code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Machine Learning Engineer Nanodegree
# ## Capstone Project
# <NAME>
# July 11 2017
# ## I. Definition
# ### Project Overview
#
# Banks struggle with many issues these days. One of these is a thorough understanding of their clients. Not only their background information is important but also their behaviour. Interest arises in whether they are involved in possibly unwanted behaviour (e.g. money laundering or terrorism financing) or increasing their risk of default (e.g. missing monthly payments).
# Machine learning is starting to help banks investigate their clients, provide new and better credit scores and predict which clients may run into payment issues.
# In this project we look at a bank's data. We use an old data set (from 1999) which includes almost all properties real banks have. Even though this data is not as rich as that of a bank nowadays, it will provide a nice introductions into the techniques used by data analysts working for large banks.
# The dataset is from the PKDD'99 and can be found here:
# http://lisp.vse.cz/pkdd99/berka.htm.
# The competition had the following original description:
# *The bank wants to improve their services. For instance, the bank managers have only vague idea, who is a good client (whom to offer some additional services) and who is a bad client (whom to watch carefully to minimize the bank loses). Fortunately, the bank stores data about their clients, the accounts (transactions within several months), the loans already granted, the credit cards issued. The bank managers hope to improve their understanding of customers and seek specific actions to improve services. A mere application of a discovery tool will not be convincing for them.*
#
#
# ## Problem Statement
# Our research will try to answer the following question:
# Using a customer's properties and transactional behaviour, can we predict whether or not he/she is able to pay future loan payments?
# To answer this question, we need to make some assumptions before we can attempt to answer this question. Some of these will not directly be clear but will once we dive deeper into the data.
# - We make a clear seperation between running and finished loans. In this, we see finsihed loans as those of which we have full information and running loans as the ones we want to have classification on.
# - All loans are equal in time. We look at all loans from start to finish. In this, we do not take into conern the seasonality between different years.
# Using these assumptions, we will manipulate the data to create features which we hope can be predictive for running and new loans.
# This will require a large amount of preprocessing. In this usecase, we will also try to use network features to predict whether we have enough interaction to build network-based features.
#
# ### Metrics
# In this project, we will use an F1-score to evaluate how well our model will be able to predict a future defaults. We will use the finished loans for both traning and testing data. As we will see that we have a relatively small amount of defaults and F1 score will give us a nice balance over accuracy and false positive ratio.
# The F1 score is defined as:
# $$F_1 = 2\cdot \frac{Precision \cdot Recall}{Precision + Recall}$$
# where
# $$ Precision = \frac{tp}{tp+fp}$$
# and
# $$ Recall = \frac{tp}{tp+fn}$$
# ## II. Analysis
#
# ### Data Exploration
# The data for this project is provided on the competition website. In this project, it is downloaded and handled in the `Get_data.py` file. The data has the following structure:
# <img src="images/data_struct.gif">
# In the `Get_data.py` file we deal with the merging and checking of the quality of the data. Any reader interested in downloading and checking the data for themselves can follow the steps in that file.
# The data is provided to us in a flat text file with `;` delimiter and `\n` EOL statements. Many of the headers/cells in the dataset are provided in Czech and translated to English for further use. The data is then loaded into pandas dataframes and merged into five files:
# - `client_info.cv` containing all the info on the client
# - `demographic_info` containing all the info on the demographics of the Czech republic
# - `Transaction_info` A file containing all transactions made for the accounts in the dataset
# - `order_info` A file containing info on the ordered products per client
# - `loan_info` A file containing the loan information
#
# All of these files are in the further steps reduced to clients and accounts which have actual loans. These steps are described in `Loan_payment_feature_engineering.ipynb`.
# For some relevant features, we can provide the plots as found in the data. This allows us to check the distribution and whether or not there is a connection to missed payments. In these plots, we compare the distributions for loans which are past debt and those which are not.
# We can give some quick statistics and plots:
#
# | Feature| mean | std | min | 25% | 50%| 75% | max |
# | ----- | ----- | ---| ----| ----| --- | --- | --- |
# | Amount | 151410 |113372 |4980| 66732 | 116928 | 210654 | 590820|
# | Balance | 44023 | 13793 |6690 | 33725| 44879 | 54396 | 79272|
# | Birthyear | 1958 | 13 |1935 | 1947| 1958 | 1969 | 1980|
# | Payments | 4190 | 2215 | 304 | 2477 | 3934 | 5813 | 9910|
#
# <img src="images/balance_amount.png" alt="Drawing" style="width: 400px;"/>
# <img src="images/birthyear_owner_payments.png" alt="Drawing" style="width: 400px;"/>
#
# We can see the following things:
# - Clients in debt seem to have higher monthly paymets but less balance on the bank.
#
# With these, we hope to be able to predict which loans will be in default, and which will run along just fine.
#
# ### Algorithms and Techniques
# In this section, we will try to answer the following questions:
# - What will be the metric to which we test?
# - How will we define our training and test sets?
# - Which features will we use?
# - Which models will we use?
#
# To tackle the first, remember that we want to predict future debt using past debt. Our data is highly skewed however, as we have 606 accounts without any debt and only 76 with debt. Hence, using our accuracy score may not be the best choice. We will go with the F1 score as it will provide us with a nice balance between precision, recall and accuracy.
# Now actually configuring the data to do what we want is quite difficult. As we want to predict 'future missed payments' we need to use data in which we can predict future payments (as our past_debt only shows us if there is any past debt). We do however have the missed counts in all the quarters which we can use. Thus the setup will be as follows:
# 1. Use the finished data as training set
# 2. Use the finshed data as test set.
# 3. For both these datasets, use the first 3 quarters as prediction for missed payment in the last.
# 4. Use a few running tests as vaildation.
# 5. Finish with predictions on the full running dataset.
#
# We will use the following features:
# - Loan amount
# - Loan duration
# - Loan payment heigth
# - Bank balance
# - Birthyear account owner
# - Gender
# - Startyear of the owner
#
# As for the models, we will use the usual suspects:
# - **Naive Bayes**: A model that leverages conditional probability. Using known distributions of data to predict related distributions. It is run by the main rule:
# $$ p(C_k \mid \mathbf {x}) = \frac {p(C_{k})\ p(\mathbf {x} \mid C_{k})}{p(\mathbf {x} )} $$ or in layman's terms:
# $$ \mbox{posterior} = \frac {{\mbox{prior}}\times {\mbox{likelihood}}}{\mbox{evidence}} $$
# In our case, we will use a gaussian kernel that makes the assumption that the data is somewhat normally distributed to estimate the probability density function.
#
# - **SVM**: A linear SVM tries to lay a line (or in more dimension a (hyper)plane) inbetween data in order to create a seperation. Each side of the line receives a different classification (in our case payment miss or not). This is then optimized to build the most robust and least error prone line.
# - **Logistic regression**: A classifier version of Linear regression. Instead of the normal formula
# $$ y = \beta_0 + \beta_1x + \epsilon $$
# with normally distributed epsilon we have
# $$ y={\begin{cases}1&\beta _{0}+\beta _{1}x+\varepsilon >0\\0&{\text{else}}\end{cases}}$$ with epsilon distributed as in the logistic distribution.
# - **Decision Tree**: An algorithm which in each step slices data based on the feature leaving the least entropy (a measure of impurity of data). It is called a decision tree as one can make decisions based on every step and builds a tree by splitting the data, leaving the leaves as the classified data.
# - **Bagging**: An algorithm that subsets the data and trains decision trees on those subsets. The idea is that this procedure improves the stability of the algorithm. The final output is the average classification over all sampled classifiers.
# - **AdaBoost**: Adaptive boosting, an algorithm that only selects only features that show predictive power. It then builds a weighted tree based on the weighted predictive power for each feature.
# - **Random Forest**: Extension of bagging by also subsetting random features and pruning the tree.
#
#
# ### Benchmark
# A a benchmark we will use the Naive Bayes (GaussianNB) classifier. As we will see later, it has an F1 score of 0.44. Even though this model is far more advanced than the industry standard, it is a good baseline.
# ## III. Methodology
#
#
# ### Data Preprocessing
# This project took some preprocessing. As described earlier a lot of effort had to be done wrestling the data into a single dataset and translating the Czech to English.
# At first the idea was to use the transaction network (with accounts as nodes and transactions as edges) to provide us with a lot of information. The dataset however, does not have the same key for the receiving and sending parties, making this network very sparsely connected. The small exercise in doing this can be found in the file `Network_analysis.ipynb`. As a matter of fact, the largest connected component only had 7 nodes as seen in the picture below:
# <img src="images/network.png" alt="Drawing" style="width: 400px;"/>
# In this picture, the bold parts are the receiving accounts and we see that no features can be derived from the features of the network. Finding datasets in which this is possible seems very difficult. One could try to use the Bitcoin ledger. By property of the crypto ledger, no other information can be derieved from the network, making the analysis quite useless.
# Therefore, I focused on the loan problem at hand. For this I made a lot of steps on the transactions and especially on the missing of payments. I started with all loans and looked at whether those who had missed payments actually had 'flat' months (months in which no loan was paid) to check if the loans were correctly identified. The following picture shows that this is true:
# <img src="images/flat_months.png" alt="Drawing" style="width: 400px;"/>
# Next, I decided to look at the difference between the ordered and paid amount each month. If this was smaller than zero, this would indicate a missed payment. If it was larger, it was a possible pre-payment (in the first month). This seemed to also happen:
# <img src="images/ordere_diff.png" alt="Drawing" style="width: 400px;"/>
#
# As we assumed that all loans are equal, we push them all back to start-date and see that all prepayments happen in the first month and we can use this data to construct our features:
# <img src="images/ordered_pushed.png" alt="Drawing" style="width: 400px;"/>
#
# After this, we can take the first 75% of the time for each loan to predict future missed payment, and use the last 25% as the input for the target variable.
#
# In the end, we did choose to not include card/demographic behaviour due to the lack of variance in the data.
#
# ### Implementation and refinement
# After all the feature engineering and exploratory analysis shown above, I moved on to looking at the global correlation structure. I rebuild a plot I often use when working in R to give the first insights.
# <img src="images/correlation.png" alt="Drawing" style="width: 600px;"/>
# Based on this, I was able to quickly spot correlation structures and together with the plots based on distributions with the target variable substet the required features.
# After this, there was the phase in which we could start doing machine learning. I decided to not go for a CV scheme, as with the low amount of actual delayed payments in the data, there was the risk of creating subsets with no accounts with delayed payments.
# To do a grid search I wrote my own little implementation (based on the GridSearchCV in skLearn) which does the job without cross-validation. It can be found in `gridSearch.py`. I have used the gridsearch for the SVM, logistic regression and decision trees. This greatly improved performance compared with naive implementations (often giving an F1-score of 0.0). In the results section, the best scoring parameter settings are given using Occam's razor.
#
# As stated above, I will test the following algorithms on the data:
# - Naive Bayes
# - SVM
# - Logistic regression
# - Decision Tree
# - Bagging
# - AdaBoost
# - Random Forest
#
# ##### Naive Bayes
# I decided to run a standard naive bayes to see how it would perform compared with other algorithms.
#
# ##### SVM
# I ran the SVM with both the linear and rbf kernal, a C-value of either 1 or 10 and balanced and unweighted classes.
#
# ##### Logistic regression
# With Logistic regression I went with both the L1 and L2 loss functions, a C-value of either 1 or 10 and balanced and unweighted classes.
#
# ##### Decision tree
# For the Decision tree, I ran with 0.25, 0.5 and 0.75 max features combined with a max depth of either 1, 2 or 3.
#
# ##### Bagging, Boosting and Random Forest
# All these will run on baisc settings with 1000 estimators.
#
# For all algorithms, I will use the skLearn implementations combined with the gridsearch function for those for whom it may help.
# Running the models gave no complications as my macbook pro was sufficiently fast to run them. It was however hard to find a model which gave a satisfying F1-score, but this will be discussed in more detail in the Results part of this report.
#
# #### Complications when running the algorithm
# The most difficult part of this project was preprocessing the data into a file that we could use. As loans run through time, have different length and different loan amounts and start/end dates, the data can't be simply fed to algorithms.
# I also had to choose whether or not to use past missed payments in the classifier. I ended up choosing not to (as then it won't be usefull in an onboarding setting) leaving somewhat troublesome model performance.
# ## IV. Results
#
# ### Model Evaluation and Validation
#
# Below is a table which shows both the F1-score for our models. We see that the F1-score is generally low. To deduce these scores, I have done several different tran/test combinations. In all of these combinations, the best score was 0.67 for the F1-score.
#
# | Model | Config | Features | Score |
# | ----- | ------- | -------- | ------|
# | Naive Bayes | Gaussian/default | all | 0.44 |
# | SVM | {'kernel': 'rbf', 'C': 1, 'class_weight': 'balanced'} | Startyear client, Duration, payments | 0.67 |
# | Logistic Regression | 'C': 1, 'class_weight': 'balanced', 'penalty': 'l2' | Amount, Payments, Startyear Client, Balance | 0.39|
# | Decision Tree | {'max_depth': 3, 'max_features': 0.75} | Amount, Balance | 0.33|
# | Bagging | base_estimatior: Cart, n_estimators: 1000 | all | 0.4 |
# | AdaBoost | n_estimators: 1000 | all | 0.5 |
# | Random Forest | n_estimators: 1000 | all | 0.4 |
#
# As our SVM seems to perfom the best, we choose to use it as our final model. In other checks, we found it to be a quite robust method, performing well in combination with the gridsearch function for various subsets.
#
# We chose the SVM with a radial basis function as kernel and a simple C parameter (the misclassification/simplicity tradeoff) of value 1. This combined with balanced class weights (due to skewness) provides a very solid model.
#
# The combination of both training and testing sets gives us a robust model, which I have tested on different splits. The score of only 0.67 makes the results not trustwordy enough to use in production. We do however have a low benchmark (no machine learning based predictions are often done by banks). Therefore, we may even say it is acceptable (but far from optimal).
#
# I would have liked a better model and this sure is work for future improvement. More data and more customers would cetrainly help in this case.
#
# ### Justification
#
# Our simple SVM outperforms the Naive Bayes classifier with an F1-score of 0.67 vs 0.44. This means that it has a significant improvement over the other model and we can therefore justify changing to an SVM implementation.
# ## V. Conclusion
#
# ### Free-Form Visualization
# Our final question is whether we can predict if running loans will get into trouble. Remember that we did not use past payment issues as predictor, loans which lack this information (newly formed) can then not be correctly classified. Therefore, we visualize how we classify running loans, based on their past debt.
# We see that this is not very good. No loans with past debt are classified correctly.
#
# <img src="images/freeform.png" alt="Drawing" style="width: 400px;"/>
#
#
# ### Reflection
# I started this project hoping to find payment issues using network properties. I found however that the data (or any data openly available) was not sufficient to provide me with both a loan/transactional dataset and interesting relationships between parties. Therefore I settled with a more traditional project.
# The research question *Using a customer's properties and transactional behaviour, can we predict whether or not he/she is able to pay future loan payments?* has shown difficult to answer with the given dataset.
# A lot of work has gone into manipulating data on the start of the project. Therefore, the data crunching was maybe on the heavy side and I could have spent more time in machine learning. But it was good to go through all the steps in Python for a change (our default language is R) and see where it may be more suitable than our default goto tools.
# The machine learning part was therefore somewhat straightforward, different models were tested but none seemed to be a very good predictor. For this, we may need more samples and features.
# The final model is somewhat dissapointing, but it is a good example of default machine learning not working for all problems.
#
# ### Improvement
# The model can be improved in several ways. The data can be cut up in a different way to generate other features. We can take into account the skewness better by either building a skewed training set or generating more samples (with bootstraps, autoencoders or a GAN). This may improve the model and provide us a better prediction for running loans.
# In the end, the model would benefit from more personal data, often not disclosed in public datasets which better captivates properties of clients.
#
| Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 119} id="WJykK-B_rnbw" outputId="21a203e4-7daf-4a98-cbc1-9d70da3190b7"
# Install TensorFlow
# # !pip install -q tensorflow-gpu==2.0.0-beta1
try:
# %tensorflow_version 2.x # Colab only.
except Exception:
pass
import tensorflow as tf
print(tf.__version__)
# + id="DkR-FgUGrt3f"
from tensorflow.keras.layers import Input, Dense
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, Adam
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="NBgpKjxXsBZn" outputId="f5026231-3dec-4517-d6bc-3de52d85b707"
# make the original data
series = np.sin(0.1*np.arange(200)) #+ np.random.randn(200)*0.1
# plot it
plt.plot(series)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="qnR4MAM9sGF1" outputId="910194bc-9e2d-4c49-8e2b-bc16ed4723ea"
### build the dataset
# let's see if we can use T past values to predict the next value
T = 10
X = []
Y = []
for t in range(len(series) - T):
x = series[t:t+T]
X.append(x)
y = series[t+T]
Y.append(y)
X = np.array(X).reshape(-1, T)
Y = np.array(Y)
N = len(X)
print("X.shape", X.shape, "Y.shape", Y.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="EESqey3TsODi" outputId="6a78666e-2dfd-4808-95be-c4123c799fce"
### try autoregressive linear model
i = Input(shape=(T,))
x = Dense(1)(i)
model = Model(i, x)
model.compile(
loss='mse',
optimizer=Adam(lr=0.1),
)
# train the RNN
r = model.fit(
X[:-N//2], Y[:-N//2],
epochs=80,
validation_data=(X[-N//2:], Y[-N//2:]),
)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="2BA0rLqrsw7i" outputId="3cb4fe21-fd26-4979-8b92-764ea3cdc2e4"
# Plot loss per iteration
import matplotlib.pyplot as plt
plt.plot(r.history['loss'], label='loss')
plt.plot(r.history['val_loss'], label='val_loss')
plt.legend()
# + id="DyyEx1Gx4Q7t"
# "Wrong" forecast using true targets
validation_target = Y[-N//2:]
validation_predictions = []
# index of first validation input
i = -N//2
while len(validation_predictions) < len(validation_target):
p = model.predict(X[i].reshape(1, -1))[0,0] # 1x1 array -> scalar
i += 1
# update the predictions list
validation_predictions.append(p)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="hb18Dr0O4ec9" outputId="42ab7683-fac0-4760-ea83-5f558e426781"
plt.plot(validation_target, label='forecast target')
plt.plot(validation_predictions, label='forecast prediction')
plt.legend()
# + id="j9Idhr4ss3g_"
# Forecast future values (use only self-predictions for making future predictions)
validation_target = Y[-N//2:]
validation_predictions = []
# first validation input
last_x = X[-N//2] # 1-D array of length T
while len(validation_predictions) < len(validation_target):
p = model.predict(last_x.reshape(1, -1))[0,0] # 1x1 array -> scalar
# update the predictions list
validation_predictions.append(p)
# make the new input
last_x = np.roll(last_x, -1)
last_x[-1] = p
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="i0QEZgwV3WPI" outputId="910810ee-6f5a-4d5d-d719-00824da69a6b"
plt.plot(validation_target, label='forecast target')
plt.plot(validation_predictions, label='forecast prediction')
plt.legend()
| 4. Time Series/AZ/Misc/TF2_0_Autoregressive_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Search for AIMA 4th edition
#
# Implementation of search algorithms and search problems for AIMA.
#
# # Problems and Nodes
#
# We start by defining the abstract class for a `Problem`; specific problem domains will subclass this. To make it easier for algorithms that use a heuristic evaluation function, `Problem` has a default `h` function (uniformly zero), and subclasses can define their own default `h` function.
#
# We also define a `Node` in a search tree, and some functions on nodes: `expand` to generate successors; `path_actions` and `path_states` to recover aspects of the path from the node.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import random
import heapq
import math
import sys
from collections import defaultdict, deque, Counter
from itertools import combinations
class Problem(object):
"""The abstract class for a formal problem. A new domain subclasses this,
overriding `actions` and `results`, and perhaps other methods.
The default heuristic is 0 and the default action cost is 1 for all states.
When you create an instance of a subclass, specify `initial`, and `goal` states
(or give an `is_goal` method) and perhaps other keyword args for the subclass."""
def __init__(self, initial=None, goal=None, **kwds):
self.__dict__.update(initial=initial, goal=goal, **kwds)
def actions(self, state): raise NotImplementedError
def result(self, state, action): raise NotImplementedError
def is_goal(self, state): return state == self.goal
def action_cost(self, s, a, s1): return 1
def h(self, node): return 0
def __str__(self):
return '{}({!r}, {!r})'.format(
type(self).__name__, self.initial, self.goal)
class Node:
"A Node in a search tree."
def __init__(self, state, parent=None, action=None, path_cost=0):
self.__dict__.update(state=state, parent=parent, action=action, path_cost=path_cost)
def __repr__(self): return '<{}>'.format(self.state)
def __len__(self): return 0 if self.parent is None else (1 + len(self.parent))
def __lt__(self, other): return self.path_cost < other.path_cost
failure = Node('failure', path_cost=math.inf) # Indicates an algorithm couldn't find a solution.
cutoff = Node('cutoff', path_cost=math.inf) # Indicates iterative deepening search was cut off.
def expand(problem, node):
"Expand a node, generating the children nodes."
s = node.state
for action in problem.actions(s):
s1 = problem.result(s, action)
cost = node.path_cost + problem.action_cost(s, action, s1)
yield Node(s1, node, action, cost)
def path_actions(node):
"The sequence of actions to get to this node."
if node.parent is None:
return []
return path_actions(node.parent) + [node.action]
def path_states(node):
"The sequence of states to get to this node."
if node in (cutoff, failure, None):
return []
return path_states(node.parent) + [node.state]
# -
# # Queues
#
# First-in-first-out and Last-in-first-out queues, and a `PriorityQueue`, which allows you to keep a collection of items, and continually remove from it the item with minimum `f(item)` score.
# +
FIFOQueue = deque
LIFOQueue = list
class PriorityQueue:
"""A queue in which the item with minimum f(item) is always popped first."""
def __init__(self, items=(), key=lambda x: x):
self.key = key
self.items = [] # a heap of (score, item) pairs
for item in items:
self.add(item)
def add(self, item):
"""Add item to the queuez."""
pair = (self.key(item), item)
heapq.heappush(self.items, pair)
def pop(self):
"""Pop and return the item with min f(item) value."""
return heapq.heappop(self.items)[1]
def top(self): return self.items[0][1]
def __len__(self): return len(self.items)
# -
# # Search Algorithms: Best-First
#
# Best-first search with various *f(n)* functions gives us different search algorithms. Note that A\*, weighted A\* and greedy search can be given a heuristic function, `h`, but if `h` is not supplied they use the problem's default `h` function (if the problem does not define one, it is taken as *h(n)* = 0).
# +
def best_first_search(problem, f):
"Search nodes with minimum f(node) value first."
node = Node(problem.initial)
frontier = PriorityQueue([node], key=f)
reached = {problem.initial: node}
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
for child in expand(problem, node):
s = child.state
if s not in reached or child.path_cost < reached[s].path_cost:
reached[s] = child
frontier.add(child)
return failure
def best_first_tree_search(problem, f):
"A version of best_first_search without the `reached` table."
frontier = PriorityQueue([Node(problem.initial)], key=f)
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
for child in expand(problem, node):
if not is_cycle(child):
frontier.add(child)
return failure
def g(n): return n.path_cost
def astar_search(problem, h=None):
"""Search nodes with minimum f(n) = g(n) + h(n)."""
h = h or problem.h
return best_first_search(problem, f=lambda n: g(n) + h(n))
def astar_tree_search(problem, h=None):
"""Search nodes with minimum f(n) = g(n) + h(n), with no `reached` table."""
h = h or problem.h
return best_first_tree_search(problem, f=lambda n: g(n) + h(n))
def weighted_astar_search(problem, h=None, weight=1.4):
"""Search nodes with minimum f(n) = g(n) + weight * h(n)."""
h = h or problem.h
return best_first_search(problem, f=lambda n: g(n) + weight * h(n))
def greedy_bfs(problem, h=None):
"""Search nodes with minimum h(n)."""
h = h or problem.h
return best_first_search(problem, f=h)
def uniform_cost_search(problem):
"Search nodes with minimum path cost first."
return best_first_search(problem, f=g)
def breadth_first_bfs(problem):
"Search shallowest nodes in the search tree first; using best-first."
return best_first_search(problem, f=len)
def depth_first_bfs(problem):
"Search deepest nodes in the search tree first; using best-first."
return best_first_search(problem, f=lambda n: -len(n))
def is_cycle(node, k=30):
"Does this node form a cycle of length k or less?"
def find_cycle(ancestor, k):
return (ancestor is not None and k > 0 and
(ancestor.state == node.state or find_cycle(ancestor.parent, k - 1)))
return find_cycle(node.parent, k)
# -
# # Other Search Algorithms
#
# Here are the other search algorithms:
# +
def breadth_first_search(problem):
"Search shallowest nodes in the search tree first."
node = Node(problem.initial)
if problem.is_goal(problem.initial):
return node
frontier = FIFOQueue([node])
reached = {problem.initial}
while frontier:
node = frontier.pop()
for child in expand(problem, node):
s = child.state
if problem.is_goal(s):
return child
if s not in reached:
reached.add(s)
frontier.appendleft(child)
return failure
def iterative_deepening_search(problem):
"Do depth-limited search with increasing depth limits."
for limit in range(1, sys.maxsize):
result = depth_limited_search(problem, limit)
if result != cutoff:
return result
def depth_limited_search(problem, limit=10):
"Search deepest nodes in the search tree first."
frontier = LIFOQueue([Node(problem.initial)])
result = failure
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
elif len(node) >= limit:
result = cutoff
elif not is_cycle(node):
for child in expand(problem, node):
frontier.append(child)
return result
def depth_first_recursive_search(problem, node=None):
if node is None:
node = Node(problem.initial)
if problem.is_goal(node.state):
return node
elif is_cycle(node):
return failure
else:
for child in expand(problem, node):
result = depth_first_recursive_search(problem, child)
if result:
return result
return failure
# -
path_states(depth_first_recursive_search(r2))
# # Bidirectional Best-First Search
# +
def bidirectional_best_first_search(problem_f, f_f, problem_b, f_b, terminated):
node_f = Node(problem_f.initial)
node_b = Node(problem_f.goal)
frontier_f, reached_f = PriorityQueue([node_f], key=f_f), {node_f.state: node_f}
frontier_b, reached_b = PriorityQueue([node_b], key=f_b), {node_b.state: node_b}
solution = failure
while frontier_f and frontier_b and not terminated(solution, frontier_f, frontier_b):
def S1(node, f):
return str(int(f(node))) + ' ' + str(path_states(node))
print('Bi:', S1(frontier_f.top(), f_f), S1(frontier_b.top(), f_b))
if f_f(frontier_f.top()) < f_b(frontier_b.top()):
solution = proceed('f', problem_f, frontier_f, reached_f, reached_b, solution)
else:
solution = proceed('b', problem_b, frontier_b, reached_b, reached_f, solution)
return solution
def inverse_problem(problem):
if isinstance(problem, CountCalls):
return CountCalls(inverse_problem(problem._object))
else:
inv = copy.copy(problem)
inv.initial, inv.goal = inv.goal, inv.initial
return inv
# +
def bidirectional_uniform_cost_search(problem_f):
def terminated(solution, frontier_f, frontier_b):
n_f, n_b = frontier_f.top(), frontier_b.top()
return g(n_f) + g(n_b) > g(solution)
return bidirectional_best_first_search(problem_f, g, inverse_problem(problem_f), g, terminated)
def bidirectional_astar_search(problem_f):
def terminated(solution, frontier_f, frontier_b):
nf, nb = frontier_f.top(), frontier_b.top()
return g(nf) + g(nb) > g(solution)
problem_f = inverse_problem(problem_f)
return bidirectional_best_first_search(problem_f, lambda n: g(n) + problem_f.h(n),
problem_b, lambda n: g(n) + problem_b.h(n),
terminated)
def proceed(direction, problem, frontier, reached, reached2, solution):
node = frontier.pop()
for child in expand(problem, node):
s = child.state
print('proceed', direction, S(child))
if s not in reached or child.path_cost < reached[s].path_cost:
frontier.add(child)
reached[s] = child
if s in reached2: # Frontiers collide; solution found
solution2 = (join_nodes(child, reached2[s]) if direction == 'f' else
join_nodes(reached2[s], child))
#print('solution', path_states(solution2), solution2.path_cost,
# path_states(child), path_states(reached2[s]))
if solution2.path_cost < solution.path_cost:
solution = solution2
return solution
S = path_states
#A-S-R + B-P-R => A-S-R-P + B-P
def join_nodes(nf, nb):
"""Join the reverse of the backward node nb to the forward node nf."""
#print('join', S(nf), S(nb))
join = nf
while nb.parent is not None:
cost = join.path_cost + nb.path_cost - nb.parent.path_cost
join = Node(nb.parent.state, join, nb.action, cost)
nb = nb.parent
#print(' now join', S(join), 'with nb', S(nb), 'parent', S(nb.parent))
return join
# +
#A , B = uniform_cost_search(r1), uniform_cost_search(r2)
#path_states(A), path_states(B)
# +
#path_states(append_nodes(A, B))
# -
# # TODO: RBFS
# # Problem Domains
#
# Now we turn our attention to defining some problem domains as subclasses of `Problem`.
# # Route Finding Problems
#
# 
#
# In a `RouteProblem`, the states are names of "cities" (or other locations), like `'A'` for Arad. The actions are also city names; `'Z'` is the action to move to city `'Z'`. The layout of cities is given by a separate data structure, a `Map`, which is a graph where there are vertexes (cities), links between vertexes, distances (costs) of those links (if not specified, the default is 1 for every link), and optionally the 2D (x, y) location of each city can be specified. A `RouteProblem` takes this `Map` as input and allows actions to move between linked cities. The default heuristic is straight-line distance to the goal, or is uniformly zero if locations were not given.
# +
class RouteProblem(Problem):
"""A problem to find a route between locations on a `Map`.
Create a problem with RouteProblem(start, goal, map=Map(...)}).
States are the vertexes in the Map graph; actions are destination states."""
def actions(self, state):
"""The places neighboring `state`."""
return self.map.neighbors[state]
def result(self, state, action):
"""Go to the `action` place, if the map says that is possible."""
return action if action in self.map.neighbors[state] else state
def action_cost(self, s, action, s1):
"""The distance (cost) to go from s to s1."""
return self.map.distances[s, s1]
def h(self, node):
"Straight-line distance between state and the goal."
locs = self.map.locations
return straight_line_distance(locs[node.state], locs[self.goal])
def straight_line_distance(A, B):
"Straight-line distance between two points."
return sum(abs(a - b)**2 for (a, b) in zip(A, B)) ** 0.5
# +
class Map:
"""A map of places in a 2D world: a graph with vertexes and links between them.
In `Map(links, locations)`, `links` can be either [(v1, v2)...] pairs,
or a {(v1, v2): distance...} dict. Optional `locations` can be {v1: (x, y)}
If `directed=False` then for every (v1, v2) link, we add a (v2, v1) link."""
def __init__(self, links, locations=None, directed=False):
if not hasattr(links, 'items'): # Distances are 1 by default
links = {link: 1 for link in links}
if not directed:
for (v1, v2) in list(links):
links[v2, v1] = links[v1, v2]
self.distances = links
self.neighbors = multimap(links)
self.locations = locations or defaultdict(lambda: (0, 0))
def multimap(pairs) -> dict:
"Given (key, val) pairs, make a dict of {key: [val,...]}."
result = defaultdict(list)
for key, val in pairs:
result[key].append(val)
return result
# +
# Some specific RouteProblems
romania = Map(
{('O', 'Z'): 71, ('O', 'S'): 151, ('A', 'Z'): 75, ('A', 'S'): 140, ('A', 'T'): 118,
('L', 'T'): 111, ('L', 'M'): 70, ('D', 'M'): 75, ('C', 'D'): 120, ('C', 'R'): 146,
('C', 'P'): 138, ('R', 'S'): 80, ('F', 'S'): 99, ('B', 'F'): 211, ('B', 'P'): 101,
('B', 'G'): 90, ('B', 'U'): 85, ('H', 'U'): 98, ('E', 'H'): 86, ('U', 'V'): 142,
('I', 'V'): 92, ('I', 'N'): 87, ('P', 'R'): 97},
{'A': ( 76, 497), 'B': (400, 327), 'C': (246, 285), 'D': (160, 296), 'E': (558, 294),
'F': (285, 460), 'G': (368, 257), 'H': (548, 355), 'I': (488, 535), 'L': (162, 379),
'M': (160, 343), 'N': (407, 561), 'O': (117, 580), 'P': (311, 372), 'R': (227, 412),
'S': (187, 463), 'T': ( 83, 414), 'U': (471, 363), 'V': (535, 473), 'Z': (92, 539)})
r0 = RouteProblem('A', 'A', map=romania)
r1 = RouteProblem('A', 'B', map=romania)
r2 = RouteProblem('N', 'L', map=romania)
r3 = RouteProblem('E', 'T', map=romania)
r4 = RouteProblem('O', 'M', map=romania)
# -
path_states(uniform_cost_search(r1)) # Lowest-cost path from Arab to Bucharest
path_states(breadth_first_search(r1)) # Breadth-first: fewer steps, higher path cost
# # Grid Problems
#
# A `GridProblem` involves navigating on a 2D grid, with some cells being impassible obstacles. By default you can move to any of the eight neighboring cells that are not obstacles (but in a problem instance you can supply a `directions=` keyword to change that). Again, the default heuristic is straight-line distance to the goal. States are `(x, y)` cell locations, such as `(4, 2)`, and actions are `(dx, dy)` cell movements, such as `(0, -1)`, which means leave the `x` coordinate alone, and decrement the `y` coordinate by 1.
# +
class GridProblem(Problem):
"""Finding a path on a 2D grid with obstacles. Obstacles are (x, y) cells."""
def __init__(self, initial=(15, 30), goal=(130, 30), obstacles=(), **kwds):
Problem.__init__(self, initial=initial, goal=goal,
obstacles=set(obstacles) - {initial, goal}, **kwds)
directions = [(-1, -1), (0, -1), (1, -1),
(-1, 0), (1, 0),
(-1, +1), (0, +1), (1, +1)]
def action_cost(self, s, action, s1): return straight_line_distance(s, s1)
def h(self, node): return straight_line_distance(node.state, self.goal)
def result(self, state, action):
"Both states and actions are represented by (x, y) pairs."
return action if action not in self.obstacles else state
def actions(self, state):
"""You can move one cell in any of `directions` to a non-obstacle cell."""
x, y = state
return {(x + dx, y + dy) for (dx, dy) in self.directions} - self.obstacles
class ErraticVacuum(Problem):
def actions(self, state):
return ['suck', 'forward', 'backward']
def results(self, state, action): return self.table[action][state]
table = dict(suck= {1:{5,7}, 2:{4,8}, 3:{7}, 4:{2,4}, 5:{1,5}, 6:{8}, 7:{3,7}, 8:{6,8}},
forward= {1:{2}, 2:{2}, 3:{4}, 4:{4}, 5:{6}, 6:{6}, 7:{8}, 8:{8}},
backward={1:{1}, 2:{1}, 3:{3}, 4:{3}, 5:{5}, 6:{5}, 7:{7}, 8:{7}})
# +
# Some grid routing problems
# The following can be used to create obstacles:
def random_lines(X=range(15, 130), Y=range(60), N=150, lengths=range(6, 12)):
"""The set of cells in N random lines of the given lengths."""
result = set()
for _ in range(N):
x, y = random.choice(X), random.choice(Y)
dx, dy = random.choice(((0, 1), (1, 0)))
result |= line(x, y, dx, dy, random.choice(lengths))
return result
def line(x, y, dx, dy, length):
"""A line of `length` cells starting at (x, y) and going in (dx, dy) direction."""
return {(x + i * dx, y + i * dy) for i in range(length)}
random.seed(42) # To make this reproducible
frame = line(-10, 20, 0, 1, 20) | line(150, 20, 0, 1, 20)
cup = line(102, 44, -1, 0, 15) | line(102, 20, -1, 0, 20) | line(102, 44, 0, -1, 24)
d1 = GridProblem(obstacles=random_lines(N=100) | frame)
d2 = GridProblem(obstacles=random_lines(N=150) | frame)
d3 = GridProblem(obstacles=random_lines(N=200) | frame)
d4 = GridProblem(obstacles=random_lines(N=250) | frame)
d5 = GridProblem(obstacles=random_lines(N=300) | frame)
d6 = GridProblem(obstacles=cup | frame)
d7 = GridProblem(obstacles=cup | frame | line(50, 35, 0, -1, 10) | line(60, 37, 0, -1, 17) | line(70, 31, 0, -1, 19))
# -
# # 8 Puzzle Problems
#
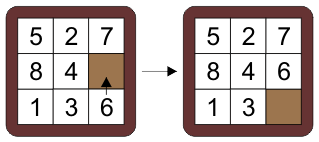
# 
#
# A sliding tile puzzle where you can swap the blank with an adjacent piece, trying to reach a goal configuration. The cells are numbered 0 to 8, starting at the top left and going row by row left to right. The pieces are numebred 1 to 8, with 0 representing the blank. An action is the cell index number that is to be swapped with the blank (*not* the actual number to be swapped but the index into the state). So the diagram above left is the state `(5, 2, 7, 8, 4, 0, 1, 3, 6)`, and the action is `8`, because the cell number 8 (the 9th or last cell, the `6` in the bottom right) is swapped with the blank.
#
# There are two disjoint sets of states that cannot be reached from each other. One set has an even number of "inversions"; the other has an odd number. An inversion is when a piece in the state is larger than a piece that follows it.
#
#
#
# +
class EightPuzzle(Problem):
""" The problem of sliding tiles numbered from 1 to 8 on a 3x3 board,
where one of the squares is a blank, trying to reach a goal configuration.
A board state is represented as a tuple of length 9, where the element at index i
represents the tile number at index i, or 0 if for the empty square, e.g. the goal:
1 2 3
4 5 6 ==> (1, 2, 3, 4, 5, 6, 7, 8, 0)
7 8 _
"""
def __init__(self, initial, goal=(0, 1, 2, 3, 4, 5, 6, 7, 8)):
assert inversions(initial) % 2 == inversions(goal) % 2 # Parity check
self.initial, self.goal = initial, goal
def actions(self, state):
"""The indexes of the squares that the blank can move to."""
moves = ((1, 3), (0, 2, 4), (1, 5),
(0, 4, 6), (1, 3, 5, 7), (2, 4, 8),
(3, 7), (4, 6, 8), (7, 5))
blank = state.index(0)
return moves[blank]
def result(self, state, action):
"""Swap the blank with the square numbered `action`."""
s = list(state)
blank = state.index(0)
s[action], s[blank] = s[blank], s[action]
return tuple(s)
def h1(self, node):
"""The misplaced tiles heuristic."""
return hamming_distance(node.state, self.goal)
def h2(self, node):
"""The Manhattan heuristic."""
X = (0, 1, 2, 0, 1, 2, 0, 1, 2)
Y = (0, 0, 0, 1, 1, 1, 2, 2, 2)
return sum(abs(X[s] - X[g]) + abs(Y[s] - Y[g])
for (s, g) in zip(node.state, self.goal) if s != 0)
def h(self, node): return self.h2(node)
def hamming_distance(A, B):
"Number of positions where vectors A and B are different."
return sum(a != b for a, b in zip(A, B))
def inversions(board):
"The number of times a piece is a smaller number than a following piece."
return sum((a > b and a != 0 and b != 0) for (a, b) in combinations(board, 2))
def board8(board, fmt=(3 * '{} {} {}\n')):
"A string representing an 8-puzzle board"
return fmt.format(*board).replace('0', '_')
class Board(defaultdict):
empty = '.'
off = '#'
def __init__(self, board=None, width=8, height=8, to_move=None, **kwds):
if board is not None:
self.update(board)
self.width, self.height = (board.width, board.height)
else:
self.width, self.height = (width, height)
self.to_move = to_move
def __missing__(self, key):
x, y = key
if x < 0 or x >= self.width or y < 0 or y >= self.height:
return self.off
else:
return self.empty
def __repr__(self):
def row(y): return ' '.join(self[x, y] for x in range(self.width))
return '\n'.join(row(y) for y in range(self.height))
def __hash__(self):
return hash(tuple(sorted(self.items()))) + hash(self.to_move)
# +
# Some specific EightPuzzle problems
e1 = EightPuzzle((1, 4, 2, 0, 7, 5, 3, 6, 8))
e2 = EightPuzzle((1, 2, 3, 4, 5, 6, 7, 8, 0))
e3 = EightPuzzle((4, 0, 2, 5, 1, 3, 7, 8, 6))
e4 = EightPuzzle((7, 2, 4, 5, 0, 6, 8, 3, 1))
e5 = EightPuzzle((8, 6, 7, 2, 5, 4, 3, 0, 1))
# +
# Solve an 8 puzzle problem and print out each state
for s in path_states(astar_search(e1)):
print(board8(s))
# -
# # Water Pouring Problems
#
# 
#
# In a [water pouring problem](https://en.wikipedia.org/wiki/Water_pouring_puzzle) you are given a collection of jugs, each of which has a size (capacity) in, say, litres, and a current level of water (in litres). The goal is to measure out a certain level of water; it can appear in any of the jugs. For example, in the movie *Die Hard 3*, the heroes were faced with the task of making exactly 4 gallons from jugs of size 5 gallons and 3 gallons.) A state is represented by a tuple of current water levels, and the available actions are:
# - `(Fill, i)`: fill the `i`th jug all the way to the top (from a tap with unlimited water).
# - `(Dump, i)`: dump all the water out of the `i`th jug.
# - `(Pour, i, j)`: pour water from the `i`th jug into the `j`th jug until either the jug `i` is empty, or jug `j` is full, whichever comes first.
class PourProblem(Problem):
"""Problem about pouring water between jugs to achieve some water level.
Each state is a tuples of water levels. In the initialization, also provide a tuple of
jug sizes, e.g. PourProblem(initial=(0, 0), goal=4, sizes=(5, 3)),
which means two jugs of sizes 5 and 3, initially both empty, with the goal
of getting a level of 4 in either jug."""
def actions(self, state):
"""The actions executable in this state."""
jugs = range(len(state))
return ([('Fill', i) for i in jugs if state[i] < self.sizes[i]] +
[('Dump', i) for i in jugs if state[i]] +
[('Pour', i, j) for i in jugs if state[i] for j in jugs if i != j])
def result(self, state, action):
"""The state that results from executing this action in this state."""
result = list(state)
act, i, *_ = action
if act == 'Fill': # Fill i to capacity
result[i] = self.sizes[i]
elif act == 'Dump': # Empty i
result[i] = 0
elif act == 'Pour': # Pour from i into j
j = action[2]
amount = min(state[i], self.sizes[j] - state[j])
result[i] -= amount
result[j] += amount
return tuple(result)
def is_goal(self, state):
"""True if the goal level is in any one of the jugs."""
return self.goal in state
# In a `GreenPourProblem`, the states and actions are the same, but instead of all actions costing 1, in these problems the cost of an action is the amount of water that flows from the tap. (There is an issue that non-*Fill* actions have 0 cost, which in general can lead to indefinitely long solutions, but in this problem there is a finite number of states, so we're ok.)
class GreenPourProblem(PourProblem):
"""A PourProblem in which the cost is the amount of water used."""
def action_cost(self, s, action, s1):
"The cost is the amount of water used."
act, i, *_ = action
return self.sizes[i] - s[i] if act == 'Fill' else 0
# +
# Some specific PourProblems
p1 = PourProblem((1, 1, 1), 13, sizes=(2, 16, 32))
p2 = PourProblem((0, 0, 0), 21, sizes=(8, 11, 31))
p3 = PourProblem((0, 0), 8, sizes=(7,9))
p4 = PourProblem((0, 0, 0), 21, sizes=(8, 11, 31))
p5 = PourProblem((0, 0), 4, sizes=(3, 5))
g1 = GreenPourProblem((1, 1, 1), 13, sizes=(2, 16, 32))
g2 = GreenPourProblem((0, 0, 0), 21, sizes=(8, 11, 31))
g3 = GreenPourProblem((0, 0), 8, sizes=(7,9))
g4 = GreenPourProblem((0, 0, 0), 21, sizes=(8, 11, 31))
g5 = GreenPourProblem((0, 0), 4, sizes=(3, 5))
# -
# Solve the PourProblem of getting 13 in some jug, and show the actions and states
soln = breadth_first_search(p1)
path_actions(soln), path_states(soln)
# # Pancake Sorting Problems
#
# Given a stack of pancakes of various sizes, can you sort them into a stack of decreasing sizes, largest on bottom to smallest on top? You have a spatula with which you can flip the top `i` pancakes. This is shown below for `i = 3`; on the top the spatula grabs the first three pancakes; on the bottom we see them flipped:
#
#
# 
#
# How many flips will it take to get the whole stack sorted? This is an interesting [problem](https://en.wikipedia.org/wiki/Pancake_sorting) that <NAME> has [written about](https://people.eecs.berkeley.edu/~christos/papers/Bounds%20For%20Sorting%20By%20Prefix%20Reversal.pdf). A reasonable heuristic for this problem is the *gap heuristic*: if we look at neighboring pancakes, if, say, the 2nd smallest is next to the 3rd smallest, that's good; they should stay next to each other. But if the 2nd smallest is next to the 4th smallest, that's bad: we will require at least one move to separate them and insert the 3rd smallest between them. The gap heuristic counts the number of neighbors that have a gap like this. In our specification of the problem, pancakes are ranked by size: the smallest is `1`, the 2nd smallest `2`, and so on, and the representation of a state is a tuple of these rankings, from the top to the bottom pancake. Thus the goal state is always `(1, 2, ..., `*n*`)` and the initial (top) state in the diagram above is `(2, 1, 4, 6, 3, 5)`.
#
class PancakeProblem(Problem):
"""A PancakeProblem the goal is always `tuple(range(1, n+1))`, where the
initial state is a permutation of `range(1, n+1)`. An act is the index `i`
of the top `i` pancakes that will be flipped."""
def __init__(self, initial):
self.initial, self.goal = tuple(initial), tuple(sorted(initial))
def actions(self, state): return range(2, len(state) + 1)
def result(self, state, i): return state[:i][::-1] + state[i:]
def h(self, node):
"The gap heuristic."
s = node.state
return sum(abs(s[i] - s[i - 1]) > 1 for i in range(1, len(s)))
c0 = PancakeProblem((2, 1, 4, 6, 3, 5))
c1 = PancakeProblem((4, 6, 2, 5, 1, 3))
c2 = PancakeProblem((1, 3, 7, 5, 2, 6, 4))
c3 = PancakeProblem((1, 7, 2, 6, 3, 5, 4))
c4 = PancakeProblem((1, 3, 5, 7, 9, 2, 4, 6, 8))
# Solve a pancake problem
path_states(astar_search(c0))
# # Jumping Frogs Puzzle
#
# In this puzzle (which also can be played as a two-player game), the initial state is a line of squares, with N pieces of one kind on the left, then one empty square, then N pieces of another kind on the right. The diagram below uses 2 blue toads and 2 red frogs; we will represent this as the string `'LL.RR'`. The goal is to swap the pieces, arriving at `'RR.LL'`. An `'L'` piece moves left-to-right, either sliding one space ahead to an empty space, or two spaces ahead if that space is empty and if there is an `'R'` in between to hop over. The `'R'` pieces move right-to-left analogously. An action will be an `(i, j)` pair meaning to swap the pieces at those indexes. The set of actions for the N = 2 position below is `{(1, 2), (3, 2)}`, meaning either the blue toad in position 1 or the red frog in position 3 can swap places with the blank in position 2.
#
# 
class JumpingPuzzle(Problem):
"""Try to exchange L and R by moving one ahead or hopping two ahead."""
def __init__(self, N=2):
self.initial = N*'L' + '.' + N*'R'
self.goal = self.initial[::-1]
def actions(self, state):
"""Find all possible move or hop moves."""
idxs = range(len(state))
return ({(i, i + 1) for i in idxs if state[i:i+2] == 'L.'} # Slide
|{(i, i + 2) for i in idxs if state[i:i+3] == 'LR.'} # Hop
|{(i + 1, i) for i in idxs if state[i:i+2] == '.R'} # Slide
|{(i + 2, i) for i in idxs if state[i:i+3] == '.LR'}) # Hop
def result(self, state, action):
"""An action (i, j) means swap the pieces at positions i and j."""
i, j = action
result = list(state)
result[i], result[j] = state[j], state[i]
return ''.join(result)
def h(self, node): return hamming_distance(node.state, self.goal)
JumpingPuzzle(N=2).actions('LL.RR')
j3 = JumpingPuzzle(N=3)
j9 = JumpingPuzzle(N=9)
path_states(astar_search(j3))
# # Reporting Summary Statistics on Search Algorithms
#
# Now let's gather some metrics on how well each algorithm does. We'll use `CountCalls` to wrap a `Problem` object in such a way that calls to its methods are delegated to the original problem, but each call increments a counter. Once we've solved the problem, we print out summary statistics.
# +
class CountCalls:
"""Delegate all attribute gets to the object, and count them in ._counts"""
def __init__(self, obj):
self._object = obj
self._counts = Counter()
def __getattr__(self, attr):
"Delegate to the original object, after incrementing a counter."
self._counts[attr] += 1
return getattr(self._object, attr)
def report(searchers, problems, verbose=True):
"""Show summary statistics for each searcher (and on each problem unless verbose is false)."""
for searcher in searchers:
print(searcher.__name__ + ':')
total_counts = Counter()
for p in problems:
prob = CountCalls(p)
soln = searcher(prob)
counts = prob._counts;
counts.update(actions=len(soln), cost=soln.path_cost)
total_counts += counts
if verbose: report_counts(counts, str(p)[:40])
report_counts(total_counts, 'TOTAL\n')
def report_counts(counts, name):
"""Print one line of the counts report."""
print('{:9,d} nodes |{:9,d} goal |{:5.0f} cost |{:8,d} actions | {}'.format(
counts['result'], counts['is_goal'], counts['cost'], counts['actions'], name))
# -
# Here's a tiny report for uniform-cost search on the jug pouring problems:
report([uniform_cost_search], [p1, p2, p3, p4, p5])
report((uniform_cost_search, breadth_first_search),
(p1, g1, p2, g2, p3, g3, p4, g4, p4, g4, c1, c2, c3))
# # Comparing heuristics
#
# First, let's look at the eight puzzle problems, and compare three different heuristics the Manhattan heuristic, the less informative misplaced tiles heuristic, and the uninformed (i.e. *h* = 0) breadth-first search:
# +
def astar_misplaced_tiles(problem): return astar_search(problem, h=problem.h1)
report([breadth_first_search, astar_misplaced_tiles, astar_search],
[e1, e2, e3, e4, e5])
# -
# We see that all three algorithms get cost-optimal solutions, but the better the heuristic, the fewer nodes explored.
# Compared to the uninformed search, the misplaced tiles heuristic explores about 1/4 the number of nodes, and the Manhattan heuristic needs just 2%.
#
# Next, we can show the value of the gap heuristic for pancake sorting problems:
report([astar_search, uniform_cost_search], [c1, c2, c3, c4])
# We need to explore 300 times more nodes without the heuristic.
#
# # Comparing graph search and tree search
#
# Keeping the *reached* table in `best_first_search` allows us to do a graph search, where we notice when we reach a state by two different paths, rather than a tree search, where we have duplicated effort. The *reached* table consumes space and also saves time. How much time? In part it depends on how good the heuristics are at focusing the search. Below we show that on some pancake and eight puzzle problems, the tree search expands roughly twice as many nodes (and thus takes roughly twice as much time):
report([astar_search, astar_tree_search], [e1, e2, e3, e4, r1, r2, r3, r4])
# # Comparing different weighted search values
#
# Below we report on problems using these four algorithms:
#
# |Algorithm|*f*|Optimality|
# |:---------|---:|:----------:|
# |Greedy best-first search | *f = h*|nonoptimal|
# |Extra weighted A* search | *f = g + 2 × h*|nonoptimal|
# |Weighted A* search | *f = g + 1.4 × h*|nonoptimal|
# |A* search | *f = g + h*|optimal|
# |Uniform-cost search | *f = g*|optimal|
#
# We will see that greedy best-first search (which ranks nodes solely by the heuristic) explores the fewest number of nodes, but has the highest path costs. Weighted A* search explores twice as many nodes (on this problem set) but gets 10% better path costs. A* is optimal, but explores more nodes, and uniform-cost is also optimal, but explores an order of magnitude more nodes.
# +
def extra_weighted_astar_search(problem): return weighted_astar_search(problem, weight=2)
report((greedy_bfs, extra_weighted_astar_search, weighted_astar_search, astar_search, uniform_cost_search),
(r0, r1, r2, r3, r4, e1, d1, d2, j9, e2, d3, d4, d6, d7, e3, e4))
# -
# We see that greedy search expands the fewest nodes, but has the highest path costs. In contrast, A\* gets optimal path costs, but expands 4 or 5 times more nodes. Weighted A* is a good compromise, using half the compute time as A\*, and achieving path costs within 1% or 2% of optimal. Uniform-cost is optimal, but is an order of magnitude slower than A\*.
#
# # Comparing many search algorithms
#
# Finally, we compare a host of algorihms (even the slow ones) on some of the easier problems:
report((astar_search, uniform_cost_search, breadth_first_search, breadth_first_bfs,
iterative_deepening_search, depth_limited_search, greedy_bfs,
weighted_astar_search, extra_weighted_astar_search),
(p1, g1, p2, g2, p3, g3, p4, g4, r0, r1, r2, r3, r4, e1))
# This confirms some of the things we already knew: A* and uniform-cost search are optimal, but the others are not. A* explores fewer nodes than uniform-cost.
# # Visualizing Reached States
#
# I would like to draw a picture of the state space, marking the states that have been reached by the search.
# Unfortunately, the *reached* variable is inaccessible inside `best_first_search`, so I will define a new version of `best_first_search` that is identical except that it declares *reached* to be `global`. I can then define `plot_grid_problem` to plot the obstacles of a `GridProblem`, along with the initial and goal states, the solution path, and the states reached during a search.
# +
def best_first_search(problem, f):
"Search nodes with minimum f(node) value first."
global reached # <<<<<<<<<<< Only change here
node = Node(problem.initial)
frontier = PriorityQueue([node], key=f)
reached = {problem.initial: node}
while frontier:
node = frontier.pop()
if problem.is_goal(node.state):
return node
for child in expand(problem, node):
s = child.state
if s not in reached or child.path_cost < reached[s].path_cost:
reached[s] = child
frontier.add(child)
return failure
def plot_grid_problem(grid, solution, reached=(), title='Search', show=True):
"Use matplotlib to plot the grid, obstacles, solution, and reached."
reached = list(reached)
plt.figure(figsize=(16, 10))
plt.axis('off'); plt.axis('equal')
plt.scatter(*transpose(grid.obstacles), marker='s', color='darkgrey')
plt.scatter(*transpose(reached), 1**2, marker='.', c='blue')
plt.scatter(*transpose(path_states(solution)), marker='s', c='blue')
plt.scatter(*transpose([grid.initial]), 9**2, marker='D', c='green')
plt.scatter(*transpose([grid.goal]), 9**2, marker='8', c='red')
if show: plt.show()
print('{} {} search: {:.1f} path cost, {:,d} states reached'
.format(' ' * 10, title, solution.path_cost, len(reached)))
def plots(grid, weights=(1.4, 2)):
"""Plot the results of 4 heuristic search algorithms for this grid."""
solution = astar_search(grid)
plot_grid_problem(grid, solution, reached, 'A* search')
for weight in weights:
solution = weighted_astar_search(grid, weight=weight)
plot_grid_problem(grid, solution, reached, '(b) Weighted ({}) A* search'.format(weight))
solution = greedy_bfs(grid)
plot_grid_problem(grid, solution, reached, 'Greedy best-first search')
def transpose(matrix): return list(zip(*matrix))
# -
plots(d3)
plots(d4)
# # The cost of weighted A* search
#
# Now I want to try a much simpler grid problem, `d6`, with only a few obstacles. We see that A* finds the optimal path, skirting below the obstacles. Weighterd A* with a weight of 1.4 finds the same optimal path while exploring only 1/3 the number of states. But weighted A* with weight 2 takes the slightly longer path above the obstacles, because that path allowed it to stay closer to the goal in straight-line distance, which it over-weights. And greedy best-first search has a bad showing, not deviating from its path towards the goal until it is almost inside the cup made by the obstacles.
plots(d6)
# In the next problem, `d7`, we see a similar story. the optimal path found by A*, and we see that again weighted A* with weight 1.4 does great and with weight 2 ends up erroneously going below the first two barriers, and then makes another mistake by reversing direction back towards the goal and passing above the third barrier. Again, greedy best-first makes bad decisions all around.
plots(d7)
# # Nondeterministic Actions
#
# To handle problems with nondeterministic problems, we'll replace the `result` method with `results`, which returns a collection of possible result states. We'll represent the solution to a problem not with a `Node`, but with a plan that consist of two types of component: sequences of actions, like `['forward', 'suck']`, and condition actions, like
# `{5: ['forward', 'suck'], 7: []}`, which says that if we end up in state 5, then do `['forward', 'suck']`, but if we end up in state 7, then do the empty sequence of actions.
# +
def and_or_search(problem):
"Find a plan for a problem that has nondterministic actions."
return or_search(problem, problem.initial, [])
def or_search(problem, state, path):
"Find a sequence of actions to reach goal from state, without repeating states on path."
if problem.is_goal(state): return []
if state in path: return failure # check for loops
for action in problem.actions(state):
plan = and_search(problem, problem.results(state, action), [state] + path)
if plan != failure:
return [action] + plan
return failure
def and_search(problem, states, path):
"Plan for each of the possible states we might end up in."
if len(states) == 1:
return or_search(problem, next(iter(states)), path)
plan = {}
for s in states:
plan[s] = or_search(problem, s, path)
if plan[s] == failure: return failure
return [plan]
# +
class MultiGoalProblem(Problem):
"""A version of `Problem` with a colllection of `goals` instead of one `goal`."""
def __init__(self, initial=None, goals=(), **kwds):
self.__dict__.update(initial=initial, goals=goals, **kwds)
def is_goal(self, state): return state in self.goals
class ErraticVacuum(MultiGoalProblem):
"""In this 2-location vacuum problem, the suck action in a dirty square will either clean up that square,
or clean up both squares. A suck action in a clean square will either do nothing, or
will deposit dirt in that square. Forward and backward actions are deterministic."""
def actions(self, state):
return ['suck', 'forward', 'backward']
def results(self, state, action): return self.table[action][state]
table = {'suck':{1:{5,7}, 2:{4,8}, 3:{7}, 4:{2,4}, 5:{1,5}, 6:{8}, 7:{3,7}, 8:{6,8}},
'forward': {1:{2}, 2:{2}, 3:{4}, 4:{4}, 5:{6}, 6:{6}, 7:{8}, 8:{8}},
'backward': {1:{1}, 2:{1}, 3:{3}, 4:{3}, 5:{5}, 6:{5}, 7:{7}, 8:{7}}}
# -
# Let's find a plan to get from state 1 to the goal of no dirt (states 7 or 8):
and_or_search(ErraticVacuum(1, {7, 8}))
# This plan says "First suck, and if we end up in state 5, go forward and suck again; if we end up in state 7, do nothing because that is a goal."
#
# Here are the plans to get to a goal state starting from any one of the 8 states:
{s: and_or_search(ErraticVacuum(s, {7,8}))
for s in range(1, 9)}
# # Comparing Algorithms on EightPuzzle Problems of Different Lengths
# +
from functools import lru_cache
def build_table(table, depth, state, problem):
if depth > 0 and state not in table:
problem.initial = state
table[state] = len(astar_search(problem))
for a in problem.actions(state):
build_table(table, depth - 1, problem.result(state, a), problem)
return table
def invert_table(table):
result = defaultdict(list)
for key, val in table.items():
result[val].append(key)
return result
goal = (0, 1, 2, 3, 4, 5, 6, 7, 8)
table8 = invert_table(build_table({}, 25, goal, EightPuzzle(goal)))
# +
def report8(table8, M, Ds=range(2, 25, 2), searchers=(breadth_first_search, astar_misplaced_tiles, astar_search)):
"Make a table of average nodes generated and effective branching factor"
for d in Ds:
line = [d]
N = min(M, len(table8[d]))
states = random.sample(table8[d], N)
for searcher in searchers:
nodes = 0
for s in states:
problem = CountCalls(EightPuzzle(s))
searcher(problem)
nodes += problem._counts['result']
nodes = int(round(nodes/N))
line.append(nodes)
line.extend([ebf(d, n) for n in line[1:]])
print('{:2} & {:6} & {:5} & {:5} && {:.2f} & {:.2f} & {:.2f}'
.format(*line))
def ebf(d, N, possible_bs=[b/100 for b in range(100, 300)]):
"Effective Branching Factor"
return min(possible_bs, key=lambda b: abs(N - sum(b**i for i in range(1, d+1))))
def edepth_reduction(d, N, b=2.67):
from statistics import mean
def random_state():
x = list(range(9))
random.shuffle(x)
return tuple(x)
meanbf = mean(len(e3.actions(random_state())) for _ in range(10000))
meanbf
# -
{n: len(v) for (n, v) in table30.items()}
# %time table30 = invert_table(build_table({}, 30, goal, EightPuzzle(goal)))
# %time report8(table30, 20, range(26, 31, 2))
# %time report8(table30, 20, range(26, 31, 2))
# +
from itertools import combinations
from statistics import median, mean
# Detour index for Romania
L = romania.locations
def ratio(a, b): return astar_search(RouteProblem(a, b, map=romania)).path_cost / sld(L[a], L[b])
nums = [ratio(a, b) for a,b in combinations(L, 2) if b in r1.actions(a)]
mean(nums), median(nums) # 1.7, 1.6 # 1.26, 1.2 for adjacent cities
# -
sld
| search4e.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="KU5so-4hURax"
# link colab to google drive directory where this project data is placed
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
# + id="sLiADzRYUDtm"
################ Need to set project path here !! #################
projectpath = # "/content/gdrive/MyDrive/GraphAttnProject/SpanTree [with start node]_[walklen=3]_[p=1,q=1]_[num_walks=50]/NIPS_Submission/"
# + id="0yY7zMECTaI5"
import os
os.chdir(projectpath)
os.getcwd()
# + id="lDTFxrCdpk6a"
# ! pip install dgl
import dgl
# + [markdown] id="GhSvd4yHmmaJ"
# # Load data
# + id="xIXBUNULmxtV"
from tqdm.notebook import tqdm, trange
import networkx as nx
import pickle
import numpy as np
import tensorflow as tf
import torch
print(tf.__version__)
# + id="dVBtU5vAm7Ik"
# load all train and validation graphs
train_graphs = pickle.load(open(f'graph_data/train_graphs.pkl', 'rb'))
val_graphs = pickle.load(open(f'graph_data/val_graphs.pkl', 'rb'))
# load all labels
train_labels = np.load('graph_data/train_labels.npy')
val_labels = np.load('graph_data/val_labels.npy')
# + id="i1RHKaJMnKpl"
#################. NEED TO SPECIFY THE RANDOM WALK LENGTH WE WANT TO USE ################
walk_len = 6 # we use GKAT with random walk length of 6 in this code file
# we could also change this parameter to load GKAT kernel generated from random walks with different lengths from 2 to 10.
#########################################################################################
# + id="rzPVFV3lntjd"
# here we load the frequency matriies (we could use this as raw data to do random feature mapping )
train_freq_mat = pickle.load(open(f'graph_data/GKAT_freq_mats_train_len={walk_len}.pkl', 'rb'))
val_freq_mat = pickle.load(open(f'graph_data/GKAT_freq_mats_val_len={walk_len}.pkl', 'rb'))
# + id="ydogONEsntqC"
# here we load the pre-calculated GKAT kernel
train_GKAT_kernel = pickle.load(open(f'graph_data/GKAT_dot_kernels_train_len={walk_len}.pkl', 'rb'))
val_GKAT_kernel = pickle.load(open(f'graph_data/GKAT_dot_kernels_val_len={walk_len}.pkl', 'rb'))
# + id="hcFF2MIuot12"
GKAT_masking = [train_GKAT_kernel, val_GKAT_kernel]
# + id="YRCTLcQdot17"
train_graphs = [ dgl.from_networkx(g) for g in train_graphs]
val_graphs = [ dgl.from_networkx(g) for g in val_graphs]
info = [train_graphs, train_labels, val_graphs, val_labels, GKAT_masking]
# + [markdown] id="X92f3fstqGTC"
# # START Training
# + id="RCTlzoRDhafQ"
import networkx as nx
import matplotlib.pyplot as plt
import time
import random
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader
from tqdm.notebook import tqdm, trange
import seaborn as sns
from random import shuffle
from multiprocessing import Pool
import multiprocessing
from functools import partial
from networkx.generators.classic import cycle_graph
import sys
import scipy
import scipy.sparse
#from CodeZip_ST import *
# + id="F-PpelEHvaUF"
from prettytable import PrettyTable
# this function will count the number of parameters in GKAT (will be used later in this code file)
def count_parameters(model):
table = PrettyTable(["Modules", "Parameters"])
total_params = 0
for name, parameter in model.named_parameters():
if not parameter.requires_grad: continue
param = parameter.numel()
table.add_row([name, param])
total_params+=param
print(table)
print(f"Total Trainable Params: {total_params}")
return total_params
# + [markdown] id="sHofaHfFnlmQ"
# # GKAT Testing
# + [markdown] id="1_6nPfVFXe25"
# ## GKAT model
# + id="bRkNVQccXg3U"
# this is the GKAT version adapted from the paper "graph attention networks"
class GKATLayer(nn.Module):
def __init__(self,
in_dim,
out_dim,
feat_drop=0.,
attn_drop=0.,
alpha=0.2,
agg_activation=F.elu):
super(GKATLayer, self).__init__()
self.feat_drop = nn.Dropout(feat_drop)
self.fc = nn.Linear(in_dim, out_dim, bias=False)
#torch.nn.init.xavier_uniform_(self.fc.weight)
#torch.nn.init.zeros_(self.fc.bias)
self.attn_l = nn.Parameter(torch.ones(size=(out_dim, 1)))
self.attn_r = nn.Parameter(torch.ones(size=(out_dim, 1)))
self.attn_drop = nn.Dropout(attn_drop)
self.activation = nn.LeakyReLU(alpha)
self.softmax = nn.Softmax(dim = 1)
self.agg_activation=agg_activation
def clean_data(self):
ndata_names = ['ft', 'a1', 'a2']
edata_names = ['a_drop']
for name in ndata_names:
self.g.ndata.pop(name)
for name in edata_names:
self.g.edata.pop(name)
def forward(self, feat, bg, counting_attn):
self.g = bg
h = self.feat_drop(feat)
head_ft = self.fc(h).reshape((h.shape[0], -1))
a1 = torch.mm(head_ft, self.attn_l) # V x 1
a2 = torch.mm(head_ft, self.attn_r) # V x 1
a = self.attn_drop(a1 + a2.transpose(0, 1))
a = self.activation(a)
maxes = torch.max(a, 1, keepdim=True)[0]
a_ = a - maxes # we could subtract max to make the attention matrix bounded. (not feasible for random feature mapping decomposition)
a_nomi = torch.mul(torch.exp(a_), counting_attn.float())
a_deno = torch.sum(a_nomi, 1, keepdim=True)
a_nor = a_nomi/(a_deno+1e-9)
ret = torch.mm(a_nor, head_ft)
if self.agg_activation is not None:
ret = self.agg_activation(ret)
return ret
# this is the GKAT version adapted from the paper "attention is all you need"
class GKATLayer(nn.Module):
def __init__(self, in_dim, out_dim, feat_drop=0., attn_drop=0., alpha=0.2, agg_activation=F.elu):
super(GKATLayer, self).__init__()
self.feat_drop = feat_drop #nn.Dropout(feat_drop, training=self.training)
self.attn_drop = attn_drop #nn.Dropout(attn_drop)
self.fc_Q = nn.Linear(in_dim, out_dim, bias=False)
self.fc_K = nn.Linear(in_dim, out_dim, bias=False)
self.fc_V = nn.Linear(in_dim, out_dim, bias=False)
self.softmax = nn.Softmax(dim = 1)
self.agg_activation=agg_activation
def forward(self, feat, bg, counting_attn):
h = F.dropout(feat, p=self.feat_drop, training=self.training)
Q = self.fc_Q(h).reshape((h.shape[0], -1))
K = self.fc_K(h).reshape((h.shape[0], -1))
V = self.fc_V(h).reshape((h.shape[0], -1))
logits = F.dropout( torch.matmul( Q, torch.transpose(K,0,1) ) , p=self.attn_drop, training=self.training) / np.sqrt(Q.shape[1])
maxes = torch.max(logits, 1, keepdim=True)[0]
logits = logits - maxes
a_nomi = torch.mul(torch.exp( logits ), counting_attn.float())
a_deno = torch.sum(a_nomi, 1, keepdim=True)
a_nor = a_nomi/(a_deno+1e-9)
ret = torch.mm(a_nor, V)
if self.agg_activation is not None:
ret = self.agg_activation(ret)
return ret
# + id="vE9B2J45XiJd"
class GKATClassifier(nn.Module):
def __init__(self, in_dim, hidden_dim, num_heads, n_classes, feat_drop_=0.,
attn_drop_=0.,):
super(GKATClassifier, self).__init__()
self.num_heads = num_heads
self.hidden_dim = hidden_dim
self.layers = nn.ModuleList([
nn.ModuleList([GKATLayer(in_dim, hidden_dim[0], feat_drop = feat_drop_, attn_drop = attn_drop_, agg_activation=F.elu) for _ in range(num_heads)]),
nn.ModuleList([GKATLayer(hidden_dim[0] * num_heads, hidden_dim[-1], feat_drop = feat_drop_, attn_drop = attn_drop_, agg_activation=F.elu) for _ in range(1)])
])
self.classify = nn.Linear(hidden_dim[-1] * 1, n_classes)
self.softmax = nn.Softmax(dim = 1)
def forward(self, bg, counting_attn, normalize = 'normal'):
h = bg.in_degrees().view(-1, 1).float() # use degree as features
num_nodes = h.shape[0]
features = h.numpy().flatten()
if normalize == 'normal':
mean_ = np.mean(features)
std_ = np.std(features)
h = (h - mean_)/(std_+1e-9)
for i, gnn in enumerate(self.layers):
all_h = []
for j, att_head in enumerate(gnn):
all_h.append(att_head(h, bg, counting_attn))
h = torch.squeeze(torch.cat(all_h, dim=1))
bg.ndata['h'] = h
hg = dgl.mean_nodes(bg, 'h')
return self.classify(hg)
# + id="eQY1EcS7xtYT"
# the following are the parameters we used in GKAT version adapted from "graph attention networks"
method = 'GKAT'
runtimes = 15 # the number of repeats
num_classes = 2
num_features = [4, 4] # use hidden dimension of 4 in each attention head
num_heads = 8 # use 8 heads
num_layers = 2 # use a two layer GKAT model
feature_drop = 0
atten_drop = 0
epsilon = 1e-4
start_tol = 499
tolerance = 80
max_epoch = 500
batch_size = 128
learning_rate = 0.005
# + id="TwRYodt8zf7S"
# the following are the parameters we used in GKAT version adapted from "attention is all you need"
method = 'GKAT'
runtimes = 15
num_classes = 2
num_features = [4, 2]
num_heads = 7
num_layers = 2
feature_drop = 0
atten_drop = 0
epsilon = 1e-4
start_tol = 499
tolerance = 80
max_epoch = 500
batch_size = 128
learning_rate = 0.005
# + id="LT0BHPfQ0IWZ"
all_GKAT_train_losses = []
all_GKAT_train_acc = []
all_GKAT_val_losses = []
all_GKAT_val_acc = []
ckpt_file = f'results_{num_layers}layers/{method}/{method}_ckpt.pt'
for runtime in trange(runtimes):
train_graphs, train_labels, val_graphs, val_labels, GKAT_masking = info
train_GKAT_masking, val_GKAT_masking = GKAT_masking
# Create model
model = GKATClassifier(1, num_features, num_heads, num_classes, feat_drop_ = feature_drop, attn_drop_ = atten_drop)
for p in model.parameters():
if p.dim() > 1:
nn.init.xavier_uniform(p)
count_parameters(model)
#model.apply(init_weights)
loss_func = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=learning_rate, weight_decay=1e-5)
model.train()
epoch_train_losses_GKAT = []
epoch_train_acc_GKAT = []
epoch_val_losses_GKAT = []
epoch_val_acc_GKAT = []
num_batches = int(len(train_graphs)/batch_size)
epoch = 0
nan_found = 0
tol = 0
while True:
if nan_found:
break
epoch_loss = 0
epoch_acc = 0
''' Training '''
for iter in range(num_batches):
#for iter in range(2):
predictions = []
labels = torch.empty(batch_size)
rand_indices = np.random.choice(len(train_graphs), batch_size, replace=False)
for b in range(batch_size):
predictions.append(model(train_graphs[rand_indices[b]], torch.Tensor(train_GKAT_masking[rand_indices[b]])))
if torch.isnan(predictions[b][0])[0]:
print('NaN found.')
break
#print(predictions[b].detach().numpy())
labels[b] = train_labels[rand_indices[b]]
acc = 0
for k in range(len(predictions)):
if predictions[k][0][0]>predictions[k][0][1] and labels[k]==0:
acc += 1
elif predictions[k][0][0]<=predictions[k][0][1] and labels[k]==1:
acc += 1
acc /= len(predictions)
epoch_acc += acc
predictions = torch.squeeze(torch.stack(predictions))
if torch.any(torch.isnan(predictions)):
print('NaN found.')
nan_found = 1
break
loss = loss_func(predictions, labels.long())
optimizer.zero_grad()
loss.backward()
optimizer.step()
epoch_loss += loss.detach().item()
epoch_acc /= (iter + 1)
epoch_loss /= (iter + 1)
val_acc = 0
val_loss = 0
predictions_val = []
for b in range(len(val_graphs)):
predictions_val.append(model(val_graphs[b], torch.Tensor(val_GKAT_masking[b])))
for k in range(len(predictions_val)):
if predictions_val[k][0][0]>predictions_val[k][0][1] and val_labels[k]==0:
val_acc += 1
elif predictions_val[k][0][0]<=predictions_val[k][0][1] and val_labels[k]==1:
val_acc += 1
val_acc /= len(val_graphs)
predictions_val = torch.squeeze(torch.stack(predictions_val))
loss = loss_func(predictions_val, torch.tensor(val_labels).long())
val_loss += loss.detach().item()
if len(epoch_val_losses_GKAT) ==0:
try:
os.remove(f'{projectpath}{ckpt_file}')
except:
pass
torch.save(model, f'{projectpath}{ckpt_file}')
print('Epoch {}, acc{:.2f}, loss {:.4f}, tol {}, val_acc{:.2f}, val_loss{:.4f} -- checkpoint saved'.format(epoch, epoch_acc, epoch_loss, tol, val_acc, val_loss))
elif (np.min(epoch_val_losses_GKAT) >= val_loss) and (np.max(epoch_val_acc_GKAT) <= val_acc):
torch.save(model, f'{projectpath}{ckpt_file}')
print('Epoch {}, acc{:.2f}, loss {:.4f}, tol {}, val_acc{:.2f}, val_loss{:.4f} -- checkpoint saved'.format(epoch, epoch_acc, epoch_loss, tol, val_acc, val_loss))
else:
print('Epoch {}, acc{:.2f}, loss {:.4f}, tol {}, val_acc{:.2f}, val_loss{:.4f}'.format(epoch, epoch_acc, epoch_loss, tol, val_acc, val_loss))
if epoch > start_tol:
if np.min(epoch_val_losses_GKAT) <= val_loss:
tol += 1
if tol == tolerance:
print('Loss do not decrease')
break
else:
if np.abs(epoch_val_losses_GKAT[-1] - val_loss)<epsilon:
print('Converge steadily')
break
tol = 0
if epoch > max_epoch:
print("Reach Max Epoch Number")
break
epoch += 1
epoch_train_acc_GKAT.append(epoch_acc)
epoch_train_losses_GKAT.append(epoch_loss)
epoch_val_acc_GKAT.append(val_acc)
epoch_val_losses_GKAT.append(val_loss)
all_GKAT_train_acc.append(epoch_train_acc_GKAT)
all_GKAT_train_losses.append(epoch_train_losses_GKAT)
all_GKAT_val_acc.append(epoch_val_acc_GKAT)
all_GKAT_val_losses.append(epoch_val_losses_GKAT)
# save results from current repeat to the following file
np.save(f'{projectpath}results_{num_layers}layers/epoch_train_acc_{method}_walklen{walk_len}_run{runtime}.npy', epoch_train_acc_GKAT)
np.save(f'{projectpath}results_{num_layers}layers/epoch_val_acc_{method}_walklen{walk_len}_run{runtime}.npy', epoch_val_acc_GKAT)
np.save(f'{projectpath}results_{num_layers}layers/epoch_train_losses_{method}_walklen{walk_len}_run{runtime}.npy', epoch_train_losses_GKAT)
np.save(f'{projectpath}results_{num_layers}layers/epoch_val_losses_{method}_walklen{walk_len}_run{runtime}.npy', epoch_val_losses_GKAT)
# all all results to the following file
np.save(f'{projectpath}results_{num_layers}layers/all_{method}_walklen{walk_len}_train_losses.npy', all_GKAT_train_losses)
np.save(f'{projectpath}results_{num_layers}layers/all_{method}_walklen{walk_len}_train_acc.npy', all_GKAT_train_acc)
np.save(f'{projectpath}results_{num_layers}layers/all_{method}_walklen{walk_len}_val_losses.npy', all_GKAT_val_losses)
np.save(f'{projectpath}results_{num_layers}layers/all_{method}_walklen{walk_len}_val_acc.npy', all_GKAT_val_acc)
# + id="16gycyQwFWDt"
| Synthetic_InducedCycle/InducedCycle_GKAT_2layer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering for dataset exploration
# > A Summary of lecture "Unsupervised Learning with scikit-learn", via datacamp
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Datacamp, Machine_Learning]
# - image: images/kmeans-centroid.png
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# ## Unsupervised Learning
# - Unsupervised Learning
# - Finds patterns in data (E.g., clustering customers by their purchase)
# - Compressing the data using purchase patterns (dimension reduction)
# - Supervised vs unsupervised learning
# - Supervised learning finds patterns for a prediction task
#
# e.g., classify tumors as benign or cancerous (labels)
# - Unsupervised learning finds patterns in data, but without a specific prediction task in mind
# - K-means clustering
# - Finds clusters of samples
# - Number of clusters must be specified
# - Cluster labels for new samples
# - New samples can be assigned to existing clusters
# - k-means remembers the mean of each cluster (the "centroids")
# - Finds the nearest centroid to each new sample
# ### How many clusters?
points = np.array([[ 0.06544649, -0.76866376],
[-1.52901547, -0.42953079],
[ 1.70993371, 0.69885253],
[ 1.16779145, 1.01262638],
[-1.80110088, -0.31861296],
[-1.63567888, -0.02859535],
[ 1.21990375, 0.74643463],
[-0.26175155, -0.62492939],
[-1.61925804, -0.47983949],
[-1.84329582, -0.16694431],
[ 1.35999602, 0.94995827],
[ 0.42291856, -0.7349534 ],
[-1.68576139, 0.10686728],
[ 0.90629995, 1.09105162],
[-1.56478322, -0.84675394],
[-0.0257849 , -1.18672539],
[ 0.83027324, 1.14504612],
[ 1.22450432, 1.35066759],
[-0.15394596, -0.71704301],
[ 0.86358809, 1.06824613],
[-1.43386366, -0.2381297 ],
[ 0.03844769, -0.74635022],
[-1.58567922, 0.08499354],
[ 0.6359888 , -0.58477698],
[ 0.24417242, -0.53172465],
[-2.19680359, 0.49473677],
[ 1.0323503 , -0.55688 ],
[-0.28858067, -0.39972528],
[ 0.20597008, -0.80171536],
[-1.2107308 , -0.34924109],
[ 1.33423684, 0.7721489 ],
[ 1.19480152, 1.04788556],
[ 0.9917477 , 0.89202008],
[-1.8356219 , -0.04839732],
[ 0.08415721, -0.71564326],
[-1.48970175, -0.19299604],
[ 0.38782418, -0.82060119],
[-0.01448044, -0.9779841 ],
[-2.0521341 , -0.02129125],
[ 0.10331194, -0.82162781],
[-0.44189315, -0.65710974],
[ 1.10390926, 1.02481182],
[-1.59227759, -0.17374038],
[-1.47344152, -0.02202853],
[-1.35514704, 0.22971067],
[ 0.0412337 , -1.23776622],
[ 0.4761517 , -1.13672124],
[ 1.04335676, 0.82345905],
[-0.07961882, -0.85677394],
[ 0.87065059, 1.08052841],
[ 1.40267313, 1.07525119],
[ 0.80111157, 1.28342825],
[-0.16527516, -1.23583804],
[-0.33779221, -0.59194323],
[ 0.80610749, -0.73752159],
[-1.43590032, -0.56384446],
[ 0.54868895, -0.95143829],
[ 0.46803131, -0.74973907],
[-1.5137129 , -0.83914323],
[ 0.9138436 , 1.51126532],
[-1.97233903, -0.41155375],
[ 0.5213406 , -0.88654894],
[ 0.62759494, -1.18590477],
[ 0.94163014, 1.35399335],
[ 0.56994768, 1.07036606],
[-1.87663382, 0.14745773],
[ 0.90612186, 0.91084011],
[-1.37481454, 0.28428395],
[-1.80564029, -0.96710574],
[ 0.34307757, -0.79999275],
[ 0.70380566, 1.00025804],
[-1.68489862, -0.30564595],
[ 1.31473221, 0.98614978],
[ 0.26151216, -0.26069251],
[ 0.9193121 , 0.82371485],
[-1.21795929, -0.20219674],
[-0.17722723, -1.02665245],
[ 0.64824862, -0.66822881],
[ 0.41206786, -0.28783784],
[ 1.01568202, 1.13481667],
[ 0.67900254, -0.91489502],
[-1.05182747, -0.01062376],
[ 0.61306599, 1.78210384],
[-1.50219748, -0.52308922],
[-1.72717293, -0.46173916],
[-1.60995631, -0.1821007 ],
[-1.09111021, -0.0781398 ],
[-0.01046978, -0.80913034],
[ 0.32782303, -0.80734754],
[ 1.22038503, 1.1959793 ],
[-1.33328681, -0.30001937],
[ 0.87959517, 1.11566491],
[-1.14829098, -0.30400762],
[-0.58019755, -1.19996018],
[-0.01161159, -0.78468854],
[ 0.17359724, -0.63398145],
[ 1.32738556, 0.67759969],
[-1.93467327, 0.30572472],
[-1.57761893, -0.27726365],
[ 0.47639 , 1.21422648],
[-1.65237509, -0.6803981 ],
[-0.12609976, -1.04327457],
[-1.89607082, -0.70085502],
[ 0.57466899, 0.74878369],
[-0.16660312, -0.83110295],
[ 0.8013355 , 1.22244435],
[ 1.18455426, 1.4346467 ],
[ 1.08864428, 0.64667112],
[-1.61158505, 0.22805725],
[-1.57512205, -0.09612576],
[ 0.0721357 , -0.69640328],
[-1.40054298, 0.16390598],
[ 1.09607713, 1.16804691],
[-2.54346204, -0.23089822],
[-1.34544875, 0.25151126],
[-1.35478629, -0.19103317],
[ 0.18368113, -1.15827725],
[-1.31368677, -0.376357 ],
[ 0.09990129, 1.22500491],
[ 1.17225574, 1.30835143],
[ 0.0865397 , -0.79714371],
[-0.21053923, -1.13421511],
[ 0.26496024, -0.94760742],
[-0.2557591 , -1.06266022],
[-0.26039757, -0.74774225],
[-1.91787359, 0.16434571],
[ 0.93021139, 0.49436331],
[ 0.44770467, -0.72877918],
[-1.63802869, -0.58925528],
[-1.95712763, -0.10125137],
[ 0.9270337 , 0.88251423],
[ 1.25660093, 0.60828073],
[-1.72818632, 0.08416887],
[ 0.3499788 , -0.30490298],
[-1.51696082, -0.50913109],
[ 0.18763605, -0.55424924],
[ 0.89609809, 0.83551508],
[-1.54968857, -0.17114782],
[ 1.2157457 , 1.23317728],
[ 0.20307745, -1.03784906],
[ 0.84589086, 1.03615273],
[ 0.53237919, 1.47362884],
[-0.05319044, -1.36150553],
[ 1.38819743, 1.11729915],
[ 1.00696304, 1.0367721 ],
[ 0.56681869, -1.09637176],
[ 0.86888296, 1.05248874],
[-1.16286609, -0.55875245],
[ 0.27717768, -0.83844015],
[ 0.16563267, -0.80306607],
[ 0.38263303, -0.42683241],
[ 1.14519807, 0.89659026],
[ 0.81455857, 0.67533667],
[-1.8603152 , -0.09537561],
[ 0.965641 , 0.90295579],
[-1.49897451, -0.33254044],
[-0.1335489 , -0.80727582],
[ 0.12541527, -1.13354906],
[ 1.06062436, 1.28816358],
[-1.49154578, -0.2024641 ],
[ 1.16189032, 1.28819877],
[ 0.54282033, 0.75203524],
[ 0.89221065, 0.99211624],
[-1.49932011, -0.32430667],
[ 0.3166647 , -1.34482915],
[ 0.13972469, -1.22097448],
[-1.5499724 , -0.10782584],
[ 1.23846858, 1.37668804],
[ 1.25558954, 0.72026098],
[ 0.25558689, -1.28529763],
[ 0.45168933, -0.55952093],
[ 1.06202057, 1.03404604],
[ 0.67451908, -0.54970299],
[ 0.22759676, -1.02729468],
[-1.45835281, -0.04951074],
[ 0.23273501, -0.70849262],
[ 1.59679589, 1.11395076],
[ 0.80476105, 0.544627 ],
[ 1.15492521, 1.04352191],
[ 0.59632776, -1.19142897],
[ 0.02839068, -0.43829366],
[ 1.13451584, 0.5632633 ],
[ 0.21576204, -1.04445753],
[ 1.41048987, 1.02830719],
[ 1.12289302, 0.58029441],
[ 0.25200688, -0.82588436],
[-1.28566081, -0.07390909],
[ 1.52849815, 1.11822469],
[-0.23907858, -0.70541972],
[-0.25792784, -0.81825035],
[ 0.59367818, -0.45239915],
[ 0.07931909, -0.29233213],
[-1.27256815, 0.11630577],
[ 0.66930129, 1.00731481],
[ 0.34791546, -1.20822877],
[-2.11283993, -0.66897935],
[-1.6293824 , -0.32718222],
[-1.53819139, -0.01501972],
[-0.11988545, -0.6036339 ],
[-1.54418956, -0.30389844],
[ 0.30026614, -0.77723173],
[ 0.00935449, -0.53888192],
[-1.33424393, -0.11560431],
[ 0.47504489, 0.78421384],
[ 0.59313264, 1.232239 ],
[ 0.41370369, -1.35205857],
[ 0.55840948, 0.78831053],
[ 0.49855018, -0.789949 ],
[ 0.35675809, -0.81038693],
[-1.86197825, -0.59071305],
[-1.61977671, -0.16076687],
[ 0.80779295, -0.73311294],
[ 1.62745775, 0.62787163],
[-1.56993593, -0.08467567],
[ 1.02558561, 0.89383302],
[ 0.24293461, -0.6088253 ],
[ 1.23130242, 1.00262186],
[-1.9651013 , -0.15886289],
[ 0.42795032, -0.70384432],
[-1.58306818, -0.19431923],
[-1.57195922, 0.01413469],
[-0.98145373, 0.06132285],
[-1.48637844, -0.5746531 ],
[ 0.98745828, 0.69188053],
[ 1.28619721, 1.28128821],
[ 0.85850596, 0.95541481],
[ 0.19028286, -0.82112942],
[ 0.26561046, -0.04255239],
[-1.61897897, 0.00862372],
[ 0.24070183, -0.52664209],
[ 1.15220993, 0.43916694],
[-1.21967812, -0.2580313 ],
[ 0.33412533, -0.86117761],
[ 0.17131003, -0.75638965],
[-1.19828397, -0.73744665],
[-0.12245932, -0.45648879],
[ 1.51200698, 0.88825741],
[ 1.10338866, 0.92347479],
[ 1.30972095, 0.59066989],
[ 0.19964876, 1.14855889],
[ 0.81460515, 0.84538972],
[-1.6422739 , -0.42296206],
[ 0.01224351, -0.21247816],
[ 0.33709102, -0.74618065],
[ 0.47301054, 0.72712075],
[ 0.34706626, 1.23033757],
[-0.00393279, -0.97209694],
[-1.64303119, 0.05276337],
[ 1.44649625, 1.14217033],
[-1.93030087, -0.40026146],
[-2.37296135, -0.72633645],
[ 0.45860122, -1.06048953],
[ 0.4896361 , -1.18928313],
[-1.02335902, -0.17520578],
[-1.32761107, -0.93963549],
[-1.50987909, -0.09473658],
[ 0.02723057, -0.79870549],
[ 1.0169412 , 1.26461701],
[ 0.47733527, -0.9898471 ],
[-1.27784224, -0.547416 ],
[ 0.49898802, -0.6237259 ],
[ 1.06004731, 0.86870008],
[ 1.00207501, 1.38293512],
[ 1.31161394, 0.62833956],
[ 1.13428443, 1.18346542],
[ 1.27671346, 0.96632878],
[-0.63342885, -0.97768251],
[ 0.12698779, -0.93142317],
[-1.34510812, -0.23754226],
[-0.53162278, -1.25153594],
[ 0.21959934, -0.90269938],
[-1.78997479, -0.12115748],
[ 1.23197473, -0.07453764],
[ 1.4163536 , 1.21551752],
[-1.90280976, -0.1638976 ],
[-0.22440081, -0.75454248],
[ 0.59559412, 0.92414553],
[ 1.21930773, 1.08175284],
[-1.99427535, -0.37587799],
[-1.27818474, -0.52454551],
[ 0.62352689, -1.01430108],
[ 0.14024251, -0.428266 ],
[-0.16145713, -1.16359731],
[-1.74795865, -0.06033101],
[-1.16659791, 0.0902393 ],
[ 0.41110408, -0.8084249 ],
[ 1.14757168, 0.77804528],
[-1.65590748, -0.40105446],
[-1.15306865, 0.00858699],
[ 0.60892121, 0.68974833],
[-0.08434138, -0.97615256],
[ 0.19170053, -0.42331438],
[ 0.29663162, -1.13357399],
[-1.36893628, -0.25052124],
[-0.08037807, -0.56784155],
[ 0.35695011, -1.15064408],
[ 0.02482179, -0.63594828],
[-1.49075558, -0.2482507 ],
[-1.408588 , 0.25635431],
[-1.98274626, -0.54584475]])
# +
xs = points[:, 0]
ys = points[:, 1]
plt.scatter(xs, ys)
# -
# ### Clustering 2D points
# From the scatter plot of the previous exercise, you saw that the points seem to separate into 3 clusters. You'll now create a KMeans model to find 3 clusters, and fit it to the data points from the previous exercise. After the model has been fit, you'll obtain the cluster labels for some new points using the ```.predict()``` method.
new_points = np.array([[ 4.00233332e-01, -1.26544471e+00],
[ 8.03230370e-01, 1.28260167e+00],
[-1.39507552e+00, 5.57292921e-02],
[-3.41192677e-01, -1.07661994e+00],
[ 1.54781747e+00, 1.40250049e+00],
[ 2.45032018e-01, -4.83442328e-01],
[ 1.20706886e+00, 8.88752605e-01],
[ 1.25132628e+00, 1.15555395e+00],
[ 1.81004415e+00, 9.65530731e-01],
[-1.66963401e+00, -3.08103509e-01],
[-7.17482105e-02, -9.37939700e-01],
[ 6.82631927e-01, 1.10258160e+00],
[ 1.09039598e+00, 1.43899529e+00],
[-1.67645414e+00, -5.04557049e-01],
[-1.84447804e+00, 4.52539544e-02],
[ 1.24234851e+00, 1.02088661e+00],
[-1.86147041e+00, 6.38645811e-03],
[-1.46044943e+00, 1.53252383e-01],
[ 4.98981817e-01, 8.98006058e-01],
[ 9.83962244e-01, 1.04369375e+00],
[-1.83136742e+00, -1.63632835e-01],
[ 1.30622617e+00, 1.07658717e+00],
[ 3.53420328e-01, -7.51320218e-01],
[ 1.13957970e+00, 1.54503860e+00],
[ 2.93995694e-01, -1.26135005e+00],
[-1.14558225e+00, -3.78709636e-02],
[ 1.18716105e+00, 6.00240663e-01],
[-2.23211946e+00, 2.30475094e-01],
[-1.28320430e+00, -3.93314568e-01],
[ 4.94296696e-01, -8.83972009e-01],
[ 6.31834930e-02, -9.11952228e-01],
[ 9.35759539e-01, 8.66820685e-01],
[ 1.58014721e+00, 1.03788392e+00],
[ 1.06304960e+00, 1.02706082e+00],
[-1.39732536e+00, -5.05162249e-01],
[-1.09935240e-01, -9.08113619e-01],
[ 1.17346758e+00, 9.47501092e-01],
[ 9.20084511e-01, 1.45767672e+00],
[ 5.82658956e-01, -9.00086832e-01],
[ 9.52772328e-01, 8.99042386e-01],
[-1.37266956e+00, -3.17878215e-02],
[ 2.12706760e-02, -7.07614194e-01],
[ 3.27049052e-01, -5.55998107e-01],
[-1.71590267e+00, 2.15222266e-01],
[ 5.12516209e-01, -7.60128245e-01],
[ 1.13023469e+00, 7.22451122e-01],
[-1.43074310e+00, -3.42787511e-01],
[-1.82724625e+00, 1.17657775e-01],
[ 1.41801350e+00, 1.11455080e+00],
[ 1.26897304e+00, 1.41925971e+00],
[ 8.04076494e-01, 1.63988557e+00],
[ 8.34567752e-01, 1.09956689e+00],
[-1.24714732e+00, -2.23522320e-01],
[-1.29422537e+00, 8.18770024e-02],
[-2.27378316e-01, -4.13331387e-01],
[ 2.18830387e-01, -4.68183120e-01],
[-1.22593414e+00, 2.55599147e-01],
[-1.31294033e+00, -4.28892070e-01],
[-1.33532382e+00, 6.52053776e-01],
[-3.01100233e-01, -1.25156451e+00],
[ 2.02778356e-01, -9.05277445e-01],
[ 1.01357784e+00, 1.12378981e+00],
[ 8.18324394e-01, 8.60841257e-01],
[ 1.26181556e+00, 1.46613744e+00],
[ 4.64867724e-01, -7.97212459e-01],
[ 3.60908898e-01, 8.44106720e-01],
[-2.15098310e+00, -3.69583937e-01],
[ 1.05005281e+00, 8.74181364e-01],
[ 1.06580074e-01, -7.49268153e-01],
[-1.73945723e+00, 2.52183577e-01],
[-1.12017687e-01, -6.52469788e-01],
[ 5.16618951e-01, -6.41267582e-01],
[ 3.26621787e-01, -8.80608015e-01],
[ 1.09017759e+00, 1.10952558e+00],
[ 3.64459576e-01, -6.94215622e-01],
[-1.90779318e+00, 1.87383674e-01],
[-1.95601829e+00, 1.39959126e-01],
[ 3.18541701e-01, -4.05271704e-01],
[ 7.36512699e-01, 1.76416255e+00],
[-1.44175162e+00, -5.72320429e-02],
[ 3.21757168e-01, -5.34283821e-01],
[-1.37317305e+00, 4.64484644e-02],
[ 6.87225910e-02, -1.10522944e+00],
[ 9.59314218e-01, 6.52316210e-01],
[-1.62641919e+00, -5.62423280e-01],
[ 1.06788305e+00, 7.29260482e-01],
[-1.79643547e+00, -9.88307418e-01],
[-9.88628377e-02, -6.81198092e-02],
[-1.05135700e-01, 1.17022143e+00],
[ 8.79964699e-01, 1.25340317e+00],
[ 9.80753407e-01, 1.15486539e+00],
[-8.33224966e-02, -9.24844368e-01],
[ 8.48759673e-01, 1.09397425e+00],
[ 1.32941649e+00, 1.13734563e+00],
[ 3.23788068e-01, -7.49732451e-01],
[-1.52610970e+00, -2.49016929e-01],
[-1.48598116e+00, -2.68828608e-01],
[-1.80479553e+00, 1.87052700e-01],
[-2.01907347e+00, -4.49511651e-01],
[ 2.87202402e-01, -6.55487415e-01],
[ 8.22295102e-01, 1.38443234e+00],
[-3.56997036e-02, -8.01825807e-01],
[-1.66955440e+00, -1.38258505e-01],
[-1.78226821e+00, 2.93353033e-01],
[ 7.25837138e-01, -6.23374024e-01],
[ 3.88432593e-01, -7.61283497e-01],
[ 1.49002783e+00, 7.95678671e-01],
[ 6.55423228e-04, -7.40580702e-01],
[-1.34533116e+00, -4.75629937e-01],
[-8.03845106e-01, -3.09943013e-01],
[-2.49041295e-01, -1.00662418e+00],
[-1.41095118e+00, -7.06744127e-02],
[-1.75119594e+00, -3.00491336e-01],
[-1.27942724e+00, 1.73774600e-01],
[ 3.35028183e-01, 6.24761151e-01],
[ 1.16819649e+00, 1.18902251e+00],
[ 7.15210457e-01, 9.26077419e-01],
[ 1.30057278e+00, 9.16349565e-01],
[-1.21697008e+00, 1.10039477e-01],
[-1.70707935e+00, -5.99659536e-02],
[ 1.20730655e+00, 1.05480463e+00],
[ 1.86896009e-01, -9.58047234e-01],
[ 8.03463471e-01, 3.86133140e-01],
[-1.73486790e+00, -1.49831913e-01],
[ 1.31261499e+00, 1.11802982e+00],
[ 4.04993148e-01, -5.10900347e-01],
[-1.93267968e+00, 2.20764694e-01],
[ 6.56004799e-01, 9.61887161e-01],
[-1.40588215e+00, 1.17134403e-01],
[-1.74306264e+00, -7.47473959e-02],
[ 5.43745412e-01, 1.47209224e+00],
[-1.97331669e+00, -2.27124493e-01],
[ 1.53901171e+00, 1.36049081e+00],
[-1.48323452e+00, -4.90302063e-01],
[ 3.86748484e-01, -1.26173400e+00],
[ 1.17015716e+00, 1.18549415e+00],
[-8.05381721e-02, -3.21923627e-01],
[-6.82273156e-02, -8.52825887e-01],
[ 7.13500028e-01, 1.27868520e+00],
[-1.85014378e+00, -5.03490558e-01],
[ 6.36085266e-02, -1.41257040e+00],
[ 1.52966062e+00, 9.66056572e-01],
[ 1.62165714e-01, -1.37374843e+00],
[-3.23474497e-01, -7.06620269e-01],
[-1.51768993e+00, 1.87658302e-01],
[ 8.88895911e-01, 7.62237161e-01],
[ 4.83164032e-01, 8.81931869e-01],
[-5.52997766e-02, -7.11305016e-01],
[-1.57966441e+00, -6.29220313e-01],
[ 5.51308645e-02, -8.47206763e-01],
[-2.06001582e+00, 5.87697787e-02],
[ 1.11810855e+00, 1.30254175e+00],
[ 4.87016164e-01, -9.90143937e-01],
[-1.65518042e+00, -1.69386383e-01],
[-1.44349738e+00, 1.90299243e-01],
[-1.70074547e-01, -8.26736022e-01],
[-1.82433979e+00, -3.07814626e-01],
[ 1.03093485e+00, 1.26457691e+00],
[ 1.64431169e+00, 1.27773115e+00],
[-1.47617693e+00, 2.60783872e-02],
[ 1.00953067e+00, 1.14270181e+00],
[-1.45285636e+00, -2.55216207e-01],
[-1.74092917e+00, -8.34443177e-02],
[ 1.22038299e+00, 1.28699961e+00],
[ 9.16925397e-01, 7.32070275e-01],
[-1.60754185e-03, -7.26375571e-01],
[ 8.93841238e-01, 8.41146643e-01],
[ 6.33791961e-01, 1.00915134e+00],
[-1.47927075e+00, -6.99781936e-01],
[ 5.44799374e-02, -1.06441970e+00],
[-1.51935568e+00, -4.89276929e-01],
[ 2.89939026e-01, -7.73145523e-01],
[-9.68154061e-03, -1.13302207e+00],
[ 1.13474639e+00, 9.71541744e-01],
[ 5.36421406e-01, -8.47906388e-01],
[ 1.14759864e+00, 6.89915205e-01],
[ 5.73291902e-01, 7.90802710e-01],
[ 2.12377397e-01, -6.07569808e-01],
[ 5.26579548e-01, -8.15930264e-01],
[-2.01831641e+00, 6.78650740e-02],
[-2.35512624e-01, -1.08205132e+00],
[ 1.59274780e-01, -6.00717261e-01],
[ 2.28120356e-01, -1.16003549e+00],
[-1.53658378e+00, 8.40798808e-02],
[ 1.13954609e+00, 6.31782001e-01],
[ 1.01119255e+00, 1.04360805e+00],
[-1.42039867e-01, -4.81230337e-01],
[-2.23120182e+00, 8.49162905e-02],
[ 1.25554811e-01, -1.01794793e+00],
[-1.72493509e+00, -6.94426177e-01],
[-1.60434630e+00, 4.45550868e-01],
[ 7.37153979e-01, 9.26560744e-01],
[ 6.72905271e-01, 1.13366030e+00],
[ 1.20066456e+00, 7.26273093e-01],
[ 7.58747209e-02, -9.83378326e-01],
[ 1.28783262e+00, 1.18088601e+00],
[ 1.06521930e+00, 1.00714746e+00],
[ 1.05871698e+00, 1.12956519e+00],
[-1.12643410e+00, 1.66787744e-01],
[-1.10157218e+00, -3.64137806e-01],
[ 2.35118217e-01, -1.39769949e-01],
[ 1.13853795e+00, 1.01018519e+00],
[ 5.31205654e-01, -8.81990792e-01],
[ 4.33085936e-01, -7.64059042e-01],
[-4.48926156e-03, -1.30548411e+00],
[-1.76348589e+00, -4.97430739e-01],
[ 1.36485681e+00, 5.83404699e-01],
[ 5.66923900e-01, 1.51391963e+00],
[ 1.35736826e+00, 6.70915318e-01],
[ 1.07173397e+00, 6.11990884e-01],
[ 1.00106915e+00, 8.93815326e-01],
[ 1.33091007e+00, 8.79773879e-01],
[-1.79603740e+00, -3.53883973e-02],
[-1.27222979e+00, 4.00156642e-01],
[ 8.47480603e-01, 1.17032364e+00],
[-1.50989129e+00, -7.12318330e-01],
[-1.24953576e+00, -5.57859730e-01],
[-1.27717973e+00, -5.99350550e-01],
[-1.81946743e+00, 7.37057673e-01],
[ 1.19949867e+00, 1.56969386e+00],
[-1.25543847e+00, -2.33892826e-01],
[-1.63052058e+00, 1.61455865e-01],
[ 1.10611305e+00, 7.39698224e-01],
[ 6.70193192e-01, 8.70567001e-01],
[ 3.69670156e-01, -6.94645306e-01],
[-1.26362293e+00, -6.99249285e-01],
[-3.66687507e-01, -1.35310260e+00],
[ 2.44032147e-01, -6.59470793e-01],
[-1.27679142e+00, -4.85453412e-01],
[ 3.77473612e-02, -6.99251605e-01],
[-2.19148539e+00, -4.91199500e-01],
[-2.93277777e-01, -5.89488212e-01],
[-1.65737397e+00, -2.98337786e-01],
[ 7.36638861e-01, 5.78037057e-01],
[ 1.13709081e+00, 1.30119754e+00],
[-1.44146601e+00, 3.13934680e-02],
[ 5.92360708e-01, 1.22545114e+00],
[ 6.51719414e-01, 4.92674894e-01],
[ 5.94559139e-01, 8.25637315e-01],
[-1.87900722e+00, -5.21899626e-01],
[ 2.15225041e-01, -1.28269851e+00],
[ 4.99145965e-01, -6.70268634e-01],
[-1.82954176e+00, -3.39269731e-01],
[ 7.92721403e-01, 1.33785606e+00],
[ 9.54363372e-01, 9.80396626e-01],
[-1.35359846e+00, 1.03976340e-01],
[ 1.05595062e+00, 8.07031927e-01],
[-1.94311010e+00, -1.18976964e-01],
[-1.39604137e+00, -3.10095976e-01],
[ 1.28977624e+00, 1.01753365e+00],
[-1.59503139e+00, -5.40574609e-01],
[-1.41994046e+00, -3.81032569e-01],
[-2.35569801e-02, -1.10133702e+00],
[-1.26038568e+00, -6.93273886e-01],
[ 9.60215981e-01, -8.11553694e-01],
[ 5.51803308e-01, -1.01793176e+00],
[ 3.70185085e-01, -1.06885468e+00],
[ 8.25529207e-01, 8.77007060e-01],
[-1.87032595e+00, 2.87507199e-01],
[-1.56260769e+00, -1.89196712e-01],
[-1.26346548e+00, -7.74725237e-01],
[-6.33800421e-02, -7.59400611e-01],
[ 8.85298280e-01, 8.85620519e-01],
[-1.43324686e-01, -1.16083678e+00],
[-1.83908725e+00, -3.26655515e-01],
[ 2.74709229e-01, -1.04546829e+00],
[-1.45703573e+00, -2.91842036e-01],
[-1.59048842e+00, 1.66063031e-01],
[ 9.25549284e-01, 7.41406406e-01],
[ 1.97245469e-01, -7.80703225e-01],
[ 2.88401697e-01, -8.32425551e-01],
[ 7.24141618e-01, -7.99149200e-01],
[-1.62658639e+00, -1.80005543e-01],
[ 5.84481588e-01, 1.13195640e+00],
[ 1.02146732e+00, 4.59657799e-01],
[ 8.65050554e-01, 9.57714887e-01],
[ 3.98717766e-01, -1.24273147e+00],
[ 8.62234892e-01, 1.10955561e+00],
[-1.35999430e+00, 2.49942654e-02],
[-1.19178505e+00, -3.82946323e-02],
[ 1.29392424e+00, 1.10320509e+00],
[ 1.25679630e+00, -7.79857582e-01],
[ 9.38040302e-02, -5.53247258e-01],
[-1.73512175e+00, -9.76271667e-02],
[ 2.23153587e-01, -9.43474351e-01],
[ 4.01989100e-01, -1.10963051e+00],
[-1.42244158e+00, 1.81914703e-01],
[ 3.92476267e-01, -8.78426277e-01],
[ 1.25181875e+00, 6.93614996e-01],
[ 1.77481317e-02, -7.20304235e-01],
[-1.87752521e+00, -2.63870424e-01],
[-1.58063602e+00, -5.50456344e-01],
[-1.59589493e+00, -1.53932892e-01],
[-1.01829770e+00, 3.88542370e-02],
[ 1.24819659e+00, 6.60041803e-01],
[-1.25551377e+00, -2.96172009e-02],
[-1.41864559e+00, -3.58230179e-01],
[ 5.25758326e-01, 8.70500543e-01],
[ 5.55599988e-01, 1.18765072e+00],
[ 2.81344439e-02, -6.99111314e-01]])
# +
from sklearn.cluster import KMeans
# Create a KMeans instance with 3 clusters: model
model = KMeans(n_clusters=3)
# Fit model to points
model.fit(points)
# Determine the cluster labels of new_points: labels
labels = model.predict(new_points)
# Print cluster labels of new_points
print(labels)
# -
# ### Inspect your clustering
# Let's now inspect the clustering you performed in the previous exercise!
#
#
# +
# Assign the columns of new_points: xs and ys
xs = new_points[:, 0]
ys = new_points[:, 1]
# Make a scatter plot of xs and ys, using labels to define the colors
plt.scatter(xs, ys, c=labels, alpha=0.5)
# Assign the cluster centers: centroids
centroids = model.cluster_centers_
# Assign the columns of centroids: centroids_x, centroids_y
centroids_x = centroids[:, 0]
centroids_y = centroids[:, 1]
# Make a scatter plot of centroids_x and centroids_y
plt.scatter(centroids_x, centroids_y, marker='D', s=50, color='red')
plt.savefig('../images/kmeans-centroid.png')
# -
# ## Evaluating a clustering
# - Evaluating a clustering
# - Can check correspondence with e.g. iris species, but what if there are no species to check against?
# - Measure quality of a clustering
# - Informs choice of how many clusters to look for
# - Cross-tabulation with pandas
# - Clusters vs species is a "cross-tabulation"
# - Measuring clustering quality
# - Using only samples and their cluster labels
# - A good clustering has tight clusters
# - Samples in each cluster bunched together
# - Inertia measures clustering quality
# - Measures how spread out the clusters are (lower is better)
# - Distance from each sample to centroid of its cluster
# - k-means attempts to minimize the inertia when choosing clusters
# - How many clusters to choose?
# - Choose an "elbow" in the inertia plot
# - Where inertia begins to decrease more slowly
# ### How many clusters of grain?
# In the video, you learned how to choose a good number of clusters for a dataset using the k-means inertia graph. You are given an array ```samples``` containing the measurements (such as area, perimeter, length, and several others) of samples of grain. What's a good number of clusters in this case?
