code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: all
# notebook_metadata_filter: all
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# language_info:
# codemirror_mode:
# name: ipython
# version: 3
# file_extension: .py
# mimetype: text/x-python
# name: python
# nbconvert_exporter: python
# pygments_lexer: ipython3
# version: 3.6.7
# ---
# # Generate stimuli for Sound-Switch Experiment 1 <a class="tocSkip">
# C and G guitar chords generated [here](https://www.apronus.com/music/onlineguitar.htm) and subsequently recorded and amplified (25.98 db and 25.065 db respectively) using Audacity version 2.3.0 (using Effect --> Amplify) via Windows 10's Stereo Mix drivers.
# # Imports
from pydub import AudioSegment
from pydub.generators import Sine
from pydub.playback import play
import random
import numpy as np
import os
# # Functions
def generate_songs(path_prefix):
for switch_probability in switch_probabilities:
for exemplar in range(num_exemplars):
# Begin with silence
this_song = silence
# Choose random tone to start with
which_tone = round(random.random())
for chunk in range(num_chunks):
this_probability = random.random()
# Change tones if necessary
if this_probability < switch_probability:
which_tone = 1 - which_tone
this_segment = songs[which_tone][:chunk_size]
# Add intervening silence
this_song = this_song.append(silence, crossfade=crossfade_duration)
# Add tone
this_song = this_song.append(this_segment, crossfade=crossfade_duration)
# Add final silence
this_song.append(silence, crossfade=crossfade_duration)
song_name = f"{path_prefix}switch-{str(round(switch_probability,2))}_chunk-{str(chunk_size)}_C_G_alternating_{str(exemplar).zfill(2)}.mp3"
this_song.export(song_name, format="mp3", bitrate="192k")
# # Stimulus Generation
# ## Guitar chords
# +
songs = [
AudioSegment.from_mp3("guitar_chords/guitar_C.mp3"),
AudioSegment.from_mp3("guitar_chords/guitar_G.mp3"),
]
chunk_size = 500 # in ms
num_chunks = 20
crossfade_duration = 50 # in ms
silence_duration = 100 # in ms
switch_probabilities = np.linspace(0.1, 0.9, num=9)
num_exemplars = 10
silence = AudioSegment.silent(duration=silence_duration)
# Generate the songs
generate_songs(path_prefix="guitar_chords/")
# -
# ## Tones
# +
# Create sine waves of given freqs
frequencies = [261.626, 391.995] # C4, G4
sample_rate = 44100 # sample rate
bit_depth = 16 # bit depth
# Same params as above for guitar
chunk_size = 500 # in ms
num_chunks = 20
crossfade_duration = 50 # in ms
silence_duration = 100 # in ms
switch_probabilities = np.linspace(0.1, 0.9, num=9)
num_exemplars = 10
silence = AudioSegment.silent(duration=silence_duration)
sine_waves = []
songs = []
for i, frequency in enumerate(frequencies):
sine_waves.append(Sine(frequency, sample_rate=sample_rate, bit_depth=bit_depth))
#Convert waveform to audio_segment for playback and export
songs.append(sine_waves[i].to_audio_segment(duration=chunk_size*2)) # just to make sure it's long enough
generate_songs(path_prefix="pure_tones/")
# -
# # Practice Stimulus
# Just choose one of the above stimuli to be a practice stimulus, and remake the stimuli so that it doesn't get repeated.
| stimuli/generate_stimuli.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HMyxX1Gmh7cl" colab_type="text"
# https://colab.research.google.com/drive/1jKzKN_7Fj-OIwTZlVBDaxVj5ct1uGAva
# + [markdown] id="c0x9z68Byl2u" colab_type="text"
# # GAN
# + id="7aivHtGkBOKw" colab_type="code" outputId="a77b70ad-c995-4ab9-9194-1540e5104ebb" colab={"base_uri": "https://localhost:8080/", "height": 34}
import numpy as np
from keras.datasets import mnist
from keras.layers import Input, Dense, Reshape, Flatten, Dropout
from keras.layers import BatchNormalization
from keras.layers.advanced_activations import LeakyReLU
from keras.models import Sequential
from keras.optimizers import Adam
import matplotlib.pyplot as plt
# %matplotlib inline
plt.switch_backend('agg')
# + id="2slim96TPTcH" colab_type="code" colab={}
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Reshape
from keras.layers.core import Activation
from keras.layers.normalization import BatchNormalization
from keras.layers.convolutional import UpSampling2D
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.layers.core import Flatten
from keras.optimizers import SGD
from keras.datasets import mnist
import numpy as np
from PIL import Image
import argparse
import math
# + id="ePrTaaRlFErE" colab_type="code" outputId="515301ae-daef-412f-c13e-da2abe795462" colab={"base_uri": "https://localhost:8080/", "height": 170}
# !pip install logger
from logger import logger
# + id="VdgeZQ3PDQsY" colab_type="code" colab={}
shape = (28, 28, 1)
epochs = 400
batch = 32
save_interval = 100
# + id="o_NdO2D5Duwn" colab_type="code" colab={}
def generator():
model = Sequential()
model.add(Dense(256, input_shape=(100,)))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(512))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(1024))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(28 * 28 * 1, activation='tanh'))
model.add(Reshape(shape))
return model
# + id="5JsKSvY0Dw0s" colab_type="code" colab={}
def discriminator():
model = Sequential()
model.add(Flatten(input_shape=shape))
model.add(Dense((28 * 28 * 1), input_shape=shape))
model.add(LeakyReLU(alpha=0.2))
model.add(Dense(int((28 * 28 * 1) / 2)))
model.add(LeakyReLU(alpha=0.2))
model.add(Dense(1, activation='sigmoid'))
return model
# + id="f8c7eJjbdS0v" colab_type="code" outputId="bbf302bc-c5ad-47dc-c501-c5accdde9afe" colab={"base_uri": "https://localhost:8080/", "height": 527}
Generator = generator()
Generator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8))
Discriminator = discriminator()
Discriminator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8),metrics=['accuracy'])
# + id="wVT8owFodTzH" colab_type="code" outputId="5de7d734-bee4-4c19-da1a-87e21dfbbebf" colab={"base_uri": "https://localhost:8080/", "height": 884}
print(Discriminator.summary(), Generator.summary())
# + id="t26rWMb9-lRV" colab_type="code" outputId="a676bb50-2d19-48c5-a89a-d59162953b8c" colab={"base_uri": "https://localhost:8080/", "height": 527}
Generator.summary()
# + id="osVaP6pqDy5-" colab_type="code" colab={}
def stacked_generator_discriminator(D, G):
D.trainable = False
model = Sequential()
model.add(G)
model.add(D)
return model
# + id="ObcGi9PdD6EI" colab_type="code" colab={}
def plot_images(samples=16, step=0):
filename = "mnist_%d.png" % step
noise = np.random.normal(0, 1, (samples, 100))
images = Generator.predict(noise)
plt.figure(figsize=(5, 5))
for i in range(images.shape[0]):
plt.subplot(4, 4, i + 1)
image = images[i, :, :, :]
image = np.reshape(image, [28, 28])
plt.imshow(image, cmap='gray')
plt.axis('off')
plt.tight_layout()
plt.show()
#plt.close('all')
# + id="REhNMaBOEGJw" colab_type="code" colab={}
stacked_generator_discriminator = stacked_generator_discriminator(Discriminator, Generator)
stacked_generator_discriminator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8))
# + id="PyBn269FPquS" colab_type="code" outputId="caf84757-ac2b-4581-a228-8f3585b27d2c" colab={"base_uri": "https://localhost:8080/", "height": 221}
stacked_generator_discriminator.summary()
# + id="_Zzp5_9zEd6T" colab_type="code" outputId="6a35f310-8bdd-442b-991c-0f791243dae5" colab={"base_uri": "https://localhost:8080/", "height": 51}
(X_train, _), (_, _) = mnist.load_data()
X_train = (X_train.astype(np.float32) - 127.5) / 127.5
X_train = np.expand_dims(X_train, axis=3)
# + id="OfSz3mV_RkPH" colab_type="code" colab={}
save_interval = 250
# + id="X2bZzvGIEg54" colab_type="code" colab={}
# %matplotlib inline
disc_loss = []
gen_loss = []
for cnt in range(4000):
random_index = np.random.randint(0, len(X_train) - batch / 2)
legit_images = X_train[random_index: random_index + batch // 2].reshape(batch // 2, 28, 28, 1)
gen_noise = np.random.normal(-1, 1, (batch // 2, 100))/2
syntetic_images = Generator.predict(gen_noise)
x_combined_batch = np.concatenate((legit_images, syntetic_images))
y_combined_batch = np.concatenate((np.ones((batch // 2, 1)), np.zeros((batch // 2, 1))))
d_loss = Discriminator.train_on_batch(x_combined_batch, y_combined_batch)
noise = np.random.normal(-1, 1, (batch, 100))/2
y_mislabled = np.ones((batch, 1))
g_loss = stacked_generator_discriminator.train_on_batch(noise, y_mislabled)
logger.info('epoch: {}, [Discriminator: {}], [Generator: {}]'.format(cnt, d_loss[0], g_loss))
disc_loss.append(d_loss[0])
gen_loss.append(g_loss)
if cnt % save_interval == 0:
plot_images(step=cnt)
# + id="kIdr7A8BN5sO" colab_type="code" outputId="e8bd0d72-1bc2-4c8d-916a-d894bdff6e82" colab={"base_uri": "https://localhost:8080/", "height": 346}
import matplotlib.ticker as mtick
import matplotlib.pyplot as plt
# %matplotlib inline
epochs = range(1, 4001)
plt.plot(epochs, disc_loss, 'bo', label='Discriminator loss')
plt.plot(epochs, gen_loss, 'r', label='Generator loss')
plt.title('Generator and Discriminator loss values')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
# + id="OkF6yBXSE_L2" colab_type="code" outputId="01d302b2-35d1-43ab-c7a7-fad10bda1dd5" colab={"base_uri": "https://localhost:8080/", "height": 369}
plot_images(step=cnt)
# + id="cqp8oeGi68jK" colab_type="code" outputId="3a4f32ec-600c-417f-9016-8c07a640d62b" colab={"base_uri": "https://localhost:8080/", "height": 202}
noise = np.random.normal(0, 1, (1, 100))
images = Generator.predict(noise)
plt.figure(figsize=(10, 10))
for i in range(images.shape[0]):
plt.subplot(4, 4, i + 1)
image = images[i, :, :, :]
image = np.reshape(image, [28, 28])
plt.imshow(image, cmap='gray')
plt.axis('off')
plt.tight_layout()
plt.show()
# + id="zJzlYFffVW5W" colab_type="code" colab={}
# + id="eRfJZxjaVW7r" colab_type="code" colab={}
# + [markdown] id="H-3IlCV2VXMY" colab_type="text"
# # DCGAN
# + id="eqPphZ_9HeNE" colab_type="code" colab={}
def generator():
model = Sequential()
model.add(Dense(input_dim=100, output_dim=1024))
model.add(Activation('tanh'))
model.add(Dense(128*7*7))
model.add(BatchNormalization())
model.add(Activation('tanh'))
model.add(Reshape((7, 7, 128), input_shape=(128*7*7,)))
model.add(UpSampling2D(size=(2, 2)))
model.add(Conv2D(64, (5, 5), padding='same'))
model.add(Activation('tanh'))
model.add(UpSampling2D(size=(2, 2)))
model.add(Conv2D(1, (5, 5), padding='same'))
model.add(Activation('tanh'))
return model
def discriminator():
model = Sequential()
model.add(
Conv2D(64, (5, 5),
padding='same',
input_shape=(28, 28, 1))
)
model.add(Activation('tanh'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, (5, 5)))
model.add(Activation('tanh'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(1024))
model.add(Activation('tanh'))
model.add(Dense(1))
model.add(Activation('sigmoid'))
return model
# + id="BXtQvpKgECRx" colab_type="code" outputId="d79d77a6-dc1a-4787-cefb-a63f4aa9dcca" colab={"base_uri": "https://localhost:8080/", "height": 187}
Generator = generator()
Generator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8))
Discriminator = discriminator()
Discriminator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8),metrics=['accuracy'])
def stacked_generator_discriminator(D, G):
D.trainable = False
model = Sequential()
model.add(G)
model.add(D)
return model
stacked_generator_discriminator = stacked_generator_discriminator(Discriminator, Generator)
stacked_generator_discriminator.compile(loss='binary_crossentropy', optimizer=Adam(lr=0.0002, beta_1=0.5, decay=8e-8))
# + id="UCj7YcjpVoTF" colab_type="code" colab={}
# %matplotlib inline
disc_loss = []
gen_loss = []
for cnt in range(4000):
random_index = np.random.randint(0, len(X_train) - batch / 2)
legit_images = X_train[random_index: random_index + batch // 2].reshape(batch // 2, 28, 28, 1)
gen_noise = np.random.normal(-1, 1, (batch // 2, 100))/2
syntetic_images = Generator.predict(gen_noise)
x_combined_batch = np.concatenate((legit_images, syntetic_images))
y_combined_batch = np.concatenate((np.ones((batch // 2, 1)), np.zeros((batch // 2, 1))))
d_loss = Discriminator.train_on_batch(x_combined_batch, y_combined_batch)
noise = np.random.normal(-1, 1, (batch, 100))/2
y_mislabled = np.ones((batch, 1))
g_loss = stacked_generator_discriminator.train_on_batch(noise, y_mislabled)
logger.info('epoch: {}, [Discriminator: {}], [Generator: {}]'.format(cnt, d_loss[0], g_loss))
disc_loss.append(d_loss[0])
gen_loss.append(g_loss)
if cnt % save_interval == 0:
plot_images(step=cnt)
# + id="5z484ecEVqWb" colab_type="code" outputId="f0b0b8a4-f98f-479e-b759-7668184b119b" colab={"base_uri": "https://localhost:8080/", "height": 346}
import matplotlib.ticker as mtick
import matplotlib.pyplot as plt
# %matplotlib inline
epochs = range(1, 4001)
plt.plot(epochs, disc_loss, 'bo', label='Discriminator loss')
plt.plot(epochs, gen_loss, 'r', label='Generator loss')
plt.title('Generator and Discriminator loss values')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
# + id="vwEZDCVGPrSR" colab_type="code" colab={}
| Chapter08/Vanilla_and_DC_GAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#Preliminary feature selection for Merge Table 1
# -
#import necessary packages
import pandas as pd
import datetime as dt
import seaborn as sns
import matplotlib.pyplot as plt
import yellowbrick as yb
import warnings
warnings.filterwarnings("ignore")
path = 'MergeTable1_FirmsAndSmWfigs2Round.csv'
df1 = pd.read_csv('MergeTable1_FirmsAndSmWfigs2Round.csv', error_bad_lines=False)
#Identifying the percentage of missing values present in each respective column of the dataframe
percent_missing = df1.isnull().sum() * 100 / len(df1)
print(percent_missing)
#Drop all columns containing more than 50 null values.
df1drop = df1.dropna(how='all', thresh=50, axis=1)
#Converting daynight and satellite values to numerical values.
daynight_map = {"D": 1, "N": 0}
satellite_map = {"Terra": 1, "Aqua": 0}
df1drop['daynight'] = df1drop['daynight'].map(daynight_map)
df1drop['satellite'] = df1drop['satellite'].map(satellite_map)
#Seperate acq_date column into seperate 'day', 'month', and 'year'columns.
df1drop['acq_date'] = pd.to_datetime(df1drop['acq_date'])
df1drop['year'] = df1drop['acq_date'].dt.year
df1drop['month'] = df1drop['acq_date'].dt.month
df1drop['day'] = df1drop['acq_date'].dt.day
#Now that we have seperated our 'acq_date' data into day, month, and year columns, we can drop the 'acq_date' column to eliminate redundancy.
df1drop.drop("acq_date", axis=1, inplace=True)
#Drop columns unneccesary to our exploration.
df1drop.drop("instrument", axis=1, inplace=True)
df1drop.drop("date_loc", axis=1, inplace=True)
df1drop.drop("date_hour_loc", axis=1, inplace=True)
df1drop.drop("acq_time", axis=1, inplace=True)
df1drop.drop("type", axis=1, inplace=True)
df1drop.drop("Unnamed: 0", axis=1, inplace=True)
df1drop.drop("latitude", axis=1, inplace=True)
df1drop.drop("longitude", axis=1, inplace=True)
#Create a new dataframe that does not contain null values.
df1drop = df1drop.dropna()
#Convert data in dataframe to string type to work with Yellowbrick visualizations.
df1drop1 = df1drop.astype(str)
#Seperate data sets as labels and features.
X = df1drop1.drop('FIRE_DETECTED', axis=1)
y = df1drop1['FIRE_DETECTED']
features = ["brightness", "scan", "track", "satellite", "confidence", "version", "bright_t31", "frp", "daynight", "lat", "long", "hour_x", "day", "month", "year"]
# +
#Use RandomForestClassifier to plot feature importance for our dataset. As we can see from the output, hour_x, daynight, version, and brightness do not seem to contribute significantly to our model.
from sklearn.ensemble import RandomForestClassifier
from yellowbrick.features import FeatureImportances
model = RandomForestClassifier(n_estimators=10)
viz = FeatureImportances(model, labels=features, size=(1080, 720))
viz.fit(X, y)
viz.show()
# +
#Create a Shaprio ranking of our features.
from yellowbrick.features import Rank1D
fig, ax = plt.subplots(1, figsize=(8, 12))
vzr = Rank1D(ax=ax)
vzr.fit(X, y)
vzr.transform(X)
sns.despine(left=True, bottom=True)
vzr.poof()
# -
#Convert data in the dataframe to float type.
df1drop2 = df1drop.astype(float)
df1drop2.head()
# +
#seperate data sets as labels and features
X = df1drop2.drop('FIRE_DETECTED', axis=1)
y = df1drop2['FIRE_DETECTED']
features = ["brightness", "scan", "track", "satellite", "confidence", "version", "bright_t31", "frp", "daynight", "lat", "long", "hour_x", "day", "month", "year"]
# +
#Use Yellowbrick's Rank2D to create a Pearson ranking of features.
from yellowbrick.features import Rank2D
fig, ax = plt.subplots(1, figsize=(12, 12))
vzr = Rank2D(ax=ax)
vzr.fit(X, y)
vzr.transform(X)
sns.despine(left=True, bottom=True)
vzr.poof()
# +
#Analyze parallel coordinates for our features using Yellowbricks' ParallelCoordinates.
from yellowbrick.features import ParallelCoordinates
features = ["brightness", "scan", "track", "satellite", "confidence", "version", "bright_t31", "frp", "daynight", "lat", "long", "hour_x", "day", "month", "year"
""
]
classes = ["True", "False"]
# Instantiate the visualizer
visualizer = ParallelCoordinates(
classes=classes, features=features, sample=0.05,
shuffle=True, size=(1080, 720)
)
# Fit and transform the data to the visualizer
visualizer.fit(X, y)
visualizer.transform(X)
# Finalize the title and axes then display the visualization
visualizer.show()
# +
#Analyze parallel coordinates for our features using Yellowbricks' ParallelCoordinates, using a data normalizer this time.
visualizer = ParallelCoordinates(
classes=classes, features=features,
normalize='standard', # This time we'll specify a normalizer
sample=0.05, shuffle=True, size=(1080, 720)
)
# Fit the visualizer and display it
visualizer.fit(X, y)
visualizer.transform(X)
visualizer.show()
# -
#Analyze feature importance using Sklearn's ExtraTreesClassifier.
from sklearn.ensemble import ExtraTreesClassifier
array = df1drop2.values
X = array[:,0:41663]
Y = array[:,15]
model = ExtraTreesClassifier(n_estimators = 100)
model.fit(X, Y)
print(model.feature_importances_)
| ML/Merge Table 1 Feature Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Data analysis on EDGAR Asset Backed Security(ABS) SEC Filings
# +
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['font.sans-serif']=['SimHei']
# -
df = pd.read_csv(r'D:\data\edgar\edgar_abs_universal.csv')
df
df['year'] = df['effective'].apply(lambda effective: str(effective)[0:4])
df['year'].dropna(inplace=True)
df['year'] = df['year'].apply(lambda year: int(year))
df
recentDf = df[df['year'] > 2014]
recentDf = recentDf[recentDf['doc_type'] != 'GRAPHIC']
recentDf = recentDf[recentDf['file_desc'] != 'Complete submission text file']
recentDf
recentDf.to_csv(r'D:\data\edgar\edgar_abs_since_2015.csv',index=False)
# +
# SPV 层面分析
spv_df = df[['cik','company','loc']]
spv_df = spv_df.drop_duplicates()
spv_df.set_index('cik', inplace=True)
def get_company_category(company):
company = company.lower()
if 'student' in company or 'educat' in company:
return 'ABS - Student Loan'
if 'card' in company:
return 'ABS - CARD'
if 'auto' in company or 'vehicle' in company or 'moto' in company or 'toyota' in company:
return 'ABS - AUTO'
if 'mbs' in company:
return 'MBS'
if 'home' in company:
return 'MBS'
if 'mor' in company:
return 'MBS'
if '-bnk' in company:
return 'MBS'
if 'hous' in company:
return 'MBS'
if 'receiv' in company or 'asset' in company:
return 'ABS - Others'
return 'OTHER'
spv_df['company_category'] = spv_df['company'].apply(get_company_category)
# other_df = spv_df[spv_df['company_category'] == 'OTHER']
# print(list(other_df['company']))
spv_group_by_type_df = spv_df.groupby(['company_category']).count()
spv_group_by_type_df.drop('loc', axis=1, inplace=True)
spv_group_by_type_df.columns = ['Count']
spv_group_by_type_df
# -
# products = spv_group_by_type_df['category']
counts = spv_group_by_type_df['Count']
colors = ['#eccc68','#1e90ff','#ff6348','#70a1ff','#2ed573']
fig = counts.plot(kind='pie', # 图形类型
title = 'SPV 类型分布',
fontsize = 20,
autopct='%.1f%%', # 数值标签
colors = colors,
radius = 1, # 饼图半径
startangle = 180, # 初始角度
textprops= {'fontsize':14,'color':'0.5'}
)
fig.axes.title.set_size('20')
# plt.title('SPV 类型分布')
plt.show()
# SPV 公司Filing 文件分析
spv_filing_df = df[['cik','company','filing_id','filing_type','idx_url','filing_desc','effective','file_num']]
spv_filing_df.drop_duplicates(inplace=True)
spv_filing_count_df = spv_filing_df.groupby(['company']).count()
spv_filing_count_df = spv_filing_count_df['cik']
# spv_filing_count_df.columns = ['Count']
spv_filing_count_df.sort_values(ascending=False).head(10)
# SPV Filing 数量分布
filing_cnt_df = spv_filing_df.groupby(['company']).count()
filing_bins = [0,5,10,15,20,25,30,40,50,100,500,1000]
filing_cnt_df['category'] = pd.cut(filing_cnt_df['cik'], filing_bins)
by_filing_count = filing_cnt_df.groupby('category').count()
by_filing_count['cik'].plot.bar(rot=45)
# SPV 公司Filing 时间分析
spv_filing_df['filing_year'] = spv_filing_df['effective'].apply(lambda effective: str(effective)[0:4])
by_year_count = spv_filing_df.groupby('filing_year').count()
by_year_count.drop(['nan'],inplace=True)
by_year_count.index = by_year_count.index.astype('int64')
by_year_count['cik'].plot(kind= 'line',xticks = by_year_count.index, rot=45)
by_type = spv_filing_df.groupby('filing_type').count()
by_type.reset_index(inplace=True)
by_type['category'] = by_type.apply(lambda row: 'OTHERS' if row['cik'] < 2000 else row['filing_type'], axis=1)
by_type = by_type.groupby('category').sum()
file_type_cnt = by_type['cik']
file_type_cnt.name = 'count'
file_type_cnt = file_type_cnt.sort_values(ascending=False)
file_type_cnt.plot(kind='pie',
autopct='%.1f%%', # 数值标签
colors = colors,
# radius = 2, # 饼图半径
startangle = 180, # 初始角度
textprops= {'fontsize':12,'color':'#2f3542'}
)
# Filing Files 分析
spv_files = df[['cik','company','effective','filing_id','filing_type','file_idx']]
spv_files_df = spv_files.groupby(['cik','company','filing_id','effective','filing_type']).count().reset_index()
spv_files_df.sort_values('file_idx', ascending=False).head(10)
# Filing File 数量分布
files_cnt_bins = [0,1,2,3,4,5,6,7,8,9,10,20,30,40,50,100,1000]
spv_files_df['cate'] = pd.cut(spv_files_df['file_idx'], files_cnt_bins)
by_spv_files_count = spv_files_df.groupby('cate').count()
by_spv_files_count['cik'].plot.bar(rot=45)
import pandas as pd
df = pd.DataFrame([10,20,30,40], columns=['nums'],index=['a','b','c','d'])
df
df.index = ['a','b','b','d']
df
df.loc['b']
df['nums']
| analysis/edgar-abs-filings-universe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import keras
import matplotlib.pyplot as plt
import csv
import cv2
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
from keras.models import *
from keras.layers import *
from keras.optimizers import Adam
# %matplotlib inline
# +
#data folder
log_path = os.getcwd() + '/data/driving_log.csv'
data_folder = os.getcwd() + '/'
#resized image dimension in training
img_rows = 16
img_cols = 32
#batch size and epoch
batch_size=256
nb_epoch=100
# +
# readin log files
logs = []
with open(log_path,'rt') as f:
reader = csv.reader(f)
for line in reader:
logs.append(line)
log_labels = logs.pop(0)
# +
# image preprocessing, take only the S channel of HSV color space
img = plt.imread(data_folder + (logs[10][0]).strip())
img_processed = (cv2.cvtColor(cv2.resize(img,(32,16)), cv2.COLOR_RGB2HSV))[:,:,1]
plt.subplot(2,1,1)
plt.imshow(img)
plt.subplot(2,1,2)
plt.imshow(img_processed,cmap='gray')
# -
def image_preprocessing(img):
"""preproccesing training data to keep only S channel in HSV color space, and resize to 16X32"""
resized = cv2.resize((cv2.cvtColor(img, cv2.COLOR_RGB2HSV))[:,:,1],(img_cols,img_rows))
return resized
def load_data(X,y,data_folder):
log_path = data_folder + 'data/' + 'driving_log.csv'
logs = []
with open(log_path,'rt') as f:
reader = csv.reader(f)
for line in reader:
logs.append(line)
log_labels = logs.pop(0)
for i in range(len(logs)):
img_path = logs[i][0]
img_path = data_folder+'/data/IMG'+(img_path.split('IMG')[1]).strip()
img = plt.imread(img_path)
X.append(image_preprocessing(img))
y.append(float(logs[i][2]))
# +
data={}
data['features'] = []
data['labels'] = []
load_data(data['features'], data['labels'],data_folder)
# -
X_train = np.array(data['features']).astype('float32')
y_train = np.array(data['labels']).astype('float32')
X_train = np.append(X_train,X_train[:,:,::-1],axis=0)
y_train = np.append(y_train,-y_train,axis=0)
X_train, y_train = shuffle(X_train, y_train)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, random_state=0, test_size=0.1)
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols, 1)
X_val = X_val.reshape(X_val.shape[0], img_rows, img_cols, 1)
# +
model = Sequential([
Lambda(lambda x: x/127.5 - 1.,input_shape=(img_rows,img_cols,1)),
Conv2D(2, 3, 3, border_mode='valid', input_shape=(img_rows,img_cols,1), activation='relu'),
MaxPooling2D((4,4),(4,4),'valid'),
Dropout(0.25),
Flatten(),
Dense(1)])
model.summary()
# -
model.compile(loss='mean_squared_error',optimizer=Adam(1e-5))
history = model.fit(X_train, y_train,batch_size=batch_size, nb_epoch=nb_epoch,verbose=1, validation_data=(X_val, y_val))
| iPythonNb/TinyModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## N PROTEIN BINDING MODELLING
# In this notebook, I will create a model for predicting the binding affinity of coronavirus N_proteins to host-cell RNA. A base model would be built for comparison purposes using the whole sequence of detected RNAs. However, the main model would use only sequences of the binding regions of N_protein. The dataset to be used is a CRAC data which has binding affinities of a particular N protien to a particular mRNA with 9522 RNAs and 4 N proteins.
# from sklearn. Also, the gene sequences are in fasta format; `biomart_transcriptome_all.fasta`. CRAC data is in `SB20201008_hittable_unique.xlsx`.
# First I import the relevant libraries for the work
# +
# python
import itertools
import joblib
# sklearn
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import OrdinalEncoder, OneHotEncoder
# data processing and visualisation
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from Bio import SeqIO
from scipy import stats
# -
# Next I write helper function; `possible_kmers` to output the possible kmers of *ATGC* of length k and `generate_kmers` to generate kmers from an input sequence
# +
def possible_kmers(k):
"""All permutation of DNA sequence based on size
Args:
- k (int); number of bases
Return:
- (list) all possible kmers of lenght k
"""
kmers = []
for output in itertools.product('ATGC', repeat=k):
kmers.append(''.join(list(output)))
return kmers
def generate_kmers(sequence, window, slide=1):
"""Make kmers from a sequence
Args:
- sequence (str); sequence to compute kmer
- window (int); size of kmers
- slide (int); no of bases to move 'window' along sequence
default = 1
Return:
- 'list' object of kmers
Example:
- >>> DNA_kmers('ATGCGTACC', window=4, slide=4)
['ATGC', 'GTAC']
"""
all_possible_seq = []
kmers = []
for base in range(0, len(sequence), slide): # indices
# extend by window
all_possible_seq.append(sequence[base:base + window])
# remove all kmers != window
for seq in all_possible_seq:
if len(seq) == window:
kmers.append(seq)
return kmers
# -
# Next, I read and process the data, `biomart_transcriptome_all` which contains all the transcripts and sequence of humans. Also, I read the CRAC data `SB20201008_hittable_unique.xlsx`
# +
# read and parse the fasta file as SeqIO object
file = SeqIO.parse(
'biomart_transcriptome_all.fasta',
'fasta'
)
# select ids and corresponding sequences
sequence_ids = []
sequences = []
for gene in file:
sequence_ids.append(gene.id)
sequences.append(str(gene.seq))
# create a table of gene ids; select only gene short name and type
id_tab = pd.Series(sequence_ids).str.split('|', expand=True).iloc[:, 2]
# join gene_id_tab with corresponding seqs
transcripts = pd.concat([id_tab, pd.Series(sequences)], axis=1)
# set column names
transcripts.columns = ['gene', 'seq']
# read N_protein CRAC data
N_protein = pd.read_excel('SB20201008_hittable_unique.xlsx', sheet_name='rpm > 10')
# -
# Next, I select for transcripts that appear in N_protein data from the transcriptome data. I also remove duplicated transcripts.
# +
# select common genes between N_protein and transcripts
N_genes = set(N_protein['Unnamed: 0'])
t_genes = set(transcripts.gene)
common_genes = N_genes.intersection(t_genes)
# filter transcripts data with common genes and remove duplicates
transcripts_N = transcripts.drop_duplicates(
subset='gene').set_index('gene').loc[common_genes]
# -
transcripts_N[1, 'seq']
# Next I use the `generate_kmers` function to make kmers from each sequence
# +
# create kmers from seq
transcripts_N['kmers'] = transcripts_N.seq.apply(generate_kmers, window=4, slide=4)
# view of kmers data
transcripts_N.kmers
# -
# From the output, it can be seen that the kmers have been produced nicely. Next, I would seperate each kmer into a feature and pad the short sequences with `'_'`
# seperate kmers into columns. pad short seqs with '_'
kmer_matrix = transcripts_N.kmers.apply(pd.Series).fillna('_')
# Now I can use `sklearn.OneHotEncoder` to convert my strings to floats for my feature matrix `ohe_kmers` and create my response vector `y` from `133_FH-N_229E` values in the CRAC data
# +
# convert kmers to ints
ohe = OneHotEncoder(sparse=True)
ohe_kmers = ohe.fit_transform(kmer_matrix)
# response vector
y = pd.concat([kmer_matrix[0],
N_protein.drop_duplicates(subset='Unnamed: 0').set_index('Unnamed: 0')], axis=1)['133_FH-N_229E']
# -
# Next, I split the data into **80%** training and **20%** testing testing sets
# split data into train and test sets
XTrain, XTest, yTrain, yTest = train_test_split(ohe_kmers, y, test_size=0.2, random_state=1)
# Now I am ready to train the model. I would use `sklearn.linear_model.LinearRegression` as my algorithm and use `r2_score` as my evaluation metric
# +
# instantiate the regressor
linreg = LinearRegression()
# train on data
linreg.fit(XTrain, yTrain)
# check performance on test set
yPred = linreg.predict(XTest)
metrics.r2_score(y_true=yTest, y_pred=yPred)
# -
# An `r2_score` of **0.71** is not bad for a base model. Next, I can save the model as a file to avoid retraining it.
# save model
_ = joblib.dump(linreg, 'BaseModel.sav')
# Next, I make a correlation plot of my predicted and testing values
# +
# plot of yTest vs yPred
g = sns.regplot(x = yTest, y = zzz, scatter_kws={'alpha':0.2})
# set axes labels
_ = plt.xlabel('yTest')
_ = plt.ylabel('yPred')
# pearson correlation test
r, p = stats.pearsonr(yTest, yPred)
_ = g.annotate('r={}, p={}'.format(r, p), (-8, 2))
# -
# Surprisingly, the pearson correlation was **0.72** with a significant p-value
#
# Next, I would use a peak calling program to select the actual sequence to which the N_proteins bind on the RNA. Hopefully that would produce a model.
plt.figure(figsize=(10, 8))
sns.barplot(x='Kmer Encoding', y='Pearson Correlation', data=kmer_data, color='blue')
plt.xticks(rotation=45, size=15)
plt.ylabel('Pearson Correlation ', size=20, rotation=360)
plt.xlabel('Kmer Encoding Type', size=20)
| N_Protein_Modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generate `nearest` file
#
# Compare synthetic PUF to its training and holdout sets.
# ## Setup
#
# ### Imports
import pandas as pd
import numpy as np
import synthimpute as si
import synpuf
# **UPDATE!**
SYNTHESIS_ID = 13
PCT_TRAIN = 50
# Folders.
PUF_SAMPLE_DIR = '~/Downloads/puf/'
SYN_DIR = '~/Downloads/syntheses/'
NEAREST_DIR = '~/Downloads/nearest/'
# ### Load data
synth = pd.read_csv(SYN_DIR + 'synpuf' + str(SYNTHESIS_ID) + '.csv')
train = pd.read_csv(PUF_SAMPLE_DIR + 'train' + str(PCT_TRAIN) + '.csv')
test = pd.read_csv(PUF_SAMPLE_DIR + 'test' + str(100 - PCT_TRAIN) + '.csv')
# ## Preprocessing
# Drop calculated features used as seeds, and drop s006.
synpuf.add_subtracted_features(train)
synpuf.add_subtracted_features(test)
DROPS = ['S006', 'e00600_minus_e00650', 'e01500_minus_e01700',
'RECID', 'E00100', 'E09600']
train.drop(DROPS, axis=1, inplace=True)
test.drop(DROPS, axis=1, inplace=True)
synth.columns = [x.upper() for x in synth.columns]
synth = synth[train.columns]
synth.reset_index(drop=True, inplace=True)
train.reset_index(drop=True, inplace=True)
test.reset_index(drop=True, inplace=True)
# ## Nearest calculation
#
# Compare nearest standardized Euclidean distance. Takes ~10 hours.
# %%time
nearest = si.nearest_synth_train_test(synth, train, test)
nearest.to_csv(NEAREST_DIR + 'nearest' + str(SYNTHESIS_ID) + '.csv',
index=False)
| analysis/disclosure/nearest13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An Analysis of Obesisy in First Class Children in Ireland
#
# ## Introduction
#
# In this notebook I am creating a dataset to simulate a hypothetical study on the BMI of primary school first class children in 2016 based on their gender, age, height, and weight. I have based the simulation on a subset of the findings of the Childhood Obesity Surveillance Initiative (COSI).
#
# The National Nutrition Centre in UCD was commisioned by the HSE to carry out the surveillance work in COSI. This research was undertaken as part of the World Health Organsiation European Childhood Obesity Surveillance Initiative. This survey has been conducted to date over four waves in 2008; 2010; 2012; and 2015. The data collected has been based on over 17,000 examinations in over 150 randomly selected primary schools. [Ref I COSI](https://www.hse.ie/eng/about/who/healthwellbeing/our-priority-programmes/heal/heal-docs/cosi-in-the-republic-of-ireland-findings-from-2008-2010-2012-and-2015.pdf)
#
# Overall the trend shows the levels of overweight and obesity in first class children is stabilising, although at a high level. In non-Deis schools the levels have fallen slightly, whereas in Deis schools levels continue to rise. More girls tend to be overweight than boys. Summary findings from the 2012 study are outlined below:
#
# [COSI Report 2012 Ref II](http://www.ucd.ie/t4cms/COSI%20report%20(2014).pdf)
#
#
#
# | BMI Classification |Thin-Normal | Overweight | Obese |
# | :-----------: |:----: |:-------------: | :-------: |
# | First Class Boys | 85.6% |12.2% | 2.2% |
# | First Class Girls | 78.6% |15.9% | 5.5% |
#
#
# I used the 2012 report to generate the BMI and weight variables, as it broke out the results by gender.
#
# +
# import python packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# magic command to use matplotlib within this notebook
# %matplotlib inline
# -
# ## Variables
#
# The dataset has a population of 1000. It contains the following variables:
#
#
# ### Names
# I created two random lists of names, by gender, combining the most popular childrens names with the most popular Irish surnames. 500 names were selected randomly for each gender.
#
#
# ### Gender
# The gender was generated in the dataset based on the condition, whether or not the name of the pupil was contained in the girls list.
#
#
# ### Age
# The ages of the children in the study I a focusing on are those in first class. The ages are chosen randomly from a list I generated, based on the 2016 first class ages breakdown from the CSO below [[Ref III]](https://www.cso.ie/px/pxeirestat/Statire/SelectVarVal/Define.asp?maintable=EDA42&PLanguage=0):
#
# 6 Years old 34%
# 7 Years old 65%
# 8 Years old 01%
#
#
# +
"""
NAMES
List for the random name generator for the dataset. I am generating two lists. A list of sample girls and boys names
from which a random selection will be chosen for the dataset.
"""
# The first names were extracted from excel from the CSO website, the most popular baby names from 2012 (https://www.cso.ie/en/releasesandpublications/er/ibn/irishbabiesnames2012/")
girl_firstname = ('Emily ','Sophie ','Emma ','Grace ','Lily ','Mia ','Ella ','Ava ','Lucy ','Sarah ','Aoife ','Amelia ','Hannah ','Katie ','Chloe ','Caoimhe ','Saoirse ','Kate ','Holly ','Ruby ','Sophia ','Anna ','Lauren ','Leah ','Amy ','Isabelle ','Molly ','Ellie ','Jessica ','Olivia ','Roisin ','Ciara ','Kayla ','Julia ','Zoe ','Laura ','Niamh ','Abbie ','Erin ','Rachel ','Robyn ','Aisling ','Faye ','Rebecca ','Eva ','Layla ','Ellen ','Cara ','Freya ','Abigail ','Eve ','Isabella ','Megan ','Aine ','Clodagh ','Aoibhinn ','Millie ','Nicole ','Aoibheann ','Maja ','Sadhbh ','Eabha ','Charlotte ','Amber ','Caitlin ','Sofia ','Alannah ','Zara ','Alice ','Maria ','Elizabeth ','Lena ','Mary ','Emilia ','Aimee ','Lilly ','Hollie ','Aoibhe ','Victoria ','Eimear ','Maya ','Isabel ','Orla ','Evie ','Kayleigh ','Brooke ','Clara ','Meabh ','Lexi ','Tara ','Daisy ','Katelyn ','Ailbhe ','Amelie ','Natalia ','Sara ','Hanna ','Laoise ','Ruth ','Madison ','Maeve ','Maisie ','Rose ',)
boy_firstname = ('Jack ','James ','Daniel ','Sean ','Conor ','Adam ','Harry ','Ryan ','Dylan ','Michael ','Luke ','Charlie ','Liam ','Oisin ','Cian ','Jamie ','Thomas ','Alex ','Noah ','Darragh ','Patrick ','Aaron ','Cillian ','Matthew ','John ','Nathan ','David ','Fionn ','Evan ','Ethan ','Jake ','Kyle ','Rian ','Ben ','Max ','Eoin ','Tadhg ','Finn ','Callum ','Samuel ','Joshua ','Rory ','Jayden ','Joseph ','Tyler ','Sam ','Shane ','Mark ','Robert ','Aidan ','William ','Ronan ','Eoghan ','Alexander ','Leon ','Cathal ','Mason ','Tom ','Oliver ','Andrew ','Oscar ','Ciaran ','Bobby ','Jacob ','Senan ','Rhys ','Scott ','Benjamin ','Cormac ','Kevin ','Lucas ','Alan ','Donnacha ','Jakub ','Christopher ','Filip ','Killian ','Josh ','Alfie ','Tommy ','Ruairi ','Odhran ','Oran ','Leo ','Isaac ','Dara ','Jason ','Zach ','Martin ','Peter ','Brian ','Danny ','Niall ','Tomas ','Edward ','Stephen ','Logan ','Kacper ','Anthony ','Billy ',)
# Surnames top 100 Irish surnames from (https://meanwhileinireland.com/ranked-top-100-irish-surnames-and-meanings/)
surname = ('Murphy','Kelly','<NAME>','Walsh','Smith','<NAME>','Byrne','Ryan','<NAME>','<NAME>','<NAME>','Doyle','McCarthy','Gallagher','O Doherty','Kennedy','Lynch','Murray','Quinn','Moore','McLoughlin','O Carroll','Connolly','Daly','O Connell','Wilson','Dunne','Brennan','Burke','Collins','Campbell','Clarke','Johnston','Hughes','O Farrell','Fitzgerald','Brown','Martin','Maguire','Nolan','Flynn','Thompson','O Callaghan','O Donnell','Duffy','O Mahony','Boyle','Healy','O Shea','White','Sweeney','Hayes','Kavanagh','Power','McGrath','Moran','Brady','Stewart','Casey','Foley','Fitzpatrick','<NAME>','McDonnell','MacMahon','Donnelly','Regan','Donovan','Burns','Flanagan','Mullan','Barry','Kane','Robinson','Cunningham','Griffin','Kenny','Sheehan','Ward','Whelan','Lyons','Reid','Graham','Higgins','Cullen','Keane','King','Maher','MacKenna','Bell','Scott','Hogan','O Keeffe','Magee','MacNamara','MacDonald','MacDermott','Molony','<NAME>','Buckley','O Dwyer',)
# Create empty lists for girls and boys names
girlsname = []
boysname = []
# Generate a list of girls names - this list contains 10300 girlsnames
for i in girl_firstname:
for j in surname:
girlsname.append(i+j)
# Generate a list of boys names - the list contains 10,000 boysnames
for i in boy_firstname:
for j in surname:
boysname.append(i+j)
# Randomly choose 500 girls and the 500 boys names from the lists above
gnames = (np.random.choice(girlsname,500))
bnames = (np.random.choice(boysname,500))
# Combine the two lists into a single array
names = np.concatenate([gnames,bnames])
"""
AGE
List for age generator for the dataset. Randomely generate 1000 ages of 1st class students based on the CSO 2016 first class ages.
"""
age = np.random.choice([6,7,8], 1000, p=[0.34, 0.65, 0.01])
plt.hist(age, bins=3)
plt.title("Ages")
plt.xlabel("Years")
plt.ylabel("Number")
plt.show()
# -
# ### Height
# The height of each child is randomly selected based on age and gender. I have used the UK WHO growth charts as a basis for these as Ireland is moving to this model currently. Data is not available in Ireland yet for children ages 6 -8.[Ref VI](https://www.hse.ie/eng/health/child/growthmonitoring/)
#
# [[Ref IV]](https://www.rcpch.ac.uk/sites/default/files/Boys_2-18_years_growth_chart.pdf)
#
# | Boys Aged | Min cm | Median cm | Max cm |
# | :-----------: |:-------------: | :-------: | :----: |
# | 6 | 103 | 119 | 129 |
# | 7 | 108 | 122 | 136 |
# | 8 | 113 | 128 | 142 |
#
#
# [[Ref V]](https://www.rcpch.ac.uk/sites/default/files/Girls_2-18_years_growth_chart.pdf)
#
#
# | Girls Aged | Min cm | Median cm | Max cm |
# | :-----------: |:-------------: | :-------: | :----: |
# | 6 | 102 | 115 | 128 |
# | 7 | 107 | 121 | 135 |
# | 8 | 113 | 127 | 141 |
#
# It is widely accepted that human heights follow a normal distribution. (See <NAME>. Foetus Into Man: Physical Growth From Conception to Maturity. Cambridge, MA: Harvard University Press; 1990. & Snedecor GW, Cochran WG. Statistical Methods. Ames, IA: Iowa University; 1989. )I have used the numpy.random normal package to generate the heights based on gender and age according to the tables above. VII. [Ref VII](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2831262/#b27-dem-46-0001)
#
# The python code to generate heights is below:
#
# +
"""
HEIGHTS
Generate a random sample of lists of heights based on gender and age, on the normal distribution - per the UK WHO growth charts.
The random lists are generated for each age and gender. The height function randomly selects
"""
height_boys_6 = np.round( np.random.normal(1.19, 0.029,10000 ), 2)
height_boys_7 = np.round( np.random.normal(1.22, 0.032,10000 ), 2)
height_boys_8 = np.round( np.random.normal(1.28, 0.032,10000 ), 2)
height_girls_6 = np.round( np.random.normal(1.15, 0.031,10000 ), 2)
height_girls_7 = np.round( np.random.normal(1.21, 0.032,10000 ), 2)
height_girls_8 = np.round( np.random.normal(1.27, 0.032,10000 ), 2)
"""
This function is called by the dataset to populate the height column. The random.choice function is used to randomly
select a height from the correct list above dependig on age and gender
"""
def height (row):
if row['Age']==6:
if row['Gender'] == 'Female':
return np.random.choice(height_girls_6,1)
if row['Age']==6:
if row['Gender'] == 'Male':
return np.random.choice(height_boys_6,1)
if row['Age']==7:
if row['Gender'] == 'Female':
return np.random.choice(height_girls_7,1)
if row['Age']==7:
if row['Gender'] == 'Male':
return np.random.choice(height_boys_7,1)
if row['Age']==8:
if row['Gender'] == 'Female':
return np.random.choice(height_girls_8,1)
if row['Age']==8:
if row['Gender'] == 'Male':
return np.random.choice(height_boys_8,1)
# -
# ### BMI
#
# For the purpose of this simulation I have broken BMI into 3 categories: Thin-Normal; Overweight; Obese.
#
# BMI is calculated using the same formula for adults and children.
#
# Formula: BMI = weight (kg) / [height (m)]2
#
# Adult BMI cutoff's are standard. For children it depends on gender and age. I used the childrens cutoffs at their age plus 6 months. [Adult BMI Ref VIII](https://en.wikipedia.org/wiki/Body_mass_index)
# [Boys BMI Ref IX](https://www.who.int/growthref/sft_bmifa_boys_z_5_19years.pdf?ua=1)
# [Girls BMI Ref X](https://www.who.int/growthref/sft_bmifa_girls_z_5_19years.pdf?ua=1)
#
#
#
# | BMI Cut-Offs | Thin-Normal | Overweight | Obese |
# | :-----------: |:-------------: | :-------: | :----: |
# | Adult | 15- 25 | 25 - 30 | 30-60 |
# | Girl aged 6 | 11.7 - 17.1| 17.1 - 19.5 | 19.5 - 30 |
# | Girl aged 7 | 11.8 - 17.5 | 17.5 - 20.1 | 20.1 - 30 |
# | Girl aged 8 | 12.0- 18.0 | 18.0 - 21.0 | 21.0 - 30 |
# | Boy aged 6 | 12.2 - 16.9 | 16.9 -18.7 | 18.7 - 30 |
# | Boy aged 7 | 12.3 - 17.2 | 17.2 - 19.3 | 19.3 - 30 |
# | Boy aged 8 | 12.5 - 17.7 | 17.7 - 20.1 | 20.1 - 30 |
#
# I created the BMI's in two steps. The classification was generated first (based on gender and % from COSI 2012), and then BMI score generated depending on this classification, the student's age, and gender.
#
# A variety of different studies suggest BMI follows different probability distributions: normal; log normal, skew student t etc. There is no definitive study on children's BMIs distribution. [Ref XI](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1636707/)I have chosen log normal from the numpy.random package to generate the BMI data.
#
# The COSI study in 2012 recorded BMI in first class students with a median value of 16.25; min value of 12.3; and max value of 26.1 averaged for boys and girls. [Ref II](http://www.ucd.ie/t4cms/COSI%20report%20(2014).pdf)
#
# Creating the BMI data
# 1. I used log normal numpy.random to generate a list of values for BMI.
# 2. I filtered this list excluding all values outside the min and max BMIs of the 2012 COSI survey.
# 3. 18 individual lists were created for each classification gender age combination based on the WHO BMI cutoffs in the table above.
# 4. A function was written to check gender, age, and classification and chose a random sample from the appropriate list.
#
# The code generating the BMI is below:
#
# +
"""
BMI CLASSIFICATION
BMI classification generator was based on the percentages in the 2012 study in the introduction.
I generated a 1000 random classification list according to the proportions from the 2012 study.
"""
girlBmiClass = np.random.choice(['Thin-Normal','Overweight','Obese'], 1000, p=[0.786, 0.159, 0.055])
boyBmiClass = np.random.choice(['Thin-Normal','Overweight','Obese'], 1000, p=[0.856, 0.122, 0.022])
"""
This function is called by the dataset to randomly select a BMI class from the lists above based on gender.
"""
def bmic (row):
# if gender female - randomly choose a classification from the girls BMI class list
if row['Gender'] == 'Female':
return np.random.choice(girlBmiClass,1)
if row['Gender'] == 'Male':
return np.random.choice(boyBmiClass,1)
"""
BMI
Generate a random sample of lists of BMI based on gender and age, on the log normal distribution - per the UK WHO BMI growth
cutoffs.
The random lists are generated for each unique age, gender and BMI classification. The get BMI function randomly selects
elements from the correct list according to age, gender and BMI classification.
"""
# Create a log normal random list rounded to one decimal place
mu, sigma = 16.25 , 2 # mean and standard deviation - standard deviation was chosen by trial and error
bmiNum = np.round(np.random.normal(mu, sigma, 10000),1)
#Filter out the numbers outside the min and max values
bmiNum = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 26.1)]
# filter the lists per the WHO cutoffs by category. TN -Thin-Normal; OW - Overweight; OB - Obese
bmi_TN_boys_6 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 16.9)]
bmi_TN_boys_7 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 17.2)]
bmi_TN_boys_8 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 17.7)]
bmi_TN_girls_6 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 17.1)]
bmi_TN_girls_7 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 17.5)]
bmi_TN_girls_8 = bmiNum [(bmiNum >= 12.3) & (bmiNum <= 18.0)]
bmi_OW_boys_6 = bmiNum [(bmiNum > 16.9) & (bmiNum <= 18.7)]
bmi_OW_boys_7 = bmiNum [(bmiNum > 17.2) & (bmiNum <= 19.3)]
bmi_OW_boys_8 = bmiNum [(bmiNum > 17.7) & (bmiNum <= 20.1)]
bmi_OW_girls_6 = bmiNum [(bmiNum > 17.1) & (bmiNum <= 19.5)]
bmi_OW_girls_7 = bmiNum [(bmiNum > 17.5) & (bmiNum <= 20.1)]
bmi_OW_girls_8 = bmiNum [(bmiNum > 18.0) & (bmiNum <= 21.0)]
bmi_OB_boys_6 = bmiNum [(bmiNum > 18.7) & (bmiNum <= 26.1)]
bmi_OB_boys_7 = bmiNum [(bmiNum > 19.3) & (bmiNum <= 26.1)]
bmi_OB_boys_8 = bmiNum [(bmiNum > 20.1) & (bmiNum <= 26.1)]
bmi_OB_girls_6 = bmiNum [(bmiNum > 19.5) & (bmiNum <= 26.1)]
bmi_OB_girls_7 = bmiNum [(bmiNum > 20.1) & (bmiNum <= 26.1)]
bmi_OB_girls_8 = bmiNum [(bmiNum > 21.0) & (bmiNum <= 26.1)]
"""
This function is called by the dataset to populate the bmi column. The random.choice function is used to randomly
select a bmi from the correct list above dependig on age, gender and BMI classification.
"""
def bmi (row):
#Thin-Normal BMI classification
if row['Age']==6:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_girls_6,1)
if row['Age']==6:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_boys_6,1)
if row['Age']==7:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_girls_7,1)
if row['Age']==7:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_boys_7,1)
if row['Age']==8:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_girls_8,1)
if row['Age']==8:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Thin-Normal':
return np.random.choice(bmi_TN_boys_8,1)
# Overweight Classification
if row['Age']==6:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_girls_6,1)
if row['Age']==6:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_boys_6,1)
if row['Age']==7:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_girls_7,1)
if row['Age']==7:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_boys_7,1)
if row['Age']==8:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_girls_8,1)
if row['Age']==8:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Overweight':
return np.random.choice(bmi_OW_boys_8,1)
# Obese Classification
if row['Age']==6:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_girls_6,1)
if row['Age']==6:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_boys_6,1)
if row['Age']==7:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_girls_7,1)
if row['Age']==7:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_boys_7,1)
if row['Age']==8:
if row['Gender'] == 'Female':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_girls_8,1)
if row['Age']==8:
if row['Gender'] == 'Male':
if row['BMI Class'] == 'Obese':
return np.random.choice(bmi_OB_boys_8,1)
# plot the overall BMI classification lists by gender
plt.subplot(1, 2, 1)
plt.title("Girls")
plt.hist(girlBmiClass)
plt.subplot(1, 2, 2)
plt.title("Boys")
plt.hist(boyBmiClass)
# -
# ### Weight
#
# To generate the BMI I used the classification porportions from the COSI 2012 survey.
#
# I calculated the weight in Kg from the height and BMI of each first class student, using the values in the columns and the formula below:
#
# Formula: weight (kg) = BMI * [height (m)]2
#
#
# ## Dataset Creation
#
# The dataset is created in the cell below, populated using the variables and functions coded above. Note the values in columns: Age is in years; Height is in meters; & Weight is in kg.
#
# There was an error when I attempted to change the datatype of the column Weight from object to float. I commented out this line of code as I have been unable to reproduce this error.
#
#
# +
# Create the dataset containing the list of names and ages and an empy column for each of the other variables.
d = {'Name': names, 'Age': age, 'Gender': '', 'Height': '', 'Weight': '', 'BMI Class': '', 'BMI' : '' }
df = pd.DataFrame(data=d)
# Populate the gender variable in the dataset. If the name is part of the gnames list return female, else return male.
df['Gender'] = np.where(np.isin(df['Name'],gnames), 'Female', 'Male')
# Populate the height variable
df['Height'] = df.apply(lambda row: height(row),axis=1)
# Change datatype to float
df['Height'] = df['Height'].str[0]
# Populate the BMI classification variable according to the 2012 proportions
df['BMI Class'] = df.apply(lambda row: bmic(row),axis=1)
df['BMI Class'] = df['BMI Class'].str[0]
# Populate the bmi variable
df['BMI'] = df.apply(lambda row: bmi(row),axis=1)
# Change datatype to float
df['BMI'] = df['BMI'].str[0]
#Populate the weight variable based on the BMI value and height
df['Weight'] = (df['Height']*df['Height']*df['BMI'])
# Change datatype to float - caused error when cell rerun
#df['Weight'] = df['Weight'].str[0]
# Round weights to one decimal place
df['Weight'] = round(df['Weight'],1)
df
# -
# ## Analysis
#
# I have analysed the dataset below, both graphically and using the summary stats of the dataframe. For the summary stats I have split the dataframe into two, one for girls and one for boys. The second table for both girls and boys was to show a count of the BMI breakdown by gender.
#
# **Summary stats of the numerical variables in the dataset**
#
# **Girls summary statistics**
# +
# Create a subset of the dataframe by gender
df_girl = (df[(df['Gender'] == 'Female')])
df_boy = (df[(df['Gender'] == 'Male')])
# Girsl summary stats
df_girl.describe()
# -
# Display a count of each BMI class for girls
grpG = df_girl.groupby('BMI Class')
grpG.describe()
# **Boys summary statistics**
#Boys summary stats
df_boy.describe()
# Display a count of each BMI class
grpB = df_boy.groupby('BMI Class')
grpB.describe()
# To visualise the releationships I have created 5 plots of the dataset below. The BMI classification, height weight and age followed the expected trends. However due to how the data is randomly generated the data does not fully mirror the original survey.
#
# Visually you can see the trend of a higher number of obese and overweight girls than boys. This is most clearly evident on the Obese plot of the three plots side by side below. Where each plot is by BMI classification type.
# Plot BMI classification by gender
sns.lmplot('Height', 'Weight', data=df, fit_reg=False, hue='BMI Class', col='Gender')
# +
sns.lmplot('Height', 'Weight', data=df, fit_reg=False, hue='Gender', col='BMI Class')
# -
# ## References
#
# I. The Childhood Obesity Surveillance Initiative (COSI) in the Republic of Ireland Findings 2015 [COSI Report 2015](https://www.hse.ie/eng/about/who/healthwellbeing/our-priority-programmes/heal/heal-docs/cosi-in-the-republic-of-ireland-findings-from-2008-2010-2012-and-2015.pdf)
#
# II. The Childhood Obesity Surveillance Initiative (COSI) in the Republic of Ireland Findings 2012 [COSI Report 2012](http://www.ucd.ie/t4cms/COSI%20report%20(2014).pdf)
#
# III. CSO - [Ages of children in first class] (https://www.cso.ie/px/pxeirestat/Statire/SelectVarVal/Define.asp?maintable=EDA42&PLanguage=0)
#
# IV. Heights [UK WHO growth chart boys](https://www.rcpch.ac.uk/sites/default/files/Boys_2-18_years_growth_chart.pdf)
#
# V. Heights [UK WHO growth chart girls](https://www.rcpch.ac.uk/sites/default/files/Girls_2-18_years_growth_chart.pdf)
#
# VI. Heights [Growth monitoring resources](https://www.hse.ie/eng/health/child/growthmonitoring/)
#
# VII. Heights [Normally distributed](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2831262/#b27-dem-46-0001)
#
# VIII. BMI Wiki [BMI Wiki](https://en.wikipedia.org/wiki/Body_mass_index)
#
# IX. WHO BMI Boys [Boys BMI](https://www.who.int/growthref/sft_bmifa_boys_z_5_19years.pdf?ua=1)
#
# x. WHO BMI Girls [Girls BMI](https://www.who.int/growthref/sft_bmifa_girls_z_5_19years.pdf?ua=1)
#
# XI. BMI distribution curve [BMI distribution](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1636707/)
| Childhood Obesity Analysis Ireland.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib widget
import os
import sys
sys.path.insert(0, os.getenv('HOME')+'/pycode/MscThesis/')
import pandas as pd
from amftrack.util import get_dates_datetime, get_dirname, get_plate_number, get_postion_number,get_begin_index
import ast
from amftrack.plotutil import plot_t_tp1
from scipy import sparse
from datetime import datetime
from amftrack.pipeline.functions.node_id import orient
import pickle
import scipy.io as sio
from pymatreader import read_mat
from matplotlib import colors
import cv2
import imageio
import matplotlib.pyplot as plt
import numpy as np
from skimage.filters import frangi
from skimage import filters
from random import choice
import scipy.sparse
import os
from amftrack.pipeline.functions.extract_graph import from_sparse_to_graph, generate_nx_graph, sparse_to_doc
from skimage.feature import hessian_matrix_det
from amftrack.pipeline.functions.experiment_class_surf import Experiment, Edge, Node, Hyphae, plot_raw_plus
from amftrack.pipeline.paths.directory import run_parallel, find_state, directory_scratch, directory_project
from amftrack.notebooks.analysis.util import *
from scipy import stats
from scipy.ndimage.filters import uniform_filter1d
from statsmodels.stats import weightstats as stests
from amftrack.pipeline.functions.hyphae_id_surf import get_pixel_growth_and_new_children
from collections import Counter
from IPython.display import clear_output
from amftrack.notebooks.analysis.data_info import *
# -
exp = get_exp((39,269,329),directory_project)
def get_hyph_infos(exp):
select_hyph = {}
for hyph in exp.hyphaes:
select_hyph[hyph] = []
for i,t in enumerate(hyph.ts[:-1]):
tp1=hyph.ts[i+1]
pixels,nodes = get_pixel_growth_and_new_children(hyph,t,tp1)
speed = np.sum([get_length_um(seg) for seg in pixels])/get_time(exp,t,tp1)
select_hyph[hyph].append((t,hyph.ts[i+1],speed,pixels))
return(select_hyph)
# + jupyter={"outputs_hidden": true}
select_hyph = get_hyph_infos(exp)
# -
rh2 = [hyph for hyph in exp.hyphaes if np.any(np.array([c[2] for c in select_hyph[hyph]])>=300)]
hyph = [rh for rh in rh2 if rh.end.label == 1][0]
# hyph = choice(rh2)
speeds = [c[2] for c in select_hyph[hyph]]
ts = [c[0] for c in select_hyph[hyph]]
tp1s = [c[1] for c in select_hyph[hyph]]
plt.close('all')
plt.rcParams.update({
"font.family": "verdana",
'font.weight' : 'normal',
'font.size': 20})
fig=plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
ax.plot(ts,speeds)
ax.set_xlabel('time (h)')
ax.set_ylabel('speed ($\mu m .h^{-1}$)')
plot_raw_plus(exp,hyph.ts[-1],[hyph.end.label]+[hyph.root.label])
counts = []
for t in range(exp.ts):
count = 0
for hyph in rh2:
if int(hyph.end.ts()[-1])==int(t):
count+=1
counts.append(count)
# + jupyter={"outputs_hidden": true}
counts
# -
plot_raw_plus(exp,hyph.ts[-1]+1,[hyph.end.label]+[hyph.root.label]+[5107,5416])
# +
nx_graph_t = exp.nx_graph[35]
nx_graph_tm1 = exp.nx_graph[34]
Sedge = sparse.csr_matrix((30000, 60000))
for edge in nx_graph_t.edges:
pixel_list = nx_graph_t.get_edge_data(*edge)["pixel_list"]
pixela = pixel_list[0]
pixelb = pixel_list[-1]
Sedge[pixela[0], pixela[1]] = edge[0]
Sedge[pixelb[0], pixelb[1]] = edge[1]
tip = 2326
pos_tm1 = exp.positions[34]
pos_t = exp.positions[35]
mini1 = np.inf
posanchor = pos_tm1[tip]
window = 1000
potential_surrounding_t = Sedge[
max(0, posanchor[0] - 2 * window) : posanchor[0] + 2 * window,
max(0, posanchor[1] - 2 * window) : posanchor[1] + 2 * window,
]
# potential_surrounding_t=Sedge
# for edge in nx_graph_t.edges:
# pixel_list=nx_graph_t.get_edge_data(*edge)['pixel_list']
# if np.linalg.norm(np.array(pixel_list[0])-np.array(pos_tm1[tip]))<=5000:
# distance=np.min(np.linalg.norm(np.array(pixel_list)-np.array(pos_tm1[tip]),axis=1))
# if distance<mini1:
# mini1=distance
# right_edge1 = edge
# print('t1 re',right_edge)
mini = np.inf
for node_root in potential_surrounding_t.data:
for edge in nx_graph_t.edges(int(node_root)):
pixel_list = nx_graph_t.get_edge_data(*edge)["pixel_list"]
if (
np.linalg.norm(np.array(pixel_list[0]) - np.array(pos_tm1[tip]))
<= 5000
):
distance = np.min(
np.linalg.norm(
np.array(pixel_list) - np.array(pos_tm1[tip]), axis=1
)
)
if distance < mini:
mini = distance
right_edge = edge
# -
right_edge,mini
origin = np.array(
orient(
nx_graph_tm1.get_edge_data(*list(nx_graph_tm1.edges(tip))[0])[
"pixel_list"
],
pos_tm1[tip],
)
)
origin_vector = origin[0] - origin[-1]
branch = np.array(
orient(
nx_graph_t.get_edge_data(*right_edge)["pixel_list"],
pos_t[right_edge[0]],
)
)
candidate_vector = branch[-1] - branch[0]
dot_product = np.dot(origin_vector, candidate_vector)
if dot_product >= 0:
root = right_edge[0]
next_node = right_edge[1]
else:
root = right_edge[1]
next_node = right_edge[0]
last_node = root
current_node = next_node
last_branch = np.array(
orient(
nx_graph_t.get_edge_data(root, next_node)["pixel_list"],
pos_t[current_node],
)
)
i = 0
loop = []
while (
nx_graph_t.degree(current_node) != 1
and not current_node in nx_graph_tm1.nodes
): # Careful : if there is a cycle with low angle this might loop indefinitely but unprobable
i += 1
if i >= 100:
print(
"identified infinite loop",
i,
tip,
current_node,
pos_t[current_node],
)
break
mini = np.inf
origin_vector = (
last_branch[0] - last_branch[min(length_id, len(last_branch) - 1)]
)
unit_vector_origin = origin_vector / np.linalg.norm(origin_vector)
candidate_vectors = []
for neighbours_t in nx_graph_t.neighbors(current_node):
if neighbours_t != last_node:
branch_candidate = np.array(
orient(
nx_graph_t.get_edge_data(current_node, neighbours_t)[
"pixel_list"
],
pos_t[current_node],
)
)
candidate_vector = (
branch_candidate[min(length_id, len(branch_candidate) - 1)]
- branch_candidate[0]
)
unit_vector_candidate = candidate_vector / np.linalg.norm(
candidate_vector
)
candidate_vectors.append(unit_vector_candidate)
dot_product = np.dot(unit_vector_origin, unit_vector_candidate)
angle = np.arccos(dot_product)
if angle < mini:
mini = angle
next_node = neighbours_t
if len(candidate_vectors) < 2:
print(
"candidate_vectors < 2",
nx_graph_t.degree(current_node),
pos_t[current_node],
[node for node in nx_graph_t.nodes if nx_graph_t.degree(node) == 2],
)
competitor = np.arccos(np.dot(candidate_vectors[0], -candidate_vectors[1]))
if mini < competitor:
current_node, last_node = next_node, current_node
current_node
| amftrack/notebooks/Draft_1/Figure_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd0fc2c00f0e2c44cb4028bd693f18a1b5d93d1de4cd12db71fca36ff691a163044
# ---
import pandas as pd
import random
# +
def func_random(jugador1,jugador2):
x = random.randint(0,6)
y = random.randint(0,6)
print("1º número:", x)
print("2º número", y)
if x > y:
return jugador1, "empieza"
else:
return jugador2 "empieza"
func_random(input("Nombre 1"),input("Nombre 2"))
# -
tablero1 = pd.DataFrame({1: "~", 2: "~",3: "~",4: "~",5: "~",6: "~",7: "~",8: "~",9: "~",10: "~"},index=[1, 2,3,4,5,6,7,8,9,10])
tablero1
tablero2 = pd.DataFrame({1: "~", 2: "~",3: "~",4: "~",5: "~",6: "~",7: "~",8: "~",9: "~",10: "~"},index=[1, 2,3,4,5,6,7,8,9,10])
tablero2
# +
""" Definimos la Clase Barco con la función colocar barco, la cual coloca los barcos en el tablero:
df = tablero1
size = dimensiones del barco ( pequeño: 2, mediano: 3, grande: 4, gigante: 5)
fil = fila
col = columna"""
class Barco:
def __init__(self, df):
self.df = df
def colocar_barco(self, size, df): # Introducimos el DataFrame (tablero que escogemos) y el size
self.size = size
fil = int(input("Introduce la coordenada fila"))
col = int(input("Introduce la coordenada columna"))
# Si tenemos ya barco en la coordenada introducida:
if df.loc[fil,col] == "#" :
print("Ops! There´s already a boat,try again!")
else:
# Pedimos cómo queremos colocar el barco a partir de la coordenada inicial:
h_v = input("escribe 'h' si quieres horizontal o 'v' si quieres vertical")
# Ponemos las condiciones para que el barco no se sobreponga o no se sobresalga del tablero para el caso horizontal:
if h_v == "h":
if col == 10:
print("You´re out of range")
elif size == 5 and (col == 7 or col == 8 or col == 9):
print("You´re out of range")
elif size == 4 and (col == 9 or col == 8):
print("You´re out of range")
elif size == 3 and (col == 9):
print("You´re out of range")
elif size == 2 and df.loc[fil,col + 1 ] == "#":
print("You´re crushing with another boat. Try again!")
elif size == 3 and (df.loc[fil,col + 1 ] == "#" or df.loc[fil,col + 2] == "#"):
print("You´re crushing with another boat. Try again!")
elif size == 4 and (df.loc[fil,col + 1 ] == "#" or df.loc[fil,col + 2] == "#" or df.loc[fil,col + 3] == "#"):
print("You´re crushing with another boat. Try again!")
elif size == 5 and (df.loc[fil,col + 1 ] == "#" or df.loc[fil,col + 2] == "#" or df.loc[fil,col + 3] == "#" or df.loc[fil,col + 4] == "#"):
print("You´re crushing with another boat. Try again!")
# Si todo está bien, entonces rellena el tablero con el barco indicado:
else:
fil = fil
col_f = col + size - 1
df.loc[fil,col:col_f] = "#"
print(tablero1)
return df
# Ponemos las condiciones para que el barco no se sobreponga o no se sobresalga del tablero para el caso horizontal:
if h_v == "v":
if fil == 10:
print("You´re out of range")
elif size == 5 and (fil == 7 or fil == 8 or fil == 9):
print("You´re out of range")
elif size == 4 and (fil == 9 or fil == 8):
print("You´re out of range")
elif size == 3 and (fil == 9):
print("You´re out of range")
elif size == 2 and (df.loc[fil + 1, col] == "#"):
print("You´re crushing with another boat. Try again!")
elif size == 3 and (df.loc[fil + 1, col] == "#" or df.loc[fil + 2, col] == "#"):
print("You´re crushing with another boat. Try again!")
elif size == 4 and (df.loc[fil + 1, col] == "#" or df.loc[fil + 2, col] == "#" or df.loc[fil + 3, col] == "#"):
print("You´re crushing with another boat. Try again!")
elif size == 5 and (df.loc[fil + 1, col] == "#" or df.loc[fil + 2, col] == "#" or df.loc[fil + 3, col] == "#" or df.loc[fil + 4 , col] == "#"):
print("You´re crushing with another boat. Try again!")
# Si todo está bien, entonces rellena el tablero con el barco indicado:
else:
fil_f = fil + size -1
col = col
df.loc[fil:fil_f,col] = "#"
print(tablero1)
return df
# Definimos una función para que te vaya pidiendo los barcos que hay que colocar:
# n_barcos = número de barcos que hay que introducir
# tamaño: pequeño, mediano, grande o gigante
# n: la dimensión del barco (2, 3, 4, 5)
# tablero: el tablero sobre el que colocamos los barcos
def coloca_tus_barcos(n_barcos,barco,tamaño,n, tablero):
contador = 1
print("contador1:", contador)
while contador < n_barcos:
print(f'Coloca el barco{contador} de tamaño {tamaño} en el tablero')
if type(barco.colocar_barco(n,tablero1)) != pd.core.frame.DataFrame:
print("Try Again!")
contador -= 1
contador += 1
return tablero
# -
barco_pequeño = Barco(tablero1)
barco_mediano = Barco(tablero1)
barco_grande = Barco(tablero1)
barco_gigante = Barco(tablero1)
# +
""" Definimos una función para que te vaya pidiendo los barcos que hay que colocar:
n_barcos = número de barcos que hay que introducir
tamaño: pequeño, mediano, grande o gigante
n: la dimensión del barco (2, 3, 4, 5)
tablero: el tablero sobre el que colocamos los barcos"""
def coloca_tus_barcos(n_barcos,barco,tamaño,n, tablero):
contador = 1
print("contador1:", contador)
while contador < n_barcos:
print(f'Coloca el barco{contador} de tamaño {tamaño} en el tablero')
if type(barco.colocar_barco(n,tablero1)) != pd.core.frame.DataFrame:
print("Try Again!")
contador -= 1
contador += 1
return tablero
# -
def coloca_jugador1():
Barco.coloca_tus_barcos(2, barco_gigante, "gigante", 5, tablero1)
Barco.coloca_tus_barcos(3, barco_grande, "grande", 4, tablero1)
Barco.coloca_tus_barcos(4, barco_mediano, "mediano", 3, tablero1)
Barco.coloca_tus_barcos(5, barco_pequeño, "pequeño", 2, tablero1)
def coloca_jugador2():
Barco.coloca_tus_barcos(2, barco_gigante, "gigante", 5, tablero2)
Barco.coloca_tus_barcos(3, barco_grande, "grande", 4, tablero2)
Barco.coloca_tus_barcos(4, barco_mediano, "mediano", 3, tablero2)
Barco.coloca_tus_barcos(5, barco_pequeño, "pequeño", 2, tablero2)
| hundir la flota/definitivo_copy_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Script that sets up training repository
#
# Change the settings in the first cell and run all to create the repository and send the invitations.
# +
import os, sys, json,importlib, base64,urllib
#This notebook requires you to install https://github.com/PyGithub/PyGithub first
from github import Github, InputGitAuthor
#Add parent folder to library paths
module_path = os.path.abspath(os.path.join('../../Common-Resources'))
if module_path not in sys.path:
sys.path.append(module_path)
import github_tools as ghTools
# +
accessToken = ghTools.getTokenAttribute(attribute="token")
ghSession = Github(accessToken)
#get user
user = ghSession.get_user()
username = user.login
# -
# ### Manually input name of repository
#The name of the repository to create, use random 4 letters to get unique url
reponame = "lyrics-"
# ## Creating the repo
#making sure the repo is not already tere
try:
del_repo = ghSession.get_repo(username+"/"+reponame)
print("Deleted: " + del_repo.full_name)
del_repo.delete()
except:
print("No repo with that name")
#Create the repo
training_repo = user.create_repo(reponame, auto_init=True)
print(training_repo)
print(training_repo.html_url)
# ### Replace the auto initialized repo with the prepared README and add .gitignore
# #### update README
# +
#Get the commit sha from where the README was created in the auto_init
contents = training_repo.get_contents("README.md")
#Read the README file in this repo to get the content for the commit
file_content = open('repo-content-files/README.md', 'rb').read()
#print(file_content)
#Update the README file
readme_update_result = training_repo.update_file("README.md",
"Update README.md",
file_content,
contents.sha,
branch="main")
readme_update_sha = readme_update_result["commit"].sha
print(readme_update_result)
# -
# #### Create the initial-branch
training_repo.create_git_ref("refs/heads/initial-branch", readme_update_sha)
# #### add .gitignore
# +
#Read the README file in this repo to get the content for the commit
file_content = open("repo-content-files/no.gitignore", 'rb').read()
#Update the README file
create_ignore_result = training_repo.create_file(".gitignore", "add a ignore file", file_content, branch="main")
# -
# ### Start adding lyrics files
# +
list_of_files = [
["lyrics/pop/backstreets-back.txt","Adding Backstreet Back - Go BSB!"],
["lyrics/rock/stairway-to-heaven.txt","Stairway to Heaven! So GOOOD."],
["lyrics/rock/november-rain.txt","November Rain - The solo is AMAZING!"],
["lyrics/pop/mamma-mia.txt","ABBA is the best! And this song is their best!"]
]
for song_list in list_of_files:
#Read the README file in this repo to get the content for the commit
file_content = open("repo-content-files/" + song_list[0], 'rb').read()
#print(file_content)
#Update the README file
create_result = training_repo.create_file(song_list[0], song_list[1], file_content, branch="main")
last_sha = create_result["commit"].sha
print(last_sha)
# -
#Create the queen branch
training_repo.create_git_ref("refs/heads/queen-branch", last_sha)
#Remove Abba from the queen branch
contents = training_repo.get_contents("lyrics/pop/mamma-mia.txt", ref="refs/heads/queen-branch")
training_repo.delete_file(contents.path, "ABBA is not that great", contents.sha, branch="queen-branch")
# +
list_of_files = [
["lyrics/pop/dont-stop-me-now.txt","Dont try to stop me!, Yass Queen","queen-branch"]
]
for song_list in list_of_files:
#Read the README file in this repo to get the content for the commit
file_content = open("repo-content-files/" + song_list[0], 'rb').read()
#print(file_content)
#Update the README file
create_result = training_repo.create_file(song_list[0], song_list[1], file_content, branch=song_list[2])
last_sha = create_result["commit"].sha
print(create_result)
| GitHub-trainings/Intro-Git-GitHub-Contributor/jupyter-repo-setup/Setup TrainingRepo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
n= 5
x = torch.zeros((1,n))
torch.cat([x, x], dim=1).size(1)
import torch.nn as nn
import torch.nn.functional as F
x
x = torch.Tensor
F.relu([1,2])
torch.manual_seed(99)
import numpy as np
w = torch.empty((3,3)).uniform_(-1,1)
x = torch.rand(3,1)
x = x.type_as(w)
y = torch.matmul(w,x)
display(y)
display(torch.relu(y))
w.type()
5//2
# torch.bmm()
display(torch.zeros((3,4)).unsqueeze(-1))
# display(torch.zeros((3,4)).size())
from functools import partial
basetwo = partial(int, baseeeeeeeeeeeeee=2)
basetwo('10010')
# +
class parentClass(object):
def __init__(self,input):
self.input = input
print('__Init__')
def __call__(self, later,power):
out = self.input ** power
self.later = later
print(self.input)
print(out)
print("__call__")
# return out
# parentclass = parentClass(2)
# call = parentClass()
# parentclass()
parentclass(power=3)
# -
ls_x = "hi" is not None
x # do something if ls_x else do other things
if "hi" is not None:
#do something with x
else:
# do other thing with x
basetwo.keywords
| scatch_paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JimKing100/DS-Unit-3-Sprint-1-Software-Engineering/blob/master/module1-python-modules-packages-and-environments/module1-assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="einGn0z3kIVa" colab_type="code" outputId="6e7d74f4-25db-41f1-eed2-e232807b80df" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !pip uninstall ds_helper
# + id="MaLEcN9w4CH2" colab_type="code" outputId="f0f487e6-986a-40e8-ea22-2c10cdc573da" colab={"base_uri": "https://localhost:8080/", "height": 207}
# !pip install --index-url https://test.pypi.org/simple/ ds_helper
# + id="vg-tDhep3kKS" colab_type="code" colab={}
#import ds_helper_data as ds
#import ds_helper_data.df_utils
#ds.df_utils.TEST_DF
# + id="DHyWCnnEHQKF" colab_type="code" outputId="53797262-b521-430c-db72-dcfb2e9fdf91" colab={"base_uri": "https://localhost:8080/", "height": 142}
from ds_helper_data import df_utils
df_utils.TEST_DF
# + id="PMM1kBKbImQS" colab_type="code" colab={}
import pandas as pd
# + id="rOEaN7_kJtKc" colab_type="code" outputId="5c433c46-8ca6-4b7b-aa14-fd938d0c9e2a" colab={"base_uri": "https://localhost:8080/", "height": 204}
df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/DS-Unit-3-Sprint-1-Software-Engineering/master/module1-python-modules-packages-and-environments/drink_test.csv')
df.head()
# + id="4LDkrD6sMNej" colab_type="code" outputId="c8d7af7a-c9dd-48b5-a464-1cc18ca49758" colab={"base_uri": "https://localhost:8080/", "height": 153}
def nulls(df):
columns = list(df)
col_num = 0
print('Nulls by Column')
for i in columns:
print(columns[col_num], df[i].isna().sum())
col_num = col_num + 1
nulls(df)
# + id="k_p1KqLnjmYF" colab_type="code" outputId="a23b0be6-d133-4970-cc21-51d6ef148c38" colab={"base_uri": "https://localhost:8080/", "height": 153}
df_utils.nulls(df)
# + id="PcqNvGd-p1PH" colab_type="code" outputId="80540605-d9ba-40fc-d9ba-6bab31999e50" colab={"base_uri": "https://localhost:8080/", "height": 306}
df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/DS-Unit-3-Sprint-1-Software-Engineering/master/module1-python-modules-packages-and-environments/MarinSalesJanJune2019.csv')
df.head()
# + id="xjwxVDzSqNRV" colab_type="code" colab={}
# from datetime import datetime
# def date_conv(df, col):
# df[col] = pd.to_datetime(df[col], infer_datetime_format=True)
# col_year = col + '_year'
# col_month = col + '_month'
# col_day = col + '_day'
# df[col_year] = df[col].dt.year
# df[col_month] = df[col].dt.month
# df[col_day] = df[col].dt.day
# return df
# df = date_conv(df, 'Listing Date')
# df.head()
# + id="MieE9C-fLJAL" colab_type="code" outputId="ec1664d1-4955-4709-d357-e089da25bf73" colab={"base_uri": "https://localhost:8080/", "height": 649}
df_utils.date_conv(df, 'Listing Date')
df.head()
# + id="rgNLg7EJv96I" colab_type="code" outputId="bfab7f69-f889-4707-bfaa-902f7aac1035" colab={"base_uri": "https://localhost:8080/", "height": 34}
def state_conv(st_abbrev):
us_state_abbrev = {
'Alabama': 'AL',
'Alaska': 'AK',
'Arizona': 'AZ',
'Arkansas': 'AR',
'California': 'CA',
'Colorado': 'CO',
'Connecticut': 'CT',
'Delaware': 'DE',
'District of Columbia': 'DC',
'Florida': 'FL',
'Georgia': 'GA',
'Hawaii': 'HI',
'Idaho': 'ID',
'Illinois': 'IL',
'Indiana': 'IN',
'Iowa': 'IA',
'Kansas': 'KS',
'Kentucky': 'KY',
'Louisiana': 'LA',
'Maine': 'ME',
'Maryland': 'MD',
'Massachusetts': 'MA',
'Michigan': 'MI',
'Minnesota': 'MN',
'Mississippi': 'MS',
'Missouri': 'MO',
'Montana': 'MT',
'Nebraska': 'NE',
'Nevada': 'NV',
'New Hampshire': 'NH',
'New Jersey': 'NJ',
'New Mexico': 'NM',
'New York': 'NY',
'North Carolina': 'NC',
'North Dakota': 'ND',
'Northern Mariana Islands':'MP',
'Ohio': 'OH',
'Oklahoma': 'OK',
'Oregon': 'OR',
'Palau': 'PW',
'Pennsylvania': 'PA',
'Puerto Rico': 'PR',
'Rhode Island': 'RI',
'South Carolina': 'SC',
'South Dakota': 'SD',
'Tennessee': 'TN',
'Texas': 'TX',
'Utah': 'UT',
'Vermont': 'VT',
'Virgin Islands': 'VI',
'Virginia': 'VA',
'Washington': 'WA',
'West Virginia': 'WV',
'Wisconsin': 'WI',
'Wyoming': 'WY',
}
abbrev_us_state = dict(map(reversed, us_state_abbrev.items()))
if len(st_abbrev) == 2:
try:
return_value = abbrev_us_state[st_abbrev]
except KeyError:
return_value = "NA"
else:
try:
return_value = us_state_abbrev[st_abbrev]
except KeyError:
return_value = "NA"
return return_value
print(state_conv('WX'))
# + id="ptQTFqLbVzIs" colab_type="code" outputId="7f90a2b7-5c60-46b0-de64-9a181758afb0" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(df_utils.state_conv('Washington'))
| module1-python-modules-packages-and-environments/module1-assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <table>
# <tr align=left><td><img align=left src="./images/CC-BY.png">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) <NAME></td>
# </table>
# + slideshow={"slide_type": "skip"}
from __future__ import print_function
# %matplotlib inline
import numpy
import matplotlib.pyplot as plt
import matplotlib.animation
from IPython.display import HTML
import os
import pyclaw
path = os.path.join(os.environ.get("CLAW", os.getcwd()), "pyclaw", "fvmbook", "chap6")
os.chdir(path)
import advection
# + [markdown] slideshow={"slide_type": "slide"}
# # Boundary Conditions
#
# Up until now we have yet to address a way to enfore boundary conditions on our methods. In this lecture we will address this and study how we might handle both simple and complex boundary conditions.
# + [markdown] slideshow={"slide_type": "subslide"}
# The primary way we will implement boundary conditions is by adding **ghost cells** to the domain. These cells are outside of the domain and can be set arbitrarily to enforce many types of boundary conditions depending on what is desired. Keep in mind a couple of things though:
#
# 1. For many hyperbolic PDEs we only need BCs on one end of the domain.
# 1. Enforcing non-reflecting BCs is important for many applications is highly non-trivial.
# 1. Setting BCs may require the solution of more complex equations so that an incoming boundary condition is correctly set.
#
# The rest of this lecture will use examples to illustrate different BCs.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: Advection
#
# Consider the scalar advection equation $q_t + u q_x$ on $x\in[a,b]$ with BC
# $$
# q(a, t) = g_a(t)
# $$
# that can easily be generalized to also be at $x=b$. Note also that we only have one BC needed.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Outflow Boundary Conditions
#
# Analytically we have no BC at $x = b$ and if we simply use an upwind method we do not need one numerically. However, if we are using a Lax-Wendroff like method we still require a value at $x=b$ to be specified. One of the easiest ways to specify this is to use **zero-order extrapolation** boundary conditions defined by
# $$
# Q^n_{N+1} = Q^n_N \quad Q^n_{N+2} = Q^n_N.
# $$
# This formulation then implies that the gradient outside of the domain is zero and that in most cases any wave approaching the boundary will simply propagate out of the domain.
# + [markdown] slideshow={"slide_type": "subslide"}
# What about the case where we may want to use a first-order extrapolation given by
# $$
# Q^n_{N+1} = Q^n_N + (Q^n_N - Q^n_{N-1} ) = 2 Q^n_{N} - Q^n_{N-1}?
# $$
# This leads to $\Delta Q^n_{N+1/2} = \Delta Q^n_{N-1/2}$ and the correction terms cancel leading to an update that again reduces to first-order upwind.
# + [markdown] slideshow={"slide_type": "subslide"}
# This extrapolation seems simplistic but turns out to be extremely powerful. This extrapolation, in conjunction with our finite volume methods and characteristic decomposition, tend to be very robust when considering non-reflecting boundary conditions. In fact first-order extrapolation can even lead to issues not found in the zero-order extrapolation and is generally not used.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Inflow Boundary Conditions
#
# At the inflow boundary it is a bit more clear what to do. Depending on how the BC is specified there are multiple ways to numerically represent the BC. The most straight forward perhaps is the case when a direct flux is specified that would easily integrate into the methods discussed already.
# + [markdown] slideshow={"slide_type": "subslide"}
# Based off of this we could want to integrate along the boundary such that
# $$\begin{aligned}
# F^n_{1/2} &= \frac{1}{\Delta t} \int^{t_{n+1}}_{t_n} u q(a, t) dt \\
# &=\frac{u}{\Delta t} \int^{t_{n+1}}_{t_n} g_a(t) dt
# \end{aligned}$$
# or
# $$
# F^n_{1/2} = u g_a(t_n + \Delta t / 2)
# $$
# as an approximation.
# + [markdown] slideshow={"slide_type": "subslide"}
# One draw back to the flux integration approach is that we may need to have fluxes also specified further from the boundary for larger stencil methods.
# + [markdown] slideshow={"slide_type": "subslide"}
# What if instead we tried to find a way to directly specify the values for $Q$ in the ghost cells? This allows the method that we are using interior to the domain to be applied everywhere where we have two cells sharing a boundary. For instance
# $$
# Q^n_0 = \frac{1}{\Delta x} \int^a_{a - \Delta x} q(x, t_n) dx.
# $$
# The drawback here is that we do not know in general $q(x, t_n)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Instead we can use the fact we know the characteristics as they head into the domain. We could for instance use
# $$\begin{aligned}
# q(x, t_n) &= q(a, t_n + (a -x) / u)\\
# &= g_a(t_n + (a - x) / u)
# \end{aligned}$$
# leading to
# $$\begin{aligned}
# Q^n_0 &= \frac{1}{\Delta x} \int^a_{a - \Delta x} g_a\left( t_n + \frac{a - x}{u} \right ) dx \\
# &= \frac{u}{\Delta x} \int^{t_n + \Delta x / u}_{t_n} g_a(\tau) d\tau
# \end{aligned}$$
# We can again approximate these integrals with higher-order approximations.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: Acoustics
#
# Systems of equations are slightly more complex to specify BCs for but follow a similar pattern. There are also special types of BCs that can arise for systems for which we will discuss. We will use the linear acoustics equations for this discussion.
# + [markdown] slideshow={"slide_type": "subslide"}
# Recall that the acoustics equations linearized around a state where $u=0$ takes the form
# $$\begin{aligned}
# &p_t + K_0 u_x = 0 \\
# \rho_0 &u_t + p_x = 0
# \end{aligned}$$
# and has characteristic variables
# $$\begin{aligned}
# w^1(x,t) &= \frac{1}{2 Z_0} (-p + Z_0 u) \\
# w^2(x,t) &= \frac{1}{2 Z_0} (p + Z_0 u)
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Non-Reflecting Boundary Conditions
#
# **Non-reflecting boundary conditions**, also called **absorbing boundary conditions**, are useful when you would like to have waves that leave a finite domain to not impact what is going on interior to the domain. One good example of this is when you want an infinite domain, such as the case with a Cauchy problem, but of course cannot represent this in a numerical example. The non-reflecting boundary conditions will preserve the steady state solution that would evolve between say waves moving in opposite directions. Let us study this in the context of the acoustics equations.
# + [markdown] slideshow={"slide_type": "subslide"}
# With Godunov-type methods that solve a Riemann problem it turns out that zero-order extrapolation works well as non-reflecting boundary conditions. One way to see that this is indeed a good idea is to consider the update formulas in terms of the characteristic variables:
# $$
# W^1 = \frac{-Q^1 + Z_0 Q^2}{2 Z_0} \\
# W^2 = \frac{Q^1 + Z_0 Q^2}{2 Z_0}.
# $$
# This separation into characteristic fields then has the same properties as the scalar advection case. This also leads to the conclusion that
# $$
# Q^n_0 = Q^n_1 \quad \quad Q^n_{-1} = Q^n_1.
# $$
# Note that this causes the gradient to be zero and therefore the flux.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Incoming Waves
#
# Say we want to impose the boundary condition
# $$
# w^2(a, t) = \sin(\omega t)
# $$
# without any reflection of waves hitting this boundary. Decomposing $Q_1$ into the characteristic variables such that
# $$
# Q_1 = W^1_1 r^1 + W^2_1 r^2
# $$
# then the ghost cell should be set to
# $$
# Q_0 = W^1_1 r^1 + \sin(\omega(t_n + \Delta x / 2 c_0)) r^2.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# If we want to generalize this to a system of equations with the boundary condition of $g_a(t)$ then we can also write
# $$
# Q_0 = Q_1 + \left [g_a(t_n + \Delta x / 2 \lambda^j) - W^j_1\right ] r^j.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solid Walls
#
# A particular type of boundary condition comes up often in fluid dynamics, that of a solid wall. If we consider a tube of gas that has a wall at one end we would expect a sound wave to reflect off of the end of the tube with the wall and reverse its direction. The effective solid wall boundary condition should be then
# $$
# u(a, t) = 0.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The key to enforcing a wall boundary condition is to observe that if we set the ghost cell value to an equivalent value for the pressure but an opposite $u$ that this will satisfy the above condition. In other words
# $$
# p^0(a - \xi) = p^0(a + \xi) \\
# u^0(a - \xi) = -u^0(a + \xi)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# This then suggests that we set the ghost cells to
# $$
# Q_0 = \begin{bmatrix} p_0 \\ u_0 \end{bmatrix} = \begin{bmatrix} p_1 \\ -u_1 \end{bmatrix} \\
# Q_{-1} = \begin{bmatrix} p_{-1} \\ u_{-1} \end{bmatrix} = \begin{bmatrix} p_2 \\ -u_2 \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Oscillating Walls
#
# Instead of a solid wall we may want to specify a wall that is oscillating. This could represent a source of acoustic waves, such as a speaker. Here we will only consider small amplitude motions.
