code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Integrate Data Validation Into Your Pipeline
#
#
# +
# Prep environment and logging
import json
import os
import logging
import great_expectations as ge
import great_expectations.jupyter_ux
import pandas as pd
import uuid # used to generate run_id
from datetime import datetime
import tzlocal
great_expectations.jupyter_ux.setup_notebook_logging()
# -
# ## Integrate data validation into your pipeline
#
# [**Watch a short tutorial video**](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#video)
#
#
# [**Read more in the tutorial**](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation)
#
# **Reach out for help on** [**Great Expectations Slack**](https://greatexpectations.io/slack)
#
#
#
# ### Get a DataContext object
#
context = ge.data_context.DataContext()
# ### Get a pipeline run id
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#set-a-run-id)
#
# Generate a run-id that GE will use to key shared parameters
run_id = datetime.utcnow().isoformat().replace(":", "") + "Z"
run_id
# ### Choose data asset name and expectation suite name
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#choose-data-asset-and-expectation-suite)
#
context.list_expectation_suite_keys()
data_asset_name = "REPLACE ME!" # TODO: replace with your value!
expectation_suite_name = "my_suite" # TODO: replace with your value!
# ### Obtain the batch to validate
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#obtain-a-batch-to-validate)
#
#
# ##### If your pipeline processes Pandas Dataframes:
#
# ```
# df = pd.read_csv(file_path_to_validate)
# df.head()
# batch = context.get_batch(data_asset_name, expectation_suite_name, df)
# ```
#
# ##### If your pipeline processes Spark Dataframes:
# ```
# from pyspark.sql import SparkSession
# from great_expectations.dataset import PandasDataset, SqlAlchemyDataset, SparkDFDataset
# spark = SparkSession.builder.getOrCreate()
# df = SparkDFDataset(spark.read.csv(file_path_to_validate))
# df.spark_df.show()
# batch = context.get_batch(data_asset_name, expectation_suite_name, df)
# ```
#
# ##### If your pipeline processes SQL querues:
# ```
# batch = context.get_batch(data_asset_name, expectation_suite_name, query="SELECT * from ....") # the query whose result set you want to validate
# ```
#
# ### Validate the batch
#
# This is the "workhorse" method of Great Expectations. Call it in your pipeline code after loading the file and just before passing it to your computation.
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#validate)
#
#
# +
validation_result = batch.validate(run_id=run_id)
if validation_result["success"]:
print("This file meets all expectations from a valid batch of {0:s}".format(data_asset_name))
else:
print("This file is not a valid batch of {0:s}".format(data_asset_name))
# -
# ### Review the validation results
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#review-validation-results)
#
print(json.dumps(validation_result, indent=4))
# ### Finishing touches - notifications and saving validation results and validated batches
#
# #### Notifications
# You want to be notified when the pipeline validated a batch, especially when the validation failed.
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#send-notifications)
#
# #### Saving validation results
#
# To enable the storing of validation results, uncomment the `result_store` section in your great_expectations.yml file.
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#save-validation-results)
#
# #### Saving failed batches
#
# When a batch fails validation (it does not pass all the expectations of the data asset), it is useful to save the batch along with the validation results for future review. You can enable this option in your project's great_expectations.yml file.
#
# [Read more in the tutorial](https://docs.greatexpectations.io/en/latest/getting_started/pipeline_integration.html?utm_source=notebook&utm_medium=integrate_validation#save-failed-batches)
#
#
| tests/data_context/fixtures/post_init_project_v0.8.0_A/great_expectations/notebooks/integrate_validation_into_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Exercise 1
from sklearn.datasets import fetch_mldata
import numpy as np
mnist = fetch_mldata('MNIST original')
X, y = mnist['data'], mnist['target']
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
shuffle_index = np.random.permutation(60000)
X_train, y_train = X_train[shuffle_index], y_train[shuffle_index]
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
knn_class = KNeighborsClassifier()
knn_class.fit(X_train, y_train)
# +
knn_scores = cross_val_score(knn_class, X_train, y_train, cv=3)
knn_rmse_scores = np.sqrt(-knn_scores)
print(knn_rmse_scores)
print('Mean:', scores.mean())
print('Standard Dev:', scores.std())
# +
from sklearn.model_selection import GridSearchCV
param_grid = [
{ 'weights': []}
]
| learning_exercises/hands_on_machine_learning/hands_on_ml_ch_3_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Performance
# ## Profile Decoder
#
# The default Python and Cython decoder can be profiled with Python's standard `cprofile`. The output can be a sorted table and a flame graph. Both are generated below:
# + language="bash"
# python -m openpifpaf.predict coco/000000081988.jpg --no-download-progress --debug --profile-decoder
# -
# !flameprof profile_decoder.prof > profile_decoder_flame.svg
# 
# There is a second output that is generated from the Autograd Profiler. This can only be viewed in the Chrome browser:
# * open `chrome://tracing`
# * click "Load" in the top left corner
# * select `decoder_profile.1.json`
#
# This is the same type of plot that is used to trace the training of a batch. An example of such a plot is shown below.
#
#
# ## Profile Training
#
# For a training batch, the Chrome trace looks like this:
#
# 
| guide/performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df1 = pd.DataFrame()
df2 = pd.DataFrame()
df1['viewers'] = ["Sushmita", "Aditya", "Bala", "Anurag"]
df2['users'] = ["Aditya", "Anurag", "Bala", "Sushmita", "Apoorva"]
np.random.seed(1729)
df1 = df1.assign(views = np.random.normal(100, 100, 4))
df2 = df2.assign(cost = [20, np.nan, 15, 2, 7])
df1.head()
df2.head()
df = df1.merge(df2, left_on="viewers", right_on="users", how="left")
df.head()
df.fillna(df.mean())
| Lesson01/Exercise 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# # Image segmentation with a U-Net-like architecture
#
# **Author:** [fchollet](https://twitter.com/fchollet)<br>
# **Date created:** 2019/03/20<br>
# **Last modified:** 2020/04/20<br>
# **Description:** Image segmentation model trained from scratch on the Oxford Pets dataset.
# + [markdown] colab_type="text"
# ## Download the data
#
# + colab_type="code"
# !curl -O http://www.robots.ox.ac.uk/~vgg/data/pets/data/images.tar.gz
# !curl -O http://www.robots.ox.ac.uk/~vgg/data/pets/data/annotations.tar.gz
# !tar -xf images.tar.gz
# !tar -xf annotations.tar.gz
# + [markdown] colab_type="text"
# ## Prepare paths of input images and target segmentation masks
#
# + colab_type="code"
import os
input_dir = "images/"
target_dir = "annotations/trimaps/"
img_size = (160, 160)
num_classes = 4
batch_size = 32
input_img_paths = sorted(
[
os.path.join(input_dir, fname)
for fname in os.listdir(input_dir)
if fname.endswith(".jpg")
]
)
target_img_paths = sorted(
[
os.path.join(target_dir, fname)
for fname in os.listdir(target_dir)
if fname.endswith(".png") and not fname.startswith(".")
]
)
print("Number of samples:", len(input_img_paths))
for input_path, target_path in zip(input_img_paths[:10], target_img_paths[:10]):
print(input_path, "|", target_path)
# + [markdown] colab_type="text"
# ## What does one input image and corresponding segmentation mask look like?
#
# + colab_type="code"
from IPython.display import Image, display
from tensorflow.keras.preprocessing.image import load_img
import PIL
from PIL import ImageOps
# Display input image #7
display(Image(filename=input_img_paths[9]))
# Display auto-constrast version of corresponding target (per-pixel categories)
img = PIL.ImageOps.autocontrast(load_img(target_img_paths[9]))
display(img)
# + [markdown] colab_type="text"
# ## Prepare `Sequence` class to load & vectorize batches of data
#
# + colab_type="code"
from tensorflow import keras
import numpy as np
from tensorflow.keras.preprocessing.image import load_img
class OxfordPets(keras.utils.Sequence):
"""Helper to iterate over the data (as Numpy arrays)."""
def __init__(self, batch_size, img_size, input_img_paths, target_img_paths):
self.batch_size = batch_size
self.img_size = img_size
self.input_img_paths = input_img_paths
self.target_img_paths = target_img_paths
def __len__(self):
return len(self.target_img_paths) // self.batch_size
def __getitem__(self, idx):
"""Returns tuple (input, target) correspond to batch #idx."""
i = idx * self.batch_size
batch_input_img_paths = self.input_img_paths[i : i + self.batch_size]
batch_target_img_paths = self.target_img_paths[i : i + self.batch_size]
x = np.zeros((batch_size,) + self.img_size + (3,), dtype="float32")
for j, path in enumerate(batch_input_img_paths):
img = load_img(path, target_size=self.img_size)
x[j] = img
y = np.zeros((batch_size,) + self.img_size + (1,), dtype="uint8")
for j, path in enumerate(batch_target_img_paths):
img = load_img(path, target_size=self.img_size, color_mode="grayscale")
y[j] = np.expand_dims(img, 2)
return x, y
# + [markdown] colab_type="text"
# ## Perpare U-Net Xception-style model
#
# + colab_type="code"
from tensorflow.keras import layers
def get_model(img_size, num_classes):
inputs = keras.Input(shape=img_size + (3,))
### [First half of the network: downsampling inputs] ###
# Entry block
x = layers.Conv2D(32, 3, strides=2, padding="same")(inputs)
x = layers.BatchNormalization()(x)
x = layers.Activation("relu")(x)
previous_block_activation = x # Set aside residual
# Blocks 1, 2, 3 are identical apart from the feature depth.
for filters in [64, 128, 256]:
x = layers.Activation("relu")(x)
x = layers.SeparableConv2D(filters, 3, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.Activation("relu")(x)
x = layers.SeparableConv2D(filters, 3, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.MaxPooling2D(3, strides=2, padding="same")(x)
# Project residual
residual = layers.Conv2D(filters, 1, strides=2, padding="same")(
previous_block_activation
)
x = layers.add([x, residual]) # Add back residual
previous_block_activation = x # Set aside next residual
### [Second half of the network: upsampling inputs] ###
previous_block_activation = x # Set aside residual
for filters in [256, 128, 64, 32]:
x = layers.Activation("relu")(x)
x = layers.Conv2DTranspose(filters, 3, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.Activation("relu")(x)
x = layers.Conv2DTranspose(filters, 3, padding="same")(x)
x = layers.BatchNormalization()(x)
x = layers.UpSampling2D(2)(x)
# Project residual
residual = layers.UpSampling2D(2)(previous_block_activation)
residual = layers.Conv2D(filters, 1, padding="same")(residual)
x = layers.add([x, residual]) # Add back residual
previous_block_activation = x # Set aside next residual
# Add a per-pixel classification layer
outputs = layers.Conv2D(num_classes, 3, activation="sigmoid", padding="same")(x)
# Define the model
model = keras.Model(inputs, outputs)
return model
# Free up RAM in case the model definition cells were run multiple times
keras.backend.clear_session()
# Build model
model = get_model(img_size, num_classes)
model.summary()
# + [markdown] colab_type="text"
# ## Set aside a validation split
#
# + colab_type="code"
import random
# Split our img paths into a training and a validation set
val_samples = 1000
random.Random(1337).shuffle(input_img_paths)
random.Random(1337).shuffle(target_img_paths)
train_input_img_paths = input_img_paths[:-val_samples]
train_target_img_paths = target_img_paths[:-val_samples]
val_input_img_paths = input_img_paths[-val_samples:]
val_target_img_paths = target_img_paths[-val_samples:]
# Instantiate data Sequences for each split
train_gen = OxfordPets(
batch_size, img_size, train_input_img_paths, train_target_img_paths
)
val_gen = OxfordPets(batch_size, img_size, val_input_img_paths, val_target_img_paths)
# + [markdown] colab_type="text"
# ## Train the model
#
# + colab_type="code"
# Configure the model for training.
# We use the "sparse" version of categorical_crossentropy
# because our target data is integers.
model.compile(optimizer="rmsprop", loss="sparse_categorical_crossentropy")
callbacks = [
keras.callbacks.ModelCheckpoint("oxford_segmentation.h5", save_best_only=True)
]
# Train the model, doing validation at the end of each epoch.
epochs = 15
model.fit(train_gen, epochs=epochs, validation_data=val_gen, callbacks=callbacks)
# + [markdown] colab_type="text"
# ## Visualize predictions
#
# + colab_type="code"
# Generate predictions for all images in the validation set
val_gen = OxfordPets(batch_size, img_size, val_input_img_paths, val_target_img_paths)
val_preds = model.predict(val_gen)
def display_mask(i):
"""Quick utility to display a model's prediction."""
mask = np.argmax(val_preds[i], axis=-1)
mask = np.expand_dims(mask, axis=-1)
img = PIL.ImageOps.autocontrast(keras.preprocessing.image.array_to_img(mask))
display(img)
# Display results for validation image #10
i = 10
# Display input image
display(Image(filename=val_input_img_paths[i]))
# Display ground-truth target mask
img = PIL.ImageOps.autocontrast(load_img(val_target_img_paths[i]))
display(img)
# Display mask predicted by our model
display_mask(i) # Note that the model only sees inputs at 150x150.
| examples/vision/ipynb/oxford_pets_image_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### News Mood
#
# In this activity, we are going to review Tweepy.
#
# BBC, CBS, CNN, Fox, and New York times
# - - -
# +
# Dependencies
import numpy as np
import pandas as pd
import tweepy
import json
import time
from datetime import datetime
import matplotlib.pyplot as plt
from matplotlib import style
style.use('ggplot')
from config import (consumer_key, consumer_secret,
access_token, access_token_secret)
# Setup Tweepy API Authentication
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, parser=tweepy.parsers.JSONParser())
# Import and Initialize Sentiment Analyzer
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
# -
# "Real Person" Filters
min_tweets = 5
max_tweets = 10000
max_followers = 2500
max_following = 2500
lang = "en"
api.search("g<NAME>")
# +
# Search for People Tweeting about <NAME>
search_term = "<NAME>"
# Create variable for holding the oldest tweet
oldest_tweet = None
# List to hold unique IDs
unique_ids = []
results = []
# Counter to keep track of the number of tweets retrieved
counter = 0
# Loop through 5 times (total of 500 tweets)
for x in range(5):
# Retrieve 100 most recent tweets -- specifying a max_id
public_tweets = api.search(search_term,
count=100,
result_type="recent",
max_id=oldest_tweet)
# Print Tweets
for tweet in public_tweets["statuses"]:
tweet_id = tweet_id = tweet["id"]
# Use filters to check if user meets conditions
if (tweet["user"]["followers_count"] < max_followers and
tweet["user"]["statuses_count"] > min_tweets and
tweet["user"]["statuses_count"] < max_tweets and
tweet["user"]["friends_count"] < max_following and
tweet["user"]["lang"] == lang):
# Print the username
#print(tweet["user"]["screen_name"])
# Print the tweet id
#print(tweet["id_str"])
# Print the tweet text
#print(tweet["text"])
#print()
# Print the tweet date
#print(tweet["created_at"])
#print()
# Append tweet_id to ids list if it doesn't already exist
# This allows checking for duplicate tweets
if tweet_id not in unique_ids:
unique_ids.append(tweet_id)
# Increase counter by 1
counter += 1
results.append({"User": tweet["user"]["screen_name"],
"Tweet": tweet["text"],
"Date": tweet["created_at"],
"Tweet ID Str": tweet["id_str"],
"Tweet ID": tweet["id"],
"Candidate": search_term})
# Reassign the the oldest tweet (i.e. the max_id)
# Subtract 1 so the previous oldest isn't included
# in the new search
oldest_tweet = tweet_id - 1
# -
len(results)
results_pd = pd.DataFrame.from_dict(results)
results_pd.head()
# Print total number of tweets retrieved
print(counter)
# Print the number of unique ids retrieved
print(len(unique_ids))
# Export to CSV
file_name = str(time.strftime("%m-%d-%y")) + "-Tweets.csv"
results_pd.to_csv(file_name, encoding="utf-8")
| 18.06.21 - Project1/Archive (Delete)/Project1 - Carla.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://www.upgto.edu.mx/itm/images/upg.png" width="200px"><img></center>
# **Universidad Politécnica de Guanajuato**
# **Ingeniería Robótica**
# **Mecanismos y máquinas**
# **Análisis cinemático de mecanismos planos utilizando software**
#
# # Ejemplo Clase 06/11/2017 01. Mecanismos
# +
import numpy as np
import matplotlib.pyplot as plt
from numpy import sin,cos,tan,pi,arcsin
# %matplotlib inline
def plotvector(p0, u, *args, **kwargs):
plt.plot([p0[0],p0[0]+u[0]], [p0[1],p0[1]+u[1]], *args, **kwargs)
w = -5
r = 1
for theta in np.linspace(0,2*pi,10):
R = np.array([r*cos(theta), r*sin(theta)])
V = np.array([-r*w*sin(theta),r*w*cos(theta)])/15
A = np.array([-r*(w**2)*cos(theta),-r*(w**2)*sin(theta)])/100
plotvector([0,0], R, 'r')
plotvector(R, V, 'g-*')
plotvector(R, A, 'b-*')
plt.axis('equal');
# -
# # <NAME>
#
r2,r3 = 3,8
for t2 in np.linspace(0,2*pi,6,endpoint=False):
color = np.array([t2,t2,t2])/(2*pi)
t3 = np.arcsin(-r2*sin(t2)/r3)
r1 = r2*cos(t2) + r3*cos(t3)
R2 = np.array([r2*cos(t2),r2*sin(t2)])
R3 = np.array([r3*cos(t3),r3*sin(t3)])
plotvector([0,0], R2, color=color)
plotvector(R2, R3, color=color)
plt.axis('equal');
| nbooks/Clase_06112017.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. hybrid-vocal-classifier autolabel workflow
# Here's the steps in the workflow for autolabeling vocalizations.
#
# First we import the library, since in Python you need to `import` a library before you can work with it.
import hvc # in Python we have to import a library before we can use it
# ### 0. Label a small set of songs to provide **training data** for the models, typically ~20 songs.
# Here we download the data from a repository.
# ** You don't need to run this if you've already downloaded the data.**
hvc.utils.fetch('gy6or6.032212')
hvc.utils.fetch('gy6or6.032612')
# ### 1. Pick a machine learning algorithm/**model** and the **features** used to train the model.
#
# In this case we'll use the k-Nearest Neighbors (k-NN) algorithm because it's fast to apply to our data. We'll use the features built into the library that have been tested with k-NN.
#
# Picking a model and the features that go with it is simple:
# 1. In a text editor, open `gy6or6_autolabel.example.knn.extract.config.yml`
# 2. Below the line that says `feature group:` add `knn` after the dash.
# 3. Below the line that says `data_dirs:` add the path to the data you downloaded after the dash.
# ### 2. Extract features for that model from song files that will be used to train the model.
#
# We call the `extract` function and we pass it the name of the `yaml` config file as an argument.
#
# ```Python
# # 1. pick a model and 2. extract features for that model
# # Model and features are defined in extract.config.yml file.
# hvc.extract('gy6or6_autolabel.example.extract.knn.config.yml')
# ```
hvc.extract('gy6or6_autolabel.example.extract.knn.config.yml')
# ### 3. Pick the **hyperparameters** used by the algorithm as it trains the model on the data.
# Now in Python we use some convenience functions to figure out which "hyperparameters" will give us the best accuracy when we train our machine learning models.
# ```Python
# # 3. pick hyperparameters for model
# # Load summary feature file to use with helper functions for
# # finding best hyperparameters.
# from glob import glob
# summary_file = glob('./extract_output*/summary*')
# summary_data = hvc.load_feature_file(summary_file)
# # In this case, we picked a k-nearest neighbors model
# # and we want to find what value of k will give us the highest accuracy
# X = summary_data['features']
# y = summary_data['labels']
# cv_scores, best_k = hvc.utils.find_best_k(X,y,k_range=range(1, 11))
# ```
# ### 4. Train, i.e., fit the **model** to the data
# ### 5. Select the **best** model based on some measure of accuracy.
#
# 1. In a text editor, open `gy6or6_autolabel.example.knn.select.config.yml`
# 2. On the line that says `feature_file:` paste the name of the feature file after the colon. The name will have a format like `summary_file_bird_ID_date`.
#
# Then run the following code in the cell below:
# ```Python
# # 4. Fit the **model** to the data and 5. Select the **best** model
# hvc.select('gy6or6_autolabel.example.select.knn.config.yml')
# ```
# !gedit gy6or6_autolabel.example.select.knn.config.yml
# cd hybrid-vocal-classifier-tutorial/
hvc.select('gy6or6_autolabel.example.select.knn.config.yml')
# ### 6. Using the fit model, **Predict** labels for unlabeled data.
# 1. In a text editor, open `gy6or6_autolabel.example.knn.predict.config.yml`
# 2. On the line that says `model_meta_file:`, after the colon, paste the name of a meta file from the `select` output. The name will have a format like `summary_file_bird_ID_date`.
# 3. Below the line that says `data_dirs:`, after the dash, add the path to the other folder of data that you downloaded.
#
# Then run the following code in the cell below.
# ```Python
# # 6. **Predict** labels for unlabeled data using the fit model.
# hvc.predict('gy6or6_autolabel.example.predict.knn.config.yml')
# ```
# cd select_output_171205_193932/knn_k4/
# ls
# cd hybrid-vocal-classifier-tutorial/
hvc.predict('gy6or6_autolabel.example.predict.knn.config.yml')
# Congratulations! You have auto-labeled an entire day's worth of data.
| notebooks/05-autolabel-workflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# ---
# # 落書き認識Webアプリを作ろう
#
# - Author: <NAME> ([github](https://github.com/ornew), [facebook](https://www.facebook.com/old.r.new))
# - Contributor: <NAME> ([github](https://github.com/hideya))
# ---
# このノートブックは、セミナーの資料として作成されています。ノートブックは、自由な編集、実行が可能です。Markdown形式でドキュメントも書き込めるため、必要に応じてメモを追記するなど、工夫してご利用ください。
#
# 編集も含めて保存したい場合は、画面上部のツールバーから、【File】タブを選び、【Download as】を選ぶことでローカルマシン上に保存することが可能です。
#
# このノートブックは自由にご利用頂けますが、インターネット上への無断での転載だけはご遠慮くださいますようお願いします。
# ## ハンズオンの概要
#
# Google が ["the Quick, Draw!"データセット](https://quickdraw.withgoogle.com/data) として公開している多量の落書きの画像データを用いてニューラルネットワークを訓練し、何の落書きかを認識するニューラルネットを生成します。そしてその学習済みニューラルネットワークを組み込んだ、落書きの分類をする Web App を作成します。以下は、最終的な Web App のスクリーンショットです。実際のアプリは [ここをクリックすることで実行](https://tfjs-doodle-recognition-pwa.netlify.com/) できます。
#
# 
#
# ニューラルネットワークのモデルとしては、手書き数字([MNIST](https://en.wikipedia.org/wiki/MNIST_database))の認識で実績のある [Convolutional Neural Network (CNN) ](https://en.wikipedia.org/wiki/Convolutional_neural_network) を用います。
#
# モデルの構築と学習には、[AWS SageMaker](https://aws.amazon.com/jp/sagemaker/) 上で、
# [TensorFlow](https://www.tensorflow.org/) を使用します。
#
# Web App の構築には、JavaScript でニューラルネットワークの実装とブラウザ上での実行を可能とする [TensorFlow.js](https://js.tensorflow.org/) を利用します。
#
# 以下に続くノートブックで、それぞれの過程を詳しく説明します(なお、ニューラルネットワークの学習を行う際、数十円の課金が発生します)。
# ---
# #### ご参考:ノートブックの操作方法
#
# ノートブックは、自由な編集、実行が可能です。Markdown形式でドキュメントも書き込めるため、必要に応じてメモを追記するなど、工夫してご利用ください。
#
# 以下に良く使うキー操作を列挙します:
#
# |キー操作| 説 明 | | キー操作 | 説 明 |
# |--|--| |--|--|
# | Enter | 編集モードに入る | | Esc → A | 新規セルを上に追加 |
# | Shift + Enter | セルを実行し / 編集モードから抜け、下のセルに移動 | | Esc → B | 新規セルを下に追加 |
# | Cntl + Enter | セルを実行する / 編集モードから抜ける | |Esc → D, D | セルを削除 |
# | Esc → M | セルをマークダウンモードに変更 | | Esc → L | セルの行番号の表示・非表示 |
# | Esc → Y | セルをコードモードに変更 | | Esc → H | キーボード・ショートカットの一覧の表示 |
#
# - ノートブックを初期状態に戻したい(全ての実行結果の消去とカーネルのリスタートをしたい)場合は、
# 画面上部のツールバーから **Kernel → Restart & Clear Output** を選択します。
#
# - 編集済みのノートブックをローカルにセーブしたい場合は、ツールバーから **File → Download as → Notebook**
# を選択します。
#
# ---
# ## モデルの概要
# 今回、認識する落書きは、以下の10クラス(種類)です。
#
# 1. りんご (apple)
# 2. ベッド (bed)
# 3. 猫 (cat)
# 4. 犬 (dog)
# 5. 目 (eye)
# 6. 魚 (fish)
# 7. 草 (grass)
# 8. 手 (hand)
# 9. アイスクリーム (ice cream)
# 10. ジャケット (jacket)
# 28x28ピクセルのグレースケール画像から、上記のいずれの落書きであるかを**確率的に**予測します。
#
# 
# ### ディープラーニング
#
# モデルは(ディープ)ニューラルネットワークで実装します。
#
# ニューラルネットワークとは、生物のニューロン(神経細胞)のネットワークを数理モデルで模倣することで、特定の課題解決能力を機械的に学習する、機械学習アルゴリズムの一種です。深い層で構成されるニューラルネットワークの学習を行うことをディープラーニングといいます。
#
# ディープラーニングにおけるモデルの学習は、以下の流れで行います。
#
# - ⓪ モデルのパラメータを初期化する
# - ① 学習用データに対する予測を計算する
# - ② 教師ラベルと予測結果の誤差を計算する
# - ③ 誤差を最小化するようにモデルのパラメータを更新する
# - ④ **誤差が十分に小さくなるまで**①-③を繰り返す
#
# 
# ## 実装の流れ
#
# このノートブックでは、以下の手順で、ディープラーニングを用いた落書き(Doodle)認識を行うWebアプリを作成します。
#
# 1. ["the Quick, Draw!"データセット](https://quickdraw.withgoogle.com/data)を学習用データとして準備する
# 2. [TensorFlow](https://www.tensorflow.org/)で落書きを認識するディープニューラルネットワークのモデルを実装する
# 3. [Amazon SageMaker](https://aws.amazon.com/jp/sagemaker/)でモデルを学習する
# 4. [TensorFlow.js](https://js.tensorflow.org/)を使ったWebアプリに学習済みモデルを組み込む
# 5. [Amazon S3](https://aws.amazon.com/jp/s3/)でWebアプリを公開する
#
# 
# ## 実装する
# まず、作業に必要な Python のモジュールをノートブック・インスタンスに読み込みます。
#
# 実行ログの出力が始まるまで、少々(30秒程度)時間がかかりますので、しばらく反応がなくてもすこし様子を見てください。
# +
import tensorflow as tf
import six # Python 2と3の互換性を保つためのライブラリです
import numpy as np # 行列などの科学数値計算をするためのライブラリです
import matplotlib.pyplot as plt # グラフを描画するライブラリです
# %matplotlib inline
# 繰り返し処理の進捗をプログレスバーで表示するためのライブラリです
# !pip install tqdm msgpack
from tqdm import tqdm_notebook as tqdm
# -
# ### ① 学習用データを準備する
# 学習データは、Google社が[クリエイティブ・コモンズ ライセンス バージョン4.0](https://creativecommons.org/licenses/by/4.0/)で公開している["the Quick, Draw!"データセット](https://quickdraw.withgoogle.com/data)を利用します。
# #### データをダウンロードする
#
# データをダウンロードして、`./raw_data`ディレクトリに保存します。
#
# ちなみに、Jupyterノートブックでは、「`!`」を先頭につけると、シェルコマンドを実行できます(Pythonの機能ではありません)。出力をPythonで使ったり、Pythonの変数を引数に使ったりも出来るので便利です。ここでは`wget`コマンドを使ってファイルをダウンロードします。
URL = 'https://storage.googleapis.com/quickdraw_dataset/full/numpy_bitmap'
LABELS = [
'apple', 'bed', 'cat', 'dog', 'eye',
'fish', 'grass', 'hand', 'ice cream', 'jacket',
]
# !rm -rf ./data ./raw_data
# !mkdir -p ./data ./raw_data
for l in LABELS:
url = '{}/{}.npy'.format(URL, l)
# !wget -P raw_data "$url"
# 各ラベルのデータファイルがダウンロードできていることを確認します。
# !ls -l ./raw_data
# ダウンロードしたデータを配列 (numpy.ndarray) に読み込みます。
raw_data = {label: np.load('raw_data/{}.npy'.format(label)) for label in tqdm(LABELS)}
# 各データの数を確認してみましょう。
for label, data in six.iteritems(raw_data):
print('{:10}: {}'.format(label, len(data)))
# ためしに、1番目の「猫」の画像を表示してみます。
plt.imshow(np.reshape(raw_data['cat'][0], [28, 28]), cmap='gray')
plt.show()
# #### 学習用と評価用のデータを準備する
# 次に、データを学習用と評価用に分けます。
#
# 学習に使ったデータは、モデルがすでに「知っている」データなので、そのモデルが本当に役に立つのかを評価するためには学習に使っていない「未知のデータ」に対する精度を確認する必要があります。ですので、ダウンロードしたデータから、学習用と評価用の2種類のデータを予め準備します。
#
# 1. ダウンロードしたデータセットのうち1万件を取り出す *1
# - クラスごとに数にばらつきがあると、学習で用いられる頻度がクラスごとに変わってしまうため、揃えます
# 2. それぞれ画像データと教師ラベルの組み合わせに変換する
# - 教師ラベルは、クラスの名前(例:apple)ではなく、それぞれクラスごとにユニークな数字を割り当てます(下で確認)
# 3. 学習用と評価用に7:3で分ける
# - 学習に使われていないデータで精度の評価を行いたいため、3割を評価用のデータとして使います
# 4. ランダムにシャッフル
#
# *1 … もとは「10万」だったのですが、時間単価の安い(初期無料枠のある)「t2.medium」ノートブック・インスタンスで
# メモリ不足を起こさず扱えるように少なくしています。
# 結果として、認識の精度(描画のブレへの対応力)は少々下がります。
# (SageMaker の料金一覧は[こちら](https://aws.amazon.com/jp/sagemaker/pricing/))
# 学習に先立って、画像の各クラスを出力ニューロンと対応づけます。
# ニューラルネットワーク (CNN) の認識結果は、各出力ニューロンの状態(0から1の値)により得られます。
# そのため、それぞれの出力ニューロンを、判定するクラスと1対1に対応づけます。
# つまり、10個の出力ニューロンを用意し、そのn番目のニューロンを、n番目のクラスと対応づけます。
# ここでは、ラベル配列のインデクスをそのまま両者を対応付ける序数として使います。
for i, label_name in enumerate(LABELS):
print(u'出力ニューロン 番号: {} クラス名: {}'.format(i, label_name))
# 次に、データを前処理して、ニューラルネットワークの訓練に用いるのに適した形式に変換します。
# 今回は、メモリ不足を起こさないように、また学習スピードを上げるために、最初にデータの個数を1万に削減します。
# もともとは「10万」個のデータを使用していたのですが、
# 無料枠のある「t2.medium」ノートブック・インスタンスで処理をしてもメモリ不足を起こさないようにするために、
# 1万個に削減しています。 またこれにより、学習時間が短縮できます。
# 結果として、認識の精度(描画のブレへの対応力)は少々下がります。
# +
for label, data in six.iteritems(raw_data):
raw_data[label] = raw_data[label][:10000]
for label, data in six.iteritems(raw_data):
print('{:10}: {}'.format(label, len(data)))
# -
# 各ピクセルの値が0から1に収まるように正規化し、学習用と評価用のデータに分割します。
train_data = []
test_data = []
for label_name, value in six.iteritems(raw_data):
label_index = LABELS.index(label_name)
print('proccessing label class {}: "{}"'.format(label_index, label_name))
# 各ピクセルの値を、0-255から0-1に修正します
value = np.asarray(value) / 255.
# 7万件を学習用のデータとして画像データと教師ラベルの組み合わせにしてリストに追加します
train_data.extend(zip(value[:7000], np.full(7000, label_index)))
# 3万件を評価用のデータとして画像データと教師ラベルの組み合わせにしてリストに追加します
test_data.extend(zip(value[7000:10000], np.full(3000, label_index)))
np.random.shuffle(train_data)
np.random.shuffle(test_data)
# 次に、これら学習用と評価用のデータを、ニューラルネットワークの訓練での利用に適した
# TFRecord 形式のファイルに変換して出力します。
# TFRecord は [Protocol Buffers](https://developers.google.com/protocol-buffers/) というフォーマットを用いたデータファイルで、構造化されている・圧縮効率が高い・読み書きの速度が非常に速い・非同期のストリーミング読み込みが可能
# とった長所があり、機械学習で用いられる大規模データセットの保存に向いています。
# まず、ヘルパー関数を定義します。
# +
train_filename = './data/train.tfr'
test_filename = './data/test.tfr'
def get_example_proto(image, label):
"""
画像とラベルをProtocol Buffers形式のtf.train.Exampleに変換します
"""
return tf.train.Example(features=tf.train.Features(feature={
'image' : tf.train.Feature(float_list=tf.train.FloatList(value=image)),
'label' : tf.train.Feature(int64_list=tf.train.Int64List(value=label)),
})).SerializeToString()
# -
# 以下の変換処理は30秒ほどかかります。
# %%time
tfr_options = tf.python_io.TFRecordOptions(tf.python_io.TFRecordCompressionType.GZIP)
with tf.python_io.TFRecordWriter(train_filename, tfr_options) as train_tfr, \
tf.python_io.TFRecordWriter(test_filename, tfr_options) as test_tfr:
print('Converting train data...')
for data, label in tqdm(train_data):
train_tfr.write(get_example_proto(data, [label]))
print('Converting test data...')
for data, label in tqdm(test_data):
test_tfr.write(get_example_proto(data, [label]))
# `train.tfr`と`test.tfr`が生成されていれば成功です。
# !ls -l data
# これで、データの準備が完了しました。
#
# 生成したデータは、学習モデルの設定を終えた後、学習プロセスを開始する前に、
# S3 にアップロードして、学習用インスタンスからアクセスできるようにします。
# ### ② TensorFlowでモデルの定義プログラムを実装する
# モデルの実装には、[TensorFlow](https://www.tensorflow.org/)を利用します。TensorFlowは、Google社が主体となって開発している、オープンソースの汎用的な分散数値演算ライブラリです。TensorFlowにはディープラーニング向けのライブラリが用意されています。GitHubのスターは10万近くあり、現在世界で最も人気のディープラーニングフレームワークとも言われています。
# 以下の4つの関数を定義したプログラムを用意すると、Amazon SageMakerを使ってモデルの学習を行うことができます。
# ```python
# def train_input_fn(training_dir, hyperparameters):
# """
# 学習用の入力データを読み込みます。
#
# training_dir: 学習の実行時に指定したS3のファイルがこの文字列のディレクトリにマウントされています。
# hyperparameters: 学習の実行時に指定したハイパーパラメータが渡されます。
#
# 基本的には、以下のことを実装するだけです。
# ① hyperparametersで指定した挙動に従って、
# ② training_dirから学習データを読み込み、データを返す。
# """
#
# def eval_input_fn(training_dir, hyperparameters):
# """
# 評価用の入力データを読み込みます。
# やることはtrain_input_fnと同じですが、評価用のデータを読み込むことや、
# 評価用に挙動を変える(例えば評価データはシャッフルしないなど)ことが可能です。
# """
#
# def serving_input_fn(hyperparameters):
# """
# モデルの入力データの形式を定義します。
# サービングと付いている通り、SageMakerでAPIサーバにデプロイしたときの入力データ定義にもなります。
# """
#
# def model_fn(features, labels, mode, hyperparameters):
# """
# モデルの定義をします
#
# features: モデルの入力と成る特徴データです *_input_fnで返した値がそのまま渡されます。
# labels: モデルの教師ラベルデータです。
# mode: モデルの実行モードです。実行モードには「学習」「評価」「推論」があり、挙動を切り替えることが可能です。
# hyperparameters: 実行時に指定したハイパーパラメータが渡されます。
# """
# ```
# 最後の`model_fn`が、その名のとおり、ニューラルネットワークの定義の本体です。
#
# `model_fn`の中では、以下の3つを定義します。
#
# 1. **モデル**: ニューラルネットワーク
# 2. **誤差**: 教師データと予測結果がどの程度違ったのかを定式化する
# 3. **最適化アルゴリズム**: 誤差を最小化するようにモデルを最適化するアルゴリズム
#
# つまり、データの入力方法と、上記3つのモデル関連の定義を行うだけで、機械学習を行うことができてしまいます。
# 今回、セミナー用のモデル定義は予め実装してあります(`src/doodle.py`ファイル)。
#
# 以下を実行して、その内容を確認してみましょう。
# コメントなどを含めても200行弱程度しかありません。
# コードには多くのコメントが付けてありますので、ざっと目を通してみてください。
# !cat src/doodle.py
# ニューラルネットワークのモデルの定義や学習に関する詳細は、別途ノートブック`model.ipynb`で解説しています。ニューラルネットワークの実装に興味がある方はそちらをご参照ください。
# ### ③ Amazon SageMakerでモデルを学習する
# Amazon SageMaker SDKを使い、ここまでで準備したデータとプログラムを指定して学習を実行します。
#
# 
# #### 設定情報を定義する
#
# モデルの学習を始めるにあたり、学習に使用するデータの保存先などの設定情報を変数で定義します。
# ここで、学習ジョブの名前を定義しますので、もし再度、学習を繰り返したい場合には、ここから以下を再実行します。
# +
import sagemaker
from datetime import datetime
import six
role = sagemaker.get_execution_role()
session = sagemaker.Session()
bucket = session.default_bucket()
timestamp = datetime.now().strftime("%Y-%m-%d-%H-%M-%S")
def _s3(path):
return 's3://{}/doodle-{}/model/{}'.format(bucket, timestamp, path)
data_key_prefix = 'doodle-{}/model/data'.format(timestamp)
config = dict(
data_dir = _s3('data'),
output_path = _s3('export'),
checkpoint_path = _s3('ckpt'),
code_location = _s3('src'),
public_dir = _s3('public'),
job_name = 'doodle-training-job-{}'.format(timestamp)
)
# -
# 確認のため、設定した変数を表示します。
for k, v in six.iteritems(config):
print('key: {:20}, value: {:20}'.format(k, v))
# #### S3 にデータをアップロードする
#
# 上記で設定した S3 パスに、①で作成したデータセットをアップロードし、学習インスタンスが学習データにアクセスできるようにします。
# +
uploaded_data_dir = session.upload_data(
'data', # ローカルディレクトリ
bucket=bucket, # アップロードするS3バケット名
key_prefix=data_key_prefix) # アップロードするパスのプリフィクス
# 設定と同じ場所になったか念のため確認します
assert uploaded_data_dir == config['data_dir']
# -
# これで、学習用と評価用のデータの準備がおわりました。
# 次に、学習を行うモデルを構築します。
# モデルの学習では、「エスティメータ(Estimator)」を利用します。 エスティメータとは、モデルの学習や評価、保存やデプロイといった一連の処理を簡便に行うための、高レベルのインターフェイスです。
#
# 用意したパスなどを設定として渡して、エスティメータを作成します。
# エスティメータへのパラメータとして、`entry_point`に`doodle.py`が指定されていることに注目してください。
# この Python のプログラム`doodle.py`で、②で述べた、ニューラルネットワークのモデルやそれに関わるシステムの挙動が定義されています。
# +
from sagemaker.tensorflow import TensorFlow
estimator = TensorFlow(
# ハイパーパラメータ
# ②で定義したプログラムの各関数の引数に渡されます
# プログラムの挙動を切り替えるのに利用できます
hyperparameters={
'save_summary_steps': 100,
'throttle_secs': 120,
},
# 先程設定した、各データの保存先のパス
output_path = config['output_path'],
checkpoint_path = config['checkpoint_path'],
code_location = config['code_location'],
# 学習用プログラムに関する設定
source_dir='./src', # 学習用のプログラムが保存されたローカルディレクトリ
entry_point='doodle.py', # ②で定義した学習用プログラムのファイル名
framework_version='1.6', # 利用したいTensorFlowのバージョン
# 学習と評価の回数
training_steps=10000,
evaluation_steps=1000,
# AWSでの実行に関する設定
role=role,
train_instance_count=1,
train_instance_type='ml.p2.xlarge') # ml.p2.xlargeはGPUの搭載されたインスタンスです
# -
# このエスティメータに対して、学習用データのパス名を指定して`fit`関数を呼び出すと、学習ジョブが作成され、クラウド上でモデルの学習を実行します。
# この学習には10分弱かかります。
# その間、学習中の状態を確認するために、ノートブック上で
# [TensorBoard](https://www.tensorflow.org/programmers_guide/summaries_and_tensorboard)
# を起動します
# (そのために`run_tensorboard_locally`引数に`True`を渡します)。
#
# なお、この学習では、1回で、数十円の課金が発生します
# (ご参考:「[Amazon SageMaker の料金](https://aws.amazon.com/jp/sagemaker/pricing/)」)。
#
# それでは、以下のセルで、学習をスタートしてみましょう。
# %%time
estimator.fit(config['data_dir'], job_name=config['job_name'],
wait=True, run_tensorboard_locally=True)
# ノートブック上で実行される TensorBoard は、学習経過を表示するように設定してあります。
#
# [ここをクリックすることにより](/proxy/6006/)、TensorBoard の画面がブラウザで開き、学習経過が確認できます
# (実行ログ中に表示される`http://localhost:6006`ではアクセスできません。
# `https://(ノートブックのURL)/`[proxy/6006/](/proxy/6006/) にアクセスする必要があります)。
#
# 学習のセットアップをしている最初のうちは「No dashboards are active for the current data set」とだけ表示されますが、
# 4〜5分経って、上のセルにログが出力され始めると、学習経過を示すグラフが表示されるようになります。
# ログが表示されはじめるのを待ってから、TensorBoard のページを確認してみてください。
# 上の学習ジョブが終了したら、学習したモデルをチェックしてみましょう。
#
# 学習済みモデルのファイルは、エスティメータのoutput_path引数で指定した場所に保存されています。
output_dir_url = '{}/{}/output/'.format(config['output_path'], config['job_name'])
# !echo $output_dir_url
# !aws s3 ls $output_dir_url
# ### ④ 学習したモデルをダウンロードして、Webアプリに組み込む
# 学習したモデルは、エスティメータの`output_path`引数で指定した場所にGZIP圧縮されたTarアーカイブとして保存されています。中身はTesnorFlow SavedModelと呼ばれるデータ形式です。
#
# 
#
# このモデルデータを使えば、Pythonで実行したり、TensorFlow ServingでAPIサーバを構築したり、TensorFlow Liteを使ってAndroidやiOSで実行したりすることが可能です。
#
# 今回は、TensorFlow.jsを使って、Webブラウザ上でモデルの推論を実行してみましょう。
# まず、学習済みモデルデータをS3からノートブック・インスタンスにダウンロードして解凍します。
model_url = '{}/{}/output/model.tar.gz'.format(config['output_path'], config['job_name'])
# !rm -rf ./export ./model.tar.gz
# !aws s3 cp "$model_url" ./model.tar.gz
# !tar xvzf ./model.tar.gz
# 次に、TensorFlow.js で読み込め、推論が実行できる形式に、学習済みモデルデータのフォーマットを変更します。
#
# そために、変換用ツールをインストールし、これを上で解凍したモデルデータに対して適用します。
# 変換用ツールをインストールします
# !pip install tensorflowjs
# +
# 変換したモデルの保存先ディレクトリを作成します
# !rm -rf ./webapp
# !mkdir -p ./webapp/model
# 変換ツールを実行します
# !tensorflowjs_converter \
# --input_format=tf_saved_model \
# --output_node_names='probabilities,classes' \
# --saved_model_tags=serve \
# ./export/Servo/* \
# ./webapp/model
# -
# これで、`./webapp/model`ディレクトリにTensorFlow.jsで読み込めるモデルデータが生成されました!
# !ls -l ./webapp/model/
# このモデルデータを TensorFlow.js で読み込み、推論を実行します。
#
# 以下に、読み込みおよび推論実行コードの主要部をあげます:
#
# ```javascript
# // ライブラリを読み込みます
# import * as tf from '@tensorflow/tfjs-core';
# import {loadFrozenModel} from '@tensorflow/tfjs-converter';
# ...
# // モデルを読み込みます
# const model = await loadFrozenModel(modelUrl, weightsUrl);
# ...
# // モデルで推論を実行し、結果を獲得します
# const output = model.execute({'image_1': input/* 画像データ */}, 'probabilities');
# const probabilities = output.dataSync();
# ```
# 非常に簡単です!
# とはいえ実際には、諸設定を行ったりアプリケーションとしての体裁を整えたりする「ガワ」を作らねばなりません。
# その詳細な説明は、ここでは割愛します。
# その代わり、既に用意されている、モデルデータを簡単に組み込めるWebアプリケーションのテスト用コードに、
# 学習済みのモデルデータを組み込んで実行することにより、データの検証をしてみましょう。
# まず、作成済みのWebアプリケーションのZIPをダウンロードして展開します。
# !wget -O webapp.zip https://github.com/hideya/tfjs-doodle-recognition-pwa/releases/download/0.0.3/nonpwa-webapp.zip
# !unzip webapp.zip -d webapp
# これで、`webapp`ディレクトリ以下にWebアプリケーションに必要なものが全て揃いました!
#
# 確認してみましょう。
# !ls -Rl ./webapp/
# なお、このWebアプリケーションのソースコードは
# [https://github.com/hideya/tfjs-doodle-recognition-pwa/tree/simple](https://github.com/hideya/tfjs-doodle-recognition-pwa/tree/simple)
# にて公開しています(simpleブランチです。masterブランチはsimpleバージョンを [PWA](https://developers.google.com/web/progressive-web-apps/) 化したものになります)。
#
# また、[Vue.js](https://vuejs.org/) を用いたプログラムサンプルも [maru-labo/doodle/examples/tensorflow_js](https://github.com/maru-labo/doodle/tree/master/examples/tensorflow_js) にて公開しています。ぜひ参考にしてください。
# ### ⑤ WebアプリをS3でホスティングして公開する
# `webapp`ディレクトリに必要なものが揃ったので、Web上に公開してみましょう。S3の静的ホスティング機能を使うと簡単にWebアプリケーションを公開できます。`aws s3 sync`コマンドで`webapp`ディレクトリを`public_dir`変数に格納したURLにアップロードします。
public_dir = config['public_dir']
# !aws s3 sync ./webapp $public_dir
# これで必要なファイルが S3 にアップロードされました。
# このアプリを公開し、ブラウザからアクセスできるようにするためには、ファイルが格納された S3 のバケットの「Access」を「Public」にする必要があります(必要なファイルのみをPublicにする方法もあるのですが、今回は簡便のためバケット全体をPublicに設定します)。
# そのためには、まず、下記セルを実行して表示される URL をクリックして、学習結果が格納されたバケットを開きます
print('https://s3.console.aws.amazon.com/s3/buckets/{}'.format(bucket))
# まず「プロパティ Properties」タブを開き、「Static website hosting」を選択し、さらに「このバケットを使用してウェブサイトをホストする Use this bucket to host a website」を選択します。
# 「インデックスドキュメント Index document」には「index.html」と書き込みます。その他は空欄のままでOKです。
# 「保存 Save」をクリックします。
# さらに、もう一つ、設定が必要です。
#
# 「アクセス権限 Permissions」タブを開きます。
# 次に、タブのすぐ下に表示されるメニューから「バケットポリシー Buket Policy」を選択します。
#
# 「バケットポリシーエディター Bucket policy editor」が開いたら、以下のテキストをコピペし、くわえて、「sagemaker-us-west-2-000000000000」の部分を、
# 今対象としているバケット名に合うように書き換えます。
# 「保存 Save」をクリックします。
#
# 「このバケットにはパブリックアクセス権限があります This bucket has public access」と警告が表示され、「バケットポリシー Buket Policy」の下に「パブリック Public」と表示されれば完了です。
# ```
# {
# "Version": "2012-10-17",
# "Statement": [
# {
# "Sid": "PublicReadGetObject",
# "Effect": "Allow",
# "Principal": "*",
# "Action": "s3:GetObject",
# "Resource": "arn:aws:s3:::sagemaker-us-west-2-000000000000/*"
# }
# ]
# }
# ```
# 以上で、S3 バケットの設定は完了です。これで、アップロードされたWebアプリケーションがブラウザからアクセスできるようになりました。
#
# それでは早速、アップロードされたWebアプリケーションの動作を確認してみましょう!
#
# 下記セルを実行するとURLが表示されますので、クリックして開いてみてください。
# 大きなモデルデータを読み込む必要があるため、起動に少々時間がかかります。
# +
print('https://s3-{}.amazonaws.com/{}/index.html'.format(session.boto_region_name, public_dir[5:]))
print('\nもし上のURLで「Error PermanentRedirect」が発生してうまく表示されない場合は、下のURLを試してみてください。')
print('https://s3.amazonaws.com/{}/index.html'.format(public_dir[5:]))
# -
# 本ノートブック冒頭のスクリーンショットのようなアプリが、ちゃんと表示されましたでしょうか?
# 落書きの認識精度はいかがでしょうか?
#
# これで、本ハンズオンは終了です。おつかれさまでした。
#
# 認識率の改善を目指して学習データの数を増やしたり、学習パラメータを変更したり、いろいろ実験してみてください。
# ### 後始末
#
# 実験の後、インスタンスを動いたままにしていたり、巨大なデータを放置していると、思わぬ課金が発生することがあります。
#
# 実験がひととおり終わったら、以下の後始末をすることをおすすめします:
#
# - ノートブックを閉じたら、ノートブック・インスタンスを停止する(これは忘れやすいので気をつけてください)。
# - 実験等で多量に溜まったデータファイルのうち不要なものは、S3 やノートブックインスタンスから削除する。
# - Public になっている S3 の設定を Private にもどす(外部からの意図しないアクセスによるデータ送信の課金を避ける)。
# 課金の状態は、「Billing & Cost Management Dashboard」で簡単に確認することができます。
#
# こちらのURL https://console.aws.amazon.com/billing/home をクリックして、確認してみてください
# (前日までの利用料が表示されますので、今日の分を知りたい場合は、翌日まで待ってからチェックしてみてください)。
# ## まとめ
#
# - ✔ 簡単なデータセットを作りました
# - ✔ SageMakerでモデルを学習しました
# - ✔ SageMakerで学習したモデルをWebアプリケーションで実行しました
#
# なお、落書き認識モデルについては、本日使用したサンプルを含めて全て[GitHub](https://github.com/maru-labo/doodle)上で公開していますので、より詳しい情報をご希望の方はぜひご参照ください。今後、LiteのサンプルやServingの使い方などもリポジトリに追加する予定です。MITライセンスですので、ご自由にご利用いただけます。お気軽にIssueやPull Requestをお寄せくださいませ。
| doodle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nested Statements and Scope
#
# Now that we have gone over writing our own functions, it's important to understand how Python deals with the variable names you assign. When you create a variable name in Python the name is stored in a *name-space*. Variable names also have a *scope*, the scope determines the visibility of that variable name to other parts of your code.
#
# Let's start with a quick thought experiment; imagine the following code:
# +
x = 25
def printer():
x = 50
return x
# print(x)
# print(printer())
# -
# What do you imagine the output of printer() is? 25 or 50? What is the output of print x? 25 or 50?
print(x)
print(printer())
# Interesting! But how does Python know which **x** you're referring to in your code? This is where the idea of scope comes in. Python has a set of rules it follows to decide what variables (such as **x** in this case) you are referencing in your code. Lets break down the rules:
# This idea of scope in your code is very important to understand in order to properly assign and call variable names.
#
# In simple terms, the idea of scope can be described by 3 general rules:
#
# 1. Name assignments will create or change local names by default.
# 2. Name references search (at most) four scopes, these are:
# * local
# * enclosing functions
# * global
# * built-in
# 3. Names declared in global and nonlocal statements map assigned names to enclosing module and function scopes.
#
#
# The statement in #2 above can be defined by the LEGB rule.
#
# **LEGB Rule:**
#
# L: Local — Names assigned in any way within a function (def or lambda), and not declared global in that function.
#
# E: Enclosing function locals — Names in the local scope of any and all enclosing functions (def or lambda), from inner to outer.
#
# G: Global (module) — Names assigned at the top-level of a module file, or declared global in a def within the file.
#
# B: Built-in (Python) — Names preassigned in the built-in names module : open, range, SyntaxError,...
# ## Quick examples of LEGB
#
# ### Local
# x is local here:
f = lambda x:x**2
# ### Enclosing function locals
# This occurs when we have a function inside a function (nested functions)
#
# +
name = 'This is a global name'
def greet():
# Enclosing function
name = 'Sammy'
def hello():
print('Hello '+name)
hello()
greet()
# -
# Note how Sammy was used, because the hello() function was enclosed inside of the greet function!
# ### Global
# Luckily in Jupyter a quick way to test for global variables is to see if another cell recognizes the variable!
print(name)
# ### Built-in
# These are the built-in function names in Python (don't overwrite these!)
len
# ## Local Variables
# When you declare variables inside a function definition, they are not related in any way to other variables with the same names used outside the function - i.e. variable names are local to the function. This is called the scope of the variable. All variables have the scope of the block they are declared in starting from the point of definition of the name.
#
# Example:
# +
x = 50
def func(x):
print('x is', x)
x = 2
print('Changed local x to', x)
func(x)
print('x is still', x)
# -
# The first time that we print the value of the name **x** with the first line in the function’s body, Python uses the value of the parameter declared in the main block, above the function definition.
#
# Next, we assign the value 2 to **x**. The name **x** is local to our function. So, when we change the value of **x** in the function, the **x** defined in the main block remains unaffected.
#
# With the last print statement, we display the value of **x** as defined in the main block, thereby confirming that it is actually unaffected by the local assignment within the previously called function.
#
# ## The <code>global</code> statement
# If you want to assign a value to a name defined at the top level of the program (i.e. not inside any kind of scope such as functions or classes), then you have to tell Python that the name is not local, but it is global. We do this using the <code>global</code> statement. It is impossible to assign a value to a variable defined outside a function without the global statement.
#
# You can use the values of such variables defined outside the function (assuming there is no variable with the same name within the function). However, this is not encouraged and should be avoided since it becomes unclear to the reader of the program as to where that variable’s definition is. Using the <code>global</code> statement makes it amply clear that the variable is defined in an outermost block.
#
# Example:
# +
x = 50
def func():
global x
print('This function is now using the global x!')
print('Because of global x is: ', x)
x = 2
print('Ran func(), changed global x to', x)
print('Before calling func(), x is: ', x)
func()
print('Value of x (outside of func()) is: ', x)
# -
# The <code>global</code> statement is used to declare that **x** is a global variable - hence, when we assign a value to **x** inside the function, that change is reflected when we use the value of **x** in the main block.
#
# You can specify more than one global variable using the same global statement e.g. <code>global x, y, z</code>.
# ## Conclusion
# You should now have a good understanding of Scope (you may have already intuitively felt right about Scope which is great!) One last mention is that you can use the **globals()** and **locals()** functions to check what are your current local and global variables.
#
# Another thing to keep in mind is that everything in Python is an object! I can assign variables to functions just like I can with numbers! We will go over this again in the decorator section of the course!
| Python-Programming/Python-3-Bootcamp/03-Methods and Functions/.ipynb_checkpoints/04-Nested Statements and Scope-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational Inference: Bayesian Neural Networks
#
# (c) 2017 by <NAME>, updated by <NAME>
#
# Original blog post: http://twiecki.github.io/blog/2016/06/01/bayesian-deep-learning/
# ## Current trends in Machine Learning
#
# There are currently three big trends in machine learning: **Probabilistic Programming**, **Deep Learning** and "**Big Data**". Inside of PP, a lot of innovation is in making things scale using **Variational Inference**. In this blog post, I will show how to use **Variational Inference** in PyMC3 to fit a simple Bayesian Neural Network. I will also discuss how bridging Probabilistic Programming and Deep Learning can open up very interesting avenues to explore in future research.
#
# ### Probabilistic Programming at scale
# **Probabilistic Programming** allows very flexible creation of custom probabilistic models and is mainly concerned with **insight** and learning from your data. The approach is inherently **Bayesian** so we can specify **priors** to inform and constrain our models and get uncertainty estimation in form of a **posterior** distribution. Using [MCMC sampling algorithms](http://twiecki.github.io/blog/2015/11/10/mcmc-sampling/) we can draw samples from this posterior to very flexibly estimate these models. PyMC3 and [Stan](http://mc-stan.org/) are the current state-of-the-art tools to consruct and estimate these models. One major drawback of sampling, however, is that it's often very slow, especially for high-dimensional models. That's why more recently, **variational inference** algorithms have been developed that are almost as flexible as MCMC but much faster. Instead of drawing samples from the posterior, these algorithms instead fit a distribution (e.g. normal) to the posterior turning a sampling problem into and optimization problem. [ADVI](http://arxiv.org/abs/1506.03431) -- Automatic Differentation Variational Inference -- is implemented in PyMC3 and [Stan](http://mc-stan.org/), as well as a new package called [Edward](https://github.com/blei-lab/edward/) which is mainly concerned with Variational Inference.
#
# Unfortunately, when it comes to traditional ML problems like classification or (non-linear) regression, Probabilistic Programming often plays second fiddle (in terms of accuracy and scalability) to more algorithmic approaches like [ensemble learning](https://en.wikipedia.org/wiki/Ensemble_learning) (e.g. [random forests](https://en.wikipedia.org/wiki/Random_forest) or [gradient boosted regression trees](https://en.wikipedia.org/wiki/Boosting_(machine_learning)).
#
# ### Deep Learning
#
# Now in its third renaissance, deep learning has been making headlines repeatadly by dominating almost any object recognition benchmark, [kicking ass at Atari games](https://www.cs.toronto.edu/~vmnih/docs/dqn.pdf), and [beating the world-champion Lee Sedol at Go](http://www.nature.com/nature/journal/v529/n7587/full/nature16961.html). From a statistical point, Neural Networks are extremely good non-linear function approximators and representation learners. While mostly known for classification, they have been extended to unsupervised learning with [AutoEncoders](https://arxiv.org/abs/1312.6114) and in all sorts of other interesting ways (e.g. [Recurrent Networks](https://en.wikipedia.org/wiki/Recurrent_neural_network), or [MDNs](http://cbonnett.github.io/MDN_EDWARD_KERAS_TF.html) to estimate multimodal distributions). Why do they work so well? No one really knows as the statistical properties are still not fully understood.
#
# A large part of the innoviation in deep learning is the ability to train these extremely complex models. This rests on several pillars:
# * Speed: facilitating the GPU allowed for much faster processing.
# * Software: frameworks like [Theano](http://deeplearning.net/software/theano/) and [TensorFlow](https://www.tensorflow.org/) allow flexible creation of abstract models that can then be optimized and compiled to CPU or GPU.
# * Learning algorithms: training on sub-sets of the data -- stochastic gradient descent -- allows us to train these models on massive amounts of data. Techniques like drop-out avoid overfitting.
# * Architectural: A lot of innovation comes from changing the input layers, like for convolutional neural nets, or the output layers, like for [MDNs](http://cbonnett.github.io/MDN_EDWARD_KERAS_TF.html).
#
# ### Bridging Deep Learning and Probabilistic Programming
# On one hand we have Probabilistic Programming which allows us to build rather small and focused models in a very principled and well-understood way to gain insight into our data; on the other hand we have deep learning which uses many heuristics to train huge and highly complex models that are amazing at prediction. Recent innovations in variational inference allow probabilistic programming to scale model complexity as well as data size. We are thus at the cusp of being able to combine these two approaches to hopefully unlock new innovations in Machine Learning. For more motivation, see also [<NAME>'s](https://twitter.com/dustinvtran) recent [blog post](http://dustintran.com/blog/a-quick-update-edward-and-some-motivations/).
#
# While this would allow Probabilistic Programming to be applied to a much wider set of interesting problems, I believe this bridging also holds great promise for innovations in Deep Learning. Some ideas are:
# * **Uncertainty in predictions**: As we will see below, the Bayesian Neural Network informs us about the uncertainty in its predictions. I think uncertainty is an underappreciated concept in Machine Learning as it's clearly important for real-world applications. But it could also be useful in training. For example, we could train the model specifically on samples it is most uncertain about.
# * **Uncertainty in representations**: We also get uncertainty estimates of our weights which could inform us about the stability of the learned representations of the network.
# * **Regularization with priors**: Weights are often L2-regularized to avoid overfitting, this very naturally becomes a Gaussian prior for the weight coefficients. We could, however, imagine all kinds of other priors, like spike-and-slab to enforce sparsity (this would be more like using the L1-norm).
# * **Transfer learning with informed priors**: If we wanted to train a network on a new object recognition data set, we could bootstrap the learning by placing informed priors centered around weights retrieved from other pre-trained networks, like [GoogLeNet](https://arxiv.org/abs/1409.4842).
# * **Hierarchical Neural Networks**: A very powerful approach in Probabilistic Programming is hierarchical modeling that allows pooling of things that were learned on sub-groups to the overall population (see my tutorial on [Hierarchical Linear Regression in PyMC3](http://twiecki.github.io/blog/2014/03/17/bayesian-glms-3/)). Applied to Neural Networks, in hierarchical data sets, we could train individual neural nets to specialize on sub-groups while still being informed about representations of the overall population. For example, imagine a network trained to classify car models from pictures of cars. We could train a hierarchical neural network where a sub-neural network is trained to tell apart models from only a single manufacturer. The intuition being that all cars from a certain manufactures share certain similarities so it would make sense to train individual networks that specialize on brands. However, due to the individual networks being connected at a higher layer, they would still share information with the other specialized sub-networks about features that are useful to all brands. Interestingly, different layers of the network could be informed by various levels of the hierarchy -- e.g. early layers that extract visual lines could be identical in all sub-networks while the higher-order representations would be different. The hierarchical model would learn all that from the data.
# * **Other hybrid architectures**: We can more freely build all kinds of neural networks. For example, Bayesian non-parametrics could be used to flexibly adjust the size and shape of the hidden layers to optimally scale the network architecture to the problem at hand during training. Currently, this requires costly hyper-parameter optimization and a lot of tribal knowledge.
# ## Bayesian Neural Networks in PyMC3
# ### Generating data
#
# First, lets generate some toy data -- a simple binary classification problem that's not linearly separable.
# %matplotlib inline
import theano
floatX = theano.config.floatX
import pymc3 as pm
import theano.tensor as T
import sklearn
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from warnings import filterwarnings
filterwarnings('ignore')
sns.set_style('white')
from sklearn import datasets
from sklearn.preprocessing import scale
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_moons
X, Y = make_moons(noise=0.2, random_state=0, n_samples=1000)
X = scale(X)
X = X.astype(floatX)
Y = Y.astype(floatX)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=.5)
fig, ax = plt.subplots()
ax.scatter(X[Y==0, 0], X[Y==0, 1], label='Class 0')
ax.scatter(X[Y==1, 0], X[Y==1, 1], color='r', label='Class 1')
sns.despine(); ax.legend()
ax.set(xlabel='X', ylabel='Y', title='Toy binary classification data set');
# ### Model specification
#
# A neural network is quite simple. The basic unit is a [perceptron](https://en.wikipedia.org/wiki/Perceptron) which is nothing more than [logistic regression](http://pymc-devs.github.io/pymc3/notebooks/posterior_predictive.html#Prediction). We use many of these in parallel and then stack them up to get hidden layers. Here we will use 2 hidden layers with 5 neurons each which is sufficient for such a simple problem.
# +
def construct_nn(ann_input, ann_output):
n_hidden = 5
# Initialize random weights between each layer
init_1 = np.random.randn(X.shape[1], n_hidden).astype(floatX)
init_2 = np.random.randn(n_hidden, n_hidden).astype(floatX)
init_out = np.random.randn(n_hidden).astype(floatX)
with pm.Model() as neural_network:
# Trick: Turn inputs and outputs into shared variables using the data container pm.Data
# It's still the same thing, but we can later change the values of the shared variable
# (to switch in the test-data later) and pymc3 will just use the new data.
# Kind-of like a pointer we can redirect.
# For more info, see: http://deeplearning.net/software/theano/library/compile/shared.html
ann_input = pm.Data('ann_input', X_train)
ann_output = pm.Data('ann_output', Y_train)
# Weights from input to hidden layer
weights_in_1 = pm.Normal('w_in_1', 0, sigma=1,
shape=(X.shape[1], n_hidden),
testval=init_1)
# Weights from 1st to 2nd layer
weights_1_2 = pm.Normal('w_1_2', 0, sigma=1,
shape=(n_hidden, n_hidden),
testval=init_2)
# Weights from hidden layer to output
weights_2_out = pm.Normal('w_2_out', 0, sigma=1,
shape=(n_hidden,),
testval=init_out)
# Build neural-network using tanh activation function
act_1 = pm.math.tanh(pm.math.dot(ann_input,
weights_in_1))
act_2 = pm.math.tanh(pm.math.dot(act_1,
weights_1_2))
act_out = pm.math.sigmoid(pm.math.dot(act_2,
weights_2_out))
# Binary classification -> Bernoulli likelihood
out = pm.Bernoulli('out',
act_out,
observed=ann_output,
total_size=Y_train.shape[0] # IMPORTANT for minibatches
)
return neural_network
neural_network = construct_nn(X_train, Y_train)
# -
# That's not so bad. The `Normal` priors help regularize the weights. Usually we would add a constant `b` to the inputs but I omitted it here to keep the code cleaner.
# ### Variational Inference: Scaling model complexity
#
# We could now just run a MCMC sampler like [NUTS](../api/inference.rst) which works pretty well in this case, but as I already mentioned, this will become very slow as we scale our model up to deeper architectures with more layers.
#
# Instead, we will use the brand-new [ADVI](../api/inference.rst) variational inference algorithm which was recently added to `PyMC3`, and updated to use the operator variational inference (OPVI) framework. This is much faster and will scale better. Note, that this is a mean-field approximation so we ignore correlations in the posterior.
from pymc3.theanof import set_tt_rng, MRG_RandomStreams
set_tt_rng(MRG_RandomStreams(42))
# +
# %%time
with neural_network:
inference = pm.ADVI()
approx = pm.fit(n=30000, method=inference)
# -
# ~ 12 seconds on my laptop. That's pretty good considering that NUTS is having a really hard time. Further below we make this even faster. To make it really fly, we probably want to run the Neural Network on the GPU.
#
# As samples are more convenient to work with, we can very quickly draw samples from the variational approximation using the `sample` method (this is just sampling from Normal distributions, so not at all the same like MCMC):
trace = approx.sample(draws=5000)
# Plotting the objective function (ELBO) we can see that the optimization slowly improves the fit over time.
plt.plot(-inference.hist, label='ADVI', alpha=.3)
plt.legend()
plt.ylabel('ELBO')
plt.xlabel('iteration');
# Now that we trained our model, lets predict on the hold-out set using a posterior predictive check (PPC).
#
# 1. We can use [`sample_posterior_predictive()`](../api/inference.rst) to generate new data (in this case class predictions) from the posterior (sampled from the variational estimation).
# 2. It is better to get the node directly and build theano graph using our approximation (`approx.sample_node`) , we get a lot of speed up
# We can get predicted probability from model
neural_network.out.distribution.p
# +
# create symbolic input
x = T.matrix('X')
# symbolic number of samples is supported, we build vectorized posterior on the fly
n = T.iscalar('n')
# Do not forget test_values or set theano.config.compute_test_value = 'off'
x.tag.test_value = np.empty_like(X_train[:10])
n.tag.test_value = 100
_sample_proba = approx.sample_node(neural_network.out.distribution.p,
size=n,
more_replacements={neural_network['ann_input']: x})
# It is time to compile the function
# No updates are needed for Approximation random generator
# Efficient vectorized form of sampling is used
sample_proba = theano.function([x, n], _sample_proba)
# Create bechmark functions
def production_step1():
pm.set_data(new_data={'ann_input': X_test, 'ann_output': Y_test}, model=neural_network)
ppc = pm.sample_posterior_predictive(trace, samples=500, progressbar=False, model=neural_network)
# Use probability of > 0.5 to assume prediction of class 1
pred = ppc['out'].mean(axis=0) > 0.5
def production_step2():
sample_proba(X_test, 500).mean(0) > 0.5
# -
# See the difference
# %timeit production_step1()
# %timeit production_step2()
# Let's go ahead and generate predictions:
pred = sample_proba(X_test, 500).mean(0) > 0.5
fig, ax = plt.subplots()
ax.scatter(X_test[pred==0, 0], X_test[pred==0, 1])
ax.scatter(X_test[pred==1, 0], X_test[pred==1, 1], color='r')
sns.despine()
ax.set(title='Predicted labels in testing set', xlabel='X', ylabel='Y');
print('Accuracy = {}%'.format((Y_test == pred).mean() * 100))
# Hey, our neural network did all right!
# ## Lets look at what the classifier has learned
#
# For this, we evaluate the class probability predictions on a grid over the whole input space.
grid = pm.floatX(np.mgrid[-3:3:100j,-3:3:100j])
grid_2d = grid.reshape(2, -1).T
dummy_out = np.ones(grid.shape[1], dtype=np.int8)
ppc = sample_proba(grid_2d ,500)
# ### Probability surface
cmap = sns.diverging_palette(250, 12, s=85, l=25, as_cmap=True)
fig, ax = plt.subplots(figsize=(16, 9))
contour = ax.contourf(grid[0], grid[1], ppc.mean(axis=0).reshape(100, 100), cmap=cmap)
ax.scatter(X_test[pred==0, 0], X_test[pred==0, 1])
ax.scatter(X_test[pred==1, 0], X_test[pred==1, 1], color='r')
cbar = plt.colorbar(contour, ax=ax)
_ = ax.set(xlim=(-3, 3), ylim=(-3, 3), xlabel='X', ylabel='Y');
cbar.ax.set_ylabel('Posterior predictive mean probability of class label = 0');
# ### Uncertainty in predicted value
#
# So far, everything I showed we could have done with a non-Bayesian Neural Network. The mean of the posterior predictive for each class-label should be identical to maximum likelihood predicted values. However, we can also look at the standard deviation of the posterior predictive to get a sense for the uncertainty in our predictions. Here is what that looks like:
cmap = sns.cubehelix_palette(light=1, as_cmap=True)
fig, ax = plt.subplots(figsize=(16, 9))
contour = ax.contourf(grid[0], grid[1], ppc.std(axis=0).reshape(100, 100), cmap=cmap)
ax.scatter(X_test[pred==0, 0], X_test[pred==0, 1])
ax.scatter(X_test[pred==1, 0], X_test[pred==1, 1], color='r')
cbar = plt.colorbar(contour, ax=ax)
_ = ax.set(xlim=(-3, 3), ylim=(-3, 3), xlabel='X', ylabel='Y');
cbar.ax.set_ylabel('Uncertainty (posterior predictive standard deviation)');
# We can see that very close to the decision boundary, our uncertainty as to which label to predict is highest. You can imagine that associating predictions with uncertainty is a critical property for many applications like health care. To further maximize accuracy, we might want to train the model primarily on samples from that high-uncertainty region.
# ## Mini-batch ADVI
#
# So far, we have trained our model on all data at once. Obviously this won't scale to something like ImageNet. Moreover, training on mini-batches of data (stochastic gradient descent) avoids local minima and can lead to faster convergence.
#
# Fortunately, ADVI can be run on mini-batches as well. It just requires some setting up:
minibatch_x = pm.Minibatch(X_train, batch_size=50)
minibatch_y = pm.Minibatch(Y_train, batch_size=50)
neural_network_minibatch = construct_nn(minibatch_x, minibatch_y)
with neural_network_minibatch:
approx = pm.fit(40000, method=pm.ADVI())
plt.plot(inference.hist)
plt.ylabel('ELBO')
plt.xlabel('iteration');
# As you can see, mini-batch ADVI's running time is much lower. It also seems to converge faster.
#
# For fun, we can also look at the trace. The point is that we also get uncertainty of our Neural Network weights.
pm.traceplot(trace);
# ## Summary
#
# Hopefully this blog post demonstrated a very powerful new inference algorithm available in PyMC3: [ADVI](http://pymc-devs.github.io/pymc3/api.html#advi). I also think bridging the gap between Probabilistic Programming and Deep Learning can open up many new avenues for innovation in this space, as discussed above. Specifically, a hierarchical neural network sounds pretty bad-ass. These are really exciting times.
#
# ## Next steps
#
# [`Theano`](http://deeplearning.net/software/theano/), which is used by `PyMC3` as its computational backend, was mainly developed for estimating neural networks and there are great libraries like [`Lasagne`](https://github.com/Lasagne/Lasagne) that build on top of `Theano` to make construction of the most common neural network architectures easy. Ideally, we wouldn't have to build the models by hand as I did above, but use the convenient syntax of `Lasagne` to construct the architecture, define our priors, and run ADVI.
#
# You can also run this example on the GPU by setting `device = gpu` and `floatX = float32` in your `.theanorc`.
#
# You might also argue that the above network isn't really deep, but note that we could easily extend it to have more layers, including convolutional ones to train on more challenging data sets.
#
# I also presented some of this work at PyData London, view the video below:
# <iframe width="560" height="315" src="https://www.youtube.com/embed/LlzVlqVzeD8" frameborder="0" allowfullscreen></iframe>
#
# Finally, you can download this NB [here](https://github.com/twiecki/WhileMyMCMCGentlySamples/blob/master/content/downloads/notebooks/bayesian_neural_network.ipynb). Leave a comment below, and [follow me on twitter](https://twitter.com/twiecki).
#
# ## Acknowledgements
#
# [<NAME>](https://github.com/taku-y) did a lot of work on ADVI in PyMC3, including the mini-batch implementation as well as the sampling from the variational posterior. I'd also like to the thank the Stan guys (specifically <NAME> and <NAME>) for deriving ADVI and teaching us about it. Thanks also to <NAME>, <NAME>, <NAME>, and <NAME> for useful comments on an earlier draft.
| docs/source/notebooks/bayesian_neural_network_advi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('linear.csv')
df.head()
df.tipe.unique()
df1 = df[df.tipe == 'data1']
df2 = df[df.tipe == 'data2']
plt.figure()
plt.scatter(df1.X, df1.y, label = 'label1');
plt.scatter(df2.X, df2.y, label = 'label2');
# # Seaborn
sns.scatterplot('X', 'y', data = df, hue = 'tipe');
sns.lmplot('X', 'y', data = df, hue = 'tipe');
sns.distplot(df1.y, bins = 30);
# # Kombinasi seaborn - pandas
# EDA (Exploratory Data Analysis), Statistical Summary
df = pd.read_csv('cereal.csv', index_col = 'name')
df.head()
df.carbo.isna()
sns.heatmap(df.isna())
plt.figure(figsize = (6, 6))
sns.heatmap(df.isna(), cmap = 'Blues', cbar = False);
plt.figure(figsize=(8, 8))
sns.heatmap(df.corr(), cmap = 'bwr', vmin = -1, vmax = 1, annot = True, cbar = False);
# # Pairplot
sns.pairplot(df);
sns.pairplot(df.dropna(), vars = ['calories', 'fat', 'rating']);
# # Plot kategori
# ## Persebaran
sns.catplot('mfr', 'rating', data=df);
sns.boxplot('mfr', 'rating', data=df);
# ## Frequency
sns.catplot('mfr', data=df, kind='count');
sns.countplot('mfr', data=df, hue = 'shelf');
| 03 - Matplotlib Intro/data/Part 02 - Seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Horizontal Current Loop over a Layered Earth (time domain)
from geoscilabs.em.TDEMHorizontalLoopCylWidget import TDEMHorizontalLoopCylWidget
APP = TDEMHorizontalLoopCylWidget()
from matplotlib import rcParams
rcParams['font.size'] = 16
# # Introduction
#
# Here, we show the transient fields and fluxes that result from placing a vertical magnetic dipole (VMD) source over a layered Earth. The transient response in this case refers to the fields and fluxes that are produced once a long-standing primary magnetic field is removed.
#
# There are [two commonly used models](https://em.geosci.xyz/content/maxwell1_fundamentals/dipole_sources_in_homogeneous_media/magnetic_dipole_time/index.html) for describing the VMD source that produces a transient response: 1) as an infinitessimally small bar magnet that experiences a long-standing vertical magnetization which is then instantaneously removed at $t=0$, and 2) as an infinitessimally small horizontal loop of wire carrying a constant current which is then instantaneously shut off at $t=0$ (step-off current waveform).
#
# True dipole sources do not exist in nature however they can be approximated in practice. For geophysical applications, we use small current loops to approximate transient VMD sources. These EM sources may be placed on the Earth's surface (ground-based surveys) or flown through the air (airborne surveys). According to the Biot-Savart law, a primary magnetic field is produced whenever there is current in the loop. When the current is shut-off, the sudden change in magnetic flux induces anomalous currents in the Earth which propagate and diffuse over time. The distribution and propagation of the induced currents depends on the subsurface conductivity distribution and how much time has passed since the current in the VMD source was shut off. The induced currents ultimately produce secondary magnetic fields which can be measured by a receiver.
#
# In this app, we explore the following:
#
# - How do the fields and currents produced by the transient VMD source change over time?
# - For a layered Earth, how does changing layer thickness and conductivity impact the fields and currents produced by the transient VMD source?
# - How do the secondary fields measured above the Earth's surface change over time?
# - For a layered Earth, how does changing layer thickness and conductivity impact secondary fields measured above the Earth's surface?
# # Setup
#
# The geological scenario being modeled is shown in the figure below. Here, we assume the Earth is comprised of 3 layers. Each layer can have a different electrical conductivity ($\sigma$). However, a constant magnetic susceptibility ($\chi$) is used for all layers; where $\mu_0$ is the magnetic permeability of free space and $\mu = \mu_0 (1 +\chi)$. The thicknesses of the top two layers are given by $h_1$ and $h_2$, respectively.
#
# In this case, a transient VMD source (*Tx*) is used to excite the Earth, and the Earth's TEM response (secondary magnetic field) is measured by a receiver (*Rx*). In practice, the transmitter and receiver may be placed near the Earth's surface or in the air. The receiver may also measure secondary fields at a variety of times after the source is shut off.
#
# To understand the fields and currents resulting from a transient VMD source we have two apps:
#
# - **Fields app:** Models the fields and currents everywhere at a particular time after shutoff
# - **Data app:** Models the secondary magnetic field observed by the receiver as a function of off-time
#
#
# <img src="https://github.com/geoscixyz/geosci-labs/blob/main/images/em/LayeredEarthTEM.png?raw=true"></img>
#
# # Exercise
#
# **Follow the exercise in a linear fashion. Some questions may use parameters set in a previous question.**
#
# - **Q1:** Set $\sigma_1$, $\sigma_2$ and $\sigma_3$ to arbitrary conductivity values. Based on the geometry of the problem, which components (x, y, z) of each field (E, B, dBdt or J) are zero? Run the *Fields app* and set *AmpDir = None*. Next, try different combinations of *Field* and *Comp*. Does the simulation match what you initially thought?
#
#
# - **Q2:** Re-run the *Fields app* to set parameters back to default. What happens to the *Ey* and *Jy* as you increase *time index* starting at 1? How does the diffusion and propagation of the EM signal change if the conductivity of all the layers is increased to 1 S/m?
#
#
# - **Q3:** Re-run the *Fields app* to set parameters back to default. Set $\sigma_1 = 0.01$ S/m, $\sigma_2 = 1$ S/m and $\sigma_3 = 0.01$ S/m. Now increase *time index* starting at 1. Is the signal able to effectively penetrate the conductive layer? Why/why not? What if the layer was resistive (i.e. $\sigma_2 = 0.0001$ S/m) instead?
#
#
# - **Q4:** Repeat Q3 but examine the current density. Where is the highest concentration of current density at late time channels? Does this support your answer to Q3?
#
#
# - **Q5:** Re-run the *Fields app* to set parameters back to default. Set *Field = B*, *AmpDir = Direction*. What happens to the magnetic flux density as the *time index* is increased starting at 1? At (x,z)=(0,0), what is the vector direction of the magnetic flux density? Repeat Q5 for dBdt.
#
#
# - **Q6:** Re-run the *Fields app* to set parameters back to default. Set $\sigma_1 = 0.01$ S/m, $\sigma_2 = 1$ S/m and $\sigma_3 = 0.01$ S/m. Examine how B and dBdt are impacted by the conductive layer.
# # Fields app
#
# We use this app to simulate the fields and currents everywhere due to a transient VMD source. The fields and induced currents depend on time and the subsurface conductivity distribution. You will use the app to change various parameters in the model and see how the fields and currents change.
#
# ## Parameters:
#
# - **Update:** If *True* is selected, the simulation is re-run as parameters are being changed. If *False* is selected, the simulation will not be re-fun as parameters are being changed.
# - **Field:** Type of EM fields ("E": electric field, "B": total magnetic flux density, "dBdt": time-derivative of the magnetic flux density, "J": current density and "Model": conductivity model)
# - **AmpDir:** If *None* is selected, then the *x*, *y* or *z* component chosen on the next line is plotted. If *Direction* is chosen, a vector plot is plotted (only possible for B and dB/dt)
# - **Comp.:** If *None* is selected on the previous line, the user chooses whether the *x*, *y* or *z* component is plotted.
# - Time index: The time channel at which fields are being plotted
# - $\boldsymbol{\sigma_0}$: Conductivity of 0th layer in S/m
# - $\boldsymbol{\sigma_1}$: Conductivity of 1st layer in S/m
# - $\boldsymbol{\sigma_2}$: Conductivity of 2nd layer in S/m
# - $\boldsymbol{\sigma_3}$: Conductivity of 3rd layer in S/m
# - $\boldsymbol{\chi}$: Susceptibility of 1-3 layers in SI
# - $\boldsymbol{h_1}$: Thickness of the first layer in metres
# - $\boldsymbol{h_2}$: Thickness of the second layer in metres
# - **Scale:** Plot data values on *log-scale* or *linear-scale*
# - $\boldsymbol{\Delta x}$ (m): Horizontal separation distance between the transmitter and receiver
# - $\boldsymbol{\Delta z}$ (m): Height of the transmitter and receiver above the Earth's surface
# - **Time index:** Time index for the set of frequencies models by this app
APP.InteractivePlane_Layer()
# # Data app
#
# Using this app, we show how the fields observed at the receiver location depend on the parameters set in the previous app. *Note that if you want to see changes in the data due to changes in the model, you MUST* re-run the previous app.
#
# ## Parameters:
#
# - **Field:** Type of EM fields ("E": electric field, "B": magnetic flux density, "dBdt": time-derivative of the magnetic flux density)
# - **Comp.:** Direction of EM field at Rx locations
# - **Scale:** Scale of y-axis values ("log" or "linear")
APP.InteractiveData_Layer()
# # Explore
#
# EM fields will be depenent upon a number of parameters, using a simple half-space model ($\sigma_1=\sigma_2=\sigma_3$) explore how EM fields and data changes upon below four parameters.