# #### Preprocess
df = pd.read_csv('./dataset/seeds.csv', header=None)
df[7] = df[7].map({1:'Kama wheat', 2:'Rosa wheat', 3:'Canadian wheat'})
df.head()
samples = df.iloc[:, :-1].values
varieties = df.iloc[:, -1].values
# +
ks = range(1,6)
inertias = []
for k in ks:
# Create a KMeans instance with k clusters: model
model = KMeans(n_clusters=k)
# Fit model to samples
model.fit(samples)
# Append the inertia to the list of inertias
inertias.append(model.inertia_)
# Plot ks vs inertias
plt.plot(ks, inertias, '-o')
plt.xlabel('number of clusters, k')
plt.ylabel('inertia')
plt.xticks(ks)
# -
# ### Evaluating the grain clustering
# In the previous exercise, you observed from the inertia plot that 3 is a good number of clusters for the grain data. In fact, the grain samples come from a mix of 3 different grain varieties: "Kama", "Rosa" and "Canadian". In this exercise, cluster the grain samples into three clusters, and compare the clusters to the grain varieties using a cross-tabulation.
#
# +
# Create a KMeans model with 3 clusters: model
model = KMeans(n_clusters=3)
# Use fit_predict to fit model and obtain cluster labels: labels
labels = model.fit_predict(samples)
# Create a DataFrame with labels and varieties as columns: df
df = pd.DataFrame({'labels': labels, 'varieties': varieties})
# Create crosstab: ct
ct = pd.crosstab(df['labels'], df['varieties'])
# Display ct
print(ct)
# -
# ## Transforming features for better clusterings
# - StandardScaler
# - In kmeans, feature variance = feature influence
# - ```StandardScaler``` transforms each feature to have mean 0 and variance 1
# - Features are said to be "standardized"
# - StandardScaler, then KMeans
# - Need to perform two steps: ```StandardScaler```, then ```KMeans```
# - Use ```sklearn``` pipeline to combine multiple steps
# - Data flows from one step into the next
# ### Scaling fish data for clustering
# You are given an array ```samples``` giving measurements of fish. Each row represents an individual fish. The measurements, such as weight in grams, length in centimeters, and the percentage ratio of height to length, have very different scales. In order to cluster this data effectively, you'll need to standardize these features first. In this exercise, you'll build a pipeline to standardize and cluster the data.
#
# These fish measurement data were sourced from the [Journal of Statistics Education](http://ww2.amstat.org/publications/jse/jse_data_archive.htm).
# #### Preprocess
df = pd.read_csv('./dataset/fish.csv', header=None)
df.head()
samples = df.iloc[:, 1:].values
species = df.iloc[:, 0].values
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import KMeans
# Create scaler: scaler
scaler = StandardScaler()
# Create KMeans instance: kmeans
kmeans = KMeans(n_clusters=4)
# Create pipeline: pipeline
pipeline = make_pipeline(scaler, kmeans)
# -
# ### Clustering the fish data
# You'll now use your standardization and clustering pipeline from the previous exercise to cluster the fish by their measurements, and then create a cross-tabulation to compare the cluster labels with the fish species.
# +
# Fit the pipeline to samples
pipeline.fit(samples)
# Calculate the cluster labels: labels
labels = pipeline.predict(samples)
# Create a DataFrame with labels and species as columns: df
df = pd.DataFrame({'labels': labels, 'species': species})
# Create crosstab: ct
ct = pd.crosstab(df['labels'], df['species'])
# Display ct
print(ct)
# -
# ### Clustering stocks using KMeans
# In this exercise, you'll cluster companies using their daily stock price movements (i.e. the dollar difference between the closing and opening prices for each trading day). You are given a NumPy array ```movements``` of daily price movements from 2010 to 2015 (obtained from Yahoo! Finance), where each row corresponds to a company, and each column corresponds to a trading day.
#
# Some stocks are more expensive than others. To account for this, include a ```Normalizer``` at the beginning of your pipeline. The Normalizer will separately transform each company's stock price to a relative scale before the clustering begins.
#
# Note that ```Normalizer()``` is different to ```StandardScaler()```, which you used in the previous exercise. While ```StandardScaler()``` standardizes features (such as the features of the fish data from the previous exercise) by removing the mean and scaling to unit variance, ```Normalizer()``` rescales each sample - here, each company's stock price - independently of the other.
# #### Preprocess
df = pd.read_csv('./dataset/company-stock-movements-2010-2015-incl.csv', index_col=0)
df.head()
movements = df.values
companies = df.index.values
# +
from sklearn.preprocessing import Normalizer
# Create a normalizer: normalizer
normalizer = Normalizer()
# Create a KMeans model with 10 clusters: kmeans
kmeans = KMeans(n_clusters=10)
# Make a pipeline chaining normalizer and kmeans: pipeline
pipeline = make_pipeline(normalizer, kmeans)
# Fit pipeline to the daily price movements
pipeline.fit(movements)
# -
# ### Which stocks move together?
# In the previous exercise, you clustered companies by their daily stock price movements. So which company have stock prices that tend to change in the same way? You'll now inspect the cluster labels from your clustering to find out.
# +
# Predict the cluster labels: labels
labels = pipeline.predict(movements)
# Create a DataFrame aligning labels and companies: df
df = pd.DataFrame({'labels': labels, 'companies': companies})
# Display df sorted by cluster label
print(df.sort_values('labels'))
| _notebooks/2020-06-01-01-Clustering-for-dataset-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit (windows store)
# language: python
# name: python3
# ---
# +
# import pandas as pd
# import os
# from dotenv import load_dotenv
# load_dotenv()
# PWD = os.getenv('PWD')
# db_name = PWD+'\\database'+'\\RVNUSDT.db'
# df = pd.read_sql(
# '''SELECT * FROM m1
# ''',
# 'sqlite:///' + db_name,
# index_col='id',
# )
# print(len(df))
# df.head(3)
# df.to_csv('../../filename.csv')
# +
import pandas as pd
import numpy as np
import pickle
import os
from dotenv import load_dotenv
load_dotenv()
PWD = os.getenv('PWD')
db_name = PWD+'\\database'+'\\RVNUSDT.db'
import sys
sys.path.insert(1, PWD+'\\modules')
from alg_modules.alg_handler import AlgHandler
from plot_modules.candle_plot import CandlePlot
from collections import deque
from paper_trade import PaperTrader
import logging
DEBUG = __debug__
LOG_FILE_NAME = 'log_file_name.log'
format = '%(asctime)s [%(levelname)s]: %(message)s'
logger = logging.basicConfig(
filename=LOG_FILE_NAME if not DEBUG else None,
format=format,
encoding='utf-8',
level=logging.INFO,
)
if not DEBUG:
logging.getLogger(logger).addHandler(logging.StreamHandler())
from stop_loss import StopLoss
from trade_strategy import TradeStrategy
# +
import numpy as np
CALCULATE_NEW = True
# SINGLE_LOOP = 1
STOP_LOSS_ENABLED = True
STOP_LOSS_THRESHOLD = -1.3
MA_list = (1, 25, 100)
stop_loss_data = {}
df = pd.read_sql(
'''
SELECT * FROM m5
WHERE date_created > "2021-12-00 00:00:00.000000"
''',
'sqlite:///' + db_name,
index_col='id',
)
if CALCULATE_NEW:
stop_loss_count, trade_data, p_trdr = TradeStrategy.static_trade_strategy(
STOP_LOSS_ENABLED=STOP_LOSS_ENABLED,
STOP_LOSS_THRESHOLD=STOP_LOSS_THRESHOLD,
df=df,
open_col='open_',
close_col='close_',
low_col='low_',
high_col='high_',
date_col='date_created',
MA_list=MA_list,
)
else:
# read data from file
with open(f'{PWD}\\temp\\p_trdr_file', 'rb') as f:
p_trdr = pickle.load(f)
trade_data = pd.read_csv(f'{PWD}\\temp\\trade_data_file')
if p_trdr.buy_price is not None:
p_trdr.convert_to_main(
price=p_trdr.buy_price
)
logging.info(f'SL_TR: {STOP_LOSS_THRESHOLD if STOP_LOSS_ENABLED else ""} SL_count {stop_loss_count}, {p_trdr.main_currency_amount} {p_trdr.main_currency_label} {p_trdr.secondary_currency_amount} {p_trdr.secondary_currency_label}')
stop_loss_data[STOP_LOSS_THRESHOLD] = p_trdr.main_currency_amount
# -
# - 2022-02-18 06:30:48,831 [INFO]: SL_TR: -1.3 SL_count 21, 120.19499068447104 RVN 0.0 USD
trade_data.head(30)
0.1>0
assert 0
with open('../../trade_data_raw.csv', 'w') as f:
f.write(trade_data.to_csv())
# print data
stop_loss_data_str = {}
for key, val in stop_loss_data.items():
stop_loss_data_str[str(key)] = str(val)
print(key, '|', val)
# plot the data
import plotly.express as px
fig__ = px.line(
x=stop_loss_data.keys(),
y=stop_loss_data.values(),
)
fig__.update_layout(
xaxis = dict(
tickmode = 'linear',
tick0 = 0,
dtick = 0.25
)
)
fig__
# save data to the dictionary
import yaml
MA_log = (25, 100)
name_proto = '_'.join(str(MA_log).split(', '))
start_date = '2021-12-00'
end_date = '_'
with open(PWD+f'\\temp\\MA_{name_proto}_SL_{start_date}_{end_date}.yaml', 'w') as f:
f.write(yaml.dump( stop_loss_data_str, sort_keys=False))
f'MA_{name_proto}_SL_{start_date}_{end_date}'
# +
# провести исследование по величине стоп лосса
# best -1.1 -10:10 step = 0.1
pd.set_option('display.max_rows', 30)
trade_data
# -
# save calculated data
if CALCULATE_NEW:
with open(PWD+'\\temp\\'+'p_trdr_file', 'wb') as f:
pickle.dump(p_trdr, f)
trade_data.to_csv(PWD+'\\temp\\'+'trade_data_file')
# +
# calc amplitiude and append it to df
# >>> max_ = 0.07392
# >>> min_ = 0.07158
# >>> (max_ - min_)/min_*100
# 3.2690695725062793
# >>>
# implement MACD
# implement EMA
## use this https://technical-analysis-library-in-python.readthedocs.io/en/latest/index.html
# implement saving currency and NOT trade w/ full amount
# -
trade_data.head(30)
# +
# calc profit
# trade_data['proift_abs'] = ''
# trade_data['proift_rel'] = ''
for i, row in trade_data.iterrows():
# try:
if row['type'] == 'sell':
try:
trade_data.loc[i, 'profit_abs'] = float(row['USD']) - float(trade_data['USD'].loc[i-2])
# (curr_buy_pr - prev_buy_pr)*100/prev_buy_pr
trade_data.loc[i, 'profit_rel'] = round((float(row['USD']) - float(trade_data['USD'].loc[i-2])) *100/float(trade_data['USD'].loc[i-2]),2)
except KeyError: print(i)
if row['type'] == 'buy':
try:
trade_data.loc[i, 'profit_abs'] = float(row['RVN']) - float(trade_data['RVN'].loc[i-2])
# (curr_buy_pr - prev_buy_pr)*100/prev_buy_pr
trade_data.loc[i, 'profit_rel'] = round((float(row['RVN']) - float(trade_data['RVN'].loc[i-2]))*100/float(trade_data['RVN'].loc[i-2]),2)
except KeyError: print(i)
# except KeyError as e: print(e)
# -
trade_data
df = pd.read_sql(
'''
SELECT * FROM m5
WHERE date_created > "2021-07-15 00:00:00.000000"
AND date_created < "2021-07-17 21:55:00.000000"
''',
'sqlite:///' + db_name,
index_col='id',
)
df.head(1)
# +
pd.set_option('display.max_rows', 470)
trade_data.head(3)
# -
for i, row in df.iterrows():
df.loc[i, 'amplitude'] = (float(row['high_']) - float(row['low_']))/float(row['low_'])*100
with open('../../df_raw.csv', 'w') as f:
f.write(df.to_csv())
df.head() # open_ < close_ -> open{GREEN}^
# +
import plotly.graph_objects as go
# region
plot = CandlePlot(
df=df,
open_col='open_',
close_col='close_',
low_col='low_',
high_col='high_',
date_col='date_created',
)
settings = {
'candle_plot': 1,
'MA_lines': 1,
'add_trades': 1,
'add_profit': 1,
'profit_annotations': 0,
'amplitude': 0,
'MACD_lines': True, # to do
'EMA_lines': True, # to do
}
fig = plot.use_settings(
settings=settings,
GO_WIDTH=1300,
GO_HEIGHT=750,
title='MA algoritm',
pair='RVNUSDT',
interval='--',
limit='no limit',
MA_list=(2, 25, 100, 200),
trade_data=trade_data,
)
# endregion
# draw MACD_lines
fig
# -
fig
trade_flags = trade_data.itertuples(name='Row', index=False)
for i in trade_flags:
print(i._6)
# p_trdr.get_info()
p_trdr.get_df()
# +
# p_trdr.convert_to_main(price=0.09488)
# df
| modules/logic_modules/ALG_TEST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1
# Add the specified code for each code cell, running the cells _in order_.
# Create a variable `food` that stores your favorite kind of food. Print or output the variable.
food = "pizza"
print(food)
# Create a variable `restaurant` that stores your favorite place to eat that kind of food.
restaurant = "Pagliacci's"
# Print the message `"I'm going to RESTAURANT for some FOOD"`, replacing the restaurant and food with your variables.
print("I'm going to " + restaurant + " for some " + food)
# Create a variable `num_friends` equal to the number of friends you would like to eat with.
num_friends = 3
# Print a message `"I'm going with X friends"`, replacing the X with the number of friends.
print("I'm going with " + str(num_friends) + " friends")
# Create a variable `meal_price`, which is how expensive you think one meal at the restaurant would be. This price should be a `float`.
meal_price = 25.00
# Update (re-assign) the `meal_price` variable so it includes a 15% tip—that is, so the price is 15% higher. Output the variable.
meal_price = round(meal_price * 1.15, 2)
print(meal_price)
# Create a variable `total_cost` that has the total estimated cost of the bill for you and all of your friends. Output or print the variable
total_cost = meal_price * (num_friends+1)
print(total_cost)
# Create a variable `budget` representing your spending budget for a night out.
budget = 200.00
# Create a variable `max_friends`, which is the maximum number of friends you can invite, at the estimated meal price, while staying within your budget. Output or print this value.
# - Be carefully that you only invite whole people!
max_friends = int((budget//meal_price)-1) #double slash rounds down to nearest whole
print(max_friends)
# Bonus: Create a variable `chorus` that is the string `"FOOD time!"` _repeated_ once for each of the friends you are able to bring. _Hint_ use the **`*`** operator. Print out the variable.
chorus = (food +" time!\n") * max_friends
print(chorus)
# Modify the above cell so that each `"FOOD time!"` is on a separate line (_hint_: use a newline character!), then rerun the cell.
| exercise-1/exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (GEE)
# language: python
# name: google_earth_engine
# ---
# %load_ext autoreload
# %autoreload 2
import ee
# +
# Trigger the authentication flow.
ee.Authenticate()
# +
# Initialize the library.
ee.Initialize()
# +
# Import the MODIS land cover collection.
lc = ee.ImageCollection('MODIS/006/MCD12Q1')
# Import the MODIS land surface temperature collection.
lst = ee.ImageCollection('MODIS/006/MOD11A1')
# Import the USGS ground elevation image.
elv = ee.Image('USGS/SRTMGL1_003')
# +
# Initial date of interest (inclusive).
i_date = '2017-01-01'
# Final date of interest (exclusive).
f_date = '2020-01-01'
# Selection of appropriate bands and dates for LST.
lst = lst.select('LST_Day_1km', 'QC_Day').filterDate(i_date, f_date)
# +
# Define the urban location of interest as a point near Lyon, France.
u_lon = 4.8148
u_lat = 45.7758
u_poi = ee.Geometry.Point(u_lon, u_lat)
# Define the rural location of interest as a point away from the city.
r_lon = 5.175964
r_lat = 45.574064
r_poi = ee.Geometry.Point(r_lon, r_lat)
# +
scale = 1000 # scale in meters
# Print the elevation near Lyon, France.
elv_urban_point = elv.sample(u_poi, scale).first().get('elevation').getInfo()
print('Ground elevation at urban point:', elv_urban_point, 'm')
# Calculate and print the mean value of the LST collection at the point.
lst_urban_point = lst.mean().sample(u_poi, scale).first().get('LST_Day_1km').getInfo()
print('Average daytime LST at urban point:', round(lst_urban_point*0.02 -273.15, 2), '°C')
# Print the land cover type at the point.
lc_urban_point = lc.first().sample(u_poi, scale).first().get('LC_Type1').getInfo()
print('Land cover value at urban point is:', lc_urban_point)
# +
# Get the data for the pixel intersecting the point in urban area.
lst_u_poi = lst.getRegion(u_poi, scale).getInfo()
# Get the data for the pixel intersecting the point in rural area.
lst_r_poi = lst.getRegion(r_poi, scale).getInfo()
# Preview the result.
lst_u_poi[:5]
# +
import pandas as pd
def ee_array_to_df(arr, list_of_bands):
"""Transforms client-side ee.Image.getRegion array to pandas.DataFrame."""
df = pd.DataFrame(arr)
# Rearrange the header.
headers = df.iloc[0]
df = pd.DataFrame(df.values[1:], columns=headers)
# Remove rows without data inside.
df = df[['longitude', 'latitude', 'time', *list_of_bands]].dropna()
# Convert the data to numeric values.
for band in list_of_bands:
df[band] = pd.to_numeric(df[band], errors='coerce')
# Convert the time field into a datetime.
df['datetime'] = pd.to_datetime(df['time'], unit='ms')
# Keep the columns of interest.
df = df[['time','datetime', *list_of_bands]]
return df
# +
lst_df_urban = ee_array_to_df(lst_u_poi,['LST_Day_1km'])
def t_modis_to_celsius(t_modis):
"""Converts MODIS LST units to degrees Celsius."""
t_celsius = 0.02*t_modis - 273.15
return t_celsius
# Apply the function to get temperature in celsius.
lst_df_urban['LST_Day_1km'] = lst_df_urban['LST_Day_1km'].apply(t_modis_to_celsius)
# Do the same for the rural point.
lst_df_rural = ee_array_to_df(lst_r_poi,['LST_Day_1km'])
lst_df_rural['LST_Day_1km'] = lst_df_rural['LST_Day_1km'].apply(t_modis_to_celsius)
lst_df_urban.head()
# +
import matplotlib.pyplot as plt
import numpy as np
from scipy import optimize
# %matplotlib inline
# Fitting curves.
## First, extract x values (times) from the dfs.
x_data_u = np.asanyarray(lst_df_urban['time'].apply(float)) # urban
x_data_r = np.asanyarray(lst_df_rural['time'].apply(float)) # rural
## Secondly, extract y values (LST) from the dfs.
y_data_u = np.asanyarray(lst_df_urban['LST_Day_1km'].apply(float)) # urban
y_data_r = np.asanyarray(lst_df_rural['LST_Day_1km'].apply(float)) # rural
## Then, define the fitting function with parameters.
def fit_func(t, lst0, delta_lst, tau, phi):
return lst0 + (delta_lst/2)*np.sin(2*np.pi*t/tau + phi)
## Optimize the parameters using a good start p0.
lst0 = 20
delta_lst = 40
tau = 365*24*3600*1000 # milliseconds in a year
phi = 2*np.pi*4*30.5*3600*1000/tau # offset regarding when we expect LST(t)=LST0
params_u, params_covariance_u = optimize.curve_fit(
fit_func, x_data_u, y_data_u, p0=[lst0, delta_lst, tau, phi])
params_r, params_covariance_r = optimize.curve_fit(
fit_func, x_data_r, y_data_r, p0=[lst0, delta_lst, tau, phi])
# Subplots.
fig, ax = plt.subplots(figsize=(14, 6))
# Add scatter plots.
ax.scatter(lst_df_urban['datetime'], lst_df_urban['LST_Day_1km'],
c='black', alpha=0.2, label='Urban (data)')
ax.scatter(lst_df_rural['datetime'], lst_df_rural['LST_Day_1km'],
c='green', alpha=0.35, label='Rural (data)')
# Add fitting curves.
ax.plot(lst_df_urban['datetime'],
fit_func(x_data_u, params_u[0], params_u[1], params_u[2], params_u[3]),
label='Urban (fitted)', color='black', lw=2.5)
ax.plot(lst_df_rural['datetime'],
fit_func(x_data_r, params_r[0], params_r[1], params_r[2], params_r[3]),
label='Rural (fitted)', color='green', lw=2.5)
# Add some parameters.
ax.set_title('Daytime Land Surface Temperature Near Lyon', fontsize=16)
ax.set_xlabel('Date', fontsize=14)
ax.set_ylabel('Temperature [C]', fontsize=14)
ax.set_ylim(-0, 40)
ax.grid(lw=0.2)
ax.legend(fontsize=14, loc='lower right')
plt.show()
# +
import folium
# Define the center of our map.
lat, lon = 45.77, 4.855
my_map = folium.Map(location=[lat, lon], zoom_start=10)
my_map
# +
def add_ee_layer(self, ee_image_object, vis_params, name):
"""Adds a method for displaying Earth Engine image tiles to folium map."""
map_id_dict = ee.Image(ee_image_object).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles=map_id_dict['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
name=name,
overlay=True,
control=True
).add_to(self)
# Add Earth Engine drawing method to folium.
folium.Map.add_ee_layer = add_ee_layer
# +
# Select a specific band and dates for land cover.
lc_img = lc.select('LC_Type1').filterDate(i_date).first()
# Set visualization parameters for land cover.
lc_vis_params = {
'min': 1,'max': 17,
'palette': ['05450a','086a10', '54a708', '78d203', '009900', 'c6b044',
'dcd159', 'dade48', 'fbff13', 'b6ff05', '27ff87', 'c24f44',
'a5a5a5', 'ff6d4c', '69fff8', 'f9ffa4', '1c0dff']
}
# Create a map.
lat, lon = 45.77, 4.855
my_map = folium.Map(location=[lat, lon], zoom_start=7)
# Add the land cover to the map object.
my_map.add_ee_layer(lc_img, lc_vis_params, 'Land Cover')
# Add a layer control panel to the map.
my_map.add_child(folium.LayerControl())
# Display the map.
display(my_map)
# -
# Define a region of interest with a buffer zone of 1000 km around Lyon.
roi = u_poi.buffer(1e6)
# +
# Reduce the LST collection by mean.
lst_img = lst.mean()
# Adjust for scale factor.
lst_img = lst_img.select('LST_Day_1km').multiply(0.02)
# Convert Kelvin to Celsius.
lst_img = lst_img.select('LST_Day_1km').add(-273.15)
# +
from IPython.display import Image
# Create a URL to the styled image for a region around France.
url = lst_img.getThumbUrl({
'min': 10, 'max': 30, 'dimensions': 512, 'region': roi,
'palette': ['blue', 'yellow', 'orange', 'red']})
print(url)
# Display the thumbnail land surface temperature in France.
print('\nPlease wait while the thumbnail loads, it may take a moment...')
Image(url=url)
# +
# Make pixels with elevation below sea level transparent.
elv_img = elv.updateMask(elv.gt(0))
# Display the thumbnail of styled elevation in France.
Image(url=elv_img.getThumbURL({
'min': 0, 'max': 2000, 'dimensions': 512, 'region': roi,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}))
# +
# Set visualization parameters for ground elevation.
elv_vis_params = {
'min': 0, 'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Set visualization parameters for land surface temperature.
lst_vis_params = {
'min': 0, 'max': 40,
'palette': ['white', 'blue', 'green', 'yellow', 'orange', 'red']}
# Arrange layers inside a list (elevation, LST and land cover).
ee_tiles = [elv_img, lst_img, lc_img]
# Arrange visualization parameters inside a list.
ee_vis_params = [elv_vis_params, lst_vis_params, lc_vis_params]
# Arrange layer names inside a list.
ee_tiles_names = ['Elevation', 'Land Surface Temperature', 'Land Cover']
# Create a new map.
lat, lon = 45.77, 4.855
my_map = folium.Map(location=[lat, lon], zoom_start=5)
# Add layers to the map using a loop.
for tile, vis_param, name in zip(ee_tiles, ee_vis_params, ee_tiles_names):
my_map.add_ee_layer(tile, vis_param, name)
folium.LayerControl(collapsed = False).add_to(my_map)
my_map
# -
| remote_sensing_python_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Beautiful Python
# + slideshow={"slide_type": "slide"}
import this
# + [markdown] slideshow={"slide_type": "slide"}
# # Strings
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Cadenas muy largas
# + slideshow={"slide_type": "-"}
my_very_big_string = 'For a long time I used to go to bed early. ' + \
'Sometimes, when I had put out my candle, my eyes would close ' + \
'so quickly that I had not even time to say Im going to sleep.'
# + slideshow={"slide_type": "fragment"}
my_very_big_string = '''For a long time I used to go to bed early.
Sometimes, when I had put out my candle, my eyes would close
so quickly that I had not even time to say Im going to sleep.'''
print(my_very_big_string)
# + slideshow={"slide_type": "fragment"}
# pythonico
my_very_big_string = ('For a long time I used to go to bed early. '
'Sometimes, when I had put out my candle, my eyes would close '
'so quickly that I had not even time to say Im going to sleep.')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Comprobar si una cadena contiene otra
# + slideshow={"slide_type": "-"}
my_string = 'Beautiful is better than ugly'
# + slideshow={"slide_type": "-"}
if my_string.find('ugly') != -1:
print('encontrado!')
# + slideshow={"slide_type": "fragment"}
# pythonico
if 'ugly' in my_string:
print('encontrado!')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Formato de strings
# + slideshow={"slide_type": "-"}
name = 'Alicia'
cats = 2
print('My name is ' + name + ' and I have ' + str(cats) + ' cats')
# + slideshow={"slide_type": "fragment"}
# pythonico
print('My name is %s and I have %s cats' %(name, cats))
print(f'My name is {name} and I have {cats} cats')
# + [markdown] slideshow={"slide_type": "slide"}
# # Booleanos
# + slideshow={"slide_type": "subslide"}
start = 1
end = 5
x = 3
# + slideshow={"slide_type": "-"}
if x >= start and x <= end:
print('todo ok')
# + slideshow={"slide_type": "fragment"}
# pythonico
if start <= x <= end:
print('todo ok')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Operador ternario
# + slideshow={"slide_type": "-"}
lang = 'English'
# + slideshow={"slide_type": "-"}
if lang == 'Spanish':
welcome = 'Bienvenido'
else:
welcome = 'Welcome'
# + slideshow={"slide_type": "fragment"}
# pythonico
welcome = 'Bienvenido' if lang == 'Spanish' else 'Welcome'
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Truth Value Testing
# + slideshow={"slide_type": "-"}
items = [1, 2]
if len(items) != 0:
print('Tiene algo')
# + slideshow={"slide_type": "fragment"}
# pythonico
if items:
print('Tiene algo')
# + [markdown] slideshow={"slide_type": "subslide"}
# Los siguientes valores serán evaluados como False en una comparación:
# - `None`
# - `False`
# - Cero en cualquier tipo numérico, por ejemplo: `0`, `0L`, `0.0`, `0j`
# - Cualquier secuencia vacía, por ejemplo: `''`, `()`, `[]`
# - Los diccionarios vacíos: `{}`
# - Las instancias de cualquier clase, si esta define los métodos `__nonzero__()` o `__len__()`, cuando estos métodos devuelven `0` o `False`
#
# En cualquier otro caso, serán evaluados como `True`
# + [markdown] slideshow={"slide_type": "subslide"}
# ### ¿algún True en la lista?
# + slideshow={"slide_type": "-"}
values = [True, True, False, False]
i = 0
flag = False
while not flag and i < len(values):
flag = values[i]
i += 1
flag
# + slideshow={"slide_type": "fragment"}
# pythonico
any(values)
# + slideshow={"slide_type": "-"}
# pythonico
all(values)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### ¿algún 0 en la lista?
# + slideshow={"slide_type": "-"}
values = range(10) # [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
i = 0
flag = False
while not flag and i < len(values):
flag = (values[i] == 0)
i += 1
flag
# + slideshow={"slide_type": "fragment"}
#pythonico
any(values)
# + slideshow={"slide_type": "-"}
#pythonico
all(values)
# + [markdown] slideshow={"slide_type": "slide"}
# # Variables
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Intercambiar dos variables
# + slideshow={"slide_type": "-"}
a = 1
b = 5
print(f'a: {a} - b: {b}')
temp = a
a = b
b = temp
print(f'a: {a} - b: {b}')
# + slideshow={"slide_type": "subslide"}
# pythonico
a = 1
b = 5
print(f'a: {a} - b: {b}')
a, b = b, a
print(f'a: {a} - b: {b}')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Unpacking
# + slideshow={"slide_type": "-"}
l = ['Alicia', 'Pythonisa', '555-55-55']
name = l[0]
title = l[1]
phone = l[2]
# + slideshow={"slide_type": "fragment"}
# pythonico
name, title, phone = l
name, _, phone = l
# + [markdown] slideshow={"slide_type": "slide"}
# # Colecciones
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Listas
# + slideshow={"slide_type": "-"}
# Crear una listas
arr = ['one', 'two', 'three']
print(arr[0])
# + slideshow={"slide_type": "-"}
# Soportan múltiples tipos de datos:
arr.append(23)
print(arr)
# + slideshow={"slide_type": "-"}
# Son mutables:
arr[1] = 'hello'
print(arr)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Tuplas
# + slideshow={"slide_type": "-"}
# Crear una tupla
arr = 'one', 'two', 'three'
print(arr[0])
# + slideshow={"slide_type": "-"}
# Soportan múltiples tipos de datos:
print(arr + (23,))
# + slideshow={"slide_type": "-"}
# Son immutables:
arr[1] = 'hello'
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Named tuples
# + slideshow={"slide_type": "-"}
# namedtuples
from collections import namedtuple
Car = namedtuple('Car', 'color mileage automatic')
car1 = Car('red', 3812.4, True)
# + slideshow={"slide_type": "-"}
# Acceder a los campos:
car1.mileage
# + slideshow={"slide_type": "-"}
# Recuerda: Los campos son inmutables:
car1.mileage = 12
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Array Slicing
# + slideshow={"slide_type": "-"}
# array slicing
a = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
# + slideshow={"slide_type": "-"}
a[5]
# + slideshow={"slide_type": "-"}
a[-1]
# + slideshow={"slide_type": "-"}
a[2:8]
# + slideshow={"slide_type": "subslide"}
a[-4:-2]
# + slideshow={"slide_type": "-"}
a[2:8:2]
# + slideshow={"slide_type": "-"}
a[::-1]
# + slideshow={"slide_type": "-"}
a = [1, 2, 3, 4, 5]
# + slideshow={"slide_type": "-"}
a[2:3] = [0, 0]
a
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Sets
# + slideshow={"slide_type": "-"}
A = {1, 2, 3, 3}
print(A)
B = {3, 4, 5, 6, 7}
print(B)
# + slideshow={"slide_type": "-"}
A | B
# + slideshow={"slide_type": "-"}
A & B
# + slideshow={"slide_type": "-"}
A - B
# + slideshow={"slide_type": "-"}
A ^ B
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Counters
# + slideshow={"slide_type": "-"}
import collections
A = collections.Counter([1, 2, 2])
B = collections.Counter([2, 2, 3])
C = collections.Counter([1, 1, 2, 2, 3, 3, 3, 3, 4, 5, 6, 7])
# + slideshow={"slide_type": "subslide"}
print(A)
print(B)
print(C)
# + slideshow={"slide_type": "-"}
A | B
# + slideshow={"slide_type": "-"}
A & B
# + slideshow={"slide_type": "-"}
A + B
# + slideshow={"slide_type": "-"}
C.most_common()
# + slideshow={"slide_type": "-"}
C.most_common(1)
# + slideshow={"slide_type": "-"}
C.most_common(3)
# + slideshow={"slide_type": "-"}
C.most_common()[-1]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Bucles
# + [markdown] slideshow={"slide_type": "subslide"}
# Imprimir los elementos de una lista y su índice
# + slideshow={"slide_type": "-"}
a = ['Hello', 'world', '!']
i = 0
for x in a:
print(f'{i}: {x}')
i += 1
# + slideshow={"slide_type": "fragment"}
# pythonico
for i, x in enumerate(a):
print(f'{i}: {x}')
# + [markdown] slideshow={"slide_type": "subslide"}
# Obtener todos los cuadrados de los números del 0 al 9
# + slideshow={"slide_type": "-"}
squares = []
for x in range(10):
squares.append(x * x)
squares
# + slideshow={"slide_type": "fragment"}
# pythonico
[x * x for x in range(10)]
# + [markdown] slideshow={"slide_type": "subslide"}
# Obtener todos los cuadrados de los números pares del 0 al 9
# + slideshow={"slide_type": "-"}
even_squares = []
for x in range(10):
if x % 2 == 0:
even_squares.append(x * x)
even_squares
# + slideshow={"slide_type": "fragment"}
# pythonico
[x * x for x in range(10) if x % 2 == 0]
# + [markdown] slideshow={"slide_type": "subslide"}
# Obtener en una sola lista los pares nombre-apellido
# + slideshow={"slide_type": "-"}
names = ['John', 'Eric', 'Terry']
surnames = ['Cleese', 'Idle', 'Gilliam']
# + slideshow={"slide_type": "-"}
people = []
for i in range(len(names)):
people.append((names[i], surnames[i]))
print(people)
# + slideshow={"slide_type": "fragment"}
# list(zip(names, surnames))
[x for x in zip(names, surnames)]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Generadores
# + slideshow={"slide_type": "-"}
squares = (i*i for i in range(1000000))
squares
# + slideshow={"slide_type": "-"}
next(squares)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Deque
# + slideshow={"slide_type": "-"}
from collections import deque
q = deque(squares)
# + slideshow={"slide_type": "-"}
q.pop()
# + slideshow={"slide_type": "-"}
q.popleft()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Diccionarios
# + [markdown] slideshow={"slide_type": "subslide"}
# Valores por defecto
# + slideshow={"slide_type": "-"}
name_for_userid = {
382: 'Alice',
950: 'Bob',
590: 'Dilbert',
}
# + slideshow={"slide_type": "subslide"}
def greeting(userid):
if userid in name_for_userid:
return f'Hi {name_for_userid[userid]}!'
else:
return 'Hi there!'
# + slideshow={"slide_type": "fragment"}
def greeting(userid):
try:
return f'Hi {name_for_userid[userid]}!'
except KeyError:
return 'Hi there'
# + slideshow={"slide_type": "fragment"}
# pythonico
def greeting(userid):
return f'Hi {name_for_userid.get(userid)}!'
print(greeting(382))
print(greeting(999))
# + [markdown] slideshow={"slide_type": "subslide"}
# Editar los valores de un diccionario
# + slideshow={"slide_type": "-"}
visits = {
'Alice': 0,
'Bob': 0,
'Dilbert': 0,
}
# + slideshow={"slide_type": "subslide"}
def add_visit(user):
if user not in visits:
visits[user] = 0
visits[user] += 1
# + slideshow={"slide_type": "-"}
def add_visit(user):
visits[user] = visits.get(user, 0) + 1
# + slideshow={"slide_type": "fragment"}
def add_visit(user):
visits.setdefault(user, 0)
visits[user] += 1
# + slideshow={"slide_type": "fragment"}
from collections import defaultdict
visits = defaultdict(int)
def add_visit(user):
visits[user] += 1
add_visit('Alice')
add_visit('Juan')
visits
# + [markdown] slideshow={"slide_type": "subslide"}
# Comprensión de diccionarios
# + slideshow={"slide_type": "-"}
m = {x: x ** 2 for x in range(5)}
m
# + slideshow={"slide_type": "-"}
m = {x: 'A' + str(x) for x in range(5)}
m
# + slideshow={"slide_type": "-"}
# Invertir las clave-valor
m = {'a': 1, 'b': 2, 'c': 3, 'd': 4}
{v: k for k, v in m.items()}
#m.items()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Asserts
# + slideshow={"slide_type": "-"}
def apply_discount(product, discount):
price = int(product['price'] * (1.0 - discount))
assert 0 <= price <= product['price']
return price
# + slideshow={"slide_type": "subslide"}
shoes = {'name': 'Fancy Shoes', 'price': 14900}
apply_discount(shoes, 0.25)
# + slideshow={"slide_type": "-"}
apply_discount(shoes, 2.0)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Contextos
# + slideshow={"slide_type": "-"}
with open('../readme.md', 'r') as file_in:
print(file_in.read())
# + slideshow={"slide_type": "slide"}
import antigravity
| teoria/1.5_beautiful_python.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
// <h2>--- Day 17: Trick Shot ---</h2>
// [](https://mybinder.org/v2/gh/oddrationale/AdventOfCode2021FSharp/main?urlpath=lab%2Ftree%2FDay17.ipynb)
// <p>You finally decode the Elves' message. <code><span title="Maybe you need to turn the message 90 degrees counterclockwise?">HI</span></code>, the message says. You continue searching for the sleigh keys.</p>
// <p>Ahead of you is what appears to be a large <a href="https://en.wikipedia.org/wiki/Oceanic_trench" target="_blank">ocean trench</a>. Could the keys have fallen into it? You'd better send a probe to investigate.</p>
// <p>The probe launcher on your submarine can fire the probe with any <a href="https://en.wikipedia.org/wiki/Integer" target="_blank">integer</a> velocity in the <code>x</code> (forward) and <code>y</code> (upward, or downward if negative) directions. For example, an initial <code>x,y</code> velocity like <code>0,10</code> would fire the probe straight up, while an initial velocity like <code>10,-1</code> would fire the probe forward at a slight downward angle.</p>
// <p>The probe's <code>x,y</code> position starts at <code>0,0</code>. Then, it will follow some trajectory by moving in <em>steps</em>. On each step, these changes occur in the following order:</p>
// <ul>
// <li>The probe's <code>x</code> position increases by its <code>x</code> velocity.</li>
// <li>The probe's <code>y</code> position increases by its <code>y</code> velocity.</li>
// <li>Due to drag, the probe's <code>x</code> velocity changes by <code>1</code> toward the value <code>0</code>; that is, it decreases by <code>1</code> if it is greater than <code>0</code>, increases by <code>1</code> if it is less than <code>0</code>, or does not change if it is already <code>0</code>.</li>
// <li>Due to gravity, the probe's <code>y</code> velocity decreases by <code>1</code>.</li>
// </ul>
// <p>For the probe to successfully make it into the trench, the probe must be on some trajectory that causes it to be within a <em>target area</em> after any step. The submarine computer has already calculated this target area (your puzzle input). For example:</p>
// <pre><code>target area: x=20..30, y=-10..-5</code></pre>
// <p>This target area means that you need to find initial <code>x,y</code> velocity values such that after any step, the probe's <code>x</code> position is at least <code>20</code> and at most <code>30</code>, <em>and</em> the probe's <code>y</code> position is at least <code>-10</code> and at most <code>-5</code>.</p>
// <p>Given this target area, one initial velocity that causes the probe to be within the target area after any step is <code>7,2</code>:</p>
// <pre><code>.............#....#............
// .......#..............#........
// ...............................
// S........................#.....
// ...............................
// ...............................
// ...........................#...
// ...............................
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTT#TT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// </code></pre>
// <p>In this diagram, <code>S</code> is the probe's initial position, <code>0,0</code>. The <code>x</code> coordinate increases to the right, and the <code>y</code> coordinate increases upward. In the bottom right, positions that are within the target area are shown as <code>T</code>. After each step (until the target area is reached), the position of the probe is marked with <code>#</code>. (The bottom-right <code>#</code> is both a position the probe reaches and a position in the target area.)</p>
// <p>Another initial velocity that causes the probe to be within the target area after any step is <code>6,3</code>:</p>
// <pre><code>...............#..#............
// ...........#........#..........
// ...............................
// ......#..............#.........
// ...............................
// ...............................
// S....................#.........
// ...............................
// ...............................
// ...............................
// .....................#.........
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................T#TTTTTTTTT
// ....................TTTTTTTTTTT
// </code></pre>
// <p>Another one is <code>9,0</code>:</p>
// <pre><code>S........#.....................
// .................#.............
// ...............................
// ........................#......
// ...............................
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTT#
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// ....................TTTTTTTTTTT
// </code></pre>
// <p>One initial velocity that <em>doesn't</em> cause the probe to be within the target area after any step is <code>17,-4</code>:</p>
// <pre><code>S..............................................................
// ...............................................................
// ...............................................................
// ...............................................................
// .................#.............................................
// ....................TTTTTTTTTTT................................
// ....................TTTTTTTTTTT................................
// ....................TTTTTTTTTTT................................
// ....................TTTTTTTTTTT................................
// ....................TTTTTTTTTTT..#.............................
// ....................TTTTTTTTTTT................................
// ...............................................................
// ...............................................................
// ...............................................................
// ...............................................................
// ................................................#..............
// ...............................................................
// ...............................................................
// ...............................................................
// ...............................................................
// ...............................................................
// ...............................................................
// ..............................................................#
// </code></pre>
// <p>The probe appears to pass through the target area, but is never within it after any step. Instead, it continues down and to the right - only the first few steps are shown.</p>
// <p>If you're going to fire a highly scientific probe out of a super cool probe launcher, you might as well do it with <em>style</em>. How high can you make the probe go while still reaching the target area?</p>
// <p>In the above example, using an initial velocity of <code>6,9</code> is the best you can do, causing the probe to reach a maximum <code>y</code> position of <code><em>45</em></code>. (Any higher initial <code>y</code> velocity causes the probe to overshoot the target area entirely.)</p>
// <p>Find the initial velocity that causes the probe to reach the highest <code>y</code> position and still eventually be within the target area after any step. <em>What is the highest <code>y</code> position it reaches on this trajectory?</em></p>
// + dotnet_interactive={"language": "fsharp"}
let input = File.ReadAllText @"input/17.txt"
// + dotnet_interactive={"language": "fsharp"}
type Position = {
X: int
Y: int
}
type Velocity = {
dX: int
dY: int
}
// + dotnet_interactive={"language": "fsharp"}
let parse (input: string) =
let split1 = input.Split(": ").[1].Split(", ")
let splitX = split1.[0].Split("=").[1].Split("..")
let splitY = split1.[1].Split("=").[1].Split("..")
{
X = int splitX.[0]
Y = int splitY.[0]
},
{
X = int splitX.[1]
Y = int splitY.[1]
}
// + dotnet_interactive={"language": "fsharp"}
let rec steps (position, velocity) =
let drag = function
| dx when dx > 0 -> dx - 1
| dx when dx < 0 -> dx + 1
| _ -> 0
let step (position, velocity) =
{
X = position.X + velocity.dX
Y = position.Y + velocity.dY
},
{
dX = drag velocity.dX
dY = velocity.dY - 1
}
seq {
yield position, velocity
yield! steps (step (position, velocity))
}
// + dotnet_interactive={"language": "fsharp"}
let withinTargets steps (targetStart, targetEnd) =
steps
|> Seq.map (fun (p, _) -> p)
|> Seq.takeWhile (fun p ->
p.X <= targetEnd.X &&
p.Y >= targetStart.Y)
|> Seq.exists (fun p ->
targetStart.X <= p.X && p.X <= targetEnd.X &&
targetStart.Y <= p.Y && p.Y <= targetEnd.Y)
// -
let calcHeight velocity =
(velocity.dY*(velocity.dY + 1))/2
// + dotnet_interactive={"language": "fsharp"}
#!time
let (targetStart, targetEnd) = input |> parse
let hits =
([1..targetEnd.X], [targetStart.Y..abs targetStart.Y])
||> Seq.allPairs
|> Seq.map (fun (x, y) -> {dX = x; dY = y})
|> Seq.filter (fun v -> withinTargets (steps ({X = 0; Y = 0}, v)) (targetStart, targetEnd))
|> Seq.cache
hits
|> Seq.map calcHeight
|> Seq.max
// -
// <h2 id="part2">--- Part Two ---</h2>
// <p>Maybe a fancy trick shot isn't the best idea; after all, you only have one probe, so you had better not miss.</p>
// <p>To get the best idea of what your options are for launching the probe, you need to find <em>every initial velocity</em> that causes the probe to eventually be within the target area after any step.</p>
// <p>In the above example, there are <code><em>112</em></code> different initial velocity values that meet these criteria:</p>
// <pre><code>23,-10 25,-9 27,-5 29,-6 22,-6 21,-7 9,0 27,-7 24,-5
// 25,-7 26,-6 25,-5 6,8 11,-2 20,-5 29,-10 6,3 28,-7
// 8,0 30,-6 29,-8 20,-10 6,7 6,4 6,1 14,-4 21,-6
// 26,-10 7,-1 7,7 8,-1 21,-9 6,2 20,-7 30,-10 14,-3
// 20,-8 13,-2 7,3 28,-8 29,-9 15,-3 22,-5 26,-8 25,-8
// 25,-6 15,-4 9,-2 15,-2 12,-2 28,-9 12,-3 24,-6 23,-7
// 25,-10 7,8 11,-3 26,-7 7,1 23,-9 6,0 22,-10 27,-6
// 8,1 22,-8 13,-4 7,6 28,-6 11,-4 12,-4 26,-9 7,4
// 24,-10 23,-8 30,-8 7,0 9,-1 10,-1 26,-5 22,-9 6,5
// 7,5 23,-6 28,-10 10,-2 11,-1 20,-9 14,-2 29,-7 13,-3
// 23,-5 24,-8 27,-9 30,-7 28,-5 21,-10 7,9 6,6 21,-5
// 27,-10 7,2 30,-9 21,-8 22,-7 24,-9 20,-6 6,9 29,-5
// 8,-2 27,-8 30,-5 24,-7
// </code></pre>
// <p><em>How many distinct initial velocity values cause the probe to be within the target area after any step?</em></p>
// + dotnet_interactive={"language": "fsharp"}
#!time
hits
|> Seq.length
// -
// [Prev](Day16.ipynb) | [Next](Day18.ipynb)
| Day17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# These notes follow the official python tutorial pretty closely: http://docs.python.org/3/tutorial/
from __future__ import print_function
# # Lists
#
# Lists group together data. Many languages have arrays (we'll look at those in a bit in python). But unlike arrays in most languages, lists can hold data of all different types -- they don't need to be homogeneos. The data can be a mix of integers, floating point or complex #s, strings, or other objects (including other lists).
#
# A list is defined using square brackets:
a = [1, 2.0, "my list", 4]
print(a)
# We can index a list to get a single element -- remember that python starts counting at 0:
print(a[2])
# Like with strings, mathematical operators are defined on lists:
print(a*2)
# The `len()` function returns the length of a list
print(len(a))
# Unlike strings, lists are _mutable_ -- you can change elements in a list easily
a[1] = -2.0
a
a[0:1] = [-1, -2.1] # this will put two items in the spot where 1 existed before
a
# Note that lists can even contain other lists:
a[1] = ["other list", 3]
a
# ## Slicing
#
# We already mentioned above that it is possible to access individual elements from a string or a list using the square bracket notation. You will also find this notation for other object types in Python, for example tuples or Numpy arrays, so it's worth spending a bit of time looking at this in more detail.
#
# In addition to using positive integers, where ``0`` is the first item, it is possible to access list items with *negative* indices, which counts from the end: ``-1`` is the last element, ``-2`` is the second to last, etc:
li = [4, 67, 4, 2, 4, 6]
li[-1]
# You can also select **slices** from a list with the ``start:end:step`` syntax. Be aware that the last element is *not* included!
li[0:2]
li[:2] # ``start`` defaults to zero
li[2:] # ``end`` defaults to the last element
li[::2] # specify a step size
# Just like everything else in python, a list is an object that is the instance of a class. Classes have methods (functions) that know how to operate on an object of that class.
#
# There are lots of methods that work on lists. Two of the most useful are append, to add to the end of a list, and pop, to remove the last element:
a.append(6)
a
a.pop()
a
# <div style="background-color:yellow; padding: 10px"><h3><span class="fa fa-flash"></span> Quick Exercise:</h3></div>
#
# An operation we'll see a lot is to begin with an empty list and add elements to it. An empty list is created as:
# ```
# a = []
# ```
#
# * Create an empty list
# * Append the integers 1 through 10 to it.
# * Now pop them out of the list one by one.
#
# <hr>
# ### copying lists
#
# copying may seem a little counterintuitive at first. The best way to think about this is that your list lives in memory somewhere and when you do
#
# ```
# a = [1, 2, 3, 4]
# ```
#
# then the variable `a` is set to point to that location in memory, so it refers to the list.
#
# If we then do
# ```
# b = a
# ```
# then `b` will also point to that same location in memory -- the exact same list object.
#
# Since these are both pointing to the same location in memory, if we change the list through `a`, the change is reflected in `b` as well:
a = [1, 2, 3, 4]
b = a # both a and b refer to the same list object in memory
print(a)
a[0] = "changed"
print(b)
# if you want to create a new object in memory that is a copy of another, then you can either index the list, using `:` to get all the elements, or use the `list()` function:
c = list(a) # you can also do c = a[:], which basically slices the entire list
a[1] = "two"
print(a)
print(c)
# Things get a little complicated when a list contains another mutable object, like another list. Then the copy we looked at above is only a _shallow copy_. Look at this example—the list within the list here is still the same object in memory for our two copies:
f = [1, [2, 3], 4]
print(f)
g = list(f)
print(g)
# Now we are going to change an element of that list `[2, 3]` inside of our main list. We need to index `f` once to get that list, and then a second time to index that list:
f[1][0] = "a"
print(f)
print(g)
# Note that the change occured in both—since that inner list is shared in memory between the two. Note that we can still change one of the other values without it being reflected in the other list—this was made distinct by our shallow copy:
f[0] = -1
print(g)
print(f)
# Again, this is what is referred to as a shallow copy. If the original list had any special objects in it (like another list), then the new copy and the old copy will still point to that same object. There is a deep copy method when you really want everything to be unique in memory.
#
# When in doubt, use the `id()` function to figure out where in memory an object lies (you shouldn't worry about the what value of the numbers you get from `id` mean, but just whether they are the same as those for another object)
print(id(a), id(b), id(c))
# There are lots of other methods that work on lists (remember, ask for help)
my_list = [10, -1, 5, 24, 2, 9]
my_list.sort()
print(my_list)
print(my_list.count(-1))
my_list
help(a.insert)
a.insert(3, "my inserted element")
a
# joining two lists is simple. Like with strings, the `+` operator concatenates:
b = [1, 2, 3]
c = [4, 5, 6]
d = b + c
print(d)
# # Dictionaries
# A dictionary stores data as a key:value pair. Unlike a list where you have a particular order, the keys in a dictionary allow you to access information anywhere easily:
my_dict = {"key1":1, "key2":2, "key3":3}
print(my_dict["key1"])
# you can add a new key:pair easily, and it can be of any type
my_dict["newkey"] = "new"
print(my_dict)
# Note that a dictionary is unordered.
#
# You can also easily get the list of keys that are defined in a dictionary
keys = list(my_dict.keys())
print(keys)
# and check easily whether a key exists in the dictionary using the `in` operator
print("key1" in keys)
print("invalidKey" in keys)
# <div style="background-color:yellow; padding: 10px"><h3><span class="fa fa-flash"></span> Quick Exercise:</h3></div>
#
# Create a dictionary where the keys are the string names of the numbers zero to nine and the values are their numeric representation (0, 1, ... , 9)
#
# <hr>
# # List Comprehensions
#
# list comprehensions provide a compact way to initialize lists. Some examples from the tutorial
squares = [x**2 for x in range(10)]
squares
# here we use another python type, the tuple, to combine numbers from two lists into a pair
[(x, y) for x in [1,2,3] for y in [3,1,4] if x != y]
# <div style="background-color:yellow; padding: 10px"><h3><span class="fa fa-flash"></span> Quick Exercise:</h3></div>
#
# Use a list comprehension to create a new list from `squares` containing only the even numbers. It might be helpful to use the modulus operator, `%`
#
# <hr>
# # Tuples
#
# tuples are immutable -- they cannot be changed, but they are useful for organizing data in some situations. We use () to indicate a tuple:
a = (1, 2, 3, 4)
print(a)
# We can unpack a tuple:
w, x, y, z = a
print(w)
print(w, x, y, z)
# Since a tuple is immutable, we cannot change an element:
a[0] = 2
# But we can turn it into a list, and then we can change it
z = list(a)
z[0] = "new"
print(z)
# It is often not clear how tuples differ from lists. The most obvious way is that they are immutable. Often we'll see tuples used to store related data that should all be interpreted together. A good example is a Cartesian point, (x, y). Here is a list of points:
points = []
points.append((1,2))
points.append((2,3))
points.append((3,4))
points
# we can even generate these for a curve using a list comprehension:
points = [(x, 2*x + 5) for x in range(10)]
points
| day1/02. Python - Advanced datatypes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 条件生成式对抗网络
#
# 在[深度卷积生成式对抗网络](./dcgan.ipynb)这一节中,我们介绍了如何使用噪声数据生成人脸图像。使用DCGAN,我们可以用随机的一些向量生成一些图像。但是我们得到的是什么样的图像呢?我们能不能指定生成一张男人的脸或者女人的脸?
#
# 在本节中,我们将介绍[条件生成式对抗网络](https://arxiv.org/abs/1411.1784),它可以接受标签作为生成模型和判别模型的输入。这样,你就可以通过选择相应的标签来生成对应的数据。我们将以MNIST数据集为例进行训练。
# +
from __future__ import print_function
import numpy as np
from matplotlib import pyplot as plt
import mxnet as mx
from mxnet import gluon, test_utils, autograd
from mxnet import ndarray as nd
from mxnet.gluon import nn, utils
# -
# ## 设置训练参数
# +
epochs = 1000
batch_size = 64
label_size = 10
latent_z_size = 100
hidden_units = 128
img_wd = 28
img_ht = 28
use_gpu = True
ctx = mx.gpu() if use_gpu else mx.cpu()
lr = 0.001
# -
# ## 下载和处理MNIST数据集
# Pixel values of mnist image are normalized to be from 0 to 1.
mnist_data = test_utils.get_mnist()
train_data = mnist_data['train_data']
train_label = nd.one_hot(nd.array(mnist_data['train_label']), 10)
train_iter = mx.io.NDArrayIter(data=train_data, label=train_label, batch_size=batch_size)
# 可视化其中的四张图像:
# +
def visualize(img_arr):
plt.imshow((img_arr.asnumpy().reshape(img_wd, img_ht) * 255).astype(np.uint8), cmap='gray')
plt.axis('off')
for i in range(4):
plt.subplot(1,4,i+1)
visualize(nd.array(train_data[i + 10]))
plt.show()
# -
# ## 设计网络
#
# 在生成模型中,将随机的噪音向量和数字标签的独热编码向量拼接在一起作为其输入。随后是一个relu激活的全连接层。输出层则由另外一个sigmoid激活的全连接层构成。
#
# 和生成模型类似,将图像展平后的向量和数字标签的独热编码向量拼接在一起后作为判别模型的输入。随后是一个relu激活的全连接层。输出层则由另外一个全连接层构成。**(此处的英文教程有误)**在本教程中,我们在输出层中不使用sigmod激活函数,这样训练过程的数值稳定性会更好。
#
# 
# +
w_init = mx.init.Xavier()
# Build the generator
netG = nn.HybridSequential()
with netG.name_scope():
netG.add(nn.Dense(units=hidden_units, activation='relu', weight_initializer=w_init))
netG.add(nn.Dense(units=img_wd * img_ht, activation='sigmoid', weight_initializer=w_init))
# Build the discriminator
netD = nn.HybridSequential()
with netD.name_scope():
netD.add(nn.Dense(units=hidden_units, activation='relu', weight_initializer=w_init))
netD.add(nn.Dense(units=1, weight_initializer=w_init))
# -
# ## 设计损失函数和优化器
#
# 我们使用二分类的交叉熵损失函数作为损失函数,使用Adam进行优化。网络的初始化使用正态分布完成。
# +
# Loss
loss = gluon.loss.SigmoidBinaryCrossEntropyLoss()
# Initialize the generator and the discriminator
netG.initialize(ctx=ctx)
netD.initialize(ctx=ctx)
# Trainer for the generator and the discriminator
trainerG = gluon.Trainer(netG.collect_params(), 'adam', {'learning_rate': lr})
trainerD = gluon.Trainer(netD.collect_params(), 'adam', {'learning_rate': lr})
# -
# ## 训练
#
# 我们推荐你使用GPU进行训练,这样在几轮训练之后你就能够看到生成的数字图像。**(此处的英文教程有误)**
# +
from datetime import datetime
import time
import logging
real_label = nd.ones((batch_size,), ctx=ctx)
fake_label = nd.zeros((batch_size,),ctx=ctx)
def facc(label, pred):
pred = pred.ravel()
label = label.ravel()
return ((pred > 0.5) == label).mean()
metric = mx.metric.CustomMetric(facc)
stamp = datetime.now().strftime('%Y_%m_%d-%H_%M')
logging.basicConfig(level=logging.INFO)
for epoch in range(epochs):
tic = time.time()
btic = time.time()
train_iter.reset()
iter = 0
for batch in train_iter:
############################
# (1) Update D network: maximize log(D(x)) + log(1 - D(G(z)))
###########################
data = batch.data[0].as_in_context(ctx)
label = batch.label[0].as_in_context(ctx)
latent_z = mx.nd.random_normal(0, 1, shape=(batch_size, latent_z_size), ctx=ctx)
D_input = nd.concat(data.reshape((data.shape[0], -1)), label)
G_input = nd.concat(latent_z, label)
with autograd.record():
# train with real image
output = netD(D_input)
errD_real = loss(output, real_label)
metric.update([real_label,], [output,])
# train with fake image
fake = netG(G_input)
D_fake_input = nd.concat(fake.reshape((fake.shape[0], -1)), label)
output = netD(D_fake_input.detach())
errD_fake = loss(output, fake_label)
errD = errD_real + errD_fake
errD.backward()
metric.update([fake_label,], [output,])
trainerD.step(batch.data[0].shape[0])
############################
# (2) Update G network: maximize log(D(G(z)))
###########################
with autograd.record():
fake = netG(G_input)
D_fake_input = nd.concat(fake.reshape((fake.shape[0], -1)), label)
output = netD(D_fake_input)
errG = loss(output, real_label)
errG.backward()
trainerG.step(batch.data[0].shape[0])
# Print log infomation every ten batches
if iter % 10 == 0:
name, acc = metric.get()
logging.info('speed: {} samples/s'.format(batch_size / (time.time() - btic)))
logging.info('discriminator loss = %f, generator loss = %f, binary training acc = %f at iter %d epoch %d'
%(nd.mean(errD).asscalar(),
nd.mean(errG).asscalar(), acc, iter, epoch))
iter = iter + 1
btic = time.time()
name, acc = metric.get()
metric.reset()
logging.info('\nbinary training acc at epoch %d: %s=%f' % (epoch, name, acc))
logging.info('time: %f' % (time.time() - tic))
# Visualize one generated image for each epoch
fake_img = fake[0]
visualize(fake_img)
plt.show()
# -
# ## 结果
#
# 使用训练好的生成模型,我们生成几张数字图片。
num_image = 4
for digit in range(10):
for i in range(num_image):
latent_z = mx.nd.random_normal(0, 1, shape=(1, latent_z_size), ctx=ctx)
label = nd.one_hot(nd.array([[digit]]), 10).as_in_context(ctx)
img = netG(nd.concat(latent_z, label.reshape((1, 10))))
plt.subplot(10, 4, digit * 4 + i + 1)
visualize(img[0])
plt.show()
# For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| 03_conditional-gan_generate digits with different labels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <p align="center">
# <img src="https://github.com/GeostatsGuy/GeostatsPy/blob/master/TCG_color_logo.png?raw=true" width="220" height="240" />
#
# </p>
#
# ## Subsurface Data Analytics
#
# ### Tabular Data Structures / DataFrames in Python
#
# #### <NAME>, Associate Professor, University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
# ### Exercise: Tabular Data Structures / DataFrames in Python
#
# This is a tutorial for / demonstration of **Tabular Data Structures in Python**. In Python, the common tool for dealing with Tabular Data Structures is the DataFrame from the pandas Python package.
#
# * Tabular Data in subsurface data analytics includes any data set with a limited number of samples as oposed to gridded maps that provide exhaustively sampled data.
#
# This tutorial includes the methods and operations that would commonly be required for and Geoscientists, Engineers and Data Scientists working with Tabular Data Structures for the purpose of:
#
# 1. Data Checking and Cleaning
# 2. Data Mining / Inferential Data Analysis
# 3. Data Analytics / Building Predictive Models with Geostatistics and Machine Learning
#
# Learning to work with Pandas DataFrames is essential for dealing with tabular data (e.g. well data) in subsurface modeling workflows and for subsurface machine learning.
#
# ##### Tabular Data Structures
#
# In Python we will commonly store our data in two formats, tables and arrays. For sampled data with typically multiple features $1,\ldots,m$ over $1,\ldots,n$ samples we will work with tables. For exhaustive maps and models usually representing a single feature on a regular grid over $1,\ldots,n_{i}$ for $i = 1,\ldots,n_{dim}$ we will work with arrays.
#
# pandas package provides a convenient DataFrame object for working with data in a table and numpy package provides a convenient ndarray object for working with gridded data. In the following tutorial we will focus on DataFrames although we will utilize ndarrays a couple of times. There is another section on Gridded Data Structures that focuses on ndarrays.
#
# #### Additional Resources
#
# These workflows are based on standard methods with their associated limitations and assumptions. For more information see:
#
# * [pandas DataFrames Lecture](https://www.youtube.com/watch?v=cggieFcKdiM&list=PLG19vXLQHvSDUmEOmBoaxGbFAbvaLdfx4&index=8&t=0s)
#
# I have provided various workflows for subsurface data analytics, geostatistics and machine learning:
#
# * [Python](https://git.io/fh4eX)
#
# * [Excel](https://github.com/GeostatsGuy/LectureExercises/blob/master/Lecture7_CI_Hypoth_eg_R.xlsx)
# * [R](https://github.com/GeostatsGuy/LectureExercises/blob/master/Lecture7_CI_Hypoth_eg.R)
#
# and all of my University of Texas at Austin
#
# * [Lectures](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig/featured?view_as=subscriber)
#
# #### Workflow Goals
#
# Learn the basics for working with Tabular Data Structures in Python. This includes:
#
# * Loading tabular data
# * Visualizing tabular data
# * Data QC and Cleaning
# * Interacting with the tabular data
#
# #### Objective
#
# I want to provide hands-on experience with building subsurface modeling workflows. Python provides an excellent vehicle to accomplish this. I have coded a package called GeostatsPy with GSLIB: Geostatistical Library (Deutsch and Journel, 1998) functionality that provides basic building blocks for building subsurface modeling workflows.
#
# The objective is to remove the hurdles of subsurface modeling workflow construction by providing building blocks and sufficient examples. This is not a coding class per se, but we need the ability to 'script' workflows working with numerical methods.
#
# #### Getting Started
#
# Here's the steps to get setup in Python with the GeostatsPy package:
#
# 1. Install Anaconda 3 on your machine (https://www.anaconda.com/download/).
# 2. From Anaconda Navigator (within Anaconda3 group), go to the environment tab, click on base (root) green arrow and open a terminal.
# 3. In the terminal type: pip install geostatspy.
# 4. Open Jupyter and in the top block get started by copy and pasting the code block below from this Jupyter Notebook to start using the geostatspy functionality.
#
# You will need to copy the data file to your working directory. They are available here:
#
# * Tabular data - 2D_MV_200wells.csv at [here].(https://github.com/GeostatsGuy/GeoDataSets/blob/master/2D_MV_200wells.csv)
#
# I have put together various subsurface workflows for data analytics, geostatistics and machine learning. Go [here](https://git.io/fh4eX) for other example workflows and source code.
#
# #### Load the required libraries
#
# The following code loads the required libraries.
#
import os # to set current working directory
import numpy as np # arrays and matrix math
import pandas as pd # DataFrames
# If you get a package import error, you may have to first install some of these packages. This can usually be accomplished by opening up a command window on Windows and then typing `python -m pip install [package-name]`. More assistance is available with the respective package docs.
#
#
# #### Set the working directory
#
# I always like to do this so I don't lose files and to simplify subsequent read and writes (avoid including the full address each time). Also, in this case make sure to place the required (see below) data file in this directory. When we are finished with this tutorial we will write our new dataset back to this directory.
os.chdir("c:/PGE383") # set the working directory
# #### Loading Data
#
# Let's load the provided multivariate, spatial dataset. '2D_MV_200wells.csv' is available [here](https://github.com/GeostatsGuy/GeoDataSets/blob/master/2D_MV_200wells.csv). It is a comma delimited file with:
#
# * X and Y coordinates ($m$)
# * facies 1 and 2 (1 is sandstone and 2 interbedded sand and mudstone)
# * porosity (fraction)
# * permeability ($mD$)
# * acoustic impedance ($\frac{kg}{m^3} \cdot \frac{m}{s} \cdot 10^6$).
#
# We load it with the pandas 'read_csv' function into a data frame we called 'df' and then preview it by printing a slice and by utilizing the 'head' DataFrame member function (with a nice and clean format, see below).
#
# **Python Tip: using functions from a package** just type the label for the package that we declared at the beginning:
#
# ```python
# import pandas as pd
# ```
#
# so we can access the pandas function 'read_csv' with the command:
#
# ```python
# pd.read_csv()
# ```
#
# but read csv has required input parameters. The essential one is the name of the file. For our circumstance all the other default parameters are fine. If you want to see all the possible parameters for this function, just go to the docs [here](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html).
#
# * The docs are always helpful
# * There is often a lot of flexibility for Python functions, possible through using various inputs parameters
#
# also, the program has an output, a pandas DataFrame loaded from the data. So we have to specficy the name / variable representing that new object.
#
# ```python
# df = pd.read_csv("2D_MV_200wells.csv")
# ```
#
# Let's run this command to load the data and then look at the resulting DataFrame to ensure that we loaded it.
df = pd.read_csv("2D_MV_200wells.csv") # read a .csv file in as a DataFrame
#print(df.iloc[0:5,:]) # display first 4 samples in the table as a preview
df.head() # we could also use this command for a table preview
# #### Summary Statistics
#
# It is useful to review the summary statistics of our loaded DataFrame. That can be accomplished with the 'describe' DataFrame member function. We transpose to switch the axes for ease of visualization.
df.describe()
# #### Rename a Variable / Features
#
# Let's rename the facies, permeability and acoustic impedance for convenience.
df = df.rename(columns={'facies_threshold_0.3': 'facies','permeability':'perm','acoustic_impedance':'ai'}) # rename columns of the
df.head()
# #### Slicing and Subsets
#
# It is straightforward to extract subsets from a DataFrame to make a new DataFrame.
#
# * This is useful for cleaning up data by removing features that are no longer of interest.
#
# If the samples are in random order then the first $n_{s}$ samples are a random sample of size $n_{s}$. Below we make a new DataFrame, 'df_subset', with the rows 0 to 4 and columns 2 to 6 and the **X and Y coordinates removed**.
df_subset = df.iloc[0:5,2:7] # make a new dataframe with just the first 4 samples and no X,Y
print(df_subset)
# Let's demonstrate some more complicated slicing options. We demonstrate two methods:
#
# * list the exact indexes that you want
#
# ```python
# df_subset2 = df.iloc[[0,2,4,5,10,43],:] # extract rows 0,2,4,5...,43 for all columns
# ```
# +
df_subset2 = df.iloc[[0,2,4,5,10,43],:] # new dataframe with samples 0, 2 ,...,43 and all features
print(df_subset2)
df_subset3 = df.iloc[2:,[2,4,5]] # new dataframe with all samples from 2 and features 2,4,5
print(df_subset3)
# -
# #### Adding a Variable / Features
#
# It is also easy to add a column to our data frame.
#
# * Note, we assume that the array is in the same order as the DataFrame. This could be an issue if any rows were removed form either before adding etc.
#
# To demonstrate we make a 1D numpy array of zeros using the 'zeros' function and add it to our DataFrame with the feature name indicated as 'zero'.
zeros = np.zeros(200) # make a array of zeros
df['zero'] = pd.Series(zeros) # add the array to our DataFrame
df.head()
# We can also remove unwanted columns without having to subset the DataFrame.
#
# * That's why we just added a column of zeros, I wanted to also demonstrated removing a column.
#
# We do this with the 'drop' member function of the DataFrame object. We just have the give the column name and by indicating axis=1 we specify to drop a column instead of a row.
df = df.drop('zero',axis=1) # remove the zero column
df.head()
# #### Standardizing / Manipulating Variables / Features
#
# We may want to make new features by using mathematical operators applied to existing features.
#
# * We can use any combinations of features, constants, math and even other data tables
#
# For example, we can make a porosity feature that is in percentage instead of fraction (called 'porosity100') or a ratio of permeability divided by porosity (called 'permpor') may be useful for subsequent calculations such as the Lorenz Coefficient.
df['porosity100'] = df['porosity']*100 # add a new column with porosity in percentage
df['permpor'] = df['perm']/df['porosity'] # add a new feature with ratio of perm / por
df.head()
# #### Truncating and Categorizing Variables / Features
#
# We could also use conditional statements when assigning values to a new feature.
#
# * We could use any condition with any combination of features and variables from any sourc
#
# For example, we could have a categorical porosity measure for high and low porosity, called 'tporosity' for truncated porosity.
df['tporosity'] = np.where(df['porosity']>=0.12, 'high', 'low') # conditional statement assign a new feature
df.head()
# #### Truncating to Remove Nonphysical Values
#
# Here's an example where we use a conditional statement to assign a very low permeability value (0.0001 mD) for all porosity values below a threshold. Of course, this is for demonstration, in practice a much lower porosity threshold would likely be applied.
df['perm_cutoff'] = np.where(df['porosity']>=0.12, df['perm'],0.0001) # conditional statement assign a new feature
df.head()
# #### Dealing with Missing Data
#
# What about missing or invalid values? Let's assign a single porosity value to NaN, 'not a number', indicating a missing or eroneous value.
#
# **Python Tip: manipulating DataFrames manually**
#
# We us this command to access anyone of the individual records in our data table.
#
# ```p
# df.at[irow,'feature name']
# ```
#
# We can get the value or set the value to a new value:
#
# ```p
# df.at[irow,'feature name'] = new_value
# ```
#
# We will then check for the number of NaN values in our DataFrame. Then we can search for and display the sample with the NaN porosity value.
# +
df.at[1,'porosity'] = np.NaN # let's give ourselves a NaN / missing value in our table
# Count the number of samples with missing values
print('Number of null values in our DataFrame = ', str(df.isnull().sum().sum()))
# Find the row(s) with missing values and look at them
nan_rows = df[df['porosity'].isnull()] # find the row with missing values
print(nan_rows)
# -
# We can see that sample 1 (see the '1' on the left hand side, that is the sample index in the table) has a NaN porosity value.
#
# #### Drop Samples with Missing Values
#
# Now we may choose to remove the sample with the NaN. The 'dropna' DataFrame member function will remove all samples with NaN entries from the entire DataFrame. By visualizing the index at the left of the DataFrame preview you can confirm that sample 1 is removed (it jumps from 0 to 2).
df = df.dropna() # drop any rows (samples) with atleast one missing value
df.head()
# #### Searching with Tabular Data
#
# One could extract samples into a new DataFrame with multiple criteria. This is shown below.
df_extract = df.loc[(df['porosity'] > 0.12) & (df['perm'] > 10.0)] # extract with multiple conditions to a new table
df_extract.head()
# #### Making Data Frames
#
# We already covered the idea of making a DataFrame by loading data from a file.
#
# It is also possible to build a brandnew DataFrame from a set of 1D arrays.
#
# * Note, they must have the same size and be sorted consistently.
#
# We will extract 'porosity' and 'perm' features as arrays.
#
# **Python Tip: extracting data from DataFrames**
#
# We can extract the data for a single feature with this command:
#
# ```python
# 1D_series = df['feature_name']
# ```
#
# The 'series' retains information about the feature that was included in the DataFrame including the name and the indexing. This is fine, but some methods don't work with series so we can also extract the data as a 1D ndarray. By adding the '.values' the series is converted to a 1D array.
#
# ```python
# 1D_ndarray = df['feature_name'].values
# ```
#
# We then use the pandas DataFrame command to make a new DataFrame with each 1D array and the column names specified as 'porosity' and 'permeabilty'.
por = df['porosity'].values # extract porosity column as vector
perm = df['perm'].values # extract permeability column as vector
df_new = pd.DataFrame({'porosity': por, 'permeability': perm}) # make a new DataFrame from the vectors
df_new.head()
# #### Information About Our Tabular Data
#
# We can reach in and retrieve the actual raw information in the DataFrame including the column names and actual values as an numpy array.
#
# * We can't edit them like this, but we can access and use this information.
#
# This includs:
#
# 1. 'index' with information about the index (i.e. index from start to stop with step)
# 2. 'columns' with the names of the features
# 3. 'values' with the data table entries as a 2D array.
#
# Let's look at these components of our DataFrame:
print(df.index) # get information about the index
print(df.columns) # get the list of feature names
print(df.values) # get the 2D array with all the table data
# Here's a method for getting a list of the DataFrame feature names:
list(df) # get a list with the feature names
# #### More Precise Information from Tabular Data
#
# Let's interact with the DataFrame more surgically, one feature and sample at a time. Here we retrieve the 4th column feature name and the porosity value for sample \#1.
col2_name = df.columns[3] # get the name of the 4th feature (porosity)
print(col2_name)
por1 = df.values[1,3] # get the value for sample 1 of the 4th feature (porosity)
print('Porosity value for sample number 1 is ' + str(por1) + '.')
# We can also manually change values.
#
# * We can use the 'at' pandas DataFrame member function to get and set manually individual records.
#
# We look up the porosity value for sample 1 and then we use the 'at' again DataFrame member function to change the value to 0.1000.
por = df.at[2,'porosity'] # get the value for sample 1 of the porosity feature
print('The value of porosity for sample 2 is ' + str(por) + '.')
df.at[2,'porosity'] = 0.10 # set the value for sample 1 of the porosity feature
print('The value of porosity for sample 2 is now 0.1000.')
df.head()
# #### Sorting a DataFrame
#
# Let's try sorting our unconventional wells in descending order. An example application would be to calculate the Lorenz ceofficient.
#
# We can use the command
#
# ```python
# df.sort_values('permpor')
# ```
df = df.sort_values('permpor', ascending = False)
df.head(n=13)
# #### Reseting Indices
#
# The DataFrame indices are now out of order. Let's reset the DataFrame indices with:
#
# ```python
# df.reset_index()
# ```
df = df.reset_index()
df
# #### Shallow and Deep Copies
#
# Let's explore the topic of shallow and deep copies with data frames.
#
# * many Python methods assumes shallow copies, this confuses many people
#
# * this means when we do this:
#
# ```python
# data_frame2 = data_frame
# ```
#
# we are creating a pointer to the original DataFrame, not a new DataFrame!
#
# * the pointer to the object, is known as a shallow copy
#
# * anything we do to the shallow copy will be done to the original object
#
# To demonstrate this, let's make a copy of the DataFrame, as a slice with facies, permeability and acoustic impedance.
df2 = df
df_subset2.head()
# Now let's go ahead and change the facies for sample '0' to 5
#
# * we can use the .at member function again to change a single value in a Pandas DataFrame
df2.loc[0,'facies'] = 5
df2.head(n=5)
# Now let's check the original DataFrame
df.head(n=5)
# Let's make a new feature, add it to the copy of the DataFrame and find out if it is in the original DataFrame.
df2['random'] = np.random.rand(len(df2))
df.head(n=5)
# We added an array to the copy of the DataFrame and it was added to the original DataFrame!
#
# Let's make a deep copy and try again.
df3 = df.copy()
df3.loc[1,'facies'] = 7
df3.head()
# Let's check the original.
df.head()
# We made a deep copy so now the edits to the copy do not impact the original object.
#
# * be careful when copying and check if you have a deep or shallow copy
#
# #### Writing the Tabular Data to a File
#
# It may be useful to write the DataFrame out for storage or curation and / or to be utilize with another platform (even R or Excel!). It is easy to write the DataFrame back to a comma delimited file. We have the 'to_csv' DataFrame member function to accomplish this. The file will write to the working directory (another reason we set that at the beginning). Go to that folder and open this new file with TextPad, Excel or any other program that opens .txt files to check it out.
df.to_csv("2D_MV_200wells_out.csv") # write out the df DataFrame to a comma delimited file
# #### More Exercises
#
# There are so many more exercises and tests that one could attempt to gain experience with the pandas package, DataFrames objects in Python. I'll end here for brevity, but I invite you to continue. Check out the docs at https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html. I'm always happy to discuss,
#
# *Michael*
#
# #### The Author:
#
# ### <NAME>, Associate Professor, University of Texas at Austin
# *Novel Data Analytics, Geostatistics and Machine Learning Subsurface Solutions*
#
# With over 17 years of experience in subsurface consulting, research and development, Michael has returned to academia driven by his passion for teaching and enthusiasm for enhancing engineers' and geoscientists' impact in subsurface resource development.
#
# For more about Michael check out these links:
#
# #### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
# #### Want to Work Together?
#
# I hope this content is helpful to those that want to learn more about subsurface modeling, data analytics and machine learning. Students and working professionals are welcome to participate.
#
# * Want to invite me to visit your company for training, mentoring, project review, workflow design and / or consulting? I'd be happy to drop by and work with you!
#
# * Interested in partnering, supporting my graduate student research or my Subsurface Data Analytics and Machine Learning consortium (co-PIs including Profs. Foster, Torres-Verdin and van Oort)? My research combines data analytics, stochastic modeling and machine learning theory with practice to develop novel methods and workflows to add value. We are solving challenging subsurface problems!
#
# * I can be reached at <EMAIL>.
#
# I'm always happy to discuss,
#
# *Michael*
#
# <NAME>, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
| SubsurfaceDataAnalytics_DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Density-based clustering
#
# In density-based clustering the approach is different compared to distributed clustering. We need to implement all functions from scratch. The libraries that we are going to use are the same as in previous example, but in this case we have also the random package that is used to shuffle the objects in the data set.
import random
import numpy as np
import pandas as pd
from math import sqrt
# DBScan is an example of a density-based clustering method. The goal is to find all element where the neighborhood is defined as:
# \begin{equation}
# N_{\epsilon}:{q|d(p,q)\leq\epsilon},
# \end{equation}
# where $p$ and $q$ are two elements of the training data set and $\epsilon$ is the neighborhood distance. For the data set used before and $\epsilon$ to 0.25 we get the regions like in figure below.
#
# 
#
# Let's setup the variables as in previous examples. The are three new ones like ```distance_matrix```, ```max_distance```, ```number_of_cluster```, and ``min_points``. The first one is clear, the second is a parameter that can be changed, depending on that how many neighborhood elements we would like to concider. The next variable is about the number of clusters that are calculated during clustering. It's not the exact number of clusters, but allow us count the clusters during clustering. Last variable is the number of points that needs to within a neighbourhood to be classified as non-border object. Boarded points are the points that are the farest points from the cluster, but it's not the noise.
# +
# %store -r data_set
assignation = np.zeros(len(data_set))
distance_matrix = np.zeros((len(data_set), len(data_set)))
max_distance = 0.25
number_of_cluster = 0
min_points = 2
# -
# We need the distance function that we used in previous example to calculate the distance matrix:
def calculate_distance(x,v):
return sqrt((x[0]-v[0])**2+(x[1]-v[1])**2)
# To calculate the distance matrix we use the calculate_distance that we used previously:
def calculate_distance_matrix():
distance_matrix = np.zeros((len(data_set),len(data_set)))
for i in range(len(data_set)):
for j in range(len(data_set)):
distance_matrix[i, j] = calculate_distance(data_set[i], data_set[j])
return distance_matrix
# The next step is to get closest elements in the feature space:
def get_closest_elements(distance_matrix, element_id):
element_distances = distance_matrix[element_id]
filtered = {}
iter = 0
for element in element_distances:
if element < max_distance:
filtered[iter] = element
iter = iter + 1
return filtered
# The last step before cluster function is to define funtions that mark the elements in our data set that are known to be a noise or were already visited by our method.
# +
def set_as_noise(assignation,element_id):
assignation[element_id] = -1
return assignation
def set_visited(elements, assignation, number_of_clusters):
for element_id in elements.keys():
assignation[element_id] = number_of_clusters
return assignation
# -
# Combine it all together:
def cluster_density(assignation):
number_of_cluster = 0
distance_matrix = calculate_distance_matrix()
element_ids = list(range(len(data_set)))
random.shuffle(element_ids)
for i in element_ids:
if assignation[i] != 0:
continue
closest = get_closest_elements(distance_matrix, i)
if len(closest) < min_points:
assignation = set_as_noise(assignation,i)
else:
assignation = set_visited(closest, assignation, number_of_cluster)
number_of_cluster = number_of_cluster + 1
return assignation
# Ready to cluster:
new_assignation_density = cluster_density(assignation)
# The number of cluster is the size of unique cluster ids that are in ``new_assignation_density`` minus noise.
print("Number of clusters: "+ str(len(np.unique(new_assignation_density))-1))
# The noise is marked with -1. The other objects have the cluster number assigned.
print(new_assignation_density)
# %store new_assignation_density
| ML1/clustering/043Clustering_Density.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/ml-unison/regresion-logistica/master/imagenes/ml-unison.png" width="250">
# # Regularización en regresión logística
#
# [**<NAME>**](http://mat.uson.mx/~juliowaissman/), 1 de octubre de 2020.
#
# ## Curso Reconocimiento de Patrones
# ### Licenciatura en Ciencias de la Computación
# ### Universidad de Sonora
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (10,5)
plt.style.use('ggplot')
# -
# ## 1. La regresión logística ya programada
#
# Esto, antes de agregarle la regularización
# La función logística está dada por
#
# $$
# \sigma(z) = \frac{1}{1 + e^{-z}},
# $$
#
# la cual es importante que podamos calcular en forma vectorial. Si bien el calculo es de una sola linea, el uso de estas funciones auxiliares facilitan la legibilidad del código.
#
# #### Desarrolla la función logística, la cual se calcule para todos los elementos de un ndarray.
def logistica(z):
"""
Calcula la función logística para cada elemento de z
@param z: un ndarray
@return: un ndarray de las mismas dimensiones que z
"""
return 1 / (1 + np.exp(-z))
# Y ahora vamos a ver como se ve la función logística
z = np.linspace(-5, 5, 100)
plt.plot( z, logistica(z))
plt.title(u'Función logística', fontsize=20)
plt.xlabel(r'$z$', fontsize=20)
plt.ylabel(r'$\frac{1}{1 + \exp(-z)}$', fontsize=26)
plt.show()
# Una vez establecida la función logística, vamos a implementar la función de error *sin regularizar* (error *en muestra*), la cual está dada por
#
# $$
# E_{in}(w, b) = -\frac{1}{M} \sum_{i=1}^M \left[ y^{(i)}\log(a^{(i)}) + (1 - y^{(i)})\log(1 - a^{(i)})\right],
# $$
#
# donde
#
# $$
# a^{(i)} = \sigma(z^{(i)}), \quad\quad z^{(i)} = w^T x^{(i)} + b
# $$
# + tags=[]
def error_in(x, y, w, b):
"""
Calcula el error en muestra para la regresión logística
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param w: un ndarray de dimensión (n, ) con los pesos
@param b: un flotante con el sesgo
@return: un flotante con el valor de pérdida
"""
y_est = logistica(x @ w + b)
return -np.nansum([
np.log(y_est[y > 0.5]).sum(),
np.log(1 - y_est[y < 0.5]).sum()
]) / y_est.shape[0]
# El testunit del pobre (ya lo calcule yo, pero puedes hacerlo a mano para estar seguro)
w = np.array([1])
b = 1.0
x = np.array([[10],
[-5]])
y1 = np.array([1, 0])
y2 = np.array([0, 1])
y3 = np.array([0, 0])
y4 = np.array([1, 1])
y_est = logistica(x @ w + b)
assert abs(error_in(x, y1, w, b) - (-np.log(y_est[0]) - np.log(1 - y_est[1])) / 2) < 1e-2
assert abs(error_in(x, y2, w, b) - (-np.log(1 - y_est[0]) - np.log(y_est[1])) / 2) < 1e-2
assert abs(error_in(x, y3, w, b) - (-np.log(1 - y_est[0]) - np.log(1 - y_est[1])) / 2) < 1e-2
assert abs(error_in(x, y4, w, b) - (-np.log(y_est[0]) - np.log(y_est[1])) / 2) < 1e-2
# -
# De la misma manera, para poder implementar las funciones de aprendizaje, vamos a implementar el gradiente de la función de pérdida. El gradiente de la función de pérdida respecto a $\omega$, $\nabla E_{in}(\omega)$, y la derividada parcial respecto al sesgo se obtienen con las siguientes ecuaciones:
#
# $$
# \frac{\partial E_{in}(w, b)}{\partial w_j} = -\frac{1}{M} \sum_{i=1}^M \left(y^{(i)} - a^{(i)}\right)x_j^{(i)}
# $$
#
# $$
# \frac{\partial E_{in}(w, b)}{\partial b} = -\frac{1}{M} \sum_{i=1}^M \left(y^{(i)} - a^{(i)}\right)
# $$
#
# Todo esto **para el caso sin regularizar**.
def gradiente_error(x, y, w, b):
"""
Calcula el gradiente de la función de error en muestra.
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param w: un ndarray de dimensión (n, ) con los pesos
@param b: un flotante con el sesgo
@return: dw, db, un ndarray de mismas dimensiones que w y un flotnte con el cálculo de
la dervada evluada en el punto w y b
"""
M = x.shape[0]
error = y - logistica(x @ w + b)
dw = -x.T @ error / M
db = - error.mean()
return dw, db
# +
# Otra vez el testunit del pobre (ya lo calcule yo, pero puedes hacerlo a mano para estar seguro)
w = np.array([1])
b = 1.0
x = np.array([[10],
[-5]])
y1 = np.array([1, 0])
y2 = np.array([0, 1])
y3 = np.array([0, 0])
y4 = np.array([1, 1])
assert abs(0.00898475 - gradiente_error(x, y1, w, b)[1]) < 1e-4
assert abs(7.45495097 - gradiente_error(x, y2, w, b)[0]) < 1e-4
assert abs(4.95495097 - gradiente_error(x, y3, w, b)[0]) < 1e-4
assert abs(-0.49101525 - gradiente_error(x, y4, w, b)[1]) < 1e-4
# -
# ## 2. Descenso de gradiente
# Ahora vamos a desarrollar las funciones necesarias para realizar el entrenamiento y encontrar la mejor $\omega$ de acuero a la función de costos y un conjunto de datos de aprendizaje.
#
# Para ilustrar el problema, vamos a utilizar una base de datos sintética proveniente del curso de [Andrew Ng](www.andrewng.org/) que se encuentra en [Coursera](https://www.coursera.org).
#
# Supongamos que pertenecemos al departamente de servicios escolares de la UNISON y vamos a modificar el procedimiento de admisión. En lugar de utilizar un solo exámen (EXCOBA) y la información del cardex de la preparatoria, hemos decidido aplicar dos exámenes, uno sicométrico y otro de habilidades estudiantiles. Dichos exámenes se han aplicado el último año aunque no fueron utilizados como criterio. Así, tenemos un historial entre estudiantes aceptados y resultados de los dos exámenes. El objetivo es hacer un método de regresión que nos permita hacer la admisión a la UNISON tomando en cuenta únicamente los dos exámenes y simplificar el proceso. *Recuerda que esto no es verdad, es solo un ejercicio*.
#
# Bien, los datos se encuentran en el archivo `admision.txt` el cual se encuentra en formato `csv` (osea los valores de las columnas separados por comas. Vamos a leer los datos y graficar la información para entender un poco los datos.
# +
datos = np.loadtxt('admision.csv', comments='%', delimiter=',')
x, y = datos[:,0:-1], datos[:,-1]
plt.plot(x[y == 1, 0], x[y == 1, 1], 'sr', label='aceptados')
plt.plot(x[y == 0, 0], x[y == 0, 1], 'ob', label='rechazados')
plt.title(u'Ejemplo sintético para regresión logística')
plt.xlabel(u'Calificación del primer examen')
plt.ylabel(u'Calificación del segundo examen')
plt.axis([20, 100, 20, 100])
plt.legend(loc=0)
# -
# Vistos los datos un clasificador lineal podría ser una buena solución. Ahora vamos a implementar el método de descenso de gradiente.
def dg(x, y, alpha, max_iter=10000, tol=1e-6, historial=False):
"""
Descenso de gradiente por lotes
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param alpha: Un flotante (típicamente pequeño) con la tasa de aprendizaje
@param tol: Un flotante pequeño como criterio de paro. Por default 1e-6
@param max_iter: Máximo numero de iteraciones. Por default 1e4
@param historial: Un booleano para saber si guardamos el historial de errores
@return: w, b, hist donde
- w es ndarray de dimensión (n, ) con los pesos;
- b es un float con el sesgo
- hist, un ndarray de dimensión (max_iter,)
con el valor del error en muestra en cada iteración.
Si historial == False, entonces perdida_hist = None.
"""
#Obtención del tamaño de x
M, n = x.shape
#w y b se inicializan como ceros
w = np.zeros(n)
b = 0.0
hist = [error_in(x, y, w, b)] if historial else None
for epoch in range(1, max_iter):
dw, db = gradiente_error(x, y, w, b)
w -= alpha * dw
b -= alpha * db
error = error_in(x, y, w, b)
if historial:
hist.append(error)
if np.abs(np.max(dw)) < tol:
break
return w, b, hist
# Para probar la función de aprendizaje, vamos a aplicarla a nuestro problema de admisión. Primero recuerda que tienes que hacer una exploración para encontrar el mejor valor de $\epsilon$. Así que utiliza el código de abajo para ajustar $\alpha$.
# +
# Distintos valores para alpha
alpha_vec = [1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 0.1, 1.0]
#número máximo de iteraciones
mi = 50
#Obtención de las curvas de aprendizaje
w0, b0, hist_0 = dg(x, y, alpha_vec[0], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-6
w0, b0, hist_1 = dg(x, y, alpha_vec[1], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-5
w0, b0, hist_2 = dg(x, y, alpha_vec[2], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-4
w0, b0, hist_3 = dg(x, y, alpha_vec[3], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-3
# w0, b0, hist_4 = dg(x, y, alpha_vec[4], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-2
# w0, b0, hist_5 = dg(x, y, alpha_vec[5], max_iter=mi, tol=1e-4, historial=True) #alpha=1e-1
# w0, b0, hist_6 = dg(x, y, alpha_vec[6], max_iter=mi, tol=1e-4, historial=True) #alpha=1
#Los últimos tres valores presentan errores relacionados con divisiones entre cero :OO !!!
# +
plt.plot(np.arange(mi), hist_0, 'magenta') #alpha=1e-6
plt.plot(np.arange(mi), hist_1, 'purple') #alpha=1e-5
plt.plot(np.arange(mi), hist_2, 'cyan') #alpha=1e-4
plt.plot(np.arange(mi), hist_3, 'green') #alpha=1e-3
# plt.plot(np.arange(mi), hist_4, 'yellow') #alpha=1e-2
# plt.plot(np.arange(mi), hist_5, 'orange') #alpha=1e-1
# plt.plot(np.arange(mi), hist_6, 'red') #alpha=1
# plt.title(r'Evolucion del valor del error en muestra para distintos valores de $\alpha$')
plt.title(r'Evolucion del valor de la función de error para distintos valores de $\alpha$')
plt.xlabel('Iteraciones')
# plt.ylabel('Error en muestra')
plt.ylabel('Pérdida')
# -
# El valor de $\alpha=10^{-4}$ será el elegido por ser el que decrece con mayor rapidez.
# Una vez encontrado el mejor $\epsilon$, entonces podemos calcular $\omega$ (esto va a tardar bastante), recuerda que el costo final debe de ser lo más cercano a 0 posible, así que agrega cuantas iteraciones sean necesarias (a partir de una función de pérdida con un valor de al rededor de 0.22 ya está bien). Puedes ejecutar la celda cuandas veces sea necesario con un número limitado de iteraciones (digamos unas 10,000) para ver como evoluciona. Esto podría mejorar sensiblementa si se normalizan los datos de entrada. Tambien puedes variar alpha.
# + tags=[]
alpha = 1e-4
w, b, hist_error = dg(x, y, 20*alpha, max_iter = 100000, tol=1e-4, historial=False)
print("Los pesos obtenidos son: \n{}".format(w))
print("El sesgo obtenidos es: \n{}".format(b))
print("El valor final de la función de pérdida es: {}".format(error_in(x, y, w, b)))
# -
# Es interesante ver como el descenso de gradiente no es muy eficiente en este tipo de problemas, a pesar de ser problemas de optimización convexos.
#
# Bueno, este método nos devuelve $\omega$, pero esto no es suficiente para decir que tenemos un clasificador, ya que un método de clasificación se compone de dos métodos, uno para **aprender** y otro para **predecir**.
#
# Recuerda que $a^{(i)} = \Pr[y^{(i)} = 1 | x^{(i)} ; w, b]$, y a partir de esta probabilidad debemos tomar una desición. Igualmente recuerda que para tomar la desicion no necesitamos calcular el valor de la logística, si conocemos el umbral.
# #### Desarrolla una función de predicción.
def predictor(x, w, b):
"""
Predice los valores de y_hat (que solo pueden ser 0 o 1), utilizando el criterio MAP.
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param w: un ndarray de dimensión (n, ) con los pesos
@param b: un flotante con el sesgo
@return: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0 con la salida estimada
"""
return np.where(logistica(x @ w + b) > 0, 1, 0)
# ¿Que tan bueno es este clasificador? ¿Es que implementamos bien el método?
#
# Vamos a contestar esto por partes. Primero, vamos a graficar los mismos datos pero vamos a agregar la superficie de separación, la cual en este caso sabemos que es una linea recta. Como sabemos el criterio para decidir si un punto pertenece a la clase distinguida o no es si el valor de $w^T x^{(i)} + b \ge 0$, por lo que la frontera entre la región donde se escoge una clase de otra se encuentra en:
#
# $$
# 0 = b + w_1 x_1 + w_2 x_2,
# $$
#
# y despejando:
#
# $$
# x_2 = -\frac{b}{w_2} -\frac{w_1}{w_2}x_1
# $$
#
# son los pares $(x_1, x_2)$ los valores en la forntera. Al ser estos (en este caso) una linea recta solo necesitamos dos para graficar la superficie de separación.
#
# + tags=[]
x1_frontera = np.array([20, 100]) #Los valores mínimo y máximo que tenemos en la gráfica de puntos
x2_frontera = -(b / w[1]) - (w[0] / w[1]) * x1_frontera
print(x1_frontera)
print(x2_frontera)
plt.plot(x[y == 1, 0], x[y == 1, 1], 'sr', label='aceptados')
plt.plot(x[y == 0, 0], x[y == 0, 1], 'ob', label='rechazados')
plt.plot(x1_frontera, x2_frontera, 'm')
plt.title(u'Ejemplo sintético para regresión logística')
plt.xlabel(u'Calificación del primer examen')
plt.ylabel(u'Calificación del segundo examen')
plt.axis([20, 100, 20, 100])
plt.legend(loc=0)
# -
# ## 3. Clasificación polinomial
#
# Como podemos ver en la gráfica de arriba, parece ser que la regresión logística aceptaría a algunos estudiantes rechazados y rechazaría a algunos que si fueron en realidad aceptados. En todo método de clasificación hay un grado de error, y eso es parte del poder de generalización de los métodos.
#
# Sin embargo, una simple inspección visual muestra que, posiblemente, la regresión lineal no es la mejor solución, ya que la frontera entre las dos clases parece ser más bien una curva.
#
# ¿Que tal si probamos con un clasificador cuadrático? Un clasificador cuadrático no es más que la regresión lineal pero a la que se le agregan todos los atributos que sean una combinación de dos de los atributos.
#
# Por ejemplo, si un ejemplo $x = (x_1, x_2, x_3)^T$ se aumenta con todas sus componentes cuadráticas, entonces tenemos los atributos
#
# $$
# \phi_2(x) = (x_1, x_2, x_3, x_1 x_2, x_1 x_3, x_2 x_3, x_1^2, x_2^2, x_3^2)^T.
# $$
#
# De la misma manera se pueden obtener clasificadores de orden tres, cuatro, cinco, etc. En general a estos clasificadores se les conoce como **clasificadores polinomiales**. Ahora, para entender bien la idea, vamos a resolver el problema anterior con un clasificador de orden 2.
#
# Sin embargo, si luego se quiere hacer el reconocimiento de otros objetos, o cambiar el orden del polinomio, pues se requeriría de reclcular cada vez la expansión polinomial. Vamos a generalizar la obtención de atributos polinomiales con la función `map_poly`, la cual la vamos a desarrollar a continuación.
#
# En este caso, la normalización de los datos es muy importante, por lo que se agregan las funciones pertinentes.
#
# +
from itertools import combinations_with_replacement
def map_poly(grad, x):
"""
Encuentra las características polinomiales hasta el grado grad de la matriz de datos x,
asumiendo que x[:n, 0] es la expansión de orden 1 (los valores de cada atributo)
@param grad: un entero positivo con el grado de expansión
@param x: un ndarray de dimension (M, n) donde n es el número de atributos
@return: un ndarray de dimensión (M, n_phi) donde
n_phi = \sum_{i = 1}^grad fact(i + n - 1)/(fact(i) * fact(n - 1))
"""
if int(grad) < 2:
raise ValueError('grad debe de ser mayor a 1')
M, n = x.shape
atrib = x.copy()
x_phi = x.copy()
for i in range(2, int(grad) + 1):
for comb in combinations_with_replacement(range(n), i):
x_phi = np.c_[x_phi, np.prod(atrib[:, comb], axis=1)]
return x_phi
def medias_std(x):
"""
Obtiene un vector de medias y desviaciones estandar para normalizar
@param x: Un ndarray de (M, n) con una matriz de diseño
@return: mu, des_std dos ndarray de dimensiones (n, ) con las medias y desviaciones estandar
"""
return np.mean(x, axis=0), np.std(x, axis=0)
def normaliza(x, mu, des_std):
"""
Normaliza los datos x
@param x: un ndarray de dimension (M, n) con la matriz de diseño
@param mu: un ndarray (n, ) con las medias
@param des_std: un ndarray (n, ) con las desviaciones estandard
@return: un ndarray (M, n) con x normalizado
"""
return (x - mu) / des_std
# -
# **Realiza la clasificación de los datos utilizando un clasificador cuadrático (recuerda ajustar primero el valor de $\alpha$)**
# +
# Encuentra phi_x (x son la expansión polinomial de segundo orden, utiliza la función map_poly
phi_x = map_poly(2, x)
mu, de = medias_std(phi_x)
phi_x_norm = normaliza(phi_x, mu, de)
# Distintos valores para alpha
alpha_vec = [1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 0.1, 1.0]
# Número de iteraciones
N = 500
#Pocas para poder revisar varios valores para alpha
# Utiliza la regresión logística
#Obtención de las curvas de aprendizaje
w, b, hist_0 = dg(phi_x_norm, y, alpha_vec[0], max_iter=N, tol=1e-4, historial=True)
w, b, hist_1 = dg(phi_x_norm, y, alpha_vec[1], max_iter=N, tol=1e-4, historial=True)
w, b, hist_2 = dg(phi_x_norm, y, alpha_vec[2], max_iter=N, tol=1e-4, historial=True)
w, b, hist_3 = dg(phi_x_norm, y, alpha_vec[3], max_iter=N, tol=1e-4, historial=True)
w, b, hist_4 = dg(phi_x_norm, y, alpha_vec[4], max_iter=N, tol=1e-4, historial=True)
w, b, hist_5 = dg(phi_x_norm, y, alpha_vec[5], max_iter=N, tol=1e-4, historial=True)
w, b, hist_6 = dg(phi_x_norm, y, alpha_vec[6], max_iter=N, tol=1e-4, historial=True)
#Gráficas
plt.plot(np.arange(N), hist_0, 'magenta') #alpha=1e-6
plt.plot(np.arange(N), hist_1, 'purple') #alpha=1e-5
plt.plot(np.arange(N), hist_2, 'cyan') #alpha=1e-4
plt.plot(np.arange(N), hist_3, 'green') #alpha=1e-3
plt.plot(np.arange(N), hist_4, 'yellow') #alpha=1e-2
# plt.plot(np.arange(N), hist_5, 'orange') #alpha=1e-1
# plt.plot(np.arange(N), hist_6, 'red') #alpha=1
# plt.title(r'Curva de aprendizaje para distintos valores de $\alpha$ ')
plt.xlabel('Iteraciones')
plt.ylabel(r'$E_{in}$')
plt.title('Evaluación del parámetro alpha')
# + tags=[]
alpha = 1e-3
N = 50000
w_norm, b_norm, historial_error = dg(phi_x_norm, y, alpha, N, tol=1e-4, historial=False)
print("Los pesos obtenidos son: \n{}".format(w_norm))
print("El sesgo obtenidos es: \n{}".format(b_norm))
print("El error en muestra es: {}".format(error_in(phi_x_norm, y, w_norm, b_norm)))
# -
# donde se puede encontrar un error en muestra de aproximadamente 0.03.
#
# Esto lo tenemos que graficar. Pero graficar la separación de datos en una proyección en las primeras dos dimensiones, no es tan sencillo como lo hicimos con una separación lineal, así que vamos atener que generar un `contour`, y sobre este graficar los datos. Para esto vamos a desarrollar una función.
def plot_separacion2D(x, y, grado, mu, de, w, b):
"""
Grafica las primeras dos dimensiones (posiciones 1 y 2) de datos en dos dimensiones
extendidos con un clasificador polinomial así como la separación dada por theta_phi
"""
if grado < 2:
raise ValueError('Esta funcion es para graficar separaciones con polinomios mayores a 1')
x1_min, x1_max = np.min(x[:,0]), np.max(x[:,0])
x2_min, x2_max = np.min(x[:,1]), np.max(x[:,1])
delta1, delta2 = (x1_max - x1_min) * 0.1, (x2_max - x2_min) * 0.1
spanX1 = np.linspace(x1_min - delta1, x1_max + delta1, 600)
spanX2 = np.linspace(x2_min - delta2, x2_max + delta2, 600)
X1, X2 = np.meshgrid(spanX1, spanX2)
X = normaliza(map_poly(grado, np.c_[X1.ravel(), X2.ravel()]), mu, de)
Z = predictor(X, w, b)
Z = Z.reshape(X1.shape[0], X1.shape[1])
# plt.contour(X1, X2, Z, linewidths=0.2, colors='k')
plt.contourf(X1, X2, Z, 1, cmap=plt.cm.binary_r)
plt.plot(x[y > 0.5, 0], x[y > 0.5, 1], 'sr', label='clase positiva')
plt.plot(x[y < 0.5, 0], x[y < 0.5, 1], 'oy', label='clase negativa')
plt.axis([spanX1[0], spanX1[-1], spanX2[0], spanX2[-1]])
# Y ahora vamos a probar la función `plot_separacion2D` con los datos de entrenamiento. El comando tarda, ya que estamos haciendo un grid de 200 $\times$ 200, y realizando evaluaciones individuales.
plot_separacion2D(x, y, 2, mu, de, w_norm, b_norm)
plt.title(u"Separación con un clasificador cuadrático")
plt.xlabel(u"Calificación del primer examen")
plt.ylabel(u"Calificación del segundo examen")
# Como podemos ver, un clasificador polinomial de orden 2 clasifica mejor los datos de aprendizaje, y además parece suficientemente simple para ser la mejor opción para hacer la predicción. Claro, esto lo sabemos porque pudimos visualizar los datos, y en el fondo estamos haciendo trampa, al seleccionar la expansión polinomial a partir de una inspección visual de los datos.
#
# Tomemos ahora una base de datos que si bien es sintética es representativa de una familia de problemas a resolver. Supongamos que estámos opimizando la fase de pruebas dentro de la linea de producción de la empresa Microprocesadores del Noroeste S.A. de C.V.. La idea es reducir el banco de pruebas de cada nuevo microprocesador fabricado y en lugar de hacer 50 pruebas, reducirlas a 2. En el conjunto de datos tenemos los valores que obtuvo cada componente en las dos pruebas seleccionadas, y la decisión que se tomo con cada dispositivo (esta desición se tomo con el banco de 50 reglas). Los datos los podemos visualizar a continuación.
# +
datos = np.loadtxt('prod_test.csv', comments='%', delimiter=',')
x, y = datos[:,0:-1], datos[:,-1]
plt.plot(x[y == 1, 0], x[y == 1, 1], 'or', label='cumple calidad')
plt.plot(x[y == 0, 0], x[y == 0, 1], 'ob', label='rechazado')
plt.title(u'Ejemplo de pruebas de un producto')
plt.xlabel(u'Valor obtenido en prueba 1')
plt.ylabel(u'Valor obtenido en prueba 2')
plt.legend(loc=0)
# -
# Cláramente este problema no se puede solucionar con un clasificador lineal (1 orden), por lo que hay que probar otros tipos de clasificadores.
#
# **Completa el código para hacer regresión polinomial para polinomios de orden 2, 4, 6 y 8, y muestra los resultados en una figura. Recuerda que este ejercicio puede tomar bastante tiempo de cómputo. Posiblemente tengas que hacer ajustes en el código para manejar diferentes valores de alpha y max_iter de acuerdo a cada caso**
for (i, grado) in enumerate([2, 6, 10, 14]):
# Genera la expansión polinomial
phi_x = map_poly(grado, x)
# Normaliza
mu, dev = medias_std(phi_x)
phi_x_norm = normaliza( phi_x, mu, dev )
# Entrena
alpha = 1e-2
N = 1000
w0, b0, hist = dg(phi_x_norm, y, alpha, N, tol=1e-4, historial=False)
# Muestra resultados con plot_separacion2D
plt.subplot(2, 2, i + 1)
plt.title(f"Polinomio de grado {grado}")
plot_separacion2D(x, y, grado, mu, dev, w0, b0)
# ## 4. Regularización
#
# Como podemos ver del ejercicio anterior, es dificil determinar el grado del polinomio, y en algunos casos es demasiado general (subaprendizaje) y en otros demasiado específico (sobreaprendizaje). ¿Que podría ser la solución?, bueno, una solución posible es utilizar un polinomio de alto grado (o relativamente alto), y utilizar la **regularización** para controlar la generalización del algoritmo, a través de una variable $\lambda$.