# + [markdown] slideshow={"slide_type": "subslide"}
# Consider again the fixed domain $x \in [a, b]$ with the boundary condition
# $$
# u(a, t) = U(t).
# $$
# Note that this will degenerate into our solid wall case if $U(t) = 0$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Consider a single frequency oscillation prescribed by
# $$
# U(t) = \epsilon \sin(\omega t) \quad \epsilon \ll 1.
# $$
# Using what we had from before to prescribe the boundary condition we could then specify
# $$
# Q_0 = \begin{bmatrix} p_0 \\ u_0 \end{bmatrix} = \begin{bmatrix} p_1 \\ 2 U(t_n) - u_1 \end{bmatrix} \\
# Q_{-1} = \begin{bmatrix} p_{-1} \\ u_{-1} \end{bmatrix} = \begin{bmatrix} p_2 \\ 2 U(t_n)-u_2 \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Periodic Boundary Conditions
#
# Last but not least we also have periodic BCs to cover. This is perhaps the easiest of the BCs to specify where with our previous definitions set
# $$
# Q^n_{-1} = Q^n_{N-1} \quad Q^n_0 = Q^n_N \quad Q^n_{N+1} = Q^n_1 \quad Q^n_{N+2} Q^n_2.
# $$
| 04_bc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Summary" data-toc-modified-id="Summary-1"><span class="toc-item-num">1 </span>Summary</a></div><div class="lev1 toc-item"><a href="#Version-Control" data-toc-modified-id="Version-Control-2"><span class="toc-item-num">2 </span>Version Control</a></div><div class="lev1 toc-item"><a href="#Change-Log" data-toc-modified-id="Change-Log-3"><span class="toc-item-num">3 </span>Change Log</a></div><div class="lev1 toc-item"><a href="#Setup" data-toc-modified-id="Setup-4"><span class="toc-item-num">4 </span>Setup</a></div><div class="lev1 toc-item"><a href="#Secure-Credentials-File" data-toc-modified-id="Secure-Credentials-File-5"><span class="toc-item-num">5 </span>Secure Credentials File</a></div><div class="lev1 toc-item"><a href="#Inspect-the-XML-returned" data-toc-modified-id="Inspect-the-XML-returned-6"><span class="toc-item-num">6 </span>Inspect the XML returned</a></div><div class="lev3 toc-item"><a href="#Data-inspection-(root)" data-toc-modified-id="Data-inspection-(root)-601"><span class="toc-item-num">6.0.1 </span>Data inspection (root)</a></div><div class="lev3 toc-item"><a href="#Get-data--(token)" data-toc-modified-id="Get-data--(token)-602"><span class="toc-item-num">6.0.2 </span>Get data (token)</a></div><div class="lev1 toc-item"><a href="#Client" data-toc-modified-id="Client-7"><span class="toc-item-num">7 </span>Client</a></div>
# -
# # Summary
#
# * Master post for the blog series that holds all the links related to making web service calls to Eoddata.com. Overview of the web service can be found [here](http://ws.eoddata.com/data.asmx)
# * Download the [class definition file](https://adriantorrie.github.io/downloads/code/eoddata.py) for an easy to use client, which is demonstrated below
# * This post shows you how to create a secure credentials file to hold the username and password so you don't have to keep entering it, and will allow for automation later.
# * A quick overview is given below of establishing a session using the `requests` module, and parsing the xml response using `xml.etree.cElementTree`. Then a quick inspection of the objects created follows.
#
# The following links were used to help get these things working.
#
# * http://stackoverflow.com/a/17378332/893766
# * http://stackoverflow.com/a/1912483/893766
# * hidden password entry: https://docs.python.org/2/library/getpass.html
# # Version Control
# %run ../../code/version_check.py
# # Change Log
# Date Created: 2017-03-25
#
# Date of Change Change Notes
# -------------- ----------------------------------------------------------------
# 2017-03-25 Initial draft
# 2017-04-02 Added "file saved: <location>" output
# [[Top]](#Table-of-Contents)
# # Setup
# +
# %run ../../code/eoddata.py
from getpass import getpass
import json
import os
import os.path
import requests as r
import stat
import xml.etree.cElementTree as etree
ws = 'http://ws.eoddata.com/data.asmx'
ns='http://ws.eoddata.com/Data'
session = r.Session()
# -
username = getpass()
password = <PASSWORD>()
# [[Top]](#Table-of-Contents)
# # Secure Credentials File
# Create credentials file for later usage. The file will have permissions created so only the current user can access the file. The following [SO post](http://stackoverflow.com/a/15015748/893766) was followed.
#
# The following directory will be created if it doesn't exist:
# * Windows: `%USERPROFILE%/.eoddata`
# * Linux: `~/.eoddata`
# +
# gather credentials
credentials = {'username': username, 'password': password}
# set filename variables
credentials_dir = os.path.join(os.path.expanduser("~"), '.eoddata')
credentials_file_name = 'credentials'
credentials_path = os.path.join(credentials_dir, credentials_file_name)
# set security variables
flags = os.O_WRONLY | os.O_CREAT | os.O_EXCL # Refer to "man 2 open".
mode = stat.S_IRUSR | stat.S_IWUSR # This is 0o600 in octal and 384 in decimal.
# create directory for file if not exists
if not os.path.exists(credentials_dir):
os.makedirs(credentials_dir)
# for security, remove file with potentially elevated mode
try:
os.remove(credentials_path)
except OSError:
pass
# open file descriptor
umask_original = os.umask(0)
try:
fdesc = os.open(credentials_path, flags, mode)
finally:
os.umask(umask_original)
# save credentials in secure file
with os.fdopen(fdesc, 'w') as f:
json.dump(credentials, f)
f.write("\n")
print("file saved: {}".format(credentials_path))
# -
# [[Top]](#Table-of-Contents)
# # Inspect the XML returned
# +
call = 'Login'
url = '/'.join((ws, call))
payload = {'Username': username, 'Password': password}
response = session.get(url, params=payload, stream=True)
if response.status_code == 200:
root = etree.parse(response.raw).getroot()
# -
# ### Data inspection (root)
dir(root)
for child in root.getchildren():
print (child.tag, child.attribute)
for item in root.items():
print (item)
for key in root.keys():
print (key)
print (root.get('Message'))
print (root.get('Token'))
print (root.get('DataFormat'))
print (root.get('Header'))
print (root.get('Suffix'))
# ### Get data (token)
token = root.get('Token')
# [[Top]](#Table-of-Contents)
# # Client
# client can be opened using a with statement
with (Client()) as eoddata:
print('token: {}'.format(eoddata.get_token()))
# +
# initialise using secure credentials file
eoddata = Client()
# client field accessors
ws = eoddata.get_web_service()
ns = eoddata.get_namespace()
token = eoddata.get_token()
session = eoddata.get_session()
print('ws: {}'.format(ws))
print('ns: {}'.format(ns))
print('token: {}'.format(token))
print(session)
# -
# the client has a list of exchange codes avaiable once intialised
eoddata.get_exchange_codes()
# client must be closed if opened outside a with block
session.close()
eoddata.close_session()
# [[Top]](#Table-of-Contents)
| content/downloads/notebooks/eoddata/eoddata_web_service_series_master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing
#
# * Null Values
# * Encoding Categorical Columns.
# * Standardisation and Normalisation.
# * Feature Generation.
# * Feature Selection – (Multicolinearity, Dimensionality Reduction).
# * Handling Noisy Data – (Binning, CLustering)
# * Handling Class Imbalance - Covered in CLassification.
# ## A. Null Values
#
# Types of null values:
# * Unit non-response - Entire entry missing. NaNs not necessarily visible in dataset.
# * Item non-response - When specific cells of a column are missing. (Typical missing value definition).
# * **MCAR** - Missing Completely At Randon - Missing values miss independently. No systematic difference between missing and available data.
# * **MAR** - Missing At Random. When missing values are dependent on some variable but independent on itself. e.g When a sensor breaks then we have missing temperature values with no systematic difference with existing data.
# * **MNAR** - Missing Not At Random. When a missing value is dependent on the value itself.Thus the difference is sytematic. e.g when a low income group fails to answer a questionnaire because its low.
#
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import stats
import scipy
# +
# # !pip install stats
# +
# # !pip install scipy
# -
data = pd.read_csv('/home/ada/teacherOn/<NAME>/Intermediate ML/datasets/train.csv', sep = ',')
print('the dataset has', data.shape[0], 'rows and', data.shape[1], 'columns.')
data.sample(5)
# +
# (data.isna().sum()/data.shape[0])*100
# +
# check for null values and do a plot
nulls = (data.isna().sum()/data.shape[0])*100
nulls = pd.DataFrame(nulls)
plt.figure(figsize = (16,6))
plt.xticks(rotation = 60)
n = nulls.head(20)
sns.barplot(x = n.index, y = n.iloc[:,0])
# nulls
# -
# ### Mean, Median, and Mode Imputation.
#
# * Pros - Easy
# * Cons - Distorts Variance.
# * Handles: MCAR and MAR Item Non-Response, sensitive to outliers.
#
# **PS**: Shouldn't be used in MNAR cases.
# +
# data.dtypes
# data['galaxy'].fillna(np.mode(data['galaxy']))
# +
# imputation
# strategy = mean, median, most_frequent(mode), constant. default = 'mean'
# numerical and categorical imputers
from sklearn.impute import SimpleImputer
#NaN
numerical_imputer = SimpleImputer(missing_values=np.nan, strategy = 'mean')
categorical_imputer = SimpleImputer(missing_values=np.nan, strategy = 'most_frequent')
# numerical and categorical columns
categorical = []
numerical = []
for col in data.columns:
if data[col].dtype in ['int8', 'int16', 'int32', 'int64', 'float16', 'float32', 'float64']:
numerical.append(col)
elif data[col].dtype == object:
categorical.append(col)
columns = numerical + categorical
new_data = data[columns]
# new_data.info()
# get index locations of the columns
numerical_indices = []
cat_indices = []
for col_name in numerical:
i = new_data.columns.get_loc(col_name)
numerical_indices.append(i)
for col_name in categorical:
i = new_data.columns.get_loc(col_name)
cat_indices.append(i)
# actual imputation
# numerical
for col in numerical_indices:
x = new_data.iloc[:, col].values
x = x.reshape(-1,1)
x = numerical_imputer.fit_transform(x)
# imputer = numerical_imputer.fit(x)
# x = imputer.transform(x)
new_data.iloc[:, col] = x
# categorical
for col in cat_indices:
x = new_data.iloc[:, col].values
x = x.reshape(-1,1)
imputer = categorical_imputer.fit(x)
x = imputer.transform(x)
new_data.iloc[:, col] = x
# checking again for null values
nulls = (new_data.isna().sum()/data.shape[0])*100
nulls = pd.DataFrame(nulls)
plt.figure(figsize = (12,3))
plt.xticks(rotation = 60)
n = nulls.head(20)
sns.barplot(x = n.index, y = n.iloc[:,0])
# -
# fillna function
data['column name'].fillna(0, inplace = True)
# Notes on Null values imputation:
# * If you replace missings with means, naturally the mean is preserved.
# * median best for data with robust outliers because mean would distort variance alot.
# * median reflects the central tendency the best implying that for continuous data, the use of the median is better than mean.
# ### Model Based Imputation
#
# Column with missing value is set as the target variable then the rest of the features act as predictor variables.
#
# * Pros: Improvement over Mean/Median/Mode Imputation.
#
# * Cons: Still distorts histograms — Underestimates variance.
#
# * Handles: MCAR and MAR Item Non-Response
# ### KNN Imputation
# KNNImputer helps to impute missing values by finding the k nearest neighbors with the Euclidean distance matrix. (Shortest distance possible - straight line between data points).
#
# Each sample’s missing values are imputed using the mean value from n_neighbors nearest neighbors found in the training set. Two samples are close if the features that neither is missing are close.
#
# * Pros - more accurate than simple imputation.
# * cons - computationally expensive, sensitive to outliers.
# +
# import numpy as np
# from sklearn.impute import KNNImputer
# imputer = KNNImputer(n_neighbors=2) #define the k nearest neighbors
# imputer.fit_transform(data)
# +
import numpy as np
from sklearn.impute import KNNImputer
X = [[1, 2, np.nan], [3, 4, 3], [np.nan, 6, 5], [8, 8, 7]]
print(np.array(X))
imputer = KNNImputer(n_neighbors=2)
y = imputer.fit_transform(X)
y
# -
# ## B. Encoding Categorical Columns
# Most ML models cannot handle categorical data, thus they have to be converted to numerical form.
# Exception of **catboost** which actually gives room for categorical columns.
#
# **Nominal**: Categories do not imply any quantitative measure and there is typically no order in the data. For example, race, gender, languages are categorical variables but we cannot order different categories of these variables.
#
# **Ordinal**: Unlike nominal data, there is an order between categories. One category can be superior to another category and vice versa. e.g .*Low, medium, high, Cold, warm, hot*
# ### 1. One Hot Encoding
# Each category is mapped into a vector containing zeros and ones.
#
# * pros - doesn't assign hierarchy to categories.
# * cons - computationally expensive.
data = pd.read_csv("/home/ada/teacherOn/<NAME>/Intermediate ML/datasets/house.csv")
data.sample(5)
# +
# get dummies function
encoded = pd.get_dummies(data, drop_first = True) # defaults to OHE
print('shape before encoding:', data.shape)
print('shape after get dummies function:', encoded.shape)
# # OneHotEncoder function
# from sklearn.preprocessing import OneHotEncoder
# ohe = OneHotEncoder()
# encoded_features = ohe.fit_transform(data['Street']).reshape(-1,1)
# print('shape after the OHE function:', encoded_features.shape)
# -
encoded.head()
# ### 2. Label Encoding
# Each category is assigned a value from 1 through N where N is the number of categories for the feature/variable.
#
# * cons - assigns hierarchy to categories.
# * pros - computationally cheap.
data['SaleCondition'].unique()
# +
# label encoder function
from sklearn.preprocessing import LabelEncoder
print(data['SaleCondition'].head())
le = LabelEncoder()
data['sales_transformed'] = le.fit_transform(data['SaleCondition'])
print(data['sales_transformed'].head())
# -
data['sales_transformed'].unique()
# ### 3. Simple Replacing
# Replacing categories with desired numerical values.
# +
# customised function
print('entries before encoding:', data['SaleCondition'].unique())
data['SaleCondition'] = data['SaleCondition'].replace({'Normal':1, 'Abnorml':2, 'Partial':3,
'AdjLand':4, 'Alloca':5, 'Family':5})
print('entries after encoding:', data['SaleCondition'].unique())
# -
# ## C. Standardisation and Normalisation
# > **Normalization** usually means to scale a variable to have a values between 0 and 1, while **standardization** transforms data to have a mean of zero and a standard deviation of 1.
#
# > The result of **standardization** (or Z-score normalization) is that the features will be rescaled to ensure the mean and the standard deviation to be 0 and 1, respectively.
# <img src="https://miro.medium.com/max/1400/1*xIilonr1kVdoQRRvcpc1ng.png">
#
# > This technique is to re-scale features value with the distribution value between 0 and 1 is useful for the optimization algorithms, such as gradient descent, that are used within machine learning algorithms that weight inputs (e.g., regression and neural networks).
# > Rescaling is also used for algorithms that use distance measurements, for example, K-Nearest-Neighbours (KNN).
#
# > Variables that are measured at different scales do not contribute equally to the analysis and might end up creating a bais. (End goal is to get a uniform scale for all the numerical variables).
# ### 0. Log Transformation
f, ax = plt.subplots(figsize=(6, 5))
sns.distplot(data['SalePrice'], bins = 20, color = 'Magenta')
ax.set(ylabel="Frequency")
ax.set(xlabel="SalePrice")
ax.set(title="SalePrice distribution before transformation")
# +
# log transformation
'''returns an array with natural logarithmic value of x + 1.
The plus 1 formulae is used because they're more accurate for smaller values '''
data1 = data.copy()
data1['SalePrice'] = np.log1p(data1['SalePrice'])
f, ax = plt.subplots(figsize=(7, 5))
sns.distplot(data1['SalePrice'], bins = 20, color = 'Magenta')
ax.set(ylabel="Frequency")
ax.set(xlabel="Age")
ax.set(title="SalePrice distribution after log transformation")
# +
# # boxcox transformation
# from scipy.stats import skew, norm
# from scipy.special import boxcox1p
# from scipy.stats import boxcox_normmax
# data2 = data.copy()
# data2['SalePrice'] = boxcox1p(data2['SalePrice'], boxcox_normmax(data2['SalePrice'] + 1))
# f, ax = plt.subplots(figsize=(7, 5))
# sns.distplot(data1['SalePrice'], bins = 20, color = 'Magenta')
# ax.set(ylabel="Frequency")
# ax.set(xlabel="Age")
# ax.set(title="Age distribution after box cox transformation")
# -
# ### 1. Standard Scaler
# StandardScaler removes the mean (0) and scales the data to unit variance(1). However, the outliers have an influence when computing the empirical mean and standard deviation which shrink the range of the feature.
#
# ** Mean is used to compute the standardised output.
#
# StandardScaler therefore cannot guarantee balanced feature scales in the presence of outliers.
# +
# define data
data = np.array([[100, 0.001],
[8, 0.05],
[50, 0.005],
[88, 0.07],
[4, 0.1]])
print(data)
# define standard scaler
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# transform data
scaled = scaler.fit_transform(data)
print(scaled)
# -
# ### 2. MinMax Scaler
# MinMaxScaler rescales the data set such that all feature values are in the range [0, 1].
#
# A MinMaxScaler is very sensitive to the presence of outliers.
# define min max scaler
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
# transform data
scaled = scaler.fit_transform(data)
print(scaled)
# ### 3. Robust Scaler
# This Scaler removes the median and scales the data according to the quantile range (defaults to IQR: Interquartile Range). The IQR is the range between the 1st quartile (25th quantile) and the 3rd quartile (75th quantile).
# +
# define robust scaler
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
# transform data
scaled = scaler.fit_transform(data)
print(scaled)
# -
# ## D. Feature Generation
# Generating new features based on the ones available.
data = pd.read_csv("/home/ada/teacherOn/<NAME>/Intermediate ML/datasets/trains.csv")
data.sample(5)
# ### Extraction from datetime columns
# +
# feature generation from date columns
print('dtype before conversion', data['Date'].dtype)
#changing the date column dtype to datetime
data['Date'] = pd.to_datetime(data['Date'])
print('dtype after conversion', data['Date'].dtype)
# data['Date'].head()
# +
# feature extraction/generation
data['Month'] = data['Date'].dt.month.to_list()
data['Year'] = data['Date'].dt.year.to_list()
data['Day'] = data['Date'].dt.day.to_list()
data['WeekOfYear'] = data['Date'].dt.weekofyear.to_list()
data['DayOfWeek'] = data['Date'].dt.dayofweek.to_list()
# weekday or not weekday
data['weekday'] = 1 # Initialize the column with default value of 1
data.loc[data['DayOfWeek'] == 5, 'weekday'] = 0
data.loc[data['DayOfWeek'] == 6, 'weekday'] = 0
# +
# data.sample(5)
# -
data.columns
# ### Extraction from arithmetic operations
data['customer_sales_ration'] = data['Customers'] / data['Sales']
data.head()
# ### Conditional Feature Generation
data['double_holiday'] = 0
data.loc[(data['StateHoliday'] == 1) & (data['SchoolHoliday'] == 1), 'double_holiday'] = 1
data.shape
# ## E. Feature Selection
data.head()
# # droping useless columns
data = data.drop(['Date', 'Store'], axis = 1)
# data = data.drop(['Store'], axis = 1)
data.shape
# check for multicolinearity
corr = data.corr()
corr
plt.figure(figsize = (12,7))
sns.heatmap(corr, cmap = 'BuPu')
data = data.drop(['WeekOfYear'], axis = 1)
data.shape
# ### Dimensionality Reduction (To e Covered Later in detail.)
# ## F. Handling Noisy Data (Binning)
# Grouping technique for numerical data. Sometimes ranges maybe more important than actual values. e.g age groups, salary groups, ...
data = pd.read_csv("/home/ada/teacherOn/<NAME>/Intermediate ML/datasets/store.csv")
data.sample(2)
# +
# pandas cut methods
'''Bins based on actual values. Values are ordered then groupeed.'''
data['distance_rank'] = pd.cut(data['CompetitionDistance'], 5, labels = False) # labels argument takes a list.
data['distance_rank'].unique()
# +
# pandas qcut method
'bins indicate quartile membership'
data['dist_rank'] = pd.qcut(data['CompetitionDistance'], 5, labels = False)
data['dist_rank'].value_counts()
# -
# ## G. Class Imbalance
#
# ** To be covered under classification.
| .ipynb_checkpoints/2 - Preprocessing & Feature Engineering-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```sh
# pip3 install sortedcontainers
# ```
from sortedcontainers import SortedSet, SortedList
from bisect import bisect
sl = SortedList(i * 10 for i in range(1000000))
ss = SortedSet(i * 10 for i in range(1000000))
s = set(i * 10 for i in range(1000000))
list(ss.irange(123, 456))
def isin():
n = 0
for i in range(100000):
if i * 2 in s:
n += 1
return n
isin()
def issin():
cnts = ss.__contains__
n = 0
for i in range(100000):
if cnts(i * 2):
n += 1
return n
issin()
def issin2():
s = ss._set
n = 0
for i in range(100000):
if i * 2 in s:
n += 1
return n
issin2()
def issiin():
cnts = ss.__contains__
return sum(1 for i in range(100000) if cnts(i * 2))
issiin()
def islin():
n = 0
for i in range(100000):
if i * 2 in sl:
n += 1
return n
islin()
from utils import prof
def perf(func, *args, **kwargs):
return prof(func, locals(), globals(), *args, **kwargs)
result = perf(isin)
result = perf(issin)
result = perf(issin2)
result = perf(issiin)
result = perf(islin)
# +
import pickle
import gzip
GZIP_LEVEL = 2
PICKLE_PROTOCOL = 4
# +
def save(data, fileName):
with gzip.open(fileName, "wb", compresslevel=GZIP_LEVEL) as f:
pickle.dump(data, f, protocol=PICKLE_PROTOCOL)
def load(fileName):
with gzip.open(fileName, "rb") as f:
return pickle.load(f)
# -
fileName = '_temp/ss.tfx'
perf(save, ss, '_temp/ss.tfx')
ss2 = perf(load, fileName)
import collections
data = {i * 3: i * 10 for i in range(1_000_000)}
# +
def none():
return None
data2 = collections.defaultdict(none, data)
# -
fileName = '_temp/defaultdict.tfx'
save(data2, fileName)
data3 = load(fileName)
data3 == data2
def v(data, n):
if n in data:
return data[n]
return None
def v2(data, n):
return data[n]
def getVal():
myData = data
n = 0
for i in range(100_000):
if v(data, i) is None:
n += 1
return n
getVal()
def getVal2():
myData = data2
n = 0
for i in range(100_000):
if v2(data2, i) is None:
n += 1
return n
getVal2()
def getVal3():
myData = data2
n = 0
for i in range(100_000):
if data2[i] is None:
n += 1
return n
getVal3()
perf(getVal)
perf(getVal2)
perf(getVal333)
from tf.app import use
A = use('bhsa:clone', checkout='clone', hoist=globals())
# +
otypeLines = '''
1-426584 word
426585-426623 book
426624-427552 chapter
427553-515673 clause
515674-606361 clause_atom
606362-651541 half_verse
651542-904748 phrase
904749-1172289 phrase_atom
1172290-1236016 sentence
1236017-1300541 sentence_atom
1300542-1414353 subphrase
1414354-1437566 verse
1437567-1446799 lex
'''.strip().split('\n')
otypeItems = [x.split('\t') for x in otypeLines]
otypeData = [tuple(int(y) for y in x[0].split('-')) + (x[1],) for x in otypeItems]
otypeData
# -
otypeBounds = tuple(x[1] for x in otypeData)
otypeBoundaries = SortedList(otypeBounds)
otypeBoundaries
otypeTypes = [x[2] for x in otypeData]
otypeTypes
def otype(n):
i = otypeBoundaries.bisect_left(n)
return otypeTypes[i]
otype(1)
otype(426584)
otype(426585)
from random import randrange
from itertools import repeat
maxNode = 1446799
F.otype.maxNode
testIndices = tuple(randrange(1, maxNode) for _ in repeat(None, 100_000))
testIndices[0:10]
def getOtype():
fOtype = F.otype.v
WORD = 'word'
n = 0
for i in testIndices:
if fOtype(i) == WORD:
n += 1
return n
getOtype()
def getOtype2():
WORD = 'word'
n = 0
for i in testIndices:
j = otypeBoundaries.bisect_left(i)
thisOtype = otypeTypes[j]
if thisOtype == WORD:
n += 1
return n
getOtype2()
def getOtype3():
fOtype = otype
WORD = 'word'
n = 0
for i in testIndices:
if fOtype(i) == WORD:
n += 1
return n
getOtype3()
def getOtype4():
WORD = 'word'
bs = otypeBoundaries.bisect_left
otypes = otypeTypes
n = 0
for i in testIndices:
j = bs(i)
thisOtype = otypes[j]
if thisOtype == WORD:
n += 1
return n
getOtype4()
def getOtype5():
WORD = 'word'
bs = bisect
obounds = otypeBounds
otypes = otypeTypes
n = 0
for i in testIndices:
j = bs(otypeBounds, i)
thisOtype = otypes[j]
if thisOtype == WORD:
n += 1
return n
getOtype5()
perf(getOtype)
perf(getOtype5)
perf(getOtype2)
perf(getOtype3)
perf(getOtype4)
| test/query/containers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sqlite3
import matplotlib.pyplot as plot
# +
def subset(org_and_repo):
org, repo = org_and_repo.split('/')
return data.loc[(data.organization == org) & (data.repository == repo)]
def merge_repos(repos, metric='stargazers'):
data = pd.DataFrame(columns=['Date'])
for repo in repos:
repo_data = subset(repo)
repo_data = repo_data[['date', metric]].rename(columns={metric: repo, 'date': 'Date'})
data = data.merge(repo_data, on='Date', how='outer')
return data.sort_values(by='Date')
def plot_repos(title, repos, metric='stargazers'):
plot_data = merge_repos(repos, metric)
plot_data.plot(x='Date', y=repos, title=title, figsize=(15,10))
def plot_all_repos(title, org, metric='stargazers'):
repo_data = data.loc[data.organization == org]
repos = [org + '/' + repo for repo in repo_data.repository.unique()]
plot_repos(title, repos, metric)
# -
conn = sqlite3.connect('data-vol-1/github.db')
data = pd.read_sql_query("""
SELECT strftime('%Y-%m-01', date) AS date, organization, repository,
MAX(stargazers) AS stargazers, MAX(forks) AS forks
FROM github_stats
GROUP BY organization, repository, strftime('%Y-%m', date)
""", conn)
# +
repos = [
'operator-framework/operator-sdk',
'kubernetes-sigs/kubebuilder',
'kubernetes-sigs/controller-runtime',
'kudobuilder/kudo',
'bluek8s/kubedirector',
'kubedb/operator',
'rook/operator-kit',
'GoogleCloudPlatform/metacontroller',
'zalando-incubator/kopf'
]
plot_repos('Operator Tools', repos)
# +
import requests
import re
r = requests.get('https://twitter.com/kudobuilder')
m = re.search('title="([0-9]+) Followers"', r.text)
followers = int(m.group(1))
cursor = conn.cursor()
cursor.execute('''INSERT INTO twitter_stats (date, followers) VALUES (date('now'), ?)''', (followers,))
conn.commit()
print("@kudobuilder has {} Twitter followers".format(followers))
# -
plot_all_repos('Operator Framework', 'operator-framework')
# +
awesome_repo_names = [l.rstrip('\n') for l in open('data-vol-1/awesome_operators.txt').readlines()]
def name_and_stars(name):
awesome_data = subset(name)
max_stars = awesome_data['stargazers'].max()
return (name, max_stars)
awesome_data = pd.DataFrame(map(name_and_stars, awesome_repo_names), columns=['repository', 'stargazers']).dropna()
n = 50
top_n_operators = awesome_data.sort_values(by='stargazers', ascending=False).head(n)
top_n_operators
# -
top_n_operators.plot(x='repository', y='stargazers', kind='bar', figsize=(20,10), title='Github stars for operators listed in awesome-operators')
# +
repos = [
'operator-framework/operator-sdk',
'kubernetes-sigs/kubebuilder',
'kudobuilder/kudo',
'bluek8s/kubedirector',
'kubedb/operator',
'rook/operator-kit',
'GoogleCloudPlatform/metacontroller',
'zalando-incubator/kopf'
]
plot_data = merge_repos(repos, 'forks')
plot_data
| notebooks/k8s-analytics/KUDO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exponential Moving Average
# t_i = alpha * t_{i-1} + (1 - alpha) * s_i, with a value of alpha = 0.99
import os
os.chdir(os.path.join(os.getcwd(), '..'))
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from src.model import mean_teacher
from keras.applications.resnet50 import ResNet50
import keras.backend as K
config = K.tf.ConfigProto()
config.gpu_options.allow_growth = True
session = K.tf.InteractiveSession(config=config)
K.set_learning_phase(1)
configs = {
'input_shape': (224,224,3),
'num_of_classes': 12,
'lr':1e-3,
'ratio':0.5
}
mean_teacher_model, student_model, teacher_model = mean_teacher(configs)
mean_teacher_weights = mean_teacher_model.layers[-1].get_weights()
teacher_weights = teacher_model.get_weights()
student_weights = student_model.get_weights()
for i in range(len(mean_teacher_weights)):
if not np.array_equal(mean_teacher_weights[i], student_weights[i]):
print(np.array_equal(mean_teacher_weights[i], student_weights[i]))
len(mean_teacher_weights), len(student_weights), len(teacher_weights)
temp = student_model.get_weights()
ema(student_model, student_model)
def ema(student_model, teacher_model, alpha = 0.99):
'''
Calculates the exponential moving average of the student model weights and updates the teacher model weights\
formula:
t_i = alpha * t_{i-1} + (1 - alpha) * s_i, with default alpha = 0.99
t_i = weights of teacher model in current epoch
s_i = weights of student model in current epoch
'''
student_weights = student_model.get_weights()
teacher_weights = teacher_model.get_weights()
assert len(student_weights) == len(teacher_weights), 'length of student and teachers weights are not equal Please check. \n Student: {}, \n Teacher:{}'.format(len(student_weights), len(teacher_weights))
new_layers = []
for i, layers in enumerate(student_weights):
new_layer = alpha*(teacher_weights[i]) + (1-alpha)*layers
new_layers.append(new_layer)
teacher_model.set_weights(new_layers)
# # EMA lambda callback
from keras.callbacks import LambdaCallback
ema_callback = LambdaCallback(on_epoch_end=lambda epoch, logs : ema(student_model, teacher_model, alpha = 0.99))
# ## Loss function
# ### model architecture
#
# 
#
# useful links:
# - https://keras.io/layers/about-keras-layers/
# - https://keras.io/getting-started/functional-api-guide/
# - https://github.com/keras-team/keras/blob/master/keras/losses.py
# - https://keras.io/getting-started/functional-api-guide/
# - https://towardsdatascience.com/advanced-keras-constructing-complex-custom-losses-and-metrics-c07ca130a618
# - https://stackoverflow.com/questions/38972380/keras-how-to-use-fit-generator-with-multiple-outputs-of-different-type
from keras.optimizers import Adam
from keras.models import Model
from keras.preprocessing.image import ImageDataGenerator
def categorical_crossentropy(y_true, y_pred):
return K.categorical_crossentropy(y_true, y_pred)
def weighted_sum_loss(squared_difference_layer, ratio = 0.5):
def categorical_crossentropy_custom(y_true, y_pred):
return ratio * K.categorical_crossentropy(y_true, y_pred) + (1 - ratio)*squared_difference_layer
return categorical_crossentropy_custom
# # Data Generator
#
# useful links:
# - https://medium.com/@ensembledme/writing-custom-keras-generators-fe815d992c5a
def get_distribution(data_path, label, color):
walker = os.walk(data_path)
next(walker) # skip the first row
class_freq = dict()
for r,d,f in walker:
class_freq[r.split('/')[-1]] = len(f)
class_freq_df = pd.DataFrame.from_dict(class_freq, orient = 'index', columns = ['count'])
class_freq_df.reset_index(inplace = True)
class_freq_df.columns = [label, 'count']
class_freq_df.sort_values('count', axis = 0, ascending=False, inplace=True)
sns.catplot(x = 'count', y = label, kind = 'bar', data=class_freq_df, color = color)
color_1 = sns.xkcd_rgb['denim blue']
color_2 = sns.xkcd_rgb['dusty purple']
syn_path = os.path.join(os.getcwd(), 'reduced_data', 'synthetic')
real_path = os.path.join(os.getcwd(), 'reduced_data', 'real')
get_distribution(syn_path, 'synthetic', color_1)
get_distribution(real_path, 'real', color_2)
# +
sup_gen = ImageDataGenerator()
unsup_gen = ImageDataGenerator()
sup_data_gen = sup_gen.flow_from_directory(syn_path)
# -
configs = {'batch_size': 32,
'target_size': (224,224)}
def mean_teacher_data_gen(sup_gen, unsup_gen, batch_size, target_size):
'''
'''
syn_path = os.path.join(os.getcwd(), 'reduced_data', 'synthetic')
real_path = os.path.join(os.getcwd(), 'reduced_data', 'real')
sup_data_gen = sup_gen.flow_from_directory(real_path,
target_size=target_size,
class_mode='categorical')
unsup_data_gen = unsup_gen.flow_from_directory(syn_path,
target_size=target_size,
class_mode='categorical')
while True:
syn_img, syn_labels = sup_data_gen.next()
real_img, _ = unsup_data_gen.next()
yield [syn_img, real_img] , syn_labels
mean_teacher_generator = mean_teacher_data_gen(sup_gen, unsup_gen, configs['batch_size'], configs['target_size'])
inputs, outputs = next(mean_teacher_generator)
syn_img, real_img = inputs
class_dict = dict[(v,k) for (k,v) in sup_data_gen.class_indices]
fig,ax = plt.subplots(1,2)
ax[0].set_axis_off()
ax[1].set_axis_off()
ax[0].imshow(syn_img[0].astype(np.uint8))
ax[1].imshow(real_img[0].astype(np.uint8))
plt.title(class)
plt.show()
for i, syn_i in enumerate(syn_img):
ax, fig = plt.subplot([2,1])
# # Training
total_samples
total_samples = sup_data_gen.n
batch_size = sup_data_gen.batch_size
total_samples//batch_size + 1
mean_teacher_model.fit_generator(mean_teacher_generator,
steps_per_epoch=total_samples // batch_size + 1,
epochs=1,
callbacks=[ema_callback])
| notebook/ema and loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from tpot import TPOTRegressor, TPOTClassifier
from sklearn.model_selection import train_test_split
import numpy as np
np.random.seed(0)
import warnings
warnings.filterwarnings("ignore")
import time as tm
import pandas as pd
from sklearn.metrics import f1_score, recall_score, accuracy_score, confusion_matrix
from sklearn.model_selection import LeaveOneGroupOut
from sklearn import preprocessing
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.colors as colors
from mpl_toolkits.axes_grid1 import make_axes_locatable
from scipy.signal import medfilt
# %matplotlib inline
# -
# ## Load dataset
pe_fv = pd.read_csv('../facies_vectors.csv')
pe_nf = pd.read_csv('../nofacies_data.csv')
pe_fv.columns
# +
Xfv = pe_fv[pe_fv["PE"].notnull()].drop(['Formation', 'Well Name', 'Depth', 'Facies', 'PE'], axis=1).values
Xnf = pe_nf[pe_nf["PE"].notnull()].drop(['Formation', 'Well Name', 'Depth', 'PE'], axis=1).values
Xpe = np.concatenate((Xfv, Xnf))
Yfv = pe_fv[pe_fv["PE"].notnull()]["PE"].values
Ynf = pe_nf[pe_nf["PE"].notnull()]["PE"].values
Ype = np.concatenate((Yfv, Ynf))
# -
Xpetr, Xpete, Ypetr, Ypete = train_test_split(Xpe, Ype, train_size=0.7, test_size=0.3, random_state=0)
# +
# # peReg = TPOTRegressor(generations=10, population_size=5, max_eval_time_mins=0.5, max_time_mins=1, verbosity=3)
# peReg = TPOTRegressor(generations=50, population_size=10, max_time_mins=60, verbosity=3)
# peReg.fit(Xpetr, Ypetr)
# print(peReg.score(Xpete, Ypete))
# peReg.export('pe_imputer_pipeline0.py')
# +
from sklearn.ensemble import ExtraTreesRegressor, VotingClassifier
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline, make_union
from sklearn.preprocessing import FunctionTransformer
pe_imputer = make_pipeline(
ExtraTreesRegressor(max_features=0.74, n_estimators=500)
)
from sklearn.decomposition import FastICA
from sklearn.ensemble import ExtraTreesRegressor, VotingClassifier
from sklearn.linear_model import ElasticNet
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline, make_union
from sklearn.preprocessing import FunctionTransformer
pe_imputer = make_pipeline(
FastICA(tol=2.0),
make_union(VotingClassifier([("est", ElasticNet(alpha=0.02, l1_ratio=0.96))]), FunctionTransformer(lambda X: X)),
ExtraTreesRegressor(max_features=0.44, n_estimators=500)
)
pe_imputer.fit(Xpe, Ype)
# results = exported_pipeline.predict(testing_features)
# -
training_data = pd.read_csv("../facies_vectors.csv")
XimpPE = training_data[training_data["PE"].isnull()].drop(['Formation', 'Well Name', 'Depth', 'Facies', 'PE'], axis=1).values
training_data["PE"][training_data["PE"].isnull()] = pe_imputer.predict(XimpPE)
training_data["PE"][training_data["PE"].isnull()].head()
# ### Utilities function
# +
def accuracy(conf):
total_correct = 0.
nb_classes = conf.shape[0]
for i in np.arange(0,nb_classes):
total_correct += conf[i][i]
acc = total_correct/sum(sum(conf))
return acc
adjacent_facies = np.array([[1], [0, 2], [1], [4], [3, 5], [4, 6, 7], [5, 7], [5, 6, 8], [6, 7]])
def accuracy_adjacent(conf, adjacent_facies):
nb_classes = conf.shape[0]
total_correct = 0.
for i in np.arange(0,nb_classes):
total_correct += conf[i][i]
for j in adjacent_facies[i]:
total_correct += conf[i][j]
return total_correct / sum(sum(conf))
def mad_based_outlier(points, thresh=4.5):
median = np.median(points, axis=0)
diff = (points - median)**2
diff = np.sqrt(diff)
med_abs_deviation = np.median(diff)
modified_z_score = 0.6745 * diff / med_abs_deviation
return abs(modified_z_score),abs(modified_z_score) > thresh
# +
# 1=sandstone 2=c_siltstone 3=f_siltstone
# 4=marine_silt_shale 5=mudstone 6=wackestone 7=dolomite
# 8=packstone 9=bafflestone
facies_colors = ['#F4D03F', '#F5B041','#DC7633','#6E2C00',
'#1B4F72','#2E86C1', '#AED6F1', '#A569BD', '#196F3D']
facies_labels = ['SS', 'CSiS', 'FSiS', 'SiSh', 'MS',
'WS', 'D','PS', 'BS']
#facies_color_map is a dictionary that maps facies labels
#to their respective colors
facies_color_map = {}
for ind, label in enumerate(facies_labels):
facies_color_map[label] = facies_colors[ind]
def label_facies(row, labels):
return labels[ row['Facies'] -1]
training_data.loc[:,'FaciesLabels'] = training_data.apply(lambda row: label_facies(row, facies_labels), axis=1)
# -
def make_facies_log_plot(logs, facies_colors):
#make sure logs are sorted by depth
logs = logs.sort_values(by='Depth')
cmap_facies = colors.ListedColormap(
facies_colors[0:len(facies_colors)], 'indexed')
ztop=logs.Depth.min(); zbot=logs.Depth.max()
cluster=np.repeat(np.expand_dims(logs['Facies'].values,1), 100, 1)
f, ax = plt.subplots(nrows=1, ncols=6, figsize=(8, 12))
ax[0].plot(logs.GR, logs.Depth, '.g')
ax[1].plot(logs.ILD_log10, logs.Depth, '.')
ax[2].plot(logs.DeltaPHI, logs.Depth, '.', color='0.5')
ax[3].plot(logs.PHIND, logs.Depth, '.', color='r')
ax[4].plot(logs.PE, logs.Depth, '.', color='black')
im=ax[5].imshow(cluster, interpolation='none', aspect='auto',
cmap=cmap_facies,vmin=1,vmax=9)
divider = make_axes_locatable(ax[5])
cax = divider.append_axes("right", size="20%", pad=0.05)
cbar=plt.colorbar(im, cax=cax)
cbar.set_label((17*' ').join([' SS ', 'CSiS', 'FSiS',
'SiSh', ' MS ', ' WS ', ' D ',
' PS ', ' BS ']))
cbar.set_ticks(range(0,1)); cbar.set_ticklabels('')
for i in range(len(ax)-1):
ax[i].set_ylim(ztop,zbot)
ax[i].invert_yaxis()
ax[i].grid()
ax[i].locator_params(axis='x', nbins=3)
ax[0].set_xlabel("GR")
ax[0].set_xlim(logs.GR.min(),logs.GR.max())
ax[1].set_xlabel("ILD_log10")
ax[1].set_xlim(logs.ILD_log10.min(),logs.ILD_log10.max())
ax[2].set_xlabel("DeltaPHI")
ax[2].set_xlim(logs.DeltaPHI.min(),logs.DeltaPHI.max())
ax[3].set_xlabel("PHIND")
ax[3].set_xlim(logs.PHIND.min(),logs.PHIND.max())
ax[4].set_xlabel("PE")
ax[4].set_xlim(logs.PE.min(),logs.PE.max())
ax[5].set_xlabel('Facies')
ax[1].set_yticklabels([]); ax[2].set_yticklabels([]); ax[3].set_yticklabels([])
ax[4].set_yticklabels([]); ax[5].set_yticklabels([])
ax[5].set_xticklabels([])
f.suptitle('Well: %s'%logs.iloc[0]['Well Name'], fontsize=14,y=0.94)
# ### Remove Outlier
# Comment this block to delete outlier removal
[Scores,indices] = mad_based_outlier(training_data['GR'].values,3.5)
ind = np.where(indices==True)
training_data.drop(training_data.index[ind[0]],inplace=True)
[Scores,indices] = mad_based_outlier(training_data['ILD_log10'].values,3.5)
ind = np.where(indices==True)
training_data.drop(training_data.index[ind[0]],inplace=True)
[Scores,indices] = mad_based_outlier(training_data['DeltaPHI'].values,3.5)
ind = np.where(indices==True)
training_data.drop(training_data.index[ind[0]],inplace=True)
# ## Extract data
X = training_data.drop(['Formation', 'Well Name', 'Depth', 'Facies', 'FaciesLabels'], axis=1).values
y = training_data['Facies'].values - 1
wells = training_data["Well Name"].values
# ### Feature Augmentation method from Bestagini
# Feature windows concatenation function
def augment_features_window(X, N_neig):
# Parameters
N_row = X.shape[0]
N_feat = X.shape[1]
# Zero padding
X = np.vstack((np.zeros((N_neig, N_feat)), X, (np.zeros((N_neig, N_feat)))))
# Loop over windows
X_aug = np.zeros((N_row, N_feat*(2*N_neig+1)))
for r in np.arange(N_row)+N_neig:
this_row = []
for c in np.arange(-N_neig,N_neig+1):
this_row = np.hstack((this_row, X[r+c]))
X_aug[r-N_neig] = this_row
return X_aug
# Feature gradient computation function
def augment_features_gradient(X, depth):
# Compute features gradient
d_diff = np.diff(depth).reshape((-1, 1))
d_diff[d_diff==0] = 0.001
X_diff = np.diff(X, axis=0)
X_grad = X_diff / d_diff
# Compensate for last missing value
X_grad = np.concatenate((X_grad, np.zeros((1, X_grad.shape[1]))))
return X_grad
# Feature augmentation function
def augment_features(X, well, depth, N_neig=1):
# Augment features
X_aug = np.zeros((X.shape[0], X.shape[1]*(N_neig*2+2)))
for w in np.unique(well):
w_idx = np.where(well == w)[0]
X_aug_win = augment_features_window(X[w_idx, :], N_neig)
X_aug_grad = augment_features_gradient(X[w_idx, :], depth[w_idx])
X_aug[w_idx, :] = np.concatenate((X_aug_win, X_aug_grad), axis=1)
# Find padded rows
padded_rows = np.unique(np.where(X_aug[:, 0:7] == np.zeros((1, 7)))[0])
return X_aug, padded_rows
well = training_data['Well Name'].values
depth = training_data['Depth'].values
X, padded_rows = augment_features(X, well, depth, N_neig=1)
scaler = preprocessing.RobustScaler().fit(X)
X = scaler.transform(X)
Xtrain, Xtest, Ytrain, Ytest = train_test_split(X, y, train_size=0.7, test_size=0.3, random_state=0)
# +
# tpot = TPOTClassifier(scoring='f1_micro', random_state=0, max_eval_time_mins=1, max_time_mins=5, verbosity=1, num_cv_folds=2)
# tpot.fit(Xtrain, Ytrain)
# print(tpot.score(Xtest, Ytest))
# tpot.export('clf_pipeline0.py')
# -
from sklearn.ensemble import ExtraTreesClassifier, VotingClassifier, RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline, make_union
from sklearn.preprocessing import FunctionTransformer
# ## Validation with Leave One Well Out on Training Dataset
logo = LeaveOneGroupOut()
t0 = tm.time()
# +
f1s_ls = []
acc_ls = []
adj_ls = []
for train, test in logo.split(X, y, groups=wells):
well_name = wells[test[0]]
X_tr = X[train]
X_te = X[test]
Y_tr = y[train]
exported_pipeline = make_pipeline(
make_union(VotingClassifier([("est", RandomForestClassifier(n_estimators=250, n_jobs=4, random_state=42, min_samples_split=10,
max_depth=None, criterion='entropy', class_weight='balanced',
min_samples_leaf=5, max_features=15))]), FunctionTransformer(lambda X: X)),
ExtraTreesClassifier(criterion="entropy", max_features=1.0, n_estimators=500)
)
exported_pipeline.fit(X_tr, Y_tr)
y_hat = exported_pipeline.predict(X_te)
# y_hat = medfilt(y_hat, kernel_size=5)
try:
f1s = f1_score(y[test], y_hat, average="weighted", labels=[0, 1, 2, 3, 4, 5, 6, 7, 8])
except:
f1s = 0
try:
conf = confusion_matrix(y[test], y_hat, labels=[0, 1, 2, 3, 4, 5, 6, 7, 8])
acc = f1_score(y[test], y_hat, average="micro", labels=[0, 1, 2, 3, 4, 5, 6, 7, 8])
except:
acc = 0
try:
acc_adj = accuracy_adjacent(conf, adjacent_facies)
except:
acc_adj = 0
f1s_ls += [f1s]
acc_ls += [acc]
adj_ls += [acc_adj]
print("{:>20s} f1w:{:.3f} | f1m:{:.3f} | acc_adj:{:.3f}".format(well_name, f1s, acc, acc_adj))
t1 = tm.time()
print("Avg F1w", np.average(f1s_ls)*100, "Avg F1m", np.average(acc_ls)*100, "Avg Adj", np.average(adj_ls)*100)
print((t1-t0), "seconds")
# -
# ## Applying to Test Dataset
# +
blind_data = pd.read_csv('../nofacies_data.csv')
X_blind = blind_data.drop(['Formation', 'Well Name', 'Depth'], axis=1).values
well_blind = blind_data['Well Name'].values
depth_blind = blind_data['Depth'].values
# Removed padded rows
X = np.delete(X, padded_rows, axis=0)
y = np.delete(y, padded_rows, axis=0)
X_blind, padded_rows = augment_features(X_blind, well_blind, depth_blind, N_neig=1)
# -
# Scaling
X_train = X
X_blind = scaler.transform(X_blind)
# +
# # Method initialization
exported_pipeline = make_pipeline(
make_union(VotingClassifier([("est", RandomForestClassifier(n_estimators=250, n_jobs=4, random_state=42, min_samples_split=10,
max_depth=None, criterion='entropy', class_weight='balanced',
min_samples_leaf=5, max_features=15))]), FunctionTransformer(lambda X: X)),
ExtraTreesClassifier(criterion="entropy", max_features=1.0, n_estimators=500)
)
exported_pipeline.fit(X_train, y)
# Predict
y_blind = exported_pipeline.predict(X_blind)
y_blind = medfilt(y_blind, kernel_size=5)
blind_data["Facies"] = y_blind + 1 # return the original value (1-9)
# -
blind_data.to_csv("PA_Team_Submission_7_RF_01.csv")
make_facies_log_plot(
blind_data[blind_data['Well Name'] == 'STUART'],
facies_colors)
make_facies_log_plot(
blind_data[blind_data['Well Name'] == 'CRAWFORD'],
facies_colors)
| PA_Team_Submission_7_RF_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Right Angle Robot
# Before we jump into trigonometry, I want to familiarize you with the Vehicle class you'll be using in this lesson (and get you thinking about motion in general).
#
# In this notebook you will complete a `Vehicle` class by filling out two methods: `drive_forward` and `turn_right`.
#
# Note that this version of a Vehicle class can ONLY face in one of 4 directions: (E)ast, (N)orth, (W)est, or (S)outh. The vehicle's current direction is stored in its `heading` property.
#
# When you've implemented the two methods below you can run the testing cells at the bottom of the Notebook to ensure everything is behaving as expected.
#
# ### TODO - Implement `drive_forward` and `turn_right`
# **Solution code is provided in the next notebook.**
# +
from matplotlib import pyplot as plt
class Vehicle:
def __init__(self):
"""
Creates new vehicle at (0,0) with a heading pointed East.
"""
self.x = 0 # meters
self.y = 0
self.heading = "E" # Can be "N", "S", "E", or "W"
self.history = []
# TODO-1 - Implement this function
def drive_forward(self, displacement):
"""
Updates x and y coordinates of vehicle based on
heading and appends previous (x,y) position to
history.
"""
# this line appends the current (x,y) coordinates
# to the vehicle's history. Useful for plotting
# the vehicle's trajectory. You shouldn't need to
# change this line.
self.history.append((self.x, self.y))
# vehicle currently pointing east...
if self.heading == "E":
self.x += displacement
# north
elif self.heading == "N":
# FILL THIS OUT
self.y += displacement
# west
elif self.heading == "W":
# FILL THIS OUT
self.x -= displacement
# south
else:
# FILL THIS OUT
self.y -= displacement
def turn(self, direction):
if direction == "L":
self.turn_left()
elif direction == "R":
self.turn_right()
else:
print("Error. Direction must be 'L' or 'R'")
return
def turn_left(self):
"""
Updates heading (for a left turn) based on current heading
"""
next_heading = {
"N" : "W",
"W" : "S",
"S" : "E",
"E" : "N",
}
self.heading = next_heading[self.heading]
# TODO-2 - implement this function
def turn_right(self):
next_heading = {
"N" : "E",
"W" : "N",
"S" : "W",
"E" : "S",
}
self.heading = next_heading[self.heading]
def show_trajectory(self):
"""
Creates a scatter plot of vehicle's trajectory.
"""
X = [p[0] for p in self.history]
Y = [p[1] for p in self.history]
X.append(self.x)
Y.append(self.y)
plt.scatter(X,Y)
plt.plot(X,Y)
plt.show()
# +
# TESTING CODE 1
# instantiate vehicle
v = Vehicle()
# drive in spirals of decreasing size
v.drive_forward(8)
v.turn("L")
v.drive_forward(5)
v.turn("L")
v.drive_forward(5)
v.turn("L")
v.drive_forward(4)
v.turn("L")
v.drive_forward(4)
v.turn("L")
v.drive_forward(3)
v.turn("L")
v.drive_forward(3)
v.turn("L")
v.drive_forward(2)
v.turn("L")
v.drive_forward(2)
v.turn("L")
v.drive_forward(1)
v.turn("L")
v.drive_forward(1)
# show the trajectory. It should look like a spiral
v.show_trajectory()
# TESTING
assert(v.x == 5)
assert(v.y == 3)
first_5 = [
(0, 0),
(8, 0),
(8, 5),
(3, 5),
(3, 1)
]
assert(first_5 == v.history[:5])
print("Nice job! Your vehicle is behaving as expected!")
# -
# TESTING CODE Part 2
def test_zig_zag():
v = Vehicle()
for i in range(10):
for _ in range(4):
v.drive_forward(2)
v.turn("R")
v.drive_forward(1)
v.turn("L")
v.drive_forward(10)
v.turn("R")
first_six = [
(0,0),
(2,0),
(2,-1),
(4,-1),
(4,-2),
(6,-2)
]
v.show_trajectory()
assert(v.x == 14)
assert(v.y == -22)
assert(v.history[:6] == first_six)
print("Nice job! Your vehicle passed the zig zag test.")
test_zig_zag()
# ## What's Next?
#
# We want to be able to keep track of vehicle trajectory for ANY heading, not just the four compass directions.
| geometry_refresher/right_angle_robot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table border="0">
# <tr>
# <td>
# <img src="https://ictd2016.files.wordpress.com/2016/04/microsoft-research-logo-copy.jpg" style="width 30px;" />
# </td>
# <td>
# <img src="https://www.microsoft.com/en-us/research/wp-content/uploads/2016/12/MSR-ALICE-HeaderGraphic-1920x720_1-800x550.jpg" style="width 100px;"/></td>
# </tr>
# </table>
# # ForestDML, ForestDRLearner, OrthoForest and CausalForest: Basic Example
#
# We depict the performance of our `ForestDML`, `ForestDRLearner`, `OrthoForest` and `CausalForest` estimators on the same data generating process as the one used in the tutorial page of the grf package (see https://github.com/grf-labs/grf#usage-examples). This is mostly for qualitative comparison and verification purposes among our implementation of variants of Causal Forests and the implementation in the grf R package.
# %load_ext autoreload
# %autoreload 2
## Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# Helper imports
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
# # DGP
#
# We use the following data generating process (DGP) from [here](https://github.com/grf-labs/grf#usage-examples):
#
# \begin{align}
# X \sim& \text{Normal}(0,\, I_{p})\\
# T =& \text{Binomial}(1, .4 + .2 \cdot 1\{X[0] > 0\})\\
# Y =& (X[0] \cdot 1\{X[0] > 0\}) \cdot T + X[1] + X[2] \cdot 1\{X[2] < 0\} + \epsilon, &\; \epsilon \sim \text{Normal}(0, 1)\\
# \end{align}
#
# We use $p=10$ and draw $n=2000$ samples from this DGP.
import numpy as np
import scipy.special
np.random.seed(123)
n = 2000
p = 10
X = np.random.normal(size=(n, p))
true_propensity = lambda x: .4 + .2 * (x[:, 0] > 0)
true_effect = lambda x: (x[:, 0] * (x[:, 0] > 0))
true_conf = lambda x: x[:, 1] + np.clip(x[:, 2], - np.inf, 0)
T = np.random.binomial(1, true_propensity(X))
Y = true_effect(X) * T + true_conf(X) + np.random.normal(size=(n,))
# ### Cross-Validated Forest Nuisance Models
#
# We use forest based estimators (Gradient Boosted Forests or Random Forests) as nuisance models. For the meta-learner versions of our forest based estimators, we also use a generic forest estimator even as a final model. The hyperparameters of the forest models (e.g. number of estimators, max depth, min leaf size) is chosen via cross validation. We also choose among Gradient or Random Forests via cross validation
# +
from econml.sklearn_extensions.model_selection import GridSearchCVList
from sklearn.linear_model import Lasso, LogisticRegression
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.base import clone
from econml.sklearn_extensions.linear_model import WeightedLasso
def first_stage_reg():
return GridSearchCVList([Lasso(),
RandomForestRegressor(n_estimators=100, random_state=123),
GradientBoostingRegressor(random_state=123)],
param_grid_list=[{'alpha': [.001, .01, .1, 1, 10]},
{'max_depth': [3, None],
'min_samples_leaf': [10, 50]},
{'n_estimators': [50, 100],
'max_depth': [3],
'min_samples_leaf': [10, 30]}],
cv=5,
scoring='neg_mean_squared_error')
def first_stage_clf():
return GridSearchCVList([LogisticRegression(),
RandomForestClassifier(n_estimators=100, random_state=123),
GradientBoostingClassifier(random_state=123)],
param_grid_list=[{'C': [0.01, .1, 1, 10, 100]},
{'max_depth': [3, 5],
'min_samples_leaf': [10, 50]},
{'n_estimators': [50, 100],
'max_depth': [3],
'min_samples_leaf': [10, 30]}],
cv=5,
scoring='neg_mean_squared_error')
def final_stage():
return GridSearchCVList([WeightedLasso(),
RandomForestRegressor(n_estimators=100, random_state=123)],
param_grid_list=[{'alpha': [.001, .01, .1, 1, 10]},
{'max_depth': [3, 5],
'min_samples_leaf': [10, 50]}],
cv=5,
scoring='neg_mean_squared_error')
# -
model_y = clone(first_stage_reg().fit(X, Y).best_estimator_)
model_y
model_t = clone(first_stage_clf().fit(X, T).best_estimator_)
model_t
# # DML Estimators
# +
from econml.dml import ForestDML
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from sklearn.ensemble import GradientBoostingRegressor, GradientBoostingClassifier
from sklearn.dummy import DummyRegressor, DummyClassifier
est = ForestDML(model_y=model_y,
model_t=model_t,
discrete_treatment=True,
n_crossfit_splits=3,
n_estimators=4000,
subsample_fr='auto',
min_samples_leaf=10,
verbose=0,
min_weight_fraction_leaf=.005)
est.fit(Y, T, X=X)
# -
from econml.dml import NonParamDML
est2 = NonParamDML(model_y=model_y,
model_t=model_t,
n_splits=3,
discrete_treatment=True,
model_final=final_stage())
est2.fit(Y, T, X=X)
X_test = np.zeros((100, p))
X_test[:, 0] = np.linspace(-2, 2, 100)
pred = est.effect(X_test)
lb, ub = est.effect_interval(X_test, alpha=0.01)
pred2 = est2.effect(X_test)
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.plot(X_test[:, 0], true_effect(X_test), '--')
plt.plot(X_test[:, 0], pred2, label='nonparamdml')
plt.plot(X_test[:, 0], pred, label='forestdml (causal forest)')
plt.fill_between(X_test[:, 0], lb, ub, alpha=.4, label='honestrf_ci')
plt.legend()
plt.show()
# ### First Stage Learned Models
# +
# Model T
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.title('honestrf')
for mdl in est.models_t:
plt.plot(X_test[:, 0], mdl.predict_proba(X_test)[:, 1])
plt.plot(X_test[:, 0], true_propensity(X_test), '--', label='truth')
plt.legend()
plt.subplot(1, 2, 2)
plt.title('rf')
for mdl in est2.models_t:
plt.plot(X_test[:, 0], mdl.predict_proba(X_test)[:, 1])
plt.plot(X_test[:, 0], true_propensity(X_test), '--', label='truth')
plt.legend()
plt.show()
# +
# Model Y
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.title('honestrf')
for mdl in est.models_y:
plt.plot(X_test[:, 0], mdl.predict(X_test))
plt.plot(X_test[:, 0], true_effect(X_test) * true_propensity(X_test) + true_conf(X_test), '--', label='truth')
plt.legend()
plt.subplot(1, 2, 2)
plt.title('rf')
for mdl in est2.models_y:
plt.plot(X_test[:, 0], mdl.predict(X_test))
plt.plot(X_test[:, 0], true_effect(X_test) * true_propensity(X_test) + true_conf(X_test), '--', label='truth')
plt.legend()
plt.show()
# -
# ### Interpretability of CATE Model of NonParamDML with SHAP
import shap
import string
feature_names=list(string.ascii_lowercase)[:X.shape[1]]
# explain the model's predictions using SHAP values
shap_values = est.shap_values(X[:100],feature_names=feature_names)
# visualize the first prediction's explanation (use matplotlib=True to avoid Javascript)
shap.force_plot(shap_values["Y0"]["T0"][0], matplotlib=True)
shap.summary_plot(shap_values["Y0"]["T0"])
# # DRLearner
model_regression = clone(first_stage_reg().fit(np.hstack([T.reshape(-1, 1), X]), Y).best_estimator_)
model_regression
# +
from econml.drlearner import ForestDRLearner
from sklearn.dummy import DummyRegressor, DummyClassifier
est = ForestDRLearner(model_regression=model_regression,
model_propensity=model_t,
n_crossfit_splits=3,
n_estimators=4000,
subsample_fr='auto',
min_samples_leaf=10,
verbose=0,
min_weight_fraction_leaf=.005)
est.fit(Y, T, X=X)
# -
from econml.drlearner import DRLearner
est2 = DRLearner(model_regression=model_regression,
model_propensity=model_t,
model_final=final_stage(),
n_splits=3)
est2.fit(Y, T, X=X)
X_test = np.zeros((100, p))
X_test[:, 0] = np.linspace(-2, 2, 100)
pred = est.effect(X_test)
lb, ub = est.effect_interval(X_test, alpha=0.01)
pred2 = est2.effect(X_test)
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.plot(X_test[:, 0], true_effect(X_test), '--')
plt.plot(X_test[:, 0], pred2, label='nonparamdml')
plt.plot(X_test[:, 0], pred, label='forestdml (causal forest)')
plt.fill_between(X_test[:, 0], lb, ub, alpha=.4, label='honestrf_ci')
plt.legend()
plt.show()
# ### First stage nuisance models
# +
# Model T
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.title('honestrf')
for mdl in est.models_propensity:
plt.plot(X_test[:, 0], mdl.predict_proba(X_test)[:, 1])
plt.plot(X_test[:, 0], true_propensity(X_test), '--', label='truth')
plt.legend()
plt.subplot(1, 2, 2)
plt.title('rf')
for mdl in est2.models_propensity:
plt.plot(X_test[:, 0], mdl.predict_proba(X_test)[:, 1])
plt.plot(X_test[:, 0], true_propensity(X_test), '--', label='truth')
plt.legend()
plt.show()
# +
# Model Y
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.title('honestrf')
for mdl in est.models_regression:
plt.plot(X_test[:, 0], mdl.predict(np.hstack([X_test, np.ones((X_test.shape[0], 1))])))
plt.plot(X_test[:, 0], true_effect(X_test) + true_conf(X_test), '--', label='truth')
plt.legend()
plt.subplot(1, 2, 2)
plt.title('rf')
for mdl in est2.models_regression:
plt.plot(X_test[:, 0], mdl.predict(np.hstack([X_test, np.ones((X_test.shape[0], 1))])))
plt.plot(X_test[:, 0], true_effect(X_test) + true_conf(X_test), '--', label='truth')
plt.legend()
plt.show()
# -
# ### Interpretability of CATE Model of DRLearner with SHAP
# explain the model's predictions using SHAP values
shap_values = est.shap_values(X[:100],feature_names=feature_names)
# visualize the first prediction's explanation (use matplotlib=True to avoid Javascript)
shap.force_plot(shap_values["Y0"]["T0"][0], matplotlib=True)
shap.summary_plot(shap_values["Y0"]["T0"])
# # OrthoForest and CausalForest
# +
from econml.ortho_forest import DROrthoForest
from sklearn.linear_model import Lasso, LassoCV, LogisticRegression, LogisticRegressionCV
from econml.sklearn_extensions.linear_model import WeightedLassoCV
est3 = DROrthoForest(model_Y=Lasso(alpha=0.01),
propensity_model=LogisticRegression(C=1),
model_Y_final=WeightedLassoCV(cv=3),
propensity_model_final=LogisticRegressionCV(cv=3),
n_trees=1000, min_leaf_size=10)
est3.fit(Y, T, X=X)
# -
pred3 = est3.effect(X_test)
# +
from econml.causal_forest import CausalForest
from sklearn.linear_model import Lasso, LassoCV, LogisticRegression, LogisticRegressionCV
from econml.sklearn_extensions.linear_model import WeightedLassoCV
est4 = CausalForest(model_Y=model_y,
model_T=model_t,
n_trees=1000, min_leaf_size=10,
discrete_treatment=True,
cv=5)
est4.fit(Y, T, X=X)
# -
pred4 = est4.effect(X_test)
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.plot(X_test[:, 0], true_effect(X_test), '--')
plt.plot(X_test[:, 0], pred, label='forestdr')
plt.plot(X_test[:, 0], pred2, label='nonparamdr')
plt.plot(X_test[:, 0], pred3, label='discreteorf')
plt.plot(X_test[:, 0], pred4, label='causalforest')
plt.fill_between(X_test[:, 0], lb, ub, alpha=.4, label='forest_dr_ci')
plt.legend()
plt.show()
| notebooks/ForestLearners Basic Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HEiOAwQPW0qb" colab_type="code" colab={}
import pandas as pd
# + id="j7whidfaYjns" colab_type="code" colab={}
file_url = 'http://raw.githubusercontent.com/PacktWorkshops/The-Data-Science-Workshop/master/Chapter11/dataset/horse-colic.data'
# + id="nFMz2jNVt-xy" colab_type="code" colab={}
df = pd.read_csv(file_url, header=None, sep='\s+', prefix='X')
# + id="n0xY9ATvN6-M" colab_type="code" outputId="8deac724-2a1e-4a73-abde-a2cca7f9cfe3" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + id="ASEdozexcGbY" colab_type="code" colab={}
df = pd.read_csv(file_url, header=None, sep='\s+', prefix='X', na_values='?')
# + id="NLiPeTsPcHpg" colab_type="code" outputId="26468262-5891-4d2a-b801-a2af377f0f7f" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + id="Rv1a7YLL63I8" colab_type="code" outputId="4e5dd925-59d8-43b2-ac37-5221de09052c" colab={"base_uri": "https://localhost:8080/", "height": 510}
df.dtypes
# + id="q1a8onHi79Z7" colab_type="code" outputId="5c3137bc-455e-4f93-f9c2-9aebffd34141" colab={"base_uri": "https://localhost:8080/", "height": 510}
df.isna().sum()
# + id="1zTWk7dtBbMe" colab_type="code" colab={}
x0_mask = df['X0'].isna()
# + id="apH4JpnGBgNG" colab_type="code" outputId="c808828a-7081-478e-b4a2-1de161ae1619" colab={"base_uri": "https://localhost:8080/", "height": 34}
x0_mask.sum()
# + id="I0wuvkFHBlml" colab_type="code" outputId="4bb32466-71e6-496b-fceb-d0d3e138d25f" colab={"base_uri": "https://localhost:8080/", "height": 34}
x0_median = df['X0'].median()
print(x0_median)
# + id="Q_m65ZKLB-jx" colab_type="code" colab={}
df['X0'].fillna(x0_median, inplace=True)
# + id="Jr6BfU45CDQw" colab_type="code" outputId="01236dae-58f5-4768-81f7-66515325fb7b" colab={"base_uri": "https://localhost:8080/", "height": 34}
df['X0'].isna().sum()
# + id="MH3Qd9LIHeKF" colab_type="code" outputId="4bd4e62a-1bef-4301-faa8-07607c134a66" colab={"base_uri": "https://localhost:8080/", "height": 969}
for col_name in df.columns:
col_median = df[col_name].median()
df[col_name].fillna(col_median, inplace=True)
print(col_name)
print(col_median)
# + id="T-CLjjkVHrfq" colab_type="code" outputId="04e8837c-39ad-49e6-ada4-0f0352151761" colab={"base_uri": "https://localhost:8080/", "height": 510}
df.isna().sum()
| Chapter11/Exercise11.04/Exercise11_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Natural Language Analytics
#
# ## Exercise 3 - Topic Modeling and NER
#
# ### <NAME> - DSC18014
# ## Question A
#
# First let's load our libraries and our data
import pandas as pd
import glob
import os
from sklearn.model_selection import GridSearchCV
from sklearn.decomposition import NMF, LatentDirichletAllocation, TruncatedSVD
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
from sklearn.metrics.pairwise import euclidean_distances
import pyLDAvis.sklearn
# After we download we take a peek
subdir = os.listdir('nipstxt')
print(subdir)
# Let's exclude the files and folders that notes, indices and the orig folder
subdir = [ item for item in subdir if "nips" in item]
print(subdir)
lst_docs = []
for dirs in subdir:
path = 'nipstxt/' + dirs + '/*.txt'
files = glob.glob(path)
for name in files:
with open(name, encoding="utf8", errors='ignore') as f:
lst_docs.append(f.read())
# Ok, now we have all our documents as items in a list.
# As a next step we will process the texts and transform the to vectors.
# We will experiment with computing both the inverse term frequency (TFIDF vectorizer)and Term frequency (sklearn's Count Vectorizer).
# #### TFIDF Vectorizer
# We choose to tokenize by english words that are 3 lettered and longer. We also consider to tokenizer by unigrams
extr = TfidfVectorizer(min_df=5, max_df=0.9, max_features=None, strip_accents='unicode', lowercase=True, analyzer='word',
token_pattern=r'\w{3,}', ngram_range=(1, 1), use_idf=True, smooth_idf=True,
sublinear_tf=True, stop_words="english")
# Let's vectorize our data
transformed_data = extr.fit_transform(lst_docs)
# Sklearn assumes alpha=1/n_components if no value is entered for doc_topic_prior. So in our exercise we will test for n_components 10 and 100. So our alpha will be 0.1 and 0.01
# +
NUM_TOPICS = 10
# Build a Latent Dirichlet Allocation Model
lda_model = LatentDirichletAllocation(n_components=NUM_TOPICS, max_iter=10, learning_method='online')
lda_Z = lda_model.fit_transform(transformed_data)
print(lda_Z.shape) # (NO_DOCUMENTS, NO_TOPICS)
# -
# Let's see how the first document in the corpus looks like in different topic spaces
print(lda_Z[0])
# +
def print_topics(model, vectorizer, top_n=10):
for idx, topic in enumerate(model.components_):
print("Topic %d:" % (idx))
print([(vectorizer.get_feature_names()[i], topic[i]) for i in topic.argsort()[:-top_n - 1:-1]])
print("Most representative document: ", np.argmax(topic))
print_topics(lda_model, extr)
# -
# wow, only one describes best the tokens. Let's see the distribution of topics across the documents
win_topic=[np.argmax(topics) for topics in lda_Z]
plt.plot(win_topic)
plt.show()
# As expected... all documents could belong to topic 7
# Let's assume a new text and try to find the most similar document from our dataset
text = "the vlsi network makes no sense"
x = lda_model.transform(extr.transform([text]))[0]
print(x)
# +
def most_similar(x, Z, top_n=5):
dists = euclidean_distances(x.reshape(1, -1), Z)
pairs = enumerate(dists[0])
most_similar = sorted(pairs, key=lambda item: item[1])[:top_n]
return most_similar
similarities = most_similar(x, lda_Z)
document_id, similarity = similarities[0]
print(lst_docs[document_id][:1000])
# -
# To visualise the results of LDA let's import pyLDAvis
pyLDAvis.enable_notebook()
panel = pyLDAvis.sklearn.prepare(lda_model, transformed_data, extr, mds='tsne', sort_topics=False)
panel
# Well, one topic seems to dominate the majority of tokens. Here counting starts from 1, so it's the same as the previously found
# +
# Log Likelyhood: Higher the better
print("Log Likelihood: ", lda_model.score(transformed_data))
# Perplexity: Lower the better. Perplexity = exp(-1. * log-likelihood per word)
print("Perplexity: ", lda_model.perplexity(transformed_data))
# -
# Let's do a grid search
# +
# Define Search Param
search_params = {'doc_topic_prior':[.01,.1,.2,.5],'n_components': [5, 10, 50], 'learning_decay': [.2, .5, .7]}
# Init the Model
lda = LatentDirichletAllocation(max_iter=10, learning_method='online')
# Init Grid Search Class
model = GridSearchCV(lda, param_grid=search_params)
# Do the Grid Search
model.fit(transformed_data)
# Best Model
best_lda_model = model.best_estimator_
# Model Parameters
print("Best Model's Params: ", model.best_params_)
# Log Likelihood Score
print("Best Log Likelihood Score: ", model.best_score_)
# Perplexity
print("Model Perplexity: ", best_lda_model.perplexity(transformed_data))
lda_Z = best_lda_model.fit_transform(transformed_data)
#distribution
win_topic=[np.argmax(topics) for topics in lda_Z]
# -
from matplotlib.pyplot import figure
figure(num=None, figsize=(15, 6), dpi=80, facecolor='w', edgecolor='k')
plt.plot(win_topic)
plt.show()
print_topics(lda_model, extr)
# +
# Log Likelyhood: Higher the better
print("Log Likelihood: ", lda_model.score(transformed_data))
# Perplexity: Lower the better. Perplexity = exp(-1. * log-likelihood per word)
print("Perplexity: ", lda_model.perplexity(transformed_data))
# -
# Let's try to change the way we represent our features and see how LDA responds
# Again for a =0.1
# +
extr = CountVectorizer(min_df=5, max_df=0.9, stop_words='english', lowercase=True, token_pattern=r'\w{3,}')
transformed_data = extr.fit_transform(lst_docs)
NUM_TOPICS = 10
# Build a Latent Dirichlet Allocation Model
lda_model = LatentDirichletAllocation(n_components=NUM_TOPICS, max_iter=10, learning_method='online')
lda_Z = lda_model.fit_transform(transformed_data)
#distribution
win_topic=[np.argmax(topics) for topics in lda_Z]
# -
figure(num=None, figsize=(15, 6), dpi=80, facecolor='w', edgecolor='k')
plt.plot(win_topic)
plt.show()
print_topics(lda_model, extr)
# +
# Log Likelyhood: Higher the better
print("Log Likelihood: ", lda_model.score(transformed_data))
# Perplexity: Lower the better. Perplexity = exp(-1. * log-likelihood per word)
print("Perplexity: ", lda_model.perplexity(transformed_data))
# -
# pyLDAvis.enable_notebook()
panel = pyLDAvis.sklearn.prepare(lda_model, transformed_data, extr, mds='tsne', sort_topics=False)
panel
# With the Count Vectorizer we have a more meaningful result. Topic 7 is slightly more popular that the others.
# Let's do a grid search.
# +
# Define Search Param
search_params = {'doc_topic_prior':[.01,.1,.2,.5],'n_components': [5, 10, 50], 'learning_decay': [.2, .5, .7]}
# Init the Model
lda = LatentDirichletAllocation(max_iter=10, learning_method='online')
# Init Grid Search Class
model = GridSearchCV(lda, param_grid=search_params)
# Do the Grid Search
model.fit(transformed_data)
# Best Model
best_lda_model = model.best_estimator_
# Model Parameters
print("Best Model's Params: ", model.best_params_)
# Log Likelihood Score
print("Best Log Likelihood Score: ", model.best_score_)
# Perplexity
print("Model Perplexity: ", best_lda_model.perplexity(transformed_data))
lda_Z = best_lda_model.fit_transform(transformed_data)
#distribution
win_topic=[np.argmax(topics) for topics in lda_Z]
figure(num=None, figsize=(15, 6), dpi=80, facecolor='w', edgecolor='k')
plt.plot(win_topic)
plt.show()
# -
# ## Question B
# +
import collections
ner_tags = collections.Counter()
corpus_root = "gmb-2.2.0"
for root, dirs, files in os.walk(corpus_root):
for filename in files:
if filename.endswith(".tags"):
with open(os.path.join(root, filename), 'rb') as file_handle:
file_content = file_handle.read().decode('utf-8').strip()
annotated_sentences = file_content.split('\n\n') # Split sentences
for annotated_sentence in annotated_sentences:
annotated_tokens = [seq for seq in annotated_sentence.split('\n') if seq] # Split words
standard_form_tokens = []
for idx, annotated_token in enumerate(annotated_tokens):
annotations = annotated_token.split('\t') # Split annotation
word, tag, ner = annotations[0], annotations[1], annotations[3]
# Get only the primary category
if ner != 'O':
ner = ner.split('-')[0]
ner_tags[ner] += 1
print(ner_tags)
print("Words=", sum(ner_tags.values()))
# +
import string
from nltk.stem.snowball import SnowballStemmer
def features(tokens, index, history):
"""
`tokens` = a POS-tagged sentence [(w1, t1), ...]
`index` = the index of the token we want to extract features for
`history` = the previous predicted IOB tags
"""
# init the stemmer
stemmer = SnowballStemmer('english')
# Pad the sequence with placeholders
tokens = [('[START2]', '[START2]'), ('[START1]', '[START1]')] + list(tokens) + [('[END1]', '[END1]'), ('[END2]', '[END2]')]
history = ['[START2]', '[START1]'] + list(history)
# shift the index with 2, to accommodate the padding
index += 2
word, pos = tokens[index]
prevword, prevpos = tokens[index - 1]
prevprevword, prevprevpos = tokens[index - 2]
nextword, nextpos = tokens[index + 1]
nextnextword, nextnextpos = tokens[index + 2]
previob = history[index - 1]
contains_dash = '-' in word
contains_dot = '.' in word
allascii = all([True for c in word if c in string.ascii_lowercase])
allcaps = word == word.capitalize()
capitalized = word[0] in string.ascii_uppercase
prevallcaps = prevword == prevword.capitalize()
prevcapitalized = prevword[0] in string.ascii_uppercase
nextallcaps = prevword == prevword.capitalize()
nextcapitalized = prevword[0] in string.ascii_uppercase
return {
'word': word,
'lemma': stemmer.stem(word),
'pos': pos,
'all-ascii': allascii,
'next-word': nextword,
'next-lemma': stemmer.stem(nextword),
'next-pos': nextpos,
'next-next-word': nextnextword,
'nextnextpos': nextnextpos,
'prev-word': prevword,
'prev-lemma': stemmer.stem(prevword),
'prev-pos': prevpos,
'prev-prev-word': prevprevword,
'prev-prev-pos': prevprevpos,
'prev-iob': previob,
'contains-dash': contains_dash,
'contains-dot': contains_dot,
'all-caps': allcaps,
'capitalized': capitalized,
'prev-all-caps': prevallcaps,
'prev-capitalized': prevcapitalized,
'next-all-caps': nextallcaps,
'next-capitalized': nextcapitalized,
}
# +
def to_conll_iob(annotated_sentence):
"""
`annotated_sentence` = list of triplets [(w1, t1, iob1), ...]
Transform a pseudo-IOB notation: O, PERSON, PERSON, O, O, LOCATION, O
to proper IOB notation: O, B-PERSON, I-PERSON, O, O, B-LOCATION, O
"""
proper_iob_tokens = []
for idx, annotated_token in enumerate(annotated_sentence):
tag, word, ner = annotated_token
if ner != 'O':
if idx == 0:
ner = "B-" + ner
elif annotated_sentence[idx - 1][2] == ner:
ner = "I-" + ner
else:
ner = "B-" + ner
proper_iob_tokens.append((tag, word, ner))
return proper_iob_tokens
def read_gmb(corpus_root):
for root, dirs, files in os.walk(corpus_root):
for filename in files:
if filename.endswith(".tags"):
with open(os.path.join(root, filename), 'rb') as file_handle:
file_content = file_handle.read().decode('utf-8').strip()
annotated_sentences = file_content.split('\n\n')
for annotated_sentence in annotated_sentences:
annotated_tokens = [seq for seq in annotated_sentence.split('\n') if seq]
standard_form_tokens = []
for idx, annotated_token in enumerate(annotated_tokens):
annotations = annotated_token.split('\t')
word, tag, ner = annotations[0], annotations[1], annotations[3]
if ner != 'O':
ner = ner.split('-')[0]
if tag in ('LQU', 'RQU'): # Make it NLTK compatible
tag = "``"
standard_form_tokens.append((word, tag, ner))
conll_tokens = to_conll_iob(standard_form_tokens)
# Make it NLTK Classifier compatible - [(w1, t1, iob1), ...] to [((w1, t1), iob1), ...]
# Because the classfier expects a tuple as input, first item input, second the class
yield [((w, t), iob) for w, t, iob in conll_tokens]
reader = read_gmb(corpus_root)
# -
print (next(reader))
# +
import pickle
from collections import Iterable
from nltk.tag import ClassifierBasedTagger
from nltk.chunk import ChunkParserI, conlltags2tree, tree2conlltags
class NamedEntityChunker(ChunkParserI):
def __init__(self, train_sents, **kwargs):
assert isinstance(train_sents, Iterable)
self.feature_detector = features
self.tagger = ClassifierBasedTagger(
train=train_sents,
feature_detector=features,
**kwargs)
def parse(self, tagged_sent):
chunks = self.tagger.tag(tagged_sent)
# Transform the result from [((w1, t1), iob1), ...]
# to the preferred list of triplets format [(w1, t1, iob1), ...]
iob_triplets = [(w, t, c) for ((w, t), c) in chunks]
# Transform the list of triplets to nltk.Tree format
return conlltags2tree(iob_triplets)
# +
reader = read_gmb(corpus_root)
data = list(reader)
training_samples = data[:int(len(data) * 0.9)]
test_samples = data[int(len(data) * 0.9):]
print ("#training samples = %s" % len(training_samples))
print ("#test samples = %s" % len(test_samples) )
# -
chunker = NamedEntityChunker(training_samples[:2000])
from nltk import pos_tag, word_tokenize
print (chunker.parse(pos_tag(word_tokenize("I'm going to Germany this Monday."))))
| Python/Topic Modeling and NER.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing HLS Fast Corner Detection Module
#
# The HLS module performs 'fast corner detection'
#
# The project builds on the VDMA demo.
#
# ## Project sources can be found here
#
# [HLS Corner Detection](https://github.com/CospanDesign/pynq-hdl/tree/master/Projects/Simple%20HLS%20Corners%20VDMA)
# + deletable=true editable=true
import cv2
import numpy as np
def cvtcolor_rgb2yuv422(rgb):
yuv422 =np.zeros((rgb.shape[0], rgb.shape[1], 2)).astype(np.uint8)
yuv444 = cv2.cvtColor(rgb, cv2.COLOR_BGR2YUV);
# chroma subsampling: yuv444 -> yuv422;
for row in range(yuv444.shape[0]):
for col in range(0, yuv444.shape[1], 2):
p0_in = yuv444[row, col]
p1_in = yuv444[row, col + 1]
p0_out = [p0_in[0], p0_in[1]]
p1_out = [p1_in[0], p0_in[2]]
yuv422[row, col] = p0_out
yuv422[row, col + 1] = p1_out
return yuv422
# -
# # Open and Convert the Image to a usable format
#
# Open the image and convert it to YUV422
#
# Perform the conversion in a seperate cell than below because the conversion takes a long time.
# + deletable=true editable=true
# # %matplotlib inline
from matplotlib import pyplot as plt
#Create a YUV422 Image So we don't need to keep regenerating it
IMAGE_FILE = "../data/test_1080p.bmp"
image_in = cv2.imread(IMAGE_FILE)
image_yuv = cvtcolor_rgb2yuv422(image_in)
#SHOW IMAGE
image_out = cv2.cvtColor(image_yuv, cv2.COLOR_YUV2BGR_YUYV)
plt.imshow(image_out)
plt.show()
# -
# # Perform the Image Processing
#
# 1. Program the FPGA.
# 2. Configure the Egress and Ingress Video DMA cores and configure them to take in images with the with and height the same as the image opened.
# 3. Configure the Image Processor.
# 4. Send down the image to the memory accessable by the FPGA.
# 5. Intitate the VDMA Transfer.
# 6. Wait for the transfer to finish.
# 7. Read back and display the image
# + deletable=true editable=true
# # %matplotlib inline
from time import sleep
from pynq import Overlay
from pynq.drivers import VDMA
from image_processor import ImageProcessor
import cv2
from matplotlib import pyplot as plt
from IPython.display import Image
import numpy as np
#Constants
BITFILE_NAME = "hls_corners.bit"
EGRESS_VDMA_NAME = "SEG_axi_vdma_0_Reg"
INGRESS_VDMA_NAME = "SEG_axi_vdma_1_Reg"
HLS_NAME = "SEG_image_filter_0_Reg"
# Set Debug to true to enable debug messages from the VDMA core
DEBUG = False
#DEBUG = True
# Set Verbose to true to dump a lot of messages about
VERBOSE = False
#VERBOSE = True
#These can be set between 0 - 2, the VDMA can also be configured for up to 32 frames in 32-bit memspace and 16 in 64-bit memspace
EGRESS_FRAME_INDEX = 0
INGRESS_FRAME_INDEX = 0
IMAGE_WIDTH = image_yuv.shape[1]
IMAGE_HEIGHT = image_yuv.shape[0]
print ("Image Size: %dx%d" % (IMAGE_WIDTH, IMAGE_HEIGHT))
#Download Images
ol = Overlay(BITFILE_NAME)
ol.download()
vdma_egress = VDMA(name = EGRESS_VDMA_NAME, debug = DEBUG)
vdma_ingress = VDMA(name = INGRESS_VDMA_NAME, debug = DEBUG)
image_processor = ImageProcessor(HLS_NAME)
image_processor.set_image_width(IMAGE_WIDTH)
image_processor.set_image_height(IMAGE_HEIGHT)
image_processor.enable(True)
#print ("Image Processor Enabled? %s" % image_processor.is_enabled())
#Set the size of the image
vdma_egress.set_image_size(IMAGE_WIDTH, IMAGE_HEIGHT, color_depth = 2)
vdma_ingress.set_image_size(IMAGE_WIDTH, IMAGE_HEIGHT, color_depth = 2)
#The above functions created the video frames
#Populate the frame
frame = vdma_egress.get_frame(EGRESS_FRAME_INDEX)
frame.set_bytearray(bytearray(image_yuv.astype(np.int8).tobytes()))
print ("Frame width, height: %d, %d" % (frame.width, frame.height))
print ("")
print ("Running? Egress:Ingress %s:%s" % (vdma_egress.is_egress_enabled(), vdma_ingress.is_ingress_enabled()))
if VERBOSE:
vdma_egress.dump_egress_registers()
vdma_ingress.dump_ingress_registers()
print ("")
print ("Enabling One of the Engine")
#Open Up the Ingress Side
vdma_ingress.start_ingress_engine( continuous = False,
num_frames = 1,
frame_index = INGRESS_FRAME_INDEX,
interrupt = False)
if VERBOSE:
vdma_egress.dump_egress_registers()
vdma_ingress.dump_ingress_registers()
print ("Running? Egress:Ingress %s:%s" % (vdma_egress.is_egress_enabled(), vdma_ingress.is_ingress_enabled()))
print ("")
print ("Enabling Both Engines")
#Quick Start
vdma_egress.start_egress_engine( continuous = False,
num_frames = 1,
frame_index = EGRESS_FRAME_INDEX,
interrupt = False)
print ("")
print ("Both of the engines should be halted after transferring one frame")
#XXX: I think this sleep isn't needed but the core erroniously reports an engine isn't finished even though it is.
#XXX: This sleep line can be commented out but the egress core may report it is not finished.
sleep(0.1)
if VERBOSE:
vdma_egress.dump_egress_registers()
vdma_ingress.dump_ingress_registers()
print ("Running? Egress:Ingress %s:%s" % (vdma_egress.is_egress_enabled(), vdma_ingress.is_ingress_enabled()))
if VERBOSE:
print ("Egress WIP: %d" % vdma_egress.get_wip_egress_frame())
print ("Ingress WIP: %d" % vdma_ingress.get_wip_ingress_frame())
#Check to see if the egress frame point progressed
print ("")
print ("Disabling both engines")
#Disable both
vdma_egress.stop_egress_engine()
vdma_ingress.stop_ingress_engine()
print ("Running? Egress:Ingress %s:%s" % (vdma_egress.is_egress_enabled(), vdma_ingress.is_ingress_enabled()))
if VERBOSE:
vdma_egress.dump_egress_registers()
vdma_ingress.dump_ingress_registers()
print ("Egress Error: 0x%08X" % vdma_egress.get_egress_error())
print ("Ingress Error: 0x%08X" % vdma_ingress.get_ingress_error())
frame = vdma_ingress.get_frame(INGRESS_FRAME_INDEX)
#frame.save_as_jpeg("./image.jpg")
image_yuv_out = np.ndarray( shape = (IMAGE_HEIGHT, IMAGE_WIDTH, 2),
dtype=np.uint8,
buffer = frame.get_bytearray())
image_rgb_out = cv2.cvtColor(image_yuv_out, cv2.COLOR_YUV2BGR_YUYV)
#SHOW IMAGE
plt.imshow(image_rgb_out)
plt.show()
| hls_corners/hls_corners.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="24XqHBcdwaXt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 666} outputId="be116d38-9c63-4315-8b64-1e6b128d63ea" executionInfo={"status": "ok", "timestamp": 1583496799524, "user_tz": -60, "elapsed": 14610, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# !pip install hyperopt
# + id="nNYV0YbMw8ev" colab_type="code" colab={}
import pandas as pd
import numpy as np
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score
from hyperopt import hp, fmin, tpe, STATUS_OK
import eli5
from eli5.sklearn import PermutationImportance
# + id="akVhUV931IgJ" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="y4tzxCPHxjsW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="025a7907-1435-4c15-c132-963cfd3f03f2" executionInfo={"status": "ok", "timestamp": 1583497027064, "user_tz": -60, "elapsed": 1149, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>FQ=s64", "userId": "16732949555547479927"}}
# cd "/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car"
# + id="tBg8jh8xxsl0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="96883fdf-16b5-4d1b-c461-0c1b168f2c8c" executionInfo={"status": "ok", "timestamp": 1583498284474, "user_tz": -60, "elapsed": 4385, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>VRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="C_aobVsmx4pc" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0],list):continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="JI_1DzGrySM7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="2470e282-929f-4ecf-b2c9-c2886f22ee24" executionInfo={"status": "error", "timestamp": 1583498301345, "user_tz": -60, "elapsed": 635, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x:-1 if str(x) == 'None' else int(x))
df['param_moc']= df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split('cm')[0].replace(' ', '')))
# + id="-RZPyn3u0Prz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="bee58d94-9481-4cd9-b9e1-c45e3bcbd30a" executionInfo={"status": "error", "timestamp": 1583498295342, "user_tz": -60, "elapsed": 685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
df['param_rok-produkcji']=df['param_rok-produkcji'].map(lambda x: -1 if str(x)=='None' else int(x))
df['param_moc']=df['param_moc'].map(lambda x: -1 if str(x)=='None' else int(x.split(' ')[0]))
df['param_pojemność-skokowa']=df['param_pojemność-skokowa'].map(lambda x: -1 if str(x)=='None' else int(x.split('cm')[0].replace(' ','')))
# + id="OCGTCjDM1Z-o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="b70d85ed-861e-4533-c039-177768435e6a" executionInfo={"status": "error", "timestamp": 1583497980840, "user_tz": -60, "elapsed": 720, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
df['param_moc']= df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x:-1 if str(x) == 'None' else int(x))
feats=['param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
# + id="SabCTN5I29C1" colab_type="code" colab={}
def run_model(model,feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="Ip5wx0EL3FVY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="f41c2250-f4e8-4463-ebfe-ae2cd38984d0" executionInfo={"status": "ok", "timestamp": 1583498494685, "user_tz": -60, "elapsed": 13315, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
feats=['param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
xgb_params = {
'max_depth': 5,
'n_estimators': 50,
'learning_rate': 0.1,
'seed': 0
}
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="P3aKy4yu3aAU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 921} outputId="ae2d52d0-e5c0-4a09-d30a-7a20c1f0e8a7" executionInfo={"status": "ok", "timestamp": 1583500700252, "user_tz": -60, "elapsed": 1400991, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhuWnWPrjl7WYXKOwdYHcyYxZBDLVRLuSUioCyfFQ=s64", "userId": "16732949555547479927"}}
def obj_func(params):
print("Training with params: ")
print(params)
mean_mae, score_std=run_model(xgb.XGBRegressor(**params),feats)
return {'loss':np.abs(mean_mae), 'status':STATUS_OK}
#space
xgb_reg_params = {
'learning_rate': hp.choice("learning_rate", np.arange(0.05, 0.31, 0.05)),
'max_depth': hp.choice("max_depth", np.arange(5,16,1,dtype=int)),
'subsample': hp.quniform('subsample', 0.5,1,0.05),
'colsample_bytree': hp.quniform("colsample_bytree", 0.5,1,0.05),
'objective': 'reg:squarederror',
'n_estimators': 100,
'seed': 0,
}
#chleb
#run
best=fmin(obj_func,xgb_reg_params,algo=tpe.suggest,max_evals=25)
best
| day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={}
# # CLT vs PJ
# + pycharm={"is_executing": false}
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] pycharm={}
# ## Cálculos CLT
# + [markdown] pycharm={}
# ### Cálculos Mensais
# + pycharm={"is_executing": false, "metadata": false, "name": "#%%\n"}
salario_mensal_bruto=[1_903.98, 2_826.66, 3_115.0, 4_664.68, 6_400, 9_000, 9_200, 15_000, 25_000, 30_000, 12_000 ]
# + pycharm={"is_executing": false}
def mostrar_descontos_mensais(funcao):
"""
Formata saída para mostrar descontos mensais em formato de tabela
"""
results = [(f'{salario:,.2f}', f'{funcao(salario):,.2f}', f'{funcao(salario) / salario:.2%}') for salario in salario_mensal_bruto]
df = pd.DataFrame(results, columns=["Salário","Imposto Descontado","Porcentagem"])
return df
# + pycharm={"is_executing": false}
def imposto_de_renda_mensal(salario_mensal:float) -> float:
"""
Calcula imposto de renda PF de acordo com faixa do ano de 2019
https://impostoderenda2019.net.br/tabela-imposto-de-renda-2019/
"""
saldo = salario_mensal
aliquotas_faixas = {0.275: 4_664.68, 0.225: 3_751.06, 0.15: 2_826.66, 0.075: 1_903.98, 0: 0}
ir = 0
desconto_simplificado = 0.20 # 20% de desconto simplificado
for aliquota, faixa in aliquotas_faixas.items():
delta = max(saldo - faixa, 0)
ir += delta * aliquota * (1 - desconto_simplificado)
saldo = min(saldo, faixa)
return ir
# + [markdown] pycharm={}
# Para o cálculo do imposto de renda, foi usada a [tabela desse site](https://impostoderenda2019.net.br/tabela-imposto-de-renda-2019/), que está reproduzida abaixo:
#
# | Base de Cálculo | Alíquota |
# | --------------- | -------- |
# |`De R$1.903,99 até R$2.826,65` | 7,5% |
# |`De R$2.826,66 até R$3.751,05` | 15% |
# |`De R$3.751,06 até R$4.664,68` | 22,5% |
# |`Acima de R$4.664,68` | 27,5% |
# + [markdown] pycharm={}
# Em seguida exibimos o salário, quanto foi descontado e o percentual real retido pelo imposto considerando o desconto simplificado.