#
# - E1: Effects of frequency?
#
#
# - E2: Effects of Tx height?
#
#
# - E3: Effects of Conductivity?
#
#
# - E4: Effects of Susceptibility?
#
| notebooks/em/TDEM_HorizontalLoop_LayeredEarth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf-gpu
# language: python
# name: tf-gpu
# ---
# importing required libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set()
# importing the dataset
dataset = pd.read_csv('Position_Salaries.csv')
dataset.head()
x = dataset.iloc[:, 1:-1].values
x
y = dataset.iloc[:, -1].values
y
# +
# reshaping y
# y = y.reshape(-1,1)
# +
# y
# -
# another way to reshape
y = y.reshape(len(y),1)
y
# visualizing the x and y
plt.figure(figsize=(10,6))
plt.scatter(x,y)
plt.xlabel('Predictor - Years', fontsize=16)
plt.ylabel('Target - Salaries', fontsize=16)
plt.show()
# # Applying Linear Regression
# importing Linear Regression
from sklearn.linear_model import LinearRegression
# Training the model
lm = LinearRegression()
lm.fit(x.reshape(-1,1), y.reshape(-1,1,))
# appling prediction to entire x datapoints, will used these for comparision later
y_pred = lm.predict(x.reshape(-1,1))
# predicting the new value using Linear Regression
y_hat_lm= lm.predict([[6.5]])
print("Predicted Salary for 6.5 years using Linear Model is => ", y_hat_lm)
# # Visualising the Linear Regression results
# Plotting the predicted value with the actual value
plt.figure(figsize=(10,6))
plt.scatter(x, y, s=15)
plt.plot(x, y_pred, color='r')
plt.title('Truth or Bluff (SVR)')
plt.xlabel('Position level (Experience in Years)')
plt.ylabel('Salary')
plt.show()
# # Applying Polynomial Regression
from sklearn.preprocessing import PolynomialFeatures
# applying polynomial degree of 4
poly_reg = PolynomialFeatures(degree=4)
x_poly = poly_reg.fit_transform(x)
lm_2 = LinearRegression()
lm_2.fit(x_poly, y)
# predicting the new value using Polynomial Linear Regression
y_ht_poly = lm_2.predict(poly_reg.fit_transform([[6.5]]))
print("Predicted Salary for 6.5 years using Poly Linear Model is => ",y_ht_poly)
# appling prediction to entire x datapoints, will used these for comparision later
y_poly_pred = lm_2.predict(poly_reg.fit_transform(x.reshape(-1,1)))
y_poly_pred
# # Visualising the Polynomial Regression results
plt.scatter(x, y, color = 'red')
plt.plot(x, y_poly_pred, color = 'blue')
plt.title('Truth or Bluff (Polynomial Regression)')
plt.xlabel('Position level')
plt.ylabel('Salary')
plt.show()
# # Support Vector Regression (SVR)
# <b>SVR required to have all the datapoints in same scale. </b>
#
# <b> So, we have to apply Feature Scaling</b>
# # Feature Scaling
from sklearn.preprocessing import StandardScaler
sc_x = StandardScaler()
sc_y = StandardScaler()
x = sc_x.fit_transform(x)
y = sc_y.fit_transform(y)
print(x)
print(y)
# <b> Training the SVR model on the whole dataset </b>
from sklearn.svm import SVR
lm_svr = SVR(kernel='rbf')
lm_svr.fit(x,y.ravel())
# predicting the new value using SVR Linear Regression
# we need to apply the inverse transform to get the out put into real form
y_ht_svr = sc_y.inverse_transform(lm_svr.predict(sc_x.transform([[6.5]])))
print("Predicted Salary for 6.5 years using SVR Linear Model is => ",y_ht_svr)
# Let see how the SVR regression performed on the non-linear data
# we will use this predicted value for creating the graph for comparison.
y_svr_pred = lm_svr.predict(sc_x.transform(x.reshape(-1,1)))
y_svr_pred
# # SVR Graph
plt.scatter(sc_x.inverse_transform(x), sc_y.inverse_transform(y), color = 'red')
plt.plot(sc_x.inverse_transform(x), sc_y.inverse_transform(lm_svr.predict(x)), color = 'blue')
plt.title('Truth or Bluff (SVR)')
plt.xlabel('Position level (Experience in Years)')
plt.ylabel('Salary')
plt.show()
# # comparing unseen data using three models
print("Predicted Salary for 6.5 years using Linear Model is => ", y_hat_lm)
print("Predicted Salary for 6.5 years using Poly Linear Model is => ",y_ht_poly)
print("Predicted Salary for 6.5 years using SVR Linear Model is => ",y_ht_svr)
# +
y_hat_lm= lm.predict([[7]])
print("Predicted Salary for 7 years using Linear Model is => ", y_hat_lm)
y_ht_poly = lm_2.predict(poly_reg.fit_transform([[7]]))
print("Predicted Salary for 7 years using Poly Linear Model is => ",y_ht_poly)
y_ht_svr = sc_y.inverse_transform(lm_svr.predict(sc_x.transform([[7]])))
print("Predicted Salary for 7 years using SVR Linear Model is => ",y_ht_svr)
# -
# ## As we can see SVR give better result compared to Linear and Poly Linear regressor
| Part 1 - Regression/4. Support Vector Regression (SVR)/.ipynb_checkpoints/Support_Vector_Regression (SVR)_salary_prediction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="_2112Jrm3xF8"
import torch
import random
import numpy as np
random.seed(0)
np.random.seed(0)
torch.manual_seed(0)
torch.cuda.manual_seed(0)
torch.backends.cudnn.deterministic = True
# + colab={"base_uri": "https://localhost:8080/", "height": 352, "referenced_widgets": ["30285d63d2074e04a96ba21c303e977a", "621aa6d451204313a639ea167ba0726e", "548e45ab2e494befbd3edc9b1539dd79", "<KEY>", "60a09efbff444239a18b102aca1f4736", "<KEY>", "fe1f49cbe1024c16877aa4dc2896dccc", "ed354b92d7144515acf6d5ec8a8eed94", "<KEY>", "b00851bc9e9342378c44ef869d637357", "089a419fcee040af802ce81733a337e2", "<KEY>", "5fe8c0c0a06b495c85c75faa769d59da", "<KEY>", "<KEY>", "757c7edbdb184df8a8095eb25e783895", "9823b82ba31d4a17bc65493abc2e9d2b", "<KEY>", "<KEY>", "ad32917ae6f54431a7f76a29ccce1e66", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "326cc5e736e24289ae0ddc83c5c0513d", "<KEY>", "<KEY>", "233940e8d14647ab9fa7e52884dd4ad3", "dbdafe958877430d83ca8962d4f9ae5d", "90b7a3d938a44be980338656c69b69b0", "c49f1dfdfedb42c58779d586ec29a5fb", "589293c79b71491f9b8a2821a65f7971"]} colab_type="code" id="BXBHFv_tT_8R" outputId="f5071926-1d74-44d6-b08d-f6235f4d7ef0"
import torchvision.datasets
MNIST_train = torchvision.datasets.MNIST('./', download=True, train=True)
MNIST_test = torchvision.datasets.MNIST('./', download=True, train=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="OJ2fCCdz325s" outputId="12e288a9-21dc-43a9-ea8a-593d160a3f90"
X_train = MNIST_train.train_data
y_train = MNIST_train.train_labels
X_test = MNIST_test.test_data
y_test = MNIST_test.test_labels
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="uGkDy3UyePNT" outputId="82e2978b-06ed-4281-f0a4-d4b0a1708638"
len(y_train), len(y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="_SAXlRjgeW0e" outputId="310f57b2-3150-4fd8-8676-723f8377c25f"
import matplotlib.pyplot as plt
plt.imshow(X_train[0, :, :])
plt.show()
print(y_train[0])
# + colab={} colab_type="code" id="D8KUxWJmeW_c"
X_train = X_train.unsqueeze(1).float()
X_test = X_test.unsqueeze(1).float()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="yQrlKXireXDy" outputId="091d7d5d-7abd-4582-bb9a-8404b1d7d783"
X_train.shape
# + colab={} colab_type="code" id="-KG_N6hueXJA"
class LeNet5(torch.nn.Module):
def __init__(self,
activation='tanh',
pooling='avg',
conv_size=5,
use_batch_norm=False):
super(LeNet5, self).__init__()
self.conv_size = conv_size
self.use_batch_norm = use_batch_norm
if activation == 'tanh':
activation_function = torch.nn.Tanh()
elif activation == 'relu':
activation_function = torch.nn.ReLU()
else:
raise NotImplementedError
if pooling == 'avg':
pooling_layer = torch.nn.AvgPool2d(kernel_size=2, stride=2)
elif pooling == 'max':
pooling_layer = torch.nn.MaxPool2d(kernel_size=2, stride=2)
else:
raise NotImplementedError
if conv_size == 5:
self.conv1 = torch.nn.Conv2d(
in_channels=1, out_channels=6, kernel_size=5, padding=2)
elif conv_size == 3:
self.conv1_1 = torch.nn.Conv2d(
in_channels=1, out_channels=6, kernel_size=3, padding=1)
self.conv1_2 = torch.nn.Conv2d(
in_channels=6, out_channels=6, kernel_size=3, padding=1)
else:
raise NotImplementedError
self.act1 = activation_function
self.bn1 = torch.nn.BatchNorm2d(num_features=6)
self.pool1 = pooling_layer
if conv_size == 5:
self.conv2 = self.conv2 = torch.nn.Conv2d(
in_channels=6, out_channels=16, kernel_size=5, padding=0)
elif conv_size == 3:
self.conv2_1 = torch.nn.Conv2d(
in_channels=6, out_channels=16, kernel_size=3, padding=0)
self.conv2_2 = torch.nn.Conv2d(
in_channels=16, out_channels=16, kernel_size=3, padding=0)
else:
raise NotImplementedError
self.act2 = activation_function
self.bn2 = torch.nn.BatchNorm2d(num_features=16)
self.pool2 = pooling_layer
self.fc1 = torch.nn.Linear(5 * 5 * 16, 120)
self.act3 = activation_function
self.fc2 = torch.nn.Linear(120, 84)
self.act4 = activation_function
self.fc3 = torch.nn.Linear(84, 10)
def forward(self, x):
if self.conv_size == 5:
x = self.conv1(x)
elif self.conv_size == 3:
x = self.conv1_2(self.conv1_1(x))
x = self.act1(x)
if self.use_batch_norm:
x = self.bn1(x)
x = self.pool1(x)
if self.conv_size == 5:
x = self.conv2(x)
elif self.conv_size == 3:
x = self.conv2_2(self.conv2_1(x))
x = self.act2(x)
if self.use_batch_norm:
x = self.bn2(x)
x = self.pool2(x)
x = x.view(x.size(0), x.size(1) * x.size(2) * x.size(3))
x = self.fc1(x)
x = self.act3(x)
x = self.fc2(x)
x = self.act4(x)
x = self.fc3(x)
return x
# + colab={} colab_type="code" id="XB7nYe8QeXGw"
def train(net, X_train, y_train, X_test, y_test):
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
net = net.to(device)
loss = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(net.parameters(), lr=1.0e-3)
batch_size = 100
test_accuracy_history = []
test_loss_history = []
X_test = X_test.to(device)
y_test = y_test.to(device)
for epoch in range(30):
order = np.random.permutation(len(X_train))
for start_index in range(0, len(X_train), batch_size):
optimizer.zero_grad()
net.train()
batch_indexes = order[start_index:start_index+batch_size]
X_batch = X_train[batch_indexes].to(device)
y_batch = y_train[batch_indexes].to(device)
preds = net.forward(X_batch)
loss_value = loss(preds, y_batch)
loss_value.backward()
optimizer.step()
net.eval()
test_preds = net.forward(X_test)
test_loss_history.append(loss(test_preds, y_test).data.cpu())
accuracy = (test_preds.argmax(dim=1) == y_test).float().mean().data.cpu()
test_accuracy_history.append(accuracy)
print(accuracy)
print('---------------')
return test_accuracy_history, test_loss_history
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="tMNlB5AqeonW" outputId="69215e9f-79f5-4f2a-a690-550025f5719d"
accuracies = {}
losses = {}
accuracies['tanh'], losses['tanh'] = \
train(LeNet5(activation='tanh', conv_size=5),
X_train, y_train, X_test, y_test)
accuracies['relu'], losses['relu'] = \
train(LeNet5(activation='relu', conv_size=5),
X_train, y_train, X_test, y_test)
accuracies['relu_3'], losses['relu_3'] = \
train(LeNet5(activation='relu', conv_size=3),
X_train, y_train, X_test, y_test)
accuracies['relu_3_max_pool'], losses['relu_3_max_pool'] = \
train(LeNet5(activation='relu', conv_size=3, pooling='max'),
X_train, y_train, X_test, y_test)
accuracies['relu_3_max_pool_bn'], losses['relu_3_max_pool_bn'] = \
train(LeNet5(activation='relu', conv_size=3, pooling='max', use_batch_norm=True),
X_train, y_train, X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="zONjrbhnes0x" outputId="c025f42e-9a58-43b5-99d6-dc5000ef3ce4"
for experiment_id in accuracies.keys():
plt.plot(accuracies[experiment_id], label=experiment_id)
plt.legend()
plt.title('Validation Accuracy');
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="lZd4h3fneurR" outputId="6a1d193f-d585-4fe1-d10f-5b6cda817de9"
for experiment_id in losses.keys():
plt.plot(losses[experiment_id], label=experiment_id)
plt.legend()
plt.title('Validation Loss');
# + colab={} colab_type="code" id="Z4U3kF49evTH"
| Mnist_batchnorm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # https://github.com/crflynn/fbm - <NAME>
# """Generate realizations of fractional Brownian motion."""
# import warnings
# import numpy as np
# class FBM(object):
# """The FBM class.
# After instantiating with n = number of increments, hurst parameter, length
# of realization (default = 1) and method of generation
# (default daviesharte), call fbm() for fBm, fgn()
# for fGn, or times() to get corresponding time values.
# """
# def __init__(self, n, hurst, length=1, method="daviesharte"):
# """Instantiate the FBM."""
# self._methods = {"daviesharte": self._daviesharte, "cholesky": self._cholesky, "hosking": self._hosking}
# self.n = n
# self.hurst = hurst
# self.length = length
# self.method = method
# self._fgn = self._methods[self.method]
# # Some reusable values to speed up Monte Carlo.
# self._cov = None
# self._eigenvals = None
# self._C = None
# # Flag if some params get changed
# self._changed = False
# def __str__(self):
# """Str method."""
# return (
# "fBm ("
# + str(self.method)
# + ") on [0, "
# + str(self.length)
# + "] with Hurst value "
# + str(self.hurst)
# + " and "
# + str(self.n)
# + " increments"
# )
# def __repr__(self):
# """Repr method."""
# return (
# "FBM(n="
# + str(self.n)
# + ", hurst="
# + str(self.hurst)
# + ", length="
# + str(self.length)
# + ', method="'
# + str(self.method)
# + '")'
# )
# @property
# def n(self):
# """Get the number of increments."""
# return self._n
# @n.setter
# def n(self, value):
# if not isinstance(value, int) or value <= 0:
# raise TypeError("Number of increments must be a positive int.")
# self._n = value
# self._changed = True
# @property
# def hurst(self):
# """Hurst parameter."""
# return self._hurst
# @hurst.setter
# def hurst(self, value):
# if not isinstance(value, float) or value <= 0 or value >= 1:
# raise ValueError("Hurst parameter must be in interval (0, 1).")
# self._hurst = value
# self._changed = True
# @property
# def length(self):
# """Get the length of process."""
# return self._length
# @length.setter
# def length(self, value):
# if not isinstance(value, (int, float)) or value <= 0:
# raise ValueError("Length of fbm must be greater than 0.")
# self._length = value
# self._changed = True
# @property
# def method(self):
# """Get the algorithm used to generate."""
# return self._method
# @method.setter
# def method(self, value):
# if value not in self._methods:
# raise ValueError("Method must be 'daviesharte', 'hosking' or 'cholesky'.")
# self._method = value
# self._fgn = self._methods[self.method]
# self._changed = True
# def fbm(self):
# """Sample the fractional Brownian motion."""
# return np.insert(self.fgn().cumsum(), [0], 0)
# def fgn(self):
# """Sample the fractional Gaussian noise."""
# scale = (1.0 * self.length / self.n) ** self.hurst
# gn = np.random.normal(0.0, 1.0, self.n)
# # If hurst == 1/2 then just return Gaussian noise
# if self.hurst == 0.5:
# return gn * scale
# else:
# fgn = self._fgn(gn)
# # Scale to interval [0, L]
# return fgn * scale
# def times(self):
# """Get times associated with the fbm/fgn samples."""
# return np.linspace(0, self.length, self.n + 1)
# def _autocovariance(self, k):
# """Autocovariance for fgn."""
# return 0.5 * (abs(k - 1) ** (2 * self.hurst) - 2 * abs(k) ** (2 * self.hurst) + abs(k + 1) ** (2 * self.hurst))
# def _daviesharte(self, gn):
# """Generate a fgn realization using Davies-Harte method.
# Uses Davies and Harte method (exact method) from:
# Davies, <NAME>., and <NAME>. "Tests for Hurst effect."
# Biometrika 74, no. 1 (1987): 95-101.
# Can fail if n is small and hurst close to 1. Falls back to Hosking
# method in that case. See:
# Wood, <NAME>, and <NAME>. "Simulation of stationary Gaussian
# processes in [0, 1] d." Journal of computational and graphical
# statistics 3, no. 4 (1994): 409-432.
# """
# # Monte carlo consideration
# if self._eigenvals is None or self._changed:
# # Generate the first row of the circulant matrix
# row_component = [self._autocovariance(i) for i in range(1, self.n)]
# reverse_component = list(reversed(row_component))
# row = [self._autocovariance(0)] + row_component + [0] + reverse_component
# # Get the eigenvalues of the circulant matrix
# # Discard the imaginary part (should all be zero in theory so
# # imaginary part will be very small)
# self._eigenvals = np.fft.fft(row).real
# self._changed = False
# # If any of the eigenvalues are negative, then the circulant matrix
# # is not positive definite, meaning we cannot use this method. This
# # occurs for situations where n is low and H is close to 1.
# # Fall back to using the Hosking method. See the following for a more
# # detailed explanation:
# #
# # Wood, <NAME>, and <NAME>. "Simulation of stationary Gaussian
# # processes in [0, 1] d." Journal of computational and graphical
# # statistics 3, no. 4 (1994): 409-432.
# if np.any([ev < 0 for ev in self._eigenvals]):
# warnings.warn(
# "Combination of increments n and Hurst value H "
# "invalid for Davies-Harte method. Reverting to Hosking method."
# " Occurs when n is small and Hurst is close to 1. "
# )
# # Set method to hosking for future samples.
# self.method = "hosking"
# # Don"t need to store eigenvals anymore.
# self._eigenvals = None
# return self._hosking(gn)
# # Generate second sequence of i.i.d. standard normals
# gn2 = np.random.normal(0.0, 1.0, self.n)
# # Resulting sequence from matrix multiplication of positive definite
# # sqrt(C) matrix with fgn sample can be simulated in this way.
# w = np.zeros(2 * self.n, dtype=complex)
# for i in range(2 * self.n):
# if i == 0:
# w[i] = np.sqrt(self._eigenvals[i] / (2 * self.n)) * gn[i]
# elif i < self.n:
# w[i] = np.sqrt(self._eigenvals[i] / (4 * self.n)) * (gn[i] + 1j * gn2[i])
# elif i == self.n:
# w[i] = np.sqrt(self._eigenvals[i] / (2 * self.n)) * gn2[0]
# else:
# w[i] = np.sqrt(self._eigenvals[i] / (4 * self.n)) * (gn[2 * self.n - i] - 1j * gn2[2 * self.n - i])
# # Resulting z is fft of sequence w. Discard small imaginary part (z
# # should be real in theory).
# z = np.fft.fft(w)
# fgn = z[: self.n].real
# return fgn
# def _cholesky(self, gn):
# """Generate a fgn realization using the Cholesky method.
# Uses Cholesky decomposition method (exact method) from:
# <NAME>. (1998). Stochastic simulation with a view towards
# stochastic processes. University of Aarhus. Centre for Mathematical
# Physics and Stochastics (MaPhySto)[MPS].
# """
# # Monte carlo consideration
# if self._C is None or self._changed:
# # Generate covariance matrix
# G = np.zeros([self.n, self.n])
# for i in range(self.n):
# for j in range(i + 1):
# G[i, j] = self._autocovariance(i - j)
# # Cholesky decomposition
# self._C = np.linalg.cholesky(G)
# self._changed = False
# # Generate fgn
# fgn = np.dot(self._C, np.array(gn).transpose())
# fgn = np.squeeze(fgn)
# return fgn
# def _hosking(self, gn):
# """Generate a fGn realization using Hosking's method.
# Method of generation is Hosking's method (exact method) from his paper:
# <NAME>. (1984). Modeling persistence in hydrological time series
# using fractional differencing. Water resources research, 20(12),
# 1898-1908.
# """
# fgn = np.zeros(self.n)
# phi = np.zeros(self.n)
# psi = np.zeros(self.n)
# # Monte carlo consideration
# if self._cov is None or self._changed:
# self._cov = np.array([self._autocovariance(i) for i in range(self.n)])
# self._changed = False
# # First increment from stationary distribution
# fgn[0] = gn[0]
# v = 1
# phi[0] = 0
# # Generate fgn realization with n increments of size 1
# for i in range(1, self.n):
# phi[i - 1] = self._cov[i]
# for j in range(i - 1):
# psi[j] = phi[j]
# phi[i - 1] -= psi[j] * self._cov[i - j - 1]
# phi[i - 1] /= v
# for j in range(i - 1):
# phi[j] = psi[j] - phi[i - 1] * psi[i - j - 2]
# v *= 1 - phi[i - 1] * phi[i - 1]
# for j in range(i):
# fgn[i] += phi[j] * fgn[i - j - 1]
# fgn[i] += np.sqrt(v) * gn[i]
# return fgn
# def fbm(n, hurst, length=1, method="daviesharte"):
# """One off sample of fBm."""
# f = FBM(n, hurst, length, method)
# return f.fbm()
# def fgn(n, hurst, length=1, method="daviesharte"):
# """One off sample of fGn."""
# f = FBM(n, hurst, length, method)
# return f.fgn()
# def times(n, length=1):
# """Generate the times associated with increments n and length."""
# return np.linspace(0, length, n + 1)
# +
# # General use
# # Estimate std scaling factor for h=[0.1,0.8] , n=1024
# import numpy as np
# import matplotlib.pyplot as plt
# import scipy.stats as stats
# data_std=[]
# h_arr= np.arange(0.1,0.8,0.05)
# for h in h_arr:
# # generate some random data
# data=[]
# print('h =',h)
# for j in range(1000):
# f = FBM(1024, h)
# fbm_sample = f.fbm()
# data.append(fbm_sample[4]-fbm_sample[3])
# # calc mean
# meanval = np.mean(data)
# print('mean =',meanval)
# # calc standard deviation
# data_cen = data - np.mean(data)
# # or use numpy function
# stdval = np.std(data,ddof=1) # note the second input to provide an unbiased estimate
# print('std =',stdval)
# # calc variance
# varval1 = stdval**2
# # or use numpy function
# varval2 = np.var(data,ddof=1)
# print('varval1 =',varval1)
# print('varval2 =',varval2)
# data_std.append(stdval)
# print('scale correction array h=',h_arr,'=> std_arr =',data_std)
# plt.plot(data_std)
# plt.plot(0.5**(h_arr*10))
# -
# # fBm - 1D
# +
# import matplotlib.pyplot as plt
# x=[]
# y=[]
# for i in range(3):
# h=0.25*(i+1)
# f = FBM(n=1024, hurst=h, length=1, method='daviesharte')
# x.append(f.times())
# y.append(f.fbm())
# plt.subplot(3, 1, 1)
# plt.plot(x[0], y[0], '-')
# plt.title('1D fBm - H=0.25 ; H=0.5 ; H=0.75')
# plt.ylabel('H=0.25')
# plt.subplot(3, 1, 2)
# plt.plot(x[1], y[1], '-')
# plt.xlabel('time (s)')
# plt.ylabel('H=0.5')
# plt.subplot(3, 1, 3)
# plt.plot(x[2], y[2], '-')
# plt.xlabel('time (s)')
# plt.ylabel('H=0.75')
# plt.tight_layout()
# plt.show()
# +
# # fBm simulation below is based on the davies-harte method
# # Davies, <NAME>., and <NAME>. "Tests for Hurst effect." Biometrika 74, no. 1 (1987): 95-101
# import warnings
# import numpy as np
# from fbm import FBM
# from fbm import fbm
# from fbm import times
# import matplotlib.pyplot as plt
# alpha_arr=np.array([1.67,0.8])
# h_arr=0.5*alpha_arr
# x=[]
# y=[]
# for h in h_arr:
# f = FBM(n=1024, hurst=h, length=1, method='daviesharte')
# scale=0.5**(h*10)
# x.append(f.fbm()/scale)
# y.append(f.fbm()/scale)
# fig_min,fig_max=-0.5,0.5
# plt.figure(figsize=(5,10))
# plt.subplot(2, 1, 1)
# plt.plot(x[0], y[0], '-')
# plt.title('2D fBm')
# plt.ylabel('alpha=1.67 ; H=0.835')
# # plt.xlim(fig_min,fig_max)
# # plt.ylim(fig_min,fig_max)
# plt.subplot(2, 1, 2)
# plt.plot(x[1], y[1], '-')
# plt.ylabel('alpha=0.8 ; H=0.4')
# # plt.xlim(fig_min,fig_max)
# # plt.ylim(fig_min,fig_max)
# plt.tight_layout()
# plt.show()
# -
# # MBM - Multifractional Brownian Motion
# # 1D MBM
# ## alpha persistent = 1.67 , alpha antipersistent = 0.8
# ### A self-avoiding walk with neural delays as a model of fxational eye movements <NAME>, <NAME> & <NAME>
# +
# import inspect
# from math import gamma
# import numpy as np
# from mbm import MBM
# alpha_arr=np.array([1.67,0.8])
# trans_time=0.02 # the transition time from persistent to antipersistent motion
# h_arr=alpha_arr/2
# # Hurst function with respect to time.
# def h(t):
# return h_arr[0]+(h_arr[1]-h_arr[0])*(np.tanh(t-trans_time)+1)/2
# m = MBM(n=1024, hurst=h, length=1, method='riemannliouville')
# t_values = m.times()
# x=m.mbm()
# plt.plot(t_values,x)
# plt.show()
# -
# # 2D MBM
#
# ## According to Engbert 2017, Nature:
# ## alpha persistent = 1.67 , alpha antipersistent = 0.8
# ## h = 0.835 , h = 0.4
# +
# import inspect
# from math import gamma
# import numpy as np
# from mbm import MBM
# alpha_arr=np.array([1.67,0.8])
# trans_time=0.02 # the transition time from persistent to antipersistent motion
# h_arr=alpha_arr/2
# # Hurst function with respect to time.
# def h(t):
# return h_arr[0]+(h_arr[1]-h_arr[0])*(np.tanh(t-trans_time)+1)/2
# m = MBM(n=1024, hurst=h, length=1, method='riemannliouville')
# # Get the times associated with the mBm
# t_values = m.times()
# x=[]
# y=[]
# for i in range(3):
# x.append(m.mbm())
# y.append(m.mbm())
# fig_min,fig_max=-0.5,0.5
# plt.figure(figsize=(5,10))
# plt.subplot(3, 1, 1)
# plt.plot(x[0], y[0], '-')
# plt.title('Traj 1')
# plt.ylabel('')
# plt.subplot(3, 1, 2)
# plt.plot(x[1], y[1], '-')
# plt.title('Traj 2')
# plt.ylabel('')
# plt.subplot(3, 1, 3)
# plt.plot(x[2], y[2], '-')
# plt.title('Traj 3')
# plt.ylabel('')
# plt.tight_layout()
# plt.show()
# +
# # test time
# import time
# # fBm simulation below is based on the davies-harte method
# # Davies, <NAME>., and <NAME>. "Tests for Hurst effect." Biometrika 74, no. 1 (1987): 95-101
# import warnings
# import numpy as np
# from fbm import FBM
# from fbm import fbm
# from fbm import times
# import matplotlib.pyplot as plt
# # Function output - x,y coordinates [float] arrays of size n+1
# # h = alpha/2 ; Engbert 2017 alpha values were: alpha=1.67 for 20 ms persistent traj and alpha=0.8 for 100-400ms anti-persistent trajectory
# def ocdr_fbm(bm_steps=1024,h=0.5,ocdr_period_sec=0.5,n_samples=10,fov_len_pix=8,scale_sample_step=1):
# f = FBM(n=bm_steps, hurst=h, length=ocdr_period_sec, method='daviesharte')
# scale_std = (ocdr_period_sec/bm_steps)**h
# scale_bm_step = n_samples/bm_steps
# x,y = scale_sample_step*scale_bm_step*f.fbm()/scale_std, scale_sample_step*scale_bm_step*f.fbm()/scale_std # scale to normal gausian distribution of simulation step size
# max_pos_dis_arr = np.ones(x.shape[0])*fov_len_pix/2
# max_neg_dis_arr = -np.ones(x.shape[0])*fov_len_pix/2
# x = np.minimum(x,max_pos_dis_arr)
# x = np.maximum(x,max_neg_dis_arr)
# y = np.minimum(y,max_pos_dis_arr)
# y = np.maximum(y,max_neg_dis_arr)
# sample_ind=np.arange(0,bm_steps,bm_steps//n_samples)
# # return x,y # return the full trajectory n+1 points
# return x[sample_ind],y[sample_ind] # returns sampled trajectory array of size (n_samples+1)
# # User input:
# h = 0.4 # set 0.1<h<0.9 ; brownian motion: h=0.5 ; persistent motion: h>0.5 ; anti-persistent motion: h<0.5
# bm_steps = 1024 # number of small brownian motion steps
# n_samples = 10 # number of samples from the brownian motion trajectory
# ocdr_period_sec=0.5 # ocular drift period [sec]
# fov_len_pix = 8 # fov_len_pix value corresponds with foveal pixl length and sets the motion max displacment to be +-(fov_len_pix/2)
# scale_sample_step = 1 # scales the brownian motion step size std
# print(t_end-t_start)
# data=[]
# for j in range(1000):
# t_start=time.time()
# # Generate 2D fractional brownian motion trajectory
# x , y = ocdr_fbm(bm_steps,h,ocdr_period_sec,n_samples,max_dis_pix,scale_sample_step)
# t_end=time.time()
# data.append((t_end-t_start)*1000)
# # calc mean
# meanval = np.mean(data)
# print('mean =',meanval)
# # calc standard deviation
# data_cen = data - np.mean(data)
# # or use numpy function
# stdval = np.std(data,ddof=1) # note the second input to provide an unbiased estimate
# print('std =',stdval)
# # calc variance
# varval1 = stdval**2
# # or use numpy function
# varval2 = np.var(data,ddof=1)
# print('varval1 =',varval1)
# print('varval2 =',varval2)
# plt.plot(x,y)
# plt.show()
# -
# # ICLR Ocular drift simulation with fBm
# # Load the FBM class - by <NAME>
#
# https://github.com/crflynn/fbm
#
# MIT License
#
# Copyright (c) 2017-2018 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
# # fBm - 2D
# # Sasha please use h = 0.4 for the 0.5 sec ocular drift period
# +
# Sasha - This the function that you use to produce random fractional brownian motion trajectories
# fBm simulation below is based on the davies-harte method
# Davies, <NAME>., and <NAME>. "Tests for Hurst effect." Biometrika 74, no. 1 (1987): 95-101
import warnings
import numpy as np
from fbm import FBM
from fbm import fbm
from fbm import times
import matplotlib.pyplot as plt
import warnings
import sys
import os
sys.path.insert(1, os.getcwd()+'/fbm_mbm_lic')
import numpy as np
from fbm import FBM
from fbm import fbm
from fbm import times
import matplotlib.pyplot as plt
# Function output - x,y coordinates [float] arrays of size n+1
# h = alpha/2 ; Engbert 2017 alpha values were: alpha=1.67 for 20 ms persistent traj and alpha=0.8 for 100-400ms anti-persistent trajectory
def ocdr_fbm(bm_steps=1024,h=0.5,ocdr_period_sec=0.5,n_samples=10,fov_len_pix=8,scale_sample_step=1):
f = FBM(n=bm_steps, hurst=h, length=ocdr_period_sec, method='daviesharte')
scale_std = (ocdr_period_sec/bm_steps)**h
scale_bm_step = n_samples/bm_steps
x,y = scale_sample_step*scale_bm_step*f.fbm()/scale_std, scale_sample_step*scale_bm_step*f.fbm()/scale_std # scale to normal gausian distribution of simulation step size
max_pos_dis_arr = np.ones(x.shape[0])*fov_len_pix/2
max_neg_dis_arr = -np.ones(x.shape[0])*fov_len_pix/2
x = np.minimum(x,max_pos_dis_arr)
x = np.maximum(x,max_neg_dis_arr)
y = np.minimum(y,max_pos_dis_arr)
y = np.maximum(y,max_neg_dis_arr)
sample_ind=np.arange(0,bm_steps,bm_steps//n_samples)
# return x,y # return the full trajectory n+1 points
return x[sample_ind],y[sample_ind] # returns sampled trajectory array of size (n_samples+1)
# +
# User input:
h = 0.4 # set 0.1<h<0.9 ; brownian motion: h=0.5 ; persistent motion: h>0.5 ; anti-persistent motion: h<0.5
bm_steps = 128 # number of small brownian motion steps
n_samples = 10 # number of samples from the brownian motion trajectory
ocdr_period_sec=0.5 # ocular drift period [sec]
fov_len_pix = 8 # fov_len_pix value corresponds with foveal pixl length and sets the motion max displacment to be +-(fov_len_pix/2)
scale_sample_step = 1 # scales the brownian motion step size std
# Generate 2D fractional brownian motion trajectory
x , y = ocdr_fbm(bm_steps,h,ocdr_period_sec,n_samples,fov_len_pix,scale_sample_step)
# plt.plot(x,y) # uncomment to plot the trjectory
# plt.show()
| .ipynb_checkpoints/iclr_ocular_drift_fbm_sasha_v5-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
tf.config.list_physical_devices('GPU')
import tensorflow_datasets as tfds
print("TensorFlow version:", tf.__version__)
(ds_train, ds_test), ds_info = tfds.load(
'mnist',
split=['train', 'test'],
shuffle_files=True,
as_supervised=True,
with_info=True,
)
def normalize_img(image, label):
"""Normalizes images: `uint8` -> `float32`."""
return tf.cast(image, tf.float32) / 255., label
batch_size = 128
ds_train = ds_train.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_train = ds_train.cache()
ds_train = ds_train.shuffle(ds_info.splits['train'].num_examples)
ds_train = ds_train.batch(batch_size)
ds_train = ds_train.prefetch(tf.data.experimental.AUTOTUNE)
ds_test = ds_test.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_test = ds_test.batch(batch_size)
ds_test = ds_test.cache()
ds_test = ds_test.prefetch(tf.data.experimental.AUTOTUNE)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, kernel_size=(3, 3),
activation='relu'),
tf.keras.layers.Conv2D(64, kernel_size=(3, 3),
activation='relu'),
tf.keras.layers.MaxPooling2D(pool_size=(2, 2)),
# tf.keras.layers.Dropout(0.25),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
# tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(0.001),
metrics=['accuracy'],
)
model.fit(
ds_train,
epochs=12,
validation_data=ds_test,
)
#there are 469 batches of labeled training examples
train_list=list(ds_train)
len(list(train_list))
#there are 128 labeled examples per batch
#each image is 28x28
train_list[0]
#the labels are digits 0-9
train_list[0][1]
#image array corresponding to the first example from the first batch
test_img=np.array(train_list[0][0])[0]; test_img
#we can plot the image
import matplotlib.pyplot as plt
test_img=np.reshape(test_img,(28,28))
plt.imshow(test_img)
#the label for the first example
np.array(train_list[0][1])[0]
#the prediction output values for each digit for the first example
pred=model.predict(train_list[0][0])[0]; pred
np.argmax(pred)
| mnist_benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Created hyperparameter tuning job on the GUI (which now fails), attempted to run tuning job with code below.
#
# ### So far, no examples of full hyperparameter tuning code available, that I have found at least.
#
# #### Documentation from Sagemaker avaiable [here]()
# ## Specify where your training and testing data are located
s3_input_train = 's3://{}/{}/data/train'.format(bucket, prefix)
s3_input_test= 's3://{}/{}/data/test'.format(bucket, prefix)
# ## Specify your training image location (your region)
training_image = containers[region]
# +
HyperParameterTuningJobConfig={
'Strategy': 'Bayesian',
'HyperParameterTuningJobObjective': {
'Type': 'Minimize',
'MetricName': 'test:RMSE' # metric could either be Bayesian or RMSE, unclear.
},
'ResourceLimits': {
'MaxNumberOfTrainingJobs': 50,
'MaxParallelTrainingJobs': 3
},
'ParameterRanges': {
'IntegerParameterRanges': [
{
'Name': 'mini_batch_size',
'MinValue': '32',
'MaxValue': '50'
},
{
'Name': 'epochs',
'MinValue': '32',
'MaxValue': '50'
},
{
'Name': 'context_length',
'MinValue': '20',
'MaxValue': '50'
},
{
'Name': 'num_cells',
'MinValue': '30',
'MaxValue': '70'
},
{
'Name': 'num_layers',
'MinValue': '1',
'MaxValue': '5'
},
],
'ContinuousParameterRanges': [
{
'Name': 'dropout_rate',
'MinValue': '0.00',
'MaxValue': '0.2'
},
{
'Name': 'learning_rate',
'MinValue': '0.0001',
'MaxValue': '0.01'
}
]
}
}
TrainingJobDefinition={
'AlgorithmSpecification': {
'TrainingImage': training_image,
'TrainingInputMode': 'File',
},
'RoleArn': role,
'StaticHyperParameters': [{
"prediction_length": '50',
"time_freq": 'min',
"test_quantiles": '[0.5, 0.9]'
# "_tuning_objective_metric": 'test:RMSE' # can't figure out where this call needs to go.
}],
'InputDataConfig': [
{
'ChannelName': 'train',
'DataSource': {
'S3DataSource': {
'S3DataType': 'S3Prefix',
'S3Uri': s3_input_train,
'S3DataDistributionType': 'FullyReplicated'
}
},
'ContentType': 'json',
'CompressionType': 'None',
'RecordWrapperType': 'None'
,
'ChannelName': 'test',
'DataSource': {
'S3DataSource': {
'S3DataType': 'S3Prefix',
'S3Uri': s3_input_test,
'S3DataDistributionType': 'FullyReplicated'
}
},
'ContentType': 'json',
'CompressionType': 'None',
'RecordWrapperType': 'None'
}
],
'OutputDataConfig': {
"S3OutputPath": "s3://{}/{}/output".format(bucket,prefix)
},
'ResourceConfig': {
'InstanceType': 'ml.m4.xlarge', # depending on size of data and number of TS, might need larger or smaller
'InstanceCount': 1, # You can change this limit, I had to request a limit
'VolumeSizeInGB': 1,
},
'StoppingCondition': {
'MaxRuntimeInSeconds': 86400
}
}
# -
# ## Run tuning job
smclient.create_hyper_parameter_tuning_job(HyperParameterTuningJobName = 'TUNING-JOB-NAME',
HyperParameterTuningJobConfig = HyperParameterTuningJobConfig,
TrainingJobDefinition = TrainingJobDefinition1)
# ## Above command will run tuning job where you can access the best job parameters on the GUI or call them here to use for model training.
| HyperparameterTuning_DeepAR_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import numpy as np
# +
def calc_score_diff(play):
"""Calculate the score differential of the team with possession.
Parameters:
play(Dict): The play object.
Returns:
int: The score differential of the team with possession.
"""
return int(play['posteam_score']) - int(play['defteam_score'])
def get_yrdln_int(play):
"""Given a play, get the line of scrimmage as an integer.
Parameters:
play(Dict): The play object.
Returns:
int: The yard line as an integer.
"""
return int(play['yrdln'].split(' ')[-1])
def calc_seconds_since_halftime(play, year, is_postseason):
"""Calculate the number of seconds elapsed since halftime.
Parameters:
play(Dict): The play object.
year(int): The year (season) of the game.
is_postseason(bool): If the game is a postseason game.
Returns:
int: The number of seconds elapsed since halftime of that play.
"""
if int(play['qtr']) <= 4:
if play['game_seconds_remaining'] == '1e3':
return 1800 - 1000
return max(0, 1800-int(play['game_seconds_remaining']))
# Check if game before 2017 or is postseason (when overtime rules change)
if (year < 2017 or is_postseason):
seconds_per_overtime = 900
else:
seconds_per_overtime = 600
# Handle overtime
if int(play['qtr']) == 5:
return 1800 + seconds_per_overtime - int(play['game_seconds_remaining'])
elif int(play['qtr']) == 6:
return 3600 - int(play['game_seconds_remaining'])
elif int(play['qtr']) == 7:
return 4500 - int(play['game_seconds_remaining'])
# Default
return 0
def calc_field_pos_score(play):
"""Calculate the field position score for a play.
Parameters:
play(Dict): The play object.
Returns:
float: The "field position score" for a given play,
used to calculate the surrender index.
"""
try:
if '50' in play['yrdln']:
return (1.1) ** 10.
if play['posteam'] in play['yrdln']:
return max(1., (1.1)**(get_yrdln_int(play) - 40))
else:
return (1.2)**(50 - get_yrdln_int(play)) * ((1.1)**(10))
except BaseException:
return 0.
def calc_yds_to_go_multiplier(play):
"""Calculate the yards to go multiplier for a play.
Parameters:
play(Dict): The play object.
Returns:
float: The "yards to go multiplier" for a given play,
used to calculate the surrender index.
"""
if int(play['ydstogo']) >= 10:
return 0.2
elif int(play['ydstogo']) >= 7:
return 0.4
elif int(play['ydstogo']) >= 4:
return 0.6
elif int(play['ydstogo']) >= 2:
return 0.8
else:
return 1.
def calc_score_multiplier(play):
"""Calculate the score multiplier for a play.
Parameters:
play(Dict): The play object.
Returns:
float: The "score multiplier" for a given play,
used to calculate the surrender index.
"""
score_diff = calc_score_diff(play)
if score_diff > 0:
return 1.
elif score_diff == 0:
return 2.
elif score_diff < -8.:
return 3.
else:
return 4.
def calc_clock_multiplier(play, year, is_postseason):
"""Calculate the clock multiplier for a play.
Parameters:
play(Dict): The play object.
year(int): The year (season) of the game.
is_postseason(bool): If the game is a postseason game.
Returns:
float: The "clock multiplier" for a given play,
used to calculate the surrender index.
"""
if calc_score_diff(play) <= 0 and int(play['qtr']) > 2:
seconds_since_halftime = calc_seconds_since_halftime(play, year, is_postseason)
return ((seconds_since_halftime * 0.001) ** 3.) + 1.
else:
return 1.
def calc_surrender_index(play, year, is_postseason):
"""Calculate the surrender index for a play.
Parameters:
play(Dict): The play object.
year(int): The year (season) of the game.
is_postseason(bool): If the game is a postseason game.
Returns:
float: The surrender index for a given play.
"""
return calc_field_pos_score(play) * calc_yds_to_go_multiplier(
play) * calc_score_multiplier(play) * calc_clock_multiplier(play, year, is_postseason)
# +
surrender_indices = []
for year in range(1999, 2021):
with open('nflfastR-data/play_by_play_' + str(year) + '.csv', 'r') as f:
print(year)
data = csv.DictReader(f)
for play in data:
if play['play_type'] == 'punt':
surrender_indices.append(calc_surrender_index(play, year, play['season_type'] == 'POST'))
# -
len(surrender_indices)
np_surrender_indices = np.array(surrender_indices)
np.save('1999-2020_surrender_indices.npy', np_surrender_indices)
| Historical Surrender Indices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Generators cannot be pickled, so this should become an exploding variable.
x=(i*i for i in range(10))
# Large array of >50MB in size should become an exploding variable.
y=list(range(2000000))
| test/testdata/features/exploding_variables/exploding_variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
assert sys.version_info.major == 3
assert sys.version_info.minor == 7
import numpy
assert numpy.version.version == "1.19.5"
import matplotlib
assert matplotlib.__version__ == "3.2.2"
import scipy
assert scipy.__version__ == "1.4.1"
import Cython
assert Cython.__version__ == "0.29.24"
import sympy
assert sympy.__version__ == "1.7.1"
| colab/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="width:1000 px">
#
# <div style="float:right; width:98 px; height:98px;">
# <img src="https://raw.githubusercontent.com/Unidata/MetPy/master/metpy/plots/_static/unidata_150x150.png" alt="Unidata Logo" style="height: 98px;">
# </div>
#
# <h1>Using Siphon to query the NetCDF Subset Service</h1>
# <h3>Unidata Python Workshop</h3>
#
# <div style="clear:both"></div>
# </div>
#
# <hr style="height:2px;">
#
# ### Objectives
# 1. Learn what Siphon is
# 2. Employ Siphon's NCSS class to retrieve data from a THREDDS Data Server (TDS)
# 3. Plot a map using numpy arrays, matplotlib, and cartopy!
#
# ### Introduction:
# Siphon is a python package that makes downloading data from Unidata data technologies a breeze! In our examples, we'll focus on interacting with the netCDF Subset Service (NCSS) as well as the radar server to retrieve grid data and radar data.
#
# **But first!**
# Bookmark these resources for when you want to use Siphon later!
# + [latest Siphon documentation](http://siphon.readthedocs.org/en/latest/)
# + [Siphon github repo](https://github.com/Unidata/siphon)
# + [TDS documentation](http://www.unidata.ucar.edu/software/thredds/v4.6/tds/TDS.html)
# + [netCDF subset service documentation](http://www.unidata.ucar.edu/software/thredds/current/tds/reference/NetcdfSubsetServiceReference.html)
# ### Let's get started!
# First, we'll import the TDSCatalog class from Siphon and put the special 'matplotlib' line in so our map will show up later in the notebook. Let's construct an instance of TDSCatalog pointing to our dataset of interest. In this case, I've chosen the TDS' "Best" virtual dataset for the GFS global 0.25 degree collection of GRIB files. This will give us a good resolution for our map. This catalog contains a single dataset.
# %matplotlib inline
from siphon.catalog import TDSCatalog
best_gfs = TDSCatalog('http://thredds.ucar.edu/thredds/catalog/grib/NCEP/GFS/'
'Global_0p25deg/catalog.xml?dataset=grib/NCEP/GFS/Global_0p25deg/Best')
best_gfs.datasets
# We pull out this dataset and call `subset()` to set up requesting a subset of the data.
best_ds = list(best_gfs.datasets.values())[0]
ncss = best_ds.subset()
# We can then use the `ncss` object to create a new query object, which
# facilitates asking for data from the server.
query = ncss.query()
# We can look at the `ncss.variables` object to see what variables are available from the dataset:
ncss.variables
# We construct a query asking for data corresponding to a latitude and longitude box where 43 lat is the northern extent, 35 lat is the southern extent, 260 long is the western extent and 249 is the eastern extent. Note that longitude values are the longitude distance from the prime meridian. We request the data for the current time. This request will return all surface temperatures for points in our bounding box for a single time. Note the string representation of the query is a properly encoded query string.
from datetime import datetime
query.lonlat_box(north=43, south=35, east=260, west=249).time(datetime.utcnow())
query.accept('netcdf4')
query.variables('Temperature_surface')
# We now request data from the server using this query. The `NCSS` class handles parsing this NetCDF data (using the `netCDF4` module). If we print out the variable names, we see our requested variables, as well as a few others (more metadata information)
# +
from xarray.backends import NetCDF4DataStore
import xarray as xr
data = ncss.get_data(query)
data = xr.open_dataset(NetCDF4DataStore(data))
list(data)
# -
# We'll pull out the temperature variable.
temp_3d = data['Temperature_surface']
# We'll pull out the useful variables for latitude, and longitude, and time (which is the time, in hours since the forecast run). Notice the variable names are labeled to show how many dimensions each variable is. This will come in to play soon when we prepare to plot. Try printing one of the variables to see some info on the data!
# Helper function for finding proper time variable
def find_time_var(var, time_basename='time'):
for coord_name in var.coords:
if coord_name.startswith(time_basename):
return var.coords[coord_name]
raise ValueError('No time variable found for ' + var.name)
time_1d = find_time_var(temp_3d)
lat_1d = data['lat']
lon_1d = data['lon']
time_1d
# Now we make our data suitable for plotting. We'll import numpy so we can combine lat/longs (meshgrid) and remove one-dimensional entities from our arrays (squeeze). Also we'll use netCDF4's num2date to change the time since the model run to an actual date.
# +
import numpy as np
from netCDF4 import num2date
from metpy.units import units
# Reduce the dimensions of the data and get as an array with units
temp_2d = temp_3d.metpy.unit_array.squeeze()
# Combine latitude and longitudes
lon_2d, lat_2d = np.meshgrid(lon_1d, lat_1d)
# -
# Now we can plot these up using matplotlib. We import cartopy and matplotlib classes, create our figure, add a map, then add the temperature data and grid points.
# +
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import cartopy.feature as cfeature
from metpy.plots import ctables
# Create a new figure
fig = plt.figure(figsize=(15, 12))
# Add the map and set the extent
ax = plt.axes(projection=ccrs.PlateCarree())
ax.set_extent([-100.03, -111.03, 35, 43])
# Retrieve the state boundaries using cFeature and add to plot
ax.add_feature(cfeature.STATES, edgecolor='gray')
# Contour temperature at each lat/long
contours = ax.contourf(lon_2d, lat_2d, temp_2d.to('degF'), 200, transform=ccrs.PlateCarree(),
cmap='RdBu_r')
#Plot a colorbar to show temperature and reduce the size of it
fig.colorbar(contours)
# Make a title with the time value
ax.set_title(f'Temperature forecast (\u00b0F) for {time_1d[0].values}Z', fontsize=20)
# Plot markers for each lat/long to show grid points for 0.25 deg GFS
ax.plot(lon_2d.flatten(), lat_2d.flatten(), linestyle='none', marker='o',
color='black', markersize=2, alpha=0.3, transform=ccrs.PlateCarree());
# -
# ### Exercise
# Create your own map using the same projection as above but plot different data variables such as dewpoint or relative humidity.
# 1. Explore the variables available in the NCSS dataset by printing NCSS.variables
# 2. Change the latitude/longitude values for the request and the map to a region of your own interest!
# 3. If you're feeling bold, pass in a different TDSCatalog reference url (such as the GFS half degree). Take a look at the full TDS catalog [here](http://thredds.ucar.edu/thredds/catalog.html).
| notebooks/Bonus/Downloading GFS with Siphon.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # minesweeper
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/minesweeper.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/minesweeper.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Minesweeper in Google CP Solver.
From gecode/examples/minesweeper.cc:
'''
A specification is a square matrix of characters. Alphanumeric
characters represent the number of mines adjacent to that field.
Dots represent fields with an unknown number of mines adjacent to
it (or an actual mine).
'''
E.g.
'..2.3.'
'2.....'
'..24.3'
'1.34..'
'.....3'
'.3.3..'
Also see:
* http://www.janko.at/Raetsel/Minesweeper/index.htm
* http://en.wikipedia.org/wiki/Minesweeper_(computer_game)
* <NAME> on Minesweeper:
http://www.claymath.org/Popular_Lectures/Minesweeper/
* Richard Kaye's Minesweeper Pages
http://web.mat.bham.ac.uk/R.W.Kaye/minesw/minesw.htm
* Some Minesweeper Configurations
http://web.mat.bham.ac.uk/R.W.Kaye/minesw/minesw.pdf
Compare with the following models:
* MiniZinc: http://www.hakank.org/minizinc/minesweeper.mzn
* Choco : http://www.hakank.org/choco/MineSweeper.java
* JaCoP : http://www.hakank.org/JaCoP/MineSweeper.java
* Gecode/R: http://www.hakank.org/gecode_r/minesweeper.rb
* Comet : http://www.hakank.org/comet/minesweeper.co
* ECLiPSe : http://www.hakank.org/eclipse/minesweeper.ecl
* SICStus : http://www.hakank.org/sicstus/minesweeper.pl
* Tailor/Essence': http://www.hakank.org/tailor/minesweeper.eprime
* Zinc: http://www.hakank.org/minizinc/minesweeper.zinc
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
import sys
from ortools.constraint_solver import pywrapcp
default_r = 8
default_c = 8
X = -1
default_game = [[2, 3, X, 2, 2, X, 2, 1], [X, X, 4, X, X, 4, X, 2],
[X, X, X, X, X, X, 4, X], [X, 5, X, 6, X, X, X, 2],
[2, X, X, X, 5, 5, X, 2], [1, 3, 4, X, X, X, 4, X],
[0, 1, X, 4, X, X, X, 3], [0, 1, 2, X, 2, 3, X, 2]]
# Create the solver.
solver = pywrapcp.Solver("Minesweeper")
#
# data
#
# Set default problem
if game == "":
game = default_game
r = default_r
c = default_c
else:
print("rows:", r, " cols:", c)
#
# Default problem from "Some Minesweeper Configurations",page 3
# (same as problem instance minesweeper_config3.txt)
# It has 4 solutions
#
# r = 8
# c = 8
# X = -1
# game = [
# [2,3,X,2,2,X,2,1],
# [X,X,4,X,X,4,X,2],
# [X,X,X,X,X,X,4,X],
# [X,5,X,6,X,X,X,2],
# [2,X,X,X,5,5,X,2],
# [1,3,4,X,X,X,4,X],
# [0,1,X,4,X,X,X,3],
# [0,1,2,X,2,3,X,2]
# ]
S = [-1, 0, 1] # for the neighbors of "this" cell
# print problem instance
print("Problem:")
for i in range(r):
for j in range(c):
if game[i][j] == X:
print("X", end=" ")
else:
print(game[i][j], end=" ")
print()
print()
# declare variables
mines = {}
for i in range(r):
for j in range(c):
mines[(i, j)] = solver.IntVar(0, 1, "mines %i %i" % (i, j))
#
# constraints
#
for i in range(r):
for j in range(c):
if game[i][j] >= 0:
solver.Add(mines[i, j] == 0)
# this cell is the sum of all the surrounding cells
solver.Add(game[i][j] == solver.Sum([
mines[i + a, j + b]
for a in S
for b in S
if i + a >= 0 and j + b >= 0 and i + a < r and j + b < c
]))
if game[i][j] > X:
# This cell cannot be a mine
solver.Add(mines[i, j] == 0)
#
# solution and search
#
solution = solver.Assignment()
solution.Add([mines[(i, j)] for i in range(r) for j in range(c)])
collector = solver.AllSolutionCollector(solution)
solver.Solve(
solver.Phase([mines[(i, j)] for i in range(r) for j in range(c)],
solver.INT_VAR_SIMPLE, solver.ASSIGN_MIN_VALUE), [collector])
num_solutions = collector.SolutionCount()
print("num_solutions: ", num_solutions)
if num_solutions > 0:
for s in range(num_solutions):
minesval = [
collector.Value(s, mines[(i, j)]) for i in range(r) for j in range(c)
]
for i in range(r):
for j in range(c):
print(minesval[i * c + j], end=" ")
print()
print()
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
else:
print("No solutions found")
#
# Read a problem instance from a file
#def read_problem(file):
f = open(file, "r")
rows = int(f.readline())
cols = int(f.readline())
game = []
for i in range(rows):
x = f.readline()
row = [0] * cols
for j in range(cols):
if x[j] == ".":
tmp = -1
else:
tmp = int(x[j])
row[j] = tmp
game.append(row)
return [game, rows, cols]
#
# Print the mines
#
def print_mines(mines, rows, cols):
for i in range(rows):
for j in range(cols):
print(mines[i, j], end=" ")
print("")
def print_game(game, rows, cols):
for i in range(rows):
for j in range(cols):
print(game[i][j], end=" ")
print("")
| examples/notebook/contrib/minesweeper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## README
# O cartpy é uma ferramenta para facilitar o acesso às bases cartográficas do IBGE e às estatísticas históricas dos territórios dos municípios brasileiros. Abaixo seguem instruções de instalação e uso
#
# ### 1- Instalação
#
# O pacote está disponível no índice de pacotes do python ([PyPi](https://pypi.org/)) e, assim, pode facilmente ser instalado usando:<br>
# ```python
# pip install cartpy
# ```
#
# ### 2- Uso
#
# O cartpy possui duas classes, Year e Municipio, vamos apresentar o uso básico do pacote abordando cada classe individualmente.
#
# #### 2.1 -Year
#
# Essa classe serve basicamente para dar acesso direto às bases do IBGE. Assim, digamos que estejamos interessados em baixar os dados do ano de 1872, data do primeiro censo brasileiro. Para isso, basta fazer:
import cartpy
data1872=cartpy.Year(1872)
# Com método get_geodata temos acesso à base em formato de dataframe do geopandas:
sf=data1872.get_geodata()
sf.plot()
# Se o usuário estiver interessado em apenas um estado, basta especificar o código ou a sigla:
data1872.get_geodata(state='BA').plot()
data1872.get_geodata(state=35).plot()
# Também é possível ter acesso à base de municípios específicos, caso em que também é preciso especificar o estado. Novamente é possível usa o código ou o nome da unidade federativa em questão
data1872.get_geodata(state='MG',county='Juiz De Fora').plot()
data1872.get_geodata(state='SP',county=3526902).plot() #Limeira
# Qualquer das bases pode ser salva como um shapefile usando:<br>
# ```python
# sf.to_file('filename.shp')
# ```
# #### 2.2 - Municipio
#
# A classe Municipio() oferece uma forma simplificada de ter acesso a diversos dados cartográficos dos municípios brasileiros em perspectiva histórica. <br>
# Essa classe aceita qualquer string quando é instanciada, mas só aceitará a aplicação de seus respectivos métodos se a string usada corresponder ao município. Para evitar frustrações, é interessante usar o método search para verificar qual a ortografia da base:
mun=cartpy.Municipio('Barreiro')
mun.search(year=1991)
# Agora instanciamos novamente a classe, corrigindo a ortografia para ficar igual a base:
mun=cartpy.Municipio('Sao Jose Do Barreiro')
# Na versão atual, a classe Municipio tem os seguintes métodos:
mun.get_code(state='SP',year=1991)
mun.all_names(code=3549607)
mun.get_map(state='SP',year=1991)
# Essa é apenas a primeira versão do pacote, outras funcionalidades serão acrescentadas ao longo do tempo por mim e/ou pela comunidade de usuários
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
np.random.seed(1789)
from IPython.core.display import HTML
def css_styling():
styles = open("styles/custom.css", "r").read()
return HTML(styles)
css_styling()
# -
# # Statistical Data Modeling
#
# Pandas, NumPy and SciPy provide the core functionality for building statistical models of our data. We use models to:
#
# - Concisely **describe** the components of our data
# - Provide **inference** about underlying parameters that may have generated the data
# - Make **predictions** about unobserved data, or expected future observations.
#
# This section of the tutorial illustrates how to use Python to build statistical models of low to moderate difficulty from scratch, and use them to extract estimates and associated measures of uncertainty.
# Estimation
# ==========
#
# An recurring statistical problem is finding estimates of the relevant parameters that correspond to the distribution that best represents our data.
#
# In **parametric** inference, we specify *a priori* a suitable distribution, then choose the parameters that best fit the data.
#
# * e.g. the mean $\mu$ and the variance $\sigma^2$ in the case of the normal distribution
x = np.array([ 1.00201077, 1.58251956, 0.94515919, 6.48778002, 1.47764604,
5.18847071, 4.21988095, 2.85971522, 3.40044437, 3.74907745,
1.18065796, 3.74748775, 3.27328568, 3.19374927, 8.0726155 ,
0.90326139, 2.34460034, 2.14199217, 3.27446744, 3.58872357,
1.20611533, 2.16594393, 5.56610242, 4.66479977, 2.3573932 ])
_ = plt.hist(x, bins=7)
# ### Fitting data to probability distributions
#
# We start with the problem of finding values for the parameters that provide the best fit between the model and the data, called point estimates. First, we need to define what we mean by ‘best fit’. There are two commonly used criteria:
#
# * **Method of moments** chooses the parameters so that the sample moments (typically the sample mean and variance) match the theoretical moments of our chosen distribution.
# * **Maximum likelihood** chooses the parameters to maximize the likelihood, which measures how likely it is to observe our given sample.
# ### Discrete Random Variables
#
# $$X = \{0,1\}$$
#
# $$Y = \{\ldots,-2,-1,0,1,2,\ldots\}$$
#
# **Probability Mass Function**:
#
# For discrete $X$,
#
# $$Pr(X=x) = f(x|\theta)$$
#
# 
# ***e.g. Poisson distribution***
#
# The Poisson distribution models unbounded counts:
#
# <div style="font-size: 150%;">
# $$Pr(X=x)=\frac{e^{-\lambda}\lambda^x}{x!}$$
# </div>
#
# * $X=\{0,1,2,\ldots\}$
# * $\lambda > 0$
#
# $$E(X) = \text{Var}(X) = \lambda$$
# ### Continuous Random Variables
#
# $$X \in [0,1]$$
#
# $$Y \in (-\infty, \infty)$$
#
# **Probability Density Function**:
#
# For continuous $X$,
#
# $$Pr(x \le X \le x + dx) = f(x|\theta)dx \, \text{ as } \, dx \rightarrow 0$$
#
# 
# ***e.g. normal distribution***
#
# <div style="font-size: 150%;">
# $$f(x) = \frac{1}{\sqrt{2\pi\sigma^2}}\exp\left[-\frac{(x-\mu)^2}{2\sigma^2}\right]$$
# </div>
#
# * $X \in \mathbf{R}$
# * $\mu \in \mathbf{R}$
# * $\sigma>0$
#
# $$\begin{align}E(X) &= \mu \cr
# \text{Var}(X) &= \sigma^2 \end{align}$$
# ### Example: Nashville Precipitation
#
# The dataset `nashville_precip.txt` contains [NOAA precipitation data for Nashville measured since 1871](http://bit.ly/nasvhville_precip_data).
#
# 
#
# The gamma distribution is often a good fit to aggregated rainfall data, and will be our candidate distribution in this case.
precip = pd.read_table("../data/nashville_precip.txt", index_col=0, na_values='NA', delim_whitespace=True)
precip.head()
_ = precip.hist(sharex=True, sharey=True, grid=False)
plt.tight_layout()
# The first step is recognizing what sort of distribution to fit our data to. A couple of observations:
#
# 1. The data are skewed, with a longer tail to the right than to the left
# 2. The data are positive-valued, since they are measuring rainfall
# 3. The data are continuous
#
# There are a few possible choices, but one suitable alternative is the **gamma distribution**:
#
# <div style="font-size: 150%;">
# $$x \sim \text{Gamma}(\alpha, \beta) = \frac{\beta^{\alpha}x^{\alpha-1}e^{-\beta x}}{\Gamma(\alpha)}$$
# </div>
#
# 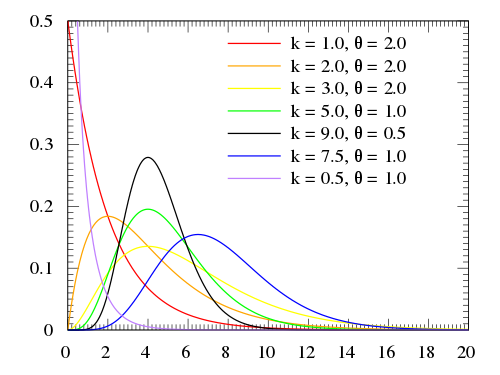
# The ***method of moments*** simply assigns the empirical mean and variance to their theoretical counterparts, so that we can solve for the parameters.
#
# So, for the gamma distribution, the mean and variance are:
#
# <div style="font-size: 150%;">
# $$ \hat{\mu} = \bar{X} = \alpha \beta $$
# $$ \hat{\sigma}^2 = S^2 = \alpha \beta^2 $$
# </div>
# So, if we solve for these parameters, we can use a gamma distribution to describe our data:
#
# <div style="font-size: 150%;">
# $$ \alpha = \frac{\bar{X}^2}{S^2}, \, \beta = \frac{S^2}{\bar{X}} $$
# </div>
# Let's deal with the missing value in the October data. Given what we are trying to do, it is most sensible to fill in the missing value with the average of the available values. We will learn more sophisticated methods for handling missing data later in the course.
precip.fillna(value={'Oct': precip.Oct.mean()}, inplace=True)
# Now, let's calculate the sample moments of interest, the means and variances by month:
precip_mean = precip.mean()
precip_mean
precip_var = precip.var()
precip_var
# We then use these moments to estimate $\alpha$ and $\beta$ for each month:
alpha_mom = precip_mean ** 2 / precip_var
beta_mom = precip_var / precip_mean
alpha_mom, beta_mom
# We can use the `gamma.pdf` function in `scipy.stats.distributions` to plot the ditribtuions implied by the calculated alphas and betas. For example, here is January:
# +
from scipy.stats.distributions import gamma
precip.Jan.hist(normed=True, bins=20)
plt.plot(np.linspace(0, 10), gamma.pdf(np.linspace(0, 10), alpha_mom[0], beta_mom[0]))
# -
# Looping over all months, we can create a grid of plots for the distribution of rainfall, using the gamma distribution:
# +
axs = precip.hist(normed=True, figsize=(12, 8), sharex=True, sharey=True, bins=15, grid=False)
for ax in axs.ravel():
# Get month
m = ax.get_title()
# Plot fitted distribution
x = np.linspace(*ax.get_xlim())
ax.plot(x, gamma.pdf(x, alpha_mom[m], beta_mom[m]))
# Annotate with parameter estimates
label = 'alpha = {0:.2f}\nbeta = {1:.2f}'.format(alpha_mom[m], beta_mom[m])
ax.annotate(label, xy=(10, 0.2))
plt.tight_layout()
# -
# Maximum Likelihood
# ==================
#
# **Maximum likelihood** (ML) fitting is usually more work than the method of moments, but it is preferred as the resulting estimator is known to have good theoretical properties.
#
# There is a ton of theory regarding ML. We will restrict ourselves to the mechanics here.
#
# Say we have some data $y = y_1,y_2,\ldots,y_n$ that is distributed according to some distribution:
#
# <div style="font-size: 120%;">
# $$Pr(Y_i=y_i | \theta)$$
# </div>
# Here, for example, is a **Poisson distribution** that describes the distribution of some discrete variables, typically *counts*:
y = np.random.poisson(5, size=100)
plt.hist(y, bins=12, normed=True)
plt.xlabel('y'); plt.ylabel('Pr(y)')
# The product $\prod_{i=1}^n Pr(y_i | \theta)$ gives us a measure of how **likely** it is to observe values $y_1,\ldots,y_n$ given the parameters $\theta$.
#
# Maximum likelihood fitting consists of choosing the appropriate function $l= Pr(Y|\theta)$ to maximize for a given set of observations. We call this function the *likelihood function*, because it is a measure of how likely the observations are if the model is true.
#
# > Given these data, how likely is this model?
# In the above model, the data were drawn from a Poisson distribution with parameter $\lambda =5$.
#
# $$L(y|\lambda=5) = \frac{e^{-5} 5^y}{y!}$$
#
# So, for any given value of $y$, we can calculate its likelihood:
# +
poisson_like = lambda x, lam: np.exp(-lam) * (lam**x) / (np.arange(x)+1).prod()
lam = 6
value = 10
poisson_like(value, lam)
# -
np.sum(poisson_like(yi, lam) for yi in y)
lam = 8
np.sum(poisson_like(yi, lam) for yi in y)
# We can plot the likelihood function for any value of the parameter(s):
lambdas = np.linspace(0,15)
x = 5
plt.plot(lambdas, [poisson_like(x, l) for l in lambdas])
plt.xlabel('$\lambda$')
plt.ylabel('L($\lambda$|x={0})'.format(x))
# How is the likelihood function different than the probability distribution function (PDF)? The likelihood is a function of the parameter(s) *given the data*, whereas the PDF returns the probability of data given a particular parameter value. Here is the PDF of the Poisson for $\lambda=5$.
lam = 5
xvals = np.arange(15)
plt.bar(xvals, [poisson_like(x, lam) for x in xvals], width=0.2)
plt.xlabel('x')
plt.ylabel('Pr(X|$\lambda$=5)')
# *Why are we interested in the likelihood function?*
#
# A reasonable estimate of the true, unknown value for the parameter is one which **maximizes the likelihood function**. So, inference is reduced to an optimization problem.
# Going back to the rainfall data, if we are using a gamma distribution we need to maximize:
#
# $$\begin{align}l(\alpha,\beta) &= \sum_{i=1}^n \log[\beta^{\alpha} x^{\alpha-1} e^{-x/\beta}\Gamma(\alpha)^{-1}] \cr
# &= n[(\alpha-1)\overline{\log(x)} - \bar{x}\beta + \alpha\log(\beta) - \log\Gamma(\alpha)]\end{align}$$
#
# *N.B.: Its usually easier to work in the log scale*
#
# where $n = 2012 − 1871 = 141$ and the bar indicates an average over all *i*. We choose $\alpha$ and $\beta$ to maximize $l(\alpha,\beta)$.
#
# Notice $l$ is infinite if any $x$ is zero. We do not have any zeros, but we do have an NA value for one of the October data, which we dealt with above.
# ### Finding the MLE
#
# To find the maximum of any function, we typically take the *derivative* with respect to the variable to be maximized, set it to zero and solve for that variable.
#
# $$\frac{\partial l(\alpha,\beta)}{\partial \beta} = n\left(\frac{\alpha}{\beta} - \bar{x}\right) = 0$$
#
# Which can be solved as $\beta = \alpha/\bar{x}$. However, plugging this into the derivative with respect to $\alpha$ yields:
#
# $$\frac{\partial l(\alpha,\beta)}{\partial \alpha} = \log(\alpha) + \overline{\log(x)} - \log(\bar{x}) - \frac{\Gamma(\alpha)'}{\Gamma(\alpha)} = 0$$
#
# This has no closed form solution. We must use ***numerical optimization***!
# Numerical optimization alogarithms take an initial "guess" at the solution, and **iteratively** improve the guess until it gets "close enough" to the answer.
#
# Here, we will use *Newton-Raphson* method, which is a **root-finding algorithm**:
#
# <div style="font-size: 120%;">
# $$x_{n+1} = x_n - \frac{f(x_n)}{f'(x_n)}$$
# </div>
# which is available to us via SciPy:
from scipy.optimize import newton
# Here is a graphical example of how Newton-Raphson converges on a solution, using an arbitrary function:
# %run newton_raphson_plot.py
# To apply the Newton-Raphson algorithm, we need a function that returns a vector containing the **first and second derivatives** of the function with respect to the variable of interest. The second derivative of the gamma distribution with respect to $\alpha$ is:
#
# $$\frac{\partial^2 l(\alpha,\beta)}{\partial \alpha^2} = \frac{1}{\alpha} - \frac{\partial}{\partial \alpha} \left[ \frac{\Gamma(\alpha)'}{\Gamma(\alpha)} \right]$$
# +
from scipy.special import psi, polygamma
dlgamma = lambda a, log_mean, mean_log: np.log(a) - psi(a) - log_mean + mean_log
dl2gamma = lambda a, *args: 1./a - polygamma(1, a)
# -
# where `log_mean` and `mean_log` are $\log{\bar{x}}$ and $\overline{\log(x)}$, respectively. `psi` and `polygamma` are complex functions of the Gamma function that result when you take first and second derivatives of that function.
# Calculate statistics
log_mean = precip.mean().apply(np.log)
mean_log = precip.apply(np.log).mean()
# Time to optimize!
# Alpha MLE for December
alpha_mle = newton(dlgamma, 2, dl2gamma, args=(log_mean[-1], mean_log[-1]))
alpha_mle
# And now plug this back into the solution for beta:
#
# <div style="font-size: 120%;">
# $$ \beta = \frac{\alpha}{\bar{X}} $$
# </div>
beta_mle = alpha_mle/precip.mean()[-1]
beta_mle
# We can compare the fit of the estimates derived from MLE to those from the method of moments:
dec = precip.Dec
dec.hist(normed=True, bins=10, grid=False)
x = np.linspace(0, dec.max())
plt.plot(x, gamma.pdf(x, alpha_mom[-1], beta_mom[-1]), 'm-', label='Moment estimator')
plt.plot(x, gamma.pdf(x, alpha_mle, beta_mle), 'r--', label='ML estimator')
plt.legend()
# For some common distributions, SciPy includes methods for fitting via MLE:
# +
from scipy.stats import gamma
gamma.fit(precip.Dec)
# -
# This fit is not directly comparable to our estimates, however, because SciPy's `gamma.fit` method fits an odd 3-parameter version of the gamma distribution.
# ### Model checking
#
# An informal way of checking the fit of our parametric model is to compare the observed quantiles of the data to those of the theoretical model we are fitting it to. If the model is a good fit, the points should fall on a 45-degree reference line. This is called a **probability plot**.
#
# SciPy includes a `probplot` function that generates probability plots based on the data and a specified distribution.
# +
from scipy.stats import probplot
probplot(precip.Dec, dist=gamma(3.51, scale=0.84), plot=plt);
# -
# ### Example: truncated distribution
#
# Suppose that we observe $Y$ truncated below at $a$ (where $a$ is known). If $X$ is the distribution of our observation, then:
#
# $$ P(X \le x) = P(Y \le x|Y \gt a) = \frac{P(a \lt Y \le x)}{P(Y \gt a)}$$
#
# (so, $Y$ is the original variable and $X$ is the truncated variable)
#
# Then X has the density:
#
# $$f_X(x) = \frac{f_Y (x)}{1−F_Y (a)} \, \text{for} \, x \gt a$$
#
# Suppose $Y \sim N(\mu, \sigma^2)$ and $x_1,\ldots,x_n$ are independent observations of $X$. We can use maximum likelihood to find $\mu$ and $\sigma$.
# First, we can simulate a truncated distribution using a `while` statement to eliminate samples that are outside the support of the truncated distribution.
# +
x = np.random.normal(size=10000)
# Truncation point
a = -1
# Resample until all points meet criterion
x_small = x < a
while x_small.sum():
x[x_small] = np.random.normal(size=x_small.sum())
x_small = x < a
_ = plt.hist(x, bins=100)
# -
# We can construct a log likelihood for this function using the conditional form:
#
# $$f_X(x) = \frac{f_Y (x)}{1−F_Y (a)} \, \text{for} \, x \gt a$$
#
# The denominator normalizes the truncated distribution so that it integrates to one.
# +
from scipy.stats.distributions import norm
trunc_norm = lambda theta, a, x: -(np.log(norm.pdf(x, theta[0], theta[1])) -
np.log(1 - norm.cdf(a, theta[0], theta[1]))).sum()
# -
# For this example, we will use an optimization algorithm, the **Nelder-Mead simplex algorithm**. It has a couple of advantages:
#
# - it does not require derivatives
# - it can optimize (minimize) a vector of parameters
#
# SciPy implements this algorithm in its `fmin` function:
# +
from scipy.optimize import fmin
fmin(trunc_norm, np.array([1,2]), args=(-1, x))
# -
# In general, simulating data is a terrific way of testing your model before using it with real data.
# ## Kernel density estimates
#
# In some instances, we may not be interested in the parameters of a particular distribution of data, but just a smoothed representation of the data at hand. In this case, we can estimate the disribution *non-parametrically* (i.e. making no assumptions about the form of the underlying distribution) using kernel density estimation.
# Some random data
y = np.random.normal(10, size=15)
y
# The kernel estimator is a sum of symmetric densities centered at each observation. The selected kernel function determines the shape of each component while the **bandwidth** determines their spread. For example, if we use a Gaussian kernel function, the variance acts as the bandwidth.
x = np.linspace(7, 13, 100)
# Smoothing parameter
s = 0.3
# Calculate the kernels
kernels = np.transpose([norm.pdf(x, yi, s) for yi in y])
plt.plot(x, kernels, 'k:')
plt.plot(x, kernels.sum(1))
plt.plot(y, np.zeros(len(y)), 'ro', ms=10)
# SciPy implements a Gaussian KDE that automatically chooses an appropriate bandwidth. Let's create a bi-modal distribution of data that is not easily summarized by a parametric distribution:
# +
# Create a bi-modal distribution with a mixture of Normals.
x1 = np.random.normal(0, 2, 50)
x2 = np.random.normal(5, 1, 50)
# Append by row
x = np.r_[x1, x2]
# -
plt.hist(x, bins=10, normed=True)
# +
from scipy.stats import kde
density = kde.gaussian_kde(x)
xgrid = np.linspace(x.min(), x.max(), 100)
plt.hist(x, bins=8, normed=True)
plt.plot(xgrid, density(xgrid), 'r-')
# -
# ### Exercise: Comparative Chopstick Effectiveness
#
# A few researchers set out to determine what the optimal length for chopsticks is. The dataset `chopstick-effectiveness.csv` includes measurements of "Food Pinching Efficiency" across a range of chopstick lengths for 31 individuals.
#
# Use the method of moments or MLE to calculate the mean and variance of food pinching efficiency for each chopstick length. This means you need to select an appropriate distributional form for this data.
# +
# Write your answer here
| notebooks/2. Density Estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="CKfkSFu7nlVg"
import tensorflow as tf
# + id="pznVa656noFn" outputId="6d59a2d3-912d-4d6c-ada3-4ec34ac04d0e" colab={"base_uri": "https://localhost:8080/"}
# Load MNIST data
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
# Preprocessing
x_train = x_train / 255.0
x_test = x_test / 255.0
# Add one domention to make 3D images
x_train = x_train[...,tf.newaxis]
x_test = x_test[...,tf.newaxis]
# Track the data type
dataType, dataShape = x_train.dtype, x_train.shape
print(f"Data type and shape x_train: {dataType} {dataShape}")
labelType, labelShape = y_train.dtype, y_train.shape
print(f"Data type and shape y_train: {labelType} {labelShape}")
# + id="zSZB8zVGq46i" outputId="43d2600e-378c-4c50-c509-c22223df0e5e" colab={"base_uri": "https://localhost:8080/", "height": 268}
im_list = []
n_samples_to_show = 16
c = 0
for i in range(n_samples_to_show):
im_list.append(x_train[i])
# Visualization
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import ImageGrid
fig = plt.figure(figsize=(4., 4.))
# Ref: https://matplotlib.org/3.1.1/gallery/axes_grid1/simple_axesgrid.html
grid = ImageGrid(fig, 111, # similar to subplot(111)
nrows_ncols=(4, 4), # creates 2x2 grid of axes
axes_pad=0.1, # pad between axes in inch.
)
# Show image grid
for ax, im in zip(grid, im_list):
# Iterating over the grid returns the Axes.
ax.imshow(im[:,:,0], 'gray')
plt.show()
# + [markdown] id="6cjdho1KupbS"
# ## Training
# + id="Z9X1PzYstWKe" outputId="ab33d448-044e-40c0-ddd7-e925a6c8a2ef" colab={"base_uri": "https://localhost:8080/"}
# Model building
NUM_CLASSES = 10
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(16, (3, 3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Conv2D(32, (3, 3), activation='relu'),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation='relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(32, activation='relu'),
tf.keras.layers.Dense(NUM_CLASSES, activation='sigmoid')]
)
# Compiling the model with the high-level keras
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=False),
metrics=['accuracy'])
# Model training
model.fit(x_train, y_train, epochs=5)
# + [markdown] id="oD2sc_rNurqN"
# ## Evaluation
# + id="EfO3wC__tdj5" outputId="ab7b244a-3c81-4a9b-ae82-47c73b7b7d59" colab={"base_uri": "https://localhost:8080/"}
eval_loss, eval_acc = model.evaluate(x_test, y_test, verbose=1)
print('Eval accuracy percentage: {:.2f}'.format(eval_acc * 100))
# + id="6gf9IHenvlTH"
| codes/ipython/neural_networks/CNNs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="title_ID"></a>
# # JWST calwebb_image3, tweakreg unit tests
#
# <span style="color:red"> **Instruments Affected**</span>: NIRCam, NIRISS, MIRI, FGS
#
# ### Table of Contents
#
# <div style="text-align: left">
#
# <br> [Introduction](#intro)
# <br> [JWST Unit Tests](#unit)
# <br> [Defining Terms](#terms)
# <br> [Test Description](#description)
# <br> [Data Description](#data_descr)
# <br> [Imports](#imports)
# <br> [Convenience Functions](#functions)
# <br> [Perform Tests](#testing)
# <br> [About This Notebook](#about)
# <br>
#
# </div>
# <a id="intro"></a>
# # Introduction
#
# This is the validation notebook that displays the unit tests for the Tweakreg step in calwebb_image3. This notebook runs and displays the unit tests that are performed as a part of the normal software continuous integration process. For more information on the pipeline visit the links below.
#
# * Pipeline description: https://jwst-pipeline.readthedocs.io/en/latest/jwst/tweakreg/index.html
#
# * Pipeline code: https://github.com/spacetelescope/jwst/tree/master/jwst/
#
# [Top of Page](#title_ID)
# <a id="unit"></a>
# # JWST Unit Tests
#
# JWST unit tests are located in the "tests" folder for each pipeline step within the [GitHub repository](https://github.com/spacetelescope/jwst/tree/master/jwst/), e.g., ```jwst/tweakreg/tests```.
#
# * Unit test README: https://github.com/spacetelescope/jwst#unit-tests
#
#
# [Top of Page](#title_ID)
# <a id="terms"></a>
# # Defining Terms
#
# These are terms or acronymns used in this notebook that may not be known a general audience.
#
# * JWST: <NAME> Space Telescope
#
# * NIRCam: Near-Infrared Camera
#
#
# [Top of Page](#title_ID)
# <a id="description"></a>
# # Test Description
#
# Unit testing is a software testing method by which individual units of source code are tested to determine whether they are working sufficiently well. Unit tests do not require a separate data file; the test creates the necessary test data and parameters as a part of the test code.
#
#
# [Top of Page](#title_ID)
# <a id="data_descr"></a>
# # Data Description
#
# Data used for unit tests is created on the fly within the test itself, and is typically an array in the expected format of JWST data with added metadata needed to run through the pipeline.
#
#
# [Top of Page](#title_ID)
# <a id="imports"></a>
# # Imports
#
# * tempfile for creating temporary output products
# * pytest for unit test functions
# * jwst for the JWST Pipeline
# * IPython.display for display pytest reports
#
# [Top of Page](#title_ID)
import tempfile
import os
import pytest
import jwst
from IPython.display import IFrame
from IPython.core.display import HTML
# <a id="functions"></a>
# # Convenience Functions
#
# Here we define any convenience functions to help with running the unit tests.
#
# [Top of Page](#title_ID)
# <a id="testing"></a>
# # Perform Tests
#
# Below we run the unit tests for the Tweakreg step.
#
# [Top of Page](#title_ID)
# +
print("Testing JWST Pipeline {}".format(jwst.__version__))
jwst_dir = os.path.dirname(jwst.__file__)
tweakreg = os.path.join(jwst_dir, 'tweakreg')
associations = os.path.join(jwst_dir, 'associations')
datamodels = os.path.join(jwst_dir, 'datamodels')
stpipe = os.path.join(jwst_dir, 'stpipe')
regtest = os.path.join(jwst_dir, 'regtest')
with tempfile.TemporaryDirectory() as tmpdir:
outdir = os.path.join(tmpdir, 'regtest_report.html')
# !pytest {tweakreg} -v --ignore={associations} --ignore={datamodels} --ignore={stpipe} --ignore={regtest} --html={outdir} --self-contained-html
with open(os.path.join(tmpdir, "regtest_report.html")) as report_file:
html_report = "".join(report_file.readlines())
# -
HTML(html_report)
# <a id="about"></a>
# ## About This Notebook
# **Author:** <NAME>, Staff Scientist, NIRCam
# <br>**Updated On:** 01/07/2021
# [Top of Page](#title_ID)
# <img style="float: right;" src="./stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="stsci_pri_combo_mark_horizonal_white_bkgd" width="200px"/>
| jwst_validation_notebooks/regression_test/tweakreg/jwst_tweakreg_unit_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cheers!! Stats with Beers
#
# This notebook contains the solutions to the exercises that are in the lesson [1_Cheers_Stats_Beers](http://nbviewer.jupyter.org/github/engineersCode/EngComp/blob/master/modules/2_takeoff/1_Cheers_Stats_Beers.ipynb).
# ##### Exercise
#
# Write a function that calculates the percentage of missing values for a certain array. Use the function to calculate the percentage of missing values for the `abv` and `ibu` data sets.
#
# For the original array, before cleaning, rememeber that you can access the values of a Serie by doing `serie.value` (eg. `abv_serie.values`).
#
# We need the clean and original data from the original notebook. We will replicate those steps here.
import pandas
import numpy
beers = pandas.read_csv("../../../data/beers.csv")
abv_serie = beers['abv']
abv_clean = abv_serie.dropna()
ibu_serie = beers['ibu']
ibu_clean = ibu_serie.dropna()
# ##### <span style="color:green"> Solution </span>
def percentage_missing_val(original, clean):
"""Calculates the percentage of missing values of an array
Arguments:
----------
original : array, original array before removing NaN.
clean : array, clean array after removing NaN.
Returns:
--------
pct : float, percentage of missing values.
"""
number_of_missing_values = len(original) - len(clean)
ratio_of_missing_values = number_of_missing_values / len(original)
pct = ratio_of_missing_values * 100
return pct
# +
pct_abv = percentage_missing_val(abv_serie.values, abv_clean.values)
print(pct_abv)
# +
pct_ibu = percentage_missing_val(ibu_serie.values, ibu_clean.values)
print(pct_ibu)
# -
# Execute this cell to load the notebook's style sheet, then ignore it
from IPython.core.display import HTML
css_file = '../../../style/custom.css'
HTML(open(css_file, "r").read())
| modules/2_takeoff/exercises_solutions/1_Cheers_Stats_Beers_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Automated Machine Learning
# _**Prepare Data using `azureml.dataprep` for Remote Execution (DSVM)**_
#
# ## Contents
# 1. [Introduction](#Introduction)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Results](#Results)
# 1. [Test](#Test)
# ## Introduction
# In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).
#
# Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.
#
# In this notebook you will learn how to:
# 1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.
# 2. Pass the `Dataflow` to AutoML for a local run.
# 3. Pass the `Dataflow` to AutoML for a remote run.
# ## Setup
#
# Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros.
# Opt-in diagnostics for better experience, quality, and security of future releases.
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics = True)
# As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
# +
import logging
import time
import pandas as pd
import azureml.core
from azureml.core.compute import DsvmCompute
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
import azureml.dataprep as dprep
from azureml.train.automl import AutoMLConfig
# +
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = 'automl-dataprep-remote-dsvm'
# project folder
project_folder = './sample_projects/automl-dataprep-remote-dsvm'
experiment = Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ## Data
# +
# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.
# The data referenced here was pulled from `sklearn.datasets.load_digits()`.
simple_example_data_root = 'https://dprepdata.blob.core.windows.net/automl-notebook-data/'
X = dprep.auto_read_file(simple_example_data_root + 'X.csv').skip(1) # Remove the header row.
# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)
# and convert column types manually.
# Here we read a comma delimited file and convert all columns to integers.
y = dprep.read_csv(simple_example_data_root + 'y.csv').to_long(dprep.ColumnSelector(term='.*', use_regex = True))
# -
# You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets.
X.skip(1).head(5)
# ## Train
#
# This creates a general AutoML settings object applicable for both local and remote runs.
automl_settings = {
"iteration_timeout_minutes" : 10,
"iterations" : 2,
"primary_metric" : 'AUC_weighted',
"preprocess" : False,
"verbosity" : logging.INFO,
"n_cross_validations": 3
}
# ### Create or Attach a Remote Linux DSVM
# +
dsvm_name = 'mydsvmc'
try:
while ws.compute_targets[dsvm_name].provisioning_state == 'Creating':
time.sleep(1)
dsvm_compute = DsvmCompute(ws, dsvm_name)
print('Found existing DVSM.')
except:
print('Creating a new DSVM.')
dsvm_config = DsvmCompute.provisioning_configuration(vm_size = "Standard_D2_v2")
dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)
dsvm_compute.wait_for_completion(show_output = True)
print("Waiting one minute for ssh to be accessible")
time.sleep(60) # Wait for ssh to be accessible
# +
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
conda_run_config = RunConfiguration(framework="python")
conda_run_config.target = dsvm_compute
cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])
conda_run_config.environment.python.conda_dependencies = cd
# -
# ### Pass Data with `Dataflow` Objects
#
# The `Dataflow` objects captured above can also be passed to the `submit` method for a remote run. AutoML will serialize the `Dataflow` object and send it to the remote compute target. The `Dataflow` will not be evaluated locally.
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
path = project_folder,
run_configuration=conda_run_config,
X = X,
y = y,
**automl_settings)
remote_run = experiment.submit(automl_config, show_output = True)
remote_run
# ## Results
# #### Widget for Monitoring Runs
#
# The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
# #### Retrieve All Child Runs
# You can also use SDK methods to fetch all the child runs and see individual metrics that we log.
# +
children = list(remote_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ### Retrieve the Best Model
#
# Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
best_run, fitted_model = remote_run.get_output()
print(best_run)
print(fitted_model)
# #### Best Model Based on Any Other Metric
# Show the run and the model that has the smallest `log_loss` value:
lookup_metric = "log_loss"
best_run, fitted_model = remote_run.get_output(metric = lookup_metric)
print(best_run)
print(fitted_model)
# #### Model from a Specific Iteration
# Show the run and the model from the first iteration:
iteration = 0
best_run, fitted_model = remote_run.get_output(iteration = iteration)
print(best_run)
print(fitted_model)
# ## Test
#
# #### Load Test Data
# +
from sklearn import datasets
digits = datasets.load_digits()
X_test = digits.data[:10, :]
y_test = digits.target[:10]
images = digits.images[:10]
# -
# #### Testing Our Best Fitted Model
# We will try to predict 2 digits and see how our model works.
# +
#Randomly select digits and test
from matplotlib import pyplot as plt
import numpy as np
for index in np.random.choice(len(y_test), 2, replace = False):
print(index)
predicted = fitted_model.predict(X_test[index:index + 1])[0]
label = y_test[index]
title = "Label value = %d Predicted value = %d " % (label, predicted)
fig = plt.figure(1, figsize=(3,3))
ax1 = fig.add_axes((0,0,.8,.8))
ax1.set_title(title)
plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')
plt.show()
# -
# ## Appendix
# ### Capture the `Dataflow` Objects for Later Use in AutoML
#
# `Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage.
# sklearn.digits.data + target
digits_complete = dprep.auto_read_file('https://dprepdata.blob.core.windows.net/automl-notebook-data/digits-complete.csv')
# `digits_complete` (sourced from `sklearn.datasets.load_digits()`) is forked into `dflow_X` to capture all the feature columns and `dflow_y` to capture the label column.
print(digits_complete.to_pandas_dataframe().shape)
labels_column = 'Column64'
dflow_X = digits_complete.drop_columns(columns = [labels_column])
dflow_y = digits_complete.keep_columns(columns = [labels_column])
| how-to-use-azureml/automated-machine-learning/dataprep-remote-execution/auto-ml-dataprep-remote-execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="3M2Fdn6cHTaT"
import torch
import numpy as np
import pandas as pd
import torch.nn as nn
from torch.utils.data import DataLoader, TensorDataset
# + id="6XFw-u7KIPGA"
df = pd.read_csv('data.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="j8JIhUWyIBA1" outputId="3d6941f4-06bb-424a-f172-68182067aa65"
device = "cuda" if torch.cuda.is_available() else "cpu"
print("Using {} device".format(device))
# + id="dmxFX_gd09SA"
class NeuralNetwork(nn.Module):
def __init__(self):
super(NeuralNetwork, self).__init__()
self.layer_1 = nn.Sequential(
nn.Linear(1, 1)
)
def forward(self, x):
logits = self.layer_1(x)
return logits
# + id="qqo4GsemIAhn"
model = NeuralNetwork().to(device)
# + colab={"base_uri": "https://localhost:8080/"} id="NXYD0rZSIKnO" outputId="d25a48f0-c6b3-40de-adad-69e79869f60b"
print(model)
# + id="gUj17q20I1yt"
loss_fn = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.005)
# + id="4NqBIsDnJEgZ"
X_train = df['Celsius']
Y_train = df['Fahrenheit']
# + colab={"base_uri": "https://localhost:8080/"} id="t6xdPsRTLUl2" outputId="75d324a7-669a-45d4-ab13-1f29164013c7"
train_target = torch.tensor(Y_train.values.astype(np.float32))
train = torch.tensor(X_train.values.astype(np.float32))
train_tensor = TensorDataset(train, train_target)
train_loader = DataLoader(dataset=train_tensor, batch_size=1, shuffle=True)
for X, y in train_loader:
print("Shape of X: ", X.shape, X.dtype)
print("Shape of y: ", y.shape, y.dtype)
break
# + id="kV92h9DgMGF7"
def train(dataloader, model, loss_fn, optimizer):
size = len(dataloader.dataset)
for batch, (X, y) in enumerate(dataloader):
X, y = X.to(device), y.to(device)
# Compute prediction error
pred = model(X)
loss = loss_fn(pred, y)
# Backpropagation
optimizer.zero_grad() # set the gradients to zero before backpropragation
loss.backward()
optimizer.step()
if batch % 100 == 0:
loss, current = loss.item(), batch * len(X)
print(f"loss: {loss:>7f} [{current:>5d}/{size:>5d}]")
# + colab={"base_uri": "https://localhost:8080/"} id="RsY87OMbMYT_" outputId="7d6a8acb-3f0b-4b5e-a5a9-49c79acac1a3"
epochs = 450
for t in range(epochs):
print(f"Epoch {t+1}\n-------------------------------")
train(train_loader, model, loss_fn, optimizer)
print("Done!")
# + id="YysoaSZlNw9F"
def c_to_f(c):
return 9/5 * c + 32
# + colab={"base_uri": "https://localhost:8080/"} id="cYaMjBE1NyES" outputId="33d6d6b4-45a8-4fb8-b8fd-ece695b999dc"
Temp_C = [0, 15, 35] # 32, 59, 95
for c in Temp_C:
with torch.no_grad():
model.eval()
Temp_F = model(torch.tensor([np.float32(c)]))
print(f'C {c} = F {np.around(Temp_F[0])}')
# + colab={"base_uri": "https://localhost:8080/"} id="x7vdC6nzSv9I" outputId="4d5f6360-f846-4427-b1a5-12971f3c1a7c"
for c in Temp_C:
print(f'C {c} = F {c_to_f(c)}')
# + colab={"base_uri": "https://localhost:8080/"} id="Le6lMe_ZUKe4" outputId="1c3ff555-2de4-44e1-a78c-c3def115f6ca"
s = torch.sum(model.layer_1[0].weight.data)
s # 9/5 = 1.8
# + colab={"base_uri": "https://localhost:8080/"} id="Zvq2OXABUWvi" outputId="910c933a-5e14-491b-96d8-ed4ffc30c7dd"
s = torch.sum(model.layer_1[0].bias.data)
s # 32
| ai/notebooks/C_to_F_(pytorch).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # semivariance module
#
# Available classes and functions:
#
# - ArealSemivariance: Class calculates semivariance of areas for Poisson Kriging (area to area and area to point),
# - RegularizedSemivariogram: Class performs deconvolution of semivariogram of areal data,
# - calculate_covariance: Function calculates covariance of a given set of points,
# - calculate_semivariance: Function calculates semivariance of a given set of points,
# - calculate_weighted_semivariance: Function calculates weighted semivariance,
# - calculate_directional_semivariogram: Function calculates semivariogram within specified ellipse,
# - build_variogram_point_cloud: Function creates OrderedDict with lags and variances between points within specific lag,
# - show_variogram_cloud: function shows boxplots of lags and squared differences between ponts' values within specific lag,
# - calc_semivariance_from_pt_cloud: based on Point Cloud semivariogram is calculated,
# - remove_outliers: removes outliers from point cloud variogram,
# - TheoreticalSemivariogram: Class calculates theoretical semivariogram.
# ## ```ArealSemivariance```
#
# ### Class initialization
#
# ```python
# pyinterpolate.semivariance.ArealSemivariance(
# areal_data,
# areal_step_size,
# max_areal_range,
# areal_points_data,
# weighted_semivariance=False,
# verbose=False)
#
# ```
#
# Class calculates semivariance of areas for Poisson Kriging (area to area and area to point).
#
#
# INITIALIZATION PARAMS:
#
# - **areal_data**: (_numpy array_ / _list_)
#
# ```python
# [area_id, area_geometry, centroid x,centroid y, value]
# ```
# - **areal_step_size**: (_float_) step size for search radius,
# - **max_areal_range**: (*float*) max distance to perform distance and semivariance calculations,
# - **areal_points_data**: (_numpy array_ / _list_)
#
# ```python
# [
# area_id,
# [point_position_x, point_position_y, value]
# ]
# ```
# - **weighted_semivariance**: (_bool_) if ```False``` then each distance is treated equally when calculating theoretical semivariance; if ```True``` then semivariances closer to the point of origin have more weight,
# - **verbose**: (_bool_) if ```True``` then all messages are printed, otherwise nothing.
#
# ### Class public methods:
#
# - **regularize_semivariogram**: Function calculates regularized point support semivariogram,
# - **show_semivariograms**: Function shows semivariograms calculated by the class: Empirical semivariogram, Theoretical model, Inblock Semivariance, Within-block semivariogram, Between blocks semivariogram, Regularized output.
#
# ---
#
# ### ```ArealSemivariance.regularize_semivariogram()```
#
# ```python
# ArealSemivariance.regularize_semivariogram(
# self,
# within_block_semivariogram=None,
# between_blocks_semivariogram=None,
# empirical_semivariance=None,
# theoretical_semivariance_model=None)
# ```
#
# Function calculates regularized point support semivariogram in the form given in:
#
# > <NAME>., Kriging and Semivariogram Deconvolution in the Presence of Irregular Geographical Units, Mathematical Geology 40(1), 101-128, 2008
#
# Function has the form:
#
# $$\gamma_{v(h)} = \gamma(v, v_h) - \gamma_{h}(v, v)$$
#
# where:
#
# - $\gamma_{v(h)}$ - regularized semivariogram,
# - $\gamma(v, v_h)$ - semivariogram value between any two blocks separated by the distance $h$,
# - $\gamma_{h}(v, v)$ - arithmetical average of within-block semivariogram.
#
# INPUT:
#
# - **within_block_semivariogram**: (_numpy array_) mean semivariance between the blocks:
#
# $$ \gamma_{h}(v, v) = \frac{1}{2*N(h)} \sum^{N(h)}_{a=1} [\gamma(v\alpha, v\alpha) + \gamma(v\alpha+h, v\alpha+h)]$$
#
# where:
#
# $\gamma(v\alpha, v\alpha)$ and $\gamma(v\alpha+h, v\alpha+h)$ are the inblock semivariances of block $\alpha$ and block $\alpha+h$ separated by the distance $h$ weighted by the inblock population.
#
# - **between_blocks_semivariogram**: (_numpy array_) semivariance between all blocks calculated from the theoretical model,
# - **empirical_semivariance**: (_numpy array_) empirical semivariance between area centroids, ```default=None```, if ```None``` is provided then empirical semivariance is computed by the ```_calculate_empirical_semivariance``` method from area centroids,
# - **theoretical_semivariance_model**: (_TheoreticalSemivariogram_) theoretical semivariance model from ```TheoreticalSemivariance``` class, default is ```None```, if ```None``` is provided then theoretical model is derived from area centroids and empirical semivariance.
#
# OUTPUT:
#
# - **semivariance**: (```numpy array```) of lag, semivariance values and number of areas within lag where:
#
# ```semivariance[0] = array of lags```;
#
# ```semivariance[1] = array of lag's values```;
#
# ```semivariance[2] = array of number of points in each lag```.
#
# ---
#
# ### ```ArealSemivariance.show_semivariograms()```
#
# ```python
# ArealSemivariance.show_semivariograms(self)
# ```
#
# Function shows semivariograms calculated by the class: Empirical semivariogram, Theoretical model, Inblock Semivariance, Within-block semivariogram, Between blocks semivariogram, Regularized output.
# ***
# ## ```RegularizedSemivariogram```
#
# ### Class initialization
#
# ```python
# pyinterpolate.semivariance.RegularizedSemivariogram(self)
#
# ```
#
# Class performs deconvolution of semivariogram of areal data. Whole procedure is based on the iterative process described in:
#
# > <NAME>., Kriging and Semivariogram Deconvolution in the Presence of Irregular Geographical Units, Mathematical Geology 40(1), 101-128, 2008
#
# Class works as follow:
#
# - initialize your object (no parameters),
# - then use ```fit()``` method to build initial point support model,
# - then use ```transform()``` method to perform semivariogram regularization,
# - save semivariogram model with ```export_model()``` method.
#
# ### Class public methods:
#
# - **fit** - fits areal data and point support data into a model, initialize experimental semivariogram, theoretical semivariogram model, regularized point support model and deviation.
# - **transform** - performs semivariogram regularization, which is an iterative process,
# - **export_regularized_model** - Function exports final regularized model parameters into specified csv file.
# - **show_baseline_semivariograms** - Function shows experimental semivariogram, initial theoretical semivariogram and initial regularized semivariogram after fit() operation.
# - **show_semivariograms** - plots experimental semivariogram of area data, theoretical curve of area data, regularized model values and regularized model theoretical curve.
#
# ---
#
# ### ```RegularizedSemivariogram.fit()```
#
# ```python
# RegularizedSemivariogram.fit(self,
# areal_data,
# areal_step_size,
# max_areal_range,
# point_support_data,
# weighted_lags=True,
# store_models=False)
# ```
#
# Function fits area and point support data to the initial regularized models.
#
# INPUT:
#
# - **areal_data**: (_numpy array_) areal data prepared with the function ```prepare_areal_shapefile()```, where data is a ```numpy array```in the form:
#
# ```python
# [area_id, area_geometry, centroid x, centroid y, value]
# ```
# - **areal_step_size**: (_float_) step size between each lag, usually it is a half of distance between lags,
# - **max_areal_range**: (*float*) max distance to perform distance and semivariance calculations,
# - **point_support_data**: (_numpy array_) point support data prepared with the function ```get_points_within_area()```, where data is a ```numpy array``` in the form:
#
# ```python
# [
# area_id,
# [point_position_x, point_position_y, value]
# ]
# ```
# - **weighted_lags**: (_bool_) lags weighted by number of points; if ```True``` then during semivariogram fitting error of each model is weighted by number of points for each lag. In practice it means that more reliable data (lags) have larger weights and semivariogram is modeled to better fit to those lags,
# - **store_models**: (_bool_) if ```True``` then experimental, regularized and theoretical models are stored in lists after each iteration. It is important for a debugging process.
#
# OUTPUT:
#
# None, class is updating its internal parameters. Usually after fitting you should perform regularization with ```transform()``` method.
#
# ---
#
# ### ```RegularizedSemivariogram.transform()```
#
# ```python
# RegularizedSemivariogram.transform(self,
# max_iters=25,
# min_deviation_ratio=0.01,
# min_diff_decrease=0.01,
# min_diff_decrease_reps=3)
# ```
#
# Function transofrms fitted data and performs semivariogram regularization iterative procedure.
#
# INPUT:
#
# - **max_iters**: (_int_) maximum number of iterations,
# - **min_deviation_ratio**: (_float_) minimum ratio between deviation and initial deviation (D(i) / D(0)) below each algorithm is stopped,
# - **min_diff_decrease**: (_float_) minimum absolute difference between new and optimal deviation divided by optimal deviation: ABS(D(i) - D(opt)) / D(opt). If it is recorded ```n``` times (controled by the ```min_diff_d_stat_reps``` param) then algorithm is stopped,
# - **min_diff_decrease_reps**: (_int_) number of iterations when algorithm is stopped if condition ```min_diff_d_stat``` is fulfilled.
#
# OUTPUT:
#
# None, class is updating its internal parameters. Usually after transforming you should export your theoretical model with ```export_regularized_model()``` method.
#
# ---
#
# ### ```RegularizedSemivariogram.export_regularized_model()```
#
# ```python
# RegularizedSemivariogram.export_regularized_model(self,
# filename)
# ```
#
# Function exports final regularized model parameters into specified csv file.
#
# INPUT:
#
# - **filename**: (_str_) filename for model parameters (nugget, sill, range, model type).
#
# OUTPUT:
#
# Method saves regularized model into csv file.
#
# ---
#
# ### ```RegularizedSemivariogram.show_baseline_semivariograms()```
#
# ```python
# RegularizedSemivariogram.show_baseline_semivariograms(self)
# ```
#
# Function shows experimental semivariogram, initial theoretical semivariogram and initial regularized semivariogram after ```fit()``` operation.
#
# ---
#
# ### ```RegularizedSemivariogram.show_semivariograms()```
#
# ```python
# RegularizedSemivariogram.show_semivariograms(self)
# ```
#
# Function shows experimental semivariogram, theoretical semivariogram and regularized semivariogram after semivariogram regularization with ```transform()``` method.
# ***
# ## ```calculate_covariance()```
#
# ```python
# pyinterpolate.semivariance.calculate_covariance(
# data,
# step_size,
# max_range)
# ```
#
# Function calculates covariance of a given set of points.
#
# Equation for calculation is:
#
# $$covariance = \frac{1}{N} * \sum_{i=1}^{N} [z(x_{i} + h) * z(x_{i})] - u^{2}$$
#
# where:
#
# $N$ - number of observation pairs,
#
# $h$ - distance (lag),
#
# $z(x_{i})$ - value at location $z_{i}$,
#
# $(x_{i} + h)$ - location at a distance $h$ from $x_{i}$,
#
# $u$ - mean of observations at a given lag distance.
#
#
# INPUT:
#
# - **data**: (_numpy array_) coordinates and their values,
# - **step_size**: (_float_) distance between lags within each points are included in the calculations,
# - **max_range**: (*float*) maximum range of analysis.
#
#
# OUTPUT:
#
# - (```numpy array```) covariance - array of pair of lag and covariance values where:
#
# ```covariance[0] = array of lags```;
#
# ```covariance[1] = array of lag's values```;
#
# ```covariance[2] = array of number of points in each lag```.
# ***
# ## ```calculate_semivariance()```
#
# ```python
# pyinterpolate.semivariance.calculate_semivariance(
# data,
# step_size,
# max_range)
# ```
#
# Function calculates semivariance of a given set of points.
#
# Equation for calculation is:
#
# $$semivariance = \frac{1}{2N} * \sum_{i=1}^{N} [z(x_{i} + h) - z(x_{i})]^{2}$$
#
# where:
#
# $N$ - number of observation pairs,
#
# $h$ - distance (lag),
#
# $z(x_{i})$ - value at location $z_{i}$,
#
# $(x_{i} + h)$ - location at a distance $h$ from $x_{i}$,
#
#
# INPUT:
#
# - **data**: (_numpy array_) coordinates and their values,
# - **step_size**: (_float_) distance between lags within each points are included in the calculations,
# - **max_range**: (*float*) maximum range of analysis.
#
#
# OUTPUT:
#
# - (```numpy array```) semivariance - array of pair of lag and semivariance values where:
#
# ```semivariance[0] = array of lags```;
#
# ```semivariance[1] = array of lag's values```;
#
# ```semivariance[2] = array of number of points in each lag```.
# ***
# ## ```calculate_weighted_semivariance()```
#
# ```python
# pyinterpolate.semivariance.calculate_weighted_semivariance(
# data,
# step_size,
# max_range)
# ```
#
# Function calculates weighted semivariance following _Monestiez et al._:
#
# > <NAME>, <NAME>, <NAME>, <NAME>, <NAME>: Comparison of model based geostatistical methods in ecology: application to fin whale spatial distribution in northwestern Mediterranean Sea. In Geostatistics Banff 2004 Volume 2. Edited by: <NAME>, Deutsch CV. Dordrecht, The Netherlands, Kluwer Academic Publishers; 2005:777-786.
#
#
# > <NAME>, <NAME>, <NAME>, <NAME>, <NAME>: Geostatistical modelling of spatial distribution of Balenoptera physalus in the northwestern Mediterranean Sea from sparse count data and heterogeneous observation efforts. Ecological Modelling 2006 in press.