#
# Recordemos, la función de costos de la regresión logística con regularización es:
#
# $$
# costo(w, b) = E_{in}(w, b) + \frac{\lambda}{M} regu(w),
# $$
#
# donde $regu(w)$ es una regularización, la cual puede ser $l_1$, $l_2$ u otras, tal como vimos en clase.
#
# Además:
# $$
# E_{in}(w, b) = -\frac{1}{M} \sum_{i=1}^M \left[ y^{(i)}\log(a^{(i)}) + (1 - y^{(i)})\log(1 - a^{(i)})\right],
# $$
#
# donde
#
# $$
# a^{(i)} = \sigma(z^{(i)}), \quad\quad z^{(i)} = w^T x^{(i)} + b
# $$
#
#
# **Completa el siguiente código, utilizando una regularización en $L_2$:**
#
# $$
# regu(w) = \sum_{i=0}^M W_i^2
# $$
def costo(x, y, w, b, lambd):
"""
Calcula el costo de una w dada para el conjunto dee entrenamiento dado por y y x,
usando regularización
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param w: un ndarray de dimensión (n, ) con los pesos
@param b: un flotante con el sesgo
@param lambd: un flotante con el valor de lambda en la regularizacion
@return: un flotante con el valor de pérdida
"""
costo = 0
M = x.shape[0]
#------------------------------------------------------------------------
E_in = error_in(x, y, w, b)
Reg = w.T @ w
costo = E_in + (lambd/M)*Reg
#------------------------------------------------------------------------
return costo
def grad_regu(x, y, w, b, lambd):
"""
Calcula el gradiente de la función de costo regularizado para clasificación binaria,
utilizando una neurona logística, para w y b y conociendo un conjunto de aprendizaje.
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param w: un ndarray de dimensión (n, ) con los pesos
@param b: un flotante con el sesgo
@param lambd: un flotante con el peso de la regularización
@return: dw, db, un ndarray de mismas dimensiones que w y un flotnte con el cálculo de
la dervada evluada en el punto w y b
"""
M = x.shape[0]
dw = np.zeros_like(w)
db = 0.0
#------------------------------------------------------------------------
# Agregua aqui tu código
error = y - logistica(x @ w + b)
dw = (-x.T @ error / M) + (2*lambd/M)*w
db = - error.mean()
#------------------------------------------------------------------------
return dw, db
def dg_regu(x, y, w, b, alpha, lambd, max_iter=10_000, tol=1e-4, historial=False):
"""
Descenso de gradiente con regularización l2
@param x: un ndarray de dimensión (M, n) con la matriz de diseño
@param y: un ndarray de dimensión (M, ) donde cada entrada es 1.0 o 0.0
@param alpha: Un flotante (típicamente pequeño) con la tasa de aprendizaje
@param lambd: Un flotante con el valor de la regularización
@param max_iter: Máximo numero de iteraciones. Por default 10_000
@param tol: Un flotante pequeño como criterio de paro. Por default 1e-4
@param historial: Un booleano para saber si guardamos el historial. Falso por default
@return:
- w: ndarray de dimensión (n, ) con los pesos;
- b: float con el sesgo
- hist: ndarray de (max_iter,) el historial de error.
Si historial == False, entonces hist = None.
"""
M, n = x.shape
hist = [costo(x, y, w, b, lambd)] if historial else None
for epoch in range(1, max_iter):
dw, db = grad_regu(x, y, w, b, lambd)
w -= alpha * dw
b -= alpha * db
error = costo(x, y, w, b, lambd)
if historial:
hist.append(error)
if np.abs(np.max(dw)) < tol:
break
return w, b, hist
#
# **Desarrolla las funciones y scriprs necesarios para realizar la regresión logística con un polinomio de grado 14 y con cuatro valores de regularización diferentes. Grafica la superficie de separación para cuatro valores diferentes de $\lambda$. Escribe tus conclusiones**
phi_x = map_poly(14, x)
# Normaliza
mu, dev = medias_std(phi_x)
phi_x_norm = normaliza(phi_x, mu, dev)
# +
phi_x = map_poly(14, x)
# Normaliza
mu, dev = medias_std(phi_x)
phi_x_norm = normaliza(phi_x, mu, dev)
for (i, lambd) in enumerate([0, 1, 10, 100]):
# Entrena
w = np.zeros(phi_x.shape[1])
b = 0
alpha = 1e-2
N = 1000
w0, b0, hist = dg_regu(phi_x_norm, y, w, b, alpha, lambd, N, tol=1e-4, historial=False)
# Muestra resultados con plot_separacion2D
plt.subplot(2, 2, i + 1)
plt.title("Polinomio de grado 14,\n lambda = {}.".format(lambd))
plot_separacion2D(x, y, 14, mu, dev, w0, b0)
# -
# **Conclusiones:** No alcanzo a ver la diferencia entre las imágenes de arriba :(
#
# Aún así, el valor de $\lambda$ debe ajustarse para reducir o aumentar el error dentro de muestra.
| regularizacion_logistica.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# ## Empty World
#
# Spawn a robot in an empty world.
roslaunch fetch_gazebo simulation.launch
# ## Playground world
#
# Spawns a robot inside a lab-like test environment. This environment has some tables with items that may be picked up and manipulated. It also has a pre-made map which can be used to test out robot navigation and some simple demonstrations of object grasping.
roslaunch fetch_gazebo playground.launch
# ## Simulating a Freight
#
# Freight uses the same launch files as Fetch, simply pass the robot argument:
roslaunch fetch_gazebo simulation.launch robot:=freight
| notebooks/fetch_world.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (pytorch_src2)
# language: python
# name: pytorch_src2
# ---
import torch
import pandas as pd
import numpy as np
import seaborn as sns
import os
sns.set(style="darkgrid")
import matplotlib.pyplot as plt
from glob import glob
# %matplotlib inline
def get_title(filename):
"""
>>> get_title("logs/0613/0613-q1-0000.train")
'0613-q1-0000'
"""
return os.path.splitext(os.path.basename(filename))[0]
def get_df_from_file(f):
df = pd.read_csv(f)
df = df[df["is_end_of_epoch"]].reset_index()
return df
result_files = sorted(glob("../../../data/logs/0708*.train"))
titles = [get_title(f) for f in result_files]
dfes = (get_df_from_file(f) for f in result_files)
def do_plot(df, title):
dfval = df[df["is_val"]]
dftrain = df[df["is_val"] != True]
sns.lineplot(x=dftrain.index, y="drmsd", data=dftrain, label="train-drmsd")
sns.lineplot(x=dfval.index, y="drmsd", data=dfval, label="val-drmsd",color="lightblue")
sns.lineplot(x=dftrain.index, y="rmsd", data=dftrain, label="train-rmsd")
sns.lineplot(x=dfval.index, y="rmsd", data=dfval, label="val-rmsd", color="orange")
sns.lineplot(x=dftrain.index, y="rmse", data=dftrain, label="rmse")
sns.lineplot(x=dftrain.index, y="combined", data=dftrain, label="drmsd+mse")
plt.ylabel("Loss Value")
plt.xlabel("Epoch")
plt.legend(loc=(1.04,.7))
plt.title("{} Training Loss".format(title))
# plt.savefig("../figs/transtrain.pdf", pad_inches=1, bbox_inches="tight")
do_plot(get_df_from_file(result_files[0]), titles[0])
min_key = "rmsd"
mins = []
for df, title in zip(dfes, titles):
try:
dfval = df
except KeyError as e:
print(e)
continue
try:
row = dfval[dfval[min_key] == dfval[min_key].min()]
except KeyError:
print(title)
continue
row["title"] = title[:]
mins.append(row)
mins_df = pd.concat(mins)
mins_df.sort_values(min_key, inplace=True)
mins_df
names = [t for t in mins_df["title"][:10]]
for n in names:
loc = None
for i in range(len(result_files)):
if n == os.path.splitext(os.path.basename(result_files[i]))[0]:
loc = i
do_plot(get_df_from_file(result_files[loc]), titles[loc])
plt.show()
mean_rmsds = mins_df[mins_df["title"].str.contains("mean")]["rmsd"].values
rand_rmsds = mins_df[mins_df["title"].str.contains("rand")]["rmsd"].values
mean_rmsds
sns.boxplot(x=["w/angle means", "random"], y=[mean_rmsds, rand_rmsds])
plt.ylabel("RMSD")
plt.xlabel("Initialization Strategy")
plt.ylim((.75, 5))
plt.title("Models trained on DRMSD+RMSE loss")
plt.savefig("model_overfit_initialization_strategies_combined.png",dpi=300)
plt.savefig("model_overfit_initialization_strategies_combined.svg")
dfes_mean = (get_df_from_file(f) for f in result_files if "mean" in f)
dfes_rand = (get_df_from_file(f) for f in result_files if "rand" in f)
first = True
for dm, dr in zip(dfes_mean, dfes_rand):
if first:
lr = "random"
lm = "mean"
first = False
else:
lr = None
lm = None
try:
sns.lineplot(x=dr.index, y="rmsd", data=dr, alpha=0.4, color="C1", label=lr)
except ValueError:
pass
try:
sns.lineplot(x=dm.index, y="rmsd", data=dm, alpha=0.4, color="C0", label=lm)
except ValueError:
continue
plt.legend()
plt.title("Training Loss Over Time - DRMSD+RMSE")
plt.xlabel("Epoch")
plt.savefig("mean_vs_random_training_over_time_combined.png", dpi=300)
plt.savefig("mean_vs_random_training_over_time_combined.svg")
| research/analysis/190710/190710 Comparison of Angle Mean vs Random Ititialization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import json
import pickle
import random
import math
import re
import tweepy
import string
from tweepy.api import TooManyRequests
import time
# +
#Twitter API credentials
consumer_key = "9Wj6RDG1XbCiIUf6DOrk6siyA"
consumer_secret = "<KEY>"
access_key = "<KEY>"
access_secret = "<KEY>"
#authorize twitter, initialize tweepy
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_key, access_secret)
api = tweepy.API(auth)
# -
df_train_all = pd.read_json('raw_data/train.json')
df_test_all = pd.read_json('raw_data/test.json')
df_val_all = pd.read_json('raw_data/val.json')
df_train_all.columns
columns = ['OriginalTweetID', 'otweet_content', 'url']
df_train = df_train_all[columns]
df_test = df_test_all[columns]
df_val = df_val_all[columns]
df_train.head()
print(df_train.shape)
print(df_test.shape)
print(df_val.shape)
df_train.iloc[7]
punc = ['.',',','!','?']
def clean_up(s: str):
if type(s) is float and math.isnan(s):
# handling nans
return ""
else:
# remove URLs
s = re.sub('((www\.[^\s]+)|(https?://[^\s]+)|(http?://[^\s]+))', '', s)
s = re.sub(r'http\S+', '', s)
# remove content in <...>
s = re.sub(r'<[\w\d]+>', '', s)
# remove special characters - ()#%^&*@! etc.
s = ''.join(e for e in s if e.isalnum() or e is ' ')
# remove multiple spaces
s = re.sub("\s+"," ",s)
return s.strip()
def fetch_tweet(df: pd.DataFrame):
idxs_not_found = []
i = 0
while i < df.shape[0]:
if i%100 == 0:
print(f"At Index => {i}")
id = df.iloc[i]['OriginalTweetID']
try:
status = api.get_status(id, tweet_mode="extended")
tweet_polluted = status.full_text
df_train.loc[i,'otweet_content'] = tweet_polluted
i+=1
except TooManyRequests:
# Catches Rate Limit Exception
print(f"Too Many Requests => {i}")
time.sleep(60*16) # Delay for 16 mins (1 minute buffer) as only 900 requests per 15 mins is allowed
except Exception:
# Catches Exception if tweet ID is not found or forbidden
idxs_not_found.append(i)
# tweet_polluted = df.iloc[i]['otweet_content']
# df_train.loc[i,'otweet_content'] = clean_up(tweet_polluted)
i+=1
return df, idxs_not_found
# Creating Validation Json file
df_val_final, val_records_not_found = fetch_tweet(df_val)
df_val_final.to_json('val_tweet.json')
print(f"Out of {df_val_final.shape[0]} data samples, {val_records_not_found} tweets were not found in API")
# Creating Test Json file
df_test_final, test_records_not_found = fetch_tweet(df_test)
df_test_final.to_json('test_tweet.json')
print(f"Out of {df_test_final.shape[0]} data samples, {test_records_not_found} tweets were not found in API")
# Creating Training Json file
df_train_final, train_records_not_found = fetch_tweet(df_train)
df_train_final.to_json('train_tweet.json')
print(f"Out of {df_train_final.shape[0]} data samples, {train_records_not_found} tweets were not found in API")
print(f"Out of {df_val_final.shape[0]} data samples, {len(val_records_not_found)} tweets were not found in API")
print(f"Out of {df_test_final.shape[0]} data samples, {len(test_records_not_found)} tweets were not found in API")
print(f"Out of {df_train_final.shape[0]} data samples, {len(train_records_not_found)} tweets were not found in API")
| blink/preprocess/fetch_complete_tweet_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ColdCoffee21/Foundations-of-Data-Science/blob/master/plotting_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ewqcNuIcZ2w9"
# ##Class 4
# + [markdown] id="UbPP1Datl05d"
# ###bar chart
# + id="L5aPMPgvZ4Tx" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="8d056f59-7e28-46a9-8c54-732a0833dd84"
id1 = np.arange(1,10)
score = np.arange(20,110,10)
plt.bar(id1,score)
plt.xlabel('Student ID')
plt.ylabel('Score')
plt.show()
# + id="j0Yaa4YLaWkX" colab={"base_uri": "https://localhost:8080/", "height": 318} outputId="599e054c-a91e-4d4d-b179-aadf812baff8"
#get commands from others
#barh
plt.bar(id1,score,color="Red")
ax = plt.axes()
ax.set_facecolor("#ECF0FF") # Setting the background color by specifying the HEX Codeplt.bar(id1,score,color = '#FFA726')
# + id="BEZpGeMxfdFn" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="728e42a7-fe4b-4f7e-e93a-fcef127a8366"
#Plotting multiple sets of data
x1= [1,3,5,7]
x2=[2,4,6,8]
y1 = [7,7,7,7]
y2= [17,18,29,40]
plt.figure(figsize=(8,6))
ax = plt.axes()
ax.set_facecolor("white")
plt.bar(x1,y1,label = "First",color = '#42B300')
# First set of data
plt.bar(x2,y2,label = "Second",color = '#94E413')
# Second set of data
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + [markdown] id="zHoKHf0nl9wR"
# ###horizontal bar chart
# + id="ftYttDjzgHDK" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="10a978d9-c530-4efb-c538-601d47aa822c"
# Horizontal Bar Chart
Age = [28,33,43,45,57]
Name = ["Asif", "Steve", 'John', "Ravi", "Basit"]
plt.barh(Name,Age, color ="yellowgreen")
plt.show()
# + id="hm_L73zGgbog" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="7176a571-3849-4be1-9fe7-c48ce8061fd3"
# Changing the width of Bars
num1 = np.array([1,3,5,7,9])
num2 = np.array([2,4,6,8,10])
plt.figure(figsize=(8,4))
plt.bar(num1, num1**2, width=0.2 , color = '#FF6F00')
plt.bar(num2, num2**2, width=0.2 , color = '#FFB300')
plt.plot()
# + id="B9TMXhdqgoOX" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="afe08b82-0d77-44cc-d89f-b235ba5960be"
num1 = np.array([1,3,5,7,9])
num2 = np.array([2,4,6,8,10])
plt.figure(figsize=(8,4))
plt.bar(num1, num1**2, width=0.9 , color = '#FF6F00',edgecolor='black')
plt.bar(num2, num2**2, width=0.5 , color = '#FFB300',edgecolor='black')
for x,y in zip(num1,num1**2):
plt.text(x, y-5, '%d' % y, ha='center' , va= 'bottom',fontsize = 10)
for x,y in zip(num2,num2**2):
plt.text(x, y+0.05, '%d' % y, ha='center' , va= 'bottom',fontsize = 5)
plt.plot()
# + [markdown] id="dai3WljHmD71"
# ###Stacked charts
# + id="GDACnj8Wg6pY" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="932b4dec-babc-4f05-df73-ada6daa42884"
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + id="9OB4eg9Djtv8" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="eecdd7e8-7acd-43e1-949c-e933048cc07b"
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= np.array([17,18,29,40])
y2 =np.array([20,21,22,23])
y3 =np.array([5,9,11,12])
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.bar(x1,y3,label = "Cancelled Tickets",width = 0.5 ,bottom = y1+y2 , color = '#F7DC6F')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + id="ygUj1AdMko48" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="eec1f4a5-8fe9-47ef-8654-89fc26c91e55"
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= np.array([17,18,29,40])
y2 =np.array([20,21,22,23])
y3 =np.array([5,9,11,12])
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.bar(x1,y3,label = "Cancelled Tickets",width = 0.5 ,bottom = y1+y2 , color = '#F7DC6F')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + id="031zNDfik_8A" colab={"base_uri": "https://localhost:8080/", "height": 349} outputId="64b37777-1e54-4ed7-d60a-e10cbd621740"
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(8,5))
plt.barh(x1,y1,label = "Open Tickets",color = '#FF6F00')
plt.barh(x1,y2,label = "Closed Tickets", left = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + id="5zYAZrrvlqKT" colab={"base_uri": "https://localhost:8080/", "height": 354} outputId="658c0492-a098-42c1-e0ec-b570ef179fcb"
#display text in horizontal bar chart
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(8,5))
plt.barh(x1,y1,label = "Open Tickets",color = '#FF6F00')
plt.barh(x1,y2,label = "Closed Tickets", left = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
for x,y in zip(x1,y1):
plt.text(y-10, x, '%d' % y, ha='center' , va= 'bottom')
for x,y,z in zip(x1,y2,y1):
plt.text(y+z-10, x, '%d' % y, ha='center' , va= 'bottom')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# + [markdown] id="qV8hlG-LnFRF"
# ###Seaborn barchart
# + id="xUQDBJUJRyGN" colab={"base_uri": "https://localhost:8080/", "height": 262} outputId="845c45c2-5cd4-4d4e-a826-0e17e74aa516"
x1=[1,2,3,4,5,6,7]
y1=[10,20,30,40,50,60,70]
sns.barplot(x=x1,y=y1)
plt.show()
# + id="SO4i-wn6UAKn" colab={"base_uri": "https://localhost:8080/", "height": 262} outputId="dee72809-353a-44a5-97de-df949c983bd6"
x1=[1,2,3,3,3,4,5]
y1=[10,20,30,40,50,60,70]
sns.barplot(x=x1,y=y1, ci=None)
plt.show()
# + id="tnxgnckjURHJ" colab={"base_uri": "https://localhost:8080/", "height": 120} outputId="f2c1727d-5c54-4f4b-ff88-7789137633dd"
from google.colab import drive
drive.mount('/content/drive')
# + id="oYRV9tqLVr5w" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="b40fec43-3b81-42ab-b1ab-180eb99cba5b"
pokemon = pd.read_csv('/content/drive/My Drive/Data Science/pokemon_updated.csv')
pokemon.head()
# + id="5-bjRY4YV9J7" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="6ac9be2d-7c20-4998-8154-8cc795c50105"
plt.figure(figsize=(20,7))
sns.barplot(x=pokemon['Type 1'], y = pokemon['Defense'])
plt.show()
# + id="OIgQn5J4WMlc" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="5d23a7cc-d015-4a42-b23a-086e8053177b"
plt.figure(figsize=(20,7))
order = pokemon.groupby(['Type 1']).mean().sort_values('Defense', ascending= False).index.values
sns.barplot(x=pokemon['Type 1'], y = pokemon['Defense'], order = order)
plt.show()
# + id="05jjwyPGW0eG" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="1a605eca-07e0-48b4-fa2c-b9eb1f2a5d9e"
order
# + id="R-Y2iOcnW1vw" colab={"base_uri": "https://localhost:8080/", "height": 602} outputId="a6aa10ae-043d-4687-f928-f37798817fd9"
plt.figure(figsize=(8,10))
sns.barplot(x=pokemon['Speed'], y = pokemon['Type 1'])
plt.show()
# + id="6ZzXYr5EX15f" colab={"base_uri": "https://localhost:8080/", "height": 292} outputId="0ef3ce88-635d-4714-ce82-0fc8a9e55c3f"
sns.barplot(x=pokemon['Defense'], y = pokemon['Type 1'])
# + id="QYJGt0g7X_r4" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="936ba68f-7fbf-4673-8959-33eae6f7ab36"
plt.figure(figsize=(20,7))
sns.countplot(x=pokemon['Type 1'])
plt.show()
# + id="eG_Rkn-JYrtF" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="8279f808-06f6-4b5b-f483-a02f1ca7d548"
plt.figure(figsize=(20,7))
sns.barplot(x=pokemon['Type 1'], y = pokemon['Defense'], hue = pokemon['Legendary'])
sns.despine() #right and top lines can be removed
plt.show()
# + id="fenk2oTbZgr7" colab={"base_uri": "https://localhost:8080/", "height": 452} outputId="9518cc95-51b5-42a2-c8b1-1edf07d816e8"
plt.figure(figsize=(20,7))
sns.set(rc={"axes.facecolor": "#283747", "axes.grid": False, 'xtick.labelsize':14, 'ytick.labelsize':14})
sns.barplot(x=pokemon['Type 1'], y = pokemon['Defense'])
plt.show()
# + [markdown] id="M_Rvre6Am5XZ"
# ###Scatter plot
# + id="H7xc4Ko5aqAr" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="676489f6-b786-48a2-eeff-c79673ab09e7"
x1 = np.array([250,150,350,252,450,550,455,358,158,355])
y1 =np.array([40,50,80, 90, 100,50,60,88,54,45])
x2 = np.array([200,100,300,220,400,500,450,380,180,350])
y2 = np.array([400,500,800, 900, 1000,500,600,808,504,405])
plt.scatter(x1,y1)
plt.xlabel("$Time $ $Spent$",fontsize=12)
plt.ylabel("$Score$",fontsize=12)
plt.title("Scatter Graph")
plt.show()
# + id="rlI90GcidzDH" colab={"base_uri": "https://localhost:8080/", "height": 309} outputId="0098e6e6-386a-4343-ef9c-034ea3ac1f8e"
plt.scatter(x1,y1,color='r')
plt.xlabel("$Time $ $Spent$",fontsize=12)
plt.ylabel("$Score$",fontsize=12)
plt.title("Scatter Graph")
plt.show()
# + id="qmSjxVQWd4qE" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="c0a4142f-6468-4e30-e918-5a99be7c0e4f"
plt.scatter(x1,y1,label = 'Class 1')
plt.scatter(x2,y2,label='Class 2', color='r')
plt.xlabel("$Time $ $Spent$",fontsize=12)
plt.ylabel("$Score$",fontsize=12)
plt.title("Scatter Graph")
plt.legend()
plt.show()
# + id="rCgeKcLKe7ME" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="5b6f0e11-f975-48cd-e8c2-434541853685"
plt.scatter(x1,y1,label = 'Class 1',marker='+', color='b', s=50, alpha = 0.6)
plt.scatter(x2,y2,label='Class 2', marker= 'v',color='r', s=100, alpha = 1)
plt.xlabel("$Time $ $Spent$",fontsize=12)
plt.ylabel("$Score$",fontsize=12)
plt.title("Scatter Graph")
plt.legend(bbox_to_anchor=(1.0,1.0), shadow = True, fontsize="x-large")
plt.show()
# + id="V75NUdX7fxVn" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="8cc356be-71d8-4410-9ea2-6f02a3fef274"
ins = pd.read_csv('/content/drive/My Drive/Data Science/insurance.csv')
ins.head()
# + id="WS4U8Fo5jg3D" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="097ad5a9-ed1f-4edf-86cc-a6d297e0a220"
plt.figure(figsize=(7,7))
sns.scatterplot(x=ins.bmi,y=ins.charges,color='g',marker='o')
plt.show()
# + id="UEnaIrYsjtSm" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="f5ccd714-6492-4ed9-908d-5cb47b78b101"
plt.figure(figsize=(7,7))
sns.scatterplot(x=ins.bmi,y=ins.charges,color='g',hue=ins.sex)
plt.show()
# + id="UQezqUk3kRVe" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="9b7efed2-6234-438f-adc5-9ce415be9c66"
plt.figure(figsize=(7,7))
sns.scatterplot(x=ins.bmi,y=ins.charges,hue=ins.smoker)
plt.show()
# + id="AtZCdkMpk28s" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="952ee379-c3b2-41b3-af08-51f2f311736a"
plt.figure(figsize=(7,7))
plt.xlim([0,50])
sns.scatterplot(x=ins.bmi,y=ins.charges,hue=ins.smoker)
plt.show()
# + id="GDHmGIgslofS" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="c6469b69-fb99-4a89-8e5b-4cff562d7dc7"
plt.figure(figsize=(7,7))
plt.xlim([0,50])
sns.scatterplot(x=ins.bmi,y=ins.charges,hue=ins.smoker, size=ins.children)
plt.show()
# + [markdown] id="q9huxNBEs_0b"
# ###Quiz
# + id="P4TvQXQnmO3K" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="0a44314d-b8aa-4a76-d04f-42e1b93cb5f6"
df1 = pd.read_csv('https://raw.githubusercontent.com/shanealynn/Pandas-Merge-Tutorial/master/user_usage.csv')
df1.head()
# + id="oBns-F8PosZK" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="d9ac609a-1dc0-47bd-c651-0e63256ed271"
df2 = pd.read_csv('https://raw.githubusercontent.com/shanealynn/Pandas-Merge-Tutorial/master/user_device.csv')
df2.head()
# + id="TNbjKXDFoxoZ" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="d3975b12-262a-4f5f-9a39-cec4ea6627f5"
df3 = pd.read_csv('https://raw.githubusercontent.com/shanealynn/Pandas-Merge-Tutorial/master/android_devices.csv')
df3.head()
# + id="S--pacXCo3mD"
df4 = pd.merge(df1,df2,on="use_id")
# + id="UylEAhZ7pWO9" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="18ab1fc2-0472-44d7-afd0-718b57b78714"
df4.head()
# + id="WgSBB3qKpXiA" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="40168af9-1aeb-4721-fe16-cb98ae777c3a"
df4.groupby('platform').get_group('android')['outgoing_mins_per_month'].sum()
# + id="3KjM3nWJpg2a" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="434d4e59-2be7-4f24-b63a-962e6cae5dfe"
df4.groupby('device').get_group('SM-G930F')['outgoing_sms_per_month'].sum()
# + id="ijZ8uC4UqYUC"
data1 = {'key': ['K0', 'K1', 'K2', 'K3'],
'key1': ['K0', 'K1', 'K0', 'K1'],
'Name':['Jai', 'Princi', 'Gaurav', 'Anuj'],
'Age':[27, 24, 22, 32],}
data2 = {'key': ['K0', 'K1', 'K2', 'K3'],
'key1': ['K0', 'K0', 'K0', 'K0'],
'Address':['Nagpur', 'Kanpur', 'Allahabad', 'Kannuaj'],
'Qualification':['Btech', 'B.A', 'Bcom', 'B.hons']}
# + id="GMByuD-Yq8R_"
df=pd.DataFrame.from_dict(data1)
# + id="m7SzJAyeq9qE"
df1=pd.DataFrame.from_dict(data2)
# + id="d6W-THgUq-To" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="ca0d0d3b-f0b6-4294-92bc-84a40d7610a2"
df.head()
# + id="N80TSnLjrcdT" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="7220cd67-a2d3-4f2c-d5bd-25a6bdef807e"
df1.head()
# + id="CiSoNGeordG9" colab={"base_uri": "https://localhost:8080/", "height": 231} outputId="1bfe4da3-61d5-4376-aeb6-a2ade3dabd2d"
pd.merge(df,df1,how="outer",on=['key','key1'])
# + id="UrcNFKO2rumQ"
one = pd.DataFrame({
'Name': ['Alex', 'Amy', 'Allen', 'Alice', 'Ayoung'],
'subject_id':['sub1','sub2','sub4','sub6','sub5'],
'Marks_scored':[98,90,87,69,78]},
index=[1,2,3,4,5])
two = pd.DataFrame({
'Name': ['Billy', 'Brian', 'Bran', 'Bryce', 'Betty'],
'subject_id':['sub2','sub4','sub3','sub6','sub5'],
'Marks_scored':[89,80,79,97,88]},
index=[1,2,3,4,5])
# + id="eyBfUZVxsLkb" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="722efb46-6ad0-476f-e292-3b594acfe767"
pd.concat([one,two])
# + id="PyhYwyjEsPcp" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="628f18d6-db58-4bc0-84cf-eeb992945ad8"
pd.merge(one,two,how='outer')
# + id="WcZT-InrsdVZ"
| plotting_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
from intern.remote.boss import BossRemote
import numpy as np
import time
import pandas as pd
boss = BossRemote({
"protocol": "http",
"host": "localhost:5000",
"token": "<PASSWORD>"
})
# +
def generate_random_data(shape):
my_data = np.random.randint(0, 100, shape, dtype="uint8")
for i in range(my_data.shape[0]):
my_data[i] += int(i/4)
for j in range(my_data.shape[2]):
my_data[:,:,j] += int(j/4)
return my_data
def run_upload(data):
boss.create_cutout(
boss.get_channel("my_channel", "my_collection", "my_experiment"),
0,
[0, data.shape[0]],
[0, data.shape[1]],
[0, data.shape[2]],
data
)
def run_download(shape):
new_data = boss.get_cutout(
boss.get_channel("my_channel", "my_collection", "my_experiment"),
0,
[0, shape[0]],
[0, shape[1]],
[0, shape[2]]
)
# -
sizes_to_test = [
20,
50,
75,
100,
200,
250,
300,
250,
400,
500,
800,
]
upload_times = []
download_times = []
for size in sizes_to_test:
for i in range(3):
shape = (size, size, size)
data = generate_random_data(shape)
tic = time.time()
run_upload(data)
upload_times.append({
"size": size,
"time": time.time() - tic,
"attempt": i
})
tic = time.time()
run_download(shape)
download_times.append({
"size": size,
"time": time.time() - tic,
"attempt": i
})
upload_df = pd.DataFrame(upload_times)
download_df = pd.DataFrame(download_times)
# +
plt.title("Upload Times")
plt.scatter(upload_df['size'], upload_df.time)
plt.ylabel("Time (s)")
plt.xlabel("Size (edge-length)")
plt.show()
plt.title("Download Times")
plt.scatter(download_df['size'], download_df.time)
plt.ylabel("Time (s)")
plt.xlabel("Size (edge-length)")
plt.show()
# -
| notebooks/Bossphorus Benchmarks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using deep features to train an image classifier
import turicreate
# # Load some data
image_train = turicreate.SFrame('../../data/image_train_data/')
image_test = turicreate.SFrame('../../data/image_test_data/')
# # Explore this image data
image_train
# # Train an image classifier on raw image pixels
raw_pixel_model = turicreate.logistic_classifier.create(image_train,
target = 'label',
features = ['image_array'])
# # Make predictions using simple raw pixel model
image_test[0:3]['image'].explore()
image_test[0:3]['label']
raw_pixel_model.predict(image_test[0:3])
# # Evaluate the raw pixel model on the test data
raw_pixel_model.evaluate(image_test)
# # Train image classifier using deep features
len(image_train)
image_train
# # Given the deep features, train a logistic classifier
deep_features_model = turicreate.logistic_classifier.create(image_train,
target='label',
features = ['deep_features'])
# # Apply the deep features classifier on the first few images
image_test[0:3]['image'].explore()
deep_features_model.predict(image_test[0:3])
# # Quantitatively evaluate deep features classifier on test data
deep_features_model.evaluate(image_test)
| course-exercises/week6-Neural-Networks/Class-examples-deep-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Training MFA on fuaimeanna.ie
#
# > "Training Monreal Forced Aligner using fuaimeanna.ie data on Kaggle"
#
# - toc: false
# - branch: master
# - badges: false
# - comments: true
# - hidden: true
# - categories: [kaggle, mfa, fuaimeanna]
# [Original](https://www.kaggle.com/jimregan/train-irish-mfa-model-fuaimeanna)
| _drafts/2021-08-25-mfa-fionn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/TwistedAlex/TensorFlow2Practice/blob/main/EAGER_EXE_TUTORIAL.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EBgHaCxIeKX-"
# **Eager execution**
# Evaluate operations immediately, without building graphs: operations *return concrete values* instead of constructing a compuational graph to run later.
# *An intuitive interface*: Structure your code naturally and use Python data structures. Quickly iterate on small models and small data.
# *Easier debugging*: Call ops directly to inspect running models and test changes. Use standard Python debugging toools for immediate error report.
# *Natural control flow*: Use Python control flow instead of graph control flow, simplifying the specification of dynamic models.
# Ref: https://www.tensorflow.org/guide/eager
# + [markdown] id="-jadZi_NfET-"
# **Setup and basic usage**
# + id="iYSr91h9fFHE"
import os
import tensorflow as tf
import cProfile
tf.compat.v1.enable_eager_execution()
# + colab={"base_uri": "https://localhost:8080/"} id="3XAQxdWLjWDL" outputId="18a330e6-e731-4475-cad0-f8af79f29741"
# Eager execution is enabled by default in TF2.0
tf.executing_eagerly()
# Run TF ops and the results will return immediately:
x = [[2.]]
m = tf.matmul(x, x)
print("hello, {}".format(m))
a = tf.constant([[1, 2], [3, 4]])
print(a)
# + [markdown] id="f1RIAi5p95Pp"
# Broadcasting
# + colab={"base_uri": "https://localhost:8080/"} id="noyh6Fax96ei" outputId="43d504dc-a3a3-43f3-ff69-e54c78977f3b"
b = tf.add(a, 1)
print(b)
# + [markdown] id="SVmLsK7_963i"
# Operator overloading
# + colab={"base_uri": "https://localhost:8080/"} id="CqnMdIA6-izf" outputId="b704a6e8-4a90-49d3-e681-2853c80837f3"
print(a * b)
# + [markdown] id="q27eIQ5k-thE"
# Use NumPy values
# + colab={"base_uri": "https://localhost:8080/"} id="7dw1DZfE-uVY" outputId="33e2732c-b4c7-4ea5-9dbb-d08ab95de365"
import numpy as np
c = np.multiply(a, b)
print(c)
# + [markdown] id="7SyjI92w_Boh"
# Obtain numpy value from a tensor:
# + colab={"base_uri": "https://localhost:8080/"} id="sDGadXb3_ErT" outputId="c92b43f9-db41-4b32-ec92-df373c551222"
print(a.numpy())
# + [markdown] id="5Je2GJFr_V0t"
# **Dynamic control flow**
#
# A major benefit of eager execution is that all the functionality of the host language is available while your model is executing. So, for example, it is easy to write fizzbuzz:
# + id="-T2gO-_8_53C"
def fizzbuzz(max_num):
counter = tf.constant(0)
max_num = tf.convert_to_tensor(max_num)
for num in range(1, max_num.numpy()+1):
num = tf.constant(num)
if int(num % 3) == 0 and int(num % 5) == 0:
print('FizzBuzz')
elif int(num % 3) == 0:
print('Fizz')
elif int(num % 5) == 0:
print('Buzz')
else:
print(num.numpy())
counter += 1
# + [markdown] id="Rr8OK7BAABRQ"
# This has conditionals that depend on tensor values and it prints these values at runtime.
# + colab={"base_uri": "https://localhost:8080/"} id="zGxXjvcx_88m" outputId="718cc0e8-81b7-431a-cf7e-5c8dab652aea"
fizzbuzz(15)
# + [markdown] id="MBKkprX7FiQI"
# **Eager training**
#
# Computing gradients
#
# **Automatic differentiation** is useful for implementing machine learning algorithms such as backpropagation for training neural networks. During eager execution, use tf.GradientTape to trace operations for computing gradients later.
#
# You can use [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) to train and/or compute gradients in eager. It is especially useful for complicated training loops.
#
#
# Args
#
# persistent: Boolean controlling whether a persistent gradient tape is created. False by default, which means at most one call can be made to the gradient() method on this object.
#
# watch_accessed_variables: Boolean controlling whether the tape will automatically watch any (trainable) variables accessed while the tape is active. Defaults to True meaning gradients can be requested from any result computed in the tape derived from reading a trainable Variable. If False users must explicitly watch any Variables they want to request gradients from.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="H_KPSMOkanMt" outputId="61689924-2eb5-4d64-cf9b-0c3d3b5b68df"
x = tf.constant(3.0)
with tf.GradientTape() as g:
g.watch(x)
y = x * x
dy_dx = g.gradient(y, x)
print(dy_dx)
# + [markdown] id="gOUkEO3V0KUq"
# **Train a model**
#
# The following example creates a **multi-layer model** that **classifies** the standard MNIST handwritten digits. It demonstrates the optimizer and layer APIs to build trainable graphs in an eager execution environment.
#
# Trainable variables (created by tf.Variable or tf.compat.v1.get_variable, where trainable=True is default in both cases) are **automatically watched**. Tensors can be manually watched by invoking the watch method on this context manager.
# + colab={"base_uri": "https://localhost:8080/"} id="5yUA8EbJFkAr" outputId="91fbb7aa-1260-4692-aa72-4ed9648ea42f"
# Fetch and format the mnist data
(mnist_images, mnist_labels), _ = tf.keras.datasets.mnist.load_data()
# Normalize mnist images pixel value
dataset = tf.data.Dataset.from_tensor_slices(
(tf.cast(mnist_images[...,tf.newaxis]/255, tf.float32),
tf.cast(mnist_labels,tf.int64)))
dataset = dataset.shuffle(1000).batch(32)
# + id="VwMsq1bn0h5V"
# Build the model
mnist_model = tf.keras.Sequential([
tf.keras.layers.Conv2D(16,[3,3], activation='relu',
input_shape=(None, None, 1)),
tf.keras.layers.Conv2D(16,[3,3], activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(10)
])
# + [markdown] id="zSF2Ir540tIK"
# Even without training, call the model and inspect the output in eager execution:
# + colab={"base_uri": "https://localhost:8080/"} id="pLNNRTx10t24" outputId="d49c3d78-3a69-47b7-d815-c8bba1d57602"
for images,labels in dataset.take(1):
print("Logits: ", mnist_model(images[0:1]).numpy())
# + [markdown] id="nhSaQt2bz6yg"
# While keras models have a builtin training loop (using the **fit**(next example) method), sometimes you need more **customization**(below). Here's an example, of a training loop implemented with eager:
# + id="io1QBgSI1NN2"
optimizer = tf.keras.optimizers.Adam()
# The loss function to be optimized
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
loss_history = []
# + id="ot3iUhnU1RRF"
def train_step(images, labels):
with tf.GradientTape() as tape:
logits = mnist_model(images, training=True)
# Add asserts to check the shape of the output.
tf.debugging.assert_equal(logits.shape, (32, 10))
loss_value = loss_object(labels, logits)
loss_history.append(loss_value.numpy().mean())
# grads = d(loss)/d(param)
grads = tape.gradient(loss_value, mnist_model.trainable_variables)
optimizer.apply_gradients(zip(grads, mnist_model.trainable_variables))
# + id="5Ots2NuJ1XUD"
def train(epochs):
for epoch in range(epochs):
for (batch, (images, labels)) in enumerate(dataset):
train_step(images, labels)
print ('Epoch {} finished'.format(epoch))
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="Lk0-Uys11ZOK" outputId="8b40837a-807a-463a-ddd0-c6316c5b5eec"
train(epochs = 5)
# + colab={"background_save": true} id="iXLvGY5P1eFH" outputId="a0038635-d34a-4ae9-9518-35d5dc3be3db"
import matplotlib.pyplot as plt
plt.plot(loss_history)
plt.xlabel('Batch #')
plt.ylabel('Loss [entropy]')
# + [markdown] id="U31POQY4JuGH"
# **Variables and optimizers**
# [tf.Variable](https://www.tensorflow.org/api_docs/python/tf/Variable) objects store mutable [tf.Tensor](https://www.tensorflow.org/api_docs/python/tf/Tensor)-like values accessed during training to make automatic differentiation easier.
#
# The collections of variables can be encapsulated into layers or models, along with methods that operate on them. See Custom Keras layers and models for details. The main difference between layers and models is that models add methods like Model.fit, Model.evaluate, and Model.save.
#
# For example, the automatic differentiation example above can be rewritten:
# + id="B3Iw48WkdOyG"
# Model class
class Linear(tf.keras.Model):
def __init__(self):
super(Linear, self).__init__()
self.W = tf.Variable(5., name='weight')
self.B = tf.Variable(10., name='bias')
def call(self, inputs):
return inputs * self.W + self.B
# + id="-rnMxOUFdOgu"
# A toy dataset of points around 3 * x + 2
NUM_EXAMPLES = 2000
training_inputs = tf.random.normal([NUM_EXAMPLES])
noise = tf.random.normal([NUM_EXAMPLES])
training_outputs = training_inputs * 3 + 2 + noise
# The loss function to be optimized
def loss(model, inputs, targets):
error = model(inputs) - targets
return tf.reduce_mean(tf.square(error))
def grad(model, inputs, targets):
with tf.GradientTape() as tape:
loss_value = loss(model, inputs, targets)
return tape.gradient(loss_value, [model.W, model.B])
# + [markdown] id="xK2OIj7Enam6"
#
#
# 1. Create the model.
# 2. The Derivatives of a loss function with respect to model parameters.
# 3. A strategy for updating the variables based on the derivatives.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="mfdMMUp4dOO6" outputId="25e3c80e-2e86-46f3-f7e0-70c55a2f3c97"
model = Linear()
optimizer = tf.keras.optimizers.SGD(learning_rate=0.01)
print("Initial loss: {:.3f}".format(loss(model, training_inputs, training_outputs)))
steps = 300
for i in range(steps):
grads = grad(model, training_inputs, training_outputs)
optimizer.apply_gradients(zip(grads, [model.W, model.B]))
if i % 20 == 0:
print("Loss at step {:03d}: {:.3f}".format(i, loss(model, training_inputs, training_outputs)))
# + colab={"base_uri": "https://localhost:8080/"} id="bEkXF1cGn1k-" outputId="7265915e-f0a9-471a-ed21-ea792053c2d3"
print("Final loss: {:.3f}".format(loss(model, training_inputs, training_outputs)))
# + colab={"base_uri": "https://localhost:8080/"} id="2ooiw82In3RL" outputId="64f7862a-602c-4b75-b869-8f4e9718a0b8"
print("W = {}, B = {}".format(model.W.numpy(), model.B.numpy()))
# + colab={"base_uri": "https://localhost:8080/"} id="05mRnwlyoLKK" outputId="cfa6e937-a220-43e4-96e6-4b79043cc052"
print(model.call(3))
# + [markdown] id="zi0DjzgQomDS"
# **Object-based saving**
#
# A tf.keras.Model includes a convenient save_weights method allowing you to easily create a checkpoint:
# + id="Frqml4NaopEk"
model.save_weights('weights')
status = model.load_weights('weights')
# + [markdown] id="aE6vB5Z6o038"
# Using [tf.train.Checkpoint](https://www.tensorflow.org/api_docs/python/tf/train/Checkpoint) you can take full control over this process.
#
# This section is an abbreviated version of the guide to training [checkpoints](https://www.tensorflow.org/guide/checkpoint).
# + id="mvuY6Uq8pt5Y"
x = tf.Variable(10.)
checkpoint = tf.train.Checkpoint(x=x)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="6xdttHVUpuXH" outputId="2f87901f-76f9-45d2-cbe0-8fbd54562c23"
x.assign(2.) # Assign a new value to the variables and save.
checkpoint_path = './ckpt/'
checkpoint.save('./ckpt/')
# + colab={"base_uri": "https://localhost:8080/"} id="1XFF2ZYFp9N7" outputId="51ad0142-a459-41fc-8c8f-a071b32e60f0"
x.assign(11.) # Change the variable after saving.
# Restore values from the checkpoint
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_path))
print(x) # => 2.0
# + [markdown] id="LcSsk5w6qBgL"
# To save and load models, tf.train.Checkpoint stores the internal state of objects, without requiring hidden variables. To record the state of a model, an optimizer, and a global step, pass them to a tf.train.Checkpoint:
# + colab={"base_uri": "https://localhost:8080/"} id="SdGPldqEqDr3" outputId="fdc9123c-3be9-4ff9-fa1f-4bd67c403c82"
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(16,[3,3], activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(10)
])
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
checkpoint_dir = 'path/to/model_dir'
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
root = tf.train.Checkpoint(optimizer=optimizer,
model=model)
root.save(checkpoint_prefix)
root.restore(tf.train.latest_checkpoint(checkpoint_dir))
# + [markdown] id="sXxO5ZbTqKN4"
# **Object-oriented metrics**
#
# tf.keras.metrics are stored as objects. Update a metric by passing the new data to the callable, and retrieve the result using the tf.keras.metrics.result method, for example:
# + colab={"base_uri": "https://localhost:8080/"} id="Jav1ZfD_qTGS" outputId="742c9d37-0e06-40eb-baac-cf6d8bebf74d"
m = tf.keras.metrics.Mean("loss")
m(0)
m(5)
m.result() # => 2.5
m([8, 9])
m.result() # => 5.5
# + [markdown] id="EyTVG3cgqgiY"
# **Summaries and TensorBoard**
#
# [TensorBoard](https://www.tensorflow.org/tensorboard) is a visualization tool for understanding, debugging and optimizing the model training process. It uses summary events that are written while executing the program.
#
# You can use [tf.summary](https://www.tensorflow.org/api_docs/python/tf/summary) to record summaries of variable in eager execution. For example, to record summaries of loss once every 100 training steps:
# + id="DTTO005RrBnA"
logdir = "./tb/"
writer = tf.summary.create_file_writer(logdir)
steps = 1000
with writer.as_default(): # or call writer.set_as_default() before the loop.
for i in range(steps):
step = i + 1
# Calculate loss with your real train function.
loss = 1 - 0.001 * step
if step % 100 == 0:
tf.summary.scalar('loss', loss, step=step)
# + [markdown] id="AhRPmfy8raAx"
# **Advanced automatic differentiation topics**
#
# Dynamic models
#
# tf.GradientTape can also be used in dynamic models. This example for a backtracking line search algorithm looks like normal NumPy code, except there are gradients and is differentiable, despite the complex control flow:
# + id="HRhMCohwr0hU"
def line_search_step(fn, init_x, rate=1.0):
with tf.GradientTape() as tape:
# Variables are automatically tracked.
# But to calculate a gradient from a tensor, you must `watch` it.
tape.watch(init_x)
value = fn(init_x)
grad = tape.gradient(value, init_x)
grad_norm = tf.reduce_sum(grad * grad)
init_value = value
while value > init_value - rate * grad_norm:
x = init_x - rate * grad
value = fn(x)
rate /= 2.0
return x, value
# + [markdown] id="pvUqFy4IvBh7"
# Custom gradients
#
# Custom gradients are an easy way to override gradients. Within the forward function, define the gradient with respect to the inputs, outputs, or intermediate results. For example, here's an easy way to clip the norm of the gradients in the backward pass:
#
# #TODO: Custom Gradients Examples
# + id="iHtVuwcovDe-"
@tf.custom_gradient
def clip_gradient_by_norm(x, norm):
y = tf.identity(x)
def grad_fn(dresult):
return [tf.clip_by_norm(dresult, norm), None]
return y, grad_fn
# + [markdown] id="jISxdNnYvPzu"
# Custom gradients are commonly used to provide a numerically stable gradient for a sequence of operations:
# + id="uokAg67IvO3n"
def log1pexp(x):
return tf.math.log(1 + tf.exp(x))
def grad_log1pexp(x):
with tf.GradientTape() as tape:
tape.watch(x)
value = log1pexp(x)
# Grad = d(value)/d(x)
return tape.gradient(value, x)
# + colab={"base_uri": "https://localhost:8080/"} id="1vwhElEDvVcq" outputId="f4bf28af-dccc-4809-a6fd-8072d6434683"
# The gradient computation works fine at x = 0.
grad_log1pexp(tf.constant(0.)).numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="mKkki0qfvWvI" outputId="2cad7fd1-58c9-42d9-fc31-f95013006636"
# However, x = 100 fails because of numerical instability.
grad_log1pexp(tf.constant(100.)).numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="V7QNR7E5945i" outputId="bd4936fa-2efe-4b09-ee3d-33c0e3f213ff"
print(tf.exp(100.).numpy())
# + [markdown] id="ZGDWrhL5vYg6"
# Here, the log1pexp function can be analytically simplified with a custom gradient. The implementation below reuses the value for tf.exp(x) that is computed during the forward pass—making it more efficient by eliminating redundant calculations:
# + id="6tGKyC5WvZyf"
@tf.custom_gradient
def log1pexp(x):
e = tf.exp(x)
def grad(dy):
return dy * (1 - 1 / (1 + e))
return tf.math.log(1 + e), grad
def grad_log1pexp(x):
with tf.GradientTape() as tape:
tape.watch(x)
value = log1pexp(x)
return tape.gradient(value, x)
# + id="nqCeVgCMva9w"
# As before, the gradient computation works fine at x = 0.
grad_log1pexp(tf.constant(0.)).numpy()
# + id="ptWxUfNxvcHP"
# And the gradient computation also works at x = 100.
grad_log1pexp(tf.constant(100.)).numpy()
# + [markdown] id="zgDQPRb5vU1o"
# **Performance**
#
# Computation is automatically offloaded to GPUs during eager execution. If you want control over where a computation runs you can enclose it in a tf.device('/gpu:0') block (or the CPU equivalent):
# + colab={"base_uri": "https://localhost:8080/"} id="6CkDCIEA_KLU" outputId="5a5b7ce0-9d10-4520-c9ba-c91cf0c62930"
import time
def measure(x, steps):
# TensorFlow initializes a GPU the first time it's used, exclude from timing.
tf.matmul(x, x)
start = time.time()
for i in range(steps):
x = tf.matmul(x, x)
# tf.matmul can return before completing the matrix multiplication
# (e.g., can return after enqueing the operation on a CUDA stream).
# The x.numpy() call below will ensure that all enqueued operations
# have completed (and will also copy the result to host memory,
# so we're including a little more than just the matmul operation
# time).
_ = x.numpy()
end = time.time()
return end - start
shape = (1000, 1000)
steps = 200
print("Time to multiply a {} matrix by itself {} times:".format(shape, steps))
# Run on CPU:
with tf.device("/cpu:0"):
print("CPU: {} secs".format(measure(tf.random.normal(shape), steps)))
# Run on GPU, if available:
if tf.config.experimental.list_physical_devices("GPU"):
with tf.device("/gpu:0"):
print("GPU: {} secs".format(measure(tf.random.normal(shape), steps)))
else:
print("GPU: not found")
# + [markdown] id="LGSr_DfZ_ex7"
# A tf.Tensor object can be copied to a different device to execute its operations:
#
# #TODO CHECK AVAILABILITY IN CURRENT VERSION
# + id="aCyXZe_N_fuT"
if tf.config.experimental.list_physical_devices("GPU"):
x = tf.random.normal([10, 10])
x_gpu0 = x.gpu()
x_cpu = x.cpu()
_ = tf.matmul(x_cpu, x_cpu) # Runs on CPU
_ = tf.matmul(x_gpu0, x_gpu0) # Runs on GPU:0
# + [markdown] id="hI2adrV0mn7Y"
# Disable eager execution
# + colab={"base_uri": "https://localhost:8080/"} id="vFZvyYaolywK" outputId="a27e995f-0a21-4e9a-b3a4-8ffca0d2e4f2"
tf.compat.v1.disable_eager_execution()
a = tf.constant([[1, 2], [3, 4]])
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="ghyt0LWxl1bY" outputId="c223e361-78ab-4875-b50a-aaae388b1dd7"
b = tf.add(a, 1)
print(b)
# + id="i_OVp02V-xi5"
import numpy as np
c = np.multiply(a, b)
print(c)
| EAGER_EXE_TUTORIAL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## homework 5
# 1) repeat exercise from the 11/4 session, but use the Cash-Karp Runge Kutta Method with adaptive stepwise control.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# 2) evolve the system of equations
# - (dy/dx) = z
# - (dz/dx) = -y
# use intital conditions y(x=0)=0 and dy/dx(x=0) = 1, <br><br>
# evolve over the range [0, 2*pi].
def dydx(x,y):
#set the derivatives
#our equation is d^2y/dx^2 = -y
#so, we can write
#dydx = z
#dz/dx = -y
#we will set y = y[0]
#and we will set z = y[1]
#declare an array
y_derivs = np.zeros(2)
#set dydx = z
y_derivs[0] = y[1]
#set dydx = -y
y_derivs[1] = -1*y[0]
#here we have to retirn an array
return y_derivs
def rk6_mv_core(dydx, xi, yi, nv, h):
#dfdx is the function of the derivatives
#xi is the valye of z at step 1
#yi is the array of variables at step i
#nv is the number of variables
#h is the step size
#declare k arrays
k1 = np.zeros(nv)
k2 = np.zeros(nv)
k3 = np.zeros(nv)
k4 = np.zeros(nv)
k5 = np.zeros(nv)
k6 = np.zeros(nv)
#half step
x_ipoh = xi + 0.5*h
#define x at 1 step
x_ipo = xi + h
#declare a temp y array
y_temp = np.zeros(nv)
#get k1 values
y_derivs = dydx(xi, yi)
k1[:] = h*y_derivs[:]
#get k2 values
y_temp[:] = yi[:] + 0.5*k1[:]
y_derivs = dydx(x_ipoh, y_temp)
k2[:] = h*y_derivs[:]
#get k3 values
y_temp[:] = yi[:] + 0.5*k2[:]
y_derivs = dydx(x_ipoh, y_temp)
k3[:] = h*y_derivs[:]
#get k4 values
y_temp[:] = yi[:] + 0.5*k3[:]
y_derivs = dydx(x_ipoh, y_temp)
k4[:] = h*y_derivs[:]
#get k5 values
y_temp[:] = yi[:] + 0.5*k4[:]
y_derivs = dydx(x_ipoh, y_temp)
k5[:] = h*y_derivs[:]
#get k6 values
y_temp[:] = yi[:] + k5[:]
y_derivs = dydx(x_ipo, y_temp)
k6[:] = h*y_derivs[:]
#advance y by a step h
yipo = yi + (k1 + 2*k2 + 2*k3 + 2*k4 + 2*k5 + k6)/6.
return yipo
# +
#this is going to be us defining an adaptive step size driver for RK4
def rk6_mv_ad(dydx, x_i, y_i, nv, h, tol):
#define saftey scale
SAFETY = 0.9
H_NEW_FAC = 2.0
#set a maximum number of iterations
imax = 10000
#set an iteration variable
i = 0
#create an error
Delta = np.full(nv, 2*tol)
#remember the step
h_step = h
#adjust step
while(Delta.max()/tol>1.0):
#estimate our error by
#taking one step of size h
#and compare with
#two steps of size h/2
#one big step
y_2 = rk6_mv_core(dydx, x_i, y_i, nv, h_step)
#two small steps
y_1 = rk6_mv_core(dydx, x_i, y_i, nv, 0.5*h_step)
y_11 = rk6_mv_core(dydx, x_i+0.5*h_step, y_1, nv, 0.5*h_step)
#compute an error
Delta = np.fabs(y_2 - y_11)
#if the error is too large, take a smaller step
if(Delta.max()/tol > 1.0):
#our error is too large
#decrease the step
h_step*= SAFETY * (Delta.max()/tol)**(-0.25)
#check iteration
if(i>=imax):
print("Too many iterations in rk4_mv_ad()")
raise StopIteration("Ending after i =", i)
#iterate
i += 1
#next time, try to take a bigger step
h_new = np.fmin(h_step * (Delta.max()/tol)**(-0.9), h_step*H_NEW_FAC)
#return the answer, a new step, and the step we actually took
return y_2, h_new, h_step
# +
#to wrap or not to wrap
def rk6_mv(dfdx, a, b, y_a, tol):
#dfdx is the derivative wrt x
#a is the lower bound
#b is the upper bound
#y_a are the boundary conditions
#tol is the tolerance for integrating y
#define our starting step
xi = a
yi = y_a.copy()
#an inital step size == make very small
h = 1.0e-4 * (b-a)
#set a maximum number of iterations
imax = 10000
#set an iteration variable
i = 0
#set the number of coupled odes to the size of y_a
nv = len(y_a)
#set the inital conditions
x = np.full(1,a)
y = np.full((1,nv),y_a)
#set a flag
flag = 1
#loop until we reach the right side
while(flag):
#calculate y_i+1 usninf an adaptive step
yi_new, h_new, h_step = rk6_mv_ad(dydx, xi, yi, nv, h, tol)
#update the step
h = h_new
#prevent an overshoot
if(xi+h_step>b):
#take a smaller step
h = b-xi
#recalculate y_i+1
yi_new, h_new, h_step = rk6_mv_ad(dydx, xi, yi, nv, h, tol)
#break
flag = 0
#update the values
xi += h_step
yi[:] = yi_new[:]
#add the step to the arrays
x = np.append(x, xi)
y_new = np.zeros((len(x), nv))
y_new[0:len(x)-1, :] = y
y_new[-1, :] = yi[:]
del y
y = y_new
#prevent too many iterations
if(i>=imax):
print("Maximum iterations reached.")
raise StopIteration("Iteration number = ", i)
#iterate
i += 1
#output some information
s = "i = %3d\tx = %9.8f\th = %9.8f\tb= %9.8f" % (i, xi, h_step, b)
print(s)
#break if new xi is == b
if(xi==b):
flag = 0
#outside of while loop
return x,y
# +
##perform the integration
a = 0.0
b = 2.0*np.pi
y_0 = np.zeros(2)
y_0[0] = 0.0
y_0[1] = 1.0
nv = 2
tolerance = 1.0e-6
#perform the integration
x,y = rk6_mv(dydx, a, b, y_0, tolerance)
# -
# 3) plot the analytical solutions for y(x) and dy/dx(x) over the specified range, and the numerical solution.
# +
#this is currently just a cut and paste from the "runge_kutta_mv"
#might need to edit for it to work in this case
#plotting y first
plt.plot(x,y[:, 0], 'o', label='y(x)')
#plot z second
plt.plot(x,y[:,1], 'o', label='z = dy/dx(x)')
xx = np.linspace(0, 2*np.pi,1000)
plt.plot(xx, np.sin(xx), 'tab:purple', label='sin(x)')
plt.plot(xx, np.cos(xx), 'tab:blue', label='cos(x)')
plt.xlabel('x')
plt.ylabel('y, dy/dx')
plt.legend(frameon=False)
# -
# 4) plot the absolute error for the numerical solutions of y(x) and dy/dx(x) over the specified range.
# +
#this is currently just a cut and paste from the "runge_kutta_mv"
#might need to edit for it to work in this case
sine = np.sin(x)
cosine = np.cos(x)
y_error = (y[:,0]-sine)
dydx_error = (y[:,1]-cosine)
plt.plot(x, y_error, 'tab:pink', label='y(x) Error')
plt.plot(x, dydx_error, 'tab:orange', label = 'dydx(x) Error')
plt.legend(frameon=False)
| astr-119-hw-6/.gitignore/hw_#6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import unittest
def encrypt(text, n):
if n < 1:
return text
for i in range(0, n):
text = text[1::2] + text[::2]
return text
def decrypt(encrypted_text, n):
if n < 1:
return encrypted_text
add = ""
if len(encrypted_text) % 2 == 1:
add = encrypted_text[-1]
for i in range(0, n):
mid = len(encrypted_text) / 2
a = encrypted_text[:mid]
b = encrypted_text[mid:]
# problem is that zip will ignore an unpaired value...
encrypted_text = ''.join([x+y for x, y in zip(b, a)])
return encrypted_text + add
# -
print encrypt("This is a test!", 0) #, "This is a test!")
print encrypt("This is a test!", 1) #, "hsi etTi sats!")
print encrypt("This is a test!", 2) #, "s eT ashi tist!")
print encrypt("This is a test!", 3) #, " Tah itse sits!")
print encrypt("This is a test!", 4) #, "This is a test!")
print encrypt("This is a test!", -1) #, "This is a test!")
a = "This is a test!"[::2]
b = "This is a test!"[1::2]
print b+a
decrypt("hsi etTi sats!", 1)
decrypt("s eT ashi tist!", 2)
# +
# code wars top solution
def decrypt_cw(text, n):
if text in ("", None):
return text
ndx = len(text) // 2 # this is floor division
for i in range(n):
a = text[:ndx]
b = text[ndx:]
text = "".join(b[i:i+1] + a[i:i+1] for i in range(ndx + 1))
return text
def encrypt_cw(text, n):
for i in range(n):
text = text[1::2] + text[::2]
return text
# -
if None in ("", None, "abs"):
print "Something"
ndx = len("this is") // 2
print ndx
| Simple Encryption Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sociation2Vec
# Embedding-и слов аналогичные word2vec из базы ассоциаций [Sociation.org](http://sociation.org)
import os
from operator import itemgetter
import pickle
import pandas as pd
import numpy as np
from tqdm import tqdm_notebook as progress
import gensim
# +
source_dir = "./source_corpus"
data_dir = "/Users/ur001/Documents/Sociation/sociation2vec_model"
random_state = 777
# %pylab inline
# %config InlineBackend.figure_format = 'svg'
# %config InlineBackend.figure_format = 'retina'
# отключаем ворнинги
import warnings
warnings.simplefilter('ignore')
# дефолтный размер графиков
from pylab import rcParams
rcParams['figure.figsize'] = 14,9
# -
# ### Лучший на данный момент вариант
# - corpus
# - normalize_matrix
# - ppmi (0.75, 5.5)
# - tfidf (по словам)
# - svd 790
# - normalize_matrix
# - normalize_l2 (при получении вектора слова)
#
# ```python
# corpus, words_dict, assoc_dict = load_source_corpus()
# corpus_csc = corpus_to_sparse(corpus)
# corpus_csc = normalize_matrix(corpus_csc, with_mean=False)
# corpus_ppmi = get_ppmi_weight(corpus_csc, cds=0.75, k=5.5)
# tfidf, corpus_ppmi = get_tfidf(corpus_ppmi, smooth_idf=False)
# svd, corpus_ppmi_svd = get_svd(corpus_ppmi, 790)
# corpus_ppmi_svd = normalize_matrix(corpus_ppmi_svd)
# similarity_index = create_similarity_index(corpus_ppmi_svd)
# sociation2vec = Sociation2Vec(svd, corpus_ppmi_svd, similarity_index, words_dict, assoc_dict)
# ```
# ## Загрузка словарей и корпуса ассоциаций
# %load_ext autoreload
# %autoreload 2
from sociation2vec.utils import DictEncoder, save_word2vec, read_word2vec
from sociation2vec.model import Sociation2Vec
from sociation2vec import mathutils, model_builder, pandas_utils
corpus, words_dict, assoc_dict = model_builder.load_source_corpus(source_dir)
words_count = len(words_dict.decode)
assoc_count = len(assoc_dict.decode)
print("Слов: {}, Слов-ассоциаций: {}".format(words_count, assoc_count))
pandas_utils.get_associations('система', corpus=corpus, words_dict=words_dict, assoc_dict=assoc_dict).head()
# ## Создание модели вручную
# ### Нормализация, PPMI, TF-IDF
corpus_csc = mathutils.corpus_to_sparse(corpus, words_count, assoc_count)
corpus_csc.shape, (words_count, assoc_count)
corpus_csc = mathutils.normalize_matrix(corpus_csc, with_mean=False)
corpus_ppmi = mathutils.get_ppmi_weight(corpus_csc, cds=0.75, k=5)
corpus_ppmi.shape
tfidf, corpus_ppmi = mathutils.get_tfidf(corpus_ppmi)
corpus_ppmi.shape
pandas_utils.get_associations(
'система',
corpus=mathutils.sparse_to_corpus(corpus_ppmi),
words_dict=words_dict,
assoc_dict=assoc_dict
).head(10)
# ### Факторизация PPMI-матрицы с помощью SVD
# Загрузка корпуса и SVD модели
corpus_ppmi_svd = model_builder.load_corpus('corpus_norm_ppmi_0.75_5.5_tfidf_svd_norm_790', data_dir=data_dir)
svd = model_builder.load_model('model_norm_ppmi_0.75_5.5_tfidf_svd_norm_790', data_dir=data_dir)
# **ИЛИ**
# Создание и сохранение корпуса и SVD модели
svd, corpus_ppmi_svd = mathutils.get_svd(corpus_ppmi, 800)
corpus_ppmi_svd = mathutils.normalize_matrix(corpus_ppmi_svd)
model_builder.save_corpus(corpus_ppmi_svd, 'corpus_norm_ppmi_svd_norm_800', data_dir=data_dir)
model_builder.save_model(svd, 'svd_800_norm_ppmi_model', data_dir=data_dir)
corpus_ppmi_svd.shape
# ### Загрузка или создание/сохранение индекса
similarity_index = model_builder.create_or_load_similarity_index(
corpus_ppmi_svd,
'norm_ppmi_0.75_5.5_tfidf_svd_norm_790',
data_dir=data_dir
)
# ## Смотрим что вышло
sociation2vec = Sociation2Vec(svd, corpus_ppmi_svd, similarity_index, words_dict, assoc_dict)
get_similar = pandas_utils.build_get_similar(sociation2vec)
get_similar('король,-мужчина,женщина')
# ### Recommendations
# http://blog.aylien.com/overview-word-embeddings-history-word2vec-cbow-glove/
#
# - DON’T use shifted PPMI with SVD. (**враньё!**)
# - DON’T use SVD “correctly”, i.e. without eigenvector weighting (performance drops 15 points compared to with eigenvalue weighting with p=0.5). (**не помогло**)
# - DO use PPMI and SVD with short contexts (window size of 2).
# - DO use many negative samples with SGNS.
# - DO always use context distribution smoothing (raise unigram distribution to the power of α=0.75) for all methods.
# - DO use SGNS as a baseline (robust, fast and cheap to train).
# - DO try adding context vectors in SGNS and GloVe.
# ## Автоматизация создания модели
sociation2vec_builder = model_builder.ModelBuilder(source_path=source_dir, out_path=data_dir)
sociation2vec = sociation2vec_builder.get_model(
ppmi_k=5.5,
svd_components=790,
tfidf_smooth_tf=False
)
# +
# sociation2vec.save_word2vec('sociation2vec820.vec')
# -
# python evaluate.py ../sociation2vec820.vec 'russian'
#
# SimLex-999 score and coverage: 0.556 599
# WordSim overall score and coverage: 0.808 305
# WordSim Similarity score and coverage: 0.795 177
# WordSim Relatedness score and coverage: 0.807 220
#
# python evaluate.py ../araneum_upos_skipgram_300_2_2018.vec 'russian'
#
# SimLex-999 score and coverage: 0.398 654
# WordSim overall score and coverage: 0.636 315
# WordSim Similarity score and coverage: 0.722 182
# WordSim Relatedness score and coverage: 0.546 225
get_similar = pandas_utils.build_get_similar(sociation2vec)
compare = pandas_utils.build_compare(sociation2vec)
get_similar('король,-мужчина,женщина')
get_similar('работа,-труд')
get_similar('труд,работа')
compare('работа', 'труд', 8, similarity_degree=0.75, separate=True, min_score=0.45)
compare('мужчина,мужик', 'женщина,девушка', 6, similarity_degree=0.44, separate=True, min_score=0.43)
compare('самец', 'самка', 8, similarity_degree=0.62, separate=True, min_score=0.36)
compare('день,-день недели', 'ночь', 5, similarity_degree=0.63, min_score=0.39)
compare('жизнь', 'смерть', 5, similarity_degree=0.6, min_score=0.35)
compare('мозг', 'сознание', similarity_degree=0.7, min_score=0.5)
compare('любовь', 'секс', 6, similarity_degree=0.7, min_score=0.34)
compare('лист', 'листья', 6, similarity_degree=0.7, min_score=0.3)
compare('сталин', 'гитлер', 9, similarity_degree=0.6, min_score=0.45)
get_similar('король,-брат,сестра', 1)
get_similar('король,-мальчик,девочка', count=1)
get_similar('ассоциация', 15)
get_similar('жизнь', count=3)
get_similar('симбиоз,-паразит', 15)
get_similar('система,-операционная система')
get_similar('ватник,быдло', count=2)
get_similar('ватник,-быдло', count=10)
get_similar('бесконечность')
get_similar('вечность')
get_similar('плотность,объём', count=1)
get_similar('кошка,-женщина,мужчина', count=3)
get_similar('мама,-женщина,мужчина', count=2)
get_similar('папа,-мужчина,женщина', count=2)
get_similar('сын,-мужчина,женщина', count=1)
get_similar('дочь,-женщина,мужчина', count=1)
get_similar('брат,-мальчик,девочка', count=1)
get_similar('принц,-мальчик,девочка', count=1)
get_similar('жена,-девочка,мальчик', count=2)
get_similar('ворон,-мужчина,женщина', count=1)
get_similar('волк,-мужчина,женщина', count=2)
get_similar('собака,-женщина,мужчина', count=1)
get_similar('бог,-христианство,индуизм', count=3)
get_similar('человек,-человечность')
get_similar('человек,человечность')
get_similar('помидоры,-красный цвет,зелёный цвет', count=2)
get_similar('огурцы,-зелёный цвет,красный цвет', count=2)
get_similar('закат,-вечер,утро', count=2)
get_similar('свет,-день,ночь', count=2)
get_similar('похолодание,-осень,весна', count=2)
get_similar('ключ,вода', count=1)
get_similar('ключ,болт', count=1)
get_similar('ключ,криптография', count=1)
get_similar('замок,ключ', count=3)
get_similar('замок,средневековье', count=2)
get_similar('папа,мама', count=4)
get_similar('квадрат', count=7)
get_similar('роберт шекли', count=10)
get_similar('тимур и его команда', count=10)
get_similar('инструмент', count=10)
get_similar('медвежуть', count=10)
get_similar('доброта')
get_similar('безвозмездность', count=4)
get_similar('странность', count=5)
get_similar('женщина,девушка,-мужчина,-мужик', count=2)
get_similar('мужчина,мужик,-женщина,-девушка', count=3)
get_similar('соль,нота', count=5)
get_similar('соль,-нота')
get_similar('рикки-тикки-тави', count=3)
get_similar('фотосессия', count=5)
get_similar('научный сотрудник')
def diminutive(word, count=3):
"""Уменьшительно-ласкательная форма"""
return get_similar(word + ',коробочка,-коробка,пуговка,-пуговица,носик,-нос,пёсик,-пёс,ножка,-нога,речка,-река', count=count)
diminutive('крышка', count=1)
diminutive('медведь', count=1)
diminutive('мышь', count=1)
diminutive('рана', count=1)
diminutive('трусы', count=1)
diminutive('заяц', count=1)
diminutive('ночь', count=3)
get_similar('фифа', count=5)
get_similar('фифа,-футбол', count=5)
get_similar('няша', count=5)
get_similar('инь-ян')
get_similar('мука,-мучения', count=5)
get_similar('мука,-хлеб', count=5)
get_similar('своды')
get_similar('резон')
get_similar('брут')
get_similar('фейсбук')
get_similar('воодушевление')
get_similar('наставник')
get_similar('кот')
get_similar('гекльберри финн')
get_similar('лён', 7)
get_similar('глава,-начальник', 5)
get_similar('глава,-параграф', 5)
get_similar('бревно')
get_similar('опасность')
get_similar('хенд-мейд')
get_similar('дженис джоплин')
get_similar('свежий воздух')
get_similar('клоун')
get_similar('дороговизна')
get_similar('дешёвка')
get_similar('предчувствие')
get_similar('многоугольник')
get_similar('Передача')
get_similar('головная боль', 15)
get_similar('Лазурит')
get_similar('Хардкор')
get_similar('Лаконизм')
get_similar('шарик,-бобик')
get_similar('шарик,-шар')
get_similar('А<NAME>') # педагог
get_similar('Пётр нестеров') # лётчик
get_similar('Коллективный разум', 6)
get_similar('гладь')
get_similar('тишь')
compare('гладь', 'тишь', 25)
get_similar('метафора')
get_similar('Хофман')
get_similar('Три-Д,-трёхмерность')
get_similar('Три-Д,-задний ряд')
get_similar('Почта России')
get_similar('Рэмбо', 5)
get_similar('допинг,стимулятор', 7)
get_similar('Зубчики', 3)
get_similar('Мимолётность')
get_similar('впуск')
get_similar('елена малышева')
get_similar('я-центризм', 20)
get_similar('обязанности', 20)
| svd_ppmi_tfidf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Text classification with Reuters-21578 datasets
# Downloaded the from reuters21578 data set first. https://archive.ics.uci.edu/ml/machine-learning-databases/reuters21578-mld/
#
# * Tested with Python 3.5
# * Keras with Tensorflow 1.3 backend
# %pylab inline
# +
import re
import xml.sax.saxutils as saxutils
from bs4 import BeautifulSoup
from keras.models import Sequential
from keras.layers.embeddings import Embedding
from keras.layers.recurrent import GRU
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from pandas import DataFrame
from random import random
import numpy as np
# Set Numpy random seed
np.random.seed(1)
# -
# ## General constants (modify them according to you environment)
# +
# TODO: download the dataset and change the data_folder to match with yours
# Downloaded from https://archive.ics.uci.edu/ml/machine-learning-databases/reuters21578-mld/
data_folder = 'D:\\Learning\\zoomtail\\reuters21578\\'
sgml_number_of_files = 22
sgml_file_name_template = 'reut2-{}.sgm'
# Category files
category_files = {
'to_': ('Topics', 'all-topics-strings.lc.txt'),
'pl_': ('Places', 'all-places-strings.lc.txt'),
'pe_': ('People', 'all-people-strings.lc.txt'),
'or_': ('Organizations', 'all-orgs-strings.lc.txt'),
'ex_': ('Exchanges', 'all-exchanges-strings.lc.txt')
}
# -
# ## Prepare documents and categories
# +
# Read all categories
category_data = []
for category_prefix in category_files.keys():
with open(data_folder + category_files[category_prefix][1], 'r') as file:
for category in file.readlines():
category_data.append([category_prefix + category.strip().lower(),
category_files[category_prefix][0],
0])
# Create category dataframe
news_categories = DataFrame(data=category_data, columns=['Name', 'Type', 'Newslines'])
# +
def update_frequencies(categories):
for category in categories:
idx = news_categories[news_categories.Name == category].index[0]
f = news_categories.get_value(idx, 'Newslines')
news_categories.set_value(idx, 'Newslines', f+1)
def to_category_vector(categories, target_categories):
vector = zeros(len(target_categories)).astype(float32)
for i in range(len(target_categories)):
if target_categories[i] in categories:
vector[i] = 1.0
return vector
# +
# Those are the top 20 categories we will use for the classification
selected_categories = ['pl_usa', 'to_earn', 'to_acq', 'pl_uk', 'pl_japan', 'pl_canada', 'to_money-fx',
'to_crude', 'to_grain', 'pl_west-germany', 'to_trade', 'to_interest',
'pl_france', 'or_ec', 'pl_brazil', 'to_wheat', 'to_ship', 'pl_australia',
'to_corn', 'pl_china']
# Parse SGML files
document_X = []
document_Y = []
def strip_tags(text):
return re.sub('<[^<]+?>', '', text).strip()
def unescape(text):
return saxutils.unescape(text)
# Iterate all files
for i in range(sgml_number_of_files):
file_name = sgml_file_name_template.format(str(i).zfill(3))
print('Reading file: %s' % file_name)
with open(data_folder + file_name, 'rb') as file:
content = BeautifulSoup(file.read().lower(), "html.parser")
for newsline in content('reuters'):
document_categories = []
# News-line Id
document_id = newsline['newid']
# News-line text
document_body = strip_tags(str(newsline('text')[0].text)).replace('reuter\n', '')
document_body = unescape(document_body)
# News-line categories
topics = newsline.topics.contents
places = newsline.places.contents
people = newsline.people.contents
orgs = newsline.orgs.contents
exchanges = newsline.exchanges.contents
for topic in topics:
document_categories.append('to_' + strip_tags(str(topic)))
for place in places:
document_categories.append('pl_' + strip_tags(str(place)))
for person in people:
document_categories.append('pe_' + strip_tags(str(person)))
for org in orgs:
document_categories.append('or_' + strip_tags(str(org)))
for exchange in exchanges:
document_categories.append('ex_' + strip_tags(str(exchange)))
# Create new document
update_frequencies(document_categories)
document_X.append(document_body)
document_Y.append(to_category_vector(document_categories, selected_categories))
# -
# ## Select top 20 categories (by number of newslines)
news_categories.sort_values(by='Newslines', ascending=False, inplace=True)
# Selected categories
selected_categories = np.array(news_categories["Name"].head(20))
num_categories = 20
news_categories.head(num_categories)
# ### Take a look at the input and output data
# The output data **document_Y** looks good, it is a list of 20 (1 or 0) representing 20 categories.
#
# The input data still need to be cleaned up for the model input.
print(document_X[220])
print(document_Y[220])
# ## Clean up the data
# Function to clean up data
# * Only take characters inside A-Za-z0-9
# * remove stop words
# * lemmatize
# +
lemmatizer = WordNetLemmatizer()
strip_special_chars = re.compile("[^A-Za-z0-9 ]+")
stop_words = set(stopwords.words("english"))
def cleanUpSentence(r, stop_words = None):
r = r.lower().replace("<br />", " ")
r = re.sub(strip_special_chars, "", r.lower())
if stop_words is not None:
words = word_tokenize(r)
filtered_sentence = []
for w in words:
w = lemmatizer.lemmatize(w)
if w not in stop_words:
filtered_sentence.append(w)
return " ".join(filtered_sentence)
else:
return r
# -
# ### Here we are cleaning up the data
totalX = []
totalY = np.array(document_Y)
for i, doc in enumerate(document_X):
totalX.append(cleanUpSentence(doc, stop_words))
# ### Take a look at the input and output data now
# The input data looks clean and ready to be turned to ids
print(totalX[220])
print(totalY[220])
# ## Show input max sequence length
# If the max input sequence length is too long, we can put a limit to it in order to reduce the training time.
xLengths = [len(word_tokenize(x)) for x in totalX]
h = sorted(xLengths) #sorted lengths
maxLength =h[len(h)-1]
print("max input length is: ",maxLength)
# ### Choose a smaller max length for input sequence to reduce the training time
maxLength = h[int(len(h) * 0.70)]
print("70% cover input sequence length up to",maxLength)
# ## Convert input words to ids
# **max_vocab_size**: the maximum number of words to keep, we choose 30000 since it is big enough to keep all words in this case.
#
# Pad each input sequence to max input length **maxLength** if it is shorter.
max_vocab_size = 200000
input_tokenizer = Tokenizer(max_vocab_size)
input_tokenizer.fit_on_texts(totalX)
input_vocab_size = len(input_tokenizer.word_index) + 1
print("input_vocab_size:",input_vocab_size)
totalX = np.array(pad_sequences(input_tokenizer.texts_to_sequences(totalX), maxlen=maxLength))
import _pickle as cPickle
with open(r"input_tokenizer.pickle", "wb") as output_file:
cPickle.dump(input_tokenizer, output_file)
# ## Create Keras model
#
# * Embedding layer embed a sequence of vectors of size 256
# * GRU layers(recurrent network) which process the sequence data
# * Dense layer output the classification result of 20 categories
embedding_dim = 256
model = Sequential()
model.add(Embedding(input_vocab_size, embedding_dim,input_length = maxLength))
model.add(GRU(256, dropout=0.9, return_sequences=True))
model.add(GRU(256, dropout=0.9))
model.add(Dense(num_categories, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# ## Train model
history = model.fit(totalX, totalY, validation_split=0.1, batch_size=128, epochs=10)
model.save("classification_model.h5")
# ## Visualize the training performance
# +
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# ## Lets predict from one of our news data
# Here is the raw text from news at index 220
doc220 = document_X[220]
print(doc220)
# ### First clean up the input data
input_x_220 = cleanUpSentence(doc220, stop_words)
print(input_x_220)
# ### Turn the text to padded ids then feed to the model
textArray = np.array(pad_sequences(input_tokenizer.texts_to_sequences([input_x_220]), maxlen=maxLength))
from keras.models import load_model
new_model = load_model("classification_model.h5")
predicted = new_model.predict(textArray)[0]
for i, prob in enumerate(predicted):
print(prob, selected_categories[i])
# ### Lets take a look at the result
for i, prob in enumerate(predicted):
if prob > 0.2:
print(selected_categories[i])
# ### And here is the ground truth
# Looks like the model got 2 out of 3 right for the given text
categories_220 = document_Y[220]
for i, prob in enumerate(categories_220):
if prob > 0.2:
print(selected_categories[i])
| Text Multi-class multi-label Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Learning the Alphabet
#
# This is an example of a simple [LSTM](https://en.wikipedia.org/wiki/Long_short-term_memory) that is powerful enough to learn the alphabet. It is trained with strings that look like the alphabet.
#
# While this seems trivial, RNNs are capable of learning more complex text sequences, such as the works of Shakespeare or computer source code. Although training RNNs is computationally more expensive compared with other network structures, this example is simple enough to train on a CPU.
# See https://machinelearningmastery.com/text-generation-lstm-recurrent-neural-networks-python-keras/
# Small LSTM Network to Generate Text for Alice in Wonderland
import sys
import numpy
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import LSTM
from keras.callbacks import ModelCheckpoint
from keras.utils import np_utils
from matplotlib import pyplot as plt
# ## Training Data
#
# The training data is generated using characters from 'a' to 'z'. Several sequences are concatenated to produce a longer sequence of characters.
# +
# load ascii text and covert to lowercase
raw_text = ''.join([chr(x) for x in range(ord('a'), ord('z')+1)])
raw_text += raw_text
raw_text += raw_text
raw_text += raw_text
# create mapping of unique chars to integers
chars = sorted(list(set(raw_text)))
char_to_int = dict((c, i) for i, c in enumerate(chars))
int_to_char = dict((i, c) for i, c in enumerate(chars))
print("Subset of training data: %s" % raw_text[0:64])
# summarize the loaded data
n_chars = len(raw_text)
n_vocab = len(chars)
print("Total Characters: ", n_chars)
print("Total Vocab: ", n_vocab)
# -
# ## Prepare Sequences
#
# Input and expected output sequences are prepared and encoded as integers.
seq_length = 10
dataX = []
dataY = []
for i in range(0, n_chars - seq_length, 1):
seq_in = raw_text[i:i + seq_length]
seq_out = raw_text[i + seq_length]
dataX.append([char_to_int[char] for char in seq_in])
dataY.append(char_to_int[seq_out])
n_patterns = len(dataX)
print("Total Patterns: ", n_patterns)
# ## Train Model
# reshape X to be [samples, time steps, features]
X = numpy.reshape(dataX, (n_patterns, seq_length, 1))
# normalize
X = X / float(n_vocab)
# one hot encode the output variable
y = np_utils.to_categorical(dataY)
# define the LSTM model
model = Sequential()
model.add(LSTM(16, input_shape=(X.shape[1], X.shape[2])))
#model.add(Dropout(0.2))
model.add(Dense(y.shape[1], activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam')
# define the checkpoint
#filepath="weights-improvement-{epoch:02d}-{loss:.4f}.hdf5"
#checkpoint = ModelCheckpoint(filepath, monitor='loss', verbose=1, save_best_only=True, mode='min')
#callbacks_list = [checkpoint]
# fit the model
#model.fit(X, y, epochs=20, batch_size=128, callbacks=callbacks_list)
history = model.fit(X, y, epochs=100, batch_size=1, verbose=2)
# ## Model Loss
plt.plot(history.history['loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train'], loc='upper left')
plt.show()
print("Minimum loss: %.3f" % history.history['loss'][-1])
# ## Prediction
#
# A random starting sequence is used to seed prediction. The output is almost perfect.
# pick a random seed
start = numpy.random.randint(0, len(dataX)-1)
pattern = dataX[start]
#pattern = [start]
print("Seed:")
print("\"", ''.join([int_to_char[value] for value in pattern]), "\"")
# generate characters
for i in range(100):
x = numpy.reshape(pattern, (1, len(pattern), 1))
x = x / float(n_vocab)
prediction = model.predict(x, verbose=0)
index = numpy.argmax(prediction)
result = int_to_char[index]
seq_in = [int_to_char[value] for value in pattern]
sys.stdout.write(result)
pattern.append(index)
pattern = pattern[1:len(pattern)]
print("\nAll done!")
| Text/Alphabet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrdbourke/pytorch-deep-learning/blob/main/extras/exercises/03_pytorch_computer_vision_exercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Vex99np2wFVt"
# # 03. PyTorch Computer Vision Exercises
#
# The following is a collection of exercises based on computer vision fundamentals in PyTorch.
#
# They're a bunch of fun.
#
# You're going to get to write plenty of code!
#
# ## Resources
#
# 1. These exercises are based on [notebook 03 of the Learn PyTorch for Deep Learning course](https://www.learnpytorch.io/03_pytorch_computer_vision/).
# 2. See a live [walkthrough of the solutions (errors and all) on YouTube](https://youtu.be/_PibmqpEyhA).
# * **Note:** Going through these exercises took me just over 3 hours of solid coding, so you should expect around the same.
# 3. See [other solutions on the course GitHub](https://github.com/mrdbourke/pytorch-deep-learning/tree/main/extras/solutions).
# + colab={"base_uri": "https://localhost:8080/"} id="GaeYzOTLwWh2" outputId="17dd5453-9639-4b01-aa18-7ddbfd5c3253"
# Check for GPU
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="DNwZLMbCzJLk" outputId="9c150c50-a092-4f34-9d33-b45247fb080d"
# Import torch
import torch
# Exercises require PyTorch > 1.10.0
print(torch.__version__)
# TODO: Setup device agnostic code
# + [markdown] id="FSFX7tc1w-en"
# ## 1. What are 3 areas in industry where computer vision is currently being used?
# + id="VyWRkvWGbCXj"
# + [markdown] id="oBK-WI6YxDYa"
# ## 2. Search "what is overfitting in machine learning" and write down a sentence about what you find.
# + id="d1rxD6GObCqh"
# + [markdown] id="XeYFEqw8xK26"
# ## 3. Search "ways to prevent overfitting in machine learning", write down 3 of the things you find and a sentence about each.
# > **Note:** there are lots of these, so don't worry too much about all of them, just pick 3 and start with those.
# + id="ocvOdWKcbEKr"
# + [markdown] id="DKdEEFEqxM-8"
# ## 4. Spend 20-minutes reading and clicking through the [CNN Explainer website](https://poloclub.github.io/cnn-explainer/).
#
# * Upload your own example image using the "upload" button on the website and see what happens in each layer of a CNN as your image passes through it.
# + id="TqZaJIRMbFtS"
# + [markdown] id="lvf-3pODxXYI"
# ## 5. Load the [`torchvision.datasets.MNIST()`](https://pytorch.org/vision/stable/generated/torchvision.datasets.MNIST.html#torchvision.datasets.MNIST) train and test datasets.
# + id="SHjeuN81bHza"
# + [markdown] id="qxZW-uAbxe_F"
# ## 6. Visualize at least 5 different samples of the MNIST training dataset.
# + id="QVFsYi1PbItE"
# + [markdown] id="JAPDzW0wxhi3"
# ## 7. Turn the MNIST train and test datasets into dataloaders using `torch.utils.data.DataLoader`, set the `batch_size=32`.
# + id="ALA6MPcFbJXQ"
# + [markdown] id="bCCVfXk5xjYS"
# ## 8. Recreate `model_2` used in notebook 03 (the same model from the [CNN Explainer website](https://poloclub.github.io/cnn-explainer/), also known as TinyVGG) capable of fitting on the MNIST dataset.
# + id="5IKNF22XbKYS"
# + [markdown] id="sf_3zUr7xlhy"
# ## 9. Train the model you built in exercise 8. for 5 epochs on CPU and GPU and see how long it takes on each.
# + id="jSo6vVWFbNLD"
# + [markdown] id="w1CsHhPpxp1w"
# ## 10. Make predictions using your trained model and visualize at least 5 of them comparing the prediciton to the target label.
# + id="_YGgZvSobNxu"
# + [markdown] id="qQwzqlBWxrpG"
# ## 11. Plot a confusion matrix comparing your model's predictions to the truth labels.
# + id="vSrXiT_AbQ6e"
# + [markdown] id="lj6bDhoWxt2y"
# ## 12. Create a random tensor of shape `[1, 3, 64, 64]` and pass it through a `nn.Conv2d()` layer with various hyperparameter settings (these can be any settings you choose), what do you notice if the `kernel_size` parameter goes up and down?
# + id="leCTsqtSbR5P"
# + [markdown] id="VHS20cNTxwSi"
# ## 13. Use a model similar to the trained `model_2` from notebook 03 to make predictions on the test [`torchvision.datasets.FashionMNIST`](https://pytorch.org/vision/main/generated/torchvision.datasets.FashionMNIST.html) dataset.
# * Then plot some predictions where the model was wrong alongside what the label of the image should've been.
# * After visualing these predictions do you think it's more of a modelling error or a data error?
# * As in, could the model do better or are the labels of the data too close to each other (e.g. a "Shirt" label is too close to "T-shirt/top")?
# + id="78a8LjtdbSZj"
| extras/exercises/03_pytorch_computer_vision_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## El modelo del objeto documento (*DOM*).
#
# Al abrir un documento *HTML* en cualquier navegador, dicho documento es considerado com un objeto, el cual a su vez contiene otros objetos o elementos.
#
# El modelo del objeto documento (*DOM*) es una manera estandarizada en la que se puede acceder a los elementos de un documento *HTML* mediante *Javascript*.
# ### Árbol de nodos.
#
# El *DOM* permite identificar a los elementos de un documento *HTML* como una serie de nodos que a su vez podrían contener a otros nodos, los cuales se van bifurcando a partir de la raiz del documento tal como lo hacen las ramas de un árbol a partir de su tronco.
#
# ### El objeto ```document```.
#
# El objeto ```document``` contiene al documento *HTML* que se despiega en la ventana de un navegador y es mediante el acceso a sus propiedades y métodos como se puede acceder a los elementos que contiene.
#
#
# ### Tipos de nodos que puede contener el *DOM*.
#
# * ```Document```, el cual corresponde al nodo raíz.
# * ```Element```, el cual representa a cada elemento por medio de su etiqueta.
# * ```Attr```, correspondiente a un atributo del elemento.
# * ```Text```, contiene el texto que está dentro del elemento.
# * ```Comment```, comentarios del documento.
#
# ### Manipulación de nodos.
#
# Mediante el *DOM*, es posible acceder a cada elemento del documento *HTML* y realizar las siguientes operaciones:
#
# * Acceder a las propiedades de un elemento.
# * Establecer valores de las propiedades de un elemento.
# * Mover un elemento de la página.
# * Crear o eliminar elementos.
#
# **NOTA:** El acceso a los elementos de un documento *HTML* solamente pueden ser realizados cuando la página se haya cargado por completo.
#
# ### Acceso a los nodos.
#
# Como se mencionó previamente, el *DOM* presenta una estructura similar a la del tronco de un árbol a partir del cual se bifurcan ramas. Se dice que el elemento que contiene a otros elementos es el padre de estos.
#
# ```
# padre (parent)
# ├─hermano (sibling)
# ├─elemento
# │ └──── hijos (children)
# └───hermano (sibling)
# ```
# ## El método ```document.getRootNode()```.
#
# Este método permite acceder al nodo raíz del documento. El elemento resultante contiene a los elementos ```head``` y ```body``` como nodos hijo.
# **Ejemplo:**:
#
# * Abrir una ventana en blanco dentro de un navegador (se recomienda Firefox).
# * Seleccionar la consola del navegador a partir de la inspección de los elementos.
# * Al ejecutar el siguente código se asignará el nodo raíz será asignado a la variable ```documento```.
#
# ``` javascript
# const documento = document.getRootNode();
# ```
#
# * Para acceder al elemento ```body``` del documento *HTML* se ejecuta lo siguiente:
#
# ``` javascript
# documento.body;
# ```
#
# Se desplegará el contenido y los atributos del elemento ```body```.
# ### Propiedades básicas de un elemento.
#
# Los elementos contienen múltiples propiedades emanadas de ```document```, pero las más características son:
#
# * ```textContent```, la cual corresponde al texto del elemento.
# * ```children```, la cual corresponde a un arreglo que enlista a los elementos que contiene el elemento.
# * ```childNodes```, la cual corresponde a un arreglo que enlista los nodos que contiene el elemento.
# * ```innerHTML```, la cual corresponde a un objeto de tipo ```String``` que contiene el código *HTML* del elemento.
# * ```innerText```, la cual corresponde a un objeto de tipo ```String``` que contiene el texto del elemento y sus elementos contenidos.
# * ```id```, la cual corresponde a un objeto de tipo ```String``` que contiene el atributo ```id``` del elemento.
# * ```className```, la cual corresponde al valor del atributo ```class``` del elemento.
# * ```parentElement``` la cual corresponde al elemento padre del elemento actual.
# * ```parentNode``` la cual corresponde al nodo padre del elemento actual.
# * ```style``` la cual contiene los propiedades de estilo del elemento.
# **Ejemplo:**
# + language="html"
# <div id="contenedor-1">
# <div id="ejemplo-1">Texto de ejemplo <b>con efectos</b> y <i>más efectos.</i></div>
# <div>
# + language="javascript"
# let contenedor_1 = document.getElementById("contenedor-1");
# let elemento_1 = document.getElementById("ejemplo-1");
# let parrafo = document.createElement("p");
# parrafo.textContent = elemento_1.innerHTML;
# contenedor_1.appendChild(parrafo);
# + language="html"
# <div id="contenedor-2">
# <div id="ejemplo-2">Texto de ejemplo <b>con efectos</b> y <i>más efectos.</i></div>
# <div>
# + language="javascript"
# let contenedor_2 = document.getElementById("contenedor-2");
# let elemento_2 = document.getElementById("ejemplo-2");
# let hijos = elemento_2.children;
# for (let item in hijos) {
# let parrafo = document.createElement("p");
# parrafo.textContent = "Elemento: " + hijos[item].nodeName + ", texto: " + hijos[item].textContent;
# contenedor_2.appendChild(parrafo);
# }
# -
# ## Creación de un elemento.
#
# Es posible crear un elemento mendiante el método ```document.createElement()```, indicando el tipo de elemento mediante el nombre de su etiqueta.
# **Ejemplo:**
#
# * Se creará un elemento de tipo ```p``` y se asignará a la variable ```parrafo```.
# + language="html"
# <div id="contenedor-3">
# <div id="ejemplo-3"></div>
# <div>
# + language="javascript"
# let ejemplo_3 = document.getElementById("ejemplo-3");
# let parrafo = document.createElement("p");
# parrafo.textContent="Hola, Mundo."
# ejemplo_3.append(parrafo);
# -
# ## Métodos de acceso a elementos emanado de ```document```.
#
# * ```getElementsByTagName()```, el cual regresa un arreglo con todos los elementos con el tipo de etiqueta ingresada como argumento.
# * ```getElementsByName()```, el cual regresa un arreglo con todos los elementos con el nombre ingresado como argumento.
# * ```getElementById()```, el cual regresa al elmentos cuyo valor del atributo ```id``` coincida con la cadena de caravteres ingresada como argumento.
# * ```getElementsByClassName()```, el cual regresa un arrgelo con todos los elementos cuyo valro del atributo ```class``` coincida con la cadena de caracteres ingresada como argumento.
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2022.</p>
| 08_modelo_del_objeto_documento.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Casual Coded Correspondence: The Project
#
# In this project, you will be working to code and decode various messages between you and your fictional cryptography enthusiast pen pal Vishal. You and Vishal have been exchanging letters for quite some time now and have started to provide a puzzle in each one of your letters. Here is his most recent letter:
#
# Hey there! How have you been? I've been great! I just learned about this really cool type of cipher called a Caesar Cipher. Here's how it works: You take your message, something like "hello" and then you shift all of the letters by a certain offset. For example, if I chose an offset of 3 and a message of "hello", I would code my message by shifting each letter 3 places to the left (with respect to the alphabet). So "h" becomes "e", "e" becomes, "b", "l" becomes "i", and "o" becomes "l". Then I have my coded message,"ebiil"! Now I can send you my message and the offset and you can decode it. The best thing is that Julius Caesar himself used this cipher, that's why it's called the Caesar Cipher! Isn't that so cool! Okay, now I'm going to send you a longer coded message that you have to decode yourself!
#
# xuo jxuhu! jxyi yi qd unqcfbu ev q squiqh syfxuh. muhu oek qrbu je tusetu yj? y xefu ie! iudt cu q cuiiqwu rqsa myjx jxu iqcu evviuj!
#
# This message has an offset of 10. Can you decode it?
#
#
# #### Step 1: Decode Vishal's Message
# In the cell below, use your Python skills to decode Vishal's message and print the result. Hint: you can account for shifts that go past the end of the alphabet using the modulus operator, but I'll let you figure out how!
alphabet = "abcdefghijklmnopqrstuvwxyz"
punctuation = ".,?'! "
message = "xuo jxuhu! jxyi yi qd unqcfbu ev q squiqh syfxuh. muhu oek qrbu je tusetu yj? y xefu ie! iudt cu q cuiiqwu rqsa myjx jxu iqcu evviuj!"
translated_message = ""
for letter in message:
if not letter in punctuation:
letter_value = alphabet.find(letter)
translated_message += alphabet[(letter_value + 10) % 26]
else:
translated_message += letter
print(translated_message)
# #### Step 2: Send Vishal a Coded Message
# Great job! Now send Vishal back a message using the same offset. Your message can be anything you want! Remember, coding happens in opposite direction of decoding.
message_for_v = "hey vishal! This is a super cool cipher, thanks for showing me! What else you got?"
translated_message = ""
for letter in message_for_v:
if not letter in punctuation:
letter_value = alphabet.find(letter)
translated_message += alphabet[(letter_value - 14) % 26]
else:
translated_message += letter
print(translated_message)
# #### Step 3: Make functions for decoding and coding
#
# Vishal sent over another reply, this time with two coded messages!
#
# You're getting the hang of this! Okay here are two more messages, the first one is coded just like before with an offset of ten, and it contains the hint for decoding the second message!
#
# First message:
#
# jxu evviuj veh jxu iusedt cuiiqwu yi vekhjuud.
#
# Second message:
#
# bqdradyuzs ygxfubxq omqemd oubtqde fa oapq kagd yqeemsqe ue qhqz yadq eqogdq!
#
# Decode both of these messages.
#
# If you haven't already, define two functions `decoder(message, offset)` and `coder(message, offset)` that can be used to quickly decode and code messages given any offset.
# +
# both of these functions need the strings `alphabet` and `punctuation` defined before being run
def decoder(message, offset):
translated_message = ""
for letter in message:
if not letter in punctuation:
letter_value = alphabet.find(letter)
translated_message += alphabet[(letter_value + offset) % 26]
else:
translated_message += letter
return translated_message
def coder(message, offset):
translated_message = ""
for letter in message:
if not letter in punctuation:
letter_value = alphabet.find(letter)
translated_message += alphabet[(letter_value - offset) % 26]
else:
translated_message += letter
return translated_message
message_one = "jxu evviuj veh jxu iusedt cuiiqwu yi vekhjuud."
# Now we'll print the output of `decoder` for the first message with an offset of 10
print(decoder(message_one, 10))
# +
# Now we know what offset to use for the second message, so we use that to solve.
message_two = "bqdradyuzs ygxfubxq omqemd oubtqde fa oapq kagd yqeemsqe ue qhqz yadq eqogdq!"
print(decoder(message_two, 14))
# -
# #### Step 4: Solving a Caesar Cipher without knowing the shift value
#
# Awesome work! While you were working to decode his last two messages, Vishal sent over another letter! He's really been bitten by the crytpo-bug. Read it and see what interesting task he has lined up for you this time.
#
# Hello again friend! I knew you would love the Caesar Cipher, it's a cool simple way to encrypt messages. Did you know that back in Caesar's time, it was considered a very secure way of communication and it took a lot of effort to crack if you were unaware of the value of the shift? That's all changed with computers! Now we can brute force these kinds of ciphers very quickly, as I'm sure you can imagine.
#
# To test your cryptography skills, this next coded message is going to be harder than the last couple to crack. It's still going to be coded with a Caesar Cipher but this time I'm not going to tell you the value of the shift. You'll have to brute force it yourself.
#
# Here's the coded message:
#
# vhfinmxkl atox kxgwxkxw tee hy maxlx hew vbiaxkl tl hulhexmx. px'ee atox mh kxteer lmxi ni hnk ztfx by px ptgm mh dxxi hnk fxlltzxl ltyx.
#
# Good luck!
#
# Decode Vishal's most recent message and see what it says!
# +
coded_message = "vhfinmxkl atox kxgwxkxw tee hy maxlx hew vbiaxkl tl hulhexmx. px'ee atox mh kxteer lmxi ni hnk ztfx by px ptgm mh dxxi hnk fxlltzxl ltyx."
# The easiest way to break this code is to simply brute force though all of the possible shifts.
# We'll only need to try 25 different shifts, so it's not computationally expensive. Then we can
# look through all of the outputs and look for the one that in english, and we've decoded our message!
for i in range(1,26):
print("offset: " + str(i))
print("\t " + decoder(coded_message, i) + "\n")
# -
# #### Step 5: The Vigenère Cipher
#
# Great work! While you were working on the brute force cracking of the cipher, Vishal sent over another letter. That guy is a letter machine!
#
# Salutations! As you can see, technology has made brute forcing simple ciphers like the Caesar Cipher extremely easy, and us crypto-enthusiasts have had to get more creative and use more complicated ciphers. This next cipher I'm going to teach you is the Vigenère Cipher, invented by an Italian cryptologist named <NAME> (cool name eh?) in the 16th century, but named after another cryptologist from the 16th century, Blaise de Vigenère.
#
# The Vigenère Cipher is a polyalphabetic substitution cipher, as opposed to the Caesar Cipher which was a monoalphabetic substitution cipher. What this means is that opposed to having a single shift that is applied to every letter, the Vigenère Cipher has a different shift for each individual letter. The value of the shift for each letter is determined by a given keyword.
#
# Consider the message
#
# barryisthespy
#
# If we want to code this message, first we choose a keyword. For this example, we'll use the keyword
#
# dog
#
# Now we use the repeat the keyword over and over to generate a _keyword phrase_ that is the same length as the message we want to code. So if we want to code the message "barryisthespy" our _keyword phrase_ is "dogdogdogdogd". Now we are ready to start coding our message. We shift the each letter of our message by the place value of the corresponding letter in the keyword phrase, assuming that "a" has a place value of 0, "b" has a place value of 1, and so forth. Remember, we zero-index because this is Python we're talking about!
#
# message: b a r r y i s t h e s p y
#
# keyword phrase: d o g d o g d o g d o g d
#
# resulting place value: 4 14 15 12 16 24 11 21 25 22 22 17 5
#
# So we shift "b", which has an index of 1, by the index of "d", which is 3. This gives us an place value of 4, which is "e". Then continue the trend: we shift "a" by the place value of "o", 14, and get "o" again, we shift "r" by the place value of "g", 15, and get "x", shift the next "r" by 12 places and "u", and so forth. Once we complete all the shifts we end up with our coded message:
#
# eoxumovhnhgvb
#
# As you can imagine, this is a lot harder to crack without knowing the keyword! So now comes the hard part. I'll give you a message and the keyword, and you'll see if you can figure out how to crack it! Ready? Okay here's my message:
#
# dfc jhjj ifyh yf hrfgiv xulk? vmph bfzo! qtl eeh gvkszlfl yyvww kpi hpuvzx dl tzcgrywrxll!
#
# and the keyword to decode my message is
#
# friends
#
# Because that's what we are! Good luck friend!
#
# And there it is. Vishal has given you quite the assignment this time! Try to decode his message. It may be helpful to create a function that takes two parameters, the coded message and the keyword and then work towards a solution from there.
# +
def vigenere_decoder(coded_message, keyword):
keyword_repeated = ''
while len(keyword_repeated) < len(coded_message):
keyword_repeated += keyword
keyword_final = keyword_repeated[0:len(coded_message)]
translated_message = ''
for i in range(0,len(coded_message)):
if not coded_message[i] in punctuation:
ln = alphabet.find(coded_message[i]) - alphabet.find(keyword_final[i])
translated_message += alphabet[ln % 26]
else:
translated_message += coded_message[i]
return translated_message
message = "dfc jhjj ifyh yf hrfgiv xulk? vmph bfzo! qtl eeh gvkszlfl yyvww kpi hpuvzx dl tzcgrywrxll!"
keyword = "friends"
print(vigenere_decoder(message, keyword))
# -
# #### Step 6: Send a message with the Vigenère Cipher
# Great work decoding the message. For your final task, write a function that can encode a message using a given keyword and write out a message to send to Vishal!
