code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Assignement Details
# Name : **<NAME>** <br>
# Student ID : **L00150445** <br>
# Course : MSc in Big Data Analytics and Artificial Intelligence <br>
# Module : Artificial Intelligence 2 <br>
# File used : Dracula.txt
#
# +
# Importing spaCy Module
import spacy
# Loading the English language library
nlp = spacy.load("en_core_web_sm")
# -
# ## Q1 : Number of sentences
# +
# Reading the dracula text file
text_file = open("Dracula.txt", encoding="utf-8")
entire_text = text_file.read()
# -
# Displaying first 1000 character which have unneccesary line breaks (\n) because conversion from PDF to text file
entire_text [10000:10500]
# +
# Replacing the line breaks with space to mitigate unneccesary line breaks
entire_text = entire_text.replace('\n', ' ')
entire_text [10000:10500]
# -
# **Note** : The '\n' characters is replaced with spaces
# +
# Using NLP creating the doc_object
doc_object = nlp(entire_text)
### stripping the sentences
sentences = [sent.string.strip() for sent in doc_object.sents]
# -
# **Note**: The doc_object creation is part of Q2 also
# ### Answer
print ( 'The number of sentences in the text file is ' + str(len(sentences)))
#
# ### Q 2 : POS Tags
##Taking a random sentance
sentences[200]
# +
def show_noun_chunks(sentence):
doc_object = nlp(sentence)
print(f" {'Token':{10}} {'POS Tag':{10}} {'Token Dependency':{20}} {'Explanation':{30}} {'Stop word'}")
for token in doc_object:
print(f" {str(token):{10}} {token.pos_:{10}} {token.dep_:{20}} {spacy.explain(token.pos_):{30}} {nlp.vocab[token.text].is_stop}")
# -
# ### Answer
show_noun_chunks(sentences[200])
# ### Q3: Regular Expression
# Regular expression is representation style for sequence of characters which is used for pattern matching in a text based searching. It allows the programmers to search for patterns like phone number, area code etc.
#
# +
import re
## searching for dates avaialble. search methods returns the position of the first match
search_pattern = r'\d\d:\d\d'
time = re.search(search_pattern, entire_text)
print('The first time : ' + str(time))
## The findall funcctions returns al the matches in the text
search_pattern = r'\d\d:\d\d'
time = re.findall(search_pattern, entire_text)
time
# -
# ### Answer
#
# The count of each characters mentioned is counted using the regular expression
# +
search_patterns = [r'Godalming', r'Morris', r'Helsing' , r'Lucy' ]
for pattern in search_patterns:
occurance = re.findall(pattern, entire_text)
print ('The character' , pattern , 'mentioned ' , len(occurance) , 'times')
# -
# ### Q4: POS Frequency
#
# POS taggging represents Part of Speech, where the we can assign the POS tags using the nlp() function of the spacy library. The spacy will assign tags such as nouns, verbs etc to each token present in the text.
POS_frequency = doc_object.count_by(spacy.attrs.POS)
# Using the ".items()" command accesses each item in the dictionary
print(f" {'Tag Id':>{10}} {'Token Type':{15}} {'Token Explanation':{30}} {'Occurrences':>10} ")
for tag_id, occurrences in POS_frequency.items():
print(f" {tag_id:>{10}} {doc_object.vocab[tag_id].text:{15}} {spacy.explain(doc_object.vocab[tag_id].text):{30}} {occurrences: >{10}}")
# ### Q5 Rule based matching
# Import the Matcher library
from spacy.matcher import Matcher
matcher = Matcher(nlp.vocab)
# Four sets of patterns is searched: <br>
# 1) long <br>
# 2) remember <br>
# 3) tonight <br>
# 4) policestation <br>
# +
# match for "longer"
token_match1 = [{"LOWER": "longer"}]
# match for "long"
token_match2 = [{"LOWER": "long"}]
# match for "longest"
token_match3 = [{"LOWER": "longest"}]
matcher.add("long", None, token_match1, token_match2, token_match3 )
# +
# match for "remember"
token_match1 = [{"LOWER": "remember"}]
# match for "remembered"
token_match2 = [{"LOWER": "remembered"}]
# match for "remembering"
token_match3 = [{"LOWER": "remembering"}]
matcher.add("remember", None, token_match1, token_match2, token_match3 )
# +
# match for "tonight"
token_match1 = [{"LOWER": "tonight"}]
# match for "tonight"
token_match3 = [{"LOWER": "to"}, {"IS_PUNCT": True}, {"LOWER": "night"}]
matcher.add("tonight", None, token_match1, token_match2, token_match3 )
# +
# match for "stopword"
token_match1 = [{"LOWER": "police station"}]
# match for "stopwords"
token_match2 = [{"LOWER": "policestation"}]
# match for stop-word
token_match3 = [{"LOWER": "police"}, {"IS_PUNCT": True}, {"LOWER": "station"}]
matcher.add("policestation", None, token_match1, token_match2, token_match3 )
# -
token_matches = matcher(doc_object)
# +
def find_matches(text):
pd = [];
# find all matches within the doc object
token_matches = matcher(text)
# For each item in the token_matches provide the following
# match_id is the hash value of the identified token match
print(f"{'Match ID':<{30}} {'String':<{15}} {'Start':{5}} {'End':{5}} {'Match word':{20}}")
for match_id, start, end in token_matches:
string_id = nlp.vocab.strings[match_id]
pd.append(string_id)
matched_span = doc_object[start:end]
print(f"{match_id:<{30}} {string_id:<{15}} {start:{5}} {end:{5}} {matched_span.text:{20}}")
return pd
# -
counter = find_matches(doc_object)
print ("The pattern 'long' has occured :" + str(counter.count('long')))
print ("The pattern 'remember' has occured :" + str(counter.count('remember')))
print ("The pattern 'tonight' has occured :" + str(counter.count('tonight')))
print ("The pattern 'policestation' has occured :" + str(counter.count('policestation')))
# ### Q6: Previous 5 words and next 5 words
#
# The previous 5 words and next words are appended to the output of the Q5
def prev_next_words(text):
# find all matches within the doc object
token_matches = matcher(text)
# For each item in the token_matches provide the following
# match_id is the hash value of the identified token match
print(f"{'Match ID':<{30}} {'String':<{15}} {'Start':{5}} {'End':{5}} {'Match word':{5}}")
for match_id, start, end in token_matches:
string_id = nlp.vocab.strings[match_id]
matched_span = doc_object[start:end]
print(f"{match_id:<{30}} {string_id:<{15}} {start:{5}} {end:{5}} {matched_span.text:{5}} \
{'previous 5 words'}\
{doc_object[start-5]} {doc_object[start-4]} {doc_object[start-3]} {doc_object[start-2]} {doc_object[start-1]}\
{'next 3 words'}\
{doc_object[end]} {doc_object[end+1]} {doc_object[end+2]} ")
prev_next_words(doc_object)
# ### Q7 : Phrase matching
#
#
# Import the PhraseMatcher library
from spacy.matcher import PhraseMatcher
phrase_matcher = PhraseMatcher(nlp.vocab)
phrase_list = ["longer", "long", "longest"]
phrase_patterns = [nlp.make_doc(word) for word in phrase_list]
phrase_matcher.add("long", None, *phrase_patterns)
phrase_list = ["remember", "remembered", "remembering"]
phrase_patterns = [nlp.make_doc(word) for word in phrase_list]
phrase_matcher.add("remember", None, *phrase_patterns)
phrase_list = ["tonight", "to night", "to-night"]
phrase_patterns = [nlp.make_doc(word) for word in phrase_list]
phrase_matcher.add("tonight", None, *phrase_patterns)
phrase_list = ["policestation", "police-station", "police station"]
phrase_patterns = [nlp.make_doc(word) for word in phrase_list]
phrase_matcher.add("policestation", None, *phrase_patterns)
def find_matches(text):
# find all matches within the doc object
token_matches = phrase_matcher(text)
# For each item in the token_matches provide the following
# match_id is the hash value of the identified token match
print(f"{'Match ID':<{30}} {'String':<{15}} {'Start':{5}} {'End':{5}} {'Match word':{20}}")
for match_id, start, end in token_matches:
string_id = nlp.vocab.strings[match_id]
matched_span = doc_object[start:end]
print(f"{match_id:<{30}} {string_id:<{15}} {start:{5}} {end:{5}} {matched_span.text:{20}}")
find_matches(doc_object)
# ### Q8: Previous 5 words and next 5 words
def prev_next_words(text):
# find all matches within the doc object
token_matches = phrase_matcher(text)
# For each item in the token_matches provide the following
# match_id is the hash value of the identified token match
print(f"{'Match ID':<{30}} {'String':<{15}} {'Start':{5}} {'End':{5}} {'Match word':{5}}")
for match_id, start, end in token_matches:
string_id = nlp.vocab.strings[match_id]
matched_span = doc_object[start:end]
print(f"{match_id:<{30}} {string_id:<{15}} {start:{5}} {end:{5}} {matched_span.text:{5}}\
{'previous 5 words'}\
{doc_object[start-5]} {doc_object[start-4]} {doc_object[start-3]} {doc_object[start-2]} {doc_object[start-1]}\
{'next 3 words'}\
{doc_object[end]} {doc_object[end+1]} {doc_object[end+2]} ")
prev_next_words(doc_object)
# ### Q9: Lemmatisation
# Lemmatization converts words in the second or third forms to their first form variants.
def create_lemmatization(text_to_convert):
for token in text_to_convert:
print (f"{token.text:{15}} {token.lemma_:{30}}")
random_sentence= nlp(sentences[300])
create_lemmatization(random_sentence)
# ### Q10 : Displacy
# +
from spacy import displacy
displacy.render(random_sentence,style = "dep", jupyter = True, options= {"distance" :50,
"font": "Ariel",
"bg": "black",
"color" : "Red",
"arrow_stroke" : 1,
"arrow_spacing": 50,
"arrow_width" : 5,
"word_spacing": 50,
"compact" : False})
# -
| Assessments/Artificial Intelligence 2 - CA 1.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Querying portia - Data analysis with R
# ## Specific time frame data analysis - Last 24 hours
#
#
# We are authenticating sucessfully, so let's dive into the data...
# +
headers <- httr::add_headers(Authorization = "Bearer bdb6e780b43011e7af0b67cba486057b", Accept = "text/csv")
params <- list(order = "-1")
response <- httr::GET("http://io.portia.supe.solutions/api/v1/device/HytTDwUp-j8yrsh8e/port/2/sensor/1", headers, query = params)
content = httr::content(response, "text", encoding = "UTF-8")
readings.temperature = read.csv(text=content, sep=";")
summary(readings.temperature$dimension_value)
# +
response <- httr::GET("http://io.portia.supe.solutions/api/v1/device/HytTDwUp-j8yrsh8e/port/2/sensor/2", headers, query = params)
content = httr::content(response, "text", encoding = "UTF-8")
readings.umidity = read.csv(text=content, sep=";")
summary(readings.umidity$dimension_value)
# -
# +
readings.temperature <- transform(readings.temperature, ts = server_timestamp / 1000, ts_local = package_local_timestamp / 1000, ts_text = as.POSIXct(server_timestamp / 1000, origin="1970-01-01" ) )
readings.temperature <- subset( readings.temperature, select = -c( X, dimension_code, dimension_unity_code, dimension_thing_code, package_device_hash, dimension_port_id, dimension_sensor_id, package_local_timestamp, package_local_timestamp, server_timestamp))
readings.temperature <- subset( readings.temperature, ts > 1508536800)
readings.umidity <- transform(readings.umidity, ts = server_timestamp / 1000, ts_local = package_local_timestamp / 1000, ts_text = as.POSIXct(server_timestamp / 1000, origin="1970-01-01" ) )
readings.umidity <- subset( readings.umidity, select = -c( X, dimension_code, dimension_unity_code, dimension_thing_code, package_device_hash, dimension_port_id, dimension_sensor_id, package_local_timestamp, package_local_timestamp, server_timestamp))
readings.umidity <- subset( readings.umidity, ts > 1508536800)
# -
head(readings.temperature, n=5)
head(readings.umidity, n=5)
# +
diff_ts_temperature = diff(readings.temperature$ts)
summary(diff_ts_temperature)
avgdiff = mean(diff_ts_temperature)
paste("Temperatura - Diferença média entre um pacote e outro: ", avgdiff, " segundos no servidor")
paste("Número de pacotes:", nrow(readings.temperature))
diff_ts_umidity = diff(readings.umidity$ts)
summary(diff_ts_umidity)
avgdiff = mean(diff_ts_umidity)
paste("Umidade - Diferença média entre um pacote e outro: ", avgdiff, " segundos no servidor")
paste("Número de pacotes:", nrow(readings.umidity))
diff_ts_local_temperature = diff(readings.temperature$ts_local)
summary(diff_ts_local_temperature)
avgdiff = mean(diff_ts_local_temperature)
paste("Temperatura - Diferença média entre um pacote e outro: ", avgdiff, " segundos locais")
diff_ts_local_umidity = diff(readings.umidity$ts_local)
summary(diff_ts_local_umidity)
avgdiff = mean(diff_ts_local_umidity)
paste("Umidade - Diferença média entre um pacote e outro: ", avgdiff, "segundos locais")
plot(diff_ts_temperature,type = "l",col = "red", xlab = "intervalo")
lines(diff_ts_umidity, type = "l", col = "blue")
lines(diff_ts_local_temperature, type = "l", col = "black")
lines(diff_ts_local_umidity, type = "l", col = "green")
# legend(2000,9.5,c("Health","Defense"),lwd=c(2.5,2.5),col=c("blue","red"))
# -
summary(readings.temperature$dimension_value)
plot(readings.temperature$ts_text,readings.temperature$dimension_value, "s", col="red")
readings.temperature$index <- seq.int(nrow(readings.temperature))
linear.model = lm(readings.temperature$dimension_value ~ readings.temperature$ts)
abline(linear.model)
# plot(readings.temperature$ts_text,readings.temperature$dimension_value, "o", col="red")
summary(readings.umidity$dimension_value)
plot(readings.umidity$ts_text,readings.umidity$dimension_value, "s", col="blue")
# plot(readings.umidity$ts_text,readings.umidity$dimension_value, "o", col="blue")
par(mfrow=c(2,2))
plot(readings.temperature$ts_text,readings.temperature$dimension_value, "s", col="red")
plot(readings.umidity$ts_text,readings.umidity$dimension_value, "s", col="blue")
m <-mean(readings.temperature$dimension_value);
std<-sqrt(var(readings.temperature$dimension_value))
hist(readings.temperature$dimension_value,prob=T,main="Temperature")
curve(dnorm(x, mean=m, sd=std), col="darkblue", lwd=3, add=TRUE)
m<-mean(readings.umidity$dimension_value);std<-sqrt(var(readings.umidity$dimension_value))
hist(readings.umidity$dimension_value,prob=T,main="Umidity")
curve(dnorm(x, mean=m, sd=std), col="red", lwd=2, add=TRUE)
box()
# +
#readings <- merge(readings.temperature,readings.umidity, by="ts")
# readings
# Define 2 vectors
readings.temperature$dimension_value <- readings.temperature$dimension_value
readings.umidity$dimension_value <- readings.umidity$dimension_value
# Calculate range from 0 to max value of readings.temperature$dimension_value and readings.umidity$dimension_value
g_range <- range(0, readings.temperature$dimension_value, readings.umidity$dimension_value)
# Graph autos using y axis that ranges from 0 to max
# value in readings.temperature$dimension_value or readings.umidity$dimension_value vector. Turn off axes and
# annotations (axis labels) so we can specify them ourself
plot(readings.temperature$dimension_value, type="s", col="red", ylim=g_range,
ann=TRUE)
# axis(2, las=1, at=5*0:g_range[2])
# Graph readings.umidity$dimension_value with red dashed line and square points
lines(readings.umidity$dimension_value, type="s", col="blue")
# Create a title with a red, bold/italic font
#title(main="Umidade e Temperatura")
# Label the x and y axes with dark green text
# title(xlab="Tempo", col.lab=rgb(0,0.5,0))
# Create a legend at (1, g_range[2]) that is slightly smaller
# (cex) and uses the same line colors and points used by
# the actual plots
#legend(1, g_range[2], c("temperatura","umidade"), cex=0.8, col=c("red","blue"), pch=21:22, lty=1:2);
| examples/R-External.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import cPickle as pickle
import numpy as np
import peakutils
from change_point_detector.density_ratio_estimator import DRChangeRateEstimator
import ROOT
ROOT.enableJSVis()
with open("../test/data/test.pkl") as file_:
data = pickle.load(file_)
c1 = ROOT.TCanvas("time series", "time series", 1000,800)
c1.Divide(1,2)
# +
detector = DRChangeRateEstimator(sliding_window=3,
pside_len=3,
cside_len=3,
mergin=-1,
trow_offset=1,
tcol_offset=1)
detector.build(estimation_method="von_mises_fisher",
options=detector.MEAN_OPTION)
change_rates = detector.transform(data["y"])
# +
time_series_graph = ROOT.TGraph(len(data["x"]), data["x"], data["y"])
time_series_graph.SetMarkerStyle(6)
peak_indexes = peakutils.indexes(np.nan_to_num(change_rates), thres=0.1, min_dist=10)
change_point_graph = ROOT.TGraph(len(peak_indexes), data["x"][peak_indexes], data["y"][peak_indexes])
change_point_graph.SetMarkerStyle(23)
change_point_graph.SetMarkerColor(2)
change_rate_graph = ROOT.TGraph(len(data["x"]), data["x"], np.nan_to_num(change_rates))
# -
c1.cd(1)
time_series_graph.Draw("apl")
change_point_graph.Draw("p")
c1.cd(2)
change_rate_graph.Draw("apl")
c1.Draw()
| notebook/test_real.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Aggregating GEO Datasets for TCGA Validation
import NotebookImport
from DX_screen import *
store = pd.HDFStore(MICROARRAY_STORE)
microarray = store['data']
tissue = store['tissue']
tissue.value_counts()
dx = microarray.xs('01',1,1) - microarray.xs('11',1,1)
tt = tissue[:,'01'].replace('COAD','COADREAD')
genes = ti(dx.notnull().sum(1) > 500)
dx = dx.ix[genes]
# Simple average
dx_simple = binomial_test_screen(microarray.ix[genes])
fig, ax = subplots(figsize=(4,4))
s1, s2 = match_series(dx_rna.frac, dx_simple.frac)
plot_regression(s1, s2, density=True, rad=.02, ax=ax, rasterized=True,
line_args={'lw':0})
ax.set_ylabel("GEO microarray")
ax.set_xlabel("TCGA mRNASeq")
ann = ax.get_children()[4]
ann.set_text(ann.get_text().split()[0])
ax.set_xticks([0, .5, 1])
ax.set_yticks([0, .5, 1])
fig.tight_layout()
# Group by tissue type first and then average. This is to limit the heavy skew of liver cancer samples in the GEO dataset.
pos = (dx>0).groupby(tt, axis=1).sum()
count = dx.groupby(tt, axis=1).count().replace(0, np.nan)
count = count[count.sum(1) > 500]
frac_df = 1.*pos / count
frac_microarray = frac_df.mean(1)
fig, ax = subplots(figsize=(4,4))
s1, s2 = match_series(dx_rna.frac, frac_microarray)
plot_regression(s1, s2, density=True, rad=.02, ax=ax, rasterized=True,
line_args={'lw':0})
ax.set_ylabel("GEO microarray")
ax.set_xlabel("TCGA mRNASeq")
ann = ax.get_children()[4]
ann.set_text(ann.get_text().split()[0])
ax.set_xticks([0, .5, 1])
ax.set_yticks([0, .5, 1])
fig.tight_layout()
# Grouping both TCGA and GEO based on tissue first. This is not the approach we use for the TCGA data in the rest of the analyses, so I'm just doing this to show that it does not effect performance of the replication.
# +
dx2 = (rna_df.xs('01',1,1) - rna_df.xs('11',1,1)).dropna(1)
cc = codes.ix[dx2.columns]
cc = cc[cc.isin(ti(cc.value_counts() > 10))]
pos = (dx2>0).groupby(cc, axis=1).sum()
count = dx2.replace(0, np.nan).groupby(cc, axis=1).count()
count = count[count.sum(1) > 500]
frac_df = 1.*pos / count
frac_tcga= frac_df.mean(1)
# -
fig, ax = subplots(figsize=(4,4))
s1, s2 = match_series(frac_tcga, frac_microarray)
plot_regression(s1, s2, density=True, rad=.02, ax=ax, rasterized=True,
line_args={'lw':0})
ax.set_ylabel("GEO microarray")
ax.set_xlabel("TCGA mRNASeq")
ann = ax.get_children()[4]
ann.set_text(ann.get_text().split()[0])
ax.set_xticks([0, .5, 1])
ax.set_yticks([0, .5, 1])
fig.tight_layout()
| Notebooks/microarray_validation_aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# What is `torch.nn` *really*?
# ============================
# by <NAME>, `fast.ai <https://www.fast.ai>`_. Thanks to <NAME> and <NAME>.
#
#
# We recommend running this tutorial as a notebook, not a script. To download the notebook (.ipynb) file,
# click the link at the top of the page.
#
# PyTorch provides the elegantly designed modules and classes `torch.nn <https://pytorch.org/docs/stable/nn.html>`_ ,
# `torch.optim <https://pytorch.org/docs/stable/optim.html>`_ ,
# `Dataset <https://pytorch.org/docs/stable/data.html?highlight=dataset#torch.utils.data.Dataset>`_ ,
# and `DataLoader <https://pytorch.org/docs/stable/data.html?highlight=dataloader#torch.utils.data.DataLoader>`_
# to help you create and train neural networks.
# In order to fully utilize their power and customize
# them for your problem, you need to really understand exactly what they're
# doing. To develop this understanding, we will first train basic neural net
# on the MNIST data set without using any features from these models; we will
# initially only use the most basic PyTorch tensor functionality. Then, we will
# incrementally add one feature from ``torch.nn``, ``torch.optim``, ``Dataset``, or
# ``DataLoader`` at a time, showing exactly what each piece does, and how it
# works to make the code either more concise, or more flexible.
#
# **This tutorial assumes you already have PyTorch installed, and are familiar
# with the basics of tensor operations.** (If you're familiar with Numpy array
# operations, you'll find the PyTorch tensor operations used here nearly identical).
#
# MNIST data setup
# ----------------
#
# We will use the classic `MNIST <http://deeplearning.net/data/mnist/>`_ dataset,
# which consists of black-and-white images of hand-drawn digits (between 0 and 9).
#
# We will use `pathlib <https://docs.python.org/3/library/pathlib.html>`_
# for dealing with paths (part of the Python 3 standard library), and will
# download the dataset using
# `requests <http://docs.python-requests.org/en/master/>`_. We will only
# import modules when we use them, so you can see exactly what's being
# used at each point.
#
#
# +
from pathlib import Path
import requests
DATA_PATH = Path("data")
PATH = DATA_PATH / "mnist"
PATH.mkdir(parents=True, exist_ok=True)
URL = "http://deeplearning.net/data/mnist/"
FILENAME = "mnist.pkl.gz"
if not (PATH / FILENAME).exists():
content = requests.get(URL + FILENAME).content
(PATH / FILENAME).open("wb").write(content)
# -
# This dataset is in numpy array format, and has been stored using pickle,
# a python-specific format for serializing data.
#
#
# +
import pickle
import gzip
with gzip.open((PATH / FILENAME).as_posix(), "rb") as f:
((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding="latin-1")
# -
# Each image is 28 x 28, and is being stored as a flattened row of length
# 784 (=28x28). Let's take a look at one; we need to reshape it to 2d
# first.
#
#
# +
from matplotlib import pyplot
import numpy as np
pyplot.imshow(x_train[0].reshape((28, 28)), cmap="gray")
print(x_train.shape)
# -
# PyTorch uses ``torch.tensor``, rather than numpy arrays, so we need to
# convert our data.
#
#
# +
import torch
x_train, y_train, x_valid, y_valid = map(
torch.tensor, (x_train, y_train, x_valid, y_valid)
)
n, c = x_train.shape
x_train, x_train.shape, y_train.min(), y_train.max()
print(x_train, y_train)
print(x_train.shape)
print(y_train.min(), y_train.max())
# -
# Neural net from scratch (no torch.nn)
# ---------------------------------------------
#
# Let's first create a model using nothing but PyTorch tensor operations. We're assuming
# you're already familiar with the basics of neural networks. (If you're not, you can
# learn them at `course.fast.ai <https://course.fast.ai>`_).
#
# PyTorch provides methods to create random or zero-filled tensors, which we will
# use to create our weights and bias for a simple linear model. These are just regular
# tensors, with one very special addition: we tell PyTorch that they require a
# gradient. This causes PyTorch to record all of the operations done on the tensor,
# so that it can calculate the gradient during back-propagation *automatically*!
#
# For the weights, we set ``requires_grad`` **after** the initialization, since we
# don't want that step included in the gradient. (Note that a trailling ``_`` in
# PyTorch signifies that the operation is performed in-place.)
#
# <div class="alert alert-info"><h4>Note</h4><p>We are initializing the weights here with
# `Xavier initialisation <http://proceedings.mlr.press/v9/glorot10a/glorot10a.pdf>`_
# (by multiplying with 1/sqrt(n)).</p></div>
#
#
# +
import math
weights = torch.randn(784, 10) / math.sqrt(784)
weights.requires_grad_()
bias = torch.zeros(10, requires_grad=True)
# -
# Thanks to PyTorch's ability to calculate gradients automatically, we can
# use any standard Python function (or callable object) as a model! So
# let's just write a plain matrix multiplication and broadcasted addition
# to create a simple linear model. We also need an activation function, so
# we'll write `log_softmax` and use it. Remember: although PyTorch
# provides lots of pre-written loss functions, activation functions, and
# so forth, you can easily write your own using plain python. PyTorch will
# even create fast GPU or vectorized CPU code for your function
# automatically.
#
#
# +
def log_softmax(x):
return x - x.exp().sum(-1).log().unsqueeze(-1)
def model(xb):
return log_softmax(xb @ weights + bias)
# -
# In the above, the ``@`` stands for the dot product operation. We will call
# our function on one batch of data (in this case, 64 images). This is
# one *forward pass*. Note that our predictions won't be any better than
# random at this stage, since we start with random weights.
#
#
# +
bs = 64 # batch size
xb = x_train[0:bs] # a mini-batch from x
preds = model(xb) # predictions
preds[0], preds.shape
print(preds[0], preds.shape)
# -
# As you see, the ``preds`` tensor contains not only the tensor values, but also a
# gradient function. We'll use this later to do backprop.
#
# Let's implement negative log-likelihood to use as the loss function
# (again, we can just use standard Python):
#
#
# +
def nll(input, target):
return -input[range(target.shape[0]), target].mean()
loss_func = nll
# -
# Let's check our loss with our random model, so we can see if we improve
# after a backprop pass later.
#
#
yb = y_train[0:bs]
print(loss_func(preds, yb))
# Let's also implement a function to calculate the accuracy of our model.
# For each prediction, if the index with the largest value matches the
# target value, then the prediction was correct.
#
#
def accuracy(out, yb):
preds = torch.argmax(out, dim=1)
return (preds == yb).float().mean()
# Let's check the accuracy of our random model, so we can see if our
# accuracy improves as our loss improves.
#
#
print(accuracy(preds, yb))
# We can now run a training loop. For each iteration, we will:
#
# - select a mini-batch of data (of size ``bs``)
# - use the model to make predictions
# - calculate the loss
# - ``loss.backward()`` updates the gradients of the model, in this case, ``weights``
# and ``bias``.
#
# We now use these gradients to update the weights and bias. We do this
# within the ``torch.no_grad()`` context manager, because we do not want these
# actions to be recorded for our next calculation of the gradient. You can read
# more about how PyTorch's Autograd records operations
# `here <https://pytorch.org/docs/stable/notes/autograd.html>`_.
#
# We then set the
# gradients to zero, so that we are ready for the next loop.
# Otherwise, our gradients would record a running tally of all the operations
# that had happened (i.e. ``loss.backward()`` *adds* the gradients to whatever is
# already stored, rather than replacing them).
#
# .. tip:: You can use the standard python debugger to step through PyTorch
# code, allowing you to check the various variable values at each step.
# Uncomment ``set_trace()`` below to try it out.
#
#
#
# +
from IPython.core.debugger import set_trace
lr = 0.5 # learning rate
epochs = 2 # how many epochs to train for
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
# set_trace()
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
with torch.no_grad():
weights -= weights.grad * lr
bias -= bias.grad * lr
weights.grad.zero_()
bias.grad.zero_()
# -
# That's it: we've created and trained a minimal neural network (in this case, a
# logistic regression, since we have no hidden layers) entirely from scratch!
#
# Let's check the loss and accuracy and compare those to what we got
# earlier. We expect that the loss will have decreased and accuracy to
# have increased, and they have.
#
#
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
# Using torch.nn.functional
# ------------------------------
#
# We will now refactor our code, so that it does the same thing as before, only
# we'll start taking advantage of PyTorch's ``nn`` classes to make it more concise
# and flexible. At each step from here, we should be making our code one or more
# of: shorter, more understandable, and/or more flexible.
#
# The first and easiest step is to make our code shorter by replacing our
# hand-written activation and loss functions with those from ``torch.nn.functional``
# (which is generally imported into the namespace ``F`` by convention). This module
# contains all the functions in the ``torch.nn`` library (whereas other parts of the
# library contain classes). As well as a wide range of loss and activation
# functions, you'll also find here some convenient functions for creating neural
# nets, such as pooling functions. (There are also functions for doing convolutions,
# linear layers, etc, but as we'll see, these are usually better handled using
# other parts of the library.)
#
# If you're using negative log likelihood loss and log softmax activation,
# then Pytorch provides a single function ``F.cross_entropy`` that combines
# the two. So we can even remove the activation function from our model.
#
#
# +
import torch.nn.functional as F
loss_func = F.cross_entropy
def model(xb):
return xb @ weights + bias
# -
# Note that we no longer call ``log_softmax`` in the ``model`` function. Let's
# confirm that our loss and accuracy are the same as before:
#
#
print(loss_func(model(xb), yb), accuracy(model(xb), yb))
# Refactor using nn.Module
# -----------------------------
# Next up, we'll use ``nn.Module`` and ``nn.Parameter``, for a clearer and more
# concise training loop. We subclass ``nn.Module`` (which itself is a class and
# able to keep track of state). In this case, we want to create a class that
# holds our weights, bias, and method for the forward step. ``nn.Module`` has a
# number of attributes and methods (such as ``.parameters()`` and ``.zero_grad()``)
# which we will be using.
#
# <div class="alert alert-info"><h4>Note</h4><p>``nn.Module`` (uppercase M) is a PyTorch specific concept, and is a
# class we'll be using a lot. ``nn.Module`` is not to be confused with the Python
# concept of a (lowercase ``m``) `module <https://docs.python.org/3/tutorial/modules.html>`_,
# which is a file of Python code that can be imported.</p></div>
#
#
# +
from torch import nn
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.weights = nn.Parameter(torch.randn(784, 10) / math.sqrt(784))
self.bias = nn.Parameter(torch.zeros(10))
def forward(self, xb):
return xb @ self.weights + self.bias
# -
# Since we're now using an object instead of just using a function, we
# first have to instantiate our model:
#
#
model = Mnist_Logistic()
# Now we can calculate the loss in the same way as before. Note that
# ``nn.Module`` objects are used as if they are functions (i.e they are
# *callable*), but behind the scenes Pytorch will call our ``forward``
# method automatically.
#
#
print(loss_func(model(xb), yb))
# Previously for our training loop we had to update the values for each parameter
# by name, and manually zero out the grads for each parameter separately, like this:
# ::
# with torch.no_grad():
# weights -= weights.grad * lr
# bias -= bias.grad * lr
# weights.grad.zero_()
# bias.grad.zero_()
#
#
# Now we can take advantage of model.parameters() and model.zero_grad() (which
# are both defined by PyTorch for ``nn.Module``) to make those steps more concise
# and less prone to the error of forgetting some of our parameters, particularly
# if we had a more complicated model:
# ::
# with torch.no_grad():
# for p in model.parameters(): p -= p.grad * lr
# model.zero_grad()
#
#
# We'll wrap our little training loop in a ``fit`` function so we can run it
# again later.
#
#
# +
def fit():
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
with torch.no_grad():
for p in model.parameters():
p -= p.grad * lr
model.zero_grad()
fit()
# -
# Let's double-check that our loss has gone down:
#
#
print(loss_func(model(xb), yb))
# Refactor using nn.Linear
# -------------------------
#
# We continue to refactor our code. Instead of manually defining and
# initializing ``self.weights`` and ``self.bias``, and calculating ``xb @
# self.weights + self.bias``, we will instead use the Pytorch class
# `nn.Linear <https://pytorch.org/docs/stable/nn.html#linear-layers>`_ for a
# linear layer, which does all that for us. Pytorch has many types of
# predefined layers that can greatly simplify our code, and often makes it
# faster too.
#
#
class Mnist_Logistic(nn.Module):
def __init__(self):
super().__init__()
self.lin = nn.Linear(784, 10)
def forward(self, xb):
return self.lin(xb)
# We instantiate our model and calculate the loss in the same way as before:
#
#
model = Mnist_Logistic()
print(loss_func(model(xb), yb))
# We are still able to use our same ``fit`` method as before.
#
#
# +
fit()
print(loss_func(model(xb), yb))
# -
# Refactor using optim
# ------------------------------
#
# Pytorch also has a package with various optimization algorithms, ``torch.optim``.
# We can use the ``step`` method from our optimizer to take a forward step, instead
# of manually updating each parameter.
#
# This will let us replace our previous manually coded optimization step:
# ::
# with torch.no_grad():
# for p in model.parameters(): p -= p.grad * lr
# model.zero_grad()
#
# and instead use just:
# ::
# opt.step()
# opt.zero_grad()
#
# (``optim.zero_grad()`` resets the gradient to 0 and we need to call it before
# computing the gradient for the next minibatch.)
#
#
from torch import optim
# We'll define a little function to create our model and optimizer so we
# can reuse it in the future.
#
#
# +
def get_model():
model = Mnist_Logistic()
return model, optim.SGD(model.parameters(), lr=lr)
model, opt = get_model()
print(loss_func(model(xb), yb))
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
start_i = i * bs
end_i = start_i + bs
xb = x_train[start_i:end_i]
yb = y_train[start_i:end_i]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(xb), yb))
# -
# Refactor using Dataset
# ------------------------------
#
# PyTorch has an abstract Dataset class. A Dataset can be anything that has
# a ``__len__`` function (called by Python's standard ``len`` function) and
# a ``__getitem__`` function as a way of indexing into it.
# `This tutorial <https://pytorch.org/tutorials/beginner/data_loading_tutorial.html>`_
# walks through a nice example of creating a custom ``FacialLandmarkDataset`` class
# as a subclass of ``Dataset``.
#
# PyTorch's `TensorDataset <https://pytorch.org/docs/stable/_modules/torch/utils/data/dataset.html#TensorDataset>`_
# is a Dataset wrapping tensors. By defining a length and way of indexing,
# this also gives us a way to iterate, index, and slice along the first
# dimension of a tensor. This will make it easier to access both the
# independent and dependent variables in the same line as we train.
#
#
from torch.utils.data import TensorDataset
# Both ``x_train`` and ``y_train`` can be combined in a single ``TensorDataset``,
# which will be easier to iterate over and slice.
#
#
train_ds = TensorDataset(x_train, y_train)
# Previously, we had to iterate through minibatches of x and y values separately:
# ::
# xb = x_train[start_i:end_i]
# yb = y_train[start_i:end_i]
#
#
# Now, we can do these two steps together:
# ::
# xb,yb = train_ds[i*bs : i*bs+bs]
#
#
#
# +
model, opt = get_model()
for epoch in range(epochs):
for i in range((n - 1) // bs + 1):
xb, yb = train_ds[i * bs: i * bs + bs]
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(xb), yb))
# -
# Refactor using DataLoader
# ------------------------------
#
# Pytorch's ``DataLoader`` is responsible for managing batches. You can
# create a ``DataLoader`` from any ``Dataset``. ``DataLoader`` makes it easier
# to iterate over batches. Rather than having to use ``train_ds[i*bs : i*bs+bs]``,
# the DataLoader gives us each minibatch automatically.
#
#
# +
from torch.utils.data import DataLoader
train_ds = TensorDataset(x_train, y_train)
train_dl = DataLoader(train_ds, batch_size=bs)
# -
# Previously, our loop iterated over batches (xb, yb) like this:
# ::
# for i in range((n-1)//bs + 1):
# xb,yb = train_ds[i*bs : i*bs+bs]
# pred = model(xb)
#
# Now, our loop is much cleaner, as (xb, yb) are loaded automatically from the data loader:
# ::
# for xb,yb in train_dl:
# pred = model(xb)
#
#
# +
model, opt = get_model()
for epoch in range(epochs):
for xb, yb in train_dl:
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
print(loss_func(model(xb), yb))
# -
# Thanks to Pytorch's ``nn.Module``, ``nn.Parameter``, ``Dataset``, and ``DataLoader``,
# our training loop is now dramatically smaller and easier to understand. Let's
# now try to add the basic features necessary to create effecive models in practice.
#
# Add validation
# -----------------------
#
# In section 1, we were just trying to get a reasonable training loop set up for
# use on our training data. In reality, you **always** should also have
# a `validation set <https://www.fast.ai/2017/11/13/validation-sets/>`_, in order
# to identify if you are overfitting.
#
# Shuffling the training data is
# `important <https://www.quora.com/Does-the-order-of-training-data-matter-when-training-neural-networks>`_
# to prevent correlation between batches and overfitting. On the other hand, the
# validation loss will be identical whether we shuffle the validation set or not.
# Since shuffling takes extra time, it makes no sense to shuffle the validation data.
#
# We'll use a batch size for the validation set that is twice as large as
# that for the training set. This is because the validation set does not
# need backpropagation and thus takes less memory (it doesn't need to
# store the gradients). We take advantage of this to use a larger batch
# size and compute the loss more quickly.
#
#
# +
train_ds = TensorDataset(x_train, y_train)
train_dl = DataLoader(train_ds, batch_size=bs, shuffle=True)
valid_ds = TensorDataset(x_valid, y_valid)
valid_dl = DataLoader(valid_ds, batch_size=bs * 2)
# -
# We will calculate and print the validation loss at the end of each epoch.
#
# (Note that we always call ``model.train()`` before training, and ``model.eval()``
# before inference, because these are used by layers such as ``nn.BatchNorm2d``
# and ``nn.Dropout`` to ensure appropriate behaviour for these different phases.)
#
#
# +
model, opt = get_model()
for epoch in range(epochs):
model.train()
for xb, yb in train_dl:
pred = model(xb)
loss = loss_func(pred, yb)
loss.backward()
opt.step()
opt.zero_grad()
model.eval()
with torch.no_grad():
valid_loss = sum(loss_func(model(xb), yb) for xb, yb in valid_dl)
print(epoch, valid_loss / len(valid_dl))
# -
# Create fit() and get_data()
# ----------------------------------
#
# We'll now do a little refactoring of our own. Since we go through a similar
# process twice of calculating the loss for both the training set and the
# validation set, let's make that into its own function, ``loss_batch``, which
# computes the loss for one batch.
#
# We pass an optimizer in for the training set, and use it to perform
# backprop. For the validation set, we don't pass an optimizer, so the
# method doesn't perform backprop.
#
#
def loss_batch(model, loss_func, xb, yb, opt=None):
loss = loss_func(model(xb), yb)
if opt is not None:
loss.backward()
opt.step()
opt.zero_grad()
return loss.item(), len(xb)
# ``fit`` runs the necessary operations to train our model and compute the
# training and validation losses for each epoch.
#
#
# +
import numpy as np
def fit(epochs, model, loss_func, opt, train_dl, valid_dl):
for epoch in range(epochs):
model.train()
for xb, yb in train_dl:
loss_batch(model, loss_func, xb, yb, opt)
model.eval()
with torch.no_grad():
losses, nums = zip(
*[loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl]
)
val_loss = np.sum(np.multiply(losses, nums)) / np.sum(nums)
print(epoch, val_loss)
# -
# ``get_data`` returns dataloaders for the training and validation sets.
#
#
def get_data(train_ds, valid_ds, bs):
return (
DataLoader(train_ds, batch_size=bs, shuffle=True),
DataLoader(valid_ds, batch_size=bs * 2),
)
# Now, our whole process of obtaining the data loaders and fitting the
# model can be run in 3 lines of code:
#
#
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
model, opt = get_model()
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# You can use these basic 3 lines of code to train a wide variety of models.
# Let's see if we can use them to train a convolutional neural network (CNN)!
#
# Switch to CNN
# -------------
#
# We are now going to build our neural network with three convolutional layers.
# Because none of the functions in the previous section assume anything about
# the model form, we'll be able to use them to train a CNN without any modification.
#
# We will use Pytorch's predefined
# `Conv2d <https://pytorch.org/docs/stable/nn.html#torch.nn.Conv2d>`_ class
# as our convolutional layer. We define a CNN with 3 convolutional layers.
# Each convolution is followed by a ReLU. At the end, we perform an
# average pooling. (Note that ``view`` is PyTorch's version of numpy's
# ``reshape``)
#
#
# +
class Mnist_CNN(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1)
self.conv2 = nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1)
self.conv3 = nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1)
def forward(self, xb):
xb = xb.view(-1, 1, 28, 28)
xb = F.relu(self.conv1(xb))
xb = F.relu(self.conv2(xb))
xb = F.relu(self.conv3(xb))
xb = F.avg_pool2d(xb, 4)
return xb.view(-1, xb.size(1))
lr = 0.1
# -
# `Momentum <https://cs231n.github.io/neural-networks-3/#sgd>`_ is a variation on
# stochastic gradient descent that takes previous updates into account as well
# and generally leads to faster training.
#
#
# +
model = Mnist_CNN()
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# -
# nn.Sequential
# ------------------------
#
# ``torch.nn`` has another handy class we can use to simply our code:
# `Sequential <https://pytorch.org/docs/stable/nn.html#torch.nn.Sequential>`_ .
# A ``Sequential`` object runs each of the modules contained within it, in a
# sequential manner. This is a simpler way of writing our neural network.
#
# To take advantage of this, we need to be able to easily define a
# **custom layer** from a given function. For instance, PyTorch doesn't
# have a `view` layer, and we need to create one for our network. ``Lambda``
# will create a layer that we can then use when defining a network with
# ``Sequential``.
#
#
# +
class Lambda(nn.Module):
def __init__(self, func):
super().__init__()
self.func = func
def forward(self, x):
return self.func(x)
def preprocess(x):
return x.view(-1, 1, 28, 28)
# -
# The model created with ``Sequential`` is simply:
#
#
# +
model = nn.Sequential(
Lambda(preprocess),
nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.AvgPool2d(4),
Lambda(lambda x: x.view(x.size(0), -1)),
)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# -
# Wrapping DataLoader
# -----------------------------
#
# Our CNN is fairly concise, but it only works with MNIST, because:
# - It assumes the input is a 28\*28 long vector
# - It assumes that the final CNN grid size is 4\*4 (since that's the average
# pooling kernel size we used)
#
# Let's get rid of these two assumptions, so our model works with any 2d
# single channel image. First, we can remove the initial Lambda layer but
# moving the data preprocessing into a generator:
#
#
# +
def preprocess(x, y):
return x.view(-1, 1, 28, 28), y
class WrappedDataLoader:
def __init__(self, dl, func):
self.dl = dl
self.func = func
def __len__(self):
return len(self.dl)
def __iter__(self):
batches = iter(self.dl)
for b in batches:
yield (self.func(*b))
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
train_dl = WrappedDataLoader(train_dl, preprocess)
valid_dl = WrappedDataLoader(valid_dl, preprocess)
# -
# Next, we can replace ``nn.AvgPool2d`` with ``nn.AdaptiveAvgPool2d``, which
# allows us to define the size of the *output* tensor we want, rather than
# the *input* tensor we have. As a result, our model will work with any
# size input.
#
#
# +
model = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1),
nn.ReLU(),
nn.AdaptiveAvgPool2d(1),
Lambda(lambda x: x.view(x.size(0), -1)),
)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# -
# Let's try it out:
#
#
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# Using your GPU
# ---------------
#
# If you're lucky enough to have access to a CUDA-capable GPU (you can
# rent one for about $0.50/hour from most cloud providers) you can
# use it to speed up your code. First check that your GPU is working in
# Pytorch:
#
#
print(torch.cuda.is_available())
# And then create a device object for it:
#
#
dev = torch.device(
"cuda") if torch.cuda.is_available() else torch.device("cpu")
# Let's update ``preprocess`` to move batches to the GPU:
#
#
# +
def preprocess(x, y):
return x.view(-1, 1, 28, 28).to(dev), y.to(dev)
train_dl, valid_dl = get_data(train_ds, valid_ds, bs)
train_dl = WrappedDataLoader(train_dl, preprocess)
valid_dl = WrappedDataLoader(valid_dl, preprocess)
# -
# Finally, we can move our model to the GPU.
#
#
model.to(dev)
opt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# You should find it runs faster now:
#
#
fit(epochs, model, loss_func, opt, train_dl, valid_dl)
# Closing thoughts
# -----------------
#
# We now have a general data pipeline and training loop which you can use for
# training many types of models using Pytorch. To see how simple training a model
# can now be, take a look at the `mnist_sample` sample notebook.
#
# Of course, there are many things you'll want to add, such as data augmentation,
# hyperparameter tuning, monitoring training, transfer learning, and so forth.
# These features are available in the fastai library, which has been developed
# using the same design approach shown in this tutorial, providing a natural
# next step for practitioners looking to take their models further.
#
# We promised at the start of this tutorial we'd explain through example each of
# ``torch.nn``, ``torch.optim``, ``Dataset``, and ``DataLoader``. So let's summarize
# what we've seen:
#
# - **torch.nn**
#
# + ``Module``: creates a callable which behaves like a function, but can also
# contain state(such as neural net layer weights). It knows what ``Parameter`` (s) it
# contains and can zero all their gradients, loop through them for weight updates, etc.
# + ``Parameter``: a wrapper for a tensor that tells a ``Module`` that it has weights
# that need updating during backprop. Only tensors with the `requires_grad` attribute set are updated
# + ``functional``: a module(usually imported into the ``F`` namespace by convention)
# which contains activation functions, loss functions, etc, as well as non-stateful
# versions of layers such as convolutional and linear layers.
# - ``torch.optim``: Contains optimizers such as ``SGD``, which update the weights
# of ``Parameter`` during the backward step
# - ``Dataset``: An abstract interface of objects with a ``__len__`` and a ``__getitem__``,
# including classes provided with Pytorch such as ``TensorDataset``
# - ``DataLoader``: Takes any ``Dataset`` and creates an iterator which returns batches of data.
#
#
| docs/_downloads/d9398fce39ca80dc4bb8b8ea55b575a8/nn_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# +
using PyPlot
#workspace()
include("HistokatControllerImageLoader.jl")
using HistokatControllerImageLoader: getTile, getImageObject, getVoxelSize, getExtent
filenameR = "/Users/jo/data/example-data-LL1_1_CD146-2014.tif"
filenameT = "/Users/jo/data/example-data-LL1_4_KL1-2014.tif"
imageR = getImageObject(filenameR)
imageT = getImageObject(filenameT)
using PyCall
@pyimport histokat
getVoxelSize(imageR)
im = getTile(imageR, 9, 1,1,0)
# -
im.data.data
figure()
arr = getTile(imageR, 9, 0,0,0)
removeBlack!(arr)
imshow(arr, cmap="gray")
colorbar()
figure()
arr = getTile(imageT, 9, 0,0,0)
removeBlack!(arr)
imshow(arr, cmap="gray")
colorbar()
a = rand(10,10,3)
sum(a,3)
ENV["PYTHON"]="/usr/local/anaconda3/bin/python"
Pkg.build("PyCall")
| histo/load_histo_image_jl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # COMP 562 – Lecture 3
#
#
# $$
# \renewcommand{\xx}{\mathbf{x}}
# \renewcommand{\yy}{\mathbf{y}}
# \renewcommand{\loglik}{\log\mathcal{L}}
# \renewcommand{\likelihood}{\mathcal{L}}
# \renewcommand{\Data}{\textrm{Data}}
# \renewcommand{\given}{ | }
# \renewcommand{\MLE}{\textrm{MLE}}
# \renewcommand{\Gaussian}[2]{\mathcal{N}\left(#1,#2\right)}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding $\mu^{MLE}$ of Gaussian Distribution
#
# We left as an exercise a problem to come up with a maximum likelihood estimate for parameter $\mu$ of a Gaussian distribution
#
# $$
# p(x\given\mu,\sigma^2)= \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x-\mu)^2}
# $$
#
# So we will do that now
# + [markdown] slideshow={"slide_type": "slide"}
# Likelihood function is
#
# $$
# \likelihood(\mu,\sigma^2\given\xx) = \prod_{i=1}^N p(x_i\given\mu,\sigma^2) = \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x_i-\mu)^2}
# $$
#
# Log-likelihood function is
#
# $$
# \log\likelihood(\mu,\sigma^2\given\xx) = \log \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x_i-\mu)^2} = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2} -\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding $\mu^{MLE}$ of Gaussian Distribution
#
# Our recipe is:
#
# 1. Take the function you want to maximize:
#
# $$
# f(\mu) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
#
# 2. Compute its first derivative: $\frac{\partial}{\partial \mu} f(\mu)$
# 3. Equate that derivative to zero and solve: $\frac{\partial}{\partial \mu} f(\mu) = 0$
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# The first derivative is
#
# $$
# \frac{\partial}{\partial \mu} f(\mu) = \sum_{i=1}^N \left[ \frac{1}{\sigma^2}(x_i - \mu)\right]
# $$
#
# We equate it to zero and solve
#
# $$
# \sum_{i=1}^N \left[ \frac{1}{\sigma^2}(x_i - \mu)\right] = 0
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# \begin{aligned}
# \frac{\sum_{i=1}^N x_i}{N} &= \mu
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding ${\sigma^{2}}^{MLE}$ of Gaussian Distribution
#
# Our recipe is:
#
# 1. Take the function you want to maximize:
#
# $$
# f(\sigma^{2}) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
#
# 2. Compute its first derivative: $\frac{\partial}{\partial \sigma^{2}} f(\sigma^{2})$
# 3. Equate that derivative to zero and solve: $\frac{\partial}{\partial \sigma^{2}} f(\sigma^{2}) = 0$
#
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# f(\sigma^{2}) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right] = - \frac{N}{2}\log{2\pi} - \frac{N}{2}\log{\sigma^2} - \frac{1}{2\sigma^2} \sum_{i=1}^N \left[(x_i-\mu)^2\right]
# $$
#
# The first derivative is
#
# $$
# \frac{\partial}{\partial \sigma^{2}} f(\sigma^{2}) = - \frac{N}{2\sigma^{2}} - \left(\frac{1}{2} \sum_{i=1}^N \left[{(x_i - \mu)}^{2}\right]\right)\frac{\partial}{\partial \sigma^{2}}\left(\frac{1}{\sigma^{2}}\right) \\
# = - \frac{N}{2\sigma^{2}} - \left(\frac{1}{2} \sum_{i=1}^N \left[{(x_i - \mu)}^{2}\right]\right)\left(-\frac{1}{{(\sigma^{2})}^{2}}\right) = \frac{1}{2\sigma^{2}} \left(\frac{1}{\sigma^{2}} \sum_{i=1}^N \left[{(x_i - \mu)}^{2} \right] - N \right)
# $$
#
# Which, if we rule out $\sigma^{2} = 0$, is equal to zero only if
#
# $$
# \sigma^{2} = \frac{1}{N} \sum_{i=1}^N \left[{(x_i - \mu)}^{2} \right]
# $$
#
# **<font color='red'> Please Verify both ${\mu}^{MLE}$ and ${\sigma^{2}}^{MLE}$ using seconed derivative test </font>**
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression
#
# Formally, we would write
#
# $$
# \begin{aligned}
# y &= \beta_0 + \sum_j x_j \beta_j + \epsilon \\
# \epsilon &\sim \Gaussian{0}{\sigma^2}
# \end{aligned}
# $$
#
# or more compactly
#
# $$
# y \given \xx \sim \Gaussian{\beta_0 + \sum_j x_j \beta_j}{ \sigma^2}
# $$
#
# Notice that the function is linear in the parameters $\beta=(\beta_0,\beta_1,…,\beta_p)$, not necessarily in terms of the covariates
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression
#
# Probability of target variable $y$
#
# $$
# p(y\given\xx,\beta_0,\beta,\sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-\underbrace{(\beta_0 + \sum_j x_j \beta_j)}_{\textrm{mean of the Gaussian}}\right)^2\right\}
# $$
#
# In the case of the 6th grader's height, we made **the same** prediction for any other 6th grader (58.5 inches)
#
# In our COMP 562 grade example, we compute a potentially different mean for every student
#
# $$
# \beta_0 + \beta_{\textrm{COMP410}}*\textrm{COMP410} + \beta_{\textrm{MATH233}}*\textrm{MATH233} + \beta_{\textrm{STOR435}}*\textrm{STOR435} + \beta_{\textrm{beers}}* \textrm{beers}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Likelihood
#
# We start by writing out a likelihood for linear regression is
#
# $$
# \likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) =
# \prod_{i=1}^N p(y\given\xx,\beta_0,\beta,\sigma^2) =
# \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right\}
# $$
#
# Log-likelihood for linear regression is
#
# $$
# \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) = \sum_{i=1}^N \left[ -\frac{1}{2}\log 2\pi\sigma^2 -\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right] \\ = - \frac{N}{2}\log(2\pi\sigma^2) -\frac{1}{2\sigma^2} \sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2 = - \frac{N}{2}\log(2\pi\sigma^2) -\frac{RSS}{2\sigma^2}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# We will refer to expression $y_i-(\beta_0 + \sum_j x_j \beta_j)$ as **residual**, and hence **RSS** stands for **residual sum of squares** or **sum of squared errors** and is defined by
#
# $$
# RSS = \sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2
# $$
#
# And RSS/N is called the **mean squared error** or **MSE**
#
# $$
# MSE = \frac{1}{N}\sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2
# $$
#
# Hence, maximizing log-likelihood is equivalent to minimizing RSS or MSE
# + [markdown] slideshow={"slide_type": "slide"}
# Another way to see this is to consider a very simplified version of Taylor's theorem
#
# **Theorem.** Given a function $f(\cdot)$ which is smooth at $x$
#
# $$
# f(x + d) = f(x) + f'(x)d + O(d^2)
# $$
#
# In words, close to $x$ function $f(\cdot)$ is very close to being a linear function of $d$
#
# $$
# f(x + \color{blue}{d}) = f(x) + f'(x)\color{blue}{d}
# $$
#
# Slope of the best linear approximation is $f'(x)$, i.e.,$\hspace{0.5em}$$f'(x)$ tells us in which direction function grows
#
# + [markdown] slideshow={"slide_type": "slide"}
# * Gradient Ascent\Descent: Choose initial ${\mathbf{\theta}^{(0)}} \in \mathbb{R}^{n}$, repeat:
# $$ \;
# \begin{aligned}
# {\mathbf{\theta}^{(k)}} = {\mathbf{\theta}^{(k-1)}} \pm t_{k}.\nabla f({\mathbf{\theta}^{(k-1)}}), k =1,2,3,\ldots
# \end{aligned}
# $$
# Where $t_{k}$ is the step size (learning rate) at step $k$
#
# * Stop at some point using a stopping criteria (depend on the problem we are solving), for example:
# * Maximum number of iterations reached
# * $| f({\mathbf{\theta}^{(k)}}) − f({\mathbf{\theta}^{(k-1)}}) | < \epsilon$
# + [markdown] slideshow={"slide_type": "slide"}
# 2. Use Line search Strategy
# * At each iteration, do the best you can along the direction of the gradient,
#
# $$
# \begin{aligned}
# t = \mathop{\textrm{argmax}}_{s \geq 0} f(\mathbf{\theta} + s.\nabla f({\mathbf{\theta}}))
# \end{aligned}
# $$
#
# * Usually, it is not possible to do this minimization exactly, and approximation methods are used
# * Backtracking Line Search:
# * Choose an initial learning rate ($t_{k} = t_{init})$, and update your parameters ${\mathbf{\theta}^{(k)}} = {\mathbf{\theta}^{(k-1)}} \pm t_{k}.\nabla f({\mathbf{\theta}^{(k-1)}})$
# * Reduce learning rate $t_{k} = \alpha . t_{init}$, where $0< \alpha <1 $
# * Repeat by reducing $\alpha$ till you see an improvmnet in $f({\mathbf{\theta}^{(k)}})$
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Likelihood
#
# We start by writing out a likelihood for linear regression is
#
# $$
# \likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) =
# \prod_{i=1}^N p(y\given\xx,\beta_0,\beta,\sigma^2) =
# \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right\}
# $$
#
# Log-likelihood for linear regression is
#
# $$
# \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) = \sum_{i=1}^N \left[ -\frac{1}{2}\log 2\pi\sigma^2 -\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right].
# $$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Gradient of Log-Likelihood
#
# Partial derivatives
#
# $$
# \begin{aligned}
# \frac{\partial}{\partial \beta_0} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-1)\\
# \frac{\partial}{\partial \beta_k} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-x_k)&,k\in\{1,\dots,p\}
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# Hence gradient (with respect to $\beta$s)
# $$
# \nabla \loglik(\beta_0,\beta,\sigma^2\given\xx,\yy) = \left[\begin{array}{c}
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-1) \\
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_1 + \sum_j x_j \beta_j)\right)(-x_1) \\
# \vdots\\
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-x_p)
# \end{array}
# \right]
# $$
| CourseMaterial/COMP562_Lect3/3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Install Packages
# %pip install seaborn
# %matplotlib inline
# +
import numpy as np
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score,recall_score,precision_score,f1_score,roc_auc_score
from sklearn.preprocessing import RobustScaler
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
sns.set_style("whitegrid")
import pandas as pd
import matplotlib.pyplot as plt
# -
# ## Import CSV Files
df = pd.read_csv("/Users/isa/Desktop/healthcare/aiproject/train_data.csv")
df_test = pd.read_csv("/Users/isa/Desktop/healthcare/aiproject/test_data.csv")
df_test_y = pd.read_csv("/Users/isa/Desktop/healthcare/aiproject/sample_sub.csv")
data_dict = pd.read_csv("/Users/isa/Desktop/healthcare/aiproject/train_data_dictionary.csv")
df.head()
df_test_cp = df_test.copy()
df_test_fe = df_test.copy()
df_test.head()
df_test_y.head()
df["Stay"].nunique()
# ### Merge Dataset
df_test = df_test.merge(df_test_y, how='inner', left_on=['case_id'], right_on=['case_id'])
df_test.info()
# create new data frame
mdf = df.append(df_test)
mdf.case_id.is_unique
# ## DATA PREPROCESSING FOR mdf
object_cols = mdf.select_dtypes(include='object').columns.to_list()
num_cols = mdf.drop(object_cols, axis=1).columns
object_cols.remove('Stay')
# convert object data to numerical using label encoding
les = {}
for col in object_cols:
les[col] = LabelEncoder()
data = mdf[col].values
mdf[col] = les[col].fit_transform(data)
print("{}: {} \n".format(col, les[col].classes_))
# ### Correlation Matrix
fig, ax = plt.subplots(figsize=(12, 12))
sns.heatmap(ax=ax, data=mdf.corr(), cmap="YlGnBu", annot=True, cbar=False)
mdf.isnull().sum()
bed_grade_mean = mdf["Bed Grade"].mean()
city_code_patient_mean = mdf["City_Code_Patient"].mean()
mdf.loc[mdf["Bed Grade"].isnull(), "Bed Grade"] = bed_grade_mean
mdf.loc[mdf["City_Code_Patient"].isnull(), "City_Code_Patient"] = city_code_patient_mean
mdf.isna().sum()
# ## MLOPS
from time import time
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import randint
df_test = df_test.merge(df_test_y, how='inner', left_on=['case_id'], right_on=['case_id'])
df_selection = df.append(df_test)
x = mdf.drop(['Stay', 'case_id', 'patientid'], axis=1)
y = mdf.Stay
x[x.columns] = RobustScaler().fit_transform(x[x.columns].values)
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.3, stratify=y)
# +
values= [RandomForestClassifier(), KNeighborsClassifier(), LogisticRegression(), DecisionTreeClassifier(), GaussianNB()]
keys= ['RandomForsetClassifier',
'KNeighborsClassifier',
'LogisticRegression',
'DecisionTreeClassifier',
'GaussianNB']
models= dict(zip(keys,values))
accuracy_scores=[]
train_times=[]
for key,value in models.items():
t = time()
value.fit(x_train,y_train)
duration = (time() - t) / 60
y_pred= value.predict(x_test)
accuracy= accuracy_score(y_test, y_pred)
accuracy_scores.append(accuracy)
train_times.append(duration)
print(key)
print(round(accuracy * 100, 2))
# +
param_dist = {"max_depth": [3, None],
"max_features": randint(1, 9),
"min_samples_leaf": randint(1, 9),
"criterion": ["gini", "entropy"]}
tree = DecisionTreeClassifier()
# Instantiate the RandomizedSearchCV object: tree_cv
tree_cv = RandomizedSearchCV(tree, param_dist, cv=5)
# Fit it to the data
tree_cv.fit(x_train, y_train)
# -
tree_cv.best_score_
tree_cv.best_estimator_
print("Precision: "+str(precision_score(y_test,y_pred, average='micro')))
print("Recall: "+str(recall_score(y_test,y_pred, average='micro')))
print("F1 Puanı: "+str(precision_score(y_test,y_pred, average='micro')))
print()
print("Sınıflandırma Matrisi: \n "+str(confusion_matrix(y_test,y_pred)))
def f_importances(coef, names, top=-1):
imp = coef
imp, names = zip(*sorted(list(zip(imp, names))))
# Show all features
if top == -1:
top = len(names)
plt.barh(range(top), imp[::-1][0:top], align='center')
plt.yticks(range(top), names[::-1][0:top])
plt.title('feature importances for mdf')
plt.show()
features_names = x_train.columns
f_importances(abs(tree_cv.best_estimator_.feature_importances_), features_names, top=4)
x_train,x_val,y_train,y_val=train_test_split(x,y,test_size=0.25,random_state=0,stratify=y)
dtc=DecisionTreeClassifier()
dtc.fit(x_train,y_train)
y_pred_test=dtc.predict(x_val)
y_pred_train=dtc.predict(x_train)
print("Eğitim için Doğruluk: "+str(accuracy_score(y_train,y_pred_train)))
print("Validasyon için Doğruluk: "+str(accuracy_score(y_val,y_pred_test)))
print()
print("Precision: "+str(precision_score(y_val,y_pred_test,average='micro')))
# print("ROC Eğrisi Altındaki Alan: "+str(roc_auc_score(y_test,y_pred,average='micro')))
print("Recall: "+str(recall_score(y_val,y_pred_test,average='micro')))
print("F1 Puanı: "+str(precision_score(y_val,y_pred_test,average='micro')))
print()
print("Test Sınıflandırma Matrisi: \n "+str(confusion_matrix(y_val,y_pred_test)))
print()
print("Train Sınıflandırma Matrisi: \n "+str(confusion_matrix(y_train,y_pred_train)))
# ## FEATURE SELECTION
from scipy.stats import uniform
from scipy.stats import uniform as sp_randFloat
from scipy.stats import randint as sp_randInt
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
# +
# TRAIN dataset converted to digitized
## alternatively labelEncoder() can be used
# -
bed_grade_mean = df["Bed Grade"].mean()
city_code_patient_mean = df["City_Code_Patient"].mean()
df.loc[df["Bed Grade"].isnull(), "Bed Grade"] = bed_grade_mean
df.loc[df["City_Code_Patient"].isnull(), "City_Code_Patient"] = city_code_patient_mean
df_train = df[
["case_id","Hospital_code","Department", "Age", "Severity of Illness", "Type of Admission", "Stay"]].copy()
# print("TRAIN DATA")
# 0 -> gynecology / 1 -> anesthesia / 2-> radiotherapy / 3 -> TB & Chest disease / 4 -> surgery
# print(df["Department"].value_counts())
df_train = df_train.replace(['gynecology'], '0')
df_train = df_train.replace(['anesthesia'], '1')
df_train = df_train.replace(['radiotherapy'], '2')
df_train = df_train.replace(['TB & Chest disease'], '3')
df_train = df_train.replace(['surgery'], '4')
# print(df_train["Department"].value_counts())
# 0 -> Moderate / 1 -> Minor / 2 -> Extreme / 3 -> Severity of Illness
# print(df["Severity of Illness"].value_counts())
df_train = df_train.replace(['Moderate'], '0')
df_train = df_train.replace(['Minor'], '1')
df_train = df_train.replace(['Extreme'], '2')
# print(df_train["Severity of Illness"].value_counts())
# 0 -> Trauma / 1 -> Emergency / 2 -> Urgent
# print(df["Type of Admission"].value_counts())
df_train = df_train.replace(['Trauma'], '0')
df_train = df_train.replace(['Emergency'], '1')
df_train = df_train.replace(['Urgent'], '2')
# print(df_train["Type of Admission"].value_counts())
# 0 -> 41-50 / 1 -> 31-40 / 2 -> 51-60 / 3 -> 21-30 / 4 -> 71-80 / 5 -> 61-70
# / 6 -> 11-20 / 7 -> 81-90 / 8 -> 0-10 / 9 -> 91-100
# print(df["Age"].value_counts())
df_train = df_train.replace(['41-50'], '0')
df_train = df_train.replace(['31-40'], '1')
df_train = df_train.replace(['51-60'], '2')
df_train = df_train.replace(['21-30'], '3')
df_train = df_train.replace(['71-80'], '4')
df_train = df_train.replace(['61-70'], '5')
df_train = df_train.replace(['11-20'], '6')
df_train = df_train.replace(['81-90'], '7')
df_train = df_train.replace(['0-10'], '8')
df_train = df_train.replace(['91-100'], '9')
# print(df_train["Age"].value_counts())
# 0 -> 21-30 / 1 -> 11-20 / 2 -> 31-40 / 3 -> 51-60 / 4 -> 0-10 / 5 -> 41-50
# 6 -> 71-80 / 7 -> More than 100 Days / 8 -> 81-90 / 9 -> 91-100 / 10 -> 61-70
# print(df["Stay"].value_counts())
df_train = df_train.replace(['21-30'], '0')
df_train = df_train.replace(['11-20'], '1')
df_train = df_train.replace(['31-40'], '2')
df_train = df_train.replace(['51-60'], '3')
df_train = df_train.replace(['0-10'], '4')
df_train = df_train.replace(['41-50'], '5')
df_train = df_train.replace(['71-80'], '6')
df_train = df_train.replace(['More than 100 Days'], '7')
df_train = df_train.replace(['81-90'], '8')
df_train = df_train.replace(['91-100'], '9')
df_train = df_train.replace(['61-70'], '10')
# print(df_train["Stay"].value_counts())
# +
# TEST dataset converted to digitized
## alternatively labelEncoder() can be used
# -
bed_grade_mean = df_test_cp["Bed Grade"].mean()
city_code_patient_mean = df_test_cp["City_Code_Patient"].mean()
df_test_cp.loc[df_test_cp["Bed Grade"].isnull(), "Bed Grade"] = bed_grade_mean
df_test_cp.loc[df_test_cp["City_Code_Patient"].isnull(), "City_Code_Patient"] = city_code_patient_mean
df_test_cp = df_test_cp[
["case_id","Hospital_code","Department", "Age", "Severity of Illness", "Type of Admission"]].copy()
# 0 -> gynecology / 1 -> anesthesia / 2-> radiotherapy / 3 -> TB & Chest disease / 4 -> surgery
# print(df_test_cp["Department"].value_counts())
df_test_cp = df_test_cp.replace(['gynecology'], '0')
df_test_cp = df_test_cp.replace(['anesthesia'], '1')
df_test_cp = df_test_cp.replace(['radiotherapy'], '2')
df_test_cp = df_test_cp.replace(['TB & Chest disease'], '3')
df_test_cp = df_test_cp.replace(['surgery'], '4')
# print(df_test_cp["Department"].value_counts())
# 0 -> Moderate / 1 -> Minor / 2 -> Extreme / 3 -> Severity of Illness
# print(df_test_cp["Severity of Illness"].value_counts())
df_test_cp = df_test_cp.replace(['Moderate'], '0')
df_test_cp = df_test_cp.replace(['Minor'], '1')
df_test_cp = df_test_cp.replace(['Extreme'], '2')
# print(df_test_cp["Severity of Illness"].value_counts())
# 0 -> Trauma / 1 -> Emergency / 2 -> Urgent
# print(df_test_cp["Type of Admission"].value_counts())
df_test_cp = df_test_cp.replace(['Trauma'], '0')
df_test_cp = df_test_cp.replace(['Emergency'], '1')
df_test_cp = df_test_cp.replace(['Urgent'], '2')
# print(df_test_cp["Type of Admission"].value_counts())
# 0 -> 41-50 / 1 -> 31-40 / 2 -> 51-60 / 3 -> 21-30 / 4 -> 71-80 / 5 -> 61-70
# / 6 -> 11-20 / 7 -> 81-90 / 8 -> 0-10 / 9 -> 91-100
# print(df_test_cp["Age"].value_counts())
df_test_cp = df_test_cp.replace(['41-50'], '0')
df_test_cp = df_test_cp.replace(['31-40'], '1')
df_test_cp = df_test_cp.replace(['51-60'], '2')
df_test_cp = df_test_cp.replace(['21-30'], '3')
df_test_cp = df_test_cp.replace(['71-80'], '4')
df_test_cp = df_test_cp.replace(['61-70'], '5')
df_test_cp = df_test_cp.replace(['11-20'], '6')
df_test_cp = df_test_cp.replace(['81-90'], '7')
df_test_cp = df_test_cp.replace(['0-10'], '8')
df_test_cp = df_test_cp.replace(['91-100'], '9')
# print(df_test_cp["Age"].value_counts())
df_test_cp = df_test_cp.merge(df_test_y, how='inner', left_on=['case_id'], right_on=['case_id'])
df_sn = df_train.append(df_test_cp)
x = df_sn.drop(['Stay', 'case_id'], axis=1)
y = df_sn.Stay
x[x.columns] = RobustScaler().fit_transform(x[x.columns].values)
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.3, stratify=y)
# +
values= [RandomForestClassifier(), KNeighborsClassifier(), LogisticRegression(), DecisionTreeClassifier(), GaussianNB()]
keys= ['RandomForsetClassifier',
'KNeighborsClassifier',
'LogisticRegression',
'DecisionTreeClassifier',
'GaussianNB']
models= dict(zip(keys,values))
accuracy_scores=[]
train_times=[]
for key,value in models.items():
t = time()
value.fit(x_train,y_train)
duration = (time() - t) / 60
y_pred= value.predict(x_test)
accuracy= accuracy_score(y_test, y_pred)
accuracy_scores.append(accuracy)
train_times.append(duration)
print(key)
print(round(accuracy * 100, 2))
# -
# ## FEATURE EXTRACTION
# +
# -----------------------------FEATURE EXTRACTION WITH TRAIN --------------------------------------
## Severity of Illness -> Extreme and Age -> 61... if greater then create column named priority and 1 will be set
## otherwise priority status will be 0
# -
df_copy_train = df_train
options_sol = ['2']
rslt_df = df_copy_train.loc[df_copy_train['Severity of Illness'].isin(options_sol)]
options_age = ['4', '5', '7', '9']
rslt_df_age = df_copy_train.loc[df_copy_train['Age'].isin(options_age)]
df_feature_ext = df_copy_train.copy()
print("rslt_df size:" + str(rslt_df.shape))
common = rslt_df.merge(rslt_df_age, left_index=True, right_index=True, how='outer', suffixes=('', '_drop'))
common.drop(common.filter(regex='_y$').columns.tolist(), axis=1, inplace=False)
# print("merged two column : ", common["Stay"])
# print(common.isnull().sum())
common.loc[common["case_id"].isnull(), "case_id"] = "0"
common.loc[common["Hospital_code"].isnull(), "Hospital_code"] = "0"
common.loc[common["Department"].isnull(), "Department"] = "0"
common.loc[common["Age"].isnull(), "Age"] = "0"
common.loc[common["Severity of Illness"].isnull(), "Severity of Illness"] = "0"
common.loc[common["Type of Admission"].isnull(), "Type of Admission"] = "0"
common.loc[common["Stay"].isnull(), "Stay"] = "0"
# print(common.isnull().sum())
f = open("train_join.csv", "w")
f.write("case_id,Hospital_code,Department,Age,Severity of Illness,Type of Admission,priority,Stay\n")
for (i, row) in common.iterrows():
if common["Hospital_code"][i] == "0" and common["Department"][i] == "0" and \
common["Age"][i] == "0" and common["Severity of Illness"][i] == "0" and common["Type of Admission"][
i] == "0" and common["Stay"][i] == "0":
row["case_id"] = df_copy_train["case_id"][i]
row["Hospital_code"] = df_copy_train["Hospital_code"][i]
row["Department"] = df_copy_train["Department"][i]
row["Age"] = df_copy_train["Age"][i]
row["Severity of Illness"] = df_copy_train["Severity of Illness"][i]
row["Type of Admission"] = df_copy_train["Type of Admission"][i]
row["Stay"] = df_copy_train["Stay"][i]
# row["priority"] = "NO"
row["priority"] = "0"
else:
# row["priority"] = "YES"
row["priority"] = "1"
f.write(str(row["case_id"]) + "," + str(row["Hospital_code"]) + "," + str(row["Department"]) + "," + str(
row["Age"]) + "," + str(row["Severity of Illness"]) + "," + str(row["Type of Admission"]) + ","
+ str(row["priority"]) + "," + str(row["Stay"]) + "\n")
file = open("train_join.csv", "r")
df_common = pd.read_csv(file)
df_common.head()
# +
# -----------------------------FEATURE EXTRACTION WITH TEST --------------------------------------
## Severity of Illness -> Extreme and Age -> 61... if greater then create column named priority and 1 will be set
## otherwise priority status will be 0
# -
df_test_fe = df_test_cp.copy()
options_sol = ['2']
rslt_df_test = df_test_fe.loc[df_test_fe['Severity of Illness'].isin(options_sol)]
# print('\nResult Severity of Illness :\n', rslt_df_test)
options_age = ['4', '5', '7', '9']
rslt_df_test_age = df_test_fe.loc[df_test_fe['Age'].isin(options_age)]
# print('\nResult Age :\n', rslt_df_test_age)
common = rslt_df_test.merge(rslt_df_test_age, left_index=True, right_index=True, how='outer',
suffixes=('', '_drop'))
common.drop(common.filter(regex='_y$').columns.tolist(), axis=1, inplace=False)
common.loc[common["case_id"].isnull(), "case_id"] = "0"
common.loc[common["Hospital_code"].isnull(), "Hospital_code"] = "0"
common.loc[common["Department"].isnull(), "Department"] = "0"
common.loc[common["Age"].isnull(), "Age"] = "0"
common.loc[common["Severity of Illness"].isnull(), "Severity of Illness"] = "0"
common.loc[common["Type of Admission"].isnull(), "Type of Admission"] = "0"
# print(common.isnull().sum())
f = open("test_join.csv", "w")
f.write("case_id,Hospital_code,Department,Age,Severity of Illness,Type of Admission,priority\n")
for (i, row) in common.iterrows():
if common["Hospital_code"][i] == "0" and common["Department"][i] == "0" and \
common["Age"][i] == "0" and common["Severity of Illness"][i] == "0" and common["Type of Admission"][
i] == "0":
row["case_id"] = df_test_fe["case_id"][i]
row["Hospital_code"] = df_test_fe["Hospital_code"][i]
row["Department"] = df_test_fe["Department"][i]
row["Age"] = df_test_fe["Age"][i]
row["Severity of Illness"] = df_test_fe["Severity of Illness"][i]
row["Type of Admission"] = df_test_fe["Type of Admission"][i]
# row["priority"] = "NO"
row["priority"] = "0"
else:
# row["priority"] = "YES"
row["priority"] = "1"
f.write(str(row["case_id"]) + "," + str(row["Hospital_code"]) + "," + str(row["Department"]) + "," + str(
row["Age"]) + "," + str(row["Severity of Illness"]) + "," + str(row["Type of Admission"]) + ","
+ str(row["priority"]) + "\n")
file_test = open("test_join.csv", "r")
df_test_common = pd.read_csv(file_test)
df_test_common.head(10)
# ## MLOPS with F.Extraction
df_test_fe = df_test_fe.merge(df_test_y, how='inner', left_on=['case_id'], right_on=['case_id'])
df_feat = df_train.append(df_test_cp)
x = df_feat.drop(['Stay', 'case_id'], axis=1)
y = df_feat.Stay
x[x.columns] = RobustScaler().fit_transform(x[x.columns].values)
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.3, stratify=y)
# +
values= [RandomForestClassifier(), KNeighborsClassifier(), LogisticRegression(), DecisionTreeClassifier(), GaussianNB()]
keys= ['RandomForsetClassifier',
'KNeighborsClassifier',
'LogisticRegression',
'DecisionTreeClassifier',
'GaussianNB']
models= dict(zip(keys,values))
accuracy_scores=[]
train_times=[]
for key,value in models.items():
t = time()
value.fit(x_train,y_train)
duration = (time() - t) / 60
y_pred= value.predict(x_test)
accuracy= accuracy_score(y_test, y_pred)
accuracy_scores.append(accuracy)
train_times.append(duration)
print(key)
print(round(accuracy * 100, 2))
# -
# ## VALIDATION
x_train,x_val,y_train,y_val=train_test_split(x,y,test_size=0.25,random_state=0,stratify=y)
dtc=DecisionTreeClassifier()
dtc.fit(x_train,y_train)
y_pred_test=dtc.predict(x_val)
y_pred_train=dtc.predict(x_train)
print("Eğitim için Doğruluk: "+str(accuracy_score(y_train,y_pred_train)))
print("Validasyon için Doğruluk: "+str(accuracy_score(y_val,y_pred_test)))
print()
print("Precision: "+str(precision_score(y_val,y_pred_test,average='micro')))
# print("ROC Eğrisi Altındaki Alan: "+str(roc_auc_score(y_test,y_pred,average='micro')))
print("Recall: "+str(recall_score(y_val,y_pred_test,average='micro')))
print("F1 Puanı: "+str(precision_score(y_val,y_pred_test,average='micro')))
print()
print("Test Sınıflandırma Matrisi: \n "+str(confusion_matrix(y_val,y_pred_test)))
print()
print("Train Sınıflandırma Matrisi: \n "+str(confusion_matrix(y_train,y_pred_train)))
| .ipynb_checkpoints/aiproject-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis, LinearDiscriminantAnalysis
from sklearn.pipeline import Pipeline
from tqdm import tqdm_notebook
import warnings
import multiprocessing
from scipy.optimize import minimize
import time
warnings.filterwarnings('ignore')
# STEP 2
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
cols = [c for c in train.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
print(train.shape, test.shape)
# STEP 3
oof = np.zeros(len(train))
preds = np.zeros(len(test))
for i in tqdm_notebook(range(512)):
train2 = train[train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train2.index; idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]; test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=42)
for train_index, test_index in skf.split(train2, train2['target']):
clf = QuadraticDiscriminantAnalysis(0.5)
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
oof[idx1[test_index]] = clf.predict_proba(train3[test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof)
print(f'AUC: {auc:.5}')
# STEP 4
for itr in range(4):
test['target'] = preds
test.loc[test['target'] > 0.955, 'target'] = 1 # initial 94
test.loc[test['target'] < 0.045, 'target'] = 0 # initial 06
usefull_test = test[(test['target'] == 1) | (test['target'] == 0)]
new_train = pd.concat([train, usefull_test]).reset_index(drop=True)
print(usefull_test.shape[0], "Test Records added for iteration : ", itr)
new_train.loc[oof > 0.995, 'target'] = 1 # initial 98
new_train.loc[oof < 0.005, 'target'] = 0 # initial 02
oof2 = np.zeros(len(train))
preds = np.zeros(len(test))
for i in tqdm_notebook(range(512)):
train2 = new_train[new_train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train[train['wheezy-copper-turtle-magic']==i].index
idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]
test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=time.time)
for train_index, test_index in skf.split(train2, train2['target']):
oof_test_index = [t for t in test_index if t < len(idx1)]
clf = QuadraticDiscriminantAnalysis(0.5)
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
if len(oof_test_index) > 0:
oof2[idx1[oof_test_index]] = clf.predict_proba(train3[oof_test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof2)
print(f'AUC: {auc:.5}')
# STEP 5
sub1 = pd.read_csv('../input/sample_submission.csv')
sub1['target'] = preds
# sub.to_csv('submission.csv',index=False)
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis, LinearDiscriminantAnalysis
from sklearn.pipeline import Pipeline
from tqdm import tqdm_notebook
import warnings
import multiprocessing
from scipy.optimize import minimize
warnings.filterwarnings('ignore')
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
cols = [c for c in train.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
print(train.shape, test.shape)
oof = np.zeros(len(train))
preds = np.zeros(len(test))
for i in tqdm_notebook(range(512)):
train2 = train[train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train2.index; idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]; test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=42)
for train_index, test_index in skf.split(train2, train2['target']):
clf = QuadraticDiscriminantAnalysis(0.5)
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
oof[idx1[test_index]] = clf.predict_proba(train3[test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof)
print(f'AUC: {auc:.5}')
for itr in range(4):
test['target'] = preds
test.loc[test['target'] > 0.94, 'target'] = 1
test.loc[test['target'] < 0.06, 'target'] = 0
usefull_test = test[(test['target'] == 1) | (test['target'] == 0)]
new_train = pd.concat([train, usefull_test]).reset_index(drop=True)
print(usefull_test.shape[0], "Test Records added for iteration : ", itr)
new_train.loc[oof > 0.98, 'target'] = 1
new_train.loc[oof < 0.02, 'target'] = 0
oof2 = np.zeros(len(train))
preds = np.zeros(len(test))
for i in tqdm_notebook(range(512)):
train2 = new_train[new_train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train[train['wheezy-copper-turtle-magic']==i].index
idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]
test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=42)
for train_index, test_index in skf.split(train2, train2['target']):
oof_test_index = [t for t in test_index if t < len(idx1)]
clf = QuadraticDiscriminantAnalysis(0.5)
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
if len(oof_test_index) > 0:
oof2[idx1[oof_test_index]] = clf.predict_proba(train3[oof_test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof2)
print(f'AUC: {auc:.5}')
sub2 = pd.read_csv('../input/sample_submission.csv')
sub2['target'] = preds
# sub.to_csv('submission.csv',index=False)
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis, LinearDiscriminantAnalysis
from sklearn.pipeline import Pipeline
from tqdm import tqdm_notebook
import warnings
import multiprocessing
from scipy.optimize import minimize
import time
from sklearn.model_selection import GridSearchCV, train_test_split
warnings.filterwarnings('ignore')
# STEP 2
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
cols = [c for c in train.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
print(train.shape, test.shape)
# STEP 3
oof = np.zeros(len(train))
preds = np.zeros(len(test))
params = [{'reg_param': [0.1, 0.2, 0.3, 0.4, 0.5]}]
# 512 models
reg_params = np.zeros(512)
for i in tqdm_notebook(range(512)):
train2 = train[train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train2.index; idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]; test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=42)
for train_index, test_index in skf.split(train2, train2['target']):
qda = QuadraticDiscriminantAnalysis()
clf = GridSearchCV(qda, params, cv=4)
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
reg_params[i] = clf.best_params_['reg_param']
oof[idx1[test_index]] = clf.predict_proba(train3[test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof)
print(f'AUC: {auc:.5}')
# STEP 4
for itr in range(10):
test['target'] = preds
test.loc[test['target'] > 0.955, 'target'] = 1 # initial 94
test.loc[test['target'] < 0.045, 'target'] = 0 # initial 06
usefull_test = test[(test['target'] == 1) | (test['target'] == 0)]
new_train = pd.concat([train, usefull_test]).reset_index(drop=True)
print(usefull_test.shape[0], "Test Records added for iteration : ", itr)
new_train.loc[oof > 0.995, 'target'] = 1 # initial 98
new_train.loc[oof < 0.005, 'target'] = 0 # initial 02
oof2 = np.zeros(len(train))
preds = np.zeros(len(test))
for i in tqdm_notebook(range(512)):
train2 = new_train[new_train['wheezy-copper-turtle-magic']==i]
test2 = test[test['wheezy-copper-turtle-magic']==i]
idx1 = train[train['wheezy-copper-turtle-magic']==i].index
idx2 = test2.index
train2.reset_index(drop=True,inplace=True)
data = pd.concat([pd.DataFrame(train2[cols]), pd.DataFrame(test2[cols])])
pipe = Pipeline([('vt', VarianceThreshold(threshold=2)), ('scaler', StandardScaler())])
data2 = pipe.fit_transform(data[cols])
train3 = data2[:train2.shape[0]]
test3 = data2[train2.shape[0]:]
skf = StratifiedKFold(n_splits=11, random_state=time.time)
for train_index, test_index in skf.split(train2, train2['target']):
oof_test_index = [t for t in test_index if t < len(idx1)]
clf = QuadraticDiscriminantAnalysis(reg_params[i])
clf.fit(train3[train_index,:],train2.loc[train_index]['target'])
if len(oof_test_index) > 0:
oof2[idx1[oof_test_index]] = clf.predict_proba(train3[oof_test_index,:])[:,1]
preds[idx2] += clf.predict_proba(test3)[:,1] / skf.n_splits
auc = roc_auc_score(train['target'], oof2)
print(f'AUC: {auc:.5}')
# STEP 5
sub3 = pd.read_csv('../input/sample_submission.csv')
sub3['target'] = preds
# sub.to_csv('submission.csv',index=False)
# -
sub1.head()
sub2.head()
sub3.head()
sub = pd.read_csv('../input/sample_submission.csv')
sub.head()
sub['target'] = 0.5*sub1.target + 0.3*sub2.target + 0.2*sub3.target
sub.to_csv('submission1.csv', index = False)
sub['target'] = 0.2*sub1.target + 0.3*sub2.target + 0.5*sub3.target
sub.to_csv('submission2.csv', index = False)
sub['target'] = 0.2*sub1.target + 0.4*sub2.target + 0.4*sub3.target
sub.to_csv('submission3.csv', index = False)
| Kaggle_Instant_Gratification/script/instant-gratification-ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Solving M Models without Loops
# A type III loop is a cycle with no exchange of metabolites. Therefore, type III loops do not have flux through exchange, demand and biomass reactions, which exchange mass with the system. If flux flows through a type III loop, it has no thermodynamic driving force, which is not realistic. As long as the reaction bounds do not force flux through any reactions in the loop, a solution with flux through a loop can be collapsed into a solution without it which still allows flux through the biomass.
#
# Proof: Take the following model, where all fluxes are positive or zero (conversion of a model with negative fluxes to this form is trivial), with a flux solution $v$. All exchange, demand, and biomass, reactions (which by definition can not be a part of a loop) are denoted with $_i$ subscripts, while all others are denoted with $_j$.
# $$ \mathbf S \cdot v = 0$$
# $$ ub \ge v \ge 0 $$
# $$ ub_i \ge v_i \ge lb_i $$
#
# Let $v_L$ be any flux loop within $v$. Because exchange, demand, and biomass can not be in the loop, we have the following:
# $$v_{L,i} = 0 $$
#
# By definition, no metabolites are exchanged, so
# $$ \mathbf S \cdot v_L = 0 $$
# $$ \mathbf S \cdot (v - v_L) = 0 $$
#
# Because $v_L$ is a loop within $v$, the following are true:
# $$ ub \ge v \ge v_L \ge 0 $$
# $$ ub \ge v - v_L \ \ge 0 $$
#
# $\therefore$ $\forall v$ and $\forall v_L$ within it, $v - v_L$ is also a valid solution which will produce biomass flux.
# This proof does not apply to models where flux is forced through a reaction (i.e. where the lower bound is greater than 0). To optain a flux state with no flux through thermodynamically infeasible loops for all models, loopless FBA was developed by [Schellenberger et al.](http://dx.doi.org/10.1016/j.bpj.2010.12.3707) To compute this for all of these correctly parsed-models, we use the loopless FBA [implementation](http://cobrapy.readthedocs.org/en/latest/loopless.html) in [cobrapy](http://opencobra.github.io/cobrapy/) (version 0.4.0b1 or later) and the [gurobi](http://www.gurobi.com) solver (version 6.0). Because loopless FBA is an MILP, it is not guarenteed to find the optimum, but for every model we are able to find a feasible loopless solution with the algorithm.
# +
import os
import pandas
import cobra
pandas.set_option("display.max_rows", 100)
# -
# Load in the models, as parsed by this [script](load_models.ipynb). We will also break up all reversible reactions into the model into two irreversible reactions in opposite directions.
models = cobra.DictList()
for filename in sorted(os.listdir("sbml3")):
if not filename.endswith(".xml"):
continue
models.append(cobra.io.read_sbml_model(os.path.join("sbml3", filename)))
# Construct the loopless models, which impose additional constraints which prevent flux from going around one of these loops.
loopless_models = cobra.DictList((cobra.flux_analysis.construct_loopless_model(m) for m in models))
# Solve the models using gurobi, allowing the branch-and-bound solver to proceed for two minutes.
loopless_solutions = {}
for loopless_model in loopless_models:
solution = loopless_model.optimize(solver="gurobi", time_limit=120)
loopless_solutions[loopless_model.id] = solution
# Constrain the models based the loopless result. Only reactions which carry flux in the loopless solution will be allowed to carry flux.
constrained_loopless_models = cobra.DictList()
for model, loopless_model in zip(models, loopless_models):
constrained = model.copy()
constrained_loopless_models.append(constrained)
for reaction in constrained.reactions:
loopless_flux = loopless_model.reactions.get_by_id(reaction.id).x
reaction.upper_bound = min(reaction.upper_bound, loopless_flux + 1e-6)
# Solve these problems which are constrained to be loopless exactly.
loopless_exact_solutions = {
model.id: model.optimize(solver="esolver", rational_solution=True)
for model in constrained_loopless_models}
# Format the results as a table. This demonstrates these models have a feasible flux state which produce biomass without using these loops, and this loopless flux state can also be solved using exact arithmetic.
results = pandas.DataFrame.from_dict(
{model.id: {"loopless": loopless_solutions[model.id].f,
"FBA": model.optimize().f,
"constrained looples exact": str(loopless_exact_solutions[model.id].f)}
for model in models},
orient="index")
results
| loopless_m_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import torch
def read_file(fname: str) -> pd.DataFrame:
"""Reads a filename and formats it properly for simpletransformers"""
df = pd.read_table(fname, sep="\t", header=None, names="text,labels,role".split(","))
offensive_ids = df.labels != "Acceptable speech"
df.labels[offensive_ids] = 1
df.labels[~offensive_ids] = 0
df["labels"] = df.labels.astype(np.int8)
df = df.drop(columns=["role"])
return df
# -
# # English
# +
# %%time
train_fname = "../data/lgbt-en.train.tsv"
test_fname = "../data/lgbt-en.test.tsv"
train = read_file(train_fname)
test = read_file(test_fname)
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer(ngram_range=(1,3))
X_train_counts = count_vect.fit_transform(train.text.values)
from sklearn.feature_extraction.text import TfidfTransformer
#tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
#X_train_tf = tf_transformer.transform(X_train_counts)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
from sklearn.svm import SVC
clf = SVC().fit(X=X_train_tfidf, y=train.labels)
docs_new = test.text.values.tolist()
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
from sklearn.metrics import accuracy_score, f1_score
y_true = test["labels"]
accuracy = accuracy_score(y_true, predicted)
print("Accuracy: ", accuracy)
f1 = f1_score(y_true, predicted)
print("F1 score: ", f1)
# -
# # Slovenian
# +
# %%time
train_fname = "../data/lgbt-sl.train.tsv"
test_fname = "../data/lgbt-sl.test.tsv"
train = read_file(train_fname)
test = read_file(test_fname)
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer(ngram_range=(1,3))
X_train_counts = count_vect.fit_transform(train.text.values)
from sklearn.feature_extraction.text import TfidfTransformer
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
from sklearn.svm import SVC
clf = SVC().fit(X=X_train_tfidf, y=train.labels)
docs_new = test.text.values.tolist()
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
from sklearn.metrics import accuracy_score, f1_score
y_true = test["labels"]
accuracy = accuracy_score(y_true, predicted)
print("Accuracy: ", accuracy)
f1 = f1_score(y_true, predicted)
print("F1 score: ", f1)
# -
# # Croatian
# +
# %%time
train_fname = "../data/lgbt-hr.train.tsv"
test_fname = "../data/lgbt-hr.test.tsv"
train = read_file(train_fname)
test = read_file(test_fname)
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer(ngram_range=(1,3))
X_train_counts = count_vect.fit_transform(train.text.values)
from sklearn.feature_extraction.text import TfidfTransformer
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
from sklearn.svm import SVC
clf = SVC().fit(X=X_train_tfidf, y=train.labels)
docs_new = test.text.values.tolist()
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
from sklearn.metrics import accuracy_score, f1_score
y_true = test["labels"]
accuracy = accuracy_score(y_true, predicted)
print("Accuracy: ", accuracy)
f1 = f1_score(y_true, predicted)
print("F1 score: ", f1)
# -
# # Adding dummy classifier data
# +
from sklearn.dummy import DummyClassifier
languages = ["en", "sl", "hr"]
for strategy in {"stratified", "most_frequent", "prior", "uniform"}:
results = dict()
for lang in languages:
train_fname = f"../data/lgbt-{lang}.train.tsv"
test_fname = f"../data/lgbt-{lang}.test.tsv"
train = read_file(train_fname)
test = read_file(test_fname)
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer(ngram_range=(1,3))
X_train_counts = count_vect.fit_transform(train.text.values)
from sklearn.feature_extraction.text import TfidfTransformer
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
from sklearn.svm import SVC
clf = DummyClassifier(strategy=strategy)
clf.fit(X=X_train_tfidf, y=train.labels)
docs_new = test.text.values.tolist()
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
from sklearn.metrics import accuracy_score, f1_score
y_true = test["labels"]
accuracy = accuracy_score(y_true, predicted)
f1 = f1_score(y_true, predicted)
results[lang] = {"acc": accuracy, "f1": f1}
print(f"""Strategy: {strategy}
| language | accuracy | f1 |
|---|---|---|""")
for lang in languages:
print(f"|{lang}| {results[lang].get('acc', -1.0):0.3} | {results[lang].get('f1', -1.0):0.3} |")
# -
# # Word n-grams
# +
from sklearn.dummy import DummyClassifier
languages = ["en", "sl", "hr"]
results = dict()
for lang in languages:
train_fname = f"../data/lgbt-{lang}.train.tsv"
test_fname = f"../data/lgbt-{lang}.test.tsv"
train = read_file(train_fname)
test = read_file(test_fname)
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer(ngram_range=(3,7), analyzer="char")
X_train_counts = count_vect.fit_transform(train.text.values)
from sklearn.feature_extraction.text import TfidfTransformer
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
from sklearn.svm import SVC
clf = SVC().fit(X=X_train_tfidf, y=train.labels)
docs_new = test.text.values.tolist()
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
from sklearn.metrics import accuracy_score, f1_score
y_true = test["labels"]
accuracy = accuracy_score(y_true, predicted)
f1 = f1_score(y_true, predicted)
results[lang] = {"acc": accuracy, "f1": f1}
print(f"""
| language | accuracy | f1 |
|---|---|---|""")
for lang in languages:
print(f"|{lang}| {results[lang].get('acc', -1.0):0.3} | {results[lang].get('f1', -1.0):0.3} |")
# -
| sklearn count vectorizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## For Checking GPU properties.