# + pycharm={"is_executing": false}
mostrar_descontos_mensais(imposto_de_renda_mensal)
# + pycharm={"is_executing": false}
def fgts_mensal(salario_mensal: float)-> float:
"""
Calcula fgts a ser pago de imposto pelo empregador em 1 ano.
Aliquota de 8% + 40% disso em caso de demissão com justa causa -> 11,2%
:param salario_mensal: salário mensal bruto do empregado
:return: float
"""
return salario_mensal * 0.112
# + pycharm={"is_executing": false}
mostrar_descontos_mensais(fgts_mensal)
# + pycharm={"is_executing": false}
def inss_empregado(salario_mensal):
"""
Calcula inss para 2019
:param salario_mensal:
:return:
"""
if salario_mensal < 1_751.81:
return salario_mensal * 0.08
elif salario_mensal < 2_919.72:
return salario_mensal * 0.09
teto = 5_839.45
return min(salario_mensal, teto) * 0.11
# + pycharm={"is_executing": false}
mostrar_descontos_mensais(inss_empregado)
# + [markdown] pycharm={}
# ### Cálculos Anuais
# + pycharm={"is_executing": false}
def imposto_de_renda_anual(salario):
"""
Calcula imposto de renda anual considerando Décimo terceiro e terço de ferias
:param salario:
:return:
"""
ir = imposto_de_renda_mensal(salario) * 11
salario_de_ferias = salario + salario / 3
ir += imposto_de_renda_mensal(salario_de_ferias - inss_empregado(salario_de_ferias)) / 0.80
ir += imposto_de_renda_mensal(salario - inss_empregado(salario)) / 0.80 # Tributação exlusiva de décimo terceiro
return ir
# + pycharm={"is_executing": false}
def mostrar_descontos_anuais(funcao):
salarios = salario_mensal_bruto
salarios_anuais = [s * 13 + s / 3 for s in salario_mensal_bruto]
results = [
(f'{salario:,.2f}', f'{salario_anual:,.2f}', f'{funcao(salario):,.2f}', f'{funcao(salario) / salario_anual:.2%}') for
salario, salario_anual in zip(salarios, salarios_anuais)]
df = pd.DataFrame(results, columns=["Salário Mensal", "Salário Anual", "Imposto Descontado", "Porcentagem"])
return df
# + pycharm={"is_executing": false}
mostrar_descontos_anuais(imposto_de_renda_anual)
# + pycharm={"is_executing": false}
def inss_anual(salario):
"""
Calcula valor total decontado do empregado anualmente
:param salario:
:return:
"""
ONZE_MESES_MAIS_DECIMO_TERCEIRO = 12
inss = inss_empregado(salario) * ONZE_MESES_MAIS_DECIMO_TERCEIRO
salario_ferias = salario + salario / 3
return inss + inss_empregado(salario_ferias)
# + pycharm={"is_executing": false}
mostrar_descontos_anuais(inss_anual)
# + pycharm={"is_executing": false}
def impostos_anuais(salario):
return imposto_de_renda_anual(salario)+inss_empregado(salario)
# + pycharm={"is_executing": false}
def resumo_empregado(salario):
"""
Retorna tupla com (Salario Mensal, Salario Mensal liquido,Salario Anual Liquido, Salario Liquido +fgts,Salario Bruto Anual , Impostos, FGTS,
:param funcao:
:return:
"""
salario_anual_bruto = salario * 13 + salario / 3
impostos = impostos_anuais(salario)
fgts_anual = fgts_mensal(salario)
fgts_anual = fgts_anual * 13 + fgts_anual / 3
salario_mensal_liquido = salario - imposto_de_renda_mensal(salario) - fgts_mensal(salario) - inss_empregado(salario)
salario_anual_menos_impostos = salario_anual_bruto - impostos
return salario, salario_mensal_liquido, salario_anual_menos_impostos, salario_anual_menos_impostos + fgts_anual, salario_anual_bruto, impostos, fgts_anual,
# + pycharm={}
result = [map(lambda v: f'{v:<15,.2f}', resumo_empregado(salario)) for salario in salario_mensal_bruto]
df = pd.DataFrame(result, columns=["Salário Mensal", "Salário Mensal Líquido","Salário Anual Líquido","Salário Liquido +FGTS", "Salário Bruto Anual", "Imposto Descontado", "FGTS"])
display(df)
# + pycharm={"is_executing": false}
faturamento_bruto = [(s+fgts_mensal(s)) * (13 + 1 / 3) for s in salario_mensal_bruto]
# + pycharm={"is_executing": false}
display(pd.DataFrame([f'{v:,.2f}' for v in faturamento_bruto], columns=["Faturamento Bruto"]))
# + [markdown] pycharm={}
# # PJ SIMPLES ANEXO III
# + pycharm={"is_executing": false}
def custos_simples_anexo_iii(faturamento_anual, contador=89, prolabore=1_100) -> float:
"""
Calcula custos de impostos para empresa simples de anexo III
DAS, Prólabore e Contador
Aliquotas 2019: https://www.contabilizei.com.br/contabilidade-online/anexo-3-simples-nacional/
:param faturamento_anual:
:param contador:
:param prolabore:
:return:
"""
saldo = faturamento_anual
aliquotas_faixas = {0.33: 3_600_000, 0.21: 1_800_000, 0.16: 720_000, 0.135: 360_000, 0.112: 180_000, 0.06: 0}
imposto = 0
for aliquota, faixa in aliquotas_faixas.items():
delta = max(saldo - faixa, 0)
imposto += delta * aliquota
saldo = min(saldo, faixa)
inss = prolabore * 0.11
return imposto + 12 * (contador + inss)
# + [markdown] pycharm={}
# Para o cálculo de impostos PJ foi utilizada a tabela [desse site](https://www.contabilizei.com.br/contabilidade-online/anexo-3-simples-nacional/) reproduzida abaixo
#
# | Faixa | Alíquota| Valor a Deduzir | Receita Bruta em 12 Meses |
# | ------- | ------- | ----------------- | --------------------------------- |
# |1a Faixa | 6,00% | - | Até 180.000,00 |
# |2a Faixa | 11,20% | 9.360,00 | De 180.000,01 a 360.000,00 |
# |3a Faixa | 13,50% | 17.640,00 | De 360.000,01 a 720.000,00 |
# |4a Faixa | 16,00% | 35.640,00 | De 720.000,01 a 1.800.000,00 |
# |5a Faixa | 21,00% | 125.640,00 | De 1.800.000,01 a 3.600.000,00 |
# |6a Faixa | 33,00% | 648.000,00 | De 3.600.000,01 a 4.800.000,00 |
# + pycharm={"is_executing": false}
def resumo_pj(faturamento):
custo_anual = custos_simples_anexo_iii(faturamento)
return faturamento / 12, (faturamento - custo_anual) / 12, faturamento - custo_anual, faturamento
# + pycharm={"is_executing": false}
results = [map(lambda v: f'{v:<15,.2f}', resumo_pj(faturamento)) for faturamento in faturamento_bruto]
display(pd.DataFrame(results, columns=["Faturamento Mensal", "<NAME>", "<NAME>", "Faturamento Anual"]))
# + [markdown] pycharm={}
# # Resultado
# + pycharm={"is_executing": false}
data = []
for salario, faturamento in zip(salario_mensal_bruto, faturamento_bruto):
result = []
result.append(f'{salario:,.2f}')
clt = resumo_empregado(salario)[2]
result.append(f'{clt :<15,.2f}')
pj = resumo_pj(faturamento)[2]
result.append(f'{pj :<15,.2f}')
result.append(f'{pj - clt :<15,.2f}')
data.append(result)
df = pd.DataFrame(data, columns=["<NAME>", "CLT", "PJ", "DIFF"])
display(df)
# + pycharm={}
data = []
index = []
for salario, faturamento in zip(salario_mensal_bruto, faturamento_bruto):
result = []
index.append(f'R$ {salario:,.2f}')
clt = resumo_empregado(salario)[2]
result.append(clt)
pj = resumo_pj(faturamento)[2]
result.append(pj)
result.append(pj - clt)
data.append(result)
df = pd.DataFrame(data, columns=[ "CLT", "PJ", "DIFF"], index=index)
# + pycharm={}
ax = df.plot.bar(figsize=(9.5,5));
vals = ax.get_yticks();
ax.set_yticklabels([f'R${x:,.2f}' for x in vals]);
ax.set_ylabel("<NAME>");
ax.set_xlabel("Salário Mensal");
# + pycharm={}
# + pycharm={"is_executing": false, "metadata": false, "name": "#%%\n"}
def custos_simples_anexo_v(faturamento_anual, contador=89) -> float:
"""
Calcula custos de impostos para empresa simples de anexo V
manipulando pro labore para 28% DAS, Prólabore e Contador
Aliquotas 2019:
:param faturamento_anual:
:param contador:
:param prolabore:
:return: imposto
"""
saldo = faturamento_anual
aliquotas_faixas = {0.33: 3_600_000, 0.21: 1_800_000, 0.16: 720_000, 0.135: 360_000, 0.112: 180_000, 0.06: 0}
imposto = 0
for aliquota, faixa in aliquotas_faixas.items():
delta = max(saldo - faixa, 0)
imposto += delta * aliquota
saldo = min(saldo, faixa)
prolabore= 0.28 * faturamento_anual/12
prolabore = max(prolabore, 1000) # prolabore nao pode ser menor que salario minimo
inss = prolabore * 0.11
ir = imposto_de_renda_mensal(prolabore)
return imposto + 12 * (contador + inss + ir)
# + pycharm={"metadata": false, "name": "#%%\n"}
def resumo_pj_v(faturamento):
custo_anual = custos_simples_anexo_v(faturamento)
return faturamento / 12, (faturamento - custo_anual) / 12, faturamento - custo_anual, faturamento
# + pycharm={"is_executing": false, "metadata": false, "name": "#%%\n"}
results = [map(lambda v: f'{v:<15,.2f}', resumo_pj_v(faturamento)) for faturamento in faturamento_bruto]
display(pd.DataFrame(results, columns=["Faturamento Mensal", "<NAME>", "<NAME>", "Faturamento Anual"]))
# + pycharm={}
data = []
for salario, faturamento in zip(salario_mensal_bruto, faturamento_bruto):
result = []
result.append(f'{salario:,.2f}')
clt = resumo_empregado(salario)[2]
result.append(f'{clt :<15,.2f}')
pj = resumo_pj(faturamento)[2]
result.append(f'{pj :<15,.2f}')
result.append(f'{pj - clt :<15,.2f}')
pj_v = resumo_pj_v(faturamento)[2]
result.append(f'{pj_v :<15,.2f}')
result.append(f'{pj_v - clt :<15,.2f}')
data.append(result)
df = pd.DataFrame(data, columns=["Salário Mensal", "CLT", "PJ - III", "DIFF - III", "PJ - V", "DIFF - V"])
display(df)
# + pycharm={}
# + pycharm={}
| clt_vs_pj.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from IPython.core.display import HTML
with open ('style.css', 'r') as file:
css = file.read()
HTML(css)
# # How to Check that a Formula is a Tautology
# In this notebook we develop a function <tt>tautology</tt> that takes a formula $f$ from propositional logic and checks whether $f$ is a tautology. As we represent tautologies as nested tuples, we first have to import the parser for propositional logic.
import propLogParser as plp
# As we represent propositional valuations as sets of variables, we need a function to compute all subsets of a given set. The module <tt>power</tt> provides a function called <tt>allSubsets</tt> such that for a given set $M$ the function call $\texttt{allSubsets}(M)$ computes a list containing all subsets of $M$, that is we have:
# $$ \texttt{allSubsets}(M) = \bigl[A \mid A \in 2^M\bigr] $$
import power
power.allSubsets({'p', 'q'})
# To be able to compute all propositional valuations for a given formula $f$ we first need to determine the set of all variables that occur in $f$. The function $\texttt{collectVars}(f)$ takes a formula $f$ from propositional logic and computes all propositional variables occurring in $f$. This function is defined recursively.
# + code_folding=[]
def collectVars(f):
"Collect all propositional variables occurring in the formula f."
if f[0] in ['⊤', '⊥']:
return set()
if isinstance(f, str): # at this point, f has to be a propositional variable
return { f }
if f[0] == '¬':
return collectVars(f[1])
# In all other cases, f is build using a binary logical connective.
return collectVars(f[1]) | collectVars(f[2])
# -
# We have discussed the function <tt>evaluate</tt> previously. The call
# $\texttt{evaluate}(f, I)$ takes a propsitional formula $f$ and a propositional valuation $I$, where $I$ is represented as a set of propositional variables. It evaluates $f$ given $I$.
def evaluate(f, I):
"""
Evaluate the propositional formula f using the propositional valuation I.
I is represented as a set of variables.
"""
if isinstance(f, str):
return f in I
if f[0] == '⊤': return True
if f[0] == '⊥': return False
if f[0] == '¬': return not evaluate(f[1], I)
if f[0] == '∧': return evaluate(f[1], I) and evaluate(f[2], I)
if f[0] == '∨': return evaluate(f[1], I) or evaluate(f[2], I)
if f[0] == '→': return not evaluate(f[1], I) or evaluate(f[2], I)
if f[0] == '↔': return evaluate(f[1], I) == evaluate(f[2], I)
# Now we are ready to define the function $\texttt{tautology}(f)$ that takes a propositional formula $f$ and checks whether $f$ is a tautology. If $f$ is a tautology, the function returns <tt>True</tt>, otherwise a set of variables $I$ is returned such that $f$ evaluates to <tt>False</tt> if all variables in $I$ are <tt>True</tt>, while all variables not in $I$ are <tt>False</tt>.
def tautology(f):
"Check, whether the formula f is a tautology."
P = collectVars(f)
A = power.allSubsets(P)
if all(evaluate(f, I) for I in A):
return True
else:
return [I for I in A if not evaluate(f, I)][0]
# The function $\texttt{test}(s)$ takes a string $s$ that can be parsed as a propositionl formula and checks whether this formula is a tautology.
def test(s):
f = plp.LogicParser(s).parse()
counterExample = tautology(f);
if counterExample == True:
print('The formula', s, 'is a tautology.')
else:
P = collectVars(f)
print('The formula ', s, ' is not a tautology.')
print('Counter example: ')
for x in P:
if x in counterExample:
print(x, "↦ True")
else:
print(x, "↦ False")
# Let us run a few tests.
# The first example is DeMorgan's rule.
test('¬(p ∨ q) ↔ ¬p ∧ ¬q')
test('(p → q) → (¬p → q) → q')
test('(p → q) → (¬p → ¬q)')
test('¬p ↔ (p → ⊥)')
| Python/Tautology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image classification with CNN
# In this lab we will teach computer to distinguish between images of cats and dogs using Convolutional Neural Network.
# The input dataset consists of 10,000 images manually labeled as ''cats'' and ''dogs''. The original dataset was downloaded from kaggle.
# ## 1. Building the model
# ### 1.1. Import all the required libraries.
# +
import numpy as np
import matplotlib.pyplot as plt
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
from PIL import Image
from keras import preprocessing
import sys
# -
# ### 1.2. Load images
# +
import os
cwd = os.getcwd()
os.chdir(cwd)
data_dir = "../data_set/cat_dog_data"
print(os.listdir(data_dir))
# +
train_cats_files = []
train_path_cats = data_dir +"/training_set/cats/"
for path in os.listdir(train_path_cats):
if '.jpg' in path:
train_cats_files.append(os.path.join(train_path_cats, path))
train_dogs_files = []
train_path_dogs = data_dir +"/training_set/dogs/"
for path in os.listdir(train_path_dogs):
if '.jpg' in path:
train_dogs_files.append(os.path.join(train_path_dogs, path))
print(len(train_cats_files), len(train_dogs_files))
test_cats_files = []
test_path_cats = data_dir +"/test_set/cats/"
for path in os.listdir(test_path_cats):
if ".jpg" in path:
test_cats_files.append(os.path.join(test_path_cats,path))
test_dogs_files = []
test_path_dogs = data_dir +"/test_set/dogs/"
for path in os.listdir(test_path_dogs):
if '.jpg' in path:
test_dogs_files.append(os.path.join(test_path_dogs, path))
print(len(test_cats_files), len(test_dogs_files))
# -
k = 200
sample_dog_file = train_dogs_files[k]
img = preprocessing.image.load_img(sample_dog_file, target_size=(64, 64))
img_array = preprocessing.image.img_to_array(img)
plt.imshow(np.uint8(img_array))
# ### 1.3. Images to numpy arrays
# Now we create training and testing sets for cats and for dogs and then concatenate them into a single `X_train`, `X_test` dataset of features.
# +
d = 64
X_train_orig = np.zeros((8000, d, d, 3), dtype='float32')
for i in range(4000):
path = train_cats_files[i]
img = preprocessing.image.load_img(path, target_size=(d, d))
X_train_orig[i] = preprocessing.image.img_to_array(img)
for i in range(4000,8000):
path = train_dogs_files[i-4000]
img = preprocessing.image.load_img(path, target_size=(d, d))
X_train_orig[i] = preprocessing.image.img_to_array(img)
X_test_orig = np.zeros((2000, d, d, 3), dtype='float32')
for i in range(1000):
img = preprocessing.image.load_img(test_cats_files[i], target_size=(d, d))
X_test_orig[i] = preprocessing.image.img_to_array(img)
for i in range(1000, 2000):
img = preprocessing.image.load_img(test_dogs_files[i-1000], target_size=(d, d))
X_test_orig[i] = preprocessing.image.img_to_array(img)
print(X_train_orig.shape)
print(X_test_orig.shape)
# -
# ### 1.4. Normalize image array
X_train = X_train_orig / 255.0
X_test = X_test_orig / 255.0
print(X_train.shape)
print(X_test.shape)
# ### 1.5. Create class labels
# Now we need to create the corresponding class label vectors. We will mark the cats as class 1, and the dogs as class 0 (not cats).
# +
from keras.utils.np_utils import to_categorical
Y_train_orig = np.ones((4000,))
Y_train_orig = np.concatenate((Y_train_orig, np.zeros((4000,))))
Y_train = Y_train_orig.reshape(-1)
print("At position 3 should be a cat:", Y_train[3])
print("At position 4002 should be a dog:", Y_train[4002])
Y_train = to_categorical(Y_train, num_classes = 2)
print(Y_train.shape)
Y_test_orig = np.ones((1000,))
Y_test_orig = np.concatenate((Y_test_orig, np.zeros((1000,))))
Y_test = Y_test_orig.reshape(-1)
Y_test = to_categorical(Y_test, num_classes = 2)
print(Y_test.shape)
# -
# ### 1.6 Build CNN Model
# model structure: conv => max pool => dropout => conv => max pool => dropout => conv => max pool => dropout => fully connected (3 layer)
#
# +
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop,Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
model = Sequential()
#
model.add(Conv2D(filters = 8, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (64,64,3)))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
#
model.add(Conv2D(filters = 16, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
#
model.add(Conv2D(filters = 32, kernel_size = (2,2),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
# fully connected
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(2, activation = "softmax"))
# -
# Define the optimizer
optimizer = Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
# +
# Fit the model
epochs = 15
batch_size = 150
history = model.fit(X_train,Y_train, epochs = epochs, validation_data = (X_test,Y_test), steps_per_epoch=X_train.shape[0] // batch_size)
# -
# ### 1.7. Evaluate the model
# +
# visualize the loss function
plt.plot(history.history['loss'], color='r', label="training loss")
plt.plot(history.history['val_loss'], color='b', label="test loss")
plt.title("Loss")
plt.xlabel("Number of Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
# visualize the accuracy function
plt.plot(history.history['accuracy'], color='r', label="training accuracy")
plt.plot(history.history['val_accuracy'], color='b', label="test accuracy")
plt.title("Accuracy")
plt.xlabel("Number of Epochs")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
# -
# We can find out that after 10 epochs there are signs for over-fitting because the training accuracy is increasing but the test accuracy is dropping significantly. That's to say, the best test accuracy we can achieve is 79.85% with 10 epochs. After manipulating all parameters, adding more layers(which decreases performance over 3 layers), changing batch size(which decreases performance over 150), changing models(add batch normalization which doesn't improve performance), changing learning rate(which decreases performance when we change it in either direction), a lucky choice of initial weight(start with 76% test accuracy), we achieve around 80% accuracy.
# +
# compute the confusion matrix
import seaborn as sns
Y_pred = model.predict(X_test)
Y_pred_classes = np.argmax(Y_pred,axis = 1)
Y_true = np.argmax(Y_test,axis = 1)
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
f,ax = plt.subplots(figsize=(8, 8))
sns.heatmap(confusion_mtx, annot=True, linewidths=0.01,cmap="Greens",linecolor="gray", fmt= '.1f',ax=ax)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plt.title("Confusion Matrix")
plt.show()
# -
# ### 1.8 Predict the new image
# +
# predict random image of cat and dog
d = 64
test_dir = "./"
test_path_files = []
for path in os.listdir(test_dir):
if ".jpg" in path:
test_path_files.append(os.path.join(test_dir,path))
img_arr = np.zeros((2, d, d, 3), dtype='float32')
for i in range(2):
img = preprocessing.image.load_img(test_path_files[i], target_size=(d, d))
img_arr[i] = preprocessing.image.img_to_array(img)
X_new = img_arr / 255
Y_new = model.predict(X_new)
print(Y_new)
Y_new_classes = np.argmax(Y_new,axis = 1)
display = ['dog','cat']
# %matplotlib inline
plt.figure()
f, axarr = plt.subplots(1,2)
for i in range(2):
axarr[i].imshow(np.uint8(img_arr[i]))
axarr[i].set_title("predicted as "+display[int(Y_new_classes[i])])
| cats_lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../')
import controller
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# +
np.random.seed(149)
ci = controller.Car_Interface()
ci.set_gear(ci.FORWARD)
def apply_alternating_controls(ci, num, time_per):
commands = []
cmdtype = [ci.BRAKE, ci.ACCELERATOR]
cmd_name = ['brake', 'accelerate']
'''
commands should be a list of tuples (cmd_type, amt) where
cmd_type is either 'accelerate' or 'brake' and amt is a
random number between 0 and 1 that denotes how much the
corresponding pedal was depressed.
In total there should be num commands and each should
be executed for time_per seconds. The car interface log
will be used later for accessing the data, so make sure
to actually execute the commands on the vehicle.
(see controller.py)
'''
#CODE HERE
for i in range(num):
cmd = np.random.randint(0,2)
amt = np.random.random()
commands.append((cmd_name[cmd], amt))
ci.apply_control_for_time(cmdtype[cmd], amt, time_per)
ci.log_data()
return commands
N = 20
TIME_PER = 50
commands = apply_alternating_controls(ci, num = N, time_per = TIME_PER)
for i in range(10):
print(commands[i][0], f"{commands[i][1] * 100:.2f}%")
# +
vel_log = ci.log["velocity"]
acc_log = ci.log["acceleration"]
time_log = ci.log["time"]
NUM_READINGS = len(time_log)
NUM_READINGS_PER = int(NUM_READINGS / N)
filt_times = []
filt_inps = []
filt_outs = []
all_times = []
all_inps = []
all_outs = []
for i in range(NUM_READINGS):
cmd = commands[int(i / NUM_READINGS_PER)]
vel = vel_log[i]
acc = acc_log[i]
t = time_log[i]
inp = [0,0,0]
if (cmd[0] == 'accelerate'):
'''
inp = ? should be a list of length 3 corresponding
to the three input variable in the linear equation
for determining acceleration
'''
inp = [cmd[1], 0, vel]
elif(cmd[0] == 'brake'):
'''
inp = ? remember to order input variables exactly
as done in the previous statement
'''
inp = [0, cmd[1], vel]
all_inps.append(inp)
all_outs.append(acc)
all_times.append(t)
'''
At first you can leave this code section blank.
Once you run Linear Regression on the basic inputs
and outputs you will notice the regression struggles to
accurately model data. Here you will populate the filtered
lists (filt_inps, filt_outs, filt_times) similar to how
the all lists are populate above, but censoring out certain
(input, output) data points that are undesirable.
'''
#CODE HERE
if inp != [0,0,0] :
filt_inps.append(inp)
plt.title("Collected Data")
plt.xlabel("Time")
plt.ylabel("Acceleration")
plt.scatter(all_times, all_outs, color = 'b')
plt.show()
print(f"Actual Coefficients:")
'''
Fill in the blanks in the next 4 lines with
reasonable names for the 4 coefficients and
in the curly braces put the corresponding
variable name to display it. The variables
should attributes of the car interface object
that correspond to those in the linear
equation presented in class.
'''
print(f"rolling bias: {ci.rolling_bias}")
print(f"friction: {ci.friction_constant}")
print(f"accel_weight: {ci.accelerator_weight}")
print(f"brake_weight: {ci.brake_weight}")
print("___________________________________________")
# +
model = LinearRegression()
'''
First fit the model with the generated data.
Then assign fit_score to sklearn LinearRegression
score function. Finally retrieve the learned coefficients.
There should be 4, 3 linear dependencies and one constant.
(See the coef_ and intercept_ attributes in the LinearRegression
documentation)
'''
#CODE HERE
model.fit(all_inps, all_outs)
fit_score = model.score(all_inps, all_outs)
coef = model.coef_
const = model.intercept_
print(str(coef) + " | " + str(const))
print(f"Model Fit {fit_score}")
print(f"Model Predictions:")
'''
Fill in the blanks in the next 4 lines with
reasonable names for the 4 coefficients and
in the curly braces put the corresponding
variable name to display it
'''
print(f": {_}")
print(f"_______: {_}")
print(f"_______: {_}")
print(f"_______: {_}")
print("___________________________________________")
plt.clf()
plt.title("Model Predictions on all Data")
plt.xlabel("Time")
plt.ylabel("Acceleration")
plt.scatter(all_times, all_outs, color = 'b')
plt.scatter(all_times, model.predict(all_inps), color = 'r')
plt.show()
# +
'''
First fit the model with the filtered generated data.
Then assign fit_score to sklearn LinearRegression
score function. Finally retrieve the learned coefficients.
There should be 4, 3 linear dependencies and one constant.
Code should be similar to the previous section but now using
the filtered data.
'''
print(f"Model Fit {fit_score}")
'''
Fill in the blanks as in the previous section
'''
print(f"_______: {_}")
print(f"_______: {_}")
print(f"_______: {_}")
print(f"_______: {_}")
print("___________________________________________")
plt.clf()
plt.title("Filtered Data")
plt.scatter(filt_times, filt_outs, color = 'b')
plt.xlabel("Time")
plt.ylabel("Acceleration")
plt.show()
plt.clf()
plt.title("Model Predictions on Filtered Data")
plt.xlabel("Time")
plt.ylabel("Acceleration")
plt.scatter(filt_times, filt_outs, color = 'b')
plt.scatter(filt_times, model.predict(filt_inps), color = 'r')
plt.show()
# -
| Car Interface Weeks 2-3/SystemID/.ipynb_checkpoints/hw2_system_id-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#importing necessary packages
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
import numpy as np
data = pd.read_csv('Nursery/nursery.data',header=None,skipinitialspace=True)
data.shape
data.info()
data.columns = ['parents','has_nurs','form','children','housing','finance','social','health','recomendation']
data.head()
data.children.replace("1","one",inplace=True)
data.children.replace("2","two",inplace=True)
data.children.replace("3","three",inplace=True)
from sklearn.preprocessing import LabelEncoder
enc = LabelEncoder()
data.parents = enc.fit_transform(data.parents)
data.has_nurs = enc.fit_transform(data.has_nurs)
data.form = enc.fit_transform(data.form)
data.children = enc.fit_transform(data.children)
data.housing = enc.fit_transform(data.housing)
data.finance = enc.fit_transform(data.finance)
data.social = enc.fit_transform(data.social)
data.health = enc.fit_transform(data.health)
data.recomendation = enc.fit_transform(data.recomendation)
X = data.iloc[:,:-1]
y = data.recomendation
from sklearn.model_selection import train_test_split
X_train, X_test,y_train, y_test = train_test_split(X,y,test_size=0.3,
random_state=10)
#Defining and fitting
model = DecisionTreeClassifier(max_depth=6,criterion='gini',random_state=15,max_features=5)
model.fit(X_train,y_train)
#Visualizing
dot_data = StringIO()
export_graphviz(model, out_file=dot_data,
filled=True,
feature_names = X_train.columns,
class_names=['not_recom','priority','recomended','spec_prior','very_recom'],
rounded=True,
special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
from collections import Counter
Counter(data.loc[data.recomendation<5.0,'recomendation'])
y_predict = model.predict(X_test)
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, y_predict))
pd.crosstab(y_test, y_predict)
# +
# 1 - ((50/104)**2+(48/104)**2+(6/104)**2)
# +
# 1 - ((10/90)**2+(40/90)**2+(40/90)**2)
# +
# 1 - ((50/70)**2+(10/70)**2+(10/70)**2)
# -
| NurseryDataDicisionTreeClassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <!--HEADER-->
# [*NBBinder test on a collection of notebooks about some thermodynamic properperties of water*](https://github.com/rmsrosa/nbbinder)
# + [markdown] slideshow={"slide_type": "skip"}
# <!--BADGES-->
# <a href="https://colab.research.google.com/github/rmsrosa/nbbinder/blob/master/tests/nb_builds/nb_water/03.00-Low_Dim_Fittings.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Google Colab" title="Open in Google Colab"></a><a href="https://mybinder.org/v2/gh/rmsrosa/nbbinder/master?filepath=tests/nb_builds/nb_water/03.00-Low_Dim_Fittings.ipynb"><img align="left" src="https://mybinder.org/badge.svg" alt="Open in binder" title="Open in binder"></a><a href="https://nbviewer.jupyter.org/github/rmsrosa/nbbinder/blob/master/tests/nb_builds/nb_water/03.00-Low_Dim_Fittings.ipynb"><img align="left" src="https://img.shields.io/badge/view%20in-nbviewer-orange" alt="View in NBViewer" title="View in NBViewer"></a><a href="https://nbviewer.jupyter.org/github/rmsrosa/nbbinder/blob/master/tests/nb_builds/nb_water_slides/03.00-Low_Dim_Fittings.slides.html"><img align="left" src="https://img.shields.io/badge/view-slides-darkgreen" alt="View Slides" title="View Slides"></a>
# + [markdown] slideshow={"slide_type": "skip"}
# <!--NAVIGATOR-->
# [<- Reading the Data](02.00-Data.ipynb) | [Water Contents](00.00-Water_Contents.ipynb) | [References](BA.00-References.ipynb) | [High-Dimensional Fittings ->](04.00-High_Dim_Fittings.ipynb)
#
# ---
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Low-Dimensional Fittings
#
# We use the classical least-square method to fit low degree polynomials to the data.
#
# See e.g. [<NAME> (1996)](BA.00-References.ipynb) and [<NAME> (1997)](BA.00-References.ipynb) for details on the least-square method.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Importing the libraries
# + slideshow={"slide_type": "fragment"}
import csv
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "slide"}
# ## Loading the data
#
# This time we use the `csv` library to read the data from file and use `numpy` to work with the data.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Loading with csv
#
# We first load the data into a python list with `csv`.
# + slideshow={"slide_type": "fragment"}
file = open('water.csv',"r")
water_csv = list(csv.reader(file, delimiter=","))
water_csv
# + [markdown] slideshow={"slide_type": "slide"}
# ### Header and datapoints
#
# We separate the header from the datapoints and have the datapoints put into a `numpy` array as follows.
# + slideshow={"slide_type": "fragment"}
header = dict([(water_csv[0][i],water_csv[1][i]) for i in range(3)])
print(header)
# + slideshow={"slide_type": "subslide"}
datapoints = np.array(water_csv[2:]).astype("float")
print(datapoints)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Linear approximation
#
# Given a set of temperature and density data in the form $(T_j, \rho_j)$, we look for a linear relation $\rho^{(1)}(T) = c + m T$ which is the "best fit" for the data.
#
# One way to approach this problem is to interpret the best fit as minimizing the sum of squares of the residuals. The **residual** for each $j$ measurement is
#
# $$
# r_j = \rho_j - \rho^{(1)}(T_j),
# $$
#
# and the **sum of squares of the residuals** is
#
# $$
# \min_{c,m\in \mathbb{R}} \sum_j (\rho_j - \rho^{(1)}(T))^2.
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Matrix form
#
# This can be written in matrix form as
#
# $$
# \displaystyle \min_{\mathbf{u}\in \mathbb{R}^2} \|A\mathbf{u} - \mathbf{f}\|_2^2,
# $$
#
# where $\|\cdot\|_2$ is the Euclidian norm of a vector and
#
# $$ A = \left[ \begin{matrix} T_1 & 1 \\ \vdots & 1 \\ T_n & 1 \end{matrix}\right], \qquad \mathbf{u} = \left( \begin{matrix} m \\ c \end{matrix}\right), \qquad \mathbf{f} = \left( \begin{matrix} \rho_1 \\ \vdots \\ \rho_n \end{matrix} \right).
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# The matrix $A$ is a simple **Vandermonde** type matrix obtained from the temperature data, $\mathbf{u}$ is the unknown vector with the desired coefficients for the approximation, and $\mathbf{f}$ is the vector with the density measurements.
# + [markdown] slideshow={"slide_type": "slide"}
# ### The Vandermonde matrix
#
# We use the [`numpy`](https://docs.scipy.org/doc/numpy/) function [`numpy.vstack()`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.vstack.html) to build the Vandermonde matrix $A$.
# + slideshow={"slide_type": "fragment"}
T = datapoints[:,0]
A1 = np.vstack([T,np.ones(len(T))]).T
print(A1)
# + [markdown] slideshow={"slide_type": "slide"}
# ### The density measurements
#
# The density data is the second column of the data array.
# + slideshow={"slide_type": "fragment"}
f = datapoints[:,1]
print(f)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Solution of the least-square problem for the linear approximation
#
# We use the `numpy` function [numpy.linagl.lstsq()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.lstsq.html) to solve the least-square problem.
# + slideshow={"slide_type": "fragment"}
m, d = np.linalg.lstsq(A1, f, rcond=None)[0]
print(m,d)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Visualizing the result
#
# Now we plot the linear approximation along with the data to visualize the quality of the approximation
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(10,5))
plt.plot(T, f, 'o', label='Data', color='tab:blue')
plt.plot(T, m*T + d, 'b', label='Linear approximation', color='tab:red')
plt.title('Plot of the data and of the linear approximation', fontsize=14)
plt.xlabel(header['temp'], fontsize=12)
plt.ylabel(header['density'], fontsize=12)
plt.legend()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Second-degree approximation
#
# One can see from the plot above that the first-degree approximation doesn't seem to be a very good approximation. We look, now, for a second-degree approximation.
# + [markdown] slideshow={"slide_type": "slide"}
# ### The least-square problem
#
# For the second-degree approximation, we look for a second-degree polynomial $\rho^{(2)}(T) = aT^2 + bT + c$ that best approximates the data in the sense of minimizing the sum of the square of the residuals
#
# $$
# r_j = \rho_j - \rho^{(2)}(T_j)
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Matrix form
#
# This can be written in matrix form as
#
# $$
# \displaystyle \min_{\mathbf{u}\in \mathbb{R}^2} \|A\mathbf{u} - \mathbf{f}\|_2^2,
# $$
#
# where $\mathbf{f}$ is as before but the Vandermonde matrix and the vector of unknowns take the form
#
# $$ A = \left[ \begin{matrix} T_1^2 & T_1 & 1 \\ \vdots & 1 \\ T_n^2 & T_n & 1 \end{matrix}\right], \qquad \mathbf{u} = \left( \begin{matrix} a \\ b \\ c \end{matrix}\right).
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Vandermonde matrix
#
# In this case, the Vandermonde matrix is
# + slideshow={"slide_type": "fragment"}
A2 = np.vstack([T**2, T,np.ones(len(T))]).T
print(A2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Solution
# + slideshow={"slide_type": "fragment"}
a, b, c = np.linalg.lstsq(A2, f, rcond=None)[0]
print(a,b,c)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Visualizing the result
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(10,5))
plt.plot(T, f, 'o', label='Data', color='tab:blue')
plt.plot(T, a*T**2 + b*T + c, 'r', label='Quadratic approximation', color='tab:green')
plt.title('Plot of the data and of the quadratic approximation', fontsize=14)
plt.xlabel(header['temp'], fontsize=12)
plt.ylabel(header['viscosity'], fontsize=12)
plt.legend()
plt.show()
# + [markdown] slideshow={"slide_type": "fragment"}
# This seems much better
# + [markdown] slideshow={"slide_type": "slide"}
# ## Comparing the two approximations
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Visual comparison
#
# Visually, the second-degree approximation is way better.
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(10,5))
plt.plot(T, f, 'o', label='Data', color='tab:blue')
plt.plot(T, m*T + d, 'b', label='Linear approximation', color='tab:red')
plt.plot(T, a*T**2 + b*T + c, 'r', label='Quadratic approximation', color='tab:green')
plt.title('Plot of the data along with the approximations', fontsize=14)
plt.xlabel(header['temp'], fontsize=12)
plt.ylabel(header['viscosity'], fontsize=12)
plt.legend()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Comparing the quadratic error
#
# The **quadratic error** is the sum of the square of the residual errors for each measurement, i.e.
#
# $$
# \Delta = \sum_j r_j^2 = \|A\mathbf{u} - \mathbf{f}\|_2^2,
# $$
#
# for the best approximation obtained. But this error is not normalized by the number of measurements. This is achieved with the **mean quadratic error**:
#
# $$
# E = \frac{1}{N} \sum_j r_j^2.
# $$
# + slideshow={"slide_type": "fragment"}
N = len(f)
print(f'Number of measurements: {N}\n')
print(f'Quadratic error for the linear approximation: {np.linalg.lstsq(A1, f, rcond=None)[1][0]:.2e}')
print(f'Quadratic error for the quadratic approximation: {np.linalg.lstsq(A2, f, rcond=None)[1][0]:.2e}\n')
print(f'Mean quadratic error for the linear approximation: {np.linalg.lstsq(A1, f, rcond=None)[1][0]/N:.2e}')
print(f'Mean quadratic error for the quadratic approximation: {np.linalg.lstsq(A2, f, rcond=None)[1][0]/N:.2e}')
# + [markdown] slideshow={"slide_type": "slide"}
# <!--NAVIGATOR-->
#
# ---
# [<- Reading the Data](02.00-Data.ipynb) | [Water Contents](00.00-Water_Contents.ipynb) | [References](BA.00-References.ipynb) | [High-Dimensional Fittings ->](04.00-High_Dim_Fittings.ipynb)
| tests/nb_builds/nb_water/03.00-Low_Dim_Fittings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mlos_python_environment]
# language: python
# name: conda-env-mlos_python_environment-py
# ---
# # Optimizing Smart Cache with Bayesian Optimization
#
# The goal of this notebook is to optimize SmartCache using Bayesian Optimization approach.
#
# We're using a sequential model-based optimization approach, that consists of the following loop:
# 1. Get suggested config from optimizer,
# 2. Apply suggested config to ``SmartCache``,
# 3. Execute a fixed workload,
# 4. Collect the metrics from ``SmartCache``,
# 5. Register an observation with the optimizer.
#
# +
# import the required classes and tools
import grpc
import pandas as pd
import logging
from mlos.Logger import create_logger
from mlos.Examples.SmartCache import HitRateMonitor, SmartCache, SmartCacheWorkloadGenerator, SmartCacheWorkloadLauncher
from mlos.Mlos.SDK import MlosExperiment
from mlos.Optimizers.BayesianOptimizerFactory import BayesianOptimizerFactory
from mlos.Optimizers.OptimizationProblem import OptimizationProblem, Objective
from mlos.Spaces import Point, SimpleHypergrid, ContinuousDimension
# The optimizer will be in a remote process via grpc, we pick the port here:
grpc_port = 50051
# -
# Launch the optimizer service in a different process:
import subprocess
optimizer_microservice = subprocess.Popen(f"start_optimizer_microservice launch --port {grpc_port}", shell=True)
# Now the optimizer service that runs the surrogate model and suggests new points is started in the background.
# Next, we instantiate an object that connects to it over grpc using the ``BayesianOptimizerFactory``.
logger = create_logger('Optimizing Smart Cache', logging_level=logging.WARN)
optimizer_service_grpc_channel = grpc.insecure_channel(f'localhost:{grpc_port}')
bayesian_optimizer_factory = BayesianOptimizerFactory(grpc_channel=optimizer_service_grpc_channel, logger=logger)
# # The optimization problem
# Then we can instantiate our optimization problem. We want to optimize the configuration of the ``SmartCache`` component that contains two implementations: an LRU (least recently used) cache and an MRU cache (most recently used).
# The ``SmartCache`` component has two parameters that we can adjust, the type of cache and the cache size. We are using some synthetic workloads for the cache and try to find what the optimum configuration for each workload is.
#
# Here, we measure 'optimum' by the number of cache hits. Another option would be to measure runtime; however, this is a toy example with a trivial workload and there is likely substantial runtime difference.
# The parameter search space is declared in ``SmartCache.parameter_search_space``:
SmartCache.parameter_search_space
# The optimization problem is constructed using this parameter space as the input to optimize, and defines a single continuous objective, 'hit_rate' between 0 and 1.
# Optimization Problem
#
optimization_problem = OptimizationProblem(
parameter_space=SmartCache.parameter_search_space,
objective_space=SimpleHypergrid(name="objectives", dimensions=[ContinuousDimension(name="hit_rate", min=0, max=1)]),
objectives=[Objective(name="hit_rate", minimize=False)]
)
# create an optimizer proxy that connects to the remote optimizer via grpc:
# here we could also configure the optimizer
optimizer = bayesian_optimizer_factory.create_remote_optimizer(optimization_problem=optimization_problem)
# # Defining workloads
# Now we can instantiate our workloads and stand up the MLOS infrastructure, both of which are orchestrated by``SmartCacheWorkloadLauncher``. The MLOS infrastructure consists of the MlosAgent and a communication channel, which are available to both the ``SmartCacheWorkloadGenerator`` and the ``SmartCache``.
# The ``SmartCacheWorkloadLauncher`` launches workloads in ``SmartCacheWorkloadGenerator`` in a separate thread, which will actually generate and run the workloads for the smart cache.
# The SmartCacheWorkloadLauncher also connects the ``SmartCacheWorkLloadGenerator`` to the optimization problem via a ``MlosAgent`` that will consume the configurations.
workload_launcher = SmartCacheWorkloadLauncher(logger=logger)
mlos_agent = workload_launcher.mlos_agent
# We set up the agent to consume configurations for the ``SmartCacheWorkloadGenerator``, and we configure the workload to be sequential keys from a range from 0 to 2048.
mlos_agent.set_configuration(
component_type=SmartCacheWorkloadGenerator,
new_config_values=Point(
workload_type='cyclical_key_from_range',
cyclical_key_from_range_config=Point(
min=0,
range_width=2048
)
)
)
# # Launching the experiment (measurement)
# Now we build the experiment, which collects hit-rate statistics from the ``SmartCacheWorkloadGenerator`` via the ``HitRateMonitor``. This architecture reflects the native architecture for the C++ interface in which communication is done via shared memory between MLOS and the worker.
hit_rate_monitor = HitRateMonitor()
smart_cache_experiment = MlosExperiment(
smart_component_types=[SmartCache],
telemetry_aggregators=[hit_rate_monitor]
)
mlos_agent.start_experiment(smart_cache_experiment)
# # Performing the optimization
# Now that we have all the pieces in place, we can iterate our main optimization loop.
# Our workload will run in the same process as this notebook, but in a separate thread, which we block on.
# In a real example, the workload might run completely independent of our optimization procedure.
#
# We run the optimization for 20 iterations, in each of which we obtain a new configuration from the optimizer (that interfaces the remote optimizer service).
# The configuration is passed to ``SmartCacheWorkloadGenerator`` via the ``MlosAgent``, after which we start a blocking workload for 0.2 seconds.
# Then, the hit-rate (our objective) is read from the ``HitRateMonitor`` and the suggested configuration together with the resulting hit-rate are passed to the optimizer.
num_iterations = 100
data = []
for i in range(num_iterations):
# suggest runs a 'cheap' search on the surrogate model to find a good candidate configuration
new_config_values = optimizer.suggest()
# set_configuration communicates the proposed configuration to the SmartCache
mlos_agent.set_configuration(component_type=SmartCache, new_config_values=new_config_values)
hit_rate_monitor.reset()
# start_workload will actually run the worker, here for 0.2 seconds
workload_launcher.start_workload(duration_s=0.2, block=True)
# obtain hit-rate as quality measure for configuration
hit_rate = hit_rate_monitor.get_hit_rate()
objectives_df = pd.DataFrame({'hit_rate': [hit_rate]})
# pass configuration and observed hit-rate to the optimizer to update the surrogate model
features_df = new_config_values.to_dataframe()
optimizer.register(features_df, objectives_df)
print(f"[{i+1}/{num_iterations}] current_config: {new_config_values.to_json()}, hit_rate: {hit_rate:.3f}")
# # Analyzing results
#
# For a cyclical workload with 2048 keys, we assume that a MRU cache with a size of at least 2048 will perform best, and get 100% hits once the cache is filled.
# Now lets see the suggestions and results from the current experiment.
# some pandas wrangling
features, targets = optimizer.get_all_observations()
data = pd.concat([features, targets], axis=1)
data
# +
# group by implementation, then plot
lru_data, mru_data = data.groupby('implementation')
import matplotlib.pyplot as plt
line_lru = lru_data[1].plot(x='lru_cache_config.cache_size', y='hit_rate', label='LRU', marker='o', linestyle='none', alpha=.6)
mru_data[1].plot(x='mru_cache_config.cache_size', y='hit_rate', label='MRU', marker='o', linestyle='none', alpha=.6, ax=plt.gca())
plt.ylabel("Cache hitrate")
plt.xlabel("Cache Size")
plt.legend()
# -
# We can see that if the cache size is over 2048 keys, it means everything can fit into the cache and the strategy does not matter.
# However, for smaller cache sizes, the MRU strategy has an obvious advantage over the LRU strategy.
# # Going Further
#
# 1) Log how the optimum evolves over time. How many iterations are needed?
#
# 2) Can you adjust options in the Optimizer to improve convergence (see the BayesianOptimization notebook for suggestions).
#
# 3) Choose a different workload in the ``SmartCacheWorkloadGenerator``. How do the workloads change the optimum strategy?
# # Clean up
# We need to stop all processes & separate threads after running the experiments:
# +
# Clean up
#
mlos_agent.stop_experiment(smart_cache_experiment)
mlos_agent.stop_all()
# Stop the optimizer service
import signal
optimizer_microservice.send_signal(signal.SIGTERM)
| source/Mlos.Notebooks/SmartCacheOptimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing Libraries
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib notebook
# ## Importing the Dataset
df=pd.read_csv("Desktop/Shops.csv",encoding='latin1')
df
X1=df['X1']
X2=df['X2']
Y1=df['Y1']
Y2=df['Y2']
Name=df['Name']
Type=df['Type']
# +
#Making Diconary of Shops
dict_shops={}
for i in range(df.shape[0]):
dict_shops[Name[i]]=Type[i]
dict_shops["Relax"]="Relax"
# -
# ## Plotting the MAP of the Airport
# +
fig = plt.figure()
ax = fig.add_subplot(111)
for i in range (12):
rect1 = matplotlib.patches.Rectangle((X1[i], Y1[i]),X2[i]-X1[i], Y2[i]-Y1[i], facecolor="grey", edgecolor='black')
ax.add_patch(rect1)
ax.annotate(Name[i], (X1[i]+30, Y1[i]+10),color='w', weight='bold',
fontsize=6, ha='center', va='center')
plt.xlim([50, 600])
plt.ylim([50, 500])
plt.show()
# -
# ## Importing Dataset (Location of the passengers)
pdf=pd.read_csv("Desktop/location.csv",encoding='latin1')
pdf
# ## Fetching the Precise location of the Passenger
# +
k=pdf.shape[0]
lat=pdf['Lat']
long=pdf['Long']
pid=pdf['Pid']
current_location=[]
for i in range(k):
l1=lat[i]
l2=long[i]
flag=0;
for j in range(12):
if(l1>=X1[j] and l1<=X2[j] and l2>=Y1[j] and l2<=Y2[j]):
flag=1
print(Name[j])
current_location.append(Name[j])
break
if(flag==0):
current_location.append("Relax")
print("Relax")
# -
print(current_location)
# ## Declaring the lists
# +
# Generalised Lists
Restaurant=[]
Shopping=[]
Electronics=[]
Cosmetics=[]
Clothing=[]
Relax=[]
# Specific Lists
Mcdonalds=[]
Subway=[]
KFC=[]
Pizza_Hut=[]
Cloth_store=[]
Liquor=[]
Cafe=[]
Haldiram=[]
Samsung=[]
Apple=[]
Lakme=[]
Zara=[]
general_type=[]
# -
for i in range(pdf.shape[0]):
temp=dict_shops[current_location[i]]
print(temp)
if(temp=="Shopping"):
Shopping.append(pid[i])
general_type.append("Shopping")
elif(temp=="Restaurant"):
Restaurant.append(pid[i])
general_type.append("Restaurant")
elif(temp=="Electronics"):
Electronics.append(pid[i])
general_type.append("Electronics")
elif(temp=="Cosmetics"):
Cosmetics.append(pid[i])
general_type.append("Cosmetics")
elif(temp=="Clothing"):
Clothing.append(pid[i])
general_type.append("Clothing")
else:
Relax.append(pid[i])
general_type.append("Relax")
# ## Finding the Percentage
p_Restaurant=(len(Restaurant)/len(current_location))*100
p_Shopping=(len(Shopping)/len(current_location))*100
p_Electronics=(len(Electronics)/len(current_location))*100
p_Cosmetics=(len(Restaurant)/len(current_location))*100
p_Clothing=(len(Cosmetics)/len(current_location))*100
p_Relax=(len(Clothing)/len(current_location))*100
p_Relax=(len(Relax)/len(current_location))*100
# ## Visualizing the Results
# +
# %matplotlib notebook
height = [p_Restaurant, p_Shopping, p_Electronics, p_Cosmetics, p_Clothing, p_Relax]
bars = ('Restaurant', 'Shopping', 'Electronics', 'Cosmetics', 'Clothing', 'Relax')
colors = ['gold', 'yellowgreen', 'lightcoral', 'lightskyblue', 'green', 'orange']
explode = (0.1, 0.1, 0.1, 0.1,0.1,0.1) # explode 1st slice
# Plot
plt.pie(height, explode=explode, labels=bars, colors=colors,autopct='%1.1f%%', shadow=True, startangle=140)
plt.axis('equal')
plt.show()
# -
# ## Implementation of Algorithm
# +
from pandas import DataFrame
df1 = DataFrame (current_location ,columns=['Current Location'])
df2 = DataFrame (general_type ,columns=['Type'])
print (df1)
# -
df_new =pdf
df_new['Current location'] = df1
df_new['type'] = df2
df_new
# ## Pre-processing
# +
from sklearn import preprocessing
label_encoder = preprocessing.LabelEncoder()
df_new['Gender']= label_encoder.fit_transform(df_new['Gender'])
df_new['F Type']= label_encoder.fit_transform(df_new['F Type'])
df_new['type']= label_encoder.fit_transform(df_new['type'])
X1 = df_new.drop('type', axis=1)
X = X1.drop('Current location', axis=1)
X = X.drop('Lat', axis=1)
X = X.drop('Long', axis=1)
X = X.drop('Pid', axis=1)
y = df_new['type']
# -
temp = df_new
temp = temp.drop('Current location', axis=1)
temp = temp.drop('Lat', axis=1)
temp = temp.drop('Long', axis=1)
temp = temp.drop('Pid', axis=1)
temp.to_csv('desktop/new_data.csv', index = None)
# ## Splitting the dataset
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# -
# ## Implementing Decision Tree
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier()
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
from sklearn.metrics import classification_report, accuracy_score
print(accuracy_score(y_pred,y_test))
# ## Implementing Naive Bayes
from sklearn.naive_bayes import MultinomialNB
model = MultinomialNB().fit(X_train, y_train)
y_pred = model.predict(X_test)
print(accuracy_score(y_pred,y_test))
# ### Since Decision Tree seems more accurate than Naive Bayes, So we will proceed with Decision Tree
Age=int(input("Enter Age: "))
Gender=input("Enter Gender (M or F): ")
F_type=input("Enter Flight Type (D or I): ")
# +
g=0
if(Gender=='M'):
g=1
f=0
if(F_type=='I'):
f=1
# -
# ## Predicting the Category
demo=[]
demo.append([Age,g,f])
demo
# +
y=classifier.predict(demo)
temp=y[0]
ans="Relax"
if(temp==0):
ans="Clothing"
print("Clothing")
elif(temp==1):
ans="Cosmetics"
print("Cosmetics")
elif(temp==2):
ans="Electronics"
print("Electronics")
elif(temp==3):
ans="Relax"
print("Relax")
elif(temp==4):
ans="Restaurant"
print("Restaurant")
elif(temp==5):
ans="Shopping"
print("Shopping")
# -
# ## Picking the best Deals
deals=df=pd.read_csv("Desktop/deals.csv")
k=deals.shape[0]
deals
deals['Promotions']
# +
d_type=deals['Type']
d_prom=deals['Promotions']
for i in range(k):
if(d_type[i]==ans or d_type[i]=='Relax'):
print(d_prom[i])
| Location Based Tracking and Personalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Scrape National Monument Areas
#
# The List of National Monuments doesn't include their areas, but each of the pages does. So I scrape them.
import time
import urllib2
import pickle
# %run load_nm.ipynb
nm.columns
nm['url'] = nm['National Monument Name'].apply(
lambda x: 'https://en.wikipedia.org/wiki/{:}_National_Monument'.format(x))
nm['url'] = nm['url'].str.replace(' ', '_')
data = [[], []]
i = 0
for row in nm.iterrows():
if row[1]['National Monument Name'] in data[0]:
continue
print i, row[1]['url'],
try:
a = pd.read_html(row[1]['url'])
except urllib2.HTTPError as e:
print 'not found?', e
dd = {b[0]: b[1] for b in zip(a[0][0], a[0][1])}
data[0].append(row[1]['National Monument Name'])
if 'Area' in dd:
print 'Area =', dd['Area']
data[1].append(dd['Area'])
else:
print 'no area'
data[1].append(0.)
time.sleep(1)
i += 1
dd = {a: b for a, b in zip(data[0], data[1])}
# Unfortunately, not all the pages are formatted equally. I have to manually fixup some of the data.
for k in dd:
if dd[k] == 0.:
print "dd['{:}'] = {:0.0f}".format(k, dd[k])
dd['Fort Ord'] = '14,658 acres'
dd['Newberry'] = '55,500 acres (225 km2)'
dd['El Malpais'] = '114,276 acres (462.46 km2)'
dd['<NAME>'] = '2,526 acres (10.22 km2)'
dd['Charles Young Buffalo Soldiers'] = 0.
dd['John Day Fossil Beds'] = '13,944 acres (5,643 ha)'
dd['<NAME>'] = '43.26 acres (17.51 ha)'
dd['Pullman'] = 0
dd['Statue of Liberty'] = 0
dd['Pacific Remote Islands Marine'] = '313,818,892 acres (1,269,980.00 km2)'
dd['Wupatki'] = '35,422 acres (143.35 km2)'
dd['<NAME>'] = '2,294,343 acres (9,246 km2)'
dd['Gold Butte'] = '300,000 acres'
dd['Military Working Dog Teams'] = 0
dd['Sunset Crater Volcano'] = '3,040 acres (12.3 km2)'
dd['Devils Tower'] = '1,347 acres'
dd['Mount St. Helens'] = '110,000 acres (450 km2)'
dd['Marianas Trench Marine'] = '60,938,240 acres'
if False: # only do this if you have to
with open('national_monuments_areas.pkl', 'w') as ofile:
pickle.dump(dd, ofile)
| NationalProtectedLands/Scrape National Moments From Wikipedia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark_Python3
# language: python
# name: pyspark3
# ---
# # Remote Service Control Manager Handle
#
# ## Playbook Tags
#
# **ID:** WINDISC1908260101
#
# **Author:** <NAME> [@Cyb3rWard0g](https://twitter.com/Cyb3rWard0g)
#
# **References:**
#
# ## ATT&CK Tags
#
# **Tactic:** Discovery
#
# **Technique:** Permissions Level Check
#
# ## Applies To
#
# All Windows Versions
#
# ## Technical Description
#
# Often times, when an adversary lands on an endpoint, the current user does not have local administrator privileges over the compromised system. While some adversaries consider this situation a dead end, others find it very interesting to identify which machines on the network the current user has administrative access to. One common way to accomplish this is by attempting to open up a handle to the service control manager (SCM) database on remote endpoints in the network with SC_MANAGER_ALL_ACCESS (0xF003F) access rights.
#
# The Service Control Manager (SCM) is a remote procedure call (RPC) server, so that service configuration and service control programs can manipulate services on remote machines. Only processes with Administrator privileges are able to open a handle to the SCM database. This database is also known as the `ServicesActive database`. Therefore, it is very effective to check if the current user has administrative or local admin access to other endpoints in the network.
#
# An adversary can simply use the Win32 API function [OpenSCManagerA](https://docs.microsoft.com/en-us/windows/win32/api/winsvc/nf-winsvc-openscmanagera) to attempt to establish a connection to the service control manager (SCM) on the specified computer and open the service control manager database. If this succeeds (A non-zero handle is returned), the current user context has local administrator acess to the remote host.
#
# ### Additional Reading
#
# * [Service Control Manager (SCM)](https://github.com/Cyb3rWard0g/ThreatHunter-Playbook/tree/master/library/service_control_manager.md)
#
# ## Permission Required
#
# User, Administrator
#
# ## Hypothesis
#
# Adversaries might be attempting to open up a handle to the service control manager (SCM) database on remote endpoints to check for local admin access in my environment.
#
# ## Attack Simulation Dataset
#
# | Environment| Name | Description |
# |--------|---------|---------|
# | [Shire](https://github.com/Cyb3rWard0g/mordor/tree/acf9f6be6a386783a20139ceb2faf8146378d603/environment/shire) | [empire_find_local_admin](https://github.com/Cyb3rWard0g/mordor/blob/master/small_datasets/windows/lateral_movement/remote_services_T1021/empire_find_local_admin.md) | A mordor dataset to simulate the use of the OpenSCManagerW Win32API call to establish a handle to a remote host |
#
# ## Recommended Data Sources
#
# | Event ID | Event Name | Log Provider | Audit Category | Audit Sub-Category | ATT&CK Data Source |
# |---------|---------|----------|----------|---------|---------|
# | [4656](https://github.com/Cyb3rWard0g/OSSEM/blob/master/data_dictionaries/windows/security/events/event-4656.md) | A handle to an object was requested | Microsoft-Windows-Security-Auditing | Object Access | Kernel Object | Windows Event Logs |
# | [4674](https://docs.microsoft.com/en-us/windows/security/threat-protection/auditing/event-4674) | An operation was attempted on a privileged object | Microsoft-Windows-Security-Auditing | Privilege Use | Sensitive Privilege Use | Windows Event Logs |
# | [5156](https://docs.microsoft.com/en-us/windows/security/threat-protection/auditing/event-5156) | The Windows Filtering Platform has permitted a connection. | Microsoft-Windows-Security-Auditing | Object Access | Filtering Platform Connection | Process use of network |
# | [3](https://github.com/Cyb3rWard0g/OSSEM/blob/master/data_dictionaries/windows/sysmon/event-3.md) | Network connection | Microsoft-Windows-Sysmon | | | Process use of network |
# ## Data Analytics
# ### Initialize Analytics Engine
from openhunt.logparser import winlogbeat
from pyspark.sql import SparkSession
win = winlogbeat()
spark = SparkSession.builder.appName("Mordor").config("spark.sql.caseSensitive", "True").getOrCreate()
print(spark)
# #### Prepare & Process Mordor File
mordor_file = win.extract_nested_fields("mordor/small_datasets/empire_find_local_admin_2019-05-18224039.json",spark)
# #### Register Mordor DataFrame as a SQL temporary view
mordor_file.createOrReplaceTempView("mordor_file")
# ### Validate Analytic I
# | FP Rate | Source | Analytic Logic | Description |
# |--------|---------|---------|---------|
# | Low | Security | SELECT `@timestamp`, computer_name, SubjectUserName, SubjectLogonId, ProcessName, ObjectName, AccessMask FROM mordor_file WHERE channel = "Security" AND event_id = 4656 AND AccessMask = "0xf003f" AND NOT SubjectLogonId = "0x3e4" AND ObjectName = "ServicesActive" | Detects non-system users failing to get a handle of the SCM database. |
security_handle_df = spark.sql(
'''
SELECT `@timestamp`, computer_name, SubjectUserName, ProcessName, ObjectName
FROM mordor_file
WHERE channel = "Security"
AND event_id = 4656
AND ObjectType = "SC_MANAGER OBJECT"
AND ObjectName = "ServicesActive"
AND AccessMask = "0xf003f"
AND NOT SubjectLogonId = "0x3e4"
'''
)
security_handle_df.show(10,False)
# ### Validate Analytic II
# | FP Rate | Source | Analytic Logic | Description |
# |--------|---------|---------|---------|
# | Low | Security | SELECT `@timestamp`, computer_name, SubjectUserName, SubjectLogonId, ProcessName, PrivilegeList, ObjectServer, ObjectName FROM mordor_file WHERE channel = "Security" AND event_id = 4674 AND ObjectType = "SC_MANAGER OBJECT" AND ObjectName = "ServicesActive" AND PrivilegeList = "SeTakeOwnershipPrivilege" AND NOT SubjectLogonId = "0x3e4" | Look for non-system accounts performing privileged operations on protected subsystem objects such as the SCM database |
security_privileged_df = spark.sql(
'''
SELECT `@timestamp`, computer_name, SubjectUserName, ProcessName, ObjectName, PrivilegeList, ObjectServer
FROM mordor_file
WHERE channel = "Security"
AND event_id = 4674
AND ObjectType = "SC_MANAGER OBJECT"
AND ObjectName = "ServicesActive"
AND PrivilegeList = "SeTakeOwnershipPrivilege"
AND NOT SubjectLogonId = "0x3e4"
'''
)
security_privileged_df.show(10,False)
# ### Validate Analytic III
# | FP Rate | Source | Analytic Logic | Description |
# |--------|---------|---------|---------|
# | Low | Security | SELECT `@timestamp`, computer_name, Application, SourcePort, SourceAddress, DestPort, DestAddress FROM mordor_file WHERE channel = "Security" AND event_id = 5156 AND Application LIKE "%\\\services.exe" AND LayerRTID = 44 | Look for inbound network connections to services.exe from other endpoints in the network. Same SourceAddress, but different computer_name |
security_network_df = spark.sql(
'''
SELECT `@timestamp`, computer_name, Application, SourcePort, SourceAddress, DestPort, DestAddress
FROM mordor_file
WHERE channel = "Security"
AND event_id = 5156
AND Application LIKE "%\\\services.exe"
AND LayerRTID = 44
'''
)
security_network_df.show(10,False)
# ### Validate Analytic IV
# | FP Rate | Source | Analytic Logic | Description |
# |--------|---------|---------|---------|
# | High | Sysmon | SELECT `@timestamp`, computer_name, User, SourcePort, SourceIp, DestinationPort, DestinationIp FROM mordor_file WHERE channel = "Microsoft-Windows-Sysmon/Operational" AND event_id = 3 AND Image LIKE "%\\\services.exe" | Look for several network connection maded by services.exe from different endpoints to the same destination |
security_network_df = spark.sql(
'''
SELECT `@timestamp`, computer_name, User, SourcePort, SourceIp, DestinationPort, DestinationIp
FROM mordor_file
WHERE channel = "Microsoft-Windows-Sysmon/Operational"
AND event_id = 3
AND Image LIKE "%\\\services.exe"
'''
)
security_network_df.show(10,False)
# ### Validate Analytic V
# | FP Rate | Source | Analytic Logic | Description |
# |--------|---------|---------|---------|
# | Low | Security | SELECT o.`@timestamp`, o.computer_name, o.SubjectUserName, o.ObjectType,o.ObjectName, o.PrivilegeList, a.IpAddress FROM mordor_file o INNER JOIN (SELECT computer_name,TargetUserName,TargetLogonId,IpAddress FROM mordor_file WHERE channel = "Security" AND LogonType = 3 AND IpAddress is not null AND NOT TargetUserName LIKE "%$") a ON o.SubjectLogonId = a.TargetLogonId WHERE o.channel = "Security" AND o.event_id = 4674 AND o.ObjectType = "SC_MANAGER OBJECT" AND o.ObjectName = "ServicesActive" AND NOT o.SubjectLogonId = "0x3e4" | Look for non-system accounts performing privileged operations on protected subsystem objects such as the SCM database from other endpoints in the network |
security_4674_4624 = spark.sql(
'''
SELECT o.`@timestamp`, o.computer_name, o.SubjectUserName, o.ObjectType,o.ObjectName, o.PrivilegeList, a.IpAddress
FROM mordor_file o
INNER JOIN (
SELECT computer_name,TargetUserName,TargetLogonId,IpAddress
FROM mordor_file
WHERE channel = "Security"
AND LogonType = 3
AND IpAddress is not null
AND NOT TargetUserName LIKE "%$"
) a
ON o.SubjectLogonId = a.TargetLogonId
WHERE o.channel = "Security"
AND o.event_id = 4674
AND o.ObjectType = "SC_MANAGER OBJECT"
AND o.ObjectName = "ServicesActive"
AND NOT o.SubjectLogonId = "0x3e4"
'''
)
security_4674_4624.show(10,False)
# ## False Positives
#
# ## Detection Blind Spots
#
# ## Hunter Notes
#
# * Event id 4656 gets generated only when the OpenSCManager API call fails to get a handle to the SCM database. There is not SACL for SCM database so success attempts will not be logged.
# * Event id 4674 gets triggered when the SCM database is accessed. Filter known or common accounts that obtain a handle to SCM on a regular basis (i.e vulnerability scanners)
# * You can join security events 4674 and security events 4624 on the LogonID field and filter results on logon type 3 or network to add more context to your query and look for handles to SCM from remote endpoints.
# * Look for the same endpoint or IP address to many remote hosts to find potential aggressive attempts.
# * You can also join security events 4674 where the object name is `servicesactive` (SCM database) with other security events on the object handle. This will allow you to identify what was actually done after the handle was opened. For example, the same handle can be used to create a service (i.e. PSEXESVC)
# * Event id 5156 gets generated on the target as an inbound network event with process name services.exe. You might have to stack the `SourceAddress` field value based on your environment noise.
#
# ## Hunt Output
#
# | Category | Type | Name |
# |--------|---------|---------|
# | Signature | Sigma Rule | [win_scm_database_handle_failure.yml](https://github.com/Cyb3rWard0g/ThreatHunter-Playbook/tree/master/signatures/sigma/win_scm_database_handle_failure.yml) |
# | Signature | Sigma Rule | [win_scm_database_privileged_operation.yml](https://github.com/Cyb3rWard0g/ThreatHunter-Playbook/tree/master/signatures/sigma/win_scm_database_privileged_operation.yml) |
#
# ## References
#
# * https://docs.microsoft.com/en-us/windows/win32/services/service-security-and-access-rights
# * https://github.com/EmpireProject/Empire/blob/dev/data/module_source/situational_awareness/network/powerview.ps1#L15473
# * https://github.com/rapid7/metasploit-framework/blob/master/modules/post/windows/gather/local_admin_search_enum.rb#L217
# * https://github.com/nettitude/PoshC2_Python/blob/master/Modules/Get-System.ps1#L222
# * https://www.pentestgeek.com/metasploit/find-local-admin-with-metasploit
# * http://www.harmj0y.net/blog/penetesting/finding-local-admin-with-the-veil-framework/
# * https://www.slideshare.net/harmj0y/derbycon-the-unintended-risks-of-trusting-active-directory
# * https://docs.microsoft.com/en-us/dotnet/api/system.serviceprocess.servicebase.servicehandle?view=netframework-4.8
# * https://community.rsa.com/community/products/netwitness/blog/2019/04/10/detecting-lateral-movement-in-rsa-netwitness-winexe
| playbooks/windows/07_discovery/T0000_permissions_level_check/remote_service_control_manager_handle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Calling the Cloudmesh Shell
# We provide a simple api that allows cloudmesh to be called just like form a terminal with the commandline. The arguments of the `Shell.cm` command is a simple list. that are typically space separated.
# ## Default command
# Here we show how to pragmatically get the output from the command
# cm default list --format=json
from cloudmesh_client.common.Shell import Shell
result = Shell.cm("default", "list", "--format=json")
print (result)
| notebooks/cm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cifar10 Outlier Detection with NVIDIA Triton
# 
#
# In this example we will deploy an image classification model along with an outlier detector trained on the same dataset. For in depth details on creating an outlier detection model for your own dataset see the [alibi-detect project](https://github.com/SeldonIO/alibi-detect) and associated [documentation](https://docs.seldon.io/projects/alibi-detect/en/latest/). You can find details for this [CIFAR10 example in their documentation](https://docs.seldon.io/projects/alibi-detect/en/latest/examples/od_vae_cifar10.html) as well.
#
#
# Prequisites:
#
# * [Knative eventing installed](https://knative.dev/docs/install/)
# * Ensure the istio-ingressgateway is exposed as a loadbalancer (no auth in this demo)
# * [Seldon Core installed](https://docs.seldon.io/projects/seldon-core/en/latest/workflow/install.html)
# * Ensure you install for istio, e.g. for the helm chart `--set istio.enabled=true`
# * A cluster with NVIDIA GPUs available compatible with Triton Inference Server.
#
# Tested on GKE and Kind with Knative 0.18 and Istio 1.7.3
# !pip install -r requirements_notebook.txt
# Ensure istio gateway installed
# !kubectl apply -f ../../../notebooks/resources/seldon-gateway.yaml
# ## Setup Resources
# !kubectl create namespace cifar10
# %%writefile broker.yaml
apiVersion: eventing.knative.dev/v1
kind: broker
metadata:
name: default
namespace: cifar10
# !kubectl create -f broker.yaml
# +
# %%writefile event-display.yaml
apiVersion: apps/v1
kind: Deployment
metadata:
name: hello-display
namespace: cifar10
spec:
replicas: 1
selector:
matchLabels: &labels
app: hello-display
template:
metadata:
labels: *labels
spec:
containers:
- name: event-display
image: gcr.io/knative-releases/knative.dev/eventing-contrib/cmd/event_display
---
kind: Service
apiVersion: v1
metadata:
name: hello-display
namespace: cifar10
spec:
selector:
app: hello-display
ports:
- protocol: TCP
port: 80
targetPort: 8080
# -
# !kubectl apply -f event-display.yaml
# Create the SeldonDeployment image classification model for Cifar10. We add in a `logger` for requests - the default destination is the namespace Knative Broker.
# %%writefile cifar10.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: triton-cifar10
namespace: cifar10
spec:
predictors:
- componentSpecs:
- metadata: {}
spec:
containers:
- image: nvcr.io/nvidia/tritonserver:21.08-py3
name: cifar10
resources:
limits:
cpu: "1"
memory: 20Gi
nvidia.com/gpu: "1"
requests:
cpu: "1"
memory: 10Gi
nvidia.com/gpu: "1"
graph:
implementation: TRITON_SERVER
logger:
mode: all
url: http://broker-ingress.knative-eventing.svc.cluster.local/cifar10/default
modelUri: gs://seldon-models/triton/tf_cifar10
name: cifar10
type: MODEL
name: default
replicas: 1
protocol: kfserving
# !kubectl apply -f cifar10.yaml
# Create the pretrained VAE Cifar10 Outlier Detector. We forward replies to the message-dumper we started.
# %%writefile cifar10od.yaml
apiVersion: serving.knative.dev/v1
kind: Service
metadata:
name: vae-outlier
namespace: cifar10
spec:
template:
metadata:
annotations:
autoscaling.knative.dev/minScale: "1"
spec:
containers:
- image: seldonio/alibi-detect-server:1.6.0-dev
imagePullPolicy: IfNotPresent
args:
- --model_name
- cifar10od
- --http_port
- '8080'
- --protocol
- kfserving.http
- --storage_uri
- gs://seldon-models/alibi-detect/od/OutlierVAE/cifar10
- --reply_url
- http://hello-display.cifar10
- --event_type
- io.seldon.serving.inference.outlier
- --event_source
- io.seldon.serving.cifar10od
- OutlierDetector
# !kubectl apply -f cifar10od.yaml
# Create a Knative trigger to forward logging events to our Outlier Detector.
# %%writefile trigger.yaml
apiVersion: eventing.knative.dev/v1
kind: Trigger
metadata:
name: vaeoutlier-trigger
namespace: cifar10
spec:
broker: default
filter:
attributes:
type: io.seldon.serving.inference.request
subscriber:
ref:
apiVersion: serving.knative.dev/v1
kind: Service
name: vae-outlier
namespace: cifar10
# !kubectl apply -f trigger.yaml
# Get the IP address of the Istio Ingress Gateway. This assumes you have installed istio with a LoadBalancer.
CLUSTER_IPS = !(kubectl -n istio-system get service istio-ingressgateway -o jsonpath='{.status.loadBalancer.ingress[0].ip}')
CLUSTER_IP = CLUSTER_IPS[0]
print(CLUSTER_IP)
# Optionally add an authorization token here if you need one.Acquiring this token will be dependent on your auth setup.
TOKEN = "Bearer <<PASSWORD>>"
# If you are using Kind or Minikube you will need to port-forward to the istio ingressgateway and uncomment the following
# +
# CLUSTER_IP="localhost:8004"
# -
SERVICE_HOSTNAMES = !(kubectl get ksvc -n cifar10 vae-outlier -o jsonpath='{.status.url}' | cut -d "/" -f 3)
SERVICE_HOSTNAME_VAEOD = SERVICE_HOSTNAMES[0]
print(SERVICE_HOSTNAME_VAEOD)
# +
import json
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
tf.keras.backend.clear_session()
import requests
from alibi_detect.od.vae import OutlierVAE
from alibi_detect.utils.perturbation import apply_mask
from alibi_detect.utils.visualize import plot_feature_outlier_image
train, test = tf.keras.datasets.cifar10.load_data()
X_train, y_train = train
X_test, y_test = test
X_train = X_train.astype("float32") / 255
X_test = X_test.astype("float32") / 255
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
classes = (
"plane",
"car",
"bird",
"cat",
"deer",
"dog",
"frog",
"horse",
"ship",
"truck",
)
def show(X):
plt.imshow(X.reshape(32, 32, 3))
plt.axis("off")
plt.show()
def predict(X):
formData = {
"inputs": [
{
"name": "input_1",
"datatype": "FP32",
"shape": [1, 32, 32, 3],
"data": X.flatten().tolist(),
}
]
}
headers = {"Authorization": TOKEN, "Content-Type": "application/json"}
res = requests.post(
"http://"
+ CLUSTER_IP
+ "/seldon/cifar10/triton-cifar10/v2/models/cifar10/infer",
json=formData,
headers=headers,
)
if res.status_code == 200:
return classes[np.array(res.json()["outputs"][0]["data"]).argmax()]
else:
print("Failed with ", res.status_code)
return []
def outlier(X):
formData = {
"inputs": [
{
"name": "input_1",
"datatype": "FP32",
"shape": [1, 32, 32, 3],
"data": X.flatten().tolist(),
}
]
}
headers = {
"Alibi-Detect-Return-Feature-Score": "true",
"Alibi-Detect-Return-Instance-Score": "true",
"ce-namespace": "default",
"ce-modelid": "cifar10",
"ce-type": "io.seldon.serving.inference.request",
"ce-id": "1234",
"ce-source": "localhost",
"ce-specversion": "1.0",
}
headers["Host"] = SERVICE_HOSTNAME_VAEOD
headers["Authorization"] = TOKEN
res = requests.post("http://" + CLUSTER_IP + "/", json=formData, headers=headers)
if res.status_code == 200:
od = res.json()
od["data"]["feature_score"] = np.array(od["data"]["feature_score"])
od["data"]["instance_score"] = np.array(od["data"]["instance_score"])
return od
else:
print("Failed with ", res.status_code)
return []
# -
# ## Normal Prediction
idx = 1
X = X_train[idx : idx + 1]
show(X)
predict(X)
# Lets check the message dumper for an outlier detection prediction. This should be false.
# res = !kubectl logs -n cifar10 $(kubectl get pod -n cifar10 -l app=hello-display -o jsonpath='{.items[0].metadata.name}')
data = []
for i in range(0, len(res)):
if res[i] == "Data,":
data.append(res[i + 1])
j = json.loads(json.loads(data[0]))
print("Outlier", j["data"]["is_outlier"] == [1])
# ## Outlier Prediction
np.random.seed(0)
X_mask, mask = apply_mask(
X.reshape(1, 32, 32, 3),
mask_size=(10, 10),
n_masks=1,
channels=[0, 1, 2],
mask_type="normal",
noise_distr=(0, 1),
clip_rng=(0, 1),
)
show(X_mask)
predict(X_mask)
# Now lets check the message dumper for a new message. This should show we have found an outlier.
# res = !kubectl logs -n cifar10 $(kubectl get pod -n cifar10 -l app=hello-display -o jsonpath='{.items[0].metadata.name}')
data = []
for i in range(0, len(res)):
if res[i] == "Data,":
data.append(res[i + 1])
j = json.loads(json.loads(data[-1]))
print("Outlier", j["data"]["is_outlier"] == [1])
# We will now call our outlier detector directly and ask for the feature scores to gain more information about why it predicted this instance was an outlier.
od_preds = outlier(X_mask)
# We now plot those feature scores returned by the outlier detector along with our original image.
plot_feature_outlier_image(od_preds, X_mask, X_recon=None)
# ## Tear Down
# !kubectl delete ns cifar10
| components/outlier-detection/nvidia-triton-cifar10/cifar10_outlier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from datetime import datetime
from matplotlib import pyplot as plt
import sys, os
sys.path.append("../..")
sys.path.append("..")
sys.path.append(os.getcwd())
from tslb.src.lzw import *
from tslb.src.utils import *
from tslb.src.regModel import regModel as regModel
plt.rcParams.update({'font.size': 14})
# +
############### dropping some rows and pivoting ############
df = pd.read_pickle("../data/nba_scores_2103-2018.pkl")
##### create df_int : 5 sec interval scores
fr = '5S'
def get_year(df, yr=213):
return df[(df.nbaId > yr*100000) & (df.nbaId < (yr+1)*100000)]
def get_matrix(df, fr='5S'):
df_int = df.copy()
df_int = df_int[df_int.TIME_INT.shift(-1) != df_int.TIME_INT] # remove the rows with the same TIME_INT
### Create Matrix only with Q1-Q4
# 1230 rows : 1230 games in total
# 193 columns : 48 mins (4 Quarters) / 15 secs interval
# only quarter 4
df_q4 = df_int[df_int.TIME_INT <= pd.to_timedelta("00:48:00")]
df_q4.loc[:,'TIME_INT'] = pd.to_datetime(df_q4.loc[:,'TIME_INT'])
# time_index = pd.timedelta_range(start = pd.to_timedelta("00:00:00"), end = pd.to_timedelta("00:48:00"), freq='15s')
df_q4_home = pd.pivot_table(df_q4, values='HOME_SCORE', columns=['nbaId'],index=['TIME_INT'])
df_q4_home = df_q4_home.fillna(method = 'ffill')
df_q4_home = df_q4_home.asfreq(freq=fr, method='ffill')
df_q4_away = pd.pivot_table(df_q4, values='AWAY_SCORE', columns=['nbaId'],index=['TIME_INT'])
df_q4_away = df_q4_away.fillna(method = 'ffill')
df_q4_away = df_q4_away.asfreq(freq=fr, method='ffill')
df_q4_home = df_q4_home.T
df_q4_away = df_q4_away.T
return df_q4_home, df_q4_away
# -
# # First Diff
def get_first_diff(seq):
return (seq.shift(-1) - seq).values[:-1].astype(int)
# +
# get data
fr = '15S'
yr = 214
#####
df_q4_home, df_q4_away = get_matrix(get_year(df, yr=yr), fr=fr)
#####
df_q4 = pd.concat([df_q4_home, df_q4_away])
# +
score = df_q4.iloc[0,:]
plt.title("NBA game score")
plt.plot(score.values)
plt.xlabel("time")
plt.ylabel("score")
plt.show()
plt.title("NBA game score difference (15s interval)")
plt.scatter(range(len(get_first_diff(score))),get_first_diff(score))
plt.xlabel("time")
plt.ylabel("score")
plt.show()
# +
# get data
fr = '15S'
yr = 213
# lbs_df_all = pd.DataFrame()
for yr in [217,218]:
#####
df_q4_home, df_q4_away = get_matrix(get_year(df, yr=yr), fr=fr)
samples = 100
size = df_q4_home.shape[1]-1
myRegModel3 = regModel(3, size, samples)
myRegModel4 = regModel(4, size, samples)
myRegModel5 = regModel(5, size, samples)
myRegModel6 = regModel(6, size, samples)
myRegModel7 = regModel(7, size, samples)
myRegModel3.fit(plot=False)
myRegModel4.fit(plot=False)
myRegModel5.fit(plot=False)
myRegModel6.fit(plot=False)
myRegModel7.fit(plot=False)
lbs_home=[]
for i in range(df_q4_home.shape[0]):
seq = df_q4_home.astype(int).iloc[i]
uncomp_numbers = get_first_diff(seq)
n = max(uncomp_numbers)+1
print(n)
if n==3:
myRegModel = myRegModel3
elif n==4:
myRegModel = myRegModel4
elif n==5:
myRegModel = myRegModel5
elif n==6:
myRegModel = myRegModel6
elif n==7:
myRegModel = myRegModel7
if np.sum(uncomp_numbers <0) !=0:
continue
ratio = lzw_compression_ratio(uncomp_numbers, n)
ent = myRegModel.get_entropy(ratio, "a multinomial sequence", False)
lb = h_inverse(ent, n, a=0.001)
lbs_home.append(lb)
lbs_away=[]
for i in range(df_q4_away.shape[0]):
seq = df_q4_away.astype(int).iloc[i]
uncomp_numbers = get_first_diff(seq)
n = max(uncomp_numbers)+1
print(n)
if n==3:
myRegModel = myRegModel3
elif n==4:
myRegModel = myRegModel4
elif n==5:
myRegModel = myRegModel5
elif n==6:
myRegModel = myRegModel6
elif n==7:
myRegModel = myRegModel7
if np.sum(uncomp_numbers <0) !=0:
continue
ratio = lzw_compression_ratio(uncomp_numbers, n)
ent = myRegModel.get_entropy(ratio, "a multinomial sequence", False)
lb = h_inverse(ent, n, a=0.001)
lbs_away.append(lb)
lbs = np.append(np.array(lbs_home), np.array(lbs_away))
lbs_df = pd.DataFrame(lbs, columns=[yr])
lbs_df_all = pd.concat([lbs_df_all, lbs_df], axis=1)
# -
lbs_df_all
lbs_df
plt.title("Error lower bound histogram \n NBA season 2013")
plt.hist(lbs)
plt.axvline(np.mean(lbs), color='red')
plt.xlim(0,1)
plt.xlabel("classification error")
plt.show()
print("mean : ", np.mean(lbs))
print("median : ", np.median(lbs))
print("min : ", np.min(lbs))
print("max : ", np.max(lbs))
# +
# get data
fr = '15S'
yr = 218
#####
df_q4_home, df_q4_away = get_matrix(get_year(df, yr=yr), fr=fr)
samples = 100
size = df_q4_home.shape[1]-1
myRegModel3 = regModel(3, size, samples)
myRegModel4 = regModel(4, size, samples)
myRegModel5 = regModel(5, size, samples)
myRegModel6 = regModel(6, size, samples)
myRegModel7 = regModel(7, size, samples)
myRegModel3.fit(plot=False)
myRegModel4.fit(plot=False)
myRegModel5.fit(plot=False)
myRegModel6.fit(plot=False)
myRegModel7.fit(plot=False)
lbs=[]
for i in range(df_q4_home.shape[0]):
seq = df_q4_home.astype(int).iloc[i]
uncomp_numbers = get_first_diff(seq)
n = max(uncomp_numbers)+1
if (np.sum(uncomp_numbers<0) != 0):
continue
print(n)
if n==3:
myRegModel = myRegModel3
elif n==4:
myRegModel = myRegModel4
elif n==5:
myRegModel = myRegModel5
elif n==6:
myRegModel = myRegModel6
elif n==7:
myRegModel = myRegModel7
ratio = lzw_compression_ratio(uncomp_numbers, n)
ent = myRegModel.get_entropy(ratio, "a multinomial sequence", False)
lb = h_inverse(ent, n, a=0.001)
lbs.append(lb)
# -
plt.title("Error lower bound histogram \n Season 2018")
plt.hist(lbs)
plt.axvline(np.mean(lbs), color='red')
plt.xlim(0,1)
plt.xlabel("classification error")
plt.show()
print("mean : ", np.mean(lbs))
print("median : ", np.median(lbs))
print("min : ", np.min(lbs))
print("max : ", np.max(lbs))
lbs_df_all.columns=["2013","2014","2015","2016", "2017", "2018"]
plt.title("P(e) distribution per season")
lbs_df_all.boxplot()
plt.xlabel("season")
plt.ylabel("probability of error")
# plt.ylim(0.1,0.4)
plt.show()
| test/nba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Data-Science-and-Data-Analytics-Courses/UniMelb---Database-Systems-Information-Modelling-INFO90002_2019_SM1/blob/master/Resources.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="UdDQ4H0VmPpJ"
# # Clone remote
# + colab_type="code" id="cXqTKogHXEr9" outputId="4267a0d2-1db9-48ef-90d8-f49b4f675ee2" colab={"base_uri": "https://localhost:8080/", "height": 241}
import os, sys
from google.colab import drive
from urllib.parse import urlsplit
from pathlib import Path
URL = "https://github.com/Data-Science-and-Data-Analytics-Courses/UniMelb---Database-Systems-Information-Modelling-INFO90002_2019_SM1"
DRIVE = Path("/drive", "My Drive")
nbdir_rel = ""
def clone(url, dest=".", name="", options="--single-branch -b master", reloc=True):
"""
Clone url into dest
name: if provided, rename repository
options: supported by git-clone (https://git-scm.com/docs/git-clone)
reloc: if True, relocate to repository
"""
rurl = urlsplit(url)
dest = Path(dest).resolve()
repo = dest / (name or Path(rurl.path).name)
# Nested repositories not allowed
# out = !git -C "{dest}" rev-parse
if not out: # inside repository
raise ValueError("Can't clone into existing repository")
# Clone
# !git clone {options} "{rurl.geturl()}" "{repo}"
# Relocate
if reloc:
os.chdir(repo)
return repo
REPO = clone(URL)
drive.mount(DRIVE.parent.as_posix())
NBDIR = REPO/nbdir_rel
os.chdir(NBDIR)
# + [markdown] id="fRrOetKsg-RC" colab_type="text"
# # Import libraries
# + id="XfwrINzwdmkn" colab_type="code" outputId="9eb5ec11-6931-40df-eb7e-a32d5a3326e9" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %run "{REPO/'Packages'/'.Importable.ipynb'}"
sys.path.append(REPO.as_posix())
from Packages import Git, Files
# + [markdown] colab_type="text" id="5sl8tFrDP67Z"
# # Download
# + id="oknoyNQHF1nB" colab_type="code" colab={}
authfile_uni = DRIVE / "UniMelb" / "login.json" # path to UniMelb authentication file
# + [markdown] id="ORA_qxjTnZKg" colab_type="text"
# ## Week 01 Introduction
# Extra materials:
# * [Hoffer video](http://mediaplayer.pearsoncmg.com/_ph_bp2_cc_set.title.Chapter_1%26%2358%3B_Introduction_to_Databases__/bp_hoffer_mdm_10/hoffer10e_ch1_video.m4v)
# + id="4gWwsdjJnpok" colab_type="code" outputId="38fa9d8f-411c-4901-c537-b7ffb2eb7670" colab={"base_uri": "https://localhost:8080/", "height": 34}
urls = {"W01_1-IntroToSubject.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57703905_2/courses/INFO90002_2019_SM1/lectures/W01_1-IntroToSubject.pdf",
"W01_2-IntroToDatabases.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57703906_2/courses/INFO90002_2019_SM1/lectures/W01_2-IntroToDatabases.pdf",
"W01_3-Design.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57703907_2/courses/INFO90002_2019_SM1/lectures/W01_3-Design.pdf"}
dest = "Week 01 Introduction"
Files.download2(urls["W01_1-IntroToSubject.pdf"], dest=dest, authfile=authfile_uni)
Files.download2(urls["W01_2-IntroToDatabases.pdf"], dest=dest, authfile=authfile_uni)
Files.download2(urls["W01_3-Design.pdf"], dest=dest, authfile=authfile_uni)
# + [markdown] colab_type="text" id="ME1QvAuBN0Tc"
# ## Week 02 Implementing a Database
# + colab_type="code" id="rlGx8aeNN0Tm" colab={}
urls = {"W02_1-DesignImplement.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59203564_2/courses/INFO90002_2019_SM1/lectures/W02_1-DesignImplement.pdf",
"W02_2-DataModelling1.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59203565_2/courses/INFO90002_2019_SM1/lectures/W02_2-DataModelling1.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57842726_2/courses/INFO90002_2019_SM1/Wk%202%20Tutorial%20INFO90002%202019%20S1%20v2.pdf",
"TutorialSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942837-dt-content-rid-59566552_2/courses/INFO90002_2019_SM1/Wk%202%20Tutorial%20INFO90002%202019%20v3%20S1%20Solution.pdf"}
dest = "Week 02 Implementing a Database"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="KdODAMjOk5FF"
# ## Week 03 Data Modelling
# Extra materials:
# * [Hoffer video](http://mediaplayer.pearsoncmg.com/_ph_bp2_cc_set.title.Chapter_6%26%2358%3B_Writing_SQL_Queries%26%2358%3B_Part_1_%26%23150%3B_Single_Table_Queries__/bp_hoffer_mdm_10/hoffer10e_ch6_video.m4v)
# + colab_type="code" id="g5lu1XETk5FG" colab={}
urls = {"W03_1-DataModelling2.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59550657_2/courses/INFO90002_2019_SM1/lectures/W03_1-DataModelling2.pdf",
"W03_2-SQL1.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59550658_2/courses/INFO90002_2019_SM1/lectures/W03_2-SQL1.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57655075_2/courses/INFO90002_2019_SM1/Wk%203%20Tutorial%20INFO90002%202019%20S1%281%29.pdf",
"TutorialSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942837-dt-content-rid-59644302_2/courses/INFO90002_2019_SM1/Wk%203%20Tutorial%20INFO90002%202019%20S1%20Solution.pdf"}
dest = "Week 03 Data Modelling"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="4gVfKuHb4_dN"
# ## Week 04 Physical Design
# Extra materials:
# * [Hoffer video](http://mediaplayer.pearsoncmg.com/_ph_bp2_cc_set.title.Chapter_7%26%2358%3B_Writing_SQL_Queries%26%2358%3B_Part_2_%26%23150%3B_Multiple_Table_Queries__/bp_hoffer_mdm_10/hoffer10e_ch7_video.m4v)
# + colab_type="code" id="pm58EFX_4_dP" colab={}
urls = {"W04_1-PhysicalDesign.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59630905_2/courses/INFO90002_2019_SM1/lectures/W04_1-PhysicalDesign.pdf",
"W04_2-DataDictionaries.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59630906_2/courses/INFO90002_2019_SM1/lectures/W04_2-DataDictionaries.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57655093_2/courses/INFO90002_2019_SM1/Wk%204%20Tutorial%20INFO90002%202019%20S1%281%29.pdf",
"TutorialSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942837-dt-content-rid-59796842_2/courses/INFO90002_2019_SM1/Wk%204%20Tutorial%20Answer%20INFO90002%202019%20S1.pdf"}
dest = "Week 04 Physical Design"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="y88ntQjT6Xz3"
# ## Week 05 Normalisation
# Extra materials:
# * [SE Radio](http://www.se-radio.net/2009/06/episode-137-sql-with-jim-melton/)
# + colab_type="code" id="laO4p9f86Xz4" colab={}
urls = {"W05_1-Normalisation.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59704329_2/courses/INFO90002_2019_SM1/lectures/W05_1-Normalisation.pdf",
"W05_2-SQL2.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59704330_2/courses/INFO90002_2019_SM1/lectures/W05_2-SQL2.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57655094_2/courses/INFO90002_2019_SM1/Wk%205%20Tutorial%20INFO90002%202019%20S1.pdf",
"TutorialSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942837-dt-content-rid-59972460_2/courses/INFO90002_2019_SM1/Wk%205%20Tutorial%20INFO90002%202019%20S1%20Solution.pdf"}
dest = "Week 05 Normalisation"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="9DAPJ5tNQ_iZ"
# ## Week 06 SQL
# Extra materials:
# * [Hoffer video](http://mediaplayer.pearsoncmg.com/_ph_bp2_cc_set.title.Chapter_4%26%2358%3B_Normalization__/bp_hoffer_mdm_10/hoffer10e_ch4_video.m4v)
# * [MySQL data types](https://dev.mysql.com/doc/refman/8.0/en/data-types.html)
# + colab_type="code" id="vAAj3pmbQ_ia" colab={}
urls = {"W06_1-SQL3.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59782693_2/courses/INFO90002_2019_SM1/W06_1-SQL3.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59810471_2/courses/INFO90002_2019_SM1/Wk%206%20Tutorial%20INFO90002%202019%20S1%20v2.pdf",
"TutorialSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942837-dt-content-rid-60004213_2/courses/INFO90002_2019_SM1/Wk%206%20Tutorial%20INFO90002%202019%20S1%20Solution%281%29.pdf"}
dest = "Week 06 SQL"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="KHOLMy3oSvL0"
# ## Week 07 Databases in Applications
# Extra materials:
# * [O'Reilly video](http://player.oreilly.com/videos/9781491923320?toc_id=211283)
#
# + colab_type="code" id="9cs1UY5GSvL0" colab={}
urls = {"W07-Applications_fzk.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59884006_2/courses/INFO90002_2019_SM1/W07-Applications_fzk.pdf",
"W07-WebApps_fzk.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59884007_2/courses/INFO90002_2019_SM1/W07-WebApps_fzk.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57718629_2/courses/INFO90002_2019_SM1/Wk%207%20Tutorial%20INFO90002%202019%20S1.pdf"}
dest = "Week 07 Databases in Applications"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="_lFC-Z8j-stX"
# ## Week 08 Data and Database Administration
# Extra materials:
# * [MySQL](https://dev.mysql.com/doc/refman/8.0/en/server-administration.html)
# * [Oracle](https://docs.oracle.com/cd/E11882_01/nav/portal_4.htm)
#
# + colab_type="code" id="CdVTBJ-T-stZ" colab={}
urls = {"W08-Transactions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59980331_2/courses/INFO90002_2019_SM1/W08-Transactions%20v3b.pdf",
"W08-Database Administration.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59980339_2/courses/INFO90002_2019_SM1/W08%20DBA%20ADMIN_DE%20v4%281%29.pdf",
"Tutorial.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-57718631_2/courses/INFO90002_2019_SM1/Wk%208%20Tutorial%20%20INFO90002%202019.pdf"}
dest = "Week 08 Data and Database Administration"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="hsS-XhedC8xu"
# ## Week 09 Distributed Databases
# Extra materials:
# * [Discussion](https://youtu.be/zxwsOueJU4Q)
#
# + colab_type="code" id="u2DnKeyrC8xx" colab={}
urls = {"MelbourneUniPwCPresentation.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-60192887_2/courses/INFO90002_2019_SM1/Melbourne%20Uni%20-%20PwC%20Presentation%2020190506%281%29.pdf",
"W09-DistributedDatabases.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-59980381_2/courses/INFO90002_2019_SM1/W09-DistributedDatabases%20v4.pdf"}
dest = "Week 09 Distributed Databases"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="Fa_zXOqRYoqA"
# ## Week 10 Security & Ethics
# Extra materials:
# * [Security & Ethics: A podcast on Immaneul Kant's Categorical Imperative](https://app.lms.unimelb.edu.au/bbcswebdav/pid-7466182-dt-content-rid-60174135_2/xid-60174135_2)
#
# + colab_type="code" id="DuWU6cWkYoqP" colab={}
urls = {"DWBAIntroINFO90002-2019.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-60192886_2/courses/INFO90002_2019_SM1/DW%26BAIntro%20INFO90002%20-%202019.pdf",
"W10-Security_1_.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-60192888_2/courses/INFO90002_2019_SM1/W10-Security_1_.pdf",
"W10_2-Cloud_DBs.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-60182323_2/courses/INFO90002_2019_SM1/lectures/W10_2-Cloud_DBs.pdf"}
dest = "Week 10 Security & Ethics"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="1TAhuEZFy5Ja"
# ## Week 11 NoSQL Databases
# + colab_type="code" id="iHFgO69Ny5Ji" colab={}
urls = {"W11-NoSQL.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-6942842-dt-content-rid-60315491_2/courses/INFO90002_2019_SM1/W11-NoSQL.pdf",
"Week11ReviewSolutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7478898-dt-content-rid-60669755_2/courses/INFO90002_2019_SM1/Week%2011%20Review%20Solutions.pdf"}
dest = "Week 11 NoSQL Databases"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="_MdMgyC5Wt-U"
# ## Revision
# + colab_type="code" id="pssyFDsDWt-Y" colab={}
urls = {"INFO90002SubjectRevision1.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7478898-dt-content-rid-60310261_2/courses/INFO90002_2019_SM1/INFO90002%20Subject%20Revision%201.pdf",
"INFO90002SubjectRevision1Solutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7478898-dt-content-rid-60669755_2/courses/INFO90002_2019_SM1/Week%2011%20Review%20Solutions.pdf",
"INFO90002RevisionNo2.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7495828-dt-content-rid-60669766_2/courses/INFO90002_2019_SM1/INFO90002%20Revision%20No%202.pdf",
"INFO90002RevisionNo2Answers.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7495828-dt-content-rid-61543624_2/courses/INFO90002_2019_SM1/Revision%20No%202%20Answers.pdf",
"INFO90002RevisionNo3.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7497564-dt-content-rid-61380605_2/courses/INFO90002_2019_SM1/INFO90002%20Revision%20No%203.pdf",
"INFO90002RevisionNo3Solutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7497564-dt-content-rid-61853700_2/courses/INFO90002_2019_SM1/INFO90002%20Revision%20No%203%20Solutions.pdf",
"INFO90002ExamRevisionNo4.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7508140-dt-content-rid-62474282_2/courses/INFO90002_2019_SM1/INFO90002%20Exam%20Revision%20No%204.pdf"}
dest = "Revision"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="BI_WQK_5ohsb"
# ## Assignment 1 ER Modelling
# + colab_type="code" id="oNMv8K61ohsf" colab={}
urls = {"Assignment 1 ER Modelling.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7405982-dt-content-rid-59226013_2/courses/INFO90002_2019_SM1/INFO90002%20S1%202019%20A1.pdf",
"INFO90002 A1 Suggested Solution.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7469209-dt-content-rid-60236936_2/courses/INFO90002_2019_SM1/INFO90002%20A1%20Suggested%20Solution.pdf"}
dest = "Assignment 1 ER Modelling"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] colab_type="text" id="Wy-2mENaTiq_"
# ## Assignment 2 SQL
# + colab_type="code" id="mnOAk04ZTirA" colab={}
urls = {"Assignment 2 SQL.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7434770-dt-content-rid-59816927_2/courses/INFO90002_2019_SM1/INFO90002%20S1%202019%20A2.pdf",
"INFO90002A2S12019Solutions.pdf": "https://app.lms.unimelb.edu.au/bbcswebdav/pid-7497293-dt-content-rid-60918513_2/courses/INFO90002_2019_SM1/INFO90002%20A2%20S1%202019%20Solutions.pdf"}
dest = "Assignment 2 SQL"
for name, url in urls.items():
Files.download2(url, dest=dest, name=name, authfile=authfile_uni)
# + [markdown] id="7ILM4JyuLzue" colab_type="text"
# # Push repository
# + id="IctLlekpKkaO" colab_type="code" colab={}
confile_git = DRIVE.joinpath("GitHub/config.json") # path to GitHub configuration file
authfile_git = DRIVE.joinpath("GitHub/login.json") # path to GitHub authentication file
# + id="RqifDMhBglwC" colab_type="code" outputId="0590f7b7-d6c5-4064-db64-fe0096737a71" colab={"base_uri": "https://localhost:8080/", "height": 238}
message = "Update Revision"
# Configure
Git.config(confile=confile_git)
# Track
# !git add -A
# Commit
# !git commit -m "{message}"
# Push
Git.push(URL, authfile=authfile_git)
| Resources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Read sample text data
# +
import pandas as pd
from rake_nltk import Rake
data = pd.read_csv('./tennis.csv')
# -
# ## Generate summarized text using keyphrases extraction and then take only those sentences which contain any of top 10 keyphrases
# +
print(data.columns)
key_phrases_summary = []
article_ids = []
article_text = []
sources = []
for row in range(data.shape[0]):
rake_processor = Rake()
text = data.iloc[row][1]
print(text)
article_ids.append(data.iloc[row][0])
article_text.append(data.iloc[row][1])
sources.append(data.iloc[row][2])
rake_processor.extract_keywords_from_text(text)
extracted_data = rake_processor.get_ranked_phrases_with_scores()
if len(extracted_data) > 10:
extracted_data = extracted_data[:10]
for kw in extracted_data:
key_phrases.append(kw[1])
result = ''
stmt_count = 0
for stmt in text.split('.'):
isPartOfSummary = False
for item in key_phrases:
if stmt.find(item) != -1:
isPartOfSummary = True
break
if isPartOfSummary:
result += stmt
result += "."
else:
if stmt_count == 0:
result += stmt
result += "."
stmt_count += 1
key_phrases_summary.append(result)
response = {'article_id': article_ids, 'article_text': article_text, 'source': sources, 'key_phrases_summary': key_phrases_summary}
df = pd.DataFrame(response)
df.to_csv('key_phrases_summary_results.csv', index = False)
# -
| NLP/Text_Summarizer/key_phrases_text_summarizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BigQuery basics
#
# [BigQuery](https://cloud.google.com/bigquery/docs/) is a petabyte-scale analytics data warehouse that you can use to run SQL queries over vast amounts of data in near realtime. This page shows you how to get started with the Google BigQuery API using the Python client library.