#
# Equation for calculation is:
#
# $$s(h) = \frac{1}{2*\sum_{a=1}^{N(h)} c_{a}} * \sum_{a=1}^{N(h)} c_{a}*(z(u_{a}) - z(u_{a} + h))^2 - m'$$
#
# where:
#
# $$c_{a} = \frac{n(u_{a}) * n(u_{a} + h)}{n(u_{a}) + n(u_{a} + h)}$$
#
# where:
#
# $s(h)$ Semivariogram of the risk,
#
# $n(u_{a})$ - size of the population at risk in the unit a,
#
# $z(u_{a})$ - mortality rate at the unit a,
#
# $u_{a} + h$ - area at the distance (h) from the analyzed area,
#
# $m'$ - population weighted mean of rates.
#
# INPUT:
#
# - **data**: (_numpy array_) coordinates and their values and weights:
#
# ```python
# [coordinate x, coordinate y, value, weight]
# ```
# - **step_size**: (_float_) distance between lags within each points are included in the calculations,
# - **max_range**: (*float*) maximum range of analysis.
#
#
# OUTPUT:
#
# - (```numpy array```) semivariance - array of pair of lag and semivariance values where:
#
# ```semivariance[0] = array of lags```;
#
# ```semivariance[1] = array of lag's values```;
#
# ```semivariance[2] = array of number of points in each lag```.
# ***
# + [markdown] pycharm={"name": "#%% md\n"}
# ## ```calculate_directional_semivariogram()```
#
# ```python
# pyinterpolate.semivariance.calculate_directional_semivariogram(
# data,
# step_size,
# max_range,
# direction=0,
# tolerance=0.1)
# ```
#
# Function calculates directional semivariogram of points. Semivariance is calculated as:
#
# $$semivariance = \frac{1}{2N} * \sum_{i=1}^{N} [z(x_{i} + h) - z(x_{i})]^{2}$$
#
# where:
#
# - $N$ - number of observation pairs,
# - $h$ - distance (lag),
# - $z(x_{i})$ - value at location $z_{i}$,
# - $(x_{i} + h)$ - location at a distance $h$ from $x_{i}$.
#
# INPUT:
#
# - **data**: (*numpy array8) coordinates and their values,
# - **step_size**: (*float*) distance between lags within each points are included in the calculations,
# - **max_range**: (*float*) maximum range of analysis,
# - **direction**: (*float*) direction of semivariogram, values from 0 to 360 degrees:
#
# -- 0 or 180: is NS direction,
#
# -- 90 or 270 is EW direction,
#
# -- 30 or 210 is NE-SW direction,
#
# -- 120 or 300 is NW-SE direction,
#
# - tolerance: (float) value in range (0-1) normalized to [0 : 0.5] to select tolerance of semivariogram. If tolerance
# is 0 then points must be placed at a single line with beginning in the origin of coordinate system and angle
# given by y axis and direction parameter. If tolerance is greater than 0 then semivariance is estimated
# from elliptical area with major axis with the same direction as the line for 0 tolerance and minor axis
# of a size:
#
# $$(tolerance * step\_size)$$
#
# and major axis (pointed in NS direction):
#
# $$((1 - tolerance) * step\_size)$$
#
# and baseline point at a center of ellipse. Tolerance == 1 (normalized to 0.5) creates omnidirectional semivariogram.
#
# OUTPUT:
#
# - (*numpy array*) **semivariance** - array of pair of lag and semivariance values where:
#
# -- semivariance[0] = array of lags;
#
# -- semivariance[1] = array of lag's values;
#
# -- semivariance[2] = array of number of points in each lag.
# + [markdown] pycharm={"name": "#%% md\n"}
# ***
# -
# ## ```build_variogram_point_cloud()```
#
# ```python
# pyinterpolate.semivariance.build_variogram_point_cloud(
# data,
# step_size,
# max_range)
# ```
#
# Function calculates variogram point cloud of a given set of points for a given set of distances. Variogram is calculated as a squared difference of each point against other point within range specified by step_size parameter.
#
# INPUT:
#
# - **data**: (*numpy array*) coordinates and their values and weights:
#
# ```python
# [coordinate x, coordinate y, value, weight]
# ```
# - **step_size**: (*float*) distance between lags within each points are included in the calculations,
# - **max_range**: (*float*) maximum range of analysis.
#
#
# OUTPUT:
#
# - (*OrderedDict*) variogram_cloud - dict with pairs {lag: list of squared differences}.
# ---
# ## ```show_variogram_cloud()```
#
# ```python
# pyinterpolate.semivariance.show_variogram_cloud(
# variogram_cloud,
# figsize=None)
# ```
#
# Function shows boxplots of variogram lags. It is especially useful when you want to check outliers in your dataset.
#
# INPUT:
#
# - **variogram_cloud**: (*OrderedDict*) lags and halved squared differences between points,
# - **figsize**: (*tuple*), default is `None`.
# ---
# ## ```calc_semivariance_from_pt_cloud()```
#
# ```python
# pyinterpolate.semivariance.calc_semivariance_from_pt_cloud(
# pt_cloud_dict)
# ```
#
# Function calculates experimental semivariogram from point cloud variogram.
#
# INPUT:
#
# - **pt_cloud_dict**: (*OrderedDict*) {lag: [values]}.
#
# OUTPUT:
#
# - (*numpy array*) [lag, semivariance, number of points].
# ---
# ## ```remove_outliers()```
#
# ```python
# pyinterpolate.semivariance.remove_outliers(data_dict,
# exclude_part='top',
# weight=1.5)
# ```
#
# Function removes outliers from the variogram point cloud for each lag and returns dict without extreme values from the top, bottom or both parts of the variogram point cloud for a given lag. Algorithm uses quartiles to remove outliers:
#
# (1)
# $$BottomOutlier < Q1 - w*(Q3-Q1)$$
#
# (2)
# $$Q3 + w*(Q3-Q1) < TopOutlier$$
#
# where:
#
# - $Q1$ - 1st quantile (25%)
# - $Q3$ - 3rd quantile (75%)
# - $w$ - weight associated with the algorithm, larger weight => less number of values treated as an outlier.
#
#
# INPUT:
#
# - **data_dict**: (*OrderedDict*) with {lag: list of values},
# - **exclude_part**: (*str*) default = `'top'`, available `'top'`, `'both'` or `'bottom'` - part of the variogram point cloud which is excluded from a given lag.
# - **weight**: (*float*) default=1.5, affects number of values which are removed.
#
# OUTPUT:
#
# - (*OrderedDict*) {lag: [variances between point pairs within a given lag]}
# ---
# ## ```TheoreticalSemivariogram```
#
# ### Class initialization
#
# ```python
# pyinterpolate.semivariance.TheoreticalSemivariogram(
# self,
# points_array=None,
# empirical_semivariance=None,
# verbose=False)
#
# ```
#
# Class calculates theoretical semivariogram.
#
# Available theoretical models:
#
# - spherical_model(distance, nugget, sill, semivar_range)
# - gaussian_model(distance, nugget, sill, semivar_range)
# - exponential_model(distance, nugget, sill, semivar_range)
# - linear_model(distance, nugget, sill, semivar_range)
# - cubic_model(distance, nugget, sill, semivar_range)
# - circular_model(distance, nugget, sill, semivar_range)
# - power_model(distance, nugget, sill, semivar_range)
#
#
# INITIALIZATION PARAMS:
#
# - **points_array**: (_numpy array_) analysed points where the last column is representing values, typically x, y, value,
# - **empirical_semivariance**: (_numpy array_) semivariance where first row of array represents lags and the second row represents semivariance's values for given lag.
#
# ### Class public methods:
#
# - **fit_semivariance**: Method fits experimental points into chosen semivariance model type,
# - **find_optimal_model**: Method fits experimental points into all available models and choose one with the lowest error,
# - **export_model**: Function exports semivariance model to the csv file,
# - **import_model**: Function imports semivariance model and updates it's parameters,
# - **export_semivariance**: Method exports theoretical semivariance and experimental semivariance to csv file,
# - **show_experimental_semivariogram**: Function shows experimental semivariogram of a given model,
# - **show_semivariogram**: Function shows experimental and theoretical semivariogram in one plot.
#
# ---
#
# ### ```TheoreticalSemivariogram.fit_semivariance()```
#
# ```python
# TheoreticalSemivariogram.fit_semivariance(
# self,
# model_type,
# number_of_ranges=16)
# ```
#
# Method fits experimental points into chosen semivariance model type.
#
# INPUT:
#
# - **model_type**: (_str_) 'exponential', 'gaussian', 'linear', 'spherical',
# - **number_of_ranges**: (_int_) deafult = 16. Used to create an array of equidistant ranges between minimal range of empirical semivariance and maximum range of empirical semivariance.
#
# OUTPUT:
#
# - (model_type, model parameters)
#
# ---
#
# ### ```TheoreticalSemivariogram.find_optimal_model()```
#
# ```python
# TheoreticalSemivariogram.find_optimal_model(
# self,
# weighted=False,
# number_of_ranges=16)
# ```
#
# Method fits experimental points into all available models and choose one with the lowest error.
#
# INPUT:
#
# - **weighted**: (_bool_) default=```False```. If ```True``` then each lag is weighted by:
#
# $$\frac{\sqrt{N(h)}}{\gamma_{experimental}(h)}$$
#
# where:
#
# $N(h)$ - number of point pairs in a given range,
#
# $\gamma_{experimental}(h)$ - value of experimental semivariogram for $h$.
# - **number_of_ranges**: (_int_) default=16. Used to create an array of equidistant ranges between minimal range of empirical semivariance and maximum range of empirical semivariance.
#
# OUTPUT:
#
# - model_type
#
# Function updates class parameters with model properties.
#
# ---
#
# ### ```TheoreticalSemivariogram.export_model()```
#
# ```python
# TheoreticalSemivariogram.export_model(self, filename)
# ```
#
# Function exports semivariance model to the csv file. Columns of csv file are: name, nugget, sill, range, model_error.
#
# ---
#
# ### ```TheoreticalSemivariogram.import_model()```
#
# ```python
# TheoreticalSemivariogram.import_model(self, filename)
# ```
#
# Function imports semivariance model and updates its parameters (model name, nugget, sill, range, model_error).
#
# ---
#
# ### ```TheoreticalSemivariogram.export_semivariance()```
#
# ```python
# TheoreticalSemivariogram.export_semivariance(self, filename)
# ```
#
# Function exports semivariance data into csv file. Exported data has three columns: `lags`, `experimental`, `theoretical` where theoretical values are calculated from the fitted model and lags given by experimental semivariogram.
#
# ---
#
# ### ```TheoreticalSemivariogram.show_experimental_semivariogram()```
#
# ```python
# TheoreticalSemivariogram.show_experimental_semivariogram(self)
# ```
#
# Function shows experimental semivariogram of a given model.
#
# ---
#
# ### ```TheoreticalSemivariogram.show_semivariogram()```
#
# ```python
# TheoreticalSemivariogram.show_semivariogram(self)
# ```
#
# Function shows experimental and theoretical semivariogram in one plot.
| docs/build/html/code_documentation/semivariance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# First create a Azure Web App from Container
# +
import requests
import json
import numpy as np
scoring_uri = "https://playgroundsklearn-mnist-svc.azurewebsites.net/score"
from utils import load_data
# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the neural network converge faster
X_test = load_data('./data/test-images.gz', False) / 255.0
y_test = load_data('./data/test-labels.gz', True).reshape(-1)
# send a random row from the test set to score
random_index = np.random.randint(0, len(X_test)-1)
input_data = "{\"data\": [" + str(list(X_test[random_index])) + "]}"
#print("input data:", input_data)
headers = {'Content-Type':'application/json'}
resp = requests.post(scoring_uri, input_data, headers=headers)
print("POST to url", scoring_uri)
print("label:", y_test[random_index])
print("prediction:", resp.text)
# -
| GetStarted/AzureML - 06_Test Model on WebApp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear models
# In this notebook I will try and implement linear regression for 2-dim data and some of it's basic implementations, such as: <br>
# -LSM<br>
# -Gradient descent<br>
# -Stohastic Gradient descent<br>
# -Gradient descent with momentum<br>
# -Nesterov Accelerated Gradient descent<br>
# -AdaGrad
# -AdaDelta
#
#
#
# @article {url = {https://ruder.io/optimizing-gradient-descent/index.html#stochasticgradientdescent}}
#
# @article {url = {https://habr.com/ru/post/318970/}}
#
# @article {url = {https://ruder.io/optimizing-gradient-descent/index.html#stochasticgradientdescent}}
# ### Step 1: generating data
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
np.random.seed(0) # fix seed for reproducibility.
mean = [-1, 1] # mean of normal distribution. Try others.
cov = [[1, -0.8],
[-0.8, 1]] # covariance of normal distribution
sample = np.random.multivariate_normal(mean, cov, size=1000) # 1000 normally distributed samples
# TODO: compare with normalized normal distribution; also with other distributions; also add outliers
# +
plt.rcParams['figure.figsize'] = [15,8]
x, y = sample[:, 0], sample[:, 1] # separate samples on feature x and target y
# Visualize dataset
plt.scatter(x, y)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
X = np.array([np.array([1, xi]) for xi in x])
X[:3], y[:3], X.shape, y.shape
# ### Some basic functions
#
# -prediction ( X, theta): $y = \theta^TX$ <br>
# -MSE score (y, $\hat{y}$)
def predict(X, theta):
return np.dot(X, theta)
def l2_score(y, y_hat):
return sum((y - y_hat) ** 2) / (2 * len(y))
# ### Simple MSE
# Note: This is an analytical algorhytm. Thought it finds the best solution, it's complexity is: O(n) = $(n^\theta) + n^{2.4} $
# +
def MSE_weights(X, y):
return np.linalg.inv(X.T @ X) @ (X.T @ y)
theta_mse = MSE_weights(X, y)
#plot
edges = np.array([x.min(), x.max()])
line = theta_mse[0] + theta_mse[1] * edges
plt.plot(edges, line, 'r-')
plt.scatter(x, y)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# #### MSE score
# This is as good as it gets
prediction = predict(X, theta_mse)
l2_score(prediction, y)
# ### Gradient descent
# The main idea behind gradient descent is that we can easily find the direction of function's fastest decrease - Jacobian matrix transforms our space so that (1, 1, ..., 1) turns into the fector pointing to the fastest increase direction. We multiply it by -1 and get decrease.
#
#
# $\theta = \theta + \alpha \nabla J(\theta)$
#
# $\nabla J(\theta) = \sum_m(y - h_\theta(X))X$
#
# Note: $ O(n) = i * n^3 $
# +
class GradientDescent:
def __init__(self, X, y, l_rate=0.001, iters=100, starting_point=np.array([0]*2)):
self.le_rate = l_rate
self.iters = iters
self.starting_point = starting_point
self.X = X
self.y = y
def fit(self):
theta = np.array(self.starting_point) # init theta with any scalar or vector
m = len(X)
history = [theta] # list of theta values on each iteration
cost = [l2_score(y, predict(X, theta))] # list of cost function values on each iteration
for i in range(self.iters):
history.append(theta)
cost.append(l2_score(y, predict(X, theta)))
theta = theta + self.le_rate * (np.sum((y - np.dot(X, theta.T)) * X.T, axis = 1)) # gradient descent itself, vectorized
self.theta, self.cost, self.history = theta, np.array(cost), np.array(history)
def predict(self):
return self.theta, np.array(self.cost), np.array(self.history)
gd = GradientDescent(X, y, 0.0001, 100, [5, 10])
gd.fit()
gd_theta, gd_cost, gd_hishory = gd.predict()
extremes = np.array([np.min(x), np.max(x)])
line = gd_theta[0] + extremes * gd_theta[1]
plt.plot(extremes, line, 'r-')
plt.scatter(x, y)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
prediction = predict(X, gd_theta)
l2_score(prediction, y)
# FOR FURTHER EXTENSIONS HERE
# ## Stohastic gradient
class StohasticGD:
def __init__(self, X, y, le_rate, starting_point, iters=100, batch_size=1):
self.X = X
self.y = y
self.le_rate = le_rate
self.iters = iters
self.starting_point = starting_point
self.batch_size = batch_size
def fit(self):
theta = np.array(self.starting_point)
history = []
cost = []
data = []
for i in range(self.iters):
history.append(theta)
cost.append(l2_score(predict(self.X, theta), self.y))
batch_indeces = np.random.choice(X.shape[0], size=self.batch_size, replace=False)
data.append((self.X[batch_indeces][:,1], self.y[batch_indeces]))
theta = theta + self.le_rate * (np.sum((y[batch_indeces] - np.dot(X[batch_indeces], theta.T))\
* X[batch_indeces].T, axis = 1))
return theta, np.array(cost), np.array(history), np.array(data)
# +
sgd = StohasticGD(X, y, 0.001, [5, 10], iters=10000, batch_size=5)
sgd_theta, sgd_cost, sgd_hishory, sgd_data = sgd.fit()
print(sgd_theta)
prediction = predict(X, sgd_theta)
score = l2_score(prediction, y)
print(score)
extremes = np.array([np.min(x), np.max(x)])
line = sgd_theta[0] + extremes * sgd_theta[1]
plt.plot(extremes, line, 'r-')
plt.scatter(x, y)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# ## Nesterov's gradient
# $\theta_i = \theta_i - v_t $ <br>
# $ v_t = \gamma v_{t-1} + \eta \bigtriangledown_\theta J(\theta)$ ,where $\eta = 1 - \gamma$
| Linear_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/python3
import urllib.request
from bs4 import BeautifulSoup
website="http://php.net"
#data downloading
web_data=urllib.request.urlopen(website)
#printing data
#print (web_data.read())
#reading web data with html tags
clean_data=web_data.read()
#applying lib of html5 to scrap
get_clean=BeautifulSoup(clean_data, 'html5lib')
#getting only text formt data
final_data=get_clean.get_text()
#removing un-necessary space
good_data=final_data.strip()
#print good-data
new_data=[]
for i in good_data:
j=i.split()
new_data.append(j)
print(new_data)
# -
| data-collect_tokenize_trim-eg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from sklearn.inspection import permutation_importance
import time
from catboost import CatBoostClassifier, CatBoostRegressor, Pool, cv
import pickle
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import shap
# -
os.getcwd()
shap.initjs()
# +
random_seed = 123
print("start")
df_eco = pd.read_csv('../../tables/kgmicrobe_table.tsv', sep='\t', encoding='utf-8')#index_col=1,
print(df_eco.head())
# -
df_eco.shape
subjects = df_eco.loc[:,"subject"]
df_eco_small = df_eco
df_eco_small.drop('subject', axis=1, inplace=True)
df_eco_small = df_eco.fillna(0)
print(df_eco_small.describe())
df_eco_small = df_eco_small.loc[~(df_eco_small==0).all(axis=1)]
df_eco_small.shape
# +
#value.Shape:bacillus
y = df_eco_small['value.Shape:bacillus']
print(y)
print("df_eco "+str(df_eco_small.shape))
X = df_eco_small.drop('value.Shape:bacillus', 1)#df_eco.iloc[:,:-1]
# -
df_eco_small[df_eco_small.isin(['NCBITaxon:1095661']).any(1)]
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=random_seed) #, random_state=9# The seed was 'chosen' so test and training contain all labels: rn=3,4,8,9
print("train label deficit:",len(set(y)-set(y_train)),"test label deficit:",len(set(y)-set(y_test)))
print("shapes "+str(X_train.shape)+"\t"+str(X_test.shape)+"\t"+str(y_train.shape)+"\t"+str(y_test.shape))
train_dataset = Pool(X_train, y_train)
test_dataset = Pool(X_test, y_test)
input_data_dump = [X, y, X_train, X_test, y_train, y_test]
pickle.dump(input_data_dump,open("input_data_dump", "wb" ) )
# +
modelstart = time.time()
print(f"Starting search at {modelstart}")
cb_model = CatBoostRegressor(loss_function='MAE',
iterations = 200,
verbose = 5,
learning_rate = 0.1,
depth = 3,
l2_leaf_reg = 0.5,
#eval_metric = 'MCC',
random_seed = random_seed,
#bagging_temperature = 0.2,
#od_type = 'Iter',
#od_wait = 100
)
grid = {#'iterations': [100, 150, 200],
'learning_rate': [0.6, 0.8, 1],
'depth': [3,4,5],
'l2_leaf_reg': [2,3,4]}
grid_search_result = cb_model.grid_search(grid, train_dataset)
lr = grid_search_result['params']['learning_rate']
de = grid_search_result['params']['depth']
l2 = grid_search_result['params']['l2_leaf_reg']
print(f"Trained grid search in {time.time() - modelstart}s")
print("lr, de, l2 "+str(lr)+", "+str(de)+", "+str(l2))
# +
modelstart = time.time()
print(f"Starting at {modelstart}")
cb_model = CatBoostRegressor(loss_function='MAE',
iterations = 200,
verbose = 5,
learning_rate = lr,
depth = de,
l2_leaf_reg = l2,
#eval_metric = 'MCC',
random_seed = random_seed,
#bagging_temperature = 0.2,
#od_type = 'Iter',
#od_wait = 100
)
cbmf = cb_model.fit(X_train, y_train)
print(f"Trained in {time.time() - modelstart}s")
# -
pred_train = cb_model.predict(X_train)
rmseT = (np.sqrt(mean_squared_error(y_train, pred_train)))
r2T = r2_score(y_train, pred_train)
print("Testing performance:")
print('RMSE training: {:.2f}'.format(rmseT))
print('R2 training: {:.2f}'.format(r2T))
cbmf.feature_names = X.columns
pred_test = cb_model.predict(X_test)
rmse = (np.sqrt(mean_squared_error(y_test, pred_test)))
r2 = r2_score(y_test, pred_test)
print("Testing performance:")
print('RMSE: {:.2f}'.format(rmse))
print('R2: {:.2f}'.format(r2))
explainer_model = shap.TreeExplainer(cb_model)
explainer_fit = shap.TreeExplainer(cbmf)
data_output = [random_seed, cb_model, cbmf, pred_train, explainer_model, pred_test, explainer_fit]
pickle.dump(data_output,open("data_output", "wb" ) )
sorted_feature_importance = cb_model.feature_importances_.argsort()
plt.figure(figsize=(20,10))
plt.barh(cb_model.feature_names[sorted_feature_importance][-50:],
cb_model.feature_importances_[sorted_feature_importance][-50:],
color='turquoise')
plt.xlabel("CatBoost Feature Importance")
shap_values = explainer_model.shap_values(X_train)
shap.summary_plot(shap_values, X_train, feature_names = cb_model.feature_names[sorted_feature_importance],max_display=X.shape[1])#,matplotlib=True).savefig('SHAP.pdf',bbox_inches = 'tight')
shap_values = explainer_fit.shap_values(X_train)
shap.force_plot(explainer_fit.expected_value, shap_values[0:20,:], X_train.iloc[0:20,:])
cbmf_all = cb_model.fit(X,y)
pred_all = cbmf_all.predict(X)
rmseA = (np.sqrt(mean_squared_error(y, pred_all)))
r2A = r2_score(y, pred_all)
print("All performance:")
print('RMSE training: {:.2f}'.format(rmseA))
print('R2 training: {:.2f}'.format(r2A))
# +
explainer_all = shap.TreeExplainer(cbmf_all)
# -
shap_values_all = explainer_all.shap_values(X)
shap.summary_plot(shap_values_all, X, feature_names = cb_model.feature_names[sorted_feature_importance],max_display=X.shape[1])#,matplotlib=True).savefig('SHAP.pdf',bbox_inches = 'tight')
_feature_importance = cbmf_all.feature_importances_.argsort()
plt.figure(figsize=(20,10))
plt.barh(cb_model.feature_names[sorted_feature_importance][-50:],
cb_model.feature_importances_[sorted_feature_importance][-50:],
color='turquoise')
plt.xlabel("CatBoost Feature Importance")
| notebooks/classifiers/catboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # From tokens to numbers: the document-term matrix
# -
# The bag of words model represents a document based on the frequency of the terms or tokens it contains. Each document becomes a vector with one entry for each token in the vocabulary that reflects the token’s relevance to the document.
#
# The document-term matrix is straightforward to compute given the vocabulary. However, it is also a crude simplification because it abstracts from word order and grammatical relationships. Nonetheless, it often achieves good results in text classification quickly and, thus, a very useful starting point.
#
# There are several ways to weigh a token’s vector entry to capture its relevance to the document. We will illustrate below how to use sklearn to use binary flags that indicate presence or absence, counts, and weighted counts that account for differences in term frequencies across all documents, i.e., in the corpus.
# ## Imports & Settings
import warnings
warnings.filterwarnings('ignore')
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
from collections import Counter
from pathlib import Path
import numpy as np
import pandas as pd
from scipy import sparse
from scipy.spatial.distance import pdist
# Visualization
import matplotlib.pyplot as plt
from matplotlib.ticker import ScalarFormatter
import seaborn as sns
from ipywidgets import interact, FloatRangeSlider
# spacy for language processing
import spacy
# sklearn for feature extraction & modeling
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer, TfidfTransformer
from sklearn.model_selection import train_test_split
# -
sns.set_style('white')
# + [markdown] slideshow={"slide_type": "skip"}
# ## Load BBC data
# + slideshow={"slide_type": "skip"}
path = Path('..', 'data', 'bbc')
files = sorted(list(path.glob('**/*.txt')))
doc_list = []
for i, file in enumerate(files):
topic = file.parts[-2]
article = file.read_text(encoding='latin1').split('\n')
heading = article[0].strip()
body = ' '.join([l.strip() for l in article[1:]]).strip()
doc_list.append([topic, heading, body])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Convert to DataFrame
# + slideshow={"slide_type": "skip"}
docs = pd.DataFrame(doc_list, columns=['topic', 'heading', 'body'])
docs.info()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inspect results
# + slideshow={"slide_type": "fragment"}
docs.sample(10)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Data drawn from 5 different categories
# + slideshow={"slide_type": "fragment"}
docs.topic.value_counts(normalize=True).to_frame('count').style.format({'count': '{:,.2%}'.format})
# -
# ## Explore Corpus
# ### Token Count via Counter()
# + slideshow={"slide_type": "fragment"}
# word count
word_count = docs.body.str.split().str.len().sum()
print(f'Total word count: {word_count:,d} | per article: {word_count/len(docs):,.0f}')
# -
token_count = Counter()
for i, doc in enumerate(docs.body.tolist(), 1):
if i % 500 == 0:
print(i, end=' ', flush=True)
token_count.update([t.strip() for t in doc.split()])
tokens = (pd.DataFrame(token_count.most_common(), columns=['token', 'count'])
.set_index('token')
.squeeze())
n = 50
(tokens
.iloc[:50]
.plot
.bar(figsize=(14, 4), title=f'Most frequent {n} of {len(tokens):,d} tokens'))
sns.despine()
plt.tight_layout();
# + [markdown] slideshow={"slide_type": "slide"}
# ## Document-Term Matrix with `CountVectorizer`
# -
# The scikit-learn preprocessing module offers two tools to create a document-term matrix. The [CountVectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) uses binary or absolute counts to measure the term frequency tf(d, t) for each document d and token t.
#
# The [TfIDFVectorizer](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html), in contrast, weighs the (absolute) term frequency by the inverse document frequency (idf). As a result, a term that appears in more documents will receive a lower weight than a token with the same frequency for a given document but lower frequency across all documents.
#
# The resulting tf-idf vectors for each document are normalized with respect to their absolute or squared totals (see the sklearn documentation for details). The tf-idf measure was originally used in information retrieval to rank search engine results and has subsequently proven useful for text classification or clustering.
# Both tools use the same interface and perform tokenization and further optional preprocessing of a list of documents before vectorizing the text by generating token counts to populate the document-term matrix.
#
# Key parameters that affect the size of the vocabulary include:
#
# - `stop_words`: use a built-in or provide a list of (frequent) words to exclude
# - `ngram_range`: include n-grams in a range for n defined by a tuple of (nmin, nmax)
# - `lowercase`: convert characters accordingly (default is True)
# - `min_df `/ max_df: ignore words that appear in less / more (int) or a smaller / larger share of documents (if float [0.0,1.0])
# - `max_features`: limit number of tokens in vocabulary accordingly
# - `binary`: set non-zero counts to 1 True
# ### Key parameters
# + slideshow={"slide_type": "fragment"}
print(CountVectorizer().__doc__)
# -
# ### Document Frequency Distribution
# +
binary_vectorizer = CountVectorizer(max_df=1.0,
min_df=1,
binary=True)
binary_dtm = binary_vectorizer.fit_transform(docs.body)
# -
binary_dtm
n_docs, n_tokens = binary_dtm.shape
tokens_dtm = binary_vectorizer.get_feature_names()
# #### CountVectorizer skips certain tokens by default
tokens.index.difference(pd.Index(tokens_dtm))
# #### Persist Result
results_path = Path('results', 'bbc')
if not results_path.exists():
results_path.mkdir(parents=True)
dtm_path = results_path / 'binary_dtm.npz'
if not dtm_path.exists():
sparse.save_npz(dtm_path, binary_dtm)
token_path = results_path / 'tokens.csv'
if not token_path.exists():
pd.Series(tokens_dtm).to_csv(token_path, index=False)
else:
tokens = pd.read_csv(token_path, header=None, squeeze=True)
doc_freq = pd.Series(np.array(binary_dtm.sum(axis=0)).squeeze()).div(n_docs)
max_unique_tokens = np.array(binary_dtm.sum(axis=1)).squeeze().max()
# ### `min_df` vs `max_df`: Interactive Visualization
# The notebook contains an interactive visualization that explores the impact of the min_df and max_df settings on the size of the vocabulary. We read the articles into a DataFrame, set the CountVectorizer to produce binary flags and use all tokens, and call its .fit_transform() method to produce a document-term matrix:
# The visualization shows that requiring tokens to appear in at least 1% and less than 50% of documents restricts the vocabulary to around 10% of the almost 30K tokens.
# This leaves a mode of slightly over 100 unique tokens per document (left panel), and the right panel shows the document frequency histogram for the remaining tokens.
# +
df_range = FloatRangeSlider(value=[0.0, 1.0],
min=0,
max=1,
step=0.0001,
description='Doc. Freq.',
disabled=False,
continuous_update=True,
orientation='horizontal',
readout=True,
readout_format='.1%',
layout={'width': '800px'})
@interact(df_range=df_range)
def document_frequency_simulator(df_range):
min_df, max_df = df_range
keep = doc_freq.between(left=min_df, right=max_df)
left = keep.sum()
fig, axes = plt.subplots(ncols=2, figsize=(14, 6))
updated_dtm = binary_dtm.tocsc()[:, np.flatnonzero(keep)]
unique_tokens_per_doc = np.array(updated_dtm.sum(axis=1)).squeeze()
sns.distplot(unique_tokens_per_doc, ax=axes[0], kde=False, norm_hist=False)
axes[0].set_title('Unique Tokens per Doc')
axes[0].set_yscale('log')
axes[0].set_xlabel('# Unique Tokens')
axes[0].set_ylabel('# Documents (log scale)')
axes[0].set_xlim(0, max_unique_tokens)
axes[0].yaxis.set_major_formatter(ScalarFormatter())
term_freq = pd.Series(np.array(updated_dtm.sum(axis=0)).squeeze())
sns.distplot(term_freq, ax=axes[1], kde=False, norm_hist=False)
axes[1].set_title('Document Frequency')
axes[1].set_ylabel('# Tokens')
axes[1].set_xlabel('# Documents')
axes[1].set_yscale('log')
axes[1].set_xlim(0, n_docs)
axes[1].yaxis.set_major_formatter(ScalarFormatter())
title = f'Document/Term Frequency Distribution | # Tokens: {left:,d} ({left/n_tokens:.2%})'
fig.suptitle(title, fontsize=14)
sns.despine()
fig.tight_layout()
fig.subplots_adjust(top=.9)
# -
# ### Most similar documents
# The CountVectorizer result lets us find the most similar documents using the `pdist()` function for pairwise distances provided by the `scipy.spatial.distance` module.
#
# It returns a condensed distance matrix with entries corresponding to the upper triangle of a square matrix.
#
# We use `np.triu_indices()` to translate the index that minimizes the distance to the row and column indices that in turn correspond to the closest token vectors.
m = binary_dtm.todense()
pairwise_distances = pdist(m, metric='cosine')
closest = np.argmin(pairwise_distances)
rows, cols = np.triu_indices(n_docs)
rows[closest], cols[closest]
docs.iloc[6].to_frame(6).join(docs.iloc[245].to_frame(245)).to_csv(results_path / 'most_similar.csv')
docs.iloc[6]
pd.DataFrame(binary_dtm[[6, 245], :].todense()).sum(0).value_counts()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Baseline document-term matrix
# + slideshow={"slide_type": "fragment"}
# Baseline: number of unique tokens
vectorizer = CountVectorizer() # default: binary=False
doc_term_matrix = vectorizer.fit_transform(docs.body)
doc_term_matrix
# + slideshow={"slide_type": "fragment"}
doc_term_matrix.shape
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inspect tokens
# + slideshow={"slide_type": "fragment"}
# vectorizer keeps words
words = vectorizer.get_feature_names()
words[:10]
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inspect doc-term matrix
# + slideshow={"slide_type": "fragment"}
# from scipy compressed sparse row matrix to sparse DataFrame
doc_term_matrix_df = pd.DataFrame.sparse.from_spmatrix(doc_term_matrix, columns=words)
doc_term_matrix_df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Most frequent terms
# + slideshow={"slide_type": "fragment"}
word_freq = doc_term_matrix_df.sum(axis=0).astype(int)
word_freq.sort_values(ascending=False).head()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Compute relative term frequency
# + slideshow={"slide_type": "fragment"}
vectorizer = CountVectorizer(binary=True)
doc_term_matrix = vectorizer.fit_transform(docs.body)
doc_term_matrix.shape
# + slideshow={"slide_type": "slide"}
words = vectorizer.get_feature_names()
word_freq = doc_term_matrix.sum(axis=0)
# reduce to 1D array
word_freq_1d = np.squeeze(np.asarray(word_freq))
pd.Series(word_freq_1d, index=words).div(
docs.shape[0]).sort_values(ascending=False).head(10)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Visualize Doc-Term Matrix
# + slideshow={"slide_type": "fragment"}
sns.heatmap(pd.DataFrame(doc_term_matrix.todense(), columns=words), cmap='Blues')
plt.gcf().set_size_inches(14, 8);
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using thresholds to reduce the number of tokens
# + slideshow={"slide_type": "fragment"}
vectorizer = CountVectorizer(max_df=.2, min_df=3, stop_words='english')
doc_term_matrix = vectorizer.fit_transform(docs.body)
doc_term_matrix.shape
# + [markdown] slideshow={"slide_type": "slide"}
# ### Use CountVectorizer with Lemmatization
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Building a custom `tokenizer` for Lemmatization with `spacy`
# + slideshow={"slide_type": "fragment"}
nlp = spacy.load('en')
def tokenizer(doc):
return [w.lemma_ for w in nlp(doc)
if not w.is_punct | w.is_space]
# + run_control={"marked": false} slideshow={"slide_type": "slide"}
vectorizer = CountVectorizer(tokenizer=tokenizer, binary=True)
doc_term_matrix = vectorizer.fit_transform(docs.body)
doc_term_matrix.shape
# + slideshow={"slide_type": "fragment"}
lemmatized_words = vectorizer.get_feature_names()
word_freq = doc_term_matrix.sum(axis=0)
word_freq_1d = np.squeeze(np.asarray(word_freq))
word_freq_1d = pd.Series(word_freq_1d, index=lemmatized_words).div(docs.shape[0])
word_freq_1d.sort_values().tail(20)
# + [markdown] slideshow={"slide_type": "fragment"}
# Unlike verbs and common nouns, there's no clear base form of a personal pronoun. Should the lemma of "me" be "I", or should we normalize person as well, giving "it" — or maybe "he"? spaCy's solution is to introduce a novel symbol, -PRON-, which is used as the lemma for all personal pronouns.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Document-Term Matrix with `TfIDFVectorizer`
# -
# The TfIDFTransfomer computes the tf-idf weights from a document-term matrix of token counts like the one produced by the CountVectorizer.
#
# The TfIDFVectorizer performs both computations in a single step. It adds a few parameters to the CountVectorizer API that controls the smoothing behavior.
# ### Key Parameters
# The `TfIDFTransformer` builds on the `CountVectorizer` output; the `TfIDFVectorizer` integrates both
# + slideshow={"slide_type": "fragment"}
print(TfidfTransformer().__doc__)
# -
# ### How Term Frequency - Inverse Document Frequency works
# The TFIDF computation works as follows for a small text sample
sample_docs = ['call you tomorrow',
'Call me a taxi',
'please call me... PLEASE!']
# #### Compute term frequency
vectorizer = CountVectorizer()
tf_dtm = vectorizer.fit_transform(sample_docs).todense()
tokens = vectorizer.get_feature_names()
term_frequency = pd.DataFrame(data=tf_dtm,
columns=tokens)
print(term_frequency)
# #### Compute document frequency
vectorizer = CountVectorizer(binary=True)
df_dtm = vectorizer.fit_transform(sample_docs).todense().sum(axis=0)
document_frequency = pd.DataFrame(data=df_dtm,
columns=tokens)
print(document_frequency)
# #### Compute TfIDF
tfidf = pd.DataFrame(data=tf_dtm/df_dtm, columns=tokens)
print(tfidf)
# #### The effect of smoothing
# The TfidfVectorizer uses smoothing for document and term frequencies:
# - `smooth_idf`: add one to document frequency, as if an extra document contained every token in the vocabulary
# once to prevents zero divisions
# - `sublinear_tf`: scale term Apply sublinear tf scaling, i.e. replace tf with 1 + log(tf)
vect = TfidfVectorizer(smooth_idf=True,
norm='l2', # squared weights sum to 1 by document
sublinear_tf=False, # if True, use 1+log(tf)
binary=False)
print(pd.DataFrame(vect.fit_transform(sample_docs).todense(),
columns=vect.get_feature_names()))
# ### TfIDF with new articles
# Due to their ability to assign meaningful token weights, TFIDF vectors are also used to summarize text data. E.g., reddit's autotldr function is based on a similar algorithm.
# + slideshow={"slide_type": "slide"}
tfidf = TfidfVectorizer(stop_words='english')
dtm_tfidf = tfidf.fit_transform(docs.body)
tokens = tfidf.get_feature_names()
dtm_tfidf.shape
# + slideshow={"slide_type": "fragment"}
token_freq = (pd.DataFrame({'tfidf': dtm_tfidf.sum(axis=0).A1,
'token': tokens})
.sort_values('tfidf', ascending=False))
# -
token_freq.head(10).append(token_freq.tail(10)).set_index('token')
# ### Summarizing news articles using TfIDF weights
# #### Select random article
article = docs.sample(1).squeeze()
article_id = article.name
print(f'Topic:\t{article.topic.capitalize()}\n\n{article.heading}\n')
print(article.body.strip())
# #### Select most relevant tokens by tfidf value
article_tfidf = dtm_tfidf[article_id].todense().A1
article_tokens = pd.Series(article_tfidf, index=tokens)
article_tokens.sort_values(ascending=False).head(10)
# #### Compare to random selection
pd.Series(article.body.split()).sample(10).tolist()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Create Train & Test Sets
# -
# ### Stratified `train_test_split`
# + slideshow={"slide_type": "fragment"}
train_docs, test_docs = train_test_split(docs,
stratify=docs.topic,
test_size=50,
random_state=42)
# + slideshow={"slide_type": "fragment"}
train_docs.shape, test_docs.shape
# + slideshow={"slide_type": "fragment"}
pd.Series(test_docs.topic).value_counts()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Vectorize train & test sets
# + slideshow={"slide_type": "fragment"}
vectorizer = CountVectorizer(max_df=.2,
min_df=3,
stop_words='english',
max_features=2000)
train_dtm = vectorizer.fit_transform(train_docs.body)
words = vectorizer.get_feature_names()
train_dtm
# + slideshow={"slide_type": "fragment"}
test_dtm = vectorizer.transform(test_docs.body)
test_dtm
| 14_working_with_text_data/03_document_term_matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# File name: Investigation_1.ipynb
# Author: <NAME>
# Latest Update: 16.07.2019
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import math
from sklearn.cluster import KMeans
from bokeh.plotting import figure, show, output_notebook, gridplot
from bokeh.tile_providers import CARTODBPOSITRON
# # 0) Task
# **Questions:**
#
# The businesses stored inside the YELP dataset are located widely across North America. In which regions are the most businesses? Does the geographical location have influence on the ration behavior of the users?
#
# **Approach:**
#
# Step 1: Cluster the businesses into different regions. <br>
# Step 2: Investigate via descriptive analysis possible differences between the regions.
# # 1) Preprocessing
# ## 1.1) Load the data
path_dataset_business = '../Data/yelp_dataset/business.json'
df = pd.read_json(path_dataset_business, lines=True)
df.head()
# ## 1.2) EDA (exploratory data analysis)
df.describe()
# Distribution of the star ratings
star_count = df['stars'].value_counts()
star_count.reindex([1, 1.5, 2, 2.5, 3, 3.5, 4, 4.5, 5]).plot.bar()
# +
# Number of businesses
len_business = len(df['business_id'])
# Number of cities
len_unique_cities = len(df['city'].unique())
# Number of postal codes
len_unique_postal_code = len(df['postal_code'].unique())
print('Number of businesses: ', len_business)
print('Number of cities: ', len_unique_cities)
print('Number of postal codes: ', len_unique_postal_code)
# -
# ## 1.3) Visualize the locations
def transform_mercator(Coords):
""" Transformation of longitude and latitude coordinates into the mercator format.
Parameters:
Coords: List ([latitute, logitute]) with the coordinates of one point
Return:
(x,y): Transformation of Coords
"""
lat = Coords[0]
lon = Coords[1]
r_major = 6378137.000
x = r_major * math.radians(lon)
scale = x/lon
y = 180.0/math.pi * math.log(math.tan(math.pi/4.0 + lat * (math.pi/180.0)/2.0)) * scale
return (x, y)
def transform_mercator_dataframe(Coords):
""" Transformation of longitude and latitude coordinates into the mercator format.
Parameters:
Coords: DataFrame with the columns 'latitute' and 'logitute'
Return:
(x,y): Transformation of Coords
"""
for i in range(len(Coords['latitude'])):
lat = Coords['latitude']
lon = Coords['longitude']
r_major = 6378137.000
x = r_major * lon.apply(lambda x: math.radians(x))
scale = x/lon
y = 180.0/math.pi * lat.apply(lambda x: math.log(math.tan(math.pi/4.0 + x * (math.pi/180.0)/2.0))) * scale
return (x, y)
# Boundaries of the US (130°W - 60°W, 30°N - 55°N)
x_b1, y_b1= transform_mercator([30, -130])
x_b2, y_b2= transform_mercator([55, -60])
# +
# Plot the business locations
p = figure(x_range=(x_b1, x_b2), y_range=(y_b1, y_b2), x_axis_type="mercator", y_axis_type="mercator")
p.sizing_mode = 'scale_width'
p.plot_height = 400
# Load businesses as circles in the map
# Info: Not all businesses are loaded into the table
x_p, y_p = transform_mercator_dataframe(df.sample(n=1000, random_state=42))
p.circle(x = x_p, y = y_p)
p.add_tile(CARTODBPOSITRON)
p.title.text = "Locations of businesses (representative sample)"
output_notebook()
show(p)
# -
# The visualization of the data shows, that the businesses are located in a just few areas. They are not wildly distributed across the America.
# # 2) Clustering
# Task: Cluster the businesses. Businesses in the same area should be grouped together.
# The number of clusters k -- which should be build up -- has to be defined.
# Set up a range for k, to determine the best parameter:
k_min = 2
k_max = 20
# +
score_kmeans = []
# Build up the different clusters with k in [k_min, k_max]
for i in range(k_min, k_max+1):
print('Build up Kmeans for k =', i)
kmeans = KMeans(n_clusters=i, random_state=42).fit(df[['latitude', 'longitude']])
score_kmeans.append(kmeans.inertia_)
# +
# Plot the kmeans error for different k
p = figure(plot_width=400, plot_height=400)
p.plot_width=800
p.plot_height=400
p.line(range(k_min, k_max), score_kmeans, line_width=2)
p.title.text = "Kmeans Error"
p.xaxis.axis_label = 'New xlabel'
p.yaxis.axis_label = 'Error'
show(p)
# -
# The upper plot visualizes the clustering error dependent on the different k values.
#
# In analogy of the 'elbow method' the best Cluster is gained with k = 8. With a higher value k the error will decrease further - but only very slowly. Simultaneously the efficiency will decrease.
# # 3) Deskriptiv Snalysis
# Build up clusters with k = 8
k = 8
kmeans_final = KMeans(n_clusters=k, random_state=0).fit(df[['latitude', 'longitude']])
cluster_center = kmeans_final.cluster_centers_
# +
# Transform the cluster center into mercator format
cc_x = []
cc_y = []
for i in range(k):
x, y = transform_mercator(cluster_center[i])
cc_x.append(x)
cc_y.append(y)
# +
# Plot the business locations
p = figure(x_range=(x_b1, x_b2), y_range=(y_b1, y_b2), x_axis_type="mercator", y_axis_type="mercator")
p.sizing_mode = 'scale_width'
p.plot_height = 400
# Load cluster center as circles in the map
colors = ['red', 'navy', 'gray', 'brown', 'cyan', 'lime', 'orange', 'darkgreen']
for i in range(len(cc_x)):
p.circle(x = cc_x[i], y = cc_y[i], size=50, color=colors[i], alpha=0.7, legend="Cluster "+str(i))
x_p, y_p = transform_mercator_dataframe(df.sample(n=500, random_state=42))
p.circle(x = x_p, y = y_p, color="black")
p.add_tile(CARTODBPOSITRON)
p.title.text = "Locations of businesses (representative sample)"
output_notebook()
show(p)
# -
# The map locates the business with the color black. The big, colored circle represent the clusters.
# Create a dictionary to describe the area of each cluster:
cluster_dict = {0:'Montreal' ,
1:'Las Vegas' ,
2:'Pittsburg_Cleaveland',
3:'Calgary',
4:'Phoenix',
5:'Charlotte',
6:'Toronto',
7:'Madision_Urbana'}
# Add the cluster as a new feature to the dataframe df
df['cluster'] = kmeans_final.predict(df[['latitude', 'longitude']])
df['cluster_name'] = [cluster_dict[x] for x in df['cluster']]
# Numer of businesses per cluster
cluster_grouped = df[['business_id', 'cluster_name', 'cluster']].groupby(['cluster_name', 'cluster'])
cluster_grouped.count().sort_values(by =['business_id'], ascending=False)
# The right column of the upper column shows the number of businesses in each cluster. It can be detected, that there are big differences. Example given: Cluster 4 has 8 times more businesses than cluster 7.
# +
# Calculate the number and the distribution of star ratings
star_count_cluster_0 = df[df['cluster'] == 0]['stars'].value_counts()
star_count_cluster_1 = df[df['cluster'] == 1]['stars'].value_counts()
star_count_cluster_2 = df[df['cluster'] == 2]['stars'].value_counts()
star_count_cluster_3 = df[df['cluster'] == 3]['stars'].value_counts()
star_count_cluster_4 = df[df['cluster'] == 4]['stars'].value_counts()
star_count_cluster_5 = df[df['cluster'] == 5]['stars'].value_counts()
star_count_cluster_6 = df[df['cluster'] == 6]['stars'].value_counts()
star_count_cluster_7 = df[df['cluster'] == 7]['stars'].value_counts()
star_count_cluster_0 = star_count_cluster_0.reset_index()
star_count_cluster_0.columns = ['stars', 'count']
star_count_cluster_0['stars'] = star_count_cluster_0['stars'].astype(float)
star_count_cluster_1 = star_count_cluster_1.reset_index()
star_count_cluster_1.columns = ['stars', 'count']
star_count_cluster_1['stars'] = star_count_cluster_1['stars'].astype(float)
star_count_cluster_2 = star_count_cluster_2.reset_index()
star_count_cluster_2.columns = ['stars', 'count']
star_count_cluster_2['stars'] = star_count_cluster_2['stars'].astype(float)
star_count_cluster_3 = star_count_cluster_3.reset_index()
star_count_cluster_3.columns = ['stars', 'count']
star_count_cluster_3['stars'] = star_count_cluster_3['stars'].astype(float)
star_count_cluster_4 = star_count_cluster_4.reset_index()
star_count_cluster_4.columns = ['stars', 'count']
star_count_cluster_4['stars'] = star_count_cluster_4['stars'].astype(float)
star_count_cluster_5 = star_count_cluster_5.reset_index()
star_count_cluster_5.columns = ['stars', 'count']
star_count_cluster_5['stars'] = star_count_cluster_5['stars'].astype(float)
star_count_cluster_6 = star_count_cluster_6.reset_index()
star_count_cluster_6.columns = ['stars', 'count']
star_count_cluster_6['stars'] = star_count_cluster_6['stars'].astype(float)
star_count_cluster_7 = star_count_cluster_7.reset_index()
star_count_cluster_7.columns = ['stars', 'count']
star_count_cluster_7['stars'] = star_count_cluster_7['stars'].astype(float)
# +
# Plot the distribution of star ratings for the 8 clusters
WIDTH=250
HEIGHT=250
p0 = figure(title="Cluster 0", width=WIDTH, plot_height=HEIGHT)
p0.vbar(x=star_count_cluster_0['stars'], top=star_count_cluster_0['count'], width=0.3)
p0.xaxis.axis_label = 'Stars'
p0.yaxis.axis_label = 'Count'
p1 = figure(title="Cluster 1", width=WIDTH, plot_height=HEIGHT)
p1.vbar(x=star_count_cluster_1['stars'], top=star_count_cluster_1['count'], width=0.3)
p1.xaxis.axis_label = 'Stars'
p1.yaxis.axis_label = 'Count'
p2 = figure(title="Cluster 2", width=WIDTH, plot_height=HEIGHT)
p2.vbar(x=star_count_cluster_2['stars'], top=star_count_cluster_2['count'], width=0.3)
p2.xaxis.axis_label = 'Stars'
p2.yaxis.axis_label = 'Count'
p3 = figure(title="Cluster 3", width=WIDTH, plot_height=HEIGHT)
p3.vbar(x=star_count_cluster_3['stars'], top=star_count_cluster_3['count'], width=0.3)
p3.xaxis.axis_label = 'Stars'
p3.yaxis.axis_label = 'Count'
p4 = figure(title="Cluster 4", width=WIDTH, plot_height=HEIGHT)
p4.vbar(x=star_count_cluster_4['stars'], top=star_count_cluster_4['count'], width=0.3)
p4.xaxis.axis_label = 'Stars'
p4.yaxis.axis_label = 'Count'
p5 = figure(title="Cluster 5", width=WIDTH, plot_height=HEIGHT)
p5.vbar(x=star_count_cluster_5['stars'], top=star_count_cluster_5['count'], width=0.3)
p5.xaxis.axis_label = 'Stars'
p5.yaxis.axis_label = 'Count'
p6 = figure(title="Cluster 6", width=WIDTH, plot_height=HEIGHT)
p6.vbar(x=star_count_cluster_6['stars'], top=star_count_cluster_6['count'], width=0.3)
p6.xaxis.axis_label = 'Stars'
p6.yaxis.axis_label = 'Count'
p7 = figure(title="Cluster 7", width=WIDTH, plot_height=HEIGHT)
p7.vbar(x=star_count_cluster_7['stars'], top=star_count_cluster_7['count'], width=0.3)
p7.xaxis.axis_label = 'Stars'
p7.yaxis.axis_label = 'Count'
p01 = gridplot([[p0, p1]])
p23 = gridplot([[p2, p3]])
p45 = gridplot([[p4, p5]])
p67 = gridplot([[p6, p7]])
show(p01)
show(p23)
show(p45)
show(p67)
# -
# Regional differences and similarities between the ratings and the businesses can be recognized.
#
# Example given:
# * The largest number of businesses in Cluster 1 and Cluster 4 are rated with 5 stars.
# * The group of 5 star rated businesses in Cluster 0 and Cluster 6 are relatively low.
# * In all clusters the 1 and 1.5 star ratings are approximately equal. Furthermore those ratings are comparatively low in contrast to the other ratings.
# # 4) Outlook
# The upper investigation can be extended for example with the following questions:
# * Are businesses (and therefor the categories) changing with the region? (Concrete Example: Has cluster 1 more Sushi Bars and have therefor the Sushi Bars a percentage higher rating?
# * Are businesses rated differently in different regions? (Concrete Example: Is there a correlation between the business attributes and the regions?)
# * What features should a business have, to be successful in a region?
# * ...
| Notebooks/Investigation_1.ipynb |
# ## Tutorial 0: Open a Jupyter notebook from a remote server.
#
# #### Step 1: On your local machine(e.g. Desktop) ssh into the cluster using: "ssh -L localhost:8888:localhost:8889 'remote server address'"
#
# * The port number: 8888 or 8889 may be changed if they are used by others.
# * Once you login, make sure you have activated a conda environment and you are at the folder where the .ipynb file lies.
#
# #### Step 2: On the remote server (your cluster), Run command "jupyter notebook --no-browser --port=8889". This "8889" should match with the localhost:8889.
#
# * You will see many output messages came out. Make sure you locate the token number (like <PASSWORD>) buried inside the message
#
# #### Step 3: On your local machine, Open the url: http://localhost:8888 from your web browser. The port number "8888" should match with the localhost:8888 upon login
#
# * You will be prompted to give a token number for loging in. Copy and paste the token number to log in.
#
# #### Step 4: Once you give a correct token number, you should be able open the Jupyter notebook in your local browser.
#
| tutorial/How_to_launch_jupyter_notebook/How_to.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Train neural network to predict significant wave height from SAR spectra.
# Train with heteroskedastic regression uncertainty estimates.
# Author: <NAME>, Dec 2020
import os, sys
os.environ['TF_FORCE_GPU_ALLOW_GROWTH'] = 'true' # Needed to avoid cudnn bug.
import numpy as np
import h5py
import tensorflow as tf
from tensorflow.keras.utils import Sequence, plot_model
from tensorflow.keras.callbacks import *
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers import *
from tensorflow.keras.models import Model
sys.path = ['../'] + sys.path
from sarhs.generator import SARGenerator
from sarhs.heteroskedastic import Gaussian_NLL, Gaussian_MSE
# -
def define_model():
# Low-level features.
inputs = Input(shape=(72, 60, 2))
x = Conv2D(64, (3, 3), activation='relu')(inputs)
x = MaxPooling2D(pool_size=(2, 2))(x)
x = Conv2D(128, (3, 3), activation='relu')(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
x = Conv2D(256, (3, 3), activation='relu')(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
x = GlobalMaxPooling2D()(x)
x = Dense(256, activation='relu')(x)
#x = Dropout(0.5)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.5)(x)
cnn = Model(inputs, x)
# High-level features.
inp = Input(shape=(32, )) # 'hsSM', 'hsWW3v2', 'hsALT', 'altID', 'target' -> dropped
x = Dense(units=256, activation='relu')(inp)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
x = Dense(units=256, activation='relu')(x)
#x = Dropout(0.5)(x)
x = Dense(units=256, activation='relu')(x)
#x = Dropout(0.5)(x)
x = Dense(units=256, activation='relu')(x)
x = Dropout(0.5)(x)
ann = Model(inputs=inp, outputs=x)
# Combine
combinedInput = concatenate([cnn.output, ann.output])
x = Dense(256, activation="relu")(combinedInput)
x = Dropout(0.5)(x)
x = Dense(256, activation="relu", name='penultimate')(x)
x = Dropout(0.5)(x)
x = Dense(2, activation="softplus", name='output')(x)
model = Model(inputs=[cnn.input, ann.input], outputs=x)
return model
# +
# Train
file_model = '../models/heteroskedastic_2017.h5'
model = define_model()
model.compile(loss=Gaussian_NLL, optimizer=Adam(lr=0.0001), metrics=[Gaussian_MSE])
# Dataset
batch_size = 128
epochs = 123
#filename = '../../data/alt/sar_hs.h5'
filename = '/mnt/tmp/psadow/sar/sar_hs.h5'
train = SARGenerator(filename=filename,
subgroups=['2015_2016', '2017'],
batch_size=batch_size)
valid = SARGenerator(filename=filename, subgroups=['2018'], batch_size=batch_size)
# filename = '/mnt/tmp/psadow/sar/sar_hs.h5'
# epochs = 25
# train = SARGenerator(filename=filename,
# subgroups=['2015_2016', '2017', '2018'], # Train on all data without early stopping.
# batch_size=batch_size)
# Callbacks
# This LR schedule is slower than in the paper.
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.9, patience=1)
check = ModelCheckpoint(file_model, monitor='val_loss', verbose=0,
save_best_only=True, save_weights_only=False,
mode='auto', save_freq='epoch')
stop = EarlyStopping(monitor='val_loss', min_delta=0, patience=10, verbose=0,
mode='auto', baseline=None, restore_best_weights=False)
clbks = [reduce_lr, check, stop]
history = model.fit(train,
epochs=epochs,
validation_data=valid,
callbacks=clbks,
verbose=1)
| notebooks/train_model_heteroskedastic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danielmlow/tutorials/blob/main/make_GIF.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="D6x91DKj-VhR"
import glob
from PIL import Image
# filepaths
fp_in = output_dir+"mental_map/participant_215_*.png"
fp_out = output_dir+"mental_map/participant_215.gif"
# https://pillow.readthedocs.io/en/stable/handbook/image-file-formats.html#gif
imgs = (Image.open(f) for f in sorted(glob.glob(fp_in)))
img = next(imgs) # extract first image from iterator
# img = Image.open(glob.glob(fp_in[0]))
img.save(fp=fp_out, format='GIF', append_images=imgs,
save_all=True, duration=1000, loop=0)
| make_GIF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/IgorBaratta/wave-direction/blob/master/notebooks/generate_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="sOKDk6tlnCIR" colab_type="code" colab={}
#Imports
# %matplotlib inline
import numpy
import matplotlib.pyplot as plt
from scipy.special import hankel1
# + id="de9wJzbAnPs_" colab_type="code" colab={}
N = 101
u = numpy.linspace(-1, 1, N)
v = numpy.linspace(-1, 1, N)
U, V = numpy.meshgrid(u, v)
X = numpy.hstack((U.reshape((N**2, 1)), V.reshape((N**2, 1))))
omega = 2*numpy.pi # frequency
# + id="ith7Xncc1nlu" colab_type="code" outputId="6ff3206f-e29a-4309-d14a-f62ef4fc5c5f" colab={"base_uri": "https://localhost:8080/", "height": 339}
x0 = numpy.array([0.5, 0.5])
field = numpy.zeros(N**2, dtype=numpy.complex128)
dist = numpy.sqrt(numpy.sum((X - x0)**2, axis=1))
field += 1j/4*numpy.sqrt(omega)*hankel1(0, omega*dist)
field = field.reshape(N, N)
fig, ax = plt.subplots(1,2, figsize=(10, 5))
ax[0].pcolor(field.real)
ax[1].pcolor(field.imag)
# + id="JXLVB2lK1IVm" colab_type="code" outputId="b6f16bd1-3117-45b5-f981-0264d16a1c55" colab={"base_uri": "https://localhost:8080/", "height": 338}
## Generate plane wave from point sources:
Npoints = 5
x0 = numpy.vstack((2 *numpy.random.rand(Npoints), 2 * numpy.random.rand(Npoints))).T
field = numpy.zeros(N**2, dtype=numpy.complex128)
# TODO: vectorize this:
for x in x0:
dist = numpy.sqrt(numpy.sum((X - x)**2, axis=1))
field += 1j/4*numpy.sqrt(omega)*hankel1(0, omega*dist)
field = field.reshape(N, N)
fig, ax = plt.subplots(1,2, figsize=(10, 5))
ax[0].pcolor(field.real)
ax[1].pcolor(field.imag)
# + id="HPXpm_3j6y9_" colab_type="code" colab={}
N = 32
u = numpy.linspace(-1, 1, N)
v = numpy.linspace(-1, 1, N)
U, V = numpy.meshgrid(u, v)
X = numpy.hstack((U.reshape((N**2, 1)), V.reshape((N**2, 1))))
omega = 10*numpy.pi # frequency
def calculate_field(x0):
field = numpy.zeros(N**2, dtype=numpy.complex128)
dist = numpy.sqrt(numpy.sum((X - x0)**2, axis=1))
field += 1j/4*numpy.sqrt(omega)*hankel1(0, omega*dist)
field = field.reshape(N, N)
return field
def generate_data(num_samples, N):
# N is the height and width of the image
# generate data with num_sample samples
x = (numpy.random.rand(num_samples, 2) - 0.5) * 10
img_input = numpy.zeros((num_samples, N, N, 2))
label = numpy.zeros(num_samples)
for i in range(num_samples):
field = calculate_field(x[i])
img_input[i,:,:,0] = field.real
img_input[i,:,:,1] = field.imag
label[i] = ((math.atan2(-x[i][1],-x[i][0]) + 2*numpy.pi) * 180/numpy.pi) % 360
return img_input, label
# + id="F7cO0wtR4-uW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 338} outputId="27994a75-623f-4ef3-cecb-a8998d1e6fe8"
import math
# Number of samples:
training_size = 5000
test_size = 200
train_input, train_label = generate_data(training_size, N)
train_label = train_label/ 360
test_input, test_label = generate_data(test_size, N)
test_label = test_label/ 360
fig, ax = plt.subplots(1,2, figsize=(10, 5))
ax[0].pcolor(train_input[0,:,:,0])
ax[0].plot(N/2, N/2, 'ko')
ax[1].pcolor(train_input[0,:,:,1])
ax[1].plot(N/2, N/2, 'ko')
print("Estimated angle:", train_label[0]*360)
# + id="qc6XoXh_n6HA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="1ca77388-6316-40e4-c141-827f674809f0"
hist, bin_edges = numpy.histogram(train_label)
# An "interface" to matplotlib.axes.Axes.hist() method
n, bins, patches = plt.hist(x=train_label, bins='auto', color='#0504aa',
alpha=0.7, rwidth=0.85)
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Value')
plt.ylabel('Frequency')
maxfreq = n.max()
# Set a clean upper y-axis limit.
plt.ylim(ymax=numpy.ceil(maxfreq / 10) * 10 if maxfreq % 10 else maxfreq + 10)
# + id="c82wSRkREJpW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a0e677e0-c834-4b54-d62e-da89fe23bc71"
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
print(tf.__version__)
# + id="MBp0Rv3bGBwH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 449} outputId="f3153df9-b7b2-49b8-929b-4a4201f2d75e"
model = models.Sequential()
model.add(layers.Conv2D(64, (3, 3), activation='relu', input_shape=(N, N, 2)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
# model.add(layers.Conv2D(64, (3, 3), activation='relu'))
# model.add(layers.MaxPooling2D((2, 2)))
# model.add(layers.InputLayer(input_shape=(N, N, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(256, activation='relu'))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# Display archictecture
model.summary()
# + id="EL4IYDyBO8Ey" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="645c6b64-b187-4fed-ffe5-ae7fc4915c8b"
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=0.0001),
loss=tf.keras.losses.MeanAbsoluteError())
history = model.fit(train_input, train_label, epochs=100,
validation_data=(test_input, test_label))
# + id="VD7vfipDcBhD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 315} outputId="f303fdf8-c47e-42f9-9bf1-12a3e40f779f"
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label = 'val_loss')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
# plt.ylim([0, 1])
plt.legend(loc='upper right')
test_label = test_label.reshape(test_size, 1)
test_loss = model.evaluate(test_input, test_label, verbose=2)
print(test_loss*360)
direction = model.predict(test_input)
# print(numpy.hstack((direction, test_label))*360)
# + id="rVGuRHIBsf83" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="0cfeefa7-31d6-4972-a68d-239e2697883f"
prf = numpy.linspace(0, 360, 100)
plt.plot(prf, prf, linewidth=3)
plt.plot(direction*360, test_label*360, 'o')
| notebooks/generate_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# metadata:
# interpreter:
# hash: 767d51c1340bd893661ea55ea3124f6de3c7a262a8b4abca0554b478b1e2ff90
# name: python3
# ---
# # Analyse results for query dissemination experiment
# + tags=[]
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.set_option('display.max_rows', 500)
# -
# # 1. Parse Logfiles
# +
paths = {
"10": "../../logs/query-dissemination/29-03-2021/selectivity_10",
"25": "../../logs/query-dissemination/29-03-2021/selectivity_25",
"80": "../../logs/query-dissemination/29-03-2021/selectivity_80"
}
num_nodes = 73 # number of nodes per run
num_runs = {
10:3,
25:3,
80:3
}
# -
# The function stores the lat and lon values for each selectivity.
def determine_values(selectivity, idx):
selectivity_to_values = {
10:(90.0,0.0),
25:(50.0, 50.0),
80:(20.0,0.0)
}
val = selectivity_to_values[selectivity]
return val[idx]
# + tags=[]
sels = []
categories = []
node_ids = []
lats = []
lons = []
runs = []
connect_outliers = []
received = []
runs_rec = []
sels_received = []
query_times = []
startup_times = {}
for selectivity in paths:
for root, dirs, files in os.walk(paths[selectivity]):
#print("Root: " + root)
run = root.split("/")[-1].split("_")[-1]
for file in files :
#print("File: " + file)
n_id = file[:6]
if n_id[:4] != 'base' :
# get node_id
if n_id[-1:] == '_' :
node_id = int(n_id[4:5])
else :
node_id = int(n_id[4:6])
#print(node_id)
with open(os.path.join(root, file)) as log:
connect_attempts = 0
query_time = 100000
for line in log :
log_time = int(line.split(" ")[0])
elem = line.split( )
if "initial attribute range" in line:
tmp = str(elem[-1:])
t = tmp[3:-3]
coords = t.split(',')
lat = float(coords[0])
lon = float(coords[1])
node_ids.append( node_id)
lats.append(lat)
lons.append(lon)
runs.append(run)
sels.append(selectivity)
if "! RECEIVED QUERY" in line :
received.append(int(node_id))
runs_rec.append(run)
sels_received.append(selectivity)
query_time = elem[8]
query_times.append(query_time)
elif "STARTUP TIME" in line:
elem = line.split( )
startup_times[(selectivity, run)] = int(elem[-1])
# sanity checks
if "ACTOR DOWN" in line and log_time < query_time and time < 59000:
print("Actor down in: selectivity " + selectivity + ", run " + str(run) + ", node" + str(node_id) + ", query time: "
+ str(query_time) + ", log_time: " + str(log_time))
if "RECONNECTING TO INITIAL PARENT" in line:
print("Initial connection issues in: selectivity " + selectivity + ", run " + str(run) + ", node" + str(node_id))
if "could not reach" in line:
print("Found Error in: selectivity " + selectivity + ", run " + run + ", node" + node_id)
if (connect_attempts > 5):
m_connect_outliers.append("run " + run + " node " + node_id)
#print(received)
#print('node_ids ' + str(len(node_ids)))
#print('categories ' + str(len(categories)))
#print('lat ' + str(len(lats)))
#print('lon ' + str(len(lons)))
#print(len(sels))
initial_ranges_tmp = pd.DataFrame(np.column_stack([sels,runs,node_ids, lats, lons]), columns=['selectivity','run', 'node_id','lat', 'lon'])
initial_ranges = initial_ranges_tmp.astype({
'selectivity':'int32',
'run':'int32',
'node_id':'int32',
'lat':'float',
'lon':'float'
})
print(initial_ranges.dtypes)
initial_ranges
# -
# ## Outliers & Sanity checks
print("Nodes with connection issues:")
connect_outliers
nan = initial_ranges[initial_ranges.isna().any(axis=1)]
nan
nul = initial_ranges[initial_ranges.isnull().any(axis=1)]
nul
# # 2. Determine which nodes received the query
# For all selectivities, create a df that contains all nodes that received the query in each run
# +
received_tmp = pd.DataFrame(np.column_stack([sels_received, runs_rec, received, query_times]),
columns=['selectivity','run', 'node_id', 'time_received'])
received = received_tmp.astype({
'selectivity':'int64',
'run':'int64',
'node_id':'int64',
'time_received':'int64'
})
received['time_received'] = received.apply(lambda row: row['time_received'] - startup_times[(str(row['selectivity']),str(row['run']))], axis=1)
received.head()
# -
received['time_received'].max()
# # 3. Delete all nodes from the df that did not match the query
# + tags=[]
nodes_applying = initial_ranges[
(initial_ranges['lat'] >= initial_ranges.apply(lambda x: determine_values(x['selectivity'],0), axis=1) ) &
(initial_ranges['lon'] >= initial_ranges.apply(lambda x: determine_values(x['selectivity'],1), axis=1) )
]
nodes_applying.groupby(['selectivity']).size().reset_index(name='applies_to')
# -
# ### Sanity check
#
# Find the nodes where the query applied according to (lat,lon), but did not receive it
a_waldo = nodes_applying.merge(received, how='left', on=['selectivity', 'run', 'node_id'])
len(a_waldo)
w = a_waldo[a_waldo.isnull().any(axis=1)]
w.head()
# # 4. Count the nodes that the respective query applies to in each run for every selectivity
# + tags=[]
grouped = nodes_applying.groupby(['selectivity','run']).size().reset_index(name='applies_to')
print(grouped.dtypes)
grouped
# -
# # 5. Count the nodes that received the query in each run for every selectivity
received_grouped = received.groupby(['selectivity','run']).size().reset_index(name='num_received')
print(received_grouped.dtypes)
received_grouped
# # 6. Compute Overhead etc.
#
# ### Table columns
# - applies_to = number of nodes where the query applies
# - received = number of nodes that received the query
# - overhead = received - applies
# - baseline_ovehead = num_nodes - applies
# - num_nodes = topology size
# - overhead_ratio = overhead/baseline_overhead = ratio
# - overhead_ratio_pct = overhead_ratio * 100
overview_tmp = grouped.merge(received_grouped, how='left', on=['selectivity', 'run'])#.astype({'num_received':'int64'})
print(overview_tmp.dtypes)
overview_tmp
# +
overview = overview_tmp
overview['rime_overhead_abs'] = overview.apply(lambda row: row.num_received - row.applies_to, axis=1 )
overview['num_nodes'] = num_nodes
overview['baseline_overhead'] = overview.apply(lambda row: num_nodes - row.applies_to, axis=1)
overview['overhead_ratio'] = overview.apply(lambda row: row.rime_overhead_abs/row.baseline_overhead, axis=1)
overview['overhead_ratio_pct'] = overview.apply(lambda row: row.overhead_ratio*100, axis=1)
print(overview.dtypes)
overview
# + tags=[]
summary = pd.DataFrame()
summary['mean_overhead_ratio'] = overview.groupby('selectivity')['overhead_ratio_pct'].mean()
summary['median_overhead_ratio'] = overview.groupby('selectivity')['overhead_ratio_pct'].median()
summary['applies_pct'] = overview.groupby('selectivity')['applies_to'].mean() / num_nodes
summary['received_pct'] = overview.groupby('selectivity')['num_received'].mean() / num_nodes
summary['base_overhead'] = 1 - summary['applies_pct']
summary['rime_overhead'] = summary['received_pct'] - summary['applies_pct']
summary['overhead_saved_by_rime'] = (summary['base_overhead'] - summary['rime_overhead']) * 100
# -
summary
| benchmarks/notebooks/query-dissemination/query-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SaQphvIrtYdw"
# # Technology Explorers Course 2, Lab 1: Practice with Pandas
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# <br>
#
# ---
#
# <br>
#
# In this lab we will continue to practice with pandas
#
# <br>
#
# ---
#
#
#
# + id="rM5rDI1dt403" executionInfo={"status": "ok", "timestamp": 1627522126907, "user_tz": 420, "elapsed": 194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}}
import pandas as pd
import numpy as np
# + [markdown] id="0PPaZvlct0fH"
# # Q1
#
# Convert the two series into the columns of a DataFrame
# + id="9lphFb2ksnEC" executionInfo={"status": "ok", "timestamp": 1627522131926, "user_tz": 420, "elapsed": 159, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="17ade1db-a0cd-4c3d-ec0f-2f76bc93ca6b" colab={"base_uri": "https://localhost:8080/", "height": 860}
ser1 = pd.Series(list('abcedfghijklmnopqrstuvwxyz'))
ser2 = pd.Series(np.arange(26))
pd.DataFrame([ser1,ser2]).T
# + [markdown] id="0gi9EjLNuLGK"
# # Q2
#
# Conver the series into a DataFrame with 7 rows and 5 columns
# + id="EXkjgWoOt395"
ser = pd.Series(np.random.randint(1, 10, 35))
# + [markdown] id="HK6nG-UPuvkT"
# # Q3
#
# Compute the difference of differences between consecutive numbers in a series using `ser.diff()`
#
# + id="kAh1JuzOu1A6"
ser = pd.Series([1, 3, 6, 10, 15, 21, 27, 35])
# + [markdown] id="08Phen9ru9fb"
# # Q4
#
# Convert a series of dates to `datetime` format using `pd.to_datetime()`
# + id="kz7TvktXu1ai"
ser = pd.Series(['01 Jan 2010', '02-02-2011', '20120303', '2013/04/04', '2014-05-05', '2015-06-06T12:20'])
# + [markdown] id="Yt6pOFN818x_"
# # Q5
#
# Compute the mean of weights grouped by fruit
#
# + colab={"base_uri": "https://localhost:8080/"} id="KFdCPjZWvEy_" executionInfo={"status": "ok", "timestamp": 1626733030500, "user_tz": 420, "elapsed": 162, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="045abf56-082b-4645-c05c-d473915e54e1"
fruit = pd.Series(np.random.choice(['apple', 'banana', 'carrot'], 10))
weights = pd.Series(np.linspace(1, 10, 10))
print(weights.tolist())
print(fruit.tolist())
# + [markdown] id="M8Igu4sK2L1y"
# # Q6
#
# Compute the euclidian distance between vectors p and q (pythagorean theorem)
# + id="1SWVo8vn2Ay9"
p = pd.Series([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
q = pd.Series([10, 9, 8, 7, 6, 5, 4, 3, 2, 1])
# + [markdown] id="HWuS2x092yxA"
# # Q7
#
# Fill in missing values for dates with the previous dates' value using `ser.bfill()` or `ser.ffill()`
# + colab={"base_uri": "https://localhost:8080/"} id="HO0gcmvJ2ObL" executionInfo={"status": "ok", "timestamp": 1626733250745, "user_tz": 420, "elapsed": 165, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="61f7bc19-0085-4a27-a7c7-f02b4c94445f"
ser = pd.Series([1,10,3,np.nan], index=pd.to_datetime(['2000-01-01', '2000-01-03', '2000-01-06', '2000-01-08']))
print(ser)
# + [markdown] id="x1H949kQ3dvJ"
# # Q8
#
# Check if there are missing values in a dataframe using `.isnull()`, `.values`, and `.any()`
# + id="N12aXr6v24cu"
df = pd.read_csv('https://raw.githubusercontent.com/selva86/datasets/master/Cars93_miss.csv')
# + [markdown] id="EboeGY0p4mb7"
# # Q9
#
# Grab the first column and return it as a DataFrame rather than as a series
# + id="LMpa73403jce"
df = pd.DataFrame(np.arange(20).reshape(-1, 5), columns=list('abcde'))
# + [markdown] id="EdgvQfqq4vux"
# # Q10
#
# In `df`, interchange columns 'a' and 'c'.
# + id="d5Oz2gzL4pz2"
df = pd.DataFrame(np.arange(20).reshape(-1, 5), columns=list('abcde'))
# + id="yc34SOU140oq"
| Fundamentals II/SOLUTIONS/SOLUTION_Tech_Fun_C2_L1__Practice_with_Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] colab_type="text" id="4embtkV0pNxM"
# Deep Learning
# =============
#
# Assignment 4
# ------------
#
# Previously in `2_fullyconnected.ipynb` and `3_regularization.ipynb`, we trained fully connected networks to classify [notMNIST](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html) characters.
#
# The goal of this assignment is make the neural network convolutional.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="tm2CQN_Cpwj0"
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 11948, "status": "ok", "timestamp": 1446658914837, "user": {"color": "", "displayName": "", "isAnonymous": false, "isMe": true, "permissionId": "", "photoUrl": "", "sessionId": "0", "userId": ""}, "user_tz": 480} id="y3-cj1bpmuxc" outputId="016b1a51-0290-4b08-efdb-8c95ffc3cd01"
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# + [markdown] colab_type="text" id="L7aHrm6nGDMB"
# Reformat into a TensorFlow-friendly shape:
# - convolutions need the image data formatted as a cube (width by height by #channels)
# - labels as float 1-hot encodings.
# -
(np.arange(10) == test_labels[0]).astype(float)
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 11952, "status": "ok", "timestamp": 1446658914857, "user": {"color": "", "displayName": "", "isAnonymous": false, "isMe": true, "permissionId": "", "photoUrl": "", "sessionId": "0", "userId": ""}, "user_tz": 480} id="IRSyYiIIGIzS" outputId="650a208c-8359-4852-f4f5-8bf10e80ef6c"
image_size = 28
num_labels = 10
num_channels = 1 # grayscale
import numpy as np
def reformat(dataset, labels):
dataset = dataset.reshape(
(-1, image_size, image_size, num_channels)).astype(np.float32)
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="AgQDIREv02p1"
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
# + [markdown] colab_type="text" id="5rhgjmROXu2O"
# Let's build a small network with two convolutional layers, followed by one fully connected layer. Convolutional networks are more expensive computationally, so we'll limit its depth and number of fully connected nodes.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="IZYv70SvvOan"
batch_size = 16
patch_size = 5
depth = 16
num_hidden = 64
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, num_channels, depth], stddev=0.1))
layer1_biases = tf.Variable(tf.zeros([depth]))
layer2_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, depth, depth], stddev=0.1))
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth]))
layer3_weights = tf.Variable(tf.truncated_normal(
[image_size // 4 * image_size // 4 * depth, num_hidden], stddev=0.1))
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden]))
layer4_weights = tf.Variable(tf.truncated_normal(
[num_hidden, num_labels], stddev=0.1))
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]))
# Model.
def model(data):
conv = tf.nn.conv2d(data, layer1_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer1_biases)
conv = tf.nn.conv2d(hidden, layer2_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer2_biases)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer3_weights) + layer3_biases)
return tf.matmul(hidden, layer4_weights) + layer4_biases
# Training computation.
logits = model(tf_train_dataset)
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.05).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset))
test_prediction = tf.nn.softmax(model(tf_test_dataset))
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 37}]} colab_type="code" executionInfo={"elapsed": 63292, "status": "ok", "timestamp": 1446658966251, "user": {"color": "", "displayName": "", "isAnonymous": false, "isMe": true, "permissionId": "", "photoUrl": "", "sessionId": "0", "userId": ""}, "user_tz": 480} id="noKFb2UovVFR" outputId="28941338-2ef9-4088-8bd1-44295661e628"
num_steps = 1001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size), :]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 50 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
# + [markdown] colab_type="text" id="KedKkn4EutIK"
# ---
# Problem 1
# ---------
#
# The convolutional model above uses convolutions with stride 2 to reduce the dimensionality. Replace the strides by a max pooling operation (`nn.max_pool()`) of stride 2 and kernel size 2.
#
# ---
# +
batch_size = 16
patch_size = 5
depth = 16
num_hidden = 64
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, num_channels, depth], stddev=0.1))
layer1_biases = tf.Variable(tf.zeros([depth]))
layer2_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, depth, depth], stddev=0.1))
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth]))
layer3_weights = tf.Variable(tf.truncated_normal(
[image_size // 4 * image_size // 4 * depth, num_hidden], stddev=0.1))
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden]))
layer4_weights = tf.Variable(tf.truncated_normal(
[num_hidden, num_labels], stddev=0.1))
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]))
# Model.
def model(data):
conv = tf.nn.conv2d(data, layer1_weights, [1, 1, 1, 1], padding='SAME')
pool = tf.nn.max_pool(conv, [1, 2, 2, 1], [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(pool + layer1_biases)
conv = tf.nn.conv2d(hidden, layer2_weights, [1, 1, 1, 1], padding='SAME')
pool = tf.nn.max_pool(conv, [1, 2, 2, 1], [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(pool + layer2_biases)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer3_weights) + layer3_biases)
return tf.matmul(hidden, layer4_weights) + layer4_biases
# Training computation.
logits = model(tf_train_dataset)
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.05).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset))
test_prediction = tf.nn.softmax(model(tf_test_dataset))
# +
num_steps = 5001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size), :]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
# + [markdown] colab_type="text" id="klf21gpbAgb-"
# ---
# Problem 2
# ---------
#
# Try to get the best performance you can using a convolutional net. Look for example at the classic [LeNet5](http://yann.lecun.com/exdb/lenet/) architecture, adding Dropout, and/or adding learning rate decay.
#
# ---
# +
batch_size = 128
num_hidden = 84
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
global_step = tf.Variable(0)
C1_weights = tf.Variable(tf.truncated_normal([5, 5, num_channels, 6], stddev=0.1))
C1_biases = tf.Variable(tf.zeros([6]))
C3_weights = tf.Variable(tf.truncated_normal([5, 5, 6, 16], stddev=0.1))
C3_biases = tf.Variable(tf.constant(1.0, shape=[16]))
C5_weights = tf.Variable(tf.truncated_normal([5, 5, 16, 120], stddev=0.1))
C5_biases = tf.Variable(tf.constant(1.0, shape=[120]))
FC6_weights = tf.Variable(tf.truncated_normal([120, num_hidden], stddev=0.1))
FC6_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden]))
Output_weights = tf.Variable(tf.truncated_normal([num_hidden, num_labels], stddev=0.1))
Output_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]))
# Model inspired by LeNET5
def model(data, dropout_prob):
conv = tf.nn.conv2d(data, C1_weights, [1, 1, 1, 1], padding='SAME')
hidden = tf.nn.dropout(tf.nn.relu(conv + C1_biases), dropout_prob)
pool = tf.nn.max_pool(hidden, [1, 2, 2, 1], [1, 2, 2, 1], padding='SAME')
conv = tf.nn.conv2d(pool, C3_weights, [1, 1, 1, 1], padding='VALID')
hidden = tf.nn.dropout(tf.nn.relu(conv + C3_biases), dropout_prob)
pool = tf.nn.max_pool(hidden, [1, 2, 2, 1], [1, 2, 2, 1], padding='VALID')
conv = tf.nn.conv2d(pool, C5_weights, [1, 1, 1, 1], padding='VALID')
hidden = tf.nn.dropout(tf.nn.relu(conv + C5_biases), dropout_prob)
#print(hidden.get_shape().as_list())
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1]*shape[2]*shape[3]])
fc = tf.nn.relu(tf.matmul(reshape, FC6_weights) + FC6_biases)
hidden = tf.nn.dropout(fc, dropout_prob)
return tf.matmul(hidden, Output_weights) + Output_biases
# Training computation.
logits = model(tf_train_dataset, 0.5)
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
learning_rate = tf.train.exponential_decay(0.05, global_step, 2000, 0.9)
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss, global_step=global_step)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset, 1))
test_prediction = tf.nn.softmax(model(tf_test_dataset, 1))
# +
num_steps = 5001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size), :]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions, lr = session.run(
[optimizer, loss, train_prediction, learning_rate], feed_dict=feed_dict)
if (step % 500 == 0):
print('Learning Rate: %.5f' % lr)
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
| udacity_Deep_Learning/.ipynb_checkpoints/4_convolutions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MNIST with SciKit-Learn and skorch
#
# This notebooks shows how to define and train a simple Neural-Network with PyTorch and use it via skorch with SciKit-Learn.
from sklearn.datasets import fetch_mldata
from sklearn.model_selection import train_test_split
import numpy as np
# ## Loading Data
# Using SciKit-Learns ```fetch_mldata``` to load MNIST data.
mnist = fetch_mldata('MNIST original')
mnist
mnist.data.shape
# ## Preprocessing Data
#
# Each image of the MNIST dataset is encoded in a 784 dimensional vector, representing a 28 x 28 pixel image. Each pixel has a value between 0 and 255, corresponding to the grey-value of a pixel.<br />
# The above ```featch_mldata``` method to load MNIST returns ```data``` and ```target``` as ```uint8``` which we convert to ```float32``` and ```int64``` respectively.
X = mnist.data.astype('float32')
y = mnist.target.astype('int64')
# As we will use ReLU as activation in combination with softmax over the output layer, we need to scale `X` down. An often use range is [0, 1].
X /= 255.0
X.min(), X.max()
# Note: data is not normalized.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
assert(X_train.shape[0] + X_test.shape[0] == mnist.data.shape[0])
X_train.shape, y_train.shape
# ## Build Neural Network with Torch
# Simple, fully connected neural network with one hidden layer. Input layer has 784 dimensions (28x28), hidden layer has 98 (= 784 / 8) and output layer 10 neurons, representing digits 0 - 9.
import torch
from torch import nn
import torch.nn.functional as F
torch.manual_seed(0);
mnist_dim = X.shape[1]
hidden_dim = int(mnist_dim/8)
output_dim = len(np.unique(mnist.target))
mnist_dim, hidden_dim, output_dim
# A Neural network in PyTorch's framework.
class ClassifierModule(nn.Module):
def __init__(
self,
input_dim=mnist_dim,
hidden_dim=hidden_dim,
output_dim=output_dim,
dropout=0.5,
):
super(ClassifierModule, self).__init__()
self.dropout = nn.Dropout(dropout)
self.hidden = nn.Linear(input_dim, hidden_dim)
self.output = nn.Linear(hidden_dim, output_dim)
def forward(self, X, **kwargs):
X = F.relu(self.hidden(X))
X = self.dropout(X)
X = F.softmax(self.output(X), dim=-1)
return X
# Skorch allows to use PyTorch's networks in the SciKit-Learn setting.
from skorch.net import NeuralNetClassifier
net = NeuralNetClassifier(
ClassifierModule,
max_epochs=20,
lr=0.1,
# device='cuda', # uncomment this to train with CUDA
)
net.fit(X_train, y_train);
# ## Prediction
predicted = net.predict(X_test)
np.mean(predicted == y_test)
# An accuracy of nearly 96% for a network with only one hidden layer is not too bad
# # Convolutional Network
# PyTorch expects a 4 dimensional tensor as input for its 2D convolution layer. The dimensions represent:
# * Batch size
# * Number of channel
# * Height
# * Width
#
# As initial batch size the number of examples needs to be provided. MNIST data has only one channel. As stated above, each MNIST vector represents a 28x28 pixel image. Hence, the resulting shape for PyTorch tensor needs to be (x, 1, 28, 28).
XCnn = X.reshape(-1, 1, 28, 28)
XCnn.shape
XCnn_train, XCnn_test, y_train, y_test = train_test_split(XCnn, y, test_size=0.25, random_state=42)
XCnn_train.shape, y_train.shape
class Cnn(nn.Module):
def __init__(self):
super(Cnn, self).__init__()
self.conv1 = nn.Conv2d(1, 32, kernel_size=3)
self.conv2 = nn.Conv2d(32, 64, kernel_size=3)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(1600, 128) # 1600 = number channels * width * height
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, x.size(1) * x.size(2) * x.size(3)) # flatten over channel, height and width = 1600
x = F.relu(self.fc1(x))
x = F.dropout(x, training=self.training)
x = self.fc2(x)
x = F.softmax(x, dim=-1)
return x
cnn = NeuralNetClassifier(
Cnn,
max_epochs=15,
lr=1,
optimizer=torch.optim.Adadelta,
# device='cuda', # uncomment this to train with CUDA
)
cnn.fit(XCnn_train, y_train);
cnn_pred = cnn.predict(XCnn_test)
np.mean(cnn_pred == y_test)
# An accuracy of 99.1% should suffice for this example!
| examples/MNIST/MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf2.0
# language: python
# name: tf2.0
# ---
# # 模型构造
#
# 让我们回顾一下在[“多层感知机的简洁实现”](../chapter_deep-learning-basics/mlp-gluon.ipynb)一节中含单隐藏层的多层感知机的实现方法。我们首先构造`Sequential`实例,然后依次添加两个全连接层。其中第一层的输出大小为256,即隐藏层单元个数是256;第二层的输出大小为10,即输出层单元个数是10。我们在上一章的其他
# 节中也使用了`Sequential`类构造模型。这里我们介绍另外一种基于`tf.keras.Model`类的模型构造方法:它让模型构造更加灵活。
#
#
# ## 4.1.1 build model from block
#
# `tf.keras.Model`类是`tf.keras`模块里提供的一个模型构造类,我们可以继承它来定义我们想要的模型。下面继承`tf.keras.Model`类构造本节开头提到的多层感知机。这里定义的`MLP`类重载了`tf.keras.Model`类的`__init__`函数和`call`函数。它们分别用于创建模型参数和定义前向计算。前向计算也即正向传播。
import tensorflow as tf
import numpy as np
print(tf.__version__)
class MLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.flatten = tf.keras.layers.Flatten() # Flatten层将除第一维(batch_size)以外的维度展平
self.dense1 = tf.keras.layers.Dense(units=256, activation=tf.nn.relu)
self.dense2 = tf.keras.layers.Dense(units=10)
def call(self, inputs):
x = self.flatten(inputs)
x = self.dense1(x)
output = self.dense2(x)
return output
# 以上的`MLP`类中无须定义反向传播函数。系统将通过自动求梯度而自动生成反向传播所需的`backward`函数。
#
# 我们可以实例化`MLP`类得到模型变量`net`。下面的代码初始化`net`并传入输入数据`X`做一次前向计算。其中,`net(X)`将调用`MLP`类定义的`call`函数来完成前向计算。
X = tf.random.uniform((2,20))
net = MLP()
net(X)
# ## 4.1.2 Sequential
#
# 我们刚刚提到,`tf.keras.Model`类是一个通用的部件。事实上,`Sequential`类继承自`tf.keras.Model`类。当模型的前向计算为简单串联各个层的计算时,可以通过更加简单的方式定义模型。这正是`Sequential`类的目的:它提供`add`函数来逐一添加串联的`Block`子类实例,而模型的前向计算就是将这些实例按添加的顺序逐一计算。
#
# 我们用Sequential类来实现前面描述的MLP类,并使用随机初始化的模型做一次前向计算。
# +
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(10),
])
model(X)
# -
# ## 4.1.3 build complex model
# 虽然`Sequential`类可以使模型构造更加简单,且不需要定义`call`函数,但直接继承`tf.keras.Model`类可以极大地拓展模型构造的灵活性。下面我们构造一个稍微复杂点的网络`FancyMLP`。在这个网络中,我们通过`constant`函数创建训练中不被迭代的参数,即常数参数。在前向计算中,除了使用创建的常数参数外,我们还使用`tensor`的函数和Python的控制流,并多次调用相同的层。
class FancyMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.flatten = tf.keras.layers.Flatten()
self.rand_weight = tf.constant(
tf.random.uniform((20,20)))
self.dense = tf.keras.layers.Dense(units=20, activation=tf.nn.relu)
def call(self, inputs):
x = self.flatten(inputs)
x = tf.nn.relu(tf.matmul(x, self.rand_weight) + 1)
x = self.dense(x)
while tf.norm(x) > 1:
x /= 2
if tf.norm(x) < 0.8:
x *= 10
return tf.reduce_sum(x)
# 在这个`FancyMLP`模型中,我们使用了常数权重`rand_weight`(注意它不是模型参数)、做了矩阵乘法操作(`tf.matmul`)并重复使用了相同的`Dense`层。下面我们来测试该模型的随机初始化和前向计算。
net = FancyMLP()
net(X)
# 因为`FancyMLP`和`Sequential`类都是`tf.keras.Model`类的子类,所以我们可以嵌套调用它们。
# +
class NestMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.net = tf.keras.Sequential()
self.net.add(tf.keras.layers.Flatten())
self.net.add(tf.keras.layers.Dense(64, activation=tf.nn.relu))
self.net.add(tf.keras.layers.Dense(32, activation=tf.nn.relu))
self.dense = tf.keras.layers.Dense(units=16, activation=tf.nn.relu)
def call(self, inputs):
return self.dense(self.net(inputs))
net = tf.keras.Sequential()
net.add(NestMLP())
net.add(tf.keras.layers.Dense(20))
net.add(FancyMLP())
net(X)
| code/chapter04_DL-computation/4.1_model-construction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import numpy as np
import cv2
from PIL import Image
import torch.nn as nn
from torch.utils.data import Dataset
import os
os.environ["KMP_DUPLICATE_LIB_OK"] = "TRUE"
# -
import albumentations as A
from albumentations.pytorch import ToTensorV2
# +
class ImageFolder(Dataset):
def __init__(self, root_dir, transform=None):
super(ImageFolder, self).__init__()
self.data = []
self.root_dir = root_dir
self.transform = transform
self.class_names = os.listdir(root_dir)
for index, name in enumerate(self.class_names):
files = os.listdir(os.path.join(root_dir, name))
self.data += list(zip(files, [index]*len(files)))
def __len__(self):
return len(self.data)
def __getitem__(self, index):
img_file, label = self.data[index]
root_and_dir = os.path.join(self.root_dir, self.class_names[label])
image = np.array(Image.open(os.path.join(root_and_dir, img_file)))
if self.transform is not None:
augmentations = self.transform(image=image)
image = augmentations["image"]
return image, label
transform = A.Compose(
[
A.Resize(width=1920, height=1080),
A.RandomCrop(width=1280, height=720),
A.Rotate(limit=40, p=0.9, border_mode=cv2.BORDER_CONSTANT),
A.HorizontalFlip(p=0.5),
A.VerticalFlip(p=0.1),
A.RGBShift(r_shift_limit=25, g_shift_limit=25, b_shift_limit=25, p=0.9),
A.OneOf([
A.Blur(blur_limit=3, p=0.5),
A.ColorJitter(p=0.5),
], p=1.0),
A.Normalize(
mean=[0, 0, 0],
std=[1, 1, 1],
max_pixel_value=255,
),
ToTensorV2(),
]
)
# -
# +
dataset = ImageFolder(root_dir="cat_dogs", transform=transform)
for x,y in dataset:
print(x.shape)
| 12_Data_augmentation/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression
#
# *Supervised* machine learning techniques involve training a model to operate on a set of *features* and predict a *label* using a dataset that includes some already-known label values. The training process *fits* the features to the known labels to define a general function that can be applied to new features for which the labels are unknown, and predict them. You can think of this function like this, in which ***y*** represents the label we want to predict and ***x*** represents the features the model uses to predict it.
#
# $$y = f(x)$$
#
# In most cases, *x* is actually a *vector* that consists of multiple feature values, so to be a little more precise, the function could be expressed like this:
#
# $$y = f([x_1, x_2, x_3, ...])$$
#
# The goal of training the model is to find a function that performs some kind of calculation to the *x* values that produces the result *y*. We do this by applying a machine learning *algorithm* that tries to fit the *x* values to a calculation that produces *y* reasonably accurately for all of the cases in the training dataset.
#
# There are lots of machine learning algorithms for supervised learning, and they can be broadly divided into two types:
#
# - **_Regression_ algorithms**: Algorithms that predict a *y* value that is a numeric value, such as the price of a house or the number of sales transactions.
# - **_Classification_ algorithms**: Algorithms that predict to which category, or *class*, an observation belongs. The *y* value in a classification model is a vector of probability values between 0 and 1, one for each class, indicating the probability of the observation belonging to each class.
#
# In this notebook, we'll focus on *regression*, using an example based on a real study in which data for a bicycle sharing scheme was collected and used to predict the number of rentals based on seasonality and weather conditions. We'll use a simplified version of the dataset from that study.
#
# > **Citation**: The data used in this exercise is derived from [Capital Bikeshare](https://www.capitalbikeshare.com/system-data) and is used in accordance with the published [license agreement](https://www.capitalbikeshare.com/data-license-agreement).
#
# ## Explore the Data
#
# The first step in any machine learning project is to explore the data that you will use to train a model. The goal of this exploration is to try to understand the relationships between its attributes; in particular, any apparent correlation between the *features* and the *label* your model will try to predict. This may require some work to detect and fix issues in the data (such as dealing with missing values, errors, or outlier values), deriving new feature columns by transforming or combining existing features (a process known as *feature engineering*), *normalizing* numeric features (values you can measure or count) so they're on a similar scale, and *encoding* categorical features (values that represent discrete categories) as numeric indicators.
