = mask.unsqueeze(1)
nbatches = query.size(0)
# 1) Do all the linear projections in batch from d_model => h x d_k
query, key, value = \
[l(x).view(nbatches, -1, self.h, self.d_k).transpose(1, 2)
for l, x in zip(self.linears, (query, key, value))]
# 2) Apply attention on all the projected vectors in batch.
x, self.attn = attention(query, key, value, mask=mask,
dropout=self.dropout)
# 3) "Concat" using a view and apply a final linear.
x = x.transpose(1, 2).contiguous() \
.view(nbatches, -1, self.h * self.d_k)
return self.linears[-1](x)
# ### Applications of Attention in our Model
# The Transformer uses multi-head attention in three different ways:
# 1) In "encoder-decoder attention" layers, the queries come from the previous decoder layer, and the memory keys and values come from the output of the encoder. This allows every position in the decoder to attend over all positions in the input sequence. This mimics the typical encoder-decoder attention mechanisms in sequence-to-sequence models such as [(cite)](https://arxiv.org/abs/1609.08144).
#
#
# 2) The encoder contains self-attention layers. In a self-attention layer all of the keys, values and queries come from the same place, in this case, the output of the previous layer in the encoder. Each position in the encoder can attend to all positions in the previous layer of the encoder.
#
#
# 3) Similarly, self-attention layers in the decoder allow each position in the decoder to attend to all positions in the decoder up to and including that position. We need to prevent leftward information flow in the decoder to preserve the auto-regressive property. We implement this inside of scaled dot-product attention by masking out (setting to $-\infty$) all values in the input of the softmax which correspond to illegal connections.
# ## Position-wise Feed-Forward Networks
#
# In addition to attention sub-layers, each of the layers in our encoder and decoder contains a fully connected feed-forward network, which is applied to each position separately and identically. This consists of two linear transformations with a ReLU activation in between.
#
# $$\mathrm{FFN}(x)=\max(0, xW_1 + b_1) W_2 + b_2$$
#
# While the linear transformations are the same across different positions, they use different parameters from layer to layer. Another way of describing this is as two convolutions with kernel size 1. The dimensionality of input and output is $d_{\text{model}}=512$, and the inner-layer has dimensionality $d_{ff}=2048$.
class PositionwiseFeedForward(nn.Module):
"Implements FFN equation."
def __init__(self, d_model, d_ff, dropout=0.1):
super(PositionwiseFeedForward, self).__init__()
self.w_1 = nn.Linear(d_model, d_ff)
self.w_2 = nn.Linear(d_ff, d_model)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
return self.w_2(self.dropout(F.relu(self.w_1(x))))
# ## Embeddings and Softmax
# Similarly to other sequence transduction models, we use learned embeddings to convert the input tokens and output tokens to vectors of dimension $d_{\text{model}}$. We also use the usual learned linear transformation and softmax function to convert the decoder output to predicted next-token probabilities. In our model, we share the same weight matrix between the two embedding layers and the pre-softmax linear transformation, similar to [(cite)](https://arxiv.org/abs/1608.05859). In the embedding layers, we multiply those weights by $\sqrt{d_{\text{model}}}$.
class Embeddings(nn.Module):
def __init__(self, d_model, vocab):
super(Embeddings, self).__init__()
self.lut = nn.Embedding(vocab, d_model)
self.d_model = d_model
def forward(self, x):
return self.lut(x) * math.sqrt(self.d_model)
# ## Positional Encoding
# Since our model contains no recurrence and no convolution, in order for the model to make use of the order of the sequence, we must inject some information about the relative or absolute position of the tokens in the sequence. To this end, we add "positional encodings" to the input embeddings at the bottoms of the encoder and decoder stacks. The positional encodings have the same dimension $d_{\text{model}}$ as the embeddings, so that the two can be summed. There are many choices of positional encodings, learned and fixed [(cite)](https://arxiv.org/pdf/1705.03122.pdf).
#
# In this work, we use sine and cosine functions of different frequencies:
# $$PE_{(pos,2i)} = sin(pos / 10000^{2i/d_{\text{model}}})$$
#
# $$PE_{(pos,2i+1)} = cos(pos / 10000^{2i/d_{\text{model}}})$$
# where $pos$ is the position and $i$ is the dimension. That is, each dimension of the positional encoding corresponds to a sinusoid. The wavelengths form a geometric progression from $2\pi$ to $10000 \cdot 2\pi$. We chose this function because we hypothesized it would allow the model to easily learn to attend by relative positions, since for any fixed offset $k$, $PE_{pos+k}$ can be represented as a linear function of $PE_{pos}$.
#
# In addition, we apply dropout to the sums of the embeddings and the positional encodings in both the encoder and decoder stacks. For the base model, we use a rate of $P_{drop}=0.1$.
#
#
class PositionalEncoding(nn.Module):
"Implement the PE function."
def __init__(self, d_model, dropout, max_len=5000):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(p=dropout)
# Compute the positional encodings once in log space.
pe = torch.zeros(max_len, d_model)
position = torch.arange(0, max_len).unsqueeze(1)
div_term = torch.exp(torch.arange(0, d_model, 2) *
-(math.log(10000.0) / d_model))
pe[:, 0::2] = torch.sin(position * div_term)
pe[:, 1::2] = torch.cos(position * div_term)
pe = pe.unsqueeze(0)
self.register_buffer('pe', pe)
def forward(self, x):
x = x + Variable(self.pe[:, :x.size(1)],
requires_grad=False)
return self.dropout(x)
# > Below the positional encoding will add in a sine wave based on position. The frequency and offset of the wave is different for each dimension.
plt.figure(figsize=(15, 5))
pe = PositionalEncoding(20, 0)
y = pe.forward(Variable(torch.zeros(1, 100, 20)))
plt.plot(np.arange(100), y[0, :, 4:8].data.numpy())
plt.legend(["dim %d"%p for p in [4,5,6,7]])
None
# We also experimented with using learned positional embeddings [(cite)](https://arxiv.org/pdf/1705.03122.pdf) instead, and found that the two versions produced nearly identical results. We chose the sinusoidal version because it may allow the model to extrapolate to sequence lengths longer than the ones encountered during training.
# ## Full Model
#
# > Here we define a function from hyperparameters to a full model.
def make_model(src_vocab, tgt_vocab, N=6,
d_model=512, d_ff=2048, h=8, dropout=0.1):
"Helper: Construct a model from hyperparameters."
c = copy.deepcopy
attn = MultiHeadedAttention(h, d_model)
ff = PositionwiseFeedForward(d_model, d_ff, dropout)
position = PositionalEncoding(d_model, dropout)
model = EncoderDecoder(
Encoder(EncoderLayer(d_model, c(attn), c(ff), dropout), N),
Decoder(DecoderLayer(d_model, c(attn), c(attn),
c(ff), dropout), N),
nn.Sequential(Embeddings(d_model, src_vocab), c(position)),
nn.Sequential(Embeddings(d_model, tgt_vocab), c(position)),
Generator(d_model, tgt_vocab))
# This was important from their code.
# Initialize parameters with Glorot / fan_avg.
for p in model.parameters():
if p.dim() > 1:
nn.init.xavier_uniform(p)
return model
# Small example model.
tmp_model = make_model(10, 10, 2)
None
# # Training
#
# This section describes the training regime for our models.
# > We stop for a quick interlude to introduce some of the tools
# needed to train a standard encoder decoder model. First we define a batch object that holds the src and target sentences for training, as well as constructing the masks.
# ## Batches and Masking
class Batch:
"Object for holding a batch of data with mask during training."
def __init__(self, src, trg=None, pad=0):
self.src = src
self.src_mask = (src != pad).unsqueeze(-2)
if trg is not None:
self.trg = trg[:, :-1]
self.trg_y = trg[:, 1:]
self.trg_mask = \
self.make_std_mask(self.trg, pad)
self.ntokens = (self.trg_y != pad).data.sum()
@staticmethod
def make_std_mask(tgt, pad):
"Create a mask to hide padding and future words."
tgt_mask = (tgt != pad).unsqueeze(-2)
tgt_mask = tgt_mask & Variable(
subsequent_mask(tgt.size(-1)).type_as(tgt_mask.data))
return tgt_mask
# > Next we create a generic training and scoring function to keep track of loss. We pass in a generic loss compute function that also handles parameter updates.
# ## Training Loop
def run_epoch(data_iter, model, loss_compute):
"Standard Training and Logging Function"
start = time.time()
total_tokens = 0
total_loss = 0
tokens = 0
for i, batch in enumerate(data_iter):
out = model.forward(batch.src, batch.trg,
batch.src_mask, batch.trg_mask)
loss = loss_compute(out, batch.trg_y, batch.ntokens)
total_loss += loss
total_tokens += batch.ntokens
tokens += batch.ntokens
if i % 50 == 1:
elapsed = time.time() - start
print("Epoch Step: %d Loss: %f Tokens per Sec: %f" %
(i, loss / batch.ntokens, tokens / elapsed))
start = time.time()
tokens = 0
return total_loss / total_tokens
# ## Training Data and Batching
#
# We trained on the standard WMT 2014 English-German dataset consisting of about 4.5 million sentence pairs. Sentences were encoded using byte-pair encoding, which has a shared source-target vocabulary of about 37000 tokens. For English-French, we used the significantly larger WMT 2014 English-French dataset consisting of 36M sentences and split tokens into a 32000 word-piece vocabulary.
#
#
# Sentence pairs were batched together by approximate sequence length. Each training batch contained a set of sentence pairs containing approximately 25000 source tokens and 25000 target tokens.
# > We will use torch text for batching. This is discussed in more detail below. Here we create batches in a torchtext function that ensures our batch size padded to the maximum batchsize does not surpass a threshold (25000 if we have 8 gpus).
global max_src_in_batch, max_tgt_in_batch
def batch_size_fn(new, count, sofar):
"Keep augmenting batch and calculate total number of tokens + padding."
global max_src_in_batch, max_tgt_in_batch
if count == 1:
max_src_in_batch = 0
max_tgt_in_batch = 0
max_src_in_batch = max(max_src_in_batch, len(new.src))
max_tgt_in_batch = max(max_tgt_in_batch, len(new.trg) + 2)
src_elements = count * max_src_in_batch
tgt_elements = count * max_tgt_in_batch
return max(src_elements, tgt_elements)
# ## Hardware and Schedule
# We trained our models on one machine with 8 NVIDIA P100 GPUs. For our base models using the hyperparameters described throughout the paper, each training step took about 0.4 seconds. We trained the base models for a total of 100,000 steps or 12 hours. For our big models, step time was 1.0 seconds. The big models were trained for 300,000 steps (3.5 days).
# ## Optimizer
#
# We used the Adam optimizer [(cite)](https://arxiv.org/abs/1412.6980) with $\beta_1=0.9$, $\beta_2=0.98$ and $\epsilon=10^{-9}$. We varied the learning rate over the course of training, according to the formula:
# $$
# lrate = d_{\text{model}}^{-0.5} \cdot
# \min({step\_num}^{-0.5},
# {step\_num} \cdot {warmup\_steps}^{-1.5})
# $$
# This corresponds to increasing the learning rate linearly for the first $warmup\_steps$ training steps, and decreasing it thereafter proportionally to the inverse square root of the step number. We used $warmup\_steps=4000$.
# > Note: This part is very important. Need to train with this setup of the model.
# +
class NoamOpt:
"Optim wrapper that implements rate."
def __init__(self, model_size, factor, warmup, optimizer):
self.optimizer = optimizer
self._step = 0
self.warmup = warmup
self.factor = factor
self.model_size = model_size
self._rate = 0
def step(self):
"Update parameters and rate"
self._step += 1
rate = self.rate()
for p in self.optimizer.param_groups:
p['lr'] = rate
self._rate = rate
self.optimizer.step()
def rate(self, step = None):
"Implement `lrate` above"
if step is None:
step = self._step
return self.factor * \
(self.model_size ** (-0.5) *
min(step ** (-0.5), step * self.warmup ** (-1.5)))
def get_std_opt(model):
return NoamOpt(model.src_embed[0].d_model, 2, 4000,
torch.optim.Adam(model.parameters(), lr=0, betas=(0.9, 0.98), eps=1e-9))
# -
#
# > Example of the curves of this model for different model sizes and for optimization hyperparameters.
# Three settings of the lrate hyperparameters.
opts = [NoamOpt(512, 1, 4000, None),
NoamOpt(512, 1, 8000, None),
NoamOpt(256, 1, 4000, None)]
plt.plot(np.arange(1, 20000), [[opt.rate(i) for opt in opts] for i in range(1, 20000)])
plt.legend(["512:4000", "512:8000", "256:4000"])
None
# ## Regularization
#
# ### Label Smoothing
#
# During training, we employed label smoothing of value $\epsilon_{ls}=0.1$ [(cite)](https://arxiv.org/abs/1512.00567). This hurts perplexity, as the model learns to be more unsure, but improves accuracy and BLEU score.
# > We implement label smoothing using the KL div loss. Instead of using a one-hot target distribution, we create a distribution that has `confidence` of the correct word and the rest of the `smoothing` mass distributed throughout the vocabulary.
class LabelSmoothing(nn.Module):
"Implement label smoothing."
def __init__(self, size, padding_idx, smoothing=0.0):
super(LabelSmoothing, self).__init__()
self.criterion = nn.KLDivLoss(size_average=False)
self.padding_idx = padding_idx
self.confidence = 1.0 - smoothing
self.smoothing = smoothing
self.size = size
self.true_dist = None
def forward(self, x, target):
assert x.size(1) == self.size
true_dist = x.data.clone()
true_dist.fill_(self.smoothing / (self.size - 2))
true_dist.scatter_(1, target.data.unsqueeze(1), self.confidence)
true_dist[:, self.padding_idx] = 0
mask = torch.nonzero(target.data == self.padding_idx)
if mask.dim() > 0:
true_dist.index_fill_(0, mask.squeeze(), 0.0)
self.true_dist = true_dist
return self.criterion(x, Variable(true_dist, requires_grad=False))
# > Here we can see an example of how the mass is distributed to the words based on confidence.
# +
#Example of label smoothing.
crit = LabelSmoothing(5, 0, 0.4)
predict = torch.FloatTensor([[0, 0.2, 0.7, 0.1, 0],
[0, 0.2, 0.7, 0.1, 0],
[0, 0.2, 0.7, 0.1, 0]])
v = crit(Variable(predict.log()),
Variable(torch.LongTensor([2, 1, 0])))
# Show the target distributions expected by the system.
plt.imshow(crit.true_dist)
None
# -
# > Label smoothing actually starts to penalize the model if it gets very confident about a given choice.
crit = LabelSmoothing(5, 0, 0.1)
def loss(x):
d = x + 3 * 1
predict = torch.FloatTensor([[0, x / d, 1 / d, 1 / d, 1 / d],
])
#print(predict)
return crit(Variable(predict.log()),
Variable(torch.LongTensor([1]))).data[0]
plt.plot(np.arange(1, 100), [loss(x) for x in range(1, 100)])
None
# # A First Example
#
# > We can begin by trying out a simple copy-task. Given a random set of input symbols from a small vocabulary, the goal is to generate back those same symbols.
# ## Synthetic Data
def data_gen(V, batch, nbatches):
"Generate random data for a src-tgt copy task."
for i in range(nbatches):
data = torch.from_numpy(np.random.randint(1, V, size=(batch, 10)))
data[:, 0] = 1
src = Variable(data, requires_grad=False)
tgt = Variable(data, requires_grad=False)
yield Batch(src, tgt, 0)
# ## Loss Computation
class SimpleLossCompute:
"A simple loss compute and train function."
def __init__(self, generator, criterion, opt=None):
self.generator = generator
self.criterion = criterion
self.opt = opt
def __call__(self, x, y, norm):
x = self.generator(x)
loss = self.criterion(x.contiguous().view(-1, x.size(-1)),
y.contiguous().view(-1)) / norm
loss.backward()
if self.opt is not None:
self.opt.step()
self.opt.optimizer.zero_grad()
return loss.data[0] * norm
# ## Greedy Decoding
# +
# Train the simple copy task.
V = 11
criterion = LabelSmoothing(size=V, padding_idx=0, smoothing=0.0)
model = make_model(V, V, N=2)
model_opt = NoamOpt(model.src_embed[0].d_model, 1, 400,
torch.optim.Adam(model.parameters(), lr=0, betas=(0.9, 0.98), eps=1e-9))
for epoch in range(10):
model.train()
run_epoch(data_gen(V, 30, 20), model,
SimpleLossCompute(model.generator, criterion, model_opt))
model.eval()
print(run_epoch(data_gen(V, 30, 5), model,
SimpleLossCompute(model.generator, criterion, None)))
# -
# > This code predicts a translation using greedy decoding for simplicity.
# +
def greedy_decode(model, src, src_mask, max_len, start_symbol):
memory = model.encode(src, src_mask)
ys = torch.ones(1, 1).fill_(start_symbol).type_as(src.data)
for i in range(max_len-1):
out = model.decode(memory, src_mask,
Variable(ys),
Variable(subsequent_mask(ys.size(1))
.type_as(src.data)))
prob = model.generator(out[:, -1])
_, next_word = torch.max(prob, dim = 1)
next_word = next_word.data[0]
ys = torch.cat([ys,
torch.ones(1, 1).type_as(src.data).fill_(next_word)], dim=1)
return ys
model.eval()
src = Variable(torch.LongTensor([[1,2,3,4,5,6,7,8,9,10]]) )
src_mask = Variable(torch.ones(1, 1, 10) )
print(greedy_decode(model, src, src_mask, max_len=10, start_symbol=1))
# -
# # A Real World Example
#
# > Now we consider a real-world example using the IWSLT German-English Translation task. This task is much smaller than the WMT task considered in the paper, but it illustrates the whole system. We also show how to use multi-gpu processing to make it really fast.
# +
# #!pip install torchtext spacy
# #!python -m spacy download en
# #!python -m spacy download de
# -
# ## Data Loading
# > We will load the dataset using torchtext and spacy for tokenization.
# +
# For data loading.
from torchtext import data, datasets
if True:
import spacy
spacy_de = spacy.load('de')
spacy_en = spacy.load('en')
def tokenize_de(text):
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
return [tok.text for tok in spacy_en.tokenizer(text)]
BOS_WORD = '<s>'
EOS_WORD = '</s>'
BLANK_WORD = "<blank>"
SRC = data.Field(tokenize=tokenize_de, pad_token=BLANK_WORD)
TGT = data.Field(tokenize=tokenize_en, init_token = BOS_WORD,
eos_token = EOS_WORD, pad_token=BLANK_WORD)
MAX_LEN = 100
train, val, test = datasets.IWSLT.splits(
exts=('.de', '.en'), fields=(SRC, TGT),
filter_pred=lambda x: len(vars(x)['src']) <= MAX_LEN and
len(vars(x)['trg']) <= MAX_LEN)
MIN_FREQ = 2
SRC.build_vocab(train.src, min_freq=MIN_FREQ)
TGT.build_vocab(train.trg, min_freq=MIN_FREQ)
# -
# > Batching matters a ton for speed. We want to have very evenly divided batches, with absolutely minimal padding. To do this we have to hack a bit around the default torchtext batching. This code patches their default batching to make sure we search over enough sentences to find tight batches.
# ## Iterators
# +
class MyIterator(data.Iterator):
def create_batches(self):
if self.train:
def pool(d, random_shuffler):
for p in data.batch(d, self.batch_size * 100):
p_batch = data.batch(
sorted(p, key=self.sort_key),
self.batch_size, self.batch_size_fn)
for b in random_shuffler(list(p_batch)):
yield b
self.batches = pool(self.data(), self.random_shuffler)
else:
self.batches = []
for b in data.batch(self.data(), self.batch_size,
self.batch_size_fn):
self.batches.append(sorted(b, key=self.sort_key))
def rebatch(pad_idx, batch):
"Fix order in torchtext to match ours"
src, trg = batch.src.transpose(0, 1), batch.trg.transpose(0, 1)
return Batch(src, trg, pad_idx)
# -
# ## Multi-GPU Training
#
# > Finally to really target fast training, we will use multi-gpu. This code implements multi-gpu word generation. It is not specific to transformer so I won't go into too much detail. The idea is to split up word generation at training time into chunks to be processed in parallel across many different gpus. We do this using pytorch parallel primitives:
#
# * replicate - split modules onto different gpus.
# * scatter - split batches onto different gpus
# * parallel_apply - apply module to batches on different gpus
# * gather - pull scattered data back onto one gpu.
# * nn.DataParallel - a special module wrapper that calls these all before evaluating.
#
# Skip if not interested in multigpu.
class MultiGPULossCompute:
"A multi-gpu loss compute and train function."
def __init__(self, generator, criterion, devices, opt=None, chunk_size=5):
# Send out to different gpus.
self.generator = generator
self.criterion = nn.parallel.replicate(criterion,
devices=devices)
self.opt = opt
self.devices = devices
self.chunk_size = chunk_size
def __call__(self, out, targets, normalize):
total = 0.0
generator = nn.parallel.replicate(self.generator,
devices=self.devices)
out_scatter = nn.parallel.scatter(out,
target_gpus=self.devices)
out_grad = [[] for _ in out_scatter]
targets = nn.parallel.scatter(targets,
target_gpus=self.devices)
# Divide generating into chunks.
chunk_size = self.chunk_size
for i in range(0, out_scatter[0].size(1), chunk_size):
# Predict distributions
out_column = [[Variable(o[:, i:i+chunk_size].data,
requires_grad=self.opt is not None)]
for o in out_scatter]
gen = nn.parallel.parallel_apply(generator, out_column)
# Compute loss.
y = [(g.contiguous().view(-1, g.size(-1)),
t[:, i:i+chunk_size].contiguous().view(-1))
for g, t in zip(gen, targets)]
loss = nn.parallel.parallel_apply(self.criterion, y)
# Sum and normalize loss
l = nn.parallel.gather(loss,
target_device=self.devices[0])
l = l.sum()[0] / normalize
total += l.data[0]
# Backprop loss to output of transformer
if self.opt is not None:
l.backward()
for j, l in enumerate(loss):
out_grad[j].append(out_column[j][0].grad.data.clone())
# Backprop all loss through transformer.
if self.opt is not None:
out_grad = [Variable(torch.cat(og, dim=1)) for og in out_grad]
o1 = out
o2 = nn.parallel.gather(out_grad,
target_device=self.devices[0])
o1.backward(gradient=o2)
self.opt.step()
self.opt.optimizer.zero_grad()
return total * normalize
# > Now we create our model, criterion, optimizer, data iterators, and paralelization
# GPUs to use
devices = [0, 1, 2, 3]
if True:
pad_idx = TGT.vocab.stoi["<blank>"]
model = make_model(len(SRC.vocab), len(TGT.vocab), N=6)
model.cuda()
criterion = LabelSmoothing(size=len(TGT.vocab), padding_idx=pad_idx, smoothing=0.1)
criterion.cuda()
BATCH_SIZE = 12000
train_iter = MyIterator(train, batch_size=BATCH_SIZE, device=0,
repeat=False, sort_key=lambda x: (len(x.src), len(x.trg)),
batch_size_fn=batch_size_fn, train=True)
valid_iter = MyIterator(val, batch_size=BATCH_SIZE, device=0,
repeat=False, sort_key=lambda x: (len(x.src), len(x.trg)),
batch_size_fn=batch_size_fn, train=False)
model_par = nn.DataParallel(model, device_ids=devices)
None
# > Now we train the model. I will play with the warmup steps a bit, but everything else uses the default parameters. On an AWS p3.8xlarge with 4 Tesla V100s, this runs at ~27,000 tokens per second with a batch size of 12,000
# ## Training the System
# +
# #!wget https://s3.amazonaws.com/opennmt-models/iwslt.pt
# -
if False:
model_opt = NoamOpt(model.src_embed[0].d_model, 1, 2000,
torch.optim.Adam(model.parameters(), lr=0, betas=(0.9, 0.98), eps=1e-9))
for epoch in range(10):
model_par.train()
run_epoch((rebatch(pad_idx, b) for b in train_iter),
model_par,
MultiGPULossCompute(model.generator, criterion,
devices=devices, opt=model_opt))
model_par.eval()
loss = run_epoch((rebatch(pad_idx, b) for b in valid_iter),
model_par,
MultiGPULossCompute(model.generator, criterion,
devices=devices, opt=None))
print(loss)
else:
model = torch.load("iwslt.pt")
# > Once trained we can decode the model to produce a set of translations. Here we simply translate the first sentence in the validation set. This dataset is pretty small so the translations with greedy search are reasonably accurate.
for i, batch in enumerate(valid_iter):
src = batch.src.transpose(0, 1)[:1]
src_mask = (src != SRC.vocab.stoi["<blank>"]).unsqueeze(-2)
out = greedy_decode(model, src, src_mask,
max_len=60, start_symbol=TGT.vocab.stoi["<s>"])
print("Translation:", end="\t")
for i in range(1, out.size(1)):
sym = TGT.vocab.itos[out[0, i]]
if sym == "</s>": break
print(sym, end =" ")
print()
print("Target:", end="\t")
for i in range(1, batch.trg.size(0)):
sym = TGT.vocab.itos[batch.trg.data[i, 0]]
if sym == "</s>": break
print(sym, end =" ")
print()
break
# # Additional Components: BPE, Search, Averaging
# > So this mostly covers the transformer model itself. There are four aspects that we didn't cover explicitly. We also have all these additional features implemented in [OpenNMT-py](https://github.com/opennmt/opennmt-py).
#
#
# > 1) BPE/ Word-piece: We can use a library to first preprocess the data into subword units. See <NAME>'s [subword-nmt](https://github.com/rsennrich/subword-nmt) implementation. These models will transform the training data to look like this:
# ▁Die ▁Protokoll datei ▁kann ▁ heimlich ▁per ▁E - Mail ▁oder ▁FTP ▁an ▁einen ▁bestimmte n ▁Empfänger ▁gesendet ▁werden .
# > 2) Shared Embeddings: When using BPE with shared vocabulary we can share the same weight vectors between the source / target / generator. See the [(cite)](https://arxiv.org/abs/1608.05859) for details. To add this to the model simply do this:
if False:
model.src_embed[0].lut.weight = model.tgt_embeddings[0].lut.weight
model.generator.lut.weight = model.tgt_embed[0].lut.weight
# > 3) Beam Search: This is a bit too complicated to cover here. See the [OpenNMT-py](https://github.com/OpenNMT/OpenNMT-py/blob/master/onmt/translate/Beam.py) for a pytorch implementation.
#
#
# > 4) Model Averaging: The paper averages the last k checkpoints to create an ensembling effect. We can do this after the fact if we have a bunch of models:
def average(model, models):
"Average models into model"
for ps in zip(*[m.params() for m in [model] + models]):
p[0].copy_(torch.sum(*ps[1:]) / len(ps[1:]))
# # Results
#
# On the WMT 2014 English-to-German translation task, the big transformer model (Transformer (big)
# in Table 2) outperforms the best previously reported models (including ensembles) by more than 2.0
# BLEU, establishing a new state-of-the-art BLEU score of 28.4. The configuration of this model is
# listed in the bottom line of Table 3. Training took 3.5 days on 8 P100 GPUs. Even our base model
# surpasses all previously published models and ensembles, at a fraction of the training cost of any of
# the competitive models.
#
# On the WMT 2014 English-to-French translation task, our big model achieves a BLEU score of 41.0,
# outperforming all of the previously published single models, at less than 1/4 the training cost of the
# previous state-of-the-art model. The Transformer (big) model trained for English-to-French used
# dropout rate Pdrop = 0.1, instead of 0.3.
#
#
Image(filename="images/results.png")
# > The code we have written here is a version of the base model. There are fully trained version of this system available here [(Example Models)](http://opennmt.net/Models-py/).
# >
# > With the addtional extensions in the last section, the OpenNMT-py replication gets to 26.9 on EN-DE WMT. Here I have loaded in those parameters to our reimplemenation.
# !wget https://s3.amazonaws.com/opennmt-models/en-de-model.pt
model, SRC, TGT = torch.load("en-de-model.pt")
model.eval()
sent = "▁The ▁log ▁file ▁can ▁be ▁sent ▁secret ly ▁with ▁email ▁or ▁FTP ▁to ▁a ▁specified ▁receiver".split()
src = torch.LongTensor([[SRC.stoi[w] for w in sent]])
src = Variable(src)
src_mask = (src != SRC.stoi["<blank>"]).unsqueeze(-2)
out = greedy_decode(model, src, src_mask,
max_len=60, start_symbol=TGT.stoi["<s>"])
print("Translation:", end="\t")
trans = "<s> "
for i in range(1, out.size(1)):
sym = TGT.itos[out[0, i]]
if sym == "</s>": break
trans += sym + " "
print(trans)
# ## Attention Visualization
#
# > Even with a greedy decoder the translation looks pretty good. We can further visualize it to see what is happening at each layer of the attention
# +
tgt_sent = trans.split()
def draw(data, x, y, ax):
seaborn.heatmap(data,
xticklabels=x, square=True, yticklabels=y, vmin=0.0, vmax=1.0,
cbar=False, ax=ax)
for layer in range(1, 6, 2):
fig, axs = plt.subplots(1,4, figsize=(20, 10))
print("Encoder Layer", layer+1)
for h in range(4):
draw(model.encoder.layers[layer].self_attn.attn[0, h].data,
sent, sent if h ==0 else [], ax=axs[h])
plt.show()
for layer in range(1, 6, 2):
fig, axs = plt.subplots(1,4, figsize=(20, 10))
print("Decoder Self Layer", layer+1)
for h in range(4):
draw(model.decoder.layers[layer].self_attn.attn[0, h].data[:len(tgt_sent), :len(tgt_sent)],
tgt_sent, tgt_sent if h ==0 else [], ax=axs[h])
plt.show()
print("Decoder Src Layer", layer+1)
fig, axs = plt.subplots(1,4, figsize=(20, 10))
for h in range(4):
draw(model.decoder.layers[layer].self_attn.attn[0, h].data[:len(tgt_sent), :len(sent)],
sent, tgt_sent if h ==0 else [], ax=axs[h])
plt.show()
# -
# # Conclusion
#
# > Hopefully this code is useful for future research. Please reach out if you have any issues. If you find this code helpful, also check out our other OpenNMT tools.
#
# ```
# @inproceedings{opennmt,
# author = {<NAME> and
# <NAME> and
# <NAME> and
# <NAME> and
# <NAME>},
# title = {OpenNMT: Open-Source Toolkit for Neural Machine Translation},
# booktitle = {Proc. ACL},
# year = {2017},
# url = {https://doi.org/10.18653/v1/P17-4012},
# doi = {10.18653/v1/P17-4012}
# }
# ```
#
# > Cheers,
# > srush
# {::options parse_block_html="true" /}
# <div id="disqus_thread"></div>
# <script>
#
# /**
# * RECOMMENDED CONFIGURATION VARIABLES: EDIT AND UNCOMMENT THE SECTION BELOW TO INSERT DYNAMIC VALUES FROM YOUR PLATFORM OR CMS.
# * LEARN WHY DEFINING THESE VARIABLES IS IMPORTANT: https://disqus.com/admin/universalcode/#configuration-variables*/
# /*
# var disqus_config = function () {
# this.page.url = PAGE_URL; // Replace PAGE_URL with your page's canonical URL variable
# this.page.identifier = PAGE_IDENTIFIER; // Replace PAGE_IDENTIFIER with your page's unique identifier variable
# };
# */
# (function() { // DON'T EDIT BELOW THIS LINE
# var d = document, s = d.createElement('script');
# s.src = 'https://harvard-nlp.disqus.com/embed.js';
# s.setAttribute('data-timestamp', +new Date());
# (d.head || d.body).appendChild(s);
# })();
# </script>
# <noscript>Please enable JavaScript to view the <a href="https://disqus.com/?ref_noscript">comments powered by Disqus.</a></noscript>
#
# <div id="disqus_thread"></div>
# <script>
# /**
# * RECOMMENDED CONFIGURATION VARIABLES: EDIT AND UNCOMMENT THE SECTION BELOW TO INSERT DYNAMIC VALUES FROM YOUR PLATFORM OR CMS.
# * LEARN WHY DEFINING THESE VARIABLES IS IMPORTANT: https://disqus.com/admin/universalcode/#configuration-variables
# */
# /*
# var disqus_config = function () {
# this.page.url = PAGE_URL; // Replace PAGE_URL with your page's canonical URL variable
# this.page.identifier = PAGE_IDENTIFIER; // Replace PAGE_IDENTIFIER with your page's unique identifier variable
# };
# */
# (function() { // REQUIRED CONFIGURATION VARIABLE: EDIT THE SHORTNAME BELOW
# var d = document, s = d.createElement('script');
#
# s.src = 'https://EXAMPLE.disqus.com/embed.js'; // IMPORTANT: Replace EXAMPLE with your forum shortname!
#
# s.setAttribute('data-timestamp', +new Date());
# (d.head || d.body).appendChild(s);
# })();
# </script>
# <noscript>Please enable JavaScript to view the <a href="https://disqus.com/?ref_noscript" rel="nofollow">comments powered by Disqus.</a></noscript>
| The Annotated Transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rY0wXimFDpyK"
# [](https://colab.research.google.com/github/eirasf/GCED-AA2/blob/main/lab1/4_Numpy.ipynb)
# + id="Xrn19tJL2xbq"
import tensorflow as tf
# + id="RG8Hyj5UFWxi"
import numpy as np
many_ones = np.ones([3, 3])
# + colab={"base_uri": "https://localhost:8080/"} id="BtrHHEUaFs33" outputId="de55e168-c109-40f7-ab12-d492c4fdfaae"
tensor = tf.multiply(many_ones, 42)
print(tensor)
| lab1/4_Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.10 64-bit ('neurolib')
# language: python
# name: python3710jvsc74a57bd040c7a7921e4911bf33743545bc23717ac3cf742f9e21e92f1609fffc896830ac
# ---
# # Creating new model from scratch
#
# The real power of `MultiModel` framework is in fast prototyping of heterogeneous models. To showcase this, in this notebook we create a brand new model in the framework (the famous Jansen-Rit model), and then create a thalamocortical mini network with 1 node representing a thalamus and 1 node Jansen-Rit model representing a cortical column.
#
# ## Jansen-Rit model
# The Jansen-Rit model is a neural population model of a local cortical circuit. It contains three interconnected neural populations: one for the pyramidal projection neurons and two for excitatory and inhibitory interneurons forming feedback loops.
#
# The equations for Jansen-Rit model reads:
# \begin{align}
# \ddot{x}_{0} & = Aa\cdot\mathrm{Sigm}\left(x_{1} - x_{2}\right) - 2a\dot{x}_{0} - a^{2}x_{0} \\
# \ddot{x}_{1} & = Aa[p + C_{2}\mathrm{Sigm}\left(C_{1}x_{0}\right)] - 2a\dot{x}_{1} - a^2x_{1} \\
# \ddot{x}_{2} & = BbC_{4}\mathrm{Sigm}\left( C_{3}x_{o} \right) - 2b\dot{x}_{2} - b^2x_{2} \\
# \mathrm{Sigm}(x) & = \frac{v_{max}}{1 + \mathrm{e}^{r(v_{0} - x)}}
# \end{align}
# Of course, in order to implement the above equations numerically, the system of three second-order ODEs will be rewritten into a system of six first-order ODEs.
#
# ## `MultiModel` strategy
# The actual implementation will be a bit more involved than simply writing down the above equations. The building block of any proper `MultiModel` is a `NeuralMass`. Jansen-Rit model actually summarises an activity of a cortical column consisting of three populations: a population of pyramidal cells interacting with two populations of interneurons - one excitatory and one inhibitory. Moreover, the $x$ represent the average membrane potential, but typically, neuronal models (at least in `neurolib`) are coupled via firing rate. For this reason, our main output variable would actually be a firing rate of a main, pyramidal population. The average membrane potential of a pyramidal population is $x = x_{1} - x_{2}$ and its firing rate is then $r = \mathrm{Sigm}(x) = \mathrm{Sigm}(x_{1} - x_{2})$. Similar strategy (sigmoidal transfer function for average membrane potential) is used for the thalamic model.
#
# The coupling variable in `MultiModel` must be the same across all hierarchical levels. Individial populations in Jansen-Rit model are coupled via their average membrane potentials $x_{i}$, $i\in[0, 1, 2]$. However, the "global" coupling variable for the node would be the firing rate of pyramidal population $r$, introduced in the paragraph above. To reconcile this, two options exists in `MultiModel`:
# * create a `NeuralMass` representing a pyramidal population with two coupling variables: $x_{0}$ and $r$; `NeuralMass` representing interneurons would have one coupling variable, $x_{1,2}$
# * advantages: cleaner (implementation), more modular (can create J-R `Node` with more masses than 3 very easily)
# * disadvantages: more code, might be harder to navigate for the beginner
# * create one `NeuralMass` representing all three populations with one coupling variable $r$, since the others are not really coupling since the whole dynamics is contained in one object
# * advantages: less code, easier to grasp
# * disadvantages: less modular, cannot simply edit J-R model, since everything is "hardcoded" into one `NeuralMass` object
#
# In order to build a basic understanding of building blocks, we will follow the second option here (less code, easier to grasp). For interested readers, the first option (modular one) will be implemented in `MultiModel`, so you can follow the model files.
#
# Our strategy for this notebook then would be:
# 1. implement single `NeuralMass` object representing all three populations in the Jansen-Rit model, with single coupling variable $r$
# 2. implement a "dummy" `Node` with single `NeuralMass` (requirement, one cannot couple `Node` to a `NeuralMass` to build a network)
# 3. experiment with connecting this Jansen-Rit cortical model to the thalamic population model
#
# Last note, all model's dynamics and parameters in `MultiModel` are defined in milliseconds, therefore we will scale the default parameters.
#
# Let us start with the imports:
import matplotlib.pyplot as plt
import numpy as np
import symengine as se
from IPython.display import display
from jitcdde import input as system_input
from neurolib.models.multimodel import MultiModel, ThalamicNode
from neurolib.models.multimodel.builder.base.constants import LAMBDA_SPEED
from neurolib.models.multimodel.builder.base.network import Network, Node
from neurolib.models.multimodel.builder.base.neural_mass import NeuralMass
from neurolib.utils.functions import getPowerSpectrum
from neurolib.utils.stimulus import Input, OrnsteinUhlenbeckProcess, StepInput
# A quick detour before we dive into the model itself. Jansen-Rit model is typically driven with a uniformly distributed noise, as the authors wanted to model nonspecific input (they used the term *background spontaneous activity*). For this we quickly create our model input using the `ModelInput` class (the tutorial on how to use stimuli in `neurolib` is given elsewhere).
class UniformlyDistributedNoise(Input):
"""
Uniformly distributed noise process between two values.
"""
def __init__(self, low, high, n=1, seed=None):
# save arguments as attributes for later
self.low = low
self.high = high
# init super
super().__init__(n=n, seed=seed)
def generate_input(self, duration, dt):
# generate time vector
self._get_times(duration=duration, dt=dt)
# generate noise process itself with the correct shape
# as (time steps x num. processes)
return np.random.uniform(
self.low, self.high, (self.n, self.times.shape[0])
)
# let us build a proper hierarchy, i.e. we firstly build a Jansen-Rit mass
class SingleJansenRitMass(NeuralMass):
"""
Single Jansen-Rit mass implementing whole three population dynamics.
Reference:
<NAME>., & <NAME>. (1995). Electroencephalogram and visual evoked potential
generation in a mathematical model of coupled cortical columns. Biological cybernetics,
73(4), 357-366.
"""
# all these attributes are compulsory to fill in
name = "Jansen-Rit mass"
label = "JRmass"
num_state_variables = 7 # 6 ODEs + firing rate coupling variable
num_noise_variables = 1 # single external input
# NOTE
# external inputs (so-called noise_variables) are typically background noise drive in models,
# however, this can be any type of stimulus - periodic stimulus, step stimulus, square pulse,
# anything. Therefore you may want to add more stimuli, e.g. for Jansen-Rit model three to each
# of its population. Here we do not stimulate our Jansen-Rit model, so only use actual noise
# drive to excitatory interneuron population.
# as dictionary {index of state var: it's name}
coupling_variables = {6: "r_mean_EXC"}
# as list
state_variable_names = [
"v_pyr",
"dv_pyr",
"v_exc",
"dv_exc",
"v_inh",
"dv_inh",
# to comply with other `MultiModel` nodes
"r_mean_EXC",
]
# as list
# note on parameters C1 - C4 - all papers use one C and C1-C4 are
# defined as various rations of C, typically: C1 = C, C2 = 0.8*C
# C3 = C4 = 0.25*C, therefore we use only `C` and scale it in the
# dynamics definition
required_params = [
"A",
"a",
"B",
"b",
"C",
"v_max",
"v0",
"r",
"lambda",
]
# list of required couplings when part of a `Node` or `Network`
# `network_exc_exc` is the default excitatory coupling between nodes
required_couplings = ["network_exc_exc"]
# here we define the default noise input to Jansen-Rit model (this can be changed later)
# for a quick test, we follow the original Jansen and Rit paper and use uniformly distributed
# noise between 120 - 320 Hz; but we do it in kHz, hence 0.12 - 0.32
# fix seed for reproducibility
_noise_input = [UniformlyDistributedNoise(low=0.12, high=0.32, seed=42)]
def _sigmoid(self, x):
"""
Sigmoidal transfer function which is the same for all populations.
"""
# notes:
# - all parameters are accessible as self.params - it is a dictionary
# - mathematical definition (ODEs) is done in symbolic mathematics - all functions have to be
# imported from `symengine` module, hence se.exp which is a symbolic exponential function
return self.params["v_max"] / (
1.0 + se.exp(self.params["r"] * (self.params["v0"] - x))
)
def __init__(self, params=None, seed=None):
# init this `NeuralMass` - use passed parameters or default ones
# parameters are now accessible as self.params, seed as self.seed
super().__init__(params=params or JR_DEFAULT_PARAMS, seed=seed)
def _initialize_state_vector(self):
"""
Initialize state vector.
"""
np.random.seed(self.seed)
# random in average potentials around zero
self.initial_state = (
np.random.normal(size=self.num_state_variables)
# * np.array([10.0, 0.0, 10.0, 0.0, 10.0, 0.0, 0.0])
).tolist()
def _derivatives(self, coupling_variables):
"""
Here the magic happens: dynamics is defined here using symbolic maths package symengine.
"""
# first, we need to unwrap state vector
(
v_pyr,
dv_pyr,
v_exc,
dv_exc,
v_inh,
dv_inh,
firing_rate,
) = self._unwrap_state_vector() # this function does everything for us
# now we need to write down our dynamics
# PYR dynamics
d_v_pyr = dv_pyr
d_dv_pyr = (
self.params["A"] * self.params["a"] * self._sigmoid(v_exc - v_inh)
- 2 * self.params["a"] * dv_pyr
- self.params["a"] ** 2 * v_pyr
)
# EXC dynamics: system input comes into play here
d_v_exc = dv_exc
d_dv_exc = (
self.params["A"]
* self.params["a"]
* (
# system input as function from jitcdde (also in symengine) with proper index:
# in our case we have only one noise input (can be more), so index 0
system_input(self.noise_input_idx[0])
# C2 = 0.8*C, C1 = C
+ (0.8 * self.params["C"]) * self._sigmoid(self.params["C"] * v_pyr)
)
- 2 * self.params["a"] * dv_exc
- self.params["a"] ** 2 * v_exc
)
# INH dynamics
d_v_inh = dv_inh
d_dv_inh = (
self.params["B"] * self.params["b"]
# C3 = C4 = 0.25 * C
* (0.25 * self.params["C"])
* self._sigmoid((0.25 * self.params["C"]) * v_pyr)
- 2 * self.params["b"] * dv_inh
- self.params["b"] ** 2 * v_inh
)
# firing rate computation
# firing rate as dummy dynamical variable with infinitely fast
# fixed point dynamics
firing_rate_now = self._sigmoid(v_exc - v_inh)
d_firing_rate = -self.params["lambda"] * (firing_rate - firing_rate_now)
# now just return a list of derivatives in the correct order
return [d_v_pyr, d_dv_pyr, d_v_exc, d_dv_exc, d_v_inh, d_dv_inh, d_firing_rate]
# And we are done with the basics! Only thing we really need is to define attributes (such as how many variables we have, what couplings we have, what about noise, etc.) and the actual dynamics as symbolic expressions. Symbolic expressions are easy: are basic operators like `+`, `-`, `*`, `/`, or `**` are overloaded, which means you can simply use them and do not think about. Functions such as `sin`, `log`, or `exp` must be imported from `symengine` and used. Now we define a default set of parameters. Do not forget - `MultiModel` defines all in ms, therefore the parameters needs to be in ms, kHz, and similar.