# ## Import the libraries used in this tutorial
from google.cloud import bigquery
import pandas
# ## Initialize a client
#
# To use the BigQuery Python client library, start by initializing a client. The BigQuery client is used to send and receive messages from the BigQuery API.
#
# ### Client project
# The `bigquery.Client` object uses your default project. Alternatively, you can specify a project in the `Client` constructor. For more information about how the default project is determined, see the [google-auth documentation](https://google-auth.readthedocs.io/en/latest/reference/google.auth.html).
#
#
# ### Client location
# Locations are required for certain BigQuery operations such as creating a dataset. If a location is provided to the client when it is initialized, it will be the default location for jobs, datasets, and tables.
#
# Run the following to create a client with your default project:
client = bigquery.Client(location="US")
print("Client creating using default project: {}".format(client.project))
# To explicitly specify a project when constructing the client, set the `project` parameter:
# +
# client = bigquery.Client(location="US", project="your-project-id")
# -
# ## Run a query on a public dataset
#
# The following example queries the BigQuery `usa_names` public dataset to find the 10 most popular names. `usa_names` is a Social Security Administration dataset that contains all names from Social Security card applications for births that occurred in the United States after 1879.
#
# Use the [Client.query](https://googleapis.github.io/google-cloud-python/latest/bigquery/generated/google.cloud.bigquery.client.Client.html#google.cloud.bigquery.client.Client.query) method to run the query, and the [QueryJob.to_dataframe](https://googleapis.github.io/google-cloud-python/latest/bigquery/generated/google.cloud.bigquery.job.QueryJob.html#google.cloud.bigquery.job.QueryJob.to_dataframe) method to return the results as a pandas [`DataFrame`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html).
# +
query = """
SELECT name, SUM(number) as total
FROM `bigquery-public-data.usa_names.usa_1910_current`
GROUP BY name
ORDER BY total DESC
LIMIT 10
"""
query_job = client.query(
query,
# Location must match that of the dataset(s) referenced in the query.
location="US",
) # API request - starts the query
df = query_job.to_dataframe()
df
# -
# ## Run a parameterized query
#
# BigQuery supports query parameters to help prevent [SQL injection](https://en.wikipedia.org/wiki/SQL_injection) when you construct a query with user input. Query parameters are only available with [standard SQL syntax](https://cloud.google.com/bigquery/docs/reference/standard-sql/). Query parameters can be used as substitutes for arbitrary expressions. Parameters cannot be used as substitutes for identifiers, column names, table names, or other parts of the query.
#
# To specify a parameter, use the `@` character followed by an [identifier](https://cloud.google.com/bigquery/docs/reference/standard-sql/lexical#identifiers), such as `@param_name`. For example, the following query finds all the words in a specific Shakespeare corpus with counts that are at least the specified value.
#
# For more information, see [Running parameterized queries](https://cloud.google.com/bigquery/docs/parameterized-queries) in the BigQuery documentation.
# +
# Define the query
sql = """
SELECT word, word_count
FROM `bigquery-public-data.samples.shakespeare`
WHERE corpus = @corpus
AND word_count >= @min_word_count
ORDER BY word_count DESC;
"""
# Define the parameter values in a query job configuration
job_config = bigquery.QueryJobConfig(
query_parameters=[
bigquery.ScalarQueryParameter("corpus", "STRING", "romeoandjuliet"),
bigquery.ScalarQueryParameter("min_word_count", "INT64", 250),
]
)
# Start the query job
query_job = client.query(sql, location="US", job_config=job_config)
# Return the results as a pandas DataFrame
query_job.to_dataframe()
# -
# ## Create a new dataset
#
# A dataset is contained within a specific [project](https://cloud.google.com/bigquery/docs/projects). Datasets are top-level containers that are used to organize and control access to your [tables](https://cloud.google.com/bigquery/docs/tables) and [views](https://cloud.google.com/bigquery/docs/views). A table or view must belong to a dataset. You need to create at least one dataset before [loading data into BigQuery](https://cloud.google.com/bigquery/loading-data-into-bigquery).
# +
# Define a name for the new dataset.
dataset_id = 'your_new_dataset'
# The project defaults to the Client's project if not specified.
dataset = client.create_dataset(dataset_id) # API request
# -
# ## Write query results to a destination table
#
# For more information, see [Writing query results](https://cloud.google.com/bigquery/docs/writing-results) in the BigQuery documentation.
# +
sql = """
SELECT corpus
FROM `bigquery-public-data.samples.shakespeare`
GROUP BY corpus;
"""
table_ref = dataset.table("your_new_table_id")
job_config = bigquery.QueryJobConfig(
destination=table_ref
)
# Start the query, passing in the extra configuration.
query_job = client.query(sql, location="US", job_config=job_config)
query_job.result() # Waits for the query to finish
print("Query results loaded to table {}".format(table_ref.path))
# -
# ## Load data from a pandas DataFrame to a new table
# +
records = [
{"title": "The Meaning of Life", "release_year": 1983},
{"title": "Monty Python and the Holy Grail", "release_year": 1975},
{"title": "Life of Brian", "release_year": 1979},
{"title": "And Now for Something Completely Different", "release_year": 1971},
]
# Optionally set explicit indices.
# If indices are not specified, a column will be created for the default
# indices created by pandas.
index = ["Q24980", "Q25043", "Q24953", "Q16403"]
df = pandas.DataFrame(records, index=pandas.Index(index, name="title"))
table_ref = dataset.table("monty_python")
job = client.load_table_from_dataframe(df, table_ref, location="US")
job.result() # Waits for table load to complete.
print("Loaded dataframe to {}".format(table_ref.path))
# -
# ## Load data from a local file to a table
#
# The following example demonstrates how to load a local CSV file into a new table. See [SourceFormat](https://googleapis.github.io/google-cloud-python/latest/bigquery/generated/google.cloud.bigquery.job.SourceFormat.html#google.cloud.bigquery.job.SourceFormat) in the Python client library documentation for a list of available source formats. For more information, see [Loading Data into BigQuery from a local data source](https://cloud.google.com/bigquery/docs/loading-data-local) in the BigQuery documentation.
# +
source_filename = 'resources/us-states.csv'
table_ref = dataset.table('us_states_from_local_file')
job_config = bigquery.LoadJobConfig(
source_format=bigquery.SourceFormat.CSV,
skip_leading_rows=1,
autodetect=True
)
with open(source_filename, 'rb') as source_file:
job = client.load_table_from_file(
source_file,
table_ref,
location='US', # Must match the destination dataset location.
job_config=job_config) # API request
job.result() # Waits for table load to complete.
print('Loaded {} rows into {}:{}.'.format(
job.output_rows, dataset_id, table_ref.path))
# -
# ## Load data from Cloud Storage to a table
#
# The following example demonstrates how to load a local CSV file into a new table. See [SourceFormat](https://googleapis.github.io/google-cloud-python/latest/bigquery/generated/google.cloud.bigquery.job.SourceFormat.html#google.cloud.bigquery.job.SourceFormat) in the Python client library documentation for a list of available source formats. For more information, see [Introduction to loading data from Cloud Storage](https://cloud.google.com/bigquery/docs/loading-data-cloud-storage) in the BigQuery documentation.
# +
# Configure the load job
job_config = bigquery.LoadJobConfig(
schema=[
bigquery.SchemaField('name', 'STRING'),
bigquery.SchemaField('post_abbr', 'STRING')
],
skip_leading_rows=1,
# The source format defaults to CSV. The line below is optional.
source_format=bigquery.SourceFormat.CSV
)
uri = 'gs://cloud-samples-data/bigquery/us-states/us-states.csv'
destination_table_ref = dataset.table('us_states_from_gcs')
# Start the load job
load_job = client.load_table_from_uri(
uri, destination_table_ref, job_config=job_config)
print('Starting job {}'.format(load_job.job_id))
load_job.result() # Waits for table load to complete.
print('Job finished.')
# Retreive the destination table
destination_table = client.get_table(table_ref)
print('Loaded {} rows.'.format(destination_table.num_rows))
# -
# ## Cleaning Up
#
# The following code deletes the dataset created for this tutorial, including all tables in the dataset.
# +
# Retrieve the dataset from the API
dataset = client.get_dataset(client.dataset(dataset_id))
# Delete the dataset and its contents
client.delete_dataset(dataset, delete_contents=True)
print('Deleted dataset: {}'.format(dataset.path))
| notebooks/tutorials/bigquery/BigQuery basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Version 1.0.3
# # Pandas basics
# Hi! In this programming assignment you need to refresh your `pandas` knowledge. You will need to do several [`groupby`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html)s and [`join`]()`s to solve the task.
# +
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from grader import Grader
# +
DATA_FOLDER = '../readonly/final_project_data/'
transactions = pd.read_csv(os.path.join(DATA_FOLDER, 'sales_train.csv.gz'))
items = pd.read_csv(os.path.join(DATA_FOLDER, 'items.csv'))
item_categories = pd.read_csv(os.path.join(DATA_FOLDER, 'item_categories.csv'))
shops = pd.read_csv(os.path.join(DATA_FOLDER, 'shops.csv'))
# -
# The dataset we are going to use is taken from the competition, that serves as the final project for this course. You can find complete data description at the [competition web page](https://www.kaggle.com/c/competitive-data-science-final-project/data). To join the competition use [this link](https://www.kaggle.com/t/1ea93815dca248e99221df42ebde3540).
# ## Grading
# We will create a grader instace below and use it to collect your answers. When function `submit_tag` is called, grader will store your answer *locally*. The answers will *not* be submited to the platform immediately so you can call `submit_tag` function as many times as you need.
#
# When you are ready to push your answers to the platform you should fill your credentials and run `submit` function in the <a href="#Authorization-&-Submission">last paragraph</a> of the assignment.
grader = Grader()
# # Task
# Let's start with a simple task.
#
# <ol start="0">
# <li><b>Print the shape of the loaded dataframes and use [`df.head`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.head.html) function to print several rows. Examine the features you are given.</b></li>
# </ol>
# +
# YOUR CODE GOES HERE
print(transactions.shape)
transactions.head()
# -
# Now use your `pandas` skills to get answers for the following questions.
# The first question is:
#
# 1. ** What was the maximum total revenue among all the shops in September, 2014?**
#
#
# * Hereinafter *revenue* refers to total sales minus value of goods returned.
#
# *Hints:*
#
# * Sometimes items are returned, find such examples in the dataset.
# * It is handy to split `date` field into [`day`, `month`, `year`] components and use `df.year == 14` and `df.month == 9` in order to select target subset of dates.
# * You may work with `date` feature as with strings, or you may first convert it to `pd.datetime` type with `pd.to_datetime` function, but do not forget to set correct `format` argument.
# YOUR CODE GOES HERE
transactions['date'] = pd.to_datetime(transactions['date'], format='%d.%m.%Y')
trans = transactions[(transactions['date'].dt.year == 2014) & (transactions['date'].dt.month == 9)]
# PUT YOUR ANSWER IN THIS VARIABLE
max_revenue = (trans['item_price'] * trans['item_cnt_day']).groupby(trans['shop_id']).sum().max()
grader.submit_tag('max_revenue', max_revenue)
# Great! Let's move on and answer another question:
#
# <ol start="2">
# <li><b>What item category generated the highest revenue in summer 2014?</b></li>
# </ol>
#
# * Submit `id` of the category found.
#
# * Here we call "summer" the period from June to August.
#
# *Hints:*
#
# * Note, that for an object `x` of type `pd.Series`: `x.argmax()` returns **index** of the maximum element. `pd.Series` can have non-trivial index (not `[1, 2, 3, ... ]`).
# YOUR CODE GOES HERE
trans = transactions[(transactions['date'].dt.year == 2014) & (transactions['date'].dt.month >= 6) & (transactions['date'].dt.month <= 8)]
trans = pd.merge(trans, items, how='left', on='item_id')
# PUT YOUR ANSWER IN THIS VARIABLE
category_id_with_max_revenue = (trans['item_price'] * trans['item_cnt_day']).groupby(trans['item_category_id']).sum().idxmax()
grader.submit_tag('category_id_with_max_revenue', category_id_with_max_revenue)
# <ol start="3">
# <li><b>How many items are there, such that their price stays constant (to the best of our knowledge) during the whole period of time?</b></li>
# </ol>
#
# * Let's assume, that the items are returned for the same price as they had been sold.
# YOUR CODE GOES HERE
trans = transactions.groupby('item_id')['item_price']
# PUT YOUR ANSWER IN THIS VARIABLE
num_items_constant_price = (trans.nunique() == 1).sum()
grader.submit_tag('num_items_constant_price', num_items_constant_price)
# Remember, the data can sometimes be noisy.
# <ol start="4">
# <li><b>What was the variance of the number of sold items per day sequence for the shop with `shop_id = 25` in December, 2014? Do not count the items, that were sold but returned back later.</b></li>
# </ol>
#
# * Fill `total_num_items_sold` and `days` arrays, and plot the sequence with the code below.
# * Then compute variance. Remember, there can be differences in how you normalize variance (biased or unbiased estimate, see [link](https://math.stackexchange.com/questions/496627/the-difference-between-unbiased-biased-estimator-variance)). Compute ***unbiased*** estimate (use the right value for `ddof` argument in `pd.var` or `np.var`).
# * If there were no sales at a given day, ***do not*** impute missing value with zero, just ignore that day
# +
shop_id = 25
trans = transactions[(transactions['shop_id'] == shop_id) & (transactions['date'].dt.year == 2014) & (transactions['date'].dt.month == 12)]
trans = trans.groupby('date')['item_cnt_day'].sum()
total_num_items_sold = trans.values # YOUR CODE GOES HERE
days = trans.index # YOUR CODE GOES HERE
# Plot it
plt.plot(days, total_num_items_sold)
plt.ylabel('Num items')
plt.xlabel('Day')
plt.title("Daily revenue for shop_id = 25")
plt.show()
total_num_items_sold_var = trans.var() # PUT YOUR ANSWER IN THIS VARIABLE
grader.submit_tag('total_num_items_sold_var', total_num_items_sold_var)
# -
# ## Authorization & Submission
# To submit assignment to Cousera platform, please, enter your e-mail and token into the variables below. You can generate token on the programming assignment page. *Note:* Token expires 30 minutes after generation.
STUDENT_EMAIL = '<EMAIL>'
STUDENT_TOKEN = '<KEY>'
grader.status()
grader.submit(STUDENT_EMAIL, STUDENT_TOKEN)
# Well done! :)
| PandasBasics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv("addresses.csv")
df.columns
df.head()
df.borough.value_counts()
df.building_category.value_counts()
df.neighborhood.value_counts()
df['age'] = 2020 - df['year_built']
df.head()
df = df.rename(columns = {'latitude': 'longitude', 'longitude': 'latitude'})
df.head()
# +
import geopy.distance
from geopy.distance import lonlat, distance
from tqdm import tqdm
tqdm.pandas()
central_park = (40.785091, -73.968285)
df['distance'] = df.progress_apply(lambda row: geopy.distance.great_circle(central_park, (row.latitude, row.longitude)).km, axis=1)
# -
import numpy as np
df['distance'] = df['distance'].apply(lambda x: np.nan if x >= 100 else x)
df.head()
df.to_csv("test_data.csv", index = False)
# +
## PARSE OUT SALE PRICE DATA INTO CATEGORIES
def price_parse(x):
if x <= 1000000:
return '0-1m'
elif x > 1000000 and x <= 10000000:
return '1m-10m'
elif x > 10000000 and x <= 100000000:
return '10m-100m'
elif x > 100000000:
return '100m+'
else:
return 'na'
##Put categorized sale price data in df
df['cat_price'] = df['sale_price'].apply(lambda x: price_parse(x))
# -
df['square_ft'] = df['square_ft'].str.replace(',', '')
df_num = df[['sale_price', 'distance', 'res_units', 'comm_units', 'total_units','age', 'square_ft']].astype(float)
df = df[(df['sale_price'] > ((100000))) & (df['sale_price'] < ((5000000)))]
# +
#df_num = df_num.reset_index(drop=True)
# -
for i, col in enumerate(df_num.columns):
plt.figure(i)
sns.distplot(df_num[col].dropna())
# from scipy.stats import skew
# skewed = df[df_num.columns].apply(lambda x: skew(x.dropna().astype(float)))
# skewed = skewed[skewed > 0.75]
# skewed = skewed.index
# df[skewed] = np.log1p(df[skewed].astype(float))
# +
# from sklearn.preprocessing import StandardScaler
# scaler = StandardScaler()
# scaler.fit(df[df_num.columns].dropna())
# scaled = scaler.transform(df[df_num.columns])
# for i, col in enumerate(df_num.columns):
# print(i)
# df_num[col] = scaled[:,i]
# -
# df_num['sale_price'] = df_num['sale_price'].apply(lambda x: np.log1p(x))
# df_num['square_ft'] = df_num['square_ft'].apply(lambda x: np.log1p(x))
for i in df_num:
df_num[i] = df_num[i].apply(lambda x: np.log1p(x))
df.head()
for i, col in enumerate(df_num.columns):
plt.figure(i)
sns.distplot(df_num[col].dropna())
corr = df_num.dropna().corr()
cmap = sns.diverging_palette(220,10,as_cmap=True)
sns.heatmap(corr,vmax=.3,center=0,cmap=cmap,square=True, linewidth=.5, cbar_kws={"shrink":.5})
# +
from datetime import datetime
strDate = '7/16/2020'
df1 = df.copy()
df1['sale_date'] = df['sale_date'].apply(lambda x: datetime.strptime(x, '%m/%d/%Y'))
# -
df1.head()
d1 = datetime(2020, 3, 1)
df['date'] = pd.to_datetime(df['sale_date'])
df['day'] = (df['date'] - d1).dt.days
df_cat = df[['borough', 'neighborhood', 'building_category', 'cat_price']]
# +
##Rename Buroughs
def borough_parser(x):
if x == 1:
return 'Manhattan'
elif x == 2:
return 'Bronx'
elif x == 3:
return 'Brooklyn'
elif x == 4:
return 'Queens'
elif x == 5:
return 'Staten Island'
else:
return 'na'
df_cat['borough'] = df['borough'].apply(lambda x: borough_parser(x))
# -
df_cat.head()
df['borough'] = df_cat['borough']
for i in df_cat.columns:
chart = sns.barplot(x=df_cat[i].value_counts()[:20].index,y=df_cat[i].value_counts()[:20], data=df_cat)
print("Bar Graph for %s total = %d" % (i,len(df_cat[i].value_counts()[:20])))
chart.set_xticklabels(chart.get_xticklabels(),rotation=75,fontweight='light',
fontsize='x-small',horizontalalignment='right')
plt.show()
b_pivot = pd.pivot_table(df, index = 'borough',
values='sale_price').sort_values('sale_price',ascending=False)
sale_chart = df[['borough', 'neighborhood', 'building_category', 'zip_code']]
for i in sale_chart.columns:
plot_order = df.groupby(i)['sale_price'].mean().sort_values(ascending=False).index.values
chart = sns.barplot(data=df, x= sale_chart[i], y='sale_price',estimator=np.mean,order=plot_order[:20])
chart.set_xticklabels(chart.get_xticklabels(),rotation=75,fontweight='light',
fontsize='x-small',horizontalalignment='right')
plt.show()
df.corr()
df['borough'] = df_cat['borough']
for i in df_num:
df[i] = df_num[i]
df.head()
corr = df.dropna().corr()
cmap = sns.diverging_palette(220,10,as_cmap=True)
sns.heatmap(corr,vmax=.3,center=0,cmap=cmap,square=True, linewidth=.5, cbar_kws={"shrink":.5})
df.to_csv("explored_data.csv", index = False)
| eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy.io as sio
# +
data_dir = '/media/nsaleheen/BCE46043E460024E/research/data/Smoking_data/data/Memphis_Smoking_Lab/basicfeature/'
filename = 'p01_s02_basicfeature.mat'
def append_file(filename, txt):
fh = open(filename, 'a')
fh.write(txt + '\n')
fh.close()
# +
def process_basicfeature_mat(data_dir, filename, pid, sid):
B = sio.loadmat(data_dir + filename)
B=B['B']
B = B[0,0]
smoking_epis = B['smoking_episode'][0]
print(len(smoking_epis))
for i, se in enumerate(smoking_epis):
starttimestamp = int(se[0]['starttimestamp'][0][0][0])
endtimestamp = int(se[0]['endtimestamp'][0][0][0])
append_file(data_dir + 'ground_truth/' + pid+'_'+sid+'_smoking_epi.csv', str(starttimestamp) + ',' + str(endtimestamp))
# print(starttimestamp)
# print(endtimestamp)
puffs = se[0]['puff'][0][0][0][0][0]
# print(puffs)
for p in puffs:
append_file(data_dir + 'ground_truth/' + pid+'_'+sid+'_smoking_puff.csv', str(int(p)) + ',' + str(i+1))
pid = 'p06'
sid = 's01'
filename = pid + '_' + sid + '_basicfeature.mat'
process_basicfeature_mat(data_dir, filename, pid, sid)
# from os import listdir
# from os.path import isfile, join
# onlyfiles = [f for f in listdir(data_dir) if isfile(join(data_dir, f))]
# for fn in onlyfiles:
# print(fn)
# pid = fn[:3]
# sid = fn[4:7]
# process_basicfeature_mat(data_dir, fn, pid, sid)
# print(onlyfiles)
# -
fn = 'p01_s02_basicfeature.mat'
pid = fn[:3]
sid = fn[4:7]
print(pid)
print(sid)
| read_6smoker_matlabfile_storesmokingtimes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pylab as pl
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from sklearn.utils import shuffle
from sklearn.svm import SVC
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.model_selection import cross_val_score, GridSearchCV
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
# print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
# -
# Training data
train= pd.read_csv(r'C:\Users\nguyent2\Desktop\Kaggle-Competition-on-HR-Analytics-Employee-Promotion-Data\dataset\train.csv')
train.head()
# **Features** <br/>
# employee_id<br/>
# department<br/>
# region<br/>
# education<br/>
# gender<br/>
# recruitment_channel<br/>
# no_of_trainings<br/>
# age<br/>
# previous_year_rating<br/>
# length_of_service<br/>
# awards_won?<br/>
# avg_training_score<br/>
# is_promoted<br/>
# Visualization
# +
#Count how many different departments
#Check if there is ISNULL information in any column
print(train['employee_id'].nunique(), "different employee_id items")
print(train['employee_id'].isnull().sum())
print(train['department'].nunique(), "different departments")
print(train['department'].isnull().sum())
print(train['region'].nunique(), "different regions")
print(train['region'].isnull().sum())
print(train['education'].nunique(), "different education items")
print(train['education'].isnull().sum())
print(train['gender'].nunique(), "different genders")
print(train['gender'].isnull().sum())
print(train['recruitment_channel'].nunique(), "different recruitment channels")
print(train['recruitment_channel'].isnull().sum())
print(train['no_of_trainings'].nunique(), "different no of trainings")
print(train['no_of_trainings'].isnull().sum())
print(train['age'].nunique(), "different ages")
print(train['age'].isnull().sum())
print(train['previous_year_rating'].nunique(), "different previous year ratings")
print(train['previous_year_rating'].isnull().sum())
print(train['length_of_service'].nunique(), "different lengths of service")
print(train['length_of_service'].isnull().sum())
print(train['awards_won?'].nunique(), "different awards_won")
print(train['awards_won?'].isnull().sum())
print(train['avg_training_score'].nunique(), "different avg_training_score")
print(train['avg_training_score'].isnull().sum())
print(train['is_promoted'].nunique(), "different is_promoted")
print(train['is_promoted'].isnull().sum())
# -
# Since there are so many ISNULL in the column of "education" (and also the column of "previous year ratings"), I tried to calculate the percentage between 1 and 0 (of the column "is_promoted").
# Barplots showing the frequency of each category separated by label
# +
#barplots showing the frequency of each category separated by label
plt.figure(figsize=[15,20])
fft=["department", "region","education", "gender","recruitment_channel","no_of_trainings", "age","previous_year_rating", "length_of_service", "awards_won?", "avg_training_score"]
n=1
for f in fft:
plt.subplot(12,1,n)
sns.countplot(x=f, hue='is_promoted', edgecolor="black", alpha=0.5, data=train)
sns.despine()
plt.title("Countplot of {} by is_promoted".format(f))
n=n+1
plt.tight_layout()
plt.show()
| Model_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import matplotlib.pyplot as plt
import glob
from PIL import Image
import numpy as np
from sklearn.utils import shuffle
from tensorflow.python import keras
from tensorflow.python.keras import Sequential
from tensorflow.python.keras.layers import Dense, InputLayer, Conv2D, MaxPool2D, Flatten, BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from sklearn.model_selection import train_test_split
import keras.backend as K
from keras.callbacks import History
loss_ges = np.array([])
val_loss_ges = np.array([])
# %matplotlib inline
np.set_printoptions(precision=4)
np.set_printoptions(suppress=True)
# +
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
x_data = []
y_data = []
for aktfile in files:
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
test_image = np.reshape(test_image, (32,32,3))
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
x_data.append(test_image)
zw = np.array([zahl])
y_data.append(zw)
x_data = np.array(x_data)
y_data = np.array(y_data)
print(x_data.shape)
print(y_data.shape)
x_data, y_data = shuffle(x_data, y_data)
X_train, X_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.2)
# +
model = Sequential()
model.add(BatchNormalization(input_shape=(32,32,3)))
model.add(Conv2D(32, (5, 5), input_shape=(32,32,3), padding='same', activation="relu"))
model.add(MaxPool2D(pool_size=(4,4)))
model.add(Conv2D(16, (5, 5), padding='same'))
model.add(MaxPool2D(pool_size=(4,4)))
model.add(Conv2D(16, (3, 3), padding='same'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(64,activation="relu"))
model.add(Dense(8,activation="linear"))
model.add(Dense(1))
model.summary()
def Periodic_Loss(y_true, y_pred):
dif1 = K.abs(y_pred - y_true)
dif2 = K.abs(dif1 + K.constant(1))
# dif3 = K.abs(dif1 + K.constant(-1))
dif = K.minimum(dif1, dif2)
# dif = K.minimum(dif, dif3)
ret = K.mean(K.square(dif), axis=-1)
return ret
model.compile(loss=Periodic_Loss, optimizer=keras.optimizers.Adadelta(), metrics = ["accuracy"])
#model.compile(loss=keras.losses.mean_squared_error, optimizer=keras.optimizers.Adadelta(), metrics = ["accuracy"])
# +
Batch_Size = 8
Epoch_Anz = 20
Shift_Range = 0
Brightness_Range = 0.3
datagen = ImageDataGenerator(width_shift_range=[-Shift_Range,Shift_Range], height_shift_range=[-Shift_Range,Shift_Range],brightness_range=[1-Brightness_Range,1+Brightness_Range])
train_iterator = datagen.flow(X_train, y_train, batch_size=Batch_Size)
validation_iterator = datagen.flow(X_test, y_test, batch_size=Batch_Size)
history = model.fit_generator(train_iterator, validation_data = validation_iterator, epochs = Epoch_Anz)
# +
loss_ges = np.append(loss_ges, history.history['loss'])
val_loss_ges = np.append(val_loss_ges, history.history['val_loss'])
plt.semilogy(history.history['loss'])
plt.semilogy(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.show()
# +
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
res = []
for aktfile in files:
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
img = np.reshape(test_image,[1,32,32,3])
classes = model.predict(img)
zw1 = zahl
zw2 = round(classes[0][0], 2)
zw3 = round(classes[0][0] - zahl, 2)
zw4a = abs(zw3)
zw4b = abs(zw3+1)
zw4c = abs(zw3-1)
zw4 = zw3
if zw4b < zw4a:
zw4 = zw3+1
zw4a = zw4b
if zw4c < zw4a:
zw4 = zw3-1
res.append(np.array([zw1, zw2, zw3, zw4]))
# print(base, ', ', zw1, ', ', round(zw2, 2), ', ', round(zw3, 2), ', ', round(zw4, 2))
res = np.asarray(res)
statistic = np.array([np.mean(res[:,3]), np.std(res[:,3]), np.min(res[:,3]), np.max(res[:,3])])
print(statistic)
res_step_1 = res
# -
plt.plot(res[:,0])
plt.plot(res[:,1])
plt.title('Result')
plt.ylabel('Analog Value')
plt.xlabel('#Picture')
plt.legend(['real','model'], loc='upper left')
plt.show()
plt.plot(res[:,3])
plt.title('Deviation')
plt.ylabel('Deviation from expected value')
plt.xlabel('#Picture')
plt.legend(['model'], loc='upper left')
plt.show()
# +
Batch_Size = 8
Epoch_Anz = 40
Shift_Range = 1
Brightness_Range = 0.3
datagen = ImageDataGenerator(width_shift_range=[-Shift_Range,Shift_Range], height_shift_range=[-Shift_Range,Shift_Range],brightness_range=[1-Brightness_Range,1+Brightness_Range])
train_iterator = datagen.flow(X_train, y_train, batch_size=Batch_Size)
validation_iterator = datagen.flow(X_test, y_test, batch_size=Batch_Size)
history = model.fit_generator(train_iterator, validation_data = validation_iterator, epochs = Epoch_Anz)
# +
loss_ges = np.append(loss_ges, history.history['loss'])
val_loss_ges = np.append(val_loss_ges, history.history['val_loss'])
plt.semilogy(history.history['loss'])
plt.semilogy(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.show()
# +
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
res = []
for aktfile in files:
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
img = np.reshape(test_image,[1,32,32,3])
classes = model.predict(img)
zw1 = zahl
zw2 = round(classes[0][0], 2)
zw3 = round(classes[0][0] - zahl, 2)
zw4a = abs(zw3)
zw4b = abs(zw3+1)
zw4c = abs(zw3-1)
zw4 = zw3
if zw4b < zw4a:
zw4 = zw3+1
zw4a = zw4b
if zw4c < zw4a:
zw4 = zw3-1
res.append(np.array([zw1, zw2, zw3, zw4]))
# print(base, ', ', zw1, ', ', round(zw2, 2), ', ', round(zw3, 2), ', ', round(zw4, 2))
res = np.asarray(res)
statistic = np.array([np.mean(res[:,3]), np.std(res[:,3]), np.min(res[:,3]), np.max(res[:,3])])
print(statistic)
res_step_1 = res
# -
plt.plot(res[:,0])
plt.plot(res[:,1])
plt.title('Result')
plt.ylabel('Analog Value')
plt.xlabel('#Picture')
plt.legend(['real','model'], loc='upper left')
plt.show()
plt.plot(res[:,3])
plt.title('Deviation')
plt.ylabel('Deviation from expected value')
plt.xlabel('#Picture')
plt.legend(['model'], loc='upper left')
plt.ylim(-0.3, 0.3)
plt.show()
model.save("test.h5")
plt.semilogy(loss_ges)
plt.semilogy(val_loss_ges)
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.ylim(1E-1, 5E-5)
plt.show()
| background_info/jyupiter_files/06a_CNN-Smaller-0.5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Create a Ground Truth Streaming Labeling Job
#
# You can use a streaming labeling job to perpetually send new data objects to Amazon SageMaker Ground Truth to be labeled. Ground Truth streaming labeling jobs remain active until they are manually stopped or have been idle for more than 10 days. You can intermittently send new data objects to workers while the labeling job is active.
#
# Use this notebook to create a Ground Truth streaming labeling job using any of the [built-in task types](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-task-types.html). You can make necessary parameter changes for the custom workflow. You can either configure the notebook to create a labeling job using your own input data, or run the notebook on *default* mode and use provided, image input data. **To use your own input data, set `DEFAULT` to `False`**.
DEFAULT=True
# To read more about streaming labeling jobs, see the Amazon SageMaker documentation on [Ground Truth Streaming Labeling Jobs](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-streaming-labeling-job.html).
#
# To learn more about each step in this notebook, refer to [Create a Streaming Labeling Job](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-streaming-create-job.html).
# ## Get latest version of AWS python SDK
# +
# !pip install -q --upgrade pip
# !pip install awscli -q --upgrade
# !pip install botocore -q --upgrade
# !pip install boto3 -q --upgrade
# !pip install sagemaker -q --upgrade
# NOTE: Restart Kernel after the above command
# -
import boto3
import botocore
import json
import time
import sagemaker
import re
# ## Prerequisites
#
# You will create some of the resources you need to launch a Ground Truth streaming labeling job in this notebook. You must create the following resources before executing this notebook:
#
# * A work team. A work team is a group of workers that complete labeling tasks. If you want to preview the worker UI and execute the labeling task you will need to create a private work team, add yourself as a worker to this team, and provide the work team ARN below. If you do not want to use a private or vendor work team ARN, set `private_work_team` to `False` to use the Amazon Mechanical Turk workforce. To learn more about private, vendor, and Amazon Mechanical Turk workforces, see [Create and Manage Workforces
# ](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-workforce-management.html).
# * **IMPORTANT**: 3D point cloud and video frame labeling jobs only support private and vendor workforces. If you plan to use 3D point cloud or video frame input data, specify a private or vendor workforce below for WORKTEAM_ARN.
# +
private_work_team = True # Set it to false if using Amazon Mechanical Turk Workforce
if(private_work_team):
WORKTEAM_ARN = '<<ADD WORK TEAM ARN HERE>>'
else :
region = boto3.session.Session().region_name
WORKTEAM_ARN = f'arn:aws:sagemaker:{region}:394669845002:workteam/public-crowd/default'
print(f'This notebook will use the work team ARN: {WORKTEAM_ARN}')
# -
# Make sure workteam arn is populated if private work team is chosen
assert (WORKTEAM_ARN != '<<ADD WORK TEAM ARN HERE>>')
# * The IAM execution role you used to create this notebook instance must have the following permissions:
# * AWS managed policy [AmazonSageMakerGroundTruthExecution](https://console.aws.amazon.com/iam/home#policies/arn:aws:iam::aws:policy/AmazonSageMakerGroundTruthExecution). Run the following code-block to see your IAM execution role name. This [GIF](add-policy.gif) demonstrates how to add this policy to an IAM role in the IAM console. You can also find instructions in the IAM User Guide: [Adding and removing IAM identity permissions](https://docs.aws.amazon.com/IAM/latest/UserGuide/access_policies_manage-attach-detach.html#add-policies-console).
# * When you create your role, you specify Amazon S3 permissions. Make sure that your IAM role has access to the S3 bucket that you plan to use in this example. If you do not specify an S3 bucket in this notebook, the default bucket in the AWS region you are running this notebook instance will be used. If you do not require granular permissions, you can attach [AmazonS3FullAccess](https://console.aws.amazon.com/iam/home#policies/arn:aws:iam::aws:policy/AmazonS3FullAccess) to your role.
role = sagemaker.get_execution_role()
role_name = role.split('/')[-1]
print('IMPORTANT: Make sure this execution role has the AWS Managed policy AmazonGroundTruthExecution attached.')
print('********************************************************************************')
print('The IAM execution role name:', role_name)
print('The IAM execution role ARN:', role)
print('********************************************************************************')
sess = sagemaker.Session()
BUCKET = '<< YOUR S3 BUCKET NAME >>'
if(BUCKET=='<< YOUR S3 BUCKET NAME >>'):
BUCKET=sess.default_bucket()
region = boto3.session.Session().region_name
s3 = boto3.client('s3')
# Make sure the bucket is in the same region as this notebook.
bucket_region = s3.head_bucket(Bucket=BUCKET)['ResponseMetadata']['HTTPHeaders']['x-amz-bucket-region']
assert bucket_region == region, f'Your S3 bucket {BUCKET} and this notebook need to be in the same region.'
print(f'IMPORTANT: make sure the role {role_name} has the access to read and write to this bucket.')
print('********************************************************************************************************')
print(f'This notebook will use the following S3 bucket: {BUCKET}')
print('********************************************************************************************************')
# ## Create SNS Topics for Input and Output Data
# You can send data objects to your streaming labeling job using Amazon Simple Notification Service (Amazon SNS). Amazon SNS is a web service that coordinates and manages the delivery of messages to and from endpoints (for example, an email address or AWS Lambda function). An Amazon SNS topic acts as a communication channel between two or more endpoints. You use Amazon SNS to send, or publish, new data objects to the topic specified in the CreateLabelingJob parameter SnsTopicArn in InputConfig.
#
# The following cells will create a name for your labeling job and use this name to create Amazon SNS input and output topics. This labeling job name and these topics will be used in your `CreateLabelingJob` request later in this notebook.
# Job Name
LABELING_JOB_NAME = 'GroundTruth-streaming-' + str(int(time.time()))
print('Your labeling job name will be :', LABELING_JOB_NAME)
# +
# Make sure role has "Sns:CreateTopic" access
sns = boto3.client('sns')
# Create Input Topic
input_response = sns.create_topic(Name= LABELING_JOB_NAME + '-Input')
INPUT_SNS_TOPIC_ARN = input_response['TopicArn']
print('input_sns_topic :', INPUT_SNS_TOPIC_ARN)
# Create Output Topic
output_response = sns.create_topic(Name= LABELING_JOB_NAME + '-Output')
OUTPUT_SNS_TOPIC_ARN = output_response['TopicArn']
print('output_sns_topic :', OUTPUT_SNS_TOPIC_ARN)
# -
# ## Choose Labeling Job Type
#
# Ground Truth supports a variety of built-in task types which streamline the process of creating image, text, video, video frame, and 3D point cloud labeling jobs. You can use this notebook on *default* mode if you do not want to bring your own input data.
#
# If you have input data and an input manifest file in an S3 bucket, set `DEFAULT` to `False` and, optionally, choose the **Labeling Job Task Type** you want to use below. To learn more about each task type, see [Built-in Task Types](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-task-types.html).
# ### Choose Labeling Job Built-In Task Type
#
# Copy one of the following task types and use it to set the value for `task_type`. If you set **`DEFAULT`** to `True`, at the beginning of this notebook, the image bounding box task type will be used by default.
# +
## Choose from following:
## Bounding Box
## Image Classification (Single Label)
## Image Classification (Multi-label)
## Image Semantic Segmentation
## Text Classification (Single Label)
## Text Classification (Multi-label)
## Named Entity Recognition
## Video Classification
## Video Frame Object Detection
## Video Frame Object Tracking
## 3D Point Cloud Object Detection
## 3D Point Cloud Object Detection
## 3D Point Cloud Semantic Segmentation
task_type = "<<COPY AND PASTE TASK TYPE FROM LIST ABOVE>>"
if(DEFAULT):
task_type = "Bounding Box"
print(f'Your task type: {task_type}')
# -
# The following cells will configure the lambda functions Ground Truth uses to pre-process your input data and output data. These cells will configure your [PreHumanTaskLambdaArn](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_HumanTaskConfig.html#sagemaker-Type-HumanTaskConfig-PreHumanTaskLambdaArn) and [AnnotationConsolidationLambdaArn](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_AnnotationConsolidationConfig.html#sagemaker-Type-AnnotationConsolidationConfig-AnnotationConsolidationLambdaArn).
# +
task_type_map = {
"Bounding Box" : "BoundingBox",
"Image Classification (Single Label)" : "ImageMultiClass",
"Image Classification (Multi-label)" : "ImageMultiClassMultiLabel",
"Image Semantic Segmentation" : "SemanticSegmentation",
"Text Classification (Single Label)" : "TextMultiClass",
"Text Classification (Multi-label)" : "TextMultiClassMultiLabel",
"Named Entity Recognition" : "NamedEntityRecognition",
"Video Classification" : "VideoMultiClass",
"Video Frame Object Detection" : "VideoObjectDetection",
"Video Frame Object Tracking" : "VideoObjectTracking",
"3D Point Cloud Object Detection" : "3DPointCloudObjectDetection",
"3D Point Cloud Object Tracking" : "3DPointCloudObjectTracking",
"3D Point Cloud Semantic Segmentation" : "3DPointCloudSemanticSegmentation"
}
arn_region_map = {'us-west-2': '081040173940',
'us-east-1': '432418664414',
'us-east-2': '266458841044',
'eu-west-1': '568282634449',
'eu-west-2': '487402164563',
'ap-northeast-1': '477331159723',
'ap-northeast-2': '845288260483',
'ca-central-1': '918755190332',
'eu-central-1': '203001061592',
'ap-south-1': '565803892007',
'ap-southeast-1': '377565633583',
'ap-southeast-2': '454466003867'
}
# -
task_type_suffix = task_type_map[task_type]
region_account = arn_region_map[region]
PRE_HUMAN_TASK_LAMBDA = f'arn:aws:lambda:{region}:{region_account}:function:PRE-{task_type_suffix}'
POST_ANNOTATION_LAMBDA = f'arn:aws:lambda:{region}:{region_account}:function:ACS-{task_type_suffix}'
print(PRE_HUMAN_TASK_LAMBDA)
print(POST_ANNOTATION_LAMBDA)
# 3D point cloud and video frame task types have special requirements. The following variables will be used to configure your labeling job for these task types. To learn more, see the following topics in the documentation:
# * [3D Point Cloud Labeling Jobs Overview](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-point-cloud-general-information.html)
# * [Video Frame Labeling Job Overview](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-video-overview.html)
point_cloud_task = re.search(r'Point Cloud', task_type) is not None
video_frame_task = re.search(r'Video Frame', task_type) is not None
# ### Create Custom Labeling Workflow
#
# If you want to create a custom labeling workflow, you can create your own lambda functions to pre-process your input data and post-process the labels returned from workers. To learn more, see [Step 3: Processing with AWS Lambda](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-custom-templates-step3.html).
#
# To use this notebook to run a custom flow, set `CUSTOM` to `True` and specify your pre- and post-processing lambdas below.
CUSTOM = False
if(CUSTOM):
PRE_HUMAN_TASK_LAMBDA = '<ADD-PRE-PROCESSING-LABMDA-ARN>'
POST_ANNOTATION_LAMBDA = '<ADD-POST-PROCESSING-LABMDA-ARN>'
# ## Specify Labels
#
# You specify the labels that you want workers to use to annotate your data in a label category configuration file. When you create a 3D point cloud or video frame labeling job, you can add label category attributes to your labeling category configruation file. Workers can assign one or more attributes to annotations to give more information about that object.
#
# For all task types, you can use the following cell to identify the labels you use for your labeling job. To create a label category configuration file with label category attributes, see [Create a Labeling Category Configuration File with Label Category Attributes
# ](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-label-cat-config-attributes.html) in the Amazon SageMaker developer guide.
# Add label categories of your choice
LABEL_CATEGORIES = []
if(DEFAULT):
LABEL_CATEGORIES = ['Pedestrian', 'Street Car', 'Biker']
# The following cell will create a label category configuration file using the labels specified above.
#
# **IMPORTANT**: Make sure you have added label categories above and they appear under `labels` when you run the following cell.
# +
# Specify labels and this notebook will upload and a label category configuration file to S3.
json_body = {
"document-version": "2018-11-28",
'labels': [{'label': label} for label in LABEL_CATEGORIES]
}
with open('class_labels.json', 'w') as f:
json.dump(json_body, f)
print("Your label category configuration file:")
print("\n",json.dumps(json_body, indent=2))
# -
s3.upload_file('class_labels.json', BUCKET, 'class_labels.json')
LABEL_CATEGORIES_S3_URI = f's3://{BUCKET}/class_labels.json'
print(f'You should now see class_labels.json in {LABEL_CATEGORIES_S3_URI}')
# ## Create A Worker Task Template
#
# Part or all of your images will be annotated by human annotators. It is essential to provide good instructions. Good instructions are:
#
# 1. Concise. We recommend limiting verbal/textual instruction to two sentences and focusing on clear visuals.
# 2. Visual. In the case of object detection, we recommend providing several labeled examples with different numbers of boxes.
# 3. When used through the AWS Console, Ground Truth helps you create the instructions using a visual wizard. When using the API, you need to create an HTML template for your instructions.
#
# NOTE: If you use any images in your template (as we do), they need to be publicly accessible. You can enable public access to files in your S3 bucket through the S3 Console, as described in S3 Documentation.
#
# ### Specify Resources Used for Human Task UI
#
# The human task user interface (UI) is the interface that human workers use to label your data. Depending on the type of labeling job you create, you will specify a resource that is used to generate the human task UI in the `UiConfig` parameter of `CreateLabelingJob`.
#
# For 3D point cloud and video frame labeling tasks, you will specify a pre-defined `HumanTaskUiARN`. For all other labeling job task types, you will specify a `UiTemplateS3Uri`.
# #### Bounding Box Image Labeling Job (Default)
#
# If you set `DEFAULT` to `True`, use the following to create a worker task template and upload it to your S3 bucket. Ground Trust uses this template to generate your human task UI.
# +
from IPython.core.display import HTML, display
def make_template(save_fname='instructions.template'):
template = r"""<script src="https://assets.crowd.aws/crowd-html-elements.js"></script>
<crowd-form>
<crowd-bounding-box
name="boundingBox"
src="{{{{ task.input.taskObject | grant_read_access }}}}"
header="Dear Annotator, please draw a tight box around each object you see (if there are more than 8 objects, draw boxes around at least 8)."
labels="{{{{ task.input.labels | to_json | escape }}}}"
>
<full-instructions header="Please annotate each object">
<ol>
<li><strong>Inspect</strong> the image</li>
<li><strong>Determine</strong> if the specified label is/are visible in the picture.</li>
<li><strong>Outline</strong> each instance of the specified label in the image using the provided “Box” tool.</li>
</ol>
</full-instructions>
<short-instructions>
<ul>
<li>Boxes should fit tightly around each object</li>
<li>Do not include parts of the object are overlapping or that cannot be seen, even though you think you can interpolate the whole shape.</li>
<li>Avoid including shadows.</li>
<li>If the target is off screen, draw the box up to the edge of the image.</li>
</ul>
</short-instructions>
</crowd-bounding-box>
</crowd-form>
""".format()
with open(save_fname, 'w') as f:
f.write(template)
if(DEFAULT):
make_template(save_fname='instructions.template')
# -
if(DEFAULT):
result = s3.upload_file('instructions.template', BUCKET, 'instructions.template')
# #### Image, Text, and Custom Labeling Jobs (Non Default)
#
# For all image and text based built-in task types, you can find a sample worker task template on that task type page. Find the page for your task type on [Built-in Task Types](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-task-types.html). You will see an example template under the section **Create a {Insert-Task-Type} Job (API)**.
#
# Update `<full-instructions></full-instructions>` and `<short-instructions></short-instructions>`. Add your template to the following code block and run the code blocks below to generate your worker task template and upload it to your S3 bucket.
#
# For custom labeling workflows, you can provide a custom HTML worker task template using Crowd HTML Elements. To learn more, see [Step 2: Creating your custom labeling task template](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-custom-templates-step2.html).
#
# Ground Trust uses this template to generate your human task UI.
# **Important**: If you use the following `make_template` function to create and upload a worker task template to Amazon S3, you must add an extra pair of `{}` brackets around each Liquid element. For example, if the template contains `{{ task.input.labels | to_json | escape }}`, this line should look as follows in the `make_template` variable `template`: `{{{{ task.input.labels | to_json | escape }}}}`.
# +
from IPython.core.display import HTML, display
def make_template(save_fname='instructions.template'):
template = r"""
<<<ADD-TEMPLATE-HTML-CODE-HERE>>>
""".format()
with open(save_fname, 'w') as f:
f.write(template)
#This will upload your template to S3 if you are not running on DEFAULT mode, and if your take type
#does not use video frames or 3D point clouds.
if(not DEFAULT and not video_frame_task and not point_cloud_task):
make_template(save_fname='instructions.html')
s3.upload_file('instructions.template', BUCKET, 'instructions.template')
# -
# #### 3D Point Cloud and Video Frame Task Types
#
# If you are creating a 3D point cloud or video frame task type, your worker UI is configured by Ground Truth. If you chose one of these task types above, the following cell will specify the correct template.
# +
import re
if(not DEFAULT):
if (point_cloud_task):
task_type_suffix_humanuiarn = task_type_suffix.split('3D')[-1]
HUMAN_UI_ARN = f'arn:aws:sagemaker:{region}:394669845002:human-task-ui/{task_type_suffix_humanuiarn}'
if (video_frame_task):
HUMAN_UI_ARN = f'arn:aws:sagemaker:{region}:394669845002:human-task-ui/{task_type_suffix}'
print(f'The Human Task UI ARN is: {HUMAN_UI_ARN}')
# -
# ## (Optional) Create an Input Manifest File
#
# You can optionally specify an input manifest file Amazon S3 URI in ManifestS3Uri when you create the streaming labeling job. Ground Truth sends each data object in the manifest file to workers for labeling as soon as the labeling job starts.
#
# Each line in an input manifest file is an entry containing an object, or a reference to an object, to label. An entry can also contain labels from previous jobs and for some task types, additional information.
#
# To learn how to create an input manifest file, see [Use an Input Manifest File](https://docs.aws.amazon.com/sagemaker/latest/dg/sms-input-data-input-manifest.html). Copy the S3 URI of the file below.
# [Optional] The path in Amazon S3 to your input manifest file.
INPUT_MANIFEST = ''
# ## Specify Parameters for Labeling Job
#
# If you set `DEFAULT` to `False`, you must specify the following parameters. These will be used to configure and create your lableing job. If you set `DEFAULT` to `True`, default parameters will be used.
#
# To learn more about these parameters, use the following documentation:
# * [TaskTitle](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_HumanTaskConfig.html#sagemaker-Type-HumanTaskConfig-TaskTitle)
# * [TaskDescription](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_HumanTaskConfig.html#sagemaker-Type-HumanTaskConfig-TaskDescription)
# * [TaskKeywords](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_HumanTaskConfig.html#sagemaker-Type-HumanTaskConfig-TaskKeywords)
# +
TASK_TITLE = '<<ADD-TASK-TITLE>>'
if(DEFAULT):
TASK_TITLE = 'Add bounding boxes to detect objects in an image'
TASK_DESCRIPTION = '<<ADD-TASK-DESCRIPTION>>'
if(DEFAULT):
TASK_DESCRIPTION = 'Categorize images into classes using bounding boxes'
# Keywords for your task, in a string-array. ex) ['image classification', 'image dataset']
TASK_KEYWORDS = ['<<ADD-KEYWODS>>']
if(DEFAULT):
TASK_KEYWORDS = ['bounding box', 'image dataset']
# -
# Run the following to specify the rest of the parameters required to configure your labeling job.
# +
# The path in Amazon S3 to your worker task template or human task UI
HUMAN_UI = []
if(point_cloud_task or video_frame_task):
HUMAN_TASK_UI_ARN = HUMAN_UI_ARN
HUMAN_UI.append(HUMAN_TASK_UI_ARN)
UI_CONFIG_PARAM = 'HumanTaskUiArn'
else:
UI_TEMPLATE_S3_URI = f's3://{BUCKET}/instructions.template'
HUMAN_UI.append(UI_TEMPLATE_S3_URI)
UI_CONFIG_PARAM = 'UiTemplateS3Uri'
print(f'{UI_CONFIG_PARAM} resource that will be used: {HUMAN_UI[0]}')
# +
# If you want to store your output manifest in a different folder, provide an OUTPUT_PATH.
OUTPUT_FOLDER_PREFIX = '/gt-streaming-demo-output'
OUTPUT_BUCKET = 's3://' + BUCKET + OUTPUT_FOLDER_PREFIX
print("Your output data will be stored in:", OUTPUT_BUCKET)
# An IAM role with AmazonGroundTruthExecution policies attached.
# This must be the same role that you used to create this notebook instance.
ROLE_ARN = role
# -
# ## Use the CreateLabelingJob API to create a streaming labeling job
# +
if(re.search(r'Semantic Segmentation', task_type) is not None or re.match(r'Object Tracking', task_type) is not None or video_frame_task):
LABEL_ATTRIBUTE_NAME = LABELING_JOB_NAME + '-ref'
else:
LABEL_ATTRIBUTE_NAME = LABELING_JOB_NAME
human_task_config = {
"PreHumanTaskLambdaArn": PRE_HUMAN_TASK_LAMBDA,
"MaxConcurrentTaskCount": 100, # Maximum of 100 objects will be available to the workteam at any time
"NumberOfHumanWorkersPerDataObject": 1, # We will obtain and consolidate 1 human annotationsfor each image.
"TaskAvailabilityLifetimeInSeconds": 21600, # Your workteam has 6 hours to complete all pending tasks.
"TaskDescription": TASK_DESCRIPTION,
# If using public workforce, specify "PublicWorkforceTaskPrice"
"WorkteamArn": WORKTEAM_ARN,
"AnnotationConsolidationConfig": {
"AnnotationConsolidationLambdaArn": POST_ANNOTATION_LAMBDA
},
"TaskKeywords": TASK_KEYWORDS,
"TaskTimeLimitInSeconds": 600, # Each image must be labeled within 10 minutes.
"TaskTitle": TASK_TITLE,
"UiConfig": {
UI_CONFIG_PARAM : HUMAN_UI[0]
}
}
#if you are using the Amazon Mechanical Turk workforce, specify the amount you want to pay a
#worker to label a data object. See https://aws.amazon.com/sagemaker/groundtruth/pricing/ for recommendations.
if (not private_work_team):
human_task_config["PublicWorkforceTaskPrice"] = {
"AmountInUsd": {
"Dollars": 0,
"Cents": 3,
"TenthFractionsOfACent": 6,
}
}
human_task_config["WorkteamArn"] = WORKTEAM_ARN
else:
human_task_config["WorkteamArn"] = WORKTEAM_ARN
ground_truth_request = {
"InputConfig": {
"DataSource": {
"SnsDataSource": {
"SnsTopicArn": INPUT_SNS_TOPIC_ARN
}
}
},
"HumanTaskConfig" : human_task_config,
"LabelAttributeName": LABEL_ATTRIBUTE_NAME,
"LabelCategoryConfigS3Uri" : LABEL_CATEGORIES_S3_URI,
"LabelingJobName": LABELING_JOB_NAME,
"OutputConfig": {
"S3OutputPath": OUTPUT_BUCKET,
"SnsTopicArn": OUTPUT_SNS_TOPIC_ARN
},
"RoleArn": ROLE_ARN
}
if(INPUT_MANIFEST is not ''):
ground_truth_request["InputConfig"]["DataSource"]["S3DataSource"] = {"ManifestS3Uri": INPUT_MANIFEST}
# -
# #### DataAttributes
# You should not share explicit, confidential, or personal information or protected health information with the Amazon Mechanical Turk workforce.
#
# If you are using Amazon Mechanical Turk workforce, you must verify that your data is free of personal, confidential, and explicit content and protected health information using this code cell.
if (not private_work_team):
ground_truth_request["InputConfig"]["DataAttributes"]={"ContentClassifiers": ["FreeOfPersonallyIdentifiableInformation","FreeOfAdultContent"]}
print("Your create labeling job request:\n",json.dumps(ground_truth_request,indent=4))
sagemaker_client = boto3.client('sagemaker')
sagemaker_client.create_labeling_job(**ground_truth_request)
# ## Use the DescribeLabelingJob API to describe a streaming labeling job
sagemaker_client.describe_labeling_job(LabelingJobName=LABELING_JOB_NAME)
# ### Wait until the labeling job status equals `InProgress` before moving forward in this notebook
sagemaker_client.describe_labeling_job(LabelingJobName=LABELING_JOB_NAME)['LabelingJobStatus']
# ## Check for LabelingJobStatus and interpreting describe response
#
# * If you specified "S3DataSource.ManifestS3Uri" in the above request, the objects in the S3 file will automatically make their way to the labeling job. You will see counters incrementing from the objects from the file.
# * Streaming jobs create a SQS queue in your account. You can check for existence of the queue by name "GroundTruth-LABELING_JOB_NAME" via console or through below command
sqs = boto3.client('sqs')
response = sqs.get_queue_url(QueueName='GroundTruth-' + LABELING_JOB_NAME.lower())
print("Queue url is :", response['QueueUrl'])
# ## Publish a new object to your labeling job once it has started
#
# Once you start a labeling job, you an publish a new request to it using Amazon SNS.
#
# ### Configure your Request
#
# You will need to specify `REQUEST` in the following format:
#
# **For non-text objects**
#
# First, make sure that your object is located in `s3_bucket_location`
#
# `{"source-ref": "s3_bucket_location"}`
#
# **For text objects**
#
# `{"source": "Lorem ipsum dolor sit amet"}`
#
# Modify one of these examples to specify your request in the next cell.
REQUEST = '<Populate your object as shown above>'
# If you set `Default` to `True` use the following cell upload a sample-image to your S3 bucket and send that image to labeling job.
if(DEFAULT):
# !wget https://aws-ml-blog.s3.amazonaws.com/artifacts/gt-labeling-job-resources/example-image.jpg
s3.upload_file('example-image.jpg', BUCKET, 'example-image.jpg')
REQUEST = str({"source-ref": f"s3://{BUCKET}/example-image.jpg"})
print(f'Your request: {REQUEST}')
# ### Publish Your Request
#
# First, check the `LabelCounters` variable for your labeling job using `DescribeLabelingJob`. After you publish your request, you'll see `Unlabeled` increases to `1` (or the number of objects you send to your labeling job).
sagemaker_client.describe_labeling_job(LabelingJobName=LABELING_JOB_NAME)['LabelCounters']
# The following will publish your request to your Amazon SNS input topic.
print(f'Your Request: {REQUEST}\n')
if(REQUEST != '<Populate your object as shown above>'):
published_message = sns.publish(TopicArn=INPUT_SNS_TOPIC_ARN,Message=REQUEST)
print(f'Published Message: {published_message}')
# You may need to wait 1 to 2 minutes for your request to appear in `LabelCounters`.
sagemaker_client.describe_labeling_job(LabelingJobName=LABELING_JOB_NAME)['LabelCounters']
# ## Call StopLabelingJob for your previously launched job
#
# To stop your Streaming job, call StopLabelingJob with the `LABELING_JOB_NAME`.
#
#
sagemaker_client.stop_labeling_job(LabelingJobName=LABELING_JOB_NAME)
| ground_truth_labeling_jobs/ground_truth_streaming_labeling_jobs/ground_truth_create_streaming_labeling_job.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import csv
import pandas as pd
from sklearn import svm
from nltk.corpus import stopwords
import re
import nltk
from nltk.tokenize import *
from nltk.data import load
# ### Reading Files Line by Line
def sentenceExtractionForTraining(dirName, fileName, classes):
sentencesClass = []
for i in range(0,len(classes)):
sentences = readFile(dirName+fileName[i])
sentencesClass.append(sentences)
return sentencesClass
def readFile(fileName):
f = open(fileName,"r+")
sentences = []
for line in f.readlines():
line = line.strip()
if len(line)==0 :
continue
if "TIMEX3" in line:
continue
sentences.append(line.lower().rstrip())
f.close()
return sentences
# ### Removing non word characters
#
def removeFancyChars(sentences):
lengthPhrase = len(sentences)
for i in range(lengthPhrase):
sentences[i] = re.sub(r'([^\s\w]|_)+', '', sentences[i])
return sentences
def removeFC(sentencesClass):
for i in range(0, len(sentencesClass)):
sentencesClass[i] = removeFancyChars(sentencesClass[i])
return sentencesClass
# ### Parts of Speech
def POSForSentence(sentence):
text = word_tokenize(sentence)
posSentence = nltk.pos_tag(text)
posSentence = [y for x, y in posSentence]
return posSentence
def getUniquePOS():
tagdict = load('help/tagsets/upenn_tagset.pickle')
return len(tagdict), tagdict.keys()
# ### Removing StopWords
def removeSWFromSent(sentence):
words = sentence.split()
sentence = []
for w in words:
if w not in stopwords.words('english'):
sentence.append(w)
if len(sentence) == 0:
sentence = [""]
return convertlistToString(sentence)
def removeSWFromPar(sentences):
sents = []
for i in range(0, len(sentences)):
sent = removeSWFromSent(sentences[i])
sents.append(sent)
return sents
def removeSWFromClass(sentencesClass):
sentClass = []
for i in range(0, len(sentencesClass)):
sentClass.append(removeSWFromPar(sentencesClass[i]))
return sentClass
def convertlistToString(sentence):
sentence = " ".join(sentence)
return sentence
# ### Count unique words
def uniqueWordsCount(sentencesClass):
uniqueWords = set()
for i in range(0,len(sentencesClass)):
for j in range(0,len(sentencesClass[i])):
words = sentencesClass[i][j].split()
for k in words:
uniqueWords.add(k)
return list(uniqueWords)
# ### Bag of words
def totalSentences(sentencesClass):
size = 0
for i in range(0, len(sentencesClass)):
size += len(sentencesClass[i])
return size;
def defineBOWM(sentencesClass, vocab):
vocabSize = len(vocab)
n = totalSentences(sentencesClass)
labels = np.zeros(n)
mat = np.zeros((n,vocabSize))
k = 0
for i in range(0, len(sentencesClass)):
for j in range(0, len(sentencesClass[i])):
words = sentencesClass[i][j].split()
for w in words:
mat[k, vocab.index(w)] = 1.0
labels[k] = i+1
k+=1
return mat, labels
def defineBOWMPOS(originalSentencesClass, sentencesClass, vocab):
vocabSize = len(vocab)
n = totalSentences(sentencesClass)
labels = np.zeros(n)
sizePOS, POSList = getUniquePOS()
mat = np.zeros((n, vocabSize + sizePOS))
matFromBOWM, labels = defineBOWM(sentencesClass, vocab)
for i in range(0, n):
mat[i,:vocabSize] = matFromBOWM[i]
k = 0
for i in range(0, len(originalSentencesClass)):
for j in range(0, len(originalSentencesClass[i])):
pos = POSForSentence(originalSentencesClass[i][j])
for p in pos:
mat[k, vocabSize + POSList.index(p)] = 1.0
k+=1
return mat, labels
# ### Save features
def finalFeaturesLabel(X,y):
n, d = X.shape
finalMat = np.zeros((n,d+1))
for i in range(0, n):
finalMat[i, 0] = y[i]
finalMat[i, 1:] = X[i]
return finalMat
def saveBOW(fileName, finalMat):
np.save(fileName, finalMat)
def saveVocab(fileName, vocab):
np.save(fileName, vocab)
# ## Main
def main():
dirName = "Email-classification_dataset/"
classes = [1,2,3,4,5]
fileName = ["RD-positive-800.txt", "meetings-positive-800.txt", "negative-800.txt", "fyi-positive-800.txt", "tp-positive-500.txt",]
removeStopWords = True
originalsentencesClass = sentenceExtractionForTraining(dirName, fileName, classes)
sentencesClass = removeFC(originalsentencesClass)
vocab = uniqueWordsCount(sentencesClass)
XBOW, yBOW = defineBOWM(sentencesClass, vocab)
finalMatBOW = finalFeaturesLabel(XBOW, yBOW)
saveBOW("bow.npy", finalMatBOW)
saveVocab("vocab.npy", vocab)
sentencesClassWOSW = removeSWFromClass(sentencesClass)
vocabWOSW = uniqueWordsCount(sentencesClassWOSW)
XBOWWOSW, yBOWWOSW = defineBOWM(sentencesClassWOSW, vocabWOSW)
finalMatBOWWOSW = finalFeaturesLabel(XBOWWOSW, yBOWWOSW)
saveBOW("bowsw.npy", finalMatBOWWOSW)
saveVocab("vocabsw.npy", vocabWOSW)
XBOWPOS, yBOWPOS = defineBOWMPOS(originalsentencesClass, sentencesClass, vocab)
finalMatBOWPOS = finalFeaturesLabel(XBOWPOS, yBOWPOS)
saveBOW("bowpos.npy", finalMatBOWPOS)
XBOWPOSSW, yBOWPOSSW = defineBOWMPOS(originalsentencesClass, sentencesClassWOSW, vocabWOSW)
finalMatBOWPOSSW = finalFeaturesLabel(XBOWPOSSW, yBOWPOSSW)
saveBOW("bowpossw.npy", finalMatBOWPOSSW)
if __name__=="__main__":
main()
| PRNN/Assignment/Assignment3/BOW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# HIDDEN
from datascience import *
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import math
import numpy as np
from scipy import stats
import ipywidgets as widgets
import nbinteract as nbi
# ### The Central Limit Theorem ###
# Very few of the data histograms that we have seen in this course have been bell shaped. When we have come across a bell shaped distribution, it has almost invariably been an empirical histogram of a statistic based on a random sample.
# **The Central Limit Theorem says that the probability distribution of the sum or average of a large random sample drawn with replacement will be roughly normal, *regardless of the distribution of the population from which the sample is drawn*.**
#
# As we noted when we were studying Chebychev's bounds, results that can be applied to random samples *regardless of the distribution of the population* are very powerful, because in data science we rarely know the distribution of the population.
#
# The Central Limit Theorem makes it possible to make inferences with very little knowledge about the population, provided we have a large random sample. That is why it is central to the field of statistical inference.
# ### Proportion of Purple Flowers ###
# Recall Mendel's probability model for the colors of the flowers of a species of pea plant. The model says that the flower colors of the plants are like draws made at random with replacement from {Purple, Purple, Purple, White}.
#
# In a large sample of plants, about what proportion will have purple flowers? We would expect the answer to be about 0.75, the proportion purple in the model. And, because proportions are means, the Central Limit Theorem says that the distribution of the sample proportion of purple plants is roughly normal.
#
# We can confirm this by simulation. Let's simulate the proportion of purple-flowered plants in a sample of 200 plants.
# +
colors = make_array('Purple', 'Purple', 'Purple', 'White')
model = Table().with_column('Color', colors)
model
# +
props = make_array()
num_plants = 200
repetitions = 1000
for i in np.arange(repetitions):
sample = model.sample(num_plants)
new_prop = np.count_nonzero(sample.column('Color') == 'Purple')/num_plants
props = np.append(props, new_prop)
props[:5]
# -
opts = {
'title': 'Distribution of sample proportions',
'xlabel': 'Sample Proportion',
'ylabel': 'Percent per unit',
'xlim': (0.64, 0.84),
'ylim': (0, 25),
'bins': 20,
}
nbi.hist(props, options=opts)
# There's that normal curve again, as predicted by the Central Limit Theorem, centered at around 0.75 just as you would expect.
#
# How would this distribution change if we increased the sample size? We can copy our sampling code into a function and then use interaction to see how the distribution changes as the sample size increases.
#
# We will keep the number of `repetitions` the same as before so that the two columns have the same length.
def empirical_props(num_plants):
props = make_array()
for i in np.arange(repetitions):
sample = model.sample(num_plants)
new_prop = np.count_nonzero(sample.column('Color') == 'Purple')/num_plants
props = np.append(props, new_prop)
return props
nbi.hist(empirical_props, options=opts,
num_plants=widgets.ToggleButtons(options=[100, 200, 400, 800]))
# All of the above distributions are approximately normal but become more narrow as the sample size increases. For example, the proportions based on a sample size of 800 are more tightly clustered around 0.75 than those from a sample size of 200. Increasing the sample size has decreased the variability in the sample proportion.
| packages/nbinteract-core/example-notebooks/examples_central_limit_theorem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Modules
# +
import warnings
warnings.filterwarnings('ignore')
from src import detect_faces, show_bboxes
from PIL import Image
import torch
from torchvision import transforms, datasets
import numpy as np
import os
# -
# # Path Definition
dataset_path = '../Dataset/emotiw/'
face_coordinates_directory = '../Dataset/FaceCoordinates/'
processed_dataset_path = '../Dataset/CroppedFaces/'
# # Load Train and Val Dataset
# +
image_datasets = {x : datasets.ImageFolder(os.path.join(dataset_path, x))
for x in ['train', 'val']}
class_names = image_datasets['train'].classes
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# -
class_names
training_dataset = image_datasets['train']
validation_dataset = image_datasets['val']
# +
neg_train = sorted(os.listdir(dataset_path + 'train/Negative/'))
neu_train = sorted(os.listdir(dataset_path + 'train/Neutral/'))
pos_train = sorted(os.listdir(dataset_path + 'train/Positive/'))
neg_val = sorted(os.listdir(dataset_path + 'val/Negative/'))
neu_val = sorted(os.listdir(dataset_path + 'val/Neutral/'))
pos_val = sorted(os.listdir(dataset_path + 'val/Positive/'))
# +
neg_train_filelist = [x.split('.')[0] for x in neg_train]
neu_train_filelist = [x.split('.')[0] for x in neu_train]
pos_train_filelist = [x.split('.')[0] for x in pos_train]
neg_val_filelist = [x.split('.')[0] for x in neg_val]
neu_val_filelist = [x.split('.')[0] for x in neu_val]
pos_val_filelist = [x.split('.')[0] for x in pos_val]
# +
print(neg_train_filelist[:10])
print(neu_train_filelist[:10])
print(pos_train_filelist[:10])
print(neg_val_filelist[:10])
print(neu_val_filelist[:10])
print(pos_val_filelist[:10])
# -
train_filelist = neg_train_filelist + neu_train_filelist + pos_train_filelist
val_filelist = neg_val_filelist + neu_val_filelist + pos_val_filelist
print(len(training_dataset))
print(len(validation_dataset))
# # Crop Faces
for i in range(len(training_dataset)):
try:
image, label = training_dataset[i]
face_list = []
landmarks_new_coordinates = []
if label == 0:
if os.path.isfile(processed_dataset_path + 'train/Negative/' + train_filelist[i] + '.npz'):
print(train_filelist[i] + ' Already present')
continue
bbox_lm = np.load(face_coordinates_directory + 'train/Negative/' + train_filelist[i] +'.npz')
bounding_boxes = bbox_lm['a']
if bounding_boxes.size == 0 or (bounding_boxes[0] == 0).all():
print("No bounding boxes for " + train_filelist[i] + ". Adding empty file for the same")
np.savez(processed_dataset_path + 'train/Negative/' + train_filelist[i], a = np.zeros(1), b = np.zeros(1))
continue
landmarks = bbox_lm['b']
for j in range(len(bounding_boxes)):
bbox_coordinates = bounding_boxes[j]
landmark = landmarks[j]
img_face = image.crop((bbox_coordinates[0], bbox_coordinates[1], bbox_coordinates[2], bbox_coordinates[3]))
x = bbox_coordinates[0]
y = bbox_coordinates[1]
for k in range(5):
landmark[k] -= x
landmark[k+5] -= y
img_face = np.array(img_face)
landmark = np.array(landmark)
if len(face_list) != 0:
if img_face.shape[0] == face_list[-1].shape[0]:
img_face = image.crop((bbox_coordinates[0] - 1, bbox_coordinates[1] - 1, bbox_coordinates[2], bbox_coordinates[3]))
img_face = np.array(img_face)
landmark +=1
face_list.append(img_face)
landmarks_new_coordinates.append(landmark)
face_list = np.asarray(face_list)
landmarks_new_coordinates = np.asarray(landmarks_new_coordinates)
np.savez(processed_dataset_path + 'train/Negative/' + train_filelist[i], a = face_list, b = landmarks_new_coordinates)
elif label == 1:
if os.path.isfile(processed_dataset_path + 'train/Neutral/' + train_filelist[i] + '.npz'):
print(train_filelist[i] + ' Already present')
continue
bbox_lm = np.load(face_coordinates_directory + 'train/Neutral/' + train_filelist[i] +'.npz')
bounding_boxes = bbox_lm['a']
if bounding_boxes.size == 0 or (bounding_boxes[0] == 0).all():
print("No bounding boxes for " + train_filelist[i] + ". Adding empty file for the same")
np.savez(processed_dataset_path + 'train/Neutral/' + train_filelist[i], a = np.zeros(1), b = np.zeros(1))
continue
landmarks = bbox_lm['b']
for j in range(len(bounding_boxes)):
bbox_coordinates = bounding_boxes[j]
landmark = landmarks[j]
img_face = image.crop((bbox_coordinates[0], bbox_coordinates[1], bbox_coordinates[2], bbox_coordinates[3]))
x = bbox_coordinates[0]
y = bbox_coordinates[1]
for k in range(5):
landmark[k] -= x
landmark[k+5] -= y
img_face = np.array(img_face)
landmark = np.array(landmark)
if len(face_list) != 0:
if img_face.shape[0] == face_list[-1].shape[0]:
img_face = image.crop((bbox_coordinates[0] - 1, bbox_coordinates[1] - 1, bbox_coordinates[2], bbox_coordinates[3]))
img_face = np.array(img_face)
landmark += 1
face_list.append(img_face)
landmarks_new_coordinates.append(landmark)
face_list = np.asarray(face_list)
landmarks_new_coordinates = np.asarray(landmarks_new_coordinates)
np.savez(processed_dataset_path + 'train/Neutral/' + train_filelist[i], a = face_list, b = landmarks_new_coordinates)
else:
if os.path.isfile(processed_dataset_path + 'train/Positive/' + train_filelist[i] + '.npz'):
print(train_filelist[i] + ' Already present')
continue
bbox_lm = np.load(face_coordinates_directory + 'train/Positive/' + train_filelist[i] +'.npz')
bounding_boxes = bbox_lm['a']
if bounding_boxes.size == 0 or (bounding_boxes[0] == 0).all():
print("No bounding boxes for " + train_filelist[i] + ". Adding empty file for the same")
np.savez(processed_dataset_path + 'train/Positive/' + train_filelist[i], a = np.zeros(1), b = np.zeros(1))
continue
landmarks = bbox_lm['b']
for j in range(len(bounding_boxes)):
bbox_coordinates = bounding_boxes[j]
landmark = landmarks[j]
img_face = image.crop((bbox_coordinates[0], bbox_coordinates[1], bbox_coordinates[2], bbox_coordinates[3]))
x = bbox_coordinates[0]
y = bbox_coordinates[1]
for k in range(5):
landmark[k] -= x
landmark[k+5] -= y
img_face = np.array(img_face)
landmark = np.array(landmark)
if len(face_list) != 0:
if img_face.shape[0] == face_list[-1].shape[0]:
img_face = image.crop((bbox_coordinates[0] - 1, bbox_coordinates[1] - 1, bbox_coordinates[2], bbox_coordinates[3]))
img_face = np.array(img_face)
landmark += 1
face_list.append(img_face)
landmarks_new_coordinates.append(landmark)
face_list = np.asarray(face_list)
landmarks_new_coordinates = np.asarray(landmarks_new_coordinates)
np.savez(processed_dataset_path + 'train/Positive/' + train_filelist[i], a = face_list, b = landmarks_new_coordinates)
if i % 100 == 0:
print(i)
except:
print("Error/interrput at validation dataset file " + val_filelist[i])
print(bounding_boxes)
print(landmarks)
print(bounding_boxes.shape)
print(landmarks.shape)
break
| MTCNN/.ipynb_checkpoints/Face_Cropper_TrainValDataset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### String Problems
# ##### https://leetcode.com/explore/challenge/card/30-day-leetcoding-challenge/528/week-1/3283/
# ##### Q.1. Given a non-empty array of integers, every element appears twice except for one. Find that single one.
from typing import List
"""
For O(1) space complexity use math operation or XOR.
a^a = 0
a^0 = a
a^b^c = a^a^b = 0^b = b
"""
class Solution(object):
def singleNumber(self, nums: List[int]) -> int:
"""
:type nums: List[int]
:rtype: int
"""
idx = {}
for i in range(len(nums)):
if nums[i] not in idx:
idx[nums[i]] = 1
else:
idx[nums[i]] += 1
for k in idx.keys():
if idx[k] == 1:
return k
print(Solution().singleNumber([4,1,2,1,2]))
# ##### Q.2. Write an algorithm to determine if a number n is "happy".
#
# ##### A happy number is a number defined by the following process: Starting with any positive integer, replace the number by the sum of the squares of its digits, and repeat the process until the number equals 1 (where it will stay), or it loops endlessly in a cycle which does not include 1. Those numbers for which this process ends in 1 are happy numbers.
#
# ##### Return True if n is a happy number, and False if not.
class Solution(object):
def ifHappy(self, n: int) -> bool:
"""
:type n: int
:rtype: bool
"""
l = 0
while (n != 1):
add = 0
for i in str(n):
add += int(i) ** 2
n = add
l += 1
if l > 100:
return False
return True
print(Solution().ifHappy(19))
# #### Q.3. Given an integer array nums, find the contiguous subarray (containing at least one number) which has the largest sum and return its sum.
class Solution(object):
def maxSubArray(self, nums: List[int]) -> int:
"""
:type nums: List[int]
:rtype int
"""
# Special case is when all values in num are negative.
if max(nums) < 0:
return max(nums)
max_sum = 0; curr = 0
for i in range(len(nums)):
if curr + nums[i] > 0:
curr = curr + nums[i]
else:
curr = 0 # Reset the sum.
if curr > max_sum:
max_sum = curr
return max_sum
print(Solution().maxSubArray([-2,1,-3,4,-1,2,1,-5,4]))
# #### Q.4. Given an array nums, write a function to move all 0's to the end of it while maintaining the relative order of the non-zero elements.
class Solution(object):
def moveZeroes(self, nums: List[int]) -> None:
"""
:type nums: List[int]
:rtype None
Perform inplace ordering.
Method: Apply a form of insert sort that moves each non-negative value
to its right place in the list.
"""
for i in range(len(nums)):
if nums[i] != 0:
j = i
while j > 0 and nums[j - 1] == 0:
nums[j], nums[j-1] = nums[j-1], nums[j]
j -= 1
nums = [0,1,0,3,12]
Solution().moveZeroes(nums)
print(nums)
# #### Q.5. Say you have an array prices for which the ith element is the price of a given stock on day i. Design an algorithm to find the maximum profit. You may complete as many transactions as you like (i.e., buy one and sell one share of the stock multiple times).
# ##### Note: You may not engage in multiple transactions at the same time (i.e., you must sell the stock before you buy again).
class Solution(object):
def maxProfit(self, prices: List[int]) -> int:
"""
:type prices: List[int]
:rtype: int
Maximum Profit is the cumulation of all positive differences.
"""
profit = 0
for i in range(1, len(prices)):
diff = prices[i] - prices[i-1]
if diff > 0:
profit += diff
return profit
print(Solution().maxProfit([7,6,4,3,1]))
# #### Q.6. Given an array of strings, group anagrams together.
class Solution(object):
def groupAnagrams(self, strs: List[str]) -> List[List[str]]:
"""
:type strs: List[str]
:rtype: List[List[str]]
Method: Build a dictionary of words creating a bag of characters representation.
Generate a has for that representation and add words with a similar hash.
"""
words = {}
# Build a dictionary of words.
for word in strs:
boc_vec = [0 for i in range(26)]
for char in word:
boc_vec[ord(char) - 97] += 1
# Check if the representation if present in the dict.
hval = hash(tuple(boc_vec))
if hval not in words:
words[hval] = [word]
else:
words[hval].append(word)
# Once, the dictionary is built, generate list.
fin = []
for key in words.keys():
fin.append(words[key])
return fin
print(Solution().groupAnagrams(["eat", "tea", "tan", "ate", "nat", "bat"]))
# #### Q.7. Given an integer array arr, count how many elements x there are, such that x + 1 is also in arr. If there're duplicates in arr, count them seperately.
class Solution(object):
def countElements(self, arr: List[int]) -> int:
"""
:type arr: List[int]
:rtype: int
Method: Build a dictionary of all numbers in the list and then separately
verify if (n+1) number exists in the dictionary for every n.
"""
nums = {}
for n in arr:
if n not in nums:
nums[n] = 1
cnt = 0
for n in arr:
if n+1 in nums:
cnt += 1
return cnt
print(Solution().countElements([1,3,2,3,5,0]))
if root.left is None and root.right is None:
return 0
def get_longest_path(root):
if root.left is None and root.right is None:
return 0
elif root.left is None:
return 1 + get_longest_path(root.right)
elif root.right is None:
return 1 + get_longest_path(root.left)
else:
return max(1 + get_longest_path(root.left), 1 + get_longest_path(root.right))
return get_longest_path(root.left) + get_longest_path(root.right)
| Python/leetcode/easy/strings-arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 语言翻译
#
# 在此项目中,你将了解神经网络机器翻译这一领域。你将用由英语和法语语句组成的数据集,训练一个序列到序列模型(sequence to sequence model),该模型能够将新的英语句子翻译成法语。
#
# ## 获取数据
#
# 因为将整个英语语言内容翻译成法语需要大量训练时间,所以我们提供了一小部分的英语语料库。
#
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
source_path = 'data/small_vocab_en'
target_path = 'data/small_vocab_fr'
source_text = helper.load_data(source_path)
target_text = helper.load_data(target_path)
# -
# ## 探索数据
#
# 研究 view_sentence_range,查看并熟悉该数据的不同部分。
#
# +
view_sentence_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in source_text.split()})))
sentences = source_text.split('\n')
word_counts = [len(sentence.split()) for sentence in sentences]
print('Number of sentences: {}'.format(len(sentences)))
print('Average number of words in a sentence: {}'.format(np.average(word_counts)))
print()
print('English sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(source_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
print()
print('French sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(target_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
# -
# ## 实现预处理函数
#
# ### 文本到单词 id
#
# 和之前的 RNN 一样,你必须首先将文本转换为数字,这样计算机才能读懂。在函数 `text_to_ids()` 中,你需要将单词中的 `source_text` 和 `target_text` 转为 id。但是,你需要在 `target_text` 中每个句子的末尾,添加 `<EOS>` 单词 id。这样可以帮助神经网络预测句子应该在什么地方结束。
#
#
# 你可以通过以下代码获取 `<EOS> ` 单词ID:
#
# ```python
# target_vocab_to_int['<EOS>']
# ```
#
# 你可以使用 `source_vocab_to_int` 和 `target_vocab_to_int` 获得其他单词 id。
#
# +
def text_to_ids(source_text, target_text, source_vocab_to_int, target_vocab_to_int):
"""
Convert source and target text to proper word ids
:param source_text: String that contains all the source text.
:param target_text: String that contains all the target text.
:param source_vocab_to_int: Dictionary to go from the source words to an id
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: A tuple of lists (source_id_text, target_id_text)
"""
# TODO: Implement Function
source_letter_ids = [[source_vocab_to_int.get(word, source_vocab_to_int['<UNK>']) for word in line.split(' ')] for line in source_text.split('\n')]
target_letter_ids = [[target_vocab_to_int.get(word, target_vocab_to_int['<UNK>']) for word in line.split(' ')] + [target_vocab_to_int['<EOS>']] for line in target_text.split('\n')]
return source_letter_ids, target_letter_ids
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_text_to_ids(text_to_ids)
# -
# ### 预处理所有数据并保存
#
# 运行以下代码单元,预处理所有数据,并保存到文件中。
#
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
helper.preprocess_and_save_data(source_path, target_path, text_to_ids)
# # 检查点
#
# 这是你的第一个检查点。如果你什么时候决定再回到该记事本,或需要重新启动该记事本,可以从这里继续。预处理的数据已保存到磁盘上。
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
import helper
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
# -
# ### 检查 TensorFlow 版本,确认可访问 GPU
#
# 这一检查步骤,可以确保你使用的是正确版本的 TensorFlow,并且能够访问 GPU。
#
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
from distutils.version import LooseVersion
import warnings
import tensorflow as tf
# Check TensorFlow Version
assert LooseVersion(tf.__version__) in [LooseVersion('1.0.0'), LooseVersion('1.0.1')], 'This project requires TensorFlow version 1.0 You are using {}'.format(tf.__version__)
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
# -
# ## 构建神经网络
#
# 你将通过实现以下函数,构建出要构建一个序列到序列模型所需的组件:
#
# - `model_inputs`
# - `process_decoding_input`
# - `encoding_layer`
# - `decoding_layer_train`
# - `decoding_layer_infer`
# - `decoding_layer`
# - `seq2seq_model`
#
# ### 输入
#
# 实现 `model_inputs()` 函数,为神经网络创建 TF 占位符。该函数应该创建以下占位符:
#
# - 名为 “input” 的输入文本占位符,并使用 TF Placeholder 名称参数(等级(Rank)为 2)。
# - 目标占位符(等级为 2)。
# - 学习速率占位符(等级为 0)。
# - 名为 “keep_prob” 的保留率占位符,并使用 TF Placeholder 名称参数(等级为 0)。
#
# 在以下元祖(tuple)中返回占位符:(输入、目标、学习速率、保留率)
#
# +
def model_inputs():
"""
Create TF Placeholders for input, targets, and learning rate.