# + id="RrtEYeollPWz"
# # !nvidia-smi
# -
# ## Important libraries to Install before start
# + id="-rCPJjq8pALX"
# # !pip install overrides==3.1.0
# # !pip install allennlp==0.8.4
# # !pip install pytorch-pretrained-bert
# # !pip install transformers==4.4.2
# # !pip install entmax
# -
# ## Static arguments and other model hyper-parameters declaration.
# + id="fHnawMktqM--"
args = {}
args['origin_path'] = '/content/drive/MyDrive/Thesis/DeepDifferentialDiagnosis/data/MIMIC-III/mimic-iii-clinical-database-1.4/'
args['out_path'] = '/content/drive/MyDrive/Thesis/DeepDifferentialDiagnosis/data/25.10.2021-Old-Compare/test/'
args['min_sentence_len'] = 3
args['random_seed'] = 1
args['vocab'] = '%s%s.csv' % (args['out_path'], 'vocab')
args['vocab_min'] = 3
args['Y'] = 'full' #'50'/'full'
args['data_path'] = '%s%s_%s.csv' % (args['out_path'], 'train', args['Y']) #50/full
args['version'] = 'mimic3'
args['model'] = 'KG_MultiResCNN' #'KG_MultiResCNNLSTM','bert_we_kg'
args['gpu'] = 0
args['embed_file'] = '%s%s_%s.embed' % (args['out_path'], 'processed', 'full')
args['use_ext_emb'] = False
args['dropout'] = 0.2
args['num_filter_maps'] = 50
args['conv_layer'] = 2
args['filter_size'] = '3,5,7,9,13,15,17,23,29' #'3,5,9,15,19,25', '3,5,7,9,13,15,17,23,29'
args['test_model'] = None
args['weight_decay'] = 0 #Adam, 0.01 #AdamW
args['lr'] = 0.001 #0.0005, 0.001, 0.00146 best 3 for adam and Adamw, 1e-5 for Bert
args['tune_wordemb'] = True
args['MAX_LENGTH'] = 3000 #2500, 512 for bert #1878 is the avg length and max length is 10504 for only discharge summary. 238438 is the max length, 3056 is the avg length combined DS+PHY+NUR
args['batch_size'] = 6 #8,16
args['n_epochs'] = 15
args['MODEL_DIR'] = '/content/drive/MyDrive/Thesis/DeepDifferentialDiagnosis/data/model_output'
args['criterion'] = 'prec_at_8'
args['for_test'] = False
args['bert_dir'] = '/content/drive/MyDrive/Thesis/DeepDifferentialDiagnosis/data/Bert/'
args['pretrained_bert'] = 'bert-base-uncased' # 'emilyalsentzer/Bio_ClinicalBERT''bert-base-uncased' 'dmis-lab/biobert-base-cased-v1.1'
args['instance_count'] = 'full' #if not full then the number specified here will be the number of samples.
args['graph_embedding_file'] = '/home/pgoswami/DifferentialEHR/data/Pytorch-BigGraph/wikidata_translation_v1.tsv.gz'
args['entity_dimention'] = 200 #pytorch biggraph entity has dimention size of 200
# args['entity_selected'] = 5
args['MAX_ENT_LENGTH'] = 30 #mean value is 27.33, for DS+PY+NR max 49, avg 29
args['use_embd_layer'] = True
args['add_with_wordrap'] = True
args['step_size'] = 8
args['gamma'] = 0.1
args['patience'] = 10 #if does not improve result for 5 epochs then break.
args['use_schedular'] = True
args['grad_clip'] = False
args['use_entmax15'] = False
args['use_sentiment']=False
args['sentiment_bert'] = 'siebert/sentiment-roberta-large-english'
args['use_tfIdf'] = True
args['use_proc_label'] = True
args['notes_type'] = 'Discharge summary' # 'Discharge summary,Nursing,Physician ' / 'Nursing,Physician '
args['comment'] = """ My changes with 3000 token+30 ent embed. Tf-idf weight. Diag_ICD+Prod_ICD used, 50 codes """
args['save_everything'] = True
# -
# ## Data Processing ##
# +
#wikidump creation process and indexing
# import wikimapper
# wikimapper.download_wikidumps(dumpname="enwiki-latest", path="/home/pgoswami/DifferentialEHR/data/Wikidata_dump/")
# wikimapper.create_index(dumpname="enwiki-latest",path_to_dumps="/home/pgoswami/DifferentialEHR/data/Wikidata_dump/",
# path_to_db= "/home/pgoswami/DifferentialEHR/data/Wikidata_dump/index_enwiki-latest.db")
# +
# Pytorch Biggraph pre-trained embedding file downloaded from
#https://github.com/facebookresearch/PyTorch-BigGraph#pre-trained-embeddings
# to '/home/pgoswami/DifferentialEHR/data/Pytorch-BigGraph/wikidata_translation_v1.tsv.gz'
# -
class ProcessedIter(object):
def __init__(self, Y, filename):
self.filename = filename
def __iter__(self):
with open(self.filename) as f:
r = csv.reader(f)
next(r)
for row in r:
yield (row[2].split()) #after group-by with subj_id and hadm_id, text is in 3rd column
# +
import pandas as pd
import numpy as np
from collections import Counter, defaultdict
import csv
import sys
import operator
# import operator
from scipy.sparse import csr_matrix
from tqdm import tqdm
import gensim.models
import gensim.models.word2vec as w2v
import gensim.models.fasttext as fasttext
import nltk
nltk.download('punkt')
from nltk.tokenize import RegexpTokenizer
nlp_tool = nltk.data.load('tokenizers/punkt/english.pickle')
tokenizer = RegexpTokenizer(r'\w+')
import re
from transformers import pipeline #for entity extraction
from wikimapper import WikiMapper #creating wikidata entity id
import pickle
import smart_open as smart
class DataProcessing:
def __init__(self, args):
# step 1: process code-related files
dfdiag = pd.read_csv(args['origin_path']+'DIAGNOSES_ICD.csv')
if args['use_proc_label']:
dfproc = pd.read_csv(args['origin_path']+'PROCEDURES_ICD.csv')
dfdiag['absolute_code'] = dfdiag.apply(lambda row: str(self.reformat(str(row[4]), True)), axis=1)
if args['use_proc_label']:
dfproc['absolute_code'] = dfproc.apply(lambda row: str(self.reformat(str(row[4]), False)), axis=1)
dfcodes = pd.concat([dfdiag, dfproc]) if args['use_proc_label'] else dfdiag
dfcodes.to_csv(args['out_path']+'ALL_CODES.csv', index=False,
columns=['ROW_ID', 'SUBJECT_ID', 'HADM_ID', 'SEQ_NUM', 'absolute_code'],
header=['ROW_ID', 'SUBJECT_ID', 'HADM_ID', 'SEQ_NUM', 'ICD9_CODE']) #columns: 'ROW_ID', 'SUBJECT_ID', 'HADM_ID', 'SEQ_NUM', 'ICD9_CODE'
print("unique ICD9 code: {}".format(len(dfcodes['absolute_code'].unique())))
del dfcodes
if args['use_proc_label']:
del dfproc
del dfdiag
# step 2: process notes
# min_sentence_len = 3
disch_full_file = self.write_discharge_summaries(args['out_path']+'disch_full_acc.csv', args['min_sentence_len'], args['origin_path']+'NOTEEVENTS.csv')
dfnotes = pd.read_csv(args['out_path']+'disch_full_acc.csv')
dfnotes = dfnotes.sort_values(['SUBJECT_ID', 'HADM_ID'])
dfnotes = dfnotes.drop_duplicates()
dfnotes = dfnotes.groupby(['SUBJECT_ID','HADM_ID']).apply(lambda x: pd.Series({'TEXT':' '.join(str(v) for v in x.TEXT)})).reset_index()
dfnotes.to_csv(args['out_path']+'disch_full.csv', index=False) #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT'
# step 3: filter out the codes that not emerge in notes
subj_ids = set(dfnotes['SUBJECT_ID'])
self.code_filter(args['out_path'], subj_ids)
dfcodes_filtered = pd.read_csv(args['out_path']+'ALL_CODES_filtered_acc.csv', index_col=None)
dfcodes_filtered = dfcodes_filtered.sort_values(['SUBJECT_ID', 'HADM_ID'])
dfcodes_filtered.to_csv(args['out_path']+'ALL_CODES_filtered.csv', index=False) #columns: 'SUBJECT_ID', 'HADM_ID', 'ICD9_CODE', 'ADMITTIME', 'DISCHTIME'
del dfnotes
del dfcodes_filtered
# step 4: link notes with their code
# labeled = self.concat_data(args['out_path']+'ALL_CODES_filtered.csv', args['out_path']+'disch_full.csv', args['out_path']+'notes_labeled.csv')
labeled = self.concat_data_new(args['out_path']+'ALL_CODES_filtered.csv', args['out_path']+'disch_full.csv', args['out_path']+'notes_labeled.csv')
#columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS'
labled_notes = pd.read_csv(labeled) #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS'
labled_notes = labled_notes.drop_duplicates()
labled_notes.to_csv(labeled, index=False) #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS'
# step 5: statistic unique word, total word, HADM_ID number
types = set()
num_tok = 0
for row in labled_notes.itertuples():
for w in row[3].split(): #TEXT in 4rd column when used itertuples
types.add(w)
num_tok += 1
print("num types", len(types), "num tokens", num_tok)
print("HADM_ID: {}".format(len(labled_notes['HADM_ID'].unique())))
print("SUBJECT_ID: {}".format(len(labled_notes['SUBJECT_ID'].unique())))
del labled_notes
#important step for entity extraction and finding their entity id from wikidata.
fname_entity = self.extract_entity('%snotes_labeled.csv' % args['out_path'], '%snotes_labeled_entity.csv' % args['out_path'])
#columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID'
#important step to create embedding file from Pytorch Biggraph pretrained embedding file for our dataset entities.
self.extract_biggraph_embedding(fname_entity, args['graph_embedding_file'], '%sentity2embedding.pickle' % args['out_path'])
# step 6: split data into train dev test
# step 7: sort data by its note length, add length to the last column
tr, dv, te = self.split_length_sort_data(fname_entity, args['out_path'], 'full') #full data split and save
#columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID', 'length'
# vocab_min = 3
vname = '%svocab.csv' % args['out_path']
self.build_vocab(args['vocab_min'], tr, vname)
# step 8: train word embeddings via word2vec and fasttext
Y = 'full'
#if want to create vocabulary from w2v model then pass the vocabulary file name where you want to save the vocabulary
w2v_file = self.word_embeddings('full', '%sdisch_full.csv' % args['out_path'], 100, 0, 5)
self.gensim_to_embeddings('%sprocessed_full.w2v' % args['out_path'], '%svocab.csv' % args['out_path'], Y)
self.fasttext_file = self.fasttext_embeddings('full', '%sdisch_full.csv' % args['out_path'], 100, 0, 5)
self.gensim_to_fasttext_embeddings('%sprocessed_full.fasttext' % args['out_path'], '%svocab.csv' % args['out_path'], Y)
# step 9: statistic the top 50 code
Y = 50
counts = Counter()
dfnl = pd.read_csv(fname_entity)
for row in dfnl.itertuples(): #for read_csv and iteratuples, the first column (row[0]) is the index column
for label in str(row[4]).split(';'): #lables are in 4th position
counts[label] += 1
codes_50 = sorted(counts.items(), key=operator.itemgetter(1), reverse=True)
codes_50 = [code[0] for code in codes_50[:Y]]
with open('%sTOP_%s_CODES.csv' % (args['out_path'], str(Y)), 'w') as of:
w = csv.writer(of)
for code in codes_50:
w.writerow([code])
with open(fname_entity, 'r') as f: #columns: 'SUBJECT_ID', 'TEXT', 'LABELS', 'ENTITY_ID'
with open('%snotes_labeled_50.csv' % args['out_path'], 'w') as fl:
r = csv.reader(f)
w = csv.writer(fl)
#header
w.writerow(next(r))
newrow = False
for row in r:
newrow = True
for code in codes_50:
if code in str(row[3]).split(';'):
if newrow:
w.writerow(row)
newrow = False
fname_50 = '%snotes_labeled_50.csv' % args['out_path'] #input dataframe
tr, dv, te = self.split_length_sort_data(fname_50, args['out_path'], str(Y))
#columns: 'SUBJECT_ID', 'TEXT', 'LABELS', 'ENTITY_ID', 'length'
def reformat(self, code, is_diag):
"""
Put a period in the right place because the MIMIC-3 data files exclude them.
Generally, procedure codes have dots after the first two digits,
while diagnosis codes have dots after the first three digits.
"""
code = ''.join(code.split('.'))
if is_diag:
if code.startswith('E'):
if len(code) > 4:
code = code[:4] + '.' + code[4:]
else:
if len(code) > 3:
code = code[:3] + '.' + code[3:]
else:
code = code[:2] + '.' + code[2:]
return code
def write_discharge_summaries(self, out_file, min_sentence_len, notes_file):
print("processing notes file")
with open(notes_file, 'r') as csvfile:
with open(out_file, 'w') as outfile:
print("writing to %s" % (out_file))
outfile.write(','.join(['SUBJECT_ID', 'HADM_ID', 'CHARTTIME', 'TEXT']) + '\n')
notereader = csv.reader(csvfile)
next(notereader)
for line in tqdm(notereader):
subj = int(float(line[1]))
category = line[6]
if category in args['notes_type'].split(','): #can Includes "Nursing" and "Physician".
note = line[10]
all_sents_inds = []
generator = nlp_tool.span_tokenize(note)
for t in generator:
all_sents_inds.append(t)
text = ""
for ind in range(len(all_sents_inds)):
start = all_sents_inds[ind][0]
end = all_sents_inds[ind][1]
sentence_txt = note[start:end]
sentence_txt = re.sub(r'[[**].+?[**]]', '', sentence_txt) #adding to remove texts between [** **]
tokens = [t.lower() for t in tokenizer.tokenize(sentence_txt) if not t.isnumeric()]
if ind == 0:
text += '[CLS] ' + ' '.join(tokens) + ' [SEP]'
else:
text += ' [CLS] ' + ' '.join(tokens) + ' [SEP]'
text = '"' + text + '"'
outfile.write(','.join([line[1], line[2], line[4], text]) + '\n')
return out_file
def code_filter(self, out_path, subj_ids):
with open(out_path+'ALL_CODES.csv', 'r') as lf:
with open(out_path+'ALL_CODES_filtered_acc.csv', 'w') as of:
w = csv.writer(of)
w.writerow(['SUBJECT_ID', 'HADM_ID', 'ICD9_CODE', 'ADMITTIME', 'DISCHTIME'])
r = csv.reader(lf)
#header
next(r)
for i,row in enumerate(r):
subj_id = int(float(row[1]))
if subj_id in subj_ids:
w.writerow(row[1:3] + [row[-1], '', ''])
def concat_data_new(self, labelsfile, notes_file, outfilename):
print("labelsfile=",labelsfile) #columns: 'SUBJECT_ID', 'HADM_ID', 'ICD9_CODE', 'ADMITTIME', 'DISCHTIME'
print("notes_file=",notes_file) #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT'
mydf_label = pd.read_csv(labelsfile)
mydf_label = mydf_label.groupby(['SUBJECT_ID','HADM_ID']).apply(lambda x: pd.Series({'ICD9_CODE':';'.join(str(v) for v in x.ICD9_CODE)})).reset_index()
mydf_notes = pd.read_csv(notes_file) #already groupby with [subj,hadm]
merged_df = pd.merge(mydf_notes, mydf_label, how='inner', on=['SUBJECT_ID','HADM_ID']).rename(columns={"ICD9_CODE": "LABELS"})
merged_df.to_csv(outfilename, index=False)
del merged_df
return outfilename
#used in old data process.
def concat_data(self, labelsfile, notes_file, outfilename):
"""
INPUTS:
labelsfile: sorted by hadm id, contains one label per line
notes_file: sorted by hadm id, contains one note per line
"""
csv.field_size_limit(sys.maxsize)
print("labelsfile=",labelsfile) #columns: 'SUBJECT_ID', 'HADM_ID', 'ICD9_CODE', 'ADMITTIME', 'DISCHTIME'
print("notes_file=",notes_file) #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT'
with open(labelsfile, 'r') as lf:
print("CONCATENATING")
with open(notes_file, 'r') as notesfile:
with open(outfilename, 'w') as outfile:
w = csv.writer(outfile)
w.writerow(['SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS'])
labels_gen = self.next_labels(lf)
notes_gen = self.next_notes(notesfile)
for i, (subj_id, text, hadm_id) in enumerate(notes_gen):
if i % 10000 == 0:
print(str(i) + " done")
cur_subj, cur_labels, cur_hadm = next(labels_gen)
if cur_hadm == hadm_id:
w.writerow([subj_id, str(hadm_id), text, ';'.join(cur_labels)])
else:
print("couldn't find matching hadm_id. data is probably not sorted correctly")
break
return outfilename
def next_labels(self, labelsfile): #columns: 'SUBJECT_ID', 'HADM_ID', 'ICD9_CODE', 'ADMITTIME', 'DISCHTIME'
"""
Generator for label sets from the label file
"""
labels_reader = csv.reader(labelsfile)
# header
next(labels_reader)
first_label_line = next(labels_reader)
cur_subj = int(first_label_line[0])
cur_hadm = int(first_label_line[1])
cur_labels = [first_label_line[2]]
for row in labels_reader:
subj_id = int(row[0])
hadm_id = int(row[1])
code = row[2]
# keep reading until you hit a new hadm id
if hadm_id != cur_hadm or subj_id != cur_subj:
yield cur_subj, cur_labels, cur_hadm
cur_labels = [code]
cur_subj = subj_id
cur_hadm = hadm_id
else:
# add to the labels and move on
cur_labels.append(code)
yield cur_subj, cur_labels, cur_hadm
def next_notes(self, notesfile): #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT'
"""
Generator for notes from the notes file
This will also concatenate discharge summaries and their addenda, which have the same subject and hadm id
"""
nr = csv.reader(notesfile)
# header
next(nr)
first_note = next(nr)
cur_subj = int(first_note[0])
cur_hadm = int(first_note[1])
cur_text = first_note[2]
for row in nr:
subj_id = int(row[0])
hadm_id = int(row[1])
text = row[2]
# keep reading until you hit a new hadm id
if hadm_id != cur_hadm or subj_id != cur_subj:
yield cur_subj, cur_text, cur_hadm
cur_text = text
cur_subj = subj_id
cur_hadm = hadm_id
else:
# concatenate to the discharge summary and move on
cur_text += " " + text
yield cur_subj, cur_text, cur_hadm
def extract_entity(self, data_file, out_file): #data file columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS'
#Pre-trained Entity extraction model from Huggingface
unmasker = pipeline('ner', model='samrawal/bert-base-uncased_clinical-ner')
#wikimapper from downloaded and indexed wikidump
mapper = WikiMapper("/home/pgoswami/DifferentialEHR/data/Wikidata_dump/index_enwiki-latest.db")
csv.field_size_limit(sys.maxsize)
with open(data_file, 'r') as lf:
with open(out_file, 'w') as of:
w = csv.writer(of)
w.writerow(['SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID'])
r = csv.reader(lf)
#header
next(r)
for i,row in enumerate(r):
if i % 1000 == 0:
print(str(i) + " entity extraction done")
text = str(row[2])
extracted_entities = ' '.join([x for x in [obj['word'] for obj in unmasker(text)[0:50]]])
fine_text = extracted_entities.replace(' ##', '').split()
entity_ids = ' '.join([mapper.title_to_id(m.title()) for m in fine_text if mapper.title_to_id(m.title()) is not None]) #getting the title ids from wikidata
w.writerow(row + [entity_ids])
return out_file
def extract_biggraph_embedding(self, data_file, embedding_file_path, out_file):
#datafile columns :'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID
selected_entity_ids = set()
with open(data_file, 'r') as lf:
r = csv.reader(lf)
#header
next(r)
for i,row in enumerate(r):
entity_ids = str(row[4]).split()
selected_entity_ids.update(entity_ids)
print(f'Total {len(selected_entity_ids)} QIDs for Entities')
entity2embedding = {}
with smart.open(embedding_file_path, encoding='utf-8') as fp: # smart open can read .gz files
for i, line in enumerate(fp):
cols = line.split('\t')
entity_id = cols[0]
if entity_id.startswith('<http://www.wikidata.org/entity/Q') and entity_id.endswith('>'):
entity_id = entity_id.replace('<http://www.wikidata.org/entity/', '').replace('>', '')
if entity_id in selected_entity_ids:
entity2embedding[entity_id] = np.array(cols[1:]).astype(np.float)
if not i % 100000:
print(f'Lines completed {i}')
# Save
with open(out_file, 'wb') as f:
pickle.dump(entity2embedding, f)
print(f'Embeddings Saved to {out_file}')
#datasetType = full/50,
#labeledfile=inputfilepath,
#base_name=outputfilename
def split_length_sort_data(self, labeledfile, base_name, datsetType):
print("SPLITTING")
labeledDf = pd.read_csv(labeledfile)
labeledDf['length'] = labeledDf.apply(lambda row: len(str(row['TEXT']).split()), axis=1)
labeledDf_train = labeledDf.sample(frac = 0.7) #70% train data
labeledDf_remain = labeledDf.drop(labeledDf_train.index)
labeledDf_dev = labeledDf_remain.sample(frac = 0.5) #15% val data
labeledDf_test = labeledDf_remain.drop(labeledDf_dev.index) #15% test data
filename_list = []
for splt in ['train', 'dev', 'test']:
filename = '%s%s_full.csv' % (base_name, splt) if datsetType == 'full' else '%s%s_%s.csv' % (base_name, splt, '50')
conv_df = eval('labeledDf_'+splt) #getting the variable
conv_df = conv_df.sort_values(['length'])
print('saving to ..'+filename)
filename_list.append(filename)
conv_df.to_csv(filename, index=False)
#gc the dataframes
del labeledDf_train
del labeledDf_remain
del labeledDf_dev
del labeledDf_test
return filename_list[0], filename_list[1], filename_list[2]
def build_vocab(self, vocab_min, infile, vocab_filename): #columns : 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID', 'length'
"""
INPUTS:
vocab_min: how many documents a word must appear in to be kept
infile: (training) data file to build vocabulary from. CSV reader also need huge memory to load the file.
vocab_filename: name for the file to output
"""
csv.field_size_limit(sys.maxsize)
with open(infile, 'r') as csvfile: #columns: 'SUBJECT_ID', 'HADM_ID', 'TEXT', 'LABELS', 'ENTITY_ID', 'length'
reader = csv.reader(csvfile)
# header
next(reader)
# 0. read in data
print("reading in data...")
# holds number of terms in each document
note_numwords = []
# indices where notes start
note_inds = [0]
# indices of discovered words
indices = []
# holds a bunch of ones
data = []
# keep track of discovered words
vocab = {}
# build lookup table for terms
num2term = {}
# preallocate array to hold number of notes each term appears in
note_occur = np.zeros(400000, dtype=int)
i = 0
for row in reader:
text = row[2] #chnage Prantik: after merging same subject values, text is in third (2) position
numwords = 0
for term in text.split():
# put term in vocab if it's not there. else, get the index
index = vocab.setdefault(term, len(vocab))
indices.append(index)
num2term[index] = term
data.append(1)
numwords += 1
# record where the next note starts
note_inds.append(len(indices))
indset = set(indices[note_inds[-2]:note_inds[-1]])
# go thru all the word indices you just added, and add to the note occurrence count for each of them
for ind in indset:
note_occur[ind] += 1
note_numwords.append(numwords)
i += 1
# clip trailing zeros
note_occur = note_occur[note_occur > 0]
# turn vocab into a list so indexing doesn't get fd up when we drop rows
vocab_list = np.array([word for word, ind in sorted(vocab.items(), key=operator.itemgetter(1))])
# 1. create sparse document matrix
C = csr_matrix((data, indices, note_inds), dtype=int).transpose()
# also need the numwords array to be a sparse matrix
note_numwords = csr_matrix(1. / np.array(note_numwords))
# 2. remove rows with less than 3 total occurrences
print("removing rare terms")
# inds holds indices of rows corresponding to terms that occur in < 3 documents
inds = np.nonzero(note_occur >= vocab_min)[0]
print(str(len(inds)) + " terms qualify out of " + str(C.shape[0]) + " total")
# drop those rows
C = C[inds, :]
note_occur = note_occur[inds]
vocab_list = vocab_list[inds]
print("writing output")
with open(vocab_filename, 'w') as vocab_file:
for word in vocab_list:
vocab_file.write(word + "\n")
def word_embeddings(self, Y, notes_file, embedding_size, min_count, n_iter, outfile=None):
modelname = "processed_%s.w2v" % (Y)
sentences = ProcessedIter(Y, notes_file)
print("Model name %s..." % (modelname))
model = w2v.Word2Vec(vector_size=embedding_size, min_count=min_count, workers=4, epochs=n_iter)
print("building word2vec vocab on %s..." % (notes_file))
model.build_vocab(sentences)
print("training...")
model.train(sentences, total_examples=model.corpus_count, epochs=model.epochs)
out_file = '/'.join(notes_file.split('/')[:-1] + [modelname])
print("writing embeddings to %s" % (out_file))
model.save(out_file)
#if want to create vocab from w2v model, pass the vocab file name
if outfile is not None:
print("writing vocab to %s" % (outfile))
with open(outfile, 'w') as vocab_file:
for word in model.wv.key_to_index:
vocab_file.write(word + "\n")
return out_file
def gensim_to_embeddings(self, wv_file, vocab_file, Y, outfile=None):
model = gensim.models.Word2Vec.load(wv_file)
wv = model.wv
#free up memory
del model
vocab = set()
with open(vocab_file, 'r') as vocabfile:
for i,line in enumerate(vocabfile):
line = line.strip()
if line != '':
vocab.add(line)
ind2w = {i+1:w for i,w in enumerate(sorted(vocab))}
W, words = self.build_matrix(ind2w, wv)
if outfile is None:
outfile = wv_file.replace('.w2v', '.embed')
#smash that save button
self.save_embeddings(W, words, outfile)
def build_matrix(self, ind2w, wv):
"""
Go through vocab in order. Find vocab word in wv.index2word, then call wv.word_vec(wv.index2word[i]).
Put results into one big matrix.
Note: ind2w starts at 1 (saving 0 for the pad character), but gensim word vectors starts at 0
"""
W = np.zeros((len(ind2w)+1, len(wv.get_vector(wv.index_to_key[0])) ))
print("W shape=",W.shape)
words = ["**PAD**"]
W[0][:] = np.zeros(len(wv.get_vector(wv.index_to_key[0])))
for idx, word in tqdm(ind2w.items()):
if idx >= W.shape[0]:
break
W[idx][:] = wv.get_vector(word)
words.append(word)
print("W shape final=",W.shape)
print("Word list length=",len(words))
return W, words
def fasttext_embeddings(self, Y, notes_file, embedding_size, min_count, n_iter):
modelname = "processed_%s.fasttext" % (Y)
sentences = ProcessedIter(Y, notes_file)
print("Model name %s..." % (modelname))
model = fasttext.FastText(vector_size=embedding_size, min_count=min_count, epochs=n_iter)
print("building fasttext vocab on %s..." % (notes_file))
model.build_vocab(sentences)
print("training...")
model.train(sentences, total_examples=model.corpus_count, epochs=model.epochs)
out_file = '/'.join(notes_file.split('/')[:-1] + [modelname])
print("writing embeddings to %s" % (out_file))
model.save(out_file)
return out_file
def gensim_to_fasttext_embeddings(self, wv_file, vocab_file, Y, outfile=None):
model = gensim.models.FastText.load(wv_file)
wv = model.wv
#free up memory
del model
vocab = set()
with open(vocab_file, 'r') as vocabfile:
for i,line in enumerate(vocabfile):
line = line.strip()
if line != '':
vocab.add(line)
ind2w = {i+1:w for i,w in enumerate(sorted(vocab))}
W, words = self.build_matrix(ind2w, wv)
if outfile is None:
outfile = wv_file.replace('.fasttext', '.fasttext.embed')
#smash that save button
self.save_embeddings(W, words, outfile)
def save_embeddings(self, W, words, outfile):
with open(outfile, 'w') as o:
#pad token already included
for i in range(len(words)):
line = [words[i]]
line.extend([str(d) for d in W[i]])
o.write(" ".join(line) + "\n")
# -
# ## Approach
# ## Everything in one cell for easy running.
# + id="_9ckqsQhqWyE"
#############Imports###############
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.nn.init import xavier_uniform_ as xavier_uniform
import torch.optim as optim
from torch.utils.data import DataLoader
from torch.optim.lr_scheduler import StepLR, ReduceLROnPlateau
from torch.utils.data import Dataset
from torchsummary import summary
from entmax import sparsemax, entmax15, entmax_bisect
from pytorch_pretrained_bert.modeling import BertLayerNorm
from pytorch_pretrained_bert import BertModel, BertConfig
from pytorch_pretrained_bert import WEIGHTS_NAME, CONFIG_NAME
from pytorch_pretrained_bert import BertAdam
from pytorch_pretrained_bert import BertTokenizer
import transformers as tr
from transformers import AdamW
from allennlp.data.token_indexers.elmo_indexer import ELMoTokenCharactersIndexer
from allennlp.data import Token, Vocabulary, Instance
from allennlp.data.fields import TextField
from allennlp.data.dataset import Batch
from sklearn.metrics import f1_score, precision_recall_fscore_support, accuracy_score
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import roc_curve, auc
from keras.preprocessing.sequence import pad_sequences
from keras.preprocessing.text import Tokenizer
from typing import Tuple,Callable,IO,Optional
from collections import defaultdict
from urllib.parse import urlparse
from functools import wraps
from hashlib import sha256
from typing import List
from math import floor
from tqdm import tqdm
import pandas as pd
import numpy as np
import requests
import tempfile
import tarfile
import random
import shutil
import struct
import pickle
import time
import json
import csv
import sys
import os
#Models
class ModelHub:
def __init__(self, args, dicts):
self.pick_model(args, dicts)
def pick_model(self, args, dicts):
Y = len(dicts['ind2c'])
if args['model'] == 'KG_MultiResCNN':
model = KG_MultiResCNN(args, Y, dicts)
elif args['model'] == 'KG_MultiResCNNLSTM':
model = KG_MultiResCNNLSTM(args, Y, dicts)
elif args['model'] == 'bert_se_kg':
model = Bert_SE_KG(args, Y, dicts)
elif args['model'] == 'bert_we_kg':
model = Bert_WE_KG(args, Y, dicts)
elif args['model'] == 'bert_l4_we_kg':
model = Bert_L4_WE_KG(args, Y, dicts)
elif args['model'] == 'bert_mcnn_kg':
model = Bert_MCNN_KG(args, Y, dicts)
else:
raise RuntimeError("wrong model name")
if args['test_model']:
sd = torch.load(args['test_model'])
model.load_state_dict(sd)
if args['gpu'] >= 0:
model.cuda(args['gpu'])
return model
class WordRep(nn.Module):
def __init__(self, args, Y, dicts):
super(WordRep, self).__init__()
self.gpu = args['gpu']
self.isTfIdf = False
if args['use_tfIdf']:
self.isTfIdf = True
if args['embed_file']:
print("loading pretrained embeddings from {}".format(args['embed_file']))
if args['use_ext_emb']:
pretrain_word_embedding, pretrain_emb_dim = self.build_pretrain_embedding(args['embed_file'], dicts['w2ind'],
True)
W = torch.from_numpy(pretrain_word_embedding)
else:
W = torch.Tensor(self.load_embeddings(args['embed_file']))
self.embed = nn.Embedding(W.size()[0], W.size()[1], padding_idx=0)
self.embed.weight.data = W.clone()
else:
# add 2 to include UNK and PAD
self.embed = nn.Embedding(len(dicts['w2ind']) + 2, args['embed_size'], padding_idx=0)
self.feature_size = self.embed.embedding_dim
self.embed_drop = nn.Dropout(p=args['dropout'])
self.conv_dict = {
1: [self.feature_size, args['num_filter_maps']],
2: [self.feature_size, 100, args['num_filter_maps']],
3: [self.feature_size, 150, 100, args['num_filter_maps']],
4: [self.feature_size, 200, 150, 100, args['num_filter_maps']]
}
def forward(self, x, tfIdf_inputs): #tfIdf_inputs
if self.gpu >= 0:
x = x if x.is_cuda else x.cuda(self.gpu)
if self.isTfIdf and tfIdf_inputs != None:
tfIdf_inputs = tfIdf_inputs if tfIdf_inputs.is_cuda else tfIdf_inputs.cuda(self.gpu)
try:
features = [self.embed(x)]
except:
print(x)
raise
out = torch.cat(features, dim=2)
if self.isTfIdf and tfIdf_inputs != None:
weight = tfIdf_inputs.unsqueeze(dim=2)
out = out * weight
out = self.embed_drop(out)
del x
del tfIdf_inputs
return out
def load_embeddings(self, embed_file):
#also normalizes the embeddings
W = []
with open(embed_file) as ef:
for line in ef:
line = line.rstrip().split()
vec = np.array(line[1:]).astype(np.float)
vec = vec / float(np.linalg.norm(vec) + 1e-6)
W.append(vec)
#UNK embedding, gaussian randomly initialized
print("adding unk embedding")
vec = np.random.randn(len(W[-1]))
vec = vec / float(np.linalg.norm(vec) + 1e-6)
W.append(vec)
W = np.array(W)
return W
def build_pretrain_embedding(self, embedding_path, word_alphabet, norm):
embedd_dict, embedd_dim = self.load_pretrain_emb(embedding_path)
scale = np.sqrt(3.0 / embedd_dim)
pretrain_emb = np.zeros([len(word_alphabet)+2, embedd_dim], dtype=np.float32) # add UNK (last) and PAD (0)
perfect_match = 0
case_match = 0
digits_replaced_with_zeros_found = 0
lowercase_and_digits_replaced_with_zeros_found = 0
not_match = 0
for word, index in word_alphabet.items():
if word in embedd_dict:
if norm:
pretrain_emb[index,:] = self.norm2one(embedd_dict[word])
else:
pretrain_emb[index,:] = embedd_dict[word]
perfect_match += 1
elif word.lower() in embedd_dict:
if norm:
pretrain_emb[index,:] = self.norm2one(embedd_dict[word.lower()])
else:
pretrain_emb[index,:] = embedd_dict[word.lower()]
case_match += 1
elif re.sub('\d', '0', word) in embedd_dict:
if norm:
pretrain_emb[index,:] = self.norm2one(embedd_dict[re.sub('\d', '0', word)])
else:
pretrain_emb[index,:] = embedd_dict[re.sub('\d', '0', word)]
digits_replaced_with_zeros_found += 1
elif re.sub('\d', '0', word.lower()) in embedd_dict:
if norm:
pretrain_emb[index,:] = self.norm2one(embedd_dict[re.sub('\d', '0', word.lower())])
else:
pretrain_emb[index,:] = embedd_dict[re.sub('\d', '0', word.lower())]
lowercase_and_digits_replaced_with_zeros_found += 1
else:
if norm:
pretrain_emb[index, :] = self.norm2one(np.random.uniform(-scale, scale, [1, embedd_dim]))
else:
pretrain_emb[index,:] = np.random.uniform(-scale, scale, [1, embedd_dim])
not_match += 1
# initialize pad and unknown
pretrain_emb[0, :] = np.zeros([1, embedd_dim], dtype=np.float32)
if norm:
pretrain_emb[-1, :] = self.norm2one(np.random.uniform(-scale, scale, [1, embedd_dim]))
else:
pretrain_emb[-1, :] = np.random.uniform(-scale, scale, [1, embedd_dim])
print("pretrained word emb size {}".format(len(embedd_dict)))
print("prefect match:%.2f%%, case_match:%.2f%%, dig_zero_match:%.2f%%, "
"case_dig_zero_match:%.2f%%, not_match:%.2f%%"
%(perfect_match*100.0/len(word_alphabet), case_match*100.0/len(word_alphabet), digits_replaced_with_zeros_found*100.0/len(word_alphabet),
lowercase_and_digits_replaced_with_zeros_found*100.0/len(word_alphabet), not_match*100.0/len(word_alphabet)))
return pretrain_emb, embedd_dim
def load_pretrain_emb(self, embedding_path):
embedd_dim = -1
embedd_dict = dict()
# emb_debug = []
if embedding_path.find('.bin') != -1:
with open(embedding_path, 'rb') as f:
wordTotal = int(self._readString(f, 'utf-8'))
embedd_dim = int(self._readString(f, 'utf-8'))
for i in range(wordTotal):
word = self._readString(f, 'utf-8')
# emb_debug.append(word)
word_vector = []
for j in range(embedd_dim):
word_vector.append(self._readFloat(f))
word_vector = np.array(word_vector, np.float)
f.read(1) # a line break
embedd_dict[word] = word_vector
else:
with codecs.open(embedding_path, 'r', 'UTF-8') as file:
for line in file:
# logging.info(line)
line = line.strip()
if len(line) == 0:
continue
# tokens = line.split()
tokens = re.split(r"\s+", line)
if len(tokens) == 2:
continue # it's a head
if embedd_dim < 0:
embedd_dim = len(tokens) - 1
else:
# assert (embedd_dim + 1 == len(tokens))
if embedd_dim + 1 != len(tokens):
continue
embedd = np.zeros([1, embedd_dim])
embedd[:] = tokens[1:]
embedd_dict[tokens[0]] = embedd
return embedd_dict, embedd_dim
def _readString(self, f, code):
# s = unicode()
s = str()
c = f.read(1)
value = ord(c)
while value != 10 and value != 32:
if 0x00 < value < 0xbf:
continue_to_read = 0
elif 0xC0 < value < 0xDF:
continue_to_read = 1
elif 0xE0 < value < 0xEF:
continue_to_read = 2
elif 0xF0 < value < 0xF4:
continue_to_read = 3
else:
raise RuntimeError("not valid utf-8 code")
i = 0
# temp = str()
# temp = temp + c
temp = bytes()
temp = temp + c
while i<continue_to_read:
temp = temp + f.read(1)
i += 1
temp = temp.decode(code)
s = s + temp
c = f.read(1)
value = ord(c)
return s
def _readFloat(self,f):
bytes4 = f.read(4)
f_num = struct.unpack('f', bytes4)[0]
return f_num
def norm2one(self,vec):
root_sum_square = np.sqrt(np.sum(np.square(vec)))
return vec/root_sum_square
class SentimentOutput():
def __init__(self, args):
self.gpu = args['gpu']
cache_path = os.path.join(args['bert_dir'], args['sentiment_bert'])
savedModel = None
if os.path.exists(cache_path):
print("model path exist")
savedModel = tr.AutoModelForSequenceClassification.from_pretrained(cache_path)
else:
print("Downloading and saving model")
savedModel = tr.AutoModelForSequenceClassification.from_pretrained(str(args['sentiment_bert']))
savedModel.save_pretrained(save_directory = cache_path, save_config=True)
self.bert = savedModel
self.config = savedModel.config
def forward(self, x):
encoded_input = dict()
if self.gpu >= 0:
x[0] = x[0] if x[0].is_cuda else x[0].cuda(self.gpu)
x[1] = x[1] if x[1].is_cuda else x[1].cuda(self.gpu)
model = self.bert
model = model.cuda(self.gpu)
encoded_input['input_ids'] = x[0]
encoded_input['attention_mask'] = x[1]
senti_output = model(**encoded_input, output_hidden_states=True)
all_hidden_states = senti_output.hidden_states
out = all_hidden_states[-1] #last hidden state. [#batch_size, sequence(m), 1024]
del all_hidden_states
del senti_output
del encoded_input
del x
del model
return out
class OutputLayer(nn.Module):
def __init__(self, args, Y, dicts, input_size):
super(OutputLayer, self).__init__()
self.gpu = args['gpu']
self.use_entmax15 = False
if args['use_entmax15']:
self.use_entmax15 = True
self.U = nn.Linear(input_size, Y)
xavier_uniform(self.U.weight)
self.final = nn.Linear(input_size, Y)
xavier_uniform(self.final.weight)
self.loss_function = nn.BCEWithLogitsLoss()
def forward(self, x, target):
if self.gpu >= 0:
target = target if target.is_cuda else target.cuda(self.gpu)
x = x if x.is_cuda else x.cuda(self.gpu)
if self.use_entmax15:
alpha = entmax15(self.U.weight.matmul(x.transpose(1, 2)), dim=2)
else:
alpha = F.softmax(self.U.weight.matmul(x.transpose(1, 2)), dim=2)
m = alpha.matmul(x)
y = self.final.weight.mul(m).sum(dim=2).add(self.final.bias)
loss = self.loss_function(y, target)
del x
del target
return y, loss
class MRCNNLayer(nn.Module):
def __init__(self, args, feature_size):
super(MRCNNLayer, self).__init__()
self.gpu = args['gpu']
self.feature_size = feature_size
self.conv_dict = {
1: [self.feature_size, args['num_filter_maps']],
2: [self.feature_size, 100, args['num_filter_maps']],
3: [self.feature_size, 150, 100, args['num_filter_maps']],
4: [self.feature_size, 200, 150, 100, args['num_filter_maps']]
}
self.conv = nn.ModuleList()
filter_sizes = args['filter_size'].split(',')
self.filter_num = len(filter_sizes)
for filter_size in filter_sizes:
filter_size = int(filter_size)
one_channel = nn.ModuleList()
tmp = nn.Conv1d(self.feature_size, self.feature_size, kernel_size=filter_size,
padding=int(floor(filter_size / 2)))
xavier_uniform(tmp.weight)
one_channel.add_module('baseconv', tmp)
conv_dimension = self.conv_dict[args['conv_layer']]
for idx in range(args['conv_layer']):
tmp = ResidualBlock(conv_dimension[idx], conv_dimension[idx + 1], filter_size, 1, True,
args['dropout'])
one_channel.add_module('resconv-{}'.format(idx), tmp)
self.conv.add_module('channel-{}'.format(filter_size), one_channel)
def forward(self, x):
if self.gpu >= 0:
x = x if x.is_cuda else x.cuda(self.gpu)
x = x.transpose(1, 2)
conv_result = []
for conv in self.conv:
tmp = x
for idx, md in enumerate(conv):
if idx == 0:
tmp = torch.tanh(md(tmp))
else:
tmp = md(tmp)
tmp = tmp.transpose(1, 2)
conv_result.append(tmp)
out = torch.cat(conv_result, dim=2)
del x
return out
class ResidualBlock(nn.Module):
def __init__(self, inchannel, outchannel, kernel_size, stride, use_res, dropout):
super(ResidualBlock, self).__init__()
self.left = nn.Sequential(
nn.Conv1d(inchannel, outchannel, kernel_size=kernel_size, stride=stride, padding=int(floor(kernel_size / 2)), bias=False),
nn.BatchNorm1d(outchannel),
nn.Tanh(),
nn.Conv1d(outchannel, outchannel, kernel_size=kernel_size, stride=1, padding=int(floor(kernel_size / 2)), bias=False),
nn.BatchNorm1d(outchannel)
)
self.use_res = use_res
if self.use_res:
self.shortcut = nn.Sequential(
nn.Conv1d(inchannel, outchannel, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm1d(outchannel)
)
self.dropout = nn.Dropout(p=dropout)
def forward(self, x):
out = self.left(x)
if self.use_res:
out += self.shortcut(x)
out = torch.tanh(out)
out = self.dropout(out)
return out
class KG_MultiResCNN(nn.Module):
def __init__(self, args, Y, dicts):
super(KG_MultiResCNN, self).__init__()
self.word_rep = WordRep(args, Y, dicts)
self.feature_size = self.word_rep.feature_size
self.is_sentiment = False
if args['use_sentiment']:
self.sentiment_model = SentimentOutput(args)
self.S_U = nn.Linear(self.sentiment_model.config.hidden_size, self.feature_size)
self.is_sentiment = True
if args['use_embd_layer'] and args['add_with_wordrap']:
self.kg_embd = EntityEmbedding(args, Y)
self.kg_embd.dim_red = nn.Linear(self.kg_embd.feature_size, self.feature_size)
self.kg_embd.feature_red = nn.Linear(args['MAX_ENT_LENGTH'], args['MAX_LENGTH'])
self.add_emb_with_wordrap = True
self.dropout = nn.Dropout(args['dropout'])
self.conv = MRCNNLayer(args, self.feature_size)
self.feature_size = self.conv.filter_num * args['num_filter_maps']
self.output_layer = OutputLayer(args, Y, dicts, self.feature_size)
def forward(self, x, target, text_inputs, embeddings, tfIdf_inputs): #inputs_id, labels, text_inputs, embeddings, tfIdf_inputs
x = self.word_rep(x, tfIdf_inputs) #(batch, sequence, 100)
if self.is_sentiment:
senti_out = self.sentiment_model.forward(text_inputs)
s_alpha = self.S_U(senti_out)
del senti_out
x = torch.mul(x, s_alpha) #(batch, sequence, 100)
del s_alpha
if hasattr(self, 'add_emb_with_wordrap') and (self.add_emb_with_wordrap):
# with embedding layer
out = self.kg_embd(embeddings) #torch.Size([batch, seq len(n), embedding dim(200)])
out = self.kg_embd.dim_red(out) #torch.Size([batch, seq len(n), embedding dim(100)])
x = torch.cat((x, out), dim=1) # new shape (batch_size, sequence_length(m+n), feature_size (100))
del out
x = self.dropout(x)
x = self.conv(x)
y, loss = self.output_layer(x, target)
del x
return y, loss
def freeze_net(self):
for p in self.word_rep.embed.parameters():
p.requires_grad = False
class KG_MultiResCNNLSTM(nn.Module):
def __init__(self, args, Y, dicts):
super(KG_MultiResCNNLSTM, self).__init__()
self.word_rep = WordRep(args, Y, dicts)
self.embedding_size = self.word_rep.embed.weight.data.size()[0]
self.conv = nn.ModuleList()
filter_sizes = args['filter_size'].split(',')
self.filter_num = len(filter_sizes)
for filter_size in filter_sizes:
filter_size = int(filter_size)
one_channel = nn.ModuleList()
tmp = nn.Conv1d(self.word_rep.feature_size, self.word_rep.feature_size, kernel_size=filter_size,
padding=int(floor(filter_size / 2)))
xavier_uniform(tmp.weight)
one_channel.add_module('baseconv', tmp)
conv_dimension = self.word_rep.conv_dict[args['conv_layer']]
for idx in range(args['conv_layer']):
tmp = ResidualBlock(conv_dimension[idx], conv_dimension[idx + 1], filter_size, 1, True,
args['dropout'])
one_channel.add_module('resconv-{}'.format(idx), tmp)
lstm = torch.nn.LSTM(
input_size= args['num_filter_maps'],
hidden_size= args['num_filter_maps'],
num_layers=1
)
one_channel.add_module('LSTM', lstm)
self.conv.add_module('channel-{}'.format(filter_size), one_channel)
self.output_layer = OutputLayer(args, Y, dicts, self.filter_num * args['num_filter_maps'])
def forward(self, x, target, text_inputs, embeddings, tfIdf_inputs):
x = self.word_rep(x, tfIdf_inputs)
x = x.transpose(1, 2)
conv_result = []
for conv in self.conv:
tmp = x
for idx, md in enumerate(conv):
if idx == 0:
tmp = torch.tanh(md(tmp))
else:
if idx == 2:
tmp = tmp.transpose(1, 2)
tmp, (h,c) = md(tmp)
tmp = tmp.transpose(1, 2)
else:
tmp = md(tmp)
tmp = tmp.transpose(1, 2)
conv_result.append(tmp)
x = torch.cat(conv_result, dim=2)
y, loss = self.output_layer(x, target)
return y, loss
def freeze_net(self):
for p in self.word_rep.embed.parameters():
p.requires_grad = False
class KGEntityToVec:
@staticmethod
def getEntityToVec():
with open('%sentity2embedding.pickle' % args['out_path'], 'rb') as f:
entity2vec = pickle.load(f)
return entity2vec
class EntityEmbedding(nn.Module):
def __init__(self, args, Y):
super(EntityEmbedding, self).__init__()
self.gpu = args['gpu']
entity2vec = KGEntityToVec().getEntityToVec()
embedding_matrix = self.create_embedding_matrix(entity2vec)
vocab_size=embedding_matrix.shape[0]
vector_size=embedding_matrix.shape[1]
self.embed = nn.Embedding(num_embeddings=vocab_size,embedding_dim=vector_size)
self.embed.weight=nn.Parameter(torch.tensor(embedding_matrix,dtype=torch.float32))
# self.embed.weight.requires_grad=False
self.feature_size = self.embed.embedding_dim
self.conv_dict = {
1: [self.feature_size, args['num_filter_maps']],
2: [self.feature_size, 100, args['num_filter_maps']],
3: [self.feature_size, 150, 100, args['num_filter_maps']],
4: [self.feature_size, 200, 150, 100, args['num_filter_maps']]
}
self.embed_drop = nn.Dropout(p=args['dropout'])
def forward(self, x):
if self.gpu >= 0:
x = x if x.is_cuda else x.cuda(self.gpu)
features = [self.embed(x)]
output = torch.cat(features, dim=2)
output = self.embed_drop(output)
del x
return output
def create_embedding_matrix(self, ent2vec):
embedding_matrix=np.zeros((len(ent2vec)+2,200))
for index, key in enumerate(ent2vec):
vec = ent2vec[key]
vec = vec / float(np.linalg.norm(vec) + 1e-6)
embedding_matrix[index+1]=vec
return embedding_matrix
class Bert_SE_KG(nn.Module): #bert with sentence embedding
def __init__(self, args, Y, dicts):
super(Bert_SE_KG, self).__init__()
cache_path = os.path.join(args['bert_dir'], args['pretrained_bert'])
savedModel = None
if os.path.exists(cache_path):
print("model path exist")
savedModel = tr.BertModel.from_pretrained(cache_path)
else:
print("Downloading and saving model")
savedModel = tr.BertModel.from_pretrained(str(args['pretrained_bert']))
savedModel.save_pretrained(save_directory = cache_path, save_config=True)
self.bert = savedModel
self.config = savedModel.config
# print("Model config {}".format(self.config))
self.feature_size = self.config.hidden_size
if args['use_embd_layer']:
self.kg_embd = EntityEmbedding(args, Y)
self.kg_embd.embed.weight.requires_grad=False
filetrs = [3]
self.convs = nn.ModuleList([nn.Conv1d(self.kg_embd.feature_size, self.kg_embd.feature_size, int(filter_size)) for filter_size in filetrs])
self.dim_reduction = nn.Linear(self.feature_size, self.kg_embd.feature_size)
self.feature_size = self.kg_embd.feature_size*2
self.dropout = nn.Dropout(args['dropout'])
self.classifier = nn.Linear(self.feature_size, Y)
self.apply(self.init_bert_weights)
def forward(self, input_ids, token_type_ids, attention_mask, entity_embeddings, target):
last_hidden_state, x = self.bert(input_ids=input_ids, token_type_ids=token_type_ids, attention_mask=attention_mask, return_dict=False)
if hasattr(self, 'kg_embd'):
# with embedding layer
out = self.kg_embd(entity_embeddings) #torch.Size([batch, seq len(n), embedding dim(200)])
embedded = out.permute(0,2,1) #torch.Size([batch, embedding dim (200), seq len])#if want sentence embedding
conved = [torch.relu(conv(embedded)) for conv in self.convs] #if want sentence embedding
pooled = [conv.max(dim=-1).values for conv in conved] #if want sentence embedding
cat = self.dropout(torch.cat(pooled, dim=-1)) #if want sentence embedding
x = self.dim_reduction(x)
x = x / float(torch.norm(x) + 1e-6)
x = torch.cat((x, cat), dim=1) #if want sentence embedding
x = self.dropout(x) #(batch_size, sequence_length(m), hidden_size(200/756))
y = self.classifier(x)
loss = F.binary_cross_entropy_with_logits(y, target)
return y, loss
def init_bert_weights(self, module):
BertLayerNorm = torch.nn.LayerNorm
if isinstance(module, (nn.Linear, nn.Embedding)):
module.weight.data.normal_(mean=0.0, std=self.config.initializer_range)
elif isinstance(module, BertLayerNorm):
module.bias.data.zero_()
module.weight.data.fill_(1.0)
if isinstance(module, nn.Linear) and module.bias is not None:
module.bias.data.zero_()
def freeze_net(self):
pass
class Bert_WE_KG(nn.Module): #bert with word embedding
def __init__(self, args, Y, dicts):
super(Bert_WE_KG, self).__init__()
cache_path = os.path.join(args['bert_dir'], args['pretrained_bert'])
savedModel = None
if os.path.exists(cache_path):
savedModel = tr.BertModel.from_pretrained(cache_path)
else:
savedModel = tr.BertModel.from_pretrained(str(args['pretrained_bert']))
savedModel.save_pretrained(save_directory = cache_path, save_config=True)
self.bert = savedModel
self.config = savedModel.config
self.feature_size = self.config.hidden_size
if args['use_embd_layer']:
self.kg_embd = EntityEmbedding(args, Y)
self.kg_embd.embed.weight.requires_grad=False
self.dim_reduction = nn.Linear(self.feature_size, self.kg_embd.feature_size)
self.feature_size = self.kg_embd.feature_size
self.dropout = nn.Dropout(args['dropout'])
self.output_layer = OutputLayer(args, Y, dicts, self.feature_size)
self.apply(self.init_bert_weights)
def forward(self, input_ids, token_type_ids, attention_mask, entity_embeddings, target):
last_hidden_state, pooled_output = self.bert(input_ids=input_ids, token_type_ids=token_type_ids, attention_mask=attention_mask, return_dict=False)
x = self.dropout(last_hidden_state) #(batch_size, sequence_length(m), hidden_size(786))
if hasattr(self, 'kg_embd'):
out = self.kg_embd(entity_embeddings) #torch.Size([batch, seq len(n), embedding dim(200)])
x = self.dim_reduction(x) #torch.Size([batch, seq len(m), embedding dim(200)])
x = x / float(torch.norm(x) + 1e-6)
x = torch.cat((x, out), dim=1) # new shape (batch_size, sequence_length(m+n), feature_size (200))
y, loss = self.output_layer(x, target)
return y, loss
def init_bert_weights(self, module):
BertLayerNorm = torch.nn.LayerNorm
if isinstance(module, (nn.Linear, nn.Embedding)):
module.weight.data.normal_(mean=0.0, std=self.config.initializer_range)
elif isinstance(module, BertLayerNorm):
module.bias.data.zero_()
module.weight.data.fill_(1.0)
if isinstance(module, nn.Linear) and module.bias is not None:
module.bias.data.zero_()
def freeze_net(self):
pass
class Bert_L4_WE_KG(nn.Module): #adding last 5 layers output of bert
def __init__(self, args, Y, dicts):
super(Bert_L4_WE_KG, self).__init__()
cache_path = os.path.join(args['bert_dir'], args['pretrained_bert'])
savedModel = None
if os.path.exists(cache_path):
savedModel = tr.BertModel.from_pretrained(cache_path, return_dict=True)
else:
savedModel = tr.BertModel.from_pretrained(str(args['pretrained_bert']), return_dict=True)
savedModel.save_pretrained(save_directory = cache_path, save_config=True)
self.bert = savedModel
self.config = savedModel.config
self.feature_size = self.config.hidden_size*4
if args['use_embd_layer']:
self.kg_embd = EntityEmbedding(args, Y)
self.kg_embd.embed.weight.requires_grad=False
self.dim_reduction = nn.Linear(self.feature_size, self.kg_embd.feature_size)
self.feature_size = self.kg_embd.feature_size
self.dropout = nn.Dropout(args['dropout'])
self.output_layer = OutputLayer(args, Y, dicts, self.feature_size)
self.apply(self.init_bert_weights)
def forward(self, input_ids, token_type_ids, attention_mask, entity_embeddings, target):
output = self.bert(input_ids=input_ids, token_type_ids=token_type_ids, attention_mask=attention_mask, output_hidden_states=True)
#*************experiment*************
hidden_states = output.hidden_states
# concatenate last four layers
x = torch.cat([hidden_states[i] for i in [-1,-2,-3,-4]], dim=-1) #[batch_size, sequence_length, hidden_size(786)*4]
#***********experiment***************
x = self.dropout(x)
if hasattr(self, 'kg_embd'):
out = self.kg_embd(entity_embeddings) #torch.Size([batch, seq len(n), embedding dim(200)])
x = self.dim_reduction(x) #torch.Size([batch, seq len(m), embedding dim(200)])
x = x / float(torch.norm(x) + 1e-6)
x = torch.cat((x, out), dim=1) # new shape (batch_size, sequence_length(m+n), feature_size (200))
x = self.dropout(x)
y, loss = self.output_layer(x, target)
return y, loss
def loss_fn(self, outputs, target):
return nn.BCEWithLogitsLoss()(outputs, target)
def init_bert_weights(self, module):
BertLayerNorm = torch.nn.LayerNorm
if isinstance(module, (nn.Linear, nn.Embedding)):
module.weight.data.normal_(mean=0.0, std=self.config.initializer_range)
elif isinstance(module, BertLayerNorm):
module.bias.data.zero_()
module.weight.data.fill_(1.0)
if isinstance(module, nn.Linear) and module.bias is not None:
module.bias.data.zero_()
def freeze_net(self):
pass
class Bert_MCNN_KG(nn.Module): #Bert with KG and CNN
def __init__(self, args, Y, dicts):
super(Bert_MCNN_KG, self).__init__()
cache_path = os.path.join(args['bert_dir'], args['pretrained_bert'])
savedModel = None
if os.path.exists(cache_path):
savedModel = tr.BertModel.from_pretrained(cache_path)
else:
savedModel = tr.BertModel.from_pretrained(str(args['pretrained_bert']))
savedModel.save_pretrained(save_directory = cache_path, save_config=True)
self.bert = savedModel
self.config = savedModel.config
self.dim_reduction1 = nn.Linear(self.config.hidden_size*4, self.config.hidden_size)
self.feature_size = self.config.hidden_size
if args['use_embd_layer']:
self.kg_embd = EntityEmbedding(args, Y)
self.kg_embd.embed.weight.requires_grad=False
self.dim_reduction2 = nn.Linear(self.feature_size, self.kg_embd.feature_size)
self.feature_size = self.kg_embd.feature_size
self.dropout = nn.Dropout(args['dropout'])
self.conv = MRCNNLayer(args, self.feature_size)
self.feature_size = self.conv.filter_num * args['num_filter_maps']
self.output_layer = OutputLayer(args, Y, dicts, self.feature_size)
# self.apply(self.init_bert_weights)
def forward(self, input_ids, token_type_ids, attention_mask, entity_embeddings, target):
output = self.bert(input_ids=input_ids, token_type_ids=token_type_ids, attention_mask=attention_mask, output_hidden_states=True)
#*************experiment*************
hidden_states = output.hidden_states
# concatenate last four layers
x = torch.cat([hidden_states[i] for i in [-1,-2,-3,-4]], dim=-1) #[batch_size, sequence_length, hidden_size(786)*4]
#***********experiment***************
x = x / float(torch.norm(x) + 1e-6) #normalize
x = self.dim_reduction1(x) #[batch_size, sequence_length, hidden_size(786]
if hasattr(self, 'kg_embd'):
out = self.kg_embd(entity_embeddings) #torch.Size([batch, seq len(n), embedding dim(200)])
x = self.dim_reduction2(x)
x = torch.cat((x, out), dim=1) # new shape (batch_size, sequence_length(m+n), feature_size (200))
x = self.dropout(x) #(batch_size, sequence_length, hidden_size(786 or 200))
x = self.conv(x)
y, loss = self.output_layer(x, target)
return y, loss
def freeze_net(self):
pass
#############Train-Test###############
class Train_Test:
def __init__(self):
print("Train--Test")
def train(self, args, model, optimizer, scheduler, epoch, gpu, data_loader):
# print("EPOCH %d" % epoch)
print('Epoch:', epoch,'LR:', optimizer.param_groups[0]['lr'])
print('Epoch:', epoch,'LR:', scheduler.get_last_lr())
losses = []
model.train()
# loader
data_iter = iter(data_loader)
num_iter = len(data_loader)
for i in tqdm(range(num_iter)):
if args['model'].find("bert") != -1:
inputs_id, segments, masks, ent_embeddings, labels = next(data_iter)
inputs_id, segments, masks, labels = torch.LongTensor(inputs_id), torch.LongTensor(segments), \
torch.LongTensor(masks), torch.FloatTensor(labels)
if args['use_embd_layer']:
#for embedding layer
ent_embeddings = torch.LongTensor(ent_embeddings)
else:
ent_embeddings = None
if gpu >= 0:
if args['use_embd_layer']:
ent_embeddings = ent_embeddings.cuda(gpu)
else:
ent_embeddings = None
inputs_id, segments, masks, labels = inputs_id.cuda(gpu), segments.cuda(gpu), \
masks.cuda(gpu), labels.cuda(gpu)
try:
optimizer.zero_grad()
output, loss = model(inputs_id, segments, masks, ent_embeddings, labels)
except:
print("Unexpected error:", sys.exc_info()[0])
raise
else:
inputs_id, labels, text_inputs, embeddings, tfIdf_inputs = next(data_iter)
if args['use_embd_layer']:
embeddings = torch.LongTensor(embeddings)
if args['use_sentiment']:
input_ids = torch.stack([x_[0][0] for x_ in text_inputs])
attention = torch.stack([x_[1][0] for x_ in text_inputs])
text_inputs = [input_ids,attention]
if args['use_tfIdf']:
tfIdf_inputs = torch.FloatTensor(tfIdf_inputs)
inputs_id, labels = torch.LongTensor(inputs_id), torch.FloatTensor(labels)
optimizer.zero_grad()
output, loss = model(inputs_id, labels, text_inputs, embeddings, tfIdf_inputs)
loss.backward()
if args['grad_clip']:
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
losses.append(loss.item())
return losses
def test(self, args, model, data_path, fold, gpu, dicts, data_loader):
self.model_name = args['model']
filename = data_path.replace('train', fold)
print('file for evaluation: %s' % filename)
num_labels = len(dicts['ind2c'])
y, yhat, yhat_raw, hids, losses = [], [], [], [], []
model.eval()
# loader
data_iter = iter(data_loader)
num_iter = len(data_loader)
for i in tqdm(range(num_iter)):
with torch.no_grad():
if args['model'].find("bert") != -1:
inputs_id, segments, masks, ent_embeddings, labels = next(data_iter)
inputs_id, segments, masks, labels = torch.LongTensor(inputs_id), torch.LongTensor(segments), \
torch.LongTensor(masks), torch.FloatTensor(labels)
if args['use_embd_layer']:
#for embedding layer
ent_embeddings = torch.LongTensor(ent_embeddings)
else:
ent_embeddings = None
if gpu >= 0:
if args['use_embd_layer']:
ent_embeddings = ent_embeddings.cuda(gpu)
else:
ent_embeddings = None
inputs_id, segments, masks, labels = inputs_id.cuda(
gpu), segments.cuda(gpu), masks.cuda(gpu), labels.cuda(gpu)
try:
output, loss = model(inputs_id, segments, masks, ent_embeddings, labels)
except:
print("Unexpected error:", sys.exc_info()[0])
raise
else:
inputs_id, labels, text_inputs, embeddings, tfIdf_inputs = next(data_iter)
if args['use_embd_layer']:
embeddings = torch.LongTensor(embeddings)
if args['use_sentiment']:
input_ids = torch.stack([x_[0][0] for x_ in text_inputs])
attention = torch.stack([x_[1][0] for x_ in text_inputs])
text_inputs = [input_ids,attention]
if args['use_tfIdf']:
tfIdf_inputs = torch.FloatTensor(tfIdf_inputs)
inputs_id, labels = torch.LongTensor(inputs_id), torch.FloatTensor(labels)
output, loss = model(inputs_id, labels, text_inputs, embeddings, tfIdf_inputs)
output = torch.sigmoid(output)
output = output.data.cpu().numpy()
losses.append(loss.item())
target_data = labels.data.cpu().numpy()
yhat_raw.append(output)
output = np.round(output)
y.append(target_data)
yhat.append(output)
y = np.concatenate(y, axis=0)
yhat = np.concatenate(yhat, axis=0)
yhat_raw = np.concatenate(yhat_raw, axis=0)
k = 5 if num_labels == 50 else [8,15]
self.new_metric_calc(y, yhat_raw) #checking my metric values #considering 0 detection as TN
self.calculate_print_metrics(y, yhat_raw) #checking sklearn metric values considering 0 detection as TP
metrics = self.all_metrics(yhat, y, k=k, yhat_raw=yhat_raw)
print()
print("Metric calculation by <NAME> and Hong Yu start")
self.print_metrics(metrics)
print("Metric calculation by Fei Li and Hong Yu end")
print()
metrics['loss_%s' % fold] = np.mean(losses)
print('loss_%s' % fold, metrics['loss_%s' % fold])
return metrics
def new_metric_calc(self, y, yhat):
names = ["acc", "prec", "rec", "f1"]
yhat = np.round(yhat) #rounding the vaues
#Macro
macro_accuracy = np.mean([accuracy_score(y[i], yhat[i]) for i in range(len(y))])
macro_precision = np.mean([self.getPrecision(y[i], yhat[i]) for i in range(len(y))])
macro_recall = np.mean([self.getRecall(y[i], yhat[i]) for i in range(len(y))])
macro_f_score = np.mean([self.getFScore(y[i], yhat[i]) for i in range(len(y))])
#Micro
ymic = y.ravel()
yhatmic = yhat.ravel()
micro_accuracy = accuracy_score(ymic, yhatmic)
micro_precision = self.getPrecision(ymic, yhatmic)
micro_recall = self.getRecall(ymic, yhatmic)
micro_f_score = self.getFScore(ymic, yhatmic)
macro = (macro_accuracy, macro_precision, macro_recall, macro_f_score)
micro = (micro_accuracy, micro_precision, micro_recall, micro_f_score)
metrics = {names[i] + "_macro": macro[i] for i in range(len(macro))}
metrics.update({names[i] + "_micro": micro[i] for i in range(len(micro))})
print()
print("Metric calculation for all labels together start")
self.print_metrics(metrics)
print("Metric calculation for all labels together end")
print()
return metrics
def getFScore(self, y, yhat):
prec = self.getPrecision(y, yhat)
rec = self.getRecall(y, yhat)
if prec + rec == 0:
f1 = 0.
else:
f1 = (2*(prec*rec))/(prec+rec)
return f1
def getRecall(self, y, yhat):
return self.getTP(y, yhat)/(self.getTP(y, yhat) + self.getFN(y, yhat) + 1e-10)
def getPrecision(self, y, yhat):
return self.getTP(y, yhat)/(self.getTP(y, yhat) + self.getFP(y, yhat) + 1e-10)
def getTP(self, y, yhat):
return np.multiply(y, yhat).sum().item()
def getFN(self, y, yhat):
return np.multiply(y, np.logical_not(yhat).astype(float)).sum().item()
def getFP(self, y, yhat):
return np.multiply(np.logical_not(y).astype(float), y).sum().item()
def calculate_print_metrics(self, y, yhat):
names = ["acc", "prec", "rec", "f1"]
yhat = np.round(yhat) #rounding the vaues
macro_precision, macro_recall, macro_f_score, macro_support = precision_recall_fscore_support(y, yhat, average = 'macro', zero_division=1)
# macro_accuracy = ((np.concatenate(np.round(yhat), axis=0) == np.concatenate(y, axis=0)).sum().item()) / len(y) #accuracy_score(y, np.round(yhat))
# macro_accuracy = ((np.round(yhat) == y).sum().item() / len(y[0])) / len(y)
macro_accuracy = np.mean([accuracy_score(y[i], yhat[i]) for i in range(len(y))])
ymic = y.ravel()
yhatmic = yhat.ravel()
micro_precision, micro_recall, micro_f_score, micro_support = precision_recall_fscore_support(ymic, yhatmic, average='micro', zero_division=1)
micro_accuracy = accuracy_score(ymic, yhatmic)
macro = (macro_accuracy, macro_precision, macro_recall, macro_f_score)
micro = (micro_accuracy, micro_precision, micro_recall, micro_f_score)
metrics = {names[i] + "_macro": macro[i] for i in range(len(macro))}
metrics.update({names[i] + "_micro": micro[i] for i in range(len(micro))})
print()
print("Sklearn Metric calculation start")
self.print_metrics(metrics)
print("Sklearn Metric calculation end")
print()
return metrics
def all_metrics(self, yhat, y, k=8, yhat_raw=None, calc_auc=True):
"""
Inputs:
yhat: binary predictions matrix
y: binary ground truth matrix
k: for @k metrics
yhat_raw: prediction scores matrix (floats)
Outputs:
dict holding relevant metrics
"""
names = ["acc", "prec", "rec", "f1"]
#macro
macro = self.all_macro(yhat, y)
#micro
ymic = y.ravel()
yhatmic = yhat.ravel()
micro = self.all_micro(yhatmic, ymic)
metrics = {names[i] + "_macro": macro[i] for i in range(len(macro))}
metrics.update({names[i] + "_micro": micro[i] for i in range(len(micro))})
#AUC and @k
if yhat_raw is not None and calc_auc:
#allow k to be passed as int or list
if type(k) != list:
k = [k]
for k_i in k:
rec_at_k = self.recall_at_k(yhat_raw, y, k_i)
metrics['rec_at_%d' % k_i] = rec_at_k
prec_at_k = self.precision_at_k(yhat_raw, y, k_i)
metrics['prec_at_%d' % k_i] = prec_at_k
metrics['f1_at_%d' % k_i] = 2*(prec_at_k*rec_at_k)/(prec_at_k+rec_at_k)
roc_auc = self.auc_metrics(yhat_raw, y, ymic)
metrics.update(roc_auc)
return metrics
def auc_metrics(self, yhat_raw, y, ymic):
if yhat_raw.shape[0] <= 1:
return
fpr = {}
tpr = {}
roc_auc = {}
#get AUC for each label individually
relevant_labels = []
auc_labels = {}
for i in range(y.shape[1]):
#only if there are true positives for this label
if y[:,i].sum() > 0:
fpr[i], tpr[i], _ = roc_curve(y[:,i], yhat_raw[:,i])
if len(fpr[i]) > 1 and len(tpr[i]) > 1:
auc_score = auc(fpr[i], tpr[i])
if not np.isnan(auc_score):
auc_labels["auc_%d" % i] = auc_score
relevant_labels.append(i)
#macro-AUC: just average the auc scores
aucs = []
for i in relevant_labels:
aucs.append(auc_labels['auc_%d' % i])
roc_auc['auc_macro'] = np.mean(aucs)
#micro-AUC: just look at each individual prediction
yhatmic = yhat_raw.ravel()
fpr["micro"], tpr["micro"], _ = roc_curve(ymic, yhatmic)
roc_auc["auc_micro"] = auc(fpr["micro"], tpr["micro"])
return roc_auc
def precision_at_k(self, yhat_raw, y, k):
#num true labels in top k predictions / k
sortd = np.argsort(yhat_raw)[:,::-1]
topk = sortd[:,:k]
#get precision at k for each example
vals = []
for i, tk in enumerate(topk):
if len(tk) > 0:
num_true_in_top_k = y[i,tk].sum()
denom = len(tk)
vals.append(num_true_in_top_k / float(denom))
return np.mean(vals)
def recall_at_k(self,yhat_raw, y, k):
#num true labels in top k predictions / num true labels
sortd = np.argsort(yhat_raw)[:,::-1]
topk = sortd[:,:k]
#get recall at k for each example
vals = []
for i, tk in enumerate(topk):
num_true_in_top_k = y[i,tk].sum()
denom = y[i,:].sum()
vals.append(num_true_in_top_k / float(denom))
vals = np.array(vals)
vals[np.isnan(vals)] = 0.
return np.mean(vals)
def all_micro(self, yhatmic, ymic):
return self.micro_accuracy(yhatmic, ymic), self.micro_precision(yhatmic, ymic), self.micro_recall(yhatmic, ymic), self.micro_f1(yhatmic, ymic)
def micro_f1(self, yhatmic, ymic):
prec = self.micro_precision(yhatmic, ymic)
rec = self.micro_recall(yhatmic, ymic)
if prec + rec == 0:
f1 = 0.
else:
f1 = 2*(prec*rec)/(prec+rec)
return f1
def micro_recall(self, yhatmic, ymic):
return self.intersect_size(yhatmic, ymic, 0) / (ymic.sum(axis=0) + 1e-10) #NaN fix
def micro_precision(self, yhatmic, ymic):
return self.intersect_size(yhatmic, ymic, 0) / (yhatmic.sum(axis=0) + 1e-10) #NaN fix
def micro_accuracy(self, yhatmic, ymic):
return self.intersect_size(yhatmic, ymic, 0) / (self.union_size(yhatmic, ymic, 0) + 1e-10) #NaN fix
def all_macro(self,yhat, y):
return self.macro_accuracy(yhat, y), self.macro_precision(yhat, y), self.macro_recall(yhat, y), self.macro_f1(yhat, y)
def macro_f1(self, yhat, y):
prec = self.macro_precision(yhat, y)
rec = self.macro_recall(yhat, y)
if prec + rec == 0:
f1 = 0.
else:
f1 = 2*(prec*rec)/(prec+rec)
return f1
def macro_recall(self, yhat, y):
num = self.intersect_size(yhat, y, 0) / (y.sum(axis=0) + 1e-10)
return np.mean(num)
def macro_precision(self, yhat, y):
num = self.intersect_size(yhat, y, 0) / (yhat.sum(axis=0) + 1e-10)
return np.mean(num)
def macro_accuracy(self, yhat, y):
num = self.intersect_size(yhat, y, 0) / (self.union_size(yhat, y, 0) + 1e-10)
return np.mean(num)
def intersect_size(self, yhat, y, axis):
#axis=0 for label-level union (macro). axis=1 for instance-level
return np.logical_and(yhat, y).sum(axis=axis).astype(float)
def union_size(self, yhat, y, axis):
#axis=0 for label-level union (macro). axis=1 for instance-level
return np.logical_or(yhat, y).sum(axis=axis).astype(float)
def print_metrics(self, metrics):
print()
if "auc_macro" in metrics.keys():
print("[MACRO] accuracy, precision, recall, f-measure, AUC")
print(" %.4f, %.4f, %.4f, %.4f, %.4f" % (metrics["acc_macro"], metrics["prec_macro"], metrics["rec_macro"], metrics["f1_macro"], metrics["auc_macro"]))
else:
print("[MACRO] accuracy, precision, recall, f-measure")
print(" %.4f, %.4f, %.4f, %.4f" % (metrics["acc_macro"], metrics["prec_macro"], metrics["rec_macro"], metrics["f1_macro"]))
if "auc_micro" in metrics.keys():
print("[MICRO] accuracy, precision, recall, f-measure, AUC")
print(" %.4f, %.4f, %.4f, %.4f, %.4f" % (metrics["acc_micro"], metrics["prec_micro"], metrics["rec_micro"], metrics["f1_micro"], metrics["auc_micro"]))
else:
print("[MICRO] accuracy, precision, recall, f-measure")
print(" %.4f, %.4f, %.4f, %.4f" % (metrics["acc_micro"], metrics["prec_micro"], metrics["rec_micro"], metrics["f1_micro"]))
for metric, val in metrics.items():
if metric.find("rec_at") != -1:
print("%s: %.4f" % (metric, val))
print()
#############Model Summary###############
import torch
import torch.nn as nn
from torch.autograd import Variable
from collections import OrderedDict
import numpy as np
def My_Summary(model, input_size, batch_size=-1, device="cuda"):
def register_hook(module):
def hook(module, input, output):
class_name = str(module.__class__).split(".")[-1].split("'")[0]
module_idx = len(summary)
m_key = "%s-%i" % (class_name, module_idx + 1)
summary[m_key] = OrderedDict()
summary[m_key]["input_shape"] = list(input[0].size())
summary[m_key]["input_shape"][0] = batch_size
if isinstance(output, (list, tuple)):
summary[m_key]["output_shape"] = [[-1] + list(o.size())[1:] for o in output if len(list(o.size())) > 0][0]
else:
summary[m_key]["output_shape"] = list(output.size())
if len(summary[m_key]["output_shape"]) > 0:
summary[m_key]["output_shape"][0] = batch_size
else:
summary[m_key]["output_shape"] = [-1]
params = 0
if hasattr(module, "weight") and hasattr(module.weight, "size"):
params += torch.prod(torch.LongTensor(list(module.weight.size())))
summary[m_key]["trainable"] = module.weight.requires_grad
if hasattr(module, "bias") and hasattr(module.bias, "size"):
params += torch.prod(torch.LongTensor(list(module.bias.size())))
summary[m_key]["nb_params"] = params
if (
not isinstance(module, nn.Sequential)
and not isinstance(module, nn.ModuleList)
and not (module == model)
):
hooks.append(module.register_forward_hook(hook))
device = device.lower()
assert device in [
"cuda",
"cpu",
], "Input device is not valid, please specify 'cuda' or 'cpu'"
# if device == "cuda" and torch.cuda.is_available():
# dtype = torch.cuda.FloatTensor
# else:
# dtype = torch.FloatTensor
# multiple inputs to the network
if isinstance(input_size, tuple):
input_size = [input_size]
# batch_size of 2 for batchnorm
x = [torch.rand(2, *in_size[0]).type(in_size[1]) if in_size[1] != 0 else None for in_size in input_size]
# print(type(x[0]))
# create properties
summary = OrderedDict()
hooks = []
# register hook
model.apply(register_hook)
# make a forward pass
# print(x.shape)
model(*x)
# remove these hooks
for h in hooks:
h.remove()
print("----------------------------------------------------------------")
line_new = "{:>20} {:>25} {:>15}".format("Layer (type)", "Output Shape", "Param #")
print(line_new)
print("================================================================")
total_params = 0
total_output = 0
trainable_params = 0
for layer in summary:
# input_shape, output_shape, trainable, nb_params
line_new = "{:>20} {:>25} {:>15}".format(
layer,
str(summary[layer]["output_shape"]),
"{0:,}".format(summary[layer]["nb_params"]),
)
total_params += summary[layer]["nb_params"]
total_output += np.prod(summary[layer]["output_shape"])
if "trainable" in summary[layer]:
if summary[layer]["trainable"] == True:
trainable_params += summary[layer]["nb_params"]
print(line_new)
# assume 4 bytes/number (float on cuda).
total_input_size = abs(np.prod([in_size[0][0] for in_size in input_size]) * batch_size * 4. / (1024 ** 2.))
total_output_size = abs(2. * total_output * 4. / (1024 ** 2.)) # x2 for gradients
total_params_size = abs(total_params.numpy() * 4. / (1024 ** 2.))
total_size = total_params_size + total_output_size + total_input_size
print("================================================================")
print("Total params: {0:,}".format(total_params))
print("Trainable params: {0:,}".format(trainable_params))
print("Non-trainable params: {0:,}".format(total_params - trainable_params))
print("----------------------------------------------------------------")
print("Input size (MB): %0.2f" % total_input_size)
print("Forward/backward pass size (MB): %0.2f" % total_output_size)
print("Params size (MB): %0.2f" % total_params_size)
print("Estimated Total Size (MB): %0.2f" % total_size)
print("----------------------------------------------------------------")
# return summary
#############Main###############
class MyDataset(Dataset):
def __init__(self, X):
self.X = X
def __len__(self):
return len(self.X)
def __getitem__(self, idx):
return self.X[idx]
class Run:
def __init__(self, args):
if args['random_seed'] != 0:
random.seed(args['random_seed'])
np.random.seed(args['random_seed'])
torch.manual_seed(args['random_seed'])
torch.cuda.manual_seed_all(args['random_seed'])
print("loading lookups...")
dicts = self.load_lookups(args)
modelhub = ModelHub(args, dicts)
model = modelhub.pick_model(args, dicts)
print(model)
My_Summary(model,
[(tuple([args['MAX_LENGTH']]),torch.LongTensor), (tuple([len(dicts['ind2c'])]),torch.FloatTensor), (tuple([0]),0), (tuple([args['MAX_ENT_LENGTH']]),torch.LongTensor), (tuple([0]),0)],
device="cpu") #inputs_id, labels, text_inputs, embeddings, tfIdf_inputs
if not args['test_model']:
optimizer = optim.Adam(model.parameters(), weight_decay=args['weight_decay'], lr=args['lr'])
# optimizer = optim.AdamW(model.parameters(), lr=args['lr'], betas=(0.9, 0.999), eps=1e-08, weight_decay=args['weight_decay'], amsgrad=True)
else:
optimizer = None
if args['tune_wordemb'] == False:
model.freeze_net()
metrics_hist = defaultdict(lambda: [])
metrics_hist_te = defaultdict(lambda: [])
metrics_hist_tr = defaultdict(lambda: [])
if args['model'].find("bert") != -1:
prepare_instance_func = self.prepare_instance_bert
else:
prepare_instance_func = self.prepare_instance
train_instances = prepare_instance_func(dicts, args['data_path'], args, args['MAX_LENGTH'])
print("train_instances {}".format(len(train_instances)))
dev_instances = prepare_instance_func(dicts, args['data_path'].replace('train','dev'), args, args['MAX_LENGTH'])
print("dev_instances {}".format(len(dev_instances)))
test_instances = prepare_instance_func(dicts, args['data_path'].replace('train','test'), args, args['MAX_LENGTH'])
print("test_instances {}".format(len(test_instances)))
if args['model'].find("bert") != -1:
collate_func = self.my_collate_bertf
else:
collate_func = self.my_collate
train_loader = DataLoader(MyDataset(train_instances), args['batch_size'], shuffle=True, collate_fn=collate_func)
dev_loader = DataLoader(MyDataset(dev_instances), 1, shuffle=False, collate_fn=collate_func)
test_loader = DataLoader(MyDataset(test_instances), 1, shuffle=False, collate_fn=collate_func)
if not args['test_model'] and args['model'].find("bert") != -1:
#original start
param_optimizer = list(model.named_parameters())
param_optimizer = [n for n in param_optimizer if 'pooler' not in n[0]]
no_decay = ['bias', 'LayerNorm.bias', 'LayerNorm.weight']
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay': 0.01},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)], 'weight_decay': 0.0}
]
num_train_optimization_steps = int(
len(train_instances) / args['batch_size'] + 1) * args['n_epochs']
# optimizer = AdamW(optimizer_grouped_parameters, lr=args['lr'], eps=1e-8)
# optimizer = BertAdam(optimizer_grouped_parameters,
# lr=args['lr'],
# warmup=0.1,
# e=1e-8,
# t_total=num_train_optimization_steps)
optimizer = BertAdam(optimizer_grouped_parameters,
lr=args['lr'],
warmup=0.1,
t_total=num_train_optimization_steps)
#original end
scheduler = StepLR(optimizer, step_size=args['step_size'], gamma=args['gamma'])
test_only = args['test_model'] is not None
train_test = Train_Test()
for epoch in range(args['n_epochs']):
if epoch == 0 and not args['test_model'] and args['save_everything']:
model_dir = os.path.join(args['MODEL_DIR'], '_'.join([args['model'], time.strftime('%b_%d_%H_%M_%S', time.localtime())]))
os.makedirs(model_dir)
elif args['test_model']:
model_dir = os.path.dirname(os.path.abspath(args['test_model']))
if not test_only:
epoch_start = time.time()
losses = train_test.train(args, model, optimizer, scheduler, epoch, args['gpu'], train_loader)
loss = np.mean(losses)
epoch_finish = time.time()
print("epoch finish in %.2fs, loss: %.4f" % (epoch_finish - epoch_start, loss))
else:
loss = np.nan
if epoch == args['n_epochs'] - 1:
print("last epoch: testing on dev and test sets")
test_only = True
# test on dev
evaluation_start = time.time()
metrics = train_test.test(args, model, args['data_path'], "dev", args['gpu'], dicts, dev_loader)
evaluation_finish = time.time()
print("evaluation finish in %.2fs" % (evaluation_finish - evaluation_start))
if test_only or epoch == args['n_epochs'] - 1:
metrics_te = train_test.test(args, model, args['data_path'], "test", args['gpu'], dicts, test_loader)
else:
metrics_te = defaultdict(float)
if args['use_schedular']:
#Update scheduler
scheduler.step()
metrics_tr = {'loss': loss}
metrics_all = (metrics, metrics_te, metrics_tr)
for name in metrics_all[0].keys():
metrics_hist[name].append(metrics_all[0][name])
for name in metrics_all[1].keys():
metrics_hist_te[name].append(metrics_all[1][name])
for name in metrics_all[2].keys():
metrics_hist_tr[name].append(metrics_all[2][name])
metrics_hist_all = (metrics_hist, metrics_hist_te, metrics_hist_tr)
if args['save_everything']:
self.save_everything(args, metrics_hist_all, model, model_dir, None, args['criterion'], test_only)
sys.stdout.flush()
if test_only:
break
if args['criterion'] in metrics_hist.keys():
if self.early_stop(metrics_hist, args['criterion'], args['patience']):
#stop training, do tests on test and train sets, and then stop the script
print("%s hasn't improved in %d epochs, early stopping..." % (args['criterion'], args['patience']))
test_only = True
args['test_model'] = '%s/model_best_%s.pth' % (model_dir, args['criterion'])
model = modelhub.pick_model(args, dicts)
def load_lookups(self, args):
csv.field_size_limit(sys.maxsize)
ind2w, w2ind = self.load_vocab_dict(args, args['vocab'])
#get code and description lookups
if args['Y'] == 'full':
ind2c = self.load_full_codes(args['data_path'], version=args['version'])
else:
codes = set()
with open("%sTOP_%s_CODES.csv" % (args['out_path'], str(args['Y'])), 'r') as labelfile:
lr = csv.reader(labelfile)
for i,row in enumerate(lr):
codes.add(row[0])
ind2c = {i:c for i,c in enumerate(sorted(codes))}
c2ind = {c:i for i,c in ind2c.items()}
dicts = {'ind2w': ind2w, 'w2ind': w2ind, 'ind2c': ind2c, 'c2ind': c2ind}
return dicts
def load_vocab_dict(self, args, vocab_file):
vocab = set()
with open(vocab_file, 'r') as vocabfile:
for i, line in enumerate(vocabfile):
line = line.rstrip()
# if line.strip() in vocab:
# print(line)
if line != '':
vocab.add(line.strip())
ind2w = {i + 1: w for i, w in enumerate(sorted(vocab))}
w2ind = {w: i for i, w in ind2w.items()}
return ind2w, w2ind
def load_full_codes(self,train_path, version='mimic3'):
csv.field_size_limit(sys.maxsize)
codes = set()
for split in ['train', 'dev', 'test']:
with open(train_path.replace('train', split), 'r') as f:
lr = csv.reader(f)
next(lr)
for row in lr:
for code in row[3].split(';'): #codes are in 3rd position after removing hadm_id, 3 when hadm id
codes.add(code)
codes = set([c for c in codes if c != ''])
ind2c = defaultdict(str, {i:c for i,c in enumerate(sorted(codes))})
return ind2c
def prepare_instance(self, dicts, filename, args, max_length):
#columns : SUBJECT_ID, HADM_ID, TEXT, LABELS, ENTITY_ID, length
print("reading from file=",filename)
csv.field_size_limit(sys.maxsize)
ind2w, w2ind, ind2c, c2ind = dicts['ind2w'], dicts['w2ind'], dicts['ind2c'], dicts['c2ind']
instances = []
num_labels = len(dicts['ind2c'])
if args['use_embd_layer']:
ent2vec = KGEntityToVec().getEntityToVec()
keys_list = list(ent2vec.keys())
if args['use_sentiment']:
tokenizer = tr.AutoTokenizer.from_pretrained(str(args['sentiment_bert']))
if args['use_tfIdf']:
data_to_use = pd.read_csv(filename)
X_data = data_to_use['TEXT']
X_data = [text.replace('[CLS]','').replace('[SEP]','') for text in X_data]
vectorizer = TfidfVectorizer(max_features=300)
df_data = vectorizer.fit_transform(X_data)
sequences_data = dict(zip(vectorizer.get_feature_names_out(), df_data.toarray()[0]+1))
del data_to_use
del X_data
del vectorizer
del df_data
with open(filename, 'r') as infile:
r = csv.reader(infile)
#header
next(r)
count = 0
for row in tqdm(r):
text = row[2] #text is in 2nd column after removing hadm_id, 2 if HADM
labels_idx = np.zeros(num_labels)
labelled = False
for l in row[3].split(';'): #labels are in 3rd column after removing hadm_id, 3 if HADM
if l in c2ind.keys():
code = int(c2ind[l])
labels_idx[code] = 1
labelled = True
if not labelled:
continue
tokens_ = text.split()
tokens = []
tokens_id = []
for token in tokens_:
if token == '[CLS]' or token == '[SEP]':
continue
tokens.append(token)
token_id = w2ind[token] if token in w2ind else len(w2ind) + 1
tokens_id.append(token_id)
if len(tokens) > max_length:
tokens = tokens[:max_length]
tokens_id = tokens_id[:max_length]
if args['use_sentiment']:
tokens = text.replace('[CLS]', '').replace('[SEP]', '')
#Bert models can use max 512 tokens
tokens = tokenizer(tokens,
padding='max_length',
truncation=True,
max_length=512,
return_tensors='pt')
if args['use_tfIdf']:
tf_idf = [sequences_data[token] if token in sequences_data else 1.0 for token in tokens]
else:
tf_idf = None
if args['use_embd_layer']:
#getting entity embeddings from KG. Each entity embd is of 200d. Extending to to create a single array.
entities = row[4] #entities are stored in 4th column
entities_ = entities.split()
ent_found = False
#for use in embedding layer
entities_id = set()
for entity in entities_[:args['MAX_ENT_LENGTH']]:
ent_id = keys_list.index(entity)+1 if entity in keys_list else len(keys_list) + 1
entities_id.add(ent_id)
ent_found = True
if not ent_found:
continue
entity_embeddings = list(entities_id)
else:
entity_embeddings = None
dict_instance = {'label': labels_idx,
'tokens': tokens,
"entity_embd":entity_embeddings,
"tokens_id": tokens_id,
"tf_idf": tf_idf
}
instances.append(dict_instance)
count += 1
if args['instance_count'] != 'full' and count == int(args['instance_count']):
break
return instances
def prepare_instance_bert(self, dicts, filename, args, max_length):
#columns : SUBJECT_ID, HADM_ID, TEXT, LABELS, ENTITY_ID, length
csv.field_size_limit(sys.maxsize)
ind2w, w2ind, ind2c, c2ind = dicts['ind2w'], dicts['w2ind'], dicts['ind2c'], dicts['c2ind']
instances = []
num_labels = len(dicts['ind2c'])
wp_tokenizer = tr.BertTokenizer.from_pretrained(args['pretrained_bert'], do_lower_case=True)
ent2vec = KGEntityToVec().getEntityToVec()
if args['use_embd_layer']:
keys_list = list(ent2vec.keys())
with open(filename, 'r') as infile:
r = csv.reader(infile)
#header
next(r)
count = 0
for row in tqdm(r):
text = row[2] #text is in 2nd column now after removing hadm_id, if HADM_ID then text is in 3rd column
labels_idx = np.zeros(num_labels)
labelled = False
for l in row[3].split(';'): #labels are in 3rd column after removing hadm_id
if l in c2ind.keys():
code = int(c2ind[l])
labels_idx[code] = 1
labelled = True
if not labelled:
continue
# original 2 start
##Changes made by prantik for obove code start
tokens = wp_tokenizer.tokenize(text)
tokens = list(filter(lambda a: (a != "[CLS]" and a != "[SEP]"), tokens))[0:max_length-2]
tokens.insert(0, '[CLS]')
tokens.append('[SEP]')
##Changes made by prantik for obove code end
tokens_id = wp_tokenizer.convert_tokens_to_ids(tokens)
masks = [1] * len(tokens)
segments = [0] * len(tokens)
# original 2 end
#getting entity embeddings from KG. Each entity embd is of 200d. Extending to to create a single array.
entities = row[4] #entities are stored in 4th column
entities_ = entities.split()
ent_found = False
if args['use_embd_layer']:
#for use in embedding layer
entities_id = set()
for entity in entities_[:args['MAX_ENT_LENGTH']]:
ent_id = keys_list.index(entity)+1 if entity in keys_list else len(keys_list) + 1
entities_id.add(ent_id)
ent_found = True
if not ent_found:
continue
entity_embeddings = list(entities_id)
else:
entity_embeddings = None
dict_instance = {'label':labels_idx, 'tokens':tokens, "entity_embd":entity_embeddings,
"tokens_id":tokens_id, "segments":segments, "masks":masks}
instances.append(dict_instance)
count += 1
if args['instance_count'] != 'full' and count == int(args['instance_count']):
break
return instances
def my_collate(self, x):
words = [x_['tokens_id'] for x_ in x]
max_seq_len = max([len(w) for w in words])
if max_seq_len < args['MAX_LENGTH']:
max_seq_len = args['MAX_LENGTH']
inputs_id = self.pad_sequence(words, max_seq_len)
labels = [x_['label'] for x_ in x]
if args['use_sentiment']:
text_inputs = [[x_['tokens']['input_ids'], x_['tokens']['attention_mask']] for x_ in x]
else:
text_inputs = []
embeddings = None
if args['use_embd_layer']:
embeddings = [x_['entity_embd'] for x_ in x]
emb_list = [len(x) for x in embeddings]
max_embd_len = max(emb_list)
if max_embd_len < args['MAX_ENT_LENGTH']:
max_embd_len = args['MAX_ENT_LENGTH']
embeddings = self.pad_sequence(embeddings, max_embd_len)
tfIdf_inputs = None
if args['use_tfIdf']:
tfIdf_inputs = [x_['tf_idf'] for x_ in x]
tfIdf_inputs = self.pad_sequence(tfIdf_inputs, max_seq_len, np.float)
return inputs_id, labels, text_inputs, embeddings, tfIdf_inputs
def my_collate_bert(self, x):
words = [x_['tokens_id'] for x_ in x]
segments = [x_['segments'] for x_ in x]
masks = [x_['masks'] for x_ in x]
embeddings = [x_['entity_embd'] for x_ in x]
seq_len = [len(w) for w in words]
max_seq_len = max(seq_len)
if args['use_embd_layer']:
#for embedding layer
max_embd_len = max([len(x) for x in embeddings])
if max_embd_len < args['MAX_ENT_LENGTH']:
max_embd_len = args['MAX_ENT_LENGTH']
try:
inputs_id = self.pad_sequence(words, max_seq_len)
segments = self.pad_sequence(segments, max_seq_len)
masks = self.pad_sequence(masks, max_seq_len)
if args['use_embd_layer']:
#for embedding layer
embeddings = self.pad_sequence(embeddings, max_embd_len)
except:
print("Unexpected error:", sys.exc_info()[0])
raise
labels = [x_['label'] for x_ in x]
return inputs_id, segments, masks, embeddings, labels
def pad_sequence(self, x, max_len, type=np.int):
padded_x = np.zeros((len(x), max_len), dtype=type)
for i, row in enumerate(x):
if max_len >= len(row):
padded_x[i][:len(row)] = row
else:
padded_x[i][:max_len] = row[:max_len] #trancate
return padded_x
def save_metrics(self, metrics_hist_all, model_dir):
with open(model_dir + "/metrics.json", 'w') as metrics_file:
#concatenate dev, train metrics into one dict
data = metrics_hist_all[0].copy()
data.update({"%s_te" % (name):val for (name,val) in metrics_hist_all[1].items()})
data.update({"%s_tr" % (name):val for (name,val) in metrics_hist_all[2].items()})
json.dump(data, metrics_file, indent=1)
def save_everything(self, args, metrics_hist_all, model, model_dir, params, criterion, evaluate=False):
self.save_args(args, model_dir)
self.save_metrics(metrics_hist_all, model_dir)
if not evaluate:
#save the model with the best criterion metric
if not np.all(np.isnan(metrics_hist_all[0][criterion])):
if criterion == 'loss_dev':
eval_val = np.nanargmin(metrics_hist_all[0][criterion])
else:
eval_val = np.nanargmax(metrics_hist_all[0][criterion])
if eval_val == len(metrics_hist_all[0][criterion]) - 1:
print("saving model==")
sd = model.cpu().state_dict()
torch.save(sd, model_dir + "/model_best_%s.pth" % criterion)
if args['gpu'] >= 0:
model.cuda(args['gpu'])
print("saved metrics, params, model to directory %s\n" % (model_dir))
def save_args(self, args, model_path):
file_path = model_path + "/args.json"
if not os.path.exists(file_path):
with open(file_path, 'w') as args_file:
json.dump(args, args_file)
def early_stop(self, metrics_hist, criterion, patience):
if not np.all(np.isnan(metrics_hist[criterion])):
if len(metrics_hist[criterion]) >= patience:
if criterion == 'loss_dev':
return np.nanargmin(metrics_hist[criterion]) < len(metrics_hist[criterion]) - patience
else:
return np.nanargmax(metrics_hist[criterion]) < len(metrics_hist[criterion]) - patience
else:
return False
# -
#Set proper path values in args{} and hit for data processig and saving.
DataProcessing(args)
# + colab={"base_uri": "https://localhost:8080/"} id="lLu1OLM0gtqe" outputId="eb289489-e36e-460d-cea3-fd803c32fcd4"
#Set proper values in args{} and hit for training, validating and testing.
Run(args)
| KG_MultiResCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# HIDDEN CELL
import sys, os
# Importing argopy in dev mode:
on_rtd = os.environ.get('READTHEDOCS', None) == 'True'
if not on_rtd:
sys.path.insert(0, "/Users/gmaze/git/github/euroargodev/argopy")
import git
import argopy
from argopy.options import OPTIONS
print("argopy:", argopy.__version__,
"\nsrc:", argopy.__file__,
"\nbranch:", git.Repo(search_parent_directories=True).active_branch.name,
"\noptions:", OPTIONS)
else:
sys.path.insert(0, os.path.abspath('..'))
import xarray as xr
# xr.set_options(display_style="html");
xr.set_options(display_style="text");
# -
import argopy
from argopy import DataFetcher as ArgoDataFetcher
# + raw_mimetype="text/restructuredtext" active=""
# .. _user-mode:
# + [markdown] raw_mimetype="text/restructuredtext"
# # User mode: standard vs expert
# -
# **Problem**
#
# For beginners or non-experts of the Argo dataset, it can be quite complicated to get access to Argo measurements. Indeed, the Argo data set is very complex, with thousands of different variables, tens of reference tables and a [user manual](https://doi.org/10.13155/29825) more than 100 pages long.
#
# This is mainly due to:
#
# - Argo measurements coming from many different models of floats or sensors,
# - quality control of *in situ* measurements of autonomous platforms being really a matter of ocean and data experts,
# - the Argo data management workflow being distributed between more than 10 Data Assembly Centers all around the world,
# - the Argo autonomous profiling floats, despite quite a simple principle of functionning, is a rather complex robot that needs a lot of data to be monitored and logged.
#
# **Solution**
#
# In order to ease Argo data analysis for the vast majority of standard users, we implemented in **argopy** different levels of verbosity and data processing to hide or simply remove variables only meaningful to experts.
# ## Which type of user are you ?
#
# If you don't know in which user category you would place yourself, try to answer the following questions:
#
# - what is a WMO number ?
# - what is the difference between Delayed and Real Time data mode ?
# - what is an adjusted parameter ?
# - what a QC flag of 3 means ?
#
# If you answered to no more than 1 question, you probably would feel more confortable with the **standard** user mode.
# Otherwise, you can give a try to the **expert** mode.
#
# In **standard** mode, fetched data are automatically filtered to account for their quality (only good are retained) and level of processing by the data centers (wether they looked at the data briefly or not).
# ## Setting the user mode
# By default, all **argopy** data fetchers are set to work with a **standard** user mode.
#
# If you want to change the user mode, or simply makes it explicit, you can use:
#
# - **argopy** global options:
argopy.set_options(mode='standard')
# - a temporary context:
with argopy.set_options(mode='standard'):
ArgoDataFetcher().profile(6902746, 34)
# - option when instantiating the data fetcher:
ArgoDataFetcher(mode='standard').profile(6902746, 34)
# ## Differences in user modes
#
# To highlight that, let's compare data fetched for one profile with each modes.
#
# You will note that the **standard** mode has fewer variables to let you focus on your analysis.
# For **expert**, all Argo variables for you to work with are here.
#
# The difference is the most visible when fetching Argo data from a local copy of the GDAC ftp, so let's use a sample of this provided by **argopy** tutorial datasets:
ftproot, flist = argopy.tutorial.open_dataset('localftp')
argopy.set_options(local_ftp=ftproot)
# In **standard** mode:
with argopy.set_options(mode='standard'):
ds = ArgoDataFetcher(src='localftp').profile(6901929, 2).to_xarray()
print(ds.data_vars)
# In **expert** mode:
with argopy.set_options(mode='expert'):
ds = ArgoDataFetcher(src='localftp').profile(6901929, 2).to_xarray()
print(ds.data_vars)
| docs/user_mode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sklearn,numpy,scipy,matplotlib
from matplotlib.pylab import plt
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.datasets import make_blobs
plt.rcParams['image.cmap'] = "gray"
plt.rcParams['axes.xmargin'] = 0.05
plt.rcParams['axes.ymargin'] = 0.05
cancer = load_breast_cancer()
print("cancer.data=",cancer.data.shape)
print("cancer.target=", cancer.target.shape)
X_train, X_test, y_train, y_test = train_test_split(cancer.data, cancer.target,random_state=1)
print("train X=",X_train.shape)
print("train y=",y_train.shape)
print("test X=",X_test.shape)
print("test y=",y_test.shape)
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
print(X_test_scaled.shape)
print(X_test_scaled.min(axis=0).shape)
print(X_test_scaled.max(axis=0).shape)
# +
"""Examples illustrating the use of plt.subplots().
This function creates a figure and a grid of subplots with a single call, while
providing reasonable control over how the individual plots are created. For
very refined tuning of subplot creation, you can still use add_subplot()
directly on a new figure.