JR_DEFAULT_PARAMS = {
"A": 3.25, # mV
"B": 22.0, # mV
# `a` and `b` are originally 100Hz and 50Hz
"a": 0.1, # kHz
"b": 0.05, # kHz
"v0": 6.0, # mV
# v_max is originally 5Hz
"v_max": 0.005, # kHz
"r": 0.56, # m/V
"C": 135.0,
# parameter for dummy `r` dynamics
"lambda": LAMBDA_SPEED,
}
# The next step is to create a `Node`. `Node` is second level in the hierarchy and already it can be wrapped into `MultiModel` and treated as any other `neurolib` model. On our case, creating the `Node` is really simple: it has only one mass, no delays, and no connectivity.
class JansenRitNode(Node):
"""
Jansen-Rit node with 1 neural mass representing 3 population model.
"""
name = "Jansen-Rit node"
label = "JRnode"
# if Node is integrated isolated, what network input we should use
# zero by default = no network input for one-node model
default_network_coupling = {"network_exc_exc": 0.0}
# default output is the firing rate of pyramidal population
default_output = "r_mean_EXC"
# list of all variables that are accessible as outputs
output_vars = ["r_mean_EXC", "v_pyr", "v_exc", "v_inh"]
def __init__(self, params=None, seed=None):
# in `Node` __init__, the list of masses is created and passed
jr_mass = SingleJansenRitMass(params=params, seed=seed)
# each mass has to have index, in this case it is simply 0
jr_mass.index = 0
# call super and properly initialize a Node
super().__init__(neural_masses=[jr_mass])
self.excitatory_masses = np.array([0])
def _sync(self):
# this function typically defines the coupling between masses
# within one node, but in our case there is nothing to define
return []
# And we are done. At this point, we can integrate our Jansen-Rit model and see some results.
#
# ## Test and simulate Jansen-Rit model
# In order to simulate our newly created model, we just need to wrap it with `MultiModel` as in the last example and see how it goes.
# +
# init model - fix seed for reproducibility (random init. conditions)
jr_model = MultiModel.init_node(JansenRitNode(seed=42))
# see parameters
print("Parameters:")
display(jr_model.params)
print("")
# see describe
print("Describe:")
display(jr_model.model_instance.describe())
# -
# run model for 5 seconds - all in ms
jr_model.params["sampling_dt"] = 1.0
jr_model.params["duration"] = 5000
jr_model.params["backend"] = "jitcdde"
jr_model.run()
# +
_, axs = plt.subplots(
nrows=2, ncols=2, figsize=(12, 6), gridspec_kw={"width_ratios": [2, 1]}
)
axs[0, 0].plot(jr_model.t, jr_model.v_exc.T - jr_model.v_inh.T, color="k")
axs[0, 0].set_ylabel("LFP")
axs[1, 0].plot(jr_model.t, jr_model.r_mean_EXC.T * 1000, color="navy")
axs[1, 0].set_ylabel("PYR r [Hz]")
axs[1, 0].set_xlabel("time [sec]")
fr, po = getPowerSpectrum(
(jr_model.v_exc.T - jr_model.v_inh.T).squeeze(),
dt=jr_model.params["sampling_dt"],
maxfr=40.0,
)
axs[0, 1].semilogy(fr, po, color="indianred")
axs[0, 1].set_xlabel("frequency [Hz]")
axs[0, 1].set_ylabel("log power")
axs[1, 1].hist(jr_model.v_exc.T - jr_model.v_inh.T, bins=15, color="gray")
axs[1, 1].set_xlabel("LFP")
axs[1, 1].set_ylabel("#")
plt.suptitle("`jitcdde` backend")
# -
# The results looks good! With the same parameters as original Jansen and Rit paper, we got the $\alpha$ activity with spectral peak around 10 Hz. Just as a proof of concept - let us try the second `MultiModel` backend.
# run model for 5 seconds - all in ms
jr_model.params["sampling_dt"] = 1.0
jr_model.params["dt"] = 0.1
jr_model.params["duration"] = 5000
jr_model.params["backend"] = "numba"
jr_model.run()
# +
_, axs = plt.subplots(
nrows=2, ncols=2, figsize=(12, 6), gridspec_kw={"width_ratios": [2, 1]}
)
axs[0, 0].plot(jr_model.t, jr_model.v_exc.T - jr_model.v_inh.T, color="k")
axs[0, 0].set_ylabel("LFP")
axs[1, 0].plot(jr_model.t, jr_model.r_mean_EXC.T * 1000, color="navy")
axs[1, 0].set_ylabel("PYR r [Hz]")
axs[1, 0].set_xlabel("time [sec]")
fr, po = getPowerSpectrum(
(jr_model.v_exc.T - jr_model.v_inh.T).squeeze(),
dt=jr_model.params["sampling_dt"],
maxfr=40.0,
)
axs[0, 1].semilogy(fr, po, color="indianred")
axs[0, 1].set_xlabel("frequency [Hz]")
axs[0, 1].set_ylabel("log power")
axs[1, 1].hist(jr_model.v_exc.T - jr_model.v_inh.T, bins=15, color="gray")
axs[1, 1].set_xlabel("LFP")
axs[1, 1].set_ylabel("#")
plt.suptitle("`numba` backend")
# -
# All works as it should, we have our Jansen-Rit model! In the next step, we will showcase how the new model can be connected and coupled to other models (similar as in the first example).
#
# ## Couple Jansen-Rit to thalamic model
# Here we practically copy the previous example where we coupled ALN node with a thalamic node, but instead of ALN representing a cortical column, we use our brand new Jansen-Rit model.
# +
# let us start by subclassing the Network
class JansenRitThalamusMiniNetwork(Network):
"""
Simple thalamocortical motif: 1 cortical node Jansen-Rit + 1 NMM thalamus.
"""
# provide basic attributes as name and label
name = "Jansen-Rit 1 node + Thalamus"
label = "JRThlmNet"
# define which variables are used to sync, i.e. what coupling variables our nodes need
sync_variables = [
# both nodes are connected via excitatory synapses
"network_exc_exc",
# and INH mass in thalamus also receives excitatory coupling
"network_inh_exc",
]
# lastly, we need to define what is default output of the network (this has to be the
# variable present in all nodes)
# for us it is excitatory firing rates
default_output = f"r_mean_EXC"
# define all output vars of any interest to us - EXC and INH firing rates
output_vars = [f"r_mean_EXC", f"r_mean_INH"]
def __init__(self, connectivity_matrix, delay_matrix, seed=None):
# self connections are resolved within nodes, so zeroes at the diagonal
assert np.all(np.diag(connectivity_matrix) == 0.0)
# init Jansen-Rit node with index 0
jr_node = JansenRitNode(seed=seed)
jr_node.index = 0
# index where the state variables start - for first node it is always 0
jr_node.idx_state_var = 0
# set correct indices for noise input - in JR we have only one noise source
jr_node[0].noise_input_idx = [0]
# init thalamus node with index 1
thalamus = ThalamicNode()
thalamus.index = 1
# thalamic state variables start where ALN state variables end - easy
thalamus.idx_state_var = jr_node.num_state_variables
# set correct indices of noise input - one per mass, after ALN noise
# indices
for mass in thalamus:
mass.noise_input_idx = [jr_node.num_noise_variables + mass.index]
# now super.__init__ network with these two nodes:
super().__init__(
nodes=[jr_node, thalamus],
connectivity_matrix=connectivity_matrix,
delay_matrix=delay_matrix,
)
# done! the only other thing we need to do, is to set the coupling variables
# thalamus vs. Jansen-Rit are coupled via their firing rates and here we setup the
# coupling matrices; the super class `Network` comes with some convenient
# functions for this
def _sync(self):
"""
Set coupling variables - the ones we defined in `sync_variables`
_sync returns a list of tuples where the first element in each tuple is the coupling "symbol"
and the second is the actual mathematical expression
for the ease of doing this, `Network` class contains convenience functions for this:
- _additive_coupling
- _diffusive_coupling
- _no_coupling
here we use additive coupling only
"""
# get indices of coupling variables from all nodes
exc_indices = [
next(
iter(
node.all_couplings(
mass_indices=node.excitatory_masses.tolist()
)
)
)
for node in self
]
assert len(exc_indices) == len(self)
return (
# basic EXC <-> EXC coupling
# within_node_idx is a list of len 2 (because we have two nodes)
# with indices of coupling variables within the respective state vectors
self._additive_coupling(
within_node_idx=exc_indices, symbol="network_exc_exc"
)
# EXC -> INH coupling (only to thalamus)
+ self._additive_coupling(
within_node_idx=exc_indices,
symbol="network_inh_exc",
connectivity=self.connectivity,
)
+ super()._sync()
)
# -
# lets check what we have
# in the ALN-thalamus case the matrix was [0.0, 0.15], [1.2, 0.0] - JR produces firing rates around 5Hz (5 times lower than ALN)
SC = np.array([[0.0, 0.15], [6., 0.0]])
delays = np.array([[0.0, 13.0], [13.0, 0.0]]) # thalamocortical delay = 13ms
thalamocortical = MultiModel(JansenRitThalamusMiniNetwork(connectivity_matrix=SC, delay_matrix=delays, seed=42))
# original `MultiModel` instance is always accessible as `MultiModel.model_instance`
display(thalamocortical.model_instance.describe())
# fix parameters for interesting regime
thalamocortical.params["*g_LK"] = 0.032 # K-leak conductance in thalamus
thalamocortical.params["*TCR*input*sigma"] = 0.005 # noise in thalamus
thalamocortical.params["*input*tau"] = 5.0 # timescale of OU process
thalamocortical.params["duration"] = 20000. # 20 seconds simulation
thalamocortical.params["sampling_dt"] = 1.0
thalamocortical.run()
_, axs = plt.subplots(nrows=2, ncols=1, sharex=True, figsize=(12, 6))
axs[0].plot(thalamocortical.t, thalamocortical.r_mean_EXC[0, :].T)
axs[0].set_ylabel("Jansen-Rit firing rate [kHz]")
axs[1].plot(thalamocortical.t, thalamocortical.r_mean_EXC[1, :].T, color="C1")
axs[1].set_ylabel("thalamus firing rate [kHz]")
axs[1].set_xlabel("time [sec]")
# So the model works, just like that! Of course, in this case we do not see anything interesting in the modelled dynamics, since we would need to proper investigation of various parameters. Consider this as a proof-of-concept, where we can very easily couple two very different population models.
| examples/example-4.1-multimodel-custom-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains course material from [CBE30338](https://jckantor.github.io/CBE30338)
# by <NAME> (jeff at nd.edu); the content is available [on Github](https://github.com/jckantor/CBE30338.git).
# The text is released under the [CC-BY-NC-ND-4.0 license](https://creativecommons.org/licenses/by-nc-nd/4.0/legalcode),
# and code is released under the [MIT license](https://opensource.org/licenses/MIT).*
# <!--NAVIGATION-->
# < [Linearization](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/03.00-Linearization.ipynb) | [Contents](toc.ipynb) | [Linear Approximation of a Process Model](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/03.02-Linear-Approximation-of-a-Process-Model.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE30338/blob/master/notebooks/03.01-Step-Response-of-a-Gravity-Drained-Tank.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE30338/master/notebooks/03.01-Step-Response-of-a-Gravity-Drained-Tank.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
# + [markdown] slideshow={"slide_type": "slide"}
# # Step Response of a Gravity Drained Tank
#
# by <NAME> (jeff at nd.edu). The latest version of this notebook is available at [https://github.com/jckantor/CBE30338](https://github.com/jckantor/CBE30338).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Summary
#
# In the example we show how to fit the step response of a nonlinear system, a gravity drained tank, to a first order linear system.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Mass Balance for Tank with Constant Cross-Sectional Area
#
# For a tank with constant cross-sectional area, such as a cylindrical or rectangular tank, the liquid height is described by a differential equation
#
# $$A\frac{dh}{dt} = q_{in}(t) - q_{out}(t)$$
#
# where $q_{out}$ is a function of liquid height. Torricelli's law tells the outlet flow from the tank is proportional to square root of the liquid height
#
# $$ q_{out}(h) = C_v\sqrt{h} $$
# + [markdown] slideshow={"slide_type": "slide"}
# Dividing by area we obtain a nonlinear ordinary differential equation
#
# $$ \frac{dh}{dt} = - \frac{C_V}{A}\sqrt{h} + \frac{1}{A}q_{in}(t) $$
#
# in our standard form where the LHS derivative appears with a constant coefficient of 1.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Nonlinear Step Response
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
Cv = 0.1 # Outlet valve constant [cubic meters/min/meter^1/2]
A = 1.0 # Tank area [meter^2]
# inlet flow rate in cubic meters/min
def qin(t):
return 0.15
def deriv(h,t):
return qin(t)/A - Cv*np.sqrt(h)/A
IC = [0.0]
t = np.linspace(0,200,101)
h = odeint(deriv,IC,t)
plt.plot(t,h)
plt.xlabel('Time [min]')
plt.ylabel('Level [m]')
plt.grid();
# + [markdown] slideshow={"slide_type": "slide"}
# ## Linear Approximation of the Step Response
#
# The step response of the gravity drained to a change in flowrate looks similar to the step response of a firat order linear system. Let's try a linear approximation
#
# $$\tau\frac{dx}{dt} + x = Ku$$
#
# which has a step response solution that can be written
#
# $$x(t) = x_{ss} + (x_0 - x_{ss})\exp(-t/\tau)$$
#
# where $x_{ss} = Ku_{ss}$. There are two parameters, $K$ and $\tau$, which we need to estimate in order to fit the linear approximation to the nonlinear simulation results computed above.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fit for $K$
#
# The steady state gain $K$ of the linear system is given by
#
# $$ K = \frac{x_{ss} - x(0)}{u_{ss} - u_0}$$
#
# where $u_0$ is the initial input, $u_{ss}$ is the steady-state input, and $x_0$ and $x_{ss}$ are corresponding values of the state variable. In the case of liquid level, $h\sim x$ and $q_{in}\sim u$, therefore an estimate of $K$ is
#
# $$ K = \frac{h_{ss} - h_0}{q_{in,ss} - q_{in,0}}$$
# + slideshow={"slide_type": "slide"}
q0,h0 = 0,0 # initial conditions
qss = qin(t[-1]) # final input
hss = h[-1] # python way to get the last element in a list
K = (hss-h0)/(qss-q0) # step change in output divided by step change in input
print('Steady-State Gain is approximately = ', K)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fit for $\tau$
#
# From the formula for the solution of a first-order linear equation with constant input,
#
# $$\frac{x_{ss} - x(t)}{x_{ss} - x_0} = \exp(-t/\tau) \qquad \implies \qquad \tau = \frac{-t}{\ln\frac{x_{ss} - x(t)}{x_{ss} - x_0}}$$
#
# We pick one point representative of the transient portion of the nonlinear response. In this case the response at $t = 25$ minutes accounts for $\approx$60% of the ultimate response, so we choose point as a representative point.
# + slideshow={"slide_type": "slide"}
k = sum(t<25) # find index in t corresponding to 25 minutes
tk = t[k]
hk = h[k]
tau = -tk/np.log((hss-hk)/(hss-h0))
print('Estimated time constant is ', tau)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Comparing the linear approximation to the nonlinear simulation
# +
u0 = q0
uss = qss
xss = K*(uss - u0)
xpred = xss - xss*np.exp(-t/tau)
plt.plot(t,h)
plt.plot(t,xpred)
plt.legend(['Nonlinear Simulation','Linear Approximation'])
plt.xlabel('Time [min]')
plt.ylabel('Level [m]')
plt.title('Nonlinear Simulation vs. Linear Approximation');
# -
# <!--NAVIGATION-->
# < [Linearization](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/03.00-Linearization.ipynb) | [Contents](toc.ipynb) | [Linear Approximation of a Process Model](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/03.02-Linear-Approximation-of-a-Process-Model.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE30338/blob/master/notebooks/03.01-Step-Response-of-a-Gravity-Drained-Tank.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE30338/master/notebooks/03.01-Step-Response-of-a-Gravity-Drained-Tank.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
| Mathematics/Mathematical Modeling/03.01-Step-Response-of-a-Gravity-Drained-Tank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Fitting Models Exercise 2
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt
# + [markdown] nbgrader={}
# ## Fitting a decaying oscillation
# + [markdown] nbgrader={}
# For this problem you are given a raw dataset in the file `decay_osc.npz`. This file contains three arrays:
#
# * `tdata`: an array of time values
# * `ydata`: an array of y values
# * `dy`: the absolute uncertainties (standard deviations) in y
#
# Your job is to fit the following model to this data:
#
# $$ y(t) = A e^{-\lambda t} \cos{\omega t + \delta} $$
#
# First, import the data using NumPy and make an appropriately styled error bar plot of the raw data.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
f = np.load('decay_osc.npz', mmap_mode='r')
# -
list(f)
ydata = f['ydata']
dy = f['dy']
tdata = f['tdata']
plt.figure(figsize=(10,5))
plt.errorbar(tdata, ydata, dy, fmt='.k', ecolor='lightgray')
plt.xlabel("t", fontsize=14)
plt.ylabel("y", fontsize=14)
ax = plt.gca()
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.spines['bottom'].set_color('#a2a7ff')
ax.spines['left'].set_color('#a2a7ff')
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
plt.title("Decaying Oscillations Plot with Error")
plt.show()
# + deletable=false nbgrader={"checksum": "8fe685c8222cc4b0b71fde4d0409d50f", "grade": true, "grade_id": "fittingmodelsex02a", "points": 5}
assert True # leave this to grade the data import and raw data plot
# + [markdown] nbgrader={}
# Now, using `curve_fit` to fit this model and determine the estimates and uncertainties for the parameters:
#
# * Print the parameters estimates and uncertainties.
# * Plot the raw and best fit model.
# * You will likely have to pass an initial guess to `curve_fit` to get a good fit.
# * Treat the uncertainties in $y$ as *absolute errors* by passing `absolute_sigma=True`.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def model(t, A, lamb, omega, delta):
return A*np.exp(-lamb*t)*np.cos(omega*t) + delta
# -
theta_best, theta_cov = opt.curve_fit(model, tdata, ydata, sigma=dy)
print('A = {0:.3f} +/- {1:.3f}'.format(theta_best[0],np.sqrt(theta_cov[0,0])))
print('lambda = {0:.3f} +/- {1:.3f}'.format(theta_best[1],np.sqrt(theta_cov[1,1])))
print('omega = {0:.3f} +/- {1:.3f}'.format(theta_best[2],np.sqrt(theta_cov[2,2])))
print('delta = {0:.3f} +/- {1:.3f}'.format(theta_best[3],np.sqrt(theta_cov[3,3])))
Y = theta_best[0]*np.exp(-theta_best[1]*tdata)*np.cos(theta_best[2]*tdata) + theta_best[3]
plt.figure(figsize=(10,5))
plt.plot(tdata,Y)
plt.errorbar(tdata, ydata, dy, fmt='.k', ecolor='lightgray')
plt.xlabel("t", fontsize=14)
plt.ylabel("y", fontsize=14)
ax = plt.gca()
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.spines['bottom'].set_color('#a2a7ff')
ax.spines['left'].set_color('#a2a7ff')
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
plt.title("Curve Fit for Decaying Oscillation Plot with Error")
plt.show()
# + deletable=false nbgrader={"checksum": "abacc1ad72e3412252e4ed47c8f65897", "grade": true, "grade_id": "fittingmodelsex02b", "points": 5}
assert True # leave this cell for grading the fit; should include a plot and printout of the parameters+errors
| assignments/assignment12/FittingModelsEx02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# language: python
# name: python3
# ---
class Solution(object):
def reverse(self, x):
"""
:type x: int
:rtype: int
"""
#check and store if the number if positive or negative
positive_num=1
abs_x = x
if (x<0):
positive_num=0
abs_x = -x
#calculate and store each digit in a list in reverse order
reverse_list = []
remaining=abs_x
while (remaining>=10):
last_num = remaining%10
remaining = remaining//10
reverse_list.append(last_num)
reverse_list.append(remaining)
#construct the reverse number
result=0
for i in range(len(reverse_list)):
result = result + reverse_list[i]*(10**(len(reverse_list)-i-1))
#if the original number if negative, then the reverse number is also negative
if positive_num==0:
result = 0-result
#set to 0 if the obtained reverse number goes outside the signed 32-bit integer range
if (result > 2**31-1 or result < -2**31):
result = 0
return result
sol = Solution()
print(sol.reverse(132))
print(sol.reverse(12345678999))
| Problems/7-ReverseInteger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Bootcamp Ciência de Dados - IGTI
# Esse notebook se refere ao segundo módulo do bootcamp ciência de dados e tem como objetivo responder ao segundo trabalho prático.
#
# * Originalmente o trabalho proposto utilizava o mysql e foi adaptado para o MongoDB
# ## Módulo 2 - Coleta e obtenção de dados
from pymongo import MongoClient
import csv
from glob import glob
import pandas as pd
import json
# ### Criando o Database e Conectando ao MongoDB
mongo_client=MongoClient()
db = mongo_client['bootcamp']
db.segment.drop()
# ### Carga individualizada dos arquivos .csv locais
with open('imoveis.csv', 'r', encoding="latin-1") as arquivo_csv:
leitor = csv.DictReader(arquivo_csv)
for coluna in leitor:
db.imoveis.insert(coluna)
with open('caracteristicasgerais.csv', 'r', encoding="latin-1") as arquivo_csv:
leitor = csv.DictReader(arquivo_csv)
for coluna in leitor:
db.caracteristicasgerais.insert(coluna)
with open('caracteristicageralimovel.csv', 'r', encoding="latin-1") as arquivo_csv:
leitor = csv.DictReader(arquivo_csv)
for coluna in leitor:
db.caracteristicageralimovel.insert(coluna)
with open('cidades.csv', 'r', encoding="latin-1") as arquivo_csv:
leitor = csv.DictReader(arquivo_csv)
for coluna in leitor:
db.cidade.insert(coluna)
with open('estados.csv', 'r', encoding="latin-1") as arquivo_csv:
leitor = csv.DictReader(arquivo_csv, delimiter=';')
for coluna in leitor:
db.estado.insert(coluna)
# ### Consultas para responder as questões
#
# +
#Q1. Quantos registros são retornados ao executar o seguinte comando sql no banco de dados após ter povoado a tabela de estados: SELECT * FROM estado WHERE NomeEstado like 'P%'
db.cidade.find({$in: "CodEstadoIBGE": "11"}).count()
# +
#Q2. Após executar todos os passos do enunciado do trabalho prático e ter povoado todas as tabelas do banco de dados, qual o total de registros da tabela ‘caracteristicasgerais’?
# -
#Q3. Crie uma consulta sql para verificar a quantidade de imóveis cadastrados no estado de RS. Após executar esse sql, qual o valor retornado?
# +
#Crie uma consulta sql para contar quantos estados possuem imóveis cadastrados em que o valor do IPTU é igual a R$ 0,00. Após executar esse sql, qual o valor retornado?
# +
#Quantos registros foram inseridos na tabela ‘caracteristicageralimovel’? Crie um sql que realize a consulta.
# +
#Qual o número do registro do imóvel (coluna codRegistro) que possui o maior valor de condomínio? Crie um sql que realize a consulta.
| Modulo2_ColetaeObtencaoDados/TrabalhoPratico/ColetaDados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: shamo
# language: python
# name: shamo
# ---
# + raw_mimetype="text/restructuredtext" active=""
# HD-tDCS simulation
# ==================
#
# To simulate HD-tDCS is very easy in *shamo*.
#
# The first step is to load the finite element model created before.
# +
from shamo import FEM
model = FEM.load("../../derivatives/fem_from_labels/fem_from_labels.json")
# + raw_mimetype="text/restructuredtext" active=""
# Next we import the :py:class:`~shamo.hd_tdcs.simulation.single.problem.ProbHDTDCSSim` class and create an instance of it.
# +
from shamo.hd_tdcs import ProbHDTDCSSim
problem = ProbHDTDCSSim()
# + raw_mimetype="text/restructuredtext" active=""
# Then, as for the EEG leadfield computation, we set the electrical conductivity of the tissues, setup the reference electrodes and source. We also set the injected current (A).
# -
problem.sigmas.set("scalp", 0.4137)
problem.sigmas.set("gm", 0.4660)
problem.sigmas.set("wm", 0.2126)
problem.source.add("P3")
problem.references.adds(["T5", "C3", "PZ", "O1"])
problem.current = (0.004)
# +
import nibabel as nib
img = nib.load(model.path / f"{model.name}.nii")
problem.grid.set(img.affine, img.shape)
# + raw_mimetype="text/restructuredtext" active=""
# Finally, we solve the problem and obtain the different field generated in the whole brain both in the form of POS files that can be opened in Gmsh and in the form of NII files that can be visualised in FSLeyes or any other nifti file opener.
# -
solution = problem.solve("single", "../../derivatives/hd_tdcs_simulation", model)
| code/hd_tdcs/hd_tdcs_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Code is developed on Visual Studio Code (Ubuntu OS)
Created by: <NAME>
Time: 11:00 pm 8 September
Task Provider: The Sparks Foundation (Machine Leaning Intern - Task 2)
Feel free to download the code and check it
Any errors or questions can contact me(details in readme)
Note: Check the requirements.txt file before executing the code
"""
# this code snippet manages the voice pace and rate and checks whether the modules are available or not.
from os import system
system("clear")
import pyttsx3
voiceEngine = pyttsx3.init()
rate = voiceEngine.getProperty('rate')
volume = voiceEngine.getProperty('volume')
newVoiceRate,newVolume = 160, 0.4
voiceEngine.setProperty('rate', newVoiceRate)
voiceEngine.setProperty('volume', newVolume)
pyttsx3.speak("Hello I am Sai's personal assistant")
pyttsx3.speak("Am here to assist you through out the execution.")
pyttsx3.speak(" ")
try:
pyttsx3.speak(" importing the modules ")
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import pyttsx3
print("-"*42, "Successfully imported the modules", "-"*42)
pyttsx3.speak("Successfully imported the modules")
except:
print("Oops!, few modules are missing\nInstall the modules by executing `pip install -r requirements.txt`")
pyttsx3.speak("Oops!, few modules are missing kindly install the modules")
# -
# this snippet does the preprcessing work, like data shape, head, info, describe, isnull, dtypes..
def preprocessing(data):
pyttsx3.speak("Before applying the algorithm its good practise to have a look at the data")
print(" ")
pyttsx3.speak("would you like to see dataset information")
user_input = input("would you like to see dataset information (y/n)?..")
user_input = user_input.lower()
if(('y' in user_input) or ('yes' in user_input)):
print(" ")
print("-"*45, "DataSet Information", "-"*45)
print('\nThe shape of the data is: ',data.shape)
print(" ")
print('\nDataset: \n',data.head())
print(" ")
print("\nChecking for null values: \n",data.isnull().sum())
print(" ")
print("\nChecking the dataypes: \n",data.dtypes)
print(" ")
print("\nDescription about the data: \n",data.describe())
print(" ")
data.info()
pyttsx3.speak(" As I have manually checked, no null values are present in the data, more information about the dataset is available on scree, go thorugh it")
# this snippet holds the logic of regression task
def task2(data):
pyttsx3.speak("Were dealing with regression task")
print("-"*48, "Regression Task", "-"*48)
preprocessing(data)
pyttsx3.speak("lets have visualisation of the data")
plt.scatter(x ='Hours', y = 'Scores', data = data)
plt.title('Hours vs Score')
plt.xlabel('Hours Studied')
plt.ylabel('Score obtained')
plt.show()
pyttsx3.speak("As you seen the plot, you can observe that it has good correlation")
sns.pairplot(data)
pyttsx3.speak("We see that there is linear relationship between 'Scores' and 'Hours'. So our assumption of Linearity for linear regession is verified.")
sns.heatmap(data.corr(),annot=True,cmap='winter')
pyttsx3.speak("Lets find out the correlation of them")
pyttsx3.speak("sci kitlearn module is used for the implementation of regression model")
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
print("\nDividing the dataset into attributes and labels :")
pyttsx3.speak("Dividing the dataset into attributes and labels")
x = data.iloc[:, :-1].values
y = data.iloc[:, 1].values
print("\nAttributes:\n",x)
print("\nLables :",y)
pyttsx3.speak("Now lets split the data into train set and test set")
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2, random_state = 0)
pyttsx3.speak("Default test size is used to split")
print("\nDataset has got split into train and test sets, these are the shapes")
print("XTrain set : ", x_train.shape)
print("XTest set : ", x_test.shape)
print("YTrain set : ", y_train.shape)
print("YTest set : ", y_test.shape)
pyttsx3.speak("Successfully the data got split lets train the algorithm by creating its instance")
print("-"*48, "Training the Algorithm", "-"*48)
pyttsx3.speak("lets start training the model")
print(" ")
regr = LinearRegression()
pyttsx3.speak("model instance is created")
print(" ")
regr.fit(x_train, y_train)
pyttsx3.speak("we fitted the data to the model")
print(" ")
line = regr.coef_*x+regr.intercept_
pyttsx3.speak("lets have look at the data and the regression line")
print(" ")
plt.scatter(x, y)
plt.plot(x, line)
plt.title('Hours vs Score')
plt.xlabel('Hours Studied')
plt.ylabel('Score obtained')
plt.show()
pyttsx3.speak("model has been trained successfully")
print(" ")
print("-"*48, "Testing the Algorithm", "-"*48)
pyttsx3.speak("lets start testing the Algorithm")
y_pred = regr.predict(x_test)
pyttsx3.speak("making predictions")
print(" ")
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
pyttsx3.speak("successfully predicted the test data, lets look at few eavluation metrics")
print('Mean Absolute Error :', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error :', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error :', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
pyttsx3.speak("These are the different errors caluculated")
print("\nWhat will be predicted score if a student study for 9.25 hrs in a day?")
pyttsx3.speak(" But still we havent answered the actual question What will be predicted score if a student study for 9.25 hours in a day?")
y_pred = regr.predict([[9.25]])
print("The predicted score is:", y_pred)
pyttsx3.speak("Prediction done on the trained regression model")
flag1 = True
while flag1 == True:
pyttsx3.speak("Why dont we try with different hours")
user_input = int(input("\nLets try with different value of hours:"))
y_pred = regr.predict([[user_input]])
print("The predicted score is:", y_pred)
pyttsx3.speak("Prediction is done, check the value")
fl = input("Do you want to try it again (y/n)?..")
if('yes' in fl or 'y' in fl):
flag1 = True
else:
flag1 = False
pyttsx3.speak("Its good to assist you I learned a lot about simple linear regression thank you")
# this snippet holds the logic of clustering task.
def task3(data):
pyttsx3.speak("Were dealing with Clustering task")
print("-"*48, "\nClustering Task", "-"*48)
preprocessing(data)
x = data.iloc[:, [0, 1, 2, 3]].values
from sklearn.cluster import KMeans
ls = []
for i in range(1, 11):
kmeans = KMeans(n_clusters = i, init = 'k-means++', max_iter = 300, n_init = 10, random_state = 0)
kmeans.fit(x)
ls.append(kmeans.inertia_)
print("Plotting the Elbow curve")
pyttsx3.speak("Lets plot the eblow curve")
plt.plot(range(1, 11), ls)
plt.title('The elbow method')
plt.xlabel('Number of clusters')
plt.ylabel('')
plt.show()
pyttsx3.speak(" The optimal values of k using elbow curve is 3, so we can set the k value to 3.")
print("\nApplying kmeans instance on the dataset")
pyttsx3.speak("Applying kmeans instance on the dataset")
kmeans = KMeans(n_clusters = 3, init = 'k-means++', max_iter = 300, n_init = 10, random_state = 0)
pyttsx3.speak("Predictions are made")
y_kmeans = kmeans.fit_predict(x)
pyttsx3.speak("Now lets visualise the Clusters foremd")
plt.scatter(x[y_kmeans == 0, 0], x[y_kmeans == 0, 1], s = 100, c = 'red', label = 'Iris-setosa')
plt.scatter(x[y_kmeans == 1, 0], x[y_kmeans == 1, 1], s = 100, c = 'blue', label = 'Iris-versicolour')
plt.scatter(x[y_kmeans == 2, 0], x[y_kmeans == 2, 1], s = 100, c = 'green', label = 'Iris-virginica')
pyttsx3.speak("Plotting the centroids of the cluster")
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:,1], s = 100, c = 'yellow', label = 'Centroids')
plt.legend()
print(" ")
pyttsx3.speak("Its good to assist you I learned a lot about simple Clustering techniques thank you")
# this snippet holds the logic of decision tree task.
def task4(data):
print("Still in developing process")
pyttsx3.speak("Still in developing process..")
# this snippet holds the logic of decision tree task.
def task5(data):
print("Still in developing process")
pyttsx3.speak("Still in developing process..")
# snippet loads the data and initiates the each tasks.
if __name__=="__main__":
pyttsx3.speak("Which task you want me to per")
print("TASKs available are \n1. TASK2 \n2. TASK3 \n3. TASK4 \n4. TASK5")
flag = True
while flag:
user_input = input("Feed me the task name : ")
if(('task2' in user_input) or ('2' in user_input)):
data = pd.read_csv("data.csv")
task2(data)
elif(('task3' in user_input) or ('3' in user_input)):
data = pd.read_csv("iris.csv")
task3(data)
elif(('task4' in user_input) or ('4' in user_input)):
data = pd.read_csv("iris.csv")
task4(data)
elif(('task5' in user_input) or ('5' in user_input)):
data = pd.read_csv("SampleStore.csv")
task5(data)
else:
pass
pyttsx3.speak("Do you want to try another tasks?")
usin = input("Do you want to try another tasks (y/n)? ")
if 'Y' in usin or 'y' in usin or 'yes' in usin:
pass
else:
flag = False
| alltasks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Tutorial #1: Train an image classification model with Azure Machine Learning
#
# In this tutorial, you train a machine learning model on remote compute resources. You'll use the training and deployment workflow for Azure Machine Learning service (preview) in a Python Jupyter notebook. You can then use the notebook as a template to train your own machine learning model with your own data. This tutorial is **part one of a two-part tutorial series**.
#
# This tutorial trains a simple logistic regression using the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset and [scikit-learn](http://scikit-learn.org) with Azure Machine Learning. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of 28x28 pixels, representing a number from 0 to 9. The goal is to create a multi-class classifier to identify the digit a given image represents.
#
# Learn how to:
#
# > * Set up your development environment
# > * Access and examine the data
# > * Train a simple logistic regression model on a remote cluster
# > * Review training results, find and register the best model
#
# You'll learn how to select a model and deploy it in [part two of this tutorial](deploy-models.ipynb) later.
#
# ## Prerequisites
#
# See prerequisites in the [Azure Machine Learning documentation](https://docs.microsoft.com/azure/machine-learning/service/tutorial-train-models-with-aml#prerequisites).
# ## Set up your development environment
#
# All the setup for your development work can be accomplished in a Python notebook. Setup includes:
#
# * Importing Python packages
# * Connecting to a workspace to enable communication between your local computer and remote resources
# * Creating an experiment to track all your runs
# * Creating a remote compute target to use for training
#
# ### Import packages
#
# Import Python packages you need in this session. Also display the Azure Machine Learning SDK version.
# + tags=["check version"]
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import azureml.core
from azureml.core import Workspace
# check core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
# -
# ### Connect to workspace
#
# Create a workspace object from the existing workspace. `Workspace.from_config()` reads the file **config.json** and loads the details into an object named `ws`.
# + tags=["load workspace"]
# load workspace configuration from the config.json file in the current folder.
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, ws.location, sep='\t')
# -
# ### Create experiment
#
# Create an experiment to track the runs in your workspace. A workspace can have muliple experiments.
# + tags=["create experiment"]
experiment_name = 'sklearn-mnist'
from azureml.core import Experiment
exp = Experiment(workspace=ws, name=experiment_name)
# -
# ### Create or Attach existing compute resource
# By using Azure Machine Learning Compute, a managed service, data scientists can train machine learning models on clusters of Azure virtual machines. Examples include VMs with GPU support. In this tutorial, you create Azure Machine Learning Compute as your training environment. The code below creates the compute clusters for you if they don't already exist in your workspace.
#
# **Creation of compute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace the code will skip the creation process.
# + tags=["create mlc", "amlcompute"]
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
import os
# choose a name for your cluster
compute_name = os.environ.get("AML_COMPUTE_CLUSTER_NAME", "cpu-cluster")
compute_min_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MIN_NODES", 0)
compute_max_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MAX_NODES", 4)
# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6
vm_size = os.environ.get("AML_COMPUTE_CLUSTER_SKU", "STANDARD_D2_V2")
if compute_name in ws.compute_targets:
compute_target = ws.compute_targets[compute_name]
if compute_target and type(compute_target) is AmlCompute:
print('found compute target. just use it. ' + compute_name)
else:
print('creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,
min_nodes = compute_min_nodes,
max_nodes = compute_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current AmlCompute status, use get_status()
print(compute_target.get_status().serialize())
# -
# You now have the necessary packages and compute resources to train a model in the cloud.
#
# ## Explore data
#
# Before you train a model, you need to understand the data that you are using to train it. You also need to copy the data into the cloud so it can be accessed by your cloud training environment. In this section you learn how to:
#
# * Download the MNIST dataset
# * Display some sample images
# * Upload data to the cloud
#
# ### Download the MNIST dataset
#
# Download the MNIST dataset and save the files into a `data` directory locally. Images and labels for both training and testing are downloaded.
# +
import urllib.request
data_folder = os.path.join(os.getcwd(), 'data')
os.makedirs(data_folder, exist_ok=True)
urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz', filename=os.path.join(data_folder, 'train-images.gz'))
urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz', filename=os.path.join(data_folder, 'train-labels.gz'))
urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename=os.path.join(data_folder, 'test-images.gz'))
urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename=os.path.join(data_folder, 'test-labels.gz'))
# -
# ### Display some sample images
#
# Load the compressed files into `numpy` arrays. Then use `matplotlib` to plot 30 random images from the dataset with their labels above them. Note this step requires a `load_data` function that's included in an `util.py` file. This file is included in the sample folder. Please make sure it is placed in the same folder as this notebook. The `load_data` function simply parses the compresse files into numpy arrays.
# +
# make sure utils.py is in the same directory as this code
from utils import load_data
# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the model converge faster.
X_train = load_data(os.path.join(data_folder, 'train-images.gz'), False) / 255.0
X_test = load_data(os.path.join(data_folder, 'test-images.gz'), False) / 255.0
y_train = load_data(os.path.join(data_folder, 'train-labels.gz'), True).reshape(-1)
y_test = load_data(os.path.join(data_folder, 'test-labels.gz'), True).reshape(-1)
# now let's show some randomly chosen images from the traininng set.
count = 0
sample_size = 30
plt.figure(figsize = (16, 6))
for i in np.random.permutation(X_train.shape[0])[:sample_size]:
count = count + 1
plt.subplot(1, sample_size, count)
plt.axhline('')
plt.axvline('')
plt.text(x=10, y=-10, s=y_train[i], fontsize=18)
plt.imshow(X_train[i].reshape(28, 28), cmap=plt.cm.Greys)
plt.show()
# -
# Now you have an idea of what these images look like and the expected prediction outcome.
#
# ### Upload data to the cloud
#
# Now make the data accessible remotely by uploading that data from your local machine into Azure so it can be accessed for remote training. The datastore is a convenient construct associated with your workspace for you to upload/download data, and interact with it from your remote compute targets. It is backed by Azure blob storage account.
#
# The MNIST files are uploaded into a directory named `mnist` at the root of the datastore.
# + tags=["use datastore"]
ds = ws.get_default_datastore()
print(ds.datastore_type, ds.account_name, ds.container_name)
ds.upload(src_dir=data_folder, target_path='mnist', overwrite=True, show_progress=True)
# -
# ## Train on a remote cluster
#
# For this task, submit the job to the remote training cluster you set up earlier. To submit a job you:
# * Create a directory
# * Create a training script
# * Create an estimator object
# * Submit the job
#
# ### Create a directory
#
# Create a directory to deliver the necessary code from your computer to the remote resource.
import os
script_folder = os.path.join(os.getcwd(), "sklearn-mnist")
os.makedirs(script_folder, exist_ok=True)
# ### Create a training script
#
# To submit the job to the cluster, first create a training script. Run the following code to create the training script called `train.py` in the directory you just created.
# +
# %%writefile $script_folder/train.py
import argparse
import os
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.externals import joblib
from azureml.core import Run
from utils import load_data
# let user feed in 2 parameters, the location of the data files (from datastore), and the regularization rate of the logistic regression model
parser = argparse.ArgumentParser()
parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')
parser.add_argument('--regularization', type=float, dest='reg', default=0.01, help='regularization rate')
args = parser.parse_args()
data_folder = args.data_folder
print('Data folder:', data_folder)
# load train and test set into numpy arrays
# note we scale the pixel intensity values to 0-1 (by dividing it with 255.0) so the model can converge faster.
X_train = load_data(os.path.join(data_folder, 'train-images.gz'), False) / 255.0
X_test = load_data(os.path.join(data_folder, 'test-images.gz'), False) / 255.0
y_train = load_data(os.path.join(data_folder, 'train-labels.gz'), True).reshape(-1)
y_test = load_data(os.path.join(data_folder, 'test-labels.gz'), True).reshape(-1)
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape, sep = '\n')
# get hold of the current run
run = Run.get_context()
print('Train a logistic regression model with regularization rate of', args.reg)
clf = LogisticRegression(C=1.0/args.reg, solver="liblinear", multi_class="auto", random_state=42)
clf.fit(X_train, y_train)
print('Predict the test set')
y_hat = clf.predict(X_test)
# calculate accuracy on the prediction
acc = np.average(y_hat == y_test)
print('Accuracy is', acc)
run.log('regularization rate', np.float(args.reg))
run.log('accuracy', np.float(acc))
os.makedirs('outputs', exist_ok=True)
# note file saved in the outputs folder is automatically uploaded into experiment record
joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')
# -
# Notice how the script gets data and saves models:
#
# + The training script reads an argument to find the directory containing the data. When you submit the job later, you point to the datastore for this argument:
# `parser.add_argument('--data-folder', type=str, dest='data_folder', help='data directory mounting point')`
#
# + The training script saves your model into a directory named outputs. <br/>
# `joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')`<br/>
# Anything written in this directory is automatically uploaded into your workspace. You'll access your model from this directory later in the tutorial.
# The file `utils.py` is referenced from the training script to load the dataset correctly. Copy this script into the script folder so that it can be accessed along with the training script on the remote resource.
import shutil
shutil.copy('utils.py', script_folder)
# ### Create an estimator
#
# An estimator object is used to submit the run. Azure Machine Learning has pre-configured estimators for common machine learning frameworks, as well as generic Estimator. Create SKLearn estimator for scikit-learn model, by specifying
#
# * The name of the estimator object, `est`
# * The directory that contains your scripts. All the files in this directory are uploaded into the cluster nodes for execution.
# * The compute target. In this case you will use the AmlCompute you created
# * The training script name, train.py
# * Parameters required from the training script
#
# In this tutorial, this target is AmlCompute. All files in the script folder are uploaded into the cluster nodes for execution. The data_folder is set to use the datastore (`ds.path('mnist').as_mount()`).