# +
def vigenere_coder(message, keyword):
keyword_repeated = ''
while len(keyword_repeated) < len(message):
keyword_repeated += keyword
keyword_final = keyword_repeated[0:len(message)]
translated_message = ''
for i in range(0,len(message)):
if message[i] not in punctuation:
ln = alphabet.find(message[i]) + alphabet.find(keyword_final[i])
translated_message += alphabet[ln % 26]
else:
translated_message += message[i]
return translated_message
message_for_v = "thanks for teaching me all these cool ciphers! you really are the best!"
keyword = "besties"
print(vigenere_coder(message_for_v,keyword))
# -
# #### Conclusion
# Over the course of this project you've learned about two different cipher methods and have used your Python skills to code and decode messages. There are all types of other facinating ciphers out there to explore, and Python is the perfect language to implement them with, so go exploring!
| Cumulative_Projects/Wk 6 coded_correspondence_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imminent ICU Admission and Prolonged Stay Prediction using Gradient Boosting Machines
# ## Imports & Inits
# %load_ext autoreload
# %autoreload 2
# +
import sys
sys.path.append('../')
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("darkgrid")
# %matplotlib inline
import numpy as np
np.set_printoptions(precision=2)
import pandas as pd
import pickle
from sklearn.feature_extraction.text import TfidfVectorizer
import lightgbm
from utils.splits import set_group_splits
from utils.metrics import BinaryAvgMetrics, get_best_model
from utils.plots import *
from args import args
vars(args)
# -
# ## GBM Dev
# +
seed = 643
ori_df = pd.read_csv(args.dataset_csv, usecols=args.cols, parse_dates=args.dates)
ia_df = ori_df.loc[(ori_df['imminent_adm_label'] != -1)][args.imminent_adm_cols].reset_index(drop=True)
ps_df = ori_df.loc[(ori_df['chartinterval'] != 0)][args.prolonged_stay_cols].reset_index(drop=True)
print(ia_df.shape)
print(ps_df.shape)
# -
# ### Imminent ICU Admission
df = set_group_splits(ia_df.copy(), group_col='hadm_id', seed=seed)
vectorizer = TfidfVectorizer(sublinear_tf=True, ngram_range=(1,2), binary=True, max_features=60_000)
x_train = vectorizer.fit_transform(df.loc[(df['split'] == 'train')]['processed_note'])
x_test = vectorizer.transform(df.loc[(df['split'] == 'test')]['processed_note'])
y_train = df.loc[(df['split'] == 'train')]['imminent_adm_label'].to_numpy()
y_test = df.loc[(df['split'] == 'test')]['imminent_adm_label'].to_numpy()
df.shape, x_train.shape, x_test.shape, y_train.shape, y_test.shape
params = {
"objective": "binary",
"metric": "binary_logloss",
"bagging_fraction": 0.5,
"bagging_freq": 5,
"boosting": "dart",
"feature_fraction": 0.5,
"is_unbalance": True,
"learning_rate": 0.1,
"min_data_in_leaf": 3,
"num_iterations": 150,
"num_leaves": 50,
}
clf = lightgbm.LGBMClassifier(**params, n_threads=32)
clf.fit(x_train, y_train)
prob = clf.predict_proba(x_test)[:, 1]
fig, ax = plt.subplots(figsize=(20, 10))
plot_youden(ax, y_test, prob, 0.1, 0.9, 40)
fig, ax = plt.subplots(figsize=(20, 10))
plot_thresh_range(ax, y_test, prob, 0.1, 0.9, 40)
fig, ax = plt.subplots(figsize=(10,8))
plot_roc(ax, y_test, prob)
# +
threshold = 0.42
y_pred = (prob > threshold).astype(np.int64)
cm = confusion_matrix(y_test, y_pred)
tn,fp,fn,tp = cm[0][0],cm[0][1],cm[1][0],cm[1][1]
sensitivity = tp/(tp+fn)
specificity = tn/(tn+fp)
ppv = tp/(tp+fp)
npv = tn/(tn+fn)
f1 = (2*ppv*sensitivity)/(ppv+sensitivity)
auroc = roc_auc_score(y_test, prob)
d = {
'sensitivity': np.round(sensitivity, 3),
'specificity': np.round(specificity, 3),
'ppv': np.round(ppv, 3),
'npv': np.round(npv, 3),
'f1': np.round(f1, 3),
'auroc': np.round(auroc, 3),
}
metrics = pd.DataFrame(d.values(), index=d.keys(), columns=['Value'])
metrics
# -
fig, ax = plt.subplots(figsize=(10, 8))
plot_confusion_matrix(ax, cm, classes=['Delayed', 'Imminent'], normalize=False, title='Confusion matrix')
# +
import re
from spacy.lang.en.stop_words import STOP_WORDS
def get_top_markers(feature_names, scores, n_markers=50):
p = re.compile('^[a-z\s]+$')
sorted_idxs = np.argsort(scores)
markers = []
wts = []
for i in sorted_idxs:
marker = feature_names[i]
if len(marker) > 7 and marker not in STOP_WORDS and p.match(marker):
markers.append(marker)
wts.append(scores[i])
wts = np.asarray(wts)
df = pd.DataFrame({'markers': markers, 'score': wts/wts.sum()}, columns=['markers', 'score'])
return df.head(n_markers), df.tail(n_markers).sort_values(by='score').reset_index(drop=True)
# +
# fig, ax = plt.subplots(1, 2, figsize=(15, 15))
# sns.barplot(x='score', y='markers', data=pos_df, color='b', ax=ax[0])
# sns.barplot(x='score', y='markers', data=neg_df, color='b', ax=ax[1])
# ax[0].set_xlabel('')
# ax[1].set_xlabel('')
# ax[1].set_ylabel('')
# plt.subplots_adjust(wspace = 0.3)
# fig.text(0.5, 0.1, 'Score', ha='center')
# +
# scores = clf.feature_importances_ / clf.feature_importances_.sum()
# fig, ax = plt.subplots(1, 2, figsize=(20, 10))
# neg_cloud, pos_cloud = get_wordcloud(vectorizer.get_feature_names(), scores, n_words=50)
# ax[0].imshow(neg_cloud)
# ax[0].axis('off')
# ax[0].set_title('Delayed')
# ax[1].imshow(pos_cloud)
# ax[1].axis('off')
# ax[1].set_title('Imminent')
# -
# ### Prolonged ICU Stay
df = set_group_splits(ps_df.copy(), group_col='hadm_id', seed=seed)
vectorizer = TfidfVectorizer(sublinear_tf=True, ngram_range=(1,2), binary=True, max_features=60_000)
x_train = vectorizer.fit_transform(df.loc[(df['split'] == 'train')]['processed_note'])
x_test = vectorizer.transform(df.loc[(df['split'] == 'test')]['processed_note'])
y_train = df.loc[(df['split'] == 'train')]['prolonged_stay_label'].to_numpy()
y_test = df.loc[(df['split'] == 'test')]['prolonged_stay_label'].to_numpy()
df.shape, x_train.shape, x_test.shape, y_train.shape, y_test.shape
params = {
"objective": "binary",
"metric": "binary_logloss",
"is_unbalance": True,
"bagging_fraction": 0.7,
"bagging_freq": 6,
"boosting": "gbdt",
"feature_fraction": 0.5,
"learning_rate": 0.25,
"min_data_in_leaf": 5,
"num_iterations": 172,
}
clf = lightgbm.LGBMClassifier(**params, n_threads=32)
clf.fit(x_train, y_train)
prob = clf.predict_proba(x_test)[:, 1]
fig, ax = plt.subplots(figsize=(20, 10))
plot_youden(ax, y_test, prob, 0.1, 0.9, 40)
fig, ax = plt.subplots(figsize=(20, 10))
plot_thresh_range(ax, y_test, prob, 0.1, 0.9, 40)
fig, ax = plt.subplots(figsize=(10,8))
plot_roc(ax, y_test, prob)
# +
threshold = 0.34
y_pred = (prob > threshold).astype(np.int64)
cm = confusion_matrix(y_test, y_pred)
tn,fp,fn,tp = cm[0][0],cm[0][1],cm[1][0],cm[1][1]
sensitivity = tp/(tp+fn)
specificity = tn/(tn+fp)
ppv = tp/(tp+fp)
npv = tn/(tn+fn)
f1 = (2*ppv*sensitivity)/(ppv+sensitivity)
auroc = roc_auc_score(y_test, prob)
d = {
'sensitivity': np.round(sensitivity, 3),
'specificity': np.round(specificity, 3),
'ppv': np.round(ppv, 3),
'npv': np.round(npv, 3),
'f1': np.round(f1, 3),
'auroc': np.round(auroc, 3),
}
metrics = pd.DataFrame(d.values(), index=d.keys(), columns=['Value'])
metrics
# -
fig, ax = plt.subplots(figsize=(10, 8))
plot_confusion_matrix(ax, cm, classes=['Short Stay', 'Prolonged Stay'], normalize=False, title='Confusion matrix')
# +
# scores = clf.feature_importances_ / clf.feature_importances_.sum()
# fig, ax = plt.subplots(1, 2, figsize=(20, 10))
# neg_cloud, pos_cloud = get_wordcloud(vectorizer.get_feature_names(), clf.feature_importances_/clf.feature_importances_.sum(), n_words=50)
# ax[0].imshow(neg_cloud)
# ax[0].axis('off')
# ax[0].set_title('Negative Class')
# ax[1].imshow(pos_cloud)
# ax[1].axis('off')
# ax[1].set_title('Positive Class')
# -
# ## Metrics
# Taken from [here](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/):
#
# 1. Prevalence: `(fn + tp) / total`
# 2. Sensitivity: AKA recall, true positive rate `tp / (tp + fn)`
# 3. Specificity: AKA true negative rate `tn / (tn + fp)`
# 4. Positive Predictive Value (PPV): AKA precision `tp / (tp + fp)`
# 5. Negative Predictive Value (NPV): `tn / (tn + fn)`
# ### Imminent ICU Admission
# +
with open(args.workdir/f'imminent_adm_preds.pkl', 'rb') as f:
targs = pickle.load(f)
preds = pickle.load(f)
probs = pickle.load(f)
fnames = [f'imminent_adm_seed_{seed}.pkl' for seed in range(args.start_seed, args.start_seed + 100)]
bam = BinaryAvgMetrics(targs, preds, probs)
# -
bam.get_avg_metrics(conf=0.95)
get_best_model(bam, fnames)
fig, ax = plt.subplots(figsize=(10, 8))
plot_mean_roc(ax, bam.targs, bam.probs)
# +
fig, ax = plt.subplots(1, 2, figsize=(15, 6))
plot_confusion_matrix(ax[0], bam.cm_avg, classes=['not imminent', 'imminent'], normalize=False,\
title='Confusion Matrix Over Runs')
plot_confusion_matrix(ax[1], bam.cm_avg, classes=['not imminent', 'imminent'], normalize=True,\
title='Normalized Confusion Matrix Over Runs')
plt.show()
# -
# ### Prolonged ICU Stay
# +
with open(args.workdir/f'prolonged_stay_preds.pkl', 'rb') as f:
targs = pickle.load(f)
preds = pickle.load(f)
probs = pickle.load(f)
fnames = [f'prolonged_stay_seed_{seed}.pkl' for seed in range(args.start_seed, args.start_seed + 100)]
bam = BinaryAvgMetrics(targs, preds, probs)
# -
bam.get_avg_metrics(conf=0.95)
get_best_model(bam, fnames)
fig, ax = plt.subplots(figsize=(10, 8))
plot_mean_roc(ax, bam.targs, bam.probs)
# +
fig, ax = plt.subplots(1, 2, figsize=(15, 6))
plot_confusion_matrix(ax[0], bam.cm_avg, classes=['Discharge within 5 days', 'Discharge after 5 days'], normalize=False, title='Confusion Matrix Over Runs')
plot_confusion_matrix(ax[1], bam.cm_avg, classes=['Discharge within 5 days', 'Discharge after 5 days'], normalize=True, title='Normalized Confusion Matrix Over Runs')
plt.show()
# -
# ## Full Run
# +
seed = 643
ori_df = pd.read_csv(args.dataset_csv, usecols=args.cols, parse_dates=args.dates)
ori_df['relative_charttime'] = (ori_df['charttime'] - ori_df['intime'])
ia_df = ori_df.loc[(ori_df['imminent_adm_label'] != -1)][args.imminent_adm_cols + ['relative_charttime']].reset_index(drop=True)
ps_df = ori_df.loc[(ori_df['chartinterval'] != 0)][args.prolonged_stay_cols + ['relative_charttime']].reset_index(drop=True)
# -
interval_hours = 12
starting_day = -20
ending_day = -1
# ### Imminent ICU Admission
# +
df = set_group_splits(ia_df.copy(), pct=0.25, group_col='hadm_id', seed=seed)
df['prob'] = -1
vectorizer = TfidfVectorizer(min_df=args.min_freq, analyzer=str.split, sublinear_tf=True, ngram_range=(2,2))
x_train = vectorizer.fit_transform(df.loc[(df['split'] == 'train')]['processed_note'])
x_test = vectorizer.transform(df.loc[(df['split'] == 'test')]['processed_note'])
y_train = df.loc[(df['split'] == 'train')]['imminent_adm_label'].to_numpy()
y_test = df.loc[(df['split'] == 'test')]['imminent_adm_label'].to_numpy()
df.shape, x_train.shape, x_test.shape, y_train.shape, y_test.shape
# +
from args import ia_params
clf = lightgbm.LGBMClassifier(**ia_params)
clf.fit(x_train, y_train)
df.loc[(df['split'] == 'test'), 'prob'] = clf.predict_proba(x_test)[:, 1]
# -
fig, ax = plt.subplots(figsize=(15, 8))
plot_prob(ax, df.loc[(df['split'] == 'test')], args.ia_thresh, starting_day, ending_day, interval_hours)
scores = clf.feature_importances_ / clf.feature_importances_.sum()
fig, ax = plt.subplots(1, 2, figsize=(20, 10))
neg_cloud, pos_cloud = get_wordcloud(vectorizer.get_feature_names(), clf.feature_importances_, n_words=50)
ax[0].imshow(neg_cloud)
ax[0].axis('off')
ax[0].set_title('Negative Class')
ax[1].imshow(pos_cloud)
ax[1].axis('off')
ax[1].set_title('Positive Class')
# ### Prolonged ICU Stay
# +
df = set_group_splits(ps_df.copy(), pct=0.25, group_col='hadm_id', seed=seed)
df['prob'] = -1
vectorizer = TfidfVectorizer(min_df=args.min_freq, analyzer=str.split, sublinear_tf=True, ngram_range=(2,2))
x_train = vectorizer.fit_transform(df.loc[(df['split'] == 'train')]['processed_note'])
x_test = vectorizer.transform(df.loc[(df['split'] == 'test')]['processed_note'])
y_train = df.loc[(df['split'] == 'train')]['prolonged_stay_label'].to_numpy()
y_test = df.loc[(df['split'] == 'test')]['prolonged_stay_label'].to_numpy()
df.shape, x_train.shape, x_test.shape, y_train.shape, y_test.shape
# +
from args import ps_params
clf = lightgbm.LGBMClassifier(**ps_params)
clf.fit(x_train, y_train)
df.loc[(df['split'] == 'test'), 'prob'] = clf.predict_proba(x_test)[:, 1]
# -
fig, ax = plt.subplots(figsize=(15, 8))
plot_prob(ax, df.loc[(df['split'] == 'test')], args.ps_thresh, starting_day, ending_day, interval_hours)
scores = clf.feature_importances_ / clf.feature_importances_.sum()
fig, ax = plt.subplots(1, 2, figsize=(20, 10))
neg_cloud, pos_cloud = get_wordcloud(vectorizer.get_feature_names(), clf.feature_importances_, n_words=50)
ax[0].imshow(neg_cloud)
ax[0].axis('off')
ax[0].set_title('Negative Class')
ax[1].imshow(pos_cloud)
ax[1].axis('off')
ax[1].set_title('Positive Class')
| gbm/gbm_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Change directory to VSCode workspace root so that relative path loads work correctly. Turn this addition off with the DataSciece.changeDirOnImportExport setting
import os
try:
os.chdir(os.path.join(os.getcwd(), '..'))
print(os.getcwd())
except:
pass
# ## Run preprocessor.py manually
# in all other cases add 'from preprocess import csv_2_days'
import os
import sys
nb_dir = os.path.split(os.getcwd())[0]
if nb_dir not in sys.path:
sys.path.append(nb_dir)
from homesensors.preprocess import *
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# not local, not yet shared on github
filename = "balcony_temperature.csv"
days = csv_2_days_map(filename)
print(f"nb days loaded = {len(days)}")
# +
np_time = np.array(days["2018-05-27"]["timestamp"]).astype("datetime64")
np_temp = np.array(days["2018-05-27"]["temperature"]).astype("float")
plt.plot(np_time, np_temp)
plt.show()
# -
np_days = csv_2_days_np(filename)
plt.plot(np_days["2018-05-28"]["timestamp"], np_days["2018-05-28"]["temperature"])
plt.show()
days_add_params(np_days)
print(f'average = {np_days["2018-05-28"]["mean"]}')
a = np_days["2018-05-28"]["temperature"]
plt.hist(a, bins='auto') # arguments are passed to np.histogram
plt.title("Histogram with 'auto' bins")
plt.show()
# +
series = days_to_array(np_days)
df = pd.DataFrame(series)
df.plot('day', 'min')
plt.show()
# -
plt.plot( 'day', 'min', data=df, marker='', color='skyblue', linewidth=2)
plt.plot( 'day', 'mean', data=df, marker='', color='olive', linewidth=4)
plt.plot( 'day', 'max', data=df, marker='', color='red', linewidth=2)
plt.show()
| homesensors/analyse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from silx.gui import qt
# %gui qt
# # simple plot of a 2D image
#
# - using Plot2D
# ## load data from data/lena.hdf5
# input using .hdf5
import h5py
import numpy
dataPath='../data/ascent.h5'
f=h5py.File(dataPath)
from silx.io.utils import h5ls
h5ls(dataPath)
image=numpy.array(f['data'], dtype='float32')
# ## plot the image
# 
from silx.gui.plot import Plot2D
plotImage=Plot2D()
plotImage.addImage(image, origin=(0, 0), legend='sino')
plotImage.show()
# # display the pixel intensity distribution
# ## create the histogramnd
# - using silx.math.histogram.Histogramnd
#
# - http://www.silx.org/doc/silx/dev/modules/math/histogram.html
from silx.math.histogram import Histogramnd
histo, w_histo, edges = Histogramnd(image.flatten(),
n_bins=256,
histo_range=[0,256])
# ## plot the histogram
# - using silx.gui.plot.Plot1d
from silx.gui.plot import Plot1D
plotHisto = Plot1D()
plotHisto.addCurve(range(256), histo, legend='intensity')
plotHisto.show()
# # create a PlotAction which plot the histogram for the current image
#
# - using silx.gui.plot.PlotActions.PlotAction
#
# - <EMAIL>@ http://www.silx.org/doc/silx/dev/modules/gui/plot/plotactions_examples.html
#
# 
# +
from silx.gui.plot.PlotActions import PlotAction
from silx.math.histogram import Histogramnd
from silx.gui.plot import Plot1D
class ComputeHistogramAction(PlotAction):
"""Computes the intensity distribution on the current image
:param plot: :class:`.PlotWidget` instance on which to operate
:param parent: See :class:`QAction`
"""
def __init__(self, plot, parent=None):
PlotAction.__init__(self,
plot,
icon='shape-circle',
text='pixels intensity',
tooltip='Compute image intensity distribution',
triggered=self.computeIntensityDistribution,
parent=parent)
self.plotHistogram=Plot1D()
def computeIntensityDistribution(self):
"""Get the active image and compute the image
intensity distribution"""
# By inheriting from PlotAction, we get access to attribute
# self.plot
# which is a reference to the PlotWindow
activeImage = self.plot.getActiveImage()
if activeImage is not None:
histo, w_histo, edges = Histogramnd(activeImage[0].flatten(),
n_bins=256,
histo_range=[0,256])
self.plotHistogram.addCurve(range(256),
histo,
legend='pixel intensity')
self.plotHistogram.show()
# -
# ## Add this action into the toolBar of the window
plotHisto.clear()
myaction=ComputeHistogramAction(plotImage)
toolBar=plotImage.toolBar()
toolBar.addAction(myaction)
plotImage.show()
# # show automatically the histogram when the image change
#
# - using plotImage.sigActiveImageChanged.connect(plotHisto)
# +
from silx.math.histogram import Histogramnd
def computeIntensityDistribution():
"""Get the active image and compute the image
intensity distribution"""
# By inheriting from PlotAction, we get access to attribute
# self.plot
# which is a reference to the PlotWindow
activeImage = plotImage.getActiveImage()
if activeImage is not None:
histo, w_histo, edges = Histogramnd(activeImage[0].flatten(),
n_bins=256,
histo_range=[0,256])
from silx.gui.plot import Plot1D
plotHistogram = Plot1D()
plotHistogram.addCurve(range(256),
histo,
legend='pixel intensity')
plotHistogram.show()
# -
plotImage=Plot2D()
plotImage.sigActiveImageChanged.connect(computeIntensityDistribution)
plotImage.addImage(image, origin=(0, 0), legend='lena')
# +
from silx.gui.plot.PlotActions import PlotAction
from silx.math.histogram import Histogramnd
from silx.gui.plot import Plot1D
class ComputeHistogramAction(PlotAction):
"""Computes the intensity distribution on the current image
:param plot: :class:`.PlotWidget` instance on which to operate
:param parent: See :class:`QAction`
"""
def __init__(self, plot, parent=None):
PlotAction.__init__(self,
plot,
icon='shape-circle',
text='pixels intensity',
tooltip='Compute image intensity distribution',
triggered=self.computeIntensityDistribution,
checkable=True,
parent=parent)
self.plotHistogram=Plot1D()
self.plot.sigActiveImageChanged.connect(self.update)
def update(self):
# By inheriting from PlotAction, we get access to attribute
# self.plot
# which is a reference to the PlotWindow
activeImage = self.plot.getActiveImage()
if activeImage is not None:
histo, w_histo, edges = Histogramnd(activeImage[0].flatten(),
n_bins=256,
histo_range=[0,256])
self.plotHistogram.addCurve(range(256),
histo,
legend='pixel intensity')
def computeIntensityDistribution(self):
"""Get the active image and compute the image intensity distribution"""
if self.isChecked():
self.update()
self.plotHistogram.show()
else:
self.plotHistogram.hide()
# -
plotImage=Plot2D()
myaction=ComputeHistogramAction(plotImage)
toolBar=plotImage.toolBar()
toolBar.addAction(myaction)
plotImage.addImage(image, origin=(0, 0), legend='ascent')
plotImage.show()
plotImage.addImage(image/2.0, origin=(0, 0), legend='ascent')
# Note : This feature has been added into silx as the 'intensityHistoAction'
#
# Some code to access it :
# +
import scipy.misc
image=scipy.misc.ascent()
plotImage=Plot2D()
plotImage.addImage(image, scale=(1, 1))
plotImage.getIntensityHistogramAction().setVisible(True)
plotImage.show()
# -
# # For information : the class diagram of the Plot module
# 
| silx/plot/solution/PlotInteraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 02_02 Preparing Data for Sentiment Analysis
#Import the movie reviews corpus
with open("Movie-Reviews.txt", 'r') as fh:
reviews = fh.readlines()
print(reviews[:2])
# ## 02_03 Finding Sentiments by Review
# +
#install textblob if not already installed using "pip install -U textblob"
from textblob import TextBlob
print('{:40} : {:10} : {:10}'.format("Review", "Polarity", "Subjectivity") )
for review in reviews:
#Find sentiment of a review
sentiment = TextBlob(review)
#Print individual sentiments
print('{:40} : {: 01.2f} : {:01.2f}'.format(review[:40]\
, sentiment.polarity, sentiment.subjectivity) )
# -
# ## 02_04 Summarizing Sentiment
# +
#Categorize Polarity into Positive, Neutral or Negative
labels = ["Negative", "Neutral", "Positive"]
#Initialize count array
values =[0,0,0]
#Categorize each review
for review in reviews:
sentiment = TextBlob(review)
#Custom formula to convert polarity
# 0 = (Negative) 1 = (Neutral) 2=(Positive)
polarity = round(( sentiment.polarity + 1 ) * 3 ) % 3
#add the summary array
values[polarity] = values[polarity] + 1
print("Final summarized counts :", values)
import matplotlib.pyplot as plt
#Set colors by label
colors=["Green","Blue","Red"]
print("\n Pie Representation \n-------------------")
#Plot a pie chart
plt.pie(values, labels=labels, colors=colors, \
autopct='%1.1f%%', shadow=True, startangle=140)
plt.axis('equal')
plt.show()
# -
| Sentiment Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # House Price prediction with AutoML (Accelerated Data Science Python SDK)
# +
# Load a number of libraries, including the Accelerated Data Science DatasetFactory and initialize it.
import pandas as pd
import logging
import numpy as np
import ads
import oci
from oci.data_science import DataScienceClient
from ads.dataset.factory import DatasetFactory
# OCI ADS API
ads.set_auth(auth='resource_principal')
resource_principal = oci.auth.signers.get_resource_principals_signer()
dsc = DataScienceClient(config={},signer=resource_principal)
# +
# Load the House Price data set using the ADS library.
# This library is capable of loading data from a variety of sources, including OCI Object Storage,
# Autonomous Database (ADW and ATP), other Oracle databases, Hadoop, S3, and many more.
# In this case the file is simply loaded from the block storage of the OCI Data Science service
# itself.
logging.basicConfig(format='%(levelname)s:%(message)s', level=logging.ERROR)
ds = DatasetFactory.open("housesales.csv", target="SalePrice")
# +
# Automatic Data Preparation with ADS:
# Last time we investigated every feature to decide whether it needed to be included in the model
# training. We manually had to check for things like number of unique values and missing values.
# Instead of doing this manually, let's ask ADS for suggestions on what changes to make to the
# data set.
# The following will show a summary of the recommendations. For example, ADS can recommend how
# to handle missing values. And it will recommend what to do with columsn that appear identifiers.
# Note that this can take up to a minute (depending on the shape you've chosen).
suggestion_df = ds.suggest_recommendations()
# +
# Imagine that we decide to simply take the recommendations made by ADS. Before we run the
# transformations, let's have a look at the original situation.
# For example, ADS suggests that we drop the columns Id, Alley, FireplaceQu, et ceters.
# ADS also suggests that we set default values for missing values in LotFrontage, MasVnrArea,
# et cetera.
# Now, let's apply the recommended transformations.
# We will also visualize the transformations.
ds_transformed = ds.auto_transform()
ds_transformed.visualize_transforms()
# +
# Let's visualize the end result.
# We see that there are now only 74 features left.
ds_transformed.summary()
# +
# AutoML:
# Now let's move on to building the model.
# We will run an experiment in which we create 3 models:
# 1: Linear Regression manually configured
# 2: Ridge Regression manually configured
# 3: AutoML
# Once we've trained the models, we will compare their performance.
#
# Preparation:
# 1. Resolve the skew in the target SalePrice by doing a log operation.
ds_transformed = ds_transformed.assign_column('SalePrice', lambda x: np.log(x))
# 2. Convert categorical attributes to numeric attributes
from ads.dataset.label_encoder import DataFrameLabelEncoder
ds_encoded = DataFrameLabelEncoder().fit_transform(ds_transformed.to_pandas_dataframe())
# 3. Separate the dataset in target and input features and split into test and train.
y = ds_encoded['SalePrice']
X = ds_encoded.drop(['SalePrice'], axis=1)
ds_done = DatasetFactory.open(ds_encoded, target="SalePrice")
train, test = ds_done.train_test_split(test_size=0.2)
# +
# First train two models manually (without AutoML): Linear Regression and Ridge Regression.
# We'll use these later to compare their performance against the model created by AutoML.
from sklearn.linear_model import Ridge, Lasso, LinearRegression
from ads.common.model import ADSModel
X = train.X.copy()
y = train.y.copy()
lr = LinearRegression()
rr = Ridge(alpha=13.5)
lr_model_fit = lr.fit(X, y)
rr_model_fit = rr.fit(X, y)
lr_model = ADSModel.from_estimator(lr_model_fit, name="LinearRegression")
rr_model = ADSModel.from_estimator(rr_model_fit, name="Ridge")
# +
# Now, let's see how well AutoML can perform the same task.
# AutoML will evaluate several algorithms in combination with different values for their
# hyperparameters.
# Note the time parameter of 160 seconds, this means we allow AutoML this amount of time to find the best solution.
from ads.automl.provider import OracleAutoMLProvider
from ads.automl.driver import AutoML
ml_engine = OracleAutoMLProvider(n_jobs = -1) # n_jobs = -1 means that we are not going to limit the resources and let the AutoML to do this for us and take as much as possible
automl = AutoML(train, provider=ml_engine)
model, baseline = automl.train(time_budget=160, random_state=42)
# +
# Have a look at the top candidate that was selected by AutoML. What algorithm is it?
# What are the hyperparameters it's using?
#
# Now, let's compare the performance of the manual and AutoML models
from ads.evaluations.evaluator import ADSEvaluator
evaluator = ADSEvaluator(test, models=[model, baseline, lr_model, rr_model], training_data=train)
evaluator.metrics
# +
# Conclusion:
# The models have been compared using various metrics.
# The best performing metrics are displayed in green.
#
# For example, look at the Mean Square Error, this is a measure of how close the predicted values
# match the actual values. The value of the AutoML model is significantly lower than that of the
# manually configured models, meaning that the AutoML model is performing better.
# The same is the case for the other metrics.
#
| oci-library/oci-hol/oci-datascience/introduction-to-data-science/linear-regression/files/lab100-bonus-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
"""
1. Plot a pie chart for columns: ‘cyl’ and ‘model’ form the mtcars.csv data frame
"""
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
mtcars = pd.read_csv("C:\\Users\\black\\Desktop\\PyforDS\\datasets\\mtcars.csv")
# +
labels = mtcars['model']
sizes = mtcars['cyl']
plt.pie(sizes, labels=labels)
plt.show()
| matplotlib_pie_chart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Apparel Recommendations usin Convolutional Neural Network
# ## Get the feature vectors of all apparel images
# <pre>
# Running this cell will take time, you can skip running this cell. you can download the feature vectors from given link
# 16k_data_cnn_features.npy: https://drive.google.com/open?id=0BwNkduBnePt2c1BkNzRDQ1dOVFk
# bottleneck_features_cnn.npy : https://drive.google.com/open?id=0BwNkduBnePt2ODRxWHhUVzIyWDA
# </pre>
import numpy as np
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dropout, Flatten, Dense
from keras import applications
from sklearn.metrics import pairwise_distances
import matplotlib.pyplot as plt
import requests
from PIL import Image
import pandas as pd
import pickle
# +
# https://gist.github.com/fchollet/f35fbc80e066a49d65f1688a7e99f069
# https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html
# dimensions of our images.
img_width, img_height = 224, 224
top_model_weights_path = 'bottleneck_fc_model.h5'
train_data_dir = 'images2/'
nb_train_samples = 16042
epochs = 50
batch_size = 1
def save_bottlebeck_features():
asins = []
datagen = ImageDataGenerator(rescale=1. / 255)
# build the VGG16 network
model = applications.VGG16(include_top=False, weights='imagenet')
generator = datagen.flow_from_directory(
train_data_dir,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode=None,
shuffle=False)
for i in generator.filenames:
asins.append(i[2:-5])
bottleneck_features_train = model.predict_generator(generator, nb_train_samples // batch_size)
bottleneck_features_train = bottleneck_features_train.reshape((16042,25088))
np.save(open('workshop/models/16k_data_cnn_features.npy', 'wb'), bottleneck_features_train)
np.save(open('workshop/models/16k_data_cnn_feature_asins.npy', 'wb'), np.array(asins))
save_bottlebeck_features()
# -
# # load the extracted features
bottleneck_features_train = np.load('workshop/models/16k_data_cnn_features.npy')
asins = np.load('workshop/models/16k_data_cnn_feature_asins.npy')
# ## get the most similar apparels using euclidean distance measure
data = pd.read_pickle('workshop/pickels/16k_apperal_data_preprocessed')
df_asins = list(data['asin'])
asins = list(asins)
# +
from IPython.display import display, Image, SVG, Math, YouTubeVideo
def get_similar_products_cnn(doc_id, num_results):
doc_id = asins.index(df_asins[doc_id])
pairwise_dist = pairwise_distances(bottleneck_features_train, bottleneck_features_train[doc_id].reshape(1,-1))
indices = np.argsort(pairwise_dist.flatten())[0:num_results]
pdists = np.sort(pairwise_dist.flatten())[0:num_results]
for i in range(len(indices)):
rows = data[['medium_image_url','title']].loc[data['asin']==asins[indices[i]]]
for indx, row in rows.iterrows():
display(Image(url=row['medium_image_url'], embed=True))
print('Product Title: ', row['title'])
print('Euclidean Distance from input image:', pdists[i])
print('Amazon Url: www.amzon.com/dp/'+ asins[indices[i]])
get_similar_products_cnn(12566, 10)
# +
# with GPU => per image 3.5/20 (0.175 sec), for whole data set its taking around 40min
# with CPU => per image 12.3/20 (0.615 sec), for whole data set its taking around 160min
| image_similarity_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NetCDF files
#
# NetCDF is a binary storage format for many different kinds of rectangular data. Examples include atmosphere and ocean model output, satellite images, and timeseries data. NetCDF files are intended to be device independent, and the dataset may be queried in a fast, random-access way. More information about NetCDF files can be found [here](http://www.unidata.ucar.edu/software/netcdf/). The [CF conventions](http://cfconventions.org) are used for storing NetCDF data for earth system models, so that programs can be aware of the coordinate axes used by the data cubes.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import cartopy
#import cmocean.cm as cmo
import netCDF4
# -
# ### Sea surface temperature example
#
# An example NetCDF file containing monthly means of sea surface temperature over 160 years can be found [here](http://www.esrl.noaa.gov/psd/data/gridded/data.noaa.ersst.v4.html). We'll use the NetCDF4 package to read this file, which has already been saved into the `data` directory.
nc = netCDF4.Dataset('../data/sst.mnmean.v4.nc')
nc['sst'].shape
print(nc)
# The representation of the object shows some of the attributes of the netCDF file. The final few lines show the dimensions and the variable names (with corresponding dimensions). Another representation of the file can be seen using the `ncdump` command. This is similar to the output of the command (at a command-line prompt, not within python)
#
# $ ncdump -h ../data/sst.mnmean.v4.nc
#
# netcdf sst.mnmean.v4 {
# dimensions:
# lon = 180 ;
# lat = 89 ;
# nbnds = 2 ;
# time = UNLIMITED ; // (1946 currently)
# variables:
# float lat(lat) ;
# lat:units = "degrees_north" ;
# lat:long_name = "Latitude" ;
# lat:actual_range = 88.f, -88.f ;
# lat:standard_name = "latitude" ;
# lat:axis = "Y" ;
# lat:coordinate_defines = "center" ;
# float lon(lon) ;
# lon:units = "degrees_east" ;
# lon:long_name = "Longitude" ;
# lon:actual_range = 0.f, 358.f ;
# lon:standard_name = "longitude" ;
# lon:axis = "X" ;
# lon:coordinate_defines = "center" ;
# double time_bnds(time, nbnds) ;
# time_bnds:long_name = "Time Boundaries" ;
# double time(time) ;
# time:units = "days since 1800-1-1 00:00:00" ;
# time:long_name = "Time" ;
# time:delta_t = "0000-01-00 00:00:00" ;
# time:avg_period = "0000-01-00 00:00:00" ;
# time:prev_avg_period = "0000-00-07 00:00:00" ;
# time:standard_name = "time" ;
# time:axis = "T" ;
# time:actual_range = 19723., 78923. ;
# float sst(time, lat, lon) ;
# sst:long_name = "Monthly Means of Sea Surface Temperature" ;
# sst:units = "degC" ;
# sst:var_desc = "Sea Surface Temperature" ;
# sst:level_desc = "Surface" ;
# sst:statistic = "Mean" ;
# sst:missing_value = -9.96921e+36f ;
# sst:actual_range = -1.8f, 33.95f ;
# sst:valid_range = -5.f, 40.f ;
# sst:dataset = "NOAA Extended Reconstructed SST V4" ;
# sst:parent_stat = "Individual Values" ;
#
# // global attributes:
# :history = "created 10/2014 by CAS using NCDC\'s ERSST V4 ascii values" ;
# [....and so on....]
# You can access terminal commands from with the Jupyter notebook by putting "!" first:
# !ncdump -h ../data/sst.mnmean.v4.nc
# ### Mapping the netcdf object to the python object
#
# We can query the data within the NetCDF file using the NetCDF object. The structure of the object (the composition of the methods and attributes) is designed to mirror the data structure in the file. See how these queries give the same information as the textual representation above.
# `Global` attributes of the file
nc.history
# Variables are stored in a dictionary
nc.variables['lon'] # this is a variable object, just a pointer to the variable. NO DATA HAS BEEN LOADED!
# Variable objects also have attributes
nc.variables['lon'].units
# we can also query the dimensions
nc.dimensions['lon']
# to find the length of a dimension, do
len(nc.dimensions['lon'])
# A list of the dimensions can be found by looking at the keys in the dimensions dictionary
nc.dimensions.keys()
# Same for variables
nc.variables.keys()
# Let's take a look at the main 3D variable
nc['sst'] # A shorthand for nc.variables['sst']
nc['sst'].units
# ---
# ### *Exercise*
#
# > Inspect the NetCDF object.
#
# > 1. What are the units of the time variable?
# > 1. What are the dimensions of the latitude variable?
# > 1. What is the length of the latitude dimension?
#
# ---
# We can extract data from the file by indexing:
# This reads in the data so be careful how much you read in at once
lon = nc['lon'][:]
lat = nc['lat'][:]
sst = nc['sst'][0] # same as nc['sst'][0, :, :], gets the first 2D time slice in the series.
# Extract the time variable using the convenient num2date, which converts from time numbers to datetime objects
time = netCDF4.num2date(nc['time'][:], nc['time'].units)
time
# There are lots of operations you can do with `datetime` objects, some of which you've used before, possibly in other packages like `pandas`.
#
# For example, you can find the difference between two datetimes. This is given in a `datetime.timedelta` object.
time[2] - time[1]
# You can also specify the time unit of measurement you get out of this difference:
(time[2] - time[1]).days
# Note that asking for the number of seconds (from 0 to 60 for that datetime object) is different than asking for the number of total seconds (total number of seconds in the time measurement):
(time[2] - time[1]).seconds
(time[2] - time[1]).total_seconds()
# ---
# ### *Exercise*
#
# > Practice with `datetime`:
#
# > 1. Find the number of days between several successive `datetimes` in the `time` variable. You will need to extract this number from the `timedelta` object.
# > 1. One way you want present the date and time contained within a `datetime` object is with:
# time[0].isoformat()
# Test this, and also try using the following to display your datetime as a string:
# time[0].strftime([input formats])
# where you choose time formats from the options which can be seen at `strftime.org`.
#
# ---
# Let's use the data that we have read in to make a plot.
# +
proj = cartopy.crs.Mollweide(central_longitude=180)
pc = cartopy.crs.PlateCarree()
fig = plt.figure(figsize=(14,6))
ax = fig.add_subplot(111, projection=proj)
mappable = ax.contourf(lon, lat, sst, 100, cmap=plt.get_cmap('magma'), transform=pc)
# -
# ---
# ### *Exercise*
#
# > Finish the plot above. Add:
# > * Land
# > * Colorbar with proper label and units
# > * Title with nicely formatting date and time
#
# ---
# ### THREDDS example. Loading data from a remote dataset.
#
# The netCDF library can be compiled such that it is 'THREDDS enabled', which means that you can put in a URL instead of a filename. This allows access to large remote datasets, without having to download the entire file. You can find a large list of datasets served via an OpenDAP/THREDDs server [here](http://apdrc.soest.hawaii.edu/data/data.php).
#
# Let's look at the ESRL/NOAA 20th Century Reanalysis – Version 2. You can access the data by the following link (this is the link of the `.dds` and `.das` files without the extension.):
loc = 'http://apdrc.soest.hawaii.edu/dods/public_data/Reanalysis_Data/esrl/daily/monolevel/V2c/cprat'
nc_cprat = netCDF4.Dataset(loc)
nc_cprat['cprat'].long_name
time = netCDF4.num2date(nc_cprat['time'][:], nc_cprat['time'].units) # convert to datetime objects
time
cprat = nc_cprat['cprat'][-1] # get the last time, datetime.datetime([year], 12, 31, 0, 0)
lon = nc_cprat['lon'][:]
lat = nc_cprat['lat'][:]
# +
proj = cartopy.crs.Sinusoidal(central_longitude=180)
fig = plt.figure(figsize=(14,6))
ax = fig.add_subplot(111, projection=proj)
ax.coastlines(linewidth=0.25)
mappable = ax.contourf(lon, lat, cprat, 20, cmap=cmo.tempo, transform=pc)
ax.set_title(time[-1].isoformat()[:10])
fig.colorbar(mappable).set_label('%s' % nc_cprat['cprat'].long_name)
# -
# ---
# ### *Exercise*
#
# > Pick another [variable](http://apdrc.soest.hawaii.edu/dods/public_data/Reanalysis_Data/esrl/daily/monolevel) from this dataset. Inspect and plot the variable in a similar manner to precipitation.
#
# > Find another dataset on a THREDDS server at SOEST (or elsewhere), pick a variable, and plot it.
#
# ---
# ### Creating NetCDF files
#
# We can also create a NetCDF file to store data. It is a bit of a pain. Later we will see an easier way to do this.
# +
from matplotlib import tri
Ndatapoints = 1000
Ntimes = 20
Nbad = 200
xdata = np.random.rand(Ndatapoints)
ydata = np.random.rand(Ndatapoints)
time = np.arange(Ntimes)
# create a progressive wave
fdata = np.sin((xdata+ydata)[np.newaxis, :]*5.0 +
time[:, np.newaxis]/3.0)
# remove some random 'bad' data.
idx = np.arange(fdata.size)
np.random.shuffle(idx)
fdata.flat[idx[:Nbad]] = np.nan
ygrid, xgrid = np.mgrid[0:1:60j, 0:1:50j]
fgrid = np.ma.empty((Ntimes, 60, 50), 'd')
# interpolate
for n in range(Ntimes):
igood = ~np.isnan(fdata[n])
t = tri.Triangulation(xdata[igood], ydata[igood])
interp = tri.LinearTriInterpolator(t, fdata[n][igood])
fgrid[n] = interp(xgrid, ygrid)
# create netCDF file
nc = netCDF4.Dataset('foo.nc', 'w')
nc.author = 'Me'
nc.createDimension('x', 50)
nc.createDimension('y', 60)
nc.createDimension('time', None) # An 'unlimited' dimension.
nc.createVariable('f', 'd', ('time', 'y', 'x'))
nc.variables['f'][:] = fgrid
nc.variables['f'].units = 'meters sec-1'
nc.createVariable('x', 'd', ('x',))
nc.variables['x'][:] = xgrid[0, :]
nc.variables['x'].units = 'meters'
nc.createVariable('y', 'd', ('y',))
nc.variables['y'][:] = ygrid[:, 0]
nc.variables['y'].units = 'meters'
nc.createVariable('time', 'd', ('time',))
nc.variables['time'][:] = time
nc.variables['time'].units = 'seconds'
nc.close()
# -
nc = netCDF4.Dataset('foo.nc')
nc
# ### See also
#
# - [Xarray](http://xarray.pydata.org/en/stable/): NetCDF + PANDAS + CF conventions. Awesome.
# - [pygrib](https://github.com/jswhit/pygrib): Reading GRIB files.
# - [ncview](http://meteora.ucsd.edu/~pierce/ncview_home_page.html): Not python, but a very useful NetCDF file viewer.
# # `xarray`
#
# `xarray` expands the utility of the time series analysis package `pandas` into more than one dimension. It is actively being developed so some functionality isn't yet available, but for certain analysis it is very useful.
import xarray as xr
# In the previous material, we used `netCDF` directly to read in a data file, then access the data:
# +
nc = netCDF4.Dataset('../data/sst.mnmean.v4.nc')
print(nc['sst'].shape)
# -
# However, as was pointed out in class, in this approach if we want to pull out the sea surface temperature data at a particular time, we need to first know which time index that particular time corresponds to. How can we find this?
#
# First we convert the time numbers from the file into datetimes, like before:
# Extract the time variable using the convenient num2date
time = netCDF4.num2date(nc['time'][:], nc['time'].units)
time
time.fill_value = 9
# Say we want to search for the time index corresponding to May 1, 1954.
# +
from datetime import datetime
date = datetime(1954, 5, 1, 0, 0)
# -
# Now we search for the time index:
tind = np.where(time==date)[0][0]
print(tind)
# Great! So the time index we want is 1204. We can now make our sea surface temperature plot:
# +
proj = cartopy.crs.Mollweide(central_longitude=180)
pc = cartopy.crs.PlateCarree()
fig = plt.figure(figsize=(14,6))
ax = fig.add_subplot(111, projection=proj)
mappable = ax.contourf(nc['lon'][:], nc['lat'][:], nc['sst'][tind], 100, cmap=plt.get_cmap('inferno'), transform=pc)
# -
# What if instead we want the index corresponding to May 23, 1954
date = datetime(1954, 5, 23, 0, 0)
np.where(time==date)
# What is the problem here? There is no data at that exact time.
#
# So what should we do?
#
# A few options:
# index of date that minimizes time between model times and desired date
tidx = np.abs(time - date).argmin()
tidx
time[tidx]
np.where(time<=date)[0][-1]
# So, you can do this but it's a little annoying and takes extra effort.
# Now let's access this data using a different package called `xarray`:
ds = xr.open_dataset('../data/sst.mnmean.v4.nc') # similar way to read in — also works for nonlocal data addresses
ds
# Now we can search for data in May 1954:
ds['sst'].sel(time=slice('1954-05','1954-05'))
# Or we can search for the nearest output to May 23, 1954:
ds['sst'].sel(time='1954-05-23', method='nearest')
# Let's plot it!
# +
sst = ds['sst'].sel(time='1954-05-23', method='nearest')
fig = plt.figure(figsize=(14,6))
ax = fig.add_subplot(111, projection=proj)
mappable = ax.contourf(nc['lon'][:], nc['lat'][:], sst, 10, cmap=plt.get_cmap('inferno'), transform=pc)
# -
# Note that you can also just plot against the included coordinates with built-in convenience functions (this is analogous to `pandas` which was for one dimension):
fig = plt.figure(figsize=(14,6))
ax = fig.add_subplot(111, projection=proj)
sst.plot(transform=pc) # the plot's projection
# ## GroupBy
#
# Like in `pandas`, we can use the `groupby` method to do some neat things. Let's group by season and save a new file.
seasonal_mean = ds.groupby('time.season').mean('time')
seasonal_mean
# Do you remember how many lines of code were required to save a netCDF file from scratch? It is straight-forward, but tedious. Once you are working with data using `xarray`, you can save new, derived files very easily from your data array:
fname = 'test.nc'
seasonal_mean.to_netcdf(fname)
d = netCDF4.Dataset(fname)
d
# ---
# ### *Exercise*
#
# > Plot the difference between summer and winter mean sea surface temperature.
#
# ---
| materials/Module 7/7_netcdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 输入一个pandas dataframe(t-sne数据),看聚类效果
# #### 导入所需包
import json
from pyecharts import Scatter3D
# #### 画图方法
def echart3D_tsne(pddata,label,x_title,y_title,z_title):
scatter3D = Scatter3D("T-SNE", width=1200, height=600)
for lab in set(pddata[label]):
data=pddata[pddata[label]==lab]
jsondata = json.loads(data[[x_title,y_title,z_title]].to_json(orient='values'))
scatter3D.add(str(lab),
jsondata,
xaxis3d_name='TSNE1',
yaxis3d_name='TSNE2',
zaxis3d_name='TSNE3',)
return scatter3D
# #### 一个例子
# +
import pandas as pd
import random
data = [
[random.randint(0, 2),
random.randint(0, 100),
random.randint(0, 100),
random.randint(0, 100)] for _ in range(10)
]
pddata = pd.DataFrame(data,columns=['label','t1','t2','t3'])
print(pddata)
s3d=echart3D_tsne(pddata,'label','t1','t2','t3')
s3d
# -
# ## GC test
# +
def one_sample_depth(file):
'''
:param file:
:return:
'''
data = []
with open(file, 'r') as f:
line = f.readline()
line = f.readline()
while line:
cells = line.strip('\n').split(',')
name_eles = cells[0].split('-')
data.append(['{}:{}-{}'.format(*name_eles), int(cells[1])])
line = f.readline()
pd_data = pd.DataFrame(data, columns=['loc', 'depth'])
return pd_data
def get_name(file):
return (os.path.basename(file).split('.'))[0]
def echart3D(e3d,pddata,label,x_title,y_title,z_title):
# print(label)
jsondata = json.loads(pddata[[x_title,y_title,z_title]].to_json(orient='values'))
e3d.add(str(label),
jsondata,
xaxis3d_name='GC',
yaxis3d_name='LENGTH',
zaxis3d_name='DEPTH',)
# +
import os
import pandas as pd
files=['demo_data/controls/FH_1923.stat.csv',
'demo_data/controls/FH_1925.stat.csv',
'demo_data/controls/FH_1926.stat.csv',
# 'demo_data/controls/FH_2128.stat.csv',
# 'demo_data/controls/FH_2182.stat.csv',
# 'demo_data/controls/FH_2250.stat.csv'
]
gc_length_file = 'demo_data/controls/bed_gc.csv'
glf=pd.read_csv(gc_length_file)
scatter3D = Scatter3D("GC", width=1200, height=600)
# -
for file in files:
depth = one_sample_depth(file)
depth['depth'] = depth['depth']/depth['depth'].mean()
test = pd.merge(glf,depth,how='outer',on='loc',left_index=True)
label = get_name(file)
# print(test.head())
echart3D(scatter3D,test,label,'gc','length','depth')
scatter3D
depth = one_sample_depth(files[0])
test=pd.merge(glf,depth,how='outer',on='loc',left_index=True)
glf.head()
| pyechart-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_lusa
# language: python
# name: venv_lusa
# ---
# +
from IPython.display import display, Markdown, HTML
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import missingno
# -
sns.set()
pd.options.mode.chained_assignment = None
# # Lendo os dados
# A base de dados utilizada está no [formato csv](https://www.linkedin.com/pulse/lendo-arquivos-csv-com-pandas-rog%C3%A9rio-guimar%C3%A3es-de-campos-j%C3%BAnior/)
#
df = pd.read_csv("https://raw.githubusercontent.com/nubank/diversidados-curso-ds/master/iniciante/base_imoveis.csv")
# Olhando conteúdo das 5 primeiras linhas
df.head()
# Olhando nome das colunas
df.columns
# Olhando informações gerais sobre a base
df.info()
# Plotando matriz de nulos
missingno.matrix(df)
# Acima, vimos que não temos nenhum nulo na base! **Mas será que temos na base outros valores que poderiam ser considerados como nulo?**
# Imprimindo tipo de cada coluna
df.dtypes
target = "preco_dolares"
# # Explorando as variáveis
# <img src="imgs/tipos_variaveis.png" alt="tipos variaveis" width="100%"/>
# Para printar numero de linhas por cada valor possível da coluna `tem_porao`: `df["tem_porao"].value_counts()`
df["tem_porao"].value_counts()
# ## Categoricas
# ### Cardinalidade
cardinalidades = [] # lista que vai guardar valor das cardinalidades
vars_categoricas_raw = [] # lista que vai guardar nomes das variaveis categoricas
for i, row in df.dtypes.reset_index().iterrows():
nome_var = row["index"]
tipo_var = row[0]
if tipo_var==object:
cardinalidade = df[nome_var].nunique() # Funcao nunique retorna número de valores diferentes na coluna
cardinalidades.append(cardinalidade) # Adc valor na lista
vars_categoricas_raw.append(nome_var) # Adc nome na lista
df_cardinalidades = pd.DataFrame({"Variavel": vars_categoricas_raw,
"Cardinalidade": cardinalidades,}) # Criando uma tabela com valores calculados
df_cardinalidades
# Plotando cardinalidades
sns.barplot(x="Cardinalidade", y="Variavel", data=df_cardinalidades, orient="h", palette="coolwarm")
plt.title("Cardinalidade das variaveis categoricas");
# Colocando na escala logaritmica
g = sns.barplot(x="Cardinalidade", y="Variavel", data=df_cardinalidades, orient="h", palette="coolwarm")
g.set_xscale("log")
plt.title("Cardinalidade das variaveis categoricas");
# ### Rank e distribuição
for nome_var in vars_categoricas_raw:
# Contando numero de casos em cada categoria e ordenando pelas classes mais frequentes
df_rank = df.groupby(nome_var).agg({"id": "count"}) \
.sort_values("id", ascending=False)
# Calculando porcentagem em cada categoria para ver distribuição
df_rank["% ids"] = df_rank["id"]/df_rank["id"].sum()*100
display(df_rank)
# ### Filtrando variáveis
vars_categoricas_raw
vars_categoricas_raw.pop(0)
vars_categoricas_raw
categoricas_nominais = ['orla_mar', 'tem_porao', 'teve_reforma']
categoricas_ordinais = [ 'nota_vista', 'nota_condicao',]
# ## Numericas
# O método `.describe()` já calcula algumas métricas comuns nas variáveis numéricas
df.describe()
# Plot de distribuição tradicional
vars_numericas_raw = []
for i, row in df.dtypes.reset_index().iterrows():
nome_var = row["index"]
tipo_var = row[0]
if tipo_var in (np.float64, np.int64):
print(nome_var)
vars_numericas_raw.append(nome_var)
fig, ax = plt.subplots()
fig.set_size_inches(10, 6)
sns.distplot(df[nome_var], hist=True, kde=False, norm_hist=False, ax=ax)
plt.show()
for nome_var in vars_numericas_raw:
print(nome_var)
fig, ax = plt.subplots()
fig.set_size_inches(10, 6)
sns.boxplot(df[nome_var], ax=ax)
plt.show()
# ### Filtrando variáveis
vars_numericas_raw
vars_numericas_raw.pop(0)
vars_numericas_raw.pop(0)
vars_numericas_raw
# # Separando a base
# <img src="imgs/splits.png" alt="separacao da base" width="80%"/>
#
# Não existe um número mágico, alguns indicam 60%-20%-20%, e vamos usar essa proporção
vars_categoricas_raw
vars_numericas_raw
# Transformando campo de data pois estava como string
df["data"] = pd.to_datetime(df["data"], format="%Y-%m-%d")
# Contando numero de imoveis por dia
df_datas = df.groupby("data").agg({"id": "count"})
df_datas["media_data"] = df_datas.rolling(7).mean()
df_datas = df_datas.reset_index()
# Plotando número de imoveis por dia ao longo do tempo
fig, ax = plt.subplots()
fig.set_size_inches(10, 6)
ax.plot(df_datas["data"], df_datas["media_data"])
plt.title("Numero de imoveis por data");
df = df.sort_values("data").reset_index(drop="index")
# ## Divisão no tempo
# Usando [`train_test_split`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) da biblioteca sklearn.
# Primeiro, fazemos a divisão no tempo, deixando 20% para nossa base de validação
treino_teste, validacao = train_test_split(df, test_size=.2, random_state=0, shuffle=False)
# Verificando se a divisão no tempo funcionou
treino_teste.data.describe()
validacao.data.describe()
# ## Divisão no espaço
# Agora, dos 80% iniciais, vamos deixar 25% para teste e 75% para treino, misturando os dados para fazer apenas a divisão no espaço
treino, teste = train_test_split(treino_teste, test_size=.25, random_state=0, shuffle=True)
treino.data.describe()
teste.data.describe()
# ## Check de tamanho das bases
treino.shape, teste.shape, validacao.shape
# # Tratando variáveis categóricas
# ## Ordinais
categoricas_ordinais
# Olhando categorias existentes
treino["nota_vista"].unique()
# Coloque os valores que representam a ordem das categorias
nota_vista_map = {'Vista completamente bloqueada': 1,
'Vista parcialmente bloqueada': 2,
'Vista quase sem bloqueios': 3,
'Vista boa': 4,
'Excelente - vista sem bloqueios': 5}
treino["nota_vista_encoded"] = treino["nota_vista"].map(nota_vista_map)
teste["nota_vista_encoded"] = teste["nota_vista"].map(nota_vista_map)
validacao["nota_vista_encoded"] = validacao["nota_vista"].map(nota_vista_map)
# Faça o mesmo para variável `nota_condicao`
treino["nota_condicao"].unique()
nota_condicao_map = {'Excelente estado': 5,
'Bom estado' : 4,
'Pronto para morar' : 3,
'Mau estado - pequena reforma necessária' : 2,
'Mau estado - reforma completa necessária' : 1}
treino["nota_condicao_encoded"] = treino["nota_condicao"].map(nota_condicao_map)
teste["nota_condicao_encoded"] = teste["nota_condicao"].map(nota_condicao_map)
validacao["nota_condicao_encoded"] = validacao["nota_condicao"].map(nota_condicao_map)
# ## Nominais
categoricas_nominais
for var_n in categoricas_nominais:
# Pegando média do target no treino para cada categoria
var_map = treino.groupby(var_n).agg({target: "mean"}).to_dict()[target]
treino[f"{var_n}_encoded"] = treino[f"{var_n}"].map(var_map) # Aplicando no treino
teste[f"{var_n}_encoded"] = teste[f"{var_n}"].map(var_map) # Aplicando no teste
validacao[f"{var_n}_encoded"] = validacao[f"{var_n}"].map(var_map) # Aplicando na validacao
vars_categoricas = [x+"_encoded" for x in categoricas_nominais+categoricas_ordinais]
# # Tratando variáveis numéricas
# ## Normalizando
for var_n in vars_numericas_raw+vars_categoricas:
scaler = MinMaxScaler()
scaler.fit(treino[[var_n]]) # Usando treino como base para transformações
treino[f"{var_n}_scaled"] = scaler.transform(treino[[var_n]]).reshape(-1) # Aplicando no treino
teste[f"{var_n}_scaled"] = scaler.transform(teste[[var_n]]).reshape(-1) # Aplicando no teste
validacao[f"{var_n}_scaled"] = scaler.transform(validacao[[var_n]]).reshape(-1) # Aplicando na validacao
# # Organizando variáveis
# Criamos várias colunas novas com as variáveis transformadas, precisamos filtrar só as novas, abaixo, temos todas as colunas que existem agora
treino.columns
# Agora vamos pegar só as que transformamos
todas_variaveis = [x+"_scaled" for x in vars_numericas_raw+vars_categoricas]
todas_variaveis
# Salvando quais eram as variaveis originais (raw)
todas_variaveis_raw = vars_numericas_raw+vars_categoricas_raw
# Olhando variáveis transformadas pra checar se estão ok
validacao[todas_variaveis].head()
validacao[todas_variaveis_raw].head()
# # Modelinho teste
todas_variaveis
reg = DecisionTreeRegressor(random_state=0)
reg.fit(treino[vars_numericas_raw+vars_categoricas], treino[target])
predictions_teste = reg.predict(teste[vars_numericas_raw+vars_categoricas])
predictions_validacao = reg.predict(validacao[vars_numericas_raw+vars_categoricas])
r2_score(teste[target], predictions_teste), r2_score(validacao[target], predictions_validacao)
r2_score(teste[target], predictions_teste), r2_score(validacao[target], predictions_validacao)
| iniciante/Dataprep/Dataprep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# TODO: This needs to be updated to use mutation AA positions rather than residue position
from biopandas.pdb import PandasPdb
# pdf = PandasPdb().read_pdb('/home/pphaneuf/iML1515_GP/iML1515_GP/genes/b3926/b3926_protein/structures/GLPK_ECOLI_model1_clean.pdb')
pdf = PandasPdb().read_pdb('../data/pdb_files/KPYK1_ECOLI_model1_clean.pdb') # From iML1515
adf = pdf.df['ATOM']
cadf = adf[adf["atom_name"]=="CA"]
cadf = cadf.set_index("residue_number")
cadf = cadf[cadf["alt_loc"].isin(['A', ''])] # removing the 'B' residue alternative locations. I could also find the average between the two positions if necessary to account for both.
cadf.head()
# +
import pandas as pd
aa_mut_cnt_df = pd.read_csv("./pykF_pub_aa_muts.csv")
aa_mut_cnt_df.head()
# -
gff_df = pd.read_csv("./PykF_feats.csv")
gff_df = gff_df[~(gff_df.feature.isin(["Turn", "Helix", "Beta strand", "Barrel domain", "Alpha/beta domain"]))].copy()
gff_df.head()
aa_mut_cnt_df = aa_mut_cnt_df[aa_mut_cnt_df["color"] != '#CF000F'].copy() # filtering out all truncating mutations since currently not considering truncations for 3D clustering
# +
dist_aa_df = aa_mut_cnt_df.copy()
# don't care about about multiple muts on same AA for now
dist_aa_df = dist_aa_df[["AA position", "name"]]
dist_aa_df = dist_aa_df.drop_duplicates()
# display(dist_aa_df)
for _, r in gff_df.iterrows():
if r["feature"] != "Chain":
for aa in range(r["start"], r["end"] + 1):
dist_aa_df = dist_aa_df.append({"AA position":aa, "name": r["feature"] + " AA " + str(aa)}, ignore_index=True)
dist_aa_df["AA position"] = dist_aa_df["AA position"].astype(int)
# PykF using includes the initiator methionine.
dist_aa_df["residue index"] = dist_aa_df["AA position"] # - 1 # this really depends on the structure being used. Double check if the initiator methionine is included in structure.
dist_aa_df['coords'] = dist_aa_df.apply(lambda r:
[
cadf.at[r["residue index"], "x_coord"],
cadf.at[r["residue index"], "y_coord"],
cadf.at[r["residue index"], "z_coord"],
],
axis=1)
dist_aa_df.head()
# +
import numpy as np
def _get_dist_mat_label(r):
lbl = r["name"]
if ("SNP" in r["name"]) or ("INS" in r["name"]):
lbl = "mut AA " + str(r["AA position"])
return lbl
dist_aa_df['dist mat label'] = dist_aa_df.apply(lambda r: _get_dist_mat_label(r), axis=1)
# dist_aa_df = dist_aa_df.set_index("AA position")
dist_aa_df = dist_aa_df.set_index("dist mat label")
dist_aa_df.head()
# +
import numpy as np
def get_3D_res_dist(feat_res_1, feat_res_2):
dist = np.nan
feat_res_1_coords = dist_aa_df.at[feat_res_1, "coords"]
feat_res_2_coords = dist_aa_df.at[feat_res_2, "coords"]
dist = np.sqrt((feat_res_2_coords[0] - feat_res_1_coords[0])**2
+ (feat_res_2_coords[1] - feat_res_1_coords[1])**2
+ (feat_res_2_coords[2] - feat_res_1_coords[2])**2)
return dist
dist_mat = pd.DataFrame(np.nan, columns=dist_aa_df.index, index=dist_aa_df.index)
for ir in dist_mat.index:
for ic in dist_mat.columns:
dist_mat.at[ir, ic] = get_3D_res_dist(ir, ic)
dist_mat.head()
# -
df = gff_df[["feature", "color"]].drop_duplicates()
feat_to_color_d = dict()
for _, r in df.iterrows():
feat_to_color_d[r["feature"]] = r["color"]
feat_to_color_d
# +
AA_feat_dist_df = dist_mat[dist_mat.index.str.contains("mut")]
AA_feat_dist_df = AA_feat_dist_df.T
AA_feat_dist_df = AA_feat_dist_df[~(AA_feat_dist_df.index.str.contains("mut"))]
col_colors = []
for _, r in AA_feat_dist_df.iterrows():
col_colors.append(feat_to_color_d[r.name[:r.name.find(" AA")]])
# col_colors
# +
import seaborn as sns
import matplotlib
import matplotlib.pyplot as mpl
# %matplotlib inline
mpl.rcParams["figure.dpi"] = 200
mpl.rcParams['font.sans-serif'] = ["FreeSans"]
sns.set_context("paper")
sns.set_style("white")
sns.set(font="FreeSans")
df = AA_feat_dist_df.T
cm = sns.clustermap(
df,
# linewidths=0.1,
cbar_kws={'label': 'distance (Å)'},
cbar_pos=(0.05, 0.85, 0.03, 0.10),
yticklabels=True,
xticklabels=True,
# annot=True,
cmap="rocket_r",
col_colors=col_colors,
# figsize=(10,10)
)
# Fix for recent matplotlib/seaborn issue
cm.ax_heatmap.set_ylim(len(df)+0, -0)
# Below is for adjusting the sizes of the dendograms and heatmaps
H = 0.425
W = 1.7
y0_offset = 0.29
x0_offset = 0.01
hm = cm.ax_heatmap.get_position()
# mpl.setp(cm.ax_heatmap.yaxis.get_majorticklabels(),fontsize=6)
# mpl.setp(cm.ax_heatmap.xaxis.get_majorticklabels(),fontsize=6)
col_dendo = cm.ax_col_dendrogram.get_position()
cm.ax_col_dendrogram.set_position([
col_dendo.x0 + x0_offset,
col_dendo.y0,
col_dendo.width*W,
col_dendo.height*1
])
row_dendo = cm.ax_row_dendrogram.get_position()
cm.ax_row_dendrogram.set_position([
row_dendo.x0 + 0.015,
row_dendo.y0 + y0_offset,
row_dendo.width*1,
row_dendo.height*H
])
cm.ax_heatmap.set_position([
hm.x0 + x0_offset,
hm.y0 + y0_offset,
hm.width*W,
hm.height*H
])
col_color_pos = cm.ax_col_colors.get_position()
cm.ax_col_colors.set_position([
col_color_pos.x0 + 0.01,
col_color_pos.y0 + 0.01,
col_color_pos.width*W,
col_color_pos.height*H
])
# mpl.setp(cm.ax_heatmap.get_xticklabels(), rotation=45, ha="right", rotation_mode="anchor")
cm.ax_heatmap.set_ylabel('')
cm.ax_heatmap.set_xlabel('')
cm.ax_heatmap.set_xticks([])
cm.ax_heatmap.tick_params(axis='both', which='both', length=0)
# cm.fig.suptitle("3D distance between mutated AA and small active sites on GlpK", y=0.92, x=0.6)
# +
mut_aa_feat_prox_df = pd.DataFrame()
df = AA_feat_dist_df.T
for mut_aa, r in df.iterrows():
min_dist_feats = r[r==r.min()]
for feat_AA, min_dist in min_dist_feats.iteritems():
feat = feat_AA[:feat_AA.find(" AA ")]
mut_aa_feat_prox_df = mut_aa_feat_prox_df.append({"mut AA": mut_aa.replace("mut AA ", ''), "nearest feature": feat}, ignore_index=True)
mut_aa_feat_prox_df
mut_aa_feat_prox_df.to_pickle("./mut_aa_feat_prox_df.pkl")
# -
gene_mut_df = pd.read_pickle("./pykF_mut_df.pkl")
gene_mut_df.head()
mut_aa_feat_prox_df = mut_aa_feat_prox_df.set_index("mut AA")
mut_aa_feat_prox_df
# +
non_trunc_aa_mut_effect_df = pd.DataFrame(columns=["mutated AA", "functionally disruptive", "structurally disruptive", "nearest feature"])
for _, m in gene_mut_df.iterrows():
if m["AA range"][0] == m["AA range"][1]:
aa = m["AA range"][0]
for _, r in mut_aa_feat_prox_df[mut_aa_feat_prox_df.index==str(aa)].iterrows():
non_trunc_aa_mut_effect_df = non_trunc_aa_mut_effect_df.append({
"mutated AA": aa,
"functionally disruptive": m["conservation (SIFT)"],
"structurally disruptive": m["stability (ΔΔGpred)"],
"nearest feature": r["nearest feature"]},
ignore_index=True)
else:
for aa in m["AA range"]:
for _, r in mut_aa_feat_prox_df[mut_aa_feat_prox_df.index==str(aa)].iterrows():
non_trunc_aa_mut_effect_df = non_trunc_aa_mut_effect_df.append({
"mutated AA": aa,
"functionally disruptive": m["conservation (SIFT)"],
"structurally disruptive": m["stability (ΔΔGpred)"],
"nearest feature": r["nearest feature"]},
ignore_index=True)
non_trunc_aa_mut_effect_df["unknown"] = non_trunc_aa_mut_effect_df.apply(lambda r: True if ((r["functionally disruptive"]==False) & (r["structurally disruptive"]==False)) else False, axis=1)
non_trunc_aa_mut_effect_df.head()
# -
non_trunc_aa_mut_effect_df.to_pickle("./PykF_mut_res_df.pkl") # Since this also describes the frequency of mutated residues, using it to described freq mut res per feat.
# +
cnt_columns = ["unknown", "functionally disruptive", "structurally disruptive"]
feat_mut_eff_cnt_d = {f:{"functionally disruptive":0, "structurally disruptive":0, "unknown":0} for f in non_trunc_aa_mut_effect_df["nearest feature"].unique()}
for nf, gdf in non_trunc_aa_mut_effect_df.groupby("nearest feature"):
for _, r in gdf.iterrows():
f = r["nearest feature"]
for c in cnt_columns:
feat_mut_eff_cnt_d[f][c] += int(r[c])
feat_mut_eff_cnt_df = pd.DataFrame(feat_mut_eff_cnt_d).T
feat_mut_eff_cnt_df["total"] = feat_mut_eff_cnt_df.sum(axis=1)
feat_mut_eff_cnt_df
# -
# feat_mut_eff_cnt_df.to_pickle("./pykF_mut_nearest_feat_df.pkl") # The old name given is of an inappropraite context. Keeping this as comment to search out in other NBs and fix.
feat_mut_eff_cnt_df.to_pickle("./pykF_feat_mut_eff_cnt_df.pkl")
| pykF/pykF_mut_struct_clust.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> Graph Element</dd>
# <dt>Dependencies</dt> <dd>Bokeh</dd>
# <dt>Backends</dt> <dd><a href='./Graph.ipynb'>Bokeh</a></dd> <dd><a href='../matplotlib/Graph.ipynb'>Matplotlib</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
from holoviews import opts
hv.extension('bokeh')
# The ``Graph`` element provides an easy way to represent and visualize network graphs. It differs from other elements in HoloViews in that it consists of multiple sub-elements. The data of the ``Graph`` element itself are the abstract edges between the nodes. By default the element will automatically compute concrete ``x`` and ``y`` positions for the nodes and represent them using a ``Nodes`` element, which is stored on the Graph. The abstract edges and concrete node positions are sufficient to render the ``Graph`` by drawing straight-line edges between the nodes. In order to supply explicit edge paths we can also declare ``EdgePaths``, providing explicit coordinates for each edge to follow.
#
# To summarize a ``Graph`` consists of three different components:
#
# * The ``Graph`` itself holds the abstract edges stored as a table of node indices.
# * The ``Nodes`` hold the concrete ``x`` and ``y`` positions of each node along with a node ``index``. The ``Nodes`` may also define any number of value dimensions, which can be revealed when hovering over the nodes or to color the nodes by.
# * The ``EdgePaths`` can optionally be supplied to declare explicit node paths.
#
# This reference document describes only basic functionality, for a more detailed summary on how to work with network graphs in HoloViews see the [User Guide](../../../user_guide/Network_Graphs.ipynb).
#
# To make the visualizations in this notebook easier to view, the first thing we will do is increase the default width and height:
opts.defaults(opts.Graph(width=400, height=400))
# #### A simple Graph
#
# Let's start by declaring a very simple graph connecting one node to all others. If we simply supply the abstract connectivity of the ``Graph``, it will automatically compute a layout for the nodes using the ``layout_nodes`` operation, which defaults to a circular layout:
# +
# Declare abstract edges
N = 8
node_indices = np.arange(N)
source = np.zeros(N)
target = node_indices
padding = dict(x=(-1.2, 1.2), y=(-1.2, 1.2))
simple_graph = hv.Graph(((source, target),)).redim.range(**padding)
simple_graph
# -
# #### Directed graphs
#
# The Graph element also allows indicating the directionality of graph edges using arrows. To enable the arrows set ``directed=True`` and to optionally control the ``arrowhead_length`` provide a length as a fraction of the total graph extent:
simple_graph.opts(directed=True, arrowhead_length=0.05)
# #### Accessing the nodes and edges
#
# We can easily access the ``Nodes`` and ``EdgePaths`` on the ``Graph`` element using the corresponding properties:
#
simple_graph.nodes + simple_graph.edgepaths
# #### Additional features
#
# Next we will extend this example by supplying explicit edges, node information and edge weights. By constructing the ``Nodes`` explicitly we can declare an additional value dimensions, which are revealed when hovering and/or can be mapped to the color by specifying the ``color_index``. We can also associate additional information with each edge by supplying a value dimension to the ``Graph`` itself, which we can map to a color using the ``edge_color_index``.
# +
# Node info
np.random.seed(7)
x, y = simple_graph.nodes.array([0, 1]).T
node_labels = ['Output']+['Input']*(N-1)
edge_weights = np.random.rand(8)
# Compute edge paths
def bezier(start, end, control, steps=np.linspace(0, 1, 100)):
return (1-steps)**2*start + 2*(1-steps)*steps*control+steps**2*end
paths = []
for node_index in node_indices:
ex, ey = x[node_index], y[node_index]
paths.append(np.column_stack([bezier(x[0], ex, 0), bezier(y[0], ey, 0)]))
# Declare Graph
nodes = hv.Nodes((x, y, node_indices, node_labels), vdims='Type')
graph = hv.Graph(((source, target, edge_weights), nodes, paths), vdims='Weight')
graph.redim.range(**padding).opts(color_index='Type', edge_color_index='Weight',
cmap=['blue', 'red'], edge_cmap='viridis')
# -
# For full documentation and the available style and plot options, use ``hv.help(hv.Graph).``
| examples/reference/elements/bokeh/Graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter, defaultdict
class Solution:
def maxCount(self, m: int, n: int, ops):
if not ops:
return m * n
count = defaultdict(int)
for x, y in ops:
count[(x, y)] += 1
s_count = sorted(count.items(), key=lambda x: (x[0][0] * x[0][1], -x[1]))
print(s_count)
item = s_count.pop(0)
return item[0][0] * item[0][1]
# -
class Solution:
def maxCount(self, m: int, n: int, ops):
if ops == []:
return m*n
row = []
column = []
for i in ops:
row.append(i[0])
column.append(i[1])
print(row, column)
return min(row) * min(column)
solution = Solution()
solution.maxCount(18,3,[[16,1],[14,3],[14,2],[4,1],[10,1],[11,1],[8,3],[16,2],[13,1],[8,3],[2,2],[9,1],[3,1],[2,2],[6,3]])
| Math/1226/598. Range Addition II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Pandas Data Visualization Exercise
#
# This is just a quick exercise for you to review the various plots we showed earlier. Use **df3** to replicate the following plots.
import pandas as pd
import matplotlib.pyplot as plt
df3 = pd.read_csv('df3')
# %matplotlib inline
df3.info()
df3.head()
# ** Recreate this scatter plot of b vs a. Note the color and size of the points. Also note the figure size. See if you can figure out how to stretch it in a similar fashion. Remeber back to your matplotlib lecture...**
# ** Create a histogram of the 'a' column.**
# ** These plots are okay, but they don't look very polished. Use style sheets to set the style to 'ggplot' and redo the histogram from above. Also figure out how to add more bins to it.***
# ** Create a boxplot comparing the a and b columns.**
# ** Create a kde plot of the 'd' column **
# ** Figure out how to increase the linewidth and make the linestyle dashed. (Note: You would usually not dash a kde plot line)**
# ** Create an area plot of all the columns for just the rows up to 30. (hint: use .ix).**
# ## Bonus Challenge!
# Note, you may find this really hard, reference the solutions if you can't figure it out!
# ** Notice how the legend in our previous figure overlapped some of actual diagram. Can you figure out how to display the legend outside of the plot as shown below?**
#
# ** Try searching Google for a good stackoverflow link on this topic. If you can't find it on your own - [use this one for a hint.](http://stackoverflow.com/questions/23556153/how-to-put-legend-outside-the-plot-with-pandas)**
| Data Visualization/Pandas/02-Pandas Data Visualization Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **The notebook is for preprocessing the dataset and then build the suitable ML model on top of the dataset**
# **Initially the dataset is preprocessed and then model/algorithm is applied over it to predict the results**
#Import the libraries
import numpy as np
import pandas as pd
# Import the dataset
dataset = pd.read_csv('credit_card_defaults.csv')
dataset.head()
# **Data Preprocessing**
# Drop Id column, as it is not required
dataset = dataset.drop(['ID'], axis=1)
# Rename the column PAY_0 to PAY_1
dataset.rename(columns={'PAY_0':'PAY_1'}, inplace=True)
# ***Now,remove the unwanted categorical levels of features as seen in DataExploration notebook***
#Education
dataset['EDUCATION'].value_counts()
dataset['EDUCATION']=dataset['EDUCATION'].map({0:4,1:1,2:2,3:3,4:4,5:4,6:4})
dataset.head(2)
dataset['EDUCATION'].value_counts()
#Marriage
dataset['MARRIAGE'].value_counts()
dataset['MARRIAGE']=dataset['MARRIAGE'].map({0:3,1:1,2:2,3:3})
# Feature Scaling
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
scaler = scaler.fit(dataset)
# +
# Check for null values
#dataset.isnull().sum()
dataset.apply(lambda x:sum(x.isnull()), axis=0)
# +
# Split the train and test set
X = dataset.iloc[:,:-1].values
y = dataset.iloc[:,23].values
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state=0)
# -
# **Machine Learning Algorithms/Models**
# +
# Base model- with Logistic Regression technique
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state=10)
classifier.fit(X_train, y_train)
# +
# Prediction and evaluation metrics with Logistic Regression
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
#prediction
y_pred = classifier.predict(X_test)
y_pred
# -
#Evaluation
accuracy = accuracy_score(y_test,y_pred)
precision = precision_score(y_test, y_pred, average='weighted')
recall = recall_score(y_test, y_pred, average='weighted')
f1score = f1_score(y_test, y_pred, average='weighted')
print("Accuracy: ",accuracy)
print("Precision: ",precision)
print("Recall: ",recall)
print("F1Score: ",f1score)
# +
# Let's try ensemble techniques for better prediction
# +
#Random Forest
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(random_state=10)
classifier.fit(X_train,y_train)
# +
y_pred = classifier.predict(X_test)
#Evaluation
accuracy = accuracy_score(y_test, y_pred)
precision = precision_score(y_test, y_pred, average='weighted')
recall = recall_score(y_test, y_pred, average='weighted')
f1score = f1_score(y_test, y_pred, average='weighted')
print("Accuracy: ",accuracy)
print("Precision: ",precision)
print("Recall: ",recall)
print("F1Score: ",f1score)
# +
#GradientBoosting
from sklearn.ensemble import GradientBoostingClassifier
classifier = GradientBoostingClassifier(random_state=10)
classifier.fit(X_train,y_train)
# +
y_pred = classifier.predict(X_test)
#Evaluation
accuracy = accuracy_score(y_test,y_pred)
precision = precision_score(y_test, y_pred, average='weighted')
recall = recall_score(y_test, y_pred, average='weighted')
f1score = f1_score(y_test, y_pred, average='weighted')
print("Accuracy: ",accuracy)
print("Precision: ",precision)
print("Recall: ",recall)
print("F1Score: ",f1score)
# +
#Adaboost
from sklearn.ensemble import AdaBoostClassifier
classifier = AdaBoostClassifier(random_state=10)
classifier.fit(X_train, y_train)
# +
y_pred = classifier.predict(X_test)
accuracy = accuracy_score(y_test,y_pred)
precision = precision_score(y_test,y_pred,average='weighted')
recall = recall_score(y_test,y_pred,average='weighted')
f1score = f1_score(y_test,y_pred,average='weighted')
print("Accuracy: ",accuracy)
print("Precision: ",precision)
print("Recall: ",recall)
print("F1Score: ",f1score)
# -
| ModelFromMachineLearning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.image import imread
#Copy and paste the directory of the data
my_data_dir = 'C:\\Users\\yasas\\Desktop\\cell_images\\cell_images'
#Check what folders are in the data
os.listdir(my_data_dir)
test_path = my_data_dir+'\\test\\'
train_path = my_data_dir+'\\train\\'
os.listdir(test_path)
os.listdir(train_path)
#Infected Cell image
para_cell = train_path+'\\parasitized\\'+ os.listdir(train_path+'\\parasitized')[0]
para_img = imread(para_cell)
plt.imshow(para_img)
para_img.shape
#Regular cell image
unifected_cell_path = train_path+'\\uninfected\\'+os.listdir(train_path+'\\uninfected')[0]
unifected_cell = imread(unifected_cell_path)
plt.imshow(unifected_cell)
unifected_cell.shape
len(os.listdir(train_path+'\\parasitized'))
len(os.listdir(train_path+'\\uninfected'))
# +
dim1 = []
dim2 = []
for image_filename in os.listdir(test_path+'\\uninfected'):
img = imread(test_path+'\\uninfected'+'\\'+image_filename)
d1,d2,colors = img.shape
dim1.append(d1)
dim2.append(d2)
# -
np.mean(dim1)
np.mean(dim2)
image_shape = (131, 131, 3)
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# +
#Randomly transforms images to help CNN perform better on rotated/scaled images
image_gen = ImageDataGenerator(rotation_range=20, # rotate the image 20 degrees
width_shift_range=0.10, # Shift the pic width by a max of 5%
height_shift_range=0.10, # Shift the pic height by a max of 5%
rescale=1/255, # Rescale the image by normalzing it.
shear_range=0.1, # Shear means cutting away part of the image (max 10%)
zoom_range=0.1, # Zoom in by 10% max
horizontal_flip=True, # Allo horizontal flipping
fill_mode='nearest' # Fill in missing pixels with the nearest filled value
)
# -
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Activation, Dropout, Flatten, Dense, Conv2D, MaxPooling2D
# +
model = Sequential()
model.add(Conv2D(filters=32, kernel_size=(3,3),input_shape=image_shape, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=64, kernel_size=(3,3),input_shape=image_shape, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=64, kernel_size=(3,3),input_shape=image_shape, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# -
from tensorflow.keras.callbacks import EarlyStopping
early_stop = EarlyStopping(monitor='val_loss',patience=2)
batch_size = 16
train_image_gen = image_gen.flow_from_directory(train_path,
target_size=image_shape[:2],
color_mode='rgb',
batch_size=batch_size,
class_mode='binary')
test_image_gen = image_gen.flow_from_directory(test_path,
target_size=image_shape[:2],
color_mode='rgb',
batch_size=batch_size,
class_mode='binary',shuffle=False)
model.fit_generator(train_image_gen,epochs=20,
validation_data=test_image_gen,
callbacks=[early_stop])
losses = pd.DataFrame(model.history.history)
losses[['loss','val_loss']].plot()
losses[['acc','val_acc']].plot()
from tensorflow.keras.preprocessing import image
pred_probabilities = model.predict_generator(test_image_gen)
pred_probabilities
test_image_gen.classes
predictions = pred_probabilities > 0.5
predictions
# +
from sklearn.metrics import classification_report
print(classification_report(test_image_gen.classes,predictions))
# -
from tensorflow.keras.models import load_model
model.save('infected_cell_detector.h5')
| Malaria_Detection_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 파이썬 데이터 모델 설명
# Namedtuple
# namedtuple attrs
# Namedtuple method
# List Comprehesion
# 구조화된 모델 설명
# NamedTuple
# 데이터 모델 (Data Model)
# NamedTuple
# 파이썬의 중요한 핵심 프레임워크 -> 시퀀스(Sequence), 반복(iterator), 함수(Function), 클래스(Class)
# Iterator --> Generator
# Sequence
# 객체 -> 파이썬의 데이터를 추상화
# 모든 객체 -> id, type -> value
# 파이썬 -> 일관성
# +
# NamedTuple
# 일반적인 튜플 사용
pt1 = (1.0, 5.0)
pt2 = (2.5, 1.5)
from math import sqrt
line_leng1 = sqrt((pt2[0] - pt1[0]) ** 2 + (pt2[1] - pt1[1]) ** 2)
print('Ex1-1 -', line_leng1)
# +
# 네임드 튜플 사용
from collections import namedtuple
# 네임드 튜플 선언
Point = namedtuple('Point', 'x y')
# 두 점 선언
pt1 = Point(1.0, 5.0)
pt2 = Point(2.5, 1.5)
# 출력
line_leng2 = sqrt((pt1.x - pt2.x) ** 2 + (pt1.y - pt2.y) ** 2)
print('EX1-2 -', line_leng2)
print('EX1-3', line_leng1 == line_leng2)
# +
# 네임드 튜플 선언 방법
Point1 = namedtuple('Point', ['x', 'y'])
Point2 = namedtuple('Point', 'x, y')
Point3 = namedtuple('Point', 'x y')
Point4 = namedtuple('Point', 'x y x class', rename=True) # Default=False
# 출력
print('EX2-1', Point1, Point2, Point3, Point4)
# 객체 생성
p1 = Point1(x=10, y=35)
p2 = Point2(20, 40)
p3 = Point3(45, y=20)
p4 = Point4(10, 20, 30, 40)
# Dict to Unpacking
temp_dict = {'x': 75, 'y': 55}
p5 = Point3(**temp_dict)
# 출력
print('EX2-2 - ', p1, p2, p3, p4, p5)
# -
# 사용
print('EX3-1 - ', p1[0] + p2[1]) # Index Error 주의
print('EX3-2 - ', p1.x + p2.y) # 클래스 변수 접근 방식
# +
# Unpacking
x, y = p3
print('EX3-3 - ', x + y)
# Rename 테스트
print('EX3-4 - ', p4)
print()
print()
# +
# Namedtuple 메서드
temp = [52, 38]
# _make() : 새로운 객체 생성
p4 = Point1._make(temp)
print('EX4-1', p4)
# _fields : 필드 네임 확인
print('EX4-2 -', p1._fields, p2._fields, p3._fields)
# _asdict() : OrderedDict 로 반환
print('EX4-3', p1._asdict(), p2._asdict(), p4._asdict())
# _replace() : 수정된 새로운 객체 반환 (아이디값이 바뀜), 튜플은 불변
print('EX4-4 -', p2._replace(y=100))
# +
# 실사용 실습
# 학생 전체 그룹 새엇ㅇ
# 반20명, 4개의 반 -> (A, B, C, D) 번호
# 네임드 튜플 선언
Classes = namedtuple('Classes', ['rank', 'number'])
# 그룹 리스트 선언
# 지능형 리스트 (List Comprehension)
numbers = [str(n) for n in range(1, 21)]
print(numbers)
ranks = 'A B C D'.split()
print(rank, numbers)
# List Comprehension 사용
students = [Classes(rank, number) for rank in ranks for number in numbers]
print(students)
print('EX5-1 - ', len(students))
print('EX5-2 - ', students)
# +
# 가독성 x
students2 = [Classes(rank, number)
for rank in 'A B C D'.split()
for number in [str(n) for n in range(1,21)]]
print(students2)
# +
# 출력
for s in students:
print('Ex7-1', s)
# -
| python/data_model_special_method_named_tuple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from bayes_opt import BayesianOptimization
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import gridspec
import sqlite3
# -
def queryTarget(x):
return x*x
bo = BayesianOptimization(queryTarget, {'x':(-2,2)})
bo.maximize(init_points=2, n_iter=0, acq='ei', kappa=5)
bo.maximize(init_points=0, n_iter=1, acq='ei', kappa=5)
bo.maximize(init_points=0, n_iter=1, acq='ei', kappa=5)
bo.maximize(init_points=0, n_iter=1, acq='ei', kappa=5)
| examples/Untitled1.ipynb |
# # Relationship Extraction
#
# In this notebook, we'll train, deploy and use an relationship extraction
# model using transformers from the
# [transformers](https://huggingface.co/transformers/) library which uses
# PyTorch.
#
# **Note**: When running this notebook on SageMaker Studio, you should make
# sure the 'SageMaker JumpStart PyTorch 1.0' image/kernel is used. When
# running this notebook on SageMaker Notebook Instance, you should make
# sure the 'sagemaker-soln' kernel is used.
# We start by importing a variety of packages that will be used throughout
# the notebook. One of the most important packages is the Amazon SageMaker
# Python SDK (i.e. `import sagemaker`). We also import modules from our own
# custom (and editable) package that can be found at `../package`.
# +
import boto3
from pathlib import Path
import sagemaker
from sagemaker.pytorch import PyTorch
from sagemaker.predictor import json_serializer, json_deserializer
import sys
sys.path.insert(0, '../package')
from package import config, utils
# -
# Up next, we define the current folder and create a SageMaker client (from
# `boto3`). We can use the SageMaker client to call SageMaker APIs
# directly, as an alternative to using the Amazon SageMaker SDK. We'll use
# it at the end of the notebook to delete certain resources that are
# created in this notebook.
current_folder = utils.get_current_folder(globals())
sagemaker_client = boto3.client('sagemaker')
sagemaker_session = sagemaker.Session()
# !aws s3 cp --recursive --quiet $config.SOURCE_S3_PATH/data ../data
# !aws s3 cp --recursive --quiet ../data s3://$config.S3_BUCKET/$config.DATASETS_S3_PREFIX
# +
hyperparameters = {
"learning-rate": 0.0007
}
current_folder = utils.get_current_folder(globals())
estimator = PyTorch(
framework_version='1.5.1',
entry_point='entry_point.py',
source_dir=str(Path(current_folder, '../containers/relationship_extraction').resolve()),
hyperparameters=hyperparameters,
role=config.IAM_ROLE,
train_instance_count=1,
train_instance_type=config.TRAINING_INSTANCE_TYPE,
output_path='s3://' + str(Path(config.S3_BUCKET, config.OUTPUTS_S3_PREFIX)),
code_location='s3://' + str(Path(config.S3_BUCKET, config.OUTPUTS_S3_PREFIX)),
base_job_name=config.SOLUTION_PREFIX,
tags=[{'Key': config.TAG_KEY, 'Value': config.SOLUTION_PREFIX}],
sagemaker_session=sagemaker_session,
train_volume_size=30
)
# -
estimator.fit({
'train': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'semeval', 'train')),
'test': 's3://' + str(Path(config.S3_BUCKET, config.DATASETS_S3_PREFIX, 'semeval', 'test'))
})
# We'll use the unique solution prefix to name the model and endpoint.
model_name = "{}-relationship-extraction".format(config.SOLUTION_PREFIX)
predictor = estimator.deploy(
endpoint_name=model_name,
instance_type=config.HOSTING_INSTANCE_TYPE,
initial_instance_count=1
)
# When calling our new endpoint from the notebook, we use a Amazon
# SageMaker SDK
# [`Predictor`](https://sagemaker.readthedocs.io/en/stable/predictors.html).
# A `Predictor` is used to send data to an endpoint (as part of a request),
# and interpret the response. Our `estimator.deploy` command returned a
# `Predictor` but, by default, it will send and receive numpy arrays. Our
# endpoint expects to receive (and also sends) JSON formatted objects, so
# we modify the `Predictor` to use JSON instead of the PyTorch endpoint
# default of numpy arrays. JSON is used here because it is a standard
# endpoint format and the endpoint response can contain nested data
# structures.
predictor.content_type = 'application/json'
predictor.accept = 'application/json'
predictor.serializer = json_serializer
predictor.deserializer = json_deserializer
# With our model successfully deployed and our predictor configured, we can
# try out the relationship extraction model out on example inputs.
predictor.predict(
data={
'sequence': 'Amazon SageMaker is a fully managed service that provides every developer and data scientist with the ability to build, train, and deploy machine learning (ML) models quickly.',
'entity_one_start': 0,
'entity_one_end': 6,
'entity_two_start': 7,
'entity_two_end': 16
}
)
# ## Clean Up
#
# When you've finished with the relationship extraction endpoint (and associated
# endpoint-config), make sure that you delete it to avoid accidental
# charges.
predictor.delete_endpoint()
predictor.delete_model()
| sagemaker_notebook_instance/notebooks/4_relationship_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercícios com Reais
# ## Exercício 1
# Uma pessoa aplicou um capital de $x$ complexos a juros mensais de $z$ durante $1$ ano. Determinar o montante de cada mês durante este período
# + pycharm={"is_executing": true}
def main():
capital = float(input('Digite o capital aplicado: '))
taxa_de_juros = float(input('Digite a taxa de Juros: '))
montante = capital + (capital * taxa_de_juros / 100)
for i in range(12):
print(f'mês: {i} / montante: {montante}')
montante = montante + (montante*taxa_de_juros/100)
main()
# -
# ## Exercício 2
# Dado um natural $n$, determine o número harmônico $H_{n}$ definido por: 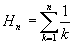
# + pycharm={"is_executing": true}
def main():
n = int(input('Digite um número natural: '))
harm = 0
for k in range(1, n+1):
harm += 1/k
print(f'O harnônico do numero {n} é {harm}')
main()
# -
# ## Exercício 3
# Os pontos $(x,y)$ que pertencem à figura $H$ (abaixo) são tais que $x \geq 0$, $y \geq 0$ e $ x^{2} + y^{2} \leq 1$. Dados $n$ pontos reais $(x,y)$, verifique se cada ponto pertence ou não a $H$. 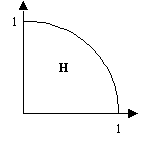
# + pycharm={"is_executing": true}
def verficar_h(x, y):
if x >= 0 and y >= 0:
if (x**2 + y**2) <= 1:
return True
else:
return False
else:
return False
def main():
n = int(input('n: '))
for i in range(n):
x = float(input('Digite a cordenada X: '))
y = float(input('Digite a cordenada Y: '))
if verficar_h(x, y):
print(f'({x},{y}) pertence a H')
else:
print(f'({x},{y}) não pertence a H')
main()
# -
# ## Exercício 4
# (GEO 84) Considere o conjunto 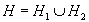 de pontos reais, onde: 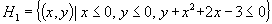 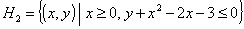
# Faça um programa que lê uma sequência de $n$ pontos reais $(x,y)$ e verifica se cada ponto pertence ou não ao conjunto $H$. O programa deve também contar o número de pontos da sequência que pertencem a $H$.
# + pycharm={"is_executing": true}
def main():
n = int(input('n: '))
cont = 0
for i in range(n):
x = float(input('Insira um valor de X: '))
y = float(input('Insira um valor de Y: '))
if x <= 0 and y <= 0 and (y + x**2 + 2 * x - 3) <= 0:
print(f'O ponto ({x},{y}) pertence ao conjunto H')
cont += 1
print(f'{cont} pontos da sequencia que pertencem a H')
elif x >= 0 and (y + x**2 - 2 * x - 3) <= 0:
cont += 1
print(f'O ponto ({x},{y}) pertence ao conjunto H')
print(f'{cont} pontos da sequencia que pertencem a H')
main()
# -
# ## Exercício 5
# Dados números reais $a$, $b$ e $c$, calcular as raízes de uma equação do 2º grau da forma $ax^2 + bx + c = 0$.
# + pycharm={"is_executing": true}
def main():
print('Digite valores que satisfazem a equação ax^2 + bx + c = 0')
a = float(input('Digite um valor de a: '))
b = float(input('Digite um valor de b: '))
c = float(input('Digite um valor de c: '))
delta = b**2 - (4*a*c)
x_1 = (-b + delta**0.5) / (2*a)
x_2 = (-b - delta ** 0.5) / (2 * a)
if delta > 0:
print(f'RAIZES DISTINTAS')
print(f'Raiz 1: {x_1}')
print(f'Raiz 2: {x_2}')
if delta < 0:
print(f'COMPLEXAS')
print(f'Parte Real: {-b / (2*a)}')
print(f'Parte Imaginária: ±{((-delta)**0.5) / (2*a)}')
else:
print('DUPLA')
print(f'Raiz: {x_1}')
main()
# -
# ## Exercício 6
# Dados $x$ real e $n$ natural, calcular uma aproximação para $cos(x)$ através dos $n$ primeiros termos da seguinte série: 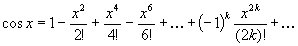
# + pycharm={"is_executing": true}
def fatorial(x):
prod = 1
for i in range(1, x+1):
prod *= i
return prod
def cos(x, n):
soma = 0
for i in range(0, n):
soma += (((-1) ** i)/fatorial(2 * i)) * (x ** (2 * i))
return soma
def main():
x = float(input('Digite o angulo: '))
n = int(input('Ordem na série de taylor: '))
print(f'O cosseno do ângulo {x} é {cos(x,n)}')
main()
# -
# ## Exercício 7
# Dados $x$ e $\varepsilon$ reais, $\varepsilon > 0$, calcular uma aproximação para $sen(x)$ através da seguinte série infinita:
# 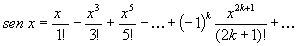
# incluindo todos os termos até que:
# 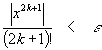
# + pycharm={"is_executing": true}
def fatorial(x):
prod = 1
for i in range(1, x+1):
prod *= i
return prod
def sen(x, epi):
soma = 0
i = 0
while (abs(x ** (2 * i + 1)/fatorial(2 * i + 1))) > epi:
soma += (((-1) ** i)/fatorial(2 * i + 1)) * (x ** (2 * i + 1))
i += 1
return soma
def main():
x = float(input('Digite o angulo: '))
epi = float(input('Digite um epsilon: '))
print(f'O seno do ângulo {x} é {sen(x, epi)}')
main()
# -
# ## Exercício 8
# Para $n$ alunos de uma determinada classe são dadas as $3$ notas das provas. Calcular a média aritmética das provas de cada aluno, a média da classe, o número de aprovados e o número de reprovados (critério de aprovação: média maior ou igual a cinco).
# + pycharm={"is_executing": true}
def main():
n = int(input('Quantidade de alunos na sala: '))
reprovados = 0
aprovados = 0
med_classe = 0
for i in range(1, n+1):
nota_um = float(input(f'Digite a primeira nota do aluno {i}: '))
nota_dois = float(input(f'Digite a segunda nota do aluno {i}: '))
nota_tres = float(input(f'Digite a terceira nota do aluno {i}: '))
media = (nota_um + nota_dois + nota_tres) / 3.0
med_classe += media
if media >= 5:
print(f'ALUNO APROVADO COM MÉDIA: {media}')
aprovados += 1
else:
print(f'ALUNO REPROVADO COM MÉDIA: {media}')
reprovados += 1
print(f'A média da classe: {med_classe / n}')
print(f'Números de aprovados: {aprovados}')
print(f'Números de reporvados: {reprovados}')
main()
# -
# ## Exercício 9
# Dadas $n$ triplas compostas por um símbolo de operação $(+, -, * ou /)$ e dois números reais, calcule o resultado ao efetuar a operação indicada para os dois números (Sugestão: use switch)
# + pycharm={"is_executing": true}
def main():
tripla = input('Digite uma operação: ')
if '+' in tripla:
numeros = tripla.split('+')
print(int(numeros[0]) + int(numeros[1]))
elif '-' in tripla:
numeros = tripla.split('-')
print(int(numeros[0]) - int(numeros[1]))
elif '*' in tripla:
numeros = tripla.split('*')
print(int(numeros[0]) * int(numeros[1]))
elif '/' in tripla:
numeros = tripla.split('/')
print(int(numeros[0]) / int(numeros[1]))
main()
# -
# ## Exercício 10
# Dadas as populações de Uauá (BA) e Nova York (PI) e sabendo que a população de Uauá tem um crescimento anual de $x$ e a população de Nova York tem um crescimento anual de $y$ determine:
#
# - Se a população da cidade menor ultrapassa a da maior.
# - Quantos anos passarão antes que isso ocorra.
# + pycharm={"is_executing": true}
def main():
pop_uaua = 25853
pop_ny = 8399000
ano = 2020
x = float(input('Digite o crescimento anual de Uauá: '))
y = float(input('Digite o crescimento anual de Nova Iorque: '))
while pop_ny >= pop_uaua:
pop_uaua += pop_uaua*x/100
pop_ny += pop_ny*y/100
ano += 1
print(f'Em {ano} a população de Uauá vai passar a de Nova Iorque')
main()
# -
# ## Exercício 11
# Dado um inteiro positivo $n$, calcular e imprimir o valor da seguinte soma: 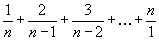
# + pycharm={"is_executing": true}
def main():
n = int(input('Digite um numero interio: '))
soma = 0
for i in range(0, n):
soma += (1+i)/(n-i)
print(soma)
main()
# -
# ## Exercício 12
# Faça um programa que calcula a soma:
# 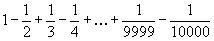
# pelas seguintes maneiras:
#
# * adição dos termos da direita para a esquerda;
# * adição dos termos da esquerda para a direita;
# * adição separada dos termos positivos e dos termos negativos da esquerda para a direita;
# * adição separada dos termos positivos e dos termos negativos da direita para a esquerda.
#
# + pycharm={"is_executing": true}
def soma_dir_esq():
# adição dos termos da direita para a esquerda
soma = 0
for i in range(10000, 0, -1):
if i % 2 == 0:
soma -= 1 / i
else:
soma += 1 / i
print(f'A soma dos termos da direita para a esquerda é: {soma}')
def soma_esq_dir():
# adição dos termos da esquerda para a direita
soma = 0
for i in range(1, 10001):
if i % 2 == 0:
soma -= 1 / i
else:
soma += 1 / i
print(f'A soma dos termos da esquerda pare a direita é: {soma}')
def soma_sep_esq_dir():
# adição separada dos termos positivos e dos temos negativos da esquerda para a direita
pos = 0
neg = 0
for i in range(1, 10001):
if i % 2 == 0:
neg -= 1 / i
else:
pos += 1 / i
print(
f'A soma dos temos positivos é: {pos} / A soma dos temos negativos é {neg} / A soma da esquerda para direita é {pos + neg}')
def soma_sep_dir_esq():
# adição separada dos termos positivos e dos termos negativos da dirreita para a esquerda
pos = 0
neg = 0
for i in range(10000, 0, -1):
if i % 2 == 0:
neg -= 1 / i
else:
pos += 1 / i
print(
f'A soma dos temos positivos é: {pos} / A soma dos temos negativos é {neg} / A soma da direita para esquerda é {pos + neg}')
def main():
soma_dir_esq()
soma_esq_dir()
soma_sep_esq_dir()
soma_sep_dir_esq()
main()
| Macmulti/exercicios_com_reais.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Factors that affect student performance
#
# ### By : <NAME>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Investigation Overview
#
# >In this investigation, I wanted to look at the characteristics of students that could be affect their grades. The main focus and the main attributes i worked on are
# ( the parent education level ,gender ,Test Preparation course ,the race groubs, and Of course
# students grades in 3 subjects)
#
# ## Dataset Overview
#
# >student performance dataset from Kaggle it consist of 1000 student informations like grades in 3 subjects (maths , reading ,
# writing )and the parent education level
# and 4 other attributes (gender, race, launch, test_preparation_course),
# I modified the names of the columns to be easy to use and modified the data by adding 6 more attributes (pass_math,pass_reading,pass_writing,Total_score,percentage_score,result_score) converted the 3 subjects grades to be shown as
# (fail/pass) and added the 3 subjects grades and make a total and percentage columns with result column as (fail /pass)
#
# + slideshow={"slide_type": "skip"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
# suppress warnings from final output
import warnings
warnings.simplefilter("ignore")
# + slideshow={"slide_type": "skip"}
#load data
df= pd.read_csv("StudentsPerformance.csv")
# + slideshow={"slide_type": "skip"}
# #copy from orginal df
df2=df.copy()
df2.head()
# + slideshow={"slide_type": "skip"}
#renaming the columns
df2.rename(columns = {"race/ethnicity": "race",
"parental level of education":"parent_education",
"test preparation course": "test_preparation_course",
"math score":"maths_score",
"reading score":"reading_score",
"writing score":"writing_score"},inplace=True)
df2.head()
# + slideshow={"slide_type": "skip"}
# creating a new columns pass_math,reading_math,writing_math this columns will tell us whether the students are pass or fail
passmarks = 40 #assume passmark=40
df2['pass_math'] = np.where(df2['maths_score']< passmarks, 'Fail', 'Pass')
df2['pass_reading'] = np.where(df2['reading_score']< passmarks, 'Fail', 'Pass')
df2['pass_writing'] = np.where(df2['writing_score']< passmarks, 'Fail', 'Pass')
# + slideshow={"slide_type": "skip"}
#creating a new columns of total score of each student
df2['Total_score']= df2['maths_score']+df2['writing_score']+df2['reading_score']
# + slideshow={"slide_type": "skip"}
#creating a new column of percentage of each student
df2['percentage_score']=df2['Total_score']/300
# + slideshow={"slide_type": "skip"}
#creating a new columns pass/fail result of each student
def result(percentage_score):
if percentage_score >=0.33:
return "Pass"
else:
return "Fail"
df2['result_score']=df2['percentage_score'].apply(result)
# + slideshow={"slide_type": "skip"}
#the new data
df2.head()
# + slideshow={"slide_type": "skip"}
# convert parental level of education and race/ethnicity into ordered categorical types
ordinal_var_dict = {'parent_education': ["some high school","high school","some college","associate's degree"
,"bachelor's degree","master's degree"],
'race': ['group E', 'group D', 'group C', 'group B', 'group A']}
for var in ordinal_var_dict:
ordered_var = pd.api.types.CategoricalDtype(ordered = True,
categories = ordinal_var_dict[var])
df2[var] = df2[var].astype(ordered_var)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Comparison of parent education level
#
# at the Bar chart we figure out at (parental level of education) :
# the highest category in it is "some collage" with prop = 22.6% , and the next from it is ""associate's degree" with prop =22.2%
# and the lowest category is "master's degree" with prop = 5.9% .
#
# + slideshow={"slide_type": "subslide"}
# create the plot
default_color = sb.color_palette('viridis',9)[3]
plt.figure(figsize = [9, 6])
sb.countplot(data = df2, x = 'parent_education', color = default_color);
# add annotations
n_df=df2.shape[0]
parental_counts = df2['parent_education'].value_counts()
locs, labels = plt.xticks()
# loop through each pair of locations and labels
for loc, label in zip(locs, labels):
# get the text property for the label to get the correct count
count = parental_counts[label.get_text()]
pct_string = '{:0.1f}%'.format(100*count/n_df)
# print the annotation just below the top of the bar
plt.text(loc, count-8, pct_string, ha = 'center', color = 'w',fontweight = 30,fontsize=12)
plt.xticks(rotation=15);
plt.title('Comparison of parent education level',fontweight = 30,fontsize=13);
plt.yticks([0,25,50,75,100,125,150,175,200,225]);
plt.xlabel("Education levels");
# + [markdown] slideshow={"slide_type": "slide"}
# ## relations between parent education and maths score,reading score,writing score,Total score
#
# we asked at the first if the parent education affects with the result or not now we will know the answer with the point plots below:
#
#
# the avarge highest grades at all 3 subjects and the total score happened with parent education of master 's degree
# and the lowest avarge grade at math happened with parent education of high school
#
# so the the parent education is realy affects with the result.
#
#
# + slideshow={"slide_type": "subslide"}
#relations between parent education and maths score,reading score,writing score,Total score
p1=df2.groupby('parent_education')['maths_score'].mean().sort_values()
p2=df2.groupby('parent_education')['reading_score'].mean().sort_values()
p3=df2.groupby('parent_education')['writing_score'].mean().sort_values()
p4=df2.groupby('parent_education')['Total_score'].mean().sort_values()
color=sb.color_palette()[0]
plt.figure(figsize=(18,8))
plt.subplot(1, 2, 1);
sb.pointplot(data=df2,x=p1.index, y=p1,color=color)
plt.title("Maths Score avarge VS parent education",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
plt.subplot(1, 2, 2);
sb.pointplot(data=df2,x=p2.index, y=p2,color=color)
plt.title("Reading Score avarge VS parent education",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(18,8))
plt.subplot(1, 2, 1);
sb.pointplot(data=df2,x=p3.index, y=p3,color=color)
plt.title("Writing Score avarge VS parent education",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
plt.subplot(1, 2, 2);
sb.pointplot(data=df2,x=p4.index, y=p4,color=color)
plt.title("Total Score avarge VS parent education", fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multivariate relations between (result score vs (parent education & Total score))
#
# we see at a high level of education like master's degree and bachelor's degree no one fail as a result score
#
# thats conform that parent education affect the grades of the students .
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=[9,6])
sb.pointplot(data = df2, x = 'parent_education', y = 'Total_score', hue = 'result_score',
dodge = 0.3, linestyles = "")
plt.xticks(rotation=90);
plt.legend(loc = 5, title = 'result_score');
plt.title("parent education & Total score .VS. result score ",fontweight = 30,fontsize=14);
# + [markdown] slideshow={"slide_type": "slide"}
# # let's move on and see another attribute like gender and know if that affect the grades of the students .
#
#
# ### Student Numbers of Males VS Femals
#
# >the females is more than the males at the data
# females = 51.8% , males= 48.2%
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = [8, 6])
colors=( 'mistyrose', 'skyblue')
plt.pie(df2['gender'].value_counts()/n_df,labels=('Female','Male'),
explode = [0.08,0.08],autopct ='%1.1f%%'
,shadow = True,startangle = 90, textprops={'fontsize': 12},colors=colors);
plt.title('Students Numbers of Males VS Femals ',fontweight = 30,fontsize=14 );
plt.axis('equal');
plt.ylabel('count')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## relation between gender and marks scored in each subject
#
#
# at realations between the gender and the 3 subjects with the boxs plots above we discoverd that :
# the average of maths score at the males is higher than at females (male=68.7>female=63.6),
# the average of reading score at the female is higher than at males (male=65.4<female=72.6),
# the average of writing score at the female is higher than at males (male=63.3<female=72.4)
#
#
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(18,8))
plt.subplot(1, 3, 1);
default_color = sb.color_palette('vlag',9)[5]
#plot for maths score
sb.boxplot(data=df2,x='gender', y='maths_score',color=default_color)
plt.title("Maths Score .VS. gender",fontweight = 30,fontsize=14);
#plot for reading score
plt.subplot(1, 3, 2);
sb.boxplot(data=df2,x='gender', y='reading_score',color=default_color)
plt.title("Reading Score .VS. gender",fontweight = 30,fontsize=14);
#plot for writing score
plt.subplot(1, 3, 3);
sb.boxplot(data=df2,x='gender', y='writing_score',color=default_color)
plt.title("writing score .VS. gender",fontweight = 30,fontsize=14);
plt.show();
# + [markdown] slideshow={"slide_type": "slide"}
# ## relations between gender and total,percentage score
#
#
# at realations between the gender and total,percentage score with the boxs plots above we discoverd that :
# the average of total score at the females is higher than at males (male=197.5<female=208.7),
# the average of percentage score at the female is higher than at males (male=0.69<female=0.65),
#
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(18,8))
plt.subplot(1, 2, 1);
default_color = sb.color_palette('vlag',9)[5]
#plot for gender & total score score
sb.boxplot(data=df2,x='gender', y='Total_score',color=default_color)
plt.title("Total Score .VS. gender",fontweight = 30,fontsize=14);
#plot for gender & percentage score
plt.subplot(1, 2, 2);
sb.boxplot(data=df2,x='gender', y='percentage_score',color=default_color)
plt.title("percentage Score .VS. gender",fontweight = 30,fontsize=14);
plt.show();
# + [markdown] slideshow={"slide_type": "slide"}
# ## Result score .VS. gender
#
# from the plot we see that at the result of the 3 subjects together (Pass/Fail) the females is higher than males at the pass rate
# but it also is higher than at fail rate
# + slideshow={"slide_type": "subslide"}
#plot for result score and gender
plt.figure(figsize=(10,5))
default_color = sb.color_palette('vlag',9)[8]
sb.countplot(data = df2, x = 'gender', hue = 'result_score'
,palette = 'Greens')
plt.title("Result score .VS. gender",fontweight = 30,fontsize=14);
# + [markdown] slideshow={"slide_type": "slide"}
# ## let's move on and see another attribute like( test preparation course) and know if that affect the grades of the students .¶
#
#
#
#
# >## test preparation course(none/completed)
# at this Pie chart we discoverd that most of the students didn't completed the test preparation course .
# 64.2% didn't completed the preparation and 35.8% completed the preparation.
#
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = [8, 6])
colors=( 'silver', 'teal')
plt.pie(df2['test_preparation_course'].value_counts()/n_df,labels=('none','completed'),
explode = [0.08,0.08],autopct ='%1.1f%%'
,shadow = True,startangle = 90, textprops={'fontsize': 12},colors=colors)
plt.title('test preparation course(none/completed)) ',fontweight = 30,fontsize=14 );
plt.axis('equal')
plt.ylabel('count')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multivariate relations between (test preparation course & Total score .VS. result score))
#
# and from the plot shown it look like the students who completed test preparation course no one fail otherswise from the students who didn't completed it there is a percentage of fail
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = [9, 6])
sb.pointplot(data = df2, x = 'test_preparation_course', y = 'Total_score', hue = 'result_score',
dodge = 0.3, linestyles = "")
plt.xticks(rotation=90);
plt.legend(loc = 5, title = 'result_score')
plt.title("test preparation course & Total score .VS. result score ",fontweight = 30,fontsize=14);
# + [markdown] slideshow={"slide_type": "slide"}
# ## finally we here with last attribute we intersted in( race groubs ) and trying to know if that affect the grades of the students or which groub have the highest grades.