"""
import matplotlib.pyplot as plt
import numpy as np
# Simple data to display in various forms
x = np.linspace(0, 2 * np.pi, 400)
y = np.sin(x ** 2)
plt.close('all')
# Just a figure and one subplot
f, ax = plt.subplots()
ax.plot(x, y)
ax.set_title('Simple plot')
# Two subplots, the axes array is 1-d
f, axarr = plt.subplots(2, sharex=True)
axarr[0].plot(x, y)
axarr[0].set_title('Sharing X axis')
axarr[1].scatter(x, y)
# Two subplots, unpack the axes array immediately
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True)
ax1.plot(x, y)
ax1.set_title('Sharing Y axis')
ax2.scatter(x, y)
# Three subplots sharing both x/y axes
f, (ax1, ax2, ax3) = plt.subplots(3, sharex=True, sharey=True)
ax1.plot(x, y)
ax1.set_title('Sharing both axes')
ax2.scatter(x, y)
ax3.scatter(x, 2 * y ** 2 - 1, color='r')
# Fine-tune figure; make subplots close to each other and hide x ticks for
# all but bottom plot.
f.subplots_adjust(hspace=0)
plt.setp([a.get_xticklabels() for a in f.axes[:-1]], visible=False)
# row and column sharing
f, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, sharex='col', sharey='row')
ax1.plot(x, y)
ax1.set_title('Sharing x per column, y per row')
ax2.scatter(x, y)
ax3.scatter(x, 2 * y ** 2 - 1, color='r')
ax4.plot(x, 2 * y ** 2 - 1, color='r')
# Four axes, returned as a 2-d array
f, axarr = plt.subplots(2, 2)
axarr[0, 0].plot(x, y)
axarr[0, 0].set_title('Axis [0,0]')
axarr[0, 1].scatter(x, y)
axarr[0, 1].set_title('Axis [0,1]')
axarr[1, 0].plot(x, y ** 2)
axarr[1, 0].set_title('Axis [1,0]')
axarr[1, 1].scatter(x, y ** 2)
axarr[1, 1].set_title('Axis [1,1]')
# Fine-tune figure; hide x ticks for top plots and y ticks for right plots
plt.setp([a.get_xticklabels() for a in axarr[0, :]], visible=False)
plt.setp([a.get_yticklabels() for a in axarr[:, 1]], visible=False)
# Four polar axes
f, axarr = plt.subplots(2, 2, subplot_kw=dict(projection='polar'))
axarr[0, 0].plot(x, y)
axarr[0, 0].set_title('Axis [0,0]')
axarr[0, 1].scatter(x, y)
axarr[0, 1].set_title('Axis [0,1]')
axarr[1, 0].plot(x, y ** 2)
axarr[1, 0].set_title('Axis [1,0]')
axarr[1, 1].scatter(x, y ** 2)
axarr[1, 1].set_title('Axis [1,1]')
# Fine-tune figure; make subplots farther from each other.
f.subplots_adjust(hspace=0.3)
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
import sklearn,numpy,scipy,matplotlib
#x = np.linspace(0, 2* np.pi, 400)
x = np.arange(0, 2* np.pi, 0.01)
y = np.sin(x)
y2 = np.cos(x**2)
plt.close('all')
f, ax = plt.subplots(2,2, subplot_kw=dict(projection='polar'))
ax[0,0].plot(x,y)
ax[0,0].set_title("axis [0,0]")
ax[0,1].scatter(x,y)
ax[0,1].set_title("axis [0,1]")
ax[1,0].plot(x,y2)
ax[1,0].set_title("axis [1,0]")
ax[1,1].scatter(x,y2)
ax[1,1].set_title("axis [1,1]")
plt.show()
# +
from matplotlib.pylab import plt
from sklearn.datasets import make_blobs
import sklearn,numpy,scipy,matplotlib
import mglearn
X, _ = make_blobs( n_samples=50, centers=5, random_state=4, cluster_std=2)
# 훈련 세트와 테스트 세트로 나눕니다
X_train, X_test = train_test_split(X, random_state=5, test_size=.1)
# 훈련 세트와 테스트 세트의 산점도를 그립니다
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
axes[0].scatter(X_train[:, 0], X_train[:, 1],
c=mglearn.cm2(0), label="Train", s=60)
axes[0].scatter(X_test[:, 0], X_test[:, 1], marker='^',
c=mglearn.cm2(1), label="Test", s=60)
axes[0].legend(loc='upper left')
axes[0].set_title("Original")
plt.show()
# -
import numpy
a = numpy.arange(0,100, 10)
print(a)
| Test3/Working.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
# ### Define Model
class DBN:
def __init__(self, n_v, layers, k=1, mean_field=True):
if n_v is None or layers is None: raise ValueError("Incorrect inputs for layer 0.")
n_hs = [n_v]
n_layer = 0
training_done = True
rbms = []
for (n_h, model) in layers:
n_layer += 1
if model is None:
if n_h <= 0: raise ValueError("Incorrect inputs for layer %d" % (n_layer))
else: n_hs.append(n_h)
rbm = RBM(n_hs[n_layer-1], n_h, k=k)
training_done = False
else:
assert training_done
rbm = RBM.load_model(model)
assert rbm.n_v == n_hs[n_layer-1]
n_hs.append(rbm.n_h)
rbms.append(rbm)
self.n_hs = n_hs
self.n_layer = n_layer
self.training_done = training_done
self.rbms = rbms
self.mean_field = mean_field
return
def forward(self, X):
assert self.training_done
Hp = X
for i in range(self.n_layer):
Hp, Hs = self.rbms[i].forward(Hp)
return Hp, Hs
def backward(self, H):
assert self.training_done
Vp = H
for i in reversed(range(self.n_layer)):
Vp, Vs = self.rbms[i].backward(Vp)
return Vp, Vs
def train_1layer_1batch(self, V, layer):
Hp = V
for i in range(layer):
Hp, Hs = self.rbms[i].forward(Hp)
if self.mean_field or layer==0:
self.rbms[layer].contrastive_divergence(Hp, self.learning)
else:
for j in range(5):
Vs = np.random.binomial(1, Hp, size=Hp.shape)
self.rbms[layer].contrastive_divergence(Vs, self.learning)
return
def train_1layer(self, X, layer, epochs, batch_size=10, learning=0.01, save_file=None):
assert not self.rbms[layer].training_done
self.learning = learning
n_x = X.shape[0]
n_batch = n_x//batch_size
startrate = learning
for e in range(epochs):
learning = startrate * math.pow(0.1, e//50)
for i in range(n_batch):
s = i*batch_size
V = X[s:s+batch_size]
self.train_1layer_1batch(V, layer)
self.rbms[layer].save_model(save_file+"-"+str(layer+1)+".epochs"+str(epochs))
self.rbms[layer].training_done = True
return
# this is internal API for more complex network to use
def train_model(self, X, epochs=1, batch_size=10, learning=0.01, save_file=None):
for layer in range(self.n_layer):
if self.rbms[layer].training_done: continue
self.train_1layer(X, layer, epochs, batch_size, learning, save_file)
return
# this is the API for app to use
def train(self, X, epochs=1, batch_size=10, learning=0.01, save_file=None):
if self.training_done: return
save_file += ".dbn.layer" + str(self.n_layer)
self.train_model(X, epochs, batch_size, learning, save_file)
self.training_done = True
return
def reconstruct(self, X):
h_layer = self.n_layer - 1
Hp = X
for i in range(h_layer):
Hp, Hs = self.rbms[i].forward(Hp)
Vp, Vs = self.rbms[h_layer].reconstruct(Hp)
for i in reversed(range(h_layer)):
Hp, Hs = self.rbms[i].backward(Hp)
return Hp, Hs
def show_features(self, shapes, title, count=-1):
#for i in range(self.n_layer):
# rbm = self.rbms[i]
# rbm.show_features(shapes[i], "MNIST learned features in DBN layer %d" % (i+1), count)
lower_rbm = None
for i in range(self.n_layer):
rbm = self.rbms[i]
if i!=0:
W = np.dot(rbm.W, lower_rbm.W)
rbm = RBM(W.shape[1], W.shape[0], W=W)
rbm.show_features(shapes[0], title +" learned features in layer %d" % (i+1), count)
lower_rbm = rbm
return lower_rbm
# + run_control={"marked": false}
class MNIST_DBN:
def __init__(self, n_v, layers, data_path):
self.dbn = DBN(n_v, layers)
self.train_input = MnistInput("train", data_path)
self.test_input = MnistInput("test", data_path)
return
def train(self, train_size=-1, n_epoch=1, batch_size=10, learning=0.01):
X = []
n_x = 0
for x, y in self.train_input.read(train_size):
X.append(x)
n_x += 1
X = np.array(X).reshape(n_x, -1) > 30
X = X*1 / 255
self.dbn.train(X, n_epoch, batch_size, learning, save_file="mnist")
return
def test_reconstruct(self, n):
X=[]; i=2*n
for x, y in self.test_input.read(n):
x *= np.random.binomial(1, i/(n*2), size=(28,28))
x = x * 2*n/i
x /= 255
X.append(x)
i -=1
recon_X = []
for i in range(n):
Vp, Vs = self.dbn.reconstruct(X[i].reshape(1, -1))
recon_X.append(Vp)
return np.array(X).reshape(-1, 784), np.array(recon_X).reshape(-1, 784)
# -
def mult(tuple):
prod = 1
for i in tuple:
prod *= i
return prod
# ### Training
# +
# %run "RBM-backup.ipynb"
mnist_dbn = None
if __name__ == "__main__" and '__file__' not in globals():
np.seterr(all='raise')
plt.close('all')
v = (28,28); h1 = (20,25); h2 = (20,25); h3 = (1000,1)
#layers = [
# (mult(h1), None), # (dimension, "model_file") of hidden layer 1
# (mult(h2), None), # (dimension, "model_file") of hidden layer 2
# (mult(h3), None) # (dimension, "model_file") of hidden layer 3
#]
layers = [
(mult(h1), "trained_models/mnist_rbm.784x500.epochs100"), # hidden layer 1
(mult(h2), "trained_models/mnist.dbn.layer3-2.epochs100.500x500"),
(mult(h3), "trained_models/mnist.dbn.layer3-3.epochs100.500x1000")
]
#set_trace()
mnist_dbn = MNIST_DBN(mult(v), layers, "..")
mnist_dbn.train(train_size=60000, n_epoch=200, batch_size=100,)
feature_shapes = (v, h1, h2)
mnist_dbn.dbn.show_features(feature_shapes,"MNIST %d-layer DBN " % (len(layers)), 14*3)
x_sample, recon_x = mnist_dbn.test_reconstruct(100)
# -
# ### Image Reconstruction
# +
plt.figure(figsize=(4, 6))
for i in range(5):
plt.subplot(5, 2, 2*i + 1)
plt.imshow(x_sample[i].reshape(28, 28), vmin=0, vmax=1, cmap="gray")
plt.title("Input")
plt.colorbar()
plt.subplot(5, 2, 2*i + 2)
plt.imshow(recon_x[i].reshape(28, 28), vmin=0, vmax=1, cmap="gray")
plt.title("Reconstruction")
plt.colorbar()
plt.tight_layout()
# -
def gen_mnist_image(X):
return np.rollaxis(np.rollaxis(X[0:200].reshape(20, -1, 28, 28), 0, 2), 1, 3).reshape(-1, 20 * 28)
plt.figure(figsize=(10,20))
plt.imshow(gen_mnist_image(recon_x))
# ### Calculate Reconstruction Error
def calculate_recon_error(X, recon_X):
"""
Compute the reconstruction error.
"""
rec_loss = - np.sum(X * np.log(1e-8 + recon_X)
+ (1 - X) * np.log(1e-8 + 1 - recon_X), 1)
return np.mean(rec_loss)
calculate_recon_error(x_sample, recon_x)
| RBM_DBN/DBN_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.models import load_model
from tkinter import *
import tkinter as tk
import win32gui
from PIL import Image, ImageGrab
import numpy as np
model=load_model("mnist.h5")
def predict_digit(img):
#resize the picture to 28x28 pixels
img=img.resize((28,28))
#convert rgb to grayscale
img=img.convert('L')
img=np.array(img)
#reshaping to support our model and input normalizing
img=img.reshape(1,28,28,1)
img=img/255.0
#predicting the class
res=model.predict([img])[0]
return np.argmax(res),max(res)
# +
class App(tk.Tk):
def __init__(self):
tk.Tk.__init__(self)
self.x=self.y=0
#creating elements
self.canvas=tk.Canvas(self,width=300,height=300,bg="white",cursor="cross")
#self.title=tk.Title(self,"Handwritten Digit Recognizer")
self.label=tk.Label(self,text="Draw",font=("Arial", 48))
self.classify_btn=tk.Button(self,text="Recognize",command=self.classify_handwriting)
self.button_clear=tk.Button(self,text="Clear",command=self.clear_all)
#Grid Structure
self.canvas.grid(row=0, column=0, pady=2, sticky=W,)
self.label.grid(row=0, column=1,pady=2, padx=2)
self.classify_btn.grid(row=1, column=1, pady=2, padx=2)
self.button_clear.grid(row=1, column=0, pady=2)
#self.canvas.bind("<Motion>", self.start_pos)
self.canvas.bind("<B1-Motion>",self.draw_lines)
def clear_all(self):
self.canvas.delete("all")
def classify_handwriting(self):
HWND=self.canvas.winfo_id() #for getting the handle of the canvas
rect=win32gui.GetWindowRect(HWND) #for getting cordinates of canvas
a,b,c,d=rect
rect=(a+4,b+4,c-4,d-4)
im=ImageGrab.grab(rect)
digit,acc=predict_digit(im)
self.label.configure(text=str(digit)+', '+str(int(acc*100))+'%')
def draw_lines(self,event):
self.x=event.x
self.y=event.y
r=8
self.canvas.create_oval(self.x-r,self.y-r, self.x + r, self.y + r, fill='black')
app = App()
mainloop()
# -
| Handwritten Recognition/digit_recognizer_gui.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Alibaba Cloud Container Service for Kubernetes (ACK) Deep MNIST Example
# In this example we will deploy a tensorflow MNIST model in the Alibaba Cloud Container Service for Kubernetes.
#
# This tutorial will break down in the following sections:
#
# 1) Train a tensorflow model to predict mnist locally
#
# 2) Containerise the tensorflow model with our docker utility
#
# 3) Test model locally with docker
#
# 4) Set-up and configure Alibaba Cloud environment
#
# 5) Deploy your model and visualise requests
#
# Let's get started! 🚀🔥
#
# ## Dependencies:
#
# * Helm v3.0.0+
# * kubectl v1.14+
# * Python 3.6+
# * Python DEV requirements
#
# ## 1) Train a tensorflow model to predict mnist locally
# We will load the mnist images, together with their labels, and then train a tensorflow model to predict the right labels
# +
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot = True)
import tensorflow as tf
if __name__ == '__main__':
x = tf.placeholder(tf.float32, [None,784], name="x")
W = tf.Variable(tf.zeros([784,10]))
b = tf.Variable(tf.zeros([10]))
y = tf.nn.softmax(tf.matmul(x,W) + b, name="y")
y_ = tf.placeholder(tf.float32, [None, 10])
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y), reduction_indices=[1]))
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
init = tf.initialize_all_variables()
sess = tf.Session()
sess.run(init)
for i in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(100)
sess.run(train_step, feed_dict={x: batch_xs, y_: batch_ys})
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print(sess.run(accuracy, feed_dict = {x: mnist.test.images, y_:mnist.test.labels}))
saver = tf.train.Saver()
saver.save(sess, "model/deep_mnist_model")
# -
# ## 2) Containerise the tensorflow model with our docker utility
# Create a wrapper file that exposes the functionality through a `predict` function:
# +
# %%writefile DeepMnist.py
import tensorflow as tf
import numpy as np
class DeepMnist(object):
def __init__(self):
self.class_names = ["class:{}".format(str(i)) for i in range(10)]
self.sess = tf.Session()
saver = tf.train.import_meta_graph("model/deep_mnist_model.meta")
saver.restore(self.sess,tf.train.latest_checkpoint("./model/"))
graph = tf.get_default_graph()
self.x = graph.get_tensor_by_name("x:0")
self.y = graph.get_tensor_by_name("y:0")
def predict(self,X,feature_names):
predictions = self.sess.run(self.y,feed_dict={self.x:X})
return predictions.astype(np.float64)
# -
# Define the dependencies for the wrapper in the requirements.txt:
# %%writefile requirements.txt
tensorflow>=1.12.0
# You need to make sure that you have added the .s2i/environment configuration file in this folder with the following content:
# !mkdir .s2i
# %%writefile .s2i/environment
MODEL_NAME=DeepMnist
API_TYPE=REST
SERVICE_TYPE=MODEL
PERSISTENCE=0
# Now we can build a docker image named "deep-mnist" with the tag 0.1
# !s2i build . seldonio/seldon-core-s2i-python36:1.3.0-dev deep-mnist:0.1
# ## 3) Test model locally with docker
# We first run the docker image we just created as a container called "mnist_predictor"
# !docker run --name "mnist_predictor" -d --rm -p 5000:5000 deep-mnist:0.1
# Send some random features that conform to the contract
import matplotlib.pyplot as plt
import numpy as np
# This is the variable that was initialised at the beginning of the file
i = [0]
x = mnist.test.images[i]
y = mnist.test.labels[i]
plt.imshow(x.reshape((28, 28)), cmap='gray')
plt.show()
print("Expected label: ", np.sum(range(0,10) * y), ". One hot encoding: ", y)
# +
from seldon_core.seldon_client import SeldonClient
import math
import numpy as np
# We now test the REST endpoint expecting the same result
endpoint = "0.0.0.0:5000"
batch = x
payload_type = "ndarray"
sc = SeldonClient(microservice_endpoint=endpoint)
# We use the microservice, instead of the "predict" function
client_prediction = sc.microservice(
data=batch,
method="predict",
payload_type=payload_type,
names=["tfidf"])
for proba, label in zip(client_prediction.response.data.ndarray.values[0].list_value.ListFields()[0][1], range(0,10)):
print(f"LABEL {label}:\t {proba.number_value*100:6.4f} %")
# -
# !docker rm mnist_predictor --force
# ## 4) Set-up and configure Alibaba Cloud environment
# ### 4.1 create a managed kubernetes cluster
# We need to first create a cluster in Alibaba Cloud - you should follow the following instructions (make sure you expose the cluster with an elastic IP by checking the tickbox): https://www.alibabacloud.com/help/doc-detail/95108.htm
#
# You should follow up the instructions but the finished cluster should look as follows:
#
# 
# ### 4.2 Copy the kubectl configuration to access the cluster
# Once you have the cluster created, you will be able to use your local kubectl by copying the configuration details on the overview page, and copy it to your ~/.kube/config
# ### 4.3 Create an Alibaba Container Registry to push the image
# Finally we need to create a container registry repository by following this guide: https://www.alibabacloud.com/blog/up-and-running-with-alibaba-cloud-container-registry_593765
# ## Setup Seldon Core
#
# Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Ambassador Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Ambassador) and [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
# ### Finally we install the Seldon Analytics Package
# + jupyter={"outputs_hidden": true}
# !helm install seldon-core-analytics seldon-core-analytics --repo https://storage.googleapis.com/seldon-charts
# !kubectl rollout status deployment.apps/grafana-prom-deployment
# !kubectl patch svc grafana-prom --type='json' -p '[{"op":"replace","path":"/spec/type","value":"LoadBalancer"}]'
# -
# ### 4.5 Push docker image
# We'll now make sure the image is accessible within the Kubernetes cluster by pushing it to the repo that we created in step 4.3. This should look as follows in your dashboard:
#
# 
# #### To push the image we first tag it
# !docker tag deep-mnist:0.1 registry-intl.eu-west-1.aliyuncs.com/seldonalibaba/deep-mnist:0.1
# #### And then we push it
# !docker push registry-intl.eu-west-1.aliyuncs.com/seldonalibaba/deep-mnist:0.1
# ## 5 - Deploy your model and visualise requests
#
# **IMPORTANT: Make sure you replace the URL for your repo in the format of:**
# * registry-intl.**[REPO]**.aliyuncs.com/**[REGISTRY]**/**[REPO]**:0.1
# %%writefile deep_mnist_deployment.yaml
---
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
labels:
app: seldon
name: deep-mnist
spec:
annotations:
project_name: Tensorflow MNIST
deployment_version: v1
name: deep-mnist
oauth_key: oauth-key
oauth_secret: oauth-secret
predictors:
- componentSpecs:
- spec:
containers:
# REPLACE_FOR_IMAGE_AND_TAG
- image: registry-intl.eu-west-1.aliyuncs.com/seldonalibaba/deep-mnist:0.1
imagePullPolicy: IfNotPresent
name: classifier
resources:
requests:
memory: 1Mi
terminationGracePeriodSeconds: 20
graph:
children: []
name: classifier
endpoint:
type: REST
type: MODEL
name: single-model
replicas: 1
annotations:
predictor_version: v1
# ### Run the deployment in your cluster
kubectl apply -f deep_mnist_deployment.yaml
# ### And let's check that it's been created.
# !kubectl get pods
# ### Test the model
# We'll use a random example from our dataset
import matplotlib.pyplot as plt
import numpy as np
# This is the variable that was initialised at the beginning of the file
i = [0]
x = mnist.test.images[i]
y = mnist.test.labels[i]
plt.imshow(x.reshape((28, 28)), cmap='gray')
plt.show()
print("Expected label: ", np.sum(range(0,10) * y), ". One hot encoding: ", y)
# ### First we need to find the URL that we'll use
# You need to add it to the script in the next block
# !kubectl get svc ambassador -o jsonpath='{.status.loadBalancer.ingress[0].ip}'
# We can now add the URL above to send our request:
# +
from seldon_core.seldon_client import SeldonClient
import math
import numpy as np
import subprocess
# Add the URL you found above, here:
HOST = "192.168.127.12"
port = "80" # Make sure you use the port above
batch = x
payload_type = "ndarray"
sc = SeldonClient(
gateway="ambassador",
gateway_endpoint=HOST + ":" + port,
namespace="default",
oauth_key="oauth-key",
oauth_secret="oauth-secret")
client_prediction = sc.predict(
data=batch,
deployment_name="deep-mnist",
names=["text"],
payload_type=payload_type)
print(client_prediction.response)
# -
# ### Let's see the predictions for each label
# It seems that it correctly predicted the number 7
for proba, label in zip(client_prediction.response.data.ndarray.values[0].list_value.ListFields()[0][1], range(0,10)):
print(f"LABEL {label}:\t {proba.number_value*100:6.4f} %")
# #### Finally let's visualise the metrics that seldon provides out of the box
# For this we can access the URL with the command below, it will request an admin and password which by default are set to the following:
# * Username: admin
# * Password: <PASSWORD>
#
# **You will be able to access it at http://[URL]/d/ejAHFXIWz/prediction-analytics?orgId=1**
# !kubectl get svc grafana-prom -o jsonpath='{.status.loadBalancer.ingress[0].ip}'
# The metrics include requests per second, as well as latency. You are able to add your own custom metrics, and try out other more complex deployments by following furher guides at https://docs.seldon.io/projects/seldon-core/en/latest/examples/notebooks.html
# 
| examples/models/alibaba_ack_deep_mnist/alibaba_cloud_ack_deep_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
import pandas as pd
# +
# # La Liga
# league = "laliga"
# abbr = "ES1"
# # Ligue 1
# league = "ligue-1"
# abbr = "FR1"
# Premier League
league = "premier-league"
abbr = "GB1"
r = requests.get(f"https://www.transfermarkt.co.uk/{league}/legionaereeinsaetze/wettbewerb/{abbr}/saison_id/2020/altersklasse/alle/option/spiele/plus/1",
headers= {"User-Agent":"Mozilla/5.0 (Windows NT 10.0; Win64; x64; rv:74.0) Gecko/20100101 Firefox/74.0"})
soup = BeautifulSoup(r.content, "html.parser")
# -
results = soup.find("table",class_="items")
df = pd.read_html(str(results))[0]
df.drop(columns=['wappen',"% minutes foreign players"],inplace=True)
df.columns=['Club', 'Players used', 'Non-foreigners played',
'Used foreign players', '% minutes non-foreigners',
'% minutes foreign players']
df = df[:20]
df = df.sort_values(by="% minutes foreign players", ascending=False)
df
# df.to_csv("laliga_foreign_player_minutes.csv", index=False)
# df.to_csv("ligue1_foreign_player_minutes.csv", index=False)
df.to_csv("premierleague_foreign_player_minutes.csv", index=False)
| foreign_player_minutes_scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import print_function
from collections import defaultdict
import sys
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
from Bio import PDB
# -
repository = PDB.PDBList()
parser = PDB.PDBParser()
repository.retrieve_pdb_file('1TUP', pdir='.')
p53_1tup = parser.get_structure('P 53', 'pdb1tup.ent')
# +
atom_cnt = defaultdict(int)
atom_chain = defaultdict(int)
atom_res_types = defaultdict(int)
for atom in p53_1tup.get_atoms():
my_residue = atom.parent
my_chain = my_residue.parent
atom_chain[my_chain.id] += 1
if my_residue.resname != 'HOH':
atom_cnt[atom.element] += 1
atom_res_types[my_residue.resname] += 1
print(dict(atom_res_types))
print(dict(atom_chain))
print(dict(atom_cnt))
# -
res_types = defaultdict(int)
res_per_chain = defaultdict(int)
for residue in p53_1tup.get_residues():
res_types[residue.resname] += 1
res_per_chain[residue.parent.id] +=1
print(dict(res_types))
print(dict(res_per_chain))
def get_bounds(my_atoms):
my_min = [sys.maxint] * 3
my_max = [-sys.maxint] * 3
for atom in my_atoms:
for i, coord in enumerate(atom.coord):
if coord < my_min[i]:
my_min[i] = coord
if coord > my_max[i]:
my_max[i] = coord
return my_min, my_max
chain_bounds = {}
for chain in p53_1tup.get_chains():
print(chain.id, get_bounds(chain.get_atoms()))
chain_bounds[chain.id] = get_bounds(chain.get_atoms())
print(get_bounds(p53_1tup.get_atoms()))
#matplotlib 3d plot
fig = plt.figure(figsize=(16, 9))
ax3d = fig.add_subplot(111, projection='3d')
ax_xy = fig.add_subplot(331)
ax_xy.set_title('X/Y')
ax_xz = fig.add_subplot(334)
ax_xz.set_title('X/Z')
ax_zy = fig.add_subplot(337)
ax_zy.set_title('Z/Y')
color = {'A': 'r', 'B': 'g', 'C': 'b', 'E': '0.5', 'F': '0.75'}
zx, zy, zz = [], [], []
for chain in p53_1tup.get_chains():
xs, ys, zs = [], [], []
for residue in chain.get_residues():
ref_atom = next(residue.get_iterator())
x, y, z = ref_atom.coord
if ref_atom.element == 'ZN':
zx.append(x)
zy.append(y)
zz.append(z)
continue
xs.append(x)
ys.append(y)
zs.append(z)
ax3d.scatter(xs, ys, zs, color=color[chain.id])
ax_xy.scatter(xs, ys, marker='.', color=color[chain.id])
ax_xz.scatter(xs, zs, marker='.', color=color[chain.id])
ax_zy.scatter(zs, ys, marker='.', color=color[chain.id])
ax3d.set_xlabel('X')
ax3d.set_ylabel('Y')
ax3d.set_zlabel('Z')
ax3d.scatter(zx, zy, zz, color='k', marker='v', s=300)
ax_xy.scatter(zx, zy, color='k', marker='v', s=80)
ax_xz.scatter(zx, zz, color='k', marker='v', s=80)
ax_zy.scatter(zz, zy, color='k', marker='v', s=80)
for ax in [ax_xy, ax_xz, ax_zy]:
ax.get_yaxis().set_visible(False)
ax.get_xaxis().set_visible(False)
| notebooks/06_Prot/Stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="jCIIG4eCyMrP"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn as sk
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
# + colab={} colab_type="code" id="0NkLKSbfyMrX"
# Генерируем уникальный seed
my_code = "Soloviev"
seed_limit = 2 ** 32
my_seed = int.from_bytes(my_code.encode(), "little") % seed_limit
np.random.seed(my_seed)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} colab_type="code" id="VAABYAScyMrc" outputId="71cd5b6d-4d97-4638-91ac-ae674e6bcdef"
# Формируем случайную нормально распределенную выборку sample
N = 10000
sample = np.random.normal(0, 1, N)
plt.hist(sample, bins=100)
plt.show()
# -
# Формируем массив целевых метока классов: 0 - если значение в sample меньше t и 1 - если больше
t = 0
target_labels = np.array([0 if i < t else 1 for i in sample])
plt.hist(target_labels, bins=100)
plt.show()
# + colab={} colab_type="code" id="lpiBPPw1yMr_"
# Используя данные заготовки (или, при желании, не используя),
# реализуйте функции для рассчета accuracy, precision, recall и F1
def confusion_matrix(target_labels, model_labels) :
tp = 0
tn = 0
fp = 0
fn = 0
for i in range(len(target_labels)) :
if target_labels[i] == 1 and model_labels[i] == 1 :
tp += 1
if target_labels[i] == 0 and model_labels[i] == 0 :
tn += 1
if target_labels[i] == 0 and model_labels[i] == 1 :
fp += 1
if target_labels[i] == 1 and model_labels[i] == 0 :
fn += 1
return tp, tn, fp, fn
def metrics_list(target_labels, model_labels):
metrics_result = []
metrics_result.append(sk.metrics.accuracy_score(target_labels, model_labels))
metrics_result.append(sk.metrics.precision_score(target_labels, model_labels))
metrics_result.append(sk.metrics.recall_score(target_labels, model_labels))
metrics_result.append(sk.metrics.f1_score(target_labels, model_labels))
return metrics_result
# +
# Первый эксперимент: t = 0, модель с вероятностью 50% возвращает 0 и 1
t = 0
target_labels = np.array([0 if i < t else 1 for i in sample])
model_labels = np.random.randint(2, size=N)
# Рассчитайте и выведите значения метрик accuracy, precision, recall и F1.
metrics_list(target_labels, model_labels)
# +
# Второй эксперимент: t = 0, модель с вероятностью 25% возвращает 0 и с 75% - 1
t = 0
target_labels = np.array([0 if i < t else 1 for i in sample])
labels = np.random.randint(4, size=N)
model_labels = np.array([0 if i == 0 else 1 for i in labels])
np.random.shuffle(model_labels)
# Рассчитайте и выведите значения метрик accuracy, precision, recall и F1.
metrics_list(target_labels, model_labels)
# +
# Проанализируйте, какие из метрик применимы в первом и втором экспериментах.
# +
# Третий эксперимент: t = 2, модель с вероятностью 50% возвращает 0 и 1
t = 2
target_labels = np.array([0 if i < t else 1 for i in sample])
model_labels = np.random.randint(2, size=N)
# Рассчитайте и выведите значения метрик accuracy, precision, recall и F1.
metrics_list(target_labels, model_labels)
# +
# Четвёртый эксперимент: t = 2, модель с вероятностью 100% возвращает 0
t = 2
target_labels = np.array([0 if i < t else 1 for i in sample])
model_labels = np.zeros(N)
# Рассчитайте и выведите значения метрик accuracy, precision, recall и F1.
metrics_list(target_labels, model_labels)
# +
# Проанализируйте, какие из метрик применимы в третьем и четвёртом экспериментах.
| 2020 Осенний семестр/Практическое задание 3/Соловьёв-задание 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + pycharm={"is_executing": true}
import sys
import os
from pathlib import Path
from glob import glob
# from datetime import timedelta
import datetime as dt
import pandas as pd
from fameio.scripts.convert_results import run as convert_results
from fameio.source.cli import Config
from fameio.source.time import FameTime, Constants
import math
import matplotlib.pyplot as plt
# %matplotlib notebook
# + pycharm={"is_executing": true}
CONFIG = {
Config.LOG_LEVEL: "info",
Config.LOG_FILE: None,
Config.AGENT_LIST: None,
Config.OUTPUT: 'FameResults_converted',
Config.SINGLE_AGENT_EXPORT: False
}
# + pycharm={"is_executing": true}
def process_file(filepath: str) -> pd.DataFrame:
"""Process single AMIRIS csv file"""
df = pd.read_csv(filepath, sep=';')
object_class = Path(filepath).stem
assert df.columns[1] == 'TimeStep'
assert df.columns[0] == 'AgentId'
# Convert times steps
df['TimeStamp'] = df['TimeStep'].apply(roundup)
df['TimeStamp'] = df['TimeStamp'].apply(convert_fame_time_step_to_datetime)
df['ObjectClass'] = object_class
return df.drop('TimeStep', axis=1).melt(['ObjectClass', 'AgentId', 'TimeStamp']).dropna()
# + pycharm={"is_executing": true}
def roundup(x):
return round(x / 100.0) * 100
# + pycharm={"is_executing": true}
def convert_fame_time_step_to_datetime(fame_time_steps: int) -> str:
"""Converts given `fame_time_steps` to corresponding real-world datetime string"""
years_since_start_time = math.floor(fame_time_steps / Constants.STEPS_PER_YEAR)
current_year = years_since_start_time + Constants.FIRST_YEAR
beginning_of_year = dt.datetime(year=current_year, month=1, day=1, hour=0, minute=0, second=0)
steps_in_current_year = (fame_time_steps - years_since_start_time * Constants.STEPS_PER_YEAR)
seconds_in_current_year = steps_in_current_year / Constants.STEPS_PER_SECOND
simulatedtime = beginning_of_year + dt.timedelta(seconds=seconds_in_current_year)
tiemporounded = simulatedtime.replace(second=0, microsecond=0, minute=0, hour=simulatedtime.hour) + dt.timedelta(
hours=simulatedtime.minute // 30)
#tiemporounded.strftime('%Y-%m-%dT%H:%M:%S')
return tiemporounded
# -
# Get input file from cmd line arguments
# + pycharm={"is_executing": true}
input_pb_file = "C:\\Users\\isanchezjimene\\Documents\\TraderesCode\\toolbox-amiris-emlab\\scripts\\AMIRIS.FameResult.pb"
#parent = os.path.basename(os.getcwd())
#complete = os.path.join(Path(os.getcwd()).parent, "data", input_pb_file)
# -
# Convert Proto Buffer file to csv's
# + pycharm={"is_executing": true}
convert_results(input_pb_file, CONFIG)
# Combine csv files into one data frame
csv_files = glob(f'{CONFIG[Config.OUTPUT]}/*.csv')
originaldata = pd.concat(map(process_file, csv_files))
# -
# Plot results
# + pycharm={"is_executing": true, "name": "#%%\n"}
data = originaldata
data.dtypes
data.info()
data.head(5)
# + pycharm={"is_executing": true}
data.head(5)
# + pycharm={"is_executing": true}
data.set_index('TimeStamp').resample('Y')
# -
data.resample('Y').agg(dict(ObjectClass='sum', variable='sum', AgentId='sum'))
# + pycharm={"is_executing": true}
grouped = data.groupby(["ObjectClass", "variable", "AgentId" ]).sum()
# + pycharm={"is_executing": true}
data.groupby(pd.Grouper(key='TimeStamp',freq='Y')).sum()
# + pycharm={"is_executing": true}
grouped.info()
# + pycharm={"is_executing": true}
grouped.head(5)
# + [markdown] pycharm={"name": "#%% md\n"}
# Write results
# + pycharm={"is_executing": true, "name": "#%%\n"}
grouped.to_csv('AMIRIS_combined.csv', index=True)
# -
| scripts/ignore/combine_AMIRIS_results_ingrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Synapse PySpark
# name: synapse_pyspark
# ---
# Copyright (c) Microsoft Corporation.
#
# Licensed under the MIT License.
#
# # DISCLAIMER
# By accessing this code, you acknowledge that the code is not designed, intended, or made available: (1) as a medical device(s); (2) for the diagnosis of disease or other conditions, or in the cure, mitigation, treatment or prevention of a disease or other conditions; or (3) as a substitute for professional medical advice, diagnosis, treatment, or judgment. Do not use this code to replace, substitute, or provide professional medical advice, diagnosis, treatment, or judgement. You are solely responsible for ensuring the regulatory, legal, and/or contractual compliance of any use of the code, including obtaining any authorizations or consents, and any solution you choose to build that incorporates this code in whole or in part.
#
# # Recommendations
# Load the test data and apply to the model
# # Library Imports
#
# +
import numpy as np
import pandas as pd
from pyspark.sql.types import *
data_lake_account_name = ""
file_system_name = "raw"
subscription_id = ""
resource_group = ""
workspace_name = ""
workspace_region = ""
experiment_name = "DiabetesPredictionExperiment"
autoMLRunId = ""
aks_target_name = ''
# -
# set transformed data schema
transformedSchema = StructType([StructField("race", StringType(), True),
StructField("gender", StringType(), True),
StructField("age", StringType(), True) ,
StructField("admission_type_id", StringType(), True),
StructField("discharge_disposition_id", StringType(), True),
StructField("admission_source_id", StringType(), True),
StructField("time_in_hospital", StringType(), True),
StructField("payer_code", StringType(), True),
StructField("num_lab_procedures", StringType(), True),
StructField("num_procedures", StringType(), True),
StructField("num_medications", StringType(), True),
StructField("number_outpatient", StringType(), True),
StructField("number_emergency", StringType(), True),
StructField("number_inpatient", StringType(), True),
StructField("number_diagnoses", StringType(), True),
StructField("max_glu_serum", StringType(), True),
StructField("A1Cresult", StringType(), True),
StructField("metformin", StringType(), True),
StructField("repaglinide", StringType(), True),
StructField("nateglinide", StringType(), True),
StructField("chlorpropamide", StringType(), True),
StructField("glimepiride", StringType(), True),
StructField("glipizide", StringType(), True),
StructField("glyburide", StringType(), True),
StructField("tolbutamide", StringType(), True),
StructField("pioglitazone", StringType(), True),
StructField("rosiglitazone", StringType(), True),
StructField("acarbose", StringType(), True),
StructField("miglitol", StringType(), True),
StructField("tolazamide", StringType(), True),
StructField("insulin", StringType(), True),
StructField("glyburide-metformin", StringType(), True),
StructField("metformin-rosiglitazone", StringType(), True),
StructField("change", StringType(), True),
StructField("diabetesMed", StringType(), True),
StructField("FirstName", StringType(), True),
StructField("LastName", StringType(), True),
StructField("Id", StringType(), True),
StructField("spec_InternalMedicine", BooleanType(), True),
StructField("spec_Emergency/Trauma", BooleanType(), True),
StructField("spec_Family/GeneralPractice", BooleanType(), True),
StructField("spec_Cardiology", BooleanType(), True),
StructField("spec_Surgery-General", BooleanType(), True),
StructField("diag_428", BooleanType(), True),
StructField("diag_250", BooleanType(), True),
StructField("diag_276", BooleanType(), True),
StructField("diag_414", BooleanType(), True),
StructField("diag_401", BooleanType(), True),
StructField("diag_427", BooleanType(), True),
StructField("diag_599", BooleanType(), True),
StructField("diag_496", BooleanType(), True),
StructField("diag_403", BooleanType(), True),
StructField("diag_486", BooleanType(), True),
StructField("is_readmitted", BooleanType(), True)
])
# # Load Data from Azure Data Lake
#
# +
from sklearn.model_selection import train_test_split
import pandas as pd
df_train = spark.read.format("csv").load(f"abfss://{file_system_name}@{data_lake_account_name}.dfs.core.windows.net/DatasetDiabetes/preparedtraindata/",header=True,schema=transformedSchema)
df_train = df_train.toPandas()
outcome_column = 'is_readmitted'
id_column = 'Id'
df_train = df_train.drop(id_column,axis=1)
#%% Split data for validation
X = df_train.drop(outcome_column, axis=1)
y = df_train[outcome_column]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=0)
# -
# # Connect to Azure Machine Learning Workspace, Experiment and Load Best Run
#
#
# +
#save the model to a local file
import azureml.core
from azureml.core import Workspace
ws = Workspace(workspace_name = workspace_name,
subscription_id = subscription_id,
resource_group = resource_group)
ws.write_config()
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl.run import AutoMLRun
from azureml.train.automl import AutoMLConfig
experiment = Experiment(workspace = ws, name = experiment_name)
previous_automl_run = AutoMLRun(experiment, autoMLRunId, outputs = None)
automl_run = previous_automl_run
best_run, fitted_model = automl_run.get_output()
from sklearn.externals import joblib
model_path = 'diabetesmodel'
joblib.dump(fitted_model, model_path)
# +
from azureml.interpret.scoring.scoring_explainer import TreeScoringExplainer, save
from interpret.ext.glassbox import LGBMExplainableModel
from azureml.interpret.mimic_wrapper import MimicWrapper
from azureml.train.automl.runtime.automl_explain_utilities import AutoMLExplainerSetupClass, automl_setup_model_explanations
automl_explainer_setup_obj = automl_setup_model_explanations(fitted_model, X=X_train,
X_test=X_test, y=y_train,
task='classification')
explainer = MimicWrapper(ws, automl_explainer_setup_obj.automl_estimator, LGBMExplainableModel,
init_dataset=automl_explainer_setup_obj.X_transform, run=best_run,
features=automl_explainer_setup_obj.engineered_feature_names,
feature_maps=[automl_explainer_setup_obj.feature_map],
classes=automl_explainer_setup_obj.classes)
#Initialize the ScoringExplainer
scoring_explainer = TreeScoringExplainer(explainer.explainer, feature_maps=[automl_explainer_setup_obj.feature_map])
#Pickle scoring explainer locally
save(scoring_explainer, exist_ok=True)
# -
# # Register the Model
#
# +
from azureml.core.model import Model
model_name = "diabetesmodel"
registered_model = Model.register(model_path = model_path, # this points to a local file
model_name = model_name, # name the model is registered as
tags = {'type': "classification"},
description = "Diabetes Classifier",
workspace = ws)
exp_model_name = "scoring_explainer.pkl"
exp_model_path = "scoring_explainer.pkl"
exp_registered_model = Model.register(model_path = exp_model_path, # this points to a local file
model_name = exp_model_name, # name the model is registered as
tags = {'type': "scoring explainer"},
description = "Diabetes Readmission Classifier Explainer",
workspace = ws)
# -
def create_additional_features(df):
to_drop = ['acetohexamide', 'troglitazone', 'examide', 'citoglipton',
'glipizide-metformin', 'glimepiride-pioglitazone',
'metformin-pioglitazone', 'weight', 'patient_nbr', 'encounter_id']
df.drop(to_drop, axis=1, inplace=True, errors = 'ignore')
df_transformed = df.replace('?', np.nan)
spec_counts_raw = {"specs": ['InternalMedicine', 'Emergency/Trauma', 'Family/GeneralPractice','Cardiology',
'Surgery-General'], "num patients": [14635, 7565, 7440, 5352, 3099]}
df_transformed['medical_specialty'] = df_transformed['medical_specialty'].replace(np.nan, "NaNSpec")
spec_counts = pd.DataFrame(data = spec_counts_raw)
spec_thresh = 5
for (index, row) in spec_counts.head(spec_thresh).iterrows():
spec = row['specs']
new_col = 'spec_' + str(spec)
df_transformed[new_col] = (df_transformed.medical_specialty == spec)
diag_counts_raw = {"icd9value": ['428', '250', '276', '414', '401', '427', '599', '496', '403', '486'],
'num patients w diag': [18101., 17861., 13816., 12895., 12371., 11757., 6824., 5990.,5693., 5455.]}
diag_counts = pd.DataFrame(diag_counts_raw, columns = [ 'icd9value', 'num patients w diag'])
diag_thresh = 10
for (index, row) in diag_counts.head(diag_thresh).iterrows():
icd9 = row['icd9value']
new_col = 'diag_' + str(icd9)
df_transformed[new_col] = (df_transformed.diag_1 == icd9)|(df_transformed.diag_2 == icd9)|(df_transformed.diag_3 == icd9)
df_transformed = df_transformed.reset_index(drop=True)
df_transformed2 = pd.DataFrame(df_transformed, copy=True) #preserve df_transformed so I can rerun this step
df_transformed2['age'] = df_transformed2.age.str.extract('(\d+)-\d+')
to_drop = ['acetohexamide', 'troglitazone', 'examide', 'citoglipton',
'glipizide-metformin', 'glimepiride-pioglitazone',
'metformin-pioglitazone', 'weight', 'medical_specialty', 'diag_2',
'diag_1', 'diag_3', 'patient_nbr', 'encounter_id']
df_transformed2.drop(to_drop, axis=1, inplace=True,errors = 'ignore')
df_transformed2 = df_transformed2.reset_index(drop=True)
df_transformed2['readmitted'].value_counts()
df = pd.DataFrame(df_transformed2)
#Imputing with outlying value since we are focusing on tree based methods
df = df.fillna(-9999)
df = df.reset_index(drop=True)
df.dtypes
return df
# +
import pandas as pd
df_test = spark.read.format("csv").load(f"abfss://{file_system_name}@{data_lake_account_name}.dfs.<EMAIL>.windows.net/DatasetDiabetes/preparedtestdata/",header=True,multiLine=True)
df_test = df_test.toPandas()
outcome_column = 'readmitted'
df_test = df_test.drop(outcome_column,axis=1)
df_test = df_test.head(2)
id_column = 'Id'
df_test = df_test.drop(id_column,axis=1)
# +
scoring_script = """
import json
import pickle
import numpy as np
import pandas as pd
import azureml.train.automl
from sklearn.externals import joblib
from azureml.core.model import Model
from azureml.train.automl.runtime.automl_explain_utilities import automl_setup_model_explanations
def create_additional_features(df):
to_drop = ['acetohexamide', 'troglitazone', 'examide', 'citoglipton',
'glipizide-metformin', 'glimepiride-pioglitazone',
'metformin-pioglitazone', 'weight', 'patient_nbr', 'encounter_id']
df.drop(to_drop, axis=1, inplace=True, errors = 'ignore')
df_transformed = df.replace('?', np.nan)
spec_counts_raw = {"specs": ['InternalMedicine', 'Emergency/Trauma', 'Family/GeneralPractice','Cardiology',
'Surgery-General'], "num patients": [14635, 7565, 7440, 5352, 3099]}
df_transformed['medical_specialty'] = df_transformed['medical_specialty'].replace(np.nan, "NaNSpec")
spec_counts = pd.DataFrame(data = spec_counts_raw)
spec_thresh = 5
for (index, row) in spec_counts.head(spec_thresh).iterrows():
spec = row['specs']
new_col = 'spec_' + str(spec)
df_transformed[new_col] = (df_transformed.medical_specialty == spec)
diag_counts_raw = {"icd9value": ['428', '250', '276', '414', '401', '427', '599', '496', '403', '486'],
'num patients w diag': [18101., 17861., 13816., 12895., 12371., 11757., 6824., 5990.,5693., 5455.]}
diag_counts = pd.DataFrame(diag_counts_raw, columns = [ 'icd9value', 'num patients w diag'])
diag_thresh = 10
for (index, row) in diag_counts.head(diag_thresh).iterrows():
icd9 = row['icd9value']
new_col = 'diag_' + str(icd9)
df_transformed[new_col] = (df_transformed.diag_1 == icd9)|(df_transformed.diag_2 == icd9)|(df_transformed.diag_3 == icd9)
df_transformed = df_transformed.reset_index(drop=True)
df_transformed2 = pd.DataFrame(df_transformed, copy=True) #preserve df_transformed so I can rerun this step
df_transformed2['age'] = df_transformed2.age.str.extract('(\d+)-\d+')
to_drop = ['acetohexamide', 'troglitazone', 'examide', 'citoglipton',
'glipizide-metformin', 'glimepiride-pioglitazone',
'metformin-pioglitazone', 'weight', 'medical_specialty', 'diag_2',
'diag_1', 'diag_3', 'patient_nbr', 'encounter_id']
df_transformed2.drop(to_drop, axis=1, inplace=True,errors = 'ignore')
df_transformed2 = df_transformed2.reset_index(drop=True)
# df_transformed2['readmitted'].value_counts()
df = pd.DataFrame(df_transformed2)
#Imputing with outlying value since we are focusing on tree based methods
df = df.fillna(-9999)
df = df.reset_index(drop=True)
df.dtypes
return df
def init():
global model
global scoring_explainer
# This name is model.id of model that we want to deploy deserialize the model file back
model_path = Model.get_model_path(model_name = 'diabetesmodel')
model = joblib.load(model_path)
scoring_explainer_path = Model.get_model_path(model_name = 'scoring_explainer.pkl')
scoring_explainer = joblib.load(scoring_explainer_path)
def run(input_json):
try:
data_df = pd.read_json(input_json, orient='records').head(1)
data_df = create_additional_features(data_df)
stacked_data = pd.DataFrame(data_df.stack().reset_index())
stacked_data.columns = ['Ind','Column','Value']
stacked_data = stacked_data[['Column','Value']]
# Get the predictions...
# prediction = model.predict(data_df)
prediction = pd.DataFrame(model.predict_proba(data_df),columns=model.y_transformer.inverse_transform(model.classes_)).T.iloc[0,0]
prediction = np.round(prediction * 100,2)
automl_explainer_setup_obj = automl_setup_model_explanations(model,X_test=data_df, task='classification')
raw_local_importance_values = scoring_explainer.explain(automl_explainer_setup_obj.X_test_transform, get_raw=True)
stacked_data['raw_imp'] = raw_local_importance_values[0]
stacked_data = stacked_data.sort_values('raw_imp',ascending = False).head(10)
#stacked_data['raw_imp'] = stacked_data['raw_imp'] * 100
stacked_data = stacked_data.round(2)
except Exception as e:
prediction = np.array([str(e)])
stacked_data = pd.DataFrame([str(e)])
return {'predictions': prediction.tolist(),
'raw_local_importance_values': stacked_data.values.tolist()}
"""
exec(scoring_script)
with open("scoring_script.py", "w") as file:
file.write(scoring_script)
scoring_script_file_name = 'scoring_script.py'
# -
json_test_data = df_test.head(1).to_json(orient='records')
print(json_test_data)
init()
run(json_test_data)
# +
# obtain conda dependencies from the automl run and save the file locally
from azureml.core import Environment
from azureml.core.environment import CondaDependencies
conda_dep = CondaDependencies()
environment_config_file = 'diabetes_conda_env.yml'
best_run.download_file('outputs/conda_env_v_1_0_0.yml', environment_config_file)
with open('diabetes_conda_env.yml', 'r') as f:
print(f.read())
# create the environment based on the saved conda dependencies file
myenv = Environment.from_conda_specification(name="diabetesenv", file_path=environment_config_file)
conda_dep.add_pip_package("shap==0.35.0")
conda_dep.add_pip_package("azureml-train-automl-runtime==1.32.0")
conda_dep.add_pip_package("inference-schema")
conda_dep.add_pip_package("azureml-interpret==1.32.0")
conda_dep.add_pip_package("azureml-defaults==1.32.0")
conda_dep.add_conda_package("numpy>=1.16.0,<1.19.0")
conda_dep.add_conda_package("pandas==0.25.1")
conda_dep.add_conda_package("scikit-learn==0.22.1")
conda_dep.add_conda_package("py-xgboost<=0.90")
conda_dep.add_conda_package("fbprophet==0.5")
conda_dep.add_conda_package("holidays==0.9.11")
conda_dep.add_conda_package("psutil>=5.2.2,<6.0.0")
myenv.python.conda_dependencies=conda_dep
myenv.register(workspace=ws)
# -
# # Deploy Model to AKS Cluster
#
# +
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AksWebservice
from azureml.core.webservice import Webservice
# Configure and deploy the web service to Azure Container Instances
inference_config = InferenceConfig(environment=myenv, entry_script=scoring_script_file_name)
aks_config = AksWebservice.deploy_configuration(cpu_cores = 1, memory_gb= 2, tags = { 'type' : 'automl-classification'}, description='AutoML Diabetes Readmission Classifier Service')
aks_service_name = 'diabetes-readmission-service-aks'
aks_target = AksCompute(ws,aks_target_name)
aks_service = Model.deploy(ws, aks_service_name, [exp_registered_model,registered_model], inference_config, aks_config, aks_target)
# -
aks_service.wait_for_deployment(show_output = True)
print(aks_service.state)
json_test_data = df_test.head(1).to_json()
aks_service.run(json_test_data)
| Analytics_Deployment/Notebooks/02_deploy_AKS_diabetes_readmission_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jZwZr4SRjJl1"
# # Core 2020
# # Taller Introducción al aprendizaje por refuerzo profundo para juegos en atari
# ## Libreta de entornos en python y métodos de programación dinámica.
# *Por <NAME>*
#
#
# + id="J6BZADOEjkS9"
import matplotlib.pyplot as plt
import numpy as np
from abc import ABC
# + [markdown] id="M0EOd8EsK-oL"
# ### Entornos y clases base
# Las siguientes son ejemplos de clases base para comenzar a crear un entorno de interés en el lenguaje de programacion de Python. Hacer estos no es estrictamente necesario, pero sirven como base para los métodos que todos los entornos necesitan tener para que las funciones escritas en base a estas clases funcionen sin problema. Algunos de estos estan basados en una interfaz popular de OpenAI llamada gym.
#
# Hacer entornos se puede hacer con una mentalidad desconectada a usar algoritmos de aprendizaje por refuerzo, el objetivo es representar lo más fiel el fénomeno a estudiar. Como segunda meta, es adaptar este modelo o simulación del fenomeno con estas clases para generar una interfaz (también conocidas como API). Esto con la finalidad que diferentes algoritmos, personas o entornos diseñados alrededor de la API sufran pocos o ningún cambio para funcionar entre si.
# + id="aQni7aNaSKZg"
class actionSpace(ABC):
"""
Discrete Action Space class.
Stores all the posible actions for an environment and can generate
samples with the method sample().
Parameters
----------
n: int
Number of total actions. These are consider to be sequential.
minValue: int
Default 0. The minimum value that the action can take. It's
the lower inclusive of the action space intervals
[minValue, minValue + n)
"""
def __init__(self, n:int, minValue:int=0):
assert n > 0, "Number of actions must be greater than 0"
self.n = n
self.mV = minValue
@property
def shape(self):
return self.n
def __iter__(self):
self.actions = []
self.i = 0
for i in range(self.n):
self.actions += [self.mV + i]
return self
def __next__(self):
if self.i == self.n:
raise StopIteration
nx = self.actions[self.i]
self.i += 1
return nx
def sample(self):
"""
Returns a random sample with an uniform distribution of the
actions.
"""
return np.random.randint(self.mV, self.mV + self.n)
class Environment(ABC):
"""
Environment base class.
"""
def step(self, action):
"""
Executes the action, updates the environment, calculates de reward and observation output.
Returns
-------
observation, reward, done
"""
raise NotImplemented
def reset(self):
"""
Restart the initial state of the environment, in a deterministic or stochastic manner
Returns
------
obeservation
"""
raise NotImplemented
def getObservation(self):
"""
Calculates and returns the observation of the actual state of the environment.
"""
raise NotImplemented
def calculateReward(self, state):
"""
Calculate with the actual mode the reward from the last observation made in the environment
Returns
-------
reward
"""
raise NotImplemented
@property
def actionSpace(self):
"""
Returns the ActionShape class designed of the environment.
"""
raise NotImplemented
@property
def stateSpace(self):
"""
Returns a list or generator of all the states availables.
"""
raise NotImplemented
def transProb(self, state, action):
"""
Returns the probabilities and states of the transitions from the
state and action given.
"""
raise NotImplemented
def isTerminal(self, state):
"""
Returns the bool that expreses if the actual state is a terminal one
or not.
"""
raise NotImplemented
class Policy(ABC):
"""
Policy base class.
"""
def getAction(self, state):
"""
Calculates and returns the corresponding action for the state given.
"""
raise NotImplemented
def update(self, state, action):
"""
Update the action per state manner of the policy
"""
raise NotImplemented
# + [markdown] id="saXCWTeBNEvg"
# ### Grid World; Primer entorno sencillo.
# Un "mundo reticula", es un caso básico a analizar y encontrarle analogos a un entorno real podria resultar sencillo. Aunque se describa como sencillo, nos permite observar fenomenos interesantes en el contexto de aprendizaje por refuerzo.
#
# Preguntas interesantes acerca de este entorno serían:
# - ¿Que representación del estado seria suficiente?
# - De crecer una columna o fila al entorno, ¿incrementa mucha nuestra complejidad computacional?
# - ¿Qué utilidad podría tener un modelo así?
# + id="YMKltSddjtRV"
class gridWorld(Environment):
"""
Little and simple environment for a deterministic grid world.
Parameters
----------
width: int
First dimension of the grid
height: int
Second dimension of the grid
initPos: tuple of int
Initial position of the agent.
goal: tuple of int
Position of the first goal to create the grid.
One can add more goals later if required.
movement: str
Default 8C. Refer to step method.
horizon: int
Default 10**6. Number of steps to run the environment before it
terminates.
"""
# All gfx related
EMPTYC = (255, 255, 255)
OBST = 2
OBSTC = (128, 64, 64)
VORTEX = 3
VORTEXC = (230, 0, 10)
GOAL = 1
GOALC = (0, 200, 20)
AGENTC = (230, 150, 240)
POLICYC = (0, 1, 0.5)
CELLSIZE = 4
GRAPHSCALE = 1.2
VORTEXD = [[False, True, True, False],
[True, False, False, True],
[True, False, False, True],
[False, True, True, False]]
AGENTD = [[False, True, True, False],
[True, True, True, True],
[True, True, True, True],
[False, True, True, False]]
GOALD = [[True, False, True, False],
[False, True, False, True],
[True, False, True, False],
[False, True, False, True]]
# Action meaning related
actions = [(-1,-1),(-1, 0),(-1,1),
(0, -1),(0, 0),(0, 1),
(1, -1),(1, 0),(1, 1)]
actions4C = [1,3,4,5,7]
def __init__(self, width:int, height:int, initPos:tuple, goal:tuple, movement:str = "4C", horizon:int = 10**6):
# Grid Related
self.grid = np.zeros((width, height), dtype=np.uint8)
self._w = width
self._h = height
self.obstacles = []
self.vortex = []
self.goal = [goal]
self.steps = 0
self.gameOver = False
self.horizon = horizon
# Agent related
self.movMode = movement
self.validateTuple(initPos)
self.initX, self.initY = initPos
self.posX, self.posY = initPos
self.__actionSpace = actionSpace(9 if movement == "8C" else 5, 1)
# Graphics related
self.frame = np.zeros((width * self.CELLSIZE, height * self.CELLSIZE, 3), dtype=np.uint8)
# Initialize the grid
self.reset()
def validateTuple(self, T:tuple):
assert len(T) == 2, "Tuple needs to have 2 items for x and y"
if (T[0] < 0) or (T[1] < 0):
raise ValueError("Values of the tuple must be non-negative")
if (T[0] >= self._w) or (T[1] >= self._h):
raise ValueError("Value of the tuple need to be in the interval x[0, {}), y[0, {})".format(self._w, self._h))
return True
def addVortex(self, *vortex):
"""
Add a vortex on the grid
Parameters
---------
vortex: tuple of int
A tuple of integers that cointains the position in which one
desire to put a new vortex.
"""
for v in vortex:
self.validateTuple(v)
self.vortex += [v]
def addObstacles(self, *obstacles):
"""
Add an obstacle on the grid
Parameters
---------
obstacles: tuple of int
A tuple of integers that cointains the position in which one
desire to put a new obstacle.
"""
for o in obstacles:
self.validateTuple(o)
self.obstacles += [o]
def addGoals(self, *goals):
"""
Add a goal on the grid
Parameters
---------
goal: tuple of int
A tuple of integers that cointains the position in which one
desire to put an additional goal.
"""
for g in goals:
self.validateTuple(g)
self.goals += [g]
def reset(self, initialPos = None):
self.grid[:,:] = 0
for o in self.obstacles:
self.grid[o] = self.OBST
for v in self.vortex:
self.grid[v] = self.VORTEX
for g in self.goal:
self.grid[g] = self.GOAL
if initialPos is None:
self.posX = self.initX
self.posY = self.initY
else:
self.validateTuple(initialPos)
self.posX, self.posY = initialPos
self.steps = 0
self.gameOver = False
self.lastObs = self.getObservation()
self.lastReward = 0
self.lastAction = 5
return self.lastObs
def step(self, action:int = 5):
"""
Excecute a step on the environment.
The actions on the grid that the agent can take on mode 8C are
integers from 1 to 9.
[1 2 3]
|4 5 6|
[7 8 9]
5 being the neutral action or "Do nothing"
In mode 4C, the action space is reduced to just move in a cross
pattern with integers from 1 to 5
[- 1 -]
|2 3 4|
[- 5 -]
3 being the "do nothing" action.
Parameters
----------
action: int
Returns
-------
observation , reward, done
"""
# If the environment has reached a terminal state
if self.gameOver:
return self.lastObs, 0, True
# Select the action from the corresponding transition probabilities
randomSelect = np.random.uniform(0,1)
probs, states = self.transProb(self.lastObs, action)
lastP = 0
for p, s in zip(probs, states):
if randomSelect <= (p + lastP):
self.posX, self.posY = s
break
else:
lastP += p
self.steps += 1
# Check the horizon
if self.steps > self.horizon:
self.gameOver = True
# Get new state and reward
self.lastObs = self.getObservation(copy = False)
self.lastReward = self.calculateReward(self.lastObs)
return self.lastObs, self.lastReward, self.gameOver
def validateAction(self, state, action:int):
if self.movMode == "8C":
assert (action > 0) and (action < 10), "Action must be an integer between 1 and 9"
dx, dy = self.actions[action - 1]
elif self.movMode == "4C":
assert (action > 0) and (action < 6), "Action must be an integer between 1 and 5"
dx, dy = self.actions[self.actions4C[action - 1]]
self.lastAction = action
posX, posY = state["agent"]
# Tentative new position
posX += dx
posY += dy
# Checking the movements be inside the grid
if (posX < 0) or (posX >= self._w) or (posY < 0) or (posY >= self._h):
# Is not inside the grid, this does nothing
return state["agent"]
# Checking if the movement gets it to an obstacle
elif self.grid[posX, posY] == self.OBST:
# Returns the same position as before
return state["agent"]
else:
# No obstacle the new position is returned
return posX, posY
def calculateReward(self, state):
# For each movement
reward = -1
cellAgent = self.grid[state["agent"]]
if cellAgent == self.VORTEX:
# The agent has enter a vortex.
reward += - 14
self.gameOver = True
elif cellAgent == self.GOAL:
reward += 11
self.gameOver = True
return reward
def getObservation(self, copy:bool = True):
if copy:
return {"agent":(self.posX, self.posY),
"grid": np.copy(self.grid)}
else:
return {"agent":(self.posX, self.posY),
"grid": self.grid}
def render(self, values=None, policy=None):
# Suboptimal but simple to understand graphics for the environment
fig = plt.figure(figsize=(self._w * self.GRAPHSCALE, self._h * self.GRAPHSCALE), clear = True)
self.frame[:,:] = self.EMPTYC
for i in range(self._w):
for j in range(self._h):
cell = self.grid[i,j]
ni, nj = self.CELLSIZE * i, self.CELLSIZE * j
f = self.frame[ni:ni+self.CELLSIZE,nj:nj+self.CELLSIZE]
if cell == self.OBST:
f[:,:,:] = self.OBSTC
elif cell == self.VORTEX:
f[self.VORTEXD,:] = self.VORTEXC
elif cell == self.GOAL:
f[self.GOALD,:] = self.GOALC
if values is not None:
plt.text(nj + 1.5, ni + 1.5, str(np.round(values[i,j], 2)),
horizontalalignment='center',
verticalalignment='center',)
if policy is not None and (cell == 0):
action = policy.getAction((i,j)) - 1
if self.movMode == "4C":
action = self.actions4C[action]
dx, dy = self.actions[action]
plt.arrow(nj + 1.5, ni + 1.5, 1.5 * dy, 1.5 * dx, width=0.2, color=self.POLICYC)
ni, nj = self.posX * self.CELLSIZE, self.posY * self.CELLSIZE
f = self.frame[ni:ni+self.CELLSIZE,nj:nj+self.CELLSIZE,:]
f[self.AGENTD,:] = self.AGENTC
plt.title("GridWorld {}x{} Action {} Reward {}".format(self._w, self._h,
self.lastAction,
self.lastReward))
plt.imshow(self.frame)
plt.axis("off")
def updateGrid(self):
pass
@property
def actionSpace(self):
return self.__actionSpace
@property
def stateSpace(self):
states = []
for i in range(self._w):
for j in range(self._h):
if not self.grid[i,j] == self.OBST:
states += [(i,j)]
return states
def transProb(self, state, action):
# Deterministic Environment
state = self.validateAction(state, action)
return [1], [state]
def isTerminal(self, state):
if isinstance(state, dict):
cellAgent = self.grid[state["agent"]]
else:
cellAgent = self.grid[state]
if cellAgent == self.VORTEX or cellAgent == self.GOAL:
return True
else:
return False
@property
def shape(self):
return self.grid.shape
# + [markdown] id="Ele4ydUGPS2R"
# ### Controlador del agente
# Los agentes en RL en realidad son un conjunto de varios elementos disintos para lograr tomar las decisiones con el entorno. Para el entorno anterior, se generan dos clases nuevas de politicas sencillas. Una que es siempre aleatoria con probabilidad uniforme, y otra que se inicializa aleatoriamente pero se comporta de manera deterministica.
#
#
# + id="1VEkupipmGmO"
class uniformRandomPolicy(Policy):
def __init__(self, env:Environment):
self.pi = env.actionSpace
self.env = env
def getAction(self, state):
return self.pi.sample()
def update(self, state, action):
pass # Do nothing
class gridPolicy(Policy):
def __init__(self, env:Environment):
self.pi = np.zeros(env.shape, dtype=np.uint8)
# or could be a dict() as well
self.env = env
self.randomInit()
def randomInit(self):
for state in self.env.stateSpace:
self.pi[state] = self.env.actionSpace.sample()
def update(self, state, action):
if isinstance(state, dict):
state = state["agent"]
self.pi[state] = action
def getAction(self, state):
if isinstance(state, dict):
state = state["agent"]
return self.pi[state]
# + [markdown] id="Iim7We7gP8Pv"
# ### A comenzar a jugar con el entorno GridWorld!
# Las celdas ejecutadas son del código muestra como el autor las diseño, sus configuraciones no tienen justificación en particulas más que mostrar el funcionamiento.
#
# Los resultados de los códigos a escribir están basados en esos, pero aun asi cambia todo lo que haga contento tu corazón. Resultados a otras reticulas pueden ser comparados en el classroom.
# + id="puTYAxjqK04u"
env = gridWorld(6,6,(0,0),(4,4),movement="8C") # Iniciar el entorno
randomPolicy = uniformRandomPolicy(env) # Crear una politica
env.addObstacles((4,3),(2,2),(0,1),(0,2)) # Añadimos algunos obstaculos
env.addVortex((4,2)) # Añadimos otros obstaculos que se comportan como un torbellino
obs = env.reset() # Reiniciamos el entorno para que vuelva a cargar todo lo anterior
# + colab={"base_uri": "https://localhost:8080/", "height": 438} id="rK-XxCaMNchZ" executionInfo={"status": "ok", "timestamp": 1606711075265, "user_tz": 360, "elapsed": 1899, "user": {"displayName": "<NAME>\u00f3pez", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="e38cbd2c-ce1d-48ee-9206-c02e8851ef1c"
obs, reward, done = env.step(randomPolicy.getAction(obs))
env.render()
# + [markdown] id="n7mVzRwlQxmT"
# ## Evaluación de la politica
# Todos pseudo códigos fueron extraidos del libro de Suton Y Barto.
# #### Pseudocódigo de Iterative Policy Evaluation
# Input $\pi$, the policy to be evaluated
# Algorithm parameter: a small threshold $\theta > 0$ determining accuracy of estimation
# Initialize $V(s)$, for al $s\in \mathit{S^+}$, arbitrarilly except that $V(terminal) = 0$
#
# Loop:
# - $\Delta \leftarrow 0$
# - Loop for each $s\in \mathit{S}$:
# - $v \leftarrow V(s)$
# - $V(s) \leftarrow \Sigma_a \pi(a|s)\Sigma_{s',r}p(s',r | s, a)[r+\gamma V(s')]$
# - $\Delta \leftarrow \max(\Delta, |v-V(s)|)$
#
# until $\Delta < \theta$
# + id="kltZJwR2eWRf"
def policyEvaluation(env:Environment, policy:Policy, k:int, thres:float = 0.01, gamma:float = 0.99):
assert (gamma >= 0) and (gamma <= 1), "Gamma discount factor must be in the interval [0,1]"
assert k > 0, "k needs to be am integer greater or equal to 1 iteration."
# Initialization
V = np.zeros(env.shape, dtype=np.float32)
states = env.stateSpace
actions = env.actionSpace
diff = 0
# Policy evaluation
# Iteration as stop condition
for _ in range(k):
# Do a copy of V_t to be V_{t - 1}
Vpass = np.copy(V)
# Iterate the states of the environment
for state in states:
# Check if the state is terminal
if env.isTerminal(state):
V[state] = 0
continue # With this the next code is not not executed
# Get the action from the policy
action = policy.getAction(state)
# Get the probabilities and next states corresponding to the
# actual state and action
probs, nextStates = env.transProb({"agent":state}, action)
sum = 0
for p, s in zip(probs, nextStates):
r = env.calculateReward({"agent":s})
sum += p * (r + gamma * Vpass[s])
# Update the value function given pi
V[state] = sum
diff = max(diff, Vpass[state] - sum)
# First condition
if diff < thres:
return V
return V
# + colab={"base_uri": "https://localhost:8080/", "height": 438} id="Wyw9j0S3RhWG" executionInfo={"status": "ok", "timestamp": 1606711075820, "user_tz": 360, "elapsed": 2440, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/<KEY>g=s64", "userId": "01862305152975853555"}} outputId="f8c6ecd9-cbca-4078-8b36-dfca4c5b832d"
policy = gridPolicy(env)
vp = policyEvaluation(env, policy, 10, 0.1)
env.render(vp, policy)
# + [markdown] id="n7iWXz_sQ0hC"
# ## Mejora de la politica con Iteración de politica
# 1. Initialization $V(s)\in \mathbb{R} $ and $ \pi(s)\in \mathit{A}(s)$
# 2. Policy Evaluation.
# 3. Policy improvemente
# - $\text{policy-stable} \leftarrow \text{true}$
# - For each $s\in \mathit(S)$:
# - $old-action \leftarrow \pi(s)$
# - $\pi(s)\leftarrow \arg \max_a \Sigma_{s',r}p(s', r | s,a)[r + \gamma V(s')] $
# - If $\text{old-action} \neq \pi(s)$, $\text{policy-stable} \leftarrow false$
#
# if $\text{policy-stable}$, then stop and return $V$, else return to step 2.
# + id="GjhHP8jtvAmA"
def policyIteration(env:Environment, policy:Policy, k_eva:int, k_improvement:int, thres:float = 0.01, gamma:float = 0.99,):
assert (gamma >= 0) and (gamma <= 1), "Gamma discount factor must be in the interval [0,1]"
assert k_improvement > 0, "k needs to be am integer greater or equal to 1 iteration."
# Initialization
states = env.stateSpace
actions = env.actionSpace
# Iteration as alternate stop condition
for improvement in range(k_improvement):
# Policy evaluation
V = policyEvaluation(env, policy, k_eva, thres = thres, gamma = gamma)
# Policy improvement
policyStable = True
# Iterate the states
for state in states:
oldAction = policy.getAction(state)
maxSum = - np.inf
maxAction = None
for action in actions:
probs, nextStates = env.transProb({"agent":state}, action)
sum = 0
for p, s in zip(probs, nextStates):
r = env.calculateReward({"agent":s})
sum += p * (r + gamma * V[s])
if sum > maxSum:
maxSum = sum
maxAction = action
# Update the policy with the new action which gives
# more reward
policy.update(state, maxAction)
if maxAction != oldAction:
policyStable = False
# Stop condition check
if policyStable == True:
print("Policy stable afer {} iterations".format(improvement))
return V
print("Stopped after all the iterations")
return V
# + colab={"base_uri": "https://localhost:8080/", "height": 456} id="_yhRq48Rz4oC" executionInfo={"status": "ok", "timestamp": 1606711076218, "user_tz": 360, "elapsed": 2822, "user": {"displayName": "<NAME>\u00f3pez", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="da4f1f0d-32f9-4bed-89f4-8e639487ea8b"
vp = policyIteration(env, policy, 10, 50, 0.99)
env.render(vp, policy)
# + [markdown] id="2GGoDDUfezLB"
# ## Value Iteration
# ### Pseudocode
# Algorithm parameter: a small threshold $\theta > 0$ determining accuracy of estimation.
#
# Initialize $V(s)$, for all $a\in \mathit{S^+}$, arbitrarily except that $V(terminal)=0$
#
# Loop:
# - $\Delta \leftarrow 0$
# - Loop for each $s\in \mathit{S}$:
# - $v \leftarrow V(s)$
# - $V(S) \leftarrow \max_a \Sigma_{s',r}p(s', r | s,a)[r + \gamma V(s')]$
# - $\Delta \leftarrow \max(\Delta, |v-V(s)|)$
#
# until $\Delta < 0$
#
# Output a deterministic policy, $\pi \approx \pi_*$, such that
#
# $\pi(s) = \arg \max_a \Sigma_{s',r} p(s',r| s, a)[r+\gamma V(s')]$
# + id="4QaQVrfHNtww"
def valueIteration(env:Environment, k:int, thres:float = 0.01, gamma:float = 0.99):
assert (gamma >= 0) and (gamma <= 1), "Gamma discount factor must be in the interval [0,1]"
assert k > 0, "k needs to be am integer greater or equal to 1 iteration."
actions = env.actionSpace
states = env.stateSpace
V = np.zeros(env.shape, dtype=np.float32)
diff = 0
# Start the iterations
for iteration in range(k):
# Do a copy of V_t as is now V_{t-1}
Vpass = np.copy(V)
# For each state of all available
for state in states:
maxSum = - np.inf
maxAction = None
stateDict = {"agent": state}
# If the state is terminal assign V(s) = 0
if env.isTerminal(stateDict):
V[state] = 0
continue # With this the next code is not not executed
# For each action available
for action in actions:
# Get from the mode the probabilities and their
# corresponding next states given the actual state
probs, nextStates = env.transProb(stateDict, action)
sum = 0
# For each pair of the probability and its state
for p, s in zip(probs, nextStates):
r = env.calculateReward({"agent":s})
sum += p * (r + gamma * Vpass[s])
if sum > maxSum:
maxSum = sum
maxAction = action
V[state] = maxSum
diff = max(diff, Vpass[state] - maxSum)
# First stop condition
if diff < thres:
return V
return V
# + colab={"base_uri": "https://localhost:8080/", "height": 438} id="gPanwM2Pdhsr" executionInfo={"status": "ok", "timestamp": 1606711076897, "user_tz": 360, "elapsed": 3486, "user": {"displayName": "<NAME>\u00f3pez", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="135bb1a1-80cc-4d22-be2d-2bdbb0fdd4d0"
vi = valueIteration(env, 100, 0.99)
env.render(vi)
# + [markdown] id="o5PtOGvHl9z4"
# ### Una nueva variante, un mundo con probabilidades
# + id="gJMur9_yObDN"
class stochasticGridWorld(gridWorld):
"""
A modification to the GridWorld to add moving vortex with random directions.
This movements follow the same type of movement as the agent.
Parameters
----------
width: int
First dimension of the grid
height: int
Second dimension of the grid
initPos: tuple of int
Initial position of the agent.
goal: tuple of int
Position of the first goal to create the grid.
One can add more goals later if required.
movement: str
Default 8C. Refer to step method.
horizon: int
Default 10**6. Number of steps to run the environment before it
terminates.
"""
def __init__(self, width:int, height:int, initPos:tuple, goal:tuple, movement:str = "4C", horizon:int = 10**6):
super().__init__(width, height, initPos, goal, movement, horizon)
self.vortexProb = []
def addVortex(self, *vortex):
"""
Add a stochastic atraction vortex on the grid
Parameters
---------
vortex: tuple
A tuple with the form (x,y,p). x and y are integers
which cointain the initial position to put a new vortex.
While p is a float in [0,1) that the vortex will attract the
agent to it even if it's not leading to it.
"""
for v in vortex:
assert len(v) == 3, "The tuple must cointain two integers as position and third float number to express the probability"
self.validateTuple(v[:2])
self.vortex += [v[:2]]
p = v[2]
assert (p >= 0) and (p < 1), "The probability; third item on the tuple needs to be between 0 and 1"
self.vortexProb += [v[2]]
def transProb(self, state, action):
# Local function
def nearby(pos:tuple, vortex:tuple, diag:bool):
d1 = abs(pos[0] - vortex[0])
d2 = abs(pos[1] - vortex[1])
if (d1 <= 1) and (d2 <= 1) and (diag == True):
return True
elif ((d1 == 1 and d2 == 0) or (d1 == 0 and d2 == 1)) and (diag == False):
return True
else:
return False
# Checking state type
if isinstance(state, dict):
agent = state["agent"]
else:
agent = state
# Init
states = []
probs = []
# Check if the agent is nearby 1 cell of the effect of the vortex
for v, p in zip(self.vortex, self.vortexProb):
if nearby(agent, v, True if self.movMode == "8C" else False):
states += [v]
probs += [p]
# Add the action state
states += [self.validateAction(state, action)]
n = len(states) - 1
if n == 0:
probs = [1]
else:
probs += [n - sum(probs)]
# Normalize the probabilities
probs = np.array(probs, dtype=np.float32)
probs = probs / n
return probs, states
# + id="tXRC6hMwYCPO"
envs = stochasticGridWorld(6,6,(0,0), (5,5), movement="8C")
envs.addVortex((1,2,0.9), (2,2,0.7), (5,4, 0.5))
envs.addObstacles((4,3),(2,2),(0,1),(0,2))
# + colab={"base_uri": "https://localhost:8080/", "height": 438} id="BDH-bsOaYMHc" executionInfo={"status": "ok", "timestamp": 1606711076906, "user_tz": 360, "elapsed": 3478, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="97612933-3e0e-4e6a-d413-22adcb547dd3"
envs.reset()
envs.render()
# + colab={"base_uri": "https://localhost:8080/"} id="QFk51-sitMP-" executionInfo={"status": "ok", "timestamp": 1606711076907, "user_tz": 360, "elapsed": 3474, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="d0f0909c-be83-49ed-afd5-c6df9b65cfcb"
envs.transProb({"agent":(1,1)},5)
# + colab={"base_uri": "https://localhost:8080/", "height": 456} id="aq0tZW_FxHjT" executionInfo={"status": "ok", "timestamp": 1606711077622, "user_tz": 360, "elapsed": 4180, "user": {"displayName": "<NAME>\u00f3pez", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="8c20baa4-8f57-475e-f6dd-ecc34be0b8f0"
spolicy = gridPolicy(envs)
vp = policyIteration(envs, spolicy, 20, 20)
envs.render(vp, spolicy)
# + colab={"base_uri": "https://localhost:8080/", "height": 438} id="CAaEdwFyxkLW" executionInfo={"status": "ok", "timestamp": 1606711078704, "user_tz": 360, "elapsed": 5252, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiSsX1x6xJBOqtfATqdKNcynw2HNLV_3DUkdYyJq9g=s64", "userId": "01862305152975853555"}} outputId="2a676bab-186a-4270-ee96-d47aa59457f9"
vi = valueIteration(envs, 100)
envs.render(vi)
| TallerCore2020 DRL Complete-s1v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
from unidecode import unidecode
#options
pd.set_option('display.float_format', lambda x: '%.2f' % x)
# -
# # Trenes - Transporte de pasajeros
pasajeros = pd.read_csv("https://servicios.transporte.gob.ar/gobierno_abierto/descargar.php?t=trenes&d=pasajeros", sep=";", encoding="UTF-8")
pasajeros
# ## DATA CLEANING
#DEJAMOS TODO EN MAYUS - NOMBRES DE COLUMNAS
pasajeros.columns = pasajeros.columns.str.upper()
#QUITAMOS TILDES Y DEJAMOS TODO EN MAYUS - VALORES
pasajeros[["LINEA", "ESTACION"]]= pasajeros[["LINEA", "ESTACION"]].applymap(lambda x: str.upper(unidecode(x)))
#DAMOS FORMATO A LA FECHA
pasajeros["FECHA"] = pd.to_datetime(pasajeros["MES"])
#CORREGIMOS UN PROBLEMA DE ESPACIO
pasajeros = pasajeros.replace({"BELGRANOSUR":"BELGRANO SUR", "BELGRANONORTE":"BELGRANO NORTE", "SANMARTIN": "SAN MARTIN"})
#ASUMIMOS EL MES COMO LA REFERENCIA DE ORDEN TEMPORAL
pasajeros = pasajeros.sort_values("FECHA").reset_index(drop=True)
# **Nota:** Parece que fueron agregando datos sin tomar en cuenta la referencia temporal, por eso la lo reordenamos
pasajeros = pasajeros.drop(columns="MES")
pasajeros = pasajeros.rename(columns={"CANTIDAD":"TOTAL"})
pasajeros.to_csv("./data/07-TRENES-PASAJEROS.csv", index_label="INDEX")
pasajeros.dtypes
# # INSPECCION VISUAL
pasajeros_mes_linea = pasajeros.groupby(["FECHA", "LINEA"]).aggregate("sum").unstack()
# +
#PARAMETRO
fig, ax = plt.subplots(figsize=(40,15))
#AGRUPAMOS
pasajeros_mes_linea.plot(ax=ax)
#ETIQUETAS
ax.set_title("PASAJEROS - TRENES ARGENTINOS", size=40)
plt.xlabel("FECHA", size=15)
plt.ylabel("CANTIDAD DE VIAJES", size=15)
plt.show();
# -
# Pese que habría que preprocesar los datos para emitir un resultado riguroso, del gráfico podemos observar varias cosas:
#
# 1. Hay una marcada diferencia en el comprtamiento de todas las redes en el año `2018`.
# 2. La línea de `BELGRANO` tiene un comportamiento bastante diferente y las otras se parecen entre sí, tomando un lenguaje de física moderna y ondas, diria que "están en fase".
# 3. La línea `GENERAL ROCA` a transportado significativamente más personas desde sus inicios.
# 4. Exeptuando las lineas de `GENERAL ROCA` las otras parecen oscilar al rededor de un valor medio.
# 5. ¿Estuvieron sin servicio las líneas de `BELGRANO`?
| 7-SERVICIOS-Trenes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.applications.inception_v3 import InceptionV3
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.models import Model, load_model
from keras.layers import *
from keras import backend as K
from keras import optimizers, callbacks
import numpy as np
import pandas as pd
import cv2, h5py
# ## Model (Inception V3)
# +
batch_size = 128
train_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
'train_aug/',
target_size=(299, 299),
batch_size=batch_size,
class_mode='categorical')
validation_datagen = ImageDataGenerator(rescale=1./255)
validation_generator = validation_datagen.flow_from_directory(
'data2/validation',
target_size=(299, 299),
batch_size=batch_size,
class_mode='categorical')
# +
base_model = InceptionV3(weights='imagenet', include_top=False)
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dense(512, activation='relu')(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.4)(x)
predictions = Dense(128, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=predictions)
for layer in base_model.layers:
layer.trainable = False
# compile the model (should be done *after* setting layers to non-trainable)
adadelta = optimizers.Adadelta(lr=1.0, rho=0.95, epsilon=None, decay=0.001)
model.compile(loss='categorical_crossentropy',
optimizer=adadelta,
metrics=['acc'])
# -
# ### Training
tensorboard = callbacks.TensorBoard(log_dir='./logs', histogram_freq=0, batch_size=16, write_grads=True , write_graph=True)
model_checkpoints = callbacks.ModelCheckpoint("inception-{val_loss:.3f}.h5", monitor='val_loss', verbose=0, save_best_only=True, save_weights_only=False, mode='auto', period=0)
# !rm -R logs
# !ls
print("Training Progress:")
model_log = model.fit_generator(train_generator, validation_data=validation_generator,
epochs=5,
callbacks=[tensorboard, model_checkpoints])
pd.DataFrame(model_log.history).to_csv("inception-history.csv")
# ### Fine Tuning
"""# let's visualize layer names and layer indices to see how many layers
# we should freeze:
for i, layer in enumerate(base_model.layers):
print(i, layer.name)
# we chose to train the top 2 inception blocks, i.e. we will freeze
# the first 249 layers and unfreeze the rest:
for layer in model.layers[:249]:
layer.trainable = False
for layer in model.layers[249:]:
layer.trainable = True
# we need to recompile the model for these modifications to take effect
# we use SGD with a low learning rate
from keras.optimizers import SGD
model.compile(optimizer=SGD(lr=0.0001, momentum=0.9), loss='categorical_crossentropy')
# we train our model again (this time fine-tuning the top 2 inception blocks
# alongside the top Dense layers
model.fit_generator(...)"""
# ## Evaluation
# +
from keras.models import load_model
from sklearn.metrics import classification_report, confusion_matrix
import matplotlib.pyplot as plt
import numpy as np
# %config InlineBackend.figure_format = 'retina'
import itertools, pickle
classes = [str(x) for x in range(0,129)]
# -
model_test = load_model('best_weights.h5')
Y_test = np.argmax(y_val, axis=1) # Convert one-hot to index
y_pred = model_test.predict(x_val)
y_pred_class = np.argmax(y_pred,axis=1)
cnf_matrix = confusion_matrix(Y_test, y_pred_class)
print(classification_report(Y_test, y_pred_class, target_names=classes))
| furniture/model-inception.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
data=[]
for i in range(10):
data.append(2**i)
data
# -
plt.plot(data)
fig=plt.figure()
ax1=fig.add_subplot(2,2,1)
ax2=fig.add_subplot(2,2,2)
ax3=fig.add_subplot(2,2,3)
fig
_=ax1.hist(np.random.randn(100),bins=20,color='K',alpha=0.3)
ax2.scatter(np.arange(30),np.arange(30)+3*np.random.randn(30))
ax2
x=np.arange(30)
y=np.arange(30)+3*np.random.rand(30)
fig1=plt.scatter(x,y)
x
y
data=np.random.rand(50)
fig2=plt.hist(data)
data
data=np.random.randn(10)
plt.plot(data,'k*--')
data
data.cumsum()
for N in range(1,100):
flag=1
for i in range(2,N//2+1):
if(N%i==0):
flag=0
break
if(flag==1):
print(N)
fig=plt.figure()
ax=fig.add_subplot(1,1,1,)
ax.plot(np.random.randn(1000).cumsum())
ax.set_xlabel("Stages")
ax.set_title("My first matplotlib plot")
fig
# # Object
class student:
id=1
name='Khushbu'
def __init__(self,d_id,d_name):
self.id=d_id
self.name=d_name
def display(self):
print(self.id)
print(self.name)
s=student(50,'Parixit')
s.display()
# +
from sklearn import tree
features=[[100,0],[80,0],[120,0],[125,1],[140,1],[160,1]]
labels=[0,0,0,1,1,1]
# -
classifier=tree.DecisionTreeClassifier()
classifier.fit(features,labels)
predict=classifier.predict([[90,0]])
predict
from numpy.random import randn
fig=plt.figure();
ax=fig.add_subplot(1,1,1)
ax.plot(randn(1000).cumsum(),'k',label='One')
ax.plot(randn(1000).cumsum(),'k--',label='Two')
ax.plot(randn(1000).cumsum(),'k.',label='Three')
ax.legend(loc='best')
# # Woking with Pandas
import pandas as pd
data1=pd.read_csv("D:\\Machine learning\\BVM\\Day-1_16092018\\DATASET\\tsv_data.txt",sep='\t')
data1
data2=pd.read_table('D:\\Machine learning\\BVM\\Day-1_16092018\\DATASET\\tsv_data.txt')
data2
data3=pd.read_csv('D:\\Machine learning\\BVM\\Day-1_16092018\\DATASET\\Data.csv')
data3
# +
import numpy as np
df=pd.DataFrame({'a':[4,5,6],
'b':[7,8,9],
'c':[10,11,12]},index=[1,2,6])
# -
df
df2=pd.DataFrame(np.random.randint(low=0,high=10,size=(5,5)),columns=['a','b','c','d','e'])
df2
np.random.randint(low=0,high=10,size=(5,5))
df.to_excel('D:\\Machine learning\\BVM\\Day-1_16092018\\DATASET\\sqltoxcel.xlsx')
df.to_clipboard()
# # Pivot Library
temp=df2.pivot(index="a",columns='b',values='c')
temp
temp.corr()
temp.describe()
| BVM-Day 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
import numpy as np
from torch.utils.tensorboard import SummaryWriter
from torch.utils.data import SubsetRandomSampler
import random
import math
import copy
# +
class Arguments():
def __init__(self):
self.batch_size = 64
self.test_batch_size = 64
self.epochs = 20
self.best_lr_list = []
self.no_cuda = False
self.seed = 1
self.log_interval = 10
self.save_model = False
self.gamma = 0.1
self.alpha_max = 0.1
self.epsilon = 8
self.clip_threshold = 0.01
self.split = 600
args = Arguments()
use_cuda = not args.no_cuda and torch.cuda.is_available()
torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
kwargs = {'num_workers': 1, 'pin_memory': True} if use_cuda else {}
# +
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('~/data', train=True, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.batch_size,
pin_memory = True
)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('~/data', train=False, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.test_batch_size,
pin_memory = True
)
# +
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 34, 5, 1)
self.conv2 = nn.Conv2d(34, 64, 5, 1)
self.fc1 = nn.Linear(20*20*64, 512)
self.fc2 = nn.Linear(512, 10)
self.drop = nn.Dropout(p=0.3)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = x.view(-1, 20*20*64)
x = self.drop(x)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=1)
# model is not exactully the same as the paper since it did not mention the unit of fc
# -
np.random.laplace(0, 1.5/1.3, 1)
def load_grad(temp, model):
for net1,net2 in zip(model.named_parameters(),temp.named_parameters()):
net2[1].grad = net1[1].grad.clone()
def noisy_max (loss_list, p_nmax, clip_threshold):
neg_loss_array = np.array([-x for x in loss_list])
noise = np.random.laplace(0, clip_threshold/p_nmax, len(neg_loss_array))
noisy_loss = neg_loss_array + noise
best_loss_index = np.argmax(noisy_loss)
return best_loss_index
# +
def add_grad_noise(model, noise):
for i, param in enumerate(model.parameters()):
param.grad.add_(noise[i])
def sub_grad_noise(model, noise):
for i, param in enumerate(model.parameters()):
param.grad.sub_(noise[i])
# -
def create_grad_Gaussian_noise(model, device, p_ng, clip_threshold, batch_size):
noise = []
# remembe that torch.normal(mean, std) use std
for param in model.parameters():
noise.append(torch.normal(0, clip_threshold/math.sqrt(2 * p_ng), param.grad.size(), device=device)/batch_size)
return noise
# +
model = Net()
r = np.random.randint(920)
sampler = SubsetRandomSampler(list(range(r*args.batch_size, (r+5)*args.batch_size)))
step_size_loader = torch.utils.data.DataLoader(
datasets.MNIST('~/data', train=True, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
sampler=sampler,
batch_size=args.batch_size,
pin_memory = True
)
for i,j in step_size_loader:
o = model(i)
loss = F.nll_loss(o, j)
loss.backward()
noise = create_noise(model, device, args.epsilon, args.clip_threshold, args.batch_size)
print(noise[0])
add_grad_noise(model, noise)
# -
def best_step_size_model(args, model, device, train_loader, p_ng):
r = np.random.randint(920)
sampler = SubsetRandomSampler(list(range(r*args.batch_size, (r+5)*args.batch_size)))
step_size_loader = torch.utils.data.DataLoader(
datasets.MNIST('~/data', train=True, download=True, transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
sampler=sampler,
batch_size=args.batch_size,
pin_memory = True
)
best_loss = math.inf
best_lr = 0
best_model = Net().to(device)
if not args.best_lr_list:
args.alpha_max = min(args.alpha_max, 0.1)
elif len(args.best_lr_list) % 10 == 0:
args.alpha_max = (1+args.gamma) * max(args.best_lr_list)
del args.best_lr_list[:]
#while lr_index == 0, means choose the noise add on gradient again.
noise = create_grad_Gaussian_noise(model, device, p_ng, args.clip_threshold, args.batch_size)
index = 0
args.epsilon -= p_ng
if args.epsilon < 0:
return model, p_ng
while index == 0:
temp_loss_list = []
temp_model_list = []
temp_lr_list = []
add_grad_noise(model, noise)
for i in np.linspace(0, args.alpha_max, 21):
temp = Net().to(device)
temp_loss = 0
temp.load_state_dict(model.state_dict())
#load_state_dict will not copy the grad, so you need to copy it here.
load_grad(temp, model)
temp_optimizer = optim.SGD(temp.parameters(), lr=i)
temp_optimizer.step()
#optimizer will be new every time, so if you have state in optimizer, it will need load state from the old optimzer.
for (data, target) in step_size_loader:
data,target = data.to(device), target.to(device)
output = model(data)
temp_loss += F.nll_loss(output, target).item()
temp_loss_list.append(temp_loss)
temp_model_list.append(temp)
temp_lr_list.append(i)
#choose the best lr with noisy max
index = noisy_max(temp_loss_list, math.sqrt(2*p_nmax), args.clip_threshold)
args.epsilon -= p_nmax
if args.epsilon < 0:
return model, p_ng
# if index == 0, means we need to add the noise again and cost more epsilon
if index == 0:
#delete the original noise and add new noise
sub_grad_noise(model, noise)
# create new noise, and also sub the epsilon of new noise
p_ng = (1+args.gamma) * p_ng
noise = create_grad_Gaussian_noise(model, device, p_ng, args.clip_threshold, args.batch_size)
args.epsilon -= (args.gamma * p_ng)
if args.epsilon < 0:
break
else :
best_model.load_state_dict(temp_model_list[index].state_dict())
best_loss = temp_loss_list[index]
best_lr = temp_lr_list[index]
args.best_lr_list.append(best_lr)
# print("best learning rate:", best_lr)
# print("best loss:", best_loss)
return best_model, p_ng
def train(args, device, model, train_loader, epoch, p_ng):
#init the p_nmax and p_ng
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data,target = data.to(device), target.to(device)
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
# Chose the best step size(learning rate)
batch_best_model, p_ng = best_step_size_model(args, model, device, train_loader, p_ng)
if args.epsilon < 0:
break
model.load_state_dict(batch_best_model.state_dict())
model.zero_grad()
#remember to zero_grad or the grad will accumlate and the model will explode
if batch_idx % args.log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}\talpha_max: {:.6f}\tepsilon: {:.2f}'.format(
epoch, batch_idx * args.batch_size, len(train_loader.dataset) ,
100. * batch_idx * args.batch_size / len(train_loader.dataset), loss.item(), args.alpha_max, args.epsilon))
return model, p_ng
def test(args, device, model, test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader)*(args.batch_size)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\tepsilon: {:.2f}\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / (len(test_loader.dataset)), args.epsilon))
# +
# #%%time
model = Net().to(device)
args.best_lr_list = []
args.alpha_max = 0.1
args.epsilon = 8
p_ng, p_nmax = args.epsilon / (2 * args.split), args.epsilon / (2 * args.split) # 30 is split number, change if we need to
epoch = 0
# for epoch in range(1, args.epochs + 1):
while args.epsilon > 0:
epoch += 1
epoch_best_model, p_ng = train(args, device, model, train_loader , epoch, p_ng)
model.load_state_dict(epoch_best_model.state_dict())
test(args, device, model, test_loader)
if (args.save_model):
torch.save(model.state_dict(), "mnist_cnn.pt")
# -
| DP_FL_recreate/C_DPAGD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Simulation Parameters
# =====================
#
# Manage parameters for creating simulated power spectra.
#
# +
# Import simulation functions for creating spectra
from fooof.sim.gen import gen_power_spectrum, gen_group_power_spectra
# Import simulation utilities for managing parameters
from fooof.sim.params import param_sampler, param_iter, param_jitter, Stepper
# Import plotting functions to visualize spectra
from fooof.plts.spectra import plot_spectra
# -
# Simulation Parameters
# ---------------------
#
# One of the useful things about using simulated data is being able to compare results
# to ground truth values - but in order to do that, one needs to keep track of the
# simulation parameters themselves.
#
# To do so, there is the :obj:`~.SimParams` object to manage
# and keep track of simulation parameters.
#
# For example, when you simulate power spectra, the parameters for each spectrum are stored
# in a :obj:`~.SimParams` object, and then these objects are collected and returned.
#
# SimParams objects are named tuples with the following fields:
#
# - ``aperiodic_params``
# - ``periodic_params``
# - ``nlv``
#
#
#
# Set up settings for simulating a group of power spectra
n_spectra = 2
freq_range = [3, 40]
ap_params = [[0.5, 1], [1, 1.5]]
pe_params = [[10, 0.4, 1], [10, 0.2, 1, 22, 0.1, 3]]
nlv = 0.02
# Simulate a group of power spectra
freqs, powers, sim_params = gen_group_power_spectra(n_spectra, freq_range, ap_params,
pe_params, nlv, return_params=True)
# Print out the SimParams objects that track the parameters used to create power spectra
for sim_param in sim_params:
print(sim_param)
# You can also use a SimParams object to regenerate a particular power spectrum
cur_params = sim_params[0]
freqs, powers = gen_power_spectrum(freq_range, *cur_params)
# Managing Parameters
# -------------------
#
# There are also helper functions for managing and selecting parameters for
# simulating groups of power spectra.
#
# These functions include:
#
# - :func:`~.param_sampler` which can be used to sample parameters from possible options
# - :func:`~.param_iter` which can be used to iterate across parameter ranges
# - :func:`~.param_jitter` which can be used to add some 'jitter' to simulation parameters
#
#
#
# param_sampler
# ~~~~~~~~~~~~~
#
# The :func:`~.param_sampler` function takes a list of parameter options and
# randomly selects from the parameters to create each power spectrum. You can also optionally
# specify the probabilities with which to sample from the options.
#
#
#
# +
# Create a sampler to choose from two options for aperiodic parameters
ap_opts = param_sampler([[1, 1.25], [1, 1]])
# Create sampler to choose from two options for periodic parameters, and specify probabilities
pe_opts = param_sampler([[10, 0.5, 1], [[10, 0.5, 1], [20, 0.25, 2]]],
probs=[0.75, 0.25])
# -
# Generate some power spectra, using param_sampler
freqs, powers = gen_group_power_spectra(10, freq_range, ap_opts, pe_opts)
# Plot some of the spectra that were generated
plot_spectra(freqs, powers[0:4, :], log_powers=True)
# param_iter
# ~~~~~~~~~~
#
# The :func:`~.param_iter` function can be used to create iterators that
# can 'step' across a range of parameter values to be simulated.
#
# The :class:`~.Stepper` object needs to be used in conjunction with
# :func:`~.param_iter`, as it specifies the values to be iterated across.
#
#
#
# +
# Set the aperiodic parameters to be stable
ap_params = [1, 1]
# Use a stepper object to define the range of values to step across
# Stepper is defined with `start, stop, step`
# Here we'll define a step across alpha center frequency values
cf_steps = Stepper(8, 12, 1)
# We can use use param_iter, with our Stepper object, to create the full peak params
# The other parameter values will be held constant as we step across CF values
pe_params = param_iter([cf_steps, 0.4, 1])
# -
# Generate some power spectra, using param_iter
freqs, powers = gen_group_power_spectra(len(cf_steps), freq_range, ap_params, pe_params)
# Plot the generated spectra
plot_spectra(freqs, powers, log_freqs=True, log_powers=True)
# param_jitter
# ~~~~~~~~~~~~
#
# The :func:`~.param_jitter` function can be used to create iterators that
# apply some 'jitter' to the defined parameter values.
#
#
#
# +
# Define default aperiodic values, with some jitter
# The first input is the default values, the second the scale of the jitter
# You can set zero for any value that should not be jittered
ap_params = param_jitter([1, 1], [0.0, 0.15])
# Define the peak parameters, to be stable, with an alpha and a beta
pe_params = [10, 0.2, 1, 22, 0.1, 3]
# -
# Generate some power spectra, using param_jitter
freqs, powers = gen_group_power_spectra(5, freq_range, ap_params, pe_params)
# Plot the generated spectra
plot_spectra(freqs, powers, log_freqs=True, log_powers=True)
# We can see that in the generated spectra above, there is some jitter
# to the simulated aperiodic exponent values.
#
#
#
| doc/auto_examples/sims/plot_sim_params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hideCode=true hidePrompt=true
# <font size = "5"> **Chapter 4: [Spectroscopy](Ch4-Spectroscopy.ipynb)** </font>
#
#
# <hr style="height:1px;border-top:4px solid #FF8200" />
#
# # Edge Onset
#
# part of
#
# <font size = "5"> **[Analysis of Transmission Electron Microscope Data](_Analysis_of_Transmission_Electron_Microscope_Data.ipynb)**</font>
#
#
# by <NAME>, 2020
#
# Microscopy Facilities<br>
# Joint Institute of Advanced Materials<br>
# The University of Tennessee, Knoxville
#
# Model based analysis and quantification of data acquired with transmission electron microscopes
#
# ## Goal
#
# There are goals that we want to accomplish in this notebook.
#
# We want to determine which edges are present in a spectrum, that means ideally we want to know which element and which symmetry the excitation of this edge is associated with.
#
# Also, we want to determine as precisely as possible where the edge start. The onset of the edge gives us some indication into the oxidation state, charge transfer of an element in a compound and/or bandgap changes. The change of this edge onset in a compound from the edge onset of a single atom is called ``chemical shift``. Please look at the [chemical shift notebook](CH4-Chemical_Shift.ipynb) for more information.
#
# ## Relevant packages
# ### Install the newest version of sidpy, and pyTEMlib
# This notebook will only run with ``version 0.0.3`` or higher.
# +
import sys
from IPython.lib.deepreload import reload as dreload
try:
import sidpy
except ModuleNotFoundError:
# !pip3 install sidpy
if sidpy.__version__ < '0.0.3':
# !{sys.executable} -m pip install --upgrade sidpy
dreload(sidpy)
try:
import pyNSID
except ModuleNotFoundError:
# !{sys.executable} -m pip install --upgrade pyNSID
try:
import pyTEMlib
except ModuleNotFoundError:
# !{sys.executable} -m pip install --upgrade pyTEMlib
if pyTEMlib.__version__ < '0.2020.10.3':
# !{sys.executable} -m pip install --upgrade pyTEMlib
dreload(pyTEMlib)
# + [markdown] hideCode=false hidePrompt=false
# ### Import the packages needed
# + hideCode=true hidePrompt=false
# %pylab --no-import-all notebook
# %gui qt
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.insert(0,'../pyTEMlib/')
import pyTEMlib
import pyTEMlib.file_tools as ft
import pyTEMlib.eels_tools as eels # EELS methods
import pyTEMlib.interactive_eels as ieels # Dialogs for EELS input and quantification
import scipy
# For archiving reasons it is a good idea to print the version numbers out at this point
print('pyTEM version: ',pyTEMlib.__version__)
__notebook__ = 'analyse_core_loss'
__notebook_version__ = '2020_10_06'
# -
# ## Definition of Edge Onset
#
# If we consider an edge as more or less a sawtooth like feature, then any convolution with a resolution function (for simplicity we use a Gaussian). Change the resolution and watch the edge onset.
#
# Only the inflection point of the ionization edge stays at the same energy. This is why it makes sense to define the inflection point as the edge onset. The ``second derivative at an inflection point is zero`` (or infinite, a case that can be ignored for EELS) and so the ``second derivative is used to define the onset of an ionization edge``.
#
# The behaviour of a saw-tooth like edge is different from a peak, where the start or inflection point changes with width of smearing, while the peak position remains unchanged. I added a delta-like feature (one channel is higher)
# before the onset of the sawtooth-like edge.
# +
# Input
resolution = 15. #in eV
######################################
# calculate h in channels.
h = int(resolution/0.25)
# make energy scale
energy_scale = np.arange(1024)*.25+200
# make spectrum with powerlaw background
A = 1e10
r = 3.
spectrum = A* np.power(energy_scale,-r)
# add edge
spectrum[500:] = spectrum[500:]*2.4
spectrum[200] = spectrum[200]*10
# plot
plt.figure()
plt.plot(energy_scale[h:-h], spectrum[h:-h], linewidth=2, label='original simulated spectrum')
plt.plot(energy_scale[h:-h], scipy.ndimage.gaussian_filter(spectrum,h)[h:-h], linewidth=2,
label=f'broadened spectrum with resolution {resolution:.1f} eV')
plt.scatter(energy_scale[500], spectrum[500]*.7, color='red')
plt.ylim(0,spectrum[0]*1.05)
plt.legend();
# -
# ## Load and plot a spectrum
#
# Let's look at a real spectrum, the higher the signal background ratio the better, but that is true for any spectrum.
#
# As an example, we load the spectrum **1EELS Acquire (high-loss).dm3** from the *example data* folder.
#
# Please see [Loading an EELS Spectrum](LoadEELS.ipynb) for details on storage and plotting.
#
# First a dialog to select a file will apear.
#
# Then the spectrum plot and ``Spectrum Info`` dialog will appear, in which we set the experimental parameters.
#
# Please use the ``Set Energy Scale`` button to change the energy scale. When pressed a new dialog and a cursor will appear in which one is able to set the energy scale based on known features in the spectrum.
#
# +
try:
current_dataset.h5_dataset.file.close()
except:
pass
Qt_app = ft.get_qt_app()
current_dataset = ft.open_file()
current_channel = current_dataset.h5_dataset.parent
eels.set_previous_quantification(current_dataset)
# US 200 does not set acceleration voltage correctly.
# comment out next line for other microscopes
current_dataset.metadata['experiment']['acceleration_voltage'] = 200000
info = ieels.InfoDialog(current_dataset)
# + [markdown] hideCode=true hidePrompt=true
# ## Content
#
# The second derivative of an ionization edge is used as the definition of its onset.
#
# We will use as an approximation of an second derivative, the ``final difference`` of 2$^{\rm nd}$ order.
#
# -
# ## Finite Difference
#
# ### First Derivative
# The derivative of a function $f$ at a point $x$ is defined by the limit of a function.
#
# $$ f'(x) = \lim_{h\to0} \frac{f(x+h) - f(x)}{h} \approx \frac{f(x+h) - f(x)}{h} $$
#
# In the finite difference, we approximate this limit by a small integer, so that the derivative of a discrete list of values can be derived.