# + tags=["configure estimator"]
from azureml.train.sklearn import SKLearn
script_params = {
'--data-folder': ds.path('mnist').as_mount(),
'--regularization': 0.5
}
est = SKLearn(source_directory=script_folder,
script_params=script_params,
compute_target=compute_target,
entry_script='train.py')
# -
# This is what the mounting point looks like:
print(ds.path('mnist').as_mount())
# ### Submit the job to the cluster
#
# Run the experiment by submitting the estimator object. And you can navigate to Azure portal to monitor the run.
# + tags=["remote run", "amlcompute", "scikit-learn"]
run = exp.submit(config=est)
run
# -
# Since the call is asynchronous, it returns a **Preparing** or **Running** state as soon as the job is started.
#
# ## Monitor a remote run
#
# In total, the first run takes **approximately 10 minutes**. But for subsequent runs, as long as the dependencies (`conda_packages` parameter in the above estimator constructor) don't change, the same image is reused and hence the container start up time is much faster.
#
# Here is what's happening while you wait:
#
# - **Image creation**: A Docker image is created matching the Python environment specified by the estimator. The image is built and stored in the ACR (Azure Container Registry) associated with your workspace. Image creation and uploading takes **about 5 minutes**.
#
# This stage happens once for each Python environment since the container is cached for subsequent runs. During image creation, logs are streamed to the run history. You can monitor the image creation progress using these logs.
#
# - **Scaling**: If the remote cluster requires more nodes to execute the run than currently available, additional nodes are added automatically. Scaling typically takes **about 5 minutes.**
#
# - **Running**: In this stage, the necessary scripts and files are sent to the compute target, then data stores are mounted/copied, then the entry_script is run. While the job is running, stdout and the files in the ./logs directory are streamed to the run history. You can monitor the run's progress using these logs.
#
# - **Post-Processing**: The ./outputs directory of the run is copied over to the run history in your workspace so you can access these results.
#
#
# You can check the progress of a running job in multiple ways. This tutorial uses a Jupyter widget as well as a `wait_for_completion` method.
#
# ### Jupyter widget
#
# Watch the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
# + tags=["use notebook widget"]
from azureml.widgets import RunDetails
RunDetails(run).show()
# -
# By the way, if you need to cancel a run, you can follow [these instructions](https://aka.ms/aml-docs-cancel-run).
# ### Get log results upon completion
#
# Model training happens in the background. You can use `wait_for_completion` to block and wait until the model has completed training before running more code.
# + tags=["remote run", "amlcompute", "scikit-learn"]
# specify show_output to True for a verbose log
run.wait_for_completion(show_output=True)
# -
# ### Display run results
#
# You now have a model trained on a remote cluster. Retrieve all the metrics logged during the run, including the accuracy of the model:
# + tags=["get metrics"]
print(run.get_metrics())
# -
# In the next tutorial you will explore this model in more detail.
#
# ## Register model
#
# The last step in the training script wrote the file `outputs/sklearn_mnist_model.pkl` in a directory named `outputs` in the VM of the cluster where the job is executed. `outputs` is a special directory in that all content in this directory is automatically uploaded to your workspace. This content appears in the run record in the experiment under your workspace. Hence, the model file is now also available in your workspace.
#
# You can see files associated with that run.
# + tags=["query history"]
print(run.get_file_names())
# -
# Register the model in the workspace so that you (or other collaborators) can later query, examine, and deploy this model.
# + tags=["register model from history"]
# register model
model = run.register_model(model_name='sklearn_mnist', model_path='outputs/sklearn_mnist_model.pkl')
print(model.name, model.id, model.version, sep='\t')
# -
# ## Next steps
#
# In this Azure Machine Learning tutorial, you used Python to:
#
# > * Set up your development environment
# > * Access and examine the data
# > * Train multiple models on a remote cluster using the popular scikit-learn machine learning library
# > * Review training details and register the best model
#
# You are ready to deploy this registered model using the instructions in the next part of the tutorial series:
#
# > [Tutorial 2 - Deploy models](img-classification-part2-deploy.ipynb)
# 
| tutorials/img-classification-part1-training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Evaluation, Cross-Validation, and Model Selection
# #### By <NAME> - [<EMAIL>](mailto:<EMAIL>) - http://github.com/karlnapf - http://herrstrathmann.de.
# Based on the model selection framework of his [Google summer of code 2011 project](http://www.google-melange.com/gsoc/project/google/gsoc2011/XXX) | <NAME> - [github.com/Saurabh7](https://github.com/Saurabh7) as a part of [Google Summer of Code 2014 project](http://www.google-melange.com/gsoc/project/details/google/gsoc2014/saurabh7/5750085036015616) mentored by - <NAME>
# This notebook illustrates the evaluation of prediction algorithms in Shogun using <a href="http://en.wikipedia.org/wiki/Cross-validation_(statistics)">cross-validation</a>, and selecting their parameters using <a href="http://en.wikipedia.org/wiki/Hyperparameter_optimization">grid-search</a>. We demonstrate this for a toy example on <a href="http://en.wikipedia.org/wiki/Binary_classification">Binary Classification</a> using <a href="http://en.wikipedia.org/wiki/Support_vector_machine">Support Vector Machines</a> and also a [regression](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLOOCrossValidationSplitting.html) problem on a real world dataset.
# 1. [General Idea](#General-Idea)
# 2. [Splitting Strategies](#Types-of-splitting-strategies)
# 1. [K-fold cross-validation](#K-fold-cross-validation)
# 2. [Stratified cross-validation](#Stratified-cross-validation)
# 3. [Example: Binary classification](#Toy-example:-Binary-Support-Vector-Classification)
# 4. [Example: Regression](#Regression-problem-and-cross-validation)
# 5. [Model Selection: Grid Search](#Model-selection-using-Grid-Search)
# ## General Idea
# Cross validation aims to estimate an algorithm's performance on unseen data. For example, one might be interested in the average classification accuracy of a Support Vector Machine when being applied to new data, that it was not trained on. This is important in order to compare the performance different algorithms on the same target. Most crucial is the point that the data that was used for running/training the algorithm is not used for testing. Different algorithms here also can mean different parameters of the same algorithm. Thus, cross-validation can be used to tune parameters of learning algorithms, as well as comparing different families of algorithms against each other. Cross-validation estimates are related to the marginal likelihood in Bayesian statistics in the sense that using them for selecting models avoids overfitting.
# Evaluating an algorithm's performance on training data should be avoided since the learner may adjust to very specific random features of the training data which are not very important to the general relation. This is called [overfitting](http://en.wikipedia.org/wiki/Overfitting). Maximising performance on the training examples usually results in algorithms explaining the noise in data (rather than actual patterns), which leads to bad performance on unseen data. This is one of the reasons behind splitting the data and using different splits for training and testing, which can be done using cross-validation.
# Let us generate some toy data for binary classification to try cross validation on.
# +
# %pylab inline
# %matplotlib inline
# include all Shogun classes
import os
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
from shogun import *
# generate some ultra easy training data
gray()
n=20
title('Toy data for binary classification')
X=hstack((randn(2,n), randn(2,n)+1))
Y=hstack((-ones(n), ones(n)))
_=scatter(X[0], X[1], c=Y , s=100)
p1 = Rectangle((0, 0), 1, 1, fc="w")
p2 = Rectangle((0, 0), 1, 1, fc="k")
legend((p1, p2), ["Class 1", "Class 2"], loc=2)
# training data in Shogun representation
feats=features(X)
labels=BinaryLabels(Y)
# -
# ### Types of splitting strategies
# As said earlier Cross-validation is based upon splitting the data into multiple partitions. Shogun has various strategies for this. The base class for them is [CSplittingStrategy](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CSplittingStrategy.html).
# #### K-fold cross-validation
# Formally, this is achieved via partitioning a dataset $X$ of size $|X|=n$ into $k \leq n$ disjoint partitions $X_i\subseteq X$ such that $X_1 \cup X_2 \cup \dots \cup X_n = X$ and $X_i\cap X_j=\emptyset$ for all $i\neq j$. Then, the algorithm is executed on all $k$ possibilities of merging $k-1$ partitions and subsequently tested on the remaining partition. This results in $k$ performances which are evaluated in some metric of choice (Shogun support multiple ones). The procedure can be repeated (on different splits) in order to obtain less variance in the estimate. See [1] for a nice review on cross-validation using different performance measures.
k=5
normal_split=CrossValidationSplitting(labels, k)
# #### Stratified cross-validation
# On classificaiton data, the best choice is [stratified cross-validation](http://en.wikipedia.org/wiki/Cross-validation_%28statistics%29#Common_types_of_cross-validation). This divides the data in such way that the fraction of labels in each partition is roughly the same, which reduces the variance of the performance estimate quite a bit, in particular for data with more than two classes. In Shogun this is implemented by [CStratifiedCrossValidationSplitting](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CStratifiedCrossValidationSplitting.html) class.
stratified_split=StratifiedCrossValidationSplitting(labels, k)
# #### Leave One Out cross-validation
# [Leave One Out Cross-validation](http://en.wikipedia.org/wiki/Cross-validation_%28statistics%29#Leave-one-out_cross-validation) holds out one sample as the validation set. It is thus a special case of K-fold cross-validation with $k=n$ where $n$ is number of samples. It is implemented in [LOOCrossValidationSplitting](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLOOCrossValidationSplitting.html) class.
# Let us visualize the generated folds on the toy data.
split_strategies=[stratified_split, normal_split]
# +
#code to visualize splitting
def get_folds(split, num):
split.build_subsets()
x=[]
y=[]
lab=[]
for j in range(num):
indices=split.generate_subset_indices(j)
x_=[]
y_=[]
lab_=[]
for i in range(len(indices)):
x_.append(X[0][indices[i]])
y_.append(X[1][indices[i]])
lab_.append(Y[indices[i]])
x.append(x_)
y.append(y_)
lab.append(lab_)
return x, y, lab
def plot_folds(split_strategies, num):
for i in range(len(split_strategies)):
x, y, lab=get_folds(split_strategies[i], num)
figure(figsize=(18,4))
gray()
suptitle(split_strategies[i].get_name(), fontsize=12)
for j in range(0, num):
subplot(1, num, (j+1), title='Fold %s' %(j+1))
scatter(x[j], y[j], c=lab[j], s=100)
_=plot_folds(split_strategies, 4)
# -
# Stratified splitting takes care that each fold has almost the same number of samples from each class. This is not the case with normal splitting which usually leads to imbalanced folds.
# ## Toy example: Binary Support Vector Classification
# Following the example from above, we will tune the performance of a SVM on the binary classification problem. We will
#
# * demonstrate how to evaluate a loss function or metric on a given algorithm
# * then learn how to estimate this metric for the algorithm performing on unseen data
# * and finally use those techniques to tune the parameters to obtain the best possible results.
#
# The involved methods are
#
# * [LibSVM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLibSVM.html) as the binary classification algorithms
# * the area under the ROC curve (AUC) as performance metric
# * three different kernels to compare
# +
# define SVM with a small rbf kernel (always normalise the kernel!)
C=1
kernel=GaussianKernel(2, 0.001)
kernel.init(feats, feats)
kernel.set_normalizer(SqrtDiagKernelNormalizer())
classifier=LibSVM(C, kernel, labels)
# train
_=classifier.train()
# -
# Ok, we now have performed classification on the training data. How good did this work? We can easily do this for many different performance measures.
# +
# instanciate a number of Shogun performance measures
metrics=[ROCEvaluation(), AccuracyMeasure(), ErrorRateMeasure(), F1Measure(), PrecisionMeasure(), RecallMeasure(), SpecificityMeasure()]
for metric in metrics:
print(metric.get_name(), metric.evaluate(classifier.apply(feats), labels))
# -
# Note how for example error rate is 1-accuracy. All of those numbers represent the training error, i.e. the ability of the classifier to explain the given data.
# Now, the training error is zero. This seems good at first. But is this setting of the parameters a good idea? No! A good performance on the training data alone does not mean anything. A simple look up table is able to produce zero error on training data. What we want is that our methods generalises the input data somehow to perform well on unseen data. We will now use cross-validation to estimate the performance on such.
#
# We will use [CStratifiedCrossValidationSplitting](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CStratifiedCrossValidationSplitting.html), which accepts a reference to the labels and the number of partitions as parameters. This instance is then passed to the class [CCrossValidation](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CCrossValidation.html), which does the estimation using the desired splitting strategy. The latter class can take all algorithms that are implemented against the [CMachine](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CMachine.html) interface.
# +
metric=AccuracyMeasure()
cross=CrossValidation(classifier, feats, labels, stratified_split, metric)
# perform the cross-validation, note that this call involved a lot of computation
result=cross.evaluate()
# the result needs to be casted to CrossValidationResult
result=CrossValidationResult.obtain_from_generic(result)
# this class contains a field "mean" which contain the mean performance metric
print("Testing", metric.get_name(), result.get_real('mean'))
# -
# Now this is incredibly bad compared to the training error. In fact, it is very close to random performance (0.5). The lesson: Never judge your algorithms based on the performance on training data!
#
# Note that for small data sizes, the cross-validation estimates are quite noisy. If we run it multiple times, we get different results.
print("Testing", metric.get_name(), [CrossValidationResult.obtain_from_generic(cross.evaluate()).get_real('mean') for _ in range(10)])
# It is better to average a number of different runs of cross-validation in this case. A nice side effect of this is that the results can be used to estimate error intervals for a given confidence rate.
# +
# 25 runs and 95% confidence intervals
cross.put('num_runs', 25)
# perform x-validation (now even more expensive)
cross.evaluate()
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
print("Testing cross-validation mean %.2f " \
% (result.get_real('mean')))
# -
# Using this machinery, it is very easy to compare multiple kernel parameters against each other to find the best one. It is even possible to compare a different kernel.
# +
widths=2**linspace(-5,25,10)
results=zeros(len(widths))
for i in range(len(results)):
kernel.set_width(widths[i])
result=CrossValidationResult.obtain_from_generic(cross.evaluate())
results[i]=result.get_mean()
plot(log2(widths), results, 'blue')
xlabel("log2 Kernel width")
ylabel(metric.get_name())
_=title("Accuracy for different kernel widths")
print("Best Gaussian kernel width %.2f" % widths[results.argmax()], "gives", results.max())
# compare this with a linear kernel
classifier.set_kernel(LinearKernel())
lin_k=CrossValidationResult.obtain_from_generic(cross.evaluate())
plot([log2(widths[0]), log2(widths[len(widths)-1])], [lin_k.get_mean(),lin_k.get_mean()], 'r')
# please excuse this horrible code :)
print("Linear kernel gives", lin_k.get_mean())
_=legend(["Gaussian", "Linear"], loc="lower center")
# -
# This gives a brute-force way to select paramters of any algorithm implemented under the [CMachine](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CMachine.html) interface. The cool thing about this is, that it is also possible to compare different model families against each other. Below, we compare a a number of regression models in Shogun on the Boston Housing dataset.
# ### Regression problem and cross-validation
# Various regression models in Shogun are now used to predict house prices using the [boston housing dataset](https://archive.ics.uci.edu/ml/datasets/Housing). Cross-validation is used to find best parameters and also test the performance of the models.
# +
feats=features(CSVFile(os.path.join(SHOGUN_DATA_DIR, 'uci/housing/fm_housing.dat')))
labels=RegressionLabels(CSVFile(os.path.join(SHOGUN_DATA_DIR, 'uci/housing/housing_label.dat')))
preproc=RescaleFeatures()
preproc.init(feats)
feats.add_preprocessor(preproc)
feats.apply_preprocessor(True)
#Regression models
ls=LeastSquaresRegression(feats, labels)
tau=1
rr=LinearRidgeRegression(tau, feats, labels)
width=1
tau=1
kernel=GaussianKernel(feats, feats, width)
kernel.set_normalizer(SqrtDiagKernelNormalizer())
krr=KernelRidgeRegression(tau, kernel, labels)
regression_models=[ls, rr, krr]
# -
# Let us use cross-validation to compare various values of tau paramter for ridge regression ([Regression notebook](http://www.shogun-toolbox.org/static/notebook/current/Regression.html)). We will use [MeanSquaredError](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CMeanSquaredError.html) as the performance metric. Note that normal splitting is used since it might be impossible to generate "good" splits using Stratified splitting in case of regression since we have continous values for labels.
# +
n=30
taus = logspace(-4, 1, n)
#5-fold cross-validation
k=5
split=CrossValidationSplitting(labels, k)
metric=MeanSquaredError()
cross=CrossValidation(rr, feats, labels, split, metric)
cross.put('num_runs', 50)
errors=[]
for tau in taus:
#set necessary parameter
rr.put('tau', tau)
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
#Enlist mean error for all runs
errors.append(result.get_mean())
figure(figsize=(20,6))
suptitle("Finding best (tau) parameter using cross-validation", fontsize=12)
p=subplot(121)
title("Ridge Regression")
plot(taus, errors, linewidth=3)
p.set_xscale('log')
p.set_ylim([0, 80])
xlabel("Taus")
ylabel("Mean Squared Error")
cross=CrossValidation(krr, feats, labels, split, metric)
cross.put('num_runs', 50)
errors=[]
for tau in taus:
krr.put('tau', tau)
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
#print tau, "error", result.get_mean()
errors.append(result.get_mean())
p2=subplot(122)
title("Kernel Ridge regression")
plot(taus, errors, linewidth=3)
p2.set_xscale('log')
xlabel("Taus")
_=ylabel("Mean Squared Error")
# -
# A low value of error certifies a good pick for the tau paramter which should be easy to conclude from the plots. In case of Ridge Regression the value of tau i.e. the amount of regularization doesn't seem to matter but does seem to in case of Kernel Ridge Regression. One interpretation of this could be the lack of over fitting in the feature space for ridge regression and the occurence of over fitting in the new kernel space in which Kernel Ridge Regression operates. </br> Next we will compare a range of values for the width of [Gaussian Kernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGaussianKernel.html) used in [Kernel Ridge Regression](http://shogun-toolbox.org/doc/en/latest/classshogun_1_1CKernelRidgeRegression.html)
# +
n=50
widths=logspace(-2, 3, n)
krr.put('tau', 0.1)
metric=MeanSquaredError()
k=5
split=CrossValidationSplitting(labels, k)
cross=CrossValidation(krr, feats, labels, split, metric)
cross.put('num_runs', 10)
errors=[]
for width in widths:
kernel.set_width(width)
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
#print width, "error", result.get_mean()
errors.append(result.get_mean())
figure(figsize=(15,5))
p=subplot(121)
title("Finding best width using cross-validation")
plot(widths, errors, linewidth=3)
p.set_xscale('log')
xlabel("Widths")
_=ylabel("Mean Squared Error")
# -
# The values for the kernel parameter and tau may not be independent of each other, so the values we have may not be optimal. A brute force way to do this would be to try all the pairs of these values but it is only feasible for a low number of parameters.
# +
n=40
taus = logspace(-3, 0, n)
widths=logspace(-1, 4, n)
cross=CrossValidation(krr, feats, labels, split, metric)
cross.put('num_runs', 1)
x, y=meshgrid(taus, widths)
grid=array((ravel(x), ravel(y)))
print(grid.shape)
errors=[]
for i in range(0, n*n):
krr.put('tau', grid[:,i][0])
kernel.set_width(grid[:,i][1])
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
errors.append(result.get_real('mean'))
errors=array(errors).reshape((n, n))
# +
from mpl_toolkits.mplot3d import Axes3D
#taus = logspace(0.5, 1, n)
jet()
fig=figure(figsize(15,7))
ax=subplot(121)
c=pcolor(x, y, errors)
_=contour(x, y, errors, linewidths=1, colors='black')
_=colorbar(c)
xlabel('Taus')
ylabel('Widths')
ax.set_xscale('log')
ax.set_yscale('log')
ax1=fig.add_subplot(122, projection='3d')
ax1.plot_wireframe(log10(y),log10(x), errors, linewidths=2, alpha=0.6)
ax1.view_init(30,-40)
xlabel('Taus')
ylabel('Widths')
_=ax1.set_zlabel('Error')
# -
# Let us approximately pick the good parameters using the plots. Now that we have the best parameters, let us compare the various regression models on the data set.
# +
#use the best parameters
rr.put('tau', 1)
krr.put('tau', 0.05)
kernel.set_width(2)
title_='Performance on Boston Housing dataset'
print("%50s" %title_)
for machine in regression_models:
metric=MeanSquaredError()
cross=CrossValidation(machine, feats, labels, split, metric)
cross.put('num_runs', 25)
result=cross.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
print("-"*80)
print("|", "%30s" % machine.get_name(),"|", "%20s" %metric.get_name(),"|","%20s" %result.get_mean() ,"|" )
print("-"*80)
# -
# ### Model selection using Grid Search
# A standard way of selecting the best parameters of a learning algorithm is by Grid Search. This is done by an exhaustive search of a specified parameter space. [CModelSelectionParameters](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CModelSelectionParameters.html) is used to select various parameters and their ranges to be used for model selection. A tree like structure is used where the nodes can be [CSGObject](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CSGObject.html) or the parameters to the object. The range of values to be searched for the parameters is set using `build_values()` method.
# +
#Root
param_tree_root=ModelSelectionParameters()
#Parameter tau
tau=ModelSelectionParameters("tau")
param_tree_root.append_child(tau)
# also R_LINEAR/R_LOG is available as type
min_value=0.01
max_value=1
type_=R_LINEAR
step=0.05
base=2
tau.build_values(min_value, max_value, type_, step, base)
# -
# Next we will create [CModelSelectionParameters](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CModelSelectionParameters.html) instance with a kernel object which has to be appended the root node. The kernel object itself will be append with a kernel width parameter which is the parameter we wish to search.
# +
#kernel object
param_gaussian_kernel=ModelSelectionParameters("kernel", kernel)
gaussian_kernel_width=ModelSelectionParameters("log_width")
gaussian_kernel_width.build_values(0.1, 6.0, R_LINEAR, 0.5, 2.0)
#kernel parameter
param_gaussian_kernel.append_child(gaussian_kernel_width)
param_tree_root.append_child(param_gaussian_kernel)
# -
# cross validation instance used
cross_validation=CrossValidation(krr, feats, labels, split, metric)
cross_validation.put('num_runs', 1)
# model selection instance
model_selection=GridSearchModelSelection(cross_validation, param_tree_root)
# +
print_state=False
# TODO: enable it once crossval has been fixed
#best_parameters=model_selection.select_model(print_state)
#best_parameters.apply_to_machine(krr)
#best_parameters.print_tree()
result=cross_validation.evaluate()
result=CrossValidationResult.obtain_from_generic(result)
print('Error with Best parameters:', result.get_mean())
# -
# The error value using the parameters obtained from Grid Search is pretty close (and should be better) to the one we had seen in the last section. Grid search suffers from the [curse of dimensionality](http://en.wikipedia.org/wiki/Curse_of_dimensionality) though, which can lead to huge computation costs in higher dimensions.
# ## References
# [1] <NAME>. and <NAME>. (2009). Apples-to-apples in cross-validation studies: Pitfalls in classifier performance measurement. Technical report, HP Laboratories.
| doc/ipython-notebooks/evaluation/xval_modelselection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# Ideas for processing MIT Airline Data Project data:
# * Could be done as a class that processes the SEC data spreadsheet and returns an object where you can call different parts of the spreadsheet
# * look for places where the data has already been processed into easier to handle
#
#
#
# http://web.mit.edu/airlinedata/www/Revenue&Related.html
#
#
# +
jetblueProfit_dat = pd.read_excel('JetBlue Airways Corp.xls', sheet_name='Sheet1', header=1)
jetblueProfit_dat
# -
# create column names for income statement DF
indexTry = list(jetblueProfit_dat.columns[1:])
indexTry.insert(0,jetblueProfit_dat.iloc[0,0])
| AdvStrat_Jupyter/ADP Data Processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:test1]
# language: python
# name: conda-env-test1-py
# ---
# ## Introduction
#
# In this notebook we demonstrate how to use previously computed spectra to create new gold-standard spectra which can be used to categorize the quality of new samples.
import PaSDqc
freq, nd, sample_list = PaSDqc.extra_tools.mk_ndarray("../data/gold_standard/")
sample_list
labels = ['bad', 'bad', 'bad', 'good', 'good', 'good', 'good']
cat_spec = PaSDqc.extra_tools.mk_categorical_spectra(freq, nd, labels)
cat_spec.to_csv("categorical_spectra_Qiagen_heat_1x.txt", sep="\t")
# These new categorical spectra could be used by supplying the command line option ```-c /path/to/categorical_spectra_Qiagen_heat_1x.txt``` to PaSDqc
| examples/08_example_make_gold_standard_spectra/Gold_standard_spectra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
with open ('style.css', 'r') as file:
css = file.read()
HTML(css)
# # Computing with Unlimited Precision
# *Python* provides the module <tt>fractions</tt> that implements *rational numbers* through the function <tt>Fraction</tt> that is implemented in this module. We can load this function as follows:
from fractions import Fraction
# The function <tt>Fraction</tt> expects two arguments, the *nominator* and the *denominator*. Mathematically, we have
# $$ \texttt{Fraction}(p, q) = \frac{p}{q}. $$
# For example, we can compute the sum $\frac{1}{2} + \frac{1}{3}$ as follows:
sum = Fraction(1, 2) + Fraction(1, 3)
print(sum)
1/2 + 1/3
# Let us compute <a href="https://en.wikipedia.org/wiki/E_(mathematical_constant)">Euler's number</a> $e$. The easiest way to compute $e$ is as inifinite series. We have that
# $$ e = \sum\limits_{n=0}^\infty \frac{1}{n!}. $$
# Here $n!$ denotes the *factorial* of $n$, which is defined as follows:
# $$ n! = 1 \cdot 2 \cdot 3 \cdot {\dots} \cdot n. $$
# The function `factorial` takes a natural number `n` and returns `n!`.
def factorial(n):
"returns the factorial of n"
result = 1
for i in range(1, n+1):
result *= i
return result
# Let's check that our definition of the factorial works as expected.
for i in range(10):
print(i, '! = ', factorial(i), sep='')
# Lets approximate $e$ by the following sum:
# $$ e = \sum\limits_{i=0}^n \frac{1}{i!} $$
# Setting $n=100$ should be sufficient to compute $e$ to a hundred decimal places.
n = 100
e = 0
for i in range(n+1):
e += Fraction(1, factorial(i))
print(e)
# As a fraction, that result is not helpful. Let us convert it into a floating point representation by
# multiply $e$ by $10^{100}$ and rounding so that we get the first 100 decimal places of $e$:
eTimesBig = e * 10 ** n
eTimesBig
s = str(round(eTimesBig))
s
# Insert a '.' after the first digit:
print(s[0], '.', s[1:], sep='')
| Python/Unlimited-Precision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Here we'll import our Turtle object. This lets us create specific turtles.
from mobilechelonian import Turtle
help(Turtle)
# We'll create a new turtle like this. It should open a window with our turtle inside.
t = Turtle()
# ## Controlling your turtle
# You can control your turtle's speed, position, and whether it draws a line using a few extra commands.
#
# Play around with different values below to control the turtle above.
# t.speed(N) will control the speed of the turtle when it moves (N from 1 - 10)
t.speed(1)
# t.penup() will "raise the pen" meaning the turtle won't draw lines anymore
t.penup()
# And this will "lower the pen" so the turtle draws when it moves
t.pendown()
# This changes the color of the turtle's path, if the pen is down
t.pencolor("red") # try "blue", "yellow", "brown", "black", "purple", "green"
# This will rotate the turtle counter-clockwise
t.left(90)
# And this rotates the turtle clockwise
t.right(90)
# This makes the turtle move forward
t.forward(100)
# Finally, this will bring the turtle back home (to the center)
t.home()
# Try drawing different shapes.
#
# # Keeping track of the turtle's path
# You can see the history of the turtle's position here
t.points
| 4.programmer/resources/tp3/try.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import stanfordnlp
model_dir='/pi/ai/corenlp'
nlp_zh = stanfordnlp.Pipeline(models_dir=model_dir,
lang='zh',
treebank='zh_gsd')
nlp_ja = stanfordnlp.Pipeline(models_dir=model_dir,
lang='ja',
treebank='ja_gsd')
# +
def analyse(sents, nlp):
doc = nlp(sents)
print(*[f'text: {word.text+" "}\tlemma: {word.lemma}\tupos: {word.upos}\txpos: {word.xpos}' for sent in doc.sentences for word in sent.words], sep='\n')
doc.sentences[0].print_dependencies()
# sents='我是一个中学生。'
# sents='我是一個高中生。'
sents='我是一個高中生'
analyse(sents, nlp_zh)
# -
sents='私は中学生です。'
analyse(sents, nlp_ja)
analyse('私は高校生です。', nlp_ja)
from sagas.nlu.corenlp_helper import CoreNlp, CoreNlpViz, langs
nlp_ja=langs['ja']()
ana=lambda sents: CoreNlpViz().analyse(sents, nlp_ja)
ana('私は高校生です。')
ana('私はドイツ出身です')
ana('b')
ana('私たちはすべてを可能にします。')
import sagas
sagas.dia('ja').ana_s('私たちはすべてを可能にします。')
sagas.dia('en').ana_s('it is a cat')
sagas.dia('zh').ana_s('達沃斯世界經濟論壇是每年全球政商界領袖聚在一起的年度盛事。')
import random
random.randint(0,1)
from sagas.nlu.uni_viz import viz
viz('I am a student', 'en')
viz('it is a cat', 'en')
| notebook/procs-stanfordnlp-ja.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Using code from <NAME> et al.
# https://nbviewer.jupyter.org/github/barbagroup/AeroPython/blob/master/lessons/11_Lesson11_vortexSourcePanelMethod.ipynb
# +
# import libraries and modules needed
import os
import numpy
from scipy import integrate, linalg
from matplotlib import pyplot
# integrate plots into the notebook
# %matplotlib inline
# -
# load geometry from data file
naca_filepath = os.path.join('naca2412.dat')
with open(naca_filepath, 'r') as infile:
x, y = numpy.loadtxt(infile, dtype=float, unpack=True)
# plot geometry
width = 10
pyplot.figure(figsize=(width, width))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.plot(x, y, color='k', linestyle='-', linewidth=2)
pyplot.axis('scaled', adjustable='box')
pyplot.xlim(-0.1, 1.1)
pyplot.ylim(-0.1, 0.1);
class Panel:
"""
Contains information related to a panel.
"""
def __init__(self, xa, ya, xb, yb):
"""
Initializes the panel.
Sets the end-points and calculates the center-point, length,
and angle (with the x-axis) of the panel.
Defines if the panel is located on the upper or lower surface of the geometry.
Initializes the source-strength, tangential velocity, and pressure coefficient
of the panel to zero.
Parameters
---------_
xa: float
x-coordinate of the first end-point.
ya: float
y-coordinate of the first end-point.
xb: float
x-coordinate of the second end-point.
yb: float
y-coordinate of the second end-point.
"""
self.xa, self.ya = xa, ya # panel starting-point
self.xb, self.yb = xb, yb # panel ending-point
self.xc, self.yc = (xa + xb) / 2, (ya + yb) / 2 # panel center
self.length = numpy.sqrt((xb - xa)**2 + (yb - ya)**2) # panel length
# orientation of panel (angle between x-axis and panel's normal)
if xb - xa <= 0.0:
self.beta = numpy.arccos((yb - ya) / self.length)
elif xb - xa > 0.0:
self.beta = numpy.pi + numpy.arccos(-(yb - ya) / self.length)
# panel location
if self.beta <= numpy.pi:
self.loc = 'upper' # upper surface
else:
self.loc = 'lower' # lower surface
self.sigma = 0.0 # source strength
self.vt = 0.0 # tangential velocity
self.cp = 0.0 # pressure coefficient
def define_panels(x, y, N=40):
"""
Discretizes the geometry into panels using 'cosine' method.
Parameters
----------
x: 1D array of floats
x-coordinate of the points defining the geometry.
y: 1D array of floats
y-coordinate of the points defining the geometry.
N: integer, optional
Number of panels;
default: 40.
Returns
-------
panels: 1D Numpy array of Panel objects.
The list of panels.
"""
R = (x.max() - x.min()) / 2.0 # circle radius
x_center = (x.max() + x.min()) / 2.0 # x-coordinate of circle center
theta = numpy.linspace(0.0, 2.0 * numpy.pi, N + 1) # array of angles
x_circle = x_center + R * numpy.cos(theta) # x-coordinates of circle
x_ends = numpy.copy(x_circle) # x-coordinate of panels end-points
y_ends = numpy.empty_like(x_ends) # y-coordinate of panels end-points
# extend coordinates to consider closed surface
x, y = numpy.append(x, x[0]), numpy.append(y, y[0])
# compute y-coordinate of end-points by projection
I = 0
for i in range(N):
while I < len(x) - 1:
if (x[I] <= x_ends[i] <= x[I + 1]) or (x[I + 1] <= x_ends[i] <= x[I]):
break
else:
I += 1
a = (y[I + 1] - y[I]) / (x[I + 1] - x[I])
b = y[I + 1] - a * x[I + 1]
y_ends[i] = a * x_ends[i] + b
y_ends[N] = y_ends[0]
# create panels
panels = numpy.empty(N, dtype=object)
for i in range(N):
panels[i] = Panel(x_ends[i], y_ends[i], x_ends[i + 1], y_ends[i + 1])
return panels
# discretize geoemetry into panels
panels = define_panels(x, y, N=100)
# plot discretized geometry
width = 10
pyplot.figure(figsize=(width, width))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.plot(x, y, color='k', linestyle='-', linewidth=2)
pyplot.plot(numpy.append([panel.xa for panel in panels], panels[0].xa),
numpy.append([panel.ya for panel in panels], panels[0].ya),
linestyle='-', linewidth=1, marker='o', markersize=6, color='#CD2305')
pyplot.axis('scaled', adjustable='box')
pyplot.xlim(-0.1, 1.1)
pyplot.ylim(-0.1, 0.1);
class Freestream:
"""
Freestream conditions.
"""
def __init__(self, u_inf=1.0, alpha=0.0):
"""
Sets the freestream speed and angle (in degrees).
Parameters
----------
u_inf: float, optional
Freestream speed;
default: 1.0.
alpha: float, optional
Angle of attack in degrees;
default 0.0.
"""
self.u_inf = u_inf
self.alpha = numpy.radians(alpha) # degrees to radians
# define freestream conditions
freestream = Freestream(u_inf=1.0, alpha=4.0)
def integral(x, y, panel, dxdk, dydk):
"""
Evaluates the contribution from a panel at a given point.
Parameters
----------
x: float
x-coordinate of the target point.
y: float
y-coordinate of the target point.
panel: Panel object
Panel whose contribution is evaluated.
dxdk: float
Value of the derivative of x in a certain direction.
dydk: float
Value of the derivative of y in a certain direction.
Returns
-------
Contribution from the panel at a given point (x, y).
"""
def integrand(s):
return (((x - (panel.xa - numpy.sin(panel.beta) * s)) * dxdk +
(y - (panel.ya + numpy.cos(panel.beta) * s)) * dydk) /
((x - (panel.xa - numpy.sin(panel.beta) * s))**2 +
(y - (panel.ya + numpy.cos(panel.beta) * s))**2) )
return integrate.quad(integrand, 0.0, panel.length)[0]
def source_contribution_normal(panels):
"""
Builds the source contribution matrix for the normal velocity.
Parameters
----------
panels: 1D array of Panel objects
List of panels.
Returns
-------
A: 2D Numpy array of floats
Source contribution matrix.
"""
A = numpy.empty((panels.size, panels.size), dtype=float)
# source contribution on a panel from itself
numpy.fill_diagonal(A, 0.5)
# source contribution on a panel from others
for i, panel_i in enumerate(panels):
for j, panel_j in enumerate(panels):
if i != j:
A[i, j] = 0.5 / numpy.pi * integral(panel_i.xc, panel_i.yc,
panel_j,
numpy.cos(panel_i.beta),
numpy.sin(panel_i.beta))
return A
def vortex_contribution_normal(panels):
"""
Builds the vortex contribution matrix for the normal velocity.
Parameters
----------
panels: 1D array of Panel objects
List of panels.
Returns
-------
A: 2D Numpy array of floats
Vortex contribution matrix.
"""
A = numpy.empty((panels.size, panels.size), dtype=float)
# vortex contribution on a panel from itself
numpy.fill_diagonal(A, 0.0)
# vortex contribution on a panel from others
for i, panel_i in enumerate(panels):
for j, panel_j in enumerate(panels):
if i != j:
A[i, j] = -0.5 / numpy.pi * integral(panel_i.xc, panel_i.yc,
panel_j,
numpy.sin(panel_i.beta),
-numpy.cos(panel_i.beta))
return A
A_source = source_contribution_normal(panels)
B_vortex = vortex_contribution_normal(panels)
def kutta_condition(A_source, B_vortex):
"""
Builds the Kutta condition array.
Parameters
----------
A_source: 2D Numpy array of floats
Source contribution matrix for the normal velocity.
B_vortex: 2D Numpy array of floats
Vortex contribution matrix for the normal velocity.
Returns
-------
b: 1D Numpy array of floats
The left-hand side of the Kutta-condition equation.
"""
b = numpy.empty(A_source.shape[0] + 1, dtype=float)
# matrix of source contribution on tangential velocity
# is the same than
# matrix of vortex contribution on normal velocity
b[:-1] = B_vortex[0, :] + B_vortex[-1, :]
# matrix of vortex contribution on tangential velocity
# is the opposite of
# matrix of source contribution on normal velocity
b[-1] = - numpy.sum(A_source[0, :] + A_source[-1, :])
return b
def build_singularity_matrix(A_source, B_vortex):
"""
Builds the left-hand side matrix of the system
arising from source and vortex contributions.
Parameters
----------
A_source: 2D Numpy array of floats
Source contribution matrix for the normal velocity.
B_vortex: 2D Numpy array of floats
Vortex contribution matrix for the normal velocity.
Returns
-------
A: 2D Numpy array of floats
Matrix of the linear system.
"""
A = numpy.empty((A_source.shape[0] + 1, A_source.shape[1] + 1), dtype=float)
# source contribution matrix
A[:-1, :-1] = A_source
# vortex contribution array
A[:-1, -1] = numpy.sum(B_vortex, axis=1)
# Kutta condition array
A[-1, :] = kutta_condition(A_source, B_vortex)
return A
def build_freestream_rhs(panels, freestream):
"""
Builds the right-hand side of the system
arising from the freestream contribution.
Parameters
----------
panels: 1D array of Panel objects
List of panels.
freestream: Freestream object
Freestream conditions.
Returns
-------
b: 1D Numpy array of floats
Freestream contribution on each panel and on the Kutta condition.
"""
b = numpy.empty(panels.size + 1, dtype=float)
# freestream contribution on each panel
for i, panel in enumerate(panels):
b[i] = -freestream.u_inf * numpy.cos(freestream.alpha - panel.beta)
# freestream contribution on the Kutta condition
b[-1] = -freestream.u_inf * (numpy.sin(freestream.alpha - panels[0].beta) +
numpy.sin(freestream.alpha - panels[-1].beta) )
return b
A = build_singularity_matrix(A_source, B_vortex)
b = build_freestream_rhs(panels, freestream)
# +
# solve for singularity strengths
strengths = numpy.linalg.solve(A, b)
# store source strength on each panel
for i , panel in enumerate(panels):
panel.sigma = strengths[i]
# store circulation density
gamma = strengths[-1]
# -
def compute_tangential_velocity(panels, freestream, gamma, A_source, B_vortex):
"""
Computes the tangential surface velocity.
Parameters
----------
panels: 1D array of Panel objects
List of panels.
freestream: Freestream object
Freestream conditions.
gamma: float
Circulation density.
A_source: 2D Numpy array of floats
Source contribution matrix for the normal velocity.
B_vortex: 2D Numpy array of floats
Vortex contribution matrix for the normal velocity.
"""
A = numpy.empty((panels.size, panels.size + 1), dtype=float)
# matrix of source contribution on tangential velocity
# is the same than
# matrix of vortex contribution on normal velocity
A[:, :-1] = B_vortex
# matrix of vortex contribution on tangential velocity
# is the opposite of
# matrix of source contribution on normal velocity
A[:, -1] = -numpy.sum(A_source, axis=1)
# freestream contribution
b = freestream.u_inf * numpy.sin([freestream.alpha - panel.beta
for panel in panels])
strengths = numpy.append([panel.sigma for panel in panels], gamma)
tangential_velocities = numpy.dot(A, strengths) + b
for i, panel in enumerate(panels):
panel.vt = tangential_velocities[i]
# tangential velocity at each panel center.
compute_tangential_velocity(panels, freestream, gamma, A_source, B_vortex)
def compute_pressure_coefficient(panels, freestream):
"""
Computes the surface pressure coefficients.
Parameters
----------
panels: 1D array of Panel objects
List of panels.
freestream: Freestream object
Freestream conditions.
"""
for panel in panels:
panel.cp = 1.0 - (panel.vt / freestream.u_inf)**2
# surface pressure coefficient
compute_pressure_coefficient(panels, freestream)
# plot surface pressure coefficient
pyplot.figure(figsize=(10, 6))
pyplot.grid()
pyplot.xlabel('$x$', fontsize=16)
pyplot.ylabel('$C_p$', fontsize=16)
pyplot.plot([panel.xc for panel in panels if panel.loc == 'upper'],
[panel.cp for panel in panels if panel.loc == 'upper'],
label='upper surface',
color='r', linestyle='-', linewidth=2, marker='o', markersize=6)
pyplot.plot([panel.xc for panel in panels if panel.loc == 'lower'],
[panel.cp for panel in panels if panel.loc == 'lower'],
label= 'lower surface',
color='b', linestyle='-', linewidth=1, marker='o', markersize=6)
pyplot.legend(loc='best', prop={'size':16})
pyplot.xlim(-0.1, 1.1)
pyplot.ylim(1.0, -2.0)
pyplot.title('Number of panels: {}'.format(panels.size), fontsize=16);
# calculate the accuracy
accuracy = sum([panel.sigma * panel.length for panel in panels])
print('sum of singularity strengths: {:0.6f}'.format(accuracy))
# compute the chord and lift coefficient
c = abs(max(panel.xa for panel in panels) -
min(panel.xa for panel in panels))
cl = (gamma * sum(panel.length for panel in panels) /
(0.5 * freestream.u_inf * c))
print('lift coefficient: CL = {:0.3f}'.format(cl))
| basic_source_vortex_method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
'''This holds the machinery for making measurment equations for a given antenna layout and pointing and sky brightness.