:return: Tuple (input, targets, learning rate, keep probability)
"""
# TODO: Implement Function
input_data = tf.placeholder(tf.int32, [None, None], name='input')
targets = tf.placeholder(tf.int32, [None, None], name='targets')
lr = tf.placeholder(tf.float32, name='learning_rate')
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
return input_data,targets,lr,keep_prob
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_inputs(model_inputs)
# -
# ### 处理解码输入
#
# 使用 TensorFlow 实现 `process_decoding_input`,以便删掉 `target_data` 中每个批次的最后一个单词 ID,并将 GO ID 放到每个批次的开头。
# +
def process_decoding_input(target_data, target_vocab_to_int, batch_size):
"""
Preprocess target data for dencoding
:param target_data: Target Placehoder
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param batch_size: Batch Size
:return: Preprocessed target data
"""
# TODO: Implement Function
ending = tf.strided_slice(target_data, [0, 0], [batch_size, -1], [1, 1])
dec_input = tf.concat([tf.fill([batch_size, 1], target_vocab_to_int['<GO>']), ending], 1)
return dec_input
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_process_decoding_input(process_decoding_input)
# -
# ### 编码
#
# 实现 `encoding_layer()`,以使用 [`tf.nn.dynamic_rnn()`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn) 创建编码器 RNN 层级。
# +
def encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob):
"""
Create encoding layer
:param rnn_inputs: Inputs for the RNN
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param keep_prob: Dropout keep probability
:return: RNN state
"""
# TODO: Implement Function
enc_cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicLSTMCell(rnn_size)] * num_layers)
enc_cell = tf.contrib.rnn.DropoutWrapper(enc_cell, output_keep_prob=keep_prob)
rnn, enc_state = tf.nn.dynamic_rnn(enc_cell, rnn_inputs, dtype=tf.float32)
return enc_state
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_encoding_layer(encoding_layer)
# -
# ### 解码 - 训练
#
# 使用 [`tf.contrib.seq2seq.simple_decoder_fn_train()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_train) 和 [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder) 创建训练分对数(training logits)。将 `output_fn` 应用到 [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder) 输出上。
# +
def decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope,
output_fn, keep_prob):
"""
Create a decoding layer for training
:param encoder_state: Encoder State
:param dec_cell: Decoder RNN Cell
:param dec_embed_input: Decoder embedded input
:param sequence_length: Sequence Length
:param decoding_scope: TenorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Train Logits
"""
# TODO: Implement Function
dynamic_fn_train = tf.contrib.seq2seq.simple_decoder_fn_train(encoder_state)
train_pred, _, _ = tf.contrib.seq2seq.dynamic_rnn_decoder(cell = dec_cell,
decoder_fn = dynamic_fn_train,
inputs = dec_embed_input,
sequence_length = sequence_length,
scope = decoding_scope)
train_logits = output_fn(train_pred)
droupout = tf.nn.dropout(train_logits, keep_prob)
return droupout;
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_train(decoding_layer_train)
# -
# ### 解码 - 推论
#
# 使用 [`tf.contrib.seq2seq.simple_decoder_fn_inference()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_inference) 和 [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder) 创建推论分对数(inference logits)。
# +
def decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id,
maximum_length, vocab_size, decoding_scope, output_fn, keep_prob):
"""
Create a decoding layer for inference
:param encoder_state: Encoder state
:param dec_cell: Decoder RNN Cell
:param dec_embeddings: Decoder embeddings
:param start_of_sequence_id: GO ID
:param end_of_sequence_id: EOS Id
:param maximum_length: The maximum allowed time steps to decode
:param vocab_size: Size of vocabulary
:param decoding_scope: TensorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Inference Logits
"""
# TODO: Implement Function
dynamic_fn_inference = tf.contrib.seq2seq.simple_decoder_fn_inference(
output_fn, encoder_state, dec_embeddings, start_of_sequence_id, end_of_sequence_id,
maximum_length, vocab_size)
train_logits, _, _ = tf.contrib.seq2seq.dynamic_rnn_decoder(cell = dec_cell,
decoder_fn = dynamic_fn_inference,
scope = decoding_scope)
droupout = tf.nn.dropout(train_logits, keep_prob)
return droupout
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_infer(decoding_layer_infer)
# -
# ### 构建解码层级
#
# 实现 `decoding_layer()` 以创建解码器 RNN 层级。
#
# - 使用 `rnn_size` 和 `num_layers` 创建解码 RNN 单元。
# - 使用 [`lambda`](https://docs.python.org/3/tutorial/controlflow.html#lambda-expressions) 创建输出函数,将输入,也就是分对数转换为类分对数(class logits)。
# - 使用 `decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope, output_fn, keep_prob)` 函数获取训练分对数。
# - 使用 `decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id, maximum_length, vocab_size, decoding_scope, output_fn, keep_prob)` 函数获取推论分对数。
#
# 注意:你将需要使用 [tf.variable_scope](https://www.tensorflow.org/api_docs/python/tf/variable_scope) 在训练和推论分对数间分享变量。
def decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size,
num_layers, target_vocab_to_int, keep_prob):
"""
Create decoding layer
:param dec_embed_input: Decoder embedded input
:param dec_embeddings: Decoder embeddings
:param encoder_state: The encoded state
:param vocab_size: Size of vocabulary
:param sequence_length: Sequence Length
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param keep_prob: Dropout keep probability
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
dec_cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicLSTMCell(rnn_size)] * num_layers)
with tf.variable_scope("decoding") as decoding_scope:
output_fn = lambda x: tf.contrib.layers.fully_connected(x, vocab_size, None, scope=decoding_scope)
train_logits = decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope,
output_fn, keep_prob)
with tf.variable_scope("decoding", reuse=True) as decoding_scope:
infer_logits = decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, target_vocab_to_int['<GO>'], target_vocab_to_int['<EOS>'],
sequence_length, vocab_size, decoding_scope, output_fn, keep_prob)
return train_logits, infer_logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer(decoding_layer)
# ### 构建神经网络
#
# 应用你在上方实现的函数,以:
#
# - 向编码器的输入数据应用嵌入。
# - 使用 `encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob)` 编码输入。
# - 使用 `process_decoding_input(target_data, target_vocab_to_int, batch_size)` 函数处理目标数据。
# - 向解码器的目标数据应用嵌入。
# - 使用 `decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size, num_layers, target_vocab_to_int, keep_prob)` 解码编码的输入数据。
# +
def seq2seq_model(input_data, target_data, keep_prob, batch_size, sequence_length, source_vocab_size, target_vocab_size,
enc_embedding_size, dec_embedding_size, rnn_size, num_layers, target_vocab_to_int):
"""
Build the Sequence-to-Sequence part of the neural network
:param input_data: Input placeholder
:param target_data: Target placeholder
:param keep_prob: Dropout keep probability placeholder
:param batch_size: Batch Size
:param sequence_length: Sequence Length
:param source_vocab_size: Source vocabulary size
:param target_vocab_size: Target vocabulary size
:param enc_embedding_size: Decoder embedding size
:param dec_embedding_size: Encoder embedding size
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
enc_embed_input = tf.contrib.layers.embed_sequence(input_data, source_vocab_size, enc_embedding_size)
encoder_state = encoding_layer(enc_embed_input, rnn_size, num_layers, keep_prob)
target_data = process_decoding_input(target_data, target_vocab_to_int, batch_size)
dec_embeddings = tf.Variable(tf.random_uniform([target_vocab_size, dec_embedding_size]))
target_embed = tf.nn.embedding_lookup(dec_embeddings, target_data)
return decoding_layer(target_embed, dec_embeddings, encoder_state, target_vocab_size, sequence_length, rnn_size, \
num_layers, target_vocab_to_int, keep_prob)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_seq2seq_model(seq2seq_model)
# -
# ## 训练神经网络
#
# ### 超参数
#
# 调试以下参数:
#
# - 将 `epochs` 设为 epoch 次数。
# - 将 `batch_size` 设为批次大小。
# - 将 `rnn_size` 设为 RNN 的大小。
# - 将 `num_layers` 设为层级数量。
# - 将 `encoding_embedding_size` 设为编码器嵌入大小。
# - 将 `decoding_embedding_size` 设为解码器嵌入大小
# - 将 `learning_rate` 设为训练速率。
# - 将 `keep_probability` 设为丢弃保留率(Dropout keep probability)。
# Number of Epochs
epochs = 4
# Batch Size
batch_size = 256
# RNN Size
rnn_size = 512
# Number of Layers
num_layers = 3
# Embedding Size
encoding_embedding_size = 256
decoding_embedding_size = 256
# Learning Rate
learning_rate = 0.001
# Dropout Keep Probability
keep_probability = 0.5
# ### 构建图表
#
# 使用你实现的神经网络构建图表。
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_path = 'checkpoints/dev'
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
max_source_sentence_length = max([len(sentence) for sentence in source_int_text])
train_graph = tf.Graph()
with train_graph.as_default():
input_data, targets, lr, keep_prob = model_inputs()
sequence_length = tf.placeholder_with_default(max_source_sentence_length, None, name='sequence_length')
input_shape = tf.shape(input_data)
train_logits, inference_logits = seq2seq_model(
tf.reverse(input_data, [-1]), targets, keep_prob, batch_size, sequence_length, len(source_vocab_to_int), len(target_vocab_to_int),
encoding_embedding_size, decoding_embedding_size, rnn_size, num_layers, target_vocab_to_int)
tf.identity(inference_logits, 'logits')
with tf.name_scope("optimization"):
# Loss function
cost = tf.contrib.seq2seq.sequence_loss(
train_logits,
targets,
tf.ones([input_shape[0], sequence_length]))
# Optimizer
optimizer = tf.train.AdamOptimizer(lr)
# Gradient Clipping
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None]
train_op = optimizer.apply_gradients(capped_gradients)
# -
# ### 训练
#
# 利用预处理的数据训练神经网络。如果很难获得低损失值,请访问我们的论坛,看看其他人是否遇到了相同的问题。
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import time
def get_accuracy(target, logits):
"""
Calculate accuracy
"""
max_seq = max(target.shape[1], logits.shape[1])
if max_seq - target.shape[1]:
target = np.pad(
target,
[(0,0),(0,max_seq - target.shape[1])],
'constant')
if max_seq - logits.shape[1]:
logits = np.pad(
logits,
[(0,0),(0,max_seq - logits.shape[1]), (0,0)],
'constant')
return np.mean(np.equal(target, np.argmax(logits, 2)))
train_source = source_int_text[batch_size:]
train_target = target_int_text[batch_size:]
valid_source = helper.pad_sentence_batch(source_int_text[:batch_size])
valid_target = helper.pad_sentence_batch(target_int_text[:batch_size])
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
for epoch_i in range(epochs):
for batch_i, (source_batch, target_batch) in enumerate(
helper.batch_data(train_source, train_target, batch_size)):
start_time = time.time()
_, loss = sess.run(
[train_op, cost],
{input_data: source_batch,
targets: target_batch,
lr: learning_rate,
sequence_length: target_batch.shape[1],
keep_prob: keep_probability})
batch_train_logits = sess.run(
inference_logits,
{input_data: source_batch, keep_prob: 1.0})
batch_valid_logits = sess.run(
inference_logits,
{input_data: valid_source, keep_prob: 1.0})
train_acc = get_accuracy(target_batch, batch_train_logits)
valid_acc = get_accuracy(np.array(valid_target), batch_valid_logits)
end_time = time.time()
print('Epoch {:>3} Batch {:>4}/{} - Train Accuracy: {:>6.3f}, Validation Accuracy: {:>6.3f}, Loss: {:>6.3f}'
.format(epoch_i, batch_i, len(source_int_text) // batch_size, train_acc, valid_acc, loss))
# Save Model
saver = tf.train.Saver()
saver.save(sess, save_path)
print('Model Trained and Saved')
# -
# ### 保存参数
#
# 保存 `batch_size` 和 `save_path` 参数以进行推论(for inference)。
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Save parameters for checkpoint
helper.save_params(save_path)
# # 检查点
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import tensorflow as tf
import numpy as np
import helper
import problem_unittests as tests
_, (source_vocab_to_int, target_vocab_to_int), (source_int_to_vocab, target_int_to_vocab) = helper.load_preprocess()
load_path = helper.load_params()
# -
# ## 句子到序列
#
# 要向模型提供要翻译的句子,你首先需要预处理该句子。实现函数 `sentence_to_seq()` 以预处理新的句子。
#
# - 将句子转换为小写形式
# - 使用 `vocab_to_int` 将单词转换为 id
# - 如果单词不在词汇表中,将其转换为`<UNK>` 单词 id
# +
def sentence_to_seq(sentence, vocab_to_int):
"""
Convert a sentence to a sequence of ids
:param sentence: String
:param vocab_to_int: Dictionary to go from the words to an id
:return: List of word ids
"""
# TODO: Implement Function
sentence = sentence.lower()
words = sentence.split(' ')
ids = []
for w in words:
if(w in vocab_to_int):
ids.append(vocab_to_int[w])
else:
ids.append(vocab_to_int['<UNK>'])
return ids
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_sentence_to_seq(sentence_to_seq)
# -
# ## 翻译
#
# 将 `translate_sentence` 从英语翻译成法语。
# +
translate_sentence = 'he saw a old yellow truck .'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
translate_sentence = sentence_to_seq(translate_sentence, source_vocab_to_int)
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load saved model
loader = tf.train.import_meta_graph(load_path + '.meta')
loader.restore(sess, load_path)
input_data = loaded_graph.get_tensor_by_name('input:0')
logits = loaded_graph.get_tensor_by_name('logits:0')
keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
translate_logits = sess.run(logits, {input_data: [translate_sentence], keep_prob: 1.0})[0]
print('Input')
print(' Word Ids: {}'.format([i for i in translate_sentence]))
print(' English Words: {}'.format([source_int_to_vocab[i] for i in translate_sentence]))
print('\nPrediction')
print(' Word Ids: {}'.format([i for i in np.argmax(translate_logits, 1)]))
print(' French Words: {}'.format([target_int_to_vocab[i] for i in np.argmax(translate_logits, 1)]))
# -
# ## 不完美的翻译
#
# 你可能注意到了,某些句子的翻译质量比其他的要好。因为你使用的数据集只有 227 个英语单词,但实际生活中有数千个单词,只有使用这些单词的句子结果才会比较理想。对于此项目,不需要达到完美的翻译。但是,如果你想创建更好的翻译模型,则需要更好的数据。
#
# 你可以使用 [WMT10 French-English corpus](http://www.statmt.org/wmt10/training-giga-fren.tar) 语料库训练模型。该数据集拥有更多的词汇,讨论的话题也更丰富。但是,训练时间要好多天的时间,所以确保你有 GPU 并且对于我们提供的数据集,你的神经网络性能很棒。提交此项目后,别忘了研究下 WMT10 语料库。
#
#
# ## 提交项目
#
# 提交项目时,确保先运行所有单元,然后再保存记事本。保存记事本文件为 “dlnd_language_translation.ipynb”,再通过菜单中的“文件” ->“下载为”将其另存为 HTML 格式。提交的项目文档中需包含“helper.py”和“problem_unittests.py”文件。
#
| language-translation/.ipynb_checkpoints/dlnd_language_translation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Huffman coding
# ## Zadanie polega na implementacji dwóch algorytmów kompresji:
# __- statycznego algorytmu Huffmana (1 punkty),__
#
# __- dynamicznego algorytmu Huffmana (2 punkty).__
# ## Dla każdego z algorytmów należy wykonać następujące zadania:
# __1) Opracować format pliku przechowującego dane,__
#
# __2) Zaimplementować algorytm kompresji i dekompresji danych dla tego formatu pliku,__
#
# __3) Zmierzyć współczynnik kompresji (wyrażone w procentach: 1 - plik_skompresowany / plik_nieskompresowany) dla plików tekstowych o rozmiarach: 1kB, 10kB, 100kB, 1MB,__
#
# __4) Zmierzyć czas kompresji i dekompresji dla plików z punktu 3.__
# ### Imports
from heapq import heappop, heappush, heapify
from bitarray import bitarray
from bitarray.util import ba2int
# ### Statyczny algorytm Huffmana
# +
class Node:
def __init__(self, value, char=None, left=None, right=None):
self.value = value
self.char = char
self.left = left
self.right = right
def __gt__(self, other):
return self.value > other.value
class StaticHuffmanTree:
def __init__(self, text):
self.root = self.build_static_huffman_tree(text)
self.codes = dict()
self.create_huffman_codes(self.root, self.codes, bitarray())
def build_static_huffman_tree(self, text):
letters = dict()
for char in text:
letters[char] = letters.get(char, 0) + 1
leafs = [Node(weight, char) for char, weight in letters.items()]
while len(leafs) > 1:
first, second = heappop(leafs), heappop(leafs)
heappush(leafs, Node(first.value + second.value, left=first, right=second))
return leafs[0]
def create_huffman_codes(self, node, codes, code):
if node.char is not None:
codes[node.char] = code
code_cpy = code.copy()
if node.left is not None:
code.append(0)
self.create_huffman_codes(node.left, codes, code)
if node.right is not None:
code = code_cpy
code.append(1)
self.create_huffman_codes(node.right, codes, code)
def encode_static_huffman_tree(self, text):
result = bitarray()
for char in text: # TODO check append below
result.extend(self.codes[char])
return result
def decode_static_huffman_tree(self, encoded_text):
node = self.root
decoded_text = ""
for bit in encoded_text:
if not bit:
node = node.left
else:
node = node.right
if not node.left and not node.right:
decoded_text += node.char
node = self.root
return decoded_text
# -
# __Example test__
text = "Hello world. I'm Huffman encoder and decoder!"
tree = StaticHuffmanTree(text)
result = tree.encode_static_huffman_tree(text)
tree.decode_static_huffman_tree(result)
# ### Dynamiczny algorytm Huffmana
# +
class Node2:
def __init__(self, weight=0, index=0, char=None, left=None, right=None, parent=None):
self.weight = weight
self.index = index
self.char = char
self.left = left
self.right = right
self.parent = parent
class AdaptiveHuffmanTree:
def __init__(self):
self.index = 520
NYT = Node2(weight=0, index=self.index + 1, char='NYT')
self.NYT = NYT
self.root = NYT
self.leaves = {"NYT": self.root}
self.weights = {0: {self.root}, 1: set()}
def add_new_node(self, char):
node = self.NYT
left_node = Node2(weight=0, index=self.index - 1, parent=node, char="NYT")
node.left = left_node
right_node = Node2(weight=1, index=self.index, parent=node, char=char)
node.right = right_node
node.char = None
self.index -= 2
self.NYT = left_node
self.weights[0].add(left_node)
self.weights[1].add(right_node)
self.leaves[char] = right_node
self.leaves["NYT"] = left_node
self.increment_and_swap(node)
def increment_and_swap(self, node):
while node != self.root:
node = node.parent
max_index_node = max(self.weights[node.weight], key=lambda nd: nd.index)
if node != max_index_node:
node.index, max_index_node.index = max_index_node.index, node.index
if node.parent == max_index_node.parent:
if node == node.parent.left:
node.parent.right = node
node.parent.left = max_index_node
else:
node.parent.right = max_index_node
node.parent.left = node
else:
if node == node.parent.left:
node.parent.left = max_index_node
else:
node.parent.right = max_index_node
if max_index_node.parent.left == max_index_node:
max_index_node.parent.left = node
else:
max_index_node.parent.right = node
if node.parent != max_index_node.parent:
max_index_node.parent, node.parent = node.parent, max_index_node.parent
self.weights[node.weight].remove(node)
node.weight += 1
if node.weight not in self.weights:
self.weights[node.weight] = set()
self.weights[node.weight].add(node)
def get_code(self, char):
node = self.leaves[char]
code = bitarray()
while node != self.root:
if node == node.parent.left:
code.append(0)
else:
code.append(1)
node = node.parent
code.reverse()
return code
def encode_adaptive_huffman_tree(self, text):
coded_text = bitarray()
for char in text:
if char in self.leaves:
coded_text += self.get_code(char)
self.increment_and_swap(self.leaves[char])
else:
coded_char = self.get_code('NYT')
coded_char.frombytes(char.encode("utf-8"))
coded_text += coded_char
self.add_new_node(char)
end_bits = 8 - len(coded_text) % 8
coded_text = bitarray(f"{end_bits:08b}") + coded_text + bitarray(end_bits)
return coded_text
def decode_adaptive_huffman_tree(encoded_text):
tree = AdaptiveHuffmanTree()
node = tree.root
idx = 0
encoded_text = encoded_text[8:-ba2int(encoded_text[:8])]
decoded_text = ""
while idx < len(encoded_text):
while not (node.left is None and node.right is None):
if not encoded_text[idx]:
node = node.left
else:
node = node.right
idx += 1
if node.char == "NYT":
char_coded = encoded_text[idx:idx + 8]
char_decoded = char_coded.tobytes().decode("utf-8")
tree.add_new_node(char_decoded)
idx += 8
else:
char_decoded = node.char
tree.increment_and_swap(tree.leaves[char_decoded])
node = tree.root
decoded_text += char_decoded
return decoded_text
# -
# __Example test__
text = "Hello world. I'm Huffman encoder and decoder!"
tree = AdaptiveHuffmanTree()
result = tree.encode_adaptive_huffman_tree(text)
print(decode_adaptive_huffman_tree(result))
# ### Tests
from timeit import default_timer as timer
import os
# __Compression test__
def compression_ratio(read_file, write_file):
original_size = os.path.getsize(read_file)
coded_size = os.path.getsize(write_file)
return 1 - coded_size / original_size
def compression_test(read_file, size):
save_file = f"output_files/compression_static_{size}.txt"
static_tree = StaticHuffmanTree(text)
encoded = static_tree.encode_static_huffman_tree(text)
with open(save_file, "wb+") as f:
encoded.tofile(f)
static_compression = compression_ratio(read_file, save_file)
print(f"Compression ratio for {read_file} for StaticHuffmanTree is {static_compression * 100}%.")
save_file = f"output_files/compression_adaptive_{size}.txt"
adaptive_tree = AdaptiveHuffmanTree()
encoded = adaptive_tree.encode_adaptive_huffman_tree(text)
with open(save_file, "wb+") as f:
encoded.tofile(f)
adaptive_compression = compression_ratio(read_file, save_file)
print(f"Compression ratio for {read_file} for AdaptiveHuffmanTree is {adaptive_compression * 100}%.")
# 1kb file
compression_test("test_files/1kB.txt", "1kb")
# 10kb file
compression_test("test_files/10kB.txt", "10kb")
# 100kb file
compression_test("test_files/100kB.txt", "100kb")
# 1MB file
compression_test("test_files/1MB.txt", "1Mb")
# __Time test__
def time_test(filename, n):
with open(filename, "r") as f:
text = f.read()
static_tree = StaticHuffmanTree(text)
average_time = 0
for _ in range(n):
start = timer()
result = static_tree.encode_static_huffman_tree(text)
static_tree.decode_static_huffman_tree(result)
end = timer()
average_time += (end - start)
average_time /= n
print(f"Average time execution for {filename} for StaticHuffmanTree is {average_time} s.")
average_time = 0
for _ in range(n):
adaptive_tree = AdaptiveHuffmanTree()
start = timer()
result = adaptive_tree.encode_adaptive_huffman_tree(text)
decode_adaptive_huffman_tree(result)
end = timer()
average_time += (end - start)
average_time /= n
print(f"Average time execution for {filename} AdaptiveHuffmanTree is {average_time} s.")
# 1kb file
time_test("test_files/1kB.txt", 100)
# 10kb file
time_test("test_files/10kB.txt", 100)
# 100kb file
time_test("test_files/100kB.txt", 50)
# 1MB file
time_test("test_files/1MB.txt", 10)
| Lab_3/Lab_3_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from scipy.spatial.distance import pdist
from scipy.cluster.hierarchy import *
from matplotlib import pyplot as plt
from matplotlib import rc
import numpy as np
from sklearn.cluster import KMeans
from sklearn import preprocessing
from scipy.cluster.hierarchy import fcluster
data = pd.read_excel('Задача.xlsx')
# -
data.head()
data.describe()
from pandas.plotting import scatter_matrix
scatter_matrix(data, alpha=0.05, figsize=(10, 10));
data.corr()
data_for_clust=data.drop(data.columns[0], axis=1).values
dataNorm = preprocessing.scale(data_for_clust)
data_dist = pdist(dataNorm, 'euclidean')
data_linkage = linkage(data_dist, method='average')
# +
last = data_linkage[-10:, 2]
last_rev = last[::-1]
idxs = np.arange(1, len(last) + 1)
plt.plot(idxs, last_rev)
acceleration = np.diff(last, 2)
acceleration_rev = acceleration[::-1]
plt.plot(idxs[:-2] + 1, acceleration_rev)
plt.show()
k = acceleration_rev.argmax() + 2
print("clusters:", k)
# -
km = KMeans(k).fit(dataNorm)
plt.figure(figsize=(10, 8))
plt.scatter(data_for_clust[:,0], data_for_clust[:,2], c=km.labels_, cmap='flag')
plt.show()
k=4
clusters=fcluster(data_linkage, k, criterion='maxclust')
dataK=data
dataK['group_no']=clusters
writer = pd.ExcelWriter('result.xlsx')
dataK.to_excel(writer,'KMeans')
writer.save()
data_clust4 = pd.read_excel('cluster4.xlsx')
data_clust4.drop(data_clust4.columns[[0,1,6]],axis=1,inplace=True)
data_clust4.head()
data_clust4.describe()
data_clust1 = pd.read_excel('cluster1.xlsx')
data_clust1.drop(data_clust1.columns[[0,1,6]],axis=1,inplace=True)
data_clust1.describe()
data_clust2 = pd.read_excel('cluster2.xlsx')
data_clust2.drop(data_clust2.columns[[0,1,6]],axis=1,inplace=True)
data_clust2.describe()
data_clust3 = pd.read_excel('cluster3.xlsx')
data_clust3.drop(data_clust3.columns[[0,1,6]],axis=1,inplace=True)
data_clust3.describe()
| data analysis JN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tensorflow)
# language: python
# name: tensorflow
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import os
from scipy.optimize import curve_fit
def sigmoid(x, L ,x0, k, b):
y = L / (1 + np.exp(-k*(x-x0)))+b
return (y)
def sigmoidfit(xdata,ydata):
p0 = [max(ydata), np.median(xdata),1,min(ydata)] # this is an mandatory initial guess
popt, pcov = curve_fit(sigmoid, xdata, ydata, p0,maxfev = 3000)
return popt
plt.figure(figsize=(5,10))
for n,i in enumerate(filter(lambda v: ".npy" in v,os.listdir())):
plt.subplot(5,1,n+1)
a = np.load(f"{i}",allow_pickle=True)
a = a.item()
values = np.array([temp[0] for temp in a['cv_ll_scores']])
plt.plot(a['x_dims'],values,'+')
opts = sigmoidfit(a['x_dims'][~np.isnan(values)],values[~np.isnan(values)])
plt.plot(a['x_dims'],sigmoid(a['x_dims'],*opts),'k--')
name = i.split('.')[0]
plt.title(f"Session {name.split('_')[1]} ({name.split('_')[0]} Region)")
plt.box(on=None)
plt.ylabel("Max Log-Likelihood")
plt.xlabel("Number of Latent Dimensions")
plt.tight_layout()
# -
plt.figure(figsize=(7,5))
xys = []
for n,i in enumerate(filter(lambda v: ".npy" in v,os.listdir())):
#plt.subplot(5,1,n+1)
a = np.load(f"{i}",allow_pickle=True)
a = a.item()
values = np.array([temp[0] for temp in a['cv_ll_scores']])
values = (values-values[0])/(values[~np.isnan(values)][-1]-values[0])
plt.plot(a['x_dims'],values,'o', mfc='none',ms=5)
xys.append([a['x_dims'],values])
xys = np.concatenate(xys,axis=1).T
order = np.argsort(xys[:,0])
plt.plot(xys[order,0],sigmoid(xys[order,0],*sigmoidfit(xys[:,0][~np.isnan(xys[:,1])],xys[:,1][~np.isnan(xys[:,1])])),'k--',linewidth=3)
plt.xlabel("Number of Latent Dimensions")
plt.ylabel("End-Normalised Maximum Log-Likelihood")
plt.box(on=None)
plt.tight_layout()
| Latent Information Approach/results/GFPA-maxLL-data/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/priyanshgupta1998/WebScraping/blob/master/Scrape_DOB_Name_URL.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PnWoVQLDrXTL" colab_type="code" colab={}
from urllib.request import urlopen as ureq
from bs4 import BeautifulSoup
import requests
import pandas as pd
# + id="k9XtaiIf-1qg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 638} outputId="221471fb-cd43-4a61-bbfb-686002472a92"
links = ['https://nettv4u.com/celebrity/english/producer/deborah-snyder' ,
'https://nettv4u.com/celebrity/english/producer/benjamin-melniker' ,
'https://nettv4u.com/celebrity/english/producer/wesley-coller',
'https://nettv4u.com/celebrity/english/movie-actress/demi-kazanis' ,
'https://nettv4u.com/celebrity/english/movie-actor/david-midura',
'https://nettv4u.com/celebrity/english/movie-actor/dante-briggins',
'https://nettv4u.com/celebrity/english/stunt-director/ele-bardha',
'https://nettv4u.com/celebrity/english/movie-actor/gordon-michaels',
'https://nettv4u.com/celebrity/english/movie-actress/christina-wren',
'https://nettv4u.com/celebrity/english/movie-actor/brian-boland',
'https://nettv4u.com/celebrity/english/movie-actress/heather-fairbanks',
'https://nettv4u.com/celebrity/english/movie-actress/jade-chynoweth',
'https://nettv4u.com/celebrity/english/movie-actress/nicole-santini',
'https://nettv4u.com/celebrity/english/movie-actor/roger-wiggins',
'https://nettv4u.com/celebrity/english/movie-actress/sandhya-chandel',
'https://nettv4u.com/celebrity/english/movie-actress/tao-okamoto',
'https://nettv4u.com/celebrity/english/movie-actress/terri-partyka',
'https://nettv4u.com/celebrity/english/producer/denise-ream',
'https://nettv4u.com/celebrity/english/executive-producer/scott-niemeyer']
name = []
post = []
img = []
for i in range(len(links)):
name.append(links[i].split('/')[-1])
page = requests.get(links[i])
html = BeautifulSoup(page.content,'html.parser')
div = html.findAll("div",{"class":"actor-photo"})
post.append(div[0].img.get('alt'))
img.append(div[0].img.get('src'))
data = pd.DataFrame()
data['Name'] = name
data['KnownAs'] = post
data['Links'] = links
data['Image_Link'] = img
data
# + id="ysHw1-9VZ1A4" colab_type="code" colab={}
data.to_csv('data.csv',index=False)
# + id="8P5WYhdR-1eQ" colab_type="code" colab={}
# + [markdown] id="l6FdnWzf2rhe" colab_type="text"
# #Full Details ---> Producer/Actor/Director....'s Name with their DOB and Link
# + id="zuJNkUKxrXW1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="55091367-8828-49a3-9c27-539979ee9405"
#For page number 0
url = 'https://nettv4u.com/celebrity/english/producer/'
page = requests.get(url)
html = BeautifulSoup(page.content,'html.parser')
div= html.findAll("div",{"id":"celeb-item-all"})
liss = div[0].findAll('a')
producer = []
producer = producer + div[0].text.strip().split('\n\n\n\n\n\n\n\n')
links =[]
for i in range(0 , len(liss) , 2):
links.append('https://nettv4u.com' + liss[i].get('href'))
images = []
for i in range(len(liss)):
if(liss[i].img!=None):
images += [str(liss[i].img.get('data-src'))]
#lin = list(set(lin))
#Now for the rest of the pages
num_of_pages = int(input("Enter the number of pages you want to scrape ...less than or equal to 53 : ")) + 1
for i in range(1,num_of_pages):
url = 'https://nettv4u.com/celebrity/english/producer/?page={}'.format(i)
page = requests.get(url)
html = BeautifulSoup(page.content,'html.parser')
# html.findAll("a",{"class":"LinkNormal"})
div2 = html.findAll("div",{"id":"celeb-item-all"})
#For post , Name and DOB
producer += div2[0].text.strip().split('\n\n\n\n\n\n\n\n')
#For links
liss = div2[0].findAll('a')
for i in range(0 ,len(liss),2):
links += ['https://nettv4u.com' + liss[i].get('href')]
#urls = list(set(urls))
#links += urls
for i in range(len(liss)):
if(liss[i].img!=None):
images += [str(liss[i].img.get('data-src'))]
# + id="VZuBRd9XGm1V" colab_type="code" colab={}
names = []
dob =[]
work = []
for i in range(len(producer)):
k = producer[i].strip().split('\n\n')
if(len(k)<2):
k +=['DOB : ?']
names.append(k[0])
dob.append(k[1].split(':')[1])
work.append(links[i].split('https://nettv4u.com/celebrity/english/')[1].split('/')[0])
# + id="bj4gRZAYrXYY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="6709bc8c-b9b9-4430-e369-f5303181dcc4"
df = pd.DataFrame()
df['Name'] = names
df['DOB : DD-MM-YYYY'] = dob
df['Task & Job'] = work
df['Url_Web_page_Link'] = links
df['Image_Link'] = images
print(df.shape)
df.head()
# + id="LGgmN9580riY" colab_type="code" colab={}
df.to_csv('Intership_ssignment.csv' , index=False)
# + id="Cw0ypqeD0rry" colab_type="code" colab={}
# + id="l6vDk4ya0rp2" colab_type="code" colab={}
# + id="a-0UYztF0rnB" colab_type="code" colab={}
# + id="fYuBR_JE0rgf" colab_type="code" colab={}
# + id="EeMYJH3C0ret" colab_type="code" colab={}
# + id="qEITQ5l80rcd" colab_type="code" colab={}
# + id="38ZK-5GL0rYv" colab_type="code" colab={}
# + id="W_L0FYX10rVh" colab_type="code" colab={}
# + id="tdmCTOEVrXaO" colab_type="code" colab={}
# + [markdown] id="xB0726sO63Fz" colab_type="text"
# #Practice Work .........
# + id="bCG-Aw21IxJV" colab_type="code" colab={}
# + id="vTTLVsPWIxBJ" colab_type="code" colab={}
print(len(producer) , len(links) , len(images))
i= 35
print(links[i])
print(producer[i])
images[i]
# + id="A54T4FavrXNn" colab_type="code" colab={}
from urllib.request import urlopen as ureq
from bs4 import BeautifulSoup
import requests
# + id="7SczgdMyrXP5" colab_type="code" colab={}
url = 'https://nettv4u.com/celebrity/english/'
page = requests.get(url)
html = BeautifulSoup(page.content,'html.parser')
# html.findAll("a",{"class":"LinkNormal"})
div1 = html.findAll("a",{"class":"dropdown-item"})
# + id="LN1nnRmOrXRo" colab_type="code" outputId="5bcc7c69-42a3-4996-9ffd-e2a63c988bf7" colab={"base_uri": "https://localhost:8080/", "height": 153}
globe = []
for i in range(len(div1)):
if "https://nettv4u.com/celebrity/" in div1[i].get('href'):
globe.append(div1[i].get('href'))
globe
# + id="4oIwXkOyrXdC" colab_type="code" colab={}
url = 'https://nettv4u.com/celebrity/english/producer/page=1'
page = requests.get(url)
html = BeautifulSoup(page.content,'html.parser')
# html.findAll("a",{"class":"LinkNormal"})
div = html.findAll("a")
producer = []
for i in range(len(div)):
if(len(str(div[i].get('href')))>0):
if "/celebrity/english/producer/" in str(div[i].get('href')):
producer.append('https://nettv4u.com' + div[i].get('href'))
producer
| Scrape_DOB_Name_URL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 如何使用和开发微信聊天机器人的系列教程
# # A workshop to develop & use an intelligent and interactive chat-bot in WeChat
# ### WeChat is a popular social media app, which has more than 800 million monthly active users.
#
# <img src='http://www.kudosdata.com/wp-content/uploads/2016/11/cropped-KudosLogo1.png' width=30% style="float: right;">
# <img src='reference/WeChat_SamGu_QR.png' width=10% style="float: right;">
#
# ### http://www.KudosData.com
#
# by: <EMAIL>
#
#
# May 2017 ========== Scan the QR code to become trainer's friend in WeChat ========>>
# ### 第四课:自然语言处理:语义和情感分析
# ### Lesson 4: Natural Language Processing 2
# * 消息文字中名称实体的识别 (Name-Entity detection)
# * 消息文字中语句的情感分析 (Sentiment analysis, Sentence level)
# * 整篇消息文字的情感分析 (Sentiment analysis, Document level)
# * 语句的语法分析 (Syntax / Grammer analysis)
# ### Flag to indicate the environment to run this program:
# parm_runtime_env_GCP = True
parm_runtime_env_GCP = False
# ### Using Google Cloud Platform's Machine Learning APIs
# From the same API console, choose "Dashboard" on the left-hand menu and "Enable API".
#
# Enable the following APIs for your project (search for them) if they are not already enabled:
# <ol>
# <li> Google Translate API </li>
# <li> Google Cloud Vision API </li>
# <li> Google Natural Language API </li>
# <li> Google Cloud Speech API </li>
# </ol>
#
# Finally, because we are calling the APIs from Python (clients in many other languages are available), let's install the Python package (it's not installed by default on Datalab)
# +
# Copyright 2016 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License");
# import subprocess
# retcode = subprocess.call(['pip', 'install', '-U', 'google-api-python-client'])
# retcode = subprocess.call(['pip', 'install', '-U', 'gTTS'])
# Below is for GCP only: install audio conversion tool
# retcode = subprocess.call(['apt-get', 'update', '-y'])
# retcode = subprocess.call(['apt-get', 'install', 'libav-tools', '-y'])
# -
# ### 导入需要用到的一些功能程序库:
import io, os, subprocess, sys, re, codecs, time, datetime, requests, itchat
from itchat.content import *
from googleapiclient.discovery import build
# ### GCP Machine Learning API Key
#
# First, visit <a href="http://console.cloud.google.com/apis">API console</a>, choose "Credentials" on the left-hand menu. Choose "Create Credentials" and generate an API key for your application. You should probably restrict it by IP address to prevent abuse, but for now, just leave that field blank and delete the API key after trying out this demo.
#
# Copy-paste your API Key here:
# +
# Here I read in my own API_KEY from a file, which is not shared in Github repository:
with io.open('../../../API_KEY.txt') as fp:
for line in fp: APIKEY = line
# You need to un-comment below line and replace 'APIKEY' variable with your own GCP API key:
# APIKEY='<KEY>'
# +
# Below is for Google Speech synthesis: text to voice API
# from gtts import gTTS
# Below is for Google Speech recognition: voice to text API
# speech_service = build('speech', 'v1', developerKey=APIKEY)
# Below is for Google Language Tranlation API
# service = build('translate', 'v2', developerKey=APIKEY)
# Below is for Google Natual Language Processing API
# nlp_service = build('language', 'v1', developerKey=APIKEY)
nlp_service = build('language', 'v1beta2', developerKey=APIKEY)
# -
# ### 多媒体二进制base64码转换 (Define media pre-processing functions)
# +
# Import the base64 encoding library.
import base64
# Pass the image data to an encoding function.
def encode_image(image_file):
with io.open(image_file, "rb") as image_file:
image_content = image_file.read()
# Python 2
if sys.version_info[0] < 3:
return base64.b64encode(image_content)
# Python 3
else:
return base64.b64encode(image_content).decode('utf-8')
# Pass the audio data to an encoding function.
def encode_audio(audio_file):
with io.open(audio_file, 'rb') as audio_file:
audio_content = audio_file.read()
# Python 2
if sys.version_info[0] < 3:
return base64.b64encode(audio_content)
# Python 3
else:
return base64.b64encode(audio_content).decode('utf-8')
# -
# ### 机器智能API接口控制参数 (Define control parameters for API)
# +
# API control parameter for Image API:
parm_image_maxResults = 10 # max objects or faces to be extracted from image analysis
# API control parameter for Language Translation API:
parm_translation_origin_language = 'zh' # original language in text: to be overwriten by TEXT_DETECTION
parm_translation_target_language = 'zh' # target language for translation: Chinese
# API control parameter for 消息文字转成语音 (Speech synthesis: text to voice)
parm_speech_synthesis_language = 'zh' # speech synthesis API 'text to voice' language
# parm_speech_synthesis_language = 'zh-tw' # speech synthesis API 'text to voice' language
# parm_speech_synthesis_language = 'zh-yue' # speech synthesis API 'text to voice' language
# API control parameter for 语音转换成消息文字 (Speech recognition: voice to text)
# parm_speech_recognition_language = 'en' # speech API 'voice to text' language
parm_speech_recognition_language = 'cmn-Hans-CN' # speech API 'voice to text' language
# API control parameter for 自然语言处理:语义和情感分析
parm_nlp_extractDocumentSentiment = True # 情感分析 (Sentiment analysis)
parm_nlp_extractEntities = True # 消息文字中名称实体的识别 (Name-Entity detection)
parm_nlp_extractEntitySentiment = False # Only available in v1beta2. But Chinese language zh is not supported yet.
parm_nlp_extractSyntax = True # 语句的语法分析 (Syntax / Grammer analysis)
# -
# ### 定义一个调用自然语言处理接口的小功能
# Running Speech API
def KudosData_nlp(text, extractDocumentSentiment, extractEntities, extractEntitySentiment, extractSyntax):
# Python 2
# if sys.version_info[0] < 3:
# tts = gTTS(text=text2voice.encode('utf-8'), lang=parm_speech_synthesis_language, slow=False)
# Python 3
# else:
# tts = gTTS(text=text2voice, lang=parm_speech_synthesis_language, slow=False)
request = nlp_service.documents().annotateText(body={
"document":{
"type": "PLAIN_TEXT",
"content": text
},
"features": {
"extractDocumentSentiment": extractDocumentSentiment,
"extractEntities": extractEntities,
"extractEntitySentiment": extractEntitySentiment, # only available in v1beta2
"extractSyntax": extractSyntax,
},
"encodingType":"UTF8"
})
responses = request.execute(num_retries=3)
print('\nCompeleted: NLP analysis API')
return responses
# < Start of interactive demo >
text4nlp = 'As a data science consultant and trainer with Kudos Data, <NAME> (Sam) engages communities and schools ' \
'to help organizations making sense of their data using advanced data science , machine learning and ' \
'cloud computing technologies. Inspire next generation of artificial intelligence lovers and leaders.'
text4nlp = '作为酷豆数据科学的顾问和培训师,<NAME> (白黑) 善长联络社群和教育资源。' \
'促进各大公司组织使用先进的数据科学、机器学习和云计算技术来获取数据洞见。激励下一代人工智能爱好者和领导者。'
responses = KudosData_nlp(text4nlp
, parm_nlp_extractDocumentSentiment
, parm_nlp_extractEntities
, parm_nlp_extractEntitySentiment
, parm_nlp_extractSyntax)
# +
# print(responses)
# -
# ### * 消息文字中名称实体的识别 (Name-Entity detection)
# +
# print(responses['entities'])
# -
for i in range(len(responses['entities'])):
# print(responses['entities'][i])
print('')
print(u'[ 实体 {} : {} ]\n 实体类别 : {}\n 重要程度 : {}'.format(
i+1
, responses['entities'][i]['name']
, responses['entities'][i]['type']
, responses['entities'][i]['salience']
))
# print(responses['entities'][i]['name'])
# print(responses['entities'][i]['type'])
# print(responses['entities'][i]['salience'])
if 'sentiment' in responses['entities'][i]:
print(u' 褒贬程度 : {}\n 语彩累积 : {}'.format(
responses['entities'][i]['sentiment']['score']
, responses['entities'][i]['sentiment']['magnitude']
))
# print(responses['entities'][i]['sentiment'])
if responses['entities'][i]['metadata'] != {}:
if 'wikipedia_url' in responses['entities'][i]['metadata']:
print(' ' + responses['entities'][i]['metadata']['wikipedia_url'])
# ### * 消息文字中语句的情感分析 (Sentiment analysis, Sentence level)
# +
# print(responses['sentences'])
# -
for i in range(len(responses['sentences'])):
print('')
print(u'[ 语句 {} : {} ]\n( 褒贬程度 : {} | 语彩累积 : {} )'.format(
i+1
, responses['sentences'][i]['text']['content']
, responses['sentences'][i]['sentiment']['score']
, responses['sentences'][i]['sentiment']['magnitude']
))
# https://cloud.google.com/natural-language/docs/basics
#
# * **score 褒贬程度** of the sentiment ranges between -1.0 (negative) and 1.0 (positive) and corresponds to the overall emotional leaning of the text.
# * **magnitude 语彩累积** indicates the overall strength of emotion (both positive and negative) within the given text, between 0.0 and +inf. Unlike score, magnitude is not normalized; each expression of emotion within the text (both positive and negative) contributes to the text's magnitude (so longer text blocks may have greater magnitudes).
#
#
# | Sentiment | Sample Values |
# |:-------------:|:-------------:|
# | 明显褒义 Clearly Positive | "score 褒贬程度": 0.8, "magnitude 语彩累积": 3.0 |
# | 明显贬义 Clearly Negative | "score 褒贬程度": -0.6, "magnitude 语彩累积": 4.0 |
# | 中性 Neutral | "score 褒贬程度": 0.1, "magnitude 语彩累积": 0.0 |
# | 混合 Mixed | "score 褒贬程度": 0.0, "magnitude 语彩累积": 4.0 |
#
# ### * 整篇消息文字的情感分析 (Sentiment analysis, Document level)
# +
# print(responses['documentSentiment'])
# -
print(u'[ 整篇消息 语种 : {} ]\n( 褒贬程度 : {} | 语彩累积 : {} )'.format(
responses['language']
, responses['documentSentiment']['score']
, responses['documentSentiment']['magnitude']
))
# ### * 语句的语法分析 (Syntax / Grammer analysis)
for i in range(len(responses['tokens'])):
print('')
print(responses['tokens'][i]['text']['content'])
print(responses['tokens'][i]['partOfSpeech'])
print(responses['tokens'][i]['dependencyEdge'])
# print(responses['tokens'][i]['text'])
# print(responses['tokens'][i]['lemma'])
# < End of interactive demo >
# ### 定义一个输出为NLP分析结果的文本消息的小功能,用于微信回复:
def KudosData_nlp_generate_reply(responses):
nlp_reply = u'[ NLP 自然语言处理结果 ]'
# 1. 整篇消息文字的情感分析 (Sentiment analysis, Document level)
nlp_reply += '\n'
nlp_reply += '\n' + u'[ 整篇消息 语种 : {} ]\n( 褒贬程度 : {} | 语彩累积 : {} )'.format(
responses['language']
, responses['documentSentiment']['score']
, responses['documentSentiment']['magnitude']
)
# 2. 消息文字中语句的情感分析 (Sentiment analysis, Sentence level)
nlp_reply += '\n'
for i in range(len(responses['sentences'])):
nlp_reply += '\n' + u'[ 语句 {} : {} ]\n( 褒贬程度 : {} | 语彩累积 : {} )'.format(
i+1
, responses['sentences'][i]['text']['content']
, responses['sentences'][i]['sentiment']['score']
, responses['sentences'][i]['sentiment']['magnitude']
)
# 3. 消息文字中名称实体的识别 (Name-Entity detection)
nlp_reply += '\n'
for i in range(len(responses['entities'])):
nlp_reply += '\n' + u'[ 实体 {} : {} ]\n 实体类别 : {}\n 重要程度 : {}'.format(
i+1
, responses['entities'][i]['name']
, responses['entities'][i]['type']
, responses['entities'][i]['salience']
)
if 'sentiment' in responses['entities'][i]:
nlp_reply += '\n' + u' 褒贬程度 : {}\n 语彩累积 : {}'.format(
responses['entities'][i]['sentiment']['score']
, responses['entities'][i]['sentiment']['magnitude']
)
if responses['entities'][i]['metadata'] != {}:
if 'wikipedia_url' in responses['entities'][i]['metadata']:
nlp_reply += '\n ' + responses['entities'][i]['metadata']['wikipedia_url']
# 4. 语句的语法分析 (Syntax / Grammer analysis)
# nlp_reply += '\n'
# for i in range(len(responses['tokens'])):
# nlp_reply += '\n' + str(responses['tokens'][i])
return nlp_reply
print(KudosData_nlp_generate_reply(responses))
# ### 用微信App扫QR码图片来自动登录
itchat.auto_login(hotReload=True) # hotReload=True: 退出程序后暂存登陆状态。即使程序关闭,一定时间内重新开启也可以不用重新扫码。
# Obtain my own Nick Name
MySelf = itchat.search_friends()
NickName4RegEx = '@' + MySelf['NickName'] + '\s*'
# 单聊模式,自动进行自然语言分析,以文本形式返回处理结果:
@itchat.msg_register([TEXT, MAP, CARD, NOTE, SHARING])
def text_reply(msg):
text4nlp = msg['Content']
# call NLP API:
nlp_responses = KudosData_nlp(text4nlp
, parm_nlp_extractDocumentSentiment
, parm_nlp_extractEntities
, parm_nlp_extractEntitySentiment
, parm_nlp_extractSyntax)
# Format NLP results:
nlp_reply = KudosData_nlp_generate_reply(nlp_responses)
print(nlp_reply)
return nlp_reply
# 群聊模式,如果收到 @ 自己的文字信息,会自动进行自然语言分析,以文本形式返回处理结果:
@itchat.msg_register(TEXT, isGroupChat=True)
def text_reply(msg):
if msg['isAt']:
text4nlp = re.sub(NickName4RegEx, '', msg['Content'])
# call NLP API:
nlp_responses = KudosData_nlp(text4nlp
, parm_nlp_extractDocumentSentiment
, parm_nlp_extractEntities
, parm_nlp_extractEntitySentiment
, parm_nlp_extractSyntax)
# Format NLP results:
nlp_reply = KudosData_nlp_generate_reply(nlp_responses)
print(nlp_reply)
return nlp_reply
itchat.run()
# interupt kernel, then logout
itchat.logout() # 安全退出
# ### 第四课:自然语言处理:语义和情感分析
# ### Lesson 4: Natural Language Processing 2
# * 消息文字中名称实体的识别 (Name-Entity detection)
# * 消息文字中语句的情感分析 (Sentiment analysis, Sentence level)
# * 整篇消息文字的情感分析 (Sentiment analysis, Document level)
# * 语句的语法分析 (Syntax / Grammer analysis)
# ### 下一课是:
# ### 第五课:视频识别和处理
# ### Lesson 5: Video Recognition & Processing
# * 识别视频消息中的物体名字 (Recognize objects in video)
# * 识别视频的场景 (Detect scenery in video)
# * 直接搜索视频内容 (Search content in video)
# <img src='http://www.kudosdata.com/wp-content/uploads/2016/11/cropped-KudosLogo1.png' width=30% style="float: right;">
# <img src='reference/WeChat_SamGu_QR.png' width=10% style="float: left;">
#
#
| uat_wechat/wechat_tool_testing/.ipynb_checkpoints/lesson_4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
import pandas as pd
import os
df = pd.read_csv('jena_climate_2009_2016.csv')
df.head()
# -
uni_data = df['T (degC)']
uni_data.index = df['Date Time']
uni_data.head()
TRAIN_SPLIT = 300000
uni_train_mean = uni_data[:TRAIN_SPLIT].mean()
uni_train_std = uni_data[:TRAIN_SPLIT].std()
uni_data = (uni_data-uni_train_mean)/uni_train_std
uni_data = uni_data.values
print(uni_data)
| jupyter/LSTM/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Data validation
#
# * https://adventofcode.com/2020/day/4
#
# We get to validate passports. Part 1 asks us to validate the fields; there are a number of required fields, and one optional. This is mostly a parsing task, however.
#
# The data for each passport is separated from the next by a blank line, so we just split the whole text by the doubled newline character (`\n`). Each passport is then trivially split into key-value pairs by splitting on arbitrary whitespace; the `str.split()` method doesn't care if the separators are newlines, spaces or some other whitespace! Each key-value pair is then split once on `:`, turning each passport entry into a dictionary.
#
# Now that we have dictionaries, we need to validate the keys in them. I'm making use of the fact that Python's [`dict.keys()` keys view object](https://docs.python.org/library/stdtypes.html#dict-views) acts as a *set*, and testing each is a [superset](https://docs.python.org/3/library/stdtypes.html#frozenset.issuperset) of the required field names, as well as being a subset of all possible field names. Python's chained operators make this a very simple and elegant expression:
#
# ```python
# required = frozenset(...) # all required keys
# all_ = required | frozenset({"cid"}) # required plus optional keys
# all_ >= passport.keys() >= required # true if all required keys are there, and no unknown keys
# ```
# +
from typing import Callable, Iterable, Mapping
PassportData = Mapping[str, str]
Validator = Callable[[PassportData], bool]
required = frozenset({"byr", "iyr", "eyr", "hgt", "hcl", "ecl", "pid"})
all_ = required | frozenset({"cid"})
def valid_passport(passport: PassportData) -> bool:
return all_ >= passport.keys() >= required
def read_passports(data: str) -> Iterable[PassportData]:
for block in data.split("\n\n"):
yield dict(f.split(':', 1) for f in block.split())
def count_valid(passports: Iterable[PassportData], validator: Validator=valid_passport) -> int:
return sum(1 for _ in filter(validator, passports))
testdata = """\
ecl:gry pid:860033327 eyr:2020 hcl:#fffffd
byr:1937 iyr:2017 cid:147 hgt:183cm
iyr:2013 ecl:amb cid:350 eyr:2023 pid:028048884
hcl:#cfa07d byr:1929
hcl:#ae17e1 iyr:2013
eyr:2024
ecl:brn pid:760753108 byr:1931
hgt:179cm
hcl:#cfa07d eyr:2025 pid:166559648
iyr:2011 ecl:brn hgt:59in
"""
assert count_valid(read_passports(testdata)) == 2
# -
import aocd
passportdata = aocd.get_data(day=4, year=2020)
print("Part 1:", count_valid(read_passports(passportdata)))
# ## Value validation
#
# To validate the values, I reached for a tool I use quite often: a schema validation library called [Marshmallow](https://marshmallow.readthedocs.io/). It makes it trivial to define validators for each field; only the height validation required 'custom' code:
# +
from marshmallow import fields, validate, RAISE, Schema, ValidationError
def validate_height(height: str) -> bool:
try:
value = int(height[:-2])
except ValueError:
raise ValidationError("Invalid height")
if height[-2:] == "cm" and (150 <= value <= 193):
return
elif height[-2:] == "in" and (59 <= value <= 76):
return
raise ValidationError("Invalid height")
class PassportSchema(Schema):
class Meta:
unknown = 'RAISE'
byr = fields.Int(required=True, validate=validate.Range(1920, 2003))
iyr = fields.Int(required=True, validate=validate.Range(2010, 2021))
eyr = fields.Int(required=True, validate=validate.Range(2020, 2031))
hgt = fields.Str(required=True, validate=validate_height)
hcl = fields.Str(required=True, validate=validate.Regexp(r"^#[0-9a-fA-F]{6}$"))
ecl = fields.Str(
required=True,
validate=validate.OneOf(frozenset("amb blu brn gry grn hzl oth".split())),
)
pid = fields.Str(required=True, validate=validate.Regexp(r"^\d{9}$"))
cid = fields.Str()
def valid_passport_fields(passport: Mapping):
try:
PassportSchema().load(passport)
return True
except ValidationError:
return False
testinvalid = """\
eyr:1972 cid:100
hcl:#18171d ecl:amb hgt:170 pid:186cm iyr:2018 byr:1926
iyr:2019
hcl:#602927 eyr:1967 hgt:170cm
ecl:grn pid:012533040 byr:1946
hcl:dab227 iyr:2012
ecl:brn hgt:182cm pid:021572410 eyr:2020 byr:1992 cid:277
hgt:59cm ecl:zzz
eyr:2038 hcl:74454a iyr:2023
pid:3556412378 byr:2007
"""
assert count_valid(read_passports(testinvalid), valid_passport_fields) == 0
testvalid = """\
pid:087499704 hgt:74in ecl:grn iyr:2012 eyr:2030 byr:1980
hcl:#623a2f
eyr:2029 ecl:blu cid:129 byr:1989
iyr:2014 pid:896056539 hcl:#a97842 hgt:165cm
hcl:#888785
hgt:164cm byr:2001 iyr:2015 cid:88
pid:545766238 ecl:hzl
eyr:2022
iyr:2010 hgt:158cm hcl:#b6652a ecl:blu byr:1944 eyr:2021 pid:093154719
"""
count_valid(read_passports(testvalid), valid_passport_fields) == 4
# -
print("Part 2:", count_valid(read_passports(passportdata), valid_passport_fields))
| 2020/Day 04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GraphNets (Pytorch)
# ## Utils
# +
import itertools
def pairwise(iterable):
"""s -> (s0,s1), (s1,s2), (s2, s3), ..."""
a, b = itertools.tee(iterable)
next(b, None)
return zip(a, b)
# +
# GraphTuple
from collections import namedtuple
from functools import partial
from typing import Callable
from typing import List
from typing import Tuple
from typing import Dict
import networkx as nx
import numpy as np
import torch
GraphTuple = namedtuple(
"GraphTuple",
[
"node_attr", # node level attributes
"edge_attr", # edge level attributes
"global_attr", # global level attributes
"edges", # node-to-node connectivity
"node_indices", # tensor where each element indicates the index of the graph the node_attr belongs to
"edge_indices",
# tensor where each element indicates the index of the graph that the edge_attr and edges belong to.
],
)
def pick_edge(graphs):
for g in graphs:
for x in g.edges(data=True):
if x[-1] is not None:
return x
def pick_node(graphs):
for g in graphs:
for x in g.nodes(data=True):
if x[-1] is not None:
return x
def to_graph_tuple(
graphs: List[nx.DiGraph],
feature_key: str = "features",
global_attr_key: str = "data",
device: str = None,
) -> GraphTuple:
"""Convert a list og networkx graphs into a GraphTuple.
:param graphs: list of graphs
:param feature_key: key to find the node, edge, and global features
:param global_attr_key: attribute on the NetworkX graph to find the global data (default: 'data')
:return: GraphTuple, a namedtuple of ['node_attr', 'edge_attr', 'global_attr',
'edges', 'node_inices', 'edge_indices']
"""
n_edges = 0
n_nodes = 0
for graph in graphs:
n_edges += graph.number_of_edges()
n_nodes += graph.number_of_nodes()
n = len(graphs)
node_idx = np.empty(n_nodes)
edge_idx = np.empty(n_edges)
edge = pick_edge(graphs)
if edge:
edata = edge[-1][feature_key]
else:
edata = np.empty(1)
node = pick_node(graphs)
if node:
vdata = node[-1][feature_key]
else:
vdata = np.empty(1)
if hasattr(graph, global_attr_key):
udata = getattr(graph, global_attr_key)[feature_key]
else:
udata = np.zeros(1)
connectivity = np.empty((n_edges, 2))
v = np.empty((n_nodes, *tuple(vdata.shape)))
e = np.empty((n_edges, *tuple(edata.shape)))
u = np.empty((n, *tuple(udata.shape)))
_v = 0
_e = 0
ndict = {}
for gidx, graph in enumerate(graphs):
for node, ndata in graph.nodes(data=True):
v[_v] = ndata[feature_key]
ndict[node] = _v
node_idx[_v] = gidx
_v += 1
for n1, n2, edata in graph.edges(data=True):
e[_e] = edata[feature_key]
edge_idx[_e] = gidx
connectivity[_e] = [ndict[n1], ndict[n2]]
_e += 1
if hasattr(graph, global_attr_key):
u[gidx] = getattr(graph, global_attr_key)[feature_key]
else:
u[gidx] = 0
result = GraphTuple(
torch.tensor(v, dtype=torch.float),
torch.tensor(e, dtype=torch.float),
torch.tensor(u, dtype=torch.float),
torch.tensor(connectivity, dtype=torch.long),
torch.tensor(node_idx, dtype=torch.long),
torch.tensor(edge_idx, dtype=torch.long),
)
if device:
return GraphTuple(*[x.to(device) for x in result])
return result
def from_graph_tuple(gt: GraphTuple, feature_key: str = 'features') -> Dict[int, nx.DiGraph]:
graph_dict = {}
for node, (gidx, ndata) in enumerate(zip(gt.node_indices, gt.node_attr)):
graph_dict.setdefault(gidx.item(), nx.DiGraph())
g = graph_dict[gidx.item()]
g.add_node(node, **{'features': ndata})
for gidx, (n1, n2), edata in zip(gt.edge_indices, gt.edges, gt.edge_attr):
graph_dict.setdefault(gidx.item(), nx.DiGraph())
g = graph_dict[gidx.item()]
g.add_edge(n1.item(), n2.item(), **{'features': edata})
return graph_dict
def collate_tuples(tuples: List[Tuple], func: Callable[[List[Tuple]], Tuple]):
"""Collate elements of many tuples using a function. All of the first
elements for all the tuples will be collated using the function, then all
of the second elements of all tuples, and so on.
:param tuples: list of tuples
:param func: callable
:return: tuple of same type
"""
def f(x):
if x is None:
return None
else:
return func(x)
t = type(tuples[0])
if t is tuple:
def t(*args):
return tuple(args)
return t(*[f(x) for x in zip(*tuples)])
def replace_key(graph_tuple, data: dict):
"""Replace the values of the graph tuple.
DOES NOT REPLACE IN PLACE.
"""
values = []
for k, v in zip(graph_tuple._fields, graph_tuple):
if k in data:
v = data[k]
values.append(v)
return GraphTuple(*values)
def apply_to_tuple(x, func: Callable[[List[Tuple]], Tuple]):
"""Apply function to each element of the tuple."""
return type(x)(*[func(x) for x in x])
def print_graph_tuple_shape(graph_tuple):
for field, x in zip(graph_tuple._fields, graph_tuple):
print(field, " ", x.shape)
def cat_gt(*gts: Tuple[GraphTuple, ...]) -> GraphTuple:
"""Concatenate graph tuples along dimension=1.
Edges, node idx and edge idx are simply copied over.
"""
cat = partial(torch.cat, dim=1)
return GraphTuple(
cat([gt.node_attr for gt in gts]),
cat([gt.edge_attr for gt in gts]),
cat([gt.global_attr for gt in gts]),
gts[0].edges,
gts[0].node_indices,
gts[0].edge_indices,
)
def gt_to_device(x: Tuple, device):
return GraphTuple(*[v.to(device) for v in x])
class InvalidGraphTuple(Exception):
pass
def validate_gt(gt: GraphTuple):
if not isinstance(gt, GraphTuple):
raise InvalidGraphTuple("{} is not a {}".format(gt, GraphTuple))
if not gt.edge_attr.shape[0] == gt.edges.shape[0]:
raise InvalidGraphTuple(
"Edge attribute shape {} does not match edges shape {}".format(
gt.edge_attr.shape, gt.edges.shape
)
)
if not gt.edge_attr.shape[0] == gt.edge_indices.shape[0]:
raise InvalidGraphTuple(
"Edge attribute shape {} does not match edge idx shape {}".format(
gt.edge_attr.shape, gt.edge_indices.shape
)
)
if not gt.node_attr.shape[0] == gt.node_indices.shape[0]:
raise InvalidGraphTuple(
"Node attribute shape {} does not match node idx shape {}".format(
gt.node_attr.shape, gt.node_indices.shape
)
)
# edges cannot refer to non-existent nodes
if gt.edges.shape[0] and not (gt.edges.max().item() < gt.node_attr.shape[0]):
raise InvalidGraphTuple(
"Edges reference node {} which does not exist nodes of size {}".format(
gt.edges.max(), gt.node_attr.shape[0]
)
)
if gt.edges.shape[0] and not (gt.edges.min().item() >= 0):
raise InvalidGraphTuple(
"Node index must be greater than 0, not {}".format(gt.edges.min())
)
# -
# ## Data
# +
from torch import nn
from torch.utils.data import DataLoader
class GraphMask(object):
def __init__(self, node_mask, edge_mask, global_mask):
self.x_mask = node_mask,
self.e_mask = edge_mask
self.g_mask = global_mask
class GraphData(object):
"""Data representing a single graph"""
__slots__ = ['x', 'e', 'g', 'edges']
def __init__(self, node_attr, edge_attr, global_attr, edges):
self.x = node_attr
self.e = edge_attr
self.g = global_attr
self.edges = edges
GraphData.debug(self)
def debug(self):
if self.edges.max() >= self.x.shape[0]:
raise RuntimeError("Edge coordinate {} is greater than number of nodes {}".format(self.edges.max(), self.x.shape[0
]))
if not self.edges.shape[1] == self.e.shape[0]:
raise RuntimeError("Number of edges {} must match number of edge attributes {}".format(
self.edges.shape[1],
self.e.shape[0]
))
if not self.edges.dtype == torch.long:
raise RunTimeError("Wrong tensor type. `edges` must be dtype={} not {}".format(self.edges.dtype, torch.long))
def apply(self, func):
for f in self.__slots__:
func(f)
def to(self, device: str):
self.apply(lambda x: x.to(device))
def contiguous(self):
self.apply(lambda x: x.contiguous())
@property
def num_graphs(self):
return self.g.shape[0]
@property
def num_nodes(self):
return self.x.shape[0]
@property
def node_shape(self):
return self.x.shape[1:]
@property
def edge_shape(self):
return self.e.shape[1:]
@property
def global_shape(self):
return self.g.shape[1:]
@property
def shape(self):
return self.x.shape[1:] + self.e.shape[1:] + self.g.shape[1:]
@property
def size(self):
return self.x.shape[:1] + self.e.shape[:1] + self.g.shape[:1]
def _mask_fields(self, masks: Dict[str, torch.tensor]):
for m in masks:
if m not in self.__slots__:
raise RuntimeError("{} is not a valid field".format(m))
masked_fields = []
for field in self.__slots__:
if field not in masks or masks[field] is None:
masked_fields.append(getattr(self, field))
else:
masked_fields.append(getattr(self, field)[:, masks[field]])
return masked_fields
def mask(self, node_mask, edge_mask, global_mask, invert: bool = False):
d = {'x': node_mask, 'e': edge_mask, 'g': global_mask}
if invert:
d = {k: ~v for k, v in d.items()}
return self.__class__(
*self._mask_fields(d)
)
def clone(self):
return self.__class__(
*[getattr(self, field).clone() for field in self.__class__.__slots__]
)
def __repr__(self):
return "<{cls} size(n,e,g)={size} features(n,e,g)={shape}>".format(
cls=self.__class__.__name__,
n_graphs=self.num_graphs,
size=self.x.shape[:1] + self.e.shape[:1] + self.g.shape[:1],
shape=self.shape
)
class GraphBatch(GraphData):
__slots__ = GraphData.__slots__ + ['node_idx', 'edge_idx']
def __init__(self, node_attr, edge_attr, global_attr, edges, node_idx, edge_idx):
super(GraphBatch, self).__init__(node_attr, edge_attr, global_attr, edges)
self.node_idx = node_idx
self.edge_idx = edge_idx
GraphBatch.debug(self)
@staticmethod
def _same(a):
return min(a) == max(a)
def debug(self):
super().debug()
if not self.node_idx.dtype == torch.long:
raise RunTimeError("Wrong tensor type. `node_idx` must be dtype={} not {}".format(self.node_idx.dtype, torch.long))
if not self.edge_idx.dtype == torch.long:
raise RunTimeError("Wrong tensor type. `edge_idx` must be dtype={} not {}".format(self.edge_idx.dtype, torch.long))
if not self.node_idx.max() == self.edge_idx.max():
raise RuntimeError("Number of graphs in node_idx and edge_idx mismatch")
if not self.node_idx.min() == 0:
raise RuntimeError("Minimum graph index (node_idx.min()) must start at 0, not {}".format(self.node_idx.min()))
if not self.edge_idx.min() == 0:
raise RuntimeError("Minimum graph index (edge_idx.min()) must start at 0, not {}".format(self.edge_idx.min()))
@classmethod
def from_data_list(cls, data_list):
# checks
n_features = []
e_features = []
g_features = []
for data in data_list:
n_features.append(data.x.shape[1])
e_features.append(data.e.shape[1])
g_features.append(data.g.shape[0])
if not cls._same(n_features):
raise RuntimeError("Node feature dimensions must all be the same")
if not cls._same(e_features):
raise RuntimeError("Edge feature dimensions must all be the same")
if not cls._same(g_features):
raise RuntimeError("Global feature dimensions must all be the same")
node_repeats = torch.tensor([data.x.shape[0] for data in data_list])
edge_repeats = torch.tensor([data.e.shape[0] for data in data_list])
node_idx = torch.repeat_interleave(torch.range(0, node_repeats.shape[0]-1, dtype=torch.long), node_repeats)
edge_idx = torch.repeat_interleave(torch.range(0, edge_repeats.shape[0]-1, dtype=torch.long), edge_repeats)
# concatenate edges
edges = torch.cat([data.edges for data in data_list], dim=1)
# cumulated shapes
c = torch.cumsum(torch.tensor([0] + [data.x.shape[0] for data in data_list[:-1]]), dim=0)
delta = torch.repeat_interleave(c, edge_repeats).repeat(2, 1)
# shift concatenated edges
edges = edges + delta
return cls(
node_attr=torch.cat([data.x for data in data_list]),
edge_attr=torch.cat([data.e for data in data_list]),
global_attr = torch.cat([data.g for data in data_list]),
edges=edges,
node_idx=node_idx,
edge_idx=edge_idx
)
# def mask(self, node_mask = None, edge_mask = None, global_mask = None):
# masked = self._mask_fields({'x': node_mask, 'e': edge_mask, 'g': global_mask})
# return self.__class__(*masked)
# def __repr__(self):
# return "<{cls} size(n,e,g)={size} features(n,e,g)={shape}>".format(
# cls=self.__class__.__name__,
# n_graphs=self.num_graphs,
# size=self.x.shape[:1] + self.e.shape[:1] + self.g.shape[:1],
# shape=self.shape
# )
data1 = GraphData(
torch.randn(10, 5),
torch.randn(3, 4),
torch.randn(1, 3),
torch.randint(0, 10, torch.Size([2, 3]))
)
data2 = GraphData(
torch.randn(10, 5),
torch.randn(5, 4),
torch.randn(1, 3),
torch.randint(0, 10, torch.Size([2, 5]))
)
batch = GraphBatch.from_data_list([data1, data2])
# +
def foo(*args):
print(*args)
foo(*(3,2,3), *(3,))
# -
# test clone
data1.clone()
batch.clone()
# +
# test masks
data1.mask(
torch.tensor([True, True, False, False, False]),
torch.tensor([True, True, False, False]),
torch.tensor([True, False, False])
)
batch.mask(
torch.tensor([True, True, False, False, False]),
torch.tensor([True, True, False, False]),
torch.tensor([True, False, False])
)
# +
def random_data(n_features, e_features, g_features):
n_nodes = torch.randint(1, 10, torch.Size([])).item()
n_edges = torch.randint(1, 20, torch.Size([])).item()
return GraphData(
torch.randn(n_nodes, n_features),
torch.randn(n_edges, e_features),
torch.randn(1, g_features),
torch.randint(0, n_nodes, torch.Size([2, n_edges]))
)
random_data(5, 3, 4)
# +
from torch.utils.data import DataLoader
def collate(data_list):
if isinstance(data_list[0], tuple):
if issubclass(type(data_list[0][0]), GraphData):
return tuple([collate([x[i] for x in data_list]) for i in range(len(data_list[0]))])
else:
raise RuntimeError("Cannot collate {}({})({})".format(type(data_list), type(data_list[0]), type(data_list[0][0])))
return GraphBatch.from_data_list(data_list)
class GraphDataLoader(DataLoader):
def __init__(self, dataset, batch_size=1, shuffle=False,
**kwargs):
super().__init__(dataset, batch_size, shuffle,
collate_fn=collate, **kwargs)
# +
data_list = [random_data(5, 4, 3) for _ in range(2000)]
# test data list loader
loader = GraphDataLoader(data_list)
for s in loader:
pass
# test data list loader of tuples
loader = GraphDataLoader(list(zip(data_list, data_list)))
for s in loader:
pass
# -
# ### From Networkx
# +
from itertools import tee
next(tee(g.edges(data=True))[0])
next((x for x in g.edges(data=True)))
# +
import networkx as nx
import itertools
def _first(i):
"""Select the first element in an iterable"""
return next((x for x in itertools.tee(i)[0]))
def to_graph_data(g: nx.DiGraph,
n_node_feat: Optional[int] = None,
n_edge_feat: Optional[int] = None,
n_glob_feat: Optional[int] = None):
if hasattr(g, 'data'):
gdata = g.data
else:
gdata = {}
if n_node_feat is None:
_, ndata = _first(g.nodes(data=True))
n_node_feat = ndata['features'].shape[0]
if n_edge_feat is None:
_, _, edata = _first(g.edges(data=True))
n_edge_feat = edata['features'].shape[0]
if n_glob_feat is None:
n_glob_feat = gdata['features'].shape[0]
n_nodes = g.number_of_nodes()
n_edges = g.number_of_edges()
node_attr = np.empty((n_nodes, n_node_feat))
edge_attr = np.empty((n_edges, n_edge_feat))
glob_attr = np.empty((1, n_glob_feat))
nodes = sorted(list(g.nodes(data=True)))
ndict = {}
for i, (n, ndata) in enumerate(nodes):
node_attr[i] = ndata['features']
ndict[n] = i
edges = np.empty((2, n_edges))
for i, (n1, n2, edata) in enumerate(g.edges(data=True)):
edges[:, i] = np.array([ndict[n1], ndict[n2]])
edge_attr[i] = edata['features']
glob_attr[0] = g.data['features']
return GraphData(torch.tensor(node_attr, dtype=torch.float),
torch.tensor(edge_attr, dtype=torch.float),
torch.tensor(glob_attr, dtype=torch.float),
torch.tensor(edges, dtype=torch.long))
g = nx.DiGraph()
g.add_node(0, features=np.array([0, 0, 0]))
g.add_node(1, features=np.array([1,2,3]))
g.add_edge(0, 1, features=np.array([0, 1]))
g.data = {'features': np.array([0,])}
data = to_graph_data(g)
batch = GraphBatch.from_data_list([data, data, data])
# +
# optimized methods for grouping tensors
import torch
from typing import Dict, List, Tuple
@torch.jit.script
def unique_with_counts(idx, grouped: Dict[int, int]):
for x in idx:
if x.item() not in grouped:
grouped[x.item()] = 1
else:
grouped[x.item()] += 1
counts = torch.zeros(len(grouped), dtype=torch.long)
values = torch.empty(len(grouped), dtype=idx.dtype)
for i, (k, v) in enumerate(grouped.items()):
values[i] = k
counts[i] = v
a = torch.argsort(values)
return values[a], counts[a]
@torch.jit.script
def _jit_scatter_group(x: torch.Tensor, idx: torch.Tensor, d: Dict[int, int]) -> Tuple[torch.Tensor, List[torch.Tensor]]:
x = x[torch.argsort(idx)]
groups, b = unique_with_counts(idx, d)
i_a = 0
arr_list = []
for i_b in b:
arr_list.append(x[i_a:i_a + i_b.item()])
i_a += i_b.item()
return groups, arr_list
def scatter_group(x: torch.Tensor, idx: torch.Tensor) -> Tuple[torch.Tensor, List[torch.Tensor]]:
return _jit_scatter_group(x, idx, {})
# +
def graph_batch_to_data_list(batch: GraphBatch):
assert issubclass(type(batch), GraphBatch)
gidx_n, node_attr = scatter_group(batch.x, batch.node_idx)
gidx_e, edge_attr = scatter_group(batch.e, batch.edge_idx)
gidx_edge, edges = scatter_group(batch.edges.T, batch.edge_idx)
def to_dict(a, b):
return dict(zip([x.item() for x in a], b))
ndict = to_dict(gidx_n, node_attr)
edict = to_dict(gidx_e, edge_attr)
edgesdict = to_dict(gidx_edge, edges)
datalist = []
for k in ndict:
_edges = edgesdict[k].T - edgesdict[k].min()
data = GraphData(ndict[k], edict[k], batch.g[k], _edges)
datalist.append(data)
return datalist
def graph_data_to_nx(data: GraphData):
g = nx.DiGraph()
for n, ndata in enumerate(data.x):
g.add_node(n, **{'features': ndata})
for i, e in enumerate(data.edges.T):
g.add_edge(e[0], e[1], **{'features': data.e[i]})
g.data = {'features': data.g}
return g
for data in graph_batch_to_data_list(batch):
g = graph_data_to_nx(data)
nx.draw(g)
# -
print(np.random.uniform(0, 10, (4)))
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Blocks
# -
# ### Common Blocks
# +
from functools import wraps
from typing import *
import torch
import torch_scatter
from torch import nn
class MLPBlock(nn.Module):
"""A multilayer perceptron block."""
def __init__(self, input_size: int, output_size: int = None):
super().__init__()
if output_size is None:
output_size = input_size
self.blocks = nn.Sequential(
nn.Linear(input_size, output_size), nn.ReLU(), nn.LayerNorm(output_size)
)
def forward(self, x):
return self.blocks(x)
class BlockRepeat(nn.Module):
"""A repeating block"""
def __init__(self, module: nn.Module, *latent_sizes: List[int]):
super().__init__()
self.blocks = nn.Sequential(
*[module(n1, n2) for n1, n2 in pairwise(latent_sizes)]
)
def forward(self, x):
return self.blocks(x)
class MLP(BlockRepeat):
"""
Repeating MLP block
Usage:
.. code-block:: python
MLP([10, 16, 10])
"""
def __init__(self, *latent_sizes: List[int]):
for size in latent_sizes:
assert isinstance(size, int)
super().__init__(MLPBlock, *latent_sizes)
class Aggregator(nn.Module):
"""Aggregation layer."""
def __init__(self, aggregator: str, dim: int = None, dim_size: int = None):
super().__init__()
self.valid_aggregators = {
"mean": torch_scatter.scatter_mean,
"max": self.scatter_max,
"min": self.scatter_min,
"add": torch_scatter.scatter_add,
}
if aggregator not in self.valid_aggregators:
raise ValueError(
"Aggregator '{}' not not one of the valid aggregators {}".format(
aggregator, self.valid_aggregators
)
)
self.aggregator = aggregator
self.kwargs = dict(dim=dim, dim_size=dim_size)
def forward(self, x, indices, **kwargs):
func_kwargs = dict(self.kwargs)
func_kwargs.update(kwargs)
func = self.valid_aggregators[self.aggregator]
result = func(x, indices, **func_kwargs)
return result
@staticmethod
@wraps(torch_scatter.scatter_max)
def scatter_max(*args, **kwargs):
return torch_scatter.scatter_max(*args, **kwargs)[0]
@staticmethod
@wraps(torch_scatter.scatter_min)
def scatter_min(*args, **kwargs):
return torch_scatter.scatter_min(*args, **kwargs)[0]
class Block(nn.Module):
def __init__(self, module_dict: Dict[str, nn.Module], independent: bool):
super().__init__()
self._independent = independent
self.block_dict = nn.ModuleDict(
{name: mod for name, mod in module_dict.items() if mod is not None}
)
@property
def out_dim(self):
pass
# -
# ### Flexible Blocks
# +
from functools import wraps
from typing import Any
from typing import Dict
from typing import Tuple
from typing import Type
import torch
class FlexDim:
def __init__(self, pos: int = 0, dim: int = 1):
"""Flexible dimension to be used in conjunction with `FlexBlock`
:param pos: position of the input arguments that contains the input data
:param dim: dimension to use for the input shape
"""
self.pos = pos
self.dim = dim
def resolve(self, input_args, input_kwargs):
d = input_args[self.pos].shape[self.dim]
if d == 0:
raise ValueError("Dimension cannot be zero")
return d
class FlexBlock(torch.nn.Module):
def __init__(self, module_fn, *args, **kwargs):
super().__init__()
self.module = module_fn
self.args = args
self.kwargs = kwargs
self.resolved_module = None
def resolve_args(self, input_args: Tuple[Any, ...], input_kwargs: Dict[str, Any]):
rargs = []
for i, a in enumerate(self.args):
if isinstance(a, FlexDim):
rargs.append(a.resolve(input_args, input_kwargs))
elif a is FlexDim:
raise ValueError("Found {}. Initialize FlexDim to use flexible dimensions, `Flex.d()` or `FlexDim()`".format(
a))
else:
rargs.append(a)
return rargs
def resolve_kwargs(self, input_args: Tuple[Any, ...], input_kwargs: Dict[str, Any]):
return self.kwargs
def resolve(self, args: Tuple[Any, ...], kwargs: Dict[str, Any]):
resolved_args = self.resolve_args(args, kwargs)
resolved_kwargs = self.resolve_kwargs(args, kwargs)
self.resolved_module = self.module(*resolved_args, **resolved_kwargs)
def forward(self, *args, **kwargs):
if self.resolved_module is None:
self.resolve(args, kwargs)
return self.resolved_module(*args, **kwargs)
class Flex:
d = FlexDim
def __init__(self, module_type: Type[torch.nn.Module]):
"""Initialize a module as a FlexBlock with flexible dimensions.
Usage:
.. code-block:: python
Flex(torch.nn.Linear)(Flex.d(), 25)
:param module_type: module type (e.g. `torch.nn.Linear`
"""
self.module_type = module_type
self.__call__ = wraps(module_type.__init__)(self.__class__.__call__)
def __call__(self, *args, **kwargs) -> torch.nn.Module:
"""Initialize the flexible module.
:param args: the initialization arguments
:param kwargs: the initialization keyword arguments
:return: initialized torch.nn.Module
"""
return FlexBlock(self.module_type, *args, **kwargs)
# -
Flex(MLP)(Flex.d(), 16, 10)(torch.randn(10, 5))
# ### Graph Blocks
# +
class EdgeBlock(Block):
def __init__(self, mlp: nn.Module, independent: bool):
super().__init__({"mlp": mlp}, independent=independent)
def forward(self, edge_attr: torch.tensor, src: torch.tensor = None, dest: torch.tensor = None):
if not self._independent:
out = torch.cat([src, dest, edge_attr], 1)
else:
out = edge_attr
results = self.block_dict["mlp"](out)
return results
# TODO: concatenate global features for Edge and Node block
class NodeBlock(Block):
def __init__(
self, mlp: nn.Module, independent: bool, edge_aggregator: Aggregator = None
):
"""
:param input_size:
:param layers:
:param edge_aggregator:
:param independent:
"""
super().__init__(
{"edge_aggregator": edge_aggregator, "mlp": mlp}, independent=independent
)
def forward(self, v, edge_index, edge_attr, u, node_idx, edge_idx):
if not self._independent:
row, col = edge_index
aggregator_fn = self.block_dict["edge_aggregator"]
if aggregator_fn:
aggregated = self.block_dict["edge_aggregator"](
edge_attr, col, dim=0, dim_size=v.size(0)
)
out = torch.cat([aggregated, v], dim=1)
else:
out = torch.cat([v], dim=1)
else:
out = v
return self.block_dict["mlp"](out)
class GlobalBlock(Block):
def __init__(
self,
mlp,
independent: bool,
node_aggregator: Aggregator = None,
edge_aggregator: Aggregator = None,
):
super().__init__(
{
"node_aggregator": node_aggregator,
"edge_aggregator": edge_aggregator,
"mlp": mlp,
},
independent=independent,
)
def forward(self, node_attr, edge_index, edge_attr, u, node_idx, edge_idx):
if not self._independent:
node_agg = self.block_dict["node_aggregator"]
edge_agg = self.block_dict["edge_aggregator"]
to_cat = [u]
if node_agg is not None:
to_cat.append(node_agg(node_attr, node_idx, dim=0, dim_size=u.shape[0]))
if edge_agg is not None:
to_cat.append(edge_agg(edge_attr, edge_idx, dim=0, dim_size=u.shape[0]))
try:
out = torch.cat(to_cat, dim=1)
except RuntimeError as e:
raise e
else:
out = u
return self.block_dict["mlp"](out)
# -
# ## Models
class Block(nn.Module):
def __init__(self, module_dict: Dict[str, nn.Module], independent: bool):
super().__init__()
self._independent = independent
self.block_dict = nn.ModuleDict(
{name: mod for name, mod in module_dict.items() if mod is not None}
)
@property
def independent(self):
return self._independent
# ### Edge Block
# +
class EdgeBlock(Block):
def __init__(self, mlp: nn.Module):
super().__init__({"mlp": mlp}, independent=True)
def forward(self, edge_attr: torch.tensor, node_attr: torch.tensor = None, edges: torch.tensor = None):
results = self.block_dict["mlp"](edge_attr)
return results
def forward_from_data(self, data: GraphData):
return self(data.e, data.x, data.edges)
class AggregatingEdgeBlock(EdgeBlock):
def __init__(self, mlp: nn.Module):
super().__init__(mlp)
self._independent = False
def forward(self, edge_attr: torch.tensor, node_attr: torch.tensor, edges: torch.tensor):
out = torch.cat([node_attr[edges[0]], node_attr[edges[1]], edge_attr], 1)
return self.block_dict['mlp'](out)
def forward_from_data(self, data: GraphData):
return self(data.e, data.x. data.edges)
# +
# demonstrate independent edge block
edge_encoder = EdgeBlock(MLP(3, 10, 16))
x = torch.randn(20, 3)
out = edge_encoder(x)
print(out.shape)
assert out.shape == torch.Size([20, 16])
for p in edge_encoder.parameters():
print(p)
print(p.requires_grad)
# +
# demonstrate dependent edge block
edge_model = AggregatingEdgeBlock(MLP(7, 10, 16))
x = torch.randn(20, 3)
edges = torch.randint(0, 40, torch.Size([2, 20]))
n = torch.randn(40, 2)
assert edge_model(x, n, edges).shape == torch.Size([20, 16])
for p in edge_model.parameters():
print(p)
print(p.requires_grad)
# -
# ### Node Block
# +
# TODO: this is redundent with EdgeBlock
class NodeBlock(Block):
def __init__(self, mlp: nn.Module):
super().__init__({
'mlp': mlp
}, independent=True)
def forward(self, node_attr):
return self.block_dict['mlp'](node_attr)
def forward_from_data(self, data: GraphBatch):
return self(data.x)
class AggregatingNodeBlock(NodeBlock):
def __init__(self, mlp: nn.Module, edge_aggregator: Aggregator):
super().__init__(mlp)
self.block_dict['edge_aggregator'] = edge_aggregator
self._independent = False
# TODO: source_to_target, target_to_source
def forward(self, node_attr, edge_attr, edges):
aggregated = self.block_dict['edge_aggregator'](edge_attr, edges[1], dim=0, dim_size=node_attr.size(0))
out = torch.cat([node_attr, aggregated], dim=1)
return self.block_dict['mlp'](out)
def forward_from_data(self, data: GraphBatch):
return self(data.x, data.e, data.edges)
# +
# test NodeBlock
node_encoder = NodeBlock(MLP(5, 16, 10))
node_attr = torch.randn(10, 5)
for p in node_encoder.parameters():
print(p)
print(p.requires_grad)
# +
# test AggregatingNodeBlock
node_model = AggregatingNodeBlock(MLP(5, 16, 10), Aggregator('mean'))
edge_attr = torch.randn(20, 3)
edges = torch.randint(0, 40, torch.Size([2, 20]))
node_attr = torch.randn(40, 2)
node_model(node_attr, edge_attr, edges).shape
for p in node_model.parameters():
print(p)
print(p.requires_grad)
# -
# ### Global Block
# +
class GlobalBlock(Block):
def __init__(self, mlp):
super().__init__({'mlp': mlp}, independent=True)
def forward(self, global_attr):
return self.block_dict['mlp'](global_attr)
def forward_from_data(self, data: GraphBatch):
return self(data.g)
class AggregatingGlobalBlock(GlobalBlock):
def __init__(self, mlp, edge_aggregator = None, node_aggregator = None):
super().__init__(mlp)
self.block_dict['edge_aggregator'] = edge_aggregator
self.block_dict['node_aggregator'] = node_aggregator
self._independent = False
def forward(self, global_attr, node_attr, edge_attr, edges, node_idx, edge_idx):
aggregated = [global_attr]
if 'node_aggregator' in self.block_dict:
aggregated.append(self.block_dict['node_aggregator'](node_attr, node_idx, dim=0, dim_size=global_attr.shape[0]))
if 'edge_aggregator' in self.block_dict:
aggregated.append(self.block_dict['edge_aggregator'](edge_attr, edge_idx, dim=0, dim_size=global_attr.shape[0]))
out = torch.cat(aggregated, dim=1)
return self.block_dict['mlp'](out)
def forward_from_data(self, data: GraphBatch):
return self(data.g, data.x, data.e, data.edges, data.node_idx, data.edge_idx)
# +
# test GlobalBlock
# +
# test GlobalBlock
global_encoder = GlobalBlock(MLP(3, 10))
for p in global_encoder.parameters():
print(p)
print(p.requires_grad)
global_attr = torch.randn(10, 3)
global_encoder(global_attr).shape
for p in global_encoder.parameters():
print(p)
print(p.requires_grad)
# +
# test AggregatingGlobalBlock
global_attr = torch.randn(10, 3)
edge_attr = torch.randn(20, 3)
edges = torch.randint(0, 40, torch.Size([2, 20]))
node_attr = torch.randn(40, 2)
node_idx = torch.randint(0, 3, torch.Size([40]))
edge_idx = torch.randint(0, 3, torch.Size([20]))
global_model = AggregatingGlobalBlock(MLP(8, 16, 10), Aggregator('mean'), Aggregator('mean'))
out = global_model(global_attr, node_attr, edge_attr, edges, node_idx, edge_idx)
for p in global_model.parameters():
print(p)
print(p.requires_grad)
print(list(global_model.parameters()))
# -
Flex(MLP)(Flex.d(), 16, 10)(torch.randn(10, 5))
# +
global_attr = torch.randn(10, 3)
edge_attr = torch.randn(20, 3)
edges = torch.randint(0, 40, torch.Size([2, 20]))
node_attr = torch.randn(40, 2)
node_idx = torch.randint(0, 3, torch.Size([40]))
edge_idx = torch.randint(0, 3, torch.Size([20]))
global_model = AggregatingGlobalBlock(Flex(MLP)(Flex.d(), 16, 10), Aggregator('mean'), Aggregator('mean'))
global_model(global_attr, node_attr, edge_attr, edges, node_idx, edge_idx)
for p in global_model.parameters():
print(p)
print(p.requires_grad)
# +
x = torch.zeros(10)
# x.to("cuda")
# -
x.contiguous()
# ## Test False Training with Blocks
# +
def edge_forward(model, data):
return model(data.e, data.x, data.edges)
def node_forward(model, data):
return model()
# +
# AggregatingNodeBlock.forward?
# +
from os.path import isdir
from torch.utils.tensorboard import SummaryWriter
def new_writer(directory: str, suffix = ''):
i = 0
def name(index):
return directory + "%04d" % index + suffix
while isdir(name(i)):
i += 1
dirname = name(i)
print("New writer at '{}'".format(dirname))
return SummaryWriter(dirname)
# -
# ### Testining on GlobalBlock
# +
from tqdm.auto import tqdm
# new writer
writer = new_writer('runs/global_block')
# initialize random data
test_data = [random_data(5, 4, 3) for _ in range(1000)]
test_loader = GraphDataLoader(test_data, batch_size=32, shuffle=True)
test_mask = (
torch.tensor([True, True, True, True, False]),
torch.tensor([True, True, False, False]),
torch.tensor([True, True, False])
)
# model
# model = GlobalBlock(MLP(2, 16, 1))
model = GlobalBlock(Flex(MLP)(Flex.d(), 16, 1))
model = AggregatingGlobalBlock(Flex(MLP)(Flex.d(), 16, 1), Aggregator('add'), Aggregator('add'))
# global_attr, node_attr, edge_attr, edges, node_idx, edge_idx):
for batch in test_loader:
batch = batch.mask(*test_mask)
model(batch.g, batch.x, batch.e, batch.edges, batch.node_idx, batch.edge_idx)
# model = torch.nn.Sequential(
# torch.nn.Linear(2, 16),
# torch.nn.ReLU(),
# torch.nn.Linear(16, 1),
# torch.nn.ReLU()
# )
# setup initializer
optimizer = torch.optim.Adam(model.parameters(), lr=0.001, weight_decay=5e-4)
loss_fn = torch.nn.MSELoss()
#training
for epoch in tqdm(range(100)):
running_loss = 0.
for batch_idx, batch in enumerate(test_loader):
test_data = batch.mask(*test_mask)
target_data = batch.mask(*test_mask, invert=True)
out = model(test_data.g, test_data.x, test_data.e, test_data.edges, test_data.node_idx, test_data.edge_idx)
loss = loss_fn(out, target_data.g)
loss.backward()
optimizer.step()
running_loss += loss.item()
writer.add_scalar('test loss', running_loss, epoch)
if epoch % 1 == 0:
k = 'block_dict.mlp.resolved_module.blocks.0.blocks.0.weight'
writer.add_histogram(k, model.state_dict()[k])
# -
model.state_dict()
# ### Test GraphEncoder
class GraphEncoder(torch.nn.Module):
def __init__(self, edge_block, node_block, global_block):
assert issubclass(type(edge_block), EdgeBlock)
assert issubclass(type(node_block), NodeBlock)
assert issubclass(type(global_block), GlobalBlock)
super().__init__()
self.node_block = node_block
self.edge_block = edge_block
self.global_block = global_block
def forward(self, data: GraphBatch):
a = self.node_block.forward_from_data(data)
b = self.edge_block.forward_from_data(data)
c = self.global_block.forward_from_data(data)
return a, b, c
class GraphCore(torch.nn.Module):
def __init__(self, edge_block, node_block, global_block):
assert issubclass(type(edge_block), AggregatingEdgeBlock)
assert issubclass(type(node_block), AggregatingNodeBlock)
assert issubclass(type(global_block), AggregatingGlobalBlock)
super().__init__()
self.node_block = node_block
self.edge_block = edge_block
self.global_block = global_block
def forward(self, data: GraphBatch):
edge_attr = self.edge_block(data.e, data.x, data.edges)
node_attr = self.node_block(data.x, edge_attr, data.edges)
global_attr = self.global_block(data.g, data.x, data.e, data.edges, data.node_idx, data.edge_idx)
return edge_attr, node_attr, global_attr
# +
# initialize random data
test_data = [random_data(5, 4, 3) for _ in range(1000)]
test_loader = GraphDataLoader(test_data, batch_size=32, shuffle=True)
test_mask = (
torch.tensor([True, True, True, True, False]),
torch.tensor([True, True, False, False]),
torch.tensor([True, True, False])
)
# define the model
model = GraphCore(
AggregatingEdgeBlock(Flex(MLP)(Flex.d(), 16, 2)),
AggregatingNodeBlock(Flex(MLP)(Flex.d(), 16, 1), Aggregator('add')),
AggregatingGlobalBlock(Flex(MLP)(Flex.d(), 8, 1), Aggregator('add'), Aggregator('add'))
)
# prime the model
for batch in test_loader:
pass
model(batch.mask(*test_mask))
# setup initializer
optimizer = torch.optim.Adam(model.parameters(), lr=0.001, weight_decay=5e-4)
loss_fn = torch.nn.MSELoss()
writer = new_writer('runs/full_gcn_block')
#training
for epoch in tqdm(range(100)):
running_loss = 0.
for batch_idx, batch in enumerate(test_loader):
test_data = batch.mask(*test_mask)
target_data = batch.mask(*test_mask, invert=True)
edge_attr, node_attr, global_attr = model(test_data)
loss1 = loss_fn(edge_attr, target_data.e)
loss2 = loss_fn(node_attr, target_data.x)
loss3 = loss_fn(global_attr, target_data.g)
loss = loss1 + loss2 + loss3
loss.backward()
optimizer.step()
running_loss += loss.item()
writer.add_scalar('test loss', running_loss, epoch)
if epoch % 1 == 0:
k = list(model.state_dict())[0]
writer.add_histogram(k, model.state_dict()[k])
# -
list(model.state_dict())[0]
# # CircuitSeq
# +
import numpy as np
import seaborn as sns
import pylab as plt
# %matplotlib inline
# +
n_parts = 5
ka = np.zeros(n_parts)
ka[0] = 1
def sigmoid(x, a, kd, n, offset):
return a - (a) / (1. + np.exp((-x + kd)*n)) + offset
x = np.linspace(0, 20, 30)
y = sigmoid(x, 10, 10, 0.2, 0)
sns.scatterplot(x=x, y=y)
x = np.linspace(0, 20, 30)
y = sigmoid(x, 10, 10, 1, 10)
sns.scatterplot(x=x, y=y)
# +
num = 20
A = np.random.uniform(5, 20, size=(num))
K = np.random.uniform(5, 20, size=(num))
n = np.random.uniform(0.1, 1, size=(num))
o = np.random.uniform(0, A.max() / 10., size=(num))
A = np.expand_dims(A, 1)
K = np.expand_dims(K, 1)
n = np.expand_dims(n, 1)
o = np.expand_dims(o, 1)
# [n_parts, n_params]
params = np.hstack([A, K, n, o])
print(params.T.shape)
x = np.repeat(np.expand_dims(np.linspace(0, 40, 30), 0), num, axis=0)
y = sigmoid(x, *tuple(np.expand_dims(params.T, 2)))
plt.plot(x.T, y.T);
# +
import networkx as nx
g = nx.DiGraph()
g.add_edge(0, 1)
g.add_edge(1, 2)
g.add_edge(2, 3)
x = np.linspace(0, 30, 30)
def sigmoid_partial(x, node):
return sigmoid(x, *tuple(np.expand_dims(params[node:node+1].T, 2)))
def steady_state(g, acc='sum', node_to_part = lambda x: int(x)):
acc_dict = {
'sum': lambda x: np.sum(np.concatenate(x))
}
# in topological order, we evaluate the sigmoid function at each node
for node in nx.topological_sort(g):
idx = node_to_part(node)
# gather all parents
# accumulate outputs 'y' using the provided accumulation function
parents = list(g.predecessors(node))
if not parents:
p = np.expand_dims(params[idx:idx+1].T, 2)
x = np.array([[0.]])
else:
a = []
for p in parents:
_x = g.nodes[p]['y']
a.append(_x)
x = acc_dict[acc](a)
x = x.reshape(_x.shape)
y = sigmoid_partial(x, idx)
# if 'y' in g.nodes[node]:
# g.nodes[node]['y'] += y
# else:
g.nodes[node]['y'] = y
for _ in range(5):
steady_state(g, node_to_part = lambda x: int(x))
out = []
for n, ndata in g.nodes(data=True):
out.append(ndata['y'])
print(np.array(out).flatten())
# +
from itertools import permutations
from typing import *
from functools import partial
def has_cycle(g):
try:
nx.find_cycle(g)
return True
except nx.NetworkXNoCycle:
return False
def random_circuit(part_range):
g = nx.DiGraph()
n = np.random.randint(1, 10)
nodes = np.random.randint(0, 20, size=(n, 3))
for n1, n2 in permutations(nodes, r=2):
if n1[-1] in (n2[0], n2[1]):
g.add_edge(tuple(n1), tuple(n2))
return g
def iter_random_circuit(limit, part_range: Tuple[int, int], cycles: bool = True):
new_circuit = partial(random_circuit, part_range=part_range)
for i in range(limit):
c = new_circuit()
if cycles is True:
yield c
else:
while has_cycle(c):
c = new_circuit()
yield c
# +
import torch
from tqdm.auto import tqdm
from torch_geometric.data import DataLoader
def add_features(g):
new_g = nx.DiGraph()
for n, data in g.nodes(data=True):
new_g.add_node(n, **data)
for n1, n2, edata in g.edges(data=True):
edata['features'] = np.array([0.])
print(edata)
new_g.add_edge(n1, n2, **edata)
steady_state(new_g, node_to_part= lambda x: x[-1])
for n, ndata in new_g.nodes(data=True):
ndata['features'] = torch.tensor(list(n), dtype=torch.float)
ndata['target'] = torch.tensor([ndata['y'].flatten()], dtype=torch.float)
return new_g
graphs = [add_features(g) for g in iter_random_circuit(3000, (2, 20), cycles=False)]
# -
for _, _, edata in graphs[0].edges(data=True):
print(edata)
to_graph_data(graphs[0])
| ipython/PytorchGraphNetsRewrite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This script performs the equity calculation that is generally to be used in the figures and results of the paper. It should then be followed by script '03_...' starting not from the beginning but skipping the first few cells.