# +
h = 3
# We smooth the spectrum first a little
f_x = scipy.ndimage.gaussian_filter(current_dataset,h)
spec_dim = ft.get_dimensions_by_type('SPECTRAL', current_dataset)[0]
energy_scale = spec_dim[1].values
f_x_plus_h = np.roll(f_x,-h)
first_derivative = (f_x_plus_h - f_x) / h
first_derivative[:h] = 0
first_derivative[-h:] = 0
plt.figure()
plt.plot(energy_scale, f_x/20 , label='spectrum')
plt.plot(energy_scale,first_derivative, label='1$^{\rm st}$ derivative')
# -
# ### Second Derivative
# Again we use the finite differene, but now of order 2 to approximate the 2$^{\rm nd}$ derivative.
#
# $$ f''(x) \approx \frac{\delta_h^2[f](x)}{h^2} = \frac{ \frac{f(x+h) - f(x)}{h} - \frac{f(x) - f(x-h)}{h} }{h} = \frac{f(x+h) - 2 f(x) + f(x-h)}{h^{2}} . $$
#
# +
h = 3
# We smooth the spectrum first a little
f_x = scipy.ndimage.gaussian_filter(current_dataset,h)
spec_dim = ft.get_dimensions_by_type('SPECTRAL', current_dataset)[0]
energy_scale = spec_dim[1].values
f_x_plus_h = np.roll(f_x,-h)
f_x_minus_h = np.roll(f_x,+h)
second_derivative = (f_x_plus_h - 2*f_x + f_x_minus_h)/ h**2
second_derivative[:3] = 0
second_derivative[-3:] = 0
plt.figure()
plt.plot(energy_scale, f_x/10/h**2, label='spectrum')
#plt.plot(energy_scale, first_dif, label='first derivative')
plt.plot(energy_scale, -second_derivative+1000, label='second derivative')
plt.axhline(0, color='gray')
plt.legend();
# -
# ## Edge Detection
#
# ### Second Derivative and Edge Onset.
# The edge onset is defined as a zero of a second derivative. Actually, to be a correct onset, the 2$^{\rm nd}$ derivative has to go through zero with a negative slope. So, we neew first a maximum and then a minimum.
#
# First we need to locate the peaks in the second derivative, that are higher than the noise level. For that we determine the noise level at the start and end of the spectrum and approximate it linearly in between. Any maximum higher than the noise level is considered significant. We do the same for the minima.
#
# We define the noise level as a the product of a constant called ``sensitivity`` and the standard deviation in an energy window. Change the sensitivity around to see the effect (we start with 2.5 as a good first try)
# +
def second_derivative(dataset):
dim = ft.get_dimensions_by_type('spectral', dataset)
energy_scale = np.array(dim[0][1])
if dataset.data_type.name == 'SPECTRAL_IMAGE':
spectrum = dataset.view.get_spectrum()
else:
spectrum = np.array(dataset)
spec = scipy.ndimage.gaussian_filter(spectrum, 3)
dispersion = ft.get_slope(energy_scale)
second_dif = np.roll(spec, -3) - 2 * spec + np.roll(spec, +3)
second_dif[:3] = 0
second_dif[-3:] = 0
# find if there is a strong edge at high energy_scale
noise_level = 2. * np.std(second_dif[3:50])
[indices, peaks] = scipy.signal.find_peaks(second_dif, noise_level)
width = 50 / dispersion
if width < 50:
width = 50
start_end_noise = int(len(energy_scale) - width)
for index in indices[::-1]:
if index > start_end_noise:
start_end_noise = index - 70
noise_level_start = sensitivity * np.std(second_dif[3:50])
noise_level_end = sensitivity * np.std(second_dif[start_end_noise: start_end_noise + 50])
slope = (noise_level_end - noise_level_start) / (len(energy_scale) - 400)
noise_level = noise_level_start + np.arange(len(energy_scale)) * slope
return second_dif, noise_level
second_dif, noise_level = second_derivative(current_dataset)
plt.figure()
plt.plot(energy_scale, current_dataset/ 10, label='spectrum')
#plt.plot(energy_scale, first_dif, label='first derivative')
plt.plot(energy_scale, second_dif, label='second derivative')
plt.axhline(0, color='gray')
plt.plot(energy_scale, noise_level, color='gray', linewidth=2, label='noise level')
plt.plot(energy_scale, -noise_level, color='gray', linewidth=2)
# +
sensitivity = 2.5
import scipy
dim = ft.get_dimensions_by_type('spectral', current_dataset)
energy_scale = np.array(dim[0][1])
second_dif, noise_level = second_derivative(current_dataset)
[indices, peaks] = scipy.signal.find_peaks(second_dif,noise_level)
peaks['peak_positions']=energy_scale[indices]
peaks['peak_indices']=indices
edge_energies = [energy_scale[50]]
edge_indices = []
[indices, _] = scipy.signal.find_peaks(-second_dif,noise_level)
minima = energy_scale[indices]
plt.figure()
plt.plot(energy_scale, spec/ 10, label='spectrum')
#plt.plot(energy_scale, first_dif, label='first derivative')
plt.plot(energy_scale, second_dif, label='second derivative')
plt.axhline(0, color='gray')
plt.plot(energy_scale, noise_level, color='gray', linewidth=2, label='noise level')
plt.plot(energy_scale, -noise_level, color='gray', linewidth=2)
plt.scatter(peaks['peak_positions'], peaks['peak_heights'])
plt.scatter(energy_scale[indices], second_dif[indices], color='red')
plt.legend();
# -
# ### Determine Edge
#
# Now we can sort through the maxima and make sure a minimum is right behind it, but none at nearby lower energies.
#
# For the edge onset, we just make a linear interpolation between maximum and minimum, to determine the zero of this second derivative.
#
# +
edge_energies = [energy_scale[50]]
edge_indices = []
for peak_number in range(len(peaks['peak_positions'])):
position = peaks['peak_positions'][peak_number]
if position - edge_energies[-1]> 20:
impossible = minima[minima < position]
impossible = impossible[impossible > position-10]
if len(impossible) == 0:
possible = minima[minima > position]
possible = possible[possible < position+5]
if len(possible) > 0:
edge_energies.append((position + possible[0])/2)
edge_indices.append(np.searchsorted(energy_scale, (position + possible[0])/2))
plt.figure()
plt.plot(energy_scale, spec, label='spectrum')
#plt.plot(energy_scale, first_dif, label='first derivative')
plt.plot(energy_scale, second_dif, label='second derivative')
plt.scatter(energy_scale[edge_indices], spec[edge_indices], color='red', label='onsets')
plt.axhline(0, color='gray')
plt.legend();
# -
# ## Identify Edge
#
# We can now look up which major edge is close each of the onsets that we found.
#
# I'll handle the oxygen edge seprately, because there is always a lot of chemical shift in that edge, and it is very often present.
#
# The we look within all the major edges to determine which one is closest. We will use the function ``find_major_edges`` of **eels_tools** from **pyTEMlib** for that task.
selected_edges = []
for peak in edge_indices:
if 525 < energy_scale[peak] < 533:
selected_edges.append('O-K1')
else:
selected_edge = ''
edges = eels.find_major_edges(energy_scale[peak], 20)
edges = edges.split(('\n'))
minimum_dist = 100.
for edge in edges[1:]:
edge = edge[:-3].replace(' ','').split(':')
name = edge[0].strip()
energy = float(edge[1].strip())
if np.abs(energy-energy_scale[peak])<minimum_dist:
minimum_dist = np.abs(energy-energy_scale[peak])
selected_edge = name
if selected_edge != '':
selected_edges.append(selected_edge)
print('Found edges: ', selected_edges)
# ### Plot Identified Edge Onsets
#
# Here we do everything we explained above in function ``find_edges`` of **eels_tools** from **pyTEMlib** and then we plot this information. You can now accurately determine the chemical shift, by just taking the difference between the actual and tabulated edge onset.
# +
dim = ft.get_dimensions_by_type('spectral', current_dataset)
energy_scale = np.array(dim[0][1])
spec = scipy.ndimage.gaussian_filter(current_dataset,2.5)
selected_edges = eels.find_edges(current_dataset, sensitivity=3)
print(selected_edges)
plt.figure()
plt.plot(energy_scale, spec)
for edge in selected_edges:
atomic_number = eels.get_z(edge.split('-')[0])
edge_info = eels.get_x_sections(atomic_number)
plt.axvline(edge_info[edge.split('-')[1]]['onset'], color='gray')
_, y_max = plt.gca().get_ylim()
plt.text(edge_info[edge.split('-')[1]]['onset'], y_max*1.01, edge)
# -
# ## Peak Detection
# Using the second derivative, can also get a good start for the peak detection in EELS spectra. The second derivative test says that the minima of the second derivative coninceide with maxima and so using that we have already determined most relevant peaks for the electron energy-loss near edge structure (ELNES)
# ## Interpretation of Chemical Shift
#
# While the interpretation of the chemical shift is rather complicated, the determination of the real edge onset may be obscured by a sharp feature at the onset of an edge.
#
# Please change the position (approximately from -10 to 10) and see what the resulting shape and edge onset does.
#
# +
# Input
resolution = 2. #in eV
peak_position = -0 # in eV relative to edge onset
######################################
# calculate h in channels.
h = int(resolution/0.25)
peak_channel = int(500+peak_position/0.25)
# make energy scale
energy_scale = np.arange(1024)*.25+200
# make spectrum with powerlaw background
A = 1e10
r = 3.
spectrum = A* np.power(energy_scale,-r)
# add edge
spectrum[500:] = spectrum[500:]*2.4
original_inflection_point = spectrum[500]*.7
print(original_inflection_point)
spectrum[peak_channel:peak_channel+3] = spectrum[peak_channel:peak_channel+3]*4
f_x = broadened_spectrum = scipy.ndimage.gaussian_filter(spectrum,h)[h:-h]
second_derivative = np.roll(f_x, -3) - 2 * f_x + np.roll(f_x, +3)
second_derivative[:3] = 0
second_derivative[-3:] = 0
# plot
plt.figure()
plt.plot(energy_scale[h:-h], spectrum[h:-h], linewidth=2, label='original simulated spectrum')
plt.plot(energy_scale[h:-h], broadened_spectrum, linewidth=2,
label=f'broadened spectrum with resolution {resolution:.1f} eV')
plt.scatter(energy_scale[500], original_inflection_point, color='red')
plt.ylim(0,spectrum[0]*1.05)
plt.plot(energy_scale[h:-h],second_derivative[h:-h])
plt.legend();
# -
# ## Summary
#
# The only input parameter that we used was the sensitivity factor (to the standard deviation) and we got a list of edges present in the spectra. Because of overlapping of edges of different elements, that may not always work flawlessly and so in a compositional analysis, we will have to verify that those elements are present in the investigated material.
#
# We also determined the chemical shift of the edges. Obviously that can only be as accurate as the accuracy of the energy scale. A reference edge in the spectrum and/or a well characterized experimental setup are essential for high quality chemical shift measurements.
#
# The chemical shift is not easy to interpret, but is further complicated if there are sharp features near or at the edge onsets like excitons, d- or f-bands.
#
# + [markdown] hideCode=false hidePrompt=true
# ## Close File
# File needs to be closed to be used with other notebooks
# + hideCode=false hidePrompt=true
current_dataset.h5_dataset.file.close()
# + [markdown] hideCode=true hidePrompt=true
# ## Navigation
# <font size = "5"> **Back: [Calculating Dielectric Function II: Silicon](DielectricDFT2.ipynb)** </font>
#
# <font size = "5"> **Next: [ELNES](ELNES.ipynb)** </font>
#
# <font size = "5"> **Chapter 4: [Spectroscopy](Spectroscopy.ipynb)** </font>
#
# <font size = "5"> **Index: [Index](Analysis_of_Transmission_Electron_Microscope_Data.ipynb)** </font>
# + hideCode=true hidePrompt=true
| notebooks/.ipynb_checkpoints/4_2_1_Edge_Onset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Making multipanel plots with matplotlib
#
# first, we inport numpy and matplotlib as usual
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Then we define an array of angles, and their sines and cosines using numpy. This time we will use linspace
# +
x = np.linspace(0,2*np.pi,100)
print(x[-1],2*np.pi)
y = np.sin(x)
z = np.cos(x)
w = np.sin(4*x)
v = np.cos(4*x)
# -
# Now, let's make a two panel plot side-by-side
# +
f, axarr = plt.subplots(1,2)
axarr[0].plot(x, y)
axarr[0].set_xlabel('x')
axarr[0].set_ylabel('sin(x)')
axarr[0].set_title(r'$\sin(x)$')
axarr[1].plot(x, z)
axarr[1].set_xlabel('x')
axarr[1].set_ylabel('cos(x)')
axarr[1].set_title(r'$\cos(x)$')
f.subplots_adjust(wspace=0.4)
axarr[0].set_aspect('equal')
axarr[1].set_aspect(np.pi)
# -
# ### Let's keep the figure, merge them, remove the titles and add legends
# +
#adjust the size of the figure
fig = plt.figure(figsize=(6,6))
plt.plot(x, y, label=r'$y = \sin(x)$')
plt.plot(x, z, label=r'$y = \cos(x)$')
plt.plot(x, w, label=r'$y = \sin(4x)$')
plt.plot(x, v, label=r'$y = \cos(4x)$')
plt.xlabel(r'$x$')
plt.ylabel(r'$y(x)$')
plt.xlim([0,2*np.pi])
plt.ylim([-1.2,1.2])
plt.legend(lcc=1, framealpha=0.95)
plt.gca().set_aspect(np.pi/1.2)
# +
#adjust the size of the figure
fig = plt.figure(figsize=(6,6))
plt.plot(x, y, label=r'$y = \sin(x)$')
plt.plot(x, z, label=r'$y = \cos(x)$')
plt.plot(x, w, label=r'$y = \sin(4x)$')
plt.plot(x, v, label=r'$y = \cos(4x)$')
plt.xlabel(r'$x$')
plt.ylabel(r'$y(x)$')
plt.xlim([0,2*np.pi])
plt.ylim([-1.2,1.2])
plt.legend(loc=1, framealpha=0.95)
plt.gca().set_aspect(np.pi/1.2)
# -
| Multiplanel Figures and legend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3-stat-ML
# language: python
# name: python3-stat-ml
# ---
# %pylab inline
import numpy as np
import matplotlib.pyplot as plt
# +
def sigma(x):
return 1 / (1 + np.exp(-x))
plt.figure(figsize(8,5))
X = np.linspace(-10, 10, 100)
plt.plot(X, sigma(X),'b')
plt.xlabel('$x = \sum_{j=1} w_jx_j + w_0$', fontsize=12)
plt.ylabel('out', fontsize=12)
plt.title('Sigmoid Function', fontsize=20)
plt.axvline(0, color='k', lw=.5)
plt.axhline(0, color='k', lw=.5)
#plt.grid()
plt.text(4, 0.9, r'$\sigma(x)=\frac{1}{1+e^{-x}}$', fontsize=16)
plt.tight_layout()
plt.savefig('figures/Sigmoid.png', dpi=300, transparency=True)
# +
def tanh(x):
return np.tanh(x)
plt.figure(figsize(8,5))
X = np.linspace(-10, 10, 100)
plt.plot(X, tanh(X),'b', label='tanh(x)')
plt.xlabel('$x = \sum_{j=1} w_jx_j + w_0$', fontsize=12)
plt.ylabel('out', fontsize=12)
plt.title('Tanh Function', fontsize=20)
plt.axvline(0, color='k', lw=.5)
plt.axhline(0, color='k', lw=.5)
#plt.grid()
#plt.text(4, 0.8, r'$\tanh(x)=\frac{1}{1+e^{-x}}$', fontsize=16)
plt.tight_layout()
plt.savefig('figures/Tanh.png', dpi=300)
# +
# alternative activation function
def ReLU(x):
return np.maximum(0.0, x)
# derivation of relu
def ReLU_derivation(x):
if x <= 0:
return 0
else:
return 1
plt.figure(figsize(8,5))
X = np.linspace(-10, 10, 100)
plt.plot(X, ReLU(X),'b')
plt.xlabel('$x = \sum_{j=1} w_jx_j + w_0$', fontsize=12)
plt.ylabel('out', fontsize=12)
plt.title('ReLU Function', fontsize=20)
plt.axvline(0, color='k', lw=.5)
plt.axhline(0, color='k', lw=.5)
#plt.grid()
plt.text(2.2, 9, r'$ReLU(x)=max(0.0, x)$', fontsize=12)
plt.tight_layout()
plt.savefig('figures/RELU.png', dpi=300)
# -
| notebooks/activation_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#export
from local.torch_basics import *
from local.test import *
from local.core import *
from local.data.all import *
from local.notebook.showdoc import show_doc
#export
pd.set_option('mode.chained_assignment','raise')
# +
#default_exp tabular.core
# -
# # Tabular core
#
# > Basic function to preprocess tabular data before assembling it in a `DataBunch`.
# ## TabularProc -
# We use this class to preprocess tabular data. `cat_names` should contain the names of the categorical variables in your dataframe, `cont_names` the names of the continuous variables. If you don't need any state, you can initiliaze a `TabularProc` with a `func` to be applied on the dataframes. Otherwise you should subclass and implement `setup` and `__call__`.
#export
class Tabular(CollBase):
def __init__(self, df, cat_names=None, cont_names=None, y_names=None, is_y_cat=True, splits=None):
super().__init__(df)
self.splits = L(ifnone(splits,slice(None)))
store_attr(self, 'y_names,is_y_cat')
self.cat_names,self.cont_names = L(cat_names),L(cont_names)
self.cat_y = None if not is_y_cat else y_names
self.cont_y = None if is_y_cat else y_names
def _new(self, df):
return Tabular(df, self.cat_names, self.cont_names, y_names=self.y_names, is_y_cat=self.is_y_cat, splits=self.splits)
def set_col(self,k,v): super().__setitem__(k, v)
def transform(self, cols, f): self.set_col(cols, self.loc[:,cols].transform(f))
def show(self, max_n=10, **kwargs): display_df(self.all_cols[:max_n])
def __getitem__(self, idxs): return self._new(self.items.iloc[idxs])
def __getattr__(self,k):
if k.startswith('_') or k=='items': raise AttributeError
return getattr(self.items,k)
@property
def __array__(self): return self.items.__array__
@property
def iloc(self): return self
@property
def loc(self): return self.items.loc
@property
def targ(self): return self.loc[:,self.y_names]
@property
def all_cont_names(self): return self.cont_names + self.cont_y
@property
def all_cat_names (self): return self.cat_names + self.cat_y
@property
def all_col_names (self): return self.all_cont_names + self.all_cat_names
# +
#export
def _add_prop(cls, nm):
prop = property(lambda o: o.items[list(getattr(o,nm+'_names'))])
setattr(cls, nm+'s', prop)
def _f(o,v): o.set_col(getattr(o,nm+'_names'), v)
setattr(cls, nm+'s', prop.setter(_f))
_add_prop(Tabular, 'cat')
_add_prop(Tabular, 'all_cat')
_add_prop(Tabular, 'cont')
_add_prop(Tabular, 'all_cont')
_add_prop(Tabular, 'all_col')
# -
df = pd.DataFrame({'a':[0,1,2,0,2], 'b':[0,0,0,0,0]})
to = Tabular(df, 'a')
t = pickle.loads(pickle.dumps(to))
test_eq(t.items,to.items)
to.show() # only shows 'a' since that's the only col in `Tabular`
#export
class TabularProc(InplaceTransform):
"Base class to write a tabular processor for dataframes"
def process(self, *args,**kwargs): return self(*args,**kwargs)
#export
class Categorify(TabularProc, CollBase):
"Transform the categorical variables to that type."
order = 1
def setup(self, to):
to.classes = self.items = {n:CategoryMap(to.loc[to.splits[0],n], add_na=True)
for n in to.all_cat_names}
def _apply_cats(self, c): return c.cat.codes+1 if is_categorical_dtype(c) else c.map(self[c.name].o2i)
def encodes(self, to): to.transform(to.all_cat_names, self._apply_cats)
def _decode_cats(self, c): return c.map(dict(enumerate(self[c.name].items)))
def decodes(self, to): to.transform(to.all_cat_names, self._decode_cats)
show_doc(Categorify, title_level=3)
# +
cat = Categorify()
df = pd.DataFrame({'a':[0,1,2,0,2]})
to = Tabular(df, 'a')
cat.setup(to)
test_eq(cat['a'], ['#na#',0,1,2])
cat(to)
test_eq(df['a'], [1,2,3,1,3])
df1 = pd.DataFrame({'a':[1,0,3,-1,2]})
to1 = Tabular(df1, 'a')
cat(to1)
#Values that weren't in the training df are sent to 0 (na)
test_eq(df1['a'], [2,1,0,0,3])
to2 = cat.decode(to1)
test_eq(to2.a, [1,0,'#na#','#na#',2])
# -
#test with splits
cat = Categorify()
df = pd.DataFrame({'a':[0,1,2,3,2]})
to = Tabular(df, 'a', splits=[range(3)])
cat.setup(to)
test_eq(cat['a'], ['#na#',0,1,2])
cat(to)
test_eq(df['a'], [1,2,3,0,3])
df = pd.DataFrame({'a':pd.Categorical(['M','H','L','M'], categories=['H','M','L'], ordered=True)})
to = Tabular(df, 'a')
cat = Categorify()
cat.setup(to)
test_eq(cat['a'], ['#na#','H','M','L'])
cat(to)
test_eq(df.a, [2,1,3,2])
to2 = cat.decode(to)
test_eq(to2.a, ['M','H','L','M'])
#export
class Normalize(TabularProc):
"Normalize the continuous variables."
order = 2
def setup(self, to):
df = to.loc[to.splits[0], to.cont_names]
self.means,self.stds = df.mean(),df.std(ddof=0)+1e-7
def encodes(self, to): to.conts = (to.conts-self.means) / self.stds
def decodes(self, to): to.conts = (to.conts*self.stds ) + self.means
show_doc(Normalize, title_level=3)
norm = Normalize()
df = pd.DataFrame({'a':[0,1,2,3,4]})
to = Tabular(df, cont_names='a')
norm.setup(to)
x = np.array([0,1,2,3,4])
m,s = x.mean(),x.std()
test_eq(norm.means['a'], m)
test_close(norm.stds['a'], s)
norm(to)
test_close(df['a'].values, (x-m)/s)
df1 = pd.DataFrame({'a':[5,6,7]})
to1 = Tabular(df1, cont_names='a')
norm(to1)
test_close(df1['a'].values, (np.array([5,6,7])-m)/s)
to2 = norm.decode(to1)
test_close(to2.a.values, [5,6,7])
norm = Normalize()
df = pd.DataFrame({'a':[0,1,2,3,4]})
to = Tabular(df, cont_names='a', splits=[range(3)])
norm.setup(to)
x = np.array([0,1,2])
m,s = x.mean(),x.std()
test_eq(norm.means['a'], m)
test_close(norm.stds['a'], s)
norm(to)
test_close(df['a'].values, (np.array([0,1,2,3,4])-m)/s)
#export
class FillStrategy:
"Namespace containing the various filling strategies."
def median (c,fill): return c.median()
def constant(c,fill): return fill
def mode (c,fill): return c.dropna().value_counts().idxmax()
#export
class FillMissing(TabularProc):
"Fill the missing values in continuous columns."
def __init__(self, fill_strategy=FillStrategy.median, add_col=True, fill_vals=None):
if fill_vals is None: fill_vals = defaultdict(int)
store_attr(self, 'fill_strategy,add_col,fill_vals')
def setup(self, to):
df = to.loc[to.splits[0], to.cont_names]
self.na_dict = {n:self.fill_strategy(df[n], self.fill_vals[n])
for n in pd.isnull(to.conts).any().keys()}
def encodes(self, to):
missing = pd.isnull(to.conts)
for n in missing.any().keys():
assert n in self.na_dict, f"nan values in `{n}` but not in setup training set"
to.loc[:,n].fillna(self.na_dict[n], inplace=True)
if self.add_col:
to.loc[:,n+'_na'] = missing[n]
if n+'_na' not in to.cat_names: to.cat_names.append(n+'_na')
show_doc(FillMissing, title_level=3)
# +
fill1,fill2,fill3 = (FillMissing(fill_strategy=s)
for s in [FillStrategy.median, FillStrategy.constant, FillStrategy.mode])
df = pd.DataFrame({'a':[0,1,np.nan,1,2,3,4]})
df1 = df.copy(); df2 = df.copy()
to,to1,to2 = Tabular(df, cont_names='a'),Tabular(df1, cont_names='a'),Tabular(df2, cont_names='a')
fill1.setup(to); fill2.setup(to1); fill3.setup(to2)
test_eq(fill1.na_dict, {'a': 1.5})
test_eq(fill2.na_dict, {'a': 0})
test_eq(fill3.na_dict, {'a': 1.0})
fill1(to); fill2(to1); fill3(to2)
for t in [to, to1, to2]: test_eq(t.cat_names, ['a_na'])
for to_,v in zip([to, to1, to2], [1.5, 0., 1.]):
test_eq(to_.a.values, np.array([0, 1, v, 1, 2, 3, 4]))
test_eq(to_.a_na.values, np.array([0, 0, 1, 0, 0, 0, 0]))
dfa = pd.DataFrame({'a':[np.nan,0,np.nan]})
dfa1 = dfa.copy(); dfa2 = dfa.copy()
to,to1,to2 = Tabular(dfa, cont_names='a'),Tabular(dfa1, cont_names='a'),Tabular(dfa2, cont_names='a')
fill1(to); fill2(to1); fill3(to2)
for to_,v in zip([to, to1, to2], [1.5, 0., 1.]):
test_eq(to_.a.values, np.array([v, 0, v]))
test_eq(to_.a_na.values, np.array([1, 0, 1]))
# -
# ## Tabular Pipelines -
# +
procs = [Normalize(), Categorify(), FillMissing(), noop]
proc = Pipeline(procs)
#Test reordering and partialize
test_eq(L(proc.fs).mapped(type), [FillMissing, Transform, Categorify, Normalize])
df = pd.DataFrame({'a':[0,1,2,1,1,2,0], 'b':[0,1,np.nan,1,2,3,4]})
to = Tabular(df, 'a', 'b')
#Test setup and apply on df_trn
proc.setup(to)
test_eq(to.cat_names, ['a', 'b_na'])
test_eq(to.a, [1,2,3,2,2,3,1])
test_eq(to.b_na, [1,1,2,1,1,1,1])
x = np.array([0,1,1.5,1,2,3,4])
m,s = x.mean(),x.std()
test_close(to.b.values, (x-m)/s)
test_eq(proc.classes, {'a': ['#na#',0,1,2], 'b_na': ['#na#',False,True]})
# +
#Test apply on y_names
procs = [Normalize(), Categorify(), FillMissing(), noop]
proc = Pipeline(procs)
df = pd.DataFrame({'a':[0,1,2,1,1,2,0], 'b':[0,1,np.nan,1,2,3,4], 'c': ['b','a','b','a','a','b','a']})
to = Tabular(df, 'a', 'b', y_names='c')
proc.setup(to)
test_eq(to.cat_names, ['a', 'b_na'])
test_eq(to.a, [1,2,3,2,2,3,1])
test_eq(to.b_na, [1,1,2,1,1,1,1])
test_eq(to.c, [2,1,2,1,1,2,1])
x = np.array([0,1,1.5,1,2,3,4])
m,s = x.mean(),x.std()
test_close(to.b.values, (x-m)/s)
test_eq(proc.classes, {'a': ['#na#',0,1,2], 'b_na': ['#na#',False,True], 'c': ['#na#','a','b']})
# -
#export
@delegates(Tabular)
def process_df(df, procs, inplace=True, **kwargs):
"Process `df` with `procs` and returns the processed dataframe and the `TabularProcessor` associated"
to = Tabular(df if inplace else df.copy(), **kwargs)
proc = Pipeline(procs)
proc.setup(to)
return to,proc
procs = [Normalize(), Categorify(), FillMissing(), noop]
df = pd.DataFrame({'a':[0,1,2,1,1,2,0], 'b':[0,1,np.nan,1,2,3,4], 'c': ['b','a','b','a','a','b','a']})
to,proc = process_df(df, procs, cat_names='a', cont_names='b', y_names='c', inplace=False)
test_eq(to.cat_names, ['a', 'b_na'])
test_eq(to.a, [1,2,3,2,2,3,1])
test_eq(df.a.dtype,int)
test_eq(to.b_na, [1,1,2,1,1,1,1])
test_eq(to.c, [2,1,2,1,1,2,1])
# Pass the same `splits` as you will use for splitting the data, so that the setup is only done on the training set. `cat_names` are the names of the categorical variables, `cont_names` the continous ones, `y_names` are the names of the dependent variables that are categories. If `inplace=True`, processing is applied inplace, otherwis it creates a copy of `df`.
#export
class ReadTabBatch(ItemTransform):
def __init__(self, proc): self.proc = proc
def encodes(self, to):
return (tensor(to.cats.values).long(),tensor(to.conts.values).float()), tensor(to.targ.values).long()
def decodes(self, o):
(cats,conts),targs = to_np(o)
df = pd.DataFrame({**{c: cats [:,i] for i,c in enumerate(self.proc.cat_names )},
**{c: conts[:,i] for i,c in enumerate(self.proc.cont_names)},
self.proc.y_names: targs})
to = Tabular(df, self.proc.cat_names, self.proc.cont_names, self.proc.y_names, is_y_cat=self.proc.cat_y is not None)
to = self.proc.decode(to)
return to
#export
@delegates()
class TabDataLoader(TfmdDL):
do_item = noops
def __init__(self, dataset, proc, bs=16, shuffle=False, after_batch=None, num_workers=0, **kwargs):
after_batch = L(after_batch)+ReadTabBatch(proc)
super().__init__(dataset, bs=bs, shuffle=shuffle, after_batch=after_batch, num_workers=num_workers, **kwargs)
def create_batch(self, b): return self.dataset.items[b]
# ## Integration example
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
df_trn,df_tst = df.iloc[:10000].copy(),df.iloc[10000:].copy()
df_trn.head()
# +
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['age', 'fnlwgt', 'education-num']
procs = [Categorify(), FillMissing(), Normalize()]
splits = RandomSplitter()(range_of(df_trn))
to,proc = process_df(df_trn, procs, splits=splits, cat_names=cat_names, cont_names=cont_names, y_names="salary")
# -
dsrc = DataSource(to, filts=splits, tfms=[None])
dl = TabDataLoader(dsrc.valid, proc, bs=64, num_workers=0)
dl.show_batch()
to_tst = Tabular(df_tst, cat_names, cont_names, y_names="salary")
proc(to_tst)
to_tst.all_cols.head()
# ## Not being used now - for multi-modal
# +
class TensorTabular(TupleBase):
def get_ctxs(self, max_n=10, **kwargs):
n_samples = min(self[0].shape[0], max_n)
df = pd.DataFrame(index = range(n_samples))
return [df.iloc[i] for i in range(n_samples)]
def display(self, ctxs): display_df(pd.DataFrame(ctxs))
class TabularLine(pd.Series):
"A line of a dataframe that knows how to show itself"
def show(self, ctx=None, **kwargs): return self if ctx is None else ctx.append(self)
class ReadTabLine(ItemTransform):
def __init__(self, proc): self.proc = proc
def encodes(self, row):
cats,conts = (o.mapped(row.__getitem__) for o in (self.proc.cat_names,self.proc.cont_names))
return TensorTabular(tensor(cats).long(),tensor(conts).float())
def decodes(self, o):
to = Tabular(o, self.proc.cat_names, self.proc.cont_names, self.proc.y_names)
to = self.proc.decode(to)
return TabularLine(pd.Series({c: v for v,c in zip(to.items[0]+to.items[1], self.proc.cat_names+self.proc.cont_names)}))
class ReadTabTarget(ItemTransform):
def __init__(self, proc): self.proc = proc
def encodes(self, row): return row[self.proc.y_names].astype(np.int64)
def decodes(self, o): return Category(self.proc.classes[self.proc.y_names][o])
# +
# tds = TfmdDS(to.items, tfms=[[ReadTabLine(proc)], ReadTabTarget(proc)])
# enc = tds[1]
# test_eq(enc[0][0], tensor([2,1]))
# test_close(enc[0][1], tensor([-0.628828]))
# test_eq(enc[1], 1)
# dec = tds.decode(enc)
# assert isinstance(dec[0], TabularLine)
# test_close(dec[0], pd.Series({'a': 1, 'b_na': False, 'b': 1}))
# test_eq(dec[1], 'a')
# test_stdout(lambda: print(tds.show_at(1)), """a 1
# b_na False
# b 1
# category a
# dtype: object""")
# -
# ## Export -
#hide
from local.notebook.export import notebook2script
notebook2script(all_fs=True)
| dev/40_tabular_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Whale off the Port(folio)
# ---
#
# In this assignment, you'll get to use what you've learned this week to evaluate the performance among various algorithmic, hedge, and mutual fund portfolios and compare them against the S&P 500 Index.
# +
# Initial imports
import pandas as pd
import numpy as np
import datetime as dt
from pathlib import Path
# %matplotlib inline
# -
# # Data Cleaning
#
# In this section, you will need to read the CSV files into DataFrames and perform any necessary data cleaning steps. After cleaning, combine all DataFrames into a single DataFrame.
#
# Files:
#
# * `whale_returns.csv`: Contains returns of some famous "whale" investors' portfolios.
#
# * `algo_returns.csv`: Contains returns from the in-house trading algorithms from Harold's company.
#
# * `sp500_history.csv`: Contains historical closing prices of the S&P 500 Index.
# ## Whale Returns
#
# Read the Whale Portfolio daily returns and clean the data
# +
# Reading whale returns
csv_path_whale = Path("./Resources/whale_returns.csv")
whale_daily_df = pd.read_csv(csv_path_whale)
whale_daily_df.set_index(pd.to_datetime(whale_daily_df['Date'], infer_datetime_format = True), inplace = True)
whale_daily_df.drop(columns = ["Date"], inplace = True)
whale_daily_df.head()
# -
# Count nulls
whale_daily_df.isnull().sum()
# +
# Drop nulls
whale_daily_df = whale_daily_df.dropna().copy()
whale_daily_df.isnull().sum()
# -
# ## Algorithmic Daily Returns
#
# Read the algorithmic daily returns and clean the data
# +
# Reading algorithmic returns
csv_path_algo = Path("./Resources/algo_returns.csv")
algo_daily_df = pd.read_csv(csv_path_algo)
algo_daily_df.set_index(pd.to_datetime(algo_daily_df['Date'], infer_datetime_format = True),inplace = True)
algo_daily_df.drop(columns = ["Date"], inplace = True)
algo_daily_df.head()
# -
# Count nulls
algo_daily_df.isnull().sum()
# +
# Drop nulls
algo_daily_df = algo_daily_df.dropna().copy()
algo_daily_df.isnull().sum()
# -
# ## S&P 500 Returns
#
# Read the S&P 500 historic closing prices and create a new daily returns DataFrame from the data.
# +
# Reading S&P 500 Closing Prices
csv_path_sp500 = Path("./Resources/sp500_history.csv")
sp500_history_df = pd.read_csv(csv_path_sp500)
sp500_history_df.set_index(pd.to_datetime(sp500_history_df['Date'], infer_datetime_format = True), inplace = True)
sp500_history_df.drop(columns = ["Date"], inplace = True)
sp500_history_df = sp500_history_df.sort_index()
sp500_history_df.head()
# -
# Check Data Types
sp500_history_df.dtypes
# +
# Fix Data Types
# Drops "$" from Close column
sp500_history_df["Close"] = sp500_history_df["Close"].str.replace('$', '', regex = False)
# Converts close column from object to float
sp500_history_df["Close"] = sp500_history_df["Close"].astype('float')
sp500_history_df.dtypes
# +
# Calculate Daily Returns
sp500_daily_return_df = sp500_history_df.pct_change()
sp500_daily_return_df
# +
# Drop nulls
sp500_daily_return_df = sp500_daily_return_df.dropna().copy()
sp500_daily_return_df.isnull().sum()
# +
# Rename `Close` Column to be specific to this portfolio.
sp500_daily_return_df = sp500_daily_return_df.rename(columns = {"Close":"SP 500"})
sp500_daily_return_df.head()
# -
# ## Combine Whale, Algorithmic, and S&P 500 Returns
# +
# Join Whale Returns, Algorithmic Returns, and the S&P 500 Returns into a single DataFrame with columns for each portfolio's returns.
all_portfolios_df = pd.concat([whale_daily_df, algo_daily_df, sp500_daily_return_df], axis = "columns", join = "inner")
all_portfolios_df.tail()
# -
# ---
# # Conduct Quantitative Analysis
#
# In this section, you will calculate and visualize performance and risk metrics for the portfolios.
# ## Performance Anlysis
#
# #### Calculate and Plot the daily returns.
# +
# Plot daily returns of all portfolios
all_portfolios_df.plot(title = "Daily for Returns for Portfolios and S&P", figsize = (10,5))
# -
# #### Calculate and Plot cumulative returns.
# +
# Calculate cumulative returns of all portfolios
cumulative_return = (1+all_portfolios_df).cumprod()
print(cumulative_return.tail())
# Plot cumulative returns
cumulative_return.plot(title = "Cumulative returns", figsize=(10,5))
# -
# ---
# ## Risk Analysis
#
# Determine the _risk_ of each portfolio:
#
# 1. Create a box plot for each portfolio.
# 2. Calculate the standard deviation for all portfolios
# 4. Determine which portfolios are riskier than the S&P 500
# 5. Calculate the Annualized Standard Deviation
# ### Create a box plot for each portfolio
#
# +
# Box plot to visually show risk
all_portfolios_df.plot(kind = "box", figsize =(20,10))
# -
# ### Calculate Standard Deviations
# +
# Calculate the daily standard deviations of all portfolios
all_portfolios_df_sdev = all_portfolios_df.iloc[0:,0:6]
all_portfolios_df_sdev = all_portfolios_df_sdev.std(ddof = 1)
all_portfolios_df_sdev = pd.DataFrame(all_portfolios_df_sdev)
all_portfolios_df_sdev
# + [markdown] tags=[]
# ### Determine which portfolios are riskier than the S&P 500
# +
# Calculate the daily standard deviation of S&P 500
sp500_daily_return_df_std = all_portfolios_df.iloc[0:,-1:]
sp500_daily_return_df_std = float(sp500_daily_return_df_std.std(ddof = 1))
# print(sp500_daily_return_df_std)
# Determine which portfolios are riskier than the S&P 500
for portfolios in all_portfolios_df_sdev.iterrows():
if float(portfolios[1]) > sp500_daily_return_df_std:
print(f" {portfolios[0]} is more risky than S&P 500")
# -
# ### Calculate the Annualized Standard Deviation
# +
# Calculate the annualized standard deviation (252 trading days)
all_portfolio_annual_std = (all_portfolios_df.std())*(252**.5)
all_portfolio_annual_std
# -
# ---
# ## Rolling Statistics
#
# Risk changes over time. Analyze the rolling statistics for Risk and Beta.
#
# 1. Calculate and plot the rolling standard deviation for the S&P 500 using a 21-day window
# 2. Calculate the correlation between each stock to determine which portfolios may mimick the S&P 500
# 3. Choose one portfolio, then calculate and plot the 60-day rolling beta between it and the S&P 500
# ### Calculate and plot rolling `std` for all portfolios with 21-day window
# +
# Calculate the rolling standard deviation for all portfolios using a 21-day window
all_portfolios_rolling_std = all_portfolios_df.rolling(21).std()
all_portfolios_rolling_std.head(25)
# Plot the rolling standard deviation
all_portfolios_rolling_std.plot(figsize = (20,10))
# -
# ### Calculate and plot the correlation
# +
# Calculate the correlation
all_portfolio_correlation = all_portfolios_df.corr()
# Display de correlation matrix
all_portfolio_correlation
# -
# ### Calculate and Plot Beta for a chosen portfolio and the S&P 500
# +
# Calculate covariance of a single portfolio
algo1_covar = all_portfolios_df.iloc[0:,4].cov(all_portfolios_df.iloc[0:,6])
print(algo1_covar)
# Calculate variance of S&P 500
variance = all_portfolios_df.iloc[0:,6].var()
print(variance)
# Computing beta
beta = algo1_covar/variance
print(beta)
# Plot beta trend
rolling_covariance = all_portfolios_df['Algo 1'].rolling(window=60).cov(all_portfolios_df["SP 500"].rolling(window=60))
rolling_variance = all_portfolios_df['SP 500'].rolling(window=60).var()
rolling_beta = rolling_covariance/rolling_variance
rolling_beta.plot(figsize = (20,10))
# -
# ## Rolling Statistics Challenge: Exponentially Weighted Average
#
# An alternative way to calculate a rolling window is to take the exponentially weighted moving average. This is like a moving window average, but it assigns greater importance to more recent observations. Try calculating the [`ewm`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.ewm.html) with a 21-day half-life.
# Use `ewm` to calculate the rolling window
# ---
# # Sharpe Ratios
# In reality, investment managers and thier institutional investors look at the ratio of return-to-risk, and not just returns alone. After all, if you could invest in one of two portfolios, and each offered the same 10% return, yet one offered lower risk, you'd take that one, right?
#
# ### Using the daily returns, calculate and visualize the Sharpe ratios using a bar plot
# +
# Annualized Sharpe Ratios
sharpe_ratios = (all_portfolios_df.mean()* 252) / all_portfolio_annual_std
print(sharpe_ratios)
# +
# Visualize the sharpe ratios as a bar plot
sharpe_ratios.plot(kind="bar", figsize = (20,10))
# -
### Determine whether the algorithmic strategies outperform both the market (S&P 500) and the whales portfolios.
# The first algorithmic strategy outperformed the whale portfolios and the market. It's cumulative return of 93% was the highest of all portfolios and it did so with the lowest measure of risk .121 other than Paulson which had an annualized standard deviation of .11. However, Paulson only had a cumulative return of less than one which means they lost money in this time frame. Algo 1 trading strategy also had the lowest correlation relative to all the other strategies and had the highest Sharpe ratio of 1.38. So based on the highest Sharpe ratio it had the best risk adjusted return and the best cumulative return over this period.
# The second algorithmic strategy had a lower cumulative return than Berkshire Hathaway and the S&P 500, but a higher return than the rest of the whale portfolios. However, it had a lower level of risk, .132 than the S&P 500, .136 and Berkshire Hathaway, .21. So Algo 2 strategy was less volatile than Berkshire Hathaway, the S&P 500, and the rest of the whale portfolios, other than Paulson & Co. The Algo 2 portfolio also had a lower Sharpe Ratio .50 than S&P 500, .65 and Berkshire Hathaway, .62.
# In conclusion, the Algo 1 portfolio had the best risk-adjusted return compared to the S&P 500 and the whale portfolios. The Algo 2 portfolio outperformed the Soros Fund, Paulson & Co, and Tiger Global Management with its risk adjusted return, but did worse than the S&P 500 and Berkshire Hathaway.
#
# # Create Custom Portfolio
#
# In this section, you will build your own portfolio of stocks, calculate the returns, and compare the results to the Whale Portfolios and the S&P 500.
#
# 1. Choose 3-5 custom stocks with at last 1 year's worth of historic prices and create a DataFrame of the closing prices and dates for each stock.
# 2. Calculate the weighted returns for the portfolio assuming an equal number of shares for each stock
# 3. Join your portfolio returns to the DataFrame that contains all of the portfolio returns
# 4. Re-run the performance and risk analysis with your portfolio to see how it compares to the others
# 5. Include correlation analysis to determine which stocks (if any) are correlated
# ## Choose 3-5 custom stocks with at last 1 year's worth of historic prices and create a DataFrame of the closing prices and dates for each stock.
#
# For this demo solution, we fetch data from three companies listes in the S&P 500 index.
#
# * `GOOG` - [Google, LLC](https://en.wikipedia.org/wiki/Google)
#
# * `AAPL` - [Apple Inc.](https://en.wikipedia.org/wiki/Apple_Inc.)
#
# * `COST` - [Costco Wholesale Corporation](https://en.wikipedia.org/wiki/Costco)
# +
# Reading data from 1st stock
csv_path_aapl = Path("./Resources/aapl_historical.csv")
aapl_daily_df = pd.read_csv(csv_path_aapl)
aapl_daily_df.set_index(pd.to_datetime(aapl_daily_df['Trade DATE'], infer_datetime_format = True), inplace = True)
aapl_daily_df.drop(columns = ["Trade DATE"], inplace = True)
aapl_daily_df = aapl_daily_df.sort_index()
aapl_daily_df.head()
# +
# Reading data from 2nd stock
csv_path_goog = Path("./Resources/goog_historical.csv")
goog_daily_df = pd.read_csv(csv_path_goog)
goog_daily_df.set_index(pd.to_datetime(goog_daily_df['Trade DATE'], infer_datetime_format = True), inplace = True)
goog_daily_df.drop(columns = ["Trade DATE"], inplace = True)
goog_daily_df = goog_daily_df.sort_index(ascending = True)
goog_daily_df.head()
# +
# Reading data from 3rd stock
csv_path_cost = Path("./Resources/cost_historical.csv")
cost_daily_df = pd.read_csv(csv_path_cost)
cost_daily_df.set_index(pd.to_datetime(cost_daily_df['Trade DATE'], infer_datetime_format = True), inplace = True)
cost_daily_df.drop(columns = ["Trade DATE"], inplace = True)
cost_daily_df = cost_daily_df.sort_index()
cost_daily_df.head()
# -
# Combine all stocks in a single DataFrame
my_portfolio_df = pd.concat([aapl_daily_df, goog_daily_df, cost_daily_df], axis = "columns", join = "inner")
my_portfolio_df
# Reset Date index
my_portfolio_dff = my_portfolio_df.reset_index()
my_portfolio_dff
# +
# Reorganize portfolio data by having a column per symbol
my_portfolio_df = my_portfolio_df.drop(columns = "Symbol")
columns = ["AAPL", "GOOG", "COST"]
my_portfolio_df.columns = columns
my_portfolio_df
# +
# Calculate daily returns
my_daily_return = my_portfolio_df.pct_change()
# Drop NAs
my_daily_return = my_daily_return.dropna()
# Display sample data
my_daily_return
# -
# ## Calculate the weighted returns for the portfolio assuming an equal number of shares for each stock
# +
# Set weights
weights = [1/3, 1/3, 1/3]
# Calculate portfolio return
my_portfolio_return_df = my_daily_return.dot(weights)
my_portfolio_cumulative = (1+my_portfolio_return_df).cumprod()
print(my_portfolio_cumulative)
# Display sample data
my_portfolio_return_df.head()
# -
# ## Join your portfolio returns to the DataFrame that contains all of the portfolio returns
# +
# Join your returns DataFrame to the original returns DataFrame
total_portfolios_df = pd.concat([all_portfolios_df,my_portfolio_return_df], axis = "columns", join = "inner")
total_portfolios_df
# +
# Only compare dates where return data exists for all the stocks (drop NaNs)
total_portfolios_df.dropna()
total_portfolios_df
# -
# ## Re-run the risk analysis with your portfolio to see how it compares to the others
# ### Calculate the Annualized Standard Deviation
# +
# Calculate the annualized `std`
total_portfolios_std = total_portfolios_df.std()*np.sqrt(252)
total_portfolios_std
# -
# ### Calculate and plot rolling `std` with 21-day window
# +
# Calculate rolling standard deviation
total_portfolios_rolling_std = total_portfolios_df.rolling(21).std()
total_portfolios_rolling_std.tail()
# Plot rolling standard deviation
total_portfolios_rolling_std.plot(figsize = (20,10))
# -
# ### Calculate and plot the correlation
# +
# Calculate and plot the correlation
total_portfolios_corr = total_portfolios_df.corr()
total_portfolios_corr
# -
# ### Calculate and Plot Rolling 60-day Beta for Your Portfolio compared to the S&P 500
# +
# Calculate and plot Beta
rolling_covariance = total_portfolios_df[0].rolling(window=60).cov(total_portfolios_df["SP 500"].rolling(window=60))
rolling_variance = total_portfolios_df['SP 500'].rolling(window=60).var()
rolling_beta = rolling_covariance/rolling_variance
rolling_beta.plot(figsize = (20,10))
# -
# ### Using the daily returns, calculate and visualize the Sharpe ratios using a bar plot
# Calculate Annualzied Sharpe Ratios
sharpe_ratios = (total_portfolios_df.mean()* 252) / total_portfolios_std
sharpe_ratios
# +
# Visualize the sharpe ratios as a bar plot
sharpe_ratios.plot(kind = "bar", figsize = (20,10))
# -
# ### How does your portfolio do?
#
# My portfolio had a cumulative return of 13% this period and had a higher level of volatility, .21, than the rest of the portfolios except for Tiger Management, .23 and Berkshire Hathaway, .25. My portfolio had a higher Sharpe ratio, .93 than the portfolios and S&P 500, except for the Algo 1 portfolio. Overall, despite a comparatively higher level of risk, its risk-adjusted return was better that the portfolios other than Algo 1.
| whale_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 0: Import Statements
import numpy as np
import pandas as pd
import pubchempy
# # Part 1: Read in data and parse it
df = pd.read_csv('NF2 Synodos MIPE qHTS March2016.txt', sep='\t')
df
# get unique drugs and the number of drugs tested
unique_drugs = np.unique(list(df['Sample Name']))
num_drugs = len(np.unique(list(df['Sample Name'])))
# get unique cell lines and the number of cell lines tested
unique_lines = np.unique(list(df['Cell line']))
num_lines = len(np.unique(list(df['Cell line'])))
# +
# get array of Max Resp and AUC for each drug for each of 6 cell lines
drug_array_max_resp = np.zeros((num_drugs, num_lines))
# loop through all unique drugs
for drug in range(0, num_drugs):
drug_name = str(unique_drugs[drug])
curr_drug = df.loc[df['Sample Name'] == drug_name]
# loop through each of the cell lines
for line in range(0, num_lines):
val = list(curr_drug.loc[curr_drug['Cell line'] == unique_lines[line], 'Max Resp'])
if (val != []):
val = val[0]
else:
val = None
drug_array_max_resp[drug][line] = val #populate with the max resp
# +
# get array of Max Resp and AUC for each drug for each of 6 cell lines
drug_array_auc = np.zeros((num_drugs, num_lines))
# loop through all unique drugs
for drug in range(0, num_drugs):
drug_name = str(unique_drugs[drug])
curr_drug = df.loc[df['Sample Name'] == drug_name]
# loop through each of the cell lines
for line in range(0, num_lines):
val = list(curr_drug.loc[curr_drug['Cell line'] == unique_lines[line], 'AUC'])
if (val != []):
val = val[0]
else:
val = None
drug_array_auc[drug][line] = val #populate with the max resp
# -
# make pandas dataframes of these drugs + relevant values
max_resp = pd.DataFrame(drug_array_max_resp)
auc = pd.DataFrame(drug_array_auc)
drugs = pd.DataFrame(unique_drugs)
# save dataframes
max_resp.to_csv('max_resp_9_15.csv')
auc.to_csv('auc_9_15.csv')
drugs.to_csv('drugs_9_15.csv')
# # Part 2: Get Pubchem data for each of the drugs
# now get pubchem data- just make sure it works for one example drug
# try printing and accessing different parts of the structural info
for compound in pubchempy.get_compounds(str(unique_drugs[1]), 'name'):
print(compound)
cid = compound.cid
print(cid)
c = pubchempy.Compound.from_cid(cid)
print(unique_drugs[0])
frame = pubchempy.compounds_to_frame(c, properties=['molecular_weight', 'exact_mass', 'monoisotopic_mass', 'xlogp', 'tpsa', 'complexity', 'charge',
'h_bond_donor_count','h_bond_acceptor_count','rotatable_bond_count','heavy_atom_count','isotope_atom_count',
'atom_stereo_count','bond_stereo_count','covalent_unit_count'])
print(frame)
print(c.isomeric_smiles)
print(c.molecular_formula)
print(c.molecular_weight)
print(c.exact_mass)
print(c.monoisotopic_mass)
print(c.xlogp)
print(c.tpsa)
print(c.complexity)
print(c.charge)
print(c.h_bond_donor_count)
print(c.h_bond_acceptor_count)
print(c.rotatable_bond_count)
print(c.heavy_atom_count)
print(c.isotope_atom_count)
print(c.atom_stereo_count)
print(c.bond_stereo_count)
print(c.covalent_unit_count)
# # WARNING: TAKES VERY LONG TO RUN! Do not recommend if you don't have 1-2 hours to wait around!!
# now get pubchem data- for real, for each of the unique drugs
unique_drugs = [str(x) for x in unique_drugs]
drug_dict = {}
for curr_drug in unique_drugs:
print(curr_drug)
try: # need try catch block for any drug not in the dataframe
compound = pubchempy.get_compounds(curr_drug, 'name')
if (compound != []):
compound = compound[0]
cid = compound.cid
c = pubchempy.Compound.from_cid(cid)
frame = pubchempy.compounds_to_frame(c, properties=['molecular_weight', 'exact_mass', 'monoisotopic_mass', 'xlogp', 'tpsa', 'complexity', 'charge',
'h_bond_donor_count','h_bond_acceptor_count','rotatable_bond_count','heavy_atom_count','isotope_atom_count',
'atom_stereo_count','bond_stereo_count','covalent_unit_count'])
drug_dict[curr_drug] = frame
except:
print('not found in dataframe')
# make dataframe out of the dictionary
drug_dict_df = pd.DataFrame.from_dict(drug_dict, orient="index")
# get shape of the dictionary
drug_dict_df.shape
# save the dataframe
drug_dict_df.to_csv('drug_dict_9_15.csv')
# let's see what the data looks like
for i in range(0, 1784):
drug_dict_df.iloc[i,0]
# have to parse the weird column of the frame because i saved it weird
processed_drug_dict = drug_dict_df.iloc[0:1784,0]
processed_drug_dict.to_csv('processed_dict.csv')
# and check out the head of the data
processed_drug_dict.head(5)
# flatten the dataframe saved for each drug into values in columns
drug_name_list = []
drug_vals = np.zeros((1784,15))
counter = 0
for key, value in drug_dict.items():
drug_name_list.append(key)
print(np.array(value))
drug_vals[counter] = np.array(value)
counter = counter + 1
# let's see the rownames of the dataframe
drug_name_list
# let's see the actual contents of the dataframe
drug_vals
# save the final list of drug names
pd.DataFrame(drug_name_list).to_csv('drug_name_list_final_9_15.csv')
# save the final list of drug properties
pd.DataFrame(drug_vals).to_csv('drug_properties_final_9_15.csv')
# notate the drug properties that we analyzed
val_names = ['atom_stereo_count', 'bond_stereo_count', 'charge', 'complexity', 'covalent_unit_count', 'exact_mass', 'h_bond_acceptor_count', 'h_bond_donor_count','heavy_atom_count','isotope_atom_count', 'molecular_weight', 'monoisotopic_mass', 'rotable_bond_count', 'tpsa', 'xlogp']
# save the drug properties to their own dataframe
pd.DataFrame(val_names).to_csv('val_names_9_15.csv')
| get_drug_data_pubchempy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''anaconda3'': conda)'
# metadata:
# interpreter:
# hash: 6c03733f887ef21fbe0f240902c4a619cd92f5ab72ba254b59a7f0b51984db09
# name: 'Python 3.7.7 64-bit (''anaconda3'': conda)'
# ---
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# -
dataset = pd.read_csv('../input/Flight_Data.csv')
dataset.head()
dataset.Airline.unique()
dataset.isna().sum()
dataset[dataset.Route.isna()]
# ### As we can see, there is a single row with both Route and Total_Stops as null.
# so we can drop it.
dataset.dropna(inplace=True)
dataset.isna().sum()
# +
# Converting all the date and time data to numbers
dataset["Journey_day"] = pd.to_datetime(dataset.Date_of_Journey, format="%d/%m/%Y").dt.day
dataset["Journey_month"] = pd.to_datetime(dataset["Date_of_Journey"], format = "%d/%m/%Y").dt.month
dataset.drop('Date_of_Journey', 1, inplace=True)
dataset["Dep_hour"] = pd.to_datetime(dataset["Dep_Time"]).dt.hour
dataset["Dep_min"] = pd.to_datetime(dataset["Dep_Time"]).dt.minute
dataset.drop(["Dep_Time"], axis = 1, inplace = True)
dataset["Arr_hour"] = pd.to_datetime(dataset["Arrival_Time"]).dt.hour
dataset["Arr_min"] = pd.to_datetime(dataset["Arrival_Time"]).dt.minute
dataset.drop(["Arrival_Time"], axis = 1, inplace = True)
# -
dataset
# +
duration = list(dataset['Duration'])
for i in range(len(duration)):
if len(duration[i].split()) != 2:
if "h" in duration[i]:
duration[i] = duration[i].strip() + " 0m"
else:
duration[i] = "0h " + duration[i]
duration_hours = []
duration_mins = []
for i in range(len(duration)):
duration_hours.append(int(duration[i].split(sep = "h")[0]))
duration_mins.append(int(duration[i].split(sep = "m")[0].split()[-1]))
# +
dataset["Duration_hrs"] = duration_hours
dataset["Duration_mins"] = duration_mins
dataset.drop('Duration', 1, inplace=True)
# -
dataset
dataset = pd.concat([dataset, pd.get_dummies(dataset[['Airline', 'Source', 'Destination']],
drop_first = True
)], axis=1)
dataset.drop(['Airline', 'Source', 'Destination'], 1, inplace=True)
# drop route and total_steps
dataset.drop(['Route', 'Additional_Info'], 1, inplace=True)
dataset
dataset.Total_Stops.unique()
stops_mapping = {
'non-stop': 0,
'1 stop': 1,
'2 stops': 2,
'3 stops': 3,
'4 stops': 4,
}
dataset['Stops'] = dataset.Total_Stops.map(stops_mapping)
dataset.drop('Total_Stops', 1, inplace=True)
| notebooks/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Set partitioning
#
# Given a list of numbers, partition the values into two sets of equal sum.
values = [1, 2, 3, 4, 5, 6, 7, 8]
# # Create the BQM object
#
# - Use one binary variable $x_i$ for each value $v_i$. If $x_i$ = 1, the value $v_i$ belongs to one set (call it Set1), otherwise it belongs to the other set (Set2).
# - There is no objective, only a constraint: the sum of the values in each set must be equal
#
# $$ \sum_i v_i x_i = \sum_i v_i (1 - x_i) $$
#
# After simplifying:
#
# $$ \sum_i 2 v_i x_i - \sum_i v_i = 0$$
# +
from dimod import BinaryQuadraticModel
bqm = BinaryQuadraticModel('BINARY')
n = len(values)
x = {i: bqm.add_variable(f'x_{i}') for i in range(n)}
bqm.add_linear_equality_constraint(
[(x[i], 2.0 * values[i]) for i in range(n)],
constant=-sum(values),
lagrange_multiplier=10
)
# +
from dimod import ExactSolver
response = ExactSolver().sample(bqm).truncate(5)
solution = response.first.sample
print(response)
# -
set1 = {values[i] for i in x if solution[x[i]]}
set2 = {values[i] for i in x if not solution[x[i]]}
print(f'{sum(set1)} = sum{tuple(set1)}')
print(f'{sum(set2)} = sum{tuple(set2)}')
# # Partitioning to more than two sets
#
# - We will need one binary variable for each number and set combination
# - The binary value xij = 1 if value i belongs to set j
# - Each value can only be assigned to one set
# +
values = [7, 2, 3, 1, 8, 3, 1, 2, 9]
bqm = BinaryQuadraticModel('BINARY')
n = len(values)
m = 3 # num_partitions
x = {(i, k): bqm.add_variable((f'x_{i}', k))
for i in range(n)
for k in range(m)
}
# -
# # No objective, only constraints
# For each pair of sets, ensure that the sum of the values in one set is equal to the sum of the values in the other set
#
# $$ \sum_i v_i x_{ij} = \sum_i v_i x_{ik} $$ for all j and k
#
# Or equally:
#
# $$ \sum_i v_i x_{ij} - \sum_i v_i x_{ik} = 0$$
from itertools import combinations
for k, l in combinations(range(m), r=2):
bqm.add_linear_equality_constraint(
[(x[i, k], values[i]) for i in range(n)] + [(x[i, l], -values[i]) for i in range(n)],
constant=0,
lagrange_multiplier=10)
# Add a constraint to make sure each value is assign to exactly one set
for i in range(n):
bqm.add_linear_equality_constraint(
[(x[i, k], 1.0) for k in range(m)],
constant=-1.0,
lagrange_multiplier=10)
# Solve using one of the solvers. You may have to run it a few times.
# +
from neal import SimulatedAnnealingSampler
res = SimulatedAnnealingSampler().sample(bqm, num_reads=100, num_sweeps=1000).truncate(5)
print(res)
# -
# # Result
# +
sample = res.first.sample
print(sum(values))
for k in range(m):
set1 = [values[i] for (i, l) in x if sample[x[i, l]] if k == l]
print(sum(set1), set1)
| part04_example3_number_partitioning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.012534, "end_time": "2022-01-28T13:26:49.478482", "exception": false, "start_time": "2022-01-28T13:26:49.465948", "status": "completed"} tags=[]
# ## cloudFPGA TRIERES example
# ### Case study: Give an input string to FPGA and get the upper-case string from FPGA
# ### You don't need FPGA knowledge, just basic Python syntax !!!
# + [markdown] papermill={"duration": 0.014214, "end_time": "2022-01-28T13:26:49.504535", "exception": false, "start_time": "2022-01-28T13:26:49.490321", "status": "completed"} tags=[]
# 
#
# Assuming that the FPGA is already flashed
# + [markdown] papermill={"duration": 0.007696, "end_time": "2022-01-28T13:26:49.522837", "exception": false, "start_time": "2022-01-28T13:26:49.515141", "status": "completed"} tags=[]
# Configure the Python path to look for FPGA aceleration library
# + papermill={"duration": 0.01505, "end_time": "2022-01-28T13:26:49.545953", "exception": false, "start_time": "2022-01-28T13:26:49.530903", "status": "completed"} tags=[]
import time
import sys
import os
# + papermill={"duration": 0.013514, "end_time": "2022-01-28T13:26:49.565785", "exception": false, "start_time": "2022-01-28T13:26:49.552271", "status": "completed"} tags=[]
trieres_lib=os.environ['cFpRootDir'] + "/HOST/custom/uppercase/languages/python/build"
sys.path.append(trieres_lib)
# + [markdown] papermill={"duration": 0.009598, "end_time": "2022-01-28T13:26:49.584311", "exception": false, "start_time": "2022-01-28T13:26:49.574713", "status": "completed"} tags=[]
# Import the FPGA accelerator library
# + papermill={"duration": 0.014257, "end_time": "2022-01-28T13:26:49.605887", "exception": false, "start_time": "2022-01-28T13:26:49.591630", "status": "completed"} tags=[]
import _trieres
# + papermill={"duration": 0.013967, "end_time": "2022-01-28T13:26:49.629352", "exception": false, "start_time": "2022-01-28T13:26:49.615385", "status": "completed"} tags=[]
input = "HelloWorld"
# + [markdown] papermill={"duration": 0.007099, "end_time": "2022-01-28T13:26:49.644030", "exception": false, "start_time": "2022-01-28T13:26:49.636931", "status": "completed"} tags=[]
# Currently we reserve space on host for output (can also be done in library)
# + papermill={"duration": 0.010294, "end_time": "2022-01-28T13:26:49.660926", "exception": false, "start_time": "2022-01-28T13:26:49.650632", "status": "completed"} tags=[]
fpga_output = "1111111111"
# + [markdown] papermill={"duration": 0.006166, "end_time": "2022-01-28T13:26:49.673681", "exception": false, "start_time": "2022-01-28T13:26:49.667515", "status": "completed"} tags=[]
# Execute the FPGA accelerator as a Python function
# + papermill={"duration": 0.011343, "end_time": "2022-01-28T13:26:49.691298", "exception": false, "start_time": "2022-01-28T13:26:49.679955", "status": "completed"} tags=[]
#fpga_ip=os.environ['FPGA_IP']
fpga_ip="localhost"
print(fpga_ip)
# + papermill={"duration": 0.25582, "end_time": "2022-01-28T13:26:49.954451", "exception": false, "start_time": "2022-01-28T13:26:49.698631", "status": "completed"} tags=[]
start_fpga = time.time()
out, fpga_output = _trieres.uppercase(fpga_ip, "2718", input, True)
done_fpga = time.time()
elapsed_fpga = done_fpga - start_fpga
# + papermill={"duration": 0.022061, "end_time": "2022-01-28T13:26:49.988214", "exception": false, "start_time": "2022-01-28T13:26:49.966153", "status": "completed"} tags=[]
print("Output from FPGA : "+fpga_output)
# + papermill={"duration": 0.022207, "end_time": "2022-01-28T13:26:50.020555", "exception": false, "start_time": "2022-01-28T13:26:49.998348", "status": "completed"} tags=[]
start_cpu = time.time()
cpu_output=input.upper()
done_cpu = time.time()
print("Output from CPU : "+cpu_output)
elapsed_cpu = done_cpu - start_cpu
# + papermill={"duration": 0.016638, "end_time": "2022-01-28T13:26:50.053930", "exception": false, "start_time": "2022-01-28T13:26:50.037292", "status": "completed"} tags=[]
print("FPGA time = "+'{0:.10f}'.format(elapsed_fpga)+"\nCPU time = "+'{0:.10f}'.format(elapsed_cpu))
| HOST/custom/uppercase/languages/python/trieres_uppercase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="rX8mhOLljYeM"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="BZSlp3DAjdYf" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + colab_type="code" id="RXZT2UsyIVe_" colab={}
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/horse-or-human.zip \
# -O /tmp/horse-or-human.zip
# + [markdown] id="9brUxyTpYZHy" colab_type="text"
# The following python code will use the OS library to use Operating System libraries, giving you access to the file system, and the zipfile library allowing you to unzip the data.
# + colab_type="code" id="PLy3pthUS0D2" colab={}
import os
import zipfile
local_zip = '/tmp/horse-or-human.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp/horse-or-human')
zip_ref.close()
# + [markdown] colab_type="text" id="o-qUPyfO7Qr8"
# The contents of the .zip are extracted to the base directory `/tmp/horse-or-human`, which in turn each contain `horses` and `humans` subdirectories.
#
# In short: The training set is the data that is used to tell the neural network model that 'this is what a horse looks like', 'this is what a human looks like' etc.
#
# One thing to pay attention to in this sample: We do not explicitly label the images as horses or humans. If you remember with the handwriting example earlier, we had labelled 'this is a 1', 'this is a 7' etc. Later you'll see something called an ImageGenerator being used -- and this is coded to read images from subdirectories, and automatically label them from the name of that subdirectory. So, for example, you will have a 'training' directory containing a 'horses' directory and a 'humans' one. ImageGenerator will label the images appropriately for you, reducing a coding step.
#
# Let's define each of these directories:
# + colab_type="code" id="NR_M9nWN-K8B" colab={}
# Directory with our training horse pictures
train_horse_dir = os.path.join('/tmp/horse-or-human/horses')
# Directory with our training human pictures
train_human_dir = os.path.join('/tmp/horse-or-human/humans')
# + [markdown] colab_type="text" id="LuBYtA_Zd8_T"
# Now, let's see what the filenames look like in the `horses` and `humans` training directories:
# + colab_type="code" id="4PIP1rkmeAYS" colab={}
train_horse_names = os.listdir(train_horse_dir)
print(train_horse_names[:10])
train_human_names = os.listdir(train_human_dir)
print(train_human_names[:10])
# + [markdown] colab_type="text" id="HlqN5KbafhLI"
# Let's find out the total number of horse and human images in the directories:
# + colab_type="code" id="H4XHh2xSfgie" colab={}
print('total training horse images:', len(os.listdir(train_horse_dir)))
print('total training human images:', len(os.listdir(train_human_dir)))
# + [markdown] colab_type="text" id="C3WZABE9eX-8"
# Now let's take a look at a few pictures to get a better sense of what they look like. First, configure the matplot parameters:
# + colab_type="code" id="b2_Q0-_5UAv-" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# Parameters for our graph; we'll output images in a 4x4 configuration
nrows = 4
ncols = 4
# Index for iterating over images
pic_index = 0
# + [markdown] colab_type="text" id="xTvHzGCxXkqp"
# Now, display a batch of 8 horse and 8 human pictures. You can rerun the cell to see a fresh batch each time:
# + colab_type="code" id="Wpr8GxjOU8in" colab={}
# Set up matplotlib fig, and size it to fit 4x4 pics
fig = plt.gcf()
fig.set_size_inches(ncols * 4, nrows * 4)
pic_index += 8
next_horse_pix = [os.path.join(train_horse_dir, fname)
for fname in train_horse_names[pic_index-8:pic_index]]
next_human_pix = [os.path.join(train_human_dir, fname)
for fname in train_human_names[pic_index-8:pic_index]]
for i, img_path in enumerate(next_horse_pix+next_human_pix):
# Set up subplot; subplot indices start at 1
sp = plt.subplot(nrows, ncols, i + 1)
sp.axis('Off') # Don't show axes (or gridlines)
img = mpimg.imread(img_path)
plt.imshow(img)
plt.show()
# + [markdown] colab_type="text" id="5oqBkNBJmtUv"
# ## Building a Small Model from Scratch
#
# But before we continue, let's start defining the model:
#
# Step 1 will be to import tensorflow.
# + id="qvfZg3LQbD-5" colab_type="code" colab={}
import tensorflow as tf
# + [markdown] colab_type="text" id="BnhYCP4tdqjC"
# We then add convolutional layers as in the previous example, and flatten the final result to feed into the densely connected layers.
# + [markdown] id="gokG5HKpdtzm" colab_type="text"
# Finally we add the densely connected layers.
#
# Note that because we are facing a two-class classification problem, i.e. a *binary classification problem*, we will end our network with a [*sigmoid* activation](https://wikipedia.org/wiki/Sigmoid_function), so that the output of our network will be a single scalar between 0 and 1, encoding the probability that the current image is class 1 (as opposed to class 0).
# + id="PixZ2s5QbYQ3" colab_type="code" colab={}
model = tf.keras.models.Sequential([
# Note the input shape is the desired size of the image 300x300 with 3 bytes color
# This is the first convolution
tf.keras.layers.Conv2D(16, (3,3), activation='relu', input_shape=(300, 300, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
# The second convolution
tf.keras.layers.Conv2D(32, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The third convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The fourth convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The fifth convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# Flatten the results to feed into a DNN
tf.keras.layers.Flatten(),
# 512 neuron hidden layer
tf.keras.layers.Dense(512, activation='relu'),
# Only 1 output neuron. It will contain a value from 0-1 where 0 for 1 class ('horses') and 1 for the other ('humans')
tf.keras.layers.Dense(1, activation='sigmoid')
])
# + [markdown] colab_type="text" id="s9EaFDP5srBa"
# The model.summary() method call prints a summary of the NN
# + colab_type="code" id="7ZKj8392nbgP" colab={}
model.summary()
# + [markdown] colab_type="text" id="DmtkTn06pKxF"
# The "output shape" column shows how the size of your feature map evolves in each successive layer. The convolution layers reduce the size of the feature maps by a bit due to padding, and each pooling layer halves the dimensions.
# + [markdown] colab_type="text" id="PEkKSpZlvJXA"
# Next, we'll configure the specifications for model training. We will train our model with the `binary_crossentropy` loss, because it's a binary classification problem and our final activation is a sigmoid. (For a refresher on loss metrics, see the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/descending-into-ml/video-lecture).) We will use the `rmsprop` optimizer with a learning rate of `0.001`. During training, we will want to monitor classification accuracy.
#
# **NOTE**: In this case, using the [RMSprop optimization algorithm](https://wikipedia.org/wiki/Stochastic_gradient_descent#RMSProp) is preferable to [stochastic gradient descent](https://developers.google.com/machine-learning/glossary/#SGD) (SGD), because RMSprop automates learning-rate tuning for us. (Other optimizers, such as [Adam](https://wikipedia.org/wiki/Stochastic_gradient_descent#Adam) and [Adagrad](https://developers.google.com/machine-learning/glossary/#AdaGrad), also automatically adapt the learning rate during training, and would work equally well here.)
# + colab_type="code" id="8DHWhFP_uhq3" colab={}
from tensorflow.keras.optimizers import RMSprop
model.compile(loss='binary_crossentropy',
optimizer=RMSprop(lr=0.001),
metrics=['acc'])
# + [markdown] colab_type="text" id="Sn9m9D3UimHM"
# ### Data Preprocessing
#
# Let's set up data generators that will read pictures in our source folders, convert them to `float32` tensors, and feed them (with their labels) to our network. We'll have one generator for the training images and one for the validation images. Our generators will yield batches of images of size 300x300 and their labels (binary).
#
# As you may already know, data that goes into neural networks should usually be normalized in some way to make it more amenable to processing by the network. (It is uncommon to feed raw pixels into a convnet.) In our case, we will preprocess our images by normalizing the pixel values to be in the `[0, 1]` range (originally all values are in the `[0, 255]` range).
#
# In Keras this can be done via the `keras.preprocessing.image.ImageDataGenerator` class using the `rescale` parameter. This `ImageDataGenerator` class allows you to instantiate generators of augmented image batches (and their labels) via `.flow(data, labels)` or `.flow_from_directory(directory)`. These generators can then be used with the Keras model methods that accept data generators as inputs: `fit_generator`, `evaluate_generator`, and `predict_generator`.
# + colab_type="code" id="ClebU9NJg99G" colab={}
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# All images will be rescaled by 1./255
train_datagen = ImageDataGenerator(rescale=1/255)
# Flow training images in batches of 128 using train_datagen generator
train_generator = train_datagen.flow_from_directory(
'/tmp/horse-or-human/', # This is the source directory for training images
target_size=(300, 300), # All images will be resized to 150x150
batch_size=128,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
# + [markdown] colab_type="text" id="mu3Jdwkjwax4"
# ### Training
# Let's train for 15 epochs -- this may take a few minutes to run.
#
# Do note the values per epoch.
#
# The Loss and Accuracy are a great indication of progress of training. It's making a guess as to the classification of the training data, and then measuring it against the known label, calculating the result. Accuracy is the portion of correct guesses.
# + colab_type="code" id="Fb1_lgobv81m" colab={}
history = model.fit_generator(
train_generator,
steps_per_epoch=8,
epochs=15,
verbose=1)
# + [markdown] id="o6vSHzPR2ghH" colab_type="text"
# ###Running the Model
#
# Let's now take a look at actually running a prediction using the model. This code will allow you to choose 1 or more files from your file system, it will then upload them, and run them through the model, giving an indication of whether the object is a horse or a human.
# + id="DoWp43WxJDNT" colab_type="code" colab={}
import numpy as np
from google.colab import files
from keras.preprocessing import image
uploaded = files.upload()
for fn in uploaded.keys():
# predicting images
path = '/content/' + fn
img = image.load_img(path, target_size=(300, 300))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
images = np.vstack([x])
classes = model.predict(images, batch_size=10)
print(classes[0])
if classes[0]>0.5:
print(fn + " is a human")
else:
print(fn + " is a horse")
# + [markdown] colab_type="text" id="-8EHQyWGDvWz"
# ### Visualizing Intermediate Representations
#
# To get a feel for what kind of features our convnet has learned, one fun thing to do is to visualize how an input gets transformed as it goes through the convnet.
#
# Let's pick a random image from the training set, and then generate a figure where each row is the output of a layer, and each image in the row is a specific filter in that output feature map. Rerun this cell to generate intermediate representations for a variety of training images.
# + colab_type="code" id="-5tES8rXFjux" colab={}
import numpy as np
import random
from tensorflow.keras.preprocessing.image import img_to_array, load_img
# Let's define a new Model that will take an image as input, and will output
# intermediate representations for all layers in the previous model after
# the first.
successive_outputs = [layer.output for layer in model.layers[1:]]
#visualization_model = Model(img_input, successive_outputs)
visualization_model = tf.keras.models.Model(inputs = model.input, outputs = successive_outputs)
# Let's prepare a random input image from the training set.
horse_img_files = [os.path.join(train_horse_dir, f) for f in train_horse_names]
human_img_files = [os.path.join(train_human_dir, f) for f in train_human_names]
img_path = random.choice(horse_img_files + human_img_files)
img = load_img(img_path, target_size=(300, 300)) # this is a PIL image
x = img_to_array(img) # Numpy array with shape (150, 150, 3)
x = x.reshape((1,) + x.shape) # Numpy array with shape (1, 150, 150, 3)
# Rescale by 1/255
x /= 255
# Let's run our image through our network, thus obtaining all
# intermediate representations for this image.
successive_feature_maps = visualization_model.predict(x)
# These are the names of the layers, so can have them as part of our plot
layer_names = [layer.name for layer in model.layers]
# Now let's display our representations
for layer_name, feature_map in zip(layer_names, successive_feature_maps):
if len(feature_map.shape) == 4:
# Just do this for the conv / maxpool layers, not the fully-connected layers
n_features = feature_map.shape[-1] # number of features in feature map
# The feature map has shape (1, size, size, n_features)
size = feature_map.shape[1]
# We will tile our images in this matrix
display_grid = np.zeros((size, size * n_features))
for i in range(n_features):
# Postprocess the feature to make it visually palatable
x = feature_map[0, :, :, i]
x -= x.mean()
x /= x.std()
x *= 64
x += 128
x = np.clip(x, 0, 255).astype('uint8')
# We'll tile each filter into this big horizontal grid
display_grid[:, i * size : (i + 1) * size] = x
# Display the grid
scale = 20. / n_features
plt.figure(figsize=(scale * n_features, scale))
plt.title(layer_name)
plt.grid(False)
plt.imshow(display_grid, aspect='auto', cmap='viridis')
# + [markdown] colab_type="text" id="tuqK2arJL0wo"
# As you can see we go from the raw pixels of the images to increasingly abstract and compact representations. The representations downstream start highlighting what the network pays attention to, and they show fewer and fewer features being "activated"; most are set to zero. This is called "sparsity." Representation sparsity is a key feature of deep learning.
#
#
# These representations carry increasingly less information about the original pixels of the image, but increasingly refined information about the class of the image. You can think of a convnet (or a deep network in general) as an information distillation pipeline.
# + [markdown] colab_type="text" id="j4IBgYCYooGD"
# ## Clean Up
#
# Before running the next exercise, run the following cell to terminate the kernel and free memory resources:
# + colab_type="code" id="651IgjLyo-Jx" colab={}
import os, signal
os.kill(os.getpid(), signal.SIGKILL)
| Tensorflow/Course 1 - Part 8 - Lesson 2 - Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
import loader_celebA as loader
import os
from glob import glob
import numpy as np
from matplotlib import pyplot
import tensorflow as tf
# %matplotlib inline
# +
# Let's download the dataset from http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html under the link "Align&Cropped Images
# alternatively You can use https://www.dropbox.com/sh/8oqt9vytwxb3s4r/AADIKlz8PR9zr6Y20qbkunrba/Img/img_align_celeba.zip?dl=1&pv=1
# This function unzips the data and kreep all the images in celebA folder
loader.download_celeb_a()
# Let us explore the images
data_dir = os.getcwd()
test_images = loader.get_batch(glob(os.path.join(data_dir, 'celebA/*.jpg'))[:10], 56, 56)
pyplot.imshow(loader.plot_images(test_images))
# -
def discriminator(images, reuse=False):
"""
Create the discriminator network
"""
alpha = 0.2
with tf.variable_scope('discriminator', reuse=reuse):
# using 4 layer network as in DCGAN Paper
# First convolution layer
conv1 = tf.layers.conv2d(images, 64, 5, 2, 'SAME')
lrelu1 = tf.maximum(alpha * conv1, conv1)
# Second convolution layer
conv2 = tf.layers.conv2d(lrelu1, 128, 5, 2, 'SAME')
batch_norm2 = tf.layers.batch_normalization(conv2, training=True)
lrelu2 = tf.maximum(alpha * batch_norm2, batch_norm2)
# Third convolution layer
conv3 = tf.layers.conv2d(lrelu2, 256, 5, 1, 'SAME')
batch_norm3 = tf.layers.batch_normalization(conv3, training=True)
lrelu3 = tf.maximum(alpha * batch_norm3, batch_norm3)
# Flatten layer
flat = tf.reshape(lrelu3, (-1, 4*4*256))
# Logits
logits = tf.layers.dense(flat, 1)
# Output
out = tf.sigmoid(logits)
return out, logits
def generator(z, out_channel_dim, is_train=True):
"""
Create the generator network
"""
alpha = 0.2
with tf.variable_scope('generator', reuse=False if is_train==True else True):
# First fully connected layer
x_1 = tf.layers.dense(z, 2*2*512)
# Reshape it to start the convolutional stack
deconv_2 = tf.reshape(x_1, (-1, 2, 2, 512))
batch_norm2 = tf.layers.batch_normalization(deconv_2, training=is_train)
lrelu2 = tf.maximum(alpha * batch_norm2, batch_norm2)
# Deconv 1
deconv3 = tf.layers.conv2d_transpose(lrelu2, 256, 5, 2, padding='VALID')
batch_norm3 = tf.layers.batch_normalization(deconv3, training=is_train)
lrelu3 = tf.maximum(alpha * batch_norm3, batch_norm3)
# Deconv 2
deconv4 = tf.layers.conv2d_transpose(lrelu3, 128, 5, 2, padding='SAME')
batch_norm4 = tf.layers.batch_normalization(deconv4, training=is_train)
lrelu4 = tf.maximum(alpha * batch_norm4, batch_norm4)
# Output layer
logits = tf.layers.conv2d_transpose(lrelu4, out_channel_dim, 5, 2, padding='SAME')
out = tf.tanh(logits)
return out
def model_loss(input_real, input_z, out_channel_dim):
"""
Get the loss for the discriminator and generator
"""
label_smoothing = 0.9
g_model = generator(input_z, out_channel_dim)
d_model_real, d_logits_real = discriminator(input_real)
d_model_fake, d_logits_fake = discriminator(g_model, reuse=True)
d_loss_real = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_real,
labels=tf.ones_like(d_model_real) * label_smoothing))
d_loss_fake = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_fake,
labels=tf.zeros_like(d_model_fake)))
d_loss = d_loss_real + d_loss_fake
g_loss = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_fake,
labels=tf.ones_like(d_model_fake) * label_smoothing))
return d_loss, g_loss
def model_opt(d_loss, g_loss, learning_rate, beta1):
"""
Get optimization operations
"""
t_vars = tf.trainable_variables()
d_vars = [var for var in t_vars if var.name.startswith('discriminator')]
g_vars = [var for var in t_vars if var.name.startswith('generator')]
# Optimize
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
d_train_opt = tf.train.AdamOptimizer(learning_rate, beta1=beta1).minimize(d_loss, var_list=d_vars)
g_train_opt = tf.train.AdamOptimizer(learning_rate, beta1=beta1).minimize(g_loss, var_list=g_vars)
return d_train_opt, g_train_opt
def generator_output(sess, n_images, input_z, out_channel_dim):
"""
Show example output for the generator
"""
z_dim = input_z.get_shape().as_list()[-1]
example_z = np.random.uniform(-1, 1, size=[n_images, z_dim])
samples = sess.run(
generator(input_z, out_channel_dim, False),
feed_dict={input_z: example_z})
pyplot.imshow(loader.plot_images(samples))
pyplot.show()
def train(epoch_count, batch_size, z_dim, learning_rate, beta1, get_batches, data_shape, data_files):
"""
Train the GAN
"""
w, h, num_ch = data_shape[1], data_shape[2], data_shape[3]
X = tf.placeholder(tf.float32, shape=(None, w, h, num_ch), name='input_real')
Z = tf.placeholder(tf.float32, (None, z_dim), name='input_z')
#model_inputs(data_shape[1], data_shape[2], data_shape[3], z_dim)
D_loss, G_loss = model_loss(X, Z, data_shape[3])
D_solve, G_solve = model_opt(D_loss, G_loss, learning_rate, beta1)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
train_loss_d = []
train_loss_g = []
for epoch_i in range(epoch_count):
num_batch = 0
lossD, lossG = 0,0
for batch_images in get_batches(batch_size, data_shape, data_files):
# values range from -0.5 to 0.5 so we scale to range -1, 1
batch_images = batch_images * 2
num_batch += 1
batch_z = np.random.uniform(-1, 1, size=(batch_size, z_dim))
_,d_loss = sess.run([D_solve,D_loss], feed_dict={X: batch_images, Z: batch_z})
_,g_loss = sess.run([G_solve,G_loss], feed_dict={X: batch_images, Z: batch_z})
lossD += (d_loss/batch_size)
lossG += (g_loss/batch_size)
if num_batch % 500 == 0:
# After every 500 batches
print("Epoch {}/{} For Batch {} Discriminator Loss: {:.4f} Generator Loss: {:.4f}".
format(epoch_i+1, epochs, num_batch, lossD/num_batch, lossG/num_batch))
generator_output(sess, 9, Z, data_shape[3])
train_loss_d.append(lossD/num_batch)
train_loss_g.append(lossG/num_batch)
return train_loss_d, train_loss_g
# +
# Data Parameters
IMAGE_HEIGHT = 28
IMAGE_WIDTH = 28
data_files = glob(os.path.join(data_dir, 'celebA/*.jpg'))
#Hyper parameters
batch_size = 16
z_dim = 100
learning_rate = 0.0002
beta1 = 0.5
epochs = 2
shape = len(data_files), IMAGE_WIDTH, IMAGE_HEIGHT, 3
with tf.Graph().as_default():
Loss_D, Loss_G = train(epochs, batch_size, z_dim, learning_rate, beta1, loader.get_batches, shape, data_files)
# -
pyplot.plot(Loss_D, label = 'Discriminator Loss')
pyplot.plot(Loss_G, label = 'Generator Loss')
pyplot.legend()
pyplot.xlabel('epochs')
pyplot.ylabel('Loss')
| Chapter07/DCGAN_CelebA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Freya-LR/Leetcode/blob/main/35.%20Search%20Insert%20Position.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PJp5ZXsd3oKY"
# Array
# hint: Tme complexity is O(log n),
# if the target in nums, return the index, if not, add target into nums and resort, then return index
# but this method time complexity O(N), might not meet the reqirement runtime complexity O(log n)
class Solution:
def searchInsert(self, nums: List[int], target: int) -> int:
if target in nums:
return nums.index(target)
else:
nums.append(target)
nums.sort()
return nums.index(target)
# + colab={"base_uri": "https://localhost:8080/"} id="ygoa0KCfK22Z" outputId="f92762e3-330e-4d5e-cd15-768136566beb"
#
def searchInsert(nums,target):
if target in nums:
return nums.index(target)
right=len(nums)-1
left=0
if nums[right]<target:
return right+1
if nums[left]>target:
return left
while left < right:
middle=int(abs((right+left)/2))
if nums[middle] > target: # target in left half list
right = middle
else:
left = middle+1
return right
nums=[3,5,6,7,9,11]
target = 8
searchInsert(nums,target)
| 35. Search Insert Position.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (Nopimal)
# language: python
# name: pycharm-28004bf6
# ---
# + pycharm={"name": "#%%\n"}
# 读取数据
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
data=pd.read_csv('./data/train_tag.csv',encoding='gbk')
data.drop_duplicates(inplace=True)
# + pycharm={"name": "#%%\n"}
# 删除无关特征
drop_list=[]
for i in data.columns:
count=data[i].count()
if len(list(data[i].unique())) in [1,count,count-1]:
drop_list.append(i)
print(drop_list)
data.drop(drop_list,axis=1,inplace=True)
# + pycharm={"name": "#%%\n"}
# 了解数据整体情况
data.info()
# + pycharm={"name": "#%%\n"}
# 分析数值型数据缺失情况
# % matplotlib inline
data_num=data.select_dtypes('number').copy()
data_num_miss_rate=1-(data_num.count()/len(data_num))
data_num_miss_rate.sort_values(ascending=False,inplace=True)
print(data_num_miss_rate[:20])
data_num_miss_rate.plot()
# -
# 画图了解缺失情况
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['font.family']=['SimHei']
plt.rcParams['axes.unicode_minus']=False
data_num=data.select_dtypes('number').copy()
data_num_miss_rate=1-(data_num.count()/len(data_num))
data_num_miss_rate.sort_values(ascending=False,inplace=True)
fig,ax1=plt.subplots(figsize=(10,6))
sns.barplot([1,2,3,4,5,6,7,8,9,10],data_num_miss_rate[:10].values,ax=ax1)
ax1.set_title('特征缺失情况')
ax1.set_xlabel('缺失特征排名')
ax1.set_ylabel('缺失占比')
# + pycharm={"name": "#%%\n"}
# 数据处理
data_str=data.select_dtypes(exclude='number').copy()
data_str.describe()
data_str['reg_preference_for_trad'].fillna(data_str['reg_preference_for_trad'].mode()[0],inplace=True)
dic={}
for i,val in enumerate(list(data_str['reg_preference_for_trad'].unique())):
dic[val]=i
data_str['reg_preference_for_trad']=data_str['reg_preference_for_trad'].map(dic)
data_str['latest_query_time_month']=pd.to_datetime(data_str['latest_query_time']).dt.month
data_str['latest_query_time_weekday']=pd.to_datetime(data_str['latest_query_time']).dt.weekday
data_str['loans_latest_time_month']=pd.to_datetime(data_str['loans_latest_time']).dt.month
data_str['loans_latest_time_weekday']=pd.to_datetime(data_str['loans_latest_time']).dt.weekday
data_str.drop(['latest_query_time','loans_latest_time'],axis=1,inplace=True)
for i in data_str.columns:
data_str[i].fillna(data_str[i].mode()[0],inplace=True)
# + pycharm={"name": "#%%\n"}
# 划分训练集测试集
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
features=[x for x in data_all.columns if x not in ['status']]
X=data_all[features]
y=data_all['status']
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3)
# + pycharm={"name": "#%%\n"}
# 特征归一化
std=StandardScaler()
X_train=std.fit_transform(X_train)
X_test=std.fit_transform(X_test)
# + pycharm={"name": "#%%\n"}
# 模型评估
from sklearn.metrics import accuracy_score,precision_score,recall_score,f1_score
from sklearn.metrics import roc_auc_score,roc_curve,auc
import matplotlib.pyplot as plt
def model_metrics(clf,X_train,X_test,y_train,y_test):
# 预测
y_train_pred=clf.predict(X_train)
y_test_pred=clf.predict(X_test)
y_train_proba=clf.predict_proba(X_train)[:,1]
y_test_proba=clf.predict_proba(X_test)[:,1]
# 准确率
print('[准确率]',end='')
print('训练集:{:.4f}'.format(accuracy_score(y_train,y_train_pred)),end='')
print('测试集:{:.4f}'.format(accuracy_score(y_test,y_test_pred)),end='')
# 精准率
print('[精准率]',end='')
print('训练集:{:.4f}'.format(precision_score(y_train,y_train_pred)),end='')
print('测试集:{:.4f}'.format(precision_score(y_test,y_test_pred)),end='')
# 召回率
print('[召回率]',end='')
print('训练集:{:.4f}'.format(recall_score(y_train,y_train_pred)),end='')
print('测试集:{:.4f}'.format(recall_score(y_test,y_test_pred)),end='')
# f1-score
print('[f1-score]',end='')
print('训练集:{:.4f}'.format(f1_score(y_train,y_train_pred)),end='')
print('测试集:{:.4f}'.format(f1_score(y_test,y_test_pred)),end='')
# auc取值:用roc_auc_score或auc
print('[auc值]',end='')
print('训练集:{:.4f}'.format(roc_auc_score(y_train,y_train_proba)),end='')
print('测试集:{:.4f}'.format(roc_auc_score(y_test,y_test_proba)),end='')
# roc曲线
fpr_train,tpr_train,thresholds_train=roc_curve(y_train,y_train_proba,pos_label=1)
fpr_test,tpr_test,thresholds_test=roc_curve(y_test,y_test_proba,pos_label=1)
label=['Train - AUC:{:.4f'.format(auc(fpr_train,tpr_train)),
'Test - AUC:{:.4f'.format(auc(fpr_test,tpr_test))]
plt.plot(fpr_train,tpr_train)
plt.plot(fpr_test,tpr_test)
plt.plot([0,1],[0,1],'d--')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(label,loc=4)
plt.title('ROC curve')
# + pycharm={"name": "#%%\n"}
# 模型
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn import svm
from sklearn.tree import DecisionTreeClassifier
from lightgbm.sklearn import LGBMClassifier
from sklearn.metrics import confusion_matrix
from mlxtend.classifier import StackingClassifier
from sklearn.ensemble import RandomForestClassifier
plt.rcParams['font.family']=['SimHei']
plt.rcParams['axes.unicode_minus']=False
# rf
rf=RandomForestClassifier(random_state=2018)
param={'n_estimators':[40,60,800],'max_depth':[i for i in range(6,10)],
'criterion':['entropy'],'min_samples_split':[5,6,7,8]}
gsearch=GridSearchCV(rf,param_grid=param,scoring='roc_auc',cv=4)
gsearch.fit(X_train,y_train)
print('最佳参数: ',gsearch.best_params_)
print('训练集的最佳分数:',gsearch.best_score_)
print('测试集的最佳分数: ',gsearch.score(X_test,y_test))
rf=RandomForestClassifier(criterion='entropy',max_depth=9,min_samples_split=7,n_estimators=800)
rf.fit(X_train,y_train)
model_metrics(rf,X_train,X_test,y_train,y_test)
# svm_linear
svm_linear=svm.SVC(kernel='linear',probability=True).fit(X_train,y_train)
model_metrics(svm_linear,X_train,X_test,y_train,y_test)
# svm poly
svm_poly=svm.SVC(C=0.01,kernel='poly',probability=True).fit(X_train,y_train)
model_metrics(svm_poly,X_train,X_test,y_train,y_test)
# svm_rbf
svm_rbf=svm.SVC(kernel='rbf',probability=True,gamma=0.01,C=0.1)
svm_rbf.fit(X_train,y_train)
model_metrics(svm_rbf,X_train,X_test,y_train,y_test)
# svm_sigmoid
svm_sigmoid=svm.SVC(C=0.05,kernel='sigmoid',probability=True)
svm_sigmoid.fit(X_train,y_train)
model_metrics(svm_sigmoid,X_train,X_test,y_train,y_test)
# dt
dt=DecisionTreeClassifier(max_depth=9,min_samples_split=100,min_samples_leaf=90,max_features=9)
dt.fit(X_train,y_train)
model_metrics(dt,X_train,X_test,y_train,y_test)
# lr
lr=LogisticRegression(C=0.04,penalty='l1')
model_metrics(lr,X_train,X_test,y_train,y_test)
# lgb
lgb=LGBMClassifier(learning_rate=0.1,n_estimators=50,max_depth=3,
min_child_weight=7,gamma=0,subsample=0.5,colsample_bytree=0.8,
reg_alpha=1e-5,nthread=4,scale_pos_weight=1)
model_metrics(lgb,X_train,X_test,y_train,y_test)
# 模型融合
sclf_lr=StackingClassifier(classifiers=[lr,svm_linear,svm_rbf,rf,lgb],
meta_classifier=lr,
use_probas=True,
average_probas=True,
use_features_in_secondary=True)
sclf_lr.fit(X_train,y_train.values)
model_metrics(sclf_lr,X_train,X_test,y_train,y_test)
| code/feature/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import copy
from skimage.color import rgb2gray
import warnings
warnings.filterwarnings("ignore")
# -
def bilinear_interpolate_1(img, pixel):
"""
Bilinear Interpolation
Parameters:
img(matrix) - initial unchanged image
pixel(tuple or list of 2 elements)
Return:
float(interpolated value)
"""
H, W = img.shape
x = np.asarray(pixel[0])
y = np.asarray(pixel[1])
x_0 = np.floor(x).astype(int)
x_1 = x_0 + 1
y_0 = np.floor(y).astype(int)
y_1 = y_0 + 1
x_0 = np.clip(x_0, 0, H - 1)
x_1 = np.clip(x_1, 0, H - 1)
y_0 = np.clip(y_0, 0, W - 1)
y_1 = np.clip(y_1, 0, W - 1)
a_0 = img[ x_0, y_0 ]
a_1 = img[ x_0, y_1 ]
a_2 = img[ x_1, y_0 ]
a_3 = img[ x_1, y_1 ]
w_a_0 = (x - x_0) * (y - y_0)
w_a_1 = (x - x_0) * (y_1 - y)
w_a_2 = (x_1 - x) * (y - y_0)
w_a_3 = (x_1 - x) * (y_1 - y)
return w_a_0 * a_3 + w_a_1 * a_2 + w_a_2 * a_1 + w_a_3 * a_0
# return w_a_0 * a_0 + w_a_1 * a_1 + w_a_2 * a_2 + w_a_3 * a_3
def bilinear_interpolate_2(img, pixel):
"""
Bilinear Interpolation with 2 sequential 1 dimensional interpolation.
Parameters:
img(matrix) - initial unchanged image
pixel(tuple or list of 2 elements)
Return:
float(interpolated value)
"""
H, W = img.shape
x = np.asarray(pixel[0])
y = np.asarray(pixel[1])
x_0 = np.floor(x).astype(int)
x_1 = x_0 + 1
y_0 = np.floor(y).astype(int)
y_1 = y_0 + 1
x_0 = np.clip(x_0, 0, H - 1)
x_1 = np.clip(x_1, 0, H - 1)
y_0 = np.clip(y_0, 0, W - 1)
y_1 = np.clip(y_1, 0, W - 1)
f_q_1_1 = img[ x_0, y_0 ]
f_q_1_2 = img[ x_0, y_1 ]
f_q_2_1 = img[ x_1, y_0 ]
f_q_2_2 = img[ x_1, y_1 ]
p_x_y_1 = f_q_1_1 * (x-x_1) / (x_0-x_1) + f_q_2_1 * (x-x_0) / (x_1-x_0)
p_x_y_2 = f_q_1_2 * (x-x_1) / (x_0-x_1) + f_q_2_2 * (x-x_0) / (x_1-x_0)
return p_x_y_1 * (y-y_1) / (y_0-y_1) + p_x_y_2 * (y-y_0) / (y_1-y_0)
def Sampling(shape, i, j):
"""
Uniform Sampling
Parameters:
shape(list or tuple) - shape of the image we want sample from
i(int) - i-th index
j(int) - j-th index
Returns:
tuple(of floats) - sampled pixel
"""
H, W = shape
return (2*i+1)/(2*H), (2*j+1)/(2*W)
def Image_Resizing(initial_img, new_img_shape):
"""
Image Resizing
Parameters:
initial_img(matrix) - initial unchanged image
new_img_shape(list or tuple) - shape of the new image
Returns:
matrix(resized image)
"""
H, W = initial_img.shape
H_n, W_n = new_img_shape
new_image = np.zeros(new_img_shape, dtype=float)
for x in range(H_n):
for y in range(W_n):
pixel = Sampling(new_img_shape, x, y)
pixel = (pixel[0] * H, pixel[1] * W)
new_image[x, y] = bilinear_interpolate_1(initial_img, pixel)
return new_image
# +
img=mpimg.imread('inputs/20190926/2.jpg')
img = rgb2gray(img)
plt.subplot(1,2,1)
plt.title('Origin image')
plt.imshow(img)
plt.subplot(1,2,2)
plt.title('Resized image')
# new_image_shape = tuple([2*x for x in img.shape])
new_image_shape = [440, 300]
resized = Image_Resizing(img, new_image_shape)
plt.imshow(resized)
plt.show();
# +
plt.subplot(1,2,1)
plt.title('Origin image')
plt.imshow(img)
plt.subplot(1,2,2)
plt.title('Resized image')
new_image_shape = tuple([2*x for x in img.shape])
# new_image_shape = [440, 300]
resized = Image_Resizing(img, new_image_shape)
plt.imshow(resized)
plt.show();
| Homeworks/2019.09.26. Bilinear Interpolation Algorithm for Image Resizing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
run_checks = False
# + [markdown] _cell_guid="e1aad90f-aee7-428a-b692-0c13cb713e28" _uuid="6240d42e-a73e-4104-9076-71a4f5f4f732"
# ### Overview
# This notebook works on the IEEE-CIS Fraud Detection competition. Here I build a simple XGBoost model based on a balanced dataset.