Uses the linear equation formulation.'''
import numpy as np
import multiprocessing as mp
from itertools import product
import astropy.coordinates as ac
import astropy.units as au
import astropy.time as at
from Geometry import itrs2uvw
def vec(A):
'''Stack columns aka use F ordering (col-major)'''
return np.ndarray.flatten(A,order='F')
def reshape(v,shape):
return np.ndarray.reshape(v,shape,order='F')
def khatrirao(A,B,out=None):
if len(A.shape) != len(B.shape):
print("Not same size")
return
if A.shape[0] != B.shape[0]:
print("Not same number of rows")
return
if A.shape[1] != B.shape[1]:
print("Not same number of columns")
return
if out is not None:
if out.shape[0] == A.shape[0] and out.shape[1] == A.shape[0]*A.shape[1]:
K = out
else:
print("out matrix not right size")
return
else:
K = np.zeros([A.shape[0]*B.shape[0],A.shape[1]],dtype=type(1j))
n = 0
while n<A.shape[1]:
#print A[:,n].shape,B[:,n].shape,K[:,n*A.shape[0]:(n+1)*A.shape[0]].shape
K[:,n:n+1] = np.kron(A[:,n],B[:,n]).reshape([A.shape[0]*B.shape[0],1])
#K[:,n:n+1] = reshape(np.outer(A[:,n],B[:,n])[A.shape[0]*B.shape[0],1])
n += 1
return K
def createPointing(lvec,mvec):
'''returns the pointings of all combinations of l,m in l m vec'''
L,M = np.meshgrid(lvec,mvec)
N = np.sqrt(1-L**2-M**2)
P = np.array([vec(L),vec(M),vec(N)])
return P
def phaseTracking(ra,dec,obsLoc,obsTime):
s = ac.SkyCoord(ra=ra*au.deg,dec=dec*au.deg,frame='icrs')
frame = ac.AltAz(location = obsLoc, obstime = obsTime, pressure=None, copy = True)
s_ = s.transform_to(frame)
return s_
def responseMatrix(Z,P):
'''Returns the array response matrix, where each column is the array response vector:
Z =[x_1,...,x_N] are antenna locations in lambda in uvw coordinates
P = [p_1,...,p_Q] where each p_i is (l_i,m_i,sqrt(1-l_i^2-m_i^2))
a_q = exp(-1j*Z^T.p_q)
'''
#Q = P.shape[1]#number of pointings
#N = X.shape[1]#number of antennas
A = np.exp(-1j*2*np.pi*Z.transpose().dot(P))
return A
def measurementEquationMatrix(skyBrightness,X,P,noiseCovariance=None):
if np.size(skyBrightness) != np.size(P)/3:
print("Not equal components")
return
if noiseCovariance is not None:
if np.size(noiseCovariance) == 1:
sigmaN = np.eye(X.shape[1])*noiseCovariance
else:
if np.size(noiseCovariance) == np.size(skyBrightness):
sigmaN = np.diag(noiseCovariance)
else:
print("sigma sizes not equal")
return
else:
sigmaN = 0.
sigmaS = np.diag(skyBrightness)
A = responseMatrix(X,P)
R = A.dot(sigmaS).dot(A.conjugate().transpose()) + sigmaN
return R
def measurementEquationVector(skyBrightness,X,P,noiseCovariance=None):
if np.size(skyBrightness) != P.shape[1]:
print("Not equal components")
return
sigmaS = skyBrightness
A = responseMatrix(X,P)
r = np.zeros(A.shape[0]*A.shape[0],dtype=type(1j))
n = 0
#pool = mp.Pool(4)
#print pool
#r = pool.map(lambda p: np.kron(p[0].conjugate(),p[0])*p[1],product(list(A.transpose()),list(sigmaS)))
#print r
while n<A.shape[1]:
#print A[:,n].shape,B[:,n].shape,K[:,n*A.shape[0]:(n+1)*A.shape[0]].shape
r += np.kron(A[:,n],A[:,n])*sigmaS[n]
#r += vec(np.outer(A[:,n],A[:,n]))*sigmaS[n]
n += 1
#Ms = khatrirao(A.conjugate(),A)
if noiseCovariance is not None:
if np.size(noiseCovariance) == np.size(skyBrightness):
sigmaN = noiseCovariance
Eye = np.eye(np.size(skyBrightness))
Mn = khatrirao(Eye,Eye)
else:
if np.size(noiseCovariance) == 1:
sigmaN = np.ones_like(skyBrightness)*noiseCovariance
Mn = vec(np.eye(np.size(skyBrightness)))
else:
print("sigma sizes not equal")
return
r += Mn.dot(sigmaN)
return r
def catalog2ms(simTk,catalogName):
#load catalog
cat = np.genfromtxt(catalogName,comments='#',skip_header=5,names=True)
print (cat)
ra = cat['RA']
dec = cat['DEC']
skyBrightness = cat['Total_flux']
if np.size(ra) == 1:
ra = np.array([ra])
dec = np.array([dec])
skyBrightness = np.array([skyBrightness])
#Maj = cat['Maj']
noise = np.mean(cat['E_Total_flux'])
#pointings of sources in catalog
P = ac.SkyCoord(ra=ra*au.deg,dec = dec*au.deg,frame='icrs')
# RadioArray with all frame info
radioArray = simTk.radioArray
#compare with ms 0-1 baseline at first timestamp, make sure fieldId is correct in json
print(radioArray.baselines[:,0,1,:])
s = radioArray.pointing
lon = radioArray.center.earth_location.geodetic[0].rad
lat = radioArray.center.earth_location.geodetic[1].rad
#response matrix, visibilities
R = np.zeros([len(simTk.timeSlices),radioArray.Nantenna,radioArray.Nantenna],dtype=type(1j))
plotBl = []
times = []
i = 0
while i < len(simTk.timeSlices):
#print('Calculating visibilities for: {0}'.format(simTk.timeSlices[i]))
lmst = simTk.timeSlices[i].sidereal_time('mean',lon)
hourangle = lmst.to(au.rad).value - s.ra.rad
Ruvw = itrs2uvw(hourangle,s.dec.rad,lon,lat)
#l,m,n 3xM
responseDirections = Ruvw.dot(P.itrs.cartesian.xyz)
#uvw in lambda
X = radioArray.antennaLocs[i,:,:].transpose()/simTk.wavelength
R[i,:,:] = measurementEquationMatrix(skyBrightness,X,responseDirections,noiseCovariance=noise)
times.append(simTk.timeSlices[i].gps)
plotBl.append(R[i,0,1])
#print R[i,0,:]
i += 1
import pylab as plt
#baselines = np.sqrt(np.sum(radioArray.baselines[i,:,1,:]**2,axis=1))
plt.plot(times,np.abs(plotBl))
#plt.title("UV:{0}".format(P.itrs.cartesian.xyz[]))
plt.show()
if __name__=='__main__':
N = 5*10
M = 25
X = np.random.uniform(size=[3,N])
s = np.array([0,0,1])
lvec = np.linspace(-1./np.sqrt(2),1./np.sqrt(2),int(np.sqrt(M)))
mvec = np.copy(lvec)
P = createPointing(lvec,mvec)
wavelength = 1
skyBrightness = np.random.uniform(size=M)
#R = measurementEquationMatrix(skyBrightness,X,s,P,wavelength)
#r = measurementEquationVector(skyBrightness,X,s,P,wavelength)
#print r,R
obsTime = at.Time('2016-12-31T00:00:00.000',format='isot',scale='utc')
obsLoc = ac.ITRS(x=0*au.m,y=0*au.m,z=0*au.m)#stick your location here
from SimulationToolkit import Simulation
simTk = Simulation(simConfigJson='SimulationConfig.json',logFile='logs/0.log')
#catalogName = '/net/para34/data1/albert/casa/images/plckg004-19_150.fits.gaul'
catalogName = 'test.gaul'
catalog2ms(simTk,catalogName)
| src/ionotomo/notebooks/MeasurmentEquation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# name: python3
# ---
# !which python
import csv
import numpy as np
from collections import Counter
from nltk.corpus import brown
from mittens import GloVe, Mittens
from sklearn.feature_extraction import _stop_words
from sklearn.feature_extraction.text import CountVectorizer
import pickle
import re
import json
import os
import pandas as pd
import spacy
import nltk
from nltk.tokenize import sent_tokenize, word_tokenize
# +
def glove2dict(glove_filename):
with open(glove_filename, encoding='utf-8') as f:
reader = csv.reader(f, delimiter=' ', quoting=csv.QUOTE_NONE)
embed = {line[0]: np.array(list(map(float, line[1:])))
for line in reader}
return embed
def get_rareoov(xdict, val):
return [k for (k,v) in Counter(xdict).items() if v<=val]
# -
# # Load Data
# +
# Load the orignial dataset to train glove. Larger dataset can give better results for word embedding
# NOTE: For current fast development use the whole_filtered dataset.
# Later should change this to whole_cleaned dataset.
dir_path = '../Dataset/wine/'
# Load train dataset
train_review = []
cnt = 0
file_path = os.path.join(dir_path, 'whole_filtered.json')
with open(file_path) as f:
for line in f:
line_data = json.loads(line)
user_id = line_data['user']
item_id = line_data['item']
rating = line_data['rating']
review = line_data['review']
train_review.append([item_id, user_id, rating, review])
cnt += 1
if cnt % 100000 == 0:
print('{} lines loaded.'.format(cnt))
print('Finish loading train dataset, totally {} lines.'.format(len(train_review)))
# -
df_train_data = pd.DataFrame(train_review, columns=['item', 'user', 'rating', 'review'])
train_review_doc = list(df_train_data['review'])
len(train_review_doc)
with open('./embeddings/wine/train-review.txt', 'w') as f:
cnt = 0
for review in train_review_doc:
# write this into file
f.write(review.strip())
f.write('\n')
cnt += 1
print('Write {} lines of review into file'.format(cnt))
# +
from collections import Counter
import simplejson
from nltk.tokenize import word_tokenize
import string
import re
import nltk
nltk.download('punkt')
def tokenize(infname="embeddings/wine/train-review.txt"):
outfname = open(infname + ".tokenized", "w")
sents = open(infname).read().split("\n")
for sent in sents:
# convert characters into lower cases
text = sent.lower()
# tokenize the raw text using nltk.tokenize
text = word_tokenize(text)
# remove all punctuation
text = list(filter(lambda token: token not in string.punctuation, text))
# # replace all numbers with a special token <num>
# text = [re.sub('[0-9]+','<num>', token) for token in text]
# write this processed text into the output file
simplejson.dump(text, outfname)
outfname.write("\n")
outfname.close()
def token_filter(infname="embeddings/wine/train-review.txt.tokenized", thresh=5):
outfname = open(infname.replace(".tokenized", ".filtered"), 'w')
vocab = []
vocab_cnt = Counter()
tokenized_sents = []
with open(infname) as f_in:
for line in f_in:
tokenized_text = simplejson.loads(line)
vocab_cnt.update(tokenized_text)
tokenized_sents.append(tokenized_text)
for tokenized_sent in tokenized_sents:
# filter word that has less than 5 TF over the whole dataset
filtered_text = []
for token in tokenized_sent:
if vocab_cnt[token] < thresh:
pass
else:
filtered_text.append(token)
# write this filtered data into file
# ignore the last line of the initial imdb-small dataset since it's an empty line
if len(tokenized_sent) > 0:
new_sentence = " ".join(filtered_text).strip()+'\n'
outfname.write(new_sentence)
# bulld the vocabulary
## filter out token with low TF
vocab_cnt_filter = {token: cnt for token, cnt in vocab_cnt.items() if cnt >= thresh}
vocab_cnt_filter = Counter(vocab_cnt_filter)
## convert this vocabulary counter to a list
for token,cnt in vocab_cnt_filter.most_common():
vocab.append(token)
outfname.close()
return vocab
# -
tokenize()
vocab = token_filter()
print("The vocab size = {}".format(len(vocab)))
# ## Load Feature Words
dataset_name = 'wine'
feature_2_id_file = '../Dataset/{}/train/feature/feature2id.json'.format(dataset_name)
with open(feature_2_id_file, 'r') as f:
print("Load file: {}".format(feature_2_id_file))
feature_vocab = json.load(f)
for fea_word in list(feature_vocab.keys()):
if fea_word in vocab:
pass
else:
print(fea_word)
print("Length of feature vocab: {}".format(len(feature_vocab)))
total_vocab = set(vocab) | set(list(feature_vocab.keys()))
print("The total vocab size = {}".format(len(total_vocab)))
total_vocab_list = list(total_vocab)
cv = CountVectorizer(ngram_range=(1,1), vocabulary=total_vocab_list)
X = cv.fit_transform(train_review_doc)
Xc = (X.T * X)
Xc.setdiag(0)
coocc_ar = Xc.toarray()
coocc_ar.shape
X.shape
len(cv.vocabulary_)
train_vocab = cv.get_feature_names()
print(train_vocab[:20])
cv.vocabulary_["aroma"]
train_vocab == total_vocab_list
# +
from mittens import GloVe
import numpy as np
# mittens_model = Mittens(n=256, max_iter=1000)
# new_embeddings = mittens_model.fit(
# coocc_ar,
# vocab=train_vocab)
# NOTE: For fast development, set the max_iter to be 100.
# Should set it to be 1000 for more accurate word embedding.
glove_model = GloVe(n=256, max_iter=100)
embeddings = glove_model.fit(coocc_ar)
# -
embeddings.shape
len(cv.vocabulary_)
len(cv.get_feature_names())
vocabulary_words = cv.get_feature_names()
print(list(feature_vocab.keys())[:20])
print(vocabulary_words[:20])
# construct an embedding dict which has a format of {'word':'wine', 'embedding': array([...])}
word_embedding_dict = dict()
for i in range(len(embeddings)):
current_word = vocabulary_words[i]
current_embedding = embeddings[i]
word_embedding_dict[current_word] = current_embedding
print("Totally {} words begin added in the word embedding dict".format(len(word_embedding_dict)))
# Save the Embedding dict into pickle file
import pickle
with open('./embeddings/wine/glove.pickle', 'wb') as handle:
pickle.dump(word_embedding_dict, handle, protocol=pickle.HIGHEST_PROTOCOL)
# extract the embedding vector for each feature word
feature_embedding = dict()
emb_cnt = 0
for feature_word in list(feature_vocab.keys()):
feature_word_id = cv.vocabulary_[feature_word]
feature_vocab_id = feature_vocab[feature_word]
feature_word_emb = embeddings[feature_word_id]
feature_embedding[feature_vocab_id] = feature_word_emb
emb_cnt += 1
print("Totally {} feature words".format(emb_cnt))
# extract the embedding vector for each feature word
feature_embed_dict = dict()
for emb_chunk in list(feature_embedding.items()):
feature_id = emb_chunk[0] # this is a str
feature_emb_np = emb_chunk[1].tolist()
feature_embed_dict[feature_id] = feature_emb_np
len(feature_embed_dict)
feature_emb_filepath = '../Dataset/wine/train/feature/featureid2embedding.json'
with open(feature_emb_filepath, 'w') as f:
print("Write file: {}".format(feature_emb_filepath))
json.dump(feature_embed_dict, f)
| 5-FineTuneGloveFeatureWords_wine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from hcipy import *
import numpy as np
from matplotlib import pyplot as plt
N = 64
D = 9.96
aperture = circular_aperture(D)
pupil_grid = make_pupil_grid(N, D)
sps = 40 * N // 128
pupsep = 1
outgrid_size = int(np.ceil(sps * (pupsep + 1)))
D_grid = 3.6e-3
pyramid_grid = make_pupil_grid(N, D_grid)
outgrid = np.zeros((N, N)).tolist()
buffer = (N - 2 * outgrid_size)//2
def aberration_basis(N):
wfref = Wavefront(aperture(pupil_grid))
wfref.electric_field.shape = (N, N)
wfrefl = wfref.electric_field.tolist()
aberration_mode_basis = []
for i in range(N):
for j in range(N):
l = np.zeros((N, N), dtype=complex)
if np.complex(wfrefl[i][j]) != 0:
l[i][j] = 1j
aberration_mode_basis.append(Wavefront(Field(np.round(np.asarray(l).ravel(), 3) * aperture(pupil_grid), wfref.grid)))
return aberration_mode_basis
basis = aberration_basis(N)
# +
imshow_field(basis[0].electric_field)
plt.show()
#okay so the basis is getting created correctly that's not the problem. Let's propagate each of these.
# +
def get_sub_images(electric_field):
pyramid_grid = make_pupil_grid(N, D_grid)
images = Field(np.asarray(electric_field).ravel(), pyramid_grid)
pysize = int(np.sqrt(images.size))
images.shape = (pysize, pysize)
sub_images = [images[pysize-sps-1:pysize-1, 0:sps], images[pysize-sps-1:pysize-1, pysize-sps-1:pysize-1],
images[0:sps, 0:sps], images[0:sps, pysize-sps-1:pysize-1]]
subimage_grid = make_pupil_grid(sps, D_grid * sps / N)
for count, img in enumerate(sub_images):
img = img.ravel()
img.grid = subimage_grid
sub_images[count] = img
return sub_images
def pyramid_prop(wf):
# Given a wavefront, returns the result of a pyramid propagation and splitting into sub-images,
# as a list of hcipy Field objects.
keck_pyramid = PyramidWavefrontSensorOptics(pupil_grid, pupil_separation=pupsep, num_pupil_pixels=sps)
return get_sub_images(keck_pyramid.forward(wf).electric_field)
def estimate(images_list):
EstimatorObject = PyramidWavefrontSensorEstimator(aperture, make_pupil_grid(sps*2, D_grid*sps*2/N))
I_b = images_list[0]
I_a = images_list[1]
I_c = images_list[2]
I_d = images_list[3]
norm = I_a + I_b + I_c + I_d
I_x = (I_a + I_b - I_c - I_d) / norm
I_y = (I_a - I_b - I_c + I_d) / norm
pygrid = make_pupil_grid(sps)
return Field(I_x.ravel(), pygrid), Field(I_y.ravel(), pygrid)
def make_slopes(wf):
x, y = estimate(pyramid_prop(wf))
return np.concatenate((x, y))
# -
slopes_basis = [make_slopes(basis_element) for basis_element in basis]
imshow_field(slopes_basis[0][0:sps*sps], make_pupil_grid(sps))
plt.show()
imshow_field(slopes_basis[0][sps*sps:2*sps*sps], make_pupil_grid(sps))
plt.show()
len(basis)
flat_slopes = make_slopes(Wavefront(aperture(pupil_grid)))
spsg = make_pupil_grid(sps)
flat_x = Field(flat_slopes[0:sps*sps], spsg)
flat_y = Field(flat_slopes[sps*sps:2*sps*sps], spsg)
def least_inv(A):
# given a matrix A such that Ax = b, makes a least-squares matrix Y such that
# x^ = Yb.
return np.linalg.inv(A.T.dot(A)).dot(A.T)
for i in range(10):
imshow_field(slopes_basis[i][0:sps*sps] - flat_x)
plt.show()
imshow_field(slopes_basis[sps-5][0:sps*sps] - slopes_basis[0][0:sps*sps], spsg)
plt.show()
# +
# let's do a simple test. Adding together the first two basis elements:
test = Wavefront(basis[0].electric_field + basis[sps-1].electric_field)
imshow_field(test.electric_field)
plt.show()
# +
# Cool. Now let's propagate this through the pyramid
test_slopes = make_slopes(test)
imshow_field(test_slopes[0:25], make_pupil_grid(5))
plt.show()
test_x = Field(test_slopes[0:25], make_pupil_grid(5))
test_y = Field(test_slopes[25:50], make_pupil_grid(5))
# +
# Now for some comparisons. First the x:
imshow_field(slopes_basis[0][0:25] + slopes_basis[sps-1][0:25], make_pupil_grid(5))
plt.show()
imshow_field(test_x)
plt.show()
# +
# Now the y:
imshow_field(slopes_basis[0][25:50] + slopes_basis[sps-1][25:50], make_pupil_grid(5))
plt.colorbar()
plt.show()
imshow_field(test_y)
plt.colorbar()
plt.show()
# +
# Weird. Linearity in x is fine, in y is not. Let's up the N.
| pwfs-keck/t/Simple PWFS Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Process Synonyms
#
# This notebook uses a combination of Python data science libraries and the Google Natural Language API (machine learning) to expand the vocabulary of the chatbot by generating synonyms for topics created in the previous notebook.
# + deletable=true editable=true
# !pip uninstall -y google-cloud-datastore
# + deletable=true editable=true
# !pip install google-cloud-datastore
# + deletable=true editable=true
# !pip install inflect
# -
# Hit Reset Session > Restart, then resume with the following cells.
# + deletable=true editable=true
# Only need to do this once...
import nltk
nltk.download('stopwords')
nltk.download('wordnet')
# + deletable=true editable=true
from nltk.corpus import stopwords
stop = set(stopwords.words('english'))
# + deletable=true editable=true
from google.cloud import datastore
# + deletable=true editable=true
datastore_client = datastore.Client()
# + deletable=true editable=true
client = datastore.Client()
query = client.query(kind='Topic')
results = list(query.fetch())
# + deletable=true editable=true
import inflect
plurals = inflect.engine()
# + [markdown] deletable=true editable=true
# ## Extract Synonyms with Python
# Split the topic into words and use PyDictionary to look up synonyms in a "thesaurus" for each word. Store these in Datastore and link them back to the topic. Note this section uses the concept of "stop words" to filter out articles and other parts of speech that don't contribute to meaning of the topic.
# + deletable=true editable=true
from nltk.corpus import wordnet
from sets import Set
for result in results:
for word in result.key.name.split():
if word in stop:
continue
synonyms = Set()
for syn in wordnet.synsets(word):
if ".n." in str(syn):
for l in syn.lemmas():
lemma = l.name()
if (lemma.isalpha()):
synonyms.add(lemma)
synonyms.add(plurals.plural(lemma))
if ".a." in str(syn):
synonyms = Set()
break
print result.key.name, word, synonyms
kind = 'Synonym'
synonym_key = datastore_client.key(kind, result.key.name)
synonym = datastore.Entity(key=synonym_key)
synonym['synonym'] = result.key.name
datastore_client.put(synonym)
synonym_key = datastore_client.key(kind, word)
synonym = datastore.Entity(key=synonym_key)
synonym['synonym'] = result.key.name
datastore_client.put(synonym)
for dictionary_synonym in synonyms:
synonym_key = datastore_client.key(kind, dictionary_synonym)
synonym = datastore.Entity(key=synonym_key)
synonym['synonym'] = result.key.name
datastore_client.put(synonym)
synonym_key = datastore_client.key(kind, plurals.plural(word))
synonym = datastore.Entity(key=synonym_key)
synonym['synonym'] = result.key.name
datastore_client.put(synonym)
# + deletable=true editable=true
| courses/dialogflow-chatbot/notebooks/ProcessSynonyms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib as plt
from sklearn import metrics
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
import tensorflow as tf
import datetime
import time
from datetime import timedelta
import os
# %matplotlib inline
# -
df = pd.read_csv(r"C:\Users\rg654th\Downloads\Flight Delay\Clean Data\Clean Historical Data.csv", parse_dates=[0], infer_datetime_format=True)
# New Column and Format
df['Scheduled DateTime'] = pd.to_datetime(df['Scheduled DateTime'], infer_datetime_format=True)
df['Delay? (15 min)'] = np.where(df['Minute Delay'] >= 15, 'Yes', 'No')
# df['Sched_mins_since_midnight']
df['Sched_mins_since_midnight'] = (df['Scheduled DateTime'] - df['Scheduled DateTime'].apply(lambda dt: dt.replace(hour=0, minute=0, second=0, microsecond=0)))
df['Sched_mins_since_midnight'] = df['Sched_mins_since_midnight'] / np.timedelta64(1, 'm')
traindf = df.drop(['Flight No.', 'Scheduled Date','Scheduled DateTime', 'Scheduled Time', 'Real Departure Time'], axis=1)
traindf = traindf.dropna()
traindf.shape
traindf.head(1)
newtraindf = pd.get_dummies(traindf)
newtraindf.head(1)
# +
# Split Data
x_data = newtraindf.drop(['Minute Delay', 'Delay Type_None', 'Delay Type_Short', 'Delay Type_Long', 'Delay Type_Medium', 'Delay? (15 min)_No', 'Delay? (15 min)_Yes'],axis=1)
y_data = np.vstack((newtraindf['Delay Type_Long'], newtraindf['Delay Type_Medium'], newtraindf['Delay Type_None'], newtraindf['Delay Type_Short'])).T
X_train, X_test, y_train, y_test = train_test_split(x_data,y_data,test_size=0.2)
n_samples, n_features = x_data.shape
n_outputs = y_data.shape[1]
n_classes = 4
# -
# SGD Regressor, Lasso, Elastic Net, Ridge Regression, SVR(kernel = 'linear'or'rbf'), Ensemble
from sklearn.ensemble import RandomForestClassifier
from sklearn.multioutput import MultiOutputClassifier
forest = RandomForestClassifier(n_estimators=100)
model = MultiOutputClassifier(forest, n_jobs=-1)
model.fit(X_train, y_train)
# +
y_pred = model.predict(X_test)
score2 = metrics.accuracy_score(y_test, y_pred)
print("Accuracy", "", score2)
# -
| Baseline Models/Model_Training Baseline Delay Type (too hard to predict).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import nltk
import re
nltk.download('punkt')
df = pd.read_csv("tennis_articles_v4.csv")
df.head
df['article_text'][0]
from nltk.tokenize import sent_tokenize
sentences = []
for st in df['article_text']:
sentences.append(sent_tokenize(st))
sentences
word_embeddings = {}
f = open('glove.6B.100d.txt', encoding='utf-8')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:],dtype='float32')
word_embeddings[word] = coefs
for article in sentences:
for i in range(len(article)):
article[i] = re.sub(r'[^a-zA-Z]', ' ', article[i])
article[i] = article[i].lower()
sentences
nltk.download('stopwords')
from nltk.corpus import stopwords
stopwords = stopwords.words('english')
stopwords
def remove_stopwords(sen):
ret_sen = ' '.join([word for word in sen if word not in stopwords])
return ret_sen
clean_senteces = []
for i in range(len(sentences)):
clean_senteces.extend(sentences[i])
for i in range(len(clean_senteces)):
clean_senteces[i] = remove_stopwords(clean_senteces[i].split(' '))
clean_senteces
word_embeddings = {}
f = open('glove.6B.100d.txt', encoding='utf-8')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
word_embeddings[word] = coefs
f.close()
sentece_vector = []
for i in clean_senteces:
if len(i)!=0:
v = sum([word_embeddings.get(w, np.zeros((100,))) for w in i.split()])/(len(i.split())+0.001)
else:
v = np.zeroes(100,)
sentece_vector.append(v)
sim_mat = np.zeros([len(sentences), len(sentences)])
sim_mat
from sklearn.metrics.pairwise import cosine_similarity
for i in range(len(sentences)):
for j in range(len(sentences)):
if i != j:
sim_mat[i][j] = cosine_similarity(sentece_vector[i].reshape(1,100), sentece_vector[i].reshape(1,100))[0,0]
sim_mat
import networkx as nx
nx_graph = nx.from_numpy_array(sim_mat)
scores = nx.pagerank(nx_graph)
scores
ranked_sentences = sorted(((scores[i],s) for i,s in enumerate(sentences)), reverse=True)
for i in range(10):
print(ranked_sentences[i][1])
| examples/TextRank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import sklearn
import pandas as pd
import os
import sys
import time
import tensorflow as tf
from tensorflow import keras
print(tf.__version__)
print(sys.version_info)
for module in mpl, np, pd, sklearn, tf, keras:
print(module.__name__, module.__version__)
# -
@tf.function(input_signature=[tf.TensorSpec([None], tf.int32, name='x')])
def cube(z):
return tf.pow(z, 3)
cube_func_int32 = cube.get_concrete_function(
tf.TensorSpec([None], tf.int32))
print(cube_func_int32)
print(cube_func_int32 is cube.get_concrete_function(
tf.TensorSpec([5], tf.int32)))
print(cube_func_int32 is cube.get_concrete_function(
tf.constant([1, 2, 3])))
cube_func_int32.graph
print(cube(tf.constant([1, 2, 3])))
to_export = tf.Module()
to_export.cube = cube
tf.saved_model.save(to_export, "./signature_to_saved_model")
# !saved_model_cli show --dir ./signature_to_saved_model --all
imported = tf.saved_model.load('./signature_to_saved_model')
imported.cube(tf.constant([2]))
| JupyterNotebookCode/signature_to_saved_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Zencity_env
# language: python
# name: zencity_env
# ---
# # Create Insights
# ### Imports and constants
import random
import pandas as pd
import ast
import datetime
from wordcloud import WordCloud
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from collections import Counter, OrderedDict
from transformers import AutoConfig, AutoTokenizer, AutoModel, pipeline
from summarizer import Summarizer
import warnings
warnings.filterwarnings("ignore")
# +
FILE_NAME = './results/cluster_results.csv'
CLUSTER_NUMBER = 1
# +
PREV = "Efforts to develop prophylaxis clinical studies and prioritize in healthcare workers"
THERAPEUTIC_1 = "Effectiveness of drugs being developed and tried to treat COVID-19 patients."
THERAPEUTIC_2 = "Clinical and bench trials to investigate less common viral inhibitors against COVID-19 such as naproxen, clarithromycin, and minocyclinethat that may exert effects on viral replication."
THERAPEUTIC_3 = "Capabilities to discover a therapeutic for the disease, and clinical effectiveness studies to discover therapeutics, to include antiviral agents."
VACCINE_1 = "Methods evaluating potential complication of Antibody-Dependent Enhancement (ADE) in vaccine recipients."
VACCINE_2 = "Exploration of use of best animal models and their predictive value for a human vaccine."
VACCINE_3 = "Efforts targeted at a universal coronavirus vaccine."
VACCINE_4 = "Approaches to evaluate risk for enhanced disease after vaccination"
VACCINE_5 = "Assays to evaluate vaccine immune response and process development for vaccines, alongside suitable animal models [in conjunction with therapeutics]"
DISTRIBUTION = "Alternative models to aid decision makers in determining how to prioritize and distribute scarce, newly proven therapeutics as production ramps up. This could include identifying approaches for expanding production capacity to ensure equitable and timely distribution to populations in need."
QUESTIONS = [PREV, THERAPEUTIC_1, THERAPEUTIC_2, THERAPEUTIC_3,
VACCINE_1, VACCINE_2, VACCINE_3, VACCINE_4, VACCINE_5, DISTRIBUTION]
# -
QUESTIONS[CLUSTER_NUMBER]
# ### Import and read clustered data
# +
df_nn = pd.read_csv(FILE_NAME)
df_nn = df_nn.drop(columns=['Unnamed: 0'])
df_nn['disease_on_abstract'] = [ast.literal_eval(a) for a in df_nn['disease_on_abstract']]
df_nn['chemicals_on_abstract'] = [ast.literal_eval(a) for a in df_nn['chemicals_on_abstract']]
df_nn['cluster_vote'] = [ast.literal_eval(a) for a in df_nn['cluster_vote']]
# -
df_nn.head()
indexes = []
for i in range(len(df_nn)):
if CLUSTER_NUMBER in df_nn['cluster_vote'][i]:
indexes.append(i)
df_cluster = df_nn.loc[indexes].reset_index(drop=True)
# ### Vizualization of groups and advising what to read
plt.figure(figsize=(16, 12))
plt.scatter(df_nn['tsne_0'], df_nn['tsne_1'], color='lightgrey')
plt.scatter(df_cluster['tsne_0'], df_cluster['tsne_1'], color='red')
df_cluster[df_cluster['license']!='question']
# ### Number of Articles through time
df_cluster_articles = df_cluster[df_cluster['license']!='question']
for i in range(len(df_cluster_articles)):
try:
df_cluster_articles['publish_time'][i] = datetime.datetime.strptime(df_cluster_articles['publish_time'][i],
"%Y-%m-%d").date().year
except:
df_cluster_articles['publish_time'][i] = datetime.datetime.strptime(df_cluster_articles['publish_time'][i],
"%Y").date().year
# +
x = Counter(df_cluster_articles['publish_time'].values)
labels, values = zip(*OrderedDict(x.most_common()).items())
plt.figure(figsize=(10, 3))
ax = plt.subplot(111)
barlist = ax.bar(labels, values, width=0.5, align='center', tick_label=labels, color='sandybrown')
if 2020 in labels:
barlist[0].set_color('saddlebrown')
plt.xticks(rotation='vertical')
plt.show()
# -
# ### Keywords Extraction
# https://towardsdatascience.com/textrank-for-keyword-extraction-by-python-c0bae21bcec0
# +
ARTICLE = '. '.join(df_cluster[df_cluster['license']!='question']['abstract'][1:25].values)
from gensim.summarization import keywords
c = keywords(ARTICLE, words=20, lemmatize=True, pos_filter=('NP', 'NN'), scores=True, split=True)
# -
wordcloud = WordCloud(prefer_horizontal=1.0,
width=800,
height=300,
max_font_size=200,
max_words=25,
colormap="copper",
background_color='white').generate_from_frequencies(dict(c))
plt.figure()
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
plt.close()
# ### NER
# +
chemical_flat_list = [item.lower() for sublist in df_cluster['chemicals_on_abstract'] for item in sublist]
c = Counter(chemical_flat_list)
c.most_common(10)
# +
disease_flat_list = [item.lower() for sublist in df_cluster['disease_on_abstract'] for item in sublist]
c = Counter(disease_flat_list)
c.most_common(10)
# -
# ## Text Summarization
# ### Extractive
# https://huggingface.co/emilyalsentzer/Bio_Discharge_Summary_BERT
#
# https://pypi.org/project/bert-extractive-summarizer/
# Load model, model config and tokenizer via Transformers
custom_config = AutoConfig.from_pretrained('emilyalsentzer/Bio_Discharge_Summary_BERT')
custom_config.output_hidden_states=True
custom_tokenizer = AutoTokenizer.from_pretrained('emilyalsentzer/Bio_Discharge_Summary_BERT')
custom_model = AutoModel.from_pretrained('emilyalsentzer/Bio_Discharge_Summary_BERT',
config=custom_config)
model = Summarizer(custom_model=custom_model, custom_tokenizer=custom_tokenizer)
model(ARTICLE, min_length=60, max_length=150)
# ### Abstractive
# https://huggingface.co/transformers/usage.html#summarization
#
# https://arxiv.org/pdf/1910.13461.pdf
summarizer = pipeline("summarization")
summarizer(ARTICLE, max_length=160, min_length=100)[0]['summary_text']
# ## Conclusion:
#
# - The abstractive summarization seems to be much better and coherent
| notebooks/.ipynb_checkpoints/6_insights-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import urllib.request
print('Beginning file download with urllib2...')
url = 'https://ww2.icb.usp.br/icb/wp-content/uploads/2019/10/Jacqueline_Alves_06-11-19.pdf?x89681'
urllib.request.urlretrieve(url, '../data/temp.pdf')
# +
from pdf2image import convert_from_path
pages = convert_from_path('../data/temp.pdf', 500)
for page in pages:
page.save('../data/temp.png', 'PNG')
# +
import pytesseract as ocr
from PIL import Image
phrase = ocr.image_to_string(Image.open('../data/temp.png'), lang='por')
print(phrase)
| src/converting_pdf_to_image.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + [markdown] dotnet_interactive={"language": "markdown"}
// # Dividing By Zero
//
// We all know that mathematically speaking, dividing by zero is an illegal operation. Right?
//
// Let's look at dividing integers first:
// + dotnet_interactive={"language": "csharp"}
int three = 3;
int zero = 0;
Console.WriteLine($"{three / zero}");
// + [markdown] dotnet_interactive={"language": "markdown"}
// Dividing decimals should have the same output:
// + dotnet_interactive={"language": "csharp"}
decimal three = 3;
decimal zero = 0;
Console.WriteLine($"{three / zero}");
// + [markdown] dotnet_interactive={"language": "markdown"}
// Finally, doubles should return the same:
// + dotnet_interactive={"language": "csharp"}
double three = 3;
double zero = 0;
Console.WriteLine($"{three / zero}");
| puzzles/1_DivideByZero.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:LHLFinal]
# language: python
# name: conda-env-LHLFinal-py
# ---
# + [markdown] id="b53e7677-ab48-4a81-88b7-61188dbf4f5f"
# ### Import packages
#
# Import of the packages that will be needed for the project. This includes packages for data manipulation, sklearn modules and custom functions.
# + id="3650a5d6-1e9a-4c25-81e3-aa4eea52a763" executionInfo={"status": "ok", "timestamp": 1638026634487, "user_tz": 300, "elapsed": 448, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
import pandas as pd
import numpy as np
import pickle
from sklearn.metrics import mean_squared_error, mean_absolute_error, r2_score
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder, RobustScaler, StandardScaler, FunctionTransformer, PowerTransformer
from sklearn.base import TransformerMixin
from sklearn.svm import SVR
# + id="b616207c-eef1-4e3d-9707-05e3c9edcc50" executionInfo={"status": "ok", "timestamp": 1638026634488, "user_tz": 300, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
import warnings
from pandas.core.common import SettingWithCopyWarning
warnings.simplefilter(action = 'ignore')
# + colab={"base_uri": "https://localhost:8080/"} id="xxjONhHPZ3nB" executionInfo={"status": "ok", "timestamp": 1638026637523, "user_tz": 300, "elapsed": 3039, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="567f5e14-3bd7-4824-b35f-7d2d474e6c38"
# !pip install pandas==1.2.0
# + [markdown] id="99fd408d-6acd-4210-a609-c8496356f996"
# ### Import Data
#
# Let's import the dataframe that we will be using for modelling
# + colab={"base_uri": "https://localhost:8080/"} id="Z-JEp08q7EEw" executionInfo={"status": "ok", "timestamp": 1638026637523, "user_tz": 300, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="c20ec194-2145-4e52-9ec2-43b192bedbe2"
from google.colab import drive
drive.mount('/content/drive')
# + id="JOvLNWbK7Cwt" executionInfo={"status": "ok", "timestamp": 1638026637757, "user_tz": 300, "elapsed": 237, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
def offensive_contribution(team_yards, player_yards):
"""
Calculate a percentage for the percentage of team yards that a player contributes to.
Input:
- Dataframe to use in the calculation
Output:
- New dataframe column with the desired contribution score
"""
contribution = player_yards / team_yards
if contribution > 1.0:
return 1.0
else:
return contribution
#--------------------------------------------------------------------
def get_contribution(df):
"""
Apply offensive_contribution(), taking in the whole dataframe as input.
"""
df['YardageContribution'] = df.apply(lambda x: offensive_contribution(x['YardsFor'],
x['TotalYards'],
), axis = 1)
return df
#---------------------------------------------------------
# Define the stats for which we need to calculate trailing averages
stats_for_trailing = ['TotalTouchdowns','RushingYards','PassingInterceptions','PassingTouchdowns','PassingRating','PassingYards',
'PassingCompletionPercentage', 'PassingLong','RushingYards', 'RushingTouchdowns', 'RushingLong',
'RushingYardsPerAttempt', 'ReceivingYardsPerReception', 'PuntReturns', 'PuntReturnTouchdowns',
'Receptions','ReceivingYards','ReceivingTargets', 'ReceivingTouchdowns', 'ExtraPointsMade', 'FieldGoalsMade',
'FieldGoalsMade40to49','FieldGoalsMade50Plus','Fumbles','FumblesLost', 'TeamPoints', 'OpponentPoints', 'YardsFor', 'YardsAgainst']
def trailing_stats_mean(df):
"""
Function to create a dataframe with a trailing aggregate mean
as a new feature for prediction. Does so for each column in the global
variable stats_for_trailing
Inputs:
- df: The dataframe on which the function will be applied
- Column: The column on which to apply the function
- Window: The number of past values to consider when apply the function
Output:
- An aggregate value
"""
#Access the column names in stats_for_trailing
global stats_for_trailing
# Get all unique players in the DataFrame
players = df['Name'].unique().tolist()
# Define a DataFrame to hold our values
df_out = pd.DataFrame()
# Loop through the unique players
for player in players:
# Create a temporary dataframe for each player
temp_df = df[df['Name'] == player]
# Calculate the n game trailing average for all players. Set closed parameter to 'left'
# so that the current value for fantasy points is not included in the calculation.
# Backfill the two resulting NaN values
for column in stats_for_trailing:
temp_df[f'TA7{column}'] = temp_df[column].fillna(method = 'ffill').rolling(window = 7,
closed = 'left').mean().fillna(method = 'bfill')
temp_df[f'TA3{column}'] = temp_df[column].rolling(window = 3,
closed = 'left').mean().fillna(method = 'bfill')
# Append the temporary dataframe to the output
df_out = df_out.append(temp_df)
# Return a dataframe with the values sorted by the original index
df_out.sort_index(inplace = True)
return df_out
#---------------------------------------------------------
def trailing_stats_single_column(df, column):
"""
Function to create a new column with a trailing aggregate mean
as a new feature for prediction.
Inputs:
- df: The dataframe on which the function will be applied
- Column: The column on which to apply the function
- Window: The number of past values to consider when apply the function
Output:
- An aggregate value
"""
# Get all unique players in the DataFrame
players = df['Name'].unique().tolist()
# Make a dataframe to store the output
df_out = pd.DataFrame()
# Loop through the unique players
for player in players:
# Create a temporary dataframe for each player
temp_df = df[df['Name'] == player]
# Calculate the n game trailing average for all players. Set closed parameter to 'left'
# so that the current value for fantasy points is not included in the calculation.
# Backfill the two resulting NaN values
temp_df[f'TA7{column}'] = temp_df[column].fillna(method='ffill').rolling(window = 7,
closed = 'left').mean().fillna(method = 'bfill')
temp_df[f'TA3{column}'] = temp_df[column].rolling(window = 3,
closed = 'left').mean().fillna(method = 'bfill')
# Append the temporary dataframe to the output
df_out = df_out.append(temp_df)
# Return a dataframe with the values sorted by the original index
df_out.sort_index(inplace = True)
return df_out
#---------------------------------------------------------
def tier_maker(position, points):
"""
Take in two arguments:
- Position: Column of the dataframe holding the player position
- Points: Trailing average of fantasy points for a given player
Classify players at every position to a tier based on their recent
performance (Trailing average fantasy points). Classifications will work
as follows.
Running Back:
-RB1: Trailing average greater than 18pts
-RB2: Trailing average between 12 and 18 pts
-RB3: Trailing average between 8 and 12 pts
-RB4: Trailing average below 8 pts
Wide Receiver:
-WR1: Trailing average greater than 18pts
-WR2: Trailing average between 12 and 18 pts
-WR3: Trailing average between 8 and 12 pts
-WR4: Trailing average below 8 pts
Tight End:
-TE1: Trailing average greater than 15pts
-TE2: Trailing average between 10 and 15 pts
-TE3: Trailing average below 10 pts
Quarterback:
-QB1: Trailing average greater than 24pts
-QB2: Trailing average between 18 and 24pts
-QB3: Trailing average between 12 and 18pts
-QB4: Trailing average below 12 pts
Kicker:
- K1: Trailing average greater than 10 pts
- K2: Trailing average between 7 and 10 points
- K3: Trailing average below 7 points
"""
# Let's make tier assignments for running backs
if position == 'RB':
if points > 18:
return 'RB1'
elif 12 < points <= 18:
return 'RB2'
elif 8 < points <= 12:
return 'RB3'
else:
return 'RB4'
# Let's make tier assignments for wide receivers
if position == 'WR':
if points > 18:
return 'WR1'
elif 12 < points <= 18:
return 'WR2'
elif 8 < points <= 12:
return 'WR3'
else:
return 'WR4'
# Let's make tier assignments for tight ends
if position == 'TE':
if points > 15:
return 'TE1'
elif 10 < points <= 15:
return 'TE2'
else:
return 'TE3'
# Let's make tier assignments for quarterbacks
if position == 'QB':
if points > 24:
return 'QB1'
elif 16 < points <= 22:
return 'QB2'
elif 10 < points <= 16:
return 'QB3'
else:
return 'QB4'
# Let's make tier assignments for kickers
if position == 'K':
if points > 10:
return 'K1'
elif 7 < points <= 10:
return 'K2'
else:
return 'K3'
# If noting is returned before this point, return np.nan
return np.nan
#---------------------------------------------------------------------
def get_tiers(df):
"""
Apply the tier_maker function to the entire dataframe.
"""
df['PlayerTier'] = df.apply(lambda x: tier_maker(x['Position'], x['TA7FantasyPointsPPR']),
axis = 1)
return df
#---------------------------------------------------------------------
def get_touchdowns(df):
"""
Get the total number of touchdowns for a player in a given week.
Input:
- Dataframe
Output:
- Dataframe with a new column representing total touchdowns"""
TD_sum = df['ReceivingTouchdowns'] + df['RushingTouchdowns'] + df['PassingTouchdowns']
df['TotalTouchdowns'] = TD_sum
return df
def get_yards(df):
"""
Get the total number of yards for a player in a given week.
Input:
- Dataframe
Output:
- Dataframe with a new column representing total touchdowns"""
yardage_sum = df['ReceivingYards'] + df['RushingYards'] + df['PassingYards']
df['TotalYards'] = yardage_sum
return df
#---------------------------------------------------------------------
def LogShift(X):
X_10 = X + 10
X_log = np.log(X_10)
return X_log
#---------------------------------------------------------------------
from sklearn.base import TransformerMixin
# Build a new transformer class to convert the sparse matrix output of the
# pipeline to a dense matrix compatible with the model
class DenseTransformer(TransformerMixin):
def fit(self, X, y = None, **fit_params):
return self
def transform(self, X, y = None, **fit_params):
return X.todense()
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="9e62a442-6037-4bb3-9f13-94325a7ca85b" executionInfo={"status": "ok", "timestamp": 1638026638125, "user_tz": 300, "elapsed": 376, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="ab341129-2e36-49b2-b7d8-96fa0a4a744a"
data = pd.read_csv('drive/MyDrive/LHL_Final_Project/Data/weekly_data.csv')
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="a9249b88-7eae-46d4-a4cf-bba7b3976a00" executionInfo={"status": "ok", "timestamp": 1638026638126, "user_tz": 300, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="3b66b709-4ce4-4ffd-a36c-bc89d20664d4"
data['Week'] = data['Week'].astype(str)
data.isna().sum()
# + id="pNRdIEjqAQ3t" executionInfo={"status": "ok", "timestamp": 1638026638126, "user_tz": 300, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# + [markdown] id="671af809-974a-42ef-a3b3-f1dc504b70d4"
# Before the train test split, we have to calculate the trailing average fantasy points for each observation, as we cannot incorporate this step into the pipeline without causing data leakage.