#
# Let's start by loading the bicycle sharing data as a **Pandas** DataFrame and viewing the first few rows.
# +
import pandas as pd
# load the training dataset
bike_data = pd.read_csv('data/daily-bike-share.csv')
bike_data.head()
# -
# The data consists of the following columns:
#
# - **instant**: A unique row identifier
# - **dteday**: The date on which the data was observed - in this case, the data was collected daily; so there's one row per date.
# - **season**: A numerically encoded value indicating the season (1:spring, 2:summer, 3:fall, 4:winter)
# - **yr**: The year of the study in which the observation was made (the study took place over two years - year 0 represents 2011, and year 1 represents 2012)
# - **mnth**: The calendar month in which the observation was made (1:January ... 12:December)
# - **holiday**: A binary value indicating whether or not the observation was made on a public holiday)
# - **weekday**: The day of the week on which the observation was made (0:Sunday ... 6:Saturday)
# - **workingday**: A binary value indicating whether or not the day is a working day (not a weekend or holiday)
# - **weathersit**: A categorical value indicating the weather situation (1:clear, 2:mist/cloud, 3:light rain/snow, 4:heavy rain/hail/snow/fog)
# - **temp**: The temperature in celsius (normalized)
# - **atemp**: The apparent ("feels-like") temperature in celsius (normalized)
# - **hum**: The humidity level (normalized)
# - **windspeed**: The windspeed (normalized)
# - **rentals**: The number of bicycle rentals recorded.
#
# In this dataset, **rentals** represents the label (the *y* value) our model must be trained to predict. The other columns are potential features (*x* values).
#
# As mentioned previously, you can perform some *feature engineering* to combine or derive new features. For example, let's add a new column named **day** to the dataframe by extracting the day component from the existing **dteday** column. The new column represents the day of the month from 1 to 31.
bike_data['day'] = pd.DatetimeIndex(bike_data['dteday']).day
bike_data.head(32)
# OK, let's start our analysis of the data by examining a few key descriptive statistics. We can use the dataframe's **describe** method to generate these for the numeric features as well as the **rentals** label column.
numeric_features = ['temp', 'atemp', 'hum', 'windspeed']
bike_data[numeric_features + ['rentals']].describe()
# The statistics reveal some information about the distribution of the data in each of the numeric fields, including the number of observations (there are 731 records), the mean, standard deviation, minimum and maximum values, and the quartile values (the threshold values for 25%, 50% - which is also the median, and 75% of the data). From this, we can see that the mean number of daily rentals is around 848; but there's a comparatively large standard deviation, indicating a lot of variance in the number of rentals per day.
#
# We might get a clearer idea of the distribution of rentals values by visualizing the data. Common plot types for visualizing numeric data distributions are *histograms* and *box plots*, so let's use Python's **matplotlib** library to create one of each of these for the **rentals** column.
# +
import pandas as pd
import matplotlib.pyplot as plt
# This ensures plots are displayed inline in the Jupyter notebook
# %matplotlib inline
# Get the label column
label = bike_data['rentals']
# Create a figure for 2 subplots (2 rows, 1 column)
fig, ax = plt.subplots(2, 1, figsize = (9,12))
# Plot the histogram
ax[0].hist(label, bins=100)
ax[0].set_ylabel('Frequency')
# Add lines for the mean, median, and mode
ax[0].axvline(label.mean(), color='magenta', linestyle='dashed', linewidth=2)
ax[0].axvline(label.median(), color='cyan', linestyle='dashed', linewidth=2)
# Plot the boxplot
ax[1].boxplot(label, vert=False)
ax[1].set_xlabel('Rentals')
# Add a title to the Figure
fig.suptitle('Rental Distribution')
# Show the figure
fig.show()
# -
# The plots show that the number of daily rentals ranges from 0 to just over 3,400. However, the mean (and median) number of daily rentals is closer to the low end of that range, with most of the data between 0 and around 2,200 rentals. The few values above this are shown in the box plot as small circles, indicating that they are *outliers* - in other words, unusually high or low values beyond the typical range of most of the data.
#
# We can do the same kind of visual exploration of the numeric features. Let's create a histogram for each of these.
# Plot a histogram for each numeric feature
for col in numeric_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
feature = bike_data[col]
feature.hist(bins=100, ax = ax)
ax.axvline(feature.mean(), color='magenta', linestyle='dashed', linewidth=2)
ax.axvline(feature.median(), color='cyan', linestyle='dashed', linewidth=2)
ax.set_title(col)
plt.show()
# The numeric features seem to be more *normally* distributed, with the mean and median nearer the middle of the range of values, coinciding with where the most commonly occurring values are.
#
# > **Note**: The distributions are not truly *normal* in the statistical sense, which would result in a smooth, symmetric "bell-curve" histogram with the mean and mode (the most common value) in the center; but they do generally indicate that most of the observations have a value somewhere near the middle.
#
# We've explored the distribution of the numeric values in the dataset, but what about the categorical features? These aren't continuous numbers on a scale, so we can't use histograms; but we can plot a bar chart showing the count of each discrete value for each category.
# +
import numpy as np
# plot a bar plot for each categorical feature count
categorical_features = ['season','mnth','holiday','weekday','workingday','weathersit', 'day']
for col in categorical_features:
counts = bike_data[col].value_counts().sort_index()
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
counts.plot.bar(ax = ax, color='steelblue')
ax.set_title(col + ' counts')
ax.set_xlabel(col)
ax.set_ylabel("Frequency")
plt.show()
# -
# Many of the categorical features show a more or less *uniform* distribution (meaning there's roughly the same number of rows for each category). Exceptions to this include:
#
# - **holiday**: There are many fewer days that are holidays than days that aren't.
# - **workingday**: There are more working days than non-working days.
# - **weathersit**: Most days are category *1* (clear), with category *2* (mist and cloud) the next most common. There are comparatively few category *3* (light rain or snow) days, and no category *4* (heavy rain, hail, or fog) days at all.
#
# Now that we know something about the distribution of the data in our columns, we can start to look for relationships between the features and the **rentals** label we want to be able to predict.
#
# For the numeric features, we can create scatter plots that show the intersection of feature and label values. We can also calculate the *correlation* statistic to quantify the apparent relationship..
for col in numeric_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
feature = bike_data[col]
label = bike_data['rentals']
correlation = feature.corr(label)
plt.scatter(x=feature, y=label)
plt.xlabel(col)
plt.ylabel('Bike Rentals')
ax.set_title('rentals vs ' + col + '- correlation: ' + str(correlation))
plt.show()
# The results aren't conclusive, but if you look closely at the scatter plots for **temp** and **atemp**, you can see a vague diagonal trend showing that higher rental counts tend to coincide with higher temperatures; and a correlation value of just over 0.5 for both of these features supports this observation. Conversely, the plots for **hum** and **windspeed** show a slightly negative correlation, indicating that there are fewer rentals on days with high humidity or windspeed.
#
# Now let's compare the categorical features to the label. We'll do this by creating box plots that show the distribution of rental counts for each category.
# plot a boxplot for the label by each categorical feature
for col in categorical_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
bike_data.boxplot(column = 'rentals', by = col, ax = ax)
ax.set_title('Label by ' + col)
ax.set_ylabel("Bike Rentals")
plt.show()
# The plots show some variance in the relationship between some category values and rentals. For example, there's a clear difference in the distribution of rentals on weekends (**weekday** 0 or 6) and those during the working week (**weekday** 1 to 5). Similarly, there are notable differences for **holiday** and **workingday** categories. There's a noticeable trend that shows different rental distributions in summer and fall months compared to spring and winter months. The **weathersit** category also seems to make a difference in rental distribution. The **day** feature we created for the day of the month shows little variation, indicating that it's probably not predictive of the number of rentals.
# ## Train a Regression Model
#
# Now that we've explored the data, it's time to use it to train a regression model that uses the features we've identified as potentially predictive to predict the **rentals** label. The first thing we need to do is to separate the features we want to use to train the model from the label we want it to predict.
# Separate features and labels
X, y = bike_data[['season','mnth', 'holiday','weekday','workingday','weathersit','temp', 'atemp', 'hum', 'windspeed']].values, bike_data['rentals'].values
print('Features:',X[:10], '\nLabels:', y[:10], sep='\n')
# After separating the dataset, we now have numpy arrays named **X** containing the features, and **y** containing the labels.
#
# We *could* train a model using all of the data; but it's common practice in supervised learning to split the data into two subsets; a (typically larger) set with which to train the model, and a smaller "hold-back" set with which to validate the trained model. This enables us to evaluate how well the model performs when used with the validation dataset by comparing the predicted labels to the known labels. It's important to split the data *randomly* (rather than say, taking the first 70% of the data for training and keeping the rest for validation). This helps ensure that the two subsets of data are statistically comparable (so we validate the model with data that has a similar statistical distribution to the data on which it was trained).
#
# To randomly split the data, we'll use the **train_test_split** function in the **scikit-learn** library. This library is one of the most widely used machine learning packages for Python.
# +
from sklearn.model_selection import train_test_split
# Split data 70%-30% into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
print ('Training Set: %d rows\nTest Set: %d rows' % (X_train.shape[0], X_test.shape[0]))
# -
# Now we have the following four datasets:
#
# - **X_train**: The feature values we'll use to train the model
# - **y_train**: The corresponding labels we'll use to train the model
# - **X_test**: The feature values we'll use to validate the model
# - **y_test**: The corresponding labels we'll use to validate the model
#
# Now we're ready to train a model by fitting a suitable regression algorithm to the training data. We'll use a *linear regression* algorithm, a common starting point for regression that works by trying to find a linear relationship between the *X* values and the *y* label. The resulting model is a function that conceptually defines a line where every possible X and y value combination intersect.
#
# In Scikit-Learn, training algorithms are encapsulated in *estimators*, and in this case we'll use the **LinearRegression** estimator to train a linear regression model.
# +
# Train the model
from sklearn.linear_model import LinearRegression
# Fit a linear regression model on the training set
model = LinearRegression().fit(X_train, y_train)
print (model)
# -
# ### Evaluate the Trained Model
#
# Now that we've trained the model, we can use it to predict rental counts for the features we held back in our validation dataset. Then we can compare these predictions to the actual label values to evaluate how well (or not!) the model is working.
# +
import numpy as np
predictions = model.predict(X_test)
np.set_printoptions(suppress=True)
print('Predicted labels: ', np.round(predictions)[:10])
print('Actual labels : ' ,y_test[:10])
# -
# Comparing each prediction with its corresponding "ground truth" actual value isn't a very efficient way to determine how well the model is predicting. Let's see if we can get a better indication by visualizing a scatter plot that compares the predictions to the actual labels. We'll also overlay a trend line to get a general sense for how well the predicted labels align with the true labels.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# There's a definite diagonal trend, and the intersections of the predicted and actual values are generally following the path of the trend line; but there's a fair amount of difference between the ideal function represented by the line and the results. This variance represents the *residuals* of the model - in other words, the difference between the label predicted when the model applies the coefficients it learned during training to the validation data, and the actual value of the validation label. These residuals when evaluated from the validation data indicate the expected level of *error* when the model is used with new data for which the label is unknown.
#
# You can quantify the residuals by calculating a number of commonly used evaluation metrics. We'll focus on the following three:
#
# - **Mean Square Error (MSE)**: The mean of the squared differences between predicted and actual values. This yields a relative metric in which the smaller the value, the better the fit of the model
# - **Root Mean Square Error (RMSE)**: The square root of the MSE. This yields an absolute metric in the same unit as the label (in this case, numbers of rentals). The smaller the value, the better the model (in a simplistic sense, it represents the average number of rentals by which the predictions are wrong!)
# - **Coefficient of Determination (usually known as *R-squared* or R<sup>2</sup>**: A relative metric in which the higher the value, the better the fit of the model. In essence, this metric represents how much of the variance between predicted and actual label values the model is able to explain.
#
# > **Note**: You can find out more about these and other metrics for evaluating regression models in the [Scikit-Learn documentation](https://scikit-learn.org/stable/modules/model_evaluation.html#regression-metrics)
#
# Let's use Scikit-Learn to calculate these metrics for our model, based on the predictions it generated for the validation data.
# +
from sklearn.metrics import mean_squared_error, r2_score
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# -
# So now we've quantified the ability of our model to predict the number of rentals. It definitely has *some* predictive power, but we can probably do better!
#
# ## Experiment with Algorithms
#
# The linear regression algorithm we used to train the model has some predictive capability, but there are many kinds of regression algorithm we could try, including:
#
# - **Linear algorithms**: Not just the Linear Regression algorithm we used above (which is technically an *Ordinary Least Squares* algorithm), but other variants such as *Lasso* and *Ridge*.
# - **Tree-based algorithms**: Algorithms that build a decision tree to reach a prediction.
# - **Ensemble algorithms**: Algorithms that combine the outputs of multiple base algorithms to improve generalizability.
#
# > **Note**: For a full list of Scikit-Learn estimators that encapsulate algorithms for supervised machine learning, see the [Scikit-Learn documentation](https://scikit-learn.org/stable/supervised_learning.html). There are many algorithms to choose from, but for most real-world scenarios, the [Scikit-Learn estimator cheat sheet](https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html) can help you find a suitable starting point.
#
# ### Try Another Linear Algorithm
#
# Let's try training our regression model by using a **Lasso** algorithm. We can do this by just changing the estimator in the training code.
# +
from sklearn.linear_model import Lasso
# Fit a lasso model on the training set
model = Lasso().fit(X_train, y_train)
print (model, "\n")
# Evaluate the model using the test data
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# ### Try a Decision Tree Algorithm
#
# As an alternative to a linear model, there's a category of algorithms for machine learning that uses a tree-based approach in which the features in the dataset are examined in a series of evaluations, each of which results in a *branch* in a *decision tree* based on the feature value. At the end of each series of branches are leaf-nodes with the predicted label value based on the feature values.
#
# It's easiest to see how this works with an example. Let's train a Decision Tree regression model using the bike rental data. After training the model, the code below will print the model definition and a text representation of the tree it uses to predict label values.
# +
from sklearn.tree import DecisionTreeRegressor
from sklearn.tree import export_text
# Train the model
model = DecisionTreeRegressor().fit(X_train, y_train)
print (model, "\n")
# Visualize the model tree
tree = export_text(model)
print(tree)
# -
# So now we have a tree-based model; but is it any good? Let's evaluate it with the test data.
# +
# Evaluate the model using the test data
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# The tree-based model doesn't seem to have improved over the linear model, so what else could we try?
#
# ### Try an Ensemble Algorithm
#
# Ensemble algorithms work by combining multiple base estimators to produce an optimal model, either by applying an aggregate function to a collection of base models (sometimes referred to a *bagging*) or by building a sequence of models that build on one another to improve predictive performance (referred to as *boosting*).
#
# For example, let's try a Random Forest model, which applies an averaging function to multiple Decision Tree models for a better overall model.
# +
from sklearn.ensemble import RandomForestRegressor
# Train the model
model = RandomForestRegressor().fit(X_train, y_train)
print (model, "\n")
# Evaluate the model using the test data
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# For good measure, let's also try a *boosting* ensemble algorithm. We'll use a Gradient Boosting estimator, which like a Random Forest algorithm builds multiple trees, but instead of building them all independently and taking the average result, each tree is built on the outputs of the previous one in an attempt to incrementally reduce the *loss* (error) in the model.
# +
# Train the model
from sklearn.ensemble import GradientBoostingRegressor
# Fit a lasso model on the training set
model = GradientBoostingRegressor().fit(X_train, y_train)
print (model, "\n")
# Evaluate the model using the test data
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# ## Optimize Hyperparameters
#
# Take a look at the **GradientBoostingRegressor** estimator definition in the output above, and note that it, like the other estimators we tried previously, includes a large number of parameters that control the way the model is trained. In machine learning, the term *parameters* refers to values that can be determined from data; values that you specify to affect the behavior of a training algorithm are more correctly referred to as *hyperparameters*.
#
# The specific hyperparameters for an estimator vary based on the algorithm that the estimator encapsulates. In the case of the **GradientBoostingRegressor** estimator, the algorithm is an ensemble that combines multiple decision trees to create an overall predictive model. You can learn about the hyperparameters for this estimator in the [Scikit-Learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingRegressor.html).
#
# We won't go into the details of each hyperparameter here, but they work together to affect the way the algorithm trains a model. In many cases, the default values provided by Scikit-Learn will work well; but there may be some advantage in modifying hyperparameters to get better predictive performance or reduce training time.
#
# So how do you know what hyperparameter values you should use? Well, in the absence of a deep understanding of how the underlying algorithm works, you'll need to experiment. Fortunately, SciKit-Learn provides a way to *tune* hyperparameters by trying multiple combinations and finding the best result for a given performance metric.
#
# Let's try using a *grid search* approach to try combinations from a grid of possible values for the **learning_rate** and **n_estimators** hyperparameters of the **GradientBoostingRegressor** estimator.
# +
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer, r2_score
# Use a Gradient Boosting algorithm
alg = GradientBoostingRegressor()
# Try these hyperparameter values
params = {
'learning_rate': [0.1, 0.5, 1.0],
'n_estimators' : [50, 100, 150]
}
# Find the best hyperparameter combination to optimize the R2 metric
score = make_scorer(r2_score)
gridsearch = GridSearchCV(alg, params, scoring=score, cv=3, return_train_score=True)
gridsearch.fit(X_train, y_train)
print("Best parameter combination:", gridsearch.best_params_, "\n")
# Get the best model
model=gridsearch.best_estimator_
print(model, "\n")
# Evaluate the model using the test data
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
# overlay the regression line
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# > **Note**: The use of random values in the Gradient Boosting algorithm results in slightly different metrics each time. In this case, the best model produced by hyperparameter tuning is unlikely to be significantly better than one trained with the default hyperparameter values; but it's still useful to know about the hyperparameter tuning technique!
#
# ## Preprocess the Data
#
# We trained a model with data that was loaded straight from a source file, with only moderately successful results.
#
# In practice, it's common to perform some preprocessing of the data to make it easier for the algorithm to fit a model to it. There's a huge range of preprocessing transformations you can perform to get your data ready for modeling, but we'll limit ourselves to a few common techniques:
#
# ### Scaling numeric features
#
# Normalizing numeric features so they're on the same scale prevents features with large values from producing coefficients that disproportionately affect the predictions. For example, suppose your data includes the following numeric features:
#
# | A | B | C |
# | - | --- | --- |
# | 3 | 480 | 65 |
#
# Normalizing these features to the same scale may result in the following values (assuming A contains values from 0 to 10, B contains values from 0 to 1000, and C contains values from 0 to 100):
#
# | A | B | C |
# | -- | --- | --- |
# | 0.3 | 0.48| 0.65|
#
# There are multiple ways you can scale numeric data, such as calculating the minimum and maximum values for each column and assigning a proportional value between 0 and 1, or by using the mean and standard deviation of a normally distributed variable to maintain the same *spread* of values on a different scale.
#
# ### Encoding categorical variables
#
# Machine learning models work best with numeric features rather than text values, so you generally need to convert categorical features into numeric representations. For example, suppose your data includes the following categorical feature.
#
# | Size |
# | ---- |
# | S |
# | M |
# | L |
#
# You can apply *ordinal encoding* to substitute a unique integer value for each category, like this:
#
# | Size |
# | ---- |
# | 0 |
# | 1 |
# | 2 |
#
# Another common technique is to use *one hot encoding* to create individual binary (0 or 1) features for each possible category value. For example, you could use one-hot encoding to translate the possible categories into binary columns like this:
#
# | Size_S | Size_M | Size_L |
# | ------- | -------- | -------- |
# | 1 | 0 | 0 |
# | 0 | 1 | 0 |
# | 0 | 0 | 1 |
#
# To apply these preprocessing transformations to the bike rental, we'll make use of a Scikit-Learn feature named *pipelines*. These enable us to define a set of preprocessing steps that end with an algorithm. You can then fit the entire pipeline to the data, so that the model encapsulates all of the preprocessing steps as well as the regression algorithm. This is useful, because when we want to use the model to predict values from new data, we need to apply the same transformations (based on the same statistical distributions and category encodings used with the training data).
#
# >**Note**: The term *pipeline* is used extensively in machine learning, often to mean very different things! In this context, we're using it to refer to pipeline objects in Scikit-Learn, but you may see it used elsewhere to mean something else.
# +
# Train the model
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.linear_model import LinearRegression
import numpy as np
# Define preprocessing for numeric columns (scale them)
numeric_features = [6,7,8,9]
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
# Define preprocessing for categorical features (encode them)
categorical_features = [0,1,2,3,4,5]
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
# Combine preprocessing steps
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('regressor', GradientBoostingRegressor())])
# fit the pipeline to train a linear regression model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# -
# OK, the model is trained, including the preprocessing steps. Let's see how it performs with the validation data.
# +
# Get predictions
predictions = model.predict(X_test)
# Display metrics
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# The pipeline is composed of the transformations and the algorithm used to train the model. To try an alternative algorithm you can just change that step to a different kind of estimator.
# +
# Use a different estimator in the pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('regressor', RandomForestRegressor())])
# fit the pipeline to train a linear regression model on the training set
model = pipeline.fit(X_train, (y_train))
print (model, "\n")
# Get predictions
predictions = model.predict(X_test)
# Display metrics
mse = mean_squared_error(y_test, predictions)
print("MSE:", mse)
rmse = np.sqrt(mse)
print("RMSE:", rmse)
r2 = r2_score(y_test, predictions)
print("R2:", r2)
# Plot predicted vs actual
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Daily Bike Share Predictions - Preprocessed')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test,p(y_test), color='magenta')
plt.show()
# -
# We've now seen a number of common techniques used to train predictive models for regression. In a real project, you'd likely try a few more algorithms, hyperparameters, and preprocessing transformations; but by now you should have got the general idea. Let's explore how you can use the trained model with new data.
#
# ### Use the Trained Model
#
# First, let's save the model.
# +
import joblib
# Save the model as a pickle file
filename = './models/bike-share.pkl'
joblib.dump(model, filename)
# -
# Now, we can load it whenever we need it, and use it to predict labels for new data. This is often called *scoring* or *inferencing*.
# +
# Load the model from the file
loaded_model = joblib.load(filename)
# Create a numpy array containing a new observation (for example tomorrow's seasonal and weather forecast information)
X_new = np.array([[1,1,0,3,1,1,0.226957,0.22927,0.436957,0.1869]]).astype('float64')
print ('New sample: {}'.format(list(X_new[0])))
# Use the model to predict tomorrow's rentals
result = loaded_model.predict(X_new)
print('Prediction: {:.0f} rentals'.format(np.round(result[0])))
# -
# The model's **predict** method accepts an array of observations, so you can use it to generate multiple predictions as a batch. For example, suppose you have a weather forecast for the next five days; you could use the model to predict bike rentals for each day based on the expected weather conditions.
# +
# An array of features based on five-day weather forecast
X_new = np.array([[0,1,1,0,0,1,0.344167,0.363625,0.805833,0.160446],
[0,1,0,1,0,1,0.363478,0.353739,0.696087,0.248539],
[0,1,0,2,0,1,0.196364,0.189405,0.437273,0.248309],
[0,1,0,3,0,1,0.2,0.212122,0.590435,0.160296],
[0,1,0,4,0,1,0.226957,0.22927,0.436957,0.1869]])
# Use the model to predict rentals
results = loaded_model.predict(X_new)
print('5-day rental predictions:')
for prediction in results:
print(np.round(prediction))
# -
# ## Further Reading
#
# To learn more about Scikit-Learn, see the [Scikit-Learn documentation](https://scikit-learn.org/stable/user_guide.html).
#
# ## Challenge: Predict Real Estate Prices
#
# Think you're ready to create your own regression model? Try the challenge of predicting real estate property prices in the [/challenges/02 - Real Estate Regression Challenge.ipynb](./challenges/02%20-%20Real%20Estate%20Regression%20Challenge.ipynb) notebook!
#
# > **Note**: The time to complete this optional challenge is not included in the estimated time for this exercise - you can spend as little or as much time on it as you like!
| .ipynb_checkpoints/02 - Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# %matplotlib inline
import re
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import accuracy_score
from sklearn.multiclass import OneVsRestClassifier
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
from sklearn.svm import LinearSVC
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
import seaborn as sns
# -
df = pd.read_csv("../input/comment/1613373921-5e748a2d5fc288e9f69c5f86.csv")
label_cols = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
df['none'] = 1-df[label_cols].max(axis=1)
df.describe()
df_toxic = df.drop(['id', 'comment_text'], axis=1)
counts = []
categories = list(df_toxic.columns.values)
for i in categories:
counts.append((i, df_toxic[i].sum()))
df_stats = pd.DataFrame(counts, columns=['category', 'number_of_comments'])
df_stats
df_stats.plot(x='category', y='number_of_comments', kind='bar', legend=False, grid=True, figsize=(8, 5))
plt.title("Number of comments per category")
plt.ylabel('# of Occurrences', fontsize=12)
plt.xlabel('category', fontsize=12)
# +
rowsums = df.iloc[:,2:].sum(axis=1)
x=rowsums.value_counts()
#plot
plt.figure(figsize=(8,5))
ax = sns.barplot(x.index, x.values)
plt.title("Multiple categories per comment")
plt.ylabel('# of Occurrences', fontsize=12)
plt.xlabel('# of categories', fontsize=12)
# -
lens = df.comment_text.str.len()
lens.hist(bins = np.arange(0,5000,50))
print('Number of missing comments in comment text:')
df['comment_text'].isnull().sum()
df['comment_text'][0]
categories = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate','none']
# +
def clean_text(text):
text = text.lower()
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "can not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub(r"\'scuse", " excuse ", text)
text = re.sub('\W', ' ', text)
text = re.sub('\s+', ' ', text)
text = text.strip(' ')
return text
# -
df['comment_text'] = df['comment_text'].map(lambda com : clean_text(com))
df['comment_text'][0]
train, test = train_test_split(df, random_state=42, test_size=0.20, shuffle=True)
X_train = train.comment_text
X_test = test.comment_text
print(X_train.shape)
print(X_test.shape)
NB_pipeline = Pipeline([
('tfidf', TfidfVectorizer(stop_words=stop_words)),
('clf', OneVsRestClassifier(MultinomialNB(
fit_prior=True, class_prior=None))),
])
for category in categories:
print('... Processing {}'.format(category))
# train the model using X_dtm & y
NB_pipeline.fit(X_train, train[category])
# compute the testing accuracy
prediction = NB_pipeline.predict(X_test)
print('Test accuracy is {}'.format(accuracy_score(test[category], prediction)))
# +
SVC_pipeline = Pipeline([
('tfidf', TfidfVectorizer(stop_words=stop_words)),
('clf', OneVsRestClassifier(LinearSVC(), n_jobs=1)),
])
# -
for category in categories:
print('... Processing {}'.format(category))
# train the model using X_dtm & y
SVC_pipeline.fit(X_train, train[category])
# compute the testing accuracy
prediction = SVC_pipeline.predict(X_test)
print('Test accuracy is {}'.format(accuracy_score(test[category], prediction)))
LogReg_pipeline = Pipeline([
('tfidf', TfidfVectorizer(stop_words=stop_words)),
('clf', OneVsRestClassifier(LogisticRegression(solver='sag'), n_jobs=1)),
])
for category in categories:
print('... Processing {}'.format(category))
# train the model using X_dtm & y
LogReg_pipeline.fit(X_train, train[category])
# compute the testing accuracy
prediction = LogReg_pipeline.predict(X_test)
print('Test accuracy is {}'.format(accuracy_score(test[category], prediction)))
| one-vs-rest-ml-approach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="tOVbsNcpcVPI"
# # **ESS314: Lab5: Earthquake Location**
#
# we will be using two different methods (triangulation and grid search) to find the location of an earthquake epicenter.
#
# Distance $d$, time $t$, and velocity $v$ are simply related by
#
# $d = vt$. (1)
#
# Therefore, a P-wave traveling for time $t$ at velocity $V_P$ will move across a distance $r$, where $r$ is given by
#
# $𝑟 = 𝑉_Pt$. (2)
#
# 
#
# The time necessary for the P-wave to travel a distance r can be obtained by dividing equation (2) by $V_P$:
#
# $𝑡 = 𝑟 /𝑉_P$ (3)
#
# 
#
# If a P-wave is generated from an earthquake (EQ) at the origin time $𝑇_0$ (the time the earthquake starts), the first wiggle on the seismogram will arrive at the observation station labeled “seismometer 1”, at a distance $r_1$ from the earthquake, at the time $T_0$ given by
#
# $T_p^1 = T_0 + r_1/V_P$ (4)
#
# where the superscript 1 indicates the seismometer.
#
# In equation (4), the first term on the right-hand side (RHS) is the start time of the EQ, and the second term on the RHS is the time it takes the P-wave ray to reach the seismometer located at
# a distance $r_0$ from the EQ.
#
# Similarly, an S-wave from the same earthquake traveling at a velocity $V_S$ will arrive at seismometer 1 at time $T_S^1$, given by
#
# $T_s^1 = T_0 + r_1/V_S$ (5)
#
#
# Since S-waves travel more slowly than P-waves, $T_S^1 > T_P^1$.
#
# Now we have a problem. Equations (4) and (5) both contain the unknown origin time $T_0$ . However, if we subtract equation (4) from equation (5) we eliminate $T_0$ and obtain the following very useful result:
#
# $T_S^1 - T_P^1 = r_1/V_S - t_1/V_P = r_1 (1/V_S - 1/V_P)$ (6)
#
#
# Equation (6) defines the S-P time (“S minus P time”) for the event as recorded at seismometer 1: $T_{s-p} = T_S^1 - T_P^1$. We will use $T_{s-p}^1$ to represent the S-P time for this event at station 1.
#
# 
#
#
# The S-P time depends on the distance from the earthquake to the seismic station and the relative times between the P and S wave, which can be variable based on 3D earth structure.
#
# As part of Lab 2: you calculated P and S wave traveltime curves for an earthquake located 10 km below the surface. Your plot should have looked something like this:
#
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="x9AoOy1eitWb" outputId="920fbc66-716c-4424-aacb-15ca2e199911"
import matplotlib.pyplot as plt
import numpy as np
r = np.linspace(0,50,50) # km
vp = 5 # km/s, this is 5000 m/s
vs = 3.5 # km/s, this is 3500 m/s
tp = r/vp
ts = r/vs
fig = plt.plot(r,tp, 'b-',label='P')
fig = plt.plot(r,ts, 'r-', label='S')
plt.xlabel('Epicentral Distance (km)')
plt.ylabel('Travel Time to Seismometer (sec)')
plt.title("Earthquake Travel Times")
plt.legend(loc=1)
plt.grid(True)
plt.show()
# + [markdown] id="kgKPdTowcVM4"
# ### **Question 1:**
#
# > a. *What happens to the S-P travel time as you get farther away from the epicenter?*
# + [markdown] id="PUCklfpScVKw"
# answer here:
# + [markdown] id="k0Rl6ojNcVIe"
# > b. *An earthquake is recorded at two seismic stations, station A and station B. If station A is 10 km from the epicenter (point on Earth’s surface directly above where the earthquake rupture began), and station B is 100 km from the epicenter, which station will exhibit a larger S-P time?*
# + [markdown] id="NAuuBBx4cVFo"
# answer here:
# + [markdown] id="f6ic9iENcVD6"
# Comparing equations (3) and (6), you may notice that the time separation between the **P-wave arrival** (peak on the seismogram corresponding to the first arrival of a P-wave at the seismometer) and the **S-wave arrival** (peak on the seismogram corresponding to the first arrival of an S-wave at the seismometer) has an effective velocity by which it increases with distance:
#
# $V_{s-p} = \dfrac{1}{1/V_S - 1/V_P}$ . (7)
#
# The quantity $V_{s-p}$ is called the **effective S-P velocity**.
#
# Therefore, the distance r1 from seismometer 1 to the earthquake can be computed from seismograms (P-wave and S-wave arrival times) as
#
# $r_1 = V_{s-p}T^1_{s-p}$, (8)
#
# where $ T^1_{s-p} $ is measured on a seismogram, but $ V_{s-p} $ is based on knowledge of Earth properties, as values for both $V_P$ and $V_S$ are necessary to make this calculation (eqn 7). In other words, we need to have some idea of the P-wave and S-wave velocities in the Earth.
#
# We are going to download data from the [IRIS DMC](https://ds.iris.edu/ds/nodes/dmc/) data center. The data was collected from the [Pacific Northwest Seismic Network](https://pnsn.org/). We focus on the M 7.8 October 27, 2012 Haida Gwaii earthquake.
#
# + colab={"base_uri": "https://localhost:8080/"} id="Og5r_zu_nnLK" outputId="4fcaa2a5-91d2-4cd3-c53e-357d2e51bf87"
import obspy
from obspy.clients.fdsn import Client
from obspy import UTCDateTime
# create the link between you and the IRIS-DMC
client = Client('IRIS')
# Origin time of the Nisqually earthquake
s1 = obspy.UTCDateTime(2001,2,28,18,54,37)
# collect the information about what data is available
inv = client.get_stations(network="UW",station="*",channel="BHZ",location="*",starttime=s1,endtime=s1+86400)
print(inv)
# create a list of the stations you found
sta=list()
for k in inv:
for sta1 in k:
sta.append(sta1.code)
# print the list of the stations we would be using
print("The stations we will be using are:")
print(sta)
# -
# Mapping in python requires a few more packages, so we found a map of where these stations are on the IRIS-GMAP tool https://ds.iris.edu/gmap and annotated the stations.
#
# 
#
# marine will add a legend for a scale of distance.
# + colab={"base_uri": "https://localhost:8080/"} id="Zx8lE9DKqcjc" outputId="631be77f-cf76-443a-c327-dd83313006c9"
# download data:
tr=obspy.Stream()
# we will update sta to make sure we have all of the stations
# create a list of the stations you found
sta1=list()
# s1 = obspy.UTCDateTime(2012,10,28,3,4,0) # this is the Haida Gwaii M7.8 2012 earthquake.
# s1 = obspy.UTCDateTime(2001,2,28,18,54,37)
for ista in sta:
try:
tr += client.get_waveforms(network="UW",station=ista,channel="*HZ",location="*",starttime=s1,endtime=s1+100)
sta1.append(tr[-1].stats.station)
print(tr[-1].stats.station)
except:
pass
# we actually have fewer stations
print(sta1)
sta=sta1
# +
# we can use matplotlib
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams["figure.figsize"] = (8,6) #bigger size of figures as defaults
####
itr=0 # you can change and make it range from 0 to len(tr)-1
###
# create a time vector of 60 seconds (1 minute), sampled at the sampling rate of the data.
t=np.linspace(0,60,int(60/tr[itr].stats.delta))
# plot the data
plt.plot(t,tr[itr].data[:len(t)]/np.max(np.abs(tr[itr].data)),'k') # normalized data
plt.title(tr[itr].stats.station)
plt.xlabel('Time in (s)')
plt.grid(True)
# zoom in on specific time
plt.xlim([0,20])
# plt.xlim([20,100])
# -
# Now we can plot each of the data one by one. ``tr`` is an **array** of seismic ***streams***. ***Streams*** are basically **Obspy** objects that combine seismic station information and the time series. We can *iterate* through ``tr`` to plot each waveform. We can also zoom in on specific times to focus on different part of the waveforms.
print("there are "+str(len(tr))+" seismograms from these stations") # here we use "str" to convert an integer (len) into a string
# Now we are going to see how the overall waves compare to each others.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams["figure.figsize"] = (18,12)
for ii,itr in enumerate(tr):
if itr.stats.station == "SEP":continue # this station "clipped", meaning that the shaking was too intense for the instrument
t=np.linspace(0,60,int(60/itr.stats.delta))
if len(itr.data)<len(t):continue
crap=itr.copy()
crap = crap.filter('bandpass',freqmin=1,freqmax=10) # you can play with that.
# plot
plt.plot(t,crap.data[:len(t)]/np.max(np.abs(crap.data))+ii,'k')
plt.text(5,ii+0.3,itr.stats.station,fontsize=14,c='r')
plt.grid(True)
plt.xlim([0,100])
# plt.xticks(np.linspace(0,60,60))
plt.xlabel('Time in s')
plt.savefig('waveforms.png')
# + [markdown] id="bEWu4CE0nnT7"
#
#
# The figure above shows seismograms (data from seismometers) recording ground motion from the same earthquake at 5 different stations. A few things to note about this data:
#
# * The vertical axis on the plot is the amplitude of ground motion. Individual seismograms or traces are scaled differently, and actually normalized.
#
# * The waveforms may look jaggeted and not pure smooth oscillations. This is because this earthquake shaking was really, really strong and even damaged the soil. This was a very strongly felt earthquake with extensive damage.
#
# * There is a high-frequency signal first, that is the P. The lower frequency that arrives next is likely the S wave.
#
#
#
#
# -
# ## Earthquake Location using triangulation
# + [markdown] id="MFCKMcIvcVBS"
# ## **Question 2**:
# For this question, assume the earthquake is at the surface, so depth is not a factor.
#
# > a. *The plot of all of the waveforms is saved into a waveforms.png file. Open the file and annotate it with what you think the P and S waves are on these seismograms. Save the file into another figure.*
#
# Add the code line to import your figure within the notebook using the command: ````
# -
# Import the figure here
# + [markdown] id="f-5oceDJcU-1"
# > b. *Measure the S-P times for each station and list below in text of python variables. There will be 2 times 5 measurements, a total of 10 measurements to find.* You can pick an approximate time on the plot, or use an annotation to your previous figure using circles in Preview or other. Another way is to plot individual traces as shown above and "zoom in" by setting the command ``plt.xlim([t_1,t_2])``, where ``t_1`` and ``t_2`` are times of your choice.
# + [markdown] id="vyxB-I8pcU8V"
# > c. *Suppose $V_P$ = 6.0 km/s and $V_S$ = 3.47 km/s. These velocities are based on properties of the Earth’s crust. Use equation (7) to compute the effective $V_{s-p}$.*
# + id="gXnN53GQ31PT"
# + [markdown] id="OVCEPsypcU3a"
#
# > d) *Use equation (8), your computed value of $V_{s-p}$, and the S-P times that you measured for each station to populate the r column in the table below, where r is the distance from each station to the EQ that caused the ground motion recorded in the seismograms.*
# -
print(sta)
# note that LON and SEP are missing for some reason
r_gnw =
# + [markdown] id="LxW3w-h4cUvI"
#
# + [markdown] id="MahSCrQP5pzZ"
# ### **Question 3:**
# > a. *On the map provided, annotate with circles around each station with a radius corresponding to the r that you computed for that station.*
#
# The earthquake epicenter is where all the circles (or the most) intersect. Mark this single point with a star. This method of locating the earthquake epicenter is called triangulation. Note that you only have data for some of the stations on the map.
#
# + [markdown] id="thRYuAWE52i1"
# > b. *Give at least two reasons why the circles may not exactly cross at a single point. Don't cite poor drawing – think about the myriad sources of error throughout this process.*
# + [markdown] id="KgOmFwcz56yD"
# answer below:
# + [markdown] id="Yoa9C2gn573X"
# > c. *What is the minimum number of S-P times needed to find the epicenter of an earthquake? Draw a diagram and explain in words why your answer is correct.*
# + [markdown] id="fV_zhIuO6A6K"
# ## Earthquake Location using grid search algorithm
# + [markdown] id="6f5AWqSS6EQl"
#
#
# Now we are going to locate an earthquake in a different way. The difference between the observed travel time, or the travel time observed from seismic data, and the predicted travel time, or the travel time predicted from a model, is called the travel time residual. One method
# to find the location is to calculate the predicted travel times on a grid and find the grid point with the smallest residual. This is what we will do today.
#
# The predicted travel time from an earthquake at ($x,y$) to the k$^{th}$ station with coordinates ($x_k,y_k$)
#
# $t^p_k = \dfrac{\sqrt{ (x-x_k)^2 + (y-y_k)^2 }}{v}$, (9)
#
# where $v$ is the appropriate seismic velocity. Note that the superscript p here is not a power, but just denotes the predicted travel time.
#
# You can also use the seismogram recorded at the k$^{th}$ station to find the real, or, **observed travel time**. One way to locate an earthquake is to minimize the difference between the travel times predicted from the model, tp, and observed travel times, to. The expression for the travel time residual at the kth station is
#
# $r_k = t_k^0 - t_k^p$ (10)
#
# where $t_p^0$ is the observed travel time at station k and $t_k^p$ is the predicted travel time at station k.
#
# We are going to minimize the residuals for a whole network of stations using the method of **least squares**. Here we will define the **misfit** of a certain grid point as the sum of the squared residuals of each station
#
# $\epsilon = \sum_{k=1}^n r_k^2$, (11)
#
# where $n$ is the number of stations.
#
# Our goal is to find the grid point with minimum misfit. We will do this by assuming an earthquake location (x,y) and calculating the misfit 𝜖 for that location. We do this for many trial locations, and then find the location that fits the data the best.
#
# The table below has the location (x and y coordinates) of 13 stations, as well as the observed P wave travel times at each station.
#
# We are going to read the travel-time data from the CSV file "travel_times.csv" using Pandas. Pandas are a wonderful open-source python package to handle structured (tables) data.
# + id="M3oumltIcUGR"
import pandas as pd
pd = pd.read_csv("travel_times.csv")
print(pd)
# et voila!
# this reads the CSV file (a table) into a Pandas "data frame"
# -
# Pandas data frames are great.
pd.head() # shows the top 5 rows
pd.describe() # runs some basic stats on the data in the file.
pd['x'] # how to select the locations.
np.asarray(pd['x']) # convert the data frame column into a numpy array.
# ### **Question 4:**
#
# Assume the P wave velocity is 6 km/s. The steps outlined below will help you write a program in Python that uses this information to locate the earthquake using a grid search. Perform your grid search on a 100x100 km grid (10,000 locations).
#
# > 1. *Define your variables.*
#
# In the following, define: i) a P-wave velocity as a single scalar, ii) the ``x`` and ``y`` arrays of spatial coordinate, with values ranging from 1 to 100 with increment of 1, iii) convert the stations coordinates and travel times from the pands column into numpy arrays using the function above
# +
# type here is vp
# type here is the spatial x
# type here the spatial y
# type here the zero matrix
# -
# Now we will initialize an abnormally large value of what residuals could be. We will update this value within a loop to only find the minimum residual among all possible locations.
resmin = 1e9
# > 2. *Use nested FOR loops to go through the x and y coordinates of your grid (e.g., outer
# loop goes through the x coordinates, inner loop goes through the y coordinates).*
#
# With the for loop where we select ``i`` and ``j`` as possible index for ``x`` and ``y``, you can inquire each potential earthquake source location of x and y. Calculate for each the distance between your trial earthquake location and each source-station distance. For each potential earthquake location, please calculate:
#
# * Distance between the potential earthquake location and each station
# * Predict the travel time between the potential earthquake location and the station for P waves
# * Compute the residual ``res`` between the observation and the prediction (that is, the residual)
# * Sum all of the residuals over the 13 stations.
# * If that residual is less than ``resmin``, update ``resmin=res`` and save the value of the indices (or location as ``i_best`` and ``j_best``.
# * Save that scalar into the matrix ``res`` at that specific indices. You can use the numpy function ``np.sum()``.
#
# The best earthquake location is that found when the residuals are minimized. So where is the earthquake in the ``x`` and ``y`` space?
# answer here
| LABS/LAB5/.ipynb_checkpoints/Lab5-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.stats import norm
from scipy.stats import t
from scipy.stats import chi2
import seaborn as sns
import statsmodels.graphics.gofplots as sm
import scipy.stats as sp
import pandas as pd
import math
import sklearn as skl
import random as rand
import time
import matplotlib.pyplot as plt
import statistics as stats
#We are mostly interested in Frequentist Statistics. Bayesian Statistics is for another time
# -
# # Heights Experiment
# To get a good grasp on fundamentals, we are going to be working with randomly generated population distributions so I can finally distinguish between the concepts of Sample Distribution, Sampling Distribution, and Population Distribution and finally understand Inferential Statistics
rand.seed(1623362164)
pop = [round(rand.gauss(63, 3), 1) for i in range(600)]
#outliers
pop.extend([round(rand.gauss(50, 0.8), 1) for i in range(20)])
pop.extend([round(rand.gauss(72, 0.4), 1) for i in range(10)])
pop.sort()
print(pop)
# # Population Distribution
# Basically, this is impossible to know in almost all practical cases. This is here to just get an overall idea of the population distribution for comparison with the sampling and sample distributions
plt.hist(pop, bins=30)
# We see this population is roughly normal. In reality, it may very well not be,
# But that doesn't matter usually because of the Central Limit theorem applied to the Sampling Distribution
# Roughly speaking, say we have a single sample X drawn from this distribution. This means X can take on any
# value as drawn in this histogram, in roughly this frequency.
# Population Mean, standard deviation, variance
# Round to two decimals please
mu = round(stats.mean(pop), 2)
sigma_2 = round(stats.pvariance(pop), 2)
sigma = round(stats.pstdev(pop), 2)
print(mu, sigma, sigma_2)
# # Sampling Distribution
# This is the distribution of sample means. Basically, the random variable X_bar = (X_1 + X_2 + ... + X_N)/N
# will take on a roughly normal distribution due to Central Limit Theorem. Basically all of inferential stats
# depends on this distribution, and NOT the population distribution since we almost NEVER know the population
# distribution
N = 50
sampling_dist = [round(stats.mean(rand.sample(pop, k=N)), 2) for i in range(3000)]
plt.hist(sampling_dist, bins=20)
# A small sample from a normal distribution allows t-distribution to be applied
N_small = 15
sampling_dist = [round(stats.mean(rand.sample(pop, k=N_small)), 2) for i in range(3000)]
plt.hist(sampling_dist, bins=20)
# # Sample Distribution
# Now we're getting to the real statistics. Note: this is not the sampling distribution, as the sampling distribution is based on the mean of a sample for a certain number of samples N
# Get an SRS without replacement of the population
# This is used when you can't get any reads on population, otherwise, cluster or stratified sampling may be better
sample = rand.sample(pop, k=N)
small_sample = rand.sample(pop, k=N_small)
print(small_sample)
# ## Exploratory Statistics
# Basically, we want to investigate our population distribution, but we only have the sample distribution.
# Among the most important things we need to do is check visually/test for normality as that unlocks the
# powers of Gaussian/parametric statistics for us. If we are unable to assume normality, then we must turn
# to nonparametric methods
#
# Though most of this will be in the context of investigating normality, the visualizations are also
# extraordinarily important for general data visualization
# Descriptive Statistics/Data Exploration
# At a minimum:
# Graph histogram, get 5-number summary, sample mean, stdev, and mode. Maybe do a box plot
# As we have here, we have an outlier in our sample. Perfect. But there is a roughly normal looking distribution sample
plt.hist(sample, bins=9)
# +
def five_num_sum(sample):
Q = np.percentile(sample, [25, 75])
return [min(sample), Q[0], stats.median(sample), Q[1], max(sample)]
#five_num_sum(sample)
X_bar = round(stats.mean(sample), 2)
s = round(stats.stdev(sample), 2)
print(five_num_sum(sample), X_bar, s)
small_X_bar = round(stats.mean(small_sample), 2)
small_s = round(stats.stdev(small_sample), 2)
print(five_num_sum(small_sample), small_X_bar, small_s)
# -
# ### Tests for Normality
# There are two categories: visual and significance tests
# Both are recommended to be done: visual is more ad-hoc but gives greater understanding,
# significance testing is more rigorous.
# This and most of what follows is according to the paper:
# Normality Tests for Statistical Analysis: A Guide for Non-Statisticians
#
# First off, if we have N > 30 or 40, we have no fear of nonnormal population distributions, due to the CLT.
# If you have hundreds of observations, then the population can basically be ignored. According to the CLT,
# 1. If you have a normal population, then samples will be normal
# 2. In large samples of 30-40, you have roughly normal data regardless of the population.
# 3. Means from any distribution are normal
#
# #### Visual Methods
# Unreliable and doesn't guarantee normality, but can be quickly appreciated and understood by humans
# histograms, stem and leaf plot, boxplot, P-P plot, Q-Q plots are good for checking normality
# Scores greater than 1.5 times the interquartile range are out of the boxplot and are considered as outliers,
# and those greater than 3 times the interquartile range are extreme outliers. A boxplot that is symmetric with
# the median line at approximately the center of the box and with symmetric whiskers that are slightly longer
# than the subsections of the center box suggests that the data may have come from a normal distribution
plt.boxplot(sample, vert=False)
# As we see here, we do have roughly normal data,but we have not only outliers, but extreme outliers
# This is a pretty poor sample, but thanks to the size, it is usable. For a small sample, outliers
# and extremes pretty much ruin data.
fig, ax = plt.subplots(1, 3, figsize=(15, 4))
sns.histplot(sample,kde=True, color ='blue',ax=ax[0])
# Need to normalize data as (x - X_bar) / s for QQ plot
normalized_sample = [(x - X_bar) / s for x in sample]
sm.ProbPlot(np.asarray(normalized_sample)).qqplot(line='s', ax=ax[1])
cumulative_prob = [norm.cdf(x) for x in normalized_sample]
sm.ProbPlot(np.asarray(cumulative_prob)).ppplot(line='s', ax=ax[2])
plt.show()
# We compare the QQ plot against the PP plot, and we can see a difference in how to interpret it.
# Perhaps the PP plot is more obvious as to whether or not something is normal, but the QQ plot
# appears more understandable.
# #### Normal Significance Tests
#
# Many tests exist: Kolmogorov-Smirnov (K-S) test, Lilliefors corrected K-S test, Shapiro-Wilk test,
# Anderson-Darling test, Cramer-von Mises test, D’Agostino skewness test, Anscombe-Glynn kurtosis test,
# D’Agostino-Pearson omnibus test, and the Jarque-Bera test
# Shall compare K-S and Shapiro-Wilk as these two are the most popular, the former being the most popular.
# It is recommended to use Shapiro-Wilk as K-S is frequently low in power even with the Lilliefors correction
# the K-S test uses the emperical cumulative distribution, and compares the difference between either the
# actual cumulative distribution function, or another function's distribution function. It is a nonparametric
# test. But it is also low in power, so not recommended.
# It is an exact test, so the number of samples does not matter to its validity. But it does have limitations:
# 1. only applies to continuous distribution, 2. more sensitive in center than tails, 3. most serious limitation:
# requires a fully specified distribution (like one we know or another eCDF), which means it is not estimable.
# The statistic is basically the maximum positive difference between each distribution. The KS is most commonly
# used for normality, but can apply to any continuous distribution that we know
# null hypothesis is that the data follow the given distibution. The alternative is not
# source: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3693611/#:~:text=The%20main%20tests%20for%20the,test%20(7)%2C%20and%20the
sp.kstest(sample, lambda x: norm.cdf(x, loc=X_bar, scale=s))
# Shapiro-Wilk Test. It is biased by sample size, the larger it is, the more likely it will find a significant result.
# But it has more power than KS. It has some random formula, don't even worry about it because it's complicated and
# we mostly monte carlo the distribution anyways.
# The H_0 is the data came from a normal distribution
sp.shapiro(sample)
# Iis correct, the distribution is not normal technically, so I'm rather impresssed
# # Inferential Statistics
# Now we'll be doing the real cool stuff
# ## Interval Estimation (for mean)
# Confidence Intervals, do we have a value that makes sense here for the mean?
# The idea is that when you calculate a say 95% confidence interval, when you resample
# 100 times and calculate these 95% confidence intervals, about 95 of the samples'
# Confidence intervals shall contain the population mean
# Typically, we want confidence of either 90%, 95%, 99%, or 99.9%, or critical value
# alpha = 0.1. 0.05, 0.01, 0.001 respectively
# Now when estimating a point estimator interval that involves a sum like the mean,
# we can use the CLT to say that the distribution is normal and do all our Gaussian
# stuff. For N small, this assumption does not hold and we need to use the
# unique distribution obtained from summing together the samples or however they're
# combined. Of course, that's a bit hard if you don't even know the population
# distribution. But when you assume or know that the population distribution is normal,
# then we have a few things like Chi-squared distribution and the (Student's) t-distribution
#
# Now, the confidence interval we are working with is based on the distribution for
# the estimator, in this case the mean. Thus, with large N, we have a normal
# distribution and it basically boils down to finding z-scores until
# P(mu in [lower bound, higher bound]) >= 1 - alpha
#
# Note: the misunderstanding of confidence intervals is that there is a 95% chance the
# calculated interval contains the mean. This is INCORRECT. The 95% refers to the
# reliability of repeated experimentation, that about 95% of our confidence intervals
# shall have the true mean somewhere. It is wholly possible for the mean to be completely
# out of the confidence interval, as once it is calculated, the mean is either inside or
# out, it is not a matter of probability. Said another way, each confidence interval is
# like a trial. By Law of Large Numbers, it trends towards containing the true mean
# 95% percent of the time, but this is not the case for any individual trial.
alpha = 0.05
# X_bar ~ Normal(mu, sqrt(1/n)), so normalizing gets us (X_bar - mu)/(1/sqrt(n)) = sqrt(n) * (X_bar - mu) ~ N(0, 1)
# Now, you do Z-scores where the tails in total sum up to alpha, so each tail should contain alpha/2. From this
# we obtain the general formula: [X_bar - z_alpha/2 * sigma/sqrt(n), X_bar + z_alpha/2 * sigma/sqrt(n)].
# Of course, if we have the population stdev, this is easy, otherwise, replace sigma with sample stdev = s
# Here, we use s, as we usually don't know the stdev of all people's heights
# To obtain z_alpha/2, we have CDF^-1(1-alpha/2)
z = norm.ppf(1 - alpha / 2)
[X_bar - z * s / math.sqrt(N), X_bar + z * s / math.sqrt(N)]
# For comparison, this is with the small sample, but this one will actually be incorrect way more often
# because it is too narrow at such a small N
[small_X_bar - z * small_s / math.sqrt(N_small), small_X_bar + z * small_s / math.sqrt(N_small)]
# For N small, we must use Chi squared and t-distribution. These only apply for when
# the population distribution is about or is normal
# Now if your n is small, but you have the population stdev, you can use the above procedure. Else, use the t-distribution
# You have this pivot for the T-value: (X_bar - mu)/(s/sqrt(n)) ~ T(n - 1). You can then derive:
# mu in [X_bar - t_alpha/2,n-1 * s/sqrt(n), X_bar + t_alpha/2,n-1 * s/sqrt(n)]
T = t.ppf(1 - alpha / 2, N_small - 1)
[small_X_bar - T * small_s / math.sqrt(N_small), small_X_bar + T * small_s / math.sqrt(N_small)]
# For N small, variance is in particular strange. At large N, since variance is also a sum
# the CLT applies, but because it's hard to find variance of variance, it's annoying to do Gaussian pivots,
# so it's best to resort to the Chi-squared distribution. When N is small,
# variance is approximately the Chi Squared function.
# If you don't know both mu and stdev of population, you must use this for interval estimation,
# but even if you knew mu, you'd still have to use this.
# You get the pivot: (n-1)s^2/stdev^2 which gives you the formula
# [(n-1)s^2/chi-squared_1 - alpha/2,n-1, (n-1)s^2/chi-squared_alpha/2,n-1] as a confidence interval for
# variance
Y_upper = chi2.ppf(1 - alpha / 2, N_small - 1)
Y_lower = chi2.ppf(alpha/2, N_small - 1)
[(N_small-1) * small_s**2 / Y_upper, (N_small-1) * small_s**2 / Y_lower]
# ## Null Hypothesis Significance Testing
# Remember the null hypothesis is the one with the equality
#
# While this focuses on significance of a statistic to the mean, we can also do either paired or between
# individually random samples. For paired, its simple, you simply subtract values and use the variance of
# the difference. For individually random samples, the test statistic is
# (X_bar_1 - X_bar_2)/sqrt(s_1^2/n_1 + s_2^2/n_2). This is used in A/B testing. Two sample z-tests exist but
# the stdev formula is super biased, so t-tests are almost exclusively used for two samples. Of course, you
# need data that is roughly normal, although I'm sure at high enough samples, the two sample t-test is still
# valid. If you have more than 2 groups, ANOVA, Tukey-Kramer, ANOM, and Dunnett's test are all good options.
#
# If you don't have normal data, or your sample size is too small to have normality tests conducted on it,
# you may have to resort to nonparametric methods.
#
# Since in practice, the stdev of the population is unknown, the statistic is only distributed according to the
# t-distribution. The recommendation to use the normal on n > 30 only applied when it was difficult to perform
# t-tests by hand, but now, you should almost always use t-tests.
#
# Other things you can do: given a sample proportion p_hat, you can do a Z-test on the statistic:
# p_hat - p/ sqrt(p(1-p)/n). This is based on binomial distributions, so have np and n(1-p) > 10 respectively,
# to be able to use the approximate normal distribution. Otherwise, this does not really apply. SRS sampling
# is necessary, and it actually can't be more than 10% of the population size since that meses with independence
# (apparently some more advanced methods exist for this however).
#
# To use the T-distribution, on N >= 40, no assumptions need to be made about the population distribution.
# On 15 < N < 40, sample should show no extreme values and little, if any, skewness; or you can assume the
# population distribution is normal. For N <= 15, sample should contain no outliers and no skewness, or assume
# parent distribution is normal.
#
# Standard errors: of a proportion p_hat, we have SE(p_hat) = sqrt(p_hat (1-p_hat)/n), of a mean, SE(X_bar) = s /sqrt(n)
#
# ### Fisherian P-Value Method
# The idea here is to take your null hypothesis and see if you can reject it. When you take your test, you get to be able to say your results were statistically significant or not, but if it's not, you can't say that the null hypothesis is wrong, rather you don't have the evidence enough for it yet. This is arguably more used today, and is the p-value method in some people's textbooks. It can reject hypotheses by contradiction, but not prove/accept them without more evidence. H_0 is the hypothesis you wanna disprove. H_1 is your alternative hypothesis. In Fisher, you don't really need the alternative hypothesis or need to accept it, you can only say if you have a significant result or not. Usually, you say you fail to reject the null.
#
# Of course in Significance testing, you choose a cutoff alpha level, but the P-value is the lowest alpha for which you can reject your null hypothesis
# One-sided test (lower tail)
# H_0: mu >= mu_0 (remember we want to reject this)
# H_1: mu < mu_0
# Of course use S if you don't know population stdev. Mu_0, your hypothesized
# value is always a given
# The test statistic is:
# W = (X_bar - mu_0) / (S/sqrt(n))
# for large N (CLT applies), use the acceptance region W >= z_alpha (opposite for other sided tail)
# Else, if you have a small N, and your population is normal,
# do a t-test. (W <= t_alpha,n-1)
mu_0 = 62.68 # we have omniscience here, it's kinda a guess in real life
W = (X_bar - mu_0) / (s / math.sqrt(N))
p = norm.cdf(W)
print(p, alpha) # In this case, we do not have a significant result, we don't have the evidence to reject
# Two-sided test
# H_0: mu = mu_0 (remember we want to reject this)
# H_1: mu != mu_0
W = (X_bar - mu_0) / (s / math.sqrt(N))
p = 2 * norm.cdf(-abs(W))
print(p, alpha) # Again failed to reject. This is a good sign, our sample mean is pretty close.
# Under Fisherian assumptions however, this just means we can't say anything meaningful
# ### Neyman-Pearson (traditional) method
# In this case, when you reject your null hypothesis, you MUST accept the other hypothesis.
# In most intro statistics classes, these are hybridized, and so you accept the other hypothesis
# when the p-value rejects it, but this way is more clear. You choose your cutoff and then look at
# whether or not the test statistic is in the acceptance region. In this paradigm, your null can
# really be anything (it technically works in Fisher too, but you need to do a little math magic)
#
# You basically use your alpha to obtain a value c, known as the critical value, and compare the
# statistic to c. The picture is regions of acceptance and rejection, so ideally check yourself
# as you do it.
# One-sided test
# H_0: mu >= mu_0 (remember we want to reject this)
# H_1: mu != mu_0
# Of course use S if you don't know population stdev. Mu_0, your hypothesized
# value is always a given
# The test statistic is:
# W = (X_bar - mu_0) / (S/sqrt(n))
# for large N (CLT applies), use the acceptance region W >= -z_alpha (opposite for other sided tail)
# Else, if you have a small N, and your population is normal,
# do a t-test. (W <= t_alpha,n-1)
mu_0 = 62.68 # we have omniscience here, it's kinda a guess in real life
W = (X_bar - mu_0) / (s / math.sqrt(N))
c = norm.ppf(alpha)
print(W, c) # In this case, we do not have a significant result, so we accept the null. This makes sense,
# as the mean really IS
# Two-sided test
# H_0: mu = mu_0 (remember we want to reject this)
# H_1: mu != mu_0
W = (X_bar - mu_0) / (s / math.sqrt(N))
c = norm.ppf(1 - alpha / 2)
print(abs(W), c) # Again we accept. Statistics is working!
# ## Neyman-Pearson (Hypothesis Testing) vs. Fisher (Significance Testing)
# This debate has not ever been settled and is why Statistics is so confusing. Basically, both
# methodologies should yield the same results, though extreme edge cases may differ. The idea
# is that Fisher is more philosophically sound, while Neyman-Pearson is more mathematically
# sound. Pick your poison, but Fisher is used more in science, especially, though p values
# come with their misunderstandings and confusions.
#
# ### Power and P Values
# Before we discuss P Value misunderstandings, we must talk about type I and type II errors and power.
# A type I error is rejecting H_0 when it is true. Type II is accepting H_0 when it is false. If
# P(making type I error) <= alpha then we have a test of significance level alpha. We also have beta,
# which is equal to P(making type II error). The Power of a test is defined as 1-beta. The power is
# equivalently the probability of getting a true positive result. Thus, we seek to maximize power.
# In experiment design, power is usually made to be 80% or greater (so beta <= 20). Power analysis
# is ideally done before experiementation, and can do things like getting the minimum sample number.
# Power analysis basically gives you minimum numbers as in sample size, but it can do so in
# significance level and power effect size as well. There is always a trade off between power
# and alpha. Sample size can increase power with increasing size. Small effect size means less
# power, bigger effect size means more power.
# Minimum value of power is alpha. You basically have to choose a minimum beta to use to calculate
# power. (manipulate the probability into a pivot, by subtracting and adding the true statistic, like mean,
# and then assuming the alternative is true). It can be used to usually find the minimum sample number.
# Note: this current setting is frequentist, but Bayesian may be more appropriate here.
#
# Based on The Fickle P Value Generates Irreproducible Results:
# A p value is not good enough alone for research. High enough power is arguably more important, as without
# sufficient power, your p value is not useful enough. (you won't get a false positive, but you may get a
# false negative). In fact, unless power is high, the p-value itself fluctuates a lot.
#
# 'In the real world, the power of a study is not
# known; at best it can be estimated.'
#
# Power is a meausure of the repeatability of the p value, so 80% power means about 80% of the time it shall
# be deemed significant.
#
# In Fisher's original development of P-value analysis, his idea was that the lower the p value, the greater
# reason to doubt the null. He in fact, wanted people to use the p value as a continuous variable to help
# judgement (though not to determine it).
#
# Even with 90% power, though, P-values vary by A LOT. Let's say you obtain a P-value of 0.03. A repeat could
# give you p values anywhere between 0 and 0.6, and the chance of p < 0.05 is just 56%. This is argument
# against low power of 80% (which is standard due to our perception that false negative are much more
# acceptable compared to false positive)
#
# 'Most scientific studies have much
# less than 80% power, often around 50%
# in psychological research and averaging 21% in neuroscience'
#
# 'We must consider alternative methods of
# statistical interpretation that could be used.
# Several options are available, and although
# no one approach is perfect, perhaps the
# most intuitive and tractable is to report
# effect size estimates and their precision
# (95% confidence intervals)'
#
# 'When interpreting data, many scientists
# appreciate that an estimate of effect size is
# relevant only within the context of a specific
# study. We should take this further and not
# only include effect sizes and their 95% CIs
# in analyses but also focus our attention on
# these values and discount the fickle P value.'
#
# 'Power analysis can be replaced with
# ‘planning for precision’, which calculates
# the sample size required for estimating the
# effect size to reach a defined degree of precision'
#
# Also, if sample size grows too large, significance tests are likely to even detect tiny variations (overfitting?)
# # Bivariate Data
#
# ## Exploratory Statistics
# Graph with a scatterplot, effect is always on y, independent variable on x. We measure effect/correlation with the Pearson's correlation coefficient or R for a population:
# R = Cov(X, Y)/ (sigma_X * sigma_Y).
# for a sample, it's basically this but times 1/ (n-1) to correct bias. Since correlation relies on mean and stdev,
# it will fluctuate based on outliers.
#
# The R^2 is the coefficient of determination is the percentage of variation in y that is explained by variation in x.
# Must interpret R^2 in context of problem. Fo scientific experiments, R^2 of 90% or greater is standard. Observational
# studies can be informative in the 10-20% range however.
#
# Of course, the line of best fit is done by least squares, which can be extended for many nonlinear relationships that are
# linear in the coefficients. Key thing is that the plot of residuals is normally distributed to have a good fit.
#
# Clusters and outliers are important to note
sample.sort()
effect = [(x - mu)**2/ 4 + rand.gauss(0, 1.5) for x in sample]
plt.scatter(sample, effect)
# In this case we don't have a linear relationship. It seems to be quadratic.
# We can however perform least squares to fit this: Ax = b => x=(A^T*A)^-1A^T * b
k = 2 #num of predictors, for quadratic it's 2. Constant term is not counted
A = np.asmatrix([[x**2, x, 1] for x in sample])
b = np.asmatrix(effect)
b = np.reshape(b, (50, 1))
coeff = np.linalg.inv(A.T*A) * A.T * b
quad = [coeff[0].item()* x **2 + coeff[1].item() * x + coeff[2].item() for x in sample]
m, b = np.polyfit(sample, effect, 1)
lin = [m * x + b for x in sample]
plt.plot(sample, effect, 'o')
plt.plot(sample, quad)
plt.plot(sample, lin)
# Now for Pearson's R:
# Strictly speaking, Pearson only measures a linear relationship (why transformations are so important)
r = sp.pearsonr(sample, effect)[0]
print(r, r **2)
# The sample is poor mostly due to the nonlinearity. In a measurable sense, however, the R^2 is too low
# R^2 also overestimates population determination, rho^2
# Pearson's R also works in multiple linear regression, where we use x**2, x as explanatory variables . The R we get is the
# coefficient of multiple correlation. In this case, we must find the R^2 first and square root it for R:
# R^2 = c.T * R_xx.inv * c, where c is the vector c = [r_x1y, r_x2y, ...].T, the correlations between predictors and predicted
# R_xx = the matrix of predictor correlations, [[rx1x1 rx1x2 ....], [rx2x1 rx2x2 ...] ...]
sq_sample = [x **2 for x in sample]
r_x2x = sp.pearsonr(sample, sq_sample)[0]
r_xx = sp.pearsonr(sample, sample)[0]
r_x2x2 = sp.pearsonr(sq_sample, sq_sample)[0]
R_xx = np.asmatrix([[r_x2x2, r_x2x],
[r_x2x, r_xx]])
r_x2y = sp.pearsonr(sq_sample, effect)[0]
r_xy = sp.pearsonr(sample, effect)[0]
c = np.asmatrix([r_x2y, r_xy]).T
R2 = c.T * np.linalg.inv(R_xx) * c
R = math.sqrt(R2)
print(R, R2)
# Unfortunately, overfitting is a problem. As more variables are added, R^2 grows. It is best to use the adjusted R^2.
# We can adjust though according to the formula: 1 - (1 - R^2) * (n - 1) / (n - p - 1) where p is the number of predictor
# variables, n being the number of samples. This is known as the Ezekiel adjustment. However this is a biased estimator.
R_adj2 = 1 - (1 - R2) * (N - 1) / (N - 2 - 1)
print(R_adj2)
# The better adjustment is known as the Olkin-Pratt estimator or an approximation of it, confirmed through emperical comparisons.
# OPK involves a hypergeometric series, but has valid approximations. The Pratt approximation is my favored one:
# (N - 3)/ (N - k - 1) * (1-R^2) * (1 + 2(1-R^2)/(N - k - 2.3))
rho2 = (N - 3) / (N - k - 1) * (1 - R2) * (1 + 2 * (1 - R2) / (N - k - 2.3))
print(rho2)
# https://online.ucpress.edu/collabra/article/6/1/45/114458/Improving-on-Adjusted-R-Squared
# this paper has more info
# Graph residuals
lin_res = [(lin[i] - effect[i])**2 for i in range(N)]
quad_res = [(quad[i] - effect[i])**2 for i in range(N)]
plt.plot(sample, lin_res, 'o')
plt.plot(sample, quad_res, 'x')
# residuals are always centered on 0 for a good model. For our purposes, we assume Gaussian distribution of those residuals
# which we can test via tests for normality, but in some cases, there may not be a normal distribution
# Confidence Intervals
# Now, we shall do confidence intervals with this graph. Officially, this test only works if you have a normally distributed,
# zero residual random sample with linear scatterplots. The confidence interval is for the true slope of the line.
# The formula for the sample stdev of the slope is sqrt(sum((y - y_hat)**2) / (n-2))/sqrt((sum(x-X_bar)**2)),
# which is also the standard error. This test works for multivariate regression by giving a confidence interval for each
# coefficient.
T_reg = t.ppf(1 - alpha / 2, N - 2)
s_b = math.sqrt(sum([(effect[i] - lin[i])**2 for i in range(N)])/ (N-2))
s_b = s_b / math.sqrt(sum([(x - X_bar)**2 for x in sample]))
[m - T_reg * s_b, m + T_reg * s_b]
# For multivariate regression you compute CI for each coefficient. Here's how:
# e = the vector of residuals. We have the variance of the sample:
# e.T * e / (n - k) = var_hat
# Also for the least squares problem: X*beta = y, we obtain
# C = var_hat * (X.T * X).inv
# To obtain confidence intervals then, we just do
# beta_j +-t_alpha/2, n - p - 1 * sqrt(C_jj)
e = np.asmatrix([(effect[i] - quad[i]) for i in range(N)]).T
var_hat = (e.T * e / (N - k))[0, 0]
C = var_hat * np.linalg.inv(A.T * A)
t_reg = t.ppf(1 - alpha/2, N - k - 1)
size_of_interval = np.asmatrix([t_reg * math.sqrt(C[i, i]) for i in range(3)]).T
print(coeff, size_of_interval) # +- size of interval for each row is the confidence intervals
# You can do confidence intervals for each point already along the graph, which is known as finding the mean response CI. Our formula is given x_h, we have y_hat_h +- t_alpha/2,n-2 * s_e * sqrt(1/n + (x_h - X_bar)^2/sum((x_i - X_bar)^2))
# You can use this to plot a confidence interval curve around your model.
#
# For multivariate regression, given a vector x, with X * beta = y as our model, we have residual standard error:
# s_e = sqrt((y+i - y_bar)^2/(n - k - 1)), and from this we can obtain (remember k is one less than the coefficients being used)
# y_h +- t_alpha/2, n - k - 1 * s_e * sqrt(x.T * (X.T * X).inv * x)
# Lastly we have prediction intervals, or confidence intervals for new observations to our model
# The formula is y_n+1 has the interval Y_n+1_hat +- t_alpha/2,n-2 * s_e * sqrt(1 + 1\n + (x_n+1 - X_n_bar)^2/sum((x_i - X_bar)^2))
#
# For Multivariate:
# Y_n+1_hat +- t_alpha/2,n-k-1 * s_e * sqrt(1 + x.T * (X.T * X).inv * x)
#
# Some remarks: prediction intervals are always wider than confidence intervals. Furthermore, prediction intervals also require a lot more conditions to go right, basically your model must be nearly ideally normal in every way to have valid prediction. See the below guides to read up more, especially on the conditions for these predictions. (remember that predictions must be made in a valid region of your model, and determining that valid region is very difficult)
# https://online.stat.psu.edu/stat501/lesson/3/3.3
# https://daviddalpiaz.github.io/appliedstats/multiple-linear-regression.html
#
# ## Hypothesis Testing on Regression
# t-tests on coefficients. Basically this lets us determine if a new predictor added to the model is useful or not.
# For linear regression, it measures the slope, and determines the existence of correlation
# We create the t-statistic for linear regression as follows:
# T = (beta_hat - beta_null)/ sqrt(sum of residuals squared / ((n-2) * sum((x_i - X_bar)^2)) )
# Beta_null is often 0, as our null hypothesis is basically always, do we need this coefficient. Do your choice of Fisherian
# or Neyman-Pearson testing.
#
# For multiple regression, we test individual coefficients.
# Our statistic:
# T = (beta_j_hat - beta_j_null)/sqrt(C_jj) where C is the matrix in the above discussion of multiple regression.
#
# The problem is that these don't test the overall fit. For that there are two popular approaches: F-tests and ANOVA
#
# ### F-Tests
# You would want to use the f distribution from scipy.
# For f-distributions we are measuring the sums of squares if the null were true. That is, we look at both the residual squares (SSE) and the differences of the predicted y_values to the mean y (summed and squared) (SSR). However, we must also take into account degrees of freedom, so for SS_R we have dof of 1, so our MSR = SSR / 1. MSE is similar with dof N-2, so MSE = SSE / (n-2)
# Our f-statistic is
# F = MSR / MSE
# using the f distribution with f_alpha, 1, n-2 (f-tests are single-tailed). Btw, in linear regression, squaring the t statistic
# gives the same f-statistic
#
# For multivariable regression, it's almost same expression for F, but the null hypothesis is that all the coefficients (not
# including the constant term) are equal to 0. Thus the alternative is that at least one coefficient has a linear relationship.
# MSR = SSR / (p - 1) and MSE = SSE / (n - p) and we use the F_p-1,n-p distribution for our value.
#
# You can also run Partial F-tests on groups of coefficients. Say we have a model with q predictors < p. We have the Sums of squares of differences between each models predictions = SSD, and the SSE between the model with p predictors. We have MSD = SSD / (p - q), MSE = SSE / (n - p). Do F = MSD / MSE on F_p-q,n-p
# Our H_0 is that beta_q ,....,beta_p-1 are all equal to 0, the alternative being that at least one is linearly related.
#
# F-tests are generally run on continuous independent variables. ANOVA is for discrete independent variables. Discrete or continuous response variables don't really matter.
# ### ANOVA
# ANOVA is the other way of significance testing in multivariate analysis. It uses F-tests as a component.
# There are a few forms of ANOVA, one-way ANOVA is used when one explanatory variable is in use, while factorial ANOVA is when more than one explanatory variable exists. ANOVA works for categorical variables.
# There's also ANCOVA and Repeated Measures ANOVA (repeated measures is used when same subjects receive each factor, like in longitudinal study). ANOVA is good for a sort of omnibus test check to make sure if there is even a statistical difference in your data. If you find a difference, then pairwise t-tests are kinda the only way to figure out what is significant from what, although graphing side by side can also help.
#
# Caution: as with the above, these are assuming the standard linear assumptions: Guassian random error, heteroescedasticity (variation across predictors). Furthermore, if your treatment and experiment has groups that are unbalanced (with severely different numbers for each group), then ANOVA may go bonkers.
#
# #### One Way ANOVA
# One Way ANOVA works with categorical variables with one response variable and one predictor per group (say k groups).
# Say we have a table of bunnies we see and the weather on that day. We might model the bunnies seen based on the weather, and weather might only have two values: 1 for raining and 0 for normal. This is generally what one way ANOVA works with. But it builds a model by treating the weather as if it were a continuous one. You need the sum of squares in between groups, and the sum of squares of error and will perform an F-test. Between (also known as Model) Sum of Squares is obtained by first finding the overall mean of all data points. Then you add the squared difference of each group's mean from the overall mean multipled by the number of samples per group. This is the SSB. The SSE is the sums of squares between each point in a group and that group mean and sum across all groups. Then we need MSB and MSE. MSB = SSB / (k - 1) and MSE = SSE / (n - k), where n is the overall number of points. F = MSB / MSE, then perform your F-test.
#
# #### Factorial ANOVA (Two-way ANOVA, Three-way ANOVA, when there are 2/3 factors in the test. 4 and up are almost never used because of difficulties in interpretation. This will focus on two-way)
# In this ANOVA, we have to account for possible interaction terms. If you have independent (orthogonal) factors, then you can assume your model has a coefficient for each factor. If you have interactions, then you must include a coefficient for each interaction. Degrees of freedom per factor is number of levels (num of groups) - 1. You essentially perform an F-test for each factor, so if we want color and manufacturer influence on cars, then we do one ANOVA on color, and one ANOVA on manufacturer using the above one way ANOVA. For interactions, we first need the sums of squares between groups. This is done by taking all the combinations of the factors and treating each as a group, then subtracting each of those group means from the overall mean. To obtain SSI or the interaction SS, we subtract sums of squares for each factor. For the MSI, we divide by degrees of freedom of each factor multiplied together. MSE is SSE / N - factor a levels * factor b levels where N is the overall number of samples.
#
# To check for interactions, use interaction plots. The groups are not interactive if you have about parallel lines in the interaction plot.
#
# #### ANCOVA
# In ANCOVA, we account for a covariate (it blends regression and ANOVA). It controls for a continuous variable to increase statistical power, though adding willy nilly covariates to control for can reduce power if one is not careful. You essentially run a regression and then take the residuals, and run ANOVA on the residuals
#
# #### RMA
# This is used to control within individuals. It basically calculates a sum of squares for each subject and removes that from the SSE.
# ### Chi Square Tests
# These are nonparametric tests
# #### Test of Goodness of Fit
# This tests for whether sample data is representative of the population data
# Your chi square is the sum of obs - exp squared divided by the expected.
# You run on Chi2_n-1 where n is number of groups.
# #### Test for Independence
# This tests for when two categorical variables are related or not
# df = (r-1)* (c-1) where r and c are number of rows and columns respectively.
# Have to calculate expected table, and the chi square is the same based on this. To calculate the
# expected, you need to sum up totals of each row and column. For each column (the dependent variable), you take the proportion of the total that its sum is and apply that proportion across the totals for each row to get expected.
# #### Test for Homogeneity of Proportions
# This tests for if two categorical variables are drawn from the same population or not
# Essentially run the same as the test for independence, but you are sampling from (hypothesized) two different populations rather than from one in the above.
# ### Effect Size
# There are three types that these metrics generally fall into. For the correlation family, R^2 and eta squared are standard, explaining variance in a model. In the difference family, differences between means is measured, the primary metric being Cohen's d. The last are for categorical variables and are measures of association, the phi coefficient being the main one for things like Chi-squared.
# #### Eta squared
# The equivalent for R2 but for ANOVA. Measures amount of variation explained by model
#
# #### Cohen's d
# Measure of the perceived difference between two means
#
# #### Phi Coefficient
# Measure of the association between categorical variables
# ## Power Analysis
# I think this being forced to derive these equations is the best way of learning elementary statistics. Power is arguably more important than P-values, yet we focus so little on it. Power is one of the four parts to conducting any statistical experiment:
# effect size, number of observations, alpha/significance level, and power.
# These four quantities are interdependent in one equation, though that equation may be implicit and therefore have no closed solution.
#
# Power Analysis is commonly run to determine your necessary sample size. Running it for effect size is common too, but we shall focus on sample size in particular; understanding power analysis for one variable means it is theoretically similar on the others.
#
# The method for power analysis is this: given desired effect size, power, and alpha, find the inequality for N to be sufficient for all these minimums to be satisfied. Since we are doing this in general, the sampling distributions require standardization. The test-statistic you find is for the standardized distribution. But this means we can just assume mu_0 is 0 itself, the test doesn't change. The test-statistic, in terms of the effect size (cohen's d) and sample size is equal to d* sqrt(N). You assume that your alternative hypothesis sampling distribution is centered at that value. Then you proceed to use alpha to obtain the critical value. You then want to find the beta, which is the tail of the alternative hypothesis. We can apply shifts to then use the same CDF to find the Beta. Using this, you should be able to get an equation in terms of N, d, alpha, and Pow. It's then a matter of manipulating terms to find your desired value. For sample size of a one tailed test, we have
# N >= (phi^-1(1-alpha) - phi^-1(1 - Pow))^2 / d^2
#
# It's common to also find power curves, where power is dependent on one value varied while the others held constant. For the power curve for sample size, we manipulate things to find
#
# 1 - phi(phi^-1(1-alpha) - d * sqrt(N)) = pow
#
# This is however for one tailed tests. For two tailed tests, it is a bit more complicated. or two tailed, you want to take the absolute value for d. It doesn't really matter whether d * sqrt(N) is higher or lower, so we typically choose it higher. Then you want to apply the same process to get critical values for both tails and with half alphas. Then you should get a rather unwieldy equation for the power curve. In the olden days, it was basically just holding alpha and d constant as you painstakingly calculated the curve for different N's until you found the threshold for a good power level, but nowadays we just use solvers. It would be hard to get the minimum N explicitly, but using a solver should give you the minimum N as power curves are monotonic.
# The equation for the power curves are
# -phi(phi^-1(alpha/2) - |d|sqrt(N)) + phi(phi^-1(1-alpha/2) - |d|sqrt(N)) = Pow
#
# Practically, statsmodel has a power solver that can solve for any one of the variables left blank.
#
# But that concludes Everyday Statistics the practical guide for all the standard and elementary statistics any stats major needs to know!
# +
#TODO in other jupyter notebooks:
# Bayesian Statistics https://en.wikipedia.org/wiki/Bayesian_statistics, https://amstat.tandfonline.com/doi/abs/10.1080/00031305.1986.10475342#.YPYDcZhKjIU, https://projecteuclid.org/journals/bayesian-analysis/volume-3/issue-3/Objections-to-Bayesian-statistics/10.1214/08-BA318.full
# Bayesian hierarchical modeling, Bayesan Network, Emperical Bayes, Monte Carlo Markov Chains
# https://www.nature.com/articles/s43586-020-00001-2, https://www.nature.com/articles/s43586-020-00001-2.pdf
# Generalized Linear Models (Bayesian and Variance Stabilized Least Squares are alternatives), Stat 151a, dummy variable regression
# Nonparametric Statistics and robust methods for modelling: https://www.statisticshowto.com/probability-and-statistics/statistics-definitions/parametric-and-non-parametric-data/,
# https://jsteinhardt.stat.berkeley.edu/teaching/stat240-spring-2021, http://mlss.tuebingen.mpg.de/2015/slides/ghahramani/gp-neural-nets15.pdf
# https://arxiv.org/pdf/1906.10221.pdf
# https://en.wikipedia.org/wiki/Robust_statistics
# https://en.wikipedia.org/wiki/Robust_regression
# https://en.wikipedia.org/wiki/Robust_confidence_intervals, Kohonen Self organizing map (KSOM)
# Causal Inference: https://www.ucbbiostat.com/
# Decision theory: https://data102.org/, game theory: https://bcourses.berkeley.edu/courses/1454200/assignments/syllabus,
# Sampling + Large Sampling theory + Monte Carlo and stuff + experiments?, theoretical statistics: STAT 210 (combine with Bayesian?)
# Experiment design: http://statweb.stanford.edu/~owen/courses/363/
# -
| Everyday Statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import re
sns.set(color_codes=True)
sns.set(font_scale=True)
sns.set_color_codes()
# -
df = pd.read_csv("../../milky_white_mushroom/results/cazymes/shiitake/filtered_hits.tsv",delimiter="\t")
df
# +
from typing import Dict
from collections import Counter
def clean_label(element):
indexes_removed = re.sub(r"\([0-9]+-[0-9]+\)","",element)
return re.sub(r"_[0-9]+","",indexes_removed)
def get_counts(df):
total = []
for row in df.values:
prediction1 = clean_label(row[1]).split('+')
prediection2 = clean_label(row[3]).split('+')
list(set(prediction1+prediection2))
total += [s if "_" not in s else s.split("_")[0] for s in set(prediction1+prediection2) if s != "N" and s != "-" and "." not in s]
return Counter(total)
def count_cazymes(all_counts: Dict[str,int]):
cazyme_count = {}
for sub_group,count in all_counts.items():
group = sub_group[:2]
if group[:2] == "CB":
group = sub_group[:3]
cazyme_count[group] = cazyme_count.get(group,0) + count
return cazyme_count
def count_families(all_counts: Dict[str,int]):
family_counts = {}
for sub_group,count in all_counts.items():
group = sub_group[:2]
if group[:2] == "CB":
group = sub_group[:3]
sub_groups =family_counts.get(group,{})
sub_groups[sub_group] = sub_groups.get(sub_group,0) + count
family_counts[group] = sub_groups
return family_counts
# -
all_counts = get_counts(df)
cazyme_count = count_cazymes(all_counts)
family_count = count_families(all_counts)
cazyme_count
family_count
sum(value for value in cazyme_count.values())
# +
figure,axes = plt.subplots(nrows=1,ncols=1,figsize=(15,6))
labels_sorted = sorted(list(family_count['GH'].items()),key=lambda pair: pair[0][2:])
labels,counts = list(zip(*labels_sorted))
axes.bar(list(range(len(labels))),counts,tick_label=labels)
plt.tight_layout()
#s = pd.Series(family_count["GH"])
#s.plot(ax=axes,kind="bar")
plt.ylabel("Count")
plt.xlabel("Glycoside Hydrolase family")
plt.yticks(range(1,21))
plt.xticks(rotation=90)
plt.savefig('GH-Counts.png',bbox_inches="tight")
plt.show()
# +
figure,axes = plt.subplots(nrows=1,ncols=1,figsize=(15,6))
labels_sorted = sorted(list(family_count['GH'].items()),key=lambda pair: pair[1])
labels,counts = list(zip(*labels_sorted))
axes.bar(list(range(len(labels))),counts,tick_label=labels)
plt.tight_layout()
#s = pd.Series(family_count["GH"])
#s.plot(ax=axes,kind="bar")
plt.ylabel("Count")
plt.xlabel("Glycoside Hydrolase family")
plt.yticks(range(1,21))
plt.xticks(rotation=90)
plt.savefig('GH-Counts.png',bbox_inches="tight")
plt.show()
# -
| notebooks/Cazymes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Monolayer: Complex hierarchies, patterns, tiling and writing to files
# ---------------------------------------------------------------
#
# __Note__: mBuild expects all distance units to be in nanometers.
#
# In this example, we'll cover assembling more complex hierarchies of components using patterns, tiling and how to output systems to files. To illustrate these concepts, let's build an alkane monolayer on a crystalline substrate.
#
# First, let's build our monomers and functionalized them with a silane group which we can then attach to the substrate. The `Alkane` example uses the `polymer` tool to combine `CH2` and `CH3` repeat units. You also have the option to cap the front and back of the chain or to leave a `CH2` group with a dangling port. The `Silane` compound is a Si(OH)<sub>2</sub> group with two ports facing out from the central Si. Lastly, we combine `alkane` with `silane` and add a label to `AlkylSilane` which points to, `silane['down']`. This allows us to reference it later using `AlkylSilane['down']` rather than `AlkylSilane['silane']['down']`.
#
# __Note:__ In `Compounds` with multiple `Ports`, by convention, we try to label every `Port` successively as 'up', 'down', 'left', 'right', 'front', 'back' which should roughly correspond to their relative orientations. This is a bit tricky to enforce because the system is so flexible so use your best judgement and try to be consistent! The more components we collect in our library with the same labeling conventions, the easier it becomes to build ever more complex structures.
# +
import mbuild as mb
from mbuild.lib.recipes import Alkane
from mbuild.lib.moieties import Silane
class AlkylSilane(mb.Compound):
"""A silane functionalized alkane chain with one Port. """
def __init__(self, chain_length):
super(AlkylSilane, self).__init__()
alkane = Alkane(chain_length, cap_end=False)
self.add(alkane, 'alkane')
silane = Silane()
self.add(silane, 'silane')
mb.force_overlap(self['alkane'], self['alkane']['down'], self['silane']['up'])
# Hoist silane port to AlkylSilane level.
self.add(silane['down'], 'down', containment=False)
# -
AlkylSilane(5).visualize()
# Now let's create a substrate to which we can later attach our monomers:
# +
import mbuild as mb
from mbuild.lib.surfaces import Betacristobalite
surface = Betacristobalite()
tiled_surface = mb.lib.recipes.TiledCompound(surface, n_tiles=(2, 1, 1))
# -
# Here we've imported a beta-cristobalite surface from our component library. The `TiledCompound` tool allows you replicate any `Compound` in the x-, y-
# and z-directions by any number of times - 2, 1 and 1 for our case.
#
# Next, let's create our monomer and a hydrogen atom that we'll place on unoccupied surface sites:
from mbuild.lib.atoms import H
alkylsilane = AlkylSilane(chain_length=10)
hydrogen = H()
# Then we need to tell mBuild how to arrange the chains on the surface. This is accomplished with the "pattern" tools. Every pattern is just a collection of points. There are all kinds of patterns like spherical, 2D, regular, irregular etc. When you use the `apply_pattern` command, you effectively superimpose the pattern onto the host compound, mBuild figures out what the closest ports are to the pattern points and then attaches copies of the guest onto the binding sites identified by the pattern:
#
# +
pattern = mb.Grid2DPattern(8, 8) # Evenly spaced, 2D grid of points.