#
#
#
#
# >## Comparison of race Groups
# at this Bar chart too we figure out at (race) :
# the highest numbers of students in "group C" with prop = 31.9% , and the next from it is "group D" with prop =26.2%
# and the lowestnumbers of students in A" with prop = 8.9% .
# + slideshow={"slide_type": "subslide"}
# create the plot
default_color = sb.color_palette('viridis',9)[3]
plt.figure(figsize = [9, 6])
sb.countplot(data = df2, x = 'race', color = default_color);
# add annotations
parental_counts = df2['race'].value_counts()
locs, labels = plt.xticks()
# loop through each pair of locations and labels
for loc, label in zip(locs, labels):
# get the text property for the label to get the correct count
count = parental_counts[label.get_text()]
pct_string = '{:0.1f}%'.format(100*count/n_df)
# print the annotation just below the top of the bar
plt.text(loc, count-8, pct_string, ha = 'center', color = 'w')
plt.xticks(rotation=90);
plt.title('Comparison of race Groups',fontweight = 30,fontsize=14);
plt.yticks([0,50,100,150,200,250,300,350]);
plt.xlabel("Groups");
# + [markdown] slideshow={"slide_type": "slide"}
# ## relations between Race and maths score,reading score,writing score,Total score
#
# from the the point plots :
#
#
# the avarge highest grades at all 3 subjects and the total score happened with the race group "E"
# and the lowest avarge grade at all 3 subjects and the total score happened with the race group "A"
#
# + slideshow={"slide_type": "subslide"}
#relations between Race and maths score,reading score,writing score,Total score
g1=df2.groupby('race')['maths_score'].mean().sort_values()
g2=df2.groupby('race')['reading_score'].mean().sort_values()
g3=df2.groupby('race')['writing_score'].mean().sort_values()
g4=df2.groupby('race')['Total_score'].mean().sort_values()
color=sb.color_palette()[0]
plt.figure(figsize=(18,8))
plt.subplot(1, 2, 1);
sb.pointplot(data=df2,x=g1.index, y=g1,color=color)
plt.title("Maths Score avarge VS race",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
plt.subplot(1, 2, 2);
sb.pointplot(data=df2,x=g2.index, y=g2,color=color)
plt.title("Reading Score avarge VS race",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(18,8))
plt.subplot(1, 2, 1);
sb.pointplot(data=df2,x=g3.index, y=g3,color=color)
plt.title("Writing Score avarge VS race",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
plt.subplot(1, 2, 2);
sb.pointplot(data=df2,x=g4.index, y=g4,color=color)
plt.title("Total Score avarge VS race",fontweight = 30,fontsize=14);
plt.xticks(rotation=15);
# + [markdown] slideshow={"slide_type": "slide"}
# # Result summary
#
# ## percentage (Pass/Fail ) of Total Score of students
# -percentage of success = 98.5% and percentage of failed 1.5%
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = [8, 6])
plt.pie(df2['result_score'].value_counts()/n_df,labels=('Pass','Fail'),
explode = [0.08,0.08],autopct ='%1.1f%%'
,shadow = True,startangle = 45, textprops={'fontsize': 12},colors=colors)
plt.title('Pass/Fail Total Score of students ',fontweight = 30,fontsize=14)
plt.ylabel('count')
plt.axis('equal')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Conclusion
#
#
# -Males are doing better than females in maths
#
# -Females are doing better than males in writing and reading
#
# -Parent education affected with the results of the students, the Students whose parents have completed master's or bachleor's degree have not failed in any subject otherwise the highest rate of failuer with the Students whose parents have some high school.
#
# -The students who completed Test Preparation course no one of them fail
#
# -the avarge highest grades at all 3 subjects and the total score happened at group "E"
# and the lowest avarge grade at all 3 subjects and the total score happened at group "A"
#
# -percentage of success = 98.5% and percentage of failed 1.5%
#
#
# -
| StudentsPerformance_analysis_visualizations-slides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to ExKaldi
#
# In this section, we will decode the test data based on HMM-DNN model and WFST graph.
# +
import exkaldi
import os
dataDir = "librispeech_dummy"
# -
# Restorage the posteiori probability of AM from file (generated in 11_train_DNN_acoustic_model_with_tensorflow).
# +
probFile = os.path.join(dataDir, "exp", "train_DNN", "amp.npy")
prob = exkaldi.load_prob(probFile)
prob
# -
prob.subset(nHead=1).data
# As above, this is naive output without log softmax activation function. We need do softmax.
#
# Exkaldi Numpy achivements have a method __.map(...)__ . We map a softmax function to all matrixs.
# +
prob = prob.map( lambda x: exkaldi.nn.log_softmax(x, axis=1) )
prob.subset(nHead=1).data
# -
# Then decode based on WFST. HCLG graph file and HMM model file have been generated (07_train_triphone_HMM-GMM_delta).
# +
HCLGFile = os.path.join(dataDir, "exp", "train_delta", "graph", "HCLG.fst")
hmmFile = os.path.join(dataDir, "exp", "train_delta", "final.mdl")
# -
# And for convenience, prepare lexicons.
# +
lexFile = os.path.join(dataDir, "exp", "lexicons.lex")
lexicons = exkaldi.decode.graph.load_lex(lexFile)
lexicons
# -
# Use __nn_decode__ function.
# +
lat = exkaldi.decode.wfst.nn_decode(prob, hmmFile, HCLGFile, symbolTable=lexicons("words"))
lat
# +
outDir = os.path.join(dataDir, "exp", "train_DNN", "decode_test")
exkaldi.utils.make_dependent_dirs(outDir, False)
lat.save( os.path.join(outDir,"test.lat") )
# -
# From lattice get 1-best result and score it.
# +
refIntFile = os.path.join(dataDir, "exp", "train_delta", "decode_test", "text.int")
for penalty in [0., 0.5, 1.0]:
for LMWT in range(10, 15):
newLat = lat.add_penalty(penalty)
result = newLat.get_1best(lexicons("words"), hmmFile, lmwt=LMWT, acwt=0.5)
score = exkaldi.decode.score.wer(ref=refIntFile, hyp=result, mode="present")
print(f"Penalty {penalty}, LMWT {LMWT}: WER {score.WER}")
# -
# In step 10_process_lattice_and_score, the best WER based on HMM-GMM model is about 135% and here is 107% in our experiment. So it is a truth that the performance of HMM-DNN got better.
# Up to here, the simple tutorial is over.
| tutorials/12_decode_back_HMM-DNN_and_WFST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from nltk import word_tokenize
from nltk import sent_tokenize
from nltk import FreqDist
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from nltk import bigrams
import re
f = open("alice.txt", "r")
alice=f.read()
filter_all=re.sub('@\w*|[!,?,#,~,&,%,£,*]|\.|\n|\ufeff|,|\'|\"|\’|\”|\“|\-|\_|the|\;|\:|\(|\)','',alice)
# +
#filter_all
# -
all_words=word_tokenize(filter_all)
stop_words=stopwords.words('english')
filtered_words=[]
for w in all_words:
if w not in stop_words:
filtered_words.append(w)
frdis=FreqDist(filtered_words)
frdis.plot(20)
A=bigrams(filtered_words)
fdis2=FreqDist(A)
fdis2.plot(10)
from nltk import ngrams
B=ngrams(filtered_words,3)
C=ngrams(filtered_words,4)
fdist_B=FreqDist(B)
fdist_C=FreqDist(C)
fdist_B.plot(10)
fdist_C.plot(12)
# +
import requests
import numpy as np # linear algebra
import pandas as pd # dataframes
from bs4 import BeautifulSoup # work with html
import nltk # natural language packages
import string # to do some work with strings
import matplotlib.pyplot as plt # data visualization
# %matplotlib inline
import seaborn as sns # data visualization
sns.set(color_codes=True) # data visualization
from textblob import TextBlob # sentiment analysis
# +
# #!pip3 install textblob
# +
def analize_sentiment_textblop(sentence):
analysis = TextBlob(sentence)
return analysis.sentiment.polarity
#Vader:
analyser = SentimentIntensityAnalyzer()
def sentiment_analyzer_positive(sentence):
score = analyser.polarity_scores(sentence)
retorno = score.get('pos')
return retorno
def sentiment_analyzer_negative(sentence):
score = analyser.polarity_scores(sentence)
retorno = score.get('neg')
return retorno
def sentiment_analyzer_vader(sentence):
score = analyser.polarity_scores(sentence)
#print("{:-<40} {}".format(sentence, str(score)))
retorno = score.get('compound')
return retorno
# -
from nltk.sentiment.vader import SentimentIntensityAnalyzer as SIA
sentiment_analyzer_negative('alice is wonderful')
| Twitter Sentiment Analysis/Alice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sklearn
import glob
from pathlib import Path
import sys
sys.path.append('../mss')
import mssmain as mss
import peakutils
from scipy.integrate import simps
from ast import literal_eval
import scipy
from tqdm import tqdm
#Read the dataset
path = '../example_data/peakdata/labelled_output/'
all_files = glob.glob(path + "/*.csv")
for i in range(len(all_files)):
if i == 0:
df = pd.read_csv(all_files[i])
df['source'] = all_files[i]
else:
df_else = pd.read_csv(all_files[i])
df_else['source'] = all_files[i]
df = df.append(df_else, ignore_index = True)
#reshape data
df.columns = ['index', 'mz', 'i array', 'label', 'source']
df = pd.DataFrame(df, columns = ['mz', 'i array', 'label', 'source', 'index'])
df.head()
# # RT conversion rate should be incorporated if any width parameters included
df_relabel = df[(df['label'] != 3) & (df['label'] != 2) & (df['label'] != 1)] #df for mislabelled peaks
df_model = df.drop(df_relabel.index) #data for modeling, now have ~3500 rows of data
rt_conversion_rate = 0.005533333
def peak_para(intensity, rt_conversion_rate, peak_thres = 0.01, thr = 0.02, min_d = 1, rt_window = 1.5, peak_area_thres = 1e5, min_scan = 15, max_scan = 200, max_peak = 5, min_scan_window = 20, sn_range = 7):
'''
firstly get rt, intensity from given mz and error out of the mzml file
Then find peak on the intensity array, represent as index --> index
Find peak range by looping from peak index forward/backward until hit the peak_base --> l_range,h_range. peakspan = h_range - l_range
Trim/correct peak range is too small or too large, using min_scan/max_scan,min_scan_window --> trimed l/h_range
Integration of peak based on the given range using simp function --> peakarea
'''
#Get rt_window corresponded scan number -- needs update later
#Get peak index
indexes = peakutils.indexes(intensity, thres=thr, min_dist = min_d)
result_dict = {}
#dev note: boundary detection refinement
for index in indexes:
h_range = index
l_range = index
base_intensity = peak_thres * intensity[index] # use relative thres, also considering S/N, 1/2 rt point?
half_intensity = 0.5 * intensity[index]
#Get the higher and lower boundary
while intensity[h_range] >= base_intensity:
h_range += 1
if intensity[h_range-1] < half_intensity: #potentially record this
if h_range - index > 4: #fit r2 score, keep record https://stackoverflow.com/questions/55649356/how-can-i-detect-if-trend-is-increasing-or-decreasing-in-time-series as alternative
x = np.linspace(h_range - 2, h_range, 3)
y = intensity[h_range - 2 : h_range + 1]
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(x, y)
# print(rt[h_range],r_value)
if abs(r_value) < 0.6:
break
elif h_range > len(intensity)-2:
break
while intensity[l_range] >= base_intensity: #Dev part 2, low priority since general peak shapes
l_range -= 1
if intensity[l_range] < half_intensity:
pass #backdoor for recording 1/2 rt point
#Output a range from the peak list
peak_range = intensity[l_range:h_range]#no filter so ignored for tailing effects
#print(index + scan_window)
#Calculate for S/N
signal = intensity[index]
neighbour_blank = intensity[l_range - sn_range : l_range] + intensity[h_range + 1 : h_range + sn_range + 1]
noise = max(neighbour_blank)
if noise != 0:
sn = round(signal/noise, 3)
else:
sn = 0
#Calculate height/width, consider log10 transform
height = signal
width = (h_range - l_range) * rt_conversion_rate
#Add rt conversion factor here to convert width in scan into rt
hw_ratio = round(height/width,0)
#------------------------------------------------new-------------------------------------------
#Additional global parameters
#1/2 peak range
h_loc = index
l_loc = index
while intensity[h_loc] > half_intensity:
h_loc += 1
while intensity[l_loc] > half_intensity:
l_loc -= 1
#calculate for slope -- interpolation included-- pay attention!
h_half = h_loc + (half_intensity - intensity[h_loc]) / (intensity[h_loc - 1] - intensity[h_loc])
l_half = l_loc + (half_intensity - intensity[l_loc]) / (intensity[l_loc + 1] - intensity[l_loc])
mb = (height - half_intensity) / ((h_half - index) * rt_conversion_rate) #when transfer back use rt[index] instead
ma = (height - half_intensity) / ((index - l_half) * rt_conversion_rate)
#------------------------------------------------new-------------------------------------------
#Intergration based on the simps function
if len(peak_range) >= min_scan:
integration_result = simps(peak_range)
if integration_result >= peak_area_thres:
#Calculate Area/background ratio, i.e, peak area vs rectangular area as whole(if =1 then peak is a pleateu)
background_area = (h_range - l_range) * height
ab_ratio = round(integration_result/background_area, 3)
#appending to result
if len(result_dict) == 0:
result_dict.update({index : [l_range, h_range, integration_result, sn, hw_ratio, ab_ratio, h_half, l_half, height, ma, mb, ma+mb, mb/ma]})
elif integration_result != list(result_dict.values())[-1][2]: #Compare with previous item
s_window = abs(index - list(result_dict.keys())[-1])
if s_window > min_scan_window:
result_dict.update({index : [l_range, h_range, integration_result, sn, hw_ratio, ab_ratio, h_half, l_half, height, ma, mb, ma+mb, mb/ma]})
#Filtering:
#1. delete results that l_range/h_range within 5 scans
#3. If still >5 then select top 5 results
#list(result_dict.values())[-1]
#Noise filter
if len(result_dict) > max_peak:
result_dict = {}
return result_dict
df_para = pd.DataFrame(columns = ['mz',
'i array',
'peak width in min',
'half intensity width in min',
'left width' ,
'right width',
'assymetric factor',
'integration',
'sn',
'hw',
'ab',
'peak height',
'ma',
'mb',
'broad rate',
'skewness',
'variance',
'label'])
for i, row in tqdm(df_model.iterrows()):
try:
i_array = literal_eval(row['i array'])
para = peak_para(i_array, rt_conversion_rate)
for i in para.items():
index = i[0]
l_range = i[1][0]
h_range = i[1][1]
integration = i[1][2]
sn = i[1][3]
hw = i[1][4]
ab = i[1][5]
h_half = i[1][6]
l_half = i[1][7]
height = i[1][8]
ma = i[1][9]
mb = i[1][10]
broad_rate = i[1][11]
skewness = i[1][12]
w = (h_range - l_range) * rt_conversion_rate
l_width = (index - l_range) * rt_conversion_rate
r_width = (h_range - index) * rt_conversion_rate
t_r = (h_half - l_half) * rt_conversion_rate
paradict = {'mz' : row['mz'],
'i array' : row['i array'],
'peak width in min' : w,
'half intensity width in min' : t_r,
'left width' : l_width,
'right width' : r_width,
'assymetric factor' : ((h_range - index) * rt_conversion_rate) / ((index - l_range) * rt_conversion_rate),
'integration' : integration,
'sn' : sn,
'hw' : hw,
'ab' : ab,
'peak height' : height,
'ma' : ma,
'mb' : mb,
'broad rate' : broad_rate,
'skewness' : skewness,
'variance' : w ** 2 / (1.764 * ((r_width / l_width) ** 2) - 11.15 * (r_width / l_width) + 28),
'label': row['label']}
df_para = df_para.append(paradict, ignore_index = True)
except:
continue
df_para.head()
df_para.to_csv('../example_data/peakdata/labelled_output/summary-4rd-newbatchincluded.csv')
| dev/D-archive/peak_picking_modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109B Data Science 2: Advanced Topics in Data Science
# ## Homework 6 - RNNs
#
#
#
# **Harvard University**<br/>
# **Fall 2020**<br/>
# **Instructors**: <NAME>, <NAME>, & <NAME>
#
#
# <hr style="height:2pt">
#RUN THIS CELL
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# ### INSTRUCTIONS
#
# <span style="color:red">**Model training can be very slow; start doing this HW early**</span>
#
# - To submit your assignment follow the instructions given in Canvas.
#
# - This homework can be submitted in pairs.
#
# - If you submit individually but you have worked with someone, please include the name of your **one** partner below.
# - Please restart the kernel and run the entire notebook again before you submit.
#
# **Names of person you have worked with goes here:**
# <br><BR>
# <div class="theme"> Overview: Named Entity Recognition Challenge</div>
# Named entity recognition (NER) seeks to locate and classify named entities present in unstructured text into predefined categories such as organizations, locations, expressions of times, names of persons, etc. This technique is often used in real use cases such as classifying content for news providers, efficient search algorithms over large corpora and content-based recommendation systems.
#
# This represents an interesting "many-to-many" problem, allowing us to experiment with recurrent architectures and compare their performances against other models.
#
# +
import pandas as pd
import numpy as np
import os
from sklearn.metrics import f1_score, confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
plt.style.use("ggplot")
# +
import tensorflow as tf
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.utils import to_categorical
from tensorflow.keras import backend
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras import Model, Sequential
from tensorflow.keras.models import model_from_json
from tensorflow.keras.layers import Input, SimpleRNN, Embedding, Dense, TimeDistributed, GRU, \
Dropout, Bidirectional, Conv1D, BatchNormalization
print(tf.keras.__version__)
print(tf.__version__)
# -
# Set seed for repeatable results
np.random.seed(123)
tf.random.set_seed(456)
# + [markdown] colab_type="text" id="rUkgUGwJXUcH"
# <div class="theme"> Part 1: Data </div>
# Read `HW6_data.csv` into a pandas dataframe using the provided code below.
# -
# Given code
path_dataset = './data/HW6_data.csv'
data = pd.read_csv(path_dataset,
encoding="latin1")
data = data.fillna(method="ffill")
print(data.shape)
data.head(15)
# As you can see, we have a dataset with sentences (```Sentence #``` column), each composed of words (```Word``` column) with part-of-speech tagging (```POS``` tagging) and inside–outside–beginning (IOB) named entity tags (```Tag``` column) attached. ```POS``` will not be used for this homework. We will predict ```Tag``` using only the words themselves.
#
# Essential info about entities:
# * geo = Geographical Entity
# * org = Organization
# * per = Person
# * gpe = Geopolitical Entity
# * tim = Time indicator
# * art = Artifact
# * eve = Event
# * nat = Natural Phenomenon
#
# IOB prefix:
# * B: beginning of named entity
# * I: inside of named entity
# * O: outside of named entity
#
# <div class='exercise'><b> Question 1: Data [20 points total]</b></div>
#
# **1.1** Create a list of unique words found in the 'Word' column and sort it in alphabetic order. Then append the special word "ENDPAD" to the end of the list, and assign it to the variable ```words```. Store the length of this list as ```n_words```. **Print your results for `n_words`**
#
# **1.2** Create a list of unique tags and sort it in alphabetic order. Then append the special word "PAD" to the end of the list, and assign it to the variable ```tags```. Store the length of this list as ```n_tags```. **Print your results for `n_tags`**
#
# **1.3** Process the data into a list of sentences where each sentence is a list of (word, tag) tuples. Here is an example of how the first sentence in the list should look:
#
# [('Thousands', 'O'),
# ('of', 'O'),
# ('demonstrators', 'O'),
# ('have', 'O'),
# ('marched', 'O'),
# ('through', 'O'),
# ('London', 'B-geo'),
# ('to', 'O'),
# ('protest', 'O'),
# ('the', 'O'),
# ('war', 'O'),
# ('in', 'O'),
# ('Iraq', 'B-geo'),
# ('and', 'O'),
# ('demand', 'O'),
# ('the', 'O'),
# ('withdrawal', 'O'),
# ('of', 'O'),
# ('British', 'B-gpe'),
# ('troops', 'O'),
# ('from', 'O'),
# ('that', 'O'),
# ('country', 'O'),
# ('.', 'O')]
#
# **1.4** Find out the number of words in the longest sentence, and store it to variable ```max_len```. **Print your results for `max_len`.**
#
# **1.5** It's now time to convert the sentences data in a suitable format for the RNNs training/evaluation procedures. Create a ```word2idx``` dictionary mapping distinct words from the dataset into distinct integers. Also create a ```idx2word``` dictionary.
#
# **1.6** Prepare the predictors matrix ```X```, as a list of lists, where each inner list is a sequence of words mapped into integers accordly to the ```word2idx``` dictionary.
#
# **1.7** Apply the keras ```pad_sequences``` function to standardize the predictors. You should retrieve a matrix with all padded sentences and length equal to ```max_len``` previously computed. The dimensionality should therefore be equal to ```[# of sentences, max_len]```. Run the provided cell to print your results. Your ```X[i]``` now should be something similar to this:
#
# `[ 8193 27727 31033 33289 22577 33464 23723 16665 33464 31142 31319 28267
# 27700 33246 28646 16052 21 16915 17349 7924 32879 32985 18238 23555
# 24 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178]`
#
# **1.8** Create a ```tag2idx``` dictionary mapping distinct named entity tags from the dataset into distinct integers. Also create a ```idx2tag``` dictionary.
#
# **1.9** Prepare targets matrix ```Y```, as a list of lists,where each inner list is a sequence of tags mapped into integers accordly to the ```tag2idx``` dictionary.
#
# **1.10** apply the keras ```pad_sequences``` function to standardize the targets. Inject the ```PAD``` tag for the padding words. You should retrieve a matrix with all padded sentences'tags and length equal to ```max_length``` previously computed.
#
# **1.11** Use the ```to_categorical``` keras function to one-hot encode the tags. Now your ```Y``` should have dimension ```[# of sentences, max_len, n_tags]```. Run the provided cell to print your results.
#
# **1.12** Split the dataset into train and test sets (test 10%).
# + [markdown] colab_type="text" id="hBtmANNuuS6h"
# ## Answers
# -
# **1.1** Create a list of unique words found in the 'Word' column and sort it in alphabetic order. Then append the special word "ENDPAD" to the end of the list, and assign it to the variable ```words```. Store the length of this list as ```n_words```. **Print your results for `n_words`**
word_extract = list(data.Word)
words = list(set(word_extract))
words.sort()
words.append("ENDPAD")
n_words = len(words)
# Run this cell to show your results for n_words
print(n_words)
# **1.2** Create a list of unique tags and sort it in alphabetic order. Then append the special word "PAD" to the end of the list, and assign it to the variable ```tags```. Store the length of this list as ```n_tags```. **Print your results for `n_tags`**
tag_extract = list(data.Tag)
tags = list(set(tag_extract))
tags.sort()
tags.append("PAD")
n_tags = len(tags)
# Run this cell to show your results for n_tags
print(n_tags)
# **1.3** Process the data into a list of sentences where each sentence is a list of (word, tag) tuples. Here is an example of how the first sentence in the list should look:
#
# [('Thousands', 'O'),
# ('of', 'O'),
# ('demonstrators', 'O'),
# ('have', 'O'),
# ('marched', 'O'),
# ('through', 'O'),
# ('London', 'B-geo'),
# ('to', 'O'),
# ('protest', 'O'),
# ('the', 'O'),
# ('war', 'O'),
# ('in', 'O'),
# ('Iraq', 'B-geo'),
# ('and', 'O'),
# ('demand', 'O'),
# ('the', 'O'),
# ('withdrawal', 'O'),
# ('of', 'O'),
# ('British', 'B-gpe'),
# ('troops', 'O'),
# ('from', 'O'),
# ('that', 'O'),
# ('country', 'O'),
# ('.', 'O')]
#
data['word_tag'] = list(zip(data.Word, data.Tag)) # create tuple of words and tags
data.head()
sent_tag_list = data.word_tag.groupby(data['Sentence #']).apply(list)
print(f'Length of the list: {len(sent_tag_list)}\n')
print(f'First sentence:\n {(sent_tag_list[0])}')
# **1.4** Find out the number of words in the longest sentence, and store it to variable ```max_len```. **Print your results for `max_len`.**
max_len = data['Sentence #'].value_counts().sort_values(ascending = False)[0]
# Run this cell to show your results for max_len
print(max_len)
# **1.5** It's now time to convert the sentences data in a suitable format for the RNNs training/evaluation procedures. Create a ```word2idx``` dictionary mapping distinct words from the dataset into distinct integers. Also create a ```idx2word``` dictionary.
word2idx = dict(zip(words, range(len(words))))
print(f'Length of idx2word: {len(word2idx)}')
print(f'1st 10 items in the dictionary: {list(word2idx.items())[:10]}')
idx2word = dict(enumerate(words))
print(f'Length of word2idx: {len(idx2word)}')
print(f'1st 10 items in the dictionary: {list(idx2word.items())[:10]}')
# **1.6** Prepare the predictors matrix ```X```, as a list of lists, where each inner list is a sequence of words mapped into integers accordly to the ```word2idx``` dictionary.
# +
sent_word_list = list(data['Word'].groupby(data['Sentence #']).apply(list)) # create list of lists for sentences
temp_check = len(sent_word_list) == len(data['Sentence #'].value_counts())
print(f'Length equals number of sentences in provided df: {temp_check}')
print(f'1st sentence below: \n{sent_word_list[1]}')
# +
x = []
for i in range(len(sent_word_list)):
# get index values from word2idx dictionary and append to a list
x.append(list(map(word2idx.get, sent_word_list[i])))
X = x.copy() # Predictor matrix
# -
# **1.7** Apply the keras ```pad_sequences``` function to standardize the predictors. You should retrieve a matrix with all padded sentences and length equal to ```max_len``` previously computed. The dimensionality should therefore be equal to ```[# of sentences, max_len]```. Run the provided cell to print your results. Your ```X[i]``` now should be something similar to this:
#
# `[ 8193 27727 31033 33289 22577 33464 23723 16665 33464 31142 31319 28267
# 27700 33246 28646 16052 21 16915 17349 7924 32879 32985 18238 23555
# 24 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178 35178
# 35178 35178 35178 35178 35178 35178 35178 35178]`
X = pad_sequences(x, maxlen = max_len, padding = 'post', value = len(words)-1)
# Run this cell to show your results #
print("The index of word 'Harvard' is: {}\n".format(word2idx["Harvard"]))
print("Sentence 1: {}\n".format(X[1]))
print(X.shape)
# **1.8** Create a ```tag2idx``` dictionary mapping distinct named entity tags from the dataset into distinct integers. Also create a ```idx2tag``` dictionary.
# +
tag2idx = dict(zip(tags, range(len(tags))))
print(f'Length of tag2idx: {len(tag2idx)}')
print(f'1st 10 items in the dictionary: {list(tag2idx.items())[:10]}\n')
idx2tag = dict(zip(range(len(tags)), tags))
print(f'Length of idx2word: {len(idx2tag)}')
print(f'1st 10 items in the dictionary: {list(idx2tag.items())[:10]}')
# -
# **1.9** Prepare targets matrix ```Y```, as a list of lists,where each inner list is a sequence of tags mapped into integers accordly to the ```tag2idx``` dictionary.
# +
sent_tagged = list(data.Tag.groupby(data['Sentence #']).apply(list))
y = []
for i in range(len(sent_tagged)):
# Lookup tag values from each dictionary and append to target list y
y.append(list(map(tag2idx.get, sent_tagged[i])))
temp_check = len(y) == len(data['Sentence #'].value_counts())
print(f'Length equals number of sentences in provided df: {temp_check}')
# -
# **1.10** apply the keras ```pad_sequences``` function to standardize the targets. Inject the ```PAD``` tag for the padding words. You should retrieve a matrix with all padded sentences'tags and length equal to ```max_length``` previously computed.
# +
y_train = pad_sequences(y, maxlen = max_len, padding = 'post', value = len(tags)-1)
temp_check = len(y_train) == len(data['Sentence #'].value_counts())
print(f'Length equals number of sentences in provided df: {temp_check}')
# -
# **1.11** Use the ```to_categorical``` keras function to one-hot encode the tags. Now your ```Y``` should have dimension ```[# of sentences, max_len, n_tags]```. Run the provided cell to print your results.
Y = to_categorical(y_train)
# Run this cell to show your results #
print("The index of tag 'B-gpe' is: {}\n".format(tag2idx["B-gpe"]))
print("The tag of the last word in Sentence 1: {}\n".format(Y[0][-1]))
print(np.array(Y).shape)
# **1.12** Split the dataset into train and test sets (test 10%).
X_tr, X_te, y_tr, y_te = train_test_split(X, Y, test_size = 0.1, random_state = 123)
X_tr.shape, y_tr.shape, X_te.shape, y_te.shape
# ## Part 2: Modelling
#
# After preparing the train and test sets, we are ready to build five models:
# * frequency-based baseline
# * vanilla feedforward neural network
# * recurrent neural network
# * gated recurrent neural network
# * bidirectional gated recurrent neural network
#
# More details are given about architecture in each model's section. The input/output dimensionalities will be the same for all models:
# * input: ```[# of sentences, max_len]```
# * output: ```[# of sentences, max_len, n_tags]```
#
# Follow the information in each model's section to set up the architecture of each model. And the end of each training, use the given ```store_model``` function to store the weights and architectures in the ```./models``` path for later testing;```load_keras_model()``` is also provided to you
#
# A further ```plot_training_history``` helper function is given in case you need to check the training history.
#
# +
# Store model
def store_keras_model(model, model_name):
model_json = model.to_json() # serialize model to JSON
with open("./models/{}.json".format(model_name), "w") as json_file:
json_file.write(model_json)
model.save_weights("./models/{}.h5".format(model_name)) # serialize weights to HDF5
print("Saved model to disk")
# Plot history
def plot_training_history(history):
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1,len(loss)+1)
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('epoch')
plt.legend()
plt.show()
# -
# Load model
def load_keras_model(model_name):
# Load json and create model
json_file = open('./models/{}.json'.format(model_name), 'r')
loaded_model_json = json_file.read()
json_file.close()
model = tf.keras.models.model_from_json(loaded_model_json)
# Load weights into new model
model.load_weights("./models/{}.h5".format(model_name))
return model
# <div class='exercise'><b>Question 2: Models [40 points total]</b></div>
#
# **2.1** **Model 1: Baseline Model**
#
# Predict the tag of a word simply with the most frequently-seen named entity tag of this word from the training set.
#
# e.g. word "Apple" appears 10 times in the training set; 7 times it was tagged as "Corporate" and 3 times it was tagged as "Fruit". If we encounter the word "Apple" in the test set, we predict it as "Corporate".
#
# **Create an np.array ```baseline``` of length [n_words]**
# where the ith element ```baseline[i]``` is the index of the most commonly seen named entity tag of word i summarised from training set. (e.g. [16, 16, 16, ..., 0, 16, 16])
#
#
# **2.2** **Model 2: Vanilla Feed Forward Neural Network**
#
# This model is provided for you. Please pay attention to the architecture of this neural network, especially the input/output dimensionalities and the Embedding layer.
#
#
# **2.2a** Explain what the embedding layer is and why we need it here.
#
# **2.2b** Explain why the Param # of Embedding layer is 1758950 (as shown in `print(model.summary())`).
#
# **2.3** **Model 3: RNN**
#
# Set up a simple RNN model by stacking the following layers in sequence:
#
# an input layer
# a simple Embedding layer transforming integer words into vectors
# a dropout layer to regularize the model
# a SimpleRNN layer
# a TimeDistributed layer with an inner Dense layer which output dimensionality is equal to n_tag
#
# *(For hyperparameters, use those provided in Model 2)*
#
# **2.3a** Define, compile, and train an RNN model. Use the provided code to save the model and plot the training history.
#
# **2.3b** Visualize outputs from the SimpleRNN layer, one subplot for B-tags and one subplot for I-tags. Comment on the patterns you observed.
#
# **2.4** **Model 4: GRU**
#
# **2.4a** Briefly explain what a GRU is and how it's different from a simple RNN.
#
# **2.4b** Define, compile, and train a GRU architecture by replacing the SimpleRNN cell with a GRU one. Use the provided code to save the model and plot the training history.
#
# **2.4c** Visualize outputs from GRU layer, one subplot for **B-tags** and one subplot for **I-tags**. Comment on the patterns you observed.
#
# **2.5** **Model 5: Bidirectional GRU**
#
# **2.5a** Explain how a Bidirectional GRU differs from GRU model above.
#
# **2.5b** Define, compile, and train a bidirectional GRU by wrapping your GRU layer in a Bidirectional one. Use the provided code to save the model and plot the training history.
#
# **2.5c** Visualize outputs from bidirectional GRU layer, one subplot for **B-tags** and one subplot for **I-tags**. Comment on the patterns you observed.
# + [markdown] colab_type="text" id="hBtmANNuuS6h"
# ## Answers
# -
# **2.1** **Model 1: Baseline Model**
#
# Predict the tag of a word simply with the most frequently-seen named entity tag of this word from the training set.
#
# e.g. word "Apple" appears 10 times in the training set; 7 times it was tagged as "Corporate" and 3 times it was tagged as "Fruit". If we encounter the word "Apple" in the test set, we predict it as "Corporate".
#
# **Create an np.array ```baseline``` of length [n_words]**
# where the ith element ```baseline[i]``` is the index of the most commonly seen named entity tag of word i summarised from training set. (e.g. [16, 16, 16, ..., 0, 16, 16])
#
# +
baseline_unique_wordidx = np.unique(X_tr) # list of unique words in training set
baseline_unique_word = list(map(idx2word.get, baseline_unique_wordidx)) # map numbers to words
print(f'1st 5 unique words in the training set: {baseline_unique_word[:5]}')
# -
# Create df with words and max tags
base_grouped = data.groupby('Word').Tag.max().reset_index()
print(base_grouped.shape)
base_grouped.head()
# +
# Convert to dictionary for lookup
base_grouped_dict = dict(zip(base_grouped.Word, base_grouped.Tag))
# map words to tags and convert to list
baseline_tag = list(map(base_grouped_dict.get, baseline_unique_word))
temp_check = len(baseline_tag) == len(baseline_unique_wordidx)
print(f'Length of baseline tags in training set equals number of unique words: {temp_check}\n')
print(f'1st 5 tags of traning set:\n {baseline_tag[:5]}')
# -
# address shape mismatch here due to all words not in traning set
for word in words:
if word not in baseline_unique_word:
baseline_tag.append('O')
# define baseline
baseline = np.array(list(map(tag2idx.get, baseline_tag)))
baseline[255:260]
# Run this cell to show your results #
print(baseline[X].shape,'\n')
print('Sentence:\n {}\n'.format([idx2word[w] for w in X[0]]))
print('Predicted Tags:\n {}'.format([idx2tag[i] for i in baseline[X[0]]]))
# **2.2** **Model 2: Vanilla Feed Forward Neural Network**
#
# This model is provided for you. Please pay attention to the architecture of this neural network, especially the input/output dimensionalities and the Embedding layer.
#
# ### Use these hyperparameters for all NN models
# +
n_units = 100
drop_rate = .1
dim_embed = 50
optimizer = "rmsprop"
loss = "categorical_crossentropy"
metrics = ["accuracy"]
batch_size = 32
epochs = 10
validation_split = 0.1
verbose = 1
# +
# Define model
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(input_dim=n_words, output_dim=dim_embed, input_length=max_len))
model.add(tf.keras.layers.Dropout(drop_rate))
model.add(tf.keras.layers.Dense(n_tags, activation="softmax"))
# Compile model
model.compile(optimizer=optimizer, loss=loss, metrics=metrics)
# -
print(model.summary())
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs,
validation_split=validation_split, verbose=verbose)
store_keras_model(model, 'model_FFNN')
plot_training_history(history)
# **2.2a** Explain what the embedding layer is and why we need it here.
# Embeddings convert the given inputs indexes into dense vectors that perform better than sparse matrices that result from text being replaced by indices of unique words.
#
# This heps improve performance since it also captures relationships between words as well.
# **2.2b** Explain why the Param # of Embedding layer is 1758950 (as shown in `print(model.summary())`).
# This is the number of inputs words - 35179 multiplied by the dimension of the embedding 50.
# ### Viewing Hidden Layers
# In addition to the final result, we also want to see the intermediate results from hidden layers. Below is an example showing how to get outputs from a hidden layer, and visualize them on the reduced dimension of 2D by PCA. (**Please note that this code and the parameters cannot be simply copied and pasted for other questions; some adjustments need to be made**)
# +
FFNN = load_keras_model("model_FFNN")
def create_truncated_model_FFNN(trained_model):
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(input_dim=n_words, output_dim=dim_embed, input_length=max_len))
model.add(tf.keras.layers.Dropout(drop_rate))
# set weights of first few layers using the weights of trained model
for i, layer in enumerate(model.layers):
layer.set_weights(trained_model.layers[i].get_weights())
model.compile(optimizer=optimizer, loss=loss, metrics=metrics)
return model
truncated_model = create_truncated_model_FFNN(FFNN)
hidden_features = truncated_model.predict(X_te)
# flatten data
hidden_features = hidden_features.reshape(-1,50)
# find first two PCA components
pca = PCA(n_components=2)
pca_result = pca.fit_transform(hidden_features)
print('Variance explained by PCA: {}'.format(np.sum(pca.explained_variance_ratio_)))
# -
# visualize hidden features on first two PCA components
# this plot only shows B-tags
def visualize_hidden_features(pca_result):
color=['r', 'C1', 'y', 'C3', 'b', 'g', 'm', 'orange']
category = np.argmax(y_te, axis=1)
fig, ax = plt.subplots()
fig.set_size_inches(6,6)
for cat in range(8):
indices_B = np.where(category==cat)[0]
#length=min(1000,len(indices_B))
#indices_B=indices_B[:length]
ax.scatter(pca_result[indices_B,0], pca_result[indices_B, 1], label=idx2tag[cat],s=2,color=color[cat],alpha=0.5)
legend=ax.legend(markerscale=3)
legend.get_frame().set_facecolor('w')
plt.show()
visualize_hidden_features(pca_result)
# ### Full function for other questions ###
def get_hidden_output_PCA(model,X_te,y_te,layer_index,out_dimension):
output = tf.keras.backend.function([model.layers[0].input],[model.layers[layer_index].output])
hidden_feature=np.array(output([X_te]))
hidden_feature=hidden_feature.reshape(-1,out_dimension)
pca = PCA(n_components=2)
pca_result = pca.fit_transform(hidden_feature)
print('Variance explained by PCA: {}'.format(np.sum(pca.explained_variance_ratio_)))
return pca_result
def visualize_B_I(pca_result):
color = ['r', 'C1', 'y', 'C3', 'b', 'g', 'm', 'orange']
category = np.argmax(y_te.reshape(-1,18), axis=1)
fig, ax = plt.subplots(1,2)
fig.set_size_inches(12,6)
for i in range(2):
for cat in range(8*i,8*(i+1)):
indices = np.where(category==cat)[0]
ax[i].scatter(pca_result[indices,0], pca_result[indices, 1], label=idx2tag[cat],s=2,color=color[cat-8*i],alpha=0.5)
legend = ax[i].legend(markerscale=3)
legend.get_frame().set_facecolor('w')
ax[i].set_xlabel("first dimension")
ax[i].set_ylabel("second dimension")
fig.suptitle("visualization of hidden feature on reduced dimension by PCA")
plt.show()
h = get_hidden_output_PCA(FFNN,X_te,y_te,1,50)
visualize_B_I(h)
# **2.3** **Model 3: RNN**
#
# Set up a simple RNN model by stacking the following layers in sequence:
#
# an input layer
# a simple Embedding layer transforming integer words into vectors
# a dropout layer to regularize the model
# a SimpleRNN layer
# a TimeDistributed layer with an inner Dense layer which output dimensionality is equal to n_tag
#
# *(For hyperparameters, use those provided in Model 2)*
#
# **2.3a** Define, compile, and train an RNN model. Use the provided code to save the model and plot the training history.
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(100, return_sequences = True))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# -
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs,
validation_split=validation_split, verbose=verbose)
# save your mode #
store_keras_model(model, 'model_RNN')
# run this cell to show your results #
print(model.summary())
plot_training_history(history)
# **2.3b** Visualize outputs from the SimpleRNN layer, one subplot for B-tags and one subplot for I-tags. Comment on the patterns you observed.
# +
RNN = load_keras_model("model_RNN")
h = get_hidden_output_PCA(RNN,X_te,y_te,1,50)
visualize_B_I(h)
# -
# ### Comment
#
# * B-per and I-per tags appear to be dominating the PCA while other classes are overlapped. While the 1st 2 principal components explain 98% of the variation this could be on account of class unbalance.
# **2.4** **Model 4: GRU**
#
# **2.4a** Briefly explain what a GRU is and how it's different from a simple RNN.
# * GRU - Gradient Recurring Unit is a more sophisticated implementation of an RNN.
#
# * RNN has a simple hidden state while a GRU has a reset gate and an update gate.
# **2.4b** Define, compile, and train a GRU architecture by replacing the SimpleRNN cell with a GRU one. Use the provided code to save the model and plot the training history.
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(GRU(100, return_sequences = True))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# -
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs,
validation_split=validation_split, verbose=verbose)
# save your model #
store_keras_model(model, 'model_GRU')
# run this cell to show your results #
print(model.summary())
# run this cell to show your results #
plot_training_history(history)
# **2.4c** Visualize outputs from GRU layer, one subplot for **B-tags** and one subplot for **I-tags**. Comment on the patterns you observed.
# +
GRU = load_keras_model("model_GRU")
h = get_hidden_output_PCA(GRU,X_te,y_te,1,50)
visualize_B_I(h)
# -
# ### Comment
#
# * The modeal starts to overfit from training history chart.
#
# * Variance explained by PCA drops to 96%
#
# * Classes now appear to be more equally distributed for 'B' tags while the I-per tag still dominates. We believe that this could still be on account of the unbalanced which still needs to be addressed.
# **2.5** **Model 5: Bidirectional GRU**
#
# **2.5a** Explain how a Bidirectional GRU differs from GRU model above.
#
#
# Bidirectional GRU takes into account the previous input as well as the subsequent input. This is useful for cases such as language translation wherein we need the previous and subsequent words for context.
# **2.5b** Define, compile, and train a bidirectional GRU by wrapping your GRU layer in a Bidirectional one. Use the provided code to save the model and plot the training history.
#
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(Bidirectional(GRU(100, return_sequences = True)))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# -
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs,
validation_split=validation_split, verbose=verbose)
# save your model #
store_keras_model(model, 'model_BiGRU')
# run this cell to show your results #
print(model.summary())
# run this cell to show your results #
plot_training_history(history)
# **2.5c** Visualize outputs from bidirectional GRU layer, one subplot for **B-tags** and one subplot for **I-tags**. Comment on the patterns you observed.
# +
BGRU = load_keras_model("model_BiGRU")
h = get_hidden_output_PCA(BGRU,X_te,y_te,1,50)
visualize_B_I(h)
# -
# ### Comment
#
# * We can see from the plot_history that the model continues to overfit since the validation curve starts to curve upwards while the training curve is trending downwards.
#
# * This is suppoerted by the fact that the overall variance explained by PCA also drops from the 90s in earlier charts to 86% here.
#
# * From the PCA chart we see that the B-per tags are overlapping while hte I-per tag continues to dominate.
#
# * Values are also less spread out as compared to the previous charts.
# <div class='exercise'><b> Question 3: Analysis [40pt]</b></div>
#
# **3.1** For each model, iteratively:
#
# - Load the model using the given function ```load_keras_model```
#
# - Apply the model to the test dataset
#
# - Compute an F1 score for each ```Tag``` and store it
#
# **3.2** Plot the F1 score per Tag and per model making use of a grouped barplot.
#
# **3.3** Briefly discuss the performance of each model
#
#
# **3.4** Which tags have the lowest f1 score? For instance, you may find from the plot above that the test accuracy on "B-art", and "I-art" are very low (just an example, your case maybe different). Here is an example when models failed to predict these tags right
#
# <img src="data/B_art.png" alt="drawing" width="600"/>
#
# **3.5** Write functions to output another example in which the tags of the lowest accuracy was predicted wrong in a sentence (include both "B-xxx" and "I-xxx" tags). Store the results in a DataFrame (same format as the above example) and use styling functions below to print out your df.
#
# **3.6** Choose one of the most promising models you have built, improve this model to achieve an f1 score higher than 0.8 for as many tags as possible (you have lots of options here, e.g. data balancing, hyperparameter tuning, changing the structure of NN, a different optimizer, etc.)
#
# **3.7** Explain why you chose to change certain elements of the model and how effective these adjustments were.
#
# + [markdown] colab_type="text" id="hBtmANNuuS6h"
# ## Answers
# -
# **3.1** For each model, iteratively:
#
# - Load the model using the given function ```load_keras_model```
#
# - Apply the model to the test dataset
#
# - Compute an F1 score for each ```Tag``` and store it
# +
# define lists for each model
# Have not listed the baseline model here since the question asks us to use the load_keras_model
f1_ffnn = []
f1_rnn = []
f1_gru = []
f1_bgru = []
f1_list = [f1_ffnn, f1_rnn, f1_gru, f1_bgru]
type(f1_list)
# -
y_te_2d = np.argmax(y_te, axis = 2)
print(y_te_2d.shape)
print(y_te_2d[1])
# +
# Note - I'm sure there is a better way to do this so any guidance here would be appreciated.
# https://stackoverflow.com/questions/33326704/scikit-learn-calculate-f1-in-multilabel-classification
from sklearn.preprocessing import MultiLabelBinarizer
m = MultiLabelBinarizer().fit(y_te_2d)
model_list = [item for item in os.listdir('./models/') if '.json' in item]
for i in range(len(model_list)):
print(i)
print(model_list[i][:-5])
model = load_keras_model(model_list[i][:-5])
pred = model.predict_classes(X_te, verbose = 1)
print(pred.shape)
f = f1_score(m.transform(y_te_2d), m.transform(pred), average = None)
print(type(f), len(f))
f1_list[i].append(f)
print(type(f1_list[i]), len(f1_list[i]),'\n')
# -
# **3.2** Plot the F1 score per Tag and per model making use of a grouped barplot.
f1_df = pd.DataFrame(list(zip(f1_ffnn[0].T, f1_rnn[0].T, f1_gru[0].T, f1_bgru[0].T)), columns = ['FFNN', 'RNN', 'GRU', 'BGRU'], index = tags)
print(f1_df.shape)
f1_df.head()
# +
# https://chrisalbon.com/python/data_visualization/matplotlib_grouped_bar_plot/
# set posiiton and limits
width = 0.25
pos = np.arange(f1_df.shape[0])*2
# define the object create plot
fig, ax = plt.subplots(figsize=(15,8))
plt.bar(pos, f1_df.FFNN, width, alpha = 0.5, color = 'r')
plt.bar([p+width for p in pos], f1_df.RNN, width, alpha = 0.5, color = 'b')
plt.bar([p+width*2 for p in pos], f1_df.GRU, width, alpha = 0.5, color = 'g')
plt.bar([p+width*3 for p in pos], f1_df.BGRU, width, alpha = 0.5, color = 'y' )
# Set labels, ticks and other parameters
ax.set_ylabel('F1_Score')
ax.set_title('Model wise F1 Score')
ax.set_xticks([p + 1.5 * width for p in pos])
ax.set_xticklabels(f1_df.index)
plt.xlim(min(pos)-width, max(pos)+width*4)
# Adding the legend and showing the plot
plt.legend(['FFNN', 'RNN', 'GRU', 'BGRU'], loc='best')
plt.grid()
plt.show()
# -
# **3.3** Briefly discuss the performance of each model
# * All models have high accuray for most of the tags
# * FFNN has low accuracy for I-art, I-nat and I-tim tags
# * RNN and GRU have 0 values for I-art and I-nat tags
# * We should try to fine tune a less complex model such as FFNN or RNN since the results are mostly similar.
# **3.4** Which tags have the lowest f1 score? For instance, you may find from the plot above that the test accuracy on "B-art", and "I-art" are very low (just an example, your case maybe different). Here is an example when models failed to predict these tags right
#
# <img src="data/B_art.png" alt="drawing" width="600"/>
# #### Comment
#
# * B-art, I-art have the lowest scores
# * I-eve and I-nat have higher scores than the tags above but the scores are much lower relative to the other tags.
# **3.5** Write functions to output another example in which the tags of the lowest accuracy was predicted wrong in a sentence (include both "B-xxx" and "I-xxx" tags). Store the results in a DataFrame (same format as the above example) and use styling functions below to print out your df.
def highlight_errors(s):
is_max = s == s.y_true
return ['' if v or key=='Word' else 'color: red' for key,v in is_max.iteritems()]
# +
# choose random x value and get word indexes
sample = 50
x_sample = X_te[sample]
#print(f'Shape of sample value: {x_sample.shape}')
x_sent = [x for x in x_sample if x != 35178] # remove padding
#print(len(x_sent), type(x_sent))
# get word from index
x_word = list(map(idx2word.get, x_sent))
#x_word
# define sample sentence output; get index values from vector and remove padding
y_sample = y_te[sample]
#print(y_sample.shape)
y_true = np.argmax(y_sample, axis = 1)
y_true = [y for y in y_true if y != 17]
#print(len(y_true), type(y_true))
# get baseline prediction
y_baseline = baseline[x_sent]
#len(y_baseline)
# get FFNN prediction
y_ffnn = FFNN.predict_classes(np.array(x_sample).reshape(1,-1))
y_ffnn = y_ffnn[0][:len(x_sent)]
#len(y_ffnn)
# get RNN prediction
y_rnn = RNN.predict_classes(np.array(x_sample).reshape(1,-1))
y_rnn = y_rnn[0][:len(x_sent)]
#len(y_rnn)
# get GRU prediction
y_gru = GRU.predict_classes(np.array(x_sample).reshape(1,-1))
y_gru = y_gru[0][:len(x_sent)]
#len(y_gru)
# get bidirectional GRU prediction
y_bgru = BGRU.predict_classes(np.array(x_sample).reshape(1,-1))
y_bgru = y_bgru[0][:len(x_sent)]
#len(y_bgru)
# +
df2 = pd.DataFrame(list(zip(x_word, y_true, y_baseline, y_ffnn, y_rnn, y_gru, y_bgru)), columns = ['Word', 'y_true', 'y_baseline', 'y_ffnn', 'y_rnn', 'y_gru', 'y_bgru'])
print(df2.head())
df2[['y_true', 'y_baseline', 'y_ffnn', 'y_rnn', 'y_gru', 'y_bgru']] = df2[['y_true', 'y_baseline', 'y_ffnn', 'y_rnn', 'y_gru', 'y_bgru']].applymap(idx2tag.get)
df2.head()
# -
df2.style.applymap(highlight_errors) #, axis = 1) #, inplace = True)
df2
# **3.6** Choose one of the most promising models you have built, improve this model to achieve an f1 score higher than 0.8 for as many tags as possible (you have lots of options here, e.g. data balancing, hyperparameter tuning, changing the structure of NN, a different optimizer, etc.)
# #### Approach
#
# * We make 4 different attempts here
# * Increase dropout to 0.20
# * Increase dim_embed to 100
# * Add 1 layer of RNN and droput
# * Add didirectional RNN layer
# * Earlystopping is also addedd to check overfitting
# * f1_score plots for all attempts are also plotted at the end
# #### Approach 1 - Increase dropout to 0.20
# +
# Increase dropout to 0.20
n_units = 100
drop_rate = .20 # changed from 0.10 to 0.20
dim_embed = 50
optimizer = "rmsprop"
loss = "categorical_crossentropy"
metrics = ["accuracy"]
batch_size = 32
epochs = 10
validation_split = 0.1
verbose = 1
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(100, return_sequences = True))
#model.add(Dropout(drop_rate))
#model.add(SimpleRNN(50, return_sequences = True))
#model.add(Bidirectional(SimpleRNN(50, return_sequences = True)))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# +
# earlystopping callback
callbacks = [
EarlyStopping(
# Stop training when `val_accuracy` is no longer improving
monitor='val_accuracy',
# "no longer improving" being further defined as "for at least 2 epochs"
patience=2,
verbose=1)
]
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs, callbacks = callbacks,
validation_split=validation_split, verbose=verbose)
# -
pred = model.predict_classes(X_te, verbose = 0)
print(pred.shape)
f1_drop20 = f1_score(m.transform(y_te_2d), m.transform(pred), average = None)
f1_drop20.shape
f1mod_df = pd.DataFrame(list(zip(f1_rnn[0].T, f1_drop20.T)), columns = ['RNN_base','Drop20'], index = tags)
print(f1mod_df.shape)
f1mod_df.head()
# #### Approach 2 - Change dim_embed from 50 to 100
# +
# change dim-embed from 50 to 100
n_units = 100
drop_rate = .10
dim_embed = 100 # 50
optimizer = "rmsprop"
loss = "categorical_crossentropy"
metrics = ["accuracy"]
batch_size = 32
epochs = 10
validation_split = 0.1
verbose = 1
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(100, return_sequences = True))
#model.add(Dropout(drop_rate))
#model.add(SimpleRNN(50, return_sequences = True))
#model.add(Bidirectional(SimpleRNN(50, return_sequences = True)))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# +
# earlystopping callback
callbacks = [
EarlyStopping(
# Stop training when `val_accuracy` is no longer improving
monitor='val_accuracy',
# "no longer improving" being further defined as "for at least 2 epochs"
patience=2,
verbose=1)
]
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs, callbacks = callbacks,
validation_split=validation_split, verbose=verbose)
# -
pred = model.predict_classes(X_te, verbose = 0)
print(pred.shape)
f1_embed100 = f1_score(m.transform(y_te_2d), m.transform(pred), average = None)
f1_embed100.shape
f1mod_df['Embed100'] = f1_embed100
f1mod_df.head()
# #### Approach 3 - Add dropuout and simple RNN layers
# +
n_units = 100
drop_rate = .10
dim_embed = 50
optimizer = "rmsprop"
loss = "categorical_crossentropy"
metrics = ["accuracy"]
batch_size = 32
epochs = 10
validation_split = 0.1
verbose = 1
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(100, return_sequences = True))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(50, return_sequences = True))
#model.add(Bidirectional(SimpleRNN(50, return_sequences = True)))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# +
# earlystopping callback
callbacks = [
EarlyStopping(
# Stop training when `val_accuracy` is no longer improving
monitor='val_accuracy',
# "no longer improving" being further defined as "for at least 2 epochs"
patience=2,
verbose=1)
]
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs, callbacks = callbacks,
validation_split=validation_split, verbose=verbose)
# -
pred = model.predict_classes(X_te, verbose = 0)
print(pred.shape)
f1_dense_rnn = f1_score(m.transform(y_te_2d), m.transform(pred), average = None)
f1_dense_rnn.shape
f1mod_df['denseRNN'] = f1_dense_rnn
f1mod_df.head()
# #### Approach 4 - Add 1 Bidirectional layer
# +
n_units = 100
drop_rate = .10
dim_embed = 50
optimizer = "rmsprop"
loss = "categorical_crossentropy"
metrics = ["accuracy"]
batch_size = 32
epochs = 10
validation_split = 0.1
verbose = 1
# +
# define model
model = Sequential()
model.add(Embedding(input_dim = n_words, output_dim = dim_embed, input_length = max_len))
model.add(Dropout(drop_rate))
model.add(SimpleRNN(100, return_sequences = True))
model.add(Dropout(drop_rate))
#model.add(SimpleRNN(50, return_sequences = True))
model.add(Bidirectional(SimpleRNN(50, return_sequences = True)))
model.add(TimeDistributed(Dense(n_tags, activation = 'softmax')))
# compile model
model.compile(optimizer = optimizer, loss = loss, metrics = metrics )
# +
# earlystopping callback
callbacks = [
EarlyStopping(
# Stop training when `val_accuracy` is no longer improving
monitor='val_accuracy',
# "no longer improving" being further defined as "for at least 2 epochs"
patience=2,
verbose=1)
]
# Train model
history = model.fit(X_tr, y_tr, batch_size=batch_size, epochs=epochs, callbacks = callbacks,
validation_split=validation_split, verbose=verbose)
# -
pred = model.predict_classes(X_te, verbose = 0)
print(pred.shape)
f1_bi_rnn = f1_score(m.transform(y_te_2d), m.transform(pred), average = None)
f1_bi_rnn.shape
f1mod_df['BiD_RNN'] = f1_bi_rnn
f1mod_df.head()
# ### Plot
# +
# https://chrisalbon.com/python/data_visualization/matplotlib_grouped_bar_plot/
# set posiiton and limits
width = 0.25
pos = np.arange(f1_df.shape[0])*2
# define the object create plot
fig, ax = plt.subplots(figsize=(10,10))
plt.bar(pos, f1mod_df.RNN_base, width, alpha = 0.5, color = 'r')
plt.bar([p+width for p in pos], f1mod_df.RNN_base, width, alpha = 0.5, color = 'b')
plt.bar([p+width*2 for p in pos], f1mod_df.Drop20, width, alpha = 0.5, color = 'g')
plt.bar([p+width*3 for p in pos], f1mod_df.Embed100, width, alpha = 0.5, color = 'y' )
plt.bar([p+width*3 for p in pos], f1mod_df.denseRNN, width, alpha = 0.5, color = 'y' )
plt.bar([p+width*4 for p in pos], f1mod_df.BiD_RNN, width, alpha = 0.5, color = 'orange' )
# Set labels, ticks and other parameters
ax.set_ylabel('F1_Score')
ax.set_title('Model wise F1 Score')
ax.set_xticks([p + 1.5 * width for p in pos])
ax.set_xticklabels(f1_df.index)
plt.xlim(min(pos)-width, max(pos)+width*4)
# Adding the legend and showing the plot
#plt.legend(['RNN', 'Modified'], loc='best')
plt.grid()
plt.show()
# -
# **3.7** Explain why you chose to change certain elements of the model and how effective these adjustments were.
# #### Comment
#
# * We make 4 different attempts here
# * Increase dropout to 0.20 - To explore if increased layers being dropped can bring focus to the unbalanced classes.
# * Increase dim_embed to 100 - More dense embeddings may bring deeper relationships to the fore
# * Add 1 layer of RNN and droput - More deep embeddings may bring deeper relationships to the fore
# * Add bidirectional RNN layer - Greater feedback from before and after to provide sentence context for beter tagging
# * Earlystopping is also addedd to check overfitting
#
# +
# END
| content/homeworks/hw06/cs109b_hw6-submisison-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GeoData
import ipyleaflet as ipy
import geopandas
import json
m = ipy.Map(center=(52.3,8.0), zoom = 3, basemap= ipy.basemaps.Esri.WorldTopoMap)
countries = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
cities = geopandas.read_file(geopandas.datasets.get_path('naturalearth_cities'))
rivers = geopandas.read_file("https://www.naturalearthdata.com/http//www.naturalearthdata.com/download/10m/physical/ne_10m_rivers_lake_centerlines.zip")
# ## GeoDataFrame
countries.head()
geo_data = ipy.GeoData(geo_dataframe = countries,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'Countries')
m.add_layer(geo_data)
m.add_control(ipy.LayersControl())
m
rivers_data = ipy.GeoData(geo_dataframe = rivers,
style={'color': 'purple', 'opacity':3, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'Rivers')
m.add_layer(rivers_data)
# +
#geo_data.geo_dataframe = cities
# +
#m.remove_layer(geo_data)
# -
# ## Continent
africa_countries = countries[countries['continent'] == 'Africa']
africa_countries.head()
africa_data = ipy.GeoData(geo_dataframe = africa_countries,
style={'color': 'black', 'fillColor': 'red', 'opacity':7, 'weight':1.9, 'dashArray':'7', 'fillOpacity':0.2},
hover_style={'fillColor': 'grey', 'fillOpacity': 0.6},
name = 'Africa')
m.add_layer(africa_data)
m
# +
#m.remove_layer(africa_data)
# -
m.add_control(ipy.FullScreenControl())
| examples/GeoData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
pi = np.pi
for x in range(0,360):
y=np.sin(pi*x/180)
print(x,y)
import matplotlib.pyplot as plt
import numpy as np
x=np.linspace(0,10,100)
plt.plot(x, np.sin(x))
plt.plot(x,np.cos(x))
plt.show
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ml-training] *
# language: python
# name: conda-env-ml-training-py
# ---
# # Encoder / Decoder
#
# <img src="img/encoder-decoder.png" style="width: 95%; margin-left: auto; margin-right: auto;"/>
# # Transformer
# <img src="img/transformer.png" style="width: 45%; margin-left: auto; margin-right: auto;"/>
# ## Multi-Head Attention
#
# <img src="img/multi-head-attention.png" style="width: 35%; margin-left: auto; margin-right: auto;"/>
# ## Scaled Dot-Product Attention
#
# <img src="img/scaled-dot-prod-attention.png" style="width: 30%; margin-left: auto; margin-right: auto;"/>
# !nvidia-smi
from transformers import GPT2LMHeadModel, GPT2TokenizerFast
pretrained_weights = 'gpt2'
tokenizer = GPT2TokenizerFast.from_pretrained(pretrained_weights)
model = GPT2LMHeadModel.from_pretrained(pretrained_weights)
ids = tokenizer.encode('This is an example of text, and')
ids
tokenizer.decode(ids)
import torch
t = torch.LongTensor(ids).unsqueeze(0)
preds = model.generate(t)
preds.shape, preds[0]
tokenizer.decode(preds[0].numpy())
preds[:, -10:]
preds2 = model.generate(preds[:, -10:])
torch.cat((preds[0, :-10], preds2[0]))
tokenizer.decode(torch.cat((preds[0, :-10], preds2[0])).numpy())
from fastai.text.all import *
path = untar_data(URLs.WIKITEXT_TINY)
path.ls()
df_train = pd.read_csv(path/'train.csv', header=None)
df_valid = pd.read_csv(path/'test.csv', header=None)
df_train.head()
all_texts = np.concatenate([df_train[0].values, df_valid[0].values])
class TransformersTokenizer(Transform):
def __init__(self, tokenizer):
self.tokenizer = tokenizer
def encodes(self, x):
toks = self.tokenizer.tokenize(x)
return tensor(self.tokenizer.convert_tokens_to_ids(toks))
def decodes(self, x):
return TitledStr(self.tokenizer.decode(x.cpu().numpy()))
splits = [range_of(df_train), list(range(len(df_train), len(all_texts)))]
tfmd_lists = TfmdLists(all_texts, TransformersTokenizer(tokenizer), splits=splits, dl_type=LMDataLoader)
tfmd_lists.train[0],tfmd_lists.valid[0]
tfmd_lists.tfms(tfmd_lists.train.items[0]).shape, tfmd_lists.tfms(tfmd_lists.valid.items[0]).shape
batch_size, seq_len = 6, 1024
dls = tfmd_lists.dataloaders(bs=batch_size, seq_len=seq_len)
dls.show_batch(max_n=2)
# +
def tokenize(text):
toks = tokenizer.tokenize(text)
return tensor(tokenizer.convert_tokens_to_ids(toks))
tokenized = [tokenize(t) for t in progress_bar(all_texts)]
# -
class TransformersTokenizer(Transform):
def __init__(self, tokenizer): self.tokenizer = tokenizer
def encodes(self, x):
return x if isinstance(x, Tensor) else tokenize(x)
def decodes(self, x):
return TitledStr(self.tokenizer.decode(x.cpu().numpy()))
tfmd_lists = TfmdLists(tokenized, TransformersTokenizer(tokenizer), splits=splits, dl_type=LMDataLoader)
dls = tfmd_lists.dataloaders(bs=batch_size, seq_len=seq_len)
dls.show_batch(max_n=2)
class DropOutput(Callback):
def after_pred(self): self.learn.pred = self.learn.pred[0]
learn = Learner(dls, model, loss_func=CrossEntropyLossFlat(), cbs=[DropOutput], metrics=Perplexity()).to_fp16()
learn.validate()
learn.lr_find()
learn.fit_one_cycle(1, 1e-4)
df_valid.head(1)
prompt = "\n = Unicorn = \n \n A unicorn is a magical creature with a rainbow tail and a horn"
prompt_ids = tokenizer.encode(prompt)
inp = tensor(prompt_ids)[None].cuda()
inp.shape
preds = learn.model.generate(inp, max_length=40, num_beams=5, temperature=1.5)
preds[0]
tokenizer.decode(preds[0].cpu().numpy())
| 050 Transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib - Data Visualisation with 2D Arrays
# +
# Importing the required packages
from dateutil import parser
from time import time
import datetime as dt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sb
import math
# Generate the plots and charts within this window instead of opening a new window for each chart/plot
# %matplotlib inline
# -
# # 1. Data Visualisation with 2D Arrays
# +
u = np.linspace(-2, 2, 3)
v = np.linspace(-1, 1, 5)
X , Y = np.meshgrid(u, v)
Z = X**2/25 + Y**2/4
print("Z: \n", Z)
plt.set_cmap('gray')
plt.pcolor(Z)
plt.show()
# +
# Orientations of 2D Arrays and images
Z = np.array([[1,2,3],[4,5,6]])
print("Z: \n", Z)
plt.pcolor(Z)
plt.show()
# -
# Psuedocolor plot
u = np.linspace(-2, 2 , 65)
v = np.linspace (-1, 1, 33)
X, Y = np.meshgrid(u, v)
Z = X**2/25 + Y**2/4
plt.pcolor(Z)
plt.colorbar()
plt.set_cmap("viridis")
plt.show()
# color map
u = np.linspace(-2, 2 , 65)
v = np.linspace (-1, 1, 33)
X, Y = np.meshgrid(u, v)
Z = X**2/25 + Y**2/4
plt.pcolor(Z, cmap = "autumn")
plt.axis("tight")
plt.colorbar()
plt.show()
# Plot using mesh grid
u = np.linspace(-2, 2 , 65)
v = np.linspace (-1, 1, 33)
X, Y = np.meshgrid(u, v)
Z = X**2/25 + Y**2/4
plt.pcolor(X, Y , Z) # X, Y are 2D Meshgrid
plt.colorbar()
plt.show()
# Contour plots - Used when data is used continuously
plt.axes([1,1,0.7,1]) # Adjust the size of the figure
plt.contour(Z, 30) # Draw 30 contours - number of contours passed in as args
plt.show()
# Contour plots - Used when data is used continuously
plt.axes([1,1,0.7,1])
plt.contour(X, Y,Z, 30) # Draw 30 contours - number of contours passed in as args
plt.show()
# Filled contour plots
# Contour plots - Used when data is used continuously
plt.axes([1,1,0.8,1])
plt.contourf(X, Y,Z, 30) # Draw 30 contours - number of contours passed in as args
plt.colorbar()
plt.show()
# +
# Create a filled contour plot with a color map of 'viridis'
plt.subplot(2,2,1)
plt.contourf(X,Y,Z,20, cmap='viridis')
plt.colorbar()
plt.title('Viridis')
# Create a filled contour plot with a color map of 'gray'
plt.subplot(2,2,2)
plt.contourf(X,Y,Z,20, cmap='gray')
plt.colorbar()
plt.title('Gray')
# Create a filled contour plot with a color map of 'autumn'
plt.subplot(2,2,3)
plt.contourf(X, Y, Z, 20, cmap="autumn")
plt.colorbar()
plt.title('Autumn')
# Create a filled contour plot with a color map of 'winter'
plt.subplot(2,2,4)
plt.contourf(X, Y, Z, 20, cmap="winter")
plt.colorbar()
plt.title('Winter')
# Improve the spacing between subplots and display them
plt.tight_layout()
plt.show()
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# sphinx_gallery_thumbnail_number = 2
vegetables = ["cucumber", "tomato", "lettuce", "asparagus",
"potato", "wheat", "barley"]
farmers = ["<NAME>", "Upland Bros.", "<NAME>",
"Agrifun", "Organiculture", "BioGoods Ltd.", "Cornylee Corp."]
harvest = np.array([[0.8, 2.4, 2.5, 3.9, 0.0, 4.0, 0.0],
[2.4, 0.0, 4.0, 1.0, 2.7, 0.0, 0.0],
[1.1, 2.4, 0.8, 4.3, 1.9, 4.4, 0.0],
[0.6, 0.0, 0.3, 0.0, 3.1, 0.0, 0.0],
[0.7, 1.7, 0.6, 2.6, 2.2, 6.2, 0.0],
[1.3, 1.2, 0.0, 0.0, 0.0, 3.2, 5.1],
[0.1, 2.0, 0.0, 1.4, 0.0, 1.9, 6.3]])
fig, ax = plt.subplots()
im = ax.imshow(harvest)
# We want to show all ticks...
ax.set_xticks(np.arange(len(farmers)))
ax.set_yticks(np.arange(len(vegetables)))
# ... and label them with the respective list entries
ax.set_xticklabels(farmers)
ax.set_yticklabels(vegetables)
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
for i in range(len(vegetables)):
for j in range(len(farmers)):
text = ax.text(j, i, harvest[i, j],
ha="center", va="center", color="w")
ax.set_title("Harvest of local farmers (in tons/year)")
fig.tight_layout()
plt.show();
| Datacamp Assignments/Data Science Track/Matplotlib/Matplotlib - Data Visualisation with 2D Arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Document Image Classification using Transfer Learning from VGG16 (pretrained with ImageNet dataset)
#
# This exercise classifies document image files, using pre-trained ConvNet VGG16. The project was conducted on private dataset. Print output is removed to protect provicy.
#
#
# ### Reference:
#
# Binary class image recognition using Transfer Learning:
# https://blog.keras.io/
# https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html
#
# Multi-class image recognition using Transfer Learning:
# https://www.codesofinterest.com/2017/08/bottleneck-features-multi-class-classification-keras.html
#
# Graph Model Traning History in Keras:
# https://www.codesofinterest.com/2017/03/graph-model-training-history-keras.html
#
# ## Transfer Learning
#
# Transfer learning has become popular recently in computer vision, when there is not enough labled data to train a full neural network. It is observed that the lower layers of a neural network (such as CNN) are responsible to automatically extract basic visual features such as lines and shapes etc, thus a pre-trained NN can be "transferred" to recogenize different images with higher layers re-trained on the intented images.
# +
#Using Bottleneck Features for Multi-Class Classification in Keras
import numpy as np
from keras.preprocessing.image import ImageDataGenerator, img_to_array, load_img
from keras.models import Sequential
from keras.layers import Dropout, Flatten, Dense
from keras import applications
from keras.utils.np_utils import to_categorical
import matplotlib.pyplot as plt
import math
#import cv2
# -
# ### Dataset
#
# Dataset is pre-split into training and valiation, that are under directory "data/train" and "data/validation" corespondingly
# +
# dimensions of our images when loading.
img_width, img_height = 224, 224
top_model_weights_path = 'bottleneck_fc_model.h5'
train_data_dir = 'data/Document_PNG/train'
validation_data_dir = 'data/Document_PNG/validation'
# number of epochs to train top model
epochs = 50
# batch size used by flow_from_directory and predict_generator
batch_size = 16
# -
# The basic technique of transfer learning is to get a pre-trained model (with the weights loaded) and remove final fully-connected layers from that model. We then use the remaining portion of the model as a feature extractor for our smaller dataset. These extracted features are called "Bottleneck Features". Then train the last fully-connected network block on those extracted bottleneck features in order to get the classes we need as outputs for our problem.
# <img src="VGG16.png">
# The data is split into training and validation. In each of training and validation directory, the class (category) should be used as sub-directory name. Under each sub-directory, there are the image files that belongs to that class.
def create_generator(root_path):
datagen = ImageDataGenerator(rescale=1. / 255)
generator = datagen.flow_from_directory(
root_path,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode=None,
shuffle=False)
# Comment out print statement to protect privacy
#generator.filenames contains all the filenames
#print('total number of samples = {0}'.format(len(generator.filenames)))
# generator.class_indices is the map/dictionary for the class-names and their indexes
#print('number of categories= {0}'.format(len(generator.class_indices)))
#print('\ncategory vs. index mapping:')
#print(generator.class_indices)
return generator
train_generator = create_generator(train_data_dir)
# +
validation_generator = create_generator(validation_data_dir)
# +
def save_bottlebeck_features(train_generator, validation_generator):
# build the VGG16 network, use the weights trained on imagenet data
model = applications.VGG16(include_top=False, weights='imagenet')
nb_train_samples = len(train_generator.filenames)
num_classes = len(train_generator.class_indices)
predict_size_train = int(math.ceil(nb_train_samples / batch_size))
bottleneck_features_train = model.predict_generator(
train_generator, predict_size_train)
np.save('bottleneck_features_train.npy', bottleneck_features_train)
nb_validation_samples = len(validation_generator.filenames)
predict_size_validation = int(
math.ceil(nb_validation_samples / batch_size))
bottleneck_features_validation = model.predict_generator(
validation_generator, predict_size_validation)
np.save('bottleneck_features_validation.npy',
bottleneck_features_validation)
# -
save_bottlebeck_features(train_generator, validation_generator)
# +
def train_FC_model():
datagen_top = ImageDataGenerator(rescale=1. / 255)
generator_top = datagen_top.flow_from_directory(
train_data_dir,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode='categorical',
shuffle=False)
nb_train_samples = len(generator_top.filenames)
num_classes = len(generator_top.class_indices)
# save the class indices to use use later in predictions
np.save('class_indices.npy', generator_top.class_indices)
# load the bottleneck features saved earlier
train_data = np.load('bottleneck_features_train.npy')
# get the class lebels for the training data, in the original order
train_labels = generator_top.classes
# https://github.com/fchollet/keras/issues/3467
# convert the training labels to categorical vectors
train_labels = to_categorical(train_labels, num_classes=num_classes)
generator_top = datagen_top.flow_from_directory(
validation_data_dir,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode=None,
shuffle=False)
nb_validation_samples = len(generator_top.filenames)
validation_data = np.load('bottleneck_features_validation.npy')
validation_labels = generator_top.classes
validation_labels = to_categorical(
validation_labels, num_classes=num_classes)
model = Sequential()
model.add(Flatten(input_shape=train_data.shape[1:]))
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
#model.add(Dense(num_classes, activation='sigmoid')) # to get class prediction
model.add(Dense(num_classes, activation='softmax')) # to get probability prediction
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(train_data, train_labels,
epochs=epochs,
batch_size=batch_size,
validation_data=(validation_data, validation_labels))
model.save_weights(top_model_weights_path)
(eval_loss, eval_accuracy) = model.evaluate(
validation_data, validation_labels, batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(eval_accuracy * 100))
print("[INFO] Loss: {}".format(eval_loss))
plt.figure(1)
# summarize history for accuracy
plt.subplot(211)
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
# summarize history for loss
plt.subplot(212)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
train_FC_model()
def predict(image_path):
# load the class_indices saved in the earlier step
class_dictionary = np.load('class_indices.npy').item()
num_classes = len(class_dictionary)
#load and pre-process the image
#orig = cv2.imread(image_path)
print("[INFO] loading and preprocessing image...")
image = load_img(image_path, target_size=(224, 224))
image = img_to_array(image)
# Rescale the image, this is important, otherwise the predictions will be '0'
# This is because ImageDataGenerator set rescale=1. / 255,
# which means all data is re-scaled from a [0 - 255] range to [0 - 1.0]
image = image / 255
image = np.expand_dims(image, axis=0)
# build the VGG16 network
model = applications.VGG16(include_top=False, weights='imagenet')
# get the bottleneck prediction from the pre-trained VGG16 model
bottleneck_prediction = model.predict(image)
# build top FC model block
model = Sequential()
model.add(Flatten(input_shape=bottleneck_prediction.shape[1:]))
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
#model.add(Dense(num_classes, activation='sigmoid')) # to get class prediction
model.add(Dense(num_classes, activation='softmax')) # to get probability prediction
model.load_weights(top_model_weights_path)
# use the bottleneck prediction on the FC model to get the final classification
class_predicted = model.predict_classes(bottleneck_prediction)
#proba = model.predict_proba(bottleneck_prediction)
output_y = model.predict(bottleneck_prediction)
inID = class_predicted[0]
inv_map = {v: k for k, v in class_dictionary.items()}
label = inv_map[inID]
# get the prediction label
print("Prediction: class ID: {}, Label: {}".format(inID, label))
#print(proba)
predict1 = {}
for i in range(len(output_y[0])):
predict1[inv_map[i]] = output_y[0][i]
print(predict1)
print('sum of probability = {0}'.format(sum(output_y[0])))
# display the predictions with the image
'''
cv2.putText(orig, "Predicted: {}".format(label), (10, 30),
cv2.FONT_HERSHEY_PLAIN, 1.5, (43, 99, 255), 2)
cv2.imshow("Classification", orig)
cv2.waitKey(0)
cv2.destroyAllWindows()
'''
# +
# add the path to your test image below
image_path = './data/Document_PNG/validation/Bank Statement/Chase-bank-statement-2.png'
predict(image_path)
#cv2.destroyAllWindows()
# -
image_path = './data/Document_PNG/validation/W2/W2 C-20173-0.PNG'
predict(image_path)
| ImageClassifier/Document_Image_Transfer_Learning_VGG16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Start-to-Finish Example: [TOV](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation) Neutron Star Simulation: The "Hydro without Hydro" Test
#
# ## Authors: <NAME> & <NAME>
# ### Formatting improvements courtesy <NAME>
#
# ## This module sets up initial data for a neutron star on a spherical numerical grid, using the approach [documented in the previous NRPy+ module](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb), and then evolves these initial data forward in time. The aim is to reproduce the results from [Baumgarte, Hughes, and Shapiro]( https://arxiv.org/abs/gr-qc/9902024) (which were performed using Cartesian grids); demonstrating that the extrinsic curvature and Hamiltonian constraint violation converge to zero with increasing numerical resolution
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This module has been validated to exhibit convergence to zero of the Hamiltonian constraint violation at the expected order to the exact solution (see [plot](#convergence) at bottom). Note that convergence in the region causally influenced by the surface of the star will possess lower convergence order due to the sharp drop to zero in $T^{\mu\nu}$.
#
# ### NRPy+ Source Code for this module:
#
# * [TOV/TOV_Solver.py](../edit/TOV/TOV_Solver.py); ([**NRPy+ Tutorial module reviewing mathematical formulation and equations solved**](Tutorial-ADM_Initial_Data-TOV.ipynb)); ([**start-to-finish NRPy+ Tutorial module demonstrating that initial data satisfy Hamiltonian constraint**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb)): Tolman-Oppenheimer-Volkoff (TOV) initial data; defines all ADM variables and nonzero $T^{\mu\nu}$ components in Spherical basis.
# * [BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py](../edit/BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb): *Numerical* Spherical ADM$\to$Curvilinear BSSN converter function
# * [BSSN/BSSN_constraints.py](../edit/BSSN/BSSN_constraints.py); [\[**tutorial**\]](Tutorial-BSSN_constraints.ipynb): Hamiltonian constraint in BSSN curvilinear basis/coordinates
#
# ## Introduction:
# Here we use NRPy+ to evolve initial data for a [simple polytrope TOV star](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation), keeping the $T^{\mu\nu}$ source terms fixed. As the hydrodynamical fields that go into $T^{\mu\nu}$ are not updated, this is called the "Hydro without Hydro" test.
#
# The entire algorithm is outlined as follows, with links to the relevant NRPy+ tutorial notebooks listed at each step:
#
# 1. Allocate memory for gridfunctions, including temporary storage for the Method of Lines time integration [(**NRPy+ tutorial on NRPy+ Method of Lines algorithm**)](Tutorial-Method_of_Lines-C_Code_Generation.ipynb).
# 1. Set gridfunction values to initial data
# * [**NRPy+ tutorial on TOV initial data**](Tutorial-ADM_Initial_Data-TOV.ipynb)
# * [**NRPy+ tutorial on validating TOV initial data**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb).
# 1. Next, integrate the initial data forward in time using the Method of Lines coupled to a Runge-Kutta explicit timestepping algorithm:
# 1. At the start of each iteration in time, output the Hamiltonian constraint violation
# * [**NRPy+ tutorial on BSSN constraints**](Tutorial-BSSN_constraints.ipynb).
# 1. At each RK time substep, do the following:
# 1. Evaluate BSSN RHS expressions
# * [**NRPy+ tutorial on BSSN right-hand sides**](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb)
# * [**NRPy+ tutorial on BSSN gauge condition right-hand sides**](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb)
# * [**NRPy+ tutorial on adding stress-energy source terms to BSSN RHSs**](Tutorial-BSSN_stress_energy_source_terms.ipynb).
# 1. Apply singular, curvilinear coordinate boundary conditions [*a la* the SENR/NRPy+ paper](https://arxiv.org/abs/1712.07658)
# * [**NRPy+ tutorial on setting up singular, curvilinear boundary conditions**](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
# 1. Enforce constraint on conformal 3-metric: $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$
# * [**NRPy+ tutorial on enforcing $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint**](Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb)
# 1. Repeat above steps at two numerical resolutions to confirm convergence to zero.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Set core NRPy+ parameters for numerical grids and reference metric
# 1. [Step 1.a](#cfl) Output needed C code for finding the minimum proper distance between grid points, needed for [CFL](https://en.wikipedia.org/w/index.php?title=Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition&oldid=806430673)-limited timestep
# 1. [Step 2](#adm_id_tov): Set up ADM initial data for polytropic TOV Star
# 1. [Step 2.a](#tov_interp): Interpolate the TOV data file as needed to set up ADM spacetime quantities in spherical basis (for input into the `Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear` module) and $T^{\mu\nu}$ in the chosen reference metric basis
# 1. [Step 3](#adm_id_spacetime): Convert ADM spacetime quantity initial data to BSSN-in-curvilinear-coordinates
# 1. [Step 4](#bssn): Output C code for BSSN spacetime solve
# 1. [Step 4.a](#bssnrhs): Set up the BSSN right-hand-side (RHS) expressions, and add the *rescaled* $T^{\mu\nu}$ source terms
# 1. [Step 4.b](#hamconstraint): Output C code for Hamiltonian constraint
# 1. [Step 4.c](#enforce3metric): Enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$
# 1. [Step 4.d](#ccodegen): Generate C code kernels for BSSN expressions, in parallel if possible
# 1. [Step 4.e](#cparams_rfm_and_domainsize): Output C codes needed for declaring and setting Cparameters; also set `free_parameters.h`
# 1. [Step 5](#bc_functs): Set up boundary condition functions for chosen singular, curvilinear coordinate system
# 1. [Step 6](#mainc): `TOV_Playground.c`: The Main C Code
# 1. [Step 7](#visualize): Data Visualization Animations
# 1. [Step 7.a](#installdownload): Install `scipy` and download `ffmpeg` if they are not yet installed/downloaded
# 1. [Step 7.b](#genimages): Generate images for visualization animation
# 1. [Step 7.c](#genvideo): Generate visualization animation
# 1. [Step 8](#convergence): Validation: Convergence of numerical errors (Hamiltonian constraint violation) to zero
# 1. [Step 9](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Set core NRPy+ parameters for numerical grids and reference metric \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# +
# Step P1: Import needed NRPy+ core modules:
from outputC import * # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# Step P2: Create C code output directory:
Ccodesdir = os.path.join("BSSN_Hydro_without_Hydro_Ccodes/")
# First remove C code output directory if it exists
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
# # !rm -r ScalarWaveCurvilinear_Playground_Ccodes
shutil.rmtree(Ccodesdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesdir)
# Step P3: Create executable output directory:
outdir = os.path.join(Ccodesdir,"output/")
cmd.mkdir(outdir)
# Step 1: Set the spatial dimension parameter
# to three this time, and then read
# the parameter as DIM.
par.set_parval_from_str("grid::DIM",3)
DIM = par.parval_from_str("grid::DIM")
# Step 2: Set some core parameters, including CoordSystem MoL timestepping algorithm,
# FD order, floating point precision, and CFL factor:
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
CoordSystem = "Spherical"
# Step 2.a: Set defaults for Coordinate system parameters.
# These are perhaps the most commonly adjusted parameters,
# so we enable modifications at this high level.
# domain_size = 7.5 # SET BELOW BASED ON TOV STELLAR RADIUS
# sinh_width sets the default value for:
# * SinhSpherical's params.SINHW
# * SinhCylindrical's params.SINHW{RHO,Z}
# * SinhSymTP's params.SINHWAA
sinh_width = 0.4 # If Sinh* coordinates chosen
# sinhv2_const_dr sets the default value for:
# * SinhSphericalv2's params.const_dr
# * SinhCylindricalv2's params.const_d{rho,z}
sinhv2_const_dr = 0.05# If Sinh*v2 coordinates chosen
# SymTP_bScale sets the default value for:
# * SinhSymTP's params.bScale
SymTP_bScale = 0.5 # If SymTP chosen
# Step 2.b: Set the order of spatial and temporal derivatives;
# the core data type, and the CFL factor.
# RK_method choices include: Euler, "RK2 Heun", "RK2 MP", "RK2 Ralston", RK3, "RK3 Heun", "RK3 Ralston",
# SSPRK3, RK4, DP5, DP5alt, CK5, DP6, L6, DP8
RK_method = "RK4"
FD_order = 4 # Finite difference order: even numbers only, starting with 2. 12 is generally unstable
REAL = "double" # Best to use double here.
CFL_FACTOR= 0.5 # (GETS OVERWRITTEN WHEN EXECUTED.) In pure axisymmetry (symmetry_axes = 2 below) 1.0 works fine. Otherwise 0.5 or lower.
# Set the lapse & shift to be consistent with the original Hydro without Hydro paper.
LapseCondition = "HarmonicSlicing"
ShiftCondition = "Frozen"
# Step 3: Generate Runge-Kutta-based (RK-based) timestepping code.
# As described above the Table of Contents, this is a 3-step process:
# 3.A: Evaluate RHSs (RHS_string)
# 3.B: Apply boundary conditions (post_RHS_string, pt 1)
# 3.C: Enforce det(gammabar) = det(gammahat) constraint (post_RHS_string, pt 2)
import MoLtimestepping.C_Code_Generation as MoL
from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
RK_order = Butcher_dict[RK_method][1]
cmd.mkdir(os.path.join(Ccodesdir,"MoLtimestepping/"))
MoL.MoL_C_Code_Generation(RK_method,
RHS_string = """
Ricci_eval(&rfmstruct, ¶ms, RK_INPUT_GFS, auxevol_gfs);
rhs_eval(&rfmstruct, ¶ms, auxevol_gfs, RK_INPUT_GFS, RK_OUTPUT_GFS);""",
post_RHS_string = """
apply_bcs_curvilinear(¶ms, &bcstruct, NUM_EVOL_GFS, evol_gf_parity, RK_OUTPUT_GFS);
enforce_detgammabar_constraint(&rfmstruct, ¶ms, RK_OUTPUT_GFS);\n""",
outdir = os.path.join(Ccodesdir,"MoLtimestepping/"))
# Step 4: Set the coordinate system for the numerical grid
par.set_parval_from_str("reference_metric::CoordSystem",CoordSystem)
rfm.reference_metric() # Create ReU, ReDD needed for rescaling B-L initial data, generating BSSN RHSs, etc.
# Step 5: Set the finite differencing order to FD_order (set above).
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order)
# Step 6: Copy SIMD/SIMD_intrinsics.h to $Ccodesdir/SIMD/SIMD_intrinsics.h
cmd.mkdir(os.path.join(Ccodesdir,"SIMD"))
shutil.copy(os.path.join("SIMD/")+"SIMD_intrinsics.h",os.path.join(Ccodesdir,"SIMD/"))
# Step 7: Set the direction=2 (phi) axis to be the symmetry axis; i.e.,
# axis "2", corresponding to the i2 direction.
# This sets all spatial derivatives in the phi direction to zero.
par.set_parval_from_str("indexedexp::symmetry_axes","2")
# -
# <a id='cfl'></a>
#
# ## Step 1.a: Output needed C code for finding the minimum proper distance between grid points, needed for [CFL](https://en.wikipedia.org/w/index.php?title=Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition&oldid=806430673)-limited timestep \[Back to [top](#toc)\]
# $$\label{cfl}$$
#
# In order for our explicit-timestepping numerical solution to the scalar wave equation to be stable, it must satisfy the [CFL](https://en.wikipedia.org/w/index.php?title=Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition&oldid=806430673) condition:
# $$
# \Delta t \le \frac{\min(ds_i)}{c},
# $$
# where $c$ is the wavespeed, and
# $$ds_i = h_i \Delta x^i$$
# is the proper distance between neighboring gridpoints in the $i$th direction (in 3D, there are 3 directions), $h_i$ is the $i$th reference metric scale factor, and $\Delta x^i$ is the uniform grid spacing in the $i$th direction:
# Output the find_timestep() function to a C file.
rfm.out_timestep_func_to_file(os.path.join(Ccodesdir,"find_timestep.h"))
# <a id='adm_id_tov'></a>
#
# # Step 2: Set up ADM initial data for polytropic TOV Star \[Back to [top](#toc)\]
# $$\label{adm_id_tov}$$
#
# As documented [in the TOV Initial Data NRPy+ Tutorial Module](Tutorial-TOV_Initial_Data.ipynb) ([older version here](Tutorial-GRMHD_UnitConversion.ipynb)), we will now set up TOV initial data, storing the densely-sampled result to file (***Courtesy <NAME>***).
#
# The TOV solver uses an ODE integration routine provided by scipy, so we first make sure that scipy is installed:
# !pip install scipy > /dev/null
# Next we call the [TOV.TOV_Solver() function](../edit/TOV/TOV_Solver.py) ([NRPy+ Tutorial module](Tutorial-ADM_Initial_Data-TOV.ipynb)) to set up the initial data, using the default parameters for initial data. This function outputs the solution to a file named "outputTOVpolytrope.txt".
# +
############################
# Single polytrope example #
############################
import TOV.Polytropic_EOSs as ppeos
# Set neos = 1 (single polytrope)
neos = 1
# Set rho_poly_tab (not needed for a single polytrope)
rho_poly_tab = []
# Set Gamma_poly_tab
Gamma_poly_tab = [2.0]
# Set K_poly_tab0
K_poly_tab0 = 1. # ZACH NOTES: CHANGED FROM 100.
# Set the eos quantities
eos = ppeos.set_up_EOS_parameters__complete_set_of_input_variables(neos,rho_poly_tab,Gamma_poly_tab,K_poly_tab0)
import TOV.TOV_Solver as TOV
M_TOV, R_Schw_TOV, R_iso_TOV = TOV.TOV_Solver(eos,
outfile="outputTOVpolytrope.txt",
rho_baryon_central=0.129285,
return_M_RSchw_and_Riso = True,
verbose = True)
domain_size = 2.0 * R_iso_TOV
# -
# <a id='tov_interp'></a>
#
# ## Step 2.a: Interpolate the TOV data file as needed to set up ADM spacetime quantities in spherical basis (for input into the `Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear` module) and $T^{\mu\nu}$ in the chosen reference metric basis \[Back to [top](#toc)\]
# $$\label{tov_interp}$$
#
# The TOV data file just written stored $\left(r,\rho(r),P(r),M(r),e^{\nu(r)}\right)$, where $\rho(r)$ is the total mass-energy density (cf. $\rho_{\text{baryonic}}$).
#
# **METRIC DATA IN TERMS OF ADM QUANTITIES**
#
# The [TOV line element](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation) in *Schwarzschild coordinates* is written (in the $-+++$ form):
# $$
# ds^2 = - c^2 e^\nu dt^2 + \left(1 - \frac{2GM}{rc^2}\right)^{-1} dr^2 + r^2 d\Omega^2.
# $$
#
# In *isotropic coordinates* with $G=c=1$ (i.e., the coordinate system we'd prefer to use), the ($-+++$ form) line element is written:
# $$
# ds^2 = - e^{\nu} dt^2 + e^{4\phi} \left(d\bar{r}^2 + \bar{r}^2 d\Omega^2\right),
# $$
# where $\phi$ here is the *conformal factor*.
#
# The ADM 3+1 line element for this diagonal metric in isotropic spherical coordinates is given by:
# $$
# ds^2 = (-\alpha^2 + \beta_k \beta^k) dt^2 + \gamma_{\bar{r}\bar{r}} d\bar{r}^2 + \gamma_{\theta\theta} d\theta^2+ \gamma_{\phi\phi} d\phi^2,
# $$
#
# from which we can immediately read off the ADM quantities:
# \begin{align}
# \alpha &= e^{\nu(\bar{r})/2} \\
# \beta^k &= 0 \\
# \gamma_{\bar{r}\bar{r}} &= e^{4\phi}\\
# \gamma_{\theta\theta} &= e^{4\phi} \bar{r}^2 \\
# \gamma_{\phi\phi} &= e^{4\phi} \bar{r}^2 \sin^2 \theta \\
# \end{align}
#
# **STRESS-ENERGY TENSOR $T^{\mu\nu}$**
#
# We will also need the stress-energy tensor $T^{\mu\nu}$. [As discussed here](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation), the stress-energy tensor is diagonal:
#
# \begin{align}
# T^t_t &= -\rho \\
# T^i_j &= P \delta^i_j \\
# \text{All other components of }T^\mu_\nu &= 0.
# \end{align}
#
# Since $\beta^i=0$ the inverse metric expression simplifies to (Eq. 4.49 in [Gourgoulhon](https://arxiv.org/pdf/gr-qc/0703035.pdf)):
# $$
# g^{\mu\nu} = \begin{pmatrix}
# -\frac{1}{\alpha^2} & \frac{\beta^i}{\alpha^2} \\
# \frac{\beta^i}{\alpha^2} & \gamma^{ij} - \frac{\beta^i\beta^j}{\alpha^2}
# \end{pmatrix} =
# \begin{pmatrix}
# -\frac{1}{\alpha^2} & 0 \\
# 0 & \gamma^{ij}
# \end{pmatrix},
# $$
#
# and since the 3-metric is diagonal we get
#
# \begin{align}
# \gamma^{\bar{r}\bar{r}} &= e^{-4\phi}\\
# \gamma^{\theta\theta} &= e^{-4\phi}\frac{1}{\bar{r}^2} \\
# \gamma^{\phi\phi} &= e^{-4\phi}\frac{1}{\bar{r}^2 \sin^2 \theta}.
# \end{align}
#
# Thus raising $T^\mu_\nu$ yields a diagonal $T^{\mu\nu}$
#
# \begin{align}
# T^{tt} &= -g^{tt} \rho = \frac{1}{\alpha^2} \rho = e^{-\nu(\bar{r})} \rho \\
# T^{\bar{r}\bar{r}} &= g^{\bar{r}\bar{r}} P = \frac{1}{e^{4 \phi}} P \\
# T^{\theta\theta} &= g^{\theta\theta} P = \frac{1}{e^{4 \phi}\bar{r}^2} P\\
# T^{\phi\phi} &= g^{\phi\phi} P = \frac{1}{e^{4\phi}\bar{r}^2 \sin^2 \theta} P
# \end{align}
# +
thismodule = "HydrowithoutHydro"
rbar,theta,rho,P,expnu,exp4phi = par.Cparameters("REAL",thismodule,
["rbar","theta","rho","P","expnu","exp4phi"],1e300)
IDalpha = sp.sqrt(expnu)
gammaSphDD = ixp.zerorank2(DIM=3)
gammaSphDD[0][0] = exp4phi
gammaSphDD[1][1] = exp4phi*rbar**2
gammaSphDD[2][2] = exp4phi*rbar**2*sp.sin(theta)**2
T4SphUU = ixp.zerorank2(DIM=4)
T4SphUU[0][0] = rho/expnu
T4SphUU[1][1] = P/exp4phi
T4SphUU[2][2] = P/(exp4phi*rbar**2)
T4SphUU[3][3] = P/(exp4phi*rbar**2*sp.sin(theta)**2)
# +
expr_list = [IDalpha]
name_list = ["*alpha"]
for i in range(3):
for j in range(i,3):
expr_list.append(gammaSphDD[i][j])
name_list.append("*gammaDD"+str(i)+str(j))
desc = """This function takes as input either (x,y,z) or (r,th,ph) and outputs
all ADM quantities in the Cartesian or Spherical basis, respectively."""
name = "ID_TOV_ADM_quantities"
outCparams = "preindent=1,outCverbose=False,includebraces=False"
outCfunction(
outfile=os.path.join(Ccodesdir, name + ".h"), desc=desc, name=name,
params=""" const REAL xyz_or_rthph[3],
const ID_inputs other_inputs,
REAL *gammaDD00,REAL *gammaDD01,REAL *gammaDD02,REAL *gammaDD11,REAL *gammaDD12,REAL *gammaDD22,
REAL *KDD00,REAL *KDD01,REAL *KDD02,REAL *KDD11,REAL *KDD12,REAL *KDD22,
REAL *alpha,
REAL *betaU0,REAL *betaU1,REAL *betaU2,
REAL *BU0,REAL *BU1,REAL *BU2""",
body="""
// Set trivial metric quantities:
*KDD00 = *KDD01 = *KDD02 = 0.0;
/**/ *KDD11 = *KDD12 = 0.0;
/**/ *KDD22 = 0.0;
*betaU0 = *betaU1 = *betaU2 = 0.0;
*BU0 = *BU1 = *BU2 = 0.0;
// Next set gamma_{ij} in spherical basis
const REAL rbar = xyz_or_rthph[0];
const REAL theta = xyz_or_rthph[1];
const REAL phi = xyz_or_rthph[2];
REAL rho,rho_baryon,P,M,expnu,exp4phi;
TOV_interpolate_1D(rbar,other_inputs.Rbar,other_inputs.Rbar_idx,other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_Schw_arr,other_inputs.rho_arr,other_inputs.rho_baryon_arr,other_inputs.P_arr,other_inputs.M_arr,
other_inputs.expnu_arr,other_inputs.exp4phi_arr,other_inputs.rbar_arr,
&rho,&rho_baryon,&P,&M,&expnu,&exp4phi);\n"""+
outputC(expr_list,name_list, "returnstring",outCparams),
opts="DisableCparameters")
# -
# As all input quantities are functions of $r$, we will simply read the solution from file and interpolate it to the values of $r$ needed by the initial data.
#
# 1. First we define functions `ID_TOV_ADM_quantities()` and `ID_TOV_TUPMUNU()` that call the [1D TOV interpolator function](../edit/TOV/tov_interp.h) to evaluate the ADM spacetime quantities and $T^{\mu\nu}$, respectively, at any given point $(r,\theta,\phi)$ in the Spherical basis. All quantities are defined as above.
# 1. Next we will construct the BSSN/ADM source terms $\{S_{ij},S_{i},S,\rho\}$ in the Spherical basis
# 1. Then we will perform the Jacobian transformation on $\{S_{ij},S_{i},S,\rho\}$ to the desired `(xx0,xx1,xx2)` basis
# 1. Next we call the *Numerical* Spherical ADM$\to$Curvilinear BSSN converter function to conver the above ADM quantities to the rescaled BSSN quantities in the desired curvilinear coordinate system: [BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py](../edit/BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb).
#
# $$
# {\rm Jac\_dUSph\_dDrfmUD[mu][nu]} = \frac{\partial x^\mu_{\rm Sph}}{\partial x^\nu_{\rm rfm}},
# $$
#
# via exact differentiation (courtesy SymPy), and the inverse Jacobian
# $$
# {\rm Jac\_dUrfm\_dDSphUD[mu][nu]} = \frac{\partial x^\mu_{\rm rfm}}{\partial x^\nu_{\rm Sph}},
# $$
#
# using NRPy+'s `generic_matrix_inverter3x3()` function. In terms of these, the transformation of BSSN tensors from Spherical to `"reference_metric::CoordSystem"` coordinates may be written:
#
# $$
# T^{\mu\nu}_{\rm rfm} =
# \frac{\partial x^\mu_{\rm rfm}}{\partial x^\delta_{\rm Sph}}
# \frac{\partial x^\nu_{\rm rfm}}{\partial x^\sigma_{\rm Sph}} T^{\delta\sigma}_{\rm Sph}
# $$
# +
r_th_ph_or_Cart_xyz_oID_xx = []
CoordType_in = "Spherical"
if CoordType_in == "Spherical":
r_th_ph_or_Cart_xyz_oID_xx = rfm.xxSph
elif CoordType_in == "Cartesian":
r_th_ph_or_Cart_xyz_oID_xx = rfm.xxCart
else:
print("Error: Can only convert ADM Cartesian or Spherical initial data to BSSN Curvilinear coords.")
exit(1)
# Next apply Jacobian transformations to convert into the (xx0,xx1,xx2) basis
# rho and S are scalar, so no Jacobian transformations are necessary.
Jac4_dUSphorCart_dDrfmUD = ixp.zerorank2(DIM=4)
Jac4_dUSphorCart_dDrfmUD[0][0] = sp.sympify(1)
for i in range(DIM):
for j in range(DIM):
Jac4_dUSphorCart_dDrfmUD[i+1][j+1] = sp.diff(r_th_ph_or_Cart_xyz_oID_xx[i],rfm.xx[j])
Jac4_dUrfm_dDSphorCartUD, dummyDET = ixp.generic_matrix_inverter4x4(Jac4_dUSphorCart_dDrfmUD)
# Perform Jacobian operations on T^{mu nu} and gamma_{ij}
T4UU = ixp.register_gridfunctions_for_single_rank2("AUXEVOL","T4UU","sym01",DIM=4)
IDT4UU = ixp.zerorank2(DIM=4)
for mu in range(4):
for nu in range(4):
for delta in range(4):
for sigma in range(4):
IDT4UU[mu][nu] += \
Jac4_dUrfm_dDSphorCartUD[mu][delta]*Jac4_dUrfm_dDSphorCartUD[nu][sigma]*T4SphUU[delta][sigma]
lhrh_list = []
for mu in range(4):
for nu in range(mu,4):
lhrh_list.append(lhrh(lhs=gri.gfaccess("auxevol_gfs","T4UU"+str(mu)+str(nu)),rhs=IDT4UU[mu][nu]))
desc = """This function takes as input either (x,y,z) or (r,th,ph) and outputs
all ADM quantities in the Cartesian or Spherical basis, respectively."""
name = "ID_TOV_TUPMUNU_xx0xx1xx2"
outCparams = "preindent=1,outCverbose=False,includebraces=False"
outCfunction(
outfile=os.path.join(Ccodesdir, name + ".h"), desc=desc, name=name,
params="""const paramstruct *restrict params,REAL *restrict xx[3],
const ID_inputs other_inputs,REAL *restrict auxevol_gfs""",
body=outputC([rfm.xxSph[0],rfm.xxSph[1],rfm.xxSph[2]],
["const REAL rbar","const REAL theta","const REAL ph"],"returnstring",
"CSE_enable=False,includebraces=False")+"""
REAL rho,rho_baryon,P,M,expnu,exp4phi;
TOV_interpolate_1D(rbar,other_inputs.Rbar,other_inputs.Rbar_idx,other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_Schw_arr,other_inputs.rho_arr,other_inputs.rho_baryon_arr,other_inputs.P_arr,other_inputs.M_arr,
other_inputs.expnu_arr,other_inputs.exp4phi_arr,other_inputs.rbar_arr,
&rho,&rho_baryon,&P,&M,&expnu,&exp4phi);\n"""+
fin.FD_outputC("returnstring",lhrh_list,params="outCverbose=False,includebraces=False").replace("IDX4","IDX4S"),
loopopts="AllPoints,Read_xxs")
# -
# <a id='adm_id_spacetime'></a>
#
# # Step 3: Convert ADM initial data to BSSN-in-curvilinear coordinates \[Back to [top](#toc)\]
# $$\label{adm_id_spacetime}$$
#
# This is an automated process, taken care of by [`BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear`](../edit/BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py), and documented [in this tutorial notebook](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb).
import BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear as AtoBnum
AtoBnum.Convert_Spherical_or_Cartesian_ADM_to_BSSN_curvilinear("Spherical","ID_TOV_ADM_quantities",
Ccodesdir=Ccodesdir,loopopts="")
# <a id='bssn'></a>
#
# # Step 4: Output C code for BSSN spacetime solve \[Back to [top](#toc)\]
# $$\label{bssn}$$
#
# <a id='bssnrhs'></a>
#
# ## Step 4.a: Set up the BSSN right-hand-side (RHS) expressions, and add the *rescaled* $T^{\mu\nu}$ source terms \[Back to [top](#toc)\]
# $$\label{bssnrhs}$$
#
# `BSSN.BSSN_RHSs()` sets up the RHSs assuming a spacetime vacuum: $T^{\mu\nu}=0$. (This might seem weird, but remember that, for example, *spacetimes containing only single or binary black holes are vacuum spacetimes*.) Here, using the [`BSSN.BSSN_stress_energy_source_terms`](../edit/BSSN/BSSN_stress_energy_source_terms.py) ([**tutorial**](Tutorial-BSSN_stress_energy_source_terms.ipynb)) NRPy+ module, we add the $T^{\mu\nu}$ source terms to these equations.