# + id="9c4ee866-1436-4aea-b508-f2497c0b9872" executionInfo={"status": "ok", "timestamp": 1638026638127, "user_tz": 300, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
def trailing_stats(df):
"""
Function to create a new column with a trailing aggregate mean
as a new feature for prediction.
Inputs:
- df: The dataframe on which the function will be applied
- Column: The column on which to apply the function
- Window: The number of past values to consider when apply the function
Output:
- An aggregate value
"""
#Access the column names in stats_for_trailing
global stats_for_trailing
# Get all unique players in the DataFrame
players = df['Name'].unique().tolist()
# Define a DataFrame to hold our values
df_out = pd.DataFrame()
# Loop through the unique players
for player in players:
# Create a temporary dataframe for each player
temp_df = df[df['Name'] == player]
# Calculate the n game trailing average for all players. Set closed parameter to 'left'
# so that the current value for fantasy points is not included in the calculation.
# Backfill the two resulting NaN values
for column in stats_for_trailing:
temp_df[f'TA7{column}'] = temp_df[column].rolling(window = 7, closed = 'left').mean().fillna(method = 'bfill')
temp_df[f'TA3{column}'] = temp_df[column].rolling(window = 3, closed = 'left').mean().fillna(method = 'bfill')
# Append the temporary dataframe to the output
df_out = df_out.append(temp_df)
# Return a dataframe with the values sorted by the original index
df_out = df_out.sort_index()
return df_out
# + id="b7c74494-19e2-483e-984a-103e813508a7" executionInfo={"status": "ok", "timestamp": 1638026638229, "user_tz": 300, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
stats_for_trailing = ['FantasyPointsPPR']
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="bf194a3f-92c0-4203-ac41-af5e5c1d4fe0" executionInfo={"status": "ok", "timestamp": 1638026671941, "user_tz": 300, "elapsed": 33715, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="f82a8387-42a5-4804-fe5b-30b6a26a2ded"
# Prepare the trailing average fantasy points column
data = trailing_stats(data)
data.head()
# + [markdown] id="31959661-c748-44e3-96a1-2bfe5fc39424"
# ### Train Test Split
# + id="zh_YUzcwZofn" executionInfo={"status": "ok", "timestamp": 1638026671942, "user_tz": 300, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
data['LogFantasyPointsPPR'] = LogShift(data['FantasyPointsPPR'])
# + id="0407499a-b8dd-4433-99e9-11d70ffc1d80" executionInfo={"status": "ok", "timestamp": 1638026671943, "user_tz": 300, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Separate data from the target
y = data['LogFantasyPointsPPR']
data.drop(columns = ['FantasyPointsPPR', 'LogFantasyPointsPPR'],
inplace = True)
X = data
# + id="5d378f8e-2c4b-4b88-877f-5a3a3c855965" executionInfo={"status": "ok", "timestamp": 1638026672076, "user_tz": 300, "elapsed": 140, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Execute the train, test split
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size = 0.2,
shuffle = False)
# + [markdown] id="afdacb97-f333-46e6-a454-d886a5e31cd1"
# ### Feature Engineering
#
# The main features that we will be engineering to predict a player's fantasy output will be the 5-game trailing average of various statistics as well as the binning of players into their respective tiers based on recent performance.
# + id="0518879c-5553-48bd-b01e-f0319dabe27c" executionInfo={"status": "ok", "timestamp": 1638026672076, "user_tz": 300, "elapsed": 15, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# + id="5c9ffb5a-873b-46f6-aea3-bbf99fb42883" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638026672077, "user_tz": 300, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="87b1b519-6301-4842-fbcf-a843c7171a69"
# Define the columns for which we want a 5 game trailing average.
stats_for_trailing = ['TotalTouchdowns','RushingYards','PassingInterceptions','PassingTouchdowns','PassingRating','PassingYards',
'PassingCompletionPercentage', 'PassingLong','RushingYards', 'RushingTouchdowns', 'RushingLong',
'RushingYardsPerAttempt', 'ReceivingYardsPerReception', 'PuntReturns', 'PuntReturnTouchdowns',
'Receptions','ReceivingYards','ReceivingTargets', 'ReceivingTouchdowns', 'ExtraPointsMade', 'FieldGoalsMade',
'FieldGoalsMade40to49','FieldGoalsMade50Plus','Fumbles','FumblesLost', 'TeamPoints', 'OpponentPoints', 'YardsFor', 'YardsAgainst']
trailing_stats = []
for col in stats_for_trailing:
trailing_stats.append('TA3' + col)
trailing_stats.append('TA7' + col)
trailing_stats.append('TA3FantasyPointsPPR')
trailing_stats.append('TA7FantasyPointsPPR')
trailing_stats
# + id="cbc659c2-805a-4466-9160-943bba379e23" executionInfo={"status": "ok", "timestamp": 1638026672078, "user_tz": 300, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Instantiate the function transformers for the feature engineering pipeline
touchdown_transformer = FunctionTransformer(get_touchdowns) # Get total touchdowns per week per player
yard_transformer = FunctionTransformer(get_yards) # Get total yardage per week per player
trailing_transformer = FunctionTransformer(trailing_stats_mean) # Get the 5 game trailing averages of appropriate statistics
tier_transformer = FunctionTransformer(get_tiers) # Bin players into the appropriate tiers based on recent performance
contribution_transformer = FunctionTransformer(get_contribution) # Calculate the offensive contribution of a given player relative to the team's offense
# Instantiate the pipeline for the necessary transformations
engineering = Pipeline([('touchdown', touchdown_transformer),
('yards', yard_transformer),
('trailing', trailing_transformer),
('tier', tier_transformer),
('contribution', contribution_transformer)])
# + [markdown] id="7754f786-e5f5-4959-b46a-09f7fc44c99d"
# <br>
#
# ### Preprocessing
#
# As shown above, the bulk of the null values fall into one of two categories. They are either:
# * In the InjuryStatus column
# * Here we can impute a value of healthy, as the only values in the injury column are
# * In the TA (trailing average) columns we created
# * No player with a null value played more than 5 games, therefore we cannot calculate the trailing average for them. We will impute a default value of 0 for these columns, as they represent players who likely did not have much impact. If they had an impact, they likely would have played in more games. I will explore imputing the median value as well through a grid search.
# + id="219e428c-dc9b-4ca0-a50c-acb529cdc85b" executionInfo={"status": "ok", "timestamp": 1638026672079, "user_tz": 300, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Define the groups of columns for preprocessing steps.
categorical_columns = ['Week',
'Team',
'Opponent',
'PlayerTier',
'InjuryStatus']
numerical_columns = trailing_stats
# + id="KqTUn2A2sZ7N" executionInfo={"status": "ok", "timestamp": 1638026672080, "user_tz": 300, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Build a new transformer class to convert the sparse matrix output of the
# pipeline to a dense matrix compatible with the model
class DenseTransformer(TransformerMixin):
def fit(self, X, y = None, **fit_params):
return self
def transform(self, X, y = None, **fit_params):
return X.todense()
# + id="89ab4df5-0eaf-4119-a2e3-2334bbf8afdc" executionInfo={"status": "ok", "timestamp": 1638026672080, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Create a custom function to generate a log-transformed version of continuous data with a constant 5 added prior to the transform.
LogShiftTransformer = FunctionTransformer(LogShift)
# + id="d511b177-7069-43fb-8042-3a00dc3a7f06" executionInfo={"status": "ok", "timestamp": 1638026672081, "user_tz": 300, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Define the preprocessing steps for categorical features
categorical_transform = Pipeline([('impute_cat',SimpleImputer(strategy = 'constant',
fill_value = 'Healthy')),
('one_hot_encoder', OneHotEncoder(handle_unknown = 'ignore'))])
# Define the preprocessing steps for numerical features
numerical_transform = Pipeline([('impute_num', SimpleImputer(strategy = 'mean')),
('scaler', LogShiftTransformer)])
# Instantiate the column transformer object for the preprocessing steps
preprocessing = ColumnTransformer([('num', numerical_transform, numerical_columns),
('cat', categorical_transform, categorical_columns)])
# + [markdown] id="nMfd3WFmQ4T6"
# ### Modelling
# + id="6615856c-53b2-436e-bdcd-e22f9c9ed0f7" executionInfo={"status": "ok", "timestamp": 1638026672081, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Instantiate a pipeline with a linear regression model as a baseline
pipeline = Pipeline([('engineering', engineering),
('prep', preprocessing),
('model', SVR())])
# + id="1a90931d-d2d0-4cf8-ba7d-3814daa8fc5f" executionInfo={"status": "ok", "timestamp": 1638026672082, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Set param grid values, parameters for grid search
param_grid = {'model__kernel': [ 'rbf'],
'model__C': [1, 10, 15],
'model__epsilon': [.001, .1]}
grid_search = GridSearchCV(pipeline,
param_grid = param_grid,
cv = 5,
verbose = 2,
n_jobs = -2)
# + colab={"base_uri": "https://localhost:8080/"} id="f1fdcfe7-849a-4a0a-80b6-4dc08dae0051" executionInfo={"status": "ok", "timestamp": 1638029759923, "user_tz": 300, "elapsed": 3087852, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="a34c2da5-233f-4412-daf6-051df08ae969"
# Fit the grid search to X_train and y_train
grid_search.fit(X_train, y_train)
# + id="ab80897a-b345-406e-a6a9-4b759d026252" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029759927, "user_tz": 300, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="31a74e57-5cc3-4466-9109-ea6975627432"
grid_search.best_score_
# + id="He9_P2VOlWYq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029759930, "user_tz": 300, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="e7b206fb-43ba-40d4-9812-145a34fd133e"
grid_search.best_params_
# + id="83880adc-a015-4698-9a7f-18dc397fe1fc" executionInfo={"status": "ok", "timestamp": 1638029830366, "user_tz": 300, "elapsed": 70458, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
preds = grid_search.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="CBlOYEI5bkIQ" executionInfo={"status": "ok", "timestamp": 1638029830368, "user_tz": 300, "elapsed": 40, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="5888d1ab-4f1e-4a15-9140-f1e18790d4eb"
preds
# + id="934c4b02-9873-464f-884a-2eb3e3fceb63" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029830370, "user_tz": 300, "elapsed": 37, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="df5c1273-fac8-4b7f-b879-0dd4f131101e"
mean_squared_error(y_test, preds)
# + id="b6fa8f4a-a86f-4cd7-b0a6-8061a0be294e" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029830371, "user_tz": 300, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="a7fee662-b071-4adb-b2d0-249566d4946c"
mean_absolute_error(y_test, preds)
# + id="ykb8Ru3ap4bT" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029830372, "user_tz": 300, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="acbb5964-aa22-44dc-9ae3-619592ffa81a"
r2_score(y_test, preds)
# + [markdown] id="1fa02e0a-d33e-4159-9a67-4e4e09918efb"
# Let's dig into the predictions a little bit
# + id="bbe2fbae-0426-4ece-b267-b34b3ab8c0ae" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029830373, "user_tz": 300, "elapsed": 23, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="5665d7f5-c7f0-42f9-f6e4-8a28c81db56e"
y_test
# + id="mjc5yNirQ757" executionInfo={"status": "ok", "timestamp": 1638029830883, "user_tz": 300, "elapsed": 528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Save pickle of model
import pickle
pickle.dump(grid_search, open('drive/MyDrive/LHL_Final_Project/Pickles/SVR.pickle', 'wb'))
# + id="620a5e8a-034b-4999-be72-c6383c59de18" executionInfo={"status": "ok", "timestamp": 1638029830884, "user_tz": 300, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
df_preds = pd.DataFrame(y_test)
# + id="ffb075d5-bb6b-46d6-87e4-1f27e87a2728" executionInfo={"status": "ok", "timestamp": 1638029830885, "user_tz": 300, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
df_preds['LogPredicted'] = preds
# + colab={"base_uri": "https://localhost:8080/", "height": 676} id="C6UeLXCingn6" executionInfo={"status": "ok", "timestamp": 1638029830886, "user_tz": 300, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="6f4b085c-4d38-4fa3-8a88-fcab81a3b2da"
df_preds['LogPredicted'] = preds
df_preds['Predicted'] = np.exp(df_preds['LogPredicted']) - 10
df_preds['Target'] = np.exp(df_preds['LogFantasyPointsPPR']) - 10
df_preds.sample(20)
# + id="21c5651f-28f0-4594-af6d-1c22c74bf8ff" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029830887, "user_tz": 300, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="9d03b831-3962-4c68-be98-426097b52c3b"
r2_score(df_preds['Target'], df_preds['Predicted'])
# + id="6d5e2a22-405b-4f78-96db-ebd06fe565d5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638029831021, "user_tz": 300, "elapsed": 147, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="1bb0d63f-8d02-49f0-d080-c7d9c9f2b9a4"
mean_squared_error(df_preds['Target'], df_preds['Predicted'])
# + colab={"base_uri": "https://localhost:8080/"} id="nJmNwyORn5Q4" executionInfo={"status": "ok", "timestamp": 1638029831026, "user_tz": 300, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="63e9305b-63ac-4dde-cb6f-42cfc210a6a2"
mean_absolute_error(df_preds['Target'], df_preds['Predicted'])
# + [markdown] id="jmxfwruUK-03"
# # Time to dig into the results
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="zVolEA5gK-i4" executionInfo={"status": "ok", "timestamp": 1638029831129, "user_tz": 300, "elapsed": 115, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="18000354-9bb0-41c9-8219-f4c274fc2a41"
df_review = X_test
df_review['Predicted'] = df_preds['Predicted']
df_review['Target'] = df_preds['Target']
df_review['Error'] = df_review['Target'] - df_review['Predicted']
df_review.head()
# + id="ypn9ZJ7wn7nX" executionInfo={"status": "ok", "timestamp": 1638029831353, "user_tz": 300, "elapsed": 227, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
import seaborn as sns
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 569} id="bciWZ6_QLFnD" executionInfo={"status": "ok", "timestamp": 1638029831849, "user_tz": 300, "elapsed": 499, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="288c9c37-3d88-4d79-902e-8be62d1f10c0"
# Let's visualize how the prediction error varies by player position.
plt.figure(figsize = (12, 9))
sns.violinplot(x = df_review['Position'],
y = df_review['Error'])
# + id="qEwwX8brLH3D" executionInfo={"status": "ok", "timestamp": 1638029883336, "user_tz": 300, "elapsed": 51492, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
# Let's apply our engineered features to df_review and keep on digging
df_review = trailing_stats_mean(df_review)
df_review['PlayerTier'] = df_review.apply(lambda x: tier_maker(x['Position'], x['TA7FantasyPointsPPR']), axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 645} id="j42LwhbpLN2T" executionInfo={"status": "ok", "timestamp": 1638029883338, "user_tz": 300, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}} outputId="50226acb-fb60-4697-8b50-7c591c9fb6a7"
df_review['AbsoluteError'] = abs(df_review['Error'])
mean_tier_error = df_review[['PlayerTier', 'Error', 'AbsoluteError']].groupby('PlayerTier').mean()
sum_tier_error = df_review[['PlayerTier', 'Error','AbsoluteError']].groupby('PlayerTier').sum()
count_tier_error = df_review[['PlayerTier', 'Error']].groupby('PlayerTier').count()
mean_tier_error['Total Absolute Error'] = sum_tier_error['AbsoluteError']
mean_tier_error['Total Error'] = sum_tier_error['Error']
mean_tier_error['Error Count'] = count_tier_error['Error']
mean_tier_error.rename({'Error': 'Mean Error'},
axis = 1)
# + id="57kMu31CLQLp" executionInfo={"status": "ok", "timestamp": 1638029883340, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12779563215770517109"}}
| Modeling Work/SVR_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import datetime
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import sys
from scipy.stats import dirichlet
import seaborn as sns
sns.set(style='whitegrid')
from twitpol import config
ground_truth_polls_df = pd.read_csv(config.DATA / 'ground_truth' / 'ground_truth_polling.csv', index_col=0)
best_guess_polling_df = pd.read_csv(config.DATA / 'ground_truth' / 'best_guess_polling.csv', index_col=0)
first_valid_day = datetime.date(2019, 3, 15)
twitter_counts_df = pd.read_csv(config.DATA / 'tweet_counts' / 'FINAlFINALFINALFINAL.csv', index_col=0)
ground_truth_polls_df.index = pd.to_datetime(ground_truth_polls_df.index)
best_guess_polling_df.index = pd.to_datetime(best_guess_polling_df.index)
twitter_counts_df.index = pd.to_datetime(twitter_counts_df.index)
last_valid_day = list(twitter_counts_df.index)[-1].to_pydatetime().date()
def get_tweet_columns(threshold):
return (['Number of tweets >= ' + str(threshold) + '_' + cand
for cand in list(ground_truth_polls_df.columns)])
# get tweets between latest_poll_date and number of days in advance from that date
def get_tweets(latest_poll_date, days_in_advance, columns):
days_not_nan = np.zeros(len(columns))
tweet_tot = np.zeros(len(columns))
for i in range(1, days_in_advance + 1):
tweets = np.array(twitter_counts_df.loc[latest_poll_date + datetime.timedelta(days=i), columns])
days_not_nan += [int(not np.isnan(x)) for x in tweets]
tweet_tot += [0 if np.isnan(x) else x for x in tweets]
days_not_nan = [days if days else 1 for days in days_not_nan]
return days_in_advance * (tweet_tot / days_not_nan)
# update prior distribution using Twitter info and calculate posterior likelihood of ground truth polling
def obtain_posterior_likelihood(latest_poll_date, days_in_advance, poll_scaling_factor, decay_factor,
tweet_scaling_factor):
prediction_date = latest_poll_date + datetime.timedelta(days=days_in_advance)
sum_of_alphas = poll_scaling_factor * decay_factor ** days_in_advance
polling_prior_belief = np.array(best_guess_polling_df.loc[latest_poll_date])
proportion_top_5 = np.sum(polling_prior_belief)
prior_alphas = np.array([(cand_proportion / proportion_top_5) * sum_of_alphas
for cand_proportion in polling_prior_belief])
tweet_data = get_tweets(latest_poll_date, days_in_advance, get_tweet_columns(0.6))
posterior_alphas = prior_alphas + tweet_data * tweet_scaling_factor
ground_truth_polling = np.array(ground_truth_polls_df.loc[prediction_date])
ground_truth_polling_sum_to_1 = ground_truth_polling / np.sum(ground_truth_polling)
return(dirichlet.pdf(ground_truth_polling_sum_to_1, posterior_alphas))
# +
# performing grid search to find optimal values of poll_scaling_factor and tweet_scaling_factor
poll_scaling_factors = [10000 * x for x in range(5, 12)]
tweet_scaling_factors = np.linspace(0.1, 2, ((2 - 0.1) / 0.1) + 2) / 100
decay_factors = np.linspace(0.995, 0.95, ((0.995 - 0.95) / 0.005) + 1)
sentiment_threshold = 0.6
days_in_advance = 5
starting_days = []
curr_day = first_valid_day
while curr_day < last_valid_day:
starting_days.append(curr_day)
curr_day += datetime.timedelta(days=days_in_advance)
del starting_days[-1]
last_starting_day = starting_days[-1]
last_starting_day_days_in_advance = (last_valid_day - last_starting_day).days
cv_df = pd.DataFrame(columns=['poll_scaling_factors', 'tweet_scaling_factors', 'likelihood|model'])
cnt = 0
for poll_scaling_factor in poll_scaling_factors:
for tweet_scaling_factor in tweet_scaling_factors:
likelihood_lst = []
for starting_day in starting_days:
likelihood = obtain_posterior_likelihood(starting_day, days_in_advance, poll_scaling_factor,
1, tweet_scaling_factor)
likelihood_lst.append(likelihood)
cv_df.loc[cnt] = [poll_scaling_factor, tweet_scaling_factor, np.mean(likelihood_lst)]
cnt += 1
# -
# extract optimal hyperparamters from dataframe
cv_df.loc[cv_df['likelihood|model'].idxmax()]
| notebooks/dirichlet model/dirichlet_model_grid_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import time
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
class HMM:
'''
实现一个HMM
假设我们有N个隐藏状态,1,2,...,N。隐藏状态0和N+1是START和STOP状态,
只有在开头和结束的时候出现。
假设我们有M个观测状态,1,2,...,M。观察状态0和M+1是为了开头和结束保留的。
'''
def __init__(self, a, b):
'''
初始化HMM参数
用一些随机数来初始化模型参数。
"a", 状态转移概率矩阵,是一个 (N+2) x (N+2)的矩阵。
a(i, j)代表从隐状态i转变到隐状态j的概率。
由于我们不能从任何状态回到状态0,所以a的第一列全部为0。
同理,第N+1行也全部为0。
"b", 观测生成概率矩阵,是一个(N+2) x (M+2)的矩阵。
b(i, j)代表从隐状态i生成观测状态j的概率。
由于观测状态0只会从隐状态0中产生,第一行只有第一列为1.0,
其余都为0。同理,第N+1行只有第M+1列为1.0,其余都是0。
'''
# Check input
if (a.shape[0] != a.shape[1] or
a.shape[0] != b.shape[0]):
raise Exception("Improper a or b matrix shapes.")
# N is number of regular states
self.N = a.shape[0] - 2
# M is number of regular outputs
self.M = b.shape[1] - 2
self.a = a
self.b = b
# Useful macros
self.START = 0 # start state
self.FINISH = self.N+1 # finish state
self.STATES = range(1, self.N+1) # set of regular states
# placeholders
self.O = [] # observation sequence
self.T = 0 # number of observations
self.alpha_table = None
self.beta_table = None
self.viterbi_table = None
self.viterbi_bpointers = None
def print(self):
print(f"N {self.N} M {self.M} T {self.T}")
print("Transmission probabilities 'a'")
print(self.a)
print("Emission probabilities 'b'")
print(self.b)
def set_a_b(self, a, b):
if (a.shape[0] != a.shape[1] or a.shape[0] != b.shape[0] or
a.shape[0] != self.N+2 or b.shape[1] != self.M+2):
raise Exception('Incompatible a, b shapes')
self.a = a
self.b = b
self.reset_alpha_beta_tables()
def reset_alpha_beta_tables(self):
# intialize alpha values to save recursive calls
self.alpha_table = np.array(
[[-1.0] * (self.N+2)] * (self.T+2))
self.alpha_table[self.T+1, :self.N+1] = 0.0
self.alpha_table[0, 0] = 1.0
self.alpha_table[0, 1:] = 0.0
# initialize beta values
self.beta_table = np.array(
[[-1.0] * (self.N+2)] * (self.T+1))
# Recall beta(T+1, i) is not defined.
def set_observations(self, O):
'''
设置观测序列
假设第一个观测是0,最后一个观测是M+1,其余是
[1,...,M]。
'''
# Check that observations are valid
if (not set(O[1:-1]).issubset(set(range(1, self.M+1))) or
O[0] != 0 or O[-1] != self.M+1):
print(O)
raise Exception("Invalid observation sequence.")
self.O = O
self.T = len(O)-2
self.reset_alpha_beta_tables()
def forward(self):
'''填充 alpha_table, i.e., alpha_t(j)'s'''
# This should be called after set_observation,
# else self.T == 0.
if (self.T == 0):
raise Exception("Premature call to alpha().")
# Fill t=0 values
self.alpha_table[0, self.START] = 1.0
for j in range(1, self.N+1):
self.alpha_table[0, j] = 0.0
# 从左往右填充alpha表格
##################################################
# 你的代码 ########################################
##################################################
for t in range(1, self.T + 2): # 遍历 1 -> T + 1 时刻计算P(X_{1:t}, Z_t)
for i in range(0, self.N + 2): # 遍历当前时刻所有隐藏状态
self.alpha_table[t][i] = 0.0
for j in range(0, self.N + 2): # 遍历前一时刻所有隐藏状态
# 计算转移概率
self.alpha_table[t][i] += self.alpha_table[t - 1][j] * self.a[j][i]
# 计算发射概率
self.alpha_table[t][i] *= self.b[i][self.O[t]]
def viterbi(self, t, j):
if (self.T==0 or t < 1 or t > self.T+1 or j < 0 or
j > self.N + 1 or
(t == self.T+1 and j != self.FINISH)):
raise Exception("Invalid call to viterbi().")
# Check if value already available
rv = self.viterbi_table[t, j]
if rv <= -1:
if t == 1:
rv = self.a[self.START, j] * self.b[j, self.O[1]]
self.viterbi_bpointers[t, j] = self.START
else:
if t == self.T+1:
possibles = [(self.viterbi(t-1, i) *
self.a[i,j] , i)
for i in self.STATES]
else:
possibles = [(self.viterbi(t-1, i) *
self.a[i,j] * self.b[j, self.O[t]],
i)
for i in self.STATES]
rv, max_i = max(possibles, key=lambda tup: tup[0])
self.viterbi_bpointers[t, j] = max_i
self.viterbi_table[t, j] = rv
return rv
def decode(self, obs=None):
'''给定观测序列和self.a, self.b,
返回最可能的hidden state序列,以及它的概率。'''
seq = []
prob = 0.0
##################################################
# 你的代码 ########################################
##################################################
# 你的代码应当实现VITERBI算法
# 具体来说,把概率存在一张表格X中,把上一个状态的指针存在
# 另一张表格Y中。
# 你应该从表格Y中得到最好的隐状态序列,并计算它的概率。
# 你会用到self.a, self.b和self.T,不过你不需要使用alpha
# 或者beta表格。
T = len(obs) # 观测长度
dp = np.zeros((T, self.N + 2), dtype=np.float) # 动态规划状态
path = -np.ones((T, self.N + 2), dtype=np.int) # 动态规划路径
dp[0][0] = 1.0 # 初始化起始状态
for t in range(1, T):
for i in range(self.N + 2):
for j in range(self.N + 2):
temp = dp[t - 1][j] * self.a[j][i] * self.b[i][obs[t]]
if temp > dp[t][i]:
dp[t][i] = temp
path[t][i] = j
# 计算路径
prob = np.max(dp[T - 1])
pos = np.argmax(dp[T - 1])
for i in range(T - 1, -1, -1):
seq.append(pos)
pos = path[i][pos]
seq = list(reversed(seq))
return (seq, prob)
def backward(self):
'''填充beta_table beta_t(i)'''
# Unlike alpha we don't define beta(T+1, i).
# This should be called after set_observation,
# else self.T == 0.
if (self.T == 0):
raise Exception("Premature call to beta()")
# First fill for t = T
for i in range(1, self.N+1):
self.beta_table[self.T, i] = self.a[i, self.FINISH]
# Fill for t=1 to T-1 "recursively"
# 从右往左填充beta表格
for t in range(self.T-1, 0, -1): # 计算P(X_{t+1:T}|Z_t)
##################################################
# 你的代码 ########################################
##################################################
for i in range(1, self.N + 1): # 遍历当前时刻所有隐藏状态
self.beta_table[t][i] = 0.0
for j in range(1, self.N + 1): # 遍历后一时刻所有隐藏状态
self.beta_table[t][i] += (self.beta_table[t + 1][j] *
self.a[i][j] *
self.b[j][self.O[t + 1]])
# Fill in for t = 0 and i = START
self.beta_table[0, self.START] = sum(
[self.a[self.START, j] * self.b[j, self.O[1]] *
self.beta_table[1, j] for j in self.STATES])
def xi(self, t, i, j):
'''
Return xi_t(i, j)
'''
# The basic formula works for t=0 as well. For t=T we need
# a special case because beta(t+1, j) is not defined for all j.
if self.alpha_table[self.T+1, self.FINISH] <= 0.0:
raise Exception("Impossible observation sequence")
if t >= 0 and t < self.T:
##################################################
# 你的代码 ########################################
##################################################
numerator = (self.alpha_table[t, i] *
self.a[i, j] *
self.b[j, self.O[t + 1]] *
self.beta_table[t + 1, j])
denominator = 0
for i in range(self.N + 2):
for j in range(self.N + 2):
denominator += (self.alpha_table[t, i] *
self.a[i, j] *
self.b[j, self.O[t + 1]] *
self.beta_table[t + 1, j])
rv = numerator / denominator
elif t == self.T: # non-zero only for j == FINISH
##################################################
# 你的代码 ########################################
##################################################
numerator = (self.alpha_table[t, i] *
self.a[i, j] *
self.b[j, self.O[t + 1]])
denominator = 0
for i in range(self.N + 2):
for j in range(self.N + 2):
denominator += (self.alpha_table[t, i] *
self.a[i, j] *
self.b[j, self.O[t + 1]])
rv = numerator / denominator
else:
raise Exception("Invalid call to xi().")
return rv
def gamma(self, t, j):
'''
返回 gamma_t(j)
'''
rv = 0
##################################################
# 你的代码 ########################################
##################################################
# 你的代码需要计算 gamma_t(j)
# 你会需要用到self.alpha_table, self.beta_table,
# self.T以及self.START, self.FINISH等等。
if t == 0:
rv = (1.0 if j == self.START else 0.0)
elif t == self.T + 1:
rv = (1.0 if j == self.FINISH else 0.0)
else:
rv = ((self.alpha_table[t][j] * self.beta_table[t][j]) /
np.sum(self.alpha_table[t] * self.beta_table[t]))
return rv
def a_from_xi(self, i, j, rtype='ratio'):
'''
Return new a_{ij} computed from xi
'''
if j == self.START or i == self.FINISH:
numerator = 0.0
denominator = 1.0
else:
##################################################
# 你的代码 ########################################
##################################################
numerator = 0.0
denominator = 0.0
for t in range(self.T + 1):
numerator += self.xi(t, i, j)
denominator += self.gamma(t, i)
rv = numerator / denominator
if rtype == 'separate':
rv = (numerator, denominator)
elif math.isclose(denominator, 0.0):
print("Divide by zero in i={} j={}".format(i, j))
rv = np.nan
else:
rv = numerator/denominator
return rv
def b_from_gamma(self, j, v, rtype='ratio'):
##################################################
# 你的代码 ########################################
##################################################
numerator = 0
denominator = 0
for t in range(self.T + 2):
denominator += self.gamma(t, j)
if self.O[t] == v:
numerator += self.gamma(t, j)
if rtype == 'separate':
rv = (numerator, denominator)
elif math.isclose(denominator, 0.0):
print("Divide by zero in j={}, v={}".format(j, v))
rv = np.nan
else:
rv = numerator/denominator
return rv
def new_a(self):
rv = np.array([[-1.0] * (self.N+2)] * (self.N+2))
for i in range(self.N+2):
for j in range(self.N+2):
rv[i,j] = self.a_from_xi(i, j)
return rv
def new_b(self):
rv = np.array([[-1.0] * (self.M+2)] * (self.N+2))
for j in range(self.N+2):
for v in range(self.M+2):
rv[j,v] = self.b_from_gamma(j,v)
return rv
def fit(self, O, max_iter=10, verbose=False):
self.set_observations(O)
converged = False
i = 0
while not converged and i < max_iter:
i += 1
self.forward()
self.backward()
new_a = self.new_a()
new_b = self.new_b()
if verbose:
print(f'Iteration {i}')
print(new_a)
print(new_b)
if (np.allclose(new_a, self.a) and
np.allclose(new_b, self.b)):
converged = True
else:
self.set_a_b(new_a, new_b)
return(i)
# +
# A toy example for testing your code, based on Jason Eisner's
# Workbook
transmission_prob = np.array([
[0. , 0.5, 0.5, 0. ],
[0. , 0.8, 0.1, 0.1],
[0. , 0.2, 0.7, 0.1],
[0. , 0. , 0. , 0. ]])
emission_prob = np.array([
[1. , 0. , 0. , 0. , 0. ],
[0. , 0.7, 0.2, 0.1, 0. ],
[0. , 0. , 0.3, 0.7, 0. ],
[0. , 0. , 0. , 0. , 1. ]])
observation_short = [
0,
2, 3, 3, 2,
4
]
T_observation_short = len(observation_short) -2
observation_long = [
0,
2, 3, 3, 2,
3, 2, 3, 2,
2, 3, 1, 3,
3, 1, 1, 1,
2, 1, 1, 1,
3, 1, 2, 1,
1, 1, 2, 3,
3, 2, 3, 2,
2,
4
]
T_observation_long = len(observation_long)-2
# +
hmms = HMM(transmission_prob, emission_prob)
hmms.set_observations(observation_short)
hmms.forward()
expected_alpha = np.array(
[[1. , 0. , 0. , 0. ],
[0. , 0.1 , 0.15 , 0. ],
[0. , 0.011 , 0.0805 , 0. ],
[0. , 0.00249 , 0.040215 , 0. ],
[0. , 0.002007 , 0.00851985, 0. ],
[0. , 0. , 0. , 0.00105268]])
assert np.allclose(hmms.alpha_table, expected_alpha)
hmms.backward()
expected_beta = np.array([
[ 0.00105268, -1. , -1. , -1. ],
[-1. , 0.0011457 , 0.0062541 , -1. ],
[-1. , 0.00327 , 0.01263 , -1. ],
[-1. , 0.019 , 0.025 , -1. ],
[-1. , 0.1 , 0.1 , -1. ]])
assert np.allclose(hmms.beta_table, expected_beta)
expected_decode = ([0, 2, 2, 2, 2, 3], 0.0007563149999999998)
assert hmms.decode(hmms.O) == expected_decode
# +
hmml = HMM(transmission_prob, emission_prob)
hmml.fit(observation_long, max_iter = 20, verbose=False)
expected_a = np.array([
[0.00000000e+00, 5.68740363e-15, 1.00000000e+00, 0.00000000e+00],
[0.00000000e+00, 9.33778559e-01, 6.62214406e-02, 1.21039999e-15],
[0.00000000e+00, 7.18667227e-02, 8.64943927e-01, 6.31893502e-02],
[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]])
expected_b = np.array([
[1. , 0. , 0. , 0. , 0. ],
[0. , 0.64048263, 0.14806965, 0.21144771, 0. ],
[0. , 0. , 0.53439047, 0.46560953, 0. ],
[0. , 0. , 0. , 0. , 1. ]])
assert np.allclose(hmml.a, expected_a)
assert np.allclose(hmml.b, expected_b)
expected_decode_result = ([0, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 2,
2, 2, 2, 2, 2, 2, 3],
1.4779964597903278e-16)
assert np.allclose(hmml.decode(hmml.O)[0], expected_decode_result[0])
assert np.isclose(hmml.decode(hmml.O)[1], expected_decode_result[1])
# -
class HMMMulti(HMM):
def update_new_a_num_deno(self):
for i in range(self.N+2):
for j in range(self.N+2):
(num, deno) = self.a_from_xi(i, j, 'separate')
self.new_a_num[i,j] += num
self.new_a_deno[i,j] += deno
def update_new_b_num_deno(self):
for j in range(self.N+2):
for v in range(self.M+2):
(num, deno) = self.b_from_gamma(j, v, 'separate')
self.new_b_num[j, v] += num
self.new_b_deno[j, v] += deno
def get_likelihoods(self):
'''在每一个fit/learning步骤中计算似然概率。
假设它会在fit函数中被用到。'''
prob = 0 # in log space
self.forward()
prob += np.log(self.alpha_table[
self.T+1, self.FINISH])
return np.exp(prob)
def fit(self, Oset, max_iter=10, verbose=False):
self.new_a_num = np.array([[0.0] * (self.N+2)] * (self.N+2))
self.new_a_deno = np.array([[0.0] * (self.N+2)] * (self.N+2))
self.new_b_num = np.array([[0.0] * (self.M+2)] * (self.N+2))
self.new_b_deno = np.array([[0.0] * (self.M+2)] * (self.N+2))
self.history = []
converged = False
i = 0
while not converged and i < max_iter:
i += 1
for O in Oset:
self.set_observations(O)
self.forward()
self.backward()
self.update_new_a_num_deno()
self.update_new_b_num_deno()
new_a = self.new_a_num / self.new_a_deno
new_b = self.new_b_num / self.new_b_deno
if verbose:
print(f'Iteration {i}')
print(new_b.T)
print(new_a.T)
else:
print(f'Iteration {i}')
if (np.allclose(new_a, self.a) and
np.allclose(new_b, self.b)):
converged = True
else:
self.set_a_b(new_a, new_b)
self.new_a_num.fill(0)
self.new_a_deno.fill(0)
self.new_b_num.fill(0)
self.new_b_deno.fill(0)
self.history.append(self.get_likelihoods())
return(i)
hmmml = HMMMulti(transmission_prob, emission_prob)
hmmml.fit([observation_long]*5, max_iter = 20)
expected_a = np.array([
[0.00000000e+00, 5.68740363e-15, 1.00000000e+00, 0.00000000e+00],
[0.00000000e+00, 9.33778559e-01, 6.62214406e-02, 1.21039999e-15],
[0.00000000e+00, 7.18667227e-02, 8.64943927e-01, 6.31893502e-02],
[0.00000000e+00, 0.00000000e+00, 0.00000000e+00, 0.00000000e+00]])
expected_b = np.array([
[1. , 0. , 0. , 0. , 0. ],
[0. , 0.64048263, 0.14806965, 0.21144771, 0. ],
[0. , 0. , 0.53439047, 0.46560953, 0. ],
[0. , 0. , 0. , 0. , 1. ]])
assert np.allclose(hmml.a, expected_a)
assert np.allclose(hmml.b, expected_b)
expected_likelihoods = [8.463524947168683e-21, 6.45760751286262e-19,
2.604423720662633e-18, 3.772405535111533e-18,
3.934445572171057e-18, 3.945375545182965e-18,
3.9459945227193204e-18, 3.946030509476693e-18,
3.946032758826868e-18, 3.946032909436204e-18,
3.94603292005471e-18]
assert np.allclose(hmmml.get_likelihoods(), expected_likelihoods)
import nltk
from nltk.corpus import brown
# +
class POS:
'''
Partition given sentences, or those obtained from the brown
corpus, equally into fit sentences and test sentences.
For each obtain tags using nltk tagger. Initialize "a"
with the tags from all sentences, but with uniform
transition probabilities. Initialize "b" with all words
in all sentences. But the emission probability
initialization should only use the words in fit sentences.
For each word in the fit sentence, give its most common
tag "high" probability of emitting it. All other words
get a "low" probability of being emitted by a tag. (The
high and low values are different for each tag because of
normalization.)