# Attach chains to specified binding sites. Other sites get a hydrogen.
chains, hydrogens = pattern.apply_to_compound(host=tiled_surface, guest=alkylsilane, backfill=hydrogen)
# -
# Also note the `backfill` optional argument which allows you to place a different compound on any unused ports. In this case we want to backfill with hydrogen atoms on every port without a chain.
monolayer = mb.Compound([tiled_surface, chains, hydrogens])
monolayer.visualize() # Warning: may be slow in IPython notebooks
# Save as .mol2 file
monolayer.save('monolayer.mol2', overwrite=True)
# `lib.recipes.monolayer.py` wraps many these functions into a simple, general class for generating the monolayers, as shown below:
# +
from mbuild.lib.recipes import Monolayer
monolayer = Monolayer(fractions=[1.0], chains=alkylsilane, backfill=hydrogen,
pattern=mb.Grid2DPattern(n=8, m=8),
surface=surface, tile_x=2, tile_y=1)
monolayer.visualize()
# -
| docs/tutorials/tutorial_monolayer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pickle
import json
import os
import sys
import copy
import sklearn.preprocessing
import models
from sklearn.neighbors import NearestNeighbors
import utils
from collections import Counter
import matplotlib.pyplot as plt
data='fb15k'
DATA_DIR = "../../data/"+data
DUMP_FILE = "../dumps/"+data+"_distmult_dump_norm.pkl"
MODEL_TYPE = data
# mining_dir=data+"_low_thresh"
mining_dir=data+"_rule_mining_tmp"
os.system("mkdir -p "+mining_dir)
train_data = utils.read_data(os.path.join(DATA_DIR,"train.txt"))
# dev_data = read_data(os.path.join(DATA_DIR,"valid.txt"))
# test_data = read_data(os.path.join(DATA_DIR,"test.txt"))
dump=utils.load_pickle(DUMP_FILE)
dump.keys()
model=models.TypedDM(DUMP_FILE)
mapped_train_data = utils.map_data(train_data,dump)
# mapped_dev_data = map_data(dev_data)
# mapped_test_data = map_data(test_data)
entity_to_rel=utils.get_ent_to_rel(mapped_train_data)
index_head=utils.get_head_index(mapped_train_data)
# # Length 1 Rules
relation_count=len(dump['relation_to_id'])
nbrs = NearestNeighbors(n_neighbors=relation_count,metric=model.similarity_relembedding).fit(model.relation_matrix)
print(relation_count)
count_r,set_r=utils.get_relation_dict(mapped_train_data)
print(len(count_r),len(set_r))
support=1
rules_dict_1={}
count=0
count2=0
for r1 in range(relation_count):
if(r1%100==0):
print(r1)
if(count_r[r1]<support):
continue
combined_rel=model.relation_matrix[r1].reshape((1,-1))
distances, indices = nbrs.kneighbors(combined_rel)
indices=indices[0]
bool_arr=[False for i in range(relation_count)]
for ind in indices:
if ind!=r1:
bool_arr[ind]=True
cur_dict={}
for pair_e1e2 in set_r[r1]:
if pair_e1e2 not in entity_to_rel:
print("Strange")
continue
for r2 in entity_to_rel[pair_e1e2]:
count+=1
if bool_arr[r2]==False:
continue
if r2 not in cur_dict:
count2+=1
cur_dict[r2]=0
cur_dict[r2]+=1
rules_dict_1[r1]=cur_dict
print(len(rules_dict_1))
print(count,count2)
# # Length 2 Rules
# %time count_r1_r2,set_r1_r2=utils.get_r1r2_count(mapped_train_data,index_head,get_set=True)
nbrs = NearestNeighbors(n_neighbors=500,metric=model.similarity_relembedding).fit(model.relation_matrix)
support=1
count=0
rules_dict={}
for r1 in range(relation_count):
if(r1%100==0):
print(r1)
for r2 in range(relation_count):
if(r1==r2):
continue
pair=(r1,r2)
if(pair not in count_r1_r2 or count_r1_r2[pair]<support):
continue
combined_rel=model.dot_relation(r1,r2).reshape((1,-1))
distances, indices = nbrs.kneighbors(combined_rel)
bool_arr=[False for i in range(relation_count)]
for ind in indices[0]:
if ind!=r1 and ind!=r2:
bool_arr[ind]=True
if bool_arr[r1]==True or bool_arr[r2]==True:
print("Strange")
print(indices[0],r1,r2)
cur_dict={}
entpair_lis=list(set_r1_r2[pair])
for pair_e1e2 in entpair_lis:
if pair_e1e2 not in entity_to_rel:
continue
for r in entity_to_rel[pair_e1e2]:
if(bool_arr[r]==False):
continue
if r not in cur_dict:
count+=1
cur_dict[r]=0
cur_dict[r]+=1
rules_dict[pair]=cur_dict
print(len(rules_dict))
print(relation_count)
print(count)
# # Saving rules
path1='1_sup=1.pkl'
path1=os.path.join(mining_dir,path1)
utils.dump_pickle(rules_dict_1,path1)
print(len(rules_dict_1))
path2='2_sup=1_nei=500.pkl'
path2=os.path.join(mining_dir,path2)
utils.dump_pickle(rules_dict,path2)
print(len(rules_dict))
# ## Saving auxilary data for rule2
path='set_r1_r2.pkl'
path=os.path.join(mining_dir,path)
set_len_r1_r2={}
for r1r2 in set_r1_r2:
set_len_r1_r2[r1r2]=len(set_r1_r2[r1r2])
utils.dump_pickle(set_len_r1_r2,path)
print(len(count_r1_r2))
# ## Analyse 3 Rules
# +
rules_3=[]
count=0
count_dict={}
for r1r2 in set_r1_r2:
count+=1
if(count%100==0):
print(count)
for e1e2 in set_r1_r2[r1r2]:
e1=e1e2[0]
e2=e1e2[1]
# e1-r1-e-r2-e2
count_conf_dict={}
# e1-r1-e-r2-e2-r3-e3
if e2 not in index_head:
continue
for r3e3 in index_head[e2]:
e3=r3e3[1]
triplet=(r1r2[0],r1r2[1],r3e3[0])
if triplet not in count_dict:
count_dict[triplet]=0
count_conf_dict[triplet]={}
count_dict[triplet]+=1
# -
print(len(count_dict))
value_list=list(count_dict.values())
cnt=Counter(value_list)
cnt=sorted(cnt.items())
array=np.array(cnt)
print(array.shape)
plt.plot(array[:,1])
plt.show()
temp=np.cumsum(array[:,1])/np.sum(array[:,1])
print(array[temp[:]<0.5],temp[temp[:]<0.5])
# ## Rules Length 3
min_sup=4
confidence=0
r1r2_ent=utils.get_r1r2_e1e2_dict(mapped_train_data,index_head)
# +
rules_3=[]
count=0
for r1r2 in set_r1_r2:
# prit(r1r2)
count+=1
if(count%100==0):
print(count)
print(len(rules_3))
for e1e2 in set_r1_r2[r1r2]:
e1=e1e2[0]
e2=e1e2[1]
# e1-r1-e-r2-e2
e=r1r2_ent[r1r2][e1e2]
count_conf_dict={}
count_dict={}
# e1-r1-e-r2-e2-r3-e3
if e2 not in index_head:
continue
for r3e3 in index_head[e2]:
e3=r3e3[1]
triplet=(r1r2[0],r1r2[1],r3e3[0])
if triplet not in count_dict:
count_dict[triplet]=0
count_conf_dict[triplet]={}
count_dict[triplet]+=1
pair=(e1,e3)
if pair not in entity_to_rel:
continue
for rel in entity_to_rel[pair]:
if rel not in count_conf_dict[triplet]:
count_conf_dict[triplet][rel]=0
count_conf_dict[triplet][rel]+=1
for r1r2r3,supp in count_dict.items():
if supp < min_sup:
continue
for r4,conf in count_conf_dict[r1r2r3].items():
if conf>confidence:
rules_3.append((r1r2r3,r4,supp,conf))
# -
rules_3.sort(key=lambda x:(x[3]*1.0)/x[2],reverse=True)
path_rule_3=os.path.join(mining_dir,"3_sup=4_conf=0.pkl")
utils.dump_pickle(rules_3,path_rule_3)
# ## Load Len 3 rules
rules_3=utils.load_pickle(path_rule_3)
rules_3[0:10]
print(len(rules_3))
len(rules_3)
min_sup
| Rule-Mining-Distmult/Rule_Mining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# !pip install transformers
# +
import pandas as pd
import numpy as np
from nltk.stem import PorterStemmer
# LaBSE
from transformers import AutoTokenizer, AutoModel
# facebook/bart-base
from transformers import BartTokenizer, BartModel
import torch
import plotly.express as px
import datetime
# -
# # Генерация эмбеддингов для строк
print(datetime.datetime.now())
# ## JSON to DataFrame
# !python data.py
# ## Загрузка данных
df_education = pd.read_csv('data/uuid_x_education_fields.csv', index_col=0)
df_education.head(5)
# ## Facebook/bart-base
# + active=""
# tokenizer = BartTokenizer.from_pretrained('facebook/bart-base')
# model = BartModel.from_pretrained('facebook/bart-base')
#
# encoded_input = tokenizer("Computer Sciense", padding=True, truncation=True, max_length=64, return_tensors="pt")
#
# with torch.no_grad():
# model_output = model(**encoded_input)
#
# embeddings = model_output.last_hidden_state
# tensor = torch.nn.functional.normalize(embeddings)
#
# array = tensor.cpu().detach().numpy()
# print(array.shape)
#
# array[0].shape
# + active=""
# encoded_input = tokenizer("English", padding=True, truncation=True, max_length=64, return_tensors='pt')
#
# with torch.no_grad():
# model_output = model(**encoded_input)
#
# embeddings = model_output.last_hidden_state
#
# np.array(embeddings[0].shape)
# + active=""
# def get_bart_base_embeddings(df_education):
#
# tokenizer = AutoTokenizer.from_pretrained("facebook/bart-base")
# model = AutoModel.from_pretrained("facebook/bart-base")
#
# embedding_vec = []
# df_education_study = df_education[['education_field_of_study', 'education_school_name']].drop_duplicates().reset_index(drop=True)
#
# for i, row in enumerate(df_education_study['education_field_of_study']):
#
# print('{} / {}'.format(i+1, df_education_study.shape[0]), end="\r")
#
# embeddings = []
#
# if row == row:
# encoded_input = tokenizer(row, padding=True, truncation=True, max_length=64, return_tensors='pt')
#
# with torch.no_grad():
# model_output = model(**encoded_input)
#
# embeddings = model_output.pooler_output
#
# embedding_vec.extend(np.array(embeddings))
#
# df_embeddings = pd.DataFrame(embedding_vec)
#
# df_embeddings = pd.concat(
# [
# df_embeddings,
# df_education_study
# ], axis=1
# )
#
# return df_embeddings
# + active=""
# df_education = pd.read_csv('data/uuid_x_education_fields.csv', index_col=0)
# df_embeddings = get_bart_base_embeddings(df_education)
# + active=""
# df_embeddings
# -
# ## cointegrated/LaBSE-en-ru
# https://huggingface.co/cointegrated/LaBSE-en-ru
def get_education_embeddings(df_education):
tokenizer = AutoTokenizer.from_pretrained("cointegrated/LaBSE-en-ru")
model = AutoModel.from_pretrained("cointegrated/LaBSE-en-ru")
embedding_vec = []
df_education_study = df_education[~df_education['education_field_of_study'].isna()][['education_field_of_study']].drop_duplicates().reset_index(drop=True)
for i, row in enumerate(df_education_study['education_field_of_study']):
print('{} / {}'.format(i+1, df_education_study.shape[0]), end="\r")
embeddings = []
if row == row:
encoded_input = tokenizer(row, padding=True, truncation=True, max_length=64, return_tensors='pt')
with torch.no_grad():
model_output = model(**encoded_input)
embeddings = model_output.pooler_output
#embeddings = torch.nn.functional.normalize(embeddings)
embedding_vec.extend(np.array(embeddings))
df_embeddings = pd.DataFrame(embedding_vec)
df_embeddings = pd.concat(
[
df_embeddings,
df_education_study
], axis=1
)
return df_embeddings
df_education = pd.read_csv('data/uuid_x_education_fields.csv', index_col=0)
df_embeddings = get_education_embeddings(df_education)
df_embeddings.to_csv('data/embeddings/education_field_of_study.csv')
| code/data-embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cmip6-processing
# language: python
# name: cmip6-processing
# ---
# # Nukleus - exploring submasks
# +
# %matplotlib inline
import xarray as xr
import numpy as np
# use orography to look at the data
file = "/pool/data/CORDEX/data/cordex/output/EUR-11/GERICS/ECMWF-ERAINT/evaluation/r0i0p0/GERICS-REMO2015/v1/fx/orog/v20180813/orog_EUR-11_ECMWF-ERAINT_evaluation_r0i0p0_GERICS-REMO2015_v1_fx.nc"
ds = xr.open_dataset(file)
pollat = ds.rotated_latitude_longitude.grid_north_pole_latitude
pollon = ds.rotated_latitude_longitude.grid_north_pole_longitude
mask = xr.open_dataset("NUKLEUS_MASKEN_BTU-20210510.nc")
mask
# -
# fix inconsistencies, make submask a coordinate for convenience:
# fix inconsistencies, make submask a coordinate for convenience:
#mask = mask.rename({'submask_names':'submask', 'submask_index':'submask'})
mask = mask.swap_dims({'submask_index':'submask_names'}).rename({'submask_names':'submask'})
mask['submask'] = mask.submask.astype(str)
mask
# the coordinates of the mask file and the ESGF dataset are not identical (because of different precision). we fix this here so that we can use the mask file in the where function. let's just copy the original coordinates to the mask dataset.
mask.coords['rlon'] = ds.rlon
mask.coords['rlat'] = ds.rlat
mask.coords['lon'] = ds.lon
mask.coords['lat'] = ds.lat
mask.MASK.plot(col='submask', col_wrap=4)
# ### mask all regions
def plot(da, title=''):
import cartopy.crs as ccrs
import cartopy.feature as cf
import matplotlib.pyplot as plt
plt.figure(figsize=(20,10))
projection = ccrs.PlateCarree()
transform = ccrs.RotatedPole(pole_latitude=pollat, pole_longitude=pollon)
#ax = plt.axes(projection=projection)
ax = plt.axes(projection=transform)
#ax.set_extent([ds_sub.rlon.min(), ds_sub.rlon.max(), ds_sub.rlat.min(), ds_sub.rlat.max()], crs=transform)
ax.gridlines(draw_labels=True, linewidth=0.5, color='gray',
xlocs=range(-180,180,1), ylocs=range(-90,90,1))
da.plot(ax=ax, cmap='terrain', transform=transform, vmin=-200, vmax=1000, x='rlon', y='rlat')
ax.coastlines(resolution='50m', color='black', linewidth=1)
ax.add_feature(cf.BORDERS)
plot(ds.orog.where(mask.MASK.sel(submask='REA'), drop=True))
masked_oro = ds.orog.where(mask.MASK)
masked_oro
masked_oro.sel(submask='REA').plot()
# ### combination of all regions
sum_mask = mask.MASK.sum(dim='submask')
plot(ds.orog.where(sum_mask, drop=True))
ds.orog.where(sum_mask, drop=True).plot(cmap='terrain')
rea = masked_oro.sel(submask='REA').plot()
ds.orog.where(mask.MASK.sel(submask='REA'), drop=True).plot(cmap='terrain', x='lon', y='lat')
# ### plot all regions on lat lon grid
# +
import matplotlib.pyplot as plt
# Subplots are organized in a Rows x Cols Grid
# Tot and Cols are known
Tot = len(mask.submask)
Cols = 4
# Compute Rows required
Rows = Tot // Cols
Rows += Tot % Cols
# Create a Position index
Position = range(1,Tot + 1)
# +
# Create main figure
import cartopy.crs as ccrs
import cartopy.feature as cf
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(30, 25))
transform = ccrs.RotatedPole(pole_latitude=pollat, pole_longitude=pollon)
for k in range(Tot):
# add every single subplot to the figure with a for loop
ax = fig.add_subplot(Rows,Cols,Position[k], projection=transform)
ax.gridlines(draw_labels=True, linewidth=0.5, color='gray',
xlocs=range(-180,180,1), ylocs=range(-90,90,1))
ax.coastlines(resolution='50m', color='black', linewidth=1)
ax.add_feature(cf.BORDERS)
subregion = mask.MASK.isel(submask=k)
ds.orog.where(subregion, drop=True).plot(ax=ax, cmap='terrain', x='rlon', y='rlat', transform=transform)
| nukleus-masks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # Vitessce Widget Tutorial
# -
# # Export data to local files
# ## 1. Import dependencies
#
# We need to import the classes and functions that we will be using from the corresponding packages.
# +
import os
import json
from urllib.parse import quote_plus
from os.path import join
from urllib.request import urlretrieve
from anndata import read_h5ad
import scanpy as sc
from vitessce import (
VitessceWidget,
VitessceConfig,
Component as cm,
CoordinationType as ct,
AnnDataWrapper,
)
# -
# ## 2. Download and process data
#
# For this example, we need to download a dataset from the COVID-19 Cell Atlas https://www.covid19cellatlas.org/index.healthy.html#habib17.
# +
os.makedirs("data", exist_ok=True)
adata_filepath = join("data", "habib17.processed.h5ad")
urlretrieve('https://covid19.cog.sanger.ac.uk/habib17.processed.h5ad', adata_filepath)
adata = read_h5ad(adata_filepath)
sc.pp.highly_variable_genes(adata, n_top_genes=100)
# -
# ## 3. Create the Vitessce configuration
# Set up the configuration by adding the views and datasets of interest.
vc = VitessceConfig(name='Habib et al', description='COVID-19 Healthy Donor Brain')
dataset = vc.add_dataset(name='Brain').add_object(AnnDataWrapper(adata, cell_set_obs_cols=["CellType"]))
scatterplot = vc.add_view(dataset, cm.SCATTERPLOT, mapping="X_umap")
cell_sets = vc.add_view(dataset, cm.CELL_SETS)
genes = vc.add_view(dataset, cm.GENES)
heatmap = vc.add_view(dataset, cm.HEATMAP)
vc.layout((scatterplot | (cell_sets / genes)) / heatmap);
# ## 4. Export files to a local directory
#
# The `.export(to='files')` method on the view config instance will export files to the specified directory `out_dir`. The `base_url` parameter is required so that the file URLs in the view config point to the location where you ultimately intend to serve the files.
config_dict = vc.export(to='files', base_url='http://localhost:3000', out_dir='./test')
# ## 5. Serve the files
# Now that the files have been saved to the `./test` directory, they can be served by any static web server.
#
# If you would like to serve the files locally, we recommend [http-server](https://github.com/http-party/http-server) which can be installed with NPM or Homebrew:
# ```sh
# # cd test
# http-server ./ --cors -p 3000
# ```
# ## 6. View on vitessce.io
#
# The returned view config dict can be converted to a URL, and if the files are served on the internet (rather than locally), this URL can be used to share the interactive visualizations with colleagues.
vitessce_url = f"http://vitessce.io/?url=data:," + quote_plus(json.dumps(config_dict))
print(vitessce_url)
| docs/notebooks/data_export_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy import integrate
import matplotlib.pyplot as plt
## simple model parameter for air density and surface area
r = 0.05
c = 0.47
rho_air = 1.28
A = np.pi * r**2
## parameter for resistance, mass and gravity
k = 0.5 * c * rho_air * A
m = 0.2
g = 9.81
## Key in the ball throwing problem with air resistance
## Consider two direction, x and y[hight]
def Air_resistance(t, para):
x, vx, y, vy = para
v = (vx**2+ vy**2)**(1/2)
ax = -k/m * v * vx
ay = -k/m * v * vy - g
return vx, ax, vy, ay
## Initial conditions
v0 = 40
phi0 = np.radians(70)
para = 0, v0 * np.cos(phi0), 0., v0 * np.sin(phi0)
t0, tf = 0, 15
## Solve the differential equation
soln = integrate.solve_ivp(Air_resistance, (t0, tf), para, dense_output=True)
## Get a dense output
t = np.linspace(0, 15, 1000)
sol_finer = soln.sol(t)
x, vx, y, vy, = sol_finer[0], sol_finer[1], sol_finer[2],sol_finer[3]
## Comparison of my solver with Scipy solver
x_0, vx_0, y_0, vy_0 ,Dt = 0, 40 * np.cos(phi0), 0., 40 * np.sin(phi0), 0.05
for i in range(200):
v_0 = (vx_0**2+ vy_0**2)**(1/2)
ax_0 = -k/m * v_0 * vx_0
ay_0 = -k/m * v_0 * vy_0 - g
vx_0 = vx_0 + ax_0*Dt
vy_0 = vy_0 + ay_0*Dt
x_0 = x_0 + vx_0*Dt
y_0 = y_0 + vy_0*Dt
plt.figure(0)
plt.plot(x_0,y_0,'o')
## Plot out x versus y figure
plt.figure(0)
plt.plot(x, y, label = "speed - dense output")
plt.plot(soln.y[0], soln.y[2], label = "speed")
plt.xlabel('x [meter]')
plt.ylabel('y [meter]')
plt.ylim(0, 60)
plt.xlim(.0, 80)
plt.title(r'Air resistance simulation $F_D = \frac{-1}{2} c \rho A v|v|$')
plt.legend(loc='upper right')
plt.grid(True)
plt.show()
## Plot out velocity in x and y direction
plt.figure(2)
plt.plot(t,sol_finer[1],'red', label = "The velocity in x direction")
plt.plot(t,sol_finer[3],'orange', label = "The velocity in y direction")
plt.xlabel('Time [second]')
plt.ylabel('Velocity [meter/second]')
plt.title("The velocity varience with time")
plt.legend(loc='upper right')
plt.grid(True)
plt.show()
# -
| HW6/HW6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
x = np.linspace(0, 10, 20) # linspace is another way we can create an array
# it take three args with there usage as followed
# 1st- starting point
# 2nd- ending point
# 3rd- number of points you want between the starting and the ending point
x
y = np.sin(x)
plt.plot(x,y) # how to plot a linechart
plt.show() # it helps us combine all the attributes and then show them together
plt.plot(x,y)
plt.xlabel('Time') # givinv a label to the x-axis
plt.ylabel("sin(x)") # giving a label to the y-axis
# here we can combine all the fields and a plot.show() would let you
# show the whole plot together
plt.plot(x,y)
plt.xlabel("time")
plt.ylabel("sin(x)")
plt.title("My cool chart") # adding a title to the plot
plt.show()
plt
| MatplotlibBasics/Linechart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="6ApTDxdPVgiI"
# # Scene Basic
#
# * Author: <NAME>
# * Scene for McThings framework: https://github.com/juntosdesdecasa/mcthings (0.50.0)
# * License: ASL 2.0
# * Description of the scene: The first scene in McThings history
# + colab={} colab_type="code" id="MUVPF1KoVgiK"
# Install McThings to create the Scene
# !pip install mcthings_extra --upgrade
# + colab={} colab_type="code" id="DDRRDm1OVgib"
# Import definition of blocks
import mcpi.block
# Import Vec3 for defining positions
from mcpi.vec3 import Vec3
# Be sure to use the last version
import importlib
import mcthings
importlib.reload(mcthings)
# Import Scene and Server
from mcthings.renderers.raspberry_pi import RaspberryPi
from mcthings.scene import Scene
from mcthings.world import World
# Import the Things used in the Scene
from mcthings.bridge import Bridge
from mcthings.house import House
from mcthings.river import River
from mcthings.schematic import Schematic
# + colab={} colab_type="code" id="VITvEf3PVgip"
# Connect to the Minecraft server and send a testing message to chat
MC_SEVER_HOST = "localhost"
MC_SEVER_PORT = 4711
World.renderer = RaspberryPi(MC_SEVER_HOST, MC_SEVER_PORT)
World.renderer.post_to_chat("Building a Basic Scene")
server = World.renderer.server.mc
# + colab={} colab_type="code" id="oLBbbTm3Vgiz"
# Get the initial position to create the scene
BUILDER_NAME = "ElasticExplorer"
pos = World.renderer.get_pos(BUILDER_NAME)
pos.x += 1
World.renderer.post_to_chat("Building the Scene at %s %s %s " % (pos.x, pos.y, pos.z))
# + [markdown] colab_type="text" id="g4jV-8FLVgjD"
# # The order in which the scene will be built
#
# * Create House
# * Create River
# * Create Bridge
# * Create House
# + colab={} colab_type="code" id="OuDmjbxkVgjF"
# Commons params for the Scene
river_width = 10
house_to_river = 5
house_width = 5
# -
# The first house
house = House(pos)
house.mirror = True
house.width = house_width
house.build()
# Create a river between the houses
pos.x += house_to_river + 1
river = River(pos)
river.width = river_width
river.build()
# Create a bridge over the river
pos.x -= 1
bridge = Bridge(pos)
bridge.large = river_width + 2
bridge.block = mcpi.block.STONE
bridge.build()
# + colab={} colab_type="code" id="yzZZXoEkVgjP"
# The last house
pos.x = river.end_position.x + 1 + house_to_river
house = House(pos)
house.width = house_width
house.build()
# -
# Let's persist the scene
World.scenes[0].save("mct/scene_basic.mct")
# Let's save it to a Schematic
World.scenes[0].to_schematic("schematics/scene_basic.schematic")
# Let's load the Schematic to test it
schematic = Schematic(Vec3(pos.x + 2, pos.y, pos.z))
schematic.file_path = "schematics/scene_basic.schematic"
schematic.build()
# + [markdown] colab_type="text" id="G3_0MH_oVgkM"
# Include a screenshot with the scene built
#
# 
| notebooks/scene_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decision trees (DTs)
#
# Hello and welcome to this workshop in which we will build together our first decision tree model. In this workshop we are going to create complexe tree and forest
# to solve classification problems.
#
# **What you will learn:**
# - Creation of decision tree models.
# - How to train and optimize a model.
# - Introduction to random forest.
# - Analyzing model results.
# ## 1 - Packages ##
#
# Please make sure you have the following programs installed:
#
# - [Sklearn](http://scikit-learn.org/stable/) Simple and efficient tools for predictive data analysis.
# - [Matplotlib](http://matplotlib.org) Matplotlib is a comprehensive library for creating static, animated, and interactive visualizations in Python.
# - [Numpy](https://numpy.org/) The fundamental package for scientific computing with Python
import numpy as np
import sklearn
import sklearn.datasets
import sklearn.linear_model
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
# %matplotlib inline
# ## 2 - Dataset ##
#
# First of all you must understand how to use the most important thing in machine leaning, the data.
#
# To get started, let's get the data set. The following code will load your first data
flowers = load_iris()
X = flowers.data
Y = flowers.target
# **Now that you have the dataset load we must analyze our data further.**
#
# Your data set is slip into two important things, the data and the labels.
# The data is the information about a state of an object and the label is the thing we are predicting. The label could be the future price of wheat, the kind of animal shown in a picture or in our case the kind of Iris
#
# To train a model you must split your data into 2 batches. One to train and another to test your model.
#
# **Exercise**: You must extract the data and the label from the dataset thanks to the " train_test_split " fonc and **print** their shape
#
# **Help:** The train_test_split need 3 parameters and return 4 split data , you should look at this link [Data Split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) and give it a random state of 10 the data X and the target Y
#
# +
#_need_data_container_# = #_ Need data split
#Print the shape
# -
# **Wanted output**:
#
# <table style="width:20%">
# <tr>
# <td>shape de X_train</td>
# <td> (112, 4) </td>
# </tr>
#
# <tr>
# <td>shape de x_test</td>
# <td>(38, 4) </td>
# </tr>
#
# <tr>
# <td>shape de y_train</td>
# <td>(112,) </td>
# </tr>
# <tr>
# <td>shape de y_test</td>
# <td>(38,) </td>
# </tr>
# </table>
# **If the output are similar, your data is ready to be use**
# ## 3 - Build Decision Tree
# To build our first Decision tree, you are going to use a very useful function "DecisionTreeClassifier" from the sklearn library. [Tree Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html?highlight=decisiontreeclassifier#sklearn.tree.DecisionTreeClassifier)
# ### 3.1 - Create tree ####
# +
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(random_state=0)
# -
# We have created the tree, now we need to fit it with our datas and labels (X_train, y_train).
#
# The "fit" method consists in creating the most adequate prediction model for the data given to it as parameter, this is one of the most important method of the class
#
# **Help**: you should look at tree.fit() method here [Tree Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html?highlight=decisiontreeclassifier#sklearn.tree.DecisionTreeClassifier.fit)
# +
# need one line to fit tree
# -
# **Wanted output**:
#
# <table style="width:20%">
# <tr>
# <td>DecisionTreeClassifier(random_state=0)</td>
# </tr>
# </table>
# ### 3.2 - Display tree ####
# +
from src.tree.tree_fonc import *
create_graph_tree(tree, flowers.target_names, flowers.feature_names)
display_tree("tree.dot")
# -
from IPython.display import Image
Image(filename='tree.png')
# **As you can see, you create a tree we various branches an leaf and a depth of six.**
#
# You can now see how the model is going to predict the kind of iris for each picture depending on the petal length or width.
# ### 3.3 - Test our model ####
#
# After the train phase, we can test our model. To test a tree model we use the score() build in method
#
print("Accuracy on training set: {:.3f} / 1.000".format(tree.score(X_train, y_train)))
print("Accuracy on test set: {:.3f} / 1.000".format(tree.score(X_test, y_test)))
# The output show 1 / 1 of accuracy for train set meanings that he can now find the best label for 100% of the train data and for 97% of the test batch.
#
# **Those result are good, but does the tree grown the right way? We are going to analyse these results....**
# ### 3.4 - Analyze the Decision Tree ####
#
# The goal of machine learning is to create models who must not only fit the training data well, but also accurately classify records it has never seen.
#
# When a decision tree is fully grown, it may lose some generalization capability because of the complexity of the train set. We call this the [Overfitting](https://en.wikipedia.org/wiki/Overfitting)
#
# In our case, we can see this phenomenon in the last leaves of the tree. We get leaf with only 1 samples. Meaning thas is a very particular case who can be a dataset error or genetic anomaly.
#
# We need to find a solution to make our tree more generalize. We should take a look at the "max_depth" argument in the [DecisionTreeClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html?highlight=decisiontreeclassifier#sklearn.tree.DecisionTreeClassifier) function
# **Exercise**: Rebuild the tree with more paramaters to avoid Overfitting
# +
##Use DecisionTreeClassifier again.
##You need one more line fit the tree again
# -
create_graph_tree(tree, flowers.target_names, flowers.feature_names)
display_tree("tree.dot")
Image(filename='tree.png')
# ##You must now have a tree with a max deep of threec##
# Let see if our model score change.
print("Accuracy on training set: {:.3f} / 1.000".format(tree.score(X_train, y_train)))
print("Accuracy on test set: {:.3f} / 1.000".format(tree.score(X_test, y_test)))
# We can see that with a tree with half branch the accuracy result is as well as the deeper one. That means that when the we train the tree at first we create a useless branch
# **Résultat attendu**:
#
# <table style="width:90%">
# <tr>
# <td>Accuracy on training set:</td>
# <td> 0.964 / 1.000</td>
# </tr>
# <tr>
# <td>Accuracy on test set:</td>
# <td> 0.974 / 1.000</td>
# </tr>
# </table>
#
#
# +
from matplotlib import pyplot as plt
def plot_feature_importance(model, dataset):
n_features = dataset.data.shape[1]
plt.barh(np.arange(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), dataset.feature_names)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
plot_feature_importance(tree, flowers)
# -
# This graph shows the use of the different information about the dataset.
#
#
#
# **Conclusion**
#
# Well done you have created and trained your own decision tree.
# We have optimized the results of our tree using a max depth variable to avoid overfitting.
#
# Generally dealing with the max depth variable is enough to keep a good accuracy with simple data set, but when you use more complexes one the performance becomes quickly very bad.
#
# Then we going to use **ensemble methods**
# ## 4 - Ensembles of Decision Trees ##
#
# The goal of [ensemble methods](https://scikit-learn.org/stable/modules/ensemble.html) is to combine the predictions of several base estimators built with a given learning algorithm in order to improve generalization and robustness.
#
# We will see 2 of the most famous ensembles trees:
#
# <table style="width:90%">
# <tr>
# <td>Random Forest</td>
# <td>(Regressor / Classifier)</td>
# </tr>
# <tr>
# <td>Gradient Boost</td>
# <td>(Regressor / Classifier)</td>
# </tr>
# </table>
# ### 4.1 build Random Forest
# As we have seen in previous steps, decision tree can overfit really quickly. To avoid that, we are going to train several trees and compare their results to get better accuracy for every dataset.
#
# We are now going to use a new and more complex dataset:
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import load_breast_cancer
# **Exercise**: You now need to load the dataset and split it like we did before.
cancer = load_breast_cancer()
X = cancer.data
Y = cancer.target
# **Wanted output**:
#
# <table style="width:20%">
# <tr>
# <td>shape de X_train</td>
# <td>(426, 30)</td>
# </tr>
#
# <tr>
# <td>shape de x_test</td>
# <td>(143, 30)</td>
# </tr>
#
# <tr>
# <td>shape de y_train</td>
# <td>(426,)</td>
# </tr>
# <tr>
# <td>shape de y_test</td>
# <td>(143,)</td>
# </tr>
# </table>
# **Exercise**: Let build your forest with the [RandomForestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html) and fit it with the Cancer dataset.
#
# **Help**: You must set the number of trees to 5 with n_estimators, the random_state to 2 and chose a max_depth value for the trees
##need one line to create the forest
forest = RandomForestClassifier()## Need args
##need one line to fit the forest tree with X_train and y_train
# +
from src.plotlib.plot_fonc import *
plot_forest(forest, cancer)
# -
# <img src="fire3_PIL.gif" width="750" align="center">
# There is a plot of all tree in the forest
print("Accuracy on training set: {:.3f} / 1.000".format(forest.score(X_train, y_train)))
print("Accuracy on test set: {:.3f} / 1.000".format(forest.score(X_test, y_test)))
# We have very good accuracy who is guaranteed to be generalist thanking to the tree merges.
plot_feature_importance(forest, cancer)
# As we can see on the graph must more information are used in forest to predict results.
#
# **Conclusion**
#
# Random forests are the best way to quickly classify datasets.
#
# To increase the forest accuracy you can play with the number of trees or branches. But be careful, random forest are **" random "** so if you change the "random_stats" the result will probably move a lot.
#
# You made your first forest **grow up**, well done. Now we will look at a more complex model who outperforms random forest thank to loss function...
# ### 4.2 Gradient Boosting tree ###
#
# The gradient boosting is a general technique which consists of aggregating classifiers (trees) train sequentially on a learning dataset whose individual prediction are corrected at each
# step. Classifiers are weighted according to their performance.
#
# So more a model predicts a bad answer, the more it will be corrected and vice versa. We call that [Gradient descent](https://en.wikipedia.org/wiki/Gradient_descent)
#
# A chance for us, the library sklearn does those calculations for us.
#
# **Exercise**: Let build your Gradient Boosting tree with the [GradientBoostingClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html?highlight=gradient#sklearn.ensemble.GradientBoostingClassifier) and fit it with the Cancer dataset (X_train, y_train).
#
# **Help**: By default the Gradient Boosting tree has a depth of three
# +
from sklearn.ensemble import GradientBoostingClassifier
##one line to init the GradientBoostingClassifier
gradient_tree = ##Need function
##give the train data and labels to fit the tree
# -
print("Accuracy on training set: {:.3f} / 1.000".format(gradient_tree.score(X_train, y_train)))
print("Accuracy on test set: {:.3f} / 1.000".format(gradient_tree.score(X_test, y_test)))
# If you have correctly set up the model you should have very good results in the training and the test set.
#
# If you have an accuracy of 1.000 is probably due to an overfitting. You must correct that with the [learning_rate](https://en.wikipedia.org/wiki/Learning_rate) argument from [GradientBoostingRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingRegressor.html)
# **Conclusion**:
#
# As we can see, the results are approximately the same as the **Random forest**. However the gradient leaves us a greater freedom of adaptation which allows to manage a greater number of cases and need less branch to work well, therefore accelerate the learning process.
# ## 5 - End ##
#
# Well done, you had completed each of the points of this workshop. You had acquired the necessary skills to build decision trees for your own dataset. I now encourage you to find a dataset that you like and to build a decision tree on your own. [Dataset](https://scikit-learn.org/stable/datasets.html)
#
# +
## You can code
| ai/8.DecisionTree/decisionTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shapes counting
#
#
# ## Task
# **Geometic shapes counting**: network outputs $10$ probabilities for each class, representing different numbers of objects of this class on the image. So the network should have $60$ outputs. Outputs $0..9$ should sum up to $100\%$, so outputs $10..19$, and so on. The loss function for the network is the sum of squared counting errors:
# $$J = \sum_{i=0}^5 \sum_{j=0}^9\hat y_j^i (j - r^i)^2$$ *Notation:* $r^i$ is a ground truth, $\hat y_j^i$ is predicted probability of $j$ figures of class $i$ on the image.
# ## Network
#
# At first I tried to use the same model that for the [classification task](classification.ipynb).
# But it didn't even achieve any sensible loss, so I decided to make a larger model.
class ShapesCounter(nn.Module):
def __init__(self):
super(ShapesCounter, self).__init__()
self.conv1 = nn.Sequential(
nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(2, 2),
padding=(1, 1)),
nn.BatchNorm2d(4),
nn.ReLU()
)
self.conv2 = nn.Sequential(
nn.Conv2d(in_channels=4,
out_channels=16,
kernel_size=(3, 3),
padding=(1, 1)),
nn.BatchNorm2d(16),
nn.ReLU()
)
self.conv3 = nn.Sequential(
nn.Conv2d(in_channels=16,
out_channels=32,
kernel_size=(4, 4),
padding=(1, 1)),
nn.BatchNorm2d(32),
nn.ReLU()
)
self.conv4 = nn.Sequential(
nn.Conv2d(in_channels=32,
out_channels=64,
kernel_size=(5, 5),
padding=(1, 1)),
nn.BatchNorm2d(64),
nn.ReLU()
)
self.conv5 = nn.Sequential(
nn.Conv2d(in_channels=64,
out_channels=128,
kernel_size=(6, 6),
stride=(2, 2),
padding=(1, 1)),
nn.BatchNorm2d(128),
nn.ReLU()
)
self.conv6 = nn.Sequential(
nn.Conv2d(in_channels=128,
out_channels=256,
kernel_size=(7, 7),
padding=(1, 1)),
nn.BatchNorm2d(256),
nn.ReLU()
)
self.conv7 = nn.Sequential(
nn.Conv2d(in_channels=256,
out_channels=512,
kernel_size=(8, 8)),
nn.BatchNorm2d(512),
nn.ReLU()
)
self.fc1 = nn.Sequential(
nn.Linear(512, 256),
)
def forward(self, x):
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.conv4(x)
x = self.conv5(x)
x = self.conv6(x)
x = self.conv7(x)
x = x.view(x.shape[0], -1)
x = self.fc1(x)
x = x.view(x.shape[0], 6, 10)
return x
# +
import json
import matplotlib.pyplot as plt
import numpy as np
def plot_training(run_hist):
"""Plot the training history of the classification model."""
fig, ax = plt.subplots(1, 2, figsize=(20, 6), sharex=True)
x = np.arange(len(run_hist["train/loss"])) + 1
ax[0].plot(x, run_hist["train/loss"], 'b', marker='.', label="epoch train loss")
ax[0].plot(x, run_hist["validation/loss"], 'r', marker='.', label="epoch test loss")
ax[0].legend()
ax[1].plot(x, run_hist["train/acc"], 'b', marker='.', label="epoch train accuracy")
ax[1].plot(x, run_hist["validation/acc"], 'r', marker='.', label="epoch test accuracy")
ax[1].legend()
fig, ax = plt.subplots(1, 1, figsize=(20, 6), sharex=True)
x = np.arange(len(run_hist["train/batch_loss"])) + 1
ax.plot(x, run_hist["train/batch_loss"], 'b', marker='.', label="batch train loss")
ax.legend()
# -
with open('charts/counter/basic.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
# We can see on the chart that it didn't went good. The loss is decreasing but accuracy is still 0.
# Probably our net is still too small and it can't learn more complex relations.
# Before we try adding more layers let's try the `BatchNorm` trick from classification.
# ## Batch Normalization after last layer
with open('charts/counter/batch_norm_last.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
# The accuracy went up to 45%! It's a great result, but maybe we can improve it.
# ## More linear layers
#
# Same network with 2 linear layers. `512 -> 256 -> 60`
with open('charts/counter/new_layer.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
# The results are slightly better, but the network is overfitting more then the one before.
# ## 3 linear layers
#
# I added one more layer out of curiosity `512 -> 256 -> 256 -> 60`
with open('charts/counter/3_layers.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)cd
# ## Network with 135 outputs
#
# This network has 135 outputs, because that's the different results of counting figures (${6 \choose 2}9=135$).
# We transformed this into classification task so I used `nn.CrossEntropyLoss()` as a loss function.
class ShapesCounter135(nn.Module):
def __init__(self):
super(ShapesCounter135, self).__init__()
self.conv1 = nn.Sequential(
nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(2, 2),
padding=(1, 1)),
nn.BatchNorm2d(4),
nn.ReLU()
)
self.conv2 = nn.Sequential(
nn.Conv2d(in_channels=4,
out_channels=16,
kernel_size=(3, 3),
padding=(1, 1)),
nn.BatchNorm2d(16),
nn.ReLU()
)
self.conv3 = nn.Sequential(
nn.Conv2d(in_channels=16,
out_channels=32,
kernel_size=(4, 4),
padding=(1, 1)),
nn.BatchNorm2d(32),
nn.ReLU()
)
self.conv4 = nn.Sequential(
nn.Conv2d(in_channels=32,
out_channels=64,
kernel_size=(5, 5),
padding=(1, 1)),
nn.BatchNorm2d(64),
nn.ReLU()
)
self.conv5 = nn.Sequential(
nn.Conv2d(in_channels=64,
out_channels=128,
kernel_size=(6, 6),
stride=(2, 2),
padding=(1, 1)),
nn.BatchNorm2d(128),
nn.ReLU()
)
self.conv6 = nn.Sequential(
nn.Conv2d(in_channels=128,
out_channels=256,
kernel_size=(7, 7),
padding=(1, 1)),
nn.BatchNorm2d(256),
nn.ReLU()
)
self.conv7 = nn.Sequential(
nn.Conv2d(in_channels=256,
out_channels=512,
kernel_size=(8, 8)),
nn.BatchNorm2d(512),
nn.ReLU()
)
self.fc1 = nn.Sequential(
nn.Linear(512, 135),
)
def forward(self, x):
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.conv4(x)
x = self.conv5(x)
x = self.conv6(x)
x = self.conv7(x)
x = x.view(x.shape[0], -1)
x = self.fc1(x)
return x
with open('charts/counter/135_output.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
# Model is overfitting, while loss is nearly at zero.
# ## Batch normalization after last layer
with open('charts/counter/135_batch_norm.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
with open('charts/counter/135_batch_norm_larger.json', 'r') as f:
hist = json.loads(f.read())
plot_training(hist)
# The last model appears to give the best result of **51.1%**. Both models (output 60 or 135) can be found [here](models/shapes_counter.py).
| counter.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// ## Introduction
// **Goals:** This notebook is meant to serve as an introduction to [Scala](http://scala-lang.org/) and [Apache Spark](https://spark.apache.org/).
//
// ## Scala
// This is a basic introduction to assignments, IO, and functions in Scala.
// A simple variable assignment
val x = 10
// We can print values to standard output
println(x)
// A simple function declaration to add two integers
def add(a: Int, b: Int) = {
a + b
}
add(1,5)
// ## Spark Intro
// Toree creates several objects used to interact with Spark. The most common is the [SparkContext](https://spark.apache.org/docs/latest/api/scala/index.html#org.apache.spark.SparkContext) object which is bound to the variable `sc`.
// We create a Spark RDD of the numbers 1 to 100
val numbersRDD = sc.parallelize(1 to 100)
numbersRDD.collect()
// We map the numbers to be a tuple of the number and a boolean stating if it is divisble by two
val mappedNumbersRDD = numbersRDD.map((num: Int) => {
(num, num % 2 == 0)
})
mappedNumbersRDD.collect()
// We filter the RDD to get only the even numbers
val evenNumbersRDD = mappedNumbersRDD.filter((tuple: (Int, Boolean)) => {
tuple._2
})
// Collect the numbers for output and them out with a message
evenNumbersRDD.collect().foreach( x => println(s"${x._1} is an even number"))
| work/1-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 2: Some Simple Spin Glasses
#
# We'll now investigate some simple spin glasses. Your goal is to devise an annealing schedule (or perhaps employ some other clever solution) to find the ground state of the given Ising Hamiltonian.
# +
import numpy as np
from ising_animator import IsingAnimator
from abstract_ising import AbstractIsing
# %matplotlib inline
# -
# ## 1D Random Bond Ising on a Periodic Chain
#
# $$H = J \sum_{\langle ij \rangle} B_{ij} \sigma_i \sigma_j$$
#
# where $B_{ij} = \pm 1$ which is selected randomly and independently for each bond nearest neighbour bond $ij$ when the model is initialized.
#
# Due to the intrinsic randomness of the Hamiltonian parameters, it is generally quite difficult to find the ground state of this model.
class RandomBondIsing1DPBC(AbstractIsing):
def __init__(self, N, J=1.):
self.J, self.N = J, N
self.num_spins = self.N
self.bonds = 2*(np.random.rand(self.N) < 0.5) - 1
# initialize system at infinite temperature
# i.e. spins are completely random and uncorrelated
self.spins = 2*(np.random.rand(self.N) < 0.5) - 1
def energy(self, spins=None):
"""Returns the energy of the current spin configuration"""
spins = self.spins if spins is None else spins
interactions = self.bonds * spins * np.roll(spins, 1, axis=-1)
total = self.J * np.sum(interactions, axis=-1)
return total
def energy_diff(self, i):
"""Returns the energy difference resulting from flipping the i'th site"""
# sum the nearest neighbour sites
nn_sum = (
self.bonds[i] * self.spins[i-1]
+ self.bonds[(i+1)%self.N] * self.spins[(i+1) % self.N]
)
return -2 * self.J * self.spins[i] * nn_sum
def rand_site(self):
"""Selects a site in the lattice at random"""
return (np.random.randint(self.N),)
ising = RandomBondIsing1DPBC(10, J=1)
ising.spins
ising.bonds
ising.energy()
# perform 1000 MC steps
for t in range(1000):
# take a look at the abstract_ising.py file to see how mc_step works
E = ising.mc_step(T=1.0)
if t % 50 == 0:
print(E)
ising.spins
T = 1.0
N = 1000
# +
# reinitialize so we're back at a T=infinity state
ising = RandomBondIsing1DPBC(10, J=1)
print(ising.spins) # print the starting configuration
# NOTE: sometimes the animation gets stuck displaying a single image
# The most reliable way to fix this is by restarting the notebook.
# Initializing the Ising Model in the same cell as the one where you
# run the animation also seems to work.
IsingAnimator(ising).run_animation([T] * N)
# -
dim = np.arange(2 ** ising.num_spins)
space = ((dim[:, None] & (1 << np.arange(ising.num_spins))) > 0)
space = 2*space.astype(int) - 1
ising.energy(space).min()
# Now of course, we're limited in the size of the systems for which we can compute this energy exactly, hence why we need an annealing procedure which (we hope) would help us find the ground state of the Ising model of interest (or at least a state that is close enough).
# Your task is to come up with an annealing procedure to help find the ground state (or something close enough) of this model for several different chain sizes: 10, 20, 50, 100.
#
# For the small systems, compute the ground state energy exactly, and compare it to the result of your annealer.
#
# Since it's impossible to find the exact ground state for large systems, you will be evaluated on your algorithm's performance on the small systems, as well as the overall ingenuity of your method.
# your solution here
ising = RandomBondIsing1DPBC(100, J=1.)
# # Fully Connected Random Bond Ising
#
# The Fully Connected Random Bond Ising Hamiltonian takes the form:
#
# $$H = J \sum_{i<j} B_{ij} \sigma_i \sigma_j$$
#
# where $B_{ij} = \pm 1$ which is selected randomly (uniformly) and independently for each bond $ij$ when the model is initialized.
class FullyConnectedRandomBondIsing(AbstractIsing):
def __init__(self, N, J=1.):
self.J, self.N = J, N
self.num_spins = self.N
# initialize system at infinite temperature
# i.e. spins are completely random and uncorrelated
self.spins = 2*(np.random.rand(self.N) < 0.5) - 1
self.bonds = np.zeros((self.N, self.N))
for i in range(self.N):
for j in range(i+1, self.N):
self.bonds[i, j] = 2*(np.random.rand() < 0.5) - 1
def energy(self):
"""Returns the energy of the current spin configuration"""
interaction = self.spins.dot(self.bonds.dot(self.spins))
return self.J * interaction
def energy_diff(self, i):
"""Returns the energy difference resulting from flipping the i'th site"""
return -2 * self.J * self.spins[i] * (
self.bonds[i, :].dot(self.spins)
+ self.bonds[:, i].dot(self.spins)
)
def rand_site(self):
"""Selects a site in the lattice at random"""
return (np.random.randint(self.N),)
ising = FullyConnectedRandomBondIsing(10, J=1)
ising.spins
ising.energy()
IsingAnimator(ising).run_animation([T] * n_steps)
ising.energy()
dim = np.arange(2 ** ising.num_spins)
space = ((dim[:, None] & (1 << np.arange(ising.num_spins))) > 0)
space = 2*space.astype(int) - 1
np.einsum("bi,ij,bj->b", space, ising.bonds, space).min()
# Repeat the previous problem with the Fully Connected Random Bond Ising Model for the same system sizes (10, 20, 50, 100).
# +
# your solution here
# -
# # The Fully Connected Mattis Model
#
# Another model of interest is the Mattis Model which is a specific instance of a Random Bond Ising Model:
#
# $$H = J\sum_{i<j} B_{ij}\sigma_i\sigma_j$$
#
# however, the random bonds take the form:
#
# $$B_{ij} = -\xi_i \xi_j$$
#
# where $\xi_i = \pm 1$ with the sign selected randomly (uniformly) when the model is initialized.
class FullyConnectedMattisModel(FullyConnectedRandomBondIsing):
def __init__(self, N, J=1.):
self.J, self.N = J, N
self.num_spins = self.N
self.xi = 2*(np.random.rand(self.N) < 0.5) - 1
# initialize system at infinite temperature
# i.e. spins are completely random and uncorrelated
self.spins = 2*(np.random.rand(self.N) < 0.5) - 1
self.bonds = np.zeros((self.N, self.N))
for i in range(self.N):
for j in range(i+1, self.N):
self.bonds[i, j] = -self.xi[i] * self.xi[j]
ising = FullyConnectedMattisModel(10, J=1)
ising.spins
ising.bonds
ising.energy()
T = 0.1
# perform 1000 MC steps at low temperature to find the ground state
# it's fairly easy to find in this case as we'll discuss soon
for t in range(1000+1):
# take a look at the abstract_ising.py file to see how mc_step works
E = ising.mc_step(T=T)
if t % 50 == 0:
print(E)
ising.spins
dim = np.arange(2 ** ising.num_spins)
space = ((dim[:, None] & (1 << np.arange(ising.num_spins))) > 0)
space = 2*space.astype(int) - 1
np.einsum("bi,ij,bj->b", space, ising.bonds, space).min()
# While at first glance the `bonds` matrix for the Mattis Model appears to produce the glassy model, it is in fact not a glass at all. We can convert this into a simple fully connected Ising Model using a *gauge transformation*.
# But first, make sure the MC simulation has converged to the ground state energy we computed exactly in the previous cell. If it hasn't done so yet, let it run a few more times.
# Now let's look at our ground state spin configuration:
ising.spins
# Next we'll look at the Mattis parameters $\xi$
ising.xi
# If we did indeed find the ground state, the spins should match with $\xi$ (upto a global sign). This gives us a hint as to what this Mattis Model actually is.
# Let's look again at the Hamiltonian:
#
# $$H = J \sum_{i<j} B_{ij} \sigma_i \sigma_j = -J\sum_{i < j} \xi_i \xi_j \sigma_i \sigma_j = -J\sum_{i < j} (\xi_i \sigma_i) (\xi_j\sigma_j)$$
#
# We introduce some new variables $s_i = \xi_i \sigma_i \in \lbrace 0, 1 \rbrace$. Then our Hamiltonian becomes:
#
# $$H = -J \sum_{i < j} s_i s_j$$
#
# which is just a fully connected Ising Model. If we compute $s_i$ for the state found in the previous simulation, we get:
ising.spins * ising.xi
# which is one of the ground states of the fully connected Ising Model.
# As an (unmarked) bonus problem, you can apply the annealing algorithm you developed in the two previous parts to this model.
# At the end of the annealing procedure, confirm that you've found the ground state by checking that $\xi_i$ is equal to $\sigma_i$ upto a global sign.
| Project_4_Ising_Annealer/Task_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Basics of Pinocchio
#
# Welcome in Pinocchio, a C++ library with Python API to efficiently compute all the elements of the model of a rigid robot, and their derivatives. Pinocchio is nicely tailored for generating the motion of a robot using a optimization program.
#
# Let's start by loading and display the robot model.
import pinocchio
import numpy as np
from numpy.linalg import norm,inv,pinv,svd,eig
# You can always alias the long pinocchio namespace to something shorter like pino. In the tuto, we keep the long name for clarity, feel free to shorten it.
# ## Loading the robot
#
# Pinocchio offers several possibilities to load the model of the robot as a kinematic tree of joints, masses, geometry object and other informative tree "decoration": the most classical is to parse the information from a URDF model. Here we will work with the Talos models: a fixed arm and a pair of floating leg. The collection of Talos models can be downloaded in Ubuntu with the APT package *robotpkg-talos-data*.
urdfFile = '/opt/openrobots/share/talos_data/robots/talos_left_arm.urdf'
rmodel = pinocchio.buildModelFromUrdf(urdfFile)
print(rmodel)
# This model as 7 joints, with a configuration space of dimension nq=7. Let's browse quickly the content of the kinematic tree.
# In Pinocchio, we do not store explicitly bodies, but the placement of the joint frame, needed to compute the forward kinematics. We name "universe" the first joint frame, and then name each other frame from the name of the joint. Joint information are then stored with the same numbers, 0 being the universe, 1 the first joint, etc until 8 the last joint.
for i,j in enumerate(rmodel.names): print(i,j)
# We do our best to keep the names of the URDF file, but as we do not have exactly the same convention for representing the kinematic tree, sometime information are not stored.
# Contrary to URDF, our tree is made only of joint. The kinematic order is stored in the parent map rmodel.parent. The type of the joints (revolute X, free flyer, spherical, prismatic, etc) are stored in the rmodel.joints map. Each joint is placed (ie position and orientation) with respect to its parent, and the placement is stored in rmodel.placement.
for i,(j,p,M) in enumerate(zip(rmodel.joints,rmodel.parents,rmodel.jointPlacements)):
print(i,j,"parent=%d"%p,M)
# Masses and inertias are also stored, along with other informations we will discuss later. The dimension of the configuration space is denoted rmodel.nq, while the number of degrees of freedom is rmodel.nv (sometime, nq>nv, here both are equals). The number of joints is given by rmodel.njoints: here as all joints have dimension 1 and we must account for the universe, we have njoints=nq+1.
print(rmodel.nq,rmodel.nv,rmodel.njoints)
# ## The convenient RobotWrapper and the display
# Most of the Python API simply copy the C++ API. We are using Boost::Python to simply copy the API from C++ to Python, with minor re-arragement.
# Only a helper object has been introduced in Python to make the life of the newcommer easier: the RobotWrapper. It is in particular helpful to load completely the model and display it in Gepetto-Viewer. RobotWrapper loads the URDF model but also loads the geometry models for the display and the collision checker. For that, it needs some extra path information that are gathered in the robots.py. Go and have a look at it.
import robots
robot = robots.loadTalosArm()
print(robot.model)
# Finally, it loaded the same robot model than before, but also did other interresting parsing, that we will discover later. Using the extra info we can initialize the display. Start gepetto-gui from a terminal (it has been installed from the 2 packages robotpkg-gepetto-viewer-corba and robotpkg-osg-dae). Gepetto-gui starts a graphic server, and we will open a client to this server in Python using the method implemented in RobotWrapper:
robot.initDisplay(loadModel=True)
# You can see in the GUI window that a blue world has been loaded, with all the meshes of the robot stacked in the middle of the scene. Gepetto-viewer is indeed a rigid-object viewer, that display each mesh at a given placement (gepetto-viewer has no idea of the kinematic chain). You then need pinocchio to compute the placement of all the bodies and place them at the right position and orientation. This is all done in RobotWrapper.
robot.display(robot.q0)
# where robot.q0 is a configuration of the robot. RobotWrapper was designed initially as a reference example where a newcommer can pick basic ways of using Pinocchio. Don't hesitate to go inside this Python file and have a look at how it is done.
# The robot configuration can be changed in Gepetto-viewer with:
from pinocchio.utils import rand,zero,eye
robot.display(rand(robot.model.nq)*2-1)
# ## Spatial algebra
# The scene, and later the movement and forces in the scene are modeled following Featherstone's Spatial Algebra. Placement, i.e. rotation and translation of frames (and bodies) are stored in objects of the class SE3. Rigid velocities and acceleration are stored in the class Motion, forces in the class Forces and masses/inertias in the class Inertias.
from pinocchio import SE3,Motion,Force,Inertia
M = SE3.Random()
nu = Motion.Random()
phi = Force.Random()
Y = Inertia.Random()
print(M,nu,phi,Y)
# These objects store linear and angular part asside, but we often have to come back to a plain vector/matrix representation. In that case, contrary to Featherstone, we rather store linear part first and angular second.
print(nu,nu.vector.T)
# # Forward kinematics: the first algorithm
# ## Model and data
# Before calling the algorithm, let's introduce a specificity of Pinocchio: the strict separation between constant model element in the Model class, and all the buffers for storing algorithm quantities in the Data class.
rdata = rmodel.createData()
# The RobotWrapper creates a Data object by default:
print(robot.data)
# The idea is that the same model can be used by different part of the algorithm to compute different values from different argument. For example, in a optimal-control implementation of Pinocchio, you likely want to have a single robot model for all your problem, but several data for each node of your optimal control solver. In the tuto, we will for example use the fact that the numerical algorithm has its own Data, while the RobotWrapper use another Data for computing body placements when displaying something.
# ## Calling the algorithm implementation
# The forward kinematics simply compute the placement of every joint frame for a given configuration q.
q = rand(rmodel.nq)
pinocchio.forwardKinematics(rmodel,rdata,q)
for i,M in enumerate(rdata.oMi[1:]): print(i,M)
# When calling forwardKinematics, the model rmodel has not been changed, while the results of the algorithm have been stored in the robot Data. Some algorithm have a main result that is returned by the algorithm (like for example crba that returns the robot mass matrix), but in general the main algorithm results are just several changes in the tables stored in robot Data.
#
# ## Computing the end-effector position
# In a first time, we want to control the end effector position (3D). It is given by:
print("End effector = " , rdata.oMi[-1].translation.T)
# # Optimizing the end effector position
# We can now set up the first optimization problem that compute a robot configuration minimizing the distance between the position of the end-effector and a 3D target.
# For that we will use the fmin_slsqp from SciPy. You need to define the cost function cost(q) that returns a scalar measuring this distance. Beware of a painful implementation detail!
# ## The classes Matrix and Array of NumPy
# NumPy implements matrices and vectors with the class np.array, represented as tensors i.e N-D tables, where vectors would be N=1, matrices N=2. Tensor product, that boils down to Matrix-Vector and Matrix-Matrix products in the corresponding dimension, is obtained with the np.dot operator : np.dot(A,x) to multiply $A*x$. The multiplication operator is not the matrix product but the coefficient-wise product.
#
# On the other hand, NumPy also introduces the class Matrix as a specific implementation of the 2-D array, and overload the __ mult __ operator to match the matrix operator, hence $A*x$ is directly obtain with the \* symbol.
#
# Pinocchio has been implemented with the Matrix class. Any other Python package is rather implemented with the Array class. In particular, the SciPy optimizers are with Array. So, we will painfully have to convert array to matrix before calling Pinocchio algorithms, and back to array when returning the results to the optimizer.
#
m2a = lambda m: np.array(m.flat)
a2m = lambda a: np.matrix(a).T
# ## Cost function
# That said, the cost function simply has to call forwardKinematics, and return the difference between the computed effector position and a reference.
ref = np.matrix([.3,.3,.3]).T
def cost(x):
q = a2m(x)
pinocchio.forwardKinematics(rmodel,rdata,q)
M = rdata.oMi[-1]
p = M.translation
residuals = m2a(p-ref)
return sum( residuals**2)
x0 = np.random.rand(rmodel.nq)
print(cost(x0))
# ## FMin
# The optimizer chosen for the class is SLSQP which is a SQP accepting equality, inequality and bound constraints, using BFGS for quasi-newton acceleration and a least-square QP for computing the Newton step. It is quite a good solver, although not strong enough for implementing real robotics application. It is yet quite comfortable for a class to have access to it through the easy package SciPy.
# The API of slsqp is as follows:
from scipy.optimize import fmin_slsqp
help(fmin_slsqp)
# Here we only use the initial guess and the cost function.
result = fmin_slsqp(x0=np.zeros(rmodel.nq),
func=cost)
qopt = a2m(result)
robot.display(qopt)
# ## Using the viewer to interpret the solver
# Let's use the viewer to see what the solver is doing.
# First, let's add a visual object to mark the target.
gview = robot.viewer.gui
gview.addSphere('world/target',0.1,[1.,0.,0.,1.]) # radius, [R,G,B,A]
gview.applyConfiguration('world/target',[.3,.3,.3,0.,0.,0.,1.]) # x,y,z,quaternion
gview.refresh()
# Gepetto-viewer accepts many types of simple 3d geom (see the /opt/openrobots/share/idl/gepetto/corbaserver/graphical-interface.idl API for a list), and wait for the "refresh" order before placing all of them.
#
# We can also change the robot configuration while the solver works, to render the current guess of the algorithm. For that, slsqp offers the callback interface.
def callbackDisp(x):
import time
q = a2m(x)
robot.display(q)
time.sleep(.1)
result = fmin_slsqp(x0=np.zeros(rmodel.nq),
func=cost,
callback=callbackDisp)
# # The complete program
# In general, it is recommanded to store all interesting information and data related to the optimization program inside a dedicated object, whose paramaters are initialized in the constructer. Then the cost (and later constraint, callback, etc) functions are object methods. A complete implementation of the 3D example is given in arm3d.py
# +
# # %load arm3d.py
from robots import loadTalosArm
from scipy.optimize import fmin_slsqp
import pinocchio
from pinocchio.utils import *
from numpy.linalg import norm,inv,pinv,eig,svd
m2a = lambda m: np.array(m.flat)
a2m = lambda a: np.matrix(a).T
robot = loadTalosArm()
robot.initDisplay(loadModel=True)
class OptimProblem:
def __init__(self,rmodel,rdata,gview=None):
self.rmodel = rmodel
self.rdata = rdata
self.ref = [ .3, 0.3, 0.3 ] # Target position
self.idEff = -1 # ID of the robot object to control
def cost3(self,x):
q = a2m(x)
pinocchio.forwardKinematics(self.rmodel,self.rdata,q)
M = self.rdata.oMi[self.idEff]
self.residuals = m2a(M.translation) - self.ref
return sum( self.residuals**2 )
def initDisplay(self,gview=None):
self.gview = gview
if gview is None: return
self.gobj = "world/target3d"
self.gview.addSphere(self.gobj,.03,[1,0,0,1])
self.gview.applyConfiguration(self.gobj,self.ref+[0,0,0,1])
self.gview.refresh()
def callback(self,x):
import time
q = a2m(x)
robot.display(q)
time.sleep(1e-2)
pbm = OptimProblem(robot.model,robot.model.createData(),robot.viewer.gui)
x0 = m2a(robot.q0)
result = fmin_slsqp(x0=x0,
func=pbm.cost3,
callback=pbm.callback)
qopt = a2m(result)
# -
# # Optimization of the effector placement
# The forward kinematics indeed computes the placement of the last frame, i.e the rotation R and the translation p, denoted M = \[R,p\] $\in SE(3)$.
# We need to define a metric to score the distance between to frames $M_1$ and $M_2$. Several metrics can be chosen, but a nice one is given by the SE3 logarithm function, that converts the gap between two frames into the velocity that should applied (constant) during t=1 to bridge the gap a displace $M_1$ into $M_2$.
M1 = SE3.Random()
M2 = SE3.Random()
nu = pinocchio.log(M1.inverse()*M2)
print(nu,nu.vector.T)
# The norm of the logarithm is a proper cost function: it is 0 if and only if the two frames matches, and positive otherwise ; it is smooth; compare to other fancy metrics, it is easy to differenciate (at least, there are some well founded rules to differentiate the logarithm and related operators).
#
# Modify the program above to search for the robot configuration so that the end effector is placed at a reference position and orientation (solution only if need be).
# +
# # %load arm6d.py
# -
# # Frames
# We already said that the kinematic tree is composed of a hierarchy of frames corresponding to the output of each joint. In practice, we find it useful to attach additional frames to these main frames. We name the main frames defining the kinematic tree by Joint Frames, stored in rdata.oMi. The other frames are described in the model by the rmodel.frames list, each object storing its name, the index of its parent joint frame and the fixed placement with respect to its parent.
#
for i,f in enumerate(rmodel.frames): print(i,f.name,f.parent)
# For convenience, we also describe if this frame was parsed as a body frame, a joint frame (yes, joint frames are copied again in the rmodel.frames list as it makes the access to frame more generic) or as fixed joints (that is a pretty classical trick in URDF.
#
# For example, the joint frame attached to the foot of a biped robot is often located at its ankle, i.e. 5 to 10 cm above the ground. We then also attach a second frame on the foot sole, to make it easier to write the contact constraints. And similarly for a quadruped, the last joint frame is at the knew, and we rather attach another frame at the tip of the leg.
#
# Frames are best indexed by their name:
fid = rmodel.getFrameId('gripper_left_fingertip_2_link')
print(fid)
# The joint placement are stored in rdata.oMi. The frame placements are stored in rdata.oMf. By default, the forwardKinematics does not reevaluate the oMf. Do it with:
pinocchio.updateFramePlacements(rmodel,rdata)
# Note that this method does not need q to evaluate the oMf from the oMi.
#
# Modify the above example to optimize the placement of the robot effector tip rather than its wrist.
# # With joint limits
# The joint limits are also parsed from the URDF model and stored in rmodel.lowerLimit and rmodel.upperLimit.
print(rmodel.lowerPositionLimit.T,rmodel.upperPositionLimit.T)
# fmin_slsqp accepts bound constraints as a list of 2-ples (lower,upper).
bounds=[ (l,u) for l,u in zip(robot.model.lowerPositionLimit.flat,robot.model.upperPositionLimit.flat) ]
# # Constraint or cost
# We yet set up a cost whose optimum is reached at 0. In this case, we could equivalently set it up as a constraint, and possibly optimize a second objective like the posture. Let's do that know, as later we will need to play more with constraint and cost.
#
# ## Constraints in slsqp
# Constraints should be implemented as function that returns an array of NC values, that should be 0 in a successful optimization. It seems that there is a problem in the numerical differencition scheme of slsqp that force the user to return a list of values instead of a array of value.
#
# ## Posture cost under constraint terminal position
# Implement a new cost function that minimize the squared norm between the current configuration and a reference configuration, and turn the previous cost function into a constraint function that returns the list of x,y and z errors to a 3D position target.
#
# The 0 configuration stored in rmodel.neutralConfiguration can be used for a reference configuration.
#
#
# %load arm3dconstraint.py
# # Non Euclidean configuration space
#
# The arm only has revolute joints, which are simple to model. Let's now move to the case where we have joints with 3D rotation, in particular the case of floating robots.
#
# A biped robot can be loaded from robots.py
from robots import loadTalosLegs
robot = loadTalosLegs()
rmodel = robot.model ; rdata = rmodel.createData()
print(robot.model)
# Each leg has 6 revolute joints, 12 joints in total, plus the free flyer joint that denotes the movements between a fixed "universe" frame and the root of the robot locate at the hip. The free flyer corresponds to 12 degrees of freedom but will be represented in Pinocchio with 3 translation and a unit quaternion, i.e. 7 parameters (and 1 constraint, the norm of quaternion should be one). In total, rmodel.nq is 19, while the number of degrees of freedom rmodel.nv is 18:
print(rmodel.nq,rmodel.nv)
# It is now less direct to measure the distance between 2 configurations, randomly pick a configuration and locally change a configuration.
# ## Randomly sampling a configuration
# Pinocchio implements the randomConfiguration algorithm to sample a configuration for a model where q is subject to constraints:
q = pinocchio.randomConfiguration(rmodel)
print(q.T)
# ## Distance and increment of configuration
# A velocity $v_q$ will have rmodel.nv dimension, while q as rmodel.nq>rmodel.nv dimension. It is not possible any more to add q+v.
#
# Pinocchio implements the integrate algorithm to add a displacement $v_q$ in the configuration space.
vq = rand(rmodel.nv)
print(q.shape,vq.shape)
qnext = pinocchio.integrate(rmodel,q,vq)
# We will measure a distance between two configurations $q_1$ and $q_2$ as the velocity to apply during t=1 to go from $q_1$ to $q_2$.
q1 = pinocchio.randomConfiguration(rmodel)
q2 = pinocchio.randomConfiguration(rmodel)
dq = pinocchio.difference(rmodel,q1,q2)
# ## Working with optimization and quaternion: the problem
# If we let the solver optimize over a constrained q without notifying it, the algorithm will quickly comes to a q that does not respect the constraint hence is not an interesting solution.
#
# Try to think of the expected result before running the following algorithm.
# +
# # %load bip6fail.py
from robots import loadTalosLegs
from scipy.optimize import fmin_slsqp
import pinocchio
from pinocchio.utils import *
from numpy.linalg import norm,inv,pinv,eig,svd
m2a = lambda m: np.array(m.flat)
a2m = lambda a: np.matrix(a).T
robot = loadTalosLegs()
robot.initDisplay(loadModel=True)
class OptimProblem:
def __init__(self,rmodel,rdata,gview=None):
self.rmodel = rmodel
self.rdata = rdata
self.refL = pinocchio.SE3(eye(3), np.matrix([ 0., 1.5, 1.]).T )
self.idL = rmodel.getFrameId('left_sole_link') # ID of the robot object to control
self.refR = pinocchio.SE3(eye(3), np.matrix([ 0., -1.5, 0.]).T )
self.idR = rmodel.getFrameId('right_sole_link')# ID of the robot object to control
self.initDisplay(gview)
def cost(self,x):
q = a2m(x)
pinocchio.forwardKinematics(self.rmodel,self.rdata,q)
pinocchio.updateFramePlacements(self.rmodel,self.rdata)
refMl = self.refL.inverse()*self.rdata.oMf[self.idL]
residualL = m2a(pinocchio.log(refMl).vector)
refMr = self.refR.inverse()*self.rdata.oMf[self.idR]
residualR = m2a(pinocchio.log(refMr).vector)
self.residuals = np.concatenate([residualL,residualR])
return sum( self.residuals**2 )
# --- BLABLA -------------------------------------------------------------
def initDisplay(self,gview):
if gview is None: return
self.gview = gview
self.gobjR = "world/targetR"
self.gobjL = "world/targetL"
self.gview.addBox(self.gobjR,.1,.03,.03,[1,0,0,1])
self.gview.addBox(self.gobjL,.1,.03,.03,[0,1,0,1])
self.gview.applyConfiguration(self.gobjR,se3ToXYZQUAT(self.refR))
self.gview.applyConfiguration(self.gobjL,se3ToXYZQUAT(self.refL))
self.gview.refresh()
def callback(self,x):
import time
q = a2m(x)
robot.display(q)
time.sleep(1e-2)
pbm = OptimProblem(robot.model,robot.data,robot.viewer.gui)
x0 = m2a(robot.q0)
result = fmin_slsqp(x0 = x0,
func = pbm.cost,
callback = pbm.callback)
qopt = a2m(result)
# -
# ## Working with optimization and quaternion: solution 1
# We can add a constraint to force the solver to keep the quaternion unitary. Do it! (solution only if need be).
# # %load solution_quaternion_constraint.py
# ## Working with optimization and quaternion: solution 2
# An alternative is to work in another representation of the configuration space that is minimal. To be efficient, it should also be smooth and easy to differentiate.
#
# Here, we will use a representation as $v_q$ the displacement from a reference configuration $q_0$. It is not a very good representation when $v_q$ becomes too large. But it as the advantage that the derivatives are not too complex to compute. And it is a good representation when $v_q$ is small. In more advanced algorithms, we will keep the same representation but change the reference $q_0$ from time to time. By that way, everything that we do here can be kept for a more advance numerical algorithm.
# +
# # %load bip6d.py
from robots import loadTalosLegs
from scipy.optimize import fmin_slsqp
import pinocchio
from pinocchio.utils import *
from numpy.linalg import norm,inv,pinv,eig,svd
m2a = lambda m: np.array(m.flat)
a2m = lambda a: np.matrix(a).T
robot = loadTalosLegs()
robot.initDisplay(loadModel=True)
class OptimProblem:
def __init__(self,rmodel,rdata,gview=None):
self.rmodel = rmodel
self.rdata = rdata
self.refL = pinocchio.SE3(eye(3), np.matrix([ 0., .3, 0.]).T )
self.idL = rmodel.getFrameId('left_sole_link') # ID of the robot object to control
self.refR = pinocchio.SE3(eye(3), np.matrix([ 0., -.3, 0.]).T )
self.idR = rmodel.getFrameId('right_sole_link')# ID of the robot object to control
self.refQ = rmodel.neutralConfiguration
self.initDisplay(gview)
self.neq = 12
self.eq = np.zeros(self.neq)
self.Jeq = np.zeros([self.neq, self.rmodel.nv])
# configurations are represented as velocity integrated from this point.
self.q0 = rmodel.neutralConfiguration
def vq2q(self,vq): return pinocchio.integrate(self.rmodel,self.q0,vq)
def q2vq(self,q): return pinocchio.difference(self.rmodel,self.q0,q)
def cost(self,x):
q = self.vq2q(a2m(x))
self.residuals = m2a(pinocchio.difference(self.rmodel,self.refQ,q)[6:])
return sum( self.residuals**2 )
def constraint_leftfoot(self,x,nc=0):
q = self.vq2q(a2m(x))
pinocchio.forwardKinematics(self.rmodel,self.rdata,q)
pinocchio.updateFramePlacements(self.rmodel,self.rdata)
refMl = self.refL.inverse()*self.rdata.oMf[self.idL]
self.eq[nc:nc+6] = m2a(pinocchio.log(refMl).vector)
return self.eq[nc:nc+6].tolist()
def constraint_rightfoot(self,x,nc=0):
q = self.vq2q(a2m(x))
pinocchio.forwardKinematics(self.rmodel,self.rdata,q)
pinocchio.updateFramePlacements(self.rmodel,self.rdata)
refMr = self.refR.inverse()*self.rdata.oMf[self.idR]
self.eq[nc:nc+6] = m2a(pinocchio.log(refMr).vector)
return self.eq[nc:nc+6].tolist()
def constraint(self,x):
self.constraint_rightfoot(x,0)
self.constraint_leftfoot(x,6)
return self.eq.tolist()
# --- BLABLA -------------------------------------------------------------
def initDisplay(self,gview):
if gview is None: return
self.gview = gview
self.gobjR = "world/targetR"
self.gobjL = "world/targetL"
self.gview.addBox(self.gobjR,.1,.03,.03,[1,0,0,1])
self.gview.addBox(self.gobjL,.1,.03,.03,[0,1,0,1])
self.gview.applyConfiguration(self.gobjR,se3ToXYZQUAT(self.refR))
self.gview.applyConfiguration(self.gobjL,se3ToXYZQUAT(self.refL))
self.gview.refresh()
def callback(self,x):
import time
q = self.vq2q(a2m(x))
robot.display(q)
time.sleep(1e-1)
pbm = OptimProblem(robot.model,robot.data,robot.viewer.gui)
pbm.refQ = robot.q0.copy()
x0 = m2a(pbm.q2vq(robot.q0))
result = fmin_slsqp(x0 = x0,
func = pbm.cost,
f_eqcons = pbm.constraint,
callback = pbm.callback)
qopt = pbm.vq2q(a2m(result))
# -
| 1. Optimize the effector of a manipulator robot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
# Load numpy for math/array operations
# and matplotlib for plotting
import numpy as np
import matplotlib.pyplot as plt
# + slideshow={"slide_type": "subslide"}
# %matplotlib inline
# Set up figure size and DPI for screen demo
plt.rcParams['figure.figsize'] = (4,3)
plt.rcParams['figure.dpi'] = 150
# -
# # Legends
nums = np.arange(0,10,0.1)
plt.plot(nums, np.sin(nums), label='sin')
plt.plot(nums, np.cos(nums), label='cos')
plt.plot(nums, np.tan(nums), label='tan')
plt.ylim(-2,2)
#Location
plt.plot(nums, np.sin(nums), label='sin')
plt.plot(nums, np.cos(nums), label='cos')
plt.plot(nums, np.tan(nums), label='tan')
plt.legend(
plt.ylim(-2,2)
#Number of Columns
plt.plot(nums, np.sin(nums), label='sin')
plt.plot(nums, np.cos(nums), label='cos')
plt.plot(nums, np.tan(nums), label='tan')
plt.plot(nums, np.sinh(nums), label='sinh')
plt.plot(nums, np.cosh(nums), label='cosh')
plt.plot(nums, np.tanh(nums), label='tanh')
plt.legend(
plt.gcf().set_size_inches(6,2)
plt.ylim(-2,2)
#Title
plt.plot(nums, np.sin(nums), label='sin')
plt.plot(nums, np.cos(nums), label='cos')
plt.plot(nums, np.tan(nums), label='tan')
plt.legend(
plt.ylim(-2,2)
#Appearance (frame, fancybox, shadow, font size)
plt.plot(nums, np.sin(nums), label='sin')
plt.plot(nums, np.cos(nums), label='cos')
plt.plot(nums, np.tan(nums), label='tan')
plt.legend(
plt.ylim(-2,2)
# # Colorbars
#Generate a smoothed, gaussian random field
from scipy.ndimage.filters import gaussian_filter
rands2d = gaussian_filter(np.random.normal(size=(512,512)), sigma=10)
plt.imshow(rands2d)
plt.colorbar()
# Orientation
plt.imshow(rands2d)
plt.colorbar(
# Label
plt.imshow(rands2d)
plt.colorbar()
| 6506_02_code_ACC_SB/Video 2.7- Legends and Colorbars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NICO2AI 第5回 勾配法と誤差逆伝搬法 (7/15) 実践演習
#
# 課題
#
# - 3層NNモデルのクロスエントロピー誤差最小化をミニバッチ勾配降下法で実装する
# - MNISTデータセットを用いて学習を行う
#
# `### CODE HERE ###` と記載されている部分にコードを埋めていく
# !wget "https://drive.google.com/uc?export=download&id=1FfK_OGcOU5Jy_jhkXlPYhoq0LmJBIiDB" -O utils.py
# +
# %matplotlib inline
import numpy as np
np.random.seed(111)
import matplotlib as mpl
import matplotlib.pyplot as plt
from IPython.display import display, clear_output
from sklearn.metrics import confusion_matrix
import seaborn as sns
from sklearn.datasets import fetch_mldata
from utils import to_categorical, calculate_accuracy, plot_confusion_matrix, get_image_tile
sns.set_style('ticks')
# -
# ## MNISTデータのロード
# +
mnist = fetch_mldata('MNIST original', data_home='/tmp')
np.random.seed(111)
data_idx = np.arange(70000)
np.random.shuffle(data_idx)
X_train = mnist['data'][data_idx][:50000]
X_valid = mnist['data'][data_idx][50000:60000]
X_test = mnist['data'][data_idx][60000:]
y_train = mnist['target'][data_idx][:50000]
y_valid = mnist['target'][data_idx][50000:60000]
y_test = mnist['target'][data_idx][60000:]
Y_train = to_categorical(y_train)
Y_valid = to_categorical(y_valid)
Y_test = to_categorical(y_test)
# -
im = get_image_tile(X_train, width=10, height=10)
im
# ## 各種関数の実装
# +
def softmax(U, reduce_axis=0):
shp = list(U.shape)
shp[reduce_axis] = 1
return ### CODE HERE ###
def softmax_cross_entropy(D, Y):
epsilon = 1e-8
Y = np.clip(Y, epsilon, 1-epsilon)
return ### CODE HERE ###
def sigmoid(U):
return ### CODE HERE ###
def dsigmoid_du(U):
return ### CODE HERE ###
# +
## エラーチェック
DIM = 20
NB_CLASS = 10
N = 100
U = np.random.normal(size=DIM*N).reshape(DIM, N)
D = np.zeros([NB_CLASS, N])
D[4] = 1.0
Y = np.zeros([NB_CLASS, N])
Y[:] = 1.0 / NB_CLASS
assert softmax(U).shape == (DIM, N), 'softmax(U).shape must be {}. result: {}'.format((DIM, N), softmax(U).shape)
assert softmax_cross_entropy(D, Y).shape == (), \
'softmax_cross_entropy(D, Y).shape must be {}. result: {}'.format((), softmax_cross_entropy(D, Y).shape)
assert 220 <= softmax_cross_entropy(D, Y) <= 240, \
'softmax_cross_entropy(D, Y) must approximately equal to 230, when the values of all elements of Y are equal.'
assert sigmoid(U).shape == (DIM, N), 'sigmoid(U).shape must be {}. result: {}'.format((DIM, N), sigmoid(U).shape)
assert dsigmoid_du(U).shape == (DIM, N), \
'dsigmoid_du(U).shape must be {}. result: {}'.format((DIM, N), dsigmoid_du(U).shape)
# -
# ## パラメーター初期化関数の実装
# +
x_dim = 784
h_dim = 256
nb_classes = 10
def init_params():
W_2 = np.random.normal(loc=0.0, scale=1.0, size=x_dim*h_dim).reshape([h_dim, x_dim])
b_2 = np.zeros(h_dim).reshape(h_dim, 1)
W_3 = np.random.normal(loc=0.0, scale=1.0, size=h_dim*nb_classes).reshape([nb_classes, h_dim])
b_3 = np.zeros(nb_classes).reshape(nb_classes, 1)
return W_2, b_2, W_3, b_3
# -
# ## 推論用関数の実装
def inference(X):
X = X.T
assert X.shape[0] == x_dim
U_2 = ### CODE HERE ###
Z_2 = ### CODE HERE ###
U_3 = ### CODE HERE ###
Z_3 = ### CODE HERE ###
return Z_3.T
# +
## エラーチェック
W_2, b_2, W_3, b_3 = init_params()
assert inference(X_train[:100]).shape == (100, NB_CLASS), \
'inference(X).shape must be {}. result: {}'.format((100, NB_CLASS), inference(X_train[:100]).shape)
assert inference(X_train[:100]).sum(axis=1).shape == np.ones(100).shape, \
'The sum around the class of output of inference(X) must be 1'
# -
# ## 学習
# +
nb_epoch = 10
batch_size = 100
nb_batch = int(len(X_train) / batch_size)
eta = 0.01
data_idx = np.arange(len(X_train))
fig = plt.figure(figsize=(4,4))
ax = fig.add_subplot(111)
train_epochs = []
valid_epochs = []
train_losses = []
valid_losses = []
train_accuracies = []
valid_accuracies = []
plot_freq = 50
W_2, b_2, W_3, b_3 = init_params()
for epoch in range(nb_epoch):
np.random.shuffle(data_idx)
for batch in range(nb_batch):
X_batch = X_train[data_idx[batch*batch_size:(batch+1)*batch_size]]
Y_batch = Y_train[data_idx[batch*batch_size:(batch+1)*batch_size]]
X = X_batch.T
D = Y_batch.T
## 順方向の推論
U_2 = ### CODE HERE ###
Z_2 = ### CODE HERE ###
U_3 = ### CODE HERE ###
Z_3 = ### CODE HERE ###
Y = Z_3
## デルタの計算
Delta_3 = ### CODE HERE ###
Delta_2 = ### CODE HERE ###
## 3層目のパラメーターに関する勾配
dLdW_3 = ### CODE HERE ###
dLdb_3 = ### CODE HERE ###
## 2層目のパラメーターに関する勾配
dLdW_2 = ### CODE HERE ###
dLdb_2 = ### CODE HERE ###
## 3層目のパラメーターの更新
W_3 = ### CODE HERE ###
b_3 = ### CODE HERE ###
## 2層目のパラメーターの更新
W_2 = ### CODE HERE ###
b_2 = ### CODE HERE ###
## リアルタイムの誤差の描画
if batch % plot_freq == 0:
train_epochs.append( epoch+batch/nb_batch )
train_losses.append( softmax_cross_entropy(D, Y) / batch_size )
train_accuracies.append( calculate_accuracy(D.argmax(axis=0), Y.argmax(axis=0)) )
clear_output(wait = True)
ax.plot( train_epochs, train_losses, label='Train' )
ax.plot( valid_epochs, valid_losses, label='Validation' )
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('epoch: {:02d}, batch: {:04d}'.format(epoch, batch))
plt.legend()
display(fig)
ax.cla()
Y = inference(X_valid).T
valid_epochs.append( epoch+1 )
valid_losses.append( softmax_cross_entropy(Y_valid.T, Y) / len(Y_valid) )
valid_accuracies.append( calculate_accuracy(y_valid, Y.argmax(axis=0)) )
fig.clf()
# +
fig = plt.figure(figsize=(4,4))
ax = fig.add_subplot(111)
ax.plot( train_epochs, train_accuracies, label='Train' )
ax.plot( valid_epochs, valid_accuracies, label='Validation' )
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.legend()
plt.ylim(0, 1)
# -
# ## 学習結果の確認
Y_hat = inference(X_valid)
y_hat = Y_hat.argmax(axis=1)
valacc = calculate_accuracy(y_valid, y_hat)
C = confusion_matrix(y_valid, y_hat)
plot_confusion_matrix(C, range(10))
print('Validation accuracy: {:.4f}'.format(valacc))
# ## テストデータでの確認
Y_hat = ### CODE HERE ###
y_hat = Y_hat.argmax(axis=1)
testacc = ### CODE HERE ###
C = confusion_matrix(y_test, y_hat)
plot_confusion_matrix(C, range(10))
print('Test accuracy: {:.4f}'.format(testacc))
| lecture5/nico2ai_lecture5_practice.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:miniconda3-4.3.30]
# language: R
# name: conda-env-miniconda3-4.3.30-r
# ---
# # LOGISTIC REGRESSION
# Logistic regression (aka logit regression or logit model) was developed by statistician <NAME> in 1958 and is a regression model where the response variable Y is categorical. Logistic regression allows us to estimate the probability of a categorical response based on one or more predictor variables (X). It allows one to say that the presence of a predictor increases (or decreases) the probability of a given outcome by a specific percentage. This tutorial covers the case when Y is binary — that is, where it can take only two values, “0” and “1”, which represent outcomes such as pass/fail, win/lose, alive/dead or healthy/sick. Cases where the dependent variable has more than two outcome categories may be analysed with multinomial logistic regression, or, if the multiple categories are ordered, in ordinal logistic regression. However, discriminant analysis has become a popular method for multi-class classification so our next tutorial will focus on that technique for those instances.
install.packages("ROCR")
install.packages('caTools')
library(tidyverse)
library(modelr) # provides easy pipeline modeling functions
library(broom) # helps to tidy up model outputs
library(caTools)
library(ROCR)
# This tutorial primarily leverages the Default data provided by the ISLR package. This is a simulated data set containing information on ten thousand customers such as whether the customer defaulted, is a student, the average balance carried by the customer and the income of the customer. We’ll also use a few packages that provide data manipulation, visualization, pipeline modeling functions, and model output tidying functions.
library(repr)
options(repr.plot.width=5, repr.plot.height=5)
default <- as_tibble(ISLR::Default)
head(default)
set.seed(123)
sample <- sample(c(TRUE, FALSE), nrow(default), replace = T, prob = c(0.6,0.4))
train <- default[sample, ]
test <- default[!sample, ]
head(train)
# We will fit a logistic regression model in order to predict the probability of a customer defaulting based on the average balance carried by the customer. The glm function fits generalized linear models, a class of models that includes logistic regression. The syntax of the glm function is similar to that of lm, except that we must pass the argument family = binomial in order to tell R to run a logistic regression rather than some other type of generalized linear mode
model1 <- glm(default ~ balance, family = "binomial", data = train)
default %>%
mutate(prob = ifelse(default == "Yes", 1, 0)) %>%
ggplot(aes(balance, prob)) +
geom_point(alpha = .15) +
geom_smooth(method = "glm", method.args = list(family = "binomial")) +
ggtitle("Logistic regression model fit") +
xlab("Balance") +
ylab("Probability of Default")
tidy(model1)
exp(coef(model1))
# We can further interpret the balance coefficient as - for every one dollar increase in monthly balance carried, the odds of the customer defaulting increases by a factor of 1.0057.
#
#
confint(model1)
# For example, we can measure the confidence intervals and accuracy of the coefficient estimates by computing their standard errors. For instance,
# ^
# β
# 1
# has a p-value < 2e-16 suggesting a statistically significant relationship between balance carried and the probability of defaulting.
predict(model1, data.frame(balance = c(1000, 2000)), type = "response")
# # Logistic Regression
data = read_csv("data/diabetes_log_r.csv")
head(data)
summary(data)
# - We are predicting whether a person has diabetes or not
# - Outcome 0 : Not Diabetic
# - Outcome 1 : Diabetic
split <- sample.split(data$Outcome, SplitRatio = 0.75)
#GET TRAINING DATA AND TESTING DATA
data_train <- subset(data, split == TRUE)
data_test <- subset(data, split == FALSE)
dim(data_train)
dim(data_test)
## COLUMN NAMES
names(data)
## LOGISTIC REGRESSION MODEL
logit <- glm (Outcome ~Pregnancies +Glucose+BloodPressure+SkinThickness+Insulin+BMI+DiabetesPedigreeFunction+Age,
data = data_train, family = binomial)
summary(logit)
## PREDICTING THE PROBABILITIES
predicted_prob <- predict(logit,type = 'response', newdata=data_test[1:8])
head(predicted_prob)
## CONVERTING PROBABILITY TO PREDICTED OUTCOME
predicted_outcome = ifelse(predicted_prob>0.5,1,0)
head(predicted_outcome)
# ## CONFUSION MATRIX
# 
# - **true positives** (Top-right quadrant): these are cases in which we predicted the customer would default and they did.
# - **true negatives** (Bottom-left quadrant): We predicted no default, and the customer did not default.
# - **false positives** (Bottom-right quadrant): We predicted yes, but they didn’t actually default.
# - **false negatives** (Top-left): We predicted no, but they did default.
## CREATION OF CONFUSION MATRIX
confusion_matrix = table(data_test$Outcome,predicted_outcome)
confusion_matrix
# Accuracy = 
## CALCULATION OF ACCURACY
accuracy = (112+40) / (112+13+27+40)
accuracy
# ### ROC determines the accuracy of a classification model at a user defined threshold value.
# It determines the model's accuracy using Area Under Curve (AUC). Higher the area, better the model. ROC is plotted between True Positive Rate (Y axis) and False Positive Rate (X Axis).
# +
#ROCR Curve
ROCRpred <- prediction(predicted_prob, data_test$Outcome)
ROCRperf <- performance(ROCRpred, 'tpr','fpr')
plot(ROCRperf, colorize = TRUE )
# -
| 06-Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from nbdev import *
# +
# default_exp text_transcribe
# -
# ttsdataset# VoskTranscribe
# > Class for accented word/phase transcription according to the [`VOSK`](https://alphacephei.com/vosk/models.html) library rules.
# > Supports `list`, `str` thanks to type dispatch implemented in [`fastcore`](https://fastcore.fast.ai/) library.
#export
from fastcore.all import *
#export
class VoskTranscribe():
"""Transcribe an accented word using rules of `VOSK` library.
For example:
абстракциони+стов
абстра+кцию
абстра+кция
абстракционистов a0 b s t r a0 k c i0 o0 nj i1 s t o0 v
абстракцию a0 b s t r a1 k c i0 j u0
абстракция a0 b s t r a1 k c i0 j a0
The code is adapted from `vosk-model-ru-0.10/extra/scripts/dictionary.py` https://alphacephei.com/vosk/
"""
def __init__(self, acc_before=False):
"""Create a `VoskTranscribe`r.
Args:
acc_before (bool): Accent marked with `+` before (True) or after (False) a vowel. Default=False.
Example: "сл+ива" -- use `acc_before = True`, like in `VOSK`.
"сли+ва" -- use `acc_before = False`, like in `russian_g2p`.
"""
self.acc_before = acc_before
self.softletters=set("яёюиье")
self.startsyl=set("#ъьаяоёуюэеиы-")
self.others = set("#+-ьъ")
self.softhard_cons = {
"б" : "b",
"в" : "v",
"г" : "g",
"Г" : "g",
"д" : "d",
"з" : "z",
"к" : "k",
"л" : "l",
"м" : "m",
"н" : "n",
"п" : "p",
"р" : "r",
"с" : "s",
"т" : "t",
"ф" : "f",
"х" : "h"
}
self.other_cons = {
"ж" : "zh",
"ц" : "c",
"ч" : "ch",
"ш" : "sh",
"щ" : "sch",
"й" : "j"
}
self.vowels = {
"а" : "a",
"я" : "a",
"у" : "u",
"ю" : "u",
"о" : "o",
"ё" : "o",
"э" : "e",
"е" : "e",
"и" : "i",
"ы" : "y",
}
@typedispatch
def __call__(self, word: str) -> str:
"""To call class instance as a function."""
return self.convert(word)
@typedispatch
def __call__(self, phrase: list) -> list:
"""To call class instance as a function."""
if isinstance(phrase[0],list): phrase = phrase[0]
return [self.convert(word) for word in phrase]
def __pallatize(self, phones: list) -> list:
"""Transcribe consonant phones.
Args:
phones (list): tuples of phones marked: 0 -- not stressed, 1 -- stressed.
Example: [('#', 0), ('с', 0), ('л', 0), ('и', 1), ('в', 0), ('а', 0), ('#', 0)]
Returns:
list of tuples: consonants transcribed.
Example: [('#', 0), ('s', 0), ('lj', 0), ('и', 1), ('v', 0), ('а', 0), ('#', 0)]
"""
for i, (ph, _) in enumerate(phones[:-1]):
if ph in self.softhard_cons:
if phones[i+1][0] in self.softletters:
phones[i] = (self.softhard_cons[ph] + "j", 0)
else:
phones[i] = (self.softhard_cons[ph], 0)
if ph in self.other_cons:
phones[i] = (self.other_cons[ph], 0)
return phones
def __convert_vowels(self, phones: list) -> list:
"""Transcribe vowel phones.
Args:
phones (list): tuples of phones marked: 0 -- not stressed, 1 -- stressed.
Example: [('#', 0), ('s', 0), ('lj', 0), ('и', 1), ('v', 0), ('а', 0), ('#', 0)]
Returns:
list: consonants transcribed. Ex: ['#', 's', 'lj', 'i1', 'v', 'a0', '#']
"""
new_phones = []
prev = ""
for (ph, stress) in phones:
if prev in self.startsyl:
if ph in set("яюеё"):
new_phones.append("j")
if ph in self.vowels:
new_phones.append(self.vowels[ph] + str(stress))
else:
new_phones.append(ph)
prev = ph
return new_phones
def convert(self, word: str) -> str:
""""""
if word == '<sil>': return word
phones = ("#" + word + "#")
# Assign stress marks
stress_phones = []
acc_before = False
offset = -1 if self.acc_before else 1
for i,ph in enumerate(phones[:-1]):
if ph == '+': continue
if phones[i+offset] == '+':
stress_phones.append((ph,1))
else:
stress_phones.append((ph,0))
else:
stress_phones.append((phones[-1],0))
phones = self.__convert_vowels(self.__pallatize(stress_phones))
phones = [x for x in phones if x not in self.others] # Filter
return " ".join(phones)
# ## Testing
#hide
from fastcore.test import *
from nbdev.showdoc import *
# #### VOSK-like accents
vt = VoskTranscribe(acc_before=True)
# ##### Calling with a list of words
vt([['куп+юра', 'мо+я']])
# ##### Calling with a string
vt('куп+юра')
test_eq(vt('куп+юра'), 'k u0 pj u1 r a0')
test_eq(vt('к+онь'), 'k o1 nj')
# #### Russian_G2P-like accents
# +
vt = VoskTranscribe(acc_before=False)
test_eq(vt('купю+ра'), 'k u0 pj u1 r a0')
test_eq(vt('ко+нь'), 'k o1 nj')
# -
# ##### Ignore special tag
test_eq(vt('<sil>'), '<sil>')
# ## TODO
#
# * raise a `Warning` in case no stress.
test_eq(vt('конь'), 'k o0 nj')
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/03_transcribe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from malaya_speech.utils import subword
import pandas as pd
with open('train/TRANS.txt') as fopen:
data = fopen.read().split('\n')
df = pd.read_csv('train/TRANS.txt', sep = '\t')
df.head()
import pinyin
from tqdm import tqdm
texts = df['Transcription'].tolist()
texts = [pinyin.get(t, format="strip", delimiter=" ") for t in tqdm(texts)]
tokenizer = subword.generate_tokenizer(texts, target_vocab_size = 512, max_subword_length = 3)
subword.save(tokenizer, 'mandarin-512.subword')
| pretrained-model/prepare-stt/vocab/build-mandarin-vocab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/newcooldiscoveries/AMPL/blob/master/MedNIST_tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JeC6o5OMAFN7"
#
# # Image Classification Tutorial with the MedNIST Dataset
#
# Introduction
# In this tutorial, we introduce an end-to-end training and evaluation example based on the MedNIST dataset.
# We'll go through the following steps:
#
# - Create a MONAI Dataset for training and testing
# - Use MONAI transforms to pre-process data
# - Use the DenseNet from MONAI for the classification task
# - Train the model with a PyTorch program
# - Evaluate on test dataset
#
# ### Get the dataset
# The MedNIST dataset was gathered from several sets from [TCIA](https://wiki.cancerimagingarchive.net/display/Public/Data+Usage+Policies+and+Restrictions), [the RSNA Bone Age Challenge](http://rsnachallenges.cloudapp.net/competitions/4), and [the NIH Chest X-ray dataset](https://cloud.google.com/healthcare/docs/resources/public-datasets/nih-chest).
#
# The dataset is kindly made available by [Dr. <NAME>., Ph.D.](https://www.mayo.edu/research/labs/radiology-informatics/overview) (Department of Radiology, Mayo Clinic) under the Creative Commons [CC BY-SA 4.0 license](https://creativecommons.org/licenses/by-sa/4.0/). If you use the MedNIST dataset, please acknowledge the source, e.g.