# +
import time
import BSSN.BSSN_RHSs as rhs
import BSSN.BSSN_gauge_RHSs as gaugerhs
par.set_parval_from_str("BSSN.BSSN_gauge_RHSs::LapseEvolutionOption", LapseCondition)
par.set_parval_from_str("BSSN.BSSN_gauge_RHSs::ShiftEvolutionOption", ShiftCondition)
print("Generating symbolic expressions for BSSN RHSs...")
start = time.time()
# Enable rfm_precompute infrastructure, which results in
# BSSN RHSs that are free of transcendental functions,
# even in curvilinear coordinates, so long as
# ConformalFactor is set to "W" (default).
cmd.mkdir(os.path.join(Ccodesdir,"rfm_files/"))
par.set_parval_from_str("reference_metric::enable_rfm_precompute","True")
par.set_parval_from_str("reference_metric::rfm_precompute_Ccode_outdir",os.path.join(Ccodesdir,"rfm_files/"))
# Evaluate BSSN + BSSN gauge RHSs with rfm_precompute enabled:
import BSSN.BSSN_quantities as Bq
par.set_parval_from_str("BSSN.BSSN_quantities::LeaveRicciSymbolic","True")
rhs.BSSN_RHSs()
import BSSN.BSSN_stress_energy_source_terms as Bsest
Bsest.BSSN_source_terms_for_BSSN_RHSs(T4UU)
rhs.trK_rhs += Bsest.sourceterm_trK_rhs
for i in range(DIM):
# Needed for Gamma-driving shift RHSs:
rhs.Lambdabar_rhsU[i] += Bsest.sourceterm_Lambdabar_rhsU[i]
# Needed for BSSN RHSs:
rhs.lambda_rhsU[i] += Bsest.sourceterm_lambda_rhsU[i]
for j in range(DIM):
rhs.a_rhsDD[i][j] += Bsest.sourceterm_a_rhsDD[i][j]
gaugerhs.BSSN_gauge_RHSs()
# We use betaU as our upwinding control vector:
Bq.BSSN_basic_tensors()
betaU = Bq.betaU
import BSSN.Enforce_Detgammabar_Constraint as EGC
enforce_detg_constraint_symb_expressions = EGC.Enforce_Detgammabar_Constraint_symb_expressions()
# Next compute Ricci tensor
par.set_parval_from_str("BSSN.BSSN_quantities::LeaveRicciSymbolic","False")
Bq.RicciBar__gammabarDD_dHatD__DGammaUDD__DGammaU()
# Now register the Hamiltonian as a gridfunction.
H = gri.register_gridfunctions("AUX","H")
# Then define the Hamiltonian constraint and output the optimized C code.
import BSSN.BSSN_constraints as bssncon
bssncon.BSSN_constraints(add_T4UUmunu_source_terms=False)
Bsest.BSSN_source_terms_for_BSSN_constraints(T4UU)
bssncon.H += Bsest.sourceterm_H
# Now that we are finished with all the rfm hatted
# quantities in generic precomputed functional
# form, let's restore them to their closed-
# form expressions.
par.set_parval_from_str("reference_metric::enable_rfm_precompute","False") # Reset to False to disable rfm_precompute.
rfm.ref_metric__hatted_quantities()
end = time.time()
print("Finished BSSN symbolic expressions in "+str(end-start)+" seconds.")
def BSSN_RHSs():
print("Generating C code for BSSN RHSs in "+par.parval_from_str("reference_metric::CoordSystem")+" coordinates.")
start = time.time()
# Construct the left-hand sides and right-hand-side expressions for all BSSN RHSs
lhs_names = [ "alpha", "cf", "trK"]
rhs_exprs = [gaugerhs.alpha_rhs, rhs.cf_rhs, rhs.trK_rhs]
for i in range(3):
lhs_names.append( "betU"+str(i))
rhs_exprs.append(gaugerhs.bet_rhsU[i])
lhs_names.append( "lambdaU"+str(i))
rhs_exprs.append(rhs.lambda_rhsU[i])
lhs_names.append( "vetU"+str(i))
rhs_exprs.append(gaugerhs.vet_rhsU[i])
for j in range(i,3):
lhs_names.append( "aDD"+str(i)+str(j))
rhs_exprs.append(rhs.a_rhsDD[i][j])
lhs_names.append( "hDD"+str(i)+str(j))
rhs_exprs.append(rhs.h_rhsDD[i][j])
# Sort the lhss list alphabetically, and rhss to match.
# This ensures the RHSs are evaluated in the same order
# they're allocated in memory:
lhs_names,rhs_exprs = [list(x) for x in zip(*sorted(zip(lhs_names,rhs_exprs), key=lambda pair: pair[0]))]
# Declare the list of lhrh's
BSSN_evol_rhss = []
for var in range(len(lhs_names)):
BSSN_evol_rhss.append(lhrh(lhs=gri.gfaccess("rhs_gfs",lhs_names[var]),rhs=rhs_exprs[var]))
# Set up the C function for the BSSN RHSs
desc="Evaluate the BSSN RHSs"
name="rhs_eval"
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params = """rfm_struct *restrict rfmstruct,const paramstruct *restrict params,
const REAL *restrict auxevol_gfs,const REAL *restrict in_gfs,REAL *restrict rhs_gfs""",
body = fin.FD_outputC("returnstring",BSSN_evol_rhss, params="outCverbose=False,SIMD_enable=True",
upwindcontrolvec=betaU).replace("IDX4","IDX4S"),
loopopts = "InteriorPoints,EnableSIMD,Enable_rfm_precompute")
end = time.time()
print("Finished BSSN_RHS C codegen in " + str(end - start) + " seconds.")
def Ricci():
print("Generating C code for Ricci tensor in "+par.parval_from_str("reference_metric::CoordSystem")+" coordinates.")
start = time.time()
desc="Evaluate the Ricci tensor"
name="Ricci_eval"
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params = """rfm_struct *restrict rfmstruct,const paramstruct *restrict params,
const REAL *restrict in_gfs,REAL *restrict auxevol_gfs""",
body = fin.FD_outputC("returnstring",
[lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD00"),rhs=Bq.RbarDD[0][0]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD01"),rhs=Bq.RbarDD[0][1]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD02"),rhs=Bq.RbarDD[0][2]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD11"),rhs=Bq.RbarDD[1][1]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD12"),rhs=Bq.RbarDD[1][2]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","RbarDD22"),rhs=Bq.RbarDD[2][2])],
params="outCverbose=False,SIMD_enable=True").replace("IDX4","IDX4S"),
loopopts = "InteriorPoints,EnableSIMD,Enable_rfm_precompute")
end = time.time()
print("Finished Ricci C codegen in " + str(end - start) + " seconds.")
# -
# <a id='hamconstraint'></a>
#
# ## Step 4.b: Output the Hamiltonian constraint \[Back to [top](#toc)\]
# $$\label{hamconstraint}$$
#
# Next output the C code for evaluating the Hamiltonian constraint [(**Tutorial**)](Tutorial-BSSN_constraints.ipynb). In the absence of numerical error, this constraint should evaluate to zero. However it does not due to numerical (typically truncation and roundoff) error. We will therefore measure the Hamiltonian constraint violation to gauge the accuracy of our simulation, and, ultimately determine whether errors are dominated by numerical finite differencing (truncation) error as expected.
def Hamiltonian():
start = time.time()
print("Generating optimized C code for Hamiltonian constraint. May take a while, depending on CoordSystem.")
# Set up the C function for the Hamiltonian RHS
desc="Evaluate the Hamiltonian constraint"
name="Hamiltonian_constraint"
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params = """rfm_struct *restrict rfmstruct,const paramstruct *restrict params,
REAL *restrict in_gfs, REAL *restrict auxevol_gfs, REAL *restrict aux_gfs""",
body = fin.FD_outputC("returnstring",lhrh(lhs=gri.gfaccess("aux_gfs", "H"), rhs=bssncon.H),
params="outCverbose=False").replace("IDX4","IDX4S"),
loopopts = "InteriorPoints,Enable_rfm_precompute")
end = time.time()
print("Finished Hamiltonian C codegen in " + str(end - start) + " seconds.")
# <a id='enforce3metric'></a>
#
# ## Step 4.c: Enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint \[Back to [top](#toc)\]
# $$\label{enforce3metric}$$
#
# Then enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint (Eq. 53 of [<NAME>, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)), as [documented in the corresponding NRPy+ tutorial notebook](Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb)
#
# Applying curvilinear boundary conditions should affect the initial data at the outer boundary, and will in general cause the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint to be violated there. Thus after we apply these boundary conditions, we must always call the routine for enforcing the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint:
def gammadet():
start = time.time()
print("Generating optimized C code for gamma constraint. May take a while, depending on CoordSystem.")
# Set up the C function for the det(gammahat) = det(gammabar)
EGC.output_Enforce_Detgammabar_Constraint_Ccode(Ccodesdir,exprs=enforce_detg_constraint_symb_expressions)
end = time.time()
print("Finished gamma constraint C codegen in " + str(end - start) + " seconds.")
# <a id='ccodegen'></a>
#
# ## Step 4.d: Generate C code kernels for BSSN expressions, in parallel if possible \[Back to [top](#toc)\]
# $$\label{ccodegen}$$
# +
# Step 1: Create a list of functions we wish to evaluate in parallel
funcs = [BSSN_RHSs,Ricci,Hamiltonian,gammadet]
try:
if os.name == 'nt':
# It's a mess to get working in Windows, so we don't bother. :/
# https://medium.com/@grvsinghal/speed-up-your-python-code-using-multiprocessing-on-windows-and-jupyter-or-ipython-2714b49d6fac
raise Exception("Parallel codegen currently not available in Windows")
# Step 1.a: Import the multiprocessing module.
import multiprocessing
# Step 1.b: Define master function for parallelization.
# Note that lambdifying this doesn't work in Python 3
def master_func(arg):
funcs[arg]()
# Step 1.c: Evaluate list of functions in parallel if possible;
# otherwise fallback to serial evaluation:
pool = multiprocessing.Pool()
pool.map(master_func,range(len(funcs)))
except:
# Steps 1.b-1.c, alternate: As fallback, evaluate functions in serial.
for func in funcs:
func()
# -
# <a id='cparams_rfm_and_domainsize'></a>
#
# ## Step 4.e: Output C codes needed for declaring and setting Cparameters; also set `free_parameters.h` \[Back to [top](#toc)\]
# $$\label{cparams_rfm_and_domainsize}$$
#
# Based on declared NRPy+ Cparameters, first we generate `declare_Cparameters_struct.h`, `set_Cparameters_default.h`, and `set_Cparameters[-SIMD].h`.
#
# Then we output `free_parameters.h`, which sets initial data parameters, as well as grid domain & reference metric parameters, applying `domain_size` and `sinh_width`/`SymTP_bScale` (if applicable) as set above
# +
# Step 3.d.i: Generate declare_Cparameters_struct.h, set_Cparameters_default.h, and set_Cparameters[-SIMD].h
par.generate_Cparameters_Ccodes(os.path.join(Ccodesdir))
# Step 3.d.ii: Set free_parameters.h
# Output to $Ccodesdir/free_parameters.h reference metric parameters based on generic
# domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale,
# parameters set above.
rfm.out_default_free_parameters_for_rfm(os.path.join(Ccodesdir,"free_parameters.h"),
domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale)
# Step 1.c.ii: Generate set_Nxx_dxx_invdx_params__and__xx.h:
rfm.set_Nxx_dxx_invdx_params__and__xx_h(Ccodesdir)
# Step 1.c.iii: Generate xxCart.h, which contains xxCart() for
# (the mapping from xx->Cartesian) for the chosen
# CoordSystem:
rfm.xxCart_h("xxCart","./set_Cparameters.h",os.path.join(Ccodesdir,"xxCart.h"))
# Step 1.c.iv: Generate declare_Cparameters_struct.h, set_Cparameters_default.h, and set_Cparameters[-SIMD].h
par.generate_Cparameters_Ccodes(os.path.join(Ccodesdir))
# -
# <a id='bc_functs'></a>
#
# # Step 5: Set up boundary condition functions for chosen singular, curvilinear coordinate system \[Back to [top](#toc)\]
# $$\label{bc_functs}$$
#
# Next apply singular, curvilinear coordinate boundary conditions [as documented in the corresponding NRPy+ tutorial notebook](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
import CurviBoundaryConditions.CurviBoundaryConditions as cbcs
cbcs.Set_up_CurviBoundaryConditions(os.path.join(Ccodesdir,"boundary_conditions/"),Cparamspath=os.path.join("../"))
# <a id='mainc'></a>
#
# # Step 6: `Hydro_without_Hydro_Playground.c`: The Main C Code \[Back to [top](#toc)\]
# $$\label{mainc}$$
#
# +
# Part P0: Define REAL, set the number of ghost cells NGHOSTS (from NRPy+'s FD_CENTDERIVS_ORDER),
# and set the CFL_FACTOR (which can be overwritten at the command line)
with open(os.path.join(Ccodesdir,"Hydro_without_Hydro_Playground_REAL__NGHOSTS__CFL_FACTOR.h"), "w") as file:
file.write("""
// Part P0.a: Set the number of ghost cells, from NRPy+'s FD_CENTDERIVS_ORDER
#define NGHOSTS """+str(int(FD_order/2)+1)+"""
// Part P0.b: Set the numerical precision (REAL) to double, ensuring all floating point
// numbers are stored to at least ~16 significant digits
#define REAL """+REAL+"""
// Part P0.c: Set the number of ghost cells, from NRPy+'s FD_CENTDERIVS_ORDER
REAL CFL_FACTOR = """+str(CFL_FACTOR)+"""; // Set the CFL Factor. Can be overwritten at command line.
// Part P0.d: Set TOV stellar parameters
#define TOV_Mass """+str(M_TOV)+"""
#define TOV_Riso """+str(R_iso_TOV)+"\n")
# +
# %%writefile $Ccodesdir/Hydro_without_Hydro_Playground.c
// Step P0: Define REAL and NGHOSTS; and declare CFL_FACTOR. This header is generated in NRPy+.
#include "Hydro_without_Hydro_Playground_REAL__NGHOSTS__CFL_FACTOR.h"
#include "rfm_files/rfm_struct__declare.h"
#include "declare_Cparameters_struct.h"
// All SIMD intrinsics used in SIMD-enabled C code loops are defined here:
#include "SIMD/SIMD_intrinsics.h"
// Step P1: Import needed header files
#include "stdio.h"
#include "stdlib.h"
#include "math.h"
#include "time.h"
#include "stdint.h" // Needed for Windows GCC 6.x compatibility
#ifndef M_PI
#define M_PI 3.141592653589793238462643383279502884L
#endif
#ifndef M_SQRT1_2
#define M_SQRT1_2 0.707106781186547524400844362104849039L
#endif
#define wavespeed 1.0 // Set CFL-based "wavespeed" to 1.0.
// Step P2: Declare the IDX4S(gf,i,j,k) macro, which enables us to store 4-dimensions of
// data in a 1D array. In this case, consecutive values of "i"
// (all other indices held to a fixed value) are consecutive in memory, where
// consecutive values of "j" (fixing all other indices) are separated by
// Nxx_plus_2NGHOSTS0 elements in memory. Similarly, consecutive values of
// "k" are separated by Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1 in memory, etc.
#define IDX4S(g,i,j,k) \
( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) + Nxx_plus_2NGHOSTS2 * (g) ) ) )
#define IDX4ptS(g,idx) ( (idx) + (Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2) * (g) )
#define IDX3S(i,j,k) ( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) ) ) )
#define LOOP_REGION(i0min,i0max, i1min,i1max, i2min,i2max) \
for(int i2=i2min;i2<i2max;i2++) for(int i1=i1min;i1<i1max;i1++) for(int i0=i0min;i0<i0max;i0++)
#define LOOP_ALL_GFS_GPS(ii) _Pragma("omp parallel for") \
for(int (ii)=0;(ii)<Nxx_plus_2NGHOSTS_tot*NUM_EVOL_GFS;(ii)++)
// Step P3: Set UUGF and VVGF macros, as well as xxCart()
#include "boundary_conditions/gridfunction_defines.h"
// Step P4: Set xxCart(const paramstruct *restrict params,
// REAL *restrict xx[3],
// const int i0,const int i1,const int i2,
// REAL xCart[3]),
// which maps xx->Cartesian via
// {xx[0][i0],xx[1][i1],xx[2][i2]}->{xCart[0],xCart[1],xCart[2]}
#include "xxCart.h"
// Step P5: Defines set_Nxx_dxx_invdx_params__and__xx(const int EigenCoord, const int Nxx[3],
// paramstruct *restrict params, REAL *restrict xx[3]),
// which sets params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for
// the chosen Eigen-CoordSystem if EigenCoord==1, or
// CoordSystem if EigenCoord==0.
#include "set_Nxx_dxx_invdx_params__and__xx.h"
// Step P6: Include basic functions needed to impose curvilinear
// parity and boundary conditions.
#include "boundary_conditions/CurviBC_include_Cfunctions.h"
// Step P7: Implement the algorithm for upwinding.
// *NOTE*: This upwinding is backwards from
// usual upwinding algorithms, because the
// upwinding control vector in BSSN (the shift)
// acts like a *negative* velocity.
//#define UPWIND_ALG(UpwindVecU) UpwindVecU > 0.0 ? 1.0 : 0.0
// Step P8: Include function for enforcing detgammabar constraint.
#include "enforce_detgammabar_constraint.h"
// Step P9: Find the CFL-constrained timestep
#include "find_timestep.h"
// Step P4: Declare initial data input struct:
// stores data from initial data solver,
// so they can be put on the numerical grid.
typedef struct __ID_inputs {
REAL Rbar;
int Rbar_idx;
int interp_stencil_size;
int numlines_in_file;
REAL *r_Schw_arr,*rho_arr,*rho_baryon_arr,*P_arr,*M_arr,*expnu_arr,*exp4phi_arr,*rbar_arr;
} ID_inputs;
// Part P11: Declare all functions for setting up TOV initial data.
/* Routines to interpolate the TOV solution and convert to ADM & T^{munu}: */
#include "../TOV/tov_interp.h"
#include "ID_TOV_ADM_quantities.h"
#include "ID_TOV_TUPMUNU_xx0xx1xx2.h"
/* Next perform the basis conversion and compute all needed BSSN quantities */
#include "ID_ADM_xx0xx1xx2_to_BSSN_xx0xx1xx2__ALL_BUT_LAMBDAs.h"
#include "ID_BSSN__ALL_BUT_LAMBDAs.h"
#include "ID_BSSN_lambdas.h"
// Step P10: Declare function necessary for setting up the initial data.
// Step P10.a: Define BSSN_ID() for BrillLindquist initial data
// Step P10.b: Set the generic driver function for setting up BSSN initial data
void initial_data(const paramstruct *restrict params,const bc_struct *restrict bcstruct,
const rfm_struct *restrict rfmstruct,
REAL *restrict xx[3], REAL *restrict auxevol_gfs, REAL *restrict in_gfs) {
#include "set_Cparameters.h"
// Step 1: Set up TOV initial data
// Step 1.a: Read TOV initial data from data file
// Open the data file:
char filename[100];
sprintf(filename,"./outputTOVpolytrope.txt");
FILE *in1Dpolytrope = fopen(filename, "r");
if (in1Dpolytrope == NULL) {
fprintf(stderr,"ERROR: could not open file %s\n",filename);
exit(1);
}
// Count the number of lines in the data file:
int numlines_in_file = count_num_lines_in_file(in1Dpolytrope);
// Allocate space for all data arrays:
REAL *r_Schw_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rho_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rho_baryon_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *P_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *M_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *expnu_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *exp4phi_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rbar_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
// Read from the data file, filling in arrays
// read_datafile__set_arrays() may be found in TOV/tov_interp.h
if(read_datafile__set_arrays(in1Dpolytrope, r_Schw_arr,rho_arr,rho_baryon_arr,P_arr,M_arr,expnu_arr,exp4phi_arr,rbar_arr) == 1) {
fprintf(stderr,"ERROR WHEN READING FILE %s!\n",filename);
exit(1);
}
fclose(in1Dpolytrope);
REAL Rbar = -100;
int Rbar_idx = -100;
for(int i=1;i<numlines_in_file;i++) {
if(rho_arr[i-1]>0 && rho_arr[i]==0) { Rbar = rbar_arr[i-1]; Rbar_idx = i-1; }
}
if(Rbar<0) {
fprintf(stderr,"Error: could not find rbar=Rbar from data file.\n");
exit(1);
}
ID_inputs TOV_in;
TOV_in.Rbar = Rbar;
TOV_in.Rbar_idx = Rbar_idx;
const int interp_stencil_size = 12;
TOV_in.interp_stencil_size = interp_stencil_size;
TOV_in.numlines_in_file = numlines_in_file;
TOV_in.r_Schw_arr = r_Schw_arr;
TOV_in.rho_arr = rho_arr;
TOV_in.rho_baryon_arr = rho_baryon_arr;
TOV_in.P_arr = P_arr;
TOV_in.M_arr = M_arr;
TOV_in.expnu_arr = expnu_arr;
TOV_in.exp4phi_arr = exp4phi_arr;
TOV_in.rbar_arr = rbar_arr;
/* END TOV INPUT ROUTINE */
// Step 1.b: Interpolate data from data file to set BSSN gridfunctions
ID_BSSN__ALL_BUT_LAMBDAs(params,xx,TOV_in, in_gfs);
apply_bcs_curvilinear(params, bcstruct, NUM_EVOL_GFS, evol_gf_parity, in_gfs);
enforce_detgammabar_constraint(rfmstruct, params, in_gfs);
ID_BSSN_lambdas(params, xx, in_gfs);
apply_bcs_curvilinear(params, bcstruct, NUM_EVOL_GFS, evol_gf_parity, in_gfs);
enforce_detgammabar_constraint(rfmstruct, params, in_gfs);
ID_TOV_TUPMUNU_xx0xx1xx2(params,xx,TOV_in,auxevol_gfs);
free(rbar_arr);
free(rho_arr);
free(rho_baryon_arr);
free(P_arr);
free(M_arr);
free(expnu_arr);
}
// Step P11: Declare function for evaluating Hamiltonian constraint (diagnostic)
#include "Hamiltonian_constraint.h"
// Step P12: Declare rhs_eval function, which evaluates BSSN RHSs
#include "rhs_eval.h"
// Step P13: Declare Ricci_eval function, which evaluates Ricci tensor
#include "Ricci_eval.h"
// main() function:
// Step 0: Read command-line input, set up grid structure, allocate memory for gridfunctions, set up coordinates
// Step 1: Set up initial data to an exact solution
// Step 2: Start the timer, for keeping track of how fast the simulation is progressing.
// Step 3: Integrate the initial data forward in time using the chosen RK-like Method of
// Lines timestepping algorithm, and output periodic simulation diagnostics
// Step 3.a: Output 2D data file periodically, for visualization
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
// Step 3.c: If t=t_final, output conformal factor & Hamiltonian
// constraint violation to 2D data file
// Step 3.d: Progress indicator printing to stderr
// Step 4: Free all allocated memory
int main(int argc, const char *argv[]) {
paramstruct params;
#include "set_Cparameters_default.h"
// Step 0a: Read command-line input, error out if nonconformant
if((argc != 4 && argc != 5) || atoi(argv[1]) < NGHOSTS || atoi(argv[2]) < NGHOSTS || atoi(argv[3]) < 2 /* FIXME; allow for axisymmetric sims */) {
fprintf(stderr,"Error: Expected three command-line arguments: ./BrillLindquist_Playground Nx0 Nx1 Nx2,\n");
fprintf(stderr,"where Nx[0,1,2] is the number of grid points in the 0, 1, and 2 directions.\n");
fprintf(stderr,"Nx[] MUST BE larger than NGHOSTS (= %d)\n",NGHOSTS);
exit(1);
}
if(argc == 5) {
CFL_FACTOR = strtod(argv[4],NULL);
if(CFL_FACTOR > 0.5 && atoi(argv[3])!=2) {
fprintf(stderr,"WARNING: CFL_FACTOR was set to %e, which is > 0.5.\n",CFL_FACTOR);
fprintf(stderr," This will generally only be stable if the simulation is purely axisymmetric\n");
fprintf(stderr," However, Nx2 was set to %d>2, which implies a non-axisymmetric simulation\n",atoi(argv[3]));
}
}
// Step 0b: Set up numerical grid structure, first in space...
const int Nxx[3] = { atoi(argv[1]), atoi(argv[2]), atoi(argv[3]) };
if(Nxx[0]%2 != 0 || Nxx[1]%2 != 0 || Nxx[2]%2 != 0) {
fprintf(stderr,"Error: Cannot guarantee a proper cell-centered grid if number of grid cells not set to even number.\n");
fprintf(stderr," For example, in case of angular directions, proper symmetry zones will not exist.\n");
exit(1);
}
// Step 0c: Set free parameters, overwriting Cparameters defaults
// by hand or with command-line input, as desired.
#include "free_parameters.h"
// Step 0d: Uniform coordinate grids are stored to *xx[3]
REAL *xx[3];
// Step 0d.i: Set bcstruct
bc_struct bcstruct;
{
int EigenCoord = 1;
// Step 0d.ii: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen Eigen-CoordSystem.
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0d.iii: Set Nxx_plus_2NGHOSTS_tot
#include "set_Cparameters-nopointer.h"
const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
// Step 0e: Find ghostzone mappings; set up bcstruct
#include "boundary_conditions/driver_bcstruct.h"
// Step 0e.i: Free allocated space for xx[][] array
for(int i=0;i<3;i++) free(xx[i]);
}
// Step 0f: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen (non-Eigen) CoordSystem.
int EigenCoord = 0;
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0g: Set all C parameters "blah" for params.blah, including
// Nxx_plus_2NGHOSTS0 = params.Nxx_plus_2NGHOSTS0, etc.
#include "set_Cparameters-nopointer.h"
const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
// Step 0h: Time coordinate parameters
const REAL t_final = 1.8*TOV_Mass; /* Final time is set so that at t=t_final,
* data at the origin have not been corrupted
* by the approximate outer boundary condition */
// Step 0i: Set timestep based on smallest proper distance between gridpoints and CFL factor
REAL dt = find_timestep(¶ms, xx);
//fprintf(stderr,"# Timestep set to = %e\n",(double)dt);
int N_final = (int)(t_final / dt + 0.5); // The number of points in time.
// Add 0.5 to account for C rounding down
// typecasts to integers.
int output_every_N = (int)((REAL)N_final/800.0);
if(output_every_N == 0) output_every_N = 1;
// Step 0j: Error out if the number of auxiliary gridfunctions outnumber evolved gridfunctions.
// This is a limitation of the RK method. You are always welcome to declare & allocate
// additional gridfunctions by hand.
if(NUM_AUX_GFS > NUM_EVOL_GFS) {
fprintf(stderr,"Error: NUM_AUX_GFS > NUM_EVOL_GFS. Either reduce the number of auxiliary gridfunctions,\n");
fprintf(stderr," or allocate (malloc) by hand storage for *diagnostic_output_gfs. \n");
exit(1);
}
// Step 0k: Allocate memory for gridfunctions
#include "MoLtimestepping/RK_Allocate_Memory.h"
REAL *restrict auxevol_gfs = (REAL *)malloc(sizeof(REAL) * NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS_tot);
// Step 0l: Set up precomputed reference metric arrays
// Step 0l.i: Allocate space for precomputed reference metric arrays.
#include "rfm_files/rfm_struct__malloc.h"
// Step 0l.ii: Define precomputed reference metric arrays.
{
#include "set_Cparameters-nopointer.h"
#include "rfm_files/rfm_struct__define.h"
}
// Step 1: Set up initial data to an exact solution
initial_data(¶ms,&bcstruct, &rfmstruct, xx, auxevol_gfs, y_n_gfs);
// Step 1b: Apply boundary conditions, as initial data
// are sometimes ill-defined in ghost zones.
// E.g., spherical initial data might not be
// properly defined at points where r=-1.
apply_bcs_curvilinear(¶ms, &bcstruct, NUM_EVOL_GFS,evol_gf_parity, y_n_gfs);
enforce_detgammabar_constraint(&rfmstruct, ¶ms, y_n_gfs);
// Step 2: Start the timer, for keeping track of how fast the simulation is progressing.
#ifdef __linux__ // Use high-precision timer in Linux.
struct timespec start, end;
clock_gettime(CLOCK_REALTIME, &start);
#else // Resort to low-resolution, standards-compliant timer in non-Linux OSs
// http://www.cplusplus.com/reference/ctime/time/
time_t start_timer,end_timer;
time(&start_timer); // Resolution of one second...
#endif
// Step 3: Integrate the initial data forward in time using the chosen RK-like Method of
// Lines timestepping algorithm, and output periodic simulation diagnostics
for(int n=0;n<=N_final;n++) { // Main loop to progress forward in time.
// Step 3.a: Output 2D data file periodically, for visualization
if(n%100 == 0) {
// Evaluate Hamiltonian constraint violation
Hamiltonian_constraint(&rfmstruct, ¶ms, y_n_gfs,auxevol_gfs, diagnostic_output_gfs);
char filename[100];
sprintf(filename,"out%d-%08d.txt",Nxx[0],n);
FILE *out2D = fopen(filename, "w");
LOOP_REGION(NGHOSTS,Nxx_plus_2NGHOSTS0-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS1-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS2-NGHOSTS) {
const int idx = IDX3S(i0,i1,i2);
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
REAL xCart[3];
xxCart(¶ms,xx,i0,i1,i2,xCart);
fprintf(out2D,"%e %e %e %e\n",
xCart[1]/TOV_Mass,xCart[2]/TOV_Mass,
y_n_gfs[IDX4ptS(CFGF,idx)],log10(fabs(diagnostic_output_gfs[IDX4ptS(HGF,idx)])));
}
fclose(out2D);
}
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
#include "MoLtimestepping/RK_MoL.h"
// Step 3.c: If t=t_final, output conformal factor & Hamiltonian
// constraint violation to 2D data file
if(n==N_final-1) {
// Evaluate Hamiltonian constraint violation
Hamiltonian_constraint(&rfmstruct, ¶ms, y_n_gfs,auxevol_gfs, diagnostic_output_gfs);
char filename[100];
sprintf(filename,"out%d.txt",Nxx[0]);
FILE *out2D = fopen(filename, "w");
const int i0MIN=NGHOSTS; // In spherical, r=Delta r/2.
const int i1mid=Nxx_plus_2NGHOSTS1/2;
const int i2mid=Nxx_plus_2NGHOSTS2/2;
LOOP_REGION(NGHOSTS,Nxx_plus_2NGHOSTS0-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS1-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS2-NGHOSTS) {
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
REAL xCart[3];
xxCart(¶ms,xx,i0,i1,i2,xCart);
int idx = IDX3S(i0,i1,i2);
fprintf(out2D,"%e %e %e %e\n",xCart[1]/TOV_Mass,xCart[2]/TOV_Mass, y_n_gfs[IDX4ptS(CFGF,idx)],
log10(fabs(diagnostic_output_gfs[IDX4ptS(HGF,idx)])));
}
fclose(out2D);
}
// Step 3.d: Progress indicator printing to stderr
// Step 3.d.i: Measure average time per iteration
#ifdef __linux__ // Use high-precision timer in Linux.
clock_gettime(CLOCK_REALTIME, &end);
const long long unsigned int time_in_ns = 1000000000L * (end.tv_sec - start.tv_sec) + end.tv_nsec - start.tv_nsec;
#else // Resort to low-resolution, standards-compliant timer in non-Linux OSs
time(&end_timer); // Resolution of one second...
REAL time_in_ns = difftime(end_timer,start_timer)*1.0e9+0.5; // Round up to avoid divide-by-zero.
#endif
const REAL s_per_iteration_avg = ((REAL)time_in_ns / (REAL)n) / 1.0e9;
const int iterations_remaining = N_final - n;
const REAL time_remaining_in_mins = s_per_iteration_avg * (REAL)iterations_remaining / 60.0;
const REAL num_RHS_pt_evals = (REAL)(Nxx[0]*Nxx[1]*Nxx[2]) * 4.0 * (REAL)n; // 4 RHS evals per gridpoint for RK4
const REAL RHS_pt_evals_per_sec = num_RHS_pt_evals / ((REAL)time_in_ns / 1.0e9);
// Step 3.d.ii: Output simulation progress to stderr
if(n % 10 == 0) {
fprintf(stderr,"%c[2K", 27); // Clear the line
fprintf(stderr,"It: %d t/M=%.2f dt/M=%.2e | %.1f%%; ETA %.0f s | t/M/h %.2f | gp/s %.2e\r", // \r is carriage return, move cursor to the beginning of the line
n, n * (double)dt/TOV_Mass, (double)dt/TOV_Mass, (double)(100.0 * (REAL)n / (REAL)N_final),
(double)time_remaining_in_mins*60, (double)(dt/TOV_Mass * 3600.0 / s_per_iteration_avg), (double)RHS_pt_evals_per_sec);
fflush(stderr); // Flush the stderr buffer
} // End progress indicator if(n % 10 == 0)
} // End main loop to progress forward in time.
fprintf(stderr,"\n"); // Clear the final line of output from progress indicator.
// Step 4: Free all allocated memory
#include "rfm_files/rfm_struct__freemem.h"
#include "boundary_conditions/bcstruct_freemem.h"
#include "MoLtimestepping/RK_Free_Memory.h"
free(auxevol_gfs);
for(int i=0;i<3;i++) free(xx[i]);
return 0;
}
# +
import cmdline_helper as cmd
print("Now compiling, should take ~20 seconds...\n")
start = time.time()
cmd.C_compile(os.path.join(Ccodesdir,"Hydro_without_Hydro_Playground.c"), "Hydro_without_Hydro_Playground")
end = time.time()
print("Finished in "+str(end-start)+" seconds.\n")
cmd.delete_existing_files("out96*.txt")
cmd.delete_existing_files("out96-00*.txt.png")
print("Now running, should take ~10 seconds...\n")
start = time.time()
cmd.Execute("Hydro_without_Hydro_Playground", "96 16 2 "+str(CFL_FACTOR),"out96.txt")
end = time.time()
print("Finished in "+str(end-start)+" seconds.\n")
# -
# <a id='visualize'></a>
#
# # Step 7: Data Visualization Animations \[Back to [top](#toc)\]
# $$\label{visualize}$$
# <a id='installdownload'></a>
#
# ## Step 7.a: Install `scipy` and download `ffmpeg` if they are not yet installed/downloaded \[Back to [top](#toc)\]
# $$\label{installdownload}$$
#
# Note that if you are not running this within `mybinder`, but on a Windows system, `ffmpeg` must be installed using a separate package (on [this site](http://ffmpeg.org/)), or (if running Jupyter within Anaconda, use the command: `conda install -c conda-forge ffmpeg`).
# +
print("Ignore any warnings or errors from the following command:")
# !pip install scipy > /dev/null
# check_for_ffmpeg = !which ffmpeg >/dev/null && echo $?
if check_for_ffmpeg != ['0']:
print("Couldn't find ffmpeg, so I'll download it.")
# Courtesy https://johnvansickle.com/ffmpeg/
# !wget http://astro.phys.wvu.edu/zetienne/ffmpeg-static-amd64-johnvansickle.tar.xz
# !tar Jxf ffmpeg-static-amd64-johnvansickle.tar.xz
print("Copying ffmpeg to ~/.local/bin/. Assumes ~/.local/bin is in the PATH.")
# !mkdir ~/.local/bin/
# !cp ffmpeg-static-amd64-johnvansickle/ffmpeg ~/.local/bin/
print("If this doesn't work, then install ffmpeg yourself. It should work fine on mybinder.")
# -
# <a id='genimages'></a>
#
# ## Step 7.b: Generate images for visualization animation \[Back to [top](#toc)\]
# $$\label{genimages}$$
#
# Here we loop through the data files output by the executable compiled and run in [the previous step](#mainc), generating a [png](https://en.wikipedia.org/wiki/Portable_Network_Graphics) image for each data file.
#
# **Special thanks to <NAME>. His work with the first versions of these scripts greatly contributed to the scripts as they exist below.**
# +
## VISUALIZATION ANIMATION, PART 1: Generate PNGs, one per frame of movie ##
import numpy as np
from scipy.interpolate import griddata
import matplotlib.pyplot as plt
from matplotlib.pyplot import savefig
from IPython.display import HTML
import matplotlib.image as mgimg
import glob
import sys
from matplotlib import animation
globby = glob.glob('out96-00*.txt')
file_list = []
for x in sorted(globby):
file_list.append(x)
bound=7.5
pl_xmin = -bound
pl_xmax = +bound
pl_ymin = -bound
pl_ymax = +bound
N_interp_pts = 300
N_interp_ptsj = 300j
for filename in file_list:
fig = plt.figure()
x,y,other,Ham = np.loadtxt(filename).T #Transposed for easier unpacking
plotquantity = Ham
plotdescription = "Numerical Soln."
plt.title("Single Neutron Star (Ham. constraint)")
plt.xlabel("y/M")
plt.ylabel("z/M")
grid_x, grid_y = np.mgrid[pl_xmin:pl_xmax:N_interp_ptsj, pl_ymin:pl_ymax:N_interp_ptsj]
points = np.zeros((len(x), 2))
for i in range(len(x)):
# Zach says: No idea why x and y get flipped...
points[i][0] = y[i]
points[i][1] = x[i]
grid = griddata(points, plotquantity, (grid_x, grid_y), method='nearest')
gridcub = griddata(points, plotquantity, (grid_x, grid_y), method='cubic')
im = plt.imshow(grid, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
#plt.pcolormesh(grid_y,grid_x, grid, vmin=-8, vmax=0) # Set colorbar range from -8 to 0
ax = plt.colorbar()
plt.clim(-9, -2)
ax.set_label(plotdescription)
savefig(filename+".png",dpi=150)
plt.close(fig)
sys.stdout.write("%c[2K" % 27)
sys.stdout.write("Processing file "+filename+"\r")
sys.stdout.flush()
# -
# <a id='genvideo'></a>
#
# ## Step 7.c: Generate visualization animation \[Back to [top](#toc)\]
# $$\label{genvideo}$$
#
# In the following step, [ffmpeg](http://ffmpeg.org) is used to generate an [mp4](https://en.wikipedia.org/wiki/MPEG-4) video file, which can be played directly from this Jupyter notebook.
# +
## VISUALIZATION ANIMATION, PART 2: Combine PNGs to generate movie ##
# https://stackoverflow.com/questions/14908576/how-to-remove-frame-from-matplotlib-pyplot-figure-vs-matplotlib-figure-frame
# https://stackoverflow.com/questions/23176161/animating-pngs-in-matplotlib-using-artistanimation
fig = plt.figure(frameon=False)
ax = fig.add_axes([0, 0, 1, 1])
ax.axis('off')
myimages = []
for i in range(len(file_list)):
img = mgimg.imread(file_list[i]+".png")
imgplot = plt.imshow(img)
myimages.append([imgplot])
ani = animation.ArtistAnimation(fig, myimages, interval=100, repeat_delay=1000)
plt.close()
ani.save('SingleNS.mp4', fps=5,dpi=150)
# +
## VISUALIZATION ANIMATION, PART 3: Display movie as embedded HTML5 (see next cell) ##
# https://stackoverflow.com/questions/18019477/how-can-i-play-a-local-video-in-my-ipython-notebook
# -
# %%HTML
<video width="480" height="360" controls>
<source src="SingleNS.mp4" type="video/mp4">
</video>
# <a id='convergence'></a>
#
# # Step 8: Validation: Convergence of numerical errors (Hamiltonian constraint violation) to zero \[Back to [top](#toc)\]
# $$\label{convergence}$$
#
# The equations behind these initial data solve Einstein's equations exactly, at a single instant in time. One reflection of this solution is that the Hamiltonian constraint violation should be exactly zero in the initial data.
#
# However, when evaluated on numerical grids, the Hamiltonian constraint violation will *not* generally evaluate to zero due to the associated numerical derivatives not being exact. However, these numerical derivatives (finite difference derivatives in this case) should *converge* to the exact derivatives as the density of numerical sampling points approaches infinity.
#
# In this case, all of our finite difference derivatives agree with the exact solution, with an error term that drops with the uniform gridspacing to the fourth power: $\left(\Delta x^i\right)^4$.
#
# Here, as in the [Start-to-Finish Scalar Wave (Cartesian grids) NRPy+ tutorial](Tutorial-Start_to_Finish-ScalarWave.ipynb) and the [Start-to-Finish Scalar Wave (curvilinear grids) NRPy+ tutorial](Tutorial-Start_to_Finish-ScalarWaveCurvilinear.ipynb) we confirm this convergence.
#
# First, let's take a look at the numerical error on the x-y plane at a given numerical resolution, plotting $\log_{10}|H|$, where $H$ is the Hamiltonian constraint violation:
# +
grid96 = griddata(points, plotquantity, (grid_x, grid_y), method='nearest')
grid96cub = griddata(points, plotquantity, (grid_x, grid_y), method='cubic')
# fig, ax = plt.subplots()
plt.clf()
plt.title("96^3 Numerical Err.: log_{10}|Ham|")
plt.xlabel("x/M")
plt.ylabel("y/M")
fig96cub = plt.imshow(grid96cub.T, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
cb = plt.colorbar(fig96cub)
# -
# Next, we set up the same initial data but on a lower-resolution, $48\times 8\times 2$ grid (axisymmetric in the $\phi$ direction). Since the constraint violation (numerical error associated with the fourth-order-accurate, finite-difference derivatives) should converge to zero with the uniform gridspacing to the fourth power: $\left(\Delta x^i\right)^4$, we expect the constraint violation will increase (relative to the $96\times 16\times 2$ grid) by a factor of $\left(96/48\right)^4$. Here we demonstrate that indeed this order of convergence is observed as expected, *except* in the region causally influenced by the star's surface at $\bar{r}=\bar{R}\approx 0.8$ where the stress-energy tensor $T^{\mu\nu}$ sharply drops to zero.
# +
cmd.delete_existing_files("out48*.txt")
cmd.delete_existing_files("out48-00*.txt.png")
print("Now running, should take ~10 seconds...\n")
start = time.time()
cmd.Execute("Hydro_without_Hydro_Playground", "48 8 2 "+str(CFL_FACTOR), "out48.txt")
end = time.time()
print("Finished in "+str(end-start)+" seconds.")
# +
x48,y48,valuesother48,valuesHam48 = np.loadtxt('out48.txt').T #Transposed for easier unpacking
points48 = np.zeros((len(x48), 2))
for i in range(len(x48)):
points48[i][0] = x48[i]
points48[i][1] = y48[i]
grid48 = griddata(points48, valuesHam48, (grid_x, grid_y), method='cubic')
griddiff_48_minus_96 = np.zeros((N_interp_pts,N_interp_pts))
griddiff_48_minus_96_1darray = np.zeros(N_interp_pts*N_interp_pts)
gridx_1darray_yeq0 = np.zeros(N_interp_pts)
grid48_1darray_yeq0 = np.zeros(N_interp_pts)
grid96_1darray_yeq0 = np.zeros(N_interp_pts)
count = 0
outarray = []
for i in range(N_interp_pts):
for j in range(N_interp_pts):
griddiff_48_minus_96[i][j] = grid48[i][j] - grid96[i][j]
griddiff_48_minus_96_1darray[count] = griddiff_48_minus_96[i][j]
if j==N_interp_pts/2-1:
gridx_1darray_yeq0[i] = grid_x[i][j]
grid48_1darray_yeq0[i] = grid48[i][j] + np.log10((48./96.)**4)
grid96_1darray_yeq0[i] = grid96[i][j]
count = count + 1
plt.clf()
fig, ax = plt.subplots()
plt.title("Plot Demonstrating 4th-order Convergence")
plt.xlabel("x/M")
plt.ylabel("log10(Relative error)")
ax.plot(gridx_1darray_yeq0, grid96_1darray_yeq0, 'k-', label='Nr=96')
ax.plot(gridx_1darray_yeq0, grid48_1darray_yeq0, 'k--', label='Nr=48, mult by (48/96)^4')
ax.set_ylim([-9.5,-1.5])
legend = ax.legend(loc='lower right', shadow=True, fontsize='x-large')
legend.get_frame().set_facecolor('C1')
plt.show()
# -
# <a id='latex_pdf_output'></a>
#
# # Step 9: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.pdf](Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.ipynb
# !pdflatex -interaction=batchmode Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.tex
# !pdflatex -interaction=batchmode Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.tex
# !pdflatex -interaction=batchmode Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
| Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ''
# name: ''
# ---
# *Code for figure generation not yet available. Static version of the figure below.*
#
# 
| 2-Additional_Figures/Fig04_Detector-Resolution-Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: optics
# language: python
# name: optics
# ---
# +
# -*- coding: utf-8 -*-
import sys
from felpy.model.wavefront import Wavefront
if __name__ == '__main__':
print("working")
in_directory = "/gpfs/exfel/data/user/guestt/dCache/NanoKB-Pulse/EHC/2.h5" #sys.argv[1]
out_directory = "/out" #sys.argv[2]
wfr = Wavefront()
print("wfr loaded")
wfr.load_hdf5(in_directory)
wfr.analysis(VERBOSE = True, DEBUG = True)
#print(wfr.custom_fields) ## good debug
# -
| desy/ipynb_simulated_wavefront_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
from __future__ import print_function
import sys
sys.path.append('../build/')
# %pylab inline
np.set_printoptions(precision=4, suppress=True)
import versor as vsr
from versor.drawing import *
# +
l1 = vsr.Dll(vsr.Vec(0,0,0).null(), vsr.Vec(1,0,0).null()).unit()
theta = pi/10
delta = 0.4
m = vsr.Vec(1,1,delta).trs() * vsr.Biv(theta/2,0,0).exp()
# l1 = l1.spin(m)
l2 = l1.spin(m)
m2 = l2 * l1.rev()
t = m2.trs() * 0.5
w = t.reject_from(m2.dll().biv().unit())
l3 = l2.comm(l1.rev())
l1_dir = l1.dir().vec().unit()
l1_loc = l1.loc(vsr.Vec(0,0,0).null()).vec()
l2_dir = l2.dir().vec().unit()
l2_loc = l2.loc(vsr.Vec(0,0,0).null()).vec()
l4 = vsr.Dll(l3.loc(vsr.Vec(0,0,0).null()), l3.biv().duale().unit())
p1 = l1.loc(l4.loc(vsr.Vec(0,0,0).null()))
p2 = l2.loc(l4.loc(vsr.Vec(0,0,0).null()))
p = ((p2.vec() + p1.vec()) * 0.5).null()
# bisector
lv = vsr.Dll(p, (l2_dir + l1_dir).unit())
lw = vsr.Dll(p, (l2_dir-l1_dir).unit())
l5 = l2 - l1
st = sin(theta/2)
ct = cos(theta/2)
deltahalf = delta / 2
dw = lw.drv().vec()
dw2 = np.inner(dw,dw)
print(dw2)
cost = 4 * st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + delta**2 * ct**2
print(cost)
print(4 * st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + delta**2 * (1 - st**2))
print(theta**2 + dw2 * theta**2 + delta**2)
print(theta**2 + theta**2 + )
# -
(1.0 + dw2) * theta**2 + delta**2
(1.0) * theta**2 + delta**2
# +
l1 = vsr.Dll(vsr.Vec(0,0,0).null(), vsr.Vec(1,0,0).null()).unit()
theta = pi/6
delta = 0.6
m = vsr.Vec(1,1,delta).trs() * vsr.Biv(theta/2,0,0).exp()
# l1 = l1.spin(m)
l2 = l1.spin(m)
m2 = l2 * l1.rev()
t = m2.trs() * 0.5
w = t.reject_from(m2.dll().biv().unit())
l3 = l2.comm(l1.rev())
l1_dir = l1.dir().vec().unit()
l1_loc = l1.loc(vsr.Vec(0,0,0).null()).vec()
l2_dir = l2.dir().vec().unit()
l2_loc = l2.loc(vsr.Vec(0,0,0).null()).vec()
l4 = vsr.Dll(l3.loc(vsr.Vec(0,0,0).null()), l3.biv().duale().unit())
p1 = l1.loc(l4.loc(vsr.Vec(0,0,0).null()))
p2 = l2.loc(l4.loc(vsr.Vec(0,0,0).null()))
p = ((p2.vec() + p1.vec()) * 0.5).null()
# bisector
lv = vsr.Dll(p, (l2_dir + l1_dir).unit())
lw = vsr.Dll(p, (l2_dir-l1_dir).unit())
l5 = l2 - l1
st = sin(theta/2)
ct = cos(theta/2)
deltahalf = delta / 2
dw = lw.drv().vec()
dw2 = np.inner(dw,dw)
cost = 4 * st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + delta**2 * ct**2
print(cost)
scene = Scene(children=[
frame_mesh(),
line_mesh(l1, color=Colors.DEEPSKYBLUE),
point_mesh(p1,color=Colors.DEEPSKYBLUE),
line_mesh(l2, color=Colors.DEEPPINK),
point_mesh(p2,color=Colors.DEEPPINK),
vector_mesh(t, color='red'),
vector_mesh(w, position=p1, color='green'),
# line_mesh(l3, color='lightgreen'),
# line_mesh(l5, color='lightgreen'),
# midpoint
# point_mesh(p, color='lightgreen'),
# bivector_mesh(l3.biv(), color='red'),
# vector_mesh(B1.comm(bn.comm(B2) * delta), color='red'),
# line_mesh(lv, color='lightgreen'),
AmbientLight(color='#777777')])
camera = PerspectiveCamera(position=[0, 5, 5], up=[0, 0, 1],
children=[DirectionalLight(color='white',
position=[3, 5, 1],
intensity=0.5)])
# camera.children = []
renderer = Renderer(camera=camera,
scene=scene,
controls=[TrackballControls(controlling=camera)],
background = 'white')
# renderer.width = '950'
# renderer.height = '713'
# display(renderer)
# +
p1 = vsr.Vec(1,0,1).null()
p2 = vsr.Vec(2,0.2,1).null()
l1 = vsr.Dll(vsr.Vec(0,0,0).null(), vsr.Vec(1,0,0).null()).unit()
theta = pi/6
delta = 0.6
m = vsr.Vec(1,1,delta/2).trs() * vsr.Biv(theta/2,0,0).exp()
l1 = l1.spin(m)
l2 = l1.spin(m)
l3 = l2.comm(l1.rev())
m2 = l2 /
l1_dir = l1.dir().vec().unit()
l1_loc = l1.loc(vsr.Vec(0,0,0).null()).vec()
l2_dir = l2.dir().vec().unit()
l2_loc = l2.loc(vsr.Vec(0,0,0).null()).vec()
l4 = vsr.Dll(l3.loc(vsr.Vec(0,0,0).null()), l3.biv().duale().unit())
p1 = l1.loc(l4.loc(vsr.Vec(0,0,0).null()))
p2 = l2.loc(l4.loc(vsr.Vec(0,0,0).null()))
p = ((p2.vec() + p1.vec()) * 0.5).null()
# bisector
lv = vsr.Dll(p, (l2_dir + l1_dir).unit())
lw = vsr.Dll(p, (l2_dir-l1_dir).unit())
l5 = l2 - l1
scene = Scene(children=[
frame_mesh(),
line_mesh(l1, color=Colors.DEEPSKYBLUE),
point_mesh(p1,color=Colors.DEEPSKYBLUE),
line_mesh(l2, color=Colors.DEEPPINK),
point_mesh(p2,color=Colors.DEEPPINK),
# line_mesh(l3, color='lightgreen'),
# line_mesh(l5, color='lightgreen'),
# midpoint
# point_mesh(p, color='lightgreen'),
# bivector_mesh(l3.biv(), color='red'),
# vector_mesh(B1.comm(bn.comm(B2) * delta), color='red'),
# line_mesh(lv, color='lightgreen'),
AmbientLight(color='#777777')])
camera = PerspectiveCamera(position=[0, 5, 5], up=[0, 0, 1],
children=[DirectionalLight(color='white',
position=[3, 5, 1],
intensity=0.5)])
# camera.children = []
renderer = Renderer(camera=camera,
scene=scene,
controls=[TrackballControls(controlling=camera)],
background = 'white')
# renderer.width = '950'
# renderer.height = '713'
# display(renderer)
# -
m = l2 * l1.rev()
t = m.trs()
v = t.reject_from(m.dll().biv().unit())
v
m.dll().drv().vec().reject_from(m.dll().biv().unit()) * 2
S = m.log()
A = S.biv()
b = S.drv().vec()
print(A.norm() - theta)
b.reject_from(A.unit())
a = vsr.Dll(0,0,1,0,0,0)
b = a.spin(vsr.Vec(0,0,1).trs() * vsr.Biv(0.01,0,0).exp())
m = a * b.rev()
m.trs().reject_from(m.dll().biv().unit())
print(a)
print(b)
delta * sin(theta)
-tan(theta)
# +
st = sin(theta/2)
ct = cos(theta/2)
deltahalf = delta / 2
dw = lw.drv().vec()
dw2 = np.inner(dw,dw)
4 * st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + delta**2 * ct**2
# -
np.inner(np.array((l2 * l1.rev())) - np.array([1,0,0,0,0,0,0,0]),
np.array((l2 * l1.rev())) - np.array([1,0,0,0,0,0,0,0]))
l2 * l1.rev() * (l2 * l1.rev()).rev()
l = l2 * l1.inv()
l
m = l2.comm(l1.rev()).drv().vec().norm()
np.inner([l[0],l[7]],[l[0],l[7]])
[l[0],l[7]]
l[0]*l[0]
cos(theta)**2 + delta**2 * sin(theta)**2
cos(theta)**2 + delta**2 * sin(theta)**2 + sin(theta)**2
np.inner(l,l)
np.inner([l[4],l[5],l[6]],[l[4],l[5],l[6]])
sin(theta)**2 * m + delta**2 * cos(theta)**2
B3 = np.array(l2.comm(l1.rev()).biv().unit())
D3 = np.array(l2.comm(l1.rev()).drv().vec().unit())
np.inner(B3,D3)
np.inner([l[0],l[1],l[2],l[3],l[7]],[l[0],l[1],l[2],l[3],l[7]])
dl = lambda th, d : 4 * sin(th/2)**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + d**2 * cos(th/2)**2
dl2 = lambda th, d : 4 * sin(th/2)**2 + d**2 * cos(th/2)**2
dl3 = lambda th, d: 4 * sin(th/2)**2 * (1.0 + np.inner(np.array(dw),np.array(dw)))
ds = dl(np.linspace(0,pi), 0.0)
np.linalg.norm(np.outer(l1,l2) - np.outer(l2,l1))
np.linalg.norm(np.outer(l1,l2) - np.outer(l2,l1))
np.inner(l1.comm(l2), l1.comm(l2)) * 2 * sqrt(2)
A = np.outer(l1,l2) - np.outer(l2,l1)
np.diag(np.dot(A.T,A))
l1.comm(l2)
sqrt(np.sum(np.diag(np.dot(A.T,A))))
np.outer([1,2,3],[4,5,6])
l1a = np.array(l1)
l2a = np.array(l2)
A = np.outer(l1a[:3],l2a[:3]) - np.outer(l2a[:3],l1a[:3])
(np.linalg.norm(A) / sqrt(2))**2
A = np.outer(l1a,l2a) - np.outer(l2a,l1a)
(np.linalg.norm(A) / sqrt(2))**2
np.linalg.svd(np.eye(150, 36))
np.linalg.norm(l2 * l1 - l1 * l2)
np.array
np.dot(A.T,A)
2 * sin(theta)**2
np.outer(l1a[:3],l2a[3:]) - np.outer(l2a[:3],l1a[3:])
np.outer(l1,l2) - np.outer(l2,l1)
[[( l1[k] * l2[l] - l1[l] * l2[k])**2 for k in range(6) if k != l] for l in range(6)]
d = 0
for k in range(6):
for l in range(6):
if k != l:
d += (l1[k] * l2[l] - l1[l] * l2[k])**2
print(sqrt(d))
lc = l1.comm(l2) * 2.0
np.linalg.norm(np.outer(lc,lc))
np.linspace(0,pi)
plot(dl(np.linspace(0,2 * pi,1000), 1.0))
plot(dl(np.linspace(0,2 * pi,1000), 2.0))
plot(dl(np.linspace(0,2 * pi,1000), 4.0))
plot(dl2(np.linspace(0,2 * pi,1000), 1.0))
plot(dl2(np.linspace(0,2 * pi,1000), 2.0))
plot(dl2(np.linspace(0,2 * pi,1000), 4.0))
x = arange(0,2*pi,0.01)
y = arange(0,2*pi,0.01)
dl(0,0)
X,Y = meshgrid(x,y)
Z = sqrt(dl(X,Y))
im = imshow(Z,cmap=cm.RdBu)
# adding the Contour lines with labels
cset = contour(Z,arange(0, 2 * pi,1),linewidths=2,cmap=cm.Set2)
clabel(cset,inline=True,fmt='%1.1f',fontsize=10)
colorbar(im) # adding the colobar on the right
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1,
cmap=cm.RdBu,linewidth=0, antialiased=False)
# +
def z_func(x,y):
return (1-(x**2+y**3))*exp(-(x**2+y**2)/2)
x = arange(-3.0,3.0,0.1)
y = arange(-3.0,3.0,0.1)
X,Y = meshgrid(x, y) # grid of point
Z = z_func(X, Y) # evaluation of the function on the grid
im = imshow(Z,cmap=cm.RdBu) # drawing the function
# -
plot(sqrt(dl(np.linspace(0,pi), 0.0)))
plot(sqrt(dl(np.linspace(0,pi), 1.0)))
plot(sqrt(dl(np.linspace(0,pi), 2.0)))
cos(pi/2)**2
sqrt(4 * st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + delta**2 * ct**2)
2 * sqrt( st**2 * (1.0 + np.inner(np.array(dw),np.array(dw))) + 0.25 * delta**2 * ct**2)
0.25 * delta**2 * ct**2
(0.5 * delta * ct)**2
delta**2 * (1 + cos(theta)) * 0.5
delta**2 * ct**2
0.64 * 0.64
dw[0] * dw[0] + dw[1] * dw[1] + dw[2] * dw[2]
dw[0]**2 + dw[1]**2 + dw[2]**2
np.dot(dw,dw)
np.inner(dw,dw)
2 * st**2
1.0 - cos(theta)
delta**2 * ct**2
delta*2 * (1 + ct) * 0.5
sqrt(np.inner(l5,l5))
np.inner(l5.dir().vec().unduale(),l5.dir().vec().unduale())
l5.loc(vsr.Vec(0,0,0).null())
np.cross(p.vec(),l5.dir().vec()) * 2
4 * st**2
scene.children[1].material.color='pink'
np.inner(l5,l5)
4 * st**2 * (1.0 + np.inner(dw,dw)) + delta**2 * ct**2
eps = vsr.Mot(0,0,0,0,0,0,0,-1)
eps
eps.rev()
th = vsr.Mot(theta,0,0,0,0,0,0,-delta)
th
sth = vsr.Mot(sin(theta),0,0,0,0,0,0, -cos(theta) * delta)
sth
cth
l4 * sth
l3
np.linalg.norm(l3)
n = l3 * l3.conj()
A = vsr.Dll(1,-1,1,2,1,-1)
vsr.CGA(A)
Ac = vsr.CGA(A).conj()
Ac
n[0] - n[7]
l3 * l3.rev()
l = vsr.Dll(vsr.Vec(1,0,2).null(), vsr.Vec(0,1,1).null())
np.inner(l,l)
vsr.CGA(vsr.Pnt(0,0,0,0,1)).rev()
Ei = vsr.Dll(0,0,0,1,0,0)
Ei * Ei.conj()
vsr.CGA(vsr.Dll(1,2,3,4,5,6)) * vsr.CGA(vsr.Vec(1,2,3).null())
vsr.CGA(vsr.Dll(1,2,3,4,5,6)) <= vsr.CGA(vsr.Vec(1,2,3).null())
vsr.CGA(vsr.Vec(1,2,3).null()) <= vsr.CGA(vsr.Dll(1,2,3,4,5,6))
vsr.EGA(vsr.Vec(1,2,3)) <= vsr.EGA(vsr.Biv(1,2,3))
l1 * l2.rev()
l2 * l1.inv()
l2 * l1.rev()
l1 * l2
B1 = l1.biv()
B2 = l2.biv()
D1 = p1.vec() ^ B1.duale()
D1reveps = vsr.Dll(D1.rev()[0], D1.rev()[1], D1.rev()[2],0,0,0) * eps
D2 = p2.vec() ^ B2.duale()
l1.rev()
a = vsr.EGA(vsr.Vec(1,2,3))
I = vsr.EGA(0,0,0,0,0,0,0,1)
a * I
0.5 * (a*I + I*a)
D1eps.rev()
D1reveps
l1
l1.rev()
D1.rev()
print(l1)
print(B1)
print(D1eps)
A = vsr.Mot(0,1,2,3,0,0,0,0)
B = vsr.Mot(0,-4,-5,-6,0,0,0,0)
l2 * (l1 * -1)
(l2 * l1) * -1
l2.comm(l1.rev())
acomm = lambda l1,l2 : (l1 * l2 + l2 * l1) * 0.5
acomm(l2,l1.rev())
acomm(l1.rev(),l2)
A * eps * B
B2 <= B1.rev()
(B1.duale() <= B2)
(B2.duale() <= B1.rev())
-(B2.duale() <= B1)
l3.biv().duale()
vsr.EGA(B1) * vsr.EGA(0,0,0,0,0,0,0,1)
l2 * l1.inv()
b1 = B1.duale()
b2 = B2.duale()
b1.comm(B2)
b2.comm(B1)
Tri = vsr.EGA(0,0,0,0,0,0,0,1)
b1 = vsr.EGA(b1)
b2 = vsr.EGA(b2)
p1 = vsr.EGA(p1.vec())
p1 = vsr.EGA(p2.vec())
b1 * Tri
((b1 * Tri).comm(p1.comm(b2)) - (b2*Tri).comm(p1.comm(b1))).dual()
p1.comm(b1.comm(b2))
p1.comm(Tri)
(Tri * ((b1).comm(p1.comm(b2)) - (b2).comm(p1.comm(b1)))).dual()
l3 * l3.rev()
eps * ((B2 <= (p1 ^ b1).rev()) + ((p2 ^ b2) <= B1.rev()))
eps * (((p2 ^ b2) <= B1.rev()) - ((p1 ^ b1) <= B2))
eps * ((-((p2 ^ b2) <= B1)) - ((p1 ^ b1) <= B2))
eps * ((-(p2 <= (b2 <= B1))) - (p1 <= (b1 <= B2)))
eps * ((-(p2 <= (b2 ^ b1).unduale())) - (p1 <= (b1 ^ b2).unduale()))
eps * (((p2 <= (b1 ^ b2).unduale())) - (p1 <= (b1 ^ b2).unduale()))
eps * (((p2 <= (b1 ^ b2).unduale())) - (p1 <= (b1 ^ b2).unduale()))
B2.comm(B1.rev())
l3
bivtodll = lambda b: vsr.Dll(b[0], b[1], b[2], 0,0,0)
l2.comm(l1.rev())
bivtodll(B2.comm(D1.rev()) + D2.comm(B1.rev())) * eps
vsr.EGA(p1 * b1) - vsr.EGA(b1 * p1)
B2.comm(B1.rev())
B1.comm(B2)
l2.comm(l1.rev())
sth * (l2.comm(l1.rev()).unit())
np.array((p2.vec() - p1.vec()) * sin(theta))
delta * sin(theta) * l3.biv().unit().duale()
A * B * eps
cth = vsr.Mot(cos(theta),0,0,0,0,0,0, sin(theta) * delta)
cth
np.allclose(4 * st**2 * (1.0 + np.inner(dw,dw)) + delta**2 * ct**2, np.inner(l5,l5))
S = l1 - l2
print(S)
Bs = S.biv()
print()
Bsu = Bs.unit()
ds = S.drv().vec()
print(Bsu)
(l1 * l2 + l2 * l1 ) * 0.5
d1 = vsr.EGA(l1.drv().vec())
d2 = vsr.EGA(l2.drv().vec())
B1 = vsr.EGA(l1.biv())
B2 = vsr.EGA(l2.biv())
((B1^d2) + (d1^B2) + (B2^d1) + (d2^B1)) * 0.5
B1^d2
d2^B1
d1^B2
(B2^d1)
cos(theta) , delta * sin(theta)
vsr.EGA(l1.biv()) <= vsr.EGA(l2.biv()) #acomm
(l1 * l2 - l2 * l1 ) * 0.5
l2 * l1
(vsr.EGA(l1.biv() * l2.biv()) + vsr.EGA(l2.biv() * l1.biv())) * 0.5
(vsr.EGA(l1.biv() * l2.biv()) - vsr.EGA(l2.biv() * l1.biv())) * 0.5
l1.biv() <= l2.biv()
l1.biv().comm(l2.biv())
vsr.EGA(l1.biv()) * vsr.EGA(l2.drv().vec()) + vsr.EGA(l1.drv().vec()) * vsr.EGA(l2.biv())
dsp = ds.project_onto(Bsu)
print(dsp)
dso = ds.reject_from(Bs)
print(dso)
print(dsp + dso, ds)
w = Bs.norm()
d = dso.norm()
dso.unit() <= Bsu.duale()
h = d / w
dso * h
S1 = Dll(Bs[0], Bs[1], Bs[1], dsp[0], dsp[1], dsp[2]).unit()
print(S1)
S1.unit()
S2 = Dll(0,0,0, Bsu.duale()[0], Bsu.duale()[1], Bsu.duale()[2]) * h
print(S2)
S11 = S1 + S2
print(S11 * w)
S
S
| python/Line-Difference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Семинар 1. Python, numpy
# ## Starter-pack (не для курса)
#
# 👒 Разберитесь с гитхабом. Клонируйте себе [репо](https://github.com/AsyaKarpova/ml_nes_2021) нашего курса. Необязательные [советы](https://t.me/KarpovCourses/213) по оформлению.
#
# 👒 [Leetcode](https://leetcode.com/problemset/all/https://leetcode.com/problemset/all/): тут можно решать задачки на алгоритмы на питоне (и не только)
#
# 👒 Если вы почему-то решили, что вам надо понимать алгоритмы, но не хочется ботать целый курс, почитайте «Грокаем алгоритмы..»
#
# 👒 Можно скачать Slack и вступить в ods ([тык](https://ods.ai/join-community)).
# ## Вспомнить Python за 10 минут
# ### list comprehensions — [expression for member in iterable]
# - Создайте лист с квадратами чисел от 1 до 10 с помощью list comprehensions
[i**2 for i in range(1, 11)]
# - С помощью list comprehensions создайте новый лист с квадратами только тех чисел от 1 до 10, что без остатка делятся на 3.
[i**2 for i in range(1, 11) if i%3 == 0]
# - С помощью list comprehensions замените все отрицательные значения в листе на 0
candy_prices = [35.4, 26.7, -33.8, 41.9, -100, 25]
[i if i > 0 else 0 for i in candy_prices]
# - walrus operator
#
# Cгенерируйте 20 значений случайной величины и оставьте только те, что больше 28.
import random
def get_number():
return random.randrange(17, 35)
num = 1
print(num)
print(num:=1)
[i for _ in range(20) if (i:= get_number()) > 28]
# ### lambda function
# - отсортируйте айдишники по убыванию
ids = ['id1', 'id2', 'id30', 'id3', 'id100', 'id22']
def sqrt(x):
return x * x
sqrt(2)
(lambda x: x * x)(2)
sorted(ids, key=lambda x: int(x[2:]), reverse=True)
# ### Индексация
# - Вспомните, как вывести последний элемент; все, кроме последнего; через один элемент, начиная с первого; в обратном порядке
elems = [1, 2, 3, 'b||']
elems[-1]
elems[0:-1]
elems[1::2]
elems[::-1]
# ## Задачки
# - Написать функцию, которая выдает вектор с кумулятивными суммами. Функция должна изменять старый вектор, а не возвращать новый.
#
# `Input: [1,2,3,4]`
#
# `Output: [1,3,6,10]`, т. к. `[1, 1+2, 1+2+3, 1+2+3+4]`
#
from typing import List
def runningSum(nums: List[int]) -> List[int]:
for i in range(1, len(nums)):
nums[i] += nums[i-1]
return nums
def runningSum(nums: List[int]) -> List[int]:
return [sum(nums[:i+1]) for i in range(len(nums))] # не in-place
nums = [1, 3, 2]
runningSum(nums)
nums
# - Дан вектор `prices` с ценами акций. Цена в день `[i]` — это `prices[i]`.
# Вы можете один раз купить и один раз продать акцию. Найдите максимальный профит от этой операции. Если профита нет, вверните 0 (ничего не покупайте).
#
#
# `Input: [1,10,8,16]`
#
# `Output: 15`, т. к. `16-1`
#
def maxProfit(prices: List[int]) -> int:
return max([max(prices[i:]) - prices[i] for i in range(len(prices))])
# хардкор
def maxProfit(prices: List[int]) -> int:
if len(prices) == 1:
return 0
else:
days_purchase = [-1] * (len(prices) - 1)
profit = [-1] * (len(prices) - 1)
days_purchase[0] = 0
profit[0] = prices[1] - prices[0]
for i in range(1, len(days_purchase)):
if prices[i] < prices[days_purchase[i - 1]]:
days_purchase[i] = i
profit[i] = prices[i + 1] - prices[i]
else:
days_purchase[i] = days_purchase[i - 1]
profit[i] = prices[i + 1] - prices[days_purchase[i]]
return max(max(profit), 0)
# O(N)
def maxProfit(prices: List[int]) -> int:
current_min = prices[0] # это минимальная цена
current_max = 0 # это максимальный профит
for i in range(len(prices)):
current_min = min(current_min, prices[i])
current_max = max(current_max, prices[i] - current_min)
return current_max
maxProfit([2,1,8,16])
maxProfit([15, 12, 11])
# - Дан массив чисел, в котором каждое число повторяется дважды и только одно значение — один раз.
# Верните это число.
#
# `Input: [1,1,2,3,3]`
#
# `Output: 2`
#
# **hint**: XOR
def singleNumber(nums: List[int]) -> int:
return [nums.count(nums[i]) for i in range(len(nums))]
def singleNumber(nums: List[int]) -> int:
result = 0
for i in nums:
result ^= i
return result
singleNumber([1,1,2,3,3])
# - Определите, является ли последовательность из скобочек валидной.
#
# Возможные символы: `'(', ')', '{', '}', '[', ']'.`
#
#
# `Input: '()[]'`
#
# `Output: True`
#
# ===
#
# `Input: '([{}])'`
#
# `Output: True`
#
# ===
#
# `Input: '([{])'`
#
# `Output: False`
# **hint**: stack
def ValidPar(seq: str) -> bool:
pairs = {'}':'{', ')':'(', ']':'['}
opens = []
for i in seq:
if i in pairs.values():
opens.append(i)
elif len(opens)==0 or pairs[i]!=opens[-1]:
return False
else:
opens.pop(-1)
return True
ValidPar(')')
ValidPar('()')
ValidPar('()[{}]')
# ## numpy
import numpy as np
np.random.seed(10)
# - Вспоминаем, как создавать матрицы из ноликов, едничек, диагональную, матрицу случайных величин :)
np.diag([1, 2, 3])
np.zeros((5,5))
np.ones((5, 5))
np.random.poisson(lam=5, size=(5,3))
a = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
a
# - Записать все значения матрицы в один лист
c = a.ravel()
c
c[0] = 200
print(c)
print('Исходная матрица тоже изменилась')
print(a)
b = a.flatten()
b[0] = 400
b
print(b)
print('Исходная матрица не изменилась')
print(a)
# - Из двумерного массива отберите в один лист элементы с индексами `(0,0), (1,2), (2,2)`
a[[0,1,2],[0,2,2]]
# - Создайте таблицу умножения от 1 до 10
# Перед тем как идти в бой :)
x = np.arange(4)
y = np.arange(5)
x*y
y.shape
y1 = y[:, None]
print(y1.shape)
y1 = y[:, np.newaxis]
print(y1.shape)
y1 * x
# + active=""
# (5, 1) * (1, 4) -> (5, 4)
#
# потому что
# (5, 1)
# (1, 4)
# +
def mult_table(n: int) -> np.ndarray:
return np.arange(1,n+1)[:,np.newaxis]@np.arange(1,n+1)[np.newaxis,:]
# это работает, потому что для векторов c размерностями (n, 1) и (1, n)
# предсказуемым образом выполняется матричное (@) умножение
# то есть (n, 1) @ (1, n) -> (n, n)
# +
def mult_table(n: int) -> np.ndarray:
rng = np.arange(1,n+1)
return rng[:, None] * rng
# тут перемножаются два вектора (n, 1) * (n, )
# крайние размерности не совпадают, но в одном из случаев она равна 1
# тогда python посчитает "в столбик"
# rng[:, None] n x 1
# rng x n
# result n x n
# -
mult_table(10)
# Не стоит воспринимать это как какую-то новую математику. Скорее, это логика, которая реализована в numpy, чтобы обеспечить более эффективые вычисления над векторами и избежать лишних циклов в питоне.
# - Заменить нолики на единички
purr = np.array([1, 2, 6, 0, 0 , 7])
np.where(purr==0, 1, purr)
# - Сделайте one-hot encoding вектора. То есть создайте матрицу, в которой число колонок равно числу уникальных значений вектора, а число строк — длине вектора. Заполните ячейки матрицы единичками, если на данной позиции в векторе встретилась конкретная цифра.
vec2 = np.array([1, 2, 5, 5, 4])
def one_hot(vec: np.ndarray) -> np.ndarray:
a = np.repeat(np.unique(vec)[np.newaxis,:],vec.size,axis=0)
b = np.repeat(vec[:,np.newaxis], np.unique(vec).size, axis=1)
res = np.zeros((vec.size,np.unique(vec).size))
res[a==b]=1
return res
# чуть проще
def one_hot(vec: np.ndarray) -> np.ndarray:
uniq_vals = np.unique(vec2)
return (vec2[:, None] == uniq_vals).astype(int)
one_hot(vec2)
# - Выведите вектор отношения минимума к максимум для каждой строчки матрицы
a = np.random.normal(loc=3, scale=4, size=(4,3))
np.min(a, axis=1)/np.max(a, axis=1)
np.apply_along_axis(lambda x: x.min()/x.max(), 1, a)
# - Уберите все пропущенные значения из vec
vec = np.array([1, 2, 3, np.nan, np.nan, 3])
vec[~np.isnan(vec)]
vec[vec==vec]
# - Поменяйте первую и вторую колонку местами
arr = np.arange(9).reshape(3,3)
arr[:, [0, 2, 1]]
arr[:,[0,1]]=arr[:,[1,0]]
arr
# - Найдите наиболее близкое значение вектора к заданному (по абсолютному значению)
def find_nearest(array: np.array, val: int) -> int:
return array[np.argmin(abs(array - val))]
find_nearest(np.array([-1, 2, 3]), 0)
# +
from PIL import Image
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Про картинки
# 
# В картинке 4 полосочки с пикселями :) Каждая полосочка содержит пять пикселей. Каждый пиксель содержит три бита (red, green, blue).
#
# Такое изображение обычно хранится в виде трехмерного массива размерности $height \times width \times numChannels$, где число каналов часто равно 3 (или 4 в случае RGBA). Элементами такого массива являются 8-битовые натуральные числа (то есть возможные значения от 0 до 2^8-1 :) Чиселки влияют на насыщенность цвета — от черного (0) до максимально насыщенного (255).
# - Нарисовать RGB картинку с помощью numpy :) Например, розовый квадратик
picture = np.zeros((512, 512, 3), dtype=np.uint8)
picture[:, :] = [120, 0, 60]
img = Image.fromarray(picture, 'RGB')
picture[:, :, 0]
# #### Для тех, кто не понял или хочет деталей
#
# - [Броадкаcтинг](https://numpy.org/devdocs/user/basics.broadcasting.html)
# - [Индексация](https://numpy.org/doc/stable/reference/arrays.indexing.html)
| sem1/sem1_py_numpy_done.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="FJogEfYBOfUm"
# # Chapter 8: Winningest Methods in Time Series Forecasting
#
# Compiled by: <NAME>
#
# In previous sections, we examined several models used in time series forecasting such as ARIMA, VAR, and Exponential Smoothing methods. While the main advantage of traditional statistical methods is their ability to perform more sophisticated inference tasks directly (e.g. hypothesis testing on parameters, causality testing), they usually lack predictive power because of their rigid assumptions. That is not to say that they are <i>necessarily</i> inferior when it comes to forecasting, but rather they are typically used as performance benchmarks.
#
# In this section, we demonstrate several of the fundamental ideas and approaches used in the recently concluded [`M5 Competition`](https://mofc.unic.ac.cy/m5-competition/) where challengers from all over the world competed in building time series forecasting models for both [`accuracy`](https://www.kaggle.com/c/m5-forecasting-accuracy) and [`uncertainty`](https://www.kaggle.com/c/m5-forecasting-uncertainty) prediction tasks. Specifically, we explore the machine learning model that majority of the competition's winners utilized: [`LightGBM`](https://lightgbm.readthedocs.io/en/latest/index.html), a tree-based gradient boosting framework designed for speed and efficiency.
# + [markdown] id="olkHffRbOfUr"
# ## 1. M5 Dataset
#
# You can download the M5 dataset from the Kaggle links above.
#
# Let's load the dataset and examine it.
# + id="floWkFlxOfUs"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plot_x_size = 15
plot_y_size = 2
np.set_printoptions(precision = 6, suppress = True)
date_list = [d.strftime('%Y-%m-%d') for d in pd.date_range(start = '2011-01-29', end = '2016-04-24')]
df_calendar = pd.read_csv('../data/m5/calendar.csv')
df_price = pd.read_csv('../data/m5/sell_prices.csv')
df_sales = pd.read_csv('../data/m5/sales_train_validation.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 456} id="mjsCtvwkOfUt" outputId="506c2c09-de24-4f59-956c-55fe6242e08f"
df_sales.rename(columns = dict(zip(df_sales.columns[6:], date_list)), inplace = True)
df_sales
# -
df_calendar
df_price
# + [markdown] id="u95dxURfOfUu"
# #### Sample Product
#
# Let's choose a random product and plot it.
# + colab={"base_uri": "https://localhost:8080/"} id="voNbIA1TOfUu" outputId="3ca5b139-9034-473a-f168-fdcfaa1e504b"
df_sample = df_sales.iloc[3, :]
series_sample = df_sample.iloc[6:]
df_sample
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="KCkY84feOfUu" outputId="5d2434f7-ef7a-47a8-9e80-46a4d23e91b7"
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series_sample.plot()
plt.legend()
plt.show()
# + [markdown] id="45kPq4yYOfUv"
# #### Pick a Time Series
#
# Let's try and find an interesting time series to forecast.
# + colab={"base_uri": "https://localhost:8080/", "height": 456} id="zvLKxPr9OfUv" outputId="a44bace7-fa45-489d-96fc-74585a410285"
df_sales_total_by_store = df_sales.groupby(['store_id']).sum()
df_sales_total_by_store
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="E9SaQDUZOfUv" outputId="045f4531-41c2-4125-96a7-cee43f50804b"
plt.rcParams['figure.figsize'] = [plot_x_size, 4]
df_sales_total_by_store.T.plot()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="zoxgHBtrOfUw" outputId="7eb0faf1-c21d-4ca8-80dc-3de487be64ea"
series = df_sales_total_by_store.iloc[0]
print(series.name)
print('Min Dates:' + str(series[series == series.min()].index.to_list()))
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series.plot()
plt.legend()
plt.show()
# + [markdown] id="A_B8vHefOfUw"
# ## 2. Pre-processing
#
# Before we build a forecasting model, let's check some properties of our time series.
# + [markdown] id="fVUA0chBOfUw"
# ### Is the series non-stationary?
#
# Let's check.
# + colab={"base_uri": "https://localhost:8080/"} id="wrKw7EoYOfUw" outputId="2feec086-ace4-4f21-87f2-8f5ec5ebb83f"
from statsmodels.tsa.stattools import adfuller
result = adfuller(series)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
# + [markdown] id="VQUEmUdpOfUx"
# ### Does differencing make the series stationary?
#
# Let's check.
# + colab={"base_uri": "https://localhost:8080/"} id="7i368QGsOfUx" outputId="49bd4764-d235-40c9-b0e9-b42e82e60473"
def difference(dataset, interval = 1):
diff = list()
for i in range(interval, len(dataset)):
value = dataset[i] - dataset[i - interval]
diff.append(value)
return np.array(diff)
def inverse_difference(history, yhat, interval=1):
return yhat + history[-interval]
series_d1 = difference(series)
result = adfuller(series_d1)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
# + [markdown] id="rjYYNGjTOfUx"
# ### Is the series seasonal?
#
# Let's check.
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="8Kbh-PuyOfUy" outputId="95d593d5-36e9-48f7-c353-7d8c49fb3414"
from statsmodels.graphics.tsaplots import plot_acf
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
plot_acf(series)
plt.show()
plot_acf(series, lags = 730, use_vlines = True)
plt.show()
# + [markdown] id="rb6KXvpEOfUy"
# ### Can we remove the seasonality?
#
# Let's check.
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="O-ivBmF4OfUy" outputId="13cc147b-3f17-40d8-fce8-de8b7f9203d8"
series_d7 = difference(series, 7)
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
plot_acf(series_d7)
plt.show()
plot_acf(series_d7, lags = 730, use_vlines = True)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="elfoUhemOfUy" outputId="b7bf1b77-b0dd-4303-c956-cd90fabf4b83"
series_d7_d30 = difference(series_d7, 30)
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
plot_acf(series_d7_d30)
plt.show()
plot_acf(series_d7_d30, lags = 730, use_vlines = True)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="0Fb145YaOfUz" outputId="c002effd-474b-4805-aff8-75026daf8b75"
result = adfuller(series_d7_d30)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="k5uqKkxBOfUz" outputId="669a8f7f-c6d6-4eb2-a136-89b398059d12"
series_d7_d30 = pd.Series(series_d7_d30)
series_d7_d30.index = date_list[37:]
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series_d7_d30.plot(label = 'Differenced Series')
plt.legend()
plt.show()
# + [markdown] id="jCnRtOEVOfU0"
# ### What now?
#
# At this point we have two options:
#
# - Model the seasonally differenced series, then reverse the differencing after making predictions.
#
# - Model the original series directly.
#
# While (vanilla) ARIMA requires a non-stationary and non-seasonal time series, these properties are not necessary for most non-parametric ML models.
# + [markdown] id="38Rd-9UeOfU0"
# ## 3. One-Step Prediction
#
# Let's build a model for making one-step forecasts.
#
# To do this, we first need to transform the time series data into a supervised learning dataset.
#
# In other words, we need to create a new dataset consisting of $X$ and $Y$ variables, where $X$ refers to the features and $Y$ refers to the target.
#
# ### How far do we lookback?
#
# To create the new $(X,Y)$ dataset, we first need to decide what the $X$ features are.
#
# For the moment, let's ignore any exogenous variables. In this case, what determines the $X$s is how far we <i>lookback</i>. In general, we can treat the lookback as a hyperparameter, which we will call `window_size`.
#
# <i>Advanced note:</i> Technically, we could build an entire methodology for feature engineering $X$.
#
# ### Test Set
#
# To test our model we will use the last 28 days of the series.
# + id="Xwfa6b9IOfU0"
### CREATE X,Y ####
def create_xy(series, window_size, prediction_horizon, shuffle = False):
x = []
y = []
for i in range(0, len(series)):
if len(series[(i + window_size):(i + window_size + prediction_horizon)]) < prediction_horizon:
break
x.append(series[i:(i + window_size)])
y.append(series[(i + window_size):(i + window_size + prediction_horizon)])
x = np.array(x)
y = np.array(y)
return x,y
# + colab={"base_uri": "https://localhost:8080/"} id="bkCFL3l3OfU0" outputId="55986e55-7961-441e-aafb-ba96c707fc07"
### HYPERPARAMETERS ###
window_size = 365
prediction_horizon = 1
### TRAIN VAL SPLIT ### (include shuffling later)
test_size = 28
split_time = len(series) - test_size
train_series = series[:split_time]
test_series = series[split_time - window_size:]
train_x, train_y = create_xy(train_series, window_size, prediction_horizon)
test_x, test_y = create_xy(test_series, window_size, prediction_horizon)
train_y = train_y.flatten()
test_y = test_y.flatten()
print(train_x.shape)
print(train_y.shape)
print(test_x.shape)
print(test_y.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="gHANelCHOfU1" outputId="78ca90b6-3427-4b2e-8b35-da0a942c9145"
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series[-test_size:].plot(label = 'CA_1 Test Series')
plt.legend()
plt.show()
# + [markdown] id="4iHnJwMcOfU1"
# ### LightGBM
#
# Now we can build a LightGBM model to forecast our time series.
#
# Gradient boosting is an ensemble method that combines multiple weak models to produce a single strong prediction model. The method involves constructing the model (called a <i>gradient boosting machine</i>) in a serial stage-wise manner by sequentially optimizing a differentiable loss function at each stage. Much like other boosting algorithms, the residual errors are passed to the next weak learner and trained.
#
# For this work, we use LightGBM, a gradient boosting framework designed for speed and efficiency. Specifically, the framework uses tree-based learning algorithms.
#
# To tune the model's hyperparameters, we use a combination of grid search and repeated k-fold cross validation, with some manual tuning. For more details, see the Hyperparameter Tuning notebook.
# + [markdown] id="bKh_Gu6-OfU2"
# Now we train the model on the full dataset and test it.
# + colab={"base_uri": "https://localhost:8080/"} id="9pNItUGaOfU2" outputId="a48a88e4-41e0-4b2f-deb4-fc431a3f96b6"
import lightgbm as lgb
params = {
'n_estimators': 2000,
'max_depth': 4,
'num_leaves': 2**4,
'learning_rate': 0.1,
'boosting_type': 'dart'
}
model = lgb.LGBMRegressor(first_metric_only = True, **params)
model.fit(train_x, train_y,
eval_metric = 'l1',
eval_set = [(test_x, test_y)],
#early_stopping_rounds = 10,
verbose = 0)
# + colab={"base_uri": "https://localhost:8080/", "height": 405} id="nck0f2RyOfU5" outputId="e8f8e75d-6e4f-4481-f059-f40c67695315"
forecast = model.predict(test_x)
s1_naive = series[-29:-1].to_numpy()
s7_naive = series[-35:-7].to_numpy()
s30_naive = series[-56:-28].to_numpy()
s365_naive = series[-364:-336].to_numpy()
print(' Naive MAE: %.4f' % (np.mean(np.abs(s1_naive - test_y))))
print(' s7-Naive MAE: %.4f' % (np.mean(np.abs(s7_naive - test_y))))
print(' s30-Naive MAE: %.4f' % (np.mean(np.abs(s30_naive - test_y))))
print('s365-Naive MAE: %.4f' % (np.mean(np.abs(s365_naive - test_y))))
print(' LightGBM MAE: %.4f' % (np.mean(np.abs(forecast - test_y))))
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series[-test_size:].plot(label = 'True')
plt.plot(forecast, label = 'Forecast')
plt.legend()
plt.show()
# + [markdown] id="0JM5O94QS6wY"
# ### Tuning Window Size
# How does our metric change as we extend the window size?
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="UYLLRLraTFRR" outputId="1db6b4fc-371f-489e-8e01-1ec487027b10"
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedKFold
params = {
'n_estimators': 2000,
'max_depth': 4,
'num_leaves': 2**4,
'learning_rate': 0.1,
'boosting_type': 'dart'
}
windows = [7, 30, 180, 365, 545, 730]
results = []
names = []
for w in windows:
window_size = w
train_x, train_y = create_xy(train_series, window_size, prediction_horizon)
train_y = train_y.flatten()
cv = RepeatedKFold(n_splits = 10, n_repeats = 3, random_state = 123)
scores = cross_val_score(lgb.LGBMRegressor(**params), train_x, train_y, scoring = 'neg_mean_absolute_error', cv = cv, n_jobs = -1)
results.append(scores)
names.append(w)
print('%3d --- MAE: %.3f (%.3f)' % (w, np.mean(scores), np.std(scores)))
plt.rcParams['figure.figsize'] = [plot_x_size, 5]
plt.boxplot(results, labels = names, showmeans = True)
plt.show()
# + [markdown] id="JsNAcgw5OfU7"
# ## 4. Multi-Step Prediction
#
# Suppose we were interested in forecasting the next $n$-days instead of just the next day.
#
# There are several approaches we can take to solve this problem.
# +
### HYPERPARAMETERS ###
window_size = 365
prediction_horizon = 1
### TRAIN VAL SPLIT ###
test_size = 28
split_time = len(series) - test_size
train_series = series[:split_time]
test_series = series[split_time - window_size:]
train_x, train_y = create_xy(train_series, window_size, prediction_horizon)
test_x, test_y = create_xy(test_series, window_size, prediction_horizon)
train_y = train_y.flatten()
test_y = test_y.flatten()
# + [markdown] id="JsNAcgw5OfU7"
# ### Recursive Forecasting
#
# In recursive forecasting, we first train a one-step model then generate a multi-step forecast by recursively feeding our predictions back into the model.
# +
params = {
'n_estimators': 2000,
'max_depth': 4,
'num_leaves': 2**4,
'learning_rate': 0.1,
'boosting_type': 'dart'
}
model = lgb.LGBMRegressor(first_metric_only = True, **params)
model.fit(train_x, train_y,
eval_metric = 'l1',
eval_set = [(test_x, test_y)],
#early_stopping_rounds = 10,
verbose = 0)
# +
recursive_x = test_x[0, :]
forecast_ms = []
for i in range(test_x.shape[0]):
pred = model.predict(recursive_x.reshape((1, recursive_x.shape[0])))
recursive_x = np.append(recursive_x[1:], pred)
forecast_ms.append(pred)
forecast_ms_rec = np.asarray(forecast_ms).flatten()
forecast_os = model.predict(test_x)
print(' One-Step MAE: %.4f' % (np.mean(np.abs(forecast_os - test_y))))
print('Multi-Step MAE: %.4f' % (np.mean(np.abs(forecast_ms_rec - test_y))))
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series[-test_size:].plot(label = 'True')
plt.plot(forecast_ms_rec, label = 'Forecast Multi-Step')
plt.plot(forecast_os, label = 'Forecast One-Step')
plt.legend()
plt.show()
# + [markdown] id="JsNAcgw5OfU7"
# ### Direct Forecasting
#
# In direct forecasting, we train $n$ independent models and generate a multi-step forecast by concatenating the $n$ predictions.
#
# For this implementation, we need to create a new $(X,Y)$ dataset, where $Y$ is now a vector of $n$ values.
# +
### HYPERPARAMETERS ###
window_size = 365
prediction_horizon = 28
### TRAIN VAL SPLIT ###
test_size = 28
split_time = len(series) - test_size
train_series = series[:split_time]
test_series = series[split_time - window_size:]
train_x, train_y = create_xy(train_series, window_size, prediction_horizon)
test_x, test_y = create_xy(test_series, window_size, prediction_horizon)
# +
from sklearn.multioutput import MultiOutputRegressor
model = MultiOutputRegressor(lgb.LGBMRegressor(), n_jobs = -1)
model.fit(train_x, train_y)
# +
forecast_ms_dir = model.predict(test_x)
print(' One-Step MAE: %.4f' % (np.mean(np.abs(forecast_os - test_y))))
print('Multi-Step MAE: %.4f' % (np.mean(np.abs(forecast_ms_dir - test_y))))
plt.rcParams['figure.figsize'] = [plot_x_size, plot_y_size]
series[-test_size:].plot(label = 'True')
plt.plot(forecast_ms_dir.T, label = 'Forecast Multi-Step')
plt.plot(forecast_os, label = 'Forecast One-Step')
plt.legend()
plt.show()
# + [markdown] id="JsNAcgw5OfU7"
# ### Single-Shot Forecasting
#
# In single-shot forecasting, we create a model that attempts to predict all $n$-steps simultaneously.
#
# Unfortunately, LightGBM (tree-based methods in general) does not support multi-output models.
# + [markdown] id="JsNAcgw5OfU7"
# ### Forecast Combination
#
# An easy way to improve forecast accuracy is to use several different methods on the same time series, and to average the resulting forecasts.
# +
forecast_ms_comb = 0.5*forecast_ms_dir.flatten() + 0.5*forecast_ms_rec
print(' Recursive MAE: %.4f' % (np.mean(np.abs(forecast_ms_rec - test_y))))
print(' Direct MAE: %.4f' % (np.mean(np.abs(forecast_ms_dir - test_y))))
print('Combination MAE: %.4f' % (np.mean(np.abs(forecast_ms_comb - test_y))))
series[-test_size:].plot(label = 'True')
plt.plot(forecast_ms_comb, label = 'Forecast Combination')
plt.show()
# + [markdown] id="JsNAcgw5OfU7"
# ## 5. Feature Importance
#
# One advantage of GBM models is that it can generate feature importance metrics based on the quality of the splits (or information gain).
# +
### HYPERPARAMETERS ###
window_size = 365
prediction_horizon = 1
### TRAIN VAL SPLIT ###
test_size = 28
split_time = len(series) - test_size
train_series = series[:split_time]
test_series = series[split_time - window_size:]
train_x, train_y = create_xy(train_series, window_size, prediction_horizon)
test_x, test_y = create_xy(test_series, window_size, prediction_horizon)
train_y = train_y.flatten()
test_y = test_y.flatten()
params = {
'n_estimators': 2000,
'max_depth': 4,
'num_leaves': 2**4,
'learning_rate': 0.1,
'boosting_type': 'dart'
}
model = lgb.LGBMRegressor(first_metric_only = True, **params)
feature_name_list = ['lag_' + str(i+1) for i in range(window_size)]
model.fit(train_x, train_y,
eval_metric = 'l1',
eval_set = [(test_x, test_y)],
#early_stopping_rounds = 10,
feature_name = feature_name_list,
verbose = 0)
# +
plt.rcParams['figure.figsize'] = [5, 5]
lgb.plot_importance(model, max_num_features = 15, importance_type = 'split')
plt.show()
# + [markdown] id="FJogEfYBOfUm"
# ## Summary
#
# In summary, this section has shown the following:
#
# - A quick exploration of the M5 dataset and its salient features
# - Checking the usual assumptions for classical time series analysis
# - Demonstrating the 'out-of-the-box' performance of LightGBM models
# - Demonstrating how tuning the lookback window can effect forecasting performance
# - Demonstrating how tuning the hyperparameters of a LightGBM model can improve performance
# - Summarizing the different approaches to multi-step forecasting
# - Illustrating that gradient boosting methods can measure feature importance
#
# Many of the methods and approaches shown above are easily extendable. Of note, because of the non-parametric nature of most machine learning models, performing classical inference tasks like hypothesis testing is more challenging. However, trends in ML research have been moving towards greater interpretability of so-called 'black box' models, such as SHAP for feature importance and even Bayesian approaches to neural networks for causality inference.
# + [markdown] id="FJogEfYBOfUm"
# ## References
#
# [1] <NAME>, <NAME>, and <NAME>. The M5 Accuracy competition: Results, findings and conlusions. 2020.
#
# [2] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. The M5 Uncertainty competition: Results, findings and conlusions. 2020.
#
# [3] <NAME>, and <NAME>. Evaluating quantile assessments. Operations research, 2009.
#
# [4] <NAME>, <NAME>, and <NAME>. Combining interval forecasts. Decision Analysis, 2017.
# -
#
# ```{toctree}
# :hidden:
# :titlesonly:
#
#
# lightgbm_m5_tuning
# lightgbm_jena_forecasting
# ```
#
| _build/jupyter_execute/08_WinningestMethods/lightgbm_m5_forecasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''base'': conda)'
# name: python3
# ---
# +
import pandas as pd
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# +
df_heart = pd.read_csv('../data/raw/heart.csv')
x = df_heart.drop(['target'], axis=1)
y = df_heart['target']
x_train, x_test, y_train, y_test = train_test_split(x,y, test_size=0.35)
boost_class = GradientBoostingClassifier( n_estimators=50).fit(x_train, y_train)
boost_pred = boost_class.predict(x_test)
print('BAG Accuracy: %f'%accuracy_score(boost_pred, y_test))
# -
| notebooks/boosting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Q-Learning using Monte-carlo Method
from graphics import *
import random
import numpy as np
# #### create 5x5 Grid world
win = GraphWin("My Window", 600,600)
win.setBackground(color_rgb(255,255,255))
def line(x1,y1,x2,y2):
"""
create a line between pt(x1,y1) and pt(x2,y2) and return it
"""
ln = Line(Point(x1,y1),Point(x2,y2))
ln.setOutline(color_rgb(0,0,0))
ln.setWidth(2)
return ln
# +
#border lines
border1 = line(50,50,550,50)
border1.draw(win)
border2 = line(550,50,550,550)
border2.draw(win)
border3 = line(50,550,550,550)
border3.draw(win)
border4 = line(50,50,50,550)
border4.draw(win)
#lines horizontal
ln1 = line(50,150,550,150)
ln1.draw(win)
ln2 = line(50,250,550,250)
ln2.draw(win)
ln3 = line(50,350,550,350)
ln3.draw(win)
ln4 = line(50,450,550,450)
ln4.draw(win)
#lines vertical
lnv1 = line(150,50,150,550)
lnv1.draw(win)
lnv2 = line(250,50,250,550)
lnv2.draw(win)
lnv3 = line(350,50,350,550)
lnv3.draw(win)
lnv4 = line(450,50,450,550)
lnv4.draw(win)
#create goal, obstacle and hole
rect_goal = Rectangle(Point(450,50), Point(550,150))
rect_goal.setFill(color_rgb(0,255,0))
rect_goal.draw(win)
rect_hole = Rectangle(Point(450,150), Point(550,250))
rect_hole.setFill(color_rgb(255,0,0))
rect_hole.draw(win)
rect_obs1 = Rectangle(Point(150,350), Point(250,450))
rect_obs1.setFill(color_rgb(0,0,0))
rect_obs1.draw(win)
rect_obs2 = Rectangle(Point(150,250), Point(250,350))
rect_obs2.setFill(color_rgb(0,0,0))
rect_obs2.draw(win)
# -
# #### Create agent look like circle
agent = Circle(Point(100,500), 25)
agent.setFill(color_rgb(0,0,255))
agent.draw(win)
# +
def up(agent):
val = agent.getCenter()
if (val.getY()) == 100 or (val.getY() == 500 and val.getX() == 200) :
pass
else :
agent.move(0,-100)
def down(agent):
val = agent.getCenter()
if val.getY() == 500 or (val.getX() == 200 and val.getY() == 200):
pass
else :
agent.move(0,100)
def right(agent):
val = agent.getCenter()
if val.getX() == 500 or (val.getX() == 100 and (val.getY() == 300 or val.getY() == 400)):
pass
else :
agent.move(100,0)
def left(agent):
val = agent.getCenter()
if val.getX() == 100 or (val.getX() == 300 or (val.getY() == 300 or val.getY() == 400)):
pass
else :
agent.move(-100,0)
def action(agent,val):
if val == 0:
up(agent)
elif val == 1:
down(agent)
elif val == 2:
right(agent)
else :
left(agent)
# -
# #### Reward for agent in the each state
reward = np.array([[-1,-1,-1,-1,-1],
[-1,-20,-1,-1,-1],
[-1,-20,-1,-1,-1],
[-1,-1,-1,-1,-100],
[-1,-1,-1,-1,100]])
# +
def rewards(agent):
val = agent.getCenter()
i = int(val.getX() / 100) - 1
j = int((500 - val.getY()) / 100)
return reward[j][i]
def observation(agent):
val = agent.getCenter()
i = int(val.getX() / 100) - 1
j = int((500 - val.getY()) / 100)
return [i,j]
def env_close():
win.close()
def reset(agent):
agent.move(-400,400)
def set_agent(agent,obs):
pos = agent.getCenter()
pos = [pos.getX(), pos.getY()]
refer_pos = [(obs[0]+1) * 100, (500 - (obs[1]*100))]
agent.move(refer_pos[0]-pos[0], refer_pos[1] - pos[1])
# -
# #### Create the value function and policy
value = np.array([[.0,.0,.0,.0,.0],
[.0,.0,.0,.0,.0],
[.0,.0,.0,.0,.0],
[.0,.0,.0,.0,0.0],
[.0,.0,.0,.0,100.0]])
| Basic_gridWorld/Q_Learning_using_MC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import importlib, utils2; importlib.reload(utils2)
from utils2 import *
np.set_printoptions(4)
cfg = K.tf.ConfigProto(gpu_options={'allow_growth':True})
K.set_session(K.tf.Session(config=cfg))
def tokenize(sent):
return [x.strip() for x in re.split('(\W+)?', sent) if x.strip()]
def parse_stories(lines):
data = []
story = []
for line in lines:
line = line.decode('utf-8').strip()
nid, line = line.split(' ', 1)
if int(nid) == 1: story = []
if '\t' in line:
q, a, supporting = line.split('\t')
q = tokenize(q)
substory = None
substory = [[str(i)+":"]+x for i,x in enumerate(story) if x]
data.append((substory, q, a))
story.append('')
else: story.append(tokenize(line))
return data
path = get_file('babi-tasks-v1-2.tar.gz',
origin = 'https://s3.amazonaws.com/text-datasets/babi_tasks_1-20_v1-2.tar.gz')
tar = tarfile.open(path)
challenges = {
'single_supporting_fact_10k': 'tasks_1-20_v1-2/en-10k/qa1_single-supporting-fact_{}.txt',
'two_supporting_facts_10k': 'tasks_1-20_v1-2/en-10k/qa2_two-supporting-facts_{}.txt',
'two_supporting_facts_1k': 'tasks_1-20_v1-2/en/qa2_two-supporting-facts_{}.txt',
}
challenge_type = 'single_supporting_fact_10k'
challenge = challenges[challenge_type]
def get_stories(f):
data = parse_stories(f.readlines())
return [(story, q, answer) for story, q, answer in data]
train_stories = get_stories(tar.extractfile(challenge.format('train')))
test_stories = get_stories(tar.extractfile(challenge.format('test')))
test_stories[0]
stories = train_stories + test_stories
story_maxlen = max((len(s) for x, _, _ in stories for s in x))
story_maxsents = max((len(x) for x, _, _ in stories))
query_maxlen = max(len(x) for _, x, _ in stories)
def do_flatten(el):
return isinstance(el, collections.Iterable) and not isinstance(el, (str, bytes))
def flatten(l):
for el in l:
if do_flatten(el): yield from flatten(el)
else: yield el
vocab = sorted(set(flatten(stories)))
vocab.insert(0, '<PAD>')
# +
# vocab
# -
vocab_size = len(vocab); vocab_size
test_stories[530]
word_idx = dict((c, i) for i, c in enumerate(vocab))
word_idx
def vectorize_stories(data, word_idx, story_maxlen, query_maxlen):
X = []; Xq = []; Y = []
for story, query , answer in data:
x = [[word_idx[w] for w in s] for s in story]
xq = [word_idx[w] for w in query]
y = [word_idx[answer]]
X.append(x); Xq.append(xq); Y.append(y)
return ([pad_sequences(x, maxlen=story_maxlen) for x in X],
pad_sequences(Xq, maxlen=query_maxlen), np.array(Y))
inputs_train, queries_train, answers_train = vectorize_stories(train_stories,
word_idx, story_maxlen, query_maxlen)
input_test, queries_test, answers_test = vectorize_stories(test_stories,
word_idx, story_maxlen, query_maxlen)
def stack_inputs(inputs):
for i, it in enumerate(inputs):
inputs[i] = np.concatenate([it,
np.zeros((story_maxsents-it.shape[0], story_maxlen), 'int')])
return np.stack(inputs)
inputs_train = stack_inputs(inputs_train)
input_test = stack_inputs(input_test)
inps = [inputs_train, queries_train]
val_inps = [input_test, queries_test]
| memory_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NER with BERT in Spark NLP
# !python -V
# ## Installation
# +
import os
# Install java
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# ! java -version
# Install pyspark
# ! pip install --ignore-installed pyspark==2.4.4
# Install Spark NLP
# ! pip install --ignore-installed spark-nlp==2.5.1
# -
# ## Import libraries and download datasets
# +
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
import sparknlp
from sparknlp.annotator import *
from sparknlp.common import *
from sparknlp.base import *
# -
spark = sparknlp.start()
#spark = sparknlp.start(gpu=True)
print("Spark NLP version: ", sparknlp.version())
print("Apache Spark version: ", spark.version)
# +
def start(gpu=False):
builder = SparkSession.builder \
.appName("Spark NLP") \
.master("local[*]") \
.config("spark.driver.memory", "8G") \
.config("spark.serializer", "org.apache.spark.serializer.KryoSerializer")\
.config("spark.kryoserializer.buffer.max", "1000M")
if gpu:
builder.config("spark.jars.packages", "com.johnsnowlabs.nlp:spark-nlp-gpu_2.11:2.5.1")
else:
builder.config("spark.jars.packages", "com.johnsnowlabs.nlp:spark-nlp_2.11:2.5.1")
return builder.getOrCreate()
spark = start(gpu=False)
# +
from urllib.request import urlretrieve
urlretrieve('https://github.com/JohnSnowLabs/spark-nlp/raw/master/src/test/resources/conll2003/eng.train',
'eng.train')
urlretrieve('https://github.com/JohnSnowLabs/spark-nlp/raw/master/src/test/resources/conll2003/eng.testa',
'eng.testa')
# +
with open("eng.train") as f:
c=f.read()
print (c[:500])
# -
# ## Building NER pipeline
# +
from sparknlp.training import CoNLL
training_data = CoNLL().readDataset(spark, './eng.train')
training_data.show(3)
# -
training_data.count()
# ### Loading Bert
# In Spark NLP, we have four pre-trained variants of BERT: bert_base_uncased , bert_base_cased , bert_large_uncased , bert_large_cased . Which one to use depends on your use case, train set, and the complexity of the task you are trying to model.
#
# In the code snippet above, we basically load the bert_base_cased version from Spark NLP public resources and point thesentenceand token columns in setInputCols(). In short, BertEmbeddings() annotator will take sentence and token columns and populate Bert embeddings in bert column. In general, each word is translated to a 768-dimensional vector. The parametersetPoolingLayer() can be set to 0 as the first layer and fastest, -1 as the last layer and -2 as the second-to-last-hidden layer.
#
# As explained by the authors of official BERT paper, different BERT layers capture different information. The last layer is too closed to the target functions (i.e. masked language model and next sentence prediction) during pre-training, therefore it may be biased to those targets. If you want to use the last hidden layer anyway, please feel free to set pooling_layer=-1. Intuitively, pooling_layer=-1 is close to the training output, so it may be biased to the training targets. If you don't fine-tune the model, then this could lead to a bad representation. That said, it is a matter of trade-off between model accuracy and computational resources you have.
bert_annotator = BertEmbeddings.pretrained('bert_base_cased', 'en') \
.setInputCols(["sentence",'token'])\
.setOutputCol("bert")\
.setCaseSensitive(False)\
.setPoolingLayer(0)
# +
# BertEmbeddings.load("local/path/")
# +
from sparknlp.training import CoNLL
test_data = CoNLL().readDataset(spark, './eng.testa')
test_data = bert_annotator.transform(test_data)
test_data.show(3)
# +
#test_data.limit(1000).write.parquet("test_withEmbeds.parquet")
# -
test_data.select("bert.result","bert.embeddings",'label.result').show()
# +
import numpy as np
emb_vector = np.array(test_data.select("bert.embeddings").take(1))
emb_vector
# +
nerTagger = NerDLApproach()\
.setInputCols(["sentence", "token", "bert"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(1)\
.setLr(0.001)\
.setPo(0.005)\
.setBatchSize(8)\
.setRandomSeed(0)\
.setVerbose(1)\
.setValidationSplit(0.2)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset("test_withEmbeds.parquet")
pipeline = Pipeline(
stages = [
bert_annotator,
nerTagger
])
# -
# You can also set learning rate ( setLr ), learning rate decay coefficient ( setPo ), setBatchSize and setDropout rate. Please see the official repo for the entire list.