# -
# ### Lessons:
#
# . keep the categorical variables as single items
#
# . Use a high max_depth for xgboost (maybe 40)
#
#
# ### Ideas to try:
#
# . train divergence of expected value (eg. for TransactionAmt and distance based on the non-fraud subset (not all subset as in the case now)
#
# . try using a temporal approach to CV
# + _cell_guid="8d760d7d-53b1-4e67-877f-dce1ce151823" _kg_hide-input=true _uuid="452e2475-e300-41b8-bbb6-ded3c1d99325"
# all imports necessary for this notebook
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
import gc
import copy
import missingno as msno
import xgboost
from xgboost import XGBClassifier, XGBRegressor
from sklearn.model_selection import StratifiedKFold, cross_validate, train_test_split
from sklearn.metrics import roc_auc_score, r2_score
import warnings
warnings.filterwarnings('ignore')
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# + _cell_guid="dbab167c-2ec4-481f-a561-6f01cbf288b2" _uuid="d70cb9a7-e834-459c-8386-0200e7d3b25d"
# Helpers
def seed_everything(seed=0):
'''Seed to make all processes deterministic '''
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
def drop_correlated_cols(df, threshold, cols_to_keep, sample_frac = 1):
'''Drops one of two dataframe's columns whose pairwise pearson's correlation is above the provided threshold'''
if sample_frac != 1:
dataset = df.sample(frac = sample_frac).copy()
else:
dataset = df
col_corr = set() # Set of all the names of deleted columns
corr_matrix = dataset.corr()
for i in range(len(corr_matrix.columns)):
if corr_matrix.columns[i] in col_corr:
continue
for j in range(i):
if corr_matrix.columns[j] in cols_to_keep:
continue
if (corr_matrix.iloc[i, j] >= threshold) and (corr_matrix.columns[j] not in col_corr):
colname = corr_matrix.columns[i] # getting the name of column
col_corr.add(colname)
del dataset
gc.collect()
df.drop(columns = col_corr, inplace = True)
def calc_feature_difference(df, feature_name, indep_features, min_r2 = 0.1, min_r2_improv = 0, frac1 = 0.1,
max_depth_start = 2, max_depth_step = 4):
from copy import deepcopy
print("Feature name %s" %feature_name)
#print("Indep_features %s" %indep_features)
is_imrpoving = True
curr_max_depth = max_depth_start
best_r2 = float("-inf")
clf_best = np.nan
while is_imrpoving:
clf = XGBRegressor(max_depth = curr_max_depth)
rand_sample_indeces = df[df[feature_name].notnull()].sample(frac = frac1).index
clf.fit(df.loc[rand_sample_indeces, indep_features], df.loc[rand_sample_indeces, feature_name])
rand_sample_indeces = df[df[feature_name].notnull()].sample(frac = frac1).index
pred_y = clf.predict(df.loc[rand_sample_indeces, indep_features])
r2Score = r2_score(df.loc[rand_sample_indeces, feature_name], pred_y)
print("%d, R2 score %.4f" % (curr_max_depth, r2Score))
curr_max_depth = curr_max_depth + max_depth_step
if r2Score > best_r2:
best_r2 = r2Score
clf_best = deepcopy(clf)
if r2Score < best_r2 + (best_r2 * min_r2_improv) or (curr_max_depth > max_depth_start * max_depth_step and best_r2 < min_r2 / 2):
is_imrpoving = False
print("The best R2 score of %.4f" % ( best_r2))
if best_r2 > min_r2:
pred_feature = clf_best.predict(df.loc[:, indep_features])
return (df[feature_name] - pred_feature)
else:
return df[feature_name]
# + _cell_guid="98d458c0-c733-461f-9954-21bc4d5cdfd2" _uuid="fa9e0ef8-be7e-4968-92df-8fb00fc3babc"
seed_everything()
pd.set_option('display.max_columns', 500)
# -
master_df = pd.read_csv('/kaggle/input/master-df-time-adjusted-top-100-v2csv/master_df_time_adjusted_top_100.csv')
master_df.head()
master_df_df.shape
for col in master_df.select_dtypes(exclude='number').columns:
master_df[col] = master_df[col].astype('category').cat.codes
'''
length_ones = len(master_df[master_df['isFraud']==1])
train_balanced = pd.concat([master_df[master_df['isFraud']==1], (master_df[master_df['isFraud']==0]).sample(length_ones)], axis=0)
#train_balanced = train_balanced.sample(10000)
X_train, X_test, y_train, y_test = train_test_split(
train_balanced.drop(columns=['isFraud', 'TransactionID', 'TransactionDT']), train_balanced['isFraud'],
test_size=1/4, stratify =train_balanced['isFraud'], random_state=0)
print(X_train.shape)
print(X_test.shape)
clf = XGBClassifier(max_depth=5, n_estimators=1000, verbosity=1)
clf.fit(X_train, y_train)
pred_prob = clf.predict_proba(X_test)
pred_prob[:, 1]
roc_score = roc_auc_score(y_test, pred_prob[:, 1])
print("roc_auc score %.4f" % roc_score)
xgboost.plot_importance(clf, max_num_features=20, importance_type='gain')
xgboost.plot_importance(clf, max_num_features=20, importance_type='weight')
'''
# + _cell_guid="eab8542e-85aa-489a-8b72-bdb28caf0c10" _uuid="4058dab4-441b-4670-b0bf-79d228cf212b"
train_balanced = master_df[master_df['isFraud'].notnull()]
temp_list_to_drop = []
temp_list_to_drop.extend(['isFraud', 'TransactionID', 'TransactionDT', 'is_train_df'])
print(train_balanced.shape)
clf = XGBClassifier(max_depth=50)
clf.fit(train_balanced.drop(columns=temp_list_to_drop), train_balanced['isFraud'])
# + _cell_guid="dfaa5ac4-6edc-4c88-b3f8-e81394a0d316" _uuid="87646822-7836-4e74-8d3d-b87c6d71f000"
gc.collect()
# + _cell_guid="2ccb0178-454f-4a26-bc76-face494a8bf8" _uuid="e65ca54c-111d-4b4c-82a1-d9801ee0457f"
# prepare submission
temp_list_to_drop = []
#temp_list_to_drop = list(cols_cat)
temp_list_to_drop.extend(['isFraud', 'TransactionID', 'TransactionDT'])
temp_list_to_include = list(set(master_df.columns).difference(set(temp_list_to_drop)))
temp_list_to_drop = []
#temp_list_to_drop = list(cols_cat)
temp_list_to_drop.extend(['isFraud', 'TransactionID', 'TransactionDT'])
temp_list_to_include = list(train_balanced.drop(columns=temp_list_to_drop).columns)
temp_list_to_drop = []
#temp_list_to_drop = list(cols_cat)
temp_list_to_drop.extend(['isFraud', 'TransactionID', 'TransactionDT', 'is_train_df'])
counter_from = master_df.loc[master_df['is_train_df']==0, 'isFraud'].index[0]
len_master_df = len(master_df)
print(counter_from)
print(len_master_df)
print('start!!')
while counter_from < len_master_df:
print(counter_from)
counter_to = counter_from + 10000
pred = pd.DataFrame()
#print(len(master_df['isFraud'].loc[counter_from:counter_to]))
#print(len(master_df.loc[counter_from:counter_to, [col for col in master_df.columns if col not in temp_list_to_drop]]))
master_df['isFraud'].loc[counter_from:counter_to] = clf.predict_proba(master_df.loc[counter_from:counter_to, [col for col in master_df.columns if col not in temp_list_to_drop]])[:, 1]
counter_from += 10000
gc.collect()
#print(temp_list_to_include)
# + _cell_guid="41474e8a-fa68-4611-a657-dbf0b5122e7e" _uuid="16e31eb9-e221-408e-ae8c-2fd907129870"
#sample_submission.head()
# + _cell_guid="9d1ac8b3-26cf-4094-81cc-9500b6dfade8" _uuid="92fec1ec-2f87-45a2-b9b1-3c4fc2a65f01"
counter_from = master_df.loc[master_df['is_train_df']==0, 'isFraud'].index[0]
submission = pd.DataFrame(master_df[['TransactionID', 'isFraud']].loc[counter_from:]).reset_index(drop = True)
submission.head()
# + _cell_guid="aec70b7c-6f01-451b-b305-61cd5f57b132" _uuid="a388dc2f-48ad-4d10-b2df-258f68bfdb1e"
submission.describe()
# + _cell_guid="d7b4c0d6-ac52-40c0-b73e-75c5c8fa5578" _uuid="e00a5899-3492-4f04-9e01-2a10883b1359"
submission.to_csv('submission.csv', index=False)
# -
| ieee-based-on-time-adj-top100-pure-v2-0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# tgb - 3/4/2020 - Cleaning up the code for the non-linear ACnet developed in notebook [https://github.com/tbeucler/CBRAIN-CAM/blob/master/notebooks/tbeucler_devlog/035_RH_layers.ipynb]. Includes:
# - Moist thermodynamics libraries in both tensorflow and numpy
# - Code to build & train non-linear UCnet and ACnet
# - Diagnostics of non-linear UCnet/ACnet's performances & energy/mass conservation
# # 0) Initialization
# +
from cbrain.imports import *
from cbrain.data_generator import *
from cbrain.cam_constants import *
from cbrain.losses import *
from cbrain.utils import limit_mem
from cbrain.layers import *
from cbrain.data_generator import DataGenerator
import tensorflow as tf
import tensorflow.math as tfm
import tensorflow_probability as tfp
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
import xarray as xr
import numpy as np
from cbrain.model_diagnostics import ModelDiagnostics
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.image as imag
import scipy.integrate as sin
import cartopy.crs as ccrs
import matplotlib.ticker as mticker
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import pickle
TRAINDIR = '/local/Tom.Beucler/SPCAM_PHYS/'
DATADIR = '/project/meteo/w2w/A6/S.Rasp/SP-CAM/fluxbypass_aqua/'
PREFIX = '8col009_01_'
# %cd /filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM
# Otherwise tensorflow will use ALL your GPU RAM for no reason
limit_mem()
# -
# # 1) Tensorflow library
# ## 1.1) Moist thermodynamics
# +
# Moist thermodynamics library adapted to tf
def eliq(T):
a_liq = np.float32(np.array([-0.976195544e-15,-0.952447341e-13,\
0.640689451e-10,\
0.206739458e-7,0.302950461e-5,0.264847430e-3,\
0.142986287e-1,0.443987641,6.11239921]));
c_liq = np.float32(-80.0)
T0 = np.float32(273.16)
return np.float32(100.0)*tfm.polyval(a_liq,tfm.maximum(c_liq,T-T0))
def eice(T):
a_ice = np.float32(np.array([0.252751365e-14,0.146898966e-11,0.385852041e-9,\
0.602588177e-7,0.615021634e-5,0.420895665e-3,\
0.188439774e-1,0.503160820,6.11147274]));
c_ice = np.float32(np.array([273.15,185,-100,0.00763685,0.000151069,7.48215e-07]))
T0 = np.float32(273.16)
return tf.where(T>c_ice[0],eliq(T),\
tf.where(T<=c_ice[1],np.float32(100.0)*(c_ice[3]+tfm.maximum(c_ice[2],T-T0)*\
(c_ice[4]+tfm.maximum(c_ice[2],T-T0)*c_ice[5])),\
np.float32(100.0)*tfm.polyval(a_ice,T-T0)))
def esat(T):
T0 = np.float32(273.16)
T00 = np.float32(253.16)
omtmp = (T-T00)/(T0-T00)
omega = tfm.maximum(np.float32(0.0),tfm.minimum(np.float32(1.0),omtmp))
return tf.where(T>T0,eliq(T),tf.where(T<T00,eice(T),(omega*eliq(T)+(1-omega)*eice(T))))
def qv(T,RH,P0,PS,hyam,hybm):
R = np.float32(287.0)
Rv = np.float32(461.0)
p = P0 * hyam + PS[:, None] * hybm # Total pressure (Pa)
T = tf.cast(T,tf.float32)
RH = tf.cast(RH,tf.float32)
p = tf.cast(p,tf.float32)
return R*esat(T)*RH/(Rv*p)
# DEBUG 1
# return esat(T)
def RH(T,qv,P0,PS,hyam,hybm):
R = np.float32(287.0)
Rv = np.float32(461.0)
p = P0 * hyam + PS[:, None] * hybm # Total pressure (Pa)
T = tf.cast(T,tf.float32)
qv = tf.cast(qv,tf.float32)
p = tf.cast(p,tf.float32)
return Rv*p*qv/(R*esat(T))
# -
# ## 1.2) Conversion Layers
# ### 1.2.1) From relative to specific humidity (inputs)
class RH2QV(Layer):
def __init__(self, inp_subQ, inp_divQ, inp_subRH, inp_divRH, hyam, hybm, **kwargs):
"""
Call using ([input])
Converts specific humidity to relative humidity and renormalizes all inputs
in preparation for ACnet
Assumes
prior: [RHBP,
QCBP, QIBP, TBP, VBP, Qdt_adiabatic, QCdt_adiabatic, QIdt_adiabatic,
Tdt_adiabatic, Vdt_adiabatic, PS, SOLIN, SHFLX, LHFLX]
Returns
post(erior): [QBP,
QCBP, QIBP, TBP, VBP, Qdt_adiabatic, QCdt_adiabatic, QIdt_adiabatic,
Tdt_adiabatic, Vdt_adiabatic, PS, SOLIN, SHFLX, LHFLX]
"""
self.inp_subQ, self.inp_divQ, self.inp_subRH, self.inp_divRH, self.hyam, self.hybm = \
np.array(inp_subQ), np.array(inp_divQ), np.array(inp_subRH), np.array(inp_divRH), \
np.array(hyam), np.array(hybm)
# Define variable indices here
# Input
self.QBP_idx = slice(0,30)
self.TBP_idx = slice(90,120)
self.PS_idx = 300
self.SHFLX_idx = 302
self.LHFLX_idx = 303
super().__init__(**kwargs)
def build(self, input_shape):
super().build(input_shape)
def get_config(self):
config = {'inp_subQ': list(self.inp_subQ), 'inp_divQ': list(self.inp_divQ),
'inp_subRH': list(self.inp_subRH), 'inp_divRH': list(self.inp_divRH),
'hyam': list(self.hyam),'hybm': list(self.hybm)}
base_config = super().get_config()
return dict(list(base_config.items()) + list(config.items()))
def call(self, arrs):
prior = arrs
# Denormalize T,RH,PS to get them in physical units
Tprior = prior[:,self.TBP_idx]*self.inp_divRH[self.TBP_idx]+self.inp_subRH[self.TBP_idx]
RHprior = prior[:,self.QBP_idx]*self.inp_divRH[self.QBP_idx]+self.inp_subRH[self.QBP_idx]
PSprior = prior[:,self.PS_idx]*self.inp_divRH[self.PS_idx]+self.inp_subRH[self.PS_idx]
# Calculate qv from RH,PS,T using moist thermo library & normalize
qvprior = (qv(Tprior,RHprior,P0,PSprior,self.hyam,self.hybm)-\
self.inp_subQ[self.QBP_idx])/self.inp_divQ[self.QBP_idx]
# Concatenate renormalized inputs to form final input vector
post = tf.concat([tf.cast(qvprior,tf.float32),
((prior[:,30:]*self.inp_divRH[30:]+self.inp_subRH[30:])\
-self.inp_subQ[30:])/self.inp_divQ[30:]\
], axis=1)
return post
def compute_output_shape(self,input_shape):
"""Input shape + 1"""
return (input_shape[0][0])
# ### 1.2.2) From specific to relative humidity time-tendency (output)
class dQVdt2dRHdt(Layer):
def __init__(self, inp_subQ, inp_divQ, norm_qQ, norm_TQ, inp_subRH, inp_divRH, norm_qRH, hyam, hybm, **kwargs):
"""
Call using ([input_qv,output])
Converts specific humidity tendency output to relative humidity tendency output
Assumes
prior: [PHQ, PHCLDLIQ, PHCLDICE, TPHYSTND, QRL, QRS, DTVKE, FSNT, FSNS, FLNT, FLNS, PRECT, PRECTEND, PRECST, PRECSTEN]
Returns
post(erior): [dRHdt, PHCLDLIQ, PHCLDICE, TPHYSTND, QRL, QRS, DTVKE, FSNT, FSNS, FLNT, FLNS, PRECT, PRECTEND, PRECST, PRECSTEN]
"""
self.inp_subQ, self.inp_divQ, self.norm_qQ, self.norm_TQ, \
self.inp_subRH, self.inp_divRH, self.norm_qRH, \
self.hyam, self.hybm = \
np.array(inp_subQ), np.array(inp_divQ), \
np.array(norm_qQ), np.array(norm_TQ),\
np.array(inp_subRH), np.array(inp_divRH), np.array(norm_qRH), \
np.array(hyam), np.array(hybm)
# Define variable indices here
# Input
self.PHQ_idx = slice(0,30)
self.QBP_idx = slice(0,30)
self.TBP_idx = slice(90,120)
self.PS_idx = 300
self.SHFLX_idx = 302
self.LHFLX_idx = 303
super().__init__(**kwargs)
def build(self, input_shape):
super().build(input_shape)
def get_config(self):
config = {'inp_subQ': list(self.inp_subQ), 'inp_divQ': list(self.inp_divQ),
'norm_qQ': list(self.norm_qQ),'norm_TQ':list(self.norm_TQ),
'inp_subRH': list(self.inp_subRH), 'inp_divRH': list(self.inp_divRH),
'norm_qRH': list(self.norm_qRH),
'hyam': list(self.hyam),'hybm': list(self.hybm)}
base_config = super().get_config()
return dict(list(base_config.items()) + list(config.items()))
def call(self, arrs):
inp, prior = arrs
# Denormalize specific humidity, temperature and surface pressure to convert them to physical units
Qprior = inp[:,self.QBP_idx]*self.inp_divQ[self.QBP_idx]+self.inp_subQ[self.QBP_idx]
Tprior = inp[:,self.TBP_idx]*self.inp_divQ[self.TBP_idx]+self.inp_subQ[self.TBP_idx]
PSprior = inp[:,self.PS_idx]*self.inp_divQ[self.PS_idx]+self.inp_subQ[self.PS_idx]
# Calculate specific humidity after physics using its time-tendency
dqvdtprior = prior[:,self.QBP_idx]/self.norm_qQ
Q2prior = Qprior+DT*dqvdtprior
# Calculate temperature after physics using its time-tendency
dTdtprior = prior[:,self.TBP_idx]/self.norm_TQ
T2prior = Tprior+DT*dTdtprior
# Infer the relative humidity tendency from relative humidity before & after physics
RHprior = RH(Tprior,Qprior,P0,PSprior,self.hyam,self.hybm)
RH2prior = RH(T2prior,Q2prior,P0,PSprior,self.hyam,self.hybm)
dRHdtprior = ((RH2prior-RHprior)/DT)*self.norm_qRH
# Concatenate the relative humidity tendency with the remaining outputs
post = tf.concat([dRHdtprior,prior[:,30:]], axis=1)
return post
def compute_output_shape(self,input_shape):
"""Input shape"""
return (input_shape[0][0],input_shape[0][1])
# # 2) Build UCnet_NL and ACnet_NL
# ## 2.1) Generators
# %cd /filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM
scale_dict = load_pickle('./nn_config/scale_dicts/009_Wm2_scaling.pkl')
scale_dict['dRHdt'] = 5*L_S/G, # Factor 5 in loss to give std of dRH/dt similar weight as std of dT/dt
# ### 2.1.1) Generator using RH
in_vars = ['RH', 'QCBP', 'QIBP', 'TBP', 'VBP',
'Qdt_adiabatic', 'QCdt_adiabatic', 'QIdt_adiabatic', 'Tdt_adiabatic', 'Vdt_adiabatic',
'PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_vars = ['dRHdt', 'PHCLDLIQ', 'PHCLDICE', 'TPHYSTND', 'QRL', 'QRS', 'DTVKE',
'FSNT', 'FSNS', 'FLNT', 'FLNS', 'PRECT', 'PRECTEND', 'PRECST', 'PRECSTEN']
TRAINFILE = '8col009RH_01_train.nc'
NORMFILE = '8col009RH_01_norm.nc'
VALIDFILE = '8col009RH_01_valid.nc'
TESTFILE = '8col009RH_01_test.nc'
train_gen = DataGenerator(
data_fn = TRAINDIR+TRAINFILE,
input_vars = in_vars,
output_vars = out_vars,
norm_fn = TRAINDIR+NORMFILE,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = TRAINDIR+VALIDFILE,
input_vars = in_vars,
output_vars = out_vars,
norm_fn = TRAINDIR+NORMFILE,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
test_gen = DataGenerator(
data_fn = TRAINDIR+TESTFILE,
input_vars = in_vars,
output_vars = out_vars,
norm_fn = TRAINDIR+NORMFILE,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
# ### 2.1.2) Generators using qv
TRAINFILEQ = '8col009_01_train.nc'
VALIDFILEQ = '8col009_01_valid.nc'
NORMFILEQ = '8col009_01_norm.nc'
TESTFILEQ = '8col009_01_test.nc'
scale_dictQ = load_pickle('./nn_config/scale_dicts/009_Wm2_scaling.pkl')
in_varsQ = ['QBP', 'QCBP', 'QIBP', 'TBP', 'VBP',
'Qdt_adiabatic', 'QCdt_adiabatic', 'QIdt_adiabatic', 'Tdt_adiabatic', 'Vdt_adiabatic',
'PS', 'SOLIN', 'SHFLX', 'LHFLX']
out_varsQ = ['PHQ', 'PHCLDLIQ', 'PHCLDICE', 'TPHYSTND', 'QRL', 'QRS', 'DTVKE',
'FSNT', 'FSNS', 'FLNT', 'FLNS', 'PRECT', 'PRECTEND', 'PRECST', 'PRECSTEN']
train_genQ = DataGenerator(
data_fn = TRAINDIR+TRAINFILEQ,
input_vars = in_varsQ,
output_vars = out_varsQ,
norm_fn = TRAINDIR+NORMFILEQ,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dictQ,
batch_size=1024,
shuffle=True
)
valid_genQ = DataGenerator(
data_fn = TRAINDIR+VALIDFILEQ,
input_vars = in_varsQ,
output_vars = out_varsQ,
norm_fn = TRAINDIR+NORMFILEQ,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dictQ,
batch_size=1024,
shuffle=True
)
test_genQ = DataGenerator(
data_fn = TRAINDIR+TESTFILEQ,
input_vars = in_varsQ,
output_vars = out_varsQ,
norm_fn = TRAINDIR+NORMFILEQ,
input_transform = ('mean', 'maxrs'),
output_transform = scale_dictQ,
batch_size=1024,
shuffle=True
)
# ## 2.2) Models
# ### 2.2.1) UCnet NL
inp = Input(shape=(304,))
inpQ = RH2QV(inp_subQ=train_genQ.input_transform.sub,
inp_divQ=train_genQ.input_transform.div,
inp_subRH=train_gen.input_transform.sub,
inp_divRH=train_gen.input_transform.div,
hyam=hyam, hybm=hybm)(inp)
densout = Dense(512, activation='linear')(inpQ)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
outQ = Dense(218, activation='linear')(densout)
out = dQVdt2dRHdt(inp_subQ=train_genQ.input_transform.sub,
inp_divQ=train_genQ.input_transform.div,
norm_qQ=scale_dictQ['PHQ'],
inp_subRH=train_gen.input_transform.sub,
inp_divRH=train_gen.input_transform.div,
norm_qRH=scale_dict['dRHdt'],
hyam=hyam, hybm=hybm)([inpQ, outQ])
UCnet_NL = tf.keras.models.Model(inp, out)
name = 'UCnetNL_20'
path_HDF5 = '/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/'
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint(path_HDF5+name+'.hdf5',save_best_only=True, monitor='val_loss', mode='min')
UCnet_NL.compile(tf.keras.optimizers.RMSprop(), loss=mse)
Nep = 10
UCnet_NL.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,\
callbacks=[earlyStopping, mcp_save])
# ### 2.2.2) ACnet NL
inp = Input(shape=(304,))
inpQ = RH2QV(inp_subQ=train_genQ.input_transform.sub,
inp_divQ=train_genQ.input_transform.div,
inp_subRH=train_gen.input_transform.sub,
inp_divRH=train_gen.input_transform.div,
hyam=hyam, hybm=hybm)(inp)
densout = Dense(512, activation='linear')(inpQ)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(214, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
surfout = SurRadLayer(
inp_div=train_genQ.input_transform.div,
inp_sub=train_genQ.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpQ, densout])
massout = MassConsLayer(
inp_div=train_genQ.input_transform.div,
inp_sub=train_genQ.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpQ, surfout])
enthout = EntConsLayer(
inp_div=train_genQ.input_transform.div,
inp_sub=train_genQ.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpQ, massout])
out = dQVdt2dRHdt(inp_subQ=train_genQ.input_transform.sub,
inp_divQ=train_genQ.input_transform.div,
norm_qQ=scale_dictQ['PHQ'],
norm_TQ=scale_dictQ['TPHYSTND'],
inp_subRH=train_gen.input_transform.sub,
inp_divRH=train_gen.input_transform.div,
norm_qRH=scale_dict['dRHdt'],
hyam=hyam, hybm=hybm)([inpQ, enthout])
ACnet_NL = tf.keras.models.Model(inp, out)
name = 'ACnetNL_20'
path_HDF5 = '/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/'
earlyStopping = EarlyStopping(monitor='val_loss', patience=10, verbose=0, mode='min')
mcp_save = ModelCheckpoint(path_HDF5+name+'.hdf5',save_best_only=True, monitor='val_loss', mode='min')
ACnet_NL.compile(tf.keras.optimizers.RMSprop(), loss=mse)
Nep = 10
ACnet_NL.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen,\
callbacks=[earlyStopping, mcp_save])
# # 3) Numpy library
# <a id='np_destination'></a>
# ## 3.1) Moist thermodynamics
# +
def eliq(T):
a_liq = np.array([-0.976195544e-15,-0.952447341e-13,0.640689451e-10,0.206739458e-7,0.302950461e-5,0.264847430e-3,0.142986287e-1,0.443987641,6.11239921]);
c_liq = -80
T0 = 273.16
return 100*np.polyval(a_liq,np.maximum(c_liq,T-T0))
def deliqdT(T):
a_liq = np.array([-0.599634321e-17,-0.792933209e-14,-0.604119582e-12,0.385208005e-9,0.103167413e-6,0.121167162e-4,0.794747212e-3,0.285976452e-1,0.443956472])
c_liq = -80
T0 = 273.16
return 100*np.polyval(a_liq,np.maximum(c_liq,T-T0))
def eice(T):
a_ice = np.array([0.252751365e-14,0.146898966e-11,0.385852041e-9,0.602588177e-7,0.615021634e-5,0.420895665e-3,0.188439774e-1,0.503160820,6.11147274]);
c_ice = np.array([273.15,185,-100,0.00763685,0.000151069,7.48215e-07])
T0 = 273.16
return (T>c_ice[0])*eliq(T)+\
(T<=c_ice[0])*(T>c_ice[1])*100*np.polyval(a_ice,T-T0)+\
(T<=c_ice[1])*100*(c_ice[3]+np.maximum(c_ice[2],T-T0)*(c_ice[4]+np.maximum(c_ice[2],T-T0)*c_ice[5]))
def deicedT(T):
a_ice = np.array([0.497275778e-16,0.390204672e-13,0.132073448e-10,0.255653718e-8,0.312668753e-6,0.249065913e-4,0.126710138e-2,0.377174432e-1,0.503223089])
c_ice = np.array([273.15,185,-100,0.0013186,2.60269e-05,1.28676e-07])
T0 = 273.16
return (T>c_ice[0])*deliqdT(T)+\
(T<=c_ice[0])*(T>c_ice[1])*100*np.polyval(a_ice,T-T0)+\
(T<=c_ice[1])*100*(c_ice[3]+np.maximum(c_ice[2],T-T0)*(c_ice[4]+np.maximum(c_ice[2],T-T0)*c_ice[5]))
def esat(T):
T0 = 273.16
T00 = 253.16
omega = np.maximum(0,np.minimum(1,(T-T00)/(T0-T00)))
return (T>T0)*eliq(T)+(T<T00)*eice(T)+(T<=T0)*(T>=T00)*(omega*eliq(T)+(1-omega)*eice(T))
def RH(T,qv,P0,PS,hyam,hybm):
R = 287
Rv = 461
p = np.moveaxis((hyam*P0+hybm*PS).values,0,1) # Total pressure (Pa)
return Rv*p*qv/(R*esat(T))
def qv(T,RH,P0,PS,hyam,hybm):
R = 287
Rv = 461
Bsize = np.shape(T)[0]
p = np.tile(hyam*P0,(Bsize,1))+np.tile(hybm,(Bsize,1))*np.tile(PS,(30,1)).T
return R*esat(T)*RH/(Rv*p)
# -
# ## 3.2) Mass/Energy/Radiation checkers
def mass_res_diagno(inp_div,inp_sub,norm_q,inp,pred):
# Input
PS_idx = 300
LHFLX_idx = 303
# Output
PHQ_idx = slice(0, 30)
PHCLDLIQ_idx = slice(30, 60)
PHCLDICE_idx = slice(60, 90)
PRECT_idx = 214
PRECTEND_idx = 215
# 1. Compute dP_tilde
dP_tilde = compute_dP_tilde(inp[:, PS_idx], inp_div[PS_idx], inp_sub[PS_idx], norm_q, hyai, hybi)
# 2. Compute water integral
WATINT = np.sum(dP_tilde *(pred[:, PHQ_idx] + pred[:, PHCLDLIQ_idx] + pred[:, PHCLDICE_idx]), axis=1)
# print('PHQ',np.mean(np.sum(dP_tilde*pred[:,PHQ_idx],axis=1)))
# print('PHCLQ',np.mean(np.sum(dP_tilde*pred[:,PHCLDLIQ_idx],axis=1)))
# print('PHICE',np.mean(np.sum(dP_tilde*pred[:,PHCLDICE_idx],axis=1)))
# 3. Compute latent heat flux and precipitation forcings
LHFLX = inp[:, LHFLX_idx] * inp_div[LHFLX_idx] + inp_sub[LHFLX_idx]
PREC = pred[:, PRECT_idx] + pred[:, PRECTEND_idx]
# 4. Compute water mass residual
# print('LHFLX',np.mean(LHFLX))
# print('PREC',np.mean(PREC))
# print('WATINT',np.mean(WATINT))
WATRES = LHFLX - PREC - WATINT
#print('WATRES',np.mean(WATRES))
return np.square(WATRES)
def ent_res_diagno(inp_div,inp_sub,norm_q,inp,pred):
# Input
PS_idx = 300
SHFLX_idx = 302
LHFLX_idx = 303
# Output
PHQ_idx = slice(0, 30)
PHCLDLIQ_idx = slice(30, 60)
PHCLDICE_idx = slice(60, 90)
TPHYSTND_idx = slice(90, 120)
DTVKE_idx = slice(180, 210)
FSNT_idx = 210
FSNS_idx = 211
FLNT_idx = 212
FLNS_idx = 213
PRECT_idx = 214
PRECTEND_idx = 215
PRECST_idx = 216
PRECSTEND_idx = 217
# 1. Compute dP_tilde
dP_tilde = compute_dP_tilde(inp[:, PS_idx], inp_div[PS_idx], inp_sub[PS_idx], norm_q, hyai, hybi)
# 2. Compute net energy input from phase change and precipitation
PHAS = L_I / L_V * (
(pred[:, PRECST_idx] + pred[:, PRECSTEND_idx]) -
(pred[:, PRECT_idx] + pred[:, PRECTEND_idx])
)
# 3. Compute net energy input from radiation, SHFLX and TKE
RAD = (pred[:, FSNT_idx] - pred[:, FSNS_idx] -
pred[:, FLNT_idx] + pred[:, FLNS_idx])
SHFLX = (inp[:, SHFLX_idx] * inp_div[SHFLX_idx] +
inp_sub[SHFLX_idx])
KEDINT = np.sum(dP_tilde * pred[:, DTVKE_idx], 1)
# 4. Compute tendency of vapor due to phase change
LHFLX = (inp[:, LHFLX_idx] * inp_div[LHFLX_idx] +
inp_sub[LHFLX_idx])
VAPINT = np.sum(dP_tilde * pred[:, PHQ_idx], 1)
SPDQINT = (VAPINT - LHFLX) * L_S / L_V
# 5. Same for cloud liquid water tendency
SPDQCINT = np.sum(dP_tilde * pred[:, PHCLDLIQ_idx], 1) * L_I / L_V
# 6. And the same for T but remember residual is still missing
DTINT = np.sum(dP_tilde * pred[:, TPHYSTND_idx], 1)
# 7. Compute enthalpy residual
ENTRES = SPDQINT + SPDQCINT + DTINT - RAD - SHFLX - PHAS - KEDINT
return np.square(ENTRES)
# +
def lw_res_diagno(inp_div,inp_sub,norm_q,inp,pred):
# Input
PS_idx = 300
# Output
QRL_idx = slice(120, 150)
FLNS_idx = 213
FLNT_idx = 212
# 1. Compute dP_tilde
dP_tilde = compute_dP_tilde(inp[:, PS_idx], inp_div[PS_idx], inp_sub[PS_idx], norm_q, hyai, hybi)
# 2. Compute longwave integral
LWINT = np.sum(dP_tilde *pred[:, QRL_idx], axis=1)
# 3. Compute net longwave flux from lw fluxes at top and bottom
LWNET = pred[:, FLNS_idx] - pred[:, FLNT_idx]
# 4. Compute water mass residual
LWRES = LWINT-LWNET
return np.square(LWRES)
def sw_res_diagno(inp_div,inp_sub,norm_q,inp,pred):
# Input
PS_idx = 300
# Output
QRS_idx = slice(150, 180)
FSNS_idx = 211
FSNT_idx = 210
# 1. Compute dP_tilde
dP_tilde = compute_dP_tilde(inp[:, PS_idx], inp_div[PS_idx], inp_sub[PS_idx], norm_q, hyai, hybi)
# 2. Compute longwave integral
SWINT = np.sum(dP_tilde *pred[:, QRS_idx], axis=1)
# 3. Compute net longwave flux from lw fluxes at top and bottom
SWNET = pred[:, FSNT_idx] - pred[:, FSNS_idx]
# 4. Compute water mass residual
SWRES = SWINT-SWNET
return np.square(SWRES)
# -
def tot_res_diagno(inp_div,inp_sub,norm_q,inp,pred):
return 0.25*(mass_res_diagno(inp_div,inp_sub,norm_q,inp,pred)+\
ent_res_diagno(inp_div,inp_sub,norm_q,inp,pred)+\
lw_res_diagno(inp_div,inp_sub,norm_q,inp,pred)+\
sw_res_diagno(inp_div,inp_sub,norm_q,inp,pred))
# [Link to diagnostics](#diagnostics)
# # 4) Diagnostics
# ## 4.1) Load models
path_HDF5 = '/local/Tom.Beucler/SPCAM_PHYS/HDF5_DATA/'
NNarray = ['035_UCnet.hdf5','UCnet_11.hdf5','UCnet_12.hdf5',
'UCnetNL_10.hdf5','UCnetNL_11.hdf5','UCnetNL_12.hdf5',
'ACnetNL_10.hdf5','ACnetNL_11.hdf5','ACnetNL_12.hdf5']
NNname = ['UCnet','UCnet_{NL}','ACnet_{NL}'] # TODO: Add UCnet_NL
dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer,
'RH2QV':RH2QV,'dQVdt2dRHdt':dQVdt2dRHdt,
'eliq':eliq,'eice':eice,'esat':esat,'qv':qv,'RH':RH}
NN = {}; md = {};
# %cd $TRAINDIR/HDF5_DATA
for i,NNs in enumerate(NNarray):
print('NN name is ',NNs)
path = path_HDF5+NNs
NN[NNs] = load_model(path,custom_objects=dict_lay)
# ## 4.2) Calculate square error and physical constraints residual
# [Link to numpy library for diagnostics](#np_destination)
# <a id='diagnostics'></a>
gen = test_gen
genQ = test_genQ
# +
# SE = {}
# TRES = {}
# for iNNs,NNs in enumerate(['UCnetNL_10.hdf5','ACnetNL_10.hdf5']):
# SE[NNs] = np.zeros((1,218))
# TRES[NNs] = np.zeros((1,))
# -
SE = {}
TRES = {}
MSE = {}
# +
spl = 0
while gen[spl][0].size>0: #spl is sample number
print('spl=',spl,' ',end='\r')
inp = gen[spl][0]
truth = gen[spl][1]
inp_phys = inp*gen.input_transform.div+gen.input_transform.sub
for iNNs,NNs in enumerate(NNarray):
pred = NN[NNs].predict_on_batch(inp)
se = (pred-truth)**2
pred_phys = pred/gen.output_transform.scale
QV1 = qv(inp_phys[:,90:120],inp_phys[:,:30],P0,inp_phys[:,300],hyam,hybm)
QV2 = qv(inp_phys[:,90:120]+DT*pred_phys[:,90:120],
inp_phys[:,:30]+DT*pred_phys[:,:30],P0,inp_phys[:,300],hyam,hybm)
dQVdt = train_genQ.output_transform.scale[:30]*(QV2-QV1)/DT
predQ = np.copy(pred)
predQ[:,:30] = dQVdt
tresid = tot_res_diagno(gen.input_transform.div,gen.input_transform.sub,
genQ.output_transform.scale[:30],inp,predQ)
if spl==0: SE[NNs] = se; TRES[NNs] = tresid; MSE[NNs] = np.mean(se,axis=1);
else:
SE[NNs] += se;
TRES[NNs] = np.concatenate((TRES[NNs],tresid),axis=0);
MSE[NNs] = np.concatenate((MSE[NNs],np.mean(se,axis=1)),axis=0);
spl += 1
for iNNs,NNs in enumerate(NNarray): SE[NNs] /= spl;
# -
plt.plot(np.mean(SE['ACnetNL_12.hdf5'][:,:30],axis=0),color='r')
plt.plot(np.mean(SE['UCnetNL_12.hdf5'][:,:30],axis=0),color='b')
plt.plot(np.mean(SE['UCnet_12.hdf5'][:,:30],axis=0),color='g')
plt.plot(np.mean(SE['ACnetNL_11.hdf5'][:,:30],axis=0),color='r')
plt.plot(np.mean(SE['UCnetNL_11.hdf5'][:,:30],axis=0),color='b')
plt.plot(np.mean(SE['UCnet_11.hdf5'][:,:30],axis=0),color='g')
plt.hist(TRES['ACnetNL_10.hdf5'],bins=100);
plt.hist(TRES['UCnetNL_10.hdf5'],bins=100);
plt.hist(TRES['035_UCnet.hdf5'],bins=100);
plt.hist(TRES['UCnetNL_10.hdf5'],bins=100);
np.mean(TRES['ACnetNL_10.hdf5']),np.std(TRES['ACnetNL_10.hdf5'])
np.mean(TRES['UCnetNL_10.hdf5']),np.std(TRES['UCnetNL_10.hdf5'])
np.mean(TRES['035_UCnet.hdf5']),np.std(TRES['035_UCnet.hdf5'])
# ## 4.3) Save reduced data in PKL format
pathPKL = '/home/t/Tom.Beucler/SPCAM/CBRAIN-CAM/notebooks/tbeucler_devlog/PKL_DATA/'
hf = open(pathPKL+'2020_03_04_testgen041.pkl','wb')
S = {"TRES":TRES,"MSE":MSE,"SE":SE}
pickle.dump(S,hf)
hf.close()
| notebooks/tbeucler_devlog/041_ACnet_Non_Linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: toy
# language: python
# name: toy
# ---
# %load_ext autoreload
# +
import numpy as np
from scipy.stats import itemfreq
import networkx as nx
import pandas as pd
import matplotlib
import seaborn as sns
sns.set_palette('colorblind')
import matplotlib.pyplot as plt
# %matplotlib inline
matplotlib.rcParams['font.size'] = 30
matplotlib.rcParams['xtick.major.size'] = 9
matplotlib.rcParams['ytick.major.size'] = 9
matplotlib.rcParams['xtick.minor.size'] = 4
matplotlib.rcParams['ytick.minor.size'] = 4
matplotlib.rcParams['axes.linewidth'] = 2
matplotlib.rcParams['xtick.major.width'] = 2
matplotlib.rcParams['ytick.major.width'] = 2
matplotlib.rcParams['xtick.minor.width'] = 2
matplotlib.rcParams['ytick.minor.width'] = 2
matplotlib.rcParams['figure.figsize'] = [10, 8]
matplotlib.rcParams['text.usetex'] = True
import random
from toysimulations import Network
import pickle
# -
import networkx as nx
G = nx.grid_2d_graph(5,5)
pos = {n:n for n in G}
# +
import matplotlib.patches as patches
from matplotlib.path import Path
fig, ax = plt.subplots(figsize=(8,8))
ax.axis('off')
bg_elems_color = "xkcd:light grey blue"
draw_nodes_kwargs = dict(node_color = bg_elems_color, alpha=1.0)
draw_stop_kwargs = dict(node_size=500, node_color = "xkcd:medium blue", alpha=1)
draw_edge_kwargs = dict(width=3, edge_color=bg_elems_color, alpha=1.0)
draw_path_kwargs = dict(width=7, color='black')
volume_patch_kwargs = dict(lw=2, zorder=-3, capstyle='round', joinstyle='bevel', alpha=0.6)
nx.draw_networkx_edges(G, pos=pos, ax=ax, **draw_edge_kwargs)
nx.draw_networkx_nodes(G, pos=pos, ax=ax, **draw_nodes_kwargs)
path = [(0, 1), (3,1), (3,2), (1,2), (1,3), (4,3), (4,4)]
stoplist = [(0,1), (3,2), (1,3), (4,4)]
e = 0.25
vol_polygon_vertices = [(0-e,1-e), (3+e,1-e), (3+e,3-e), (4+e,3-e), (4+e,4+e), (1-e,4+e), (1-e,2+e), (0-e,2+e), (0-e, 1-e)]
vol_rest_polygon_vertices = [(1-e,3-e), (4+e,3-e), (4+e,4+e), (1-e,4+e), (1-e,3-e)]
edges_in_path = [(u,v) for u,v in zip(path[:-1], path[1:]) for n in nx.shortest_path(G, u,v)]
nx.draw_networkx_edges(G, pos=pos, edgelist=edges_in_path, ax=ax, **draw_path_kwargs)
nx.draw_networkx_nodes(G, pos=pos, nodelist=stoplist, ax=ax, **draw_stop_kwargs)
nx.draw_networkx_labels(G, pos=pos, labels={(0,1): 1, (3,2): 2, (1,3):3, (4,4):4}, ax=ax, font_size=20, font_color='w')
# patch for v
codes_v = [Path.MOVETO] + [Path.LINETO]*(len(vol_polygon_vertices)-2) + [Path.CLOSEPOLY]
path_v = Path(vol_polygon_vertices, codes_v)
patch_v = patches.PathPatch(path_v, hatch='/', facecolor='xkcd:pale green', label = r'$V=16$', **volume_patch_kwargs)
codes_vrest = [Path.MOVETO] + [Path.LINETO]*(len(vol_rest_polygon_vertices)-2) + [Path.CLOSEPOLY]
path_vrest = Path(vol_rest_polygon_vertices, codes_vrest)
patch_vrest = patches.PathPatch(path_vrest, hatch='.', ls='--', facecolor='xkcd:sand', label = r'$V_{rest}=8$', **volume_patch_kwargs)
ax.add_patch(patch_v)
ax.add_patch(patch_vrest)
ax.legend(loc=(1.1,0.7), fontsize=18)
# -
# ## Now make a two panel plot
# +
bg_elems_color = "xkcd:light grey blue"
draw_nodes_kwargs = dict(node_color = bg_elems_color, alpha=1.0)
draw_stop_kwargs = dict(node_size=500, node_color = "xkcd:medium blue", alpha=1)
draw_edge_kwargs = dict(width=3, edge_color=bg_elems_color, alpha=1.0)
draw_path_kwargs = dict(width=7, color='black')
volume_patch_kwargs = dict(lw=2, zorder=-3, capstyle='round', joinstyle='bevel', alpha=0.6)
nx.draw_networkx_edges(G, pos=pos, ax=ax, **draw_edge_kwargs)
nx.draw_networkx_nodes(G, pos=pos, ax=ax, **draw_nodes_kwargs)
path = [(0, 1), (3,1), (3,2), (1,2), (1,3), (4,3), (4,4)]
stoplist = [(0,1), (3,2), (1,3), (4,4)]
e = 0.25
vol_polygon_vertices = [(0-e,1-e), (3+e,1-e), (3+e,3-e), (4+e,3-e), (4+e,4+e), (1-e,4+e), (1-e,2+e), (0-e,2+e), (0-e, 1-e)]
vol_rest_polygon_vertices = [(1-e,3-e), (4+e,3-e), (4+e,4+e), (1-e,4+e), (1-e,3-e)]
edges_in_path = [(u,v) for u,v in zip(path[:-1], path[1:]) for n in nx.shortest_path(G, u,v)]
# patch for v
# +
def plot_base(ax):
nx.draw_networkx_edges(G, pos=pos, ax=ax, **draw_edge_kwargs)
nx.draw_networkx_nodes(G, pos=pos, ax=ax, **draw_nodes_kwargs)
nx.draw_networkx_edges(G, pos=pos, edgelist=edges_in_path, ax=ax, **draw_path_kwargs)
nx.draw_networkx_nodes(G, pos=pos, nodelist=stoplist, ax=ax, **draw_stop_kwargs)
nx.draw_networkx_labels(G, pos=pos, labels={(0,1): 1, (3,2): 2, (1,3):3, (4,4):4}, ax=ax, font_size=20, font_color='w')
def plot_v(ax):
codes_v = [Path.MOVETO] + [Path.LINETO]*(len(vol_polygon_vertices)-2) + [Path.CLOSEPOLY]
path_v = Path(vol_polygon_vertices, codes_v)
patch_v = patches.PathPatch(path_v, hatch='/', facecolor='xkcd:pale green', label = r'$V=16$', **volume_patch_kwargs)
ax.add_patch(patch_v)
def plot_vrest(ax):
codes_vrest = [Path.MOVETO] + [Path.LINETO]*(len(vol_rest_polygon_vertices)-2) + [Path.CLOSEPOLY]
path_vrest = Path(vol_rest_polygon_vertices, codes_vrest)
patch_vrest = patches.PathPatch(path_vrest, hatch='.', ls='--', facecolor='xkcd:sand', label = r'$V_{\textsf{rest}}=8$', **volume_patch_kwargs)
ax.add_patch(patch_vrest)
# +
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(12,4), gridspec_kw={'wspace': 0.9})
ax1.axis('off')
ax2.axis('off')
plot_base(ax1)
plot_v(ax1)
ax1.legend(loc='upper center', fontsize=18, bbox_to_anchor=(0.5, 1.15), frameon=False)
ax1.text(-0.5, 4.5, "(a)", fontsize=25)
plot_base(ax2)
plot_vrest(ax2)
nx.draw_networkx_nodes(G, pos=pos, nodelist=[(1,2)], ax=ax2)
bbox_props = dict(boxstyle="round,pad=0.3", fc="xkcd:carnation", ec="xkcd:merlot", lw=2)
t = ax2.text(1, 2-0.5, "New pickup", ha="center", va="center",
size=12,
bbox=bbox_props)
ax2.legend(loc='upper center', fontsize=18, bbox_to_anchor=(0.5, 1.15), frameon=False)
ax2.text(-0.5, 4.5, "(b)", fontsize=25)
#fig.tight_layout()
fig.savefig("v_and_v_rest_illustration.pdf", bbox_inches='tight')
# -
# ## Illustrations on the coarse graining
from shapely.geometry import MultiPoint, Point
from descartes.patch import PolygonPatch
from shapely.ops import cascaded_union
# +
G = nx.Graph()
eps = 1/np.sqrt(3)
a = 5
nodes = [
# (label), coords
('a', (0,-1)),
('b', (eps,0)),
('c', (-eps,0)),
('aa', (0,-5)),
('bb', (5,1)),
('cc', (-3,1)),
]
RADIUS = 1
points = {}
for node, pos in nodes:
G.add_node(node, pos=pos)
p = Point(*pos).buffer(RADIUS)
points[node] = p
for u,v in [('a', 'b'), ('b', 'c'), ('c', 'a'), ('a', 'aa'), ('b', 'bb'), ('c', 'cc')]:
G.add_edge(u,v)
def draw_merge_patch(ax):
pa, pb, pc = points['a'], points['b'], points['c']
comp = cascaded_union(points.values())
#comp = pa.union(pb).union(pc)
patch = PolygonPatch(comp, facecolor='grey', edgecolor='k', alpha=0.5, zorder=1)
ax.add_patch(patch)
draw_node_wo_color_kwargs = dict(node_size=80, linewidths=2, edgecolors='xkcd:slate')
draw_node_kwargs = draw_node_wo_color_kwargs.copy()
draw_node_kwargs['node_color'] = 'xkcd:merlot'
draw_edge_kwargs = dict(width=2, alpha=1, edge_color='grey')
def draw_g(ax, G, nodelist = None, node_color=None):
if nodelist is None:
nodelist = G.nodes()
if node_color is None:
node_drawing_kwargs = draw_node_kwargs
else:
node_drawing_kwargs = draw_node_wo_color_kwargs.copy()
node_drawing_kwargs['node_color'] = node_color
nx.draw_networkx_nodes(G, pos={n:G.node[n]['pos'] for n in G.nodes()}, nodelist=nodelist, ax=ax, **node_drawing_kwargs)
nx.draw_networkx_edges(G, pos={n:G.node[n]['pos'] for n in G.nodes()}, ax=ax, **draw_edge_kwargs)
fig, (ax1, ax2, ax3, ax4) = plt.subplots(nrows=1, ncols=4, figsize=(12,4))
for ax in (ax1, ax2, ax3, ax4):
ax.set_aspect('equal')
ax.axis('off')
for ax in (ax1, ax2):
draw_g(ax, G)
draw_merge_patch(ax2)
H = G.copy()
H.add_node('e', pos=(0,0))
H.add_edges_from((['aa', 'e'], ['bb', 'e'], ['cc', 'e']))
H.remove_nodes_from(['a', 'b', 'c'])
draw_g(ax3, H)
draw_g(ax3, H, nodelist=['e'], node_color='xkcd:goldenrod')
J = H.copy()
for u in ['aa', 'bb', 'cc']:
v = 'e'
x, y = np.array(J.node[u]['pos']), np.array(J.node[v]['pos'])
elen = np.linalg.norm(y-x)
targen_elen = 1.8
n = 1
for int_node_num in range(1, int(elen/targen_elen)+1):
frac = targen_elen*int_node_num/elen
pos = x+frac*(y-x)
J.add_node(f'{u}_{v}_{int_node_num}', pos=pos)
draw_g(ax4, H)
new_nodes = set(J.nodes())-set(H.nodes())
draw_g(ax4, J, nodelist=new_nodes, node_color='xkcd:cerulean')
for ax in (ax1, ax2, ax3, ax4):
xmin, xmax = ax.get_xlim()
ymin, ymax = ax.get_ylim()
delta = 1.2
ax.set_xlim(xmin-delta, xmax+delta)
ax.set_ylim(ymin-delta, ymax+delta)
ax1.text(-0.1, 0.8, '(a)', horizontalalignment='center', transform=ax1.transAxes, fontsize=18)
ax2.text(-0.1, 0.8, '(b)', horizontalalignment='center', transform=ax2.transAxes, fontsize=18)
ax3.text(-0.1, 0.8, '(c)', horizontalalignment='center', transform=ax3.transAxes, fontsize=18)
ax4.text(-0.1, 0.8, '(d)', horizontalalignment='center', transform=ax4.transAxes, fontsize=18)
#fig.tight_layout(rect=(0,-300,1,300))
fig.savefig("coarse_graining_illustration.pdf", bbox_inches='tight')
# -
# ## Illustrations on the bias in pickup insertion
# ### Load the data for the 100 node ring
# +
PICKLE_FILE = '../data/ring_100.pkl'
with open(PICKLE_FILE, 'rb') as f:
result = pickle.load(f)
INS_DATA_COLUMNS = ['time', 'stoplist_len', 'stoplist_volume', 'rest_stoplist_volume',
'pickup_idx', 'dropoff_idx', 'insertion_type', 'pickup_enroute',
'dropoff_enroute']
x_range = np.array(sorted(result.keys()))
all_dfs = []
for x in x_range:
ins_df = pd.DataFrame(result[x]['insertion_data'],
columns = INS_DATA_COLUMNS)
ins_df.loc[:, 'x'] = x
# cut out transients
ins_df = ins_df[ins_df['time'] * ins_df['x'] > 80000]
all_dfs.append(ins_df)
master_ins_df = pd.concat(all_dfs)
master_ins_df.head()
# +
fig, ax = plt.subplots(figsize=(12, 9))
(master_ins_df['pickup_idx']/master_ins_df['stoplist_len']).hist(ax=ax)
mean = (master_ins_df['pickup_idx']/master_ins_df['stoplist_len']).mean()
ax.axvline(x=mean, c='k', linewidth=2, linestyle='--')
ax.set_xlabel("Pickup index/Stoplist length")
bbox_props = dict(boxstyle="rarrow,pad=0.3", fc="xkcd:beige", ec="k", lw=2)
t = ax.text(mean*0.93, 100000, f"mean={mean:.2f}", ha="right", va="center", rotation=0,
size=25,
bbox=bbox_props)
ax.set_xlim(0,1)
fig.savefig("illustration_skew_pickup_location.pdf", bbox_inches='tight')
| generate_data_and_plot/fig_extra_v_illustration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Doctest
#
# This is a test notebook for the documentation. Sometimes the creation of the automatic documentation fails as there are non-utf-8 characters. A common mistake is the use of a fancy quote "’" instead of "'".
import os
# +
ipynb_path = os.path.dirname(os.path.realpath("__file__"))
doc_path = os.path.join(ipynb_path, os.pardir, 'docs')
html_files = [_ for _ in os.listdir(doc_path) if _.endswith('.html')]
# -
for file in html_files:
myfile = open(os.path.join(doc_path, file), encoding='utf-8')
try:
for idx, l in enumerate(myfile):
pass
except Exception as e:
print(f'Exception in file {file}. After line {idx}: {l}')
print(e)
myfile = open(os.path.join(doc_path, file), encoding='cp1252')
f = open(os.path.join(doc_path, file), "w", encoding='utf-8')
f.write(myfile.read())
f.close()
| sandbox/Doctest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, transpile, Aer, IBMQ
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
# Loading your IBM Quantum account(s)
provider = IBMQ.load_account()
# -
# # IBM Quantum Challenge 2021
# ## Exercise 5 - Variational Quantum Eigensolver
# ### VQE for LiH molecule
#
#
# The goal was to simulate LiH molecule using the STO-3G basis with the PySCF driver, and find the best ansatz, in terms of the number of CNOT gates, a lower number being better.
#
# Some hints to reduce the number of qubits were
# - freezing the core electrons that do not contribute significantly to chemistry and consider only the valence electrons. Inspect the different transformers in `qiskit_nature.transformers` and find the one that performs the freeze core approximation.
# - Using `ParityMapper` with `two_qubit_reduction=True` to eliminate 2 qubits.
# - Reducucing the number of qubits by inspecting the symmetries of your Hamiltonian using `Z2Symmetries` in Qiskit.
#
# I basically studied these hints from the Qiskit Nature API reference and implemented them in the solution.
#
# In addition, I considered the LiH molecule properties to decide orbitals to remove.LiH is Lithium and Hydrogen. The orbitals are
# - H: 1s
# - Li: 1s, 2s, px, py, pz
# The Li 1s electrons do not form bonds i.e. are core electrons and thus could be removed by FreezeCore.
# Then expected H 1s electrons to interact with Li 2s electrons. Initially, I then thought I can remove p orbitals 3,4 and 5, but could not achieve ground state. It turns out that there is some mixing with Li pz orbital in the bonding so only px and py i.e. 3 and 4 can be removed. Orbital removal could be done with FreezeCore.
#
# For the ansatz, TwoLocal in combination with SPSA or SLSQP optimizer looked promising from the previous part a) of the exercise so I tried those. The 'default' with 3 repetitions and full entanglement worked fine with score 18. Then experimented with reducing the repetions, which amazingly went to just 1 and score 6!
#
# Progress next came after checking ?TwoLocal. I realized there were other entanglements like linear, circular and even a map. Mapping with [(0,1), (1, 2), (2, 3), (3, 0)] worked fantastic bringing the score to 4! From this point the returns of experimenting were diminishing. I actually thought 4 was the limit. Then I realized that somehow the entanglement (3,0) was somehow more important. This made me try mixing in such 'reversed' entanglements until finally [(3, 2), (2, 1), (1, 0)] worked! This 'reverse' linear was fantastic (actually might suggest additions of such reversed maps to TwoLocal i.e. 'reversed linear', 'reversed circular'). I still haven't figured out the 'scientific' reason why entanglement in that direction worked.
#
# The full code is below. Hope it is useful!
#
# ## Code
from qiskit_nature.drivers import PySCFDriver
from qiskit_nature.problems.second_quantization.electronic import ElectronicStructureProblem
from qiskit_nature.mappers.second_quantization import ParityMapper, BravyiKitaevMapper, JordanWignerMapper
from qiskit_nature.converters.second_quantization.qubit_converter import QubitConverter
from qiskit_nature.circuit.library import HartreeFock
from qiskit.circuit.library import TwoLocal
from qiskit_nature.circuit.library import UCCSD, PUCCD, SUCCD
from qiskit import Aer
from qiskit.algorithms.optimizers import COBYLA, L_BFGS_B, SPSA, SLSQP
from qiskit_nature.algorithms.ground_state_solvers.minimum_eigensolver_factories import NumPyMinimumEigensolverFactory
from qiskit_nature.algorithms.ground_state_solvers import GroundStateEigensolver
import numpy as np
from qiskit.algorithms import VQE
from IPython.display import display, clear_output
from qiskit_nature.transformers import FreezeCoreTransformer
from qiskit.opflow import X, Y, Z, I, PauliSumOp, Z2Symmetries
# ### Exact diagonalizer and callback
# +
def exact_diagonalizer(problem, converter):
solver = NumPyMinimumEigensolverFactory()
calc = GroundStateEigensolver(converter, solver)
result = calc.solve(problem)
return result
def callback(eval_count, parameters, mean, std):
# Overwrites the same line when printing
display("Evaluation: {}, Energy: {}, Std: {}".format(eval_count, mean, std))
clear_output(wait=True)
counts.append(eval_count)
values.append(mean)
params.append(parameters)
deviation.append(std)
# -
molecule = 'Li 0.0 0.0 0.0; H 0.0 0.0 1.5474'
driver = PySCFDriver(atom=molecule)
qmolecule = driver.run()
# +
# Definitition of the problem
## Takes a FreezeCoreTransformer with choice to freeze_core and list of orbitals to remove
problem = ElectronicStructureProblem(driver,
q_molecule_transformers=[FreezeCoreTransformer(freeze_core=True, remove_orbitals=[3,4])])
# Generate the second-quantized operators
second_q_ops = problem.second_q_ops()
# Hamiltonian
main_op = second_q_ops[0]
mapper = ParityMapper()
converter = QubitConverter(mapper=mapper, two_qubit_reduction=True, z2symmetry_reduction=[-1])
# The fermionic operators are mapped to qubit operators
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
qubit_op = converter.convert(main_op, num_particles=num_particles)
num_particles = (problem.molecule_data_transformed.num_alpha,
problem.molecule_data_transformed.num_beta)
num_spin_orbitals = 2 * problem.molecule_data_transformed.num_molecular_orbitals
init_state = HartreeFock(num_spin_orbitals, num_particles, converter)
#print(init_state)
#ansatz = UCCSD(converter,num_particles,num_spin_orbitals,initial_state = init_state)
# TwoLocal Ansatz
# Single qubit rotations that are placed on all qubits with independent parameters
rotation_blocks = ['rz','rx']
# Entangling gates
entanglement_blocks = ['cx']
# How the qubits are entangled
entangler_map1 = [(0,1), (1, 2), (2, 3), (3, 0)]
entangler_map2 = [(3, 2), (2, 1), (1, 0)]
entanglement = entangler_map2
#entanglement = 'linear'
# Repetitions of rotation_blocks + entanglement_blocks with independent parameters
repetitions = 1
# Skip the final rotation_blocks layer
skip_final_rotation_layer = False
ansatz = TwoLocal(qubit_op.num_qubits, rotation_blocks, entanglement_blocks, reps=repetitions,
entanglement=entanglement, skip_final_rotation_layer=skip_final_rotation_layer)
# Add the initial state
ansatz.compose(init_state, front=True, inplace=True)
#ansatz.draw()
backend = Aer.get_backend('statevector_simulator')
optimizer = SLSQP(maxiter=500)
result_exact = exact_diagonalizer(problem, converter)
exact_energy = np.real(result_exact.eigenenergies[0])
print("Exact electronic energy", exact_energy)
#print(result_exact)
counts = []
values = []
params = []
deviation = []
# Set initial parameters of the ansatz
# We choose a fixed small displacement
# So all participants start from similar starting point
try:
initial_point = [0.01] * len(ansatz.ordered_parameters)
except:
initial_point = [0.01] * ansatz.num_parameters
algorithm = VQE(ansatz,
optimizer=optimizer,
quantum_instance=backend,
callback=callback,
initial_point=initial_point)
result = algorithm.compute_minimum_eigenvalue(qubit_op)
# -
ansatz.draw()
| solutions by participants/ex5/ex5-JoshuaDJohn-3cnot-2.492mHa-16params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from robot import *
from scipy.stats import multivariate_normal
import random #追加
import copy
class Particle:
def __init__(self, init_pose, weight):
self.pose = init_pose
self.weight = weight
def motion_update(self, nu, omega, time, noise_rate_pdf):
ns = noise_rate_pdf.rvs()
pnu = nu + ns[0]*math.sqrt(abs(nu)/time) + ns[1]*math.sqrt(abs(omega)/time)
pomega = omega + ns[2]*math.sqrt(abs(nu)/time) + ns[3]*math.sqrt(abs(omega)/time)
self.pose = IdealRobot.state_transition(pnu, pomega, time, self.pose)
def observation_update(self, observation, envmap, distance_dev_rate, direction_dev): #変更
for d in observation:
obs_pos = d[0]
obs_id = d[1]
##パーティクルの位置と地図からランドマークの距離と方角を算出##
pos_on_map = envmap.landmarks[obs_id].pos
particle_suggest_pos = IdealCamera.observation_function(self.pose, pos_on_map)
##尤度の計算##
distance_dev = distance_dev_rate*particle_suggest_pos[0]
cov = np.diag(np.array([distance_dev**2, direction_dev**2]))
self.weight *= multivariate_normal(mean=particle_suggest_pos, cov=cov).pdf(obs_pos)
class Mcl:
def __init__(self, envmap, init_pose, num, motion_noise_stds={"nn":0.19, "no":0.001, "on":0.13, "oo":0.2}, \
distance_dev_rate=0.14, direction_dev=0.05):
self.particles = [Particle(init_pose, 1.0/num) for i in range(num)]
self.map = envmap
self.distance_dev_rate = distance_dev_rate
self.direction_dev = direction_dev
v = motion_noise_stds
c = np.diag([v["nn"]**2, v["no"]**2, v["on"]**2, v["oo"]**2])
self.motion_noise_rate_pdf = multivariate_normal(cov=c)
def motion_update(self, nu, omega, time):
for p in self.particles: p.motion_update(nu, omega, time, self.motion_noise_rate_pdf)
def observation_update(self, observation):
for p in self.particles:
p.observation_update(observation, self.map, self.distance_dev_rate, self.direction_dev)
self.resampling()
def resampling(self): ###systematicsampling
ws = np.cumsum([e.weight for e in self.particles]) #重みを累積して足していく(最後の要素が重みの合計になる)
if ws[-1] < 1e-100: ws = [e + 1e-100 for e in ws] #重みの合計が0のときの処理
step = ws[-1]/len(self.particles) #正規化されていない場合はステップが「重みの合計値/N」になる
r = np.random.uniform(0.0, step)
cur_pos = 0
ps = [] #抽出するパーティクルのリスト
while(len(ps) < len(self.particles)):
if r < ws[cur_pos]:
ps.append(self.particles[cur_pos]) #もしかしたらcur_posがはみ出るかもしれませんが例外処理は割愛で
r += step
else:
cur_pos += 1
self.particles = [copy.deepcopy(e) for e in ps] #以下の処理は前の実装と同じ
for p in self.particles: p.weight = 1.0/len(self.particles)
def draw(self, ax, elems):
xs = [p.pose[0] for p in self.particles]
ys = [p.pose[1] for p in self.particles]
vxs = [math.cos(p.pose[2])*p.weight*len(self.particles) for p in self.particles] #重みを要素に反映
vys = [math.sin(p.pose[2])*p.weight*len(self.particles) for p in self.particles] #重みを要素に反映
elems.append(ax.quiver(xs, ys, vxs, vys, \
angles='xy', scale_units='xy', scale=1.5, color="blue", alpha=0.5)) #変更
class EstimationAgent(Agent):
def __init__(self, time_interval, nu, omega, estimator):
super().__init__(nu, omega)
self.estimator = estimator
self.time_interval = time_interval
self.prev_nu = 0.0
self.prev_omega = 0.0
def decision(self, observation=None):
self.estimator.motion_update(self.prev_nu, self.prev_omega, self.time_interval)
self.prev_nu, self.prev_omega = self.nu, self.omega
self.estimator.observation_update(observation)
return self.nu, self.omega
def draw(self, ax, elems):
self.estimator.draw(ax, elems)
# +
def trial():
time_interval = 0.1
world = World(30, time_interval, debug=False)
### 地図を生成して3つランドマークを追加 ###
m = Map()
for ln in [(-4,2), (2,-3), (3,3)]: m.append_landmark(Landmark(*ln))
world.append(m)
### ロボットを作る ###
initial_pose = np.array([0, 0, 0]).T
estimator = Mcl(m, initial_pose, 100) #地図mを渡す
a = EstimationAgent(time_interval, 0.2, 10.0/180*math.pi, estimator)
r = Robot(initial_pose, sensor=Camera(m), agent=a, color="red")
world.append(r)
world.draw()
trial()
# -
| section_particle_filter/mcl13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Writing good code
#
# This is a short list of tips. [Also look at these tips](../about/tips.html) and [these tips for when you're stuck](../about/help.html).
#
# ## Naming variables and functions
#
# I try to put verbs in the names of functions (`get_returns()`) and name dataframes after their unit-levels (`monthly_df`). Avoid ambiguous abbreviations.
#
# ## Code style + Auto-Formatting!
#
# There are many ways to achieve the same things in python. Python tends to be a very readable language compared to others. Still, the _style_ of how you write the code can make python code easy to read or hard to read!
#
# ```{tip}
# 1. Obey the naming suggestions above.
# 2. Use an auto-formatter! This will rewrite your code automatically to have good style!
# ```
#
# There are a few popular auto-formatters (black, yapf, and autopep8). In my [JupyterLab set up](../01/05_jupyterlab.html#my-jupyter-lab-set-up), I explain how I set up Black, the "uncompromising Python code formatter" which is very opinionated ("any color you'd like, as long as it is black").
#
# Look at what Black does to this code:
#
# ````{tabbed} Ugly Code I wrote
#
# This function is too long to even read:
# ```python
# def very_important_function(template: str, *variables, file: os.PathLike, engine: str, header: bool = True, debug: bool = False):
# """Applies `variables` to the `template` and writes to `file`."""
# with open(file, 'w') as f:
# ...
# ```
# ````
#
# ````{tabbed} Black fixed it
#
# I hit CTRL+SHIFT+F and this was the result:
# ```python
# def very_important_function(
# template: str,
# *variables,
# file: os.PathLike,
# engine: str,
# header: bool = True,
# debug: bool = False
# ):
# """Applies `variables` to the `template` and writes to `file`."""
# with open(file, "w") as f:
# ...
# ```
# ````
# ## Use a linter + coding assistance
#
# A linter is a programming tool to detect possible errors in your code and stylistic issues.
#
# See my [JupyterLab set up](../01/05_jupyterlab.html#my-jupyter-lab-set-up), for install instructions you can follow to install [`Jupyterlab-lsp`](https://github.com/jupyter-lsp/jupyterlab-lsp). This extension provides code navigation + hover suggestions + linters + autocompletion + rename assistance.
#
# ## DRY code
#
# Don't Repeat Yourself. See the [Functions](10_Golden_5a) page for tips on how to use functions to reduce duplication.
#
# ## Premature optimization
#
# In this class, you likely won't get to the point where you try to optimize your code for speed. Our problem sets aren't quite massive enough to need that. Some student projects might tempt students to optimize for speed.
#
# Don't! Total time = your coding time (initial time, debug time, revising time) + computer time. Computer time is cheap. Yours is limited.
#
# First: Write clean, easy to use code that works.
#
# Only once your code is virtually complete should you even contemplate speed. And still, you're probably optimizing too soon. (You haven't yet realized that you need to completely reformulate the approach to the problem.)
#
# ## Magic numbers are bad
#
# A magic number is a literal number or parameter value embedded in your code. Having numbers that act as in-line inputs and parameters can easily lead to errors and make modifying code very tough.
#
# Here is an example that tries to download and stock price data.
#
# ````{tabbed} Bad
#
# If I want to change the stocks included, I'll need to change the list of stocks twice.
#
# ```python
# import pandas_datareader as pdr
# from datetime import datetime
#
# # load stock returns
# start = datetime(2004, 1, 1)
# end = datetime(2007, 12, 31)
# stock_prices = pdr.get_data_yahoo(['MSFT','AAPL'], start=start, end=end)
# stock_prices = stock_prices.filter(like='Adj Close')
# stock_prices.columns = ['MSFT','AAPL']
#
# # do more stuff...
# ```
#
# ````
#
# ````{tabbed} Good
#
# Now I only need to change the variable `stocks` to alter the entire code.
#
# ```python
# import pandas_datareader as pdr
# from datetime import datetime
#
# # load stock returns
# start = datetime(2004, 1, 1)
# end = datetime(2007, 12, 31)
# stocks = ['MSFT','AAPL']
# stock_prices = pdr.get_data_yahoo(stocks, start=start, end=end)
# stock_prices = stock_prices.filter(like='Adj Close')
# stock_prices.columns = stocks
#
# # do more stuff...
# ```
#
# ````
| content/02/10_Golden_8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/0/05/Scikit_learn_logo_small.svg/1200px-Scikit_learn_logo_small.svg.png" width=50%>
#
# # Scikit-Learn에서의 Decision Tree 예제
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#라이브러리-가져오기" data-toc-modified-id="라이브러리-가져오기-1"><span class="toc-item-num">1 </span>라이브러리 가져오기</a></span></li><li><span><a href="#하이퍼파라미터-설정하기" data-toc-modified-id="하이퍼파라미터-설정하기-2"><span class="toc-item-num">2 </span>하이퍼파라미터 설정하기</a></span></li><li><span><a href="#데이터-불러오기" data-toc-modified-id="데이터-불러오기-3"><span class="toc-item-num">3 </span>데이터 불러오기</a></span><ul class="toc-item"><li><span><a href="#데이터-설명-확인하기" data-toc-modified-id="데이터-설명-확인하기-3.1"><span class="toc-item-num">3.1 </span>데이터 설명 확인하기</a></span></li><li><span><a href="#데이터의-독립변수-확인" data-toc-modified-id="데이터의-독립변수-확인-3.2"><span class="toc-item-num">3.2 </span>데이터의 독립변수 확인</a></span></li><li><span><a href="#데이터의-종속변수-확인" data-toc-modified-id="데이터의-종속변수-확인-3.3"><span class="toc-item-num">3.3 </span>데이터의 종속변수 확인</a></span></li><li><span><a href="#데이터-시각화하기" data-toc-modified-id="데이터-시각화하기-3.4"><span class="toc-item-num">3.4 </span>데이터 시각화하기</a></span></li></ul></li><li><span><a href="#모델링하기" data-toc-modified-id="모델링하기-4"><span class="toc-item-num">4 </span>모델링하기</a></span><ul class="toc-item"><li><span><a href="#Decision-Tree-학습하기" data-toc-modified-id="Decision-Tree-학습하기-4.1"><span class="toc-item-num">4.1 </span>Decision Tree 학습하기</a></span></li></ul></li><li><span><a href="#추론하기" data-toc-modified-id="추론하기-5"><span class="toc-item-num">5 </span>추론하기</a></span><ul class="toc-item"><li><span><a href="#Decision-Tree의-노드-확인하기" data-toc-modified-id="Decision-Tree의-노드-확인하기-5.1"><span class="toc-item-num">5.1 </span>Decision Tree의 노드 확인하기</a></span></li></ul></li><li><span><a href="#참고자료" data-toc-modified-id="참고자료-6"><span class="toc-item-num">6 </span>참고자료</a></span></li></ul></div>
# -
# ## 라이브러리 가져오기
# +
import os
import sys
import random
import datetime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.tree import DecisionTreeClassifier, plot_tree
# -
# ## 하이퍼파라미터 설정하기
# Parameters
n_classes = 3
plot_colors = "ryb"
plot_step = 0.02
# ## 데이터 불러오기
# Load data
iris = load_iris()
# ### 데이터 설명 확인하기
print(iris.DESCR)
# ### 데이터의 독립변수 확인
iris.feature_names # 4C2 = 6개쌍의 관계를 살펴볼 것
iris.data
iris.data.shape
# ### 데이터의 종속변수 확인
iris.target_names
iris.target.shape
iris_class = {n:c for n,c in enumerate(iris.target_names)}
iris_class
# ### 데이터 시각화하기
# +
plt.figure(figsize=(12, 8))
for pairidx, pair in enumerate([[0, 1], [0, 2], [0, 3],
[1, 2], [1, 3], [2, 3]]):
# We only take the two corresponding features
X = iris.data[:, pair]
y = iris.target
# Train
clf = DecisionTreeClassifier().fit(X, y)
# Plot the decision boundary
plt.subplot(2, 3, pairidx + 1)
plt.tight_layout(h_pad=0.5, w_pad=0.5, pad=2.5)
plt.xlabel(iris.feature_names[pair[0]])
plt.ylabel(iris.feature_names[pair[1]])
# Plot the training points
for i, color in zip(range(n_classes), plot_colors):
idx = np.where(y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=color, label=iris.target_names[i],
cmap=plt.cm.RdYlBu, edgecolor='black', s=15)
plt.suptitle("Decision surface of a decision tree using paired features")
plt.legend(loc='lower right', borderpad=0, handletextpad=0)
plt.axis("tight")
plt.show()
# -
# ## 모델링하기
# ### Decision Tree 학습하기
# +
plt.figure(figsize=(12, 8))
for pairidx, pair in enumerate([[0, 1], [0, 2], [0, 3],
[1, 2], [1, 3], [2, 3]]):
# We only take the two corresponding features
X = iris.data[:, pair]
y = iris.target
# Train
clf = DecisionTreeClassifier().fit(X, y)
# Plot the decision boundary
plt.subplot(2, 3, pairidx + 1)
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, plot_step),
np.arange(y_min, y_max, plot_step))
plt.tight_layout(h_pad=0.5, w_pad=0.5, pad=2.5)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
cs = plt.contourf(xx, yy, Z, cmap=plt.cm.RdYlBu)
plt.xlabel(iris.feature_names[pair[0]])
plt.ylabel(iris.feature_names[pair[1]])
# Plot the training points
for i, color in zip(range(n_classes), plot_colors):
idx = np.where(y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=color, label=iris.target_names[i],
cmap=plt.cm.RdYlBu, edgecolor='black', s=15)
plt.suptitle("Decision surface of a decision tree using paired features")
plt.legend(loc='lower right', borderpad=0, handletextpad=0)
plt.axis("tight")
plt.show()
# -
# ## 추론하기
clf = DecisionTreeClassifier().fit(iris.data,iris.target)
_input = [10 , 0, 1, 10] # 원하는 값을 입력해보세요하지만
y_pred = clf.predict([_input])
iris_class[y_pred[0]]
# ### Decision Tree의 노드 확인하기
plt.figure(figsize=(15, 10))
clf = DecisionTreeClassifier().fit(iris.data, iris.target)
plot_tree(clf, filled=True)
plt.show()
# +
plt.figure(figsize=(12, 8))
for pairidx, pair in enumerate([[0, 1], [0, 2], [0, 3],
[1, 2], [1, 3], [2, 3]]):
# We only take the two corresponding features
X = iris.data[:, pair]
y = iris.target
# Train
clf = DecisionTreeClassifier().fit(X, y)
# Plot the decision boundary
plt.subplot(2, 3, pairidx + 1)
plt.tight_layout(h_pad=0.5, w_pad=0.5, pad=2.5)
plt.xlabel(iris.feature_names[pair[0]])
plt.ylabel(iris.feature_names[pair[1]])
# Plot the training points
for i, color in zip(range(n_classes), plot_colors):
idx = np.where(y == i)
plt.scatter(X[idx, 0], X[idx, 1], c=color, label=iris.target_names[i],
cmap=plt.cm.RdYlBu, edgecolor='black', s=15, alpha=0.2)
plt.scatter(_input[pair[0]], _input[pair[1]], c='black',
cmap=plt.cm.RdYlBu, edgecolor='black', s=50)
plt.suptitle("Decision surface of a decision tree using paired features")
plt.legend(loc='lower right', borderpad=0, handletextpad=0)
plt.axis("tight")
plt.show()
# -
# ## 참고자료
# - https://scikit-learn.org/stable/auto_examples/tree/plot_iris_dtc.html#sphx-glr-auto-examples-tree-plot-iris-dtc-py
| code/Day04/Day04_08_DecisionTree_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import argparse
#from Data.stock_choices import list_of_stocks
#from Visualize_Prediction.make_visualization import visualization
# -
class visualize():
'''
The goal of this function is to use the user's arguments and provide a correct
output
'''
def parser(self):
'''
The goal of this function is to add the necessary arguments to the parser
'''
visualize_parser = argparse.ArgumentParser()
#Add the Arguments
# 1. The user can choose if they want to see the predicted values for the stock or not
# 2. The user can choose what stock they want to be displayed
visualize_parser.add_argument('--data_display')
visualize_parser.add_argument('--choice')
self.arguments = visualize_parser.parse_args()
stock_choice, type = self.data_from_arguments()
return stock_choice, type
def data_from_arguments(self):
'''
The goal of this function is to read the input, and know what to do next
'''
#Check to see if the stock was an acceptable input
length = len(list_of_stocks)
for i in list_of_stocks:
if self.arguments.choice == i:
self.stock_choice = i
# Break if the stock was acceptable
break
length -= 1
#This gets activated if the stock input was not valid
if length == 0:
print("")
print("Enter a valid stock!")
sys.exit(0)
# Decides if the user wants to see the forecasted values or not
# Assigns that information to a global variable
if self.arguments.data_display == "forecast":
self.type = "forecast"
elif self.arguments.data_display == "simple":
self.type = "simple"
else:
#This activates if the user did not provide an acceptable input for the
# --data_display argument
print("Use one of these commands:")
print("")
print("--data_display [forecast] allows you to see the predicted prices,")
print("--data_display [simple] allows you to see the normal prices")
sys.exit(0)
return self.stock_choice, self.type
#if __name__ == "__main__":
# start_visualize = visualize()
# stock_choice, type = start_visualize.parser()
#Calls a differnt file to visualize based on the information provided by the user
# show_visualization = visualization(stock_choice, type)
| ! Dissertation/ARIMA Prediction/display_main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python3
# ---
# + tags=[]
from itertools import combinations_with_replacement
r = 0
for blackArray in combinations_with_replacement(["A", "B", "C", "D"], 6):
for whiteArray in combinations_with_replacement(["A", "B", "C", "D"], 6):
ab = blackArray.count("A")
bb = blackArray.count("B")
cb = blackArray.count("C")
db = blackArray.count("D")
aw = whiteArray.count("A")
bw = whiteArray.count("B")
cw = whiteArray.count("C")
dw = whiteArray.count("D")
if (bb + bw) > 0 and (cb +cw) > 0 and (db + dw) > 0:
if ab >= 4 and ab > aw:
moreBlack = 0
if bb > bw:
moreBlack += 1
if cb > cw:
moreBlack += 1
if db > dw:
moreBlack += 1
if moreBlack == 1:
r += 1
# print(blackArray, whiteArray)
# print(ab, bb, cb, db)
# print(aw, bw, cw, dw)
print("result", r)
# -
# 답 201
| 2021_Math/m29.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''web'': conda)'
# name: python385jvsc74a57bd0936961605949c8af6a082776a99176412ce25745978a797c3acd398b4d5851fa
# ---
from IPython.display import Image
import dis
# # Chater4. 사전과 셋
# 배울 내용
#
# * 사전과 셋의 용도
# * 사전과 셋의 유사점
# * 사전의 오버페드
# * 사전의 성능을 최적화 하는 방법
# * 파이썬에서 사전을 사용해서 네임스페이스를 유지하는 방법
# hashable 하다
#
# 사전과 셋은 정렬되지 않음.
# 고유하게 참조할 수 있는 별도 객체가 있는 상황에서 사용
#
# 참조하는 객체는 일반적으로는 문자열이고 hashable하다면 어떤 타입도 상관없음.
#
# 참조객체 : key
# 데이터 : value
#
# 사전과 셋은 유사하나 셋에는 value가 없다.
# 셋은 유일한 키를 저장하는 자료구조이다.
# ### Hash Function : 임의의 길이를 갖는 임의의 데이터를 고정된 길이의 데이터로 매핑하는 함수. 아무리 큰 숫자를 넣더라도 정해진 크기의 숫자가 나오는 함수
#
# ex) 수를 나누었을 때 나머지를 구하는 함수
#
# ### Hash Value, Hash Code, Hash Sum, Check Sum : 이러한 hash function을 적용해서 나온 고정된 길이의 값
#
# hash(object): object의 저장된 내용을 기준으로 한 개의 정수를 생성하여 반환하는 함수
#
# - 이 함수는 hasher라고도 부른다. hash 함수안에 인자로 들어가는 object은 하나의 값일 수도 있고, 여러 개의 값을 갖고 있는 컨테이너 일 수도 있다. 즉, 내용이 뭐든지간에(사실, 조건이 없진 않다..), 이에 해당하는 숫자 하나를 무조건 생성한다고 생각하면 된다. 아래의 예시처럼, string이 무엇이든 간에 hash function을 통과하고나면 무조건 하나의 숫자로 반환되는 함수를 의미한다. 이는 튜플일 수도 있는데 (1, 2, 3) 을 hash function에 넣어, 2366528764324이 나왔다고 하면, 이 튜플은 hashable object 라고 부른다.
#
#
# ### hashable은 왜 필요할까?
#
# 비교를 위해 사용된다. 즉, hash을 하고나면, 각 객체가 숫자로 나오기 때문에, 같은 객체인지 다른객체인지 비교가 가능하다. 같은 숫자면 같은 객체로 인식한다. 또한, 컨테이너인 객체들도, hash을 이용하면 한번에 비교가 가능하다. 가령 (a, b, c)인 객체와 (d, b, c)인 객체를 비교할 대도, 각각의 원소들이 같은지 여러번 비교를 하는 것이 아니라, hash 값만 있으면 비교가 가능하므로, 한 번의 비교 연산만이 필요하다. 즉, 1) Computationally robust함을 유지하기 하기위해 사용된다 (쉽게 말하면, 다른 객체는 다른 해시함수를 갖어야한다). 다른 객체들이 같은 해시를 가지면 충돌이 일어날 수 있다. 2) 같은 오브젝트가 일관성 있게, 같은 값을 가질 수 있도록 표식한다.
#
#
# ### [ HashTable(해시테이블)이란? ]
# 해시 테이블은 (Key, Value)로 데이터를 저장하는 자료구조 중 하나로 빠르게 데이터를 검색할 수 있는 자료구조이다. 해시 테이블이 빠른 검색속도를 제공하는 이유는 내부적으로 배열(버킷)을 사용하여 데이터를 저장하기 때문이다. 해시 테이블은 각각의 Key값에 해시함수를 적용해 배열의 고유한 index를 생성하고, 이 index를 활용해 값을 저장하거나 검색하게 된다. 여기서 실제 값이 저장되는 장소를 버킷 또는 슬롯이라고 한다.
#
#
#
#
# 예를 들어 우리가 (Key, Value)가 ("<NAME>", "521-1234")인 데이터를 크기가 16인 해시 테이블에 저장한다고 하자. 그러면 먼저 index = hash_function("<NAME>") % 16 연산을 통해 index 값을 계산한다. 그리고 array[index] = "521-1234" 로 전화번호를 저장하게 된다.
#
# 어라훈 구조로 데이터를 저장하면 Key값으로 데이터를 찾을 때 해시 함수를 1번만 수행하면 되므로 매우 빠르게 데이터를 저장/삭제/조회할 수 있다. 해시테이블의 평균 시간복잡도는 O(1)이다.
#
#
#
#
# hash가 가능한 타입 : __hash__ magic function, __eq__ 혹은 __cmp__ magic function을 구현한 타입 --> 추가
# ### 파이썬 내장 타입은 이를 모두 구현한다 ???
hash((1, 2, 3))
hash([1, 2, 3])
hash({1:2, 3:4})
hash(set([1, 2, 3]))
# 사전과 셋은 주어진 색인을 O(1) 시간 복잡도 안에 찾아준다.
# 삽입 연산의 시간복잡도는 리스트/튜플과 같은 O(1) 이다.
#
# 또한 이 시간복잡도는 개방 주소 open address 해시 테이블을 사용했을 경우이다. --> 추가
# 단점 :
# * 사전과 셋은 메모리를 많이 사용한다.
# * 사용하는 해시 함수에 속도를 전적으로 의존한다 . --> 추가
# * 해시 함수가 느리다면 사전과 셋의 연산도 느려진다.
# +
# 예제 4-1 리스트를 이용한 전화번호 검색
def find_phonenumber(phonebook, name):
for n, p in phonebook:
if n == name:
return p
return None
phonebook = [
("<NAME>", "555-5555-5555"),
("<NAME>", "212-555-5555")
]
print(f"<NAME>'s phone number is {find_phonenumber(phonebook, '<NAME>')}")
# -
# ### 만약 이 리스트를 정렬한 다음에 bisect 모듈을 이용하면 O(log n)의 시간복잡도로 검색할 수 있다.
# +
# 예제 4-2 사전을 이용한 전화번호 검색
phonebook = {
"<NAME>": "555-5555-5555",
"<NAME>" : "212-555-5555"
}
print(f"<NAME>'s phone number is {phonebook['<NAME>']}")
# -
# 색인으로 이름을 찾으면 이렇게 전화번호를 받을 수 있다.
# 전체 데이터를 살펴보는 대신 직접 참조를 통해 필요한 값을 가져올 수 있다.
# +
# 예제 4-3 리스트와 셋에서 유일한 이름 찾기
def list_unique_names(phonebook):
unique_names = []
for name, phonenumber in phonebook:
first_name, last_name = name.split(" ", 1)
for unique in unique_names:
if unique == first_name:
break
else:
unique_names.append(first_name)
return len(unique_names)
def set_unique_names(phonebook):
unique_names = set()
for name, phonenumber in phonebook:
first_name, last_name = name.split(" ", 1)
unique_names.add(first_name)
return len(unique_names)
# +
phonebook = [
("<NAME>", "555-555-5555"),
("<NAME>", "202-555-5555"),
("<NAME>", "212-555-5555"),
("<NAME>", "647-555-5555"),
("<NAME>", "301-555-5555")
]
print("Number of unique names from set method", set_unique_names(phonebook))
print("Number of unique names from list method", list_unique_names(phonebook))
# -
# ### 리스트를 이용한 알고리즘의 시간복잡도는 O(n^2) 이다.
#
# 중복되는 이름이 없는 최악의 상황에는 크기가 phonebook만큼 커진다.
# 이는 계속 커지는 리스트에서 전화번호부의 각 이름을 선형 탄색하는 작업이다.
# ### 셋을 이용한 최종 알고리즘의 시간 복잡도는 O(n)이다.
#
# set.add 연산은 전화번호부의 크기에 상관없이 O(1) 시간복잡도로 수행된다.
# 상수 시간에 수행되지 않는 연산은 전화번호부를 순회하는 루프뿐이다.
# %timeit list_unique_names(phonebook)
# %timeit set_unique_names(phonebook)
# ## 4.1 사전과 셋의 동작 원리
#
# 사전과 셋은 모두 해시 테이블을 사용해서 시간복잡도가 O(1) 이다 -> 추가
# 임의의 키를 리스트의 색인으로 변화하는 해시함수 사용
# ### 4.1.1 삽입과 검색
#
# 해시 테이블을 처음 생성하면 배열을 사용할 때 처럼 메모리부터 할당한다.
# 배열에서의 데이터 추가 : 사용하지 않은 메모리 블록을 찾아서 데이터를 추가하고 필요할 때 크기를 조정
# 해시 테이블 에서의 데이터 추가 : 이 연속적인 메모리에 데이터를 나열할 방법을 생각해 봐야함 --> 추가
# 새로운 데이터의 위치는 키의 해시값, 데이터의 값을 다른 객체와 비교하는 방법으로 결정된다.
#
# ### Data -> Hash Function -> Hash Value -> Masking -> Index ??
#
# ### 파이썬은 키/값 데이터를 표준배열(?) 에 덧붙이고 오직 이 배열의 색인만 해시 테이블에 저장한다.
# ### 이 덕분에 메모리 사용량이 30 ~ 95% 줄어든다.
#
# 새로운 색인은 단순한 선형 함수를 이용해서 계산하는데 이를 Probing이라고 한다.
# 파이썬의 Probing 메커니즘은 원래 해시값에서 더 상위 Bit를 활용한다. ---> 추가
# +
# 예제 4-4 사전 탐색 과정
def index_sequence(key, mask=0b111, PERTURB_SHIFT = 5):
perturb = hash(key)
i = perturb & mask
yield i
while True:
perturb >>= PERTURB_SHIFT
i = (i * 5 + perturb + 1) & mask
yield i
# -
Image("./img/4-1.png")
# +
def City(str):
def __hash__(self):
return ord(self[0])
# 임의 값의 도시 이름으로 사전 생성
data = {
City("Rome") : "Italy",
City("San Francisco") : "USA",
City("New York") : "USA",
City("Barcelona") : "Spain"
}
# -
a = City("Barcelona")
b = City("Rome")
a.__hash__ == b.__hash__
# ### 4.1.2 삭제
#
# 해시 테이블에서 값을 삭제할 때 단순히 해당 메모리 블록을 NULL 로 만드는 방법은 사용할 수 없다.
# NULL 을 Probing 시 해시 충돌을 위한 감싯값 Sentinel Value로 사용하기 때문이다. --> 추가
# 따라서 해당 블록이 비었음을 나타내는 특수한 값을 기록해두고 나중에 해시 충돌을 해결하는 과정에서 이 값을 활용해야 한다.
# 예를 들어 Rome이 삭제된 다음에 Barcelona를 검색하면 처음에는 Rome이 들어있던 블록이 비었음을 나태는 값이 있는 배열 항목을 먼저 만난다.
# 값을 찾지 못했다고 멈추는 대신, index_sequence 함수가 반환하는 다음 색인을 검색해야 한다.
# 해시 테이블의 빈 슬롯은 새로운 값이 쓰이거나 해시 테이블의 크기가 변경될 때 삭제된다.
# ### 4.1.3 크기 변경
#
# 해시 테이블의 2/3 이하만 채워진다면 충돌 횟수와 공간 활용이 적절함.
#
# 크기를 변경할 때는 충분히 큰 해시 테이블을 할당하고 (더 많은 메모리를 할당하고 ) 그 크기에 맞게 마스크값을 조정한다.
# -> 이후 모든 항목을 새로운 해시 테이블로 옮기는데
# 이 과정에서 바뀐 마스크값 때문에 색인을 새로 계산해야 하기 때문에 해시 테이블 크기 변경은 비싼 작업이다.
#
# 그러나 크기 변경은 여유공간이 아주 적을 때만 수행되기 때문에 보통 개별항목 추가는 시간복잡도가 O(1) 이다.
#
#
# 기본적으로 dicttionay 나 set의 최소 크기는 8이다. 그리고 사전이 2/3만큼 찰 때마다 크기를 3배 늘린다.
#
# 8, 18, 39, 81, 165, 333, 669.....
# ### 4.1.4 해시 함수와 엔트로피
#
# 파이썬 객체는 __hash__와 __cmp__ 함수를 구현하므로 해시가 가능.
# 튜플, 문자열, int, float
#
hash("str")
hash(5)
hash(5.0)
int(5).__hash__
# __hash__ 함수는 내장 함수인 id 함수를 이용해서 객체의 메모리 위치를 반환한다.
# __cmp__ 연산자는 객체의 메모리 위치를 산술 비교
class Point(object):
def __init__(self, x, y):
self.x, self.y = x, y
# 만약 x, y값이 동일한 Point 객체를 여러 개 생성하면
# 메모리에서 각 객체는 서로 다른 위치에 있으므로 해시값이 모두 다르다.
# 이 객체들을 모두 같은 set에 추가한다면 각각의 항목이 추가된다.
p1 = Point(1, 1)
p2 = Point(1, 1)
set([p1, p2])
Point(1, 1) in set([p1, p2])
# 다 다른 것을 알 수 있음.
class Point2(object):
def __init__(self, x, y):
self.x, self.y = x, y
def __hash__(self):
return hash((self.x, self.y))
def __eq__(self, other):
return self.x == other.x and self.y == other.y
# 이 해시함수는 같은 내용의 객체에 대해서는 항상 같은 결과를 반환한다.
# 인스턴스화한 객체의 메모리 주소가 아니라 Point 객체의 속성으로 dictionary 나 set에 필요한 색인을 만들 수 있다.
#
p3 = Point2(1, 1)
p4 = Point2(1, 1)
set([p3, p4])
Point2(1, 1) in set([p3, p4])
# ### 사용자 정의 해시 함수에서 충돌을 피하려면 해시값이 균일하게 분포되도록 신경 써야 한다.
# ### 충돌이 잦으면 해시 테이블의 성능에 악영향을 끼친다.
# ### 사전의 모든 키가 충돌하는 "최악의 상황" 에는 사전의 탐색 성능이 리스트와 같은 O(n)이 된다. (입력된 n의 크기와 같은)
# +
# 예제 4-6 두 소문자를 조합한 최적 해시 함수
def twoletter_hash(key):
offset = ord('a')
k1, k2 = key
return (ord(k2), - offset) + 26 * (ord(k1) - offset)
# +
# 예제 4-7 좋은 해시함수와 나쁜 해시함수의 시간 차이
import string
import timeit
class BadHash(str):
def __hash__(self):
return 42
class GoodHash(str):
def __hash__(self):
"""
아래는 twoletter hash 함수를 약간 개선한 버전
"""
return ord(self[1]) + 26 * ord(self[0]) - 2619
# +
baddict = set()
gooddict = set()
for i in string.ascii_lowercase:
for j in string.ascii_lowercase:
key = i + j
baddict.add(BadHash(key))
gooddict.add(GoodHash(key))
badtime = timeit.repeat(
"key in baddict",
setup = "from __main__ import baddict, BadHash; key = BadHash('zz')",
repeat = 3,
number = 1_000_000
)
goodtime = timeit.repeat(
"key in gooddict",
setup = "from __main__ import gooddict, GoodHash; key = GoodHash('zz')",
repeat = 3,
number = 1_000_000
)
print(f"Min lookup time for baddict: {min(badtime)}")
print(f"Min lookup time for gooddict: {min(goodtime)}")
# -
# ## 4.2 사전과 네임스페이스
# 파이썬에서 객체, 함수 모듈 찾을 때 찾는 순서 :
# ### locals() -> globals() -> __builtin__
#
# locals(), globals() : dictionary
# __builtin__ : 모듈 객체
#
# __builtin__ 은 모듈 내부에서 locals() dictionary를 탐색하여 특정 속성을 찾는다.
#
# +
# 예제 4-8 네임스페이스 탐색
import math
from math import sin
def test1(x):
res = 1
for _ in range(1000):
res += math.sin(x)
return res
def test2(x):
res = 1
for _ in range(1000):
res += sin(x)
return res
def test3(x):
res = 1
for _ in range(1000):
res += sin(x)
return res
# -
dis.dis(test1)
| chap4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
from tensorflow.compat.v1 import keras as keras
import keras.backend as K
print(tf.__version__)
import numpy as np
import utils
import loggingreporter
cfg = {}
cfg['SGD_BATCHSIZE'] = 128
cfg['SGD_LEARNINGRATE'] = 0.001
cfg['NUM_EPOCHS'] = 1000
#cfg['ACTIVATION'] = 'relu'
cfg['ACTIVATION'] = 'tanh'
# How many hidden neurons to put into each of the layers
# cfg['LAYER_DIMS'] = [1024, 20, 20, 20]
cfg['LAYER_DIMS'] = [32, 28, 24, 20, 16, 12, 8, 8]
#cfg['LAYER_DIMS'] = [128, 64, 32, 16, 16] # 0.967 w. 128
#cfg['LAYER_DIMS'] = [20, 20, 20, 20, 20, 20] # 0.967 w. 128
ARCH_NAME = '-'.join(map(str,cfg['LAYER_DIMS']))
trn, tst = utils.get_mnist()
# Where to save activation and weights data
cfg['SAVE_DIR'] = 'rawdata/' + cfg['ACTIVATION'] + '_' + ARCH_NAME
# +
input_layer = keras.layers.Input((trn.X.shape[1],))
clayer = input_layer
for n in cfg['LAYER_DIMS']:
clayer = keras.layers.Dense(n, activation=cfg['ACTIVATION'])(clayer)
output_layer = keras.layers.Dense(trn.nb_classes, activation='softmax')(clayer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
optimizer = keras.optimizers.SGD(lr=cfg['SGD_LEARNINGRATE'])
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# +
def do_report(epoch):
# Only log activity for some epochs. Mainly this is to make things run faster.
if epoch < 20: # Log for all first 20 epochs
return True
elif epoch < 100: # Then for every 5th epoch
return (epoch % 5 == 0)
elif epoch < 200: # Then every 10th
return (epoch % 10 == 0)
else: # Then every 100th
return (epoch % 100 == 0)
# reporter = loggingreporter.LoggingReporter(cfg=cfg,
# trn=trn,
# tst=tst,
# do_save_func=do_report)
r = model.fit(x=trn.X, y=trn.Y,
verbose = 2,
batch_size = cfg['SGD_BATCHSIZE'],
epochs = cfg['NUM_EPOCHS'])
# validation_data=(tst.X, tst.Y),
# callbacks = [reporter,])
# -
| MNIST_SaveActivations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="mw2VBrBcgvGa"
# # Load Keras Datasets
# -
import tensorflow as tf
# ### Load Dataset
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
# ### Shape of Data
# +
# shape of training images
print('x_train shape :', x_train.shape)
# shape of testing images
print('x_test shape :', x_test.shape)
# -
# ### Normalize Data
x_train = x_train / 255.0
x_test = x_test / 255.0
# ### Plot Sample Image
# +
import matplotlib.pyplot as plt
plt.imshow(x_train[0], cmap = 'gray')
| coding-exercise/week3/part1/exploring-mnist-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Load everything
# Import the libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image, ImageChops
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis, QuadraticDiscriminantAnalysis
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.metrics import make_scorer, confusion_matrix, accuracy_score, recall_score, precision_score, classification_report
# +
# Read the data files
train_data = pd.read_csv("data/train.csv")
test_data = pd.read_csv("data/test.csv")
print(train_data.head())
# +
# Slice the data and create X, y and test_data
X = train_data.ix[:, 1:].values
y = train_data.ix[:, 0].values
test_data = test_data.values
print(X.shape)
print(y.shape)
# +
# Visualize a digit
# %matplotlib inline
fig, ax = plt.subplots(ncols=2, figsize=(8,8))
ax1, ax2 = ax
original_im = X[7].reshape((int(np.sqrt(X[7].shape[0])), int(np.sqrt(X[7].shape[0]))))
new_im = Image.fromarray(original_im.astype('uint8')).convert("1") # Convert the values to 8-bit and convert the image to 1-bit black and white
new_im = ImageChops.invert(new_im)
ax1.imshow(original_im)
ax2.imshow(new_im)
ax1.set_title("Original Image")
ax2.set_title("Modified Image")
plt.show()
# +
# Preprocess the images
def image_preprocessing(arr):
arr = arr.reshape((int(np.sqrt(arr.shape[0])), int(np.sqrt(arr.shape[0])))).astype('uint8')
im = Image.fromarray(arr).convert("1") # Convert the values to 8-bit and convert the image to 1-bit black and white
im = ImageChops.invert(im)
new_arr = np.array(im.getdata())
return new_arr
X = np.apply_along_axis(image_preprocessing, 1, X)
test_data = np.apply_along_axis(image_preprocessing, 1, test_data)
# +
# Verify the images
fig, ax = plt.subplots(ncols=2, figsize=(8,8))
ax1, ax2 = ax
ax1.imshow(Image.fromarray(X[7].reshape((int(np.sqrt(X[7].shape[0])), int(np.sqrt(X[7].shape[0]))))))
ax2.imshow(Image.fromarray(test_data[7].reshape((int(np.sqrt(test_data[7].shape[0])), int(np.sqrt(test_data[7].shape[0]))))))
ax1.set_title("Train Image")
ax2.set_title("Test Image")
plt.show()
# -
# ### Preprocessing
# +
# Scale the data
X_scale = StandardScaler()
X = X_scale.fit_transform(X)
test_data = X_scale.transform(test_data)
# -
# ### Model Selection
# +
# Split partial data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=3)
print(X_train.shape, X_test.shape)
print(y_train.shape, y_test.shape)
# +
# Select model/classifier
names = [
"Logistic Regression",
"Nearest Neighbors",
"Linear SVM",
"RBF SVM",
"Gaussian Process",
"Decision Tree",
"Random Forest",
"Neural Net",
"AdaBoost",
"Naive Bayes",
"LDA",
"QDA",
"GBC"
]
classifiers = [
LogisticRegression(),
KNeighborsClassifier(3),
SVC(kernel="linear", C=0.025, random_state=3),
SVC(gamma=2, C=1, random_state=3),
GaussianProcessClassifier(warm_start=True, random_state=3),
DecisionTreeClassifier(max_depth=5, random_state=3),
RandomForestClassifier(max_depth=5, n_estimators=100, random_state=3),
MLPClassifier(alpha=1, max_iter=5000, random_state=3),
AdaBoostClassifier(random_state=3),
GaussianNB(),
LinearDiscriminantAnalysis(),
QuadraticDiscriminantAnalysis(),
GradientBoostingClassifier(random_state=3)
]
for name, clf in zip(names, classifiers):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
score = recall_score(y_pred, y_test, average="micro")
print("{}: {}".format(name, score))
# -
# ### Model Creation
# Create the final model/classifier for test
clf = RandomForestClassifier(500, random_state=3)
clf.fit(X_train, y_train)
# +
# Compute accuracy and create confusion matrix
y_pred = clf.predict(X_test)
accuracy = accuracy_score(y_test, y_pred)
cm = confusion_matrix(y_test, y_pred)
cr = classification_report(y_test, y_pred)
print("Accuracy: {}%".format(round(accuracy*100), 1))
print("Confusion Matrix: ")
print(cm)
print("Classification Report: ")
print(cr)
# -
# ### Prediction
# Create the final model/classifier for prediction
clf = RandomForestClassifier(500, random_state=3)
clf.fit(X, y)
# Predict and save the predicted data to csv
prediction = clf.predict(test_data)
prediction = pd.DataFrame({"ImageId": range(1, len(prediction)+1), "Label": prediction})
prediction.to_csv("data/prediction.csv", index=None)
# ### Kaggle Accuracy Result: 96.7% (0.96714)
| Digit Recognizer/digit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''py38'': conda)'
# language: python
# name: python3
# ---
import numpy as np
n = 6
a = np.array(range(n ** 4)).reshape((n,) * 4)
print(a)
# +
b = a.reshape((n * n,) * 2)
print(b)
# -
c = a.transpose([1, 0, 2, 3])
print(c)
d = c.reshape((n * n,) * 2)
print(d)
print(a[1, 0, 1, 1])
print(c[0, 1, 1, 1])
print(np.argwhere(b==223))
print(np.argwhere(d==223))
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Data Visualization Home Page](https://www.kaggle.com/learn/data-visualization)**
#
# ---
#
# In this exercise, you'll explore different chart styles, to see which color combinations and fonts you like best!
#
# ## Setup
#
# Run the next cell to import and configure the Python libraries that you need to complete the exercise.
import pandas as pd
pd.plotting.register_matplotlib_converters()
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
print("Setup Complete")
# The questions below will give you feedback on your work. Run the following cell to set up our feedback system.
# Set up code checking
import os
if not os.path.exists("../input/spotify.csv"):
os.symlink("../input/data-for-datavis/spotify.csv", "../input/spotify.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.data_viz_to_coder.ex6 import *
print("Setup Complete")
# You'll work with a chart from the previous tutorial. Run the next cell to load the data.
# +
# Path of the file to read
spotify_filepath = "../input/spotify.csv"
# Read the file into a variable spotify_data
spotify_data = pd.read_csv(spotify_filepath, index_col="Date", parse_dates=True)
# -
# # Try out seaborn styles
#
# Run the command below to try out the `"dark"` theme.
# +
# Change the style of the figure
sns.set_style("dark")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# Mark the exercise complete after the code cell is run
step_1.check()
# -
# Now, try out different themes by amending the first line of code and running the code cell again. Remember the list of available themes:
# - `"darkgrid"`
# - `"whitegrid"`
# - `"dark"`
# - `"white"`
# - `"ticks"`
#
# This notebook is your playground -- feel free to experiment as little or as much you wish here! The exercise is marked as complete after you run every code cell in the notebook at least once.