#
# https://github.com/Project-MONAI/MONAI/blob/master/examples/notebooks/mednist_tutorial.ipynb.
#
# The following commands download and unzip the dataset (~60MB).
#
# + id="BZTNbFpgA6rX"
# !wget -q https://www.dropbox.com/s/5wwskxctvcxiuea/MedNIST.tar.gz
# + id="jyDZo_qaBEyn"
# unzip the '.tar.gz' file to the current directory
import tarfile
datafile = tarfile.open("MedNIST.tar.gz")
datafile.extractall()
datafile.close()
# + [markdown] id="c3rUPlOhBsnR"
# Install MONAI
# + id="oRsBnGJeBJgW"
# !pip install -q "monai-weekly[gdown, nibabel, tqdm, itk]"
# + id="Ty_YA-cLBpyr" colab={"base_uri": "https://localhost:8080/"} outputId="26d3baf4-0c29-4174-fc3f-685ddb1c15b2"
import os
import shutil
import tempfile
import matplotlib.pyplot as plt
import PIL
import torch
import numpy as np
from sklearn.metrics import classification_report
from monai.apps import download_and_extract
from monai.config import print_config
from monai.metrics import compute_roc_auc
from monai.networks.nets import DenseNet121
from monai.transforms import (
Activations,
AddChannel,
AsDiscrete,
Compose,
LoadImage,
RandFlip,
RandRotate,
RandZoom,
ScaleIntensity,
ToTensor,
)
from monai.utils import set_determinism
print_config()
# + [markdown] id="qHhD_snaB70p"
#
# ## Read image filenames from the dataset folders
# First of all, check the dataset files and show some statistics.
# There are 6 folders in the dataset: Hand, AbdomenCT, CXR, ChestCT, BreastMRI, HeadCT,
# which should be used as the labels to train our classification model.
# + id="ZaHFhidyCBJa" colab={"base_uri": "https://localhost:8080/"} outputId="4ce6d95c-4c01-440e-f5f6-20bf8985c7e4"
data_dir = './MedNIST/'
class_names = sorted([x for x in os.listdir(data_dir) if os.path.isdir(os.path.join(data_dir, x))])
num_class = len(class_names)
image_files = [[os.path.join(data_dir, class_name, x)
for x in os.listdir(os.path.join(data_dir, class_name))]
for class_name in class_names]
image_file_list = []
image_label_list = []
for i, class_name in enumerate(class_names):
image_file_list.extend(image_files[i])
image_label_list.extend([i] * len(image_files[i]))
num_total = len(image_label_list)
image_width, image_height = Image.open(image_file_list[0]).size
print('Total image count:', num_total)
print("Image dimensions:", image_width, "x", image_height)
print("Label names:", class_names)
print("Label counts:", [len(image_files[i]) for i in range(num_class)])
# + [markdown] id="9KuPf7t-CFEV"
# ## Visualise some randomly picked examples from the dataset
# + id="p7kXrcmPCQPU" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="18310a0e-a2b3-4dec-f1ce-bcbe9d26b82d"
plt.subplots(3, 3, figsize=(8, 8))
for i,k in enumerate(np.random.randint(num_total, size=9)):
im = Image.open(image_file_list[k])
arr = np.array(im)
plt.subplot(3, 3, i + 1)
plt.xlabel(class_names[image_label_list[k]])
plt.imshow(arr, cmap='gray', vmin=0, vmax=255)
plt.tight_layout()
plt.show()
# + [markdown] id="tkguefb6CTw5"
# ## Prepare training, validation and test data lists
# Randomly select 10% of the dataset as validation and 10% as test.
# + id="WuryrHlpCYfK" colab={"base_uri": "https://localhost:8080/"} outputId="1c9a6718-3318-43cb-9147-ac53a2e554d6"
valid_frac, test_frac = 0.1, 0.1
trainX, trainY = [], []
valX, valY = [], []
testX, testY = [], []
for i in range(num_total):
rann = np.random.random()
if rann < valid_frac:
valX.append(image_file_list[i])
valY.append(image_label_list[i])
elif rann < test_frac + valid_frac:
testX.append(image_file_list[i])
testY.append(image_label_list[i])
else:
trainX.append(image_file_list[i])
trainY.append(image_label_list[i])
print("Training count =",len(trainX),"Validation count =", len(valX), "Test count =",len(testX))
# + [markdown] id="IJRqm8MNCbi4"
# ## Define MONAI transforms, Dataset and Dataloader to pre-process data
# + id="PMsUgaYNCfrw"
train_transforms = Compose([
LoadImage(image_only=True),
AddChannel(),
ScaleIntensity(),
RandRotate(range_x=15, prob=0.5, keep_size=True),
RandFlip(spatial_axis=0, prob=0.5),
RandZoom(min_zoom=0.9, max_zoom=1.1, prob=0.5, keep_size=True),
ToTensor()
])
val_transforms = Compose([
LoadImage(image_only=True),
AddChannel(),
ScaleIntensity(),
ToTensor()
])
act = Activations(softmax=True)
to_onehot = AsDiscrete(to_onehot=True, n_classes=num_class)
# + id="JJgCYleyCpTT"
class MedNISTDataset(Dataset):
def __init__(self, image_files, labels, transforms):
self.image_files = image_files
self.labels = labels
self.transforms = transforms
def __len__(self):
return len(self.image_files)
def __getitem__(self, index):
return self.transforms(self.image_files[index]), self.labels[index]
train_ds = MedNISTDataset(trainX, trainY, train_transforms)
train_loader = DataLoader(train_ds, batch_size=300, shuffle=True, num_workers=2)
val_ds = MedNISTDataset(valX, valY, val_transforms)
val_loader = DataLoader(val_ds, batch_size=300, num_workers=2)
test_ds = MedNISTDataset(testX, testY, val_transforms)
test_loader = DataLoader(test_ds, batch_size=300, num_workers=2)
# + [markdown] id="hmwtEGBbCtuL"
# ## Define network and optimizer
# 1. Set learning rate for how much the model is updated per batch.
# 2. Set total epoch number, as we have shuffle and random transforms, so the training data of every epoch is different.
# And as this is just a get start tutorial, let's just train 4 epochs.
# If train 10 epochs, the model can achieve 100% accuracy on test dataset.
# 3. Use DenseNet from MONAI and move to GPU devide, this DenseNet can support both 2D and 3D classification tasks.
# 4. Use Adam optimizer.
# + id="3efM0bwsC1wS"
device = torch.device("cuda:0")
model = DenseNet121(
spatial_dims=2,
in_channels=1,
out_channels=num_class
).to(device)
loss_function = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), 1e-5)
epoch_num = 4
val_interval = 1
# + [markdown] id="6XpGqz_oDkXP"
# ## Model training
# Execute a typical PyTorch training that run epoch loop and step loop, and do validation after every epoch.
# Will save the model weights to file if got best validation accuracy.
# + id="MpKhLo3YDpXw" colab={"base_uri": "https://localhost:8080/"} outputId="1812976a-cc28-46d8-f8a8-f1b8b428f968"
best_metric = -1
best_metric_epoch = -1
epoch_loss_values = list()
metric_values = list()
for epoch in range(epoch_num):
print('-' * 10)
print(f"epoch {epoch + 1}/{epoch_num}")
model.train()
epoch_loss = 0
step = 0
for batch_data in train_loader:
step += 1
inputs, labels = batch_data[0].to(device), batch_data[1].to(device)
optimizer.zero_grad()
outputs = model(inputs)
loss = loss_function(outputs, labels)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
print(f"{step}/{len(train_ds) // train_loader.batch_size}, train_loss: {loss.item():.4f}")
epoch_len = len(train_ds) // train_loader.batch_size
epoch_loss /= step
epoch_loss_values.append(epoch_loss)
print(f"epoch {epoch + 1} average loss: {epoch_loss:.4f}")
if (epoch + 1) % val_interval == 0:
model.eval()
with torch.no_grad():
y_pred = torch.tensor([], dtype=torch.float32, device=device)
y = torch.tensor([], dtype=torch.long, device=device)
for val_data in val_loader:
val_images, val_labels = val_data[0].to(device), val_data[1].to(device)
y_pred = torch.cat([y_pred, model(val_images)], dim=0)
y = torch.cat([y, val_labels], dim=0)
y_onehot = to_onehot(y)
y_pred_act = act(y_pred)
auc_metric = compute_roc_auc(y_pred_act, y_onehot)
del y_pred_act, y_onehot
metric_values.append(auc_metric)
acc_value = torch.eq(y_pred.argmax(dim=1), y)
acc_metric = acc_value.sum().item() / len(acc_value)
if auc_metric > best_metric:
best_metric = auc_metric
best_metric_epoch = epoch + 1
torch.save(model.state_dict(), 'best_metric_model.pth')
print('saved new best metric model')
print(f"current epoch: {epoch + 1} current AUC: {auc_metric:.4f}"
f" current accuracy: {acc_metric:.4f} best AUC: {best_metric:.4f}"
f" at epoch: {best_metric_epoch}")
print(f"train completed, best_metric: {best_metric:.4f} at epoch: {best_metric_epoch}")
# + [markdown] id="kBDNC9--DtI8"
# ## Plot the loss and metric
# + id="7P1BlRfsDuz4" colab={"base_uri": "https://localhost:8080/", "height": 404} outputId="aa1e02d8-3ae2-41a1-b6b6-0207158f6265"
plt.figure('train', (12, 6))
plt.subplot(1, 2, 1)
plt.title("Epoch Average Loss")
x = [i + 1 for i in range(len(epoch_loss_values))]
y = epoch_loss_values
plt.xlabel('epoch')
plt.plot(x, y)
plt.subplot(1, 2, 2)
plt.title("Validation: Area under the ROC curve")
x = [val_interval * (i + 1) for i in range(len(metric_values))]
y = metric_values
plt.xlabel('epoch')
plt.plot(x, y)
plt.show()
# + [markdown] id="Hrx_mtTODyOe"
# ## Evaluate the model on test dataset
# After training and validation, we already got the best model on validation test.
# We need to evaluate the model on test dataset to check whether it's robust and not over-fitting.
# We'll use these predictions to generate a classification report.
# + id="uHAA3LUxD2b6"
model.load_state_dict(torch.load('best_metric_model.pth'))
model.eval()
y_true = list()
y_pred = list()
with torch.no_grad():
for test_data in test_loader:
test_images, test_labels = test_data[0].to(device), test_data[1].to(device)
pred = model(test_images).argmax(dim=1)
for i in range(len(pred)):
y_true.append(test_labels[i].item())
y_pred.append(pred[i].item())
# + id="zOy8uzlwD8se" colab={"base_uri": "https://localhost:8080/"} outputId="e6a9bb4a-be39-4724-f5f5-6bda07190713"
from sklearn.metrics import classification_report
print(classification_report(y_true, y_pred, target_names=class_names, digits=4))
| MedNIST_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# -*- coding: utf-8 -*-
import os
import google.oauth2.credentials
import google_auth_oauthlib.flow
from googleapiclient.discovery import build
from googleapiclient.errors import HttpError
from google_auth_oauthlib.flow import InstalledAppFlow
# The CLIENT_SECRETS_FILE variable specifies the name of a file that contains
# the OAuth 2.0 information for this application, including its client_id and
# client_secret.
CLIENT_SECRETS_FILE = "client_secret.json"
# This OAuth 2.0 access scope allows for full read/write access to the
# authenticated user's account and requires requests to use an SSL connection.
SCOPES = ['https://www.googleapis.com/auth/youtube.force-ssl']
API_SERVICE_NAME = 'youtube'
API_VERSION = 'v3'
def get_authenticated_service():
flow = InstalledAppFlow.from_client_secrets_file(CLIENT_SECRETS_FILE, SCOPES)
credentials = flow.run_console()
return build(API_SERVICE_NAME, API_VERSION, credentials = credentials)
def print_response(response):
print(response)
# Build a resource based on a list of properties given as key-value pairs.
# Leave properties with empty values out of the inserted resource.
def build_resource(properties):
resource = {}
for p in properties:
# Given a key like "snippet.title", split into "snippet" and "title", where
# "snippet" will be an object and "title" will be a property in that object.
prop_array = p.split('.')
ref = resource
for pa in range(0, len(prop_array)):
is_array = False
key = prop_array[pa]
# For properties that have array values, convert a name like
# "snippet.tags[]" to snippet.tags, and set a flag to handle
# the value as an array.
if key[-2:] == '[]':
key = key[0:len(key)-2:]
is_array = True
if pa == (len(prop_array) - 1):
# Leave properties without values out of inserted resource.
if properties[p]:
if is_array:
ref[key] = properties[p].split(',')
else:
ref[key] = properties[p]
elif key not in ref:
# For example, the property is "snippet.title", but the resource does
# not yet have a "snippet" object. Create the snippet object here.
# Setting "ref = ref[key]" means that in the next time through the
# "for pa in range ..." loop, we will be setting a property in the
# resource's "snippet" object.
ref[key] = {}
ref = ref[key]
else:
# For example, the property is "snippet.description", and the resource
# already has a "snippet" object.
ref = ref[key]
return resource
# Remove keyword arguments that are not set
def remove_empty_kwargs(**kwargs):
good_kwargs = {}
if kwargs is not None:
for key, value in kwargs.iteritems():
if value:
good_kwargs[key] = value
return good_kwargs
def videos_list_most_popular(client, **kwargs):
# See full sample for function
kwargs = remove_empty_kwargs(**kwargs)
response = client.videos().list(
**kwargs
).execute()
return print_response(response)
if __name__ == '__main__':
# When running locally, disable OAuthlib's HTTPs verification. When
# running in production *do not* leave this option enabled.
os.environ['OAUTHLIB_INSECURE_TRANSPORT'] = '1'
client = get_authenticated_service()
videos_list_most_popular(client,
part='snippet,contentDetails,statistics',
chart='mostPopular',
regionCode='US',
videoCategoryId='20')
# -
| WU_HAN_INFO6210_ProjectCodeReview/DB_Project/Python_files/youtube_API_movies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import spacy
from sklearn.datasets import fetch_20newsgroups
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.base import TransformerMixin
from sklearn.feature_extraction.stop_words import ENGLISH_STOP_WORDS as stopwords
from sklearn.cluster import DBSCAN
from sklearn.feature_extraction.text import TfidfVectorizer
import hdbscan
from sklearn.decomposition import TruncatedSVD
from sklearn.preprocessing import Normalizer
import string
import time
import re
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams["figure.figsize"] = [16,9]
# -
def load20NewsGroups():
# #############################################################################
# Load some categories from the training set
categories = [
'alt.atheism',
'talk.religion.misc',
'comp.graphics',
'sci.space',
]
dataset = fetch_20newsgroups(subset='all', categories=categories,
shuffle=True, random_state=42)
print("%d documents" % len(dataset.data))
print("%d categories" % len(dataset.target_names))
return dataset.data
# +
###########create data clearner
#Custom transformer using spaCy
class CleanTextTransformer(TransformerMixin):
def transform(self, X, **transform_params):
return [cleanText(text) for text in X]
def fit(self, X, y=None, **fit_params):
return self
def get_params(self, deep=True):
return {}
# A custom function to clean the text before sending it into the vectorizer
def cleanText(text):
# get rid of newlines
text = text.strip().replace("\n", " ").replace("\r", " ")
# replace twitter @mentions
mentionFinder = re.compile(r"@[a-z0-9_]{1,15}", re.IGNORECASE)
text = mentionFinder.sub("@MENTION", text)
# replace HTML symbols
text = text.replace("&", "and").replace(">", ">").replace("<", "<")
# lowercase
text = text.lower()
return text
# +
############create tokenizer
#Create spacy tokenizer that parses a sentence and generates tokens
#these can also be replaced by word vectors
# List of symbols we don't care about
punctuations = " ".join(string.punctuation).split(" ") + ["-----", "---", "...", "“", "”", "'ve"]
parser = spacy.load('en')
def tokenizeText(sentence):
tokens = parser(sentence)
tokens = [tok.lemma_.lower().strip() if tok.lemma_ != "-PRON-" else tok.lower_ for tok in tokens]
tokens = [tok for tok in tokens if (tok not in stopwords and tok not in punctuations)]
# remove large strings of whitespace
while "" in tokens:
tokens.remove("")
while " " in tokens:
tokens.remove(" ")
while "\n" in tokens:
tokens.remove("\n")
while "\n\n" in tokens:
tokens.remove("\n\n")
return tokens
# -
##########Create preprocess pipline and run
def preProcessData(X_train):
#create vectorizer object to generate feature vectors, we will use custom spacy’s tokenizer
vectorizer = TfidfVectorizer(tokenizer = tokenizeText)
svd = TruncatedSVD(2)
#normalizer = Normalizer(copy=False)
start_time = time.time()
pipe_preprocess = Pipeline([("cleaner", CleanTextTransformer()),
("vectorizer", vectorizer),
("svd", svd)])
X_train_preprocess = pipe_preprocess.fit_transform(X_train)
end_time = time.time()
print("Preprocess done in {} Seconds".format(end_time - start_time))
return X_train_preprocess
######### visualize two dimensional cluster
def visualizeCluster(x1, x2, colors):
plt.scatter(x1, x2, c=colors, s=50, linewidths=0.5, alpha=0.7)
plt.show()
###########Training model 1:DBSCAN
X_train = load20NewsGroups()
X_train_preprocess = preProcessData(X_train)
start_time = time.time()
model = DBSCAN(eps=0.1, min_samples=20, algorithm="brute", metric="cosine")
model.fit(X_train_preprocess)
end_time = time.time()
print("Trained DBSCAN model in {} Seconds".format(end_time - start_time))
print("#of labels {},\nlabels:{}".format(len(set(model.labels_)), model.labels_))
visualizeCluster(X_train_preprocess[:, 0], X_train_preprocess[:, 1], colors = model.labels_)
model = DBSCAN(eps=0.001, min_samples=10, algorithm="brute", metric="cosine")
model.fit(X_train_preprocess)
end_time = time.time()
print("Trained DBSCAN model in {} Seconds".format(end_time - start_time))
print("#of labels {},\nlabels:{}".format(len(set(model.labels_)), model.labels_))
visualizeCluster(X_train_preprocess[:, 0], X_train_preprocess[:, 1], colors = model.labels_)
# ### 直接用DBSCAN出来的效果并不好,维度灾难
# 试一下hdbscan的效果
# +
###########Training mode2 1:hdbscan
start_time = time.time()
clusterer = hdbscan.HDBSCAN()
clusterer.fit(X_train_preprocess)
end_time = time.time()
print("Trained DBSCAN model in {} Seconds".format(end_time - start_time))
print("#of labels {},\nlabels:{}".format(len(set(clusterer.labels_)), clusterer.labels_))
visualizeCluster(X_train_preprocess[:, 0], X_train_preprocess[:, 1], colors = clusterer.labels_)
clusterer.condensed_tree_.plot()
# -
print(X_train_preprocess[:10])
| algorithms/nlp/spacy_textclustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="00001-16c1dadd-ebdb-4f10-bf4e-569a151efb65" deepnote_app_coordinates={"h": 5, "w": 12, "x": 0, "y": 0} deepnote_cell_type="markdown" tags=[]
# # SQLite3
# ## database interaction from Python
# + [markdown] cell_id="00001-df373803-219c-4ddd-a72d-d54d888ac757" deepnote_cell_type="markdown" tags=[]
#
# To interact with a databse sqlite3 and pandas are excelent starting points.
# + cell_id="00001-57225758-b89d-4cfe-82ca-ff791831a016" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=25 execution_start=1638459121601 source_hash="ff22691a" tags=[]
# import
import sqlite3
import pandas as pd
# + [markdown] cell_id="00002-66d49159-2e01-47ce-8906-1fd379159735" deepnote_cell_type="markdown" tags=[]
# As an example database we will work with a dataset downloaded from [BOLD database](https://boldsystems.org/) with information related to the canidae family.
# The database here used was converted to db format from a csv table used in our paper [4SpecID: Reference DNA Libraries Auditing and Annotation System for Forensic Applications](https://doi.org/10.3390/genes12010061) just for demonstration pruposes.
#
# The datase includes two tables:
# * species: with columns ['recordID', 'phylum_name', 'class_name', 'order_name', 'family_name',
# 'genus_name', 'species_name', 'subspecies_name']
# * bins: with columns ['recordID', 'bin_uri', 'nucleotides']
#
# Both tables connect through a common columns, 'recordid'
#
# + [markdown] cell_id="00004-6cb5979b-b6d0-41ad-be79-94398d71824c" deepnote_cell_type="markdown" tags=[]
# The first thing we need is to create a pointer (squlite3 connection object) to our database and a function to get access to its entries, by taking advantadge of pandas.
# + cell_id="00005-603e784e-b1ce-4740-896d-852c58e76694" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1638459121627 source_hash="52341309" tags=[]
# connect database
db = sqlite3.connect('Canidae_COI.db')
def get_data(query):
'''
get data from the database
'''
return pd.read_sql(query, db)
# + [markdown] cell_id="00006-815c119f-6995-4071-9933-7e548ebe3ef4" deepnote_cell_type="markdown" tags=[]
# ## Tables in database
#
# One database can contain one or more tables. The example here contain two (described above), but if we want to access data from a db without previous knowledge we can access the number of tables and their names by using the command .execute as in the example below.
# + cell_id="00006-919f3d9f-d2c4-476b-a354-8911e0544f8f" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1638459121627 source_hash="30606832" tags=[]
cursor = db.cursor()
cursor.execute("SELECT name FROM sqlite_master WHERE type='table';")
tableNames = cursor.fetchall()
tableNames = [ tableNames[i][0] for i in range(len(tableNames)) ]
print(tableNames)
# + [markdown] cell_id="00004-7238aaf1-456c-4858-8c57-2ba6741cb124" deepnote_cell_type="markdown" tags=[]
# To access specific tables and/or data inside tables several functions exist.
#
# Let's start by investigating some SQL commands such as SELECT; WHERE; LIMIT,...
#
# + [markdown] cell_id="00007-f9631f6f-404a-42dd-9b28-42f607ecb6e5" deepnote_cell_type="markdown" tags=[]
# ## Access database data
#
# + [markdown] cell_id="00010-8f722b69-f8a6-4154-8330-590cdf3e4b86" deepnote_cell_type="markdown" tags=[]
# ### SELECT
# Command "SELECT" plus wildcard "*" to get the entire table
#
# + cell_id="00005-9f5d0faa-b970-4e92-8fd2-4d2636eb111d" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=240 execution_start=1638459121628 source_hash="eda8910a" tags=[]
query = 'SELECT * FROM species;'
speciesDF = get_data(query)
speciesDF
# + [markdown] cell_id="00009-e02c45d4-5896-4802-9b17-7d8e069453f6" deepnote_cell_type="markdown" tags=[]
# ### LIMIT
# A smaller version of the database could have been retrived by using LIMIT, to limit the number of output rows; and SELECT, to retrieve just a few variables
# + cell_id="00010-f5427529-340a-43f4-9c17-cb3d2ef6b2b2" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=26 execution_start=1638459121870 source_hash="6c52ad1d" tags=[]
speciesSmallerDF = get_data('SELECT recordid, species_name, subspecies_name FROM species LIMIT 100;')
speciesSmallerDF
# + [markdown] cell_id="00014-f9393ad3-d882-4d49-8349-557b4b30e9fd" deepnote_cell_type="markdown" tags=[]
# ### ORDER BY
# + [markdown] cell_id="00013-e1061391-9bc0-4b77-a9c1-287e7bf39085" deepnote_cell_type="markdown" tags=[]
# Data can be sorted according to some column of interest using "ORDER BY" followed by ASC or DESC for ascending or descending order.
# + cell_id="00012-99042238-613e-46d2-9d2e-7e058ac48322" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1638459121920 source_hash="50f73574" tags=[]
speciesOrderSmallerDF = get_data('SELECT recordid, species_name, subspecies_name FROM species ORDER BY species_name DESC LIMIT 100;')
speciesOrderSmallerDF
# + [markdown] cell_id="00017-7956ff26-d4a0-43db-a1db-b34e49e5308f" deepnote_cell_type="markdown" tags=[]
# ### WHERE
# + [markdown] cell_id="00013-113b284a-fd53-4951-b111-a21e99660a10" deepnote_cell_type="markdown" tags=[]
# Other keywords can be used to trim the recovered database. For example "WHERE".
# Let's get just data with subspecies assigned.
# + cell_id="00014-70dc649e-e0b3-4aac-93cf-3be2b7a76a39" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=44 execution_start=1638459121927 source_hash="66aceb03" tags=[]
query = '''
SELECT recordid, species_name, subspecies_name
FROM species
WHERE ( subspecies_name != "None" )
ORDER BY subspecies_name
'''
speciesWithSubSpeciesDF = get_data(query)
speciesWithSubSpeciesDF
# + [markdown] cell_id="00015-249c5cde-8c14-4441-a960-0fb3ed5b2684" deepnote_cell_type="markdown" tags=[]
# ## Logical operators
# AND and OR operators can be used to filter even more
# + cell_id="00016-c167d82e-984e-4d9e-b5b6-3e5a154297c0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=26 execution_start=1638459121971 source_hash="7ba92a9d" tags=[]
query = '''
SELECT recordid, species_name, subspecies_name
FROM species
WHERE ( subspecies_name != "None" AND species_name == "Canis lupus")
ORDER BY subspecies_name
'''
speciesWithSubSpeciesDF = get_data(query)
speciesWithSubSpeciesDF
# + [markdown] cell_id="00020-de05208b-4429-44f8-a53d-cfab7df90637" deepnote_cell_type="markdown" tags=[]
# ## Statistics
# + [markdown] cell_id="00017-2de1105a-9241-4853-a95c-7addd13235ca" deepnote_cell_type="markdown" tags=[]
# ### COUNT
# Finally, to determine the number of entries that fulfilled a given set of criteria, we can get the table and determine its size, or, **more quickly**, use the COUNT
# + [markdown] cell_id="00017-91a775bb-8b2f-4df0-8f37-38043248362b" deepnote_cell_type="markdown" tags=[]
#
# + cell_id="00019-bae331ed-a3ad-4983-8f01-d680fda342b2" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1638459121997 source_hash="56d57b9f" tags=[]
howManyCanisLupus = get_data('SELECT COUNT(*) FROM species WHERE species_name == "Canis lupus";')
howManyCanisLupus
# + [markdown] cell_id="00021-633211eb-1a71-4758-a3d4-4a871688b835" deepnote_cell_type="markdown" tags=[]
# ### GROUP BY
# Or we can use COUNT combined with GROUP BY to get counts by groups.
# + cell_id="00022-92a49f8f-7476-4dc3-8c72-e2d5c61dccef" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1638459122012 source_hash="8aecaebc" tags=[]
query = '''
SELECT species_name, COUNT(*)
FROM species
WHERE genus_name == "Canis"
GROUP BY species_name;
'''
howManyPerGenus = get_data(query)
howManyPerGenus
# + [markdown] cell_id="00028-8021a28d-449a-47b9-8eb0-c66e842157bd" deepnote_cell_type="markdown" tags=[]
# ### MIN(), MAX(), and AVG()
# + [markdown] cell_id="00021-58407c54-5e2a-41a2-a917-233a399e7435" deepnote_cell_type="markdown" tags=[]
# Other mathematical functions can be used to retrieve results directly.
# For example, the code below will get the average, minimum and maximum recordid among all Canis adustus (just demonstrations, the value is meaningless in this context).
# + cell_id="00021-05db26a8-c1ee-4243-8c92-5da6c7edf8ad" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=21 execution_start=1638459122035 source_hash="388b785f" tags=[]
get_data('SELECT AVG(recordid), MIN(recordid), MAX(recordid) FROM species WHERE species_name == "Canis adustus";')
# + [markdown] cell_id="00027-a2dd4d99-4f6d-4a9c-80c8-8e46fda41c3e" deepnote_cell_type="markdown" tags=[]
# ## Connect/combine two tables
#
# To combine multiple tables, we can SELECT columns by naming them table.columnName, e.g. species.recordid will correspond to the column recordid of table species, while bins.recordid will correspond to the recordid column in bins table.
#
# To join two tables we can use JOIN:
# * FROM tableName1 INNER JOIN tableName2
#
# and tell it to JOIN these two tables by column tableName1.x and tableName2.y we use
# * ON tableName1.x = tableName2.y
# + [markdown] cell_id="00032-3f6d8cdc-d488-44f8-9c14-790256cfbaed" deepnote_cell_type="markdown" tags=[]
# ### INNER JOIN and ON
# + cell_id="00029-8d7516ba-136a-4854-b3f0-5e4d34cc517f" deepnote_cell_type="code" deepnote_output_heights=[232.25] deepnote_to_be_reexecuted=false execution_millis=60 execution_start=1638459122061 source_hash="ca4c902a" tags=[]
query = '''
SELECT species.recordid AS "RecordID", species.species_name AS "Species Name", bins.bin_uri AS "BIN"
FROM species INNER JOIN bins
ON species.recordid == bins.recordid
where species_name != "None"
ORDER BY bin_uri DESC
'''
allDataDF = get_data(query)
allDataDF
# + [markdown] cell_id="00027-69309777-471f-4d9d-9109-6bf48518e1ad" deepnote_cell_type="markdown" tags=[]
# Finally, we can combine previous commands and get the count of each BIN connected to a given species (e.g. "Vulpes vulpes")
# + cell_id="00028-e136cd25-243e-4b1c-90a7-942f3156e6b7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=32 execution_start=1638459122122 source_hash="16705d2c" tags=[]
query = '''
SELECT species.species_name AS Species, bins.bin_uri AS BIN, COUNT(*) AS Count
FROM species INNER JOIN bins
ON species.recordid == bins.recordid
WHERE species.genus_name == "Vulpes"
GROUP BY BIN
ORDER BY Species ASC, Count DESC
;
'''
get_data(query)
| SQLite3-database-interaction-from-Python/sqlite3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Anaconda 5)
# language: python
# name: anaconda5
# ---
from othello import *
from collections import namedtuple
# +
import time
class Tick():
def _init_(self):
self.t = time.time()
def tick(self):
self.t = time.time()
print(abs(self.t))
def tock(self):
print(abs(self.t - time.time()))
# +
GameState = namedtuple('GameState', 'to_move, utility, board, moves')
moves = [(x, y) for x in range(1, 9) for y in range(1, 9)]
boardA = {
(1, 5):'W',
(2, 5):'W',
(3, 1):'W',(3, 3):'B',(3, 4):'W',(3, 5):'W',(3, 6):'B',
(4, 1):'W',(4, 2):'W',(4, 3):'B',(4, 4):'B',(4, 5):'W',(4, 6):'B',
(5, 1):'W',(5, 2):'W',(5, 3):'W',(5, 4):'W',(5, 5):'W' ,(5, 6):'B',
(6, 3):'B',(6, 4):'W',(6, 5):'B', (6, 6):'B',
(7, 4):'B',(7, 5):'W', (7, 6):'B',
(8, 1):'B',(8, 2):'B',(8, 3):'B',(8, 4):'B',(8, 5):'B',(8, 6):'B'
}
boardB = {
(3, 6):'B',
(4, 4):'W',(4, 5):'B',(4, 6):'W',
(5, 3):'W',(5, 4):'B',(5, 5):'W',
(6, 3):'B'
}
boardC = {
(1,2):'W',(1,3):'W', (1,4):'W',(1, 5):'W',(1, 6):'W',(1, 7):'W',
(2,1):'W',(2,2):'W',(2,3):'W', (2,4):'W',(2, 5):'W',(2, 6):'W',(2, 7):'W',(2, 8):'W',
(3,1):'W',(3,2):'W',(3,3):'W', (3,4):'W',(3, 5):'W',(3, 6):'W',(3, 7):'W',(3, 8):'W',
(4, 1):'W',(4, 2):'W',(4, 3):'W',(4, 4):'B',(4, 5):'W',(4, 6):'W',(4, 7):'W',(4, 8):'W',
(5, 1):'W',(5, 2):'W',(5, 3):'W',(5, 4):'W',(5, 5):'W',(5, 6):'W',(5, 7):'W',(5, 8):'W',
(6, 1):'W',(6, 2):'W',(6, 3):'W',(6, 4):'W',(6, 5):'W',(6, 6):'W',(6, 7):'W',(6, 8):'W',
(7, 1):'W',(7, 2):'W',(7, 3):'W',(7, 4):'W',(7, 5):'W',(7, 6):'W',(7, 7):'W',(7, 8):'W',
(8, 2):'W',(8, 3):'W',(8, 4):'W',(8, 5):'W',(8, 6):'W',(8, 7):'W'
}
boardD={
(1, 2):'B',(1, 3):'W',(1, 4):'W',(1, 5):'W',(1, 6):'W',(1, 7):'W',
(2, 1):'B',(2, 2):'B',(2, 3):'W',(2, 4):'B',(2, 5):'B',(2, 6):'B',(2, 7):'B',
(3, 2):'B',(3, 3):'W',(3, 4):'W',(3, 5):'W',(3, 6):'B',(3, 7):'B',
(4, 3):'B',(4, 4):'W',(4, 5):'B',(4, 6):'B',(4, 7):'B',
(5, 3):'W',(5, 4):'B',(5, 5):'B',(5, 7):'B',
(6, 4):'B',(6, 5):'B',(6, 6):'B'
}
boardE={
(1,1):'W',(1,3):'B', (1,4):'B',(1, 5):'B',(1, 6):'B',(1, 7):'B',(1, 8):'B',
(2,1):'W',(2,2):'W',(2,3):'B', (2,4):'B',(2, 5):'B',(2, 6):'B',(2, 7):'B',(2, 8):'B',
(3,1):'W',(3,2):'W',(3,3):'W', (3,4):'B',(3, 5):'B',(3, 6):'B',(3, 7):'B',(3, 8):'B',
(4, 1):'W',(4, 2):'W',(4, 3):'B',(4, 4):'W',(4, 5):'B',(4, 6):'B',(4, 7):'B',(4, 8):'B',
(5, 1):'W',(5, 2):'W',(5, 3):'W',(5, 4):'B',(5, 5):'W',(5, 6):'B',(5, 7):'B',(5, 8):'B',
(6, 1):'W',(6, 2):'W',(6, 3):'B',(6, 4):'W',(6, 5):'B',(6, 6):'W',(6, 7):'B',(6, 8):'B',
(7, 1):'W',(7, 2):'W',(7, 3):'B',(7, 4):'B',(7, 5):'W',(7, 6):'B',(7, 7):'W',(7, 8):'B',
(8, 1):'W',(8, 2):'W',(8, 3):'W',(8, 4):'W',(8, 5):'W',(8, 6):'W',(8, 7):'W', (8, 8):'W'
}
for pos in boardD.keys():
moves.remove(pos)
state = GameState(to_move='W',utility=0,board=boardD,moves=moves)
Othello().display(state)
# -
O = Othello()
Othello().actions(state)
O = Othello()
O.display(state)
print(state.moves)
# +
def mobility(state, player):
O = Othello()
if player == 'B':
mine = len(O.legal_moves(state.moves,state.board,'B'))
other = len(O.legal_moves(state.moves,state.board,'W'))
else:
mine = len(O.legal_moves(state.moves,state.board,'W'))
other = len(O.legal_moves(state.moves,state.board,'B'))
if mine > other:
m = (100.0 * mine)/(mine + other)
elif mine < other:
m = -(100.0 * other)/(mine + other)
else:
m = 0
return m
mobility(state,'B')
# +
def corners(state,player):
corners={(1,1), (1,8), (8,1), (8,8)}
Own_Corners = 0
Other_Corners = 0
for coord in corners:
if coord in state.board:
if state.board[coord] == player:
Own_Corners += 1
else:
Other_Corners += 1
else:
pass
if((Own_Corners + Other_Corners) !=0):
Corner_Heuristic= 100 * (Own_Corners - Other_Corners) / (Own_Corners + Other_Corners)
else:
Corner_Heuristic = 0
return Corner_Heuristic
corners(state,'B')
# +
def coin_parity(state,player):
max_value = 'B' if player == 'B' else 'W'
min_value = 'W' if player == 'B' else 'B'
pieces = list(state.board.values())
max_count = pieces.count(max_value)
min_count = pieces.count(min_value)
return 100 * (max_count - min_count) / (max_count + min_count)
coin_parity(state,'B')
# +
def corner_1_1(n,y,x,player,board):
if((y,x) in board and board[y,x] == player):
return corner_1_1(n + 1,y,x+1,player,board)
else:
if(((y+1,1) not in board) or (board[y+1,1] != player)):
return n
else:
return corner_1_1(n,y+1,1,player,board)
def corner_1_8(n,y,x,player,board):
if((y,x) in board and board[y,x] == player):
return corner_1_8(n + 1,y,x-1,player,board)
else:
if(((y+1,8) not in board) or (board[y+1,8] != player)):
return n
else:
return corner_1_8(n,y+1,8,player,board)
def corner_8_1(n,y,x,player,board):
if((y,x) in board and board[y,x] == player):
return corner_8_1(n + 1,y,x+1,player,board)
else:
if(((y-1,1) not in board) or (board[y-1,1] != player)):
return n
else:
return corner_8_1(n,y-1,1,player,board)
def corner_8_8(n,y,x,player,board):
if((y,x) in board and board[y,x] == player):
return corner_8_8(n + 1,y,x-1,player,board)
else:
if(((y-1,8) not in board) or (board[y-1,8] != player)):
return n
else:
return corner_8_8(n,y-1,8,player,board)
def corners(state,player):
n = 0
board = state.board
return corner_1_1(0,1,1,player,board) + corner_1_8(n,1,8,player,board) + corner_8_1(n,8,1,player,board) + corner_8_8(n,8,8,player,board)
def stability(state,player):
max_value = 'B' if player == 'B' else 'W'
min_value = 'W' if player == 'B' else 'B'
if((corners(state,max_value) + corners(state,min_value)) !=0):
sta = 100 * (corners(state,max_value) - corners(state,min_value))/ (corners(state,max_value) + corners(state,min_value))
else:
sta = 0
return sta
stability(state,"W")
# +
def eval_fn(state, player):
return 0.50 * corners(state,player) + 0.25 * mobility(state,player) + 0.15 * stability(state,player) + 0.5 * coin_parity(state,player)
eval_fn(state,'W')
# -
Othello().display(state)
for i in range(0,42):
play_game(Othello(), alphabeta_player(3,eval_fn), random_player)
| Othello.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import torch
from IPython import display
from matplotlib import pyplot as plt
import numpy as np
import random
# ### 生成训练集
num_inputs = 2
num_examples = 1000
true_w = [2, -3.4]
true_b = 4.2
features = torch.from_numpy(np.random.normal(0,1,(num_examples, num_inputs)))
labels = true_w[0] * features[:, 0] + true_w[1] * features[:, 1] +true_b
labels += torch.from_numpy(np.random.normal(0, 0.0001 , size=labels.size()))
features = features.float()
labels = labels.float()
print(features[0], labels[0])
# +
def use_svg_display():
# 用矢量图显示
display.set_matplotlib_formats('svg')
def set_figsize(figsize=(3.5, 2.5)):
use_svg_display()
# 设置图的尺寸
plt.rcParams['figure.figsize'] = figsize
# # 在../d2lzh_pytorch里面添加上面两个函数后就可以这样导入
# import sys
# sys.path.append("..")
# from d2lzh_pytorch import *
set_figsize()
plt.scatter(features[:, 1].numpy(), labels.numpy(), 1);
# -
def data_iter(batch_size, features,labels):
num_examples = len(labels)
num_batch = num_examples // batch_size +1
indices = list (range(num_examples))
random.shuffle(indices)
for i in range(0, num_examples, batch_size):
j = torch.LongTensor(indices[i:min(i+batch_size, num_examples)])
yield features.index_select(0, j), labels.index_select(0, j)
# +
batch_size = 10
for X, y in data_iter(batch_size, features, labels):
print(X, y)
break
# -
w = torch.tensor(np.random.normal(0, 0.01, (num_inputs, 1)), dtype=torch.float32)
b = torch.zeros(1, dtype=torch.float32)
w
w.requires_grad_(requires_grad=True)
b.requires_grad_(requires_grad=True)
# ## 3.2.4 定义模型
#
# 下面是线性回归的矢量计算表达式的实现。我们使用`mm`函数做矩阵乘法。
#
#
#
def linreg(X, w, b): # 本函数已保存在d2lzh_pytorch包中方便以后使用
return torch.mm(X, w) + b
# ## 3.2.5 定义损失函数
#
# 我们使用上一节描述的平方损失来定义线性回归的损失函数。在实现中,我们需要把真实值`y`变形成预测值`y_hat`的形状。以下函数返回的结果也将和`y_hat`的形状相同。
def squared_loss(y_hat, y):
return (y_hat - y.view(y_hat.size()))**2/2
# ## 3.2.6 定义优化算法
#
# 以下的`sgd`函数实现了上一节中介绍的小批量随机梯度下降算法。它通过不断迭代模型参数来优化损失函数。这里自动求梯度模块计算得来的梯度是一个批量样本的梯度和。我们将它除以批量大小来得到平均值。
#
def sgd(params, lr , batch_size):
for param in params:
param.data -= lr * param.grad/batch_size
# ## 3.2.7 训练模型
#
# 在训练中,我们将多次迭代模型参数。在每次迭代中,我们根据当前读取的小批量数据样本(特征`X`和标签`y`),通过调用反向函数`backward`计算小批量随机梯度,并调用优化算法`sgd`迭代模型参数。由于我们之前设批量大小`batch_size`为10,每个小批量的损失`l`的形状为(10, 1)。回忆一下自动求梯度一节。由于变量`l`并不是一个标量,所以我们可以调用`.sum()`将其求和得到一个标量,再运行`l.backward()`得到该变量有关模型参数的梯度。注意在每次更新完参数后不要忘了将参数的梯度清零。
#
# 在一个迭代周期(epoch)中,我们将完整遍历一遍`data_iter`函数,并对训练数据集中所有样本都使用一次(假设样本数能够被批量大小整除)。这里的迭代周期个数`num_epochs`和学习率`lr`都是超参数,分别设3和0.03。在实践中,大多超参数都需要通过反复试错来不断调节。虽然迭代周期数设得越大模型可能越有效,但是训练时间可能过长。而有关学习率对模型的影响,我们会在后面“优化算法”一章中详细介绍。
# +
lr = 0.03
num_epochs = 3
net = linreg
loss = squared_loss
for epoch in range (num_epochs):
for X,y in data_iter(batch_size, features,labels):
L = loss (net(X, w, b), y ).sum()
L.backward()
sgd([w, b], lr, batch_size)
# 不要忘了梯度清零
w.grad.data.zero_()
b.grad.data.zero_()
train_L = loss(net(features,w,b), labels)
print('epoch %d, loss %f' % (epoch + 1, train_L.mean().item()))
# -
print(true_w, '\n', w)
print(true_b, '\n', b)
# # 3.3 线性回归的简洁实现
#
# 随着深度学习框架的发展,开发深度学习应用变得越来越便利。实践中,我们通常可以用比上一节更简洁的代码来实现同样的模型。在本节中,我们将介绍如何使用PyTorch更方便地实现线性回归的训练。
#
# ## 3.3.1 生成数据集
#
# 我们生成与上一节中相同的数据集。其中`features`是训练数据特征,`labels`是标签。
num_inputs = 2
num_examples = 1000
true_w = [2, -3.4]
true_b = 4.2
features = torch.tensor(np.random.normal(0,1,(num_examples, num_inputs)))
labels = true_w[0] * features[: , 0] +true_w[1] * features[:, 1] +true_b
print(labels.size())
labels = labels + torch.tensor(np.random.normal(0, 0.01,size = labels.size() ))
features = features.float()
labels = labels.float()
# ## 3.3.2 读取数据
#
# PyTorch提供了`data`包来读取数据。由于`data`常用作变量名,我们将导入的`data`模块用`Data`代替。在每一次迭代中,我们将随机读取包含10个数据样本的小批量。
# +
import torch.utils.data as Data
batch_size = 10
# 将训练数据的特征和标签组合
dataset = Data.TensorDataset(features, labels)
# 随机读取小批量
data_iter = Data.DataLoader(dataset, batch_size, shuffle=True)
# -
for X,y in data_iter:
print(X, y)
break
# ### 定义模型
import torch.nn as nn
# +
class LinearNet(nn.Module):
def __init__(self, n_feature):
super (LinearNet,self).__init__()
self.linear = nn.Linear(n_feature, 1)
#forward 定义前向传播
def forward(self, x):
y = self.linear(x)
return y
net = LinearNet(num_inputs)
print(net)
# -
# 事实上我们还可以用`nn.Sequential`来更加方便地搭建网络,`Sequential`是一个有序的容器,网络层将按照在传入`Sequential`的顺序依次被添加到计算图中。
# +
# # 写法一
# net = nn.Sequential(
# nn.Linear(num_inputs, 1)
# # 此处还可以传入其他层
# )
# # 写法二
# net = nn.Sequential()
# net.add_module('linear', nn.Linear(num_inputs, 1))
# # net.add_module ......
net =[3]
# 写法三
from collections import OrderedDict
net = nn.Sequential(OrderedDict([
('linear', nn.Linear(num_inputs,1))
# ......
]))
print(net)
print(net[0])
# -
for para in net.parameters():
print(para)
from torch.nn import init
init.normal_(net[0].weight , mean = 0, std =0.01)
init.constant_(net[0].bias , val=0) # 也可以直接修改bias的data: net[0].bias.data.fill_(0)
loss = nn.MSELoss()
import torch.optim as optim
optimizer = optim.SGD(net.parameters(), lr = 0.03)
print(optimizer)
'''python
optimizer =optim.SGD([
# 如果对某个参数不指定学习率,就使用最外层的默认学习率
{'params': net.subnet1.parameters()}, # lr=0.03
{'params': net.subnet2.parameters(), 'lr': 0.01}
], lr=0.03)
'''
# 调整学习率
for param_group in optimizer.param_groups:
param_group['lr'] *= 0.1 # 学习率为之前的0.1倍
num_epochs = 8
for epoch in range(1, num_epochs+1):
for X,y in data_iter:
output = net(X)
I = loss (output, y.view(-1,1))
optimizer.zero_grad()
I.backward()
optimizer.step()
print('epoch %d, loss:%f' %(epoch, I.item()))
dense = net[0]
print(true_w, dense.weight)
print(true_b, dense.bias)
# ## 小结
#
# * 使用PyTorch可以更简洁地实现模型。
# * `torch.utils.data`模块提供了有关数据处理的工具,`torch.nn`模块定义了大量神经网络的层,`torch.nn.init`模块定义了各种初始化方法,`torch.optim`模块提供了模型参数初始化的各种方法。
#
# -----------
# > 注:本节除了代码之外与原书基本相同,[原书传送门](https://zh.d2l.ai/chapter_deep-learning-basics/linear-regression-gluon.html)
| mycode/linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import numpy as np
import snsims
import healpy as hp
from astropy.cosmology import Planck15 as cosmo
zdist = snsims.PowerLawRates(rng=np.random.RandomState(1),
fieldArea=9.6,
surveyDuration=10.,
zbinEdges=np.arange(0.10001, 1.1, 0.1))
# ten years
zdist.DeltaT
# The sky is >~ 40000 sq degrees ~ 4000 * LSST field of view
zdist.skyFraction * 2000 * 2
zdist.zSampleSize().sum()
# To compare wih David's rate divide by number of days (note bins go from 0.1 to 1.0 in steps of 0.1)
zdist.zSampleSize() / 3650.
# Get samples of those numbers and histogram (consistency, should not be new information)
np.histogram(zdist.zSamples, np.arange(0.1, 1.01, 0.1))[0]/3650.
otherEstimate = np.array([0.0297949656, 0.0773033283291, 0.143372148164, 0.224245838014, 0.315868414203, 0.413859866222
, 0.513435128451, 0.609346340913, 0.696255385228])
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
fig, ax = plt.subplots()
zvals = np.arange(0.15, 0.96, 0.1)
ax.plot(zvals, zdist.zSampleSize()/ 3650., 'or', label='snsims ')
ax.plot(zvals, otherEstimate, 'bs', label='<NAME>')
ax.set_xlabel('z')
ax.set_ylabel('numbers per field per day')
# ## Parts of the calculation
zbin_edges = np.arange(0.1, 1.01, 0.1)
diff_volume = cosmo.comoving_volume(zbin_edges[1:]) - cosmo.comoving_volume(zbin_edges[:-1])
print diff_volume
fig_subs, axs = plt.subplots(3)
axs[0].plot(zvals, zdist.snRate(zvals), 'or')
axs[1].plot(zvals, diff_volume , 'or')
axs[2].plot(zvals, diff_volume * zdist.snRate(zvals)*10.0/40000. / 365.0, 'or')
axs[2].set_xlabel('z')
axs[0].set_ylabel('rate')
axs[1].set_ylabel('comoving vol')
axs[2].set_ylabel('vol X skyfrac X time')
# ## Total Number of SN
zdist = snsims.PowerLawRates(rng=np.random.RandomState(1),
fieldArea=18000.,
surveyDuration=10.,
zbinEdges=np.arange(0.010001, 0.901, 0.1))
zdist.zSampleSize().sum() /1.0e6
fig, ax = plt.subplots()
_ = ax.hist(zdist.zSamples, bins=np.arange(0.001, 1.4, 0.05), histtype='step', lw=2., alpha=1.)
arcmin = 1.0 / 60.
zdist = snsims.PowerLawRates(rng=np.random.RandomState(1),
fieldArea=10.,
surveyDuration=10.,
zbinEdges=np.arange(0.010001, 0.901, 0.05))
np.array(map(np.float, zdist.numSN())) /5.
np.pi * (1.0 / 12.)**2
10.0 / 200.
| examples/Comparison_SNRates_DavidRubin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.feature_extraction.text import TfidfTransformer, CountVectorizer, TfidfVectorizer
from sklearn.linear_model import LogisticRegression, SGDClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
from sklearn.model_selection import cross_val_score
from sklearn.pipeline import Pipeline
import numpy as np
import pandas as pd
# +
file = open('SMSSpamCollection.txt', 'r', encoding = 'utf-8')
lines = file.readlines()
labels = []
features = []
for line in lines:
labels.append(line.split('\t')[0])
features.append(line.split('\t')[1].replace('\n', ''))
map_dict = {'ham' : 0, 'spam' : 1}
# -
labels = list(map(map_dict.get, labels))
# vect1 = CountVectorizer(ngram_range=(1,3))
vect1 = TfidfVectorizer()
X = vect1.fit_transform(features)
# +
model = LogisticRegression(random_state = 2)
cross_val_score(estimator = model, scoring = 'f1', cv = 10, X = X, y = labels).mean()
# model.fit(X, labels)
# -
test = ["FreeMsg: Txt: CALL to No: 86888 & claim your reward of 3 hours talk time to use from your phone now! Subscribe6GB",
"FreeMsg: Txt: claim your reward of 3 hours talk time",
"Have you visited the last lecture on physics?",
"Have you visited the last lecture on physics? Just buy this book and you will have all materials! Only 99$",
"Only 99$"]
model.predict(vect1.transform(test))
| Yandex data science/5/Week 3/Text analysis example.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
// This notebook was inspired by [Plottting with XPlot](https://github.com/dotnet/interactive/blob/master/NotebookExamples/fsharp/Docs/Plotting%20with%20Xplot.ipynb).
open XPlot.Plotly
// +
let bar =
Bar(
name = "Bar 1",
x = ["A"; "B"; "C"],
y = [1; 3; 2])
[bar]
|> Chart.Plot
|> Chart.WithTitle "A sample bar plot"
| tests/notebooks/ipynb_fs/fsharp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TRTR and TSTR Results Comparison
# +
#import libraries
import warnings
warnings.filterwarnings("ignore")
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
pd.set_option('precision', 4)
# -
# ## 1. Create empty dataset to save metrics differences
DATA_TYPES = ['Real','GM','SDV','CTGAN','WGANGP']
SYNTHESIZERS = ['GM','SDV','CTGAN','WGANGP']
ml_models = ['RF','KNN','DT','SVM','MLP']
# ## 2. Read obtained results when TRTR and TSTR
FILEPATHS = {'Real' : 'RESULTS/models_results_real.csv',
'GM' : 'RESULTS/models_results_gm.csv',
'SDV' : 'RESULTS/models_results_sdv.csv',
'CTGAN' : 'RESULTS/models_results_ctgan.csv',
'WGANGP' : 'RESULTS/models_results_wgangp.csv'}
#iterate over all datasets filepaths and read each dataset
results_all = dict()
for name, path in FILEPATHS.items() :
results_all[name] = pd.read_csv(path, index_col='model')
results_all
# ## 3. Calculate differences of models
# +
metrics_diffs_all = dict()
real_metrics = results_all['Real']
columns = ['data','accuracy_diff','precision_diff','recall_diff','f1_diff']
metrics = ['accuracy','precision','recall','f1']
for name in SYNTHESIZERS :
syn_metrics = results_all[name]
metrics_diffs_all[name] = pd.DataFrame(columns = columns)
for model in ml_models :
real_metrics_model = real_metrics.loc[model]
syn_metrics_model = syn_metrics.loc[model]
data = [model]
for m in metrics :
data.append(abs(real_metrics_model[m] - syn_metrics_model[m]))
metrics_diffs_all[name] = metrics_diffs_all[name].append(pd.DataFrame([data], columns = columns))
metrics_diffs_all
# -
# ## 4. Compare absolute differences
# ### 4.1. Barplots for each metric
# +
metrics = ['accuracy', 'precision', 'recall', 'f1']
metrics_diff = ['accuracy_diff', 'precision_diff', 'recall_diff', 'f1_diff']
colors = ['tab:blue', 'tab:orange', 'tab:green', 'tab:red', 'tab:purple']
barwidth = 0.15
fig, axs = plt.subplots(nrows=1, ncols=4, figsize=(15, 2.5))
axs_idxs = range(4)
idx = dict(zip(metrics + metrics_diff,axs_idxs))
for i in range(0,len(metrics)) :
data = dict()
y_pos = dict()
y_pos[0] = np.arange(len(ml_models))
ax = axs[idx[metrics[i]]]
for k in range(0,len(DATA_TYPES)) :
generator_data = results_all[DATA_TYPES[k]]
data[k] = [0, 0, 0, 0, 0]
for p in range(0,len(ml_models)) :
data[k][p] = generator_data[metrics[i]].iloc[p]
ax.bar(y_pos[k], data[k], color=colors[k], width=barwidth, edgecolor='white', label=DATA_TYPES[k])
y_pos[k+1] = [x + barwidth for x in y_pos[k]]
ax.set_xticks([r + barwidth*2 for r in range(len(ml_models))])
ax.set_xticklabels([])
ax.set_xticklabels(ml_models, fontsize=10)
ax.set_title(metrics[i], fontsize=12)
ax.legend(DATA_TYPES, ncol=5, bbox_to_anchor=(-0.3, -0.2))
fig.tight_layout()
#fig.suptitle('Models performance comparisson Boxplots (TRTR and TSTR) \n Dataset F - Indian Liver Patient', fontsize=18)
fig.savefig('RESULTS/MODELS_METRICS_BARPLOTS.svg', bbox_inches='tight')
# +
metrics = ['accuracy_diff', 'precision_diff', 'recall_diff', 'f1_diff']
colors = ['tab:orange', 'tab:green', 'tab:red', 'tab:purple']
fig, axs = plt.subplots(nrows=1, ncols=4, figsize=(15,2.5))
axs_idxs = range(4)
idx = dict(zip(metrics,axs_idxs))
for i in range(0,len(metrics)) :
data = dict()
ax = axs[idx[metrics[i]]]
for k in range(0,len(SYNTHESIZERS)) :
generator_data = metrics_diffs_all[SYNTHESIZERS[k]]
data[k] = [0, 0, 0, 0, 0]
for p in range(0,len(ml_models)) :
data[k][p] = generator_data[metrics[i]].iloc[p]
ax.plot(data[k], 'o-', color=colors[k], label=SYNTHESIZERS[k])
ax.set_xticks(np.arange(len(ml_models)))
ax.set_xticklabels(ml_models, fontsize=10)
ax.set_title(metrics[i], fontsize=12)
ax.set_ylim(bottom=-0.01, top=0.43)
ax.grid()
ax.legend(SYNTHESIZERS, ncol=5, bbox_to_anchor=(-0.4, -0.2))
fig.tight_layout()
#fig.suptitle('Models performance comparisson Boxplots (TRTR and TSTR) \n Dataset F - Indian Liver Patient', fontsize=18)
fig.savefig('RESULTS/MODELS_METRICS_DIFFERENCES.svg', bbox_inches='tight')
# -
| notebooks/Dataset B - Cardiovascular Disease/Synthetic data evaluation/Utility/TRTR and TSTR Results Comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploration of ENRON Dataset
# #### by <NAME>
#
#
# ## Table of Contents
#
# 1. Introduction
# 2. Data Gathering
# 3. Data Assessing + Cleaning
# 4. Exploration Data Analysis (EDA)
# - Univariate
# - Bivariate
# - Multivariate
# 5. Conclusions
#
#
#
# <br><br>
# # 1. Introduction
#
# TStarter project code for students taking Udacity ud120.
#
# The whole project idea is from Udacity's Introduction to Machine Learning Course https://classroom.udacity.com/courses/ud120
#
# ### ENRON-DATA-Udacity
#
# The dataset can be found in http://www.cs.cmu.edu/~./enron/
#
# The [Enron Corpus](https://en.wikipedia.org/wiki/Enron_Corpus) is a large database of over 600,000 emails generated by 158 employees of the Enron Corporation and acquired by the Federal Energy Regulatory Commission during its investigation after the company's collapse.
#
# # 2. Gather the Enron data
# +
import sys
import pickle
sys.path.append("../final_project/")
from feature_format import featureFormat, targetFeatureSplit
#from tester import dump_classifier_and_data
import pandas as pd
from matplotlib import pyplot as plt
### Task 1: Select what features you'll use. Features_list is a list of strings, each of which is a feature name.
### The first feature must be "poi".
### Load the dictionary containing the dataset
with open("final_project_dataset.pkl", "rb") as data_file:
data_dict = pickle.load(data_file)
# enron_data = pickle.load(open("../final_project/final_project_dataset.pkl", "rb"))
# -
import numpy as np
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV #from sklearn.grid_search import GridSearchCV
from sklearn.model_selection import train_test_split
from time import time
# +
# converting the given pickled Enron data to a pandas dataframe
enron_df = pd.DataFrame.from_records(list(data_dict.values()))
# set the index of df to be the employees series:
employees = pd.Series(list(data_dict.keys()))
enron_df.set_index(employees, inplace=True)
enron_df.head()
# -
# <br><br>
# # 2. Wrangle Enron data (Assess + Clean)
print ("Size of the enron dataframe:",enron_df.shape)
print ("Number of data points(people) in the dataset:", len(enron_df))
print ("To find the number of Features in the Enron Dataset : ",len(enron_df.columns))
poi_t = enron_df.groupby('poi').size()
poi_t
type(poi_t)
print ("Total number of non-POI's in the given dataset : ",poi_t.iloc[0])
print ("Total number of POI's in the given dataset : ",poi_t.iloc[1])
# So, in the given dataset there are 18 POI's and 128 non-POI's.
#
enron_df.dtypes
# Coerce numeric values into floats or ints; also change NaN to zero:
enron_df_new = enron_df.apply(lambda x : pd.to_numeric(x, errors = 'coerce')).copy().fillna(0)
enron_df_new.head()
enron_df_new.dtypes
# Removing the column of __email_address__ from the enron_df as it is not of much used in this project.
# Dropping column 'email_address' as not required in analysis
enron_df_new.drop('email_address', axis = 1, inplace = True)
enron_df_new.head()
# Checking the changed shape of df
enron_df_new.shape
enron_df_new.describe()
# <br><br>
# # 3. Explore / analyzing the features of Enron Dataset
# ## 3.1. Financial Features
#
# ### 3.1.1. Bonus and Salary
# +
# Drawing scatterplot
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == True],enron_df_new['bonus'][enron_df_new['poi'] == True], color = 'r',
label = 'POI')
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == False],enron_df_new['bonus'][enron_df_new['poi'] == False],color = 'b',
label = 'Not-POI')
plt.xlabel("Salary")
plt.ylabel("Bonus")
plt.title("Scatterplot of salary vs bonus w.r.t POI")
plt.legend(loc='upper left')
plt.show()
# -
# From the above figure, one point has high value of salary and bonus.
enron_df_new['salary'].argmax()
enron_df_new.loc['TOTAL',:]
enron_df_new.iloc()
# ## Removing Outlier 1 : 'TOTAL'
# So the 'TOTAL' row is removed from the above df.
enron_df_new.drop('TOTAL', axis = 0, inplace = True)
enron_df_new.shape
enron_df_new.describe()
# So the scatterplot for the changed df is.
# +
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == True],enron_df_new['bonus'][enron_df_new['poi'] == True], color = 'r', label = 'POI')
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == False],enron_df_new['bonus'][enron_df_new['poi'] == False],color = 'b', label = 'Not-POI')
plt.xlabel("Salary")
plt.ylabel("Bonus")
plt.title("Scatterplot of salary vs bonus w.r.t POI")
plt.legend(loc='upper left')
plt.show()
# -
# As the POI's were taking larger amounts of money as bonus, in addition to their high salary so it can be stated that the ratio of bonus to salary of the POI's will be higher as compared to that of non-POI's.
# #### Feature created : bonus-to-salary_ratio
# Created a new feature
enron_df_new['bonus-to-salary_ratio'] = enron_df_new['bonus']/enron_df_new['salary']
# +
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == True], enron_df_new['bonus-to-salary_ratio'][enron_df_new['poi'] == True], color = 'r', label = 'POI')
plt.scatter(enron_df_new['salary'][enron_df_new['poi'] == False], enron_df_new['bonus-to-salary_ratio'][enron_df_new['poi'] == False],color = 'b', label = 'Not-POI')
plt.xlabel("Salary")
plt.ylabel("Bonus-to-salary ratio")
plt.title("Scatterplot of salary vs bonus-to-salary ratio - w.r.t POI")
plt.legend(loc='upper left')
plt.show()
# -
# ## Removing Outlier 2 : 'THE TRAVEL AGENCY IN THE PARK'
enron_df_new.loc['THE TRAVEL AGENCY IN THE PARK']
enron_df_new.drop('THE TRAVEL AGENCY IN THE PARK', axis = 0, inplace = True)
enron_df_new.shape
# ## 2. Deferred_income, deferred_payment and total_payment
#
# According to [BusinessDictionary.com](http://www.businessdictionary.com/definition/deferred-payment.html) :
# - Deferred payment is "a loan arrangement in which the borrower is allowed to start making payments at some specified time in the future. Deferred payment arrangements are often used in retail settings where a person buys and receives an item with a commitment to begin making payments at a future date."
#
# - [Deferred income](https://en.wikipedia.org/wiki/Deferred_income) : (also known as deferred revenue, unearned revenue, or unearned income) is, in accrual accounting, money received for goods or services which have not yet been delivered. According to the revenue recognition principle, it is recorded as a liability until delivery is made, at which time it is converted into revenue.
#
# As Enron scam involved a lot of undisclosed assets and cheating public by selling assets to shell companies at end of each month and buying them back at the start of next month to hide the acounting losses so there are chances that lot of deferred revenue by the company was used by the POI's.
enron_df_new['deferred_income'].describe()
# The __deferred_income__ feature has mostly negative values as it is the money which has to be returned by the company.
# +
# Finding out the integer index locations of POIs and non-POIs
poi_rs = []
non_poi_rs = []
for i in range(len(enron_df_new['poi'])):
if enron_df_new['poi'][i] == True:
poi_rs.append(i+1)
else:
non_poi_rs.append(i+1)
print ("length poi list : ",len(poi_rs))
print ("length non-poi list : ",len(non_poi_rs))
# +
plt.scatter(non_poi_rs,
enron_df_new['deferred_income'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(poi_rs,
enron_df_new['deferred_income'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('Employees')
plt.ylabel('deferred_income')
plt.title("Scatterplot of Employees with deferred income")
plt.legend(loc='upper right')
plt.show()
# +
plt.scatter(enron_df_new['total_payments'][enron_df_new['poi'] == False],
enron_df_new['deferral_payments'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(enron_df_new['total_payments'][enron_df_new['poi'] == True],
enron_df_new['deferral_payments'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('total_payments')
plt.ylabel('deferral_payments')
plt.title("Scatterplot of total_payments vs deferral_payments w.r.t POI")
plt.legend(loc='upper right')
plt.show()
# -
# From the above scatterplot it can be observed that majority of POIs have very low value of deferral payments as compared to the deferral_payments of non-POIs.
#
# So, from the above we can observe there are two outliers. The one having high value of total_payments is a POI and the other outlier with high value of deferral payments is a non-POI. I am removing the non-POI outlier.
enron_df_new['deferral_payments'].argmax()
# ### remove the 3rd outlier for 'deferral_payments'
enron_df_new.drop('FREVERT MARK A', axis = 0, inplace = True)
enron_df_new.shape
# Finding out the integer index locations of POIs and non-POIs
poi_rs = []
non_poi_rs = []
for i in range(len(enron_df_new['poi'])):
if enron_df_new['poi'][i] == True:
poi_rs.append(i+1)
else:
non_poi_rs.append(i+1)
# +
plt.scatter(non_poi_rs,
enron_df_new['restricted_stock'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(poi_rs,
enron_df_new['restricted_stock'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('Employees')
plt.ylabel('restricted_stock')
plt.title("Scatterplot of Employee number with restricted stock")
plt.legend(loc='upper right')
plt.show()
# -
enron_df_new['restricted_stock'].argmax()
# #### 3. long_term_incentive
# +
plt.scatter(non_poi_rs,
enron_df_new['long_term_incentive'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(poi_rs,
enron_df_new['long_term_incentive'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('Employees')
plt.ylabel('long_term_incentive')
plt.title("Scatterplot of Employee number with long_term_incentive")
plt.legend(loc='upper left')
plt.show()
# -
enron_df_new['long_term_incentive'].argmax()
# ### Remove the 4th outlier for 'long_term_incentive'
enron_df_new.drop('<NAME>', axis = 0, inplace = True)
enron_df_new.shape
# #### 4. restricted_stock and restricted_stock_deferred
enron_df_new['restricted_stock_deferred'].describe()
enron_df_new['restricted_stock'].describe()
# +
# Scatterplot of restricted_stock vs 'restricted_stock_deferred' w.r.t POI
plt.scatter(enron_df_new['restricted_stock'][enron_df_new['poi'] == False],
enron_df_new['restricted_stock_deferred'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(enron_df_new['restricted_stock'][enron_df_new['poi'] == True],
enron_df_new['restricted_stock_deferred'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('restricted_stock')
plt.ylabel('restricted_stock_deferred')
plt.title("Scatterplot of restricted_stock vs 'restricted_stock_deferred' w.r.t POI")
plt.legend(loc='upper right')
plt.show()
# -
# So obtained an outlier in the feature __restricted_stock_deferred__. Also taking a quick look at the values of __restricted_stock_deferred__ most of the values are zeros and the remaining few are negative values. The outlier found here is for the enron employee __<NAME>__ who is not a POI and in this analysis i am removing this datapoint hoping that it may aid in classification.
#
# And at the other axis of the graph, the other maximum values are of a POI and a non-POI so no need to remove them.
enron_df_new['restricted_stock_deferred'].argmax()
enron_df_new.loc['BHATNAGAR SANJAY']['poi']
# ### Did not remove the outlier for 'restricted_stock_deferred'
enron_df_new.drop('BHATNAGAR SANJAY', axis = 0, inplace = True)
enron_df_new.shape
# #### 5. expenses
# Finding out the integer index locations of POIs and non-POIs
poi_rs = []
non_poi_rs = []
for i in range(len(enron_df_new['poi'])):
if enron_df_new['poi'][i] == True:
poi_rs.append(i+1)
else:
non_poi_rs.append(i+1)
# +
plt.scatter(non_poi_rs,
enron_df_new['expenses'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(poi_rs,
enron_df_new['expenses'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('Employees')
plt.ylabel('expenses')
plt.title("Scatterplot of Employee number with expenses")
plt.legend(loc='upper right')
plt.show()
# +
plt.scatter(non_poi_rs,
enron_df_new['deferred_income'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(poi_rs,
enron_df_new['deferred_income'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('Employees')
plt.ylabel('deferred_income')
plt.title("Scatterplot of Employees with deferred income")
plt.legend(loc='upper right')
plt.show()
# -
# ## Email-Features
# Also it can be thought that for doing such a big scam the POI's might have frequent contact between them via E-mails so by checking on the number of e-mails transferred between POIs and an Employee we can be able to guess for the involvement of that person in that scam. So finding the fraction of the mail from and to this person with respect to the POI.
#
# #### 1. from_poi_to_this_person and from_this_person_to_poi
# +
plt.scatter(enron_df_new['from_poi_to_this_person'][enron_df_new['poi'] == False],
enron_df_new['from_this_person_to_poi'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(enron_df_new['from_poi_to_this_person'][enron_df_new['poi'] == True],
enron_df_new['from_this_person_to_poi'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('from_poi_to_this_person')
plt.ylabel('from_this_person_to_poi')
plt.title("Scatterplot of count of from and to mails between poi and this_person w.r.t POI")
plt.legend(loc='upper right')
plt.show()
# -
# #### Features created : fraction_mail_from_poi and fraction_mail_to_poi
enron_df_new['fraction_mail_from_poi'] = enron_df_new['from_poi_to_this_person']/enron_df_new['from_messages']
enron_df_new['fraction_mail_to_poi'] = enron_df_new['from_this_person_to_poi']/enron_df_new['to_messages']
# +
plt.scatter(enron_df_new['fraction_mail_from_poi'][enron_df_new['poi'] == False],
enron_df_new['fraction_mail_to_poi'][enron_df_new['poi'] == False],
color = 'b', label = 'Not-POI')
plt.scatter(enron_df_new['fraction_mail_from_poi'][enron_df_new['poi'] == True],
enron_df_new['fraction_mail_to_poi'][enron_df_new['poi'] == True],
color = 'r', label = 'POI')
plt.xlabel('fraction_mail_from_poi')
plt.ylabel('fraction_mail_to_poi')
plt.title("Scatterplot of fraction of mails from and to between poi and this_person w.r.t POI")
plt.legend(loc='upper right')
plt.show()
# -
# From the above figure, the difference between POIs and non-POIs points can be clearly classified.
#
# So the total number of features in the dataframe after the data analysis is.
enron_df_new.columns
enron_df_new.shape
new_features_list = enron_df_new.columns.values
new_features_list
#clean all 'inf' values which we got if the person's from_messages = 0
enron_df_new = enron_df_new.replace('inf', 0)
enron_df_new = enron_df_new.fillna(0)
# Converting the above modified dataframe to a dictionary
enron_dict = enron_df_new.to_dict('index')
print ("Features of modified data_dictionary \n")
print ("Total number of datapoints : ",len(enron_dict))
print ("Total number of features : ",len(enron_dict['METTS MARK']))
# +
enron_dict['METTS MARK']
#enron_data["PRENTICE JAMES"]
# -
for key, value in enron_dict.items():
print (key)
print ("\n")
for val in enron_dict[key].keys():
print (val)
break
# Store to my_dataset for easy export below.
my_dataset = enron_dict
## Selecting features which i think might be important
features_list = ['poi', 'salary', 'bonus', 'long_term_incentive', 'bonus-to-salary_ratio', 'deferral_payments', 'expenses',
'restricted_stock_deferred', 'restricted_stock', 'deferred_income','fraction_mail_from_poi', 'total_payments',
'other', 'fraction_mail_to_poi', 'from_poi_to_this_person', 'from_this_person_to_poi', 'to_messages',
'from_messages', 'shared_receipt_with_poi', 'loan_advances', 'director_fees', 'exercised_stock_options',
'total_stock_value', 'restricted_stock']
len(features_list)
# Extract features and labels from dataset for local testing
data = featureFormat(my_dataset, features_list, sort_keys = True)
labels, features = targetFeatureSplit(data)
# # Task 4: Try a varity of classifiers
# +
### split data into training and testing datasets
from sklearn.model_selection import cross_validate #from sklearn import cross_validation
features_train, features_test, labels_train, labels_test = train_test_split(features, labels, test_size=0.3, random_state=42)
# Stratified ShuffleSplit cross-validator
from sklearn.model_selection import StratifiedShuffleSplit
sss = StratifiedShuffleSplit(n_splits=100, test_size=0.3,random_state = 42)
# -
# Importing modules for feature scaling and selection
from sklearn.preprocessing import MinMaxScaler
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.decomposition import PCA
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
# Defining functions to be used via the pipeline
scaler = MinMaxScaler()
skb = SelectKBest(f_classif)
pca = PCA()
# ### Classifer 1 : Naive Bayes
import warnings
warnings.filterwarnings('ignore')
# +
from sklearn.naive_bayes import GaussianNB
clf_gnb = GaussianNB()
pipeline = Pipeline(steps = [("SKB", skb), ("NaiveBayes",clf_gnb)])
param_grid = {"SKB__k":[7,8,9,10,11,12,13,14,15,16,17,18,19]}
grid = GridSearchCV(pipeline, param_grid, verbose = 0, cv = sss, scoring = 'f1')
t0 = time()
grid.fit(features, labels)
print ("training time: ", round(time()-t0, 3), "s")
# best algorithm
clf = grid.best_estimator_
t0 = time()
# refit the best algorithm:
clf.fit(features_train, labels_train)
prediction = clf.predict(features_test)
print ("testing time: ", round(time()-t0, 3), "s")
# -
print ("Accuracy of GaussianNB classifer is : ",accuracy_score(labels_test, prediction))
print ("Precision of GaussianNB classifer is : ",precision_score(prediction, labels_test))
print ("Recall of GaussianNB classifer is : ",recall_score(prediction, labels_test))
print ("f1-score of GaussianNB classifer is : ",f1_score(prediction, labels_test))
grid.best_estimator_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_scores
grid.best_params_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_selected_scores = feature_scores[features_selected_bool]
feature_selected_scores
imp_features_df = pd.DataFrame({'Features_Selected':features_selected_list, 'Features_score':feature_selected_scores})
imp_features_df.sort_values('Features_score', ascending = False,inplace = True)
Rank = pd.Series(list(range(1,len(features_selected_list)+1)))
imp_features_df.set_index(Rank, inplace = True)
imp_features_df
# ## Classifier 2 : Decision Tree without PCA
# +
from sklearn.tree import DecisionTreeClassifier
clf_dtree = DecisionTreeClassifier()
pipeline = Pipeline(steps = [("SKB", skb), ("dtree",clf_dtree)])
param_grid = {"SKB__k":[7,8,9,10,11,12,13,14,15,16,17,18,19],
"dtree__criterion": ["gini", "entropy"],
"dtree__min_samples_split": [2, 4, 8, 10]}
grid = GridSearchCV(pipeline, param_grid, verbose = 0, cv = sss, scoring = 'f1')
t0 = time()
#clf = clf.fit(features_train, labels_train)
grid.fit(features, labels)
print ("training time: ", round(time()-t0, 3), "s")
# best algorithm
clf = grid.best_estimator_
t0 = time()
# refit the best algorithm:
clf.fit(features_train, labels_train)
prediction = clf.predict(features_test)
print ("testing time: ", round(time()-t0, 3), "s")
print ("Accuracy of DT classifer is : ",accuracy_score(labels_test, prediction))
print ("Precision of DT classifer is : ",precision_score(prediction, labels_test))
print ("Recall of DT classifer is : ",recall_score(prediction, labels_test))
print ("f1-score of DT classifer is : ",f1_score(prediction, labels_test))
# -
# View the best parameters for the model found using grid search
print ('Best criterion:',grid.best_estimator_.named_steps['dtree'].criterion)
grid.best_params_
grid.best_estimator_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_scores
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_selected_scores = feature_scores[features_selected_bool]
feature_selected_scores
imp_features_df = pd.DataFrame({'Features_Selected':features_selected_list, 'Features_score':feature_selected_scores})
imp_features_df.sort_values('Features_score', ascending = False,inplace = True)
Rank = pd.Series(list(range(1,len(features_selected_list)+1)))
imp_features_df.set_index(Rank, inplace = True)
imp_features_df
# ## Classifier X. DT using SKB(est) and PCA
# +
from sklearn.tree import DecisionTreeClassifier
clf_dtree = DecisionTreeClassifier()
pipeline = Pipeline(steps = [("SKB", skb), ("PCA",pca), ("dtree",clf_dtree)])
param_grid = {"SKB__k":[7,8,9,10,11,12,13,14,15,16,17,18],
"PCA__n_components":[2,3,4,5,6,7],
"PCA__whiten":[True],
"dtree__criterion": ["gini", "entropy"],
"dtree__min_samples_split": [2, 4, 8, 10]}
grid = GridSearchCV(pipeline, param_grid, verbose = 0, cv = sss, scoring = 'f1')
t0 = time()
#clf = clf.fit(features_train, labels_train)
grid.fit(features, labels)
print ("training time: ", round(time()-t0, 3), "s")
# best algorithm
clf = grid.best_estimator_
t0 = time()
# refit the best algorithm:
clf.fit(features_train, labels_train)
prediction = clf.predict(features_test)
print ("testing time: ", round(time()-t0, 3), "s")
print ("Accuracy of DT classifer is : ",accuracy_score(labels_test, prediction))
print ("Precision of DT classifer is : ",precision_score(prediction, labels_test))
print ("Recall of DT classifer is : ",recall_score(prediction, labels_test))
print ("f1-score of DT classifer is : ",f1_score(prediction, labels_test))
# -
# View the best parameters for the model found using grid search
print ('Best criterion:',grid.best_estimator_.named_steps['dtree'].criterion)
grid.best_params_
grid.best_estimator_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_scores
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_selected_scores = feature_scores[features_selected_bool]
feature_selected_scores
imp_features_df = pd.DataFrame({'Features_Selected':features_selected_list, 'Features_score':feature_selected_scores})
imp_features_df.sort_values('Features_score', ascending = False,inplace = True)
Rank = pd.Series(list(range(1,len(features_selected_list)+1)))
imp_features_df.set_index(Rank, inplace = True)
imp_features_df
# ## Classifier 5 : KNN with PCA
# +
from sklearn.neighbors import KNeighborsClassifier
clf_knn = KNeighborsClassifier()
sss = StratifiedShuffleSplit(n_splits=10, test_size=0.3,random_state = 42)
pipeline = Pipeline(steps = [("scaling", scaler), ("SKB", skb), ("PCA",pca), ("knn",clf_knn)])
param_grid = {"SKB__k":[7,8,9,10,11,12,13,14,15, 16, 17, 18],
"PCA__n_components":[2,3,4,5,6,7],
"PCA__whiten":[True],
"knn__n_neighbors": [3,4,5,6,7,8,9,11],
#"knn__weights": ['uniform','distance'],
#"knn__algorithm": ['auto', 'ball_tree', 'kd_tree', 'brute']
}
grid = GridSearchCV(pipeline, param_grid, verbose = 0, cv = sss, scoring = 'f1')
t0 = time()
#clf = clf.fit(features_train, labels_train)
grid.fit(features, labels)
print ("training time: ", round(time()-t0, 3), "s")
# best algorithm
clf = grid.best_estimator_
t0 = time()
# refit the best algorithm:
clf.fit(features_train, labels_train)
prediction = clf.predict(features_test)
print ("testing time: ", round(time()-t0, 3), "s")
print ("Accuracy of DT classifer is : ",accuracy_score(labels_test, prediction))
print ("Precision of DT classifer is : ",precision_score(prediction, labels_test))
print ("Recall of DT classifer is : ",recall_score(prediction, labels_test))
print ("f1-score of DT classifer is : ",f1_score(prediction, labels_test))
# -
grid.best_params_
grid.best_estimator_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_scores
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_selected_scores = feature_scores[features_selected_bool]
feature_selected_scores
imp_features_df = pd.DataFrame({'Features_Selected':features_selected_list, 'Features_score':feature_selected_scores})
imp_features_df.sort_values('Features_score', ascending = False,inplace = True)
Rank = pd.Series(list(range(1,len(features_selected_list)+1)))
imp_features_df.set_index(Rank, inplace = True)
imp_features_df
# ## Classifier KNN without PCA
# +
from sklearn.neighbors import KNeighborsClassifier
clf_knn = KNeighborsClassifier()
sss = StratifiedShuffleSplit(n_splits=10, test_size=0.3,random_state = 42)
pipeline = Pipeline(steps = [("scaling", scaler), ("SKB", skb), ("knn",clf_knn)])
param_grid = {"SKB__k":[7,8,9,10,11,12,13,14,15, 16, 17, 18],
"knn__n_neighbors": [3,4,5,6,7,8,9,11,12,13,15],
}
grid = GridSearchCV(pipeline, param_grid, verbose = 0, cv = sss, scoring = 'f1')
t0 = time()
#clf = clf.fit(features_train, labels_train)
grid.fit(features, labels)
print ("training time: ", round(time()-t0, 3), "s")
# best algorithm
clf = grid.best_estimator_
t0 = time()
# refit the best algorithm:
clf.fit(features_train, labels_train)
prediction = clf.predict(features_test)
print ("testing time: ", round(time()-t0, 3), "s")
print ("Accuracy of DT classifer is : ",accuracy_score(labels_test, prediction))
print ("Precision of DT classifer is : ",precision_score(prediction, labels_test))
print ("Recall of DT classifer is : ",recall_score(prediction, labels_test))
print ("f1-score of DT classifer is : ",f1_score(prediction, labels_test))
# -
grid.best_params_
grid.best_estimator_
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_scores
features_selected_bool = grid.best_estimator_.named_steps['SKB'].get_support()
features_selected_list = [x for x,y in zip(features_list[1:], features_selected_bool) if y]
print ("Selected Features : \n",features_selected_list)
feature_scores = grid.best_estimator_.named_steps['SKB'].scores_
feature_selected_scores = feature_scores[features_selected_bool]
feature_selected_scores
imp_features_df = pd.DataFrame({'Features_Selected':features_selected_list, 'Features_score':feature_selected_scores})
imp_features_df.sort_values('Features_score', ascending = False,inplace = True)
Rank = pd.Series(list(range(1,len(features_selected_list)+1)))
imp_features_df.set_index(Rank, inplace = True)
imp_features_df
# + jupyter={"outputs_hidden": true}
# + jupyter={"outputs_hidden": true}
| __Project_analysis_jeswingeorge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
import bokeh.plotting as blt
import Cython
# %load_ext Cython
# %load_ext autoreload
# %autoreload 2
import causticTools.sim as cSim
import causticTools.io as cIo
import causticTools.analytic as cAnalytic
import pyximport
pyximport.install()
from causticTools.cythonSim import massFunction
M=1e9
m=9e8
rho=1
vFactor=(4/3)*np.pi
class baryonInit(massFunction):
def evaluate(self,r):
return M
class baryonMass(massFunction):
def evaluate(self,r):
return m
class dmMass(massFunction):
def evaluate(self,r):
return vFactor*rho*(r**3)
class findEcc(massFunction):
def evaluate(self,P):
return 1-(1-P)**(1/n)
#test cell
nShells=4096
nPhase=1
nEcc=1
T=1e7
dt=1e5
rMin=50
rMax=500
n=1 # defines eccentricity distribution
nOutput=8
name='Test'
cSim.runSim(nShells,nPhase,nEcc,T,dt,rMin,rMax,name,nOutput,dmMass(),baryonInit(),baryonMass(),findEcc())
| causticsSim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Requirements:
#
# Tested with opsef003.yml (see attached file)
# opsef002 + n2v = opsef003
#
# on a GeForce RTX 2080 with 8GB RAM
# on ubuntu/18.04.3
# ### adaped from:
#
# https://github.com/MouseLand/cellpose
#
# https://github.com/CellProfiler/CellProfiler
#
# https://github.com/mpicbg-csbd/stardist
#
# https://github.com/scikit-image/scikit-image
#
# https://github.com/VolkerH/unet-nuclei/
#
# Thanks to:
#
# All developer of the above mentioned repositories.
# +
# basic libs
import os
import sys
import time
import datetime
import inspect
from glob import glob
import tifffile as tif
import cv2 as cv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import collections
import math
import pickle
import networkx as nx
# %matplotlib inline
# for lif
import readlif
from readlif.reader import LifFile
# skimage
import skimage
from skimage import transform, io, filters, measure, morphology,img_as_float
from skimage.color import label2rgb,gray2rgb
from skimage.filters import gaussian, rank, threshold_otsu
from skimage.io import imread, imsave
from skimage.measure import label, regionprops, regionprops_table
from skimage.morphology import disk, watershed
# scipy
from scipy.signal import medfilt
from scipy.ndimage import generate_binary_structure, binary_dilation
# for cellpose
from cellpose import models as cp_models
from cellpose import utils as cp_utils
from cellpose import plot, transforms
from cellpose import plot, transforms
# other
import mxnet as mx
# for cluster analysis
from sklearn.datasets import load_iris
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.cluster import AgglomerativeClustering
# +
main_folder = os.path.dirname(os.path.abspath(inspect.stack()[0][1]))
import_path = os.path.join(main_folder,"Utils_and_Configs")
if import_path not in sys.path:
sys.path.append(import_path)
# import from import_path
from Tools_002 import *
from UNet_CP01 import *
from Segmentation_Func_06 import *
from Pre_Post_Process002 import *
from N2V_DataGeneratorTR001 import *
from opsef_core_002 import *
# +
# from https://github.com/mpicbg-csbd/stardist / 3_prediction (2D)
from __future__ import print_function, unicode_literals, absolute_import, division
# %config InlineBackend.figure_format = 'retina'
from csbdeep.utils import Path, normalize
from stardist import random_label_cmap, _draw_polygons
from stardist.models import StarDist2D
# other
import pkg_resources
import keras
# We import all our dependencies.
from n2v.models import N2VConfig, N2V
from n2v.utils.n2v_utils import manipulate_val_data
# from n2v.internals.N2V_DataGenerator2 import N2V_DataGenerator2
# -
# ## Load parameter
# the parameter for processing need to be defined in the notebook.
# Opsef_Setup_000X
# this notebook will print in the end a file_path.
# Please cut and paste it below!
# +
file_path = "./Demo_Notebooks/my_runs/Parameter_muscle_mask_Run_000.pkl"
infile = open(file_path,'rb')
parameter = pickle.load(infile)
print("Loading processing pipeline from",file_path)
infile.close()
pc,input_def,run_def,initModelSettings = parameter
# -
# def rewrite_fiji_tiff(input_d):
# search_path = os.path.join(input_d["root"],"tiff_fiji")
# tiff_to_split = glob("{}/*tif".format(search_path))
# for file in tiff_to_split:
# print(file)
# fn = os.path.split(file)[1]
# img = tif.imread(file)
# print(img.shape)
# img_new = np.swapaxes(img.copy(),0,2)
# print(img_new.shape)
# tif.imsave(os.path.join(input_d["root"],"tiff",fn),img_new)
# return
# input_def["rearrange_ch_from_fiji"] = True
# if input_def["rearrange_ch_from_fiji"]:
# rewrite_fiji_tiff(input_def)
# ## Process Images
# +
# process for all
# create subfolder
make_folder_structure(pc,input_def,run_def)
# process for lif
if input_def["input_type"] == ".lif":
lifobject,input_def = define_lif_pipeline(input_def)
preprocess_1_for_lif(lifobject,input_def,pc,run_def)
preprocess_2_for_lif(lifobject,input_def,pc,run_def)
# process for tif
if input_def["input_type"] == ".tif":
fpath_list = define_tif_pipeline(input_def)
if pc["export_another_channel"]: # implement cleaner
fpath_list = [f for f in fpath_list if input_def["export_seg_ch"] in f]
preprocess_1_for_tif(fpath_list,input_def,pc,run_def)
preprocess_2_for_tif(fpath_list,input_def,pc,run_def)
# Segment
start_time = datetime.datetime.now()
segment(input_def,pc,run_def,initModelSettings)
end_time = datetime.datetime.now()
time_delta = end_time - start_time
print("The segmentatio took overall:", time_delta)
# -
# ## Export annditional channel & Quantify Results
if pc["Export_to_CSV"]:
all_combined = [] # used for quantifications of more than one intensity channel
# get a list of the masks that were produced by segmentation
mask_files = glob(os.path.join(input_def["root"],"Processed_{}".format(run_def["run_ID"]),pc["sub_f"][2])+"/*.tif")
mask_to_img_dic, mask_to_8bitimg_dic = make_mask_to_img_dic(mask_files,pc,input_def,run_def,0,pc["Intensity_Ch"])
if pc["toFiji"]:
if not pc["Export_to_CSV"]:
mask_files = glob(os.path.join(input_def["root"],"Processed_{}".format(run_def["run_ID"]),pc["sub_f"][2])+"/*.tif")
mask_to_img_dic, mask_to_8bitimg_dic = make_mask_to_img_dic(mask_files,pc,input_def,run_def,0,pc["Intensity_Ch"])
root_plus = os.path.join(input_def["root"],"Processed_{}".format(run_def["run_ID"]))
txt_fn = os.path.join(root_plus,pc["sub_f"][10],"FilePairList_{}_{}.txt".format(input_def["dataset"],run_def["run_ID"]))
with open(txt_fn,"w") as f:
for mask_fn,image_fn in mask_to_8bitimg_dic.items():
f.write("{};{}{}".format(image_fn.replace(root_plus,""),mask_fn.replace(root_plus,""),"\n"))
f.close()
# export additional channel
if pc["export_another_channel"]:
if input_def["input_type"] == ".lif":
exported_file_list = export_second_channel_for_mask(lifobject,pc,input_def,run_def)
if input_def["input_type"] == ".tif":
exported_file_list = export_second_channel_for_mask("NoneIsTiFF",pc,input_def,run_def)
# optional in case segmentation results shall be filtered by a mask:
if pc["create_filter_mask_from_channel"]:
# create new masks (by thresolding the additional input) and extract their names
new_mask_fn_list = create_mask_from_add_ch(exported_file_list,input_def["root"],pc["sub_f"],run_def["run_ID"],run_def["para_mp"],run_def)
# make a dic that has the segmentation output mask name as key, the name of the threshold mask as value
if input_def["input_type"] == ".lif":
pair_dic = make_pair_second_mask_simple(mask_files,new_mask_fn_list)
if input_def["input_type"] == ".tif":
core_match = [8,10] # use to define how to match filenames
# for documentation see: how_to_define_core_match.txt
# integrate this variable in OpSeF_Setup!!!
pair_dic = make_pair_second_mask_tiff(mask_files,new_mask_fn_list,core_match)
# create new seqmentation masks per class and return a list of file_names
class1_to_img_dic,class2_to_img_dic = split_by_mask(input_def["root"],run_def["run_ID"],pc["sub_f"],pair_dic,mask_to_8bitimg_dic,mask_to_img_dic)
# +
# print(mask_files)
# -
if pc["toFiji"]:
if pc["create_filter_mask_from_channel"]:
root_plus = os.path.join(input_def["root"],"Processed_{}".format(run_def["run_ID"]))
txt_fn = os.path.join(root_plus,pc["sub_f"][10],"FilePairList_Classes_{}_{}.txt".format(input_def["dataset"],run_def["run_ID"]))
img_to_class2_dic = dict((v,k) for k,v in class2_to_img_dic.items()) # invert dic 2
with open(txt_fn,"w") as f:
for mask_fn,image_fn in class1_to_img_dic.items():
mask2 = img_to_class2_dic[image_fn] # second seg mask
f.write("{};{};{};{}".format(image_fn.replace(root_plus,""),mask_fn.replace(root_plus,""),mask2.replace(root_plus,""),"\n"))
f.close()
# ## Export results
# +
# quantify original mask
if pc["Export_to_CSV"]:
all_combined.append(results_to_csv(mask_to_img_dic,pc["get_property"],input_def["root"],pc["sub_f"],run_def["run_ID"],4,"All_Main",input_def["subset"])) # 4 is the main result folder
if pc["plot_head_main"]:
all_combined[0].head()
if pc["create_filter_mask_from_channel"]:
# quantify class1 masks
results_to_csv(class1_to_img_dic,pc["get_property"],input_def["root"],pc["sub_f"],run_def["run_ID"],9,"Class00",input_def["post_subset"]) # 9 is the classified result folder
# quantify class2 masks
results_to_csv(class2_to_img_dic,pc["get_property"],input_def["root"],pc["sub_f"],run_def["run_ID"],9,"Class01",input_def["post_subset"]) # 9 is the classified result folder
# -
if pc["Quantify_2ndCh"]:
mask_to_img_dic, mask_to_8bitimg_dic = make_mask_to_img_dic(mask_files,pc,input_def,run_def,5,pc["Intensity_2ndCh"])
all_combined.append(results_to_csv(mask_to_img_dic,pc["get_property"],input_def["root"],pc["sub_f"],run_def["run_ID"],4,"All_2nd",input_def["subset"]))
if pc["merge_results"]:
result_summary = merge_intensity_results(all_combined,input_def,pc["sub_f"],run_def,4)
if pc["plot_merged"]:
result_summary.head()
else:
if pc["Export_to_CSV"]:
result_summary = all_combined[0]
# ## AddOn 1: Basic plotting of results
if pc["Plot_Results"]:
fig, axs = plt.subplots(len(pc["Plot_xy"]), 1, figsize=(5, 5*len(pc["Plot_xy"])), constrained_layout=True)
for i in range(0,len(pc["Plot_xy"])):
axs[i].scatter(result_summary[pc["Plot_xy"][i][0]],result_summary[pc["Plot_xy"][i][1]], c="red")
axs[i].set_title('{} vs {}'.format(*pc["Plot_xy"][i]))
axs[i].set_xlabel(pc["Plot_xy"][i][0],fontsize=15)
axs[i].set_ylabel(pc["Plot_xy"][i][1],fontsize=15)
# ## AddOn 2: Do PCA and TSNE
# ### Example pipeline auto-clustering
if pc["Cluster_How"] == "Auto":
# get data for PCA / TSNE
df_for_tsne_list = extract_values_for_TSNE_PCA(input_def["root"],run_def["run_ID"],pc["sub_f"],4,pc["include_in_tsne"])
# get cluster
data = df_for_tsne_list[0].values
auto_clustering = AgglomerativeClustering(linkage=pc["link_method"], n_clusters=pc["cluster_expected"]).fit(data)
# do analysis
result_tsne = TSNE(learning_rate=pc["tSNE_learning_rate"]).fit_transform(data)
result_pca = PCA().fit_transform(data)
# display results
fig, axs = plt.subplots(2, 1, figsize=(10, 20), constrained_layout=True)
axs[0].scatter(result_tsne[:, 0], result_tsne[:, 1], c=auto_clustering.labels_)
axs[0].set_title('tSNE')
axs[1].scatter(result_pca[:, 0], result_pca[:, 1], c=auto_clustering.labels_)
axs[1].set_title('PCA')
# ### Example pipeline mask-clustering
# get data for PCA / TSNE
if pc["Cluster_How"] == "Mask":
df_for_tsne_list_by_class = extract_values_for_TSNE_PCA(input_def["root"],run_def["run_ID"],pc["sub_f"],9,pc["include_in_tsne"])
fused_df = pd.concat(df_for_tsne_list_by_class,axis = 0,join="outer")
data_by_class = fused_df.values
class_def_by_mask = [0 for x in range (0,df_for_tsne_list_by_class[0].shape[0])] + [1 for x in range (0,df_for_tsne_list_by_class[1].shape[0])]
# do analysis
result_tsne_by_class = TSNE(learning_rate=pc["tSNE_learning_rate"]).fit_transform(data_by_class)
result_pca_by_class = PCA().fit_transform(data_by_class)
# display results
fig, axs = plt.subplots(2, 1, figsize=(10, 20), constrained_layout=True)
axs[0].scatter(result_tsne_by_class[:, 0], result_tsne_by_class[:, 1], c=class_def_by_mask)
axs[0].set_title('tSNE')
axs[1].scatter(result_pca_by_class[:, 0], result_pca_by_class[:, 1], c=class_def_by_mask)
axs[1].set_title('PCA')
# ## Results
print("Processing completed sucessfully !\n")
print("All results have been saved in this folder: \n")
print(os.path.join(input_def["root"],"Processed_{}".format(run_def["run_ID"])))
| .ipynb_checkpoints/OpSeF_IV_Run_002b-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ZsQ5PDPK01ZB" executionInfo={"status": "ok", "timestamp": 1639677277053, "user_tz": -330, "elapsed": 3054, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>w=s64", "userId": "14345249188510044799"}}
import os
import cv2
import numpy as np
from tqdm import tqdm
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Dense, Input, GlobalAveragePooling2D, Flatten
from tensorflow.keras.applications import DenseNet121
from tensorflow.keras.models import Model
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.callbacks import ModelCheckpoint, ReduceLROnPlateau, EarlyStopping
from sklearn.model_selection import train_test_split
from IPython.display import clear_output
import random
import shutil
# + id="Wnr7Zg4u37jO" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1639677829845, "user_tz": -330, "elapsed": 552805, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="3fbd8067-7933-4900-a695-13261eec3cc1"
from google.colab import drive
drive.mount("drive")
# + id="muxpUKgQ4YkT" executionInfo={"status": "ok", "timestamp": 1639677843160, "user_tz": -330, "elapsed": 13333, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
# !unzip drive/MyDrive/Datasets/FINAL_COVID_DATASET.zip
clear_output()
# + id="Iv3Ghij1ezeG" executionInfo={"status": "ok", "timestamp": 1639677843161, "user_tz": -330, "elapsed": 22, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
def split_data(data_path):
os.mkdir("dataset")
os.mkdir("dataset/train")
os.mkdir("dataset/test")
os.mkdir("dataset/val")
for i in os.listdir(data_path):
os.mkdir(f"dataset/train/{i}")
os.mkdir(f"dataset/test/{i}")
os.mkdir(f"dataset/val/{i}")
cat_path = os.path.join(data_path,i)
filenames = os.listdir(cat_path)
random.seed(2021)
random.shuffle(filenames)
train, test = train_test_split(filenames, test_size=0.25)
test, val = train_test_split(test, test_size=0.5)
for file in tqdm(train):
shutil.copy(f"{cat_path}/{file}", f"dataset/train/{i}/{file}")
for file in tqdm(test):
shutil.copy(f"{cat_path}/{file}", f"dataset/test/{i}/{file}")
for file in tqdm(val):
shutil.copy(f"{cat_path}/{file}", f"dataset/val/{i}/{file}")
# + colab={"base_uri": "https://localhost:8080/"} id="5pS-8QJtezZO" executionInfo={"status": "ok", "timestamp": 1639677846159, "user_tz": -330, "elapsed": 3017, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="941d409f-3c19-4043-bc8a-f80eca97e5ca"
split_data("FINAL_COVID_DATASET")
# + colab={"base_uri": "https://localhost:8080/"} id="50TNAAwRezVA" executionInfo={"status": "ok", "timestamp": 1639677846160, "user_tz": -330, "elapsed": 11, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="07a0186f-7f76-4cb4-ef0f-84e159a8bc00"
len(os.listdir("dataset/train/COVID"))
# + id="223K6tHR48vB" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1639677846983, "user_tz": -330, "elapsed": 829, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="6db3f127-4d74-4de1-f4a9-ccaf27c3d0d2"
IMAGE_SIZE = 224
BATCH_SIZE = 64
generator = ImageDataGenerator(rescale=1./255)
X_train = generator.flow_from_directory(
"dataset/train",
target_size = (IMAGE_SIZE, IMAGE_SIZE),
batch_size = BATCH_SIZE
)
X_test = generator.flow_from_directory(
"dataset/test",
target_size = (IMAGE_SIZE, IMAGE_SIZE),
batch_size = BATCH_SIZE
)
X_val = generator.flow_from_directory(
"dataset/val",
target_size = (IMAGE_SIZE, IMAGE_SIZE),
batch_size = BATCH_SIZE
)
print("Train:",X_train.class_indices)
print("Test:",X_test.class_indices)
print("Val:",X_val.class_indices)
# + colab={"base_uri": "https://localhost:8080/"} id="p4UFYWYxBTpK" executionInfo={"status": "ok", "timestamp": 1639677855189, "user_tz": -330, "elapsed": 8214, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="4aa0dbad-95e3-4bf6-f50e-12f3b20d7e25"
base_model = DenseNet121(
input_shape=(IMAGE_SIZE,IMAGE_SIZE,3),
include_top=False,
weights='imagenet',
)
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dense(1024, activation="relu")(x)
x = Dense(512, activation="relu")(x)
x = Dense(3, activation="softmax")(x)
# + colab={"base_uri": "https://localhost:8080/"} id="jBVdbmzaDl-l" executionInfo={"status": "ok", "timestamp": 1639677856101, "user_tz": -330, "elapsed": 922, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="ec97eee0-d6e1-48b7-fbca-4175a58ec7f1"
model = Model(inputs=base_model.input, outputs=x)
for layer in model.layers[:30]:
layer.trainable = False
for layer in model.layers[30:]:
layer.trainable = True
model.summary()
# + id="j4B-zWhlKRb0" executionInfo={"status": "ok", "timestamp": 1639677856102, "user_tz": -330, "elapsed": 58, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + id="JeijJBTQHWxq" executionInfo={"status": "ok", "timestamp": 1639677856103, "user_tz": -330, "elapsed": 59, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
checkpoint_cb = ModelCheckpoint(
"checkpoint_model.h5",
monitor='val_loss',
save_best_only=True,
verbose=1,
mode='min'
)
early_stopping_cb = EarlyStopping(
monitor='val_loss',
patience=4,
verbose=1,
restore_best_weights=True,
mode='min'
)
reduce_on_plateau_cb = ReduceLROnPlateau(
monitor='val_loss',
mode='min',
verbose=1,
patience=2,
factor=0.5,
min_lr=1e-6
)
# + colab={"base_uri": "https://localhost:8080/"} id="lAJWfDwOJqD3" executionInfo={"status": "ok", "timestamp": 1639680157219, "user_tz": -330, "elapsed": 2301174, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="67880895-325f-45ed-e294-e61aff1ed3a0"
with tf.device("/device:GPU:0"):
history = model.fit_generator(
X_train,
validation_data=X_val,
epochs=50,
callbacks=[checkpoint_cb, early_stopping_cb, reduce_on_plateau_cb]
)
# + id="lH0K5iefPUnB" colab={"base_uri": "https://localhost:8080/", "height": 621} executionInfo={"status": "ok", "timestamp": 1639680208852, "user_tz": -330, "elapsed": 1131, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="1fd08514-9665-4c73-e434-078402df8775"
plt.figure(figsize=(10,10))
for i, met in enumerate(['accuracy', 'loss']):
plt.subplot(2,1,i+1)
plt.plot(history.history[met], color="b")
plt.plot(history.history["val_"+met], color="g")
plt.title('Model '+met.capitalize())
plt.xlabel('epochs')
plt.ylabel(met)
plt.legend(['train', 'val'])
# plt.axvline(x=6, label=f"line at x = {6}", c='r',ls="--", lw=1)
# + colab={"base_uri": "https://localhost:8080/"} id="f-BwenuYnsst" executionInfo={"status": "ok", "timestamp": 1639680270267, "user_tz": -330, "elapsed": 13875, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="f98b62ac-bb3d-484a-85ce-c5b28eb704a2"
model.evaluate(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="Xdmxfse5nwVQ" executionInfo={"status": "ok", "timestamp": 1639680354680, "user_tz": -330, "elapsed": 27502, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}} outputId="b0f27fdf-15fd-43aa-90fc-a8f4a3f03aa8"
cp_model = tf.keras.models.load_model("checkpoint_model.h5")
cp_model.evaluate(X_test)
# + id="_fAvutxwD6de" executionInfo={"status": "ok", "timestamp": 1639680356249, "user_tz": -330, "elapsed": 1582, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
model.save("model.h5")
# + id="aXr385-Jtff2" executionInfo={"status": "ok", "timestamp": 1639680397822, "user_tz": -330, "elapsed": 536, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
# !cp model.h5 drive/MyDrive/Covid-Pneumonia_Detection_Model/Transfer_Learning/densenet121.h5
# + id="1hDJnDY3xg8U" executionInfo={"status": "ok", "timestamp": 1639680432125, "user_tz": -330, "elapsed": 799, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
import pickle
with open('DenseNet121trainHistory', 'wb') as file_pi:
pickle.dump(history.history, file_pi)
# To load history
# history = pickle.load(open('DenseNet121trainHistory', "rb"))
# + id="HtxsutIi0yES" executionInfo={"status": "ok", "timestamp": 1639680452282, "user_tz": -330, "elapsed": 785, "user": {"displayName": "1DS19IS027 <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhNw_C5vXvbrerf4BPWr_TB0trE_XAd8B_crxCH8w=s64", "userId": "14345249188510044799"}}
# !cp DenseNet121trainHistory drive/MyDrive/Covid-Pneumonia_Detection_Model/Transfer_Learning/DenseNet121trainHistory
# + id="lYIxHLI-1QdR"
| notebooks/Covid_Pneumonia Classification DenseNet121.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + slideshow={"slide_type": "skip"}
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from numpy import *
from IPython.html.widgets import *
from IPython.display import display
import matplotlib.pyplot as plt
from IPython.core.display import clear_output
# + [markdown] slideshow={"slide_type": "skip"}
# <style>
# .center-image { display: block; margin: auto; }
# </style>
# + [markdown] slideshow={"slide_type": "slide"}
# # Perceptron
#
# In this notebook, I will go through how to train a perceptron for binary classification problems.
#
# <img src="files/images/Perceptron/classification.png" class="image-center" style="width: 330px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is a perceptron?