# +
# %%time
ner_model = pipeline.fit(training_data.limit(1000))
# -
ner_model
# +
# on COLAB, it takes 30 min to train entire trainset (conll2003) for 10 epochs on GPU with layer = 0
# -
predictions = ner_model.transform(test_data)
predictions.show(3)
predictions.select('token.result','label.result','ner.result').show(truncate=40)
predictions.printSchema()
# +
import pyspark.sql.functions as F
predictions.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).show(truncate=False)
# -
# ### Loading from local
# +
# loading the one trained 10 epochs on GPU with entire train set
loaded_ner_model = NerDLModel.load("NER_bert_20200226")\
.setInputCols(["sentence", "token", "bert"])\
.setOutputCol("ner")
# +
predictions_loaded = loaded_ner_model.transform(test_data)
predictions_loaded.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).show(30, truncate=False)
# +
import pandas as pd
df = predictions_loaded.select('token.result','label.result','ner.result').toPandas()
df
# -
# ### Bert with poolingLayer -2
bert_annotator.setPoolingLayer(-2)
pipeline = Pipeline(
stages = [
bert_annotator,
nerTagger
])
ner_model_v2 = pipeline.fit(training_data.limit(1000))
# +
predictions_v2 = ner_model_v2.transform(test_data.limit(10))
predictions_v2.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).show(truncate=False)
# -
# ### with Glove Embeddings
# +
glove = WordEmbeddingsModel().pretrained() \
.setInputCols(["sentence",'token'])\
.setOutputCol("glove")\
.setCaseSensitive(False)
test_data = CoNLL().readDataset(spark, './eng.testa')
test_data = glove.transform(test_data.limit(1000))
test_data.write.parquet("test_withGloveEmbeds.parquet")
# +
nerTagger.setInputCols(["sentence", "token", "glove"])
nerTagger.setTestDataset("test_withGloveEmbeds.parquet")
glove_pipeline = Pipeline(
stages = [
glove,
nerTagger
])
# +
# %%time
ner_model_v3 = glove_pipeline.fit(training_data.limit(1000))
# +
predictions_v3 = ner_model_v3.transform(test_data.limit(10))
# test_data.sample(False,0.1,0)
predictions_v3.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).show(truncate=False)
# -
np.array (predictions.select('token.result').take(1))[0][0]
# +
import pandas as pd
tokens = np.array (predictions.select('token.result').take(1))[0][0]
ground = np.array (predictions.select('label.result').take(1))[0][0]
label_bert_0 = np.array (predictions.select('ner.result').take(1))[0][0]
label_bert_2 = np.array (predictions_v2.select('ner.result').take(1))[0][0]
label_glove = np.array (predictions_v3.select('ner.result').take(1))[0][0]
pd.DataFrame({'token':tokens,
'ground':ground,
'label_bert_0':label_bert_0,
'label_bert_2':label_bert_2,
'label_glove':label_glove})
# -
# ### Saving the trained model
ner_model_v3.stages
ner_model_v3.stages[1].write().overwrite().save('NER_bert_20200219')
# ## Prediction Pipeline
# +
document = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')
token = Tokenizer()\
.setInputCols(['sentence'])\
.setOutputCol('token')
bert = BertEmbeddings.pretrained('bert_base_cased', 'en') \
.setInputCols(["sentence",'token'])\
.setOutputCol("bert")\
.setCaseSensitive(False)
loaded_ner_model = NerDLModel.load("NER_bert_20200219")\
.setInputCols(["sentence", "token", "bert"])\
.setOutputCol("ner")
converter = NerConverter()\
.setInputCols(["document", "token", "ner"])\
.setOutputCol("ner_span")
ner_prediction_pipeline = Pipeline(
stages = [
document,
sentence,
token,
bert,
loaded_ner_model,
converter])
# +
empty_data = spark.createDataFrame([['']]).toDF("text")
empty_data.show()
# -
prediction_model = ner_prediction_pipeline.fit(empty_data)
text = "<NAME> is a nice guy and lives in New York."
sample_data = spark.createDataFrame([[text]]).toDF("text")
sample_data.show()
# +
preds = prediction_model.transform(sample_data)
preds.show()
# -
preds.select('ner_span.result').take(1)
# +
preds.select(F.explode(F.arrays_zip("ner_span.result","ner_span.metadata")).alias("entities")) \
.select(F.expr("entities['0']").alias("chunk"),
F.expr("entities['1'].entity").alias("entity")).show(truncate=False)
# +
document = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')
token = Tokenizer()\
.setInputCols(['sentence'])\
.setOutputCol('token')
loaded_ner_model = NerDLModel.load("NER_bert_20200219")\
.setInputCols(["sentence", "token", "glove"])\
.setOutputCol("ner")
converter = NerConverter()\
.setInputCols(["document", "token", "ner"])\
.setOutputCol("ner_span")
glove_ner_prediction_pipeline = Pipeline(
stages = [
document,
sentence,
token,
glove,
loaded_ner_model,
converter])
# -
glove_prediction_model = glove_ner_prediction_pipeline.fit(empty_data)
# +
preds = glove_prediction_model.transform(sample_data)
preds.show()
# +
preds.select(F.explode(F.arrays_zip("ner_span.result","ner_span.metadata")).alias("entities")) \
.select(F.expr("entities['0']").alias("chunk"),
F.expr("entities['1'].entity").alias("entity")).show(truncate=False)
# -
# ### Pretrained Pipelines
# +
from sparknlp.pretrained import PretrainedPipeline
pretrained_pipeline = PretrainedPipeline('recognize_entities_dl', lang='en')
#onto_recognize_entities_sm
#explain_document_dl
# +
text = "The Mona Lisa is a 16th century oil painting created by Leonardo. It's held at the Louvre in Paris."
result = pretrained_pipeline.annotate(text)
list(zip(result['token'], result['ner']))
# -
pretrained_pipeline2 = PretrainedPipeline('explain_document_dl', lang='en')
# +
text = "The Mona Lisa is a 16th centry oil painting created by Leonrdo. It's held at the Louvre in Paris."
result2 = pretrained_pipeline2.annotate(text)
result2
list(zip(result2['token'], result2['checked'], result2['pos'], result2['ner'], result2['lemma'], result2['stem']))
# +
xx= pretrained_pipeline2.fullAnnotate(text)
[(n.result, n.metadata['entity']) for n in xx['ner_span']]
# -
# ## Using your own custom Word Embedding
custom_embeddings = WordEmbeddings()\
.setInputCols(["sentence", "token"])\
.setOutputCol("glove")\
.setStoragePath('/Users/vkocaman/cache_pretrained/PubMed-shuffle-win-2.bin', "BINARY")\
.setDimension(200)
custom_embeddings.fit(training_data.limit(10)).transform(training_data.limit(10)).show()
# ## creating your own CoNLL dataset
# +
import json
import os
from pyspark.ml import Pipeline
from sparknlp.base import *
from sparknlp.annotator import *
import sparknlp
spark = sparknlp.start()
def get_ann_pipeline ():
document_assembler = DocumentAssembler() \
.setInputCol("text")\
.setOutputCol('document')
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')\
.setCustomBounds(['\n'])
tokenizer = Tokenizer() \
.setInputCols(["sentence"]) \
.setOutputCol("token")
pos = PerceptronModel.pretrained() \
.setInputCols(["sentence", "token"]) \
.setOutputCol("pos")
embeddings = WordEmbeddingsModel.pretrained()\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
ner_model = NerDLModel.pretrained() \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter()\
.setInputCols(["sentence", "token", "ner"])\
.setOutputCol("ner_chunk")
ner_pipeline = Pipeline(
stages = [
document_assembler,
sentence,
tokenizer,
pos,
embeddings,
ner_model,
ner_converter
]
)
empty_data = spark.createDataFrame([[""]]).toDF("text")
ner_pipelineFit = ner_pipeline.fit(empty_data)
ner_lp_pipeline = LightPipeline(ner_pipelineFit)
print ("Spark NLP NER lightpipeline is created")
return ner_lp_pipeline
# -
conll_pipeline = get_ann_pipeline ()
parsed = conll_pipeline.annotate ("<NAME> is a nice guy and lives in New York.")
parsed
# +
conll_lines=''
for token, pos, ner in zip(parsed['token'],parsed['pos'],parsed['ner']):
conll_lines += "{} {} {} {}\n".format(token, pos, pos, ner)
print(conll_lines)
# -
| tutorials/blogposts/3.NER_with_BERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°11
#
# <a id='p1'></a>
# ## I.- Problema 01
#
# Lista de actos delictivos registrados por el Service de police de la Ville de Montréal (SPVM).
#
#
# <img src="http://henriquecapriles.com/wp-content/uploads/2017/02/femina_detenida-1080x675.jpg" width="480" height="360" align="center"/>
#
# El conjunto de datos en estudio `interventionscitoyendo.csv` corresponde a todos los delitos entre 2015 y agosto de 2020en Montreal. Cada delito está asociado en grandes categorías, y hay información sobre la ubicación, el momento del día, etc.
#
# > **Nota**: Para más información seguir el siguiente el [link](https://donnees.montreal.ca/ville-de-montreal/actes-criminels).
# +
# librerias
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
from statsmodels.tsa.statespace.sarimax import SARIMAX
from metrics_regression import *
# graficos incrustados
plt.style.use('fivethirtyeight')
# %matplotlib inline
# parametros esteticos de seaborn
sns.set_palette("deep", desat=.6)
sns.set_context(rc={"figure.figsize": (12, 4)})
# +
# read data
validate_categorie = [
'Introduction', 'Méfait','Vol dans / sur véhicule à moteur', 'Vol de véhicule à moteur',
]
df = pd.read_csv(os.path.join("data","interventionscitoyendo.csv"), sep=",", encoding='latin-1')
df.columns = df.columns.str.lower()
df['date'] = pd.to_datetime(df['date'], format='%Y-%m-%d')
df = df.loc[lambda x: x['categorie'].isin(validate_categorie)]
df = df.sort_values(['categorie','date'])
df.head()
# -
# Como tenemos muchos datos por categoría a nivel de día, agruparemos a nivel de **semanas** y separaremos cada serie temporal.
cols = ['date','pdq']
y_s1 = df.loc[lambda x: x.categorie == validate_categorie[0] ][cols].set_index('date').resample('W').mean()
y_s2 = df.loc[lambda x: x.categorie == validate_categorie[1] ][cols].set_index('date').resample('W').mean()
y_s3 = df.loc[lambda x: x.categorie == validate_categorie[2] ][cols].set_index('date').resample('W').mean()
y_s4 = df.loc[lambda x: x.categorie == validate_categorie[3] ][cols].set_index('date').resample('W').mean()
# El objetivo de este laboratorio es poder realizar un análisis completo del conjunto de datos en estudio, para eso debe responder las siguientes preguntas:
#
# 1. Realizar un gráfico para cada serie temporal $y\_{si}, i =1,2,3,4$.
#
# graficar datos
y_s1.plot(figsize=(15, 3),color = 'blue')
plt.show()
# graficar datos
y_s2.plot(figsize=(15, 3),color = 'red')
plt.show()
# graficar datos
y_s3.plot(figsize=(15, 3),color = 'green')
plt.show()
# graficar datos
y_s4.plot(figsize=(15, 3),color = 'orange')
plt.show()
# 2. Escoger alguna serie temporal $y\_{si}, i =1,2,3,4$. Luego:
#
# * Realice un análisis exploratorio de la serie temporal escogida
# * Aplicar el modelo de pronóstico $SARIMA(p,d,q)x(P,D,Q,S)$, probando varias configuraciones de los hiperparámetros. Encuentre la mejor configuración. Concluya.
# * Para el mejor modelo encontrado, verificar si el residuo corresponde a un ruido blanco.
#
# > **Hint**: Tome como `target_date` = '2021-01-01'. Recuerde considerar que su columna de valores se llama `pdq`.
#
# Se escogió i=1
# +
# creando clase SarimaModels
class SarimaModels:
def __init__(self,params):
self.params = params
@property
def name_model(self):
return f"SARIMA_{self.params[0]}X{self.params[1]}".replace(' ','')
@staticmethod
def test_train_model(y,date):
mask_ds = y.index < date
y_train = y[mask_ds]
y_test = y[~mask_ds]
return y_train, y_test
def fit_model(self,y,date):
y_train, y_test = self.test_train_model(y,date )
model = SARIMAX(y_train,
order=self.params[0],
seasonal_order=self.params[1],
enforce_stationarity=False,
enforce_invertibility=False)
model_fit = model.fit(disp=0)
return model_fit
def df_testig(self,y,date):
y_train, y_test = self.test_train_model(y,date )
model = SARIMAX(y_train,
order=self.params[0],
seasonal_order=self.params[1],
enforce_stationarity=False,
enforce_invertibility=False)
model_fit = model.fit(disp=0)
start_index = y_test.index.min()
end_index = y_test.index.max()
preds = model_fit.get_prediction(start=start_index,end=end_index, dynamic=False)
df_temp = pd.DataFrame(
{
'y':y_test['pdq'],
'yhat': preds.predicted_mean
}
)
return df_temp
def metrics(self,y,date):
df_temp = self.df_testig(y,date)
df_metrics = summary_metrics(df_temp)
df_metrics['model'] = self.name_model
return df_metrics
# definir parametros
import itertools
p = d = q = range(0, 2)
pdq = list(itertools.product(p, d, q))
seasonal_pdq = [(x[0], x[1], x[2], 12) for x in list(itertools.product(p, d, q))]
params = list(itertools.product(pdq,seasonal_pdq))
target_date = '2021-01-01'
# -
# Análisis Exploratorio
y_s1.describe()
# diagrama de caja y bigotes
fig, ax = plt.subplots(figsize=(15,6))
sns.boxplot(y_s1.pdq.index.year, y_s1.pdq, ax=ax)
plt.show()
# +
from pylab import rcParams
import statsmodels.api as sm
import matplotlib.pyplot as plt
rcParams['figure.figsize'] = 18, 8
decomposition = sm.tsa.seasonal_decompose(y_s1, model='multiplicative')
fig = decomposition.plot()
plt.show()
# +
from statsmodels.graphics.tsaplots import plot_acf
from statsmodels.graphics.tsaplots import plot_pacf
from matplotlib import pyplot
pyplot.figure(figsize=(12,9))
# acf
pyplot.subplot(211)
plot_acf(y_s1.pdq, ax=pyplot.gca(), lags = 30)
#pacf
pyplot.subplot(212)
plot_pacf(y_s1.pdq, ax=pyplot.gca(), lags = 30)
pyplot.show()
# +
from statsmodels.tsa.stattools import adfuller
#test Dickey-Fulle:
print ('Resultados del test de Dickey-Fuller:')
pdqtest = adfuller(y_s1.pdq, autolag='AIC')
pdqoutput = pd.Series(pdqtest[0:4],
index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
print(pdqoutput)
# -
# Aplicar modelo
# +
# iterar para los distintos escenarios
frames = []
for param in params:
try:
sarima_model = SarimaModels(param)
df_metrics = sarima_model.metrics(y_s1,target_date)
frames.append(df_metrics)
except:
pass
# -
# juntar resultados de las métricas y comparar
df_metrics_result = pd.concat(frames)
df_metrics_result.sort_values(['mae','mape'])
# +
# ajustar mejor modelo
param = [(0,0,0),(1,0,1,12)]
sarima_model = SarimaModels(param)
model_fit = sarima_model.fit_model(y_s1,target_date)
best_model = sarima_model.df_testig(y_s1,target_date)
best_model.head()
# +
# graficar mejor modelo
preds = best_model['yhat']
ax = y_s1['2015':].plot(label='observed')
preds.plot(ax=ax, label='Forecast', alpha=.7, figsize=(14, 7))
ax.set_xlabel('date')
ax.set_ylabel('pdq')
plt.legend()
plt.show()
# -
# resultados del error
model_fit.plot_diagnostics(figsize=(16, 8))
plt.show()
# * **gráfico 01** (standarized residual): Este gráfico nos muestra el error estandarizado en el tiempo. En este caso se observa que esta nueva serie de tiempo corresponde a una serie estacionaria que oscila entorno al cero, es decir, un ruido blanco.
#
#
# * **gráfico 02** (histogram plus estimated density): Este gráfico nos muestra el histograma del error. En este caso, el histograma es muy similar al histograma de una variable $\mathcal{N}(0,1)$ (ruido blanco).
#
#
# * **gráfico 03** (normal QQ): el gráfico Q-Q ("Q" viene de cuantil) es un método gráfico para el diagnóstico de diferencias entre la distribución de probabilidad de una población de la que se ha extraído una muestra aleatoria y una distribución usada para la comparación. En este caso se comparar la distribución del error versus una distribución normal. Cuando mejor es el ajuste lineal sobre los puntos, más parecida es la distribución entre la muestra obtenida y la distribución de prueba (distribución normal).
#
#
#
#
# * **gráfico 04** (correlogram): Este gráfico nos muestra el gráfico de autocorrelación entre las variables del error, se observa que no hay correlación entre ninguna de las variables, por lo que se puedan dar indicios de independencia entre las variables.
#
#
# En conclusión, el error asociado al modelo en estudio corresponde a un ruido blanco.
| labs/lab_11_resuelto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `Distribute` Parallel Processing Basics
# Due to a number of limitations involving data passed to processes using `multiprocessing.Pool()`, I've implemented a similar class called `Distribute()`. The primary difference is that Distribute is meant to distribute chunks of data for parallel processing, so your map function should parse multiple values. There are currently two functions in Distribute:
#
# * `.map_chunk()` simply applies a function to a list of elements and returns a list of parsed elements.
# * `.map()` applies a function to a single element. Same as `multiprocessing.Pool().map()`.
#from IPython import get_ipython
import sys
sys.path.append('..')
import doctable
# ## `.map()` Method
# Unsurprisingly, this method works exactly like the `multiprocessing.Pool().map()`. Simply provide a sequence of elements and a function to apply to them, and this method will parse all the elements in parallel.
# +
def multiply(x, y=2):
return x * y
nums = list(range(5))
with doctable.Distribute(3) as d:
res = d.map(multiply, nums)
print(res)
# pass any argument to your function here.
# we try multiplying by 5 instead of 2.
res = d.map(multiply, nums, 5)
print(res)
# -
# ## `.map_chunk()` Method
# Allows you to write map functions that processes a chunk of your data at a time. This is the lowest-level method for distributed processing.
# +
# map function to multiply 1.275 by each num and return a list
def multiply_nums(nums):
return [num*1.275 for num in nums]
# use Distribute(3) to create three separate processes
nums = list(range(1000))
with doctable.Distribute(3) as d:
res = d.map_chunk(multiply_nums, nums)
# won't create new process at all. good for testing
with doctable.Distribute(1) as d:
res = d.map_chunk(multiply_nums, nums)
res[:3]
| examples/distributed_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fonction pour formatage dates : depuis format (lettre : jour-mois-année) vers (num)
import datetime
import time
import locale
locale.setlocale(locale.LC_ALL,"french")
from datetime import datetime
def letter_to_date(num_date):
#convertir en datetime object
date_obj = datetime.strptime(num_date,'%d %B %Y')
#formater
return date_obj.strftime('%d-%m-%Y')
# **Testing :**
date_str = '10 Décembre 2011'
letter_to_date(date_str)
| Time/Function-LetterToDatetime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.4 64-bit
# language: python
# name: python3
# ---
# + id="jIpqPLCSfifl"
# #!pip3 install vosk
# #!pip3 install pydub # for audio processing
# #!pip3 install rpunct
# #!pip3 install transformers
# + id="LCnNrCJpfifn"
from vosk import Model, KaldiRecognizer
from pydub import AudioSegment
import json
# + id="HCU7nYeSfifo" outputId="83fd5742-0d8c-4e98-f040-e6cdb603fa3d" colab={"base_uri": "https://localhost:8080/"}
FRAME_RATE = 16000 # Hz
CHANNEL = 1 # Mono
model = Model(model_name="vosk-model-en-us-0.22") # Model name is "vosk-model-en-us-0.22"
rec = KaldiRecognizer(model, FRAME_RATE)
rec.SetWords(True)
# + id="lTO1zAPsfp9W" outputId="a6396eaa-1fc8-4914-bf58-6a94de0a6e07" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/drive')
# + id="UPBW2ReKfifo"
mp3 = AudioSegment.from_mp3("/content/drive/MyDrive/data/speech2sum/marketplace.mp3")
mp3 = mp3.set_frame_rate(FRAME_RATE)
mp3 = mp3.set_channels(CHANNEL)
# + id="HPcmIylktJQN"
rec.AcceptWaveform(mp3.raw_data) #raw_data is the binary represantaion of the original file
result = rec.Result() #outputs a JSON file
raw_text = json.loads(result)
# + id="MnSpAet7tnqE" outputId="19272ae2-508b-436d-f802-a6c973beef32" colab={"base_uri": "https://localhost:8080/"}
raw_text
# + id="xi1zh391uFDS" outputId="0604ab06-2334-435c-b5da-bdfca3862fa7" colab={"base_uri": "https://localhost:8080/", "height": 140}
text=raw_text['text']
text
# + id="8iF5PtGpwylo"
import torch
from rpunct import RestorePuncts
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
#use_cuda= False
# The default language is 'english'
rpunct = RestorePuncts()
# + id="wdkdnN-z1IA_" outputId="da46f4d4-3e46-4b45-899e-1a4ddf86acfd" colab={"base_uri": "https://localhost:8080/", "height": 140}
rpunct.punctuate(text)
| Natural-Language-Processing/speech2text/sppech2summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Multiple Sequence Alignment**
# This notebook covers work done on MSA and any strategies used to improve on the alignments
# ## **Goals**
# 1. Develop a Multiple Sequence Alignment pipeline
# 2. Make decision on which tools to use for MSA and visualization
# 3. Conduct Multiple Sequence Alignment on the generated FASTA format sequences
# 4. Visualize the results using an assortment of tools
# 5. Trim and develop a clean, suitable data set for phylogenetic analysis:
# 5.1. Exclude unaligned sequences (non-homologous)
# 5.2. Exclude short sequences
#
# ## **Tasks**
# 1. [Installing the bioinformatics tools for MSA](./03.01.MSA_tools_installation.ipynb)
# 2. Setting up the tools: writting the scripts, setting parameters and testing using samples from test data
# 3. Evaluating the test data results from various tools to enable decision making on which tools to use for what purpose
# 4. Conduct MSA on the subsets of data using the best selected tools
# 5. Visuallize the aligned sequences and trim the sequences to collumns within the 658 5' region of the COI-5P barcode
# 6. Finally develop a suitable data set for phylogenetic analysis
# ### **Multiple Sequence Alignment tools.**
# 1. [MUSCLE.](http://www.drive5.com/muscle/)
# It is problematic to align large number of sequences using global alignment algorithims used by muscle as explained in [Very large alignments are usually a bad idea](http://www.drive5.com/muscle/manual/bigalignments.html). Clustering highly identical (95% or 90% identity) help reduce the the sequences and challanges faced.
# 2. [T-Coffee(Tree based Consistency Objective Function For AlignmEnt Evaluation)](https://github.com/cbcrg/tcoffee). [The regressive mode of T-Coffee](https://github.com/cbcrg/tcoffee/blob/master/docs/tcoffee_quickstart_regressive.rst) is [described as most suitable for large datasets](https://www.biorxiv.org/content/10.1101/490235v1.full) by <NAME> et. al (2018).
# 3. [MAFFT Version 7](https://mafft.cbrc.jp/alignment/software/). For large datasets: [Tips for handling a large dataset](https://mafft.cbrc.jp/alignment/software/tips.html). More published by [<NAME> et. al (2018)](https://academic.oup.com/bioinformatics/article/34/14/2490/4916099)
# 4. [SATé(Simultaneous Alignment and Tree Estimation)](http://sysbio.oxfordjournals.org/content/61/1/90.abstract?sid=58895a54-2686-4b58-a676-3cc4d73a3b76): From GitHub [source code](https://github.com/sate-dev/sate-core) using [sate-tools-linux](https://github.com/sate-dev/sate-tools-linux) tools
# 5. [PASTA(Practical Alignment using Sate and TrAnsitivity)](https://www.liebertpub.com/doi/full/10.1089/cmb.2014.0156): From GitHub [Source code](https://github.com/smirarab/pasta) [(Tutorial)](https://github.com/smirarab/pasta/blob/master/pasta-doc/pasta-tutorial.md)
# 6. Other tools; [SEPP](https://github.com/smirarab/sepp), [UPP](https://github.com/smirarab/sepp/blob/master/README.UPP.md) and [HMMER](http://hmmer.org/)
# 7. Visualized the multiple sequence alignments using [jalview](http://www.jalview.org/download) for datasets upto a few thousands and [Seaview](http://doua.prabi.fr/software/seaview) by [Gouy et al.](https://academic.oup.com/mbe/article/27/2/221/970247) for bigger data sets. Seaview uses the FLTK project (installed separately) for its user interface.
#
# #### MSA evaluation methods used in T_Coffee:
# We used the following [sequence based methods](https://tcoffee.readthedocs.io/en/latest/tcoffee_main_documentation.html#sequence-based-methods) to evaluate our MSAs:
# 1. [Computing the CORE index of any alignment
# ](https://tcoffee.readthedocs.io/en/latest/tcoffee_main_documentation.html#computing-the-local-core-index).
# 2. Evaluating the [Transitive Consistency Score (TCS)](https://tcoffee.readthedocs.io/en/latest/tcoffee_main_documentation.html#transitive-consistency-score-tcs) of an MSA. The scores generated here are usefull in filtering our sequences and in phylogenetic inference based on herogenous site evolutionary rates.
#
# ### **MSA Tools Installation**
# **MUSCLE, T-Coffee and MAFFT** are all available on conda environment bioconda channel and are easily installed to our analysis anaconda3 environment (coi_env):
# + language="bash"
# conda env list
# #conda install -n coi_env -c bioconda t-coffee muscle mafft
# -
# **SATé** and **PASTA** up-to-date source codes are available on GitHub and are installed as described below.
# SATé *(~on its own~)** was not compatible with Python3 and some of it's dependencies were no longer available for Python2. I tried to upgrade it to accommodate Python3 but gave up after a few days.
# >(~on its own~)*: SATé is a crucial part of PASTA and UPP/SEPP by extention and won't operate without some inbuilt SATé modules
# + language="bash"
# # PASTA
# #cd ~/bioinformatics/github/co1_metaanalysis/code/tools/pasta_code/
# #git clone https://github.com/smirarab/pasta.git
# #git clone https://github.com/sate-dev/sate-tools-linux.git
# cd pasta/
# sudo python3 setup.py develop
# chmod +x run_pasta.py
# chmod +x run_pasta_gui.py
# chmod +x run_seqtools.py
#
# # SATé
# #cd ~/bioinformatics/github/co1_metaanalysis/code/tools/sate/
# #git clone https://github.com/sate-dev/sate-core.git
# #git clone https://github.com/sate-dev/sate-tools-linux.git
# sudo python3 setup.py develop #
# chmod +x run_sate.py
# chmod +x run_sate_gui.py
# -
# For the **other tools**:
# 1. SEPP (SATe-enabled Phylogenetic Placement): phylogenetic placement of short reads into reference alignments and trees.
# 2. UPP (Ultra-large alignments using Phylogeny-aware Profiles): alignment of very large datasets, potentially containing fragmentary data.
# 4. HMMER (I have **NOT** used so far for alignment on it's own): Uses probabilistic models called profile hidden Markov models (profile HMMs) for sequence alignment among other functions
#
# SEPP and UPP source codes are available on GitHub as a single package and were installed as follows:
# + language="bash"
# # SEPP
# #cd ~/bioinformatics/github/co1_metaanalysis/code/tools/sepp
# #git clone https://github.com/smirarab/sepp.git
# python3 setup.py config -c
# python3 setup.py install
# -
# ### **Visualization Tools**
# 1. **Seaview**: The most suitable
# 2. **Jalview**: Good for fewer sequences
# 3. **SuiteMSA**
#
# All are java based programs and were installed using their source codes
| code/03.00.Multiple_sequence_Alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.2
# language: ''
# name: sagemath
# ---
# ## Exercise 4 ~ Nibblet Blockcipher
# TEST PARAMETERS
"""
s = [[[6, 2, 13, 7, 11, 0, 14, 8, 3, 10, 4, 1, 9, 15, 12, 5], [15, 9, 2, 3, 8, 0, 4, 1, 13, 14, 11, 12, 6, 5, 7, 10], [5, 14, 0, 12, 1, 2, 11, 3, 9, 7, 6, 4, 10, 8, 15, 13], [9, 15, 11, 6, 2, 14, 5, 3, 0, 7, 8, 12, 10, 13, 4, 1], [3, 10, 4, 1, 14, 13, 9, 7, 0, 12, 8, 2, 11, 5, 6, 15], [1, 0, 7, 3, 8, 12, 5, 9, 4, 6, 15, 13, 10, 14, 11, 2], [2, 9, 0, 4, 13, 15, 8, 12, 3, 1, 14, 7, 11, 5, 10, 6], [10, 5, 1, 13, 0, 6, 14, 15, 11, 4, 8, 3, 7, 2, 12, 9]], [[3, 10, 0, 4, 5, 11, 2, 6, 9, 15, 1, 7, 14, 12, 8, 13], [4, 10, 8, 12, 15, 14, 9, 3, 0, 11, 5, 1, 13, 6, 7, 2], [0, 5, 7, 11, 13, 12, 9, 14, 3, 1, 4, 10, 15, 6, 8, 2], [14, 5, 4, 0, 13, 10, 2, 15, 12, 8, 11, 7, 1, 6, 9, 3], [15, 5, 10, 4, 12, 6, 9, 2, 11, 13, 0, 8, 3, 1, 14, 7], [4, 9, 11, 12, 15, 6, 3, 8, 10, 2, 14, 1, 13, 7, 5, 0], [3, 10, 14, 2, 4, 8, 5, 7, 11, 12, 9, 15, 13, 1, 0, 6], [6, 2, 13, 9, 7, 14, 3, 12, 8, 5, 1, 15, 0, 10, 11, 4]], [[11, 4, 7, 1, 14, 3, 6, 0, 2, 13, 8, 9, 5, 15, 12, 10], [3, 6, 8, 1, 9, 7, 12, 15, 13, 2, 14, 11, 5, 0, 10, 4], [9, 12, 10, 2, 7, 8, 6, 1, 3, 0, 4, 14, 5, 13, 11, 15], [11, 4, 7, 0, 14, 8, 2, 3, 12, 9, 15, 1, 10, 5, 6, 13], [15, 10, 8, 13, 11, 0, 7, 5, 6, 14, 12, 1, 4, 3, 9, 2], [1, 10, 2, 9, 5, 8, 13, 14, 4, 7, 6, 11, 3, 15, 12, 0], [4, 14, 8, 5, 9, 2, 3, 11, 13, 0, 15, 12, 7, 1, 10, 6], [12, 10, 4, 13, 2, 3, 9, 14, 8, 7, 5, 0, 15, 11, 1, 6]], [[6, 15, 2, 4, 5, 0, 3, 11, 9, 7, 12, 10, 1, 8, 13, 14], [1, 6, 10, 13, 4, 8, 9, 15, 3, 0, 14, 2, 12, 11, 7, 5], [5, 1, 12, 13, 8, 3, 4, 10, 15, 0, 7, 11, 2, 6, 14, 9], [5, 6, 9, 1, 15, 11, 7, 2, 14, 12, 3, 8, 4, 0, 10, 13], [3, 5, 6, 1, 13, 11, 7, 10, 2, 12, 15, 8, 4, 14, 0, 9], [6, 3, 8, 7, 4, 15, 1, 10, 5, 12, 9, 13, 14, 2, 11, 0], [7, 15, 11, 12, 13, 8, 6, 3, 5, 10, 2, 14, 1, 0, 9, 4], [15, 0, 3, 8, 9, 14, 10, 11, 1, 6, 4, 5, 13, 2, 12, 7]], [[7, 10, 15, 9, 5, 4, 14, 11, 13, 3, 0, 1, 6, 12, 8, 2], [12, 7, 9, 13, 14, 1, 11, 4, 8, 5, 3, 0, 10, 15, 6, 2], [0, 3, 13, 4, 5, 9, 7, 2, 14, 6, 11, 12, 8, 10, 15, 1], [8, 2, 4, 1, 11, 15, 0, 5, 13, 10, 3, 9, 14, 6, 12, 7], [13, 14, 5, 15, 4, 2, 8, 1, 9, 3, 6, 12, 10, 11, 7, 0], [2, 15, 8, 12, 11, 3, 4, 14, 0, 5, 1, 6, 13, 9, 7, 10], [13, 10, 8, 3, 15, 9, 0, 5, 6, 12, 11, 1, 14, 2, 7, 4], [1, 9, 15, 0, 2, 13, 7, 12, 3, 6, 4, 10, 8, 5, 14, 11]], [[11, 6, 4, 9, 7, 2, 1, 13, 14, 15, 12, 10, 5, 0, 8, 3], [6, 1, 7, 11, 0, 9, 14, 3, 15, 10, 2, 4, 8, 5, 13, 12], [15, 9, 0, 10, 6, 11, 3, 14, 7, 1, 12, 5, 8, 13, 2, 4], [11, 1, 10, 13, 3, 7, 9, 0, 8, 14, 2, 15, 6, 4, 5, 12], [13, 12, 9, 15, 1, 8, 5, 10, 3, 11, 0, 14, 2, 4, 7, 6], [11, 14, 0, 12, 2, 8, 10, 3, 13, 7, 5, 4, 9, 1, 6, 15], [4, 6, 2, 10, 11, 9, 3, 7, 8, 0, 1, 14, 13, 5, 12, 15], [6, 4, 8, 13, 2, 10, 12, 14, 7, 1, 9, 11, 0, 15, 3, 5]], [[9, 8, 2, 5, 1, 6, 0, 13, 7, 15, 3, 12, 4, 10, 11, 14], [4, 1, 2, 15, 7, 11, 10, 5, 12, 9, 6, 8, 3, 14, 13, 0], [7, 10, 9, 0, 15, 12, 3, 13, 14, 5, 11, 4, 2, 1, 6, 8], [0, 11, 10, 6, 8, 5, 13, 1, 9, 12, 4, 15, 7, 14, 2, 3], [13, 7, 5, 3, 8, 9, 4, 12, 11, 14, 0, 10, 15, 2, 6, 1], [12, 15, 14, 3, 10, 11, 5, 8, 0, 6, 4, 13, 1, 9, 2, 7], [2, 3, 12, 1, 13, 5, 9, 4, 6, 11, 15, 14, 7, 10, 8, 0], [3, 6, 9, 5, 7, 12, 10, 11, 13, 14, 8, 15, 1, 2, 0, 4]], [[12, 1, 3, 6, 14, 5, 13, 7, 0, 4, 9, 2, 8, 10, 11, 15], [15, 14, 11, 13, 0, 5, 2, 4, 1, 3, 8, 12, 9, 7, 10, 6], [13, 8, 9, 3, 2, 11, 5, 4, 14, 15, 10, 1, 0, 12, 6, 7], [7, 2, 6, 4, 13, 9, 0, 1, 3, 15, 14, 10, 5, 11, 8, 12], [11, 12, 7, 14, 15, 4, 1, 5, 13, 2, 6, 0, 10, 3, 8, 9], [8, 11, 6, 2, 5, 14, 13, 4, 10, 1, 12, 3, 7, 9, 15, 0], [8, 5, 2, 0, 15, 3, 1, 4, 7, 11, 14, 10, 6, 12, 13, 9], [12, 4, 9, 1, 10, 14, 15, 6, 2, 13, 11, 3, 0, 7, 8, 5]], [[4, 3, 10, 2, 0, 9, 13, 14, 5, 11, 7, 1, 15, 6, 8, 12], [5, 9, 1, 0, 10, 7, 3, 15, 4, 12, 2, 13, 11, 8, 6, 14], [4, 9, 6, 7, 10, 14, 0, 8, 13, 2, 15, 3, 5, 1, 11, 12], [4, 3, 5, 8, 10, 0, 1, 14, 2, 7, 15, 13, 11, 12, 6, 9], [13, 1, 2, 8, 6, 9, 3, 7, 11, 5, 14, 15, 0, 10, 4, 12], [14, 0, 13, 8, 7, 15, 5, 2, 10, 1, 11, 12, 4, 3, 6, 9], [13, 15, 0, 3, 4, 6, 12, 10, 14, 1, 2, 5, 11, 7, 9, 8], [3, 8, 1, 2, 14, 4, 0, 6, 15, 12, 13, 7, 11, 10, 9, 5]]]
pin = [61, 24, 29, 31, 63, 47, 43, 57, 55, 7, 30, 28, 51, 22, 49, 0, 2, 37, 23, 8, 27, 46, 6, 25, 18, 54, 17, 42, 45, 1, 60, 41, 11, 56, 40, 44, 50, 48, 36, 5, 4, 53, 39, 14, 9, 32, 21, 35, 3, 33, 15, 62, 20, 26, 58, 59, 19, 38, 16, 13, 12, 34, 10, 52]
pout = [13, 35, 61, 40, 7, 0, 15, 17, 39, 31, 55, 32, 21, 11, 29, 14, 28, 16, 52, 57, 54, 50, 8, 27, 41, 9, 4, 38, 53, 24, 44, 22, 48, 18, 49, 12, 3, 5, 51, 46, 36, 23, 43, 63, 37, 34, 58, 59, 33, 42, 10, 1, 6, 2, 60, 20, 45, 56, 25, 26, 47, 19, 62, 30]
p = [[2, 3, 4, 5, 6, 7, 0, 1], [2, 3, 4, 5, 6, 7, 0, 1], [2, 3, 4, 5, 6, 7, 0, 1], [6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [2, 3, 4, 5, 6, 7, 0, 1], [6, 7, 0, 1, 2, 3, 4, 5], [2, 3, 4, 5, 6, 7, 0, 1], [6, 7, 0, 1, 2, 3, 4, 5]]
m = "Now, now my good man, this is no time to be making enemies; said Voltaire on his deathbed in response to a priest asking him that he renounce the devil."
IV = "FFFFFFFFFF100000"
"""; None
# +
# TEST ANSWER
# ct = "c9dcef337eb02270467c6e3b54239f24de07fdc7dcb6cb642c9997eb0b392e8d586b389d87b848fd79176c5c4dfe2f794e413cd6c3b4671f4b54741162123d325cd1b7b52d037e5bc349a557b8a1aadc241511098c62a0ebdb2a468e03443514c1b2de27903d66c81775498e3c10a2c6aaaab3fb428c3fce29c404d45c4bbc17eb735d9191655cf5ae13d7ac4a310ebb6aa4878dd5ee49b9"
# -
# REAL PARAMETERS
s = [[[11, 1, 2, 0, 9, 10, 12, 8, 14, 6, 7, 15, 13, 5, 4, 3], [9, 10, 11, 15, 13, 12, 8, 1, 4, 3, 5, 14, 2, 7, 6, 0], [13, 6, 0, 1, 14, 9, 10, 2, 15, 3, 4, 8, 11, 12, 7, 5], [13, 1, 10, 6, 8, 3, 12, 5, 11, 2, 0, 15, 4, 7, 9, 14], [15, 14, 6, 0, 12, 7, 2, 4, 11, 13, 1, 9, 8, 3, 10, 5], [3, 8, 6, 12, 4, 7, 13, 5, 1, 11, 10, 15, 0, 2, 9, 14], [12, 1, 5, 0, 11, 4, 15, 6, 2, 3, 13, 9, 8, 10, 14, 7], [7, 0, 3, 11, 14, 2, 10, 4, 5, 15, 13, 9, 12, 8, 1, 6]], [[1, 8, 11, 3, 5, 14, 9, 12, 4, 10, 15, 6, 2, 7, 0, 13], [3, 5, 6, 2, 15, 4, 7, 11, 0, 14, 9, 13, 8, 10, 1, 12], [8, 13, 15, 4, 10, 2, 7, 5, 14, 0, 11, 12, 6, 3, 1, 9], [14, 5, 15, 0, 4, 2, 1, 9, 12, 3, 13, 6, 8, 10, 7, 11], [6, 0, 12, 3, 2, 7, 14, 5, 8, 11, 13, 15, 4, 10, 1, 9], [7, 13, 6, 5, 3, 12, 10, 4, 0, 8, 14, 1, 2, 11, 9, 15], [11, 12, 9, 3, 7, 15, 4, 8, 5, 2, 10, 0, 13, 14, 6, 1], [2, 13, 15, 10, 8, 3, 6, 7, 9, 5, 1, 0, 4, 12, 11, 14]], [[5, 3, 11, 14, 8, 6, 7, 13, 15, 0, 1, 2, 10, 12, 4, 9], [15, 9, 10, 6, 8, 14, 13, 0, 4, 12, 11, 1, 3, 2, 7, 5], [8, 3, 4, 11, 9, 13, 14, 1, 0, 6, 7, 15, 10, 5, 12, 2], [0, 8, 1, 11, 6, 7, 3, 12, 14, 10, 5, 4, 9, 15, 2, 13], [12, 10, 9, 14, 0, 11, 6, 13, 7, 5, 1, 2, 8, 4, 15, 3], [3, 12, 10, 13, 8, 15, 14, 4, 6, 7, 0, 5, 11, 9, 2, 1], [11, 15, 13, 8, 14, 5, 10, 9, 0, 7, 4, 3, 2, 1, 6, 12], [5, 15, 4, 3, 10, 9, 14, 0, 11, 6, 12, 2, 8, 13, 1, 7]], [[2, 11, 9, 6, 10, 8, 15, 14, 12, 7, 0, 1, 3, 13, 5, 4], [0, 8, 14, 5, 13, 15, 12, 6, 11, 7, 3, 1, 2, 9, 10, 4], [6, 11, 8, 12, 1, 5, 3, 2, 10, 9, 15, 13, 0, 7, 14, 4], [14, 6, 2, 5, 8, 11, 9, 10, 13, 1, 3, 0, 12, 15, 7, 4], [6, 15, 10, 2, 8, 9, 14, 12, 3, 0, 13, 1, 11, 5, 7, 4], [7, 4, 3, 9, 11, 12, 10, 0, 14, 8, 15, 6, 2, 1, 13, 5], [9, 0, 11, 12, 15, 3, 5, 10, 14, 13, 4, 8, 6, 7, 2, 1], [3, 10, 4, 12, 2, 15, 7, 6, 9, 14, 13, 1, 8, 11, 0, 5]], [[2, 6, 1, 12, 9, 5, 13, 14, 8, 10, 3, 11, 4, 15, 7, 0], [4, 0, 9, 2, 5, 13, 8, 12, 3, 10, 6, 15, 14, 11, 7, 1], [1, 5, 4, 2, 7, 12, 6, 13, 3, 0, 11, 10, 15, 9, 14, 8], [4, 13, 8, 5, 11, 1, 2, 12, 9, 6, 10, 7, 15, 0, 3, 14], [0, 9, 13, 5, 3, 11, 6, 12, 8, 14, 7, 1, 10, 2, 15, 4], [12, 3, 8, 4, 0, 15, 10, 9, 5, 2, 7, 14, 13, 1, 6, 11], [2, 12, 13, 10, 14, 8, 4, 15, 9, 1, 6, 11, 0, 5, 7, 3], [10, 6, 5, 14, 11, 7, 15, 1, 9, 3, 12, 4, 2, 0, 13, 8]], [[12, 2, 4, 13, 7, 0, 5, 11, 3, 9, 10, 14, 1, 6, 8, 15], [1, 5, 6, 10, 12, 4, 3, 14, 9, 13, 15, 11, 0, 8, 2, 7], [1, 0, 13, 14, 12, 15, 11, 7, 6, 10, 2, 9, 4, 5, 3, 8], [7, 15, 11, 10, 0, 6, 5, 4, 13, 8, 2, 3, 1, 14, 12, 9], [15, 5, 9, 11, 14, 4, 1, 0, 6, 3, 2, 10, 8, 12, 7, 13], [2, 11, 15, 6, 12, 0, 3, 10, 5, 1, 13, 7, 8, 9, 14, 4], [10, 1, 2, 9, 5, 14, 13, 15, 12, 8, 3, 4, 6, 0, 11, 7], [14, 10, 7, 3, 5, 15, 13, 11, 9, 0, 8, 2, 12, 6, 1, 4]], [[9, 10, 14, 11, 13, 5, 1, 15, 8, 7, 12, 2, 4, 0, 6, 3], [1, 2, 9, 5, 10, 6, 8, 12, 0, 4, 14, 3, 13, 7, 15, 11], [13, 14, 7, 3, 6, 12, 4, 8, 10, 2, 1, 5, 0, 9, 11, 15], [14, 6, 5, 13, 3, 0, 7, 11, 2, 1, 4, 10, 15, 8, 9, 12], [7, 8, 6, 5, 4, 0, 14, 10, 2, 1, 12, 15, 3, 9, 11, 13], [15, 11, 4, 6, 14, 2, 9, 8, 7, 5, 10, 13, 1, 12, 0, 3], [4, 14, 2, 1, 5, 9, 6, 0, 12, 10, 11, 13, 3, 15, 8, 7], [10, 3, 12, 14, 6, 5, 2, 0, 7, 13, 1, 15, 4, 11, 8, 9]], [[3, 6, 8, 14, 12, 9, 10, 13, 4, 2, 15, 1, 7, 0, 11, 5], [4, 15, 2, 5, 8, 14, 11, 7, 6, 12, 9, 1, 10, 13, 3, 0], [3, 9, 11, 10, 7, 15, 2, 5, 14, 8, 6, 4, 1, 0, 12, 13], [14, 9, 7, 2, 0, 4, 10, 5, 6, 15, 12, 1, 3, 13, 11, 8], [2, 5, 10, 12, 3, 13, 4, 7, 15, 1, 0, 14, 9, 8, 11, 6], [13, 8, 5, 12, 1, 6, 11, 0, 15, 4, 10, 3, 9, 14, 7, 2], [7, 15, 1, 9, 3, 13, 6, 12, 0, 11, 10, 8, 4, 5, 2, 14], [2, 5, 14, 8, 1, 4, 13, 10, 9, 0, 15, 6, 11, 7, 12, 3]], [[0, 3, 4, 11, 8, 9, 6, 10, 12, 7, 14, 1, 15, 13, 2, 5], [3, 0, 13, 1, 4, 12, 5, 10, 8, 11, 7, 9, 6, 15, 2, 14], [9, 5, 8, 10, 12, 13, 7, 11, 0, 6, 14, 2, 3, 15, 1, 4], [8, 5, 15, 0, 9, 13, 10, 6, 11, 14, 3, 4, 2, 1, 7, 12], [3, 6, 9, 4, 10, 1, 14, 15, 2, 0, 8, 11, 13, 7, 5, 12], [2, 4, 8, 11, 13, 5, 6, 0, 3, 12, 7, 1, 9, 14, 10, 15], [10, 5, 7, 9, 4, 15, 6, 1, 8, 12, 14, 2, 11, 0, 13, 3], [3, 8, 6, 15, 1, 7, 2, 10, 13, 14, 12, 4, 0, 5, 11, 9]]]
pin = [32, 4, 34, 51, 8, 29, 47, 10, 43, 48, 33, 19, 0, 52, 55, 16, 56, 25, 23, 14, 26, 27, 6, 15, 36, 17, 61, 20, 63, 11, 13, 49, 12, 5, 24, 50, 38, 30, 62, 2, 60, 21, 7, 1, 40, 22, 28, 31, 35, 41, 9, 45, 58, 53, 42, 18, 39, 57, 37, 3, 54, 59, 44, 46]
pout = [50, 33, 49, 54, 4, 60, 22, 47, 59, 46, 3, 8, 57, 35, 58, 45, 52, 18, 23, 6, 13, 53, 0, 26, 44, 37, 27, 10, 16, 42, 41, 1, 56, 2, 48, 31, 24, 7, 14, 17, 51, 30, 63, 20, 19, 55, 34, 5, 21, 61, 39, 12, 9, 62, 29, 32, 25, 36, 38, 11, 15, 43, 28, 40]
p = [[6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [2, 3, 4, 5, 6, 7, 0, 1], [6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [6, 7, 0, 1, 2, 3, 4, 5], [2, 3, 4, 5, 6, 7, 0, 1]]
m = "Now, now my good man, this is no time to be making enemies; said Voltaire on his deathbed in response to a priest asking him that he renounce the devil."
IV = "FFFFFFFFFF247565"
# +
# A few helper methods
def string_to_binary(s):
return map(int, ''.join([bin(ord(i)).lstrip('0b').rjust(8,'0') for i in s]))
def nibble_to_int(n):
return int(''.join([str(x) for x in n]), 2)
def hex_to_binary(h):
return bin(int(h, 16))[2:]
def binary_to_hex(b):
h = hex(int(''.join(b), 2))[2:]
if h[-1] == 'L':
return h[:-1]
else:
return h
# decode_binary_string will decode a binary string using a given encoding [FOUND ON STACKOVERFLOW]
def decode_binary_string(s, encoding='utf-8'):
byte_string = ''.join(chr(int(s[i*8:i*8+8],2)) for i in range(len(s)//8))
return byte_string.decode(encoding)
def xor(a, b):
c = []
for i in range(len(a)):
if (str(a[i]) == str(b[i])):
c.append('0')
else:
c.append('1')
return c
# -
def RF(i, R):
b = R
B = []
C = [[] for _ in range(8)]
for j in range(8):
B.append(b[4*j:4*j+4])
# B[j] = s[i][j](B[j])
B_int = nibble_to_int(B[j])
s_bits = '{0:04b}'.format(s[i][j][B_int])
B[j] = list(s_bits)
# C[Pi[j]] = B[j]
permutation = p[i][j]
C[permutation] = B[j]
# c[4*j:4*j+4] = C[j]
# Z = c
Z = [item for sublist in C for item in sublist]
return Z
def Enc(X):
x = X
y = ['' for b in x]
for u in range(64):
y[pin[u]] = x[u]
R = []
L = []
L.append(y[0:32])
R.append(y[32:64])
for i in range(9):
L.append(R[i])
feistel = RF(i, R[i])
R.append(xor(L[i], feistel))
y = R[9] + L[9]
z = ['' for b in y]
for u in range(64):
z[pout[u]] = y[u]
Z = z
return z
def CBC(message):
message_binary = string_to_binary(message)
enc_out = hex_to_binary(IV)
output = []
for i in range(len(message_binary) / 64):
enc_in = xor(enc_out, message_binary[64*i:64*(i + 1)])
enc_out = Enc(enc_in)
output.append(enc_out)
output_bin = [item for sublist in output for item in sublist]
return binary_to_hex(output_bin)
encryption = CBC(m)
print encryption
| src/homework04/e04-blockcipher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import gpplot as gpp
from poola import core as pool
import anchors
import core_functions as fns
gpp.set_aesthetics(palette='Set2')
# +
def run_guide_residuals(lfc_df, paired_lfc_cols=[]):
'''
Calls get_guide_residuals function from anchors package to calculate guide-level residual z-scores
Inputs:
1. lfc_df: data frame with log-fold changes (relative to pDNA)
2. paired_lfc_cols: grouped list of initial populations and corresponding resistant populations
'''
lfc_df = lfc_df.drop_duplicates()
if not paired_lfc_cols:
paired_lfc_cols = fns.pair_cols(lfc_df)[1] #get lfc pairs
modified = []
unperturbed = []
#reference_df: column1 = modifier condition, column2 = unperturbed column
ref_df = pd.DataFrame(columns=['modified', 'unperturbed'])
row = 0 #row index for reference df
for pair in paired_lfc_cols:
#number of resistant pops in pair = len(pair)-1
res_idx = 1
#if multiple resistant populations, iterate
while res_idx < len(pair):
ref_df.loc[row, 'modified'] = pair[res_idx]
ref_df.loc[row, 'unperturbed'] = pair[0]
res_idx +=1
row +=1
print(ref_df)
#input lfc_df, reference_df
#guide-level
residuals_lfcs, all_model_info, model_fit_plots = anchors.get_guide_residuals(lfc_df, ref_df)
return residuals_lfcs, all_model_info, model_fit_plots
def select_top_ranks(df, rank = 5): #pick top ranks from each column of df with ranks, rank = top rank threshold (e.g. 5 if top 5)
'''
Inputs:
1. df: Dataframe with columns "Gene Symbol" and data used to rank
2. rank: top number of rows to select
Outputs:
1. final_top_rank_df: Data frame with top ranked rows
'''
rank_cols = df.columns.to_list()[1:]
prev_top_rank_rows = pd.DataFrame(columns = df.columns)
final_top_rank_df = pd.DataFrame() #for final list
for col in rank_cols:
#top_rank_rows = df.copy().loc[lambda df: df[col] <= rank, :] #pick rows with rank <= 5
top_rank_rows = df.copy().nlargest(rank, col)
top_rank_df = pd.concat([prev_top_rank_rows, top_rank_rows]) #concat with rows selected from previous column
prev_top_rank_rows = top_rank_df #set combined list as previous
final_top_rank_df = prev_top_rank_rows.drop_duplicates(subset = ['Gene Symbol']) #drop duplicate gene rows
return final_top_rank_df
# -
# ## Data summary
#
# +
reads_nopDNA = pd.read_csv('../../../Data/Reads/Goujon/Caco2/SecondaryScreens/counts-JD_GPP2845_Goujon_Plate3_CP1658.txt', sep='\t')
reads_nopDNA = reads_nopDNA.copy().drop('Construct IDs', axis=1)
CP1658_cols = ['Construct Barcode']+[col for col in reads_nopDNA.columns if 'CP1658' in col]
reads_nopDNA_CP1658 = reads_nopDNA[CP1658_cols]
pDNA_reads_all = pd.read_csv('../../../Data/Reads/Goujon/Calu3/Secondary_Library/counts-LS20210325_A02_AAHG04_RDA118_G0_CP1658_M-AM33.txt', sep='\t')
pDNA_reads = pDNA_reads_all[['Construct Barcode','A02_AAHG04_RDA118_G0_CP1658_M-AM33']].copy()
pDNA_reads = pDNA_reads.rename(columns = {'A02_AAHG04_RDA118_G0_CP1658_M-AM33': 'pDNA'})
pDNA_reads
reads_CP1658 = pd.merge(pDNA_reads, reads_nopDNA_CP1658, how = 'right', on='Construct Barcode')
empty_cols = [col for col in reads_CP1658.columns if 'EMPTY' in col]
reads_CP1658 = reads_CP1658.copy().drop(empty_cols, axis=1)
reads_CP1658
# -
# Gene Annotations
chip = pd.read_csv('../../../Data/Interim/Goujon/Secondary_Library/CP1658_GRCh38_NCBI_strict_gene_20210604.chip', sep='\t')
chip = chip.rename(columns={'Barcode Sequence':'Construct Barcode'})
chip_reads_CP1658 = pd.merge(chip[['Construct Barcode', 'Gene Symbol']], reads_CP1658, on = ['Construct Barcode'], how = 'right')
chip_reads_CP1658
# +
#Calculate lognorm
cols = chip_reads_CP1658.columns[2:].to_list() #reads columns = start at 3rd column
lognorms = fns.get_lognorm(chip_reads_CP1658.dropna(), cols = cols)
col_list = []
for col in lognorms.columns:
if 'intitial'in col:
new_col = col.replace('intitial', 'initial')
col_list.append(new_col)
else:
col_list.append(col)
lognorms.columns = col_list
lognorms
# -
# ## Quality Control
# ### Population Distributions
#Calculate log-fold change relative to pDNA
target_cols = list(lognorms.columns[3:])
pDNA_lfc = fns.calculate_lfc(lognorms,target_cols)
pDNA_lfc
pair1 = list(pDNA_lfc.columns[2:4])
pair2 = list(pDNA_lfc.columns[-2:])
paired_cols = (True, [pair1, pair2])
#Plot population distributions of log-fold changes
fns.lfc_dist_plot(pDNA_lfc, paired_cols=paired_cols, filename = 'Caco2_Cas9SecondaryLibraryKO_Goujon')
# ### Distributions of control sets
fns.control_dist_plot(pDNA_lfc, paired_cols=paired_cols, control_name=['ONE_INTERGENIC_SITE'], filename = 'Caco2_Cas9SecondaryLibraryKO_Goujon')
# ### ROC_AUC
# Essential gene set: Hart et al., 2015
# <br>
# Non-essential gene set: Hart et al., 2014
# +
ess_genes, non_ess_genes = fns.get_gene_sets()
initial_cols = [col for col in pDNA_lfc.columns if 'initial' in col]
tp_genes = ess_genes.loc[:, 'Gene Symbol'].to_list()
fp_genes = non_ess_genes.loc[:, 'Gene Symbol'].to_list()
initial_roc_dict = {}
intial_roc_auc_dict = {}
for col in initial_cols:
roc_auc, roc_df = pool.get_roc_aucs(pDNA_lfc, tp_genes, fp_genes, gene_col = 'Gene Symbol', score_col=col)
initial_roc_dict[col] = roc_df
intial_roc_auc_dict[col] = roc_auc
fig,ax=plt.subplots(figsize=(6,6))
for key, df in initial_roc_dict.items():
roc_auc = intial_roc_auc_dict[key]
ax=sns.lineplot(data=df, x='fpr',y='tpr', ci=None, label = key+',' + str(round(roc_auc,2)))
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title('ROC-AUC')
plt.xlabel('False Positive Rate (non-essential)')
plt.ylabel('True Positive Rate (essential)')
# -
# ## Gene level analysis
# ### Residual z-scores
lfc_df = pDNA_lfc.drop('Gene Symbol', axis = 1)
lfc_df
# run_guide_residuals(lfc_df.drop_duplicates(), cols)
residuals_lfcs, all_model_info, model_fit_plots = run_guide_residuals(lfc_df, paired_lfc_cols=paired_cols[1])
residuals_lfcs
guide_mapping = pool.group_pseudogenes(chip[['Construct Barcode', 'Gene Symbol']], pseudogene_size=10, gene_col='Gene Symbol', control_regex=['ONE_INTERGENIC_SITE'])
guide_mapping
gene_residuals = anchors.get_gene_residuals(residuals_lfcs.drop_duplicates(), guide_mapping)
gene_residuals
# +
gene_residual_sheet = fns.format_gene_residuals(gene_residuals, guide_min = 8, guide_max = 11)
guide_residual_sheet = pd.merge(guide_mapping, residuals_lfcs.drop_duplicates(), on = 'Construct Barcode', how = 'inner')
guide_residual_sheet
# -
with pd.ExcelWriter('../../../Data/Processed/GEO_submission_v2/SecondaryLibrary/Caco2_Cas9SecondaryLibraryKO_Goujon.xlsx') as writer:
gene_residual_sheet.to_excel(writer, sheet_name='Caco2_avg_zscore', index =False)
reads_CP1658.to_excel(writer, sheet_name='Caco2_genomewide_reads', index =False)
guide_mapping.to_excel(writer, sheet_name='Caco2_guide_mapping', index =False)
screen1_df = gene_residuals[gene_residuals['condition'].str.contains('#1')]
screen2_df = gene_residuals[gene_residuals['condition'].str.contains('#2')]
zscore_df = pd.merge(screen1_df[['Gene Symbol', 'residual_zscore']], screen2_df[['Gene Symbol', 'residual_zscore']], on = 'Gene Symbol', how = 'outer', suffixes = ['_screen#1', '_screen#2'])
zscore_df
# +
# Screen 2 vs Screen 1
fig, ax = plt.subplots(figsize = (2, 2))
ax = gpp.point_densityplot(zscore_df, 'residual_zscore_screen#1', 'residual_zscore_screen#2', s=6)
ax = gpp.add_correlation(zscore_df, 'residual_zscore_screen#1', 'residual_zscore_screen#2', fontsize=7)
top_ranked_screen1 = zscore_df.nsmallest(10, 'residual_zscore_screen#1')
top_ranked_screen2 = zscore_df.nsmallest(10, 'residual_zscore_screen#2')
bottom_ranked_screen1 = zscore_df.nlargest(10, 'residual_zscore_screen#1')
bottom_ranked_screen2 = zscore_df.nlargest(10, 'residual_zscore_screen#2')
screen1_ranked = pd.concat([top_ranked_screen1, bottom_ranked_screen1])
screen2_ranked = pd.concat([top_ranked_screen2, bottom_ranked_screen2])
# Annotate common hits
common_ranked = pd.merge(screen1_ranked, screen2_ranked, on = ['Gene Symbol', 'residual_zscore_screen#1', 'residual_zscore_screen#2'], how = 'inner')
common_ranked
sns.scatterplot(data=common_ranked, x='residual_zscore_screen#1', y='residual_zscore_screen#2', color = sns.color_palette('Set2')[0], edgecolor=None, s=6)
texts= []
for j, row in common_ranked.iterrows():
texts.append(ax.text(row['residual_zscore_screen#1']+0.25, row['residual_zscore_screen#2'], row['Gene Symbol'], fontsize=7,
color = 'black'))
# ensures text labels are non-overlapping
# adjust_text(texts)
plt.title('Caco-2 Cas9 KO Secondary Library Screen', fontsize=7)
plt.xlabel('Screen #1', fontsize=7)
plt.ylabel('Screen #2', fontsize=7)
plt.xticks(fontsize=7)
plt.yticks(fontsize=7)
sns.despine()
gpp.savefig('../../../Figures/Scatterplots/Caco2_Cas9KO_Secondary_Screen1vs2_scatterplot.pdf', dpi=300)
# -
with pd.ExcelWriter('../../../Data/Processed/Individual_screens_v2/Caco2_Cas9SecondaryLibraryKO_Goujon_indiv_screens.xlsx') as writer:
zscore_df.to_excel(writer, sheet_name='indiv_screen_zscore', index =False)
gene_residuals.to_excel(writer, sheet_name='condition_genomewide_zscore', index =False)
guide_residual_sheet.to_excel(writer, sheet_name='guide-level_zscore', index =False)
# ## Primary vs Secondary Screen
# +
Caco2_primary = pd.read_excel('../../../Data/Processed/GEO_submission_v2/Caco2_Brunello_Goujon_v2.xlsx')
Caco2_primaryvssecondary = pd.merge(Caco2_primary, gene_residual_sheet, on = 'Gene Symbol', how = 'inner', suffixes=['_primary', '_secondary'])
Caco2_primaryvssecondary
fig, ax = plt.subplots(figsize=(2,2))
ax = gpp.point_densityplot(Caco2_primaryvssecondary, 'residual_zscore_avg_primary', 'residual_zscore_avg_secondary', s=6)
ax = gpp.add_correlation(Caco2_primaryvssecondary, 'residual_zscore_avg_primary', 'residual_zscore_avg_secondary', fontsize=7)
top_ranked_primary = Caco2_primaryvssecondary.nlargest(10, 'residual_zscore_avg_primary')
top_ranked_secondary = Caco2_primaryvssecondary.nlargest(10, 'residual_zscore_avg_secondary')
bottom_ranked_primary = Caco2_primaryvssecondary.nsmallest(10, 'residual_zscore_avg_primary')
bottom_ranked_secondary = Caco2_primaryvssecondary.nsmallest(10, 'residual_zscore_avg_secondary')
primary_ranked = pd.concat([top_ranked_primary, bottom_ranked_primary])
secondary_ranked = pd.concat([top_ranked_secondary, bottom_ranked_secondary])
# Annotate common hits
common_ranked = pd.merge(primary_ranked, secondary_ranked, on = ['Gene Symbol', 'residual_zscore_avg_primary', 'residual_zscore_avg_secondary'], how = 'inner')
common_ranked
annot_df = pd.concat([common_ranked, top_ranked_secondary.head(5)]).drop_duplicates()
sns.scatterplot(data=annot_df, x='residual_zscore_avg_primary', y='residual_zscore_avg_secondary', color = sns.color_palette('Set2')[0], edgecolor=None, s=6, rasterized=True)
texts= []
for j, row in annot_df.iterrows():
texts.append(ax.text(row['residual_zscore_avg_primary']+0.25, row['residual_zscore_avg_secondary'], row['Gene Symbol'], fontsize=7,
color = 'black'))
# ensures text labels are non-overlapping
# adjust_text(texts)
plt.title('Caco-2 Cas9 KO Primary vs. Secondary Screens', fontsize=7)
plt.xlabel('Primary (Mean z-score)', fontsize=7)
plt.ylabel('Secondary (Mean z-score)', fontsize=7)
plt.xticks(fontsize=7)
plt.yticks(fontsize=7)
sns.despine()
gpp.savefig('../../../Figures/Scatterplots/Caco2_Cas9KO_PrimaryvsSecondary_scatterplot.pdf', dpi=300)
# -
| SecondaryScreens/Goujon_Caco2_Cas9KO_SecondaryScreen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# ### Read Data
# read a small number of rows
df = pd.read_csv("datasets/weather_underground.csv", nrows=2)
df
# take a subset of columns
cols = df.columns.tolist()
cols[:4]
# read selected columns, limited rows, and cast data types
dtype = {'date' : str, 'maxpressurem': int, 'maxdewptm': float, 'maxpressurei': float}
df = pd.read_csv("datasets/weather_underground.csv", nrows=2, usecols=cols[:4], dtype=dtype)
df
# check data type has been properly casted
df.maxdewptm
# ### Data Wrangling
# select subset of columns by data types
df.select_dtypes(include=['object', 'int'])
# map categorical variables
level_map = {'2011-05-01': 'day 1', '2011-05-02': 'day 2'}
df['cat_level'] = df['date'].map(level_map)
df
# +
# map numeric variables
def rule(x):
if x < 7: return 'low'
return 'high'
df['cut_level'] = df.apply(lambda x: rule(x['maxdewptm']), axis = 1)
df
# -
# aggregate columns
df['maximum'] = df[['maxdewptm','maxpressurei']].max(axis =1)
df
df = pd.read_csv("datasets/weather_underground.csv", usecols=cols[:4])
df.head()
# select certain rows
# df_filter is a boolean mask
df_filter = df['date'].isin(['2011-05-04','2011-05-05'])
df[df_filter]
# +
# smallest partition is numbered highest
cut_points = [np.percentile(df['maxdewptm'], i) for i in [25, 50, 75]]
print(cut_points)
df['group'] = 1
for i in range(3):
df['group'] = df['group'] + (df['maxdewptm'] < cut_points[i])
df.head()
# -
# save without storing numeric index
df.to_csv("datasets/weather_underground_subset.csv", index=None)
# #### NULL Handling
# count nulls in each rows
df = pd.DataFrame({ 'id': [1,2,3], 'c1':[0,0,np.nan], 'c2': [np.nan,1,1]})
df = df[['id', 'c1', 'c2']]
df['num_nulls'] = df[['c1', 'c2']].isnull().sum(axis=1)
df.head()
df.shape
# drop rows that contains any null
df.dropna().shape
# drop rows that have nulls in specific columns
df.dropna(subset=['c2']).shape
| preprocessing/pandas_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from astropy.table import Table, Column
import numpy as np
from numpy.linalg import inv
import pandas as pd
import matplotlib.pyplot as plt
import os
import urllib.request
os.chdir("/Users/<NAME>/Documents/Python Scripts")
os.getcwd()
# +
Table1 = np.loadtxt(fname = "/Users/<NAME>/Documents/Python Scripts/Table3.txt")
column_names = ['ID', 'x', 'y','Sigma y', 'Sigma x', 'Rho xy']
Table1 = pd.DataFrame(Table1, columns=column_names)
# +
Matrix_Y_Transpose = np.array([Table1['y']])
Matrix_Y = Matrix_Y_Transpose.transpose()
# +
LineOfOnes = np.linspace(1, 1, Table1['ID'].idxmax()+1)
Matrix_A_Transpose = np.array([LineOfOnes,Table1['x'],Table1['x']*Table1['x']])
Matrix_A = Matrix_A_Transpose.transpose()
Covariance_Matrix = np.diag(Table1['Sigma y']*Table1['Sigma y'])
Covariance_Matrix_Inverse = inv(Covariance_Matrix)
# +
MAT_times_CMI = np.matmul(Matrix_A_Transpose,Covariance_Matrix_Inverse)
MAT_CMI_times_MA = np.matmul(MAT_times_CMI,Matrix_A)
MAT_CMI_times_MA_Inverse = inv(MAT_CMI_times_MA)
MAT_CMI_times_MY = np.matmul(MAT_times_CMI,Matrix_Y)
Matrix_X = np.matmul(MAT_CMI_times_MA_Inverse,MAT_CMI_times_MY)
# +
b = Matrix_X[0].item()
m = Matrix_X[1].item()
q = Matrix_X[2].item()
print(q,m,b)
# +
def f(t):
return q*t**2 + t*m +b
t3 = np.arange(0.0, 300.0, 0.1)
t4 = np.arange(0.0, 300.0, 0.02)
# -
error = Table1['Sigma y']
plt.errorbar(Table1['x'], Table1['y'], yerr=error, fmt='ko',capsize=4)
plt.plot(t4, f(t4), 'k')
| Data analysis recipes - Fitting a model to data/Section 1 - Standard practice/Exercise 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lambda Expressions, Map, and Filter
#
# Now its time to quickly learn about two built in functions, filter and map. Once we learn about how these operate, we can learn about the lambda expression, which will come in handy when you begin to develop your skills further!
# ## map function
#
# The **map** function allows you to "map" a function to an iterable object. That is to say you can quickly call the same function to every item in an iterable, such as a list. For example:
def square(num):
return num**2
my_nums = [1,2,3,4,5]
map(square,my_nums)
# To get the results, either iterate through map()
# or just cast to a list
list(map(square,my_nums))
# The functions can also be more complex
def splicer(mystring):
if len(mystring) % 2 == 0:
return 'even'
else:
return mystring[0]
mynames = ['John','Cindy','Sarah','Kelly','Mike']
list(map(splicer,mynames))
# ## filter function
#
# The filter function returns an iterator yielding those items of iterable for which function(item)
# is true. Meaning you need to filter by a function that returns either True or False. Then passing that into filter (along with your iterable) and you will get back only the results that would return True when passed to the function.
def check_even(num):
return num % 2 == 0
nums = [0,1,2,3,4,5,6,7,8,9,10]
filter(check_even,nums)
list(filter(check_even,nums))
# ## lambda expression
#
# One of Pythons most useful (and for beginners, confusing) tools is the lambda expression. lambda expressions allow us to create "anonymous" functions. This basically means we can quickly make ad-hoc functions without needing to properly define a function using def.
#
# Function objects returned by running lambda expressions work exactly the same as those created and assigned by defs. There is key difference that makes lambda useful in specialized roles:
#
# **lambda's body is a single expression, not a block of statements.**
#
# * The lambda's body is similar to what we would put in a def body's return statement. We simply type the result as an expression instead of explicitly returning it. Because it is limited to an expression, a lambda is less general that a def. We can only squeeze design, to limit program nesting. lambda is designed for coding simple functions, and def handles the larger tasks.
# Lets slowly break down a lambda expression by deconstructing a function:
def square(num):
result = num**2
return result
square(2)
# We could simplify it:
def square(num):
return num**2
square(2)
# We could actually even write this all on one line.
def square(num): return num**2
square(2)
# This is the form a function that a lambda expression intends to replicate. A lambda expression can then be written as:
lambda num: num ** 2
# You wouldn't usually assign a name to a lambda expression, this is just for demonstration!
square = lambda num: num **2
square(2)
# So why would use this? Many function calls need a function passed in, such as map and filter. Often you only need to use the function you are passing in once, so instead of formally defining it, you just use the lambda expression. Let's repeat some of the examples from above with a lambda expression
list(map(lambda num: num ** 2, my_nums))
list(filter(lambda n: n % 2 == 0,nums))
# Here are a few more examples, keep in mind the more comples a function is, the harder it is to translate into a lambda expression, meaning sometimes its just easier (and often the only way) to create the def keyword function.
# ** Lambda expression for grabbing the first character of a string: **
lambda s: s[0]
# ** Lambda expression for reversing a string: **
lambda s: s[::-1]
# You can even pass in multiple arguments into a lambda expression. Again, keep in mind that not every function can be translated into a lambda expression.
lambda x,y : x + y
# You will find yourself using lambda expressions often with certain non-built-in libraries, for example the pandas library for data analysis works very well with lambda expressions.
| 03-Methods and Functions/05-Lambda-Expressions-Map-and-Filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 여러분의 정신 건강을 위해, 그냥 프로젝트 옮길 때마다 복붙 해서 쓰는 것을 추천 드립니다.
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset
from torchvision.datasets import FashionMNIST
from torchvision import transforms
from torch.utils.data import DataLoader
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import random
# ## XOR Gate?
# XOR Gate는 일반적인 선형 모델로는 해결 할 수 없습니다.
# +
x = np.array([
[0, 0, 1, 1],
[0, 1, 0, 1]
])
y = np.array([0, 1, 1, 0])
marker = ['x', 'o']
for i in range(4):
plt.scatter(x[0][i], x[1][i], color='black', marker=marker[y[i]])
plt.title("XOR Gate")
plt.xlim(-0.5, 1.5)
plt.xticks([0, 1])
plt.yticks([0, 1])
plt.ylim(-0.5, 1.5)
plt.show()
# -
# ## Multi-Layer Perceptron
# 하지만, 퍼셉트론을 여러개 이용 한다면, 문제를 해결 할 수 있습니다.
# +
new_x = np.array([x[0] + x[1] < 1.5, x[0] + x[1] > 0.5])
marker = ['x', 'o']
for i in range(4):
plt.scatter(new_x[0][i], new_x[1][i], color='black', marker=marker[y[i]])
plt.title("XOR to new dimension")
plt.xlim(-0.5, 1.5)
plt.xticks([0, 1])
plt.yticks([0, 1])
plt.ylim(-0.5, 1.5)
plt.show()
# -
# ## Pytorch Tensor
# **Pytorch**에서 **Tensor**는 계산의 기본 단위로, **Numpy**의 `ndarray`와 유사 합니다.
#
# `Tensor` 객체를 만드는 여러 가지 방법들을 알려 드리겠습니다.
# - `torch.empty(x, y)`: x * y 사이즈의 요소들의 값이 초기화 되지 않은 행렬 반환.
# - `torch.rand(x, y)`: x * y 사이즈의 요소들이 **0 ~ 1 사이의 랜덤한 값**으로 초기화 된 행렬 반환.
# - `torch.randn(x, y)`: x * y 사이즈의 요소들이 **정규분포 그래프** 상의 랜덤한 값으로 초기화 된 행렬 반환.
# - `torch.zeros(x, y, dtype=type)`: x * y 사이즈의 요소들이 **0으로 초기화** 된 행렬 반환, 요소들은 type에 맞게 초기화 된다.
# - `torch.ones(x, y, dtype=type)`: x * y 사이즈의 요소들이 **1으로 초기화** 된 행렬 반환, 요소들은 type에 맞게 초기화 된다.
# - `torch.tensor(iterable)`: `iterable`한 객체를 `Tensor` 객체로 변환한다.
# - `torch.zeros_like(tensor, dtype=type)`: 파라미터로 들어 간 `Tensor` 객체의 사이즈과 똑같은 행렬을 반환하며, 요소들은 **0으로 초기화** 되어 있다.
# - `torch.ones_like(tensor, dtype=type)`: 파라미터로 들어 간 `Tensor` 객체의 사이즈과 똑같은 행렬을 반환하며, 요소들은 **1으로 초기화** 되어 있다.
# - `torch.randn_like(tensor, dtype=type)`: 파라미터로 들어 간 `Tensor` 객체의 사이즈과 똑같은 행렬을 반환하며, 요소들은 **정규분포 그래프** 상의 랜덤한 값으로 초기화 되어 있다.
# +
empty_tensor = torch.empty(3, 3) # 3 * 3의 빈 행렬 생성
rand_tensor = torch.rand(3, 3) # 3 * 3의 요소들이 0 ~ 1의 랜덤 값으로 초기화된 행렬 생성
randn_tensor = torch.randn(3, 3, dtype=torch.double) # 3 * 3의 요소들이 정규분포 그래프 값으로 초기화된 행렬 생성
zero_tensor = torch.zeros(3, 3, dtype=torch.long) # 3 * 3의 요소들이 0으로 초기화된 행렬 생성
one_tensor = torch.ones(3, 3, dtype=torch.double) # 3 * 3의 요소들이 1으로 초기화된 행렬 생성
iterable_tensor = torch.tensor([1, 2, 3]) # list 객체를 Tensor 객체로 변환
zeros_like_tensor = torch.zeros_like(iterable_tensor, dtype=torch.double) # iterable_tensor와 사이즈가 같은, 요소들이 0으로 초기화된 행렬 생성
ones_like_tensor = torch.ones_like(iterable_tensor, dtype=torch.double) # iterable_tensor와 사이즈가 같은, 요소들이 1으로 초기화된 행렬 생성
randn_like_tensor = torch.randn_like(iterable_tensor, dtype=torch.double) # iterable_tensor와 사이즈가 같은, 요소들이 정규분포 그래프 값으로 초기화된 행렬 생성
print(empty_tensor)
print(rand_tensor)
print(randn_tensor)
print(zero_tensor)
print(one_tensor)
print(iterable_tensor)
print(zeros_like_tensor)
print(ones_like_tensor)
print(randn_like_tensor)
# -
# 일단 기본적인 **연산자**는 다음과 같이 사용 할 수 있습니다. 또한, `Numpy`에서 사용 했던, **차원별 인덱싱**과 **브로드캐스팅**이 가능합니다.
# +
x_tensor = torch.tensor([[1, 2], [3, 4]]) # 2 * 2 행렬 생성
y_tensor = torch.tensor([[5, 6], [7, 8]])
z_tensor = torch.tensor([[[1, 2], [3, 4]], [[1, 2], [3, 4]]])
print(x_tensor)
print(y_tensor)
print(x_tensor + y_tensor) # Index가 일치하는 요소 끼리 덧셈
print(x_tensor - y_tensor) # Index가 일치하는 요소 끼리 뺄셈
print(x_tensor * y_tensor) # Index가 일치하는 요소 끼리 곱셈
print(x_tensor @ y_tensor) # 행렬 곱
print(x_tensor * 3) # x_tensor 각 요소에 3을 곱해줌
print(x_tensor + z_tensor) # 일치하는 요소에 브로드캐스팅
a_tensor = torch.tensor([[1, 2, 3], [4, 5, 6]])
b_tensor = torch.tensor([[1, 2], [3, 4], [5, 6]])
print(a_tensor)
print(b_tensor)
print(a_tensor @ b_tensor) # 2 * 3 행렬과 3 * 2 행렬 곱셈
print(a_tensor[:, 1]) # 각 행의 1번째 열 추출
print(b_tensor[1, :]) # 1번째 행의 모든 열 추출
# -
# **딥러닝 모델**을 설계하다 보면 **행렬 사이즈를 재조정** 해야 하는 경우가 상당히 많습니다. 이를 위한 **몇 가지 함수**를 소개 시켜 드리겠습니다.
# - `Tensor.size()`: `Tensor` 객체의 **사이즈를 반환** 한다.
# - `Tensor.view(size)`: 파라미터로 들어간 사이즈로 `Tensor` 객체의 **사이즈를 변환** 시켜 주며, 파라미터로 -1이 들어갈 시, **행렬의 차원 수를 낮춰** 리사이징 하며, (-1, n)이 들어가면 가장 **하위 차원에서 n개씩 끊어 넣는 방식**으로 리사이징 한다. `numpy`의 `resize`와 방법이 유사하다.
# a_tensor = torch.tensor([[1, 2, 3, 4], # 4 * 4 행렬
# [5, 6, 7, 8],
# [9, 10, 11, 12],
# [13, 14, 15, 16]])
# print(a_tensor)
# print(a_tensor.size())
#
# b_tensor = a_tensor.view(16) # 사이즈가 16 array
# print(b_tensor)
# print(b_tensor.size())
#
# c_tensor = a_tensor.view(-1, 8) # 4 * 4 => 2 * 8
# print(c_tensor) # 6 * 4 일시 3 * 8 으로 리사이징 된다.
# print(c_tensor.size())
#
# d_tensor = a_tensor.view(-1) # 4 * 4 => 16
# print(d_tensor) # 6 * 4 일시 24로 리사이징 된다.
# print(d_tensor.size())
#
# e_tensor = a_tensor.view(8, 2) # 8 * 2 행렬로 사이즈 변환
# print(e_tensor)
# print(e_tensor.size())
# `Pytorch`는 `Numpy`연산을 GPU를 이용하여 더욱 빠르게 하기 위해 탄생했습니다. 그렇기 때문에 `Numpy`와 `Pytorch`를 연동할 수 있도록, `Pytorch`에서는 `API`를 제공합니다.
#
# - `Tensor.numpy()`: `Tensor` 객체를 `numpy.ndarray` 객체로 변환 하여 반환합니다.
# - `torch.from_numpy(ndarray)`: `numpy.ndarray`를 `Tensor` 객체로 변환 하여 반환합니다.
# +
a_matrix = np.array([[1, 2], [3, 4]]) # np.ndarray 객체 할당
b_matrix = torch.from_numpy(a_matrix) # np.ndarray 객체를 이용하여 Tensor 객체 할당
c_matrix = b_matrix.numpy() # Tensor 객체를 이용하여 np.ndarray 객체 할당
print(a_matrix) # np.ndarray
print(b_matrix) # Tensor
print(c_matrix) # np.ndarray
# -
# ## Autograd
# Pytorch에서는 연산을 진행 하면 전 연산에 대한 미분 또한 제공 합니다. 이는 각 `Tensor`에 `requires_grad=True`를 넣어 줌으로써 기능 합니다.
# +
x = torch.ones(2, 2, requires_grad=True)
print(x)
x = torch.ones(2, 2)
x.requires_grad_(True)
print(x)
# -
# **역전파 (Back Propagation)**를 시키는 방법은 생각보다 간단합니다. 최종 연산 된 `Tensor` 객체의 `backward()` 함수만 호출해 주면 됩니다.
#
# `out = z/4 = y * y * 3/4 = (x + 2)^2 * 3 / 4` 이고, 이를 **x**에 대해서 미분하면
# `dout/dx = 3(2x + 4) / 4` 입니다. **x = 1**일 경우 **18 / 4 = 4.5** 가 정답이므로, 잘 작동 하고 있다고 볼 수 있습니다.
#
# 주의: **out**에 한 연산은 **스칼라 연산**입니다. **행렬 곱**등을 사용하지 않았습니다.
# +
x = torch.ones(2, 2, requires_grad=True) # 최초 Tensor 객체
y = x + 2
z = y * y * 3
out = z.mean() # 미분 대상
out.backward() # out.backward(torch.tensor(1.)) 과 동일
print(x.grad) # dout/dx
# -
# 행렬 미분은 다음과 같습니다.
# +
x = torch.ones(2, 3, requires_grad=True) # 최초 Tensor 객체
y = torch.ones(3, 6)
z = x @ y # x @ y 행렬 곱, 더 이상 스칼라 값이 아니다.
out = z * 2
print(out)
out.backward(torch.ones(2, 6) * 2) # dout/dz
print(x.grad) # dout/dx
# -
# 역전파 연산이 필요 없을 때는 다음 두 가지 방법 중 하나를 선택합니다.
#
# 1. `torch.no_grad()` 를 이용하여 연산하기.
# 2. `Tensor.detech()` 를 이용하여, `autograd` 연산을 하지 않은 `Tensor` 복사하기.
# +
x = torch.ones(2, 2, requires_grad=True)
print(x.requires_grad)
print((x ** 2).requires_grad)
with torch.no_grad(): # autograd 연산 생략
print((x ** 2).requires_grad)
print('')
y = x.detach() # autograd가 없이 내용물만 복사.
print(y.requires_grad)
print(x.eq(y).all()) # x와 y의 내용물은 같다.
# -
# ## Fashion MNIST
# 우리가 오늘 해 볼 것은, **DNN을 이용한 Fashion MNIST Dataset** 분류 입니다. 여기서 말하는 **Fashion MNIST**는 입력이 28x28 크기의 행렬로 이루어져 있고 출력은 10개의 분류로 나타 냅니다. 열 개의 분류는 다음과 같습니다. (T-shirt/top, Trouser, Pullover, Dress, Coat, Sandal, Shirt, Sneaker, Bag, Ankle boot)
# 첫 번째로, 데이터를 불러 오기 전, 데이터를 전처리 하기 위한 **파이프 라인**을 구성 해 보겠습니다. 아래 코드는 **Input Data**를 **Tensor**로 만들고, 이를 **Z-Score Normalization**을 하는 모습입니다. `transforms.Compose`를 이용하여 파이프라인을 구성하고, `transforms.ToTensor()`를 통해 `Tensor`로의 변환, `transforms.Normalize`를 통해 Z-Score Normalization을 실시 합니다.
RANDOM_SEED = 123
DEVICE = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# +
custom_train_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=(0.5,), std=(0.5,))
])
custom_test_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=(0.5,), std=(0.5,))
])
# -
# 두 번째로는 만들어진 **파이프라인**에, **Fashion MNIST** 데이터를 넣어 전처리를 해 보겠습니다. 이는 `Dataset`을 불러 올 때 `transform` 파라미터에 값을 넣어 줌으로써 가능 합니다.
#
# `DataLoader`를 통해 **데이터 셋**을 **미니 배치**로 분류하고, **데이터를 섞어 줄 수** 있습니다.
# +
BATCH_SIZE = 64
train_dataset = FashionMNIST(".", train=True, download=True, transform=custom_train_transform)
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
shuffle=True,
drop_last=True,
num_workers=2)
test_dataset = FashionMNIST(".", train=False, download=True, transform=custom_test_transform)
test_loader = DataLoader(dataset=test_dataset,
batch_size=BATCH_SIZE,
shuffle=False,
num_workers=2)
# -
# 아래 창은 데이터가 잘 다운로드 되었는지 확인 하는 코드입니다.
for batch_idx, (x, y) in enumerate(train_loader):
print(' | Batch size:', y.size()[0])
x = x.to(DEVICE)
y = y.to(DEVICE)
print("X shape: ", x.shape)
print("Y shape: ", y.shape)
print('break minibatch for-loop')
break
# ## Deep Neural Network
# 이제 **DNN 모델**을 구현 해 볼 시간입니다. 한 번 구현을 해 볼까요?
#
# 일단 코드 설명을 드리자면, 가장 먼저 해야할 일은 `torch.nn.Module`을 상속 받은 class를 만드는 것입니다. 그 다음, 생성자와 `forward` 함수를 구현 해 주면 됩니다.
#
# 먼저 생성자 함수는 `super(클래스명, self).__init__()`을 통해, 모델 초기화를 해 주어야 하며, 그 다음으로는 `torch.nn.Linear`로 층을 쌓아 줍니다. `torch.nn.Linear`는 파라미터로 (input_features, output_features)를 입력 해 줍니다. 입력 차원과 출력 차원을 입력 해 주어야 한다는 것입니다. 층을 입력 해 줄때마다 **Input Vector**에 대해 (input_features X output_features) 에 대한 행렬 곱을 하는 것과 같습니다.
#
# `forward`에서는 순전파 연산을 진행 합니다. 파라미터로 데이터를 넣어 가면서 통과 시키면 되지만, 여기서 중간에 **ReLU** 함수를 넣는 것을 볼 수가 있습니다. 그 이유는, **y = ax + b** 꼴의 층을 쌓는다고 한들, 층을 하나 더 쌓으면 **y = c(ax + b) + d = acx + bc + d** 꼴이 되기 때문에, 학습이 되지 않습니다. 그렇기 때문에, **활성화 함수**를 사용 합니다.
#
# **ReLU** 함수는 0보다 작거나 같으면 0을, 0보다 크면 그대로 값을 반환 합니다.
# +
class DNN(torch.nn.Module):
def __init__(self, num_features, num_hidden_1, num_hidden_2, num_hidden_3, num_classes):
super(DNN, self).__init__()
self.num_classes = num_classes
self.linear_1 = torch.nn.Linear(num_features, num_hidden_1)
self.linear_2 = torch.nn.Linear(num_hidden_1, num_hidden_2)
self.linear_3 = torch.nn.Linear(num_hidden_2, num_hidden_3)
self.linear_out = torch.nn.Linear(num_hidden_3, num_classes)
def forward(self, x):
### activation 함수 변경 가능
### (optional)레이어간의 연결 추가, 변경 가능
out = self.linear_1(x)
out = torch.relu(out)
out = self.linear_2(out)
out = torch.relu(out)
out = self.linear_3(out)
out = torch.relu(out)
logits = self.linear_out(out)
probas = torch.sigmoid(logits)
return logits, probas
random.seed(RANDOM_SEED)
torch.manual_seed(RANDOM_SEED)
model = DNN(num_features=28*28,
num_hidden_1=1024,
num_hidden_2=128,
num_hidden_3=64,
num_classes=10)
model = model.to(DEVICE)
# -
# ## Training
# Training 과정에 대한 코드는 주석으로 설명 하겠습니다.
# +
optimizer = torch.optim.SGD(model.parameters(), lr=0.01) # 파라미터 학습을 위한 optimizer, 경사 하강법에 도움을 줌
NUM_EPOCHS = 20
def compute_accuracy_and_loss(model, data_loader, device): # 손실 계산
correct_pred, num_examples = 0, 0
cross_entropy = 0.
for i, (features, targets) in enumerate(data_loader): # 미니 배치 iteration
features = features.view(-1, 28*28).to(device) # view를 통해 [batch_size * 784] 크기로 변경
targets = targets.to(device) # 타겟
logits, probas = model(features) # 모델 결과 반환 (결과, sigmoid 적용한 결과)
cross_entropy += F.cross_entropy(logits, targets).item() # 타겟과 연산 결과의 cost function 결과
_, predicted_labels = torch.max(probas, 1) # 예측 결과 반환
num_examples += targets.size(0) # input 개수 반환
correct_pred += (predicted_labels == targets).sum() # 맞은 갯수 반환
return correct_pred.float()/num_examples * 100, cross_entropy/num_examples # 정확도, cost function 평균
start_time = time.time() # 시작 시간 계산
train_acc_lst, test_acc_lst = [], [] # 훈련 데이터 정확도, 테스트 데이터 정확도
train_loss_lst, test_loss_lst = [], [] # 훈련 데이터 손실함수, 테스트 데이터 손실함수
for epoch in range(NUM_EPOCHS): # EPOCH 만큼 반복
model.train() # 학습 모드
for batch_idx, (features, targets) in enumerate(train_loader): # 미니 배치 iteration
### PREPARE MINIBATCH
features = features.view(-1, 28*28).to(DEVICE)
targets = targets.to(DEVICE)
### FORWARD AND BACK PROP
logits, probas = model(features) # 순전파
cost = F.cross_entropy(logits, targets) # 예측 결과에 대한 cost function 계산
optimizer.zero_grad() # 기울기 0 초기화
cost.backward() # 역전파
### UPDATE MODEL PARAMETERS
optimizer.step() # 모델 파라미터 업데이트
### LOGGING
if not batch_idx % 40:
print (f'Epoch: {epoch+1:03d}/{NUM_EPOCHS:03d} | '
f'Batch {batch_idx:03d}/{len(train_loader):03d} |'
f' Cost: {cost:.4f}')
# 매 Epoch마다 evaluation을 진행합니다.
# Epoch마다 Loss를 기록하여 학습과정을 살펴보고 Underfitting, Overfitting 여부를 확인합니다.
model.eval()
with torch.set_grad_enabled(False): # Gradient 계산이 안되도록
train_acc, train_loss = compute_accuracy_and_loss(model, train_loader, device=DEVICE) # train acc, loss 계산
test_acc, test_loss = compute_accuracy_and_loss(model, test_loader, device=DEVICE) # test acc, loss 계산
# list에 train, test의 acc, loss 추가
train_acc_lst.append(train_acc)
test_acc_lst.append(test_acc)
train_loss_lst.append(train_loss)
test_loss_lst.append(test_loss)
# 로깅
print(f'Epoch: {epoch+1:03d}/{NUM_EPOCHS:03d} Train Acc.: {train_acc:.2f}%'
f' | Test Acc.: {test_acc:.2f}%')
# 1 epoch 학습 소요시간
elapsed = (time.time() - start_time)/60
print(f'Time elapsed: {elapsed:.2f} min')
# 총 학습 소요시간
elapsed = (time.time() - start_time)/60
print(f'Total Training Time: {elapsed:.2f} min')
# -
# ## Evaluation
# 테스트 데이터와 학습 데이터의 Loss변화를 확인합니다.
plt.plot(range(1, NUM_EPOCHS+1), train_loss_lst, label='Training loss')
plt.plot(range(1, NUM_EPOCHS+1), test_loss_lst, label='Test loss')
plt.legend(loc='upper right')
plt.ylabel('Cross entropy')
plt.xlabel('Epoch')
plt.show()
plt.plot(range(1, NUM_EPOCHS+1), train_acc_lst, label='Training accuracy')
plt.plot(range(1, NUM_EPOCHS+1), test_acc_lst, label='Test accuracy')
plt.legend(loc='upper left')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.show()
model.eval()
with torch.set_grad_enabled(False): # save memory during inference
test_acc, test_loss = compute_accuracy_and_loss(model, test_loader, DEVICE)
print(f'Test accuracy: {test_acc:.2f}%')
| example/09-DNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from nba_api.stats.endpoints import commonplayerinfo
from nba_api.stats.endpoints._base import Endpoint
from nba_api.stats.library.http import NBAStatsHTTP
from nba_api.stats.endpoints import playerdashboardbyteamperformance
from nba_api.stats.endpoints import leaguedashplayerstats
from nba_api.stats.endpoints import teamdashboardbyteamperformance
from nba_api.stats.endpoints import playerdashboardbylastngames
from nba_api.stats.endpoints import playerdashboardbyyearoveryear
from nba_api.stats.endpoints import teamdashboardbylastngames
from nba_api.stats.endpoints import teamplayerdashboard
from nba_api.stats.endpoints import teamdashlineups
from nba_api.stats.static import teams
from nba_api.stats.static import players
import pandas
import inspect
# +
#Player Input
player1_input = input('Enter player 1: ')
print(player1_input)
player1 = players.find_players_by_full_name(player1_input)[0]['id']
player1_name = players.find_players_by_full_name(player1_input)[0]['full_name']
player_info = playerdashboardbylastngames.PlayerDashboardByLastNGames(player_id=player1, season='2018-19')
player_headers = player_info.overall_player_dashboard.get_dict().get('headers')
player_data = player_info.overall_player_dashboard.get_dict().get('data')#[0]
# #last 20 games
# player_headers = player_info.last20_player_dashboard.get_dict().get('headers')
# player_data = player_info.last20_player_dashboard.get_dict().get('data')[0]
print(player_headers, '\n' , player_data)
# print(player_headers[5], player_data[5],
# player_headers[2], player_data[2],
# player_headers[26], player_data[26],
# player_headers[27], player_data[27])
# +
#Team Input
team1_input = input('Enter team 1: ')
print(team1_input)
team1 = teams.find_teams_by_full_name(team1_input)[0]['id']
teamdashboard = teamdashboardbylastngames.TeamDashboardByLastNGames(team_id=team1, season='2019-20')
teamdashboard_adv = teamdashboardbylastngames.TeamDashboardByLastNGames(team_id=team1, season='2019-20',
measure_type_detailed_defense='Advanced')
#team_headers = teamdashboard.overall_team_dashboard.get_dict().get('headers')
#team_data = teamdashboard.overall_team_dashboard.get_dict().get('data')[0]
##last 20 games
team_headers_basic = teamdashboard.last20_team_dashboard.get_dict().get('headers')
team_data_basic = teamdashboard.last20_team_dashboard.get_dict().get('data')[0]
team_headers = teamdashboard_adv.last20_team_dashboard.get_dict().get('headers')
team_data = teamdashboard_adv.last20_team_dashboard.get_dict().get('data')[0]
# print(team1, team_headers, team_data)
print(team1, team_headers[23], team_data[23], team_headers[8], team_data[8], team_headers[10], team_data[10],
team_headers[2], team_data[2], team_headers[3], team_data[3],
team_headers[4], team_data[4], team_headers[21], team_data[21], team_headers[6], team_data[6])
# +
roster1 = teamplayerdashboard.TeamPlayerDashboard(team_id=team1, last_n_games=2, season='2019-20').get_dict()#.PlayersSeasonTotals(team_id=team1, last_n_games=20)
team1_roster_last20 = []
team1_roster_lastn = []
#print(roster1)['resultSets'][1])#['rowSet'])
for n in roster1['resultSets'][1]['rowSet']:
print(n[2], 'GP:', n[3], 'id:', n[1], 'Min', n[7])
team1_roster_lastn.append(n[2])
print(team1_roster_lastn)
#(PER * MP)/n_players
# #roster1 = teamplayerdashboard.TeamPlayerDashboard(team_id=team1, last_n_games=20, season='2018-19').get_dict()#.PlayersSeasonTotals(team_id=team1, last_n_games=20)
# team1_roster_last20 = []
# roster1 = teamdashlineups.TeamDashLineups(team_id=1610612747, season='2019-20').get_dict()
# #print(roster1)['resultSets'][1])#['rowSet'])
# for n in roster1['resultSets'][1]['rowSet']:
# print(n[2], 'GP:', n[3], 'id:', n[1], 'Min', n[7])
# team1_roster_last20.append(n[2])
# print(team1_roster_last20)
# #(PER * MP)/n_players
# -
def off_per_calculator(player, team):
aPER_MP = 0
PER_MP = 0
PER_MP_n = 0
num_players = 0
agg_min = 0
PER = 0
agg_PER = 0
for n in player:
if players.find_players_by_full_name(n):
p_id = players.find_players_by_full_name(n)[0]['id']
p_info = playerdashboardbylastngames.PlayerDashboardByLastNGames(player_id=p_id, season='2019-20',
last_n_games=5)
##last 20 games
# player_headers = p_info.last20_player_dashboard.get_dict().get('headers')
# player_data = p_info.last20_player_dashboard.get_dict().get('data')[0]
player_headers = p_info.overall_player_dashboard.get_dict().get('headers')
player_data = p_info.overall_player_dashboard.get_dict().get('data')[0]
#####PER player stats####
MP = player_data[6]
ThrP = player_data[10]
AST = player_data[19]
TOV = player_data[20]
FGA = player_data[8]
FG = player_data[7] + player_data[10]
FTA = player_data[14]
FT = player_data[13]
TRB = player_data[18]
ORB = player_data[16]
STL = player_data[21]
BLK = player_data[22]
PF = player_data[24]
###PER league stats (2018-19)###
lg_AST = 24.6
lg_FG = 41.1
lg_PTS = 111.2
lg_FGA = 89.2
lg_FT = 17.7
lg_FTA = 23.1
lg_TRB = 45.2
lg_ORB = 10.3
lg_TOV = 14.1
lg_PF = 20.9
lg_pace = 100
###PER team stats###
team_AST = team_data_basic[19]
team_FG = team_data_basic[7]+team_data_basic[10]
team_pace = team_data[23]
factor = (2/3) - (0.5*(lg_AST/lg_FG)) / (2*(lg_FG/lg_FT))
VOP = lg_PTS / (lg_FGA-lg_ORB+lg_TOV+0.44*lg_FTA)
DRB_perc = (lg_TRB -lg_ORB) / lg_TRB
####PER Calculation####
uPER = (1 / MP) * (
ThrP
+ (2/3) * AST
+ (2 - factor * (team_AST / team_FG)) * FG
+ (FT *0.5 * (1 + (1 - (team_AST / team_FG))
+ (2/3) * (team_AST / team_FG)))
- VOP * TOV
- VOP * DRB_perc * (FGA - FG)
- VOP * 0.44 * (0.44 + (0.56 * DRB_perc)) * (FTA - FT)
+ VOP * (1 - DRB_perc) * (TRB - ORB)
+ VOP * DRB_perc * ORB
+ VOP * STL
+ VOP * DRB_perc * BLK
- PF * ((lg_FT / lg_PF) - 0.44 * (lg_FTA / lg_PF) * VOP) )
aPER = uPER * (lg_pace /team_pace)
PER = aPER*50
num_players +=1
aPER_MP = aPER*MP
PER_MP += aPER_MP*50
agg_min += MP
agg_PER += PER
PER_MP_n = PER_MP/num_players
print(n, 'Min', MP, 'aPER', aPER, 'PER', PER, 'aPER_MP', aPER_MP)
print('agg_PER', agg_PER)
print('PER_MP_n', PER_MP/(num_players*5))
print('team ast', team_AST, 'team FG', team_FG, 'team_pace', team_pace)
# print('league stats:', factor, VOP, DRB_perc)
off_per_calculator(team1_roster_lastn, team1)
# +
#Alternate Offensive Rating
def off_per_calc_alt(player, team):
agg_off_rating = 0
agg_min = 0
player_info_off = leaguedashplayerstats.LeagueDashPlayerStats(last_n_games=5, season='2019-20',
measure_type_detailed_defense='Advanced')
player_off_headers = player_info_off.league_dash_player_stats.get_dict().get('headers')
player_off_data = player_info_off.league_dash_player_stats.get_dict().get('data')
# print('headers:', player_off_headers)
#print('data:', player_def_data)
for p in team1_roster_lastn:
for n in player_off_data:
if p in n:
print(n[1], '\n', 'E_Off_Rating', n[10], 'Min', n[9])
agg_off_rating += (n[10]*n[9])
agg_min += n[9]
avg_off_rating = agg_off_rating/len(team1_roster_lastn)
print('agg_off_rating', agg_off_rating, 'avg_off_rating', avg_off_rating, 'agg_min', agg_min,
'agg_off_rat_min', agg_off_rating/agg_min)
# -
off_per_calc_alt(team1_roster_lastn, team1)
# +
#Individual Player Defense Stats
agg_def_rating = 0
agg_min = 0
player_info_def = leaguedashplayerstats.LeagueDashPlayerStats(last_n_games=5, season='2019-20',
measure_type_detailed_defense='Defense')
player_def_headers = player_info_def.league_dash_player_stats.get_dict().get('headers')
player_def_data = player_info_def.league_dash_player_stats.get_dict().get('data')
print('headers:', player_def_headers)
#print('data:', player_def_data)
for p in team1_roster_lastn:
for n in player_def_data:
if p in n:
print(n[1], '\n', 'Def Rating', n[10], 'Min', n[9])
agg_def_rating += (n[10]*n[9])
agg_min += n[9]
avg_def_rating = agg_def_rating/len(team1_roster_lastn)
print('agg_def_rating', agg_def_rating, 'avg_def_rating', avg_def_rating, 'agg_min', agg_min)
# -
avg_def_rating_min = agg_def_rating/agg_min
print(avg_def_rating_min)
# +
#Theory: If second game of back-to-back is a road game, odds of winning decrease 7-8%, multiply ratings by 0.93.
| NBA Stats 2019 - 2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
n = 100000
ans = 0
for i in range(n+1):
ans += np.cos(i*np.pi/n)
print(np.sqrt(n*n+n)+n*ans)
import numpy as np
a = 4000
0.5/a
a = -12345678987654321
b = 123
(b**2)/((a**2 + b**2)**0.5 - a)
sig = 1
x = 1.00001
ans = -1
for i in range(99):
ans = x * ans + sig
sig *= -1
ans
(1-x)*(1-x**100)/(1-x**2) - ans
a = 1
b = 9**12
c = -3
ans = -b - (b**2 - 4*a*c)**0.5
x1 = 2*c/ans
x2 = ans/2/a
print(x1)
print(x2)
import math
# +
def fx(x):
return math.exp(x*x)
def lagrange(x, y):
M = len(x)
p = 0.0
for j in range(M):
pt = y[j]
for k in range(M):
if k == j:
continue
fac = x[j]-x[k]
pt *= np.poly1d([1.0, -x[k]])/fac
p += pt
return p
xi = [0.6, 0.7, 0.8, 0.9, 1.0]
yi = []
for i in range(5):
ans = fx(xi[i])
yi.append(ans)
Lp = lagrange(xi, yi)
print(Lp)
print(Lp(0.82))
print(Lp(0.98))
# +
def f5(x):
# ans = fx(x)
# ans *= (16*(x**4) + 48*x*x + 12)
ans = fx(1) * (32+160+120)
return ans
def cal02(x):
ans = f5(x)
for i in range(5):
ans *= x-xi[i]
ans /= i+1.0
return ans
print(cal02(0.82))
print((Lp(0.82)-fx(0.82))/fx(0.82))
print(Lp(0.82)-fx(0.82))
print(cal02(0.98))
print((Lp(0.98)-fx(0.98))/fx(0.98))
print(Lp(0.98)-fx(0.98))
# +
import matplotlib.pyplot as plt
from pylab import *
def draw(a, b):
mpl.rcParams['font.sans-serif'] = ['SimHei']
mpl.rcParams['axes.unicode_minus'] = False
xj = np.linspace(a, b, 100)
y = []
for xjj in xj:
y.append(Lp(xjj)-fx(xjj))
plt.plot(xj, y, label='误差')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.show()
draw(0.5, 1.0)
draw(0.0, 2.0)
# -
xii = [1, 2, 4]
yii = [1, 1, 2]
Lpi = lagrange(xii, yii)
print(Lpi)
import numpy as np
x = np.array([-3, -2, -1, 0, 1, 2, 3])
y = np.array([4, 2, 3, 0, -1, -2, -5])
Y = np.array([y]).T
x0 = np.zeros(7, int) + 1
x2 = x**2
A = np.array([x0, x, x2])
A = A.T
print(np.linalg.inv(A.T.dot(A)).dot(A.T).dot(Y))
import numpy as np
x = np.array([1.02, 0.95, 0.87, 0.77, 0.67, 0.56, 0.44, 0.3, 0.16, 0.01])
y = np.array([0.39, 0.32, 0.27, 0.22, 0.18, 0.15, 0.13, 0.12, 0.13, 0.15])
Z = np.array([x**2]).T
x1 = y**2
x2 = x*y
x3 = x
x4 = y
x5 = np.zeros(10, int) + 1
A = np.array([x1, x2, x3, x4, x5]).T
print(np.linalg.inv(A.T.dot(A)).dot(A.T).dot(Z))
import numpy as np
-2-3*np.sqrt(6)
# +
import numpy as np
def f1(x):
return np.exp(x)
def f2(x):
return np.sin(x) ** 2
def f3(x):
return np.exp(-x**2)
def f4(x):
if x == 0:
return 1
return np.sin(x)/x
def f1_int(a, b):
return f1(b) - f1(a)
def f2_int(a, b):
return 1/2*(b-a) - 1/4*np.sin(2*b) + 1/4*np.sin(2*a)
def solve(fx, a, b):
ix1 = (fx(a) + fx(b))/2 * (b-a)
print(ix1)
ix2 = (fx(a) + fx((a+b)/2) + fx(b))/3 * (b-a)
print(ix2)
return
solve(f1, 1.1, 1.8)
print(f1_int(1.1, 1.8))
solve(f2, 0, np.pi/2)
print(f2_int(0, np.pi/2))
solve(f3, 1, 2)
solve(f4, 0, np.pi/2)
# -
14/np.exp(1)/12
(16*16-48*4+12)*np.exp(-1)/2880
# +
def r1(x):
return np.exp(-x**2)*(4*x**2-2)/12
def r2(x):
return np.exp(-x**2)*(16*x**4-48*x**2+12)/2880
def r3(x):
return ((x**2-2)*np.sin(x)+2*x*np.cos(x))/12/x**3
def r4(x):
return ((x**4-12*x**2+24)*np.sin(x)+(4*x**3-24*x)*np.cos(x))/2880/x**5
def solve2(rx, a, b):
r = 0
for i in np.arange(a, b, 0.001):
temp = np.fabs(rx(i))
if r < temp:
r = temp
print(r)
return
solve2(r1, 1, 2)
solve2(r2, 1, 2)
solve2(r3, 0, np.pi/2)
solve2(r4, 0, np.pi/2)
# -
import numpy as np
np.sqrt(np.exp(1)*np.exp(1)/6)
# +
import numpy as np
def fun(x):
return np.exp(x)*np.sin(x)
def tx(a,b,n):
h=(b-a)/n
x=a
s=fun(x)-fun(b)
for k in range(1,n+1):
x=x+h
s=s+2*fun(x)
res=(h/2)*s
return res
print(tx(1,2,1110))
# +
import math
import numpy as np
def fun(x):
return np.exp(x)*np.sin(x)
def T_2n(a, b, n, T_n):
h = (b - a)/n
sum_f = 0.
for k in range(0, n):
sum_f = sum_f + fun(a + (k + 0.5)*h)
T_2n = T_n/2. + sum_f*h/2.
return T_2n
def Romberg(a, b, err_min):
kmax = 99
tm = np.zeros(kmax,dtype = float)
tm1 = np.zeros(kmax,dtype = float)
tm[0] = 0.5*(b-a)*(fun(a) + fun(b))
err = 1.
k = 0
while(err>err_min):
n = 2**k
m = 1
tm1[0] = T_2n(a, b, n, tm[0])
while(err>err_min and m <= (k+1)):
tm1[m] = tm1[m-1]+(tm1[m-1]-(tm[m-1]))/(4.**m-1)
result = tm1[m]
err1 = abs(tm1[m]-tm[m-1])
err2 = abs(tm1[m]-tm1[m-1])
err = min(err1,err2)
m = m+1
tm = np.copy(tm1)
k = k+1
return result
print(Romberg(1, 3, 1.e-6))
# -
import numpy as np
x = np.array([[6, 2, 1, -1],
[2, 4, 1, 0],
[1, 1, 4, -1],
[-1, 0, -1, 3]])
np.linalg.solve(x, np.array([6, -1, 5, -5]))
# +
import numpy as np
import pandas as pd
def LU(A):
n=len(A[0])
for i in range(n):
if i==0:
for j in range(1,n):
A[j][0]=A[j][0]/A[0][0]
else:
for j in range(i, n):
temp=0
for k in range(0, i):
temp = temp+A[i][k] * A[k][j]
A[i][j]=A[i][j]-temp
for j in range(i+1, n):
temp = 0
for k in range(0, i ):
temp = temp + A[j][k] * A[k][i]
A[j][i] = (A[j][i] - temp)/A[i][i]
return A
x = np.zeros((10, 10))
x[0][0] = 9
for i in range(7):
x[i+1][i+1] = 6
x[8][8] = 5
x[9][9] = 1
for i in range(8):
x[i][i+1] = x[i+1][i] = -4
x[8][9] = x[9][8] = -2
for i in range(8):
x[i][i+2] = x[i+2][i] = 1
print(LU(x))
# -
| statistic/na/.ipynb_checkpoints/NA_test-checkpoint.ipynb |
# ### Data Prep
# This notebook will cover how to access existing data in S3 in a particular bucket.
# %run "/<EMAIL>/<EMAIL>/_helper.py"
# +
ACCESS_KEY = "[REPLACE_WITH_ACCESS_KEY]"
SECRET_KEY = "[REPLACE_WITH_SECRET_KEY]"
ENCODED_SECRET_KEY = SECRET_KEY.replace("/", "%2F")
AWS_BUCKET_NAME = get_bucket_name()
MOUNT_NAME = "mwc"
# Mount S3 bucket
try:
dbutils.fs.ls("/mnt/%s" % MOUNT_NAME)
except:
print "Mount not found. Attempting to mount..."
dbutils.fs.mount("s3n://%s:%s@%s/" % (ACCESS_KEY, ENCODED_SECRET_KEY, AWS_BUCKET_NAME), "/mnt/%s" % MOUNT_NAME)
# -
display(dbutils.fs.ls("/mnt/mwc"))
# %fs head dbfs:/mnt/mwc/accesslog/databricks.com-access.log
# * Create an external table against the access log data where we define a regular expression format as part of the serializer/deserializer (SerDe) definition.