# +
# %matplotlib inline
import numpy as np
import os
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
from scipy import integrate
import seaborn as sns
import itertools
from matplotlib.lines import Line2D
import matplotlib.pylab as plb
from sklearn.linear_model import LinearRegression
import matplotlib.style as style
from matplotlib.patches import Patch
data_path = ""
plot_path = ""
table_path = ""
pd.set_option('display.max_columns', 500)
# +
main_remind=pd.read_excel(data_path+'REMIND_prepINPUT_use.xls')
main_image=pd.read_excel(data_path+'IMAGE_prepINPUT_use.xls')
main_message=pd.read_excel(data_path+'MESSAGE_prepINPUT_use.xls')
# +
main_remind = main_remind.drop(columns='Unnamed: 0')
main_image = main_image.drop(columns='Unnamed: 0')
main_message = main_message.drop(columns='Unnamed: 0')
main_remind.columns
# +
# Sum regions up to match REMIND regions
main_reg_image = main_image.copy()
main_reg_message = main_message.copy()
sumColums = ['Total_CO2', 'Total_GHG', 'BECCS', 'LANDCDR',
'TOTAL_CDR', 'CO2_Emissions', 'CUM_CO2', 'CUM_CDR',
'Population', 'GDP', 'cumPopulation']
mask_reg = (main_reg_image.Region.isin(['EAF', 'RSAF', 'SAF', 'WAF']))
for sce in main_reg_image.Scenario.unique():
mask_sce = (main_reg_image.Scenario == sce)
for year in main_reg_image.Year.unique():
mask_year = (main_reg_image.Year == year)
filtered_df = main_reg_image.loc[mask_sce & mask_reg & mask_year,:]
result = main_reg_image.loc[mask_sce & mask_reg & mask_year,sumColums].sum(axis=0)
idx = len(main_reg_image)+1
main_reg_image.loc[idx,:] = filtered_df.iloc[1]
main_reg_image.loc[idx,sumColums] = result
main_reg_image.loc[idx,'Region'] = 'AFR'
mask_reg = (main_reg_image.Region.isin(['NAF', 'ME']))
for sce in main_reg_image.Scenario.unique():
mask_sce = (main_reg_image.Scenario == sce)
for year in main_reg_image.Year.unique():
mask_year = (main_reg_image.Year == year)
filtered_df = main_reg_image.loc[mask_sce & mask_reg & mask_year,:]
result = main_reg_image.loc[mask_sce & mask_reg & mask_year,sumColums].sum(axis=0)
idx = len(main_reg_image)+1
main_reg_image.loc[idx,:] = filtered_df.iloc[0]
main_reg_image.loc[idx,sumColums] = result
main_reg_image.loc[idx,'Region'] = 'MEA'
mask_reg = (main_reg_image.Region.isin(['RSAM', 'RCAM', 'MEX', 'BRA']))
for sce in main_reg_image.Scenario.unique():
mask_sce = (main_reg_image.Scenario == sce)
for year in main_reg_image.Year.unique():
mask_year = (main_reg_image.Year == year)
filtered_df = main_reg_image.loc[mask_sce & mask_reg & mask_year,:]
result = main_reg_image.loc[mask_sce & mask_reg & mask_year,sumColums].sum(axis=0)
idx = len(main_reg_image)+1
main_reg_image.loc[idx,:] = filtered_df.iloc[0]
main_reg_image.loc[idx,sumColums] = result
main_reg_image.loc[idx,'Region'] = 'LAM'
mask_reg = (main_reg_image.Region.isin(['CEU', 'WEU']))
for sce in main_reg_image.Scenario.unique():
mask_sce = (main_reg_image.Scenario == sce)
for year in main_reg_image.Year.unique():
mask_year = (main_reg_image.Year == year)
filtered_df = main_reg_image.loc[mask_sce & mask_reg & mask_year,:]
result = main_reg_image.loc[mask_sce & mask_reg & mask_year,sumColums].sum(axis=0)
idx = len(main_reg_image)+1
main_reg_image.loc[idx,:] = filtered_df.iloc[0]
main_reg_image.loc[idx,sumColums] = result
main_reg_image.loc[idx,'Region'] = 'EUR'
# Make Region-Column universal
#Image: INDIA -> IND
# JAP -> JPN
#Message: R11_AFR -> AFR
# EU -> EUR
# R11_LAM -> LAM
# R11_MEA -> MEA
# R11_FSU -> RUS
main_reg_image = main_reg_image.replace('INDIA', 'IND')
main_reg_image = main_reg_image.replace('JAP', 'JPN')
main_reg_message = main_reg_message.replace('R11_AFR', 'AFR')
main_reg_message = main_reg_message.replace('EU', 'EUR')
main_reg_message = main_reg_message.replace('R11_LAM', 'LAM')
main_reg_message = main_reg_message.replace('R11_MEA', 'MEA')
main_reg_message = main_reg_message.replace('R11_FSU', 'RUS')
main_reg_message = main_reg_message.replace('R11_PAO', 'PAO')
main_reg_message = main_reg_message.replace('R11_PAS', 'PAS')
# +
# Delete rows in dataframe that are not the same as remind model
main_equal_image = main_reg_image.copy()
indexNames = main_reg_image[(main_reg_image['Region'] == 'CAN') + (main_reg_image['Region'] == 'INDO')
+ (main_reg_image['Region'] == 'JPN') + (main_reg_image['Region'] == 'KOR') +
(main_reg_image['Region'] == 'ME') + (main_reg_image['Region'] == 'OCE') +
(main_reg_image['Region'] == 'RSAS') + (main_reg_image['Region'] == 'SEAS') +
(main_reg_image['Region'] == 'STAN') + (main_reg_image['Region'] == 'TUR') +
(main_reg_image['Region'] == 'BRA') + (main_reg_image['Region'] == 'UKR') +
(main_reg_image['Region'] == 'CEU') + (main_reg_image['Region'] == 'EAF') +
(main_reg_image['Region'] == 'MEX') + (main_reg_image['Region'] == 'NAF') +
(main_reg_image['Region'] == 'RCAM')+ (main_reg_image['Region'] == 'RSAF') +
(main_reg_image['Region'] == 'RSAM')+ (main_reg_image['Region'] == 'SAF') +
(main_reg_image['Region'] == 'WAF') + (main_reg_image['Region'] == 'WEU')].index
main_equal_image.drop(indexNames, inplace=True)
main_equal_message = main_reg_message.copy()
indexNames = main_reg_message[
(main_reg_message['Region'] == 'PAO') + (main_reg_message['Region'] == 'PAS')].index
#(main_reg_message['Region'] == 'MEA') +
main_equal_message.drop(indexNames, inplace=True)
main_equal_remind = main_remind.copy()
indexNames = main_remind[(main_remind['Region'] == 'JPN')
+ (main_remind['Region'] == 'OAS') + (main_remind['Region'] == 'ROW')].index
# #+ (main_remind['Region'] == 'MEA')
main_equal_remind.drop(indexNames, inplace=True)
allmodels_equal_df = main_equal_remind.append(main_equal_image)
allmodels_equal_df = allmodels_equal_df.append(main_equal_message)
indexNames = allmodels_equal_df[(allmodels_equal_df['Region'] == 'PAO')
+ (allmodels_equal_df['Region'] == 'PAS')+ (allmodels_equal_df['Region'] == 'MEA')].index
# #+ (main_remind['Region'] == 'MEA')
allmodels_equal_df.drop(indexNames, inplace=True)
print(allmodels_equal_df.Region.unique())
# -
print(main_equal_remind.Region.unique())
print(main_equal_image.Region.unique())
print(main_equal_message.Region.unique())
# +
regions = ['AFR', 'CHN', 'EUR', 'IND', 'LAM', 'RUS', 'USA', 'World']
regions_no_world = ['AFR', 'CHN', 'EUR', 'IND', 'LAM', 'RUS', 'USA']
def GDP_cap_calc(main_df, regions_names):
main_df['GDP'] = main_df['GDP']#*1000
main_df['Unit_GDP'] = 'USD Billion'
main_df['GDP_cap'] = 0
main_df['GDP_cap_avg'] = 0
wfilt = (main_df.Region == 'World')
for scen in main_df.Scenario.unique():
rfilt=(main_df.Region.isin(regions_names))
sfilt=(main_df.Scenario==scen)
for yr in main_df.Year.unique():
yrfilt=(main_df.Year==yr)
main_df.loc[sfilt&yrfilt&rfilt,
'GDP_cap'] = main_df.loc[
rfilt&yrfilt&sfilt,'GDP'].values/main_df.loc[rfilt&yrfilt&sfilt,'Population'].values
main_df.loc[sfilt&yrfilt&rfilt,
'GDP_cap_avg'] = sum(main_df.loc[wfilt&sfilt&yrfilt, 'GDP_cap'])
return main_df
main_image = GDP_cap_calc(main_equal_image, regions)
main_remind = GDP_cap_calc(main_equal_remind, regions)
main_message = GDP_cap_calc(main_equal_message, regions)
# -
main_image.Scenario.unique()
# +
def CoI_calc(main_df):
mask = (main_df['GDP_cap'].values > main_df['GDP_cap_avg'].values)
main_df['GDP_cap_CoI_nan'] = np.where(mask, main_df['GDP_cap'], np.nan)
main_df['GDP_cap_CoI'] = np.where(mask, main_df['GDP_cap'], 0)
main_df['GDP_CoI_nan'] = np.where(mask, main_df['GDP'], np.nan)
main_df['GDP_CoI'] = np.where(mask, main_df['GDP'], 0)
return main_df
main_image = CoI_calc(main_image)
main_remind = CoI_calc(main_remind)
main_message = CoI_calc(main_message)
# +
# Alternative GDP EQUITY CALCULATION
regions = ['AFR', 'CHN', 'EUR', 'IND', 'LAM', 'RUS', 'USA', 'World']
regions_no_world = ['AFR', 'CHN', 'EUR', 'IND', 'LAM', 'RUS', 'USA']
def GDP_eq_Winkler(main_df, regions_names):
main_df['zahler'] = 0
main_df['nenner'] = 0
for scen in main_df.Scenario.unique():
#rfilt=(main_df.Region.isin(regions_names))
wfilt = (main_df.Region == 'World')
sfilt=(main_df.Scenario==scen)
for yr in main_df.Year.unique():
yrfilt= (main_df.Year == yr)
rfilt=(main_df.Region.isin(regions_names))
main_df.loc[sfilt&yrfilt&rfilt,
'nenner'] = sum((main_df.loc[
rfilt&yrfilt&sfilt,'Population'].values/main_df.loc[wfilt&yrfilt&sfilt,'Population'].values) * main_df.loc[rfilt&yrfilt&sfilt,'GDP_cap'].values)
return main_df
main_image = GDP_eq_Winkler(main_image, regions_no_world)
main_remind = GDP_eq_Winkler(main_remind, regions_no_world)
main_message = GDP_eq_Winkler(main_message, regions_no_world)
# +
def GDP_eq_zahler(main_df, regions_names):
for scen in main_df.Scenario.unique():
#rfilt=(main_df.Region.isin(regions_names))
wfilt = (main_df.Region == 'World')
sfilt=(main_df.Scenario==scen)
for yr in main_df.Year.unique():
yrfilt= (main_df.Year == yr)
rfilt=(main_df.Region.isin(regions_names))
main_df['Tot_CDR'] = 0
# main_df.loc[sfilt & yrfilt & rfilt, 'Tot_CDR'] = main_df.loc[sfilt & yrfilt & wfilt, 'TOTAL_CDR']
# print(main_df.loc[sfilt & yrfilt & rfilt, 'Tot_CDR'])
# print(main_df.loc[sfilt & yrfilt & wfilt, 'TOTAL_CDR'])
for reg in main_df.Region.unique():
reg_filt = (main_df.Region == reg)
# main_df.loc[sfilt & yrfilt & reg_filt, 'Tot_CDR'] = main_df.loc[sfilt & yrfilt & wfilt, 'TOTAL_CDR'].values
main_df.loc[sfilt&yrfilt®_filt,'zahler'] = (main_df.loc[sfilt&yrfilt®_filt,
'Population'].values/main_df.loc[wfilt&yrfilt&sfilt,'Population'].values) * main_df.loc[reg_filt&yrfilt&sfilt,'GDP_cap'].values
main_df.loc[sfilt&yrfilt®_filt,'M'] = (main_df.loc[sfilt&yrfilt®_filt,
'zahler'] / main_df.loc[sfilt&yrfilt®_filt,
'nenner']) * main_df.loc[sfilt&yrfilt&wfilt,
'TOTAL_CDR'].values
return main_df
main_image = GDP_eq_zahler(main_image, regions_no_world)
main_remind = GDP_eq_zahler(main_remind, regions_no_world)
main_message = GDP_eq_zahler(main_message, regions_no_world)
# -
main_image.loc[main_image.Scenario == 'SSP1-19-SPA0-V17']
main_image.to_excel(data_path+'IMAGE_equity_GDP_WINKLER.xls')
print(main_image.Region.unique())
print(main_remind.Region.unique())
print(main_message.Region.unique())
# +
def GDP_CoI_frac_calc(main_df, regions_names_noworld):
main_df['GDP_CoI_frac'] = 0
main_df['CDR_per_cap_GDP'] = 0
main_df['CDR_per_cap_per_GDP_per_cap'] = 0
for scen in main_df.Scenario.unique():
rfilt=(main_df.Region.isin(regions_names_noworld))
sfilt=(main_df.Scenario==scen)
for yr in main_df.Year.unique():
yrfilt=(main_df.Year==yr)
tot_GDP_yr = main_df.loc[sfilt&yrfilt&rfilt,
'GDP_CoI'].sum()
main_df.loc[sfilt&yrfilt&rfilt,
'GDP_CoI_frac'] = (main_df.loc[rfilt&sfilt&yrfilt,'GDP_CoI'])/tot_GDP_yr
main_df.loc[sfilt&yrfilt&rfilt,
'CDR_fair_share_by_GDP'] = (main_df.loc[rfilt&sfilt&yrfilt,'GDP_CoI_frac'])*(main_df.loc[sfilt&yrfilt&(main_df.Region=='World'),'TOTAL_CDR'].values)
main_df.loc[sfilt&yrfilt&rfilt,
'CDR_per_cap_GDP'] = (main_df.loc[rfilt&sfilt&yrfilt,'CDR_fair_share_by_GDP'])/(main_df.loc[sfilt&yrfilt&rfilt,'Population'].values)
main_df.loc[sfilt&yrfilt&rfilt,
'CDR_per_cap_per_GDP_per_cap'] = (main_df.loc[rfilt&sfilt&yrfilt,'CDR_per_cap_GDP'])/(main_df.loc[sfilt&yrfilt&rfilt,'GDP_cap'].values)
main_df['CUM_CDR_fair_share_by_GDP'] = 0
main_df['Cum_M'] = 0
for scen in main_df.Scenario.unique():
for reg in main_df.Region.unique():
main_df.loc[(main_df.Year == 2020), 'CUM_CDR_fair_share_by_GDP'] = main_df.loc[(main_df.Year == 2020), 'CDR_fair_share_by_GDP']
main_df.loc[(main_df.Year == 2020), 'Cum_M'] = main_df.loc[(main_df.Year == 2020), 'M']
for reg in main_df.Region.unique():
if reg in regions_names_noworld:
mask = main_df.Region == reg
full_reg = main_df.loc[mask,:]
for scen in full_reg.Scenario.unique():
mask_scen = full_reg.Scenario == scen
full_scen = full_reg.loc[mask_scen,:]
integral = integrate.cumtrapz(y = full_scen.CDR_fair_share_by_GDP, x = full_scen.Year, initial = 0)
full_reg.loc[mask_scen,'CUM_CDR_fair_share_by_GDP'] += integral
inte_M = integrate.cumtrapz(y = full_scen.M, x = full_scen.Year, initial = 0)
full_reg.loc[mask_scen,'Cum_M'] += inte_M
main_df.loc[mask,'CUM_CDR_fair_share_by_GDP'] = full_reg.loc[:,'CUM_CDR_fair_share_by_GDP']
main_df.loc[mask,'Cum_M'] = full_reg.loc[:,'Cum_M']
return main_df
main_remind = GDP_CoI_frac_calc(main_remind, regions_no_world)
main_image = GDP_CoI_frac_calc(main_image, regions_no_world)
main_message = GDP_CoI_frac_calc(main_message, regions_no_world)
# -
main_remind
# +
"""%% EQUITY CALCULATION - CUMULATIVE EMISSIONS PER CUMULATIVE CAPITA
At each point in time calculate the cumulative emissions divided by the cumulative population for the world,
...then multiply this value by the population of each country to find the fair share of emissions
...then find the difference between fair share emissions and actual emissions - this is the required negative emissions
"""
def equity_calc(main_df):
# DERIVE WORLD FAIR SHARE BOTH GDP AND POPULATION
main_df['Global_CUM_CO2_by_CUM_POP']=np.nan
#main_df['Global_CUM_CO2_by_GDP']=np.nan
#main_df['Fair_share_by_GDP']=np.nan
main_df['Fair_share_by_CUM_POP']=np.nan
#main_df['Excess_emissions_by_GDP']=np.nan
main_df['Excess_emissions_by_CUM_POP']=np.nan
main_df['Exc_em_fac_by_CUM_POP']=np.nan
#main_df['Exc_em_fac_by_GDP']=np.nan
# Set to zero instead of NAN, for regions who have 'negative' CDR reliabilities
main_df['CDR_fair_share_by_CUM_POP']=0
#main_df['CDR_fair_share_by_GDP']=0
main_df['CUM_CDR_fair_share_by_CUM_POP']=0
#main_df['CUM_CDR_fair_share_by_GDP']=0
main_df['CDR_model_vs_fair_share_by_CUM_POP']=0
main_df['CDR_model_vs_fair_share_by_GDP']=0
main_df['CUM_CDR_model_vs_fair_share_by_CUM_POP']=0
main_df['CUM_CDR_model_vs_fair_share_by_GDP']=0
# Derive World Average
reg='World'
for scen in main_df.Scenario.unique():
rfilt=(main_df.Region==reg)
sfilt=(main_df.Scenario==scen)
for yr in main_df.Year.unique():
yrfilt=(main_df.Year==yr)
main_df.loc[sfilt&yrfilt,
'Global_CUM_CO2_by_CUM_POP']=main_df.loc[
rfilt&yrfilt&sfilt,'CUM_CO2'].values/main_df.loc[rfilt&yrfilt&sfilt,'cumPopulation'].values #cumPop & CUM_CO2of world
# main_df.loc[sfilt&yrfilt,
# 'Global_CUM_CO2_by_GDP']=main_df.loc[
# rfilt&yrfilt&sfilt,'CUM_CO2'].values/main_df.loc[rfilt&yrfilt&sfilt,'GDP'].values
# Derive country fair share population based
main_df['Fair_share_by_CUM_POP']=main_df['Global_CUM_CO2_by_CUM_POP']*main_df['cumPopulation'] #cumPop of single Region?
# main_df['Fair_share_by_GDP']=main_df['Global_CUM_CO2_by_GDP']*main_df['GDP']
# Derive difference between fair share and actual emissions
main_df['Excess_emissions_by_CUM_POP']=main_df['CUM_CO2']-main_df['Fair_share_by_CUM_POP']
# main_df['Excess_emissions_by_GDP']=main_df['CUM_CO2']-main_df['Fair_share_by_GDP']
rfilt_wo_world=(main_df.Region!='World')
# Derive total burden for countries
for yr in main_df.Year.unique():
yrfilt=(main_df.Year==yr)
for scen in main_df.Scenario.unique():
scen_filt=(main_df.Scenario==scen)
# Filter only for regions with positive excess emissions
exemfilt_POP=(main_df.Excess_emissions_by_CUM_POP>0)&rfilt_wo_world&yrfilt&scen_filt
# exemfilt_GDP=(main_df.Excess_emissions_by_GDP>0)&rfilt_wo_world&yrfilt&scen_filt
# Derive total excess emissions per year
tot_exc=main_df.loc[exemfilt_POP,'Excess_emissions_by_CUM_POP'].sum()
# Derive_country_share
main_df.loc[exemfilt_POP,
'Exc_em_fac_by_CUM_POP']=main_df.loc[exemfilt_POP,'Excess_emissions_by_CUM_POP']/tot_exc
main_df.loc[exemfilt_POP,
'CDR_fair_share_by_CUM_POP']=main_df.loc[exemfilt_POP,
'Exc_em_fac_by_CUM_POP']*main_df.loc[yrfilt&scen_filt&
(main_df.Region=='World'),'TOTAL_CDR'].values
main_df.loc[exemfilt_POP,
'CUM_CDR_fair_share_by_CUM_POP']=main_df.loc[exemfilt_POP,
'Exc_em_fac_by_CUM_POP']*main_df.loc[yrfilt&scen_filt&
(main_df.Region=='World'),'CUM_CDR'].values
#GDP CALCULATION
# Derive total excess emissions per year
# glob_exc_gdp=main_df.loc[exemfilt_GDP,'Excess_emissions_by_GDP']
# Sum only over positive contributions
# tot_exc_gdp=glob_exc_gdp[glob_exc_gdp>0].sum()
# Derive_country_share
# main_df.loc[exemfilt_GDP,
# 'Exc_em_fac_by_GDP']=main_df.loc[exemfilt_GDP,'Excess_emissions_by_GDP']/tot_exc_gdp
# main_df.loc[exemfilt_GDP,
# 'CDR_fair_share_by_GDP']=main_df.loc[exemfilt_GDP,
# 'Exc_em_fac_by_GDP']*main_df.loc[yrfilt&scen_filt&
# (main_df.Region=='World'),'TOTAL_CDR'].values
# main_df.loc[exemfilt_GDP,
# 'CUM_CDR_fair_share_by_GDP']=main_df.loc[exemfilt_GDP,
# 'Exc_em_fac_by_GDP']*main_df.loc[yrfilt&scen_filt&
# (main_df.Region=='World'),'CUM_CDR'].values
# Derive difference between model and equity CDR distribution
main_df['CDR_model_vs_fair_share_by_CUM_POP']= main_df['CDR_fair_share_by_CUM_POP']-main_df['TOTAL_CDR']
main_df['CDR_model_vs_fair_share_by_GDP']= main_df['CDR_fair_share_by_GDP']-main_df['TOTAL_CDR']
main_df['CUM_CDR_model_vs_fair_share_by_CUM_POP']= main_df['CUM_CDR_fair_share_by_CUM_POP']-main_df['CUM_CDR']
main_df['CUM_CDR_model_vs_fair_share_by_GDP']= main_df['CUM_CDR_fair_share_by_GDP']-main_df['CUM_CDR']
return main_df
equity_remind = equity_calc(main_remind)
equity_image = equity_calc(main_image)
equity_message = equity_calc(main_message)
# +
#this function basically sums up the 2100 cumulative difference between CUM_CDR according to model
# and fair share CUM_POP according to equity calculation (if difference is POSITIVE)
def total_excess_CDR_by_CUM_POP(equity_df, scenario):
scen = scenario
rfilt_wo_world=(equity_df.Region!='World')
scen_filt = (equity_df.Scenario == scen)
for yr in equity_df.Year.unique():
yrfilt=(equity_df.Year==yr)
excdrfilt_POP=(equity_df.CUM_CDR_model_vs_fair_share_by_CUM_POP>0)&rfilt_wo_world&yrfilt&scen_filt
tot_exs_CDR_by_CUM_POP = equity_df.loc[excdrfilt_POP,'CUM_CDR_model_vs_fair_share_by_CUM_POP'].sum()
return(tot_exs_CDR_by_CUM_POP)
def total_excess_CDR_by_CUM_GDP(equity_df, scenario):
scen = scenario
rfilt_wo_world=(equity_df.Region!='World')
scen_filt = (equity_df.Scenario == scen)
for yr in equity_df.Year.unique():
yrfilt=(equity_df.Year==yr)
excdrfilt_GDP=(equity_df.CUM_CDR_model_vs_fair_share_by_GDP>0)&rfilt_wo_world&yrfilt&scen_filt
tot_exs_CDR_by_CUM_GDP = equity_df.loc[excdrfilt_GDP,'CUM_CDR_model_vs_fair_share_by_GDP'].sum()
return(tot_exs_CDR_by_CUM_GDP)
# +
equity_image = equity_image.round(2)
equity_remind = equity_remind.round(2)
equity_message = equity_message.round(2)
equity_image.to_excel(data_path+'IMAGE_equity_equalreg_use.xls')
equity_remind.to_excel(data_path+'REMIND_equity_equalreg_use.xls')
equity_message.to_excel(data_path+'MESSAGE_equity_equalreg_use.xls')
# +
# Master dataframe with all models
allmodels_df = pd.concat([equity_image, equity_message, equity_remind], ignore_index=True, sort=False)
# Excel export
allmodels_df.to_excel(data_path + 'MASTERDF_equity_equalreg_use.xls')
print(allmodels_df.Region.unique())
# +
dat = allmodels_df.loc[(allmodels_df.Year == 2100) & (allmodels_df.MODEL == 'IMAGE')]
sns.barplot(x = 'Region', y = 'Cum_M', data = dat)
# -
dat = allmodels_df.loc[allmodels_df.Year == 2100]
sns.barplot(x = 'Region', y = 'CUM_CDR_fair_share_by_GDP', data = dat)
| 02_CDR_region_alloc_equity_calc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
from bs4 import BeautifulSoup
from urllib import request
import re
yrly_unemployment_data = request.urlopen('http://www.bls.gov/cps/cpsaat01.htm')
soup = BeautifulSoup(yrly_unemployment_data,'html')
yrs_initial = soup.find_all('th',id=re.compile('cps_eeann_year.r.'))
yrs_final = [int(yr.text) for yr in yrs_initial if yr.text.isnumeric()]
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
data = soup.find_all(class_="datavalue")
# Unemployment rate is the 8th value in the table data and it repeats every 10 rows
unemploy_rate_data = data[8::10]
unemploy_rate = [float(rate.text) for rate in unemploy_rate_data]
# We have a list of years and a list of unemployment rates, we need to make a dictionary out of them using zip
unemployment_dict = {year:rate for year, rate in zip(yrs_final, unemploy_rate)}
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
bar_width = 0.5
ax = subplot()
current_axis = plt.gca()
for xticklabels in current_axis.get_xticklabels():
xticklabels.set_fontsize(8)
bar(np.arange(0,25), unemploy_rate[-25:], bar_width, alpha=0.7)
title("U.S. Unemployment Rate of last 25 Years")
ylabel("Unemployment Rate %")
xticks(np.arange(0,25),yrs_final[-25:],rotation=-90)
yticks(np.arange(0,11))
grid()
show()
| web_scraping/BeautifulSoup_Unemployment_Rate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # (Robust) One Hot Encoding
# One-hot encoding is a common technique used to work with categorical features. There are multiple tools available to facilitate this preprocessing step in `Python`, but it usually becomes a much harder when you need your preprocessing code to work on new data that might have missing or additional values. That's the case if you want to deploy a model to production for instance, sometimes you don't know what new values will appear in the data you receive.
#
# In this tutorial I will present two ways of dealing with this problem. Everytime we will run one hot encoding on our training set first and save a few attributes that we can reuse later on when we need to process new data. If you deploy a model to production, the best way of saving those values is writing your own class and define them as attributes that will be set at training time, as an internal state.
#
# If you're working in a notebook, it's fine to save them as simple variables.
# # Let's create a dataset
# Let's make up a dataset containing journeys that happened in different cities in the UK, using different ways of transportation/
#
# We'll create a new `DataFrame` that contains two categorical features, `city` and `transport` as well as a numerical feature `duration` for the duration of the journey in minutes.
import pandas as pd
df = pd.DataFrame([["London", "car", 20],
["Cambridge", "car", 10],
["Liverpool", "bus", 30]],
columns=["city", "transport", "duration"])
df
# Now let's create our "unseen" test data. To make it difficult, we will simulate the case where the test data has different values for the categorical features.
df_test = pd.DataFrame([["Manchester", "bike", 30],
["Cambridge", "car", 40],
["Liverpool", "bike", 10]],
columns=["city", "transport", "duration"])
df_test
# Here our column `city` does not have the value `London` but has a new value `Cambridge`. Our column `transport` has no value `bus` but the new value `bike`. Let's see how we can build one hot encoded features for those datasets!
#
# We'll show two different methods, one using the `get_dummies` method from `pandas`, and the other with the `OneHotEncoder` class from `sklearn`.
# ## Using pandas' `get_dummies`
# ### Process our training data
# First we define the list of categorical features that we will want to process:
cat_columns = ["city", "transport"]
# We can really quickly build dummy features with pandas by calling the `get_dummies` function. Let's create a new `DataFrame` for our processed data:
df_processed = pd.get_dummies(df, prefix_sep="__", columns=cat_columns)
df_processed
# That's it for the training set part, now you have a `DataFrame` with one hot encoded features. We will need to save a few things into variables to make sure that we build the exact same columns on the test dataset.
# See how pandas created new columns with the following format: `<column__value>`. Let's create a list that looks for those new columns and store them in a new variable `cat_dummies`.
cat_dummies = [col for col in df_processed if "__" in col and col.split("__")[0] in cat_columns]
cat_dummies
# Let's also save the list of columns so we can enforce the order of columns later on
processed_columns = list(df_processed.columns[:])
processed_columns
# ### Process our unseen (test) data!
# Now let's see how to ensure our test data has the same columns, first let's call `get_dummies` on it:
df_test_processed = pd.get_dummies(df_test, prefix_sep="__", columns=cat_columns)
# Let's look at our new dataset
df_test_processed
# As expected we have new columns (`city__Manchester`) and missing ones (`transport__bus`). But we can easily clean it up!
# Remove additional columns
for col in df_test_processed.columns:
if ("__" in col) and (col.split("__")[0] in cat_columns) and col not in cat_dummies:
print("Removing additional feature {}".format(col))
df_test_processed.drop(col, axis=1, inplace=True)
# Now we need to add the missing columns. We can set all missing columns to a vector of 0s since those values did not appear in the test data.
for col in cat_dummies:
if col not in df_test_processed.columns:
print("Adding missing feature {}".format(col))
df_test_processed[col] = 0
df_test_processed
# That's it, we now have the same features. Note that the order of the columns isn't kept though, if you need to reorder the columns, reuse the list of processed columns we saved earlier:
df_test_processed = df_test_processed[processed_columns]
df_test_processed
# All good! Now let's see how to do the same with sklearn and the `OneHotEncoder`
# ## With sklearn one hot and label encoder
# ### Process our training data
# Let's start by importing what we need. The `OneHotEncoder` to build one hot features, but also the `LabelEncoder` to transform strings into integer labels (needed before using the `OneHotEncoder`)
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
# We're starting again from our initial dataframe and our list of categorical features.
cat_columns = ["city", "transport"]
df
# First let's create our `df_processed` DataFrame, we can take all the non-categorical features to start with:
df_processed = df[[col for col in df.columns if col not in cat_columns]]
df_processed
# Now we need to encode every categorical feature separately, meaning we need as many encoders as categorical features. Let's loop over all categorical features and build a dictionary that will map a feature to its encoder:
# For each categorical column
# We fit a label encoder, transform our column and
# add it to our new dataframe
label_encoders = {}
for col in cat_columns:
print("Encoding {}".format(col))
new_le = LabelEncoder()
df_processed[col] = new_le.fit_transform(df[col])
label_encoders[col] = new_le
df_processed
# Now that we have proper integer labels, we need to one hot encode our categorical features.
# Unfortunately, the one hot encoder does not support passing the list of categorical features by their names but only by their indexes, so let's get a new list, now with indexes. We can use the `get_loc` method to get the index of each of our categorical columns:
cat_columns_idx = [df_processed.columns.get_loc(col) for col in cat_columns]
# We'll need to specify `handle_unknown` as `ignore` so the `OneHotEncoder` can work later on with our unseen data.
# The `OneHotEncoder` will build a numpy array for our data, replacing our original features by one hot encoding versions. Unfortunately it can be hard to re-build the DataFrame with nice labels, but most algorithms work with numpy arrays, so we can stop there.
ohe = OneHotEncoder(categorical_features=cat_columns_idx, sparse=False, handle_unknown="ignore")
df_processed_np = ohe.fit_transform(df_processed)
df_processed_np
# ### Process our unseen (test) data
# Now we need to apply the same steps on our test data; first create a new dataframe with our non-categorical features:
df_test_processed = df_test[[col for col in df_test.columns if col not in cat_columns]]
df_test_processed
# Now we need to reuse our `LabelEncoder`s to properly assign the same integer to the same values. Unfortunately since we have new, unseen, values in our test dataset, we cannot use transform. Instead we will create a new dictionary from the `classes_` defined in our label encoder. Those classes map a value to an integer. If we then use `map` on our pandas `Series`, it set the new values as `NaN` and convert the type to float.
#
# Here we will add a new step that fills the `NaN` by a huge integer, say 9999 and converts the column to `int`.
for col in cat_columns:
print("Encoding {}".format(col))
label_map = {val: label for label, val in enumerate(label_encoders[col].classes_)}
print(label_map)
df_test_processed[col] = df_test[col].map(label_map)
# fillna and convert to int
df_test_processed[col] = df_test_processed[col].fillna(9999).astype(int)
df_test_processed
# Looks good, now we can finally apply our fitted `OneHotEncoder` "out-of-the-box" by using the transform method:
df_test_processed_np = ohe.transform(df_test_processed)
df_test_processed_np
# Double check that it has the same columns as the `pandas` version!
| misc/one_hot_encoding/(Robust) One Hot Encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.1 64-bit (''ns'': conda)'
# language: python
# name: python38164bitnscondaf7cf2969956448d1a9f2563a2ea8251d
# ---
import pandas as pd
import networkx as nx
import numpy as np
import collections
import matplotlib.pyplot as plt
plt.xkcd()
# %matplotlib inline
path_file = "data/routes_data.tsv"
routes_df = pd.read_csv(path_file, sep="\t")
routes_df.head()
G = nx.nx.from_pandas_edgelist(
routes_df, "id_from", "id_to", ["delay", "station_from", "station_from"]
)
nx.draw(G)
nx.radius(G)
nx.diameter(G)
G.number_of_nodes()
G.number_of_edges()
G.degree()
# +
degree_sequence = sorted([d for n, d in G.degree()], reverse=True) # degree sequence
degreeCount = collections.Counter(degree_sequence)
deg, cnt = zip(*degreeCount.items())
fig, ax = plt.subplots()
plt.bar(deg, cnt, width=0.80, color='b')
plt.title("Degree Histogram")
plt.ylabel("Count")
plt.xlabel("Degree")
ax.set_xticks([d + 0.4 for d in deg])
ax.set_xticklabels(deg)
# draw graph in inset
plt.axes([0.4, 0.4, 0.5, 0.5])
Gcc = G.subgraph(sorted(nx.connected_components(G), key=len, reverse=True)[0])
pos = nx.spring_layout(G)
plt.axis('off')
nx.draw_networkx_nodes(G, pos, node_size=20)
nx.draw_networkx_edges(G, pos, alpha=0.4)
plt.show()
# -
plt.loglog(deg, cnt, '.', lw=2)
# From our experimentations we can realise that the Moscow Metrow is a skewed distribution with most nodes having 2 connections. Furthermore it may take multiple stops for one to get from point `A` to point `B` depending on where they are going. Hence passenger `X` may not be affected due to the number of stops they may share in common wih passenger `Y` but rather due to human interactions passenger `X` may have in common with passenger `Y`
| experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduzione a Pandas
#
# Oggetto di tipo `DataFrame` = tabella organizzata in righe (records) e colonne intestate.
#
# `Pandas` offre tre funzionalità principali:
#
# 1. costruzione
# 1. interrogazione
# 1. aggiornamento
# ### Leggere un file `csv` con Pandas
#
# Un file `csv` è un file composto di record di campi separati da `,` di cui il primo è il record di intestazione che specifica il nome di ognuno dei campi dei record seguenti che contengono i dati.
#
# Funzione `read_csv()` per leggere un file `csv`:
#
# df = pd.read_csv(csv_file_name)
#
# `df` è il riferimento a oggetto di tipo `DataFrame`
# #### Ad esempio leggiamo il file `2017-german-election-overall.csv`
# ### Ottenere informazioni sul data frame
#
# - informazioni generali sul data frame
#
# df.info()
#
# - statistiche generali sul data frame
#
# df.describe()
# ### Ottenere la copia di un data frame
#
# df.copy()
# ### Variabili `shape` e `columns`
#
# - `shape`, tupla contenente il numero di righe e numero di colonne del data frame
# - `columns`, oggetto `Index` che contiene i nomi delle colonne del data frame
# ### Cambiare i nomi delle colonne
#
# df.rename(columns = name_dict, inplace = True|False)
#
# `name_dict`, dizionario che mappa un nome a un nuovo nome
# ### Rimuovere colonne
#
# df.drop(column_list, axis = 1, inplace = True|False)
# ### Rimuovere righe per indice
#
# df.drop(index_list, axis = 0, inplace = True|False)
# ### Ottenere le prime/ultime righe
#
# df.head(n)
# df.tail(n)
# ### Selezionare righe per posizione (*slicing*)
#
# df[start_pos:end_pos]
# ### Selezionare una colonna
#
# L'espressione:
#
# df[column_name]
#
# restituisce la colonna con nome `column_name` in un oggetto `Series`
# In alternativa si può usare la notazione con il punto:
#
# df.column_name
# ### Selezionare colonne
#
# L'espressione:
#
# df[column_list]
#
# restituisce un data frame con le colonne specificate in `column_list`.
# ### Controllare se ci sono valori nulli
#
# Le espressioni:
#
# pd.isnull(df)
# df.isnull()
#
# restituiscono un data frame di valori booleani.
# Le espressioni:
#
# pd.isnull(series_obj)
# series_obj.isnull()
#
# restituiscono un data frame di valori booleani.
# ### Metodo `unique()`
#
# Il metodo `unique()` degli oggetti `Series` restituisce l'array dei valori distinti presenti nell'oggetto invocante.
# ### Selezionare le righe che verificano una certa condizione
#
# Le istruzioni equivalenti:
#
# mask = df[column_name] cfr_op value
# mask = df.column_name cfr_op value
#
# dove `cfr_op` è un operatore di confronto, assegnano alla variabile `mask` un oggetto `Series` di valori booleani in cui l'i-esimo booleano è `True` se il valore nell'i-esima riga in corrispondenza della colonna `column_name` verifica l'espressione di confronto.
# L'espressione:
#
# df[mask]
#
# restituisce un data frame con le sole righe che corrispondono a un valore `True` in `mask`.
# ### Ottenere gli indici delle righe che verificano una certa condizione
#
# df[mask].index
# ### Localizzare righe con `iloc[]`
#
# L'espressione:
#
# df.iloc[pos_index]
#
# restituisce in un oggetto di tipo `Series` la riga in posizione di indice `pos_index`.
# L'espressione:
#
# df.iloc[start_pos_index:end_pos_index]
#
# restituisce in un oggetto di tipo `DataFrame` tutte le righe dalla posizione di indice `start_pos_index` a quella di indice `end_pos_index-1`.
# L'espressione:
#
# df.iloc[pos_index_list]
#
# restituisce in un oggetto di tipo `DataFrame` tutte le righe specificate.
# ### Uso di `loc[]`
#
# - accesso a una riga tramite il suo indice
#
# df.loc[index]
# - accesso a più righe tramite i loro indici
#
# df.loc[[index1, index2, ...]]
# - accesso a un valore del data frame
#
# df.loc[index, column_name]
# - accesso a più valori del data frame
#
# df.loc[[index1, index2, ...], column_name]
# - accesso a più valori del data frame
#
# df.loc[[index1, index2, ...], [column_name1, column_name2, ...]]
# - accesso alle righe che verificano una certa condizione
#
# df.loc[mask]
# ### Ottenere un valore tramite un indice con `at[]`
#
# df.at[index, column_name]
# ### Ordinare valori
#
# Ordinare per valori di una colonna:
#
# df.sort_values(column_name, ascending = True|False, inplace = True|False)
#
# Ordinare per valori di più colonne:
#
# df.sort_values(column_list, ascending = True|False, inplace = True|False)
# ### Raggruppare i valori
#
# L'espressione:
#
# df.groupby(column_name)
# df.groupby(column_list)
#
# restituisce un oggetto `DataFrameGroupBy`.
# ### Aggiungere una colonna
#
# df[new_column] = new_series_obj
# ### Applicare una funzione a un oggetto `Series`
#
# L'espressione:
#
# series_obj.apply(fun)
#
# applica la funzione `fun` a tutti i valori in `series_obj` e restituisce un altro oggetto di tipo `Series`.
# ### Applicare una funzione a un oggetto `DataFrame`
#
# L'espressione:
#
# df.applymap(fun)
#
# applica la funzione `fun` a tutti i valori in `df` e restituisce un altro oggetto di tipo `DataFrame`.
# ### Come iterare i record di un data frame
#
# for (index, record) in df.iterrows():
# do_something
# ### Scrivere un data frame su un file in formato `csv`
#
# df.to_csv(file_name, index=False|True)
# ### Richiamare `matplotlib` da Pandas
| laboratorio/lezione11-04nov21/.ipynb_checkpoints/lezione-pandas-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **1**. (10 points)
#
# Consider an eager (i.e. non-lazy) form of the map function that takes a list `xs` to another list `ys` by applying the same function `f` to each member of `xs`. Write a recursive version of map `rmap` and test it on the two simple cases shown below.
list(map(lambda x: x + x, []))
list(map(lambda x: x + x, [1,2,3]))
def rmap(f, xs):
"""Recursive version of map."""
if xs == []:
return []
else:
return [f(xs[0])] + rmap(f, xs[1:])
rmap(lambda x: x + x, [])
rmap(lambda x: x + x, [1,2,3])
# **2**. (25 points)
#
# - Using the `requests` package, download berries 1, 2, and 3 from `https://pokeapi.co/api/v2/berry` in JSON format (5 points)
# - Convert to a `pandas` DataFrame (5 points)
# - Create a new DataFrame that only retains numeric columns and the name column (5 points)
# - Show only rows wheere the name begins with the letter `c` (5 points)
# - Convert to a numpy array (excluding `name`) and standardize so each **row** has mean 0 and standard deviation 1 (5 points)
#
# - The necessary package imports are made for you
import requests
import numpy as np
import pandas as pd
from pandas.api.types import is_numeric_dtype
berries = []
for i in range(1, 4):
berry = requests.get('https://pokeapi.co/api/v2/berry/{}/'.format(i)).json()
berries.append(berry)
berries_df = pd.DataFrame(berries)
idx = np.nonzero([is_numeric_dtype(x) for x in berries_df.dtypes])
idx = np.r_[idx[0], [berries_df.columns.tolist().index('name')]]
df = berries_df.iloc[:, idx]
df[df.name.str.startswith('c')]
xs = df.iloc[:, :-1].values
(xs - xs.mean(axis=1)[:, None])/xs.std(axis=1)[:, None]
# **3**. (25 points)
#
# We have provided an SQLite3 database with 3 tables `dog`, `treat` and `dog_treat`. The `dog_treat` table is a linker table showing which dog ate which treat.
#
# - Show a table of ALL dogs and the treats with calories that they ate with column names `dog`, `treat`, `calorie`. A dog that did not eat any treats should still be present in the table (15 points)
# - Using a common table expression, show a table with two columns `dog` and `total_calories` where only dogs that have eaten more than 500 calories are displayed (5 points)
# %load_ext sql
# %sql sqlite:///pets.db
# %sql SELECT * FROM sqlite_master
# + language="sql"
#
# SELECT dog.name as dog, treat.name as treat, treat.calories
# FROM dog
# LEFT JOIN dog_treat
# ON dog.dog_id = dog_treat.dog_id
# LEFT JOIN treat
# ON dog_treat.treat_id = treat.treat_id
# + language="sql"
#
# with t AS
# (SELECT dog.name as dog, treat.name as treat, treat.calories
# FROM dog
# LEFT JOIN dog_treat
# ON dog.dog_id = dog_treat.dog_id
# LEFT JOIN treat
# ON dog_treat.treat_id = treat.treat_id)
# SELECT dog, SUM(calories) as total_calories
# FROM t
# GROUP BY dog
# HAVING total_calories > 500
# -
# **4**. (40 points)
#
# You want to evaluate whether a liner, quadratic or cubic polynomial is the best model for a set of data using leave-one-out cross-validation (LOOCV) and the mean squared error as evaluation metric. Write a function named `loocv` that takes the predictor variable `x`, the outcome variable `y`, a list of of degrees of polynomial models to be evaluated, and an evaluation function and returns the best model found by LOOCV. For example, you would call the function like this `loocv(x, y, [1,2,3], mse)` where `mse` is of course a function that returns the mean squared error.
#
# - Use the `x` and `y` variables provided
# - Do not use any packages except for the standard library and `numpy`
# - Code snipets for fitting and estimation of polynomials is provided
import numpy as np
x = np.load('x.npy')
y = np.load('y.npy')
coeffs = np.polyfit(x, y, 2)
ypred = np.polyval(coeffs, x)
x
x[np.ones(10).astype('bool')]
def loocv(x, y, degrees, metric):
"""Performs LOOCV to find best polynomial modle."""
n = len(x)
losses = []
for d in degrees:
loss = 0
for i in range(n):
idx = np.ones(n).astype('bool')
idx[i] = False
xx = x[idx]
yy = y[idx]
coeffs = np.polyfit(xx, yy, d)
ypred = np.polyval(coeffs, x[i])
loss += metric(y[i], ypred)
losses.append(loss)
k = np.argmin(losses)
return degrees[k]
def mse(y, ypred):
"""Returns MSE between y and ypred."""
return np.mean((y - ypred)**2)
loocv(x, y, [1,2,3, 4], mse)
| exams/Midterm 01 Mock Exam_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import os
#numpy
import numpy as np
# pandas
import pandas as pd
# matplotlib
import matplotlib.pyplot as plt
# the commonly used alias for seaborn is sns
import seaborn as sns
from pandas import DataFrame
# set a seaborn style of my taste
sns.set_style("whitegrid")
# + deletable=true editable=true
cwd = os.getcwd()
dirs = os.listdir( cwd )
df2 = []
for items in dirs:
if items.find('GSVA')>0 and items.find('.csv')>0:
x=cwd+'/'+items
temp_df = pd.read_csv((x), encoding = "ISO-8859-1")
temp_df = temp_df.loc[::,['S.No.','Item', '2014-15']]
temp_df['State']=items.split('-')[1]
df2.append(temp_df)
usable_gsva=pd.concat(df2)
# + deletable=true editable=true
usable_gsva= usable_gsva.set_index('State')
df_by_state = usable_gsva.groupby('State')
df_by_sector = usable_gsva.groupby('Item')
# + deletable=true editable=true
usable_gsva.head()
# + deletable=true editable=true
print(usable_gsva.info())
# + deletable=true editable=true
#grouping by state
df_by_state = usable_gsva.groupby('State')
# + deletable=true editable=true
#sorting by sector
usable_gsva.set_index('Item')
df_by_sector = usable_gsva.groupby('Item')
df_by_sector = df_by_sector['2014-15'].mean().sort_values(ascending=False)
# + deletable=true editable=true
df_by_sector
# + deletable=true editable=true
#grouping my sector identifying top 5 sectors that are Trade & Repair service, Agriculture Forestry and Fishing, Manufacturing
#real estate, Taxes on Products
df_by_sector['2014-15'].mean().sort_values(ascending=False)
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
| Part1-B-FINAL.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Jupyterlab下简单接触 R 语言
#
# 承接 1-basic-envir/1-get-started.ipynb 最后一段话中关于R的部分,这里稍微给几个R语言的例子。日常积累。首先,是安装R包。这里以USGS 的 dataRetrieval 为例子,如果还未执行过下面句子,就取消注释,运行即可。代码主要参考了[这里](https://usgs-r.github.io/dataRetrieval/articles/tutorial.html)
# +
# install.packages(c("dplyr","dataRetrieval","ggplot2","tidyr","RColorBrewer","EGRET", "rmarkdown", "geoknife"))
# -
# R中使用包前,同样需要先导入。
library(dataRetrieval)
MauiCo_avgdailyQ <- readNWISdata(stateCd="Hawaii", countyCd="Maui", service="dv", parameterCd="00060")
head(MauiCo_avgdailyQ)
# 指定站点id来读取数据看看。
# +
# Choptank River near Greensboro, MD
siteNumber <- "01491000"
ChoptankInfo <- readNWISsite(siteNumber)
parameterCd <- "00060"
#Raw daily data:
rawDailyData <- readNWISdv(siteNumber,parameterCd, "2010-01-01","2011-01-01")
head(rawDailyData)
# +
siteNo <- "01491000"
pCode <- "00060"
start.date <- "2010-01-03"
end.date <- "2010-01-03"
pheasant <- readNWISuv(siteNumbers = siteNo,
parameterCd = pCode,
startDate = start.date,
endDate = end.date)
# -
names(pheasant)
head(pheasant)
# 可以注意到每天的数据并不是从0点开始的,也不是到0点结束的,因为美国各地时区不同,为了统一,在日期表示上,统一采用了UTC时间,而上例中的 Choptank River near Greensboro, MD 时区是美东时区,当地早上5点是UTC的0点。所以当输入是UTC 0点时候,就是从当地5点开始计算的。因此,日均值也是基于当地时间的。
# 做一些简单的统计,可以看看summary。
summary(pheasant)
| 1-basic-envir/BonusR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML/DL techniques for Tabular Modeling PART III
# > In the last part on Tabular Modeling a Neural Network is trained on the Kaggle Dataset. Embedding from a NN are also explained.
# - toc: true
# - badges: true
# - comments: true
# +
#hide
# # !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
# +
#hide
from fastbook import *
from kaggle import api
from pandas.api.types import is_string_dtype, is_numeric_dtype, is_categorical_dtype
from fastai.tabular.all import *
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from dtreeviz.trees import *
from IPython.display import Image, display_svg, SVG
import numpy as np
import matplotlib.pyplot as plt
pd.options.display.max_rows = 20
pd.options.display.max_columns = 8
def r_mse(pred,y): return round(math.sqrt(((pred-y)**2).mean()), 6)
def m_rmse(m, xs, y): return r_mse(m.predict(xs), y)
# -
#hide
df = pd.read_csv('/home/nitish/Downloads/bluebook-bulldozers/TrainAndValid.csv', low_memory=False)
# ## Introduction
# In this, the last part on tabular modeling, we will look into training a Neural Network (NN) on the Kaggle Dataset. In the last two parts we have seen decision trees, and random forests and look into their advantages/disadvantages. Their main advantage is their simplicity, faster training, and disadvantage being unable to extrapolate on out of domain data. Firstly let’s train a NN on the random dataset to examine its extrapolation power.
# ## Extrapolation
#
# +
x = np.linspace(0, 10, 110)
y = x + np.random.randn(110)
df_lin=pd.concat([pd.DataFrame(x, columns=['x']), pd.DataFrame(y, columns=['y'])], axis = 1)
cont_names=['x']
cat_names=[]
cond = (df_lin.x<8)
train_idx = np.where( cond)[0]
valid_idx = np.where(~cond)[0]
splits = (list(train_idx),list(valid_idx))
procs_nn = [Normalize]
to_nn = TabularPandas(df_lin, procs_nn, cont_names=cont_names, cat_names=cat_names, splits=splits, y_names=['y'])
dls = to_nn.dataloaders(30)
y.min(), y.max()
learn = tabular_learner(dls, y_range=(-2,13), layers=[100,10],
n_out=1, loss_func=F.mse_loss)
learn.fit_one_cycle(30, 5e-2)
preds,targs = learn.get_preds(0)
v_preds,v_targs = learn.get_preds(0)
tr_res,_ = learn.get_preds(0)
val_res,_ = learn.get_preds(1)
df_lin.iloc[valid_idx]
fig, ax = plt.subplots(figsize=(16,8))
ax.scatter(x,y, marker='+', label='actual data')
ax.scatter(df_lin.iloc[train_idx]['x'], tr_res, label='training data prediction')
ax.scatter(df_lin.iloc[valid_idx]['x'], val_res, label='validation data prediction')
ax.xaxis.set_major_locator(mpl.ticker.MultipleLocator(1))
ax.grid(which='major', axis='both', linestyle=':', linewidth = 1, color='b')
ax.set_xlabel("x", labelpad=5, fontsize=26, fontname='serif', color="blue")
ax.set_ylabel("y", labelpad=5, fontsize=26, fontname='serif', color="blue")
ax.legend(prop={"size":15})
# -
# We can see clearly from the above figure that the neural network is giving good results, and far better than decision trees and RF, on extrapolation. This is because a neural network could fit a complex non-linear function easily, and could generalize better than on unseen data. Now let's see the neural network performance on the Kaggle dataset.
# +
df = pd.read_csv('/home/nitish/Downloads/bluebook-bulldozers/TrainAndValid.csv', low_memory=False)
sizes = ('Large','Large / Medium','Medium','Small','Mini','Compact')
df['ProductSize'] = df['ProductSize'].astype('category')
df['ProductSize'].cat.set_categories(sizes, ordered=True, inplace=True)
dep_var = 'SalePrice'
df[dep_var] = np.log(df[dep_var])
df = add_datepart(df, 'saledate')
procs = [Categorify, FillMissing]
cond = (df.saleYear<2011) | (df.saleMonth<10)
train_idx = np.where( cond)[0]
valid_idx = np.where(~cond)[0]
splits = (list(train_idx),list(valid_idx))
cont,cat = cont_cat_split(df, 1, dep_var=dep_var)
to = TabularPandas(df, procs, cat, cont, y_names=dep_var, splits=splits)
xs,y = to.train.xs,to.train.y
valid_xs,valid_y = to.valid.xs,to.valid.y
xs.loc[xs['YearMade']<1900, 'YearMade'] = 1950
valid_xs.loc[valid_xs['YearMade']<1900, 'YearMade'] = 1950
m = DecisionTreeRegressor()
m.fit(xs, y);
def rf(xs, y, n_estimators=40, max_samples=200_000,
max_features=0.5, min_samples_leaf=5, **kwargs):
return RandomForestRegressor(n_jobs=-1, n_estimators=n_estimators,
max_samples=max_samples, max_features=max_features,
min_samples_leaf=min_samples_leaf, oob_score=True).fit(xs, y)
m = rf(xs, y)
def rf_feat_importance(m, df):
return pd.DataFrame({'cols':df.columns, 'imp':m.feature_importances_}
).sort_values('imp', ascending=False)
fi = rf_feat_importance(m, xs)
fi[:10]
to_keep = fi[fi.imp>0.005].cols
xs_imp = xs[to_keep]
valid_xs_imp = valid_xs[to_keep]
to_drop = ['saleYear', 'ProductGroupDesc', 'fiBaseModel', 'Grouser_Tracks']
xs_final = xs_imp.drop(to_drop, axis=1)
valid_xs_final = valid_xs_imp.drop(to_drop, axis=1)
df_dom = pd.concat([xs_final, valid_xs_final])
is_valid = np.array([0]*len(xs_final) + [1]*len(valid_xs_final))
m = rf(df_dom, is_valid)
rf_feat_importance(m, df_dom)[:6]
m = rf(xs_final, y)
# print('orig', m_rmse(m, valid_xs_final, valid_y))
time_vars = ['SalesID','MachineID']
xs_final_time = xs_final.drop(time_vars, axis=1)
valid_xs_time = valid_xs_final.drop(time_vars, axis=1)
################ NN
df_nn = pd.read_csv('/home/nitish/Downloads/bluebook-bulldozers/TrainAndValid.csv', low_memory=False)
df_nn['ProductSize'] = df_nn['ProductSize'].astype('category')
df_nn['ProductSize'].cat.set_categories(sizes, ordered=True, inplace=True)
df_nn[dep_var] = np.log(df_nn[dep_var])
df_nn = add_datepart(df_nn, 'saledate')
df_nn_final = df_nn[list(xs_final_time.columns) + [dep_var]]
cont_nn,cat_nn = cont_cat_split(df_nn_final, dep_var=dep_var)
cont_nn.append('saleElapsed')
cat_nn.remove('saleElapsed')
cat_nn.remove('fiModelDescriptor')
procs_nn = [Categorify, FillMissing, Normalize]
to_nn = TabularPandas(df_nn_final.astype({'saleElapsed': 'int64'}), procs_nn, cat_nn, cont_nn,
splits=splits, y_names=dep_var)
dls = to_nn.dataloaders(1024)
from fastai.tabular.all import *
learn = tabular_learner(dls, y_range=(8,12), layers=[500,250],
n_out=1, loss_func=F.mse_loss)
learn.fit_one_cycle(5, 1e-2)
preds,targs = learn.get_preds()
r_mse(preds,targs)
# -
# We are getting rmse of 0.25 here, and each epoch takes ~40 seconds, which is on CPU, on GPU it will take around 1 2 seconds. If we train for 15 epochs total we could achieve ~0.23 rmse, same as rf. I didn't train for 15 more epochs because while writing the blog I was on CPU ;-)
# ## Categorical Embeddings
# If we look at the architecture of our model, we can see that before the first layer there is an embedding layer. It's for categorical variables. So rather than keeping the categorical variables category-values in the df and use one-hot encoding, NN should use embeddings instead because of two following 2 main reasons:
# 1. One-Hot encoding is expensive both space-wise and computationally.
# 1. We can use the embeddings we get to get the idea about the relationship of different categories.
# Reason for 1 is that if a categorical column has 5000 distinct categories, and there are a total of 1000 training examples, then it will use $5$x$10^6$ length array just for feeding the input to the NN, which would be infeasible if these numbers are bigger.
#
# Reason for 2 is suppose we get embeddings for words then basically we are getting a fixed dimensional representation of each of the word, and we can do various mathematical operations on the word-embeddings. E.g., we can find the distance between any two words which could be interpreted as how similar or distinct the words are, cluster the similar words together, etc.
#
#
# In Fastai, the target dimension of an embedding is found out by the learner itself during training. So, there is a total of 10 categorical variables, so 11 embedding matrices are shown below, where for example the third embedding matrix is of shape (75, 18), meaning it has 75 distinct categories and it's calculating an 18-dim vector for each of the category.
learn.model
# ## Using Embeddings in a Random Forest
# There is a paper, "Entity Embeddings of Categorical Variables" which tells about embeddings of categorical variables. The abstract of the paper is:
# > Entity embedding not only reduces memory usage and speeds up neural networks compared with one-hot encoding, but more importantly by mapping similar values close to each other in the embedding space it reveals the intrinsic properties of the categorical variables... [It] is especially useful for datasets with lots of high cardinality features, where other methods tend to overfit... As entity embedding defines a distance measure for categorical variables it can be used for visualizing categorical data and for data clustering.
# We can use the embeddings we got for different categorical variables as features and train a random forest on this new data. It can be seen that RF with embeddings instead of raw categories performs better. I have tried using NN embeddings and trained an RF on top of it, but get only slightly better results (might be some bug/issue with my code). We can get the embedding matrix like:
cat_nn[0],list(learn.model.embeds.parameters())[0]
# This is the 7x5 embedding matrix for the categorical variable ProductSize. There are 6 distinct categories in this feature, and one category (the last one) is na, for missing ones or not applicable.
#
cat=[]
emb=[]
for idx, item in enumerate(list(learn.model.embeds.parameters())):
cat.append(cat_nn[idx])
emb.append(item.shape)
df_ = pd.DataFrame()
df_['categories']=cat
df_['embeddings']=emb
df_
# ## Conclusion
# We have covered the basic ML techniques, a NN network, and using embeddings obtained from a NN in an RF.
# We have seen better extrapolation by a NN than RF. We have trained a simple, not so deep NN on the Kaggle dataset which gives the same rmse or a little better than RF.
| _notebooks/2020-11-24-DLforCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Perform Batch Inference (Predictions) using SageMaker Batch Transform
# ### Imports
from sagemaker import get_execution_role
from time import gmtime, strftime
import pandas as pd
import sagemaker
import boto3
import time
# ### 1. Essentials
BUCKET = sagemaker.Session().default_bucket()
PREFIX = 'clf'
REGION = 'us-east-1'
batch_input = f's3://{BUCKET}/{PREFIX}/batch_test/'
batch_input
batch_output = f's3://{BUCKET}/{PREFIX}/batch_test_out/'
batch_output
current_timestamp = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
# +
TRAINING_JOB_NAME = 'classifier-2020-11-08-22-53-37-824' # Copy this from the console
MODEL_NAME = f'clf-xgboost-model-{current_timestamp}'
BATCH_JOB_NAME = f'clf-xgboost-batch-job-{current_timestamp}'
session = boto3.Session()
sagemaker_execution_role = get_execution_role()
sagemaker_session = sagemaker.session.Session()
sagemaker_client = boto3.client('sagemaker', region_name=REGION)
s3_client = boto3.client('s3')
container_uri = sagemaker.image_uris.retrieve(region=session.region_name,
framework='xgboost',
version='1.0-1',
image_scope='training')
# -
# ### 2. Create a Model object using previously run training job name
info = sagemaker_client.describe_training_job(TrainingJobName=TRAINING_JOB_NAME)
info
model_artifact_url = info['ModelArtifacts']['S3ModelArtifacts']
model_artifact_url
primary_container = {
'Image': container_uri,
'ModelDataUrl': model_artifact_url
}
response = sagemaker_client.create_model(
ModelName=MODEL_NAME,
ExecutionRoleArn=sagemaker_execution_role,
PrimaryContainer=primary_container)
response
# ### 3. Create a Batch Transformer for Inference
request = {
"TransformJobName": BATCH_JOB_NAME,
"ModelName": MODEL_NAME,
"BatchStrategy": "MultiRecord",
"TransformOutput": {
"S3OutputPath": batch_output
},
"TransformInput": {
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": batch_input
}
},
"ContentType": "text/csv",
"SplitType": "Line",
"CompressionType": "None"
},
"TransformResources": {
"InstanceType": "ml.m5.xlarge",
"InstanceCount": 1
}
}
response = sagemaker_client.create_transform_job(**request)
response
while(True):
response = sagemaker_client.describe_transform_job(TransformJobName=BATCH_JOB_NAME)
status = response['TransformJobStatus']
if status == 'Completed':
print("Transform job ended with status: {}".format(status))
break
if status == 'Failed':
message = response['FailureReason']
print('Transform failed with the following error: {}'.format(message))
raise Exception('Transform job failed')
print("Transform job is still in status: {}".format(status))
time.sleep(30)
# ### 4. Evaluate Output
key = f'{PREFIX}/batch_test_out/batch_test.csv.out'
obj = s3_client.get_object(Bucket=BUCKET, Key=key)
results_df = pd.read_csv(obj['Body'], names=['Predictions'])
results_df
| 4. Batch Transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DPU example: Resnetv1 using Vitis AI Runtime (VART)
#
# This notebooks shows an example of DPU applications. The application,
# as well as the DPU IP, is pulled from the official
# [Vitis AI Github Repository](https://github.com/Xilinx/Vitis-AI).
# For more information, please refer to the
# [Xilinx Vitis AI page](https://www.xilinx.com/products/design-tools/vitis/vitis-ai.html).
#
# In this notebook, we will show how to use **VART** to run DPU tasks.
#
# ## 1. Prepare the overlay
# We will download the overlay onto the board.
#
# By default, `DpuOverlay` object will set the DPU runtime to be `dnndk`;
# in this example, we will change it to `vart`. You cannot use them both at the
# same time because of memory allocation conflicts - users need to choose either one.
#
# The `load_model()` method will automatically prepare the `graph`
# which is used by VART.
from pynq_dpu import DpuOverlay
overlay = DpuOverlay("dpu.bit")
overlay.set_runtime("vart")
overlay.load_model("dpu_tf_resnetv1_0.elf")
# ## 2. Utility functions
#
# In this section, we will prepare a few functions for later use.
import os
import time
import numpy as np
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# Let's first define a few useful preprocessing functions. These functions
# will make sure the DPU can take input images with arbitrary sizes.
# +
_R_MEAN = 123.68
_G_MEAN = 116.78
_B_MEAN = 103.94
MEANS = [_B_MEAN,_G_MEAN,_R_MEAN]
def resize_shortest_edge(image, size):
H, W = image.shape[:2]
if H >= W:
nW = size
nH = int(float(H)/W * size)
else:
nH = size
nW = int(float(W)/H * size)
return cv2.resize(image,(nW,nH))
def mean_image_subtraction(image, means):
B, G, R = cv2.split(image)
B = B - means[0]
G = G - means[1]
R = R - means[2]
image = cv2.merge([R, G, B])
return image
def BGR2RGB(image):
B, G, R = cv2.split(image)
image = cv2.merge([R, G, B])
return image
def central_crop(image, crop_height, crop_width):
image_height = image.shape[0]
image_width = image.shape[1]
offset_height = (image_height - crop_height) // 2
offset_width = (image_width - crop_width) // 2
return image[offset_height:offset_height + crop_height, offset_width:
offset_width + crop_width, :]
def normalize(image):
image=image/256.0
image=image-0.5
image=image*2
return image
def preprocess_fn(image, crop_height = 224, crop_width = 224):
image = resize_shortest_edge(image, 256)
image = mean_image_subtraction(image, MEANS)
image = central_crop(image, crop_height, crop_width)
return image
# -
# We will also define a few functions to calculate softmax and provide
# the output class after running a DPU task.
# +
def calculate_softmax(data):
result = np.exp(data)
return result
def predict_label(softmax):
with open("img/words.txt", "r") as f:
lines = f.readlines()
return lines[np.argmax(softmax)-1]
# -
# Keep in mind that our original images are 640x480 so we need to preprocess them
# later to make sure it fits our model.
image_folder = 'img'
original_images = [i for i in os.listdir(image_folder) if i.endswith("JPEG")]
total_images = len(original_images)
# ## 3. Use VART
# Now we should be able to use VART to do image classification.
# +
dpu = overlay.runner
inputTensors = dpu.get_input_tensors()
outputTensors = dpu.get_output_tensors()
outputHeight = outputTensors[0].dims[1]
outputWidth = outputTensors[0].dims[2]
outputChannel = outputTensors[0].dims[3]
outputSize = outputHeight*outputWidth*outputChannel
softmax = np.empty(outputSize)
# -
# We can define a few buffers to store input and output data. They will be reused
# during multiple runs.
shape_in = (1,) + tuple(
[inputTensors[0].dims[i] for i in range(inputTensors[0].ndim)][1:])
shape_out = (1, outputHeight, outputWidth, outputChannel)
input_data = []
output_data = []
input_data.append(np.empty((shape_in),
dtype = np.float32, order = 'C'))
output_data.append(np.empty((shape_out),
dtype = np.float32, order = 'C'))
image = input_data[0]
# Remember that we have a list of `original_images`.
# We can now define a new function `run()` which takes the image index as
# the input, and calculate the softmax as the classification result.
# With the argument `display` set to `True`, the original image as well as the
# predicted label can be rendered.
#
# It is obvious that the range of `image_index` should be [0, `total_images`-1].
def run(image_index, display=False):
preprocessed = preprocess_fn(cv2.imread(
os.path.join(image_folder, original_images[image_index])))
image[0,...] = preprocessed.reshape(
inputTensors[0].dims[1],
inputTensors[0].dims[2],
inputTensors[0].dims[3])
job_id = dpu.execute_async(input_data, output_data)
dpu.wait(job_id)
temp = [j.reshape(1, outputSize) for j in output_data]
softmax = calculate_softmax(temp[0][0])
if display:
display_image = cv2.imread(os.path.join(
image_folder, original_images[image_index]))
_, ax = plt.subplots(1)
_ = ax.imshow(cv2.cvtColor(display_image, cv2.COLOR_BGR2RGB))
print("Classification: {}".format(predict_label(softmax)))
# Let's run it for 1 image and print out the predicted label.
run(1, display=True)
# We can also run it for multiple images as shown below. In this example
# we have only used 1 thread; in principle, users should be able to boost
# the performance by employing more threads.
time1 = time.time()
[run(i) for i in range(total_images)]
time2 = time.time()
fps = total_images/(time2-time1)
print("Performance: {} FPS".format(fps))
# Copyright (C) 2020 Xilinx, Inc
| pynq_dpu/edge/notebooks/dpu_tf_resnetv1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''.venv'': venv)'
# name: python3
# ---
# +
import time
from PIL import Image, ImageOps
import numpy as np
import matplotlib.pyplot as plt
import cv2
import pyocr
import pyocr.builders
# %matplotlib inline
# -
# ## Init tesseract
# +
tools = pyocr.get_available_tools()
print('Tools:')
for t in tools:
print(f' {t.__name__}')
ocr = tools[0]
print('langs:')
for lang in ocr.get_available_languages():
print(f' {lang}')
# -
# ## Line Box
# +
origin = Image.open('../testdata/title2.jpg')
inv = ImageOps.invert(origin)
for lang in ['jpn', 'jpn_best', 'jpn_fast']:
start = time.time()
lines = ocr.image_to_string(
inv,
lang=lang,
builder=pyocr.builders.LineBoxBuilder())
elapsed = time.time() - start
img = np.array(origin)
for line in lines:
print(line.content)
# line の描画
(l, t), (r, b) = line.position
cv2.rectangle(img, (l, t), (r, b), (0, 0, 255), 10)
cv2.line(img, (l, (t + b) //2), (r, (t + b) //2), (0, 0, 255), 10)
# word box の描画
for box in line.word_boxes:
ul, lr = box.position
cv2.rectangle(img, ul, lr, (0, 255, 0), 10)
ul, lr = line.position
print('lang:', lang)
print('elapsed time:', elapsed)
plt.imshow(img)
plt.show()
# -
# ## Word Box
for lang in ['jpn', 'jpn_best', 'jpn_fast']:
img = Image.open('../testdata/title2.jpg')
start = time.time()
txt = ocr.image_to_string(
ImageOps.invert(img),
lang=lang,
builder=pyocr.builders.TextBuilder()) # type: ignore
elapsed = time.time() - start
img = np.array(img)
print('lang:', lang)
print('elapsed time:', elapsed)
print(txt)
| notebooks/ocr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# coding: utf-8
import sys, os
sys.path.append(os.pardir) # 親ディレクトリのファイルをインポートするための設定
import numpy as np
import matplotlib.pyplot as plt
from dataset.mnist import load_mnist
from deep_convnet import DeepConvNet
from common.trainer import Trainer
(x_train, t_train), (x_test, t_test) = load_mnist(flatten=False)
network = DeepConvNet()
trainer = Trainer(network, x_train, t_train, x_test, t_test,
epochs=20, mini_batch_size=100,
optimizer='Adam', optimizer_param={'lr':0.001},
evaluate_sample_num_per_epoch=1000)
trainer.train()
# パラメータの保存
network.save_params("deep_convnet_params.pkl")
print("Saved Network Parameters!")
| ch08/train_deepnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import math
# import jtplot module in notebook
from jupyterthemes import jtplot
# choose which theme to inherit plotting style from
# onedork | grade3 | oceans16 | chesterish | monokai | solarizedl | solarizedd
jtplot.style(theme='monokai')
# set "context" (paper, notebook, talk, poster)
# scale font-size of ticklabels, legend, etc.
# remove spines from x and y axes and make grid dashed
jtplot.style(context='talk', fscale=1.4, spines=False, gridlines='--')
# turn on X- and Y-axis tick marks (default=False)
# turn off the axis grid lines (default=True)
# and set the default figure size
jtplot.style(ticks=True, grid=False, figsize=(10, 10))
# -
# # Week 1
#
# ## Overview
#
# As explained in the [*Before week 1* notebook](https://nbviewer.jupyter.org/github/lalessan/comsocsci2021/blob/master/lectures/Before_week_1.ipynb), each week of this class is a Jupyter notebook like this one. **_In order to follow the class, you simply start reading from the top_**, following the instructions.
#
# **Hint**: And you can ask me for help at any point if you get stuck!
# ## Today
#
# This first lecture will go over a few different topics to get you started
#
# * First, we will learn about Computational Social Science.
# * Second, we talk a bit about APIs and how they work.
# * Third, we'll use an API to download Reddit data from the _r/wallstreetbet_ subreddit
#
#
# ## Part 1: Computational Social Science
#
#
# But _What is Computational Social Science_? Watch the video below, where I will give a short introduction to the topic.
#
# > **_Video lecture_**: Watch the video below about Computational Social Science
from IPython.display import YouTubeVideo
YouTubeVideo("qoPk_C3buD8",width=600, height=337.5)
# Now that you have learnt what Computational Social Science, read about the advantages and challenges of using _"Big Data"_ for Social Science Research in Sections 2.1 to 2.3 of the book Bit by Bit.
#
# > _Reading_: [Bit by Bit, sections 2.1 to 2.3](https://www.bitbybitbook.com/en/1st-ed/observing-behavior/observing-intro/) Read sections 2.1 and 2.3, then skim through section 2.3. The idea is for you to understand, in general terms, advantages and challenges of large observational datasets (a.k.a. Big Data) for social studies.
# > *Exercise 1*: This year, lockdowns have helped governments contain the pandemic. But they also negatively impacted our wellbeing. Imagine you had to study the following question: "_What are some of the strategies people adopt to preserve their mental and physical wellbeing during lockdown?_"
# > * Write in a couple of lines:
# >> * Which data would you collect to study this topic?
# >> * How would you collect it?
# > * Describe the data you would need more in details (also by writing down a couple of lines):
# >> * How big is the data (number of users/number of data points)?
# >> * Which variables it contains?
#
# ## Part 2: Using APIs to download Reddit data
#
# But what is an API? Find the answer in the short video below, where we get familiar with APIs to access Reddit data.
#
# > **_Video lecture_**: Watch the video below about the Reddit API
from IPython.display import YouTubeVideo
YouTubeVideo("eqBIFua00O4",width=600, height=337.5)
import datetime
from psaw import PushshiftAPI
api = PushshiftAPI()
features = ["title", "id", "score", "author", "num_comments", "created_utc"]
subreddit = "wallstreetbets"
date1 = int(datetime.datetime(2020,1,1).timestamp())
date2 = int(datetime.datetime(2021,1,25).timestamp())
query = "GME|Gamestop"
gen = api.search_submissions(subreddit = subreddit, after = date1, before = date2, q = query, filter = features)
results = list(gen)
# +
# title, id, score, date of creation, author, and number of comments
# id, title, 'score', 'created_utc', 'author', 'num_comments'
ids, title, score, created, author, num_comments = [], [], [], [], [], []
for i in range(len(results)):
ids.append(i)
title.append(results[i].d_['title'])
score.append(results[i].d_['score'])
created.append(results[i].d_['created_utc'])
author.append(results[i].d_['author'])
num_comments.append(results[i].d_['num_comments'])
data = pd.DataFrame(
{
"id" : ids,
"title" : title,
"score" : score,
"created" : created,
"author" : author,
"num_comments" : num_comments
}, index = None)
# -
from datetime import datetime
dates = [datetime.utcfromtimestamp(res.created) for res in results]
data["created"] = dates
data
data.describe()
data = data.set_index("created")
post_counts = data.resample("1D")["title"].count()
# +
plt.plot(post_counts)
plt.show()
week_authors = data.resample("1W")["author"].nunique()
plt.plot(week_authors)
plt.show()
# -
# It's time for you to get to work. Take a look at the two texts below - just to get a sense of a more technical description of how the Pushshift API works.
#
#
# > _Reading_ (just skim): [New to Pushshift? Read this! FAQ](https://www.reddit.com/r/pushshift/comments/bcxguf/new_to_pushshift_read_this_faq/)
# > _Reading_ (just skim): [Pushshift Github Repository](https://github.com/pushshift/api)
# >
# ## Prelude to part 3: Pandas Dataframes
#
# Before starting, we will also learn a bit about [pandas dataframes](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html), a very user-friendly data structure that you can use to manipulate tabular data. Pandas dataframes are implemented within the [pandas package](https://pandas.pydata.org/).
#
# Pandas dataframes should be intuitive to use. **I suggest you to go through the [10 minutes to Pandas tutorial](https://pandas.pydata.org/pandas-docs/stable/user_guide/10min.html) to learn what you need to solve the next exercise.**
# ## Part 3: Getting data from the _r/wallstreetbet_ subreddit
# There has been a lot of interest in the social platform Reddit this week, after investors from the [_r/wallstreetbet_](https://www.reddit.com/r/wallstreetbets/) subreddit managed to [give a huge boost](https://www.google.com/search?q=GME+price&oq=GME+price&aqs=chrome..69i57.1261j0j4&sourceid=chrome&ie=UTF-8) to the shares of the video game retailer's GameStop (traded as "_GME_"), causing massive losses to professional investors and established hedge funds.
#
# There is so much buzz about _Gamestop_ because it is really something unprecedented! Online discussions about stocks on social media have fuelled massive price moves that cannot be explained by traditional valuation metrics and can seriously destabilize the established market. Many ordinary investors on Reddit have coordinated to buy shares of a stock that had been losing value for a long time. __But how did this all happen?__
#
# Today and in the following classes, we will try to answer precisely this question, by studying the social network of Redditors of _r/wallstreetbet_ throughout last year.
# The starting point will be to understand how to download data from Reddit using APIs. But before we start getting our hands diry, if you feel like you don't know much about Gamestop, I suggest to watch this short video summarizing the latest events. If you already know everything about it, feel free to skip it.
#
# >
# > **_Video_**: [Stocks explained: What's going on with GameStop?](https://www.bbc.com/news/av/technology-55864312)
# >
# > *Exercise 2*: __Download submissions of the [_r/wallstreetbet_](https://www.reddit.com/r/wallstreetbets/) subreddit using the [Pushift API](https://github.com/pushshift/api)__
# > 1. Use the [psaw Python library](https://pypi.org/project/psaw/) (a wrapper for the Pushshift API) to find all the submissions in subreddit _r/wallstreetbet_', related to either "_GME_" or "_Gamestop_" (**Hint**: Use the [``q``](https://github.com/pushshift/api) parameter to search text. To search multiple words you can separate them with character "|"). Focus on the period included __between Jan,1st 2020 and Jan 25th, 2021__, where time must be provided in [Unix Timestamp](https://www.unixtimestamp.com/). _Note: The Pushift API returns at most 100 results per query, so you may need to divide your entire time period in small enough sub-periods._
# > 2. For each submission, find the following information: __title, id, score, date of creation, author, and number of comments__ (**Hint**: access the dictionary with all attributes by typing ``my_submission.d_``). Store this data in a pandas DataFrame and save it into a file. (Downloading required me 30 minutes using two cores. While you wait for the results, you can start thinking about _Exercise 3_).
# > 3. Create a figure using [``matplotlib``](https://matplotlib.org/) and plot the total number of submissions per day (**Hint**: You can use the function [``datetime.datetime.utcfromtimestamp``](https://docs.python.org/3/library/datetime.html) to convert a timestamp into a date, and you can use the function [``pd.resample``](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.resample.html) to aggregate by day). What do you observe?
# > 4. How many submissions have you downloaded in total? How many unique authors?
# > 5. _Optional_: How many unique authors are there each week in the period under study?
#
#
# > *Exercise 3*: __Download comments from the [_r/wallstreetbet_](https://www.reddit.com/r/wallstreetbets/) subreddit.__ The second task for today is to download the comments associated to each submission, which we will use to build the social network of Redditers.
# > 1. For each submission you found in _Exercise 2_, download all the comments (*Hint*: Use the [``search_comments``](https://github.com/pushshift/api) function to search comments. You can specify the parameter ``link_id``, which corresponds to the _id_ of the submission for which you require comments).
# > 2. For each comment, store the following information: __title, id, submission, score, date of creation, author, and number of comments__. Store this in a pandas DataFrame and save it into a file. We will use it in the next classes.
#
# > __Note__: It took me about a night to get the data for _Exercise 3_. I guess Pushshift servers are going through increasing stress due to the raising interest in the Gamestop saga. If you experience extremely slow downloading time, reach out to me! If you are brave, you can also check out the Reddit API, which is wrapped by [praw](https://praw.readthedocs.io/en/latest/tutorials/comments.html). It functions very much like psaw, but it requires you to first get credentials [here](https://www.reddit.com/prefs/apps) (click on _Create another app_)
from tqdm import tqdm
print(results[0].d_)
ids = []
for i in range(len(results)):
ids.append(results[i].d_['id'])
results[0].d_
features = [ "id","parent_id", "link_id", "score", "created_utc", "author"]
comments = list()
for i in tqdm(range(0,len(data),100)):
comments += list(api.search_comments(link_id = ids[i:i+100], filter = features))
# +
ids, parent_id ,link_id, score, created, author = [], [], [], [], [], []
for i in range(len(comments)):
ids.append(results[i].d_['id'])
link_id.append(results[i].d_['link_id'])
parent_id.append()
score.append(results[i].d_['score'])
created.append(results[i].d_['created_utc'])
author.append(results[i].d_['author'])
data = pd.DataFrame(
{
"id" : ids,
"link_id" : link_id,
"score" : score,
"created" : created,
"author" : author,
}, index = None)
data.to_csv('reddit_comments', encoding='utf-8')
# -
| lectures/Week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fKwkP2ix7H73"
# # Download Dataset of Google
#
# Download data from below link with mark and area and label
# ```
# https://storage.googleapis.com/openimages/web/visualizer/index.html?set=train&type=detection&c=%2Fm%2F01mqdt
# ```
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 34340, "status": "ok", "timestamp": 1637089195165, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgY2XonBoTwStMW5U7xXRwb8HO5ET2tooeBTzhP0Q=s64", "userId": "17922861448141550220"}, "user_tz": 300} id="DUs40DGb-E7F" outputId="49fe221a-f5d1-4ad9-8ad6-d85f559215e2"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="dzoOv204eoNj"
# ### download model
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2715, "status": "ok", "timestamp": 1637089197869, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgY2XonBoTwStMW5U7xXRwb8HO5ET2tooeBTzhP0Q=s64", "userId": "17922861448141550220"}, "user_tz": 300} id="ROxxeGAaem65" outputId="d0189708-ae2c-4050-f688-e85a86932f37"
# clone darknet repo
# % cd ../gdrive/MyDrive/Yolo/yolov3
# !git clone https://github.com/AlexeyAB/darknet
# + [markdown] id="nqqMQaSmeRIp"
# ### download data
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="sGKHnxS36c6b"
# ! git clone https://github.com/EscVM/OIDv4_ToolKit.git
# ! mkdir dataset/google
% cd OIDv4_ToolKit
# ! pip3 install -r requirements.txt
# ! python3 main.py downloader --classes Traffic_sign --type_csv validation --Dataset ../../dataset/google/ -y
# ! python3 main.py downloader --classes Traffic_sign --type_csv test --Dataset ../../dataset/google/ -y
# ! python3 main.py downloader --classes Traffic_sign --type_csv train --Dataset ../../dataset/google/ -y
# + [markdown] id="zeAs05FJF1re"
# ### organize the data
# + colab={"background_save": true} id="XnFKmahHGtzO"
% cd ../dataset
# ! mkdir obj
# ! cp google/test/Traffic\ sign/Label/* obj
# ! cp google/test/Traffic\ sign/*.jpg obj
# ! cp google/train/Traffic\ sign/Label/* obj
# ! cp google/train/Traffic\ sign/*.jpg obj
# ! cp google/validation/Traffic\ sign/Label/* obj
# ! cp google/validation/Traffic\ sign/*.jpg obj
# + colab={"background_save": true} id="-Vr-0hoYJf_x"
# !sed -i 's/Traffic sign/0/g' obj/*.txt
# + colab={"background_save": true} id="pCc90TxT0Ext"
# ! pwd
# + [markdown] id="7ppHhoNUHdvP"
# ### change annotation
# + colab={"background_save": true} id="fwqAPYqzKRrw"
import os
filelist = os.listdir('obj')
textfilelist = list(filter(lambda x : '.txt' in x, filelist))
# + colab={"background_save": true} id="MnzVvZ1W2J_W"
import cv2
for textfile in textfilelist :
file = open("obj/"+textfile,mode='r+')
all_of_it = file.read()
lines = all_of_it.split('\n')[:-1]
file.close()
im = cv2.imread('obj/'+textfile.split('.txt')[0]+'.jpg')
h, w, c = im.shape
print(h, w, c)
new_file = open("obj/"+textfile, "w+")
for line in lines :
print(line)
content = line.split()
print(content)
min_x = float(content[1])
min_y = float(content[2])
max_x = float(content[3])
max_y = float(content[4])
content[1] = str(float(min_x/w))
content[2] = str(float(min_y/h))
content[3] = str(float(max_x/w))
content[4] = str(float(max_y/h))
print(content)
new_line = " ".join(content)+"\n"
print(new_line)
new_file.write(new_line)
new_file.close()
# break
# + colab={"background_save": true} id="j-eDHUN8ZAWY"
# !cp -r obj ../darknet/data
# + [markdown] id="rsQfoRgUd1J6"
# ### write obj.names
# + colab={"background_save": true} id="pig1X4MhLSjJ"
# !echo "Traffic sign" > ../darknet/data/obj.names
# + [markdown] id="Ux2a3Zd4d5tR"
# ### write obj.data
#
# + colab={"background_save": true} id="nOJXy26Ad8wY"
# !echo -e """classes= 1\ntrain = data/train.txt\nvalid = data/test.txt\nnames = data/obj.names\nbackup = /content/gdrive/MyDrive/Yolo/yolov3""" > ../darknet/data/obj.data
# + [markdown] id="Fn0G4lEtrIUz"
# ### make train.txt
#
# + colab={"background_save": true} id="a6mJ5mL-B0f9"
% pwd
% cd ..