'''
def __init__(self, sents=None, category=None,
num_sents=40,
b_bound = 1*10**(-4),
tagset="universal"):
if sents is None:
self.sents = brown.sents(categories=category)[:num_sents]
self.tagged_sents = brown.tagged_sents(
categories=category, tagset=tagset)[:num_sents]
else:
self.sents = [nltk.word_tokenize(s) for s in sents]
self.tagged_sents = [nltk.pos_tag(s, tagset = tagset)
for s in self.sents]
self.fsize = int(len(self.sents) * 0.5)
self.fit_tsents = self.tagged_sents[:self.fsize]
self.test_tsents = self.tagged_sents[self.fsize:]
tagged_words = [tup for s in self.tagged_sents[:self.fsize]
for tup in s]
self.vocabulary = (['_SWORD'] + # Special observation in STAG
sorted(list(set([w.lower() for
s in self.sents for w in s]))) +
['_FWORD']) # Special observation in FTAG
self.M = len(self.vocabulary) # Number of distinct observations
print(f'Vocabulary size {self.M}')
self.tags = (['STAG'] + # Special start state
sorted(list(set([p[1] for s in self.tagged_sents
for p in s]))) +
['FTAG']) # Special end state
self.N = len(self.tags) # Number of states
self.w2i = dict(zip(self.vocabulary,
range(len(self.vocabulary))))
self.t2i = dict(zip(self.tags, range(len(self.tags))))
# Initialize a (transition probabilities)
#self.a = np.random.rand(self.N, self.N)
self.a = np.ones((self.N, self.N))
self.a[:,0] = 0.0
self.a = self.a/(self.a.sum(axis=1).reshape(
self.a.shape[0],1))
self.a[-1,:] = 0.0
cfd = nltk.ConditionalFreqDist((word.lower(), tag) for
(word, tag)
in tagged_words)
# Initialize b (emission probabilities)
#self.b = np.random.rand(self.N, self.M)*b_random_bound
self.b = np.ones((self.N, self.M))*b_bound
# STAG only produces _SWORD and FTAG only produces _FWORD
self.b[0,:] = 0.0
self.b[:,0] = 0.0
self.b[0,0] = 1.0
self.b[-1,:] = 0.0
self.b[:,-1] = 0.0
self.b[-1, -1] = 1.0
# Set non-trivial probability of a word being produced
# by its most common tag
for word in cfd.conditions():
tag = (cfd[word].most_common(1)[0][0])
self.setb(tag, word, 0.5)
self.b = self.b/(self.b.sum(axis=1).reshape(self.N,1))
# Initialize an HMM on oset with a, b
self.hmm = HMMMulti(self.a.copy(), self.b.copy())
def create_oset(self):
self.oset = []
for ts in self.tagged_sents:
s, _ = split_tagged_sent(ts)
self.oset.append(self.s2o(s))
def fit(self, max_iter=10):
# Create oset
self.create_oset()
# Fit the hmm
self.hmm.fit(self.oset, max_iter=max_iter)
def test(self):
for descrip, ts_set in [("fit set", self.fit_tsents),
("test set", self.test_tsents)]:
correct = 0
sum_accuracy = 0
for ts in ts_set:
s, tag_seq = split_tagged_sent(ts)
predicted, _ = self.decode(s)
a = accuracy(tag_seq, predicted)
if a == 1.0:
correct += 1
sum_accuracy += a
avg_accuracy = sum_accuracy/len(ts_set)
correct_ratio = correct/len(ts_set)
print(f"For {descrip}: Average accuracy = "
f"{avg_accuracy:.3f},"
f" Totally correct ratio = "
f"{correct_ratio:.3f}")
def decode(self, sentence):
if sentence[0] != 0: # Weak check if converted
O = self.s2o(sentence)
else:
O = sentence.copy()
state_seq, prob = self.hmm.decode(O)
tag_seq = [self.tags[s] for s in state_seq]
return (tag_seq, prob)
def setb(self, j, w, p):
self.b[self.t2i[j], self.w2i[w]] = p
def getb(self, j, w, b=None):
if b is None:
b = self.b
return b[self.t2i[j], self.w2i[w]]
def seta(self, i, j, p):
self.a[self.t2i[i], self.t2i[j]] = p
def geta(self, i, j, a=None):
if a is None:
a = self.a
return a[self.t2i[i], self.t2i[j]]
def s2o(self, sentence):
return([0] +
[self.w2i[w.lower()] for w in sentence] +
[self.w2i['_FWORD']])
def i2t(self, state_sequence):
return [self.tags[i] for i in state_sequence][1:-1]
def print_b_unsorted(self, b=None):
if b is None:
b = self.b
for t in self.tags:
print(f'\n\n{t}', end='')
for w in self.vocabulary:
p = self.getb(t, w, b)
if p > 10**(-5):
print(f' {w}[{p:.5f}]', end='')
def print_a_unsorted(self, a=None):
if a is None:
a = self.a
for t in self.tags:
print(f'\n\n{t}', end='')
for u in self.tags:
p = self.geta(t, u, a)
print(f' {u}[{p:.5f}]', end='')
def print_b(self, b=None):
if b is None:
b = self.b
for t in self.tags:
ptups = zip(b[self.t2i[t],:],self.vocabulary)
ptups = sorted(ptups, key = lambda tup: tup[0],
reverse = True)
print(f'\n\n{t}', end='')
for tup in ptups:
print(f' {tup[1]}[{tup[0]}]', end='')
def print_a(self, a=None):
if a is None:
a = self.a
for t in self.tags:
ptups = zip(a[self.t2i[t],:],self.tags)
ptups = sorted(ptups, key = lambda tup: tup[0],
reverse = True)
print(f'\n\n{t}', end='')
for tup in ptups:
print(f' {tup[1]}[{tup[0]}]', end='')
def create_tagged_sent(self, sentence, tags):
if sentence[0] != '_SWORD':
s = (['_SWORD'] +
[w.lower() for w in sentence] +
['_FWORD'])
else:
s = sentence
return list(zip(s, tags))
def check_sent(self, sentence_number):
n = sentence_number
s = self.sents[n]
tag_seq = [tag for (word, tag) in
self.tagged_sents[n]]
predicted_tag_seq = self.decode(s)[0][1:-1]
return list(zip(s, tag_seq, predicted_tag_seq))
def split_tagged_sent(tagged_sent):
return list(zip(*tagged_sent))
def accuracy(tag_seq, predicted_tag_seq):
if predicted_tag_seq[0] == 'STAG':
p = predicted_tag_seq[1:-1]
else:
p = predicted_tag_seq
return (sum(t == pt for (t, pt) in zip(tag_seq, p))/
len(tag_seq))
# -
# * 在下面的样例中,HMM词性标注的效果不错,这个应该是因为所有的句子的真实词性顺序都是相同的,在这种情况下模型很容易估计参数
# +
sents_example_1 = [
"Janet will back the bill.",
"Richard will eat an apple.",
"Mary will run a mile.",
"Sam will write an essay.",
]
start = time.time()
pos = POS(sents_example_1)
pos.fit(100)
pos.test()
pos.print_a(pos.hmm.a)
pos.print_b(pos.hmm.b)
finish = time.time()
elapsed = finish - start
print("time elapsed: ", elapsed)
plt.plot(pos.hmm.history)
plt.show()
############################################################
# 回答以下问题: #############################################
############################################################
# 基于HMM的POS tagger模型效果如何?如果效果不好的话,会是因为什么原
# 因?
# -
# * 在第二个案例中,模型的表现就不是很好了,没有一个句子全部预测正确
# * 这个我想主要问题在于两个方面
# * 一阶马尔科夫假设:当前隐状态只和前一个时刻的隐状态相关,序列标注问题不仅和单个词相关,而是和整个观测序列的长度,上下文信息等都相关。
# * 目标函数跟实际情况不符:HMM中学到的是状态与观测的联合分布P(X, Z),而在预测问题中我们更关心条件概率P(Z|X)
# +
sents_example_2 = [
"Janet will back the bill.",
"Janet likes music.",
"Richard will eat an apple.",
"Peter needs rest.",
"Mary will run a mile.",
"Joe hates TV.",
"Sam will write an essay.",
"Ramona loves cooking.",
]
start = time.time()
pos = POS(sents_example_2)
pos.fit(100)
pos.test()
pos.print_a(pos.hmm.a)
pos.print_b(pos.hmm.b)
finish = time.time()
elapsed = finish - start
print("time elapsed: ", elapsed)
plt.plot(pos.hmm.history)
plt.show()
############################################################
# 回答以下问题: #############################################
############################################################
# 基于HMM的POS tagger模型效果如何?如果效果不好的话,会是因为什么原
# 因?
# -
| notebooks/homework/HMM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import os
import tensorflow.keras as keras
from tensorflow.keras.models import model_from_json
import operator
json_file = open("model_bw.json", "r")
model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(model_json)
loaded_model.load_weights("Save_file.h5")
# +
cap = cv2.VideoCapture(0)
categories = {0: 'Zero', 1: 'One', 2: 'Two', 3: 'Three', 4: 'Four', 5: 'Five'}
while True:
_, frame = cap.read()
frame = cv2.flip(frame, 1) #mirror image
# coordinates of the ROI
x1 = int(0.6*frame.shape[1])
y1 = 120
x2 = frame.shape[1]-10
y2 = int(0.5*frame.shape[1])
#Drawing the ROI
# The increment/decrement by 1 is to compensate for the bounding box
cv2.rectangle(frame, (x1-1, y1-1), (x2+1, y2+1), (255, 0, 0), 1)
# Extracting the ROI
roi = frame[y1:y2, x1:x2]
roi = cv2.resize(roi, (68, 68))
roi = cv2.cvtColor(roi, cv2.COLOR_BGR2GRAY)
_, test_image = cv2.threshold(roi, 120, 255, cv2.THRESH_BINARY)
cv2.imshow("test", test_image)
# Batch of 1
result = loaded_model.predict(test_image.reshape(1,68,68,1))
prediction = {'Zero': result[0][0],
'One': result[0][1],
'Two': result[0][2],
'Three': result[0][3],
'Four': result[0][4],
'Five': result[0][5]}
# Sorting based on top prediction
prediction = sorted(prediction.items(),key=operator.itemgetter(1), reverse=True)
# Displaying the predictions
cv2.putText(frame, prediction[0][0], (x1+100, y2+30), cv2.FONT_HERSHEY_PLAIN, 2, (0,255,255), 1)
cv2.imshow("Frame", frame)
interrupt = cv2.waitKey(10)
if interrupt & 0xFF == 27: #esc key
break
cap.release()
cv2.destroyAllWindows()
# -
| Predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COURSE: Master math by coding in Python
# ## SECTION: Linear algebra
#
# #### www.udemy.com/master-math-with-python
# #### INSTRUCTOR: sincxpress.com
#
# Note about this code: Each video in this section of the course corresponds to a section of code below. Please note that this code roughly matches the code shown in the live recording, but is not exactly the same -- the variable names, order of lines, and parameters may be slightly different.
# # Linear algebra BUG HUNT!!
# +
# create a column vector
cv = np.array([ (-2, 3 )])
display(Math(sym.latex(sym.sympify(cv))))
# +
# visualize scalar-vector multiplication
v = np.array([-2,2])
s = .7
sv = np.array([s,s]).T@v
plt.plot([0,v[0]],[0,v[1]],'ro-',linewidth=3,label='v')
plt.plot([0,sv],[0,sv],'o-',linewidth=3,label='%sv')
plt.axis = 'square'
plt.legend()
plt.axis([-3,3,-3,3])
plt.grid()
plt.show()
# +
# algorithm to compute the dot product
v = np.random.randn(7)
w = np.random.randn(8)
dp1 = 0
for i in range(0,len(v)):
dp1 = dp1 + v[i]*w[1]
dp2 = np.dot(v,w)
print(str(dp1) + '\t' + str(dp2))
# +
# number of data points
n = 10
# data
data1 = np.arange(0,n) + np.random.randn(n)
data2 = np.arange(0,n) + np.random.randn(n)
# compute correlation
numer = np.dot(data1,data2)
denom = np.sqrt( np.dot(data1,data1) ) * np.sqrt(np.dot(data2,data2))
r1 = numer/denom
# confirm with numpy function
r2 = np.corrcoef(data1,data2)[1][0]
print(r1)
print(r2)
# +
# outer product computation
o1 = np.random.randint(0,10,7)
o2 = np.random.randint(0,10,4)
outermat = np.zeros((len(o1),len(o2)))
for i in range(len(o2)):
outermat[i,:] = o1*o2[i]
print(outermat-np.outer(o1,o2))
# +
# matrix multiplication
A = np.random.randn(5,5)
I = np.eye(5)
A*I
# +
# matrix multiplication
A = np.random.randn(8,5)
I = np.eye(5)
print(A)
print(' ')
print(A*I)
# +
# random matrices are invertible
A = np.random.randint(-5,6,(5,5))
Ainv = np.inv(A)
np.round(A@Ainv,4)
# +
# plot the eigenspectrum
# the matrix
M = np.random.randint(-5,5,(5,5))
M = M@M.T
# its eigendecomposition
eigvecs,eigvals = np.linalg.eig(M)
plt.plot(np.matrix.flatten(eigvals),'s-')
plt.xlabel('Components')
plt.ylabel('Eigenvalues')
plt.show()
# +
# Reconstruct a matrix based on its SVD
A = np.random.randint(-10,11,(10,20))
U,s,V = np.linalg.svd(A)
# reconstruct S
S = np.diag(s)
Arecon = U@V@S
fig,ax = plt.subplots(1,3)
ax[0].imshow(A,vmin=-10,vmax=10)
ax[0].set_title('A')
ax[1].imshow(Arecon,vmin=-10,vmax=10)
ax[1].set_title('Arecon')
ax[2].imshow(A-Arecon,vmin=-10,vmax=10)
ax[2].set_title('A-Arecon')
plt.show()
| MXC_pymath_linalg_BUGHUNT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 用于预训练BERT的数据集
# :label:`sec_bert-dataset`
#
# 为了预训练 :numref:`sec_bert`中实现的BERT模型,我们需要以理想的格式生成数据集,以便于两个预训练任务:遮蔽语言模型和下一句预测。一方面,最初的BERT模型是在两个庞大的图书语料库和英语维基百科(参见 :numref:`subsec_bert_pretraining_tasks`)的合集上预训练的,但它很难吸引这本书的大多数读者。另一方面,现成的预训练BERT模型可能不适合医学等特定领域的应用。因此,在定制的数据集上对BERT进行预训练变得越来越流行。为了方便BERT预训练的演示,我们使用了较小的语料库WikiText-2 :cite:`Merity.Xiong.Bradbury.ea.2016`。
#
# 与 :numref:`sec_word2vec_data`中用于预训练word2vec的PTB数据集相比,WikiText-2(1)保留了原来的标点符号,适合于下一句预测;(2)保留了原来的大小写和数字;(3)大了一倍以上。
#
# + origin_pos=1 tab=["mxnet"]
import os
import random
from mxnet import gluon, np, npx
from d2l import mxnet as d2l
npx.set_np()
# + [markdown] origin_pos=3
# 在WikiText-2数据集中,每行代表一个段落,其中在任意标点符号及其前面的词元之间插入空格。保留至少有两句话的段落。为了简单起见,我们仅使用句号作为分隔符来拆分句子。我们将更复杂的句子拆分技术的讨论留在本节末尾的练习中。
#
# + origin_pos=4 tab=["mxnet"]
#@save
d2l.DATA_HUB['wikitext-2'] = (
'https://s3.amazonaws.com/research.metamind.io/wikitext/'
'wikitext-2-v1.zip', '3c914d17d80b1459be871a5039ac23e752a53cbe')
#@save
def _read_wiki(data_dir):
file_name = os.path.join(data_dir, 'wiki.train.tokens')
with open(file_name, 'r') as f:
lines = f.readlines()
# 大写字母转换为小写字母
paragraphs = [line.strip().lower().split(' . ')
for line in lines if len(line.split(' . ')) >= 2]
random.shuffle(paragraphs)
return paragraphs
# + [markdown] origin_pos=5
# ## 为预训练任务定义辅助函数
#
# 在下文中,我们首先为BERT的两个预训练任务实现辅助函数。这些辅助函数将在稍后将原始文本语料库转换为理想格式的数据集时调用,以预训练BERT。
#
# ### 生成下一句预测任务的数据
#
# 根据 :numref:`subsec_nsp`的描述,`_get_next_sentence`函数生成二分类任务的训练样本。
#
# + origin_pos=6 tab=["mxnet"]
#@save
def _get_next_sentence(sentence, next_sentence, paragraphs):
if random.random() < 0.5:
is_next = True
else:
# paragraphs是三重列表的嵌套
next_sentence = random.choice(random.choice(paragraphs))
is_next = False
return sentence, next_sentence, is_next
# + [markdown] origin_pos=7
# 下面的函数通过调用`_get_next_sentence`函数从输入`paragraph`生成用于下一句预测的训练样本。这里`paragraph`是句子列表,其中每个句子都是词元列表。自变量`max_len`指定预训练期间的BERT输入序列的最大长度。
#
# + origin_pos=8 tab=["mxnet"]
#@save
def _get_nsp_data_from_paragraph(paragraph, paragraphs, vocab, max_len):
nsp_data_from_paragraph = []
for i in range(len(paragraph) - 1):
tokens_a, tokens_b, is_next = _get_next_sentence(
paragraph[i], paragraph[i + 1], paragraphs)
# 考虑1个'<cls>'词元和2个'<sep>'词元
if len(tokens_a) + len(tokens_b) + 3 > max_len:
continue
tokens, segments = d2l.get_tokens_and_segments(tokens_a, tokens_b)
nsp_data_from_paragraph.append((tokens, segments, is_next))
return nsp_data_from_paragraph
# + [markdown] origin_pos=9
# ### 生成遮蔽语言模型任务的数据
# :label:`subsec_prepare_mlm_data`
#
# 为了从BERT输入序列生成遮蔽语言模型的训练样本,我们定义了以下`_replace_mlm_tokens`函数。在其输入中,`tokens`是表示BERT输入序列的词元的列表,`candidate_pred_positions`是不包括特殊词元的BERT输入序列的词元索引的列表(特殊词元在遮蔽语言模型任务中不被预测),以及`num_mlm_preds`指示预测的数量(选择15%要预测的随机词元)。在 :numref:`subsec_mlm`中定义遮蔽语言模型任务之后,在每个预测位置,输入可以由特殊的“掩码”词元或随机词元替换,或者保持不变。最后,该函数返回可能替换后的输入词元、发生预测的词元索引和这些预测的标签。
#
# + origin_pos=10 tab=["mxnet"]
#@save
def _replace_mlm_tokens(tokens, candidate_pred_positions, num_mlm_preds,
vocab):
# 为遮蔽语言模型的输入创建新的词元副本,其中输入可能包含替换的“<mask>”或随机词元
mlm_input_tokens = [token for token in tokens]
pred_positions_and_labels = []
# 打乱后用于在遮蔽语言模型任务中获取15%的随机词元进行预测
random.shuffle(candidate_pred_positions)
for mlm_pred_position in candidate_pred_positions:
if len(pred_positions_and_labels) >= num_mlm_preds:
break
masked_token = None
# 80%的时间:将词替换为“<mask>”词元
if random.random() < 0.8:
masked_token = '<mask>'
else:
# 10%的时间:保持词不变
if random.random() < 0.5:
masked_token = tokens[mlm_pred_position]
# 10%的时间:用随机词替换该词
else:
masked_token = random.choice(vocab.idx_to_token)
mlm_input_tokens[mlm_pred_position] = masked_token
pred_positions_and_labels.append(
(mlm_pred_position, tokens[mlm_pred_position]))
return mlm_input_tokens, pred_positions_and_labels
# + [markdown] origin_pos=11
# 通过调用前述的`_replace_mlm_tokens`函数,以下函数将BERT输入序列(`tokens`)作为输入,并返回输入词元的索引(在 :numref:`subsec_mlm`中描述的可能的词元替换之后)、发生预测的词元索引以及这些预测的标签索引。
#
# + origin_pos=12 tab=["mxnet"]
#@save
def _get_mlm_data_from_tokens(tokens, vocab):
candidate_pred_positions = []
# tokens是一个字符串列表
for i, token in enumerate(tokens):
# 在遮蔽语言模型任务中不会预测特殊词元
if token in ['<cls>', '<sep>']:
continue
candidate_pred_positions.append(i)
# 遮蔽语言模型任务中预测15%的随机词元
num_mlm_preds = max(1, round(len(tokens) * 0.15))
mlm_input_tokens, pred_positions_and_labels = _replace_mlm_tokens(
tokens, candidate_pred_positions, num_mlm_preds, vocab)
pred_positions_and_labels = sorted(pred_positions_and_labels,
key=lambda x: x[0])
pred_positions = [v[0] for v in pred_positions_and_labels]
mlm_pred_labels = [v[1] for v in pred_positions_and_labels]
return vocab[mlm_input_tokens], pred_positions, vocab[mlm_pred_labels]
# + [markdown] origin_pos=13
# ## 将文本转换为预训练数据集
#
# 现在我们几乎准备好为BERT预训练定制一个`Dataset`类。在此之前,我们仍然需要定义辅助函数`_pad_bert_inputs`来将特殊的“<mask>”词元附加到输入。它的参数`examples`包含来自两个预训练任务的辅助函数`_get_nsp_data_from_paragraph`和`_get_mlm_data_from_tokens`的输出。
#
# + origin_pos=14 tab=["mxnet"]
#@save
def _pad_bert_inputs(examples, max_len, vocab):
max_num_mlm_preds = round(max_len * 0.15)
all_token_ids, all_segments, valid_lens, = [], [], []
all_pred_positions, all_mlm_weights, all_mlm_labels = [], [], []
nsp_labels = []
for (token_ids, pred_positions, mlm_pred_label_ids, segments,
is_next) in examples:
all_token_ids.append(np.array(token_ids + [vocab['<pad>']] * (
max_len - len(token_ids)), dtype='int32'))
all_segments.append(np.array(segments + [0] * (
max_len - len(segments)), dtype='int32'))
# valid_lens不包括'<pad>'的计数
valid_lens.append(np.array(len(token_ids), dtype='float32'))
all_pred_positions.append(np.array(pred_positions + [0] * (
max_num_mlm_preds - len(pred_positions)), dtype='int32'))
# 填充词元的预测将通过乘以0权重在损失中过滤掉
all_mlm_weights.append(
np.array([1.0] * len(mlm_pred_label_ids) + [0.0] * (
max_num_mlm_preds - len(pred_positions)), dtype='float32'))
all_mlm_labels.append(np.array(mlm_pred_label_ids + [0] * (
max_num_mlm_preds - len(mlm_pred_label_ids)), dtype='int32'))
nsp_labels.append(np.array(is_next))
return (all_token_ids, all_segments, valid_lens, all_pred_positions,
all_mlm_weights, all_mlm_labels, nsp_labels)
# + [markdown] origin_pos=16
# 将用于生成两个预训练任务的训练样本的辅助函数和用于填充输入的辅助函数放在一起,我们定义以下`_WikiTextDataset`类为用于预训练BERT的WikiText-2数据集。通过实现`__getitem__ `函数,我们可以任意访问WikiText-2语料库的一对句子生成的预训练样本(遮蔽语言模型和下一句预测)样本。
#
# 最初的BERT模型使用词表大小为30000的WordPiece嵌入 :cite:`Wu.Schuster.Chen.ea.2016`。WordPiece的词元化方法是对 :numref:`subsec_Byte_Pair_Encoding`中原有的字节对编码算法稍作修改。为简单起见,我们使用`d2l.tokenize`函数进行词元化。出现次数少于5次的不频繁词元将被过滤掉。
#
# + origin_pos=17 tab=["mxnet"]
#@save
class _WikiTextDataset(gluon.data.Dataset):
def __init__(self, paragraphs, max_len):
# 输入paragraphs[i]是代表段落的句子字符串列表;
# 而输出paragraphs[i]是代表段落的句子列表,其中每个句子都是词元列表
paragraphs = [d2l.tokenize(
paragraph, token='word') for paragraph in paragraphs]
sentences = [sentence for paragraph in paragraphs
for sentence in paragraph]
self.vocab = d2l.Vocab(sentences, min_freq=5, reserved_tokens=[
'<pad>', '<mask>', '<cls>', '<sep>'])
# 获取下一句子预测任务的数据
examples = []
for paragraph in paragraphs:
examples.extend(_get_nsp_data_from_paragraph(
paragraph, paragraphs, self.vocab, max_len))
# 获取遮蔽语言模型任务的数据
examples = [(_get_mlm_data_from_tokens(tokens, self.vocab)
+ (segments, is_next))
for tokens, segments, is_next in examples]
# 填充输入
(self.all_token_ids, self.all_segments, self.valid_lens,
self.all_pred_positions, self.all_mlm_weights,
self.all_mlm_labels, self.nsp_labels) = _pad_bert_inputs(
examples, max_len, self.vocab)
def __getitem__(self, idx):
return (self.all_token_ids[idx], self.all_segments[idx],
self.valid_lens[idx], self.all_pred_positions[idx],
self.all_mlm_weights[idx], self.all_mlm_labels[idx],
self.nsp_labels[idx])
def __len__(self):
return len(self.all_token_ids)
# + [markdown] origin_pos=19
# 通过使用`_read_wiki`函数和`_WikiTextDataset`类,我们定义了下面的`load_data_wiki`来下载并生成WikiText-2数据集,并从中生成预训练样本。
#
# + origin_pos=20 tab=["mxnet"]
#@save
def load_data_wiki(batch_size, max_len):
"""加载WikiText-2数据集"""
num_workers = d2l.get_dataloader_workers()
data_dir = d2l.download_extract('wikitext-2', 'wikitext-2')
paragraphs = _read_wiki(data_dir)
train_set = _WikiTextDataset(paragraphs, max_len)
train_iter = gluon.data.DataLoader(train_set, batch_size, shuffle=True,
num_workers=num_workers)
return train_iter, train_set.vocab
# + [markdown] origin_pos=22
# 将批量大小设置为512,将BERT输入序列的最大长度设置为64,我们打印出小批量的BERT预训练样本的形状。注意,在每个BERT输入序列中,为遮蔽语言模型任务预测$10$($64 \times 0.15$)个位置。
#
# + origin_pos=23 tab=["mxnet"]
batch_size, max_len = 512, 64
train_iter, vocab = load_data_wiki(batch_size, max_len)
for (tokens_X, segments_X, valid_lens_x, pred_positions_X, mlm_weights_X,
mlm_Y, nsp_y) in train_iter:
print(tokens_X.shape, segments_X.shape, valid_lens_x.shape,
pred_positions_X.shape, mlm_weights_X.shape, mlm_Y.shape,
nsp_y.shape)
break
# + [markdown] origin_pos=24
# 最后,我们来看一下词量。即使在过滤掉不频繁的词元之后,它仍然比PTB数据集的大两倍以上。
#
# + origin_pos=25 tab=["mxnet"]
len(vocab)
# + [markdown] origin_pos=26
# ## 小结
#
# * 与PTB数据集相比,WikiText-2数据集保留了原来的标点符号、大小写和数字,并且比PTB数据集大了两倍多。
# * 我们可以任意访问从WikiText-2语料库中的一对句子生成的预训练(遮蔽语言模型和下一句预测)样本。
#
# ## 练习
#
# 1. 为简单起见,句号用作拆分句子的唯一分隔符。尝试其他的句子拆分技术,比如Spacy和NLTK。以NLTK为例。你需要先安装NLTK:`pip install nltk`。在代码中先`import nltk`。然后下载Punkt语句词元分析器:`nltk.download('punkt')`。要拆分句子,比如`sentences = 'This is great ! Why not ?'`,调用`nltk.tokenize.sent_tokenize(sentences)`将返回两个句子字符串的列表:`['This is great !', 'Why not ?']`。
# 1. 如果我们不过滤出一些不常见的词元,词量会有多大?
#
# + [markdown] origin_pos=27 tab=["mxnet"]
# [Discussions](https://discuss.d2l.ai/t/5737)
#
| d2l/mxnet/chapter_natural-language-processing-pretraining/bert-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# 
# +
import numpy as np
from matplotlib import pyplot as plt
import warnings
# %matplotlib inline
warnings.filterwarnings('ignore')
# +
def l_func(x):
return 1/x
def o1_func(x):
return (9-x**2)**0.5
def o2_func(x):
return -(9-x**2)**0.5
def v_func(x):
return np.abs(-2*x)
def e_func(y):
return -3*np.abs(np.sin(y))
# L
x_first = 0e-5
x_last = 1.75
x = np.linspace(x_first, x_last, num=100)
y = list(map(l_func, x))
fig = plt.figure(figsize=(20, 7))
ax = fig.add_subplot(1, 4, 1)
ax.plot(x, y, 'r', linewidth=8)
ax.grid(True)
plt.xlim(-0.25, 2)
# O
x_first = -3
x_last = 3
x = np.linspace(x_first, x_last, num=100)
y1 = list(map(o1_func, x))
y2 = list(map(o2_func, x))
ax = fig.add_subplot(1, 4, 2)
ax.plot(x, y1, 'r', linewidth=8)
ax.plot(x, y2, 'r', linewidth=8)
ax.grid(True)
plt.xlim(-3.5, 3.5)
plt.ylim(-3.25, 3.25)
# V
x_first = -1
x_last = 1
x = np.linspace(x_first, x_last, num=100)
y = list(map(v_func, x))
ax = fig.add_subplot(1, 4, 3)
ax.plot(x, y, 'r', linewidth=8)
ax.grid(True)
plt.xlim(-1.25, 1.25)
plt.ylim(-0.1, 2.10)
# E
x_first = -np.pi
x_last = np.pi
x = np.linspace(x_first, x_last, num=100)
y = list(map(e_func, x))
ax = fig.add_subplot(1, 4, 4)
ax.plot(y, x, 'r', linewidth=8)
ax.grid(True)
# ---
plt.suptitle('MATHS PROVIDE THAT ALL YOU NEED...', fontsize=50)
plt.show()
print('MATHS AGAINST THE ANEMY...')
# -
# +
def c_func_1(x):
return ((-x)**(2/3) - np.sqrt(1-(x)**2))
def c_func_2(x):
return ((-x)**(2/3) + np.sqrt(1-(x)**2))
def c_func_3(x):
return ((x)**(2/3) - np.sqrt(1-(x)**2))
def c_func_4(x):
return ((x)**(2/3) + np.sqrt(1-(x)**2))
x_first = -1
x_last = 1
x = np.linspace(x_first, x_last, num=100)
y1 = list(map(c_func_1, x))
y2 = list(map(c_func_2, x))
y3 = list(map(c_func_3, x))
y4 = list(map(c_func_4, x))
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(1, 4, 2)
ax.plot(x, y1, 'r', linewidth=8)
ax.plot(x, y2, 'r', linewidth=8)
ax.plot(x, y3, 'r', linewidth=8)
ax.plot(x, y4, 'r', linewidth=8)
ax.grid(True)
plt.xlim(-1.25, 1.25)
plt.ylim(-1.25, 1.75)
plt.suptitle('FROM MY HEART...', fontsize=50)
plt.show()
print('MATHS...')
# -
| Data Science and Machine Learning/Machine-Learning-In-Python-THOROUGH/EXAMPLES/FUNNY_PYTHON/MATHS_AGAINST_THE_ANEMY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="KdUFcDsdzRyw"
# # Clonamos el repositorio para obtener los dataSet
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="mHReFf3_y9ms" outputId="c17545fd-c7dd-42c2-e3ad-4f55db21611f"
# !git clone https://github.com/joanby/machinelearning-az.git
# + [markdown] colab_type="text" id="vNKZXgtKzU2x"
# # Damos acceso a nuestro Drive
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="5gu7KWnzzUQ0" outputId="abe602b4-3a59-470e-d508-037c6966002b"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] colab_type="text" id="1gUxIkHWzfHV"
# # Test it
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="mIQt3jBMzYRE" outputId="d050bd10-4da5-4ff3-db48-cead7fdee3d1"
# !ls '/content/drive/My Drive'
# + [markdown] colab_type="text" id="mHsK36uN0XB-"
# # Google colab tools
# + colab={} colab_type="code" id="kTzwfUPWzrm4"
from google.colab import files # Para manejar los archivos y, por ejemplo, exportar a su navegador
import glob # Para manejar los archivos y, por ejemplo, exportar a su navegador
from google.colab import drive # Montar tu Google drive
# + [markdown] colab_type="text" id="3yFpBwmNz70v"
# # Plantilla de Pre Procesado
# + [markdown] colab_type="text" id="v8OxSXXSz-OP"
# # Cómo importar las librerías
#
# + colab={} colab_type="code" id="edZX51YLzs59"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] colab_type="text" id="8XfXlqtF0B58"
# # Importar el data set
#
# + colab={} colab_type="code" id="-nnozsHsz_-N"
dataset = pd.read_csv('/content/machinelearning-az/datasets/Part 1 - Data Preprocessing/Section 2 -------------------- Part 1 - Data Preprocessing --------------------/Data.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, 3].values
# + [markdown] colab_type="text" id="x8PABYut0i7y"
# # Dividir el data set en conjunto de entrenamiento y conjunto de testing
#
# + colab={} colab_type="code" id="oPVZUP870DoR"
from sklearn.model_selection import train_test_split
# + colab={} colab_type="code" id="3lajo7ye0lEs"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] colab_type="text" id="YRMD7_oq3J_f"
# # Escalado de variables
# + colab={} colab_type="code" id="BQ-<KEY>"
from sklearn.preprocessing import StandardScaler
# + colab={} colab_type="code" id="kt9uD3hE0nxd"
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
| datasets/Part 1 - Data Preprocessing/Section 2 -------------------- Part 1 - Data Preprocessing --------------------/data_preprocessing_template.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// <a href="https://cocl.us/Data_Science_with_Scalla_top"><img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/SC0103EN/adds/Data_Science_with_Scalla_notebook_top.png" width = 750, align = "center"></a>
// <br/>
// <a><img src="https://ibm.box.com/shared/static/ugcqz6ohbvff804xp84y4kqnvvk3bq1g.png" width="200" align="center"></a>"
//
// # Linear Methods
//
//
// After completing this lesson you should be able to:
//
// * Understand the Pipeline API for Logistic Regression and Linear Least Squares
// * Perform classification with Logistic Regression
// * Perform regression with Linear Least Squares
// * Use regularization with Logistic Regression and Linear Least Squares
//
// ## Logistic Regression
//
// * Widely used to predict binary responses
// * Can be generalized into multinomial logistic regression
//
// The benefits of Logistic Regression are:
//
// * there are no tuning parameters
// * the prediction equation is simple and easy to implement
//
// ## Continuing from previous example
// + attributes={"classes": ["scala"], "id": ""}
import org.apache.spark.sql.SparkSession
val spark = SparkSession.builder().getOrCreate()
import spark.implicits._
import org.apache.spark.sql.functions._
import org.apache.spark.mllib.util.MLUtils.{
convertVectorColumnsFromML => fromML,
convertVectorColumnsToML => toML
}
// + attributes={"classes": ["scala"], "id": ""}
import org.apache.spark.mllib.util.MLUtils
val data = toML(MLUtils.loadLibSVMFile(sc, "/resources/data/sample_libsvm_data.txt").toDF())
val splitData = data.randomSplit(Array(0.7, 0.3))
val trainingData = toML(splitData(0))
val testData = toML(splitData(1))
// + attributes={"classes": ["scala"], "id": ""}
import org.apache.spark.ml.classification.LogisticRegression
import org.apache.spark.ml.classification.BinaryLogisticRegressionSummary
val logr = new LogisticRegression().setMaxIter(10).setRegParam(0.3).setElasticNetParam(0.8)
val logrModel = logr.fit(trainingData)
println(s"Weights: ${logrModel.coefficients}\nIntercept: ${logrModel.intercept}")
// + attributes={"classes": ["scala"], "id": ""}
val trainingSummaryLR = logrModel.summary
val objectiveHistoryLR = trainingSummaryLR.objectiveHistory
println(objectiveHistoryLR)
// -
// ## Linear Least Squares
//
// - Most common formulation for regression problems
// - Average loss = Mean Squared Error
// + attributes={"classes": ["scala"], "id": ""}
import org.apache.spark.ml.regression.LinearRegression
val lr = new LinearRegression().setMaxIter(10).setRegParam(0.3).setElasticNetParam(0.8)
val lrModel = lr.fit(trainingData)
println(s"Weights: ${lrModel.coefficients}\nIntercept: ${lrModel.intercept}")
// + attributes={"classes": ["scala"], "id": ""}
val trainingSummaryLLS = lrModel.summary
val objectiveHistoryLLS = trainingSummaryLLS.objectiveHistory
println(objectiveHistoryLLS)
trainingSummaryLLS.residuals.show(3)
// -
// ## Lesson Summary
//
// Having completed this lesson, you should be able to:
//
// - Understand the Pipelines API for Logistic Regression and Linear Least Squares
// - Perform classification with Logistic Regression
// - Perform classification with Linear Least Squares
// - Use regularization with Logistic Regression and Linear Least Squares
//
// ### About the Authors
//
// [<NAME>](https://www.linkedin.com/in/vpetro) is Consulting Manager at Lightbend. He holds a Masters degree in Computer Science with specialization in Intelligent Systems. He is passionate about functional programming and applications of AI.
| Scala Programming for Data Science/Data Science with Scala/Module 4: Fitting a Model/3.4.4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
def split(txt):
return re.split("(?<=[a-z])(?=[A-Z])|_|[0-9]|(?<=[A-Z])(?=[A-Z][a-z])|\\s+", txt)
split("getContainerSystem")
split("get")
split(" get ")
| regex_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Algebra
# Now that you can store and manipulate data,
# let's briefly review the subset of basic linear algebra
# that you will need to understand most of the models.
# We will introduce all the basic concepts,
# the corresponding mathematical notation,
# and their realization in code all in one place.
# If you are already confident in your basic linear algebra,
# feel free to skim through or skip this chapter.
import torch
# ## Scalars
#
# If you never studied linear algebra or machine learning,
# you are probably used to working with one number at a time.
# And know how to do basic things like add them together or multiply them.
# For example, in Palo Alto, the temperature is $52$ degrees Fahrenheit.
# Formally, we call these values $scalars$.
# If you wanted to convert this value to Celsius (using metric system's more sensible unit of temperature measurement),
# you would evaluate the expression $c = (f - 32) * 5/9$ setting $f$ to $52$.
# In this equation, each of the terms $32$, $5$, and $9$ is a scalar value.
# The placeholders $c$ and $f$ that we use are called variables
# and they represent unknown scalar values.
#
# In mathematical notation, we represent scalars with ordinary lower-cased letters ($x$, $y$, $z$).
# We also denote the space of all scalars as $\mathcal{R}$.
# For expedience, we are going to punt a bit on what precisely a space is,
# but for now, remember that if you want to say that $x$ is a scalar,
# you can simply say $x \in \mathcal{R}$.
# The symbol $\in$ can be pronounced "in" and just denotes membership in a set.
#
# In PyTorch, we work with scalars by creating tensors with just one element.
# In this snippet, we instantiate two scalars and perform some familiar arithmetic operations with them, such as addition, multiplication, division and exponentiation.
# +
x = torch.tensor([3.0])
y = torch.tensor([2.0])
print('x + y = ', x + y)
print('x * y = ', x * y)
print('x / y = ', x / y)
print('x ** y = ', torch.pow(x,y))
# -
# We can convert any tensor to a Python float by calling its `item` method.
x.item()
# ## Vectors
#
# You can think of a vector as simply a list of numbers, for example ``[1.0,3.0,4.0,2.0]``.
# Each of the numbers in the vector consists of a single scalar value.
# We call these values the *entries* or *components* of the vector.
# Often, we are interested in vectors whose values hold some real-world significance.
# For example, if we are studying the risk that loans default,
# we might associate each applicant with a vector
# whose components correspond to their income,
# length of employment, number of previous defaults, etc.
# If we were studying the risk of heart attacks hospital patients potentially face,
# we might represent each patient with a vector
# whose components capture their most recent vital signs,
# cholesterol levels, minutes of exercise per day, etc.
# In math notation, we will usually denote vectors as bold-faced,
# lower-cased letters ($\mathbf{u}$, $\mathbf{v}$, $\mathbf{w})$.
# In PyTorch, we work with vectors via 1D tensors with an arbitrary number of components.
x = torch.arange(4)
print('x = ', x)
# We can refer to any element of a vector by using a subscript.
# For example, we can refer to the $4$th element of $\mathbf{u}$ by $u_4$.
# Note that the element $u_4$ is a scalar,
# so we do not bold-face the font when referring to it.
# In code, we access any element $i$ by indexing into the ``tensor``.
x[3]
# ## Length, dimensionality and shape
#
# Let's revisit some concepts from the previous section. A vector is just an array of numbers. And just as every array has a length, so does every vector.
# In math notation, if we want to say that a vector $\mathbf{x}$ consists of $n$ real-valued scalars,
# we can express this as $\mathbf{x} \in \mathcal{R}^n$.
# The length of a vector is commonly called its $dimension$.
# As with an ordinary Python array, we can access the length of a tensor
# by calling Python's in-built ``len()`` function.
#
# We can also access a vector's length via its `.shape` attribute.
# The shape is a tuple that lists the dimensionality of the tensor along each of its axes.
# Because a vector can only be indexed along one axis, its shape has just one element.
len(x)
x.shape
# Note that the word dimension is overloaded and this tends to confuse people.
# Some use the *dimensionality* of a vector to refer to its length (the number of components).
# However some use the word *dimensionality* to refer to the number of axes that an array has.
# In this sense, a scalar *would have* $0$ dimensions and a vector *would have* $1$ dimension.
#
# **To avoid confusion, when we say *2D* array or *3D* array, we mean an array with 2 or 3 axes respectively. But if we say *$n$-dimensional* vector, we mean a vector of length $n$.**
a = 2
x = torch.tensor([1,2,3])
y = torch.tensor([10,20,30])
print(a * x)
print(a * x + y)
# ## Matrices
#
# Just as vectors generalize scalars from order $0$ to order $1$,
# matrices generalize vectors from $1D$ to $2D$.
# Matrices, which we'll typically denote with capital letters ($A$, $B$, $C$),
# are represented in code as tensors with 2 axes.
# Visually, we can draw a matrix as a table,
# where each entry $a_{ij}$ belongs to the $i$-th row and $j$-th column.
#
#
# $$A=\begin{pmatrix}
# a_{11} & a_{12} & \cdots & a_{1m} \\
# a_{21} & a_{22} & \cdots & a_{2m} \\
# \vdots & \vdots & \ddots & \vdots \\
# a_{n1} & a_{n2} & \cdots & a_{nm} \\
# \end{pmatrix}$$
#
# We can create a matrix with $n$ rows and $m$ columns in PyTorch
# by specifying a shape with two components `(n,m)`
# when calling any of our favorite functions for instantiating an `tensor`
# such as `ones`, or `zeros`.
A = torch.arange(20, dtype=torch.float32).reshape((5,4))
print(A)
# Matrices are useful data structures: they allow us to organize data that has different modalities of variation. For example, rows in our matrix might correspond to different patients, while columns might correspond to different attributes.
#
# We can access the scalar elements $a_{ij}$ of a matrix $A$ by specifying the indices for the row ($i$) and column ($j$) respectively. Leaving them blank via a `:` takes all elements along the respective dimension (as seen in the previous section).
#
# We can transpose the matrix through `t()`. That is, if $B = A^T$, then $b_{ij} = a_{ji}$ for any $i$ and $j$.
print(A.t())
# ## Tensors
#
# Just as vectors generalize scalars, and matrices generalize vectors, we can actually build data structures with even more axes. Tensors give us a generic way of discussing arrays with an arbitrary number of axes. Vectors, for example, are first-order tensors, and matrices are second-order tensors.
#
# Using tensors will become more important when we start working with images, which arrive as 3D data structures, with axes corresponding to the height, width, and the three (RGB) color channels.
X = torch.arange(24).reshape((2, 3, 4))
print('X.shape =', X.shape)
print('X =', X)
# ## Basic properties of tensor arithmetic
#
# Scalars, vectors, matrices, and tensors of any order have some nice properties that we will often rely on.
# For example, as you might have noticed from the definition of an element-wise operation,
# given operands with the same shape,
# the result of any element-wise operation is a tensor of that same shape.
# Another convenient property is that for all tensors, multiplication by a scalar
# produces a tensor of the same shape.
# In math, given two tensors $X$ and $Y$ with the same shape,
# $\alpha X + Y$ has the same shape
# (numerical mathematicians call this the AXPY operation).
a = 2
x = torch.ones(3)
y = torch.zeros(3)
print(x.shape)
print(y.shape)
print((a * x).shape)
print((a * x + y).shape)
# Shape is not the the only property preserved under addition and multiplication by a scalar. These operations also preserve membership in a vector space. But we will postpone this discussion for the second half of this chapter because it is not critical to getting your first models up and running.
#
# ## Sums and means
#
# The next more sophisticated thing we can do with arbitrary tensors
# is to calculate the sum of their elements.
# In mathematical notation, we express sums using the $\sum$ symbol.
# To express the sum of the elements in a vector $\mathbf{u}$ of length $d$,
# we can write $\sum_{i=1}^d u_i$. In code, we can just call `torch.sum()`
print(x)
print(torch.sum(x))
# We can similarly express sums over the elements of tensors of arbitrary shape. For example, the sum of the elements of an $m \times n$ matrix $A$ could be written $\sum_{i=1}^{m} \sum_{j=1}^{n} a_{ij}$.
print(A)
print(torch.sum(A))
# A related quantity is the *mean*, which is also called the *average*.
# We calculate the mean by dividing the sum by the total number of elements.
# With mathematical notation, we could write the average
# over a vector $\mathbf{u}$ as $\frac{1}{d} \sum_{i=1}^{d} u_i$
# and the average over a matrix $A$ as $\frac{1}{n \cdot m} \sum_{i=1}^{m} \sum_{j=1}^{n} a_{ij}$.
# In code, we could just call ``torch.mean()`` on tensors of arbitrary shape:
print(torch.mean(A))
print(torch.div(torch.sum(A), torch.numel(A)))
# torch.numel() calculates number of elements in a tensor
# ## Dot products
#
# So far, we have only performed element-wise operations, sums and averages. And if this was all we could do, linear algebra probably would not deserve its own chapter. However, one of the most fundamental operations is the dot product. Given two vectors $\mathbf{u}$ and $\mathbf{v}$, the dot product $\mathbf{u}^T \mathbf{v}$ is a sum over the products of the corresponding elements: $\mathbf{u}^T \mathbf{v} = \sum_{i=1}^{d} u_i \cdot v_i$.
x = torch.arange(4, dtype = torch.float32)
y = torch.ones(4, dtype = torch.float32)
print(x, y, torch.dot(x, y))
# Note that we can express the dot product of two vectors ``torch.dot(x, y)`` equivalently by performing an element-wise multiplication and then a sum:
torch.sum(x * y)
# Dot products are useful in a wide range of contexts. For example, given a set of weights $\mathbf{w}$, the weighted sum of some values ${u}$ could be expressed as the dot product $\mathbf{u}^T \mathbf{w}$. When the weights are non-negative and sum to one $\left(\sum_{i=1}^{d} {w_i} = 1\right)$, the dot product expresses a *weighted average*. When two vectors each have length one (we will discuss what *length* means below in the section on norms), dot products can also capture the cosine of the angle between them.
#
# ## Matrix-vector products
#
# Now that we know how to calculate dot products we can begin to understand matrix-vector products. Let's start off by visualizing a matrix $A$ and a column vector $\mathbf{x}$.
#
# $$A=\begin{pmatrix}
# a_{11} & a_{12} & \cdots & a_{1m} \\
# a_{21} & a_{22} & \cdots & a_{2m} \\
# \vdots & \vdots & \ddots & \vdots \\
# a_{n1} & a_{n2} & \cdots & a_{nm} \\
# \end{pmatrix},\quad\mathbf{x}=\begin{pmatrix}
# x_{1} \\
# x_{2} \\
# \vdots\\
# x_{m}\\
# \end{pmatrix} $$
#
# We can visualize the matrix in terms of its row vectors
#
# $$A=
# \begin{pmatrix}
# \mathbf{a}^T_{1} \\
# \mathbf{a}^T_{2} \\
# \vdots \\
# \mathbf{a}^T_n \\
# \end{pmatrix},$$
#
# where each $\mathbf{a}^T_{i} \in \mathbb{R}^{m}$
# is a row vector representing the $i$-th row of the matrix $A$.
#
# Then the matrix vector product $\mathbf{y} = A\mathbf{x}$ is simply a column vector $\mathbf{y} \in \mathbb{R}^n$ where each entry $y_i$ is the dot product $\mathbf{a}^T_i \mathbf{x}$.