#
# Perceptron is a artificial neural network whose learning was invented by <NAME> in 1957.
#
# <img src="files/images/Perceptron/rosenblatt.jpg" class="image-center" style="width: 225px;"/>
#
# According to wikipedia, "In a 1958 press conference organized by the US Navy, Rosenblatt made statements about the perceptron that caused a heated controversy among the fledgling AI community; based on Rosenblatt's statements, The New York Times reported the perceptron to be "the embryo of an electronic computer that [the Navy] expects will be able to walk, talk, see, write, reproduce itself and be conscious of its existence."
# + [markdown] slideshow={"slide_type": "subslide"}
# A perceptron is a single-layer, linear classifier:
#
# <img src="files/images/Perceptron/diagram.png" class="image-center" style="width: 817px;"/>
#
# Although it is very simple (and too simple for many tasks), it forms the basis for more sophisticated networks and algorithms (backpropagation).
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="files/images/Perceptron/diagram.png" class="image-center" style="width: 817px;"/>
#
# A perceptron has $P$ input units, one output unit and $P+1$ weights (parameters) $w_n$. For a particular input (a $P$-dimensional vector ${\bf x}$), the perceptron outputs
#
# $$ t = sign( a ) = sign( {\bf x} {\bf w}^\intercal + {\bf w}_0 ) $$
#
# $a$ is the *activation* of the perceptrion. The "sign" function returns $+1$ for anything greater than zero and $-1$ for less than zero.
# + [markdown] slideshow={"slide_type": "fragment"}
# ${\bf w}_0$ is called the "bias weight". For convenience, we often change the input vector ${\bf x} = (x_1, x_2, ...)$ to have $1$ at the end (${\bf x}' = (x_1, x_2, ..., x_p, 1)$), so that we can express the activation as simply $a = {\bf x}' {\bf w}^\intercal$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## How is a perceptron trained?
#
# In binary classification problems, for each sample ${\bf x}_i$, we have a corresponding *label* $y_i \in \{ 1, -1 \}$. $1$ corresponds to one class (red points, for example), and $-1$ corresponds to the other class (blue points).
#
# <img src="files/images/Perceptron/classification.png" class="image-center" style="width: 330px;"/>
#
# By "training", I mean that I want to find ${\bf w}$ such that for all $i$, $y_i = sign({\bf x}_i' {\bf w}^\intercal)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# In other words, I want to minimize the (0/1) *loss function*
#
# $$ J_{0/1}({\bf w}) = \frac{1}{N} \sum_{i=1}^{N} (y_i == sign({\bf x}_i' {\bf w}^\intercal))$$
#
# This is an optimization problem. Unfortunately, this particular loss function is practically impossible to solve, because the gradient is flat everywhere!
#
# <img src="files/images/Perceptron/0-1loss.png" class="image-center" style="width: 330px;"/>
# + [markdown] slideshow={"slide_type": "subslide"}
# Instead, a perceptron learning rule minimizes the *perceptron criterion*:
#
# $$ J({\bf w}) = \frac{1}{N} \sum_{i=1}^{N} \max (0, - y_i a_i)$$
#
# * If the prediction was correct - say, $y_i = 1$ and $a = 0.8$, then $-y_i a_i < 0$, so $\max(0, -y_i a_i) = 0$. In other words, the loss is zero for correct examples.
# * If the prediction was wrong - say, $y_i = -1$ and $a = 0.8$, then the loss is *proportional* to $a_i$. The penalty is very large when you predict very large $a_i$ and get it wrong!
#
# It's important to note that the loss function only cares about the examples that were classified wrong.
# + [markdown] slideshow={"slide_type": "subslide"}
# So we can also rewrite the loss function as
#
# $$ J({\bf w}) = \sum_{i= \textrm{Wrong samples}} - y_i a_i $$
#
# Now we can take the derivative with respect to ${\bf w}$ and get something nicer:
#
# $$ \frac{\partial J_i({\bf w})}{\partial w_j} = -y_i x_{ij} \quad \textrm{For wrong sample } i$$
#
# <img src="files/images/Perceptron/perceptron_loss.png" class="image-center" style="width: 330px;"/>
# + [markdown] slideshow={"slide_type": "subslide"}
# And, using the stochastic gradient decent algorithm with the learning rate of $\eta$, we get the perceptron weight update rule:
#
# $${\bf w} \leftarrow {\bf w} + \eta y_i {\bf x}_i' $$
#
# ... for all misclassified examples $i$.
#
# (Exercise: take out your papers and pencils and convince yourself of this.)
# + [markdown] slideshow={"slide_type": "slide"}
# # Training a perceptron
#
# Let's generate some data `X` and binary labels `y` to test on! We'll use `sklearn.datasets` to make some test data. Note that we want the targets `y` to be `{-1, 1}`.
# + slideshow={"slide_type": "subslide"}
from sklearn.datasets import make_blobs
X = y = None # Global variables
@interact
def plot_blobs(n_samples=(10, 500),
center1_x=1.5,
center1_y=1.5,
center2_x=-1.5,
center2_y=-1.5):
centers=array([[center1_x, center1_y],[center2_x, center2_y]])
global X, y
X, y= make_blobs(n_samples=n_samples, n_features=2,
centers=centers, cluster_std=1.0)
y = y*2 - 1 # To convert to {-1, 1}
plt.scatter(X[:,0], X[:,1], c=y, edgecolor='none')
plt.xlim([-10,10]); plt.ylim([-10,10]); plt.grid()
plt.axes().set_aspect('equal')
# + slideshow={"slide_type": "skip"}
from sklearn.cross_validation import train_test_split
# Plotting routine for perceptron training
def predict(w, X):
"""Returns the predictions."""
return sign(dot(c_[X, ones((X.shape[0], 1))], w))
def error01(w, X, y):
"""Calculates the mean 0/1 error."""
return 1.0 - (predict(w, X) == y).mean()
def perceptron_training(X,y,eta=0.1):
global w, errors
# Split data to training and test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
# Plot the current predictions and the hyperplane
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
axs[0].scatter(X_train[:,0], X_train[:,1], c=predict(w, X_train), edgecolor='none')
axs[0].set_xlim([-10,10]); axs[0].set_ylim([-10,10]); axs[0].grid()
axs[0].set_aspect('equal')
# Draw the separating line
cw=-w[2]/(w[0]**2+w[1]**2)
ts=array([-100.0,100.0])
axs[0].plot(-w[1]*ts+w[0]*cw, w[0]*ts+w[1]*cw, linestyle='--', color='r')
axs[0].arrow(w[0]*cw,w[1]*cw, w[0], w[1],
head_width=0.5, head_length=0.5, fc='r', ec='r')
# Plot the classification errors
train_error, test_error = [error01(w, X_, y_) for X_, y_ in [[X_train, y_train], [X_test, y_test]]]
errors = r_[errors, array([train_error, test_error])[newaxis,:]]
axs[1].plot(errors)
axs[1].set_title('Classification Errors')
axs[1].set_ylim([0,1])
axs[1].legend(['Training','Test'])
# Update w
w = update_w_all(w, X_train, y_train, eta)
# + [markdown] slideshow={"slide_type": "slide"}
# Exercise 2. implement the code to do a single step of perceptron weight update. Press the button each time to run a single step of `update_w_all`; re-evaluating the cell resets the weight and starts over.
#
# Remember:
#
# $${\bf w} \leftarrow {\bf w} + \eta y_i {\bf x}_i' $$
#
# Note that the following code does random train-test split everytime `w` is updated.
# + slideshow={"slide_type": "subslide"}
def delta_w_single(w, x, y):
"""Calculates the gradient for w from the single sample x and the target y.
inputs:
w: 1 x (p+1) vector of current weights.
x: 1 x p vector representing the single sample.
y: the target, -1 or 1.
returns:
w: 1 x (p+1) vector of updated weights.
"""
# TODO implement this
return 0
def update_w_all(w, X, y, eta = 0.1):
"""Updates the weight vector for all training examples.
inputs:
w: 1 x (p+1) vector of current weights.
X: N x p vector representing the single sample.
y: N x 1 vector of the targets, -1 or 1.
eta: The training rate. Defaults to 0.1.
returns:
w: 1 x (p+1) vector of updated weights.
"""
for xi, yi in zip(X, y):
w += eta * delta_w_single(w, xi, yi) / X.shape[0]
return w
from numpy.random import random_sample
w = random_sample(3) # Initialize w to values from [0, 1)
errors = zeros((0, 2)) # Keeps track of error values over time
interact_manual(perceptron_training, X=fixed(X), y=fixed(y), eta=FloatSlider(min=0.01, max=1.0, value=0.1))
# + slideshow={"slide_type": "skip"}
# A sample solution:
def delta_w_single(w, x, y):
"""Updates the weight vector w from the single sample x and the target y.
inputs:
w: 1 x (p+1) vector of current weights.
x: 1 x p vector representing the single sample.
y: the target, -1 or 1.
returns:
w: 1 x (p+1) vector of updated weights.
"""
x_prime = r_[x, 1]
prediction = sign(dot(x_prime, w.T))
if prediction != y:
return y * x_prime
else:
return 0
# + [markdown] slideshow={"slide_type": "slide"}
# The perceptron only works for linearly separable data.
# + slideshow={"slide_type": "subslide"}
from sklearn.datasets import make_circles, make_moons
X_circle, y_circle = make_circles(100) # or try make_moons(100)
y_circle = y_circle * 2 - 1
X_circle*=4 # Make it a bit larger
plt.scatter(X_circle[:,0], X_circle[:,1], c=y_circle, edgecolor='none')
plt.xlim([-10,10]); plt.ylim([-10,10]); plt.grid()
plt.axes().set_aspect('equal')
# + slideshow={"slide_type": "subslide"}
w = random_sample(3) # Initialize w to values from [0, 1)
errors = zeros((0, 2)) # Keeps track of error values over time
interact_manual(perceptron_training, X=fixed(X_circle), y=fixed(y_circle), eta=fixed(0.1))
# -
# As we would have expected, the perceptron fails to converge for this problem.
| notebooks/Perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from firecloud import fiss
from firecloud import api as fapi
workspace = fapi.get_workspace("broad-firecloud-dsde-methods", "MMRF_EndToEndHardClippedUseDefaultAF")
workspace_attributes = workspace.json()
workspace_attributes
workspace_attributes["workspace"]["attributes"]["reference"]
method_configurations = fapi.get_method_configurations("LiquidBiopsyDevelopment", "BenchmarkLiquidBiopsy")
method_configurations
workflow_outputs = fapi.get_workflow_outputs("broad-firecloud-dsde-methods", "MMRF_EndToEndHardClippedUseDefaultAF", "30c74fe6-f594-4e4b-9dc1-cbd1458d3d27", "00eb253a-a18c-41ac-80db-b125e2a661e7")
workflow_outputs.json()
| fireplace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
'''
Created on Mar 18, 2016
@author: pushkar
'''
import urllib2
from bs4 import BeautifulSoup
import csv
import time
import re
def getAmazonDetails(isbn):
with open('csv_files/amazon_book_ratings.csv', 'a') as csvfile_ratings, open('csv_files/amazon_book_reviews.csv', 'a') as csvfile_reviews:
##Create file headers and writer
ratings_fieldnames = ['book_isbn', 'avg_rating', 'five_rating', 'four_rating', 'three_rating', 'two_rating', 'one_rating' ]
writer = csv.DictWriter(csvfile_ratings, delimiter=',', lineterminator='\n', fieldnames=ratings_fieldnames)
##writer.writeheader()
reviews_fieldnames = ['book_isbn', 'review']
writer_book = csv.DictWriter(csvfile_reviews, delimiter=',', lineterminator='\n', fieldnames=reviews_fieldnames)
##writer_book.writeheader()
##Get Overall details of the book
req = urllib2.Request('http://www.amazon.com/product-reviews/' + isbn + '?ie=UTF8&showViewpoints=1&sortBy=helpful&pageNumber=1', headers={ 'User-Agent': 'Mozilla/5.0 (X11; U; Linux i686) Gecko/20071127 Firefox/2.0.0.11' })
html = urllib2.urlopen(req).read()
soup = BeautifulSoup(html, 'html.parser')
avgRatingTemp = soup.find_all('div',{'class':"a-row averageStarRatingNumerical"})[0].get_text()
avgRating = re.findall('\d+\.\d+', avgRatingTemp)[0]
try:
fiveStarRatingTemp = soup.find_all('a',{'class':"a-size-small a-link-normal 5star histogram-review-count"})[0].get_text()
fiveStarRating = fiveStarRatingTemp.strip('%')
except:
fiveStarRating = 0
try:
fourStarRatingTemp = soup.find_all('a',{'class':"a-size-small a-link-normal 4star histogram-review-count"})[0].get_text()
fourStarRating = fourStarRatingTemp.strip('%')
except:
fourStarRating = 0
try:
threeStarRatingTemp = soup.find_all('a',{'class':"a-size-small a-link-normal 3star histogram-review-count"})[0].get_text()
threeStarRating = threeStarRatingTemp.strip('%')
except:
threeStarRating = 0
try:
twoStarRatingTemp = soup.find_all('a',{'class':"a-size-small a-link-normal 2star histogram-review-count"})[0].get_text()
twoStarRating = twoStarRatingTemp.strip('%')
except:
twoStarRating = 0
try:
oneStarRatingTemp = soup.find_all('a',{'class':"a-size-small a-link-normal 1star histogram-review-count"})[0].get_text()
oneStarRating = oneStarRatingTemp.strip('%')
except:
oneStarRating = 0
writer.writerow({'book_isbn': isbn, 'avg_rating': avgRating, 'five_rating': fiveStarRating,
'four_rating': fourStarRating, 'three_rating': threeStarRating, 'two_rating': twoStarRating,
'one_rating': oneStarRating})
##Get top 20 helpful review of book
for pagenumber in range(1,3):
req = urllib2.Request('http://www.amazon.com/product-reviews/' + isbn + '?ie=UTF8&showViewpoints=1&sortBy=helpful&pageNumber='+ str(pagenumber), headers={ 'User-Agent': 'Mozilla/5.0 (X11; U; Linux i686) Gecko/20071127 Firefox/2.0.0.11' })
html = urllib2.urlopen(req).read()
soup = BeautifulSoup(html, 'html.parser')
for i in range(0,10):
try:
review = soup.find_all('div',{'class':"a-section review"})[i].contents[3].get_text().encode('UTF-8')
#print review
writer_book.writerow({'book_isbn': isbn, 'review': review})
except:
print "No Reviews ISBN - " + isbn
#getAmazonDetails('0940650703')
# +
import urllib2
import csv
import time
import sys
import xml.etree.ElementTree as ET
import os
import traceback
import random
from IPython.display import clear_output
def getval(root, element):
try:
ret = root.find(element).text
if ret is None:
return ""
else:
return ret.encode("utf8")
except:
return ""
with open('csv_files/amazon_book_ratings.csv', 'w') as csvfile_ratings, open('csv_files/amazon_book_reviews.csv', 'w') as csvfile_reviews:
##Create file headers and writer
ratings_fieldnames = ['book_isbn', 'avg_rating', 'five_rating', 'four_rating', 'three_rating', 'two_rating', 'one_rating' ]
writer = csv.DictWriter(csvfile_ratings, delimiter=',', lineterminator='\n', fieldnames=ratings_fieldnames)
writer.writeheader()
reviews_fieldnames = ['book_isbn', 'review']
writer_book = csv.DictWriter(csvfile_reviews, delimiter=',', lineterminator='\n', fieldnames=reviews_fieldnames)
writer_book.writeheader()
with open('csv_files/user_data.csv', 'w') as csvfile, open('csv_files/book_data.csv', 'w') as csvfile_book, open('csv_files/book_author.csv', 'w') as csvfile_author:
fieldnames = ['id', 'name','user_name', 'profile_url','image_url', 'about', 'age', 'gender',
'location','joined','last_active' ]
writer = csv.DictWriter(csvfile, delimiter = ',', lineterminator = '\n', fieldnames=fieldnames)
writer.writeheader()
book_fieldnames = [
'user_id',
'b_id',
'shelf',
'isbn',
'isbn13',
'text_reviews_count',
'title',
'image_url',
'link',
'num_pages',
'b_format',
'publisher',
'publication_day',
'publication_year',
'publication_month',
'average_rating',
'ratings_count',
'description',
'published',
'children',
'religion',
'history',
'math',
'anatology',
'poetry',
'encyclopedia',
'dictionaries',
'comics',
'art',
'cookbook',
'diaries',
'journals',
'prayer_books',
'series',
'trilogy',
'biographies',
'autobiographies',
'fantasy',
'comic',
'self-help',
'science-fiction',
'non-fiction']
writer_book = csv.DictWriter(csvfile_book, delimiter = ',', lineterminator = '\n', fieldnames=book_fieldnames)
writer_book.writeheader()
author_fieldnames = [
'u_id',
'b_id',
'a_id',
'name',
'average_rating',
'ratings_count',
'text_reviews_count']
writer_author = csv.DictWriter(csvfile_author, delimiter = ',', lineterminator = '\n', fieldnames = author_fieldnames)
writer_author.writeheader()
lst = []
i = 0
while i < 1000:
try:
#clear_output()
#time.sleep(1)
clear_output()
c = random.randint(5000000, 5625000) #7500000 625000
print "random number: " + str(c)
if (c not in lst):
print "getting information for user id:"+ str(c)
lst.append(c)
url = 'https://www.goodreads.com/user/show/'+ str(c) +'.xml?key=i3Zsl7r13oHEQCjv1vXw'
response = urllib2.urlopen(url)
user_data_xml = response.read()
#write xml to file
print "User number:" + str(i)
i = i + 1
f = open("xml_docs/user"+ str(c) +".xml", "w")
try:
f.write(user_data_xml)
finally:
f.close()
#root = ET.fromstring()
root = ET.parse("xml_docs/user"+ str(c) +".xml").getroot()
os.remove("xml_docs/user"+ str(c) +".xml")
user_element = root.find('user')
id = getval(user_element,'id')
name = getval(user_element,'name')
user_name = getval(user_element,'user_name')
profile_url = getval(user_element,'link')
image_url = getval(user_element,'image_url')
about = getval(user_element,'about')
age = getval(user_element,'age')
gender = getval(user_element,'gender')
location = getval(user_element,'location')
joined = getval(user_element,'joined')
last_active = getval(user_element,'last_active')
writer.writerow({'id': id, 'name' : name,'user_name' : user_name,
'profile_url' : profile_url,'image_url' : image_url,
'about' : about, 'age': age, 'gender' : gender,
'location' : location, 'joined' : joined, 'last_active': last_active})
print "Saved user data for user id:" + str(c)
# get list of user shelves
user_shelves_root = user_element.find('user_shelves')
user_shelf_list = []
for user_shelf in user_shelves_root.findall("user_shelf"):
shelf = getval(user_shelf,"name")
#Books on Shelf
print "Checking for books in shelf: " + shelf + " for user id:" + str(c)
shelf_url = "https://www.goodreads.com/review/list/"+ str(c) +".xml?key=i3Zsl7r13oHEQCjv1vXw&v=2&shelf=" + shelf
#time.sleep(1)
print shelf_url
response = urllib2.urlopen(shelf_url)
shelf_data_xml = response.read()
# write xml to file
f = open("xml_docs/user_shelf_" + shelf + "_"+ str(c) + ".xml", "w")
try:
f.write(shelf_data_xml)
finally:
f.close()
shelf_root = ET.parse("xml_docs/user_shelf_" + shelf + "_"+ str(c) + ".xml").getroot()
os.remove("xml_docs/user_shelf_" + shelf + "_"+ str(c) + ".xml")
reviews = shelf_root.find("reviews")
for review in reviews.findall("review"):
for book in review.findall("book"):
b_id = getval(book,"id")
isbn = getval(book,"isbn")
print "Fetching data for book with isbn:" + str(isbn) + " and id:" + str(id)
isbn13 = getval(book,"isbn13")
text_reviews_count = getval(book,"text_reviews_count")
title = getval(book,"title")
image_url = getval(book,"image_url")
link = getval(book,"link")
num_pages = getval(book,"num_pages")
b_format = getval(book,"format")
publisher = getval(book,"publisher")
publication_day = getval(book,"publication_day")
publication_year = getval(book, "publication_year")
publication_month = getval(book,"publication_month")
average_rating = getval(book,"average_rating")
ratings_count = getval(book,"rating_count")
description = getval(book,"description")
published = getval(book,"published")
#get number of books on each type of shelf
book_url = 'https://www.goodreads.com/book/show/'+str(b_id)+'.xml?key=i3Zsl7r13oHEQCjv1vXw'
response = urllib2.urlopen(book_url)
book_data_xml = response.read()
# write xml to file
f = open("xml_docs/book_data_" + str(b_id) + ".xml", "w")
try:
f.write(book_data_xml)
finally:
f.close()
book_root = ET.parse("xml_docs/book_data_" + str(b_id) + ".xml").getroot()
os.remove("xml_docs/book_data_" + str(b_id) + ".xml")
print "checking count in shelf for book_id:" + str(b_id)
book_root = book_root.find("book")
book_shelves = book_root.find("popular_shelves")
children = 0
religion = 0
history = 0
math = 0
anatology = 0
poetry = 0
encyclopedia = 0
dictionaries = 0
comics = 0
art = 0
cookbook = 0
diaries = 0
journals = 0
prayer_books = 0
series = 0
trilogy = 0
biographies = 0
autobiographies = 0
fantasy = 0
comic = 0
self_help = 0
science_fiction = 0
non_fiction = 0
for shelf_type in book_shelves.findall("shelf"):
attributes = shelf_type.attrib
name = attributes['name']
count = attributes['count']
print name + ":" + count
if(name == 'children'):
children = count
if(name =='religion'):
religion = count
if(name =='science-fiction'):
science_fiction = count
if(name =='history'):
history = count
if(name =='math'):
math = count
if(name =='anatology'):
anatology = count
if(name =='poetry'):
poetry = count
if(name =='encyclopedia'):
encyclopedia = count
if(name =='dictionaries'):
dictionaries = count
if(name =='comics'):
comics = count
if(name =='art'):
art = count
if(name =='cookbook'):
cookbook = count
if(name =='diaries'):
diaries = count
if(name =='journals'):
journals = count
if(name =='prayer-books'):
prayer_books = count
if(name =='series'):
series = count
if(name =='trilogy'):
trilogy = count
if(name =='biographies'):
biortaphies = count
if(name =='autobiographies'):
autobiographies = count
if(name =='fantasy'):
fantasy = count
if(name =='comic'):
comic = count
if(name =='self-help'):
self_help = count
if(name == 'non-fiction'):
non_fiction = count
writer_book.writerow({
'user_id': id,
'b_id' : b_id ,
'shelf' : shelf,
'isbn' : isbn,
'isbn13': isbn13,
'text_reviews_count' : text_reviews_count,
'title' : title,
'image_url' : image_url,
'link' : link,
'num_pages' : num_pages,
'b_format' : b_format,
'publisher' : publisher,
'publication_day' : publication_day,
'publication_year' : publication_year,
'publication_month' : publication_month,
'average_rating' : average_rating,
'ratings_count' : ratings_count,
'description' : description,
'children' : children,
'religion' : religion,
'history' : history,
'math': math,
'anatology' : anatology,
'poetry' : poetry,
'encyclopedia' : encyclopedia,
'dictionaries' : dictionaries,
'comics' : comics,
'art' : art,
'cookbook' : cookbook,
'diaries' : diaries,
'journals' : journals,
'prayer_books' : prayer_books,
'series' : series ,
'trilogy' : trilogy,
'biographies' : biographies ,
'autobiographies' : autobiographies,
'fantasy' : fantasy,
'comic' : comic,
'self-help' : self_help,
'science-fiction' : science_fiction,
'non-fiction' : non_fiction})
print "Data written on csv for book:" + title
getAmazonDetails(isbn)
print "Fetched review data from Amazon for book :" + title
authors = book.find("authors")
for author in authors.findall("author"):
a_id = getval(author,"id")
name = getval(author,"name")
average_rating = getval(author,"average_rating")
ratings_count = getval(author,"ratings_count")
text_reviews_count = getval(author,"text_reviews_count")
writer_author.writerow({'u_id': id,
'b_id' : b_id,
'a_id' : a_id,
'name' : name,
'average_rating' : average_rating,
'ratings_count' : ratings_count,
'text_reviews_count' : text_reviews_count})
except:
#time.sleep(1)
print "Exception!!"
print traceback.format_exc()
print "End of Program"
# -
f = open("xml_docs/book_data_" + "520980" + ".xml", "w")
try:
f.write(book_data_xml)
finally:
f.close()
book_root = ET.parse("xml_docs/book_data_" + "520980" + ".xml").getroot()
#os.remove("xml_docs/book_data_" + str(b_id) + ".xml")
print "checking count in shelf for book_id:" + "520980"
book_shelves = book_root.find("book")
book_in_shelf = book_shelves.find("popular_shelves")
children = 0
religion = 0
history = 0
math = 0
anatology = 0
poetry = 0
encyclopedia = 0
dictionaries = 0
comics = 0
art = 0
cookbook = 0
diaries = 0
journals = 0
prayer_books = 0
series = 0
trilogy = 0
biographie = 0
autobiographies = 0
fantasy = 0
print book_in_shelf.findall("shelf")
for shelf in book_in_shelf.findall("shelf"):
print shelf
# +
attributes = shelf.attrib
name = attributes['name']
print name
count = attributes['count']
print count
if(name == 'children'):
children = count
if(name =='religion'):
religion = count
if(name =='history'):
history = count
if(name =='math'):
math = count
if(name =='anatology'):
anatology = count
if(name =='poetry'):
poetry = count
if(name =='encyclopedia'):
encyclopedia = count
if(name =='dictionaries'):
dictionaries = count
if(name =='comics'):
comics = count
if(name =='art'):
art = count
if(name =='cookbook'):
cookbook = count
if(name =='diaries'):
diaries = count
if(name =='journals'):
journals = count
if(name =='prayer-books'):
prayer_books = count
if(name =='series'):
series = count
if(name =='trilogy'):
trilogy = count
if(name =='biographies'):
biortaphies = count
if(name =='autobiographies'):
autobiographies = count
if(name =='fantasy'):
fantasy = count
# -
| Amazon Crawler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
data = pd.read_csv(r'C:\Users\Biplob\Desktop\Python for Machine Learning and Data Science\All Projects - Data Analytics\3. Data Analysis with Python Police Dataset\3. Police Data.csv')
data
# # Instruction (For Data Cleaning)
# 1. Remove the column that only contains missing values
#
data.head()
data.isnull().sum()
data.drop(columns = 'country_name', inplace = True)
data
# # Question (Based on Filtering + Value Counts)
# 2. For Speeding, were Men or Women stopped more often ?
#For Speeding indicate violation column. Men or Women indicate driver_gender column.
data.head(5)
data[data.violation == 'Speeding'].driver_gender.value_counts()
# # Question (Groupby)
# 3. Does gender affect who gets searched during a stop?
data.head(5)
data.groupby('driver_gender').search_conducted.sum()
data.search_conducted.value_counts()
2113+366
# # Question (mapping + data-type casting)
# 4. What is the mean stop_duration ?
data.head(5)
data['stop_duration'].value_counts()
data['stop_duration'] = data['stop_duration'].map({'0-15 Min' : 7.5, '16-30 Min' : 24, '30+ Min' : 45})
data
data['stop_duration'].mean()
# # Question ( Groupby, Describe )
# 5. Compare the age distributions for each violation
data.head(5)
data.groupby('violation').driver_age.describe()
| 3. Data Analysis with Python Police Dataset/Data Analysis with Python - Police Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qfold cnot 7x2 Qiskit
# <a id = 'index'></a>
# * **Step 1.** - [Initial setup](#set)
#
# * **Step 2.** - [Defining the circuit](#nc)
#
# * **Step 3.** - [Optimizing the circuit](#op)
#
# * **Step 4.** - [Running in real device and data treatment](#real)
#
# * **Step 5.** - [Tests with different inputs](#oin)
# <a id = 'set'></a>
#
# ## Initial Setup
#
# To achieve results comparable to the ones present in paper *Compiling quantamorphisms for the IBM Q-Experience*, ensure the application of the correct imports and versions.
import qiskit
import qiskit.tools.jupyter
# %qiskit_version_table
# Useful additional packages
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
from math import pi
# these imports are essential since the new circuit section
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
# this import is essential since the simulation section
from qiskit import Aer
# these imports are essential since the optimization section
from qiskit import IBMQ
from qiskit.tools.monitor import backend_monitor, backend_overview
from qiskit.compiler import transpile
# this import is essential since run in real device section
from qiskit.tools.visualization import plot_histogram
# this is essential in the PyZX section
import pyzx as zx
# this is essential in the ignis section
from qiskit.ignis.mitigation.measurement import ( complete_meas_cal, CompleteMeasFitter, MeasurementFilter )
# #### Important/Useful functions
def circuit_inf(quantum_circuit):
circuit_information={}
# total number of operations in the circuit. no unrolling is done.
circuit_size = quantum_circuit.size()
circuit_information['size']=circuit_size
# depth of circuit (number of ops on the critical path)
circuit_depth = quantum_circuit.depth()
circuit_information['depth']=circuit_depth
# number of unentangled subcircuits in this circuit.
# each subcircuit can in principle be executed on a different quantum processor!
circuit_tensor = quantum_circuit.num_tensor_factors()
circuit_information['tensor factors']= circuit_tensor
# a breakdown of operations by type
circuit_count = quantum_circuit.count_ops()
circuit_information['operations']=circuit_count
return circuit_information
def running_circuit(circuit, backend, shots=1024):
job_run = execute(circuit, backend, shots=shots)
jobID_run = job_run.job_id()
result_run = job_run.result()
counts_run = result_run.get_counts(circuit)
return jobID_run, counts_run
# +
def sum_the_target_0(counts_raw):
k=counts_raw.keys()
sum_counts_ok=sum_counts_bad=0
lk=list(k)
for x in lk:
if x[3]=='0':
sum_counts_ok=sum_counts_ok+counts_raw.get(x)
else:
sum_counts_bad=sum_counts_bad+counts_raw.get(x)
return {'good': sum_counts_ok, 'bad': sum_counts_bad}
def sum_right(target, counts_raw):
s = sum_the_target_0(counts_raw)
if target==1:
s['good_temp'] = s.pop('good')
s['good']= s.pop('bad')
s['bad']= s.pop('good_temp')
return s
# -
# [back to top](#index)
# <a id = 'nc'></a>
#
# ## New Circuit
#
#
# Recall that the output of Quipper language goes thought the translator in quipperToQiskit. In this format, it is possible to define the circuit easily.
#
# Moreover, it is essential to simulate the experiment to see what are the ideal outputs.
# Qiskit swaps the least and the most significant qubits. Therefore, to keep conformity, there was a rearrangement of the least and most significant qubits.
#
# In other words,
# * qubit 0 is now qubit 4
# * qubit 1 is now qubit 3
# * qubit 2 holds
# * qubit 3 is not qubut 1
# * and qubit 4 is now qubit 0
#
# number of qubits
n = 5
# create quantum register named 'qr'
qr = QuantumRegister(n, 'qr')
# create classical register named 'cr'
cr = ClassicalRegister(n, 'cr')
#create quantum circuit
qc= QuantumCircuit(qr,cr)
# Go to the document `circuit_cnot_7x2_qiskit.txt`, select all the unitary gates, and copy to the following cell.
qc.draw(output='mpl', scale=0.5)
circuit_inf(qc)
# ### Simulation
# This simulation can run with Aer or BasicAer.
# +
# add measure gates
m4 = QuantumCircuit(qr, cr)
m4.measure(qr[4],cr[4])
m4.measure(qr[1],cr[1])
m4.measure(qr[2],cr[2])
m4.measure(qr[3],cr[3])
m4.draw(output='mpl')
# +
qc_m = qc + m4
qc_m.draw(output='mpl')
# -
circuit_inf(qc_m)
# +
# Use Aer's qasm_simulator
backend_sim = Aer.get_backend('qasm_simulator')
# Execute the circuit on the qasm simulator.
# We've set the number of repeats of the circuit
# to be 1024, which is the default.
job_sim = execute(qc_m, backend_sim, shots=1024)
# Grab the results from the job.
result_sim = job_sim.result()
# -
counts_sim = result_sim.get_counts(qc_m)
print(counts_sim)
# [back to top](#index)
# <a id = 'op'></a>
#
# ## Optimizing the circuit
#
#
# The considerable volume of the circuit displayed points to its optimization.
#
# +
# https://qiskit.org/documentation/install.html#access-ibm-quantum-systems
#
#provider = IBMQ.save_account('token')
provider = IBMQ.load_account()
# -
my_providers=IBMQ.providers()
print(my_providers)
# +
# you may not have this access
my_provider_academic = IBMQ.get_provider(hub='my_hub', group='my_group', project='my_project')
my_provider_academic.backends()
# +
my_provider_ibmq = IBMQ.get_provider(hub='ibm-q', group='open', project='main')
my_provider_ibmq.backends()
# -
# %qiskit_backend_overview
backend_overview()
# **Boebligen** - choosen because has hight T2 comparing to the others.
backend = my_provider_academic.get_backend('ibmq_boeblingen')
qc_sim = transpile(qc_m, backend=backend)
qc_sim.draw(output='mpl')
circuit_inf(qc_sim)
backend_monitor(backend)
# ### IBM Q Transpiler
#
# One trivial approach is to apply the IBM Q transpiler.
optimized_0 = transpile(qc_m, backend=backend, optimization_level=0)
circuit_inf(optimized_0)
optimized_1 = transpile(qc_m, backend=backend, optimization_level=1)
circuit_inf(optimized_1)
optimized_2 = transpile(qc_m, backend=backend, optimization_level=2)
circuit_inf(optimized_2)
optimized_3 = transpile(qc_m, backend=backend, optimization_level=3)
circuit_inf(optimized_3)
# <div class="alert alert-block alert-info">
# try with optimization 2 and 3
#
# <p>2 has less depth.</p>
#
# <p>3 has less cnot.</p>
# </div>
# ### PyZX
# Since this optimization was insufficient, the circuit ended rewritten with PyZX.
# +
my_qc = zx.Circuit.from_quipper_file("circuit_cnot_7x2_quipper_A.txt")
zx.draw(my_qc)
# -
print(my_qc.gates)
print(my_qc.stats())
mg = my_qc.to_graph()
print(mg)
zx.simplify.full_reduce(mg)
zx.draw(mg)
print(mg)
mg.normalise()
zx.draw(mg)
print(mg)
mc = zx.extract.streaming_extract(mg.copy(), True)
zx.draw(mc)
print(mc)
# Turn graph back into circuit
mc2 = zx.extract.streaming_extract(mg).to_basic_gates()
print(mc2.stats())
mc3 = zx.optimize.full_optimize(mc2)
print(mc3.stats())
print(mc3.to_quipper())
f = open("quipper_pyzx.txt", "w")
f.write(mc3.to_quipper())
f.close()
# * Open the `quipperToQiskit.gawk` file;
# * In line 2 change "qc" to "qc_pyzx";
# * Save;
# * Run the command line:
# ```
# awk -f quipperToQiskit.gawk circuit_cnot_7x2_quipper_A.txt > circuit_cnot_7x2_qiskit_pyzx.txt
# ```
qc_pyzx = QuantumCircuit(qr, cr)
# Go to the document `circuit_cnot_7x2_qiskit_pyzx.txt`, select all the unitary gates, and copy to the following cell.
qc_pyzx.draw(output='mpl')
circuit_inf(qc_pyzx)
qc_pyzx = qc_pyzx+m4
id_temp, counts_pyzx_sim = running_circuit(qc_pyzx, backend_sim)
print(counts_pyzx_sim)
qc_sim_pyzx = transpile(qc_pyzx, backend=backend)
circuit_inf(qc_sim_pyzx)
qc_pyzx_o2 = transpile(qc_pyzx, backend=backend, optimization_level=2)
circuit_inf(qc_pyzx_o2)
qc_pyzx_o3 = transpile(qc_pyzx, backend=backend, optimization_level=3)
circuit_inf(qc_pyzx_o3)
# [back to top](#index)
# <a id='real'> </a>
#
# ## Running in the real device and data treatment
#
#
# After reaching optimization, the IBM Q Experience Ignis module ensures the filtration of the results.
#
# Furthermore, the target as the most relevant qubit in the program leads to analyzing only the target result.
# %qiskit_job_watcher
shots=1024
# Save the job Id value to recover the job information later.
id_run, counts_dev = running_circuit(qc_m, backend, shots)
print(id_run)
# +
#id_run=''
job = backend.retrieve_job(id_run)
result_run= job.result()
counts_dev= result_run.get_counts()
print(counts_dev)
# -
id_opt2, counts_opt2 = running_circuit(optimized_2, backend, shots)
print(id_opt2)
# +
#id_opt2=''
job = backend.retrieve_job(id_opt2)
result_opt2= job.result()
counts_opt2= result_opt2.get_counts()
print(counts_opt2)
# -
id_opt3, counts_opt3 = running_circuit(optimized_3, backend, shots)
print(id_opt3)
# +
#id_opt3=''
job = backend.retrieve_job(id_opt3)
result_opt3= job.result()
counts_opt3= result_opt3.get_counts()
print(counts_opt3)
# -
id_pyzx, counts_pyzx = running_circuit(qc_pyzx, backend, shots)
print(id_pyzx, counts_pyzx)
# +
#id_pyzx=''
job = backend.retrieve_job(id_pyzx)
result_pyzx= job.result()
counts_pyzx= result_pyzx.get_counts()
print(counts_pyzx)
# -
id_pyzx_2, counts_pyzx_2 = running_circuit(qc_pyzx_o2, backend)
print(id_pyzx_2)
# +
#id_pyzx_2=''
job = backend.retrieve_job(id_pyzx_2)
result_pyzx_2= job.result()
counts_pyzx_2= result_pyzx_2.get_counts()
print(counts_pyzx_2)
# -
id_pyzx_3, counts_pyzx_3 = running_circuit(qc_pyzx_o3, backend)
print(id_pyzx_3)
# +
#id_pyzx_3=''
job = backend.retrieve_job(id_pyzx_3)
result_pyzx_3= job.result()
counts_pyzx_3= result_pyzx_3.get_counts()
print(counts_pyzx_3)
# -
leg = ['simulation', 'run in real device', 'transpiler with optimization 2', 'traspiler with optimization 3', 'optimization with compiler PyZX', 'optimization with compiler PyZX and 2', 'optimization with compiler PyZX and 3']
colors = ['#061727', '#003a6d', '#00539a', '#1192e8','#33b1ff','#82cfff','#e5f6ff']
plot_histogram([counts_sim, counts_dev,counts_opt2,counts_opt3,counts_pyzx,counts_pyzx_2,counts_pyzx_3], number_to_keep = 1, color=colors, legend = leg, figsize=(11, 5))
# ### Ignis
# Generate the calibration circuits
qr_ignis = QuantumRegister(5)
meas_calibs, state_labels = complete_meas_cal(qubit_list=[0,1,2,3,4], qr=qr_ignis, circlabel='mcal')
# +
job_ignis = execute(meas_calibs, backend=backend)
cal_results = job_ignis.result()
jobID_run_ignis = job_ignis.job_id()
print('JOB ID: {}'.format(jobID_run_ignis))
# -
#id_ignis=''
job_ignis = backend.retrieve_job(id_ignis)
cal_results= job_ignis.result()
meas_fitter = CompleteMeasFitter(cal_results, state_labels, circlabel='mcal')
# Plot the calibration matrix
meas_fitter.plot_calibration()
# What is the measurement fidelity?
print("Average Measurement Fidelity: %f" % meas_fitter.readout_fidelity())
# Get the filter object
meas_filter = meas_fitter.filter
job_pyzx_3 = backend.retrieve_job(id_pyzx_3)
result_pyzx_3= job_pyzx_3.result()
mitigated_results_py3 = meas_filter.apply(result_pyzx_3)
mitigated_counts_py3 = mitigated_results_py3.get_counts(0)
print(mitigated_counts_py3)
leg = ['simulation', 'run in real device', 'optimization with compiler PyZX and 3', 'mittigation']
colors = ['#061727', '#003a6d', '#82cfff','#e5f6ff']
plot_histogram([counts_sim, counts_dev,counts_pyzx, mitigated_counts_py3], number_to_keep = 1, color=colors, legend = leg, figsize=(11, 5))
# ### Find just the ones where the target qubit holds |0>
counts_sim_0 = sum_right(0, counts_sim)
print(counts_sim_0)
counts_dev_0 = sum_right(0, counts_dev)
print(counts_dev_0)
counts_opt2_0 = sum_right(0, counts_opt2)
print(counts_opt2_0)
counts_opt3_0 = sum_right(0, counts_opt3)
print(counts_opt3_0)
counts_pyzx_0 = sum_right(0, counts_pyzx)
print(counts_pyzx_0)
counts_pyzx2_0 = sum_right(0, counts_pyzx_2)
print(counts_pyzx2_0)
counts_pyzx3_0 = sum_right(0, counts_pyzx_3)
print(counts_pyzx3_0)
mitigated_counts_0= sum_right(0, mitigated_counts_py3)
print(mitigated_counts_0)
leg = ['simulation', 'run in real device', 'optimization with compiler PyZX and 3 and mittigation']
colors = ['#061727', '#003a6d', '#1192e8']
plot_histogram([counts_sim_0, counts_dev_0, mitigated_counts_0], title='input=|0000>', color=colors, legend = leg, figsize=(5, 5))
# [back to top](#index)
# <a id = 'oin'></a>
#
# ## Tests with different inputs
#
# To obtain a faithful experience, tests with different inputs are essential.
# When the controls are |101> the target should change (the initial target value is |1>)
qc_init_1011 = QuantumCircuit(qr,cr)
# +
qc_init_1011.x(qr[2])
qc_init_1011.x(qr[4])
qc_init_1011.x(qr[1])
qc_init_1011.draw(output='mpl')
# +
qc_1011 = qc_init_1011 + qc_m
qc_1011.draw(output='mpl')
# +
qc_1011_pyzx = qc_init_1011 + qc_pyzx
qc_1011_pyzx.draw(output='mpl')
# -
id_temp, counts_sim_1011 = running_circuit(qc_1011, backend_sim)
print(counts_sim_1011)
id_temp, counts_sim_1011_p = running_circuit(qc_1011_pyzx, backend_sim)
print(counts_sim_1011_p)
id_run_1011, counts_dev_1011 = running_circuit(qc_1011, backend)
print(id_run_1011)
# +
#id_run_1011=''
job = backend.retrieve_job(id_run_1011)
result_run_1011= job.result()
counts_dev_1011= result_run_1011.get_counts()
print(counts_dev_1011)
# -
id_pyzx_1011, counts_pyzx_1011 = running_circuit(qc_1011_pyzx, backend)
print(id_pyzx_1011)
# +
#id_pyzx_1011=''
job = backend.retrieve_job(id_pyzx_1011)
result_pyzx_1011= job.result()
counts_pyzx_1011= result_pyzx_1011.get_counts()
print(counts_pyzx_1011)
# -
qc_pyzx_o3_1011 = transpile(qc_1011_pyzx, backend=backend, optimization_level=3)
id_temp, counts_sim_1011_p3 = running_circuit(qc_pyzx_o3_1011, backend_sim)
print(counts_sim_1011_p3)
id_pyzx3_1011, counts_pyzx3_1011 = running_circuit(qc_pyzx_o3_1011, backend)
print(id_pyzx3_1011)
# +
#id_pyzx3_1011=''
job = backend.retrieve_job(id_pyzx3_1011)
result_pyzx3_1011= job.result()
counts_pyzx3_1011= result_pyzx3_1011.get_counts()
print(counts_pyzx3_1011)
# -
qc_pyzx_o2_1011 = transpile(qc_1011_pyzx, backend=backend, optimization_level=2)
id_temp, counts_sim_1011_p2 = running_circuit(qc_pyzx_o2_1011, backend_sim)
print(counts_sim_1011_p2)
id_pyzx2_1011, counts_pyzx2_1011 = running_circuit(qc_pyzx_o2_1011, backend)
print(id_pyzx2_1011)
# +
#id_pyzx2_1011=''
job = backend.retrieve_job(id_pyzx2_1011)
result_pyzx2_1011= job.result()
counts_pyzx2_1011= result_pyzx2_1011.get_counts()
print(counts_pyzx2_1011)
# -
leg = ['simulation', 'run in real device','optimization with compiler PyZX','optimization with compiler PyZX and 2', 'optimization with compiler PyZX and 3']
colors = ['#061727', '#003a6d', '#1192e8','#82cfff','#e5f6ff']
plot_histogram([counts_sim_1011, counts_dev_1011, counts_pyzx_1011, counts_pyzx2_1011, counts_pyzx3_1011], target_string='10100', number_to_keep=1,color=colors, legend = leg, figsize=(11, 5), title='intup |101> with target input |1>')
job_pyzx_2_1011 = backend.retrieve_job(id_pyzx2_1011)
result_pyzx2_1011= job_pyzx_2_1011.result()
mitigated_results_py2_1011 = meas_filter.apply(result_pyzx2_1011)
mitigated_counts_py2_1011 = mitigated_results_py2_1011.get_counts(0)
print(mitigated_counts_py2_1011)
job_pyzx_1011 = backend.retrieve_job(id_pyzx_1011)
result_pyzx_1011= job_pyzx_1011.result()
mitigated_results_py_1011 = meas_filter.apply(result_pyzx_1011)
mitigated_counts_py_1011 = mitigated_results_py_1011.get_counts(0)
print(mitigated_counts_py_1011)
leg = ['simulation', 'run in real device','optimization with compiler PyZX', 'mittigation', 'optimization with compiler PyZX and 2', 'mittigation 2']
colors = ['#061727', '#003a6d', '#00539a', '#1192e8','#33b1ff','#82cfff']
plot_histogram([counts_sim_1011, counts_dev_1011, counts_pyzx_1011,mitigated_counts_py_1011, counts_pyzx2_1011,mitigated_counts_py2_1011 ], target_string='10100', number_to_keep = 1, color=colors, legend = leg, figsize=(11, 5), title='intup |101> with target input |1>')
counts_sim_2 = sum_right(0, counts_sim_1011)
print(counts_sim_2)
counts_dev_2 = sum_right(0, counts_dev_1011)
print(counts_dev_2)
counts_pyzx_2 = sum_right(0, counts_pyzx_1011)
print(counts_pyzx_2)
counts_pyzx2_2 = sum_right(0, counts_pyzx2_1011)
print(counts_pyzx2_2)
counts_pyzx3_2 = sum_right(0, counts_pyzx3_1011)
print(counts_pyzx3_2)
mitigated_counts_2= sum_right(0, mitigated_counts_py_1011)
print(mitigated_counts_2)
mitigated_counts_2_2= sum_right(0, mitigated_counts_py2_1011)
print(mitigated_counts_2_2)
leg = ['simulation', 'run in real device', 'optimization with compiter PyZX and mittigation']
colors = ['#061727', '#003a6d', '#1192e8']
plot_histogram([counts_sim_2, counts_dev_2, mitigated_counts_2], title='input=|1011>', color=colors, legend = leg, figsize=(5, 5))
# #### other important simulations
# +
qubit_controls = 3
qubit_target = 1
total_qubits = 3+1
number_inputs = 2**total_qubits
data = np.arange(number_inputs)
data_input=[]
for i in data:
data_input.append(bin(i)[2:].zfill(total_qubits))
print(data_input)
# -
for i in data_input:
print('init circuit: ', i[::-1])
count=0
qcircuit = QuantumCircuit(qr,cr)
for y in i:
if y == '1':
qcircuit.x(qr[count+1])
count = count+1
qcircuit = qcircuit + qc_m
id_temp, counts_temp = running_circuit(qcircuit, backend_sim)
print('output: ', list(counts_temp.keys())[0][:-1])
for i in data_input:
print('init circuit: ', i[::-1])
count=0
qcircuit = QuantumCircuit(qr,cr)
for y in i:
if y == '1':
qcircuit.x(qr[count+1])
count = count+1
qcircuit = qcircuit + qc_pyzx
id_temp, counts_temp = running_circuit(qcircuit, backend_sim)
print('output: ', list(counts_temp.keys())[0][:-1])
# [back to top](#index)
| qfold_cnot_7x2_qiskit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: 'Python 3.6.10 64-bit (''PythonData'': conda)'
# name: python3610jvsc74a57bd0a7795a94fcb98f890ecfbefedba6c51e1f5a02e9905bd71b2049174ccd39ef45
# ---
# install joblib. This will be used to save your model.
# Restart your kernel after installing
# !pip install joblib
import pandas as pd
# # Read the CSV and Perform Basic Data Cleaning
df = pd.read_csv("exoplanet_data.csv")
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df.head()
df = df[df.koi_disposition!="CANDIDATE"]
# # Select your features (columns)
# Set features. This will also be used as your x values.
X = df.drop("koi_disposition", axis = 1)
X
y = pd.get_dummies(df.koi_disposition)
y
print(X.shape, y.shape)
# +
# Create a Train Test Split
# Use `koi_disposition` for the y values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y.CONFIRMED)
# -
X_train.shape
# # Pre-processing
#
# Scale the data using the MinMaxScaler and perform some feature selection
# Scale your data
from numpy import asarray
from sklearn.preprocessing import MinMaxScaler
X_scaler = MinMaxScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
from tensorflow.keras.utils import to_categorical
# One-hot encoding
y_train_categorical = to_categorical(y_train)
y_test_categorical = to_categorical(y_test)
y_train_categorical
# # Define Model Architecture
from keras.wrappers.scikit_learn import KerasClassifier
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
import numpy
# Function to create model, required for KerasClassifier
def create_model():
# create model
model = Sequential()
model.add(Dense(12, input_dim=8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
# fix random seed for reproducibility
seed = 7
numpy.random.seed(seed)
# # Train the Model
#
#
# create model
model = KerasClassifier(build_fn=create_model, verbose=0)
from sklearn.model_selection import GridSearchCV
# define the grid search parameters
batch_size = [10, 20, 40, 60, 80, 100]
epochs = [10, 50, 100]
param_grid = dict(batch_size=batch_size, epochs=epochs)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1, cv=3)
grid_result = grid.fit(X_train_scaled, y_train_categorical)
model_loss, model_accuracy = model.evaluate(
X_test_scaled, y_test_categorical, verbose=2)
print(
f"Normal Neural Network - Loss: {model_loss}, Accuracy: {model_accuracy}")
print(grid.best_params_)
print(grid.best_score_)
# # Save the Model
# save your model by updating "your_name" with your name
# and "your_model" with your model variable
# be sure to turn this in to BCS
# if joblib fails to import, try running the command to install in terminal/git-bash
import joblib
filename = 'Neural Network.sav'
joblib.dump(grid.best_score_, filename)
| neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import tensorflow as tf
import os, random, time
# +
# Hyperparameters
batch_size = 128
learning_rate = 0.003
training_epochs = 40
display_step = 10
# To prevent overfitting
dropout = 0.75
summaries_dir = "./logs"
# +
# Dataset and iterator creation
in_dir = "../input/preprocessed-normalized/"
test_normal_dir = in_dir + "test/NORMAL"
test_pneumonia_dir = in_dir + "test/PNEUMONIA"
train_normal_dir = in_dir + "train/NORMAL"
train_pneumonia_dir = in_dir + "train/PNEUMONIA"
full_url = np.vectorize(lambda url,prev_url: prev_url+"/"+url)
test_normal_data = pd.DataFrame(full_url(np.array(os.listdir(test_normal_dir)),test_normal_dir), columns=["image_dir"])
test_pneumonia_data = pd.DataFrame(full_url(np.array(os.listdir(test_pneumonia_dir)),test_pneumonia_dir), columns=["image_dir"])
train_normal_data = pd.DataFrame(full_url(np.array(os.listdir(train_normal_dir)),train_normal_dir), columns=["image_dir"])
train_pneumonia_data = pd.DataFrame(full_url(np.array(os.listdir(train_pneumonia_dir)),train_pneumonia_dir), columns=["image_dir"])
test_normal_data["class"] = "NORMAL"
test_pneumonia_data["class"] = "PNEUNOMIA"
train_normal_data["class"] = "NORMAL"
train_pneumonia_data["class"] = "PNEUNOMIA"
test_data = test_normal_data.append(test_pneumonia_data)
train_data = train_normal_data.append(train_pneumonia_data)
# Total ammount of landmarks
n_classes = 2
with tf.device('/cpu:0'):
# Reads an image from a file, decodes it into a dense tensor, and resizes it
# to a fixed shape.
def _parse_function(filename, label):
image_string = tf.read_file(filename)
image_decoded = tf.image.decode_jpeg(image_string)
image_decoded = tf.cast(image_decoded, tf.float32)
image_decoded.set_shape((256, 256, 1))
return image_decoded, label
def _parse_rotate_function(filename, label):
image_string = tf.read_file(filename)
image_decoded = tf.image.decode_jpeg(image_string)
image_decoded = tf.contrib.image.rotate(image_decoded, random.uniform(-3,3))
image_decoded = tf.cast(image_decoded, tf.float32)
image_decoded.set_shape((256, 256, 1))
return image_decoded, label
train_data = tf.data.Dataset.from_tensor_slices(
(train_data["image_dir"].values,
pd.get_dummies(train_data["class"]).values))
train_data = train_data.shuffle(buffer_size=10000)
# for a small batch size
train_data = train_data.map(_parse_function, num_parallel_calls=4)
# train_data = train_data.map(_parse_rotate_function, num_parallel_calls=4)
train_data = train_data.batch(batch_size)
# for a large batch size (hundreds or thousands)
# dataset = dataset.apply(tf.contrib.data.map_and_batch(
# map_func=_parse_function, batch_size=batch_size))
# with gpu usage
train_data = train_data.prefetch(1)
test_data = tf.data.Dataset.from_tensor_slices(
(test_data["image_dir"].values,
pd.get_dummies(test_data["class"]).values))
test_data = test_data.map(_parse_function, num_parallel_calls=4)
test_data = test_data.batch(batch_size)
test_data = test_data.prefetch(1)
iterator = tf.data.Iterator.from_structure(train_data.output_types,
train_data.output_shapes)
x, y = iterator.get_next()
train_init = iterator.make_initializer(train_data) # Inicializador para train_data
test_init = iterator.make_initializer(test_data) # Inicializador para test_data
# +
# Placeholder
# x = tf.placeholder(dtype=tf.float32, shape=[None, 256, 256, 1])
# y = tf.placeholder(dtype=tf.float32, shape=[None, n_classes])
# Visualize input x
tf.summary.image("input", x, batch_size)
def conv2d(img, w, b):
return tf.nn.relu(tf.nn.bias_add\
(tf.nn.conv2d(img, w,\
strides=[1, 1, 1, 1],\
padding='SAME'),b))
def max_pool(img, k):
return tf.nn.max_pool(img, \
ksize=[1, k, k, 1],\
strides=[1, k, k, 1],\
padding='SAME')
def avg_pool(img, k):
return tf.nn.avg_pool(img, \
ksize=[1, k, k, 1],\
strides=[1, k, k, 1],\
padding='SAME')
# weights and bias
wc1 = tf.Variable(tf.random_normal([3, 3, 1, 32]))
bc1 = tf.Variable(tf.random_normal([32]))
# pool 128x128
wc2 = tf.Variable(tf.random_normal([3, 3, 32, 32]))
bc2 = tf.Variable(tf.random_normal([32]))
# pool 64x64
wd1 = tf.Variable(tf.random_normal([64*64*32, 512]))
bd1 = tf.Variable(tf.random_normal([512]))
wd2 = tf.Variable(tf.random_normal([512, 1024]))
bd2 = tf.Variable(tf.random_normal([1024]))
wout = tf.Variable(tf.random_normal([1024, n_classes]))
bout = tf.Variable(tf.random_normal([n_classes]))
tf.summary.histogram("weights", wc1)
tf.summary.histogram("bias", bc1)
tf.summary.histogram("weights", wc2)
tf.summary.histogram("bias", bc2)
tf.summary.histogram("weights", wd1)
tf.summary.histogram("bias", bd1)
tf.summary.histogram("weights", wd2)
tf.summary.histogram("bias", bd2)
tf.summary.histogram("weights", wout)
tf.summary.histogram("bias", bout)
# conv layer
conv1 = conv2d(x,wc1,bc1)
tf.summary.histogram("activations", conv1)
# Max Pooling (down-sampling), this chooses the max value from a 2*2 matrix window and outputs a 128*128 matrix.
conv1 = max_pool(conv1, k=2)
# conv1 = avg_pool(conv1, k=2)
# dropout to reduce overfitting
keep_prob = tf. placeholder(tf.float32)
conv1 = tf.nn.dropout(conv1,keep_prob)
# conv layer
conv2 = conv2d(conv1,wc2,bc2)
tf.summary.histogram("activations", conv2)
# Max Pooling (down-sampling), this chooses the max value from a 2*2 matrix window and outputs a 64*64 matrix.
conv2 = max_pool(conv2, k=2)
# conv2 = avg_pool(conv2, k=2)
# dropout to reduce overfitting
conv2 = tf.nn.dropout(conv2, keep_prob)
# fc 1
dense1 = tf.reshape(conv2, [-1, wd1.get_shape().as_list()[0]])
# dense1 = tf.reshape(conv3, [-1, wd1.get_shape().as_list()[0]])
dense1 = tf.nn.relu(tf.add(tf.matmul(dense1, wd1),bd1))
tf.summary.histogram("activations", dense1)
dense1 = tf.nn.dropout(dense1, keep_prob)
# fc 2
dense2 = tf.reshape(dense1, [-1, wd2.get_shape().as_list()[0]])
dense2 = tf.nn.relu(tf.add(tf.matmul(dense2, wd2),bd2))
tf.summary.histogram("activations", dense2)
dense2 = tf.nn.dropout(dense2, keep_prob)
# prediction
# pred = tf.add(tf.matmul(dense1, wout), bout)
pred = tf.add(tf.matmul(dense2, wout), bout)
tf.summary.histogram("activations", pred)
with tf.name_scope("cross_entropy"):
# softmax
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=pred, labels=y))
cost = tf.summary.scalar("cross_entropy", cost)
# Optimizer
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
with tf.name_scope("accuracy"):
# Accuracy
correct_pred = tf.equal(tf.argmax(pred,1), tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
tf.summary.scalar("accuracy", accuracy)
# Get all summary
summ = tf.summary.merge_all()
# +
# Session start
init = tf.global_variables_initializer()
with tf.Session() as sess:
train_writer = tf.summary.FileWriter(summaries_dir + '/train', sess.graph)
test_writer = tf.summary.FileWriter(summaries_dir + '/test', sess.graph)
# Required to get the filename matching to run.
sess.run(init)
step = 1
# Compute epochs.
for i in range(training_epochs):
print("epoch: {}".format(i))
epoch_start = time.time()
sess.run(train_init)
try:
while True:
_, acc, loss, sumaries = sess.run([optimizer, accuracy, cost, summ], feed_dict={keep_prob: dropout})
train_writer.add_summary(sumaries, step)
if step % display_step == 0:
# acc = sess.run(accuracy, feed_dict={keep_prob: 1.})
# loss = sess.run(cost, feed_dict={keep_prob: 1.})
# train_writer.add_summary(loss, step)
print("step: {}".format(step))
print("accuracy: {}".format(acc))
print("loss: {}".format(loss))
print("\n")
step += 1
except tf.errors.OutOfRangeError:
print("epoch finished in {} seconds".format(time.time() - epoch_start))
# Test
print("Test\n")
sess.run(test_init)
avg_acc = 0
avg_loss = 0
steps=0
try:
while True:
acc, loss = sess.run([accuracy, cost], feed_dict={keep_prob: 1.})
avg_acc += acc
avg_loss += loss
steps += 1
# test_writer.add_summary(loss, step)
except tf.errors.OutOfRangeError:
print("Average test set accuracy over {} iterations is {:.2f}%".format(steps,(avg_acc / steps) * 100))
print("Average test set loss over {} iterations is {:.2f}".format(steps,(avg_loss / steps)))
| nn/.ipynb_checkpoints/conv-nn-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font color=blue>NOTE: Videos and graphs do not display immediately on all devices. If this is the case for you, click the "Cell" menu option, and select "Run All". If you still do not see the videos/graphs, try a different browser. If that also does not work, please let your instructor know by email.</font>
#
# # Volumes of Solids of Revolution
#
# In this notebook, we continue the study of volumes of solids. Here, we focus on special solids, namely solids of revolution.
#
# ### 1. What is a Solid of Revolution?
#
# A solid of revolution is a symmetric object, obtained by rotation a region in the plane around a horizontal line in the plane (assuming that this line does not intersect the region).
#
# For example, the unit sphere can be understood to be the result of revolving the upper half of the unit circle around the $x$-axis. Similarly, the right triangular cone can be understood to be the result of revolving a right triangle about either one of the sides that is not the hypotenuse, as illustrated below.
#
# 
#
# The following animation nicely demonstrates the result of rotating a <i>curve</i>, resulting in a <i>surface of revolution</i>. The surface of revolution encloses a <i>solid of revolution</i>.
#
# 
# Credit: By <a href="//commons.wikimedia.org/w/index.php?title=User:Macks&action=edit&redlink=1" class="new" title="User:Macks (page does not exist)">Macks</a> - <span class="int-own-work" lang="en">Own work</span><a href="//commons.wikimedia.org/wiki/File:Mathematica_Logo.svg" title="File:Mathematica Logo.svg"></a>This diagram was created with <a href="https://en.wikipedia.org/wiki/Mathematica" class="extiw" title="w:Mathematica">Mathematica</a>, <a href="https://creativecommons.org/licenses/by-sa/2.5" title="Creative Commons Attribution-Share Alike 2.5">CC BY-SA 2.5</a>, <a href="https://commons.wikimedia.org/w/index.php?curid=1596829">Link</a>
#
# ### 2. Slicing a Solid of Revolution to Determine Its Volume
#
# In this course, we restrict ourselves to the case where a planar region is rotated around a horizontal line.
#
# The general approach to determine the volume of the resulting solid of revolution is as follows:
# 1. We focus on a typical slice of the planar region, of infinitessimal thickness $dx$.
# 2. We rotate the typical slice around the given axis of revolution.
# 3. The typical slice generates either a solid disk, or a washer (a disk with a hole).
# 4. We determine the cross-sectional area of the disk or washer, $A(x)$.
# 5. We determine the volume of the disk or washer, $dV = A(x) dx$.
# 6. We integrate to obtain the volume of the entire object, $$V = \int dV = \int_a^b A(x) \ dx.$$
#
# ### 3. Solids of Revolution with Cross-Sections that are Disks
#
# Both the sphere and the right triangular cone are solids of revolution where the cross-sections are disks.
#
# In the following video, we will use the approach outlined above to determine the volume of a sphere with radius $R$. We have known since kindergarten (give or take a few years!) that the volume of a sphere with radius $R$ is $$V_{sphere} = \frac{4}{3} \pi R^3.$$ With the calculus tools that we now have at hand, we finally can prove the origin of this formula. This is pretty powerfull stuff!
from IPython.display import YouTubeVideo
YouTubeVideo('15j8gkLivX8')
# ### 4. Solids of Revolution with Cross-Sections that are Washers
#
# In many cases, the cross-sections of a solid of revolution are disks with holes, also known as washers.
#
# In the following video, we demonstrate how the disk method from the previous example can be modified to account for the hole.
from IPython.display import YouTubeVideo
YouTubeVideo('QZkMiczJI8g')
# Remember to subtract the area of the hole from the area of the solid disk, resulting in $$A = \pi \left( R_{outer}^2 - R_{inner}^2 \right).$$
#
# > <font color=red><b>Caution:</b> A very common mistake made by students is to write $$A = \pi \left( R_{outer} - R_{inner} \right)^2,$$
# which definitely is not the same (check it by multiplying out the latter)! </font>
#
# ### 5. Summary
#
# - The volume of a solid of revolution obtained by rotating a region in the plane between $x=a$ and $x=b$ around a horizontal axis is $$V = \int dV = \int_a^b A(x) \ dx,$$ where $A(x)$ is the cross-sectional area of a <b>disk</b> or a <b>washer</b>.
#
# - The cross-sectional area of a <b>disk</b> with radius $R(x)$ is $$A(x) = \pi R^2(x).$$
#
# - The cross-secional area of a <b>washer</b> with outer radius $R_{outer}(x)$ and inner radius (radius of the hole) $R_{inner}(x)$ is $$A(x) = \pi \left( R_{outer}^2(x) - R_{inner}^2(x) \right).$$
#
# ### 6. Further Study
#
# Please refer to Section 6.4 in the textbook for additional treatment of this topic.
#
# ### 7. Don't Forget
#
# Don't forget to return to eClass to complete the pre-class quiz.
| VolumesOfSolidsOfRevolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# About the author:
# This notebook was forked from this [project](https://github.com/fonnesbeck/scipy2014_tutorial). The original author is <NAME>, Assistant Professor of Biostatistics. You can follow Chris on Twitter [@fonnesbeck](https://twitter.com/fonnesbeck).
# #### Introduction
#
# For most problems of interest, Bayesian analysis requires integration over multiple parameters, making the calculation of a [posterior](https://en.wikipedia.org/wiki/Posterior_probability) intractable whether via analytic methods or standard methods of numerical integration.
#
# However, it is often possible to *approximate* these integrals by drawing samples
# from posterior distributions. For example, consider the expected value (mean) of a vector-valued random variable $\mathbf{x}$:
#
# $$
# E[\mathbf{x}] = \int \mathbf{x} f(\mathbf{x}) \mathrm{d}\mathbf{x}\,, \quad
# \mathbf{x} = \{x_1, \ldots, x_k\}
# $$
#
# where $k$ (dimension of vector $\mathbf{x}$) is perhaps very large.
# If we can produce a reasonable number of random vectors $\{{\bf x_i}\}$, we can use these values to approximate the unknown integral. This process is known as [**Monte Carlo integration**](https://en.wikipedia.org/wiki/Monte_Carlo_integration). In general, Monte Carlo integration allows integrals against probability density functions
#
# $$
# I = \int h(\mathbf{x}) f(\mathbf{x}) \mathrm{d}\mathbf{x}
# $$
#
# to be estimated by finite sums
#
# $$
# \hat{I} = \frac{1}{n}\sum_{i=1}^n h(\mathbf{x}_i),
# $$
#
# where $\mathbf{x}_i$ is a sample from $f$. This estimate is valid and useful because:
#
# - $\hat{I} \rightarrow I$ with probability $1$ by the [strong law of large numbers](https://en.wikipedia.org/wiki/Law_of_large_numbers#Strong_law);
#
# - simulation error can be measured and controlled.
# ### Example (Negative Binomial Distribution)
#
# We can use this kind of simulation to estimate the expected value of a random variable that is negative binomial-distributed. The [negative binomial distribution](https://en.wikipedia.org/wiki/Negative_binomial_distribution) applies to discrete positive random variables. It can be used to model the number of Bernoulli trials that one can expect to conduct until $r$ failures occur.
# The [probability mass function](https://en.wikipedia.org/wiki/Probability_mass_function) reads
#
# $$
# f(k \mid p, r) = {k + r - 1 \choose k} (1 - p)^k p^r\,,
# $$
#
# where $k \in \{0, 1, 2, \ldots \}$ is the value taken by our non-negative discrete random variable and
# $p$ is the probability of success ($0 < p < 1$).
#
#
# 
# Most frequently, this distribution is used to model *overdispersed counts*, that is, counts that have variance larger
# than the mean (i.e., what would be predicted under a
# [Poisson distribution](http://en.wikipedia.org/wiki/Poisson_distribution)).
#
# In fact, the negative binomial can be expressed as a continuous mixture of Poisson distributions,
# where a [gamma distributions](http://en.wikipedia.org/wiki/Gamma_distribution) act as mixing weights:
#
# $$
# f(k \mid p, r) = \int_0^{\infty} \text{Poisson}(k \mid \lambda) \,
# \text{Gamma}_{(r, (1 - p)/p)}(\lambda) \, \mathrm{d}\lambda,
# $$
#
# where the parameters of the gamma distribution are denoted as (shape parameter, inverse scale parameter).
#
# Let's resort to simulation to estimate the mean of a negative binomial distribution with $p = 0.7$ and $r = 3$:
# +
import numpy as np
r = 3
p = 0.7
# -
# Simulate Gamma means (r: shape parameter; p / (1 - p): scale parameter).
lam = np.random.gamma(r, p / (1 - p), size=100)
# Simulate sample Poisson conditional on lambda.
sim_vals = np.random.poisson(lam)
sim_vals.mean()
# The actual expected value of the negative binomial distribution is $r p / (1 - p)$, which in this case is 7. That's pretty close, though we can do better if we draw more samples:
lam = np.random.gamma(r, p / (1 - p), size=100000)
sim_vals = np.random.poisson(lam)
sim_vals.mean()
# This approach of drawing repeated random samples in order to obtain a desired numerical result is generally known as **Monte Carlo simulation**.
#
# Clearly, this is a convenient, simplistic example that did not require simuation to obtain an answer. For most problems, it is simply not possible to draw independent random samples from the posterior distribution because they will generally be (1) multivariate and (2) not of a known functional form for which there is a pre-existing random number generator.
#
# However, we are not going to give up on simulation. Though we cannot generally draw independent samples for our model, we can usually generate *dependent* samples, and it turns out that if we do this in a particular way, we can obtain samples from almost any posterior distribution.
# ## Markov Chains
#
# A Markov chain is a special type of *stochastic process*. The standard definition of a stochastic process is an ordered collection of random variables:
#
# $$
# \{X_t:t \in T\}
# $$
#
# where $t$ is frequently (but not necessarily) a time index. If we think of $X_t$ as a state $X$ at time $t$, and invoke the following dependence condition on each state:
#
# \begin{align*}
# &Pr(X_{t+1}=x_{t+1} | X_t=x_t, X_{t-1}=x_{t-1},\ldots,X_0=x_0) \\
# &= Pr(X_{t+1}=x_{t+1} | X_t=x_t)
# \end{align*}
#
# then the stochastic process is known as a Markov chain. This conditioning specifies that the future depends on the current state, but not past states. Thus, the Markov chain wanders about the state space,
# remembering only where it has just been in the last time step.
#
# The collection of transition probabilities is sometimes called a *transition matrix* when dealing with discrete states, or more generally, a *transition kernel*.
#
# It is useful to think of the Markovian property as **mild non-independence**.
#
# If we use Monte Carlo simulation to generate a Markov chain, this is called **Markov chain Monte Carlo**, or MCMC. If the resulting Markov chain obeys some important properties, then it allows us to indirectly generate independent samples from a particular posterior distribution.
#
#
# > ### Why MCMC Works: Reversible Markov Chains
# >
# > Markov chain Monte Carlo simulates a Markov chain for which some function of interest
# > (e.g., the joint distribution of the parameters of some model) is the unique, invariant limiting distribution. An invariant distribution with respect to some Markov chain with transition kernel $Pr(y \mid x)$ implies that:
# >
# > $$\int_x Pr(y \mid x) \pi(x) dx = \pi(y).$$
# >
# > Invariance is guaranteed for any *reversible* Markov chain. Consider a Markov chain in reverse sequence:
# > $\{\theta^{(n)},\theta^{(n-1)},...,\theta^{(0)}\}$. This sequence is still Markovian, because:
# >
# > $$Pr(\theta^{(k)}=y \mid \theta^{(k+1)}=x,\theta^{(k+2)}=x_1,\ldots ) = Pr(\theta^{(k)}=y \mid \theta^{(k+1)}=x)$$
# >
# > Forward and reverse transition probabilities may be related through Bayes theorem:
# >
# > $$\frac{Pr(\theta^{(k+1)}=x \mid \theta^{(k)}=y) \pi^{(k)}(y)}{\pi^{(k+1)}(x)}$$
# >
# > Though not homogeneous in general, $\pi$ becomes homogeneous if:
# >
# > - $n \rightarrow \infty$
# >
# > - $\pi^{(i)}=\pi$ for some $i < k$
# >
# > If this chain is homogeneous it is called reversible, because it satisfies the ***detailed balance equation***:
# >
# > $$\pi(x)Pr(y \mid x) = \pi(y) Pr(x \mid y)$$
# >
# > Reversibility is important because it has the effect of balancing movement through the entire state space. When a Markov chain is reversible, $\pi$ is the unique, invariant, stationary distribution of that chain. Hence, if $\pi$ is of interest, we need only find the reversible Markov chain for which $\pi$ is the limiting distribution.
# > This is what MCMC does!
# ## Gibbs Sampling
#
# The Gibbs sampler is the simplest and most prevalent MCMC algorithm. If a posterior has $k$ parameters to be estimated, we may condition each parameter on current values of the other $k-1$ parameters, and sample from the resultant distributional form (usually easier), and repeat this operation on the other parameters in turn. This procedure generates samples from the posterior distribution. Note that we have now combined Markov chains (conditional independence) and Monte Carlo techniques (estimation by simulation) to yield Markov chain Monte Carlo.
#
# Here is a stereotypical Gibbs sampling algorithm:
#
# 1. Choose starting values for states (parameters):
# ${\bf \theta} = [\theta_1^{(0)},\theta_2^{(0)},\ldots,\theta_k^{(0)}]$.
#
# 2. Initialize counter $j=1$.
#
# 3. Draw the following values from each of the $k$ conditional
# distributions:
#
# $$\begin{aligned}
# \theta_1^{(j)} &\sim& \pi(\theta_1 | \theta_2^{(j-1)},\theta_3^{(j-1)},\ldots,\theta_{k-1}^{(j-1)},\theta_k^{(j-1)}) \\
# \theta_2^{(j)} &\sim& \pi(\theta_2 | \theta_1^{(j)},\theta_3^{(j-1)},\ldots,\theta_{k-1}^{(j-1)},\theta_k^{(j-1)}) \\
# \theta_3^{(j)} &\sim& \pi(\theta_3 | \theta_1^{(j)},\theta_2^{(j)},\ldots,\theta_{k-1}^{(j-1)},\theta_k^{(j-1)}) \\
# \vdots \\
# \theta_{k-1}^{(j)} &\sim& \pi(\theta_{k-1} | \theta_1^{(j)},\theta_2^{(j)},\ldots,\theta_{k-2}^{(j)},\theta_k^{(j-1)}) \\
# \theta_k^{(j)} &\sim& \pi(\theta_k | \theta_1^{(j)},\theta_2^{(j)},\theta_4^{(j)},\ldots,\theta_{k-2}^{(j)},\theta_{k-1}^{(j)})\end{aligned}$$
#
# 4. Increment $j$ and repeat until convergence occurs.
#
# As we can see from the algorithm, each distribution is conditioned on the last iteration of its chain values, constituting a Markov chain as advertised. The Gibbs sampler has all of the important properties outlined in the previous section: it is aperiodic, homogeneous and ergodic. Once the sampler converges, all subsequent samples are from the target distribution. This convergence occurs at a geometric rate.
# ## Example: Inferring patterns in UK coal mining disasters
#
# Let's try to model a more interesting example, a time series of recorded coal mining
# disasters in the UK from 1851 to 1962.
#
# Occurrences of disasters in the time series is thought to be derived from a
# Poisson process with a large rate parameter in the early part of the time
# series, and from one with a smaller rate in the later part. We are interested
# in locating the change point in the series, which perhaps is related to changes
# in mining safety regulations.
# +
disasters_array = np.array([4, 5, 4, 0, 1, 4, 3, 4, 0, 6, 3, 3, 4, 0, 2, 6,
3, 3, 5, 4, 5, 3, 1, 4, 4, 1, 5, 5, 3, 4, 2, 5,
2, 2, 3, 4, 2, 1, 3, 2, 2, 1, 1, 1, 1, 3, 0, 0,
1, 0, 1, 1, 0, 0, 3, 1, 0, 3, 2, 2, 0, 1, 1, 1,
0, 1, 0, 1, 0, 0, 0, 2, 1, 0, 0, 0, 1, 1, 0, 2,
3, 3, 1, 1, 2, 1, 1, 1, 1, 2, 4, 2, 0, 0, 1, 4,
0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 1, 0, 1])
n_count_data = len(disasters_array)
# -
import plotly.plotly as py
import plotly.graph_objs as pgo
data = pgo.Data([
pgo.Scatter(
x=[str(year) + '-01-01' for year in np.arange(1851, 1962)],
y=disasters_array,
mode='lines+markers'
)
])
layout = pgo.Layout(
title='UK coal mining disasters (per year), 1851--1962',
xaxis=pgo.XAxis(title='Year', type='date', range=['1851-01-01', '1962-01-01']),
yaxis=pgo.YAxis(title='Disaster count')
)
fig = pgo.Figure(data=data, layout=layout)
py.iplot(fig, filename='coal_mining_disasters')
# We are going to use Poisson random variables for this type of count data. Denoting year $i$'s accident count by $y_i$,
#
# $$y_i \sim \text{Poisson}(\lambda).$$
#
# For those unfamiliar, Poisson random variables look like this:
data2 = pgo.Data([
pgo.Histogram(
x=np.random.poisson(l, 1000),
opacity=0.75,
name=u'λ=%i' % l
) for l in [1, 5, 12, 25]
])
layout_grey_bg = pgo.Layout(
xaxis=pgo.XAxis(zeroline=False, showgrid=True, gridcolor='rgb(255, 255, 255)'),
yaxis=pgo.YAxis(zeroline=False, showgrid=True, gridcolor='rgb(255, 255, 255)'),
paper_bgcolor='rgb(255, 255, 255)',
plot_bgcolor='rgba(204, 204, 204, 0.5)'
)
layout2 = layout_grey_bg.copy()
layout2.update(
barmode='overlay',
title='Poisson Means',
xaxis=pgo.XAxis(range=[0, 50]),
yaxis=pgo.YAxis(range=[0, 400])
)
fig2 = pgo.Figure(data=data2, layout=layout2)
py.iplot(fig2, filename='poisson_means')
# The modeling problem is about estimating the values of the $\lambda$ parameters. Looking at the time series above, it appears that the rate declines over time.
#
# A **changepoint model** identifies a point (here, a year) after which the parameter $\lambda$ drops to a lower value. Let us call this point in time $\tau$. So we are estimating two $\lambda$ parameters:
# $\lambda = \lambda_1$ if $t \lt \tau$ and $\lambda = \lambda_2$ if $t \geq \tau$.
#
# We need to assign prior probabilities to both $\{\lambda_1, \lambda_2\}$. The gamma distribution not only provides a continuous density function for positive numbers, but it is also *conjugate* with the Poisson sampling distribution.
lambda1_lambda2 = [(0.1, 100), (1, 100), (1, 10), (10, 10)]
data3 = pgo.Data([
pgo.Histogram(
x=np.random.gamma(*p, size=1000),
opacity=0.75,
name=u'α=%i, β=%i' % (p[0], p[1]))
for p in lambda1_lambda2
])
layout3 = layout_grey_bg.copy()
layout3.update(
barmode='overlay',
xaxis=pgo.XAxis(range=[0, 300])
)
fig3 = pgo.Figure(data=data3, layout=layout3)
py.iplot(fig3, filename='gamma_distributions')
# We will specify suitably vague hyperparameters $\alpha$ and $\beta$ for both priors:
#
# \begin{align}
# \lambda_1 &\sim \text{Gamma}(1, 10), \\
# \lambda_2 &\sim \text{Gamma}(1, 10).
# \end{align}
#
# Since we do not have any intuition about the location of the changepoint (unless we visualize the data), we will assign a discrete uniform prior over the entire observation period [1851, 1962]:
#
# \begin{align}
# &\tau \sim \text{DiscreteUniform(1851, 1962)}\\
# &\Rightarrow P(\tau = k) = \frac{1}{111}.
# \end{align}
# ### Implementing Gibbs sampling
#
# We are interested in estimating the joint posterior of $\lambda_1, \lambda_2$ and $\tau$ given the array of annnual disaster counts $\mathbf{y}$. This gives:
#
# $$
# P( \lambda_1, \lambda_2, \tau | \mathbf{y} ) \propto P(\mathbf{y} | \lambda_1, \lambda_2, \tau ) P(\lambda_1, \lambda_2, \tau)
# $$
#
# To employ Gibbs sampling, we need to factor the joint posterior into the product of conditional expressions:
#
# $$
# P(\lambda_1, \lambda_2, \tau | \mathbf{y}) \propto P(y_{t \lt \tau} | \lambda_1, \tau) P(y_{t \geq \tau} | \lambda_2, \tau) P(\lambda_1) P(\lambda_2) P(\tau)
# $$
#
# which we have specified as:
#
# $$\begin{aligned}
# P( \lambda_1, \lambda_2, \tau | \mathbf{y} ) &\propto \left[\prod_{t=1851}^{\tau} \text{Poi}(y_t|\lambda_1) \prod_{t=\tau+1}^{1962} \text{Poi}(y_t|\lambda_2) \right] \text{Gamma}(\lambda_1|\alpha,\beta) \text{Gamma}(\lambda_2|\alpha, \beta) \frac{1}{111} \\
# &\propto \left[\prod_{t=1851}^{\tau} e^{-\lambda_1}\lambda_1^{y_t} \prod_{t=\tau+1}^{1962} e^{-\lambda_2} \lambda_2^{y_t} \right] \lambda_1^{\alpha-1} e^{-\beta\lambda_1} \lambda_2^{\alpha-1} e^{-\beta\lambda_2} \\
# &\propto \lambda_1^{\sum_{t=1851}^{\tau} y_t +\alpha-1} e^{-(\beta+\tau)\lambda_1} \lambda_2^{\sum_{t=\tau+1}^{1962} y_i + \alpha-1} e^{-\beta\lambda_2}
# \end{aligned}$$
#
# So, the full conditionals are known, and critically for Gibbs, can easily be sampled from.
#
# $$\lambda_1 \sim \text{Gamma}(\sum_{t=1851}^{\tau} y_t +\alpha, \tau+\beta)$$
# $$\lambda_2 \sim \text{Gamma}(\sum_{t=\tau+1}^{1962} y_i + \alpha, 1962-\tau+\beta)$$
# $$\tau \sim \text{Categorical}\left( \frac{\lambda_1^{\sum_{t=1851}^{\tau} y_t +\alpha-1} e^{-(\beta+\tau)\lambda_1} \lambda_2^{\sum_{t=\tau+1}^{1962} y_i + \alpha-1} e^{-\beta\lambda_2}}{\sum_{k=1851}^{1962} \lambda_1^{\sum_{t=1851}^{\tau} y_t +\alpha-1} e^{-(\beta+\tau)\lambda_1} \lambda_2^{\sum_{t=\tau+1}^{1962} y_i + \alpha-1} e^{-\beta\lambda_2}} \right)$$
#
# Implementing this in Python requires random number generators for both the gamma and discrete uniform distributions. We can leverage NumPy for this:
# +
# Function to draw random gamma variate
rgamma = np.random.gamma
def rcategorical(probs, n=None):
# Function to draw random categorical variate
return np.array(probs).cumsum().searchsorted(np.random.sample(n))
# -
# Next, in order to generate probabilities for the conditional posterior of $\tau$, we need the kernel of the gamma density:
#
# \\[\lambda^{\alpha-1} e^{-\beta \lambda}\\]
dgamma = lambda lam, a, b: lam**(a - 1) * np.exp(-b * lam)
# Diffuse hyperpriors for the gamma priors on $\{\lambda_1, \lambda_2\}$:
alpha, beta = 1., 10
# For computational efficiency, it is best to pre-allocate memory to store the sampled values. We need 3 arrays, each with length equal to the number of iterations we plan to run:
# +
# Specify number of iterations
n_iterations = 1000
# Initialize trace of samples
lambda1, lambda2, tau = np.empty((3, n_iterations + 1))
# -
# The penultimate step initializes the model paramters to arbitrary values:
lambda1[0] = 6
lambda2[0] = 2
tau[0] = 50
# Now we can run the Gibbs sampler.
# Sample from conditionals
for i in range(n_iterations):
# Sample early mean
lambda1[i + 1] = rgamma(disasters_array[:tau[i]].sum() + alpha, 1./(tau[i] + beta))
# Sample late mean
lambda2[i + 1] = rgamma(disasters_array[tau[i]:].sum() + alpha,
1./(n_count_data - tau[i] + beta))
# Sample changepoint: first calculate probabilities (conditional)
p = np.array([dgamma(lambda1[i + 1], disasters_array[:t].sum() + alpha, t + beta) *
dgamma(lambda2[i + 1], disasters_array[t:].sum() + alpha, n_count_data - t + beta)
for t in range(n_count_data)])
# ... then draw sample
tau[i + 1] = rcategorical(p/p.sum())
# Plotting the trace and histogram of the samples reveals the marginal posteriors of each parameter in the model.
color = '#3182bd'
# +
trace1 = pgo.Scatter(
y=lambda1,
xaxis='x1',
yaxis='y1',
line=pgo.Line(width=1),
marker=pgo.Marker(color=color)
)
trace2 = pgo.Histogram(
x=lambda1,
xaxis='x2',
yaxis='y2',
line=pgo.Line(width=0.5),
marker=pgo.Marker(color=color)
)
trace3 = pgo.Scatter(
y=lambda2,
xaxis='x3',
yaxis='y3',
line=pgo.Line(width=1),
marker=pgo.Marker(color=color)
)
trace4 = pgo.Histogram(
x=lambda2,
xaxis='x4',
yaxis='y4',
marker=pgo.Marker(color=color)
)
trace5 = pgo.Scatter(
y=tau,
xaxis='x5',
yaxis='y5',
line=pgo.Line(width=1),
marker=pgo.Marker(color=color)
)
trace6 = pgo.Histogram(
x=tau,
xaxis='x6',
yaxis='y6',
marker=pgo.Marker(color=color)
)
# -
data4 = pgo.Data([trace1, trace2, trace3, trace4, trace5, trace6])
import plotly.tools as tls
fig4 = tls.make_subplots(3, 2)
fig4['data'] += data4
def add_style(fig):
for i in fig['layout'].keys():
fig['layout'][i]['zeroline'] = False
fig['layout'][i]['showgrid'] = True
fig['layout'][i]['gridcolor'] = 'rgb(255, 255, 255)'
fig['layout']['paper_bgcolor'] = 'rgb(255, 255, 255)'
fig['layout']['plot_bgcolor'] = 'rgba(204, 204, 204, 0.5)'
fig['layout']['showlegend']=False
add_style(fig4)
fig4['layout'].update(
yaxis1=pgo.YAxis(title=r'$\lambda_1$'),
yaxis3=pgo.YAxis(title=r'$\lambda_2$'),
yaxis5=pgo.YAxis(title=r'$\tau$'))
py.iplot(fig4, filename='modelling_params')
# ## The Metropolis-Hastings Algorithm
#
# The key to success in applying the Gibbs sampler to the estimation of Bayesian posteriors is being able to specify the form of the complete conditionals of
# ${\bf \theta}$, because the algorithm cannot be implemented without them. In practice, the posterior conditionals cannot always be neatly specified.
#
#
# Taking a different approach, the Metropolis-Hastings algorithm generates ***candidate*** state transitions from an alternate distribution, and *accepts* or *rejects* each candidate probabilistically.
#
# Let us first consider a simple Metropolis-Hastings algorithm for a single parameter, $\theta$. We will use a standard sampling distribution, referred to as the *proposal distribution*, to produce candidate variables $q_t(\theta^{\prime} | \theta)$. That is, the generated value, $\theta^{\prime}$, is a *possible* next value for
# $\theta$ at step $t+1$. We also need to be able to calculate the probability of moving back to the original value from the candidate, or
# $q_t(\theta | \theta^{\prime})$. These probabilistic ingredients are used to define an *acceptance ratio*:
#
# $$\begin{gathered}
# \begin{split}a(\theta^{\prime},\theta) = \frac{q_t(\theta^{\prime} | \theta) \pi(\theta^{\prime})}{q_t(\theta | \theta^{\prime}) \pi(\theta)}\end{split}\notag\\\begin{split}\end{split}\notag\end{gathered}$$
#
# The value of $\theta^{(t+1)}$ is then determined by:
#
# $$\theta^{(t+1)} = \left\{\begin{array}{l@{\quad \mbox{with prob.} \quad}l}\theta^{\prime} & \text{with probability } \min(a(\theta^{\prime},\theta^{(t)}),1) \\ \theta^{(t)} & \text{with probability } 1 - \min(a(\theta^{\prime},\theta^{(t)}),1) \end{array}\right.$$
#
# This transition kernel implies that movement is not guaranteed at every step. It only occurs if the suggested transition is likely based on the acceptance ratio.
#
# A single iteration of the Metropolis-Hastings algorithm proceeds as follows:
#
# 1. Sample $\theta^{\prime}$ from $q(\theta^{\prime} | \theta^{(t)})$.
#
# 2. Generate a Uniform[0,1] random variate $u$.
#
# 3. If $a(\theta^{\prime},\theta) > u$ then
# $\theta^{(t+1)} = \theta^{\prime}$, otherwise
# $\theta^{(t+1)} = \theta^{(t)}$.
#
# The original form of the algorithm specified by Metropolis required that
# $q_t(\theta^{\prime} | \theta) = q_t(\theta | \theta^{\prime})$, which reduces $a(\theta^{\prime},\theta)$ to
# $\pi(\theta^{\prime})/\pi(\theta)$, but this is not necessary. In either case, the state moves to high-density points in the distribution with high probability, and to low-density points with low probability. After convergence, the Metropolis-Hastings algorithm describes the full target posterior density, so all points are recurrent.