# + colab={"background_save": true} id="qEckF8LHrJNb"
import os
image_files = []
os.chdir(os.path.join("dataset", "obj"))
for filename in os.listdir(os.getcwd()):
if filename.endswith(".jpg"):
image_files.append("data/obj/" + filename)
os.chdir("..")
% pwd
with open("../darknet/data/train.txt", "w") as outfile:
for image in image_files:
outfile.write(image)
outfile.write("\n")
outfile.close()
os.chdir("..")
# + [markdown] id="Wa1eX9lGmOjd"
# ### make cfg file
# + colab={"background_save": true} id="bbi6MdgFhj3G"
% cd darknet
# ! cp cfg/yolov3.cfg cfg/yolov3_traffic_signal.cfg
# !sed -i 's/batch=1/batch=8/' cfg/yolov3_traffic_signal.cfg
# !sed -i 's/subdivisions=1/subdivisions=1/' cfg/yolov3_traffic_signal.cfg
# !sed -i 's/max_batches = 500200/max_batches = 2200/' cfg/yolov3_traffic_signal.cfg
# !sed -i 's/steps=400000,450000/steps=1600,1800/' cfg/yolov3_traffic_signal.cfg
# !sed -i 's/filters=255/filters=18/g' cfg/yolov3_traffic_signal.cfg
# !sed -i 's/classes=80/classes=1/g' cfg/yolov3_traffic_signal.cfg
# + [markdown] id="C3WWoxign6C-"
# ### build darknet
# + colab={"background_save": true} id="RaleYYUMhjk1"
# change makefile to have GPU and OPENCV enabled
# !sed -i 's/OPENCV=0/OPENCV=1/' Makefile
# !sed -i 's/GPU=0/GPU=1/' Makefile
# !sed -i 's/CUDNN=0/CUDNN=1/' Makefile
# verify CUDA
# !/usr/local/cuda/bin/nvcc --version
# make darknet (build)
# !make
# + [markdown] id="RsLWD44Z89yi"
# # Download Pre-trained Weight
# + colab={"background_save": true} id="F1zxXZW19Eam"
# get yolov3 pretrained coco dataset weights
# !wget https://pjreddie.com/media/files/yolov3.weights
# + colab={"background_save": true} id="DJaw-vS3NTaM"
# upload pretrained convolutional layer weights
# !wget http://pjreddie.com/media/files/darknet53.conv.74
# + colab={"background_save": true} id="LQ_XRfYo55Jt"
# + colab={"background_save": true} id="0_hJu19t55GG"
# + colab={"background_save": true} id="U_jwrYf09JDL"
# define helper functions
def imShow(path):
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
image = cv2.imread(path)
height, width = image.shape[:2]
resized_image = cv2.resize(image,(3*width, 3*height), interpolation = cv2.INTER_CUBIC)
fig = plt.gcf()
fig.set_size_inches(18, 10)
plt.axis("off")
plt.imshow(cv2.cvtColor(resized_image, cv2.COLOR_BGR2RGB))
plt.show()
# use this to upload files
def upload():
from google.colab import files
uploaded = files.upload()
for name, data in uploaded.items():
with open(name, 'wb') as f:
f.write(data)
print ('saved file', name)
# use this to download a file
def download(path):
from google.colab import files
files.download(path)
# + [markdown] id="SrZK0LguB7cm"
# # training with my data
# + id="UtC-PuYoN8IS"
# train your custom detector
# !./darknet detector train data/obj.data cfg/yolov3_traffic_signal.cfg darknet53.conv.74 -dont_show
# + id="WBAs7CzFtNEb"
# + [markdown] id="XlHaKIW26bQB"
#
| Yolo_Trafiic_Sign_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
entrada_dados = input('Digite seu nome: '), int(input('Digite sua idade: ')), int(input('Informe seu salario: '))
nome = entrada_dados[0]
idade = entrada_dados[1]
salario = entrada_dados[2]
#print('Este é o '+ str(nome) + ' tem ' + str(idade) + ' e atualmente ganha: '+ str(salario) + 'R$')
print('Este é o ', nome, ' tem ', idade, 'anos e atualmente ganha: ', salario, 'R$')
# +
#Entrada de dados para notas
notas = (6.5, 7.8, 10.0, 9.1)
i = 0
total = 0
total_soma = sum(notas)
media = total_soma / len(notas)
#print("Temos", len(notas), "notas ao todo")
# -
#Testando quais notas são maores que a média e as abaixo da média depois da entrada de dados acima
while i < len(notas):
if notas[i] >= 7.0:
print(notas[i], "é igual ou maior que a média")
else:
print(notas[i], "é menor que a média")
i += 1
#Alternativa usando o FOR
for nota in notas:
if nota >= 7.0:
print(nota, "é igual ou maior que a média")
else:
print(nota, "é menor que a média")
#Calculando a Media pra saber se passou
for nota in notas:
total += nota
media = total / len(notas)
if media >= 7.0:
print("aprovado com média:", media)
else:
print("reprovado com média:", media)
#Agora Aproveitando as funções Total_Soma e Media
if media >= 7.0:
print("aprovado com média:", media)
else:
print("reprovado com média:", media)
animais = ('cobra', 'camaleão', 'Urubu', 'Jacaré')
print(animais[-2])
#animais = ('cobra', 'camaleão', 'Urubu', 'Jacaré')
valor_absoluto = abs(-1)
print(valor_absoluto)
#Entrada de Valres
valores = (5, 4, 7, 1)
maximo = 0
for valor in valores:
if valor > maximo:
maximo = valor
print(maximo)
| tuplasEmFuncao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Decoding sensor space data with generalization across time and conditions
#
#
# This example runs the analysis computed in:
#
# King & Dehaene (2014) 'Characterizing the dynamics of mental
# representations: the temporal generalization method', Trends In Cognitive
# Sciences, 18(4), 203-210.
# http://www.ncbi.nlm.nih.gov/pubmed/24593982
#
# The idea is to learn at one time instant and assess if the decoder
# can predict accurately over time and on a second set of conditions.
#
#
# +
# Authors: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import mne
from mne.datasets import sample
from mne.decoding import GeneralizationAcrossTime
print(__doc__)
# Preprocess data
data_path = sample.data_path()
# Load and filter data, set up epochs
raw_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw.fif'
events_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw-eve.fif'
raw = mne.io.read_raw_fif(raw_fname, preload=True)
picks = mne.pick_types(raw.info, meg=True, exclude='bads') # Pick MEG channels
raw.filter(1, 30, method='fft') # Band pass filtering signals
events = mne.read_events(events_fname)
event_id = {'AudL': 1, 'AudR': 2, 'VisL': 3, 'VisR': 4}
decim = 2 # decimate to make the example faster to run
epochs = mne.Epochs(raw, events, event_id, -0.050, 0.400, proj=True,
picks=picks, baseline=None, preload=True,
reject=dict(mag=5e-12), decim=decim, verbose=False)
# We will train the classifier on all left visual vs auditory trials
# and test on all right visual vs auditory trials.
# In this case, because the test data is independent from the train data,
# we test the classifier of each fold and average the respective predictions.
# Define events of interest
triggers = epochs.events[:, 2]
viz_vs_auditory = np.in1d(triggers, (1, 2)).astype(int)
gat = GeneralizationAcrossTime(predict_mode='mean-prediction', n_jobs=1)
# For our left events, which ones are visual?
viz_vs_auditory_l = (triggers[np.in1d(triggers, (1, 3))] == 3).astype(int)
# To make scikit-learn happy, we converted the bool array to integers
# in the same line. This results in an array of zeros and ones:
print("The unique classes' labels are: %s" % np.unique(viz_vs_auditory_l))
gat.fit(epochs[('AudL', 'VisL')], y=viz_vs_auditory_l)
# For our right events, which ones are visual?
viz_vs_auditory_r = (triggers[np.in1d(triggers, (2, 4))] == 4).astype(int)
gat.score(epochs[('AudR', 'VisR')], y=viz_vs_auditory_r)
gat.plot(title="Temporal Generalization (visual vs auditory): left to right")
| 0.13/_downloads/plot_decoding_time_generalization_conditions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## AdhereID
# #### Predicting medication adherence
# Welcome to AdhereID! This is a project I completed as part of the Insight Health Data Science Program. Since I coded the whole project .py files via PyCharm, the purpose of this notebook is to provide an overview of the project, code and associated files.
# #### Project Structure
# ```
# code/
# # Collecting and cleaning data
# n01_ncpa_import_clean_export.py
# n02_patientslikeme_webscraping.py
# n03_clean_patientslikeme.py
#
# # Models and model analysis
# n04_eval_models.py # Initial model evaluation
# n05_final_logreg.py # Final logistic regression model
# n05_final_rf.py # Random forest model (for comparison)
# n06_feature_importance.py
# n07_model_performance_plots.py
#
# data/
# raw/
# cleaned/
#
# documents/
# # NCPA dataset codebook
#
# notebooks/
# # Jupyter notebook with README information
# ```
# #### Data Sources
#
# The goal of this project was to develop a machine learning model to predict medication adherence. Medication non-adherence is a widespread problem in the healthcare industry. About 131 million Americans (half the population!) take at least one prescription medication, and 10~25% of these people don't take their prescription as prescribed.
#
# A tool to predict medication adherence could be used in the doctor's office or pharmacy to determine which patients are at risk for non-adherence; based on this prediction, the healthcare provider could take action to help the patient comply such as providing information on managing side effects, suggesting a pill reminder app, or prescribing a lower-cost or shorter-term treatment.
#
# While adherence prediction has been tackled by academia and large healthcare companies alike, typically adherence has been inferred from a combination of electronic health record data and insurance claims data. In this project, I decided to test whether publicly available demographic and patient-reported medical literacy data could be used in a similar way.
#
# I collected two datasets to use for this project:
# #### 1. NCPA (National Community Pharmacists Association) : Medication Adherence Survey (2013)
# Phone survey dataset of 1,020 adults in the U.S. See documents/ for the dataset's codebook containing the full list of survey questions.
# - Downloaded from: https://ropercenter.cornell.edu/CFIDE/cf/action/catalog/abstract.cfm?type=&start=&id=&archno=USMISC2013-NCPA&abstract (Accessed 1-14-2020)
# - Data format: SPSS
# - 1020 rows x 158 columns
# #### 2. PatientsLikeMe
# An online community for discussing medical conditions and medications.
# - Web scraped using beautifulsoup (Accessed 1-20-2020)
# - Data format: data extracted from HTML
# - 10,025 rows of drug reviews (adherence, burden(difficulty of taking) and cost) x 11 columns
#
| notebooks/AdhereID_project_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Notebook for constructiong POD-NODE NIROM approximation for a flow around a cylinder example
#
# A collection of high-fidelity snapshots are generated that sufficiently capture the time-dynamics of the simulation. POD is adopted to define a reduced basis space for the high-fidelity snaphosts. The evolution of the time dynamics in the POD-latent space is modeled using Neural ODEs (NODE).
#
# OpenFOAM is used as the high-fidelity model for simulating flow around a cylinder governed by incompressible 2D Navier Stokes.
#
# #### Note
# This notebook serves as an example of how to set up and evaluate a PODNODE model for the given dataset. However, in order to attain a desirable level of prediction accuracy, the training time is high. Please refer to
# ```
# <NAME>, <NAME>, and <NAME>, “Neural Ordinary Differential Equations for Data-Driven Reduced Order Modeling of Environmental Hydrodynamics,” in Proceedings of the AAAI 2021 Spring Symposium on Combining Artificial Intelligence and Machine Learning with Physical Sciences, 2021.
# arXiv:2104.13962 [cs.LG]
# ```
# for model configuration details.
# +
### Loading modules
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import time
from sklearn.preprocessing import MinMaxScaler, StandardScaler
import scipy
import os
import gc
import argparse
import platform
print("Python "+str(platform.python_version()))
import importlib
from importlib import reload as reload
import tensorflow as tf
print("Tensorflow "+ str(tf.__version__))
if tf.__version__ == '1.15.0':
tf.compat.v1.enable_eager_execution()
elif tf.__version__.split('.')[0] == 2:
print("Setting Keras backend datatype")
tf.keras.backend.set_floatx('float64')
from tfdiffeq import odeint,odeint_adjoint
from tfdiffeq.adjoint import odeint as adjoint_odeint
tf.keras.backend.set_floatx('float64')
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
np.random.seed(0)
basedir = os.getcwd()
srcdir = os.path.join(basedir,'../pynirom/')
workdir = os.path.join(basedir,'../examples/')
datadir = os.path.join(basedir,'../data/')
figdir = os.path.join(basedir,'../figures/podnode')
nodedir = os.path.join(basedir,'../data/')
savedir = nodedir+'cylinder/current'
import pynirom
from pynirom.pod import pod_utils as pod
from pynirom.utils import data_utils as du
from pynirom.node import main as nd
from pynirom.node import plotting as pu
from pynirom.node import node as node
# +
device = 'cpu:0' # select gpu:# or cpu:#
purpose= 'train' #'train' to train a new model,
# 'retrain' to start training from an existing model, and
# 'eval' to load a pre-trained model for evaluation
pre_trained_dir = savedir+'/model_weights/' #If 'eval' specify path for pretrained model
stacking = True #If True, Specify new stacking order of latent space vector
stack_order = 'v_x,v_y,p' #If stacking = True, specify the stacking order of the latent space vector
scale_time = False #Scale time or not (Normalize)
scale_states = True #Scale states or not
scaling_method = 'centered'#Scaling method: 'centered', 'minmax' or 'maxabs'
augmented,aug_dims = (False,5) #Augmented or not and size of augmentation
N_layers = int(1) #Only four layers supported as of now.
N_neurons = int(256) #Number of neurons per layer
act_f = 'tanh' #Activation Function ('linear', 'tanh', 'sigmoid',...), default='tanh'
learning_rate_decay = True #Use decaying learning rate or not
initial_learning_rate = float(0.001) #If 'learning_rate_decay = False' then this is the fixed learning rate
decay_steps = int(5001) #Number of steps for learning rate decay
decay_rate = float(0.5) #Rate of learning rate decay
staircase_opt = True #True for staircase decay and False for exponential
optimizer = 'RMSprop' #See pynirom.node.node.set_optimizer() for options
use_adjoint = False #Use adjoint method or not
solver = 'rk4' #Specify ODE solver. See tfdiffeq README for available options
use_minibatch, batch_size = (False,256) #Use minibatch or not and batch size
epochs = int(500) #Number of epochs of training
print("\n***** Runtime parameters: ******\n")
print(f'Mode = {purpose}, Scaling = {scale_states}, Augmenting = {augmented}, Adjoint = {use_adjoint}')
print(f'Solver = {solver}, Optimizer = {optimizer}, Stacking order = {stack_order}, Epochs = {epochs}')
print(f'# Layers = {N_layers}, # Neurons per layer = {N_neurons}, Activation fn = {act_f}')
if use_minibatch:
print(f'Use minibatch = {use_minibatch}, Batch size = {batch_size}')
if learning_rate_decay:
print(f'Init LR = {initial_learning_rate}, # LR decay steps = {decay_steps}, LR decay rate = {decay_rate}')
else:
print(f'Fixed LR = {initial_learning_rate}')
print('**********************************\n')
# +
### ------ Import Snapshot data -------------------
data = np.load(datadir + 'cylinder_Re100.0_Nn14605_Nt3001.npz')
mesh = np.load(datadir + 'OF_cylinder_mesh_Nn14605_Ne28624.npz')
print('HFM data has {0} snapshots of dimension {1} for p,u and v, spanning times [{2}, {3}]'.format(
data['time'].shape[0],data['p'].shape[0],
data['time'][0], data['time'][-1]))
## ------- Prepare training snapshots ----------------
print('\n-------Prepare training and testing data---------')
soln_names = ['p', 'v_x', 'v_y']
nodes = mesh['nodes']; node_ind = mesh['node_ind']
triangles = mesh['elems']; elem_ind = mesh['elem_ind']
snap_start = 1250
T_end = 5.0 ### 5 seconds
snap_incr = 4
snap_train, times_train = du.prepare_data(data, soln_names, start_skip=snap_start, T_end=T_end, incr=snap_incr)
print('Using {0} training snapshots for time interval [{1},{2}] seconds'.format(times_train.shape[0],
times_train[0], times_train[-1]))
## ------- Prepare testing snapshots ----------------
pred_incr = snap_incr -3
snap_pred_true, times_predict = du.prepare_data(data, soln_names, start_skip=snap_start, incr=pred_incr)
print('Using {0} testing snapshots for time interval [{1},{2}] seconds'.format(times_predict.shape[0],
times_predict[0], times_predict[-1]))
del data
del mesh
gc.collect()
# +
### ------ Compute the POD basis using the training snapshots------------------
trunc_lvl = 0.99
snap_norm, snap_mean, U, D, W = pod.compute_pod_multicomponent(snap_train)
nw, U_r = pod.compute_trunc_basis(D, U, eng_cap = trunc_lvl)
### ------ Compute the POD coefficients for training snapshots------------------
Z_train = pod.project_onto_basis(snap_train, U_r, snap_mean)
### ------ Compute the POD coefficients for the truth snapshots on the prediction interval------------------
Z_pred_true = pod.project_onto_basis(snap_pred_true, U_r, snap_mean)
npod_total = 0
for key in soln_names:
npod_total+=nw[key]
# +
### ---- Setup NODE input data
NODE = nd.NODEBase(device=device)
true_state_array, true_pred_state_array, init_state, state_len, dt_train, dt_predict = \
NODE.prepare_input_data(Z_train, nw, times_train, stack_order, times_predict, Z_pred_true)
print("Training NODE using %d modes for %d time steps with %.3f <= t <= %.3f and dt = %.4f"%(state_len,
true_state_array.shape[0], times_train[0],
times_train[-1], dt_train))
print("Predicting NODE solutions using %d modes for %d time steps with %.3f <= t <= %.3f and dt = %.4f"%(
state_len, true_pred_state_array.shape[0], times_predict[0],
times_predict[-1], dt_predict))
# +
### Preprocess training data (scale time and/or states, augment states if using ANODE)
### Set up learning rate scheduler and optimizer for training of the NODE model
true_state_tensor, times_tensor, init_tensor, learn_rate, optim = \
NODE.preprocess_data(scale_states=scale_states, scale_time=scale_time, augmented=augmented,
lr_decay=learning_rate_decay, init_lr=initial_learning_rate, opt=optimizer,
scaling_method=scaling_method, aug_dim=aug_dims,
decay_steps=decay_steps, decay_rate=decay_rate, staircase=staircase_opt, )
# +
### ---- Model Training ------
train_loss_results, train_lr, saved_ep = \
NODE.train_model(true_state_tensor, times_tensor, init_tensor, epochs, savedir,
solver=solver, purpose=purpose, adjoint=use_adjoint, minibatch=use_minibatch,
pre_trained_dir = pre_trained_dir)
# +
## --- Generate NODE predictions ---
predicted_states, times_predict = NODE.predict_time(times_predict, init_tensor, pre_trained_dir,)
## ---- Compute Mean Square Error of predictions
Z_pred = {}
ctr= 0
for key in stack_order.split(','):
Z_pred[key] = np.array(predicted_states)[:,ctr:ctr+nw[key]].T
ctr += nw[key]
snap_pred = pod.reconstruct_from_rom(Z_pred, U_r, snap_mean, nw)
error_p = np.mean(np.square(snap_pred['p']-snap_pred_true['p']))
error_vx = np.mean(np.square(snap_pred['v_x']-snap_pred_true['v_x']))
error_vy = np.mean(np.square(snap_pred['v_y']-snap_pred_true['v_y']))
print("\n---- Mean Square Error of NODE predictions ----\n")
print('Pr MSE: ' + str(error_p))
print('Vx MSE: ' + str(error_vx))
print('Vy MSE: ' + str(error_vy))
# -
def set_label(key):
if key == 'v_x':
return 'u'
elif key == 'v_y':
return 'v'
else:
return key
# +
### ----- Visualize true and predicted POD coefficients -------
comp = 0
# Visualization fluff here
fig, ax = plt.subplots(nrows=3,ncols=1,figsize=(8,15))
mnum = comp
for i, key in enumerate(soln_names):
tt = ax[i].plot(times_predict[:],true_pred_state_array[:,mnum],label='True',marker='o',markevery=20)
# Visualization of modal evolution using NODE
ln, = ax[i].plot(times_predict[:],predicted_states[:,mnum],label='NODE',color='orange',marker='D',
markevery=25)
mnum = mnum + nw[key]
ax[i].set_xlabel('Time', fontsize=18)
sv = set_label(key)+', mode '+str(comp)
ax[i].set_ylabel(sv,fontsize=18)
ax[i].legend(fontsize=14)
fig.suptitle("POD coefficients of the HFM and NODE solutions", fontsize=20)
fig.tight_layout(rect=[0, 0.03, 1, 0.98])
# +
## ---- Compute spatial RMS/Relative error
reload(nd)
reload(pu)
metric = 'rms'
err = NODE.compute_error(snap_pred_true, snap_pred, soln_names, metric=metric)
vstring = {}
for key in soln_names:
vstring[key] = set_label(key)
## ---- Visualize computed error metric
pu.plot_NODE_err(err, times_predict, soln_names, vstring, metric=metric)
# +
#### ----- Save predicted solutions -------
save_nirom_solutions = False
if save_nirom_solutions:
os.chdir(nodedir)
print("Saving results in %s"%(os.getcwd()))
np.savez_compressed('cylinder_online_node',
p=snap_pred['p'],v_x=snap_pred['v_x'], v_y=snap_pred['v_y'],
time=times_predict,loss=train_loss_results)
| examples/PODNODE_cylinder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 時尚服飾圖像辨識
# 圖像分類被廣泛地應用到很多的領域中,從醫學掃描中危及生命的疾病識別到電視卡通角色的識別。
#
# MNIST數據集(可以說)是最常用的圖像分類入門數據集。它包含手寫數字的28x28灰度圖像,每個灰度圖像都有一個關聯的標籤,用於指示圖像上手寫的數字(0到9之間的整數)。
#
# MNIST在過去幾年中越來越受歡迎, 但也引起了一些問題。
#
# 使用MNIST來進行圖像辨識的範例有什麼問題呢?
# 1. **MNIST太簡單了** - 卷積網絡(Convolution network)可以很容易地在MNIST上達到99.7%的辨識力,即使經典的機械學習(Machine learning)算法也能達到97%。
#
# 2. **MNIST被過度使用** - 幾乎每個有深度學習經驗的人都至少遇到過MNIST一次。
#
# 3. **NIST無法有效代表現代的電視視覺相關的任務** - Keras的主要貢獻者(<NAME>)也特別提及這一點。
#
# >**<NAME>**
# >
# >Many good ideas will not work well on MNIST (e.g. batch norm). Inversely many bad ideas may work on MNIST and no transfer to real CV.
# >
# > 2:51 AM - Apr 14, 2017
#
#
# 
# ## 資料集說明
#
# 時尚MNIST訓練集是由[Zalando Reserach](https://jobs.zalando.com/tech/)所發佈出來的資料集。它包含60,000個範例圖像,測試集包含10,000個範例圖像。
#
# 每個範例圖像都是28x28的灰度圖像(就像原始MNIST中的圖像一樣),與10類(T卹,褲子,套頭衫,連衣裙,大衣,涼鞋,襯衫,運動鞋,包包和踝靴)的標籤相關聯。時尚MNIST也與MNIST具有相同的訓練與測試數據分離結構,以便於學習使用。
#
# 相關的詳細資訊說明及資料集下載可以透過Zalando在Github: [https://github.com/zalandoresearch/fashion-mnist](https://github.com/zalandoresearch/fashion-mnist) 來了解更多。
#
# 在這篇文章裡頭, 我們會從[Kaggle](https://www.kaggle.com/zalando-research/fashionmnist)上取得這個數據集。
# ## 資料準備
#
# 1. 從[Kaggle](https://www.kaggle.com/zalando-research/fashionmnist/data/)點擊`Download All`下載圖像資料檔"fashionmnist.zip"。
# 2. 在這個Jupyter Notebook所在的目錄下產生一個新的子目錄"data"。
# 3. 解壓縮圖像資料檔到"data"的目錄裡頭。
# 4. 在"data/"的資料夾會看到兩個檔案
# * `fashion-mnist_test.csv`
# * `fashion-mnist_train.csv`
#
# 最後你的目錄結構看起來像這樣:
#
# 2.3-fashion-mnist-recognition.ipynb
# data/
# ├── fashion-mnist_test.csv
# └── fashion-mnist_train.csv
# ## 資料預處理 (Data Preprocessing)
# 預處理圖片的第一步是調整它們的大小。我們需要有相同大小的所有照片進行訓練。
# 我會將數據資料轉換型別為`float32`來節省一些記憶的用量並對它們進行歸一化(除以255)。
#
# 然後使用one-hot編碼來將10類別的標籤(label)轉換為向量(vector):
# +
from keras.utils import to_categorical
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
# 載入資料
data_train = pd.read_csv('data/fashion-mnist_train.csv')
data_test = pd.read_csv('data/fashion-mnist_test.csv')
num_classes = 10 # 標籤總共有10類
img_rows, img_cols, img_channels = 28, 28, 1 # 圖像是 28像素 x 28像素 (灰階: 1)
input_shape = (img_rows, img_cols, img_channels) # (圖像的height, 圖像的width, 圖像的顏色通道數channel)
X = np.array(data_train.iloc[:, 1:]) # Dataframe中 idx(1 ~ 784)的欄都是像素值
# 進行標籤的one-hot編碼
y = to_categorical(np.array(data_train.iloc[:, 0])) # Dataframe 中 idx(0)的欄是標籤
# 把訓練資料進行拆分成訓練(80%)與驗證(20%)資料集
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=13)
# 測試資料的處理
X_test = np.array(data_test.iloc[:, 1:])
y_test = to_categorical(np.array(data_test.iloc[:, 0]))
# 對向量進行shape的轉換以符合訓練的input要求
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols, 1)
X_test = X_test.reshape(X_test.shape[0], img_rows, img_cols, 1)
X_val = X_val.reshape(X_val.shape[0], img_rows, img_cols, 1)
# 對每個像素進行型別轉換與歸一化
X_train = X_train.astype('float32')/255
X_test = X_test.astype('float32')/255
X_val = X_val.astype('float32')/255
# -
# ## 網絡模型 (Model)構建
#
# 現在我們來定義我們的模型架構。我們將使用具有6個卷積層的前饋(feed-forward)網絡,然後是完全連接的隱藏層。
# 我們也將在兩者之間使用Dropout層來防止網絡"過擬合(overfitting)"。
# +
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
def create_model_six_conv(input_shape):
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same', activation='relu', kernel_initializer='he_normal', input_shape=input_shape))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.2))
model.add(Conv2D(64, (3, 3), padding='same', activation='relu'))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.2))
model.add(Conv2D(128, (3, 3), padding='same', activation='relu'))
model.add(Conv2D(128, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
return model;
# -
# 在訓練模型之前,我們需要將模型配置為學習算法並進行編譯。我們需要指定: loss: 損失函數,我們要優化。我們不能使用MSE,因為它是不連續的數值。因此,我們使用:categorical_crossentropy optimizer: 我們使用標準隨機梯度下降(Stochastic gradient descent)與涅斯捷羅夫動量(Nesterov momentum) metric: 由於我們正在處理一個分類問題,我們用度量是accuracy。
# +
from keras.optimizers import Adam
batch_size = 256
epochs = 50
#圖像的shape是 (28,28,1)
model = create_model_six_conv((img_rows, img_cols, img_channels)) # 初始化一個模型
# 秀出模型架構
model.summary()
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
# -
# ## 訓練 (Training)
#
# 現在,我們的模型已經準備好了。在訓練期間,我們的模型將進行迭代批量訓練,每個次的訓練資料的大小為batch_size。對於每批次,模型將會計算出梯度(gradient),並自動更新網絡的權重。對所有訓練集的一次迭代被稱為一次的循環(epoch)。訓練通常會一直進行到損失收斂於一個常數。
history = model.fit(X_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(X_val, y_val))
# ### 訓練過程的可視化
# +
# 透過趨勢圖來觀察訓練與驗證的走向 (特別去觀察是否有"過擬合(overfitting)"的現象)
import matplotlib.pyplot as plt
def plot_train_history(history, train_metrics, val_metrics):
plt.plot(history.history.get(train_metrics),'-o')
plt.plot(history.history.get(val_metrics),'-o')
plt.ylabel(train_metrics)
plt.xlabel('Epochs')
plt.legend(['train', 'validation'])
plt.figure(figsize=(12,4))
plt.subplot(1,2,1)
plot_train_history(history, 'loss','val_loss')
plt.subplot(1,2,2)
plot_train_history(history, 'acc','val_acc')
plt.show()
# -
# 分析一下訓練過程, 我們可以很明顯看到大約在第10個epochs之後整個模型就開始在驗證資料集有過擬合(overfitting)的現象。
# 如果要繼續調優的話, 可以朝向增加資料集或進行一些降維的處理。
# ## 驗證評估 (Evaluation)
# +
score = model.evaluate(X_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# ### 93.8%的正確率 ~~ 太神奇了, 傑克!!
#
# 事實證明,卷積神經網絡分類器比在[這里](http://fashion-mnist.s3-website.eu-central-1.amazonaws.com/)報告的最好的分類器(SVM) 89.7%還要更好。
# ### 每一種類別的預測正確率
# +
#get the predictions for the test data
predicted_classes = model.predict_classes(X_test)
#get the indices to be plotted
y_true = data_test.iloc[:, 0]
corrects = np.nonzero(predicted_classes==y_true)[0]
incorrects = np.nonzero(predicted_classes!=y_true)[0]
# +
from sklearn.metrics import classification_report
target_names = ["Class {}".format(i) for i in range(num_classes)]
print(classification_report(y_true, predicted_classes, target_names=target_names))
# -
# 顯然,我們的分類器對第6類別的圖像辨識在精度和召回方面的表現相對差。也許我們會在可視化正確和錯誤的預測之後獲得更多的洞察力。
# #### 正確辨識的圖像範例:
# +
for i, correct in enumerate(corrects[:9]):
plt.subplot(3,3,i+1)
plt.imshow(X_test[correct].reshape(28,28), cmap='gray', interpolation='none')
plt.title("Predicted {}, Class {}".format(predicted_classes[correct], y_true[correct]))
plt.tight_layout()
plt.show()
# -
# #### 辨識錯誤的圖像範例
# +
for i, incorrect in enumerate(incorrects[0:9]):
plt.subplot(3,3,i+1)
plt.imshow(X_test[incorrect].reshape(28,28), cmap='gray', interpolation='none')
plt.title("Predicted {}, Class {}".format(predicted_classes[incorrect], y_true[incorrect]))
plt.tight_layout()
plt.show()
# -
# ## 總結 (Conclusion)
#
# 在這篇文章中有一些個人學習到的一些有趣的重點:
# * 深度學習也可以很摩登很現代感
# * 只要有創造力與天馬行空的想法,很多我們眼睛看到的東西都可以變成應用AI的場景
# * 每個公司都可以擁有自己的訓練圖像資料集
# * 電腦宅男也可以跟時尚(fashion)掛上勾了, 下次我們可以說我們是"AI時尚宅男" ...太潮了!!
# ### 參考:
# * [CNN with Keras](https://www.kaggle.com/bugraokcu/cnn-with-keras)
# * [zalandoresearch/fashion-mnist](https://github.com/zalandoresearch/fashion-mnist)
# * [Classifying clothes using Tensorflow](https://medium.com/tensorist/classifying-fashion-articles-using-tensorflow-fashion-mnist-f22e8a04728a)
# * [Keras官網](http://keras.io/)
| deep-learning-with-keras-notebooks-master/2.3-fashion-mnist-recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PSM Net Baseline
#
# *See for reference: https://github.com/JiaRenChang/PSMNet*
# ## 1 Setup
#
# ----
#
# Ensure you're in <...>/argoverse-api
import os
os.chdir('..')
print(f"Current directory: {os.getcwd()}")
# +
import cv2
import numpy as np
from tqdm import tqdm
from pathlib import Path
from argoverse.data_loading.stereo_dataloader import ArgoverseStereoDataLoader
from argoverse.evaluation.stereo.eval import StereoEvaluator
from argoverse.utils.calibration import get_calibration_config
from argoverse.utils.camera_stats import RECTIFIED_STEREO_CAMERA_LIST
import time
from models import *
from dataloader import KITTI_submission_loader as DA
from PIL import Image
from pytorch_modelsize import SizeEstimator
import torch
import torch.nn as nn
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.optim as optim
import torch.utils.data
from torch.autograd import Variable
import torch.nn.functional as F
import torchvision.transforms as transforms
STEREO_FRONT_LEFT_RECT = RECTIFIED_STEREO_CAMERA_LIST[0]
STEREO_FRONT_RIGHT_RECT = RECTIFIED_STEREO_CAMERA_LIST[1]
# -
main_dir = "C:/Michael/10707/argoverse-api/"
data_dir = f"{main_dir}argoverse-stereo_v1.1/"
# ## 2 Main Model
#
# ----
#
# Goal: Predict disparity map from pair of stereo images
#
# ### 2.1 Training
# Should create/tune a PSM net and save it as "model"
# I'd say, given the time constraints, it's probably best to take one of the prebuilt ones from their
# Github page and just tune it on the Argo data? If you want to try something else though go for it.
# I think their code is input size invariant because the only operations they do are colvolution and
# spatial pyramid pooling, neither of which should have any sizes hardcoded.
maxdisp = 192
loadmodel = "./checkpoint_9.tar"
model = basic(maxdisp)
state_dict = torch.load(loadmodel, map_location='cpu')
state_dict = {(k if 'module' not in k else k[7:]): v for k, v in state_dict['state_dict'].items()}
model.load_state_dict(state_dict)
# ### 2.2 Evaluation
#
# * Probably should add checkpoint code depending on how long this takes to run (e.g. save metrics after every iteration).
def test(imgL,imgR):
model.eval()
#if args.cuda:
#imgL = imgL.cuda()
#imgR = imgR.cuda()
with torch.no_grad():
output = model(imgL,imgR)
output = torch.squeeze(output).data.cpu().numpy()
return output
# +
stereo_data_loader = ArgoverseStereoDataLoader(data_dir, "val")
metrics = []
lens = []
log_ids = [
'f9fa3960-537f-3151-a1a3-37a9c0d6d7f7',
'1d676737-4110-3f7e-bec0-0c90f74c248f',
'da734d26-8229-383f-b685-8086e58d1e05',
'6db21fda-80cd-3f85-b4a7-0aadeb14724d',
'85bc130b-97ae-37fb-a129-4fc07c80cca7',
'33737504-3373-3373-3373-633738571776',
'033669d3-3d6b-3d3d-bd93-7985d86653ea',
'f1008c18-e76e-3c24-adcc-da9858fac145',
'5ab2697b-6e3e-3454-a36a-aba2c6f27818',
'cb762bb1-7ce1-3ba5-b53d-13c159b532c8',
'70d2aea5-dbeb-333d-b21e-76a7f2f1ba1c',
'2d12da1d-5238-3870-bfbc-b281d5e8c1a1',
'64724064-6472-6472-6472-764725145600',
'00c561b9-2057-358d-82c6-5b06d76cebcf',
'cb0cba51-dfaf-34e9-a0c2-d931404c3dd8',
'e9a96218-365b-3ecd-a800-ed2c4c306c78',
'39556000-3955-3955-3955-039557148672'
]
i = 0
for log_id in log_ids:
i += 1
print(f"Now evaluating log_id :: \t\t {i}/{len(log_ids)}")
left_stereo_img_fpaths = stereo_data_loader.get_ordered_log_stereo_image_fpaths(
log_id=log_id,
camera_name=STEREO_FRONT_LEFT_RECT)
right_stereo_img_fpaths = stereo_data_loader.get_ordered_log_stereo_image_fpaths(
log_id=log_id,
camera_name=STEREO_FRONT_RIGHT_RECT)
disparity_map_fpaths = stereo_data_loader.get_ordered_log_disparity_map_fpaths(
log_id=log_id,
disparity_name="stereo_front_left_rect_disparity")
disparity_obj_map_fpaths = stereo_data_loader.get_ordered_log_disparity_map_fpaths(
log_id=log_id,
disparity_name="stereo_front_left_rect_objects_disparity")
lens += [len(left_stereo_img_fpaths)]
normal_mean_var = {'mean': [0.485, 0.456, 0.406],
'std': [0.229, 0.224, 0.225]}
infer_transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize(**normal_mean_var)])
for inx in tqdm(range(len(left_stereo_img_fpaths))):
imgL_o = Image.open(left_stereo_img_fpaths[inx]).convert('RGB')
imgR_o = Image.open(right_stereo_img_fpaths[inx]).convert('RGB')
imgL = infer_transform(imgL_o)
imgR = infer_transform(imgR_o)
# pad to width and hight to 16 times
if imgL.shape[1] % 16 != 0:
times = imgL.shape[1]//16
top_pad = (times+1)*16 -imgL.shape[1]
else:
top_pad = 0
if imgL.shape[2] % 16 != 0:
times = imgL.shape[2]//16
right_pad = (times+1)*16-imgL.shape[2]
else:
right_pad = 0
imgL = F.pad(imgL,(0,right_pad, top_pad,0)).unsqueeze(0)
imgR = F.pad(imgR,(0,right_pad, top_pad,0)).unsqueeze(0)
start_time = time.time()
pred_disp = test(imgL,imgR)
print('time = %.2f' %(time.time() - start_time))
if top_pad !=0:
if right_pad != 0:
img = pred_disp[top_pad:,:-right_pad]
else:
img = pred_disp[top_pad:, :]
else:
if right_pad != 0:
img = pred_disp[:, :-right_pad]
else:
img = pred_disp
print(img.shape)
# Load the testing image and corresponding disparity and foreground disparity maps
#stereo_front_left_rect_image = stereo_data_loader.get_rectified_stereo_image(left_stereo_img_fpaths[idx])
#stereo_front_right_rect_image = stereo_data_loader.get_rectified_stereo_image(right_stereo_img_fpaths[idx])
stereo_front_left_rect_disparity = stereo_data_loader.get_disparity_map(disparity_map_fpaths[inx])
stereo_front_left_rect_objects_disparity = stereo_data_loader.get_disparity_map(disparity_obj_map_fpaths[inx])
#left_disparity_pred = np.uint16(left_disparity)
left_disparity_pred = (img*256).astype('uint16')
img = Image.fromarray(left_disparity_pred)
#print(left_disparity_pred)
timestamp = int(Path(disparity_map_fpaths[inx]).stem.split("_")[-1])
save_dir_disp = f"{main_dir}707-files/results/psm/stereo_output/{log_id}"
Path(save_dir_disp).mkdir(parents=True, exist_ok=True)
filename = f"{save_dir_disp}/disparity_{timestamp}.png"
img.save(filename)
#cv2.imshow("image", left_disparity_pred)
#if not cv2.imwrite(filename, left_disparity_pred):
#raise Exception("Could not write image to " +filename)
pred_dir = Path(save_dir_disp)
gt_dir = Path(f"{data_dir}/disparity_maps_v1.1/val/{log_id}")
save_figures_dir = Path(f"/tmp/results/psm/figures/{log_id}/")
save_figures_dir.mkdir(parents=True, exist_ok=True)
evaluator = StereoEvaluator(
pred_dir,
gt_dir,
save_figures_dir,
)
metrics += [evaluator.evaluate()]
# +
compiled_metrics = { key : 0 for key in metrics[0] }
for i in range(0, len(metrics)):
compiled_metrics = { key : compiled_metrics[key] + lens[i] * metrics[i][key] for key in compiled_metrics }
compiled_metrics = { key : compiled_metrics[key] / sum(lens) for key in compiled_metrics }
# -
import json
print(f"{json.dumps(compiled_metrics, sort_keys=False, indent=4)}")
| 707-files/psm-net-basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:detect] *
# language: python
# name: conda-env-detect-py
# ---
# [source](../api/alibi_detect.cd.ks.rst)
# # Kolmogorov-Smirnov
#
# ## Overview
#
# The drift detector applies feature-wise two-sample [Kolmogorov-Smirnov](https://en.wikipedia.org/wiki/Kolmogorov%E2%80%93Smirnov_test) (K-S) tests. For multivariate data, the obtained p-values for each feature are aggregated either via the [Bonferroni](https://mathworld.wolfram.com/BonferroniCorrection.html) or the [False Discovery Rate](http://www.math.tau.ac.il/~ybenja/MyPapers/benjamini_hochberg1995.pdf) (FDR) correction. The Bonferroni correction is more conservative and controls for the probability of at least one false positive. The FDR correction on the other hand allows for an expected fraction of false positives to occur.
#
# For high-dimensional data, we typically want to reduce the dimensionality before computing the feature-wise univariate K-S tests and aggregating those via the chosen correction method. Following suggestions in [Failing Loudly: An Empirical Study of Methods for Detecting Dataset Shift](https://arxiv.org/abs/1810.11953), we incorporate Untrained AutoEncoders (UAE), black-box shift detection using the classifier's softmax outputs ([BBSDs](https://arxiv.org/abs/1802.03916)) and [PCA](https://en.wikipedia.org/wiki/Principal_component_analysis) as out-of-the box preprocessing methods. Preprocessing methods which do not rely on the classifier will usually pick up drift in the input data, while BBSDs focuses on label shift. The [adversarial detector](https://arxiv.org/abs/2002.09364) which is part of the library can also be transformed into a drift detector picking up drift that reduces the performance of the classification model. We can therefore combine different preprocessing techniques to figure out if there is drift which hurts the model performance, and whether this drift can be classified as input drift or label shift.
#
# Detecting input data drift (covariate shift) $\Delta p(x)$ for text data requires a custom preprocessing step. We can pick up changes in the semantics of the input by extracting (contextual) embeddings and detect drift on those. Strictly speaking we are not detecting $\Delta p(x)$ anymore since the whole training procedure (objective function, training data etc) for the (pre)trained embeddings has an impact on the embeddings we extract. The library contains functionality to leverage pre-trained embeddings from [HuggingFace's transformer package](https://github.com/huggingface/transformers) but also allows you to easily use your own embeddings of choice. Both options are illustrated with examples in the [Text drift detection on IMDB movie reviews](../examples/cd_text_imdb.nblink) notebook.
# ## Usage
#
# ### Initialize
#
#
# Parameters:
#
# * `p_val`: p-value used for significance of the K-S test for each feature. If the FDR correction method is used, this corresponds to the acceptable q-value.
#
# * `X_ref`: Data used as reference distribution.
#
# * `preprocess_X_ref`: Whether to already apply the (optional) preprocessing step to the reference data at initialization and store the preprocessed data. Dependent on the preprocessing step, this can reduce the computation time for the predict step significantly, especially when the reference dataset is large. Defaults to *True*.
#
# * `update_X_ref`: Reference data can optionally be updated to the last N instances seen by the detector or via [reservoir sampling](https://en.wikipedia.org/wiki/Reservoir_sampling) with size N. For the former, the parameter equals *{'last': N}* while for reservoir sampling *{'reservoir_sampling': N}* is passed.
#
# * `preprocess_fn`: Function to preprocess the data before computing the data drift metrics. Typically a dimensionality reduction technique.
#
# * `preprocess_kwargs`: Keyword arguments for `preprocess_fn`. Again see the notebooks for [image](../examples/cd_ks_cifar10.nblink) and [text](../examples/cd_text_imdb.nblink) data for concrete, detailed examples. The built-in *UAE*, *BBSDs* or text-specific preprocessing steps are passed here as well. See below for a brief example.
#
# * `correction`: Correction type for multivariate data. Either *'bonferroni'* or *'fdr'* (False Discovery Rate).
#
# * `alternative`: Defines the alternative hypothesis. Options are *'two-sided'* (default), *'less'* or *'greater'*.
#
# * `n_features`: Number of features used in the K-S test. No need to pass it if no preprocessing takes place. In case of a preprocessing step, this can also be inferred automatically but could be more expensive to compute.
#
# * `n_infer`: If the number of features need to be inferred after the preprocessing step, we can specify the number of instances used to infer the number of features from since this can depend on the specific preprocessing step.
#
# * `data_type`: can specify data type added to metadata. E.g. *'tabular'* or *'image'*.
#
# Initialized drift detector example:
#
# ```python
# from alibi_detect.cd import KSDrift
# from alibi_detect.cd.preprocess import UAE # Untrained AutoEncoder
#
# encoder_net = tf.keras.Sequential(
# [
# InputLayer(input_shape=(32, 32, 3)),
# Conv2D(64, 4, strides=2, padding='same', activation=tf.nn.relu),
# Conv2D(128, 4, strides=2, padding='same', activation=tf.nn.relu),
# Conv2D(512, 4, strides=2, padding='same', activation=tf.nn.relu),
# Flatten(),
# Dense(32,)
# ]
# )
# uae = UAE(encoder_net=encoder_net)
#
# cd = KSDrift(
# p_val=0.05,
# X_ref=X_ref,
# preprocess_X_ref=True,
# preprocess_kwargs={'model': uae, 'batch_size': 128},
# alternative='two-sided',
# correction='bonferroni'
# )
# ```
# ### Detect Drift
#
# We detect data drift by simply calling `predict` on a batch of instances `X`. We can return the feature-wise p-values before the multivariate correction by setting `return_p_val` to *True*. The drift can also be detected at the feature level by setting `drift_type` to *'feature'*. No multivariate correction will take place since we return the output of *n_features* univariate tests. For drift detection on all the features combined with the correction, use *'batch'*. `return_p_val` equal to *True* will also return the threshold used by the detector (either for the univariate case or after the multivariate correction).
#
# The prediction takes the form of a dictionary with `meta` and `data` keys. `meta` contains the detector's metadata while `data` is also a dictionary which contains the actual predictions stored in the following keys:
#
# * `is_drift`: 1 if the sample tested has drifted from the reference data and 0 otherwise.
#
# * `p_val`: contains feature-level p-values if `return_p_val` equals *True*.
#
# * `threshold`: for feature-level drift detection the threshold equals the p-value used for the significance of the K-S test. Otherwise the threshold after the multivariate correction (either *bonferroni* or *fdr*) is returned.
#
# * `distance`: feature-wise K-S statistics between the reference data and the new batch if `return_distance` equals *True*.
#
#
# ```python
# preds_drift = cd.predict(X, drift_type='batch', return_p_val=True, return_distance=True)
# ```
# ### Saving and loading
#
# The drift detectors can be saved and loaded in the same way as other detectors when using the built-in preprocessing steps (`alibi_detect.cd.preprocess.UAE` and `alibi_detect.cd.preprocess.HiddenOutput`) or no preprocessing at all:
#
# ```python
# from alibi_detect.utils.saving import save_detector, load_detector
#
# filepath = 'my_path'
# save_detector(cd, filepath)
# cd = load_detector(filepath)
# ```
#
# A custom preprocessing step can be passed as follows:
#
# ```python
# cd = load_detector(filepath, **{'preprocess_kwargs': preprocess_kwargs})
# ```
# ## Examples
#
# ### Image
#
# [Drift detection on CIFAR10](../examples/cd_ks_cifar10.nblink)
#
#
# ### Text
#
# [Text drift detection on IMDB movie reviews](../examples/cd_text_imdb.nblink)
| doc/source/methods/ksdrift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Branje ontologij s strežnika
#
# Primer uporabe API-ja za prenos ontologije iz strežnika, branje in izpisovanje ontologije.
# +
import os
import tempfile
import shutil
from textsemantics import OntologyAPI
from utils.ontology_utils import print_onto_tree
from owlready2 import onto_path, Thing, World
# -
# Ustvarimo povezavo s strežnikom. Pri povezovanju na strežnik projekta, ni potrebno podati naslova strežnika. Nato pridobimo imena vseh ontologij, ki so na voljo na strežniku in jih izpišemo.
api = OntologyAPI()
ontologies = api.list_ontologies()
print("\n".join(ontologies))
# Izbrali smo ontologijo `core-sskj-only.owl`. Ustvarimo začasni direktorij in v njega prenesemo ontologijo ter ontologije, ki jih dana ontologija potrebuje. Uporabimo že obstoječo povezavo do strežnika.
dirpath = tempfile.mkdtemp()
ontology_name = "core-sskj-only.owl"
api.download_ontology(ontology_name, dirpath)
# Odpremo ontologijo in izraze v njej izpišemo v drevesni strukturi.
# +
# dodaj pot do direktorija v onto_path, da knjižni owlready2 ve od kod uvoziti ontologije
# na katerih ontologija bazira
if dirpath not in onto_path:
onto_path.append(dirpath)
world = World()
onto = world.get_ontology("file://" + os.path.join(dirpath, ontology_name)).load()
with onto:
print_onto_tree(Thing, world)
# -
# Ko končamo z delom, izbrišemo začasni direktorij iz računalnika.
shutil.rmtree(dirpath)
| examples/01-02-ontologies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Waiting Time Paradox, or, Why Is My Bus Always Late?
# *This notebook originally appeared as a [post](http://jakevdp.github.io/blog/2018/09/13/waiting-time-paradox/) on the blog [Pythonic Perambulations](http://jakevdp.github.io).*
# <!-- PELICAN_BEGIN_SUMMARY -->
#
# 
# *Image Source: [Wikipedia](https://en.wikipedia.org/wiki/RapidRide#/media/File:King_County_Metro_Rapid_Ride_New_Flyer_DE60LFR_6060.JPG) License CC-BY-SA 3.0*
#
# If you, like me, frequently commute via public transit, you may be familiar with the following situation:
#
# > *You arrive at the bus stop, ready to catch your bus: a line that advertises arrivals every 10 minutes. You glance at your watch and note the time... and when the bus finally comes 11 minutes later, you wonder why you always seem to be so unlucky.*
#
# Naïvely, you might expect that if buses are coming every 10 minutes and you arrive at a random time, your average wait would be something like 5 minutes.
# In reality, though, buses do not arrive exactly on schedule, and so you might wait longer.
# It turns out that under some reasonable assumptions, you can reach a startling conclusion:
#
# **When waiting for a bus that comes on average every 10 minutes, your average waiting time will be 10 minutes.**
#
# This is what is sometimes known as the *waiting time paradox*.
#
# I've encountered this idea before, and always wondered whether it is actually true... how well do those "reasonable assumptions" match reality?
# This post will explore the waiting time paradox from the standpoint of both simulation and probabilistic arguments, and then take a look at some real bus arrival time data from the city of Seattle to (hopefully) settle the paradox once and for all.
#
# <!-- PELICAN_END_SUMMARY -->
# ## The Inspection Paradox
# If buses arrive exactly every ten minutes, it's true that your average wait time will be half that interval: 5 minutes.
# Qualitatively speaking, it's easy to convince yourself that adding some variation to those arrivals will make the average wait time somewhat longer, as we'll see here.
#
# The waiting time paradox turns out to be a particular instance of a more general phenomenon, the *inspection paradox*, which is discussed at length in this enlightening post by <NAME>: [The Inspection Paradox Is Everywhere](
# http://allendowney.blogspot.com/2015/08/the-inspection-paradox-is-everywhere.html).
#
# Briefly, the inspection paradox arises whenever the probability of observing a quantity is related to the quantity being observed.
# Allen gives one example of surveying university students about the average size of their classes. Though the school may truthfully advertise an average of 30 students per class, the average class size *as experienced by students* can be (and generally will be) much larger. The reason is that there are (of course) more students in the larger classes, and so you oversample large classes when computing the average experience of students.
#
# In the case of a nominally 10-minute bus line, sometimes the span between arrivals will be longer than 10 minutes, and sometimes shorter, and if you arrive at a random time, you have more opportunities to encounter a longer interval than to encounter a shorter interval. And so it makes sense that the average span of time *experienced by riders* will be longer than the average span of time between buses, because the longer spans are over-sampled.
#
# But the waiting time paradox makes a stronger claim than this: when the average span between arrivals is $N$ minutes, the average span *experienced by riders* is $2N$ minutes.
# Could this possibly be true?
# ## Simulating Wait Times
# To convince ourselves that the waiting time paradox is making a reasonable claim, let's start by simulating a stream of buses that arrive at an average of 10 minutes.
# For the sake of numerical accuracy, we will simulate a large number of bus arrivals: one million buses (or approximately 19 years of round-the-clock 10-minute headways):
# +
import numpy as np
N = 1000000 # number of buses
tau = 10 # average minutes between arrivals
rand = np.random.RandomState(42) # universal random seed
bus_arrival_times = N * tau * np.sort(rand.rand(N))
# -
# Just to confirm we've done things correctly, let's check that the mean interval is close to $\tau = 10$:
intervals = np.diff(bus_arrival_times)
intervals.mean()
# With these bus arrivals simulated, we can now simulate the arrival of a large number of passengers to the bus stop during this span, and compute the wait time that each of them experiences.
# Let's encapsulate this in a function for later use:
def simulate_wait_times(arrival_times,
rseed=8675309, # Jenny's random seed
n_passengers=1000000):
rand = np.random.RandomState(rseed)
arrival_times = np.asarray(arrival_times)
passenger_times = arrival_times.max() * rand.rand(n_passengers)
# find the index of the next bus for each simulated passenger
i = np.searchsorted(arrival_times, passenger_times, side='right')
return arrival_times[i] - passenger_times
# We can then simulate some wait times and compute the average:
wait_times = simulate_wait_times(bus_arrival_times)
wait_times.mean()
# The average wait time is also close to 10 minutes, just as the waiting time paradox predicted.
# ## Digging Deeper: Probabilities & Poisson Processes
# How can we understand what's going on here?
#
# Fundamentally, this is an instance of the inspection paradox, in which the probability of observing a value is related to the value itself.
# Let's denote by $p(T)$ the distribution of intervals $T$ between buses as they arrive at a bus stop. In this notation, the expectation value of the arrival times is
# $$
# E[T] = \int_0^\infty T~p(T)~dT
# $$
# In the above simulation, we had chosen $E[T] = \tau = 10$ minutes.
# When a rider arrives at a bus stop at a random time, the probability of the time interval they experience will be affected by $p(T)$, but also by $T$ itself: the longer the interval, the larger the probability is that a passenger will experience it.
#
# So we can write the distribution of arrival times experienced by passengers:
# $$
# p_{exp}(T) \propto T~p(T)
# $$
# The constant of proportionality comes from normalizing the distribution:
# $$
# p_{exp}(T) = \frac{T~p(T)}{\int_0^\infty T~p(T)~dT}
# $$
# Comparing to above we see this simplifies to
# $$
# p_{exp}(T) = \frac{T~p(T)}{E[T]}
# $$
# The expected wait time $E[W]$ will then be half of the expected interval experienced by passengers, so we can write
# $$
# E[W] = \frac{1}{2}E_{exp}[T] = \frac{1}{2}\int_0^\infty T~p_{exp}(T)~dT
# $$
# which can be rewritten in a more suggestive way:
# $$
# E[W] = \frac{E[T^2]}{2E[T]}
# $$
# and now all that remains is for us to choose a form for $p(T)$ and compute the integrals.
# ### Choosing *p(T)*
#
# With this formalism worked out, what is a reasonable distribution to use for $p(T)$?
# We can get a picture of the $p(T)$ distribution within our simulated arrivals by plotting a histogram of the intervals between arrivals:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn')
plt.hist(intervals, bins=np.arange(80), density=True)
plt.axvline(intervals.mean(), color='black', linestyle='dotted')
plt.xlabel('Interval between arrivals (minutes)')
plt.ylabel('Probability density');
# -
# The vertical dotted line here shows the mean interval of about 10 minutes.
# This looks very much like an exponential distribution, and that is no accident: our simulation of bus arrival times as uniform random numbers very closely approximates a [Poisson process](https://en.wikipedia.org/wiki/Poisson_point_process), and for such a process it can be shown that the distribution of intervals between arrivals is exponential.
#
# (Side note: In our case this is only approximately exponential; in reality the intervals $T$ between $N$ uniformly sampled points within a timespan $N\tau$ follow the [Beta distribution](https://en.wikipedia.org/wiki/Beta_distribution): $T/(N\tau) \sim \mathrm{Beta}[1, N]$, which in the large $N$ limit approaches $T \sim \mathrm{Exp}[1/\tau]$.
# See, e.g. [this StackExchange post](https://math.stackexchange.com/questions/68749/difference-of-order-statistics-in-a-sample-of-uniform-random-variables), or [this twitter thread](https://twitter.com/jakevdp/status/1038086415190351872) for more details).
#
# An exponential distribution of intervals implies that the arrival times follow a Poisson process.
# To double-check this reasoning, we can confirm that it matches another property of a Poisson process: the number of arrivals within a fixed span of time will be Poisson-distributed.
# Let's check this by binning our simulated arrivals into hourly blocks:
# +
from scipy.stats import poisson
# count the number of arrivals in 1-hour bins
binsize = 60
binned_arrivals = np.bincount((bus_arrival_times // binsize).astype(int))
x = np.arange(20)
# plot the results
plt.hist(binned_arrivals, bins=x - 0.5, density=True, alpha=0.5, label='simulation')
plt.plot(x, poisson(binsize / tau).pmf(x), 'ok', label='Poisson prediction')
plt.xlabel('Number of arrivals per hour')
plt.ylabel('frequency')
plt.legend();
# -
# The close match between the empirical and theoretical values gives confidence that our interpretation is correct: for large $N$, the arrival times we simulated above are well-described by a Poisson process, which implies exponentially-distributed arrival intervals.
# That means we can write the probability distribution:
# $$
# p(T) = \frac{1}{\tau}e^{-T/\tau}
# $$
# Plugging this into the above results, we find that the average waiting time experienced by a person is
# $$
# E[W] = \frac{\int_0^\infty T^2~e^{-T/\tau}}{2\int_0^\infty T~e^{-T/\tau}} = \frac{2\tau^3}{2(\tau^2)} = \tau
# $$
# For bus arrivals consistent with a Poisson process, the expected wait time for a passenger is identical to the average interval between arrivals.
# A complementary way to reason about this is this: a Poisson process is a *memoryless* process, meaning that the history of events has no bearing on the expected time to the next event.
# So when you arrive at the bus stop, the average waiting time until the next bus is always the same: in our case, it is 10 minutes, and this is regardless of how long it has been since the previous bus!
# Along the same lines, it does not matter how long you have been waiting already: the expected time to the next arrival is always exactly 10 minutes: for a Poisson process, you get no "credit" for time spent waiting.
# ## Waiting Times In Reality
#
# The above is well and good if real-world bus arrivals are actually described by a Poisson process, but are they?
#
# [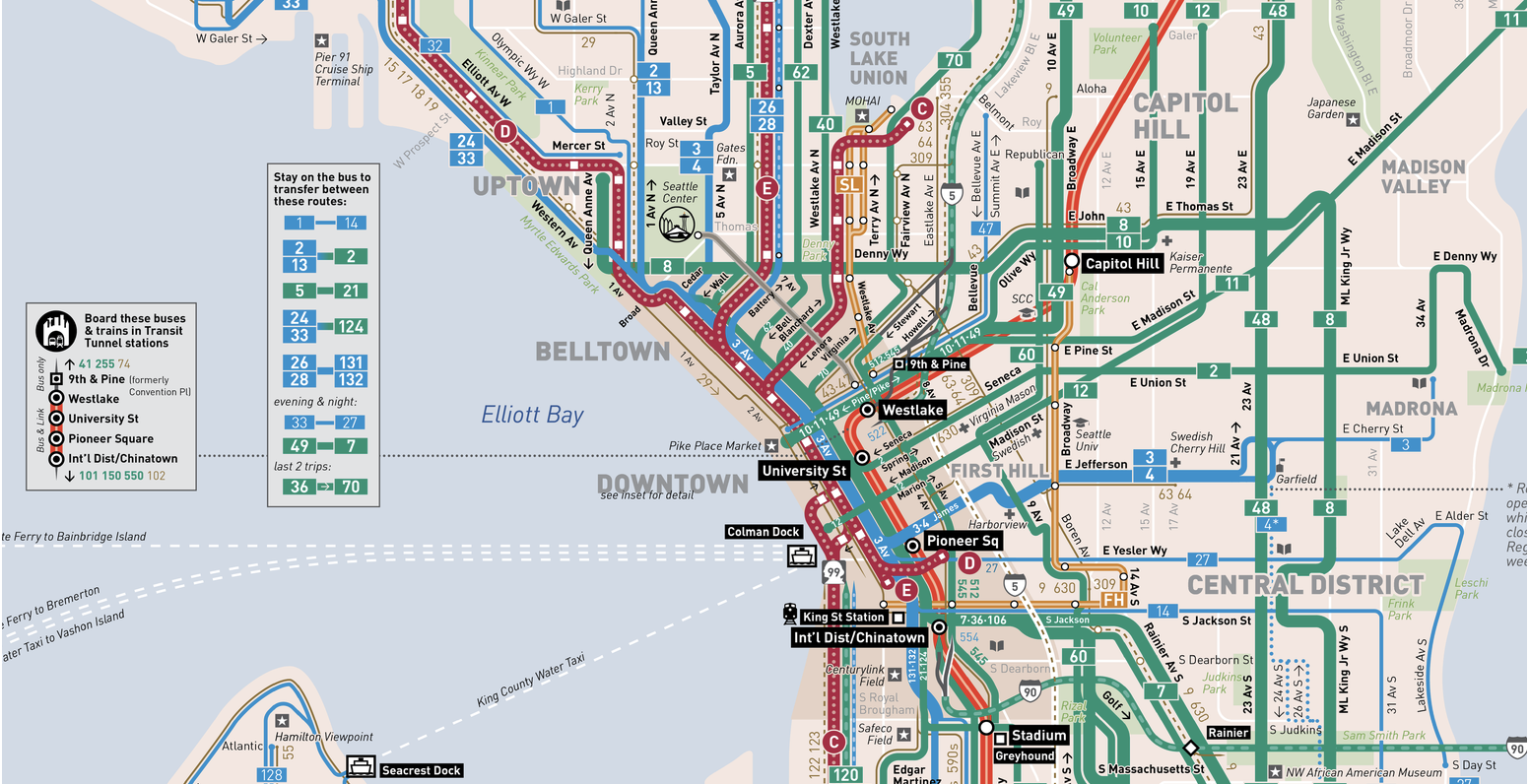](https://seattletransitmap.com/)
# *Image Source: https://seattletransitmap.com/*
#
# To determine whether the waiting time paradox describes reality, we can dig into some data, available for download here: [arrival_times.csv](https://gist.githubusercontent.com/jakevdp/82409002fcc5142a2add0168c274a869/raw/1bbabf78333306dbc45b9f33662500957b2b6dc3/arrival_times.csv) (3MB CSV file).
# The dataset contains scheduled and actual arrival times for Seattle's [Rapid Ride](https://kingcounty.gov/depts/transportation/metro/travel-options/bus/rapidride.aspx) lines C, D, and E at the 3rd & Pike bus stop in downtown Seattle, recorded during the second quarter of 2016 (huge thanks to <NAME> of the Washington State Transportation Center for providing this data!).
import pandas as pd
df = pd.read_csv('arrival_times.csv')
df = df.dropna(axis=0, how='any')
df.head()
# The reason I sought data from Rapid Ride routes in particular is that for much of the day, the buses are scheduled at regular intervals of between 10 and 15 minutes — not to mention the fact that I'm a frequent passenger on the C-line.
# ### Data Cleanup
#
# To start with, let's do a little bit of data cleanup to get it into a form that's easier to work with:
# +
# combine date and time into a single timestamp
df['scheduled'] = pd.to_datetime(df['OPD_DATE'] + ' ' + df['SCH_STOP_TM'])
df['actual'] = pd.to_datetime(df['OPD_DATE'] + ' ' + df['ACT_STOP_TM'])
# if scheduled & actual span midnight, then the actual day needs to be adjusted
minute = np.timedelta64(1, 'm')
hour = 60 * minute
diff_hrs = (df['actual'] - df['scheduled']) / hour
df.loc[diff_hrs > 20, 'actual'] -= 24 * hour
df.loc[diff_hrs < -20, 'actual'] += 24 * hour
df['minutes_late'] = (df['actual'] - df['scheduled']) / minute
# map internal route codes to external route letters
df['route'] = df['RTE'].replace({673: 'C', 674: 'D', 675: 'E'}).astype('category')
df['direction'] = df['DIR'].replace({'N': 'northbound', 'S': 'southbound'}).astype('category')
# extract useful columns
df = df[['route', 'direction', 'scheduled', 'actual', 'minutes_late']].copy()
df.head()
# -
# ### How Late Are Buses?
# There are essentially six different datasets within this table: the northbound and southbound directions for each of the C, D, and E lines.
# To get a feeling for their characteristics, let's plot a histogram of the actual minus scheduled arrival times for each of these six:
import seaborn as sns
g = sns.FacetGrid(df, row="direction", col="route")
g.map(plt.hist, "minutes_late", bins=np.arange(-10, 20))
g.set_titles('{col_name} {row_name}')
g.set_axis_labels('minutes late', 'number of buses');
# You might expect that the buses stick closer to their schedule near the beginning of each one-way trip and show more spread near the end, and this is borne out in the data: the southbound C-line and northbound D and E lines are near the beginning of their respective routes, and in the opposite direction they are near the end.
# ### Scheduled and Observed Arrival Intervals
# Let's next take a look at the observed and scheduled intervals between arrivals for these six routes.
# We'll start by using Pandas `groupby` functionality to compute these intervals:
# +
def compute_headway(scheduled):
minute = np.timedelta64(1, 'm')
return scheduled.sort_values().diff() / minute
grouped = df.groupby(['route', 'direction'])
df['actual_interval'] = grouped['actual'].transform(compute_headway)
df['scheduled_interval'] = grouped['scheduled'].transform(compute_headway)
# -
g = sns.FacetGrid(df.dropna(), row="direction", col="route")
g.map(plt.hist, "actual_interval", bins=np.arange(50) + 0.5)
g.set_titles('{col_name} {row_name}')
g.set_axis_labels('actual interval (minutes)', 'number of buses');
# It's already clear that these don't look much like the exponential distribution of our model, but that is not telling us much yet: the distributions may be affected by non-constant scheduled arrival intervals.
#
# Let's repeat the above chart, examining the scheduled rather than observed arrival intervals:
g = sns.FacetGrid(df.dropna(), row="direction", col="route")
g.map(plt.hist, "scheduled_interval", bins=np.arange(20) - 0.5)
g.set_titles('{col_name} {row_name}')
g.set_axis_labels('scheduled interval (minutes)', 'frequency');
# This shows that the buses come at variety of arrival intervals thorughout the week, so we cannot evaluate the accuracy of the waiting time paradox from the distributions of raw arrival times.
# ### Constructing Uniform Schedules
# Even though the scheduled arrival intervals are not uniform, there are a few particular intervals that have a large number of arrivals: for example, there are nearly 2000 northbound E-line buses with a scheduled interval of 10 minutes.
# In order to explore whether the waiting time paradox applies, let's group the data by line, direction, and scheduled interval, then re-stack these similar arrivals together as if they had happened in sequence.
# This should maintain all the relevant characteristics of the raw data, while making it easier to directly compare with the predictions of the waiting time paradox.
# +
def stack_sequence(data):
# first, sort by scheduled time
data = data.sort_values('scheduled')
# re-stack data & recompute relevant quantities
data['scheduled'] = data['scheduled_interval'].cumsum()
data['actual'] = data['scheduled'] + data['minutes_late']
data['actual_interval'] = data['actual'].sort_values().diff()
return data
subset = df[df.scheduled_interval.isin([10, 12, 15])]
grouped = subset.groupby(['route', 'direction', 'scheduled_interval'])
sequenced = grouped.apply(stack_sequence).reset_index(drop=True)
sequenced.head()
# -
# Using this cleaned data, we can plot the distribution of "actual" arrival intervals for each route, direction, and arrival frequency:
for route in ['C', 'D', 'E']:
g = sns.FacetGrid(sequenced.query(f"route == '{route}'"),
row="direction", col="scheduled_interval")
g.map(plt.hist, "actual_interval", bins=np.arange(40) + 0.5)
g.set_titles('{row_name} ({col_name:.0f} min)')
g.set_axis_labels('actual interval (min)', 'count')
g.fig.set_size_inches(8, 4)
g.fig.suptitle(f'{route} line', y=1.05, fontsize=14)
# We see that for each line and schedule, the distribution of observed arrival intervals is nearly Gaussian, is peaked near the scheduled arrival interval, and has a standard deviation that is smaller near the beginning of the route (southbound for C, northbound for D/E) and larger near the end.
# Even without a statistical test, it's clear by eye that the actual arrival intervals are definitely **not** exponentially distributed, which is the basic assumption on which the waiting time paradox rests.
#
# We can make use of the wait time simulation function we used above in order to find the average wait time for each bus line, direction, and schedule:
grouped = sequenced.groupby(['route', 'direction', 'scheduled_interval'])
sims = grouped['actual'].apply(simulate_wait_times)
sims.apply(lambda times: "{0:.1f} +/- {1:.1f}".format(times.mean(), times.std()))
# The average waiting times are are perhaps a minute or two longer than half the scheduled interval, but not equal to the scheduled interval as the waiting time paradox implied. In other words, the inspection paradox is confirmed, but the waiting time paradox does not appear to match reality.
# ## Final Thoughts
#
# The waiting time paradox has been an interesting launching-point for a discussion that covered simulation, probability, and comparison of statistical assumptions with reality.
# Although we confirmed that real-world bus lines do follow some version of the inspection paradox, the above analysis shows pretty definitively that the core assumption behind the waiting time paradox — that the arrival of buses follows the statistics of a Poisson process — is not well-founded.
#
# In retrospect, this is perhaps not all that surprising: a Poisson process is a memoryless process that assumes the probability of an arrival is entirely independent of the time since the previous arrival.
# In reality, a well-run bus system will have schedules deliberately structured to avoid this kind of behavior: buses don't begin their routes at random times throughout the day, but rather begin their routes on a schedule chosen to best serve the transit-riding public.
#
# The larger lesson here is that you should be careful about the assumptions you bring to any data analysis task.
# A Poisson process is a good description for arrival time data — sometimes.
# But just because one type of data sounds like another type of data, it does not mean that assumptions valid for one are necessarily valid for the other.
# Often assumptions that seem correct on their face can lead to conclusions that don't match reality.
# *This post was written entirely in the Jupyter notebook. You can
# [download](http://jakevdp.github.io/downloads/notebooks/WaitingTimeParadox.ipynb)
# this notebook, or see a static view
# [on nbviewer](http://nbviewer.jupyter.org/url/jakevdp.github.io/downloads/notebooks/WaitingTimeParadox.ipynb).*
| content/downloads/notebooks/.ipynb_checkpoints/WaitingTimeParadox-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Bytes generalization
# +
import numpy as np
import pandas as pd
import pylab as pl
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv('tmp/session_c.csv')
# -
df.dtypes
pl.figure(figsize=(12, 6))
p=df.groupby('Bytes')['Bytes'].count().plot.bar(width=2,edgecolor='blue',color='blue')
p.tick_params(labelbottom='off',top='off',bottom='off')
p.set_yscale('log')
fig = p.get_figure()
fig.savefig('tmp/bytes000.png')
df['Bytes2']=np.floor(df.Bytes/10)*10+10
pl.figure(figsize=(12, 6))
p=df.groupby('Bytes2')['Bytes2'].count().plot.bar(width=2,edgecolor='blue',color='blue')
p.tick_params(labelbottom='off',top='off',bottom='off')
p.set_yscale('log')
fig = p.get_figure()
fig.savefig('tmp/bytes001.png')
df['Bytes3']=np.floor(df.Bytes/100)*100+100
pl.figure(figsize=(12, 6))
p=df.groupby('Bytes3')['Bytes3'].count().plot.bar(width=2,edgecolor='blue',color='blue')
p.tick_params(labelbottom='off',top='off',bottom='off')
p.set_yscale('log')
fig = p.get_figure()
fig.savefig('tmp/bytes002.png')
df.to_csv('tmp/session_d.csv')
| hmac/bytes_generalization00.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/WebheadTech/QCourse511-1/blob/main/X_Y_Zgate.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="k7AcQiWFoNBG"
pip install tensorflow==2.4.1 tensorflow-quantum
# + colab={"base_uri": "https://localhost:8080/"} id="2zPU6XhpoUDi" outputId="43e92f7a-11a2-4382-f23a-5bbf4e9d59cf"
# Update package resources to account for version changes.
import importlib, pkg_resources
importlib.reload(pkg_resources)
# + colab={"base_uri": "https://localhost:8080/"} id="Uee3lKN6odrl" outputId="14de90aa-cfd6-4f93-f847-5f990f7eb644"
from google.colab import drive
drive.mount('/content/drive')
# + id="fJxNuCnKogSf"
import sys
sys.path.append('/content/drive/My Drive')
# + id="5PuTd2K7oi1T"
import eecs598
import torch
import torchvision
import matplotlib.pyplot as plt
import statistics
import numpy as np
# + id="cZtski2Soki-"
import tensorflow as tf
import tensorflow_quantum as tfq
import cirq
import sympy
import numpy as np
import seaborn as sns
import collections
# visualization tools
# %matplotlib inline
import matplotlib.pyplot as plt
from cirq.contrib.svg import SVGCircuit
# + id="DbO_hc-Qoo7U"
# Control qrid size for visualization
plt.rcParams['figure.figsize'] = (10.0, 8.0)
plt.rcParams['font.size'] = 16
# + colab={"base_uri": "https://localhost:8080/"} id="_ST2VEy6orgv" outputId="b0080373-ef78-45fd-de48-875f8289599f"
x_train, y_train, x_test, y_test = eecs598.data.cifar10()
print('Training set:', )
print(' data shape:', x_train.shape)
print(' labels shape: ', y_train.shape)
print('Test set:')
print(' data shape: ', x_test.shape)
print(' labels shape', y_test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 466} id="_0QaS22xozsh" outputId="12e34c72-cb52-4b38-a10e-d431eb55fede"
#Visualization of the dataset
import random
from torchvision.utils import make_grid
classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
samples_per_class = 12
samples = []
for y, cls in enumerate(classes):
plt.text(-4, 34 * y + 18, cls, ha='right')
idxs, = (y_train == y).nonzero(as_tuple=True)
for i in range(samples_per_class):
idx = idxs[random.randrange(idxs.shape[0])].item()
samples.append(x_train[idx])
img = torchvision.utils.make_grid(samples, nrow=samples_per_class)
plt.imshow(eecs598.tensor_to_image(img))
plt.axis('off')
plt.show()
# + id="nyACAiMgpQ2c"
# + id="cqE-lL-kpREy"
from tensorflow.keras.datasets import cifar10
(train_images, train_labels), (test_images, test_labels) = cifar10.load_data()
# + id="-Ykse73DpcB9"
y_train=train_labels.flatten()
y_test=test_labels.flatten()
# We have reduced the dimension
# + [markdown] id="X-UU4Y__5Icq"
#
# + id="jtTIg2Y6pdTB"
x_train=tf.image.rgb_to_grayscale(train_images)
x_test=tf.image.rgb_to_grayscale(test_images)
#to convert grayscale
# + id="P-lucPulpdY2"
#Filter the dataset to keep just the cat and frog, remove the other classes
def filter_36(x, y):
keep = (y == 3) | (y == 6)
x, y = x[keep], y[keep]
y = y == 3
return x,y
# + colab={"base_uri": "https://localhost:8080/"} id="Be-MhjKepdbf" outputId="ca79e3e9-c9e4-4681-9099-48598774d3cb"
x_train, y_train = filter_36(x_train, y_train)
x_test, y_test = filter_36(x_test, y_test)
print("Number of filtered training examples:", len(x_train))
print("Number of filtered test examples:", len(x_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 509} id="QmeCy__1plRz" outputId="37be68a8-f874-4582-b933-5a319e5eb688"
plt.imshow(x_train[0, :, :, 0])
plt.colorbar()
# + id="Wy-eSnfMplTy"
x_train_s = tf.image.resize(x_train, (4,4)).numpy()
x_test_s = tf.image.resize(x_test, (4,4)).numpy()
# + id="ihqMYCdEplWW"
THRESHOLD = 0.5
x_train_bin = np.array(x_train_s > THRESHOLD, dtype=np.float32)
x_test_bin = np.array(x_test_s > THRESHOLD, dtype=np.float32)
# + id="118nmrnNv_iM"
def convert_to_circuit(image):
"""Encode truncated classical image into quantum datapoint."""
values = np.ndarray.flatten(image)
qubits = cirq.GridQubit.rect(4, 4)
circuit = cirq.Circuit()
for i, value in enumerate(values):
if value:
circuit.append(cirq.Circuit(cirq.X(qubits[i]), cirq.Y(qubits[i]),cirq.Z(qubits[i]) ))
return circuit
x_train_circ = [convert_to_circuit(x) for x in x_train_bin]
x_test_circ = [convert_to_circuit(x) for x in x_test_bin]
# + colab={"base_uri": "https://localhost:8080/", "height": 841} id="LYE9Z8C7wvzw" outputId="ab3fc4b4-cd0c-47a2-8b06-404f7fa5eff9"
SVGCircuit(x_train_circ[0])
# + id="0SK8_6_apsKW"
x_train_tfcirc = tfq.convert_to_tensor(x_train_circ)
x_test_tfcirc = tfq.convert_to_tensor(x_test_circ)
# + id="IVdMNERfxYB6"
class CircuitLayerBuilder():
def __init__(self, data_qubits, readout):
self.data_qubits = data_qubits
self.readout = readout
def add_layer(self, circuit, gate, prefix):
for i, qubit in enumerate(self.data_qubits):
symbol = sympy.Symbol(prefix + '-' + str(i))
circuit.append(gate(qubit, self.readout)**symbol)
# + colab={"base_uri": "https://localhost:8080/", "height": 271} id="fWduftMNrKV5" outputId="2f2dbde7-52ca-4650-ede4-3b58eb917ea6"
demo_builder = CircuitLayerBuilder(data_qubits = cirq.GridQubit.rect(4,1),
readout=cirq.GridQubit(-1,-1))
circuit = cirq.Circuit()
demo_builder.add_layer(circuit, gate = cirq.XX, prefix='xx')
SVGCircuit(circuit)
# + id="I0CyUSn6xc_q"
def create_quantum_model():
"""Create a QNN model circuit and readout operation to go along with it."""
data_qubits = cirq.GridQubit.rect(4, 4) # a 4x4 grid.
readout = cirq.GridQubit(-1, -1) # a single qubit at [-1,-1]
circuit = cirq.Circuit()
# Prepare the readout qubit.
circuit.append(cirq.X(readout))
circuit.append(cirq.H(readout))
builder = CircuitLayerBuilder(
data_qubits = data_qubits,
readout=readout)
# Then add layers (experiment by adding more).
builder.add_layer(circuit, cirq.XX, "xx1")
builder.add_layer(circuit, cirq.ZZ, "zz1")
# Finally, prepare the readout qubit.
circuit.append(cirq.H(readout))
return circuit, cirq.Z(readout)
# + id="9PsUr6KJxgS7"
model_circuit, model_readout = create_quantum_model()
# + id="NmfTiWAMxigM"
model = tf.keras.Sequential([
# The input is the data-circuit, encoded as a tf.string
tf.keras.layers.Input(shape=(), dtype=tf.string),
# The PQC layer returns the expected value of the readout gate, range [-1,1].
tfq.layers.PQC(model_circuit, model_readout),
])
# + id="YY8EMExLxl1b"
y_train_hinge = 2.0*y_train-1.0
y_test_hinge = 2.0*y_test-1.0
# + id="XQGv-WyOxnw5"
def hinge_accuracy(y_true, y_pred):
y_true = tf.squeeze(y_true) > 0.0
y_pred = tf.squeeze(y_pred) > 0.0
result = tf.cast(y_true == y_pred, tf.float32)
return tf.reduce_mean(result)
# + id="ga10r-dZxyxn"
model.compile(
loss=tf.keras.losses.Hinge(),
optimizer=tf.keras.optimizers.Adam(),
metrics=[hinge_accuracy])
# + colab={"base_uri": "https://localhost:8080/"} id="fNOojLwHx1uY" outputId="689e5525-0bf5-45f8-8b1c-155e061d5494"
print(model.summary())
# + id="kdf-U4fyx3k_"
EPOCHS = 3
BATCH_SIZE = 128
NUM_EXAMPLES = len(x_train_tfcirc)
# + id="KMreBNgxx66F"
x_train_tfcirc_sub = x_train_tfcirc[:NUM_EXAMPLES]
y_train_hinge_sub = y_train_hinge[:NUM_EXAMPLES]
# + id="nxwUG0Lrx68i"
import time
start_time = time.time()
# + colab={"base_uri": "https://localhost:8080/"} id="_eNEAcD2x6_J" outputId="5e1830cf-b0f8-4061-af27-037a3cdedd6a"
qnn_history = model.fit(
x_train_tfcirc_sub, y_train_hinge_sub,
batch_size=32,
epochs=EPOCHS,
verbose=1,
validation_data=(x_test_tfcirc, y_test_hinge))
qnn_results = model.evaluate(x_test_tfcirc, y_test)
# + id="5ABnhw8lv4i7" outputId="b39aa638-9b14-427a-af50-0c55e4c1f6b3" colab={"base_uri": "https://localhost:8080/"}
qnn_accuracy = qnn_results[1]
qnn_accuracy
# + id="fbpSCyqkyq2e" outputId="581d7cae-d5e1-40c6-eef2-c186d8429be2" colab={"base_uri": "https://localhost:8080/"}
model.predict_classes(x_train_tfcirc[0:7])
# + id="SumHeJtSytbu" outputId="bc8bf38d-a23c-4a5b-a708-f697588eff46" colab={"base_uri": "https://localhost:8080/", "height": 687}
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.gridspec import GridSpec
def format_axes(fig):
for i, ax in enumerate(fig.axes):
ax.tick_params(labelbottom=False, labelleft=False)
fig = plt.figure(figsize=(10, 10))
gs = GridSpec(3, 3, figure=fig)
ax1 = fig.add_subplot(gs[0, 0])
# identical to ax1 = plt.subplot(gs.new_subplotspec((0, 0), colspan=3))
for i in range(3):
for j in range(3):
ax = fig.add_subplot(gs[i, j])
ax.imshow(x_train[i+j, :, :, 0])
fig.suptitle("GridSpec")
format_axes(fig)
plt.show()
# + id="9FLuon6izF-Q" outputId="b0a30603-3681-4d33-f496-c9b6a53d3c1c" colab={"base_uri": "https://localhost:8080/", "height": 507}
plt.plot(qnn_history.history['hinge_accuracy'], label='QNN')
#plt.plot(hybrid_history.history['val_custom_accuracy'], label='Hybrid CNN')
#plt.title('Quantum vs Hybrid CNN performance')
plt.xlabel('Epochs')
plt.legend()
plt.ylabel('Validation Accuracy')
plt.show()
| X_Y_Zgate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## WeRateDogs Act Report
#
# ## We rate dogs data
#
# People who have animals have always loved to talk about them and take lots of pictures. In the rate dogs twitter these people who have animals find a common place to fall in love, laugh and brighten their day with photos and comments of various dogs.
# Through these movements we can collect various data to better understand the characteristics of favorite animals and also the behavior of this community.
#
# In this report we will analyse and visualize the wrangled data collected.
# After doing all the gathering, assessing and cleaning the data, the content that was in the merged table was better to do some analysis.
#
# We found three insights that are found programmatically and two found visually for quality and tidiness issues:
#
# ### Insights
#
# 1. The quantity of people who favorite the posts is 2.039 times higher than people that retweet the posts. This shows a preference of just favorite the posts rather than retweeting them.
#
# 2. There are a strong correlation between the favorites counts and retweets. To be more precise the correlation is 0.801345. To evidence better, the most retweeted and favorite dog is a doggo labrador retriever who received 72474 retweets and 147742 favorites votes. His ID is 744234799360020481.
#
# 3. The most common dog breeds are golden retriever, labrador retriever and pembroke, respectively. They receive the most favorite counts too. In my country, in Brasil, we can see in the streets and public garden this preferences. Most dog breeds are retrievers!
#
#
# 4. We can have a visual look in the Daily tweets by favorite count chart and verify a positive trend in the amount of favorite tweets over time. This makes sense because of popularity and increase of the twitters counts and social media.
#
# 5. In the cloud chart we can see that the word pooper, dog, pup and meet are the most frequently written in the twitters posts. It matches with the most popular dog stages found in the We Rate Dogs data.
#
#
#
#
| act_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Deepmeta Train model
#
# In this notebook, we will create a hyper parameter object containing all our hyper parameters, select images and train a network.
# + pycharm={"name": "#%%\n"}
import DeepMeta.train as d_t
import DeepMeta.utils.global_vars as gv
# + [markdown] pycharm={"name": "#%% md\n"}
# First, we need to create an object containing all our hyper parameters.
# When you run the train script from Deepmeta, this object is create by parsing command line arguments.
# + pycharm={"name": "#%%\n"}
conf = {
"n_epochs": 200, # number of epochs
"batch_size": 128, # size of batches
"lr": 0.001, # learning rate value
"model_name": "small++", # name of the model to use (unet or small++)
"meta": False, # do you want to segment metas ?
"weighted": True, # Do you want to use weighted cross entropy
"w1": 2, # Inside weight
"w2": 4, # Border weight
"size": 128, # Image size
"drop_r": 0.2, # Dropout rate
"filters": 16, # Number of filters in the first conv block
"patience": 10, # Patience for early stopper
}
# + [markdown] pycharm={"name": "#%% md\n"}
# Once we have created our hyper parameters object, we just need to call the training function.
# This function will load the dataset, the model, then train the model, save the weights and plot the training curve.
# > We assume that the global vars script is correctly fulfill.
# + pycharm={"name": "#%%\n"}
d_t.train(
conf,
path_images=gv.path_img,
path_labels=gv.path_lab,
hp_search=False,
)
# + [markdown] pycharm={"name": "#%% md\n"}
# TODO : ADD FILTER VIZ
| examples/train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Library Imports
import ipywidgets as widgets
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# Slider class to combine slide and play handler
class Slider:
def __init__(self, call_back, descp, default_val=0, minimum=0, maximum=10):
# Create player and slider
self.player = widgets.Play(min=minimum, interval=1000, value=default_val)
self.slider = widgets.FloatSlider(description=descp, min=minimum, max=maximum, step=0.1, continuous_update=True,
layout=widgets.Layout(width='40%'))
self.link(call_back)
# Link slider and player
def link(self, on_value_change):
self.slider.observe(on_value_change, 'value')
widgets.jslink((self.player, 'value'), (self.slider, 'value'))
# Signal class to generate function points and hold signal parameters
class Signal:
def __init__(self, function, color, descp):
self.function = function
self.color = color
self.amplitude = 1
self.frequency = 1
self.patch = mpatches.Patch(color=self.color, label=descp)
def update(self, params):
self.amplitude = params.get("Amplitude", self.amplitude)
self.frequency = params.get("Frequency", self.frequency)
def generate_points(self, time_values):
return self.amplitude * self.function(self.frequency * time_values)
# Graph class to hold all plotting variables
class Graph:
def __init__(self, slider_list, function_list, title, interval=0.001, limit=6):
# Create title based on type of plot
temp = r' of $e^{i \Omega t}, \Omega = $' if interval < 0.5 else r' of $e^{i \omega n}, \omega = $'
self.title = title + temp
self.sliders = [Slider(self.call_back, *slide) for slide in slider_list]
self.functions = [Signal(*funct) for funct in function_list]
self.time_values = np.arange(0, limit, interval)
self.out = widgets.Output()
self.fig, self.ax = plt.subplots(figsize=(8, 6))
self.ax.grid(True)
self.ax.set_ylabel("Amplitude")
self.ax.axis([0, limit, -10, 10])
self.ax.axhline(0, linewidth=1, color='black')
self.ax.axvline(0, linewidth=1, color='black')
self.ax.legend(handles=[funct.patch for funct in self.functions])
self.initial_plot(interval)
plt.close(self.fig)
self.refresh()
# Create the first plot
def initial_plot(self, interval):
params = {} if interval < 0.01 else {'marker':'.', 'linestyle':'None'}
self.curve = {}
# Plot the functions
for funct in self.functions:
self.curve[funct.color], = self.ax.plot(
self.time_values, funct.generate_points(self.time_values), color=funct.color, **params)
# Get all elements to display
display_list = [widgets.HBox([slide.slider, slide.player]) for slide in self.sliders]
display_list.append(self.out)
self.full_output = widgets.VBox(display_list)
# Respond to slider changes
def call_back(self, change):
for funct in self.functions:
funct.update({change["owner"].description : change["new"]})
if change["owner"].description == "Frequency":
if len(self.time_values) < 50:
self.ax.set_title(self.title + " %.2f radians/sample" % (change["new"]))
else:
self.ax.set_title(self.title + " %.2f radians/second" % (change["new"]))
self.curve[funct.color].set_ydata(funct.generate_points(self.time_values))
self.refresh()
# Refresh the plot
def refresh(self):
self.out.clear_output(wait=True)
with self.out:
display(self.fig)
# Display the plot
def display_graph(self):
display(self.full_output)
self.refresh()
# Place all graphs inside tabs
def display_tab(plots, titles):
tab = widgets.Tab()
tab.children = plots
for i in range(len(titles)):
tab.set_title(i, titles[i])
display(tab)
# +
# Continuous Time Complex Exponential
cont_cplx = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.cos, "red", "Real"], [np.sin, "blue", "Imaginary"]],
"Continuous Time Complex Exponential\nBoth components")
cont_real = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.cos, "red", "Real"]],
"Continuous Time Complex Exponential\nReal component")
cont_imag = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.sin, "blue", "Imaginary"]],
"Continuous Time Complex Exponential\nImaginary component")
tab_contents = ["Real", "Imaginary", "Both"]
display_tab([cont_real.full_output, cont_imag.full_output, cont_cplx.full_output], tab_contents)
# +
# Discrete Time Complex Exponential
dcrt_cplx = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.cos, "red", "Real"], [np.sin, "blue", "Imaginary"]],
"Discrete Time Complex Exponential\nBoth components", 1, 30)
dcrt_real = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.cos, "red", "Real"]],
"Discrete Time Complex Exponential\nReal component", 1, 30)
dcrt_imag = Graph([["Amplitude", 1], ["Frequency", 1]], [[np.sin, "blue", "Imaginary"]],
"Discrete Time Complex Exponential\nImaginary component" ,1, 30)
tab_contents = ["Real", "Imaginary", "Both"]
display_tab([dcrt_real.full_output, dcrt_imag.full_output, dcrt_cplx.full_output], tab_contents)
# -
| sinusoidal_and_exponential_signals/sinusoidal_curves.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ricacero/Datascience300/blob/main/Caso_Accidentes_NYC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HjFPSXI-50BD"
# # ¿Cómo podemos controlar el creciente número de accidentes en Nueva York?
# + id="Ke4NpVH750BM"
import json
import requests
from bs4 import BeautifulSoup
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy
# + [markdown] id="N-xgSbBb50BO"
# ## Introduccion
# + [markdown] id="bJuobUO-50BP"
# **Contexto empresarial.** La ciudad de Nueva York ha experimentado un aumento en el número de accidentes en las carreteras de la ciudad. Quieren saber si el número de accidentes ha aumentado en las últimas semanas. Para todos los accidentes reportados, han recopilado detalles para cada accidente y han estado manteniendo registros durante el último año y medio (desde enero de 2018 hasta agosto de 2019).
#
# La ciudad te ha contratado para que construyas visualizaciones que les ayuden a identificar patrones en accidentes, lo que les ayudaría a tomar acciones preventivas para reducir la cantidad de accidentes en el futuro. Tienen ciertos parámetros como municipio, hora del día, motivo del accidente, etc. De los que se preocupan y de los que les gustaría obtener información específica.
# + [markdown] id="Y92vVvGw50BQ"
# **Problema comercial.** Su tarea es formatear los datos proporcionados y proporcionar visualizaciones que respondan las preguntas específicas que tiene el cliente, que se mencionan a continuación.
# + [markdown] id="xAMWU3nw50BR"
# **Contexto analítico.** Se le proporciona un archivo CSV (almacenado en la carpeta ```datos``` ya creada) que contiene detalles sobre cada accidente, como fecha, hora, ubicación del accidente, motivo del accidente, tipos de vehículos involucrados, recuento de lesiones y muertes, etc. El delimitador en el archivo CSV dado es `;` en lugar del predeterminado `,`. Realizará las siguientes tareas con los datos:
#
# 1. Extraiga datos adicionales del municipio almacenados en un archivo JSON
# 2. Leer, transformar y preparar datos para su visualización
# 3. Realizar análisis y construir visualizaciones de los datos para identificar patrones en el conjunto de datos.
#
# El cliente tiene un conjunto específico de preguntas a las que le gustaría obtener respuestas. Deberá proporcionar visualizaciones para acompañar estos:
#
# 1. ¿Cómo ha fluctuado el número de accidentes durante el último año y medio? ¿Han aumentado con el tiempo?
# 2. Para un día en particular, ¿durante qué horas es más probable que ocurran accidentes?
# 3. ¿Hay más accidentes entre semana que durante los fines de semana?
# 4. ¿Cuál es la proporción de recuento de accidentes por área por municipio? ¿Qué distritos tienen un número desproporcionadamente grande de accidentes para su tamaño?