#
# ## Keep going
#
# Learn about how to select and visualize your own datasets in the **[next tutorial](https://www.kaggle.com/alexisbcook/final-project)**!
# +
# Change the style of the figure
sns.set_style("darkgrid")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# +
# Change the style of the figure
sns.set_style("whitegrid")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# +
# Change the style of the figure
sns.set_style("white")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# +
# Change the style of the figure
sns.set_style("ticks")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# -
# ---
# **[Data Visualization Home Page](https://www.kaggle.com/learn/data-visualization)**
#
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161291) to chat with other Learners.*
| Kaggle/Data Visualization/exercise-choosing-plot-types-and-custom-styles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="f1g3BoLYJYgi"
# # <font color = green> Aula 1
#
# + [markdown] id="NObniNTKJYgk"
# ## Instalar pandas
# + id="bGa0u2fFJYgm" outputId="cd673391-a31b-4d00-c9a2-b5f62789855a"
# !pip install pandas
# + [markdown] id="0VTuf9YsJYgt"
# ## 1. 1. Importar bibliotecas
# + id="uYUgieujJYgu"
import pandas as pd
# + [markdown] id="gw20Hb5PJYgy"
# ## 1.2. Importar base de dados
# + id="UsVd5eORJYg0" outputId="c2ce2cfa-aa13-496d-81cf-bae8699580ef"
dados=pd.read_csv('dados/aluguel_rio.csv',sep=';')
dados.head(3)
# Obs: Lembrar de adicionar o separador ';'
# + id="06AATMhqJYg5" outputId="363be9cc-e64f-4efa-c898-4118fd538b9d"
dados.info()
# + [markdown] id="vGcwePD8JYg-"
# ## 1.3. Informações gerais da base de dados
# + id="1p-krBs4JYg-" outputId="7ebc3bdc-237a-4764-e01e-1f65da6f458d"
# Tipos de dados
dados.dtypes
# + id="ggk6f7uuJYhC"
#Mudar nome das colunas
tipos_de_dados=pd.DataFrame(dados.dtypes,columns=['Tipos de dados'])
# + id="XKnR_vNUJYhH" outputId="b67110ca-e6f2-4e44-e805-689bca462a8e"
#Mudar nome da primeira coluna
tipos_de_dados.columns.name='Variáveis'
tipos_de_dados
# + id="nOvFUl9XJYhM" outputId="f851c354-4a6d-4f89-ffac-25190c173c11"
#Tamanho do dataset
dados.shape
# + id="w_fWHp5HJYhQ" outputId="8d4ac241-5ca1-4f6e-9802-25defa1b3c84"
print('A base de dados tem {} entradas e {} variáveis'.format(dados.shape[0],dados.shape[1]))
# + id="xTzFeMJEJYhU" outputId="7677a6ac-214e-4e8a-d0b9-bb35a20b7d81"
#Descritiva dos dados
descritiva=dados.describe()
descritiva.columns.name='Estatística'
descritiva
# + [markdown] id="ZNVN41ccJYhY"
# # <font color = green> Aula 2
# (Tipos de movéis)
# + id="MlP5k1G8JYha" outputId="b6b9d763-e70b-4adc-9d23-904df3bd3f9c"
dados.head(3)
# + [markdown] id="z0blBEY0JYhf"
# ## 2.1. Acessar coluna epecifica
# + id="17At-ABEJYhf" outputId="d28c49e9-dbcc-4a95-824b-05dd089d5c15"
#Métódo 1
dados['Tipo']
# + id="gDNV5WF3JYhj" outputId="ab8dd3bf-f7db-4071-cd62-30ca44dc2ce9"
#Método 2
dados.Tipo
# + id="24EuZU6aJYhn" outputId="d261f434-66d9-48fb-d1e8-415b85fe5ede"
#Tipo de dado
tipo_de_imovel=dados.Tipo
type(tipo_de_imovel)
# + [markdown] id="spuKj3siJYhr"
# ## 2.2. Remover duplicatas
# + id="rX9M0npQJYhr"
tipo_de_imovel.drop_duplicates(inplace=True)
# Obs: o inplace modifica o fonte original de dados
# + id="6R_v8rqFJYhv" outputId="bdafa6b7-0cf5-478a-dfbd-eee9a4a2c24a"
tipo_de_imovel
# + [markdown] id="2UoyUTngJYh1"
# ## 2.3. Organizar visulização
# + id="KqBrw1UwJYh1" outputId="d8d04a23-57c5-4261-e713-44008c106e9a"
tipo_de_imovel=pd.DataFrame(tipo_de_imovel)
tipo_de_imovel
# + id="0jUGvlshJYh5" outputId="c08ae0b7-3f38-4768-e580-f1dd8086a3e2"
tipo_de_imovel.index
# + id="KlgS5yFgJYh8" outputId="0f02748f-4580-41fd-fed0-b0d99b9a2158"
range(tipo_de_imovel.shape[0])
# + id="O9t20ZjdJYiA" outputId="29ca2ee4-f03d-46ec-c6d3-49a9d75a2c90"
for i in range(tipo_de_imovel.shape[0]):
print(i)
# + id="-rUypi3iJYiH" outputId="182d95cf-0230-46a9-f1a2-89bf060a0fc0"
tipo_de_imovel.index=range(tipo_de_imovel.shape[0])
tipo_de_imovel.index
# + id="aVp71VsBJYiM" outputId="1f0bfa50-d93b-4d66-a58a-f3f5e9d98390"
tipo_de_imovel
# + id="mTI4KVLhJYiQ" outputId="8364615b-fc19-44e2-f949-e2b53eed7393"
tipo_de_imovel.columns.name='ID'
tipo_de_imovel
# + [markdown] id="LjtIBWwHJYiS"
# # <font color = green> Aula 3
# + [markdown] id="jrAV6uezJYiU"
# ## 3.1. Imovéis residenciais
# + id="vmKtHS4hJYiU"
#Os dados precisam se reiportados nessa etapa
import pandas as pd
dados = pd.read_csv('dados/aluguel_rio.csv', sep = ';')
# + id="WT6VnAh0JYiX" outputId="caaf6d98-1645-4c38-ee3d-26868f6c919c"
dados.head(10)
# + id="5Kttmvm4JYia" outputId="427bac1c-223f-4989-b87f-bb1e5619e0b2"
list(dados.Tipo.drop_duplicates())
# + id="ybFeoR23JYic" outputId="71fafdd5-b7fb-43cc-f569-e5ca78162b1c"
residencial = ['Quitinete',
'Casa',
'Apartamento',
'Casa de Condomínio',
'Casa de Vila']
residencial
# + id="GsfM2sjLJYif" outputId="c045d438-fb22-4942-e1ff-31ad0d101768"
dados.head(10)
# + id="xWir560KJYii" outputId="07f52813-c931-4b2f-f802-1a5146aaed1a"
#Método isin()
selecao = dados['Tipo'].isin(residencial)
selecao
# + id="FzDAGzsVJYim" outputId="688ba912-dcff-4e00-805d-a7f5be2cf71a"
dados_residencial = dados[selecao]
dados_residencial.index = range(dados_residencial.shape[0])
dados_residencial.head(10)
# + id="7WFZm-xxJYir" outputId="33d5f662-1a19-4905-a377-e17ea1c7ad1c"
print('Tamanho do data frame original : {}'.format(dados.shape[0]))
print('Tamanho do data frame residencial: {}'.format(dados_residencial.shape[0]))
# + [markdown] id="gDMU9Dm9JYiv"
# ## 3.2. Exportando base de dados
# + id="FrxBriEtJYiv" outputId="2d262170-4687-4fe2-c8c2-12e49b61d8ab"
#Gravar csv
dados_residencial.to_csv('dados/aluguel_residencial.csv', sep = ';')
#Ler e atribuir variável
dados_residencial_2 = pd.read_csv('dados/aluguel_residencial.csv', sep = ';')
dados_residencial_2.head(10)
#Obs: note que o indice do contador se repete
# + id="dZHkF0EVJYiy" outputId="83ac9c15-3748-49bf-efec-bb2b332cf406"
#Gravar csv sem duplicata de indice
dados_residencial.to_csv('dados/aluguel_residencial.csv', sep = ';',index = False)
#Ler e atribuir variável
dados_residencial_2 = pd.read_csv('dados/aluguel_residencial.csv', sep = ';')
dados_residencial_2.head(10)
# + [markdown] id="PIghOjG1JYi2"
# # <font color = green> Aula 4
#
# (Seleções e frequências)
#
# Tarefas:
#
# * Selecione somente os imóveis classificados com tipo 'Apartamento'.
#
# * Selecione os imóveis classificados com tipos 'Casa', 'Casa de Condomínio' e 'Casa de Vila'.
#
# * Selecione os imóveis com área entre 60 e 100 metros quadrados, incluindo os limites.
#
# * Selecione os imóveis que tenham pelo menos 4 quartos e aluguel menor que R$ 2.000,00.
# + [markdown] id="oT9mTYYWJYi2"
# ## 4.1. Importar base de dados criada
# + id="paxT64COJYi3" outputId="e1153002-6dc2-4269-f6a2-38a11176c082"
dados=dados_residencial_2 = pd.read_csv('dados/aluguel_residencial.csv', sep = ';')
dados_residencial_2.head(10)
# + [markdown] id="mVtyOC4HJYi7"
# ## 4.2. Selecionar somente os imóveis classificados com tipo Apartamento
# + id="3kPaCIx3JYi7" outputId="13ae5b83-0fa1-473a-f26d-73acf50a9b0a"
selecao=dados.Tipo=='Apartamento'
selecao
# + id="m6ixHj2IJYi_" outputId="74dd54bb-4aa9-4e5e-d637-573378c35780"
apartamentos=dados[selecao]
apartamentos.head(10)
# + [markdown] id="f2bXuFE2JYjD"
# ## 4.3. Selecionar os imóveis classificados com tipos Casa, Casa de Condomínio e Casa de Vila
# + id="TuWS4DCjJYjE" outputId="9b5dc831-67b6-4acc-badb-d2fe0ae7b8c9"
selecao=dados.Tipo!='Apartamento'
selecao
# + id="7bRFAGMtJYjJ" outputId="a801f15d-7503-4531-c243-dda094c724b7"
casas=dados[selecao]
casas.head(10)
# + [markdown] id="2TEvOwEBJYjO"
# ## 4.4. Selecionar os imóveis com área entre 60 e 100 metros quadrados, incluindo os limites
# + id="PzoJu5AfJYjO" outputId="112a47fa-2937-44b8-9e37-700f1a232d52"
selecao=(dados.Area>=60) & (dados.Area<=100)
selecao
# + id="YjDOlHdlJYjS" outputId="7d7cd366-bc88-4c11-8ca1-2679cae1c2e6"
areas=dados[selecao]
areas.head(10)
# + [markdown] id="rC98dxqPJYjV"
# ## 4.5. Selecionar os imóveis que tenham pelo menos 4 quartos e aluguel menor que R$ 2.000,00.
# + id="Zouu4PNRJYjW" outputId="c5618fb5-8413-4b64-eddb-fd838afc8386"
selecao=(dados.Quartos>=4) & (dados.Valor<2000)
selecao
# + id="y3U3o5s2JYja" outputId="dc5d9bec-e10d-4a7b-b888-7ee098901fc3"
quartos_aluguel=dados[selecao]
quartos_aluguel.head(10)
# + id="Wu6jo0wOJYje" outputId="d29a3377-06b6-4854-cb8a-daca45142242"
print('''
Nº de apartamentos: {}
Nº de de casas : {}
Nº de imóveis com área entre 60 e 100 metros quadrados : {}
Nº de imóveis que tenham pelo menos 4 quartos e aluguel menor que R$ 2.000: {}
'''.format(apartamentos.shape[0],casas.shape[0],areas.shape[0],quartos_aluguel.shape[0]))
# + [markdown] id="rYd2VWcJJYjh"
# # <font color = green> Aula 5
#
# + [markdown] id="W_9djIKsJYji"
# ## 5.1. Tratamento de dados faltantes
# + id="eBF1yBUCJYjj" outputId="223b8d96-2dd0-4bf1-9e39-038a3f60cd97"
dados.head(10)
# + id="Ed_tgB6SJYjl" outputId="1a44e76d-55bb-4e8a-823c-f17aba4ee8f7"
#Dados nulos
dados.isnull().head(10)
# + id="W81ve6jkJYjp" outputId="f6c55371-fc6d-4cb9-cc45-5397e2d548ed"
#Dados não nulos
dados.notnull().head(10)
# + id="yPDt1yJlJYju" outputId="52e5c91c-ca91-44b0-ab4a-bca143bfafcf"
dados.info()
# + id="bO15E4LTJYjy" outputId="f33afbaf-b71b-4bfc-ed7e-a510eaa8c6ae"
#Valore de alguel nulo
dados[dados.Valor.isnull()]
# + id="XpO10XjPJYj3" outputId="9442bf8a-2019-409f-c849-ea2f2063951a"
#Elimiar nulos
dados.dropna(subset=['Valor'],inplace=True)
dados[dados.Valor.isnull()]
# + [markdown] id="_z31BkojJYj6"
# ## 5.2. Tratamento condicional
# + id="6azeCbq3JYj9" outputId="267f1d74-98eb-43bd-f9f2-63e85058a7bf"
#Aparmentos sem condomínio
dados[dados.Condominio.isnull()].shape[0]
# + id="BIpnXrAiJYkC" outputId="4bb7a10d-4a03-4eb7-998c-9fdd9a11bc6d"
#Aparmentos sem condomínio
selecao=(dados.Tipo=='Apartamento') & (dados.Condominio.isnull())
selecao
# + id="LwS6kuYQJYkH" outputId="52753d3a-6454-40f8-c197-6c775bab05bd"
#Elimiar aparmentos sem condomínio
dados= dados[~selecao]
dados.shape[0]
# + id="RKRZA-oVJYkK" outputId="dfda3343-6941-4709-9676-913434996436"
dados.fillna(0,inplace=True)
dados.head(10)
# + id="-uKvREk6JYkN" outputId="f1708719-eab5-46b7-a4b9-113219aa19f1"
#Todos os nulos foram eliminados
dados.info()
# + id="lZamItm9JYkQ" outputId="5a4f55e3-e830-43c6-acd7-b9775bcc7412"
#Gravar dados tratados
dados.to_csv('dados/aluguel_residencial_tratado.csv', sep = ';',index=False)
dados_residencial= pd.read_csv('dados/aluguel_residencial_tratado.csv', sep = ';')
dados_residencial.head(10)
# + [markdown] id="YE2pH4mUJYkU"
# # <font color = green> Aula 6
# + [markdown] id="yCnuz7CdJYkV"
# ## 6.1. Criando a variável valor bruto
# + id="49ZP5up1JYkV" outputId="3cd1c682-a27f-413f-b4d1-b478060abed0"
dados=dados_residencial
dados['Valor Bruto']=dados.Valor+dados.Condominio+dados.IPTU
dados.head(10)
# + id="yRxd1zYRJYkY"
#Obs: Devido forma de nomear com epaço valor bruto só pode ser acessada entre chaves []
# + [markdown] id="CpkZfJkoJYka"
# ## 6.2. Criando a variável custo do m<sup>2<sup>
# + id="vnT96kwWJYkb" outputId="44d7bbb6-9e69-4f46-f75a-332b4a9873e5"
dados['Valor m²']=dados.Valor/dados.Area
dados.head(10)
# + id="lQSgLsLqJYkd" outputId="a7634f6b-ff3d-4414-d142-afc20ca1f0f2"
dados['Valor m²']=dados['Valor m²'].round(2)
dados.head(10)
# + [markdown] id="1nDyAKVSJYkg"
# ## 6.3. Criando a variável valor bruto do m<sup>2<sup>
# + id="CVDMHayFJYkg" outputId="5af8faa3-60ff-4665-b025-40ad04c883d2"
dados['Valor Bruto m²']=dados['Valor Bruto']/dados.Area
dados['Valor Bruto m²']=dados['Valor Bruto m²'].round(2)
dados.head(10)
# + [markdown] id="9VOII01aJYki"
# ## 6.4. Seprando casas dos demias imóveis
# + id="vIO0oe8vJYkj"
casas = ['Casa','Casa de Condomínio','Casa de Vila']
# + id="1lJrqLW9JYkk" outputId="3a67d250-bbb3-4c56-beb7-8e252b8a12a7"
dados['Tipo Agregado']=dados['Tipo'].apply(lambda x:'Casa' if x in casas else 'Apartamento')
dados.head(10)
# + id="IuDh7_LjJYkm" outputId="da25f609-2253-4ccd-84ad-67497d85bbe8"
#Orgizando colunas por ordem alfabética
dados.sort_index(inplace=True,axis = 1)
dados.head(10)
# + id="HtUcCSzPJYkr" outputId="829fbd65-275d-4b1b-daa8-06ffe8f36125"
#Orgnizando linhas pelo maior valor bruto
dados.sort_values(by = ['Valor Bruto'],ascending=False ,inplace=True)
dados.head(10)
# + [markdown] id="pYFmZbaQJYkt"
# ## 6.5. Excluindo variáveis
# + id="c3PFg3z4JYku" outputId="0e0fd1f1-1a27-42e2-f099-49730ae1e70f"
dados_aux = pd.DataFrame(dados[['Tipo Agregado', 'Valor m²', 'Valor Bruto', 'Valor Bruto m²']])
dados_aux.head(10)
# + id="wCE3OwyrJYky" outputId="53cfbd00-05ba-419d-b2c3-1b15bdfb9815"
#Método del
del dados_aux['Valor Bruto']
dados_aux.head(10)
# + id="wDXaXcIaJYk0" outputId="d3d91a61-dff5-458b-aadc-97161f2d17d6"
#Método pop
dados_aux.pop('Valor Bruto m²')
dados_aux.head(10)
# + id="Crutk2grJYk3"
#Metodo drop
dados.drop(['Valor Bruto','Valor Bruto m²'], axis=1,inplace=True)
# + id="gL9V1S9WJYk5" outputId="1f47a876-2a1e-4b77-f993-175a41ab9cc6"
dados.head(10)
# + id="-gBikxn0JYk8" outputId="8ab47d83-3eb8-410b-c43b-ca05fbd88197"
dados.to_csv('dados/aluguel_residencial.csv', sep=';',index=False)
dados=pd.read_csv('dados/aluguel_residencial.csv', sep=';')
dados.head(10)
# + [markdown] id="QtqgzqTkJYk-"
# # <font color = green> Aula 7
# + [markdown] id="6JwDJj9iJYk_"
# ## 7.1. Estatística descritiva
# + id="JKpPujvvJYlA" outputId="4589b1ad-a5d2-482c-c489-1a4a7ae07139"
dados.Bairro.value_counts()
# + id="zdW32UPyJYlC" outputId="221bccc4-e678-40f9-a91c-b8ab170e1f97"
dados.Tipo.value_counts()
# + id="FsJqfDwtJYlE" outputId="5eddec1d-2a16-436b-e257-acbaabc1ee34"
dados.Valor.mean()
# + id="uyM7i2LTJYlG" outputId="96bc791f-b85b-4960-f10d-60dde98511a8"
descritiva=dados.describe()
descritiva.columns.name='Estatística'
descritiva
# + [markdown] id="Kyd3IY_5JYlH"
# ### 7.1.1. Trabalhando com bairros específicos
# + id="7xmBF1D8JYlI" outputId="4f63155f-c65f-4483-d819-315d7e105239"
bairros = ['Barra da Tijuca', 'Copacabana', 'Ipanema', 'Leblon', 'Botafogo', 'Flamengo', 'Tijuca']
selecao = dados['Bairro'].isin(bairros)
dados_2 = dados[selecao]
dados_2.head(10)
# + id="Ltk_vpceJYlJ" outputId="7bd31961-1a39-4bc0-f879-1298388ed7aa"
dados_2['Bairro'].drop_duplicates()
# + id="l8mf4LMpJYlL" outputId="07224bd9-10c8-4e86-8c6f-d1368b0431cf"
bairros=dados_2.groupby('Bairro')
type(bairros)
# + id="Le_JSMVOJYlP" outputId="3624f823-9303-445c-b995-31de35223f7d"
bairros.groups
# + id="ef7ZHAz4JYlS" outputId="53b914ee-bafa-4b06-a614-2fc60e38e31f"
for bairro,data in bairros:
print('{} -> {}'.format(bairro,data.Valor.mean()))
# + id="pKRMpJu1JYlU" outputId="4310285d-adcc-4dc9-f067-ebbc4d3e452f"
bairros[['Valor','Condominio']].mean().round(2)
# + id="qdoJOq2_JYlW" outputId="714c6b57-b855-4f5f-b0b6-c113bba49a5e"
bairros.Valor.describe()
# + id="YGdCGJMDJYlY" outputId="79c344e7-5091-4b88-9e15-731baa5b2c2d"
Ex=bairros.Valor.aggregate(['min','max','std'])
Ex.columns=['Mínimo','Máximo','Desv.Pad']
Ex
# + [markdown] id="1wdiXep5JYla"
# ### 7.1.1. Trabalhando com todos bairros
# + id="LD8HH-rnJYlb" outputId="d4c09931-744b-4597-99ea-da575976f480"
dados.to_csv('dados/aluguel_residencial.csv', sep=';',index=False)
dados=pd.read_csv('dados/aluguel_residencial.csv', sep=';')
dados.tail(10)
# + id="zf7GLEuUJYlc" outputId="9e462b51-a8c5-4871-e021-7638800b1715"
todos_bairros=dados.groupby('Bairro')
Ex=todos_bairros.Valor.aggregate(['min','max','std'])
Ex.columns=['Mínimo','Máximo','Desv.Pad']
Ex
# + [markdown] id="U29F7HteJYlf"
# ## 7.2. Visulização
# + id="cY6R-bYkJYlg"
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rc('figure',figsize=(20,10))
# + id="vNUfRWt-JYli" outputId="80b8a329-6ebf-4cde-b314-94168547b54a"
fig1=todos_bairros.Valor.std().plot.bar(color='green')
fig1.set_ylabel('Valor do Aluguel')
fig1.set_title('Desvio Padrão do Aluguel',{'fontsize':20})
# + id="KhmXS80mJYlj" outputId="a9a38a58-44b1-4782-e22a-94b70fdfd50c"
fig2=bairros.Valor.std().plot.bar(color='green')
fig2.set_ylabel('Valor do Aluguel')
fig2.set_title('Desvio Padrão do Aluguel',{'fontsize':15})
# + [markdown] id="INJMky0rJYlk"
# # <font color = green> Aula 8
# + [markdown] id="U6bTZD_ZJYll"
# ## 8.1. Identificando e Removendo Outliers
# + id="_O8MOvYlJYll"
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.rc('figure', figsize = (14,6))
# + id="tKV8B7dbJYlm" outputId="4a983b49-eb68-435e-b828-a7ab70422e37"
#Pelo matplotlib
dados.boxplot(['Valor'])
# + id="g9tOKX_zJYlo" outputId="7d40563a-15e5-4323-ac96-217f1bf0c746"
dados[dados.Valor>=500000]
# + id="8mCM9OZSJYlr"
valor=dados.Valor
# + id="5vhOY-p2JYlu"
#Quantis
Q1=valor.quantile(.25)
Q3=valor.quantile(.75)
IIQ=Q3-Q1
LB=Q1-1.5*IIQ
UB=Q3+1.5*IIQ
# + id="zIqntx09JYlw" outputId="6fbfe468-244a-4efb-c2d6-a329df090159"
selecao=(valor>=LB) & (valor<=UB)
dados_novos=dados[selecao]
dados_novos.head(10)
# + id="RRAs77F_JYly" outputId="23caa761-99e4-4945-c03e-f5c2251e18ee"
dados_novos.boxplot(['Valor'])
# + id="5wRZqjEyJYl0" outputId="7e70d8d4-3e32-4d8c-873a-2fec08da8856"
#Pelo SeaBorn
sns.boxplot(x='Valor',data=dados)
# + id="e6h9mXwFJYl2" outputId="fed19e5a-4426-4362-c33e-5d5398d2c445"
sns.boxplot(x='Valor',data=dados_novos)
# + [markdown] id="XbGbblO2JYl4"
# ### 8.1.1. Histogramas
# + id="VxebnyWoJYl4" outputId="ac7e733f-a719-453d-a9a6-eb8e04f7aaba"
dados.hist(['Valor'])
dados_novos.hist(['Valor'])
# + id="9o2_pF-_JYl6" outputId="5b8c4fd2-bf12-47bc-82d5-31c1cef5e1bd"
sns.distplot(dados.Valor)
# + id="HoEuTq7oJYl7" outputId="675ce7d1-c497-4568-ca6b-5908c0e8abaa"
sns.distplot(dados_novos.Valor)
# + [markdown] id="N9AptofxJYl9"
# ## 8.2. Identificando e Removendo Outliers por grupo
# + id="9Eecwuz8JYl9" outputId="66af7844-418f-4ab9-83b0-1a7d05a0b8aa"
dados.boxplot(['Valor'],by=['Tipo'])
# + id="aq503DIcJYl-" outputId="bfb7d253-2ac5-4e9a-a401-05e4b8ae4e90"
sns.boxplot(x='Tipo',y='Valor',data=dados)
# + id="HtFuUFszJYmA" outputId="4cbb741c-5212-4a30-e576-e06a410615fd"
grupo_tipo=dados.groupby('Tipo')['Valor']
type(grupo_tipo)
# + id="w2iYL-ToJYmC"
#Quantis
Q1=grupo_tipo.quantile(.25)
Q3=grupo_tipo.quantile(.75)
IIQ=Q3-Q1
LB=Q1-1.5*IIQ
UB=Q3+1.5*IIQ
# + id="0uVeoEQJJYmF" outputId="abae047f-05df-42d4-afdc-5451f71398e4"
Q1
# + id="iGxxFMHSJYmH" outputId="f789343f-94bd-42bb-8191-a16fbf52e7cd"
Q1['Casa']
# + id="7bJpkanNJYmJ"
dados_novos=pd.DataFrame()
for tipo in grupo_tipo.groups.keys():
eh_tipo=dados.Tipo==tipo
eh_dentro_limite= (dados.Valor>=LB[tipo]) & (dados.Valor<=UB[tipo])
selecao=eh_tipo & eh_dentro_limite
dados_selecao=dados[selecao]
dados_novos=pd.concat([dados_novos,dados_selecao])
# + id="xOyXH5z4JYmN" outputId="e3c467ce-8b92-44d6-84bd-282c9edc6b38"
dados_novos.boxplot(['Valor'],by=['Tipo'])
# + id="konfihvgJYmO" outputId="a628374a-29d9-4d35-ccb2-9e944b486b28"
sns.boxplot(x='Tipo',y='Valor',data=dados_novos)
# + id="SljxiavsJYmP" outputId="e70e1d40-a5fc-403b-f959-9adda57f91bc"
dados_novos.to_csv('dados/aluguel_residencial_tratado.csv',sep=';',index=False)
residencial_tratado=pd.read_csv('dados/aluguel_residencial_tratado.csv',sep=';')
residencial_tratado.head(10)
# + id="Wl7oUgU6JYmR"
| Python/Data Science With Python/Pandas 101.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install nltk
import numpy as np
import pandas as pd
import re
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report, r2_score, mean_squared_error
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import cross_val_score
import nltk
nltk.download('stopwords')
# # Wine Points Prediction
# **Note:** a sample is being used due to the kernel running out of memory due to the jupyter notebook kernel running out of memory and dying when performing analysis otherwise as a result predictions depend on the sample that is given as it is obtained randomly. Furthermore, these predictions vary with $\pm1\%$
# +
wine_df = pd.read_csv("wine.csv")
str_cols = ['description', 'price', 'title', 'variety', 'country', 'designation', 'province', 'winery']
reviews = wine_df.sample(20000)[['points'] + str_cols].reset_index()
reviews = reviews.drop(['index'], axis=1)
reviews.head()
# -
# We first have to change features that we are going to use from categorical to numerical variables. This is done to give the features meaning when performing different forms of analysis on them to predict the points given to a bottle of wine.
# +
# assign numerical values to string columns
factorized_wine = reviews[str_cols].drop(['description'], axis=1).copy()
for col in str_cols[2:]:
factorized_wine[col] = pd.factorize(reviews[col])[0]
factorized_wine.head()
# -
# Now we assign the variables we just factorized along with the price of the wine to be our X values and our y value will be what we are trying to predict, which in this case are the points for a bottle of wine.
X = factorized_wine.to_numpy('int64')
y = reviews['points'].values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
# Below we perform several different forms of prediction to see which one produces the best result.
#
# We then need to determine how accurate this algorithm is given the estimates returned from the random forest regression. We do this by using `score()` which returns the coefficient of determination of the prediction ($r^2$). In other words, it is the observed y variation that can be explained by the and by the regression model. We also compute the residual mean squared error of the model (rmse).
# ### linear regression
# +
from sklearn import linear_model
model = linear_model.LinearRegression()
model.fit(X_train, y_train)
pred = model.predict(X_test)
print('r2 score:', model.score(X_test,y_test))
print('rmse score:', mean_squared_error(y_test, pred, squared=False))
# -
# as you can see this isnt the best prediction model so lets try some other methods and see what we get
# ### linear discriminant analysis
# +
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
lda_model = LinearDiscriminantAnalysis()
lda_model.fit(X_train, y_train)
pred = lda_model.predict(X_test)
print('r2 score:', lda_model.score(X_test,y_test))
print('rmse score:', mean_squared_error(y_test, pred, squared=False))
# -
# The results from this method are not good either so onto the next one
# ### classification tree
# +
from sklearn import tree
dt_model = tree.DecisionTreeClassifier()
dt_model.fit(X_train, y_train)
pred = dt_model.predict(X_test)
print('r2 score:', dt_model.score(X_test,y_test))
print('rmse score:', mean_squared_error(y_test, pred, squared=False))
# -
# The methods that we have done prior as well as this one are getting used nowhere and are showing very little signs of improvement so let's pivot to a different direction and try to predict the points based on the description of the wine.
# ## Incorporating description
reviews.head()
# Because we are focusing on the description (review) of the wine here is an example of one
reviews['description'][5]
# We remove punctuation and other special characters and convert everything to lower case as it is not significat that words be capitalized.
# +
descriptions = []
for descrip in reviews['description']:
line = re.sub(r'\W', ' ', str(descrip))
line = line.lower()
descriptions.append(line)
len(descriptions)
# -
# Here we use `TfidfVectorizer`, to understand what it is what term frequency-inverse document frequency (TF_IDT) is must be explained first. TF-IDF is a measure that evaluates the relevancy that a word has for a document inside a collection of other documents. Furthermore, TF-IDF can be defined as the following:
#
# $ \text{Term Frequency (TF)} = \frac{\text{Frequency of a word}}{\text{Total number of words in document}} $
#
# $ \text{Inverse Document Frequency (IDF)} = \log{\frac{\text{Total number of documents}}{\text{Number of documents that contain the word}}} $
#
# $ \text{TF-IDF} = \text{TF} \cdot \text{IDF} $
#
# In turn, what `TfidfVectorizer` gives us is a list of feature lists that we can use as estimators for prediction.
#
# The parameters for `TfidfVectorizer` are max_features, max_df, and stop_words.
# max_features tells us to only look at the top n features of the total document
# max_df causes the vectorizer to ignore terms that have a document frequency strictly higher than the given threshold. In our case because a float is its value we ignore words that appear in more than 80% of documents
# stop_words allows us to pass in a set of stop words. Stop words are words that add little to no meaning to a sentence. This includes words such as I, our, him, and her.
# Following this we fit and transform the data then we split it into training and testing data
# +
y = reviews['points'].values
vec = TfidfVectorizer(max_features=2500, min_df=7, max_df=0.8, stop_words=stopwords.words('english'))
X = vec.fit_transform(descriptions).toarray()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 12)
# -
# Now that we've split the data we use `RandomForestRegressor()` to make our prediciton given that its a random forest algorithm it takes the average of the decision trees that were created and used as estimates.
rfr = RandomForestRegressor()
rfr.fit(X_train, y_train)
pred = rfr.predict(X_test)
# Now we check to see how good our model is at predicting the points for a bottle of wine
print('r2 score:', rfr.score(X_test, y_test))
print('rmse score:', mean_squared_error(y_test, pred, squared=False))
cvs = cross_val_score(rfr, X_test, y_test, cv=10)
cvs.mean()
# This is solely based on the description of the wine. As you can see this is a large improvement in both the score and rmse for any sort of prediction that was done with any of the methods performed above. However, it is still not the best for several reasons. The first being the $r^2$ score, or how well our model is at making predictions. There is still a large portion of the data that is not being accurately predicted.
#
# The other issue pertains to when the model does fail at making the prediction. given that the rmse score is very high this can be interpreted as when we do fail we fail rather spectacularly. However, given that the context of this problem is making a prediction for determining arbitrary integer point values for bottles of wine, failing spectacularly is not necessarily what is occurring. The rmse value tells us that with each incorrect prediction we are about 2.1 points off. However, it is still less than ideal.
#
# Below we see if we can improve upon these shortcomings.
# ## Combining features
# Next we combine the features that were obtained from `TfidfVectorizer` with the features that we just factorized in there respective rows.
wine_X = factorized_wine.to_numpy('int64')
X = np.concatenate((wine_X,X),axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 12)
rfr_fac = RandomForestRegressor()
rfr_fac.fit(X_train, y_train)
fac_pred = rfr_fac.predict(X_test)
# Next we perform the same actions as above to determine the accuracy of the prediction. That is we use `score()` and perform a 10 fold cross validation and then take the mean of the scores.
print('r2 score:', rfr_fac.score(X_test, y_test))
print('rmse score:', mean_squared_error(y_test, fac_pred, squared=False))
fac_cvs = cross_val_score(rfr_fac, X_test, y_test, cv=10)
fac_cvs.mean()
# As we can see from the scores computed above the accuracy is an improvement from only using the wine description (review) as a feature. Both the $r^2$ score and RMSE value improved by about 8% and 0.15 respectively. However, the model isn't all that reliable as there is only slightly above 50% of the bottles of wine from the sample can have their score predicted accurately
# # Conclusion
# After comparing the price to the points for a bottle of wine we learned that the majority of the data is clustered towards the middle in regards to the point value a bottle was awarded and there are few outliers in either the positive or negative direction. Furthermore, most wine follows the trend of having a greater number of points awarded as the price increases.
#
# From these trends, we attempted to determine if we can actually predict how many points a bottle of wine will receive. Given the best prediction that we could obtain took into account several features, including the price of the wine, and only has 52% accuracy, we are lead to believe that the point system that results from the wine in this dataset is more subjective than objective.
| main/TFIDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#But-de-ce-notebook" data-toc-modified-id="But-de-ce-notebook-1"><span class="toc-item-num">1 </span>But de ce notebook</a></div><div class="lev1 toc-item"><a href="#Règles-du-Jap-Jap" data-toc-modified-id="Règles-du-Jap-Jap-2"><span class="toc-item-num">2 </span>Règles du <em>Jap Jap</em></a></div><div class="lev2 toc-item"><a href="#But-du-jeu" data-toc-modified-id="But-du-jeu-21"><span class="toc-item-num">2.1 </span>But du jeu</a></div><div class="lev2 toc-item"><a href="#Début-du-jeu" data-toc-modified-id="Début-du-jeu-22"><span class="toc-item-num">2.2 </span>Début du jeu</a></div><div class="lev2 toc-item"><a href="#Tour-de-jeu" data-toc-modified-id="Tour-de-jeu-23"><span class="toc-item-num">2.3 </span>Tour de jeu</a></div><div class="lev2 toc-item"><a href="#Fin-du-jeu" data-toc-modified-id="Fin-du-jeu-24"><span class="toc-item-num">2.4 </span>Fin du jeu</a></div><div class="lev1 toc-item"><a href="#Code-du-jeu" data-toc-modified-id="Code-du-jeu-3"><span class="toc-item-num">3 </span>Code du jeu</a></div><div class="lev2 toc-item"><a href="#Code-pour-représenter-une-carte-à-jouer" data-toc-modified-id="Code-pour-représenter-une-carte-à-jouer-31"><span class="toc-item-num">3.1 </span>Code pour représenter une carte à jouer</a></div><div class="lev2 toc-item"><a href="#Fin-du-jeu" data-toc-modified-id="Fin-du-jeu-32"><span class="toc-item-num">3.2 </span>Fin du jeu</a></div><div class="lev2 toc-item"><a href="#Actions" data-toc-modified-id="Actions-33"><span class="toc-item-num">3.3 </span>Actions</a></div><div class="lev2 toc-item"><a href="#Valider-un-coup" data-toc-modified-id="Valider-un-coup-34"><span class="toc-item-num">3.4 </span>Valider un coup</a></div><div class="lev2 toc-item"><a href="#Jeu-interactif" data-toc-modified-id="Jeu-interactif-35"><span class="toc-item-num">3.5 </span>Jeu interactif</a></div><div class="lev2 toc-item"><a href="#Etat-du-jeu" data-toc-modified-id="Etat-du-jeu-36"><span class="toc-item-num">3.6 </span>Etat du jeu</a></div><div class="lev2 toc-item"><a href="#Lancement-du-jeu-intéractif" data-toc-modified-id="Lancement-du-jeu-intéractif-37"><span class="toc-item-num">3.7 </span>Lancement du jeu intéractif</a></div><div class="lev1 toc-item"><a href="#Conclusion" data-toc-modified-id="Conclusion-4"><span class="toc-item-num">4 </span>Conclusion</a></div>
# -
# ----
# # But de ce notebook
#
# - Je vais expliquer les règles d'un jeu de carte, le "Jap Jap", qu'on m'a appris pendant l'été,
# - Je veux simuler ce jeu, en Python, afin de calculer quelques statistiques sur le jeu,
# - J'aimerai essayer d'écrire une petite intelligence artificielle permettant de jouer contre l'ordinateur,
# - Le but est de faire un prototype d'une application web ou mobile qui permettrait de jouer contre son téléphone !
# ----
# # Règles du *Jap Jap*
#
# ## But du jeu
# - Le *Jap Jap* se joue à $n \geq 2$ joueur-euse-s (désignées par le mot neutre "personne"), avec un jeu de $52$ cartes classiques (4 couleurs, 1 à 10 + vallet/dame/roi).
# - Chaque partie du *Jap Jap* jeu se joue en plusieurs manches. A la fin de chaque manche, une personne gagne et les autres marquent des points. Le but est d'avoir le moins de point possible, et la première personne a atteindre $90$ points a perdu !
# - On peut rendre le jeu plus long en comptant la première personne à perdre $x \geq 1$ parties.
#
# ## Début du jeu
# - Chaque personne reçoit 5 cartes,
# - et on révèle la première carte de la pioche.
#
# ## Tour de jeu
# - Chaque personne joue l'une après l'autre, dans le sens horaire (anti trigonométrique),
# - A son tour, la personne a le choix entre jouer normalement, ou déclencher la fin de jeu si elle possède une main valant $v \leq 5$ points (voir "Fin du jeu" plus bas),
# - Jouer normalement consiste à jeter *une ou plusieurs* ($x \in \{1,\dots,5\}$) cartes de sa main dans la défausse, et prendre *une* carte et la remettre dans sa main. Elle peut choisir la carte du sommet de la pioche (qui est face cachée), ou *une* des $x' \in \{1,\dots,5\}$ cartes ayant été jetées par la personne précédente, ou bien la première carte de la défausse si c'est le début de la partie.
#
# ## Fin du jeu
# - Dès qu'une personne possède une main valant $v \leq 5$ points, elle peut dire *Jap Jap !* au lieu de jouer à son tour.
# + Si elle est la seule personne à avoir une telle main de moins de $5$ points, elle gagne !
# + Si une autre personne a une main de moins de $5$ points, elle peut dire *Contre Jap Jap !*, à condition d'avoir *strictement* moins de points que le *Jap Jap !* ou le *Contre Jap Jap !* précédent. La personne qui remporte la manche est celle qui a eu le *Contre Jap Jap !* de plus petite valeur.
# - La personne qui a gagné ne marque aucun point, et les autres ajoutent à leur total actuel de point
# - Si quelqu'un atteint $90$ points, elle perd la partie.
# ----
# # Code du jeu
# ## Code pour représenter une carte à jouer
coeur = "♥"
treffle = "♣"
pique = "♠"
carreau = "♦"
couleurs = [coeur, treffle, pique, carreau]
class Carte():
def __init__(self, valeur, couleur):
assert 1 <= valeur <= 13, "Erreur : valeur doit etre entre 1 et 13."
self.valeur = valeur
assert couleur in couleurs, "Erreur : couleur doit etre dans la liste {}.".format(couleurs)
self.couleur = couleur
def __str__(self):
val = str(self.valeur)
if self.valeur > 10:
val = {11: "V" , 12: "Q" , 13: "K"}[self.valeur]
return "{:>2}{}".format(val, self.couleur)
__repr__ = __str__
def val(self):
return self.valeur
def valeur_main(liste_carte):
return sum(carte.val() for carte in liste_carte)
# +
import random
def nouveau_jeu():
jeu = [
Carte(valeur, couleur)
for valeur in range(1, 13+1)
for couleur in couleurs
]
random.shuffle(jeu)
return jeu
# -
nouveau_jeu()[:5]
valeur_main(_)
# ## Fin du jeu
# Pour représenter la fin du jeu :
class FinDuneManche(Exception):
pass
class FinDunePartie(Exception):
pass
# ## Actions
# Pour représenter une action choisie par une personne :
# +
class action():
def __init__(self, typeAction="piocher", choix=None):
assert typeAction in ["piocher", "choisir", "Jap Jap !"]
self.typeAction = typeAction
assert choix is None or choix in [0, 1, 2, 3, 4]
self.choix = choix
def __str__(self):
if self.est_piocher(): return "Piocher"
elif self.est_japjap(): return "Jap Jap !"
elif self.est_choisir(): return "Choisir #{}".format(self.choix)
def est_piocher(self):
return self.typeAction == "piocher"
def est_choisir(self):
return self.typeAction == "choisir"
def est_japjap(self):
return self.typeAction == "Jap Jap !"
action_piocher = action("piocher")
action_japjap = action("Jap Jap !")
action_choisir0 = action("choisir", 0)
action_choisir1 = action("choisir", 1)
action_choisir2 = action("choisir", 2)
action_choisir3 = action("choisir", 3)
action_choisir4 = action("choisir", 4)
# -
# ## Valider un coup
# Pour savoir si une suite de valeurs est bien continue :
def suite_valeurs_est_continue(valeurs):
vs = sorted(valeurs)
differences = [ vs[i + 1] - vs[i] for i in range(len(vs) - 1) ]
return all([d == 1 for d in differences])
suite_valeurs_est_continue([5, 6, 7])
suite_valeurs_est_continue([5, 7, 8])
# Pour valider un coup choisie par une personne :
def valide_le_coup(jetees):
if jetees is None or not (1 <= len(jetees) <= 5):
return False
# coup valide si une seule carte !
elif len(jetees) == 1:
return True
# si plus d'une carte
elif len(jetees) >= 2:
couleurs_jetees = [carte.couleur for carte in jetees]
valeurs_jetees = sorted([carte.valeur for carte in jetees])
# coup valide si une seule couleur et une suite de valeurs croissantes et continues
if len(set(couleurs_jetees)) == 1:
return suite_valeurs_est_continue(valeurs_jetees)
# coup valide si une seule valeur et différentes couleurs
elif len(set(valeurs_jetees)) == 1:
return len(set(couleurs_jetees)) == len(couleurs_jetees)
return False
# Exemples de coups valides :
valide_le_coup([Carte(4, coeur)])
valide_le_coup([Carte(4, coeur), Carte(5, coeur)])
valide_le_coup([Carte(4, coeur), Carte(5, coeur), Carte(3, coeur)])
valide_le_coup([Carte(4, coeur), Carte(5, coeur), Carte(3, coeur), Carte(2, coeur), Carte(6, coeur)])
valide_le_coup([Carte(4, coeur), Carte(4, carreau)])
valide_le_coup([Carte(4, coeur), Carte(4, carreau), Carte(4, pique)])
valide_le_coup([Carte(4, coeur), Carte(4, carreau), Carte(4, pique), Carte(4, treffle)])
# Exemples de coups pas valides :
valide_le_coup([Carte(4, coeur), Carte(9, coeur)])
valide_le_coup([Carte(4, coeur), Carte(4, coeur), Carte(3, coeur)])
valide_le_coup([Carte(4, coeur), Carte(12, carreau)])
valide_le_coup([Carte(4, coeur), Carte(4, carreau), Carte(4, pique)])
valide_le_coup([Carte(4, coeur), Carte(4, carreau), Carte(4, pique), Carte(4, treffle)])
# ## Jeu interactif
# On va utiliser les widgets ipython pour construire le jeu interactif !
# Voir https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20Asynchronous.html#Waiting-for-user-interaction
# %gui asyncio
# +
import asyncio
def wait_for_change(widget, value):
future = asyncio.Future()
def getvalue(change):
# make the new value available
future.set_result(change.new)
widget.unobserve(getvalue, value)
widget.observe(getvalue, value)
return future
# +
import ipywidgets as widgets
from IPython.display import display
style = {
'description_width': 'initial',
}
style2boutons = {
'description_width': 'initial',
'button_width': '50vw',
}
style3boutons = {
'description_width': 'initial',
'button_width': '33vw',
}
style4boutons = {
'description_width': 'initial',
'button_width': '25vw',
}
style5boutons = {
'description_width': 'initial',
'button_width': '20vw',
}
# -
# Pour savoir quoi jouer :
def piocher_ou_choisir_une_carte_visible():
return widgets.ToggleButtons(
options=["Une carte dans la pioche ", "Une carte du sommet de la défausse "],
index=0,
tooltips=["invisible", "visibles"],
icons=["question", "list-ol"],
description="Action ?",
style=style4boutons,
)
bouton = piocher_ou_choisir_une_carte_visible()
display(bouton)
print("Choix :", bouton.index)
# Pour savoir quoi jeter :
exemple_de_main = [Carte(10, coeur), Carte(11, coeur), Carte(11, pique)]
exemple_de_main
def faire_japjap(main):
return widgets.ToggleButton(
value=False,
description="Jap Jap ? ({})".format(valeur_main(main)),
button_style="success",
tooltip="Votre main vaut moins de 5 points, donc vous pouvez terminer la partie !",
icon="check",
style=style,
)
b = faire_japjap(exemple_de_main)
display(b)
print("Choix :", b.value)
def quoi_jeter(main):
return widgets.SelectMultiple(
options=main,
index=[0],
description="Quoi jeter ?",
style=style,
)
b = quoi_jeter(exemple_de_main)
display(b)
print("Choix :", b.index)
from IPython.display import display
def valider_action():
return widgets.ToggleButton(description="Valider l'action ?")
# Pour savoir quoi piocher :
exemple_de_visibles = [Carte(11, pique), Carte(10, treffle)]
exemple_de_visibles
def quoi_prendre(visibles):
return widgets.ToggleButtons(
options=visibles,
#index=0,
description="Prendre quelle carte du sommet ?",
style=style,
)
quoi_prendre(exemple_de_visibles)
# On va tricher et afficher les cartes avec `display(Markdown(...))` plutôt que `print`, pour les avoir en couleurs.
from IPython.display import Markdown
print("[Alice] Cartes en main : [ 6♦, 5♠, V♣, V♠, 1♣]")
# avec de la couleurs
display(Markdown("[Alice] Cartes en main : [ 6♦, 5♠, V♣, V♠, 1♣]"))
# Maintenant on peut tout combiner.
async def demander_action(visibles=None, main=None, stockResultat=None):
display(Markdown("- Main : {} (valeur = {})".format(main, valeur_main(main))))
display(Markdown("- Sommet de la défausse {}".format(visibles)))
fait_japjap = False
# 1.a. si on peut faire jap jap, demander si on le fait ?
if valeur_main(main) <= 5:
print("Vous pouvez faire Jap Jap !")
bouton3 = faire_japjap(main)
validation = valider_action()
display(widgets.VBox([bouton3, validation]))
await wait_for_change(validation, 'value')
# print(" ==> Choix :", bouton3.value)
if bouton3.value:
fait_japjap = True
typeAction = "Jap Jap !"
jetees = None
# 1.b. quoi jouer
if not fait_japjap:
bouton1 = piocher_ou_choisir_une_carte_visible()
validation = valider_action()
display(widgets.VBox([bouton1, validation]))
await wait_for_change(validation, 'value')
piocher = bouton1.index == 0
# print(" ==> Choix :", bouton1.value)
# 2.a. si piocher, rien à faire pour savoir quoi piocher
if piocher:
print("Okay, vous piochez.")
typeAction = "piocher"
choix = None
# 2.b. si choisir carte
else:
typeAction = "choisir"
print("Okay, vous choisissez dans le sommet de la défausse.")
if len(visibles) > 1:
bouton2 = quoi_prendre(visibles)
validation = valider_action()
display(widgets.VBox([bouton2, validation]))
await wait_for_change(validation, 'value')
# print(" ==> Choix :", bouton2.index)
choix = bouton2.index
else:
choix = 0
# 3. choisir quoi jeter
if typeAction != "Jap Jap !":
if len(main) > 1:
pas_encore_de_coup = True
jetees = None
while pas_encore_de_coup or not valide_le_coup(jetees):
bouton4 = quoi_jeter(main)
validation = valider_action()
display(widgets.VBox([bouton4, validation]))
await wait_for_change(validation, 'value')
# print(" ==> Choix :", bouton4.index)
jetees = bouton4.index
pas_encore_de_coup = False
if not valide_le_coup(jetees):
print("ERREUR ce coup n'est pas valide, on ne peut pas se débarasser de cet ensemble de cartes {}.".format(jetees))
else:
jetees = 0
action_choisie = action(typeAction=typeAction, choix=choix)
if stockResultat is not None:
stockResultat["action_choisie"] = action_choisie
stockResultat["jetees"] = jetees
return action_choisie, jetees
# + code_folding=[0]
if False:
stockResultat = {"action_choisie": None, "jetees": None}
asyncio.ensure_future(
demander_action(
visibles=exemple_de_visibles,
main=exemple_de_main,
stockResultat=stockResultat,
)
)
stockResultat
# + code_folding=[0]
def demander_action_et_donne_resultat(visibles=None, main=None):
stockResultat = {"action_choisie": None, "jetees": None}
asyncio.ensure_future(
demander_action(
visibles=visibles,
main=main,
stockResultat=stockResultat,
)
)
return stockResultat["action_choisie"], stockResultat["jetees"]
# -
# ## Etat du jeu
# Maintenant on peut représenter un état du jeu.
# on peut changer ici pour jouer moins longtemps !
scoreMax = 90
scoreMax = 10
# + code_folding=[1, 20, 31, 38, 44, 49, 53, 65]
class EtatJeu():
def __init__(self, nbPersonnes=2, nomsPersonnes=None,
scoreMax=scoreMax, malusContreJapJap=25, nbCartesMax=5):
assert 2 <= nbPersonnes <= 5, "Le nombre de personnes pouvant jouer doit etre entre 2 et 5."
self.nbPersonnes = nbPersonnes
self.nomsPersonnes = nomsPersonnes
self.scoreMax = scoreMax
self.malusContreJapJap = malusContreJapJap
self.nbCartesMax = nbCartesMax
# on initialise le stockage interne
self.personnes = [personne for personne in range(nbPersonnes)]
self.scores = [
0 for personne in self.personnes
]
self.mains = [
[ ] for personne in self.personnes
]
self.visibles = []
self.jeu = nouveau_jeu()
def montrer_information_visibles(self):
print("- Nombre de carte dans la pioche :", len(self.jeu))
print("- Cartes visibles au sommet de la défausse :", len(self.visibles))
for personne in self.personnes:
nom = self.nomsPersonnes[personne] if self.nomsPersonnes is not None else personne
main = self.mains[personne]
score = self.scores[personne]
print(" + Personne {} a {} carte{} en main, et un score de {}.".format(
nom, len(main), "s" if len(main) > 1 else "", score)
)
def montrer_information_privee(self, personne=0):
main = self.mains[personne]
nom = self.nomsPersonnes[personne] if self.nomsPersonnes is not None else personne
display(Markdown("[{}] Carte{} en main : {}".format(nom, "s" if len(main) > 1 else "", main)))
# --- Mécanique de pioche et distribution initiale
def prendre_une_carte_pioche(self):
if len(self.jeu) <= 0:
raise FinDuneManche
premiere_carte = self.jeu.pop(0)
return premiere_carte
def debut_jeu(self):
self.distribuer_mains()
premiere_carte = self.prendre_une_carte_pioche()
self.visibles = [premiere_carte]
def donner_une_carte(self, personne=0):
premiere_carte = self.prendre_une_carte_pioche()
self.mains[personne].append(premiere_carte)
def distribuer_mains(self):
self.mains = [
[ ] for personne in self.personnes
]
premiere_personne = random.choice(self.personnes)
self.personnes = self.personnes[premiere_personne:] + self.personnes[:premiere_personne]
for nb_carte in range(self.nbCartesMax):
for personne in self.personnes:
self.donner_une_carte(personne)
# --- Fin d'une manche
def fin_dune_manche(self):
self.jeu = nouveau_jeu()
self.debut_jeu()
# --- Enchainer les tours de jeux
async def enchainer_les_tours(self):
try:
indice_actuel = 0
while len(self.jeu) > 0:
# dans la même manche, on joue chaque tour, pour la personne actuelle
personne_actuelle = self.personnes[indice_actuel]
# 1. on affiche ce qui est public, et privé
self.montrer_information_visibles()
self.montrer_information_privee(personne_actuelle)
# 2. on demande l'action choisie par la personne
# action_choisie, jetees = demander_action_et_donne_resultat(
action_choisie, jetees = await demander_action(
visibles = self.visibles,
main = self.mains[personne_actuelle],
)
# 3. on joue l'action
self.jouer(
personne = personne_actuelle,
action = action_choisie,
indices = jetees,
)
# personne suivante
indice_actuel = (indice_actuel + 1) % self.nbPersonnes
if len(self.jeu) <= 0:
print("\nIl n'y a plus de cartes dans la pioche, fin de la manche sans personne qui gagne.")
raise FinDuneManche
except FinDuneManche:
print("\nFin d'une manche.")
fin_dune_manche()
except FinDunePartie:
print("\n\nFin d'une partie.")
# --- Un tour de jeu
def jouer(self, personne=0, action=action_piocher, indices=None):
print(" ? Personne {} joue l'action {} avec les indices {} ...".format(personne, action, indices)) # DEBUG
if indices is not None:
jetees = [ self.mains[personne][indice] for indice in indices ]
assert valide_le_coup(jetees)
# et on en prend une nouvelle
if action.est_piocher():
# soit celle face cachée en sommet de pioche
premiere_carte = self.prendre_une_carte_pioche()
display(Markdown("=> Vous piochez la carte {}.".format(premiere_carte)))
self.mains[personne].append(premiere_carte)
if action.est_choisir():
# soit une des cartes précedemment visibles
choix = action.choix
carte_choisie = self.visibles.pop(choix)
display(Markdown("=> Vous récupérez la carte {}.".format(carte_choisie)))
self.mains[personne].append(carte_choisie)
if action.est_japjap():
# on vérifie que cette personne a bien un Jap Jap !
valeur_du_premier_japjap = valeur_main(self.mains[personne])
assert 1 <= valeur_du_premier_japjap <= 5
gagnante = personne
display(Markdown("=> Vous faites un Jap Jap, valant {} point{}.".format(valeur_du_premier_japjap, "s" if valeur_du_premier_japjap > 1 else "")))
contre_JapJap = False
# on vérifie les valeurs des mains des autres personnes
valeurs_mains = [valeur_main(main) for main in self.mains]
plus_petite_valeur = min([valeurs_mains[autre_personne] for autre_personne in [ p for p in personnes if p != gagnante ]])
if plus_petite_valeur < valeur_du_premier_japjap:
print("CONTRE JAP JAP !")
# si une personne a un jap jap plus petit, la personne ne gagne pas
contre_JapJap = True
# la personne gagnante est la première (ordre du jeu) à obtenir le jap jap
# de valeur minimale, et en cas d'égalité c'est la personne obtenant
# cette valeur en le nombre minimal de cartes !
gagnantes = [ p for p in personnes if valeurs_mains[p] == plus_petite_valeur ]
print("Les autres personnes ayant un Jap Jap de plus petite valeur sont {} !".format(gagnantes))
nombre_min_carte = min([len(self.mains[p]) for p in gagnantes])
gagnante = min([p for p in gagnantes if len(self.mains[p]) == nombre_min_carte])
print("La personne gagnant la manche est {}.".format(gagnante))
# on marque les scores
print("\nOn marque les scores !")
print("==> La personne {} a gagné ! Avec un Jap Jap de valeur {} !".format(gagnante, plus_petite_valeur))
for autre_personne in [ p for p in personnes if p != gagnante ]:
marque_point = valeur_main(self.mains[autre_personne])
print("- La personne {} n'a pas gagné, et marque {} points".format(autre_personne, marque_point))
self.scores[autre_personne] += marque_point
# si la personne s'est prise un contre jap jap, elle marque +25 et pas son total de cartes en main
print("- La personne {} n'a pas gagné et a subi un CONTRE JAP JAP ! Elle marque +25 points.")
if contre_JapJap:
self.scores[personne] -= valeur_main(self.mains[personne])
self.scores[personne] += self.malusContreJapJap
print("\nA la fin de cette manche :")
for personne in self.personnes:
nom = self.nomsPersonnes[personne] if self.nomsPersonnes is not None else personne
score = self.scores[personne]
print(" + Personne {} a un score de {}.".format(nom, score))
# si un score est >= 90
if max(self.scores) >= self.scoreMax:
# quelqu'un a perdu cette partie !
for personne in personnes:
score = self.scores[personne]
if score == max(self.scores):
nom = self.nomsPersonnes[personne] if self.nomsPersonnes is not None else personne
print("\n==> La personne {} a perdu, avec un score de {}.".format(nom, score))
raise FinDunePartie
raise FinDuneManche
# on pose les cartes jetées
self.visibles = jetees
# et on enlève les cartes jetées de sa main
nouvelle_main = self.mains[personne]
for carte_jetee in jetees:
nouvelle_main.remove(carte_jetee)
# et ça continue
# -
# ## Lancement du jeu intéractif
jeu = EtatJeu(nomsPersonnes=["Alice", "Bob"])
jeu.debut_jeu()
import asyncio
asyncio.ensure_future(jeu.enchainer_les_tours())
# # Conclusion
| Simulations du jeu de carte Jap-Jap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
df = pd.read_excel('data for Problem1.xlsx')
x = df['x']
y = df['y']
x = (x-x.mean())/x.std()
itera = 1000
alp = 0.01
theta[0]=0
theta[1]=0
m = y.size
past_cost=[]
past_theta=[]
for i in range(itera):
weight = 0
weightt = 0
err = 0
for j in range(m):
err = err + (y[j]-(theta[0]+theta[1]*x[j]))**2
weight = weight - (y[j]-(theta[0]+theta[1]*x[j]))
weightt = weightt - (y[j]-(theta[0]+theta[1]*x[j]))*x[j]
past_cost.append(err*(1/(2*m)))
theta[0] = theta[0]-weight*alp*(1/m)
theta[1] = theta[1]-weightt*alp*(1/m)
past_theta.append(theta)
print ("Following are the thetas", theta[0],theta[1])
#print(past_theta)
plt.title('Cost Function J')
plt.xlabel('No. of iterations')
plt.ylabel('Cost')
plt.plot(past_cost)
plt.show()
plt.title('Theta')
plt.xlabel('No. of iterations')
plt.ylabel('Theta')
plt.plot(past_theta)
plt.show()
plt.plot(y,x,'bo')
plt.show()
y1 = theta[0]+theta[1]*x
#print(y1.shape)
plt.plot(y1,x)
| Gradient_descent_without_matrix_lab2_q5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# create new directory and unpacking the archive with data
# !mkdir ./data/unzipped && unzip ./data/archive.zip -d ./data/unzipped
# ## Data set from [Kaggle](https://www.kaggle.com/lava18/google-play-store-apps)
# Web scraped data of 10k Play Store apps for analysing the Android market.
#
# *This information is scraped from the Google Play Store. This app information would not be available without it.*
#
# - **googleplaystore**. details of the applications on Google Play.
# - **googleplaystore_user_reviews**
# This file contains the first 'most relevant' 100 reviews for each app. Each review text/comment has been pre-processed and attributed with 3 new features - Sentiment, Sentiment Polarity and Sentiment Subjectivity.
# install libraly
# !pip install pandas-profiling
# import modules
import os
import pandas as pd
from pandas_profiling import ProfileReport
# +
# assign path
path, dirs, files = next(os.walk(os.getcwd() + "/data/unzipped/"))
file_count = len(files)
# create empty list
dataframes_list = []
# append datasets to the list
for i in range(file_count):
temp_df = pd.read_csv(os.getcwd() + "/data/unzipped/" + files[i])
dataframes_list.append(temp_df)
# display datasets
for dataset in dataframes_list:
display(dataset)
# +
profiling_user_review = ProfileReport(dataframes_list[0]
, title="Pandas Profiling Report for User Reviews"
, explorative=True)
profiling_user_review.to_file(os.getcwd() + "/data/profiling_user_review.html")
# +
profiling_general_app = ProfileReport(dataframes_list[1]
, title="Pandas Profiling Report for all app"
, explorative=True)
profiling_general_app.to_file(os.getcwd() + "/data/profiling_general_app.html")
| jet/2_eda_google_play_app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Deploying Campaigns and Filters<a class="anchor" id="top"></a>
#
# In this notebook, you will deploy and interact with campaigns in Amazon Personalize.
#
# 1. [Introduction](#intro)
# 1. [Create campaigns](#create)
# 1. [Interact with campaigns](#interact)
# 1. [Batch recommendations](#batch)
# 1. [Wrap up](#wrapup)
#
# ## Introduction <a class="anchor" id="intro"></a>
# [Back to top](#top)
#
# At this point, you should have several solutions and at least one solution version for each. Once a solution version is created, it is possible to get recommendations from them, and to get a feel for their overall behavior.
#
# This notebook starts off by deploying each of the solution versions from the previous notebook into individual campaigns. Once they are active, there are resources for querying the recommendations, and helper functions to digest the output into something more human-readable.
#
# As you with your customer on Amazon Personalize, you can modify the helper functions to fit the structure of their data input files to keep the additional rendering working.
#
# To get started, once again, we need to import libraries, load values from previous notebooks, and load the SDK.
# +
import time
from time import sleep
import json
from datetime import datetime
import uuid
import random
import boto3
import botocore
from botocore.exceptions import ClientError
import pandas as pd
# -
# %store -r
# +
personalize = boto3.client('personalize')
personalize_runtime = boto3.client('personalize-runtime')
# Establish a connection to Personalize's event streaming
personalize_events = boto3.client(service_name='personalize-events')
# -
# ## Create campaigns <a class="anchor" id="create"></a>
# [Back to top](#top)
#
# A campaign is a hosted solution version; an endpoint which you can query for recommendations. Pricing is set by estimating throughput capacity (requests from users for personalization per second). When deploying a campaign, you set a minimum throughput per second (TPS) value. This service, like many within AWS, will automatically scale based on demand, but if latency is critical, you may want to provision ahead for larger demand. For this POC and demo, all minimum throughput thresholds are set to 1. For more information, see the [pricing page](https://aws.amazon.com/personalize/pricing/).
#
# Let's start deploying the campaigns.
# ### User Personalization
#
# Deploy a campaign for your User Personalization solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
#
# You now have the option of setting additional configuration for the campaign, which allows you to adjust the exploration Amazon Personalize does for the item recommendations and therefore adjust the results. These settings are only available if you’re creating a campaign whose solution version uses the user-personalization recipe. The configuration options are as follows:
#
# explorationWeight – Higher values for explorationWeight signify higher exploration; new items with low impressions are more likely to be recommended. A value of 0 signifies that there is no exploration and results are ranked according to relevance. You can set this parameter in a range of [0,1] and its default value is 0.3.
#
# explorationItemAgeCutoff – This is the maximum duration in days relative to the latest interaction(event) timestamp in the training data. For example, if you set explorationItemAgeCutoff to 7, the items with an age over or equal to 7 days aren’t considered cold items and there is no exploration on these items. You may still see some items older than or equal to 7 days in the recommendation list because they’re relevant to the user’s interests and are of good quality even without the help of the exploration. The default value for this parameter is 30, and you can set it to any value over 0.
#
# We will use the explorationWeight in the "06_Adding_New_Items_and_Updating_Model.ipynb", so we will set the explorationWeight to 0.9, which will cause the new (cold start) items to be recommended more often.
# +
userpersonalization_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-userpersonalization",
solutionVersionArn = userpersonalization_solution_version_arn,
minProvisionedTPS = 1,
campaignConfig={
'itemExplorationConfig': {
'explorationWeight': '0.9'
}
}
)
userpersonalization_campaign_arn = userpersonalization_create_campaign_response['campaignArn']
print(json.dumps(userpersonalization_create_campaign_response, indent=2))
# -
# ### SIMS
#
# Deploy a campaign for your SIMS solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
# +
sims_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-SIMS",
solutionVersionArn = sims_solution_version_arn,
minProvisionedTPS = 1
)
sims_campaign_arn = sims_create_campaign_response['campaignArn']
print(json.dumps(sims_create_campaign_response, indent=2))
# -
# ### Personalized Ranking
#
# Deploy a campaign for your personalized ranking solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
# +
rerank_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-rerank",
solutionVersionArn = rerank_solution_version_arn,
minProvisionedTPS = 1
)
rerank_campaign_arn = rerank_create_campaign_response['campaignArn']
print(json.dumps(rerank_create_campaign_response, indent=2))
# -
# ### View campaign creation status
#
# As promised, how to view the status updates in the console:
#
# * In another browser tab you should already have the AWS Console up from opening this notebook instance.
# * Switch to that tab and search at the top for the service `Personalize`, then go to that service page.
# * Click `View dataset groups`.
# * Click the name of your dataset group, most likely something with POC in the name.
# * Click `Campaigns`.
# * You will now see a list of all of the campaigns you created above, including a column with the status of the campaign. Once it is `Active`, your campaign is ready to be queried.
#
# Or simply run the cell below to keep track of the campaign creation status.
# +
in_progress_campaigns = [
userpersonalization_campaign_arn,
sims_campaign_arn,
rerank_campaign_arn
]
max_time = time.time() + 3*60*60 # 3 hours
while time.time() < max_time:
for campaign_arn in in_progress_campaigns:
version_response = personalize.describe_campaign(
campaignArn = campaign_arn
)
status = version_response["campaign"]["status"]
if status == "ACTIVE":
print("Build succeeded for {}".format(campaign_arn))
in_progress_campaigns.remove(campaign_arn)
elif status == "CREATE FAILED":
print("Build failed for {}".format(campaign_arn))
in_progress_campaigns.remove(campaign_arn)
if len(in_progress_campaigns) <= 0:
break
else:
print("At least one campaign build is still in progress")
time.sleep(60)
# -
# ## Create Static Filters <a class="anchor" id="interact"></a>
# [Back to top](#top)
#
# Now that all campaigns are deployed and active, we can create filters. Filters can be created for both Items and Events. Filters can also be created dynamically for "IN" and "=" operation, which is covered in the 05_Interacting_with_Campaigns_and_Filters notebook. For range queries, you need to continue to use static filters.
# Range queries use the following operations: NOT IN, <, >, <=, and >=.
#
# A few common use cases for static filters in Video On Demand are:
#
# Categorical filters based on Item Metadata (that are range based) - Often your item metadata will have information about the title such as year, user rating, available date. Filtering on these can provide recommendations within that data, such as movies that are available after a specific date, movies rated over 3 stars, movies from the 1990s etc
#
# User Demographic ranges - you may want to recommend content to specific age demographics, for this you can create a filter that is specific to a age range like over 18, over 18 AND under 30, etc).
#
# Lets look at the item metadata and user interactions, so we can get an idea what type of filters we can create.
# +
# Create a dataframe for the items by reading in the correct source CSV
items_meta_df = pd.read_csv(data_dir + '/item-meta.csv', sep=',', index_col=0)
# Render some sample data
items_meta_df.head(10)
# -
# Since there are alot of genres to filter on, we will create a filter, using the dynamic variable $GENRE, this will allow us to pass in the variable at runtime rather than create a static filter for each genre.
creategenrefilter_response = personalize.create_filter(name='Genre',
datasetGroupArn=dataset_group_arn,
filterExpression='INCLUDE ItemID WHERE Items.GENRE IN ($GENRE)'
)
genre_filter_arn = creategenrefilter_response['filterArn']
# Since we now have the year available in our item metadata, lets create a decade filter to recommend only movies releaseed in a given decade. A soft limit of Personalize at this time is 10 total filters, so we will create 7 decade filters for this workshop, leaving room for additional static and dynamic filters
# Now create a list for the metadata decade filters and then create the actual filters with the cells below. Note this will take a few minutes to complete.
decades_to_filter = [1950,1960,1970,1980,1990,2000,2010]
# Create a list for the filters:
decade_filter_arns = []
# Iterate through Decades
for decade in decades_to_filter:
# Start by creating a filter
current_decade = str(decade)
next_decade = str(decade + 10)
try:
createfilter_response = personalize.create_filter(
name=current_decade + "s",
datasetGroupArn=dataset_group_arn,
filterExpression='INCLUDE ItemID WHERE Items.YEAR >= '+ current_decade +' AND Items.YEAR < '+ next_decade +''
)
# Add the ARN to the list
decade_filter_arns.append(createfilter_response['filterArn'])
print("Creating: " + createfilter_response['filterArn'])
# If this fails, wait a bit
except ClientError as error:
# Here we only care about raising if it isnt the throttling issue
if error.response['Error']['Code'] != 'LimitExceededException':
print(error)
else:
time.sleep(120)
createfilter_response = personalize.create_filter(
name=current_decade + "s",
datasetGroupArn=dataset_group_arn,
filterExpression='INCLUDE ItemID WHERE Items.YEAR >= '+ current_decade +' AND Items.YEAR < '+ next_decade +''
)
# Add the ARN to the list
decade_filter_arns.append(createfilter_response['filterArn'])
print("Creating: " + createfilter_response['filterArn'])
# Lets look at the type of event data we can create filters for
# +
# Create a dataframe for the interactions by reading in the correct source CSV
interactions_df = pd.read_csv(data_dir + '/interactions.csv', sep=',', index_col=0)
# Render some sample data
interactions_df.head(10)
# -
# Since we may want to recommend items already watched, or not watchet yet, lets also create 2 event filters for watched and unwatched content.
# +
createwatchedfilter_response = personalize.create_filter(name='watched',
datasetGroupArn=dataset_group_arn,
filterExpression='INCLUDE ItemID WHERE Interactions.event_type IN ("watch")'
)
createunwatchedfilter_response = personalize.create_filter(name='unwatched',
datasetGroupArn=dataset_group_arn,
filterExpression='EXCLUDE ItemID WHERE Interactions.event_type IN ("watch")'
)
# -
# Before we are done we will want to add those filters to a list as well so they can be used later.
interaction_filter_arns = [createwatchedfilter_response['filterArn'], createunwatchedfilter_response['filterArn']]
# %store sims_campaign_arn
# %store userpersonalization_campaign_arn
# %store rerank_campaign_arn
# %store decade_filter_arns
# %store genre_filter_arn
# %store interaction_filter_arns
# You're all set to move on to the last exploratory notebook: `05_Interacting_with_Campaigns_and_Filters.ipynb`. Open it from the browser and you can start interacting with the Campaigns and gettign recommendations!
| next_steps/workshops/POC_in_a_box/completed/04_Deploying_Campaigns_and_Filters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-PythonData] *
# language: python
# name: conda-env-.conda-PythonData-py
# ---
# import all dependencies
from bs4 import BeautifulSoup
from splinter import Browser
from splinter.exceptions import ElementDoesNotExist
import time
import pandas as pd
executable_path = {"executable_path": "./chromedriver.exe"}
browser = Browser("chrome", **executable_path)
url = 'https://mars.nasa.gov/news/'
browser.visit(url)
# ## Step 1 - Scraping
# visit the NASA Mars News site and scrape headlines
browser.visit(url)
time.sleep(5)
nasa_html = browser.html
nasa_soup = BeautifulSoup(nasa_html, 'html.parser')
# ### NASA Mars News
news_list = nasa_soup.find('ul', class_='item_list')
first_item = news_list.find('li', class_='slide')
nasa_headline = first_item.find('div', class_='content_title').text
nasa_teaser = first_item.find('div', class_='article_teaser_body').text
print(nasa_headline)
print(nasa_teaser)
# ### JPL Mars Space Images - Featured Image
# +
# Scrape the featured image from JPL website
jpl_url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(jpl_url)
time.sleep(1)
browser.click_link_by_partial_text('FULL IMAGE')
time.sleep(1)
expand = browser.find_by_css('a.fancybox-expand')
expand.click()
time.sleep(1)
jpl_html = browser.html
jpl_soup = BeautifulSoup(jpl_html, 'html.parser')
img_relative = jpl_soup.find('img', class_='fancybox-image')['src']
image_path = f'https://www.jpl.nasa.gov{img_relative}'
print(image_path)
# -
# ### Mars Weather
# Scrape latest tweet from the mars weather report twitter
mars_tweets_url = 'https://twitter.com/marswxreport?lang=en'
browser.visit(mars_tweets_url)
time.sleep(5)
mars_tweets_html = browser.html
mars_tweets_soup = BeautifulSoup(mars_tweets_html, 'html.parser')
mars_weather_tweet = mars_tweets_soup.find_all('article')
for x in mars_weather_tweet:
tweet = x.find("div",attrs={"data-testid":"tweet"})
tweet_list=tweet.find("div",class_="css-901oao r-hkyrab r-1qd0xha r-a023e6 r-16dba41 r-ad9z0x r-bcqeeo r-bnwqim r-qvutc0")
for j in tweet_list:
print(j.parent.text)
# ### Mars Facts
# +
# Scrape Mars facts table
mars_facts_url = 'https://space-facts.com/mars/'
browser.visit(mars_facts_url)
time.sleep(5)
mars_facts_html = browser.html
mars_facts_soup = BeautifulSoup(mars_facts_html, 'html.parser')
mars_facts_soup.body.find_all('table', class_="tablepress tablepress-id-p-mars")
# -
# ### Mars Hemispheres
# +
# scrape images of Mars' hemispheres from the USGS site
mars_hemisphere_url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
hemi_dicts = []
for i in range(1,9,2):
hemi_dict = {}
browser.visit(mars_hemisphere_url)
time.sleep(5)
hemispheres_html = browser.html
hemispheres_soup = BeautifulSoup(hemispheres_html, 'html.parser')
hemi_name_links = hemispheres_soup.find_all('a', class_='product-item')
hemi_name = hemi_name_links[i].text.strip('Enhanced')
detail_links = browser.find_by_css('a.product-item')
detail_links[i].click()
time.sleep(5)
browser.find_link_by_text('Sample').first.click()
time.sleep(5)
browser.windows.current = browser.windows[-1]
hemi_img_html = browser.html
browser.windows.current = browser.windows[0]
browser.windows[-1].close()
hemi_img_soup = BeautifulSoup(hemi_img_html, 'html.parser')
hemi_img_path = hemi_img_soup.find('img')['src']
print(hemi_name)
hemi_dict['title'] = hemi_name.strip()
print(hemi_img_path)
hemi_dict['img_url'] = hemi_img_path
hemi_dicts.append(hemi_dict)
# -
| .ipynb_checkpoints/mission_to_mars-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# - spectrogram アルゴリズムの比較
# # Const
n_fft = 512
hop_length = int(n_fft/2)
# # Import everything I need :)
import numpy as np
import scipy as sp
import librosa
import matplotlib.pyplot as plt
# # EDA
path = librosa.util.example_audio_file()
signal, sr = librosa.load(path, sr=None)
f, t, spec_sp = sp.signal.spectrogram(signal, nperseg=n_fft, noverlap=hop_length)
spec_sp.shape
spec_librosa = np.abs(librosa.stft(signal, n_fft=n_fft, hop_length=hop_length))
spec_librosa.shape
(spec_sp/spec_sp.mean()).mean()
spec_librosa.mean()
| notebook/01_spectrogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pyspark
if not 'sc' in globals():
sc = pyspark.SparkContext()
text_file = sc.textFile("Spark File Words.ipynb")
counts = text_file.flatMap(lambda line: line.split(" ")) \
.map(lambda word: (word, 1)) \
.reduceByKey(lambda a, b: a + b)
for x in counts.collect():
print x
| ch10/Spark File Words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Reading and cleaning jsons
import json
import re
import numpy as np
import pandas as pd
from sklearn.cluster import AgglomerativeClustering
import matplotlib.pyplot as plt
from bert_embedding import BertEmbedding
import RAKE
import operator
from nltk.stem import PorterStemmer
# +
video_texts = []
# dict list
with open('./Channel3/transcript_channel3.json') as f:
data = json.load(f)
for i in range(len(data)):
transcript = list(data[i].values())
if transcript[2] is not None: #Change index to 2 for channel 3, 0 for others
string = transcript[2][63:]
string = re.sub("([\<\[]).*?([\>\]])", "\g<1>\g<2>", string)
string = re.sub("[\<\[].*?[\>\]]", "", string).rstrip('\n')
arr = string.split('\n')
clean_arr = []
for sentence in arr:
if sentence != '' and sentence != ' ' and 'align' not in sentence:
clean_arr.append(sentence)
clean_text = []
for j in range(0,len(clean_arr),3):
clean_text.append(clean_arr[j])
video_text = ''
for sen in string_arr:
video_text += sen+' '
video_texts.append(video)
# -
len(video_texts)
# ## Preprocessing text (extracting keywords)
bert_embedding = BertEmbedding()
# result = bert_embedding(video_texts)
cleaned_texts = []
for v in video_texts:
cleaned_texts.append(''.join([i for i in v if not i.isdigit()]))
stop_dir = "SmartStoplist.txt"
rake_object = RAKE.Rake(stop_dir)
# +
#####
# STEMMING (OPTIONAL)
####
# from nltk.tokenize import sent_tokenize, word_tokenize
# ws = word_tokenize(cleaned_texts[4].lower())
# for w in ws:
# print(ps.stem(w))
# # for cleaned_texts[4].lower()
# -
video_embeds = []
list_of_kwords = []
for text in cleaned_texts:
keywords = rake_object.run(text)
list_of_kwords.append(keywords[:30])
for k in keywords[:30]:
key_embed = bert_embedding([k[0]])
mean_avg = []
for embed in key_embed:
to_avg = []
for e in embed[1]:
to_avg.append(e)
mean_avg.append(np.mean(to_avg, axis=0))
video_embeds.append(np.mean(mean_avg,axis=0))
# keys.append(k[0])
len(video_embeds)
# ## Hierarchical clustering and plots
# +
from sklearn.cluster import AgglomerativeClustering
cluster = AgglomerativeClustering(n_clusters=3, affinity='euclidean', linkage='ward')
preds = cluster.fit_predict(video_embeds)
# -
preds
# Checking results of clustering
cluster = 2
indices = [i for i, x in enumerate(preds) if x == cluster]
print(len(indices))
# Keywords of video in specific cluster
list_of_kwords[2]
# +
# Performing tsne to project video embeddings
from sklearn.manifold import TSNE
tsne = TSNE(n_components=2, verbose=1, perplexity=42, n_iter=300)
tsne_results = tsne.fit_transform(video_embeds)
# -
# Plotting T-SNE results
plt.figure(figsize=(10, 7))
scat = plt.scatter(tsne_results[:,0], tsne_results[:,1], c=cluster.labels_, cmap='rainbow')
# +
# Dendogram from the hierarchical clustering
import scipy.cluster.hierarchy as shc
plt.figure(figsize=(10, 10))
plt.title("Video Dendograms")
dend = shc.dendrogram(shc.linkage(video_embeds, method='ward'))
| HierClustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest Analysis of Cozie Data Tier 2
# - V16
# - Tier2 data only
# - Using continuous data instead of cluster thus avoiding the hot encoding
#
# +
# %matplotlib inline
import numpy as np
import pandas as pd
from datetime import datetime
import os
# Preperation
from sklearn.model_selection import train_test_split
# Modeling
from sklearn.ensemble import RandomForestClassifier
from sklearn.cluster import KMeans
# Visualisations
from sklearn.tree import export_graphviz # Note that you need to brew install graphviz on your local machine
import pydot
import seaborn as sns
import matplotlib.pyplot as plt
# Evaluation
from sklearn import metrics
# User Defined Functions
import cozie_functions
# -
# Add Data Folder to Path
data_path = os.path.abspath(os.path.join(os.path.dirname( "__file__" ), '..', 'data'))
# # Reading Data
# In this case we are first reading the data, and then reorganising them into groups.
# - First drop all unecessary data
# - Then drop all Sensing Data
# - Then drop all mbient Data
#
# Afterwards, created normalised datasets of the average value. See `cozie_functions.py`
# The following participants took part in the experiment:
participant_ids = ['cresh' + str(id).zfill(2) for id in range(1,31)]
print(participant_ids)
# +
feature_set_df = pd.read_csv(os.path.join(data_path, '2019-11-15_cozie_full_masked.csv'))
feature_set_df.drop(['Unnamed: 0', 'index', 'comfort_cozie', 'Space_id', 'Longitude',
'Latitude', 'co2_sensing', 'voc_sensing', 'Floor', 'lat_cozie', 'lon_cozie', 'responseSpeed_cozie'], axis=1, inplace=True)
feature_set_df.dropna(subset=['thermal_cozie','light_cozie','noise_cozie',
'temperature_sensing', 'temperature_mbient',
'heartRate_cozie'], inplace=True)
# Drop User's that were trialing and not in the experiment
feature_set_df = feature_set_df[feature_set_df.user_id.isin(participant_ids)]
#feature_set_df["thermal_cozie"] = feature_set_df["thermal_cozie"].astype(str)
# +
## Added in V4
#feature_set_df.drop(["noise_cozie", "light_cozie"], axis=1, inplace=True)
feature_set_df.head()
# -
feature_set_df.describe()
# +
prefer_cooler_df = feature_set_df[feature_set_df["thermal_cozie"] == 11]
thermal_comfy_df = feature_set_df[feature_set_df["thermal_cozie"] == 10]
prefer_warmer_df = feature_set_df[feature_set_df["thermal_cozie"] == 9]
prefer_dimmer_df = feature_set_df[feature_set_df["light_cozie"] == 11]
visually_comfy_df = feature_set_df[feature_set_df["light_cozie"] == 10]
prefer_brighter_df = feature_set_df[feature_set_df["light_cozie"] == 9]
prefer_quieter_df = feature_set_df[feature_set_df["noise_cozie"] == 11]
aurally_comfy_df = feature_set_df[feature_set_df["noise_cozie"] == 10]
prefer_louder_df = feature_set_df[feature_set_df["noise_cozie"] == 9]
# -
# +
ax = sns.kdeplot(thermal_comfy_df.temperature_sensing, thermal_comfy_df.humidity_sensing,
cmap="Greens")
ax = sns.kdeplot(prefer_warmer_df.temperature_sensing, prefer_warmer_df.humidity_sensing,
cmap="Reds",)
ax = sns.kdeplot(prefer_cooler_df.temperature_sensing, prefer_cooler_df.humidity_sensing,
cmap="Blues")
# -
feature_set_df.describe()
palette=sns.color_palette("Set2", 3)
ax = sns.scatterplot(x=feature_set_df.temperature_sensing, y=feature_set_df.humidity_sensing,
hue=feature_set_df["thermal_cozie"], palette= [palette[1],palette[0],palette[2]], alpha = 0.5, legend = False)
plt.savefig("temp_hum_scatter.pdf")
palette=sns.color_palette("Set2", 3)
ax = sns.scatterplot(x=feature_set_df.temperature_sensing, y=feature_set_df.temperature_mbient,
hue=feature_set_df["thermal_cozie"], palette= [palette[1],palette[0],palette[2]], alpha = 0.5, legend = False)
plt.savefig("temp_temp_scatter.pdf")
# +
fig1, axes = plt.subplots(2, 3, figsize=(12, 6))
palette=sns.color_palette("Set2", 3)
#Plot Mbient
sns.kdeplot(prefer_cooler_df["temperature_mbient"], shade=True, color=palette[2], label = "Prefer Cooler", ax=axes[0,0])
sns.kdeplot(prefer_warmer_df["temperature_mbient"], shade=True, color=palette[1], label = "Prefer Warmer", ax=axes[0,0])
sns.kdeplot(thermal_comfy_df["temperature_mbient"], shade=True, color=palette[0], label = "Thermally Comfy", ax=axes[0,0])
# Plot Sensing
sns.kdeplot(prefer_cooler_df["humidity_sensing"], shade=True, color=palette[2], label = "Prefer Cooler", ax=axes[0,1])
sns.kdeplot(prefer_warmer_df["humidity_sensing"], shade=True, color=palette[1], label = "Prefer Warmer", ax=axes[0,1])
sns.kdeplot(thermal_comfy_df["humidity_sensing"], shade=True, color=palette[0], label = "Thermally Comfy", ax=axes[0,1])
sns.kdeplot(prefer_cooler_df["temperature_sensing"], shade=True, color=palette[2], label = "Prefer Cooler", ax=axes[0,2])
sns.kdeplot(prefer_warmer_df["temperature_sensing"], shade=True, color=palette[1], label = "Prefer Warmer", ax=axes[0,2])
sns.kdeplot(thermal_comfy_df["temperature_sensing"], shade=True, color=palette[0], label = "Thermally Comfy", ax=axes[0,2])
# Sensing Light
sns.kdeplot(prefer_dimmer_df["light_sensing"], shade=True, color=palette[2], label = "Prefer Dimmer", ax=axes[1,0])
sns.kdeplot(prefer_brighter_df["light_sensing"], shade=True, color=palette[1], label = "Prefer Brighter", ax=axes[1,0])
sns.kdeplot(visually_comfy_df["light_sensing"], shade=True, color=palette[0], label = "Visually Comfy", ax=axes[1,0])
sns.kdeplot(prefer_quieter_df["noise_sensing"], shade=True, color=palette[2], label = "Prefer Quieter", ax=axes[1,2])
sns.kdeplot(prefer_louder_df["noise_sensing"], shade=True, color=palette[1], label = "Prefer Louder", ax=axes[1,2])
sns.kdeplot(aurally_comfy_df["noise_sensing"], shade=True, color=palette[0], label = "Aurally Comfy", ax=axes[1,2])
axes[0,0].set_xlabel('Near Body Temperature (C)')
axes[0,1].set_xlabel('Humidity (%)')
axes[0,2].set_xlabel('Temperature (C)')
axes[1,0].set_xlabel('Illuminance (lux)')
axes[1,2].set_xlabel('Noise (dB)')
fig1.tight_layout()
axes[0,0].get_legend().remove()
#axes[0,1].get_legend().remove()
axes[0,2].get_legend().remove()
#sns.kdeplot(x, bw=.2, label="bw: 0.2")
#sns.kdeplot(x, bw=2, label="bw: 2")
#plt.legend();
plt.savefig("DensityPlots.pdf")
# +
fig1, axes = plt.subplots(2, 3, figsize=(12, 6))
palette=sns.color_palette("Set2", 3)
#Plot Mbient
sns.distplot(prefer_cooler_df["temperature_mbient"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,0], rug=False)
sns.distplot(prefer_warmer_df["temperature_mbient"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,0], rug=False)
sns.distplot(thermal_comfy_df["temperature_mbient"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,0], rug=False)
# Plot Sensing
sns.distplot(prefer_cooler_df["humidity_sensing"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,1], rug=False)
sns.distplot(prefer_warmer_df["humidity_sensing"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,1], rug=False)
sns.distplot(thermal_comfy_df["humidity_sensing"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,1], rug=False)
sns.distplot(prefer_cooler_df["temperature_sensing"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,2], rug=False)
sns.distplot(prefer_warmer_df["temperature_sensing"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,2], rug=False)
sns.distplot(thermal_comfy_df["temperature_sensing"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,2], rug=False)
# Sensing Light
sns.distplot(prefer_dimmer_df["light_sensing"], color=palette[2], kde=False, label = "Prefer Dimmer", ax=axes[1,0], rug=False)
sns.distplot(prefer_brighter_df["light_sensing"], color=palette[1], kde=False, label = "Prefer Brighter", ax=axes[1,0], rug=False)
sns.distplot(visually_comfy_df["light_sensing"], color=palette[0], kde=False, label = "Visually Comfy", ax=axes[1,0], rug=False)
sns.distplot(prefer_quieter_df["noise_sensing"], color=palette[2], kde=False, label = "Prefer Quieter", ax=axes[1,2], rug=False)
sns.distplot(prefer_louder_df["noise_sensing"], color=palette[1], kde=False, label = "Prefer Louder", ax=axes[1,2], rug=False)
sns.distplot(aurally_comfy_df["noise_sensing"], color=palette[0], kde=False, label = "Aurally Comfy", ax=axes[1,2], rug=False)
## Plot lines
#Plot Mbient
sns.distplot(prefer_cooler_df["temperature_mbient"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_warmer_df["temperature_mbient"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(thermal_comfy_df["temperature_mbient"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
# Plot Sensing
sns.distplot(prefer_cooler_df["humidity_sensing"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,1], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_warmer_df["humidity_sensing"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,1], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(thermal_comfy_df["humidity_sensing"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,1], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_cooler_df["temperature_sensing"], color=palette[2], kde=False, label = "Prefer Cooler", ax=axes[0,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_warmer_df["temperature_sensing"], color=palette[1], kde=False, label = "Prefer Warmer", ax=axes[0,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(thermal_comfy_df["temperature_sensing"], color=palette[0], kde=False, label = "Thermally Comfy", ax=axes[0,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
# Sensing Light
sns.distplot(prefer_dimmer_df["light_sensing"], color=palette[2], kde=False, label = "Prefer Dimmer", ax=axes[1,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_brighter_df["light_sensing"], color=palette[1], kde=False, label = "Prefer Brighter", ax=axes[1,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(visually_comfy_df["light_sensing"], color=palette[0], kde=False, label = "Visually Comfy", ax=axes[1,0], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_quieter_df["noise_sensing"], color=palette[2], kde=False, label = "Prefer Quieter", ax=axes[1,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(prefer_louder_df["noise_sensing"], color=palette[1], kde=False, label = "Prefer Louder", ax=axes[1,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
sns.distplot(aurally_comfy_df["noise_sensing"], color=palette[0], kde=False, label = "Aurally Comfy", ax=axes[1,2], rug=False, hist_kws={"histtype": "step", "linewidth": 2, "alpha": 1})
axes[0,0].set_xlabel('Near Body Temperature (C)')
axes[0,1].set_xlabel('Humidity (%)')
axes[0,2].set_xlabel('Temperature (C)')
axes[1,0].set_xlabel('Illuminance (lux)')
axes[1,2].set_xlabel('Noise (dB)')
fig1.tight_layout()
#axes[0,0].get_legend().remove()
#axes[0,1].get_legend().remove()
#axes[0,2].get_legend().remove()
#sns.kdeplot(x, bw=.2, label="bw: 0.2")
#sns.kdeplot(x, bw=2, label="bw: 2")
#plt.legend();
plt.savefig("Histograms.pdf")
# -
type(palette)
| publications-plots/PublicationPlots_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nelslindahlx/Data-Analysis/blob/master/Basic_TensorFlow_hello_world_example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bjH2uAXaDw73" colab_type="text"
# Installing TensorFlow in this Jupyter Notebook
# + id="Tr96wvcxDsvT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 658} outputId="4a175cf9-2d77-492c-dfa2-740daa5850a7"
# !pip3 install tensorflow
# + [markdown] id="F9o0siL-ERnP" colab_type="text"
# Now test your TensorFlow Installation in this Jupyter notebook
# + id="_5SKDI_2ERYC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1996cf8f-685c-4468-d49f-fb73aa839c25"
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="WdedvVUlEUT0" colab_type="text"
# You installed TensorFlow and checked to see what version is running ;)
# + [markdown] id="qaGpjs_KMFBe" colab_type="text"
# Basic TensorFlow hello world example
# + id="47c896IMMEO_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5b3ed489-185c-43f3-faf7-25fc5f62a77a"
msg = tf.constant('Fun text saying hellow world or something else in TensorFlow 2.0')
tf.print(msg)
# + [markdown] id="psQa8moYPE2-" colab_type="text"
# A little more invovled example
# + id="QJTRRV7bPFAW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="79f71a76-e957-4c69-d205-794b2b7548ba"
h = tf.constant("hello")
w = tf.constant("world!")
hw = h + w
msg = tf.constant(hw)
tf.print(msg)
# + [markdown] id="RpG2zReVPp2h" colab_type="text"
# What went wrong? Why is the pace missing between hello and world? The answer is simple.. we forgot to add it to the printing...
# + id="kwsiffHyPqBM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="18df1c1d-a4d4-4d18-de40-b0210ea33ae7"
h = tf.constant("hello")
s = tf.constant(" ")
w = tf.constant("world!")
hw = h + s + w
msg = tf.constant(hw)
tf.print(msg)
| Basic_TensorFlow_hello_world_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="cac470df-29e7-4148-9bbd-d8b9a32fa570" tags=[]
# # (not_done_review)그로킹 심층 강화학습
# > 강찬석
#
# - toc:true
# - branch: master
# - badges: true
# - comments: false
# - author: 최서연
# - categories: [Reinforcement Learning]
# -
# # $\star$ 목표: 일주일에 몇 장씩이라도 보기!
# ref: https://goodboychan.github.io/book
# ### 1
# 심층강화학습 deep reinforcement learning DRL 이란 머신 러닝 기법 중 하나.
# - 지능이 요구되는 문제를 해결할 수 있도록 인공지능 컴퓨터 프로그램을 개발하는데 사용
# - 시행착오를 통해 얻은 반응을 학습
# 심층 강화학습은 문제에 대한 접근법이다.
# - **에이전트agent**: 의사를 결정하는 객체 자체
# - ex) 사물을 집는 로봇 학습시킬때 의사 결정을 좌우하는 코드와 연관
# - **환경unvironmnet**: 에이전트(의사 결정) 이외의 모든 것
# - ex) 의사결정하는 로봇(객체) 제외한 모든 것이 환경의 일부
# - **상태 영역state space**: 변수가 가질 수 있는 모든 값들의 집합
# - **관찰observation**: 에이전트가 관찰할 수 있는 상태의 일부
# - **전이 함수transition function** 에이전트와 환경 사이의 관계를 정의한 함수
# - **보상 함수reward function**: 행동에 대한 반응으로 환경이 제공한 보상 신호와 관련된 함수
# - **모델model**: 전이와 보상함수를 표현 가능
#
# 에이전트의 3단계 과정
# 1. 환경과 상호작용을 나누고
# 2. 행동에 대해서 평가를 하며,
# 3. 받은 반응을 개선시킨다.
# - **에피소드형 업무episodic task**: 게임과 같이 자연적으로 끝나는 업무
# - **연속형업무continuing task**: 앞으로 가는 동작을 학습하는 경우
#
# - 연속형 피드백이 야기하는 문제
# - **시간적 가치 할당 문제tamporal credit assignment problem**:문제에 시간적 개념이 들어가 있고, 행동에도 지연된 속성이 담겨 있으면, 보상에 대한 가치를 부여하기 어렵다.
# - 평가 가능한 피드백이 야기하는 문제
# - **탐험과 착취 간의 트레이드 오프exploration versus explotation trade-off**: 에이전트는 현재 가지고 있는 정보에서 얻을 수 있는 가장 좋은 것과 정보를 새로 얻는 것 간의 균형을 맞출 수 있어야 한다.
# ### 2
# 에이전트의 세 단계 과정
# 1. 모든 에이전트는 *상호작용* 요소를 가지고 있고, 학습에 필요한 데이터를 수집한다.
# 2. 모든 에이전트들은 현재 취하고 있는 행동을 *평가*하고,
# 3. 전체적인 성능을 개선하기 위해 만들어진 무언가를 *개선*한다.
#
# - *상태 영역state space*: 표현할 수 있는 변수들의 모든 값에 대한 조합
# - *관찰observation*: 에이전트가 어떤 특정 시간에 얻을 수 있는 변수들의 집합
# - *관찰 영역observation space*: 변수들이 가질 수 있는 모든 값들의 조합
# - *행동 영역action space*: 모든 상태에서 취할 수 있는 모든 행동에 대한 집합
# - *전이 함수transition function*: 환경은 에이전트의 행동에 대한 반응으로 상태를 변화할 수 있는데 이와 관련된 함수
# - *보상 함수reward function*: 행동과 관련된 보상에 대한 함수
# - *모델mpdel*: 전이와 보상 함수의 표현
# 보상 신호가 밀집되어 있을수록, 에이전트의 직관성과 학습속도가 높아지지만, 에이전트에게 편견을 주입하게 되어, 결국 에이전트가 예상하지 못한 행동을 할 가능성은 적어지게 된다. 반면, 보상 신호가 적을수록 직관성이 낮아져 에이전트가 새로운 행동을 취할 확률이 높아지지만, 그만큼 에이전트르르 학습시키는데 오래 걸리게 될 것이다.