#
# ### Random-walk Metropolis-Hastings
#
# A practical implementation of the Metropolis-Hastings algorithm makes use of a random-walk proposal.
# Recall that a random walk is a Markov chain that evolves according to:
#
# $$
# \theta^{(t+1)} = \theta^{(t)} + \epsilon_t \\
# \epsilon_t \sim f(\phi)
# $$
#
# As applied to the MCMC sampling, the random walk is used as a proposal distribution, whereby dependent proposals are generated according to:
#
# $$\begin{gathered}
# \begin{split}q(\theta^{\prime} | \theta^{(t)}) = f(\theta^{\prime} - \theta^{(t)}) = \theta^{(t)} + \epsilon_t\end{split}\notag\\\begin{split}\end{split}\notag\end{gathered}$$
#
# Generally, the density generating $\epsilon_t$ is symmetric about zero,
# resulting in a symmetric chain. Chain symmetry implies that
# $q(\theta^{\prime} | \theta^{(t)}) = q(\theta^{(t)} | \theta^{\prime})$,
# which reduces the Metropolis-Hastings acceptance ratio to:
#
# $$\begin{gathered}
# \begin{split}a(\theta^{\prime},\theta) = \frac{\pi(\theta^{\prime})}{\pi(\theta)}\end{split}\notag\\\begin{split}\end{split}\notag\end{gathered}$$
#
# The choice of the random walk distribution for $\epsilon_t$ is frequently a normal or Student’s $t$ density, but it may be any distribution that generates an irreducible proposal chain.
#
# An important consideration is the specification of the **scale parameter** for the random walk error distribution. Large values produce random walk steps that are highly exploratory, but tend to produce proposal values in the tails of the target distribution, potentially resulting in very small acceptance rates. Conversely, small values tend to be accepted more frequently, since they tend to produce proposals close to the current parameter value, but may result in chains that ***mix*** very slowly.
#
# Some simulation studies suggest optimal acceptance rates in the range of 20-50%. It is often worthwhile to optimize the proposal variance by iteratively adjusting its value, according to observed acceptance rates early in the MCMC simulation .
# ## Example: Linear model estimation
#
# This very simple dataset is a selection of real estate prices \\(p\\), with the associated age \\(a\\) of each house. We wish to estimate a simple linear relationship between the two variables, using the Metropolis-Hastings algorithm.
#
# **Linear model**:
#
# $$\mu_i = \beta_0 + \beta_1 a_i$$
#
# **Sampling distribution**:
#
# $$p_i \sim N(\mu_i, \tau)$$
#
# **Prior distributions**:
#
# $$\begin{aligned}
# & \beta_i \sim N(0, 10000) \cr
# & \tau \sim \text{Gamma}(0.001, 0.001)
# \end{aligned}$$
# +
age = np.array([13, 14, 14,12, 9, 15, 10, 14, 9, 14, 13, 12, 9, 10, 15, 11,
15, 11, 7, 13, 13, 10, 9, 6, 11, 15, 13, 10, 9, 9, 15, 14,
14, 10, 14, 11, 13, 14, 10])
price = np.array([2950, 2300, 3900, 2800, 5000, 2999, 3950, 2995, 4500, 2800,
1990, 3500, 5100, 3900, 2900, 4950, 2000, 3400, 8999, 4000,
2950, 3250, 3950, 4600, 4500, 1600, 3900, 4200, 6500, 3500,
2999, 2600, 3250, 2500, 2400, 3990, 4600, 450,4700])/1000.
# -
# To avoid numerical underflow issues, we typically work with log-transformed likelihoods, so the joint posterior can be calculated as sums of log-probabilities and log-likelihoods.
#
# This function calculates the joint log-posterior, conditional on values for each parameter:
# +
from scipy.stats import distributions
dgamma = distributions.gamma.logpdf
dnorm = distributions.norm.logpdf
def calc_posterior(a, b, t, y=price, x=age):
# Calculate joint posterior, given values for a, b and t
# Priors on a,b
logp = dnorm(a, 0, 10000) + dnorm(b, 0, 10000)
# Prior on t
logp += dgamma(t, 0.001, 0.001)
# Calculate mu
mu = a + b*x
# Data likelihood
logp += sum(dnorm(y, mu, t**-2))
return logp
# -
# The `metropolis` function implements a simple random-walk Metropolis-Hastings sampler for this problem. It accepts as arguments:
#
# - the number of iterations to run
# - initial values for the unknown parameters
# - the variance parameter of the proposal distribution (normal)
# +
rnorm = np.random.normal
runif = np.random.rand
def metropolis(n_iterations, initial_values, prop_var=1):
n_params = len(initial_values)
# Initial proposal standard deviations
prop_sd = [prop_var]*n_params
# Initialize trace for parameters
trace = np.empty((n_iterations+1, n_params))
# Set initial values
trace[0] = initial_values
# Calculate joint posterior for initial values
current_log_prob = calc_posterior(*trace[0])
# Initialize acceptance counts
accepted = [0]*n_params
for i in range(n_iterations):
if not i%1000: print('Iteration %i' % i)
# Grab current parameter values
current_params = trace[i]
for j in range(n_params):
# Get current value for parameter j
p = trace[i].copy()
# Propose new value
if j==2:
# Ensure tau is positive
theta = np.exp(rnorm(np.log(current_params[j]), prop_sd[j]))
else:
theta = rnorm(current_params[j], prop_sd[j])
# Insert new value
p[j] = theta
# Calculate log posterior with proposed value
proposed_log_prob = calc_posterior(*p)
# Log-acceptance rate
alpha = proposed_log_prob - current_log_prob
# Sample a uniform random variate
u = runif()
# Test proposed value
if np.log(u) < alpha:
# Accept
trace[i+1,j] = theta
current_log_prob = proposed_log_prob
accepted[j] += 1
else:
# Reject
trace[i+1,j] = trace[i,j]
return trace, accepted
# -
# Let's run the MH algorithm with a very small proposal variance:
n_iter = 10000
trace, acc = metropolis(n_iter, initial_values=(1,0,1), prop_var=0.001)
# We can see that the acceptance rate is way too high:
np.array(acc, float)/n_iter
# +
trace1 = pgo.Scatter(
y=trace.T[0],
xaxis='x1',
yaxis='y1',
marker=pgo.Marker(color=color)
)
trace2 = pgo.Histogram(
x=trace.T[0],
xaxis='x2',
yaxis='y2',
marker=pgo.Marker(color=color)
)
trace3 = pgo.Scatter(
y=trace.T[1],
xaxis='x3',
yaxis='y3',
marker=pgo.Marker(color=color)
)
trace4 = pgo.Histogram(
x=trace.T[1],
xaxis='x4',
yaxis='y4',
marker=pgo.Marker(color=color)
)
trace5 = pgo.Scatter(
y=trace.T[2],
xaxis='x5',
yaxis='y5',
marker=pgo.Marker(color=color)
)
trace6 = pgo.Histogram(
x=trace.T[2],
xaxis='x6',
yaxis='y6',
marker=pgo.Marker(color=color)
)
# -
data5 = pgo.Data([trace1, trace2, trace3, trace4, trace5, trace6])
fig5 = tls.make_subplots(3, 2)
fig5['data'] += data5
add_style(fig5)
fig5['layout'].update(showlegend=False,
yaxis1=pgo.YAxis(title='intercept'),
yaxis3=pgo.YAxis(title='slope'),
yaxis5=pgo.YAxis(title='precision')
)
py.iplot(fig5, filename='MH algorithm small proposal variance')
# Now, with a very large proposal variance:
trace_hivar, acc = metropolis(n_iter, initial_values=(1,0,1), prop_var=100)
np.array(acc, float)/n_iter
# +
trace1 = pgo.Scatter(
y=trace_hivar.T[0],
xaxis='x1',
yaxis='y1',
marker=pgo.Marker(color=color)
)
trace2 = pgo.Histogram(
x=trace_hivar.T[0],
xaxis='x2',
yaxis='y2',
marker=pgo.Marker(color=color)
)
trace3 = pgo.Scatter(
y=trace_hivar.T[1],
xaxis='x3',
yaxis='y3',
marker=pgo.Marker(color=color)
)
trace4 = pgo.Histogram(
x=trace_hivar.T[1],
xaxis='x4',
yaxis='y4',
marker=pgo.Marker(color=color)
)
trace5 = pgo.Scatter(
y=trace_hivar.T[2],
xaxis='x5',
yaxis='y5',
marker=pgo.Marker(color=color)
)
trace6 = pgo.Histogram(
x=trace_hivar.T[2],
xaxis='x6',
yaxis='y6',
marker=pgo.Marker(color=color)
)
# -
data6 = pgo.Data([trace1, trace2, trace3, trace4, trace5, trace6])
fig6 = tls.make_subplots(3, 2)
fig6['data'] += data6
add_style(fig6)
fig6['layout'].update(
yaxis1=pgo.YAxis(title='intercept'),
yaxis3=pgo.YAxis(title='slope'),
yaxis5=pgo.YAxis(title='precision')
)
py.iplot(fig6, filename='MH algorithm large proposal variance')
# ### Adaptive Metropolis
#
# In order to avoid having to set the proposal variance by trial-and-error, we can add some tuning logic to the algorithm. The following implementation of Metropolis-Hastings reduces proposal variances by 10% when the acceptance rate is low, and increases it by 10% when the acceptance rate is high.
def metropolis_tuned(n_iterations, initial_values, f=calc_posterior, prop_var=1,
tune_for=None, tune_interval=100):
n_params = len(initial_values)
# Initial proposal standard deviations
prop_sd = [prop_var] * n_params
# Initialize trace for parameters
trace = np.empty((n_iterations+1, n_params))
# Set initial values
trace[0] = initial_values
# Initialize acceptance counts
accepted = [0]*n_params
# Calculate joint posterior for initial values
current_log_prob = f(*trace[0])
if tune_for is None:
tune_for = n_iterations/2
for i in range(n_iterations):
if not i%1000: print('Iteration %i' % i)
# Grab current parameter values
current_params = trace[i]
for j in range(n_params):
# Get current value for parameter j
p = trace[i].copy()
# Propose new value
if j==2:
# Ensure tau is positive
theta = np.exp(rnorm(np.log(current_params[j]), prop_sd[j]))
else:
theta = rnorm(current_params[j], prop_sd[j])
# Insert new value
p[j] = theta
# Calculate log posterior with proposed value
proposed_log_prob = f(*p)
# Log-acceptance rate
alpha = proposed_log_prob - current_log_prob
# Sample a uniform random variate
u = runif()
# Test proposed value
if np.log(u) < alpha:
# Accept
trace[i+1,j] = theta
current_log_prob = proposed_log_prob
accepted[j] += 1
else:
# Reject
trace[i+1,j] = trace[i,j]
# Tune every 100 iterations
if (not (i+1) % tune_interval) and (i < tune_for):
# Calculate aceptance rate
acceptance_rate = (1.*accepted[j])/tune_interval
if acceptance_rate<0.1:
prop_sd[j] *= 0.9
if acceptance_rate<0.2:
prop_sd[j] *= 0.95
if acceptance_rate>0.4:
prop_sd[j] *= 1.05
elif acceptance_rate>0.6:
prop_sd[j] *= 1.1
accepted[j] = 0
return trace[tune_for:], accepted
trace_tuned, acc = metropolis_tuned(n_iter*2, initial_values=(1,0,1), prop_var=5, tune_interval=25, tune_for=n_iter)
np.array(acc, float)/(n_iter)
# +
trace1 = pgo.Scatter(
y=trace_tuned.T[0],
xaxis='x1',
yaxis='y1',
line=pgo.Line(width=1),
marker=pgo.Marker(color=color)
)
trace2 = pgo.Histogram(
x=trace_tuned.T[0],
xaxis='x2',
yaxis='y2',
marker=pgo.Marker(color=color)
)
trace3 = pgo.Scatter(
y=trace_tuned.T[1],
xaxis='x3',
yaxis='y3',
line=pgo.Line(width=1),
marker=pgo.Marker(color=color)
)
trace4 = pgo.Histogram(
x=trace_tuned.T[1],
xaxis='x4',
yaxis='y4',
marker=pgo.Marker(color=color)
)
trace5 = pgo.Scatter(
y=trace_tuned.T[2],
xaxis='x5',
yaxis='y5',
line=pgo.Line(width=0.5),
marker=pgo.Marker(color=color)
)
trace6 = pgo.Histogram(
x=trace_tuned.T[2],
xaxis='x6',
yaxis='y6',
marker=pgo.Marker(color=color)
)
# -
data7 = pgo.Data([trace1, trace2, trace3, trace4, trace5, trace6])
fig7 = tls.make_subplots(3, 2)
fig7['data'] += data7
add_style(fig7)
fig7['layout'].update(
yaxis1=pgo.YAxis(title='intercept'),
yaxis3=pgo.YAxis(title='slope'),
yaxis5=pgo.YAxis(title='precision')
)
py.iplot(fig7, filename='adaptive-metropolis')
# 50 random regression lines drawn from the posterior:
# +
# Data points
points = pgo.Scatter(
x=age,
y=price,
mode='markers'
)
# Sample models from posterior
xvals = np.linspace(age.min(), age.max())
line_data = [np.column_stack([np.ones(50), xvals]).dot(trace_tuned[np.random.randint(0, 1000), :2]) for i in range(50)]
# Generate Scatter obejcts
lines = [pgo.Scatter(x=xvals, y=line, opacity=0.5, marker=pgo.Marker(color='#e34a33'),
line=pgo.Line(width=0.5)) for line in line_data]
data8 = pgo.Data([points] + lines)
layout8 = layout_grey_bg.copy()
layout8.update(
showlegend=False,
hovermode='closest',
xaxis=pgo.XAxis(title='Age', showgrid=False, zeroline=False),
yaxis=pgo.YAxis(title='Price', showline=False, zeroline=False)
)
fig8 = pgo.Figure(data=data8, layout=layout8)
py.iplot(fig8, filename='regression_lines')
# -
# ## Exercise: Bioassay analysis
#
# Gelman et al. (2003) present an example of an acute toxicity test, commonly performed on animals to estimate the toxicity of various compounds.
#
# In this dataset `log_dose` includes 4 levels of dosage, on the log scale, each administered to 5 rats during the experiment. The response variable is `death`, the number of positive responses to the dosage.
#
# The number of deaths can be modeled as a binomial response, with the probability of death being a linear function of dose:
#
# <div style="font-size: 150%;">
# $$\begin{aligned}
# y_i &\sim \text{Bin}(n_i, p_i) \\
# \text{logit}(p_i) &= a + b x_i
# \end{aligned}$$
# </div>
#
# The common statistic of interest in such experiments is the **LD50**, the dosage at which the probability of death is 50%.
#
# Use Metropolis-Hastings sampling to fit a Bayesian model to analyze this bioassay data, and to estimate LD50.
# +
# Log dose in each group
log_dose = [-.86, -.3, -.05, .73]
# Sample size in each group
n = 5
# Outcomes
deaths = [0, 1, 3, 5]
# +
from scipy.stats import distributions
dbin = distributions.binom.logpmf
dnorm = distributions.norm.logpdf
invlogit = lambda x: 1./(1 + np.exp(-x))
def calc_posterior(a, b, y=deaths, x=log_dose):
# Priors on a,b
logp = dnorm(a, 0, 10000) + dnorm(b, 0, 10000)
# Calculate p
p = invlogit(a + b*np.array(x))
# Data likelihood
logp += sum([dbin(yi, n, pi) for yi,pi in zip(y,p)])
return logp
# -
bioassay_trace, acc = metropolis_tuned(n_iter, f=calc_posterior, initial_values=(1,0), prop_var=5, tune_for=9000)
# +
trace1 = pgo.Scatter(
y=bioassay_trace.T[0],
xaxis='x1',
yaxis='y1',
marker=pgo.Marker(color=color)
)
trace2 = pgo.Histogram(
x=bioassay_trace.T[0],
xaxis='x2',
yaxis='y2',
marker=pgo.Marker(color=color)
)
trace3 = pgo.Scatter(
y=bioassay_trace.T[1],
xaxis='x3',
yaxis='y3',
marker=pgo.Marker(color=color)
)
trace4 = pgo.Histogram(
x=bioassay_trace.T[1],
xaxis='x4',
yaxis='y4',
marker=pgo.Marker(color=color)
)
# -
data9 = pgo.Data([trace1, trace2, trace3, trace4])
fig9 = tls.make_subplots(2, 2)
fig9['data'] += data9
add_style(fig9)
fig9['layout'].update(
yaxis1=pgo.YAxis(title='intercept'),
yaxis3=pgo.YAxis(title='slope')
)
py.iplot(fig9, filename='bioassay')
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install publisher --upgrade
import publisher
publisher.publish(
'montecarlo.ipynb', 'ipython-notebooks/computational-bayesian-analysis/',
'Computational Methods in Bayesian Analysis',
'Monte Carlo simulations, Markov chains, Gibbs sampling illustrated in Plotly',
name='Computational Methods in Bayesian Analysis')
# -
| _posts/ipython-notebooks/montecarlo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# 
#
# ---
# ## Introduction to Data Visualization in Python
#
# ### Matplotlib 101
#
# ---
#
#
# **Author list:** <NAME> & <NAME>
#
# **License Agreement:** Feel free to do whatever you want with this code
#
# ___
#
#
# Matplotlib is the standard plotting library in Python. It is well tested, well maintained and it has been around since ~2003. It also has a great support community.
#
# Matplotlib is highly customizable and sometimes it takes a lot of code to make a plot look like you want. It would take over a full week to go through all of the commands and aspects in matplotlib, therefore this notebook only works as a gentle introduciton.
#
# Many other plotting libraries (like `seaborn, pandas, bokeh, plotly, ` etc.) are built on top / integrates well with matplotlib. Therefore, it is an important package to know.
# ## Official gallery of matplotlib plot examples
# <div class='alert alert-info'>Please visit the <a href='https://matplotlib.org/gallery.html'>`matplotlib gallery`</a> and look at all the different plots you can make</div>
# # Load matplotlib
# +
# Standard way to import matplotlib from scratch: import matplotlib.pyplot as plt
# you might also see import pylab, however the procedure below is preferred
import matplotlib.pyplot as plt # always import pyplot module as plt (standard)
import numpy as np
import pandas as pd
# set seed
np.random.seed(0)
# add IPython magic command %matplotlib inline
# to print pyplot output inline in the notebook withput calling plt.show()
# %matplotlib inline
# Increase standard plot size
plt.rcParams['figure.figsize'] = (7, 4.5)
# -
# # Two plotting approaches
#
# There are two approaches to plotting the:
# 1. The functional / MATLAB approach
# 2. Object-oriented approach **(preferred)**
# # 1. Functional / MATLAB plotting
# Note that the command:
# ``` python
# plt.plot(x,y)
# ```
# will create a `figure object` ( the canvas that we plot on ) and an `axes object` that is the plot with lines and info.
plt.plot(); # creates empty plot, semi colon suppresses text output
# ## Simple line plot
# +
# Simple plot of two lists, note that they are of equal length
# plt.plot(x,y)
plt.plot([1,2,3],[-2,5,-8])
plt.show() # Note: we don't have to run plt.show()
# if we have run %matplotlib inline once in the kernel session
# -
# ## Plot 50 random numbers
# +
# every new cell where you run plt.plot() will create a new figure
N = 50
x = np.arange(1,N+1)
y = np.random.randn(N)
plt.plot(x,y,linestyle='--') # example without plt.plot(), note this does not work in scripts
# -
# # Two line plots in the same figure
# +
# if plt.plot(*args) are run in the same cell
# we will plot on the same axes
plt.plot(x,4*y)
plt.plot(x,np.abs(10*y)-x); # semi colon to suppress text output
# -
# #### New plot in one cell
# `plt.figure()` creates a new figure in the same cell
# +
# if plt.plot(*args) are run in the same cell
# we will plot on the same axes
plt.figure()
plt.plot(x,4*y)
plt.figure()
plt.plot(x,np.abs(10*y)-x);
# -
# # Scatter plot
# Relationship between two variables
plt.scatter(x,y);
# # Combine line and scatter plots
plt.plot(x,y)
plt.scatter(x,y);
plt.plot(x,y,marker='*');
# # Histograms
N=20
vals = np.random.randint(0,11,N)
print(sorted(vals))
plt.xlim(0,10) # change limits of x axis
plt.xticks(np.arange(11)) # change ticks on x-axis
plt.hist(vals);
# ### Histogram with bins
bins = np.linspace(0,10,6)
bins
# histogram with bins
#bins = np.linspace(0,10,5)
plt.hist(vals,bins=bins,width=1.9) # change width of bars, so we can see space in between
plt.xlim(0,10)
plt.show()
# # 2. Object-oriented approach (preferred approach)
# The object-oriented approach is more powerful and should be used when you want more control of your visualization. Instead of letting matplotlib handle what figure and axes that is the current one, we assign the `figure` and `axes` objects to variables and apply functions via methods on them.
# ## Figures and axes objects
# +
# simple plot
f, ax = plt.subplots() # returns tuple:
# f is the canvas object, can contain several plots i.e. axes objects (p)
ax.plot([1,2,3],[5,2,8]);
ax.hist(np.random.randint(1,4,10));
# -
# # Several subplots on the same Figure
# +
f, ax = plt.subplots(nrows=2,ncols=2)
# or
# f, (ax1, ax2, ax3, ax4) = plt.subplots(nrows=2,ncols=2)
# or,
# fig = plt.figure()
# ax1 = plt.subplot(221) #2x2 grid, first plot
# ax2 = plt.subplot(222)
# ax3 = plt.subplot(223)
# ax4 = plt.subplot(224)
# ax1.plot(x,y) etc..
ax[0,0].plot(x, y)
ax[1,0].scatter(x, y)
ax[0,1].hist(vals)
ax[1,1].barh(x,np.abs(y));
# -
# # Custom subplot grids
# +
# We can make arbitrarly complicated subplot grids with plt.subplot2grid
# example taken from Python Bootcamp https://github.com/profjsb/python-bootcamp
x = np.linspace(0,2*np.pi,10000)
y1 = np.sin(x)
y2 = np.sin(x)**2
f = plt.figure()
# subplot2grid(grid_shape, loc_in_grid, rowspan=1, colspan=1)
# 3 x 3 grid
# Specify the grid
ax1 = plt.subplot2grid((3,3), (0,0), colspan=3) # spans three cols
ax2 = plt.subplot2grid((3,3), (1,0), colspan=2) # 3
ax3 = plt.subplot2grid((3,3), (1,2), rowspan=2)
ax4 = plt.subplot2grid((3,3), (2,0))
ax5 = plt.subplot2grid((3,3), (2,1))
# remove tick labels on all axes
for ax in f.axes:
for t in ax.get_xticklabels()+ax.get_yticklabels():
t.set_visible(False)
# make a different kinds of plots!
ax1.plot(x, y1)
ax2.hist(y1)
ax3.scatter(x,y1)
ax4.boxplot(y1)
ax5.loglog(x, y1)
# Add titles
ax1.set_title('This')
ax3.set_title('plot')
ax5.set_title('works!')
f.suptitle('Grid layout!',fontsize=20, y=1.1) # y location
f.tight_layout() # great command for adding white space
# between plots
# -
# # 3. Plot styling
#
# ### Add labels, text, legend and change color
# +
# Three different line plots on same axes
N = 50
x = np.arange(1,N+1)
y = np.random.randn(N)
fig, ax = plt.subplots()
ax.plot(x,y,color='green', linestyle=':', label='line 1',linewidth = 2.5)
ax.plot(x,y*2,color='orange', linestyle='--', label = 'line 2')
ax.plot(x,y+4,color='black',marker='o', label = 'line 3')
ax.legend(loc=4) # location of legend is an integer,
# for text to be correct in the legend the plots need a label
# add plot LaTex style title, with line breaks
ax.set_title('3 Graphs (g): $\sum^3_i= g_i$\nsubtitle\n',
color='blue',fontweight='bold',fontsize=26)
ax.set_xlabel('x label') # add xlabel
ax.set_ylabel('y label'); # add ylabel
# -
# # Custom legend
# +
fig, ax = plt.subplots()
# ax.plot() returns a line object that we can assign to variables
# Note comma, in order to unpack tuple object
l1, = ax.plot(x,y,color='green', linestyle=':', label='line 1',linewidth = 2.5)
l2, = ax.plot(x,y*2,'y--', label = 'line 2')
l3, = ax.plot(x,y+4,color='black', linestyle='--',marker='o', label = 'line 3')
ax.legend(handles=[l1, l3], labels=['green vert', 'black dash dot'],
loc=[0.6,0.92],frameon=True,numpoints=3);
# -
# # Set limits and annotate values in the plot
# +
x = np.arange(0,10,2)
y = np.array([2,6,-4,3,-5])
fig, ax = plt.subplots()
ax.set_ylim(-6,8)
ax.set_xlim(0,9)
ax.plot(x,y)
for coord in zip(x, y):
ax.annotate(s = str(coord),xy = coord) # coord = coordinates
# -
# # Double axis plot
# +
fig, ax1 = plt.subplots()
N = 100 # number of samples
err=np.random.randn(N)*5
x=np.linspace(0,10,N)
y=12+x*3.5+err
y2 = np.sin(2 * np.pi * x)
# plot
ax1.set_xlim(0,10)
l1 = ax1.scatter(x,y,color='blue',label='scatter')
ax1.set_ylabel('line',color='blue')
ax1.tick_params('y',colors='blue')
ax2 = ax1.twinx() # create second axis for y2
l2, = ax2.plot(x, y2, color='red',label='$sin(x)$')
ax2.set_ylabel('sin', color='red')
ax1.set_xlim(0,10)
ax2.tick_params('y', colors='red')
plt.legend((l1,l2),('scatter','$sin(x)$'),loc=[0.8,1]);
# -
# # Plot sharing x-axis & Saving plots
# Generate data
x = np.linspace(0,2*np.pi,51)
y1 = np.sin(x)
y2 = 2*np.sin(x)**2
# +
# Create two subplots, on two rows with two axes ax1 and ax2
f, (ax1, ax2) = plt.subplots(2, 1,sharex=True)
#f, (ax1, ax2) = plt.subplots(1,2) # Would be vertical
ax1.plot(x,y1)
ax2.plot(x + np.pi,y2)
# show ticks
plt.xticks(np.linspace(0,3*np.pi,7),
('0','$\pi/2$','$\pi$','$3\pi/2$','$2\pi$','$5\pi/2$','$3\pi$'));
f.suptitle('Sharing x-axis'); # Title for both subplots
# -
# Save fig as png
f.savefig('fig.png') # easy, png, svg, pdf, etc work
f.savefig('fig.svg')
# it will take on the file format
# %ls # list files in your directory
# display picture
from IPython.display import Image
Image(filename='fig.png')
# display RANDOM image file
from IPython.display import Image
Image(filename='example.jpg')
# -----
# ## Opacity, marker size and color bar
# +
N=50
x = np.random.randn(N)
y = np.random.randn(N)
colors = np.random.rand(N)
sizes = N*10 * np.random.rand(100)
plt.scatter(x, y, c=colors, s=sizes, alpha=0.4, cmap='nipy_spectral')
plt.colorbar(); #this will add a color bar to the right
# -
# # 3d plot
# +
# Axes 3D needed to set projection='3d'
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
# Make data, spherical coordinates
u = np.linspace(0, .5*np.pi, 100)
v = np.linspace(0, 2*np.pi, 100)
x = 10 * np.outer(np.cos(u), np.sin(v))
y = 10 * np.outer(np.sin(u), np.sin(v))
z = 10 * np.outer(np.ones(np.size(u)), np.cos(v))
# Plot the surface
ax.plot_surface(x, y, z, color='r');
# -
# # Change the plot styles
#
# Here is a link to some common styles: [matplotlib style gallery](https://tonysyu.github.io/raw_content/matplotlib-style-gallery/gallery.html)
# +
# increase the figure size for all plots
IPython_default = plt.rcParams.copy() # save default styling
# +
data1 = np.random.randint(-10,5,400)
data2 = np.random.randint(-10,11,400)
data3 = np.random.randint(-10,11,400)
plt.style.use('ggplot') # for R lovers
plt.figure()
plt.hist([data1,data2,data3])
plt.title('ggplot')
plt.style.use('fivethirtyeight')
plt.figure()
plt.hist([data1,data2,data3])
plt.title('fivethirteight')
plt.style.use('dark_background')
plt.figure()
plt.hist([data1,data2,data3])
plt.title('dark-background')
plt.style.use('seaborn-white') # classic MATLAB stylingnot that pretty
plt.figure()
plt.hist([data1,data2,data3])
plt.title('seaborn-white (alex favorite!)');
# +
# You can also customize the
plt.rcParams.update(IPython_default); # restore defaults
# -
# # Seaborn and pandas
# **Note:** Anaconda does not come with seaborn, please install it in your virtual environment by running:
# ```bash
# conda intall seaborn
# ```
#
# - **seaborn:** Mostly used for statistical plotting, and better graphics.
# - **pandas:** Mostly used for quick plotting of DataFrames and time series
# +
import seaborn as sns
import pandas as pd
df = sns.load_dataset("iris") # load classic ML dataset
df.head()
# -
df.plot();
df.plot.box();
df.mean().plot.pie();
sp = list(df['species'].unique())
cols = list(['g','b','y'])
colors = dict(zip(sp,cols))
colors
df.plot.scatter('sepal_length','sepal_width',c=df['species'].apply(lambda x: colors[x]))
sns.pairplot(df, hue='species', size=2.5)
sns.distplot(df['sepal_length'],color='orange');
sns.boxplot(data=df);
# # Other packages you can check out
# Interactive plots
# - [Bokeh](https://bokeh.pydata.org/)
# - [Plotly](https://plot.ly/)
#
# Declarative plotting
# - [Altair](https://altair-viz.github.io/)
# - [Vega](https://vega.github.io/vega/)
| 04-AI-stack/matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python(oed)
# language: python
# name: oed
# ---
# """
# This script is used to caclulate the Event Mean Concentration (EMC).
# The inputs are .csv files containing concentration and flow after linear interpolation.
# """
# +
import pandas as pd
import numpy as np
from utils.concentration import rainfall_events, emc_cal, conc_interpolate, event_emc
import datetime
# read the discrete storm events
# Read daily loads and flow
# Read hourly loads and flow
from common_settings import obspath, outpath, events_name, \
obs_events, day_load_flow, hour_load_flow, conct_name, modpath, mod_load_flow
# -
from utils.concentration import cumulative_lq, excel_save
from utils.signatures import update_cumul_df, load_flow_loc
# ## Produce the event mean concentration of obs and mod
# Calculate EMC for low-frequency data
cols = [col for col in day_load_flow.columns if ('Load' in col) or ('Flow(ML)' in col)]
index_range1 = [1, 38]
index_range2 = [38, obs_events.shape[0]+1]
obs_events = event_emc(obs_events, day_load_flow, index_range1, cols[0], cols[1],
time_scale='d', multiplier=1e3)
# Calculate EMC for high-frequency data
cols = [col for col in hour_load_flow.columns if ('Load' in col) or ('ML' in col)]
index_range2 = [38, obs_events.shape[0]+1]
loads_col = cols[1]; flow_col = cols[0]
obs_events = event_emc(obs_events, hour_load_flow, index_range2, loads_col, flow_col,
time_scale='h', multiplier=1)
obs_events.to_csv(outpath + events_name, index='ID')
# +
# read the discrete storm events
filename = 'mod_storm_event_common.csv'
events = rainfall_events(f'{modpath}{filename}')
# Calculate EMC for modeling data
cols = [col for col in mod_load_flow.columns if ('Load' in col) or ('ML' in col)]
index_range = [1, events.shape[0]+1]
loads_col = cols[0]; flow_col = cols[1]
events = event_emc(events, mod_load_flow, index_range, loads_col, flow_col,
time_scale='d', multiplier=1)
events.dropna(axis=0, inplace=True)
events.to_csv(f'{outpath}DIN_{filename}', index='ID')
# -
# ## Produce the Normalized cumulative ratio of loads and flow
# ### calculate the daily data for double mass plot (Q-L)
time_ranges = [[f'{year}/7/1', f'{year+1}/6/30'] for year in range(2009, 2020)]
# time_ranges = obs_events.loc[:, ['start', 'end']].values
double_mass_ratio = {}
annual_total = pd.DataFrame(columns=day_load_flow.columns)
for ii in range(0, len(time_ranges)-2):
# for ii in range(index_range1[0]-1, index_range1[1]-1):
df_temp = load_flow_loc(time_ranges[ii], day_load_flow, timestep='d')
df_temp = update_cumul_df(df_temp, df_temp.values[:, 0], df_temp.values[:, -2])
double_mass_ratio[f'obs_year_{ii}'] = df_temp
annual_total.loc[time_ranges[ii][0][0:4]] = df_temp.sum(axis=0)
annual_total.loc['ave'] = annual_total.mean(axis=0)
# save outputs into one excel
annual_total.to_csv(outpath+'obs_annual_sum.csv')
fn = outpath +'obs_year_cumulative_ratio_day.xlsx'
excel_save(double_mass_ratio, fn, True)
# ### calculate the hourly data for double mass plot (Q-L)
double_mass_ratio = {}
for ii in range(index_range2[0]-1, index_range2[1]-1):
# for ii in range(9, len(time_ranges)):
df_temp = load_flow_loc(time_ranges[ii], hour_load_flow, timestep='h')
df_temp = update_cumul_df(df_temp, df_temp.values[:, -1], df_temp.values[:, 0])
double_mass_ratio[f'obs_storm_{ii}'] = df_temp
# save outputs into one excel
fn = outpath +'obs_storm_cumulative_ratio_hour.xlsx'
excel_save(double_mass_ratio, fn)
# ### calculate the modeling data for double mass plot (Q-L)
modpath = '../data/mod/'
annual_total = pd.DataFrame(columns=mod_load_flow.columns)
# filename = 'storm_event.csv'
# mod_events = rainfall_events(f'{modpath}{filename}')
# Calculate EMC for modeling data
cols = [col for col in mod_load_flow.columns if ('Load' in col) or ('ML' in col)]
# index_range = [1, mod_events.shape[0]]
loads_col = cols[0]; flow_col = cols[1]
double_mass_ratio = {}
time_ranges = [[f'{year}-07-01', f'{year+1}-06-30'] for year in range(2009, 2018)]
# time_ranges = mod_events.loc[:, ['start', 'end']].values
# for ii in range(index_range[0], index_range[1]):
for ii in range(len(time_ranges)):
df_temp = load_flow_loc(time_ranges[ii], mod_load_flow, timestep='d')
df_temp = update_cumul_df(df_temp, df_temp.values[:, 0], df_temp.values[:, -1])
double_mass_ratio[f'mod_storm_{ii}'] = df_temp
annual_total.loc[time_ranges[ii][0][0:4]] = df_temp.sum(axis=0)
annual_total.loc['ave'] = annual_total.mean(axis=0)
# +
# save results
annual_total.to_csv(outpath+'mod_annual_sum.csv')
fn = outpath +'mod_year_cumulative_ratio_day.xlsx'
excel_save(double_mass_ratio, fn)
# -
# ## Calculate event load coefficients
# ### Event loads for obs
obs_event_fn = 'obs_storm_event_common'
obs_events = pd.read_csv(f'{outpath}{obs_event_fn}.csv', index_col = 'ID')
time_ranges = [[f'{year}/7/1', f'{year+1}/6/30'] for year in [2009, 2010, 2011, 2012, 2013, 2014, 2018, 2019]]
year_loads = {}
# +
# for each year, calculate the yearly loads
# obs daily data
for tt in time_ranges[0:-2]:
df = load_flow_loc(tt, day_load_flow, timestep='d')
year_loads[tt[0][0:4]] = np.round(df.values[:, 0].sum(), 2)
# obs hourly data
for tt in time_ranges[-2:]:
df = load_flow_loc(tt, hour_load_flow, timestep='h')
year_loads[tt[0][0:4]] = np.round(df.values[:, 0].sum(), 2)
# +
# The event load coefficients
for ii in range(1, index_range1[1]):
df_event = load_flow_loc(obs_events.loc[ii, 'start':'end'].values, day_load_flow, timestep='d')
ymd= pd.to_datetime(obs_events.loc[ii, 'start'])
month = ymd.month; year = ymd.year
if month < 7:
obs_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / year_loads[str(year - 1)]
else:
obs_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / year_loads[str(year)]
for ii in range(index_range2[0], index_range2[1]):
df_event = load_flow_loc(obs_events.loc[ii, 'start':'end'].values, hour_load_flow, timestep='h')
ymd= pd.to_datetime(obs_events.loc[ii, 'start'])
month = ymd.month; year = ymd.year
if month < 7:
obs_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / year_loads[str(year-1)]
else:
obs_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / year_loads[str(year)]
# -
obs_events.to_csv(f'{outpath}{obs_event_fn}.csv')
# ### Event loads for mod
mod_event_fn = 'DIN_mod_storm_event_common'
mod_events = pd.read_csv(f'{outpath}{mod_event_fn}.csv', index_col = 'ID')
time_ranges = [[f'{year}/7/1', f'{year+1}/6/30'] for year in range(2009, 2014)]
# for each year, calculate the yearly loads
mod_loads = {}
# mod daily data
for tt in time_ranges:
df = load_flow_loc(tt, mod_load_flow, timestep='d')
mod_loads[tt[0][0:4]] = np.round(df.values[:, 0].sum(), 2)
# The event load coefficients
for ii in mod_events.index:
df_event = load_flow_loc(mod_events.loc[ii, 'start':'end'].values, mod_load_flow, timestep='d')
ymd= pd.to_datetime(mod_events.loc[ii, 'start'])
month = ymd.month; year = ymd.year
if month < 7:
mod_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / mod_loads[str(year - 1)]
else:
mod_events.loc[ii, 'event_load_coefficients'] = df_event.values[:, 0].sum() / mod_loads[str(year)]
mod_events.to_csv(f'{outpath}{mod_event_fn}.csv')
# ## Calculate the peaktime difference between flow and loads
# ### Mod results
mod_event_fn = 'DIN_mod_storm_event_common'
mod_events = pd.read_csv(f'{outpath}{mod_event_fn}.csv', index_col = 'ID')
# +
# find the peak time of loads
clabel, llabel, qlabel = ['Downstream Flow Concentration (mg.L^-1)', 'Loads (kg)', 'Flow_cumecs (ML.day^-1)']
for ii in mod_events.index:
df_event = load_flow_loc(mod_events.loc[ii, 'start':'end'].values, mod_load_flow, timestep='d')
peaktime_load = df_event[df_event.loc[:, llabel]==df_event.loc[:, llabel].max()].index
peaktime_conc = df_event[df_event.loc[:, clabel]==df_event.loc[:, clabel].max()].index[0]
mod_events.loc[ii, 'peaktime_load'] = peaktime_load
mod_events.loc[ii, 'peaktime_conc'] = peaktime_conc
mod_events.loc[ii, 'peakflow'] = df_event.loc[:, qlabel].max()
mod_events.loc[ii, 'peaktime'] = df_event[df_event.loc[:, qlabel]==df_event.loc[:, qlabel].max()].index
mod_events.loc[ii, 'peak_flow_time'] = df_event[df_event.loc[:, qlabel]==df_event.loc[:, qlabel].max()].index[0]
mod_events.loc[ii, 'peak_conc_time'] = df_event[df_event.loc[:, clabel]==df_event.loc[:, clabel].max()].index[0]
mod_events.loc[ii, 'peak_load_time'] = df_event[df_event.loc[:, llabel]==df_event.loc[:, llabel].max()].index[0]
# mod_events.loc[:, 'delta_time'] = mod_events.peaktime_load - mod_events.peaktime
# -
mod_events.loc[:, 'delta_load_flow_time'] = pd.to_datetime(mod_events.peak_load_time) - pd.to_datetime(mod_events.peak_flow_time)
mod_events.loc[:, 'delta_conc_flow_time'] = pd.to_datetime(mod_events.peak_conc_time) - pd.to_datetime(mod_events.peak_flow_time)
mod_events.to_csv(f'{outpath}{mod_event_fn}.csv')
# ### Obs results
obs_event_fn = 'obs_storm_event_common'
obs_events = pd.read_csv(f'{outpath}{obs_event_fn}.csv', index_col = 'ID')
# +
# find the peak time of loads
for ii in obs_events.index:
if ii < 38:
df_event = load_flow_loc(obs_events.loc[ii, 'start':'end'].values, day_load_flow, timestep='d')
clabel, llabel, qlabel = ['Concentration (mg/L)', 'Linear_Average_Load(t)', 'Flow(ML)']
else:
clabel, llabel, qlabel = ['126001A-NO3(mg/l)', 'Loads (kg)', 'Flow (ML)']
df_event = load_flow_loc(obs_events.loc[ii, 'start':'end'].values, hour_load_flow, timestep='h')
peaktime_load = df_event[df_event.loc[:, llabel]==df_event.loc[:, llabel].max()].index[0]
peaktime_conc = df_event[df_event.loc[:, clabel]==df_event.loc[:, clabel].max()].index[0]
obs_events.loc[ii, 'peaktime_load'] = peaktime_load
obs_events.loc[ii, 'peaktime_conc'] = peaktime_conc
obs_events.loc[ii, 'peakflow'] = df_event.loc[:, qlabel].max()
obs_events.loc[ii, 'peak_flow_time'] = df_event[df_event.loc[:, qlabel]==df_event.loc[:, qlabel].max()].index[0]
obs_events.loc[ii, 'peak_conc_time'] = df_event[df_event.loc[:, clabel]==df_event.loc[:, clabel].max()].index[0]
obs_events.loc[ii, 'peak_load_time'] = df_event[df_event.loc[:, llabel]==df_event.loc[:, llabel].max()].index[0]
obs_events.loc[:, 'delta_load_flow_time'] = pd.to_datetime(obs_events.peak_load_time) - pd.to_datetime(obs_events.peak_flow_time)
obs_events.loc[:, 'delta_conc_flow_time'] = pd.to_datetime(obs_events.peak_conc_time) - pd.to_datetime(obs_events.peak_flow_time)
# -
obs_events.to_csv(f'{outpath}{obs_event_fn}.csv')
# ## Variability of load-discharge ratio (seasonal average concentration)
# ### Obs results
time_ranges = [[f'{year}/7/1', f'{year}/10/1', f'{year+1}/1/1', f'{year+1}/4/1', f'{year+1}/7/1'] for year in range(2009, 2018)]
df_ratio = pd.DataFrame(index=[str(year) for year in range(2009, 2018)], columns = [1, 2, 3, 4])
for tt in time_ranges:
for ii in range(len(tt) -1):
start = pd.to_datetime(tt[ii])
end = pd.to_datetime(tt[ii + 1]) - datetime.timedelta(days=1)
df = load_flow_loc([start, end], day_load_flow, timestep ='d')
df_ratio.loc[tt[0][0:4], ii+1] = df.sum(axis=0)[0] / df.sum(axis=0)[2] * 1000
df_ratio.to_csv(f'{outpath}obs_seasonal_concentration.csv')
# ### Mod results
time_ranges = [[f'{year}/7/1', f'{year}/10/1', f'{year+1}/1/1', f'{year+1}/4/1', f'{year+1}/7/1'] for year in range(2009, 2018)]
df_ratio = pd.DataFrame(index=[str(year) for year in range(2009, 2018)], columns = [1, 2, 3, 4])
for tt in time_ranges:
for ii in range(len(tt) -1):
start = pd.to_datetime(tt[ii])
end = pd.to_datetime(tt[ii + 1]) - datetime.timedelta(days=1)
df = load_flow_loc([start, end], mod_load_flow, timestep ='d')
df_ratio.loc[tt[0][0:4], ii+1] = df.sum(axis=0)[0] / df.sum(axis=0)[2]
df_ratio.to_csv(f'{outpath}mod_seasonal_concentration.csv')
# ## Monthly loads
df_month = pd.DataFrame(columns = ['obs', 'mod'])
# calculate the monthly loads and flow
for y in range(2009, 2019):
for m in range(1, 13):
start = pd.to_datetime(f'{y}/{m}/1')
if m == 12:
end = pd.to_datetime(f'{y+1}/1/1') - datetime.timedelta(days=1)
else:
end = pd.to_datetime(f'{y}/{m+1}/1') - datetime.timedelta(days=1)
df_month.loc[f'{y}/{m}', 'obs'] = 1000 * load_flow_loc([start, end], day_load_flow, timestep ='d').sum(axis=0)[0]
df_month.loc[f'{y}/{m}', 'mod'] = load_flow_loc([start, end], mod_load_flow, timestep ='d').sum(axis=0)[0]
df_month = df_month[(df_month.obs != 0) & (df_month.loc[:, 'mod'] != 0)]
df_month.index.name = 'Month'
df_month.to_csv(f'{outpath}mod_obs_month.csv')
# ## Calculate the coefficients of variation for concentrations (CVC) and discharge (CVQ), their ratio (CVC:CVQ)
# define timeperiod
time_ranges = pd.to_datetime(['2009/7/1', '2018/6/30'])
df_cv = pd.DataFrame(columns=['obs', 'mod'], index=['cvc', 'cvq', 'cvl'])
start, end = time_ranges
# read obs time series of flow, loads and concentration
df_obs = load_flow_loc([start, end], day_load_flow, timestep ='d')
cv_all = (df_obs.std(axis=0) / df_obs.mean(axis=0))
cols = df_obs.columns
df_cv.loc[:, 'obs'] = [cv_all[cols[3]], cv_all[cols[2]], cv_all[cols[0]]]
# +
# read mod time series of flow, loads and concentration
df_mod = load_flow_loc([start, end], mod_load_flow, timestep ='d')
cv_all = (df_mod.std(axis=0) / df_mod.mean(axis=0))
cols = df_mod.columns
df_cv.loc[:, 'mod'] = [cv_all[cols[1]], cv_all[cols[2]], cv_all[cols[0]]]
df_cv.loc['cq', :] = df_cv.loc['cvc', :] / df_cv.loc['cvq', :]
df_cv.loc['lq', :] = df_cv.loc['cvl', :] / df_cv.loc['cvq', :]
# -
df_cv.to_csv(f'{outpath}cv_cql.csv')
# ## Linear regression of C-Q
# import necessary packages
from sklearn.metrics import r2_score
from utils.signatures import residual, nonlinear_fit
from utils.plotting import regression_plot
import lmfit
# define x and y
time_range = [['/7/1', '/10/1'], ['/10/1', '/1/1'], ['/1/1', '/4/1'], ['/4/1', '/7/1']]
cols = mod_load_flow.columns
# day_load_flow.loc[:, cols[0]] = day_load_flow.loc[:, cols[0]]*1000
x_dict, y_dict = {}, {}
k = 1
for tt in time_range:
x, y = np.array([]), np.array([])
for year in range(2009, 2019):
start = pd.to_datetime(f'{year}{tt[0]}')
if tt[1] == '/1/1':
end = pd.to_datetime(f'{year+1}{tt[1]}') - datetime.timedelta(days=1)
else:
end = pd.to_datetime(f'{year}{tt[1]}') - datetime.timedelta(days=1)
df_temp = load_flow_loc([start, end], mod_load_flow, timestep ='d')
x = np.append(x, df_temp.values[:, 2])
y = np.append(y, df_temp.values[:, 0])
x_dict[f'{k}_x'] = x
x_dict[f'{k}_y'] = y
k += 1
# + jupyter={"outputs_hidden": true}
# variables are x and y
coeff_regress = pd.DataFrame(columns = np.arange(1, 5), index=['R2', 'a', 'b', 'c'])
for k in range(1, 5):
x = x_dict[f'{k}_x']
y = x_dict[f'{k}_y']
p = lmfit.Parameters()
p.add_many(('a', 0.1, True, 0, 10), ('b', 2, True, 0, 2), ('c', 1, True, 0, 10))
out1, out2, ci, trace = nonlinear_fit(p, residual, x, y, opti_method='differential_evolution')# lmfit, x=x_input, y=y_output,
# compare coefficient of determination
para_values = {}
for param in ['a', 'b', 'c']:
para_values[param] = np.round(trace['a'][param][0], 4)
y_mod = para_values['a'] * x ** para_values['b']+ para_values['c']
r2 = r2_score(np.log(y), np.log(y_mod))
abs_bias = np.abs(np.average(y_mod - y))
rel_bias = abs_bias / np.average(y)
coeff_regress.loc[:, k] = [r2, para_values['a'], para_values['b'], para_values['c']]
# -
coeff_regress.to_csv(outpath+'mod_cq_regress.csv')
# ## Calculate the variation of delivery ratio-surface
# read observations, modeled outputs with delivery ratio at 0 and 25 (%)
mod_fl_fn = 'DIN_sources.csv'
mod_drs0 = pd.read_csv(modpath + mod_fl_fn, index_col='Date')
mod_drs0.index = pd.to_datetime(mod_drs0.index, dayfirst=False)
arrays = [['mod_total','mod_total', 'mod_total', 'mod_total', 'surface', 'surface', 'surface', 'surface', 'seepage', 'seepage', 'seepage', 'seepage', 'DWC', 'DWC', 'DWC', 'DWC',
'dwc_const', 'dwc_const', 'dwc_const', 'dwc_const', 'emc_const', 'emc_const', 'emc_const', 'emc_const', 'obs', 'obs', 'obs', 'obs'],
['median', 'min', 'max', 'mean', 'median', 'min', 'max', 'mean', 'median', 'min', 'max', 'mean', 'median', 'min', 'max', 'mean',
'median', 'min', 'max', 'mean', 'median', 'min', 'max', 'mean', 'median', 'min', 'max', 'mean']]
tuples = list(zip(*arrays))
index = pd.MultiIndex.from_tuples(tuples)
df_month = pd.DataFrame(columns=mod_drs0.columns)
# calculate the monthly loads and flow
for y in range(2009, 2019):
for m in range(1, 13):
start = pd.to_datetime(f'{y}/{m}/1')
if m == 12:
# import pdb; pdb.set_trace()
end = pd.to_datetime(f'{y+1}/1/1') - datetime.timedelta(days=1)
else:
end = pd.to_datetime(f'{y}/{m+1}/1') - datetime.timedelta(days=1)
#
df_month.loc[f'{y}/{m}', :] = load_flow_loc([start, end], mod_drs0, timestep ='d').sum(axis=0)
df_month = df_month[(df_month.loc[:, df_month.columns[0]] != 0)]
df_month.index.name = 'Month'
df_month.to_csv(f'{outpath}month_loads_sources.csv')
for col in df_month.columns:
df_month[col+'_ratio'] = (df_month['obs'] - df_month['mod_all'] + df_month[col]) / df_month[col]
# df_month['drs'] = (df_month['obs'] - df_month['mod_all'] + df_month[col]) / df_month[col]
df_month.head()
df_month.to_csv(outpath+'obs_mod_month_load_ratio.csv')
x_list = [*np.arange(7, 13), *np.arange(1, 7)]
drs_stats = pd.DataFrame(columns=x_list, index=index)
drs_stats.index.name = 'month'
col_ratio = df_month.columns[0:7]
for col in col_ratio:
k = list(col_ratio).index(col)
for i in range(12):
drs_stats.loc[index[(4 * k) : (4 * k + 4)], x_list[i]] = df_month[col][i::12].median(), df_month[col][i::12].min(), df_month[col][i::12].max(), df_month[col][i::12].mean()
# obs_mod = (df_month['obs'][i::12].mean() + df_month[df_month.columns[k+1]][i::12].mean() - df_month['mod_all'][i::12].mean())
# mod_mod0 = df_month[df_month.columns[k+1]][i::12].mean()
# drs_stats.loc[index[4 * k + 3], x_list[i]] = obs_mod / mod_mod0
# drs_stats.to_csv(outpath+'DeliveryRatioSurface.csv')
drs_stats.to_csv(outpath+'month_sum_loads.csv')
| src/signatures_produce.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <blockquote>
# <h1>Exercise 4.10</h1>
# <p>This question should be answered using the <code>Weekly</code> data set, which is part of the <code>ISLR</code> package. This data is similar in nature to the <code>Smarket</code> data from this chapter’s lab, except that it contains $1,089$ weekly returns for $21$ years, from the beginning of $1990$ to the end of $2010$.</p>
# <ol>
# <li>Produce some numerical and graphical summaries of the <code>Weekly</code> data. Do there appear to be any patterns?</li>
# <li>Use the full data set to perform a logistic regression with $\mathrm{Direction}$ as the response and the five lag variables plus $\mathrm{Volume}$ as predictors. Use the summary function to print the results. Do any of the predictors appear to be statistically significant? If so, which ones?</li>
# <li>Compute the confusion matrix and overall fraction of correct predictions. Explain what the confusion matrix is telling you about the types of mistakes made by logistic regression.</li>
# <li>Now fit the logistic regression model using a training data period from $1990$ to $2008$, with $\mathrm{Lag2}$ as the only predictor. Compute the confusion matrix and the overall fraction of correct predictions for the held out data (that is, the data from $2009$ and $2010$).</li>
# <li>Repeat 4 using LDA.</li>
# <li>Repeat 4 using QDA.</li>
# <li>Repeat 4 using KNN with $K = 1$.</li>
# <li>Which of these methods appears to provide the best results on this data?</li>
# <li>Experiment with different combinations of predictors, including possible transformations and interactions, for each of the methods. Report the variables, method, and associated confusion matrix that appears to provide the best results on the held out data. Note that you should also experiment with values for $K$ in the KNN classifier.</li>
# </ol>
# </blockquote>
# +
import pandas as pd
import numpy as np
# %run ../../customModules/usefulFunctions.ipynb
# https://stackoverflow.com/questions/34398054/ipython-notebook-cell-multiple-outputs
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import statsmodels.api as sm
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.neighbors import KNeighborsClassifier
# -
# <h3>Exercise 4.10.1</h3>
# <blockquote>
# <i>Produce some numerical and graphical summaries of the <code>Weekly</code> data. Do there appear to be any patterns?</i>
# </blockquote>
#
# <p>These summaries can be found in the <a href="../../DataSets/Weekly/Exploration.ipynb">Exploration</a> notebook. The $\mathrm{Today}$, $\mathrm{Lag1}$, $\mathrm{Lag2}$, $\mathrm{Lag3}$, $\mathrm{Lag4}$ and $\mathrm{Lag5}$ variables have approximately the same summary statistics which makes sense because overall they all have the same values except for the first and last $5$ samples. Let us in addition produce a scatterplot matrix of all of the variables in the data set.</p>
df = pd.read_csv("../../DataSets/Weekly/Weekly.csv")
df = df.reindex(columns=['Year', 'Today', 'Lag1', 'Lag2', 'Lag3', 'Lag4', 'Lag5', 'Volume', 'Direction'])
df.head()
_ = pd.plotting.scatter_matrix(df, figsize=(12, 12))
# <p>The only clear pattern is between the $\mathrm{Volumne}$ and $\mathrm{Year}$ variables.</p>
#
# <h3>Exercise 4.10.2</h3>
# <blockquote>
# <i>Use the full data set to perform a logistic regression with $\mathrm{Direction}$ as the response and the five lag variables plus $\mathrm{Volume}$ as predictors. Use the summary function to print the results. Do any of the predictors appear to be statistically significant? If so, which ones?</i>
# </blockquote>
df.set_index('Year', inplace=True)
df['Direction'] = np.where(df['Direction'] == 'Up', 1, 0)
df.insert(0, 'Intercept', 1)
targetColumn = ['Direction']
descriptiveColumns = removeColumnsFromList(df, targetColumn + ['Today'])
df_X = df[descriptiveColumns]
df_Y = df[targetColumn]
model = sm.Logit(df_Y, df_X)
fitted = model.fit()
fitted.summary()
# <p>We see that, with a significance level of $\alpha=10 \%$, the $\mathrm{Lag2}$ is the only variable indicating statistical significance.</p>
#
# <h3>Exercise 4.10.3</h3>
# <blockquote>
# <i>Compute the confusion matrix and overall fraction of correct predictions. Explain what the confusion matrix is telling you about the types of mistakes made by logistic regression.</i>
# </blockquote>
#
# <p>We use our custom <code>createConfusionMatrixFromLogisticModel</code> method to compute the confusion matrix and its associated percentages.</p>
df_confusion, df_confusion_pct = createConfusionMatrixFromLogisticModel(fitted_model=fitted, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
# <p>This immediately suggests that the model is better at predicting the $\mathrm{Up}$ direction than it is at preciting the $\mathrm{Down}$ direction. That is because the model predicts $\mathrm{Up}$ a lot more than $\mathrm{Down}$. To be precise, $(430 + 557)/(430 + 557 + 54 + 48) = 90.63 \%$ of the total predictions are $\mathrm{Up}$. Furthermore, let us calculate classification accuracy
# $$
# \mathrm{classification \,\, rate} = \frac{\mathrm{TP} + \mathrm{TN}}{\mathrm{TP} + \mathrm{TN} + \mathrm{FP} + \mathrm{FN}} = 56.11 \% \,,
# $$
# where $\mathrm{TN}$, $\mathrm{FP}$, $\mathrm{FN}$ and $\mathrm{TP}$ denotes the true negative, false positive, false negative and true positive, respectively.</p>
#
# <h3>Exercise 4.10.4</h3>
# <blockquote>
# <i>Now fit the logistic regression model using a training data period from $1990$ to $2008$, with $\mathrm{Lag2}$ as the only predictor. Compute the confusion matrix and the overall fraction of correct predictions for the held out data (that is, the data from $2009$ and $2010$).</i>
# </blockquote>
# +
targetColumn = ['Direction']
descriptiveColumns = ['Intercept', 'Lag2']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
# -
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
# <p>The classification accuracy is $62.5 \%$.</p>
#
# <h3>Exercise 4.10.5</h3>
# <blockquote>
# <i>Repeat 4 using LDA.</i>
# </blockquote>
# +
targetColumn = ['Direction']
descriptiveColumns = ['Lag2']
X_train = np.squeeze(df[descriptiveColumns].loc[1990:2008].to_numpy()).reshape(-1, 1)
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
# -
# <p>These confusion matrices are exactly the same as the logistic regression has yielded in subquestion 4.10.4, and so the classification accuracy is again $62.5 \%$.</p>
#
# <h3>Exercise 4.10.6</h3>
# <blockquote>
# <i>Repeat 4 using QDA.</i>
# </blockquote>
# +
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
# -
# <p>This model predicts every observation to be $\mathrm{Up}$. The classification accuracy is $58.65 \%$.</p>
#
# <h3>Exercise 4.10.7</h3>
# <blockquote>
# <i>Repeat 4 using KNN with $K = 1$.</i>
# </blockquote>
# +
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
# -
# <p>The classification accuracy is $49.04 \%$.</p>
#
# <h3>Exercise 4.10.8</h3>
# <blockquote>
# <i>Which of these methods appears to provide the best results on this data?</i>
# </blockquote>
#
# <p>Based on the classification accuracy of each of these model, the logistic regression and LDA methods have the best predictive power.</p>
#
# <h3>Exercise 4.10.9</h3>
# <blockquote>
# <i>Experiment with different combinations of predictors, including possible transformations and interactions, for each of the methods. Report the variables, method, and associated confusion matrix that appears to provide the best results on the held out data. Note that you should also experiment with values for $K$ in the KNN classifier.</i>
# </blockquote>
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag1']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag2']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
descriptiveColumns = ['Volume', 'Lag1', 'Lag2']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
df['Lag1*Lag2'] = df['Lag1']*df['Lag2']
descriptiveColumns = ['Volume', 'Lag1', 'Lag2', 'Lag1*Lag2']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
df['Volume*Lag1'] = df['Volume']*df['Lag1']
descriptiveColumns = ['Volume', 'Lag1', 'Lag2', 'Volume*Lag1']
df_X_train = df[descriptiveColumns].loc[1990:2008]
df_Y_train = df[targetColumn].loc[1990:2008]
df_X_test = df[descriptiveColumns].loc[2009:2010]
df_Y_test = df[targetColumn].loc[2009:2010]
model = sm.Logit(df_Y_train, df_X_train)
fitted = model.fit()
fitted.summary()
sr_Y_pred = fitted.predict(df_X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': np.where(sr_Y_pred > 0.5, 1, 0),
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag1']
X_train = df[descriptiveColumns].loc[1990:2008].to_numpy()
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=2)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=3)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag2']
X_train = df[descriptiveColumns].loc[1990:2008].to_numpy()
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=2)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=3)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag1', 'Lag2']
X_train = df[descriptiveColumns].loc[1990:2008].to_numpy()
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=2)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=3)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag1', 'Lag2', 'Lag1*Lag2']
X_train = df[descriptiveColumns].loc[1990:2008].to_numpy()
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=2)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=3)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
# +
targetColumn = ['Direction']
descriptiveColumns = ['Volume', 'Lag1', 'Lag2', 'Volume*Lag1']
X_train = df[descriptiveColumns].loc[1990:2008].to_numpy()
Y_train = np.squeeze(df[targetColumn].loc[1990:2008].to_numpy())
X_test = df[descriptiveColumns].loc[2009:2010].to_numpy()
model = LinearDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = QuadraticDiscriminantAnalysis()
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=1)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=2)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
'-------------------'
model = KNeighborsClassifier(n_neighbors=3)
_ = model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
df_Y_test_and_pred = pd.DataFrame({
'Observed': df_Y_test['Direction'],
'Predicted': Y_pred,
})
df_confusion, df_confusion_pct = createConfusionMatrixFromOutOfSampleData(df=df_Y_test_and_pred, binaryMap={0: 'Down', 1: 'Up'})
df_confusion
df_confusion_pct.round(2)
confusion_matrix = df_confusion.to_numpy()
TN, FP, FN, TP = confusion_matrix[0, 0], confusion_matrix[0, 1], confusion_matrix[1, 0], confusion_matrix[1, 1]
class_acc = 100 * ((TP + TN) / (TN + FP + FN + TP))
f'The classification accuracy is {class_acc:.2f}%.'
| Chapter04/Exercise10/4_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="c3T_vo6GYvFD"
# ## 5.3 季節など周期性で売り上げ予測(時系列分析)
# + [markdown] colab_type="text" id="ZO7cPPEWaA2u"
# ### 共通事前準備
# + colab={} colab_type="code" id="uOs6nswAbY9n"
# 日本語化ライブラリ導入
# !pip install japanize-matplotlib | tail -n 1
# + colab={} colab_type="code" id="7qgmy1x_gez9"
# 共通事前処理
# 余分なワーニングを非表示にする
import warnings
warnings.filterwarnings('ignore')
# 必要ライブラリのimport
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# matplotlib日本語化対応
import japanize_matplotlib
# データフレーム表示用関数
from IPython.display import display
# 表示オプション調整
# numpyの浮動小数点の表示精度
np.set_printoptions(suppress=True, precision=4)
# pandasでの浮動小数点の表示精度
pd.options.display.float_format = '{:.4f}'.format
# データフレームですべての項目を表示
pd.set_option("display.max_columns",None)
# グラフのデフォルトフォント指定
plt.rcParams["font.size"] = 14
# 乱数の種
random_seed = 123
# + [markdown] colab_type="text" id="rFsr2f59bY9t"
# オリジナルURL
# https://archive.ics.uci.edu/ml/datasets/bike+sharing+dataset
# + [markdown] colab_type="text" id="BxV4nTzNbY9t"
# #### データ項目メモ
#
# instant インデックス
# dteday 日付(yy-mm-dd)
# season 季節 (1: 冬 2: 春 3: 夏 4:秋)
# yr 年 (0: 2011, 1:2012)
# mnth 月 (1 - 12)
# hr 時間 (0 - 23)
# holiday 祝日
# weekday 曜日 (0 - 6)
# workingday 勤務日 (1: 勤務日 0: 休日)
# weathersit 天気 (1: 晴れから曇り 2: 霧 3: 小雨 4: 大雨)
# temp 気温 (正規化済み)
# atemp 体感気温 (正規化済み)
# hum 湿度 (正規化済み)
# windspeed 風速 (正規化済み)
# casual 臨時ユーザー利用数
# registered 登録ユーザー利用数
# cnt 全体ユーザー利用数
# + [markdown] colab_type="text" id="HMptNeWrbY9t"
# ### 5.3.4 データ読み込みからデータ確認まで
# + [markdown] colab_type="text" id="8_OCPaWVge0B"
# #### データ読み込み
# + colab={} colab_type="code" id="yIg1xgb-bY9u"
# ダウンロード元URL
url = 'https://archive.ics.uci.edu/ml/\
machine-learning-databases/00275/\
Bike-Sharing-Dataset.zip'
# 公開データのダウンロードと解凍
# !wget $url -O Bike-Sharing-Dataset.zip | tail -n 1
# !unzip -o Bike-Sharing-Dataset.zip | tail -n 1
# + colab={} colab_type="code" id="Ln2UK2cxge0C"
# day.csvをデータフレームに取り込み
# 日付を表す列はparse_datesで指定する
df = pd.read_csv('day.csv', parse_dates=[1])
# + colab={} colab_type="code" id="t5vqXbzmge0F"
# instant は連番で予測で不要なので削除
df = df.drop('instant', axis=1)
# 項目名の日本語化
columns = [
'日付', '季節', '年', '月', '祝日', '曜日', '勤務日', '天気',
'気温', '体感温度', '湿度', '風速',
'臨時ユーザー利用数', '登録ユーザー利用数', '全体ユーザー利用数'
]
# 項目名を日本語に置き換え
df.columns = columns
# + [markdown] colab_type="text" id="mtZrRFuAbY91"
# #### データ確認
# + colab={} colab_type="code" id="ZmEd-65Rge0I"
# 先頭5行の確認
display(df.head())
# 最終5行の確認
display(df.tail())
# + [markdown] colab_type="text" id="DGzl1rb4bY94"
# ### 5.3.5 データ前処理とデータ分割
# + [markdown] colab_type="text" id="TjD_sEDtbY94"
# #### データ前処理
# Prophet用に列名を置き換える
# + colab={} colab_type="code" id="Jlm4J8gSbY95"
# 「日付」と「登録ユーザー利用数」のみ抽出し、
# 列名を日付:ds 、登録ユーザー利用数:y に置き換えたデータフレームdf2を作る
# データフレーム全体のコピー
df2 = df.copy()
# 「日付」「登録ユーザー利用数」列の抽出
df2 = df2[['日付', '登録ユーザー利用数']]
# 列名の置き換え
df2.columns = ['ds', 'y']
# 結果確認
display(df2.head())
# + [markdown] colab_type="text" id="WmzWhLS8bY97"
# #### データ分割
# 2012年11月1日より前を訓練データ(x_train)に、後ろを検証データ(x_test)にする
# + colab={} colab_type="code" id="vohq0EA-bY97"
# 分割日 mdayの設定
mday = pd.to_datetime('2012-11-1')
# 訓練用indexと検証用indexを作る
train_index = df2['ds'] < mday
test_index = df2['ds'] >= mday
# 入力データの分割
x_train = df2[train_index]
x_test = df2[test_index]
# 日付データの分割(グラフ表示用)
dates_test = df2['ds'][test_index]
# + [markdown] colab_type="text" id="y67jXLzBbY99"
# ### 5.3.6 アルゴリズム選定
# + [markdown] colab_type="text" id="cY7lGUrbbY99"
# #### アルゴリズム選定
# + colab={} colab_type="code" id="Qz0PCJYSge0b"
# ライブラリのimport
from fbprophet import Prophet
# モデル選定
# 3つのseasonalityパラメータの設定が重要
# 今回のデータの場合、日単位のデータなのでdaily_seasonalityは不要
# weekly_seasonality とdaily_seasonalityは
# True / Falseの他に数値で指定することも可能 (三角関数の個数)
# seasonality_mode: additive(デフォルト) multiplicative
m1 = Prophet(yearly_seasonality=True, weekly_seasonality=True,
daily_seasonality=False,
seasonality_mode='multiplicative')
# + [markdown] colab_type="text" id="JUmNdUqfbY-A"
# ### 5.3.7 学習・予測
# + [markdown] colab_type="text" id="I7gcIX7AbY-A"
# #### 学習
# + colab={} colab_type="code" id="mPY8BfZCbY-A"
# 学習
m1.fit(x_train)
# + [markdown] colab_type="text" id="bugmCk5RbY-C"
# #### 予測
# + colab={} colab_type="code" id="WvzpREQsiCu4"
# 予測用データの作成
# (日付 ds だけの入ったデータフレーム)
# 61は予測したい日数 (2012-11-1 から2012-12-31)
future1 = m1.make_future_dataframe(periods=61, freq='D')
# 結果確認
display(future1.head())
display(future1.tail())
# + colab={} colab_type="code" id="pjYTY12DiC32"
# 予測
# 結果はデータフレームで戻ってくる
fcst1 = m1.predict(future1)
# + [markdown] colab_type="text" id="GgtjJNFXbY-I"
# ### 5.3.8 評価
# + colab={} colab_type="code" id="o5ec92r9iC_A"
# 要素ごとのグラフ描画
# この段階ではトレンド、週周期、年周期
fig = m1.plot_components(fcst1)
plt.show()
# + [markdown] colab_type="text" id="-s8zjolXbY-K"
# #### 訓練データ・検証データ全体のグラフ化
# + colab={} colab_type="code" id="ZSsNzEOVO1iO"
# 訓練データ・検証データ全体のグラフ化
fig, ax = plt.subplots(figsize=(10,6))
# 予測結果のグラフ表示(prophetの関数)
m1.plot(fcst1, ax=ax)
# タイトル設定など
ax.set_title('登録ユーザー利用数予測')
ax.set_xlabel('日付')
ax.set_ylabel('利用数')
# グラフ表示
plt.show()
# + [markdown] colab_type="text" id="590Hf3pfbY-N"
# #### R2値の計算
# + colab={} colab_type="code" id="NAZiYWU0ge0i"
# ypred1: fcst1から予測部分のみ抽出する
ypred1 = fcst1[-61:][['yhat']].values
# ytest1: 予測期間中の正解データ
ytest1 = x_test['y'].values
# R2値の計算
from sklearn.metrics import r2_score
score = r2_score(ytest1, ypred1)
# 結果確認
print(f'R2 score:{score:.4f}')
# + [markdown] colab_type="text" id="AjDBnhJKbY-Q"
# #### 予測期間中のグラフ表示(正解データと予測結果)
# + colab={} colab_type="code" id="Wp3jowIdj4ea"
# 時系列グラフの描画
import matplotlib.dates as mdates
fig, ax = plt.subplots(figsize=(8, 4))
# グラフ描画
ax.plot(dates_test, ytest1, label='正解データ', c='k')
ax.plot(dates_test, ypred1, label='予測結果', c='b')
# 日付目盛間隔
# 木曜日ごとに日付を表示
weeks = mdates.WeekdayLocator(byweekday=mdates.TH)
ax.xaxis.set_major_locator(weeks)
# 日付表記を90度回転
ax.tick_params(axis='x', rotation=90)
# 方眼表示など
ax.grid()
ax.legend()
ax.set_title('登録ユーザー利用数予測結果')
# 画面出力
plt.show()
# + [markdown] colab_type="text" id="xX602gIkmE6H"
# ### チューニング方針
#
# * ステップ1 「休日」を特別な日として追加
# * ステップ2 回帰モデルに「天気」「気温」「風速」「湿度」を追加
#
# + [markdown] colab_type="text" id="i2rAP825bY-T"
# ### 5.3.9 チューニング (ステップ1)
# + [markdown] colab_type="text" id="Ey34aEoVbY-T"
# #### ステップ1
# 「休日」を特別な日 (holidays)として追加
# + colab={} colab_type="code" id="n7VBQeYrI5gf"
# 休日の抽出
df_holiday = df[df['祝日']==1]
holidays = df_holiday['日付'].values
# データフレーム形式に変換
df_add = pd.DataFrame({'holiday': 'holi',
'ds': holidays,
'lower_window': 0,
'upper_window': 0
})
# 結果確認
display(df_add.head())
display(df_add.tail())
# + colab={} colab_type="code" id="vGBQjiaFI5gz"
# 休日(df_add)をモデルの入力とする
# アルゴリズム選定
# holidaysパラメータを追加してモデルm2を生成
m2 = Prophet(yearly_seasonality=True,
weekly_seasonality=True, daily_seasonality=False,
holidays = df_add, seasonality_mode='multiplicative')
# 学習
m2 = m2.fit(x_train)
# 予測
fcst2 = m2.predict(future1)
# + [markdown] colab_type="text" id="CBn5MfdEfpou"
# #### ステップ1の評価
# + colab={} colab_type="code" id="q0aXY5oSI5g7"
# 要素ごとのグラフ描画
fig = m2.plot_components(fcst2)
plt.show()
# + colab={} colab_type="code" id="IYcMzQoMI5hB"
# R値の計算
# fcst2から予測部分のみ抽出する
ypred2 = fcst2[-61:][['yhat']].values
# R2値の計算
score2 = r2_score(ytest1, ypred2)
# 結果確認
r2_text2 = f'R2 score:{score2:.4f}'
print(r2_text2)
# + colab={} colab_type="code" id="5UoWVfULbY-d"
# 時系列グラフの描画
import matplotlib.dates as mdates
fig, ax = plt.subplots(figsize=(8, 4))
# グラフ描画
ax.plot(dates_test, ytest1, label='正解データ', c='k')
ax.plot(dates_test, ypred1, label='予測結果v1', c='c')
ax.plot(dates_test, ypred2, label='予測結果v2', c='b')
# 日付目盛間隔
# 木曜日ごとに日付を表示
weeks = mdates.WeekdayLocator(byweekday=mdates.TH)
ax.xaxis.set_major_locator(weeks)
# 日付表記を90度回転
ax.tick_params(axis='x', rotation=90)
# 開始日と終了日
sday = pd.to_datetime('2012-11-1')
eday = pd.to_datetime('2013-1-1')
ax.set_xlim(sday, eday)
# 方眼表示など
ax.grid()
ax.legend()
ax.set_title('登録ユーザー利用数予測結果 ' + r2_text2)
# 画面出力
plt.show()
# + [markdown] colab_type="text" id="-eo9Poylf_DA"
# ### 5.3.10 チューニング (ステップ2)
# 「天気」「気温」「風速」「湿度」を予測モデルに組み込む
# + colab={} colab_type="code" id="aR3VF8YnbY-g"
# 学習データに「天気」「気温」「風速」「湿度」を追加
df3 = pd.concat([df2, df[['天気', '気温', '風速', '湿度']]], axis=1)
# 入力データの分割
x2_train = df3[train_index]
x2_test = df3[test_index]
# 結果確認
display(x2_train.tail())
# + colab={} colab_type="code" id="wVxwXo88OT30"
# アルゴリズム選定
m3 = Prophet(yearly_seasonality=True,
weekly_seasonality=True, daily_seasonality=False,
seasonality_mode='multiplicative', holidays = df_add)
# add_regressor関数で、「天気」「気温」「風速」「湿度」をモデルに組み込む
m3.add_regressor('天気')
m3.add_regressor('気温')
m3.add_regressor('風速')
m3.add_regressor('湿度')
# 学習
m3.fit(x2_train)
# + colab={} colab_type="code" id="0IZYQimTbY-k"
# 予測用の入力データを作る
future3 = df3[['ds', '天気', '気温', '風速', '湿度']]
# 予測
fcst3 = m3.predict(future3)
# + colab={} colab_type="code" id="Wf6UcyL9O1VM"
### ステップ2の評価
# + colab={} colab_type="code" id="UWDQJ6ILQHZJ"
# 要素ごとのグラフ描画
fig = m3.plot_components(fcst3)
plt.show()
# + colab={} colab_type="code" id="Td_bvYYKnILS"
# R値の計算
# fcstから予測部分のみ抽出する
ypred3 = fcst3[-61:][['yhat']].values
score3 = r2_score(ytest1, ypred3)
# 結果確認
r2_text3 = f'R2 score:{score3:.4f}'
print(r2_text3)
# + colab={} colab_type="code" id="nc5sT4zhbY-r"
# 時系列グラフの描画
import matplotlib.dates as mdates
fig, ax = plt.subplots(figsize=(8, 4))
# グラフ描画
ax.plot(dates_test, ytest1, label='正解データ', c='k')
ax.plot(dates_test, ypred2, label='予測結果v2', c='c')
ax.plot(dates_test, ypred3, label='予測結果v3', c='b')
# 日付目盛間隔
# 木曜日ごとに日付を表示
weeks = mdates.WeekdayLocator(byweekday=mdates.TH)
ax.xaxis.set_major_locator(weeks)
# 日付表記を90度回転
ax.tick_params(axis='x', rotation=90)
# 方眼表示など
ax.grid()
ax.legend()
ax.set_title('登録ユーザー利用数予測結果 ' + r2_text3)
# 画面出力
plt.show()
# + [markdown] colab_type="text" id="DuCmbsVagL_j"
# ### コラム アイスクリーム購買予測」で時系列分析
# + [markdown] colab_type="text" id="hUc6jWyHbY-w"
# #### オリジナルデータ
#
# アイスクリーム調査報告書
# https://www.icecream.or.jp/biz/data/expenditures.html
#
# 下記のEXCELは、この報告書の内容を元に起こして作りました。
# + colab={} colab_type="code" id="myXoKWMJbY-x"
# データ読み込み
url2 = 'https://github.com/makaishi2/\
sample-data/blob/master/data/ice-sales.xlsx?raw=true'
df = pd.read_excel(url2, sheet_name=0)
# + colab={} colab_type="code" id="JjsvEPizbY-z"
# データ確認
display(df.head())
display(df.tail())
# + colab={} colab_type="code" id="e5VpukxKbY-1"
# 時系列グラフの描画 (アイスクリーム支出金額)
fig, ax = plt.subplots(figsize=(12, 4))
# グラフ描画
ax.plot(df['年月'], df['支出'],c='b')
# 3か月区切りの目盛にする
month3 = mdates.MonthLocator(interval=3)
ax.xaxis.set_major_locator(month3)
# 日付表記を90度回転
ax.tick_params(axis='x', rotation=90)
# 開始日と終了日
sday = pd.to_datetime('2015-1-1')
eday = pd.to_datetime('2019-12-31')
ax.set_xlim(sday, eday)
# 方眼表示など
ax.grid()
ax.set_title('アイスクリーム支出金額')
# 画面出力
plt.show()
# + colab={} colab_type="code" id="lmj32S9EbY-2"
# データ前処理
# データ形式をProphet用に合わせる
x = df.copy()
x.columns = ['ds', 'y']
display(x.head())
# + colab={} colab_type="code" id="yOP8KJzwbY-4"
# データ分割
# 2019年1月を基準に訓練データと検証データを分割
# 分割日 mdayの設定
mday = pd.to_datetime('2019-1-1')
# 訓練用indexと検証用indexを作る
train_index = x['ds'] < mday
test_index = x['ds'] >= mday
# 入力データの分割
x_train = x[train_index]
x_test = x[test_index]
#日付列もグラフ描画のために分割
dates_train = x['ds'][train_index]
dates_test = x['ds'][test_index]
# + colab={} colab_type="code" id="SL3sbCwhbY-5"
# アルゴリズムの選択
# ライブラリのimport
from fbprophet import Prophet
m = Prophet(yearly_seasonality=5, weekly_seasonality=False, daily_seasonality=False)
# + colab={} colab_type="code" id="DJH7cG0TbY-8"
# 学習
m = m.fit(x_train)
# + colab={} colab_type="code" id="MxBaYjMdbY--"
# 予測
future = x[['ds']]
fcst = m.predict(future)
# + colab={} colab_type="code" id="bBJsQR6dbY_A"
# 評価
# fcstから予測部分のみ抽出する
ypred = fcst[-12:]['yhat'].values
# 正解データのリスト
ytest = x_test['y'].values
# R値の計算
from sklearn.metrics import r2_score
score = r2_score(ytest, ypred)
score_text = f'R2 score:{score:.4f}'
print(score_text)
# + colab={} colab_type="code" id="9lYDmGBRbY_C"
# 時系列グラフの描画 (アイスクリーム支出金額)
fig, ax = plt.subplots(figsize=(8, 4))
# グラフ描画
ax.plot(dates_test, ytest, label='正解データ', c='k')
ax.plot(dates_test, ypred, label='予測結果', c='b')
# 1か月区切りの目盛にする
month = mdates.MonthLocator()
ax.xaxis.set_major_locator(month)
# 日付表記を90度回転
ax.tick_params(axis='x', rotation=90)
# 開始日と終了日
sday = pd.to_datetime('2019-1-1')
eday = pd.to_datetime('2019-12-1')
ax.set_xlim(sday, eday)
# 方眼表示など
ax.grid()
ax.legend()
ax.set_title('アイスクリーム支出金額予測 ' + score_text)
# 画面出力
plt.show()
# + colab={} colab_type="code" id="RSCimcyVbY_F"
| notebooks/ch05_03_bike_sharing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NYTCovid class test
from NYTCovid import NYTCovid
n = NYTCovid()
n.dateUpdate()
n.updateState()
n.peek()
n.plot_state(state='California',last_30_days=True)
n.process()
n.plot_state(state='California',last_30_days=True)
n.plot_multi_state(states=['California', 'Michigan', 'Georgia','Illinois'],
last_30_days=True)
n.rankState(N=6)
n.rankState(N=4,daterank='2020-03-26')
| NYTCovid-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import spearmanr
from sklearn.preprocessing import StandardScaler
import torch
import tensorflow as tf
tf.compat.v1.disable_v2_behavior()
import torch.nn as nn
SEED = 1
torch.manual_seed(SEED)
np.random.seed(SEED)
tf.random.set_seed(SEED)
# +
x = np.random.normal(size=(300, 50))
y = x[:,20] + x[:,40] + np.random.normal(scale = 0.01, size=300)
ss = StandardScaler()
x = ss.fit_transform(x)
# +
X_train = x[:200]
y_train = y[:200]
X_test = x[200:]
y_test = y[200:]
X_train = torch.tensor(X_train).float()
y_train = torch.tensor(y_train).view(-1, 1).float()
X_test = torch.tensor(X_test).float()
y_test = torch.tensor(y_test).view(-1, 1).float()
datasets = torch.utils.data.TensorDataset(X_train, y_train)
train_iter = torch.utils.data.DataLoader(datasets, batch_size=10, shuffle=True)
# -
batch_size = 50
num_epochs = 200
learning_rate = 0.0001
size_hidden1 = 50
size_hidden2 = 50
size_hidden3 = 1
class MyModel(nn.Module):
def __init__(self):
super().__init__()
self.lin1 = nn.Linear(50, size_hidden1)
self.relu1 = nn.ReLU()
self.lin2 = nn.Linear(size_hidden1, size_hidden2)
self.relu2 = nn.ReLU()
self.lin3 = nn.Linear(size_hidden2, size_hidden3)
def forward(self, input):
return self.lin3(self.relu2(self.lin2(self.relu1(self.lin1(input)))))
def predict(self, input):
x = torch.Tensor(input)
return self.forward(x).detach().numpy()
model = MyModel()
model.train()
# +
criterion = nn.MSELoss(reduction='sum')
def train(model_inp, num_epochs = num_epochs):
optimizer = torch.optim.Adam(model_inp.parameters(), lr=learning_rate)
for epoch in range(num_epochs): # loop over the dataset multiple times
running_loss = 0.0
for inputs, labels in train_iter:
# forward pass
outputs = model_inp(inputs)
# defining loss
loss = criterion(outputs, labels)
# zero the parameter gradients
optimizer.zero_grad()
# computing gradients
loss.backward()
# accumulating running loss
running_loss += loss.item()
# updated weights based on computed gradients
optimizer.step()
if epoch % 20 == 0:
print('Epoch [%d]/[%d] running accumulative loss across all batches: %.3f' %
(epoch + 1, num_epochs, running_loss))
running_loss = 0.0
# -
train(model)
from sklearn.metrics import mean_squared_error
model.eval()
outputs = model(X_test).detach().numpy()
err = np.sqrt(mean_squared_error(outputs, y_test.detach().numpy()))
print(err)
outputs = pd.Series(outputs[:,0], index=range(200,300))
plt.scatter(outputs, y[200:])
plt.xlabel("Output")
plt.ylabel("Label")
def get_masked_data_for_CXPlain(model, x):
x_train = torch.FloatTensor(x)
n_feats = x.shape[1]
patch = 5
mask = np.ones((n_feats//patch, n_feats))
for i in range(n_feats//patch):
mask[i, i*patch:(i+1)*patch] = 0
y_pred = model(x_train).detach().numpy()
mask = torch.FloatTensor(mask)
list_of_masked_outs = []
for i, sample in enumerate(x_train):
masked_sample = sample*mask
list_of_masked_outs.append(model(masked_sample).unsqueeze(0).detach().numpy())
masked_outs = np.concatenate(list_of_masked_outs)
return(x, y_pred, masked_outs)
k = get_masked_data_for_CXPlain(model, x[:200])
k[2].shape
# +
from tensorflow.python.keras.losses import mean_squared_error as loss
from cxplain import CXPlain
from cxplain.backend.model_builders.custom_mlp import CustomMLPModelBuilder
model_builder = CustomMLPModelBuilder(num_layers=2, num_units=32, batch_size=32, learning_rate=0.001, n_feature_groups=10)
explainer = CXPlain(model, model_builder, None, loss)
# -
explainer.fit(x[:200], y[:200], masked_data=k)
attributions = explainer.explain_groups(x[200:])
attr = pd.DataFrame(attributions, index=range(200, 300))
attr
for i in attr.index:
plt.plot(range(10), attr.loc[i].values)
plt.show()
# # Pathway toy example
# +
x = np.random.normal(size=(300, 50))
y = np.zeros((300))
for i in range(10):
y += x[:,i*5]
# y += np.random.normal(scale = 0.01, size=300)
ss = StandardScaler()
x = ss.fit_transform(x)
# +
X_train = x[:200]
y_train = y[:200]
X_test = x[200:]
y_test = y[200:]
X_train = torch.tensor(X_train).float()
y_train = torch.tensor(y_train).view(-1, 1).float()
X_test = torch.tensor(X_test).float()
y_test = torch.tensor(y_test).view(-1, 1).float()
datasets = torch.utils.data.TensorDataset(X_train, y_train)
train_iter = torch.utils.data.DataLoader(datasets, batch_size=10, shuffle=True)
# -
model = MyModel()
model.train()
train(model, num_epochs=300)
# +
model.eval()
outputs = model(X_test).detach().numpy()
err = np.sqrt(mean_squared_error(outputs, y_test.detach().numpy()))
print(err)
outputs = pd.Series(outputs[:,0], index=range(200,300))
plt.scatter(outputs, y[200:])
plt.xlabel("Output")
plt.ylabel("Label")
# +
from cxplain import MLPModelBuilder, ZeroMasking
model_builder = MLPModelBuilder(num_layers=2, num_units=32, batch_size=32, learning_rate=0.001)
masking_operation = ZeroMasking()
explainer = CXPlain(model, model_builder, masking_operation, loss)
explainer.fit(x[:200], y[:200])
# -
attributions = explainer.explain(x[200:])
attr = pd.DataFrame(attributions, index=range(200, 300))
attr
for i in attr.index:
plt.plot(range(50), attr.loc[i].values)
plt.show()
def get_masked_pathways_for_CXPlain(model, x):
x_train = torch.FloatTensor(x)
n_pathways = 8
n_feats = 50
mask = np.zeros((n_pathways, n_feats))
mask[0] = [0 if i%5!=0 else 1 for i in range(50)] # 10 features div by 5 {all important}
mask[1] = [0 if i%10!=0 else 1 for i in range(50) ] # 5 features div by 10 {half of the important features}
mask[2] = [0 if (i+1)%5!=0 else 1 for i in range(50)] # 10 trivial features
mask[3] = [0 if (i+2)%10!=0 else 1 for i in range(50)] # 5 trivial features {8, 18, 28, 38, 48}
mask[4] = mask[1] + mask[3] # 5 trivial + 5 important
mask[5] = mask[0] + mask[2] # 10 trivial + 10 important
mask[6] = mask[0] + mask[3] # 5 trivial + 10 important
mask[7] = mask[1] + mask[2] # 10 trivial + 5 important
# expected: 0 > 6 > {5 ? 4} > 7 > {2 ? 3}
# remove those in "pathway"
mask = np.ones((n_pathways, n_feats)) - mask
y_pred = model(x_train).detach().numpy()
mask = torch.FloatTensor(mask)
list_of_masked_outs = []
for i, sample in enumerate(x_train):
masked_sample = sample*mask
list_of_masked_outs.append(model(masked_sample).unsqueeze(0).detach().numpy())
masked_outs = np.concatenate(list_of_masked_outs)
return(x, y_pred, masked_outs)
k = get_masked_pathways_for_CXPlain(model, x[:200])
print(k[2].shape)
model_builder = CustomMLPModelBuilder(num_layers=2, num_units=32, batch_size=32, learning_rate=0.001, n_feature_groups=8)
explainer = CXPlain(model, model_builder, None, loss)
explainer.fit(x[:200], y[:200], masked_data=k)
attributions = explainer.explain_groups(x[200:])
attr = pd.DataFrame(attributions, index=range(200, 300))
attr
for i in attr.index:
plt.plot(range(8), attr.loc[i].values, alpha=0.3)
plt.show()
plt.plot(range(8), attr.mean(axis=0).values, marker='o')
# expected: 0 > {1 ? 6} > {5 ? 4} > 7 > {2 ? 3}
# 0: 10 important
#
# 1: 5 important
#
# 2: 10 trivial
#
# 3: 5 trivial
#
# 4: 5 important + 5 trivial
#
# 5: 10 important + 10 trivial
#
# 6: 10 important + 5 trivial
#
# 7: 5 important + 10 trivial
plt.plot(range(8), attr.median(axis=0).values, marker='o')
# **NOTES:**
# - attr[6] > attr[0] : This is not good because group 6 has 5 nuisance features
# - attr[6] > attr[5] : This is good because group 6 has less nuisance features than group 5
# - attr[3] > attr[2] : 5 trivial > 10 trivial?
# - attr[7] > attr[4] > attr[1]: This shows the bias towards the cardinality of zeroed features
n_zero = np.asarray([10, 5, 10, 5, 10, 20, 15, 15])
k = attr.median(axis=0).values/n_zero
plt.plot(range(8), k, marker='o')
# # All groups have 10 important features and i nuisance
def get_masked_pathways_for_CXPlain2(model, x):
x_train = torch.FloatTensor(x)
n_pathways = 41
n_feats = 50
mask = np.zeros((n_pathways, n_feats))
mask[0] = [0 if i%5!=0 else 1 for i in range(50)] # 10 features div by 5 {all important}
print(np.nonzero(1-mask[0]))
for i, j in enumerate(np.nonzero(1-mask[0])[0]): # 10 important, i trivial
mask[i+1] = mask[i]
mask[i+1, j] = 1
# remove those in "pathway"
mask = np.ones((n_pathways, n_feats)) - mask
print(mask.sum(axis=1))
y_pred = model(x_train).detach().numpy()
mask = torch.FloatTensor(mask)
list_of_masked_outs = []
for i, sample in enumerate(x_train):
masked_sample = sample*mask
list_of_masked_outs.append(model(masked_sample).unsqueeze(0).detach().numpy())
masked_outs = np.concatenate(list_of_masked_outs)
return(x, y_pred, masked_outs)
k = get_masked_pathways_for_CXPlain2(model, x[:200])
k[2].shape
model_builder = CustomMLPModelBuilder(num_layers=2, num_units=32, batch_size=32, learning_rate=0.001, n_feature_groups=41)
explainer = CXPlain(model, model_builder, None, loss, num_models=3)
explainer.fit(x[:200], y[:200], masked_data=k)
attributions, conf = explainer.explain_groups(x[200:])
conf.shape
attributions.shape
# attributions = explainer.explain_groups(x[200:])
attr = pd.DataFrame(attributions, index=range(200, 300))
attr
for i in attr.index:
plt.plot(range(41), attr.loc[i].values, alpha=0.3)
plt.show()
plt.plot(range(41), attr.mean(axis=0).values, marker='o', label='mean')
plt.plot(range(41), attr.median(axis=0).values, marker='*', label='median')
plt.xlabel('# of nuisance')
plt.show()
attr.sum(axis=1)
# **NOTE:** If all "pathways" have the same 10 important features, more zeroed features can mean higher attribution even if most of them are nuisance features only. However, the differences might be a bit smaller.
# try the completely irrelevant ones
#
# some vs none (check size bias)
# # 10-i Important and i nuisance features per group
def get_masked_pathways_for_CXPlain3(model, x):
x_train = torch.FloatTensor(x)
n_pathways = 11
n_feats = 50
mask = np.zeros((n_pathways, n_feats))
mask[0] = [0 if i%5!=0 else 1 for i in range(50)] # 10 features div by 5 {all important}
for i, j in enumerate(np.nonzero(mask[0])[0]): # 10-i important, i trivial
mask[i+1] = mask[i]
mask[i+1, j] = 0
mask[i+1, j+1] = 1
# remove those in "pathway"
mask = np.ones((n_pathways, n_feats)) - mask
print(mask)
y_pred = model(x_train).detach().numpy()
mask = torch.FloatTensor(mask)
list_of_masked_outs = []
for i, sample in enumerate(x_train):
masked_sample = sample*mask
list_of_masked_outs.append(model(masked_sample).unsqueeze(0).detach().numpy())
masked_outs = np.concatenate(list_of_masked_outs)
return(x, y_pred, masked_outs)
k = get_masked_pathways_for_CXPlain3(model, x[:200])
model_builder = CustomMLPModelBuilder(num_layers=2, num_units=32, batch_size=32, learning_rate=0.001, n_feature_groups=11)
explainer = CXPlain(model, model_builder, None, loss, num_models=3)
explainer.fit(x[:200], y[:200], masked_data=k)
attributions, conf = explainer.explain_groups(x[200:])
attr = pd.DataFrame(attributions, index=range(200, 300))
attr
for i in attr.index:
plt.plot(range(11), attr.loc[i].values, alpha=0.3)
plt.show()
plt.plot(range(11), attr.mean(axis=0).values, marker='o', label='mean')
plt.plot(range(11), attr.median(axis=0).values, marker='*', label='median')
plt.xlabel('# of nuisance')
plt.show()
| ToyExampleCustomMask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
# Loading cleaned World Bank data on forest area as percent of land area...
pct_forest_coverage = pd.read_csv('pct_forest_coverage.csv', index_col=0)
pct_forest_coverage
# +
# Loading predicted data on forest area as percent of land area...
pct_forest_coverage_predictions = pd.read_csv(
'pct_forest_coverage_predictions.csv', index_col=0)
# Adding historial data from 2016 to dataframe for use in lineplots...
pct_forest_coverage_predictions['2016'] = pct_forest_coverage['2016']
pct_forest_coverage_predictions
# +
# Putting data in tidy format...
pct_forest_coverage_tidy = pct_forest_coverage.melt(
id_vars=['Country', 'Code'])
pct_forest_coverage_predictions_tidy = pct_forest_coverage_predictions.melt(
id_vars=['Country', 'Code'])
# Renaming columns...
pct_forest_coverage_tidy = pct_forest_coverage_tidy.rename(
columns={'variable': 'Year', 'value': 'Coverage'})
pct_forest_coverage_predictions_tidy = (
pct_forest_coverage_predictions_tidy.rename(
columns={'variable': 'Year', 'value': 'Coverage'}))
# +
import seaborn as sns
import matplotlib.pyplot as plt
plt.ioff()
# Getting list of country codes...
codes = list(pct_forest_coverage_tidy['Code'].unique())
# Getting list of codes for countries without data...
nulls = (pct_forest_coverage_predictions[
pct_forest_coverage_predictions['2017'].isna()]['Code']).tolist()
# Creating list of codes for countries with data...
codes = [code for code in codes if code not in nulls]
# For-loop to create a chart for each country where we have data...
for code in codes:
historical_data = pct_forest_coverage_tidy[
pct_forest_coverage_tidy['Code'] == code]
predicted_data = pct_forest_coverage_predictions_tidy[
pct_forest_coverage_predictions_tidy['Code'] == code]
plt.style.use('fivethirtyeight')
f, ax = plt.subplots(figsize=(30, 10))
sns.lineplot(x=historical_data['Year'],
y=historical_data['Coverage'],
color='green');
sns.lineplot(x=predicted_data['Year'],
y=predicted_data['Coverage'],
color='darkgray');
ax.lines[1].set_linestyle("--")
ax.set(xlabel='Year', ylabel='Forest Area as % of Land Area')
country = list(set(historical_data['Country']))[0]
ax.set_title(country + ' (' + code +
'): ' + 'Forest Area as % of Land Area by Year')
country_with_underscores = country.replace(' ', '_')
f.savefig('Country_Charts/' + country_with_underscores + '.png')
plt.close(f)
| Generating Charts with World Bank Data.ipynb |