# 5. Para cada municipio, ¿durante qué horas es más probable que ocurran accidentes?
# 6. ¿Cuáles son las 5 principales causas de accidentes en la ciudad?
# 7. ¿Qué tipos de vehículos están más involucrados en accidentes por municipio?
# 8. ¿Qué tipos de vehículos están más involucrados en las muertes?
# + [markdown] id="-ihuFfJ550BS"
# ## Obteniendo los datos relevantes
# + [markdown] id="VWtb3s_J50BU"
# El cliente ha solicitado un análisis de la relación accidentes-área de los distritos. Los datos del municipio se almacenan en un archivo JSON en la carpeta ```datos```.
# + [markdown] id="A481OPcp50BW"
# ### Pregunta
#
# Utilice la función ```json.load()``` para cargar el archivo ```borough_data.json``` como diccionario.
# + [markdown] id="r6zUG_k550BX"
# **Respuesta.**
# + colab={"base_uri": "https://localhost:8080/"} id="aB6_h0gh6F-s" outputId="453f0d5e-37c1-4575-a81d-e96e88a8e423"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="oRLTAUkk50BZ"
# ### Pregunta
# De manera similar, use la función pandas ```read_csv()``` para cargar el archivo ```accidentes.csv``` como un DataFrame. Nombra este DataFrame ```df```.
#
# + colab={"base_uri": "https://localhost:8080/"} id="XAaG09iy6uL2" outputId="98921d21-c969-4e2b-977a-e3e9295da452"
# %cd '/content/drive/MyDrive/Caso NYC'
# + colab={"base_uri": "https://localhost:8080/"} id="ApUMe2jf50BY" outputId="23114eaa-c9c3-4586-a94c-6abcefaccb06"
with open('data/borough_data.json') as f:
borough_data=json.load(f)
borough_data
# + [markdown] id="ORj4_V3z50Ba"
# **Respuesta.**
# + id="00g85RAO50Bb"
with open('data/accidents.csv') as f:
df=pd.read_csv(f, delimiter=';')
# + colab={"base_uri": "https://localhost:8080/", "height": 870} id="D8R7Riid50Bc" outputId="64ad21f2-662d-4333-d789-d452afa6c88a"
df=pd.read_csv('data/accidents.csv',delimiter=";")
df
# + [markdown] id="VZzRQhi250Bc"
# ## Overview de la data
# + [markdown] id="xcPqdhj250Bd"
# Analizemos las columnas presentes en el data frame
# + colab={"base_uri": "https://localhost:8080/"} id="kNn6JplU50Bd" outputId="67f013e4-7f29-4a35-c906-2558082f2b8d"
df.columns
# + [markdown] id="6F7RMhY250Be"
# Tenemos las siguientes columnas
#
# 1. **BOROUGH**: el municipio en el que ocurrió el accidente
# 2. **COLLISION_ID**: un identificador único para esta colisión
# 3. **CONTRIBUTING FACTOR VEHICLE (1, 2, 3, 4, 5)**: Motivos del accidente
# 4. **CROSS STREET NAME**: Calle transversal más cercana al lugar del accidente
# 5. **DATE**: Fecha del accidente
# 6. **TIME**: Hora del accidente
# 7. **LATITUDE**: Latitud del accidente
# 8. **LONGITUDE**: Longitud del accidente
# 9. **NUMBER OF (CYCLISTS, MOTORISTS, PEDESTRIANS) INJURED**: Lesión por categoría
# 10. **NUMBER OF (CYCLISTS, MOTORISTS, PEDESTRIANS) DEATHS**: Categoría muerte sabia
# 11. **ON STREET NAME**: Calle donde ocurrió el accidente
# 13. **VEHICLE TYPE CODE(1, 2, 3, 4, 5)**: Tipos de vehículos involucrados en el accidente
# 14. **ZIP CODE**: código postal del lugar del accidente
# + [markdown] id="r3utFUxl50Bf"
# ### Ejercicio 1
#
# Desde 2014, la ciudad de Nueva York ha estado implementando un plan de seguridad vial llamado [Vision Zero](https://www1.nyc.gov/content/visionzero/pages/). Su objetivo es reducir el número de muertes por accidentes de tránsito a *cero* para fines de 2024. El plan está creando nuevas y mejorando las medidas de seguridad actuales, algunas de ellas incluyen:
#
# - [ ] Detección de peatones automatizada
# - [ ] Auditorías de seguridad vial en lugares de alto riesgo
# - [ ] Ampliación de la red de carriles bici
# - [ ] Iniciativas específicas de educación y sensibilización
# - [ ] Creación de islas de refugio peatonal
# - [ ] Lanzamiento del programa integrado de reductor de velocidad basado en datos (topes y amortiguadores de velocidad)
#
# ¿Cuáles de estas iniciativas podrían beneficiarse directamente de un análisis de los datos proporcionados? Marque todo lo que corresponda.
#
# **Nota:** Puede crear marcas de verificación haciendo doble clic en esta celda y agregando una ```[x]``` en las líneas correspondientes.
# + [markdown] id="QdoGTVXa50Bf"
# **Respuesta.**
#
# - [ ] Detecciòn de peatones automatizada
#
#
# - [ ] Auditorías de seguridadvial en lugares de alto riesgo
#
# - [ ] Ampliación de la red de carriles bici
#
# - [ ] Iniciativas especificas de educación y sensibilización
#
# - [ ] Creación de islas de refugio peatonal
#
# - [ ] Lanzamiento del programa integrado de reductor de velocidad en datos (topes y amortiguadores de velocidad)
#
# + [markdown] id="GCIirr2350Bg"
# Sigamos adelante y respondamos a cada una de las preguntas del cliente.
# + [markdown] id="rcKHFt6F50Bg"
# ## Respondiendo las preguntas del cliente
# + [markdown] id="_gZdhHVr50Bh"
# ### Ejercicio 2
#
# Agrupe los datos disponibles mensualmente y genere un line plot de accidentes a lo largo del tiempo. ¿Ha aumentado el número de accidentes durante el último año y medio?
#
# **Sugerencia**: Puede encontrar útiles las funciones de pandas ```to_datetime ()``` y ```dt.to_period ()```.
# + [markdown] id="xxpPiVPe50Bh"
# **Respuesta.**
# + [markdown] id="-SxpC_uwSXJ6"
# Si ha habido una disminución de accidentalidad en el ultimo año y medio
# + colab={"base_uri": "https://localhost:8080/", "height": 433} id="nHBdWEr750Bh" outputId="bcba6108-b98d-4ea0-aaa0-b2d22a99a5c4"
df['DATE']=pd.to_datetime(df['DATE'])
monthly_accidents =df.groupby(df['DATE'].dt.to_period('M')).size()
plt.figure(figsize=(15,6))
monthly_accidents.plot.line()
plt.xlabel('Fecha')
plt.ylabel('Frecuencia')
plt.title('Evolucion de Accidentes NYC')
# + colab={"base_uri": "https://localhost:8080/", "height": 453} id="w2y1Qrs_89pk" outputId="29fcc884-b0d0-461c-ab23-2286a215395d"
df['DATE']=pd.to_datetime(df['DATE'])
monthly_accidents =df.groupby(df['DATE'].dt.to_period('M')).size()
plt.figure(figsize=(15,6))
monthly_accidents.plot.bar()
plt.xlabel('Fecha')
plt.ylabel('Frecuencia')
plt.title('Evolucion de Accidentes NYC x Meses')
# + colab={"base_uri": "https://localhost:8080/", "height": 677} id="iTXHzQxzG9KA" outputId="d7342ba0-cf31-47dc-aae1-3dd06ecbf673"
dft= pd.DataFrame()
dft['Accidentes']= monthly_accidents
dft['Fecha']=dft.index
dft=dft.reset_index()
dft
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="Lg5rYPSS50Bi" outputId="7377d9a2-96e0-47f3-8b17-0d8f9c5b6c94"
df['DATE']=pd.to_datetime(df['DATE']) # Formato datetime python
monthly_accidents =df.groupby(df['DATE'].dt.to_period('D')).size()
plt.figure(figsize=(15,6))
monthly_accidents.plot.line()
# + [markdown] id="Q-FKSTZN50Bj"
#
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="oV83_BizHVMZ" outputId="801f28dc-1bec-4249-f4e7-035c6f4aade6"
dft= pd.DataFrame()
dft['Accidentes']= monthly_accidents
dft['Fecha']=dft.index
dft=dft.reset_index()
dft
# + [markdown] id="QIkdAt2t50Bj"
# ### Ejercicio 3
#
# De la gráfica anterior, ¿qué meses parecen tener el menor número de accidentes? ¿Cuáles crees que son las razones detrás de esto?
# + [markdown] id="xD4O5y1v50Bj"
# **Respuesta.**
#
# + [markdown] id="eJPXZPesUnh0"
# Los meses con menos accidentalidad son el mes de febrero tanto del 2018 y 2019, pero hay que tener en cuenta que estos son dos meses de 28 días comparados con los demás, le siguen abril del 2019 y enero de 2019 que tienen 31 días cada uno y es un dato más confiable. Pensaría que la accidentalidad puede estar relacionada con las estaciones, en los meses de invierno tiende a disminuir comparado con los de verano, pero es muy relativo. El mes de Agosto de 2019 no es comparable porque hay solo datos hasta el 24 de ese mes por lo que faltarían datos de 7 días por eso no lo tengo en cuenta.
# + [markdown] id="wI203KnJ50Bk"
# ### Ejercicio 4
# ¿Cómo varía el número de accidentes a lo largo de un solo día? Cree una nueva columna `HOUR` basada en los datos de la columna `TIME`, luego trace un gráfico de barras de la distribución por hora a lo largo del día.
#
# **Sugerencia:** Puede encontrar útil la función ```dt.hour```.
# + [markdown] id="Sp5ev-1_50Bl"
# **Respuesta.**
# + [markdown] id="Zk60JOVlAg02"
# Segun la grafica podriamos decir que la accidentalidad es baja en horas de la noche, se puede intuir que no hay tantas personas afuera, y tambien se visualiza un ascenso en los accidentes a la hora de regreso a casa desde las 4 a 6 y alta accidentalidad en horas de la mañana cuando se presume van al trabajo.
# + colab={"base_uri": "https://localhost:8080/", "height": 974} id="7AvKMUEKwfMd" outputId="ff90591d-24ab-4dda-bf85-0222f927a0e5"
import datetime as dt
df['TIME']=pd.to_datetime(df['TIME'])
df['HOUR'] = df['TIME'].dt.hour
df
# + id="j0aaT9kj15kV"
df = df[['DATE', 'TIME','HOUR', 'BOROUGH', 'ZIP CODE', 'LATITUDE', 'LONGITUDE',
'ON STREET NAME', 'NUMBER OF PEDESTRIANS INJURED',
'NUMBER OF PEDESTRIANS KILLED', 'NUMBER OF CYCLIST INJURED',
'NUMBER OF CYCLIST KILLED', 'NUMBER OF MOTORIST INJURED',
'NUMBER OF MOTORIST KILLED', 'CONTRIBUTING FACTOR VEHICLE 1',
'CONTRIBUTING FACTOR VEHICLE 2', 'CONTRIBUTING FACTOR VEHICLE 3',
'CONTRIBUTING FACTOR VEHICLE 4', 'CONTRIBUTING FACTOR VEHICLE 5',
'COLLISION_ID', 'VEHICLE TYPE CODE 1', 'VEHICLE TYPE CODE 2',
'VEHICLE TYPE CODE 3', 'VEHICLE TYPE CODE 4', 'VEHICLE TYPE CODE 5',
]]
# + colab={"base_uri": "https://localhost:8080/", "height": 974} id="R-uhPp2R2SO9" outputId="5254bf77-6b80-431f-ffe5-a268a534f221"
df
# + colab={"base_uri": "https://localhost:8080/", "height": 315} id="-8MH5PZx4hzs" outputId="02c117b4-e190-481b-89be-a667a6dfb811"
HA=df.groupby(['HOUR']).size().reset_index(name='numero')
HA.plot.bar()
plt.xlabel('Hora')
plt.ylabel('Cantidad')
plt.title('Accidentes hora en el dia')
# + colab={"base_uri": "https://localhost:8080/", "height": 802} id="JeumQb_56Q3J" outputId="97609375-1907-4b0f-8b3e-cee1c286db38"
df.groupby(['HOUR']).size().reset_index(name='numero')
# + [markdown] id="hbtO7JIJ50Bm"
# ### Ejercicio 5
#
# En la pregunta anterior hemos agregado el número de accidentes por hora sin tener en cuenta la fecha y el lugar en que ocurrieron. ¿Qué crítica le daría a este enfoque?
# + [markdown] id="aKY2n-Uo50Bm"
# **Respuesta.**
#
# + [markdown] id="yLNtu9QWCjDg"
# Para una mejor precisión debemos saber el lugar en donde ocurrieron, hay barrios que según los datos tienen mas trafico y afluencia de personas que otros, también como sabemos que los accidentes aumentan dependiendo del clima la estación en que nos encontremos eso se debe tener en cuenta.
# + [markdown] id="GF_nCbJH50Bo"
# ### Ejercicio 6
#
# ¿Cómo varía el número de accidentes en una sola semana? Trace un gráfico de barras basado en el recuento de accidentes por día de la semana.
#
# **Sugerencia:** Puede encontrar útil la función ```dt.weekday```.
# + [markdown] id="k6z2Uv4s50Bo"
# **Respuesta.**
# + [markdown] id="e9XPS_SH50Bp"
# Podemos visualizar que los dias con mas accidentalidad es el dia viernes, y lo que menos tienen son el dia Domingo y el sabado.
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="NAk-KtqyKrN6" outputId="79b4dc7e-80fd-4185-aa76-c994ff9fc435"
df['DATE']=pd.to_datetime(df['DATE'])
WA =df.groupby(df['DATE'].dt.weekday).size()
plt.figure(figsize=(15,6))
WA.plot.bar()
plt.xlabel('Fecha')
plt.ylabel('Frecuencia')
plt.title('Evolucion de Accidentes NYC x Meses')
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="-nrSP4bCNcFi" outputId="17e7d58b-5bb2-4cf5-e852-1c6d6c659a17"
df.groupby(df['DATE'].dt.weekday).size().reset_index(name='numero')
# + [markdown] id="puag_dvN50Bp"
# ### Ejercicio 7
#
# Trace una gráfica de barras del número total de accidentes en cada municipio, así como uno de los accidentes por milla cuadrada por municipio. ¿Qué puedes concluir?
#
# **Sugerencia:** Es posible que desee actualizar algunas de las claves en el diccionario del municipio para que coincidan con los nombres en el marco de datos.
# + [markdown] id="1KoXyAmL50Bq"
# **Respuesta.**
# + [markdown] id="6R5u4sqgFTqg"
# Se puede concluir que en el municipio de Manhattan es el de mas accidentalidad con respecto al area que maneja, y el de menor accidentalidad es el municipio de STATEN ISLAND respecto al area que tiene.
#
# + id="5a2JKpMn50Bq" colab={"base_uri": "https://localhost:8080/"} outputId="d76a9fe7-f7f6-469d-fb99-cd0bd5a519d2"
df['BOROUGH'].unique()
# + [markdown] id="afMbbr4J50Bq"
#
# + id="7QygHllZgIzY" outputId="d6c881eb-6bfa-4479-d470-87cde4e7cae4" colab={"base_uri": "https://localhost:8080/"}
borough_data
# + colab={"base_uri": "https://localhost:8080/"} outputId="d76a9fe7-f7f6-469d-fb99-cd0bd5a519d2" id="BONYrHgojNUs"
df['BOROUGH'].unique()
# + id="nRJQV-IV50Bq"
borough_data.update({
'BRONX': {'area': 42.1, 'name': 'BRONX', 'population': 1471160.0}})
del borough_data[ 'the bronx']
# + [markdown] id="BPGnNPrH50Br"
#
# + id="GUqNEwM850Br" colab={"base_uri": "https://localhost:8080/"} outputId="d484d863-843e-431b-b697-9ec47bfe1613"
borough_data
# + [markdown] id="_SSuekPj50Br"
#
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="VFVhlbe1nG-J" outputId="f5d429fa-ec31-42a5-9d49-e13763f3c364"
df1 = pd.DataFrame([key for key in borough_data.keys()], columns=['BOROUGH'])
df1['area'] = [value['area'] for value in borough_data.values()]
df1['name'] = [value['name'] for value in borough_data.values()]
df1['population'] = [value['population'] for value in borough_data.values()]
df1
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="6aBlvlX2rplo" outputId="cf8d2316-f0da-4e28-d24e-15d2711f0910"
df1 ['BOROUGH'] = df1 ['BOROUGH']. str.upper ()
df1
# + colab={"base_uri": "https://localhost:8080/", "height": 922} id="aqkc3e2zmKCx" outputId="78715452-2633-47c8-bbe3-b178454a0a84"
merged_df = pd.merge(left=df,right=df1, left_on='BOROUGH', right_on='BOROUGH')
merged_df
# + colab={"base_uri": "https://localhost:8080/"} id="5A9eLNFJs4sB" outputId="e357e934-d130-416b-c3f5-a24a1467aca3"
merged_df.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 614} id="akxg2jMBwCPY" outputId="c1a04151-6391-4d90-ebef-ac0fafb5141b"
BA=pd.DataFrame({'accidentes': merged_df.groupby(['BOROUGH','area']).size()})
BA.plot.bar()
BA
# + colab={"base_uri": "https://localhost:8080/"} id="qDk7rmXcC2j1" outputId="028028ab-1216-4d9b-af8d-d9193db98563"
BA.sum()
# + [markdown] id="r_QIV58L50Br"
# ### Ejercicio 8
#
# ¿Qué horas tienen más accidentes en cada municipio? Trace un gráfico de barras para cada municipio que muestre el número de accidentes por cada hora del día.
#
# **Sugerencia:** Puede usar ```sns.FacetGrid``` para crear una cuadrícula de parcelas con los datos por hora de cada municipio.
# + [markdown] id="GhKs2dw350Bs"
# **Respuesta.**
# + id="U3xNspXo50Bs" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="02e57a44-1e23-4752-b4c6-370278a186f3"
df3 = pd.DataFrame({'count': df.groupby(['BOROUGH', 'HOUR']).size()})
df3 = df3.reset_index()
df3
# + colab={"base_uri": "https://localhost:8080/", "height": 499} id="ymMdZKCFJ4sX" outputId="c5a9b8be-ebb7-4437-dd96-40395e8ec198"
chart = sns.FacetGrid(df3, col='BOROUGH', margin_titles=True, col_wrap=3, aspect=2, row_order=df['BOROUGH'].unique)
chart
chart.map(sns.barplot, 'HOUR', 'count')
# + [markdown] id="lPzJS5zN50Bs"
# **¿Es mayor el número de accidentes en diferentes momentos en diferentes distritos? ¿Deberíamos concentrarnos en diferentes momentos para cada municipio?**
# + [markdown] id="PglMHNEt50Bs"
# En lso municipios de Broklyn, Bronx y Staten Island la hora con mas accidentalidad es las 16 horas, en MAN es las 14 horas y en QUEENS las 17 horas, pero en todos los municipios manejan la mas alta accidentalidad desde las 14 horas hasta las 18 horas.
# + [markdown] id="NeZeGGb-50Bu"
# ### Ejercicio 9
#
# ¿Qué factores provocan la mayoría de los accidentes? Evite contar dos veces los factores que contribuyen a un solo accidente.
#
# **Sugerencia:** Una forma de lidiar con las repeticiones es concatenar las columnas correspondientes conservando sus índices, puede hacerlo con las funciones ```pd.concat()``` y ```reset_index()```. Luego, use un ```group_by``` apropiado para contar el número de repeticiones de factores contribuidos por accidente.
# + [markdown] id="gDOp91ep50Bu"
# **Respuesta**.
#
# + [markdown] id="fRIqcBrs6zuu"
# Los siguientes son los factores que provocan la mayoria de accidentes:
#
# Driver Inattention/Distraction
# Failure to Yield Right-of-Way
# Following Too Closely
# Backing Unsafely
# Passing Too Closely
# Passing or Lane Usage Improper
# + colab={"base_uri": "https://localhost:8080/"} id="VnvCbuhfu5Y5" outputId="290231c4-d3b0-44cb-fd9e-f96fcdc998b9"
merged_df.value_counts(['CONTRIBUTING FACTOR VEHICLE 1'])
# + [markdown] id="aSKvNiDl6yQ7"
#
# + [markdown] id="xTVWSY-i6mM1"
#
# + colab={"base_uri": "https://localhost:8080/"} id="6HlFzzWoxuqP" outputId="888ee874-579d-422b-9fc6-3fd1cfaa05ef"
merged_df.isna().sum()
# + id="3LInziqe50Bv" colab={"base_uri": "https://localhost:8080/", "height": 2318} outputId="47d41e6a-a704-41b8-d972-03a0c9cace16"
merged_df.groupby(['CONTRIBUTING FACTOR VEHICLE 1','CONTRIBUTING FACTOR VEHICLE 2','CONTRIBUTING FACTOR VEHICLE 3','CONTRIBUTING FACTOR VEHICLE 4','CONTRIBUTING FACTOR VEHICLE 5']).size().reset_index(name='numero')
# + [markdown] id="RinfqcKf50Bv"
# ### Ejercicio 10
#
# ¿Qué tipos de vehículos están más involucrados en accidentes por municipio? Evite contar dos veces el tipo de vehículos presentes en un solo accidente.
#
# **Sugerencia:** Puede aplicar un enfoque similar al utilizado en la pregunta anterior.
# + [markdown] id="UflSBUzS50Bv"
# **Respuesta.**
# + [markdown] id="6BgzhvXh9gHf"
# Los vehiculos mas involucrados por municipio son SEDAN, Station Wagon/Sport Utility Vehicle y PASSENGER VEHICLE
# + id="c3h-MJg550Bw" colab={"base_uri": "https://localhost:8080/", "height": 397} outputId="aeb34880-19b4-461e-a5a3-38a3077792ca"
data_crosstab = pd.crosstab(merged_df['BOROUGH'],
merged_df['VEHICLE TYPE CODE 1'],
margins = True)
data_crosstab
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="n2FH0C-s1fHw" outputId="d17601f6-9047-4104-a0aa-729e5e619c8c"
df4 = pd.DataFrame({'count': merged_df.groupby(['BOROUGH', 'VEHICLE TYPE CODE 1']).size()})
df4 = df4.reset_index()
df4
# + [markdown] id="MqICskoF50Bw"
#
# + [markdown] id="fK8Yaio050Bw"
# ### Ejercicio 11
#
# En 2018 para una [entrevista](https://www.nytimes.com/2019/01/01/nyregion/traffic-deaths-decrease-nyc.html) con The New York Times, el alcalde de Blasio de Nueva York declaró que *'Vision Zero está funcionando claramente'*. Ese año, el número de muertes en accidentes de tráfico en Nueva York se redujo a un histórico 202. Sin embargo, según lo informado por [am New York Metro](https://www.amny.com/news/vision-zero-de-blasio- 1-30707464 /), el número de víctimas mortales ha aumentado un 30% en el primer trimestre de 2019 en comparación con el año anterior y el número de peatones y ciclistas heridos no ha experimentado ninguna mejora.
#
# ¿Cómo utilizaría los datos proporcionados para comprender qué salió mal en el primer trimestre de 2019?
#
#
# > - [ ] Considere los accidentes del primer trimestre de 2019. Luego, busque las causas más comunes de accidentes en los que estuvieron involucrados peatones y ciclistas. Dé una recomendación basada únicamente en esta información.
#
# > - [X ] Cree un par de mapas de calor de los accidentes que involucraron a peatones y ciclistas lesionados / muertos en el primer trimestre de 2018 y 2019. Compare estos dos para ver si hay algún cambio en la concentración de accidentes. En áreas críticas, estudie el tipo de factores involucrados en los accidentes. Dé una recomendación para visitar estas áreas para estudiar más el problema.
#
# > - [ ] Los datos proporcionados son insuficientes para mejorar nuestra comprensión de la situación.
#
# > - [ ] Ninguna de las anteriores. Haría lo siguiente: *aquí tu respuesta recomendada*.
# + [markdown] id="Ur0G9GNt50Bx"
# **Respuesta.**
#
# + id="qrEAzB8P50By"
# + [markdown] id="JgImd_Rr50By"
# ### Ejercicio 12
#
# Calcula el número de muertes provocadas por cada tipo de vehículo. Trace un gráfico de barras para los 5 vehículos principales. ¿Qué vehículos están involucrados con mayor frecuencia en las muertes y cuánto más que los demás?
#
# **Por ejemplo,** si dos personas murieron en un accidente en el que estuvieron involucrados 5 vehículos: 4 son VEHÍCULOS DE PASAJEROS y 1 es un VAGÓN DEPORTIVO / ESTACIÓN. Luego, agregaríamos dos muertes a cada tipo de VEHÍCULO DE PASAJEROS y VAGÓN DE ESTACIÓN / SERVICIO DEPORTIVO.
#
# **Sugerencia:** Es posible que desee crear una nueva columna con el número total de muertes en el accidente. Para eso, puede encontrar útil la función ```.to_numpy()```. Luego, proceda como los ejercicios anteriores para evitar contabilizar dos veces el tipo de vehículos.
# + [markdown] id="BAJmh8Qd50By"
# **Repsuesta.**
# + id="PPvAGRtl50Bz"
# + [markdown] id="amMGSqBW50Bz"
#
# + id="hVzOBoyV50Bz"
# + [markdown] id="ApE3BHeW50Bz"
#
| Caso_Accidentes_NYC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Работа с файловой системой
# ### Запись в существующий файл и чтение из файла.
# Основной синтаксис следующий:
# для записи строки в файл нужно открыть файл функцией file = **open('path_to_file', 'open_mode')**. <br> <br>
# **open_mode** означает режим, в котором нужно открыть файл. Для записи необходимо передать 'w'. Для дописывания - 'a'. Для чтения 'r'. Более подробно здесь https://www.tutorialspoint.com/python/python_files_io.htm
#
# Основные команды - write, read, readline, readlines, writeline, close.
# Последнюю команду необходимо выполнить в конце работы с фалйом.
file = open('a.txt', 'w')
file.write('Hell')
file.write('o, world!\n')
file.writelines(['abc\n', 'line2\n'])
file.close()
# +
# # !cat a.txt
# -
file = open('a.txt', 'r')
print(file.read())
file.close()
| materials/InputOutputToFile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
import math
import random
import struct
import matplotlib.pyplot as plt
from IPython.display import Audio, display
import numpy as np
import scipy.io.wavfile
def show(data):
plt.plot(range(len(data)), data)
plt.show()
def audio(data): display(Audio(data, rate=SR))
def show2(data1, data2):
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.set_size_inches(12, 4, forward=True)
ax1.plot(range(len(data1)), data1)
ax2.plot(range(len(data2)), data2)
plt.show()
from algomusic import *
# + [markdown] slideshow={"slide_type": "slide"}
# # Простые приемы звукового синтеза и алгоритмической композиции на языке Питон
#
# *<NAME>*, МИРЭА
#
# Часть 1. Синтез звука
#
# Часть 2. **Алгоритмическая композиция**
# + [markdown] slideshow={"slide_type": "slide"}
# # Алгоритмическая музыка до появления компьютеров
#
# * Китайские колокольчики.
# * Эолова арфа.
# * Музыкальная игра в кости (Моцарт и некоторые другие композиторы).
#
# <table>
# <tr>
# <td width="30%"><img src="img/chime.png" width="50%">
# <td width="30%"><img src="img/harp.png" width="70%">
# <td><img src="img/mozart.png">
# </table>
#
# + slideshow={"slide_type": "slide"}
# Вместо ветра используем ГПСЧ
MAJ_SCALE = [0, 2, 4, 5, 7, 9, 11] # Мажор
MIN_SCALE = [0, 2, 3, 5, 7, 8, 10] # Минор
MAJ_PENTA_SCALE = [0, 2, 4, 7, 9] # Мажорная пентатоника
MIN_PENTA_SCALE = [0, 3, 5, 7, 10] # Минорная пентатоника
MAJ_BLUES_SCALE = [0, 2, 3, 4, 7, 9] # Мажорная блюзовая гамма
MIN_BLUES_SCALE = [0, 3, 5, 6, 7, 10] # Минорная блюзовая гамма
# Перевод смещения ноты от С-4, с учетом гаммы и транспонирования, в герцы
def note2freq(offs, scale, trans=0):
note = scale[offs % len(scale)] + 12 * (offs // len(scale))
return midi2freq(60 + note + trans)
def get_note(): # Случайная нота из гаммы пентатоники
n = random.randint(0, 11)
return note2freq(n, MIN_PENTA_SCALE)
v1 = Voice(Sin(), Env())
d1 = Delay(sec(0.5))
out = []
for i in range(100):
vol = random.randint(3, 10) * 0.1 # Громкость варьируется
out += d1.play(v1.play(get_note(), sec(0.3), sec(0.3), amp=vol), 0.3, 0.85)
audio(out)
# + [markdown] slideshow={"slide_type": "slide"}
# # Некоторые ранние компьютерные композиции
#
# * *1956*. Песня "<NAME>" (Push-Button Bertha), <NAME> (<NAME>) и <NAME> (<NAME>), компьютер Datatron.
# * *1957*. Произведение в 4 частях для струнного квартета "<NAME>" (Illiac Suite), Лед<NAME> (<NAME>), компьютер ILLIAC I. Помимо прочего, использовались марковские цепи.
# * *1959*. Одноголосые мелодии "Уральские напевы" Р. Зарипова, компьютер "Урал".
# <center><img src="img/zaripov.png" width="30%"></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # "Кнопка Берта"
# <center><img src="img/bertha.png" width="30%"></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # "Кнопка Берта"
#
# Случайный выбор нот по правилам, на основе анализа популярных песен (TOP 10 1956 года), среди которых:
#
# * Длина песни в пределах 35-60 нот.
# * Песня имеет структуру AABA.
# * A содержит 8 тактов и насчитывает 18-25 нот.
# * B содержит 8 тактов и насчитывает 17-35 нот.
# * Не более 5 нот в одном направлении.
# * Скачки более чем на 6 ступеней запрещены.
#
# <small>http://www.musicainformatica.org/topics/push-button-bertha.php</small>
# + slideshow={"slide_type": "-"}
display(Audio(filename="mp3/bertha.mp3"))
# + [markdown] slideshow={"slide_type": "slide"}
# # "Уральские напевы"
#
# * Порождение мелодий состоит из двух этапов: 1) выбор ритма пьесы, 2) выбор высот нот.
# * Мелодии имеют структуру ABA.
# * Каждая фраза мелодии заканчивается на одной из 3 основных ступеней лада.
# * Запрещены следующие подряд два широких интервала.
# * Не более 6 нот в одном направлении.
# * Диапазон нот содержит 2.5 октавы.
#
# <small><NAME>, Об алгоритмическом описании процесса сочинения музыки, Докл. АН СССР, 1960,
# том 132, номер 6, 1283–1286</small>
# -
display(Audio(filename="mp3/ural.mp3"))
# + [markdown] slideshow={"slide_type": "slide"}
# # Triadex Muse
#
# 1972, алгоритмический секвенсер, <NAME> (<NAME>) и <NAME> (<NAME>). На основе счетчиков и 31-битного LFSR.
#
# <table>
# <tr>
# <td><center><img src="img/muse1.jpg" width="50%"></img>
# <td><center><img src="img/muse2.jpg"></img>
# </table>
#
# + slideshow={"slide_type": "slide"}
class Muse:
def __init__(self, interval, theme):
self.interval_sliders = interval
self.theme_sliders = theme
self.rows = [0, 1] + [0] * 38
self.scale = parse_track("c-3 d-3 e-3 f-3 g-3 a-3 b-3 c-4 c-4 d-4 e-4 f-4 g-4 a-4 b-4 c-5")
self.clock = 1
def get_freq(self):
values = [self.rows[i] for i in self.interval_sliders]
return self.scale[sum([x * 2**i for i, x in enumerate(values)])]
def update_lfsr(self):
xor = sum([self.rows[i] for i in self.theme_sliders]) % 2
self.rows = self.rows[:9] + [xor ^ 1] + self.rows[9:-1]
def pulse(self):
self.clock += 1
self.rows[2] = self.clock & 1
self.rows[3] ^= int(self.clock % 2 == 0)
self.rows[4] ^= int(self.clock % 4 == 0)
self.rows[5] ^= int(self.clock % 8 == 0)
self.rows[6] ^= int(self.clock % 16 == 0)
self.rows[7] ^= int(self.clock % 6 == 0)
self.rows[8] ^= int(self.clock % 12 == 0)
if self.rows[2] == 0:
self.update_lfsr()
# + slideshow={"slide_type": "slide"}
#m = Muse([15, 16, 13, 0], [0, 12, 31, 0])
#m = Muse([18, 16, 15, 0], [1, 5, 9, 10])
#m = Muse([3, 9, 39, 6], [0, 0, 9, 39])
m = Muse([9, 10, 5, 6], [0, 0, 39, 5])
tempo = sec(0.1)
v1 = Voice(Sin(), Env())
d1 = Delay(sec(0.5))
freq = None
out = []
for i in range(200):
new_freq = m.get_freq() * 4
if new_freq == freq:
new_freq = None
else:
freq = new_freq
out += d1.play(v1.play(new_freq, sec(0.3), tempo), 0.4)
m.pulse()
audio(out)
# + slideshow={"slide_type": "slide"}
# <NAME> (1995), <NAME>
step = 1
num = 0
out = []
for i in range(16):
print(format(i, "04b"), bin(num).count("1"))
num += step
# + slideshow={"slide_type": "slide"}
step = 63 # Шаг задает номер "композиции"
num = 1
tempo = sec(0.12)
v1 = Voice(LFSR(4, [3, 0]), Env(0.01))
d1 = Delay(sec(0.1))
f1 = LP1()
f2 = LP1()
out = []
for i in range(300): # Алгоритм MusiNum в действии
freq = note2freq(bin(num).count("1"), MAJ_SCALE, 24)
p = v1.play(freq, tempo, tempo, amp=1 if i % 4 == 0 else 0.7)
out += d1.play(f1.play(f2.play(p, 0.4), 0.4), 0.35, 0.7)
num += step
audio(out)
# + slideshow={"slide_type": "slide"}
# Фрактальный генератор мелодий
def similar(data, rule, times): # Порождение мелодии из исходных данных по правилу rule, times итераций
for i in range(times):
new = []
for x in data:
new += [x + offs for offs in rule] # замена очередной ноты по правилу
data = new
return data
rule = [0, 1]
for i in range(6):
print(" ".join(str(x) for x in similar([0], rule, i)))
# + slideshow={"slide_type": "slide"}
#rule = [0, 1]
rule = [0, 1, -1, 0]
#rule = [-2, 7, -7, 2]
#rule = [0, 2, 4, -7]
notes = similar([0], rule, 4)
v1 = Voice(FMSquare(), Env(0.01))
d1 = Delay(sec(0.1))
tempo = sec(0.12); out = []
for n in notes:
freq = note2freq(n, MAJ_SCALE, 12)
out += d1.play(v1.play(freq, tempo, tempo), 0.25, 0.8)
show2(rule, notes[:len(rule) * 30]); audio(out)
# + slideshow={"slide_type": "slide"}
# Вероятностные барабаны
kick_break = [
10, 0.4, None, None, 0.7, None, 0.8, None,
None, None, None, None, 0.9, None, None, None,
0.9, None, None, None, 0.9, None, 0.9, None,
None, None, 0.9, None, 0.6, None, None, None
]
snare_break = [
None, None, None, None, None, None, None, None,
0.8, None, 0.7, None, None, None, 0.7, None,
None, None, 0.7, None, None, None, None, None,
0.7, None, 0.7, None, None, None, None, None
]
hat_break = [SR, None, SR, None] * 8
tempo = sec(1/16)
v1 = Kick_voice()
v2 = Snare_voice()
v3 = Voice(LFSR(12, [10, 9, 1, 2, 3, 11]), Env())
d1 = Delay(sec(0.1))
out = []
for j in range(4): # Простое воспроизведение
for i in range(len(kick_break)):
p1 = v1.play(kick_break[i], tempo)
p2 = v2.play(snare_break[i], tempo)
p3 = v3.play(hat_break[i], sec(0.02), tempo, amp=0.4)
out += mix(p1, d1.play(p2, 0.1), p3)
audio(out)
# + slideshow={"slide_type": "slide"}
tempo = sec(1/16)
v1 = Kick_voice()
v2 = Snare_voice()
v3 = Voice(LFSR(12, [10, 9, 1, 2, 3, 11]), Env())
d1 = Delay(sec(0.1))
out = []
busy = 0.8
for j in range(8): # Барабанные вариации
for i in range(len(kick_break)):
is_kick = kick_break[i] is not None and random.random() < kick_break[i] * busy
is_snare = snare_break[i] is not None and random.random() < snare_break[i] * busy
is_hat = (hat_break[i] is not None) and (not is_kick and not is_snare)
p1 = v1.play(kick_break[i] if is_kick else None, tempo)
p2 = v2.play(snare_break[i] if is_snare else None, tempo)
p3 = v3.play(hat_break[i] if is_hat else None, sec(0.02), tempo, amp=0.4)
out += mix(p1, d1.play(p2, 0.1), p3)
audio(out)
scipy.io.wavfile.write("drums2.wav", SR, np.array(out))
# + slideshow={"slide_type": "slide"}
# Порождение ритма в духе "Уральских напевов"
def make_bar(size, durations): # Заполнение такта длительностями из durations
bar = []
while sum(bar) < size:
d = random.choice(durations)
if sum(bar) + sum(d) <= size:
bar += d
return bar
# См. пример на след. слайде
def next_note(note, intervals, note_range): # Выбор очередной ноты, случайное блуждание
while True:
ivals, iprobs, idir = intervals
direction = 2 * int(random.random() < idir) - 1
new_note = note + random.choices(ivals, iprobs)[0] * direction
if new_note in range(note_range):
return new_note
# + slideshow={"slide_type": "slide"}
# Сравнение контуров мелодий при случайном выборе нот и с помощью next_note()
intervals = [
[1, 2, 3, 4, 5, 6, 7], # Интервалы
[1, 0, 0, 0, 0, 0, 0], # Вероятность появления интервала
0.5 # Вероятность движения мелодии вверх
]
out = []
note = 0
for i in range(100):
note = next_note(note, intervals, 12)
out.append(note)
show2([random.randint(0, 11) for i in range(100)], out)
# + slideshow={"slide_type": "slide"}
# Алгоритмический фанк
intervals = [
[1, 2, 3, 4, 5, 6],
[0.5, 0.4, 0.03, 0.03, 0.03, 0.01],
0.6
]
# Набор длительностей для построения такта
durations = [[1/4], [1/2], [1/8, 1/8], [1/4 + 1/8, 1/8], [1/16, 1/16], [1/8 + 1/16, 1/16]]
v1 = Voice(LFSR(4, [3, 0]), Env())
out = []
note = 0
for i in range(8):
part = []
ab = make_bar(4/4, durations) + make_bar(4/4, durations)
for dur in ab:
note = next_note(note, intervals, 12)
part += v1.play(note2freq(note, MAJ_BLUES_SCALE, 12), sec(dur) * 1.2, sec(dur) * 2)
out += part * 2
fs, drums = scipy.io.wavfile.read("drums2.wav")
drums = list(drums) * (1 + len(out) // len(drums))
audio(mix(drums[:len(out)], out))
# + [markdown] slideshow={"slide_type": "slide"}
# # Riffology
#
# 1982, песня Song of the Grid для игры BallBlazer (Atari 400/800), <NAME> (<NAME>), LuscasFilm Games.
#
# Алгоритмическая соло-партия порождается на основе случайного выбора из набора риффов. Очередной рифф выбирается с учетом завершающей ноты предыдущего риффа.
#
# <small>http://www.langston.com/Papers/amc.pdf</small>
# -
display(Audio(filename="mp3/ballblazer.mp3"))
# + [markdown] slideshow={"slide_type": "slide"}
# # Генератор соло-партий из гитарного самоучителя
#
# 2000, Музыкальное и аппликатурное мышление гитариста, <NAME>.
#
# <table>
# <tr>
# <td><center><img src="img/popov1.png"></img>
# <td><center><img src="img/popov2.png"></img>
# </table>
#
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/popov3.png"></img></center>
# + slideshow={"slide_type": "slide"}
# 7 ступеней, 49 риффов
def split_by(lst, n): return [lst[i: i + n] for i in range(0, len(lst), n)]
riffs = [split_by(load_track("txt/riff%i.txt" % i), 9) for i in range(1, 8)]
v1 = Voice(LFSR(4, [3, 0]), Env())
d1 = Delay(sec(0.1))
f1 = LP1()
tempo = sec(1/8)
out = []
row = random.randint(0, len(riffs) - 1)
for i in range(32):
col = random.randint(0, len(riffs) - 1) # Выбор нового риффа
riff = riffs[row][col]
row = col # Выбор ступени
dur = tempo if random.random() < 0.7 else tempo * 2
for freq in riff[:-1]: # Последняя нота риффа заменяется первой нотой нового риффа
out += d1.play(f1.play(v1.play(freq * 4, dur * 1.2, dur), 0.25), 0.2, 0.7)
fs, drums = scipy.io.wavfile.read("drums1.wav")
drums = list(drums) * (1 + len(out) // len(drums))
audio(mix(drums[:len(out)], out))
# + slideshow={"slide_type": "slide"}
# Однострочные алгоритмические композиции
#def f(t): return t * (t >> 11) * t / 3
#def f(t): return div(t, (t & (t >> 12)))
def f(t): return (div(t, ( t >> 16 | t >> 8)) & (( t >> 5 | t >> 11))) -1 | t * (( t >> 16 | t >> 8))
#def f(t): return t * ((t >> 12 | t >> 8) & 63 & t >> 4)
#def f(t): return t >> 3 | t << 2 & t | int(t + 5e3) >> 4 | t - 14 >> 5
#def f(t): return (t & t // 170 * 2) + t % 31 * 0.1
#def f(t): return t << 1 >> 1 ^ t * 3 | t >> 5 | t >> 3
#def f(t): return t * 9 & t >> 4 | t * 5 & t >> 7 | t * 3 & t // 1024
def div(a, b): return 0 if b == 0 else a // b
out = [int(f(t)) & 0xff for t in range(sec(5))]
display(Audio(out, rate=8000))
# + [markdown] slideshow={"slide_type": "slide"}
# # Спасибо за внимание!
#
# Проект: https://github.com/true-grue/algomusic
# (используйте algomusic.py и examples.py, если не хотите работать в Jupyter, для запуска достаточно иметь Python 3)
#
# Обсудить компьютерную музыку с автором: <EMAIL>
| algomusic_part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: aoc
# language: python
# name: aoc
# ---
# +
class Point:
def __init__(self, x, y):
self.x = x
self.y = y
def manhatten_distance(self, other):
return abs(self.x - other.x) + abs(self.y - other.y)
def __eq__(self, other):
return self.x == other.x and self.y == other.y
def __hash__(self):
return hash((self.x, self.y))
def __repr__(self):
return f'{self.__class__.__name__}({self.x}, {self.y})'
class Segment:
def __init__(self, start, end, points):
self.start = start
self.end = end
self.points = points
def signal_delay(self, destination):
delay = 0
for point in self.points:
if point == destination:
break
delay += 1
return delay
@classmethod
def from_instruction(cls, start, instruction):
direction, length = instruction[0], int(instruction[1:])
if direction == 'R':
points = tuple(Point(start.x+i, start.y) for i in range(length+1))
if direction == 'L':
points = tuple(Point(start.x-i, start.y) for i in range(length+1))
if direction == 'U':
points = tuple(Point(start.x, start.y+i) for i in range(length+1))
if direction == 'D':
points = tuple(Point(start.x, start.y-i) for i in range(length+1))
return cls(start=points[0], end=points[-1], points=points)
class Wire:
def __init__(self, origin, segments):
self.origin = origin
self.segments = segments
self.points = set(point for segment in segments for point in segment.points)
def intersection(self, other, ignore_origin=True):
shared_points = self.points.intersection(other.points)
if ignore_origin and self.origin in shared_points:
shared_points.remove(self.origin)
return shared_points
def signal_delay(self, destination):
delay = 0
for segment in self.segments:
delay += segment.signal_delay(destination)
if destination in segment.points:
break
# the end point of one segment is also the start point
# of the next segment, so we remove the end point of
# each segment that went through a full iteration
delay -= 1
return delay
@classmethod
def from_instructions(cls, instructions, origin=Point(0,0)):
start = origin
segments = []
for instruction in instructions:
segment = Segment.from_instruction(start, instruction)
segments.append(segment)
start = segment.end
return cls(origin, segments)
# +
test_instructions_1 = ["R75", "D30", "R83", "U83", "L12", "D49", "R7", "U7", "L72"]
test_instructions_2 = ["U62", "R66", "U55", "R34", "D71", "R55", "D58", "R83"]
test_wire_1 = Wire.from_instructions(test_instructions_1, origin)
test_wire_2 = Wire.from_instructions(test_instructions_2, origin)
test_intersections = test_wire_1.intersection(test_wire_2, ignore_origin=True)
test_signal_delays = [
test_wire_1.signal_delay(intersection) + test_wire_2.signal_delay(intersection)
for intersection in test_intersections
]
print(sorted(test_signal_delays))
# +
from pathlib import Path
instruction_1, instruction_2 = Path('./input.txt').read_text().split()
instruction_1 = instruction_1.split(',')
instruction_2 = instruction_2.split(',')
origin = Point(0, 0)
wire_1 = Wire.from_instructions(instruction_1, origin)
wire_2 = Wire.from_instructions(instruction_2, origin)
# +
intersections = wire_1.intersection(wire_2, ignore_origin=True)
signal_delays = [
wire_1.signal_delay(intersection) + wire_2.signal_delay(intersection)
for intersection in intersections
]
print(sorted(signal_delays))
| puzzles/day_3/part_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting to know qubits through QEC
#
# ### <NAME>, IBM Quantum
# ## Introduction
#
# Back in 2018 I worked as a quantum error correction researcher at the University of Basel. IBM had just put a 16 qubit device online, and I wanted to see how well it could implement the basics of QEC. So I ran repetition codes.
#
#
# * *<NAME> and <NAME> Phys. Rev. A 97, 052313 (2018)*
#
# The main aim was to run codes of varying sizes, to test the idea that logical errors are suppressed with increasing code size. Here are the results from that 16 qubit device, which at that time was called *ibmqx3*.
#
# <img src="images/ibmqx3-decay.png" alt="" width="700"/>
#
# Here the points show the probability of a logical error for different codes. The *partial decoding* is a simple majority voting of the final results of the code qubits. The *full decoding* takes the full set of results into account.
#
# The points here decrease in a manner that is a but bumpy, but largely linear. Note that the y axis is logarithmic. The results are therefore consistent with the logical error decreasing exponentially with code size.
#
#
#
#
# After a few months, the device was taken out of the fridge for a while and cooled down again. It's name changed to *ibmqx5* and its behaviour changed also.
#
# <img src="images/ibmqx5-decay.png" alt="" width="700"/>
#
# Note the uptick of the partial decoding, which shows that something is not right.
#
# To see what is going on, we can look at a different kind of graph. For each qubit, what is the probability that it ends up in the `1` state?
#
# For *ibmqx3*, the results looked like the following.
#
# <img src="images/ibmqx3-1.png" alt="" width="900"/>
#
# As more qubits are added in to the code, more get affected by the additional noise. But for those already in the code, the noise they experience appears to be the same. This is what we might expect from the repetition code, since the circuit experienced by any qubit is not affected by qubits far away.
#
# Now let's take a look at *ibmqx5* instead.
#
# <img src="images/ibmqx5-1.png" alt="" width="910"/>
#
# In going from `d=7` to `d=8`, qubits 13 and 14 are added to the code. But the effects go far beyond just these. Many qubits are hit with a huge amount of noise, all the way down to qubit 9.
#
# This is what causes the uptick seen earlier, where increasing code size does not lead to less logical errors. It shows that the method used to make this device is not good enough to support large-scale quantum error correction. Because of problems such as this, IBM rethought how they arranged their qubits. And it was independently discovered by me, an external researcher running repetition codes.
#
# So what remains for us to discover in 2021?
# ## Repetiton codes with Qiskit
# Now let's make some repetition codes with Qiskit. We'll need `QuantumCircuit` objects, and `QuantumRegister` and `ClassicalRegister` objects.
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister
# We'll also need too decide how big we want the code to be. Specifcially, the number of repetitions `d` and the number of syndrome measurement rounds `T`.
#
# To be able to see the circuits easily, we'll start with the minimum for `d`.
d = 3
# Let's also increase the value of `T` as we add measurement rounds to the circuit. So since we start with none...
T = 0
# We start with an empty circuit
qc = QuantumCircuit()
# then we add `d` code qubits
code_qubit = QuantumRegister(d, 'code_qubit')
qc.add_register( code_qubit )
# and `d-1` auxilliary qubits.
link_qubit = QuantumRegister(d-1, 'link_qubit')
qc.add_register( link_qubit )
# Then what we have is a `d=3`, `T=0` code for an encoded logical `0`. Which is still basically an empty circuit.
qc.draw(output='mpl')
# Now we can add the `cx` gates required for each syndrome measurement round. Each time this cell is run, it iterates `T` by 1.
# +
qc.barrier()
# for each link qubit, add a cx for the code qubit on one side
for j in range(d-1):
qc.cx(code_qubit[j],link_qubit[j])
# and then one for the other side
for j in range(d-1):
qc.cx(code_qubit[j+1],link_qubit[j])
# add a classical register to hold results from this round
link_bit = ClassicalRegister(d-1, 'round_'+str(T)+'_link_bit')
qc.add_register(link_bit)
# measure and then rest each link qubit to extract the syndrome
for j in range(d-1):
qc.measure(link_qubit[j],link_bit[j])
qc.reset(link_qubit[j])
# iterate the number of rounds
T += 1
qc.draw(output='mpl') # uncomment to draw circuit
# -
T
# Then we add the final readout of the code qubits.
# +
qc.barrier()
# add d bits for final readout of code qubits
code_bit = ClassicalRegister(d, 'code_bit')
qc.add_register(code_bit)
# measure code qubits
for j in range(d):
qc.measure(code_qubit[j],code_bit[j])
qc.draw(output='mpl') # uncomment to draw circuit
# -
# Now we have a code, we need a backend to run it on. We'll start with just a simulator.
#
# First we get the backend object for the simulator.
# +
from qiskit import Aer
backend_sim = Aer.get_backend('aer_simulator')
# -
# Then we use this to start the process of running the circuit, by creating the job object.
job = backend_sim.run(qc)
# Once the job has run, we extract the result object from this.
result = job.result()
# From this we can get our actual result: a histogram respresented as a Python dictionary.
counts = result.get_counts()
counts
# To see what happens when errors occur, let's manually insert some. Here are some circuits I made earlier.
# +
examples_qasm = ['OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nx code_qubit[0];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nx code_qubit[0];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nx code_qubit[0];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nx code_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nx code_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nx code_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\nx code_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nx link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n', 'OPENQASM 2.0;\ninclude "qelib1.inc";\nqreg code_qubit[3];\nqreg link_qubit[2];\ncreg round_0_link_bit[2];\ncreg round_1_link_bit[2];\ncreg code_bit[3];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nmeasure link_qubit[0] -> round_0_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_0_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\ncx code_qubit[0],link_qubit[0];\ncx code_qubit[1],link_qubit[1];\ncx code_qubit[1],link_qubit[0];\ncx code_qubit[2],link_qubit[1];\nx link_qubit[1];\nmeasure link_qubit[0] -> round_1_link_bit[0];\nreset link_qubit[0];\nmeasure link_qubit[1] -> round_1_link_bit[1];\nreset link_qubit[1];\nbarrier code_qubit[0],code_qubit[1],code_qubit[2],link_qubit[0],link_qubit[1];\nmeasure code_qubit[0] -> code_bit[0];\nmeasure code_qubit[1] -> code_bit[1];\nmeasure code_qubit[2] -> code_bit[2];\n']
examples = []
for qasm in examples_qasm:
examples.append(QuantumCircuit.from_qasm_str(qasm))
# -
# The errors seen here are the following.
#
# * 0: Initialization error on `code_qubit[0]`
# * 1: Error on `code_qubit[0]` while idle during first round of measurements.
# * 2: Error on `code_qubit[0]` while idle during second round of measurements.
# * 3: Initialization error on `code_qubit[1]`
# * 4: Error on `code_qubit[1]` while idle during first round of measurements.
# * 5: Error on `code_qubit[1]` while idle during second round of measurements.
# * 6: Error on `code_qubit[1]` between `cx` gates of first syndrome measurement.
# * 7: Measurement error on `link_qubit[1]` during first round of measurements.
# * 8: Measurement error on `link_qubit[1]` during second round of measurements.
#
# Let's take a look at each circuit
example = examples[0]
example.draw(output='mpl')
# and the effect that it has
job = backend_sim.run(example)
counts = job.result().get_counts()
print(counts)
# These results have two parts:
# * On the left, a `d`-bit string of the final code qubit readouts;
# * On the right, `T` lots of `d-1`-bit strings of the syndrome measurement readouts (with the rightmost being the first).
#
# The final code qubit readouts tell us two important things:
# * A final syndrome measuremement (inferred from comparing bit values);
# * The value of the logical bit.
#
# It therefore makes sense to process the result a bit to separate out these aspects.
# +
def process_1(output):
output = output.split(' ')
logical = output[0][0]+' '+output[0][-1]
final = ''
for j in range(d-1):
if output[0][j]==output[0][j+1]:
final += '0'
else:
final += '1'
output[0] = final
output = logical + ' ' + ' '.join(output)
return output
output = list(counts.keys())[0]
process_1(output)
# -
# Here the processed output again has two parts:
# * On the left, two opinions on what the logical bit value is (taken from the code qubits at each end of the code);
# * On the right `T+1` lots of `d-1`-bit strings of the syndrome measurement readouts (including the inferred one from the final readout.
#
# If you are wondering why we take two opinions on the logical readout: it's because it will make things nice later.
#
# As we saw in the lecture, it is not actually the syndrome values themselves that are useful, but it is their changes. So there's some more processing we can do, to output the syndrome changes instead of the values themselves.
# +
def process_2(output):
output = output.split(' ')
logical = output[0]
syndrome = output[1].split(' ') [::-1]
changes = [syndrome[0]]
for t in range(T):
change = ''
for j in range(d-1):
if syndrome[t][j]==syndrome[t+1][j]:
change += '0'
else:
change += '1'
changes.append(change)
output = logical + ' ' + ' '.join(changes[::-1])
return output
process_2(process_1(output))
# -
# Now let's take a look at the effect on all the examples.
# +
example = examples[0]
job = backend_sim.run(example)
counts = job.result().get_counts()
output = list(counts.keys())[0]
process_2(process_1(output))
# -
# Each inserted single qubit error results in a pair of syndrome changes, or one syndrome change and one logical value change. Either way, a unique pair of changes happen depending on exactly where and when the error happened. It is this that allows us to figure out what errors occurred, and how to correct them.
# ## Repetition Codes with `topological_codes`
#
# Though it was good for us to see how to create repetition codes manually, the process is automated using `topological_codes`. This was originally part of Qiskit Ignis, but is now an indepenent package.
#
# You can install it using
#
# ```
# pip install git+https://github.com/NCCR-SPIN/topological_codes.git
# ```
#
# either on the terminal, or using the cell below.
# +
# #!pip install git+https://github.com/NCCR-SPIN/topological_codes.git
# -
# Now we can import the class for repetition codes.
from topological_codes import RepetitionCode
# We can create a repetition code object using
# +
d = 3
T = 2
code = RepetitionCode(d,T,resets=True)
# -
# In this we have `code.circuit['0']` which contains the circuit for a stored logical `0`, like the one we just created.
code.circuit['0'].draw(output='mpl')
# And there's also one for a stored logical `1`.
code.circuit['1'].draw(output='mpl')
# Let's combine these into a list and run them in the same job
job = backend_sim.run([code.circuit['0'], code.circuit['1']])
# and then put the results together in the same dictionary
raw_results = {'0':job.result().get_counts(code.circuit['0']), '1':job.result().get_counts(code.circuit['1'])}
raw_results
# Since there are no errors, the results aren't very interesting. Just the standard code qubit readout on the left (all reflecting the logical value), and syndrome values on the right (detecting no errors).
#
# However, interesting or not, the code object allows us to process the results in the manner discussed earlier.
results = code.process_results(raw_results)
results
# To get some more interesting results, we can run the code on real quantum hardware. For that we need to load our IBMQ account.
from qiskit import IBMQ
provider = IBMQ.load_account()
# We'll use a suitable device: the 5 qubit `'ibmq_manila'`.
backend = provider.get_backend('ibmq_manila')
# With some fancy Jupyter tools, we can take a look at it and its properties.
import qiskit.tools.jupyter
backend
# Before running on this device, we need to transpile the circuit. This means rewriting it purely in terms of the gates that the device can directly use. For this we use Qiskit's `transpile` command.
# +
from qiskit import transpile
transpiled_circuits = {
bit:transpile(code.circuit[bit], backend=backend) for bit in ['0','1']
}
# -
# Let's take a look at how the circuit has changed, the original had the following number of each type of gate
code.circuit['0'].count_ops()
# For the transpiled circuit, these numbers have changed
transpiled_circuits['0'].count_ops()
# For some reason, the transpilation has increased the number of `cx` gates. To see why, we can use the `qubits` attribute of a circuit to see what order it considers its qubits to be in.
code.circuit['0'].qubits
# This means that it will initially try to use qubit 0 on the device as `link_qubit[0]`, qubit 1 as `link_qubit[1]`, qubit 3 as `code_qubit[0]` and so on. But this presents a problem. The circuit needs to do a `cx` between `code_qubit[0]` and `link_qubit[0]`, and so will try to do one between qubits 0 and 2 of the device. But the device does not allow such `cx` gates, and so it will have to do some tricks to create an effective `cx` gate between these two. It's these tricks that require more `cx` gates to be added.
#
# In this case, we don't actually require the tricks. In fact, we actively don't want them. We want to know exactly which qubit on the device is doing which job, so we can get some insight about their errors. We don't want strange transpilation confusing things.
#
# To solve this issue we can tell the transpiler which qubit should be doing which job. Our circuit will work fine if the qubits are given jobs as follows.
#
# ```
# qubit on device 0 ------------- 1 ------------- 2 ------------- 3 ------------- 4
#
# qubut in circuit code_qubit[0] - link_qubit[0] - code_qubit[1] - link_qubit[1] - code_qubit[2]
#
# ```
#
# So we make an `initial_layout` telling this information to the transpiler, listing the qubits on the device in the order of the `qubits` list above.
# +
initial_layout = [
1,
3,
0,
2,
4
]
transpiled_circuits = {
bit:transpile(code.circuit[bit], backend=backend, initial_layout=initial_layout) for bit in ['0','1']
}
transpiled_circuits['0'].count_ops()
# -
# Now we are ready to send the job.
job = backend.run([transpiled_circuits['0'], transpiled_circuits['1']], shots=8192)
# As soon as it is sent, we can get the job id.
job_id = job.job_id()
print(job_id)
# You won't need to use the next cell, if you are running the job live. But if you want to retreive a job that ran previously, you can use the job id.
# +
#job = backend.retrieve_job(job_id)
# -
# This won't work for you, though. You'll need to submit your own jobs and get your own job ids.
#
# Once the job has run, we can extract the raw results. Here we'll see more than just the simple case with no errors.
# +
raw_results = {
'0':job.result().get_counts(transpiled_circuits['0']),
'1':job.result().get_counts(transpiled_circuits['1'])
}
#raw_results # uncomment to see results
# -
# Let's process these results to see the syndrome changes.
results = code.process_results(raw_results)
#results # uncomment to see results
# Here we can see that the most likely outcome is the one that represents no errors. Then there are many that represent a single bit flip error somewhere, which we see from the fact that there are two syndrome changes. Then there are less likely cases with more than one error.
#
# One way we can process this data to get an idea of what is happening on the device is to calculate the logical error probability. This goes through all the different results, runs decoding to see what the most likely encoded value is for that result, and then compares that to what the encoded value is known to be.
#
# To run decoding, we need the decoder object for the code.
# +
from topological_codes import GraphDecoder
decoder = GraphDecoder(code)
# -
# Then we simply feed in the results to its `get_logical_prob` method.
decoder.get_logical_prob(results)
# Typically we find that the logical error probability is much worse for an encoded `1`, due to the fact that bit flips are much more likely to flip `1`s to `0`s than vice-versa.
#
# To get a more detailed picture of life on the device, we can also look at the probabiity for specific error processes. Though the decoder gives us a way to calculate this, let's do it manually instead. Specifically, we'll do it for all the example errors.
#
# First we'll collect the outputs that are a signature of those single errors.
# +
example_outputs = []
for example in examples:
job = backend_sim.run(example)
counts = job.result().get_counts()
output = list(counts.keys())[0]
processed_output = process_2(process_1(output))
example_outputs.append(processed_output)
example_outputs
# -
# Since we are just running small codes, it's not unlikely to see runs where only these single errors occur. By just looking at how many times these single errors occur, we can see their relative likelihood. Here we'll calculate these relative to the first example.
# +
def get_ratios(results, logical='0'):
probs = {}
for output in example_outputs:
if output in results[logical]:
probs[output] = results[logical][output]/results[logical][example_outputs[0]]
else:
probs[output] = 0.0
return probs
get_ratios(results)
# -
# Here each error process is referred to by a pair of coordinates, which aren't very easy to understand at first glance. These coordinates refer to where the two syndrome changes are for this particular type of error.
#
# To get a better idea of what these mean, we can go through all the example errors from earler and find their coordinates.
# Now let's look at a different implementation of the `d=3`, `T=2` code for comparison. This time we'll remove the `reset` gates. These are pretty slow things to have in the middle of a circuit. They aren't really needed either, since their job can be done in post-processing.
# +
code_no_resets = RepetitionCode(d,T,resets=False)
transpiled_circuits = {
bit:transpile(code_no_resets.circuit[bit], backend=backend, initial_layout=initial_layout) for bit in ['0','1']
}
# +
job = backend.run([transpiled_circuits['0'], transpiled_circuits['1']], shots=8192)
job.job_id()
# -
# Again, I can retrieve the job sent when preparing this notebook.
# +
#job = backend.retrieve_job(job_id)
# -
# And however we got the job object, we can then get the results.
# +
raw_results_no_resets = {
'0':job.result().get_counts(transpiled_circuits['0']),
'1':job.result().get_counts(transpiled_circuits['1'])
}
results_no_resets = code_no_resets.process_results(raw_results_no_resets)
# -
# Sometimes resusing a qubit immediately after measurement (as was done above) is not a good idea. When we remove the resets, we might therefore need to add a delay gate in instead. This is done by initializing the code in the following way.
# +
code_delay = RepetitionCode(d,T,resets=False,delay=1600)
transpiled_circuits = {
bit:transpile(code_delay.circuit[bit], backend=backend, initial_layout=initial_layout) for bit in ['0','1']
}
# +
job = backend.run([transpiled_circuits['0'], transpiled_circuits['1']], shots=8192)
job.job_id()
# -
# Again, let's retrieve the results.
# +
#job = backend.retrieve_job(job_id)
raw_results_delay = {
'0':job.result().get_counts(transpiled_circuits['0']),
'1':job.result().get_counts(transpiled_circuits['1'])
}
results_delay = code.process_results(raw_results_delay)
# -
# With all the results, let's first lookat the logical error probabilities.
print('Logical errors (resets):', GraphDecoder(code).get_logical_prob(results))
print('Logical errors (no resets):', GraphDecoder(code_no_resets).get_logical_prob(results_no_resets))
print('Logical errors (delay):', GraphDecoder(code_delay).get_logical_prob(results_delay))
# Now let's look at the probabilities for each of our example errors in each of these cases. As a reminder, here are our example errors.
#
# * 0: Initialization error on `code_qubit[0]`
# * 1: Error on `code_qubit[0]` while idle during first round of measurements.
# * 2: Error on `code_qubit[0]` while idle during second round of measurements.
# * 3: Initialization error on `code_qubit[1]`
# * 4: Error on `code_qubit[1]` while idle during first round of measurements.
# * 5: Error on `code_qubit[1]` while idle during second round of measurements.
# * 6: Error on `code_qubit[1]` between `cx` gates of first syndrome measurement.
# * 7: Measurement error on `link_qubit[1]` during first round of measurements.
# * 8: Measurement error on `link_qubit[1]` during second round of measurements.
for logical in ['0','1']:
print('\nResults for encoded '+logical+'\n')
probs = get_ratios(results,logical=logical)
probs_no_resets = get_ratios(results_no_resets,logical=logical)
probs_delay = get_ratios(results_delay,logical=logical)
print('Processed output resets no_resets delay')
for output in example_outputs:
print(output,' ', round(probs[output],1),' ',round(probs_no_resets[output],1),' ',round(probs_delay[output],1))
# Are the results what we expected? What secrets do we find out about these qubits?
# If you want to see the probabilities calculated by the decoder, use
decoder.get_error_probs(results)
# Here each error is described by a set of coordinates, which describe where the syndrome changes are in processed outputs (such as `'0 0 00 11 00'`, etc). To find the coordinates for these outputs, you can use
dec._string2nodes('0 0 00 11 00')
# Though, note that the flips of logical values are not included in these coordinates. So for the error that causes `'0 0 00 11 00'`, you shouldn't just use
dec._string2nodes('0 1 00 00 01')
# Instead ignore the `(0,1,0)` (the coordinate for the flip of the logical value) and just duplicate the other to get ((1, 2, 1), `(1, 2, 1))`.
# Now you have everything you need to know to understand them better than we do!
| main-presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CoNLL_2.ipynb
#
# This notebook contains the first part of the model training and analysis code from our CoNLL-2020 paper, ["Identifying Incorrect Labels in the CoNLL-2003 Corpus"](https://www.aclweb.org/anthology/2020.conll-1.16/).
#
# If you're new to the Text Extensions for Pandas library, we recommend that you start
# by reading through the notebook [`Analyze_Model_Outputs.ipynb`](https://github.com/CODAIT/text-extensions-for-pandas/blob/master/notebooks/Analyze_Model_Outputs.ipynb), which explains the
# portions of the library that we use in the notebooks in this directory.
#
# ### Summary
#
# This notebook aggregates the outputs of the 16 models submitted to the CoNLL-2003 competition. Then it compares these outputs to the corpus's gold-standard labels and identifies areas where there is a strong consensus between model outputs coupled with a disagreement with the corpus labels. Finally, it writes out CSV files containing ranked lists of potentially-incorrect labels.
# +
# INITIALIZATION BOILERPLATE
# Libraries
import os
import sys
import numpy as np
import pandas as pd
from typing import *
# And of course we need the text_extensions_for_pandas library itself.
try:
import text_extensions_for_pandas as tp
except ModuleNotFoundError as e:
raise Exception("text_extensions_for_pandas package not found on the Jupyter "
"kernel's path. Please either run:\n"
" ln -s ../../text_extensions_for_pandas .\n"
"from the directory containing this notebook, or use a Python "
"environment on which you have used `pip` to install the package.")
# Code shared among notebooks is kept in tp.cleaning, in this directory.
import text_extensions_for_pandas.cleaning
#constants
PROJECT_ROOT ='../..'
# -
# Download and cache the data set.
# NOTE: This data set is licensed for research use only. Be sure to adhere
# to the terms of the license when using this data set!
data_set_info = tp.io.conll.maybe_download_conll_data("outputs")
data_set_info
# +
# Load up the same gold standard data we used in Part 1.
gold_standard = tp.io.conll.conll_2003_to_dataframes(
data_set_info["test"], ["pos", "phrase", "ent"], [False, True, True])
gold_standard = [
df.drop(columns=["pos", "phrase_iob", "phrase_type"])
for df in gold_standard
]
# Dictionary from (collection, offset within collection) to dataframe
gold_standard_spans = {("test", i):
tp.io.conll.iob_to_spans(gold_standard[i])
for i in range(len(gold_standard))}
# +
# Load up the results from all 16 teams at once.
teams = ["bender", "carrerasa", "carrerasb", "chieu", "curran",
"demeulder", "florian", "hammerton", "hendrickx",
"klein", "mayfield", "mccallum", "munro", "whitelaw",
"wu", "zhang"]
# Read all the output files into one dataframe per <document, team> pair.
outputs = {
t: tp.io.conll.conll_2003_output_to_dataframes(
gold_standard, f"{PROJECT_ROOT}/resources/conll_03/ner/results/{t}/eng.testb")
for t in teams
} # Type: Dict[str, List[pd.DataFrame]]
# As an example of what we just loaded, show the token metadata for the
# "mayfield" team's model's output on document 3.
outputs["mayfield"][3]
# +
# Convert results from IOB2 tags to spans across all teams and documents
# See https://en.wikipedia.org/wiki/Inside%E2%80%93outside%E2%80%93beginning_(tagging) for details on IOB2 format.
output_spans = {
t: {("test", i): tp.io.conll.iob_to_spans(outputs[t][i])
for i in range(len(outputs[t]))}
for t in teams
} # Type: Dict[str, Dict[Tuple[str, int], pd.DataFrame]]
# As an example, show the first 10 spans that the "florian" team's model
# found on document 2.
output_spans["florian"][("test", 2)].head(10)
# +
# Use Pandas merge to find what spans match up exactly for each team's
# results.
# Unlike Part 1, we perform the join across all entity types, looking for
# matches of both the extracted span *and* the entity type label.
#
# Text Extensions for Pandas includes a utility function
# compute_accuracy_by_document() to makes this collection-level computation
# simpler.
stats = {
t: tp.io.conll.compute_accuracy_by_document(
gold_standard_spans, output_spans[t])
for t in teams
}
# Show the result quality statistics by document for the "carrerasa" team
stats["carrerasa"]
# -
# F1 for document 4 is looking a bit low. Is that just fluke, or is it
# part of a larger trend?
# In Part 1, we showed how to drill down to and examine "problem" documents.
# Since we have all this additional data, let's try a broader, more
# quantitative approach. We'll start by building up some more fine-grained
# data about congruence between the gold standard and the model outputs.
# Pandas' outer join will tell us what entities showed up just in the gold
# standard, just in the model output, or in both sets.
# For starters, let's do this just for the "carrerasa" team and document 4.
doc_id = ("test", 4)
(gold_standard_spans[doc_id]
.merge(output_spans["carrerasa"][doc_id], how="outer", indicator=True)
.sort_values("span"))
# +
# Repeat the analysis from the previous cell across all teams and documents.
# That is, perform an outer join between the gold standard spans dataframe
# for each document and the corresponding dataframe from each team.
def merge_span_sets(team: str,
gold_results: Dict[Tuple[str, int], pd.DataFrame],
results_by_team: Dict[str, Dict[Tuple[str, int], pd.DataFrame]]):
result = {} # Type: Dict[Tuple[str, int]: pd.DataFrame]
for k in gold_results.keys():
merged = gold_results[k].merge(results_by_team[team][k],
how="outer", indicator=True)
merged["gold"] = merged["_merge"].isin(("both", "left_only"))
merged[team] = merged["_merge"].isin(("both", "right_only"))
result[k] = merged[["span", "ent_type", "gold", team]]
return result
span_flags = {t: merge_span_sets(t, gold_standard_spans, output_spans) for t in teams} # Type: Dict[Tuple[str, int]: pd.DataFrame]
# -
# Now we have indicator variables for every extracted span, telling whether
# it was in the gold standard data set and/or in each of the team's results.
# For example, here are the first 5 spans for document 2 in the "carrerasa"
# team's results:
doc_id = ("test", 2)
span_flags["carrerasa"][doc_id].head(5)
# +
# Do an n-way merge of all those indicator variables across documents.
# This operation produces a single summary dataframe per document.
indicators = {} # Type: Dict[Tuple[str, int], pd.DataFrame]
for k in gold_standard_spans.keys():
result = gold_standard_spans[k]
for t in teams:
result = result.merge(span_flags[t][k], how="outer")
indicators[k] = result.fillna(False)
# Now we have a vector of indicator variables for every span extracted
# from every document across all the model outputs and the gold standard.
# For example, let's show the results for document 10:
doc_10 = ("test", 10)
indicators[doc_10]
# +
# If you look at the above dataframe, you can see that some entities
# ("RUGBY UNION", for example) are "easy", in that almost every entry
# found them correctly. Other entities, like "CAMPESE", are "harder",
# in that few of the entrants correctly identified them. Let's add
# a column that quantifies this "difficulty level" by counting how
# many teams found each true or false positive.
for df in indicators.values():
# Convert the teams' indicator columns into a single matrix of
# Boolean values, and sum the number of True values in each row.
vectors = df[df.columns[3:]].values
counts = np.count_nonzero(vectors, axis=1)
df["num_teams"] = counts
# Show the dataframe for document 10 again, this time with the new
# "num_teams" column at the far right.
indicators[doc_10]
# -
# Now we can rank the entities in document 10 by "difficulty", either as
# true positives for the models to find...
# (just for document 10 for the moment)
ind = indicators[doc_10].copy()
ind[ind["gold"] == True].sort_values("num_teams").head(10)
# ...or as false positives to avoid:
ind[ind["gold"] == False].sort_values("num_teams", ascending=False).head(10)
# +
# To get a better picture of what entities are "difficult", we need to look
# across the entire test set. Let's combine the dataframes in
# `indicators` into a single dataframe that covers all the documents.
# First we preprocess each dataframe to make it easier to combine.
to_stack = [
pd.DataFrame({
"fold": k[0], # Keys are (collection, offset) tuples
"doc_offset": k[1],
"span" : indicators[k]["span"],
"ent_type": indicators[k]["ent_type"],
"gold": indicators[k]["gold"],
"num_teams": indicators[k]["num_teams"]
})
for k in indicators.keys()
#for i in range(len(indicators))
]
# Then we concatenate all the preprocessed dataframes into a single dataframe.
counts = pd.concat(to_stack)
counts
# -
# Now we can pull out the most difficult entities across the entire test
# set.
# First, let's find the most difficult entities from the standpoint of recall:
# entities that are in the gold standard, but not in most results.
difficult_recall = counts[counts["gold"] == True].sort_values("num_teams").reset_index(drop=True)
difficult_recall.head(10)
# Hmm, everything is zero. How many entities were found by zero teams? One team?
(counts[counts["gold"] == True][["num_teams", "span"]]
.groupby("num_teams").count()
.rename(columns={"span": "count"}))
# Yikes! 140 entities in the test set were so hard to find, they
# were extracted by 0 teams.
# Let's go back and look at some of those 0-team entities in context:
difficult_recall["context"] = difficult_recall["span"].apply(lambda t: t.context())
pd.set_option('max_colwidth', 100)
difficult_recall.head(20)
# **Some of these entities are "difficult" because the test set contains incorrect labels.**
#
# For reference, there's a copy of the CoNLL labeling rules in this repository at
# [resources/conll_03/ner/annotation.txt](../resources/conll_03/ner/annotation.txt)
#
# There are 4 incorrect labels in this first set of 20:
# * `[3289, 3299): 'Full Light'` should be "Zywiec Full Light"
# * `[11, 19): 'Honda RV'` should be tagged `ORG`
# * `[1525, 1541): 'Consumer Project'` should be "Consumer Project on Technology" and should be tagged `ORG`
# * `[244, 255): 'McDonald 's'` should be tagged `MISC` (because it's an "adjective ... derived from a word which is ... organisation")
# +
# Let's look at the entities that are difficult from the perspective of
# precision: that is, in many models' results, but not in the gold standard.
difficult_precision = counts[counts["gold"] == False].sort_values("num_teams", ascending=False).reset_index(drop=True)
# Again, we can add some context to these spans:
difficult_precision["context"] = difficult_precision["span"].apply(lambda t: t.context())
difficult_precision.head(20)
# -
difficult_precision.loc[[16]]
# **As with the entities in `difficult_recall`, some of these entities in `difficult_precision` are "difficult" because the test set has missing and incorrect labels.**
#
# **13** of these first 20 "incorrect" results are due to missing and incorrect labels:
# * `[25, 32): 'BRITISH''` in document 202 should be tagged `MISC`.
# * `[1317, 1327): 'Portsmouth'` in document 207 should be tagged `ORG`, not `LOC`.
# * `[110, 118): 'Scottish'` in document 199 should be tagged `MISC`
# (or `[28, 53): 'SCOTTISH PREMIER DIVISION'` and
# `[110, 135): 'Scottish premier division'` should both be tagged `ORG`).
# * `[146, 163): 'Santiago Bernabeu'` in document 40 should be tagged `MISC`
# (because the "s" in `[146, 171): 'Santiago Bernabeu stadium'` is not capitalized).
# * `[239, 251): 'Philadelphia'` in document 223 should be tagged `ORG`, not `LOC`.
# * `[367, 376): 'Karlsruhe'` in document 36 should be tagged `ORG`, not `LOC`.
# * `[1003, 1011): 'Congress'` in document 100 should be tagged `ORG`
# (also, `[957, 964): 'Chilean' ==> MISC` should be replaced with
# `[957, 973): 'Chilean Congress' ==> ORG`).
# * `[420, 428): 'Freiburg'` in document 36 should be tagged `ORG`, not `LOC`.
# * In document 70, `[186, 211): 'New York Commodities Desk'`, not `[186, 206): 'New York Commodities'`, should be tagged `ORG`.
# * `[263, 271): 'St Louis'` in document 223 should be tagged `ORG`, not `LOC`.
# * `[788, 795): 'Antwerp'` in document 155 should be tagged `LOC`, not `ORG`.
# * In document 112, `[178, 191): '<NAME>r'`, not `[178, 188): '<NAME>'`, should be tagged `PER`.
# * `[274, 282): 'COLORADO'` in document 223 should be tagged `ORG`.
#
# Here's the gold standard data for document 155, for example.
# Note line 12.
doc_id = ("test", 155)
gold_standard_spans[doc_id][0:60]
# The above gold standard spans in context.
gold_standard_spans[doc_id]["span"].values
# +
# Repeat the steps from the previous cells using the dev set.
# This takes a while.
dev_gold_standard = tp.io.conll.conll_2003_to_dataframes(
data_set_info["dev"], ["pos", "phrase", "ent"], [False, True, True])
dev_gold_standard = [
df.drop(columns=["pos", "phrase_iob", "phrase_type"])
for df in dev_gold_standard
]
dev_gold_standard_spans = {
("dev", i): tp.io.conll.iob_to_spans(dev_gold_standard[i])
for i in range(len(dev_gold_standard))}
dev_outputs = {
t: tp.io.conll.conll_2003_output_to_dataframes(
dev_gold_standard, f"{PROJECT_ROOT}/resources/conll_03/ner/results/{t}/eng.testa")
for t in teams
} # Type: Dict[str, List[pd.DataFrame]]
dev_output_spans = {
t: {("dev", i): tp.io.conll.iob_to_spans(dev_outputs[t][i])
for i in range(len(dev_outputs[t]))}
for t in teams
} # Type: Dict[str, Dict[Tuple[str, int], pd.DataFrame]]
dev_span_flags = {t: merge_span_sets(t, dev_gold_standard_spans,
dev_output_spans)
for t in teams} # Type: Dict[Tuple[str, int]: pd.DataFrame]
dev_indicators = {} # Type: Dict[Tuple[str, int], pd.DataFrame]
for k in dev_gold_standard_spans.keys():
result = dev_gold_standard_spans[k]
for t in teams:
result = result.merge(dev_span_flags[t][k], how="outer")
dev_indicators[k] = result.fillna(False)
for df in dev_indicators.values():
# Convert the teams' indicator columns into a single matrix of
# Boolean values, and sum the number of True values in each row.
vectors = df[df.columns[3:]].values
nonzero_counts = np.count_nonzero(vectors, axis=1)
df["num_teams"] = nonzero_counts
dev_counts = pd.concat([
pd.DataFrame({
"fold": k[0], # Keys are (collection, offset) tuples
"doc_offset": k[1],
"span" : dev_indicators[k]["span"],
"ent_type": dev_indicators[k]["ent_type"],
"gold": dev_indicators[k]["gold"],
"num_teams": dev_indicators[k]["num_teams"]
})
for k in dev_indicators.keys()
])
# -
# How many teams found entities from the dev set that are in the gold standard?
(dev_counts[dev_counts["gold"] == True][["num_teams", "span"]]
.groupby("num_teams").count()
.rename(columns={"span": "count"}))
# How many teams found entities from the dev set that aren't in the gold standard?
(dev_counts[dev_counts["gold"] == False][["num_teams", "span"]]
.groupby("num_teams").count()
.rename(columns={"span": "count"}))
# Merge the results from the two folds
all_counts = pd.concat([counts, dev_counts])
all_counts.head()
all_counts.tail()
# Write out the results that are in the gold standard but not in
# few teams' outputs.
in_gold_to_write, not_in_gold_to_write = tp.cleaning.analysis.csv_prep(all_counts, "num_teams",gold_col_name='gold',doc_col_name='doc_offset')
in_gold_to_write
not_in_gold_to_write
# Write output files.
in_gold_to_write.to_csv("outputs/CoNLL_2_in_gold.csv", index=False)
not_in_gold_to_write.to_csv("outputs/CoNLL_2_not_in_gold.csv", index=False)
| tutorials/corpus/CoNLL_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import json
import sympy
% matplotlib inline
f = open('./exerc_phyton.txt')
V=np.genfromtxt(f,skip_header=6,delimiter='')
t=V[:,0]
print(t)
Raw=V[:,1]
print(Raw)
Noisy=V[:,2]
print(Noisy)
Acell=V[:,3]
deltat=t[1]-t[0]
print (deltat)
from numpy import diff
velocity=diff(Raw)/deltat
print(velocity)
dacell2=diff(velocity)/deltat
print(dacell2)
tamanhodacell2=np.size(dacell2)
novo_tempo=t[0:tamanhodacell2]
novo_aceleracao_medida=Acell[0:tamanhodacell2]
hfig,hax=plt.subplots(1,1,sharex = True, squeeze=True, figsize=(9,5))
plt.plot(t,Acell, label='Aceleração medida')
plt.plot(novo_tempo,dacell2,label='Aceleração calculada')
hax.legend(frameon=False)
hax.set_ylabel('Amplitude [m/$s^2$]')
hax.set_xlabel('Time[s]')
velocidadeNoisy=diff(Noisy)/deltat
Aceleracaonoisy2=diff(velocidadeNoisy)/deltat
hfig,hax=plt.subplots(1,1,sharex = True, squeeze=True, figsize=(9,5))
plt.plot(t,Acell, label='Aceleração medida')
plt.plot(novo_tempo,Aceleracaonoisy2,label='Aceleração calculada Noisy')
hax.legend(frameon=False)
hax.set_ylabel('Amplitude [m/$s^2$]')
hax.set_xlabel('Time[s]')
hfig,hax=plt.subplots(1,1,sharex = True, squeeze=True, figsize=(9,5))
plt.plot(novo_tempo,dacell2,label='Aceleração calculada')
plt.plot(novo_tempo,Aceleracaonoisy2,label='Aceleração calculada Noisy')
hax.legend(frameon=False)
hax.set_ylabel('Amplitude [m/$s^2$]')
hax.set_xlabel('Time[s]')
| exercicio_aula_5-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Iyvfu4R7k9X_"
from keras.layers import Dense
from keras.models import Sequential
# + id="ClxaOCOn96eC"
for i in range(0, 21):
# Train model for 1 epoch
h = model.fit(X_train, y_train, batch_size = 16, epochs = 1, verbose = 0)
if i%4==0:
# Get the output of the first layer
layer_output = inp_to_out([X_test])[0]
# Evaluate model accuracy for this epoch
test_accuracy = model.evaluate(X_test, y_test)[1]
# Plot 1st vs 2nd neuron output
plot()
# + id="qIvozwcG_eH-"
# + [markdown] id="sAaT45Dg_nLv"
# ### autoencoder (mnist dataset)
# + id="lDhL7e2O_eKm" colab={"base_uri": "https://localhost:8080/"} outputId="4bc1e64b-b9da-49c9-c139-c26d42544058"
# Start with a sequential model
autoencoder = Sequential()
# Add a dense layer with input the original image pixels and neurons the encoded representation
autoencoder.add(Dense(28, input_shape=(784, ), activation="relu"))
# Add an output layer with as many neurons as the orginal image pixels
autoencoder.add(Dense(784, activation = "sigmoid"))
# Compile your model with adadelta
autoencoder.compile(optimizer = 'adadelta', loss = 'binary_crossentropy')
# Summarize your model structure
autoencoder.summary()
# + id="QiwA60AF_0OC"
# + [markdown] id="p-i8xSDL_0sK"
# ### denoising
# + id="d0LCDxJaAuAe"
# + id="ksL266LA_2WX"
# Build your encoder by using the first layer of your autoencoder
encoder = Sequential()
encoder.add(autoencoder.layers[0])
# Encode the noisy images and show the encodings for your favorite number [0-9]
encodings = encoder.predict(X_test_noise)
show_encodings(encodings, number = 1)
# + id="QOcPNsD4BQjx"
# Build your encoder by using the first layer of your autoencoder
encoder = Sequential()
encoder.add(autoencoder.layers[0])
# Encode the noisy images and show the encodings for your favorite number [0-9]
encodings = encoder.predict(X_test_noise)
show_encodings(encodings, number = 1)
# Predict on the noisy images with your autoencoder
decoded_imgs = autoencoder.predict(X_test_noise)
# Plot noisy vs decoded images
compare_plot(X_test_noise, decoded_imgs)
# + [markdown] id="NoR6XhzvBoUu"
# ### CNNS
#
# Building a CNN model.
# (MNIST Dataset).
# + id="tzack609A-9H"
# Import the Conv2D and Flatten layers and instantiate model
from keras.layers import Conv2D,Flatten, Dense
model = Sequential()
# Add a convolutional layer of 32 filters of size 3x3
model.add(Conv2D(filters = 32, kernel_size = 3, input_shape = (28, 28, 1), activation = 'relu'))
# Add a convolutional layer of 16 filters of size 3x3
model.add(Conv2D(16, kernel_size = 3, activation = 'relu'))
# Flatten the previous layer output
model.add(Flatten())
# Add as many outputs as classes with softmax activation
model.add(Dense(10, activation = 'softmax'))
# + id="E2pHFLzgyD5p"
# Obtain a reference to the outputs of the first layer
first_layer_output = model.layers[0].output
# Build a model using the model's input and the first layer output
first_layer_model = Model(inputs = model.layers[0].input, outputs = first_layer_output)
# Use this model to predict on X_test
activations = first_layer_model.predict(X_test)
# Plot the activations of first digit of X_test for the 15th filter
axs[0].matshow(activations[0,:,:,14], cmap = 'viridis')
# Do the same but for the 18th filter now
axs[1].matshow(activations[0,:,:,17], cmap = 'viridis')
plt.show()
# + id="OFhCLPfmzryb"
# Import image and preprocess_input
import numpy as np
from keras.preprocessing import image
from keras.applications.resnet50 import preprocess_input
# Load the image with the right target size for your model
img = image.load_img('/content/dd.jpg', target_size=(224, 224))
# Turn it into an array
img_array = image.img_to_array(img)
# Expand the dimensions of the image, this is so that it fits the expected model input format
img_expanded = np.expand_dims(img_array, axis = 0)
# Pre-process the img in the same way original images were
img_ready = preprocess_input(img_expanded)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="JRAy9w2V3KTv" outputId="be8ca54a-8a1d-408b-bc66-f16a9318c91e"
import matplotlib.pyplot as plt
plt.imshow(img)
# + colab={"base_uri": "https://localhost:8080/"} id="qpITPhjH1M_o" outputId="0de1d461-d98f-4804-bf82-42393d97262d"
# Instantiate a ResNet50 model with 'imagenet' weights
from keras.applications.resnet50 import ResNet50, decode_predictions
model = ResNet50(weights='imagenet')
# Predict with ResNet50 on your already processed img
preds = model.predict(img_ready)
# Decode the first 3 predictions
print('Predicted:', decode_predictions(preds, top=3)[0])
# + [markdown] id="-psbfQkT3xJw"
# ### LSTM
#
# Long short term memory networks. (A type of RNN - Recurrent Neural Networks).
# + colab={"base_uri": "https://localhost:8080/"} id="wa2cKIXH3yzP" outputId="dc2ceed8-be48-4f37-d437-b50f35f8e808"
# Split text into an array of words
from keras.preprocessing.text import Tokenizer
text = 'it is not the strength of the body but the strength of the spirit it is useless to meet revenge with revenge it will heal nothing even the smallest person can change the course of history all we have to decide is what to do with the time that is given us the burned hand teaches best after that advice about fire goes to the heart'
words = text.split()
# Make sentences of 4 words each, moving one word at a time
sentences = []
for i in range(4, len(words)):
sentences.append(' '.join(words[i-4:i]))
# Instantiate a Tokenizer, then fit it on the sentences
tokenizer = Tokenizer()
tokenizer.fit_on_texts(sentences)
# Turn sentences into a sequence of numbers
sequences = tokenizer.texts_to_sequences(sentences)
print("Sentences: \n {} \n Sequences: \n {}".format(sentences[:5],sequences[:5]))
# + colab={"base_uri": "https://localhost:8080/"} id="GcSfc86F6QH6" outputId="5eba5e38-de6a-4c07-882e-50a481f74367"
# Import the Embedding, LSTM and Dense layer
from keras.layers import Dense, Embedding, LSTM
model = Sequential()
vocab_size = len(tokenizer.index_word) + 1
# Add an Embedding layer with the right parameters
model.add(Embedding(input_dim = vocab_size, input_length = 3, output_dim = 8, ))
# Add a 32 unit LSTM layer
model.add(LSTM(32))
# Add a hidden Dense layer of 32 units and an output layer of vocab_size with softmax
model.add(Dense(32, activation='relu'))
model.add(Dense(vocab_size, activation='softmax'))
model.summary()
# + id="wtYpvg3q7Zbu"
def predict_text(test_text, model = model):
if len(test_text.split()) != 3:
print('Text input should be 3 words!')
return False
# Turn the test_text into a sequence of numbers
test_seq = tokenizer.texts_to_sequences([test_text])
test_seq = np.array(test_seq)
# Use the model passed as a parameter to predict the next word
pred = model.predict(test_seq).argmax(axis = 1)[0]
# Return the word that maps to the prediction
return tokenizer.index_word[pred]
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="VCym9hMV8rdL" outputId="c8d0b407-b5b8-440b-9ac6-abcaaa5df285"
predict_text('meet revenge with')
# + colab={"base_uri": "https://localhost:8080/", "height": 74} id="FgvJQ1pI9HLY" outputId="d4ceba51-3842-434c-c9f8-f075b8c293c4"
predict_text('the course need')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="RoqO3uuc9Hfu" outputId="5aa5575c-76e9-4ff1-9b91-0ef929855ced"
predict_text('strength of the')
| keras/tensors_layers_autoencoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Zadanie 2
class PermissionError(Exception):
pass
age = int(input('Podaj wiek: '))
if age > 18:
raise PermissionError('Only for kids!')
# # Zadanie 3
try:
temperature = float(input('Podaj temperature w Kelvinach: '))
print(temperature)
except ValueError:
print('Invalid temperature!')
# # Zadanie 4
class InvalidTemperature(Exception):
pass
temperature = float(input('Podaj temperature w Kelvinach: '))
if temperature < 0:
raise InvalidTemperature('Cannot be less than 0!')
| basic/Exceptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="irEeFSG-_17e" colab_type="code" colab={}
# !pip install tensorflow-gpu
# + id="FD7q5Nkl0kvs" colab_type="code" colab={}
# !pip install deeppavlov
# + id="Jh_PW7dJ-o9x" colab_type="code" colab={}
# !python -m deeppavlov install bert_sentiment.json
# + id="eJsWV2ndHFRK" colab_type="code" colab={}
# !python -m deeppavlov install fasttext_sentiment.json
# + id="GX09UGLPEmmN" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="pjFIKhli8DkZ" colab_type="code" colab={}
from deeppavlov import train_model
# + id="c73N8OMY9Tnf" colab_type="code" colab={}
model = train_model('bert_sentiment.json', download=False)
# + id="HQlht50a80DN" colab_type="code" colab={}
model = train_model('elmo_sentiment.json')
# + id="V4qLxrWdDYNk" colab_type="code" colab={}
model = train_model('fasttext_sentiment.json', download=False)
# + id="NjqmXDApDDiM" colab_type="code" colab={}
# !zip -r /content/model_bert.zip /content/model
# + id="vxPuSm7w_zoW" colab_type="code" colab={}
# !cp model_bert.zip 'drive/My Drive/Colab Notebooks'
# + id="doT7W9-0Eit8" colab_type="code" colab={}
| notebooks/dl_classification.ipynb |