#
# $$A\mathbf{x}=
# \begin{pmatrix}
# \mathbf{a}^T_{1} \\
# \mathbf{a}^T_{2} \\
# \vdots \\
# \mathbf{a}^T_n \\
# \end{pmatrix}
# \begin{pmatrix}
# x_{1} \\
# x_{2} \\
# \vdots\\
# x_{m}\\
# \end{pmatrix}
# = \begin{pmatrix}
# \mathbf{a}^T_{1} \mathbf{x} \\
# \mathbf{a}^T_{2} \mathbf{x} \\
# \vdots\\
# \mathbf{a}^T_{n} \mathbf{x}\\
# \end{pmatrix}
# $$
#
# So you can think of multiplication by a matrix $A\in \mathbb{R}^{n \times m}$ as a transformation that projects vectors from $\mathbb{R}^{m}$ to $\mathbb{R}^{n}$.
#
# These transformations turn out to be remarkably useful. For example, we can represent rotations as multiplications by a square matrix. As we will see in subsequent chapters, we can also use matrix-vector products to describe the calculations of each layer in a neural network.
#
# Expressing matrix-vector products in code with ``tensor``, we use ``torch.mv()`` function. When we call ``torch.mv(A, x)`` with a matrix ``A`` and a vector ``x``, PyTorch knows to perform a matrix-vector product. Note that the column dimension of ``A`` must be the same as the dimension of ``x``.
torch.mv(A, x)
# ## Matrix-matrix multiplication
#
# If you have gotten the hang of dot products and matrix-vector multiplication, then matrix-matrix multiplications should be pretty straightforward.
#
# Say we have two matrices, $A \in \mathbb{R}^{n \times k}$ and $B \in \mathbb{R}^{k \times m}$:
#
# $$A=\begin{pmatrix}
# a_{11} & a_{12} & \cdots & a_{1k} \\
# a_{21} & a_{22} & \cdots & a_{2k} \\
# \vdots & \vdots & \ddots & \vdots \\
# a_{n1} & a_{n2} & \cdots & a_{nk} \\
# \end{pmatrix},\quad
# B=\begin{pmatrix}
# b_{11} & b_{12} & \cdots & b_{1m} \\
# b_{21} & b_{22} & \cdots & b_{2m} \\
# \vdots & \vdots & \ddots & \vdots \\
# b_{k1} & b_{k2} & \cdots & b_{km} \\
# \end{pmatrix}$$
#
# To produce the matrix product $C = AB$, it's easiest to think of $A$ in terms of its row vectors and $B$ in terms of its column vectors:
#
# $$A=
# \begin{pmatrix}
# \mathbf{a}^T_{1} \\
# \mathbf{a}^T_{2} \\
# \vdots \\
# \mathbf{a}^T_n \\
# \end{pmatrix},
# \quad B=\begin{pmatrix}
# \mathbf{b}_{1} & \mathbf{b}_{2} & \cdots & \mathbf{b}_{m} \\
# \end{pmatrix}.
# $$
#
# Note here that each row vector $\mathbf{a}^T_{i}$ lies in $\mathbb{R}^k$ and that each column vector $\mathbf{b}_j$ also lies in $\mathbb{R}^k$.
#
# Then to produce the matrix product $C \in \mathbb{R}^{n \times m}$ we simply compute each entry $c_{ij}$ as the dot product $\mathbf{a}^T_i \mathbf{b}_j$.
#
# $$C = AB = \begin{pmatrix}
# \mathbf{a}^T_{1} \\
# \mathbf{a}^T_{2} \\
# \vdots \\
# \mathbf{a}^T_n \\
# \end{pmatrix}
# \begin{pmatrix}
# \mathbf{b}_{1} & \mathbf{b}_{2} & \cdots & \mathbf{b}_{m} \\
# \end{pmatrix}
# = \begin{pmatrix}
# \mathbf{a}^T_{1} \mathbf{b}_1 & \mathbf{a}^T_{1}\mathbf{b}_2& \cdots & \mathbf{a}^T_{1} \mathbf{b}_m \\
# \mathbf{a}^T_{2}\mathbf{b}_1 & \mathbf{a}^T_{2} \mathbf{b}_2 & \cdots & \mathbf{a}^T_{2} \mathbf{b}_m \\
# \vdots & \vdots & \ddots &\vdots\\
# \mathbf{a}^T_{n} \mathbf{b}_1 & \mathbf{a}^T_{n}\mathbf{b}_2& \cdots& \mathbf{a}^T_{n} \mathbf{b}_m
# \end{pmatrix}
# $$
#
# You can think of the matrix-matrix multiplication $AB$ as simply performing $m$ matrix-vector products and stitching the results together to form an $n \times m$ matrix. We can compute matrix-matrix products in PyTorch by using ``torch.mm()``.
B = torch.ones(size=(4, 3))
torch.mm(A, B)
# ## Norms
#
# Before we can start implementing models,
# there is one last concept we are going to introduce.
# Some of the most useful operators in linear algebra are norms.
# Informally, they tell us how big a vector or matrix is.
# We represent norms with the notation $\|\cdot\|$.
# The $\cdot$ in this expression is just a placeholder.
# For example, we would represent the norm of a vector $\mathbf{x}$
# or matrix $A$ as $\|\mathbf{x}\|$ or $\|A\|$, respectively.
#
# All norms must satisfy a handful of properties:
#
# 1. $\|\alpha A\| = |\alpha| \|A\|$
# 1. $\|A + B\| \leq \|A\| + \|B\|$
# 1. $\|A\| \geq 0$
# 1. If $\forall {i,j}, a_{ij} = 0$, then $\|A\|=0$
#
# To put it in words, the first rule says
# that if we scale all the components of a matrix or vector
# by a constant factor $\alpha$,
# its norm also scales by the *absolute value*
# of the same constant factor.
# The second rule is the familiar triangle inequality.
# The third rule simply says that the norm must be non-negative.
# That makes sense, in most contexts the smallest *size* for anything is 0.
# The final rule basically says that the smallest norm is achieved by a matrix or vector consisting of all zeros.
# It is possible to define a norm that gives zero norm to nonzero matrices,
# but you cannot give nonzero norm to zero matrices.
# That may seem like a mouthful, but if you digest it then you probably have grepped the important concepts here.
#
# If you remember Euclidean distances (think Pythagoras' theorem) from grade school,
# then non-negativity and the triangle inequality might ring a bell.
# You might notice that norms sound a lot like measures of distance.
#
# In fact, the Euclidean distance $\sqrt{x_1^2 + \cdots + x_n^2}$ is a norm.
# Specifically it is the $\ell_2$-norm.
# An analogous computation,
# performed over the entries of a matrix, e.g. $\sqrt{\sum_{i,j} a_{ij}^2}$,
# is called the Frobenius norm.
# More often, in machine learning we work with the squared $\ell_2$ norm (notated $\ell_2^2$).
# We also commonly work with the $\ell_1$ norm.
# The $\ell_1$ norm is simply the sum of the absolute values.
# It has the convenient property of placing less emphasis on outliers.
#
# To calculate the $\ell_2$ norm, we can just call ``torch.norm()``.
torch.norm(x)
# To calculate the L1-norm we can simply perform the absolute value and then sum over the elements.
torch.sum(torch.abs(x))
# ## Norms and objectives
#
# While we do not want to get too far ahead of ourselves, we do want you to anticipate why these concepts are useful.
# In machine learning we are often trying to solve optimization problems: *Maximize* the probability assigned to observed data. *Minimize* the distance between predictions and the ground-truth observations. Assign vector representations to items (like words, products, or news articles) such that the distance between similar items is minimized, and the distance between dissimilar items is maximized. Oftentimes, these objectives, perhaps the most important component of a machine learning algorithm (besides the data itself), are expressed as norms.
# ## Intermediate linear algebra
#
# If you have made it this far, and understand everything that we have covered,
# then honestly, you *are* ready to begin modeling.
# If you are feeling antsy, this is a perfectly reasonable place to move on.
# You already know nearly all of the linear algebra required
# to implement a number of many practically useful models
# and you can always circle back when you want to learn more.
#
# But there is a lot more to linear algebra, even as concerns machine learning.
# At some point, if you plan to make a career in machine learning,
# you will need to know more than what we have covered so far.
# In the rest of this chapter, we introduce some useful, more advanced concepts.
#
#
# ### Basic vector properties
#
# Vectors are useful beyond being data structures to carry numbers.
# In addition to reading and writing values to the components of a vector,
# and performing some useful mathematical operations,
# we can analyze vectors in some interesting ways.
#
# One important concept is the notion of a vector space.
# Here are the conditions that make a vector space:
#
# * **Additive axioms** (we assume that x,y,z are all vectors):
# $x+y = y+x$ and $(x+y)+z = x+(y+z)$ and $0+x = x+0 = x$ and $(-x) + x = x + (-x) = 0$.
# * **Multiplicative axioms** (we assume that x is a vector and a, b are scalars):
# $0 \cdot x = 0$ and $1 \cdot x = x$ and $(a b) x = a (b x)$.
# * **Distributive axioms** (we assume that x and y are vectors and a, b are scalars):
# $a(x+y) = ax + ay$ and $(a+b)x = ax +bx$.
# ### Special matrices
#
# There are a number of special matrices that we will use throughout this tutorial. Let's look at them in a bit of detail:
#
# * **Symmetric Matrix** These are matrices where the entries below and above the diagonal are the same. In other words, we have that $M^\top = M$. An example of such matrices are those that describe pairwise distances, i.e. $M_{ij} = \|x_i - x_j\|$. Likewise, the Facebook friendship graph can be written as a symmetric matrix where $M_{ij} = 1$ if $i$ and $j$ are friends and $M_{ij} = 0$ if they are not. Note that the *Twitter* graph is asymmetric - $M_{ij} = 1$, i.e. $i$ following $j$ does not imply that $M_{ji} = 1$, i.e. $j$ following $i$.
# * **Antisymmetric Matrix** These matrices satisfy $M^\top = -M$. Note that any square matrix can always be decomposed into a symmetric and into an antisymmetric matrix by using $M = \frac{1}{2}(M + M^\top) + \frac{1}{2}(M - M^\top)$.
# * **Diagonally Dominant Matrix** These are matrices where the off-diagonal elements are small relative to the main diagonal elements. In particular we have that $M_{ii} \geq \sum_{j \neq i} M_{ij}$ and $M_{ii} \geq \sum_{j \neq i} M_{ji}$. If a matrix has this property, we can often approximate $M$ by its diagonal. This is often expressed as $\mathrm{diag}(M)$.
# * **Positive Definite Matrix** These are matrices that have the nice property where $x^\top M x > 0$ whenever $x \neq 0$. Intuitively, they are a generalization of the squared norm of a vector $\|x\|^2 = x^\top x$. It is easy to check that whenever $M = A^\top A$, this holds since there $x^\top M x = x^\top A^\top A x = \|A x\|^2$. There is a somewhat more profound theorem which states that all positive definite matrices can be written in this form.
# ## Summary
#
# In just a few pages (or one Jupyter notebook) we have taught you all the linear algebra you will need to understand a good chunk of neural networks. Of course there is a *lot* more to linear algebra. And a lot of that math *is* useful for machine learning. For example, matrices can be decomposed into factors, and these decompositions can reveal low-dimensional structure in real-world datasets. There are entire subfields of machine learning that focus on using matrix decompositions and their generalizations to high-order tensors to discover structure in datasets and solve prediction problems. But this book focuses on deep learning. And we believe you will be much more inclined to learn more mathematics once you have gotten your hands dirty deploying useful machine learning models on real datasets. So while we reserve the right to introduce more math much later on, we will wrap up this chapter here.
#
# If you are eager to learn more about linear algebra, here are some of our favorite resources on the topic
#
# * For a solid primer on basics, check out <NAME>'s book [Introduction to Linear Algebra](http://math.mit.edu/~gs/linearalgebra/)
# * <NAME>'s [Linear Algebra Review and Reference](http://www.cs.cmu.edu/~zkolter/course/15-884/linalg-review.pdf)
| Ch04_The_Preliminaries_A_Crashcourse/Linear_Algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#
# # Machine learning tutorial
# # MLGWSC-1 kickoff meeting
# ### 12. 10. 2021, Zoom
# ## What is ML?
# **<NAME>**: *A computer program is said to learn from experience E with respect to some class of tasks T and performance measure P if its performance at tasks in T, as measured by P, improves with experience E.*
# ## Basic formulation of ML problem
# We need to approximate function $f^\star : \mathbb{R}^n \rightarrow \mathbb{R}^m,~ f^\star\left(\mathbf{x}\right) = \mathbf{y}$.
#
# Let us define a *model* $f : \mathbb{R}^n \times \mathbb{R}^k \rightarrow \mathbb{R}^m,~ f\left(\mathbf{x}, \theta\right) = \mathbf{y}$; typically: $\mathbf{x}$ = inputs, $\mathbf{y}$ = outputs, $\theta$ = weights.
#
# If the model is suitable to the problem, then a set of weights $\hat{\theta}$ exists such that $f\left(\mathbf{x}, \hat{\theta}\right) \approx f^\star\left(\mathbf{x}\right)$.
# ### How to find the right weights?
#
# Two more necessary ingredients:
#
# Dataset: $\mathbf{X}, \mathbf{Y}, \forall i: \mathbf{Y}_i = f^\star\left(\mathbf{X}_i\right) + \mathrm{noise}$, $\mathbf{Y}$ are called labels
#
# Loss/cost/error function: $\mathcal{C}\left(\mathbf{Y}, f\left(\mathbf{X}, \theta\right)\right)$ to measure deviation of the model from the labels (e.g. mean squared error)
#
# Training the model: $\hat{\theta} = \mathrm{argmin}_\theta\{\mathcal{C}\left(\mathbf{Y}, f\left(\mathbf{X}, \theta\right)\right)\} \rightarrow \boxed{f\left(\mathbf{x}, \hat{\theta}\right)}$
# ## Example: Polynomial regression
#
# We'll attempt to approximate the polynomial $f^\star\left(x\right) = 2x - 10x^5 + 15x^{10}$ (*true model*) using polynomial regression: minimization of the mean squared error $\mathcal{C}_{MSE}\left(\mathbf{Y}, f\left(\mathbf{X}, \theta\right)\right) = \frac{1}{N}\sum_{i=0}^{N-1}\left(\mathbf{Y}_i - f\left(\mathbf{X}_i, \theta\right)\right)^2$, and the weights are the coefficients of the polynomial.
#
# If the $\mathbf{X}$ inputs are non-degenerate (no two are the same) and there are more than the polynomial degree, there is a single minimum and it is easy to compute.
#
# We'll sample some values of $f^\star$ with a little white Gaussian noise as a **training dataset**:
# +
import numpy as np
import matplotlib.pyplot as plt
rng = np.random.default_rng()
### Create the true model and some values to plot
poly = np.polynomial.Polynomial([0., 2., 0., 0., 0., -10., 0., 0., 0., 0., 15.])
plot_X = np.linspace(0., 1., 10**3)
plot_Y = poly(plot_X)
### Create the plot limits
xlim = (0., 1.)
ylim = (min(plot_Y), max(plot_Y))
extend_factor = .1
xlim = (xlim[0]-extend_factor*(xlim[1]-xlim[0]), xlim[1]+extend_factor*(xlim[1]-xlim[0]))
ylim = (ylim[0]-extend_factor*(ylim[1]-ylim[0]), ylim[1]+extend_factor*(ylim[1]-ylim[0]))
# +
### Generate training data
# tr_X = rng.uniform(0., 1., 11)
tr_X = np.linspace(0., 1., 11)
tr_Y = poly(tr_X) + rng.normal(0., .3, len(tr_X))
### Plot the true model and the training data
plt.figure(figsize=(10., 5.))
plt.xlim(xlim)
plt.ylim(ylim)
plt.plot(plot_X, plot_Y, label='true model')
plt.scatter(tr_X, tr_Y, label='training data')
plt.legend(loc='best')
plt.xlabel('x')
plt.ylabel('y')
# -
# Now, let's train some polynomials. Change the line that starts with `deg` to set the orders of fitting polynomials. One might expect that since we have the 11 points required to fit a 10-degree polynomial and the data was generated by a 10-degree polynomial, that would again reproduce the true model the best.
# +
### Fit polynomials
degs = [1, 3, 5, 10]
fit_polys = [poly.fit(tr_X, tr_Y, deg) for deg in degs]
### Plot the fitted polynomials
plt.figure(figsize=(15., 10.))
plt.xlim(xlim)
plt.ylim(ylim)
for deg, fit_poly in zip(degs, fit_polys):
plt.plot(plot_X, fit_poly(plot_X), label='%i'%deg)
plt.scatter(tr_X, tr_Y, label='training data')
plt.plot(plot_X, plot_Y, label='true model')
plt.legend(loc='best')
plt.xlabel('x')
plt.ylabel('y')
# -
# Generate some **test data** and evaluate the training (*in-sample*) and test (*out-of-sample*) losses of a set of polynomials:
# +
### Fit polynomials
degs = range(11)
fit_polys = [poly.fit(tr_X, tr_Y, deg) for deg in degs]
### Generate test data, same distribution as training data
te_X = rng.uniform(0., 1., 100)
te_Y = poly(te_X) + rng.normal(0., .3, len(te_X))
### Compute the loss values
losses = []
for deg, fit_poly in zip(degs, fit_polys):
new_losses = []
for X, Y in ((tr_X, tr_Y), (te_X, te_Y)):
model_Y = fit_poly(X)
new_losses.append(np.mean((model_Y - Y)**2))
losses.append(new_losses)
losses = np.array(losses)
### Plot the loss values
plt.figure(figsize=(10., 5.))
plt.plot(degs, losses[:, 0], '.-', label='training')
plt.plot(degs, losses[:, 1], '.-', label='test')
plt.semilogy()
plt.ylim((1.e-2, 1.e1))
plt.legend(loc='best')
plt.xlabel('polynomial degree')
plt.ylabel('loss')
# -
# In fact, the degree-10 polynomial is not a good fit. The plot above demonstrates that with a limited number of noisy data points, too high complexity model gives the best training loss (so it's a very good fit to that particular set of data), but has trouble generalizing to new data from the same underlying distribution. This phenomenon is called **overfitting**. It is the difference between fitting and predicting.
#
# One must therefore, when working with more complex data and models, be careful about model complexity - increasing it may lead to a better fit on training data and at the same time, to worse predictions on unseen data. Even if a model is perfectly suited to a given underlying true model, in the presence of limited and/or noisy data, a less complex model may reproduce the true model better. The only way to prevent this issue is to keep an eye on the out-of-sample loss. This is achieved by a dataset split.
#
# In some cases, a dataset is supplied already split - in other cases, one must split the data themselves. In a lot of cases where the optimization is iterative, one must have a third separate dataset, called the **validation set**. Upon each iteration of the optimization algorithm over the training set, the loss on the validation set is evaluated to make sure that it is decreasing, and the training is typically interrupted once this is no longer the case. This in turn might also lead to slight overfitting on the validation dataset, which is why we still need the test set despite already having computed an out-of-sample validation loss.
# ## Gradient descent algorithms
#
# Polynomial regression was a simple example, which is easy to solve - the loss optimization has a single global minimum (under weak assumptions). With more complex ML models, this is no longer the case. Usually, gradient descent-based algorithms are used to find the minimum of $E\left(\theta\right) = \mathcal{C}\left(\mathbf{Y}, f\left(\mathbf{X}, \theta\right)\right)$. These algorithms require an "initial position" $\theta_0$ and attempt to find the solution iteratively.
#
# Basic gradient descent:
# $$\mathbf{v}_t = \eta\nabla_\theta E\left(\theta_t\right),~\theta_{t+1} = \theta_t - \mathbf{v}_t$$
#
# Let's try on a simple parabolic surface $E\left(\theta\right) = \theta_0^2 + \theta_1^2$. Feel free to play around with the value of `eta` to see why some choices of learning rate are unsuitable.
# +
### Parabolic surface and its gradient
def surface(theta):
return np.sum(theta**2, axis=0)
def gradient(theta):
return 2*theta
### Generate contour plot data
X = np.linspace(-5., 5., 1000)
Y = np.linspace(-5., 5., 1000)
Z = surface(np.stack((np.tile(np.expand_dims(X, 1), (1, len(Y))), np.tile(np.expand_dims(Y, 0), (len(X), 1))), axis=0)).T
### Initial position and GD parameters
theta = [np.array((-4., -2.))]
epochs = range(21)
eta = 0.5
### GD loop
for e in epochs[1:]:
v = eta*gradient(theta[-1])
theta.append(theta[-1]-v)
theta = np.stack(theta, axis=1)
### Plot results
plt.figure(figsize=(7., 7.))
plt.contour(X, Y, Z)
plt.plot(theta[0], theta[1], '.-', linewidth=1., markersize=10.)
plt.axis('square')
plt.xlim((min(X), max(X)))
plt.ylim((min(Y), max(Y)))
# -
# Usually, the landscape which we need our optimizer to navigate will be much more rugged with local minima and saddle points - and much higher-dimensional, so impossible to visualize! This makes it much harder to determine a suitable learning rate. However, smart adaptive algorithms exist which keep slowly decaying running averages of the gradients and other properties, which allows them to locally estimate the Hessian matrix and effectively adapt the learning rate to its ideal value. We won't learn about these in this tutorial - to try them out, simply replace the `torch.optim.SGD` by, e.g., `torch.optim.Adam`.
#
# To be able to work with complex data, we need complex models - so we need the datasets to be large. Computation of the gradient of $E\left(\theta\right)$ over the full training dataset is then computationally very difficult, let alone doing several tens or hundreds of optimization steps, so we need to apply one more approximation to our method.
#
# We assume that evaluating the loss gradient over a smaller subset of the training dataset is representative of the full computation. We split the training dataset into **batches** and do a single step of the gradient descent method on each of these batches. One iteration over all the batches is called a **training epoch**. This method is called **stochastic gradient descent**. The same principle is also used with the adaptive optimizers mentioned above.
# ## Neural Networks
#
# Neural networks are inspired by the human neural system. The basic element is a very simple **artificial neuron**, which merely takes a linear combination of its input values, with an additional constant term, and applies a non-linear function:
# $$y(\mathbf{x}) = \sigma\left(\sum_{i} w_ix_i + b\right).$$
#
# The $w_i$ values are usually called **weights** and $b$ is the **bias**. The function $\sigma$ is called the **activation function** and there are many different choices. Some of the most popular are, for example:
# $$\mathrm{Sigmoid}\left(x\right) = \frac{1}{1+e^{-x}},~\mathrm{ReLU}\left(x\right) = \mathrm{max}\left(0, x\right)~.$$
#
# For classification problems, the *Softmax* is very popular, because it maps $\mathbb{R}^N$ to a set of positive numbers that sum up to one. This is often used as a final activation to produce approximate probabilities as a model output (in other places in the network, however, others are typically used).
# $$\mathrm{Softmax}\left(\mathbf{x}\right)_i = \frac{\exp\left(x_i\right)}{\sum_j \exp\left(x_j\right)}$$
# +
### Plot the Softmax and ReLU activation functions
X = np.linspace(-10., 10., 10000)
fig, axes = plt.subplots(ncols=2, figsize=(10., 5.))
Y = 1./(1.+np.exp(-X))
axes[0].plot(X, Y, label='Sigmoid')
axes[0].legend(loc='best')
Y = np.amax(np.stack((X, np.zeros_like(X)), axis=0), axis=0)
axes[1].plot(X, Y, label='ReLU')
axes[1].legend(loc='best')
# -
# We can order multiple independent neurons into a **layer**, so that:
# $$y_i\left(\mathbf{x}\right) = \sigma\left(\sum_j w_{ij}x_j + b_i\right).$$
#
# Let us apply this to a real-life classification problem. We will use the MNIST dataset, which contains 60 000 annotated hand-written digits as single-channel $28\times 28$ images and 10 000 more for testing.
# +
import torch, torchvision
fname = 'MNIST'
### Download and initialize datasets
TrainDS_orig = torchvision.datasets.MNIST(fname, train=True, download=True)
TestDS_orig = torchvision.datasets.MNIST(fname, train=False)
# -
# Plot a few examples of the data:
### Plot examples
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
for axes_row in axes:
for ax in axes_row:
test_index = rng.integers(0, len(TestDS_orig))
image, orig_label = TestDS_orig[test_index]
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i' % orig_label)
# Let's try to use a simple neuron layer to classify the digits in the MNIST dataset. We first need to create a function that will transform an integer label to a set of probabilities, e.g., $1 \to (0., 1., 0., ..., 0.)$: $i$-th element represents the probability that the sample contains the digit $i$. We reinitialize the datasets using this transform as well as one which turns our input images into $28\times 28$ PyTorch tensors.
#
# Then we define the model itself as a sequence of functions. In the PyTorch vernacular, the first is `torch.nn.Flatten`; it rearranges the input $28\times 28$ tensor into a vector of length $28\cdot 28 = 784$. The next `torch.nn.Linear` is the linear combination performed by the neurons: this outputs 10 numbers, each of which is a linear combination of the 784 inputs with independent coefficients as well as constant biases, all of which are trainable weights. Finally, the `torch.nn.Softmax` performs the Softmax operation as described a few blocks above. We will use the SGD method with learning rate 0.1 and batch size 32 and as a loss function, we will use the binary cross entropy loss, which is the canonical choice for classification problems.
# +
### Define the label transform from an integer to a set of probabilities
def target_transform(inlabel):
newlabel = torch.zeros(10)
newlabel[inlabel] = 1.
return newlabel
### Reinitialize datasets with the transforms
TrainDS = torchvision.datasets.MNIST(fname, train=True, download=True,
target_transform=target_transform, transform=torchvision.transforms.ToTensor())
TestDS = torchvision.datasets.MNIST(fname, train=False,
target_transform=target_transform, transform=torchvision.transforms.ToTensor())
### Initialize DataLoaders as PyTorch convenience
TrainDL = torch.utils.data.DataLoader(TrainDS, shuffle=True, batch_size=32)
TestDL = torch.utils.data.DataLoader(TestDS, batch_size=1000)
### Choose device: 'cuda' or 'cpu'
device = 'cuda:0'
### Define the dense neuron layer
Network = torch.nn.Sequential(
torch.nn.Flatten(), # 28x28 -> 784
torch.nn.Linear(784, 10), # 784 -> 10
torch.nn.Softmax(dim=1)
)
Network.to(device=device)
### Get information about model
totpars = 0
for par in Network.parameters():
newpars = 1
for num in par.shape:
newpars *= num
totpars += newpars
print(Network)
print('%i trainable parameters' % totpars)
### Initialize loss function and optimizer
crit = torch.nn.BCELoss()
opt = torch.optim.SGD(Network.parameters(), lr=0.1)
# -
# Most of us don't really have an idea what a good value for the binary cross entropy is, so for reference, let's calculate what the loss will be if our model simply estimates for each sample that each digit has an 0.1 probability to be present:
### Baseline: just say it's anything at probability 1/N, what's the loss?
N = 10
labels = torch.zeros(1, 10, dtype=torch.float32)
labels[0, 3] = 1.
output = torch.full_like(labels, 1./N)
print(crit(output, labels))
# Now we have a baseline for the loss value.
#
# We finally get to training the model. The following block performs 10 training epochs, in each of them loops over all the 32-samples-long batches in the training dataset, repeating the following steps: set the gradient values to zero (these are stored as attributes of the relevant tensors), move the batch samples and labels to the proper device (CPU or CUDA), compute the outputs of the model on that particular batch, compute the loss value, compute the gradients using back-propagation (this is the algorithm that computes the gradients on neural networks) and performs the optimization step. At the end of each epoch, the loss values over all the training samples are averaged and printed along the epoch number.
# +
### Set model in training mode and create the epochs axis
Network.train()
epochs = range(1, 11)
### Train the model
for e in epochs:
tr_loss = 0.
samples = 0
### Loop over batches
for inputs, labels in TrainDL:
opt.zero_grad() # zero gradient values
inputs = inputs.to(device=device) # move input and label tensors to the device with the model
labels = labels.to(device=device)
outputs = Network(inputs) # compute model outputs
loss = crit(outputs, labels) # compute batch loss
loss.backward() # back-propagate the gradients
opt.step() # update the model weights
tr_loss += loss.clone().cpu().item()*len(inputs) # add the batch loss to the running loss
samples += len(inputs) # update the number of processed samples
tr_loss /= samples # compute training loss
print(e, tr_loss)
# -
# Now our model is trained. The in-sample (training) loss has decreased considerably, to much less than the uniform probabilities baseline. We need to make sure, however, that the model can generalize and hasn't merely memorized the training dataset. To achieve that, we evaluate the loss on the test dataset. In that process, let's also determine the **accuracy**: the percentage of test samples where the model has assigned the highest probability to the correct class.
# +
### Set model in evaluation mode
Network.eval()
### Compute the test loss
with torch.no_grad():
te_loss = 0.
samples = 0
accuracy = 0
### Loop over batches
for inputs, labels in TestDL:
inputs = inputs.to(device=device)
labels = labels.to(device=device)
outputs = Network(inputs)
loss = crit(outputs, labels)
te_loss += loss.clone().cpu().item()*len(inputs)
accuracy += torch.sum(torch.eq(torch.max(labels, 1)[1], torch.max(outputs, 1)[1]), dtype=int).clone().cpu().item()
samples += len(inputs)
te_loss /= samples
accuracy /= samples
print('Test loss: %f, accuracy: %f' % (te_loss, accuracy))
# -
# The test loss is close to the training loss. This means that we haven't run into issues with overfitting. The accuracy also seems pretty good - should be around 92%! Let us now see some of the results - out of the following two blocks, the first will show 12 random test examples of digits with their true as well as predicted classes and the confidence level the model has about its prediction, and the second will do the same but only draw from those the model has classified incorrectly.
### Draw some random images from the test dataset and compare the true labels to the network outputs
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
### Draw random images from the test dataset and select and show some which have been incorrectly classified
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
while True:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
if output!=orig_label:
break
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
# ## Deep neural networks
#
# This is already a simple neural network. However, to build a **deep neural network**, we need to stack several such layers, such that each subsequent layer takes the output of the previous layer as its input. This method of stacking allows us to build very complex models - while the individual structural elements are very simple, the non-linear elements weaved into the network of linear operations allow the model to approximate any function to an arbitrary degree as long as you're flexible on depth and layer size.
### Show schematic of a fully connected deep neural network
import matplotlib.image as mpimg
fig, ax = plt.subplots(figsize=(15., 7.))
ax.set_axis_off()
ax.imshow(mpimg.imread('nn.png'))
# This network will only have one additional layer to what we've used before. It is again a sequence of PyTorch modules - but there is now an additional layer. The first layer gives 50 outputs with the $\mathrm{ReLU}\left(x\right) = \mathrm{max}\left(0, x\right)$ activation function and the second gives 10 outputs with the $\mathrm{Softmax}$ activation. The rest of the code in this section works the same way as with the previous model.
# +
### Define a simple two-layer network
Network = torch.nn.Sequential(
torch.nn.Flatten(),
torch.nn.Linear(784, 50),
torch.nn.ReLU(),
torch.nn.Linear(50, 10),
torch.nn.Softmax(dim=1)
)
Network.to(device=device)
### Get information about model
totpars = 0
for par in Network.parameters():
newpars = 1
for num in par.shape:
newpars *= num
totpars += newpars
print(Network)
print('%i trainable parameters' % totpars)
### Initialize loss function and optimizer
crit = torch.nn.BCELoss()
opt = torch.optim.SGD(Network.parameters(), lr=0.1)
# +
### Set model in training mode and create the epochs axis
Network.train()
epochs = range(1, 11)
### Train the model
for e in epochs:
tr_loss = 0.
samples = 0
### Loop over batches
for inputs, labels in TrainDL:
opt.zero_grad() # zero gradient values
inputs = inputs.to(device=device) # move input and label tensors to the device with the model
labels = labels.to(device=device)
outputs = Network(inputs) # compute model outputs
loss = crit(outputs, labels) # compute batch loss
loss.backward() # back-propagate the gradients
opt.step() # update the model weights
tr_loss += loss.clone().cpu().item()*len(inputs) # add the batch loss to the running loss
samples += len(inputs) # update the number of processed samples
tr_loss /= samples # compute training loss
print(e, tr_loss)
# +
### Set model in evaluation mode
Network.eval()
### Compute the test loss
with torch.no_grad():
te_loss = 0.
samples = 0
accuracy = 0
### Loop over batches
for inputs, labels in TestDL:
inputs = inputs.to(device=device)
labels = labels.to(device=device)
outputs = Network(inputs)
loss = crit(outputs, labels)
te_loss += loss.clone().cpu().item()*len(inputs)
accuracy += torch.sum(torch.eq(torch.max(labels, 1)[1], torch.max(outputs, 1)[1]), dtype=int).clone().cpu().item()
samples += len(inputs)
te_loss /= samples
accuracy /= samples
print('Test loss: %f, accuracy: %f' % (te_loss, accuracy))
# -
# This model has about 4 times as many trainable parameters as the previous one, which brings an increased risk of overfitting. However, the test loss indicates that we haven't run into overfitting issues yet.
### Draw some random images from the test dataset and compare the true labels to the network outputs
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
### Draw random images from the test dataset and select and show some which have been incorrectly classified
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
while True:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
if output!=orig_label:
break
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
# ## Convolutional neural networks
#
# A slightly more convoluted (sorry for the bad pun) type of neural networks has been inspired by the animal visual cortex. Convolutional layers are introduced, which slide *filters* over an input image, producing another image - often with a larger number of channels, so these can be hard to visualize. Each pixel in that output image is then basically the product of a smaller fully connected layer (like we used before) applied to a small portion of the input image. These layers are used to build **convolutional neural networks**, which typically consist of a convolutional part and a dense part. The convolutional part is constructed out of alternating convolutional layers and pooling layers (these downsample the image between the convolutional layers), the result is flattened and passed to a dense part, which is typically a deep neural network as shown before.
#
# This structure removes some less crucial connections between neurons and use many weights for multiple connections as opposed to a fully connected network. This allows us to build a network just as suited to some problems as a fully connected network while dramatically reducing the number of free parameters to be optimized. They have proven to be particularly suitable to computer vision problems.
#
# Go ahead and play with the following - it's the same code as before, just a different network design.
fig, ax = plt.subplots(figsize=(15., 7.))
ax.set_axis_off()
ax.imshow(mpimg.imread('cnn.jpg'))
# +
### Create a simple convolutional neural network
Network = torch.nn.Sequential( # 1x28x28
torch.nn.Conv2d(1, 12, (9, 9)), # 12x20x20
torch.nn.MaxPool2d((2, 2)), # 12x10x10
torch.nn.ReLU(),
torch.nn.Conv2d(12, 24, (5, 5)), # 24x 6x 6
torch.nn.ReLU(),
torch.nn.MaxPool2d((3, 3)), # 24x 2x 2
torch.nn.Flatten(), # 96
torch.nn.Linear(96, 16), # 16
torch.nn.ReLU(),
torch.nn.Linear(16, 10), # 10
torch.nn.Softmax(dim=1)
)
Network.to(device=device)
### Get information about model
totpars = 0
for par in Network.parameters():
newpars = 1
for num in par.shape:
newpars *= num
totpars += newpars
print(Network)
print('%i trainable parameters' % totpars)
### Initialize loss function and optimizer
opt = torch.optim.SGD(Network.parameters(), lr=.1)
# +
### Set model in training mode and create the epochs axis
Network.train()
epochs = range(1, 11)
### Train the model
for e in epochs:
tr_loss = 0.
samples = 0
### Loop over batches
for inputs, labels in TrainDL:
opt.zero_grad() # zero gradient values
inputs = inputs.to(device=device) # move input and label tensors to the device with the model
labels = labels.to(device=device)
outputs = Network(inputs) # compute model outputs
loss = crit(outputs, labels) # compute batch loss
loss.backward() # back-propagate the gradients
opt.step() # update the model weights
tr_loss += loss.clone().cpu().item()*len(inputs) # add the batch loss to the running loss
samples += len(inputs) # update the number of processed samples
tr_loss /= samples # compute training loss
print(e, tr_loss)
# +
### Set model in evaluation mode
Network.eval()
### Compute the test loss
with torch.no_grad():
te_loss = 0.
samples = 0
accuracy = 0
### Loop over batches
for inputs, labels in TestDL:
inputs = inputs.to(device=device)
labels = labels.to(device=device)
outputs = Network(inputs)
loss = crit(outputs, labels)
te_loss += loss.clone().cpu().item()*len(inputs)
accuracy += torch.sum(torch.eq(torch.max(labels, 1)[1], torch.max(outputs, 1)[1]), dtype=int).clone().cpu().item()
samples += len(inputs)
te_loss /= samples
accuracy /= samples
print('Test loss: %f, accuracy: %f' % (te_loss, accuracy))
# -
### Draw some random images from the test dataset and compare the true labels to the network outputs
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
### Draw random images from the test dataset and select and show some which have been incorrectly classified
fig, axes = plt.subplots(nrows=2, ncols=6, figsize=(15., 6.))
### Loop over subplots
for axes_row in axes:
for ax in axes_row:
while True:
### Draw the images
test_index = rng.integers(0, len(TestDS))
sample, label = TestDS[test_index]
image, orig_label = TestDS_orig[test_index]
### Compute the predictions
with torch.no_grad():
output = Network(torch.unsqueeze(sample, dim=0).to(device=device))
certainty, output = torch.max(output[0], 0)
certainty = certainty.clone().cpu().item()
output = output.clone().cpu().item()
if output!=orig_label:
break
### Show image
ax.set_axis_off()
ax.imshow(image)
ax.set_title('True: %i, predicted: %i\nat %f' % (orig_label, output, certainty))
# Congratulations! You have managed to get to the point where your network mostly misclassifies only those digits, which are badly written and are outliers in the dataset. We've only scratched the surface of what you can do with neural networks, but you've learned the most elementary things you need to know. Go ahead and play with this notebook - with the networks, with the training parameters, etc. Good luck!
| tutorials/Machine Learning/ml_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```
# Implementation of Adaline algorithm
# ```
#
# ### Paper link: [Adaline algorithm](http://www-isl.stanford.edu/~widrow/papers/c1992artificialneural.pdf)
#
#
# Wikipedia:
#
#
# 
# ## Source: [Wikipedia](https://en.wikipedia.org/wiki/ADALINE)
# ADALINE (Adaptive Linear Neuron or later Adaptive Linear Element) is an early single-layer artificial neural network and the name of the physical device that implemented this network. The network uses memistors. It was developed by Professor <NAME> and his graduate student <NAME> at Stanford University in 1960. It is based on the McCulloch–Pitts neuron. It consists of a weight, a bias and a summation function.
#
# The difference between Adaline and the standard (McCulloch–Pitts) perceptron is that in the learning phase, the weights are adjusted according to the weighted sum of the inputs (the net). In the standard perceptron, the net is passed to the activation (transfer) function and the function's output is used for adjusting the weights.
#
# A multilayer network of ADALINE units is known as a MADALINE.
#
#
# Adaline is a single layer neural network with multiple nodes where each node accepts multiple inputs and generates one output. Given the following variables:as
#
# * x is the input vector
# * w is the weight vector
# * n is the number of inputs
# * $\theta$ some constant
# * y is the output of the model
#
# then we find that the output is  If we further assume that 
# 
#
# then the output further reduces to:
# 
#
# ## Learning algorithm
#
# Let us assume:
#
# * $ \eta$ is the learning rate (some positive constant)
# * y is the output of the model
# * o is the target (desired) output
# then the weights are updated as follows $ w\leftarrow w+\eta (o-y)x $. The ADALINE converges to the least squares error which is $ E=(o-y)^{2} $ This update rule is in fact the stochastic gradient descent update for linear regression.
# ## Perceptron vs Adaline
#
#
#
# The perceptron surely was very popular at the time of its discovery, however, it only took a few years until <NAME> and his doctoral student <NAME> proposed the idea of the Adaptive Linear Neuron (adaline) [3].
#
# In contrast to the perceptron rule, the delta rule of the adaline (also known as Widrow-Hoff” rule or Adaline rule) updates the weights based on a linear activation function rather than a unit step function; here, this linear activation function g(z) is just the identity function of the net input $ g(w^Tx)=w^Tx $. In the next section, we will see why this linear activation is an improvement over the perceptron update and where the name “delta rule” comes from.
# Importing necessary libraries
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from mlxtend.plotting import plot_decision_regions
import pandas as pd
# +
class Perceptron:
def __init__(self , lr = 0.001 , epochs = 50):
self.lr = lr
self.epochs = epochs
def train(self, X, y):
self.w_ = np.zeros(1 + X.shape[1])
self.errors_ = []
for _ in range(self.epochs):
errors = 0
for xi , target in zip(X , y):
update = self.lr * (target - self.predict(xi))
self.w_[1:] += update * xi
self.w_[0] += update
errors += int(update != 0.0)
self.errors_.append(errors)
return self
def net_input(self , X):
return np.dot(X, self.w_[1:]) + self.w_[0]
def predict(self , X):
return np.where(self.net_input(X) >= 0.0 , 1 , 0) # check for this
# +
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data', header=None)
# setosa and versicolor
y = df.iloc[0:100, 4].values
y = np.where(y == 'Iris-setosa', -1, 1)
# sepal length and petal length
X = df.iloc[0:100, [0,2]].values
# +
from mlxtend.plotting import plot_decision_regions
ppn = Perceptron(epochs=10, lr=0.1)
ppn.train(X, y)
print('Weights: %s' % ppn.w_)
plot_decision_regions(X, y, clf=ppn)
plt.title('Perceptron')
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.show()
plt.plot(range(1, len(ppn.errors_)+1), ppn.errors_, marker='o')
plt.xlabel('Iterations')
plt.ylabel('Misclassifications')
plt.show()
# -
# ### Problems with Perceptrons
# Although the perceptron classified the two Iris flower classes perfectly, convergence is one of the biggest problems of the perceptron. <NAME> proofed mathematically that the perceptron learning rule converges if the two classes can be separated by linear hyperplane, but problems arise if the classes cannot be separated perfectly by a linear classifier. To demonstrate this issue, we will use two different classes and features from the Iris dataset.
# +
# versicolor and virginica
y2 = df.iloc[50:150, 4].values
y2 = np.where(y2 == 'Iris-virginica', -1, 1)
# sepal width and petal width
X2 = df.iloc[50:150, [1,3]].values
ppn = Perceptron(epochs=25, lr=0.01)
ppn.train(X2, y2)
plot_decision_regions(X2, y2, clf=ppn)
plt.show()
plt.plot(range(1, len(ppn.errors_)+1), ppn.errors_, marker='o')
plt.xlabel('Iterations')
plt.ylabel('Misclassifications')
plt.show()
# -
print('Total number of misclassifications: %d of 100' % (y2 != ppn.predict(X2)).sum())
# ### Even at a lower training rate, the perceptron failed to find a good decision boundary since one or more samples will always be misclassified in every epoch so that the learning rule never stops updating the weights.
#
# ### It may seem paradoxical in this context that another shortcoming of the perceptron algorithm is that it stops updating the weights as soon as all samples are classified correctly. Our intuition tells us that a decision boundary with a large margin between the classes (as indicated by the dashed line in the figure below) likely has a better generalization error than the decision boundary of the perceptron. But large-margin classifiers such as Support Vector Machines are a topic for another time.