# * Instead of writing ETL logic to do this, our table definition handles this.
# * Original Format: %s %s %s [%s] \"%s %s HTTP/1.1\" %s %s
# * Example Web Log Row
# * 10.0.0.213 - 2185662 [14/Aug/2015:00:05:15 -0800] "GET /Hurricane+Ridge/rss.xml HTTP/1.1" 200 288
# %sql
DROP TABLE IF EXISTS accesslog;
CREATE EXTERNAL TABLE accesslog (
ipaddress STRING,
clientidentd STRING,
userid STRING,
datetime STRING,
method STRING,
endpoint STRING,
protocol STRING,
responseCode INT,
contentSize BIGINT,
referrer STRING,
agent STRING,
duration STRING,
ip1 STRING,
ip2 STRING,
ip3 STRING,
ip4 STRING
)
ROW FORMAT
SERDE 'org.apache.hadoop.hive.serde2.RegexSerDe'
WITH SERDEPROPERTIES (
"input.regex" = '^(\\S+) (\\S+) (\\S+) \\[([\\w:/]+\\s[+\\-]\\d{4})\\] \\"(\\S+) (\\S+) (\\S+)\\" (\\d{3}) (\\d+) \\"(.*)\\" \\"(.*)\\" (\\S+) \\"(\\S+), (\\S+), (\\S+), (\\S+)\\"'
)
LOCATION
"/mnt/mwc/accesslog/"
# %sql select ipaddress, datetime, method, endpoint, protocol, responsecode, agent from accesslog limit 10;
# ## Obtain ISO-3166-1 Three Letter Country Codes from IP address
# * Extract out the distinct set of IP addresses from the Apache Access logs
# * Make a REST web service call to freegeoip.net to get the two-letter country codes based on the IP address
# * This creates the **mappedIP2** DataFrame where the schema is encoded.
# * Create a DataFrame to extract out a mapping between 2-letter code, 3-letter code, and country name
# * This creates the **countryCodesDF** DataFrame where the schema is inferred
# * Join these two data frames together and select out only the four columns needed to create the **mappedIP3** DataFrame
# %sql
DROP TABLE IF EXISTS distinct_ips;
create table distinct_ips as select distinct ip1 from accesslog where ip1 is not null;
select count(*) from distinct_ips;
# +
import urllib2
import json
api_key = get_ip_loc_api_key()
def getCCA2(ip):
url = 'http://api.db-ip.com/addrinfo?addr=' + ip + '&api_key=%s' % api_key
str = json.loads(urllib2.urlopen(url).read())
return str['country'].encode('utf-8')
sqlContext.udf.register("mapCCA2", getCCA2)
# -
# %sql
DROP TABLE IF EXISTS mapIps;
CREATE TABLE mapIps AS SELECT ip1 AS ip, mapCCA2(ip1) AS cca2 FROM distinct_ips;
# %sql SELECT * FROM mapIps LIMIT 40
# +
from pyspark.sql import SQLContext, Row
from pyspark.sql.types import *
fields = sc.textFile("/mnt/mwc/countrycodes/").map(lambda l: l.split(","))
countrycodes = fields.map(lambda x: Row(cn=x[0], cca2=x[1], cca3=x[2]))
sqlContext.createDataFrame(countrycodes).registerTempTable("countryCodes")
# -
# %sql
SELECT * FROM countryCodes LIMIT 20
# %sql
SELECT ip, `mapIps`.cca2 as cca2, `countryCodes`.cca3 as cca3, cn FROM mapIps LEFT OUTER JOIN countryCodes where mapIps.cca2 = countryCodes.cca2
# ## Identity the Browser and OS information
# * Extract out the distinct set of user agents from the Apache Access logs
# * Use the Python Package [user-agents](https://pypi.python.org/pypi/user-agents) to extract out Browser and OS information from the User Agent strring
# * For more information on installing pypi packages in Databricks, refer to [Databricks Guide > Product Overview > Libraries](https://docs.cloud.databricks.com/docs/latest/databricks_guide/index.html#02%20Product%20Overview/07%20Libraries.html)
# +
from user_agents import parse
from pyspark.sql.types import StringType
from pyspark.sql.functions import udf
# Convert None to Empty String
def xstr(s):
if s is None:
return ''
return str(s)
# Create UDFs to extract out Browser Family and OS Family information
def browserFamily(ua_string) : return xstr(parse(xstr(ua_string)).browser.family)
def osFamily(ua_string) : return xstr(parse(xstr(ua_string)).os.family)
sqlContext.udf.register("browserFamily", browserFamily)
sqlContext.udf.register("osFamily", osFamily)
# -
# %sql
DROP TABLE IF EXISTS userAgentTable;
DROP TABLE IF EXISTS userAgentInfo;
CREATE TABLE userAgentTable AS SELECT DISTINCT agent FROM accesslog;
CREATE TABLE userAgentInfo AS SELECT agent, osFamily(agent) as OSFamily, browserFamily(agent) as browserFamily FROM userAgentTable;
# %sql
SELECT browserFamily, count(1) FROM UserAgentInfo group by browserFamily
# ## UserID, Date, and Joins
# To make finish basic preparation of these web logs, we will do the following:
# * Convert the Apache web logs date information
# * Create a userid based on the IP address and User Agent (these logs do not have a UserID)
# * We are generating the UserID (a way to uniquify web site visitors) by combining these two columns
# * Join back to the Browser and OS information as well as Country (based on IP address) information
# * Also include call to udfWeblog2Time function to convert the Apache web log date into a Spark SQL / Hive friendly format (for session calculations below)
# +
from pyspark.sql.types import DateType
from pyspark.sql.functions import udf
import time
# weblog2Time function
# Input: 04/Nov/2015:08:15:00 +0000
# Output: 2015-11-04 08:15:00
def weblog2Time(weblog_timestr):
weblog_time = time.strptime(weblog_timestr, "%d/%b/%Y:%H:%M:%S +0000")
weblog_t = time.mktime(weblog_time)
return time.strftime("%Y-%m-%d %H:%M:%S", time.gmtime(weblog_t))
# Register the UDF
sqlContext.udf.register("weblog2Time", weblog2Time)
# -
# From here I'll use the SQL notebook to continue the SQL analysis, but refer back to this notebook for any user defined functions.
| syslog/Syslog ETL - Data Prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Learning Objectives
#
# By the end of this session you should be able to...
#
# 1. Take the derivative of a function over one variable
# 1. Take the partial derivative of a function over all of its variables
# 1. Find the minimum of the function to obtain the best line that represents relationships between two variables in a dataset
# ## Why are derivatives important?
#
# Derivatives are the foundation for Linear Regression (a topic we'll cover later in the course) that allows us to obtain the best line that represents relationships between two variables in a dataset.
#
# ## Introduction to Derivatives
#
# The process of fidning a derivative is called **Differentiation**, which is a technique used to calculate the slope of a graph at different points.
#
# ### Activity - Derivative Tutorial:
#
# 1. Go through this [Derivative tutorial from Math Is Fun](https://www.mathsisfun.com/calculus/derivatives-introduction.html) (15 min)
# 1. When you're done, talk with a partner about topics you still have questions on. See if you can answer each other's questions. (5 min)
# 1. We'll then go over questions on the tutorial as a class (10 min)
#
# ### Review Diagram
#
# Review the below diagram as a class, and compare with what you just learned in the above Derivative Tutorial. Note that a Gradient Function is just another name for the Derivative of a function:
#
# <img src="diff_y_x2.png" width="600" height="600">
# <img src="diff_y_x2_gragh.png" width="600" height="600">
# ## Derivative Formula
#
# - Choose small $\Delta x$
#
# - $f^\prime(x) = \frac{d}{dx}f(x) = \frac{\Delta x}{\Delta y} = \frac{f(x + \Delta x) - f(x)}{\Delta x}$
#
# Remember that $\Delta x$ approaches 0. So if plugging in a value in the above formula, choose a _very_ small number, or simplify the equation further such that all $\Delta x = 0$, like we saw in the tutorial
# ## Activity: Write a Python function that calculates the gradient of $x^2$ at $x = 3$ and $x = -2$ using the above definition
# +
def f(x):
return x**2
eps = 1e-6
x = 3
print((f(x + eps) - f(x)) / eps)
x = -2
print((f(x + eps) - f(x)) / eps)
# -
# Note that these values match $2x$, our derivative of $x^2$:
#
# $2*3 = 6$
#
# $2 * -2 = -4$
# ## Derivative Table
#
# As a shortcut, use the second page of this PDF to find the derivative for common formulas. Utilize this as a resource going forward!
#
# - https://www.qc.edu.hk/math/Resource/AL/Derivative%20Table.pdf
# ## Extend Gradient into Two-Dimensional Space
# Now we know how to calculate a derivative of one variable. But what if we have two?
#
# To do this, we need to utilize **Partial Derivatives**. Calculating a partial derivative is essentially calculating two derivatives for a function: one for each variable, where they other variable is set to a constant.
#
# ### Activity - Partial Derivative Video
#
# Lets watch this video about Partial Derivative Intro from Khan Academy: https://youtu.be/AXqhWeUEtQU
#
# **Note:** Here are some derivative shortcuts that will help in the video:
#
# $\frac{d}{dx}x^2 = 2x$
#
# $\frac{d}{x}sin(x) = cos(x)$
#
# $\frac{d}{dx}x = 1$
#
# ### Activity - Now You Try!
# Consider the function $f(x, y) = \frac{x^2}{y}$
#
# - Calculate the first order partial derivatives ($\frac{\partial f}{\partial x}$ and $\frac{\partial f}{\partial y}$) and evaluate them at the point $P(2, 1)$.
# ## We can use the Symbolic Python package (library) to compute the derivatives and partial derivatives
from sympy import symbols, diff
# initialize x and y to be symbols to use in a function
x, y = symbols('x y', real=True)
f = (x**2)/y
# Find the partial derivatives of x and y
fx = diff(f, x, evaluate=True)
fy = diff(f, y, evaluate=True)
print(fx)
print(fy)
# print(f.evalf(subs={x: 2, y: 1}))
print(fx.evalf(subs={x: 2, y: 1}))
print(fy.evalf(subs={x: 2, y: 1}))
# ## Optional Reading: Tensorflow is a powerful package from Google that calculates the derivatives and partial derivatives numerically
# +
import tensorflow as tf
x = tf.Variable(2.0)
y = tf.Variable(1.0)
with tf.GradientTape(persistent=True) as t:
z = tf.divide(tf.multiply(x, x), y)
# Use the tape to compute the derivative of z with respect to the
# intermediate value x and y.
dz_dx = t.gradient(z, x)
dz_dy = t.gradient(z, y)
print(dz_dx)
print(dz_dy)
# All at once:
gradients = t.gradient(z, [x, y])
print(gradients)
del t
# -
# ## Optional Reading: When x and y are declared as constant, we should add `t.watch(x)` and `t.watch(y)`
# +
import tensorflow as tf
x = tf.constant(2.0)
y = tf.constant(1.0)
with tf.GradientTape(persistent=True) as t:
t.watch(x)
t.watch(y)
z = tf.divide(tf.multiply(x, x), y)
# Use the tape to compute the derivative of z with respect to the
# intermediate value y.
dz_dx = t.gradient(z, x)
dz_dy = t.gradient(z, y)
# -
# # Calculate Partial Derivative from Definition
# +
def f(x, y):
return x**2/y
eps = 1e-6
x = 2
y = 1
print((f(x + eps, y) - f(x, y)) / eps)
print((f(x, y + eps) - f(x, y)) / eps)
# -
# Looks about right! This works rather well, but it is just an approximation. Also, you need to call `f()` at least once per parameter (not twice, since we could compute `f(x, y)` just once). This makes this approach difficult to control for large systems (for example neural networks).
# ## Why Do we need Partial Gradients?
#
# In many applications, more specifically DS applications, we want to find the Minimum of a cost function
#
# - **Cost Function:** a function used in machine learning to help correct / change behaviour to minimize mistakes. Or in other words, a measure of how wrong the model is in terms of its ability to estimate the relationship between x and y. [Source](https://towardsdatascience.com/machine-learning-fundamentals-via-linear-regression-41a5d11f5220)
#
#
# Why do we want to find the minimum for a cost function? Given that a cost function mearues how wrong a model is, we want to _minimize_ that error!
#
# In Machine Learning, we frequently use models to run our data through, and cost functions help us figure out how badly our models are performing. We want to find parameters (also known as **weights**) to minimize our cost function, therefore minimizing error!
#
# We find find these optimal weights by using a **Gradient Descent**, which is an algorithm that tries to find the minimum of a function (exactly what we needed!). The gradient descent tells the model which direction it should take in order to minimize errors, and it does this by selecting more and more optimal weights until we've minimized the function! We'll learn more about models when we talk about Linear Regression in a future lesson, but for now, let's review the Gradient Descent process with the below images, given weights $w_0$ and $w_1$:
#
# <img src="gradient_descent.png" width="800" height="800">
#
# Look at that bottom right image. Looks like we're using partial derivatives to find out optimal weights. And we know exactly how to do that!
# ## Finding minimum of a function
#
# Assume we want to minimize the function $J$ which has two weights $w_0$ and $w_1$
#
# We have two options to find the minimum of $J(w_0, w_1)$:
#
# 1. Take partial derivatives of $J(w_0, w_1)$ with relation to $w_0$ and $w_1$:
#
# $\frac{\partial J(w_0, w_1)}{\partial w_0}$
#
# $\frac{\partial J(w_0, w_1)}{\partial w_1}$
#
# And find the appropriate weights such that the partial derivatives equal 0:
#
# $\frac{\partial J(w_0, w_1)}{\partial w_0} = 0$
#
# $\frac{\partial J(w_0, w_1)}{\partial w_1} = 0$
#
# In this approach we should solve system of linear or non-linear equation
#
# 2. Use the Gradient Descent algorithm:
#
# First we need to define two things:
#
# - A step-size alpha ($\alpha$) as a small number (like $1^(e-6)$ small)
# - An arbitrary random initial value for $w_0$ and $w_1$: $w_0 = np.random.randn()$ and $w_1 = np.random.randn()$
#
# Finally, we need to search for the most optimal $w_0$ and $w_1$ by using a loop to update the weights until we find the most optimal weights. We'll need to establish a threshold to compare weights to, toi know when to stop the loop. For example, if a weight from one iteration is within 0.0001 of the weight from the next iteration, we can stop the loop (0.0001 is our threshold here)
#
# Let's review some pseudocode for how to implement this algorithm:
#
# ```
# initialize the following:
# a starting weight value
# the learning rate (alpha) (very small)
# the threshold (small)
# the current threshold (start at 1)
#
# while the current threshold is less than the threshold:
# store the current values of the weights into a previous value variable
# set the weight values to new values based on the algorithm
# set the current threshold from the difference of the current weight value and the previous weight value
# ```
#
# How do we `set the weight values to new values based on the algorithm`? by using the below equations:
#
# $w_0 = w_0 - \alpha \frac{\partial J(w_0, w_1)}{\partial w_0}$
#
# $w_1 = w_1 - \alpha \frac{\partial J(w_0, w_1)}{\partial w_1}$
#
# Try to write the function yourself, creating real code from the pseudocode!
#
# **Stretch Challenge:** We may also want to limit the number of loops we do, in addition to checking the threshold. Determine how we may go about doing that
#
#
# ## Resources
#
# - [Derivative tutorial from Math Is Fun](https://www.mathsisfun.com/calculus/derivatives-introduction.html)
# - [Derivative Table](https://www.qc.edu.hk/math/Resource/AL/Derivative%20Table.pdf)
# - [Khan Academy - Partial Derivatives video](https://www.youtube.com/watch?v=AXqhWeUEtQU&feature=youtu.be)
# - [Towards Data Science - Machine Learning Fundamentals: cost functions and gradient Descent](https://towardsdatascience.com/machine-learning-fundamentals-via-linear-regression-41a5d11f5220)
| Notebooks/Calculus/.ipynb_checkpoints/partial_derivative-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CHfPKUdwtHsj" colab_type="text"
# <a href="https://colab.research.google.com/drive/1fb6wt9QKQrSKDwDK7HSXUEqfE9cQ1cUO?usp=sharing" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# By [<NAME>](https://www.linkedin.com/in/ibrahim-sobh-phd-8681757/)
#
# + [markdown] id="uzSsqB7Qymsj" colab_type="text"
# # DDPG [Deep Deterministic Policy Gradient](https://arxiv.org/pdf/1509.02971.pdf)
#
# **Deterministic Policy** means that the Actor gives the *best* believed action for any given state (no argmax).
#
# In DDPG, the ideas underlying the (DQN) are adapted to **continuous action** domains.
#
#
# + [markdown] id="X8MpsMtnyrk4" colab_type="text"
# ## Install
# + id="AK88-CxtIzwm" colab_type="code" outputId="c178e4e2-ae49-49fb-9461-c6df8e04abdb" executionInfo={"status": "ok", "timestamp": 1582016874739, "user_tz": -120, "elapsed": 13901, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 272}
# !pip install gym
# !pip install box2d-py
# # !pip install pyglet==1.3.2
# !pip install pyglet
# + id="i5RcREGXw5ed" colab_type="code" outputId="aab684de-7725-46d8-9d83-ffee0c46514f" executionInfo={"status": "ok", "timestamp": 1582016881330, "user_tz": -120, "elapsed": 4269, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 908}
# !pip install stable-baselines[mpi]
# + [markdown] id="6NlznQ0_y2jv" colab_type="text"
# ## Agent and Env.
# + id="u6bC_jjPyfoC" colab_type="code" outputId="a03ca8a4-163e-4aa5-b3ba-05d1f7808448" executionInfo={"status": "ok", "timestamp": 1582016889110, "user_tz": -120, "elapsed": 5664, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 210}
import gym
import numpy as np
from stable_baselines.ddpg.policies import MlpPolicy
from stable_baselines.common.vec_env import DummyVecEnv
from stable_baselines.ddpg.noise import NormalActionNoise, OrnsteinUhlenbeckActionNoise, AdaptiveParamNoiseSpec
from stable_baselines import DDPG
from stable_baselines.results_plotter import load_results, ts2xy
from stable_baselines import results_plotter
from stable_baselines.bench import Monitor
import time
import matplotlib.pyplot as plt
# + id="hOGZameNHuXB" colab_type="code" outputId="ffea85a8-6630-4065-c6c9-212fe1c2c9b7" executionInfo={"status": "ok", "timestamp": 1582016890014, "user_tz": -120, "elapsed": 524, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
env = gym.make('Pendulum-v0')
env = DummyVecEnv([lambda: env])
# noise objects for DDPG
n_actions = env.action_space.shape[-1]
param_noise = None
action_noise = OrnsteinUhlenbeckActionNoise(mean=np.zeros(n_actions), sigma=float(0.5) * np.ones(n_actions))
# + [markdown] id="4_cL6yu5y7l2" colab_type="text"
# ## Train
# + id="67BI65Q1sWoS" colab_type="code" colab={}
total_timesteps = 100000
best_mean_reward = -np.inf
n_steps = 0
score_list = []
avg_scores = []
episodes = 0
prev_episode_score = 0
prev_episode_step = 0
save_every = 10000
import os
# Create log dir
log_dir = "log/"
os.makedirs(log_dir, exist_ok=True)
# + id="q5iq_eE-M3Or" colab_type="code" colab={}
def callback(_locals, _globals):
"""
Callback called at each step (for DQN an others) or after n steps (see ACER or PPO2)
:param _locals: (dict)
:param _globals: (dict)
_locals['reward'], current reward
_locals['episodes'], episode number
_locals['episode_step'], steps in episode
_locals['episode_reward']), accumilative reward
_locals['total_timesteps']
_locals['episode_rewards_history'] last 100
"""
global n_steps, best_mean_reward, episodes, prev_episode_step, score_list, avg_scores, prev_episode_score
if _locals['episodes'] > episodes: # new episode
episodes = _locals['episodes']
total_timesteps = _locals['total_timesteps']
score_list.append(prev_episode_score)
avg_scores.append(np.mean(score_list[-10:]))
print("\rStep({}/{} [{:.2f}%]) - Episode ({}) - Lengh({}) - Reward({:.2f}) - Reward mean({:.2f}) ".format(
n_steps,
total_timesteps,
n_steps/total_timesteps*100.0,
episodes,
prev_episode_step,
prev_episode_score,
np.mean(_locals['episode_rewards_history']))
, end="")
prev_episode_score = _locals['episode_reward']
prev_episode_step = _locals['episode_step']
if (n_steps + 1) % save_every == 0:
_locals['self'].save(log_dir + 'best_model.pkl')
n_steps += 1
return True
# + id="p1JCG_wEN5OQ" colab_type="code" outputId="932cfbb7-a10a-4307-bf2d-9732c2562451" executionInfo={"status": "ok", "timestamp": 1582017472250, "user_tz": -120, "elapsed": 545409, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 801}
model = DDPG(MlpPolicy, env, verbose=0, param_noise=param_noise, action_noise=action_noise)
model.learn(total_timesteps=total_timesteps, callback=callback)
model.save("ddpg_mountain")
# + id="ysROpwwqoAjy" colab_type="code" outputId="5a184a9c-a811-4053-de29-2e83133bada8" executionInfo={"status": "ok", "timestamp": 1582017505867, "user_tz": -120, "elapsed": 1435, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 279}
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(score_list)), score_list)
plt.plot(np.arange(len(avg_scores)), avg_scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# + id="m3RIP9Adz_F8" colab_type="code" colab={}
del model # to demonstrate saving and loading
# + [markdown] id="UuywPEOuzAfQ" colab_type="text"
# ## Test
# + id="Mbof27bgzsIh" colab_type="code" outputId="9935914c-1747-4b0e-d07d-bbeb8d5a74f4" executionInfo={"status": "ok", "timestamp": 1582017514113, "user_tz": -120, "elapsed": 774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
env_test = gym.make('Pendulum-v0')
# + id="zVcXS9-HzCQK" colab_type="code" outputId="7036653e-3c54-4c7d-c28c-29cc130c3601" executionInfo={"status": "ok", "timestamp": 1582017522629, "user_tz": -120, "elapsed": 2127, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYeQM_1LvthZBu6TRjB85IH8s-a5ZMla2NYwtX=s64", "userId": "05222794542711757573"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
model = DDPG.load("ddpg_mountain")
r = 0.0
obs = env_test.reset()
while True:
action, _states = model.predict(obs)
obs, rewards, dones, info = env_test.step(action)
r += rewards
if dones:
break
r
# + id="XoldUpdZ5jhN" colab_type="code" colab={}
| 07_Stable_Baselines_DDPG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Rvr5GN9tFFu_"
# # **Numpy Assignment**
# + [markdown] id="HxmjggX8FZCe"
# **Write a function so that the columns of the output matrix are powers of the input vector.**
#
# **The order of the powers is determined by the increasing boolean argument. Specifically, when increasing is False, the i-th output column is the input vector raised element-wise to the power of N - i - 1.**
#
# **Hint:** *Such a matrix with a geometric progression in each row is named for <NAME>.*
# + id="x0eFwtHcFCXa" colab={"base_uri": "https://localhost:8080/"} outputId="15b183c9-a0b0-42e0-e3bb-877173f24eda"
import numpy as np
def alexaxndreTheophile(inputVector, n, increasing):
if increasing:
outMatrix = np.matrix([x**i for x in inputVector for i in range(n)]).reshape(inputVector.size, n)
else:
outMatrix = np.matrix([x**(n-i-1) for x in inputVector for i in range(n)]).reshape(inputVector.size, n)
return outMatrix
inputVector = np.array([1,2,3,4,5])
outMatrix = alexaxndreTheophile(inputVector, inputVector.size, increasing = True)
print("When Increasing = True: \n",outMatrix)
outMatrix = alexaxndreTheophile(inputVector, inputVector.size, increasing = False)
print("\nWhen Increasing = False: \n",outMatrix)
# + [markdown] id="lk57x52RFGfI"
# ## <u>**Problem Statement**</u>
# **Given a sequence of n values x1, x2, ..., xn and a window size k>0, the k-th moving average of the given sequence is defined as follows:**
#
# **The moving average sequence has n-k+1 elements as shown below.**
#
# **The moving averages with k=4 of a ten-value sequence (n=10) is shown below:**
#
# input | 10 20 30 40 50 60 70 80 90 100
# --- | ---
# y1 | 25 = (10+20+30+40)/4
# y2 | 35 = (20+30+40+50)/4
# y3 | 45 = (30+40+50+60)/4
# y4 | 55 = (40+50+60+70)/4
# y5 | 65 = (50+60+70+80)/4
# y6 | 75 = (60+70+80+90)/4
# y7 | 85 = (70+80+90+100)/4
#
# Thus, the moving average sequence has n-k+1=10-4+1=7 values.
# + [markdown] id="3S3ceH1BpTKM"
# ### **Question:**
#
# *Write a function to find moving average in an array over a window:*
#
# *Test it over [3, 5, 7, 2, 8, 10, 11, 65, 72, 81, 99, 100, 150] and window of 3.*
# + colab={"base_uri": "https://localhost:8080/"} id="cRpV64AvDZME" outputId="3f705ae4-91df-487f-91e0-90264709d4cb"
def movingAverage(inputVector, windowLength):
c = 1
movingAvg = np.convolve(inputVector, np.ones(windowLength), 'valid') / windowLength
for i in movingAvg:
print("y{0} = {1:.2f}".format(c, i))
c += 1
inputVector = np.array([3, 5, 7, 2, 8, 10, 11, 65, 72, 81, 99, 100, 150])
windowLength = 3
movingAverage(inputVector, windowLength)
| numpy_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create images ids
# +
# %run "../config/local.ipynb"
# %run "../utils/functions.ipynb"
import numpy as np
import matplotlib.pyplot as plt
import cv2
import tqdm.notebook as tq
import math
import xml.etree.ElementTree as ET
import pandas as pd
# -
# ## Build
images = os.listdir(ORIGINAL_FEATURES_DIR)
df_images_ids = pd.DataFrame(columns=['name', 'file_name'])
df_images_ids['file_name'] = images
df_images_ids['name'] = [image.replace(".png", "") for image in images]
df_images_ids = df_images_ids.set_index('name')
# ## Save
df_images_ids.to_csv(IMAGES_IDS_FILE)
# ## Check dataframe
df_images_ids.head()
| notebooks/prepare/create_images_ids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load the data and preprocessing modules
from modules.preprocess import Vocab
with open('./data/dl_history.txt') as f:
text = f.read()
vocab = Vocab(text, max_size = 100, lower = True, one_hot = True)
print(len(vocab)) # size of the vocabulary
sents = vocab.sents2id(text)
print(sents)
print(vocab.id2sents(sents))
# # No GPU
# +
import torch
import torch.nn as nn
from torch.autograd import Variable
class LSTM(nn.Module):
def __init__(self, input_size, hidden_size, num_layers):
super().__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.lstm = nn.LSTM(input_size, hidden_size, num_layers)
def forward(self, input, h, c):
output,(h,c) = self.lstm(input,(h,c))
return output,h,c
def init_h0c0(self, batch = 1):
# dimension: num_layers*num_directions, batch_size, hidden_size
h0 = Variable(torch.zeros(self.num_layers, batch_size, self.hidden_size))
c0 = Variable(torch.zeros(self.num_layers, batch_size, self.hidden_size))
return h0,c0
input_size = len(vocab)
hidden_size = len(vocab)
num_layers = 1
rnn = LSTM(input_size, hidden_size, num_layers)
# -
# Little note about the view function
seq_len = 43
batch_size = 1
dim = 50
x = torch.arange(0,seq_len*dim).view(seq_len,dim)
print(x)
print(x.view(seq_len, batch_size, -1))
# +
# Process one string with a zero-vector inital hidden state / cell state
inputs = vocab.id2emb(sents[0])
seq_len = inputs.size()[0]
batch_size = 1
inputs = Variable(inputs.view(seq_len, batch_size, -1))
h0,c0 = rnn.init_h0c0()
output,h,c = rnn(inputs, h0, c0)
print(output,h,c)
# +
# Build the Training dataset
onehots = [vocab.id2emb(sent) for sent in sents]
# Build inputs / targets as lists of tensors
inputs = [sent[:-1,:] for sent in onehots]
targets = [sent[1:,:] for sent in onehots]
# +
input_size = len(vocab)
hidden_size = len(vocab)
num_layers = 1
batch_size = 1
rnn = LSTM(input_size, hidden_size, num_layers)
import torch.optim as optim
loss_fn = nn.MSELoss()
optimizer = optim.Adam(rnn.parameters(), lr = .005)
def run_epoch(inputs, targets):
# flush the gradients
optimizer.zero_grad()
# initial hidden state(h0)
h,c = rnn.init_h0c0()
# training loss
loss = 0
# Run a RNN through the training samples
for i in range(len(inputs)):
input = inputs[i]
target = targets[i]
seq_len = input.size()[0]
input = Variable(input.view(seq_len, batch_size, -1))
target = Variable(target.view(seq_len, batch_size, -1))
# Note: new hidden layer output is generated for every loop, so we have to send the
# hidden weights to cuda for every loop
output, h, c = rnn(input, h, c)
loss += loss_fn(output, target)
loss.backward()
optimizer.step()
return output, loss.data[0]
def train(inputs, targets, n_epochs = 100, print_every = 10):
total_loss = 0.0
for epoch in range(1, n_epochs + 1):
output, loss = run_epoch(inputs, targets)
if epoch % print_every == 0:
print('Epoch: %2i / Loss: %.7f' % (epoch, loss))
def test(input_sent):
h, c = rnn.init_h0c0()
seq_len = input_sent.size()[0]
input_sent = Variable(input_sent.view(seq_len, batch_size, -1))
output, h, c = rnn(input_sent, h, c)
_, argmaxs = torch.max(output, dim = 0)
sent = argmaxs.view(-1).data.numpy().tolist()
for i in sent:
print(vocab[i],end=' ')
# -
# run_epoch
train(inputs, targets, n_epochs = 1000, print_every = 100)
# Test
torch.manual_seed(7)
for i in range(len(inputs)):
try:
test(inputs[i])
print()
except Exception as e:
print(e)
print()
# # GPU version
# ### Dealing with Variable-length sequences for cuDNN in PyTorch
# - References
# - [Simple working example how to use packing for variable-length sequence inputs for rnn](https://discuss.pytorch.org/t/simple-working-example-how-to-use-packing-for-variable-length-sequence-inputs-for-rnn/2120)
# - [Feeding Data to PyTorch RNNs](https://djosix.github.io/2017/09/05/Feeding-Data-to-Pytorch-RNNs/)
# - [How to use pad_packed_sequence in PyTorch](https://www.snip2code.com/Snippet/1950100/How-to-use-pad_packed_sequence-in-pytorc)
# - [RNN sequence padding with batch_first](https://github.com/pytorch/pytorch/issues/1176)
# - [padded_rnn.py](https://gist.github.com/MaximumEntropy/918d4ad7c931bc14b475008c00aa09f1)
# - [About the variable length input in RNN scenario](https://discuss.pytorch.org/t/about-the-variable-length-input-in-rnn-scenario/345/7)
# - [How can i compute seq2seq loss using mask?](https://discuss.pytorch.org/t/how-can-i-compute-seq2seq-loss-using-mask/861/7)
# - Steps
# 1. Pad the input sequences to the same length
# 2. Sort them by their lengths (asc order)
# 3. Use torch.nn.utils.rnn.pack_padded_sequence()
# 4. RNN
# 5. Use torch.nn.utils.rnn.pad_packed_sequence()
# 6. Unsort output sequences
# 7. Unpad output sequences
from modules.preprocess import Vocab
with open('./data/dl_history.txt') as f:
text = f.read()
vocab = Vocab(text, max_size = 100, lower = True, one_hot = True)
print(len(vocab)) # size of the vocabulary
sents = vocab.sents2id(text)
print(sents)
print(vocab.id2sents(sents))
import torch
from torch.autograd import Variable
from torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence
# ### A little exercise on `pack_padded_sequence`, and `pad_packed_sequence`
max_seq_len = 20
num_batches = 10
x = Variable(torch.randn(max_seq_len, num_batches, 30))
lens = list(range(max_seq_len, max_seq_len - num_batches, -1)) # sequence of lengths of each batches
x_packed = pack_padded_sequence(x, lens)
lens
x
x_packed
# ### Define the packing/unpacking functions
# To utilize cuDNN for variable length input on PyTorch, we need to use `torch.utils.nn.rnn.pack_padded_sequence` and `torch.utils.nn.rnn.pad_packed_sequence`.
# - `pack_padded_sequence` packs a **padded** tensor into a `PackedSequence` object, which is internally handled by nn.LSTM.
# - `pad_padded_sequence` unpacks a `PackedSequence object` into a tensor
# +
# Build the Training dataset
onehots = [vocab.id2emb(sent) for sent in sents]
# Build inputs / targets as lists of tensors
inputs = [sent[:-1,:] for sent in onehots]
targets = [sent[1:,:] for sent in onehots]
# +
# Helper functions
def pack(seq):
'''
Packs a list of variable-length tensors into a packed sequence
Args:
seq: 2 dim tensor, where each row corresponds to an individual element.
Returns:
packed: PackedSequence
orders: ordered indices for the original sequence before the sorting.
later used to retrieve the original ordering of the sequences.
'''
seq_sorted = []
orders = []
for i, tensor in sorted(enumerate(seq), key = lambda t: -t[1].size()[0]):
seq_sorted.append(tensor)
orders.append(i)
lengths = list(map(lambda t: t.size()[0], seq_sorted))
max_seq_len = lengths[0]
dim = seq_sorted[0].size()[1]
batch_size = len(seq_sorted)
# Build a padded sequence
padded_sequence = Variable(torch.zeros(max_seq_len, batch_size, dim))
if torch.cuda.is_available():
padded_sequence = padded_sequence.cuda()
for i in range(batch_size):
padded_sequence[:lengths[i], i, :] = seq_sorted[i]
# pack the padded sequence
packed = pack_padded_sequence(padded_sequence, lengths)
return packed, orders
def unpack(packed, orders):
'''
Unpacks a packed sequence
Args:
packed: PackedSequence
Returns:
unpacked_masked
'''
unpacked, lengths = pad_packed_sequence(packed)
# Masking
unpacked_masked = [unpacked[:lengths[batch], batch, :] for batch in range(len(lengths))]
# Unsort
unpacked_masked = [tensor for i, tensor in sorted(zip(orders, unpacked_masked))]
return unpacked_masked
packed, orders = pack(inputs)
print(packed)
unpacked = unpack(packed, orders)
print(unpacked)
# +
# Define the LSTM Cell
import torch
import torch.nn as nn
from torch.autograd import Variable
class LSTM(nn.Module):
def __init__(self, input_size, hidden_size, num_layers):
super().__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.lstm = nn.LSTM(input_size, hidden_size, num_layers)
if torch.cuda.is_available():
self.cuda()
def forward(self, input, h, c):
output,(h,c) = self.lstm(input,(h,c))
return output,h,c
def init_h0c0(self, batch_size = 1):
# dimension: num_layers*num_directions, batch_size, hidden_size
h0 = Variable(torch.zeros(self.num_layers, batch_size, self.hidden_size))
c0 = Variable(torch.zeros(self.num_layers, batch_size, self.hidden_size))
if torch.cuda.is_available():
h0 = h0.cuda()
c0 = c0.cuda()
return h0,c0
# +
# Test over one input
input_size = len(vocab)
hidden_size = len(vocab)
num_layers = 1
rnn = LSTM(input_size, hidden_size, num_layers).cuda()
inputs_packed, orders = pack(inputs)
h0,c0 = rnn.init_h0c0(batch_size = 5)
outputs_packed, h, c = rnn(inputs_packed, h0, c0)
outputs = unpack(outputs_packed, orders)
print(list(map(lambda t: t.size(),outputs)))
# +
# Train the network
input_size = len(vocab)
hidden_size = len(vocab)
num_layers = 1
batch_size = 1
rnn = LSTM(input_size, hidden_size, num_layers).cuda()
import torch.optim as optim
loss_fn = nn.MSELoss()
optimizer = optim.Adam(rnn.parameters(), lr = .005)
def run_epoch(inputs, targets):
# flush the gradients
optimizer.zero_grad()
# initial hidden state(h0)
h,c = rnn.init_h0c0(batch_size = 5)
# training loss
loss = 0
targets = [Variable(tensor).cuda() for tensor in targets]
# Run a RNN through the training samples
inputs_packed, orders = pack(inputs)
outputs_packed, h, c = rnn(inputs_packed, h, c)
outputs = unpack(outputs_packed, orders)
for out, target in zip(outputs, targets):
loss += loss_fn(out, target)
loss.backward()
optimizer.step()
return outputs, loss.data[0]
def train(inputs, targets, n_epochs = 100, print_every = 10):
total_loss = 0.0
for epoch in range(1, n_epochs + 1):
output, loss = run_epoch(inputs, targets)
if epoch % print_every == 0:
print('Epoch: %2i / Loss: %.7f' % (epoch, loss))
def test(input_sent):
h, c = rnn.init_h0c0()
seq_len = input_sent.size()[0]
input_sent = Variable(input_sent.view(seq_len, batch_size, -1))
output, h, c = rnn(input_sent, h, c)
_, argmaxs = torch.max(output, dim = 0)
# flatten the sorted indices
sent = argmaxs.view(-1).data.cpu().numpy().tolist()
for i in sent:
print(vocab[i],end=' ')
# run_epoch(inputs, targets)
train(inputs, targets, n_epochs = 1000, print_every = 100)
torch.manual_seed(7)
for i in range(len(inputs)):
try:
test(inputs[i].cuda())
print()
except Exception as e:
print(e)
print()
| srcs/pytorch/tutorial/03. Word-level Language Modeling with naive LSTM(with and without GPU).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression Example
#
# Let's walk through the steps of the official documentation example. Doing this will help your ability to read from the documentation, understand it, and then apply it to your own problems.
from pyspark.sql import SparkSession
# May take a little while on a local computer
spark = SparkSession.builder.appName("Linear Regression").getOrCreate()
# check (try) if Spark session variable (spark) exists and print information about the Spark context
try:
spark
except NameError:
print("Spark session does not context exist. Please create Spark session first (run cell above).")
else:
configurations = spark.sparkContext.getConf().getAll()
for item in configurations: print(item)
#import numpy as np
from pyspark.ml.regression import LinearRegression
# Load training data
training = spark.read.format("libsvm").load("data/sample_linear_regression_data.txt")
# Interesting! We haven't seen libsvm formats before. In fact the aren't very popular when working with datasets in Python, but the Spark Documentation makes use of them a lot because of their formatting. Let's see what the training data looks like:
training.show()
# This is the format that Spark expects. Two columns with the names "label" and "features".
#
# The "label" column then needs to have the numerical label, either a regression numerical value, or a numerical value that matches to a classification grouping. Later on we will talk about unsupervised learning algorithms that by their nature do not use or require a label.
#
# The feature column has inside of it a vector of all the features that belong to that row. Usually what we end up doing is combining the various feature columns we have into a single 'features' column using the data transformations we've learned about.
#
# Let's continue working through this simple example!
# These are the default values for the featuresCol, labelCol, predictionCol
lr = LinearRegression(featuresCol='features', labelCol='label', predictionCol='prediction')
# Fit the model
lrModel = lr.fit(training)
# Print the coefficients and intercept for linear regression
print("Coefficients: {}".format(str(lrModel.coefficients))) # For each feature...
print('\n')
print("Intercept:{}".format(str(lrModel.intercept)))
# There is a summary attribute that contains even more info!
# Summarize the model over the training set and print out some metrics
trainingSummary = lrModel.summary
# Lots of info, here are a few examples:
trainingSummary.residuals.show()
print("RMSE: {}".format(trainingSummary.rootMeanSquaredError))
print("r2: {}".format(trainingSummary.r2))
# ## Train/Test Splits
#
# But wait! We've commited a big mistake, we never separated our data set into a training and test set. Instead we trained on ALL of the data, something we generally want to avoid doing. Read ISLR and check out the theory lecture for more info on this, but remember we won't get a fair evaluation of our model by judging how well it does again on the same data it was trained on!
#
# Luckily Spark DataFrames have an almost too convienent method of splitting the data! Let's see it:
all_data = spark.read.format("libsvm").load("data/sample_linear_regression_data.txt")
# Pass in the split between training/test as a list.
# No correct, but generally 70/30 or 60/40 splits are used.
# Depending on how much data you have and how unbalanced it is.
train_data,test_data = all_data.randomSplit([0.7,0.3])
train_data.show()
test_data.show()
unlabeled_data = test_data.select('features')
unlabeled_data.show()
# Now we only train on the train_data
correct_model = lr.fit(train_data)
# Now we can directly get a .summary object using the evaluate method:
test_results = correct_model.evaluate(test_data)
test_results.residuals.show()
print("RMSE: {}".format(test_results.rootMeanSquaredError))
# Well that is nice, but realistically we will eventually want to test this model against unlabeled data, after all, that is the whole point of building the model in the first place. We can again do this with a convenient method call, in this case, transform(). Which was actually being called within the evaluate() method. Let's see it in action:
predictions = correct_model.transform(unlabeled_data)
predictions.show()
# Okay, so this data is a bit meaningless, so let's explore this same process with some data that actually makes a little more intuitive sense!
# # Linear Regression Code Along
# Basically what we do here is examine a dataset with Ecommerce Customer Data for a company's website and mobile app. Then we want to see if we can build a regression model that will predict the customer's yearly spend on the company's product.
# First thing to do is start a Spark Session
# Use Spark to read in the Ecommerce Customers csv file.
data = spark.read.csv("data/Ecommerce_Customers.csv",inferSchema=True,header=True)
# Print the Schema of the DataFrame
data.printSchema()
data.show()
data.head()
for item in data.head():
print(item)
# ## Setting Up DataFrame for Machine Learning
# +
# A few things we need to do before Spark can accept the data!
# It needs to be in the form of two columns
# ("label","features")
# Import VectorAssembler and Vectors
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
# -
data.columns
assembler = VectorAssembler(
inputCols=["Avg Session Length", "Time on App",
"Time on Website",'Length of Membership'],
outputCol="features")
output = assembler.transform(data)
output.select("features").show()
output.show()
final_data = output.select("features",'Yearly Amount Spent')
train_data,test_data = final_data.randomSplit([0.7,0.3])
train_data.describe().show()
test_data.describe().show()
# Create a Linear Regression Model object
lr = LinearRegression(labelCol='Yearly Amount Spent')
# Fit the model to the data and call this model lrModel
lrModel = lr.fit(train_data,)
# Print the coefficients and intercept for linear regression
print("Coefficients: {} Intercept: {}".format(lrModel.coefficients,lrModel.intercept))
test_results = lrModel.evaluate(test_data)
# Interesting results....
test_results.residuals.show()
unlabeled_data = test_data.select('features')
predictions = lrModel.transform(unlabeled_data)
predictions.show()
print("RMSE: {}".format(test_results.rootMeanSquaredError))
print("MSE: {}".format(test_results.meanSquaredError))
# Excellent results! Let's see how you handle some more realistically modeled data in the Consulting Project!
# # Data Transformations
#
# You won't always get data in a convienent format, often you will have to deal with data that is non-numerical, such as customer names, or zipcodes, country names, etc...
#
# A big part of working with data is using your own domain knowledge to build an intuition of how to deal with the data, sometimes the best course of action is to drop the data, other times feature-engineering is a good way to go, or you could try to transform the data into something the Machine Learning Algorithms will understand.
#
# Spark has several built in methods of dealing with thse transformations, check them all out here: http://spark.apache.org/docs/latest/ml-features.html
#
# Let's see some examples of all of this!
df = spark.read.csv('data/fake_customers.csv',inferSchema=True,header=True)
df.show()
# ## Data Features
#
# ### StringIndexer
#
# We often have to convert string information into numerical information as a categorical feature. This is easily done with the StringIndexer Method:
# +
from pyspark.ml.feature import StringIndexer
df = spark.createDataFrame(
[(0, "a"), (1, "b"), (2, "c"), (3, "a"), (4, "a"), (5, "c")],
["user_id", "category"])
indexer = StringIndexer(inputCol="category", outputCol="categoryIndex")
indexed = indexer.fit(df).transform(df)
indexed.show()
# -
# The next step would be to encode these categories into "dummy" variables.
# ### VectorIndexer
#
# VectorAssembler is a transformer that combines a given list of columns into a single vector column. It is useful for combining raw features and features generated by different feature transformers into a single feature vector, in order to train ML models like logistic regression and decision trees. VectorAssembler accepts the following input column types: all numeric types, boolean type, and vector type. In each row, the values of the input columns will be concatenated into a vector in the specified order.
#
# Assume that we have a DataFrame with the columns id, hour, mobile, userFeatures, and clicked:
#
# id | hour | mobile | userFeatures | clicked
# ----|------|--------|------------------|---------
# 0 | 18 | 1.0 | [0.0, 10.0, 0.5] | 1.0
#
# userFeatures is a vector column that contains three user features. We want to combine hour, mobile, and userFeatures into a single feature vector called features and use it to predict clicked or not. If we set VectorAssembler’s input columns to hour, mobile, and userFeatures and output column to features, after transformation we should get the following DataFrame:
#
# id | hour | mobile | userFeatures | clicked | features
# ----|------|--------|------------------|---------|-----------------------------
# 0 | 18 | 1.0 | [0.0, 10.0, 0.5] | 1.0 | [18.0, 1.0, 0.0, 10.0, 0.5]
# +
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
dataset = spark.createDataFrame(
[(0, 18, 1.0, Vectors.dense([0.0, 10.0, 0.5]), 1.0)],
["id", "hour", "mobile", "userFeatures", "clicked"])
dataset.show()
# +
assembler = VectorAssembler(
inputCols=["hour", "mobile", "userFeatures"],
outputCol="features")
output = assembler.transform(dataset)
print("Assembled columns 'hour', 'mobile', 'userFeatures' to vector column 'features'")
output.select("features", "clicked").show()
# -
# There ar emany more data transformations available, we will cover them once we encounter a need for them, for now these were the most important ones.
#
# Let's continue on to Linear Regression!
# # Linear Regression Project
# Congratulations! You've been contracted by Hyundai Heavy Industries to help them build a predictive model for some ships. [Hyundai Heavy Industries](http://www.hyundai.eu/en) is one of the world's largest ship manufacturing companies and builds cruise liners.
#
# You've been flown to their headquarters in Ulsan, South Korea to help them give accurate estimates of how many crew members a ship will require.
#
# They are currently building new ships for some customers and want you to create a model and use it to predict how many crew members the ships will need.
#
# Here is what the data looks like so far:
#
# Description: Measurements of ship size, capacity, crew, and age for 158 cruise
# ships.
#
#
# Variables/Columns
# Ship Name 1-20
# Cruise Line 21-40
# Age (as of 2013) 46-48
# Tonnage (1000s of tons) 50-56
# passengers (100s) 58-64
# Length (100s of feet) 66-72
# Cabins (100s) 74-80
# Passenger Density 82-88
# Crew (100s) 90-96
#
# It is saved in a csv file for you called "cruise_ship_info.csv". Your job is to create a regression model that will help predict how many crew members will be needed for future ships. The client also mentioned that they have found that particular cruise lines will differ in acceptable crew counts, so it is most likely an important feature to include in your analysis!
#
df = spark.read.csv('data/cruise_ship_info.csv',inferSchema=True,header=True)
df.printSchema()
df.show()
df.describe().show()
# ## Dealing with the Cruise_line categorical variable
# Ship Name is a useless arbitrary string, but the cruise_line itself may be useful. Let's make it into a categorical variable!
df.groupBy('Cruise_line').count().show()
from pyspark.ml.feature import StringIndexer
indexer = StringIndexer(inputCol="Cruise_line", outputCol="cruise_cat")
indexed = indexer.fit(df).transform(df)
indexed.head(5)
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
indexed.columns
assembler = VectorAssembler(
inputCols=['Age',
'Tonnage',
'passengers',
'length',
'cabins',
'passenger_density',
'cruise_cat'],
outputCol="features")
output = assembler.transform(indexed)
output.select("features", "crew").show()
final_data = output.select("features", "crew")
train_data,test_data = final_data.randomSplit([0.7,0.3])
from pyspark.ml.regression import LinearRegression
# Create a Linear Regression Model object
lr = LinearRegression(labelCol='crew')
# Fit the model to the data and call this model lrModel
lrModel = lr.fit(train_data)
# Print the coefficients and intercept for linear regression
print("Coefficients: {} Intercept: {}".format(lrModel.coefficients,lrModel.intercept))
test_results = lrModel.evaluate(test_data)
print("RMSE: {}".format(test_results.rootMeanSquaredError))
print("MSE: {}".format(test_results.meanSquaredError))
print("R2: {}".format(test_results.r2))
# R2 of 0.86 is pretty good, let's check the data a little closer
from pyspark.sql.functions import corr
df.select(corr('crew','passengers')).show()
df.select(corr('crew','cabins')).show()
# Okay, so maybe it does make sense! Well that is good news for us, this is information we can bring to the company!
#
# ## Stop The Spark Session
# stop the underlying SparkContext.
try:
spark
except NameError:
print("Spark session does not context exist - nothing to stop.")
else:
spark.stop()
| 06_Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# for plotting
import seaborn as sns
import matplotlib.pyplot as plt
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('./data'):
for filename in filenames:
print(os.path.join(dirname, filename))
# Any results you write to the current directory are saved as output.
# -
# <h3>Data Extraction</h3>
#Data Extraction
train_data = pd.read_csv("./data/train.csv")
train_data.head()
train_data.shape
train_data.info()
# <h3>Data Cleaning</h3>
# looking at the correlation of numerical values and identifying data that is not needed
plt.subplots(figsize=(12, 9))
sns.heatmap(train_data.corr(),annot=True)
# +
# we can see that passengerId has almost zero correlation to the passenger surviving
# and to us that makes sense
# we can also see a low correlation between survival and Age, SibSp, and Parch
# which we think should influence the result. so we will not remove them.
del train_data["PassengerId"]
# -
train_data.describe()
# <H3>Data Wrangling</H3>
# <br>Lets Explore more about the ages of the passenger
# <br>This is a continuous numerical field
# <br>we can also see that it has some missing values
# <br>and so, we will try to fill them
#
age_list = train_data["Age"]
age_list.head()
train_data["Age"].fillna(train_data["Age"].mean(), inplace=True)
train_data.describe()
train_data.info()
age_list.describe()
age_list.mean()
age_list.median()
# <H3>Data Analysis</H3>
gr = age_list.plot(kind='density', figsize=(14,6))
gr.axvline(age_list.mean(), color='red')
gr.axvline(age_list.median(), color='green')
# Here we are calling plot via pandas, but this is ultimately being performed by Matplotlib
def plot_density(column):
gr = column.plot(kind='density', figsize=(14,6))
gr.axvline(column.mean(), color='red')
gr.axvline(column.median(), color='green')
gr.set_xlim(column.min(),column.max())
plot_density(train_data["Fare"])
age_list.plot(kind='box', vert=False, figsize=(14,6))
plt.subplots(figsize=(12, 9))
sns.heatmap(train_data.corr(),annot=True)
# +
plt.figure(figsize=(14,6))
survive_color = train_data.Survived
plt.scatter(train_data.Age,train_data.Fare, c = train_data.Survived, alpha = 0.5, cmap = "Spectral")
plt.colorbar()
plt.show()
plt.
# survive_color.head()
# -
# <H3>Lets quickly try to get a prediction model using Sklearn</H3>
# +
from sklearn.ensemble import RandomForestClassifier
test_data = pd.read_csv("./data/test.csv")
y = train_data["Survived"]
features = ["Pclass", "Sex", "SibSp", "Parch"]
X = pd.get_dummies(train_data[features])
X_test = pd.get_dummies(test_data[features])
model = RandomForestClassifier(n_estimators=100, max_depth=5, random_state=1)
model.fit(X, y)
predictions = model.predict(X_test)
output = pd.DataFrame({'PassengerId': test_data.PassengerId, 'Survived': predictions})
output.head()
# -
| Intro/ION Quick Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Venture-Coding/Machine-Learning-under-Kiril-E/blob/main/Model_Selection/K_fold_cross_validation_on_svm_Ads.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="taLo6Q7NxKRE"
# # k-Fold Cross Validation
# + [markdown] id="xQ1Zs9MKxXJ1"
# ## Importing the libraries
# + id="D3x14CwtxcV5"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="1L1wq9evxg83"
# ## Importing the dataset
# + id="TMeZ6FKbxkU6"
dataset = pd.read_csv('Social_Network_Ads.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
# + [markdown] id="i3FBTRwzxzl7"
# ## Splitting the dataset into the Training set and Test set
# + id="V5TZ-0qSx5mX"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 0)
# + [markdown] id="WPt8zMSIxrJ7"
# ## Feature Scaling
# + id="c60y-wPDxvY-"
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + [markdown] id="1dmXtrJbAx9Q"
# Since Age and Salaries, are on completely different scales and age feature might get over-shadowed. Hence, scaling them.
# + [markdown] id="29ry3GSGx9P_"
# ## Training the Kernel SVM model on the Training set
# + id="BUM0MSyGyCjF" colab={"base_uri": "https://localhost:8080/"} outputId="b7e4c028-6218-4ed5-c8fa-fc985eccbed8"
from sklearn.svm import SVC
classifier = SVC(kernel = 'rbf', random_state = 0)
classifier.fit(X_train, y_train)
# + [markdown] id="6MNOjSkOBjzI"
# Why RBF?
#
# Bcuz non-linear separation welcomed. Benefits of KNN but Vectors stored only during training. Not many hyperparameters to tune beside sigma(inversely related to Gamma) and C (for SVM).
# + [markdown] id="fBbur5RQyTcL"
# ## Making the Confusion Matrix
# + id="_1llVHNPyWkZ" colab={"base_uri": "https://localhost:8080/"} outputId="b928ec14-6ab2-4051-bcfe-4cea636b2fcf"
from sklearn.metrics import confusion_matrix, accuracy_score
y_pred = classifier.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy_score(y_test, y_pred)
# + [markdown] id="U-eGh0nuyabx"
# ## Applying k-Fold Cross Validation
# + [markdown] id="7XjycIdVF55G"
# Which is basically creating smaller groups of training set and having a proportionally smaller test set in each group of the training set. Eg. a 1000 long training set will be divided into 10 groups of 100 entries, of which 90 can be used as train and 10 will be used as test.
#
# This way we avoid being lucky on just one big test set that we have separated out earlier during train_test_split, and get an idea of model is performing on unseen data even before going to the test set. (A bit similar to Random Forest's Out Of Bag score)
# + id="N5puHSW9ydyi" colab={"base_uri": "https://localhost:8080/"} outputId="4ed827ce-9932-4243-c096-bd581566d5cc"
accuracies = cross_val_score(estimator = classifier, X = X_train, y = y_train, cv = 10)
print("Accuracy: {:.2f} %".format(accuracies.mean()*100))
print("Standard Deviation: {:.2f} %".format(accuracies.std()*100))
# + [markdown] id="teXfwhMOHe5H"
# cross_val_score gives a "List" of accuracies. We then find the mean of these accs and std dev amongst the various accuracies so found.
# + [markdown] id="M6HUu8z8ygbT"
# ## Visualising the Training set results
# + id="wkjkHN6Qykw3" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="f6095f27-b33f-42c0-d685-3e696a387d5c"
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('yellow', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
color = ListedColormap(('yellow', 'blue'))(i), label = j)
plt.title('Kernel SVM (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# + [markdown] id="UZmvp8dAKONl"
# Using 'color', instead of 'c', in new version, to avoid the warning:
#
# **c** *argument looks like a single numeric RGB or RGBA sequence, which should be avoided as value-mapping will have precedence in case its length matches with* *x* & *y*. Please use the **color** keyword-argument or provide a 2-D array with a single row if you intend to specify the same RGB or RGBA value for all points.
# + [markdown] id="9rLgiI19yn8m"
# ## Visualising the Test set results
# + id="PSuas5JgyrpC" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="8a9d950a-3e61-4f5d-b729-f7982a404edd"
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('yellow', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
color = ListedColormap(('yellow', 'blue'))(i), label = j)
plt.title('Kernel SVM (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# + id="9_dLJPVuFUyb"
| Model_Selection/K_fold_cross_validation_on_svm_Ads.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import re
with open("fra-eng/fra.txt", 'r', encoding='utf-8') as f:
lines = f.read().split('\n')
lines
len(lines)
num_samples = 20000 # Using only 20000 pairs for this example
lines_to_use = lines[: min(num_samples, len(lines) - 1)]
lines_to_use
# +
# remove \u202f
# -
for l in range(len(lines_to_use)):
lines_to_use[l] = re.sub("\u202f", "", lines_to_use[l])
lines_to_use
# +
# Map All digits to a token: 'NUMBER_PRESENT'
# -
for l in range(len(lines_to_use)):
lines_to_use[l] = re.sub("\d", " NUMBER_PRESENT ", lines_to_use[l])
lines_to_use
# +
# Append 'BEGIN_ ' and ' _END' characters to target sequences
# Map words to integers
# -
input_texts = []
target_texts = []
input_words = set()
target_words = set()
for line in lines_to_use:
target_text, input_text = line.split('\t')
target_text = 'BEGIN_ ' + target_text + ' _END'
input_texts.append(input_text)
target_texts.append(target_text)
for word in input_text.split():
if word not in input_words:
input_words.add(word)
for word in target_text.split():
if word not in target_words:
target_words.add(word)
len(input_texts)
len(input_words)
len(target_words)
max_input_seq_length = max([len(i.split()) for i in input_texts])
max_input_seq_length
max_target_seq_length = max([len(i.split()) for i in target_texts])
max_target_seq_length
input_words = sorted(list(input_words))
target_words = sorted(list(target_words))
num_encoder_tokens = len(input_words)
num_decoder_tokens = len(target_words)
input_words
input_token_index = dict(
[(word, i) for i, word in enumerate(input_words)])
target_token_index = dict(
[(word, i) for i, word in enumerate(target_words)])
target_token_index
# +
# define encoder-decoder inputs
# -
encoder_input_data = np.zeros(
(len(input_texts), max_input_seq_length),
dtype='float32')
decoder_input_data = np.zeros(
(len(target_texts), max_target_seq_length),
dtype='float32')
decoder_target_data = np.zeros(
(len(target_texts), max_target_seq_length, num_decoder_tokens),
dtype='float32')
encoder_input_data.shape
decoder_input_data.shape
decoder_target_data.shape
for i, (input_text, target_text) in enumerate(zip(input_texts, target_texts)):
for t, word in enumerate(input_text.split()):
encoder_input_data[i, t] = input_token_index[word]
for t, word in enumerate(target_text.split()):
decoder_input_data[i, t] = target_token_index[word]
if t > 0:
# decoder_target_data is ahead of decoder_input_data by one timestep
decoder_target_data[i, t - 1, target_token_index[word]] = 1.
# # Model
embedding_size = 50
from keras.layers import Input, LSTM, Embedding, Dense
from keras.models import Model
from keras.utils import plot_model
# ## Encoder
encoder_inputs = Input(shape=(None,))
encoder_after_embedding = Embedding(num_encoder_tokens, embedding_size)(encoder_inputs)
encoder_lstm = LSTM(50, return_state=True)
_, state_h, state_c = encoder_lstm(encoder_after_embedding)
encoder_states = [state_h, state_c]
# ## Decoder
decoder_inputs = Input(shape=(None,))
decoder_after_embedding = Embedding(num_decoder_tokens, embedding_size)(decoder_inputs)
decoder_lstm = LSTM(50, return_sequences=True, return_state=True)
decoder_outputs, _, _ = decoder_lstm(decoder_after_embedding,
initial_state=encoder_states)
decoder_dense = Dense(num_decoder_tokens, activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['acc'])
model.summary()
model.fit([encoder_input_data, decoder_input_data], decoder_target_data,
batch_size=128,
epochs=8,
validation_split=0.05)
model.save("fr-eng-model.h5")
# # Inference
encoder_model = Model(encoder_inputs, encoder_states)
encoder_model.summary()
decoder_state_input_h = Input(shape=(50,))
decoder_state_input_c = Input(shape=(50,))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
decoder_outputs_inf, state_h_inf, state_c_inf = decoder_lstm(decoder_after_embedding, initial_state=decoder_states_inputs)
decoder_states_inf = [state_h_inf, state_c_inf]
decoder_outputs_inf = decoder_dense(decoder_outputs_inf)
# Multiple input, multiple output
decoder_model = Model(
[decoder_inputs] + decoder_states_inputs,
[decoder_outputs_inf] + decoder_states_inf)
# +
# Reverse-lookup token index to decode sequences
reverse_input_word_index = dict(
(i, word) for word, i in input_token_index.items())
reverse_target_word_index = dict(
(i, word) for word, i in target_token_index.items())
# -
reverse_input_word_index
def decode_sequence(input_seq):
# encode input as a state vector
states_value = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1,1))
# Populate the first character of target sequence with the start character.
target_seq[0, 0] = target_token_index['BEGIN_']
# Sampling loop for a batch of sequences
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict(
[target_seq] + states_value)
# Sample a token
sampled_token_index = np.argmax(output_tokens) #[0, -1, :]
sampled_word = reverse_target_word_index[sampled_token_index]
decoded_sentence += ' ' + sampled_word
# Exit condition: either hit max length
# or find stop character.
if (sampled_word == '_END' or
len(decoded_sentence) > 60):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1,1))
target_seq[0, 0] = sampled_token_index
# Update states
states_value = [h, c]
return decoded_sentence
for seq_index in [4077,5122,6035,7064, 8056, 9068, 9090, 9095]:
input_seq = encoder_input_data[seq_index: seq_index + 1]
decoded_sentence = decode_sequence(input_seq)
print('-')
print('Input sentence:', input_texts[seq_index: seq_index + 1])
print('Decoded sentence:', decoded_sentence)
# +
# Inference for user input: take in a word sequence, convert
# the sequence word by word into encoded
# -
text_to_translate = "Where is my car?"
# +
encoder_input_to_translate = np.zeros(
(1, max_input_seq_length),
dtype='float32')
for t, word in enumerate(text_to_translate.split()):
encoder_input_to_translate[0, t] = input_token_index[word]
# -
decode_sequence(encoder_input_to_translate)
| Lesson 07/activity/word-level-translator_fr_eng.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p><font size="6"><b>06 - Pandas: Reshaping data</b></font></p>
#
# > *© 2016-2018, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
#
# For an overview, see [Reshaping and pivot tables](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html) in the user guide.
# + run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "-"}
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# # Pivoting data
# ## Cfr. excel
# People who know Excel, probably know the **Pivot** functionality:
# 
# 
# The data of the table:
excelample = pd.DataFrame({'Month': ["January", "January", "January", "January",
"February", "February", "February", "February",
"March", "March", "March", "March"],
'Category': ["Transportation", "Grocery", "Household", "Entertainment",
"Transportation", "Grocery", "Household", "Entertainment",
"Transportation", "Grocery", "Household", "Entertainment"],
'Amount': [74., 235., 175., 100., 115., 240., 225., 125., 90., 260., 200., 120.]})
excelample
# In some cases you may wish to transform your data to a different format: use pandas [**`pivot`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.pivot.html) method.
excelample_pivot = excelample.pivot(index="Category", columns="Month", values="Amount")
excelample_pivot
# Interested in *Grand totals*?
# sum columns
excelample_pivot.sum(axis=1)
# sum rows
excelample_pivot.sum(axis=0)
# ## Pivot is just reordering your data:
# Small subsample of the titanic dataset:
# + run_control={"frozen": false, "read_only": false}
df = pd.DataFrame({'Fare': [7.25, 71.2833, 51.8625, 30.0708, 7.8542, 13.0],
'Pclass': [3, 1, 1, 2, 3, 2],
'Sex': ['male', 'female', 'male', 'female', 'female', 'male'],
'Survived': [0, 1, 0, 1, 0, 1]})
df
# + run_control={"frozen": false, "read_only": false}
df.pivot(index='Pclass', columns='Sex', values='Fare')
# + run_control={"frozen": false, "read_only": false}
df.pivot(index='Pclass', columns='Sex', values='Survived')
# -
# So far, so good...
# Let's now use the full titanic dataset:
# + run_control={"frozen": false, "read_only": false}
df = pd.read_csv("../data/titanic.csv")
# + run_control={"frozen": false, "read_only": false}
df.head()
# -
# And try the same pivot (*no worries about the try-except, this is here just used to catch a loooong error*):
# + run_control={"frozen": false, "read_only": false}
try:
df.pivot(index='Sex', columns='Pclass', values='Fare')
except Exception as e:
print("Exception!", e)
# -
# This does not work, because we would end up with multiple values for one cell of the resulting frame, as the error says: `duplicated` values for the columns in the selection. As an example, consider the following rows of our three columns of interest:
df.loc[[1, 3], ["Sex", 'Pclass', 'Fare']]
# Since `pivot` is just restructering data, where would both values of `Fare` for the same combination of `Sex` and `Pclass` need to go?
#
# Well, they need to be combined, according to an `aggregation` functionality, which is supported by another method: [**`pivot_table`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.pivot_table.html).
# <div class="alert alert-danger">
#
# <b>NOTE</b>:
#
# <ul>
# <li><b>Pivot</b> is purely restructering: a single value for each index/column combination is required.</li>
# </ul>
# </div>
# # Pivot tables - aggregating while pivoting
# + run_control={"frozen": false, "read_only": false}
df = pd.read_csv("../data/titanic.csv")
# + run_control={"frozen": false, "read_only": false}
df.pivot_table(index='Sex', columns='Pclass', values='Fare')
# -
# <div class="alert alert-info">
#
# <b>REMEMBER</b>:
#
# - By default, [`pivot_table`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.pivot_table.html) method takes the **mean** of all values that would end up into one cell. However, you can also specify other aggregation functions using the <code>aggfunc</code> keyword.
#
# </div>
# + run_control={"frozen": false, "read_only": false}
df.pivot_table(index='Sex', columns='Pclass', values='Fare', aggfunc='max')
# + run_control={"frozen": false, "read_only": false}
df.pivot_table(index='Sex', columns='Pclass', values='Fare', aggfunc='count')
# -
# <div class="alert alert-info">
#
# <b>REMEMBER</b>:
#
# - There is a shortcut function for a `pivot_table` with a `aggfunc='count'` as aggregation: [**`crosstab`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.crosstab.html)
#
# </div>
pd.crosstab(index=df['Sex'], columns=df['Pclass'])
# + [markdown] clear_cell=false
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Make a pivot table with the survival rates for Pclass vs Sex.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data1.py
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data2.py
# + [markdown] clear_cell=false
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Make a table of the median Fare payed by aged/underaged vs Sex.</li>
# </ul>
# </div>
# + clear_cell=true
# # %load _solutions/pandas_07_reshaping_data3.py
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data4.py
# -
# # Melt - from pivot table to long or tidy format
# The [**`melt`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) function performs the inverse operation of a [`pivot`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.pivot.html). This can be used to make your frame longer, i.e. to make a *tidy* version of your data.
pivoted = df.pivot_table(index='Sex', columns='Pclass', values='Fare').reset_index()
pivoted.columns.name = None
pivoted
# Assume we have a DataFrame like the above. The observations (the average Fare people payed) are spread over different columns. In a tidy dataset, each observation is stored in one row. To obtain this, we can use the [**`melt`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) function:
pd.melt(pivoted)
# As you can see above, the `melt` function puts all column labels in one column, and all values in a second column.
#
# In this case, this is not fully what we want. We would like to keep the 'Sex' column separately:
pd.melt(pivoted, id_vars=['Sex']) #, var_name='Pclass', value_name='Fare')
# # Reshaping with `stack` and `unstack`
# + [markdown] slideshow={"slide_type": "fragment"}
# The actions of the functions we learnt about above, `pivot` and `melt`, can actually be replicated with [**`stack`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.stack.html) and [**`unstack`**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.unstack.html) and `set_index`/`reset_index`. You can think of `pivot` as a special case of using `stack`; and `melt` as a special case of using `unstack`.
#
# Indeed...
# <img src="../img/schema-stack.svg" width=50%>
#
# Let's check it in practice on the following dummy example:
# + run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "subslide"}
df = pd.DataFrame({'A':['one', 'one', 'two', 'two'],
'B':['a', 'b', 'a', 'b'],
'C':range(4)})
df
# -
# To use [`stack`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.stack.html)/[`unstack`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.unstack.html), we need the values we want to shift from rows to columns or the other way around as the index:
# + run_control={"frozen": false, "read_only": false}
df = df.set_index(['A', 'B']) # Indeed, you can combine two indices
df
# + run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "subslide"}
result = df['C'].unstack()
result
# + run_control={"frozen": false, "read_only": false}
df = result.stack().reset_index(name='C')
df
# + [markdown] run_control={"frozen": false, "read_only": false}
# <div class="alert alert-info">
#
# <b>REMEMBER</b>:
#
# - [**stack**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.stack.html): make your data *longer* and *narrower*
# - [**unstack**](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.unstack.html): make your data *shorter* and *wider*
# </div>
# -
# ## Mimick pivot table
# To better understand and reason about pivot tables, we can express this method as a combination of more basic steps. In short, the pivot is a convenient way of expressing the combination of a `groupby` and `stack/unstack`.
# + run_control={"frozen": false, "read_only": false}
df = pd.read_csv("../data/titanic.csv")
# + run_control={"frozen": false, "read_only": false}
df.head()
# + run_control={"frozen": false, "read_only": false}
df.pivot_table(index='Pclass', columns='Sex', values='Survived', aggfunc='mean')
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Get the same result as above based on a combination of `groupby` and `unstack`</li>
# <li>First use `groupby` to calculate the survival ratio for all groups`unstack`</li>
# <li>Then, use `unstack` to reshape the output of the groupby operation</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data5.py
# -
# # [OPTIONAL] Exercises: use the reshaping methods with the movie data
# These exercises are based on the [PyCon tutorial of <NAME>](https://github.com/brandon-rhodes/pycon-pandas-tutorial/) (so credit to him!) and the datasets he prepared for that. You can download these data from here: [`titles.csv`](https://drive.google.com/open?id=0B3G70MlBnCgKajNMa1pfSzN6Q3M) and [`cast.csv`](https://drive.google.com/open?id=0B3G70MlBnCgKal9UYTJSR2ZhSW8) and put them in the `/data` folder.
# + run_control={"frozen": false, "read_only": false}
cast = pd.read_csv('../data/cast.csv')
cast.head()
# + run_control={"frozen": false, "read_only": false}
titles = pd.read_csv('../data/titles.csv')
titles.head()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Plot the number of actor roles each year and the number of actress roles each year over the whole period of available movie data.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data6.py
# + clear_cell=true
# # %load _solutions/pandas_07_reshaping_data7.py
# + clear_cell=true
# # %load _solutions/pandas_07_reshaping_data8.py
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Plot the number of actor roles each year and the number of actress roles each year. Use kind='area' as plot type</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data9.py
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Plot the fraction of roles that have been 'actor' roles each year over the whole period of available movie data.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data10.py
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Define a year as a "Superman year" when films of that year feature more Superman characters than Batman characters. How many years in film history have been Superman years?</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data11.py
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_07_reshaping_data12.py
| Day_1_Scientific_Python/pandas/pandas_06_reshaping_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Austrian energy system Tutorial Part 4: Investigating Many Policy Scenarios
#
# **Pre-requisites**
# - You have the *MESSAGEix* framework installed and working
# - You have run Austrian energy system baseline scenario (``austria.ipynb``) and solved it successfully
# - You have completed tht tutorial on introducing one policy scenario (``austria_single_policy.ipynb``)
#
# **Answers**
# - Answers to the exercises below can be found in another Jupyter Notebook in this folder called ``austria_multiple_policies-answers.ipynb``
#
# **Introduction**
#
# In this notebook, we investigate a number of different scenarios. This has been streamlined with a few helper functions:
#
# - `make_scenario()`: provides a `Scenario` in which to develop the new scenario
# - `read_scenario()`: provides a `Scenario` use for investigating results
#
# These functions are used as follows:
#
# ```
# with function(<arguments>) as ds:
# <do things with the datastructure
# ```
#
# All of the opening, closing, running, and committing actions are handled for you. Your job is to concentrate on identifying and updating the scenario variables and then investigating the results.
#
# You can, of course, get the `Scenario` as you have worked on previously as well.
# ## Setup and Helper Variables
# +
# load required packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
import ixmp as ix
import message_ix
from tools import Plots, make_scenario, read_scenario
# -
# launch the IX modeling platform using the local default database
mp = ix.Platform()
# +
country = 'Austria'
horizon = range(2010, 2041, 10)
plants = [
"coal_ppl",
"gas_ppl",
"oil_ppl",
"bio_ppl",
"hydro_ppl",
"wind_ppl",
"solar_pv_ppl", # actually primary -> final
]
lights = [
"bulb",
"cfl",
]
name = "Austrian energy model"
base_scen = "baseline"
# -
# # Wind Subsidies
#
# Rerun the wind subsidy scenario using this framework.
# +
new_scen = 'wind_subsidies'
subsidies = np.array([0.5, 0.5, 0.75, 1.0])
inv_cost = pd.DataFrame({
'node_loc': country,
'year_vtg': horizon,
'technology': 'wind_ppl',
'value': 1100 * subsidies * 1e6,
'unit': 'USD/GWa',
})
with make_scenario(mp, country, name, base_scen, new_scen) as scenario:
scenario.add_par('inv_cost', inv_cost)
# -
with read_scenario(mp, name, new_scen) as scenario, read_scenario(mp, name, base_scen) as base:
Plots(base, country).plot_new_capacity(baseyear=True, subset=plants)
Plots(scenario, country).plot_new_capacity(baseyear=True, subset=plants)
# # Demand-Side Learning
#
# This model does not use `cfl`s in the basline because they are too expensive. What happens if their cost reduces with time?
# +
new_scen = 'cheap_cfls'
costs = np.array([1.0, 0.6, 0.3, 0.1])
inv_cost = pd.DataFrame({
'node_loc': country,
'year_vtg': horizon,
'technology': 'cfl',
'value': 900 * costs * 1e6,
'unit': 'USD/GWa',
})
with make_scenario(mp, country, name, base_scen, new_scen) as scenario:
scenario.add_par('inv_cost', inv_cost)
# -
with read_scenario(mp, name, new_scen) as scenario, read_scenario(mp, name, base_scen) as base:
Plots(base, country).plot_new_capacity(baseyear=True, subset=lights)
Plots(scenario, country).plot_new_capacity(baseyear=True, subset=lights)
with read_scenario(mp, name, new_scen) as scenario, read_scenario(mp, name, base_scen) as base:
Plots(base, country).plot_activity(baseyear=True, subset=lights)
Plots(scenario, country).plot_activity(baseyear=True, subset=lights)
# # Exercise: Economic Assumptions
#
# What is the effect of assuming a different interest rate? What if it is higher than the baseline? Lower? How does this affect prices?
# # Exercise: Carbon Tax
#
# What effect does a carbon tax have on the system? What if it is phased in over time? What is the effect on energy prices?
#
# Hints:
#
# - what emissions parameters are available from `scenario.par_list()`?
# - find out which fields are required using `scenario.idx_names(par_name)`
# - carbon taxes are normally provided in units of USD/tCO2
# - a normal proposed carbon tax is ~30 USD/tCO2
| tutorial/Austrian_energy_system/austria_multiple_policies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit (conda)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# Dataframe HPRD new
df_hprd = pd.read_csv('hprd_new.tsv', sep='\t', header=None)
df_hprd.columns = ['id1', 'id2', 'label']
df_hprd
# set protein in HPRD new
set_hprd = set(df_hprd['id1']).union(df_hprd['id2'])
f = open('set_hprd.txt', 'w')
for id in set_hprd:
f.write(id+'\n')
f.close()
len(set_hprd)
# +
# Submit to Uniprot database to obtain fasta file then submit to CD-HIT 40
# -
# Dataframe of id, seq, len after submitting to CD-HIT40
df_hprd_id_seq_cd40 = pd.read_csv('hprd_new_cd40.tsv', header=None, sep='\t')
df_hprd_id_seq_cd40.columns = ['id', 'seq']
df_hprd_id_seq_cd40['len'] = [len(seq) for seq in df_hprd_id_seq_cd40['seq']]
df_hprd_id_seq_cd40
# Set of hprd cd40 short id
df_hprd_id_seq_cd40_short = df_hprd_id_seq_cd40[df_hprd_id_seq_cd40['len'] < 1200]
set_hprd_cd40_short = set(df_hprd_id_seq_cd40_short['id'])
set_hprd_cd40_short
# +
def mapid(id, set_id):
if id in set_id:
return id
else:
return 0
df_hprd_cd40_short = df_hprd.applymap(lambda x: mapid(x, set_hprd_cd40_short))
df_hprd_cd40_short['label'] = df_hprd['label']
df_hprd_cd40_short
# -
df_hprd_cd40_short = df_hprd_cd40_short[(df_hprd_cd40_short['id1'] != 0) & (df_hprd_cd40_short['id2'] != 0)].reset_index(drop=True)
df_hprd_cd40_short.to_csv('hprd_new_cd40_short.tsv', sep='\t', header=None, index=None)
df_hprd_cd40_short
| data/Independent-testsets/Human-sets/HPRD-2010-new/Non-redundant/prep_non_redundant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # (Temporary) Notebook that Compares Quant results to the SIMs that Mark created
# +
import multiprocessing as mp
import numpy as np
#import multiprocess as mp # A fork of multiprocessing that uses dill rather than pickle
import yaml # pyyaml library for reading the parameters.yml file
import os
import matplotlib.pyplot as plt
from microsim.opencl.ramp.run import run_headless
from microsim.opencl.ramp.snapshot_convertor import SnapshotConvertor
from microsim.opencl.ramp.snapshot import Snapshot
from microsim.opencl.ramp.params import Params, IndividualHazardMultipliers, LocationHazardMultipliers
from microsim.opencl.ramp.simulator import Simulator
from microsim.opencl.ramp.disease_statuses import DiseaseStatus
import functions # Some additional notebook-specific functions required (functions.py)
# Useful for connecting to this kernel
# %connect_info
# -
# ## Code to make plots
#
# This is taken from `sensitivity_analysis.ipynb`
# +
def plot_summaries(summaries, plot_type="error_bars"):
#fig, ax = plt.subplots(1, len(DiseaseStatus), sharey=True)
fig, ax = plt.subplots(1, 1, figsize=(10,7))
# Work out the number of repetitions and iterations
iters, reps = _get_iters_and_reps(summaries)
x = range(iters)
for d, disease_status in enumerate(DiseaseStatus):
if disease_status==DiseaseStatus.Susceptible or disease_status==DiseaseStatus.Recovered:
continue
# Calculate the mean and standard deviation
matrix = np.zeros(shape=(reps,iters))
for rep in range(reps):
matrix[rep] = summaries[rep].total_counts[d]
mean = np.mean(matrix, axis=0)
sd = np.std(matrix, axis=0)
if plot_type == "error_bars":
ax.errorbar(x, mean, sd, label=f"{disease_status}" )
elif plot_type == "lines":
for rep in range(reps):
ax.plot(x, matrix[rep], label=f"{disease_status} {rep}",
color=plt.cm.get_cmap("hsv", len(DiseaseStatus))(d) )
ax.legend()
ax.set_title("Disease Status")
ax.set_xlabel("Iteration")
ax.set_ylabel("Number of cases")
def _get_iters_and_reps(summaries):
reps = len(summaries)
iters = len(summaries[0].total_counts[0])
return (iters, reps)
def plot_disease_status_by_age(summaries):
#fig, ax = plt.subplots(1, len(DiseaseStatus), sharey=True)
fig, ax = plt.subplots(int(len(DiseaseStatus)/2), int(len(DiseaseStatus)/2),
figsize=(15,11), tight_layout=True)
iters, reps = _get_iters_and_reps(summaries)
x = range(iters)
age_thresholds = summaries[0].age_thresholds
for d, disease_status in enumerate(DiseaseStatus):
lower_age_bound = 0
for age_idx in range(len(age_thresholds)):
matrix = np.zeros(shape=(reps, iters))
for rep in range(reps):
#matrix[age_idx][rep][it] = summaries[rep].age_counts[str(disease_status)][age_idx][it]
matrix[rep] = summaries[rep].age_counts[str(disease_status)][age_idx]
mean = np.mean(matrix, axis=0)
sd = np.std(matrix, axis=0)
ax.flat[d].errorbar(x, mean, sd, label=f"{lower_age_bound} - {age_thresholds[age_idx]}" )
lower_age_bound = age_thresholds[age_idx]
ax.flat[d].legend()
ax.flat[d].set_title(f"{str(disease_status)}")
ax.flat[d].set_xlabel("Iteration")
ax.flat[d].set_ylabel("Number of cases")
#fig.set_title(f"Num {disease_status} people by age group")
# -
# ## 1. Traditional run Setup params for all runs
# Run the model using non-quant data. Note that the model needs to have been at least initialised previously to create the snapshots below
# +
USE_QUANT = "no-quant" # CHANGE THIS TO USE QUANT ('quant') OR NOT ('no-quant')
PARAMETERS_FILENAME = f"default-{USE_QUANT}.yml"
with open(os.path.join("..","model_parameters", PARAMETERS_FILENAME)) as f:
parameters = yaml.load(f, Loader=yaml.SafeLoader)
sim_params = parameters["microsim"] # Parameters for the dynamic microsim (python)
calibration_params = parameters["microsim_calibration"]
disease_params = parameters["disease"] # Parameters for the disease model (r)
# This is a snapshot created earlier
opencl_dir = f"../microsim/opencl/"
snapshot_filepath = os.path.join(opencl_dir, "snapshots", USE_QUANT, "cache.npz")
current_risk_beta = disease_params['current_risk_beta']
# The OpenCL model incorporates the current risk beta by pre-multiplying the hazard multipliers with it
location_hazard_multipliers = LocationHazardMultipliers(
retail=calibration_params["hazard_location_multipliers"]["Retail"] * current_risk_beta,
primary_school=calibration_params["hazard_location_multipliers"]["PrimarySchool"] * current_risk_beta,
secondary_school=calibration_params["hazard_location_multipliers"]["SecondarySchool"] * current_risk_beta,
home=calibration_params["hazard_location_multipliers"]["Home"] * current_risk_beta,
work=calibration_params["hazard_location_multipliers"]["Work"] * current_risk_beta,
)
# Individual hazard multipliers can be passed straight through
individual_hazard_multipliers = IndividualHazardMultipliers(
presymptomatic=calibration_params["hazard_individual_multipliers"]["presymptomatic"],
asymptomatic=calibration_params["hazard_individual_multipliers"]["asymptomatic"],
symptomatic=calibration_params["hazard_individual_multipliers"]["symptomatic"]
)
proportion_asymptomatic = disease_params["asymp_rate"]
params = Params(
location_hazard_multipliers=location_hazard_multipliers,
individual_hazard_multipliers=individual_hazard_multipliers,
proportion_asymptomatic=proportion_asymptomatic
)
# -
# Run the model
# +
iterations = 120
repetitions = 20
num_seed_days = 10
use_gpu=False
# Prepare the function arguments as lists for starmap
l_i = [i for i in range(repetitions)]
l_iterations = [iterations] * repetitions
l_snapshot_filepath = [snapshot_filepath] * repetitions
l_params = [params] * repetitions
l_opencl_dir = [opencl_dir] * repetitions
l_num_seed_days = [num_seed_days] * repetitions
l_use_gpu = [use_gpu] * repetitions
l_store_detailed_counts= [True] * repetitions # If False then model is much quicker but no age breakdown
#results = functions.run_opencl_model_multiprocess(
# l_i, l_iterations, l_snapshot_filepath, l_params, l_opencl_dir, l_num_seed_days, l_use_gpu)
import itertools # (only while I can't get multiprocessing to work)
try:
with mp.Pool(processes=int(os.cpu_count())) as pool:
#results = pool.starmap(
results = itertools.starmap(
functions._run_opencl_model, zip(
l_i, l_iterations, l_snapshot_filepath, l_params, l_opencl_dir, l_num_seed_days, l_use_gpu, l_store_detailed_counts
))
finally: # Make sure they get closed (shouldn't be necessary)
pool.close()
summaries_no_quant = [x[0] for x in results]
final_results = [x[1] for x in results]
# -
# ### Plot MSOA geodata
# +
from microsim.load_msoa_locations import load_osm_shapefile, load_msoa_shapes
import pandas as pd
data_dir = ("../devon_data")
osm_buildings = load_osm_shapefile(data_dir)
devon_msoa_shapes = load_msoa_shapes(data_dir, visualize=False)
devon_msoa_shapes.plot()
plt.show()
# +
import pandas as pd
def plot_msoa_choropleth(msoa_shapes, summary, disease_status, timestep):
# get dataframes for all statuses
msoa_data = summary.get_area_dataframes()
msoa_data_for_status = msoa_data[disease_status]
# add "Code" column so dataframes can be merged
msoa_data_for_status["Code"] = msoa_data_for_status.index
msoa_shapes = pd.merge(msoa_shapes, msoa_data_for_status, on="Code")
msoa_shapes.plot(column=f"Day{timestep}", legend=True)
plt.show()
# -
# ## 2. Quant run
# Run the model using non-quant data. Note that the model needs to have been at least initialised previously to create the snapshots below
# +
USE_QUANT = "quant" # CHANGE THIS TO USE QUANT ('quant') OR NOT ('no-quant')
PARAMETERS_FILENAME = f"default-{USE_QUANT}.yml"
with open(os.path.join("..","model_parameters", PARAMETERS_FILENAME)) as f:
parameters = yaml.load(f, Loader=yaml.SafeLoader)
sim_params = parameters["microsim"] # Parameters for the dynamic microsim (python)
calibration_params = parameters["microsim_calibration"]
disease_params = parameters["disease"] # Parameters for the disease model (r)
# This is a snapshot created earlier
opencl_dir = f"../microsim/opencl/"
snapshot_filepath = os.path.join(opencl_dir, "snapshots", USE_QUANT, "cache.npz")
current_risk_beta = disease_params['current_risk_beta']
# The OpenCL model incorporates the current risk beta by pre-multiplying the hazard multipliers with it
location_hazard_multipliers = LocationHazardMultipliers(
retail=calibration_params["hazard_location_multipliers"]["Retail"] * current_risk_beta,
primary_school=calibration_params["hazard_location_multipliers"]["PrimarySchool"] * current_risk_beta,
secondary_school=calibration_params["hazard_location_multipliers"]["SecondarySchool"] * current_risk_beta,
home=calibration_params["hazard_location_multipliers"]["Home"] * current_risk_beta,
work=calibration_params["hazard_location_multipliers"]["Work"] * current_risk_beta,
)
# Individual hazard multipliers can be passed straight through
individual_hazard_multipliers = IndividualHazardMultipliers(
presymptomatic=calibration_params["hazard_individual_multipliers"]["presymptomatic"],
asymptomatic=calibration_params["hazard_individual_multipliers"]["asymptomatic"],
symptomatic=calibration_params["hazard_individual_multipliers"]["symptomatic"]
)
proportion_asymptomatic = disease_params["asymp_rate"]
params = Params(
location_hazard_multipliers=location_hazard_multipliers,
individual_hazard_multipliers=individual_hazard_multipliers,
proportion_asymptomatic=proportion_asymptomatic
)
# -
# Run the model
# +
# Prepare the function arguments as lists for starmap
l_i = [i for i in range(repetitions)]
l_iterations = [iterations] * repetitions
l_snapshot_filepath = [snapshot_filepath] * repetitions
l_params = [params] * repetitions
l_opencl_dir = [opencl_dir] * repetitions
l_num_seed_days = [num_seed_days] * repetitions
l_use_gpu = [use_gpu] * repetitions
l_store_detailed_counts= [True] * repetitions # If False then model is much quicker but no age breakdown
#results = functions.run_opencl_model_multiprocess(
# l_i, l_iterations, l_snapshot_filepath, l_params, l_opencl_dir, l_num_seed_days, l_use_gpu)
import itertools # (only while I can't get multiprocessing to work)
try:
with mp.Pool(processes=int(os.cpu_count())) as pool:
#results = pool.starmap(
results = itertools.starmap(
functions._run_opencl_model, zip(
l_i, l_iterations, l_snapshot_filepath, l_params, l_opencl_dir, l_num_seed_days, l_use_gpu, l_store_detailed_counts
))
finally: # Make sure they get closed (shouldn't be necessary)
pool.close()
summaries_quant = [x[0] for x in results]
final_results = [x[1] for x in results]
# -
# ## Compare the Quant / Non-Quant (separate plots)
plot_summaries(summaries=summaries_no_quant, plot_type="error_bars")
plot_summaries(summaries=summaries_quant, plot_type="error_bars")
plot_disease_status_by_age(summaries_no_quant)
plot_disease_status_by_age(summaries_quant)
# Look at maps
disease_status = "exposed"
plot_msoa_choropleth(devon_msoa_shapes, summaries_no_quant[0], disease_status, 99)
plot_msoa_choropleth(devon_msoa_shapes, summaries_quant[0], disease_status, 99)
# ## Compare Quant / No_Quant on the same plots
# +
def plot_summaries_diff(summaries1, summaries2):
#fig, ax = plt.subplots(1, len(DiseaseStatus), sharey=True)
fig, ax = plt.subplots(1, 1, figsize=(10,7))
# Work out the number of repetitions and iterations
iters, reps = _get_iters_and_reps(summaries1)
x = range(iters)
for d, disease_status in enumerate(DiseaseStatus):
if disease_status==DiseaseStatus.Susceptible or disease_status==DiseaseStatus.Recovered:
continue
# Calculate the mean and standard deviation
matrix = np.zeros(shape=(reps,iters))
for rep in range(reps):
matrix[rep] = summaries1[rep].total_counts[d] - summaries2[rep].total_counts[d]
mean = np.mean(matrix, axis=0)
sd = np.std(matrix, axis=0)
ax.errorbar(x, mean, sd, label=f"{disease_status}" )
ax.legend()
ax.set_title("Disease Status")
ax.set_xlabel("Iteration")
ax.set_ylabel("Number of cases")
def _get_iters_and_reps(summaries):
reps = len(summaries)
iters = len(summaries[0].total_counts[0])
return (iters, reps)
def plot_disease_status_by_age_diff(summaries1, summaries2):
#fig, ax = plt.subplots(1, len(DiseaseStatus), sharey=True)
fig, ax = plt.subplots(int(len(DiseaseStatus)/2), int(len(DiseaseStatus)/2),
figsize=(15,11), tight_layout=True)
iters, reps = _get_iters_and_reps(summaries1)
assert (iters, reps) == _get_iters_and_reps(summaries2)
x = range(iters)
age_thresholds = summaries1[0].age_thresholds
for d, disease_status in enumerate(DiseaseStatus):
lower_age_bound = 0
for age_idx in range(len(age_thresholds)):
matrix = np.zeros(shape=(reps, iters))
for rep in range(reps):
#matrix[age_idx][rep][it] = summaries[rep].age_counts[str(disease_status)][age_idx][it]
matrix[rep] = summaries1[rep].age_counts[str(disease_status)][age_idx] - summaries2[rep].age_counts[str(disease_status)][age_idx]
mean = np.mean(matrix, axis=0)
sd = np.std(matrix, axis=0)
ax.flat[d].errorbar(x, mean, sd, label=f"{lower_age_bound} - {age_thresholds[age_idx]}" )
lower_age_bound = age_thresholds[age_idx]
ax.flat[d].legend()
ax.flat[d].set_title(f"{str(disease_status)}")
ax.flat[d].set_xlabel("Iteration")
ax.flat[d].set_ylabel("Number of cases")
#fig.set_title(f"Num {disease_status} people by age group")
# -
plot_summaries_diff(summaries_no_quant, summaries_quant)
plot_disease_status_by_age_diff(summaries_no_quant, summaries_quant)
| experiments/test_quant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tca_traff
# language: python
# name: tca_traff
# ---
# +
import numpy as np
import pandas as pd
import scipy
import scipy.linalg
import matplotlib.pyplot as plt
import sklearn.metrics
import sklearn.neighbors
import time
import torch
from torch import nn
import torch.nn.functional as F
from torch.utils.data import DataLoader, Dataset, TensorDataset
import ipdb
import bda_utils
# -
bda_utils.setup_seed(10)
# # 1. BDA Part
# ## 1.a. Define BDA methodology
# +
def kernel(ker, X1, X2, gamma):
K = None
if not ker or ker == 'primal':
K = X1
elif ker == 'linear':
if X2 is not None:
K = sklearn.metrics.pairwise.linear_kernel(
np.asarray(X1).T, np.asarray(X2).T)
else:
K = sklearn.metrics.pairwise.linear_kernel(np.asarray(X1).T)
elif ker == 'rbf':
if X2 is not None:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, np.asarray(X2).T, gamma)
else:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, None, gamma)
return K
def proxy_a_distance(source_X, target_X):
"""
Compute the Proxy-A-Distance of a source/target representation
"""
nb_source = np.shape(source_X)[0]
nb_target = np.shape(target_X)[0]
train_X = np.vstack((source_X, target_X))
train_Y = np.hstack((np.zeros(nb_source, dtype=int),
np.ones(nb_target, dtype=int)))
clf = svm.LinearSVC(random_state=0)
clf.fit(train_X, train_Y)
y_pred = clf.predict(train_X)
error = metrics.mean_absolute_error(train_Y, y_pred)
dist = 2 * (1 - 2 * error)
return dist
def estimate_mu(_X1, _Y1, _X2, _Y2):
adist_m = proxy_a_distance(_X1, _X2)
C = len(np.unique(_Y1))
epsilon = 1e-3
list_adist_c = []
for i in range(1, C + 1):
ind_i, ind_j = np.where(_Y1 == i), np.where(_Y2 == i)
Xsi = _X1[ind_i[0], :]
Xtj = _X2[ind_j[0], :]
adist_i = proxy_a_distance(Xsi, Xtj)
list_adist_c.append(adist_i)
adist_c = sum(list_adist_c) / C
mu = adist_c / (adist_c + adist_m)
if mu > 1:
mu = 1
if mu < epsilon:
mu = 0
return mu
# +
class BDA:
def __init__(self, kernel_type='primal', dim=30, lamb=1, mu=0.5, gamma=1, T=10, mode='BDA', estimate_mu=False):
'''
Init func
:param kernel_type: kernel, values: 'primal' | 'linear' | 'rbf'
:param dim: dimension after transfer
:param lamb: lambda value in equation
:param mu: mu. Default is -1, if not specificied, it calculates using A-distance
:param gamma: kernel bandwidth for rbf kernel
:param T: iteration number
:param mode: 'BDA' | 'WBDA'
:param estimate_mu: True | False, if you want to automatically estimate mu instead of manally set it
'''
self.kernel_type = kernel_type
self.dim = dim
self.lamb = lamb
self.mu = mu
self.gamma = gamma
self.T = T
self.mode = mode
self.estimate_mu = estimate_mu
def fit(self, Xs, Ys, Xt, Yt):
'''
Transform and Predict using 1NN as JDA paper did
:param Xs: ns * n_feature, source feature
:param Ys: ns * 1, source label
:param Xt: nt * n_feature, target feature
:param Yt: nt * 1, target label
:return: acc, y_pred, list_acc
'''
# ipdb.set_trace()
list_acc = []
X = np.hstack((Xs.T, Xt.T)) # X.shape: [n_feature, ns+nt]
X_mean = np.linalg.norm(X, axis=0) # why it's axis=0? the average of features
X_mean[X_mean==0] = 1
X /= X_mean
m, n = X.shape
ns, nt = len(Xs), len(Xt)
e = np.vstack((1 / ns * np.ones((ns, 1)), -1 / nt * np.ones((nt, 1))))
C = np.unique(Ys)
H = np.eye(n) - 1 / n * np.ones((n, n))
mu = self.mu
M = 0
Y_tar_pseudo = None
Xs_new = None
for t in range(self.T):
print('\tStarting iter %i'%t)
N = 0
M0 = e * e.T * len(C)
# ipdb.set_trace()
if Y_tar_pseudo is not None:
for i in range(len(C)):
e = np.zeros((n, 1))
Ns = len(Ys[np.where(Ys == C[i])])
Nt = len(Y_tar_pseudo[np.where(Y_tar_pseudo == C[i])])
# Ns = 1
# Nt = 1
alpha = 1 # bda
tt = Ys == C[i]
e[np.where(tt == True)] = 1 / Ns
# ipdb.set_trace()
yy = Y_tar_pseudo == C[i]
ind = np.where(yy == True)
inds = [item + ns for item in ind]
try:
e[tuple(inds)] = -alpha / Nt
e[np.isinf(e)] = 0
except:
e[tuple(inds)] = 0 # ?
N = N + np.dot(e, e.T)
# ipdb.set_trace()
# In BDA, mu can be set or automatically estimated using A-distance
# In WBDA, we find that setting mu=1 is enough
if self.estimate_mu and self.mode == 'BDA':
if Xs_new is not None:
mu = estimate_mu(Xs_new, Ys, Xt_new, Y_tar_pseudo)
else:
mu = 0
# ipdb.set_trace()
M = (1 - mu) * M0 + mu * N
M /= np.linalg.norm(M, 'fro')
# ipdb.set_trace()
K = kernel(self.kernel_type, X, None, gamma=self.gamma)
n_eye = m if self.kernel_type == 'primal' else n
a, b = np.linalg.multi_dot([K, M, K.T]) + self.lamb * np.eye(n_eye), np.linalg.multi_dot([K, H, K.T])
w, V = scipy.linalg.eig(a, b)
ind = np.argsort(w)
A = V[:, ind[:self.dim]]
Z = np.dot(A.T, K)
Z_mean = np.linalg.norm(Z, axis=0) # why it's axis=0?
Z_mean[Z_mean==0] = 1
Z /= Z_mean
Xs_new, Xt_new = Z[:, :ns].T, Z[:, ns:].T
global device
model = sklearn.svm.SVC(kernel='linear').fit(Xs_new, Ys.ravel())
Y_tar_pseudo = model.predict(Xt_new)
# ipdb.set_trace()
acc = sklearn.metrics.accuracy_score(Y_tar_pseudo, Yt) # Yt is already in classes
print(acc)
return Xs_new, Xt_new, A #, acc, Y_tar_pseudo, list_acc
# -
# ## 1.b. Load Data
Xs, Xt = bda_utils.load_data(if_weekday=1, if_interdet=1)
Xs = Xs[:, :1]
Xt = Xt[:, :1]
Xs, Xs_min, Xs_max = bda_utils.normalize2D(Xs)
Xt, Xt_min, Xt_max = bda_utils.normalize2D(Xt)
for i in range(Xs.shape[1]):
plt.figure(figsize=[20,4])
plt.plot(Xs[:, i])
plt.plot(Xt[:, i])
# ## 1.d. Hyperparameters
# +
label_seq_len = 3
# batch_size = full batch
seq_len = 30
reduced_dim = 4
inp_dim = min(Xs.shape[1], Xt.shape[1])
label_dim = min(Xs.shape[1], Xt.shape[1])
hid_dim = 12
layers = 1
lamb = 3
hyper = {
'inp_dim':inp_dim,
'label_dim':label_dim,
'label_seq_len':label_seq_len,
'seq_len':seq_len,
'reduced_dim':reduced_dim,
'hid_dim':hid_dim,
'layers':layers,
'lamb':lamb}
hyper = pd.DataFrame(hyper, index=['Values'])
# -
hyper
# ## 1.e. Apply BDA and get $Xs_{new}$, $Xt_{new}$
# +
# Xs = Xs[:96, :]
# -
# [sample size, seq_len, inp_dim (dets)], [sample size, label_seq_len, inp_dim (dets)]
Xs_3d, Ys_3d = bda_utils.sliding_window(Xs, Xs, seq_len, 1)
Xt_3d, Yt_3d = bda_utils.sliding_window(Xt, Xt, seq_len, 1)
print(Xs_3d.shape)
print(Ys_3d.shape)
print(Xt_3d.shape)
print(Yt_3d.shape)
# +
t_s = time.time()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
Xs_train_3d = []
Ys_train_3d = []
Xt_valid_3d = []
Xt_train_3d = []
Yt_valid_3d = []
Yt_train_3d = []
for i in range(Xs_3d.shape[2]):
print('Starting det %i'%i)
bda = BDA(kernel_type='linear', dim=seq_len-reduced_dim, lamb=lamb, mu=0.6, gamma=1, T=1) # T is iteration time
Xs_new_raw, Xt_new_raw, A = bda.fit(
Xs_3d[:, :, i], bda_utils.get_class(Ys_3d[:, :, i]), Xt_3d[:, :, i], bda_utils.get_class(Yt_3d[:, :, i])
) # input shape: ns, n_feature | ns, n_label_feature
# normalize
Xs_new, Xs_new_min, Xs_new_max = bda_utils.normalize2D(Xs_new_raw)
Xt_new, Xt_new_min, Xt_new_max = bda_utils.normalize2D(Xt_new_raw)
print(Xs_new.shape)
print(Xt_new.shape)
day_train_t = 1
Xs_train = Xs_new.copy()
Ys_train = Ys_3d[:, :, i]
Xt_valid = Xt_new.copy()[int(96*day_train_t):, :]
Xt_train = Xt_new.copy()[:int(96*day_train_t), :]
Yt_valid = Yt_3d[:, :, i].copy()[int(96*day_train_t):, :]
Yt_train = Yt_3d[:, :, i].copy()[:int(96*day_train_t), :]
Xs_train_3d.append(Xs_train)
Ys_train_3d.append(Ys_train)
Xt_valid_3d.append(Xt_valid)
Xt_train_3d.append(Xt_train)
Yt_valid_3d.append(Yt_valid)
Yt_train_3d.append(Yt_train)
Xs_train_3d = np.array(Xs_train_3d)
Ys_train_3d = np.array(Ys_train_3d)
Xt_valid_3d = np.array(Xt_valid_3d)
Xt_train_3d = np.array(Xt_train_3d)
Yt_valid_3d = np.array(Yt_valid_3d)
Yt_train_3d = np.array(Yt_train_3d)
# bda_utils.save_np(Xs_train_3d, './outputs/BDA/Xs_new_%i.csv'%(bda_utils.get_num()-14/6))
# bda_utils.save_np(Ys_train_3d, './outputs/BDA/Xt_new_%i.csv'%(bda_utils.get_num()-14/6))
# bda_utils.save_np(Xt_valid_3d, './outputs/BDA/Xs_new_%i.csv'%(bda_utils.get_num()-14/6))
# bda_utils.save_np(Xt_train_3d, './outputs/BDA/Xt_new_%i.csv'%(bda_utils.get_num()-14/6))
# bda_utils.save_np(Yt_valid_3d, './outputs/BDA/Xs_new_%i.csv'%(bda_utils.get_num()-14/6))
# bda_utils.save_np(Yt_train_3d, './outputs/BDA/Xt_new_%i.csv'%(bda_utils.get_num()-14/6))
print('Time spent:%.5f'%(time.time()-t_s))
# -
Xs_train_3d = np.transpose(Xs_train_3d, (1, 2, 0))
Ys_train_3d = np.transpose(Ys_train_3d, (1, 2, 0))
Xt_valid_3d = np.transpose(Xt_valid_3d, (1, 2, 0))
Xt_train_3d = np.transpose(Xt_train_3d, (1, 2, 0))
Yt_valid_3d = np.transpose(Yt_valid_3d, (1, 2, 0))
Yt_train_3d = np.transpose(Yt_train_3d, (1, 2, 0))
Xs_train_3d.shape
# # 2. Learning Part
# ## 2.a. Build network
from bda_utils import traff_net_clf
# ## 2.b. Assemble Dataloader
# +
batch_size = 1960
train_x = np.vstack([Xs_train_3d, Xt_train_3d])
train_y = np.vstack([Ys_train_3d, Yt_train_3d])
train_x = torch.tensor(train_x, dtype=torch.float32).to(device)
train_y = torch.tensor(train_y, dtype=torch.float32).to(device)
Xt_valid_3d = torch.tensor(Xt_valid_3d, dtype=torch.float32).to(device)
Yt_valid_3d = torch.tensor(Yt_valid_3d, dtype=torch.float32).to(device)
train_dataset = TensorDataset(train_x, train_y)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size, shuffle=False)
train_iter = iter(train_loader)
print(train_x.shape)
print(train_y.shape)
print('\n')
print(Xt_valid_3d.shape)
print(Yt_valid_3d.shape)
# -
# ## 2.c. Learn
# +
# build model
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
net = traff_net(seq_len - reduced_dim).to(device)
criterion = nn.CrossEntropyLoss()
#scheduler = torch.optim.lr_scheduler.StepLR(optimizer, 0.7)
train_loss_set = []
val_loss_set = []
det = 0 # which detector to visualize
# -
optimizer = torch.optim.SGD(net.parameters(), lr=0.3)
# +
# train
net.train()
epochs = 101
for e in range(epochs):
for i in range(len(train_loader)):
try:
data, label = train_iter.next()
except:
train_iter = iter(train_loader)
data, label = train_iter.next()
# ipdb.set_trace()
out = net(data)
loss = criterion(out, bda_utils.get_class(label[:, 0, 0]).flatten().long() ) # label.shape=[batch, 1, num_dets]
optimizer.zero_grad()
loss.backward()
optimizer.step()
val_out = net(Xt_valid_3d)
val_loss = criterion(val_out, bda_utils.get_class(Yt_valid_3d[:, 0, 0]).flatten().long() )
val_loss_set.append(val_loss.cpu().detach().numpy())
train_loss_set.append(loss.cpu().detach().numpy())
if e%50==0:
# ipdb.set_trace()
fig = plt.figure(figsize=[16,4])
ax1 = fig.add_subplot(111)
ax1.plot(bda_utils.get_class(label)[:, 0, det].cpu().detach().numpy(), label='ground truth')
ax1.plot(torch.argmax(out, dim=1).cpu().detach().numpy(), label='predict')
ax1.legend()
plt.show()
print('Epoch No. %i success, loss: %.5f, val loss: %.5f, acc: %.5f'\
%(e, loss.cpu().detach().numpy(), val_loss.cpu().detach().numpy(), \
sklearn.metrics.accuracy_score(torch.argmax(val_out, dim=1).cpu(), bda_utils.get_class(Yt_valid_3d[:, 0, det]).cpu().flatten())\
))
# -
out
torch.argsort(out, dim=1)[35, :]
out[35, :]
torch.argmax(out, dim=1)[35]
bda_utils.get_class(label).flatten()[35]
# +
# bda_utils.save_np(Xs_new, 'Xs_new.csv')
# -
fig = plt.figure(figsize = [16, 4])
ax1 = fig.add_subplot(121)
ax1.plot(train_loss_set)
ax2 = fig.add_subplot(122)
ax2.plot(val_loss_set)
# # 3. Evaluation
val_out = net(Xt_valid_3d)
plt.figure(figsize=[16,4])
plt.plot(bda_utils.get_class(Yt_valid_3d[:, 0, det]).cpu().flatten(), label='label')
plt.plot(torch.argmax(val_out, dim=1).cpu(), label='predict')
plt.legend()
# +
# sklearn.metrics.accuracy_score(torch.argmax(val_out, dim=1).cpu(), bda_utils.get_class(Yt_valid_3d[:, 0, det]).cpu().flatten())
g_t = torch.argmax(val_out, dim=1).cpu().detach().numpy()
pred = bda_utils.get_class(Yt_valid_3d[:, 0, det]).cpu().flatten().detach().numpy()
print(bda_utils.nrmse_loss_func(g_t, pred, 0))
print(bda_utils.mape_loss_func(g_t, pred, 0))
# -
# +
# torch.save(net.state_dict(), './model/bda_weekday_M1_M4_nrmse=%.5f.pth'%nrmse_loss)
# -
| .ipynb_checkpoints/BDA_clf-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from splinter import Browser
from bs4 import BeautifulSoup as bs
import pandas as pd
import requests
# -
# ## NASA Mars News - Scrape and collect the latest News Title and Paragraph Text
executable_path = {"executable_path": "C:/Users/mike1/OneDrive/Desktop/WebDriver/chromedriver.exe"}
Browser("chrome", **executable_path, headless=False)
# +
url = "https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest"
browser.visit(url)
html = browser.html
soup = bs(html, "html.parser")
news_title = soup.find("div", class_ = "content_title").text
news_p = soup.find("div", class_ = "article_teaser_body").text
# -
print(news_title)
print(news_p)
# ## JPL Mars Space Images - Featured Image
# ## Mars Facts: Use pandas to scrape table from url and convert into HTML table string
url_2 = 'https://space-facts.com/mars/'
tables = pd.read_html(url_2)
tables
# +
mars_df = tables[0]
mars_df.columns = ["Description", "Mars"]
mars_df.set_index("Description", inplace = True)
mars_df
# -
| Missions_to_Mars/.ipynb_checkpoints/Untitled-checkpoint.ipynb |