#
# - *타임 스텝time step*: 상호작용이 진행되는 사이클, 시간의 단위
# - *경험 튜플experience tuple*: 관찰 또는 상태, 행동, 보상 그리고 새로운 관찰
# - *에피소드형 업무episodic task*: 게임과 같이 자연적으로 끝나는 업무 - *에피소드episode*
# - *연속형 업무continuing task*: 앞으로 전진하는 동작과 같이 자연적으로 끝나는 업무
# - *반환값return*: 에피소드 동안 수집된 보상의 총합
# - *상태state*: 문제에 포함되어 있는 독특하고, 자기 자신만의 설정이 담긴 요소
# - *상태 영역state square*: 모든 가능한 상태, 집합 S로 표현,
# **마르코프 결정 과정Markov decision process MDP**
# - 수학 프레임워크, 이를 이용해서 강화학습 환경을 연속적인 의사결정 문제로 표현하는 방법 학습.
# - 일반적인 형태는 불확실성상에 놓여진 상황에서 어떠한 복잡한 연속적 결정을 가상으로 모델링해서, 강화학습 에이전트가 모델과 반응을 주고받고, 경험을 통해서 스스로 학습할 수 있게 해준다.
# - 모든 상태들의 집합S
# - 모든 *시작 상태starting state* 혹은 *초기 상태initial state*라고 부르는 S+의 부분집합이 있음
# - MDP와의 상호작용 시작하면서 어떤 S에서의 특정 상태에서 어떤 확률 붙포 간의 관계를 그릴 수 있는데 이때 이 확률 분포는 무엇이든 될 수 있지만, 학습이 이뤄지는 동안에는 고정되어 있어야 한다.
# - 즉, 이 확률 분포에서 샘플링된 확률은 학습과 에이전트 검증의 처름 에피소드부터 마지막 에피소드까지는 항상 동일해야 한다는 것
# - *흡수absorbing* 또는 *종료 상태terminal state* 라는 특별한 상태도 존재.
# - 종료 상태(꼭 하나는 아님) 이외의 모든 상태들을 S라고 표현
# MDP에서는
# - **A**: 상태에 따라 결정되는 행동의 집합
# - $\therefore$ 특정 상태에서 허용되지 않는 행동도 존재한다는 말.
# - 상태(s)를 인자로 받는 함수.
# - A(s); 상태(s)에서 취할 수 있는 행동들의 집합 반환, 상수 정의 가능.
# - 행동 영역은 유한 혹은 무한 가능,
# - 단일 행동에 대한 변수들의 집합은 한 개 이상의 료소를 가질 수 있고, 유한해야 함.
# - $\therefore$ 대부분의 환경에서 모든 상태에서 취할 수 있는 행동의 수는 같도록 설계됨
# *상태-전이 확률state-transition probability* = *전이 함수*
# - $T(s,a,s')$
# - 전이함수$T$는 각 전이 튜플인 $(s,a,s')$을 확률과 연결시켜준다.
# - 어떤 상태 s에서 행동 a를 취하고 다음 상태가 s'이 되었을 때, 이때의 확률을 반환해준다는 뜻
# - 이 확률의 합은 1
# $$p(s'|s,a) = P(S_t = s'|S_{t-1} = s,A_{t-1}=a)$$
# $$\sum_{s' \in S}p(s'|s,a) = a, \forall s \in S, \forall a \in A(s)$$
# *보상 함수$R$*
# - 전이 튜플 $s,a,s'$을 특정한 스칼라 값으로 매핑,
# - 양수 보상 = 수익 또는 이득
# - 음수 보상 = 비용, 벌칙, 패널티
# - $R(s,a,s')$ = $R(s,a)$ = $R(s)$
# - *보상 함수를 표현하는 가장 명확한 방법은 상태와 행동 그리고 다음 상태, 이 세 가지를 함께 쓰는 것*
# $$r(s,a) = \mathbb{E} [R_t|S_{t-1} = s,A_{t-1} = a]$$
# $$r(s,a,s') = \mathbb{E} [R_t|S_{t-1} = s, A_{t-1} = a, S_t= s']$$
# $$R_t \in \cal{R} \subset \mathbb{R}$$
# *호라이즌horixon*
# - 계획 호라이즌planning horizon: 에피소드형 업무나 연속적 업무를 에이전트의 관점에서 정의
# - 유한 호라이즌finite horizon: 에이전트가 유한한 타임 스템 내에 업무가 종료된다는 것을 알고 있는 계획 호라이즌
# - 탐욕 호라이즌greedy horizon: 계획 호라이즌이 1인 경우
# - 무한 호라이즌infinite horiaon 에이전트한테 미리 정의된 타임 스텝에 대한 제한이 없어 에이전트가 무한하게 계획할 수 있음
# - 특히, 무한한 계획 호라이즌을 가지는 업무는 무기한 호라이즌 업무indefinite horizon task 라고 부름
# - 에이전트가 타임 스텝 루프에 빠지는 것을 막기 위해 타임 스텝을 제한하기도 한다.
# *감가율discount factor = 감마gamma*
# - 받았던 보상의 시점이 미래로 더 멀어질수록, 현재 시점에서는 이에 대한 가치를 더 낮게 평가
#
# - 환경; 자세한 예제; 5개의 미끄러지는 칸을 가지는 환경slippery walk five, SWF
# $$G_t = R_{t+1} + R_{t+2} + R_{t+3} + \dots R_T$$
# $$G_t = R_{t+1} + \gamma R_{t+2} + \gamma^2 R_{t+3} + \dots +\gamma^{T-1} R_T$$
# $$G_t = \sum^{\infty}_{k=0} \gamma^k R_{t+k+1}$$
# $$G_t = R_{t+1}+ \gamma G_{t+1}$$
# ### 3
# 에이전트의 목표: 반환값(보상의 총합)을 최대화할 수 있는 행동의 집합을 찾는 것, 정책이 필요하다.
# - *정책policy*: 가능한 모든 상태를 포괄한 전체적인 계획.
# - 확률적 혹은 결정적
# - 정책을 비교하기 위해 시작 상태를 포함한 모든 상태들에 대해 기대 반환값을 계산할 수 있어야 한다.
#
#
# - 정책$\pi$를 수행할 때, 상태 s에 대한 가치를 정의할 수 있다.
# - 에이전트가 정책 $\pi$를 따르면서 상태 s에서 시작했을 때, 상태 s의 가치는 반환값의 기대치라고 할 수 있다.
# - 가치함수는 상태 s에서 정책$\pi$를 따르면서 시작했을 때의 반환값에 대한 기대치를 나타낸다.
# $$v_{\pi} (s) = \mathbb{E}_{\pi} [G_t|S_t = s]$$
# $$v_{\pi} (s) = \mathbb{E}_{\pi}[R_{t+1} + \gamma R_{t+2} + \gamma^2 R_{+3} + \dots |S_t = s]$$
# $$v_{\pi} (s) = \sum_a \pi (a|s) \sum_{s',r} p(s',r|s,a)[r + \gamma v_\pi (s')] \forall s \in S$$
# *행동-가치 함수action-value function* = Q 함수 = $Q^{\pi}(s,a)$
# - 상태 s에서 행동 a를 취했을 때, 에이전트가 정책$\pi$를 수행하면서 얻을 수 있는 기대 반환값
# - MDP 없이도 정책을 개선시킬 수 있게 해준다.
# $$q_{\pi} (s,a) = \mathbb{E}_{\pi} [G_t|S_t = s,A_t = a]$$
# $$q_{\pi}(s,a) = \mathbb{E}_{\pi}[R_t + \gamma G_{t+1} | S_t = s, A_t = a]$$
# $$q_{\pi}(s,a) = \sum_{s',r} p(s',r|s,a)[r+\gamma v_{\pi} (s')], \forall s \in S, \forall a \in A(s)$$
# *행동-이점 함수action-advamtage function* = 이점함수advantage function = $A$
# - 상태 s에서 행동를 취했을 때의 가치와 정책 $\pi$에서 상태 s에 대한 상태-가치 함수 간의 차이
# $$a_{\pi}(s,a) = q_{\pi}(s,a) - v_{\pi}(s)$$
# 이상적인 정책optimal policy
# - 모든 상태에 대해서 다른 정책들보다 기대 반환값이 같거나 더 크게 얻을 수 있는 정책
# - 벨만 이상성 공식(아래)
# $$v_{*}(s) = max_{\pi} v_{\pi} (s), \forall_s \in S$$
# $$q_{*}(s,a) = max_{\pi} q_{\pi}(s,a), \forall s \in S, \forall a \in A(s)$$
# $$v_{*}(s) = max_{a} \sum_{s',r} p(s',r|s,a)[r+\gamma v_{*} (s')]$$
# $$q_{*}(s,a) = \sum_{s',r}p(s',r|s,a) [r + \gamma max_{a'} q_{*}(s',a')]$$
# *반복 정책 평가법iteractive policy evaluation* = *정책평가법policy rvaluation*
# - 임의의 정책을 평가할 수 있는 알고리즘
# - 상태 영역을 살펴보면서 반복적으로 에측치를 개선하는 방법 사용
# - 정책을 입력으로 받고, *예측문제prediction problem*를 풀 수 있는 알고리즘에 대한 가치 함수를 출력으로 내보내는 알고리즘, 이때는 미리 정의한 정책의 가치를 계산..(?)
# $$v_{k+1}(s) = \sum_a \pi(a|s) \sum_{s', r} p(s',r|s,a) \big[ r+\gamma v_k (s') \big]$$
# - 정책 평가 알고리즘을 충분히 반복하면 정책에 대한 가치함수로 수렴시킬 수 있다.
#
# - 실제로 적용하는 경우에는 우리가 근사하고자 하는 가치 함수의 변화를 확인하기 위해서 기분보다 작은 임계값을 사용.
# - 이런 경우에는 가치 함수의 변화가 우리가 정한 임계값보다 작을경우 반복을 멈추게 된다.
#
# - SWF환경에서 항상 왼쪽 행동을 취하는 정책에 이 알고리즘이 어떻게 동작할까?
# $$v_{k+1} (s) = \sum_a \pi(a|s) \sum_{s',r} p(s',r|s,a) \big[ r + \gamma v_k (s') \big] $$
# - $\gamma$는 1이라 가정,
# - 항상 왼쪽으로 가는 정책 사용
# - $k$: 반복적 정책-평가 알고리즘의 수행횟수
# $$ v^{\pi}_{1}(5) = p(s'=4|s=5, a=\text{왼쪽}) * [R(5,\text{왼쪽},4) + v^{\pi}_{0}(4)] +$$
# $$p(s'=5|s=5, a=\text{왼쪽}) * [R(5,\text{왼쪽},5) + v^{\pi}_{0}(5)] + $$
# $$p(s'=6|s=5, a=\text{왼쪽}) * [R(5,\text{왼쪽},6) + v^{\pi}_{0}(6)]$$
# $$c^{\pi}_{1} (5) = 0.50 * (0+0) + 0.33 * (0+0) + 0.166 * (1+0) = 0.166 \dots \text{ 1번 정책 평가법을 사용했을 떄 상태에 대한 가치를 나타냄}$$
# $\rightarrow$ Bootstraping 붓스트랩 방법: 예측치를 통해서 새로운 예측치를 계산하는 방법
# ```python
# # 정책(pi) 평가 알고리즘
# def policy_evaluation(pi,P, gamma = 1.0, theta = 1e-10):
# prev_V = np.zeros(len(P),dtype = np.float64)
# while True:
# V = np.aeros(len(P),dtype = float64)
# for s in range(len(P)):
# for prob, next_state, reward, done in P[s][pi(s)]:
# V[s] +- prob * (reward + gamma * prev_V[next_state] * (nor done))
# if np.max(np.abs(prev_V - V)) < theta:
# break
# prev_V = V.copy()
# return V
#
# ```
# > Note: 정책-개선 알고리즘이 동작하는 방법, 상태-가치 함수와 MDP를 사용해서 행동-가치 함수를 계산하고, 원래 정책의 행동-가치 함수에 대한 탐용 정책을 반환
# 정책개선 공식$$\pi ' (s) argmax_a \sum_{s',r} p(s',r|s,a) \big[ r + \gamma v_{\pi} (s') \big]$$
# ```python
# # 정책 개선(new_pi) 알고리즘
# def policy_improvement(V,P, gamma+1.0):
# Q = np.zeros((len{P}, len(P[0])), dtype = np.float64)
# for s in range(len(P)):
# for a in rnage(len(P[s])):
# for prob, next_state, reward, done in P[s][a]: # 여기서 done의 의미는 다음 상태가 종료 상태인지의 여부를 나타냄
# A[s][a] += prob * (reward + gamma * V[next_state] * nor done)
# new_pi = lambda s: {s: a for s, a in enumerate(np.argmax(np.argmax(Q, axis=1)))}[s]
# return new_pi
# ```
# *잃반화된 정책 순환generalized policy iteration, GPI* - 정책 순환, 가치 순환
# - 예측된 가치 함수를 사용해서 정책을 개선시키고, 예측된 가치 함수도 현대 정책의 실제 가치 함수를 바탕으로 개선시키는 강화학습의 일반적인 아이디어
# *정책 순환policy iteration*
# ```python
# # 정책 순환 알고리즘
# def policy _iteration (P, gamma = 1.0, theta = 1e-10):
# random_actions = np.random.choice(tuple(P[0].keys()),len(P))
# pi = lambda s: {s:a for s, a in enumerate(random_actions)}[s] # 임의 행동집합 만들고 행동에 대한 상태 매핑
# while True:
# old_pi = {s: pi(s) for s in range(len(P))} # 이전 정책에 대한 복사본 만들기
# V = policy_evaluation(pi, P, gamma, theta)
# pi = policy_improvement(V, P, gamma)
# if old_pi = {s:pi(s) for s in range(len(P))}: # 새로운 정책이 달라진 점이 있나? 있으면 앞의 과정 반복적 시행
# break
# return V, pi # 없다면 루프 중단시키고 이상적인 정책과 이상적인 상태-가치 함수 반환
# ```
# *가치 순환value iteration, VI*
# - 단일 반복 이후에 부분적인 정책 평가를 하더라도, 정책 평가시 한번 상태-영역을 훑은 이후에 대한 예측된 Q-함수에 탐용 정책을 취하면, 초기의 정책을 개선시킬 수 있다.
# 가치 순환 공식
# $$v_{k+1} (s) = max_a \sum_{s',r} p(s',r|s,a) \big[ r + \gamma v_k (s') \big]$$
# ```python
# # 가치 순환 알고리즘
# def value_iteration(P, gamma = 1.0, theta = 1e-10): # tehta는 수렴에 대한 임계값
# V = np.zeros(len(P),dtype = np.float64)
# while True:
# Q = np.zeros((len(P), len(P[0])), dtype = np.float64) # Q 함수가 0이어야 예측치가 정확해진다.
# for s in range(len(P)):
# for a in range(len(P[s])):
# for prob, next_state, reward, done in P[s][a]:
# Q[s][a] += prob * (reward + gamma * V[next_state] * (not done)) # 행동-가치 함수 계산
# if np.max(np.abs(V - np.max(Q, axis = 1))) < theta: # 상태-가치 함수가 변하지 않으면 이상적인 V함수를 찾은 것이고, 루프가 멈춤
# break
# V = np.max(Q, axis=1) # 개선과 평가 단계의 혼합
# pi = lambda s: {s:a for s, a in enumerate(np.argmax(Q, axis=1))}[s]
# ```
# ### 4
# *멀티 암드 밴딧Multi-armed bandit, MAB*
# - 상태 영역과 이에 따른 호라이즌이 1인 독특한 강화학습 문제
# - 여러 개의 선택지 중 하나를 선택하는 환경
# $$G_0 = 1* 0 + 0.99 * 0 0.9801 * 0 + 0.9702 * 0 + 0.9605 * 0.9509 * 1$$
# $$q(a) = \mathbb{E} \big[ R_t | A_t = a \big]$$
# - 행동 A에 대한 Q 함수는 A 가 샘플링 되었을 때의 기대 보상치를 나타낸다.
# $$v_* = q(a_*) = max_{a \in A} q(a)$$
# $$a_* = argmax_{a \in A} q(a)$$
# $$q(a_*) = v_*$$
# *전체 후회값total regret*
# - 전체 에피소드를 통해서 얻은 보상의 총합과 전체 보상치 간의 오차를 줄이면서 에피소드 당 기대 보상치를 최대화하는 것
# - 에피소드별로 이상적인 행동을 취했을 때의 실제 기대 보상치와 현재 정책이 선택한 행동을 취했을 때의 기대 보상치 간의 차이를 존부 더한다.
# - 낮을수록 더 좋은 성능
# $$T = \sum^{E}_{e = 1} \mathbb{E} \big[ v_* - q_* (A_e) \big]$$
# MAB 환경을 풀기 위한 방법들
# 1. 임의 탐색 전략random exploration strategy
# - 대부분 탐욕적으로 행동을 취하다가 입실론이라는 임계값의 확률로 행동을 임의로 선택
# 2. 낙관적인 탐색 전략optimistic exploration strategy
# - 의사 결정 문제에서 불확실성을 수치적으로 표현하고, 높은 불확실성 상에서도 상태에 대한 선호도를 높이는 조금 더 체계적인 방법
# 3. 정보 상태 영역 탐색 전략information state-space exploration strategy
# - 환경의 일부로써 에이전트의 정보 상태를 모델링, 상태 영역의 일부로 불확실성을 해석
# - 환경의 상태가 탐색되지 않은 미지의 경우와 이미 탐색된 경우일 때 다르게 보일 수 있다는 것을 의미
# - 상태 영역을 늘림으로써 복잡성이 증가하는 단점 존재
# *그리디 전략greedy strategy* = *순수 착취 전략pure exploitation strategy*
# - 탐욕적으로 행동을 취하는 전략은 항상 가장 높은 추정 가치를 얻는 행동을 취한다.
# ```python
# # 순수 착취 전략
# def pure_exploitation(enc, n_episodes=5000):
# Q = np.zeros((env.action_space.n),dtype=np.float64)
# N = np.zeros((env.action_space.n), dtype=np.int)
# Qe = np.empty((n_episodes, env.action_space.n),dtype=np.float64)
# returns = np.empty(n_episodes, dtype=np.float64)
# actions = np.empty(n_episodes, dtype=np.int)
# name = 'Pure exploitation'
# for e in tqdm(range(n_episodes), # 메인 루프로 들어가면서 환경과 상호작용이 일어나는 구간
# desc = 'Episodes for: ' + name,
# leave=False):
# action = np.argmax(Q) # Q 값을 최대화할 수 있는 행동 선택
# _, reward, _, _ = env.step(action) # 환경에 해당 행동 적용 후 새로운 보상
# B[action] += 1
# Q[action] = Q[action] + (reward - Q[action])/N[action]
# Qe[e] = Q
# returns[e] = reward
# actions[e] = action
# return name, returns, Qe, actions
# ```
# *임의 전략random strategy* = *순수 탐색 전략 pure exploration strategy*
# - 착취하지 않고 탐색만 하는 전략,
# - 에이전트의 유일한 목표는 정보를 얻는 것
# - 착취없이 행동을 결정하는 간단한 방식
# ```python
# # 순수 탄색 전략
# def pure_exploration(env, n_episodes=5000):
# Q = np.zeros((env.action_space.n),dtype=np.float64)
# N = np.zeros((env.action_space.n), dtype=np.int)
# Qe = np.empty((n_episodes, env.action_space.n),dtype=np.float64)
# returns = np.empty(n_episodes, dtype=np.float64)
# actions = np.empty(n_episodes, dtype=np.int)
# name = 'Pure exploration'
# for e in tqdm(range(n_episodes),
# desc = 'Episodes for: ' + name,
# leave=False):
# action=np.random.rsndit(len(Q))
# _, reward, _, _ = env.step(action) # 환경에 해당 행동 적용 후 새로운 보상
# B[action] += 1
# Q[action] = Q[action] + (reward - Q[action])/N[action]
# Qe[e] = Q
# returns[e] = reward
# actions[e] = action
# return name, returns, Qe, actions
# ```
# 착취는 목표이며, 탐색은 이 목표를 달성하기 위한 정보를 제공한다.
# **입실론 그리디 전략과 입실론 그리디 감가전략이 가장 많이 쓰이는 탐색 전략, 잘 동작하면서도 내부 구조가 단순하다는 이유로 선호받는다.**
# *입실론-그리디 전략 epsilon-greedy strategy*
# - 행동을 임의로 선택,
# ```python
# def epsilon_greedy(env,epsilon=0.01, n_episodes=5000):
# Q = np.zeros((env.action_space.n),dtype=np.float64)
# N = np.zeros((env.action_space.n), dtype=np.int)
# Qe = np.empty((n_episodes, env.action_space.n),dtype=np.float64)
# returns = np.empty(n_episodes, dtype=np.float64)
# actions = np.empty(n_episodes, dtype=np.int)
# name = 'Epsilon-Greedy {}'.format(epsilon)
# for e in tqdm(range(n_episodes),
# desc='Episodes for: ' + name,
# leave=False):
# if np.random.uniform() >epsilon: # 우선 임의로 숫자를 선택하고 이 값을 하이퍼파라미터인 epsilon과 비교
# action = np.argmax(Q)
# else:
# action = np.random.randit(len(Q))
# _, reward, _, _ = env.step(action) # 환경에 해당 행동 적용 후 새로운 보상
# B[action] += 1
# Q[action] = Q[action] + (reward - Q[action])/N[action]
# Qe[e] = Q
# returns[e] = reward
# actions[e] = action
# return name, returns, Qe, actions
#
# ```
# *입실론-그리디 감가 전략 decaying epsilon-greedy strategy*
# - 처음에는 입실론을 1보다 작거나 같은 매우 큰 값으로 시작하고, 매 타임 스텝마다 그 값을 감가시키는 것
# - 선형적으로 감가
# - 기하급수적으로 감가
# *낙관적 초기화 전략optimistic initialization* = 불확실성에 맞닿은 낙관성optimosm in the face of uncertainty
# - Q함수를 낙관적인 값인 높은 값으로 초기화함으로써 탐색되지 않은 행동에 대한 탐색을 할 수 있게 도와줘서
# - 에이전트는 환경과 상호작용 하면서 추정치는 낮은 값으로 수렴하기 시작하고
# - 정확해진 추정치는 에이전트가 실제로 높은 보상을 받을 수 있는 행동을 찾고 수렴할 수 있게 해준다.
# *소프트맥스 전략 softmax strategy*
# - 행동-가치 함수에 기반한 확률 분포로부터 행동을 샘플링하는데, 이 때 행동을 선택하는 확률은 행동-가치 추정에 비례하도록 한다.
# - Q 함수에 대한 선호도를 매기고, 이 선호도를 기반한 확률 분포에서 행동을 샘플링하면 된다.
# - Q값의 추정치 차이는 높은 추정치를 가지는 행동은 자주 선택하고, 낮은 추정치를 가지는 행동을 덜 선택하는 경향을 만든다.
# $$\pi(a) = \frac{exp\big( \frac{Q(a)}{\tau} \big) }{ \sum^{B}_{b=0} exp \big( \frac{Q(b)}{\tau} \big) }$$
# - $\tau$는 온도 계수,
# - $Q$값을 $\tau$로 나눠즈면 행동을 선택하는 선호도가 계산된다.
# *신뢰 상한 전략upper confidence bound. UCB*
# - 낙관적 초기화 원칙은 그대로 유지하되, 불확실성을 추정하는 값을 계산하는데 통계적인 기법을 사용하고, 이를 탐색할 떄 일종의 보너스로 사용하늗 것
# $$A_e = argmax_a \big[ Q_e(a) + c\sqrt{\frac{\ln e}{N_e(a)}} \big]$$
# *톰슨 샘플링 전략 thormpson sampling*
# - 탐색과 착위 사이에서 균형을 잡는데, 베이지안 기법을 사용한 샘플링 기반 확류탐색 전략
# - 책의 예시에서는 Q 값을 하나의 가우시안 분포로 고려하여 구현한다.
# - 평균을 기준으로 대칭을 이루고, 가르치는 목적에 적합할만큼 간단하기 때문
# - 다른 분포도 사용될 수 있다.
# - 가우시안 분포를 사용하는 이유
# - 가우시안 평균이 Q값의 추정치이고, 가우시안 표준편차는 추정치에 대한 불확실성을 나타내는데 이 값은 매 에피소드마다 업데이트 된다.
# - 하지만 베타 분포를 더 많이 쓰이는 것처럼 보인다.
# ### 5
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# ### 6
#
| _notebooks/2022-04-03-DRL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison of chlorophyll data from different sensors
# Different ocean color sensors have been launched since 1997 to provide continuous global ocean color data. Unfortunately, because of differences in sensor design and calibration, chlorophyll-a concentration values don’t match during their periods of overlap, making it challenging to study long-term trends.
#
# As an example, we are going to plot time-series of mean chlorophyll a concentration from various sensors from 1997 to 2019 to look at the periods of overlap.
# We are going to download data from Seawifs (1997-2010), MODIS (2002-2019) and VIIRS (2012-2019) and compare it to the ESA-CCI data (1997-2019) which combines all 3 sensors into a homogeneous time-series.
#
# First, let's load all the packages needed:
# +
import urllib.request
import xarray as xr
import netCDF4 as nc
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.colors import LinearSegmentedColormap
np.warnings.filterwarnings('ignore')
# -
# The OceanWatch website has a data catalog containing documentation and links to all the datasets available:
# https://oceanwatch.pifsc.noaa.gov/doc.html
#
# Navigate to the "Ocean Color" tab. From there you can access the different datasets using ERDDAP or THREDDS.
# ## Get monthly seawifs data, which starts in 1997
# Go to ERDDAP to find the name of the dataset for monthly SeaWIFS data: sw_chla_monthly_2018_0
#
# You should always examine the dataset in ERDDAP to check the date range, names of the variables and dataset ID, to make sure your griddap calls are correct: https://oceanwatch.pifsc.noaa.gov/erddap/griddap/sw_chla_monthly_2018_0.html
#
# Notice also that for this dataset and others, the latitudes are ordered from North to South, which will affect the construction of the download URL. (ie. instead of selecting latitudes 0-40N, you need to request 40-0).
#
# - let's download data for a box around the Hawaiian Islands:
url='https://oceanwatch.pifsc.noaa.gov/erddap/griddap/sw_chla_monthly_2018_0.nc?chlor_a[(1997-10-16T12:00:00Z):1:(2010-10-16T12:00:00Z)][(25):1:(15)][(198):1:(208)]'
url
urllib.request.urlretrieve(url, "sw.nc")
# - let's use xarray to extract the data from the downloaded file:
sw_ds = xr.open_dataset('sw.nc',decode_cf=False)
sw_ds.data_vars
sw_ds.chlor_a.shape
# The downloaded data contains only one variable: chlor_a.
#
# - let's compute the monthly mean over the region and extract the dates corresponding to each month of data:
# +
swAVG=np.mean(sw_ds.chlor_a,axis=(1,2))
swdates=nc.num2date(sw_ds.time,sw_ds.time.units)
# -
sw_ds.close()
# ## Get monthly MODIS data, which starts in 2002
url2='https://oceanwatch.pifsc.noaa.gov/erddap/griddap/aqua_chla_monthly_2018_0.nc?chlor_a[(2002-07-16T12:00:00Z):1:(2019-12-16T12:00:00Z)][(25):1:(15)][(198):1:(208)]'
urllib.request.urlretrieve(url2, "aq.nc")
# +
aq_ds = xr.open_dataset('aq.nc',decode_cf=False)
aqAVG=np.mean(aq_ds.chlor_a,axis=(1,2))
aqdates=nc.num2date(aq_ds.time,aq_ds.time.units)
# -
aq_ds.chlor_a.shape
aq_ds.close()
# ## Get monthly VIIRS data, which starts in 2012
url3='https://oceanwatch.pifsc.noaa.gov/erddap/griddap/noaa_snpp_chla_monthly.nc?chlor_a[(2012-01-02T12:00:00Z):1:(2019-12-01T12:00:00Z)][(25):1:(15)][(198):1:(208)]'
urllib.request.urlretrieve(url3, "snpp.nc")
# +
snpp_ds = xr.open_dataset('snpp.nc',decode_cf=False)
snppAVG=np.mean(snpp_ds.chlor_a,axis=(1,2))
snppdates=nc.num2date(snpp_ds.time,snpp_ds.time.units)
# -
snpp_ds.chlor_a.shape
snpp_ds.close()
# ## Get OC-CCI data (September 1997 to Dec 2019)
url4='https://oceanwatch.pifsc.noaa.gov/erddap/griddap/esa-cci-chla-monthly-v4-2.nc?chlor_a[(1997-09-04):1:(2019-12-01T00:00:00Z)][(25):1:(15)][(198):1:(208)]'
urllib.request.urlretrieve(url4, "cci.nc")
cci_ds = xr.open_dataset('cci.nc',decode_cf=False)
cciAVG=np.mean(cci_ds.chlor_a,axis=(1,2))
ccidates=nc.num2date(cci_ds.time,cci_ds.time.units)
cci_ds.close()
# ## Plot the data
plt.figure(figsize=(12,5))
plt.plot(swdates,swAVG,label='sw',c='red',marker='.',linestyle='-')
plt.plot(aqdates,aqAVG,label='aq',c='blue',marker='.',linestyle='-')
plt.plot(snppdates,snppAVG,label='snpp',c='green',marker='.',linestyle='-')
plt.ylabel('Chl-a (mg/m^3)')
plt.legend()
plt.figure(figsize=(12,5))
plt.plot(ccidates,cciAVG, label='cci',c='black')
plt.scatter(swdates,swAVG,label='sw',c='red')
plt.scatter(aqdates,aqAVG,label='aq',c='blue')
plt.scatter(snppdates,snppAVG,label='snpp',c='green')
plt.ylabel('Chl-a (mg/m^3)')
plt.legend()
| OW_tutorial2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import libraries
import os
import sys
import h5py
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#from keras.utils import HDF5Matrix
#from keras.utils import np_utils
# Modify notebook settings
# %matplotlib inline
# +
# Create a variable for the project root directory
proj_root = os.path.join(os.pardir)
# Save path to the raw metadata file
# "UrbanSound8K.csv"
metadata_file = os.path.join(proj_root,
"data",
"raw",
"UrbanSound8K",
"metadata",
"UrbanSound8K.csv")
# Save path to the raw audio files
raw_audio_path = os.path.join(proj_root,
"data",
"raw",
"UrbanSound8K",
"audio")
# Save path to the raw audio files
fold1_path = os.path.join(raw_audio_path,
"fold1")
# Save the path to the folder that will contain
# the interim data sets for modeling:
# /data/interim
interim_data_dir = os.path.join(proj_root,
"data",
"interim")
# Save the path to the folder that will contain
# the interim trash data sets
# /data/interim
interim_trash_dir = os.path.join(interim_data_dir,
"trash")
## Save path to 'sample-level.hdf5'
hdf5_file_name = 'sample-level.hdf5'
hdf5_path = os.path.join(interim_trash_dir, hdf5_file_name)
# -
# add the 'src' directory as one where we can import modules
src_dir = os.path.join(proj_root, "src")
sys.path.append(src_dir)
from utils.bedtime import computer_sleep
from models.resnext1d import ResNext1D
from models.dualplotcallback import DualPlotCallback
computer_sleep(seconds_until_sleep=12, verbose=1)
| notebooks/Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this **Python Examples** series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/90_Python_Examples)**
# </i></small></small>
# # Python Program to Multiply Two Matrices
#
# In this example, we will learn to multiply matrices using two different ways: nested loop and, nested list comprehension.
#
# To understand this example, you should have the knowledge of the following **[Python programming](https://github.com/milaan9/01_Python_Introduction/blob/main/000_Intro_to_Python.ipynb)** topics:
#
# * **[Python for Loop](https://github.com/milaan9/03_Python_Flow_Control/blob/main/005_Python_for_Loop.ipynb)**
# * **[Python List](https://github.com/milaan9/02_Python_Datatypes/blob/main/003_Python_List.ipynb)**
# * **[Python Matrices and NumPy Arrays](https://github.com/milaan9/90_Python_Examples/blob/main/02_Python_Datatypes_examples/Python_Matrices_and_NumPy_Arrays.ipynb)**
# In Python, we can implement a matrix as nested list (list inside a list).
#
# We can treat each element as a row of the matrix.
#
# For example **`X = [[1, 2], [4, 5], [3, 6]]`** would represent a 3x2 matrix.
#
# The first row can be selected as **`X[0]`**. And, the element in first row, first column can be selected as **`X[0][0]`**.
#
# Multiplication of two matrices **`X`** and **`Y`** is defined only if the number of columns in **`X`** is equal to the number of rows **`Y`**.
#
# If **`X`** is a **`n x m`** matrix and **`Y`** is a **`m x l`** matrix then, **`XY`** is defined and has the dimension **`n x l`** (but **`YX`** is not defined). Here are a couple of ways to implement matrix multiplication in Python.
# +
# Example 1: multiply two matrices using nested loops
# 3x3 matrix
X = [[12,9,3],
[4,5,6],
[7,8,3]]
# 3x4 matrix
Y = [[6,8,1,3],
[5,7,3,4],
[0,6,9,1]]
# result is 3x4
result = [[0,0,0,0],
[0,0,0,0],
[0,0,0,0]]
# iterate through rows of X
for i in range(len(X)):
# iterate through columns of Y
for j in range(len(Y[0])):
# iterate through rows of Y
for k in range(len(Y)):
result[i][j] += X[i][k] * Y[k][j]
for r in result:
print(r)
'''
>>Output/Runtime Test Cases:
[117, 177, 66, 75]
[49, 103, 73, 38]
[82, 130, 58, 56]
'''
# -
# **Explanation:**
#
# In this program we have used nested **`for`** loops to iterate through each row and each column. We accumulate the sum of products in the result.
#
# This technique is simple but computationally expensive as we increase the order of the matrix.
#
# For larger matrix operations we recommend optimized software packages like **[NumPy](http://www.numpy.org/)** which is several (in the order of 1000) times faster than the above code.
# +
# Example 2: multiply two matrices using list comprehension
# 3x3 matrix
X = [[12,9,3],
[4,5,6],
[7,8,3]]
# 3x4 matrix
Y = [[6,8,1,3],
[5,7,3,4],
[0,6,9,1]]
# result is 3x4
result = [[sum(a*b for a,b in zip(X_row,Y_col)) for Y_col in zip(*Y)] for X_row in X]
for r in result:
print(r)
'''
>>Output/Runtime Test Cases:
[117, 177, 66, 75]
[49, 103, 73, 38]
[82, 130, 58, 56]
'''
# -
# **Explanation:**
#
# The output of this program is the same as above. To understand the above code we must first know about **[built-in function zip()](https://github.com/milaan9/04_Python_Functions/blob/main/002_Python_Functions_Built_in/066_Python_zip%28%29.ipynb)** and **[unpacking argument list](http://docs.python.org/3/tutorial/controlflow.html#unpacking-argument-lists)** using **`*`** operator.
#
# We have used nested list comprehension to iterate through each element in the matrix. The code looks complicated and unreadable at first. But once you get the hang of list comprehensions, you will probably not go back to nested loops.
| 02_Python_Datatypes_examples/003_multiply_two_matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Review Labels CSVs
# Amy created Labels CSVs with just 3 classes for Train, Validation, and Test
#
# These also only consider a label to be part of a crop if it's in the ROI (at least 10 pixels away from the edge of the crop)
# +
import numpy as np
import pandas as pd
import os
from datetime import datetime
import random
# -
# ## Examine train
df_train = pd.read_csv('train_labels.csv')
print(df_train.shape)
print(df_train['ground_truth'].value_counts())
df_train.head()
# ## Examine validation
df_val = pd.read_csv('validation_labels.csv')
print(df_val.shape)
print(df_val['ground_truth'].value_counts())
df_val.head()
df_val_missing = df_val.loc[df_val['0_missing'] > 0]
df_val_missing
df_val_missing['img_id'].value_counts()
# ## Examine test
df_test = pd.read_csv('test_labels.csv')
print(df_test.shape)
print(df_test['ground_truth'].value_counts())
df_test.head()
df_test_missing = df_test.loc[df_test['0_missing'] > 0]
df_test_missing
df_test_missing['img_id'].value_counts()
# # Comments
# * Missing Curb Ramps are extremely infrequent
| label-crops/2020-04-03-ReviewLabelsCSVs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Case Study: Predicting salmon returns
#
# This case study is based on a ModSim student project by <NAME> and <NAME>.
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### Can we predict salmon populations?
#
# Each year the [U.S. Atlantic Salmon Assessment Committee](https://www.nefsc.noaa.gov/USASAC/Reports/USASAC2018-Report-30-2017-Activities.pdf) reports estimates of salmon populations in oceans and rivers in the northeastern United States. The reports are useful for monitoring changes in these populations, but they generally do not include predictions.
#
# The goal of this case study is to model year-to-year changes in population, evaluate how predictable these changes are, and estimate the probability that a particular population will increase or decrease in the next 10 years.
#
# As an example, I'll use data from page 18 of the 2017 report, which provides population estimates for the Narraguagus and Sheepscot Rivers in Maine.
#
# 
#
# At the end of this notebook, I make some suggestions for extracting data from a PDF document automatically, but for this example I will keep it simple and type it in.
#
# Here are the population estimates for the Narraguagus River:
pops = [2749, 2845, 4247, 1843, 2562, 1774, 1201, 1284, 1287, 2339, 1177, 962, 1176, 2149, 1404, 969, 1237, 1615, 1201];
# To get this data into a Pandas Series, I'll also make a range of years to use as an index.
years = range(1997, 2016)
# And here's the series.
pop_series = TimeSeries(pops, index=years, dtype=float)
# Here's what it looks like:
# +
def plot_population(series):
plot(series, label='Estimated population')
decorate(xlabel='Year',
ylabel='Population estimate',
title='Narraguacus River',
ylim=[0, 5000])
plot_population(pop_series)
# -
# ## Modeling changes
#
# To see how the population changes from year-to-year, I'll use `diff` to compute the absolute difference between each year and the next.
#
# `shift` adjusts the result so each change aligns with the year it happened.
abs_diffs = np.ediff1d(pop_series, np.nan)
# We can compute relative differences by dividing by the original series elementwise.
rel_diffs = abs_diffs / pop_series
# Or we can use the `modsim` function `compute_rel_diff`:
rel_diffs = compute_rel_diff(pop_series)
# These relative differences are observed annual net growth rates. So let's drop the `NaN` and save them.
rates = rel_diffs.dropna()
# A simple way to model this system is to draw a random value from this series of observed rates each year. We can use the NumPy function `choice` to make a random choice from a series.
np.random.choice(rates)
# ## Simulation
#
# Now we can simulate the system by drawing random growth rates from the series of observed rates.
#
# I'll start the simulation in 2015.
t_0 = 2015
p_0 = pop_series[t_0]
# Create a `System` object with variables `t_0`, `p_0`, `rates`, and `duration=10` years.
#
# The series of observed rates is one big parameter of the model.
system = System(t_0=t_0,
p_0=p_0,
duration=10,
rates=rates)
# Write an update functon that takes as parameters `pop`, `t`, and `system`.
# It should choose a random growth rate, compute the change in population, and return the new population.
# +
# Solution goes here
# -
# Test your update function and run it a few times
update_func1(p_0, t_0, system)
# Here's a version of `run_simulation` that stores the results in a `TimeSeries` and returns it.
def run_simulation(system, update_func):
"""Simulate a queueing system.
system: System object
update_func: function object
"""
t_0 = system.t_0
t_end = t_0 + system.duration
results = TimeSeries()
results[t_0] = system.p_0
for t in linrange(t_0, t_end):
results[t+1] = update_func(results[t], t, system)
return results
# Use `run_simulation` to run generate a prediction for the next 10 years.
#
# The plot your prediction along with the original data. Your prediction should pick up where the data leave off.
# +
# Solution goes here
# -
# To get a sense of how much the results vary, we can run the model several times and plot all of the results.
def plot_many_simulations(system, update_func, iters):
"""Runs simulations and plots the results.
system: System object
update_func: function object
iters: number of simulations to run
"""
for i in range(iters):
results = run_simulation(system, update_func)
plot(results, color='gray', linewidth=5, alpha=0.1)
# The plot option `alpha=0.1` makes the lines semi-transparent, so they are darker where they overlap.
#
# Run `plot_many_simulations` with your update function and `iters=30`. Also plot the original data.
# +
# Solution goes here
# -
# The results are highly variable: according to this model, the population might continue to decline over the next 10 years, or it might recover and grow rapidly!
#
# It's hard to say how seriously we should take this model. There are many factors that influence salmon populations that are not included in the model. For example, if the population starts to grow quickly, it might be limited by resource limits, predators, or fishing. If the population starts to fall, humans might restrict fishing and stock the river with farmed fish.
#
# So these results should probably not be considered useful predictions. However, there might be something useful we can do, which is to estimate the probability that the population will increase or decrease in the next 10 years.
# ## Distribution of net changes
#
# To describe the distribution of net changes, write a function called `run_many_simulations` that runs many simulations, saves the final populations in a `ModSimSeries`, and returns the `ModSimSeries`.
#
def run_many_simulations(system, update_func, iters):
"""Runs simulations and report final populations.
system: System object
update_func: function object
iters: number of simulations to run
returns: series of final populations
"""
# FILL THIS IN
# +
# Solution goes here
# -
# Test your function by running it with `iters=5`.
run_many_simulations(system, update_func1, 5)
# Now we can run 1000 simulations and describe the distribution of the results.
last_pops = run_many_simulations(system, update_func1, 1000)
last_pops.describe()
# If we substract off the initial population, we get the distribution of changes.
net_changes = last_pops - p_0
net_changes.describe()
# The median is negative, which indicates that the population decreases more often than it increases.
#
# We can be more specific by counting the number of runs where `net_changes` is positive.
np.sum(net_changes > 0)
# Or we can use `mean` to compute the fraction of runs where `net_changes` is positive.
np.mean(net_changes > 0)
# And here's the fraction where it's negative.
np.mean(net_changes < 0)
# So, based on observed past changes, this model predicts that the population is more likely to decrease than increase over the next 10 years, by about 2:1.
# ## A refined model
#
# There are a few ways we could improve the model.
#
# 1. It looks like there might be cyclic behavior in the past data, with a period of 4-5 years. We could extend the model to include this effect.
#
# 2. Older data might not be as relevant for prediction as newer data, so we could give more weight to newer data.
#
# The second option is easier to implement, so let's try it.
#
# I'll use `linspace` to create an array of "weights" for the observed rates. The probability that I choose each rate will be proportional to these weights.
#
# The weights have to add up to 1, so I divide through by the total.
weights = linspace(0, 1, len(rates))
weights /= sum(weights)
plot(weights)
decorate(xlabel='Index into the rates array',
ylabel='Weight')
# I'll add the weights to the `System` object, since they are parameters of the model.
system.weights = weights
# We can pass these weights as a parameter to `np.random.choice` (see the [documentation](https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.choice.html))
np.random.choice(system.rates, p=system.weights)
# Write an update function that takes the weights into account.
# +
# Solution goes here
# -
# Use `plot_many_simulations` to plot the results.
# +
# Solution goes here
# -
# Use `run_many_simulations` to collect the results and `describe` to summarize the distribution of net changes.
# +
# Solution goes here
# -
# Does the refined model have much effect on the probability of population decline?
# +
# Solution goes here
# -
# ## Extracting data from a PDF document
#
# The following section uses PyPDF2 to get data from a PDF document. It uses features we have not seen yet, so don't worry if it doesn't all make sense.
#
# The PyPDF2 package provides functions to read PDF documents and get the data.
#
# If you don't already have it installed, and you are using Anaconda, you can install it by running the following command in a Terminal or Git Bash:
#
# ```
# conda install -c conda-forge pypdf2
# ```
import PyPDF2
# The 2017 report is in the data directory.
pdfFileObj = open('data/USASAC2018-Report-30-2017-Activities-Page11.pdf', 'rb')
# The `PdfFileReader` object knows how to read PDF documents.
pdfReader = PyPDF2.PdfFileReader(pdfFileObj)
# This file contains only one page.
pdfReader.numPages
# `getPage` selects the only page in the document.
page = pdfReader.getPage(0)
page.extractText()
# The following function iterates through the lines on the page, removes whitespace, and ignores lines that contain only whitespace.
def iter_page(page):
for item in page.extractText().splitlines():
item = item.strip()
if item:
yield item
# The following function gets the next `n` pages from the page.
def next_n(iterable, n):
"""Get the next n items from an iterable."""
return [next(iterable) for i in range(n)]
# We skip the text at the top of the page.
t = iter_page(page)
discard = next_n(t, 8)
# The next 7 strings are the column headings of the table.
columns = next_n(t, 7)
# Create an empty `Dataframe` with the column headings.
df = pd.DataFrame(columns=columns)
df
# Get the next 19 lines of the table.
for i in range(19):
year = int(next(t))
data = next_n(t, 7)
df.loc[year] = data
# The last line in the table gets messed up, so I'll do that one by hand.
df.loc[2017] = ['363', '663', '13', '2', '1041', '806', '235']
# Here's the result.
df
# In general, reading tables from PDF documents is fragile and error-prone. Sometimes it is easier to just type it in.
| code/salmon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="FevjB4sgsITg"
#
# # **Notebook for EMG data visualization collected from Galea**
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 233, "status": "ok", "timestamp": 1631733029866, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="l6qTWG2Tc8sO" outputId="fe09aa11-9a14-43d5-b51a-125bda863e64"
#Mounts your google drive into this virtual machine
from google.colab import drive
drive.mount('/content/drive')
# + id="zYFO_Ha3ZyIW"
#Now we need to access the files downloaded, copy the path where you saved the files downloaded from the github repo and replace the path below
# %cd /content/drive/MyDrive/path/to/files/cloned/from/repo/and/now/in/your/GoogleDrive/
# + id="G7WRW_OQdQUR"
# !pip install neurokit2
# !pip install mne
# !pip install -U pandas-profiling
# + id="uaUD4dRDZnCY"
import time
import numpy as np
import pandas as pd
import matplotlib
import neurokit2 as nk
import mne
import matplotlib.pyplot as plt
import os
import random
#from pylsl import StreamInfo, StreamOutlet, resolve_stream, StreamInlet
from sklearn.cross_decomposition import CCA
from scipy import signal
from scipy.signal import butter, lfilter
from scipy.fft import fft, fftfreq, ifft
import pickle
# %matplotlib inline
plt.rcParams['figure.figsize'] = [30, 15]
# + [markdown] id="547CRw1mckKH"
# ## **Offline EMG data visualization and processing**
# + colab={"base_uri": "https://localhost:8080/", "height": 643} executionInfo={"elapsed": 3692, "status": "ok", "timestamp": 1631731876794, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="J7irTzpAca0G" outputId="12d9d72a-db43-4ab8-acc8-ac4943fb7ecc"
#Replace the path below, so we can load the data
data = pd.read_csv('/content/drive/MyDrive/YOURPATH/SharedPublicly/Data/EMG_RAW-2021-08-07_10-02-37.txt',header=4 ,sep=',')
data.columns = ["Sample Index", "EMG Channel 0", "EMG Channel 1", "EMG Channel 2", "EMG Channel 3", "EOG Channel 0", "EOG Channel 1", "EEG Channel 0", "EEG Channel 1", "EEG Channel 2", "EEG Channel 3", "EEG Channel 4", "EEG Channel 5", "EEG Channel 6", "EEG Channel 7", "EEG Channel 8", "EEG Channel 9", "PPG Channel 0", "PPG Channel 1", "EDA_Channel_0", "Other", "Raw PC Timestamp", "Raw Device Timestamp", "Other.1", "Timestamp", "Marker", "Timestamp (Formatted)"]
data
# + colab={"base_uri": "https://localhost:8080/", "height": 643} executionInfo={"elapsed": 15, "status": "ok", "timestamp": 1631731876795, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="NlwJuMrVH6id" outputId="468655bf-7790-4b02-886a-f7eb0ef049ec"
#Let's grab a section of data for clarity
dt1 =data[1800:]
dt1
# + colab={"base_uri": "https://localhost:8080/", "height": 765} executionInfo={"elapsed": 2707, "status": "ok", "timestamp": 1631731896561, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="YE1lro228Bd-" outputId="36e88e77-d5bb-4284-b9de-e76ba0edb7c1"
dt1 =data[1800:]
emg_signal =dt1["EMG Channel 0"]
emg =nk.as_vector(emg_signal)
emg = emg - np.mean(emg)
emg = nk.signal_detrend(emg, method='polynomial', order=1, regularization=500, alpha=0.75, window=1.5, stepsize=0.02)
emg_signal, info = nk.emg_process(emg, sampling_rate=250)
nk.signal_plot(emg_signal.EMG_Clean)
# + colab={"base_uri": "https://localhost:8080/", "height": 762} executionInfo={"elapsed": 1995, "status": "ok", "timestamp": 1631731901591, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="wHGMBO_Hmsze" outputId="e774e60d-e002-4e3f-8828-940a65fc4552"
nk.signal_plot(emg_signal)
# + id="PDMqYWRa1fEn"
#cleaned = nk.emg_clean(emg_signal, sampling_rate=250)
nk.emg_plot(emg_signal, sampling_rate=250)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 240, "status": "ok", "timestamp": 1631732356964, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="dbkXB0chwf95" outputId="92b6233c-604a-4e6d-a145-80bc42e68c1f"
dt2 =data[4800:7000]
dt2 =dt2["EMG Channel 1"]
emg_1 =nk.as_vector(dt2)
emg_1
# + colab={"base_uri": "https://localhost:8080/", "height": 419} executionInfo={"elapsed": 284, "status": "ok", "timestamp": 1631732430810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="gywfqwOq_NuI" outputId="2bb3c90e-3e99-4623-d3b7-fa7f524c6aea"
emg_1 = emg_1 - np.mean(emg_1)
emg_1 = nk.stats.rescale(emg_1,to=[-150, 150])
# emg_1 = nk.signal_detrend(emg_1, method='polynomial', order=1, regularization=500, alpha=0.75, window=1.5, stepsize=0.02)
emg_signal_1, info = nk.emg_process(emg_1, sampling_rate=250)
emg_signal_1
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 2331, "status": "ok", "timestamp": 1631732855761, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="YaXIDMZq_Run" outputId="40e58c63-e805-44b7-986e-9ba7115a89a5"
nk.signal_plot(emg_signal_1.EMG_Clean)
#nk.emg_plot(signals_1, sampling_rate=250)
emg_signal_1.EMG_Clean = nk.stats.rescale(emg_signal_1.EMG_Clean,to=[-150, 150])
emg_signal_1.EMG_Amplitude = nk.stats.rescale(emg_signal_1.EMG_Amplitude,to=[-10, 10])
nk.signal_plot(emg_signal_1)
# + colab={"base_uri": "https://localhost:8080/", "height": 574} executionInfo={"elapsed": 1329, "status": "ok", "timestamp": 1631733403773, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09042963316942946918"}, "user_tz": 420} id="LNTfgQoxAWbT" outputId="50c776f6-4e54-471a-e509-4da52effb1c5"
plt.rcParams['figure.figsize'] = [10, 5]
image_format = 'eps' # e.g .png, .svg, etc.
image_name = 'galea_emg.eps'
fig = nk.emg_plot(emg_signal_1, sampling_rate=250)
fig.savefig(image_name, format=image_format, dpi=1200)
# -
# ### Signal Validation Procedure:
# Signal quality was compared to [4] based on algorithm available at https://www.mathworks.com/matlabcentral/fileexchange/61830-emg_signaltonoiseratio on Matlab, and the .mat EMG data that was passed through this algorithm is available in \Data\EMG_trimmed.mat
#
# [4] <NAME> and <NAME>. An Algorithm for the Estimation of the Signal-To-Noise Ratio in Surface Myoelectric Signals Generated During Cyclic Movements. IEEE Transactions on Biomedical Engineering, 59(1):219–225, Jan. 2012. Conference Name: IEEE Transactions on Biomedical Engineering. doi: 10.1109/TBME.2011.2170687
| Notebooks/EMG_dataViz_09_02_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # FMR standard problem
#
# ## Problem specification
#
# We choose a cuboidal thin film permalloy sample measuring $120 \times 120 \times 10 \,\text{nm}^{3}$. The choice of a cuboid is important as it ensures that the finite difference method employed by OOMMF does not introduce errors due to irregular boundaries that cannot be discretized well. We choose the thin film geometry to be thin enough so that the variation of magnetization dynamics along the out-of-film direction can be neglected. Material parameters based on permalloy are:
#
# - exchange energy constant $A = 1.3 \times 10^{-11} \,\text{J/m}$,
# - magnetisation saturation $M_\text{s} = 8 \times 10^{5} \,\text{A/m}$,
# - Gilbert damping $\alpha = 0.008$.
#
# An external magnetic bias field with magnitude $80 \,\text{kA/m}$ is applied along the direction $e = (1, 0.715, 0)$. We choose the external magnetic field direction slightly off the sample diagonal in order to break the system’s symmetry and thus avoid degenerate eigenmodes. First, we initialize the system with a uniform out-of-plane magnetization $m_{0} = (0, 0, 1)$. The system is allowed to relax for $5 \,\text{ns}$, which was found to be sufficient time to obtain a well-converged equilibrium magnetization configuration. We refer to this stage of simulation as the relaxation stage, and its final relaxed magnetization configuration is saved to serve as the initial configuration for the next dynamic stage. Because we want to use a well defined method that is supported by all simulation tools, we minimize the system’s energy by integrating the LLG equation with a large, quasistatic Gilbert damping $\alpha = 1$ for $5 \,\text{ns}$. In the next step (dynamic stage), a simulation is started using the equilibrium magnetisation configuration from the relaxation stage as the initial configuration. Now, the direction of an external magnetic field is altered to $e = (1, 0.7, 0)$. This simulation stage runs for $T = 20 \,\text{ns}$ while the (average and spatially resolved) magnetization $M(t)$ is recorded every $\Delta t = 5 \,\text{ps}$. The Gilbert damping in this dynamic simulation stage is $\alpha = 0.008$.
#
# Details of this standard problem specification can be found in Ref. 1.
#
# ## Relaxation stage
#
# Firstly, all required modules are imported.
import oommfc as oc
import discretisedfield as df
import micromagneticmodel as mm
# Now, we specify all simulation parameters.
# +
import numpy as np
lx = ly = 120e-9 # x and y dimensions of the sample(m)
lz = 10e-9 # sample thickness (m)
dx = dy = dz = 5e-9 # discretisation in x, y, and z directions (m)
Ms = 8e5 # saturation magnetisation (A/m)
A = 1.3e-11 # exchange energy constant (J/m)
H = 8e4 * np.array([0.81345856316858023, 0.58162287266553481, 0.0])
alpha = 0.008 # Gilbert damping
gamma0 = 2.211e5
# -
# Now, the system object can be created and mesh, magnetisation, hamiltonian, and dynamics are specified.
# +
mesh = df.Mesh(p1=(0, 0, 0), p2=(lx, ly, lz), cell=(dx, dy, dz))
system = mm.System(name='stdprobfmr')
system.energy = mm.Exchange(A=A) + mm.Demag() + mm.Zeeman(H=H)
system.dynamics = mm.Precession(gamma0=gamma0) + mm.Damping(alpha=alpha)
system.m = df.Field(mesh, dim=3, value=(0, 0, 1), norm=Ms)
# -
# Finally, the system is relaxed.
md = oc.MinDriver()
md.drive(system)
# We can now load the relaxed state to the Field object and plot the $z$ slice of magnetisation.
system.m.plane('z', n=(10, 10)).mpl()
# ## Dynamic stage
# In the dynamic stage, we use the relaxed state from the relaxation stage.
# Change external magnetic field.
H = 8e4 * np.array([0.81923192051904048, 0.57346234436332832, 0.0])
system.energy.zeeman.H = H
# Finally, we run the multiple stage simulation using `TimeDriver`.
# +
T = 20e-9
n = 4000
td = oc.TimeDriver()
td.drive(system, t=T, n=n)
# -
# ## Postprocessing
# From the obtained vector field samples, we can compute the average of magnetisation $y$ component and plot its time evolution.
# +
import matplotlib.pyplot as plt
t = system.table.data['t'].values
my = system.table.data['mx'].values
# Plot <my> time evolution.
plt.figure(figsize=(8, 6))
plt.plot(t, my)
plt.xlabel('t (ns)')
plt.ylabel('my average')
plt.grid()
# -
# From the $<m_{y}>$ time evolution, we can compute and plot its Fourier transform.
# +
import scipy.fftpack
psd = np.log10(np.abs(scipy.fftpack.fft(my))**2)
f_axis = scipy.fftpack.fftfreq(4000, d=20e-9/4000)
plt.plot(f_axis/1e9, psd)
plt.xlim([6, 12])
plt.xlabel('f (GHz)')
plt.ylabel('Psa (a.u.)')
plt.grid()
# -
# ## References
#
# [1] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, *J. Magn. Magn. Mater.* **421**, 428 (2017).
| examples/10-tutorial-standard-problem-fmr.ipynb |
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% # <center>Interpoláció</center>
% ## <center>Lagrange-féle és Newton interpolációs polinom</center>
% <br>
% <b>Példa.</b> Tekintsük az alábbi
%
% $$(t_i, f_i):\ (−3, −209),\ (−2, −43),\ (−1, −1),\ (1, −1),\ (2, −19)$$
%
% adatokat és illesszünk ezekre egy megfelelő fokú Lagrange-féle interpolációs polinomot!
%
% +
t = [-3 -2 -1 1 2];
f = [-209 -43 -1 -1 -19];
% Beepitett fuggveny hasznalata illeszteshez
p = polyfit(t,f,length(t)-1);
tt = linspace(t(1),t(end),100);
ff = polyval(p,tt);
plot(t,f, 'm*',tt,ff)
title('Az illesztett interpolacios polinom')
% -
% <br>
% <b>Runge példája</b> Tekintsük az
%
% $$f(x)=\frac{1}{1+25x^2},\ \quad x\in [-1,1]$$
%
% függvényt. Ábrázoljuk egy ábrán ekvidisztáns felosztás mellett az $L_6(x), L_{12}(x), L_{18}(x)$ Lagrange interpolációs polinomokat, az eredeti függvényt és a Csebisev alappontokra támaszkodó interpoláló polinomot!
Rungepelda(-1,1,6)
% ## <center>Spline interpolációs polinom</center>
%
%
% Az előző példát elegánsan harmadfokú spline interpolációval is meg lehet oldani
x = linspace(-1,1,100);
y = 1./(1+25*x.^2);
yspline = spline (x,y,x);
plot(x,y,'b',x,yspline,'mo');
legend('f(x)','Spline interpolacios polinom')
% <br>
% Alkalmazás:
%
% + <a href="https://www.youtube.com/watch?v=hHb7X_eriVk" target="_blank">Mitsubish CNC marás</a>
| Eloadas/Blokk#4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Atlas2
#
#
# Write first 20 graphs from the graph atlas as graphviz dot files
# Gn.dot where n=0,19.
#
# +
# Author: <NAME> (<EMAIL>)
# Date: 2005-05-19 14:23:02 -0600 (Thu, 19 May 2005)
# Copyright (C) 2006-2019 by
# <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# All rights reserved.
# BSD license.
import networkx as nx
from networkx.generators.atlas import graph_atlas_g
atlas = graph_atlas_g()[0:20]
for G in atlas:
print("graph %s has %d nodes with %d edges"
% (G.name, nx.number_of_nodes(G), nx.number_of_edges(G)))
A = nx.nx_agraph.to_agraph(G)
A.graph_attr['label'] = G.name
# set default node attributes
A.node_attr['color'] = 'red'
A.node_attr['style'] = 'filled'
A.node_attr['shape'] = 'circle'
A.write(G.name + '.dot')
| NoSQL/NetworkX/dot_atlas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/skywalker0803r/c620/blob/main/notebook/c670_transferlearning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="nOGsCEawJNva"
import pandas as pd
import numpy as np
import joblib
# !pip install autorch > log.txt
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="7UXYDDgMJWvA" outputId="1ecb2df6-c65a-4d0c-95f3-d4268f98ef10"
c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c670_col_names.pkl')
c670_df = pd.read_excel('/content/drive/MyDrive/台塑輕油案子/data/c620/明志_遷移式學習_訓練資料_寄送版/c670_data.xlsx',index_col=0)
print(c670_df.shape)
c670_df.head(3)
# + [markdown] id="weDth9iWKG8e"
# # 缺失欄位
# + colab={"base_uri": "https://localhost:8080/"} id="_p-HyDHLJq1g" outputId="3a6f8754-c3cb-407f-d40a-a30e743627c9"
miss_col = c670_df.columns[(c670_df.isnull().sum() > 0).values].tolist()
print(len(miss_col))
miss_col
# + [markdown] id="wMJG3E0fKNc-"
# # 有提供欄位
# + colab={"base_uri": "https://localhost:8080/"} id="OB9E7WrhJyoM" outputId="4ddb5f87-040d-4881-8c43-16dea87e5782"
have_col = c670_df.columns[(c670_df.isnull().sum() == 0).values].tolist()
print(len(have_col))
have_col
# + [markdown] id="sM7xuEbI9bD8"
# # 補足缺失欄位 分離係數
# + id="4oWanpDn9bLK" colab={"base_uri": "https://localhost:8080/", "height": 304} outputId="12f368d5-4d6a-4d09-b074-a45d52f70f12"
path = '/content/drive/MyDrive/台塑輕油案子/data/c620/明志_遷移式學習_訓練資料_寄送版/蒸餾塔(C620_C660_C670)取出品管資料_寄送明志科大 r2.xlsx'
df2 = pd.read_excel(path,sheet_name='資料彙整(寄送明志)r2')
df2.head()
# + id="J_tIwW5M9i-3" colab={"base_uri": "https://localhost:8080/", "height": 457} outputId="10780070-b711-4755-b6f1-af2af5143970"
c670_wt2,c670_fout2 = df2.iloc[353:353+41,2:].T,df2.iloc[[353+41],2:].T
c670_wt4,c670_fout4 = df2.iloc[397:397+41,2:].T,df2.iloc[[397+41],2:].T
c670_feed_wt,c670_feed_flow = df2.iloc[529:529+41,2:].T,df2.iloc[[529+41],2:].T
c670_wt = c670_wt2.join(c670_wt4)
c670_wt.index = c670_df.index
c670_wt.columns = c['distillate_x'] + c['bottoms_x']
s2 = np.clip((c670_wt2.values*c670_fout2.values)/(c670_feed_wt.values*c670_feed_flow.values+1e-8),0,1)
s4 = np.clip((c670_wt4.values*c670_fout4.values)/(c670_feed_wt.values*c670_feed_flow.values+1e-8),0,1)
s2_col = c670_df.filter(regex='Split Factor for Individual Component to Toluene Column C670 Distillate').columns.tolist()
s4_col = c670_df.filter(regex='Split Factor for Individual Component to Toluene Column C670 Bottoms').columns.tolist()
c670_df[s2_col] = s2
c670_df[s4_col] = s4
c670_wt_always_same_split_factor_dict = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c670_wt_always_same_split_factor_dict.pkl')
for i in c670_wt_always_same_split_factor_dict.keys():
c670_df[i] = c670_wt_always_same_split_factor_dict[i]
c670_df.update(c670_wt)
c670_df.to_excel('/content/drive/MyDrive/台塑輕油案子/data/c620/明志_遷移式學習_訓練資料_寄送版/c670_data.xlsx')
c670_df.iloc[:,-41*2:].head()
# + [markdown] id="5-E7rEFA_G3w"
# # 定義欄位
# + id="IaMoBXdSKQ83"
x_col = c['combined'] + c['upper_bf']
op_col = c['density']+c['yRefluxRate']+c['yHeatDuty']+c['yControl']
sp_col = s2_col + s4_col
y_col = c670_df[sp_col+op_col].dropna(axis=1).columns.tolist()
n_idx = [ [i,i+41] for i in range(41)]
# + [markdown] id="a18T2VrXRvtx"
# # 1. 實驗直接訓練 (不使用預訓練模型)
# + colab={"base_uri": "https://localhost:8080/", "height": 401} id="s6icTkbYQiJF" outputId="27acbc4a-6bee-415f-82a8-5e94a49e31d8"
from autorch.utils import PartBulider
c670 = PartBulider(c670_df,x_col,y_col,normalize_idx_list=n_idx,limit_y_range=True)
c670.train()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="BzLHL_YsQkMC" outputId="42101022-b85a-4190-cbec-7804d710fd33"
c670.test(e=2e-2)
# + id="1by_ll5tE8YZ" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="8edbfaa0-3960-4537-a3d5-3c0379db6a87"
from autorch.function import sp2wt
from sklearn.metrics import r2_score,mean_squared_error
c670_sp,c670_op = c670.predict(c670.data['X_test']).iloc[:,:41*2],c670.predict(c670.data['X_test']).iloc[:,41*2:]
c670_feed = c670.data['X_test'][c['combined']].values
s2,s4 = c670_sp.iloc[:,0:41].values,c670_sp.iloc[:,41:41*2].values
w2,w4 = sp2wt(c670_feed,s2),sp2wt(c670_feed,s4)
wt = np.hstack((w2,w4))
c670_wt = pd.DataFrame(wt,index=c670.data['X_test'].index,columns=c['distillate_x']+c['bottoms_x'])
c670_wt_real = c670_df.loc[c670_wt.index,c670_wt.columns]
c670_op_real = c670_df.loc[c670_op.index,c670_op.columns]
res = pd.DataFrame(index=c670_wt.columns,columns=['R2','MSE','MAPE'])
for i in c670_wt.columns:
res.loc[i,'R2'] = np.clip(r2_score(c670_wt_real[i],c670_wt[i]),0,1)
res.loc[i,'MSE'] = mean_squared_error(c670_wt_real[i],c670_wt[i])
try:
res.loc[i,'MAPE'] = c670.mape(c670_wt_real[i],c670_wt[i],e=0.02)
except:
pass
res.loc['AVG'] = res.mean(axis=0)
res
# + id="KsKYDOJPE8eK" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="4e05238f-1676-4022-f28c-1d35b6670daa"
res = pd.DataFrame(index=c670_op.columns,columns=['R2','MSE','MAPE'])
for i in c670_op.columns:
res.loc[i,'R2'] = r2_score(c670_op_real[i],c670_op[i])
res.loc[i,'MSE'] = mean_squared_error(c670_op_real[i],c670_op[i])
res.loc[i,'MAPE'] = c670.mape(c670_op_real[i],c670_op[i],e=0.02)
res.loc['AVG'] = res.mean(axis=0)
res
# + [markdown] id="hACfpgAOSCJJ"
# # 2.使用預訓練模型
# + [markdown] id="WeKSkdwaTJfO"
# 2.1 用模擬數據先預訓練一個模型
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="7Y4PRpDKQkO0" outputId="375b9835-1565-4bab-a06b-bd831b494709"
c670_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c670_train.csv',index_col=0).dropna(axis=0)
c670 = PartBulider(c670_df,x_col,y_col,normalize_idx_list=n_idx,limit_y_range=True,max_epochs=50)
c670.train()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="bixk9_ejTdCp" outputId="b3b0db40-74db-46d4-ee76-a1d794ec9215"
c670.test(e=2e-2)
# + [markdown] id="Dp2QVIWITP77"
# 2.2 把預訓練好的模型抽出來
# + colab={"base_uri": "https://localhost:8080/"} id="zVYPD8gxQkR2" outputId="0c946650-4e94-4e6d-85ee-88a990afda8e"
import copy
from copy import deepcopy
pretrain_net = deepcopy(c670.net)
pretrain_net
# + [markdown] id="LZK_slJWUVDg"
# 2.3 在預訓練模型上丟進真實資料繼續訓練
# + colab={"base_uri": "https://localhost:8080/", "height": 401} id="viiNO3m2UAIl" outputId="61ed41b5-abc0-4311-d7d3-c7dedf3e8ac8"
from torch.optim import Adam
c670_df = pd.read_excel('/content/drive/MyDrive/台塑輕油案子/data/c620/明志_遷移式學習_訓練資料_寄送版/c670_data.xlsx',index_col=0)
c670 = PartBulider(c670_df,x_col,y_col,normalize_idx_list=n_idx,limit_y_range=True)
c670.net = pretrain_net
c670.optimizer = Adam(c670.net.parameters(),lr=0.001)
c670.train()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="ANAtT_f_QkUy" outputId="09f2a7c8-dcf6-4c6a-95e5-1dc741efe68f"
c670.test(e=2e-2)
# + id="fKwBRbULGYrW" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="09eba6d4-efea-48e8-ef30-7d80d1e0503b"
from autorch.function import sp2wt
c670_sp,c670_op = c670.predict(c670.data['X_test']).iloc[:,:41*2],c670.predict(c670.data['X_test']).iloc[:,41*2:]
c670_feed = c670.data['X_test'][c['combined']].values
s2,s4 = c670_sp.iloc[:,0:41].values,c670_sp.iloc[:,41:41*2].values
w2,w4 = sp2wt(c670_feed,s2),sp2wt(c670_feed,s4)
wt = np.hstack((w2,w4))
c670_wt = pd.DataFrame(wt,index=c670.data['X_test'].index,columns=c['distillate_x']+c['bottoms_x'])
c670_wt_real = c670_df.loc[c670_wt.index,c670_wt.columns]
c670_op_real = c670_df.loc[c670_op.index,c670_op.columns]
res = pd.DataFrame(index=c670_wt.columns,columns=['R2','MSE','MAPE'])
for i in c670_wt.columns:
res.loc[i,'R2'] = np.clip(r2_score(c670_wt_real[i],c670_wt[i]),0,1)
res.loc[i,'MSE'] = mean_squared_error(c670_wt_real[i],c670_wt[i])
try:
res.loc[i,'MAPE'] = c670.mape(c670_wt_real[i],c670_wt[i],e=0.02)
except:
pass
res.loc['AVG'] = res.mean(axis=0)
res
# + id="hU2ME5doGYw9" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="0fdbd8c7-32bb-4b24-e662-179e2ff030c5"
res = pd.DataFrame(index=c670_op.columns,columns=['R2','MSE','MAPE'])
for i in c670_op.columns:
res.loc[i,'R2'] = r2_score(c670_op_real[i],c670_op[i])
res.loc[i,'MSE'] = mean_squared_error(c670_op_real[i],c670_op[i])
res.loc[i,'MAPE'] = c670.mape(c670_op_real[i],c670_op[i],e=0.02)
res.loc['AVG'] = res.mean(axis=0)
res
# + id="X-sT25-jUfOt"
c670.shrink()
# + colab={"base_uri": "https://localhost:8080/"} id="eI7nG4O-U8Am" outputId="913f2309-f6c1-43bb-e836-f51b9348f8a6"
joblib.dump(c670,'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c670_real_data.pkl')
# + id="WaCC7PUMVISe"
| notebook/c670_transferlearning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Raspberry Pi Mouse sensor monitor
#
# [raspimouse_http_controller](https://github.com/Tiryoh/raspimouse_http_controller)と組み合わせて使用するラズパイマウスのセンサ値連続受信用Jupyter Notebookです。
#
# 注がある場合を除き、本ページに掲載されているコードは[MIT](https://tiryoh.mit-license.org/)ライセンスに、文章は[CC BY 4.0](https://creativecommons.org/licenses/by/4.0/deed.ja)ライセンスに基づいて公開します。
#
# 実際に動いている様子は<a href="https://youtu.be/sBr1ngoACnU" target="_blank">YouTubeの動画</a>で確認できます。
#
# ## Requirements
#
# Python 3を必要とします。
# # step1
# Pythonのモジュールをインポートします。
# Jupyter Notebookでの描画に必要なモジュールと、ラズパイマウスとの通信に必要なモジュールと分かれています。
# +
# 数値計算及び描画に必要なモジュール
import numpy as np
import math
from matplotlib import pyplot as plt
from matplotlib import patches as patches
# 通信用
import urllib.request
import time
import argparse
import json
import threading
# JupyterNotebook用モジュール
from IPython import display
# デバッグ用
from pprint import pprint
# -
# # step2
#
# 受信用のクラス(Receiver)を作成します。ラズパイマウスとTCP通信を行います。
class Receiver(object):
def __init__(self, ip, port):
print("init")
self.url = "http://{}:{}".format(ip, port)
print ("connecting to " + self.url)
def get_sensor_val(self):
start = time.time()
request = urllib.request.Request(self.url)
response = urllib.request.urlopen(request)
sensor_dict = json.loads(response.read().decode('utf-8'))
return sensor_dict
# # step3
#
# ラズパイマウスに接続します。
# 引数にはIPアドレスと使用するTCPポートを指定します。
# r = Receiver("192.168.64.3", 5000)
r = Receiver("192.168.22.5", 5000)
# # step4
#
# 試しにラズパイマウスからセンサの値を受信し、`print` してみます。
history = []
hoge = r.get_sensor_val()
history.append(hoge)
print(hoge)
print(history)
# # step5
#
# ラズパイマウスからセンサの値を連続受信し、`pprint` してみます。
# `pprint` を用いることで `print` される内容が整形されます。
# +
for i in range(10):
hoge = r.get_sensor_val()
history.append(hoge)
time.sleep(0.1)
pprint(history)
# -
# # step6
#
# ラズパイマウスからセンサの値を連続受信し、受信するたびにグラフに描画します。
# +
history = []
for i in range(15):
# リストが長くなったらリストを初期化
if len(history) > 99:
history = []
# ラズパイマウスから情報取得
for i in range(5):
hoge = r.get_sensor_val()
history.append(hoge)
time.sleep(0.1)
# タイトル付き新規ウィンドウ、座標軸を用意
fig = plt.figure(figsize=(22, 10))
# 左側の履歴表示
graph = plt.subplot(1,2,1)
graph.set_title("sensor history")
graph.set_xlim(0,100)
graph.set_ylim(-1.5,2000)
# 右側の現在の状態表示
view = plt.subplot(1,2,2)
view.set_title("robot environment (top view)")
view.set_xlim(-10,10)
view.set_ylim(-10,10)
# 描画用にセンサ値ごとのリストを用意
x_arr = np.array([])
y1_arr = np.array([])
y2_arr = np.array([])
y3_arr = np.array([])
y0_arr = np.array([])
# リストにセンサ値を代入
for i in range(len(history)):
x_arr = np.append(x_arr, i)
y0_arr = np.append(y0_arr, history[i]["lightsensor"]["0"])
y1_arr = np.append(y1_arr, history[i]["lightsensor"]["1"])
y2_arr = np.append(y2_arr, history[i]["lightsensor"]["2"])
y3_arr = np.append(y3_arr, history[i]["lightsensor"]["3"])
# センサの履歴表示
graph.plot(x_arr, y0_arr, label = "sensor 0(right front)", color = 'blue')
graph.plot(x_arr, y1_arr, label = "sensor 1(right side)", color = 'orange')
graph.plot(x_arr, y2_arr, label = "sensor 2(left side)", color = 'green')
graph.plot(x_arr, y3_arr, label = "sensor 3(left front)", color = 'red')
# ロボットの状態を表示
rf_sensor_value = (history[len(history)-1]["lightsensor"]["0"])/100
rf = plt.Line2D(xdata=(2.0, 2.0), ydata=(0.0, rf_sensor_value), color='blue', linewidth=2)
rs_sensor_value = (history[len(history)-1]["lightsensor"]["1"])/100
rs = plt.Line2D(xdata=(0.5, 0.5+rs_sensor_value*math.cos(np.deg2rad(45))), ydata=(0.0, rs_sensor_value*math.sin(np.deg2rad(45))), color='orange', linewidth=2)
lf_sensor_value = (history[len(history)-1]["lightsensor"]["2"])/100
lf = plt.Line2D(xdata=(-2.0, -2.0), ydata=(0.0, lf_sensor_value), color='red', linewidth=2)
ls_sensor_value = (history[len(history)-1]["lightsensor"]["3"])/100
ls = plt.Line2D(xdata=(-0.5, -0.5-ls_sensor_value*math.cos(np.deg2rad(45))), ydata=(0.0, ls_sensor_value*math.sin(np.deg2rad(45))), color='green', linewidth=2)
robot = patches.Rectangle(xy=(-3, -6), width=6, height=6, ec='#000000', fill=False)
view.add_line(rf)
view.add_line(rs)
view.add_line(lf)
view.add_line(ls)
view.add_patch(robot)
view.text(0, -5 , "Raspberry Pi Mouse", horizontalalignment='center')
view.text(0, -1 , "↑ front side of the robot", horizontalalignment='center')
# 左上に凡例を追加
graph.legend(loc='upper left')
# 現在の状態を確認
display.clear_output(wait=True)
display.display(plt.gcf())
#time.sleep(0.05)
print("done.")
# -
| sensor_monitor-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Science 2
# ## Numerical analysis - Numerical integration
#
# The following material is covered in Chapter 6 - *Numerical Integration* of the book *Numerical methods in engineering with Python 3* by <NAME> (see BlackBoard).
# ### Introduction
#
# [Numerical integration](https://en.wikipedia.org/wiki/Numerical_integration), also known as *quadrature*, is intrinsically a much more accurate procedure than numerical differentiation. Quadrature approximates the definite integral
#
# $$
# I = \int_a^b f(x) \text{d}x
# $$
#
# by the sum
#
# $$
# I \approx \sum_{i=0}^n A_i f(x_i)
# $$
#
# where the *abscissas* $x_i$ and *weights* $A_i$ depend on the particular rule used for the quadrature. All rules of quadrature are derived from polynomial interpolation of the integrand. Therefore, they work best if $f(x)$ can be approximated by a polynomial.
# ### Newton-Cotes Formulas
#
# [Newton-Cotes formulas](https://en.wikipedia.org/wiki/Newton%E2%80%93Cotes_formulas) are characterized by equally spaced abscissas and include well-known methods such as the trapezoidal rule and Simpson’s rule. They are most useful if $f(x)$ has already been computed at equal intervals or can be computed at low cost. Because Newton-Cotes formulas are based on local interpolation, they require only a piecewise fit to a polynomial.
#
# Consider the definite integral $I = \int_a^b f(x) \text{d}x$. We divide the range of integration $a < x < b$ into $n$ equal intervals of length $h = \frac{b-a}{n}$, and denote the abscissas of the resulting nodes by $x_0$, $x_1$ ,... , $x_n$. Next we approximate $f(x)$ by a polynomial of degree $n$ that intersects all the nodes.
#
# If $n = 1$, we approximate the function $f(x)$ by a linear function. The area under the curve therefore corresponds with a trapezoid. Its area equals
#
# $$
# I = \left( f(a) + f(b) \right) \frac{h}{2}
# $$
#
# This is known as the [trapezoidal rule](https://en.wikipedia.org/wiki/Trapezoidal_rule). In practice the trapezoidal rule is applied in a piecewise fashion. The function $f(x)$ to be integrated is approximated by a piecewise linear function through all points $(x_i, f(x_i))$. From the trapezoidal rule we obtain for the approximate total area representing $\int_a^b f(x) \text{d}x$
#
# $$
# I = \sum_{i=0}^{n-1} I_i = \left( f(x_0) + 2 f(x_1) + 2 f(x_2) + \ldots + 2 f(x_{n-1}) + f(x_n) \right) \frac{h}{2}
# $$
#
# which is the *composite trapezoidal rule*.
#
# It can be shown that the error in each term $I_i$ is of the order $\mathcal{O}(h^3)$. However, because the number of trapezoids equals $n = \frac{b-a}{h}$, the error of the composite trapezoidal rule cumulates to $\mathcal{O}(h^2)$.
#
# **Exercise 1**
#
# Complete the below function `trapezoid()` that implements the composite trapezoidal rule. Choose a reasonable default for the number of intervals.
import numpy as np
def trapezoid(f, a, b, n=1000):
"""df = trapezoid(f, a, b, n=...).
Calculates the definite integral of the function f(x)
from a to b using the composite trapezoidal rule with
n subdivisions (with default n=...).
"""
x = np.linspace(a, b, n + 1)
y = f(x)
y_fir = f(a)
y_last = f(b)
h = (b - a ) / n
I = (h/2) * np.sum(y_first + y_last)
return I
# +
import numpy as np
def trapezoid(f, a, b, n=1000):
"""df = trapezoid(f, a, b, n=...).
Calculates the definite integral of the function f(x)
from a to b using the composite trapezoidal rule with
n subdivisions (with default n=...).
"""
h = (b - a ) / n
I = f(a) + f(b) / 2
for i in range(1, n-1):
xi = a + i * h
I += f(xi)
I *= h
return I
# -
# Below, we apply the composite trapezoidal rule to calculate the integral of the cosine function from $-\frac{\pi}{2}$ to $\frac{\pi}{2}$, which analytically evaluates to $\int_{-\pi/2}^{\pi/2} \cos(x) \text{d}x = \sin(\frac{\pi}{2}) - \sin(-\frac{\pi}{2}) = 2$. Verify that the error of the composite trapezoidal rule is of order $\mathcal{O}(h^2)$.
# +
# Example: integral of cos(x) from -pi/2 to pi/2
from math import cos, pi
ns = [1, 10, 100, 1000, 10000, 100000]
I_exact = 2.0
for n in ns:
I_trapezoid = trapezoid(cos, -0.5 * pi, 0.5 * pi, n)
print(f'n = {n:8}: {I_trapezoid:10.3e} (error={I_trapezoid-I_exact:8.1e})')
# -
# ### Recursive Trapezoidal Rule
#
# Let $I_k$ be the integral evaluated with the composite trapezoidal rule using $2^k$ panels. Note that if $k$ is increased by one, the number of panels is doubled. Using the notation $h_k=\frac{b−a}{2^k}$ for the interval size, we obtain the following results.
#
# * $k = 0$ (one panel):
#
# $$
# I_0 = \left( f(a) + f(b) \right) \frac{h_0}{2}
# $$
#
# * $k = 1$ (two panels):
#
# $$
# I_1 = \left( f(a) + 2 f(a+h_1) + f(b) \right) \frac{h_1}{2} = \frac{1}{2} I_0 + f(a+h_1) h_1
# $$
#
# * $k = 2$ (four panels):
#
# $$
# I_2 = \left( f(a) + 2 f(a+h_2) + 2 f(a+2h_2) + 2 f(a+3h_2) + f(b) \right) \frac{h_2}{2} = \frac{1}{2} I_1 + \left( f(a+h_2) + f(a+3h_2) \right) h_2
# $$
#
# We can now see that for arbitrary $k > 0$ we have
#
# $$
# I_k = \frac{1}{2} I_{k-1} + h_k \cdot \sum_{i=1, 3, 5, \ldots, 2^k-1} f(a+i \cdot h_k)
# $$
#
# which is the *recursive trapezoidal rule*. Observe that the summation contains only the new nodes that were created when the number of panels was doubled. Therefore, the computation of the entire sequence $I_0, I_1, \ldots, I_k$ involves the same amount of algebra as the calculation of $I_k$ directly.
#
# However, the advantage of using the recursive trapezoidal rule is that it allows us to monitor convergence and terminate the process when the difference between $I_{k−1}$ and $I_k$ becomes sufficiently small.
#
# **Exercise 2**
#
# Rewrite the function `trapezoid()` such that it computes $I_k$ iteratively, given $I_{k−1}$, until it achieves an estimated accuracy set by the user through the tolerance parameter `tol` (i.e., stop when $|I_k - I_{k−1}| < \text{tol}$). Again, pick a reasonable default value for that tolerance parameter.
# +
def trapezoid(f, a, b, tol=1e-8):
"""df = trapezoid(f, a, b, tol=...).
Calculates the definite integral of the function f(x)
from a to b using the recursive trapezoidal rule with
an absolute tolerance tol (with default 1e-8).
"""
h = (b - a) # Interval size
panels = 1 # No. of intervals
I_old = (f(a)+f(b)) * h/2
while True:
h /= 2
panels *= 2
I_new = 0.5 * I_old + sum(f(a +i*h) for i in range(1, panels, 2)) * h
if abs(I_new - I_old) < tol:
return I_new
else:
I_old = I_new
# -
# Below, we again apply the recursive version of the composite trapezoidal rule to calculate the integral of the cosine function from $-\frac{\pi}{2}$ to $\frac{\pi}{2}$. Verify that the specified tolerance (or better) is indeed reached.
# Example: integral of cos(x) from -pi/2 to pi/2
I_trapezoid = trapezoid(cos, -0.5 * pi, 0.5 * pi, 1e-4)
print(f'I: {I_trapezoid:13.6e} (error={I_trapezoid-I_exact:8.1e})')
# ### Simpson's Rule
#
# [Simpson's rule](https://en.wikipedia.org/wiki/Simpson%27s_rule) can be obtained from Newton-Cotes formulas with $n = 2$; that is, by passing a parabolic interpolant through three adjacent nodes, each separated by $h$. The area under the parabola, which represents an approximation of $I = \int_a^b f(x) \text{d}x$, can be shown to equal
#
# $$
# I = \left( f(a) + 4 f(\frac{a+b}{2}) + f(b) \right) \frac{h}{3}
# $$
#
# To obtain the *composite Simpson's rule*, the integration range $(a, b)$ is divided into $n$ panels (with $n$ even) of width $h = \frac{b − a}{n}$ each. Applying the above formula to two adjacent panels, we obtain
#
# $$
# I = \left( f(x_0) + 4f(x_1) + 2f(x_2) + 4f(x_3) + \ldots + 2f(x_{n−2}) + 4f(x_{n−1}) + f(x_n) \right) \frac{h}{3}
# $$
#
# The composite Simpson's rule is perhaps the best known method of numerical integration. However, its reputation is somewhat undeserved, because the trapezoidal rule is more robust and Romberg integration (below) is more efficient.
#
# **Exercise 3**
#
# Write a function `simpson()` that implements the composite Simpson's rule.
def simpson(f, a, b, n=100):
"""df = simpson(f, a, b, n=...).
Calculates the definite integral of the function f(x)
from a to b using the composite Simpson's
rule with n subdivisions (with default n=...).
"""
n += n % 2 # force to be even
h = (b -a) / n
I = f(a) + f(b)
for i in range(1, n, 2):
xi = a + i*h
I += 4*f(xi)
for i in range(2, n, 2):
xi = a + i*h
I += 2*f(xi)
I *= h/3
return I
# We once more apply the composite Simpson's rule to the cosine integral. What is the order of the method, and why does its accuracy start to break up when $n$ reaches 10000?
# Example: integral of cos(x) from -pi/2 to pi/2
for n in ns:
I_simpson = simpson(cos, -0.5 * pi, 0.5 * pi, n)
print(f'n = {n:8}: {I_simpson:10.3e} (error={I_simpson-I_exact:8.1e})')
# Simpson's rule can also be converted to a recursive form. However, this is a bit more challenging because the weights of the odd and even terms alternate.
#
# ### Romberg Integration
#
# [Romberg integration](https://en.wikipedia.org/wiki/Romberg%27s_method) is based on the trapezoidal rule. It evaluates an integral using a series of different interval sizes. Subsequently, these various answers are averaged using carefully chosen weights that are tuned in such a way that the errors tend to cancel. Thus, a solution can be found that is much more accurate than any of the individual evaluations. This approach of combining multiple solutions is called [Richardson extrapolation](https://en.wikipedia.org/wiki/Richardson_extrapolation).
#
# We will not derive the method here, but merely describe it. For more details, see the book chapter.
#
# Let us first introduce the notation $R_{k,0} = I_k$, where, as before, $I_k$ represents the approximate value of $I = \int_a^b f(x) \text{d}x$ computed by the recursive trapezoidal rule using $2^k$ panels. Romberg integration starts with the computation of $R_{0,0} = I_0$ (one panel) and $R_{1,0} = I_1$ (two panels) from the trapezoidal rule. We already know that these have an error of order $\mathcal{O}(h^2)$. These two estimates are combined linearly in order to obtain a better estimate according to $R_{1,1} = \frac{4}{3} R_{1,0} - \frac{1}{3} R_{0,0}$ that turns out to have an error $\mathcal{O}(h^4)$.
#
# It is convenient to store the results in a triangular array of the form
#
# $$
# \begin{array}{cc}
# R_{0,0} = I_0 &\\
# & R_{1,1} = \frac{4}{3} R_{1,0} - \frac{1}{3} R_{0,0}\\
# R_{1,0} = I_1 &
# \end{array}
# $$
#
# The next step is to calculate $R_{2,0} = I_2$ (four panels) and repeat the combination procedure with $R_{1,0}$ and $R_{2,0}$, storing the result as $R_{2,1} = \frac{4}{3} R_{2,0} - \frac{1}{3} R_{1,0}$. The elements $R_{2,0}$ and $R_{2,1}$ are now both $\mathcal{O}(h^4)$ approximations, which can in turn be combined to obtain $R_{2,2} = \frac{16}{15} R_{2,1} - \frac{1}{15} R_{1,1}$ with error $\mathcal{O}(h^6)$. The array has now expanded to
#
# $$
# \begin{array}{ccc}
# R_{0,0} = I_0 & &\\
# & R_{1,1} = \frac{4}{3} R_{1,0} - \frac{1}{3} R_{0,0} &\\
# R_{1,0} = I_1 & & R_{2,2} = \frac{16}{15} R_{2,1} - \frac{1}{15} R_{1,1}\\
# & R_{2,1} = \frac{4}{3} R_{2,0} - \frac{1}{3} R_{1,0} &\\
# R_{2,0} = I_2 & &
# \end{array}
# $$
#
# After another round of calculations we get
#
# $$
# \begin{array}{cccc}
# R_{0,0} = I_0 & & &\\
# & R_{1,1} = \frac{4}{3} R_{1,0} - \frac{1}{3} R_{0,0} & &\\
# R_{1,0} = I_1 & & R_{2,2} = \frac{16}{15} R_{2,1} - \frac{1}{15} R_{1,1} &\\
# & R_{2,1} = \frac{4}{3} R_{2,0} - \frac{1}{3} R_{1,0} & & R_{3, 3} = \frac{64}{63} R_{3,2} - \frac{1}{63} R_{2,2}\\
# R_{2,0} = I_2 & & R_{3,2} = \frac{16}{15} R_{3,1} - \frac{1}{15} R_{2,1} &\\
# & R_{3,1} = \frac{4}{3} R_{3,0} - \frac{1}{3} R_{2,0} & &\\
# R_{3,0} = I_3 & & &
# \end{array}
# $$
#
# where the error in $R_{3,3}$ is $\mathcal{O}(h^8)$.
#
# The general extrapolation formula used in this scheme is
#
# $$
# R_{i,j} = \frac{4^j R_{i,j−1} - R_{i−1,j−1}}{4^j - 1}
# $$
#
# **Exercise 4**
#
# Implement a function `romberg()` that performs Romberg integration until a tolerance `tol` is achieved. Note that the most accurate estimate of the integral is always the last diagonal term of the array, so the process needs to be continued until the difference between two successive diagonal terms $|R_{i,i} - R_{i-1,i-1}| < \text{tol}$. Although the triangular array is convenient for hand computations, computer implementation of the Romberg algorithm can be carried out within a one-dimensional array $\boldsymbol{r}$ (i.e. a list or a vector) that contains a diagonal row of the array $R_{i, :}$ at any time.
def romberg(f, a, b, tol = 1e-8):
"""df = simpson(f, a, b, tol=...).
Calculates the definite integral of the function f(x)
from a to b using Romberg integration based on the
trapezoidal rule until a specified tolerance tol is
reached (with default tol=...).
"""
h = (b - a) # Interval size
n = 1 # No. of intervals
Rold = [ (f(a)+f(b)) * h/2 ]
while True:
h /= 2
n *= 2
Rnew = [ 0.5 * Rold[0] + sum(f(a +o*h) for o in range(1, n, 2)) * h ]
factor = 1
for R in Rold:
factor *= 4
Rnew.append( (factor*Rnew[-1] - R) / (factor-1) )
if abs(Rnew[-1] - Rold[-1]) < tol:
return Rnew[-1]
Rold = Rnew
exact = np.pi/4
exact
def f(x):
y = 1.0 / (1.0 + x**2)
return y
R00 = (f(0) + f(1)) * 1/2
R00
R10 = (1/2)*R00 + f(1/2)* (1/2)
R10
R20 = (1/2)*R10 + (f(1/4) + f(3/4)) *(1/4)
R20
R30 = (1/2)*R20 + (f(1/8) + f(3/8) + f(5/8) + f(7/8)) * 1/8
R30
R11 = (4*R10 -R00)/ 3
R11
R21 = (4*R20 -R10)/ 3
R21
R31 = (4*R30 -R20)/ 3
R31
R22 = (16*R21 -R11)/ 15
R22
R32 = (16*R31 -R21)/ 15
R32
R33 = (64*R32 -R22)/ 63
R33
# We apply the Romberg integration rule to the cosine integral one final time. Once more, verify that the specified tolerance (or better) is indeed reached.
# Example: integral of cos(x) from -pi/2 to pi/2
I_romberg = romberg(cos, -0.5 * pi, 0.5 * pi, tol=1e-4)
print(f'I: {I_romberg:13.6e} (error={I_romberg-I_exact:8.1e})')
# ### Exercises
#
# **Exercise 5**
#
# Determine the value of the definite integral $\int_0^1 2^x \text{d}x$ to approximately six decimals using the following three methods:
#
# * analytically, using symbolic integration;
#
# * using your own functions `trapezoid()`, `simpson()` and `romberg()`;
#
# * using the functions [quadrature](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.quadrature.html) and [romberg](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.romberg.html) of the module `scipy.integrate`.
#
# Which are the most accurate?
import scipy.integrate as sc
import matplotlib.pyplot as plt
def f(x):
return 2 **x
x = np.linspace(-5, 5, 300)
plt.plot(x, f(x), '-k')
plt.axis([-1.5, 1.5, 0, 4])
plt.axvline(0)
plt.axvline(1)
plt.show()
print('trapezoid ', trapezoid(f, 0, 1, 1e-8))
print('simpson ', simpson(f, 0, 1, 1000000))
print('romberg ', romberg(f, 0, 1, 1e-8))
print('scipy quad',sc.quadrature(f, 0, 1)[0])
print('scipy rom ',sc.romberg(f, 0, 1))
# **Exercise 6**
#
# A circle with radius 1 can be described by the equation $x^2 + y^2 = 1$. From this equation, you can derive the function $y(x)$ that describes the upper half of this circle. Theoretically, the area below this curve should therefore equal $\frac{1}{2}\pi$. Using this function in combination with the recursive trapezoid method and the Romberg integration method, calculate the value of $\pi$ up to twelve decimals accuracy. How do the runtimes of these methods compare? Hint: use the `%time` [notebook magic command](https://ipython.readthedocs.io/en/stable/interactive/magics.html#magic-time).
x = np.linspace(-1, 1, 300)
def y(x):
return np.sqrt(1 - x**2)
plt.plot(x, y(x), '-k')
plt.axvline(0)
plt.axhline(0)
plt.show()
np.pi
# +
# %time romberg(y, -1, 1, 1e-12)*2
# -
# %time trapezoid(y, -1, 1, 1e-12)*2
# **Exercise 7**
#
# Plot the function $f(x) = \sqrt{x^2-x^4}$ and calculate the area under this curve between $x=-1$ and $x=1$. Use your own trapezoid and Romberg integration rules with a tolerance `tol=1e-6`. Explain why the outcomes do not seem to make sense.
import matplotlib.pyplot as plt
def f(x):
return np.sqrt(x**2 - x**4)
# +
x = np.linspace(-1, 1, 500)
y= f(x)
plt.plot(x, y, 'k')
plt.axvline(0)
plt.show()
# -
romberg(f, -1,1, tol=1e-6)
# **Exercise 8**
#
# The present functions do not seem to be able to compute integrals with bounds that involve infinity. However, this can be circumvented by means of a coordinate transformation. For instance, to calculate the integral
#
# $$
# I = \int_{-\infty}^{\infty} e^{-z^2} \text{d}z
# $$
#
# that is hopefully familiar from the gaussian distribution, we can use a transformation like for instance
#
# $$
# z = \frac{t}{1-t^2}
# $$
#
# Verify for yourself that when $t$ goes from -1 to +1, $z$ goes from $-\infty$ to $+\infty$. Now, because
#
# $$
# \frac{\text{d}z}{\text{d}t} = \frac{1+t^2}{(1-t^2)^2}
# $$
#
# the integral can be rewritten as
#
# $$
# I = \int_{-1}^1 e^{-\left( \frac{t}{1-t^2} \right)^2} \cdot \frac{1+t^2}{(1-t^2)^2} \text{d}t
# $$
#
# Compute the value of the integral $\int_{-\infty}^{\infty} e^{-z^2} \text{d}z$ to approximately nine digits accuracy using an algorithm of your own choice and compare it to the theoretical value $I = \sqrt{\pi}$. Hint: slightly adjust the integration limits to prevent division by zero errors.
# +
def z(x):
return (np.e ** -(x / (1 - x**2))**2) * ( (1 + x**2) / (1 - x**2)**2 )
x = np.linspace(-0.9, 0.9, 500)
# -
plt.plot(x, z(x), "-b")
plt.axvline(0)
plt.axhline(0)
plt.show()
# +
# Example: integral of cos(x) from -pi/2 to pi/2
I_romberg = romberg(z,-0.9, 0.9, tol=1e-9)
I_exact = np.sqrt(np.pi)
print(f'I: {I_romberg:1.10} (error={I_romberg-I_exact:8.1e})')
# -
# ***
| Numerical_analysis/Lessons/.ipynb_checkpoints/2 - Numerical integration-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys
from pathlib import Path
import pysal as ps
import geopandas as gpd
import numpy as np
from libpysal.weights import Queen, Rook, KNN
from esda.moran import Moran_Local_BV
DIR = Path('..')
sys.path.append(str(DIR))
DATA_DIR = DIR/'data/'
# +
taz_gdf = gpd.read_file(
GIS_DIR/'census_2018_MB_WGS84.geojson,
)
# Calculate the weight
w = Queen.from_dataframe(taz_gdf)
#Calculate X
x = np.array(taz_gdf['Median_Hou'])
#Calculate Y
y = np.array(taz_gdf['PT_Job_Acc'])
#LISA
lm = Moran_Local_BV(x, y, w, transformation = "r", permutations = 99)
taz_gdf['LISA_class'] = lm.q
d = {
1: 'High income, High accessibility',
2: 'Low income, High accessibility',
3: 'Low income, Low accessibility',
4: 'High income, Low accessibility',
}
taz_gdf['LISA_class'] = taz_gdf['LISA_class'].replace(d)
taz_gdf['LISA_sig'] = lm.p_sim
taz_gdf.to_file(str(DATA_DIR/'cluster.shp'))
| ipynb/LISA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FL Simulator
#
# Simulating `Federated Learning` paradigm which averages neighbor's model weights into local one.
#
# ## Features
#
# * Byzantine
#
# ## TODO
#
# * Network topology
import import_ipynb
import nn.dist as dist
import nn.ensemble as ensemble
import nn.ml as ml
import nn.nets as nets
# + tags=[]
if __name__ == "__main__":
import os
from copy import deepcopy
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
import torchvision.datasets as dset
import torchvision.transforms as transforms
from torch.utils.data import DataLoader # TODO: DistributedDataParallel
"""Hyperparams"""
numNets = 21
# numByzs = 0
numWorkers = 4
cuda = True
base_path = './simul_21_uniform_0_byzantine_fedavg'
trainFiles = [None for _ in range(numNets)]
testFiles = [None for _ in range(numNets)]
for i in range(numNets):
path = os.path.join(base_path, str(i))
os.makedirs(path, exist_ok=True)
trainFiles[i] = open(os.path.join(path, 'train.csv'), 'w')
testFiles[i] = open(os.path.join(path, 'test.csv'), 'w')
testFile_global = open(os.path.join(base_path, 'test_global.csv'), 'w')
testFile_ensemble = open(os.path.join(base_path, 'test_ensemble.csv'), 'w')
epochs = 3000
batchSz = 64
"""Datasets"""
# # gets mean and std
# transform = transforms.Compose([transforms.ToTensor()])
# dataset = dset.CIFAR10(root='cifar', train=True, download=True, transform=transform)
# normMean, normStd = dist.get_norm(dataset)
normMean = [0.49139968, 0.48215841, 0.44653091]
normStd = [0.24703223, 0.24348513, 0.26158784]
normTransform = transforms.Normalize(normMean, normStd)
trainTransform = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normTransform
])
testTransform = transforms.Compose([
transforms.ToTensor(),
normTransform
])
trainset = dset.CIFAR10(root='cifar', train=True, download=True, transform=trainTransform)
testset = dset.CIFAR10(root='cifar', train=False, download=True, transform=trainTransform)
# splits datasets
splited_trainset = dist.random_split_by_dist(
trainset,
size=numNets,
dist=dist.uniform,
# alpha=2.
)
splited_testset = dist.random_split_by_dist(
testset,
size=numNets,
dist=dist.uniform,
# alpha=2.
)
# num_workers: number of CPU cores to use for data loading
# pin_memory: being able to speed up the host to device transfer by enabling
kwargs = {'num_workers': numWorkers, 'pin_memory': cuda}
# loaders
trainLoaders = [DataLoader(
splited_trainset[i], batch_size=batchSz, shuffle=True, **kwargs
) for i in range(numNets)]
testLoaders = [DataLoader(
splited_testset[i], batch_size=batchSz, shuffle=True, **kwargs
) for i in range(numNets)]
global_testLoader = DataLoader(testset, batch_size=batchSz, shuffle=True, **kwargs)
"""Nets"""
num_classes = 10
resnets = [nets.resnet18(num_classes=num_classes) for _ in range(numNets)]
global_model = nets.resnet18(num_classes=num_classes)
criterions = [nn.CrossEntropyLoss() for _ in range(numNets)]
global_criterion = nn.CrossEntropyLoss()
ensemble_criterion = nn.CrossEntropyLoss()
optimizers = [optim.SGD(net.parameters(), lr=1e-1, momentum=0.9) for net in resnets]
if cuda:
for net in (resnets + [global_model]):
# if multi-gpus
if torch.cuda.device_count() > 1:
net = nn.DataParallel(net)
# use cuda
net.cuda()
if cuda:
s = Variable(torch.Tensor([1. / numNets]).cuda().double())
else:
s = Variable(torch.Tensor([1. / numNets]).double())
"""Train & Test models"""
for epoch in range(epochs):
# aggregation and averaging
global_state_dict = global_model.state_dict()
local_params = dict(resnets[0].named_parameters())
for name, param in global_state_dict.items():
if name in local_params.keys():
global_state_dict[name].fill_(0.).double()
for a in range(numNets):
v = dict(resnets[a].named_parameters())[name]
t = v.clone().detach()
t.mul_(s.expand(v.size()))
global_state_dict[name].add_(t)
ml.test(
global_model, global_criterion, global_testLoader,
epoch=epoch, cuda=cuda, log=True, log_file=testFile_global
)
# ensemble
avg_ensemble = ensemble.Ensemble(deepcopy(resnets), mode=ensemble.avg)
ml.test(
avg_ensemble, ensemble_criterion, global_testLoader,
epoch=epoch, cuda=cuda, log=True, log_file=testFile_ensemble
)
# # byzantines
# # random normal distribution
# for b in range(numByzs):
# # TODO
# # weights.WrapedWeights(resnets[b].named_parameters()).apply(resnets[b])
# ml.test(
# resnets[b], criterions[b], testLoaders[b],
# epoch=epoch, cuda=cuda, log=True, log_file=testFiles[b]
# )
# students
# for i in range(numByzs, numNets):
for i in range(numNets):
resnets[i].load_state_dict(global_model.state_dict())
ml.train(
resnets[i], criterions[i], optimizers[i], trainLoaders[i],
epoch=epoch, cuda=cuda, log=True, log_file=trainFiles[i]
# alpha=0.9, temperature=4
)
ml.test(
resnets[i], criterions[i], testLoaders[i],
epoch=epoch, cuda=cuda, log=True, log_file=testFiles[i]
)
| src/FL_simul.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Explore Your Environment
# ## Get Latest Code
# + language="bash"
#
# pull_force_overwrite_local
# -
# ## Helper Scripts
# ### Find Script from Anywhere
# !which pull_force_overwrite_local
# ### Show `pull_force_overwrite_local` Script
# !cat /root/scripts/pull_force_overwrite_local
# ### List `/scripts` Directory
# !ls -l /root/scripts/
# ## [PipelineAI](http://pipeline.io)
# + language="html"
#
# <iframe width=800 height=600 src="http://pipeline.io"></iframe>
# -
# ## All Code in [GitHub Repo](https://github.com/fluxcapacitor/pipeline/)
# Please Star this [GitHub Repo](https://github.com/fluxcapacitor/pipeline/)!!
# ## [Advanced Spark and TensorFlow Meetup](https://www.meetup.com/Advanced-Spark-and-TensorFlow-Meetup/)
# Please Join this [Global Meetup](https://www.meetup.com/Advanced-Spark-and-TensorFlow-Meetup/)!!
#
#
# ## Verify IP Address
# This should match your browser!
# +
import requests
url = 'http://169.254.169.254/computeMetadata/v1/instance/network-interfaces/0/access-configs/0/external-ip'
headers = {'Metadata-Flavor': 'Google'}
r = requests.get(url, headers=headers)
ip_address = r.text
print('Your IP: %s' % ip_address)
# -
# ## Get Allocation Index
# We may use this later.
# +
import requests
import json
url = 'http://allocator.community.pipeline.ai/allocation/%s' % ip_address
r = requests.get(url, timeout=5)
allocation = r.text
allocation_json = json.loads(allocation)
print(allocation_json)
print(allocation_json['index'])
# -
| jupyterhub/notebooks/gpu/01_Explore_Environment.ipynb |
---
jupyter:
jupytext:
text_representation:
extension: .md
format_name: markdown
format_version: '1.3'
jupytext_version: 1.14.4
kernelspec:
display_name: Markdown
language: markdown
name: markdown
---
# Details
<div style="position: absolute; right:0;top:0"><a href="../evaluation.py.ipynb" style="text-decoration: none"> <font size="5">↑</font></a></div>
## Initialize
`nbbox` is needed for displaying output in the Notebook whereas in the terminal output will go to std out.