# 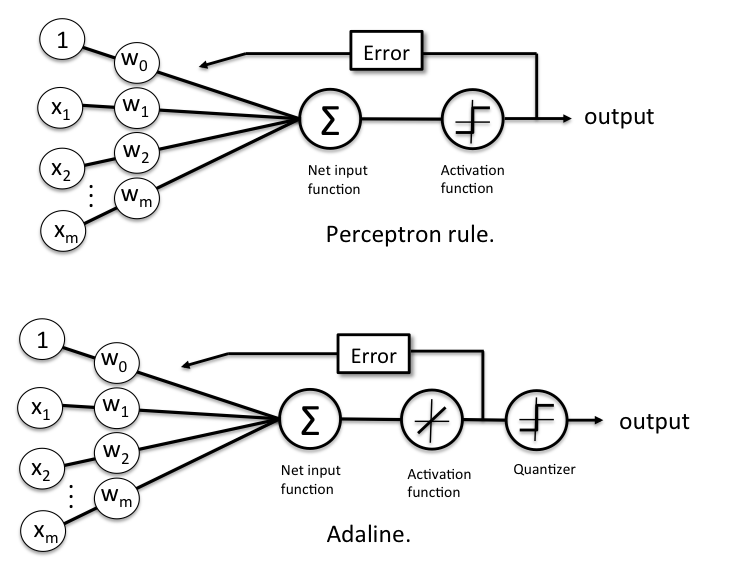
plt.rcParams['figure.figsize']= (16,9)
sns.set()
# +
# Download Dataset
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data', header=None)
print(df.isnull().sum().sum())
y = df.iloc[0:100 , 4].values
y = np.where(y == 'Iris-setosa' , -1 , 1)
# sepal length and petal length
X = df.iloc[0:100, [0,2]].values
# -
print(X.shape,y.shape)
class AdalineGD:
def __init__(self, lr = 0.01, epochs = 50):
self.lr = lr
self.epochs = epochs
def train(self, X, y):
self.w_ = np.zeros(1 + X.shape[1])
self.cost_ = []
for i in range(self.epochs):
output = self.net_input(X)
errors = (y - output)
self.w_[1:] += self.lr * X.T.dot(errors)
self.w_[0] += self.lr * errors.sum()
cost = (errors**2).sum() / 2.0
self.cost_.append(cost)
return self
def net_input(self, X):
return np.dot(X, self.w_[1:]) + self.w_[0]
def activation(self, X):
return self.net_input(X)
def predict(self, X):
return np.where(self.activation(X) > 0.0 , 1, -1)
ada = AdalineGD(epochs =10 , lr = 0.01).train(X , y)
# +
plt.plot(range(1,len(ada.cost_) +1 ), np.log10(ada.cost_), marker = 'o')
plt.xlabel("Iterations")
plt.ylabel("log(Sum squared Error)")
plt.title('Adaline - Learning rate 0.01')
plt.show()
# -
ada = AdalineGD(epochs = 10 , lr = 0.00001 ).train(X ,y)
# +
plt.plot(range(1,len(ada.cost_) + 1), np.log10(ada.cost_) ,color='red', marker = 'o')
plt.xlabel("Iterations")
plt.ylabel("log(sse)")
plt.title("Adaline - learing rate 0.0001 ")
plt.show()
# -
# ## The two plots above nicely emphasize the importance of plotting learning curves by illustrating two most common problems with gradient descent:
#
# * If the learning rate is too large, gradient descent will overshoot the minima and diverge.
# * If the learning rate is too small, the algorithm will require too many epochs to converge and can become trapped in local minima more easily.
#
#
#
# 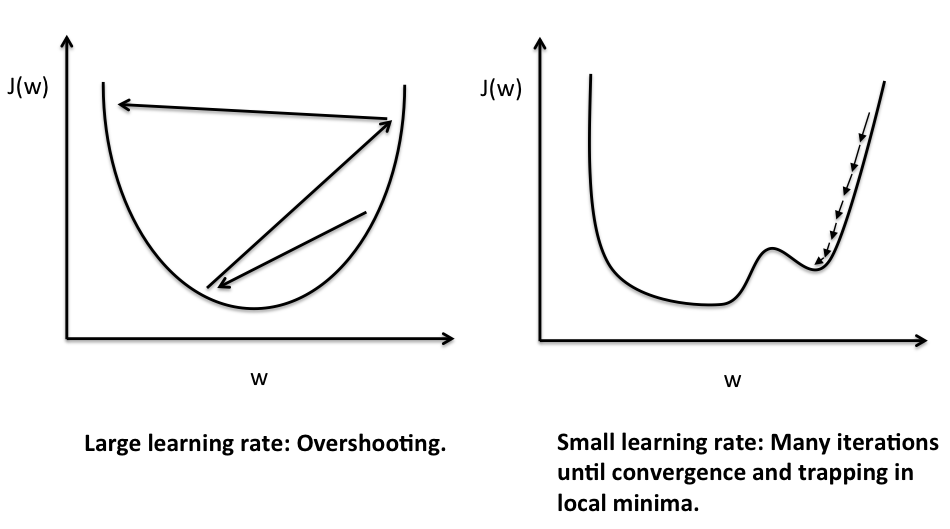
#
#
# Gradient descent is also a good example why feature scaling is important for many machine learning algorithms. It is not only easier to find an appropriate learning rate if the features are on the same scale, but it also often leads to faster convergence and can prevent the weights from becoming too small (numerical stability).
#
# A common way of feature scaling is standardization
#
#
# $$x_{j,std} = \frac{x_j - \mu_j}{\sigma_j} $$
#
# where $ \mu_j $ is the sample mean of the feature $x_j$ and σj the standard deviation, respectively. After standardization, the features will have unit variance and are centered around mean zero.
# standardize features
X_std = np.copy(X)
X_std[:,0] = (X[:,0] - X[:,0].mean()) / X[:,0].std()
X_std[:,1] = (X[:,1] - X[:,1].mean()) / X[:,1].std()
# ### Online Learning via Stochastic Gradient Descent
# The previous section was all about “batch” gradient descent learning. The “batch” updates refers to the fact that the cost function is minimized based on the complete training data set. If we think back to the perceptron rule, we remember that it performed the weight update incrementally after each individual training sample. This approach is also called “online” learning, and in fact, this is also how Adaline was first described by <NAME> et al. [3]
#
# The process of incrementally updating the weights is also called “stochastic” gradient descent since it approximates the minimization of the cost function. Although the stochastic gradient descent approach might sound inferior to gradient descent due its “stochastic” nature and the “approximated” direction (gradient), it can have certain advantages in practice. Often, stochastic gradient descent converges much faster than gradient descent since the updates are applied immediately after each training sample; stochastic gradient descent is computationally more efficient, especially for very large datasets. Another advantage of online learning is that the classifier can be immediately updated as new training data arrives, e.g., in web applications, and old training data can be discarded if storage is an issue. In large-scale machine learning systems, it is also common practice to use so-called “mini-batches”, a compromise with smoother convergence than stochastic gradient descent.
#
# In the interests of completeness let us also implement the stochastic gradient descent Adaline and confirm that it converges on the linearly separable iris dataset.
class AdalineSGD:
def __init__(self, lr =0.0001, epochs = 40):
self.lr = lr
self.epochs = epochs
def train(self , X , y , reinitialize_weights = True):
if reinitialize_weights:
self.w_ = np.zeros(1 + X.shape[1])
self.cost_ = []
for i in range(self.epochs):
for xi , target in zip(X , y):
output = self.net_input(xi)
error = (target - output)
self.w_[1:] += self.lr * xi.dot(error)
self.w_[0] += self.lr * error
cost = ((y - self.activation(X)) ** 2).sum() / 2.0
self.cost_.append(cost)
return self
def net_input(self , X):
return np.dot(X , self.w_[1:]) + self.w_[0]
def activation(self , X):
return self.net_input(X)
def predict(self , X):
return np.where(self.activation(X) >= 0.0 , 1, -1)
# One more advice before we let the adaline learn via stochastic gradient descent is to shuffle the training dataset to iterate over the training samples in random order.
#
# We shall note that the “standard” stochastic gradient descent algorithm uses sampling “with replacement,” which means that at each iteration, a training sample is chosen randomly from the entire training set. In contrast, sampling “without replacement,” which means that each training sample is evaluated exactly once in every epoch, is not only easier to implement but also shows a better performance in empirical comparisons. A more detailed discussion about this topic can be found in <NAME> and <NAME>’s paper Beneath the valley of the noncommutative arithmetic-geometric mean inequality: conjectures, case-studies, and consequences
# +
ada = AdalineSGD(epochs=15, lr=0.01)
# shuffle data
np.random.seed(123)
idx = np.random.permutation(len(y))
X_shuffled, y_shuffled = X_std[idx], y[idx]
# train and adaline and plot decision regions
ada.train(X_shuffled, y_shuffled)
plot_decision_regions(X_shuffled, y_shuffled, clf=ada)
plt.title('Adaline - Gradient Descent')
plt.xlabel('sepal length [standardized]')
plt.ylabel('petal length [standardized]')
plt.show()
plt.plot(range(1, len(ada.cost_)+1), ada.cost_, marker='o')
plt.xlabel('Iterations')
plt.ylabel('Sum-squared-error')
plt.show()
# -
# 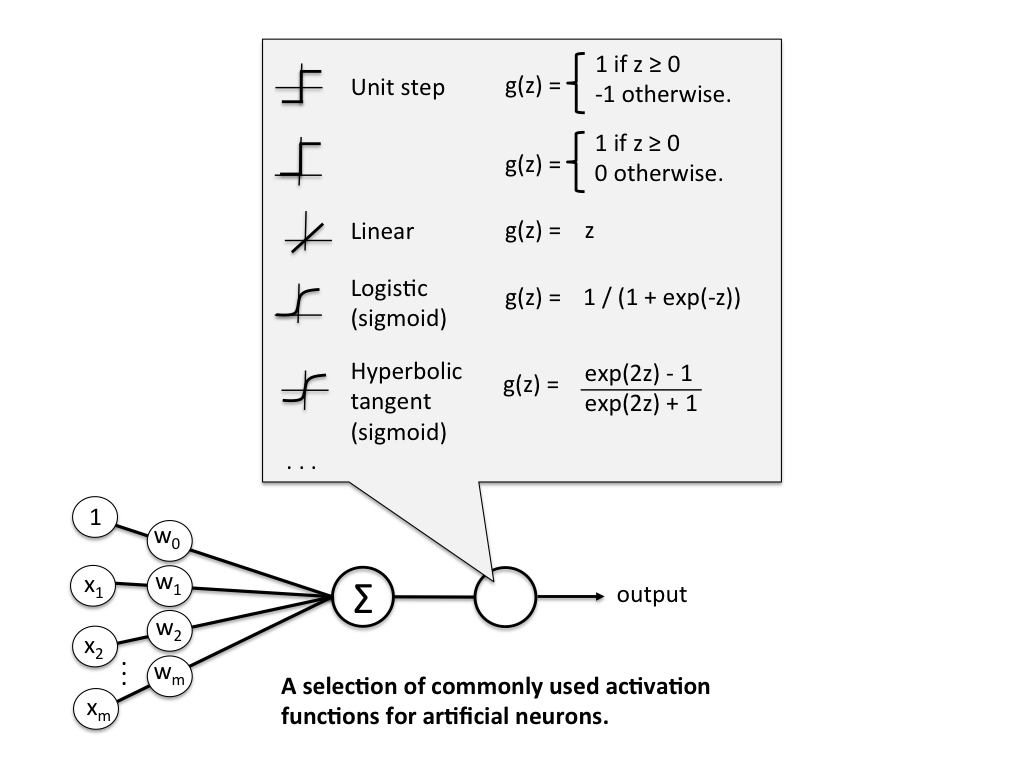
| Adaline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="guLrNzwEGP9L" colab={"base_uri": "https://localhost:8080/"} outputId="8e9e2a33-578c-49f2-8054-71c696a0f233"
import os
# Find the latest version of spark 3.0 from http://www.apache.org/dist/spark/ and enter as the spark version
# For example:
spark_version = 'spark-3.0.3'
# spark_version = 'spark-3.<enter version>'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !wget -q http://www.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + colab={"base_uri": "https://localhost:8080/"} id="llCHcEUolY5W" outputId="16bad997-85c7-4ac1-ed17-fc2e63d39a92"
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.9.jar
# + id="ztn9209sUvP3"
# starting Spark session
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
from pyspark.sql.types import IntegerType
spark = SparkSession.builder.appName("CloudETL").config("spark.driver.extraClassPath","/content/postgresql-42.2.9.jar").getOrCreate()
# + [markdown] id="a2z0OzGNXCeP"
# # Import Data
# + colab={"base_uri": "https://localhost:8080/"} id="GA397-iIb4lW" outputId="a2f3bcf8-79fc-4cf5-9fd5-6c2d25246d82"
# Read in data from S3 Buckets
from pyspark import SparkFiles
url="https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Mobile_Apps_v1_00.tsv.gz"
spark.sparkContext.addFile(url)
mobile_apps_df = spark.read.csv(SparkFiles.get("amazon_reviews_us_Mobile_Apps_v1_00.tsv.gz"), sep = r"\t", header = True, inferSchema = True)
# Show DataFrame
mobile_apps_df.show()
# + [markdown] id="7jl70yQ0XGbh"
# # Create and Clean New DataFrames for Customers, Products and Reviews
# + colab={"base_uri": "https://localhost:8080/"} id="Zztvm-t4ftaZ" outputId="9413a8ff-343a-4adb-c926-c3c021813b69"
# Create reviews dataframe to match mobile_apps_df table
apps_reviews_df = mobile_apps_df.select(["review_id", "customer_id", "product_id",
"product_parent", "review_date"])
apps_reviews_df.show()
# + colab={"base_uri": "https://localhost:8080/"} id="FTDs424AgVv4" outputId="24aed2a1-2995-45c9-a8f4-16f55d9cf0cd"
# checking the datatypes of each column
apps_reviews_df.printSchema()
# + id="2Z9l5HkmgxHZ"
# changing review_date to 'date' data type in "yyyy-mm-dd" format
apps_reviews_df = apps_reviews_df.withColumn("review_date",to_date(col("review_date"),"yyyy-mm-dd"))
# + colab={"base_uri": "https://localhost:8080/"} id="hUq3X1lJhPK5" outputId="23747ffe-097b-4bae-cead-b7eb65923adf"
# confirming change has been made
apps_reviews_df.printSchema()
# + colab={"base_uri": "https://localhost:8080/"} id="h88SaCvihTn1" outputId="1db384a7-b464-42b0-efa7-17cc3bf51f7d"
# counting number of rows in apps_reviews_df
apps_reviews_df.count()
# + colab={"base_uri": "https://localhost:8080/"} id="OQ4Gm0iXnTMP" outputId="08c50d1d-1539-4748-a22f-23df0464d1cd"
# Create products dataframe to match mobile_apps_df table and removing duplicates
apps_products_df = mobile_apps_df.select(["product_id", "product_title"])
apps_products_df = apps_products_df.drop_duplicates()
apps_products_df.show()
# + colab={"base_uri": "https://localhost:8080/"} id="NzendOO2nyjl" outputId="eaa1afc7-f2e8-4087-87cd-5bf3c60dbd9f"
# checking the datatypes of each column
apps_products_df.printSchema()
# + colab={"base_uri": "https://localhost:8080/"} id="UQA2Dbvja_mG" outputId="73b9e8bb-2757-46d2-c332-b8176a5fe743"
# counting number of rows in apps_products_df
apps_products_df.count()
# + colab={"base_uri": "https://localhost:8080/"} id="FMUvvUodrnHF" outputId="182e582d-613d-4077-9677-da9b111eb1d8"
# creating customer dataframe and adding "customer count" as a column
apps_customers_df = apps_reviews_df.groupBy("customer_id").agg({"customer_id": "count"}).withColumnRenamed("count(customer_id)", "customer_count")
apps_customers_df.show()
# + colab={"base_uri": "https://localhost:8080/"} id="HNoXq2nbbPo-" outputId="b112441c-e405-47df-ecf6-0498dfb30af0"
# checking the datatypes of each column
apps_customers_df.printSchema()
# + colab={"base_uri": "https://localhost:8080/"} id="fiQbGJBIbPzj" outputId="5ecaac1d-5015-48b4-94a9-87de7969e31d"
# counting number of rows in apps_customers_df
apps_customers_df.count()
# + [markdown] id="QPTM7ZjyXS5M"
# # Configuration for RDS Instance
# + id="mA2HeFsJeuQ3"
# Configure settings for RDS
## IMPORTANT : PLEASE ADD YOUR OWN CREDENTIALS FOR RDS ##
mode = "append"
jdbc_url="jdbc:postgresql://<connection string>:5432/<database-name>"
config = {"user":"username",
"password": "<password>",
"driver":"org.postgresql.Driver"}
# + [markdown] id="zm7KyP8fXX23"
# # Write Dataframes to RDS
# + id="cShXg837doD2"
# Write DataFrame to apps_customers table in RDS
apps_customers_df.write.jdbc(url=jdbc_url, table='apps_customers', mode=mode, properties=config)
# + id="d5cb8j01eTzj"
# Write dataframe to apps_products table in RDS
apps_products_df.write.jdbc(url=jdbc_url, table='apps_products', mode=mode, properties=config)
# + id="yAUEdsg_eWqj"
# Write dataframe to apps_review_id_table in RDS
apps_reviews_df.write.jdbc(url=jdbc_url, table='apps_review_id_table', mode=mode, properties=config)
| Level-1/mobileApplications.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="aIvb0JodYx9d"
# #### Import reina and other necessary libraries. Initialize a spark session.
# + id="9mGaEKdeU89A"
from reina.metalearners import SLearner
from reina.metalearners import TLearner
from reina.metalearners import XLearner
from pyspark.ml.regression import RandomForestRegressor
from pyspark.ml.classification import RandomForestClassifier
from pyspark.sql import SparkSession
# Initialize spark session
spark = SparkSession \
.builder \
.appName('Meta-Learner-Spark') \
.getOrCreate()
# + [markdown] id="xNHXEHMOY9PY"
# #### Read toy data. Replace .load() with the test_data.csv location -- this location could be a local one (no cluster) or it could be on a distributed storage system (e.g., HDFS)
#
# *Note: Code below assumes data generated by our script (for specifics, please refer to our toy data generation in the README). You could also modify the code accordingly to use your own data.*
# + id="DZyUrMTBVJAf"
df = spark.read \
.format("csv") \
.option('header', 'true') \
.load("test_data.csv") # replace with the location of test_data.csv
# Case variables to appropriate types
df = df.withColumn("var1", df.var1.cast("float"))
df = df.withColumn("var2", df.var2.cast("float"))
df = df.withColumn("var3", df.var3.cast("float"))
df = df.withColumn("var4", df.var4.cast("float"))
df = df.withColumn("var5", df.var5.cast("float"))
df = df.withColumn("treatment", df.treatment.cast("float"))
df = df.withColumn("outcome", df.outcome.cast("float"))
# Drop garbage column
df = df.drop("_c0")
# Print out dataframe schema
print(df.schema)
# + [markdown] id="1Xa3QohmXg2Y"
# ## S-leaner
# + id="BkMiSReQW2DE"
# Set up necessary parameters
treatments = ['treatment']
outcome = 'outcome'
# Arbitrary estimator. Can replace with other ML algo.
estimator = RandomForestRegressor()
# Fit S-learner
spark_slearner = SLearner()
spark_slearner.fit(data=df, treatments=treatments, outcome=outcome, estimator=estimator)
# Get heterogeneous treatment effects (cate for individual samples and ate for averaged treatment effect)
cate, ate = spark_slearner.effects()
print(cate)
print(ate)
# + [markdown] id="Q1fcO5LOXjH_"
# ## T-leaner
# + id="kn1QYEJPXWvM"
# Set up necessary parameters
treatments = ['treatment']
outcome = 'outcome'
# Arbitrary estimators. Can replace with other ML algo.
estimator_1 = RandomForestRegressor()
estimator_0 = RandomForestRegressor()
# Fit T-learner
spark_tlearner = TLearner()
spark_tlearner.fit(data=df, treatments=treatments, outcome=outcome,
estimator_0=estimator_0, estimator_1=estimator_1)
# Get heterogeneous treatment effects (cate for individual samples and ate for averaged treatment effect)
cate, ate = spark_tlearner.effects()
print(cate)
print(ate)
# + [markdown] id="bxZQdw1fXoPa"
# ## X-leaner
# + id="Qj1owlx-XYKo"
# Set up necessary parameters
treatments = ['treatment']
outcome = 'outcome'
# Arbitrary estimators. Can replace with other ML algo.
estimator_11 = RandomForestRegressor()
estimator_10 = RandomForestRegressor()
estimator_21 = RandomForestRegressor()
estimator_20 = RandomForestRegressor()
propensity_estimator = RandomForestClassifier()
# Fit X-learner
spark_xlearner = XLearner()
spark_xlearner.fit(data=df, treatments=treatments, outcome=outcome,
estimator_10=estimator_10, estimator_11=estimator_11,
estimator_20=estimator_20, estimator_21=estimator_21,
propensity_estimator=propensity_estimator)
# Get heterogeneous treatment effects (cate for individual samples and ate for averaged treatment effect)
cate, ate = spark_xlearner.effects()
print(cate)
print(ate)
| examples/notebooks/metalearner_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
'''
Modules required for imaging and data analysis
'''
import pandas as pd
import numpy as np
import matplotlib##
import matplotlib.pyplot as plt##
import scipy.interpolate
# %matplotlib inline
import os
import math
import seaborn as sns
import scipy
import skimage
from skimage import data
from skimage import io
import os
from skimage import filters
import math
from scipy.optimize import curve_fit
from scipy.stats.distributions import t
from skimage.filters import threshold_otsu, threshold_adaptive
from skimage.morphology import remove_small_objects
import statsmodels.api as sm
import bottleneck as bn
nanmean = bn.nanmean
# +
'''
Create a DataFrame to convert frames in minutes
'''
total_number_of_TimeIDs = 150 #number of frames
time_step = 5 #frame rate in minutes
TimeIDs = range(1, total_number_of_TimeIDs + 1, 1) #+1 because the last number is not accounted
Time_mins = []
for i in TimeIDs:
t = time_step*(i-1)
Time_mins.append(t)
time_conversion = pd.DataFrame({"TimeID" : TimeIDs, "Time" : Time_mins})
time_conversion
# +
'''
Create a list with the names of the .tif files to be loaded and analyzed
'''
PATH_to_the_TIFs = "Z:/Experiments/TORC1/20190506_wf(new)_SFP1-TOD6-pHtdGFP_agar_mm_rep4/Processed_tiffs/" #path to the folder containin the .tif files
all_files = os.listdir(PATH_to_the_TIFs)
tif_files = []
for f in all_files:
if f.endswith(".tif"):
tif_files.append(f)
tif_files
# +
'''
Creating a list of .csv files with the segmentation info created by BudJ
'''
PATH_to_the_TIFs2 = "Z:/Experiments/TORC1/20190506_wf(new)_SFP1-TOD6-pHtdGFP_agar_mm_rep4/Analysis/" # path to the folder containing the BudJ output files
files = []
for i in os.listdir(PATH_to_the_TIFs2):
if ".csv" in i:
prefix_position = "pos" + i[-22:-20]+"_"
files.append((i, prefix_position))
files
# +
'''
Creating a DataFrame containing the segmenation info from BudJ
'''
initial_table = pd.DataFrame({})
for f in files:
pos = pd.read_csv(f[0], header=0, index_col=0)
pos["Cell_pos"] = f[1] + pos["Cell"].map(str)
pos = pos.loc[:, ["TimeID", "Cell_pos", "volume", "x", "y", "major R", "minor r", "angle"]]
initial_table = pd.concat([initial_table, pos])
initial_table = pd.merge(initial_table, time_conversion, on="TimeID")
initial_table
# +
'''
Number of individual cells segmented by BudJ
'''
individual_cells = sorted(list(set(initial_table["Cell_pos"])))
len(individual_cells)
# +
'''
Creates a dictionary containing the budding events (in frame) for each cell
'''
buddings_SFP1 = {"pos01_1" : [1, 34, 53, 74, 99, 119],
"pos01_2" : [9, 34, 58, 79, 102, 125],
..........
}
# +
'''
Creates a dictionary containing the karyokinesis events (in frame) for each cell
'''
kariokinesis_SFP1 = {"pos01_1" : [17, 48, 68, 92, 113],
"pos01_2" : [26, 50, 73, 94, 116],
......
}
# -
print(len(buddings_SFP1))
print(len(kariokinesis_SFP1))
# +
'''
Creates a function that converts the ellipse parameters from BudJ into pixels unit
'''
def ellipse(time_point, BudJ_table, scaling_factor):
h = float(BudJ_table[BudJ_table["TimeID"] == time_point]['x'])/scaling_factor
k = float(BudJ_table[BudJ_table["TimeID"] == time_point]['y'])/scaling_factor
a = float(BudJ_table[BudJ_table["TimeID"] == time_point]["major R"])/scaling_factor
b = float(BudJ_table[BudJ_table["TimeID"] == time_point]["minor r"])/scaling_factor
A = float(BudJ_table[BudJ_table["TimeID"] == time_point]['angle'])*(math.pi/180)
return h, k, a, b, A
# -
def round_up_to_odd(f):
return int(np.ceil(f) // 2 * 2 + 1)
# +
### this code apply the mask defined by the BudJ segmentatio to each cell at each time point and measures
### the N/C ratio for that cell at every time point
scaling_factor = 0.16 #microns per pixel, 100x objective
initial_table_recalculated_SFP1 = pd.DataFrame({})
c = 0
for pos in ['01','02','03','04','05','06','07','08','09','10','11','12','13','14','15','16','17','18','19','20']: #list of all the XY position to be analyzed
filename = os.path.join(skimage.data_dir, PATH_to_the_TIFs+"20190506_wf(new)_sfp1-tod6_agar_mm_gl_rep3_xy"+pos+".nd2.tif")
cell_tif = io.imread(filename) # load the raw images files
for cell in individual_cells:
if 'pos'+pos in cell:
temp = initial_table[initial_table["Cell_pos"] == cell] #extract the BudJ info for a particular cell
temp = temp.sort_values(by="Time")
time_axis = list(temp["TimeID"]) #extract time axis
##List of all the parameters to be measured for each cell at each time point
GFP_cell = []
GFP_cyto1 = []
GFP_nucleus1 = []
Ratio1 = []
RFP_total = []
for t in time_axis:
t_in_tiff = int(t) - 1
h, k, a, b, A = ellipse(t, temp, scaling_factor)
#creating the mask corresponding to the BudJ ellipse
image = cell_tif[t_in_tiff,:,:,2] #Extract one channel, at time point "t_in_tiff", does not matter which channel
nrows, ncols = image.shape
row, col = np.ogrid[:nrows, :ncols]
### it creates a mask corresponding to the ellipse fitted by BudJ to segment the cell at this particular frame
inner_disk_mask = ((((col-h-1)*math.cos(A)+(row-k-1)*math.sin(A))**2)/(a**2) + (((col-h-1)*math.sin(A)-(row-k-1)*math.cos(A))**2)/(b**2) - 1 < 0)
###Calculate variables in the GFP channel like mean intensity, std,...
imageGFP = cell_tif[t_in_tiff,:,:,1] # extract GFP channel
imageGFP_corr = imageGFP[inner_disk_mask == True] # apply cell mask
mean_GFP = np.mean(imageGFP_corr) # calculate mean GFP intensity inside the cell mask
GFP_cell.append(mean_GFP)
GFP_std.append(np.std(imageGFP[inner_disk_mask == True])) # calculate the STD of the GFP intensity inside the cell mask
### Nuclear mask
imageRFP = cell_tif[t_in_tiff,:,:,2] # extract the RFP channel
imageRFP_corr = imageRFP*inner_disk_mask # apply cells mask
thr_mask = imageRFP_corr > 10 # use a simple intensity thrshold to create a mask of the nucleus
centroid1 = scipy.ndimage.measurements.center_of_mass((imageGFP*inner_disk_mask)*thr_mask) # calculate the centroid of the nuclear mask
centroid1 = np.nan_to_num(centroid1)
b1 = centroid1[1]
a1 = centroid1[0]+1
r = 3 # define radious for small nucelar mask, in pixels
r1 = 9 # define radious for big nucelar mask, in pixels
x1,y1 = np.ogrid[-a1: nrows-a1, -b1: ncols-b1]
disk_mask_nuc1 = x1*x1 +y1*y1 < r*r #create small, circular, nuclear mask
disk_mask_cyto1 = x1*x1 +y1*y1 < r1*r1 #create big, circular, nuclear mask,
diff1 = np.logical_and(disk_mask_cyto1, inner_disk_mask) #overlap big nucelar mask with whole cell mask to avoid including pixels outside the whole cell mask
mask_of_cytoplasm1 = inner_disk_mask ^ diff1 #define the cytosoli mask by subtractin the big nuclear mask from the whole cell mask
#RFP
RFP_total.append(np.sum(imageRFP_corr[thr_mask == True])) #calculate total RFP signal insied the small nuclear mask
#GFP in the nucleus
nucleus_mean1 = np.mean(imageGFP[disk_mask_nuc1 == True]) #calculate mean GFP signal inside the small nuclear mask, that is what we consider the nucelar GFP concentration
GFP_nucleus1.append(nucleus_mean1)
#GFP in the cytosol
cyto_mean1 = np.mean(imageGFP[mask_of_cytoplasm1 == True]) #calculate mean GFP signal inside the cytosolic mask, that is what we consider the cytosolic GFP concentraion
GFP_cyto1.append(cyto_mean1)
#Ratio
Ratio1.append((nucleus_mean1)/(cyto_mean1)) #calculate the N/C ratio
###save calculated variables in DataFrame
temp["GFP_cell"] = pd.Series(GFP_cell, index=temp.index)
temp["GFP_std"] = pd.Series(GFP_std, index=temp.index)
temp["GFP_nucleus1"] = pd.Series(GFP_nucleus1, index=temp.index)
temp["GFP_cyto1"] = pd.Series(GFP_cyto1, index=temp.index)
temp["Ratio1"] = pd.Series(Ratio1, index=temp.index)
temp["RFP_conc"] = pd.Series(RFP_conc, index=temp.index)
temp["RFP_total"] = pd.Series(RFP_total, index=temp.index)
initial_table_recalculated_SFP1 = pd.concat([initial_table_recalculated_SFP1, temp])
c += 1
print cell,
# -
initial_table_recalculated_SFP1 = initial_table_recalculated_SFP1.reset_index(drop="true")
initial_table_recalculated_SFP1[pd.isnull(initial_table_recalculated_SFP1).any(axis=1)]
initial_table_recalculated_SFP1.to_excel("Sfp1_agar_rep1.xlsx")
initial_table_recalculated_SFP1 = pd.read_excel("Sfp1_agar_rep1.xlsx")
initial_table_SFP1 = initial_table_recalculated_SFP1
# +
## convert budding events in minutes
buddings_control_SFP1 = {}
for cell in individual_cells_SFP1:
control = []
for i in buddings_SFP1[cell]:
budding_time = float(time_conversion[time_conversion["TimeID"] == i]["Time"])
control.append(i)
buddings_control_SFP1[cell] = control
# +
## convert karyokinesis events in minutes
kariokinesis_control_SFP1 = {}
for cell in individual_cells_SFP1:
control = []
for i in kariokinesis_SFP1[cell]:
kariokinesis_time = float(time_conversion[time_conversion["TimeID"] == i]["Time"])
control.append(i)
kariokinesis_control_SFP1[cell] = control
# +
'''alignment karyokinesis to karyokinesis
Create a list where for each cell the information regarding each cell cycle (karyokiensis event, budding event, next karyokinesis event
number of point in between the first part and the second part of each cell cycle, duration of each part of each cell cycle) are stored'''
cell_cycles_list_SFP1 = []
buddings_group_SFP1 = buddings_control_SFP1
kariokinesis_group_SFP1 = kariokinesis_control_SFP1
for cell in buddings_group_SFP1:
table = initial_table_SFP1[initial_table_SFP1["Cell_pos"] == cell] #extract all variables corresponding to a cell
buddings_of_the_cell = buddings_group_SFP1[cell]
kariokinesis_of_the_cell = kariokinesis_group_SFP1[cell]
for i in range(len(kariokinesis_of_the_cell)):
if i != len(kariokinesis_of_the_cell)-1:
print cell
start1 = float(time_conversion[time_conversion["TimeID"] == kariokinesis_of_the_cell[i]]["Time"])
end1 = float(time_conversion[time_conversion["TimeID"] == buddings_of_the_cell[i+1]]["Time"])### KEEP the FLOAT function here
start2 = float(time_conversion[time_conversion["TimeID"] == buddings_of_the_cell[i+1]]["Time"])
end2 = float(time_conversion[time_conversion["TimeID"] == kariokinesis_of_the_cell[i+1]]["Time"])### KEEP the FLOAT function here
duration1 = end1 - start1
duration2 = end2 - start2
time_points_for_aligning_at_bud1 = (1/duration1)*np.array(np.arange(start1-start1, end1-start1+1, 5))
time_points_for_aligning_at_bud2 = (1/duration2)*np.array(np.arange(start2-start2, end2-start2+1, 5))
cell_cycles_list_SFP1.append((cell, start1, end1, start2, end2, time_points_for_aligning_at_bud1, time_points_for_aligning_at_bud2, duration1, duration2))
# +
### plot the distributions of the cell cycle phases duration
# %matplotlib inline
import matplotlib.pyplot as plt
sg2m = []
g1 = []
tot = []
plt.figure(1,(9,9))
for cycles in cell_cycles_list_SFP1:
if (cycles[7]+cycles[8]) < 200: # exclude cell cycles longer than 200 minutes
dur1 = cycles[7] #G1 duration
dur2 = cycles[8] #S/G2/M duration
dur3 = cycles[7] + cycles[8]
sg2m.append(dur2)
g1.append(dur1)
tot.append(dur3)
plt.hist(tot,15,label= 'Full cell cycle duration')
plt.hist(g1,15,label = 'G1 duration')
plt.hist(sg2m,15,label = 'S/G2/M duration')
plt.xlabel("min")
plt.legend()
plt.xlim(0, 500)
plt.savefig("SFP1_cell_cycles_hist_k2k.png")
# -
print(np.mean(tot))
print(np.mean(g1))
print(np.mean(sg2m))
# +
#### in this code each variable for each cell is splitted according to the cell cycle events and each part is then interpolated
#### with a fixed number of points defined by the average duartion of G1 and S/G2/M and their ratio, the sum have to be 80.
variables = ["volume","GFP_cell","GFP_nucleus1","GFP_cyto1","Ratio1","GFP_std","RFP_total"] # variables to be aligned and interpolated
L = len(variables)
k=0
c = 1
all_time_points1 = np.linspace(0, 1, 80) ## total number of points used for the interpolation of each cell cycle series
all_time_points2 = np.linspace(0, 1, 27) ## numnber of points used to interpolate the first part (G1) of each cell cycle series,
all_time_points3 = np.linspace(0, 1, 54) ## numnber of points used to interpolate the second part (SG2M) of each cell cycle series
## +1 point that will be exclude since it is already contained in the fisrt part (G1) of the cell cycle
cc_df1_SFP1 = pd.DataFrame({})
cc_df_small1_SFP1 = pd.DataFrame({})
cell_cycle = 0
for cycle in cell_cycles_list_SFP1:
pos_cell = cycle[0]
start1 = cycle[1]
end1 = cycle[2]
start2 = cycle[3]
end2 = cycle[4]
time1 = cycle[5]
time2 = cycle[6]
new_time1 = all_time_points1
new_time2 = all_time_points2
new_time3 = all_time_points3
cell_cycle += 1
print pos_cell
for i in range(L):
variable = variables[i]
big_data_table = initial_table_SFP1
table = big_data_table[big_data_table["Cell_pos"] == pos_cell] # extract the variables corresponding to a cell
sensor1 = table[(table["Time"] >= start1) & (table["Time"] <= end1)][variable] #split a variable from karyokinesis to budding
sensor2 = table[(table["Time"] >= start2) & (table["Time"] <= end2)][variable] #split a variable from budding to karyokinesis
if len(sensor1) != len(time1):
continue
f1 = scipy.interpolate.interp1d(time1, sensor1) #linear interpolation
f2 = scipy.interpolate.interp1d(time2, sensor2) #linear interpolation
new_sensor1 = f1(new_time2)
new_sensor2 = f2(new_time3)
cc_df_small1_SFP1["TimeID"] = new_time1
cc_df_small1_SFP1[variable] = np.append(new_sensor1, new_sensor2[1:])
cc_df_small1_SFP1["Cell_cycle"] = cell_cycle
cc_df1_SFP1 = pd.concat([cc_df1_SFP1, cc_df_small1_SFP1])
| Single_cells_NC_ratio_and_alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.0 Introduction to conversational software
# +
import time
def respond(message):
print('BOT: I can hear you! You said {}'.format(message))
def send_message(message):
print('USER: {}'.format(message))
time.sleep(2)
return respond(message)
# -
send_message('Hello!')
# # 1.1 Creating a personality
# +
responses = {
"What's your name?": "My name is Echobot",
"What's the weather today?": "It's sunny!"
}
def respond(message):
if message in responses:
return responses[message]
# -
respond("What's your name?")
# +
responses = {
"What's today's weather?": "It's {} today"
}
weatherToday = 'cloudy'
def respond(message):
if message in responses:
return responses[message].format(weatherToday)
# -
respond("What's today's weather?")
# +
responses = {
"What's your name?": [
"My name echobot",
"They call me echobot",
"The name's bot, echobot"
]
}
import random
def respond(message):
if message in responses:
return random.choice(responses[message])
# -
respond("What's your name?")
# +
responses = ['Tell me more!', "Why do you think that?"]
import random
def respond(message):
return random.choice(responses)
# -
respond("I think you're really great")
# # 1.2 Text processing with regex
# +
import re
pattern = "Do you remember .*"
message = "Do you remember when you ate strawberries in the garden"
match = re.search(pattern, message)
if match:
print("String matches!")
# -
pattern = "if (.*)"
message = "What would happen if bots took over the world!"
match = re.search(pattern, message)
match.group(0)
match.group(1)
# +
import re
def swap_pronouns(phrase):
if 'I' in phrase:
return re.sub('I', 'you', phrase)
if 'my' in phrase:
return re.sub('my', 'your', phrase)
else:
return phrase
# -
swap_pronouns('I walk my dog')
pattern = "do you remember (.*)"
message = "do you remember when you ate strawberries in the garden"
phrase = re.search(pattern, message)
phrase.group(0)
phrase.group(1)
phrase = swap_pronouns(phrase.group(1))
phrase
# # 2.0 Understanding intents and entities
#
# - Intents
# - Entities: NER (Named entity recognition)
#
# ## Regular expression to recognize intents and exercises
# - Simpler than machine learning approaches
# - Highly computationally efficient
# - Debugging regex is difficult
re.search(r"(hello|hey|hi)", "hey there!") is not None
re.search(r"(hello|hey|hi)", "which one?") is not None
re.search(r"\b(hello|hey|hi)\b", "hey there!") is not None
re.search(r"\b(hello|hey|hi)\b", "which one?") is not None
# +
# Entity recognition
pattern = re.compile('[A-Z]{1}[a-z]*')
message = """
Mary is a friend of mine,
she studied at Oxford and
now works at Google
"""
# -
pattern.findall(message)
# # 2.1 Word vectors
#
# - Try to represent meaning of words
# - Words with similar context have similar vector
# - GloVe algorithm (cousin of word2vec)
# - spaCy
import spacy
# +
nlp = spacy.load('en')
nlp.vocab.vectors_length
# +
doc = nlp('hello can you help me?')
for token in doc:
print("{}: {}".format(token, token.vector[:3]))
# -
# ## Similarity
#
# - Direction of vectors matters
# - "Distance" between words = angle between the vectors
# - Cosine similarity:
# - 1: if vectors point in the same direction
# - 0: if they are perpendicular
# - -1: if they point in opposite direction
doc = nlp("cat")
doc.similarity(nlp("can"))
doc.similarity(nlp("dog"))
# # 2.2 Intents and classification
#
# - A classifier predicts the intent label from a sentence
# - Use training data to tune classifer
# - Use testing data to evaluate performance
# - Accuracy: fraction of correctly predicted labels
from collections import defaultdict
import gzip
import numpy as np
import os
import pandas as pd
import pickle
os.getcwd()
os.listdir()
os.uname()
sorted(os.listdir('data'))
# +
names = ['label', 'sentence']
intentsDF = pd.read_csv('data/atis/atis_intents.csv', names=names)
trainDF = pd.read_csv('data/atis/atis_intents_train.csv', names=names)
testDF = pd.read_csv('data/atis/atis_intents_test.csv', names=names)
# -
intentsDF.info()
intentsDF.head()
trainDF.info()
trainDF.head()
testDF.info()
testDF.head()
trainSentences = trainDF['sentence'].values.tolist()
trainSentences[:2]
trainLabels = trainDF['label'].values.tolist()
trainLabels[:2]
X_train_shape = (len(trainSentences), nlp.vocab.vectors_length)
X_train_shape
X_train = np.zeros(X_train_shape)
X_train
test = np.zeros((5000,4))
test
# ```python
# for sentence in trainSentences:
# X_train[1, :] = nlp(sentence).vector
# ```
# ## Nearest neighbor classification
#
# Simplest solution:
# - Look for the labeled example that's most similar
# - Use its intent as a best guess
from sklearn.metrics.pairwise import cosine_similarity
sampleTestMessage = testDF.iloc[0, 1]
sampleTestMessage
test_x = nlp(sampleTestMessage).vector
np.shape(test_x)
# ```python
# scores = [cosine_similarity(X[i,:], test_x) for i in range(len(trainSentences))]
#
# trainLabels[np.argmax(scores)]
# ```
# ## Support vector machines
#
# SVM / SVC
#
# ```python
# from sklearn.svm import SVC
# clf = SVC()
# clf.fit(X_train, y_train)
# y_pred = clf.predict(X_test)
# ```
# # 2.3 Entity extraction
#
# - Keywords don't work for entities you haven't seen before
# - Use contextual clues:
# - Spelling
# - Capitalization
# - Words occurring before and after
# - Pattern recognition
doc = nlp("my friend Mary has worked at Google since 2009")
doc
for ent in doc.ents:
print(ent.text, ent.label_)
# ## Roles
import re
pattern1 = re.compile('.* from (.*) to (.*)')
string1 = "I want a flight from Tel Aviv to Bucharest"
pattern1.match(string1)
pattern2 = re.compile('.* to (.*) from (.*)')
string2 = 'show me flights to Shanghai from Singapore'
pattern2.match(string2)
# ## Dependency parsing
doc = nlp('a flight to Shanghai from Singapore')
shanghai, singapore = doc[3], doc[5]
list(shanghai.ancestors)
list(singapore.ancestors)
# +
doc = nlp("let's see that jacket in red and some blue jeans")
for token in doc:
print(token.text, token.pos_, token.dep_)
# -
items = [doc[4], doc[10]]
items
colours = [doc[6], doc[9]]
colours
for color in colours:
print('The current color: {0}'.format(color))
for a in color.ancestors:
print(a)
for color in colours:
for tok in color.ancestors:
if tok in items:
print("color {} belongs to item {}".format(color, tok))
#Without this break, there will be 1 more statement
break
# # 2.4 Robust NLU with Rasa
#
# - Library for intent recognition and entity extraction
# - Based on spaCy, scikit-learn and other libraries
# - Built-in support for chatbot specific tasks
from rasa_nlu.training_data import load_data
trainDF.iloc[21:23,:]
intentsDF.iloc[21:23,:]
testDF.iloc[21:23,:]
# # 3. Virtual assistants and accessing data
# +
import sqlite3
conn = sqlite3.connect('data/hotels.db')
c = conn.cursor()
# -
c.execute('SELECT name FROM sqlite_master WHERE type="table"')
c.fetchall()
c.execute('SELECT * FROM hotels LIMIT 5')
c.fetchall()
c.execute('SELECT * FROM hotels WHERE area="south" AND price="hi"')
c.fetchall()
# #### db => pandas
import pandas as pd
def sqlite3_to_csv():
conn = sqlite3.connect('data/hotels.db')
c = conn.cursor()
c.execute('SELECT name FROM sqlite_master WHERE type="table"')
tables = c.fetchall()
for tup in tables:
tableName = tup[0]
table = pd.read_sql_query("SELECT * FROM %s" % tableName, conn)
filepath = 'data/' + tableName + '.csv'
table.to_csv(filepath, index_label='index')
conn.close()
# %%time
sqlite3_to_csv()
df = pd.read_csv('data/hotels.csv')
df.info()
df
# # 3.1. Exploring a DB with natural language
# # 3.2. Incremental slot filling and negation
doc = nlp('not sushi, maybe pizza?')
indices = [1,4]
ents, negated_ents = [], []
start = 0
for i in indices:
phrase = '{}'.format(doc[start:i])
if 'not' in phrase or "n't" in phrase:
negated_ents.append(doc[i])
else:
ents.append(doc[i])
start = i
negated_ents
ents
# # 4. Stateful bots
| building-chatbots-in-python/building-chatbots-in-python.ipynb |