Trick: Ipython magic will result in a `NameError` when executing in normal python.
| docs/details.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro
#
# `numpy`는 과학 계산을 위한 파이썬 패키지이며, 다차원 자료구조 `ndarray`를 통해 효율적인 벡터/행렬 계산 기능을 제공합니다. `pandas`는 넘파이를 기반으로 데이터 사이언스에 필요한 자료구조 및 데이터 핸들링 기능들을 제공합니다. 대표적으로 1차원 자료구조인 `Series`와 2차원 자료구조인 `DataFrame`을 지원합니다. 우리는 넘파이보다는 판다스를 집중적으로 다룰 것이지만, 판다스의 자료구조들이 결국은 넘파이의 어레이를 기반으로 만들어지므로 넘파이에 대한 최소한의 이해는 필요합니다.
#
# # 1. Numpy 튜토리얼
#
# 시작하기 전에, `numpy` 패키지를 `np`라는 이름으로 불러오겠습니다.
import numpy as np
# ## 1.1. ndarray
#
# 넘파이의 핵심은 `ndarray` 클래스입니다. 어레이는 지금까지 다룬 리스트와 유사하지만 더욱 강력한 성능과 기능을 제공합니다. `np.array()` 함수 안에 리스트를 집어넣으면 어레이를 만들 수 있습니다.
a = [1,2,3]
np.array(a)
b = [4,5]
np.array(b)
np.array([1,2,3,4,5])
# ## 1.2. 어레이와 자료형
#
# 어레이가 포함하는 요소들은 모두 같은 자료형을 가져야만 하며, 어레이를 만들 때 `dtype` 매개변수를 활용하여 자료형을 직접 지정할 수 있습니다. 데이터 타입을 지정하지 않으면 넘파이가 자동으로 데이터 타입을 판단합니다. 만들어진 어레이의 데이터 타입을 확인할 때는 `dtype` 속성을 활용합니다.
a = np.array([1,2,3], dtype="int8") # 자료가 차지하는 메모리 직접 제한 가능
a.dtype # dtype 속성을 통해 자료형에 접근 가능
# 어레이의 자료형을 변환할 때는 `astype` 메소드를 활용합니다. 괄호 안에 원하는 자료형을 적어주면 어레이를 해당 자료형으로 변환한 결과가 출력됩니다. 역시 어레이를 직접적으로 변화시키는 메소드가 아니기 때문에 결과를 저장하려면 다시 할당을 해주어야 합니다.
a = np.array([1,2,3])
a.astype('str')
a = np.array([1,2,3])
a.astype('float16')
# ## 1.3. n차원 어레이와 shape
#
# 
#
# **넘파이의 1차원 어레이는 하나의 행 벡터처럼 생각할 수 있습니다.** 따라서 여러 개의 1차원 어레이를 결합하면, 하나의 행렬을 만들 수 있을 것입니다. 예를 들어 $\vec{u} = (1,2,3)$ 이라는 벡터와 $\vec{v} = (4,5,6)$이라는 벡터를 두 개의 행으로 갖는 $2 \times 3$ 크기의 행렬 $A$를 넘파이로 구현해보겠습니다.
# $$A = \begin{pmatrix}
# \vec{u} \\
# \vec{v} \\
# \end{pmatrix} =
# \begin{pmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# \end{pmatrix}
# $$
#
#
u = np.array([1,2,3])
v = np.array([4,5,6])
A = np.array([u,v])
A
A.shape
A = np.array([[1,2,3],[4,5,6]])
A
# 위 예시 코드에서 `u`와 `v`는 각각 $(1,2,3)$, $(4,5,6)$ 을 표현하는 어레이입니다. 두 개의 어레이를 다시 `np.array` 함수로 묶은 것이 `A`입니다. `shape`은 어레이의 모양을 나타내는 속성입니다. 즉 행렬로 따지면 $m \times n$ 의 사이즈를 나타냅니다. 오른쪽 코드처럼 리스트로 행렬구조를 표현하여 즉시 어레이를 만들어줄 수도 있습니다.
#
# 차수가 서로 다른 벡터들을 가지고 행렬을 만들 수 없듯이, 길이가 서로 다른 1차원 어레이를 가지고도 2차원 어레이를 만들 수 없습니다. 길이가 서로 다른 1차원 어레이들을 쌓으면 결과는 1차원 어레이들을 요소로 갖는 1차원 어레이가 됩니다.
u = np.array([1,2,3])
v = np.array([4,5])
A = np.array([u,v])
A
A.shape
# 넘파이 배열은 3차원 이상으로도 확장될 수 있습니다. 3차원 이상의 배열로 확장할 때에도 마찬가지로 요소들의 `shape`이 일치해야 합니다. 차원이 늘어나면, `shape`의 앞쪽으로 확장된 차수가 추가됩니다.
#
# $$A =
# \begin{pmatrix}
# 1 & 2 & 3 \\
# 3 & 4 & 5 \\
# \end{pmatrix}, \space
# B =
# \begin{pmatrix}
# 5 & 6 & 7 \\
# 7 & 8 & 9 \\
# \end{pmatrix}
# $$
A = np.array([[1,2,3],[3,4,5]])
A.shape
B = np.array([[5,6,7],[7,8,9]])
B.shape
C = np.array([A,B])
C
C.shape
# 위 예시 코드에서 `A`와 `B`는 각각 $2 \times 3$ 행렬입니다. `C`는 `A` 와 `B`를 쌓아서 만든 $2 \times 2 \times 3$ 텐서입니다. 즉 2개의 행렬 안에 각각 2개의 벡터가 들어있고, 각각의 벡터 안에 3개의 스칼라가 들어있는 모양입니다.
#
# ## 1.4. 어레이 인덱싱
#
# 어레이의 인덱싱 역시 대괄호`[]`와 콜론`:`을 사용합니다. 어레이 인덱싱에서 주의할 점은 인덱싱 문법이 어레이의 `shape`에 대응한다는 점입니다. 즉 `(2,3,4)`와 같은 `shape`을 갖는 어레이에서 하나의 스칼라를 인덱싱하는 경우, `[0,0,0]` 부터 `[1,2,3]` 의 인덱스 범위가 존재하게 됩니다. 여기에서는 2차원 어레이의 인덱싱까지만 다룹니다.
#
# ### 1차원 어레이
#
# 1차원 어레이의 인덱싱은 파이썬 리스트 인덱싱과 크게 다를 것 없습니다.
a = np.array([1,2,3])
a[1]
a[1:-1]
# ### 2차원 어레이
#
# 행렬 $A$ 에 대응하는 2차원 어레이 `A`를 아래와 같이 구현해보겠습니다.
#
# $$A =
# \begin{pmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# \end{pmatrix}
# $$
A = np.array([[1,2,3],[4,5,6]])
A
# 2차원 어레이에서 하나의 스칼라를 인덱싱할 때에는 `A[행인덱스,열인덱스]`과 같이 쓰면 됩니다. 즉 2행 1열의 요소를 인덱싱히려면 다음과 같이 쓰면 됩니다. 파이썬의 인덱스는 `0`부터 시작하기 때문에 `[2,1]`이 아니라 `[1,0]`이 됩니다.
A[:2,-1]
# 이번에는 슬라이싱을 적용해서 세 번째 열 전체를 가져와보겠습니다. 전체 행에 대해서 세 번째 열의 요소만들 가져오므로 구간은 `[:,2]`와 같이 표현합니다.
A[:,2]
# **예제 1.1. 어레이 A에서 4와 5를 인덱싱해보세요.**
A = np.array([[1,2,3],[4,5,6]])
A[1,:2]
# **풀이**
#
# 두 번째 행 벡터에서 두 번째 요소까지를 가져오면 되는 문제입니다. 따라서 구간은 [1,:2]가 됩니다.
A[1,:2]
# ## 1.5. 어레이 연산
#
# ### 같은 쉐입의 연산
#
# 어레이의 쉐입이 같을 때, `+`, `-`, `*`, `/`, `**` 등 기본 연산은 같은 위치의 원소들끼리 연산한 결과를 반환합니다.
u = np.array([1,2,3])
v = np.array([4,5,6])
u + v # 덧셈
u - v # 뺄셈
u * v # 곱셈
u / v # 나눗셈
# ### 다른 `shape`의 연산: 브로드캐스팅
#
# 
#
# 넘파이는 서로 다른 쉐입의 자료들 간에도 연산을 지원합니다. 물론 모든 어레이들이 서로 호환되는 것은 아닙니다. 2차원까지의 어레이에 한정하여 말하면 다음과 같은 경우에 브로드캐스팅이 가능합니다. 여기에서는 스칼라 연산의 결과만을 확인해보겠습니다.
#
# - 스칼라와 벡터, 스칼라와 행렬 간의 연산
# - $m \times n$ 행렬과 $m \times 1$ 벡터 간의 연산
# - $m \times n$ 행렬과 $1 \times n$ 벡터 간의 연산
# - $m \times 1$ 벡터와 $1 \times n$ 벡터 간의 연산
# **덧셈**
a = np.array([1,2,3])
a + 3
#
# **뺄셈**
a = np.array([1,2,3])
a - 1
# **곱셈**
a = np.array([1,2,3])
a * 3
#
# **나눗셈**
a = np.array([1,2,3])
a / 2
# ## 1.6. 기타 함수와 메소드
#
# ### numpy 함수
#
# - `np.dot` : 두 벡터의 내적을 계산
# - `np.matmul` : 두 행렬의 곱을 계산
# - `np.power` : 배열 내 요소들의 n승
# - `np.sqrt` : 배열 내 요소들의 제곱근
#
# ### ndarray 메소드
#
# - `ndarray.transpose` : 전치
# - `ndarray.reshape` : shape 재배열
# # 2. Pandas
#
# 시작하기 전에, `pandas`를 `pd`라는 별칭으로 불러오겠습니다.
import pandas as pd
# ## 2.1. 판다스의 자료형
#
# 판다스의 핵심은 1차원 자료형 클래스 `Series`, 2차원 자료형 클래스 `DataFrame`입니다. 시리즈와 데이터프레임은 각각 넘파이의 1차원 어레이, 2차원 어레이에 더욱 다양한 기능들을 추가하여 만들어졌습니다. **시리즈는 대부분의 경우 하나의 열, 변수, 피쳐를 나타내며, 여러 개의 시리즈들을 한데 묶은 것이 데이터프레임입니다.** 아래 그림은 `apples`, `oranges`라는 두 개의 컬럼이 하나의 데이터프레임을 이루는 모습입니다.
#
# 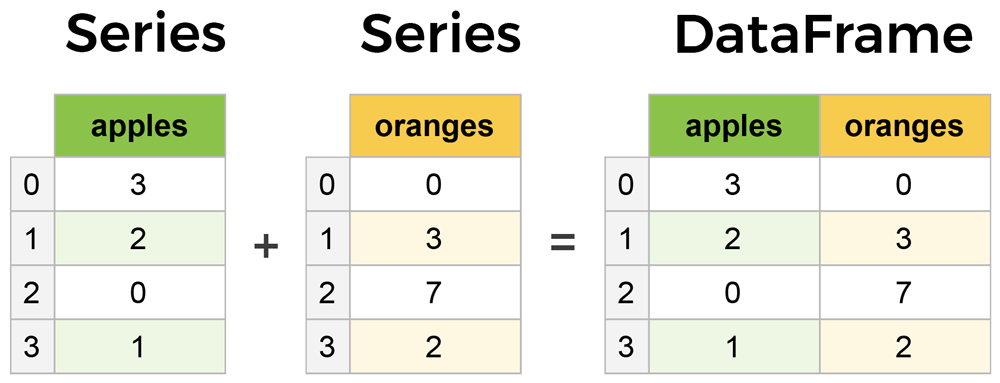
#
# ## 2.2. 시리즈
#
# 시리즈를 만들 때는 `pd.Series` 안에 리스트 혹은 넘파이 어레이를 넣어줍니다. 출력해보면 하나의 열처럼 세로로 값들이 떨어지는 것을 볼 수 있습니다. 시리즈가 하나의 컬럼이라는 것을 꼭 기억해주세요.
pd.Series([3,2,0,1])
pd.Series([3,0,2,1]).rename("apples")
# ### apply
데이터프레임.apply(함수, axis=1) # lambda 함수와 조합해서 사용하는 경우가 많습니다
# `apply` 메소드는 판다스 시리즈에서 꼭 짚고 넘어가야 할 부분입니다. 앞에서 다루었던 `map` 함수와 유사한 기능을 하는 메소드로, 시리즈의 각 요소들에 주어진 함수를 적용합니다. `for` 루프보다 빠르고 문법도 간결해서 많이 쓰이는 메소드입니다. 개인적으로 숫자 시리즈보다는 문자열 시리즈에서 유용하게 사용합니다.
mySeries = pd.Series(["서울 서대문구", "서울 중랑구", "서울 강남구"])
[x.split(" ")[1] for x in mySeries]
mySeries.apply(lambda x: x.split(" ")[1])
# 위의 예시 코드는 "서울시 ~구" 형태로 이루어진 문자열 시리즈에서 구만을 뽑아내는 조작입니다. `lambda` 함수는 문자열을 공백을 기준으로 나누고, 반환된 리스트에서 `1`번 인덱스의 값을 반환하도록 되어 있습니다. 이 함수를 `apply` 메소드와 함께 활용하면, 함수를 시리즈의 각 요소에 적용한 결과를 얻을 수 있습니다.
#
# **예제 2.1. `apply` 메소드를 활용하여 `dateTime` 시리즈 각 날짜에 해당하는 요일 시리즈를 만들어주세요.**
dateTime = pd.date_range("2020-01-01", "2020-01-30").to_series() # 날짜 생성
dateTime[0].weekday() # 힌트
# **풀이**
#
# 날짜 데이터가 생소하겠지만 크게 신경쓸 필요는 없습니다. `weekday` 메소드가 요일을 반환한다는 사실을 알고 있으므로, 먼저 날짜 데이터를 받아서 요일을 반환하는 람다 함수를 짭니다. 람다를 사용해서 메소드를 함수화하였으므로 이제 `apply`와 함께 사용할 수 있습니다.
lambda x: x.weekday()
# `dateTime` 시리즈의 `apply` 메소드에 만들어둔 람다 함수를 넣어서 실행합니다.
# %time dateTime.apply(lambda x: x.weekday()) # 출력 생략
# 앞에서 배운 리스트 컴프리헨션을 통해서도 똑같이 구현할 수 있습니다. 개인적으로 어떤 방법이 더 빠른지는 확신은 못하겠지만, 데이터가 큰 경우 리스트 컴프리헨션 쪽이 더 잘 돌아갔던 것 같습니다.
# %time pd.Series([x.weekday() for x in dateTime], dateTime)
# ### 데이터프레임
#
# 데이터프레임을 만들 때에는 `pd.DataFrame` 안에 딕셔너리를 넣어줍니다. 이 때 딕셔너리의 키는 컬럼의 이름어야 하고, 값은 컬럼에 실제로 들어갈 요소들이어야 합니다. 행 인덱스가 필요하다면 `index` 파라미터에 행 인덱스로 사용할 값들을 넣어주면 됩니다.
data = {
'apples': [3, 2, 0, 1],
'oranges': [0, 3, 7, 2]
}
purchases = pd.DataFrame(data, index=['June', 'Robert', 'Lily', 'David'])
purchases
# **예제 2.2. 아래와 같은 데이터프레임을 만들어주세요.**
# **풀이**
#
# 데이터프레임은 학점, 학년이라는 두 개의 컬럼으로 이루어져 있습니다. 따라서 딕셔너리의 키 값은 `"학점"`,`"학년"`이 됩니다. 학점의 값은 `["A","B","C","A"]` 이고, 학년의 값은 `[3,3,1,2]`입니다. 따라서 데이터프레임을 이루는 딕셔너리는 다음과 같이 만들어졌을 것입니다.
data = {"학점":["A","B","C","A"], "학년":[3,3,1,2]}
# 이 결과를 `pd.DataFrame`으로 감싸주면 데이터프레임이 완성됩니다.
pd.DataFrame(data)
# ## 2.2. 판다스 인덱싱
#
# 판다스 인덱싱에는 `loc`, `iloc` 을 사용합니다. 대괄호 안에 `[행,열]` 순서로 적어주는 것은 동일하지만, `loc`은 이름을 사용하는 반면 `iloc`은 번호(좌표)를 사용합니다.
#
# `df.loc[]`: $\textbf{loc}$ates by name, 즉 행/열 이름으로 인덱싱
#
# `df.iloc[]`: $\textbf{loc}$ates by numerical $\textbf{i}$ndex, 즉 행/열 번호로 인덱싱
purchases
purchases.loc[행,열]
purchases.iloc[행,열]
# ## 2.2. 외부 데이터 읽어오기
#
# 판다스로 데이터프레임을 직접 만들 수도 있지만, 대부분의 경우에는 `csv`, `xlsx`, `json` 등 파일로 저장된 외부 데이터를 읽어와 작업을 하게 됩니다. **앞으로 가장 많이 접하게 될 `csv` 파일을 읽어오는 함수는 `pd.read_csv` 입니다.** 가장 먼저 파일이 위치한 경로를 입력하고, `encoding` 매개변수에 파일의 인코딩을 지정해줍니다. `utf-8`이 기본 옵션이므로 `utf-8` 파일을 읽어올 때에는 따로 인코딩을 지정할 필요가 없습니다.
data = pd.read_csv("../data/dataset_for_analysis.csv", encoding="ANSI")
data
# **예제 2.2. 아래 링크의 csv 파일을 판다스로 읽어오세요.**
#
# https://raw.githubusercontent.com/agconti/kaggle-titanic/master/data/train.csv
# **풀이**
#
# 판다스는 웹에 존재하는 파일을 즉시 읽어올 수도 있습니다.
titanic = pd.read_csv(
"https://raw.githubusercontent.com/agconti/kaggle-titanic/master/data/train.csv"
)
titanic.head()
# ### 경로와 인코딩
#
# **절대경로 상대경로, 인코딩**이 무엇인지 아시면 이 내용은 건너뛰셔도 됩니다.
#
# #### 경로
#
# .|.
# ---|---
# |
#
# **경로는 파일이 존재하는 위치를 뜻하며 절대 경로, 상대 경로로 나누어 생각할 수 있습니다.** 절대 경로는 최상위 디렉토리인 `C:` 부터 시작하는 경로인 반면, 상대 경로는 특정 디렉토리를 기준으로 상대적으로 표현된 경로입니다. 위에서 우리가 읽어들인 `dataset_for_analysis.csv` 파일을 예로 들어 보겠습니다. 먼저 해당 파일이 존재하는 위치로 가서, 파일의 속성을 확인해봅니다. 그러면 위와 같은 화면을 확인할 수 있을 것입니다. 이 때 Location 항목에 표시되는 것이 파일이 위치한 디렉토리의 절대 경로이며, 파일의 절대 경로는 `C:\Users\Administrator\pyfords\data\dataset_for_analysis.csv` 가 됩니다.
#
#
# # 계층적으로 표현한 디렉토리 구조
C:/
Users/
Administrator/
pyfords/
data/
dataset_for_analysis.csv
gapminder.csv
iris.csv
# 하지만 위의 예시 코드에서는 입력한 경로는 `C:` 부터 출발하지 않는 **상대 경로**입니다. 상대 경로를 통해서 파일을 읽어올 수 있었다는 것은 분명히 기준 경로가 존재한다는 뜻입니다. 파일의 절대 경로와 상대 경로를 비교해보면, 파이썬이 인식하는 기준 경로가 `C:\Users\Administrator\pyfords\` 임을 알 수 있습니다. 만약 주피터 노트북을 사용하고 계신다면, 현재 열려있는 노트북이 실행된 위치가 기준 디렉토리가 됩니다. 파이썬 표준 패키지인 `os`를 이용하면 현재 작업중인 기준 디렉토리를 확인할 수 있습니다.
import os
os.listdir("../data")
pd.read_csv("../data/dataset_for_analysis.csv", encoding="ANSI")
utf-8
cp949
euc-kr
# #### 인코딩
#
# 컴퓨터는 숫자만을 저장할 수 있습니다. 따라서 우리가 사용하는 문자들을 컴퓨터에 파일로 저장하기 위해서는 문자를 코드로 바꾸어주는 일종의 변환이 필요합니다. 이러한 변환 과정을 인코딩이라고 합니다. 반대로 인코딩된 자료를 해독하여 다시 사람이 이해할 수 있는 문자로 바꾸어주는 것을 디코딩이라고 합니다.
#
# 문제는 인코딩에 하나의 통일된 체계가 존재하지 않는다는 점입니다. 예를 들어 `euc-kr`이라는 규칙으로 인코딩된 파일을 `utf-8`이라는 규칙으로 디코딩하려고 한다면 어떻게 될까요? 당연히 정상적으로 해독이 되지 않을 것입니다. 흔히 말해 '글자가 깨지는' 현상이 나타나거나, 혹은 아예 파일을 읽어오지 못할 것입니다. 이러한 경우에는 컴퓨터가 파일의 인코딩 규칙에 맞게 디코딩을 할 수 있도록, 파일이 어떠한 규칙으로 인코딩되었는지 알려주어야 합니다.
#
# # 3. 데이터프레임 조작
#
# 이제 본격적으로 판다스 데이터프레임을 조작하는 방법을 배워보도록 하겠습니다. 크게 아래와 같은 기능들을 학습할 것입니다.
#
# - 변수 선택: 데이터프레임의 특정 칼럼을 추출
# - 필터링: 특정 기준을 만족하는 행을 추출
# - 정렬: 행을 다시 정렬
# - 변수 추가: 새로운 변수를 추가
# - 변수 요약: 변수를 값(스칼라)으로 요약
# - 그룹별로 조작: 데이터프레임을 범주형 변수의 수준별로 묶어서 조작
#
# ## 3.1. 변수 선택
#
# **변수를 선택한다는 것은 결국 데이터프레임에서 특정 열만을 뽑아내는 작업**입니다. 변수 선택은 다음과 같은 두 가지 방법으로 실행할 수 있습니다. 속성을 사용하는 것이 타이핑은 편하지만, 변수명에 띄어쓰기나 점`.` 등이 포함되거나 기타 속성과 겹칠 경우 사용할 수 없습니다. 대괄호와 문자열을 사용해서 열을 인덱싱하는 방법은 어떠한 경우에도 사용할 수 있습니다.
# **열 인덱싱**
data['당해 GDP 성장율']
# **데이터프레임의 속성**
data.국가
# ### 복수 개의 열 인덱싱
#
# **복수 개의 열을 뽑아낼 때에는 대괄호`[]` 안에 열 이름의 리스트를 넣어줍니다.**
data[['당해 GDP 성장율', '국가']]
# **예제 3.1. 판다스 데이터프레임의 `columns` 속성을 통해 데이터프레임의 컬럼 이름들에 접근할 수 있습니다. 위에서 읽어온 `titanic` 데이터의 컬럼 이름을 확인하고, 원하는 컬럼을 추출해보세요.**
titanic.columns
# ## 3.2. 필터링
#
# ### 필터링의 원리: 마스킹
#
#
# 필터링은 특정 변수에 대해 원하는 조건을 만족하는 행을 걸러내는 조작입니다. 위에서 만들어둔 `purchases` 데이터를 예로 들어보겠습니다. `purchases` 데이터에서 사과를 1개 이상 구매한 사람들만을 걸러내려고 합니다. 이를 위해 먼저 **각 행들에 대해서 `apples` 컬럼의 값이 0보다 큰지를 판단합니다. 이 결과는 `[True, True, False, True]` 입니다. 이 결과를 행에 덮어씌워서, `True`만을 남기고 `False`는 떨어뜨립니다.**
#
#
#
# **purchases 데이터프레임**
#
# Index|apples|oranges
# ---|---|---
# June|3|0
# Robert|2|3
# Lily|0|7
# David|1|2
#
#
# **마스킹: 사과를 샀는가?**
#
# Index|apples|oranges|사과를 샀는가?
# ---|---|---|---
# June|3|0|True
# Robert|2|3|True
# ~~Lily~~|~~0~~|~~7~~|~~False~~
# David|1|2|True
#
# 위 과정을 판다스로 구현해보겠습니다. 먼저 사과 구매에 해당하는 변수인 `apples`를 선택해야 합니다. 다음으로 `purchases.apples` 에 대해서, 각각의 값이 `0` 보다 큰지를 판단해주면 됩니다. 연산을 실행한 결과는 `[True, True, False, True]` 의 불리언 시리즈입니다. 이러한 연산이 가능한 것은 넘파이의 브로드캐스팅 때문입니다.
# **1. 변수 선택**
purchases.apples
# **2. 조건 판단: 브로드캐스팅**
purchases.apples >= 1
# **3. 마스킹**
purchases
purchases[[False, True, False, True]]
# $$ Numpy \space Broadcasting:
# \begin{pmatrix}
# 3 > 0\\
# 2 > 0\\
# 0 > 0\\
# 1 > 0\\
# \end{pmatrix}
# =
# \begin{pmatrix}
# True\\
# True\\
# False\\
# True\\
# \end{pmatrix}
# $$
#
#
#
# 이제 이 결과를 통해 행을 인덱싱해주면 끝입니다. 데이터프레임 옆에 대괄호`[]`를 적고, 대괄호 안에 완성된 불리언 시리즈를 넣어주면, 조건이 `True`인 행들만 추출됩니다. 데이터 프레임 위에 `True`, `False`로 된 시리즈를 덧붙여서 `True`만 남긴다고 생각하시면 됩니다.
#
#
# **예제 3.2. 타이타닉 데이터에서 성별이 여성인 데이터만 골라낸 후, 생존 여부 변수를 선택하세요.**
titanic.Sex.unique()
titanic[titanic.Sex == "female"].Survived
# **풀이**
#
# 타이타닉 데이터에서 성별과 생존 여부를 나타내는 컬럼은 `Sex`와 `Survived` 입니다. `titanic.Sex=='female'` 이라는 조건을 걸어서 데이터프레임을 필터링해주고, 필터링된 데이터에서 `Survived` 변수를 선택해주면 끝입니다.
titanic[titanic.Sex=='female'].Survived
# ### 다중 조건으로 필터링
#
# 다중 조건을 걸 때에도, 문제는 결국 하나의 불리언 시리즈를 만들어내는 것입니다. `&`, `|` 의 논리 연산을 적절하게 활용하면 어렵지 않게 다중 조건 필터링을 구현할 수 있습니다. `purchases` 데이터의 예시를 다시 보겠습니다. 이번에는 사과와 오렌지를 모두 구매한 사람들을 골라내려고 합니다. 사과와 오렌지에 대해서 각각 0보다 큰지를 판단합니다. 이후 두 개의 조건을 연결해주면 됩니다. 우리의 관심은 **"사과와 오렌지를 모두 샀는가?"** 이므로 논리연산으로는 `&`에 해당합니다.
# **purchases 데이터프레임**
#
# Index|apples|oranges|사과를 샀는가?|오렌지를 샀는가?|사과와 오렌지를 모두 샀는가?
# ---|---|---|---|---|---
# ~~June~~|~~3~~|~~0~~|~~True~~|~~False~~|~~False~~
# Robert|2|3|True|True|True
# ~~Lily~~|~~0~~|~~7~~|~~False~~|~~True~~|~~False~~
# David|1|2|True|True|True
#
# 위와 같은 조작을 판다스로 구현해보겠습니다. 먼저 `apples` 와 `oranges`에 대해서 각각 0보다 큰지를 판단합니다. 이 두 개의 조건은 `&` 연산으로 묶어주고, 마찬가지로 대괄호 안에 넣어서 마스킹해주면 끝입니다. 역시 넘파이의 연산 특성으로 인해서 같은 위치에 있는 요소들끼리 `&` 연산이 실행됩니다.
# **1. 사과를 샀는가?**
purchases.apples > 0
# **2. 오렌지를 샀는가?**
purchases.oranges > 0
#
# **3. 사과와 오렌지를 샀는가?**
(purchases.apples > 0) & (purchases.oranges > 0)
# **4. 마스킹**
purchases[(purchases.apples > 0) & (purchases.oranges > 0)]
#
# $$
# \begin{pmatrix}
# True \space and \space False \\
# True \space and \space True \\
# False \space and \space True \\
# True \space and \space True
# \end{pmatrix}
# =
# \begin{pmatrix}
# False \\
# True \\
# False \\
# True
# \end{pmatrix}
# $$
#
# **예제 3.3. 타이타닉 데이터에서 30세 이상 남성이거나 30세 이하 여성인 행들을 골라내세요.**
#
# **풀이**
#
# 주어진 조건을 논리연산자, 비교연산자를 사용해서 표현하면 다음과 같습니다.
#
# 1. (30세 이상 남성) or (30세 이하 여성)
# 2. (30세 이상 and 남성) or (30세 이하 and 여성)
# 3. {(나이 >= 30) and (성별 == 남성)} or {(나이 <= 30) and (성별 == 여성)}
#
# 이 결과를 코드 한 줄로 적으면 너무 길어지므로, 두 개의 조건으로 분해해서 적어보겠습니다. `cond1` 은 30세 이상 남성에 해당하는 조건이고, `cond2`는 30세 이하 여성에 해당하는 조건입니다. 두 개의 조건을 `|` 으로 묶어주고 마스킹해주면 끝입니다.
cond1 = (titanic.Age >= 30) & (titanic.Sex == "male")
cond2 = (titanic.Age <= 30) & (titanic.Sex == "female")
cond1 | cond2
titanic[cond1 | cond2]
# ## 3.3. 정렬
#
# `sort_values` 메소드는 시리즈나 데이터프레임을 크기 순으로 정렬합니다. 시리즈는 건너뛰고, 데이터프레임 정렬에 대해서만 다루도록 하겠습니다. `sort_values(컬럼)` 과 같이 적으면 주어진 컬럼의 오름차순으로 데이터프레임을 정렬합니다. `ascending=False`를 전달하면 내림차순으로 정렬합니다.
purchases.sort_values("apples")
purchases.sort_values("apples", ascending=False)
# **예제 3.4. 타이타닉 데이터에서 나이가 30세 이상인 행들만을 걸러내고 나이 오름차순으로 정렬해주세요.**
#
# **풀이**
titanic[titanic.Age >= 30].sort_values("Age")
# ## 3.4. 변수 추가
#
# `assign` 메소드는 데이터프레임에 새로운 컬럼을 추가합니다. 다른 컬럼들의 정보를 반영하여 새로운 컬럼을 추가하거나, 기존에 가지고 있던 컬럼을 변형하는 경우에 사용할 수 있습니다. 예를 들어 `purchaes` 데이터프레임에서 총 과일 구매 개수를 새로운 열로 추가하려고 한다면, 다음과 같이 쓸 수 있습니다. **새롭게 추가하고자 하는 컬럼의 이름은 문자열이 아니라 변수처럼 써주어야 합니다!** `assign` 메소드는 데이터프레임 직접 변경시키지 않으므로, 결과를 저장하려면 다시 할당을 해주어야 합니다.
데이터프레임.assign(새변수명 = 데이터)
purchases.assign(total = purchases.apples + purchases.oranges)
# **예제 3.4. 사과가 하나에 500원, 오렌지가 하나에 300원이라고 가정합니다. 고객별로 얼마의 매출이 발생했는지는 나타내는 컬럼을 데이터프레임에 추가해주세요.**
# **풀이**
#
# 위에서 다루었던 예시 코드에 간단한 곱하기 연산을 추가해주면 됩니다. 사과는 500원이므로 500을 곱해주고, 오렌지는 300원이므로 300을 곱해줍니다.
purchases.assign(amount = purchases.apples * 500 + purchases.oranges * 300)
# ## 3.5. 변수 요약
#
# 사람은 평균, 표준편차, 중위수 등의 요약된 통계량을 통해서 데이터를 더 잘 이해할 수 있습니다. 판다스 시리즈와 데이터프레임은 데이터를 요약하는 다양한 메소드를 제공합니다. 메소드이므로 꼭 `()`와 함께 사용합니다.
#
# 메소드|시리즈|데이터프레임
# ---|---|---
# mean|시리즈의 평균을 반환|모든 숫자 컬럼의 평균을 반환
# std|시리즈의 표준편차를 반환|모든 숫자 컬럼의 평균을 반환
# median|시리즈의 중위수를 반환|모든 숫자 컬럼의 중위수를 반환
# min|시리즈의 최소값을 반환|모든 숫자 컬럼의 최소값을 반환
# max|시리즈의 최대값을 반환|모든 숫자 컬럼의 최대값을 반환
# describe|관측값 수, 평균,표준편차, 각 분위수 반환|관측값 수, 평균,표준편차, 각 분위수 반환
#
#
# **시리즈**
purchases.apples.mean()
purchases.apples.std()
purchases.apples.median()
purchases.apples.min()
purchases.apples.max()
# **데이터프레임**
purchases.describe()
# ## 3.6. 그룹별 조작
#
# ### groupby 기초
#
# 
#
# `groupby` 메소드는 말 그대로 데이터를 그룹별로 나누어 조작할 수 있도록 만들어주며, 데이터가 범주형 변수를 포함할 때 유용합니다. 위 그림은 A, B, C 그룹을 구분하고 그룹별로 데이터의 합을 집계하는 과정을 나타냅니다. 위 그림에서 나타난 조작을 판다스로 구현해보겠습니다.
data = pd.DataFrame(
{"key":["A","B","C","A","B","C"], "data":[1,2,3,4,5,6]}
)
data
# `groupby` 메소드의 괄호 안에 그룹으로 나누어줄 컬럼명을 전달합니다. `groupby` 메소드만를 실행하면 `DataFrameGroupBy` 객체가 반환됩니다. 값을 명시적으로 보여주지는 않지만, 그룹별로 쪼개진 상태라고 상상해주시면 됩니다. 이 상태에서 `sum` 메소드를 사용하면 합계를 구할 수 있습니다. `sum` 뿐만 아니라 위에서 다루었던 데이터 요약 메소드들을 모두 적용할 수 있습니다.
data.groupby("key") # 쪼개진 상태 !
data.groupby("key").sum() # 집계 메소드 적용
# ### 변수 선택
#
# `DataFrameGroupBy` 객체에서도 변수를 선택할 수 있으며, 데이터프레임에서 하던 것과 동일합니다. 먼저 `purchases` 데이터에 결혼 여부 컬럼을 추가하고, 결혼 여부에 따라 사과, 오렌지의 평균 구매 개수를 계산해봅니다.
#
# **데이터프레임에 변수 추가**
purchases = purchases.assign(married = [0,1,1,0])
purchases.groupby("married").mean()
# 만약 우리가 사과에는 관심이 없고, 오렌지에만 관심이 있다면 어떨까요? 먼저 평균을 집계한 이후, 그 결과에서 `oranges`만을 뽑아낼 수 있습니다. 하지만 이러한 코드는 데이터가 커진다면 시간을 상당히 낭비하는 코드일 수 있습니다. 이런 경우에는 먼저 변수를 선택한 후에 집계를 하는 편이 효율적입니다.
#
#
#
# **집계 후 변수 선택**
purchases.groupby("married").mean().oranges
# **변수 선택 후 집계: 계산 절약**
purchases.groupby("married").oranges.mean()
# ### 여러 변수로 그루핑
#
# 여러 변수를 통해 그룹을 분리하는 것도 가능합니다. `groupby` 메소드에 변수들의 리스트를 전달해주면 됩니다. `purchases` 데이터에 새로운 컬럼을 추가해보겠습니다.
purchases = purchases.assign(graduate = [True, True, True, False])
purchases.groupby(["married","graduate"]).mean()
# **예제 3.5. 타이타닉 데이터에서 선실 등급별/성별 생존률을 계산하세요. 선실 등급을 나타내는 변수는 `Pclass` 입니다.**
# **풀이**
#
# `groupby` 메소드에 그룹지을 변수들의 리스트, 즉 `['Pclass','Sex']`을 전달해줍니다. 생존 여부가 0과 1로 코딩되어 있으므로 평균을 통해서 각 그룹의 생존률을 계산할 수 있습니다. 따라서 그룹바이 객체에서 `Survived` 변수를 선택하고, 평균을 구해줍니다.
titanic.groupby(["Pclass","Sex"]).Survived.mean()
# # 4. 복수의 데이터프레임 합치기
#
# ## 4.1. pd.merge
#
# 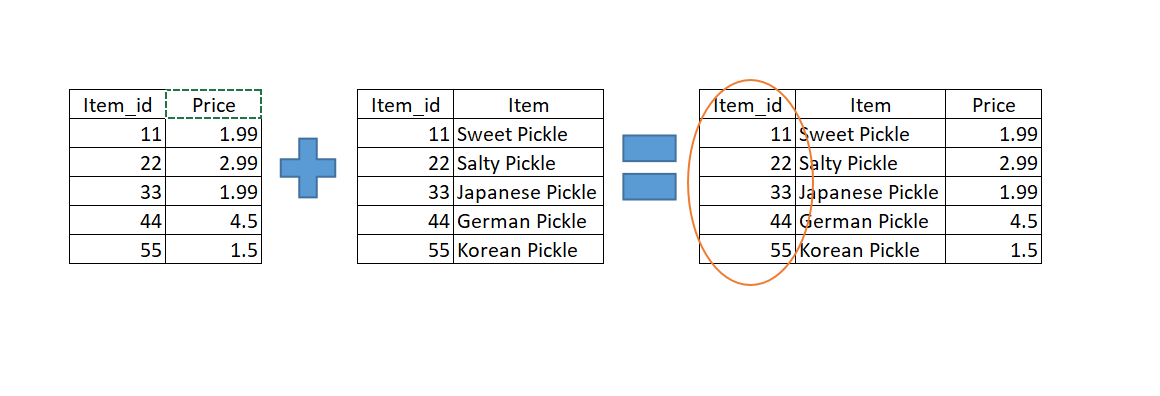
#
# 마지막으로, 복수의 데이터프레임을 합치는 방법을 배워보겠습니다. 예를 들어 위와 같은 두 개의 데이터프레임은 `item_id`라는 컬럼을 기준으로 이어붙일 수 있을 것입니다. 이렇게 기준 열을 통해서 복수의 데이터를 병합하는 조작을 SQL에서는 조인이라고 부릅니다. 판다스에서 이러한 조작은 `pd.merge` 함수를 통해서 실행할 수 있습니다. **아무런 인자 없이 `pd.merge`를 실행할때는 양쪽 데이터에 공통된 이름의 컬럼이 존재해야 하며, 양쪽 컬럼의 자료형이 일치해야 합니다.**
leftData = pd.DataFrame({"item_id":[11,22,33,44,55],"price":[1.99,2.99,1.00,4.5,1.5]})
rightData = pd.DataFrame({
"item_id":[11,22,33,44,55],
"item":["sweet","salty","japanese","german","korean"]
})
pd.merge(leftData,rightData)
# 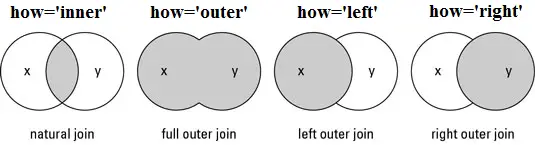
#
# 위에서는 왼쪽 데이터와 오른쪽 데이터의 행 개수가 동일했지만, 그렇지 않은 경우에도 조인이 가능합니다. 위 그림을 보면, 크게 네 가지 방식의 조인이 존재하는 것을 알 수 있습니다.
#
# 1. Inner join: 양쪽 데이터 모두에 존재하는 행들만을 남기는 조인(pd.merge 기본 옵션!)
# 2. Left join: 왼쪽 데이터에 존재하는 행들만을 남기는 조인
# 3. Right join: 오른쪽 데이터에 존재하는 행들만을 남기는 조인
# 4. Outer join: 양쪽 데이터에 존재하는 행들을 모두 남기는 조인
pd.merge(왼쪽데이터, 오른쪽데이터, how='조인방식')
# ### inner join
#
# 위의 코드를 약간 수정하여 `leftData`를 3개의 행으로, `rightData`를 4개의 행으로 줄여주었습니다. 데이터를 이렇게 수정해주면, 왼쪽 데이터와 오른쪽 데이터의 교집한은 아이디 11번, 22번 상품 뿐입니다. 따라서 inner join을 실행하면 11번, 22번 상품만 남게 됩니다. Inner join이 `pd.merge`의 기본 옵션이기 때문에 별도로 인자를 전달해줄 필요가 없습니다.
leftData = pd.DataFrame({"item_id":[11,22,33],"price":[1.99,2.99,1.00]})
rightData = pd.DataFrame({"item_id":[22,33,44,55],"item":["salty","japanese","german","korean"]})
pd.merge(leftData,rightData)
# ### left join, right join
#
# Left join은 왼쪽 데이터에 존재하는 데이터를 모두 남기는 조인입니다. 아래 예시 코드를 보겠습니다. 왼쪽에 존재하는 데이터들이 모두 보존되는 것을 확인할 수 있습니다. 하지만 11번 상품은 오른쪽 데이터에 존재하지 않는 행이기 때문에, `item` 컬럼에 `NaN`이 할당되었습니다. Right join 역시 같은 원리로 작동하기 때문에 설명은 생략하겠습니다.
pd.merge(leftData,rightData,how='left')
pd.merge(leftData,rightData,how='right')
# ### outer join
#
# Outer join은 양쪽의 데이터를 모두 남기는 조인입니다. 아래 에시 코드를 보겠습니다. 양쪽 데이터에 들어있는 행들이 모두 보존되는 것을 확인할 수 있습니다. 역시 서로 대응되는 행이 없는 경우에는 `NaN`이 할당됩니다.
pd.merge(leftData,rightData,how='outer')
# ### 기타 경우의 조인
#
# 같은 이름의 컬럼이 존재하지 않거나, 복수의 열을 기준으로 조인을 실행할 경우에는 추가적으로 인자를 전달해주어야 합니다. 다양한 경우들을 여기에서 전부 다룰 수는 없기 때문에, 간단한 문법만 짚고 넘어가겠습니다. 더 자세한 정보를 원하시면 [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.merge.html)을 참조하시기 바랍니다.
pd.merge(data1, data2, left_on="왼쪽컬럼명", right_on="오른쪽컬럼명") # 컬럼 이름이 일치하지 않는 경우
pd.merge(data1, data2, on = ['기준컬럼1','기준컬럼2', ...]) # 복수의 컬럼을 기준으로 merge
# ## 4.2. concat
#
# 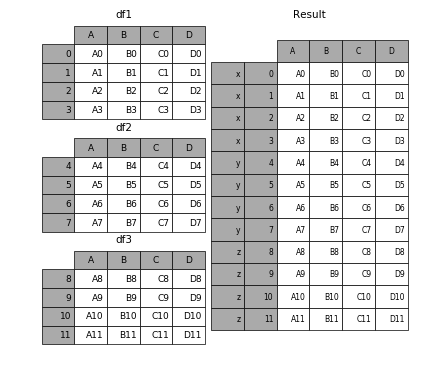
#
# `pd.merge`가 복수의 데이터를 옆으로 합쳐주었다면, `pd.concat`은 주로 복수의 데이터를 위아래로 합치는 경우에 사용합니다(양옆으로 합칠 때도 사용할 수 있습니다). 위와 같이 동일한 컬럼을 공유하는 복수의 데이터를 합쳐주는 경우가 대표적인 예시입니다. 아래 예시 코드를 보면 쉽게 이해할 수 있을 것입니다. `[]` 안에 데이터들을 넣어주시면 됩니다. `pd.concat`은 개별 데이터가 커질수록 느려지기 때문에 절대절대 `for` 문과 함께 사용하시면 안됩니다! 리스트 안에 복수의 데이터를 집어넣은 후, 한 번에 합쳐주셔야 그나마 빠르게 작업을 할 수 있습니다.
pd.concat([데이터1, 데이터2, 데이터3, ...])
data1 = pd.DataFrame({"height":[154, 184, 176], "weight":[54, 84, 76]})
data2 = pd.DataFrame({"height":[167, 170, 163, 165], "weight":[67, 70, 63, 65]})
pd.concat([data1, data2])
# # 5. 기타 판다스 함수 및 메소드, 속성
#
# ## 5.1. 판다스 함수
#
# 함수|기능
# ---|---
# pd.date_range|날짜 구간 생성
# pd.to_datetime|시리즈의 자료형을 timestamp로 변경
# pd.get_dummies|시리즈를 더미 인코딩
#
# ## 5.2. 기타 데이터프레임 메소드
#
# 메소드|기능
# ---|---
# df.astype|특정 열의 데이터 타입 변경
# df.isnull()|값이 NA인지 판단
# df.dropna()|NA를 모두 드랍
# df.set_index()|열을 인덱스로
# df.reset_index()|인덱스를 초기화
# df.duplicated()|값이 중복되었는지 판단
# df.drop_duplicates()|중복된 행 제거
# df.pivot()|피봇 테이블
#
# ## 5.3. 데이터프레임 속성
#
# 메소드|내용
# ---|---
# df.columns|데이터프레임의 열 이름
# df.values|데이터프레임의 값이 담긴 넘파이 어레이
# df.dtypes|데이터프레임의 데이터 타입
| notebook/page3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import numpy as np
import holoviews as hv
from holoviews import opts
hv.extension('bokeh')
# -
# When working with the bokeh backend in HoloViews complex interactivity can be achieved using very little code, whether that is shared axes, which zoom and pan together or shared datasources, which allow for linked cross-filtering. Separately it is possible to create custom interactions by attaching LinkedStreams to a plot and thereby triggering events on interactions with the plot. The Streams based interactivity affords a lot of flexibility to declare custom interactivity on a plot, however it always requires a live Python kernel to be connected either via the notebook or bokeh server. The ``Link`` classes described in this user guide however allow declaring interactions which do not require a live server, opening up the possibility of declaring complex interactions in a plot that can be exported to a static HTML file.
#
# ## What is a ``Link``?
#
# A ``Link`` defines some connection between a source and target object in their visualization. It is quite similar to a ``Stream`` as it allows defining callbacks in response to some change or event on the source object, however, unlike a Stream, it does not transfer data between the browser and a Python process. Instead a ``Link`` directly causes some action to occur on the ``target``, for JS based backends this usually means that a corresponding JS callback will effect some change on the target in response to a change on the source.
#
# One of the simplest examples of a ``Link`` is the ``DataLink`` which links the data from two sources as long as they match in length, e.g. below we create two elements with data of the same length. By declaring a ``DataLink`` between the two we can ensure they are linked and can be selected together:
# +
from holoviews.plotting.links import DataLink
scatter1 = hv.Scatter(np.arange(100))
scatter2 = hv.Scatter(np.arange(100)[::-1], 'x2', 'y2')
dlink = DataLink(scatter1, scatter2)
(scatter1 + scatter2).opts(
opts.Scatter(tools=['box_select', 'lasso_select']))
# -
# If we want to display the elements subsequently without linking them we can call the ``unlink`` method:
# +
dlink.unlink()
(scatter1 + scatter2)
# -
# Another example of a link is the ``RangeToolLink`` which adds a RangeTool to the ``source`` plot which is linked to the axis range on the ``target`` plot. In this way the source plot can be used as an overview of the full data while the target plot provides a more detailed view of a subset of the data:
# +
from holoviews.plotting.links import RangeToolLink
data = np.random.randn(1000).cumsum()
source = hv.Curve(data).opts(width=800, height=125, axiswise=True, default_tools=[])
target = hv.Curve(data).opts(width=800, labelled=['y'], toolbar=None)
rtlink = RangeToolLink(source, target)
(target + source).opts(merge_tools=False).cols(1)
# -
# ## Advanced: Writing a ``Link``
#
# A ``Link`` consists of two components the ``Link`` itself and a ``LinkCallback`` which provides the actual implementation behind the ``Link``. In order to demonstrate writing a ``Link`` we'll start with a fairly straightforward example, linking an ``HLine`` or ``VLine`` to the mean value of a selection on a ``Scatter`` element. To express this we declare a ``MeanLineLink`` class subclassing from the ``Link`` baseclass and declare ``ClassSelector`` parameters for the ``source`` and ``target`` with the appropriate types to perform some basic validation. Additionally we declare a ``column`` parameter to specify which column to compute the mean on.
# +
import param
from holoviews.plotting.links import Link
class MeanLineLink(Link):
column = param.String(default='x', doc="""
The column to compute the mean on.""")
_requires_target = True
# -
# Now we have the ``Link`` class we need to write the implementation in the form of a ``LinkCallback``, which in the case of bokeh will be translated into a [``CustomJS`` callback](https://bokeh.pydata.org/en/latest/docs/user_guide/interaction/callbacks.html#userguide-interaction-jscallbacks). A ``LinkCallback`` should declare the ``source_model`` we want to listen to events on and a ``target_model``, declaring which model should be altered in response. To find out which models we can attach the ``Link`` to we can create a ``Plot`` instance and look at the ``plot.handles``, e.g. here we create a ``ScatterPlot`` and can see it has a 'cds', which represents the ``ColumnDataSource``.
# +
renderer = hv.renderer('bokeh')
plot = renderer.get_plot(hv.Scatter([]))
plot.handles.keys()
# -
# In this case we are interested in the 'cds' handle, but we still have to tell it which events should trigger the callback. Bokeh callbacks can be grouped into two types, model property changes and events. For more detail on these two types of callbacks see the [Bokeh user guide](https://bokeh.pydata.org/en/latest/docs/user_guide/interaction/callbacks.html#userguide-interaction-jscallbacks).
#
# For this example we want to respond to changes to the ``ColumnDataSource.selected`` property. We can declare this in the ``on_source_changes`` class atttribute on our callback. So now that we have declared which model we want to listen to events on and which events we want to listen to, we have to declare the model on the target we want to change in response.
#
# We can once again look at the handles on the plot corresponding to the ``HLine`` element:
plot = renderer.get_plot(hv.HLine(0))
plot.handles.keys()
# We now want to change the ``glyph``, which defines the position of the ``HLine``, so we declare the ``target_model`` as ``'glyph'``. Having defined both the source and target model and the events we can finally start writing the JS callback that should be triggered. To declare it we simply define the ``source_code`` class attribute. To understand how to write this code we need to understand how the source and target models, we have declared, can be referenced from within the callback.
#
# The ``source_model`` will be made available by prefixing it with ``source_``, while the target model is made available with the prefix ``target_``. This means that the ``ColumnDataSource`` on the ``source`` can be referenced as ``source_source``, while the glyph on the target can be referenced as ``target_glyph``.
#
# Finally, any parameters other than the ``source`` and ``target`` on the ``Link`` will also be made available inside the callback, which means we can reference the appropriate ``column`` in the ``ColumnDataSource`` to compute the mean value along a particular axis.
#
# Once we know how to reference the bokeh models and ``Link`` parameters we can access their properties to compute the mean value of the current selection on the source ``ColumnDataSource`` and set the ``target_glyph.position`` to that value.
#
# A ``LinkCallback`` may also define a validate method to validate that the Link parameters and plots are compatible, e.g. in this case we can validate that the ``column`` is actually present in the source_plot ``ColumnDataSource``.
# +
from holoviews.plotting.bokeh import LinkCallback
class MeanLineCallback(LinkCallback):
source_model = 'selected'
source_handles = ['cds']
on_source_changes = ['indices']
target_model = 'glyph'
source_code = """
var inds = source_selected.indices
var d = source_cds.data
var vm = 0
if (inds.length == 0)
return
for (var i = 0; i < inds.length; i++)
vm += d[column][inds[i]]
vm /= inds.length
target_glyph.location = vm
"""
def validate(self):
assert self.link.column in self.source_plot.handles['cds'].data
# -
# Finally we need to register the ``MeanLineLinkCallback`` with the ``MeanLineLink`` using the ``register_callback`` classmethod:
MeanLineLink.register_callback('bokeh', MeanLineCallback)
# Now the newly declared Link is ready to use, we'll create a ``Scatter`` element along with an ``HLine`` and ``VLine`` element and link each one:
# +
options = opts.Scatter(
selection_fill_color='firebrick', alpha=0.4, line_color='black', size=8,
tools=['lasso_select', 'box_select'], width=500, height=500,
active_tools=['lasso_select']
)
scatter = hv.Scatter(np.random.randn(500, 2)).opts(options)
vline = hv.VLine(scatter['x'].mean()).opts(color='black')
hline = hv.HLine(scatter['y'].mean()).opts(color='black')
MeanLineLink(scatter, vline, column='x')
MeanLineLink(scatter, hline, column='y')
scatter * hline * vline
# -
# Using the 'box_select' and 'lasso_select' tools will now update the position of the HLine and VLine.
| examples/user_guide/Linking_Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parsing and Grammar
# ## Parsing
# Sentence $\rightarrow$ components of sentence
# * Input: sentence
# * Ouput: parse tree with a PoS for each word in sentence
# * Explains the "who did what to whom and why?"
#
#
# ## Grammar
# The syntatic structure of a language
#
# * Phrase: meaningful unit words
# * Clasue: subject + predicate + phrases
# * Sentence: main verb + one or more clasuses
#
# ### Context Free Grammar (CFG)
# Defined as $G = (N, \Sigma, R, S)$
# * N: Non-terminals
# * $\Sigma$: Terminals
# * R: Rules for language, ex $A_i \rightarrow B_1 \ B_2 \ ... \ B_N \quad B_i \in \{N \cup \Sigma \}$
# * S: Start symbol, $S \in N$
#
# Problem: CFGs can produce multiple valid parse trees - ambiguity problem.
#
# ### CKY Parsing
# Dynamic programming approach to building parse trees.
#
# Algorithm
# ```
# CKY(S, G) -> T
# let N = len(S)
# let T = [N][N]
#
# for j = 1...N:
#
# // FILL OUT DIAGONAL
# for all A which satisfy (A -> S[j] in G):
# T[j-1, j] = T[j-1,j] UNION {A}
#
# // FILL OUT SUPER-DIAGONAL
# for i = j-2...0:
# for k = i+1...j-1:
# for all A which satisfy (A -> BC in G)
# and (B in T[i, k]) // left
# and (C in T[k,j]): // below
# T[i,j] = T[i,j] UNION {A}
#
# ```
#
# ### PCFG
#
# # Dependency Parsing
#
# * Dependency Parsing relies on __Dependency Grammars__
# * Consituency parsing relies on __Context Free Grammars__
#
# Idea:
# * Phrase structure is not important
# * Syntatic structure is important
#
# __Typed Dependency__
# * Dependencies between words are of a sepcific class
# * ex det, root, nsub, nmod
# * The structure of a sentence is with directions between the lexical items (words)
#
# __Free word order__
# Some language have a very relaxed rule-set when it comes to ordering
# * This mean many CFG rules would be needed, which makes it infeasible
# * Dependency Grammars has 1 relation per word, pointing to another lexical item, no matter the language
#
# __Grammatical Relation (Binary relations)__
# * Head: Primary noun in NounPhrase or verb in VerbPhrase
# * Dependent: In DG, head dependent relatonship arises from links between head and word immeadiately dependent on the head
#
# Grammatical Function
# * Role of the dependent, relative to the head
# * Subject, direct object, indeirect object
# * In eglish, strongly correlated with word position
# * Not in many other languages
#
# Dependency Parsing Formalism
# * Model as Directed Graph: $G = (V,A) \quad V: vertices, \ A: arcs$
# * $V:$ words, stems, affixes, punctuation
# * $A:$ Grammatical function relationships
#
#
# Dependency Tree Constraints
# * One root node - with no incoming arcs
# * Each node has exactly 1 incoming arc (except root)
# * There exists a unique path from the root to all other vertices
#
# Projectivity
# * An arc is projective iff there exists a path from head to every word between the head and dependent
# * A Dependency Tree is projective iff all arcs are projective
# * I.e. no crossing arcs
# * Flexible word order languages = non projective tree
# * CFGs = Projective Tree
#
# Dependency parsing
# * Lexical head: N = head(NP) and V = head(VP)
# * Head is the most important word in a phrase
# ## Exercises:
#
# __What makes dependency parsing better than constituency parsing when dealing with languages with flexible
# word orders?__
# * Constituency parsing requires rules for all word orders in the form of a CFG, whereas Dep. Parsing uses one single head-dep relation which encapsulates all possible word orderings
#
# __What are the characteristics of the parses generated through dependency parsing that make them more suitable for tasks such as coreference resolution or question answering?__
# * Finds HD relationships, whereas const. parsing requires these relationships to be given beforehand.
#
# __What are the three restrictions that apply to dependency trees?__
# * Excatly one root node with no incoming arcs
# * All nodes exept for the root node has exactly 1 incoming arc
# * There is a path from the root node to every other node in the graph
#
# __An additional constraint is applied to dependency trees, projectivity. What does it mean and why is it important?__
# What is it?
# * Projectivity: Phrase is dependent iff there is a path through every word between Head and Dependent.
# * A phrase is projective if no arcs are crossing, when set up in the sentence order.
# * A tree conisting of only projective phrases is said to be projective
# Why is it important?
# * Transition based parsing produces projective trees, and non-projective trees are errorneous
# * English dependency treebanks were derived from phrase-structure treebanks through the use of head-finding rules, which are projective.
#
# __There are two dominant approaches for dependency parsing, transition-based and graph-based. What are their main advantages and disadvantages?__
# * Transition based: Linear time wrt. to word count, greedy based algorithm (except for beam-search)
# * Graph-based: Exhaustive search, much slower
#
#
| exam-prep/.ipynb_checkpoints/07-parsing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matrix_factorization_utilities
# Load user ratings
raw_training_dataset_df = pd.read_csv('movie_ratings_data_set_training.csv')
raw_testing_dataset_df = pd.read_csv('movie_ratings_data_set_testing.csv')
# Convert the running list of user ratings into a matrix
ratings_training_df = pd.pivot_table(raw_training_dataset_df, index='user_id', columns='movie_id', aggfunc=np.max)
ratings_testing_df = pd.pivot_table(raw_testing_dataset_df, index='user_id', columns='movie_id', aggfunc=np.max)
ratings_testing_df.head()
ratings_training_df.head()
# Apply matrix factorization to find the latent features
U, M = matrix_factorization_utilities.low_rank_matrix_factorization(ratings_training_df.values,
num_features=11,
regularization_amount=1.1)
# Find all predicted ratings by multiplying U and M
predicted_ratings = np.matmul(U, M)
predicted_ratings
# Measure RMSE
rmse_training = matrix_factorization_utilities.RMSE(ratings_training_df.values,
predicted_ratings)
rmse_testing = matrix_factorization_utilities.RMSE(ratings_testing_df.values,
predicted_ratings)
print("Training RMSE: {}".format(rmse_training))
print("Testing RMSE: {}".format(rmse_testing))
| Codes/Machine Learning and AI Foundations - Recommendations/7. Measure Accuracy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3 Jhub
# language: python
# name: py3-jhub
# ---
# %matplotlib inline
from matplotlib import pyplot as plt
from xarray import open_mfdataset as xrdataset
import numpy as np
data_path = "/data0/project/vortex/lahaye/DIAG/NRJ_fluxes/luckyto_pw_fluxes_M2.?.nc"
nc = xrdataset(data_path)
pwavg= nc.variables['pw_avg'][:]
nc.close()
import dask
| NRJ_flux_diag/test_plot_pw-flux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Adding measurements to `Schedule`s
#
# Measurement is clearly a very important part of building a Pulse schedule -- this is required to get the results of our program execution! The powerful low-level control we are granted by Pulse gives us more freedom than `QuantumCircuit`s in specifying how the measurement should be done, enabling you to explore readout error mitigation. This power of course comes with responsibility: we have to understand how measurement works, and accomodate certain hardware constraints.
#
# On this page, we will explore in depth how to create measurements, using several different approaches of increasing complexity.
#
# **Note: Pulse allows you to receive raw, kerneled, and disciminated readout data (whereas circuits will only return discriminated data). Documentation for these options can be found here-COMING SOON.**
#
# ### Adding a backend-default measurement with `measure`
# To add measurements as easily to `Schedule`s as to `QuantumCircuit`s, you just have to know which qubits you want to measure (below, qubits 0 and 1) and have a OpenPulse-enabled `backend`:
#
# ```
# # Appending a measurement schedule to a Schedule, sched
# from qiskit.scheduler import measure
# sched += measure([0, 1], backend) << sched.duration
# ```
# The `backend` contains a default definition for measurement, which is tailored to the qubits you are measuring.
#
# ### Basic measurement pattern and `measure_all`
# Let's use the default measurement feature to inspect a measurement and learn what each pulse does. Below, we use `measure_all`, which measures all the qubits on the backend.
# +
from qiskit import IBMQ
from qiskit.pulse import Schedule
from qiskit.scheduler import measure_all
from qiskit.test.mock import FakeAlmaden
backend = FakeAlmaden()
# +
sched = Schedule(name="Measurement scheduling example")
sched += measure_all(backend)
sched.draw()
# -
# Each qubit has two channels related to readout, as we see above. These are the readout transmit `MeasureChannel`s, and the readout receive `AcquireChannel`s. In superconducting qubit architectures, qubits are coupled to readout resonators. The `MeasureChannel` and `AcquireChannel`s label signal lines which connect to the readout resonator. The coupling between the qubit and the readout resonator hybridizes their state, so when a stimulus pulse is sent to the readout resonantor, the reflected pulse is dependent on the state of the qubit. The acquisition "pulse" is truly a trigger specifying to the analog-to-digital converter (ADC) to begin collecting data, and for how long. That data is used to classify the qubit state.
#
# ### Specifying classical memory slots
#
# If you would like to specify where your measurement results go, there is an option for that in `measure`, called `qubit_mem_slots`. It takes a dictionary mapping qubit indices to classical memory slots. For example, if you want to measure qubit 0 into memory slot 1, you would do this:
# +
from qiskit.scheduler import measure
sched = measure(qubits=[0], backend=backend, qubit_mem_slots={0: 1})
# -
# This would be equivalent to the circuit measurement `circuit.measure(qubit_reg[0], classical_reg[1])`.
# ## Build a measurement sequence from pulses
#
# Rather than use the default measurements provided by the backend, we can also build the measurement sequence up as a basic Pulse schedule. The example below is similar to a typical measurement on IBM systems.
#
# First, we'll build the measurement stimulus pulses for each of the qubits we want to measure. Below, we use a Gaussian square parametric pulse.
# +
from qiskit.pulse import MeasureChannel, AcquireChannel, MemorySlot, GaussianSquare, Acquire
# Duration (in number of cycles) for readout
duration = 16000
# Stimulus pulses for qubits 0 and 1
measure_tx = GaussianSquare(duration=duration, amp=0.2, sigma=10, width=duration - 50)(MeasureChannel(0))
measure_tx += GaussianSquare(duration=duration, amp=0.2, sigma=10, width=duration - 50)(MeasureChannel(1))
# -
# Before we build the acquisition pulses, we need to understand the measurement map.
#
# #### Acquiring qubits: the measurement map `meas_map`
#
# Due to control rack hardware constraints, some qubits may need to be acquired together. This can be the case for qubits whose readout channels are multiplexed. Any OpenPulse-enabled backend will provide a `meas_map` to notify the user of this.
#
# For instance, if we see this for a 5-qubit `backend`
#
# ```
# backend.configuration().meas_map
#
# Out: [[0, 1, 2, 3, 4]]
# ```
#
# then we know that all the qubits on this device must be acquired together. On the other hand, this output
#
# ```
# Out: [[0], [1], [2], [3, 4]]
# ```
#
# tells us that qubits 0, 1 and 2 can be acquired independently, but qubits 3 and 4 must be acquired together.
#
# When building up a pulse schedule, be sure to add all the acquire pulses required by the backend you plan to run on. This is validated at assemble time.
#
# Getting back to our example, let's imagine we plan to run on a backend with this measurement map: `[[0, 1, 2]]`. Now we can build the acquisition pulses. This is done with the `Acquire` command, which takes only a duration. We specify the channels and memory slots to acquire on.
# Acquisition instructions
acquire = Acquire(duration)
measure_rx = acquire(AcquireChannel(0), MemorySlot(0))
measure_rx += acquire(AcquireChannel(1), MemorySlot(1))
# Finally, we just combine the two parts together. Every instruction is on a different channel, so appending schedules the instructions at time 0. The `measure_schedule` can then be added to the end of any Pulse schedule to measure qubits 0 and 1 into classical memory slots 0 and 1.
# +
measure_sched = measure_tx + measure_rx
measure_sched.draw()
# -
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| qiskit/advanced/terra/programming_with_pulses/adding_measurements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PW_explorer.load_worlds import load_worlds
from PW_explorer.run_clingo import run_clingo
from DLV_Input_Parser.dlv_rules_parser import parse_dlv_rules
# %load_ext PWE_NB_Extension
# +
# %%clingo --donot-display_input -lci automorphisms --donot-run
% e(X,Y) :- e(Y,X). --> only if undirected
gnode(X) :- e(X,_).
gnode(X) :- e(_,X).
vmap(X,Y) ; vout(X,Y) :- gnode(X), gnode(Y).
:- vmap(X1,Y1), vmap(X2,Y2), e(X1,X2), not e(Y1,Y2).
:- vmap(X1,Y1), vmap(X2,Y2), not e(X1,X2), e(Y1,Y2).
% used1(X) :- vmap(X,_).
% :- gnode(X), not used1(X).
% :- vmap(X,Y),vmap(X,Z),Y!=Z.
% :- vmap(Y,X),vmap(Z,X),Y!=Z.
:- gnode(X), #count {Y: vmap(X,Y)} != 1.
:- gnode(X), #count {Y: vmap(Y,X)} != 1.
% #show vmap/2.
#show.
# -
symm_degree_rules = str(automorphisms).split('\n')
def get_incidence_graph_edge_facts(rule):
listener = parse_dlv_rules(rule, print_parse_tree=False)
edges = []
for rule in listener.rules:
head_atoms, tail_atoms = rule[0], rule[1]
atom_count = 0
for head in head_atoms+tail_atoms:
atom_node = '"{}_{}_{}"'.format(head.rel_name, head.rel_arity, atom_count)
edges.extend([('"{}"'.format(v), atom_node) for v in head.vars])
atom_count += 1
edge_facts = []
for e in edges:
edge_facts.append("e({},{}).".format(*e))
return edge_facts
def get_symm_degree(rule):
edge_facts = get_incidence_graph_edge_facts(rule)
# print(edge_facts)
asp_out, _ = run_clingo(symm_degree_rules+edge_facts)
_, _, pw_objs = load_worlds(asp_out, silent=True)
return len(pw_objs)
# tri/0
get_symm_degree('tri :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/1
get_symm_degree('tri(X) :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/2
get_symm_degree('tri(X,Y) :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/3
get_symm_degree('tri(X,Y,Z) :- e(X,Y), e(Y,Z), e(Z,X).')
# thop/0
get_symm_degree('thop :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/1
get_symm_degree('thop(X) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/2
get_symm_degree('thop(X,Y) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/3
get_symm_degree('thop(X,Y,Z1) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/4
get_symm_degree('thop(X,Y,Z1,Z2) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# +
#### ROUGH WORK FROM DEV ####
# -
test_rule = 'tri(X,Y) :- e(X,Y), e(Y,Z), e(Z,X).'
listener = parse_dlv_rules(test_rule, print_parse_tree=False)
edges = []
for rule in listener.rules:
head_atoms, tail_atoms = rule[0], rule[1]
atom_count = 0
for head in head_atoms+tail_atoms:
atom_node = '"{}_{}_{}"'.format(head.rel_name, head.rel_arity, atom_count)
edges.extend([('"{}"'.format(v), atom_node) for v in head.vars])
atom_count += 1
edges
edge_facts = []
for e in edges:
edge_facts.append("e({},{}).".format(*e))
print("\n".join(edge_facts))
# %clingo -l automorphisms edge_facts --donot-display_input
# +
# %%clingo -l automorphisms --donot-display_input
% rigid -- symm degree = 1 when undirected
e(a,b). e(b,c). e(b,d). e(d,c). e(d,e). e(e,f).
e(X,Y) :- e(Y,X).
# +
# %%clingo -l automorphisms --donot-display_input
% peterson graph -- symm degree = 120 when undirected
e(a,b). e(b,c). e(c,d). e(d,e). e(e,a).
e(a,a1). e(b,b1). e(c,c1). e(d,d1). e(e,e1).
e(a1,c1). e(a1,d1). e(b1,d1). e(b1,e1). e(c1,e1).
e(X,Y) :- e(Y,X).
# +
# %%clingo -l automorphisms --donot-display_input
% peterson graph -- symm degree = 120 when undirected
e(x1,p1). e(x2,p1). e(x2,p2). e(x3,p2). e(x3,p3). e(x4,p3).
# +
# %%clingo -l automorphisms --donot-display_input
e(u,s1). e(v,s1). e(u,s2). e(w,s2).
# +
# %%clingo -l automorphisms --donot-display_input
e(u1,r1). e(u2,r2). e(u3,r3).
# +
# %%clingo -l automorphisms --donot-display_input
e(x,q).
e(x,t). e(x,s1).
e(y,t).
e(z,s1). e(z,s2).
e(2,s2).
# +
# %%clingo -l automorphisms --donot-display_input
e(x,tri). e(y,tri). e(z,tri).
e(x,e1). e(y,e1).
e(y,e2). e(z,e2).
e(z,e3). e(x,e3).
# +
# %%clingo -l automorphisms --donot-display_input
e("X","tri_2_0").
e("Y","tri_2_0").
e("X","e_2_1").
e("Y","e_2_1").
e("Y","e_2_2").
e("Z","e_2_2").
e("Z","e_2_3").
e("X","e_2_3").
# -
| Query Analysis/Symmetry Degree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %cat 0Source_Citation.txt
# %matplotlib inline
# # %matplotlib notebook # for interactive
# For high dpi displays.
# %config InlineBackend.figure_format = 'retina'
# # 0. General note
# This example compares pressure calculated from `pytheos` and original publication for the platinum scale by Dorogokupets 2015.
# # 1. Global setup
import matplotlib.pyplot as plt
import numpy as np
from uncertainties import unumpy as unp
import pytheos as eos
# # 3. Compare
eta = np.linspace(1., 0.70, 7)
print(eta)
dorogokupets2015_pt = eos.platinum.Dorogokupets2015()
help(eos.platinum.Dorogokupets2015)
dorogokupets2015_pt.print_equations()
dorogokupets2015_pt.print_equations()
dorogokupets2015_pt.print_parameters()
v0 = 60.37930856339099
dorogokupets2015_pt.three_r
v = v0 * (eta)
temp = 3000.
p = dorogokupets2015_pt.cal_p(v, temp * np.ones_like(v))
print('for T = ', temp)
for eta_i, p_i in zip(eta, p):
print("{0: .3f} {1: .2f}".format(eta_i, p_i))
# The table is not given in this publication.
v = dorogokupets2015_pt.cal_v(p, temp * np.ones_like(p), min_strain=0.6)
print((v/v0))
| examples/6_p_scale_test_Dorogokupets2015_Pt.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
from basketball_reference_web_scraper import client
from basketball_reference_web_scraper.data import Team
import csv
import os
import random
import time
import copy
import glob
import shutil
import tqdm
user_agent_list = [ #Chrome
'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.113 Safari/537.36',
'Mozilla/5.0 (Windows NT 6.1; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.90 Safari/537.36',
'Mozilla/5.0 (Windows NT 5.1; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.90 Safari/537.36',
'Mozilla/5.0 (Windows NT 6.2; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.90 Safari/537.36',
'Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/44.0.2403.157 Safari/537.36',
'Mozilla/5.0 (Windows NT 6.3; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.113 Safari/537.36',
'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/57.0.2987.133 Safari/537.36',
'Mozilla/5.0 (Windows NT 6.1; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/57.0.2987.133 Safari/537.36',
'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/55.0.2883.87 Safari/537.36',
'Mozilla/5.0 (Windows NT 6.1; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/55.0.2883.87 Safari/537.36',
#Firefox
'Mozilla/4.0 (compatible; MSIE 9.0; Windows NT 6.1)',
'Mozilla/5.0 (Windows NT 6.1; WOW64; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (compatible; MSIE 9.0; Windows NT 6.1; WOW64; Trident/5.0)',
'Mozilla/5.0 (Windows NT 6.1; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (Windows NT 6.2; WOW64; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (Windows NT 10.0; WOW64; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (compatible; MSIE 9.0; Windows NT 6.0; Trident/5.0)',
'Mozilla/5.0 (Windows NT 6.3; WOW64; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (compatible; MSIE 9.0; Windows NT 6.1; Trident/5.0)',
'Mozilla/5.0 (Windows NT 6.1; Win64; x64; Trident/7.0; rv:11.0) like Gecko',
'Mozilla/5.0 (compatible; MSIE 10.0; Windows NT 6.1; WOW64; Trident/6.0)',
'Mozilla/5.0 (compatible; MSIE 10.0; Windows NT 6.1; Trident/6.0)',
'Mozilla/4.0 (compatible; MSIE 8.0; Windows NT 5.1; Trident/4.0; .NET CLR 2.0.50727; .NET CLR 3.0.4506.2152; .NET CLR 3.5.30729)'
]
output_base = 'scraped/'
def wait_random_time(max_wait_seconds=10.25):
import random
import time
### wait some period of time and set a new user agent string
seconds_to_wait = random.random()*max_wait_seconds
time.sleep(seconds_to_wait)
client.http_client.USER_AGENT = random.choice(user_agent_list)
# + code_folding=[]
def download_yearly_stats(year, return_download_tables=False):
downloaded_tables = {}
output_file_path = output_base+f'/stats_by_year/{year}_totals.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['totals'] = client.players_season_totals(year, output_type='csv',
output_file_path=output_file_path)
wait_random_time()
output_file_path = output_base+f'/stats_by_year/{year}_advanced.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['advanced'] = client.players_advanced_stats(year, output_type='csv',
output_file_path=output_file_path)
wait_random_time()
### repeat for the playoffs:
output_file_path = output_base+f'/stats_by_year/{year}_playoffs_totals.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['totals.playoffs'] = client.players_season_totals(year, playoffs=True, output_type='csv',
output_file_path=output_file_path)
wait_random_time()
output_file_path = output_base+f'/stats_by_year/{year}_playoffs_advanced.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['advanced.playoffs'] = client.players_advanced_stats(year, playoffs=True, output_type='csv',
output_file_path=output_file_path)
wait_random_time()
## now do the per 100 if they're aviailable
if year >= 1974:
output_file_path = output_base+f'/stats_by_year/{year}_per100.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['per100'] = client.players_season_totals_per100(year, output_type='csv',
output_file_path=output_file_path)
wait_random_time()
output_file_path = output_base+f'/stats_by_year/{year}_playoffs_per100.csv'
if not os.path.isfile(output_file_path):
downloaded_tables['per100.playoffs'] = client.players_season_totals_per100(year,
playoffs=True, output_type='csv', output_file_path=output_file_path)
wait_random_time()
if return_download_tables:
return downloaded_tables
for year in tqdm.tqdm(range(1950, 2019)):
download_yearly_stats(year)
# print(f"Done with stats for {year}")
# -
# ### Download series-by-series stats for the playoffs:
for year in range(1950, 2019):
if year == 1954:
print("Skipping 1954")
continue
print(f"Starting on the {year} playoffs")
output_directory = output_base + f'playoffs_by_series/{year}/'
os.makedirs(output_directory, exist_ok=True)
client.playoffs_series_in_one_year(year, output_directory=output_directory, output_type='csv')
wait_random_time(60)
# + code_folding=[]
from basketball_reference_web_scraper import output
from basketball_reference_web_scraper.data import OutputWriteOption
import csv
output_write_option = OutputWriteOption("w")
def combine_series_by_round(year):
directory = output_base + f'playoffs_by_series/{year}/'
for table in ['basic', 'advanced']:
if year < 1984 and table == 'advanced':
continue
files = glob.glob(directory+f'*_{table}.csv')
file_lists = {}
for fname in files:
pround = fname.split('/')[-1][:-len('.csv')].split('_')[0]
if pround in file_lists:
file_lists[pround].append(fname)
else:
file_lists[pround] = [fname]
for pround, files in file_lists.items():
if len(files) == 1:
continue
files = sorted(files)
if '_0' not in files[0]:
input_file_path = files[0][:-len('.csv')] + '_0.csv'
shutil.move(files[0], input_file_path)
files[0] = input_file_path
else:
input_file_path = files[0]
output_file_path = input_file_path[:-len(f'0_{table}.csv')] + f'{table}.csv'
print("Combining:\n\t"+'\n\t'.join(files))
all_rows = []
for file_path in files:
with open(file_path, 'r') as input_file:
CSVfile = csv.reader(input_file)
header = None
for row in CSVfile:
if header is None:
header = row
else:
all_rows.append({k: row[ii] for ii, k in enumerate(header)})
print(f"Writing {len(all_rows)} total lines to {output_file_path}")
output.playoff_stats_writer(all_rows, output_file_path, output_write_option, table)
# -
for year in range(1950, 2019):
combine_series_by_round(year)
## had to redo 2018:
combine_series_by_round(2018)
# +
### doing all the playoff series
## deprecated because I wrote a function in client to do it with less hassle (though with overwriting)
# def read_playoff_series_list(file_path):
# series_list = []
# with open(file_path, 'r') as input_file:
# CSVfile = csv.reader(input_file)
# header = None
# for row in CSVfile:
# if header is None:
# header = row
# else:
# output_dictionary = {}
# for ii, k in enumerate(header):
# if header[ii] in ['winning_team', 'losing_team']:
# output_dictionary[k] = Team(row[ii])
# else:
# output_dictionary[k] = row[ii]
# series_list.append(output_dictionary)
# return series_list
# def add_unique_name_and_round(series_list):
# series_count = {}
# for series in series_list:
# round_name = series['series_name']
# finals = 'semi' in round_name.lower()
# if True in [round_name.startswith(f'{x} ') for x in ['Western', 'Central', 'Eastern']]:
# finals = False
# if round_name in series_count:
# unique_series_name = round_name + '_{}'.format(series_count[round_name])
# series_count[round_name] = series_count[round_name] + 1
# elif finals:
# unique_series_name = copy.deepcopy(round_name)
# else:
# unique_series_name = round_name + '_0'
# series_count[round_name] = 1
# series['round_name'] = round_name
# series['unique_series_name'] = unique_series_name
# return series_list
# def download_playoff_series_in_a_year(year, return_download_tables=False):
# schedule_file_path = output_base + f'playoff_schedules/{year}_playoffs.csv'
# if os.path.isfile(schedule_file_path):
# print("Reading schedule from "+schedule_file_path)
# playoff_series_list = read_playoff_series_list(schedule_file_path)
# else:
# playoff_series_list = client.playoff_series_list(year, output_type='csv',
# output_file_path=schedule_file_path)
# wait_random_time()
# playoff_series_list = add_unique_name_and_round(playoff_series_list)
# output_directory = output_base + f'playoffs_by_series/{year}/'
# downloaded_tables = []
# for series in playoff_series_list:
# unique_name = series['unique_series_name']
# round_name = series['round_name']
# output_file_path_base = output_directory + unique_name
# if not os.path.isfile(output_file_path_base+'_basic.csv'):
# tables = client.playoff_series_stats(series, output_type='csv',
# output_file_path=output_file_path_base)
# print("Downloaded tables for {}".format(series['unique_series_name']))
# downloaded_tables.append(tables)
# wait_random_time()
# if return_download_tables:
# return downloaded_tables
# for year in range(1950, 2019):
# print(f"Beginning series from the {year} Playoffs")
# os.makedirs(output_base+f'playoffs_by_series/{year}/', exist_ok=True)
# download_playoff_series_in_a_year(year)
# print(f"Finished with series in the {year} Playoffs")
# -
| download_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing libs
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from src.visualization.visualize import categorical_eda, timeseries_eda
# %load_ext autoreload
# %autoreload 2
# ## Loading data
df_raw = pd.read_csv('../data/interim/attr_added/delivery.csv')
df_raw.head()
df_raw.info()
df = df_raw.iloc[:-1, :].copy()
# ## EDA
df.shape
# ### id
df['id'].unique()
df['id'].nunique()
df['id'].isna().sum()
# ### state
categorical_eda(df, 'state', (15, 10))
# ### district
categorical_eda(df, 'district', (15, 10))
df.info()
# ### case_no
df['case_no'].unique()
categorical_eda(df, 'case_no', (15, 10))
# #### death_cause
categorical_eda(df, 'death_cause', (15, 10))
# ### death_other
categorical_eda(df, 'death_other', (15, 5))
df.info()
# ### delivery_comp1
categorical_eda(df, 'delivery_comp1', (10, 5))
# ### delivery_comp2
categorical_eda(df, 'delivery_comp2', (10, 5))
df[df['delivery_comp1'] == 'N']['delivery_comp2'].value_counts()
df[df['delivery_comp1'] == ' ']['delivery_comp2'].value_counts()
# ### delivery_comp3
categorical_eda(df, 'delivery_comp3', (10, 5))
df[df['delivery_comp1'] == 'N']['delivery_comp3'].value_counts()
df[df['delivery_comp1'] == ' ']['delivery_comp3'].value_counts()
# ### delivery_comp4
categorical_eda(df, 'delivery_comp4', (10, 5))
df[df['delivery_comp1'] == 'N']['delivery_comp4'].value_counts()
df[df['delivery_comp1'] == ' ']['delivery_comp4'].value_counts()
# ### delivery_comp5
categorical_eda(df, 'delivery_comp5', (10, 5))
df.info()
# ### delivery_complication
categorical_eda(df, 'delivery_complication', (10, 5))
# ### delivery_conducted_by
categorical_eda(df, 'delivery_conducted_by', (10, 5))
# ### delivery_date
timeseries_eda(df, 'delivery_date', (15, 5))
df.info()
# ### delivery_outcomes
categorical_eda(df, 'delivery_outcomes', (10, 5))
# ### delivery_place
categorical_eda(df, 'delivery_place', (10, 5))
# ### delivery_type
categorical_eda(df, 'delivery_type', (10, 5))
df.info()
# ### discharge_date
timeseries_eda(df, 'discharge_date', (15, 5))
df['discharge_date']
df[df['discharge_date'] == '1900-01-01 00:00:00']
# ### registration_no
df['registration_no'].nunique()
categorical_eda(df, 'registration_no', (15, 5))
df.info()
# ### rural_urban
categorical_eda(df, 'rural_urban', (10, 5))
# ### still_birth
categorical_eda(df, 'still_birth', (10, 5))
# ### isactive
categorical_eda(df, 'isactive', (10, 5))
# ### other_delivery_complication
categorical_eda(df, 'other_delivery_complication', (10, 5))
df.info()
# ### previous_status
categorical_eda(df, 'previous_status', (10, 5))
# ### created_on
timeseries_eda(df, 'created_on', (15, 5))
# Unique dates
pd.to_datetime(df['created_on']).map(lambda t: t.date()).nunique()
# ### live_birth
categorical_eda(df, 'live_birth', (10, 5))
| notebooks/0.3-gokul-EDA_delivery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gen-methods
# language: python
# name: gen-methods
# ---
# +
# Notebooks
import nbimporter
import os
import sys
# Functions from src
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
# Defined Functions
from utils import *
# from utils import *
# Pandas, matplotlib, pickle, seaborn
import pickle
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# -
datasets = ["occutherm-reduced", "cresh", "ashrae-reduced"]
models = ["smote", "adasyn", "tgan", "ctgan"]
models_axes_baseline = ["Baseline", "SMOTE", "ADASYN", "TGAN", "CTGAN"]
# # Variability of generated samples
# 
# +
var_all_datasets = []
for dataset in datasets:
variability_all = []
var_baseline = pickle.load(open("metrics/" + dataset + "_variability_baseline.pkl", "rb"))
variability_all.append(var_baseline)
for model in models:
if model == "comfortGAN":
metric_str = "metrics/" + dataset + "-" + experiment_name + "_variability_" + model + ".pkl"
else:
metric_str = "metrics/" + dataset + "_variability_" + model + "_trials.pkl"
var = pickle.load(open(metric_str, "rb"))
variability_all.append(var)
print(variability_all)
var_all_datasets.append(variability_all)
print(var_all_datasets)
fig = plt.figure()
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
x = np.arange(len(models_axes_baseline)) # the label locations
width = 0.3 # the width of the bars
ax.bar(x - width, var_all_datasets[0], width, label='occutherm')
ax.bar(x,var_all_datasets[1], width, label='cresh')
ax.bar(x + width,var_all_datasets[2], width, label='ashrae')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set(ylim=(0, 230))
ax.set_xticks(x)
ax.set_xticklabels(models_axes_baseline)
ax.tick_params(length=20, direction="inout", labelsize='large')
ax.set_ylabel('L2 distance couple', size=15)
ax.legend(prop={'size': 15})
plt.show()
# higher the better and close to baseline (each row is one dataset)
# -
# # Diversity of generated samples
# 
# +
diversity_all_datasets = []
for dataset in datasets:
diversity_all = []
div_baseline = pickle.load(open( "metrics/" + dataset + "_diversity_baseline.pkl", "rb" ))
diversity_all.append(div_baseline)
for model in models:
if model == "comfortGAN":
metric_str = "metrics/" + dataset + "-" + experiment_name + "_diversity_" + model + ".pkl"
else:
metric_str = "metrics/" + dataset + "_diversity_" + model + "_trials.pkl"
div = pickle.load(open( metric_str, "rb" ))
diversity_all.append(div)
print(diversity_all)
diversity_all_datasets.append(diversity_all)
print(diversity_all_datasets)
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
x = np.arange(len(models_axes_baseline)) # the label locations
width = 0.3 # the width of the bars
ax.bar(x - width, diversity_all_datasets[0], width, label='occutherm')
ax.bar(x, diversity_all_datasets[1], width, label='cresh')
ax.bar(x + width, diversity_all_datasets[2], width, label='ashrae')
# ax.set(ylim=(0, 40))
ax.set_xticks(x)
ax.set_xticklabels(models_axes_baseline)
ax.tick_params(length=20, direction="inout", labelsize='large')
ax.set_ylabel('L2 distance synthetic and real', size=15)
ax.legend(prop={'size': 15})
plt.show()
# higher the better and close to baseline (each row is one dataset)
# -
# # Quality of the final Classification
# 
# +
class_acc_all_dasets = []
for dataset in datasets:
class_acc_all = []
class_acc_baseline = pickle.load(open( "metrics/" + dataset + "_rdf_classification_baseline.pkl", "rb" ))
class_acc_all.append(class_acc_baseline)
for model in models:
if model == "comfortGAN":
metric_str = "metrics/" + dataset + "-" + experiment_name + "_classification_test_" + model + ".pkl"
else:
metric_str = "metrics/" + dataset + "_classification_" + model + "_trials.pkl"
class_acc = pickle.load(open( metric_str, "rb" ))
class_acc_all.append(class_acc[3])
print(class_acc_all)
class_acc_all_dasets.append(class_acc_all)
print(class_acc_all_dasets)
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
x = np.arange(len(models_axes_baseline)) # the label locations
width = 0.3 # the width of the bars
ax.bar(x - width, class_acc_all_dasets[0], width, label='occutherm')
ax.bar(x, class_acc_all_dasets[1], width, label='cresh')
ax.bar(x + width, class_acc_all_dasets[2], width, label='ashrae')
ax.set(ylim=(0, 1))
ax.set_xticks(x)
ax.set_xticklabels(models_axes_baseline)
ax.tick_params(length=20, direction="inout", labelsize='large')
ax.set_ylabel('Accuracy', size=15)
ax.legend(prop={'size': 15})
plt.show()
# higher the better and close to baseline (each row is one dataset)
# -
| 8b-Benchmarks-Results-reduced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating a Custom Basket
#
# Welcome to the basket creation tutorial! Marquee allows you to create your own tradable basket ticker and manage it through the platform. When you create a basket it automatically gets published to Marquee, and you may also publish it to Bloomberg, Reuters, and Factset. This basket will tick live.
#
# Creating a basket requires enhanced levels of permissioning. If you are not yet permissioned to create baskets please reach out to your sales coverage or to the [Marquee sales team](mailto:<EMAIL>).
# ## Step 1: Authenticate & Initialize your session
#
# First you will import the necessary modules and add your client id and client secret.
# +
import pandas as pd
from datetime import date
from gs_quant.markets.baskets import Basket
from gs_quant.markets.indices_utils import ReturnType
from gs_quant.markets.position_set import Position, PositionSet
from gs_quant.session import Environment, GsSession
client = 'CLIENT ID'
secret = 'CLIENT SECRET'
GsSession.use(Environment.PROD, client_id=client, client_secret=secret, scopes=('read_product_data read_user_profile modify_product_data',))
# -
# ## Step 2: Define your basket metadata, publishing options, pricing options, & return type
#
# In this step you are going to define all the specifications needed to create your basket. First, instantiate an empty basket object and then you may begin defining it's settings. The below list contains all the parameters you may set.
#
# | Parameter Name | Required? | Default Value | Description |
# |:-------------------|:-----------|:--------------|:------------|
# |name |**Required**|-- |Display name of the basket|
# |ticker |**Required**|-- |Associated 8-character basket identifier (must be prefixed with "GS" in order to publish to Bloomberg). If you would like to request a custom prefix instead of using the default GSMB prefix please reach out to the [baskets team](mailto:<EMAIL>)|
# |currency |**Required**|-- |Denomination you want your basket to tick in. This can not be changed once your basket has been created|
# |return_type |**Required**|-- |Determines the index calculation methodology with respect to dividend reinvestment. One of Price Return, Gross Return, Total Return|
# |position_set |**Required**|-- |Information of constituents associated with the basket. You may provide the weight or quantity for each position. If neither is provided we will distribute the total weight evenly among each position. Please also note that any fractional shares will be rounded up to whole numbers.|
# |description |Optional |-- |Free text description of basket. Description provided will be indexed in the search service for free text relevance match.|
# |divisor |Optional |-- |Divisor to be applied to the overall position set. You need not set this unless you want to change the divisor to a specific number, which will in turn change the basket price (current notional/divisor). This might impact price continuity.|
# |initial_price |Optional |100 |Initial price the basket should start ticking at|
# |target_notional |Optional |10,000,000 |Target notional for the position set|
# |publish_to_bloomberg|Optional |True |If you'd like us to publish your basket to Bloomberg|
# |publish_to_reuters |Optional |False |If you'd like us to publish your basket to Reuters |
# |publish_to_factset |Optional |False |If you'd like us to publish your basket to Factset |
# |default_backcast |Optional |True |If you'd like us to backcast up to 5 years of pricing history and compositions, assuming constituents remained constant. Set to false if you'd like to upload your own custom history. If any IPOs are present in this composition, we will stop the backcast period accordingly.|
# |reweight |Optional |False |If you'd like us to reweight positions if input weights don't add up to 1 upon submission|
# |weighting_strategy |Optional |-- |Strategy used to price the position set (will be inferred if not indicated). One of Equal, Market Capitalization, Quantity, Weight|
# |allow_ca_restricted_assets|Optional|False |Allow your basket to have constituents that will not be corporate action adjusted in the future (You will recieve a message indicating if this action is needed when attempting to create your basket)|
# |allow_limited_access_assets|Optional|False |Allow basket to have constituents that GS has limited access to (You will recieve a message indicating if this action is needed when attempting to create your basket)|
# +
my_basket = Basket()
my_basket.name = 'My New Custom Basket'
my_basket.ticker = 'GSMBXXXX'
my_basket.currency = 'USD'
my_basket.publish_to_reuters = True
my_basket.return_type = ReturnType.PRICE_RETURN
# -
# ### Quick Tip!
# At any point, you may call the get_details() method on your basket, which will print the current state of the basket object. We recommend doing this throughout the creation process to ensure there are not any discrepancies between your preferences and the current basket settings.
my_basket.get_details() # prints out each parameters on the basket
# ## Step 3: Define your basket's composition
#
# Now you will decide what your basket composition is. If you'd like to include several positions, you may define the composition using your preferred input method (e.g., uploading an excel file) but it must then be converted to a dictionary or pandas dataframe.
#
# Your dataframe must have a column entitled 'identifier', which holds any commonly accepted identifier such as BloombergId, Cusip, Ticker, etc. for each position. You may also have a column entitled 'quantity' to store the number of shares for each position, or a column named 'weight' to represent the weight of each. If the second column is missing, we will later assign equal weight to each position when you submit your basket for creation.
#
# After uploading your composition and converting it to a dataframe, make sure to rename your columns to match our specifications if they aren't in the correct format already, and then you may use it to create a valid Position Set. You should then call get_positions() to make sure that your positions have all been mapped correctly, and can then store this composition on the basket.
# +
positions_df = pd.read_excel('path/to/excel.xlsx') # example of uploading composition from excel document
positions_df.columns = ['identifier', 'weight'] # replace weight column with 'quantity' if using number of shares
position_set = PositionSet.from_frame(positions_df)
position_set.get_positions() # returns a dataframe with each position's identifier, name, Marquee unique identifier, and weight/quantity
my_basket.position_set = position_set
# -
# ### Quick Tip!
# Wanting to quickly add one or two positions to a position set without having to modify your dataframe? You can add to a position set by inputting an identifier and an optional weight/quantity to a Position object and modify the position set directly, like below. Refer to the [position_set examples](../examples/03_basket_creation/position_set/0004_add_position_to_existing_position_set.ipynb) section for more tips like this!
# +
positions_to_add = [Position('AAPL UW', weight=0.1), Position('MSFT UW', weight=0.1)]
position_set.positions += positions_to_add
my_basket.position_set = position_set
# -
# ## Step 4: Create your basket
#
# Once you've ensured that your basket has been set up to your satisfaction, you're ready to officially create and publish to Marquee! Once you call create on your new basket, you may poll its status to make sure that it has processed successfully. This will check the report status every 30 seconds for 10 minutes by default, but you can override this option if you prefer as shown below. If you'd like to view your basket on the Marquee site, you can retrieve the link to your page by calling get_url().
# +
my_basket.get_details() # we highly recommend verifying the basket state looks correct before calling create!
my_basket.create()
my_basket.poll_status(timeout=120, step=20) # optional: constantly checks create status until report succeeds, fails, or the poll times out (this example checks every 20 seconds for 2 minutes)
my_basket.get_url() # will return a url to your Marquee basket page ex. https://marquee.gs.com/s/products/MA9B9TEMQ2RW16K9/summary
# -
# ## Step 5: Update your basket's entitlements
#
# The application you use to create your basket will initially be the only one permissioned to view, edit, and submit rebalance requests. If you'd like to entitle other users or groups with view or admin access, you may update your basket's permissions at any time.
#
# In order to add or remove permissions for a specific user, you will need either their Marquee user id or email. You may also permission groups using their group id. See the snippet below, or refer to the [baskets permissions examples](../examples/07_basket_permissions/0001_permission_application_to_basket.ipynb) for more options.
# +
from gs_quant.entities.entitlements import User
user = User.get(user_id='application_id')
basket.entitlements.view.users += [user] # update the desired entitlements block ('edit', 'admin', etc) 'users' property
basket.update()
# -
# ### You're all set, Congrats! What's next?
#
# * [How do I upload my basket's historical composition?](../examples/03_basket_creation/0001_upload_basket_position_history.ipynb)
#
# * [How do I retrieve composition data for my basket?](../examples/01_basket_composition_data/0000_get_latest_basket_composition.ipynb)
#
# * [How do I retrieve pricing data for my basket?](../examples/02_basket_pricing_data/0000_get_latest_basket_close_price.ipynb)
#
# * [How do I change my basket's current composition?](./Basket%20Rebalance.ipynb)
#
# * [How do I make other changes to my basket (name, description, etc.)?](./Basket%20Edit.ipynb)
#
# * [What else can I do with my basket?](https://developer.gs.com/docs/gsquant/api/classes/gs_quant.markets.baskets.Basket.html#gs_quant.markets.baskets.Basket)
#
# Other questions? Reach out to the [baskets team](mailto:<EMAIL>) anytime!
| gs_quant/documentation/06_baskets/tutorials/Basket Create.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jvishnuvardhan/Keras_Examples/blob/master/Untitled480.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="6FbFm1LKUP4P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 581} outputId="cdaa0262-0f05-4b70-b008-fae1d78b40d2"
# !pip install tensorflow==2.0.0rc1
# + id="Zr-GHrAPUQ5E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="8a094dfe-bede-460c-b748-1346faddadbc"
import tensorflow as tf
from tensorflow import keras
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
def create_model():
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
# Create a basic model instance
model=create_model()
history=model.fit(x_train, y_train, epochs=10,validation_data=(x_test,y_test))
loss, acc = model.evaluate(x_test, y_test,verbose=1)
print("Original model, accuracy: {:5.2f}%".format(100*acc))
# + id="c77UbEtkUXeW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cccefb04-3317-4fc1-a34f-61006dda999a"
print(history.history.keys())
# + id="68V_wyEtUrQL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="4461e7f5-7dc5-4521-a68e-f81a88f3139b"
import matplotlib.pyplot as plt
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model_Accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(["train","test"], loc="upper left")
plt.show()
# summarize history for accuracy
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model_loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(["train","test"], loc="upper left")
plt.show()
# + id="GBsJAUBtWBH-" colab_type="code" colab={}
| Untitled480.ipynb |